text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
```
import matplotlib.pyplot as plt
import sys
import os
import numpy as np
root_dir = os.path.abspath('./')
sys.path.append(root_dir)
sys.path.append(os.path.join(root_dir,'../DPMJET-III-gitlab'))
print root_dir
from impy.definitions import *
from impy.constants import *
from impy.kinematics import EventKinematics
from impy.common import impy_config, pdata
# AF: This is what the user interaction has to yield.
# It is the typical expected configuration that one
# wants to run (read pp-mode at energies not exceeding
# 7 TeV). If you want cosmic ray energies, this should
# be rather p-N at 10 EeV and lab frame (not yet defined).
event_kinematics = EventKinematics(
ecm= 7000 * GeV,
p1pdg=2212,
p2pdg=2212
# nuc2_prop=(12,6)
)
impy_config["user_frame"] = 'center-of-mass'
# impy_config['tau_stable'] = 1.
# impy_config['pi0_stable'] = False
generator = make_generator_instance(interaction_model_by_tag['URQMD34'])
make_decay_list = [211,321,111,2112, 310,130,13,-13,3122,-3212]
for pid in make_decay_list:
generator.set_stable(pid, stable=False)
generator.lib.stables.stabvec *= 0
generator.lib.stables.nstable = 0
generator.init_generator(event_kinematics)
# import IPython
# IPython.embed()
sibyll = make_generator_instance(interaction_model_by_tag['SIBYLL23C'])
sibyll.init_generator(event_kinematics)
n_bins = 30
bins = np.linspace(-1,1.,n_bins+1)
grid = 0.5*(bins[1:] + bins[:-1])
widths = (bins[1:] - bins[:-1])
urq_pions = np.zeros(n_bins)
urq_protons = np.zeros(n_bins)
eta_bins = np.linspace(-7,7,n_bins+1)
eta_grid = 0.5*(eta_bins[1:] + eta_bins[:-1])
wdths_eta = (eta_bins[1:] - eta_bins[:-1])
urq_charged_eta = np.zeros(n_bins)
n_events = 100
norm = 1./float(n_events)
# This
for event in generator.event_generator(event_kinematics, n_events):
event.filter_final_state_charged()
urq_protons += norm*np.histogram(event.xf[event.p_ids == 2212],bins=bins)[0]
urq_pions += norm*np.histogram(event.xf[np.abs(event.p_ids) == 211],bins=bins)[0]
urq_charged_eta += norm*np.histogram(event.eta,bins=eta_bins)[0]
sib_pions = np.zeros(n_bins)
sib_protons = np.zeros(n_bins)
sib_charged_eta = np.zeros(n_bins)
# This
for event in sibyll.event_generator(event_kinematics, n_events):
event.filter_final_state_charged()
sib_protons += norm*np.histogram(event.xf[event.p_ids == 2212],bins=bins)[0]
sib_pions += norm*np.histogram(event.xf[np.abs(event.p_ids) == 211],bins=bins)[0]
sib_charged_eta += norm*np.histogram(event.eta,bins=eta_bins)[0]
l=plt.plot(grid, urq_protons/widths)[0]
plt.plot(grid, sib_protons/widths,c=l.get_color(), ls='--',label='SIBYLL2.3c')
l=plt.plot(grid, urq_pions/widths)[0]
plt.plot(grid, sib_pions/widths,c=l.get_color(), ls='--',label='SIBYLL2.3c')
plt.ylim(1e-2,)
plt.semilogy()
l=plt.plot(eta_grid, urq_charged_eta/wdths_eta)[0]
plt.plot(eta_grid, sib_charged_eta/wdths_eta,c=l.get_color(), ls='--',label='SIBYLL2.3c')
plt.ylim(0,7)
# plt.semilogy()
event_kinematics = EventKinematics(
plab=1000. * GeV,
p1pdg=2212,
p2pdg=2212
# nuc1_prop=(4,2),
# nuc2_prop=(12,6)
)
generator.lib.options.ctoption[0] = 1
event_kinematics = EventKinematics(
plab=5. * GeV,
# p1pdg=2212,
# p2pdg=2212
nuc1_prop=(4,2),
nuc2_prop=(12,6)
)
n_bins = 30
# bins = np.linspace(-1,1.,n_bins+1)
bins = np.logspace(-1,1,n_bins+1)
grid = 0.5*(bins[1:] + bins[:-1])
widths = (bins[1:] - bins[:-1])
urq_pions = np.zeros(n_bins)
urq_protons = np.zeros(n_bins)
n_events = 1000
norm = 1./float(n_events)
for event in generator.event_generator(event_kinematics, n_events):
# event.filter_final_state_charged()
event.filter_final_state_charged()
urq_protons += norm*np.histogram(event.en[event.p_ids == 2212],bins=bins)[0]
urq_pions += norm*np.histogram(event.en[np.abs(event.p_ids) == 211],bins=bins)[0]
plt.semilogx(grid, urq_pions)
# plt.plot(grid, urq_protons)
```
| github_jupyter |
# <img src="https://github.com/JuliaLang/julia-logo-graphics/raw/master/images/julia-logo-color.png" height="100" /> _for Pythonistas_
> TL;DR: _Julia looks and feels a lot like Python, only much faster. It's dynamic, expressive, extensible, with batteries included, in particular for Data Science_.
This notebook is an **introduction to Julia for Python programmers**.
It will go through the most important Python features (such as functions, basic types, list comprehensions, exceptions, generators, modules, packages, and so on) and show you how to code them in Julia.
# Getting Started with Julia in Colab/Jupyter
You can either run this notebook in Google Colab, or using Jupyter on your own machine.
## Running on Google Colab
1. Work on a copy of this notebook: _File_ > _Save a copy in Drive_ (you will need a Google account). Alternatively, you can download the notebook using _File_ > _Download .ipynb_, then upload it to [Colab](https://colab.research.google.com/).
2. Execute the following cell (click on it and press Ctrl+Enter) to install Julia, IJulia (the Jupyter kernel for Julia) and other packages. You can update `JULIA_VERSION` and the other parameters, if you know what you're doing. Installation takes 2-3 minutes.
3. Reload this page (press Ctrl+R, or ⌘+R, or the F5 key) and continue to the _Checking the Installation_ section.
* _Note_: If your Colab Runtime gets reset (e.g., due to inactivity), repeat steps 2 and 3.
```
%%shell
set -e
#---------------------------------------------------#
JULIA_VERSION="1.6.0" # any version ≥ 0.7.0
JULIA_PACKAGES="IJulia BenchmarkTools PyCall PyPlot"
JULIA_PACKAGES_IF_GPU="CUDA"
JULIA_NUM_THREADS=4
#---------------------------------------------------#
if [ -n "$COLAB_GPU" ] && [ -z `which julia` ]; then
# Install Julia
JULIA_VER=`cut -d '.' -f -2 <<< "$JULIA_VERSION"`
echo "Installing Julia $JULIA_VERSION on the current Colab Runtime..."
BASE_URL="https://julialang-s3.julialang.org/bin/linux/x64"
URL="$BASE_URL/$JULIA_VER/julia-$JULIA_VERSION-linux-x86_64.tar.gz"
wget -nv $URL -O /tmp/julia.tar.gz # -nv means "not verbose"
tar -x -f /tmp/julia.tar.gz -C /usr/local --strip-components 1
rm /tmp/julia.tar.gz
# Install Packages
if [ "$COLAB_GPU" = "1" ]; then
JULIA_PACKAGES="$JULIA_PACKAGES $JULIA_PACKAGES_IF_GPU"
fi
for PKG in `echo $JULIA_PACKAGES`; do
echo "Installing Julia package $PKG..."
julia -e 'using Pkg; pkg"add '$PKG'; precompile;"' &> /dev/null
done
# Install kernel and rename it to "julia"
echo "Installing IJulia kernel..."
julia -e 'using IJulia; IJulia.installkernel("julia", env=Dict(
"JULIA_NUM_THREADS"=>"'"$JULIA_NUM_THREADS"'"))'
KERNEL_DIR=`julia -e "using IJulia; print(IJulia.kerneldir())"`
KERNEL_NAME=`ls -d "$KERNEL_DIR"/julia*`
mv -f $KERNEL_NAME "$KERNEL_DIR"/julia
echo ''
echo "Successfully installed `julia -v`!"
echo "Please reload this page (press Ctrl+R, ⌘+R, or the F5 key) then"
echo "jump to the 'Checking the Installation' section."
fi
```
## Running This Notebook Locally
If you prefer to run this notebook on your machine instead of Google Colab:
* Download this notebook (File > Download .ipynb)
* Install [Julia](https://julialang.org/downloads/)
* Run the following command in a terminal to install `IJulia` (the Jupyter kernel for Julia), and a few packages we will use:
```bash
julia -e 'using Pkg
pkg"add IJulia; precompile;"
pkg"add BenchmarkTools; precompile;"
pkg"add PyCall; precompile;"
pkg"add PyPlot; precompile;"'
```
* Next, go to the directory containing this notebook:
```julia
cd /path/to/notebook/directory
```
* Start Jupyter Notebook:
```bash
julia -e 'using IJulia; IJulia.notebook()'
```
Or replace `notebook()` with `jupyterlab()` if you prefer JupyterLab.
If you do not already have [Jupyter](https://jupyter.org/install) installed, IJulia will propose to install it. If you agree, it will automatically install a private Miniconda (just for Julia), and install Jupyter and Python inside it.
* Lastly, open this notebook and skip directly to the next section.
## Checking the Installation
The `versioninfo()` function should print your Julia version and some other info about the system (if you ever ask for help or file an issue about Julia, you should always provide this information).
```
versioninfo()
```
# Getting Help
To get help on any module, function, variable, or just about anything else, just type `?` followed by what you're interested in. For example:
```
?versioninfo
```
This works in interactive mode only: in Jupyter, Colab and in the Julia shell (called the REPL).
Here are a few more ways to get help and inspect objects in interactive mode:
|Julia|Python
|-----|------
|`?obj` | `help(obj)`
|`dump(obj)` | `print(repr(obj))`
|`names(FooModule)` | `dir(foo_module)`
|`methodswith(SomeType)` | `dir(SomeType)`
|`@which func` | `func.__module__`
|`apropos("bar")` | Search for `"bar"` in docstrings of all installed packages
|`typeof(obj)` | `type(obj)`
|`obj isa SomeType`<br />or<br />`isa(obj, SomeType)` | `isinstance(obj, SomeType)`
If you ever ask for help or file an issue about Julia, you should generally provide the output of `versioninfo()`.
And of course, you can also learn and get help here:
* Learning: https://julialang.org/learning/
* Documentation: https://docs.julialang.org/
* Questions & Discussions:
* https://discourse.julialang.org/
* http://julialang.slack.com/
* https://stackoverflow.com/questions/tagged/julia
# A First Look at Julia
This section will give you an idea of what Julia looks like and what some of its major qualities are: it's expressive, dynamic, flexible, and most of all, super fast.
## Estimating π
Let's write our first function. It will estimate π using the equation:
$π = 4 \times \left(1 - \dfrac{1}{3} + \dfrac{1}{5} - \dfrac{1}{7} + \dfrac{1}{9}-\dfrac{1}{11}+\dots\right)$
There are much better ways to estimate π, but this one is easy to implement.
```
function estimate_pi(n)
s = 1.0
for i in 1:n
s += (isodd(i) ? -1 : 1) / (2i + 1)
end
4s
end
p = estimate_pi(100_000_000)
println("π ≈ $p")
println("Error is $(p - π)")
```
**Note**: syntax highlighting for Julia does not work (yet?) on Colab, but it does in Jupyter.
Compare this with the equivalent Python 3 code:
```python
# PYTHON
import math
def estimate_pi(n):
s = 1.0
for i in range(1, n + 1):
s += (-1 if i % 2 else 1) / (2 * i + 1)
return 4 * s
p = estimate_pi(100_000_000)
print(f"π ≈ {p}") # f-strings are available in Python 3.6+
print(f"Error is {p - math.pi}")
```
Pretty similar, right? But notice the small differences:
|Julia|Python
|-----|------
|`function` | `def`
|`for i in X`<br /> `...`<br />`end` | `for i in X:`<br /> `...`
|`1:n` | `range(1, n+1)`
|`cond ? a : b` | `a if cond else b`
|`2i + 1` | `2 * i + 1`
|`4s` | `return 4 * s`
|`println(a, b)` | `print(a, b, sep="")`
|`print(a, b)` | `print(a, b, sep="", end="")`
|`"$p"` | `f"{p}"`
|`"$(p - π)"` | `f"{p - math.pi}"`
This example shows that:
* Julia can be just as concise and readable as Python.
* Indentation in Julia is _not_ meaningful like it is in Python. Instead, blocks end with `end`.
* Many math features are built in Julia and need no imports.
* There's some mathy syntactic sugar, such as `2i` (but you can write `2 * i` if you prefer).
* In Julia, the `return` keyword is optional at the end of a function. The result of the last expression is returned (`4s` in this example).
* Julia loves Unicode and does not hesitate to use Unicode characters like `π`. However, there are generally plain-ASCII equivalents (e.g., `π == pi`).
## Typing Unicode Characters
Typing Unicode characters is easy: for latex symbols like π, just type `\pi<tab>`. For emojis like 😃, type `\:smiley:<tab>`.
This works in the REPL, in Jupyter, but unfortunately not in Colab (yet?). As a workaround, you can run the following code to print the character you want, then copy/paste it:
```
using REPL.REPLCompletions: latex_symbols, emoji_symbols
latex_symbols["\\pi"]
emoji_symbols["\\:smiley:"]
```
In Julia, `using Foo.Bar: a, b` corresponds to running `from foo.bar import a, b` in Python.
|Julia|Python
|-----|------
|`using Foo` | `from foo import *; import foo`
|`using Foo.Bar` | `from foo.bar import *; from foo import bar`
|`using Foo.Bar: a, b` | `from foo.bar import a, b`
|`using Foo: Bar` | `from foo import bar`
More on this later.
## Running Python code in Julia
Julia lets you easily run Python code using the `PyCall` module. We installed it earlier, so we just need to import it:
```
using PyCall
```
Now that we have imported `PyCall`, we can use the `pyimport()` function to import a Python module directly in Julia! For example, let's check which Python version we are using:
```
sys = pyimport("sys")
sys.version
```
In fact, let's run the Python code we discussed earlier (this will take about 15 seconds to run, because Python is so slow...):
```
py"""
import math
def estimate_pi(n):
s = 1.0
for i in range(1, n + 1):
s += (-1 if i % 2 else 1) / (2 * i + 1)
return 4 * s
p = estimate_pi(100_000_000)
print(f"π ≈ {p}") # f-strings are available in Python 3.6+
print(f"Error is {p - math.pi}")
"""
```
As you can see, running arbitrary Python code is as simple as using py-strings (`py"..."`). Note that py-strings are not part of the Julia language itself: they are defined by the `PyCall` module (we will see how this works later).
Unfortunately, Python's `print()` function writes to the standard output, which is not captured by Colab, so we can't see the output of this code. That's okay, we can look at the value of `p`:
```
py"p"
```
Let's compare this to the value we calculated above using Julia:
```
py"p" - p
```
Perfect, they are exactly equal!
As you can see, it's very easy to mix Julia and Python code. So if there's a module you really love in Python, you can keep using it as long as you want! For example, let's use NumPy:
```
np = pyimport("numpy")
a = np.random.rand(2, 3)
```
Notice that `PyCall` automatically converts some Python types to Julia types, including NumPy arrays. That's really quite convenient! Note that Julia supports multi-dimensional arrays (analog to NumPy arrays) out of the box. `Array{Float64, 2}` means that it's a 2-dimensional array of 64-bit floats.
`PyCall` also converts Julia arrays to NumPy arrays when needed:
```
exp_a = np.exp(a)
```
If you want to use some Julia variable in a py-string, for example `exp_a`, you can do so by writing `$exp_a` like this:
```
py"""
import numpy as np
result = np.log($exp_a)
"""
py"result"
```
If you want to keep using Matplotlib, it's best to use the `PyPlot` module (which we installed earlier), rather than trying to use `pyimport("matplotlib")`, as `PyPlot` provides a more straightforward interface with Julia, and it plays nicely with Jupyter and Colab:
```
using PyPlot
x = range(-5π, 5π, length=100)
plt.plot(x, sin.(x) ./ x) # we'll discuss this syntax in the next section
plt.title("sin(x) / x")
plt.grid("True")
plt.show()
```
That said, Julia has its own plotting libraries, such as the `Plots` library, which you may want to check out.
As you can see, Julia's `range()` function acts much like NumPy's `linspace()` function, when you use the `length` argument. However, it acts like Python's `range()` function when you use the `step` argument instead (except the upper bound is inclusive). Julia's `range()` function returns an object which behaves just like an array, except it doesn't actually use any RAM for its elements, it just stores the range parameters. If you want to collect all of the elements into an array, use the `collect()` function (similar to Python's `list()` function):
```
println(collect(range(10, 80, step=20)))
println(collect(10:20:80)) # 10:20:80 is equivalent to the previous range
println(collect(range(10, 80, length=5))) # similar to NumPy's linspace()
step = (80-10)/(5-1) # 17.5
println(collect(10:step:80)) # equivalent to the previous range
```
The equivalent Python code is:
```python
# PYTHON
print(list(range(10, 80+1, 20)))
# there's no short-hand for range() in Python
print(np.linspace(10, 80, 5))
step = (80-10)/(5-1) # 17.5
print([i*step + 10 for i in range(5)])
```
|Julia|Python
|-----|------
|`np = pyimport("numpy")` | `import numpy as np`
|`using PyPlot` | `from pylab import *`
|`1:10` | `range(1, 11)`
|`1:2:10`<br />or<br />`range(1, 11, 2)` | `range(1, 11, 2)`
|`1.2:0.5:10.3`<br />or<br />`range(1.2, 10.3, step=0.5)` | `np.arange(1.2, 10.3, 0.5)`
|`range(1, 10, length=3)` | `np.linspace(1, 10, 3)`
|`collect(1:5)`<br />or<br />`[i for i in 1:5]` | `list(range(1, 6))`<br />or<br />`[i for i in range(1, 6)]`
## Loop Fusion
Did you notice that we wrote `sin.(x) ./ x` (not `sin(x) / x`)? This is equivalent to `[sin(i) / i for i in x]`.
```
a = sin.(x) ./ x
b = [sin(i) / i for i in x]
@assert a == b
```
This is not just syntactic sugar: it's actually a very powerful Julia feature. Indeed, notice that the array only gets traversed once. Even if we chained more than two dotted operations, the array would still only get traversed once. This is called _loop fusion_.
In contrast, when using NumPy arrays, `sin(x) / x` first computes a temporary array containing `sin(x)` and then it computes the final array. Two loops and two arrays instead of one. NumPy is implemented in C, and has been heavily optimized, but if you chain many operations, it still ends up being slower and using more RAM than Julia.
However, all the extra dots can sometimes make the code a bit harder to read. To avoid that, you can write `@.` before an expression: every operation will be "dotted" automatically, like this:
```
a = @. sin(x) / x
b = sin.(x) ./ x
@assert a == b
```
**Note**: Julia's `@assert` statement starts with an `@` sign, just like `@.`, which means that they are macros. In Julia, macros are very powerful metaprogramming tools: a macro is evaluated at parse time, and it can inspect the expression that follows it and then transform it, or even replace it. In practice, you will often _use_ macros, but you will rarely _define_ your own. I'll come back to macros later.
## Julia is fast!
Let's compare the Julia and Python implementations of the `estimate_pi()` function:
```
@time estimate_pi(100_000_000);
```
To get a more precise benchmark, it's preferable to use the `BenchmarkTools` module. Just like Python's `timeit` module, it provides tools to benchmark code by running it multiple times. This provides a better estimate of how long each call takes:
```
using BenchmarkTools
@benchmark estimate_pi(100_000_000)
```
If this output is too verbose for you, simply use `@btime` instead:
```
@btime estimate_pi(100_000_000)
```
Now let's time the Python version. Since the call is so slow, we just run it once (it will take about 15 seconds):
```
py"""
from timeit import timeit
duration = timeit("estimate_pi(100_000_000)", number=1, globals=globals())
"""
py"duration"
```
It looks like Julia is close to 100 times faster than Python in this case! To be fair, `PyCall` does add some overhead, but even if you run this code in a separate Python shell, you will see that Julia crushes (pure) Python when it comes to speed.
So why is Julia so much faster than Python? Well, **Julia compiles the code on the fly as it runs it**.
Okay, let's summarize what we learned so far: Julia is a dynamic language that looks and feels a lot like Python, you can even execute Python code super easily, and pure Julia code runs much faster than pure Python code, because it is compiled on the fly. I hope this convinces you to read on!
Next, let's continue to see how Python's main constructs can be implemented in Julia.
# Numbers
```
i = 42 # 64-bit integer
f = 3.14 # 64-bit float
c = 3.4 + 4.5im # 128-bit complex number
bi = BigInt(2)^1000 # arbitrarily long integer
bf = BigFloat(1) / 7 # arbitrary precision
r = 15//6 * 9//20 # rational number
```
And the equivalent Python code:
```python
# PYTHON
i = 42
f = 3.14
c = 3.4 + 4.5j
bi = 2**1000 # integers are seemlessly promoted to long integers
from decimal import Decimal
bf = Decimal(1) / 7
from fractions import Fraction
r = Fraction(15, 6) * Fraction(9, 20)
```
Dividing integers gives floats, like in Python:
```
5 / 2
```
For integer division, use `÷` or `div()`:
```
5 ÷ 2
div(5, 2)
```
The `%` operator is the remainder, not the modulo like in Python. These differ only for negative numbers:
```
57 % 10
(-57) % 10
```
|Julia|Python
|-----|------
|`3.4 + 4.5im` | `3.4 + 4.5j`
|`BigInt(2)^1000` | `2**1000`
|`BigFloat(3.14)` | `from decimal import Decimal`<br />`Decimal(3.14)`
|`9//8` | `from fractions import Fraction`<br />`Fraction(9, 8)`
|`5/2 == 2.5` | `5/2 == 2.5`
|`5÷2 == 2`<br />or<br />`div(5, 2)` | `5//2 == 2`
|`57%10 == 7` | `57%10 == 7`
|`(-57)%10 == -7` | `(-57)%10 == 3`
# Strings
Julia strings use double quotes `"` or triple quotes `"""`, but not single quotes `'`:
```
s = "ångström" # Julia strings are UTF-8 encoded by default
println(s)
s = "Julia strings
can span
several lines\n\n
and they support the \"usual\" escapes like
\x41, \u5bb6, and \U0001f60a!"
println(s)
```
Use `repeat()` instead of `*` to repeat a string, and use `*` instead of `+` for concatenation:
```
s = repeat("tick, ", 10) * "BOOM!"
println(s)
```
The equivalent Python code is:
```python
# PYTHON
s = "tick, " * 10 + "BOOM!"
print(s)
```
Use `join(a, s)` instead of `s.join(a)`:
```
s = join([i for i in 1:4], ", ")
println(s)
```
You can also specify a string for the last join:
```
s = join([i for i in 1:4], ", ", " and ")
```
`split()` works as you might expect:
```
split(" one three four ")
split("one,,three,four!", ",")
occursin("sip", "Mississippi")
replace("I like coffee", "coffee" => "tea")
```
Triple quotes work a bit like in Python, but they also remove indentation and ignore the first line feed:
```
s = """
1. the first line feed is ignored if it immediately follows \"""
2. triple quotes let you use "quotes" easily
3. indentation is ignored
- up to left-most character
- ignoring the first line (the one with \""")
4. the final line feed it n̲o̲t̲ ignored
"""
println("<start>")
println(s)
println("<end>")
```
## String Interpolation
String interpolation uses `$variable` and `$(expression)`:
```
total = 1 + 2 + 3
s = "1 + 2 + 3 = $total = $(1 + 2 + 3)"
println(s)
```
This means you must escape the `$` sign:
```
s = "The car costs \$10,000"
println(s)
```
## Raw Strings
Raw strings use `raw"..."` instead of `r"..."`:
```
s = raw"In a raw string, you only need to escape quotes \", but not
$ or \. There is one exception, however: the backslash \
must be escaped if it's just before quotes like \\\"."
println(s)
s = raw"""
Triple quoted raw strings are possible too: $, \, \t, "
- They handle indentation and the first line feed like regular
triple quoted strings.
- You only need to escape triple quotes like \""", and the
backslash before quotes like \\".
"""
println(s)
```
## Characters
Single quotes are used for individual Unicode characters:
```
a = 'å' # Unicode code point (single quotes)
```
To be more precise:
* A Julia "character" represents a single Unicode code point (sometimes called a Unicode scalar).
* Multiple code points may be required to produce a single _grapheme_, i.e., something that readers would recognize as a single character. Such a sequence of code points is called a "Grapheme cluster".
For example, the character `é` can be represented either using the single code point `\u00E9`, or the grapheme cluster `e` + `\u0301`:
```
s = "café"
println(s, " has ", length(s), " code points")
s = "cafe\u0301"
println(s, " has ", length(s), " code points")
for c in "cafe\u0301"
display(c)
end
```
Julia represents any individual character like `'é'` using 32-bits (4 bytes):
```
sizeof('é')
```
But strings are represented using the UTF-8 encoding. In this encoding, code points 0 to 127 are represented using one byte, but any code point above 127 is represented using 2 to 6 bytes:
```
sizeof("a")
sizeof("é")
sizeof("家")
sizeof("🏳️🌈") # this is a grapheme with 4 code points of 4 + 3 + 3 + 4 bytes
[sizeof(string(c)) for c in "🏳️🌈"]
```
You can iterate through graphemes instead of code points:
```
using Unicode
for g in graphemes("e\u0301🏳️🌈")
println(g)
end
```
## String Indexing
Characters in a string are indexed based on the position of their starting byte in the UTF-8 representation. For example, the character `ê` in the string `"être"` is located at index 1, but the character `'t'` is located at index 3, since the UTF-8 encoding of `ê` is 2 bytes long:
```
s = "être"
println(s[1])
println(s[3])
println(s[4])
println(s[5])
```
If you try to get the character at index 2, you get an exception:
```
try
s[2]
catch ex
ex
end
```
By the way, notice the exception-handling syntax (we'll discuss exceptions later):
|Julia|Python
|-----|------
|`try`<br /> `...`<br />`catch ex`<br /> `...`<br />`end`|`try`<br /> `...`<br />`except Exception as ex`<br /> `...`<br />`end`
You can get a substring easily, using valid character indices:
```
s[1:3]
```
You can iterate through a string, and it will return all the code points:
```
for c in s
println(c)
end
```
Or you can iterate through the valid character indices:
```
for i in eachindex(s)
println(i, ": ", s[i])
end
```
Benefits of representing strings as UTF-8:
* All Unicode characters are supported.
* UTF-8 is fairly compact (at least for Latin scripts).
* It plays nicely with C libraries which expect ASCII characters only, since ASCII characters correspond to the Unicode code points 0 to 127, which UTF-8 encodes exactly like ASCII.
Drawbacks:
* UTF-8 uses a variable number of bytes per character, which makes indexing harder.
* However, If the language tried to hide this by making `s[5]` search for the 5th character from the start of the string, then code like `for i in 1:length(s); s[i]; end` would be unexpectedly inefficient, since at each iteration there would be a search from the beginning of the string, leading to O(_n_<sup>2</sup>) performance instead of O(_n_).
```
findfirst(isequal('t'), "être")
findlast(isequal('p'), "Mississippi")
findnext(isequal('i'), "Mississippi", 2)
findnext(isequal('i'), "Mississippi", 2 + 1)
findprev(isequal('i'), "Mississippi", 5 - 1)
```
Other useful string functions: `ncodeunits(str)`, `codeunit(str, i)`, `thisind(str, i)`, `nextind(str, i, n=1)`, `prevind(str, i, n=1)`.
## Regular Expressions
To create a regular expression in Julia, use the `r"..."` syntax:
```
regex = r"c[ao]ff?(?:é|ee)"
```
The expression `r"..."` is equivalent to `Regex("...")` except the former is evaluated at parse time, while the latter is evaluated at runtime, so unless you need to construct a Regex dynamically, you should prefer `r"..."`.
```
occursin(regex, "A bit more coffee?")
m = match(regex, "A bit more coffee?")
m.match
m.offset
m = match(regex, "A bit more tea?")
isnothing(m) && println("I suggest coffee instead")
regex = r"(.*)#(.+)"
line = "f(1) # nice comment"
m = match(regex, line)
code, comment = m.captures
println("code: ", repr(code))
println("comment: ", repr(comment))
m[2]
m.offsets
m = match(r"(?<code>.+)#(?<comment>.+)", line)
m[:comment]
replace("Want more bread?", r"(?<verb>more|some)" => s"a little")
replace("Want more bread?", r"(?<verb>more|less)" => s"\g<verb> and \g<verb>")
```
# Control Flow
## `if` statement
Julia's `if` statement works just like in Python, with a few differences:
* Julia uses `elseif` instead of Python's `elif`.
* Julia's logic operators are just like in C-like languages: `&&` means `and`, `||` means `or`, `!` means `not`, and so on.
```
a = 1
if a == 1
println("One")
elseif a == 2
println("Two")
else
println("Other")
end
```
Julia also has `⊻` for exclusive or (you can type `\xor<tab>` to get the ⊻ character):
```
@assert false ⊻ false == false
@assert false ⊻ true == true
@assert true ⊻ false == true
@assert true ⊻ true == false
```
Oh, and notice that `true` and `false` are all lowercase, unlike Python's `True` and `False`.
Since `&&` is lazy (like `and` in Python), `cond && f()` is a common shorthand for `if cond; f(); end`. Think of it as "_cond then f()_":
```
a = 2
a == 1 && println("One")
a == 2 && println("Two")
```
Similarly, `cond || f()` is a common shorthand for `if !cond; f(); end`. Think of it as "_cond else f()_":
```
a = 1
a == 1 || println("Not one")
a == 2 || println("Not two")
```
All expressions return a value in Julia, including `if` statements. For example:
```
a = 1
result = if a == 1
"one"
else
"two"
end
result
```
When an expression cannot return anything, it returns `nothing`:
```
a = 1
result = if a == 2
"two"
end
isnothing(result)
```
`nothing` is the single instance of the type `Nothing`:
```
typeof(nothing)
```
## `for` loops
You can use `for` loops just like in Python, as we saw earlier. However, it's also possible to create nested loops on a single line:
```
for a in 1:2, b in 1:3, c in 1:2
println((a, b, c))
end
```
The corresponding Python code would look like this:
```python
# PYTHON
from itertools import product
for a, b, c in product(range(1, 3), range(1, 4), range(1, 3)):
print((a, b, c))
```
The `continue` and `break` keywords work just like in Python. Note that in single-line nested loops, `break` will exit all loops, not just the inner loop:
```
for a in 1:2, b in 1:3, c in 1:2
println((a, b, c))
(a, b, c) == (2, 1, 1) && break
end
```
Julia does not support the equivalent of Python's `for`/`else` construct. You need to write something like this:
```
found = false
for person in ["Joe", "Jane", "Wally", "Jack", "Julia"] # try removing "Wally"
println("Looking at $person")
person == "Wally" && (found = true; break)
end
found || println("I did not find Wally.")
```
The equivalent Python code looks like this:
```python
# PYTHON
for person in ["Joe", "Jane", "Wally", "Jack", "Julia"]: # try removing "Wally"
print(f"Looking at {person}")
if person == "Wally":
break
else:
print("I did not find Wally.")
```
|Julia|Python
|-----|------
|`if cond1`<br /> `...`<br/>`elseif cond2`<br /> `...`<br/>`else`<br /> `...`<br/>`end` |`if cond1:`<br /> `...`<br/>`elif cond2:`<br /> `...`<br/>`else:`<br /> `...`
|`&&` | `and`
|`\|\|` | `or`
|`!` | `not`
|`⊻` (type `\xor<tab>`) | `^`
|`true` | `True`
|`false` | `False`
|`cond && f()` | `if cond: f()`
|`cond \|\| f()` | `if not cond: f()`
|`for i in 1:5 ... end` | `for i in range(1, 6): ...`
|`for i in 1:5, j in 1:6 ... end` | `from itertools import product`<br />`for i, j in product(range(1, 6), range(1, 7)):`<br /> `...`
|`while cond ... end` | `while cond: ...`
|`continue` | `continue`
|`break` | `break`
Now lets looks at data structures, starting with tuples.
# Tuples
Julia has tuples, very much like Python. They can contain anything:
```
t = (1, "Two", 3, 4, 5)
```
Let's look at one element:
```
t[1]
```
Hey! Did you see that? **Julia is 1-indexed**, like Matlab and other math-oriented programming languages, not 0-indexed like Python and most programming languages. I found it easy to get used to, and in fact I quite like it, but your mileage may vary.
Moreover, the indexing bounds are inclusive. In Python, to get the 1st and 2nd elements of a list or tuple, you would write `t[0:2]` (or just `t[:2]`), while in Julia you write `t[1:2]`.
```
t[1:2]
```
Note that `end` represents the index of the last element in the tuple. So you must write `t[end]` instead of `t[-1]`. Similarly, you must write `t[end - 1]`, not `t[-2]`, and so on.
```
t[end]
t[end - 1:end]
```
Like in Python, tuples are immutable:
```
try
t[2] = 2
catch ex
ex
end
```
The syntax for empty and 1-element tuples is the same as in Python:
```
empty_tuple = ()
one_element_tuple = (42,)
```
You can unpack a tuple, just like in Python (it's called "destructuring" in Julia):
```
a, b, c, d, e = (1, "Two", 3, 4, 5)
println("a=$a, b=$b, c=$c, d=$d, e=$e")
```
It also works with nested tuples, just like in Python:
```
(a, (b, c), (d, e)) = (1, ("Two", 3), (4, 5))
println("a=$a, b=$b, c=$c, d=$d, e=$e")
```
However, consider this example:
```
a, b, c = (1, "Two", 3, 4, 5)
println("a=$a, b=$b, c=$c")
```
In Python, this would cause a `ValueError: too many values to unpack`. In Julia, the extra values in the tuple are just ignored.
If you want to capture the extra values in the variable `c`, you need to do so explicitly:
```
t = (1, "Two", 3, 4, 5)
a, b = t[1:2]
c = t[3:end]
println("a=$a, b=$b, c=$c")
```
Or more concisely:
```
(a, b), c = t[1:2], t[3:end]
println("a=$a, b=$b, c=$c")
```
The corresponding Python code is:
```python
# PYTHON
t = (1, "Two", 3, 4, 5)
a, b, *c = t
print(f"a={a}, b={b}, c={c}")
```
## Named Tuples
Julia supports named tuples:
```
nt = (name="Julia", category="Language", stars=5)
nt.name
dump(nt)
```
The corresponding Python code is:
```python
# PYTHON
from collections import namedtuple
Rating = namedtuple("Rating", ["name", "category", "stars"])
nt = Rating(name="Julia", category="Language", stars=5)
print(nt.name) # prints: Julia
print(nt) # prints: Rating(name='Julia', category='Language', stars=5)
```
# Structs
Julia supports structs, which hold multiple named fields, a bit like named tuples:
```
struct Person
name
age
end
```
Structs have a default constructor, which expects all the field values, in order:
```
p = Person("Mary", 30)
p.age
```
You can create other constructors by creating functions with the same name as the struct:
```
function Person(name)
Person(name, -1)
end
function Person()
Person("no name")
end
p = Person()
```
This creates two constructors: the second calls the first, which calls the default constructor. Notice that you can create multiple functions with the same name but different arguments. We will discuss this later.
These two constructors are called "outer constructors", since they are defined outside of the definition of the struct. You can also define "inner constructors":
```
struct Person2
name
age
function Person2(name)
new(name, -1)
end
end
function Person2()
Person2("no name")
end
p = Person2()
```
This time, the outer constructor calls the inner constructor, which calls the `new()` function. This `new()` function only works in inner constructors, and of course it creates an instance of the struct.
When you define inner constructors, they replace the default constructor:
```
try
Person2("Bob", 40)
catch ex
ex
end
```
Structs usually have very few inner constructors (often just one), which do the heavy duty work, and the checks. Then they may have multiple outer constructors which are mostly there for convenience.
By default, structs are immutable:
```
try
p.name = "Someone"
catch ex
ex
end
```
However, it is possible to define a mutable struct:
```
mutable struct Person3
name
age
end
p = Person3("Lucy", 79)
p.age += 1
p
```
Structs look a lot like Python classes, with instance variables and constructors, but where are the methods? We will discuss this later, in the "Methods" section.
# Arrays
Let's create a small array:
```
a = [1, 4, 9, 16]
```
Indexing and assignments work as you would expect:
```
a[1] = 10
a[2:3] = [20, 30]
a
```
## Element Type
Since we used only integers when creating the array, Julia inferred that the array is only meant to hold integers (NumPy arrays behave the same way). Let's try adding a string:
```
try
a[3] = "Three"
catch ex
ex
end
```
Nope! We get a `MethodError` exception, telling us that Julia could not convert the string `"Three"` to a 64-bit integer (we will discuss exceptions later). If we want an array that can hold any type, like Python's lists can, we must prefix the array with `Any`, which is Julia's root type (like `object` in Python):
```
a = Any[1, 4, 9, 16]
a[3] = "Three"
a
```
Prefixing with `Float64`, or `String` or any other type works as well:
```
Float64[1, 4, 9, 16]
```
An empty array is automatically an `Any` array:
```
a = []
```
You can use the `eltype()` function to get an array's element type (the equivalent of NumPy arrays' `dtype`):
```
eltype([1, 4, 9, 16])
```
If you create an array containing objects of different types, Julia will do its best to use a type that can hold all the values as precisely as possible. For example, a mix of integers and floats results in a float array:
```
[1, 2, 3.0, 4.0]
```
This is similar to NumPy's behavior:
```python
# PYTHON
np.array([1, 2, 3.0, 4.0]) # => array([1., 2., 3., 4.])
```
A mix of unrelated types results in an `Any` array:
```
[1, 2, "Three", 4]
```
If you want to live in a world without type constraints, you can prefix all you arrays with `Any`, and you will feel like you're coding in Python. But I don't recommend it: the compiler can perform a bunch of optimizations when it knows exactly the type and size of the data the program will handle, so it will run much faster. So when you create an empty array but you know the type of the values it will contain, you might as well prefix it with that type (you don't have to, but it will speed up your program).
## Push and Pop
To append elements to an array, use the `push!()` function. By convention, functions whose name ends with a bang `!` may modify their arguments:
```
a = [1]
push!(a, 4)
push!(a, 9, 16)
```
This is similar to the following Python code:
```python
# PYTHON
a = [1]
a.append(4)
a.extend([9, 16]) # or simply a += [9, 16]
```
And `pop!()` works like in Python:
```
pop!(a)
```
Equivalent to:
```python
# PYTHON
a.pop()
```
There are many more functions you can call on an array. We will see later how to find them.
## Multidimensional Arrays
Importantly, Julia arrays can be multidimensional, just like NumPy arrays:
```
M = [1 2 3 4
5 6 7 8
9 10 11 12]
```
Another syntax for this is:
```
M = [1 2 3 4; 5 6 7 8; 9 10 11 12]
```
You can index them much like NumPy arrays:
```
M[2:3, 3:4]
```
You can transpose a matrix using the "adjoint" operator `'`:
```
M'
```
As you can see, Julia arrays are closer to NumPy arrays than to Python lists.
Arrays can be concatenated vertically using the `vcat()` function:
```
M1 = [1 2
3 4]
M2 = [5 6
7 8]
vcat(M1, M2)
```
Alternatively, you can use the `[M1; M2]` syntax:
```
[M1; M2]
```
To concatenate arrays horizontally, use `hcat()`:
```
hcat(M1, M2)
```
Or you can use the `[M1 M2]` syntax:
```
[M1 M2]
```
You can combine horizontal and vertical concatenation:
```
M3 = [9 10 11 12]
[M1 M2; M3]
```
Equivalently, you can call the `hvcat()` function. The first argument specifies the number of arguments to concatenate in each block row:
```
hvcat((2, 1), M1, M2, M3)
```
`hvcat()` is useful to create a single cell matrix:
```
hvcat(1, 42)
```
Or a column vector (i.e., an _n_×1 matrix = a matrix with a single column):
```
hvcat((1, 1, 1), 10, 11, 12) # a column vector with values 10, 11, 12
hvcat(1, 10, 11, 12) # equivalent to the previous line
```
Alternatively, you can transpose a row vector (but `hvcat()` is a bit faster):
```
[10 11 12]'
```
The REPL and IJulia call `display()` to print the result of the last expression in a cell (except when it is `nothing`). It is fairly verbose:
```
display([1, 2, 3, 4])
```
The `println()` function is more concise, but be careful not to confuse vectors, column vectors and row vectors (printed with commas, semi-colons and spaces, respectively):
```
println("Vector: ", [1, 2, 3, 4])
println("Column vector: ", hvcat(1, 1, 2, 3, 4))
println("Row vector: ", [1 2 3 4])
println("Matrix: ", [1 2 3; 4 5 6])
```
Although column vectors are printed as `[1; 2; 3; 4]`, evaluating `[1; 2; 3; 4]` will give you a regular vector. That's because `[x;y]` concatenates `x` and `y` vertically, and if `x` and `y` are scalars or vectors, you just get a regular vector.
|Julia|Python
|-----|------
|`a = [1, 2, 3]` | `a = [1, 2, 3]`<br />or<br />`import numpy as np`<br />`np.array([1, 2, 3])`
|`a[1]` | `a[0]`
|`a[end]` | `a[-1]`
|`a[2:end-1]` | `a[1:-1]`
|`push!(a, 5)` | `a.append(5)`
|`pop!(a)` | `a.pop()`
|`M = [1 2 3]` | `np.array([[1, 2, 3]])`
|`M = [1 2 3]'` | `np.array([[1, 2, 3]]).T`
|`M = hvcat(1, 1, 2, 3)` | `np.array([[1], [2], [3]])`
|`M = [1 2 3`<br /> `4 5 6]`<br />or<br />`M = [1 2 3; 4 5 6]` | `M = np.array([[1,2,3], [4,5,6]])`
|`M[1:2, 2:3]` | `M[0:2, 1:3]`
|`[M1; M2]` | `np.r_[M1, M2]`
|`[M1 M2]` | `np.c_[M1, M2]`
|`[M1 M2; M3]` | `np.r_[np.c_[M1, M2], M3]`
## Comprehensions
List comprehensions are available in Julia, just like in Python (they're usually just called "comprehensions" in Julia):
```
a = [x^2 for x in 1:4]
```
You can filter elements using an `if` clause, just like in Python:
```
a = [x^2 for x in 1:5 if x ∉ (2, 4)]
```
* `a ∉ b` is equivalent to `!(a in b)` (or `a not in b` in Python). You can type `∉` with `\notin<tab>`
* `a ∈ b` is equivalent to `a in b`. You can type it with `\in<tab>`
In Julia, comprehensions can contain nested loops, just like in Python:
```
a = [(i,j) for i in 1:3 for j in 1:i]
```
Here's the corresponding Python code:
```python
# PYTHON
a = [(i, j) for i in range(1, 4) for j in range(1, i+1)]
```
Julia comprehensions can also create multi-dimensional arrays (note the different syntax: there is only one `for`):
```
a = [row * col for row in 1:3, col in 1:5]
```
# Dictionaries
The syntax for dictionaries is a bit different than Python:
```
d = Dict("tree"=>"arbre", "love"=>"amour", "coffee"=>"café")
println(d["tree"])
println(get(d, "unknown", "pardon?"))
keys(d)
values(d)
haskey(d, "love")
"love" in keys(d) # this is slower than haskey()
```
The equivalent Python code is of course:
```python
d = {"tree": "arbre", "love": "amour", "coffee": "café"}
d["tree"]
d.get("unknown", "pardon?")
d.keys()
d.values()
"love" in d
"love" in d.keys()
```
Dict comprehensions work as you would expect:
```
d = Dict(i=>i^2 for i in 1:5)
```
Note that the items (aka "pairs" in Julia) are shuffled, since dictionaries are hash-based, like in Python (although Python sorts them by key for display).
You can easily iterate through the dictionary's pairs like this:
```
for (k, v) in d
println("$k maps to $v")
end
```
The equivalent code in Python is:
```python
# PYTHON
d = {i: i**2 for i in range(1, 6)}
for k, v in d.items():
print(f"{k} maps to {v}")
```
And you can merge dictionaries like this:
```
d1 = Dict("tree"=>"arbre", "love"=>"amour", "coffee"=>"café")
d2 = Dict("car"=>"voiture", "love"=>"aimer")
d = merge(d1, d2)
```
Notice that the second dictionary has priority in case of conflict (it's `"love" => "aimer"`, not `"love" => "amour"`).
In Python, this would be:
```python
# PYTHON
d1 = {"tree": "arbre", "love": "amour", "coffee": "café"}
d2 = {"car": "voiture", "love": "aimer"}
d = {**d1, **d2}
```
Or if you want to update the first dictionary instead of creating a new one:
```
merge!(d1, d2)
```
In Python, that's:
```python
# PYTHON
d1.update(d2)
```
In Julia, each pair is an actual `Pair` object:
```
p = "tree" => "arbre"
println(typeof(p))
k, v = p
println("$k maps to $v")
```
Note that any object for which a `hash()` method is implemented can be used as a key in a dictionary. This includes all the basic types like integers, floats, as well as string, tuples, etc. But it also includes arrays! In Julia, you have the freedom to use arrays as keys (unlike in Python), but make sure not to mutate these arrays after insertion, or else things will break! Indeed, the pairs will be stored in memory in a location that depends on the hash of the key at insertion time, so if that key changes afterwards, you won't be able to find the pair anymore:
```
a = [1, 2, 3]
d = Dict(a => "My array")
println("The dictionary is: $d")
println("Indexing works fine as long as the array is unchanged: ", d[a])
a[1] = 10
println("This is the dictionary now: $d")
try
println("Key changed, indexing is now broken: ", d[a])
catch ex
ex
end
```
However, it's still possible to iterate through the keys, the values or the pairs:
```
for pair in d
println(pair)
end
```
|Julia|Python
|-----|------
|`Dict("tree"=>"arbre", "love"=>"amour")` | `{"tree": "arbre", "love": "amour"}`
|`d["arbre"]` | `d["arbre"]`
|`get(d, "unknown", "default")` | `d.get("unknown", "default")`
|`keys(d)` | `d.keys()`
|`values(d)` | `d.values()`
|`haskey(d, k)` | `k in d`
|`Dict(i=>i^2 for i in 1:4)` | `{i: i**2 for i in 1:4}`
|`for (k, v) in d` | `for k, v in d.items():`
|`merge(d1, d2)` | `{**d1, **d2}`
|`merge!(d1, d2)` | `d1.update(d2)`
# Sets
Let's create a couple sets:
```
odd = Set([1, 3, 5, 7, 9, 11])
prime = Set([2, 3, 5, 7, 11])
```
The order of sets is not guaranteed, just like in Python.
Use `in` or `∈` (type `\in<tab>`) to check whether a set contains a given value:
```
5 ∈ odd
5 in odd
```
Both of these expressions are equivalent to:
```
in(5, odd)
```
Now let's get the union of these two sets:
```
odd ∪ prime
```
∪ is the union symbol, not a U. To type this character, type `\cup<tab>` (it has the shape of a cup). Alternatively, you can just use the `union()` function:
```
union(odd, prime)
```
Now let's get the intersection using the ∩ symbol (type `\cap<tab>`):
```
odd ∩ prime
```
Or use the `intersect()` function:
```
intersect(odd, prime)
```
Next, let's get the [set difference](https://en.wikipedia.org/wiki/Complement_(set_theory)#Relative_complement) and the [symetric difference](https://en.wikipedia.org/wiki/Symmetric_difference) between these two sets:
```
setdiff(odd, prime) # values in odd but not in prime
symdiff(odd, prime) # values that are not in the intersection
```
Lastly, set comprehensions work just fine:
```
Set([i^2 for i in 1:4])
```
The equivalent Python code is:
```python
# PYTHON
odds = {1, 3, 5, 7, 9, 11}
primes = {2, 3, 5, 7, 11}
5 in primes
odds | primes # union
odds.union(primes)
odds & primes # intersection
odds.intersection(primes)
odds - primes # set difference
odds.difference(primes)
odds ^ primes # symmetric difference
odds.symmetric_difference(primes)
{i**2 for i in range(1, 5)}
```
Note that you can store any hashable object in a `Set` (i.e., any instance of a type for which the `hash()` method is implemented). This includes arrays, unlike in Python. Just like for dictionary keys, you can add arrays to sets, but make sure not to mutate them after insertion.
|Julia|Python
|-----|------
|`Set([1, 3, 5, 7])` | `{1, 3, 5, 7}`
|`5 in odd` | `5 in odd`
|`Set([i^2 for i in 1:4])` | `{i**2 for i in range(1, 5)}`
|`odd ∪ primes` | `odd | primes`
|`union(odd, primes)` | `odd.union(primes)`
|`odd ∩ primes` | `odd & primes`
|`insersect(odd, primes)` | `odd.intersection(primes)`
|`setdiff(odd, primes)` | `odd - primes` or `odd.difference(primes)`
|`symdiff(odd, primes)` | `odd ^ primes` or `odd.symmetric_difference(primes)`
# Enums
To create an enum, use the `@enum` macro:
```
@enum Fruit apple=1 banana=2 orange=3
```
This creates the `Fruit` enum, with 3 possible values. It also binds the names to the values:
```
banana
```
Or you can get a `Fruit` instance using the value:
```
Fruit(2)
```
And you can get all the instances of the enum easily:
```
instances(Fruit)
```
|Julia|Python
|-----|------
|`@enum Fruit apple=1 banana=2 orange=3` | `from enum import Enum`<br />`class Fruit(Enum):`<br /> `APPLE = 1`<br /> `BANANA = 2`<br /> `ORANGE = 3`
| `Fruit(2) === banana` | `Fruit(2) is Fruit.BANANA`
| `instances(Fruit)` | `dir(Fruit)`
# Object Identity
In the previous example, `Fruit(2)` and `banana` refer to the same object, not just two objects that happen to be equal. You can verify using the `===` operator, which is the equivalent of Python's `is` operator:
```
banana === Fruit(2)
```
You can also check this by looking at their `objectid()`, which is the equivalent of Python's `id()` function:
```
objectid(banana)
objectid(Fruit(2))
a = [1, 2, 4]
b = [1, 2, 4]
@assert a == b # a and b are equal
@assert a !== b # but they are not the same object
```
|Julia|Python
|-----|------
|`a === b` | `a is b`
|`a !== b` | `a is not b`
|`objectid(obj)` | `id(obj)`
# Other Collections
For the Julia equivalent of Python's other collections, namely `defaultdict`, `deque`, `OrderedDict`, and `Counter`, check out these libraries:
* https://github.com/JuliaCollections/DataStructures.jl
* https://github.com/JuliaCollections/OrderedCollections.jl
* https://github.com/andyferris/Dictionaries.jl
Now let's looks at various iteration constructs.
# Iteration Tools
## Generator Expressions
Just like in Python, a generator expression resembles a list comprehension, but without the square brackets, and it returns a generator instead of a list. Here's a much shorter implementation of the `estimate_pi()` function using a generator expression:
```
function estimate_pi2(n)
4 * sum((isodd(i) ? -1 : 1)/(2i+1) for i in 0:n)
end
@assert estimate_pi(100) == estimate_pi2(100)
```
That's very similar to the corresponding Python code:
```python
# PYTHON
def estimate_pi2(n):
return 4 * sum((-1 if i%2==1 else 1)/(2*i+1) for i in range(n+1))
assert estimate_pi(100) == estimate_pi2(100)
```
## `zip`, `enumerate`, `collect`
The `zip()` function works much like in Python:
```
for (i, s) in zip(10:13, ["Ten", "Eleven", "Twelve"])
println(i, ": ", s)
end
```
Notice that the parentheses in `for (i, s)` are required in Julia, as opposed to Python.
The `enumerate()` function also works like in Python, except of course it is 1-indexed:
```
for (i, s) in enumerate(["One", "Two", "Three"])
println(i, ": ", s)
end
```
To pull the values of a generator into an array, use `collect()`:
```
collect(1:5)
```
A shorter syntax for that is:
```
[1:5;]
```
The equivalent Python code is:
```python
# PYTHON
list(range(1, 6))
```
## Generators
In Python, you can easily write a generator function to create an object that will behave like an iterator. For example, let's create a generator for the Fibonacci sequence (where each number is the sum of the two previous numbers):
```python
def fibonacci(n):
a, b = 1, 1
for i in range(n):
yield a
a, b = b, a + b
for f in fibonacci(10):
print(f)
```
This is also quite easy in Julia:
```
function fibonacci(n)
Channel() do ch
a, b = 1, 1
for i in 1:n
put!(ch, a)
a, b = b, a + b
end
end
end
for f in fibonacci(10)
println(f)
end
```
The `Channel` type is part of the API for tasks and coroutines. We'll discuss these later.
Now let's take a closer look at functions.
# Functions
## Arguments
Julia functions supports positional arguments and default values:
```
function draw_face(x, y, width=3, height=4)
println("x=$x, y=$y, width=$width, height=$height")
end
draw_face(10, 20, 30)
```
However, unlike in Python, positional arguments must not be named when the function is called:
```
try
draw_face(10, 20, width=30)
catch ex
ex
end
```
Julia also supports a variable number of arguments (called "varargs") using the syntax `arg...`, which is the equivalent of Python's `*arg`:
```
function copy_files(target_dir, paths...)
println("target_dir=$target_dir, paths=$paths")
end
copy_files("/tmp", "a.txt", "b.txt")
```
Keyword arguments are supported, after a semicolon `;`:
```
function copy_files2(paths...; confirm=false, target_dir)
println("paths=$paths, confirm=$confirm, $target_dir")
end
copy_files2("a.txt", "b.txt"; target_dir="/tmp")
```
Notes:
* `target_dir` has no default value, so it is a required argument.
* The order of the keyword arguments does not matter.
You can have another vararg in the keyword section. It corresponds to Python's `**kwargs`:
```
function copy_files3(paths...; confirm=false, target_dir, options...)
println("paths=$paths, confirm=$confirm, $target_dir")
verbose = options[:verbose]
println("verbose=$verbose")
end
copy_files3("a.txt", "b.txt"; target_dir="/tmp", verbose=true, timeout=60)
```
The `options` vararg acts like a dictionary (we will discuss dictionaries later). The keys are **symbols**, e.g., `:verbose`. Symbols are like strings, less flexible but faster. They are typically used as keys or identifiers.
|Julia|Python (3.8+ if `/` is used)
|-----|------
| `function foo(a, b=2, c=3)`<br /> `...`<br />`end`<br /><br />`foo(1, 2) # positional only` | `def foo(a, b=2, c=3, /):`<br /> `...`<br /><br />`foo(1, 2) # pos only because of /`
| `function foo(;a=1, b, c=3)`<br /> `...`<br />`end`<br /><br />`foo(c=30, b=2) # keyword only` | `def foo(*, a=1, b, c=3):`<br /> `...`<br /><br />`foo(c=30, b=2) # kw only because of *`
| `function foo(a, b=2; c=3, d)`<br /> `...`<br />`end`<br /><br />`foo(1; d=4) # pos only; then keyword only` | `def foo(a, b=2, /, *, c=3, d):`<br /> `...`<br /><br />`foo(1, d=4) # pos only then kw only`
| `function foo(a, b=2, c...)`<br /> `...`<br />`end`<br /><br />`foo(1, 2, 3, 4) # positional only` | `def foo(a, b=2, /, *c):`<br /> `...`<br /><br />`foo(1, 2, 3, 4) # positional only`
| `function foo(a, b=1, c...; d=1, e, f...)`<br /> `...`<br />`end`<br /><br />`foo(1, 2, 3, 4, e=5, x=10, y=20)`<br /> | `def foo(a, b=1, /, *c, d=1, e, **f):`<br /> `...`<br /><br />`foo(1, 2, 3, 4, e=5, x=10, y=20)`
## Concise Functions
In Julia, the following definition:
```
square(x) = x^2
```
is equivalent to:
```
function square(x)
x^2
end
```
For example, here's a shorter way to define the `estimate_pi()` function in Julia:
```
estimate_pi3(n) = 4 * sum((isodd(i) ? -1 : 1)/(2i+1) for i in 0:n)
```
To define a function on one line in Python, you need to use a `lambda` (but this is generally frowned upon, since the resulting function's name is `"<lambda>"`):
```python
# PYTHON
square = lambda x: x**2
assert square.__name__ == "<lambda>"
```
This leads us to anonymous functions.
## Anonymous Functions
Just like in Python, you can define anonymous functions:
```
map(x -> x^2, 1:4)
```
Here is the equivalent Python code:
```python
list(map(lambda x: x**2, range(1, 5)))
```
Notes:
* `map()` returns an array in Julia, instead of an iterator like in Python.
* You could use a comprehension instead: `[x^2 for x in 1:4]`.
|Julia|Python
|-----|------
|`x -> x^2` | `lambda x: x**2`
|`(x,y) -> x + y` | `lambda x,y: x + y `
|`() -> println("yes")` | `lambda: print("yes")`
In Python, lambda functions must be simple expressions. They cannot contain multiple statements. In Julia, they can be as long as you want. Indeed, you can create a multi-statement block using the syntax `(stmt_1; stmt_2; ...; stmt_n)`. The return value is the output of the last statement. For example:
```
map(x -> (println("Number $x"); x^2), 1:4)
```
This syntax can span multiple lines:
```
map(x -> (
println("Number $x");
x^2), 1:4)
```
But in this case, it's probably clearer to use the `begin ... end` syntax instead:
```
map(x -> begin
println("Number $x")
x^2
end, 1:4)
```
Notice that this syntax allows you to drop the semicolons `;` at the end of each line in the block.
Yet another way to define an anonymous function is using the `function (args) ... end` syntax:
```
map(function (x)
println("Number $x")
x^2
end, 1:4)
```
Lastly, if you're passing the anonymous function as the first argument to a function (as is the case in this example), it's usually much preferable to define the anonymous function immediately after the function call, using the `do` syntax, like this:
```
map(1:4) do x
println("Number $x")
x^2
end
```
This syntax lets you easily define constructs that feel like language extensions:
```
function my_for(func, collection)
for i in collection
func(i)
end
end
my_for(1:4) do i
println("The square of $i is $(i^2)")
end
```
In fact, Julia has a similar `foreach()` function.
The `do` syntax could be used to write a Domain Specific Language (DSL), for example an infrastructure automation DSL:
```
function spawn_server(startup_func, server_type)
println("Starting $server_type server")
server_id = 1234
println("Configuring server $server_id...")
startup_func(server_id)
end
# This is the DSL part
spawn_server("web") do server_id
println("Creating HTML pages on server $server_id...")
end
```
It's also quite nice for event-driven code:
```
handlers = []
on_click(handler) = push!(handlers, handler)
click(event) = foreach(handler->handler(event), handlers)
on_click() do event
println("Mouse clicked at $event")
end
on_click() do event
println("Beep.")
end
click((x=50, y=20))
click((x=120, y=10))
```
It can also be used to create context managers, for example to automatically close an object after it has been used, even if an exception is raised:
```
function with_database(func, name)
println("Opening connection to database $name")
db = "a db object for database $name"
try
func(db)
finally
println("Closing connection to database $name")
end
end
with_database("jobs") do db
println("I'm working with $db")
#error("Oops") # try uncommenting this line
end
```
The equivalent code in Python would look like this:
```python
# PYTHON
class Database:
def __init__(self, name):
self.name = name
def __enter__(self):
print(f"Opening connection to database {self.name}")
return f"a db object for database {self.name}"
def __exit__(self, type, value, traceback):
print(f"Closing connection to database {self.name}")
with Database("jobs") as db:
print(f"I'm working with {db}")
#raise Exception("Oops") # try uncommenting this line
```
Or you could use `contextlib`:
```python
from contextlib import contextmanager
@contextmanager
def database(name):
print(f"Opening connection to database {name}")
db = f"a db object for database {name}"
try:
yield db
finally:
print(f"Closing connection to database {name}")
with database("jobs") as db:
print(f"I'm working with {db}")
#raise Exception("Oops") # try uncommenting this line
```
```
```
## Piping
If you are used to the Object Oriented syntax `"a b c".upper().split()`, you may feel that writing `split(uppercase("a b c"))` is a bit backwards. If so, the piping operation `|>` is for you:
```
"a b c" |> uppercase |> split
```
If you want to pass more than one argument to some of the functions, you can use anonymous functions:
```
"a b c" |> uppercase |> split |> tokens->join(tokens, ", ")
```
The dotted version of the pipe operator works as you might expect, applying the _i_<sup>th</sup> function of the right array to the _i_<sup>th</sup> value in the left array:
```
[π/2, "hello", 4] .|> [sin, length, x->x^2]
```
## Composition
Julia also lets you compose functions like mathematicians do, using the composition operator ∘ (`\circ<tab>` in the REPL or Jupyter, but not Colab):
```
f = exp ∘ sin ∘ sqrt
f(2.0) == exp(sin(sqrt(2.0)))
```
# Methods
Earlier, we discussed structs, which look a lot like Python classes, with instance variables and constructors, but they did not contain any methods (just the inner constructors). In Julia, methods are defined separately, like regular functions:
```
struct Person
name
age
end
function greetings(greeter)
println("Hi, my name is $(greeter.name), I am $(greeter.age) years old.")
end
p = Person("Alice", 70)
greetings(p)
```
Since the `greetings()` method in Julia is not bound to any particular type, we can use it with any other type we want, as long as that type has a `name` and an `age` (i.e., if it quacks like a duck):
```
struct City
name
country
age
end
using Dates
c = City("Auckland", "New Zealand", year(now()) - 1840)
greetings(c)
```
You could code this the same way in Python if you wanted to:
```python
# PYTHON
class Person:
def __init__(self, name, age):
self.name = name
self.age = age
class City:
def __init__(self, name, country, age):
self.name = name
self.country = country
self.age = age
def greetings(greeter):
print(f"Hi there, my name is {greeter.name}, I am {greeter.age} years old.")
p = Person("Lucy", 70)
greetings(p)
from datetime import date
c = City("Auckland", "New Zealand", date.today().year - 1840)
greetings(c)
```
However, many Python programmers would use inheritance in this case:
```python
class Greeter:
def __init__(self, name, age):
self.name = name
self.age = age
def greetings(self):
print(f"Hi there, my name is {self.name}, I am {self.age} years old.")
class Person(Greeter):
def __init__(self, name, age):
super().__init__(name, age)
class City(Greeter):
def __init__(self, name, country, age):
super().__init__(name, age)
self.country = country
p = Person("Lucy", 70)
p.greetings()
from datetime import date
c = City("Auckland", "New Zealand", date.today().year - 1840)
c.greetings()
```
## Extending a Function
One nice thing about having a class hierarchy is that you can override methods in subclasses to get specialized behavior for each class. For example, in Python you could override the `greetings()` method like this:
```python
# PYTHON
class Developer(Person):
def __init__(self, name, age, language):
super().__init__(name, age)
self.language = language
def greetings(self):
print(f"Hi there, my name is {self.name}, I am {self.age} years old.")
print(f"My favorite language is {self.language}.")
d = Developer("Amy", 40, "Julia")
d.greetings()
```
Notice that the expression `d.greetings()` will call a different method if `d` is a `Person` or a `Developer`. This is called "polymorphism": the same method call behaves differently depending on the type of the object. The language chooses which actual method implementation to call, based on the type of `d`: this is called method "dispatch". More specifically, since it only depends on a single variable, it is called "single dispatch".
The good news is that Julia can do single dispatch as well:
```
struct Developer
name
age
language
end
function greetings(dev::Developer)
println("Hi, my name is $(dev.name), I am $(dev.age) years old.")
println("My favorite language is $(dev.language).")
end
d = Developer("Amy", 40, "Julia")
greetings(d)
```
Notice that the `dev` argument is followed by `::Developer`, which means that this method will only be called if the argument has that type.
We have **extended** the `greetings` **function**, so that it now has two different implementations, called **methods**, each for different argument types: namely, `greetings(dev::Developer)` for arguments of type `Developer`, and `greetings(greeter)` for values of any other type.
You can easily get the list of all the methods of a given function:
```
methods(greetings)
```
You can also get the list of all the methods which take a particular type as argument:
```
methodswith(Developer)
```
When you call the `greetings()` function, Julia automatically dispatches the call to the appropriate method, depending on the type of the argument. If Julia can determine at compile time what the type of the argument will be, then it optimizes the compiled code so that there's no choice to be made at runtime. This is called **static dispatch**, and it can significantly speed up the program. If the argument's type can't be determined at compile time, then Julia makes the choice at runtime, just like in Python: this is called **dynamic dispatch**.
## Multiple Dispatch
Julia actually looks at the types of _all_ the positional arguments, not just the first one. This is called **multiple dispatch**. For example:
```
multdisp(a::Int64, b::Int64) = 1
multdisp(a::Int64, b::Float64) = 2
multdisp(a::Float64, b::Int64) = 3
multdisp(a::Float64, b::Float64) = 4
multdisp(10, 20) # try changing the arguments to get each possible output
```
Julia always chooses the most specific method it can, so the following method will only be called if the first argument is neither an `Int64` nor a `Float64`:
```
multdisp(a::Any, b::Int64) = 5
multdisp(10, 20)
```
Julia will raise an exception if there is some ambiguity as to which method is the most specific:
```
ambig(a::Int64, b) = 1
ambig(a, b::Int64) = 2
try
ambig(10, 20)
catch ex
ex
end
```
To solve this problem, you can explicitely define a method for the ambiguous case:
```
ambig(a::Int64, b::Int64) = 3
ambig(10, 20)
```
So you can have polymorphism in Julia, just like in Python. This means that you can write your algorithms in a generic way, without having to know the exact types of the values you are manipulating, and it will work fine, as long as these types act in the general way you expect (i.e., if they "quack like ducks"). For example:
```
function how_can_i_help(greeter)
greetings(greeter)
println("How can I help?")
end
how_can_i_help(p) # called on a Person
how_can_i_help(d) # called on a Developer
```
## Calling `super()`?
You may have noticed that the `greetings(dev::Developer)` method could be improved, since it currently duplicates the implementation of the base method `greetings(greeter)`. In Python, you would get rid of this duplication by calling the base class's `greetings()` method, using `super()`:
```python
# PYTHON
class Developer(Person):
def __init__(self, name, age, language):
super().__init__(name, age)
self.language = language
def greetings(self):
super().greetings() # <== THIS!
print(f"My favorite language is {self.language}.")
d = Developer("Amy", 40, "Julia")
d.greetings()
```
In Julia, you can do something pretty similar, although you have to implement your own `super()` function, as it is not part of the language:
```
super(dev::Developer) = Person(dev.name, dev.age)
function greetings(dev::Developer)
greetings(super(dev))
println("My favorite language is $(dev.language).")
end
greetings(d)
```
However, this implementation creates a new `Person` instance when calling `super(dev)`, copying the `name` and `age` fields. That's okay for small objects, but it's not ideal for larger ones. Instead, you can explicitely call the specific method you want by using the `invoke()` function:
```
function greetings(dev::Developer)
invoke(greetings, Tuple{Any}, dev)
println("My favorite language is $(dev.language).")
end
greetings(d)
```
The `invoke()` function expects the following arguments:
* The first argument is the function to call.
* The second argument is the type of the desired method's arguments tuple: `Tuple{TypeArg1, TypeArg2, etc.}`. In this case we want to call the base function, which takes a single `Any` argument (the `Any` type is implicit when no type is specified).
* Lastly, it takes all the arguments to be passed to the method. In this case, there's just one: `dev`.
As you can see, we managed to get the same advantages Object-Oriented programming offers, without defining classes or using inheritance. This takes a bit of getting used to, but you might come to prefer this style of generic programming. Indeed, OO programming encourage you to bundle data and behavior together, but this is not always a good idea. Let's look at one example:
```python
# PYTHON
class Rectangle:
def __init__(self, height, width):
self.height = height
self.width = width
def area(self):
return self.height * self.width
class Square(Rectangle):
def __init__(self, length):
super().__init__(length, length)
```
It makes sense for the `Square` class to be a subclass of the `Rectangle` class, since a square **is a** special type of rectangle. It also makes sense for the `Square` class to inherit from all of the `Rectangle` class's behavior, such as the `area()` method. However, it does not really make sense for rectangles and squares to have the same memory representation: a `Rectangle` needs two numbers (`height` and `width`), while a `Square` only needs one (`length`).
It's possible to work around this issue like this:
```python
# PYTHON
class Rectangle:
def __init__(self, height, width):
self.height = height
self.width = width
def area(self):
return self.height * self.width
class Square(Rectangle):
def __init__(self, length):
self.length = length
@property
def width(self):
return self.length
@property
def height(self):
return self.length
```
That's better: now, each square is only represented using a single number. We've inherited the behavior, but not the data.
In Julia, you could code this like so:
```
struct Rectangle
width
height
end
width(rect::Rectangle) = rect.width
height(rect::Rectangle) = rect.height
area(rect) = width(rect) * height(rect)
struct Square
length
end
width(sq::Square) = sq.length
height(sq::Square) = sq.length
area(Square(5))
```
Notice that the `area()` function relies on the getters `width()` and `height()`, rather than directly on the fields `width` and `height`. This way, the argument can be of any type at all, as long as it has these getters.
## Abstract Types
One nice thing about the class hierarchy we defined in Python is that it makes it clear that a square **is a** kind of rectangle. Any new function you define that takes a `Rectangle` as an argument will automatically accept a `Square` as well, but no other non-rectangle type. In contrast, our `area()` function currently accepts anything at all.
In Julia, a concrete type like `Square` cannot extend another concrete type like `Rectangle`. However, any type can extend from an abstract type. Let's define some abstract types to create a type hierarchy for our `Square` and `Rectangle` types.
```
abstract type AbstractShape end
abstract type AbstractRectangle <: AbstractShape end # <: means "subtype of"
abstract type AbstractSquare <: AbstractRectangle end
```
The `<:` operator means "subtype of".
Now we can attach the `area()` function to the `AbstractRectangle` type, instead of any type at all:
```
area(rect::AbstractRectangle) = width(rect) * height(rect)
```
Now we can define the concrete types, as subtypes of `AbstractRectangle` and `AbstractSquare`:
```
struct Rectangle_v2 <: AbstractRectangle
width
height
end
width(rect::Rectangle_v2) = rect.width
height(rect::Rectangle_v2) = rect.height
struct Square_v2 <: AbstractSquare
length
end
width(sq::Square_v2) = sq.length
height(sq::Square_v2) = sq.length
```
In short, the Julian approach to type hierarchies looks like this:
* Create a hierarchy of abstract types to represent the concepts you want to implement.
* Write functions for these abstract types. Much of your implementation can be coded at that level, manipulating abstract concepts.
* Lastly, create concrete types, and write the methods needed to give them the behavior that is expected by the generic algorithms you wrote.
This pattern is used everywhere in Julia's standard libraries. For example, here are the supertypes of `Float64` and `Int64`:
```
Base.show_supertypes(Float64)
Base.show_supertypes(Int64)
```
Note: Julia implicitly runs `using Core` and `using Base` when starting the REPL. However, the `show_supertypes()` function is not exported by the `Base` module, thus you cannot access it by just typing `show_supertypes(Float64)`. Instead, you have to specify the module name: `Base.show_supertypes(Float64)`.
And here is the whole hierarchy of `Number` types:
```
function show_hierarchy(root, indent=0)
println(repeat(" ", indent * 4), root)
for subtype in subtypes(root)
show_hierarchy(subtype, indent + 1)
end
end
show_hierarchy(Number)
```
## Iterator Interface
You will sometimes want to provide a way to iterate over your custom types. In Python, this requires defining the `__iter__()` method which should return an object which implements the `__next__()` method. In Julia, you must define at least two functions:
* `iterate(::YourIteratorType)`, which must return either `nothing` if there are no values in the sequence, or `(first_value, iterator_state)`.
* `iterate(::YourIteratorType, state)`, which must return either `nothing` if there are no more values, or `(next_value, new_iterator_state)`.
For example, let's create a simple iterator for the Fibonacci sequence:
```
struct FibonacciIterator end
import Base.iterate
iterate(f::FibonacciIterator) = (1, (1, 1))
function iterate(f::FibonacciIterator, state)
new_state = (state[2], state[1] + state[2])
(new_state[1], new_state)
end
```
Now we can iterate over a `FibonacciIterator` instance:
```
for f in FibonacciIterator()
println(f)
f > 10 && break
end
```
## Indexing Interface
You can also create a type that will be indexable like an array (allowing syntax like `a[5] = 3`). In Python, this requires implementing the `__getitem__()` and `__setitem__()` methods. In Julia, you must implement the `getindex(A::YourType, i)`, `setindex!(A::YourType, v, i)`, `firstindex(A::YourType)` and `lastindex(A::YourType)` methods.
```
struct MySquares end
import Base.getindex, Base.firstindex
getindex(::MySquares, i) = i^2
firstindex(::MySquares) = 0
S = MySquares()
S[10]
S[begin]
getindex(S::MySquares, r::UnitRange) = [S[i] for i in r]
S[1:4]
```
For more details on these interfaces, and to learn how to build full-blown array types with broadcasting and more, check out [this page](https://docs.julialang.org/en/v1/manual/interfaces/).
## Creating a Number Type
Let's create a `MyRational` struct and try to make it mimic the built-in `Rational` type:
```
struct MyRational <: Real
num # numerator
den # denominator
end
MyRational(2, 3)
```
It would be more convenient and readable if we could type `2 ⨸ 3` to create a `MyRational`:
```
function ⨸(num, den)
MyRational(num, den)
end
2 ⨸ 3
```
I chose `⨸` because it's a symbol that Julia's parser treats as a binary operator, but which is otherwise not used by Julia (see the full [list of parsed symbols](https://github.com/JuliaLang/julia/blob/master/src/julia-parser.scm) and their priorities). This particular symbol will have the same priority as multiplication and division.
If you want to know how to type it and check that it is unused, type `?⨸` (copy/paste the symbol):
```
?⨸
```
Now let's make it possible to add two `MyRational` values. We want it to be possible for our `MyRational` type to be used in existing algorithms which rely on `+`, so we must create a new method for the `Base.+` function:
```
import Base.+
function +(r1::MyRational, r2::MyRational)
(r1.num * r2.den + r1.den * r2.num) ⨸ (r1.den * r2.den)
end
2 ⨸ 3 + 3 ⨸ 5
```
It's important to import `Base.+` first, or else you would just be defining a new `+` function in the current module (`Main`), which would not be called by existing algorithms.
You can easily implement `*`, `^` and so on, in much the same way.
Let's change the way `MyRational` values are printed, to make them look a bit nicer. For this, we must create a new method for the `Base.show(io::IO, x)` function:
```
import Base.show
function show(io::IO, r::MyRational)
print(io, "$(r.num) ⨸ $(r.den)")
end
2 ⨸ 3 + 3 ⨸ 5
```
We can expand the `show()` function so it can provide an HTML representation for `MyRational` values. This will be called by the `display()` function in Jupyter or Colab:
```
function show(io::IO, ::MIME"text/html", r::MyRational)
print(io, "<sup><b>$(r.num)</b></sup>⁄<sub><b>$(r.den)</b></sub>")
end
2 ⨸ 3 + 3 ⨸ 5
```
Next, we want to be able to perform any operation involving `MyRational` values and values of other `Number` types. For example, we may want to multiply integers and `MyRational` values. One option is to define a new method like this:
```
import Base.*
function *(r::MyRational, i::Integer)
(r.num * i) ⨸ r.den
end
2 ⨸ 3 * 5
```
Since multiplication is commutative, we need the reverse method as well:
```
function *(i::Integer, r::MyRational)
r * i # this will call the previous method
end
5 * (2 ⨸ 3) # we need the parentheses since * and ⨸ have the same priority
```
It's cumbersome to have to define these methods for every operation. There's a better way, which we will explore in the next two sections.
## Conversion
It is possible to provide a way for integers to be automatically converted to `MyRational` values:
```
import Base.convert
MyRational(x::Integer) = MyRational(x, 1)
convert(::Type{MyRational}, x::Integer) = MyRational(x)
convert(MyRational, 42)
```
The `Type{MyRational}` type is a special type which has a single instance: the `MyRational` type itself. So this `convert()` method only accepts `MyRational` itself as its first argument (and we don't actually use the first argument, so we don't even need to give it a name in the function declaration).
Now integers will be automatically converted to `MyRational` values when you assign them to an array whose element type if `MyRational`:
```
a = [2 ⨸ 3] # the element type is MyRational
a[1] = 5 # convert(MyRational, 5) is called automatically
push!(a, 6) # convert(MyRational, 6) is called automatically
println(a)
```
Conversion will also occur automatically in these cases:
* `r::MyRational = 42`: assigning an integer to `r` where `r` is a local variable with a declared type of `MyRational`.
* `s.b = 42` if `s` is a struct and `b` is a field of type `MyRational` (also when calling `new(42)` on that struct, assuming `b` is the first field).
* `return 42` if the return type is declared as `MyRational` (e.g., `function f(x)::MyRational ... end`).
However, there is no automatic conversion when calling functions:
```
function for_my_rationals_only(x::MyRational)
println("It works:", x)
end
try
for_my_rationals_only(42)
catch ex
ex
end
```
## Promotion
The `Base` functions `+`, `-`, `*`, `/`, `^`, etc. all use a "promotion" algorithm to convert the arguments to the appropriate type. For example, adding an integer and a float promotes the integer to a float before the addition takes place. These functions use the `promote()` function for this. For example, given several integers and a float, all integers get promoted to floats:
```
promote(1, 2, 3, 4.0)
```
This is why a sum of integers and floats results in a float:
```
1 + 2 + 3 + 4.0
```
The `promote()` function is also called when creating an array. For example, the following array is a `Float64` array:
```
a = [1, 2, 3, 4.0]
```
What about the `MyRational` type? Rather than create new methods for the `promote()` function, the recommended approach is to create a new method for the `promote_rule()` function. It takes two types and returns the type to convert to:
```
promote_rule(Float64, Int64)
```
Let's implement a new method for this function, to make sure that any subtype of the `Integer` type will be promoted to `MyRational`:
```
import Base.promote_rule
promote_rule(::Type{MyRational}, ::Type{T}) where {T <: Integer} = MyRational
```
This method definition uses **parametric types**: the type `T` can be any type at all, as long as it is a subtype of the `Integer` abstract type. If you tried to define the method `promote_rule(::Type{MyRational}, ::Type{Integer})`, it would expect the type `Integer` itself as the second argument, which would not work, since the `promote_rule()` function will usually be called with concrete types like `Int64` as its arguments.
Let's check that it works:
```
promote(5, 2 ⨸ 3)
```
Yep! Now whenever we call `+`, `-`, etc., with an integer and a `MyRational` value, the integer will get automatically promoted to a `MyRational` value:
```
5 + 2 ⨸ 3
```
Under the hood:
* this called `+(5, 2 ⨸ 3)`,
* which called the `+(::Number, ::Number)` method (thanks to multiple dispatch),
* which called `promote(5, 2 ⨸ 3)`,
* which called `promote_rule(Int64, MyRational)`,
* which called `promote_rule(::MyRational, ::T) where {T <: Integer}`,
* which returned `MyRational`,
* then the `+(::Number, ::Number)` method called `convert(MyRational, 5)`,
* which called `MyRational(5)`,
* which returned `MyRational(5, 1)`,
* and finally `+(::Number, ::Number)` called `+(MyRational(5, 1), MyRational(2, 3))`,
* which returned `MyRational(17, 3)`.
The benefit of this approach is that we only need to implement the `+`, `-`, etc. functions for pairs of `MyRational` values, not with all combinations of `MyRational` values and integers.
If your head hurts, it's perfectly normal. ;-) Writing a new type that is easy to use, flexible and plays nicely with existing types takes a bit of planning and work, but the point is that you will not write these every day, and once you have, they will make your life much easier.
Now let's handle the case where we want to execute operations with `MyRational` values and floats. In this case, we naturally want to promote the `MyRational` value to a float. We first need to define how to convert a `MyRational` value to any subtype of `AbstractFloat`:
```
convert(::Type{T}, x::MyRational) where {T <: AbstractFloat} = T(x.num / x.den)
```
This `convert()` works with any type `T` which is a subtype of `AbstractFloat`. It just computes `x.num / x.den` and converts the result to type `T`. Let's try it:
```
convert(Float64, 3 ⨸ 2)
```
Now let's define a `promote_rule()` method which will work for any type `T` which is a subtype of `AbstractFloat`, and which will give priority to `T` over `MyRational`:
```
promote_rule(::Type{MyRational}, ::Type{T}) where {T <: AbstractFloat} = T
promote(1 ⨸ 2, 4.0)
```
Now we can combine floats and `MyRational` values easily:
```
2.25 ^ (1 ⨸ 2)
```
## Parametric Types and Functions
Julia's `Rational` type is actually a **parametric type** which ensures that the numerator and denominator have the same type `T`, subtype of `Integer`. Here's a new version of our rational struct which enforces the same constraint:
```
struct MyRational2{T <: Integer}
num::T
den::T
end
```
To instantiate this type, we can specify the type `T`:
```
MyRational2{BigInt}(2, 3)
```
Alternatively, we can use the `MyRational2` type's default constructor, with two integers of the same type:
```
MyRational2(2, 3)
```
If we want to be able to construct a `MyRational2` with integers of different types, we must write an appropriate constructor which handles the promotion rule:
```
function MyRational2(num::Integer, den::Integer)
MyRational2(promote(num, den)...)
end
```
This constructor accepts two integers of potentially different types, and promotes them to the same type. Then it calls the default `MyRational2` constructor which expects two arguments of the same type. The syntax `f(args...)` is analog to Python's `f(*args)`.
Let's see if this works:
```
MyRational2(2, BigInt(3))
```
Great!
Note that all parametrized types such as `MyRational2{Int64}` or `MyRational2{BigInt}` are subtypes of `MyRational2`. So if a function accepts a `MyRational2` argument, you can pass it an instance of any specific, parametrized type:
```
function for_any_my_rational2(x::MyRational2)
println(x)
end
for_any_my_rational2(MyRational2{BigInt}(1, 2))
for_any_my_rational2(MyRational2{Int64}(1, 2))
```
A more explicit (but verbose) syntax for this function is:
```
function for_any_my_rational2(x::MyRational2{T}) where {T <: Integer}
println(x)
end
```
It's useful to think of types as sets. For example, the `Int64` type represents the set of all 64-bit integer values, so `42 isa Int64`:
* When `x` is an instance of some type `T`, it is an element of the set `T` represents, and `x isa T`.
* When `U` is a subtype of `V`, `U` is a subset of `V`, and `U <: V`.
The `MyRational2` type itself (without any parameter) represents the set of all values of `MyRational2{T}` for all subtypes `T` of `Integer`. In other words, it is the union of all the `MyRational2{T}` types. This is called a `UnionAll` type, and indeed the type `MyRational2` itself is an instance of the `UnionAll` type:
```
@assert MyRational2{BigInt}(2, 3) isa MyRational2{BigInt}
@assert MyRational2{BigInt}(2, 3) isa MyRational2
@assert MyRational2 === (MyRational2{T} where {T <: Integer})
@assert MyRational2{BigInt} <: MyRational2
@assert MyRational2 isa UnionAll
```
If we dump the `MyRational2` type, we can see that it is a `UnionAll` instance, with a parameter type `T`, constrained to a subtype of the `Integer` type (since the upper bound `ub` is `Integer`):
```
dump(MyRational2)
```
There's a lot more to learn about Julia types. When you feel ready to explore this in more depth, check out [this page](https://docs.julialang.org/en/v1.4/manual/types/). You can also take a look at the [source code of Julia's rationals](https://github.com/JuliaLang/julia/blob/master/base/rational.jl).
# Writing/Reading Files
The `do` syntax we saw earlier is helpful when using the `open()` function:
```
open("test.txt", "w") do f
write(f, "This is a test.\n")
write(f, "I repeat, this is a test.\n")
end
open("test.txt") do f
for line in eachline(f)
println("[$line]")
end
end
```
The `open()` function automatically closes the file at the end of the block. Notice that the line feeds `\n` at the end of each line are not returned by the `eachline()` function. So the equivalent Python code is:
```python
# PYTHON
with open("test.txt", "w") as f:
f.write("This is a test.\n")
f.write("I repeat, this is a test.\n")
with open("test.txt") as f:
for line in f.readlines():
line = line.rstrip("\n")
print(f"[{line}]")
```
Alternatively, you can read the whole file into a string:
```
open("test.txt") do f
s = read(f, String)
end
```
Or more concisely:
```
s = read("test.txt", String)
```
The Python equivalent is:
```python
# PYTHON
with open("test.txt") as f:
s = f.read()
```
# Exceptions
Julia's exceptions behave very much like in Python:
```
a = [1]
try
push!(a, 2)
#throw("Oops") # try uncommenting this line
push!(a, 3)
catch ex
println(ex)
push!(a, 4)
finally
push!(a, 5)
end
println(a)
```
The equivalent Python code is:
```python
# PYTHON
a = [1]
try:
a.append(2)
#raise Exception("Oops") # try uncommenting this line
a.append(3)
except Exception as ex:
print(ex)
a.append(4)
finally:
a.append(5)
print(a)
```
There is a whole hierarchy of standard exceptions which can be thrown, just like in Python. For example:
```
choice = 1 # try changing this value (from 1 to 4)
try
choice == 1 && open("/foo/bar/i_dont_exist.txt")
choice == 2 && sqrt(-1)
choice == 3 && push!(a, "Oops")
println("Everything worked like a charm")
catch ex
if ex isa SystemError
println("Oops. System error #$(ex.errnum) ($(ex.prefix))")
elseif ex isa DomainError
println("Oh no, I could not compute sqrt(-1)")
else
println("I got an unexpected error: $ex")
end
end
```
Compare this with Python's equivalent code:
```python
# PYTHON
choice = 3 # try changing this value (from 1 to 4)
try:
if choice == 1:
open("/foo/bar/i_dont_exist.txt")
if choice == 2:
math.sqrt(-1)
if choice == 3:
#a.append("Ok") # this would actually work
raise TypeError("Oops") # so let's fail manually
print("Everything worked like a charm")
except OSError as ex:
print(f"Oops. OS error (#{ex.errno} ({ex.strerror})")
except ValueError:
print("Oh no, I could not compute sqrt(-1)")
except Exception as ex:
print(f"I got an unexpected error: {ex}")
```
A few things to note here:
* Julia only allows a single `catch` block which handles all possible exceptions.
* `obj isa SomeClass` is a shorthand for `isa(obj, SomeClass)` which is equivalent to Python's `isinstance(obj, SomeClass)`.
|Julia|Python
|-----|------
|`try`<br /> `...`<br />`catch ex`<br /> `if ex isa SomeError`<br /> `...`<br /> `else`<br /> `...`<br /> `end`<br />`finally`<br /> `...`<br />`end` | `try:`<br /> `...`<br />`except SomeException as ex:`<br /> `...`<br />`except Exception as ex:`<br /> `...`<br />`finally:`<br /> `...`
|`throw any_value` | `raise SomeException(...)`
| `obj isa SomeType`<br />or<br /> `isa(obj, SomeType`) | `isinstance(obj, SomeType)`
Note that Julia does not support the equivalent of Python's `try / catch / else` construct. You need to write something like this:
```
catch_exception = true
try
println("Try something")
#error("ERROR: Catch me!") # try uncommenting this line
catch_exception = false
#error("ERROR: Don't catch me!") # try uncommenting this line
println("No error occurred")
catch ex
if catch_exception
println("I caught this exception: $ex")
else
throw(ex)
end
finally
println("The end")
end
println("After the end")
```
The equivalent Python code is shorter, but it's fairly uncommon:
```python
# PYTHON
try:
print("Try something")
raise Exception("Catch me!") # try uncommenting this line
except Exception as ex:
print(f"I caught this exception: {ex}")
else:
raise Exception("Don't catch me!") # try uncommenting this line
print("No error occured")
finally:
print("The end")
print("After the end")
```
# Docstrings
It's good practice to add docstrings to every function you export. The docstring is placed just _before_ the definition of the function:
```
"Compute the square of number x"
square(x::Number) = x^2
```
You can retrieve a function's docstring using the `@doc` macro:
```
@doc square
```
The docstring is displayed when asking for help:
```
?square
```
Docstrings follow the [Markdown format](https://en.wikipedia.org/wiki/Markdown#:~:text=Markdown%20is%20a%20lightweight%20markup,using%20a%20plain%20text%20editor.).
A typical docstring starts with the signature of the function, indented by 4 spaces, so it will get syntax highlighted as Julia code.
It also includes an `Examples` section with Julia REPL outputs:
```
"""
cube(x::Number)
Compute the cube of `x`.
# Examples
```julia-repl
julia> cube(5)
125
julia> cube(im)
0 - 1im
```
"""
cube(x) = x^3
```
Instead of using `julia-repl` code blocks for the examples, you can use `jldoctest` to mark these examples as doctests (similar to Python's doctests).
The help gets nicely formatted:
```
?cube
```
When there are several methods for a given function, it is common to give general information about the function in the first method (usually the most generic), and only add docstrings to other methods if they add useful information (without repeating the general info).
Alternatively, you may attach the general information to the function itself:
```
"""
foo(x)
Compute the foo of the bar
"""
function foo end # declares the foo function
# foo(x::Number) behaves normally, no need for a docstring
foo(x::Number) = "baz"
"""
foo(x::String)
For strings, compute the qux of the bar instead.
"""
foo(x::String) = "qux"
?foo
```
# Macros
We have seen a few macros already: `@which`, `@assert`, `@time`, `@benchmark`, `@btime` and `@doc`. You guessed it: all macros start with an `@` sign.
What is a macro? It is a function which can fully inspect the expression that follows it, and apply any transformation to that code at parse time, before compilation.
This makes it possible for anyone to effectively extend the language in any way they please. Whereas C/C++ macros just do simple text replacement, **Julia macros are powerful meta-programming tools**.
On the flip side, this also means that **each macro has its own syntax and behavior**.
**A personal opinion**: in my experience, languages that provide great flexibility typically attract a community of programmers with a tinkering mindset, who will _love_ to experiment with all the fun features the language has to offer. This is great for creativity, but it can also be a nuisance if the community ends up producing too much experimental code, without much care for code reliability, API stability, or even for simplicity. By all means, let's be creative, let's experiment, but _with great power comes great responsibility_: let's also value reliability, stability and simplicity.
That said, to give you an idea of what macro definitions look like in Julia, here's a simple toy macro that replaces `a + b` expressions with `a - b`, and leaves other expressions alone.
```
macro addtosub(x)
if x.head == :call && x.args[1] == :+ && length(x.args) == 3
Expr(:call, :-, x.args[2], x.args[3])
else
x
end
end
@addtosub 10 + 2
```
In this macro definition, `:call`, `:+` and `:-` are **symbols**. These are similar to strings, only more efficient and less flexible. They are typically used as identifiers, such as keys in dictionaries.
If you're curious, the macro works because the parser converts `10 + 2` to `Expr(:call, :+, 10, 2)` and passes this expression to the macro (before compilation). The `if` statement checks that the expression is a function call, where the called function is the `+` function, with two arguments. If so, then the macro returns a new expression, corresponding to a call to the `-` function, with the same arguments. So `a + b` becomes `a - b`.
For more info, check out [this page](https://docs.julialang.org/en/v1/manual/metaprogramming/).
## Special Prefixed Strings
`py"..."` strings are defined by the `PyCall` module. Writing `py"something"` is equivalent to writing `@py_str "something"`. In other words, anyone can write a macro that defines a new kind of prefixed string. For example, if you write the `@ok_str` macro, it will be called when you write `ok"something"`.
Another example is the `Pkg` module which defines the `@pkg_str` macro: this is why you can use `pkg"..."` to interact with the `Pkg` module. This is how `pkg"add PyCall; precompile;"` worked (at the end of the very first cell). This downloaded, installed and precompiled the `PyCall` module.
# Modules
In Python, a module must be defined in a dedicated file. In Julia, modules are independent from the file system. You can define several modules per file, or define one module across multiple files, it's up to you. Let's create a simple module containing two submodules, each containing a variable and a function:
```
module ModA
pi = 3.14
square(x) = x^2
module ModB
e = 2.718
cube(x) = x^3
end
module ModC
root2 = √2
relu(x) = max(0, x)
end
end
```
The default module is `Main`, so whatever we define is put in this module (except when defining a package, as we will see). This is why the `ModA`'s full name is `Main.ModA`.
We can now access the contents of these modules by providing the full paths:
```
Main.ModA.ModC.root2
```
Since our code runs in the `Main` module, we can leave out the `Main.` part:
```
ModA.ModC.root2
```
Alternatively, you can use `import`:
```
import Main.ModA.ModC.root2
root2
```
Or we can use `import` with a relative path. In this case, we need to prefix `ModA` with a dot `.` to indicate that we want the module `ModA` located in the current module:
```
import .ModA.ModC.root2
root2
```
Alternatively, we can `import` the submodule:
```
import .ModA.ModC
ModC.root2
```
When you want to import more than one name from a module, you can use this syntax:
```
import .ModA.ModC: root2, relu
```
This is equivalent to this more verbose syntax:
```
import .ModA.ModC.root2, .ModA.ModC.relu
```
Nested modules do <u>not</u> automatically have access to names in enclosing modules. To import names from a parent module, use `..x`. From a grand-parent module, use `...x`, and so on.
```
module ModD
d = 1
module ModE
try
println(d)
catch ex
println(ex)
end
end
module ModF
f = 2
module ModG
import ..f
import ...d
println(f)
println(d)
end
end
end
```
Instead of `import`, you can use `using`. It is analog to Python's `from foo import *`. It only gives access to names which were explicitly exported using `export` (similar to the way `from foo import *` in Python only imports names listed in the module's `__all__` list):
```
module ModH
h1 = 1
h2 = 2
export h1
end
using .ModH
println(h1)
try
println(h2)
catch ex
ex
end
```
Note that `using Foo` not only imports all exported names (like Python's `from foo import *`), it also imports `Foo` itself (similarly, `using Foo.Bar` imports `Bar` itself):
```
ModH
```
Even if a name is not exported, you can always access it using its full path, or using `import`:
```
ModH.h2
import .ModH.h2
h2
```
You can also import individual names like this:
```
module ModG
g1 = 1
g2 = 2
export g2
end
using .ModG: g1, g2
println(g1)
println(g2)
```
Notice that this syntax gives you access to any name you want, whether or not it was exported. In other words, whether a name is exported or not only affects the `using Foo` syntax.
Importantly, when you want to expand a function which is defined in a module, you must import the function using `import`, or you must specify the function's path:
```
module ModH
double(x) = x * 2
triple(x) = x * 3
end
import .ModH: double
double(x::AbstractString) = repeat(x, 2)
ModH.triple(x::AbstractString) = repeat(x, 3)
println(double(2))
println(double("Two"))
println(ModH.triple(3))
println(ModH.triple("Three"))
```
You must never extend a function imported with `using`, unless you provide the function's path:
```
module ModI
quadruple(x) = x * 4
export quadruple
end
using .ModI
ModI.quadruple(x::AbstractString) = repeat(x, 4) # OK
println(quadruple(4))
println(quadruple("Four"))
#quadruple(x::AbstractString) = repeat(x, 4) # uncomment to see the error
```
There is no equivalent of Python's `import foo as x` ([yet](https://github.com/JuliaLang/julia/issues/1255)), but you can do something like this:
```
import .ModI: quadruple
x = quadruple
```
In general, a module named `Foo` will be defined in a file named `Foo.jl` (along with its submodules). However, if the module becomes too big for a single file, you can split it into multiple files and include these files in `Foo.jl` using the `include()` function.
For example, let's create three files: `Awesome.jl`, `great.jl` and `amazing/Fantastic.jl`, where:
* `Awesome.jl` defines the `Awesome` module and includes the other two files
* `great.jl` just defines a function
* `amazing/Fantastic.jl` defines the `Fantastic` submodule
```
code_awesome = """
module Awesome
include("great.jl")
include("amazing/Fantastic.jl")
end
"""
code_great = """
great() = "This is great!"
"""
code_fantastic = """
module Fantastic
fantastic = true
end
"""
open(f->write(f, code_awesome), "Awesome.jl", "w")
open(f->write(f, code_great), "great.jl", "w")
mkdir("amazing")
open(f->write(f, code_fantastic), "amazing/Fantastic.jl", "w")
```
If we try to execute `import Awesome` now, it won't work since Julia does not search in the current directory by default. Let's change this:
```
pushfirst!(LOAD_PATH, ".")
```
Now when we import the `Awesome` module, Julia will look for a file named `Awesome.jl` in the current directory, or for `Awesome/src/Awesome.jl`, or for `Awesome.jl/src/Awesome.jl`. If it does not find any of these, it will look in the other places listed in the `LOAD_PATH` array (we will discuss this in more details in the "Package Management" section).
```
import Awesome
println(Awesome.great())
println("Is fantastic? ", Awesome.Fantastic.fantastic)
```
Let's restore the original `LOAD_PATH`:
```
popfirst!(LOAD_PATH)
```
In short:
|Julia | Python
|------|-------
|`import Foo` | `import foo`
|`import Foo.Bar` | `from foo import bar`
|`import Foo.Bar: a, b` | `from foo.bar import a, b`
|`import Foo.Bar.a, Foo.Bar.b` | `from foo.bar import a, b`
|`import .Foo` | `import .foo`
|`import ..Foo.Bar` | `from ..foo import bar`
|`import ...Foo.Bar` | `from ...foo import bar`
|`import .Foo: a, b` | `from .foo import a, b`
||
|`using Foo` | `from foo import *; import foo`
|`using Foo.Bar` | `from foo.bar import *; from foo import bar `
|`using Foo.Bar: a, b` | `from foo.bar import a, b`
|Extending function `Foo.f()` | Result
|-----------------------------|--------
|`import Foo.f # or Foo: f` <br />`f(x::Int64) = ...` | OK
|`import Foo`<br />`Foo.f(x::Int64) = ...` | OK
|`using Foo`<br />`Foo.f(x::Int64) = ...` | OK
|`import Foo.f # or Foo: f`<br />`Foo.f(x::Int64) = ...` | `ERROR: Foo not defined`
|`using Foo`<br />`f(x::Int64) = ...` | `ERROR: Foo.f must be explicitly imported`
|`using Foo: f`<br />`f(x::Int64) = ...` | `ERROR: Foo.f must be explicitly imported`
# Scopes
Julia has two types of scopes: global and local.
Every module has its own global scope, independent from all other global scopes. There is no overarching global scope.
Modules, macros and types (including structs) can only be defined in a global scope.
Most code blocks, including `function`, `struct`, `for`, `while`, etc., have their own local scope. For example:
```
for q in 1:3
println(q)
end
try
println(q) # q is not available here
catch ex
ex
end
```
A local scope inherits from its parent scope:
```
z = 5
for i in 1:3
w = 10
println(i * w * z) # i and w are local, z is from the parent scope
end
```
An inner scope can assign to a variable in the parent scope, if the parent scope is not global:
```
for i in 1:3
s = 0
for j in 1:5
s = j # variable s is from the parent scope
end
println(s)
end
```
You can force a variable to be local by using the `local` keyword:
```
for i in 1:3
s = 0
for j in 1:5
local s = j # variable s is local now
end
println(s)
end
```
To assign to a global variable, you must declare the variable as `global` in the local scope:
```
for i in 1:3
global p
p = i
end
p
```
There is one exception to this rule: when executing code directly in the REPL (since Julia 1.5) or in IJulia, you do not need to declare a variable as `global` if the global variable already exists:
```
s = 0
for i in 1:3
s = i # implicitly global s: only in REPL Julia 1.5+ or IJulia
end
s
```
In functions, assigning to a variable which is not explicitly declared as global always makes it local (even in the REPL and IJulia):
```
s, t = 1, 2 # globals
function foo()
s = 10 * t # s is local, t is global
end
println(foo())
println(s)
```
Just like in Python, functions can capture variables from the enclosing scope (not from the scope the function is called from):
```
t = 1
foo() = t # foo() captures t from the global scope
function bar()
t = 5 # this is a new local variable
println(foo()) # foo() still uses t from the global scope
end
bar()
function quz()
global t
t = 5 # we change the global t
println(foo()) # and this affects foo()
end
quz()
```
Closures work much like in Python:
```
function create_multiplier(n)
function mul(x)
x * n # variable n is captured from the parent scope
end
end
mul2 = create_multiplier(2)
mul2(5)
```
An inner function can modify variables from its parent scope:
```
function create_counter()
c = 0
inc() = c += 1 # this inner function modifies the c from the outer function
end
cnt = create_counter()
println(cnt())
println(cnt())
```
Consider the following code, and see if you can figure out why it prints the same result multiple times:
```
funcs = []
i = 1
while i ≤ 5
push!(funcs, ()->i^2)
global i += 1
end
for fn in funcs
println(fn())
end
```
The answer is that there is a single variable `i`, which is captured by all 5 closures. By the time these closures are executed, the value of `i` is 6, so the square is 36, for every closure.
If we use a `for` loop, we don't have this problem, since a new local variable is created at every iteration:
```
funcs = []
for i in 1:5
push!(funcs, ()->i^2)
end
for fn in funcs
println(fn())
end
```
Any local variable created within a `for` loop, a `while` loop or a comprehension also get a new copy at each iteration. So we could code the above example like this:
```
funcs = []
i = 1
while i ≤ 5 # since we are in a while loop...
global i
local j = i # ...and j is created here, it's a new `j` at each iteration
push!(funcs, ()->j^2)
i += 1
end
for fn in funcs
println(fn())
end
```
Another way to get the same result is to use a `let` block, which also creates a new local variable every time it is executed:
```
funcs = []
i = 0
while i < 5
let i=i
push!(funcs, ()->i^2)
end
global i += 1
end
for fn in funcs
println(fn())
end
```
This `let i=i` block defines a new local variable `i` at every iteration, and initializes it with the value of `i` from the parent scope. Therefore each closure captures a different local variable `i`.
Variables in a `let` block are initialized from left to right, so they can access variables on their left:
```
a = 1
let a=a+1, b=a
println("a=$a, b=$b")
end
```
In this example, the local variable `a` is initialized with the value of `a + 1`, where `a` comes from the parent scope (i.e., it's the global `a` in this case). However, `b` is initialized with the value of the local `a`, since it now hides the variable `a` from the parent scope.
Default values in function arguments also have this left-to-right scoping logic:
```
a = 1
foobar(a=a+1, b=a) = println("a=$a, b=$b")
foobar()
foobar(5)
```
In this example, the first argument's default value is `a + 1`, where `a` comes from the parent scope (i.e., the global `a` in this case). However, the second argument's default value is `a`, where `a` in this case is the value of the first argument (<u>not</u> the parent scope's `a`).
Note that `if` blocks and `begin` blocks do <u>not</u> have their own local scope, they just use the parent scope:
```
a = 1
if true
a = 2 # same `a` as above
end
a
a = 1
begin
a = 2 # same `a` as above
end
a
```
# Package Management
## Basic Workflow
The simplest way to write a Julia program is to create a `.jl` file somewhere and run it using `julia`. You would usually do this with your favorite editor, but in this notebook we must do this programmatically. For example:
```
code = """
println("Hello world")
"""
open(f->write(f, code), "my_program1.jl", "w")
```
Then let's run the program using a shell command:
```
;julia my_program1.jl
```
If you need to use a package which is not part of the standard library, such as `PyCall`, you first need to install it using Julia's package manager `Pkg`:
```
using Pkg
Pkg.add("PyCall")
```
Alternatively, in interactive mode, you can enter the `Pkg` mode by typing `]`, then type a command:
```
]add PyCall
```
You can also precompile the new package to avoid the compilation delay when the package is first used:
```
]add PyCall; precompile;
```
One last alternative is to use `pkg"..."` strings to run commands in your programs:
```
pkg"add PyCall; precompile;"
```
Now you can import `PyCall` in any of your Julia programs:
```
code = """
using PyCall
py"print('1 + 2 =', 1 + 2)"
"""
open(f->write(f, code), "my_program2.jl", "w")
;julia my_program2.jl
```
You can also add packages by providing their URL (typically on github). This is useful when you want to use a package which is not in the [official Julia Package registry](https://github.com/JuliaRegistries/General), or when you want the very latest version of a package:
```
]add https://github.com/JuliaLang/Example.jl
```
You can install a specific package version like this:
```
]add PyCall@1.91.3
```
If you only specify version `1` or version `1.91`, Julia will get the latest version with that prefix. For example, `]add PyCall@0.91` would install the latest version `0.91.x`.
You can also update a package to its latest version:
```
]update PyCall
```
You can update all packages to their latest versions:
```
]update
```
If you don't want a particular package to be updated the next time you call `]update`, you can pin it:
```
]pin PyCall
```
To unpin the package:
```
]free PyCall
```
You can also run the tests defined in a package:
```
]test Example
```
Of course, you can remove a package:
```
]rm Example
```
Lastly, you can check which packages are installed using `]status` (or `]st` for short):
```
]st
```
For more `Pkg` commands, type `]help`.
|Julia (in interactive mode) | Python (in a terminal)
|-----|------
|`]status` | `pip freeze`<br />or<br />`conda list`
|`]add Foo` | `pip install foo`<br />or<br />`conda install foo`
|`]add Foo@1.2` | `pip install foo==1.2`<br />or<br />`conda install foo=1.2`
|`]update Foo` | `pip install --upgrade foo`<br />or<br />`conda update foo`
|`]pin Foo` | `foo==<version>` in `requirements.txt`<br /> or<br />`foo=<version>` in `environment.yml`
|`]free Foo` | `foo` in `requirements.txt`<br />or<br />`foo` in `environment.yml`
|`]test Foo` | `python -m unittest foo`
|`]rm Foo` | `pip uninstall foo`<br />or<br />`conda remove foo`
|`]help` | `pip --help`
This workflow is fairly simple, but it means that all of your programs will be using the same version of each package. This is analog to installing packages using `pip install` without using virtual environments.
## Projects
If you want to have multiple projects, each with different libraries and library versions, you should define **projects**. These are analog to Python virtual environments.
A project is just a directory containing a `Project.toml` file and a `Manifest.toml` file:
```
my_project/
Project.toml
Manifest.toml
```
* `Project.toml` is similar to a `requirements.txt` file (for pip) or `environment.yml` (for conda): it lists the dependencies of the project, and compatibility constraints (e.g., `SomeDependency = 2.5`).
* `Manifest.toml` is an automatically generated file which lists the exact versions and unique IDs (UUIDs) of all the packages that Julia found, based on `Project.toml`. It includes all the implicit dependencies of the project's packages. This is useful to reproduce an environment precisely. Analog to the output of `pip --freeze`.
By default, the active project is located in `~/.julia/environments/v#.#` (where `#.#` is the Julia version you are using, such as 1.4). You can set a different project when starting Julia:
```bash
# BASH
julia --project=/path/to/my_project
```
Or you can set the `JULIA_PROJECT` environment variable:
```bash
# BASH
export JULIA_PROJECT=/path/to/my_project
julia
```
Or you can just activate a project directly in Julia (this is analog to running `source my_project/env/bin/activate` when using virtualenv):
```
Pkg.activate("my_project")
```
The `my_project` directory does not exist yet, but it gets created automatically, along with the `Project.toml` and `Manifest.toml` files, when you first add a package:
```
]add PyCall
```
You can also add a package via its URL:
```
]add https://github.com/JuliaLang/Example.jl
```
Let's also add a package with a specific version:
```
]add Example@0.3
```
Now the `Project.toml` and `Manifest.toml` files were created:
```
;find my_project
```
Notice that the packages we added to the project were _not_ placed in the `my_project` directory itself. They were saved in the `~/.julia/packages` directory, the compiled files were placed in `~/.julia/compiled` director, logs were written to `~/.julia/logs` and so on.
If several projects use the same package, it will only be downloaded and built once (well, once per version). The `~/.julia/packages` directory can hold multiple versions of the same package, so it's fine if different projects use different versions of the same package. There will be no conflict, no "dependency hell".
The `Project.toml` just says that the project depends on `PyCall` and `Example`, and it specifies the UUID of this package:
```
print(read("my_project/Project.toml", String))
```
UUIDs are useful to avoid name conflicts. If several people name their package `CoolStuff`, then the UUID will clarify which one we are referring to.
The `Manifest.toml` file is much longer, since it contains all the packages which `PyCall` and `Example` depend on, along with their versions (except for the standard library packages), and the dependency graph. This file should never be modified manually:
```
print(read("my_project/Manifest.toml", String))
```
Note that `Manifest.toml` contains the precise version of the `Example` package that was installed, but the `Project.toml` file does not specify that version `0.3` is required. That's because Julia cannot know whether your project is supposed to work only with any version `0.3.x`, or whether it could work with other versions as well. So if you want to specify a version constraint for the `Example` package, you must add it manually in `Project.toml`. You would normally use your favorite editor to do this, but in this notebook we'll update `Project.toml` programmatically:
```
append_config = """
[compat]
Example = "0.3"
"""
open(f->write(f, append_config), "my_project/Project.toml", "a")
```
Here is the updated `Project.toml` file:
```
print(read("my_project/Project.toml", String))
```
Now if we try to replace `Example` 0.3 with version 0.2, we get an error:
```
try
pkg"add Example@0.2"
catch ex
ex
end
```
Now you can run a program based on this project, and it will have the possibility to use all the packages which have been added to this project, with their specific versions. If you import a package which was not explicitly added to this project, Julia will fallback to the default project:
```
code = """
import PyCall # found in the project
import PyPlot # not found, so falls back to default project
println("Success!")
"""
open(f->write(f, code), "my_program3.jl", "w")
;julia --project=my_project my_program3.jl
```
## Packages
Falling back to the default project is fine, as long as you run the code on your own machine, but if you want to share your code with other people, it would be brittle to count on packages installed in _their_ default project. Instead, if you plan to share your code, you should clearly specify which packages it depends on, and use only these packages. Such a shareable project is called a **package**.
A package is a regular project (as defined above), but with a few extras:
* the `Project.toml` file must specify a `name`, a `version` and a `uuid`.
* there must be a `src/PackageName.jl` file containing a module named `PackageName`.
* you generally want to specify the `authors` and `description`, and maybe also the `license`, `repository` (e.g., the package's github URL), and some `keywords`, but all of these are optional.
It is very easy to create a new package using the `]generate` command. To define the `authors` field, `Pkg` will look up the `user.name` and `user.email` git config entries, so let's define them before we generate the package:
```
;git config --global user.name "Alice Bob"
;git config --global user.email "alice.bob@example.com"
]generate MyPackages/Hello
```
This generated the `MyPackages/Hello/Project.toml` file (along with the enclosing directories) and the `MyPackages/Hello/src/Hello.jl` file. Let's take a look at the `Project.toml` file:
```
print(read("MyPackages/Hello/Project.toml", String))
```
Notice that the project has no dependencies yet, but it has a name, a unique UUID, and a version (plus an author).
Note: if `Pkg` does not find a your name or email in the git config, it falls back to environment variables (`GIT_AUTHOR_NAME`, `GIT_COMMITTER_NAME`, `USER`, `USERNAME`, `NAME` and `GIT_AUTHOR_EMAIL`, `GIT_COMMITTER_EMAIL`, `EMAIL`).
And let's look at the `src/Hello.jl` file:
```
print(read("MyPackages/Hello/src/Hello.jl", String))
```
Let's try to use the `greet()` function from the `Hello` package:
```
try
import Hello
Hello.greet()
catch ex
ex
end
```
Julia could not find the `Hello` package. When you're working on a package, don't forget to activate it first!
```
]activate MyPackages/Hello
import Hello
Hello.greet()
```
It works!
If the `Hello` package depends on other packages, we must add them:
```
]add PyCall Example
```
You must not use any package which has not been added to the project. If you do, you will get a warning.
Once you are happy with your package, you can deploy it to github (or anywhere else). Then you can add it to your own projects just like any other package.
If you want to make your package available to the world via the official Julia registry, you just need to send a Pull Request to https://github.com/JuliaRegistries/General. However, it's highly recommended to automate this using the [Registrator.jl](https://github.com/JuliaRegistries/Registrator.jl) github app.
If you want to use other registries (including private registries), check out [this page](https://julialang.github.io/Pkg.jl/v1.4/registries/#).
Also check out the [`PkgTemplate`](https://github.com/invenia/PkgTemplates.jl) package, which provides more sophisticated templates for creating new packages, for example with continuous integration, code coverage tests, etc.
## Fixing Issues in a Dependency
Sometimes you may run into an issue inside one of the packages your project depends on. When this happens, you can use `Pkg`'s `dev` command to fix the issue. For example, let's pretend the `Example` package has a bug:
```
]dev Example
```
This command cloned the repo into `~/.julia/dev/Example`:
```
;ls -l "~/.julia/dev"
```
It also updated the `Hello` package's `Manifest.toml` file to ensure the package now uses the `Example` clone. You can see this using `]status`:
```
]st
```
So you would now go ahead and edit the clone and fix the bug. Of course, you would also want to send a PR to the package's owners so the source package gets fixed. Once that happens, you can go back to the official `Example` package easily:
```
]free Example
]st
```
## Instantiating a Project
If you want to run someone else's project and you want to make sure you are using the exact same package versions, you can clone the project, and assuming it has a `Manifest.toml` file, you can activate the project and run `]instantiate` to install all the appropriate packages. For example, let's instantiate the `Registrator.jl` project:
```
;git clone https://github.com/JuliaRegistries/Registrator.jl
]activate Registrator.jl
]instantiate
```
Usually, that's all you need to know about projects and packages, but let's look at bit under the hood, so you can handle less common cases.
## Load Path
When you import a package, Julia searches for it in the environments listed in the `LOAD_PATH` array. An **environment** can be a project or a directory containing a bunch of packages directly. By default, the `LOAD_PATH` array contains three elements:
```
LOAD_PATH
```
Here's what these elements mean:
* `"@"` represents the active project, if any: that's the project activated via `--project`, `JULIA_PROJECT`, `]activate` or `Pkg.activate()`.
* `"@v#.#"` represents the default shared project for the version of Julia we are running. That's why it is used by default when there is no active project.
* `"@stdlib"` represents the standard library. This is not a project: it's a directory containing many packages.
If you want to see the actual paths, you can call `Base.load_path()`:
```
Base.load_path()
```
You can change the load path if you want to. For example, if you want Julia to look only in the active project and in the standard library, without looking in the default project, then you can set the `JULIA_LOAD_PATH` environment variable to `"@:@stdlib"`.
If you try to run `my_program3.jl` this way, it will successfully import `PyCall`, but it will fail to import `PyPlot`, since it is not listed in `Project.toml` (however, it would successfully import any package from the standard library):
```
try
withenv("JULIA_LOAD_PATH"=>"@:@stdlib") do
run(`julia --project=my_project my_program3.jl`)
end
catch ex
ex
end
```
You can also modify the `LOAD_PATH` array programmatically, for example to make all the packages in the `my_packages/` directory available to the project:
```
push!(LOAD_PATH, "my_packages")
```
Now any package added to this directory will be directly available to us:
```
]generate my_packages/Hello2
using Hello2
Hello2.greet()
```
This is a convenience for development, as we didn't have to push this package to a repository or even add it to the project. However, it's just for development: once you're happy with your package, make sure to push it to a repo, and add it to the project normally.
## Depots
As we saw earlier, new packages you add to a project are placed in the `~/.julia/packages` directory, logs are placed in `~/.julia/logs`, and so on.
A directory like `~/.julia` which contains `Pkg` related content is called a **depot**. Julia installs all new packages in the default depot, which is the first directory in the `DEPOT_PATH` array (this array can be modified manually in Julia, or set via the `JULIA_DEPOT_PATH` environment variable):
```
DEPOT_PATH
```
The default depot needs to be writeable for the current user, since that's where new packages will be written to (as well as logs and other stuff). The other depots can be read-only: they're typically used for private package registries.
You can occasionally run the `]gc` command, which will remove all unused package versions (`Pkg` will use the logs to located existing projects).
In summary: when some code runs `using Foo` or `import Foo`, the `LOAD_PATH` is used to determine _which_ specific package `Foo` refers to, while the `DEPOT_PATH` is used to determine _where_ it is. The exception is when the `LOAD_PATH` contains directories which directly contain packages: for these packages, the `DEPOT_PATH` is not used.
# Parallel Computing
Julia supports coroutines (aka green threads), multithreading (without a [GIL](https://en.wikipedia.org/wiki/Global_interpreter_lock#:~:text=A%20global%20interpreter%20lock%20(GIL,on%20a%20multi%2Dcore%20processor.) like CPython!), multiprocessing and distributed computing.
## Coroutines
Let's go back to the `fibonacci()` generator function:
```
function fibonacci(n)
Channel() do ch
a, b = 1, 1
for i in 1:n
put!(ch, a)
a, b = b, a + b
end
end
end
for f in fibonacci(10)
println(f)
end
```
Under the hood, `Channel() do ... end` creates a `Channel` object, and spawns an asynchronous `Task` to execute the code in the `do ... end` block. The task is scheduled to execute immediately, but when it calls the `put!()` function on the channel to yield a value, it blocks until another task calls the `take!()` function to grab that value. You do not see the `take!()` function explicitly in this code example, since it is executed automatically in the `for` loop, in the main task. To demonstrate this, we can just call the `take!()` function 10 times to get all the items from the channel:
```
ch = fibonacci(10)
for i in 1:10
println(take!(ch))
end
```
This channel is bound to the task, therefore it is automatically closed when the task ends. So if we try to get one more element, we will get an exception:
```
try
take!(ch)
catch ex
ex
end
```
Here is a more explicit version of the `fibonacci()` function:
```
function fibonacci(n)
function generator_func(ch, n)
a, b = 1, 1
for i in 1:n
put!(ch, a)
a, b = b, a + b
end
end
ch = Channel()
task = @task generator_func(ch, n) # creates a task without starting it
bind(ch, task) # the channel will be closed when the task ends
schedule(task) # start running the task asynchronously
ch
end
```
And here is a more explicit version of the `for` loop:
```
ch = fibonacci(10)
while isopen(ch)
value = take!(ch)
println(value)
end
```
Note that asynchronous tasks (also called "coroutines" or "green threads") are not actually run in parallel: they cooperate to alternate execution. Some functions, such as `put!()`, `take!()`, and many I/O functions, interrupt the current task's execution, at which point it lets Julia's scheduler decide which task should resume its execution. This is just like Python's coroutines.
For more details on coroutines and tasks, see [the manual](https://docs.julialang.org/en/v1/manual/control-flow/#man-tasks-1).
## Multithreading
Julia also supports multithreading. Currently, you need to specify the number of O.S. threads upon startup, by setting the `JULIA_NUM_THREADS` environment variable (or setting the `-t` argument in Julia 1.5+). In the first cell, we configured the IJulia kernel so that set environment variable is set:
```
ENV["JULIA_NUM_THREADS"]
```
The actual number of threads started by Julia may be lower than that, as it is limited to the number of available cores on the machine (thanks to hyperthreading, each physical core may run two threads). Here is the number of threads that were actually started:
```
using Base.Threads
nthreads()
```
Now let's run 10 tasks across these threads:
```
@threads for i in 1:10
println("thread #", threadid(), " is starting task #$i")
sleep(rand()) # pretend we're actually working
println("thread #", threadid(), " is finished")
end
```
Here is a multithreaded version of the `estimate_pi()` function. Each thread computes part of the sum, and the parts are added at the end:
```
function parallel_estimate_pi(n)
s = zeros(nthreads())
nt = n ÷ nthreads()
@threads for t in 1:nthreads()
for i in (1:nt) .+ nt*(t - 1)
@inbounds s[t] += (isodd(i) ? -1 : 1) / (2i + 1)
end
end
return 4.0 * (1.0 + sum(s))
end
@btime parallel_estimate_pi(100_000_000)
```
The `@inbounds` macro is an optimization: it tells the Julia compiler not to add any bounds check when accessing the array. It's safe in this case since the `s` array has one element per thread, and `t` varies from `1` to `nthreads()`, so there is no risk for `s[t]` to be out of bounds.
Let's compare this with the single-threaded implementation:
```
@btime estimate_pi(100_000_000)
```
If you are running this notebook on Colab, the parallel implementation is probably no faster than the single-threaded one. That's because the Colab Runtime only has a single CPU, so there is no benefit from multithreading (plus there is a bit of overhead for managing threads). However, on my 8-core machine, using 16 threads, the parallel implementation is about 6 times faster than the single-threaded one.
Julia has a `mapreduce()` function which makes it easy to implement functions like `parallel_estimate_pi()`:
```
function parallel_estimate_pi2(n)
4.0 * mapreduce(i -> (isodd(i) ? -1 : 1) / (2i + 1), +, 0:n)
end
@btime parallel_estimate_pi2(100_000_000)
```
The `mapreduce()` function is well optimized, so it's about twice faster than `parallel_estimate_pi()`.
You can also spawn a task using `Threads.@spawn`. It will get executed on any one of the running threads (it will not start a new thread):
```
task = Threads.@spawn begin
println("Thread starting")
sleep(1)
println("Thread stopping")
42 # result
end
println("Hello!")
println("The result is: ", fetch(task))
```
The `fetch()` function waits for the thread to finish, and fetches the result. You can also just call `wait()` if you don't need the result.
Last but not least, you can use channels to synchronize and communicate across tasks, even if they are running across separate threads:
```
ch = Channel()
task1 = Threads.@spawn begin
for i in 1:5
sleep(rand())
put!(ch, i^2)
end
println("Finished sending!")
close(ch)
end
task2 = Threads.@spawn begin
foreach(v->println("Received $v"), ch)
println("Finished receiving!")
end
wait(task2)
```
For more details about multithreading, check out [this page](https://docs.julialang.org/en/v1/manual/parallel-computing/#man-multithreading-1).
## Multiprocessing & Distributed Programming
Julia can spawn multiple Julia processes upon startup if you specify the number of processes via the `-p` argument. You can also spawn extra processes from Julia itself:
```
using Distributed
addprocs(4)
workers() # array of worker process ids
```
The main process has id 1:
```
myid()
```
The `@everywhere` macro lets you run any code on all workers:
```
@everywhere println("Hi! I'm worker $(myid())")
```
You can also execute code on a particular worker by using `@spawnat <worker id> <statement>`:
```
@spawnat 3 println("Hi! I'm worker $(myid())")
```
If you specify `:any` instead of a worker id, Julia chooses the worker for you:
```
@spawnat :any println("Hi! I'm worker $(myid())")
```
Both `@everywhere` and `@spawnat` return immediately. The output of `@spawnat` is a `Future` object. You can call `fetch()` on this object to wait for the result:
```
result = @spawnat 3 1+2+3+4
fetch(result)
```
If you import some package in the main process, it is <u>not</u> automatically imported in the workers. For example, the following code fails because the worker does not know what `pyimport` is:
```
using PyCall
result = @spawnat 4 (np = pyimport("numpy"); np.log(10))
try
fetch(result)
catch ex
ex
end
```
You must use `@everywhere` or `@spawnat` to import the packages you need in each worker:
```
@everywhere using PyCall
result = @spawnat 4 (np = pyimport("numpy"); np.log(10))
fetch(result)
```
Similarly, if you define a function in the main process, it is <u>not</u> automatically available in the workers. You must define the function in every worker:
```
@everywhere addtwo(n) = n + 2
result = @spawnat 4 addtwo(40)
fetch(result)
```
You can pass a `Future` to `@everywhere` or `@spawnat`, as long as you wrap it in a `fetch()` function:
```
M = @spawnat 2 rand(5)
result = @spawnat 3 fetch(M) .* 10.0
fetch(result)
```
In this example, worker 2 creates a random array, then worker 3 fetches this array and multiplies each element by 10, then the main process fetches the result and displays it.
## GPU
Julia has excellent GPU support. As you may know, GPUs are devices which can run thousands of threads in parallel. Each thread is slower and more limited than on a CPU, but there are so many of them that plenty of tasks can be executed much faster on a GPU than on a CPU, provided these tasks can be parallelized.
Let's check which GPU device is installed:
```
;nvidia-smi
```
If you're running on Colab, your runtime will generally have an Nvidia Tesla K80 GPU with 12GB of RAM installed, but sometimes other GPUs like Nvidia Tesla T4 16GB, or Nvidia Tesla P100).
If no GPU is detected, go to _Runtime_ > _Change runtime type_, set _Hardware accelerator_ to _GPU_, then go to _Runtime_ > _Factory reset runtime_, then reinstall Julia by running the first cell again, then reload the page and come back here). If you're running on your own machine, make sure you have a compatible GPU card installed, with the appropriate drivers.
Now let's create a large matrix and time how long it takes to square it on the CPU:
```
using BenchmarkTools
M = rand(2^11, 2^11)
function benchmark_matmul_cpu(M)
M * M
return
end
benchmark_matmul_cpu(M) # warm up
@btime benchmark_matmul_cpu($M)
```
Notes:
* For benchmarking, we wrapped the operation in a function which returns `nothing`.
* Why do we have a "warm up" line? Well, since Julia compiles code on the fly the first time it is executed, it's good practice to execute the operation we want to benchmark at least once before starting the benchmark, or else the benchmark will include the compilation time.
* We used `$M` instead of `M` on the last line. This is a feature of the `@btime` macro: it evaluates `M` before benchmarking takes place, to avoid the extra delay that is incurred when [benchmarking with global variables](https://docs.julialang.org/en/latest/manual/performance-tips/#Avoid-global-variables-1).
Now let's benchmark this same operation on the GPU:
```
using CUDA
# Copy the data to the GPU. Creates a CuArray:
M_on_gpu = cu(M)
# Alternatively, create a new random matrix directly on the GPU:
#M_on_gpu = CUDA.CURAND.rand(2^11, 2^11)
function benchmark_matmul_gpu(M)
CUDA.@sync M * M
return
end
benchmark_matmul_gpu(M_on_gpu) # warm up
@btime benchmark_matmul_gpu($M_on_gpu)
```
That's _much_ faster (185x faster in my test on Colab with an NVidia Tesla P100 GPU).
Importantly:
* Before the GPU can work on some data, it needs to be copied to the GPU (or generated there directly).
* the `CUDA.@sync` macro waits for the GPU operation to complete. Without it, the operation would happen in parallel on the GPU, while execution would continue on the CPU. So we would just be timing how long it takes to _start_ the operation, not how long it takes to complete.
* In general, you don't need `CUDA.@sync`, since many operations (including `cu()`) call it implicitly, and it's usually a good idea to let the CPU and GPU work in parallel. Typically, the GPU will be working on the current batch of data while the CPU works on preparing the next batch.
Of course, the speed up will vary depending on the matrix size and the GPU type. Moreover, copying the data from the CPU to the GPU is often the slowest part of the operation, but we only benchmarked the matrix multiplication itself. Let's see what we get if we include the data transfer in the benchmark:
That's still much faster than on the CPU.
Let's check how much RAM we have left on the GPU:
```
CUDA.memory_status()
```
Julia's Garbage Collector will free CUDA arrays like any other object, when there's no more reference to it. However, `CUDA.jl` uses a memory pool to make allocations faster on the GPU, so don't be surprised if the allocated memory on the GPU does not go down immediately. Moreover, IJulia keeps a reference to the output of each cell, so if you let any cell output a `CuArray`, it will only be released when you execute `Out[<cell number>]=0`. If you want to force the Garbage Collector to run, you an run `GC.gc()`. To reclaim memory from the memory pool, use `CUDA.reclaim()`:
```
GC.gc()
CUDA.reclaim()
```
Many other operations are implemented for `CuArray` (`+`, `-`, etc.) and dotted operations (`.+`, `exp.()`, etc). Importantly, loop fusion also works on the GPU. For example, if we want to compute `M .* M .+ M`, without loop fusion the GPU would first compute `M .* M` and create a temporary array, then it would add `M` to that array, like this:
```
function benchmark_without_fusion(M)
P = M .* M
CUDA.@sync P .+ M
return
end
benchmark_without_fusion(M_on_gpu) # warm up
@btime benchmark_without_fusion($M_on_gpu)
```
Instead, loop fusion ensures that the array is only traversed once, without the need for a temporary array:
```
function benchmark_with_fusion(M)
CUDA.@sync M .* M .+ M
return
end
benchmark_with_fusion(M_on_gpu) # warm up
@btime benchmark_with_fusion($M_on_gpu)
```
That's _much_ faster (almost twice as fast in my test on Colab). 😃
Lastly, you can actually **write your own GPU kernels in Julia**! In other words, rather than using GPU operations implemented in the `CUDA.jl` package (or others), you can write Julia code that will be compiled for the GPU, and executed there. This can occasionally be useful to speed up some algorithms where the standard kernels don't suffice. For example, here's a GPU kernel which implements `u .+= v`, where `u` and `v` are two (large) vectors:
```
function worker_gpu_add!(u, v)
index = (blockIdx().x - 1) * blockDim().x + threadIdx().x
index ≤ length(u) && (@inbounds u[index] += v[index])
return
end
function gpu_add!(u, v)
numblocks = ceil(Int, length(u) / 256)
@cuda threads=256 blocks=numblocks worker_gpu_add!(u, v)
return u
end
```
This code example is adapted from the [`CUDA.jl` package's documentation](https://juliagpu.gitlab.io/CUDA.jl/tutorials/introduction/), which I highly encourage you to check out if you're interested in writing your own kernels. Here are the key parts to understand this example, starting from the end:
* The `gpu_add!()` function first calculates `numblocks`, the number of blocks of threads to start, then it uses the `@cuda` macro to spawn `numblocks` blocks of GPU threads, each with 256 threads, and each thread runs `worker_gpu_add!(u, v)`.
* The `worker_gpu_add!()` function computes `u[index] += v[index]` for a single value of `index`: in other words, each thread will just update a single value in the vector! Let's see how the index is computed:
* The `@cuda` macro spawned many blocks of 256 threads each. These blocks are organized in a grid, which is one-dimensional by default, but it can be up to three-dimensional. Therefore each thread and each block have an `(x, y, z)` coordinate in this grid. See this diagram from the [Nvidia blog post](https://developer.nvidia.com/blog/even-easier-introduction-cuda/):<br />
<img src="https://juliagpu.gitlab.io/CUDA.jl/tutorials/intro1.png" width="600"/>.
* `threadIdx().x` returns the current GPU thread's `x` coordinate within its block (one difference with the diagram is that Julia is 1-indexed).
* `blockIdx().x` returns the current block's `x` coordinate in the grid.
* `blockDim().x` returns the block size along the `x` axis (in this example, it's 256).
* `gridDim().x` returns the number of blocks in the grid, along the `x` axis (in this example it's `numblocks`).
* So the `index` that each thread must update in the array is `(blockIdx().x - 1) * blockDim().x + threadIdx().x`.
* As explained earlier, the `@inbounds` macro is an optimization that tells Julia that the index is guaranteed to be inbounds, so there's no need for it to check.
Now writing your own GPU kernel won't seem like something only top experts with advanced C++ skills can do: you can do it too!
Let's check that the kernel works as expected:
```
u = rand(2^20)
v = rand(2^20)
u_on_gpu = cu(u)
v_on_gpu = cu(v)
u .+= v
gpu_add!(u_on_gpu, v_on_gpu)
@assert Array(u_on_gpu) ≈ u
```
Yes, it works well!
Note: the `≈` operator checks whether the operands are approximately equal within the float precision limit.
Let's benchmark our custom kernel:
```
function benchmark_custom_assign_add!(u, v)
CUDA.@sync gpu_add!(u, v)
return
end
benchmark_custom_assign_add!(u_on_gpu, v_on_gpu)
@btime benchmark_custom_assign_add!($u_on_gpu, $v_on_gpu)
```
Let's see how this compares to `CUDA.jl`'s implementation:
```
function benchmark_assign_add!(u, v)
CUDA.@sync u .+= v
return
end
benchmark_assign_add!(u_on_gpu, v_on_gpu)
@btime benchmark_assign_add!($u_on_gpu, $v_on_gpu)
```
How about that? Our custom kernel is just as fast as `CUDA.jl`'s kernel! But to be fair, our kernel would not work with huge vectors, since there's a limit to the number of blocks & threads you can spawn (see [Table 15](https://docs.nvidia.com/cuda/cuda-c-programming-guide/index.html#features-and-technical-specifications) in CUDA's documentation). To support such huge vectors, we need each worker to run a loop like this:
```
function worker_gpu_add!(u, v)
index = (blockIdx().x - 1) * blockDim().x + threadIdx().x
stride = blockDim().x * gridDim().x
for i = index:stride:length(u)
@inbounds u[i] += v[i]
end
return
end
```
This way, if `@cuda` is executed with a smaller number of blocks than needed to have one thread per array item, the workers will loop appropriately.
This should get you started! For more info, check out [`CUDA.jl`'s documentation](https://juliagpu.gitlab.io/CUDA.jl/).
# Command Line Arguments
Command line arguments are available via `ARGS`:
```
ARGS
```
Unlike Python's `sys.argv`, the first element of this array is <u>not</u> the program name. If you need the program name, use `PROGRAM_FILE` instead:
```
PROGRAM_FILE
```
You can get the current module, directory, file or line number:
```
@__MODULE__, @__DIR__, @__FILE__, @__LINE__
```
The equivalent of Python's `if __name__ == "__main__"` is:
```
if abspath(PROGRAM_FILE) == @__FILE__
println("Starting of the program")
end
```
# Memory Management
Let's check how many megabytes of RAM are available:
```
free() = println("Available RAM: ", Sys.free_memory() ÷ 10^6, " MB")
free()
```
If a variable holds a large object that you don't need anymore, you can either wait until the variable falls out of scope, or set it to `nothing`. Either way, the memory will only be freed when the Garbage Collector does its magic, which may not be immediate. In general, you don't have to worry about that, but if you want, you can always call the GC directly:
```
function use_ram()
M = rand(10000, 10000) # use 400+MB of RAM
println("sum(M)=$(sum(M))")
end # M will be freed by the GC eventually after this
use_ram()
M = rand(10000, 10000) # use 400+MB of RAM
println("sum(M)=$(sum(M))")
M = nothing
GC.gc() # rarely needed
free()
```
# Thanks!
I hope you enjoyed this introduction to Julia! I recommend you join the friendly and helpful Julia community on Slack or Discourse.
Cheers!
Aurélien Geron
```
```
| github_jupyter |
```
from jkg_evaluators import dragonfind_10_to_500
cow_alive_list_test_1 = [False, False, True, True, True]
def solution_katinka(dead_or_alive,
number_of_cows):
MC = int(number_of_cows / 2)
if dead_or_alive(MC):
for i in range(MC, number_of_cows):
if not dead_or_alive(i):
return i
else:
for i in range(1, MC):
if not dead_or_alive(i):
return i
dragonfind_10_to_500.evaluate(solution_katinka)
```
<a style='text-decoration:none;line-height:16px;display:flex;color:#5B5B62;padding:10px;justify-content:end;' href='https://deepnote.com?utm_source=created-in-deepnote-cell&projectId=978e47b7-a961-4dca-a945-499e8b781a34' target="_blank">
<img alt='Created in deepnote.com' style='display:inline;max-height:16px;margin:0px;margin-right:7.5px;' src='data:image/svg+xml;base64,PD94bWwgdmVyc2lvbj0iMS4wIiBlbmNvZGluZz0iVVRGLTgiPz4KPHN2ZyB3aWR0aD0iODBweCIgaGVpZ2h0PSI4MHB4IiB2aWV3Qm94PSIwIDAgODAgODAiIHZlcnNpb249IjEuMSIgeG1sbnM9Imh0dHA6Ly93d3cudzMub3JnLzIwMDAvc3ZnIiB4bWxuczp4bGluaz0iaHR0cDovL3d3dy53My5vcmcvMTk5OS94bGluayI+CiAgICA8IS0tIEdlbmVyYXRvcjogU2tldGNoIDU0LjEgKDc2NDkwKSAtIGh0dHBzOi8vc2tldGNoYXBwLmNvbSAtLT4KICAgIDx0aXRsZT5Hcm91cCAzPC90aXRsZT4KICAgIDxkZXNjPkNyZWF0ZWQgd2l0aCBTa2V0Y2guPC9kZXNjPgogICAgPGcgaWQ9IkxhbmRpbmciIHN0cm9rZT0ibm9uZSIgc3Ryb2tlLXdpZHRoPSIxIiBmaWxsPSJub25lIiBmaWxsLXJ1bGU9ImV2ZW5vZGQiPgogICAgICAgIDxnIGlkPSJBcnRib2FyZCIgdHJhbnNmb3JtPSJ0cmFuc2xhdGUoLTEyMzUuMDAwMDAwLCAtNzkuMDAwMDAwKSI+CiAgICAgICAgICAgIDxnIGlkPSJHcm91cC0zIiB0cmFuc2Zvcm09InRyYW5zbGF0ZSgxMjM1LjAwMDAwMCwgNzkuMDAwMDAwKSI+CiAgICAgICAgICAgICAgICA8cG9seWdvbiBpZD0iUGF0aC0yMCIgZmlsbD0iIzAyNjVCNCIgcG9pbnRzPSIyLjM3NjIzNzYyIDgwIDM4LjA0NzY2NjcgODAgNTcuODIxNzgyMiA3My44MDU3NTkyIDU3LjgyMTc4MjIgMzIuNzU5MjczOSAzOS4xNDAyMjc4IDMxLjY4MzE2ODMiPjwvcG9seWdvbj4KICAgICAgICAgICAgICAgIDxwYXRoIGQ9Ik0zNS4wMDc3MTgsODAgQzQyLjkwNjIwMDcsNzYuNDU0OTM1OCA0Ny41NjQ5MTY3LDcxLjU0MjI2NzEgNDguOTgzODY2LDY1LjI2MTk5MzkgQzUxLjExMjI4OTksNTUuODQxNTg0MiA0MS42NzcxNzk1LDQ5LjIxMjIyODQgMjUuNjIzOTg0Niw0OS4yMTIyMjg0IEMyNS40ODQ5Mjg5LDQ5LjEyNjg0NDggMjkuODI2MTI5Niw0My4yODM4MjQ4IDM4LjY0NzU4NjksMzEuNjgzMTY4MyBMNzIuODcxMjg3MSwzMi41NTQ0MjUgTDY1LjI4MDk3Myw2Ny42NzYzNDIxIEw1MS4xMTIyODk5LDc3LjM3NjE0NCBMMzUuMDA3NzE4LDgwIFoiIGlkPSJQYXRoLTIyIiBmaWxsPSIjMDAyODY4Ij48L3BhdGg+CiAgICAgICAgICAgICAgICA8cGF0aCBkPSJNMCwzNy43MzA0NDA1IEwyNy4xMTQ1MzcsMC4yNTcxMTE0MzYgQzYyLjM3MTUxMjMsLTEuOTkwNzE3MDEgODAsMTAuNTAwMzkyNyA4MCwzNy43MzA0NDA1IEM4MCw2NC45NjA0ODgyIDY0Ljc3NjUwMzgsNzkuMDUwMzQxNCAzNC4zMjk1MTEzLDgwIEM0Ny4wNTUzNDg5LDc3LjU2NzA4MDggNTMuNDE4MjY3Nyw3MC4zMTM2MTAzIDUzLjQxODI2NzcsNTguMjM5NTg4NSBDNTMuNDE4MjY3Nyw0MC4xMjg1NTU3IDM2LjMwMzk1NDQsMzcuNzMwNDQwNSAyNS4yMjc0MTcsMzcuNzMwNDQwNSBDMTcuODQzMDU4NiwzNy43MzA0NDA1IDkuNDMzOTE5NjYsMzcuNzMwNDQwNSAwLDM3LjczMDQ0MDUgWiIgaWQ9IlBhdGgtMTkiIGZpbGw9IiMzNzkzRUYiPjwvcGF0aD4KICAgICAgICAgICAgPC9nPgogICAgICAgIDwvZz4KICAgIDwvZz4KPC9zdmc+' > </img>
Created in <span style='font-weight:600;margin-left:4px;'>Deepnote</span></a>
| github_jupyter |
```
from __future__ import absolute_import, division, print_function, unicode_literals
import tensorflow as tf
import time
import datetime
import os
from tqdm import tqdm
import numpy as np
import matplotlib.pyplot as plt
plt.rcParams['font.sans-serif']=['SimHei'] # 用来正常显示中文标签
plt.rcParams['axes.unicode_minus']=False
os.environ["CUDA_VISIBLE_DEVICES"] = "0, 1, 2, 3" # 添加可用的gpu
physical_devices = tf.config.experimental.list_physical_devices('GPU')
for device in physical_devices:
tf.config.experimental.set_memory_growth(device, True)
from params import Params as pm
from utils_v2 import en2idx, idx2en, de2idx, idx2de, dump2record, build_dataset, LRSchedule, masking, create_masks, plot_attention_weights
from bleu import bleu_metrics
tf.__version__
```
---
```
strategy = tf.distribute.MirroredStrategy()
print('Number of device: {}'.format(strategy.num_replicas_in_sync))
def get_data(corpus_file):
return open(corpus_file, 'r', encoding='utf-8').read().splitlines()
src_train, src_val = get_data(pm.src_train), get_data(pm.src_test)
tgt_train, tgt_val = get_data(pm.tgt_train), get_data(pm.tgt_test)
dump2record(pm.train_record, src_train, tgt_train)
dump2record(pm.test_record, src_val, tgt_val)
```
---
```
from modules_v2 import positional_encoding, scaled_dot_product_attention, multihead_attention, pointwise_feedforward, EncoderBlock, DecoderBlock, Encoder, Decoder, Transformer
```
# Positional encoding
$$\Large{PE_{(pos, 2i)} = sin(pos / 10000^{2i / d_{model}})} $$
$$\Large{PE_{(pos, 2i+1)} = cos(pos / 10000^{2i / d_{model}})} $$
```
pos_encoding = positional_encoding(50, 512, True)
print(pos_encoding.shape)
```
# Masking
```
x = tf.constant([[7, 6, 0, 0, 1], [1, 2, 3, 0, 0], [0, 0, 0, 4, 5]])
masking(x, task='padding')
masking(x, task='look_ahead')
```
# Scaled dot product attention
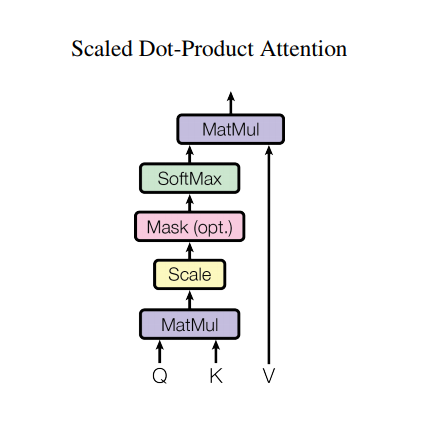
$$\Large{Attention(Q, K, V) = softmax_k(\frac{QK^T}{\sqrt{d_k}}) V} $$
```
def print_out(q, k, v):
temp_out, temp_attn = scaled_dot_product_attention(q, k, v, None)
print ('Attention weights are:')
print (temp_attn)
print ('Output is:')
print (temp_out)
np.set_printoptions(suppress=True)
temp_k = tf.constant([[10,0,0],
[0,10,0],
[0,0,10],
[0,0,10]], dtype=tf.float32)
temp_v = tf.constant([[ 1,0],
[ 10,0],
[ 100,5],
[1000,6]], dtype=tf.float32)
temp_q = tf.constant([[0, 10, 0]], dtype=tf.float32)
print_out(temp_q, temp_k, temp_v)
temp_q = tf.constant([[0, 0, 10]], dtype=tf.float32)
print_out(temp_q, temp_k, temp_v)
temp_q = tf.constant([[0, 0, 10], [0, 10, 0], [10, 10, 0]], dtype=tf.float32)
print_out(temp_q, temp_k, temp_v)
```
# Multi-head attention
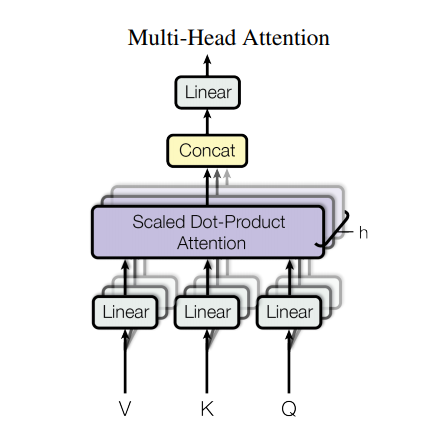
- **Tips: Dimention-level split**
```
temp_mha = multihead_attention(d_model=512, num_heads=8)
y = tf.random.uniform((1, 50, 512))
out, attn = temp_mha(y, k=y, q=y, mask=None)
out.shape, attn.shape
```
# Pointwise feed forward network
```
sample_ffn = pointwise_feedforward(512, 2048)
sample_ffn(tf.random.uniform((64, 50, 512))).shape
```
# Whole model (Encoder & Decoder)
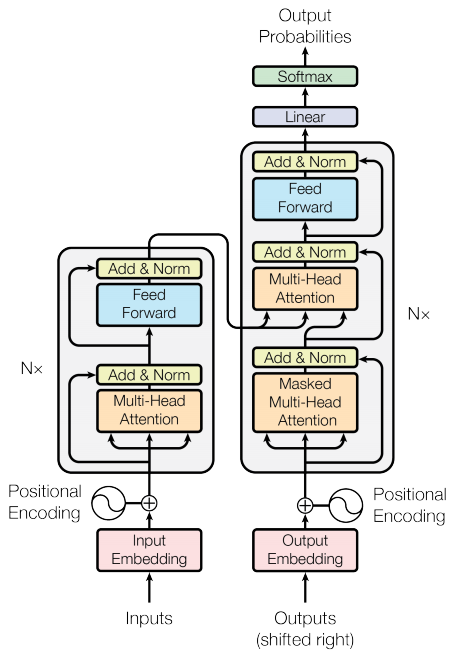
## Encoder
```
sample_encoder_layer = EncoderBlock(512, 8, 2048)
sample_encoder_layer_output, _ = sample_encoder_layer(tf.random.uniform((64, 43, 512)), False, None)
sample_encoder_layer_output.shape
```
## Decoder
```
sample_decoder_layer = DecoderBlock(512, 8, 2048)
sample_decoder_layer_output, _, _ = sample_decoder_layer(
tf.random.uniform((64, 50, 512)), sample_encoder_layer_output,
False, None, None)
sample_decoder_layer_output.shape
```
## Packed Encoder & Decoder
```
sample_encoder = Encoder(num_blocks=2, d_model=512, num_heads=8, dff=2048, input_vocab_size=8500, plot_pos_embedding=False)
attn_dict = {}
sample_encoder_output, attn_dict = sample_encoder(tf.random.uniform((64, 62)), training=False, padding_mask=None, attn_dict=attn_dict)
sample_encoder_output.shape
sample_decoder = Decoder(num_blocks=2, d_model=512, num_heads=8, dff=2048, target_vocab_size=8000, plot_pos_embedding=False)
output, attn_dict = sample_decoder(tf.random.uniform((64, 26)),
enc_output=sample_encoder_output,
training=False, look_ahead_mask=None,
padding_mask=None, attn_dict=attn_dict)
output.shape, attn_dict['decoder_layer2_block'].shape
```
# Transformer
```
sample_transformer = Transformer(num_blocks=2, d_model=512, num_heads=8, dff=2048, input_vocab_size=8500, target_vocab_size=8000, plot_pos_embedding=False)
temp_input = tf.random.uniform((64, 62))
temp_target = tf.random.uniform((64, 26))
fn_out, _ = sample_transformer(temp_input,
temp_target,
training=False,
enc_padding_mask=None,
look_ahead_mask=None,
dec_padding_mask=None)
fn_out.shape
```
# Training
```
num_layers = pm.num_block
d_model = pm.d_model
dff = pm.dff
num_heads = pm.num_heads
input_vocab_size = len(en2idx)
target_vocab_size = len(de2idx)
dropout_rate = pm.dropout_rate
EPOCHS = pm.num_epochs
```
- Learning rate schedule
$$\Large{lrate = d_{model}^{-0.5} * min(step{\_}num^{-0.5}, step{\_}num * warmup{\_}steps^{-1.5})}$$
```
temp_learning_rate_schedule = LRSchedule(d_model)
plt.figure(figsize=(12, 8))
plt.plot(temp_learning_rate_schedule(tf.range(40000, dtype=tf.float32)))
plt.ylabel("Learning Rate")
plt.xlabel("Train Step")
```
---
```
with strategy.scope():
# 1、dataset
## train_dataset = build_dataset(mode='array', batch_size=pm.batch_size * strategy.num_replicas_in_sync, cache_name='train_cache.tf-data', corpus=[src_train, tgt_train], is_training=True)
## val_dataset = build_dataset(mode='array', batch_size=pm.batch_size * strategy.num_replicas_in_sync, cache_name='val_cache.tf-data', corpus=[src_val, tgt_val], is_training=True)
train_dataset = build_dataset(mode='file', batch_size=pm.batch_size * strategy.num_replicas_in_sync, cache_name='train_cache.tf-data', filename=pm.train_record, is_training=True)
train_dist_dataset = strategy.experimental_distribute_dataset(train_dataset)
# 2、loss function
loss_object = tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True, reduction=tf.keras.losses.Reduction.NONE)
def loss_function(real, pred):
mask = tf.math.logical_not(tf.math.equal(real, 0))
loss_ = loss_object(real, pred)
mask = tf.cast(mask, dtype=loss_.dtype)
loss_ *= mask
return tf.reduce_mean(loss_), mask
# 3、metrics to track loss and accuracy
train_loss = tf.keras.metrics.Mean(name='train_loss')
train_accuracy = tf.keras.metrics.SparseCategoricalAccuracy(name='train_accuracy')
# 4、model config
transformer = Transformer(num_layers, d_model, num_heads, dff, input_vocab_size, target_vocab_size, pm.plot_pos_embedding, dropout_rate)
learning_rate = LRSchedule(d_model)
optimizer = tf.keras.optimizers.Adam(learning_rate, beta_1=pm.beta_1, beta_2=pm.beta_2, epsilon=pm.epsilon)
checkpoint_path = pm.ckpt_path
ckpt = tf.train.Checkpoint(transformer=transformer, optimizer=optimizer)
ckpt_manager = tf.train.CheckpointManager(ckpt, checkpoint_path, max_to_keep=5)
if ckpt_manager.latest_checkpoint:
ckpt.restore(ckpt_manager.latest_checkpoint)
print ('Latest checkpoint restored!!')
current_time = datetime.datetime.now().strftime("%Y%m%d-%H%M%S")
log_dir = pm.logdir + '/gradient_tape/' + current_time
summary_writer = tf.summary.create_file_writer(log_dir)
# 5、train step
def train_step(inp, tar):
tar_inp = tar[:, :-1]
tar_real = tar[:, 1:]
enc_padding_mask, combined_mask, dec_padding_mask = create_masks(inp, tar_inp)
with tf.GradientTape() as tape:
predictions, _ = transformer(inp,
tar_inp,
True,
enc_padding_mask,
combined_mask,
dec_padding_mask)
loss, istarget = loss_function(tar_real, predictions)
gradients = tape.gradient(loss, transformer.trainable_variables)
optimizer.apply_gradients(zip(gradients, transformer.trainable_variables))
train_accuracy(tar_real, predictions, sample_weight=istarget)
return loss
@tf.function
def distributed_train_step(inp, tar):
per_replica_losses = strategy.experimental_run_v2(train_step,
args=(inp, tar, ))
return strategy.reduce(tf.distribute.ReduceOp.SUM, per_replica_losses, axis=None)
# 6、for loop
total_steps = 0
for epoch in range(EPOCHS):
start = time.time()
train_loss.reset_states()
train_accuracy.reset_states()
total_loss = 0.0
num_batches = 0
for (batch, (inp, tar)) in enumerate(train_dataset):
total_loss += distributed_train_step(inp, tar)
num_batches += 1
total_steps += 1
if batch % 500 == 0:
with summary_writer.as_default():
tf.summary.scalar('loss', total_loss / num_batches, step=total_steps)
tf.summary.scalar('accuracy', train_accuracy.result() * 100, step=total_steps)
print ('Epoch {} Batch {} Loss {:.4f} Accuracy {:.4f}'.format(
epoch + 1, batch, total_loss / num_batches, train_accuracy.result() * 100))
train_loss(total_loss / num_batches)
if (epoch + 1) % 5 == 0:
ckpt_save_path = ckpt_manager.save()
print ('Saving checkpoint for epoch {} at {}'.format(epoch + 1, ckpt_save_path))
print ('Epoch {} Loss {:.4f} Accuracy {:.4f}'.format(epoch + 1, train_loss.result(), train_accuracy.result() * 100))
print ('Time taken for 1 epoch: {} secs\n'.format(time.time() - start))
```
---
```
val_dataset = build_dataset(mode='file', batch_size=pm.batch_size * strategy.num_replicas_in_sync, cache_name='val_cache.tf-data', filename=pm.test_record, is_training=True)
def evaluate(inp_sentence):
encoder_input = inp_sentence
decoder_input = [2]
output = tf.expand_dims(decoder_input, 0)
output = tf.tile(output, [tf.shape(encoder_input)[0], 1])
for i in range(pm.maxlen):
enc_padding_mask, combined_mask, dec_padding_mask = create_masks(encoder_input, output)
predictions, attention_weights = transformer(encoder_input,
output,
False,
enc_padding_mask,
combined_mask,
dec_padding_mask)
predictions = predictions[: ,-1:, :]
predicted_id = tf.cast(tf.argmax(predictions, axis=-1), tf.int32)
output = tf.concat([output, predicted_id], axis=-1)
return output, attention_weights
def cut_by_end(samples):
output_list = np.zeros(tf.shape(samples))
for i, sample in enumerate(samples):
dtype = sample.dtype
idx = tf.where(tf.equal(sample, 3))
flag = tf.where(tf.equal(tf.size(idx), 0), 1, 0)
if flag:
output_list[i] = sample
else:
indices = tf.cast(idx[0, 0], dtype)
output_list[i] = tf.concat([sample[:indices], tf.zeros(tf.shape(sample)[0] - indices, dtype=dtype)], axis=0)
return tf.cast(output_list, dtype)
eval_log = os.path.join(pm.eval_log_path, '{}_eval.tsv'.format(pm.project_name))
if not os.path.exists(pm.eval_log_path):
os.makedirs(pm.eval_log_path)
eval_file = open(eval_log, 'w', encoding='utf-8')
start = time.time()
count, scores = 0, 0
for (batch, (inp, tar)) in enumerate(val_dataset):
prediction, attention_weights = evaluate(inp)
prediction = cut_by_end(prediction)
preds, tars = [], []
for source, real_tar, pred in zip(inp, tar, prediction):
s = " ".join([idx2en.get(i, 1) for i in source.numpy() if i < len(idx2en) and i not in [0, 2, 3]])
t = "".join([idx2de.get(i, 1) for i in real_tar.numpy() if i < len(idx2de) and i not in [0, 2, 3]])
p = "".join([idx2de.get(i, 1) for i in pred.numpy() if i < len(idx2de) and i not in [0, 2, 3]])
preds.append(p)
tars.append([t])
eval_file.write('-Source : {}\n-Target : {}\n-Pred : {}\n\n'.format(s, t, p))
eval_file.flush()
scores += bleu_metrics(tars, preds, False, 3, True)
count += 1
eval_file.write('-BLEU Score : {:.4f}'.format(scores / count))
eval_file.close()
print("MSG : Done for evalutation ... Totolly {:.2f} sec.".format(time.time() - start))
def predict(inp_sentence):
start_token = [2]
end_token = [3]
inp_sentence = start_token + [en2idx.get(word, 1) for word in inp_sentence.split()] + end_token
encoder_input = tf.expand_dims(inp_sentence, 0)
decoder_input = [2]
output = tf.expand_dims(decoder_input, 0)
for i in range(pm.maxlen):
enc_padding_mask, combined_mask, dec_padding_mask = create_masks(encoder_input, output)
predictions, attention_weights = transformer(encoder_input,
output,
False,
enc_padding_mask,
combined_mask,
dec_padding_mask)
predictions = predictions[: ,-1:, :]
predicted_id = tf.cast(tf.argmax(predictions, axis=-1), tf.int32)
if tf.equal(predicted_id, 3):
return tf.squeeze(output, axis=0), attention_weights
output = tf.concat([output, predicted_id], axis=-1)
return tf.squeeze(output, axis=0), attention_weights
def translate(sentence, plot=''):
result, attention_weights = predict(sentence)
predicted_sentence = [idx2de.get(i, 1) for i in result.numpy() if i < len(idx2de) and i not in [0, 2, 3]]
print('Input: {}'.format(sentence))
print('Predicted translation: {}'.format(" ".join(predicted_sentence)))
if plot:
plot_attention_weights(attention_weights, sentence, result, plot)
translate("明 天 就 要 上 班 了", plot='decoder_layer4_block')
print("Real translation: 還好我沒工作QQ")
```
| github_jupyter |
<h1 style="text-align:center">执行</h1>
<h1 style="text-align:center">强制执行</h1>
<h1 style="text-align:center">失信被执行</h1>
```
import time
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from matplotlib.pyplot import figure
figure(figsize=(10,10), dpi=80)
import matplotlib.ticker as ticker
import seaborn as sns
from nltk.corpus import stopwords
stopwords = stopwords.words("chinese")
import jieba
import jieba.posseg as pseg
from sklearn import feature_extraction
from sklearn.feature_extraction.text import TfidfTransformer
from sklearn.feature_extraction.text import CountVectorizer
import pandas as pd
import re
## Pandas, display all row values
pd.set_option('display.max_rows', 99999)
sh_sx = pd.read_csv("../../data/sh_corp.csv")
sh_sx.sample(15)
sh_sx.dropna(inplace = True)
sh_sx = sh_sx[sh_sx.duty.str.len() >= 20]
sh_sx.shape
sh_sx.describe().T # with no surprise, other columns are non-numeric; time stamp, or string or categorical.
for i in sh_sx.columns:
print(f"{i}\n")
```
# 1. Subset the data to only contains 2015 Jan and beyond
```
## Converting reg_date&publish_date from `str` to timestamp
sh_sx['reg_date'] = pd.to_datetime(sh_sx.reg_date)
sh_sx['publish_date'] = pd.to_datetime(sh_sx.publish_date)
sh_sx.sample(20)
sh_sx_2017 = sh_sx[sh_sx["reg_date"].dt.year >= 2015] # subset to 2015 and beyond
print(f'max date\n {sh_sx_2017.reg_date.max()}\nmin date\n {sh_sx_2017.reg_date.min()}\nmedian date\n {sh_sx_2017.reg_date.median()}')
figsize=(50,10)
sh_sx_2017.groupby('reg_date')['X'].count().plot.bar(figsize=figsize)
sh_sx_2017.groupby('reg_date')['X'].count().sort_values(ascending=False).head(20)
date = pd.DataFrame(sh_sx_2017.groupby('reg_date')['X'].count())
date.reset_index(inplace = True)
date.rename(columns = {'X':'Count'}, inplace=True)
date.head(10)
plt.figure(figsize=(20,5))
ax = sns.lineplot(x = "reg_date", y= "Count", data=date)
```
# Check the str contains
## 失信 | 执行 | 被执行 | 被执行人| 强制执行
```
pd.DataFrame(sh_sx.groupby("disrupt_type_name")['iname'].count().sort_values(ascending=False)).reset_index().head(20)
pd.DataFrame(sh_sx.groupby("performance")['iname'].count().sort_values(ascending=False)).reset_index()
pd.DataFrame(sh_sx.groupby("gist_unit")['iname'].count().sort_values(ascending=False)).reset_index().head(20)
pd.DataFrame(sh_sx.groupby("iname")['X'].count().sort_values(ascending=False)).reset_index().head(20)
```
# @TODO 2 -- why duplicate
```
sh_sx_str_shixin = sh_sx[
(sh_sx.duty.str.contains("失信")) # |
# (sh_sx.duty.str.contains("被执行")) |
# (sh_sx.duty.str.contains("失信执行")) |
# (sh_sx.duty.str.contains("被执行人")) |
# (sh_sx.duty.str.contains("强制执行")) |
# (sh_sx.duty.str.contains("申请执行"))
]
print(f'Only {len(sh_sx_str_shixin)} entries contains word token 失信')
sh_sx_zhixin = sh_sx[
(sh_sx.duty.str.contains("执行")) # |
# (sh_sx.duty.str.contains("被执行")) |
# (sh_sx.duty.str.contains("失信执行")) |
# (sh_sx.duty.str.contains("被执行人")) |
# (sh_sx.duty.str.contains("强制执行")) |
# (sh_sx.duty.str.contains("申请执行"))
]
text_duty = sh_sx_zhixin.sample(10)
text_duty
sh_sx.iloc[48670]
text_duty.duty[48670]
```
### Post processed
'一 上纽 资产 管理 应 归还 原告 张显 爱本 金元 及 期内 利息 合计 具体 日期 如下 之前 归还 之前 归还 之前 归还 元二 上纽 资产 管理 应 于 之前 偿付 原告 张显 爱 逾期 利息 以本 金元 为 基数 自 起 至 实际 清偿 以本 金元 为 基数 自 起 至 实际 清偿 以本 金元 为 基数 自 起 至 实际 清偿 上述 均 按照 计算 三 上纽 资产 管理 应 于 之前 原告 张显 爱 元四如 上纽 资产 管理 届时 未 按 上述 三项 确定 的 期限 履行 付款 义务 的 则 原告 张显 爱 有权 就 上纽 资产 管理 在 本案 中未 履行 的 全部 债务 一并 向 法院 申请 强制执行 五 陈刚 对 上纽 资产 管理 上述 付款 义务 中 的 本 金元 承担 连带 清偿 责任 陈刚 在 承担 了 保证 责任 后 有权 向 上纽 资产 管理 追偿 六 王俊 对 上纽 资产 管理 上述 三项 付款 义务 承担 连带 清偿 责任 七 案件 减半 收取 计元 由 上纽 资产 管理 陈刚 王俊 共同 负担 上纽 资产 管理 陈刚 王俊 负担 之款 应于 之前 直接 给付 原告 张显 爱 '
```
print(f'length of characters: {len(text_duty.duty[48670])}')
puncts = [' ']
def clean_text(x):
x=x.strip()
for punct in puncts:
x=x.replace(punct,'')
return x
def is_chinese(xchar):
if xchar>=u'\u4e00' and xchar<=u'\u9fa5':
return True
else:
return False
def keep_chinese_text(x):
out_str=''
for i in x:
if is_chinese(i):
out_str=out_str+i
return out_str
# def clean_text(text):
# wordlist = jieba.lcut(text)
# wordlist = [w for w in wordlist if w not in stopwords and len(w)>2]
# document = " ".join(wordlist)
# return document
def seg_sentence(sentence,stopwords):
sentence_seged=jieba.cut(sentence)
outstr=''
for word in sentence_seged:
if word not in stopwords:
outstr+=word
outstr+=" "
return outstr
text_duty["duty"]=text_duty["duty"].apply(lambda x:clean_text(x))
text_duty["duty"]=text_duty["duty"].apply(lambda x:keep_chinese_text(x))
# text_duty["duty"]=text_duty["duty"].apply(lambda x:clean_text(x))
text_duty["duty"]=text_duty["duty"].apply(lambda x:seg_sentence(x,stopwords))
text_duty.duty[48670]
text_duty['duty'].str.contains("被执行人")
```
# @TODO: Vectorize the tokens
## Token as feature -- lasso feature selection model
```
# import sklean vecorization;
vectorizer = CountVectorizer()
sh_sx_blacklisted = sh_sx[
(sh_sx.duty.str.contains("纳入"))
]
sh_sx_blacklisted.duty.loc[14638]
sh_sx['duration'] = sh_sx.publish_date - sh_sx.reg_date
sh_sx[['iname','reg_date', 'publish_date', 'duration', ]].sample(20)
len(sh_sx[sh_sx.case_code.str.contains("执恢")])
sh_sx[sh_sx.case_code.str.contains("执恢")][9049]
```
| github_jupyter |
# Decision Trees for You and Me!
## Binary Classification w/ the Tips dataset!
```
# ignore warnings
import warnings
warnings.filterwarnings("ignore")
import numpy as np
import pandas as pd
from pydataset import data
from sklearn.model_selection import train_test_split
from sklearn.tree import DecisionTreeClassifier
from sklearn.tree import export_graphviz
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
import matplotlib.pyplot as plt
import seaborn as sns
import graphviz
from graphviz import Graph
def split(df, stratify_by=None):
"""
Crude train, validate, test split
To stratify, send in a column name for the stratify_by argument
"""
if stratify_by == None:
train, test = train_test_split(df, test_size=.2, random_state=123)
train, validate = train_test_split(train, test_size=.3, random_state=123)
else:
train, test = train_test_split(df, test_size=.2, random_state=123, stratify=df[stratify_by])
train, validate = train_test_split(train, test_size=.3, random_state=123, stratify=train[stratify_by])
return train, validate, test
```
## Planning Stage
- Given diner table information, predict if the table will be a smoking table or not
- Input features are bill, tip, gender, day, time of day, and table size
- Target variable is smoker status
```
# Acquire
df = data("tips")
df.head()
# We don't need to scale the continuous input variables, since we're working with a decision tree
# Let's turn strings into booleans
df["is_female"] = df.sex == "Female"
df["is_dinner"] = df.time == "Dinner"
# We'll want to encode the day variable, since there are 4 possibilities (Thursday, Friday, Saturday, Sunday)
dummy_df = pd.get_dummies(df[["day"]], drop_first=True)
dummy_df
df = pd.concat([df, dummy_df], axis=1)
# drop the old columns
df = df.drop(columns=["sex", 'time', 'day'])
df.head()
# Split the data
# stratifying means we're making representative datasets between train, validate, test
train, validate, test = split(df, stratify_by="smoker")
train.head()
# Setup our X inputs and y target variable for each split
X_train = train.drop(columns=['smoker'])
y_train = train.smoker # labeled data == supervise algorithm
X_validate = validate.drop(columns=['smoker'])
y_validate = validate.smoker
X_test = test.drop(columns=['smoker'])
y_test = test.smoker
train.head()
# Let's generate a blank, new Decision Tree model
# Be sure to set the max_depth argument
# clf = DecisionTreeClassifier(max_depth=3, random_state=123)
clf = DecisionTreeClassifier(max_depth=2, random_state=123)
# Now let's train our model on the training data
# fitting == training the model
clf = clf.fit(X_train, y_train)
clf
# Visualize the model so iut can explain itself!
dot_data = export_graphviz(clf, feature_names= X_train.columns, rounded=True, filled=True, out_file=None)
graph = graphviz.Source(dot_data)
graph.render('tips_decision_tree', view=True, format="pdf")
# Now we'll make a set of predictions using this trained model
y_pred = clf.predict(X_train)
y_pred[0:3]
# Estimate the probabilities for each class
y_pred_proba = clf.predict_proba(X_train)
y_pred_proba[0:3]
y_train.head(3)
train["most_frequent"] = "No"
baseline_accuracy = (train.smoker == train.most_frequent).mean()
baseline_accuracy
# Let's evaluate the model
print('Accuracy of Decision Tree classifier on training set: {:.2f}'
.format(clf.score(X_train, y_train)))
print(classification_report(y_train, y_pred))
```
## Takeaways so far
- Pretty high accuracy on training data.
- But how does this model perform on out-of-sample data?
```
# clf was trained on X_train, y_train
# To evaluate the model trained on new data, the arguments coming into .score()
clf.score(X_validate, y_validate)
# Let's evaluate this model on out-of-sample data
print('Accuracy of Decision Tree classifier on validate set: {:.2f}'
.format(clf.score(X_validate, y_validate)))
# Use the classification model trained on train data to make predictions on validate data
y_pred = clf.predict(X_validate)
y_pred[0:3]
y_validate.head(3)
# Compare actual y values from validate to predictions based on X_validate
print(classification_report(y_validate, y_pred))
```
| github_jupyter |
# OHLC Charts in Python
```
import plotly
plotly.__version__
```
##### Simple OHLC Chart with Pandas
```
import plotly.plotly as py
from plotly.tools import FigureFactory as FF
from datetime import datetime
import pandas.io.data as web
df = web.DataReader("aapl", 'yahoo', datetime(2008, 8, 15), datetime(2008, 10, 15))
fig = FF.create_ohlc(df.Open, df.High, df.Low, df.Close, dates=df.index)
py.iplot(fig, filename='finance/aapl-ohlc')
```
##### Customizing the Figure with Text and Annotations
```
import plotly.plotly as py
from plotly.tools import FigureFactory as FF
import pandas.io.data as web
df = web.DataReader("aapl", 'yahoo', datetime(2008, 8, 15), datetime(2008, 10, 15))
fig = FF.create_ohlc(df.Open, df.High, df.Low, df.Close, dates=df.index)
# Update the fig - all options here: https://plot.ly/python/reference/#Layout
fig['layout'].update({
'title': 'The Great Recession',
'yaxis': {'title': 'AAPL Stock'},
'shapes': [{
'x0': '2008-09-15', 'x1': '2008-09-15', 'type': 'line',
'y0': 0, 'y1': 1, 'xref': 'x', 'yref': 'paper',
'line': {'color': 'rgb(40,40,40)', 'width': 0.5}
}],
'annotations': [{
'text': "the fall of Lehman Brothers",
'x': '2008-09-15', 'y': 1.02,
'xref': 'x', 'yref': 'paper',
'showarrow': False, 'xanchor': 'left'
}]
})
py.iplot(fig, filename='finance/aapl-recession-ohlc', validate=False)
```
##### Custom OHLC Chart Colors
```
import plotly.plotly as py
from plotly.tools import FigureFactory as FF
from plotly.graph_objs import Line, Marker
from datetime import datetime
import pandas.io.data as web
df = web.DataReader("aapl", 'yahoo', datetime(2008, 1, 1), datetime(2009, 4, 1))
# Make increasing ohlc sticks and customize their color and name
fig_increasing = FF.create_ohlc(df.Open, df.High, df.Low, df.Close, dates=df.index,
direction='increasing', name='AAPL',
line=Line(color='rgb(150, 200, 250)'))
# Make decreasing ohlc sticks and customize their color and name
fig_decreasing = FF.create_ohlc(df.Open, df.High, df.Low, df.Close, dates=df.index,
direction='decreasing',
line=Line(color='rgb(128, 128, 128)'))
# Initialize the figure
fig = fig_increasing
# Add decreasing data with .extend()
fig['data'].extend(fig_decreasing['data'])
py.iplot(fig, filename='finance/aapl-ohlc-colors', validate=False)
```
##### Simple Example with `datetime` Objects
```
import plotly.plotly as py
from plotly.tools import FigureFactory as FF
from datetime import datetime
# Add data
open_data = [33.0, 33.3, 33.5, 33.0, 34.1]
high_data = [33.1, 33.3, 33.6, 33.2, 34.8]
low_data = [32.7, 32.7, 32.8, 32.6, 32.8]
close_data = [33.0, 32.9, 33.3, 33.1, 33.1]
dates = [datetime(year=2013, month=10, day=10),
datetime(year=2013, month=11, day=10),
datetime(year=2013, month=12, day=10),
datetime(year=2014, month=1, day=10),
datetime(year=2014, month=2, day=10)]
# Create ohlc
fig = FF.create_ohlc(open_data, high_data,
low_data, close_data, dates=dates)
py.iplot(fig, filename='finance/simple-ohlc', validate=False)
```
###Open, High, Low, Close Charts in Cufflinks
```
import cufflinks as cf
import pandas as pd
cf.set_config_file(world_readable=True,offline=False)
# Open, High, Low, Close Data Generation
ohlc=cf.datagen.ohlc()
ohlc.head()
ohlc.iplot(kind='ohlc')
import cufflinks as cf
import pandas as pd
cf.set_config_file(world_readable=True,offline=False)
ohlc=cf.datagen.ohlc()
ohlc.iplot(kind='ohlc', up_color='rgb(20, 140, 200)', down_color='rgb(100, 100, 100)')
```
### Reference
```
help(FF.create_ohlc)
from IPython.display import display, HTML
display(HTML('<link href="//fonts.googleapis.com/css?family=Open+Sans:600,400,300,200|Inconsolata|Ubuntu+Mono:400,700" rel="stylesheet" type="text/css" />'))
display(HTML('<link rel="stylesheet" type="text/css" href="http://help.plot.ly/documentation/all_static/css/ipython-notebook-custom.css">'))
! pip install publisher --upgrade
import publisher
publisher.publish(
'ohlc-charts.ipynb', 'pandas/ohlc-charts/', 'OHLC Charts',
'How to make interactive OHLC charts in Python with Plotly. '
'Six examples of OHLC charts with Pandas, time series, and yahoo finance data.',
title = 'Python OHLC Charts | plotly',
thumbnail='/images/ohlc.png', language='pandas',
page_type='example_index', has_thumbnail='true', display_as='chart_type', order=20)
```
| github_jupyter |
Before you begin, execute this cell to import numpy and packages from the D-Wave Ocean suite, and all necessary functions the gate-model framework you are going to use, whether that is the Forest SDK or Qiskit. In the case of Forest SDK, it also starts the qvm and quilc servers.
```
%run -i "assignment_helper.py"
```
# Mapping clustering to discrete optimization
We will have $\{x_i\}_{i=1}^N$ points lying in two distant regions:
```
import matplotlib.pyplot as plt
%matplotlib inline
np.set_printoptions(precision=3, suppress=True)
np.random.seed(0)
# Generating the data
c1 = np.random.rand(5, 2)/5
c2 = (-0.6, 0.5) + np.random.rand(5, 2)/5
data = np.concatenate((c1, c2))
plt.subplot(111, xticks=[], yticks=[])
plt.scatter(data[:, 0], data[:, 1], color='navy')
```
**Exercise 1** (3 points). The unsupervised task is identify this structure. The challenge is that if we have a high number of points compared to the number of qubits and their connections, we won't be able to solve the problem. So if we blindly follow the idea of mapping the problem to max-cut using the Eucledian distance and then solve it by some quantum optimization method, we would run out of qubit connections for this problem. We can, however, choose to be clever about how we calculate the distances classically. This is a common trick: since we are doing hybrid classical-quantum algorithms anyway, we might as well do the classical parts wisely.
In this case, we might rely on some form of [exploratory data analysis](https://en.wikipedia.org/wiki/Exploratory_data_analysis) to get a sense of the distances in the dataset. This is an easy case, since we just plotted the two-dimensional data, and we see that the density within a blob is high. You can make the task easier by thresholding the distances. Use any distance function you wish (even the Euclidean distance will do) and set distances to zero that are below a threshold you calculate. The threshold should be such that all of the distances within a blob will be set to zero. Store the symmetrix distance matrix in a two-dimensional numpy array called `w`.
```
import itertools
n_instances = data.shape[0]
###
### YOUR CODE HERE
###
w = np.zeros((n_instances, n_instances))
for i, j in itertools.product(*[range(n_instances)]*2):
dist_ij = np.linalg.norm(data[i]-data[j])
w[i, j] = np.linalg.norm(data[i]-data[j]) * (dist_ij>0.5)
labels = np.array([0, 0, 0, 0, 0, 1, 1, 1, 1, 1])
assert np.alltrue(w == w.T)
assert np.alltrue(w >= 0)
assert (w[labels==0][:, labels==0]==0).sum() == 25
assert (w[labels==1][:, labels==1]==0).sum() == 25
```
# Solving the max-cut problem by QAOA
**Exercise 2** (2 points). From here, it is rather formulaic to create the max-cut as an Ising Hamiltonian and solve either by annealing or by some variational circuit like QAOA. Set up the QAOA problem in your chosen framework and with an optimization function. Import all necessary functions you need. Call the object `qaoa`. We set the `p` parameter of the QAOA to 1.
```
p = 1
###
### YOUR CODE HERE
###
from pyquil import Program, api
from pyquil.paulis import PauliSum, PauliTerm
from scipy.optimize import fmin_bfgs
from grove.pyqaoa.qaoa import QAOA
from forest_tools import *
qvm_server, quilc_server, fc = init_qvm_and_quilc('/home/local/bin/qvm',
'/home/local/bin/quilc')
qvm = api.QVMConnection(endpoint=fc.sync_endpoint,
compiler_endpoint=fc.compiler_endpoint)
maxcut = []
for i in range(n_instances):
for j in range(i+1, n_instances):
maxcut.append(PauliSum([PauliTerm("Z", i, 1/4 * w[i, j]) *
PauliTerm("Z", j, 1.0)]))
maxcut.append(PauliSum([PauliTerm("I", i, -1/4)]))
Hm = [PauliSum([PauliTerm("X", i, 1.0)]) for i in range(n_instances)]
qaoa = QAOA(qvm,
qubits=range(n_instances),
steps=p,
ref_ham=Hm,
cost_ham=maxcut,
store_basis=True,
minimizer=fmin_bfgs,
minimizer_kwargs={'maxiter': 50})
```
Setting $p=1$ in the QAOA algorithm, we can initialize it with the max-cut problem.
```
if isinstance(qaoa, qiskit_aqua.algorithms.adaptive.qaoa.qaoa.QAOA):
assert qaoa._operator.num_qubits == 10
elif isinstance(qaoa, grove.pyqaoa.qaoa.QAOA):
assert len(qaoa.ref_ham) == 10
assert all([qaoa.ref_ham[i].terms[0].compact_str() == '(1+0j)*X{}'.format(i) for i in range(10)])
assert len(qaoa.cost_ham) == 90
else:
raise ValueError("Unknown type for Hamiltonian!")
```
**Exercise 3** (2 points). Run the QAOA on a simulator. Store the outcome in an object called `result`. In the case of the Forest SDK, this will store the return value of the `run_and_measure` method of the QVM. In the case of Qiskit, it will be the return value of the `run` method of the `qaoa` object.
```
###
### YOUR CODE HERE
###
ν, γ = qaoa.get_angles()
program = qaoa.get_parameterized_program()(np.hstack((ν, γ)))
result = qvm.run_and_measure(program, range(n_instances), trials=100)
ground_truth1 = [0, 0, 0, 0, 0, 1, 1, 1, 1, 1]
ground_truth2 = [1, 1, 1, 1, 1, 0, 0, 0, 0, 0]
if isinstance(result, list):
count = np.unique(result, return_counts=True, axis=0)
solution = count[0][np.argmax(count[1])]
assert abs(sum(solution == ground_truth1)-10) <= 1 or abs(sum(solution == ground_truth2)-10)
else:
x = maxcut.sample_most_likely(result['eigvecs'][0])
solution = maxcut.get_graph_solution(x)
assert abs(sum(solution == ground_truth1)-10) <= 1 or abs(sum(solution == ground_truth2)-10)
```
| github_jupyter |
##### Copyright 2020 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Parameter server training with ParameterServerStrategy
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/tutorials/distribute/parameter_server_training"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/tutorials/distribute/parameter_server_training.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/tutorials/distribute/parameter_server_training.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/tutorials/distribute/parameter_server_training.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
</td>
</table>
## Overview
[Parameter server training](https://www.usenix.org/system/files/conference/osdi14/osdi14-paper-li_mu.pdf) is a common data-parallel method to scale up model training on multiple machines.
A parameter server training cluster consists of _workers_ and _parameter servers_. Variables are created on parameter servers and they are read and updated by workers in each step. By default, workers read and update these variables independently without synchronizing with each other. This is why sometimes parameter server-style training is called _asynchronous training_.
In TensorFlow 2, parameter server training is powered by the `tf.distribute.experimental.ParameterServerStrategy` class, which distributes the training steps to a cluster that scales up to thousands of workers (accompanied by parameter servers).
### Supported training methods
There are two main supported training methods:
- The Keras `Model.fit` API, which is recommended when you prefer a high-level abstraction and handling of training.
- A custom training loop (you can refer to [Custom training](https://www.tensorflow.org/tutorials/customization/custom_training_walkthrough#train_the_model), [Writing a training loop from scratch
](https://www.tensorflow.org/guide/keras/writing_a_training_loop_from_scratch) and [Custom training loop with Keras and MultiWorkerMirroredStrategy](https://www.tensorflow.org/tutorials/distribute/multi_worker_with_ctl) for more details.) Custom loop training is recommended when you prefer to define the details of their training loop.
### A cluster with jobs and tasks
Regardless of the API of choice (`Model.fit` or a custom training loop), distributed training in TensorFlow 2 involves: a `'cluster'` with several `'jobs'`, and each of the jobs may have one or more `'tasks'`.
When using parameter server training, it is recommended to have:
- One _coordinator_ job (which has the job name `chief`)
- Multiple _worker_ jobs (job name `worker`); and
- Multiple _parameter server_ jobs (job name `ps`)
While the _coordinator_ creates resources, dispatches training tasks, writes checkpoints, and deals with task failures, _workers_ and _parameter servers_ run `tf.distribute.Server` that listen for requests from the coordinator.
### Parameter server training with `Model.fit` API
Parameter server training with the `Model.fit` API requires the coordinator to use a `tf.distribute.experimental.ParameterServerStrategy` object, and a `tf.keras.utils.experimental.DatasetCreator` as the input. Similar to `Model.fit` usage with no strategy, or with other strategies, the workflow involves creating and compiling the model, preparing the callbacks, followed by
a `Model.fit` call.
### Parameter server training with a custom training loop
With custom training loops, the `tf.distribute.experimental.coordinator.ClusterCoordinator` class is the key component used for the coordinator.
- The `ClusterCoordinator` class needs to work in conjunction with a `tf.distribute.Strategy` object.
- This `tf.distribute.Strategy` object is needed to provide the information of the cluster and is used to define a training step, as demonstrated in [Custom training with tf.distribute.Strategy](https://www.tensorflow.org/tutorials/distribute/custom_training#training_loop).
- The `ClusterCoordinator` object then dispatches the execution of these training steps to remote workers.
- For parameter server training, the `ClusterCoordinator` needs to work with a `tf.distribute.experimental.ParameterServerStrategy`.
The most important API provided by the `ClusterCoordinator` object is `schedule`:
- The `schedule` API enqueues a `tf.function` and returns a future-like `RemoteValue` immediately.
- The queued functions will be dispatched to remote workers in background threads and their `RemoteValue`s will be filled asynchronously.
- Since `schedule` doesn’t require worker assignment, the `tf.function` passed in can be executed on any available worker.
- If the worker it is executed on becomes unavailable before its completion, the function will be retried on another available worker.
- Because of this fact and the fact that function execution is not atomic, a function may be executed more than once.
In addition to dispatching remote functions, the `ClusterCoordinator` also helps
to create datasets on all the workers and rebuild these datasets when a worker recovers from failure.
## Tutorial setup
The tutorial will branch into `Model.fit` and custom training loop paths, and you can choose the one that fits your needs. Sections other than "Training with X" are applicable to both paths.
```
!pip install portpicker
!pip uninstall tensorflow keras -y
!pip install tf-nightly
#@title
import multiprocessing
import os
import random
import portpicker
import tensorflow as tf
```
## Cluster setup
As mentioned above, a parameter server training cluster requires a coordinator task that runs your training program, one or several workers and parameter server tasks that run TensorFlow servers—`tf.distribute.Server`—and possibly an additional evaluation task that runs side-car evaluation (see the side-car evaluation section below). The requirements to set them up are:
- The coordinator task needs to know the addresses and ports of all other TensorFlow servers except the evaluator.
- The workers and parameter servers need to know which port they need to listen to. For the sake of simplicity, you can usually pass in the complete cluster information when creating TensorFlow servers on these tasks.
- The evaluator task doesn’t have to know the setup of the training cluster. If it does, it should not attempt to connect to the training cluster.
- Workers and parameter servers should have task types as `"worker"` and `"ps"`, respectively. The coordinator should use `"chief"` as the task type for legacy reasons.
In this tutorial, you will create an in-process cluster so that the whole parameter server training can be run in Colab. You will learn how to set up [real clusters](#real_clusters) in a later section.
### In-process cluster
You will start by creating several TensorFlow servers in advance and connect to them later. Note that this is only for the purpose of this tutorial's demonstration, and in real training the servers will be started on `"worker"` and `"ps"` machines.
```
def create_in_process_cluster(num_workers, num_ps):
"""Creates and starts local servers and returns the cluster_resolver."""
worker_ports = [portpicker.pick_unused_port() for _ in range(num_workers)]
ps_ports = [portpicker.pick_unused_port() for _ in range(num_ps)]
cluster_dict = {}
cluster_dict["worker"] = ["localhost:%s" % port for port in worker_ports]
if num_ps > 0:
cluster_dict["ps"] = ["localhost:%s" % port for port in ps_ports]
cluster_spec = tf.train.ClusterSpec(cluster_dict)
# Workers need some inter_ops threads to work properly.
worker_config = tf.compat.v1.ConfigProto()
if multiprocessing.cpu_count() < num_workers + 1:
worker_config.inter_op_parallelism_threads = num_workers + 1
for i in range(num_workers):
tf.distribute.Server(
cluster_spec,
job_name="worker",
task_index=i,
config=worker_config,
protocol="grpc")
for i in range(num_ps):
tf.distribute.Server(
cluster_spec,
job_name="ps",
task_index=i,
protocol="grpc")
cluster_resolver = tf.distribute.cluster_resolver.SimpleClusterResolver(
cluster_spec, rpc_layer="grpc")
return cluster_resolver
# Set the environment variable to allow reporting worker and ps failure to the
# coordinator. This is a workaround and won't be necessary in the future.
os.environ["GRPC_FAIL_FAST"] = "use_caller"
NUM_WORKERS = 3
NUM_PS = 2
cluster_resolver = create_in_process_cluster(NUM_WORKERS, NUM_PS)
```
The in-process cluster setup is frequently used in unit testing, such as [here](https://github.com/tensorflow/tensorflow/blob/7621d31921c2ed979f212da066631ddfda37adf5/tensorflow/python/distribute/coordinator/cluster_coordinator_test.py#L437).
Another option for local testing is to launch processes on the local machine—check out [Multi-worker training with Keras](https://www.tensorflow.org/tutorials/distribute/multi_worker_with_keras) for an example of this approach.
## Instantiate a ParameterServerStrategy
Before you dive into the training code, let's instantiate a `ParameterServerStrategy` object. Note that this is needed regardless of whether you are proceeding with `Model.fit` or a custom training loop. The `variable_partitioner` argument will be explained in the [Variable sharding section](#variable-sharding).
```
variable_partitioner = (
tf.distribute.experimental.partitioners.MinSizePartitioner(
min_shard_bytes=(256 << 10),
max_shards=NUM_PS))
strategy = tf.distribute.experimental.ParameterServerStrategy(
cluster_resolver,
variable_partitioner=variable_partitioner)
```
In order to use GPUs for training, allocate GPUs visible to each worker. `ParameterServerStrategy` will use all the available GPUs on each worker, with the restriction that all workers should have the same number of GPUs available.
### Variable sharding
Variable sharding refers to splitting a variable into multiple smaller
variables, which are called _shards_. Variable sharding may be useful to distribute the network load when accessing these shards. It is also useful to distribute computation and storage of a normal variable across multiple parameter servers.
To enable variable sharding, you can pass in a `variable_partitioner` when
constructing a `ParameterServerStrategy` object. The `variable_partitioner` will
be invoked every time when a variable is created and it is expected to return
the number of shards along each dimension of the variable. Some out-of-box
`variable_partitioner`s are provided such as
`tf.distribute.experimental.partitioners.MinSizePartitioner`. It is recommended to use size-based partitioners like
`tf.distribute.experimental.partitioners.MinSizePartitioner` to avoid
partitioning small variables, which could have negative impact on model training
speed.
When a `variable_partitioner` is passed in and if you create a variable directly
under `strategy.scope()`, it will become a container type with a `variables`
property which provides access to the list of shards. In most cases, this
container will be automatically converted to a Tensor by concatenating all the
shards. As a result, it can be used as a normal variable. On the other hand,
some TensorFlow methods such as `tf.nn.embedding_lookup` provide efficient
implementation for this container type and in these methods automatic
concatenation will be avoided.
Please see the API docs of `tf.distribute.experimental.ParameterServerStrategy` for more details.
## Training with `Model.fit`
<a id="training_with_modelfit"></a>
Keras provides an easy-to-use training API via `Model.fit` that handles the training loop under the hood, with the flexibility of overridable `train_step`, and callbacks, which provide functionalities such as checkpoint saving or summary saving for TensorBoard. With `Model.fit`, the same training code can be used for other strategies with a simple swap of the strategy object.
### Input data
`Model.fit` with parameter server training requires that the input data be
provided in a callable that takes a single argument of type `tf.distribute.InputContext`, and returns a `tf.data.Dataset`. Then, create a `tf.keras.utils.experimental.DatasetCreator` object that takes such `callable`, and an optional `tf.distribute.InputOptions` object via `input_options` argument.
Note that it is recommended to shuffle and repeat the data with parameter server training, and specify `steps_per_epoch` in `fit` call so the library knows the epoch boundaries.
Please see the [Distributed input](https://www.tensorflow.org/tutorials/distribute/input#usage_2) tutorial for more information about the `InputContext` argument.
```
def dataset_fn(input_context):
global_batch_size = 64
batch_size = input_context.get_per_replica_batch_size(global_batch_size)
x = tf.random.uniform((10, 10))
y = tf.random.uniform((10,))
dataset = tf.data.Dataset.from_tensor_slices((x, y)).shuffle(10).repeat()
dataset = dataset.shard(
input_context.num_input_pipelines,
input_context.input_pipeline_id)
dataset = dataset.batch(batch_size)
dataset = dataset.prefetch(2)
return dataset
dc = tf.keras.utils.experimental.DatasetCreator(dataset_fn)
```
The code in `dataset_fn` will be invoked on the input device, which is usually the CPU, on each of the worker machines.
### Model construction and compiling
Now, you will create a `tf.keras.Model`—a trivial `tf.keras.models.Sequential` model for demonstration purposes—followed by a `Model.compile` call to incorporate components, such as an optimizer, metrics, or parameters such as `steps_per_execution`:
```
with strategy.scope():
model = tf.keras.models.Sequential([tf.keras.layers.Dense(10)])
model.compile(tf.keras.optimizers.SGD(), loss='mse', steps_per_execution=10)
```
### Callbacks and training
<a id="callbacks-and-training"> </a>
Before you call `model.fit` for the actual training, let's prepare the needed callbacks for common tasks, such as:
- `ModelCheckpoint`: to save the model weights.
- `BackupAndRestore`: to make sure the training progress is automatically backed up, and recovered if the cluster experiences unavailability (such as abort or preemption); or
- `TensorBoard`: to save the progress reports into summary files, which get visualized in TensorBoard tool.
Note: Due to performance consideration, custom callbacks cannot have batch level callbacks overridden when used with `ParameterServerStrategy`. Please modify your custom callbacks to make them epoch level calls, and adjust `steps_per_epoch` to a suitable value. In addition, `steps_per_epoch` is a required argument for `Model.fit` when used with `ParameterServerStrategy`.
```
working_dir = '/tmp/my_working_dir'
log_dir = os.path.join(working_dir, 'log')
ckpt_filepath = os.path.join(working_dir, 'ckpt')
backup_dir = os.path.join(working_dir, 'backup')
callbacks = [
tf.keras.callbacks.TensorBoard(log_dir=log_dir),
tf.keras.callbacks.ModelCheckpoint(filepath=ckpt_filepath),
tf.keras.callbacks.experimental.BackupAndRestore(backup_dir=backup_dir),
]
model.fit(dc, epochs=5, steps_per_epoch=20, callbacks=callbacks)
```
### Direct usage with `ClusterCoordinator` (optional)
Even if you choose the `Model.fit` training path, you can optionally instantiate a `tf.distribute.experimental.coordinator.ClusterCoordinator` object to schedule other functions you would like to be executed on the workers. See the [Training with a custom training loop](#training_with_custom_training_loop) section for more details and examples.
## Training with a custom training loop
<a id="training_with_custom_training_loop"> </a>
Using custom training loops with `tf.distribute.Strategy` provides great flexibility to define training loops. With the `ParameterServerStrategy` defined above (as `strategy`), you will use a `tf.distribute.experimental.coordinator.ClusterCoordinator` to dispatch the execution of training steps to remote workers.
Then, you will create a model, define a dataset and a step function, as you have done in the training loop with other `tf.distribute.Strategy`s. You can find more details in the [Custom training with tf.distribute.Strategy](https://www.tensorflow.org/tutorials/distribute/custom_training) tutorial.
To ensure efficient dataset prefetching, use the recommended distributed dataset creation APIs mentioned in the [Dispatch training steps to remote workers](https://www.tensorflow.org/tutorials/distribute/parameter_server_training#dispatch_training_steps_to_remote_workers) section below. Also, make sure to call `Strategy.run` inside `worker_fn` to take full advantage of GPUs allocated to workers. The rest of the steps are the same for training with or without GPUs.
Let’s create these components in the following steps:
### Set up the data
First, write a function that creates a dataset that includes preprocessing logic implemented by [Keras preprocessing layers](https://www.tensorflow.org/guide/keras/preprocessing_layers).
You will create these layers outside the `dataset_fn` but apply the transformation inside the `dataset_fn`, since you will wrap the `dataset_fn` into a `tf.function`, which doesn't allow variables to be created inside it.
Note: There is a known performance implication when using lookup table resources, which layers, such as `tf.keras.layers.StringLookup`, employ. Refer to the [Known limitations](#known_limitations) section for more information.
```
feature_vocab = [
"avenger", "ironman", "batman", "hulk", "spiderman", "kingkong", "wonder_woman"
]
label_vocab = ["yes", "no"]
with strategy.scope():
feature_lookup_layer = tf.keras.layers.StringLookup(
vocabulary=feature_vocab,
mask_token=None)
label_lookup_layer = tf.keras.layers.StringLookup(
vocabulary=label_vocab,
num_oov_indices=0,
mask_token=None)
raw_feature_input = tf.keras.layers.Input(
shape=(3,),
dtype=tf.string,
name="feature")
feature_id_input = feature_lookup_layer(raw_feature_input)
feature_preprocess_stage = tf.keras.Model(
{"features": raw_feature_input},
feature_id_input)
raw_label_input = tf.keras.layers.Input(
shape=(1,),
dtype=tf.string,
name="label")
label_id_input = label_lookup_layer(raw_label_input)
label_preprocess_stage = tf.keras.Model(
{"label": raw_label_input},
label_id_input)
```
Generate toy examples in a dataset:
```
def feature_and_label_gen(num_examples=200):
examples = {"features": [], "label": []}
for _ in range(num_examples):
features = random.sample(feature_vocab, 3)
label = ["yes"] if "avenger" in features else ["no"]
examples["features"].append(features)
examples["label"].append(label)
return examples
examples = feature_and_label_gen()
```
Then, create the training dataset wrapped in a `dataset_fn`:
```
def dataset_fn(_):
raw_dataset = tf.data.Dataset.from_tensor_slices(examples)
train_dataset = raw_dataset.map(
lambda x: (
{"features": feature_preprocess_stage(x["features"])},
label_preprocess_stage(x["label"])
)).shuffle(200).batch(32).repeat()
return train_dataset
```
### Build the model
Next, create the model and other objects. Make sure to create all variables under `strategy.scope`.
```
# These variables created under the `strategy.scope` will be placed on parameter
# servers in a round-robin fashion.
with strategy.scope():
# Create the model. The input needs to be compatible with Keras processing layers.
model_input = tf.keras.layers.Input(
shape=(3,), dtype=tf.int64, name="model_input")
emb_layer = tf.keras.layers.Embedding(
input_dim=len(feature_lookup_layer.get_vocabulary()), output_dim=16384)
emb_output = tf.reduce_mean(emb_layer(model_input), axis=1)
dense_output = tf.keras.layers.Dense(units=1, activation="sigmoid")(emb_output)
model = tf.keras.Model({"features": model_input}, dense_output)
optimizer = tf.keras.optimizers.RMSprop(learning_rate=0.1)
accuracy = tf.keras.metrics.Accuracy()
```
Let's confirm that the use of `FixedShardsPartitioner` split all variables into two shards and each shard was assigned to different parameter servers:
```
assert len(emb_layer.weights) == 2
assert emb_layer.weights[0].shape == (4, 16384)
assert emb_layer.weights[1].shape == (4, 16384)
assert emb_layer.weights[0].device == "/job:ps/replica:0/task:0/device:CPU:0"
assert emb_layer.weights[1].device == "/job:ps/replica:0/task:1/device:CPU:0"
```
### Define the training step
Third, create the training step wrapped into a `tf.function`:
```
@tf.function
def step_fn(iterator):
def replica_fn(batch_data, labels):
with tf.GradientTape() as tape:
pred = model(batch_data, training=True)
per_example_loss = tf.keras.losses.BinaryCrossentropy(
reduction=tf.keras.losses.Reduction.NONE)(labels, pred)
loss = tf.nn.compute_average_loss(per_example_loss)
gradients = tape.gradient(loss, model.trainable_variables)
optimizer.apply_gradients(zip(gradients, model.trainable_variables))
actual_pred = tf.cast(tf.greater(pred, 0.5), tf.int64)
accuracy.update_state(labels, actual_pred)
return loss
batch_data, labels = next(iterator)
losses = strategy.run(replica_fn, args=(batch_data, labels))
return strategy.reduce(tf.distribute.ReduceOp.SUM, losses, axis=None)
```
In the above training step function, calling `Strategy.run` and `Strategy.reduce` in the `step_fn` can support multiple GPUs per worker. If the workers have GPUs allocated, `Strategy.run` will distribute the datasets on multiple replicas.
### Dispatch training steps to remote workers
<a id="dispatch_training_steps_to_remote_workers"> </a>
After all the computations are defined by `ParameterServerStrategy`, you will use the `tf.distribute.experimental.coordinator.ClusterCoordinator` class to create resources and distribute the training steps to remote workers.
Let’s first create a `ClusterCoordinator` object and pass in the strategy object:
```
coordinator = tf.distribute.experimental.coordinator.ClusterCoordinator(strategy)
```
Then, create a per-worker dataset and an iterator. In the `per_worker_dataset_fn` below, wrapping the `dataset_fn` into `strategy.distribute_datasets_from_function` is recommended to allow efficient prefetching to GPUs seamlessly.
```
@tf.function
def per_worker_dataset_fn():
return strategy.distribute_datasets_from_function(dataset_fn)
per_worker_dataset = coordinator.create_per_worker_dataset(per_worker_dataset_fn)
per_worker_iterator = iter(per_worker_dataset)
```
The final step is to distribute the computation to remote workers using `ClusterCoordinator.schedule`:
- The `schedule` method enqueues a `tf.function` and returns a future-like `RemoteValue` immediately. The queued functions will be dispatched to remote workers in background threads and the `RemoteValue` will be filled asynchronously.
- The `join` method (`ClusterCoordinator.join`) can be used to wait until all scheduled functions are executed.
```
num_epoches = 4
steps_per_epoch = 5
for i in range(num_epoches):
accuracy.reset_states()
for _ in range(steps_per_epoch):
coordinator.schedule(step_fn, args=(per_worker_iterator,))
# Wait at epoch boundaries.
coordinator.join()
print ("Finished epoch %d, accuracy is %f." % (i, accuracy.result().numpy()))
```
Here is how you can fetch the result of a `RemoteValue`:
```
loss = coordinator.schedule(step_fn, args=(per_worker_iterator,))
print ("Final loss is %f" % loss.fetch())
```
Alternatively, you can launch all steps and do something while waiting for
completion:
```python
for _ in range(total_steps):
coordinator.schedule(step_fn, args=(per_worker_iterator,))
while not coordinator.done():
time.sleep(10)
# Do something like logging metrics or writing checkpoints.
```
For the complete training and serving workflow for this particular example, please check out this [test](https://github.com/tensorflow/tensorflow/blob/master/tensorflow/python/keras/distribute/parameter_server_training_test.py).
### More about dataset creation
The dataset in the above code is created using the `ClusterCoordinator.create_per_worker_dataset` API). It creates one dataset per worker and returns a container object. You can call the `iter` method on it to create a per-worker iterator. The per-worker iterator contains one iterator per worker and the corresponding slice of a worker will be substituted in the input argument of the function passed to the `ClusterCoordinator.schedule` method before the function is executed on a particular worker.
Currently, the `ClusterCoordinator.schedule` method assumes workers are equivalent and thus assumes the datasets on different workers are the same except they may be shuffled differently if they contain a `Dataset.shuffle` operation. Because of this, it is also recommended that the datasets to be repeated indefinitely and you schedule a finite number of steps instead of relying on the `OutOfRangeError` from a dataset.
Another important note is that `tf.data` datasets don’t support implicit serialization and deserialization across task boundaries. So it is important to create the whole dataset inside the function passed to `ClusterCoordinator.create_per_worker_dataset`.
## Evaluation
There is more than one way to define and run an evaluation loop in distributed training. Each has its own pros and cons as described below. The inline evaluation method is recommended if you don't have a preference.
### Inline evaluation
In this method, the coordinator alternates between training and evaluation and thus it is called it _inline evaluation_.
There are several benefits of inline evaluation. For example:
- It can support large evaluation models and evaluation datasets that a single task cannot hold.
- The evaluation results can be used to make decisions for training the next epoch.
There are two ways to implement inline evaluation: direct evaluation and distributed evaluation.
- **Direct evaluation**: For small models and evaluation datasets, the coordinator can run evaluation directly on the distributed model with the evaluation dataset on the coordinator:
```
eval_dataset = tf.data.Dataset.from_tensor_slices(
feature_and_label_gen(num_examples=16)).map(
lambda x: (
{"features": feature_preprocess_stage(x["features"])},
label_preprocess_stage(x["label"])
)).batch(8)
eval_accuracy = tf.keras.metrics.Accuracy()
for batch_data, labels in eval_dataset:
pred = model(batch_data, training=False)
actual_pred = tf.cast(tf.greater(pred, 0.5), tf.int64)
eval_accuracy.update_state(labels, actual_pred)
print ("Evaluation accuracy: %f" % eval_accuracy.result())
```
- **Distributed evaluation**: For large models or datasets that are infeasible to run directly on the coordinator, the coordinator task can distribute evaluation tasks to the workers via the `ClusterCoordinator.schedule`/`ClusterCoordinator.join` methods:
```
with strategy.scope():
# Define the eval metric on parameter servers.
eval_accuracy = tf.keras.metrics.Accuracy()
@tf.function
def eval_step(iterator):
def replica_fn(batch_data, labels):
pred = model(batch_data, training=False)
actual_pred = tf.cast(tf.greater(pred, 0.5), tf.int64)
eval_accuracy.update_state(labels, actual_pred)
batch_data, labels = next(iterator)
strategy.run(replica_fn, args=(batch_data, labels))
def eval_dataset_fn():
return tf.data.Dataset.from_tensor_slices(
feature_and_label_gen(num_examples=16)).map(
lambda x: (
{"features": feature_preprocess_stage(x["features"])},
label_preprocess_stage(x["label"])
)).shuffle(16).repeat().batch(8)
per_worker_eval_dataset = coordinator.create_per_worker_dataset(eval_dataset_fn)
per_worker_eval_iterator = iter(per_worker_eval_dataset)
eval_steps_per_epoch = 2
for _ in range(eval_steps_per_epoch):
coordinator.schedule(eval_step, args=(per_worker_eval_iterator,))
coordinator.join()
print ("Evaluation accuracy: %f" % eval_accuracy.result())
```
Note: Currently, the `schedule` and `join` methods of `tf.distribute.experimental.coordinator.ClusterCoordinator` don’t support visitation guarantee or exactly-once semantics. In other words, there is no guarantee that all evaluation examples in a dataset will be evaluated exactly once; some may not be visited and some may be evaluated multiple times. Visitation guarantee on evaluation dataset is being worked on.
### Side-car evaluation
Another method is called _side-car evaluation_ where you create a dedicated evaluator task that repeatedly reads checkpoints and runs evaluation on a latest checkpoint. It allows your training program to finish early if you don't need to change your training loop based on evaluation results. However, it requires an additional evaluator task and periodic checkpointing to trigger evaluation. Following is a possible side-car evaluation loop:
```python
checkpoint_dir = ...
eval_model = ...
eval_data = ...
checkpoint = tf.train.Checkpoint(model=eval_model)
for latest_checkpoint in tf.train.checkpoints_iterator(
checkpoint_dir):
try:
checkpoint.restore(latest_checkpoint).expect_partial()
except (tf.errors.OpError,) as e:
# checkpoint may be deleted by training when it is about to read it.
continue
# Optionally add callbacks to write summaries.
eval_model.evaluate(eval_data)
# Evaluation finishes when it has evaluated the last epoch.
if latest_checkpoint.endswith('-{}'.format(train_epoches)):
break
```
## Clusters in the real world
<a id="real_clusters"></a>
Note: this section is not necessary for running the tutorial code in this page.
In a real production environment, you will run all tasks in different processes on different machines. The simplest way to configure cluster information on each task is to set `"TF_CONFIG"` environment variables and use a `tf.distribute.cluster_resolver.TFConfigClusterResolver` to parse `"TF_CONFIG"`.
For a general description about `"TF_CONFIG"` environment variables, refer to the [Distributed training](https://www.tensorflow.org/guide/distributed_training#setting_up_tf_config_environment_variable) guide.
If you start your training tasks using Kubernetes or other configuration templates, it is very likely that these templates have already set `“TF_CONFIG"` for you.
### Set the `"TF_CONFIG"` environment variable
Suppose you have 3 workers and 2 parameter servers, the `"TF_CONFIG"` of worker 1 can be:
```python
os.environ["TF_CONFIG"] = json.dumps({
"cluster": {
"worker": ["host1:port", "host2:port", "host3:port"],
"ps": ["host4:port", "host5:port"],
"chief": ["host6:port"]
},
"task": {"type": "worker", "index": 1}
})
```
The `"TF_CONFIG"` of the evaluator can be:
```python
os.environ["TF_CONFIG"] = json.dumps({
"cluster": {
"evaluator": ["host7:port"]
},
"task": {"type": "evaluator", "index": 0}
})
```
The `"cluster"` part in the above `"TF_CONFIG"` string for the evaluator is optional.
### If you use the same binary for all tasks
If you prefer to run all these tasks using a single binary, you will need to let your program branch into different roles at the very beginning:
```python
cluster_resolver = tf.distribute.cluster_resolver.TFConfigClusterResolver()
if cluster_resolver.task_type in ("worker", "ps"):
# Start a TensorFlow server and wait.
elif cluster_resolver.task_type == "evaluator":
# Run side-car evaluation
else:
# Run the coordinator.
```
The following code starts a TensorFlow server and waits:
```python
# Set the environment variable to allow reporting worker and ps failure to the
# coordinator. This is a workaround and won't be necessary in the future.
os.environ["GRPC_FAIL_FAST"] = "use_caller"
server = tf.distribute.Server(
cluster_resolver.cluster_spec(),
job_name=cluster_resolver.task_type,
task_index=cluster_resolver.task_id,
protocol=cluster_resolver.rpc_layer or "grpc",
start=True)
server.join()
```
## Handling task failure
### Worker failure
`tf.distribute.experimental.coordinator.ClusterCoordinator` or `Model.fit` provide built-in fault tolerance for worker failure. Upon worker recovery, the previously provided dataset function (either to `ClusterCoordinator.create_per_worker_dataset` for a custom training loop, or `tf.keras.utils.experimental.DatasetCreator` for `Model.fit`) will be invoked on the workers to re-create the datasets.
### Parameter server or coordinator failure
However, when the coordinator sees a parameter server error, it will raise an `UnavailableError` or `AbortedError` immediately. You can restart the coordinator in this case. The coordinator itself can also become unavailable. Therefore, certain tooling is recommended in order to not lose the training progress:
- For `Model.fit`, you should use a `BackupAndRestore` callback, which handles the progress saving and restoration automatically. See [Callbacks and training](#callbacks-and-training) section above for an example.
- For a custom training loop, you should checkpoint the model variables periodically and load model variables from a checkpoint, if any, before training starts. The training progress can be inferred approximately from `optimizer.iterations` if an optimizer is checkpointed:
```python
checkpoint_manager = tf.train.CheckpointManager(
tf.train.Checkpoint(model=model, optimizer=optimizer),
checkpoint_dir,
max_to_keep=3)
if checkpoint_manager.latest_checkpoint:
checkpoint = checkpoint_manager.checkpoint
checkpoint.restore(
checkpoint_manager.latest_checkpoint).assert_existing_objects_matched()
global_steps = int(optimizer.iterations.numpy())
starting_epoch = global_steps // steps_per_epoch
for _ in range(starting_epoch, num_epoches):
for _ in range(steps_per_epoch):
coordinator.schedule(step_fn, args=(per_worker_iterator,))
coordinator.join()
checkpoint_manager.save()
```
### Fetching a `RemoteValue`
Fetching a `RemoteValue` is guaranteed to succeed if a function is executed successfully. This is because currently the return value is immediately copied to the coordinator after a function is executed. If there is any worker failure during the copy, the function will be retried on another available worker. Therefore, if you want to optimize for performance, you can schedule functions without a return value.
## Error reporting
Once the coordinator sees an error such as `UnavailableError` from parameter servers or other application errors such as an `InvalidArgument` from `tf.debugging.check_numerics`, it will cancel all pending and queued functions before raising the error. Fetching their corresponding `RemoteValue`s will raise a `CancelledError`.
After an error is raised, the coordinator will not raise the same error or any error from cancelled functions.
## Performance improvement
There are several possible reasons if you see performance issues when you train with `ParameterServerStrategy` and `ClusterResolver`.
One common reason is parameter servers have unbalanced load and some heavily-loaded parameter servers have reached capacity. There can also be multiple root causes. Some simple methods to mitigate this issue are to:
1. Shard your large model variables via specifying a `variable_partitioner` when constructing a `ParameterServerStrategy`.
2. Avoid creating a hotspot variable that is required by all parameter servers in a single step if possible. For example, use a constant learning rate or subclass `tf.keras.optimizers.schedules.LearningRateSchedule` in optimizers since the default behavior is that the learning rate will become a variable placed on a particular parameter server and requested by all other parameter servers in each step.
3. Shuffle your large vocabularies before passing them to Keras preprocessing layers.
Another possible reason for performance issues is the coordinator. Your first implementation of `schedule`/`join` is Python-based and thus may have threading overhead. Also the latency between the coordinator and the workers can be large. If this is the case,
- For `Model.fit`, you can set `steps_per_execution` argument provided at `Model.compile` to a value larger than 1.
- For a custom training loop, you can pack multiple steps into a single `tf.function`:
```python
steps_per_invocation = 10
@tf.function
def step_fn(iterator):
for _ in range(steps_per_invocation):
features, labels = next(iterator)
def replica_fn(features, labels):
...
strategy.run(replica_fn, args=(features, labels))
```
As the library is optimized further, hopefully most users won't have to manually pack steps in the future.
In addition, a small trick for performance improvement is to schedule functions without a return value as explained in the handling task failure section above.
## Known limitations
<a id="known_limitations"> </a>
Most of the known limitations are already covered in the above sections. This section provides a summary.
### `ParameterServerStrategy` general
- `os.environment["grpc_fail_fast"]="use_caller"` is needed on every task including the coordinator, to make fault tolerance work properly.
- Synchronous parameter server training is not supported.
- It is usually necessary to pack multiple steps into a single function to achieve optimal performance.
- It is not supported to load a saved_model via `tf.saved_model.load` containing sharded variables. Note loading such a saved_model using TensorFlow Serving is expected to work.
- It is not supported to load a checkpoint containing sharded optimizer slot variables into a different number of shards.
- It is not supported to recover from parameter server failure without restarting the coordinator task.
- Usage of `tf.lookup.StaticHashTable` (which is commonly employed by some Keras preprocessing layers, such as `tf.keras.layers.IntegerLookup`, `tf.keras.layers.StringLookup`, and `tf.keras.layers.TextVectorization`) results in resources placed on the coordinator at this time with parameter server training. This has performance implications for lookup RPCs from workers to the coordinator. This is a current high priority to address.
### `Model.fit` specifics
- `steps_per_epoch` argument is required in `Model.fit`. You can select a value that provides appropriate intervals in an epoch.
- `ParameterServerStrategy` does not have support for custom callbacks that have batch-level calls for performance reasons. You should convert those calls into epoch-level calls with suitably picked `steps_per_epoch`, so that they are called every `steps_per_epoch` number of steps. Built-in callbacks are not affected: their batch-level calls have been modified to be performant. Supporting batch-level calls for `ParameterServerStrategy` is being planned.
- For the same reason, unlike other strategies, progress bar and metrics are logged only at epoch boundaries.
- `run_eagerly` is not supported.
### Custom training loop specifics
- `ClusterCoordinator.schedule` doesn't support visitation guarantees for a dataset.
- When `ClusterCoordinator.create_per_worker_dataset` is used, the whole dataset must be created inside the function passed to it.
- `tf.data.Options` is ignored in a dataset created by `ClusterCoordinator.create_per_worker_dataset`.
| github_jupyter |
<a href="https://colab.research.google.com/github/ibrahimsesay/griddb/blob/master/week1_assignment1_Sorori_Shinzaemon_problem.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# PROBLEM. 1 The Number of Grain of Rice in the 100th day
```
num_days = 100
total_grain_rice = 1
for i in range(1, num_days + 1):
print(i, total_grain_rice)#show the amount of grains received for that day
total_grain_rice *= 2
#grain of rice in the 100th day
print("Sum of rice in", i, "days is: ", total_grain_rice)
num_days = 100
total_grain_rice = 1
#creating the values for the x and y axis
yaxis = []
xaxis = list(range(1, num_days + 1))
for i in range(1, num_days + 1,):
#list the change in the rice grain
print("For day: {} the total grain is: {}".format(i, total_grain_rice))
#appending the total grain rice into the list of the yaxis
yaxis.append(total_grain_rice)
total_grain_rice = 2**i
print("Sum of rice in", i, "days is: ", total_grain_rice)
#plotting the graph
import matplotlib.pyplot as plt
plt.xlabel('Number of Days')
plt.ylabel('Number of Rice')
plt.title('Change in the Number of Rice Grain')
plt.plot(xaxis, yaxis, linewidth=3.0)
plt.tick_params(labelsize=10)
plt.figure(figsize=(48, 24))
plt.show()
```
【Problem 2] Number of rice grains outside of the 100th day
```
#Create a function
def compute_sorori_shinzaemon(day=100):
#List containing number of days
list_n_grains = [(lambda x: x)(x) for x in range(1, day + 1)]
#This list show the total number of rice receive per day
list_total_grains = [2**x-1 for x in range(1, day + 1) ]
pass
return list_n_grains, list_total_grains
""""
The function to calculate the amount of rice a person can get
for the number of days he/she has taken the course.
It also take input from users.
"""
list_n_grains, list_total_grains = compute_sorori_shinzaemon(day=10)
#index function to call the total amount
print("days: {}".format(list_n_grains[-1]))
#index function to call the total amount
print("grains: {}".format(list_total_grains[-1]))
```
# version of the above code that accept input from the user
```
#Create a function
def compute_sorori_shinzaemon(day=100):
#List containing number of days
list_n_grains1 = [(lambda x: x)(x) for x in range(1, day + 1)]
#This list show the total number of rice receive per day
list_total_grains1 = [2**x-1 for x in range(1, day + 1) ]
pass
return list_n_grains1, list_total_grains1
""""
The function to calculate the amount of rice a person can get
for the number of days he/she has taken the course.
It also take input from users.
"""
list_n_grains1, list_total_grains1 = compute_sorori_shinzaemon(int(input("Days amount Entered: ")))
#index function to call the total amount
print("days: {}".format(list_n_grains1[-1]))
#index function to call the total amount
print("grains: {}".format(list_total_grains1[-1]))
```
# Plotting of the graph
The input data for the graph is going to be influence by the input the input the user is going to
give in the last task. this is due to the slight experimentations i was doing twith the code.
```
import matplotlib.pyplot as plt
plt.xlabel('Number of Days')
plt.ylabel('Number of Rice')
plt.title('Change in the Number of Rice Grain')
plt.plot(list_n_grains1, list_total_grains1 linewidth=3.0)
plt.tick_params(labelsize=10)
plt.figure(figsize=(48, 24))
plt.show()
```
# 【Problem 3] How many people can live for how many days with the rice we get?
According to a research done on Wikipedia, the united nation's world food program stated that
it takes 400 grams or about 19200 grains of rice to feed one adult for a day.
```
#Create a function
def compute_n_days(number_of_grain, number_of_people, ):
"""
A function to calculate the number of people can live for how many days with the rice we get.
Parameteres
--------------
number_of_grain: int
The number of grains of rice
number_of_people: int
number of people to live
Returns
--------------
days_to_live : int
shows the number of days a person will live
"""
days_to_live = total_grain_rice/(number_of_grain*number_of_people)
number_of_people = total_grain_rice/(number_of_grain*days_to_live)
return days_to_live
days_to_live = compute_n_days(number_of_grain=19200, number_of_people=60)
print("{} DAYS".format(days_to_live))
#Create a function
def compute_n_days(number_of_grain, number_of_people, ):
"""
A function to calculate the number of people can live for how many days with the rice we get.
Parameteres
--------------
number_of_grain: int
The number of grains of rice
number_of_people: int
number of people to live
Returns
--------------
days_to_live : int
shows the number of days a person will live
"""
#A version of problem 3 than respond to the input of user in problem 2
days_to_live = list_total_grains1[-1]/(number_of_grain*number_of_people)
number_of_people = list_total_grains1[-1]/(number_of_grain*days_to_live)
return days_to_live
days_to_live = compute_n_days(number_of_grain=19200, number_of_people=60)
print("{} DAYS".format(days_to_live))
```
| github_jupyter |
```
#importing Packages
import numpy as np
from PIL import Image
import os, glob
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras.preprocessing.image import ImageDataGenerator
import matplotlib.pyplot as plt
import zipfile
class FingerModel():
def __init__(self):
pass
#initializing the image generator of test and train
def imageGenerator(self, train_path, test_path):
#you can tweak all the parameters i'm giving the best parameters list which i found"
batch = 128 #Batch size of image generaor
train_datagenerator = ImageDataGenerator( rescale=1./255,
rotation_range=10.,
width_shift_range=0.1,
height_shift_range=0.1,
zoom_range=0.2,
horizontal_flip=True
)
test_datagenerator = ImageDataGenerator( rescale=1./255 )
training = train_datagenerator.flow_from_directory(
train_path,
target_size=(150, 150),
color_mode='grayscale',
batch_size=batch,
classes=['0','1','2','3','4','5'],
class_mode='categorical'
)
testing = test_datagenerator.flow_from_directory(
test_path,
target_size=(150, 150),
color_mode='grayscale',
batch_size=batch,
classes=['0','1','2','3','4','5'],
class_mode='categorical'
)
return training, testing
def modelArchitecture(self):
#you can tweak all the parameters i'm giving the best parameters list which i found"
"""
Here i'm using 2D Conv because we converted our dataset to gray scale images of size 150X150
"""
model = tf.keras.models.Sequential([
tf.keras.layers.Conv2D(32, (3,3), activation='relu', input_shape=(150, 150, 1)),
tf.keras.layers.MaxPooling2D(2, 2),
tf.keras.layers.Conv2D(64, (3,3), activation='relu'),
tf.keras.layers.MaxPooling2D(2, 2),
tf.keras.layers.Conv2D(128, (3,3), activation='relu'),
tf.keras.layers.MaxPooling2D(2, 2),
tf.keras.layers.Conv2D(128, (3,3), activation='relu'),
tf.keras.layers.MaxPooling2D(2, 2),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(512, activation='relu'),
tf.keras.layers.Dropout(0.3),
tf.keras.layers.Dense(6, activation='softmax')
])
print(model.summary())
model.compile(optimizer='adam', loss='categorical_crossentropy', metrics=['acc'])
return model
def trainModel(self, train_data, test_data, model):
#you can tweak all the parameters i'm giving the best parameters list which i found"
no_of_epochs = 10
no_of_steps_per_epoch = 140
no_of_validation_steps = 30
history = model.fit_generator(
train_data,
steps_per_epoch = no_of_steps_per_epoch,
epochs = no_of_epochs,
validation_data = test_data,
validation_steps = no_of_validation_steps)
model.save("my_model.h5")#saving the model for future use.
def main(self):
train_data_path = "fingers/train/"
test_data_path = "fingers/test/"
train_datagen, test_datagen = self.imageGenerator(train_data_path, test_data_path)
model = self.modelArchitecture()
self.trainModel(train_datagen, test_datagen, model)
return "success"
if __name__=="__main__":
obj = FingerModel()
obj.main()
```
| github_jupyter |
```
# Load dependencies
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
from scipy.stats import gmean
import sys
sys.path.insert(0, '../../../statistics_helper/')
from fraction_helper import *
from excel_utils import *
# Define function that runs a jypyter notebook and saves the results to the same file
def run_nb(path):
import nbformat
from nbconvert.preprocessors import ExecutePreprocessor
import os
with open(path) as f:
nb = nbformat.read(f, as_version=4)
ep = ExecutePreprocessor(timeout=6000, kernel_name='python3')
ep.preprocess(nb, {'metadata': {'path': os.path.dirname(path)}})
with open(path, 'wt') as f:
nbformat.write(nb, f)
```
# Estimating the biomass of marine arthropods
To estimate the biomass of marine arthropods, we rely on data from the marine ecosystem biomass data (MAREDAT) initiative. The MAREDAT database contains measurements of the biomass concentration for each plankton group. From this database [Buitenhuis et al.](https://doi.org/10.5194/essd-5-227-2013) generates estimates for the global biomass of each plankton group by using a characteristic biomass concentration for each depth (either a median or average of the values in the database) and applying it across the entire volume of ocean at that depth.
Buitenhuis et al. reports two estimates, one based on the arithmetic mean of concentrations at each depth, and one based on the median concentrations at each depth. The estimate based on the arithmetic mean is more susceptible to sampling bias, as even a single measurement which is not characteristic of the global population (such as samples which are in coastal waters, or samples which have some technical biases associated with them) might shift the average concentration significantly. On the other hand, the estimate based on the geometric mean might underestimate global biomass as it will reduce the effect of biologically relevant high biomass concentrations. As a compromise between these two caveats, we chose to use as our best estimate the geometric mean of the estimates from the two methodologies.
The data in the MAREDAT database is divided into plankton size classes: microzooplankton (zooplankton between 5 and 200 µm in diameter), mesozooplankton (zooplankton between 200 and 2000 µm in diameter) and macrozooplankton (zooplankton between 2 and 10 mm). We are interested in the biomass of arthropods in each class.
## Microzooplankton
Microzooplankton was defined in the MAREDAT databased as to exclude copepod biomass, and thus its contribution to the total biomass of marine arthropods is neglegible.
## Mesozooplankton
Mesozooplankton might contain several different kinds of animal and protist taxa. We argure that the main contribution to the mesozooplankton category in MAREDAT database comes from arthropods (mainly copepods). To substantiate this claim, we rely on 18S rDNA sequencing data from the *Tara* Oceans campaign reported in [de Vargas et al.](http://dx.doi.org/10.1126/science.1261605). In figure W10A in the companion [website](http://taraoceans.sb-roscoff.fr/EukDiv/#figW10) to the paper the authors detail abundance of 18S rDNA reads for different animal and protist taxa:
<img src= "http://taraoceans.sb-roscoff.fr/EukDiv/images/FigW-10-1.png">
### Can 18S rDNA data be used to estimate biomass contribution?
It is not obvious that data on the relative abundance of 18S sequences can be used in order to estimate the relative biomass contribution of different taxa. We provide two independent lines of evidence of the legitimacy of using the 18S data in de Vargas et al. in order to claim that arthropod dominate the biomass of mesozooplanktonin the MAREDAT data.
The first line of evidence is in figure W3 in the companion [website](http://taraoceans.sb-roscoff.fr/EukDiv/#figureW3): <img src="http://taraoceans.sb-roscoff.fr/EukDiv/images/FigW-03.png">
This figure shows good correlations between the rDNA content in a cell and the cell size, as well as correlation between microscopy data and 18S rDNA sequencing data.
The second line of evidence is a comparison of the relative fraction of Rhizaria in the 18S rDNA data with
data from in-situ imaging ([Biard et al.](http://dx.doi.org/10.1038/nature17652)). We estimate the average relative fraction of Rhizaria in it the 18S rDNA sequencing data:
```
# Load 18S sequecing data
seq_data = pd.read_excel('marine_arthropods_data.xlsx',sheet_name='de Vargas',skiprows=1)
print('The average fraction of Rhizaria in 18S rDNA sequencing data in surface waters is ' + '{:,.0f}%'.format(seq_data['Rhizaria surface'].mean()*100))
print('The average fraction of Rhizaria in 18S rDNA sequencing data in the deep chlorophyll maximum is ' + '{:,.0f}%'.format(seq_data['Rhizaria DCM'].mean()*100))
```
These fraction of 35-40% are very close to the average fraction reported by Biard et al. using imaging data of ≈33%.
### The biomass contribution of arthropod to mesozooplankton data in MAREDAT
To calculate the contribution of arthropods to the biomass data in the MAREDAT database, we assume the representation of Rhizaria in the MADERAT data is limited, as Biard et al. indicated that they are usually undersampled because many of them are delicate and are severely damaged by plankton nets or fixatives used in surveys such as the ones used to build the MAREDAT. Therefore, we calculate the relative contribution of arthropods to the total population of mesozooplankton excluding Rhizaria:
```
# Define the relative fraction of arthropods out of the total mesozooplankton excluding Rhizaria
arth_frac_surf = seq_data['Arthropod surface']/(1-seq_data['Rhizaria surface'])
arth_frac_dcm = seq_data['Arthropod DCM']/(1-seq_data['Rhizaria DCM'])
# Calculate the mean fraction of arthropods in surface waters and the DCM
mean_arth_frac_surf = frac_mean(arth_frac_surf)
mean_arth_frac_dcm = frac_mean(arth_frac_dcm)
print('The average fraction of arthropods out of the total biomass of mesozooplankton in surface waters excluding Rhizaria is '+'{:,.0f}%'.format(mean_arth_frac_surf*100))
print('The average fraction of arthropods out of the total biomass of mesozooplankton in the deep chlorophyll maximum excluding Rhizaria is '+'{:,.0f}%'.format(mean_arth_frac_dcm*100))
```
Overall, we use ≈80% for the fraction of arthropods out of the total biomass of mesozooplankton in the MAREDAT database.
To estimate the biomass of mesozooplankton arthropods, we rely on the global estimates made by [Buitenhuis et al.](http://search.proquest.com/openview/0e8e5672fa28111df473268e13f2f757/1?pq-origsite=gscholar&cbl=105729) based on the MAREDAT data. Buitenhuis et al. generated two estimates for the global biomass of mesozooplankton by using a characteristic biomass concentration for each depth (either a median or average of the values in the database) and applying it across the entire volume of ocean at that depth. This approach results in two types of estimates for the global biomass of mesozooplankton: a so called “minimum” estimate which uses the median concentration of biomass from the database, and a so called “maximum” estimate which uses the average biomass concentration. Because the distributions of values in the database are usually highly skewed by asymmetrically high values, the median and mean are loosely associated by the MAREDAT authors with a minimum and maximum estimate. The estimate based on the average value is more susceptible to biases in oversampling singular locations such as blooms of plankton species, or of coastal areas in which biomass concentrations are especially high, which might lead to an overestimate. On the other hand, the estimate based on the median biomass concentration might underestimate global biomass as it will reduce the effect of biologically relevant high biomass concentrations. Therefore, our best estimate of the biomass of mesozooplakton is the geometric mean of the “minimum” and “maximum” estimates. Buitenhuis et al. reports a "minimum" estimate of 0.33 Gt C and a "maximum" estimate of 0.59 Gt C. We calculate the geometric mean of those estimates:
```
# Calculate the geometric mean of the "minimum" and "maximum" estimates from Buitenhuis et al.
buitenhuis_estimate = gmean([0.33e15,0.59e15])
```
We than use 80% of the geometric mean as an estimate for the biomass of mesozooplankton arthropods:
```
# Calculate the mean fraction of arthropods between surface water and DCM
arth_frac = frac_mean(np.array([mean_arth_frac_dcm,mean_arth_frac_surf]))
# Calculate the fraction of mesozooplankton biomass that is arthropod biomass
meso_arth_biomass = buitenhuis_estimate*arth_frac
```
Most of the data in the MAREDAT databased was collected using 300 µm nets, and thus some of the lower size fraction of mesozooplankton was not collected. To correct for this fact, we use a relation between biomass estimated using 200 µm nets and 300 µm nets [O'brian 2005](https://www.st.nmfs.noaa.gov/copepod/2005/documents/fspo73_abbreviated.pdf). The relation is: $$ B_{300} = 0.619× B_{200}$$ Where $B_{300}$ is the biomass sampled with 300 µm nets and $B_{200}$ is the biomass sampled with 200 µm nets. We correct for this factor to get our best estimate for the biomass of mesozooplankton arthropods:
```
# Correct for the use of 300 µm nets when sampling mesozooplankton biomass
meso_arth_biomass /= 0.619
print('Our best estimate for the biomass of mesozooplankton arthropods is ≈%.2f Gt C' % (meso_arth_biomass/1e15))
```
## Macrozooplankton
Some arthropods are also included in the macrozooplankton size category (zooplankton between 2 and 10 mm). Macrozooplankton contains organisms from many phyla such as arthropods, cnidarians, chordates, annelids, molluscs, ctenophores and representatives from Chaetognatha (a phylum of pelagic worms). To estimate the biomass of macrozooplankton arthropods, we first estimate the total biomass of macrozooplankton, and then estimate the fraction fo this total biomass that is contributed by arthropods.
To estimate the total biomass of macrozooplankton, we rely on data from the MAREDAT database. We use the estimates of macrozooplankton biomass Buitenhuis et al. generated from the MAREDAT database. To generate these estimates, Buitenhuis et al. followed the same procedure as we detailed in the mesozooplankton section above. Buitenhuis et al. provides “minimum” and “maximum” estimates of the total biomass of macrozooplankton, which are 0.2 Gt C and 1.5 Gt C, respectively. We use the geometric mean of those estimates as our best estimate for the biomass of macrozooplankton:
```
macro_biomass = gmean([0.2e15,1.5e15])
print('Our best estimate for the biomass of macrozooplankton is ≈%.1f Gt C' %(macro_biomass/1e15))
```
From this total biomass we subtract our estimates for the biomass of pteropods, which are in the same size range as macrozooplankton. For details on the estimate of the biomass of pteropods see the molluscs section in the Supplementary Information. We estimate the total biomass of pteropods at 0.15 Gt C.
We also subtract from the total biomass of macrozooplankton the contribution by gelatinous zooplankton which also contains some species in the same size range as macrozooplankton. We estimate a global biomass of ≈0.04 Gt C (for details on the estimate of the biomass of gelatinous plankton see the cnidarians section in the Supplementary Information).
```
# Calculate the total biomass of macrozooplankton arthropods by
# subtacting the biomass of pteropods and gelatinous zooplankton
# from the total biomass of macrozooplankton
# Load biomass estimates for pteropods and gelatinous zooplankton
other_macrozooplankton = pd.read_excel('marine_arthropods_data.xlsx','Other macrozooplankton')
# In other zooplankton biomass estimate is empty, run the scripts
if(other_macrozooplankton.shape[0]<2):
run_nb('../../cnidarians/cnidarians.ipynb')
run_nb('../../molluscs/molluscs.ipynb')
other_macrozooplankton = pd.read_excel('marine_arthropods_data.xlsx','Other macrozooplankton')
macro_arth_biomass = macro_biomass - other_macrozooplankton['Value'].sum()
print('our best estimate for the total biomass of macrozooplankton arthropods is ≈%.1f Gt C' %(macro_arth_biomass/1e15))
```
We sum up the biomass of arthropods in the mesezooplankton and macrozooplankton size fractions as our best estimate for the biomass of marine arthropods:
```
best_estimate = meso_arth_biomass+macro_arth_biomass
print('Our best estimate for the biomass of marine arthropods is %.1f Gt C' %(best_estimate/1e15))
```
# Uncertanity analysis
We discuss the uncertainty of estimates based on the MAREDAT database in a dedicated section in the Supplementary Information. We crudly project an uncertainty of about an order of magnitude.
```
# We project an uncertainty of an order of magnitude (see MAREDAT consistency check section)
mul_CI = 10
```
# Estimating the total number of marine arthropods
We consider only the mesozooplankton as they are the smallest group of marine arthropods (by the definitions of the MAREDAT database they also contain microzooplankton). To estimate the total number of marine arthropods, we divide our estimate for the total biomass of mesozooplankton by an estimate for the characteristic carbon content of an individual copepod, which dominate the mesozooplankton biomass. As the basis of our estimate for the charactristic carbon content of a single copepod, we rely on data from [Viñas et al.](http://dx.doi.org/10.1590/S1679-87592010000200008) and [Dai et al.](https://doi.org/10.1016/j.jmarsys.2015.11.004), which range from 0.15 µg C to 100 µg C per individual. We use the geometric mean of this range, which is ≈4 µg C per individual, as our best estimate of the carbon content of a single copepod.
```
# The carbon content of copepods
copepod_carbon_content = 4e-6
# Calculate the total number of marine arthropods
marine_arth_num = meso_arth_biomass/copepod_carbon_content
print('Our best estimate for the total number of marine arthropods is ≈%.0e' % marine_arth_num)
# Feed results to the animal biomass data
old_results = pd.read_excel('../../animal_biomass_estimate.xlsx',index_col=0)
result = old_results.copy()
result.loc['Marine arthropods',(['Biomass [Gt C]','Uncertainty'])] = (best_estimate/1e15,mul_CI)
result.to_excel('../../animal_biomass_estimate.xlsx')
# Feed results to Table 1 & Fig. 1
update_results(sheet='Table1 & Fig1',
row=('Animals','Marine arthropods'),
col=['Biomass [Gt C]', 'Uncertainty'],
values=[best_estimate/1e15,mul_CI],
path='../../../results.xlsx')
# Feed results to Table S1
update_results(sheet='Table S1',
row=('Animals','Marine arthropods'),
col=['Number of individuals'],
values=marine_arth_num,
path='../../../results.xlsx')
```
| github_jupyter |
```
# ------------------------- #
# SET - UP #
# ------------------------- #
# ---- Requirements ----- #
!pip install datasets
!pip install sentencepiece
!pip install transformers
!pip install rouge_score
!pip install bert_score
from bert_score import score
from datasets import load_dataset, Dataset, load_metric
import sys
from google.colab import drive
import pandas as pd
import numpy as np
from transformers import BartForConditionalGeneration, BartTokenizer, DataCollatorForSeq2Seq, Seq2SeqTrainingArguments, Seq2SeqTrainer
import torch
import huggingface_hub
import matplotlib.pyplot as plt
import nltk
nltk.download("punkt")
import gc
from torch import nn
# ----- Check if GPU is connected ----- #
gpu_info = !nvidia-smi -L
gpu_info = "\n".join(gpu_info)
if gpu_info.find("failed") >= 0:
print("Not connected to a GPU")
else:
print(gpu_info)
# ----- Mounting Google Drive ----- #
drive.mount('/content/drive')
sys.path.append('/content/drive/MyDrive/CIS6930_final')
# ----- Importing TweetSum processing module ----- #
from tweet_sum_processor import TweetSumProcessor
# ----- Torch Device ----- #
torch_device = 'cuda' if torch.cuda.is_available() else 'cpu'
# ----------------------------------------------------------------------
# --- DEFINE MODEL AND TOKENIZER --- #
model_name = "facebook/bart-large"
model = BartForConditionalGeneration.from_pretrained(model_name)
tokenizer = BartTokenizer.from_pretrained(model_name)
# ----- Metric
metric = load_metric("rouge")
# ---- Freeze parameters
def freeze_params(model: nn.Module):
"""Set requires_grad=False for each of model.parameters()"""
for par in model.parameters():
par.requires_grad = False
def freeze_embeds(model):
"""Freeze token embeddings and positional embeddings for BART and PEGASUS, just token embeddings for t5."""
model_type = model.config.model_type
if model_type == "t5":
freeze_params(model.shared)
for d in [model.encoder, model.decoder]:
freeze_params(d.embed_tokens)
else:
freeze_params(model.model.shared)
for d in [model.model.encoder, model.model.decoder]:
freeze_params(d.embed_positions)
freeze_params(d.embed_tokens)
freeze_embeds(model)
# ----- Reading in the Dataset
raw_datasets = load_dataset('csv', data_files={'train': '/content/drive/MyDrive/CIS6930_final/tweetsum_train.csv',
'valid': '/content/drive/MyDrive/CIS6930_final/tweetsum_valid.csv',
'test': '/content/drive/MyDrive/CIS6930_final/tweetsum_test.csv'})
max_input_length = 512
max_target_length = 128
def preprocess_function(examples):
model_inputs = tokenizer(examples["inputs"], max_length=max_input_length, truncation=True)
# Setup the tokenizer for targets
with tokenizer.as_target_tokenizer():
labels = tokenizer(examples["summaries"], max_length=max_target_length, truncation=True)
model_inputs["labels"] = labels["input_ids"]
return model_inputs
tokenized_datasets = raw_datasets.map(preprocess_function, batched=True)
batch_size = 1
args = Seq2SeqTrainingArguments(
evaluation_strategy = "epoch",
learning_rate=2e-5,
per_device_train_batch_size=batch_size,
per_device_eval_batch_size=batch_size,
weight_decay=0.01,
save_total_limit=3,
num_train_epochs=3,
predict_with_generate=True,
fp16=True,
push_to_hub=False,
output_dir = '/content/drive/MyDrive/CIS6930_final/results/bart',
logging_dir = '/content/drive/MyDrive/CIS6930_final/logs/bart'
)
def compute_metrics(eval_pred):
predictions, labels = eval_pred
decoded_preds = tokenizer.batch_decode(predictions, skip_special_tokens=True)
# Replace -100 in the labels as we can't decode them.
labels = np.where(labels != -100, labels, tokenizer.pad_token_id)
decoded_labels = tokenizer.batch_decode(labels, skip_special_tokens=True)
# Rouge expects a newline after each sentence
decoded_preds = ["\n".join(nltk.sent_tokenize(pred.strip())) for pred in decoded_preds]
decoded_labels = ["\n".join(nltk.sent_tokenize(label.strip())) for label in decoded_labels]
result = metric.compute(predictions=decoded_preds, references=decoded_labels, use_stemmer=True)
# Extract a few results
result = {key: value.mid.fmeasure * 100 for key, value in result.items()}
# Add mean generated length
prediction_lens = [np.count_nonzero(pred != tokenizer.pad_token_id) for pred in predictions]
result["gen_len"] = np.mean(prediction_lens)
return {k: round(v, 4) for k, v in result.items()}
data_collator = DataCollatorForSeq2Seq(tokenizer, model=model)
trainer = Seq2SeqTrainer(
model,
args,
train_dataset=tokenized_datasets["train"],
eval_dataset=tokenized_datasets["valid"],
data_collator=data_collator,
tokenizer=tokenizer,
compute_metrics=compute_metrics
)
trainer.train()
```
# TESTING
```
# --------------------- #
# TEST EVALUATION #
# --------------------- #
out = trainer.predict(tokenized_datasets["test"])
generated_summaries = []
for i in range(0, 110):
generated_summaries.append(tokenizer.decode(out[0][i], skip_special_tokens = True))
ground_truth = tokenized_datasets["test"]["summaries"]
conversation = tokenized_datasets["test"]["inputs"]
```
## ROUGE SCORES
```
out.metrics
```
## BART-SCORE
```
P, R, F1 = score(generated_summaries, ground_truth, lang="en", verbose=True)
```
### SCORES
```
print(f"System level F1 score: {F1.mean():.3f}")
print(f"System level precision score: {P.mean():.3f}")
print(f"System level recall score: {R.mean():.3f}")
```
### VISUALIZATIONS
```
plt.hist(F1, bins=20)
plt.title("Distribution of F1 Scores")
plt.show()
plt.hist(P, bins=20)
plt.title("Distribution of Precision Scores")
plt.show()
plt.hist(R, bins=20)
plt.title("Distribution of Recall Scores")
plt.show()
```
## SAVE SUMMARIES TO CSV
```
# ----- Mounting Google Drive ----- #
drive.mount('/content/drive')
sys.path.append('/content/drive/MyDrive/CIS6930_final')
bart_summaries = pd.DataFrame({"candidate": generated_summaries, "reference": ground_truth, "conversation": conversation})
bart_summaries.to_csv('/content/drive/MyDrive/CIS6930_final/summaries/bart_test_summaries2.csv')
print("Done")
```
| github_jupyter |
# NYC Taxi Rides
In this notebook we show some more advanced features that are useful for building composed views, a.k.a., dashboards.
```
HOST = 'localhost'
PORT = 8003
# If using Binder, set this to the current host: https://hub.{domain}/user-{username}-{hash}
LOCATION = ''
if LOCATION:
# Use the jupyterhub proxy address to reach our server
API_URL = LOCATION.replace('/lab', '') + f'/proxy/{PORT}/api/v1'
else:
API_URL = f'http://{HOST}:{PORT}/api/v1'
%load_ext autoreload
%autoreload 2
```
### Download the data
```
import pandas as pd
from utils import download_file
nyc_data = pd.read_csv('data/nyc-taxi.csv')
for data in nyc_data.iterrows():
download_file(*data[1])
```
### Create tilesets
```
from higlass.tilesets import cooler
pickup = cooler('data/nyctaxi-pickup.count.1m.mcool', name='Total Pickups')
dropoff = cooler('data/nyctaxi-dropoff.count.1m.mcool', name='Total Dropoffs')
```
### Visualize pickups and drop-offs
```
from higlass import Track, View, display
nyc_x=[11789206, 11827005]
nyc_y=[15044734, 15081957]
view_config = {
'width': 6,
'initialXDomain': nyc_x,
'initialYDomain': nyc_y,
}
pickups_view = View(
x=0,
tracks=[Track('heatmap', tileset=pickup, height=400, server=API_URL)],
**view_config
)
dropoffs_view = View(
x=6,
tracks=[Track('heatmap', tileset=dropoff, height=400, server=API_URL)],
**view_config
)
widget, server, viewconf = display([pickups_view, dropoffs_view],
host=HOST, server_port=PORT)
widget
```
### Synchronize the viewport location and zoom level
```
widget, server, viewconf = display(
views=[pickups_view, dropoffs_view],
location_syncs=[(pickups_view, dropoffs_view)],
zoom_syncs=[(pickups_view, dropoffs_view)],
host=HOST,
server_port=PORT
)
widget
```
### Add crosshairs and tooltips
```
## Important change ##########
track_config = {
'track_type': 'heatmap',
'height': 400,
'options': {
'showMousePosition': True,
'mousePositionColor': 'black',
'showTooltip': True,
},
'server': API_URL,
}
##############################
pickups_view = View(
x=0,
tracks=[Track(tileset=pickup, **track_config)],
**view_config
)
dropoffs_view = View(
x=6,
tracks=[Track(tileset=dropoff, **track_config)],
**view_config
)
widget, server, viewconf = display(
views=[pickups_view, dropoffs_view],
location_syncs=[(pickups_view, dropoffs_view)],
zoom_syncs=[(pickups_view, dropoffs_view)],
host=HOST,
server_port=PORT
)
widget
```
### Overview & Details with Viewport projections
```
from copy import deepcopy
## Important change ##########
view_config_details = deepcopy(view_config)
view_config_details['y'] = 6
view_config_details['initialXDomain'] = [11811324, 11817197]
view_config_details['initialYDomain'] = [15057338, 15060882]
##############################
new_track_config = deepcopy(track_config)
new_track_config['height'] = 300
pickups_view_details = View(
x=0,
tracks=[Track(tileset=pickup, **new_track_config)],
**view_config_details
)
dropoffs_view_details = View(
x=6,
tracks=[Track(tileset=dropoff, **new_track_config)],
**view_config_details
)
pickups_view_overview = View(
x=0,
tracks=[
Track(tileset=pickup, **new_track_config),
## Important change ##########
Track(
'viewport-projection-center',
fromViewUid=pickups_view_details.uid,
server=API_URL
)
##############################
],
**view_config
)
dropoffs_view_overview = View(
x=6,
tracks=[
Track(tileset=dropoff, **new_track_config),
## Important change ##########
Track(
'viewport-projection-center',
fromViewUid=dropoffs_view_details.uid,
server=API_URL
)
##############################
],
**view_config
)
widget, server, viewconf = display(
views=[
pickups_view_overview, dropoffs_view_overview,
pickups_view_details, dropoffs_view_details
],
location_syncs=[
(pickups_view_overview, dropoffs_view_overview),
(pickups_view_details, dropoffs_view_details)
],
zoom_syncs=[
(pickups_view_overview, dropoffs_view_overview),
(pickups_view_details, dropoffs_view_details)
],
host=HOST,
server_port=PORT
)
widget
```
### Compare different attributes through data operations
```
pickups_track = Track(tileset=pickup, **track_config)
dropoffs_track = Track(tileset=dropoff, **track_config)
pickups_view = View(
x=0, tracks=[pickups_track], **view_config
)
dropoffs_view = View(
x=6, tracks=[dropoffs_track], **view_config
)
widget, server, viewconf = display(
views=[pickups_view, dropoffs_view],
location_syncs=[(pickups_view, dropoffs_view)],
zoom_syncs=[(pickups_view, dropoffs_view)],
host=HOST,
server_port=PORT
)
widget
```
Let's divide pickups by dropoff by directly adjusting the view config
```
from higlass import ViewConf
from higlass.viewer import HiGlassDisplay
diverging_colormap = [
'#0085cc',
'#62cef6',
'#eeeeee',
'#ff66b8',
'#bf0066',
]
diff_track = (
pickups_track
.change_options(
name='Pickups divided by dropoffs',
colorRange=diverging_colormap,
valueScaleMin=0.02,
valueScaleMax=50)
.change_attributes(
server=server.api_address,
## Important change ##########
data={
'type': 'divided',
'children': [{
'server': API_URL,
'tilesetUid': pickup.uuid
}, {
'server': API_URL,
'tilesetUid': dropoff.uuid
}]
}
##############################
)
)
diff_view_config = deepcopy(view_config)
diff_view_config['width'] = 12
diff_view = View(tracks=[diff_track], **diff_view_config)
# Finally, we need to create a new view config and pass that into the widget
HiGlassDisplay(viewconf=ViewConf([diff_view]).to_dict())
```
- **Pink** indicates areas with more *pickups*
- **Blue** indicates areas with more *dropoffs*
### Superimpose tracks
```
## Important change ##########
diverging_colormap = [
'rgba(0, 133, 204, 0.66)',
'rgba(98, 206, 246, 0.33)',
'rgba(238, 238, 238, 0.01)',
'rgba(255, 102, 184, 0.33)',
'rgba(191, 0, 102, 0.66)',
]
new_diff_track = diff_track.change_options(
# This is critical
backgroundColor='transparent',
colorRange=diverging_colormap
)
##############################
mapbox_track_config = {
'track_type': 'mapbox',
'position': 'center',
'height': 400,
'options': {
'accessToken': 'pk.eyJ1IjoiZmxla3NjaGFzIiwiYSI6ImNqeHRzMWJvZTB4dmQzZ3Q1cThqejB2dGsifQ.j_01IEuxiF8-JhX1BGKueA',
'style': 'light-v9',
'minPos': [1],
'maxPos': [40075016],
'name': '© Mapbox',
'labelPosition': 'bottomLeft',
}
}
new_diff_view = View(
tracks=[Track(**mapbox_track_config), new_diff_track],
**diff_view_config
)
HiGlassDisplay(viewconf=ViewConf([new_diff_view]).to_dict())
```
### Finally... let's turn the lights off
Everyone loves dark mode! We love it too 😍
```
heatmap_track_config = {
'track_type': 'heatmap',
'height': 400,
'options': {
'backgroundColor': 'transparent',
## Important change ##########
'colorRange': [
'rgba(0, 0, 0, 0.01)',
'rgba(34, 46, 84, 0.25)',
'rgba(68, 141, 178, 0.5)',
'rgba(104, 191, 48, 0.7)',
'rgba(253, 255, 84, 0.8)',
'rgba(253, 255, 255, 0.9)',
],
##############################
'colorbarBackgroundColor': 'black',
'colorbarBackgroundOpacity': 0.5,
'labelColor': '#ffffff',
'labelTextOpacity': 0.66,
'labelBackgroundColor': 'black',
'labelBackgroundOpacity': 0.5,
},
'server': API_URL,
}
mapbox_track_config = {
'track_type': 'mapbox',
'position': 'center',
'height': 400,
'options': {
'accessToken': 'pk.eyJ1IjoiZmxla3NjaGFzIiwiYSI6ImNqZXB2aWd4NDBmZTIzM3BjdGZudTFob2oifQ.Jnmp1xWJyS4_lRhzrZAFBQ',
'style': 'dark-v8',
'minPos': [1],
'maxPos': [40075016],
'name': '© Mapbox',
'labelPosition': 'bottomLeft',
'labelColor': '#ffffff',
'labelTextOpacity': 0.66,
'labelBackgroundColor': 'black',
'labelBackgroundOpacity': 0.5,
},
'server': API_URL,
}
pickups_in_dark = View(
tracks=[
Track(**mapbox_track_config),
Track(tileset=dropoff, **heatmap_track_config),
],
**diff_view_config
)
## Important change ##########
widget, server, viewconf = display([pickups_in_dark], dark_mode=True,
host=HOST, server_port=PORT)
##############################
widget
```
---
## Other cool NYC properties worth looking at and comparing
```
import pandas as pd
from utils import download_file
nyc_data = pd.read_csv('data/nyc-taxi-extended.csv')
for data in nyc_data.iterrows():
download_file(*data[1])
from higlass.tilesets import cooler
dropoff_more = {
'passenger_count': cooler('data/nyctaxi-dropoff.passenger_count.mean.1m.mcool', name='Dropoffs Mean Passanger Count'),
'tip_amount': cooler('data/nyctaxi-dropoff.tip_amount.mean.1m.mcool', name='Dropoffs Mean Tip'),
'total_amount': cooler('data/nyctaxi-dropoff.total_amount.mean.1m.mcool', name='Dropoffs Mean Total Amount'),
'trip_distance': cooler('data/nyctaxi-dropoff.trip_distance.mean.1m.mcool', name='Dropoffs Mean Trip Distance'),
}
pickup_more = {
'passenger_count': cooler('data/nyctaxi-pickup.passenger_count.mean.1m.mcool', name='Pickups Mean Passanger Count'),
'tip_amount': cooler('data/nyctaxi-pickup.tip_amount.mean.1m.mcool', name='Pickups Mean Tip'),
'total_amount': cooler('data/nyctaxi-pickup.total_amount.mean.1m.mcool', name='Pickups Mean Total Amount'),
'trip_distance': cooler('data/nyctaxi-pickup.trip_distance.mean.1m.mcool', name='Pickups Mean Trip Distance'),
}
## Important change ##########
track_config = {
'track_type': 'heatmap',
'height': 400,
'options': {
'showMousePosition': True,
'mousePositionColor': 'black',
'showTooltip': True,
'heatmapValueScale': 'linear',
'valueScaleMin': 0,
}
}
##############################
pickup_amount_track = Track(tileset=pickup_more['total_amount'], **track_config).change_options(valueScaleMax=100)
pickup_dist_track = Track(tileset=pickup_more['trip_distance'], **track_config).change_options(valueScaleMax=20)
dropoff_amount_track = Track(tileset=dropoff_more['total_amount'], **track_config).change_options(valueScaleMax=100)
dropoff_dist_track = Track(tileset=dropoff_more['trip_distance'], **track_config).change_options(valueScaleMax=20)
pickup_amount_view = View(x=0, y=0, tracks=[pickup_amount_track], **view_config)
pickup_dist_view = View(x=0, y=6, tracks=[pickup_dist_track], **view_config)
dropoff_amount_view = View(x=6, y=0, tracks=[dropoff_amount_track], **view_config)
dropoff_dist_view = View(x=6, y=6, tracks=[dropoff_dist_track], **view_config)
widget, server, viewconf = display(
views=[
pickup_amount_view, pickup_dist_view,
dropoff_amount_view, dropoff_dist_view
],
location_syncs=[(
pickup_amount_view, pickup_dist_view,
dropoff_amount_view, dropoff_dist_view
)],
zoom_syncs=[(
pickup_amount_view, pickup_dist_view,
dropoff_amount_view, dropoff_dist_view
)],
)
widget
from higlass import ViewConf
from higlass.viewer import HiGlassDisplay
## Important change ##########
pickup_div_track = (
pickup_amount_track
.change_options(
name='Pickups: Amount by Distance')
.change_attributes(
server=server.api_address,
# With this magic setup we tell HiGlass to divide
# total amount by the trip distance
data={
'type': 'divided',
'children': [{
'server': server.api_address,
'tilesetUid': pickup_more['total_amount'].uuid
}, {
'server': server.api_address,
'tilesetUid': pickup_more['trip_distance'].uuid
}]
}
)
)
##############################
dropoff_div_track = (
dropoff_amount_track
.change_options(
name='Dropoffs: Amount by Distance')
.change_attributes(
server=server.api_address,
# With this magic setup we tell HiGlass to divide
# total amount by the trip distance
data={
'type': 'divided',
'children': [{
'server': server.api_address,
'tilesetUid': dropoff_more['total_amount'].uuid
}, {
'server': server.api_address,
'tilesetUid': dropoff_more['trip_distance'].uuid
}]
}
)
)
pickup_amount_by_dist_view = View(
x=0, y=0, tracks=[pickup_div_track], **view_config
)
dropoff_amount_by_dist_view = View(
x=6, y=0, tracks=[dropoff_div_track], **view_config
)
## Important change ##########
div_viewconf = ViewConf([
pickup_amount_by_dist_view,
dropoff_amount_by_dist_view
],
location_syncs=[(
pickup_amount_by_dist_view,
dropoff_amount_by_dist_view
)],
zoom_syncs=[(
pickup_amount_by_dist_view,
dropoff_amount_by_dist_view
)])
HiGlassDisplay(viewconf=div_viewconf.to_dict())
##############################
```
| github_jupyter |
### Common string literals and operations
```
S = '' # Empty string
print(type(S),len(S))
S1 = "spam's" # Double quotes, same as single
S2 = 'spam\'s'
print(S1 == S2, S1 is S2)
S1 = """...""" # Triple-quoted block strings
S2 = '...'
S3 = "..."
StrGroup = [S1,S2,S3]
for i in StrGroup:
for j in StrGroup:
if j is not i:
print(i==j)
S1 = r'\temp\spam' # Raw strings
S2 = '\temp\spam'
print(S1,S2)
S1 = b'spam' # Byte strings in 3.0
print(S1,type(S1))
bytes(3) + S1
S1 = u'spam' # Unicode strings in 2.6 only
print(S1,type(S1))
S2 = 'spam'
S3 = 'spam'
print(S2,S1==S2,S1 is S2, S2 is S3)
S1 + S1 + S1 + S1 == S1*4 # Concatenate, repeat
for i in range(4):
print("S[%d] ==> %s" %(i,S1[i]))
for i in S1[0:4]:
for j in range(4):
print("S[%d] ==> %s" %(j,i))
len(S1)
parrotname=input("please type a parrot's name: ")
kind = 'good' if parrotname == 'xiaoyanzhi' else 'bad'
"a %s parrot" % kind # String formatting expression
parrotname=input("please type a parrot's name: ")
kind = 'good' if parrotname == 'xiaoyanzhi' else 'bad'
"a {0} parrot".format(kind)
print(S1)
print(S1.find('pa'),S1.find('spam'),S1.find('spa')) # Return the lowest index in S where substring sub is found, such that sub is contained within S[start:end]. Optional arguments start and end are interpreted as in slice notation.
S1.find?
S3 = 's p a m ' # remove trailing whitespace
print(S3,'\n',len(S3))
print(S3.rstrip(),'\n', len(S3.rstrip()))
S3.rstrip?
S3.replace('p a','x x') # replacement
S3.split(','),S3.split() # split on delimiter
S3.split?
print(S3,S3.isdigit())
S3.isalnum(),S3.isalpha(),S3.isascii(),S3.isdecimal(),S3.isidentifier(),S3.islower(),S3.isnumeric()
S3.isprintable(),S3.isspace(),S3.istitle(),S3.isupper()
Selection = """S3.isalnum(),S3.isalpha(),S3.isascii(),S3.isdecimal(),S3.isidentifier(),S3.islower(),S3.isnumeric(),S3.isprintable(),S3.isspace(),S3.istitle(),S3.isupper()"""
print(S3)
SelGrp = Selection.split(',')
print(SelGrp)
for i in SelGrp:
print(eval(i))
if eval(i) == True:
print("S3 is %s" % i[5:].rstrip('()'))
bool(S3.isalnum())
print(S3.lower(),S3.upper()) # case conversion
S4 = """Riyuetan-Weipai Building is the world's largest solar energy office building. It opened on November 27 in Dezhou, Shandong Province in east China. The building, which has a total area of 75,000 square meters, features MFA Degree exhibition centers, scientific research facilities, meeting and training facilities, and a hotel"""
print(S3.endswith(' '),S4.endswith('hotel')) # end test
print(S3,S4)
S4.find(' a '),S4.count(' a ')
S4.find('a'),S4.count('a') # String method calls: search
'spam'.join(S4),S3.join(S4.upper()) # delimiter join
S3.encode('latin-1'), print(S1), print(S2)
S1 = b'spam' # Unicode encoding, etc.
S2 = 'spam'
print(S1==S2,S1 is S2)
print(S1==S2.encode('latin-1'),S1 is S2.encode('latin-1'))
for i in S3: print(i)
's p a m ' in S3 # test whether sub string is in S3
[c*4 for c in S3]
map(ord,S3)
map?
'spa"m',"spa'm",'''...spam...''',"""...spam..."""
'\newline'
```
### String Literals
```
print('spa"m') # Single quotes
print("spa'm") # Double quotes
print('''...spam...''',"""...spam...""") # Triple quotes
print("s\tp\na\0m") # Escape sequences
print(r"C:\new\test.spm") # Raw strings
print(b'sp\x01am') # Byte strings in 3.0
print(u'eggs\u0020spam') # Unicode strings in 2.6 only
```
### Single-and Double-Quoted Strings Are the Same
```
S= 'shrubbery'
D= "shrubbery"
print(S,D,S==D,S is D)
S = 'a\nb\tc'
print(S)
len(S)
S[0],S[1],S[2],S[3],S[4]
```
This string is five bytes long: it contains an ASCII a byte, a newline byte, an ASCII b byte, and so on.
```
print('\newline') # Ignored (continuation line)
print('\\') # Backslash (stores one \)
print('\'') # Single quote (stores ‘)
print('\"') # Double quote (stores “)
print('\a') # Bell
print('\b') # Backspace
print('\f') # Formfeed
print('\n') # Newline(linefeed)
print('\r') # Carriage return
print('\t') # Horizontal tab
print('\v') # Vertical tab
print('\x77') # Character with hex value hh (at most 2 digits)
print('\ooo') # Character with octal value ooo (up to 3 digits)
print('\0') # Null: binary 0 charater (doesn't end string)
# print('\N{ 1423 }') # Unicode database ID
print('\u0101') # Unicode 16-bit hex
# print('\U12340231') # Unicode 32-bit hex
print('\other') # Not an escape (keeps both \ and other)
path = r'C:\new\text.dat'
path
print(path)
import sys
print(sys.argv)
print(ord('s'))
print(chr(115))
S = 'a\0b\0c'
S
print(len(S))
print(S)
mantra = """So far, you’ve seen single quotes, double quotes, escapes, and raw strings in action.
Python also has a triple-quoted string literal format, sometimes called a block string,
that is a syntactic convenience for coding multiline text data. This form begins with
three quotes (of either the single or double variety), is followed by any number of lines
of text, and is closed with the same triple-quote sequence that opened it. Single and
double quotes embedded in the string’s text may be, but do not have to be, escaped—
the string does not end until Python sees three unescaped quotes of the same kind used
to start the literal."""
print(mantra)
```
### Basic Operations
```
print(len('abc'))
print('abc' + 'def')
print('Ni!'*4)
print('--------------------------------------------------------------------------------')
print('-'*80)
myjob = "hacker"
for c in myjob: print(c,end=' ')
print("k" in myjob)
print("z" in myjob)
print('spam' in 'abcspamdef')
S = 'spam'
print(S[0], S[-2])
print(S[1:3],S[1:],S[:-1])
```
### Extended slicing: the third limit and slice objects
```
S = 'abcdefghijklmnop'
print(S)
print(S[1:10:2]) # Extract all the items in X, from offset I through J-1, by K
print(S[1::2])
print(S[::])
print(S[::2])
S = 'abcedfg'
S[5:1:-1]
print('spam'[1:3]) # Slicing syntax
print('spam'[slice(1,3)])
print('spam'[::-1])
print('spam'[slice(None,None,-1)])
import sys
print(sys.argv)
%run echo.py -a -b -c
sys.argv[1:]
```
### String Conversion Tools
```
int("42"), str(42) # Convert from / to string
repr(42) # Convert to as-code string
print(str('spam'),repr('spam'))
S = "42"
I = 1
S + I
int(S) + I # Force addition
S + str(I) # Force concatenation
```
### Charater code Conversions
```
ord?
print(ord('s'))
print(chr(115))
S = '9'
print(ord(S) + 1)
print(chr(ord(S)+1))
print(chr(9+1))
S = chr(ord(S)+1)
B = '1101'
I = 0
while B != '':
I = I * 2 + (ord(B[0])-ord('0'))
B = B[1:]
I, 1*1 + 0 + 1*2*2 + 2*2*2*1
B = '1101' # Convert binary digits to integer with ord
B[0],B[1],B[2],B[3]
ord(B[0]),ord('0'),ord(B[0])-ord('0') # Round 1
I = 0*2 + (ord(B[0])-ord('0'))
B = B[1:]
print(I,B)
print(ord(B[0]),ord(B[0])-ord('0')) # Round 2
I = 1*2 + (ord(B[0])-ord('0'))
B = B[1:]
print(I,B)
print(ord(B[0]),ord(B[0])-ord('0'))
((((0*2 + 1-0)* 2 + 1-0)* 2 + 0-0)* 2 + 1-0) # Horner scheme
I = 0
print(I << 1)
int('1101',2) # Convert binary to integer: built-in
bin(13) # Convert integer to binary
```
### Changing Strings
```
S = 'spam'
S[0] = "x"
```
#### The immutable part means that you can't change a string in-place
```
S = S + 'SPAM!' # To change a string, make a new one
S
S = S[:4] + 'Burger' + S[-1]
S
S = 'splot'
S = S.replace('pl','pamal')
S
'That is %d %s bird!' %(1, 'dead')
'That is {0} {1} bird!'.format(1,'dead')
```
##### In finer-grained detail, functions are packages of code, and method calls combine two operations at once (an atrribute fetch and a call):
###### Attribute fetches: An expression of the form object.attribute means "fetch the value of attribute in object."
###### Call expressions: An expression of the form function(arguments) means "invoke the code of function, passing zero or more commma-separated argument objects to it, and return function's result value".
```
S.capitalize()
S.ljust?
print(len(S))
S.ljust(9,'l')
S.ljust(1000,'l')
S=S.center(18,'l')
S.lower()
S.count("l",4,40)
print(S)
S.count?
S.lstrip('l')
S.rstrip('l')
S.encode?
S.encode("utf-8")
b'lllllspamalotlllll'.decode('ascii')
aa = "和平精英"
print(aa)
aa.encode("utf-8")
S.encode('gb2312')
S.maketrans?
intab = 'a b c d e f g h i j k l m n o p q r s t u v w x y z '
outtab = '1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26'
str = """as you’ll see throughout this part of the book, most objects have callable methods, and all are accessed using this same method-call syntax. To call an object method, as you’ll see in the following sections, you have to go through an existing object."""
len(intab),len(outtab)
table = str.maketrans(intab,outtab)
print(str.translate(table))
# Program for conversion betweeen words and Morse code
'''
Author: will
Date : 2019/2/21
E-mail: willgyw@126.com
'''
# dict of words2morse
dict1 = {'a':'.-','b':'-...','c':'-.-.','d':'-..','e':'.','f':'..-.','g':'--.','h':'....','i':'..','j':'.---','k':'-.-','l':'.-..','m':'--','n':'-.','o': '---','p':'.--.','q':'--.-','r':'.-.','s':'...','t':'-','u':'..-','v':'...-','w':'.--','x':'-..-','y':'-.--','z':'--..','0':'-----','1':'.----','2':'..---','3':'...--','4':'....-','5':'.....','6':'-....','7':'--...','8':'---..','9':'----.'}
# dict of morse2words
dict2 = dict(zip(dict1.values(),dict1.keys()))
def encode():
words = input("Input a sentence you want to encode, NO Punctuation:").strip().lower()
for letter in words:
if letter == ' ':
print('/', end=' ')
else:
print(dict1[letter],end=' ')
def decode():
codes = input("Input Morse code you want to decode, only morse code:").strip().split(" ")
for sign in codes:
if sign == '/':
print(' ',end = '')
else:
print(dict2[sign], end = '')
print()
def main():
while 1 == 1:
choice = input("Encode(Words to Morse codes) or Decode(Morse codes to words).please input [0/1]")
if choice == '0':
encode()
elif choice == '1':
decode()
else:
break
if __name__ == "__main__":
main()
print(dict1)
word = input("text a sentence:")
dictionary = {'a': '1', 'b': '2', 'c': '3', 'd': '4', 'e': '5', 'f': '6', 'g': '7', 'h': '8', 'i': '9', 'j': '10', 'k': '11', 'l': '12', 'm': '13', 'n': '14', 'o': '15', 'p': '16', 'q': '17', 'r': '18', 's': '19', 't': '20', 'u': '21', 'v': '22', 'w': '23', 'x': '24', 'y': '25', 'z': '26'}
transtab = str.maketrans(dictionary)
print(transtab)
st = 'as you will see throughout this part of the book most objects have callable methods and all are accessed using this same method call syntax To call an object method as you will see in the following sections you have to go through an existing object'
print(st.translate(transtab))
import Tkinter
conda install Tkinter
st = 'I love python'
print("1:", st.endswith("n"))
print("2:",st.endswith("python"))
print("3:",st.endswith('n',0,6))
print("4:",st.endswith(""))
print('5:',st[0:6].endswith('n'))
print('6:',st[0:6].endswith('e'))
print("7:",st[0:6].endswith(""))
print('8:',st.endswith(("n","z")))
print("9:",st.endswith(("k","m")))
file = "python.txt"
if file.endswith("txt"):
print("该文件是文本文件")
elif file.endswith(("AVI","WMV","RM")):
print("该文件为视频文件")
else:
print("文件格式未知")
st.partition?
S = """As you’ll see throughout this part of the book, most objects have callable methods, and
all are accessed using this same method-call syntax. To call an object method, as you’ll
see in the following sections, you have to go through an existing object."""
S.partition('-')
S.expandtabs?
srt = "this is \t string example...wow!!!"
print("original string:" + srt)
print("Default expanded tab: " + srt.expandtabs())
print("Double expanded tab: " + srt.expandtabs(16))
S.replace?
print(S.replace('you','we'))
print(S.replace('you','we',1))
print(len(S))
S.find('you',5)
S.rfind('you') # Return the highest index in S where substring sub is found, such that sub is contained within S[start:end]. Optional arguments start and end are interpreted as in slice notation.
for i in range(251):
print(S.index('you',i))
S = """As you’ll see throughout this part of the book, most objects have callable methods, and
all are accessed using this same method-call syntax. To call an object method, as you’ll
see in the following sections, you have to go through an existing object."""
S.format?
print('{} {}'.format("hello","world"))
print("{0} {1} ".format("hello","world"))
print("{1} {0} {1} {0} {1} {0} {1} {0}".format("hello","world"))
print("网站名:{name}, 地址: {url} ".format(name="菜鸟教程", url = "www.runoob.com"))
site = {"name": "菜鸟教程","url": "www.runoob.com"}
print("网站名:{name},地址:{url}".format(**site))
my_list = ['菜鸟教程','www.runoob.com']
print("网站名:{0[0]}, 地址:{0[1]}".format(my_list))
S.rindex('you') # Return the highest index in S where substring sub is found, such that sub is contained within S[start:end]. Optional arguments start and end are interpreted as in slice notation
S.index('you') # Return the lowest index in S where substring sub is found, such that sub is contained within S[start:end]. Optional arguments start and end are interpreted as in slice notation.
print(S.rjust(4) + S)
print(len(S))
S.rjust(251,"O")
S.isalnum(), S.isalpha(), S.rpartition('and')
S.rsplit()
S.isalpha(), S.isdecimal(), S.isdigit()
S.rstrip?
print(S)
print(S.split('and'))
S.isidentifier(), S.islower()
S.splitlines()
S.isnumeric(), S.strip('C')
S.strip?
S.isprintable()
S[0] + S[1:].swapcase()
S.isspace()
S.title()
S.istitle()
dictionary = {'a': '1', 'b': '2', 'c': '3', 'd': '4', 'e': '5', 'f': '6', 'g': '7', 'h': '8', 'i': '9', 'j': '10', 'k': '11', 'l': '12', 'm': '13', 'n': '14', 'o': '15', 'p': '16', 'q': '17', 'r': '18', 's': '19', 't': '20', 'u': '21', 'v': '22', 'w': '23', 'x': '24', 'y': '25', 'z': '26'}
S.translate({'a': '1'})
S.isupper(), S.upper()
S.join?
S1 = S.split()
for i in S1:
print(i)
print('.'.join(str(print(i) for i in S1)))
S.zfill(251)
```
### String Method Examples: Changing Strings
```
S = 'spammy'
S = S[:3] + 'xx' + S[5:]
S
S = S.replace('mm','xx')
S
S = 'spammy'
S = S.replace('mm','xx')
S
'aa$bb$cc$dd'.replace("$","SPAM")
S = "xxxxSPAMxxxxSPAMxxxx"
where = S.find('SPAM')
where
S = S[:where] + "EGGS" + S[(where+4):]
S
S = 'xxxxSPAMxxxxSPAMxxxx'
print(S.replace('SPAM','EGGS'))
print(S.replace('SPAM',"EGGS",2))
S = 'spammy'
L = list(S)
S = """The fact that concatenation operations and the replace method generate new string
objects each time they are run is actually a potential downside of using them to change
strings. If you have to apply many changes to a very large string, you might be able to
improve your script’s performance by converting the string to an object that does support in-place changes"""
L = list(S)
L
'.'.join(list(S))
L = list(S)
L[3] = 'K'
L[5] = 'J'
L
```
If, after your changes, you need to convert back to a string (e.g., to write to a file), use the string join method to "implode" the list back into a string:
```
S = ''.join(L)
S
S2 = """The join method may look a bit backward at first sight. Because it is a method of strings
(not of lists), it is called through the desired delimiter. join puts the strings in a list (or
other iterable) together, with the delimiter between list items; in this case, it uses an
empty string delimiter to convert from a list back to a string. More generally, any string
delimiter and iterable of strings will do."""
L = list(S2)
L
''.join(L) == S2
print(''.join(L))
```
The join method may look a bit backward at first sight. Because it is a method of strings (not of lists), it is called through the desired delimiter. join puts the strings in a list (or other iterable) together, with the delimiter between list items; in this case, it uses an empty string delimiter to convert from a list back to a string. More generally, any string delimiter and iterable of strings will do.
```
' Python '.join(['C','C++','C#','Pascal','Fortran','java','php'])
```
### String Method Examples: Parsing Text
```
line = 'aaa bbb ccc'
col1 = line[0:3]
col3 = line[8:]
col1, col3
line = 'aaa bbb ccc'
cols = line.split()
cols
line = 'bob, hacker, 40'
line.split(',')
line = "I'mSPAMaSPAMlumberjack"
line.split("SPAM")
```
Other Common String Methods in Action
```
line = "The knights who say Ni!\n"
line.rstrip()
line.upper()
line.isalpha()
line.endswith("Ni!\n")
line.startswith('The')
line
line.find('Ni')!= -1
'Ni' in line
sub = ' Ni!\n'
line.endswith(sub)
line[-len(sub):] == sub
S = 'a + b + c'
x = S.replace('+','spam')
x
import string
dir(string)
string.capwords('we')
```
String Formatting Expressions
```
'That is %d %s bird !' % (1,'dead')
exclamation = "Ni"
"The knights who say %s !" % exclamation
"%d %s %d you " % (1,'spam', 4)
"%s -- %s -- %s " % (42, 3.14159, [1,2,3])
x = 1234
res = "integers: ...%d...%-6d...%06d" % (x,x,x)
res
x = 1.23456789
x
'%e | %f | %g ' % (x,x,x)
' %E ' % x
'%f, %.2f, %.*f' %(1/3.0,1/3.0,4,1/3.0)
```
### Dictionary-Based String Formatting Expressions
```
"%(n)d %(x)s" % {"n":1,"x":"spam"}
reply = """
Greetings...
Hello %(name)s!
Your age squared is
%(age)s
"""
values = {'name':'Bob','age':40}
print(reply%values)
food = 'spam'
age = 40
vars()
"%(age)d %(food)s" % vars()
```
#### String Formatting Method Calls
```
template = '{0},{1} and {2}'
template.format('spam','ham','eggs')
template = '{motto}, {pork} and {food} '
template.format(motto = 'spam', pork = 'ham', food = 'eggs')
template = '{motto}, {0} and {food}'
template.format('ham', motto= 'spam', food = 'eggs')
'{motto}, {0} and {food}'.format(42, motto = 3.14, food = [1,2])
X = '{motto}, {0} and {food}'.format(42,motto=3.14,food=[1,2])
X
X.split('and')
Y = X.replace('and', 'but under no circumstances')
Y
```
## Adding Keys, Attributes, and Offsets
```
import sys
'My {1[spam]} runs {0.platform}'.format(sys,{'spam':'laptop'})
'My {config[spam]} runs {sys.platform}'.format(sys=sys, config={'spam':'laptop'})
somelist = list('SPAM')
somelist
'first={0[0]},third={0[2]}'.format(somelist)
'first= {0},last = {1}'.format(somelist[0],somelist[-1]) #[-1] fails in fmt
parts = somelist[0], somelist[-1], somelist[1:3]
'first = {0}, last = {1}, middle = {2}'.format(*parts)
```
### Adding Specific Formatting
```
'{0:10} = {1:10}'.format('spam',123.4567)
'{0:>10} = {1: <10}'.format('spam',123.4567)
'{0.platform: >10} = {1[item]: <10}'.format(sys,dict(item = 'laptop'))
'{0:e}, {1:.3e}, {2:g}'.format(3.14159,3.14159,3.14159)
'{0:f}, {1:.2f}, {2:06.2f}'.format(3.14159, 3.14159, 3.14159)
'{0:X}, {1:o}, {2:b}'.format(255,255,255) # Hex, octal, binary
bin(255), int('111111111', 2), 0b11111111111 # Other to/from binary
hex(255), int('FF',16), 0xFF # Other to/from hex
oct(255), int('377',8), 0o377 # Other to/from octal, 0377 works in 2.6, not 3.0!
'{0:.2f}'.format(1/3.0) # Parameters hardcoded
'%.2f' % (1/3.0)
'{0:.{1}f}'.format(1/3.0,4) # Take value from arguments
'%.*f' % (4,1/3.0) # Ditto for expression
'{0:.2f}'.format(1.2345) # String method
format(1.2345, '.2f') # Built-in function
'%.2f' % 1.2345 # Expression
```
### Comparison to the % Formatting Expression
```
print('%s = %s'%('spam', 42)) # 2.X + format expression
print('{0}={1}'.format('spam',42)) # 3.0 (and 2.6) format method
template = '%s, %s, %s'
template % ('spam','ham','eggs') # By position
template = '%(motto)s, %(pork)s and %(food)s'
template % dict(motto='spam', pork='ham', food='eggs') # By key
'%s, %s and %s' % (3.14,42,[1,2]) # Arbitrary types
import sys
'My %(spam)s runs %(platform)s ' % {'spam': 'laptop','platform': sys.platform}
'Ny %(spam)s runs %(platform)s ' % dict(spam='laptop', platform = sys.platform)
somelist = list("SPAM")
parts = somelist[0],somelist[-1],somelist[1:3]
'first = %s, last = %s, middle = %s' % parts
# Adding specific formatting
'%-10s = %10s' % ('spam',123.4567)
'%10s = %-10s' %('spam', 123.4567)
'%(plat)10s = %(item)-10s' % dict(plat=sys.platform, item = 'laptop')
# Floating-point number
'%e, %.3e,%g' % (3.14159, 3.14159, 3.14159)
' %f, %.2f. %06.2f ' % (3.14159,3.14159, 3.14159)
# Hex and octal, but not binary
'%x, %o' % (255, 255)
import sys
'My {1[spam]: <8} runs {0.platform:>8}'.format(sys,{'spam':'laptop'})
' My %(spam)-8s runs %(plat)8s ' % dict(spam='laptop',plat=sys.platform)
# Building data ahead of time in both
data = dict(platform= sys.platform, spam= 'laptop')
'My {spam: <8} runs {platform:>8}'.format(**data)
'{0:d}'.format(999999999)
'{0:,d}'.format(9999999999999999999)
'{:,d}'.format(99999999999999999999999)
'{:,d}{:,d}'.format(99999,888888888324328)
'{:,.2f}'.format(243954904890.4395894)
'{0:b}'.format((2**16)-1)
' %b ' %((2**16)-1)
bin((2**16)-1)
' %s ' % bin((2**16)-1)[2:]
" %s " % bin((2**16)-1)[2:]
'The {0} side {1} {2}'.format('bright', 'of', 'life')
'The {} side {} {}'.format('bright', 'of', 'life') # Python 3.1
'The %s side %s %s' % ('bright', 'of',"life")
'{0:f}, {1:.2f}, {2:05.2f}'.format(3.14159,3.14159, 3.14159)
from math import *
'{:f}, {:.2f}, {:06.2f}'.format(pi, pi, pi)
```
### Method names and general arguments
```
'%.2f' % pi
'%.2f %s' % (1.2345, 99)
'%s' % 1.23
' %s ' % (1.23,)
'%s' % ((1.23,),)
'{0}'.format(1.23)
'{0}'.format((1.23,))
```
# Chapter Summary
In this chapter, we took an in-depth tour of the string object type. We learned about coding string literals, and we explored string operations, including sequence expressions, string method calls, and string formatting with both expressions and method calls. Along the way, we studied a variety of concepts in depth, such as slicing, method call syntax, and triple-quoted block strings. We also defined some core ideas common to a variety of types: sequences, for example, share an entire set of operations. In the next chapter, we’ll continue our types tour with a look at the most general object collections in Python—lists and dictionaries. As you’ll find, much of what you’ve learned here will apply to those types as well. And as mentioned earlier, in the final part of this book we’ll return to Python’s string model to flesh out the details of Unicode text and binary data, which are of interest to some, but not all, Python programmers. Before moving on, though, here’s another chapter quiz to review the material covered here.
| github_jupyter |
# Synthetic experiments with Monte Carlo: Examples
```
import numpy as np
from matplotlib import pyplot as plt
# The following is specific to Jupyter Notebooks (i.e., do not copy in Spyder)
from IPython.display import set_matplotlib_formats # makes plots in SVG instead of PNG
set_matplotlib_formats('svg') # makes plots in SVG instead of PNG
```
## The Nagel-Schreckenberg traffic model
The [Nagel-Schreckenberg traffic model](https://en.wikipedia.org/wiki/Nagel%E2%80%93Schreckenberg_model) is a simple model that predicts stop-and-go patterns we observe in traffic, even when there is no particularly bad driver around.
The idea is the following.
Time is discrete.
There are $N$ drivers.
A road is split into $M$ segments and each segment can host at most one car.
Sort the drivers $i \in \{1, \ldots, N\}$ such that $i=1$ is the last driver in the road and $i=N$ is the first driver in the road.
The road has a speed limit equal to $v_{\max}$.
Collisions between vehicles are prohibited, as well as overtaking.
Each driver moves ahead or stays in place, taking her position $x_{i,t} \in \{1, \ldots, M\}$ and adding her velocity $v_{i,t}$.
At each period $t$, every driver $i \in \{1, \ldots, N\}$ takes the following actions:
1. She checks the speed at which she was travelling, $v_{i,t-t}$. If $v_{i,t-t} < v_{\max}$, then she increases her speed by 1.
1. She checks the distance $d(x_{i,t}, x_{i+1,t})$ between herself and the driver ahead. If she is going too fast, then she reduces her speed to $d_{t}-1$ to avoid collision.
1. With some probability $p$, a given driver slows down by one velocity unit if her velocity was positive.
1. Finally, the driver moves into the next period by setting $x_{i,t+1} = x_{i,t} + v_{i,t}$.
The third step is the fundamental one in this model.
It captures the idea that each driver sometimes behaves randomly.
It is also the step that allows us to create a simulation.
In this context, a simulation is a track record of the position of each driver $i$ at each period $t$.
To create a simulation, we need three basic ingredients: an initial condition, rules to govern the behavior of variables and a source of randomness.
In the traffic model above, we described the rules of the game and the source of randomness.
We need to set an initial condition.
As we have no instruction to do so, we set it arbitrarily (just keeping in mind that one "road slot" can host only one driver).
Next, we need to code the rules, from a "narrative" form (the one above) to a formal set of instructions (in Python).
Let us translate the rules above in mathematical terms:
1. $\bar{v}_{i,t} = \min\{v_{i,t-1} + 1; v_{\max}\}$
1. $\hat{v}_{i,t} = \min\{\bar{v}_{i,t}; d(x_{i,t}, x_{i+1,t}) - 1\}$
1. $v_{i,t} = \max\{0; \hat{v}_{i,t} - 1\}$
1. $x_{i,t+1} = x_{i,t} + v_{i,t}$
We are left with two edge cases: what happens to the first driver $i=N$ when she reaches the end of the road and how we compute the distance between the first driver and the last driver $i=1$.
We will assume that if $x_{N,t} + v_{N,t} > M$, then $x_{N,t+1} = x_{N,t} + v_{N,t}$, and that $d(x_{N,t}, x_{1,t}) = M + x_{1,t} - x_{N,t}$.
This effectively represents a circular road.
Now let us turn to representing the road, the drivers and their rules in Python.
We can think of the road as a vector with $M$ elements.
In this vector, we store two values: `0` to indicate that that "road slot" has no driver in it, and `1` to indicate there is a driver at that location.
We also need to keep track of the speed of each driver.
We can do this by using another vector with $M$ elements, where we find `0` if there is no driver or the driver is waiting, or the speed `v` if the driver in that location is travelling.
The rules will be represented by a function, that takes as input the state of the road at period $t$ and returns the state of the road at period $t+1$.
This function will include all the four rules above, plus the rules governing the edge cases.
It will also include the source of randomness.
It is useful to start thinking with functions.
First, we can code the rules according to which each driver adjusts her speed.
At this stage, we can forget the edge cases.
We will simply code rules 1 to 3, almost literally.
The function we will write will take the current speed of an individual driver, the distance between that driver and the next one, the speed limit and the probability of slowing down as input arguments.
As output, we will obtain the new speed at which this individual driver will travel, taking the three rules into account.
All input and output arguments of this function are expected to be scalar integers, because they pertain an individual driver.
Rule no. 4 will be taken care of later.
```
def adjust_speed(current_speed, next_driver_distance, speed_limit, prob_slowdown):
v_bar = np.min([current_speed + 1, speed_limit]) # rule no. 1
v_hat = np.min([v_bar, next_driver_distance - 1]) # rule no. 2
uniform_random_number = np.random.uniform()
if uniform_random_number < prob_slowdown: # rule no. 3
v = np.max([0, v_hat - 1])
else:
v = v_hat
new_speed = v
return new_speed
```
Next, we code the function that considers the current state of the whole road, applies the rules above to all drivers and returns the new road, after movement occurred.
We will code here rule no. 4, including the edge cases.
This function will consider the current state of the road, the current speeds of each driver, the speed limit, the probability of slowdown as input arguments.
The output will be a vector representing the state of the road in the upcoming period.
All inputs and outputs will be vectors, with the exception of the speed limit and the slowdown probability.
Inside this function, we need to apply the `adjust_speed` function to each driver on the road.
Hence, we need a `for` loop along the road.
```
def period_transition(current_road, current_speeds, speed_limit, prob_slowdown):
M = current_road.size
drivers_locations = np.where(current_road == 1)[0]
new_road = np.zeros_like(current_road, dtype=int)
new_speeds = np.zeros_like(current_speeds, dtype=int)
for l in drivers_locations:
if l != drivers_locations[-1]:
space_ahead = road[l+1] - road[l]
else: # hitting an edge case
space_ahead = M - road[l] + road[drivers_locations[0]]
new_speeds[l] = adjust_speed(current_speeds[l], space_ahead, speed_limit, prob_slowdown)
for l in drivers_locations:
if new_speeds[l] <= (M - 1) - l:
new_road[l + new_speeds[l]] = 1
else: # hitting an edge case
new_road[new_speeds[l] - M + l] = 1
return (new_road, new_speeds)
```
Finally, we can write the code that sets up the parameters of the problem and runs the simulation over periods.
For the simulation, all we need to do is to apply the functions we wrote above, in a loop that runs from period $t=1$ to $T$.
Because Python starts counting integers from zero (as opposed to one), we will loop from `t=0` to `T-1`.
```
# Parameters of simulation
T = 1000 # no. of simulated periods
# Parameters of the model
v_max = 5 # max speed
M = 1000 # road segments
N = 50 # drivers
p = 0.3 # probability of slowing down
# Creating useful vectors with placeholder content (preallocation)
road = np.zeros(M, dtype=int) # an un-oriented vector
speeds = np.zeros(M, dtype=int) # an un-oriented vector
history_road = np.zeros((T, M), dtype=int) # a [T-by-M] matrix
history_speeds = np.zeros((T, M), dtype=int) # a [T-by-M] matrix
# Creating the initial condition: equispaced drivers
initial_drivers_distance = M // N
for j in range(M):
if j % initial_drivers_distance == 0:
road[j] = 1
# Running the simulation
for t in range(T):
# Save current states to histories, for track records
history_road[t, :] = road
history_speeds[t, :]
# Compute the policy functions, i.e., new road state and speeds
new_road, new_speeds = period_transition(road, speeds, v_max, p)
# Update the model conditions for next period
road = new_road
speeds = new_speeds
```
To see the implications of the model, we can plot the history of the road state.
```
plt.imshow(history_road, aspect='equal')
plt.tight_layout()
```
| github_jupyter |
<span style='color black'>
# Stericycle
### Company
#### - Stericycle is a compliance company that specializes in collecting and disposing regulated substances, such as medical waste and sharps, pharmaceuticals, hazardous waste, and providing services for recalled and expired goods.
#### - It also provides related education and training services, and patient communication services.
```
import warnings
warnings.filterwarnings('ignore') # Hide warnings
import datetime as dt
import pandas as pd
pd.core.common.is_list_like = pd.api.types.is_list_like
import pandas_datareader.data as web
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from mpl_finance import candlestick_ohlc
import matplotlib.dates as mdates
#SRCL
#Getting stock price data
start = dt.datetime(2000, 1, 1)
end = dt.datetime.now()
df = web.DataReader("SRCL", 'yahoo', start, end) # Collects data
df.reset_index(inplace=True)
df.set_index("Date", inplace=True)
#prices in USD
df.head()
df.tail()
#Making the data as csv
df.to_csv("SRCL_data.csv")
#Reading the same data from local csv file
data=pd.read_csv("SRCL_data.csv" ,parse_dates=True, index_col=0)
plt.figure(figsize=(10, 6))
plt.plot(data["Open"])
plt.title('Stock Opening Value (USD)', fontsize = 16)
plt.xlabel('Time')
plt.ylabel('Stock Value')
plt.figure(figsize=(10, 6))
plt.plot(data["Close"])
plt.title('Stock Closing Value (USD)', fontsize = 16)
plt.xlabel('Time')
plt.ylabel('Stock Value')
```
# Stock Data Manipulation
```
data
# Create, and add 100 day moving average to dataframe
data['100ma'] = data['Close'].rolling(window=100,min_periods=0).mean()
data.head()
plt.figure(figsize=(10, 6))
plt.plot(data[["100ma","Close"]])
plt.title('Stock Closing Value (USD) and Moving Average(100 Day) ', fontsize = 16)
plt.legend(labels=data[["100ma","Close"]])
plt.xlabel('Time')
plt.ylabel('Stock Value')
# Create, and add 180 day moving average to dataframe
data['180ma'] = data['Close'].rolling(window=180,min_periods=0).mean()
data.head()
plt.figure(figsize=(10, 6))
plt.plot(data[["180ma","Close"]])
plt.title('Stock Closing Value (USD) and Moving Average(180 Day) ', fontsize = 16)
plt.legend(labels=data[["180ma","Close"]])
plt.xlabel('Time')
plt.ylabel('Stock Value')
# Create, and add 360 day moving average to dataframe
data['360ma'] = data['Close'].rolling(window=360,min_periods=0).mean()
data.head()
plt.figure(figsize=(10, 6))
plt.plot(data[["360ma","Close"]])
plt.title('Stock Closing Value (USD) and Moving Average(360 Day) ', fontsize = 16)
plt.legend(labels=data[["360ma","Close"]])
plt.xlabel('Time')
plt.ylabel('Stock Value')
```
# Resampling
```
# Resample to get open-high-low-close (OHLC) on every 50 days of data
df_ohlc_50 = data.Close.resample('50D').ohlc()
df_volume_50 = data.Volume.resample('50D').sum()
df_ohlc_50.head()
df_volume_50.head()
df_ohlc_50.reset_index(inplace=True)
df_ohlc_50.Date = df_ohlc_50.Date.map(mdates.date2num)
df_ohlc_50.head()
```
# CandleStick Charts
```
# Create and visualize candlestick charts
plt.figure(figsize=(10,5))
ax1 = plt.subplot2grid((6,1), (0,0), rowspan=5, colspan=1)
ax1.xaxis_date()
candlestick_ohlc(ax1, df_ohlc_50.values, width=2, colorup='b')
plt.show()
#taking other dateframe data
#SRCL
#Getting stock price data
start = dt.datetime(2016, 1, 1)
end = dt.datetime.now()
df = web.DataReader("SRCL", 'yahoo', start, end)
df.reset_index(inplace=True)
df.set_index("Date", inplace=True)
#prices in USD
#Making the data as csv
df.to_csv("SRCL_data_2016.csv")
#Reading the same data from local csv file
data_2016=pd.read_csv("SRCL_data_2016.csv" ,parse_dates=True, index_col=0)
data_2016.head()
```
# 10 days candlestick
```
# Resample to get open-high-low-close (OHLC) on every 10 days of data
df_ohlc_10 = data_2016.Close.resample('10D').ohlc()
df_volume_10 = data_2016.Volume.resample('10D').sum()
df_ohlc_10.reset_index(inplace=True)
df_ohlc_10.Date = df_ohlc_10.Date.map(mdates.date2num)
df_ohlc_10.head()
# Create and visualize candlestick charts
plt.figure(figsize=(10,5))
ax1 = plt.subplot2grid((6,1), (0,0), rowspan=5, colspan=1)
ax1.xaxis_date()
candlestick_ohlc(ax1, df_ohlc_10.values, width=2, colorup='Y')
plt.title("10 DAYS OHLC CANDLESTICK")
plt.show()
```
# 20 days candlestick
```
# Resample to get open-high-low-close (OHLC) on every 20 days of data
df_ohlc_20 = data_2016.Close.resample('20D').ohlc()
df_volume_20 = data_2016.Volume.resample('20D').sum()
df_ohlc_20.reset_index(inplace=True)
df_ohlc_20.Date = df_ohlc_20.Date.map(mdates.date2num)
df_ohlc_20.head()
# Create and visualize candlestick charts
plt.figure(figsize=(10,5))
ax1 = plt.subplot2grid((6,1), (0,0), rowspan=5, colspan=1)
ax1.xaxis_date()
candlestick_ohlc(ax1, df_ohlc_20.values, width=2, colorup='g')
plt.title("20 DAYS OHLC CANDLESTICK")
plt.show()
```
| github_jupyter |
**Name:** \_\_\_\_\_
**EID:** \_\_\_\_\_
# CS5489 - Tutorial 3
## Gender Classification from Face Images
In this tutorial you will train a classifier to predict whether a face image is male or female.
First we need to initialize Python. Run the below cell.
```
%matplotlib inline
import IPython.core.display
# setup output image format (Chrome works best)
IPython.core.display.set_matplotlib_formats("svg")
import matplotlib.pyplot as plt
import matplotlib
from numpy import *
from sklearn import *
import os
import zipfile
import fnmatch
random.seed(100)
```
## 1. Loading Data and Pre-processing
We first need to load the images. Download `photos-bw.zip` and put it in the same directory as this ipynb file. **Do not unzip the file.** Then run the following cell to load the images.
```
imgdata = []
genders = []
# load the zip file
filename = 'photos-bw.zip'
zfile = zipfile.ZipFile(filename, 'r')
for name in zfile.namelist():
# check file name matches
if fnmatch.fnmatch(name, "photos-bw/*.png"):
print("loading", name)
# open file in memory, and parse as an image
myfile = zfile.open(name)
img = matplotlib.image.imread(myfile)
myfile.close()
# append to data
imgdata.append(img)
genders.append( int(name[len("photos-bw/")] == 'm') ) # 0 is female, 1 is male
zfile.close()
imgsize = img.shape
print("DONE: loaded {} images".format(len(imgdata)))
```
Each image is a 45x40 array of pixel values. Run the below code to show an example:
```
print(img.shape)
plt.imshow(img, cmap='gray', interpolation='nearest')
plt.show()
```
Run the below code to show all the images!
```
# function to make an image montage
def image_montage(X, imsize=None, maxw=10):
"""X can be a list of images, or a matrix of vectorized images.
Specify imsize when X is a matrix."""
tmp = []
numimgs = len(X)
# create a list of images (reshape if necessary)
for i in range(0,numimgs):
if imsize != None:
tmp.append(X[i].reshape(imsize))
else:
tmp.append(X[i])
# add blanks
if (numimgs > maxw) and (mod(numimgs, maxw) > 0):
leftover = maxw - mod(numimgs, maxw)
meanimg = 0.5*(X[0].max()+X[0].min())
for i in range(0,leftover):
tmp.append(ones(tmp[0].shape)*meanimg)
# make the montage
tmp2 = []
for i in range(0,len(tmp),maxw):
tmp2.append( hstack(tmp[i:i+maxw]) )
montimg = vstack(tmp2)
return montimg
plt.figure(figsize=(9,9))
plt.imshow(image_montage(imgdata), cmap='gray', interpolation='nearest')
plt.show()
```
Each image is a 2d array, but the classifier algorithms work on 1d vectors. Run the following code to convert all the images into 1d vectors by flattening. The result should be a matrix where each row is a flattened image.
```
X = empty((50, prod(imgdata[0].shape))) # create empty array
for i,img in enumerate(imgdata):
X[i,:] = ravel(img) # for each image, turn it into a vector
Y = asarray(genders) # convert list to numpy array
print(X.shape)
print(Y.shape)
```
Next we will shift the pixel values so that gray is 0.0, black is -0.5 and white is 0.5.
```
print("Before: min={}, max={}".format(X.min(), X.max()))
X -= 0.5
print("After: min={}, max={}".format(X.min(), X.max()))
```
Finally, split the dataset into a training set and testing set. We select 80% for training and 20% for testing.
```
# randomly split data into 80% train and 20% test set
trainX, testX, trainY, testY = \
model_selection.train_test_split(X, Y,
train_size=0.80, test_size=0.20, random_state=4487)
print(trainX.shape)
print(testX.shape)
# compute the average image for later
avgX = mean(X,axis=0)
```
# 2. Logistic Regression
Train a logistic regression classifier. Use cross-validation to select the best C parameter.
```
### INSERT YOUR CODE HERE
```
Use the learned model to predict the genders for the training and testing data. What is the accuracy on the training set? What is the accuracy on the testing set?
```
### INSERT YOUR CODE HERE
```
### Analyzing the classifier
Run the below code to show the hyperplane parameter $\mathbf{w}$ as an image.
```
# logreg is the learned logistic regression model
wimg = logreg.coef_.reshape(imgsize) # get the w and reshape into an image
mycmap = matplotlib.colors.LinearSegmentedColormap.from_list('mycmap', ["#0000FF", "#FFFFFF", "#FF0000"])
mm = max(wimg.max(), -wimg.min())
plt.figure(figsize=(8,3))
plt.subplot(1,2,1)
plt.title('average image')
plt.imshow(avgX.reshape(imgsize), cmap='gray')
plt.subplot(1,2,2)
plt.imshow(wimg, interpolation='nearest', cmap=mycmap, vmin=-mm, vmax=mm)
plt.colorbar()
plt.title("LR weight image")
plt.show()
```
Recall that the classifier prediction is based on the sign of the function $f(\mathbf{x}) = \mathbf{w}^T\mathbf{x}+b = \sum_{i=1}^P w_ix_i + b$. Here each $x_i$ is a pixel in the face image, and $w_i$ is the corresponding weight. Hence, the function is multiplying face image by the weight image, and then summing over all pixels.
In order for $f(\mathbf{x})$ to be positive, then the positive values of the weight image (red regions) should match the positive values in the face image (white pixels), and the negative values of the weight image (blue regions) should be matched with negative values in the face image (black pixels).
Hence, we can have the following interpretation:
<table>
<tr><th>Class</th><th>red regions (positive weights)</th><th>blue regions (negative weights)</th><th>white regions (weights near 0)</th></tr>
<tr><td>+1 class (male)</td><td>white pixels in face image</td><td>black pixels in face image</td><td>region not important</td></tr>
<tr><td>-1 class (female)</td><td>black pixels in face image</td><td>white pixels in face image</td><td>region not important</td></tr>
</table>
_Looking at the weight image, what parts of the face image is the classifier looking at to determine the gender? Does it make sense?_
- **INSERT YOUR ANSWER HERE**
Now let's look at the misclassified faces in the test set. Run the below code to show the misclassifed and correctly classified faces.
```
# predYtest are the class predictions on the test set.
# find misclassified test images
inds = where(predYtest != testY) # get indices of misclassified test images
# make a montage
badimgs = image_montage(testX[inds], imsize=imgsize)
# find correctly classified test images
inds = where(predYtest == testY)
goodimgs = image_montage(testX[inds], imsize=imgsize)
plt.figure(figsize=(8,4))
plt.subplot(2,1,1)
plt.imshow(badimgs, cmap='gray', interpolation='nearest')
plt.title('misclassified faces')
plt.subplot(2,1,2)
plt.imshow(goodimgs, cmap='gray', interpolation='nearest')
plt.title('correctly classified faces')
plt.show()
```
_Why did the classifier make incorrect predictions on the misclassified faces?_
- **INSERT YOUR ANSWER HERE**
# 3. Support Vector Machine
Now train a support vector machine (SVM) on the same training and testing data. Use cross-validation to select the best $C$ parameter.
```
### INSERT YOUR CODE HERE
```
Calculate the training and test accuracy for the SVM classifier.
```
### INSERT YOUR CODE HERE
```
Similar to before, plot an image of the hyperplane parameters $w$, and view the misclassified and correctly classified test images.
```
### INSERT YOUR CODE HERE
```
_Are there any differences between the $w$ for logistic regressiona and the $w$ for SVM? Is there any interpretation for the differences?_
- **INSERT YOUR ANSWER HERE**
# 4. Classifying cropped faces
It seems that the hair around the face and forehead are discriminative enough to perform gender classifixation. Now try to perform the same task but only focusing on the face image, and not the hair.
First, we define a mask over the face.
```
imgmask = full((img.shape), False)
#imgmask[17:41,8:32] = True
#masksize = (24,24)
imgmask[18:40,11:29] = True
masksize = (22,18)
plt.imshow(imgmask)
```
Next we crop out the face image to create the new inputs. The vectors are now 576-dim, and corresponding images are 24x24.
```
Xm = X[:,imgmask.ravel()]
Xm.shape
```
Here are the cropped images.
```
plt.imshow(image_montage(Xm, imsize=masksize), cmap='gray', interpolation='nearest')
```
Generate the same training/test split as before
```
# randomly split data into 80% train and 20% test set
trainXm, testXm, trainY, testY = \
model_selection.train_test_split(Xm, Y,
train_size=0.80, test_size=0.20, random_state=4487)
print(trainXm.shape)
print(testXm.shape)
avgXm = avgX[imgmask.ravel()]
```
Now train logistic regression and SVM classifiers on the new cropped images.
```
### INSERT YOUR CODE HERE
### INSERT YOUR CODE HERE
### INSERT YOUR CODE HERE
```
Compute the train/test accuracies of the two classifiers.
```
### INSERT YOUR CODE HERE
### INSERT YOUR CODE HERE
```
Compare the classification performance? Which performs better and why?
- **INSERT YOUR ANSWER HERE**
Visualize the weights as an image and interpret what discriminative information each classifier is using.
```
### INSERT YOUR CODE HERE
### INSERT YOUR CODE HERE
```
- **INSERT YOUR ANSWER HERE**
| github_jupyter |
```
import numpy as np
import tensorflow as tf
from tcn import TCN
import os
os.chdir("../src/")
from models import tcn
from utils import auxiliary_plots, metrics
from preprocessing import normalization, data_generation
SEED = 1
tf.random.set_seed(SEED)
np.random.seed(SEED)
os.chdir("../notebooks/")
TRAIN_FILE_NAME = '../data/hourly_20140102_20191101_train.csv'
TEST_FILE_NAME = '../data/hourly_20140102_20191101_test.csv'
FORECAST_HORIZON = 24
PAST_HISTORY = 192
BATCH_SIZE = 256
BUFFER_SIZE = 10000
EPOCHS = 25
METRICS = ['mape']
TCN_PARAMS = {
'nb_filters': 128,
'kernel_size': 3,
'nb_stacks': 1,
'dilations': [1, 2, 4, 8, 16, 32, 64],
'dropout_rate': 0,
}
```
## Read data
```
# Read train file
with open(TRAIN_FILE_NAME, 'r') as datafile:
ts_train = datafile.readlines()[1:] # skip the header
ts_train = np.asarray([np.asarray(l.rstrip().split(',')[0], dtype=np.float32) for l in ts_train])
ts_train = np.reshape(ts_train, (ts_train.shape[0],))
# Read test data file
with open(TEST_FILE_NAME, 'r') as datafile:
ts_test = datafile.readlines()[1:] # skip the header
ts_test = np.asarray([np.asarray(l.rstrip().split(',')[0], dtype=np.float32) for l in ts_test])
ts_test = np.reshape(ts_test, (ts_test.shape[0],))
# Train/validation split
TRAIN_SPLIT = int(ts_train.shape[0] * 0.8)
# Normalize training data
norm_params = normalization.get_normalization_params(ts_train[:TRAIN_SPLIT])
ts_train = normalization.normalize(ts_train, norm_params)
# Normalize test data with train params
ts_test = normalization.normalize(ts_test, norm_params)
# Get x and y for training and validation
x_train, y_train = data_generation.univariate_data(ts_train, 0, TRAIN_SPLIT, PAST_HISTORY, FORECAST_HORIZON)
x_val, y_val = data_generation.univariate_data(ts_train, TRAIN_SPLIT - PAST_HISTORY, ts_train.shape[0],
PAST_HISTORY, FORECAST_HORIZON)
# Get x and y for test data
x_test, y_test = data_generation.univariate_data(ts_test, 0, ts_test.shape[0], PAST_HISTORY, FORECAST_HORIZON)
# Convert numpy data to tensorflow dataset
train_data = tf.data.Dataset.from_tensor_slices((x_train, y_train)).cache().shuffle(BUFFER_SIZE).batch(BATCH_SIZE).repeat()
val_data = tf.data.Dataset.from_tensor_slices((x_val, y_val)).batch(BATCH_SIZE).repeat()
test_data = tf.data.Dataset.from_tensor_slices((x_test, y_test)).batch(BATCH_SIZE)
```
## TCN: Create and train model
```
model = tcn(x_train.shape, FORECAST_HORIZON, 'adam', 'mae',
nb_filters=TCN_PARAMS['nb_filters'],
kernel_size=TCN_PARAMS['kernel_size'],
nb_stacks= TCN_PARAMS['nb_stacks'],
dilations=TCN_PARAMS['dilations'],
dropout_rate=TCN_PARAMS['dropout_rate'])
model.summary()
TRAIN_MODEL = True
checkpoint_path = "training_tcn/cp.ckpt"
checkpoint_dir = os.path.dirname(checkpoint_path)
if TRAIN_MODEL:
cp_callback = tf.keras.callbacks.ModelCheckpoint(filepath=checkpoint_path,
save_weights_only=True,
verbose=1)
evaluation_interval = int(np.ceil(x_train.shape[0] / BATCH_SIZE))
history = model.fit(train_data,
epochs=EPOCHS,
steps_per_epoch=evaluation_interval,
validation_data=val_data, validation_steps=evaluation_interval,
callbacks=[cp_callback])
auxiliary_plots.plot_training_history(history, ['loss'])
model.load_weights(checkpoint_path)
```
## Validation results
```
val_forecast = model.predict(x_val)
val_forecast = normalization.denormalize(val_forecast, norm_params)
y_val_denormalized = normalization.denormalize(y_val, norm_params)
val_metrics = metrics.evaluate(y_val_denormalized, val_forecast, METRICS)
print('Validation scores', val_metrics)
```
## Test results
```
test_forecast = model.predict(test_data)
test_forecast = normalization.denormalize(test_forecast, norm_params)
y_test_denormalized = normalization.denormalize(y_test, norm_params)
x_test_denormalized = normalization.denormalize(x_test, norm_params)
test_metrics = metrics.evaluate(y_test_denormalized, test_forecast, METRICS)
print('Test scores', test_metrics)
```
| github_jupyter |
```
import os
import re
import sys
import MeCab
nm = MeCab.Tagger('-Owakati')
# txtファイルを全部読み込んで分かち書きやクリーニングをしていく
files = [f for f in os.listdir('./dataset/raw_datas') if f.split('.')[1] == 'txt']
authors = {'dazai': [], 'mori': [], 'akutagawa': []}
cleaned_data = []
def clean(raw_text):
"""
ルビや入力者注を削除して一文毎に分割,分かち書き
"""
text = raw_text.replace('\n', '').replace('\u3000', '')
text = re.sub('[[《]', '/', text)
text = re.sub('[]》]', '\', text)
text = re.sub('/[^\]*?\', '', text)
text = text.replace('。', '。\n')
text = text.replace('「', '')
text = text.replace('」', '\n').split('\n')
return [nm.parse(t).split(' ') for t in text if t]
for f in files:
name = f.split('.')[0]
print(name)
with open('./dataset/raw_datas/'+str(f), 'r') as f:
clean_text = clean(f.read())
authors[name] = clean_text
cleaned_data += clean_text
def make_stopdic(lines):
"""
空白区切りの文の集まりのテキストのリストからストップワードの辞書を作成する。
"""
calc_words = {}
for line in lines:
for word in line:
if word in calc_words:
calc_words[str(word)] += 1
else:
calc_words[str(word)] = 1
sorted_stop = sorted(calc_words.items(), key=lambda x:x[1], reverse=True)
print("sorted_stop:"+str(len(sorted_stop)))
freq_num = int(len(sorted_stop)*0.03)
n = 0
stop_words = []
print("freq_num:"+str(freq_num))
for data in sorted_stop:
stop_words.append(str(data[0]))
#print('High frequency word: ',str(data[0]), str(data[1]))
n += 1
if n > freq_num:
break
# 作成したストップワードの辞書の保存
with open('./origin_stopwords.txt', 'w') as f:
f.write('\n'.join(stop_words))
make_stopdic(cleaned_data)
print("cleaned_data:"+str(len(cleaned_data)))
# ストップワードリストの作成
stop_data = {}
for f in files:
with open('./dataset/raw_datas/'+str(f)) as f1:
print(f)
nm = MeCab.Tagger('-Owakati')
text = nm.parse(f1.read()).split(' ')
for word in text:
if word in stop_data:
stop_data[str(word)] += 1
else:
stop_data[str(word)] = 1
sorted_stop = sorted(stop_data.items(), key=lambda x:x[1], reverse=True)
print(len(sorted_stop))
freq_num = int(len(sorted_stop)*0.03)
n = 0
stop_words = []
print(freq_num)
for data in sorted_stop:
stop_words.append(str(data[0]))
print('High frequency word: ',str(data[0]), str(data[1]))
n+= 1
if n > freq_num:
break
# 作成したストップワードの辞書の保存
with open('./origin_stopwords.txt', 'w') as f:
f.write('\n'.join(stop_words))
def stopword_bydic(text):
"""
辞書によるストップワードの除去
"""
# 読み込むストップワード辞書の指定。
with open('./origin_stopwords.txt') as f:
data = f.read()
stopwords = data.split('\n')
lines = []
for line in text:
words = []
for word in line:
if word not in stopwords:
words.append(word)
lines.append(words)
return lines
sum_lines = len(stopword_bydic(cleaned_data))
sum_words = []
for line in stopword_bydic(cleaned_data):
for word in line:
if word not in sum_words:
sum_words.append(word)
print('sum_lines: ', sum_lines)
print('sum_words: ', len(sum_words))
with open('origin_words.txt', 'w') as f:
f.write('\n'.join(sum_words))
cleaned_data = []
for author, data in authors.items():
data = stopword_bydic(data)
for line in data:
#if len(line) > 2:
cleaned_data.append(author+','+' '.join(line))
# 作成したコーパスの保存
with open('./dataset/corpus.csv', 'w') as f:
f.write(''.join(cleaned_data))
```
| github_jupyter |
**For tqdm progress bar to work correctly, before launching this notebook run:**
```bash
$ jupyter nbextension enable --py --sys-prefix widgetsnbextension
```
**Or simply run from project root:**
```bash
$ make jupyter
```
### imports
```
import numpy as np
import matplotlib.pyplot as plt
from sklearn.linear_model import LogisticRegression
from sklearn.svm import SVC
from sklearn.neighbors import KNeighborsClassifier as KNN
from sklearn.metrics import accuracy_score, confusion_matrix
import env
from boltzmann_machines.rbm import BernoulliRBM
from boltzmann_machines.utils import (Stopwatch,
im_plot, plot_confusion_matrix)
from boltzmann_machines.utils.dataset import load_mnist
%matplotlib inline
%load_ext autoreload
%autoreload 2
```
## load trained RBM model
```
rbm = BernoulliRBM.load_model('../models/rbm_mnist/')
```
# visualize learned filters
```
weights = rbm.get_tf_params(scope='weights')
W = weights['W']
hb = weights['hb']
fig = plt.figure(figsize=(10, 10))
im_plot(W.T, shape=(28, 28), title='First 100 filters extracted by RBM',
imshow_params={'cmap': plt.cm.gray});
plt.savefig('rbm_mnist.png', dpi=196, bbox_inches='tight');
```
## load data
```
X, y = load_mnist(mode='train', path='../data/')
X /= 255.
X_test, y_test = load_mnist(mode='test', path='../data/')
X_test /= 255.
print X.shape, y.shape, X_test.shape, y_test.shape
```
# 1) train classifiers on extracted features
## extract features $\mathbf{q}_i=p(\mathbf{h}\;|\;\mathbf{v}=\mathbf{x}_i)$
```
Q = rbm.transform(X)
Q_test = rbm.transform(X_test)
print Q.shape, Q_test.shape
```
## fit k-NN
```
knn = KNN(n_neighbors=3, p=2., weights='distance',
n_jobs=2)
with Stopwatch(verbose=True) as s:
knn.fit(Q, y)
print accuracy_score(y_test, knn.predict(Q_test))
```
## fit Logistic Regression
```
logreg = LogisticRegression(multi_class='multinomial', solver='sag', max_iter=800,
n_jobs=2, verbose=10, random_state=1337) # displays progress in terminal
logreg.fit(Q, y)
print accuracy_score(y_test, logreg.predict(Q_test))
```
## fit SVM
```
svc = SVC(C=1e3, tol=1e-6,
verbose=10, random_state=1337) # displays progress in terminal
with Stopwatch(verbose=True) as s:
svc.fit(Q, y)
print accuracy_score(y_test, svc.predict(Q_test))
```
# 2) discriminative finetuning: initialize 2-layer MLP with learned parameters $\boldsymbol{\psi}$ and train using backprop
## load predictions, targets, and fine-tuned weights
```
y_pred = np.load('../data/rbm_y_pred.npy')
y_test = np.load('../data/rbm_y_test.npy')
W_finetuned = np.load('../data/rbm_W_finetuned.npy')
print accuracy_score(y_test, y_pred)
```
## plot confusion matrix
```
C = confusion_matrix(y_test, y_pred)
fig = plt.figure(figsize=(10, 8))
ax = plot_confusion_matrix(C, fmt='d')
plt.title('Confusion matrix for fine-tuned RBM\n', fontsize=20, y=0.97)
plt.savefig('rbm_mnist_confusion_matrix.png', dpi=144, bbox_inches='tight')
```
## visualize filters after fine-tuning
```
fig = plt.figure(figsize=(10, 10))
im_plot(W_finetuned.T, shape=(28, 28), title='First 100 filters after fine-tuning',
imshow_params={'cmap': plt.cm.gray});
plt.savefig('rbm_mnist_finetuned.png', dpi=196, bbox_inches='tight');
```
# 3) classification RBMs
A third way to use RBMs for supervised problems is to jointly model distribution of the data along with the labels $\mathbb{P}(\mathbf{x},\;\mathbf{y})$. Such models are called **classification RBMs** (cRBMs). Currently, these are beyond of the scope of this repository.
| github_jupyter |
# Custom Header Routing with Seldon and Ambassador
This notebook shows how you can deploy Seldon Deployments which can have custom routing via Ambassador's custom header routing.
## Setup Seldon Core
Use the setup notebook to [Setup Cluster](https://docs.seldon.io/projects/seldon-core/en/latest/examples/seldon_core_setup.html#Setup-Cluster) with [Ambassador Ingress](https://docs.seldon.io/projects/seldon-core/en/latest/examples/seldon_core_setup.html#Ambassador) and [Install Seldon Core](https://docs.seldon.io/projects/seldon-core/en/latest/examples/seldon_core_setup.html#Install-Seldon-Core). Instructions [also online](https://docs.seldon.io/projects/seldon-core/en/latest/examples/seldon_core_setup.html).
```
!kubectl create namespace seldon
!kubectl config set-context $(kubectl config current-context) --namespace=seldon
from IPython.core.magic import register_line_cell_magic
@register_line_cell_magic
def writetemplate(line, cell):
with open(line, "w") as f:
f.write(cell.format(**globals()))
VERSION = !cat ../../../version.txt
VERSION = VERSION[0]
VERSION
```
## Launch main model
We will create a very simple Seldon Deployment with a dummy model image `seldonio/mock_classifier:1.0`. This deployment is named `example`.
```
%%writetemplate model.yaml
apiVersion: machinelearning.seldon.io/v1alpha2
kind: SeldonDeployment
metadata:
labels:
app: seldon
name: example
spec:
name: production-model
predictors:
- componentSpecs:
- spec:
containers:
- image: seldonio/mock_classifier:{VERSION}
imagePullPolicy: IfNotPresent
name: classifier
terminationGracePeriodSeconds: 1
graph:
children: []
endpoint:
type: REST
name: classifier
type: MODEL
name: single
replicas: 1
!kubectl create -f model.yaml
!kubectl rollout status deploy/$(kubectl get deploy -l seldon-deployment-id=example -o jsonpath='{.items[0].metadata.name}')
```
### Get predictions
```
from seldon_core.seldon_client import SeldonClient
sc = SeldonClient(deployment_name="example", namespace="seldon")
```
#### REST Request
```
r = sc.predict(gateway="ambassador", transport="rest")
assert r.success == True
print(r)
```
## Launch Model with Custom Routing
We will now create a new graph for our Canary with a new model `seldonio/mock_classifier_rest:1.1`. To make it a canary of the original `example` deployment we add two annotations
```
"annotations": {
"seldon.io/ambassador-header":"location:london"
"seldon.io/ambassador-service-name":"example"
},
```
The first annotation says we want to route traffic that has the header `location:london`. The second says we want to use `example` as our service endpoint rather than the default which would be our deployment name - in this case `example-canary`. This will ensure that this Ambassador setting will apply to the same prefix as the previous one.
```
%%writetemplate model_with_header.yaml
apiVersion: machinelearning.seldon.io/v1alpha2
kind: SeldonDeployment
metadata:
labels:
app: seldon
name: example-header
spec:
annotations:
seldon.io/ambassador-header: 'location: london'
seldon.io/ambassador-service-name: example
name: header-model
predictors:
- componentSpecs:
- spec:
containers:
- image: seldonio/mock_classifier:{VERSION}
imagePullPolicy: IfNotPresent
name: classifier
terminationGracePeriodSeconds: 1
graph:
children: []
endpoint:
type: REST
name: classifier
type: MODEL
name: single
replicas: 1
!kubectl create -f model_with_header.yaml
!kubectl rollout status deploy/$(kubectl get deploy -l seldon-deployment-id=example-header -o jsonpath='{.items[0].metadata.name}')
```
Check a request without a header goes to the existing model.
```
r = sc.predict(gateway="ambassador", transport="rest")
print(r)
default_count = !kubectl logs $(kubectl get pod -lseldon-app=example-single -o jsonpath='{.items[0].metadata.name}') classifier | grep "root.predict" | wc -l
print(default_count)
assert int(default_count[0]) == 2
```
Check a REST request with the required header gets routed to the new model.
```
r = sc.predict(gateway="ambassador", transport="rest", headers={"location": "london"})
print(r)
header_count = !kubectl logs $(kubectl get pod -lseldon-app=example-header-single -o jsonpath='{.items[0].metadata.name}') classifier | grep "root.predict" | wc -l
print(header_count)
assert int(header_count[0]) == 1
!kubectl delete -f model.yaml
!kubectl delete -f model_with_header.yaml
```
| github_jupyter |
## Inference using the trained model
```
import os
os.environ["CUDA_VISIBLE_DEVICES"] = ""
import math
import json
from IPython.display import Image, display
import random
import jieba
import numpy as np
import tensorflow as tf
import config
from im2txt_model import Im2TxtModel
import inference_wrapper
from inference_utils import caption_generator
from inference_utils import vocabulary
checkpoint_path="../model/semantic_attention_model_attr_only_wb/model.ckpt-36850"
#vocab_file="../data/word_counts.txt"
#attributes_file="../data/attributes.txt"
data_path="../data/"
# dev set
devset_path = os.path.join(data_path, "ai_challenger_caption_validation_20170910")
dev_image_path = os.path.join(devset_path, "caption_validation_images_20170910")
dev_annotation_path = os.path.join(devset_path, "caption_validation_annotations_20170910.json")
FLAGS = tf.flags.FLAGS
tf.flags.DEFINE_string("vocab_file", "../data/word_counts.txt", "Text file containing the vocabulary.")
tf.flags.DEFINE_string("attributes_file", "../data/attributes.txt", "Text file containing the attributes.")
FLAGS.model = "SemanticAttentionModel"
FLAGS.attributes_top_k = 15
# load model
g = tf.Graph()
with g.as_default():
tf.logging.info("Building model.")
model = Im2TxtModel(mode="inference")
model.build()
saver = tf.train.Saver()
if tf.gfile.IsDirectory(checkpoint_path):
checkpoint_path = tf.train.latest_checkpoint(checkpoint_path)
if not checkpoint_path:
raise ValueError("No checkpoint file found in: %s" % checkpoint_path)
def restore_fn(sess):
tf.logging.info("Loading model from checkpoint: %s", checkpoint_path)
saver.restore(sess, checkpoint_path)
tf.logging.info("Successfully loaded checkpoint: %s",
os.path.basename(checkpoint_path))
g.finalize()
sess = tf.Session(graph=g)
restore_fn(sess)
vocab = vocabulary.Vocabulary(FLAGS.vocab_file)
def show_example(example, image_path):
image_id = example['image_id']
caption = example['caption']
im = Image(filename=os.path.join(image_path, image_id))
display(im)
for c in caption:
c = c.strip().strip(u"。").replace('\n', '')
seg_list = jieba.cut(c, cut_all=False)
print ' '.join(seg_list)
#return image_id, caption
# read dev data
input = open(dev_annotation_path,'r')
dev_examples = json.load(input)
input.close()
dev_size = len(dev_examples)
# run a random image
index = int(random.random() * dev_size)
show_example(dev_examples[index], dev_image_path)
filename = os.path.join(dev_image_path, dev_examples[index]['image_id'])
f = tf.gfile.GFile(filename, "r")
image=f.read()
predicted_ids, scores, top_n_attributes = sess.run(
[model.predicted_ids, model.scores, model.top_n_attributes],
feed_dict={"image_feed:0": image})
predicted_ids = np.transpose(predicted_ids, (0,2,1))
scores = np.transpose(scores, (0,2,1))
attr_probs, attr_ids = top_n_attributes
attributes = [vocab.id_to_word(w) for w in attr_ids[0]]
print(" ".join(attributes))
print(attr_probs[0])
#print(top_n_attributes)
for caption in predicted_ids[0]:
print(caption)
caption = [id for id in caption if id >= 0 and id != FLAGS.end_token]
sent = [vocab.id_to_word(w) for w in caption]
print(" ".join(sent))
```
| github_jupyter |
# Modules and Packages
In this section we briefly:
* code out a basic module and show how to import it into a Python script
* run a Python script from a Jupyter cell
* show how command line arguments can be passed into a script
Check out the video lectures for more info and resources for this.
The best online resource is the official docs:
https://docs.python.org/3/tutorial/modules.html#packages
But I really like the info here: https://python4astronomers.github.io/installation/packages.html
## Writing modules
```
%%writefile file1.py
def myfunc(x):
return [num for num in range(x) if num%2==0]
list1 = myfunc(11)
```
**file1.py** is going to be used as a module.
Note that it doesn't print or return anything,
it just defines a function called *myfunc* and a variable called *list1*.
## Writing scripts
```
%%writefile file2.py
import file1
file1.list1.append(12)
print(file1.list1)
```
**file2.py** is a Python script.
First, we import our **file1** module (note the lack of a .py extension)<br>
Next, we access the *list1* variable inside **file1**, and perform a list method on it.<br>
`.append(12)` proves we're working with a Python list object, and not just a string.<br>
Finally, we tell our script to print the modified list.
## Running scripts
```
! python file2.py
```
Here we run our script from the command line. The exclamation point is a Jupyter trick that lets you run command line statements from inside a jupyter cell.
```
import file1
print(file1.list1)
```
The above cell proves that we never altered **file1.py**, we just appended a number to the list *after* it was brought into **file2**.
## Passing command line arguments
Python's `sys` module gives you access to command line arguments when calling scripts.
```
%%writefile file3.py
import sys
import file1
num = int(sys.argv[1])
print(file1.myfunc(num))
```
Note that we selected the second item in the list of arguments with `sys.argv[1]`.<br>
This is because the list created with `sys.argv` always starts with the name of the file being used.<br>
```
! python file3.py 21
```
Here we're passing 21 to be the upper range value used by the *myfunc* function in **list1.py**
## Understanding modules
Modules in Python are simply Python files with the .py extension, which implement a set of functions. Modules are imported from other modules using the <code>import</code> command.
To import a module, we use the <code>import</code> command. Check out the full list of built-in modules in the Python standard library [here](https://docs.python.org/3/py-modindex.html).
The first time a module is loaded into a running Python script, it is initialized by executing the code in the module once. If another module in your code imports the same module again, it will not be loaded twice but once only - so local variables inside the module act as a "singleton" - they are initialized only once.
If we want to import the math module, we simply import the name of the module:
```
# import the library
import math
# use it (ceiling rounding)
math.ceil(2.4)
```
## Exploring built-in modules
Two very important functions come in handy when exploring modules in Python - the <code>dir</code> and <code>help</code> functions.
We can look for which functions are implemented in each module by using the <code>dir</code> function:
```
print(dir(math))
```
When we find the function in the module we want to use, we can read about it more using the <code>help</code> function, inside the Python interpreter:
```
help(math.ceil)
```
## Writing modules
Writing Python modules is very simple. To create a module of your own, simply create a new .py file with the module name, and then import it using the Python file name (without the .py extension) using the import command.
## Writing packages
Packages are name-spaces which contain multiple packages and modules themselves. They are simply directories, but with a twist.
Each package in Python is a directory which MUST contain a special file called **\__init\__.py**. This file can be empty, and it indicates that the directory it contains is a Python package, so it can be imported the same way a module can be imported.
If we create a directory called foo, which marks the package name, we can then create a module inside that package called bar. We also must not forget to add the **\__init\__.py** file inside the foo directory.
To use the module bar, we can import it in two ways:
```
# Just an example, this won't work
import foo.bar
# OR could do it this way
from foo import bar
```
In the first method, we must use the foo prefix whenever we access the module bar. In the second method, we don't, because we import the module to our module's name-space.
The **\__init\__.py** file can also decide which modules the package exports as the API, while keeping other modules internal, by overriding the **\__all\__** variable, like so:
```
__init__.py:
__all__ = ["bar"]
```
| github_jupyter |
# Test localization and co-localization of two diseases, using network propagation
Test on simulated networks, where we can control how localized and co-localized node sets are
### Author: Brin Rosenthal (sbrosenthal@ucsd.edu)
```
import numpy as np
import matplotlib.pyplot as plt
import seaborn
import networkx as nx
import pandas as pd
import random
import scipy
import mygene
mg = mygene.MyGeneInfo()
# latex rendering of text in graphs
import matplotlib as mpl
mpl.rc('text', usetex = False)
mpl.rc('font', family = 'serif')
import sys
sys.path.append('../source')
import plotting_results
import network_prop
import imp
imp.reload(plotting_results)
imp.reload(network_prop)
% matplotlib inline
```
## First, let's create a random graph
- We will start with the connected Watts Strogatz random graph, created using the NetworkX package. This graph generator will allow us to create random graphs which are guaranteed to be fully connected, and gives us control over how connected the graph is, and how structured it is. Documentation for the function can be found here https://networkx.github.io/documentation/latest/reference/generated/networkx.generators.random_graphs.connected_watts_strogatz_graph.html#networkx.generators.random_graphs.connected_watts_strogatz_graph
<img src="screenshots/connected_watts_strogatz_graph_nx_docs.png" width="600" height="600">
## Control localization
- We can control the localization of nodes by seeding the network propagation with a focal node and that focal node's neighbors. This will guarantee that the seed nodes will be very localized in the graph
- As a first example, let's create a random network, with two localized sets.
- The network contains 100 nodes, with each node first connected to its 5 nearest neighbors.
- Once these first edges are connected, each edge is randomly rewired with probability p = 0.12 (so approximately 12 percent of the edges in the graph will be rewired)
- With this rewiring probability of 0.12, most of the structure in the graph is maintained, but some randomness has been introduced
```
# Create a random connected-Watts-Strogatz graph
Gsim = nx.connected_watts_strogatz_graph(100,5,.12)
seed1 = [0]
seed1.extend(nx.neighbors(Gsim,seed1[0]))
seed2 = [10]
seed2.extend(nx.neighbors(Gsim,seed2[0]))
#seed = list(np.random.choice(Gsim.nodes(),size=6,replace=False))
pos = nx.spring_layout(Gsim)
nx.draw_networkx_nodes(Gsim,pos=pos,node_size=100,alpha=.5,node_color = 'blue')
nx.draw_networkx_nodes(Gsim,pos=pos,nodelist=seed1,node_size=120,alpha=.9,node_color='orange',linewidths=3)
nx.draw_networkx_nodes(Gsim,pos=pos,nodelist=seed2,node_size=120,alpha=.9,node_color='red',linewidths=3)
nx.draw_networkx_edges(Gsim,pos=pos,alpha=.1)
plt.grid('off')
#plt.savefig('/Users/brin/Google Drive/UCSD/update_16_03/non_colocalization_illustration.png',dpi=300,bbox_inches='tight')
```
- In the network shown above, we plot our random connected Watts-Strogatz graph, highlighting two localized seed node sets, shown in red and orange, with bold outlines.
- These seed node sets were created by selecting two focal node, and those focal node's neighbors, thus resulting in two node sets which appear highly localized to the eye.
- Since the graph is composed of nearest neighbor relations (with some randomness added on), and it was initiated with node ids ranging from 0 to 99 (these are the default node names- they can be changed using nx.relabel_nodes()), we can control the co-localization of these node sets by selecting seed nodes which are close together, for high co-localization (e.g. 0 and 5), or which are far apart, for low co-localization (e.g. 0 and 50).
- Below, we will display node sets with both high and low co-localization
- Our ability to control the co-localization in this way will become worse as the rewiring probability increases, and the structure in the graph is destroyed.
```
# highly co-localized gene sets
seed1 = [0]
seed1.extend(nx.neighbors(Gsim,seed1[0]))
seed2 = [5]
seed2.extend(nx.neighbors(Gsim,seed2[0]))
#seed = list(np.random.choice(Gsim.nodes(),size=6,replace=False))
plt.figure(figsize=(12,5))
plt.subplot(1,2,1)
nx.draw_networkx_nodes(Gsim,pos=pos,node_size=100,alpha=.5,node_color = 'blue')
nx.draw_networkx_nodes(Gsim,pos=pos,nodelist=seed1,node_size=120,alpha=.9,node_color='orange',linewidths=3)
nx.draw_networkx_nodes(Gsim,pos=pos,nodelist=seed2,node_size=120,alpha=.9,node_color='red',linewidths=3)
nx.draw_networkx_edges(Gsim,pos=pos,alpha=.1)
plt.title('High Co-localization',fontsize=16)
plt.grid('off')
# low co-localized gene sets
seed1 = [5]
seed1.extend(nx.neighbors(Gsim,seed1[0]))
seed2 = [30]
seed2.extend(nx.neighbors(Gsim,seed2[0]))
#seed = list(np.random.choice(Gsim.nodes(),size=6,replace=False))
plt.subplot(1,2,2)
nx.draw_networkx_nodes(Gsim,pos=pos,node_size=100,alpha=.5,node_color = 'blue')
nx.draw_networkx_nodes(Gsim,pos=pos,nodelist=seed1,node_size=120,alpha=.9,node_color='orange',linewidths=3)
nx.draw_networkx_nodes(Gsim,pos=pos,nodelist=seed2,node_size=120,alpha=.9,node_color='red',linewidths=3)
nx.draw_networkx_edges(Gsim,pos=pos,alpha=.1)
plt.title('Low Co-localization',fontsize=16)
plt.grid('off')
```
# Can we quantify this concept of localization?
- Sometimes it's not easy to tell by eye if a node set is localized.
- We can use network propagation simulations to quantify this concept of localization
- Network propagation is a tool which initiates a seed node set with high 'heat', and then over the course of a number of iterations spreads this heat around to nearby nodes.
- At the end of the simulation, nodes with the highest heat are those which are most closely related to the seed nodes.
- We implemented the network propagation method described in Vanunu et. al. 2010 (Vanunu, Oron, et al. "Associating genes and protein complexes with disease via network propagation." PLoS Comput Biol 6.1 (2010): e1000641.)
<img src="screenshots/vanunu_abstracg.png">
### Localization using network propagation
- We can use network propagation to evaluate how localized a seed node set is in the network.
- If the seed node set is highly localized, the 'heat' from the network propagation simulation will be bounced around between seed nodes, and less of it will dissipate to distant parts of the network.
- We will evaluate the distribution of the heat from all the nodes, using the kurtosis (the fourth standardized moment), which measures how 'tailed' the distribution is. If our distribution has high kurtosis, this indicates that much of the 'heat' has stayed localized near the seed set. If our distribution has a low kurtosis, this indicates that the 'heat' has not stayed localized, but has diffused to distant parts of the network.
<img src="screenshots/kurtosis.png">
### Random baseline for comparison
- To evaluate localization in this way, we need a baseline to compare to.
- Te establish the baseline we take our original network, and shuffle the edges, while preserving degree (so nodes which originally had 5 neighbors will still have 5 neighbors, although these neighbors will now be spread randomly throughout the graph)
- For example, below we show the heat propagation on a non-shuffled graph, from a localized seed set (left), and the heat propagation from the same seed set, on an edge-shuffled graph (right). The nodes on the left and right have the same _number_ of neighbors, but they have different identities.
- The total amount of heat in the graph is conserved in both cases, but the heat distributions look very different- the seed nodes retain much less of their original heat in the edge-shuffled case.
<img src="screenshots/L_edge_shuffled.png">
- We will calculate the kurtosis of the heat distribution over a large number of different edge-shuffled networks (below- 1000 repetitions), to build up the baseline distribution of kurtosis values.
```
Wprime_ring = network_prop.normalized_adj_matrix(Gsim)
Fnew_ring = network_prop.network_propagation(Gsim,Wprime_ring,seed1)
plt.figure(figsize=(18,5))
plt.subplot(1,3,1)
nx.draw_networkx_nodes(Gsim,pos=pos,node_size=100,alpha=.5,node_color=Fnew_ring[Gsim.nodes()],cmap='jet',
vmin=0,vmax=max(Fnew_ring))
nx.draw_networkx_edges(Gsim,pos=pos,alpha=.2)
var_ring = plotting_results.nsf(np.var(Fnew_ring),3)
kurt_ring = plotting_results.nsf(scipy.stats.kurtosis(Fnew_ring),3)
plt.annotate('kurtosis = ' + str(kurt_ring),
xy=(.08,.1),xycoords='figure fraction')
plt.annotate('Heat: original',xy=(.08,.93),xycoords='figure fraction',fontsize=16)
plt.xticks([],[])
plt.yticks([],[])
plt.grid('off')
num_reps = 1000
var_rand_list,kurt_rand_list = [],[]
for r in range(num_reps):
G_temp = nx.configuration_model(Gsim.degree().values())
G_rand = nx.Graph() # switch from multigraph to digraph
G_rand.add_edges_from(G_temp.edges())
G_rand = nx.relabel_nodes(G_rand,dict(zip(range(len(G_rand.nodes())),Gsim.degree().keys())))
Wprime_rand = network_prop.normalized_adj_matrix(G_rand)
Fnew_rand = network_prop.network_propagation(G_rand,Wprime_rand,seed1)
var_rand_list.append(np.var(Fnew_rand))
kurt_rand_list.append(scipy.stats.kurtosis(Fnew_rand))
plt.subplot(1,3,2)
nx.draw_networkx_nodes(G_rand,pos=pos,node_size=100,alpha=.5,node_color=Fnew_rand[G_rand.nodes()],cmap='jet',
vmin=0,vmax=max(Fnew_ring))
nx.draw_networkx_edges(G_rand,pos=pos,alpha=.2)
var_rand = plotting_results.nsf(np.var(Fnew_rand),3)
kurt_rand = plotting_results.nsf(scipy.stats.kurtosis(Fnew_rand),3)
plt.annotate('kurtosis = ' + str(kurt_rand),
xy=(.40,.1),xycoords='figure fraction')
plt.annotate('Heat: edge-shuffled',xy=(.40,.93),xycoords='figure fraction',fontsize=16)
plt.xticks([],[])
plt.yticks([],[])
plt.grid('off')
plt.subplot(1,3,3)
plt.boxplot(kurt_rand_list)
z_score = (kurt_ring-np.mean(kurt_rand_list))/np.std(kurt_rand_list)
z_score = plotting_results.nsf(z_score,n=2)
plt.plot(1,kurt_ring,'*',color='darkorange',markersize=16,label='original: \nz-score = '+ str(z_score))
plt.annotate('Kurtosis',xy=(.73,.93),xycoords='figure fraction',fontsize=16)
plt.legend(loc='lower left')
#plt.savefig('/Users/brin/Google Drive/UCSD/update_16_03/localization_NWS_p1_variance.png',dpi=300,bbox_inches='tight')
```
- Above (right panel) we see that when a node set is highly localized, it has a higher kurtosis value than would be expected from a non-localized gene set (the orange star represents the kurtosis of the heat distribution on the original graph, and the boxplot represents the distribution of 1000 kurtosis values on edge-shuffled networks). The orange star is significantly higher than the baseline distribution.
# Co-localization using network propagation
- We now build on our understanding of localization using network propagation to establish a measurement of how _co-localized_ two node sets are in a network.
- In the first example we discussed (above), we came up with a general understanding of co-localization, where two node sets were co-localized if they were individually localized, _and_ were nearby in network space.
- In order to measure this co-localization using network propagation, we will first seed 2 simulations with each node set, then we will take the dot-product (or the sum of the pairwise product) of the resulting heat vectors.
- When node sets are co-localized, there will be more nodes which are hot in both heat vectors (again we compare to a distribution of heat dot-products on degree preserving edge-shuffled graphs)
```
seed1 = Gsim.nodes()[0:5] #nx.neighbors(Gsim,Gsim.nodes()[0])
seed2 = Gsim.nodes()[10:15] #nx.neighbors(Gsim,Gsim.nodes()[5]) #Gsim.nodes()[27:32]
seed3 = Gsim.nodes()[20:25]
Fnew1 = network_prop.network_propagation(Gsim,Wprime_ring,seed1,alpha=.9,num_its=20)
Fnew2 = network_prop.network_propagation(Gsim,Wprime_ring,seed2,alpha=.9,num_its=20)
F12 = Fnew1*Fnew2
F12.sort(ascending=False)
#Fnew1.sort(ascending=False)
#Fnew2.sort(ascending=False)
Fnew1_norm = Fnew1/np.linalg.norm(Fnew1)
Fnew2_norm = Fnew2/np.linalg.norm(Fnew2)
dot_12 = np.sum(F12.head(10))
print(dot_12)
plt.figure(figsize=(18,6))
plt.subplot(1,3,1)
nx.draw_networkx_nodes(Gsim,pos=pos,node_size=100,alpha=.5,node_color=Fnew1[Gsim.nodes()],
cmap='jet', vmin=0,vmax=max(Fnew1))
nx.draw_networkx_nodes(Gsim,pos=pos,nodelist=seed1,
node_size=100,alpha=.9,node_color=Fnew1[seed1],
cmap='jet', vmin=0,vmax=max(Fnew1),linewidths=3)
nx.draw_networkx_edges(Gsim,pos=pos,alpha=.2)
plt.grid('off')
plt.xticks([],[])
plt.yticks([],[])
plt.annotate('Heat: nodes A ($H_A$)',xy=(.08,.93),xycoords='figure fraction',fontsize=16)
plt.subplot(1,3,2)
nx.draw_networkx_nodes(Gsim,pos=pos,node_size=100,alpha=.5,node_color=Fnew2[Gsim.nodes()],
cmap='jet', vmin=0,vmax=max(Fnew1))
nx.draw_networkx_nodes(Gsim,pos=pos,nodelist=seed2,
node_size=100,alpha=.9,node_color=Fnew2[seed2],
cmap='jet', vmin=0,vmax=max(Fnew1),linewidths=3)
nx.draw_networkx_edges(Gsim,pos=pos,alpha=.2)
plt.grid('off')
plt.xticks([],[])
plt.yticks([],[])
plt.annotate('Heat: nodes B ($H_B$)',xy=(.4,.93),xycoords='figure fraction',fontsize=16)
plt.subplot(1,3,3)
nx.draw_networkx_nodes(Gsim,pos=pos,node_size=100,alpha=.5,node_color=Fnew1[Gsim.nodes()]*Fnew2[Gsim.nodes()],
cmap='jet', vmin=0,vmax=max(Fnew1*Fnew2))
nx.draw_networkx_nodes(Gsim,pos=pos,nodelist=seed2,
node_size=100,alpha=.9,node_color=Fnew1[seed2]*Fnew2[seed2],
cmap='jet', vmin=0,vmax=max(Fnew1*Fnew2),linewidths=3)
nx.draw_networkx_nodes(Gsim,pos=pos,nodelist=seed1,
node_size=100,alpha=.9,node_color=Fnew1[seed1]*Fnew2[seed1],
cmap='jet', vmin=0,vmax=max(Fnew1*Fnew2),linewidths=3)
nx.draw_networkx_edges(Gsim,pos=pos,alpha=.2)
plt.grid('off')
plt.xticks([],[])
plt.yticks([],[])
plt.annotate('$H_A \cdot H_B$',xy=(.73,.93),xycoords='figure fraction',fontsize=16)
```
- In the figure above, we show an example of this co-localization concept.
- In the left panel we show the heat vector of the simulation seeded by node set A (warmer colors indicate hotter nodes, and bold outlines indicate seed nodes)
- In the middle panel we show the heat vector of the simulation seeded by node set B
- The right panel shows the pairwise product of the heat vectors (note color scale is different for this panel). The nodes between the two seed sets are the hottest, meaning that these are the nodes most likely related to both seed gene sets.
- If these node sets are truly co-localized, then the sum of the heat product (the dot product) will be higher than random. This is what we will test below.
```
results_dict = network_prop.calc_3way_colocalization(Gsim,seed1,seed2,seed3,num_reps=100,num_genes=5,
replace=False,savefile=False,alpha=.5,print_flag=False,)
import scipy
num_reps = results_dict['num_reps']
dot_sfari_epi=results_dict['sfari_epi']
dot_sfari_epi_rand=results_dict['sfari_epi_rand']
#U,p = scipy.stats.mannwhitneyu(dot_sfari_epi,dot_sfari_epi_rand)
t,p = scipy.stats.ttest_ind(dot_sfari_epi,dot_sfari_epi_rand)
psig_SE = plotting_results.nsf(p,n=2)
plt.figure(figsize=(7,5))
plt.errorbar(-.1,np.mean(dot_sfari_epi_rand),2*np.std(dot_sfari_epi_rand)/np.sqrt(num_reps),fmt='o',
ecolor='gray',markerfacecolor='gray',label='edge-shuffled graph')
plt.errorbar(0,np.mean(dot_sfari_epi),2*np.std(dot_sfari_epi)/np.sqrt(num_reps),fmt='bo',
label='original graph')
plt.xlim(-.8,.5)
plt.legend(loc='lower left',fontsize=12)
plt.xticks([0],['A-B \np='+str(psig_SE)],rotation=45,fontsize=12)
plt.ylabel('$H_{A} \cdot H_{B}$',fontsize=18)
```
- In the figure above, we show the heat dot product of node set A and node set B, on the original graph (blue dot), and on 100 edge-shuffled graphs (gray dot with error bars).
- Using a two sided independent t-test, we find that the dot product on the original graph is significantly higher than on the edge-shuffled graphs, if node sets A and B are indeed co-localized.
# Can we control how co-localized two node sets are?
- We can use a parameter in our random graph generator function to control the co-localization of two node sets.
- By varying the rewiring probability, we can move from a graph which is highly structured (low p-rewire: mostly nearest neighbor connections), to a graph which is mostly random (high p-rewire: mostly random connections).
- In the following section we will sweet through values of p-rewire, ranging from 0 to 1, and measure the co-localization of identical node sets.
```
H12 = []
H12_rand = []
num_G_reps=5
for p_rewire in np.linspace(0,1,5):
print('rewiring probability = ' + str(p_rewire) + '...')
H12_temp = []
H12_temp_rand = []
for r in range(num_G_reps):
Gsim = nx.connected_watts_strogatz_graph(500,5,p_rewire)
seed1 = Gsim.nodes()[0:5]
seed2 = Gsim.nodes()[5:10]
seed3 = Gsim.nodes()[20:30]
results_dict = network_prop.calc_3way_colocalization(Gsim,seed1,seed2,seed3,num_reps=20,num_genes=5,
replace=False,savefile=False,alpha=.5,print_flag=False)
H12_temp.append(np.mean(results_dict['sfari_epi']))
H12_temp_rand.append(np.mean(results_dict['sfari_epi_rand']))
H12.append(np.mean(H12_temp))
H12_rand.append(np.mean(H12_temp_rand))
plt.plot(np.linspace(0,1,5),H12,'r.-',label='original')
plt.plot(np.linspace(0,1,5),H12_rand,'.-',color='gray',label='edge-shuffled')
plt.xlabel('link rewiring probability',fontsize=14)
plt.ylabel('$H_A \cdot H_B$',fontsize=16)
plt.legend(loc='upper right',fontsize=12)
```
- We see above, as expected, that as the rewiring probability increases (on the x-axis), and the graph becomes more random, the heat dot-product (co-localization) decreases (on the y-axis), until the co-localization on the original graph matches the edge-shuffled graph.
- We expect this to be the case because once p-rewire becomes very high, the original graph becomes essentially random, so not much is changed by shuffling the edges.
# Three-way Co-localization
- Finally, we will look at how our co-localization using network propagation method applies to three seed node sets instead of two.
- This could be useful if the user was interested in establishing if one node set provided a link between two other node sets. For example, one might find that two node sets are individually not co-localized, but each is co-localized with a third node set. This third node set would essentially provide the missing link between the two, as illustrated below, where node sets A and C are far apart, but B is close to A, and B is close to C.
<img src="screenshots/CL_triangle.png">
```
seed1 = Gsim.nodes()[0:5]
seed2 = Gsim.nodes()[5:10]
seed3 = Gsim.nodes()[10:15]
results_dict = network_prop.calc_3way_colocalization(Gsim,seed1,seed2,seed3,num_reps=100,num_genes=5,
replace=False,savefile=False,alpha=.5,print_flag=False,)
import scipy
num_reps = results_dict['num_reps']
dot_sfari_epi=results_dict['sfari_epi']
dot_sfari_epi_rand=results_dict['sfari_epi_rand']
#U,p = scipy.stats.mannwhitneyu(dot_sfari_epi,dot_sfari_epi_rand)
t,p = scipy.stats.ttest_ind(dot_sfari_epi,dot_sfari_epi_rand)
psig_SE = plotting_results.nsf(p,n=2)
dot_sfari_aem=results_dict['sfari_aem']
dot_aem_sfari_rand=results_dict['aem_sfari_rand']
#U,p = scipy.stats.mannwhitneyu(dot_sfari_aem,dot_aem_sfari_rand)
t,p = scipy.stats.ttest_ind(dot_sfari_aem,dot_aem_sfari_rand)
psig_SA = plotting_results.nsf(p,n=2)
dot_aem_epi=results_dict['aem_epi']
dot_aem_epi_rand=results_dict['aem_epi_rand']
#U,p = scipy.stats.mannwhitneyu(dot_aem_epi,dot_aem_epi_rand)
t,p = scipy.stats.ttest_ind(dot_aem_epi,dot_aem_epi_rand)
psig_AE = plotting_results.nsf(p,n=2)
plt.figure(figsize=(7,5))
plt.errorbar(-.1,np.mean(dot_sfari_epi_rand),2*np.std(dot_sfari_epi_rand)/np.sqrt(num_reps),fmt='o',
ecolor='gray',markerfacecolor='gray')
plt.errorbar(0,np.mean(dot_sfari_epi),2*np.std(dot_sfari_epi)/np.sqrt(num_reps),fmt='bo')
plt.errorbar(.9,np.mean(dot_aem_sfari_rand),2*np.std(dot_aem_sfari_rand)/np.sqrt(num_reps),fmt='o',
ecolor='gray',markerfacecolor='gray')
plt.errorbar(1,np.mean(dot_sfari_aem),2*np.std(dot_sfari_aem)/np.sqrt(num_reps),fmt='ro')
plt.errorbar(1.9,np.mean(dot_aem_epi_rand),2*np.std(dot_aem_epi_rand)/np.sqrt(num_reps),fmt='o',
ecolor='gray',markerfacecolor='gray')
plt.errorbar(2,np.mean(dot_aem_epi),2*np.std(dot_aem_epi)/np.sqrt(num_reps),fmt='go')
plt.xticks([0,1,2],['A-B \np='+str(psig_SE),'A-C \np='+str(psig_SA),'B-C\np='+str(psig_AE)],rotation=45,fontsize=12)
plt.xlim(-.5,2.5)
plt.ylabel('$H_{1} \cdot H_{2}$',fontsize=18)
```
- In the figure above, we show how three-way co-localization looks in practice.
- We have selected three node sets, two of which are distant (A and C), and one which is close to both (B).
- We find that indeed $H_A\cdot H_B$ and $H_B\cdot H_C$ (blue dot and green dot) are much higher on the original graph than on the edge shuffled graphs.
- However, we find that $H_A\cdot H_C$ is actually _lower_ than the background noise. This is telling us that node sets A and C are actually individually localized, but not co-localized at all, because more of the heat remains close to each individual seed set than would happen if each node set was not individually co-localized.
| github_jupyter |
ref: https://github.com/rickiepark/handson-ml2
# Decision Tree
```
# 파이썬 ≥3.5 필수
import sys
assert sys.version_info >= (3, 5)
# 사이킷런 ≥0.20 필수
import sklearn
assert sklearn.__version__ >= "0.20"
# 공통 모듈 임포트
import numpy as np
import os
# 노트북 실행 결과를 동일하게 유지하기 위해
np.random.seed(42)
# 깔끔한 그래프 출력을 위해
%matplotlib inline
import matplotlib as mpl
import matplotlib.pyplot as plt
mpl.rc('axes', labelsize=14)
mpl.rc('xtick', labelsize=12)
mpl.rc('ytick', labelsize=12)
# 그림을 저장할 위치
PROJECT_ROOT_DIR = "."
CHAPTER_ID = "decision_trees"
IMAGES_PATH = os.path.join(PROJECT_ROOT_DIR, "images", CHAPTER_ID)
os.makedirs(IMAGES_PATH, exist_ok=True)
def save_fig(fig_id, tight_layout=True, fig_extension="png", resolution=300):
path = os.path.join(IMAGES_PATH, fig_id + "." + fig_extension)
print("그림 저장:", fig_id)
if tight_layout:
plt.tight_layout()
plt.savefig(path, format=fig_extension, dpi=resolution)
```
# 훈련과 시각화
```
from sklearn.datasets import load_iris
from sklearn.tree import DecisionTreeClassifier
iris = load_iris()
X = iris.data[:, 2:] # 꽃잎 길이와 너비
y = iris.target
tree_clf = DecisionTreeClassifier(max_depth=2, random_state=42) #설정하지 않았으면 gini(Gini impurity)
tree_clf.fit(X, y)
from graphviz import Source
from sklearn.tree import export_graphviz #시각화 툴 (아래 tree를 보기 쉽게 그려줌)
export_graphviz(
tree_clf,
out_file=os.path.join(IMAGES_PATH, "iris_tree.dot"),
feature_names=iris.feature_names[2:],
class_names=iris.target_names,
rounded=True,
filled=True
)
Source.from_file(os.path.join(IMAGES_PATH, "iris_tree.dot")) ##fetal length와 width만을 가지고 분류
from matplotlib.colors import ListedColormap
def plot_decision_boundary(clf, X, y, axes=[0, 7.5, 0, 3], iris=True, legend=False, plot_training=True):
x1s = np.linspace(axes[0], axes[1], 100)
x2s = np.linspace(axes[2], axes[3], 100)
x1, x2 = np.meshgrid(x1s, x2s)
X_new = np.c_[x1.ravel(), x2.ravel()]
y_pred = clf.predict(X_new).reshape(x1.shape) #predict 함수를 사용해서 y_prediction 값을 얻음
custom_cmap = ListedColormap(['#fafab0','#9898ff','#a0faa0'])
plt.contourf(x1, x2, y_pred, alpha=0.3, cmap=custom_cmap)
if not iris:
custom_cmap2 = ListedColormap(['#7d7d58','#4c4c7f','#507d50'])
plt.contour(x1, x2, y_pred, cmap=custom_cmap2, alpha=0.8)
if plot_training:
plt.plot(X[:, 0][y==0], X[:, 1][y==0], "yo", label="Iris setosa")
plt.plot(X[:, 0][y==1], X[:, 1][y==1], "bs", label="Iris versicolor")
plt.plot(X[:, 0][y==2], X[:, 1][y==2], "g^", label="Iris virginica")
plt.axis(axes)
if iris:
plt.xlabel("Petal length", fontsize=14)
plt.ylabel("Petal width", fontsize=14)
else:
plt.xlabel(r"$x_1$", fontsize=18)
plt.ylabel(r"$x_2$", fontsize=18, rotation=0)
if legend:
plt.legend(loc="lower right", fontsize=14)
plt.figure(figsize=(8, 4))
plot_decision_boundary(tree_clf, X, y) ## decision boundary 그려줌 (위의 plot_decision_boundary 함수 활용)
plt.plot([2.45, 2.45], [0, 3], "k-", linewidth=2)
plt.plot([2.45, 7.5], [1.75, 1.75], "k--", linewidth=2)
plt.plot([4.95, 4.95], [0, 1.75], "k:", linewidth=2)
plt.plot([4.85, 4.85], [1.75, 3], "k:", linewidth=2)
plt.text(1.40, 1.0, "Depth=0", fontsize=15)
plt.text(3.2, 1.80, "Depth=1", fontsize=13)
plt.text(4.05, 0.5, "(Depth=2)", fontsize=11)
save_fig("decision_tree_decision_boundaries_plot")
plt.show()
```
# 클래스와 클래스 확률을 예측하기
```
tree_clf.predict_proba([[5, 1.5]])
tree_clf.predict([[5, 1.5]])
```
# 회귀 트리
```
# 2차식으로 만든 데이터셋 + 잡음
np.random.seed(42)
m = 200
X = np.random.rand(m, 1)
y = 4 * (X - 0.5) ** 2
y = y + np.random.randn(m, 1) / 10
from sklearn.tree import DecisionTreeRegressor
tree_reg1 = DecisionTreeRegressor(random_state=42, max_depth=2)
tree_reg2 = DecisionTreeRegressor(random_state=42, max_depth=3)
tree_reg1.fit(X, y)
tree_reg2.fit(X, y)
export_graphviz(
tree_reg1,
out_file=os.path.join(IMAGES_PATH, "regression_tree.dot"),
feature_names=["x1"],
rounded=True,
filled=True
)
Source.from_file(os.path.join(IMAGES_PATH, "regression_tree.dot"))
export_graphviz(
tree_reg2,
out_file=os.path.join(IMAGES_PATH, "regression_tree.dot"),
feature_names=["x1"],
rounded=True,
filled=True
)
Source.from_file(os.path.join(IMAGES_PATH, "regression_tree.dot"))
```
# Ensemble
```
# 파이썬 ≥3.5 필수
import sys
assert sys.version_info >= (3, 5)
# 사이킷런 ≥0.20 필수
import sklearn
assert sklearn.__version__ >= "0.20"
# 공통 모듈 임포트
import numpy as np
import os
import time
# 노트북 실행 결과를 동일하게 유지하기 위해
np.random.seed(42)
# 깔끔한 그래프 출력을 위해
%matplotlib inline
import matplotlib as mpl
import matplotlib.pyplot as plt
mpl.rc('axes', labelsize=14)
mpl.rc('xtick', labelsize=12)
mpl.rc('ytick', labelsize=12)
# 그림을 저장할 위치
PROJECT_ROOT_DIR = "."
CHAPTER_ID = "ensembles"
IMAGES_PATH = os.path.join(PROJECT_ROOT_DIR, "images", CHAPTER_ID)
os.makedirs(IMAGES_PATH, exist_ok=True)
def save_fig(fig_id, tight_layout=True, fig_extension="png", resolution=300):
path = os.path.join(IMAGES_PATH, fig_id + "." + fig_extension)
print("그림 저장:", fig_id)
if tight_layout:
plt.tight_layout()
plt.savefig(path, format=fig_extension, dpi=resolution)
from sklearn.model_selection import train_test_split
from sklearn.datasets import make_moons
X, y = make_moons(n_samples=500, noise=0.30, random_state=42)
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=42)
from sklearn.ensemble import BaggingClassifier
from sklearn.tree import DecisionTreeClassifier
bag_clf = BaggingClassifier(
DecisionTreeClassifier(random_state=42), n_estimators=500, #The number of base estimators in the ensemble.
max_samples=100, bootstrap=True, random_state=42)
bag_clf.fit(X_train, y_train)
y_pred = bag_clf.predict(X_test)
from sklearn.metrics import accuracy_score
print(accuracy_score(y_test, y_pred)) #bagging이 decision tree보다 test 성능이 좋다는 것을 볼 수 있다!
tree_clf = DecisionTreeClassifier(random_state=42)
tree_clf.fit(X_train, y_train)
y_pred_tree = tree_clf.predict(X_test)
print(accuracy_score(y_test, y_pred_tree)) #decision tree의 성능은 85%
from matplotlib.colors import ListedColormap
def plot_decision_boundary(clf, X, y, axes=[-1.5, 2.45, -1, 1.5], alpha=0.5, contour=True):
x1s = np.linspace(axes[0], axes[1], 100)
x2s = np.linspace(axes[2], axes[3], 100)
x1, x2 = np.meshgrid(x1s, x2s)
X_new = np.c_[x1.ravel(), x2.ravel()]
y_pred = clf.predict(X_new).reshape(x1.shape)
custom_cmap = ListedColormap(['#fafab0','#9898ff','#a0faa0'])
plt.contourf(x1, x2, y_pred, alpha=0.3, cmap=custom_cmap)
if contour:
custom_cmap2 = ListedColormap(['#7d7d58','#4c4c7f','#507d50'])
plt.contour(x1, x2, y_pred, cmap=custom_cmap2, alpha=0.8)
plt.plot(X[:, 0][y==0], X[:, 1][y==0], "yo", alpha=alpha)
plt.plot(X[:, 0][y==1], X[:, 1][y==1], "bs", alpha=alpha)
plt.axis(axes)
plt.xlabel(r"$x_1$", fontsize=18)
plt.ylabel(r"$x_2$", fontsize=18, rotation=0)
fix, axes = plt.subplots(ncols=2, figsize=(10,4), sharey=True) #bagging이 decision bound가 좀 더 smooth함
plt.sca(axes[0])
plot_decision_boundary(tree_clf, X, y)
plt.title("Decision Tree", fontsize=14)
plt.sca(axes[1])
plot_decision_boundary(bag_clf, X, y)
plt.title("Decision Trees with Bagging", fontsize=14)
plt.ylabel("")
save_fig("decision_tree_without_and_with_bagging_plot")
plt.show()
```
# Random Forest
```
start = time.time()
bag_clf = BaggingClassifier(
DecisionTreeClassifier(max_leaf_nodes=16, random_state=42),
n_estimators=500, max_samples=1.0, bootstrap=True, random_state=42)
bag_clf.fit(X_train, y_train)
print("time :", time.time() - start) # 현재시각 - 시작시간 = 실행 시간
y_pred = bag_clf.predict(X_test)
print(y_pred)
from sklearn.ensemble import RandomForestClassifier
start = time.time()
rnd_clf = RandomForestClassifier(n_estimators=500, max_leaf_nodes=16, random_state=42,max_features=2)
rnd_clf.fit(X_train, y_train)
print("time :", time.time() - start) # 현재시각 - 시작시간 = 실행 시간
y_pred_rf = rnd_clf.predict(X_test)
print(y_pred_rf)
print(accuracy_score(y_test, y_pred_rf))
plot_decision_boundary(rnd_clf, X, y)
np.sum(y_pred == y_pred_rf) / len(y_pred) #거의 예측이 동일합니다.
from sklearn.datasets import load_iris
iris = load_iris()
rnd_clf = RandomForestClassifier(n_estimators=500, random_state=42)
rnd_clf.fit(iris["data"], iris["target"])
for name, score in zip(iris["feature_names"], rnd_clf.feature_importances_):
print(name, score)
plt.figure(figsize=(6, 4))
for i in range(15):
tree_clf = DecisionTreeClassifier(max_leaf_nodes=16, random_state=42 + i)
indices_with_replacement = np.random.randint(0, len(X_train), len(X_train))
tree_clf.fit(X[indices_with_replacement], y[indices_with_replacement])
plot_decision_boundary(tree_clf, X, y, axes=[-1.5, 2.45, -1, 1.5], alpha=0.02, contour=False)
plt.show()
```
# AdaBoost
```
from sklearn.ensemble import AdaBoostClassifier
ada_clf = AdaBoostClassifier(
DecisionTreeClassifier(max_depth=1), n_estimators=200,
algorithm="SAMME.R", learning_rate=0.5, random_state=42)
ada_clf.fit(X_train, y_train)
plot_decision_boundary(ada_clf, X, y)
```
# Exercise
```
import pandas as pd
titanic = pd.read_csv('https://web.stanford.edu/class/archive' \
'/cs/cs109/cs109.1166/stuff/titanic.csv')
titanic
# drop string value
titanic.drop('Name', axis=1, inplace=True)
titanic.drop('Sex', axis=1, inplace=True)
from sklearn import tree
from graphviz import Source
# To-do
# Survived를 y에 저장, Survived를 제외한 나머지 column들을 X로 사용
# Decision Tree Classifier 학습 : model에 저장
X = titanic.drop('Survived', axis=1).values
y = titanic['Survived'].values
model = tree.DecisionTreeClassifier(max_depth=2)
model.fit(X, y)
graph = Source(tree.export_graphviz(model, out_file=None,
feature_names=titanic.columns[1:],
class_names=['Not survived', 'Survived'],
filled=True, rounded=True))
graph
```
| github_jupyter |
```
from IPython.lib.deepreload import reload
%load_ext autoreload
%autoreload 2
import re
import operator
from pathlib import Path
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import pydicom
from pymedphys_analysis.tpscompare import load_and_normalise_mephysto
from pymedphys_dicom.dicom import depth_dose, profile
ROOT_DIR = Path(r"S:\Physics\Monaco\Model vs Measurement Comparisons")
MONACO_DICOM_DIR = ROOT_DIR.joinpath(r"Beam Models\CCA Monaco Photon Monte Carlo")
DOSECHECK_DICOM_DIR = ROOT_DIR.joinpath(r"Beam Models\DoseCHECK")
MEASUREMENTS_DIR = ROOT_DIR.joinpath(r"Measurements\RCCC\Photons")
RESULTS = ROOT_DIR.joinpath(r"Results\RCCC\dosecheck\Monte Carlo")
MONACO_DOSE_DIR = MONACO_DICOM_DIR.joinpath("DICOM dose exports")
calibrated_doses_table = pd.read_csv(MEASUREMENTS_DIR.joinpath('AbsoluteDose.csv'), index_col=0)
calibrated_doses = calibrated_doses_table['d10 @ 90 SSD']
calibrated_doses
wedge_transmission_table = pd.read_csv(MEASUREMENTS_DIR.joinpath('WedgeTransmissionFactors.csv'), index_col=0)
data_column_name = wedge_transmission_table.columns[0]
wedge_transmissions = wedge_transmission_table[data_column_name]
wedge_transmissions
output_factors = pd.read_csv(MEASUREMENTS_DIR.joinpath('OutputFactors.csv'), index_col=0)
output_factors
keys = [
path.stem
for path in MONACO_DOSE_DIR.glob('*.dcm')
]
keys
regex_string = r'(\d\dMV(FFF)?) (\d\dx\d\d) ((\bOpen\b)|(\bWedge\b))'
def get_energy_field_block(key):
match = re.match(regex_string, key)
return match.group(1), match.group(3), match.group(4)
absolute_doses = {}
for key in keys:
energy, field, block = get_energy_field_block(key)
if block == 'Wedge':
wtf = wedge_transmissions[energy]
else:
wtf = 1
output_factor = output_factors[f'{field} {block}'][energy]
calibrated_dose = calibrated_doses[energy]
absolute_dose = calibrated_dose * output_factor * wtf
absolute_doses[key] = absolute_dose
absolute_doses
getter = operator.itemgetter('displacement', 'dose')
absolute_scans_per_field = load_and_normalise_mephysto(
MEASUREMENTS_DIR, r'(\d\dMV(FFF)? \d\dx\d\d Open)\.mcc', absolute_doses, 100)
new_keys = list(absolute_scans_per_field.keys())
new_keys
assert new_keys == keys
def load_dicom_files(directory, keys):
dicom_file_map = {
key: directory.joinpath(f'{key}.dcm')
for key in keys
}
dicom_dataset_map = {
key: pydicom.read_file(str(dicom_file_map[key]), force=True)
for key in keys
}
return dicom_dataset_map
monaco_dicom_dataset_map = load_dicom_files(MONACO_DOSE_DIR, keys)
dosecheck_dicom_dataset_map = load_dicom_files(DOSECHECK_DICOM_DIR, keys)
dicom_plan = pydicom.read_file(str(MONACO_DICOM_DIR.joinpath('plan.dcm')), force=True)
def plot_one_axis(ax, displacement, meas_dose, model_dose):
diff = 100 * (model_dose - meas_dose) / meas_dose
lines = []
lines += ax.plot(displacement, meas_dose, label='Measured Dose')
lines += ax.plot(displacement, model_dose, label='Model Dose')
ax.set_ylabel('Dose (Gy / 100 MU)')
x_bounds = [np.min(displacement), np.max(displacement)]
ax.set_xlim(x_bounds)
ax_twin = ax.twinx()
lines += ax_twin.plot(displacement, diff, color='C3', alpha=0.5, label=r'% Residuals [100 $\times$ (Model - Meas) / Meas]')
ax_twin.plot(x_bounds, [0, 0], '--', color='C3', lw=0.5)
ax_twin.set_ylabel(r'% Dose difference [100 $\times$ (Model - Meas) / Meas]')
labels = [l.get_label() for l in lines]
ax.legend(lines, labels, loc='lower left')
return ax_twin
def plot_tps_meas_diff(displacement, meas_dose, internal_tps_dose, external_tps_dose):
fig, ax = plt.subplots(1, 2, figsize=(16,6), sharey=True)
ax[1].yaxis.set_tick_params(which='both', labelbottom=True)
ax_twin = list()
ax_twin.append(plot_one_axis(ax[0], displacement, meas_dose, internal_tps_dose))
ax_twin.append(plot_one_axis(ax[1], displacement, meas_dose, external_tps_dose))
ax_twin[1].get_shared_y_axes().join(ax_twin[1], ax_twin[0])
ax_twin[1].set_ylim([-5, 5])
plt.tight_layout()
plt.subplots_adjust(wspace=0.4, top=0.86)
return fig, ax
def plot_pdd_diff(key, dicom_plan):
depth, meas_dose = getter(absolute_scans_per_field[key]['depth_dose'])
internal_tps_dose = depth_dose(depth, monaco_dicom_dataset_map[key], dicom_plan) / 10
external_tps_dose = depth_dose(depth, dosecheck_dicom_dataset_map[key], dicom_plan) / 10
fig, ax = plot_tps_meas_diff(depth, meas_dose, internal_tps_dose, external_tps_dose)
fig.suptitle(f'Depth Dose Comparisons | {key}', fontsize="x-large")
ax[0].set_title("Monaco Monte Carlo")
ax[1].set_title("DoseCHECK")
for key in keys:
plot_pdd_diff(key, dicom_plan)
filename = RESULTS.joinpath(f'{key}_pdd.png')
plt.savefig(filename)
plt.show()
def plot_profile_diff(key, depth, direction):
displacement, meas_dose = getter(absolute_scans_per_field[key]['profiles'][depth][direction])
internal_tps_dose = profile(displacement, depth, direction, monaco_dicom_dataset_map[key], dicom_plan) / 10
external_tps_dose = profile(displacement, depth, direction, dosecheck_dicom_dataset_map[key], dicom_plan) / 10
fig, ax = plot_tps_meas_diff(displacement, meas_dose, internal_tps_dose, external_tps_dose)
fig.suptitle(f'{direction.capitalize()} Profile Comparisons | {key} | Depth: {depth} mm', fontsize="x-large")
ax[0].set_title("Monaco Monte Carlo")
ax[1].set_title("DoseCHECK")
for key in keys:
depths = absolute_scans_per_field[key]['profiles'].keys()
for depth in depths:
for direction in ['inplane', 'crossplane']:
plot_profile_diff(key, depth, direction)
filename = RESULTS.joinpath(f'{key}_profile_{depth}mm_{direction}.png')
plt.savefig(filename)
plt.show()
```
| github_jupyter |
# Fine-tuning a Pretrained Network for Style Recognition
这个讲了从 caffe 的 model zoo 拿到别人训练好了的 model,如何把它 tune 得更适合自己的情况。
In this example, we'll explore a common approach that is particularly useful in real-world applications: take a pre-trained Caffe network and fine-tune the parameters on your custom data.
The upside of such approach is that, since pre-trained networks are learned on a large set of images, the intermediate layers capture the "semantics" of the general visual appearance. Think of it as a very powerful feature that you can treat as a black box. On top of that, only a few layers will be needed to obtain a very good performance of the data.
First, we will need to prepare the data. This involves the following parts:
(1) Get the ImageNet ilsvrc pretrained model with the provided shell scripts.
(2) Download a subset of the overall Flickr style dataset for this demo.
(3) Compile the downloaded Flickr dataset into a database that Caffe can then consume.
```
import os
os.chdir('..')
print os.path.realpath(os.getcwd())
import sys
sys.path.insert(0, './python')
import caffe
import numpy as np
from pylab import *
%matplotlib inline
# This downloads the ilsvrc auxiliary data (mean file, etc),
# and a subset of 2000 images for the style recognition task.
#!data/ilsvrc12/get_ilsvrc_aux.sh
# 还是用我们已经下载好了的数据。cp 过来就好。
!data/ilsvrc12/get_ilsvrc_aux_cvrs.sh
print "done get files.\n"
# 提前解压这个文件,用【迅雷】把图片下载了直接拷贝过去。
# examples/finetune_flickr_style/flickr_style.csv.gz
# 如果图片已经存在,这个脚本不会再下载(还得通过sha1 校验)。
!scripts/download_model_binary.py models/bvlc_reference_caffenet
print "done get model binary.\n"
!python examples/finetune_flickr_style/assemble_data.py \
--workers=-1 --images=2000 --seed=1701 --label=5
print "done assemble.\n"
```
Let's show what is the difference between the fine-tuning network and the original caffe model.
```
!diff models/bvlc_reference_caffenet/train_val.prototxt models/finetune_flickr_style/train_val.prototxt
```
For your record, if you want to train the network in pure C++ tools, here is the command:
<code>
build/tools/caffe train \
-solver models/finetune_flickr_style/solver.prototxt \
-weights models/bvlc_reference_caffenet/bvlc_reference_caffenet.caffemodel \
-gpu 0
</code>
However, we will train using Python in this example.
```
niter = 200
# losses will also be stored in the log
train_loss = np.zeros(niter)
scratch_train_loss = np.zeros(niter)
caffe.set_device(0)
caffe.set_mode_gpu()
# We create a solver that fine-tunes from a previously trained network.
solver = caffe.SGDSolver('models/finetune_flickr_style/solver.prototxt')
# 这里把之前的模型载入了。
solver.net.copy_from('models/bvlc_reference_caffenet/bvlc_reference_caffenet.caffemodel')
# 对比这个 scratch_solver,
# 没有载入新的模型。For reference, we also create a solver that does no finetuning.
scratch_solver = caffe.SGDSolver('models/finetune_flickr_style/solver.prototxt')
# We run the solver for niter times, and record the training loss.
for it in range(niter):
solver.step(1) # SGD by Caffe
scratch_solver.step(1)
# store the train loss
train_loss[it] = solver.net.blobs['loss'].data
scratch_train_loss[it] = scratch_solver.net.blobs['loss'].data
if it % 10 == 0:
print 'iter %d, finetune_loss=%f, scratch_loss=%f' % (it, train_loss[it], scratch_train_loss[it])
```
Let's look at the training loss produced by the two training procedures respectively.
train_loss 和 scratch_train_loss 都是 ndarray, 通过 `np.zeros(num)` 定义的。
```
print type(train_loss)
plot(np.vstack([train_loss, scratch_train_loss]).T)
```
Notice how the fine-tuning procedure produces a more smooth loss function change, and ends up at a better loss. A closer look at small values, clipping to avoid showing too large loss during training:
```
plot(np.vstack([train_loss, scratch_train_loss]).clip(0, 4).T)
```
Let's take a look at the testing accuracy after running 200 iterations. Note that we are running a classification task of 5 classes, thus a chance accuracy is 20%. As we will reasonably expect, the finetuning result will be much better than the one from training from scratch. Let's see.
```
test_iters = 10
accuracy = 0
scratch_accuracy = 0
for it in arange(test_iters):
solver.test_nets[0].forward()
accuracy += solver.test_nets[0].blobs['accuracy'].data
scratch_solver.test_nets[0].forward()
scratch_accuracy += scratch_solver.test_nets[0].blobs['accuracy'].data
accuracy /= test_iters
scratch_accuracy /= test_iters
print 'Accuracy for fine-tuning:', accuracy
print 'Accuracy for training from scratch:', scratch_accuracy
```
Huzzah! So we did finetuning and it is awesome. Let's take a look at what kind of results we are able to get with a longer, more complete run of the style recognition dataset. Note: the below URL might be occassionally down because it is run on a research machine.
http://demo.vislab.berkeleyvision.org/
| github_jupyter |
### Defining a cohort
A cohort is a table whose rows correspond to unique combinations of `person_id` and `index_date` where each combination is mapped to a unique `row_id`.
For downstream feature extraction and modeling, the cohort table should also contain additional columns for labeling for outcomes and group categories.
Here, we will call a set of pre-defined set of transformations to define a cohort of hospital admissions and extract relevant labels. For details of how this cohort is defined, refer to source code. In practice, a cohort can be defined arbitrarily, as long as meets the specification described above and is stored in a table in the database.
```
import os
from prediction_utils.cohorts.admissions.cohort import (
BQAdmissionRollupCohort, BQAdmissionOutcomeCohort, BQFilterInpatientCohort
)
from prediction_utils.util import patient_split
# Configuration for the extraction
config_dict = {
'gcloud_project': 'som-nero-phi-nigam-starr',
'dataset_project': 'som-rit-phi-starr-prod',
'rs_dataset_project': 'som-nero-phi-nigam-starr',
'dataset': 'starr_omop_cdm5_deid_1pcent_lite_latest',
'rs_dataset': 'temp_dataset',
'cohort_name': 'vignette_cohort',
'cohort_name_labeled': 'vignette_cohort_labeled',
'cohort_name_filtered': 'vignette_cohort_filtered',
'has_birth_datetime': True
}
cohort = BQAdmissionRollupCohort(**config_dict)
# Create the cohort table
cohort.create_cohort_table()
# Let's inspect the cohort
cohort_df = cohort.db.read_sql_query(
query="SELECT * FROM {rs_dataset_project}.{rs_dataset}.{cohort_name}".format(**config_dict)
)
cohort_df.head()
# Now let's add some labels
cohort_labeled = BQAdmissionOutcomeCohort(**config_dict)
cohort_labeled.create_cohort_table()
cohort_df_labeled = cohort_labeled.db.read_sql_query(
query="SELECT * FROM {rs_dataset_project}.{rs_dataset}.{cohort_name_labeled}".format(**config_dict)
)
cohort_df_labeled.head()
# Now let's filter down to one prediction per patient and add a row_id column called `prediction_id`
cohort_filtered = BQFilterInpatientCohort(**config_dict)
cohort_filtered.create_cohort_table()
# Get the filtered cohort
cohort_df_filtered = cohort_filtered.db.read_sql_query(
query = """
SELECT *
FROM {rs_dataset_project}.{rs_dataset}.{cohort_name_filtered}
""".format(**config_dict)
).set_index('prediction_id').reset_index()
cohort_df_filtered.head()
# Partition the dataset into folds for later
cohort_df_final = patient_split(cohort_df_filtered)
cohort_df_final.head()
# Write the result to disk
cohort_path = '/share/pi/nigam/projects/prediction_utils/scratch/cohort'
os.makedirs(cohort_path, exist_ok=True)
cohort_df_final.to_parquet(
os.path.join(cohort_path, "cohort.parquet"), engine="pyarrow", index=False
)
```
| github_jupyter |
```
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import sklearn
import seaborn as sns
?pd.read_csv
feature_train=pd.read_csv('Molecular_Descriptor.csv',index_col='SMILES')
label_train=pd.read_csv('ERα_activity.csv',index_col='SMILES')
del label_train['IC50_nM']
feature_train.info()
?pd.concat
#data=pd.concat((feature_train,label_train['pIC50']),axis=1)
#data.dtypes.value_counts()
for s in data.columns:
print(s,'的特征分布:')
print(data[s].value_counts())
sns.distplot(data['pIC50'])
corrlation['pIC50'].isna().sum()
#data['n10Ring'].value_counts()
data['hmin'].nunique()
#删除数据只有一种的特征
for s in feature_train.columns:
if feature_train[s].nunique()==1:
del feature_train[s]
feature_train.shape
corrlation=data.corr()
corrlation['pIC50'].sort_values()
cor_1=pd.DataFrame(corrlation['pIC50'])
cor_1=abs(cor_1)
#应用随机森林来选择特征
from sklearn.ensemble import RandomForestRegressor
from sklearn.model_selection import train_test_split
forest = RandomForestRegressor(n_estimators=500, random_state=0, max_features=100, n_jobs=2)
forest.fit(feature_train,label_train)
#sorted(forest.feature_importances_, reverse=True)
forest.feature_importances_[2]
feature_impor=pd.DataFrame(forest.feature_importances_)
feature_impor['rank']=range(0,504)
feature_impor.columns=['s','rank']
feature_impor
feature_impor=feature_impor.sort_values(by='s',ascending=False)
choose_feature=[]
for i in range(504):
if feature_impor.iloc[i,0]>=float(feature_impor['s'].quantile(0.89)):
choose_feature.append(feature_train.columns[feature_impor.iloc[i,1]])
len(choose_feature)
#然后通过相关分析提出自相关的变量
cor=feature_train[choose_feature].corr()
f , ax = plt.subplots(figsize = (9,9))
cor
for i in range(55):
for j in range(55):
if i !=j:
if cor.iloc[i,j]>0.5 and cor.index[i] in choose_feature and cor.columns[j] in choose_feature:
if len(choose_feature)==20:
break;
choose_feature.remove(cor.columns[j])
choose_feature
final_feature_train=feature_train[choose_feature]
final_feature_train.shape
final_feature_train.to_csv('final.csv')
from sklearn.model_selection import cross_val_score
from sklearn.metrics import mean_squared_error,make_scorer,mean_absolute_error
from sklearn.linear_model import LinearRegression
from sklearn.tree import DecisionTreeRegressor
from sklearn.ensemble import RandomForestRegressor
from sklearn.ensemble import GradientBoostingRegressor
from sklearn.neural_network import MLPRegressor
from xgboost.sklearn import XGBRegressor
from lightgbm.sklearn import LGBMRegressor
models=[LinearRegression(),DecisionTreeRegressor(),RandomForestRegressor(),
GradientBoostingRegressor(),
MLPRegressor(hidden_layer_sizes=(50),solver='lbfgs',max_iter=100),
XGBRegressor(n_estimator=100,objective='reg:squarederror'),
LGBMRegressor(n_estimator=100)
]
result=dict()
for model in models:
model_name = str(model).split('(')[0]
scores = cross_val_score(model, X=final_feature_train, y=label_train, verbose=0, cv = 5,scoring=make_scorer(mean_squared_error))
result[model_name] = scores
print(model_name + ' is finished')
#表格表示
result = pd.DataFrame(result)
result.index = ['cv' + str(x) for x in range(1, 6)]
result
result.mean()
feature_test=pd.read_csv('Molecular_Descriptor_test.csv',index_col='SMILES')
label_test=pd.read_csv('ERα_activity_test.csv',index_col='SMILES')
```
| github_jupyter |
wat.011 https://easy.dans.knaw.nl/ui/datasets/id/easy-dataset:64145/tab/2
format: asc
download 2000 phosphorus and nitrogen total load and retention data see readme.pdf for more info
Import libraries
```
# Libraries for downloading data from remote server (may be ftp)
import requests
from urllib.request import urlopen
from contextlib import closing
import shutil
# Library for uploading/downloading data to/from S3
import boto3
# Libraries for handling data
import rasterio as rio
import numpy as np
# from netCDF4 import Dataset
# import pandas as pd
# import scipy
# Libraries for various helper functions
# from datetime import datetime
import os
import threading
import sys
from glob import glob
```
s3 tools
```
s3_upload = boto3.client("s3")
s3_download = boto3.resource("s3")
s3_bucket = "wri-public-data"
s3_folder = "resourcewatch/raster/wat_011_global_riverine_N_P/"
s3_file1 = "wat_011_global_riverine_N_Load_2000.asc"
s3_file2 = "wat_011_global_riverine_P_Load_2000.asc"
s3_file3 = "wat_011_global_riverine_N_Retention_2000.asc"
s3_file4 = "wat_011_global_riverine_P_Retention_2000.asc"
s3_key_orig1 = s3_folder + s3_file1
s3_key_edit1 = s3_key_orig1[0:-4] + "_edit.tif"
s3_key_orig2 = s3_folder + s3_file2
s3_key_edit2 = s3_key_orig2[0:-4] + "_edit.tif"
s3_key_orig3 = s3_folder + s3_file3
s3_key_edit3 = s3_key_orig3[0:-4] + "_edit.tif"
s3_key_orig4 = s3_folder + s3_file4
s3_key_edit4 = s3_key_orig4[0:-4] + "_edit.tif"
s3_files_to_merge = [s3_key_edit1, s3_key_edit2, s3_key_edit3,s3_key_edit4]
band_ids = ["N_Load","P_Load","N_Retention", "P_Retention"]
s3_key_merge = s3_folder + "Wat_011_global_riverine_Merge_2000.tif"
class ProgressPercentage(object):
def __init__(self, filename):
self._filename = filename
self._size = float(os.path.getsize(filename))
self._seen_so_far = 0
self._lock = threading.Lock()
def __call__(self, bytes_amount):
# To simplify we'll assume this is hooked up
# to a single filename.
with self._lock:
self._seen_so_far += bytes_amount
percentage = (self._seen_so_far / self._size) * 100
sys.stdout.write("\r%s %s / %s (%.2f%%)"%(
self._filename, self._seen_so_far, self._size,
percentage))
sys.stdout.flush()
```
Define local file locations
```
local_folder = "/Users/Max81007/Desktop/Python/Resource_Watch/Raster/wat.011/"
file_name1 = "Nload_2000.asc"
file_name2 = "Pload_2000.asc"
file_name3 = "retention_N_2000.asc"
file_name4 = "retention_P_2000.asc"
local_orig1 = local_folder + file_name1
local_orig2 = local_folder + file_name2
local_orig3 = local_folder + file_name3
local_orig4 = local_folder + file_name4
orig_extension_length = 4 #4 for each char in .tif
local_edit1 = local_orig1[:-orig_extension_length] + "_edit.tif"
local_edit2 = local_orig2[:-orig_extension_length] + "_edit.tif"
local_edit3 = local_orig3[:-orig_extension_length] + "_edit.tif"
local_edit4 = local_orig4[:-orig_extension_length] + "_edit.tif"
merge_files = [local_edit1, local_edit2, local_edit3, local_edit4]
tmp_merge = local_folder + "Foo_039_Fertilizer_TotalConsumption_Merge.tif"
```
Use rasterio to reproject and compress
```
files = [local_orig1, local_orig2, local_orig3, local_orig4]
for file in files:
with rio.open(file, 'r') as src:
profile = src.profile
print(profile)
# Note - this is the core of Vizz's netcdf2tif function
def convert_asc_to_tif(orig_name, edit_name):
with rio.open(orig_name, 'r') as src:
# This assumes data is readable by rasterio
# May need to open instead with netcdf4.Dataset, for example
data = src.read()[0]
rows = data.shape[0]
columns = data.shape[1]
print(rows)
print(columns)
# Latitude bounds
south_lat = -90
north_lat = 90
# Longitude bounds
west_lon = -180
east_lon = 180
transform = rio.transform.from_bounds(west_lon, south_lat, east_lon, north_lat, columns, rows)
# Profile
no_data_val = -9999.0
target_projection = 'EPSG:4326'
target_data_type = np.float64
profile = {
'driver':'GTiff',
'height':rows,
'width':columns,
'count':1,
'dtype':target_data_type,
'crs':target_projection,
'transform':transform,
'compress':'lzw',
'nodata': no_data_val
}
with rio.open(edit_name, "w", **profile) as dst:
dst.write(data.astype(profile["dtype"]), 1)
convert_asc_to_tif(local_orig1, local_edit1)
convert_asc_to_tif(local_orig2, local_edit2)
convert_asc_to_tif(local_orig3, local_edit3)
convert_asc_to_tif(local_orig4, local_edit4)
with rio.open(merge_files[0]) as src:
kwargs = src.profile
kwargs.update(
count=len(merge_files)
)
with rio.open(tmp_merge, 'w', **kwargs) as dst:
for idx, file in enumerate(merge_files):
print(idx)
with rio.open(file) as src:
band = idx+1
windows = src.block_windows()
for win_id, window in windows:
src_data = src.read(1, window=window)
dst.write_band(band, src_data, window=window)
files = [tmp_merge]
for file in files:
with rio.open(file, 'r') as src:
profile = src.profile
print(profile)
```
Upload orig and edit files to s3
```
# Original
s3_upload.upload_file(local_orig1, s3_bucket, s3_key_orig1,
Callback=ProgressPercentage(local_orig1))
s3_upload.upload_file(local_orig2, s3_bucket, s3_key_orig2,
Callback=ProgressPercentage(local_orig2))
s3_upload.upload_file(local_orig3, s3_bucket, s3_key_orig3,
Callback=ProgressPercentage(local_orig3))
s3_upload.upload_file(local_orig4, s3_bucket, s3_key_orig4,
Callback=ProgressPercentage(local_orig4))
# Edit
s3_upload.upload_file(local_edit1, s3_bucket, s3_key_edit1,
Callback=ProgressPercentage(local_edit1))
s3_upload.upload_file(local_edit2, s3_bucket, s3_key_edit2,
Callback=ProgressPercentage(local_edit2))
s3_upload.upload_file(local_edit3, s3_bucket, s3_key_edit3,
Callback=ProgressPercentage(local_edit3))
s3_upload.upload_file(local_edit4, s3_bucket, s3_key_edit4,
Callback=ProgressPercentage(local_edit4))
s3_upload.upload_file(tmp_merge, s3_bucket, s3_key_merge,
Callback=ProgressPercentage(tmp_merge))
os.environ["Zs3_key"] = "s3://wri-public-data/" + s3_key_merge
os.environ["Zs3_key_inspect"] = "wri-public-data/" + s3_key_merge
os.environ["Zgs_key"] = "gs://resource-watch-public/" + s3_key_merge
!echo %Zs3_key_inspect%
!aws s3 ls %Zs3_key_inspect%
!gsutil cp %Zs3_key% %Zgs_key%
os.environ["asset_id"] = "users/resourcewatch/wat_011_global_riverine_N_P_2000"
!earthengine upload image --asset_id=%asset_id% %Zgs_key%
os.environ["band_names"] = str(band_ids)
!earthengine asset set -p band_names="%band_names%" %asset_id%
```
| github_jupyter |
```
import poppy
import matplotlib
import matplotlib.pyplot as plt
import numpy as np
import time
poppy.__version__
%matplotlib inline
matplotlib.rcParams['image.origin'] = 'lower'
D = 2.
wavelen = 1e-6
ovsamp = 8
# No FPM (control case)
fftcoron_noFPM_osys = poppy.OpticalSystem(oversample=ovsamp)
fftcoron_noFPM_osys.add_pupil( poppy.CircularAperture(radius=D/2) )
fftcoron_noFPM_osys.add_image()
fftcoron_noFPM_osys.add_pupil( poppy.CircularAperture(radius=0.9*D/2) )
fftcoron_noFPM_osys.add_detector( pixelscale=0.025, fov_arcsec=4. )
# Classical Lyot FPM, spot radius ~ 4 lam/D
fftcoron_spotFPM_osys = poppy.OpticalSystem(oversample=ovsamp)
fftcoron_spotFPM_osys.add_pupil( poppy.CircularAperture(radius=1.) )
fftcoron_spotFPM_osys.add_image( poppy.CircularOcculter(radius=0.4) )
fftcoron_spotFPM_osys.add_pupil( poppy.CircularAperture(radius=0.9*D/2) )
fftcoron_spotFPM_osys.add_detector( pixelscale=0.05, fov_arcsec=4. )
# Annular diaphragm FPM, inner radius ~ 4 lam/D, outer rad ~ 16 lam/D
fftcoron_annFPM_osys = poppy.OpticalSystem(oversample=ovsamp)
fftcoron_annFPM_osys.add_pupil( poppy.CircularAperture(radius=D/2) )
spot = poppy.CircularOcculter( radius=0.4 )
diaphragm = poppy.InverseTransmission( poppy.CircularOcculter( radius=1.6 ) )
annFPM = poppy.CompoundAnalyticOptic( opticslist = [diaphragm, spot] )
fftcoron_annFPM_osys.add_image( annFPM )
fftcoron_annFPM_osys.add_pupil( poppy.CircularAperture(radius=0.9*D/2) )
fftcoron_annFPM_osys.add_detector( pixelscale=0.05, fov_arcsec=4. )
# Re-cast as MFT coronagraph with annular diaphragm FPM
matrixFTcoron_annFPM_osys = poppy.MatrixFTCoronagraph( fftcoron_annFPM_osys, occulter_box=diaphragm.uninverted_optic.radius_inner )
noFPM_fft_psf = fftcoron_noFPM_osys.calc_psf(wavelen, display_intermediates=False)
spotFPM_fft_psf = fftcoron_spotFPM_osys.calc_psf(wavelen, display_intermediates=False)
t0_fft = time.time()
annFPM_fft_psf, annFPM_fft_interm = fftcoron_annFPM_osys.calc_psf(wavelen, display_intermediates=True,\
return_intermediates=True)
t1_fft = time.time()
plt.tight_layout()
t0_mft = time.time()
annFPM_mft_psf, annFPM_mft_interm = matrixFTcoron_annFPM_osys.calc_psf(wavelen, display_intermediates=True,\
return_intermediates=True)
t1_mft = time.time()
plt.tight_layout()
plt.figure(figsize=(14,5))
plt.subplot(121)
poppy.display_psf(noFPM_fft_psf, vmax=1e-4, title='On-axis PSF, FPM removed, FFT propagation')
plt.subplot(122)
poppy.display_psf(spotFPM_fft_psf, vmin=1e-10, vmax=1e-7, title='On-axis PSF, classical spot FPM, FFT propagation')
plt.figure(figsize=(16,3.5))
plt.subplots_adjust(left=0.10, right=0.95, bottom=0.02, top=0.98, wspace=0.2, hspace=None)
plt.subplot(131)
ax_fft, cbar_fft = poppy.display_psf(annFPM_fft_psf, vmin=1e-10, vmax=1e-7, title='Annular FPM Lyot coronagraph, FFT',
return_ax=True)
plt.subplot(132)
poppy.display_psf(annFPM_mft_psf, vmin=1e-10, vmax=1e-7, title='Annular FPM Lyot coronagraph, Matrix FT')
plt.subplot(133)
diff_vmin = np.min(annFPM_mft_psf[0].data - annFPM_fft_psf[0].data)
diff_vmax = np.max(annFPM_mft_psf[0].data - annFPM_fft_psf[0].data)
poppy.display_psf_difference(annFPM_mft_psf, annFPM_fft_psf, vmin=diff_vmin, vmax=diff_vmax, cmap='gist_heat')
plt.title('Difference (MatrixFT - FFT)')
import astropy.units as u
lamoD_asec = (wavelen*u.m/fftcoron_annFPM_osys.planes[0].pupil_diam * u.radian).to(u.arcsec)
print("System diffraction resolution element scale (lambda/D) "+ str(np.round(lamoD_asec,4)))
print("Array width in first focal plane, FFT: %d" % annFPM_fft_interm[1].amplitude.shape[0])
print("Array width in first focal plane, MatrixFT: %d" % annFPM_mft_interm[1].amplitude.shape[0])
print("Array width in Lyot plane, FFT: %d" % annFPM_fft_interm[2].amplitude.shape[0])
print("Array width in Lyot plane, MatrixFT: %d" % annFPM_mft_interm[2].amplitude.shape[0])
SoS_res = np.sum( (annFPM_mft_psf[0].data - annFPM_fft_psf[0].data)**2 )
print("Maximum absolute value of difference between MatrixFT and FFT PSF intensity arrays: %g" % \
np.max(np.abs(annFPM_mft_psf[0].data - annFPM_fft_psf[0].data)))
print("Root mean square of difference between MatrixFT and FFT PSF intensity arrays: %g" % \
(np.sqrt(SoS_res/annFPM_mft_psf[0].data.shape[0]**2)))
print("Sum-of-squares difference between MatrixFT and FFT PSF intensity arrays: %g" % SoS_res)
print("Elapsed time, FFT: %.1f s" % (t1_fft-t0_fft))
print("Elapsed time, Matrix FT: %.1f s" % (t1_mft-t0_mft))
```
| github_jupyter |

The Ames Housing dataset was compiled by Dean De Cock for use in data science education. It's an incredible alternative for data scientists looking for a modernized and expanded version of the often cited Boston Housing dataset.
## Import required libraries
```
import pandas as pd
from supervised.automl import AutoML
from supervised.preprocessing.eda import EDA
```
## Load train and test dataset
```
train_df = pd.read_csv("data/house_price_train.csv")
test_df = pd.read_csv("data/house_price_test.csv")
train_df.head()
print("\nThe train data size after dropping Id feature is : {} ".format(train_df.shape))
print("The test data size after dropping Id feature is : {} ".format(test_df.shape))
X_train = train_df.drop(['SalePrice'],axis=1)
y_train = train_df['SalePrice']
```
## Basic EDA and model training
#### Want to do basic EDA?
- AutoML contains automated Exploratory data analysis for input data, by default it performs and saved the automated eda report for the given training dataset. For more details check [here](https://supervised.mljar.com/features/eda/)
- The model training is done using `AutoML.fit()` method, you can control the and select the algorithms to be used and the training time etc, please refer [docs](https://supervised.mljar.com/features/automl/) here for more details.
#### Want to do automated feature engineering?
mljar provides [golden features](https://supervised.mljar.com/features/automl/#golden_features)
- Golden Features are new features constructed from original data which have great predictive power. Set the `golden_features` parameter to `True` and see if work.
#### What to do cross-validation?
specify your cross-validation strategy in `validation_stategy` parameter in AutoML.
#### What to do ML Explainability?
AutoML provides feature importances and SHAP value explanations for tree based models. This is controlled by `explain_level` parameter in AutoML. Refer [docs](https://supervised.mljar.com/features/explain/) for more information.
```
a = AutoML(algorithms=['Xgboost'],total_time_limit=30,
explain_level=2,golden_features=True,
validation_strategy={
"validation_type": "kfold",
"k_folds": 3,
"shuffle": False,
"stratify": True,
})
a.fit(X_train,y_train)
```
## Predict on test
```
predictions = a.predict(test_df)
submission = pd.read_csv("data/sample_submission.csv")
submission['SalePrice'] = predictions
submission.head()
```
| github_jupyter |
## Instantiate the Pipeline
```
%%time
import importlib
import cndlib.pipeline
importlib.reload(cndlib.pipeline)
cnd = cndlib.pipeline.CND()
print([name for name in cnd.nlp.pipe_names])
```
## Instantiate the Dataset
```
%%time
import importlib
from IPython.display import clear_output
import cndlib.cndobjects
importlib.reload(cndlib.cndobjects)
dirpath = r'C:\\Users\\spa1e17\\OneDrive - University of Southampton\\hostile-narrative-analysis\\dataset'
orators = cndlib.cndobjects.Dataset(cnd, dirpath)
clear_output(wait=True)
display(orators.summarise())
```
## Create .csv Files of Sentences of Each Orator for Annotation
In this experiment we test sentiment analysis to detect the ingroup elevation and outgroup other phrases. For this experiment, each sentence of Bush and bin Laden’s datasets have been annotated as either ingroup elevation or outgroup othering. Accordingly, annotation was based on two criteria. Firstly, the sentence must contain reference to a named entity, whether explicitly or by either noun phrase or pronoun. Secondly, the reference must be associated with a term in the sentence which either elevates or others the reference entity. For example, with an implicit reference to al Qaeda, the following two sentences from Bush are annotated as othering, “These terrorists don't represent peace”, “They represent evil and war”. Equally, from both datasets the clauses, “God bless America” or “Allah blessed be upon him” are annotated as elevation.
An extra annotation was also added for hostile and anti-sematic sentences. Hostile sentences are those containing a threat of violence. For example from bin Laden, “And whoever has killed our civilians, then we have the right to kill theirs”, or from Bush, “We are sending a signal to the world as we speak that if you harbor a terrorist, there will be a price to pay.”. Some hostile sentences are veiled threat, but in the context of the narrative are determined to be threatening. Bin Laden’s explicit outgroup are Jews and Israel, as such, may of his sentences have been annotated as Ant-Sematic. The International Holocaust Remembrance Alliance (IHRC) definition of anti-Semitism was used as a guide for these annotations . An example of one annotation is, “Behind them stand the Jews, who control your policies, media and economy”. In this sentence bin Laden suggests Jewish people control the wealthy Americans, which conforms with the IHRC’s anti-Semitic characterisation of Jewish control of “Jews controlling the media, economy, government or other societal institutions”.
```
%time
import os
import csv
import pandas as pd
docs = {"bush" : {"name" : "George Bush", "filename" : "bush_sentences_gold.txt", "sentences" : []},
"laden" : {"name" : "Osama bin Laden", "filename" : "bush_sentences_gold.txt", "sentences" : []},
"king" : {"name" : "Martin Luther King", "sentences" : []},
"hitler" : {"name" : "Adolf Hitler", "sentences" : []}}
dirpath = os.getcwd()
for orator, texts in orators.orators_dict.items():
for text in texts.texts:
for sentence in text.doc.sents:
sent = {"function" : "", "hostile" : "", "text" : sentence.text.replace('\n', ' ').strip()}
docs[orator]['sentences'].append(sent)
filename = f"{orator}_sentences.csv"
df = pd.DataFrame(docs[orator]['sentences'])
filepath = os.path.join(dirpath, filename)
df.to_csv(filepath, sep=',',index=False)
pd.DataFrame([{"Orator" : doc['name'],
"Number of Sentences" : len(doc['sentences'])}
for doc in docs.values()
])
```
## Import Annotation Results for Scoring by Sentiment Analysis APIs
```
import os
import csv
import pandas as pd
from cndlib.visuals import display_side_by_side
docs = {
"bush" : {"name" : "George Bush", "filename" : "bush_sentences_gold.txt", "sentences" : None},
"laden" : {"name" : "Osama bin Laden", "filename" : "laden_sentences_gold.txt", "sentences" : None}
}
for orator in docs.values():
filename = filename = os.path.join(os.getcwd(), orator['filename'])
with open(filename, newline = "") as fp:
data = csv.DictReader(fp, delimiter = '\t')
orator['sentences'] = [row for row in data]
df = pd.DataFrame()
for orator in docs.values():
summary = []
summary.append({"Number of Sentences" : len(orator['sentences'])})
df2 = pd.DataFrame(orator['sentences'])
summary.extend([{f"{k.title()} Sentences" : str(v) for k, v in df2['function'].value_counts().items() if k}])
summary.extend([{f"{k.title()} Sentences" : str(v) for k, v in df2['hostile'].value_counts().items() if k}])
df = pd.concat([df, pd.DataFrame({k:v for x in summary for k,v in x.items()}, index = [orator['name']])])
display_side_by_side([df.fillna('')], ["Elevation and Othering Annotation Results"])
```
### Get Google API Results
https://cloud.google.com/natural-language/docs/basics#:~:text=score%20of%200.8%20.-,Interpreting%20sentiment%20analysis%20values,the%20length%20of%20the%20document
```
%%time
import os
from tqdm import tqdm
import pickle
# instantiate Google Sentiment Analysis
from google.cloud import language_v1
client = language_v1.LanguageServiceClient()
# iterate through each orator() object
for orator in docs.values():
# iterate over each Text() of the orator() object
for sent_obj in tqdm(orator['sentences'], total = len(orator['sentences']), desc = orator['name']):
text = sent_obj['text']
# document = language_v1.Document(content=text, type_=language_v1.Document.Type.PLAIN_TEXT)
# sentiment = client.analyze_sentiment(request={'document': document}).document_sentiment
# sent_obj['google sentiment score'] = sentiment.score
# sent_obj['google sentiment magnitude'] = sentiment.magnitude
display(pd.DataFrame([obj for obj in docs['laden']['sentences']]))
# google_document_results = document_results
```
### Get IBM Watson API Results
```
%%time
import json
from ibm_watson import NaturalLanguageUnderstandingV1
from ibm_cloud_sdk_core.authenticators import IAMAuthenticator
from ibm_watson.natural_language_understanding_v1 import Features, SentimentOptions, EmotionOptions
apikey = 'D3ptPkoLkoQNJvIav-reiA5137cr3m8Y1f-mhX1bLile'
url = 'https://api.eu-gb.natural-language-understanding.watson.cloud.ibm.com/instances/204e6ba7-952c-41ae-99e9-fe4e8208bfde'
authenticator = IAMAuthenticator(apikey)
service = NaturalLanguageUnderstandingV1(version='2019-07-12', authenticator=authenticator)
service.set_service_url(url)
for orator in docs.values():
for sent_obj in tqdm(orator['sentences'], total = len(orator['sentences']), desc = orator['name']):
text = sent_obj['text']
analytics = service.analyze(text=text, features=Features(
sentiment=SentimentOptions(),
emotion=EmotionOptions()),
language = "en").get_result()
sent_obj['watson sentiment'] = analytics['sentiment']['document']['score']
emotion = analytics['emotion']['document']['emotion']
sent_obj.update({f"Watson {k}" : v for k, v in emotion.items()})
```
### Get Microsoft Azure API Results
https://docs.microsoft.com/en-us/azure/cognitive-services/text-analytics/quickstarts/client-libraries-rest-api?tabs=version-3-1&pivots=programming-language-python#sentiment-analysis
```
import json
from azure.core.credentials import AzureKeyCredential
from azure.ai.textanalytics import TextAnalyticsClient
filename = "C:\\Users\\spa1e17\\OneDrive - University of Southampton\\CNDWip\\APIKeys\\AzureKeys.json"
with open(filename, 'r') as fp:
keys = json.load(fp)
apikey = keys['KEY 1']
endpoint = keys['Endpoint']
credential = AzureKeyCredential(apikey)
endpoint=endpoint
text_analytics_client = TextAnalyticsClient(endpoint, credential)
for orator in docs.values():
for sent_obj in tqdm(orator['sentences'], total = len(orator['sentences']), desc = orator['name']):
text = [sent_obj['text']]
response = text_analytics_client.analyze_sentiment(text, language="en")
label = response[0].sentiment
score = response[0].confidence_scores[label]
if label == "negative":
score = score*-1
sent_obj['azure sentiment'] = score
```
## Get TextBlob API Results
https://textblob.readthedocs.io/en/dev/quickstart.html#sentiment-analysis
```
from textblob import TextBlob
for orator in docs.values():
for sent_obj in tqdm(orator['sentences'], total = len(orator['sentences']), desc = orator['name']):
text = sent_obj['text']
sent_obj['textblob sentiment'] = TextBlob(text).sentiment[0]
```
## Write Results to Disc
```
import os
from cndlib.cndutils import dump_jsonl
import json
print(type(docs))
filename = os.path.join(os.getcwd(), "gold_results.json")
with open(filename, 'w') as file:
file.write(json.dumps(docs))
df = pd.DataFrame(doc for doc in docs['laden']['sentences'])
labels = ['text', 'function', 'hostile'] + [label for label in df.keys() if 'sentiment' in label and 'magnitute' not in label]
display(df[labels][df.function.eq('elevation')])
def get_function(orator, entity):
"""
function to get the grouping of an entity from the orator's groupings
"""
if entity in docs[orator]['text']['groups']['ingroup']:
return "ingroup"
if entity in docs[orator]['text']['groups']['outgroup']:
return "outgroup"
return "not found"
def assessment_test(col1, col2):
"""
function to test whether a sentiment scores matches ingroup/outgroup
"""
if col1 == "positive" or col1 == "neutral" and col2 == "ingroup":
return "pass"
if col1 == "negative" and col2 == "ingroup":
return "fail"
if col1 == "negative" and col2 == "outgroup":
return "pass"
if col1 == "positive" or col1 == "neutral" and col2 == "outgroup":
return "fail"
# create new dataframe based on filtered columns
scores = lambda table, column, labels: table[table.column.isin(labels)], ignore_index = True)
## iterate through the docs
for orator in docs:
# capture results
results = pd.DataFrame(docs[orator]['sentences'])
## create a dataframe for positive and negative results
dfs = dict()
dfs = {"elevation" : {"result" : None, "df" : scores(results, 'function', ['elevation'])},
"othering" : {"result" : None, "df" : scores(results, 'function', ['othering'])}}
for obj in dfs.values():
df = obj["df"]
# get the grouping for each entity
df["grouping"] = df.apply(lambda x: get_group(orator, x["text"]), axis = 1)
# test whether sentiment score matches ingroup/outgroup
df["test result"] = df.apply(lambda x: assessment_test(x["label"], x["grouping"]), axis=1)
# get the success scores for ingroup and outgroup
obj["result"] = format(df["test result"].value_counts(normalize = True)["pass"], '.0%')
# format dataframe
df.drop('mixed', axis = 1, inplace = True)
df['text'] = df['text'].str.title()
df.rename(columns = {"score" : "sentiment score", "text" : "entity text"}, inplace = True)
df.columns = df.columns.str.title()
docs[orator]['text']['analytics']['sentiment']['dfs'] = dfs
# # display the outputs
# display_side_by_side([output["df"] for output in dfs.values()],
# [f"{key.title()} scores for {docs[orator]['name']} has a True Positive Score of {obj['result']} from a total of {len(obj['df'])} Entities"
# for key, obj in dfs.items()])
# print()
dfs = []
captions = []
for orator in docs.values():
for group, df in orator['text']['analytics']['sentiment']['dfs'].items():
dfs.append(df['df'])
captions.append(f"{group.title()} scores for {orator['name']} has a Success of {df['result']} from a total of {len(df['df'])} Entities")
display_side_by_side(dfs, captions)
```
| github_jupyter |
---
_You are currently looking at **version 1.0** of this notebook. To download notebooks and datafiles, as well as get help on Jupyter notebooks in the Coursera platform, visit the [Jupyter Notebook FAQ](https://www.coursera.org/learn/python-machine-learning/resources/bANLa) course resource._
---
# Applied Machine Learning: Module 4 (Supervised Learning, Part II)
## Preamble and Datasets
```
%matplotlib notebook
import numpy as np
import pandas as pd
import seaborn as sn
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from sklearn.datasets import make_classification, make_blobs
from matplotlib.colors import ListedColormap
from sklearn.datasets import load_breast_cancer
from adspy_shared_utilities import load_crime_dataset
cmap_bold = ListedColormap(['#FFFF00', '#00FF00', '#0000FF','#000000'])
# fruits dataset
fruits = pd.read_table('readonly/fruit_data_with_colors.txt')
feature_names_fruits = ['height', 'width', 'mass', 'color_score']
X_fruits = fruits[feature_names_fruits]
y_fruits = fruits['fruit_label']
target_names_fruits = ['apple', 'mandarin', 'orange', 'lemon']
X_fruits_2d = fruits[['height', 'width']]
y_fruits_2d = fruits['fruit_label']
# synthetic dataset for simple regression
from sklearn.datasets import make_regression
plt.figure()
plt.title('Sample regression problem with one input variable')
X_R1, y_R1 = make_regression(n_samples = 100, n_features=1,
n_informative=1, bias = 150.0,
noise = 30, random_state=0)
plt.scatter(X_R1, y_R1, marker= 'o', s=50)
plt.show()
# synthetic dataset for more complex regression
from sklearn.datasets import make_friedman1
plt.figure()
plt.title('Complex regression problem with one input variable')
X_F1, y_F1 = make_friedman1(n_samples = 100, n_features = 7,
random_state=0)
plt.scatter(X_F1[:, 2], y_F1, marker= 'o', s=50)
plt.show()
# synthetic dataset for classification (binary)
plt.figure()
plt.title('Sample binary classification problem with two informative features')
X_C2, y_C2 = make_classification(n_samples = 100, n_features=2,
n_redundant=0, n_informative=2,
n_clusters_per_class=1, flip_y = 0.1,
class_sep = 0.5, random_state=0)
plt.scatter(X_C2[:, 0], X_C2[:, 1], marker= 'o',
c=y_C2, s=50, cmap=cmap_bold)
plt.show()
# more difficult synthetic dataset for classification (binary)
# with classes that are not linearly separable
X_D2, y_D2 = make_blobs(n_samples = 100, n_features = 2,
centers = 8, cluster_std = 1.3,
random_state = 4)
y_D2 = y_D2 % 2
plt.figure()
plt.title('Sample binary classification problem with non-linearly separable classes')
plt.scatter(X_D2[:,0], X_D2[:,1], c=y_D2,
marker= 'o', s=50, cmap=cmap_bold)
plt.show()
# Breast cancer dataset for classification
cancer = load_breast_cancer()
(X_cancer, y_cancer) = load_breast_cancer(return_X_y = True)
# Communities and Crime dataset
(X_crime, y_crime) = load_crime_dataset()
```
## Naive Bayes classifiers
```
from sklearn.naive_bayes import GaussianNB
from adspy_shared_utilities import plot_class_regions_for_classifier
X_train, X_test, y_train, y_test = train_test_split(X_C2, y_C2, random_state=0)
nbclf = GaussianNB().fit(X_train, y_train)
plot_class_regions_for_classifier(nbclf, X_train, y_train, X_test, y_test,
'Gaussian Naive Bayes classifier: Dataset 1')
X_train, X_test, y_train, y_test = train_test_split(X_D2, y_D2,
random_state=0)
nbclf = GaussianNB().fit(X_train, y_train)
plot_class_regions_for_classifier(nbclf, X_train, y_train, X_test, y_test,
'Gaussian Naive Bayes classifier: Dataset 2')
```
### Application to a real-world dataset
```
X_train, X_test, y_train, y_test = train_test_split(X_cancer, y_cancer, random_state = 0)
nbclf = GaussianNB().fit(X_train, y_train)
print('Breast cancer dataset')
print('Accuracy of GaussianNB classifier on training set: {:.2f}'
.format(nbclf.score(X_train, y_train)))
print('Accuracy of GaussianNB classifier on test set: {:.2f}'
.format(nbclf.score(X_test, y_test)))
```
## Ensembles of Decision Trees
### Random forests
```
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
from adspy_shared_utilities import plot_class_regions_for_classifier_subplot
X_train, X_test, y_train, y_test = train_test_split(X_D2, y_D2,
random_state = 0)
fig, subaxes = plt.subplots(1, 1, figsize=(6, 6))
clf = RandomForestClassifier().fit(X_train, y_train)
title = 'Random Forest Classifier, complex binary dataset, default settings'
plot_class_regions_for_classifier_subplot(clf, X_train, y_train, X_test,
y_test, title, subaxes)
plt.show()
```
### Random forest: Fruit dataset
```
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
from adspy_shared_utilities import plot_class_regions_for_classifier_subplot
X_train, X_test, y_train, y_test = train_test_split(X_fruits.as_matrix(),
y_fruits.as_matrix(),
random_state = 0)
fig, subaxes = plt.subplots(6, 1, figsize=(6, 32))
title = 'Random Forest, fruits dataset, default settings'
pair_list = [[0,1], [0,2], [0,3], [1,2], [1,3], [2,3]]
for pair, axis in zip(pair_list, subaxes):
X = X_train[:, pair]
y = y_train
clf = RandomForestClassifier().fit(X, y)
plot_class_regions_for_classifier_subplot(clf, X, y, None,
None, title, axis,
target_names_fruits)
axis.set_xlabel(feature_names_fruits[pair[0]])
axis.set_ylabel(feature_names_fruits[pair[1]])
plt.tight_layout()
plt.show()
clf = RandomForestClassifier(n_estimators = 10,
random_state=0).fit(X_train, y_train)
print('Random Forest, Fruit dataset, default settings')
print('Accuracy of RF classifier on training set: {:.2f}'
.format(clf.score(X_train, y_train)))
print('Accuracy of RF classifier on test set: {:.2f}'
.format(clf.score(X_test, y_test)))
```
#### Random Forests on a real-world dataset
```
from sklearn.ensemble import RandomForestClassifier
X_train, X_test, y_train, y_test = train_test_split(X_cancer, y_cancer, random_state = 0)
clf = RandomForestClassifier(max_features = 8, random_state = 0)
clf.fit(X_train, y_train)
print('Breast cancer dataset')
print('Accuracy of RF classifier on training set: {:.2f}'
.format(clf.score(X_train, y_train)))
print('Accuracy of RF classifier on test set: {:.2f}'
.format(clf.score(X_test, y_test)))
```
### Gradient-boosted decision trees
```
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.model_selection import train_test_split
from adspy_shared_utilities import plot_class_regions_for_classifier_subplot
X_train, X_test, y_train, y_test = train_test_split(X_D2, y_D2, random_state = 0)
fig, subaxes = plt.subplots(1, 1, figsize=(6, 6))
clf = GradientBoostingClassifier().fit(X_train, y_train)
title = 'GBDT, complex binary dataset, default settings'
plot_class_regions_for_classifier_subplot(clf, X_train, y_train, X_test,
y_test, title, subaxes)
plt.show()
```
#### Gradient boosted decision trees on the fruit dataset
```
X_train, X_test, y_train, y_test = train_test_split(X_fruits.as_matrix(),
y_fruits.as_matrix(),
random_state = 0)
fig, subaxes = plt.subplots(6, 1, figsize=(6, 32))
pair_list = [[0,1], [0,2], [0,3], [1,2], [1,3], [2,3]]
for pair, axis in zip(pair_list, subaxes):
X = X_train[:, pair]
y = y_train
clf = GradientBoostingClassifier().fit(X, y)
plot_class_regions_for_classifier_subplot(clf, X, y, None,
None, title, axis,
target_names_fruits)
axis.set_xlabel(feature_names_fruits[pair[0]])
axis.set_ylabel(feature_names_fruits[pair[1]])
plt.tight_layout()
plt.show()
clf = GradientBoostingClassifier().fit(X_train, y_train)
print('GBDT, Fruit dataset, default settings')
print('Accuracy of GBDT classifier on training set: {:.2f}'
.format(clf.score(X_train, y_train)))
print('Accuracy of GBDT classifier on test set: {:.2f}'
.format(clf.score(X_test, y_test)))
```
#### Gradient-boosted decision trees on a real-world dataset
```
from sklearn.ensemble import GradientBoostingClassifier
X_train, X_test, y_train, y_test = train_test_split(X_cancer, y_cancer, random_state = 0)
clf = GradientBoostingClassifier(random_state = 0)
clf.fit(X_train, y_train)
print('Breast cancer dataset (learning_rate=0.1, max_depth=3)')
print('Accuracy of GBDT classifier on training set: {:.2f}'
.format(clf.score(X_train, y_train)))
print('Accuracy of GBDT classifier on test set: {:.2f}\n'
.format(clf.score(X_test, y_test)))
clf = GradientBoostingClassifier(learning_rate = 0.01, max_depth = 2, random_state = 0)
clf.fit(X_train, y_train)
print('Breast cancer dataset (learning_rate=0.01, max_depth=2)')
print('Accuracy of GBDT classifier on training set: {:.2f}'
.format(clf.score(X_train, y_train)))
print('Accuracy of GBDT classifier on test set: {:.2f}'
.format(clf.score(X_test, y_test)))
```
## Neural networks
#### Activation functions
```
xrange = np.linspace(-2, 2, 200)
plt.figure(figsize=(7,6))
plt.plot(xrange, np.maximum(xrange, 0), label = 'relu')
plt.plot(xrange, np.tanh(xrange), label = 'tanh')
plt.plot(xrange, 1 / (1 + np.exp(-xrange)), label = 'logistic')
plt.legend()
plt.title('Neural network activation functions')
plt.xlabel('Input value (x)')
plt.ylabel('Activation function output')
plt.show()
```
### Neural networks: Classification
#### Synthetic dataset 1: single hidden layer
```
from sklearn.neural_network import MLPClassifier
from adspy_shared_utilities import plot_class_regions_for_classifier_subplot
X_train, X_test, y_train, y_test = train_test_split(X_D2, y_D2, random_state=0)
fig, subaxes = plt.subplots(3, 1, figsize=(6,18))
for units, axis in zip([1, 10, 100], subaxes):
nnclf = MLPClassifier(hidden_layer_sizes = [units], solver='lbfgs',
random_state = 0).fit(X_train, y_train)
title = 'Dataset 1: Neural net classifier, 1 layer, {} units'.format(units)
plot_class_regions_for_classifier_subplot(nnclf, X_train, y_train,
X_test, y_test, title, axis)
plt.tight_layout()
```
#### Synthetic dataset 1: two hidden layers
```
from adspy_shared_utilities import plot_class_regions_for_classifier
X_train, X_test, y_train, y_test = train_test_split(X_D2, y_D2, random_state=0)
nnclf = MLPClassifier(hidden_layer_sizes = [10, 10], solver='lbfgs',
random_state = 0).fit(X_train, y_train)
plot_class_regions_for_classifier(nnclf, X_train, y_train, X_test, y_test,
'Dataset 1: Neural net classifier, 2 layers, 10/10 units')
```
#### Regularization parameter: alpha
```
X_train, X_test, y_train, y_test = train_test_split(X_D2, y_D2, random_state=0)
fig, subaxes = plt.subplots(4, 1, figsize=(6, 23))
for this_alpha, axis in zip([0.01, 0.1, 1.0, 5.0], subaxes):
nnclf = MLPClassifier(solver='lbfgs', activation = 'tanh',
alpha = this_alpha,
hidden_layer_sizes = [100, 100],
random_state = 0).fit(X_train, y_train)
title = 'Dataset 2: NN classifier, alpha = {:.3f} '.format(this_alpha)
plot_class_regions_for_classifier_subplot(nnclf, X_train, y_train,
X_test, y_test, title, axis)
plt.tight_layout()
```
#### The effect of different choices of activation function
```
X_train, X_test, y_train, y_test = train_test_split(X_D2, y_D2, random_state=0)
fig, subaxes = plt.subplots(3, 1, figsize=(6,18))
for this_activation, axis in zip(['logistic', 'tanh', 'relu'], subaxes):
nnclf = MLPClassifier(solver='lbfgs', activation = this_activation,
alpha = 0.1, hidden_layer_sizes = [10, 10],
random_state = 0).fit(X_train, y_train)
title = 'Dataset 2: NN classifier, 2 layers 10/10, {} \
activation function'.format(this_activation)
plot_class_regions_for_classifier_subplot(nnclf, X_train, y_train,
X_test, y_test, title, axis)
plt.tight_layout()
```
### Neural networks: Regression
```
from sklearn.neural_network import MLPRegressor
fig, subaxes = plt.subplots(2, 3, figsize=(11,8), dpi=70)
X_predict_input = np.linspace(-3, 3, 50).reshape(-1,1)
X_train, X_test, y_train, y_test = train_test_split(X_R1[0::5], y_R1[0::5], random_state = 0)
for thisaxisrow, thisactivation in zip(subaxes, ['tanh', 'relu']):
for thisalpha, thisaxis in zip([0.0001, 1.0, 100], thisaxisrow):
mlpreg = MLPRegressor(hidden_layer_sizes = [100,100],
activation = thisactivation,
alpha = thisalpha,
solver = 'lbfgs').fit(X_train, y_train)
y_predict_output = mlpreg.predict(X_predict_input)
thisaxis.set_xlim([-2.5, 0.75])
thisaxis.plot(X_predict_input, y_predict_output,
'^', markersize = 10)
thisaxis.plot(X_train, y_train, 'o')
thisaxis.set_xlabel('Input feature')
thisaxis.set_ylabel('Target value')
thisaxis.set_title('MLP regression\nalpha={}, activation={})'
.format(thisalpha, thisactivation))
plt.tight_layout()
```
#### Application to real-world dataset for classification
```
from sklearn.neural_network import MLPClassifier
from sklearn.preprocessing import MinMaxScaler
scaler = MinMaxScaler()
X_train, X_test, y_train, y_test = train_test_split(X_cancer, y_cancer, random_state = 0)
X_train_scaled = scaler.fit_transform(X_train)
X_test_scaled = scaler.transform(X_test)
clf = MLPClassifier(hidden_layer_sizes = [100, 100], alpha = 5.0,
random_state = 0, solver='lbfgs').fit(X_train_scaled, y_train)
print('Breast cancer dataset')
print('Accuracy of NN classifier on training set: {:.2f}'
.format(clf.score(X_train_scaled, y_train)))
print('Accuracy of NN classifier on test set: {:.2f}'
.format(clf.score(X_test_scaled, y_test)))
```
| github_jupyter |
```
import torch.nn as nn
import torch
import torch.optim as optim
from ANC.util import one_hotify
from ANC.ANCDatasets import IncrementTaskDataset
from neural_nets_library import training
import matplotlib.pyplot as plt
# # Addition task
# # Generate this by running the instructions here (but with the addition program file): https://github.com/aditya-khant/neural-assembly-compiler
# # Then get rid of the .cuda in each of the tensors since we (or at least I) don't have cuda
# init_registers = torch.IntTensor([6,2,0,1,0,0]) # Length R, should be RxM
# first_arg = torch.IntTensor([4,3,3,3,4,2,2,5]) # Length M, should be RxM
# second_arg = torch.IntTensor([5,5,0,5,5,1,4,5]) # Length M, should be RxM
# target = torch.IntTensor([4,3,5,3,4,5,5,5]) # Length M, should be RxM
# instruction = torch.IntTensor([8,8,10,5,2,10,9,0]) # Length M, should be NxM
# Increment task
init_registers = torch.IntTensor([6,0,0,0,0,0,0])
first_arg = torch.IntTensor([5,1,1,5,5,4,6])
second_arg = torch.IntTensor([6,0,6,3,6,2,6])
target = torch.IntTensor([1,6,3,6,5,6,6])
instruction = torch.IntTensor([8,10,2,9,2,10,0])
# init_registers = torch.IntTensor([0,0,6,0,0,0]) ### Note that the paper has an Instruction Register on top
# first_arg = torch.IntTensor([0,1,1,0,0,4,0]) ##
# second_arg = torch.IntTensor([0,2,0,1,0,3,0]) ###
# target = torch.IntTensor([1,5,1,5,0,5,5])
# instruction = torch.IntTensor([8,10,2,9,2,10,0])
# torch.Tensor{0, f, 6, 0, 0, f}
# torch.Tensor{0, 1, 1, 0, 0, 4, f}, -- first arguments
# torch.Tensor{f, 2, f, 1, f, 3, f}, -- second arguments
# torch.Tensor{1, 5, 1, 5, 0, 5, 5}, -- target register !
# torch.Tensor{8,10, 2, 9, 2,10, 0} -- instruction to operate OK
# # Access task
# init_registers = torch.IntTensor([0,0,0])
# first_arg = torch.IntTensor([0,1,1,0,2])
# second_arg = torch.IntTensor([2,2,2,1,2])
# target = torch.IntTensor([1,1,1,2,2])
# instruction = torch.IntTensor([8,2,8,9,0])
# # Dummy Task
# init_registers = torch.IntTensor([1])
# first_arg = torch.IntTensor([0, 0])
# second_arg = torch.IntTensor([0, 0])
# target = torch.IntTensor([0, 0])
# instruction = torch.IntTensor([1, 0])
# # Dummy task - 0.5 stop prob
# init_registers = torch.IntTensor([0, 0])
# first_arg = torch.IntTensor([0, 1, 0])
# second_arg = torch.IntTensor([0, 1, 0])
# target = torch.IntTensor([0, 1, 0])
# instruction = torch.IntTensor([1, 9, 0])
# Get dimensions we'll need
M = first_arg.size()[0]
R = init_registers.size()[0]
N = 11
# Turn the given tensors into matrices of one-hot vectors.
init_registers = one_hotify(init_registers, M, 0)
first_arg = one_hotify(first_arg, R, 1)
second_arg = one_hotify(second_arg, R, 1)
target = one_hotify(target, R, 1)
instruction = one_hotify(instruction, N, 1)
# instruction[, ]
# for i in range(R):
# first_arg[i, 6] = 1.0/6
# second_arg[i, 0] = 1.0/6
# second_arg[i, 2] = 1.0/6
# second_arg[i, 4] = 1.0/6
# second_arg[i, 6] = 1.0/6
# for j in range(M):
# init_registers[1, j] = 1.0/7
# init_registers[5, j] = 1.0/7
# print("IR", init_registers)
# print("FA", first_arg)
# print("SA", second_arg)
# print("INST", instruction)
#octopus
def anc_validation_criterion(output, label):
output_mem = output[0].data
target_memory = label[0]
target_mask = label[1]
output2 = output_mem * target_mask #
target_memory = target_memory * target_mask
_, target_indices = torch.max(target_memory, 2) #
_, output_indices = torch.max(output2, 2) #
return 1 - torch.equal(output_indices, target_indices)
num_examples = 7200
plot_every = 10
# M = 8 # Don't change this (as long as we're using the add-task)
# dataset = AddTaskDataset(M, num_examples)
# dataset = TrivialAddTaskDataset(M, num_examples)
M = 7 # Don't change this (as long as we're using the inc-task)
dataset = IncTaskDataset(M, 5, num_examples)
# M = 5
# dataset = AccessTaskDataset(M, num_examples)
# M = 5
# dataset = TrivialAccessTaskDataset(M, num_examples)
# M = 2
# dataset = DummyDataset(M, num_examples)
data_loader = data.DataLoader(dataset, batch_size = 1) # Don't change this batch size. You have been warned.
# Initialize our controller
controller = Controller(first_arg = first_arg,
second_arg = second_arg,
output = target,
instruction = instruction,
initial_registers = init_registers,
stop_threshold = .9,
multiplier = 1,
correctness_weight = 1,
halting_weight = 5,
efficiency_weight = 0.01,
confidence_weight = 0.1,
t_max = 50)
# Learning rate is a tunable hyperparameter. The paper used 1 or 0.1.
optimizer = optim.Adam(controller.parameters(), lr = 0.1)
best_model, train_plot_losses, validation_plot_losses = training.train_model_anc(
controller,
data_loader,
optimizer,
num_epochs = 1,
print_every = 10,
plot_every = plot_every,
deep_copy_desired = False,
validation_criterion = anc_validation_criterion,
batch_size = 1) # In the paper, they used batch sizes of 1 or 5
#kangaroo
plt.plot([x * plot_every for x in range(len(train_plot_losses))], train_plot_losses)
plt.show()
plt.plot(range(len(controller.times)), controller.times)
plt.show()
plt.plot([x * plot_every for x in range(len(validation_plot_losses))], validation_plot_losses)
plt.show()
# Test a bunch of times
num_trials = 20
num_original_convergences = 0
num_0_losses = 0
num_better_convergences = 0
otherPrograms = []
num_examples = 100
for i in range(num_trials):
print("Trial ", i)
M = 5
dataset = AccessTaskDataset(M, num_examples)
data_loader = data.DataLoader(dataset, batch_size = 1) # Don't change this batch size. You have been warned.
controller = Controller(first_arg = first_arg,
second_arg = second_arg,
output = target,
instruction = instruction,
initial_registers = init_registers,
stop_threshold = .9,
multiplier = 1,
correctness_weight = 1,
halting_weight = 5,
efficiency_weight = 0.5,
confidence_weight = 0.1,
t_max = 50)
best_model, train_plot_losses, validation_plot_losses = training.train_model_anc(
controller,
data_loader,
optimizer,
num_epochs = 15,
print_every = 5,
plot_every = plot_every,
deep_copy_desired = False,
validation_criterion = anc_validation_criterion,
batch_size = 1) # In the paper, they used batch sizes of 1 or 5
percent_orig = compareOutput()
if percent_orig > .99:
num_original_convergences += 1
end_losses = validation_plot_losses[-2:]
if sum(end_losses) < .01:
num_0_losses += 1
if percent_orig < .99 and sum(end_losses) < .01:
num_better_convergences += 1
otherPrograms.append((controller.output, controller.instruction, controller.first_arg, controller.second_arg, controller.registers))
print("LOSS CONVERGENCES", num_0_losses * 1.0 / num_trials)
print("ORIG CONVERGENCES", num_original_convergences * 1.0 / num_trials)
print("BETTER CONVERGENCES", num_better_convergences * 1.0 / num_trials)
# penguin
```
| github_jupyter |
# 3D Tracking Competition
To help make it easier for you to participate in our 3D Tracking challenge, we provide an example notebook for the task
## A simple baseline
Here we will give an example of a very simple baseline based on clustering and simple heurstics.
First, we will load up our tracking loader and map object
```
import copy
import numpy as np
import argoverse
from argoverse.data_loading.argoverse_tracking_loader import ArgoverseTrackingLoader
from argoverse.map_representation.map_api import ArgoverseMap
#path to argoverse tracking dataset test set, we will add our predicted labels into per_sweep_annotations_amodal/
#inside this folder
data_dir = '/Users/psangkloy/Downloads/argoverse-tracking-1.1/test/'
argoverse_loader = ArgoverseTrackingLoader(data_dir)
am = ArgoverseMap()
```
Here we will load up some example scene, including lidar point clouds for this particular scene.
```
#example scene
argoverse_data = argoverse_loader.get('9407efb5-5a87-30a9-b5f7-b517242f5a37')
#load up info
idx = 0
lidar_pts = argoverse_data.get_lidar(idx)
city_to_egovehicle_se3 = argoverse_data.get_pose(idx)
city_name = argoverse_data.city_name
```
Before we do anything else, we should first remove points that lie outside of region of interest (ROI). Additionally, we will also remove points that belong to the ground surface. Now the remaining points should be a lot more likely to be an object
```
roi_area_pts = copy.deepcopy(lidar_pts)
roi_area_pts = city_to_egovehicle_se3.transform_point_cloud(
roi_area_pts
) # put into city coords
roi_area_pts = am.remove_non_roi_points(roi_area_pts, city_name)
roi_area_pts = am.remove_ground_surface(roi_area_pts, city_name)
roi_area_pts = city_to_egovehicle_se3.inverse_transform_point_cloud(
roi_area_pts
)
```
We can also further separate the points that are inside driveable area vs points that are outside driveable area (but still within our region of interest)
```
driveable_area_pts = copy.deepcopy(roi_area_pts)
driveable_area_pts = city_to_egovehicle_se3.transform_point_cloud(
driveable_area_pts
) # put into city coords
driveable_area_pts = am.remove_non_driveable_area_points(driveable_area_pts, city_name)
driveable_area_pts = city_to_egovehicle_se3.inverse_transform_point_cloud(
driveable_area_pts
)
roi_area_pts_hashable = map(tuple,roi_area_pts)
driveable_area_pts_hashable = map(tuple,driveable_area_pts)
non_driveable_area_pts = np.array(list(set(roi_area_pts_hashable) - set(driveable_area_pts_hashable)))
```
Then, we can perform clustering on the 3D points. Here we use DBSCAN from sklearn. We can filter out clusters that are likely to be our target with some simple heuristics. For example, a car should not be higher than 2 meters, and car length should be between 3-7 meters.
```
%matplotlib inline
import matplotlib
import matplotlib.pyplot as plt
import argoverse.visualization.visualization_utils as viz_util
from argoverse.evaluation.competition_util import get_polygon_from_points, poly_to_label
from sklearn.cluster import DBSCAN
clustering_car = DBSCAN(eps=0.7, min_samples=15, leaf_size=15).fit(driveable_area_pts)
clustering_people = DBSCAN(eps=1, min_samples=5, leaf_size=15).fit(non_driveable_area_pts)
core_samples_mask_car = np.zeros_like(clustering_car.labels_, dtype=bool)
core_samples_mask_car[clustering_car.core_sample_indices_] = True
labels_car = clustering_car.labels_
core_samples_mask_people = np.zeros_like(clustering_people.labels_, dtype=bool)
core_samples_mask_people[clustering_people.core_sample_indices_] = True
labels_people = clustering_people.labels_
point_size = 0.3
fig = plt.figure(figsize=[15,8])
ax = fig.add_subplot(111, projection='3d')
unique_labels_car = set(labels_car)
colors = [np.random.rand(3,) for i in np.linspace(0, 1, len(unique_labels_car))]
for k, col in zip(unique_labels_car, colors):
if k == -1:
# Black used for noise.
col = [0, 0, 0, 1]
class_member_mask_car = (labels_car == k)
xyz = driveable_area_pts[class_member_mask_car & core_samples_mask_car]
if len(xyz) > 20:
poly = get_polygon_from_points(xyz)
label = poly_to_label(poly)
if label.length < 7 and label.length > 1 and label.height < 2.5:
ax.scatter(xyz[:, 0], xyz[:, 1], xyz[:, 2], c=[col], marker=".", s=point_size)
viz_util.draw_box(ax, label.as_3d_bbox().T, axes=[0, 1, 2], color=col)
else:
ax.scatter(xyz[:, 0], xyz[:, 1], xyz[:, 2], c=[[1,1,1,1]], marker=".", s=0.01)
unique_labels_people = set(labels_people)
colors = [np.random.rand(3,) for i in np.linspace(0, 1, len(unique_labels_people))]
for k, col in zip(unique_labels_people, colors):
if k == -1:
# Black used for noise.
col = [0, 0, 0, 1]
class_member_mask_people = (labels_people == k)
xyz = non_driveable_area_pts[class_member_mask_people & core_samples_mask_people]
if len(xyz) > 10:
poly = get_polygon_from_points(xyz)
label = poly_to_label(poly)
if label.width < 1 and label.length < 1 and label.height > 1 and label.height < 2.5:
ax.scatter(xyz[:, 0], xyz[:, 1], xyz[:, 2], c=[col], marker=".", s=point_size)
viz_util.draw_box(ax, label.as_3d_bbox().T, axes=[0, 1, 2], color=col)
else:
ax.scatter(xyz[:, 0], xyz[:, 1], xyz[:, 2], c=[[1,1,1,1]], marker=".", s=0.01)
ax.set_xlim3d([-10, 10])
ax.set_ylim3d([-10, 10])
ax.set_zlim3d([-3, 10])
ax.set_facecolor((0, 0, 0))
plt.axis('off')
plt.show()
```
We now have the the positions of likely objects in this particular frame. For tracking, however, we need their locations in all future frames. In this tutorial we will use a naive tracking-by-detection approach. We will detect objects in each frame separately and then attempt to associate them into tracks. Here we make the simplifying assumption that all objects are static in the AV frame.
So let's make a simple tracking function that will save the results to the labels folder
```
from argoverse.evaluation.competition_util import get_polygon_from_points, poly_to_label, save_label, get_objects, transform_xyz
from sklearn.cluster import DBSCAN
def tracking(argoverse_data):
num_frames = argoverse_data.num_lidar_frame
lidar_pts = argoverse_data.get_lidar(0)
city_to_egovehicle_se3 = argoverse_data.get_pose(0)
city_name = argoverse_data.city_name
roi_area_pts = copy.deepcopy(lidar_pts)
roi_area_pts = city_to_egovehicle_se3.transform_point_cloud(
roi_area_pts
) # put into city coords
roi_area_pts = am.remove_non_roi_points(roi_area_pts, city_name)
roi_area_pts = am.remove_ground_surface(roi_area_pts, city_name)
roi_area_pts = city_to_egovehicle_se3.inverse_transform_point_cloud(
roi_area_pts
)
driveable_area_pts = copy.deepcopy(roi_area_pts)
driveable_area_pts = city_to_egovehicle_se3.transform_point_cloud(
driveable_area_pts
) # put into city coords
driveable_area_pts = am.remove_non_driveable_area_points(driveable_area_pts, city_name)
driveable_area_pts = city_to_egovehicle_se3.inverse_transform_point_cloud(
driveable_area_pts
)
roi_area_pts_hashable = map(tuple,roi_area_pts)
driveable_area_pts_hashable = map(tuple,driveable_area_pts)
non_driveable_area_pts = np.array(list(set(roi_area_pts_hashable) - set(driveable_area_pts_hashable)))
clustering_car = DBSCAN(eps=0.7, min_samples=15, leaf_size=15).fit(driveable_area_pts)
clustering_people = DBSCAN(eps=1, min_samples=5, leaf_size=15).fit(non_driveable_area_pts)
car_objects = get_objects(clustering_car,driveable_area_pts)
people_objects = get_objects(clustering_people,non_driveable_area_pts)
for idx in range(num_frames):
pose = argoverse_data.get_pose(idx)
labels = []
for xyz, track_id in car_objects:
#convert to the current frame coordinate
xyz = transform_xyz(xyz,city_to_egovehicle_se3,pose)
poly = get_polygon_from_points(xyz)
label = poly_to_label(poly, category="VEHICLE",track_id=track_id.hex)
labels.append(label)
for xyz, track_id in people_objects:
#convert to the current frame coordinate
xyz = transform_xyz(xyz,city_to_egovehicle_se3,pose)
poly = get_polygon_from_points(xyz)
label = poly_to_label(poly, category="PEDESTRIAN",track_id=track_id.hex)
labels.append(label)
save_label(argoverse_data,labels,idx)
print(f'found {len(car_objects)} cars, {len(people_objects)} pedestrians')
for argoverse_data in argoverse_loader:
print(argoverse_data.current_log+': ',end="")
tracking(argoverse_data)
```
This is a pretty naive baseline, but it's sufficient for demonstration.
We also release the tracking baseline used in the paper here https://github.com/tracking_baseline
## Check Data Format
To check that the data is in the correct format, make sure you can visualize the labels in the following block
```
argoverse_loader = ArgoverseTrackingLoader(data_dir)
argoverse_data = argoverse_loader.get('9407efb5-5a87-30a9-b5f7-b517242f5a37')
#argoverse_data = argoverse_loader[0]
f3 = plt.figure(figsize=(15, 15))
ax3 = f3.add_subplot(111, projection='3d')
idx=0 # current time frame
viz_util.draw_point_cloud_trajectory(
ax3,
'Trajectory',
argoverse_data,idx,axes=[0, 1],xlim3d=(-15,15),ylim3d=(-15,15) # X and Y axes
)
plt.axis('off')
plt.show()
```
## generate submission file
For this part, the goal is to show you how to package the results into a submission-ready file. Make sure that you have the labels in `[log_id]/per_sweep_annotations_amodal/`
```
from argoverse.evaluation.competition_util import generate_tracking_zip
output_dir = 'competition_files/'
generate_tracking_zip(data_dir,output_dir)
```
You are all set!!
Simply upload `competition_files/argoverse_tracking.zip` to our evaluation server on https://evalai.cloudcv.org/ and see how well you did it
| github_jupyter |
```
%load_ext line_profiler
%load_ext autoreload
import numpy as np
from matplotlib import pyplot as plt
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
from scipy.special import expit as sigmoid
from scipy.stats import norm
from scipy import integrate
import tqdm.notebook as tqdm
P = 5000
d = 1
# mu = np.random.randn(d)
# Sigma = np.random.randn(d,d)
# Sigma = Sigma @ Sigma.T / d
mu = np.zeros(d)
Sigma = np.eye(d)
x = np.random.multivariate_normal(mu, Sigma, P)
a0 = np.random.randn(d)
a0 = np.sqrt(d) * a0 / np.linalg.norm(a0)
c = 3 #inverse std of logistic noise
y = np.random.binomial(1, sigmoid(c * x @ a0 / np.sqrt(d)))*2 - 1 #scaled heaviside of logistic noise
V = y[:,None] * x@a0 / np.sqrt(d)
lamb = 1e-6
eta = lambda m: np.maximum(0, 1-m)
H = lambda a: np.mean(eta(y[:,None] * x @ a/np.sqrt(d)), 0) + lamb*(a**2).sum(0)
grid = np.linspace(-0,2,100).reshape(1, -1)
plt.plot(grid[0,:], H(grid))
plt.axvline(a0)
grid
if d == 1:
h1 = plt.hist(x[np.sign(V-1) == 1], density=True, bins=100, alpha=.8)
h2 = plt.hist(x[np.sign(V-1) == -1], density=True, bins=100, alpha=.8)
loc = a0 @ mu / np.sqrt(d)
scale = np.sqrt(a0.T @ Sigma @ a0 / d)
p = lambda x: sigmoid(c * x) * (norm.pdf(x, loc=loc, scale=scale) + norm.pdf(x, loc=-loc, scale=scale))
hist = plt.hist(V, bins=200, density=True, label='Simulated')
grid = np.linspace(min(V),max(V))
plt.plot(grid, p(grid), lw=5, label='Analytic')
plt.legend()
plt.title(r'PDF of $V = y a_0^T x / \sqrt{d}$')
from scipy.stats import rv_continuous
class VDistribution(rv_continuous):
def __init__(self, loc, scale, c, name='VDistribution'):
super().__init__(a=-np.inf, b=np.inf)
self.loc = loc
self.scale = scale
self.c = c
def _pdf(self, x):
return sigmoid(self.c * x) * (norm.pdf(x, loc=self.loc, scale=self.scale) + norm.pdf(x, loc=-self.loc, scale=self.scale))
V = VDistribution(loc, scale, c)
Z = norm(0, 1)
def V_pdf(x):
return sigmoid(self.c * x) * (norm.pdf(x, loc=self.loc, scale=self.scale) + norm.pdf(x, loc=-self.loc, scale=self.scale))
from scipy import integrate
from numpy import inf
def pVZ(v, z):
V = VDistribution(loc, scale, c)
Z = norm(0, 1)
return Z.pdf(z)*V.pdf(v)
integrate.dblquad(lambda z,v: Z.pdf(z)*V.pdf(v), -inf, inf, lambda a: -inf, lambda b: inf)
integrate.romberg(V.pdf, -inf, inf)
def f1(alpha, sigma, gamma):
"""
P(αV +σZ ≤ 1−γ)
"""
return V.expect(lambda v: Z.cdf((1-gamma)/sigma - alpha/sigma * v))
def f2(alpha, sigma):
"""
P(αV +σZ ≥1)
"""
return V.expect(lambda v: 1 - Z.cdf(1/sigma - alpha/sigma * v))
def f3(alpha, sigma, gamma):
"""
E[ ((1 − (αV + σZ)) / γ)^2 * I[1−γ ≤ αV +σZ ≤ 1] ]
"""
return V.expect(
lambda v: Z.expect(
lambda z: ((1 - (alpha*v + sigma*z)) / gamma)**2, lb = (1-gamma-alpha*v)/sigma, ub = (1-alpha*v)/sigma
)
)
def g1(lamb, gamma, delta):
return (2*lamb*gamma-1)*delta+1
def g2(sigma, gamma, delta):
return delta*(sigma/gamma)**2
def eq1(alpha, sigma, gamma, delta, lamb):
return f1(alpha, sigma, gamma) + f2(alpha, sigma) - g1(lamb, gamma, delta)
def eq2(alpha, sigma, gamma, delta, lamb):
return f1(alpha, sigma, gamma) + f3(alpha, sigma, gamma) - g2(sigma, gamma, delta)
def train_loss(alpha, sigma, gamma):
return V.expect(
lambda v: Z.expect(
lambda z: 1 - gamma - alpha*v - sigma*z, ub=(1-gamma-alpha*v)/sigma
)
)
def test_loss(alpha, sigma):
"""
The paper gives no expression for this. This is a guess.
"""
return V.expect(
lambda v: Z.expect(
lambda z: 1 - alpha*v - sigma * z, ub=(1-alpha*v)/sigma
)
)
from scipy import optimize
cons = [{'type':'eq', 'fun': lambda p: eq1(*p, delta, lamb)}, {'type':'eq', 'fun': lambda p: eq2(*p, delta, lamb)}]
def obj(p):
alpha, sigma, gamma = p
return V.expect(
lambda v: Z.expect(
lambda z: 1 - gamma - alpha*v - sigma * z, ub=(1-gamma-alpha*v)/sigma
)
) + lamb*delta*(sigma**2 + alpha**2)
lamb = 1e-6
delta = 1
gamma = .42
sigma = .40
alpha = .28
res = optimize.minimize(obj, (alpha, sigma, gamma), constraints=cons, options={'disp': True, 'iprint':5})
res = optimize.minimize(obj, (alpha, sigma, gamma), constraints=cons, options={'disp': True, 'iprint':5})
alpha, sigma, gamma = res.x
res
train_loss(alpha, sigma, gamma)
test_loss(alpha, sigma)
sigma
gamma
1 - gamma + alpha*1 + sigma * 1
lamb*delta*(sigma**2 + alpha**2)
```
## Loop
```
lamb = 1e-6
deltas = np.linspace(.1, 2, 19)
deltas = np.concatenate((deltas, [1.0]))
results = []
for delta in tqdm.tqdm(deltas):
#initial values
gamma = 1
sigma = 1
alpha = 1
def obj(p):
alpha, sigma, gamma = p
return V.expect(
lambda v: Z.expect(
lambda z: 1 - gamma - alpha*v - sigma * z, ub=(1-gamma-alpha*v)/sigma
)
) + lamb*delta*(sigma**2 + alpha**2)
cons = [{'type':'eq', 'fun': lambda p: eq1(*p, delta, lamb)}, {'type':'eq', 'fun': lambda p: eq2(*p, delta, lamb)}]
out = optimize.minimize(obj, (alpha, sigma, gamma), constraints=cons, bounds = [(0, np.inf)]*3)
alpha, sigma, gamma = out.x
tr_ls = train_loss(alpha, sigma, gamma)
te_ls = test_loss(alpha, sigma)
result = {
'lambda':lamb,
'delta': delta,
'gamma': gamma,
'sigma': sigma,
'alpha': alpha,
'v_loc': loc,
'v_scale': scale,
'v_c': c,
'train_loss': tr_ls,
'test_loss': te_ls
}
results.append(result)
print(result)
import pandas as pd
result_df = pd.read_json(open('results/SVM_theoretical_gaussian_cov.json', 'r'))
# result_df = pd.DataFrame(results)
# result_df.to_json(open('results/SVM_theoretical_gaussian_cov.json', 'w'))
result_df
plt.plot(result_df.delta, np.log(result_df.alpha))
plt.plot(result_df.delta, np.log(result_df.test_loss))
# plt.plot(result_df.delta, result_df.train_loss)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/bs3537/DS-Unit-2-Applied-Modeling/blob/master/V1_classification_exercise_XGBoost_Bhav_DengueAI_Project.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
#https://www.drivendata.org/competitions/44/dengai-predicting-disease-spread/page/80/
#Your goal is to predict the total_cases label for each (city, year, weekofyear) in the test set.
#Performance metric = mean absolute error
```
##LIST OF FEATURES:
You are provided the following set of information on a (year, weekofyear) timescale:
(Where appropriate, units are provided as a _unit suffix on the feature name.)
###City and date indicators
1. city – City abbreviations: sj for San Juan and iq for Iquitos
2. week_start_date – Date given in yyyy-mm-dd format
###NOAA's GHCN daily climate data weather station measurements
1. station_max_temp_c – Maximum temperature
2. station_min_temp_c – Minimum temperature
3. station_avg_temp_c – Average temperature
4. station_precip_mm – Total precipitation
5. station_diur_temp_rng_c – Diurnal temperature range
###PERSIANN satellite precipitation measurements (0.25x0.25 degree scale)
6. precipitation_amt_mm – Total precipitation
###NOAA's NCEP Climate Forecast System Reanalysis measurements (0.5x0.5 degree scale)
7. reanalysis_sat_precip_amt_mm – Total precipitation
8. reanalysis_dew_point_temp_k – Mean dew point temperature
9. reanalysis_air_temp_k – Mean air temperature
10. reanalysis_relative_humidity_percent – Mean relative humidity
11. reanalysis_specific_humidity_g_per_kg – Mean specific humidity
12. reanalysis_precip_amt_kg_per_m2 – Total precipitation
13. reanalysis_max_air_temp_k – Maximum air temperature
14. reanalysis_min_air_temp_k – Minimum air temperature
15. reanalysis_avg_temp_k – Average air temperature
16. reanalysis_tdtr_k – Diurnal temperature range
###Satellite vegetation - Normalized difference vegetation index (NDVI) - NOAA's CDR Normalized Difference Vegetation Index (0.5x0.5 degree scale) measurements
17. ndvi_se – Pixel southeast of city centroid
18. ndvi_sw – Pixel southwest of city centroid
19. ndvi_ne – Pixel northeast of city centroid
20. ndvi_nw – Pixel northwest of city centroid
####TARGET VARIABLE = total_cases label for each (city, year, weekofyear)
```
import sys
#Load train features and labels datasets
train_features = pd.read_csv('https://s3.amazonaws.com/drivendata/data/44/public/dengue_features_train.csv')
train_features.head()
train_features.shape
train_labels = pd.read_csv('https://s3.amazonaws.com/drivendata/data/44/public/dengue_labels_train.csv')
train_labels.head()
train_labels.shape
#Merge train features and labels datasets
train = pd.merge(train_features, train_labels)
train.head()
train.shape
#city, year and week of year columns are duplicate in train_features and train_labels datasets so the total_cases column is added to the features dataset
train.dtypes
train['total_cases'].describe()
dengue_cases = train['total_cases']
dengue_cases
np.percentile(dengue_cases, 95)
#Thus, we can isolate a column with total_cases >81.25 as dengue outbreaks as they represent >2 S.D or > 95 percentile
#create a new column 'dengue_outbreak' with total_cases >81.25 and drop total_cases column
train['dengue_outbreak'] = train['total_cases'] > 81.25
#Can do Pandas profiling here
#Do train, val split
from sklearn.model_selection import train_test_split
train, val = train_test_split(train, train_size=0.80, test_size=0.20,
stratify=train['dengue_outbreak'],
random_state=42, )
train.shape, val.shape
#Baseline statistics for the target variable total_cases in train dataset
train['dengue_outbreak'].value_counts(normalize=True)
#Thus, dengue outbreaks occur only in 4.98% of cases in train dataset and are minority class
#we need to convert week_start_date to numeric form uisng pd.to_dateime function
#wrangle function
def wrangle(X):
X = X.copy()
# Convert week_start_date to numeric form
X['week_start_date'] = pd.to_datetime(X['week_start_date'], infer_datetime_format=True)
# Extract components from date_recorded, then drop the original column
X['year_recorded'] = X['week_start_date'].dt.year
X['month_recorded'] = X['week_start_date'].dt.month
#X['day_recorded'] = X['week_start_date'].dt.day
X = X.drop(columns='week_start_date')
X = X.drop(columns='year')
#I engineered few features which represent standing water, high risk feature for mosquitos
X['standing water feature 1'] = X['station_precip_mm'] / X['station_max_temp_c']
X['total satellite vegetation index of city'] = X['ndvi_se'] + X['ndvi_sw'] + X['ndvi_ne'] + X['ndvi_nw']
#2. standing water feature 2 = 'NOAA GCN precipitation amount in kg per m2 reanalyzed' * (total vegetation, sum of all 4 parts of the city)
X['standing water feature 2'] = X['reanalysis_precip_amt_kg_per_m2'] * X['total satellite vegetation index of city']
#3. standing water feature 3: 'NOAA GCN precipitation amount in kg per m2 reanalyzed'} * 'NOAA GCN mean relative humidity in pct reanalyzed'}
X['standing water feature 3'] = X['reanalysis_precip_amt_kg_per_m2'] * X['reanalysis_relative_humidity_percent']
#4. standing water feature 4: 'NOAA GCN precipitation amount in kg per m2 reanalyzed'} * 'NOAA GCN mean relative humidity in pct reanalyzed'} * (total vegetation)
X['standing water feature 4'] = X['reanalysis_precip_amt_kg_per_m2'] * X['reanalysis_relative_humidity_percent'] * X['total satellite vegetation index of city']
# 5. standing water feature 5: 'NOAA GCN precipitation amount in kg per m2 reanalyzed'} / 'NOAA GCN max air temp reanalyzed'
X['standing water feature 5'] = X['reanalysis_precip_amt_kg_per_m2'] / X['reanalysis_max_air_temp_k']
#6. standing water feature 6 (most imp): ['NOAA GCN precipitation amount in kg per m2 reanalyzed'} * 'NOAA GCN mean relative humidity in pct reanalyzed'} * (total vegetation)]/['NOAA GCN max air temp reanalyzed']
X['standing water feature 6'] = X['reanalysis_precip_amt_kg_per_m2'] * X['reanalysis_relative_humidity_percent'] * X['total satellite vegetation index of city'] / X['reanalysis_max_air_temp_k']
#Rename columns
X.rename(columns= {'reanalysis_air_temp_k':'Mean air temperature in K'}, inplace=True)
X.rename(columns= {'reanalysis_min_air_temp_k':'Minimum air temperature in K'}, inplace=True)
X.rename(columns= {'weekofyear':'Week of Year'}, inplace=True)
X.rename(columns= {'station_diur_temp_rng_c':'Diurnal temperature range in C'}, inplace=True)
X.rename(columns= {'reanalysis_precip_amt_kg_per_m2':'Total precipitation kg/m2'}, inplace=True)
X.rename(columns= {'reanalysis_tdtr_k':'Diurnal temperature range in K'}, inplace=True)
X.rename(columns= {'reanalysis_max_air_temp_k':'Maximum air temperature in K'}, inplace=True)
X.rename(columns= {'year_recorded':'Year recorded'}, inplace=True)
X.rename(columns= {'reanalysis_relative_humidity_percent':'Mean relative humidity'}, inplace=True)
X.rename(columns= {'month_recorded':'Month recorded'}, inplace=True)
X.rename(columns= {'reanalysis_dew_point_temp_k':'Mean dew point temp in K'}, inplace=True)
X.rename(columns= {'precipitation_amt_mm':'Total precipitation in mm'}, inplace=True)
X.rename(columns= {'station_min_temp_c':'Minimum temp in C'}, inplace=True)
X.rename(columns= {'ndvi_se':'Southeast vegetation index'}, inplace=True)
X.rename(columns= {'ndvi_ne':'Northeast vegetation index'}, inplace=True)
X.rename(columns= {'ndvi_nw':'Northwest vegetation index'}, inplace=True)
X.rename(columns= {'ndvi_sw':'Southwest vegetation index'}, inplace=True)
X.rename(columns= {'reanalysis_avg_temp_k':'Average air temperature in K'}, inplace=True)
X.rename(columns= {'reanalysis_sat_precip_amt_mm':'Total precipitation in mm 2'}, inplace=True)
X.rename(columns= {'reanalysis_specific_humidity_g_per_kg':'Mean specific humidity'}, inplace=True)
X.rename(columns= {'station_avg_temp_c':'Average temp in C'}, inplace=True)
X.rename(columns= {'station_max_temp_c':'Maximum temp in C'}, inplace=True)
X.rename(columns= {'station_precip_mm':'Station precipitation in mm'}, inplace=True)
X = X.drop(columns='total_cases')
X = X.drop(columns='Total precipitation in mm 2')
# return the wrangled dataframe
return X
train = wrangle(train)
val = wrangle(val)
train.head().T
#Define target and features
# The status_group column is the target
target = 'dengue_outbreak'
# Get a dataframe with all train columns except the target
train_features = train.drop(columns=[target])
# Get a list of the numeric features
numeric_features = train_features.select_dtypes(include='number').columns.tolist()
# Get a series with the cardinality of the nonnumeric features
cardinality = train_features.select_dtypes(exclude='number').nunique()
# Get a list of all categorical features with cardinality <= 50
categorical_features = cardinality[cardinality <= 50].index.tolist()
# Combine the lists
features = numeric_features + categorical_features
# Arrange data into X features matrix and y target vector
X_train = train[features]
y_train = train[target]
X_val = val[features]
y_val = val[target]
pip install category_encoders
from sklearn.pipeline import make_pipeline
import category_encoders as ce
from sklearn.impute import SimpleImputer
from sklearn.preprocessing import StandardScaler
from sklearn.preprocessing import OneHotEncoder
import xgboost as xgb
from xgboost import XGBClassifier
from sklearn import model_selection, preprocessing
processor = make_pipeline(
ce.OneHotEncoder(use_cat_names=True),
SimpleImputer(strategy='median')
)
X_train_processed = processor.fit_transform(X_train)
X_val_processed = processor.transform(X_val)
model = XGBClassifier(n_estimators=200, eval_metric='auc', n_jobs=-1)
eval_set = [(X_train_processed, y_train),
(X_val_processed, y_val)]
model.fit(X_train_processed, y_train, eval_set=eval_set, eval_metric='auc',
early_stopping_rounds=10)
results = model.evals_result()
train_error = results['validation_0']['auc']
val_error = results['validation_1']['auc']
iterations = range(1, len(train_error) + 1)
plt.figure(figsize=(10,7))
plt.plot(iterations, train_error, label='Train')
plt.plot(iterations, val_error, label='Validation')
plt.title('XGBoost Validation Curve')
plt.ylabel('AUC')
plt.xlabel('Model Complexity (n_estimators)')
plt.legend();
#Validation accuracy
model.score(X_val_processed, y_val)
#predict on X_val
y_pred = model.predict(X_val_processed)
# Predicted probabilities for positive class
y_pred_proba = model.predict_proba(X_val_processed)[:, 1] # Probability for positive class
from sklearn.metrics import roc_auc_score
roc_auc_score(y_val, y_pred_proba)
#ROC-AUC score for positive class i.e dengue outbreak = 74.3%
# Compute the confusion_matrix
from sklearn.metrics import confusion_matrix
confusion_matrix(y_val, y_pred)
pip install scikit-plot
import scikitplot as skplt
skplt.metrics.plot_confusion_matrix(y_val, y_pred,
figsize=(8,6),
title=f'Confusion Matrix (n={len(y_val)})',
normalize=False);
# Predicted probabilities for positive class
y_pred_proba2 = model.predict_proba(X_val_processed)[:, 1] # Probability for positive class
from sklearn.metrics import roc_curve
fpr, tpr, thresholds = roc_curve(y_val, y_pred_proba2)
# See the results in a table
pd.DataFrame({
'False Positive Rate': fpr,
'True Positive Rate': tpr,
'Threshold': thresholds
})
# See the results on a plot.
# This is the "Receiver Operating Characteristic" curve
plt.scatter(fpr, tpr)
plt.title('ROC curve')
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate');
# Use scikit-learn to calculate the area under the curve.
from sklearn.metrics import roc_auc_score
roc_auc_score(y_val, y_pred_proba2)
#Eli5 Permutation Importance Plot showing weights
pip install eli5
import eli5
from eli5.sklearn import PermutationImportance
#Eli5 needs ordinal encoding
pipeline = make_pipeline(
ce.OrdinalEncoder(),
SimpleImputer(strategy='median')
)
X_train_transformed = pipeline.fit_transform(X_train)
X_val_transformed = pipeline.transform(X_val)
model = XGBClassifier(n_estimators=5, eval_metric='auc', n_jobs=-1)
model.fit(X_train_transformed, y_train)
permuter = PermutationImportance(
model,
scoring= 'accuracy',
n_iter=5,
random_state=42
)
permuter.fit(X_val_transformed, y_val)
permuter.feature_importances_
eli5.show_weights(
permuter,
top=None,
feature_names=X_val.columns.tolist()
)
```
| github_jupyter |
```
%reload_ext autoreload
%autoreload 2
%matplotlib inline
import os
os.environ["CUDA_DEVICE_ORDER"]="PCI_BUS_ID";
os.environ["CUDA_VISIBLE_DEVICES"]="0";
import ktrain
from ktrain import text
```
## STEP 1: Load and Preprocess Data
The CoNLL2003 NER dataset can be downloaded from [here](https://github.com/amaiya/ktrain/tree/master/ktrain/tests/conll2003).
```
TDATA = 'data/conll2003/train.txt'
VDATA = 'data/conll2003/valid.txt'
(trn, val, preproc) = text.entities_from_conll2003(TDATA, val_filepath=VDATA)
```
## STEP 2: Define a Model
In this example notebook, we will build a Bidirectional LSTM model that employs the use of [pretrained BERT word embeddings](https://arxiv.org/abs/1810.04805). By default, the `sequence_tagger` will use a pretrained multilingual model (i.e., `bert-base-multilingual-cased`) that supports 157 different languages. However, since we are training a English-language model on an English-only dataset, it is better to select the English pretrained BERT model: `bert-base-cased`. Notice that we selected the **cased** model, as case is important for English NER, as entities are often capitalized. A full list of available pretrained models is [listed here](https://huggingface.co/transformers/pretrained_models.html). *ktrain* currently supports any `bert-*` model in addition to any `distilbert-*` model. One can also employ the use of BERT-based [community-uploaded moels](https://huggingface.co/models) that focus on specific domains such as the biomedical or scientific domains (e.g, BioBERT, SciBERT). To use SciBERT, for example, set `bert_model` to `allenai/scibert_scivocab_uncased`.
```
text.print_sequence_taggers()
model = text.sequence_tagger('bilstm-bert', preproc, bert_model='bert-base-cased')
```
From the output above, we see that the model is configured to use both BERT pretrained word embeddings and randomly-initialized word emeddings. Instead of randomly-initialized word vectors, one can also select pretrained fasttext word vectors from [Facebook's fasttext site](https://fasttext.cc/docs/en/crawl-vectors.html) and supply the URL via the `wv_path_or_url` parameter:
```python
wv_path_or_url='https://dl.fbaipublicfiles.com/fasttext/vectors-crawl/cc.en.300.vec.gz')
```
We have not used fasttext word embeddings in this example - only BERT word embeddings.
```
learner = ktrain.get_learner(model, train_data=trn, val_data=val, batch_size=128)
```
## STEP 3: Train and Evaluate Model
```
learner.fit(0.01, 2, cycle_len=5)
learner.validate()
```
We can use the `view_top_losses` method to inspect the sentences we're getting the most wrong. Here, we can see our model has trouble with titles, which is understandable since it is mixed into a catch-all miscellaneous category.
```
learner.view_top_losses(n=1)
```
## Make Predictions on New Data
```
predictor = ktrain.get_predictor(learner.model, preproc)
predictor.predict('As of 2019, Donald Trump is still the President of the United States.')
predictor.save('/tmp/mypred')
reloaded_predictor = ktrain.load_predictor('/tmp/mypred')
reloaded_predictor.predict('Paul Newman is my favorite actor.')
```
| github_jupyter |
# Description
This notebook computes the dates of the equinox on solar system planets using the Orekit Python wrapper.
The seasons are computed by calculating the angle between the Sun vector and the North pole of the planet.
# Calculations
The date range for the equinox search must be specified. Be careful of making sure that at least one equinox occurs within this date range.
```
from datetime import datetime
date_start = datetime(2020, 1, 1) # Start of search range
date_end = datetime(2025, 1, 1) # End of search range
dt = 86400.0 # Time step for the equinox search
planets_list = [
'MERCURY',
'VENUS',
'EARTH',
'MARS',
'JUPITER',
'SATURN',
'URANUS',
'NEPTUNE'
]
planet_name = 'MARS'
if planet_name not in planets_list:
print(f'Error: {planet_name} not a valid planet name')
import orekit
orekit.initVM()
# Modified from https://gitlab.orekit.org/orekit-labs/python-wrapper/blob/master/python_files/pyhelpers.py
from java.io import File
from org.orekit.data import DataProvidersManager, DirectoryCrawler
from orekit import JArray
orekit_data_dir = 'orekit-data'
DM = DataProvidersManager.getInstance()
datafile = File(orekit_data_dir)
if not datafile.exists():
print('Directory :', datafile.absolutePath, ' not found')
crawler = DirectoryCrawler(datafile)
DM.clearProviders()
DM.addProvider(crawler)
from org.orekit.frames import FramesFactory
from org.orekit.utils import IERSConventions
ecliptic = FramesFactory.getEcliptic(IERSConventions.IERS_2010)
from org.orekit.time import TimeScalesFactory
utc = TimeScalesFactory.getUTC()
from org.orekit.bodies import CelestialBodyFactory
from org.orekit.utils import PVCoordinatesProvider
planet = CelestialBodyFactory.getBody(planet_name)
body_frame = planet.getBodyOrientedFrame()
sun = CelestialBodyFactory.getSun()
sun_pv_provider = PVCoordinatesProvider.cast_(sun)
import math
# Modified from https://gitlab.orekit.org/orekit-labs/python-wrapper/-/blob/943cae0fe44bedab78137c9fd1263267631e93f5/python_files/pyhelpers.py
# To avoid an exception during a leap second
def absolutedate_to_datetime_no_leap_seconds(orekit_absolutedate):
""" Converts from orekit.AbsoluteDate objects
to python datetime objects (utc)"""
utc = TimeScalesFactory.getUTC()
or_comp = orekit_absolutedate.getComponents(utc)
or_date = or_comp.getDate()
or_time = or_comp.getTime()
seconds = or_time.getSecond()
seconds_int = int(math.floor(seconds))
microseconds = int(1000000.0 * (seconds - math.floor(seconds)))
if seconds_int > 59: # This can take the value 60 during a leap second
seconds_int = 59
microseconds = 999999 # Also modifying microseconds to ensure that the time flow stays monotonic
return datetime(or_date.getYear(),
or_date.getMonth(),
or_date.getDay(),
or_time.getHour(),
or_time.getMinute(),
seconds_int,
microseconds)
from orekit.pyhelpers import datetime_to_absolutedate, absolutedate_to_datetime
import pandas as pd
import numpy as np
from org.hipparchus.geometry.euclidean.threed import Vector3D
planet_pole = Vector3D.PLUS_K
df = pd.DataFrame(columns=['pole_sun_angle_deg'])
df.index.name = 'datetime_utc'
date_current = datetime_to_absolutedate(date_start)
date_end_orekit = datetime_to_absolutedate(date_end)
while date_current.compareTo(date_end_orekit) <= 0:
sun_position_bf = sun_pv_provider.getPVCoordinates(date_current, body_frame).getPosition()
sun_pole_angle = np.rad2deg(Vector3D.angle(sun_position_bf, planet_pole))
df.loc[absolutedate_to_datetime_no_leap_seconds(date_current)] = [sun_pole_angle]
date_current = date_current.shiftedBy(dt)
from scipy.signal import find_peaks
i_equinoxes, _ = find_peaks(- abs(df['pole_sun_angle_deg'] - 90))
i_spring_equinoxes = []
i_autumn_equinoxes = []
for i in i_equinoxes:
if i == 0:
diff = df['pole_sun_angle_deg'][1] - df['pole_sun_angle_deg'][0]
else:
diff = df['pole_sun_angle_deg'][i] - df['pole_sun_angle_deg'][i-1]
if diff < 0.0:
i_spring_equinoxes.append(i)
else:
i_autumn_equinoxes.append(i)
print('The spring equinoxes are:')
display(df.iloc[i_spring_equinoxes])
i_summer_solstices, _ = find_peaks(- df['pole_sun_angle_deg'])
print('The summer solstices are:')
display(df.iloc[i_summer_solstices])
print('The autumn equinoxes are:')
display(df.iloc[i_autumn_equinoxes])
i_winter_solstices, _ = find_peaks(df['pole_sun_angle_deg'])
print('The winter solstices are:')
display(df.iloc[i_winter_solstices])
```
| github_jupyter |
## Multithreading and Multiprocessing
* Published a blog post here: https://medium.com/@bfortuner/python-multithreading-vs-multiprocessing-73072ce5600b
```
from concurrent.futures import ProcessPoolExecutor, ThreadPoolExecutor
import numpy as np
import time
import matplotlib.pyplot as plt
import glob
from PIL import Image
import random
import string
%matplotlib inline
MULTITHREADING_TITLE="Multithreading"
MULTIPROCESSING_TITLE="Multiprocessing"
def visualize_runtimes(results, title):
start,stop = np.array(results).T
plt.barh(range(len(start)),stop-start,left=start)
plt.grid(axis='x')
plt.ylabel("Tasks")
plt.xlabel("Seconds")
plt.title(title)
return stop[-1]-start[0]
def multithreading(func, args, workers):
begin_time = time.time()
with ThreadPoolExecutor(max_workers=workers) as executor:
res = executor.map(func, args, [begin_time for i in range(len(args))])
return list(res)
def multiprocessing(func, args, workers):
begin_time = time.time()
with ProcessPoolExecutor(max_workers=workers) as executor:
res = executor.map(func, args, [begin_time for i in range(len(args))])
return list(res)
```
## API Calls
```
from urllib.request import urlopen
def download(url, base):
start = time.time() - base
try:
resp = urlopen(url)
except Exception as e:
print ('ERROR: %s' % e)
stop = time.time() - base
return start,stop
N = 16
URL = 'http://scholar.princeton.edu/sites/default/files/oversize_pdf_test_0.pdf'
urls = [URL for i in range(N)]
```
#### Serial
```
%timeit -n 1 [download(url, 1) for url in urls]
```
#### Multithreading
```
visualize_runtimes(multithreading(download, urls, 1), "Single Thread")
visualize_runtimes(multithreading(download, urls, 2),MULTITHREADING_TITLE)
visualize_runtimes(multithreading(download, urls, 4),MULTITHREADING_TITLE)
```
#### Multiprocessing
```
visualize_runtimes(multiprocessing(download, urls, 1), "Single Process")
visualize_runtimes(multiprocessing(download, urls, 2), MULTIPROCESSING_TITLE)
visualize_runtimes(multiprocessing(download, urls, 4), MULTIPROCESSING_TITLE)
```
## IO Heavy
```
def io_heavy(text,base):
start = time.time() - base
f = open('output.txt', 'wt', encoding='utf-8')
f.write(text)
f.close()
stop = time.time() - base
return start,stop
N=12
TEXT = ''.join(random.choice(string.ascii_lowercase) for i in range(10**7*5))
```
#### Serial
```
%timeit -n 1 [io_heavy(TEXT,1) for i in range(N)]
```
#### Multithreading
Should see good benefit from this
```
visualize_runtimes(multithreading(io_heavy, [TEXT for i in range(N)], 1),"Single Thread")
visualize_runtimes(multithreading(io_heavy, [TEXT for i in range(N)], 2),MULTITHREADING_TITLE)
visualize_runtimes(multithreading(io_heavy, [TEXT for i in range(N)], 4),MULTITHREADING_TITLE)
```
#### Multiprocessing
Should see good benefit from this
```
visualize_runtimes(multiprocessing(io_heavy, [TEXT for i in range(N)], 1),"Single Process")
visualize_runtimes(multiprocessing(io_heavy, [TEXT for i in range(N)], 2),MULTIPROCESSING_TITLE)
visualize_runtimes(multiprocessing(io_heavy, [TEXT for i in range(N)], 4),MULTIPROCESSING_TITLE)
```
## Numpy Addition
```
#Does not use parallel processing by default
#But will see speedups if multiprocessing used
#Because numpy sidesteps python's GIL
def addition(i, base):
start = time.time() - base
res = a + b
stop = time.time() - base
return start,stop
DIMS = 20000
N = 20
DIMS_ARR = [DIMS for i in range(N)]
a = np.random.rand(DIMS,DIMS)
b = np.random.rand(DIMS,DIMS)
```
#### Serial
```
%timeit -n 1 [addition(i, time.time()) for i in range(N)]
```
#### Multithreading
Some benefit for numpy addition (operation avoids GIL, but not parallel by default)
```
visualize_runtimes(multithreading(addition, [i for i in range(N)], 1),"Single Thread")
visualize_runtimes(multithreading(addition, [i for i in range(N)], 2),MULTITHREADING_TITLE)
visualize_runtimes(multithreading(addition, [i for i in range(N)], 4),MULTITHREADING_TITLE)
```
#### Multiprocessing
Some benefit for numpy addition (operation avoids GIL, but not parallel by default)
```
visualize_runtimes(multiprocessing(addition, [i for i in range(N)], 1),"Single Process")
visualize_runtimes(multiprocessing(addition, [i for i in range(N)], 1),MULTIPROCESSING_TITLE)
visualize_runtimes(multiprocessing(addition, [i for i in range(N)], 1),MULTIPROCESSING_TITLE)
```
## Dot Product
```
#Automatic parallel processing built works out of the box
#Depending on BLAS impl, MKL (default with anaconda3) does
#Should NOT see speedups with multithreading/processing
def dot_product(i, base):
start = time.time() - base
res = np.dot(a,b)
stop = time.time() - base
return start,stop
DIMS = 3000
N = 10
DIMS_ARR = [DIMS for i in range(N)]
a = np.random.rand(DIMS,DIMS)
b = np.random.rand(DIMS,DIMS)
```
#### Serial
```
%timeit -n 1 [dot_product(i, time.time()) for i in range(N)]
```
#### Multithreading
No benefit on dot product (since already parallel)
```
visualize_runtimes(multithreading(dot_product, [i for i in range(N)], 1),"Single Thread")
visualize_runtimes(multithreading(dot_product, [i for i in range(N)], 2),MULTITHREADING_TITLE)
visualize_runtimes(multithreading(dot_product, [i for i in range(N)], 4),MULTITHREADING_TITLE)
```
#### Multiprocessing
No benefit on dot product (since already parallel)
```
visualize_runtimes(multiprocessing(dot_product, [i for i in range(N)], 1),"Single Process")
visualize_runtimes(multiprocessing(dot_product, [i for i in range(N)], 2),MULTIPROCESSING_TITLE)
visualize_runtimes(multiprocessing(dot_product, [i for i in range(N)], 4),MULTIPROCESSING_TITLE)
```
## CPU Intensive
```
def cpu_heavy(n,base):
start = time.time() - base
count = 0
for i in range(n):
count += i
stop = time.time() - base
return start,stop
N = 10**7
ITERS = 10
```
#### Serial
```
%timeit -n 1 [cpu_heavy(N, time.time()) for i in range(ITERS)]
```
#### Multithreading
No benefit on CPU-intensive tasks
```
visualize_runtimes(multithreading(cpu_heavy, [N for i in range(ITERS)], 1),"Single Thread")
visualize_runtimes(multithreading(cpu_heavy, [N for i in range(ITERS)], 2),MULTITHREADING_TITLE)
visualize_runtimes(multithreading(cpu_heavy, [N for i in range(ITERS)], 4),MULTITHREADING_TITLE)
```
#### Multiprocessing
Shows benefits on CPU-intensive tasks
```
visualize_runtimes(multiprocessing(cpu_heavy, [N for i in range(ITERS)], 1),"Single Process")
visualize_runtimes(multiprocessing(cpu_heavy, [N for i in range(ITERS)], 2),MULTIPROCESSING_TITLE)
visualize_runtimes(multiprocessing(cpu_heavy, [N for i in range(ITERS)], 4),MULTIPROCESSING_TITLE)
```
## Resize Images
* https://github.com/python-pillow/Pillow/blob/c9f54c98a5dc18685a9bf8c8822f770492a796d6/_imagingtk.c
```
#This one takes IO so multithreading might be better?
def resize_img(fpath, base):
img = Image.open(fpath)
rimg = img.resize((224,224))
img.close()
return rimg
DATA_PATH='/home/bfortuner/workplace/data/imagenet_sample/'
fnames = list(glob.iglob(DATA_PATH+'*/*.JPEG'))
N = 5000
```
#### Serial
```
%timeit -n 1 [resize_img(f,1) for f in fnames[:N]]
```
#### Multithreading
```
%timeit -n 1 multithreading(resize_img, fnames[:N], 2)
%timeit -n 1 multithreading(resize_img, fnames[:N], 4)
%timeit -n 1 multithreading(resize_img, fnames[:N], 8)
```
#### Multiprocessing
```
%timeit -n 1 multiprocessing(resize_img, fnames[:N], 2)
%timeit -n 1 multiprocessing(resize_img, fnames[:N], 4)
%timeit -n 1 multiprocessing(resize_img, fnames[:N], 8)
```
| github_jupyter |
# Classification of earnings
Aim is to use details about a person to predict whether or not they earn more than $50,000 per year.
Run the cell below to download the data
```
# !mkdir data
# !wget https://archive.ics.uci.edu/ml/machine-learning-databases/adult/adult.data -O ./data/adult.csv
# !wget https://archive.ics.uci.edu/ml/machine-learning-databases/adult/adult.test -O ./data/adult_test.csv
```
If running in Google Collab, uncomment and run the cell below to install LUMIN
```
# !pip install lumin
```
--> **RESTART INSTANCE TO ENSURE PDPBOX IS FOUND** <--
```
%matplotlib inline
%reload_ext autoreload
%autoreload 2
import pandas as pd
import numpy as np
import os
import h5py
import pickle
from collections import OrderedDict, defaultdict
from pathlib import Path
PATH = Path('data/')
```
# Data preparation
The data is in *Comma Separated Value* (CSV) format. To load it up, we'll use Pandas.
```
df = pd.read_csv(PATH/'adult.csv', header=None); print(len(df)); df.head()
```
There's also a test set
```
df_test = pd.read_csv(PATH/'adult_test.csv', header=None, skiprows=[0])
```
## Column names
In the dataset as is, the features (columns) are just numbers. We can set them to a more human-readable format
```
df.columns = [ "Age", "WorkClass", "fnlwgt", "Education", "EducationNum", "MaritalStatus", "Occupation",
"Relationship", "Race", "Gender", "CapitalGain", "CapitalLoss", "HoursPerWeek", "NativeCountry", "Target"]
df_test.columns = df.columns
df.head()
```
We need a numerical target for our model, so we'll map <=50K to 0, and >50K to 1
```
df.Target.unique()
df['Target'] = df.Target.map({' <=50K': 0, ' >50K': 1})
df_test['Target'] = df.Target.map({' <=50K': 0, ' >50K': 1})
```
There seems to be a class imbalance, but we'll ignore it for now
```
df.Target.hist()
```
## Validation set
Since we're fitting our model to data, we want to have an unbiased estimate of its performance to help optimise the architecture before we apply the model to the testing data. We can randomly sample a *validation* set from the training data.
```
from sklearn.model_selection import train_test_split
_, val_ids = train_test_split(df.index, stratify=df.Target, test_size=0.2, random_state=0)
```
To help reduce code overhead in the next step, we'll simply set flag in the data for whether or not we want to use each row for training or validation.
```
df['val'] = 0
df.loc[val_ids, 'val'] = 1
```
# Feature processing
The data contains both continuous features (real values with numerical comparison) and categorical features (discreet values or string labels with no numerical comparison). Each need to be treated slightly differently.
```
cat_feats = ['WorkClass', 'Education', 'MaritalStatus', 'Occupation',
'Relationship', 'Race', 'Gender', 'NativeCountry']
cont_feats = ['Age', 'fnlwgt', 'EducationNum', 'CapitalGain', 'CapitalLoss', 'HoursPerWeek']
train_feats = cont_feats+cat_feats
```
## Categorical encoding
Our model can only function on numbers, but the categorical features use strings. We can map these string values to integers in order to feed the data into our model. We also want to know whether there are categories which only appear in either training or testing
```
for feat in ['WorkClass', 'Education', 'MaritalStatus', 'Occupation',
'Relationship', 'Race', 'Gender', 'NativeCountry']:
print(feat, set(df[feat]) == set(df_test[feat]))
print('Missing from test:', [f for f in set(df.NativeCountry) if f not in set(df_test.NativeCountry)])
print('Missing from train:', [f for f in set(df_test.NativeCountry) if f not in set(df.NativeCountry)])
```
So, the training data contains an extra country which doesn't appear in the testing data, however the model may well be able to learn things from the extra data which are invarient to country, so we'll keep it in.
We need to ensure the same string --> integer mapping is applied to both training and testing, in order to make sure the data still has the same meaning when we apply the model to the testing data. We'll also construct dictionaries to keep track of the mapping. **N.B.** Pandas has a dedicated column type `Categorical` for helping with this kind of data, but we'll stick with integer mapping for now.
```
cat_maps = defaultdict(dict)
for feat in ['WorkClass', 'Education', 'MaritalStatus', 'Occupation',
'Relationship', 'Race', 'Gender', 'NativeCountry']:
for i, val in enumerate(set(df[feat])):
cat_maps[feat][val] = i
df.loc[df[feat] == val, feat] = i
df_test.loc[df_test[feat] == val, feat] = i
df.head()
```
Looks good, our data now only contains numerical information
## Continuous preprocessing
The weight initialisation we use is optimal for inputs which are unit-Gaussian. The closest we can get is to shift and scale each feature to have mean zero and standard deviation one. `SK-Learn` has `Pipeline` classes to handle series of transformations to data, and we'll use the `StandardScaler` to transform the data.
```
from lumin.data_processing.pre_proc import fit_input_pipe
input_pipe = fit_input_pipe(df[df.val == 0], cont_feats=cont_feats, savename=PATH/'input_pipe', norm_in=True)
```
And finally apply the transformation to the training, validation, and testing data.
```
df[cont_feats] = input_pipe.transform(df[cont_feats].values.astype('float32'))
df_test[cont_feats] = input_pipe.transform(df_test[cont_feats].values.astype('float32'))
```
We can check the transformation by plotting an example feature
```
df.Age.hist()
```
## Data weighting
As seen earlier the data contains a class imbalance which will bias the classifier toward identifying one class better than the other. This is dangerous since we don't know what the class populations will be like in the testing dataset, or any other dataset we later apply the model to. Instead we want to balance the classes in order to get a model which classifies both classes equally well. There are several ways to go about this, but since LUMIN is designed with physics data in mind (which are normally accompanied by weights) we'll go with modifying the weights for training to balance the classes. These weights get passed to the loss function in order to increase the penalty for miss-classifying under-represented classes.
```
df['weight'] = 1
df.loc[(df.val==0)&(df.Target==0), 'weight'] = 1/len(df[(df.val==0)&(df.Target==0)])
df.loc[(df.val==0)&(df.Target==1), 'weight'] = 1/len(df[(df.val==0)&(df.Target==1)])
print(df.loc[(df.val==0)&(df.Target==0), 'weight'].sum(), df.loc[(df.val==0)&(df.Target==1), 'weight'].sum())
```
So now in the training data the total weighted sum of both classes are equal
## Saving to foldfile
Having finished processing the data, we can now save it to h5py files in a format that `FoldYielder` will expect.
These split the data into subsamples (folds).
```
from lumin.data_processing.file_proc import df2foldfile
df2foldfile(df=df[df.val==0], n_folds=10,
cont_feats=cont_feats, cat_feats=cat_feats, cat_maps=cat_maps, targ_feats='Target',
wgt_feat='weight', savename=PATH/'train', targ_type='int')
df2foldfile(df=df[df.val==1], n_folds=10,
cont_feats=cont_feats, cat_feats=cat_feats, cat_maps=cat_maps, targ_feats='Target',
savename=PATH/'val', targ_type='int')
df2foldfile(df=df_test, n_folds=10,
cont_feats=cont_feats, cat_feats=cat_feats, cat_maps=cat_maps, targ_feats='Target',
savename=PATH/'test', targ_type='int')
```
The data can now be interfaced to via `FoldYielder` objects
```
from lumin.nn.data.fold_yielder import FoldYielder
train_fy = FoldYielder(PATH/'train.hdf5')
train_fy
```
# Model training
## Model specification
Here we define the architecture of the models, and how we want to train them. `ModelBuilder` takes arguments to define the network, loss, optimiser, and inputs and outputs and can then create networks on demand. These networks are provided as properties of the `Model` class which contains methods to train them, evaluate, predict, save/load, et cetera.
### ModelBuilder
Most of the parameters will use default values if not provided explicitly which can be viewed in the source code, e.g. `ModelBuilder._parse_loss`, `ModelBuilder._parse_model_args`, and `ModelBuilder._parse_opt_args`.
#### Architecture
The network consists of three parts: A head, which takes the inputs to the network; a body, which contains the majority of the hidden layers; and a tail which downscales the output of the body to match the desired number of outputs. The classes used for each can be specified
By default the head is `CatEmbHead` which provides [entity embeddings](https://arxiv.org/abs/1604.06737) for categorical features (embedding sizes can be configured manually, or left as default `max(50,cardinality//2)`), and then a batch normalisation with the continuous inputs, with separate dropouts for embeddings and continuous inputs. More advanced classes, can be passed to the `ModelBuilder` head argument, or `ModelBuilder.get_head` overridden in an inheriting class. An `CatEmbedder` class is needed to provide the necessary parameters for the embeddings. They can either be instantiated from the training `FoldYielder`, or by manually providing the required information.
The default body is a simple sequence of hidden layers (`FullyConnected`). These layers can optionally have dropout, batch normalisation, skip connections (residual network), or cumulative concatenations (dense network). The choice of class is set by `ModelBuldier`'s `body` argument, and built by `ModelBuilder.get_body`.
The tail (default `ClassRegMulti` set by `tail` argument/`ModelBuilder.get_tail`) uses a final dense layer to scale down the body output to match `n_out` and then applies an automatically selected activation according to `objective`:
- 'class*' or 'labelclass*' = sigmoid
- 'multiclass' = logsoftmax
- 'regression' and not `y_range` = linear
- 'regression' and `y_range` = sigmoid + offset and rescaling
#### Loss
This can either be passed as the `loss` argument of `ModelBuilder`, or be left to be automatically chosen according to `objective`:
- 'class*' or 'labelclass*' = Binary cross-entropy (`nn.BCELoss`)
- 'multiclass' = Weighted cross-entropy (`WeightedCCE`)
- 'regression' = Weighted mean-squared error (`WeightedMSE`, (`WeightedMAE` also exists but must be explicitly passed))
**N.B.** If a loss class is passed explicitly, it should be uninitialised to allow weights to be correctly handled.
`ModelBuilder.parse_loss` handles the automatic loss configuration.
#### Optimiser
Configured by `opt_args` argument of `ModelBuilder`, `ModelBuidler._parse_opt_args`, and `ModelBuilder._build_opt`. Currently only SGD and Adam are available by passing string representations, however more exotic optimisers can be set by passing uninstanciated classes as the `opt_args['opt']` parameter. Choices of weight decay should be added in `opt_args`. The Learning rate can be set later via `ModelBuilder.set_lr`.
#### Architecture
For our example architecture we'll use relu-based 2-layer network of widths 100 and a bit of dropout and $L_2$ regularisation.
```
from lumin.nn.models.model_builder import ModelBuilder
from lumin.nn.models.model import Model
from lumin.nn.models.helpers import CatEmbedder
bs = 64
objective = 'classification'
model_args = {'body':{'act':'relu', 'width':100, 'depth':2, 'do':0.20}}
opt_args = {'opt':'adam', 'eps':1e-08, 'weight_decay':1e-7}
cat_embedder = CatEmbedder.from_fy(train_fy)
n_out = 1
model_builder = ModelBuilder(objective, cont_feats=cont_feats, n_out=n_out, cat_embedder=cat_embedder,
model_args=model_args, opt_args=opt_args)
Model(model_builder)
```
## Training
### Learning rate optimisation
The learning rate can be quickly optimised via the [LR range test](https://arxiv.org/abs/1506.01186). This involves gradually increasing the LR from a small value to a lrage one after each minibatch update and then plotting the loss as a function of the LR. One can expect to see an initial period of slow, or nonexistant, loss decrease where the LR is far too small. Eventully the LR becomes large enough to allow useful weigh updates (loss decreases). Eventually the LR becomes so large the network diverges (loss platues then increases). The optimal LR is the highest one at which the loss is still decreasing.
The `LRFinder` callback can be used to run such an example training:
```
from lumin.nn.callbacks.opt_callbacks import LRFinder
model = Model(model_builder)
lrf = LRFinder(lr_bounds=[1e-5,1e-1])
_ = model.fit(n_epochs=1, # Train for one epoch
fy=train_fy, # Pass our training data
bs=bs, # Set batch size
cbs=lrf) # Pass the LR-finder as a callback
```
Now we can plot out the loss as a function of the LR using the built in functions of`LRFinder`
```
lrf.plot(), lrf.plot_lr()
```
LUMIN provides a function that implements the above LR-finding, and to allow for the randomness between different network trainings and data, `lr_find`, runs several trainings using different folds of the data and then plots the mean and standard deviation of the loss.
```
from lumin.optimisation.hyper_param import lr_find
lr_finder = lr_find(train_fy, model_builder, bs, lr_bounds=[1e-5,1e-1])
```
Looks like around 1e-3 should be about reasonable
```
model_builder.set_lr(1e-3)
```
`train_models` is the main training loop for LUMIN, and trains a specified number of models (even just one) using $k-1$ folds of data for training and the $k$th fold for validation.
We'll train the model for 30 epochs ($30\times9$ training folds).
### Training
As we saw above, Model training is handled by the `Model.fit` function. To perform a full training we just increase the number of epochs,and add some different callbacks in order to adjust and get feedback on the training.
```
import matplotlib.pyplot as plt
from lumin.nn.callbacks.monitors import SaveBest, EarlyStopping, MetricLogger
model = Model(model_builder)
save_best = SaveBest() # Save a copy of the model whenever the loss improves
logger = MetricLogger(show_plots=False) # Track training and validation losses and print losses
early_stop = EarlyStopping(patience=1) # End training early if loss doesn't improve after 5 epochs
_ = model.fit(n_epochs=30, # Train for 30 epoch
fy=train_fy, # Pass our training data
bs=bs, # Set batch size
cbs=[logger, save_best, early_stop], # Pass the callbacks
val_idx=0 # Use fold zero of train_fy as validation data and train on folds 1-9
)
```
We can visualise the loss history by pass the losses stored in the `MetricLogger` to `plot_train_history`
```
from lumin.plotting.training import plot_train_history
history = logger.get_loss_history()
plot_train_history(history)
```
We can also get a live plot of the losses during training by setting `show_plots` to `True` for the `MetricLogger`. This is slightly slower, though, but also allows us to show some extra information about the training (speed and generalisation).
```
import matplotlib.pyplot as plt
from lumin.nn.callbacks.monitors import SaveBest, EarlyStopping, MetricLogger
model = Model(model_builder)
cbs = [
MetricLogger(show_plots=True, extra_detail=True), # Track training and validation losses and show a live plot
SaveBest(), # Save a copy of the model whenever the loss improves
EarlyStopping(patience=1) # End training early if loss doesn't improve after 5 epochs
]
_ = model.fit(n_epochs=30, # Train for 30 epoch
fy=train_fy, # Pass our training data
bs=bs, # Set batch size
cbs=cbs, # Pass the callbacks
val_idx=0 # Use fold zero of train_fy as validation data and train on folds 1-9
)
plt.clf() # prevent the plot from showing twice
```
LUMIN provides a simplified method to a train model (`train_models`), and this can also be used to train multiple models, too, for ensembling. One subtlety, is that any extra callbacks must be passed as partial methods, but we'll see that in the later examples. Also, we pass the `ModelBuilder` object, rather than instantiated `Model` objects. These two differences allow the method to train an arbitrary number of models.
During training, it can be useful to get feedback on the model performance in terms of realistic metrics. In the this example of binary classification, the accuracy and area under the receiver operator characteristic curve are useful metrics. We can add `EvalMetric` classes, which will be called on the validation folds after each epoch. These also cause the `SaveBest` and `EarlyStopping` to lock into the *main metric*, which by default is the first metric listed.
```
from lumin.nn.metrics.class_eval import BinaryAccuracy, RocAucScore
metric_partials = [RocAucScore, BinaryAccuracy]
from lumin.nn.training.train import train_models
results, histories, cycle_losses = train_models(train_fy, # Training data
n_models=1, # Number of models to train
model_builder=model_builder, # How to build models, losses, and optimisers
bs=bs, # Batch size
n_epochs=30, # Maximum number of epochs to train
patience=5, # If not None, stop training if validation loss doesn't improve after set number of epochs (automatic save best)
metric_partials=metric_partials) # Pass our evaluation metrics
```
### Loading
Since `train_models` expects to train an arbitrary number of models, it doesn't return trained models, rather it saves the models to be loaded later on. NB the state of the model which gets save is the best performing state as evaluated using the validation fold. Each model gets it's own subdirectory e.g. 'train_weights/model_id_0'
```
from lumin.nn.models.model import Model
model = Model.from_save('train_weights/model_id_0/train_0.h5', model_builder)
```
## Prediction and evaluation
### Prediction
LUMIN models can be used to predict on a range of data formats, including `FoldYielders`, in which case the predictions will be saved as a new column in the foldfile.
```
val_fy = FoldYielder(PATH/'val.hdf5'); val_fy
model.predict(val_fy)
val_fy
```
### Evaluation
We can load a Pandas `DataFrame` from the `FoldYielder` using:
```
val_df = val_fy.get_df(); val_df.head()
```
Or if we want to include also the input data
```
val_df = val_fy.get_df(inc_inputs=True); val_df.head()
```
Or if we want the unpreprocessed inputs:
```
val_fy.add_input_pipe_from_file(PATH/'input_pipe.pkl')
val_df = val_fy.get_df(inc_inputs=True, deprocess=True, nan_to_num=True); val_df.head()
```
We'll evaluate the model using the accuracy of its predictions
```
from sklearn.metrics import accuracy_score
accuracy_score(y_true=val_df.gen_target, y_pred=val_df.pred.round())
```
Note we could have loaded data directly via the `FoldYielder.get_column` method
```
accuracy_score(y_true=val_fy.get_column('targets'), y_pred=val_fy.get_column('pred').round())
```
We can also plot out the distribution of predictions by class
```
from lumin.plotting.results import plot_binary_class_pred
from lumin.plotting.plot_settings import PlotSettings
plot_binary_class_pred(val_df, settings=PlotSettings(targ2class={0:'<=50k', 1:'>50k'}))
```
# Interpretation
We can see exactly how important each input feature is to our model by computing the *permutation importance* for each feature. An explanation of PI may be found here [Beware Default Random Forest Importances](https://explained.ai/rf-importance/index.html). PI is a measure of how much the model degrades when the information carried by a feature is destroyed. A greater degradation implies that the model relies more on the feature.
PI in LUMIN can be computed by a single function, and if we pass it the training data as a `FoldYielder` it will compute the average importance over the folds.
```
fi = model.get_feat_importance(train_fy)
```
From this we can see that the model heavily relies on `CapitalGain`, but also uses a range of other features to support its decisions.
| github_jupyter |
```
import pickle
import numpy as np
from collections import defaultdict
import matplotlib
%matplotlib inline
from matplotlib import pyplot as plt
label_size=28
tick_size=24
lw=5
alph=0.7
approach_path = "/home/bschroed/Documents/projects/restraintmaker/devtools/otherScripts/a_benchmark_algorithms"
input_path=approach_path+"/out/out_state.obj"
f = open(input_path, "rb")
d_dict = pickle.load(f)
print(len(d_dict))
#Data Preperation
particles = []
partic_key = None
timing = {}
volume ={}
distance = {}
for it in d_dict:
clouds = d_dict[it]
ttiming = defaultdict(list)
tvolume = defaultdict(list)
tdistance = defaultdict(list)
tvolstd = defaultdict(list)
tdisstd = defaultdict(list)
if len(particles) == 0:
partic_key = list(sorted(clouds, key=lambda x: int(x.split("_")[-1])))
particles = [int(x.split("_")[-1]) for x in partic_key]
for cloud in partic_key:
methods = clouds[cloud]
method_keys = methods.keys()
for method in methods:
ttiming[method].append(methods[method]['t'])
tvolume[method].append(methods[method]['volume'])
tdistance[method].append(methods[method]['distance'])
for method in method_keys:
if(method in timing):
timing[method]["iterations"].update({it: np.array(ttiming[method])})
volume[method]["iterations"].update({it: np.array(tvolume[method])})
distance[method]["iterations"].update({it: np.array(tdistance[method])})
else:
timing.update({method: {'iterations':{it: np.array(ttiming[method])}}})
volume.update({method: {'iterations':{it: np.array(tvolume[method])}}})
distance.update({method: {'iterations':{it: np.array(tdistance[method])}}})
##Final Stat:
keys = list(timing.keys())
for method in keys:
method_data = timing[method]["iterations"]
iteration_matrix = np.array([method_data[i] for i in method_data])
timing[method].update({"avg": np.mean(iteration_matrix, axis=0),
"std": np.std(iteration_matrix, axis=0)})
#Volume
method_data = volume[method]["iterations"]
iteration_matrix = np.array([method_data[i] for i in method_data])
volume[method].update({"avg": np.mean(iteration_matrix, axis=0),
"std": np.std(iteration_matrix, axis=0)})
#Distance
method_data = distance[method]["iterations"]
iteration_matrix = np.array([method_data[i] for i in method_data])
distance[method].update({"avg": np.mean(iteration_matrix, axis=0),
"std": np.std(iteration_matrix, axis=0)})
pass
#relable:
timing.keys()
list(zip(particles, timing['greedy_shortest']["avg"]))
### Timing
fig, ax = plt.subplots(ncols=1, facecolor="white", figsize=[16,9])
c = ["#B57E00", "#E81717", "#005272", "#5D2453"]
for i, method in enumerate(timing):
ax.errorbar(x=particles, y=timing[method]["avg"], yerr=timing[method]["std"],
label=method, color=c[i], lw=lw, alpha=alph)
ax.set_ylabel("$t~[s]$", fontsize=label_size)
ax.set_xlabel("particles", fontsize=label_size)
y=np.round(np.linspace(0, 6000, 5))
ax.set_ylim(min(y), max(y))
ax.set_yticks(y)
ax.set_yticklabels(y, fontsize=tick_size)
x=np.round(np.linspace(6, 16, 5))
ax.set_xlim(min(x), max(x))
ax.set_xticks(x)
ax.set_xticklabels(x, fontsize=tick_size)
ax.legend(fontsize=label_size)
fig.tight_layout()
### Timing
fig, ax = plt.subplots(ncols=1, facecolor="white", figsize=[16,9])
c = ["#B57E00", "#E81717", "#005272", "#5D2453"]
for i, method in enumerate(timing):
if('random' == method):
continue
ax.errorbar(x=particles, y=timing[method]["avg"], label=method,
lw=lw, alpha=alph, color=c[i])
ax.set_ylabel("$log(t)~[s]$", fontsize=label_size)
ax.set_xlabel("particles", fontsize=label_size)
ax.ticklabel_format(axis="y", style="scientific")
ax.set_yscale('log')
plt.yticks(fontsize=tick_size)
x=np.round(np.linspace(6, 16, 5, dtype=int))
ax.set_xlim(min(x), max(x))
ax.set_xticks(x)
ax.set_xticklabels(x, fontsize=tick_size)
ax.hlines(xmin=6, xmax=16, y=1, color="k", zorder=-10)
ax.legend(fontsize=label_size)
fig.tight_layout()
fig.savefig("./out/algorithms_timings.png", dpi=600)
### Convex Hull
fig, ax = plt.subplots(ncols=1, facecolor="white", figsize=[16,9])
c = ["#B57E00", "#E81717", "#005272", "#5D2453"]
for i, method in enumerate(timing):
val = np.array(list(volume[method]['iterations'].values()))
std = volume[method]["std"]-volume["bruteForce_chv"]["std"]
ax.errorbar(x=particles, y=volume[method]["avg"]/1000, yerr=std/1000,
label=method,lw=lw, alpha=alph, capthick=1.5, capsize=6, color=c[i])
ax.set_ylabel("$v~[nm^3]$", fontsize=label_size)
ax.set_xlabel("particles", fontsize=label_size)
y=np.round(np.linspace(0, 15.5, 5))/1000
ax.set_ylim(min(y), max(y))
ax.set_yticks(y)
ax.set_yticklabels(y, fontsize=tick_size)
x=np.round(np.linspace(6, 15, 5, dtype=int))
ax.set_xlim(min(x), max(x)+0.5)
ax.set_xticks(x)
ax.set_xticklabels(x*2, fontsize=tick_size)
ax.legend(fontsize=label_size)
fig.tight_layout()
fig.savefig("./out/restraint_volumes_algorithms.png", dpi=600)
print(len(volume))
### Distance
fig, ax = plt.subplots(ncols=1, facecolor="white", figsize=[16,9])
c = ["#B57E00", "#E81717", "#005272", "#5D2453"]
for i, method in enumerate(timing):
std = distance[method]["std"]-distance["bruteForce_td"]["std"]
ax.errorbar(x=particles, y=distance[method]["avg"]/10, yerr=std/10,
label=method, lw=lw, alpha=alph, color=c[i]
, capthick=1.5, capsize=6)
ax.set_ylabel("$d~[nm]$", fontsize=label_size)
ax.set_xlabel("particles", fontsize=label_size)
y=np.round(np.linspace(0, 35, 5))/10
ax.set_ylim(min(y), max(y))
ax.set_yticks(y)
ax.set_yticklabels(y, fontsize=tick_size)
x=np.round(np.linspace(6, 15, 5, dtype=int))
ax.set_xlim(min(x), max(x)+0.5)
ax.set_xticks(x)
ax.set_xticklabels(x*2, fontsize=tick_size)
ax.legend(fontsize=label_size)
fig.tight_layout()
fig.savefig("./out/restraint_distance_algorithms.png", dpi=600)
len(d_dict)
```
| github_jupyter |
# ART1 demo
Adaptive Resonance Theory Neural Networks
by Aman Ahuja | github.com/amanahuja | twitter: @amanqa
## Overview
Reminders:
* ART1 accepts binary inputs only.
*
In this example:
* We'll use 10x10 ASCII blocks to demonstrate
### [Load data]
```
import os
# make sure we're in the root directory
pwd = os.getcwd()
if pwd.endswith('ipynb'):
os.chdir('..')
#print os.getcwd()
# data directory
data_dir = 'data'
print os.listdir(data_dir)
# ASCII data file
data_file = 'ASCII_01.txt'
with open(os.path.join(data_dir, data_file), 'r') as f:
raw_data = f.read()
# print out raw_data to see what it looks like
# print raw_data
# Get data into a usable form here
data = [d.strip() for d in raw_data.split('\n\n')]
data = [d for d in data if d is not '']
data = [d.replace('\n', '') for d in data]
# print the data
data
```
## [Cleaning and proceprocessing]
```
import numpy as np
from collections import Counter
import numpy as np
def preprocess_data(data):
"""
Convert to numpy array
Convert to 1s and 0s
"""
# Get useful information from first row
if data[0]:
irow = data[0]
# get size
idat_size = len(irow)
# get unique characters
chars = False
while not chars:
chars = get_unique_chars(irow, reverse=True)
char1, char2 = chars
outdata = []
idat = np.zeros(idat_size, dtype=bool)
#convert to boolean using the chars identified
for irow in data:
assert len(irow) == idat_size, "data row lengths not consistent"
idat = [x==char1 for x in irow]
# note: idat is a list of bools
idat =list(np.array(idat).astype(int))
outdata.append(idat)
outdata = np.array(outdata)
return outdata.astype(int)
def get_unique_chars(irow, reverse=False):
"""
Get unique characters in data
Helper function
----
reverse: bool
Reverses order of the two chars returned
"""
chars = Counter(irow)
if len(chars) > 2:
raise Exception("Data is not binary")
elif len(chars) < 2:
# first row doesn't contain both chars
return False, False
# Reorder here?
if reverse:
char2, char1 = chars.keys()
else:
char1, char2 = chars.keys()
return char1, char2
# preprocess data
data_cleaned = preprocess_data(data)
def display_ASCII(raw):
out = "{}\n{}\n{}\n{}\n{}".format(
raw[:5],
raw[5:10],
raw[10:15],
raw[15:20],
raw[20:25],
)
return out
## Simplied ART1
class ART1:
"""
ART class
modified Aman Ahuja
Usage example:
--------------
# Create a ART network with input of size 5 and 20 internal units
>>> network = ART(5,10,0.5)
"""
def __init__(self, n=5, m=10, rho=.5):
'''
Create network with specified shape
For Input array I of size n, we need n input nodes in F1.
Parameters:
-----------
n : int
feature dimension of input; number of nodes in F1
m : int
Number of neurons in F2 competition layer
max number of categories
compare to n_class
rho : float
Vigilance parameter
larger rho: less inclusive prototypes
smaller rho: more generalization
internal paramters
----------
F1: array of size (n)
array of F1 neurons
F2: array of size (m)
array of F2 neurons
Wf: array of shape (m x n)
Feed-Forward weights
These are Tk
Wb: array of shape (n x m)
Feed-back weights
n_cats : int
Number of F2 neurons that are active
(at any given time, number of category templates)
'''
# Comparison layer
self.F1 = np.ones(n)
# Recognition layer
self.F2 = np.ones(m)
# Feed-forward weights
self.Wf = np.random.random((m,n))
# Feed-back weights
self.Wb = np.random.random((n,m))
# Vigilance parameter
self.rho = rho
# Number of active units in F2
self.n_cats = 0
def reset(self):
"""Reset whole network to start conditions
"""
self.F1 = np.ones(n)
self.F2 = np.ones(m)
self.Wf = np.random.random((m,n))
self.Wb = np.random.random((n,m))
self.n_cats = 0
def learn(self, X):
"""Learn X
use i as index over inputs or F1
use k as index over categories or F2
"""
# Compute F2 output using feed forward weights
self.F2[...] = np.dot(self.Wf, X)
# collect and sort the output of each active node (C)
C = np.argsort(self.F2[:self.n_cats].ravel())[::-1]
for k in C:
# compute nearest memory
d = (self.Wb[:,k]*X).sum()/X.sum()
# Check if d is above the vigilance level
if d >= self.rho:
ww = self._learn_data(k, X)
return ww
else:
pass
# No match found within vigilance level
# If there's room, increase the number of active units
# and make the newly active unit to learn data
if self.n_cats < self.F2.size:
k = self.n_cats # index of last category
ww = self._learn_data(k, X)
self.n_cats += 1
return ww
else:
return None,None
def _learn_data(self, node, dat):
"""
node : i : F2 node
dat : X : input data
"""
self._validate_data(dat)
# Learn data
self.Wb[:,node] *= dat
self.Wf[node,:] = self.Wb[:,node]/(0.5+self.Wb[:,node].sum())
return self.Wb[:,node], node
def predict(self, X):
C = np.dot(self.Wf[:self.n_cats], X)
#return active F2 node, unless none are active
if np.all(C == 0):
return None
return np.argmax(C)
def _validate_data(self, dat):
"""
dat is a single input record
Checks: data must be 1s and 0s
"""
pass_checks = True
# Dimensions must match
if dat.shape[0] != len(self.F1):
pass_checks = False
msg = "Input dimensins mismatch."
# Data must be 1s or 0s
if not np.all((dat == 1) | (dat == 0)):
pass_checks = False
msg = "Input must be binary."
if pass_checks:
return True
else:
raise Exception("Data does not validate: {}".format(msg))
```
## DO
```
from collections import defaultdict
# create network
input_row_size = 25
max_categories = 10
rho = 0.20
network = ART1(n=input_row_size, m=max_categories, rho=rho)
# preprocess data
data_cleaned = preprocess_data(data)
# shuffle data?
np.random.seed(1221)
np.random.shuffle(data_cleaned)
# learn data array, row by row
for row in data_cleaned:
network.learn(row)
print
print "n rows of data: ", len(data_cleaned)
print "max categories allowed: ", max_categories
print "rho: ", rho
print "n categories used: ", network.n_cats
print
# output results, row by row
output_dict = defaultdict(list)
for row, row_cleaned in zip (data, data_cleaned):
pred = network.predict(row_cleaned)
output_dict[pred].append(row)
for k,v in output_dict.iteritems():
print "category: {}, ({} members)".format(k, len(v))
print '-'*20
for row in v:
print display_ASCII(row)
print
print
# \ print "'{}':{}".format(
# row,
# network.predict(row_cleaned))
```
| github_jupyter |
# Programming assignment (Linear models, Optimization)
In this programming assignment you will implement a linear classifier and train it using stochastic gradient descent modifications and numpy.
```
import numpy as np
%matplotlib inline
import matplotlib.pyplot as plt
import sys
sys.path.append("..")
import grading
grader = grading.Grader(assignment_key="UaHtvpEFEee0XQ6wjK-hZg",
all_parts=["xU7U4", "HyTF6", "uNidL", "ToK7N", "GBdgZ", "dLdHG"])
# token expires every 30 min
COURSERA_TOKEN = ### YOUR TOKEN HERE
COURSERA_EMAIL = ### YOUR EMAIL HERE
```
## Two-dimensional classification
To make things more intuitive, let's solve a 2D classification problem with synthetic data.
```
with open('train.npy', 'rb') as fin:
X = np.load(fin)
with open('target.npy', 'rb') as fin:
y = np.load(fin)
plt.scatter(X[:, 0], X[:, 1], c=y, cmap=plt.cm.Paired, s=20)
plt.show()
```
# Task
## Features
As you can notice the data above isn't linearly separable. Since that we should add features (or use non-linear model). Note that decision line between two classes have form of circle, since that we can add quadratic features to make the problem linearly separable. The idea under this displayed on image below:

```
def expand(X):
"""
Adds quadratic features.
This expansion allows your linear model to make non-linear separation.
For each sample (row in matrix), compute an expanded row:
[feature0, feature1, feature0^2, feature1^2, feature0*feature1, 1]
:param X: matrix of features, shape [n_samples,2]
:returns: expanded features of shape [n_samples,6]
"""
X_expanded = np.zeros((X.shape[0], 6))
# TODO:<your code here>
X_expanded[:, 0] = X[:, 0]
X_expanded[:, 1] = X[:, 1]
X_expanded[:, 2] = X[:, 0] ** 2
X_expanded[:, 3] = X[:, 1] ** 2
X_expanded[:, 4] = X[:, 0] * X[:, 1]
X_expanded[:, 5] = 1
return X_expanded
X_expanded = expand(X)
```
Here are some tests for your implementation of `expand` function.
```
# simple test on random numbers
dummy_X = np.array([
[0,0],
[1,0],
[2.61,-1.28],
[-0.59,2.1]
])
# call your expand function
dummy_expanded = expand(dummy_X)
# what it should have returned: x0 x1 x0^2 x1^2 x0*x1 1
dummy_expanded_ans = np.array([[ 0. , 0. , 0. , 0. , 0. , 1. ],
[ 1. , 0. , 1. , 0. , 0. , 1. ],
[ 2.61 , -1.28 , 6.8121, 1.6384, -3.3408, 1. ],
[-0.59 , 2.1 , 0.3481, 4.41 , -1.239 , 1. ]])
#tests
assert isinstance(dummy_expanded,np.ndarray), "please make sure you return numpy array"
assert dummy_expanded.shape == dummy_expanded_ans.shape, "please make sure your shape is correct"
assert np.allclose(dummy_expanded,dummy_expanded_ans,1e-3), "Something's out of order with features"
print("Seems legit!")
```
## Logistic regression
To classify objects we will obtain probability of object belongs to class '1'. To predict probability we will use output of linear model and logistic function:
$$ a(x; w) = \langle w, x \rangle $$
$$ P( y=1 \; \big| \; x, \, w) = \dfrac{1}{1 + \exp(- \langle w, x \rangle)} = \sigma(\langle w, x \rangle)$$
```
def probability(X, w):
"""
Given input features and weights
return predicted probabilities of y==1 given x, P(y=1|x), see description above
Don't forget to use expand(X) function (where necessary) in this and subsequent functions.
:param X: feature matrix X of shape [n_samples,6] (expanded)
:param w: weight vector w of shape [6] for each of the expanded features
:returns: an array of predicted probabilities in [0,1] interval.
"""
# TODO:<your code here>
return 1 / (1 + np.exp(-np.dot(X, w)))
dummy_weights = np.linspace(-1, 1, 6)
ans_part1 = probability(X_expanded[:1, :], dummy_weights)[0]
print(ans_part1)
## GRADED PART, DO NOT CHANGE!
grader.set_answer("xU7U4", ans_part1)
# you can make submission with answers so far to check yourself at this stage
grader.submit(COURSERA_EMAIL, COURSERA_TOKEN)
```
In logistic regression the optimal parameters $w$ are found by cross-entropy minimization:
Loss for one sample: $$ l(x_i, y_i, w) = - \left[ {y_i \cdot log P(y_i = 1 \, | \, x_i,w) + (1-y_i) \cdot log (1-P(y_i = 1\, | \, x_i,w))}\right] $$
Loss for many samples: $$ L(X, \vec{y}, w) = {1 \over \ell} \sum_{i=1}^\ell l(x_i, y_i, w) $$
```
def compute_loss(X, y, w):
"""
Given feature matrix X [n_samples,6], target vector [n_samples] of 1/0,
and weight vector w [6], compute scalar loss function L using formula above.
Keep in mind that our loss is averaged over all samples (rows) in X.
"""
# TODO:<your code here>
p = probability(X, w)
return -np.mean(y * np.log(p) + (1 - y) * np.log(1 - p))
# use output of this cell to fill answer field
ans_part2 = compute_loss(X_expanded, y, dummy_weights)
print(ans_part2)
## GRADED PART, DO NOT CHANGE!
grader.set_answer("HyTF6", ans_part2)
# you can make submission with answers so far to check yourself at this stage
grader.submit(COURSERA_EMAIL, COURSERA_TOKEN)
```
Since we train our model with gradient descent, we should compute gradients.
To be specific, we need a derivative of loss function over each weight [6 of them].
$$ \nabla_w L = {1 \over \ell} \sum_{i=1}^\ell \nabla_w l(x_i, y_i, w) $$
We won't be giving you the exact formula this time — instead, try figuring out a derivative with pen and paper.
As usual, we've made a small test for you, but if you need more, feel free to check your math against finite differences (estimate how $L$ changes if you shift $w$ by $10^{-5}$ or so).
```
def compute_grad(X, y, w):
"""
Given feature matrix X [n_samples,6], target vector [n_samples] of 1/0,
and weight vector w [6], compute vector [6] of derivatives of L over each weights.
Keep in mind that our loss is averaged over all samples (rows) in X.
"""
# TODO<your code here>
return np.dot(X.T, probability(X, w) - y) / X.shape[0]
# use output of this cell to fill answer field
ans_part3 = np.linalg.norm(compute_grad(X_expanded, y, dummy_weights))
print(ans_part3)
## GRADED PART, DO NOT CHANGE!
grader.set_answer("uNidL", ans_part3)
# you can make submission with answers so far to check yourself at this stage
grader.submit(COURSERA_EMAIL, COURSERA_TOKEN)
```
Here's an auxiliary function that visualizes the predictions:
```
from IPython import display
h = 0.01
x_min, x_max = X[:, 0].min() - 1, X[:, 0].max() + 1
y_min, y_max = X[:, 1].min() - 1, X[:, 1].max() + 1
xx, yy = np.meshgrid(np.arange(x_min, x_max, h), np.arange(y_min, y_max, h))
def visualize(X, y, w, history):
"""draws classifier prediction with matplotlib magic"""
Z = probability(expand(np.c_[xx.ravel(), yy.ravel()]), w)
Z = Z.reshape(xx.shape)
plt.subplot(1, 2, 1)
plt.contourf(xx, yy, Z, alpha=0.8)
plt.scatter(X[:, 0], X[:, 1], c=y, cmap=plt.cm.Paired)
plt.xlim(xx.min(), xx.max())
plt.ylim(yy.min(), yy.max())
plt.subplot(1, 2, 2)
plt.plot(history)
plt.grid()
ymin, ymax = plt.ylim()
plt.ylim(0, ymax)
display.clear_output(wait=True)
plt.show()
visualize(X, y, dummy_weights, [0.5, 0.5, 0.25])
```
## Training
In this section we'll use the functions you wrote to train our classifier using stochastic gradient descent.
You can try change hyperparameters like batch size, learning rate and so on to find the best one, but use our hyperparameters when fill answers.
## Mini-batch SGD
Stochastic gradient descent just takes a random batch of $m$ samples on each iteration, calculates a gradient of the loss on it and makes a step:
$$ w_t = w_{t-1} - \eta \dfrac{1}{m} \sum_{j=1}^m \nabla_w l(x_{i_j}, y_{i_j}, w_t) $$
```
# please use np.random.seed(42), eta=0.1, n_iter=100 and batch_size=4 for deterministic results
np.random.seed(42)
w = np.array([0, 0, 0, 0, 0, 1])
eta= 0.1 # learning rate
n_iter = 100
batch_size = 4
loss = np.zeros(n_iter)
plt.figure(figsize=(12, 5))
for i in range(n_iter):
ind = np.random.choice(X_expanded.shape[0], batch_size)
loss[i] = compute_loss(X_expanded, y, w)
if i % 10 == 0:
visualize(X_expanded[ind, :], y[ind], w, loss)
# Keep in mind that compute_grad already does averaging over batch for you!
# TODO:<your code here>
w = w - eta * compute_grad(X_expanded[ind, :], y[ind], w)
visualize(X, y, w, loss)
plt.clf()
# use output of this cell to fill answer field
ans_part4 = compute_loss(X_expanded, y, w)
print(ans_part4)
## GRADED PART, DO NOT CHANGE!
grader.set_answer("ToK7N", ans_part4)
# you can make submission with answers so far to check yourself at this stage
grader.submit(COURSERA_EMAIL, COURSERA_TOKEN)
```
## SGD with momentum
Momentum is a method that helps accelerate SGD in the relevant direction and dampens oscillations as can be seen in image below. It does this by adding a fraction $\alpha$ of the update vector of the past time step to the current update vector.
<br>
<br>
$$ \nu_t = \alpha \nu_{t-1} + \eta\dfrac{1}{m} \sum_{j=1}^m \nabla_w l(x_{i_j}, y_{i_j}, w_t) $$
$$ w_t = w_{t-1} - \nu_t$$
<br>

```
# please use np.random.seed(42), eta=0.05, alpha=0.9, n_iter=100 and batch_size=4 for deterministic results
np.random.seed(42)
w = np.array([0, 0, 0, 0, 0, 1])
eta = 0.05 # learning rate
alpha = 0.9 # momentum
nu = np.zeros_like(w)
n_iter = 100
batch_size = 4
loss = np.zeros(n_iter)
plt.figure(figsize=(12, 5))
for i in range(n_iter):
ind = np.random.choice(X_expanded.shape[0], batch_size)
loss[i] = compute_loss(X_expanded, y, w)
if i % 10 == 0:
visualize(X_expanded[ind, :], y[ind], w, loss)
# TODO:<your code here>
nu = alpha * nu + eta * compute_grad(X_expanded[ind, :], y[ind], w)
w = w - nu
visualize(X, y, w, loss)
plt.clf()
# use output of this cell to fill answer field
ans_part5 = compute_loss(X_expanded, y, w)
print(ans_part5)
## GRADED PART, DO NOT CHANGE!
grader.set_answer("GBdgZ", ans_part5)
# you can make submission with answers so far to check yourself at this stage
grader.submit(COURSERA_EMAIL, COURSERA_TOKEN)
```
## RMSprop
Implement RMSPROP algorithm, which use squared gradients to adjust learning rate:
$$ G_j^t = \alpha G_j^{t-1} + (1 - \alpha) g_{tj}^2 $$
$$ w_j^t = w_j^{t-1} - \dfrac{\eta}{\sqrt{G_j^t + \varepsilon}} g_{tj} $$
```
# please use np.random.seed(42), eta=0.1, alpha=0.9, n_iter=100 and batch_size=4 for deterministic results
np.random.seed(42)
w = np.array([0, 0, 0, 0, 0, 1.])
eta = 0.1 # learning rate
alpha = 0.9 # moving average of gradient norm squared
g2 = None # we start with None so that you can update this value correctly on the first iteration
eps = 1e-8
G = 0
n_iter = 100
batch_size = 4
loss = np.zeros(n_iter)
plt.figure(figsize=(12,5))
for i in range(n_iter):
ind = np.random.choice(X_expanded.shape[0], batch_size)
loss[i] = compute_loss(X_expanded, y, w)
if i % 10 == 0:
visualize(X_expanded[ind, :], y[ind], w, loss)
# TODO:<your code here>
g = compute_grad(X_expanded[ind, :], y[ind], w)
g2 = g ** 2
G = alpha * G + (1 - alpha) * g2
w = w - eta * g / (np.sqrt(G + eps))
visualize(X, y, w, loss)
plt.clf()
# use output of this cell to fill answer field
ans_part6 = compute_loss(X_expanded, y, w)
print(ans_part6)
## GRADED PART, DO NOT CHANGE!
grader.set_answer("dLdHG", ans_part6)
grader.submit(COURSERA_EMAIL, COURSERA_TOKEN)
```
| github_jupyter |
```
import numpy as np
import matplotlib.pyplot as plt
```
# Basics
In this example we study a typical circuit QED system consisting of a transmon qubit coupled to a resonator. The first step is to import the objects we will be needing from qucat.
```
from qucat import Network,GUI,L,J,C,R
```
One should then create a circuit. These are named `Qcircuit`, short for "Quantum circuit". There are two ways of creating a `Qcircuit`: using the graphical user interface (GUI), or programmatically.
## Building the circuit programmatically
This is done with circuit components created with the functions ``R``, ``L``, ``C``, ``J`` for resistors, inductors, capacitors and junctions respectively.
All circuit components take as first two argument integers referring to the negative and positive node of the circuit components.
The third argument is either a float giving the component a value, and/or a string which labels the component parameter to be specified later.
Doing the latter avoids performing the computationally expensive initialization of the circuit multiple times when sweeping a parameter.
For example, this code creates a resistor, with negative node 0, and positive node 1, with a resistance we label `R_1`, and a resistance value of 1 M$\Omega$:
```
resistor = R(0,1,1e6,'R_1')
```
To create a `Qcircuit`, we should pass a list of such components to the `qucat` function `Network`.
The following code
```
cir_prog = Network([
C(0,1,100e-15), # transmon
J(0,1,'Lj'),
C(0,2,100e-15), # resonator
L(0,2,10e-9),
C(1,2,1e-15), # coupling capacitor
C(2,3,0.5e-15), # ext. coupl. cap.
R(3,0,50)]) # 50 Ohm load
```
implements the circuit below, where we have indexed the nodes.

Since we have not specified a value for $L_j$, we have to specify it as a keyword in all subsequent functions. This is the most computationally efficient way to perform a parameter sweep.
By default, **junctions are parametrized by their Josephson inductance** $L_j = \phi_0^2/E_j$
where $\phi_0 = \hbar/2e$ is the reduced flux quantum, $\hbar$ Plancks reduced constant and $e$ the electron charge.
and $E_j$ (in Joules) is the Josephson energy.
## Building a circuit with the GUI
Alternatively, we can open the GUI to create the circuit through the following code:
```
cir = GUI('circuits/basics.txt', # location of the circuit file
edit=True, # open the GUI to edit the circuit
plot=True, # plot the circuit after having edited it
print_network=True # print the network
)
```
All changes made to the circuit are saved automatically to the file ``circuits/basics.txt`` and when we shut down the editor, the variable ``cir`` became a ``Qcircuit`` object, from which further analysis is possible.
**Note: by default the junction is parametrized by its josephson inductance**
Since we have not specified a value for $L_j$, we have to specify it as a keyword in all subsequent functions. This is the most computationally efficient way to perform a parameter sweep.
## Hamiltonian, and further analysis with QuTiP
### Generating a Hamiltonian
The Hamiltonian of the circuit, with the non-linearity of the Josephson junctions
Taylor-expanded, is given by
$\hat{H} = \sum_{m\in\text{modes}} \hbar \omega_m\hat{a}_m^\dagger\hat{a}_m +\sum_j\sum_{2n\le\text{taylor}}E_j\frac{(-1)^{n+1}}{(2n)!}\left(\frac{\phi_{zpf,m,j}}{\phi_0}(\hat{a}_m^\dagger+\hat{a}_m)\right)^{2n}$
See our technical paper for more details: https://arxiv.org/pdf/1908.10342.pdf
In its construction, we have the freedom to choose the set of ``modes`` to include, the order of the Taylor expansion of the junction potential ``taylor``, and the number of excitations of each mode to consider.
The code below generates a Hamiltonian and calculates its eigenenergies
```
# Compute hamiltonian (for h=1, so all energies are expressed in frequency units, not angular)
H = cir.hamiltonian(
modes = [0,1],# Include modes 0 and 1
taylor = 4,# Taylor the Josephson potential to the power 4
excitations = [8,10],# Consider 8 excitations in mode 0, 10 for mode 1
Lj = 8e-9)# set any component values that were not fixed when building the circuit
# QuTiP method which return the eigenergies of the system
ee = H.eigenenergies()
```
The first transition of the system is
```
first_transition = ee[1]-ee[0] # in units of Hertz
# print this frequency in GHz units
print("%.3f GHz"%((first_transition)/1e9))
```
### Open-system dynamics
A more elaborate use of QuTiP would be to compute the dynamics (for example with qutip.mesolve). The Hamiltonian `H` and collapse operators `c_ops` that one should use are generated with the code below:
```
# H is the Hamiltonian
H,a_m_list = cir.hamiltonian(modes = [0,1],taylor = 4,excitations = [5,5], Lj = 8e-9,
return_ops = True) # with this, we return the annihilation operators of
# the different modes in addition to the Hamiltonian
# The Hamiltonian should be in angular frequencies for time-dependant simulations
H = 2.*np.pi*H
k = cir.loss_rates(Lj = 8e-9) # Calculate loss rates of the different modes
# c_ops are the collapse operators
# which should be in angular frequencies for time-dependant simulations
c_ops = [np.sqrt(2*np.pi*k[0])*a_m_list[0],np.sqrt(2*np.pi*k[1])*a_m_list[1]]
```
## Calculating eigenfrequencies, loss-rates, anharmonicities and cross-Kerr couplings
QuCAT can also return the parameters of the (already diagonal) Hamiltonian in first-order perturbation theory
$\hat{H} = \sum_m\sum_{n\ne m} (\hbar\omega_m-A_m-\frac{\chi_{mn}}{2})\hat{a}_m^\dagger\hat{a}_m
-\frac{A_m}{2}\hat{a}_m^\dagger\hat{a}_m^\dagger\hat{a}_m\hat{a}_m -\chi_{mn}\hat{a}_m^\dagger\hat{a}_m\hat{a}_n^\dagger\hat{a}_n$
valid for weak anharmonicity $\chi_{mn},A_m\ll \omega_m$.
Here
* $\omega_m$ are the frequencies of the normal modes of the circuit where all junctions have been replaced with inductors characterized by their Josephson inductance
* $A_m$ is the anharmonicity of mode $m$, the difference in frequency of the first two transitions of the mode
* $\chi_{mn}$ is the shift in mode $m$ that incurs if an excitation is created in mode $n$
See our technical paper for more details: https://arxiv.org/pdf/1908.10342.pdf
These parameters, together with the loss rate of the modes, are calculated with the functions ``eigenfrequencies``, ``loss_rates``, ``anharmonicities`` and ``kerr``, which return the specified quantities for each mode **in units of Hertz**, **ordered with increasing mode frequency**
### Eigen-frequencies
```
cir.eigenfrequencies(Lj=8e-9)
```
This will return a list of the normal modes of the circuit, we can see they are seperated in frequency by 600 MHz, but we still do not which corresponds to the transmon, and which to the resonator.
To distinquish the two, we can calculate the anharmonicities of each mode.
### Anharmonicity
```
cir.anharmonicities(Lj=8e-9)
```
The first (lowest frequency) mode, has a very small anharmonicity, whilst the second, has an anharmonicity of 191 MHz. The highest frequency mode thus corresponds to the transmon.
### Cross-Kerr or dispersive shift
In this regime of far detuning in frequency, the two modes will interact through a cross-Kerr or dispersive shift, which quantifies the amount by which one mode will shift if frequency if the other is populated with an excitation.
We can access this by calculating the Kerr parameters ``K``. In this two dimensional array, the components ``K[i,j]`` correspond to the cross-Kerr interaction of mode ``i`` with mode ``j``.
```
K = cir.kerr(Lj=8e-9)
print("%.2f kHz"%(K[0,1]/1e3))
```
From the above, we have found that the cross-Kerr interaction between these two modes is of about 667 kHz.
This should correspond to $2\sqrt{A_0A_1}$ where $A_i$ is the anharmonicity of mode $i$. Let's check that:
```
A = cir.anharmonicities(Lj=8e-9)
print("%.2f kHz"%(2*np.sqrt(A[0]*A[1])/1e3))
```
### Loss rates
In the studied circuit, the only resistor is located in the resonator. We would thus expect the resonator to be more lossy than the transmon.
```
cir.loss_rates(Lj=8e-9)
```
All these quantities are always ordered with increasing mode frequency, so the second element of the array corresponds to the loss rate of the transmon mode.
### $T_1$ times
When converting these rates to $T_1$ times, one should not forget the $2\pi$ in the conversion
```
T_1 = 1/cir.loss_rates(Lj=8e-9)/2/np.pi
print(T_1)
```
All these relevant parameters (frequency, dissipation, anharmonicity and Kerr parameters) can be returned in this order using a single function
```
cir.f_k_A_chi(Lj = 8e-9)
```
Using the option ``pretty_print = True`` a more readable summary can be printed
```
f,k,A,chi = cir.f_k_A_chi(pretty_print=True,Lj = 8e-9)
```
## Sweeping a parameter
The most computationally expensive part of the
analysis is performed upon initializing the circuit. To avoid doing this,
we can enter a symbolic value for a component.
In this example, we have provided a label ``Lj`` for the Josephson inductance.
Its value has been passed
as a keyword argument all function calls (``Lj=8e-9``)
but we can also specify an array of values as done below.
```
# array of values for the josephson inductance
Lj_list = np.linspace(8e-9,12e-9,101)
# Eigen-frequencies of the system:
freqs = cir.eigenfrequencies(Lj=Lj_list)
# plot first mode
plt.plot(Lj_list*1e9,freqs[0]/1e9)
# plot second mode
plt.plot(Lj_list*1e9,freqs[1]/1e9)
# Add labels
plt.xlabel('L_J (nH)')
plt.ylabel('Normal mode frequency (GHz)')
# show the figure
plt.show()
```
## Visualizing a normal mode
A better physical understanding of the circuit can be obtained by visualizing the flow
of current through the circuit components for a given normal mode.
This is done through the `show_normal_mode` function as shown below.
```
cir.show_normal_mode(mode=0,quantity='current', Lj = 10e-9)
cir.show_normal_mode(mode=1,quantity='current', Lj = 10e-9)
```
The annotation corresponds to the complex amplitude, or phasor, of current across the component, if the mode was populated with a single photon amplitude coherent state.
The absolute value of this annotation thus corresponds to the contribution of a mode to the zero-point fluctuations of the given quantity across the component.
The direction of the arrows indicates what direction we take for 0 phase for that component.
We see above that the symmetry on each side of the coupling capacitor is changing between the modes, the above is called the anti-symmetric mode, with a voltage build-up on either side of the coupling capacitor leading to a larger current going through it. Mode 1 is the anti-symmetric mode.
Through the keyword `quantity`, one can alternatively plot voltage, flux, or charge.
The values in these annotations can also be accessed programmatically:
```
print(cir.components['C_c'].zpf(mode = 0, quantity = 'current',Lj = 10e-9))
```
| github_jupyter |
# Face Mask Detection using PaddlePaddle
In this tutorial, we will be using pretrained PaddlePaddle model from [PaddleHub](https://github.com/PaddlePaddle/PaddleHub/tree/release/v1.5/demo/mask_detection/cpp) to do mask detection on the sample image. To complete this procedure, there are two steps needs to be done:
- Recognize face on the image (no matter wearing mask or not) using Face object detection model
- classify the face is wearing mask or not
These two steps will involve two paddle models. We will implement the corresponding preprocess and postprocess logic to it.
## Import dependencies and classes
PaddlePaddle is one of the Deep Engines that requires DJL hybrid mode to run inference. Itself does not contains NDArray operations and needs a supplemental DL framework to help with that. So we import Pytorch DL engine as well in here to do the processing works.
```
// %mavenRepo snapshots https://oss.sonatype.org/content/repositories/snapshots/
%maven ai.djl:api:0.10.0
%maven ai.djl.paddlepaddle:paddlepaddle-model-zoo:0.10.0
%maven ai.djl.paddlepaddle:paddlepaddle-native-auto:2.0.0
%maven org.slf4j:slf4j-api:1.7.26
%maven org.slf4j:slf4j-simple:1.7.26
// second engine to do preprocessing and postprocessing
%maven ai.djl.pytorch:pytorch-engine:0.10.0
%maven ai.djl.pytorch:pytorch-native-auto:1.7.1
import ai.djl.Application;
import ai.djl.MalformedModelException;
import ai.djl.ModelException;
import ai.djl.inference.Predictor;
import ai.djl.modality.Classifications;
import ai.djl.modality.cv.*;
import ai.djl.modality.cv.output.*;
import ai.djl.modality.cv.transform.*;
import ai.djl.modality.cv.translator.ImageClassificationTranslator;
import ai.djl.modality.cv.util.NDImageUtils;
import ai.djl.ndarray.*;
import ai.djl.ndarray.types.Shape;
import ai.djl.repository.zoo.*;
import ai.djl.translate.*;
import java.io.IOException;
import java.nio.file.*;
import java.util.*;
```
## Face Detection model
Now we can start working on the first model. The model can do face detection and require some additional processing before we feed into it:
- Resize: Shrink the image with a certain ratio to feed in
- Normalize the image with a scale
Fortunatly, DJL offers a `Translator` interface that can help you with these processing. The rough Translator architecture looks like below:
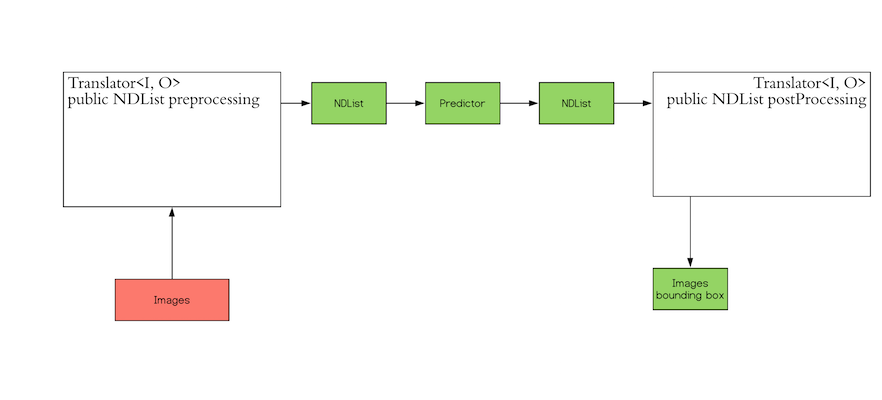
In the following sections, we will implement a `FaceTranslator` class to do the work.
### Preprocessing
In this stage, we will load an image and do some preprocessing work to it. Let's load the image first and take a look at it:
```
String url = "https://raw.githubusercontent.com/PaddlePaddle/PaddleHub/release/v1.5/demo/mask_detection/python/images/mask.jpg";
Image img = ImageFactory.getInstance().fromUrl(url);
img.getWrappedImage();
```
Then, let's try to apply some transformation to it:
```
NDList processImageInput(NDManager manager, Image input, float shrink) {
NDArray array = input.toNDArray(manager);
Shape shape = array.getShape();
array = NDImageUtils.resize(
array, (int) (shape.get(1) * shrink), (int) (shape.get(0) * shrink));
array = array.transpose(2, 0, 1).flip(0); // HWC -> CHW BGR -> RGB
NDArray mean = manager.create(new float[] {104f, 117f, 123f}, new Shape(3, 1, 1));
array = array.sub(mean).mul(0.007843f); // normalization
array = array.expandDims(0); // make batch dimension
return new NDList(array);
}
processImageInput(NDManager.newBaseManager(), img, 0.5f);
```
As you can see above, we convert the image to a NDArray with shape following (number_of_batches, channel (RGB), height, width). This is the required input for the model to run object detection.
### Postprocessing
For postprocessing, The output is in shape of (number_of_boxes, (class_id, probability, xmin, ymin, xmax, ymax)). We can store them into the prebuilt DJL `DetectedObjects` classes for further processing. Let's assume we have an inference output of ((1, 0.99, 0.2, 0.4, 0.5, 0.8)) and try to draw this box out.
```
DetectedObjects processImageOutput(NDList list, List<String> className, float threshold) {
NDArray result = list.singletonOrThrow();
float[] probabilities = result.get(":,1").toFloatArray();
List<String> names = new ArrayList<>();
List<Double> prob = new ArrayList<>();
List<BoundingBox> boxes = new ArrayList<>();
for (int i = 0; i < probabilities.length; i++) {
if (probabilities[i] >= threshold) {
float[] array = result.get(i).toFloatArray();
names.add(className.get((int) array[0]));
prob.add((double) probabilities[i]);
boxes.add(
new Rectangle(
array[2], array[3], array[4] - array[2], array[5] - array[3]));
}
}
return new DetectedObjects(names, prob, boxes);
}
NDArray tempOutput = NDManager.newBaseManager().create(new float[]{1f, 0.99f, 0.1f, 0.1f, 0.2f, 0.2f}, new Shape(1, 6));
DetectedObjects testBox = processImageOutput(new NDList(tempOutput), Arrays.asList("Not Face", "Face"), 0.7f);
Image newImage = img.duplicate(Image.Type.TYPE_INT_ARGB);
newImage.drawBoundingBoxes(testBox);
newImage.getWrappedImage();
```
### Create Translator and run inference
After this step, you might understand how process and postprocess works in DJL. Now, let's do something real and put them together in a single piece:
```
class FaceTranslator implements Translator<Image, DetectedObjects> {
private float shrink;
private float threshold;
private List<String> className;
FaceTranslator(float shrink, float threshold) {
this.shrink = shrink;
this.threshold = threshold;
className = Arrays.asList("Not Face", "Face");
}
@Override
public DetectedObjects processOutput(TranslatorContext ctx, NDList list) {
return processImageOutput(list, className, threshold);
}
@Override
public NDList processInput(TranslatorContext ctx, Image input) {
return processImageInput(ctx.getNDManager(), input, shrink);
}
@Override
public Batchifier getBatchifier() {
return null;
}
}
```
To run inference with this model, we need to load the model from Paddle model zoo. To load a model in DJL, you need to specify a `Crieteria`. `Crieteria` is used identify where to load the model and which `Translator` should apply to it. Then, all we need to do is to get a `Predictor` from the model and use it to do inference:
```
Criteria<Image, DetectedObjects> criteria =
Criteria.builder()
.optApplication(Application.CV.OBJECT_DETECTION)
.setTypes(Image.class, DetectedObjects.class)
.optArtifactId("face_detection")
.optTranslator(new FaceTranslator(0.5f, 0.7f))
.optFilter("flavor", "server")
.build();
var model = ModelZoo.loadModel(criteria);
var predictor = model.newPredictor();
DetectedObjects inferenceResult = predictor.predict(img);
newImage = img.duplicate(Image.Type.TYPE_INT_ARGB);
newImage.drawBoundingBoxes(inferenceResult);
newImage.getWrappedImage();
```
As you can see above, it brings you three faces detections.
## Mask Classification model
So, once we have the image location ready, we can crop the image and feed it to the Mask Classification model for further processing.
### Crop the image
The output of the box location is a value from 0 - 1 that can be mapped to the actual box pixel location if we simply multiply by width/height. For better accuracy on the cropped image, we extend the detection box to square. Let's try to get a cropped image:
```
int[] extendSquare(
double xmin, double ymin, double width, double height, double percentage) {
double centerx = xmin + width / 2;
double centery = ymin + height / 2;
double maxDist = Math.max(width / 2, height / 2) * (1 + percentage);
return new int[] {
(int) (centerx - maxDist), (int) (centery - maxDist), (int) (2 * maxDist)
};
}
Image getSubImage(Image img, BoundingBox box) {
Rectangle rect = box.getBounds();
int width = img.getWidth();
int height = img.getHeight();
int[] squareBox =
extendSquare(
rect.getX() * width,
rect.getY() * height,
rect.getWidth() * width,
rect.getHeight() * height,
0.18);
return img.getSubimage(squareBox[0], squareBox[1], squareBox[2], squareBox[2]);
}
List<DetectedObjects.DetectedObject> faces = inferenceResult.items();
getSubImage(img, faces.get(2).getBoundingBox()).getWrappedImage();
```
### Prepare Translator and load the model
For the face classification model, we can use DJL prebuilt `ImageClassificationTranslator` with a few transformation. This Translator brings a basic image translation process and can be extended with additional standard processing steps. So in our case, we don't have to create another `Translator` and just leverage on this prebuilt one.
```
var criteria = Criteria.builder()
.optApplication(Application.CV.IMAGE_CLASSIFICATION)
.setTypes(Image.class, Classifications.class)
.optTranslator(
ImageClassificationTranslator.builder()
.addTransform(new Resize(128, 128))
.addTransform(new ToTensor()) // HWC -> CHW div(255)
.addTransform(
new Normalize(
new float[] {0.5f, 0.5f, 0.5f},
new float[] {1.0f, 1.0f, 1.0f}))
.addTransform(nd -> nd.flip(0)) // RGB -> GBR
.build())
.optArtifactId("mask_classification")
.optFilter("flavor", "server")
.build();
var classifyModel = ModelZoo.loadModel(criteria);
var classifier = classifyModel.newPredictor();
```
### Run inference
So all we need to do is to apply the previous implemented functions and apply them all together. We firstly crop the image and then use it for inference. After these steps, we create a new DetectedObjects with new Classification classes:
```
List<String> names = new ArrayList<>();
List<Double> prob = new ArrayList<>();
List<BoundingBox> rect = new ArrayList<>();
for (DetectedObjects.DetectedObject face : faces) {
Image subImg = getSubImage(img, face.getBoundingBox());
Classifications classifications = classifier.predict(subImg);
names.add(classifications.best().getClassName());
prob.add(face.getProbability());
rect.add(face.getBoundingBox());
}
newImage = img.duplicate(Image.Type.TYPE_INT_ARGB);
newImage.drawBoundingBoxes(new DetectedObjects(names, prob, rect));
newImage.getWrappedImage();
```
| github_jupyter |
# Q* Learning with FrozenLake 🕹️⛄ (DETERMINISTIC VERSION)
<br>
## This is not the tutorial version, but a deterministic version
In this Notebook, we'll implement an agent <b>that plays FrozenLake.</b>
<img src="frozenlake.png" alt="Frozen Lake"/>
The goal of this game is <b>to go from the starting state (S) to the goal state (G)</b> by walking only on frozen tiles (F) and avoid holes (H).However, the ice is slippery, <b>so you won't always move in the direction you intend.</b>
<br><br>
This version was made by [lukewys](https://github.com/lukewys)
# This is a notebook from [Deep Reinforcement Learning Course with Tensorflow](https://simoninithomas.github.io/Deep_reinforcement_learning_Course/)
<img src="https://raw.githubusercontent.com/simoninithomas/Deep_reinforcement_learning_Course/master/docs/assets/img/DRLC%20Environments.png" alt="Deep Reinforcement Course"/>
<br>
<p> Deep Reinforcement Learning Course is a free series of articles and videos tutorials 🆕 about Deep Reinforcement Learning, where **we'll learn the main algorithms (Q-learning, Deep Q Nets, Dueling Deep Q Nets, Policy Gradients, A2C, Proximal Policy Gradients…), and how to implement them with Tensorflow.**
<br><br>
📜The articles explain the architectures from the big picture to the mathematical details behind them.
<br>
📹 The videos explain how to build the agents with Tensorflow </b></p>
<br>
This course will give you a **solid foundation for understanding and implementing the future state of the art algorithms**. And, you'll build a strong professional portfolio by creating **agents that learn to play awesome environments**: Doom© 👹, Space invaders 👾, Outrun, Sonic the Hedgehog©, Michael Jackson’s Moonwalker, agents that will be able to navigate in 3D environments with DeepMindLab (Quake) and able to walk with Mujoco.
<br><br>
</p>
## 📚 The complete [Syllabus HERE](https://simoninithomas.github.io/Deep_reinforcement_learning_Course/)
## Any questions 👨💻
<p> If you have any questions, feel free to ask me: </p>
<p> 📧: <a href="mailto:hello@simoninithomas.com">hello@simoninithomas.com</a> </p>
<p> Github: https://github.com/simoninithomas/Deep_reinforcement_learning_Course </p>
<p> 🌐 : https://simoninithomas.github.io/Deep_reinforcement_learning_Course/ </p>
<p> Twitter: <a href="https://twitter.com/ThomasSimonini">@ThomasSimonini</a> </p>
<p> Don't forget to <b> follow me on <a href="https://twitter.com/ThomasSimonini">twitter</a>, <a href="https://github.com/simoninithomas/Deep_reinforcement_learning_Course">github</a> and <a href="https://medium.com/@thomassimonini">Medium</a> to be alerted of the new articles that I publish </b></p>
## How to help 🙌
3 ways:
- **Clap our articles and like our videos a lot**:Clapping in Medium means that you really like our articles. And the more claps we have, the more our article is shared Liking our videos help them to be much more visible to the deep learning community.
- **Share and speak about our articles and videos**: By sharing our articles and videos you help us to spread the word.
- **Improve our notebooks**: if you found a bug or **a better implementation** you can send a pull request.
<br>
## Important note 🤔
<b> You can run it on your computer but it's better to run it on GPU based services</b>, personally I use Microsoft Azure and their Deep Learning Virtual Machine (they offer 170$)
https://azuremarketplace.microsoft.com/en-us/marketplace/apps/microsoft-ads.dsvm-deep-learning
<br>
⚠️ I don't have any business relations with them. I just loved their excellent customer service.
If you have some troubles to use Microsoft Azure follow the explainations of this excellent article here (without last the part fast.ai): https://medium.com/@manikantayadunanda/setting-up-deeplearning-machine-and-fast-ai-on-azure-a22eb6bd6429
## Prerequisites 🏗️
Before diving on the notebook **you need to understand**:
- The foundations of Reinforcement learning (MC, TD, Rewards hypothesis...) [Article](https://medium.freecodecamp.org/an-introduction-to-reinforcement-learning-4339519de419)
- Q-learning [Article](https://medium.freecodecamp.org/diving-deeper-into-reinforcement-learning-with-q-learning-c18d0db58efe)
- In the [video version](https://www.youtube.com/watch?v=q2ZOEFAaaI0) we implemented a Q-learning agent that learns to play OpenAI Taxi-v2 🚕 with Numpy.
```
from IPython.display import HTML
HTML('<iframe width="560" height="315" src="https://www.youtube.com/embed/q2ZOEFAaaI0?showinfo=0" frameborder="0" allow="autoplay; encrypted-media" allowfullscreen></iframe>')
```
## Step 0: Import the dependencies 📚
We use 3 libraries:
- `Numpy` for our Qtable
- `OpenAI Gym` for our FrozenLake Environment
- `Random` to generate random numbers
```
import numpy as np
import gym
import random
```
## Step 1: Create the environment 🎮
- Here we'll create the FrozenLake environment.
- OpenAI Gym is a library <b> composed of many environments that we can use to train our agents.</b>
- In our case we choose to use Frozen Lake.
```
from gym.envs.registration import register
register(
id='FrozenLakeNotSlippery-v0',
entry_point='gym.envs.toy_text:FrozenLakeEnv',
kwargs={'map_name' : '4x4', 'is_slippery': False},
max_episode_steps=100,
reward_threshold=0.8196, # optimum = .8196, changing this seems have no influence
)
env = gym.make("FrozenLakeNotSlippery-v0")
#env = gym.make("FrozenLake-v0")
```
## Step 2: Create the Q-table and initialize it 🗄️
- Now, we'll create our Q-table, to know how much rows (states) and columns (actions) we need, we need to calculate the action_size and the state_size
- OpenAI Gym provides us a way to do that: `env.action_space.n` and `env.observation_space.n`
```
action_size = env.action_space.n
state_size = env.observation_space.n
qtable = np.zeros((state_size, action_size))
```
## Step 3: Create the hyperparameters ⚙️
- Here, we'll specify the hyperparameters
```
total_episodes = 20000 # Total episodes
learning_rate = 0.8 # Learning rate
max_steps = 99 # Max steps per episode
gamma = 0.95 # Discounting rate
# Exploration parameters
epsilon = 1.0 # Exploration rate
max_epsilon = 1.0 # Exploration probability at start
min_epsilon = 0.01 # Minimum exploration probability
decay_rate = 0.001 # Exponential decay rate for exploration prob
#I find that decay_rate=0.001 works much better than 0.01
```
## Step 4: The Q learning algorithm 🧠
- Now we implement the Q learning algorithm:
<img src="qtable_algo.png" alt="Q algo"/>
```
# List of rewards
rewards = []
# 2 For life or until learning is stopped
for episode in range(total_episodes):
# Reset the environment
state = env.reset()
step = 0
done = False
total_rewards = 0
for step in range(max_steps):
# 3. Choose an action a in the current world state (s)
## First we randomize a number
exp_exp_tradeoff = random.uniform(0, 1)
## If this number > greater than epsilon --> exploitation (taking the biggest Q value for this state)
if exp_exp_tradeoff > epsilon:
action = np.argmax(qtable[state,:])
# Else doing a random choice --> exploration
else:
action = env.action_space.sample()
# Take the action (a) and observe the outcome state(s') and reward (r)
new_state, reward, done, info = env.step(action)
# Update Q(s,a):= Q(s,a) + lr [R(s,a) + gamma * max Q(s',a') - Q(s,a)]
# qtable[new_state,:] : all the actions we can take from new state
qtable[state, action] = qtable[state, action] + learning_rate * (reward + gamma * np.max(qtable[new_state, :]) - qtable[state, action])
total_rewards =total_rewards + reward
# Our new state is state
state = new_state
# If done (if we're dead) : finish episode
if done == True:
break
episode += 1
# Reduce epsilon (because we need less and less exploration)
epsilon = min_epsilon + (max_epsilon - min_epsilon)*np.exp(-decay_rate*episode)
rewards.append(total_rewards)
print ("Score over time: " + str(sum(rewards)/total_episodes))
print(qtable)
print(epsilon)
# Print the action in every place
#LEFT = 0 DOWN = 1 RIGHT = 2 UP = 3
env.reset()
env.render()
print(np.argmax(qtable,axis=1).reshape(4,4))
```
## Step 5: Use our Q-table to play FrozenLake ! 👾
- After 10 000 episodes, our Q-table can be used as a "cheatsheet" to play FrozenLake"
- By running this cell you can see our agent playing FrozenLake.
```
#All the episoded is the same
env.reset()
for episode in range(5):
state = env.reset()
step = 0
done = False
print("****************************************************")
print("EPISODE ", episode)
for step in range(max_steps):
env.render()
# Take the action (index) that have the maximum expected future reward given that state
action = np.argmax(qtable[state,:])
new_state, reward, done, info = env.step(action)
if done:
break
state = new_state
env.close()
```
| github_jupyter |
# ai4i2020 Data Visualization
- check characteristics of features and clases
- find correlation between features and classes
- with histogram
- corr()
- scatter_matrix
```
from google.colab import drive
drive.mount('/content/drive/')
cd drive/My\ Drive/Colab\ Notebooks/summer_coop
```
# Visualize ai4i2020 data from UCI
- [detail of the data](https://archive.ics.uci.edu/ml/datasets/AI4I+2020+Predictive+Maintenance+Dataset#)
```
import pandas as pd
data = pd.read_csv('ai4i2020/ai4i2020.csv')
data.head()
data.info()
data.describe()
import matplotlib.pyplot as plt
data.hist(bins = 50, figsize = (20, 15))
plt.ylabel('no. of value')
plt.xlabel('value')
plt.show()
```
## No. of Failure
- Some failure occur simultaneously
- Machine failure no : 339
- Sum of all failure : 373
```
print('Machine failure: \t' + str(data['Machine failure'].sum()))
print('Tool Wear failure: \t' + str(data['TWF'].sum()))
print('Heat Dissipation failure: \t' + str(data['HDF'].sum()))
print('Power failure: \t' + str(data['PWF'].sum()))
print('Overstrain failure: \t' + str(data['OSF'].sum()))
print('Random failure: \t' + str(data['RNF'].sum()))
print('SUM: \t' + str(data['TWF'].sum() + data['HDF'].sum() + data['PWF'].sum() + data['OSF'].sum() + data['RNF'].sum()))
```
## Features versus Machine failure
- can find from where the failure usually occur
- 1: failure
- 0: normal
### Result
- Air temperature: near 302 K
- Process temerature: 310 ~ 312 K
- Rotational speed: under 1500 rpm
- Torque: 45 ~ 70 Nm
- Tool wear: uniformly distributed, concentrated near 200 ~ 250 min
```
fig, axs = plt.subplots(1, 5, figsize = (20, 5))
axs[0].scatter(x = data['Air temperature [K]'], y = data['Machine failure'], alpha = 0.1)
axs[0].set_xlabel('Air temperature [K]')
axs[1].scatter(x = data['Process temperature [K]'], y = data['Machine failure'], alpha = 0.1)
axs[1].set_xlabel('Process temperature [K]')
axs[2].scatter(x = data['Rotational speed [rpm]'], y = data['Machine failure'], alpha = 0.1)
axs[2].set_xlabel('Rotational speed [rpm]')
axs[3].scatter(x = data['Torque [Nm]'], y = data['Machine failure'], alpha = 0.1)
axs[3].set_xlabel('Torque [Nm]')
axs[4].scatter(x = data['Tool wear [min]'], y = data['Machine failure'], alpha = 0.1)
axs[4].set_xlabel('Tool wear [min]')
plt.show()
```
## Air temp versus Process temp
- distribution is similar
```
plt.scatter(x = data['Air temperature [K]'], y = data['Process temperature [K]'], alpha= 0.1)
data.corr()
from pandas.plotting import scatter_matrix
attri = ['Air temperature [K]', 'Process temperature [K]','Rotational speed [rpm]', 'Torque [Nm]', 'Tool wear [min]']
scatter_matrix(data[attri], figsize = (12, 8))
```
| github_jupyter |
```
#Copyright Amazon.com, Inc. or its affiliates. All Rights Reserved.
#SPDX-License-Identifier: MIT-0
#install additional libraries
!pip install nltk
!pip install jsonlines
!pip install pandarallel
!pip install tensorflow==2.1
!pip install --upgrade grpcio
!pip install --upgrade s3fs
#import libraries
import os
import json
import shutil
import tensorflow as tf
import pandas as pd
import numpy as np
import boto3
import sagemaker
import nltk
from search_utils import helpers, search_preprocessing
```
# 1. Deploy a SageMaker Endpoint
```
#Creating a sagemaker session
sagemaker_session = sagemaker.Session()
#We'll be using the sagemaker default bucket
#Feel free to change this to another bucket name and make sure it's the same across all four notebooks
bucket_name = sagemaker_session.default_bucket()
#Copy the glove_job_name, this was generated automatically in step 3 of the training notebook
glove_job_name = "<replace-with-glove-job-name>"
#Copy the training_job_name, this was generated automatically in step 4 of the training notebook
training_job_name = "<replace-with-training-job-name>"
sagemaker_session.create_model_from_job(training_job_name=training_job_name,
env={'INFERENCE_PREFERRED_MODE': 'embedding'})
endpoint_config_name = sagemaker_session.create_endpoint_config(name=training_job_name,
model_name=training_job_name,
initial_instance_count=1,
instance_type='ml.m4.xlarge')
#Specify the name of the endpoint
endpoint_name = "object2vec-embeddings"
sagemaker_session.create_endpoint(endpoint_name=endpoint_name, config_name=training_job_name, tags=None, wait=True)
```
# 2. Generate predictions using the SageMaker Endpoint
```
#Reading the data from S3 and loading the vocabulary
data = pd.read_csv(f"s3://{bucket_name}/search_knn_blog/data/processed_data/data.csv")
word_to_id = helpers.read_json_from_s3(bucket_name,\
f'search_knn_blog/sagemaker-runs/{glove_job_name}/vocab.json')
#Sample a few products from the overall catalog
sub_set = data.sample(10000)
descriptions = sub_set["processed_title"]
from sagemaker.predictor import json_serializer, json_deserializer
# define encode-decode format for inference data
predictor = sagemaker.predictor.RealTimePredictor(endpoint_name)
predictor.content_type = 'application/json'
predictor.serializer = json_serializer
predictor.deserializer = json_deserializer
tokenizer = nltk.tokenize.TreebankWordTokenizer()
def l2_normalize(v):
"""
This functions normalise the embeddings based on the L2 norm.
"""
norm = np.sqrt(np.sum(np.square(v)))
return v / norm
#This is the "enc_dim" parameter you have set in the training job hyperparameters of object2vec
#By default this value is set to 512 in the training notebook
embedding_size=512
all_embeddings, labels = [], []
for i, description in enumerate(descriptions):
if i%1000==0:
print(f"Processing product {i}/{len(descriptions)}")
enc_description = search_preprocessing.sentence_to_integers(description, tokenizer, word_to_id)
if len(enc_description) != 0:
payload = {"instances" : [{"in0": enc_description}]}
result = predictor.predict(payload)
embeddings = result["predictions"][0]["embeddings"]
embeddings = l2_normalize(embeddings)
labels.append(sub_set.iloc[i]["product_category"])
all_embeddings.append(embeddings)
else:
all_embeddings.append([0]*embedding_size)
labels.append(sub_set.iloc[i]["product_category"])
#Transforming predictions to "float64" numpy array
labels = np.array(labels, dtype="str")
X = []
for em_value in all_embeddings:
X.append(em_value)
embeddings = np.array(X)
embeddings.dtype = "float64"
print(embeddings.shape)
```
# 3. Visualise the embeddings using Tensorboard projector
```
#Create a directory for storing tensorboard logs
#If a directory exists (previous runs) make sure to clean it up
if os.path.isdir("../tensorboard_logs"):
shutil.rmtree("../tensorboard_logs")
os.mkdir("../tensorboard_logs")
else:
os.mkdir("../tensorboard_logs")
from tensorboard.plugins import projector
def register_embedding(embedding_tensor_name, metadata_path, logs_dir):
config = projector.ProjectorConfig()
embedding = config.embeddings.add()
embedding.tensor_name = embedding_tensor_name
embedding.metadata_path = metadata_path
projector.visualize_embeddings(logs_dir, config)
#Setting the tensorboard logs directory and additional variables
logs_dir = '../tensorboard_logs'
metadata_path = 'metadata.tsv'
embedding_tensor_name = 'embeddings'
EMBEDDINGS_FPATH = os.path.join(logs_dir, f'{embedding_tensor_name}.ckpt')
#Registering and saving the embeddings in the logs directory
tf.compat.v1.reset_default_graph()
register_embedding(embedding_tensor_name, metadata_path, logs_dir)
tf.compat.v1.disable_eager_execution()
tensor_embeddings = tf.Variable(embeddings, name=embedding_tensor_name)
tf_session = tf.compat.v1.InteractiveSession()
tf_session.run(tf.compat.v1.global_variables_initializer())
saver = tf.compat.v1.train.Saver()
saver.save(tf_session, os.path.join(logs_dir, f'{embedding_tensor_name}.ckpt'), 0)
tf_session.close()
#Saving the labels
with open(os.path.join(logs_dir, metadata_path), 'w') as f:
for label in labels:
f.write('{}\n'.format(label))
```
# 4. Open Tensorboard
# 5. Saving enriched data for Elasticsearch
```
str_all_embeddings = [str(list(e)) for e in all_embeddings]
sub_set["embeddings"]= str_all_embeddings
def row2dict(x):
x = x.dropna().to_dict()
for key in x:
x[key] = str(x[key])
return x
records_to_save = sub_set.apply(lambda x: row2dict(x), axis=1)
records_to_save = list(records_to_save.values)
with open("./data.json", "w") as write_file:
json.dump(records_to_save, write_file)
boto3.client("s3").upload_file("./data.json",
bucket_name, "search_knn_blog/data/enriched_data/data.json")
```
# 5. Delete the endpoint (Unless you plan to continue to notebook 4)
```
#Make sure you delete your endpoint when you're done making predictions
sagemaker_session.delete_endpoint(endpoint_name="object2vec-embeddings")
```
| github_jupyter |
```
import numpy as np
import pickle
class BlackJackSolution:
def __init__(self, lr=0.1, exp_rate=0.3):
self.player_Q_Values = {} # key: [(player_value, show_card, usable_ace)][action] = value
# initialise Q values | (12-21) x (1-10) x (True, False) x (1, 0) 400 in total
for i in range(12, 22):
for j in range(1, 11):
for k in [True, False]:
self.player_Q_Values[(i, j, k)] = {}
for a in [1, 0]:
if (i == 21) and (a == 0):
self.player_Q_Values[(i, j, k)][a] = 1
else:
self.player_Q_Values[(i, j, k)][a] = 0
self.player_state_action = []
self.state = (0, 0, False) # initial state
self.actions = [1, 0] # 1: HIT 0: STAND
self.end = False
self.lr = lr
self.exp_rate = exp_rate
# give card
@staticmethod
def giveCard():
# 1 stands for ace
c_list = list(range(1, 11)) + [10, 10, 10]
return np.random.choice(c_list)
def dealerPolicy(self, current_value, usable_ace, is_end):
if current_value > 21:
if usable_ace:
current_value -= 10
usable_ace = False
else:
return current_value, usable_ace, True
# HIT17
if current_value >= 17:
return current_value, usable_ace, True
else:
card = self.giveCard()
if card == 1:
if current_value <= 10:
return current_value+11, True, False
return current_value+1, usable_ace, False
else:
return current_value+card, usable_ace, False
def chooseAction(self):
# if current value <= 11, always hit
current_value = self.state[0]
if current_value <= 11:
return 1
if np.random.uniform(0, 1) <= self.exp_rate:
action = np.random.choice(self.actions)
# print("random action", action)
else:
# greedy action
v = -999
action = 0
for a in self.player_Q_Values[self.state]:
if self.player_Q_Values[self.state][a] > v:
action = a
v = self.player_Q_Values[self.state][a]
# print("greedy action", action)
return action
# one can only has 1 usable ace
# return next state
def playerNxtState(self, action):
current_value = self.state[0]
show_card = self.state[1]
usable_ace = self.state[2]
if action:
# action hit
card = self.giveCard()
if card == 1:
if current_value <= 10:
current_value += 11
usable_ace = True
else:
current_value += 1
else:
current_value += card
else:
# action stand
self.end = True
return (current_value, show_card, usable_ace)
if current_value > 21:
if usable_ace:
current_value -= 10
usable_ace = False
else:
self.end = True
return (current_value, show_card, usable_ace)
return (current_value, show_card, usable_ace)
def winner(self, player_value, dealer_value):
# player 1 | draw 0 | dealer -1
winner = 0
if player_value > 21:
if dealer_value > 21:
# draw
winner = 0
else:
winner = -1
else:
if dealer_value > 21:
winner = 1
else:
if player_value < dealer_value:
winner = -1
elif player_value > dealer_value:
winner = 1
else:
# draw
winner = 0
return winner
def _giveCredit(self, player_value, dealer_value):
reward = self.winner(player_value, dealer_value)
# backpropagate reward
for s in reversed(self.player_state_action):
state, action = s[0], s[1]
reward = self.player_Q_Values[state][action] + self.lr*(reward - self.player_Q_Values[state][action])
self.player_Q_Values[state][action] = round(reward, 3)
reward = np.max(list(self.player_Q_Values[state].values())) # Q-learning
def reset(self):
self.player_state_action = []
self.state = (0, 0, False) # initial state
self.end = False
def deal2cards(self, show=False):
# return value after 2 cards and usable ace
value, usable_ace = 0, False
cards = [self.giveCard(), self.giveCard()]
if 1 in cards:
value = sum(cards)+10
usable_ace = True
else:
value = sum(cards)
usable_ace = False
if show:
return value, usable_ace, cards[0]
else:
return value, usable_ace
def play(self, rounds=1000):
for i in range(rounds):
if i % 1000 == 0:
print("round", i)
# give 2 cards
dealer_value, d_usable_ace, show_card = self.deal2cards(show=True)
player_value, p_usable_ace = self.deal2cards(show=False)
self.state = (player_value, show_card, p_usable_ace)
print("init", self.state)
# judge winner after 2 cards
if player_value == 21 or dealer_value == 21:
# game end
# print("reach 21 in 2 cards: player value {} | dealer value {}".format(player_value, dealer_value))
next
else:
while True:
action = self.chooseAction() # state -> action
# print("current value {}, action {}".format(self.state[0], action))
if self.state[0] >= 12:
state_action_pair = [self.state, action]
# print(state_action_pair)
self.player_state_action.append(state_action_pair)
# update next state
self.state = self.playerNxtState(action)
if self.end:
break
# dealer's turn
is_end = False
while not is_end:
dealer_value, d_usable_ace, is_end = self.dealerPolicy(dealer_value, d_usable_ace, is_end)
# judge winner
# give reward and update Q value
player_value = self.state[0]
print("player value {} | dealer value {}".format(player_value, dealer_value))
self._giveCredit(player_value, dealer_value)
# print("player state action", self.player_state_action)
self.reset()
def savePolicy(self, file="policy"):
fw = open(file, 'wb')
pickle.dump(self.player_Q_Values, fw)
fw.close()
def loadPolicy(self, file="policy"):
fr = open(file,'rb')
self.player_Q_Values = pickle.load(fr)
fr.close()
# trained robot play against dealer
def playWithDealer(self, rounds=1000):
self.reset()
self.loadPolicy()
self.exp_rate = 0
result = np.zeros(3) # player [win, draw, lose]
for _ in range(rounds):
# hit 2 cards each
# give 2 cards
dealer_value, d_usable_ace, show_card = self.deal2cards(show=True)
player_value, p_usable_ace = self.deal2cards(show=False)
self.state = (player_value, show_card, p_usable_ace)
# judge winner after 2 cards
if player_value == 21 or dealer_value == 21:
if player_value == dealer_value:
result[1] += 1
elif player_value > dealer_value:
result[0] += 1
else:
result[2] += 1
else:
# player's turn
while True:
action = self.chooseAction()
# update next state
self.state = self.playerNxtState(action)
if self.end:
break
# dealer's turn
is_end = False
while not is_end:
dealer_value, d_usable_ace, is_end = self.dealerPolicy(dealer_value, d_usable_ace, is_end)
# judge
player_value = self.state[0]
# print("player value {} | dealer value {}".format(player_value, dealer_value))
w = self.winner(player_value, dealer_value)
if w == 1:
result[0] += 1
elif w == 0:
result[1] += 1
else:
result[2] += 1
self.reset()
return result
b = BlackJackSolution()
b.play(1)
# winner test
for _ in range(10):
p_value = np.random.choice(range(12, 30))
d_value = np.random.choice(range(12, 30))
r = b.winner(p_value, d_value)
print(p_value, d_value, r)
# test dealer policy
for _ in range(10):
print("------------------")
dealer_value, d_usable_ace, is_end = 0, False, False
while not is_end:
dealer_value, d_usable_ace, is_end = b.dealerPolicy(dealer_value, d_usable_ace, is_end)
print(dealer_value, d_usable_ace)
# test deal2cards
i, j = 0, 0
for _ in range(1000):
p, _ = b.deal2cards()
d, _ = b.deal2cards()
if p == 21:
i += 1
if d == 21:
j += 1
print(i, j)
# test next state
b = BlackJackSolution()
b.state = (19, 10, True)
print(b.playerNxtState(action=1))
print(b.end)
b = BlackJackSolution()
b.state = (11, 10, True)
print(b.playerNxtState(action=1))
print(b.end)
# test play
b = BlackJackSolution()
b.play(10)
```
### Play
```
b = BlackJackSolution(exp_rate=0.2, lr=0.1)
b.play(10000)
b.savePolicy()
b.playWithDealer(10000)
for k, v in b.player_Q_Values.items():
actions = b.player_Q_Values.get(k)
action = max(actions.keys(), key=lambda k: actions[k])
action = "HIT" if action == 1 else "STAND"
print(k, action)
b.player_Q_Values.get((12, 6, True))
```
#### Play with same strategy
```
q_values = {}
for i in range(12, 22):
for j in range(1, 11):
for k in [True, False]:
q_values[(i, j, k)] = {}
for a in [1, 0]:
if i >= 17 and a == 0:
q_values[(i, j, k)][a] = 1
elif i < 17 and a == 1:
q_values[(i, j, k)][a] = 1
else:
q_values[(i, j, k)][a] = 0
b = BlackJackSolution()
b.player_Q_Values = q_values
b.playWithDealer(rounds=10000)
```
| github_jupyter |
# Predict MPG for vehicles in this dataset
```
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn.linear_model import LassoCV
from sklearn.preprocessing import StandardScaler
from sklearn.pipeline import Pipeline
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_absolute_error, r2_score, mean_squared_error, mean_absolute_percentage_error
df = pd.read_csv('data/clean_auto_mpg.csv')
print(df.describe())
df.head()
list(df.columns)
df.drop(labels=['Unnamed: 0', 'car name'],axis=1, inplace=True)
```
## Feature engineering
```
df['power_weight_ratio'] = df['horsepower']/df['weight']
df['displacement_per_cylinder'] = df['displacement']/df['cylinders']
df['power_weight'] = df['horsepower']*df['weight']
df['displacement_per_cylinder_acceleration'] = (df['displacement']/df['cylinders'])*df['acceleration']
df['displacement_power_weight'] = df['horsepower']*df['weight']*df['displacement']
```
## Exploratory Data analysis
```
sns.distplot(df['mpg'])
for col in df.columns:
sns.scatterplot(x=col,y='mpg', data=df)
plt.show()
df.corr('kendall')
list(df.columns)
```
## Model 3
```
y= df['mpg']
df.drop(['mpg','weight','horsepower','cylinders','displacement','power_weight_ratio',
'model year','power_weight','displacement_per_cylinder_acceleration'], axis=1, inplace=True)
X_train, X_test, y_train, y_test = train_test_split(df, y, test_size=0.33, random_state=42)
pipe = Pipeline(steps=[('scaler', StandardScaler(),),('lasso',LassoCV(),)])
pipe.fit(X_train, y_train)
prediction = pipe.predict(X_test)
mae = mean_absolute_error(y_test, prediction)
r2 =r2_score(y_test, prediction)
mse =mean_squared_error(y_test, prediction)
mape =mean_absolute_percentage_error(y_test, prediction)
print(r2)
print(mae)
print(mse)
print(mape)
pipe.steps[1][1].coef_
X_train.columns
pd.DataFrame(pipe.steps[1][1].coef_, index=X_train.columns,columns=['Coefficient'])
```
## Model 2
```
pipe2 = Pipeline(steps=[('scaler', StandardScaler(),),('lasso',LassoCV(),)])
X_train2 = X_train.drop(['displacement','horsepower',], axis=1)
X_test2 = X_test.drop(['displacement','horsepower',], axis=1)
pipe2.fit(X_train2, y_train)
prediction2 = pipe.predict(X_test2)
mae_2 = mean_absolute_error(y_test, prediction2)
r2_2 =r2_score(y_test, prediction2)
mse_2 =mean_squared_error(y_test, prediction2)
mape_2 =mean_absolute_percentage_error(y_test, prediction2)
r2_2
mae_2
mse_2
mape_2
pipe2.steps[1][1].coef_
pd.DataFrame(pipe2.steps[1][1].coef_, index=X_train2.columns,columns=['Coefficient'])
```
| github_jupyter |
```
# Copyright 2019 Google Inc. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ==============================================================================
```
# Local development and docker image components
- This section assumes that you have already created a program to perform the task required in a particular step of your ML workflow. This example uses an MNIST model training script.
- Then, this example packages your program as a Docker container image.
- Then, this example calls kfp.components.ContainerOp to convert it to a Kubeflow pipeline component.
Note: Ensure that you have Docker installed, if you want to build the image locally, by running the following command:
`which docker`
The result should be something like:
`/usr/bin/docker`
```
import kfp
import kfp.gcp as gcp
import kfp.dsl as dsl
import kfp.compiler as compiler
import kfp.components as comp
import datetime
import kubernetes as k8s
# Required Parameters
PROJECT_ID='<ADD GCP PROJECT HERE>'
GCS_BUCKET='gs://<ADD STORAGE LOCATION HERE>'
```
## Create client
If you run this notebook **outside** of a Kubeflow cluster, run the following command:
- `host`: The URL of your Kubeflow Pipelines instance, for example "https://`<your-deployment>`.endpoints.`<your-project>`.cloud.goog/pipeline"
- `client_id`: The client ID used by Identity-Aware Proxy
- `other_client_id`: The client ID used to obtain the auth codes and refresh tokens.
- `other_client_secret`: The client secret used to obtain the auth codes and refresh tokens.
```python
client = kfp.Client(host, client_id, other_client_id, other_client_secret)
```
If you run this notebook **within** a Kubeflow cluster, run the following command:
```python
client = kfp.Client()
```
You'll need to create OAuth client ID credentials of type `Other` to get `other_client_id` and `other_client_secret`. Learn more about [creating OAuth credentials](
https://cloud.google.com/iap/docs/authentication-howto#authenticating_from_a_desktop_app)
```
# Optional Parameters, but required for running outside Kubeflow cluster
# The host for 'AI Platform Pipelines' ends with 'pipelines.googleusercontent.com'
# The host for pipeline endpoint of 'full Kubeflow deployment' ends with '/pipeline'
# Examples are:
# https://7c021d0340d296aa-dot-us-central2.pipelines.googleusercontent.com
# https://kubeflow.endpoints.kubeflow-pipeline.cloud.goog/pipeline
HOST = '<ADD HOST NAME TO TALK TO KUBEFLOW PIPELINE HERE>'
# For 'full Kubeflow deployment' on GCP, the endpoint is usually protected through IAP, therefore the following
# will be needed to access the endpoint.
CLIENT_ID = '<ADD OAuth CLIENT ID USED BY IAP HERE>'
OTHER_CLIENT_ID = '<ADD OAuth CLIENT ID USED TO OBTAIN AUTH CODES HERE>'
OTHER_CLIENT_SECRET = '<ADD OAuth CLIENT SECRET USED TO OBTAIN AUTH CODES HERE>'
# This is to ensure the proper access token is present to reach the end point for 'AI Platform Pipelines'
# If you are not working with 'AI Platform Pipelines', this step is not necessary
! gcloud auth print-access-token
# Create kfp client
in_cluster = True
try:
k8s.config.load_incluster_config()
except:
in_cluster = False
pass
if in_cluster:
client = kfp.Client()
else:
if HOST.endswith('googleusercontent.com'):
CLIENT_ID = None
OTHER_CLIENT_ID = None
OTHER_CLIENT_SECRET = None
client = kfp.Client(host=HOST,
client_id=CLIENT_ID,
other_client_id=OTHER_CLIENT_ID,
other_client_secret=OTHER_CLIENT_SECRET)
```
## Wrap an existing Docker container image using `ContainerOp`
### Writing the program code
The following cell creates a file `app.py` that contains a Python script. The script downloads MNIST dataset, trains a Neural Network based classification model, writes the training log and exports the trained model to Google Cloud Storage.
Your component can create outputs that the downstream components can use as inputs. Each output must be a string and the container image must write each output to a separate local text file. For example, if a training component needs to output the path of the trained model, the component writes the path into a local file, such as `/output.txt`.
```
%%bash
# Create folders if they don't exist.
mkdir -p tmp/components/mnist_training
# Create the Python file that lists GCS blobs.
cat > ./tmp/components/mnist_training/app.py <<HERE
import argparse
from datetime import datetime
import tensorflow as tf
parser = argparse.ArgumentParser()
parser.add_argument(
'--model_file', type=str, required=True, help='Name of the model file.')
parser.add_argument(
'--bucket', type=str, required=True, help='GCS bucket name.')
args = parser.parse_args()
bucket=args.bucket
model_file=args.model_file
model = tf.keras.models.Sequential([
tf.keras.layers.Flatten(input_shape=(28, 28)),
tf.keras.layers.Dense(512, activation=tf.nn.relu),
tf.keras.layers.Dropout(0.2),
tf.keras.layers.Dense(10, activation=tf.nn.softmax)
])
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
print(model.summary())
mnist = tf.keras.datasets.mnist
(x_train, y_train),(x_test, y_test) = mnist.load_data()
x_train, x_test = x_train / 255.0, x_test / 255.0
callbacks = [
tf.keras.callbacks.TensorBoard(log_dir=bucket + '/logs/' + datetime.now().date().__str__()),
# Interrupt training if val_loss stops improving for over 2 epochs
tf.keras.callbacks.EarlyStopping(patience=2, monitor='val_loss'),
]
model.fit(x_train, y_train, batch_size=32, epochs=5, callbacks=callbacks,
validation_data=(x_test, y_test))
model.save(model_file)
from tensorflow import gfile
gcs_path = bucket + "/" + model_file
if gfile.Exists(gcs_path):
gfile.Remove(gcs_path)
gfile.Copy(model_file, gcs_path)
with open('/output.txt', 'w') as f:
f.write(gcs_path)
HERE
```
### Creating a Dockerfile
Now create a container that runs the script. Start by creating a Dockerfile. A Dockerfile contains the instructions to assemble a Docker image. The `FROM` statement specifies the Base Image from which you are building. `WORKDIR` sets the working directory. When you assemble the Docker image, `COPY` copies the required files and directories (for example, `app.py`) to the file system of the container. `RUN` executes a command (for example, install the dependencies) and commits the results.
```
%%bash
# Create Dockerfile.
cat > ./tmp/components/mnist_training/Dockerfile <<EOF
FROM tensorflow/tensorflow:1.15.0-py3
WORKDIR /app
COPY . /app
EOF
```
### Build docker image
Now that we have created our Dockerfile for creating our Docker image. Then we need to build the image and push to a registry to host the image. There are three possible options:
- Use the `kfp.containers.build_image_from_working_dir` to build the image and push to the Container Registry (GCR). This requires [kaniko](https://cloud.google.com/blog/products/gcp/introducing-kaniko-build-container-images-in-kubernetes-and-google-container-builder-even-without-root-access), which will be auto-installed with 'full Kubeflow deployment' but not 'AI Platform Pipelines'.
- Use [Cloud Build](https://cloud.google.com/cloud-build), which would require the setup of GCP project and enablement of corresponding API. If you are working with GCP 'AI Platform Pipelines' with GCP project running, it is recommended to use Cloud Build.
- Use [Docker](https://www.docker.com/get-started) installed locally and push to e.g. GCR.
**Note**:
If you run this notebook **within Kubeflow cluster**, **with Kubeflow version >= 0.7** and exploring **kaniko option**, you need to ensure that valid credentials are created within your notebook's namespace.
- With Kubeflow version >= 0.7, the credential is supposed to be copied automatically while creating notebook through `Configurations`, which doesn't work properly at the time of creating this notebook.
- You can also add credentials to the new namespace by either [copying credentials from an existing Kubeflow namespace, or by creating a new service account](https://www.kubeflow.org/docs/gke/authentication/#kubeflow-v0-6-and-before-gcp-service-account-key-as-secret).
- The following cell demonstrates how to copy the default secret to your own namespace.
```bash
%%bash
NAMESPACE=<your notebook name space>
SOURCE=kubeflow
NAME=user-gcp-sa
SECRET=$(kubectl get secrets \${NAME} -n \${SOURCE} -o jsonpath="{.data.\${NAME}\.json}" | base64 -D)
kubectl create -n \${NAMESPACE} secret generic \${NAME} --from-literal="\${NAME}.json=\${SECRET}"
```
```
IMAGE_NAME="mnist_training_kf_pipeline"
TAG="latest" # "v_$(date +%Y%m%d_%H%M%S)"
GCR_IMAGE="gcr.io/{PROJECT_ID}/{IMAGE_NAME}:{TAG}".format(
PROJECT_ID=PROJECT_ID,
IMAGE_NAME=IMAGE_NAME,
TAG=TAG
)
APP_FOLDER='./tmp/components/mnist_training/'
# In the following, for the purpose of demonstration
# Cloud Build is choosen for 'AI Platform Pipelines'
# kaniko is choosen for 'full Kubeflow deployment'
if HOST.endswith('googleusercontent.com'):
# kaniko is not pre-installed with 'AI Platform Pipelines'
import subprocess
# ! gcloud builds submit --tag ${IMAGE_NAME} ${APP_FOLDER}
cmd = ['gcloud', 'builds', 'submit', '--tag', GCR_IMAGE, APP_FOLDER]
build_log = (subprocess.run(cmd, stdout=subprocess.PIPE).stdout[:-1].decode('utf-8'))
print(build_log)
else:
if kfp.__version__ <= '0.1.36':
# kfp with version 0.1.36+ introduce broken change that will make the following code not working
import subprocess
builder = kfp.containers._container_builder.ContainerBuilder(
gcs_staging=GCS_BUCKET + "/kfp_container_build_staging"
)
kfp.containers.build_image_from_working_dir(
image_name=GCR_IMAGE,
working_dir=APP_FOLDER,
builder=builder
)
else:
raise("Please build the docker image use either [Docker] or [Cloud Build]")
```
#### If you want to use docker to build the image
Run the following in a cell
```bash
%%bash -s "{PROJECT_ID}"
IMAGE_NAME="mnist_training_kf_pipeline"
TAG="latest" # "v_$(date +%Y%m%d_%H%M%S)"
# Create script to build docker image and push it.
cat > ./tmp/components/mnist_training/build_image.sh <<HERE
PROJECT_ID="${1}"
IMAGE_NAME="${IMAGE_NAME}"
TAG="${TAG}"
GCR_IMAGE="gcr.io/\${PROJECT_ID}/\${IMAGE_NAME}:\${TAG}"
docker build -t \${IMAGE_NAME} .
docker tag \${IMAGE_NAME} \${GCR_IMAGE}
docker push \${GCR_IMAGE}
docker image rm \${IMAGE_NAME}
docker image rm \${GCR_IMAGE}
HERE
cd tmp/components/mnist_training
bash build_image.sh
```
```
image_name = GCR_IMAGE
```
### Define each component
Define a component by creating an instance of `kfp.dsl.ContainerOp` that describes the interactions with the Docker container image created in the previous step. You need to specify
- component name
- the image to use
- the command to run after the container starts (If None, uses default CMD in defined in container.)
- the input arguments
- the file outputs (In the `app.py` above, the path of the trained model is written to `/output.txt`.)
```
def mnist_train_op(model_file, bucket):
return dsl.ContainerOp(
name="mnist_training_container",
image='gcr.io/{}/mnist_training_kf_pipeline:latest'.format(PROJECT_ID),
command=['python', '/app/app.py'],
file_outputs={'outputs': '/output.txt'},
arguments=['--bucket', bucket, '--model_file', model_file]
)
```
### Create your workflow as a Python function
Define your pipeline as a Python function. ` @kfp.dsl.pipeline` is a required decoration including `name` and `description` properties. Then compile the pipeline function. After the compilation is completed, a pipeline file is created.
```
# Define the pipeline
@dsl.pipeline(
name='Mnist pipeline',
description='A toy pipeline that performs mnist model training.'
)
def mnist_container_pipeline(
model_file: str = 'mnist_model.h5',
bucket: str = GCS_BUCKET
):
mnist_train_op(model_file=model_file, bucket=bucket).apply(gcp.use_gcp_secret('user-gcp-sa'))
```
### Submit a pipeline run
```
pipeline_func = mnist_container_pipeline
experiment_name = 'minist_kubeflow'
arguments = {"model_file":"mnist_model.h5",
"bucket":GCS_BUCKET}
run_name = pipeline_func.__name__ + ' run'
# Submit pipeline directly from pipeline function
run_result = client.create_run_from_pipeline_func(pipeline_func,
experiment_name=experiment_name,
run_name=run_name,
arguments=arguments)
```
**As an alternative, you can compile the pipeline into a package.** The compiled pipeline can be easily shared and reused by others to run the pipeline.
```python
pipeline_filename = pipeline_func.__name__ + '.pipeline.zip'
compiler.Compiler().compile(pipeline_func, pipeline_filename)
experiment = client.create_experiment('python-functions-mnist')
run_result = client.run_pipeline(
experiment_id=experiment.id,
job_name=run_name,
pipeline_package_path=pipeline_filename,
params=arguments)
```
| github_jupyter |
## Imports
```
%load_ext autoreload
%autoreload 2
%matplotlib inline
from fastai.imports import *
from fastai.structured import *
from pandas_summary import DataFrameSummary
PATH = os.getcwd()
train_df = pd.read_csv(f'{PATH}\\train.csv', low_memory=False)
test_df = pd.read_csv(f'{PATH}\\test.csv', low_memory=False)
def display_all(df):
with pd.option_context("display.max_rows", 100):
with pd.option_context("display.max_columns", 100):
display(df)
'''
**Problem Statement**
An online question and answer platform has hired you as a data scientist to identify the best question authors on the platform.
This identification will bring more insight into increasing the user engagement. Given the tag of the question, number of views
received, number of answers, username and reputation of the question author, the problem requires you to predict the upvote
count that the question will receive.
**DATA DICTIONARY**
- Variable ----------- Definition
- ID ----------- Question ID
- Tag ----------- Anonymised tags representing question category
- Reputation ----------- Reputation score of question author
- Answers ----------- Number of times question has been answered
- Username ----------- Anonymised user id of question author
- Views ----------- Number of times question has been viewed
- Upvotes (Target) ----------- Number of upvotes for the question
''';
train_df.head()
'Train', train_df.shape, len(set(train_df.ID.values)), len(set(train_df.Username.values)), 'Test', \
test_df.shape, len(set(test_df.ID.values)), len(set(test_df.Username.values))
def Intersection(lst1, lst2):
return len(list(set(lst1).intersection(lst2)))
```
## Username
```
Intersection(train_df.Username, test_df.Username),\
Intersection(train_df.Reputation, test_df.Reputation),\
Intersection(train_df.Views, test_df.Views)
```
First of all, let's see how many different User's we have on both datasets
```
man_train_list = train_df.Username.unique()
man_test_list = test_df.Username.unique()
print("Train: {0}".format(len(man_train_list)))
print("Test: {0}".format(len(man_test_list)))
```
- **Unique User's in Test are close to $1$/$2$ of user's in Train...**
```
temp1 = train_df.groupby('Username').count().iloc[:,-1]
temp2 = test_df.groupby('Username').count().iloc[:,-1]
df_man = pd.concat([temp1,temp2], axis = 1, join = 'outer')
df_man.columns = ['train_count','test_count']
print(df_man.head(20))
```
- **Some Users have entries only in one of the two datasets**
```
print(df_man.sort_values(by = 'train_count', ascending = False).head(20))
```
- **More clearer if one looks at the plots for the cumulative distributions.**
```
fig, axes = plt.subplots(1,2, figsize = (12,5))
temp = df_man['train_count'].dropna().sort_values(ascending = False).reset_index(drop = True)
axes[0].plot(temp.index+1, temp.cumsum()/temp.sum())
axes[0].set_title('cumulative train_count');
temp = df_man['test_count'].dropna().sort_values(ascending = False).reset_index(drop = True)
axes[1].plot(temp.index+1, temp.cumsum()/temp.sum())
axes[1].set_title('cumulative test_count');
ix20 = int(len(df_man['train_count'].dropna())*0.2)
print("TRAIN: 20% of man ({0}) responsible for {1:2.2f}% of entries".format(ix20,df_man['train_count'].sort_values(ascending = False).cumsum().iloc[ix20]/df_man['train_count'].sum()*100))
ix20 = int(len(df_man['test_count'].dropna())*0.2)
print("TEST: 20% of man ({0}) responsible for {1:2.2f}% of entries".format(ix20, df_man['test_count'].sort_values(ascending = False).cumsum().iloc[ix20]/df_man['test_count'].sum()*100))
```
- **The man featuring in only one of the two datasets usually have very few entries.**
```
man_not_in_test = set(man_train_list) - set(man_test_list)
man_not_in_train = set(man_test_list) - set(man_train_list)
print("{} man are featured in train but not in test".format(len(man_not_in_test)))
print("{} man are featured in test but not in train".format(len(man_not_in_train)))
train_df.loc[list(man_not_in_test)].head()
## Need to drop them blindly...
#train_df.drop(index = train_df.loc[list(man_not_in_test)].index, inplace=True).shape
print(df_man.loc[list(man_not_in_test)]['train_count'].describe())
print(df_man.loc[list(man_not_in_train)]['test_count'].describe())
```
- **Strong Correlation among them**
```
df_man.sort_values(by = 'train_count', ascending = False).head(1000).corr()
df_man.sort_values(by = 'train_count', ascending = False).plot.scatter(x = 'train_count', y = 'test_count')
temp = df_man['train_count'].sort_values(ascending = False).head(50000)
temp = pd.concat([temp,temp.cumsum()/df_man['train_count'].sum()*100], axis = 1).reset_index()
temp.columns = ['user_id','count','percentage']
print(temp)
set(train_df.Tag)
man_list = df_man['train_count'].sort_values(ascending = False).index
ixes = train_df.Username.isin(man_list)
df10000 = train_df[ixes][['Username','Tag']]
tags_dummies = pd.get_dummies(df10000.Tag)
df10000 = pd.concat([df10000,tags_dummies[['a', 'c', 'h', 'i', 'j', 'o', 'p', 'r', 's', 'x']]], axis = 1).drop('Tag', axis = 1)
print("The contributors account for {} entries\n".format(len(df10000)))
print(df10000.head(10))
df10000.head(2)
import itertools
last_names = ['Mary', 'Patricia', 'Linda', 'Barbara', 'Elizabeth',
'Jennifer', 'Maria', 'Susan', 'Margaret', 'Dorothy',
'James', 'John', 'Robert', 'Michael', 'William', 'David',
'Richard', 'Charles', 'Joseph', 'Thomas','Smith', 'Johnson', 'Williams', 'Jones', 'Brown', 'Davis', 'Miller', 'Wilson', 'Moore',
'Taylor', 'Anderson', 'Thomas', 'Jackson', 'White', 'Harris', 'Martin', 'Thompson', 'Garcia',
'Martinez', 'Robinson', 'Clark', 'Rodriguez', 'Lewis', 'Lee', 'Walker', 'Hall', 'Allen', 'Young',
'Hernandez', 'King', 'Wright', 'Lopez', 'Hill', 'Scott', 'Green', 'Adams', 'Baker', 'Gonzalez', 'Nelson',
'Carter', 'Mitchell', 'Perez', 'Roberts', 'Turner', 'Phillips', 'Campbell', 'Parker', 'Evans', 'Edwards', 'Collins']
first_names = ['Smith', 'Johnson', 'Williams', 'Jones', 'Brown', 'Davis', 'Miller', 'Wilson', 'Moore',
'Taylor', 'Anderson', 'Thomas', 'Jackson', 'White', 'Harris', 'Martin', 'Thompson', 'Garcia',
'Martinez', 'Robinson', 'Clark', 'Rodriguez', 'Lewis', 'Lee', 'Walker', 'Hall', 'Allen', 'Young',
'Hernandez', 'King', 'Wright', 'Lopez', 'Hill', 'Scott', 'Green', 'Adams', 'Baker', 'Gonzalez', 'Nelson',
'Carter', 'Mitchell', 'Perez', 'Roberts', 'Turner', 'Phillips', 'Campbell', 'Parker', 'Evans', 'Edwards', 'Collins','Mary', 'Patricia', 'Linda', 'Barbara', 'Elizabeth',
'Jennifer', 'Maria', 'Susan', 'Margaret', 'Dorothy',
'James', 'John', 'Robert', 'Michael', 'William', 'David',
'Richard', 'Charles', 'Joseph', 'Thomas']
names = [first + ' ' + last for first,last in (itertools.product(first_names, last_names))]
# shuffle them
np.random.seed(12345)
np.random.shuffle(names)
dictionary = dict(zip(man_list, names))
df10000.loc[df10000.Username.isin(dictionary), 'Username' ] = df10000['Username'].map(dictionary)
print(df10000.head())
# see if the name coincides
print(names[:10])
print(df10000.groupby('Username').count().sort_values(by = 'a', ascending = False).head(10))
gby = pd.concat([df10000.groupby('Username').mean(),df10000.groupby('Username').count()], axis = 1).iloc[:,:-9]
gby.columns = ['a', 'c', 'h', 'i', 'j', 'o', 'p', 'r', 's', 'x', 'count']
gby.sort_values(by = 'count', ascending = False).head(10)[['a', 'c', 'h', 'i', 'j', 'o', 'p', 'r', 's', 'x', 'count']]
```
- **Their performances seem very different, even for people with similar number of entries.**
```
gby.sort_values(by = 'count', ascending = False).head(100).drop('count', axis = 1).plot(kind = 'bar', stacked = True, figsize = (15,6))
plt.figure()
gby.sort_values(by = 'count', ascending = False)['count'].head(100).plot(kind = 'bar', figsize = (15,6));
```
I think this high diversity should be accounted for when building our predictive model!
It would be interesting to rank the man based on their interest levels(the tags). For instance, we could compute their "skill" by assigning 0 points for "a", 1 for "b" and 2 for "c"...
```
gby.head(2)
pd.concat([train_df['Tag'].value_counts().sort_values(ascending=False),test_df['Tag'].value_counts().sort_values(ascending=False)],sort=False, axis =1,\
keys=['Train_Stats', 'Test_Stats'])
gby['skill'] = gby['r']*1 + gby['o']*2 + gby['h']*3 + gby['s']*4 + gby['a']*5 + gby['i']*6 + gby['p']*7 + gby['j']*8 \
+ gby['c']*9
print("Top performers")
gby.sort_values(by = 'skill', ascending = False).reset_index().head()
print("\nWorst performers")
gby.sort_values(by = 'skill', ascending = False).reset_index().tail()
gby.skill.plot(kind = 'hist', bins=10)
print(gby.mean())
```
| github_jupyter |
# DocTable File Column Types
It is often good advice to avoid storing large binary data in an SQL table because it will significantly impact the read performance of the entire table. I find, however, that it can be extremely useful in text analysis applications as a way to keep track of a large number of models with associated metadata. As an alternative to storing binary data in the table directly, `DocTable` includes a number of custom column types that can transparently store data into the filesystem and keep track of it using the schema definitions.
I provide two file storage column types: (1) `TextFileCol` for storing text data, and (2) `PickleFileCol` for storing any python data that requires pickling.
```
import numpy as np
from pathlib import Path
import sys
sys.path.append('..')
import doctable
# automatically clean up temp folder after python ends
tmpfolder = doctable.TempFolder('tmp')
```
Now I create a new table representing a matrix. Notice that I use the `PickleFileCol` column shortcut to create the column. This column is equivalent to `Col(None, coltype='picklefile', type_args=dict(folder=folder))`. See that to SQLite, this column simply looks like a text column.
```
import dataclasses
@doctable.schema(require_slots=False)
class MatrixRow:
id: int = doctable.IDCol()
array: np.ndarray = doctable.PickleFileCol('tmp/matrix_pickle_files') # will store files in the tmp directory
db = doctable.DocTable(target='tmp/test.db', schema=MatrixRow, new_db=True)
db.schema_info()
```
Now we insert a new array. It appears to be inserted the same as any other object.
```
db.insert({'array': np.random.rand(10,10)})
db.insert({'array': np.random.rand(10,10)})
print(db.count())
db.select_df(limit=3)
```
But when we actually look at the filesystem, we see that files have been created to store the array.
```
for fpath in tmpfolder.path.rglob('*.pic'):
print(str(fpath))
```
If we want to see the raw data stored in the table, we can create a new doctable without a defined schema. See that the raw filenames have been stored in the database. Recall that the directory indicating where to find these files was provided in the schema itself.
```
vdb = doctable.DocTable('tmp/test.db')
print(vdb.count())
vdb.head()
```
## Data Folder Consistency
Now we try to delete a row from the database. We can see that it was deleted as expected.
```
db.delete(where=db['id']==1)
print(db.count())
db.head()
```
However, when we check the folder where the data was stored, we find that the file was, in fact, not deleted. This is the case for technical reasons.
```
for fpath in tmpfolder.path.rglob('*.pic'):
print(str(fpath))
```
We can clean up the unused files using `clean_col_files()` though. Note that the specific column to clean must be provided.
```
db.clean_col_files('array')
for fpath in tmpfolder.path.rglob('*.pic'):
print(str(fpath))
```
There may be a situation where doctable cannot find the folder associated with an existing row. We can also use `clean_col_files()` to check for missing data. This might most frequently occur when the wrong folder is specified in the schema after moving the data file folder. For example, we delete all the pickle files in the directory and then run `clean_col_files()`.
```
[fp.unlink() for fp in tmpfolder.path.rglob('*.pic')]
for fpath in tmpfolder.path.rglob('*.pic'):
print(str(fpath))
# see that the exception was raised
try:
db.clean_col_files('array')
except FileNotFoundError as e:
print(e)
```
## Text File Types
We can also store text files in a similar way. For this, use `TextFileCol` in the folder specification.
```
@doctable.schema(require_slots=False)
class TextFileRow:
id: int = doctable.IDCol()
text: str = doctable.TextFileCol('tmp/my_text_files') # will store files in the tmp directory
tdb = doctable.DocTable(target='tmp/test_textfiles.db', schema=TextFileRow, new_db=True)
tdb.insert({'text': 'Hello world. DocTable is the most useful python package of all time.'})
tdb.insert({'text': 'Star Wars is my favorite movie.'})
tdb.head()
# and they look like text files
vdb = doctable.DocTable('tmp/test_textfiles.db')
print(vdb.count())
vdb.head()
```
See that the text files were created, and they look like normal text files so we can read them normally.
```
for fpath in tmpfolder.path.rglob('*.txt'):
print(f"{fpath}: {fpath.read_text()}")
```
| github_jupyter |
```
import pandas as pd
from google.colab import files
uploaded = files.upload()
dados_drugbank = pd.read_csv('drugbank-human (1).csv')
dados_drugbank.head()
dados_drugbank.shape
dados_drugbank.head()
dados_drugbank.shape
cod_drugbankgenes= dados_drugbank['Symbol'].value_counts().index[0:10]
cod_drugbankgenes
dados_drugbank.query('Symbol in @cod_drugbankgenes')
import seaborn as sns
import matplotlib.pyplot as plt
sns.set()
plt.figure(figsize=(35, 12))
ax = sns.countplot(x = 'Symbol', data=dados_drugbank.query('Symbol in @cod_drugbankgenes'), order= dados_drugbank['Symbol'].value_counts().index[0:10], palette='husl')
ax.set_title('Top genes')
plt.title("Top genes", fontsize = 20)
plt.xlabel("genes", fontsize = 18)
plt.ylabel("drugs", fontsize = 18)
plt.show()
cod_drugs = dados_drugbank['DRUGBANK'].value_counts().index[0:10]
cod_drugs
dados_drugbank.query('DRUGBANK in @cod_drugs')
import seaborn as sns
import matplotlib.pyplot as plt
sns.set()
plt.figure(figsize=(35, 12))
ax = sns.countplot(x = 'DRUGBANK', data=dados_drugbank.query('DRUGBANK in @cod_drugs'), order= dados_drugbank['DRUGBANK'].value_counts().index[0:10], palette='husl')
ax.set_title('Top drugs')
plt.title("Top drugs", fontsize = 20)
plt.xlabel("Top drugs", fontsize = 18)
plt.ylabel("genes", fontsize = 18)
plt.show()
cod_gse= dados_drugbank['GSECODE'].value_counts().index[0:10]
cod_gse
dados_drugbank.query('GSECODE in @cod_gse')
import seaborn as sns
import matplotlib.pyplot as plt
sns.set()
plt.figure(figsize=(35, 12))
ax = sns.countplot(x = 'GSECODE', data=dados_drugbank.query('GSECODE in @cod_gse'), order= dados_drugbank['GSECODE'].value_counts().index[0:10], palette='husl')
ax.set_title('Top GSECODE')
plt.title("Top GSECODE", fontsize = 20)
plt.xlabel("Top GSECODE", fontsize = 18)
plt.ylabel("count", fontsize = 18)
plt.show()
cod_mps = dados_drugbank['MPS_TYPE'].value_counts().index[0:10]
cod_mps
dados_drugbank.query('MPS_TYPE in @cod_mps')
import seaborn as sns
import matplotlib.pyplot as plt
sns.set()
plt.figure(figsize=(35, 12))
ax = sns.countplot(x = 'MPS_TYPE', data=dados_drugbank.query('MPS_TYPE in @cod_mps'), order= dados_drugbank['MPS_TYPE'].value_counts().index[0:10], palette='husl')
ax.set_title('Top MPS_TYPE')
plt.title("Top MPS_TYPE", fontsize = 20)
plt.xlabel("Top MPS_TYPE", fontsize = 18)
plt.ylabel("count", fontsize = 18)
plt.show()
cod_moa = dados_drugbank['MOA'].value_counts().index[0:10]
cod_moa
dados_drugbank.query('MOA in @cod_moa')
import seaborn as sns
import matplotlib.pyplot as plt
sns.set()
plt.figure(figsize=(75, 20))
ax = sns.countplot(x = 'MOA', data=dados_drugbank.query('MOA in @cod_moa'), order= dados_drugbank['MOA'].value_counts().index[0:10], palette='husl')
ax.set_title('Top MOA')
plt.title("Top MOA", fontsize = 20)
plt.xlabel("Top MOA", fontsize = 18)
plt.ylabel("count", fontsize = 18)
plt.show()
dados_drugbank.describe()
```
| github_jupyter |
# 01 - Simple ES Benchmark Function
### [Last Update: March 2022][](https://colab.research.google.com/github/RobertTLange/evosax/blob/main/examples/01_classic_benchmark.ipynb)
```
%matplotlib inline
%load_ext autoreload
%autoreload 2
%config InlineBackend.figure_format = 'retina'
!pip install git+https://github.com/RobertTLange/evosax.git@main
```
## 2D Rosenbrock with CMA-ES
`evosax` implements a set of different classic benchmark functions. These include multi-dimensional versions of `quadratic`, `rosenbrock`, `ackley`, `griewank`, `rastrigin`, `schwefel`, `himmelblau`, `six-hump`. In the following we focus on the 2D Rosenbrock case, but feel free to play around with the others.
```
import jax
import jax.numpy as jnp
from evosax import CMA_ES
from evosax.problems import ClassicFitness
# Instantiate the problem evaluator
rosenbrock = ClassicFitness("rosenbrock", num_dims=2)
# Instantiate the search strategy
rng = jax.random.PRNGKey(0)
strategy = CMA_ES(popsize=20, num_dims=2, elite_ratio=0.5)
es_params = strategy.default_params
es_params["init_min"] = -0
es_params["init_max"] = 0
state = strategy.initialize(rng, es_params)
# Run ask-eval-tell loop - NOTE: By default minimization
for t in range(50):
rng, rng_gen, rng_eval = jax.random.split(rng, 3)
x, state = strategy.ask(rng_gen, state, es_params)
fitness = rosenbrock.rollout(rng_eval, x)
state = strategy.tell(x, fitness, state, es_params)
if (t + 1) % 10 == 0:
print("CMA-ES - # Gen: {}|Fitness: {:.2f}|Params: {}".format(
t+1, state["best_fitness"], state["best_member"]))
```
## 2D Rosenbrock with Other ES
```
from evosax import Strategies
rng = jax.random.PRNGKey(0)
for s_name in ["SimpleES", "SimpleGA", "PSO", "DE", "Sep_CMA_ES",
"Full_iAMaLGaM", "Indep_iAMaLGaM", "MA_ES", "LM_MA_ES",
"RmES", "GLD", "SimAnneal"]:
strategy = Strategies[s_name](popsize=20, num_dims=2)
es_params = strategy.default_params
es_params["init_min"] = -2
es_params["init_max"] = 2
state = strategy.initialize(rng, es_params)
for t in range(30):
rng, rng_gen, rng_eval = jax.random.split(rng, 3)
x, state = strategy.ask(rng_gen, state, es_params)
fitness = rosenbrock.rollout(rng_eval, x)
state = strategy.tell(x, fitness, state, es_params)
if (t + 1) % 5 == 0:
print("{} - # Gen: {}|Fitness: {:.2f}|Params: {}".format(
s_name, t+1, state["best_fitness"], state["best_member"]))
print(20*"=")
```
# xNES on Sinusoidal Task
```
from evosax.strategies import xNES
def f(x):
"""Taken from https://github.com/chanshing/xnes"""
r = jnp.sum(x ** 2)
return -jnp.sin(r) / r
batch_func = jax.vmap(f, in_axes=0)
rng = jax.random.PRNGKey(0)
strategy = xNES(popsize=50, num_dims=2)
es_params = strategy.default_params
es_params["use_adaptive_sampling"] = True
es_params["use_fitness_shaping"] = True
es_params["eta_bmat"] = 0.01
es_params["eta_sigma"] = 0.1
state = strategy.initialize(rng, es_params)
state["mean"] = jnp.array([9999.0, -9999.0]) # a bad init guess
num_iters = 5000
for t in range(num_iters):
rng, rng_iter = jax.random.split(rng)
y, state = strategy.ask(rng_iter, state, es_params)
fitness = batch_func(y)
state = strategy.tell(y, fitness, state, es_params)
if (t + 1) % 500 == 0:
print("xNES - # Gen: {}|Fitness: {:.5f}|Params: {}".format(
t+1, state["best_fitness"], state["best_member"]))
```
| github_jupyter |
## About
This page showcases the work of [Cedric Scherer](https://www.cedricscherer.com),
built for the [TidyTuesday](https://github.com/rfordatascience/tidytuesday)
initiative. You can find the original code on his github repository
[here](https://github.com/z3tt/TidyTuesday/blob/master/R/2021_22_MarioKart.Rmd), written in [R](https://www.r-graph-gallery.com).
Thanks to him for accepting sharing his work here! Thanks also to [Tomás Capretto](https://tcapretto.netlify.app) who translated this work from R to Python! 🙏🙏
## Load libraries
As always, several libraries are needed in order to build the chart. `numpy`, `pandas` and `matplotlib` are pretty usual, but we also need some lesser known libraries like `palettable` to get some nice colors.
```
import numpy as np
import pandas as pd
import matplotlib.colors as mc
import matplotlib.pyplot as plt
from matplotlib.cm import ScalarMappable
from matplotlib.lines import Line2D
from mpl_toolkits.axes_grid1.inset_locator import inset_axes
from palettable import cartocolors
```
## Load the dataset
Today we are going to visualize world records for the Mario Kart 64 game. The game consists of 16 individual tracks and world records can be achieved for the fastest *single lap* or the fastest completed race (**three laps**). Also, through the years, players discovered **shortcuts** in many of the tracks. Fortunately, shortcut and non-shortcut world records are listed separately.
Our chart consists of a double-dumbbell plot where we visualize world record times on Mario Kart 64 with and without shortcuts. The original source of the data is [https://mkwrs.com/](https://mkwrs.com/), which holds time trial world records for all of the Mario Kart games, but we are using the version released for the [TidyTuesday](https://github.com/rfordatascience/tidytuesday) initiative on the week of 2021-05-25. You can find the original announcement and more information about the data [here](https://github.com/rfordatascience/tidytuesday/tree/master/data/2021/2021-05-25).
```
df_records = pd.read_csv('https://raw.githubusercontent.com/rfordatascience/tidytuesday/master/data/2021/2021-05-25/records.csv')
df_records.head(3)
```
<br>
<br>
<br>
<br>
From all the columns in the data, we will only use `track`, `type`, `shortcut`, `date`, and `time`.
- `track` indicates the name of the track
- `type` tells us whether the record is for single lap or a complete race
- `shortcut` is a yes/no variable that identifies records where a shortcut was used
- `date` represents the date where the record was achieved
- `time` indicates how many seconds it took to complete the track
## Prepare the dataset
Today's visualization is based on records for complete races only. Our first step is to create a `pandas.DataFrame` called `df_rank` that keeps current world records for every track.
<!-- Do you prefer this approach instead?
df_rank = df_records.query("type == 'Three Lap'")
df_rank = (
df_rank.loc[df_rank.groupby("track")["time"].idxmin()]
.sort_values("time", ascending=False)
.assign(track = lambda df: pd.Categorical(df['track'], ordered=True, categories=df["track"]))
)
-->
```
# Keep records where type is Three Lap
df_rank = df_records.query("type == 'Three Lap'")
# Keep records with the minimum time for each track
df_rank = df_rank.loc[df_rank.groupby("track")["time"].idxmin()]
# Sort by descending time
df_rank = df_rank.sort_values("time", ascending=False)
# Make "track" ordered categorical with order given by descending times
# This categorical type will be used to sort the tracks in the plot.
df_rank["track"] = pd.Categorical(df_rank["track"], ordered=True, categories=df_rank["track"])
```
Then we have `df_records_three` which holds all the records, no matter they were beaten or not. It is used to derive other data frames that used in our chart.
```
# We call '.reset_index()' to avoid SettingWithCopyWarning
df_records_three = df_records.query("type == 'Three Lap'").reset_index()
df_records_three["year"] = pd.DatetimeIndex(df_records_three["date"]).year
```
`df_connect` is the first data frame we derive. This one is used to add a dotted line that connects record times with and without shortcuts and will serve as a reference for their difference.
```
# First of all, for each track and shortcut, obtain the minimum and maximum
# value of time. These represent the most recent and first records, respectively.
df_connect = df_records_three.groupby(["track", "shortcut"]).agg(
no = ("time", min),
yes = ("time", max)
).reset_index()
# Next, put it into long format.
# Each row indicates the track, whether shortcuts were used,
# if it's the current record, and the time achieved.
df_connect = pd.melt(
df_connect,
id_vars=["track", "shortcut"],
value_vars=["no", "yes"],
var_name="record",
value_name="time"
)
# The dotted line goes from the first record without shortcut (the slowest)
# to the most recent record with shortcut (the fastest)
df_connect = df_connect.query(
"(shortcut == 'No' and record == 'no') or (shortcut == 'Yes' and record == 'yes')"
)
# Finally it is put in wide format, where there's only one row per track.
df_connect = df_connect.pivot_table(index="track", columns="record", values="time").reset_index()
```
We also have `df_longdist` and `df_shortcut`. Note each data frame consists of five columns: `track`, `year`, `max`, `min`, and `diff`. `year` refers to the year where the current record was achieved, `max` is the completetion time for the first record and `min` is the time for the current record. `diff` is simply the difference between `max` and `min`, i.e. a measurement of how much the first record was improved. `df_shortcut` and `df_longdist` refer to records with and without shortcuts, respectively.
```
# Long dist refers to records without shortcut
df_longdist = df_records_three.query("shortcut == 'No'")
# Only keep observations referring to either the first or the most recent record, by track.
grouped = df_longdist.groupby("track")
df_longdist = df_longdist.loc[pd.concat([grouped["time"].idxmax(), grouped["time"].idxmin()])]
# Create a 'group' variable that indicates whether the record
# refers to the first record, the one with maximum time,
# or to the most recent record, the one with minimum time.
df_longdist.loc[grouped["time"].idxmax(), "group"] = "max"
df_longdist.loc[grouped["time"].idxmin(), "group"] = "min"
# 'year' records the year of the most recent record
df_longdist["year"] = df_longdist.groupby("track")['year'].transform(max)
# Put the data in wide format, i.e., one observation per track.
df_longdist = df_longdist.pivot_table(index=["track", "year"], columns="group", values="time").reset_index()
df_longdist["diff"] = df_longdist["max"] - df_longdist["min"]
# Same process than above, but using records where shortcut is "Yes"
df_shortcut = df_records_three.query("shortcut == 'Yes'")
grouped = df_shortcut.groupby("track")
df_shortcut = df_shortcut.loc[pd.concat([grouped["time"].idxmax(), grouped["time"].idxmin()])]
df_shortcut.loc[grouped["time"].idxmax(), "group"] = "max"
df_shortcut.loc[grouped["time"].idxmin(), "group"] = "min"
df_shortcut["year"] = df_shortcut.groupby("track")['year'].transform(max)
df_shortcut = df_shortcut.pivot_table(index=["track", "year"], columns="group", values="time").reset_index()
df_shortcut["diff"] = df_shortcut["max"] - df_shortcut["min"]
```
All the datasets are sorted according to the order of `"track"` in `df_rank`. To do so, we first set the type of the `"track"` variable equal to the categorical type in `df_rank`, and then sort according to its levels.
```
tracks_sorted = df_rank["track"].dtype.categories.tolist()
# Sort df_connect
df_connect["track"] = df_connect["track"].astype("category")
df_connect["track"].cat.set_categories(tracks_sorted, inplace=True)
df_connect = df_connect.sort_values("track")
# Sort df_longdist
df_longdist["track"] = df_longdist["track"].astype("category")
df_longdist["track"].cat.set_categories(tracks_sorted, inplace=True)
df_longdist = df_longdist.sort_values("track")
# Sort df_shortcut
df_shortcut["track"] = df_shortcut["track"].astype("category")
df_shortcut["track"].cat.set_categories(tracks_sorted, inplace=True)
df_shortcut = df_shortcut.sort_values("track")
```
## Start building the chart
This highly customized plot demands a lot of code. It is a good practice to define the colors at the very beginning so we can refer to them by name.
```
GREY94 = "#f0f0f0"
GREY75 = "#bfbfbf"
GREY65 = "#a6a6a6"
GREY55 = "#8c8c8c"
GREY50 = "#7f7f7f"
GREY40 = "#666666"
LIGHT_BLUE = "#b4d1d2"
DARK_BLUE = "#242c3c"
BLUE = "#4a5a7b"
WHITE = "#FFFCFC" # technically not pure white
```
Today we make use of the `palettable` library to make use of the `RedOr` palette, which is the one used in the original plot. We will also make use of the `matplotlib.colors.Normalize` class to normalize values into the (0, 1) interval before we pass it to our `colormap` function and `matplotlib.colors.LinearSegmentedColormap` to create a custom colormap for blue colors.
```
# We have two colormaps, one for orange and other for blue
colormap_orange = cartocolors.sequential.RedOr_5.mpl_colormap
# And we also create a new colormap using
colormap_blue = mc.LinearSegmentedColormap.from_list("blue", [LIGHT_BLUE, DARK_BLUE], N=256)
```
`colormap_orange and` and `colormap_blue` are functions now.
```
fig, ax = plt.subplots(figsize = (15, 10))
# Add segments ---------------------------------------------------
# Dotted line connection shortcut yes/no
ax.hlines(y="track", xmin="yes", xmax="no", color=GREY75, ls=":", data=df_connect)
# Segment when shortcut==yes
# First time we use the colormap and the normalization
norm_diff = mc.Normalize(vmin=0, vmax=250)
color = colormap_orange(norm_diff(df_shortcut["diff"].values))
ax.hlines(y="track", xmin="min", xmax="max", color=color, lw=5, data=df_shortcut)
# Segment when shortcut==no. Note we are overlapping lineranges
# We use the same normalization scale.
color = colormap_orange(norm_diff(df_longdist["diff"].values))
ax.hlines(y="track", xmin="min", xmax="max", color=color, lw=4, data=df_longdist)
ax.hlines(y="track", xmin="min", xmax="max", color=WHITE, lw=2, data=df_longdist)
# Add dots -------------------------------------------------------
## Dots when shortcut==yes – first record
# zorder is added to ensure dots are on top
ax.scatter(x="max", y="track", s=200, color=GREY65, edgecolors=GREY65, lw=2.5, zorder=2, data=df_shortcut)
## Dots when shortcut==yes – latest record
# This time we normalize using the range of years in the data, and use blue colormap
norm_year = mc.Normalize(df_shortcut["year"].min(), df_shortcut["year"].max())
color = colormap_blue(norm_year(df_shortcut["year"].values))
ax.scatter(x="min", y="track", s=160, color=color, edgecolors=color, lw=2, zorder=2, data=df_shortcut)
## Dots shortcut==no – first record
color = colormap_blue(norm_year(df_longdist["year"].values))
ax.scatter(x="min", y="track", s=120, color=WHITE, edgecolors=color, lw=2, zorder=2, data=df_longdist)
## Dots shortcut==no – latest record
ax.scatter(x="max", y="track", s=120, color=WHITE, edgecolors=GREY65, lw=2, zorder=2, data=df_longdist)
# Add labels on the left side of the lollipops -------------------
# Annotations for tracks in df_shortcut
for row in range(df_shortcut.shape[0]):
ax.text(
df_shortcut["min"][row] - 7,
df_shortcut["track"][row],
df_shortcut["track"][row],
ha="right",
va="center",
size=16,
color="black",
fontname="Atlantis"
)
# Annotations for df_longdist, not in df_shortcut
for row in range(df_longdist.shape[0]):
if df_longdist["track"][row] not in df_shortcut["track"].values:
ax.text(
df_longdist["min"][row] - 7,
df_longdist["track"][row],
df_longdist["track"][row],
ha="right",
va="center",
size=17,
color="black",
fontname="Atlantis",
)
# Add labels on top of the first row of lollipops ----------------
# These labels are used to give information about the meaning of
# the different dots without having to use a legend.
# Label dots when shortcut==yes
df_shortcut_wario = df_shortcut.query("track == 'Wario Stadium'")
ax.text(
df_shortcut_wario["min"],
df_shortcut_wario["track"],
"Most recent record\nwith shortcuts\n",
color=BLUE,
ma="center",
va="bottom",
ha="center",
size=9,
fontname="Overpass"
)
ax.text(
df_shortcut_wario["max"],
df_shortcut_wario["track"],
"First record\nwith shortcuts\n",
color=GREY50,
ma="center",
va="bottom",
ha="center",
size=9,
fontname="Overpass"
)
# Label dots when shortcut==no
df_longdist_wario = df_longdist.query("track == 'Wario Stadium'")
ax.text(
df_longdist_wario["min"] - 10,
df_longdist_wario["track"],
"Most recent record\nw/o shortcuts\n",
color=BLUE,
ma="center",
va="bottom",
ha="center",
size=9,
fontname="Overpass"
)
ax.text(
df_longdist_wario["max"] + 10,
df_longdist_wario["track"],
"First record\nw/o shortcuts\n",
color=GREY50,
ma="center",
va="bottom",
ha="center",
size=9,
fontname="Overpass"
)
# Customize the layout -------------------------------------------
# Hide spines
ax.spines["left"].set_visible(False)
ax.spines["right"].set_visible(False)
ax.spines["top"].set_visible(False)
ax.spines["bottom"].set_visible(False)
# Hide y labels
ax.yaxis.set_visible(False)
# Customize x ticks
# * Remove x axis ticks
# * Put labels on both bottom and and top
# * Customize the tick labels. Only the first has the "seconds" appended.
ax.tick_params(axis="x", bottom=True, top=True, labelbottom=True, labeltop=True, length=0)
xticks = np.linspace(0, 400, num=9, dtype=int).tolist()
ax.set_xlim(-60, 400)
ax.set_xticks(xticks)
ax.set_xticklabels(["0 seconds"] + xticks[1:], fontname="Hydrophilia Iced", color=GREY40, size=9)
# Set background color for the subplot.
ax.set_facecolor(WHITE)
# Add thin vertical lines to serve as guide
# 'zorder=0' is imoprtant to they stay behind other elements in the plot.
for xtick in xticks:
ax.axvline(xtick, color=GREY94, zorder=0)
# Add vertical space to the vertical limit in the plot
x0, x1, y0, y1 = plt.axis()
plt.axis((x0, x1, y0, y1 + 0.5));
# Add custom legends ---------------------------------------------
# Legend for time difference.
# Recall the 'norm_diff()' created above.
# Create an inset axes with a given width and height.
cbaxes = inset_axes(
ax, width="0.8%", height="44%", loc=3,
bbox_to_anchor=(0.025, 0., 1, 1),
bbox_transform=ax.transAxes
)
cb = fig.colorbar(
ScalarMappable(norm=norm_diff, cmap=colormap_orange), cax=cbaxes,
ticks=[0, 50, 100, 150, 200, 250]
)
# Remove the outline of the colorbar
cb.outline.set_visible(False)
# Set label, playing with labelpad to put it in the right place
cb.set_label(
"Time difference between first and most recent record",
labelpad=-45,
color=GREY40,
size=10,
fontname="Overpass"
)
# Remove ticks in the colorbar with 'size=0'
cb.ax.yaxis.set_tick_params(
color=GREY40,
size=0
)
# Add ticklabels at given positions, with custom font and color
cb.ax.yaxis.set_ticklabels(
[0, 50, 100, 150, 200, 250],
fontname="Hydrophilia Iced",
color=GREY40,
size=10
)
# Legend for year
# We create a custom function to put the Line2D elements into a list
# that then goes into the 'handle' argument of the 'ax.legend()'
years = [2016, 2017, 2018, 2019, 2020, 2021]
def legend_dot(year):
line = Line2D(
[0],
[0],
marker="o",
markersize=10,
linestyle="none",
color=colormap_blue(norm_year(year)),
label=f"{year}"
)
return line
# Store the legend in a name because we use it to modify its elements
years_legend = ax.legend(
title="Year of Record",
handles=[legend_dot(year) for year in years],
loc=3, # lower left
bbox_to_anchor=(0.08, 0, 1, 1),
frameon=False
)
# Set font family, color and size to the elements in the legend
for text in years_legend.get_texts():
text.set_fontfamily("Hydrophilia Iced")
text.set_color(GREY40)
text.set_fontsize(10)
# Same modifications, but applied to the title.
legend_title = years_legend.get_title()
legend_title.set_fontname("Overpass")
legend_title.set_color(GREY40)
legend_title.set_fontsize(10)
# The suptitle acts as the main title.
# Play with 'x' and 'y' to get them in the place you want.
plt.suptitle(
"Let's-a-Go! You May Still Have Chances to Grab a New World Record for Mario Kart 64",
fontsize=13,
fontname="Atlantis Headline",
weight="bold",
x = 0.457,
y = 0.99
)
subtitle = [
"Most world records for Mario Kart 64 were achieved pretty recently (13 in 2020, 10 in 2021). On several tracks, the players considerably improved the time needed to complete three laps when they used shortcuts (Choco Mountain,",
"D.K.'s Jungle Parkway, Frappe Snowland, Luigi Raceway, Rainbow Road, Royal Raceway, Toad's Turnpike, Wario Stadium, and Yoshi Valley). Actually, for three out of these tracks the previous records were more than halved since 2020",
"(Luigi Raceway, Rainbow Road, and Toad's Turnpike). Four other tracks still have no records for races with shortcuts (Moo Moo Farm, Koopa Troopa Beach, Banshee Boardwalk, and Bowser's Castle). Are there none or did nobody find",
"them yet? Pretty unrealistic given the fact that since more than 24 years the game is played all around the world—but maybe you're able to find one and obtain a new world record?"
]
# And the axis title acts as a subtitle.
ax.set_title(
"\n".join(subtitle),
loc="center",
ha="center",
ma="left",
color=GREY40,
fontname="Overpass",
fontsize=9,
pad=20
)
# Add legend
fig.text(
0.8, .05, "Visualization: Cédric Scherer • Data: mkwrs.com/mk64",
fontname="Overpass",
fontsize=12,
color=GREY55,
ha="center"
)
# Set figure's background color, to match subplot background color.
fig.patch.set_facecolor(WHITE)
# Finally, save the plot!
plt.savefig(
"mario-kart-64-world-records.png",
facecolor=WHITE,
dpi=300,
bbox_inches="tight",
pad_inches=0.3
)
```
## Conclusion
That was a lot of work! And many more lines of code than using R. But here we are, a pretty good looking lollipop chart showcasing the possibilities offered by Matplotlib, which are basically infinite!
| github_jupyter |
<a href="https://colab.research.google.com/github/DaraSamii/DataDays2021/blob/main/train_category_classifier.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Dara R Samii Login
```
from google.colab import drive
drive.mount("/content/drive",force_remount=True)
%cd /content/drive/MyDrive/DataDays2021/
pwd = %pwd
pwd
```
**-------------------------------------------------------------------------------------------------------------------------------**
# installing packages
```
!pip install "dask[complete]"
!pip install -Uqq fastai
!pip install parsivar
```
# Imports
```
import dask.dataframe as dd
import pandas as pd
import os
from fastai.text.all import *
import pickle
from torch.utils.data import Dataset
from tqdm import tqdm
import torch as T
from fastai.data.core import DataLoaders
import torch.nn as nn
from torch.utils.data.dataloader import DataLoader
from helper import utils
```
# declaring paths
```
data_folder = os.path.join(pwd,"data")
final_clicked = os.path.join(data_folder,"final","final_clicked.csv")
final_products = os.path.join(data_folder,"final","final_products.csv")
category_classifier_path = os.path.join(data_folder,"models","category_classifier")
cdf = dd.read_csv(final_clicked)
cdf.head()
```
**-------------------------------------------------------------------------------------------------------------------------------**
# Creating category dataset
```
def choose_another(List,real):
while True:
choice = random.choice(List)
if choice != real:
break
return choice
def make(df, col_name,count_True=200000,count_False=200000):
unique = list(set(df[col_name]))
dd1 = df.copy()
dd2 = df.copy()
dd1["real"] = True
dd2[col_name] = df.apply(lambda x: choose_another(unique,x[col_name]),axis=1)
dd2["real"] = False
dd1 = dd1.sample(frac=1).reset_index(drop=1)[0:count_True]
dd2 = dd2.sample(frac=1).reset_index(drop=1)[0:count_False]
return pd.concat([dd1,dd2]).sample(frac=1).reset_index(drop=True)
df = cdf[cdf["is_clicked"]== True][["raw_query","products.category_name"]].compute()
df
candid_df = make(df,col_name="products.category_name",count_True = 150000,count_False=150000)
candid_df
```
## loading vocab
```
#if vocab exits:
vocab = pickle.load(open(os.path.join(category_classifier_path,"vocab"),'rb'))
vocab
```
## nlp trasnforms pipeline
```
class nlp_pipeline:
def __init__(self, vocab,):
self.vocab = vocab
self.tok = SpacyTokenizer(lang='fa')
self.num = Numericalize(vocab=self.vocab)
def encode(self,x):
x = utils._normalize_text(x)
x = tokenize1(x, self.tok)
x = self.num.encodes(x)
return x
def decode(self,x):
x = self.num.decodes(x)
x = " ".join(x)
return x
```
### testing nlp_pipeline
```
pipe = nlp_pipeline(vocab=vocab[0])
we = pipe.encode("سامسونگ گوشی")
we
pipe.decode(we)
```
## creating torch dataset
```
class categoryDataSet(Dataset):
def __init__(self, df, query_column_name, category_column_name, vocab ,target_column_name = None,test=False):
self.df = df.reset_index(drop=True)
self.query_column_name = query_column_name
self.category_column_name = category_column_name
self.test = test
if test == False:
self.target_column_name = target_column_name
self.vocab= vocab
self.nlp_pipeline = nlp_pipeline(vocab=self.vocab)
def __len__(self):
return self.df.shape[0]
def __getitem__(self, i):
x1 = self.df[self.query_column_name][i]
x1_numericalized = self.nlp_pipeline.encode(x1)
x2 = self.df[self.category_column_name][i]
x2_numericalized = self.nlp_pipeline.encode(x2)
if self.test == False:
y = self.df[self.target_column_name][i]
return (x1_numericalized,x2_numericalized), y
else:
return (x1_numericalized,x2_numericalized)
```
## spliting to valid and train
```
candid_df["is_valid"] = False
candid_df["is_valid"] = candid_df["is_valid"].apply(lambda x: True if random.random() < 0.08 else False)
```
## traind and valid dateset
```
train_ds = categoryDataSet(candid_df[candid_df["is_valid"] == False].reset_index(drop=1),"raw_query","products.category_name",vocab=vocab[0],target_column_name="real")
valid_ds = categoryDataSet(candid_df[candid_df["is_valid"] == True],"raw_query","products.category_name",vocab=vocab[0],target_column_name="real")
```
### testing dataset
```
len(train_ds),len(valid_ds)
def my_collate(batch):
b = list(zip(*batch))
x,y = b
x1,x2 = list(zip(*x))
return (nn.utils.rnn.pad_sequence(x1).T, nn.utils.rnn.pad_sequence(x2).T), T.tensor(y).to(T.long)
```
## creating torch Dataloaders
```
train_dl = DataLoader(dataset=train_ds, batch_size=180, shuffle=True, collate_fn=my_collate)
valid_dl = DataLoader(dataset=valid_ds, batch_size=180, shuffle=True, collate_fn=my_collate)
```
### testing dataloaders
```
next(iter(train_dl))
next(iter(valid_dl))
```
## fastai Dataloaders wrapper
```
fast_category_dl = DataLoaders(train_dl,valid_dl)
```
# category classifier model
```
class CatageoryClassifier(nn.Module):
def __init__(self, vocab_sz, awd_config, lin_ftrs,joint_layers,joint_drop =0.5):
super(CatageoryClassifier, self).__init__()
self.vocab_sz = vocab_sz
self.awd_config = awd_config
self.lin_ftrs = lin_ftrs
self.joint_drop = joint_drop
self.awd1 = get_text_classifier(AWD_LSTM,vocab_sz=vocab_sz, n_class=2, config=awd_config,lin_ftrs=lin_ftrs)
self.awd1[-1].layers = self.awd1[-1].layers[:-1]
self.awd2 = get_text_classifier(AWD_LSTM,vocab_sz=vocab_sz, n_class=2, config=awd_config,lin_ftrs=lin_ftrs)
self.awd2[-1].layers = self.awd2[-1].layers[:-1]
self.joint_layers = [lin_ftrs[-1]*2] + joint_layers + [2]
linBins = []
for i in range(0,len(self.joint_layers)-1):
linBins.append(LinBnDrop(self.joint_layers[i],self.joint_layers[i+1]))
self.LinBins = nn.Sequential(*linBins)
self.Softmax = nn.Softmax(dim=1)
def forward(self,x):
x1 = self.awd1(x[0])[0]
x2 = self.awd2(x[1])[0]
X = T.cat([x1,x2],dim=1)
X = self.LinBins(X)
return self.Softmax(X)
def reset(self):
self.awd1.reset()
self.awd2.reset()
awd_conf = {'bidir': True,
'emb_sz': 1000,
'embed_p': 0.05,
'hidden_p': 0.3,
'input_p': 0.4,
'n_hid': 1800,
'n_layers': 5,
'output_p': 0.4,
'pad_token': 1,
'weight_p': 0.5}
nn_model = CatageoryClassifier(len(vocab[0]),awd_config=awd_conf,lin_ftrs=[1000],joint_layers=[500])
state_dict = T.load(open(os.path.join(category_classifier_path,"models","category_classifier.pth"),'rb'))
nn_model.load_state_dict(state_dict)
a = next(iter(train_dl))
nn_model
nn_model(a[0])
loss_func = nn.CrossEntropyLoss()
learn = Learner(fast_category_dl,
nn_model,
loss_func = loss_func,
path = category_classifier_path,
metrics=[accuracy,error_rate,Recall(),Precision(),F1Score()]).to_fp16()
grp = ShowGraphCallback
svm = SaveModelCallback(at_end=False,every_epoch=False,reset_on_fit=False,monitor='f1_score',fname="category_classifier",)
esc = EarlyStoppingCallback(patience=3)
rlr = ReduceLROnPlateau(monitor="valid_loss",patience=2,factor=10,)
learn.add_cbs([grp,svm,esc,rlr,ModelResetter])
learn.cbs
learn.lr_find()
learn.fit_one_cycle(20,1.2e-05)
learn.fine_tune(15,base_lr=1e-05,freeze_epochs=10)
learn.fine_tune(15,base_lr=1e-04,freeze_epochs=10)
test_x = next(iter(valid_dl))[0]
learn.model((test_x[0].cuda(),test_x[1].cuda()))
learn.unfreeze()
learn.fit_one_cycle(1,0.02,wd=0.1)
```
| github_jupyter |
```
import numpy as np
import matplotlib.pyplot as plt
import pickle
import os
import keras
import copy
import tensorflow as tf
import seaborn as sn
import pandas as pd
import keras.backend as K
from keras.models import Sequential
from keras.layers.core import Activation, Dense
from keras.layers import Flatten, LSTM, Masking
from keras.models import Model
from keras.layers import Input
from sklearn import metrics
from model_helper import *
```
## 1. Load in dataset
```
cwd = os.getcwd()
parent_wd = cwd.replace('/model', '')
training_set_path = parent_wd + '/preprocessing/training_seq_n_12_rmrp0'
dev_set_path = parent_wd + '/preprocessing/dev_seq_n_12_rmrp0'
test_set_path = parent_wd + '/preprocessing/test_seq_n_12_rmrp0'
with open(training_set_path, 'rb') as f:
training_set = pickle.load(f)
with open(dev_set_path, 'rb') as f:
dev_set = pickle.load(f)
with open(test_set_path, 'rb') as f:
test_set = pickle.load(f)
X_train = training_set['X']
Y_train = training_set['Y']
X_dev = dev_set['X']
Y_dev = dev_set['Y']
X_test = test_set['X']
Y_test = test_set['Y']
```
## 2. Define GradeNet Model
```
np.random.seed(0)
tf.random.set_seed(0)
inputs = Input(shape = (12, 22))
mask = Masking(mask_value = 0.).compute_mask(inputs)
lstm0 = LSTM(20, activation='tanh', input_shape=(12, 22), kernel_initializer='glorot_normal', return_sequences = 'True')(
inputs, mask = mask)
dense1 = Dense(100, activation='relu', kernel_initializer='glorot_normal')(lstm0)
dense2 = Dense(80, activation='relu', kernel_initializer='glorot_normal')(dense1)
dense3 = Dense(75, activation='relu', kernel_initializer='glorot_normal')(dense2)
dense4 = Dense(50, activation='relu', kernel_initializer='glorot_normal')(dense3)
dense5 = Dense(20, activation='relu', kernel_initializer='glorot_normal')(dense4)
dense6 = Dense(10, activation='relu', kernel_initializer='glorot_normal')(dense5)
flat = Flatten()(dense6)
softmax2 = Dense(10, activation='softmax', name = 'softmax2')(flat)
lstm1 = LSTM(20, activation='tanh', kernel_initializer='glorot_normal', return_sequences = True)(dense6)
lstm2 = LSTM(20, activation='tanh', kernel_initializer='glorot_normal')(lstm1)
dense7 = Dense(15, activation='relu', kernel_initializer='glorot_normal')(lstm2)
dense8 = Dense(15, activation='relu', kernel_initializer='glorot_normal')(dense7)
softmax3 = Dense(10, activation='softmax', name = 'softmax2')(dense8)
def custom_loss(layer):
def loss(y_true,y_pred):
loss1 = K.sparse_categorical_crossentropy(y_true, y_pred)
loss2 = K.sparse_categorical_crossentropy(y_true, layer)
return K.mean(loss1 + loss2, axis=-1)
return loss
GradeNet = Model(inputs=[inputs], outputs=[softmax3])
GradeNet.compile(optimizer='adam',
loss=custom_loss(softmax2),
metrics=['sparse_categorical_accuracy'])
```
## 2-1. Training of GradeNet
### To load pretrained weights, please skip to 2-2.
```
history_GradeNet_all = []
for i in range(10):
history_GradeNet = GradeNet.fit(X_train, Y_train, epochs=10, batch_size=256, validation_data = (X_dev, Y_dev),
class_weight = {0:1, 1:1, 2:2, 3: 2, 4: 1, 5: 4, 6:2, 7: 4, 8: 8, 9: 8})
history_GradeNet_all.append(history_GradeNet)
# Change weights
for i in range(10):
history_GradeNet = GradeNet.fit(X_train, Y_train, epochs=10, batch_size=256, validation_data = (X_dev, Y_dev),
class_weight = {0:1, 1:1, 2:2, 3: 4, 4: 1, 5: 4, 6: 8, 7: 8, 8: 8, 9: 8})
history_GradeNet_all.append(history_GradeNet)
```
### Plot training history
```
GradeNet_history_package = plot_history(history_GradeNet, 'GradeNet')
```
### Save training results
```
# saving trained results
save_pickle(GradeNet_history_package, 'GradeNet_train_history')
GradeNet.save_weights("GradeNet.h5")
```
## 2-2. Loading pretrained GradeNet
```
# load model weight
GradeNet.load_weights(parent_wd + '/model/GradeNet.h5')
# load training history
history_path = parent_wd + '/model/GradeNet_train_history'
with open(history_path, 'rb') as f:
GradeNet_history_package = pickle.load(f)
plot_history_package(GradeNet_history_package, 'GradeNet')
```
## 3. Analyze GradeNet Performance
### Confusion Matrix
```
plot_confusion_matrix(Y_train, GradeNet.predict(X_train).argmax(axis=1), title = 'Confusion matrix of GradeNet(Training set)')
plot_confusion_matrix(Y_dev, GradeNet.predict(X_dev).argmax(axis=1), title = 'Confusion matrix of GradeNet(Dev set)')
plot_confusion_matrix(Y_test, GradeNet.predict(X_test).argmax(axis=1), title = 'Confusion matrix of GradeNet(Test set)')
```
### F1 score
```
F1_train = metrics.f1_score(Y_train, GradeNet.predict(X_train).argmax(axis=1), average = 'macro')
print(F1_train)
F1_dev = metrics.f1_score(Y_dev, GradeNet.predict(X_dev).argmax(axis=1), average = 'macro')
print(F1_dev)
F1_test = metrics.f1_score(Y_test, GradeNet.predict(X_test).argmax(axis=1), average = 'macro')
print(F1_test)
```
### Accuracy and Rough accuracy
```
accuracy_train = compute_accuracy(Y_train, GradeNet.predict(X_train).argmax(axis=1))
print("Exactly accuracy rate of training set = %s" %accuracy_train[0])
print("+/-1 Accuracy rate of training set= %s" %accuracy_train[1])
accuracy_dev = compute_accuracy(Y_dev, GradeNet.predict(X_dev).argmax(axis=1))
print("Exactly accuracy rate of dev set = %s" %accuracy_dev[0])
print("+/-1 Accuracy rate of dev set = %s" %accuracy_dev[1])
accuracy_test = compute_accuracy(Y_test, GradeNet.predict(X_test).argmax(axis=1))
print("Exactly accuracy rate of test set = %s" %accuracy_test[0])
print("+/-1 Accuracy rate of test set = %s" %accuracy_test[1])
```
### KL divergence
```
kl = tf.keras.losses.KLDivergence()
kld_train = kl(tf.one_hot(Y_train.astype(int), depth = 10), GradeNet.predict(X_train)).numpy()
print(kld_train)
kld_dev = kl(tf.one_hot(Y_dev.astype(int), depth = 10), GradeNet.predict(X_dev)).numpy()
print(kld_dev)
kld_test = kl(tf.one_hot(Y_test.astype(int), depth = 10), GradeNet.predict(X_test)).numpy()
print(kld_test)
```
### Mean absolute error
```
# MAE
mae_train = np.mean(np.abs(Y_train - GradeNet.predict(X_train).argmax(axis=1)))
print(mae_train)
mae_dev = np.mean(np.abs(Y_dev - GradeNet.predict(X_dev).argmax(axis=1)))
print(mae_dev)
mae_test = np.mean(np.abs(Y_test - GradeNet.predict(X_test).argmax(axis=1)))
print(mae_test)
```
### Classification report
```
sk_report_train = metrics.classification_report(
digits=4,
y_true=Y_train,
y_pred=GradeNet.predict(X_train).argmax(axis=1))
print(sk_report_train)
sk_report_dev = metrics.classification_report(
digits=4,
y_true=Y_dev,
y_pred=GradeNet.predict(X_dev).argmax(axis=1))
print(sk_report_dev)
sk_report_test = metrics.classification_report(
digits=4,
y_true=Y_test,
y_pred=GradeNet.predict(X_test).argmax(axis=1))
print(sk_report_test)
AUC_train = metrics.roc_auc_score(Y_train, GradeNet.predict(X_train), multi_class= 'ovr', average="macro")
print(AUC_train)
AUC_dev = metrics.roc_auc_score(Y_dev, GradeNet.predict(X_dev), multi_class= 'ovr', average="macro")
print(AUC_dev)
AUC_test = metrics.roc_auc_score(Y_test, GradeNet.predict(X_test), multi_class= 'ovr', average="macro")
print(AUC_test)
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import scipy
import os
import matplotlib
import matplotlib.pyplot as plt
import matplotlib.style as style
from matplotlib import gridspec
from matplotlib.colors import ListedColormap
%matplotlib inline
import seaborn as sns
import ast
import glob
import warnings
matplotlib.rcParams['font.family'] = "sans-serif"
matplotlib.rcParams['font.sans-serif'] = "Arial"
smaller_size = 13
medium_size = 14
bigger_size = 16
plt.rc('font', size=bigger_size) # controls default text sizes
plt.rc('axes', titlesize=medium_size) # fontsize of the axes title
plt.rc('axes', labelsize=medium_size) # fontsize of the x and y labels
plt.rc('xtick', labelsize=smaller_size) # fontsize of the tick labels
plt.rc('ytick', labelsize=smaller_size) # fontsize of the tick labels
plt.rc('legend', fontsize=smaller_size) # legend fontsize
plt.rc('figure', titlesize=bigger_size) # fontsize of the figure title
noises = [0, 0.01, 0.05, 0.1, 0.15, 0.2, 0.3, 0.5, 0.8]
model_summary_full = pd.read_csv("12_noise_model_summary.csv")
indexes1 = [x in ["True-True", "True-False"] for x in model_summary_full.label]
model_summary = model_summary_full[indexes1]
indexes2 = [x in noises for x in model_summary.noise]
model_summary = model_summary[indexes2]
model_summary.head()
whitenoise = pd.read_csv('whitenoise_correlation_subset.csv',index_col=0)
wn_summary = pd.read_csv('whitenoise_summary.csv',index_col=0)
# three panels
fig = plt.figure(figsize=(24, 5))
plt.gcf().subplots_adjust(left=0.25, wspace = 0.25)
gs = gridspec.GridSpec(1, 3, width_ratios=[5, 5, 5])
ax1 = plt.subplot(gs[0])
#model_summary = pd.read_csv("12_noise/12_noise_model_summary.csv")
sns.boxplot(x="noise", y="correlation", data=model_summary, width=0.38,
boxprops=dict(alpha=0.8, color="#FFE7E7", edgecolor="black", linewidth=1.5),
whiskerprops=dict(color="black", linewidth=1.5), fliersize=0, ax=ax1)
sns.swarmplot(x="noise", y="correlation", data=model_summary, s=4.8, alpha=0.7, color='#8F0202', ax=ax1)
ax1.set_xlabel("Standard deviation of multiplicative noise")
ax1.set_ylabel("Model performance (correlation)", size=14)
ax1.spines['right'].set_visible(False)
ax1.spines['top'].set_visible(False)
#ax1.set_ylim(0.7,0.95)
ax1.set_ylim(-0.1,1.0)
ax1.set_xticks(range(len(noises)))
ax1.set_xticklabels(noises)
#ax1.set_yticks([0.7,0.75,0.8,0.85,0.9,0.95])
ax1.set_yticks([0,0.2,0.4,0.6,0.8,1.0])
#ax1.set_yticklabels(['0.70','0.75','0.80','0.85','0.90','0.95'], size=14)
ax1.set_yticklabels(['0','0.2','0.4','0.6','0.8','1.0'])
ax1.text(-0.2,1,'A', weight='bold',transform=ax1.transAxes)
ax3 = plt.subplot(gs[1])
sns.boxplot(x="noise", y="correlation", data=whitenoise, width=0.38,
boxprops=dict(alpha=0.8, color="#fce7ff", edgecolor="black", linewidth=1.5),
whiskerprops=dict(color="black", linewidth=1.5), fliersize=0, ax=ax3)
# sns.swarmplot(x="noise", y="correlation", data=model_summary,
# s=4.8, alpha=0.7, color='#8F0202', ax=ax)
sns.swarmplot(x="noise", y="correlation", data=whitenoise,
s=4.8, alpha=0.7, color='#8f0250', ax=ax3)
ax3.set_xlabel("Standard deviation of additive noise")
#ax1.set_xlabel("Ratio of noise", size=14)
ax3.set_ylabel("Model performance (correlation)")
ax3.spines['right'].set_visible(False)
ax3.spines['top'].set_visible(False)
#ax1.set_ylim(0.7,0.95)
ax3.set_ylim(-0.1,1.0)
wnoises = [0,0.05,0.1,0.2,0.5,1.0,2.0,3.0,5.0]
ax3.set_xticks(range(len(wnoises)))
ax3.set_xticklabels(wnoises)
#ax1.set_yticks([0.7,0.75,0.8,0.85,0.9,0.95])
ax3.set_yticks([0,0.2,0.4,0.6,0.8,1.0])
#ax1.set_yticklabels(['0.70','0.75','0.80','0.85','0.90','0.95'], size=14)
ax3.set_yticklabels(['0','0.2','0.4','0.6','0.8','1.0'])
ax3.text(-0.2,1,'B', weight='bold',transform=ax3.transAxes)
ax2 = plt.subplot(gs[2])
fracs = [0.1,0.2,0.3,0.4,0.5,0.6,0.7,0.8,0.9]
model_summary_subset = pd.read_csv("12_subset_model_summary.csv")
indexes1 = [x in ["True-True", "True-False"] for x in model_summary_subset.label]
model_summary_subset = model_summary_subset[indexes1]
sns.swarmplot(x="frac", y="correlation", data=model_summary_subset, s=4.8, alpha=0.7, color='#1A389C', ax=ax2)
sns.boxplot(x="frac", y="correlation", data=model_summary_subset, width=0.42,
boxprops=dict(alpha=0.8, color="#C8DCFE", edgecolor="black", linewidth=1.5),
whiskerprops=dict(color="black", linewidth=1.5), fliersize=0, ax=ax2)
ax2.set_ylim(0.3,1.0)
ax2.set_xlabel("Fraction of data used in training")
ax2.set_ylabel("Model performance (correlation)")
ax2.spines['right'].set_visible(False)
ax2.spines['top'].set_visible(False)
ax2.set_xticks([0,1,2,3,4,5,6,7,8])
ax2.set_xticklabels([0.1,0.2,0.3,0.4,0.5,0.6,0.7,0.8,0.9])
ax2.set_yticks([0.3,0.4,0.5,0.6,0.7,0.8,0.9,1.0])
ax2.set_yticklabels([0.3,0.4,0.5,0.6,0.7,0.8,0.9,1.0])
ax2.text(-0.22,1,'C', weight='bold',transform=ax2.transAxes)
#fig.savefig("Figure4.pdf", format="pdf")
```
| github_jupyter |
```
%matplotlib inline
from matplotlib import pyplot
import numpy
```
All calculations in AMUSE are done with quantities having units. These quantities and their units are implemented as python classes and can be used almost everywere you would normaly use a number (or a numpy array).
As we will do in all tutorials we start by importing everything from amuse.lab:
```
from amuse.lab import *
```
You can create a quantity by typing a number and combining it with a unit using the bar `|` operator.
```
1.989e30 | units.kg
```
Most operations you can do on numbers, you can also perform on quantities. For example, assuming the earth is a sphere, let's calculate some properties:
```
earth_radius = 6384 | units.km
print("diameter :", earth_radius * 2)
print("surface area :", 4.0 * numpy.pi * earth_radius**2)
print("volume :", 4.0 / 3.0 * numpy.pi * earth_radius**3)
```
It is also possible to combine quantities with different units in a calculation. To continue our properties of the earth example, lets calcute it's density:
```
earth_mass = 5.97219e24 | units.kg
earth_volume = 4.0 * numpy.pi * earth_radius**3 / 3
density = earth_mass / earth_volume
print("density :", earth_mass / earth_volume)
```
Note that density has a *numerical value* and a *unit*
```
print("numerical value:", density.number)
print("unit:", density.unit)
print("simplified:", density.in_base())
```
You will want to interact with other python libraries that simply cannot handle units. For those cases you can convert the quantity into a value of a specific unit:
```
print("earth mass in gram :", earth_mass.value_in(units.g))
```
Astrophysical units are also supported:
```
print("earth mass in solar masses :", earth_mass.value_in(units.MSun))
```
To also print the unit, you can use a conversion function:
```
earth_mass2 = earth_mass.in_(units.MSun)
print("earth mass :", earth_mass2)
```
Note that while the two quantities have different representation, they are still the same (though this is true within numerical precision):
```
print(earth_mass == earth_mass2)
```
Numpy arrays and python lists can also be converted to vector quantities. Once converted, the resulting quantities support a lot of numpy operations
```
masses = [641.85, 4868.5, 5973.6, 102430, 86832, 568460, 1898600] | (1e21 * units.kg)
radii = [0.532, 0.950, 1, 3.86, 3.98, 9.14, 10.97] | (6384 * units.km)
print("sum of planet masses: ", masses.sum().in_(units.MSun))
print("planet diameters: ", (radii * 2))
```
You can create your own unit with a new name using the `units.named` function. This functions takes the name of the unit, a symbol for printing and the unit it is based on. You can define a unit to represent the volume of the earth
```
earth_volume_unit = units.named('Vol-Earth', 'EarthVol', earth_volume.to_unit())
print(earth_volume.in_(earth_volume_unit))
```
Do you note something odd? You have to be careful with numerical precision!
Most operations on a vector quantity are elementwise. We can do some operations on the array of planet masses and raddii we defined earlier. (As AMUSE uses numpy internally for these operations we refer to the numpy documentation, if you want to learn more)
```
volumes = 4.0 / 3.0 * numpy.pi * radii**3
earth_density = earth_mass / earth_volume
print("volumes :", volumes.in_(earth_volume_unit))
print("densities :", (masses / volumes))
```
Quantities become normal numbers or numpy arrays when the units cancel out in a calcution (You can use this fact, to replace a `value_in` function call with a division):
```
print(volumes / earth_volume)
```
Operations with incompatible units will fail:
```
print(earth_mass + earth_volume)
```
| github_jupyter |
```
from math import *
def blackcell(i,k,j):
print("module blkcell("+"G"+str(i)+"_"+str(k)+","+"P"+str(i)+"_"+str(k)+","+"G"+str(k-1)+"_"+str(j)+","+"P"+str(k-1)+"_"+str(j)+","+"G"+str(i)+"_"+str(j)+","+"P"+str(i)+"_"+str(j)+")"+";")
print(" input "+"G"+str(i)+"_"+str(k)+","+"P"+str(i)+"_"+str(k)+","+"G"+str(k-1)+"_"+str(j)+","+"P"+str(k-1)+"_"+str(j)+";")
print(" output "+"G"+str(i)+"_"+str(j)+","+"P"+str(i)+"_"+str(j)+";")
print(" wire s1 ;")
print(" assign s1 = P"+str(i)+"_"+str(k)+" & G"+str(k-1)+"_"+str(j)+";" )
print(" assign " + "G"+str(i)+"_"+str(j)+"="+"s1|"+"G"+str(i)+"_"+str(k)+";")
print(" assign " + "P"+str(i)+"_"+str(j)+"="+"P"+str(i)+"_"+str(k)+"&"+"P"+str(k-1)+"_"+str(j)+";")
print("endmodule")
blackcell(i,j,k)
i=0
k=5
j=8
def greycell(i,k,j):
print("module greycell("+"G"+str(i)+"_"+str(k)+","+"P"+str(i)+"_"+str(k)+","+"G"+str(k-1)+"_"+str(j)+","+"G"+str(i)+"_"+str(j)+""+")"+";")
print(" input "+"G"+str(i)+"_"+str(k)+","+"P"+str(i)+"_"+str(k)+","+"G"+str(k-1)+"_"+str(j)+""+";")
print(" output "+"G"+str(i)+"_"+str(j)+";")
print(" wire s1 ;")
print(" assign s1 = P"+str(i)+"_"+str(k)+" & G"+str(k-1)+"_"+str(j)+";" )
print(" assign " + "G"+str(i)+"_"+str(j)+"="+"s1|"+"G"+str(i)+"_"+str(k)+";")
print("endmodule")
greycell(i,j,k)
def buffer(i,j):
print("module buffer("+"G"+str(i)+"_"+str(j)+","+"P"+str(i)+"_"+str(j)+","+"G"+str(i)+"o_"+str(j)+"o,"+"P"+str(i)+"o_"+str(j)+"o);")
print(" input "+"G"+str(i)+"_"+str(j)+","+"P"+str(i)+"_"+str(j)+";")
print(" output "+"G"+str(i)+"o_"+str(j)+"o,"+"P"+str(i)+"o_"+str(j)+"o;")
print(" assign " + "G"+str(i)+"o_"+str(j)+"o="+"G"+str(i)+"_"+str(j)+";")
print(" assign " + "P"+str(i)+"o_"+str(j)+"o="+"P"+str(i)+"_"+str(j)+";")
print("endmodule")
buffer(9,3)
def printcell(s,i,j,k,a,b):
if(s=="black"):
printblack(i,j,k,a,b)
elif(s=="buffer"):
printbuffer(i,j,a,b)
else :
printgrey(i,j,k,a,b)
def printblack(i,j,k,a,b):
print(" blkcell cell"+str(a)+"_"+str(b)+"("+"G"+str(i)+"_"+str(k)+","+"P"+str(i)+"_"+str(k)+","+"G"+str(k-1)+"_"+str(j)+","+"P"+str(k-1)+"_"+str(j)+","+"G"+str(i)+"_"+str(j)+","+"P"+str(i)+"_"+str(j)+")"+";")
def printbuffer(i,j,a,b):
print("")
def printgrey(i,j,k,a,b):
print(" greycell cell"+str(a)+"_"+str(b)+"("+"G"+str(i)+"_"+str(k)+","+"P"+str(i)+"_"+str(k)+","+"G"+str(k-1)+"_"+str(j)+","+"G"+str(i)+"_"+str(j)+")"+";")
printblack(0,0,0,0,0)
def printverilog(n):
initializeGP()
initialiseverilog(n)
print_wires(n)
assignGP(n)
#print("assign G_1_0 = 0;")
for a in range(1,1+ceil(log(n)/log(2))):
for b in range(n):
s = findcell(a,b)
i,j,k = findijk(a,b)
printcell(s,i,j,k,a,b)
computeS(n)
print("assign C_out = (G"+str(n-1)+"_0"+"&P"+str(n)+"_"+str(n)+")|G"+str(n)+"_"+str(n)+";")
print("endmodule")
def findcell(a,b):
s = ""
if((b//(2**(a-1)))%2==0):
s = "buffer"
elif((b//(2**(a-1)))==1):
s = "grey"
else:
s = "black"
return s
def findijk(a,b):
j=0
j = (2*a)*(b//(2**a))
i = b
a = a-1
k = (2*a)*(b//(2**a))
if(a==0):
k = i
return i,j,k;
def print_wires(n):
for c in range(n+1):
for d in range(n+1):
print(" wire G"+str(c)+"_"+str(d)+";")
print(" wire P"+str(c)+"_"+str(d)+";")
def initialiseverilog(n):
blackcell(6,8,10)
print()
greycell(4,3,2)
print()
# buffer(12,14)
print()
printmodule(n)
def printmodule(n):
argin=""
argout=""
for i in range(1,n+1):
argin+="A"+str(i)+","
for i in range(1,n+1):
argin+="B"+str(i)+","
for i in range(1,n+1):
argout+="S"+str(i)+","
argout = argout[: -1]
print("module sklansky("+"C_in,"+argin+argout+",C_out)"+";")
print("input "+"C_in,"+ argin[: -1]+";")
print("output "+"C_out,"+ argout+";")
def initializeGP():
print("module initializeGP(A,B,G,P);")
print("input A,B;")
print("output G,P;")
print("assign G = A&B;")
print("assign P = A^B;")
print("endmodule")
print()
def assignGP(n):
print("assign G0_0 = C_in;")
print("assign P0_0 = 0;")
for i in range(1,n+1):
print("initializeGP a"+str(i)+"(A"+str(i)+",B"+str(i)+",G"+str(i)+"_"+str(i)+",P"+str(i)+"_"+str(i)+");")
print()
def computeS(n):
for i in range(1,n+1):
print("assign S"+str(i)+" = G"+str(i-1)+"_0^"+"P"+str(i)+"_"+str(i)+";")
printverilog(16)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_12_04_atari.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# T81-558: Applications of Deep Neural Networks
**Module 12: Deep Learning and Security**
* Instructor: [Jeff Heaton](https://sites.wustl.edu/jeffheaton/), McKelvey School of Engineering, [Washington University in St. Louis](https://engineering.wustl.edu/Programs/Pages/default.aspx)
* For more information visit the [class website](https://sites.wustl.edu/jeffheaton/t81-558/).
# Module 12 Video Material
* Part 12.1: Introduction to the OpenAI Gym [[Video]](https://www.youtube.com/watch?v=_KbUxgyisjM&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_12_01_ai_gym.ipynb)
* Part 12.2: Introduction to Q-Learning [[Video]](https://www.youtube.com/watch?v=uwcXWe_Fra0&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_12_02_qlearningreinforcement.ipynb)
* Part 12.3: Keras Q-Learning in the OpenAI Gym [[Video]](https://www.youtube.com/watch?v=Ya1gYt63o3M&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_12_03_keras_reinforce.ipynb)
* **Part 12.4: Atari Games with Keras Neural Networks** [[Video]](https://www.youtube.com/watch?v=t2yIu6cRa38&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_12_04_atari.ipynb)
* Part 12.5: How Alpha Zero used Reinforcement Learning to Master Chess [[Video]](https://www.youtube.com/watch?v=ikDgyD7nVI8&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_12_05_alpha_zero.ipynb)
# Google CoLab Instructions
The following code ensures that Google CoLab is running the correct version of TensorFlow.
```
try:
from google.colab import drive
%tensorflow_version 2.x
COLAB = True
print("Note: using Google CoLab")
except:
print("Note: not using Google CoLab")
COLAB = False
```
# Part 12.4: Atari Games with Keras Neural Networks
The Atari 2600 is a home video game console from Atari, Inc. Released on September 11, 1977. It is credited with popularizing the use of microprocessor-based hardware and games stored on ROM cartridges instead of dedicated hardware with games physically built into the unit. The 2600 was bundled with two joystick controllers, a conjoined pair of paddle controllers, and a game cartridge: initially [Combat](https://en.wikipedia.org/wiki/Combat_(Atari_2600)), and later [Pac-Man](https://en.wikipedia.org/wiki/Pac-Man_(Atari_2600)).
Atari emulators are popular and allow many of the old Atari video games to be played on modern computers. They are even available as JavaScript.
* [Virtual Atari](http://www.virtualatari.org/listP.html)
Atari games have become popular benchmarks for AI systems, particularly reinforcement learning. OpenAI Gym internally uses the [Stella Atari Emulator](https://stella-emu.github.io/). The Atari 2600 is shown in Figure 12.ATARI.
**Figure 12.ATARI: The Atari 2600**

### Installing Atari Emulator
```
pip install gym[atari]
```
### Actual Atari 2600 Specs
* CPU: 1.19 MHz MOS Technology 6507
* Audio + Video processor: Television Interface Adapter (TIA)
* Playfield resolution: 40 x 192 pixels (NTSC). Uses a 20-pixel register that is mirrored or copied, left side to right side, to achieve the width of 40 pixels.
* Player sprites: 8 x 192 pixels (NTSC). Player, ball, and missile sprites use pixels that are 1/4 the width of playfield pixels (unless stretched).
* Ball and missile sprites: 1 x 192 pixels (NTSC).
* Maximum resolution: 160 x 192 pixels (NTSC). Max resolution is only somewhat achievable with programming tricks that combine sprite pixels with playfield pixels.
* 128 colors (NTSC). 128 possible on screen. Max of 4 per line: background, playfield, player0 sprite, and player1 sprite. Palette switching between lines is common. Palette switching mid line is possible but not common due to resource limitations.
* 2 channels of 1-bit monaural sound with 4-bit volume control.
### OpenAI Lab Atari Breakout
OpenAI Gym can be used with Windows; however, it requires a special [installation procedure](https://towardsdatascience.com/how-to-install-openai-gym-in-a-windows-environment-338969e24d30) Figure 12.BREAKOUT shows the Atari Breakout Game.
**Figure 12.BREAKOUT: Atari Breakout**

(from Wikipedia)
Breakout begins with eight rows of bricks, with each two rows a different color. The color order from the bottom up is yellow, green, orange and red. Using a single ball, the player must knock down as many bricks as possible by using the walls and/or the paddle below to ricochet the ball against the bricks and eliminate them. If the player's paddle misses the ball's rebound, he or she will lose a turn. The player has three turns to try to clear two screens of bricks. Yellow bricks earn one point each, green bricks earn three points, orange bricks earn five points and the top-level red bricks score seven points each. The paddle shrinks to one-half its size after the ball has broken through the red row and hit the upper wall. Ball speed increases at specific intervals: after four hits, after twelve hits, and after making contact with the orange and red rows.
The highest score achievable for one player is 896; this is done by eliminating two screens of bricks worth 448 points per screen. Once the second screen of bricks is destroyed, the ball in play harmlessly bounces off empty walls until the player restarts the game, as no additional screens are provided. However, a secret way to score beyond the 896 maximum is to play the game in two-player mode. If "Player One" completes the first screen on his or her third and last ball, then immediately and deliberately allows the ball to "drain", Player One's second screen is transferred to "Player Two" as a third screen, allowing Player Two to score a maximum of 1,344 points if he or she is adept enough to keep the third ball in play that long. Once the third screen is eliminated, the game is over.
The original arcade cabinet of Breakout featured artwork that revealed the game's plot to be that of a prison escape. According to this release, the player is actually playing as one of a prison's inmates attempting to knock a ball and chain into a wall of their prison cell with a mallet. If the player successfully destroys the wall in-game, their inmate escapes with others following.
### Breakout (BreakoutDeterministic-v4) Specs:
* BreakoutDeterministic-v4
* State size (RGB): (210, 160, 3)
* Actions: 4 (discrete)
The video for this course demonstrated playing Breakout. The following [example code](https://github.com/wau/keras-rl2/blob/master/examples/dqn_atari.py) was used.
The following code can be used to probe an environment to see the shape of its states and actions.
```
import gym
env = gym.make("BreakoutDeterministic-v4")
print(f"Obesrvation space: {env.observation_space}")
print(f"Action space: {env.action_space}")
```
| github_jupyter |
```
from sklearn.ensemble import RandomForestRegressor
from sklearn.model_selection import train_test_split
import seaborn as sns
from __future__ import absolute_import, division, print_function, unicode_literals
import numpy as np
import pandas as pd
import os
train = pd.read_csv("data/Train_dataset.csv")
train.head()
train.shape
train_1 = train.dropna()
train_1.shape
test = pd.read_csv("data/Test_dataset.csv")
test.head()
test_1 = test.dropna()
test_1.shape
from sklearn import preprocessing
def cleanData(dataframe):
df2 = dataframe
df2 = df2.drop(['people_ID'], axis = 1)
le = preprocessing.LabelEncoder()
df2.Region = le.fit_transform(df2.Region.astype(str))
df2.Gender = le.fit_transform(df2.Gender.astype(str))
df2.Occupation = le.fit_transform(df2.Occupation.astype(str))
df2.Mode_transport = le.fit_transform(df2.Mode_transport.astype(str))
df2.Disease = le.fit_transform(df2.Disease.astype(str))
df2.COVID_Infected = le.fit_transform(df2.COVID_Infected.astype(str))
df2.Region = le.fit_transform(df2.Region)
df2.Gender = le.fit_transform(df2.Gender)
df2.Occupation = le.fit_transform(df2.Occupation)
df2.Mode_transport = le.fit_transform(df2.Mode_transport)
df2.Disease = le.fit_transform(df2.Disease)
df2.COVID_Infected = le.fit_transform(df2.COVID_Infected)
return df2
clean_train = cleanData(train_1)
clean_train.head()
clean_test = cleanData(test_1)
clean_test.columns
x = clean_train.drop(['Infect_Prob'], axis=1)
y = clean_train.Infect_Prob
X_train, X_test, y_train, y_test = train_test_split(x, y, test_size=0.2, random_state=42)
y.head()
rf = RandomForestRegressor(n_estimators = 1000, random_state = 42)
rf.fit(X_train, y_train)
from sklearn.metrics import accuracy_score
from sklearn.metrics import confusion_matrix
y_pred= list(rf.predict(X_test))
errors = abs(y_pred - y_test)
print('Mean Absolute Error:', round(np.mean(errors), 2), 'degrees.')
mape = 100 * (errors / y_test)
accuracy = 100 - np.mean(mape)
print('Accuracy:', round(accuracy, 2), '%.')
test_1
test_data = [[5942,'Delhi','Male','Driver','Public','Diabetes',52,1,'Yes']]
test_dataframe = pd.DataFrame(test_data, columns =['people_ID','Region','Gender','Occupation','Mode_transport','Disease','Age','FT/month','COVID_Infected'], dtype = float)
clean_test_data = cleanData(test_dataframe)
clean_test_data.to_csv(r'result/prob1.csv', index=False)
clean_test_data
testData = list(rf.predict(clean_test_data))
print(testData)
def priority(testData):
if testData[0] < 30 :
return 3
elif 30 < testData[0] < 70 :
return 2
else :
return 1
print("Your priority is :" ,priority(testData))
```
| github_jupyter |
```
from keras.preprocessing.image import load_img, save_img, img_to_array
import numpy as np
from scipy.optimize import fmin_l_bfgs_b
import time
from keras.applications import vgg19
from keras.applications.imagenet_utils import preprocess_input
from keras import backend as K
import tensorflow as tf
import keras
base_image_path = 'C:/Users/IS96273/Desktop/content.jpg'
style_reference_image_path = 'C:/Users/IS96273/Desktop/style.jpg'
iterations = 1
img_nrows = 400; img_ncols = 534
def preprocess_image(image_path):
img = load_img(image_path, target_size=(img_nrows, img_ncols))
img = img_to_array(img)
img = np.expand_dims(img, axis=0)
img = preprocess_input(img)
return img
def deprocess_image(x):
x = x.reshape((img_nrows, img_ncols, 3))
# Remove zero-center by mean pixel
x[:, :, 0] += 103.939
x[:, :, 1] += 116.779
x[:, :, 2] += 123.68
# 'BGR'->'RGB'
x = x[:, :, ::-1]
x = np.clip(x, 0, 255).astype('uint8')
return x
base_image = K.variable(preprocess_image(base_image_path))
x = K.variable(preprocess_image(base_image_path))
style_reference_image = K.variable(preprocess_image(style_reference_image_path))
random_pixels = np.random.randint(256, size=(img_nrows, img_ncols, 3))
combination_image = preprocess_input(np.expand_dims(random_pixels, axis=0))
combination_image = K.variable(combination_image)
content_model = vgg19.VGG19(input_tensor=base_image, weights='imagenet', include_top=False)
style_model = vgg19.VGG19(input_tensor=style_reference_image, weights='imagenet', include_top=False)
content_outputs = dict([(layer.name, layer.output) for layer in content_model.layers])
style_outputs = dict([(layer.name, layer.output) for layer in style_model.layers])
generated_model = vgg19.VGG19(input_tensor=combination_image, weights='imagenet', include_top=False)
generated_outputs = dict([(layer.name, layer.output) for layer in generated_model.layers])
def gram_matrix(x):
features = K.batch_flatten(K.permute_dimensions(x, (2, 0, 1)))
gram = K.dot(features, K.transpose(features))
return gram
def style_loss(style, combination):
S = gram_matrix(style)
C = gram_matrix(combination)
channels = 3
size = img_nrows * img_ncols
return K.sum(K.square(S - C)) / (4. * (pow(channels,2)) * (pow(size,2)))
def content_loss(base, combination):
return K.sum(K.square(combination - base))
loss = K.variable(0)
base_image_features = content_outputs['block5_conv2'][0]
combination_features = generated_outputs['block5_conv2'][0]
contentloss = content_loss(base_image_features, combination_features)
feature_layers = ['block1_conv1', 'block2_conv1', 'block3_conv1', 'block4_conv1', 'block5_conv1']
styleloss = K.variable(0)
for layer_name in feature_layers:
style_reference_features = style_outputs[layer_name][0]
combination_features = generated_outputs[layer_name][0]
styleloss = styleloss + style_loss(style_reference_features, combination_features)
alpha = 0.025; beta = 0.2
loss = alpha * contentloss + beta * styleloss
grads = K.gradients(loss, combination_image)
outputs = [loss]
outputs += grads
f_outputs = K.function([combination_image], outputs)
def eval_loss_and_grads(x):
x = x.reshape((1, img_nrows, img_ncols, 3))
outs = f_outputs([x])
loss_value = outs[0]
if len(outs[1:]) == 1:
grad_values = outs[1].flatten().astype('float64')
else:
grad_values = np.array(outs[1:]).flatten().astype('float64')
return loss_value, grad_values
class Evaluator(object):
def __init__(self):
self.loss_value = None
self.grads_values = None
def loss(self, x):
assert self.loss_value is None
loss_value, grad_values = eval_loss_and_grads(x)
self.loss_value = loss_value
self.grad_values = grad_values
return self.loss_value
def grads(self, x):
assert self.loss_value is not None
grad_values = np.copy(self.grad_values)
self.loss_value = None
self.grad_values = None
return grad_values
evaluator = Evaluator()
x = preprocess_image(base_image_path)
for i in range(0,iterations):
print("epoch ",i)
x, min_val, info = fmin_l_bfgs_b(evaluator.loss, x.flatten(), fprime=evaluator.grads, maxfun=20)
img = deprocess_image(x.copy())
fname = 'C:/Users/IS96273/Desktop/generated_%d.png' % i
save_img(fname, img)
```
| github_jupyter |
```
import numpy as np
import networkx as nx
from itertools import combinations
from random import randrange
import matplotlib.pyplot as plt
from numpy.linalg import inv
import scipy.sparse as sp
np.set_printoptions(precision=3)
def normalize_adj(adj):
"""Symmetrically normalize adjacency matrix."""
adj = sp.coo_matrix(adj)
rowsum = np.array(adj.sum(1))
d_inv_sqrt = np.power(rowsum, -0.5).flatten()
d_inv_sqrt[np.isinf(d_inv_sqrt)] = 0.
d_mat_inv_sqrt = sp.diags(d_inv_sqrt)
return adj.dot(d_mat_inv_sqrt).transpose().dot(d_mat_inv_sqrt).toarray()
def get_sparse_eigen_decomposition(graph, K):
adj = nx.adjacency_matrix(graph).toarray()
normalized_adj = normalize_adj(adj)
eigenval,eigenvectors = np.linalg.eig(normalized_adj)
eigenval_Ksparse = np.argsort(eigenval)[-K:]
V_ksparse = np.zeros(adj.shape)
V_ksparse[:,eigenval_Ksparse] = eigenvectors[:,eigenval_Ksparse]
V_ksparse = np.matrix(V_ksparse)
V_ksparse_H = V_ksparse.getH()
return V_ksparse, V_ksparse_H
def plot_graph(graph):
nx.draw_shell(graph,with_labels=True,)
def get_random_undirected_graph(N):
random_undirected_graph=nx.Graph()
for node_pair in combinations(range(N), 2):
if randrange(5) == 1:
random_undirected_graph.add_edge(node_pair[0],node_pair[1])
return random_undirected_graph
def get_H(num_nodes):
H = np.matrix(np.identity(num_nodes))
H_h = H.getH()
return H, H_h
def get_W(VH, H_h, H, V):
a = np.matmul(VH, H_h)
b = np.matmul(a, H)
W = np.matmul(b, V)
return W
def get_random_signal_from_cov(cov_matrix, mean):
return np.random.multivariate_normal(mean, cov_matrix)
def argmax(K,remaining_node):
u = (0,-1) # score, index
for candidate in remaining_node:
v_u = V_ksparse_H[:,candidate]
v_u_H = V_ksparse[candidate,:]
a = (v_u_H * K)
numerator = (((a * W) * K) * v_u)
lamda_inv = 1.0 / float(cov_w[candidate][candidate]) # get lam^(-1)_w,u should always be the same
denumerator = lamda_inv + (a * v_u)
score = numerator/denumerator
if score > u[0]:
u = (score, candidate)
return u[1]
def update_K(K, u): #Should be O(K^2)
v_u = V_ksparse_H[:,u]
v_u_H = V_ksparse[u,:]
numerator = (((K * v_u) * v_u_H)* K)
lamda_inv = 1.0 / float(cov_w[u][u]) # get lam^(-1)_w,u should always be the same
denumerator = lamda_inv + ((v_u_H * K) * v_u)
matrix = numerator/denumerator
x = (W * matrix)
return K - x
def greedy_algo():
G_subset = []
remaining_node = list(range(0,NUM_NODES))
K = cov_x
for j in l:
u = argmax(K,remaining_node)
K = update_K(K, u)
G_subset.append(u)
remaining_node.remove(u)
return G_subset, K
def get_upper_bound_trace_K():
upper_bound_matrix = np.matrix(W * cov_x)
return float(upper_bound_matrix.trace())
def get_K_trace(possible_set):
inv_cov_x = inv(cov_x)
for i in possible_set:
v_i = V_ksparse_H[:,i]
v_i_H = V_ksparse[i,:]
lamda_inv = 1.0 / float(cov_w[i][i])
inv_cov_x = inv_cov_x + lamda_inv * (v_i * v_i_H)
K = np.matrix((((H * V_ksparse) * inv(inv_cov_x)) * V_ksparse_H) * H_h)
return float(K.trace())
def brute_force_algo():
rank = {}
min_set = []
optimal_K_T = get_upper_bound_trace_K()
all_possible_set_combination = combinations(range(NUM_NODES), number_node_sampled)
for possible_set in all_possible_set_combination:
score = get_K_trace(possible_set)
rank[str(list(possible_set))] = score
if score <= optimal_K_T:
optimal_K_T = score
min_set = possible_set
return optimal_K_T, min_set, rank
NUM_NODES = 20
NOISE_CONSTANT = 10e2
K_sparse = 5
MEAN = np.zeros((NUM_NODES,))
number_node_sampled = 5
def get_relative_suboptimality(optimal_K_T, f_K_T, empty_K_T):
return (f_K_T - optimal_K_T) / (empty_K_T - optimal_K_T)
relative_sub_Erdos_greedy = []
relative_sub_Erdos_randomized = []
for simul in range(10):
graph = get_random_undirected_graph(NUM_NODES)
while(len(graph.nodes()) < NUM_NODES):
graph = get_random_undirected_graph(NUM_NODES)
V_ksparse, V_ksparse_H = get_sparse_eigen_decomposition(graph, K_sparse)
H, H_h = get_H(NUM_NODES)
cov_w = NOISE_CONSTANT * np.identity(NUM_NODES)
cov_x = np.identity(NUM_NODES)
x = get_random_signal_from_cov(cov_x, MEAN)
w = get_random_signal_from_cov(cov_w, MEAN)
y = x + w
W = get_W(V_ksparse_H, H_h, H, V_ksparse)
l = range(1,(number_node_sampled + 1))
G_subset, K = greedy_algo()
empty_set = get_K_trace([])
optimal_K_T, min_set, rank = brute_force_algo()
relative_sub_Erdos_greedy.append(get_relative_suboptimality(optimal_K_T, rank[str(list(sorted(G_subset)))]), empty_set)
relative_sub_Erdos_randomized.append(get_relative_suboptimality(optimal_K_T, rank[str(list(sorted(G_subset)))]), empty_set))
import matplotlib.patches as mpatches
from matplotlib import colors
import matplotlib.pyplot as plt
import scipy.stats as stats
n_bins = 30
fig, axs = plt.subplots(1, 3, sharey=True)
axs[0].set_ylabel('Count')
axs[0].hist(relative_sub_Erdos_greedy, bins=n_bins)
axs[0].set_title("Erdos-Renyi")
axs[1].set_ylabel('Count')
axs[1].hist(relative_sub, bins=n_bins)
axs[1].set_title("Pref. attachment")
axs[2].set_ylabel('Count')
axs[2].hist(relative_sub, bins=n_bins)
axs[2].set_title("Random")
fig.tight_layout()
plt.subplots_adjust(top=0.92, bottom=0.08, left=0.10, right=3, hspace=0.5,
wspace=0.3)
plt.show()
```
| github_jupyter |
# 3) Model Training
```
import pandas as pd
import numpy as np
heart_disease = pd.read_csv('../00.datasets/heart-diseases.csv')
heart_disease.head()
```
# 3.1) Fitting the model to the data
* `X`: features, features variables, data
* `y`: labels, target, target variables, ground truth
```
from sklearn.ensemble import RandomForestClassifier
# Features and Lables
X = heart_disease.drop('target', axis=1)
y = heart_disease['target']
# split data
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=42)
# train model
rfc = RandomForestClassifier(n_estimators=100)
rfc.fit(X_train, y_train)
rfc.score(X_test, y_test)
```
--------
# 3.2) Making Predictions
Two ways to make predictions:
* `predict()`
* `predict_proba()`
### predict()
```
y_pred = rfc.predict(X_test)
y_pred
rfc.score(X_test, y_test)
# compare predicted labels to true labels to evalute the model
# it is the same as getting score
np.mean(y_pred == y_test)
# accuracy score
from sklearn.metrics import accuracy_score
accuracy_score(y_test, y_pred)
```
### predict_proba()
- returns probabilities estimates of classification labels
```
y_pred_proba = rfc.predict_proba(X_test)
y_pred_proba[:5]
# check against with predict()
rfc.predict(X_test)
```
We can see that first prediction is 0, which means no heart disease.
- For the same prediction row using pred_proba it says 95% for label being 0 (No Heart Disease) and 5% for label being 1 (Heart Disease).
- for the second row, we can see that model is not really confident between two labels, unlike the first one.
```
# let's check against with true value
y_test[:5]
```
-------------
-------------
## Predictions on Regression Model
```
from sklearn.datasets import load_boston
boston = load_boston()
# boston
boston_df = pd.DataFrame(boston['data'], columns=boston['feature_names'])
boston_df['target'] = pd.DataFrame(boston['target'])
boston_df.head()
from sklearn.ensemble import RandomForestRegressor
X = boston_df.drop('target', axis=1)
y = boston_df['target']
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=20, random_state=42)
rf = RandomForestRegressor(n_estimators=100)
rf.fit(X_train, y_train)
y_pred = rf.predict(X_test)
# Evaluation Matrix
from sklearn.metrics import mean_absolute_error, mean_squared_error
MAE = mean_absolute_error(y_test, y_pred)
RMSE = np.sqrt(mean_squared_error(y_test, y_pred))
MAE, RMSE
```
| github_jupyter |
```
if 'google.colab' in str(get_ipython()):
!pip install -q condacolab
import condacolab
condacolab.install()
"""
You can run either this notebook locally (if you have all the dependencies and a GPU) or on Google Colab.
Instructions for setting up Colab are as follows:
1. Open a new Python 3 notebook.
2. Import this notebook from GitHub (File -> Upload Notebook -> "GITHUB" tab -> copy/paste GitHub URL)
3. Connect to an instance with a GPU (Runtime -> Change runtime type -> select "GPU" for hardware accelerator)
"""
BRANCH = 'main'
# If you're using Google Colab and not running locally, run this cell.
# install NeMo
if 'google.colab' in str(get_ipython()):
!python -m pip install git+https://github.com/NVIDIA/NeMo.git@$BRANCH#egg=nemo_toolkit[all]
if 'google.colab' in str(get_ipython()):
!conda install -c conda-forge pynini=2.1.3
! mkdir images
! wget https://github.com/NVIDIA/NeMo/blob/$BRANCH/tutorials/text_processing/images/deployment.png -O images/deployment.png
! wget https://github.com/NVIDIA/NeMo/blob/$BRANCH/tutorials/text_processing/images/pipeline.png -O images/pipeline.png
import os
import wget
import pynini
import nemo_text_processing
```
# Task Description
Inverse text normalization (ITN) is a part of the Automatic Speech Recognition (ASR) post-processing pipeline.
ITN is the task of converting the raw spoken output of the ASR model into its written form to improve the text readability. For example, `in nineteen seventy` should be changed to `in 1975` and `one hundred and twenty three dollars` to `$123`.
# NeMo Inverse Text Normalization
NeMo ITN is based on weighted finite-state
transducer (WFST) grammars. The tool uses [`Pynini`](https://github.com/kylebgorman/pynini) to construct WFSTs, and the created grammars can be exported and integrated into [`Sparrowhawk`](https://github.com/google/sparrowhawk) (an open-source version of [The Kestrel TTS text normalization system](https://www.cambridge.org/core/journals/natural-language-engineering/article/abs/kestrel-tts-text-normalization-system/F0C18A3F596B75D83B75C479E23795DA)) for production. The NeMo ITN tool can be seen as a Python extension of `Sparrowhawk`.
Currently, NeMo ITN provides support for English and the following semiotic classes from the [Google Text normalization dataset](https://www.kaggle.com/richardwilliamsproat/text-normalization-for-english-russian-and-polish):
DATE, CARDINAL, MEASURE, DECIMAL, ORDINAL, MONEY, TIME, PLAIN.
We additionally added the class `WHITELIST` for all whitelisted tokens whose verbalizations are directly looked up from a user-defined list.
The toolkit is modular, easily extendable, and can be adapted to other languages and tasks like [text normalization](https://github.com/NVIDIA/NeMo/blob/stable/tutorials/text_processing/Text_Normalization.ipynb). The Python environment enables an easy combination of text covering grammars with NNs.
The rule-based system is divided into a classifier and a verbalizer following [Google's Kestrel](https://www.researchgate.net/profile/Richard_Sproat/publication/277932107_The_Kestrel_TTS_text_normalization_system/links/57308b1108aeaae23f5cc8c4/The-Kestrel-TTS-text-normalization-system.pdf) design: the classifier is responsible for detecting and classifying semiotic classes in the underlying text, the verbalizer the verbalizes the detected text segment.
The overall NeMo ITN pipeline from development in `Pynini` to deployment in `Sparrowhawk` is shown below:

# Quick Start
## Add ITN to your Python ASR post-processing workflow
ITN is a part of the `nemo_text_processing` package which is installed with `nemo_toolkit`. Installation instructions could be found [here](https://github.com/NVIDIA/NeMo/tree/main/README.rst).
```
from nemo_text_processing.inverse_text_normalization.inverse_normalize import InverseNormalizer
inverse_normalizer = InverseNormalizer()
raw_text = "we paid one hundred and twenty three dollars for this desk, and this."
inverse_normalizer.inverse_normalize(raw_text, verbose=False)
```
In the above cell, `one hundred and twenty three dollars` would be converted to `$123`, and the rest of the words remain the same.
## Run Inverse Text Normalization on an input from a file
Use `run_predict.py` to convert a spoken text from a file `INPUT_FILE` to a written format and save the output to `OUTPUT_FILE`. Under the hood, `run_predict.py` is calling `inverse_normalize()` (see the above section).
```
# If you're running the notebook locally, update the NEMO_TEXT_PROCESSING_PATH below
# In Colab, a few required scripts will be downloaded from NeMo github
NEMO_TOOLS_PATH = '<UPDATE_PATH_TO_NeMo_root>/nemo_text_processing/inverse_text_normalization'
DATA_DIR = 'data_dir'
os.makedirs(DATA_DIR, exist_ok=True)
if 'google.colab' in str(get_ipython()):
NEMO_TOOLS_PATH = '.'
required_files = ['run_predict.py',
'run_evaluate.py']
for file in required_files:
if not os.path.exists(file):
file_path = 'https://raw.githubusercontent.com/NVIDIA/NeMo/' + BRANCH + '/nemo_text_processing/inverse_text_normalization/' + file
print(file_path)
wget.download(file_path)
elif not os.path.exists(NEMO_TOOLS_PATH):
raise ValueError(f'update path to NeMo root directory')
INPUT_FILE = f'{DATA_DIR}/test.txt'
OUTPUT_FILE = f'{DATA_DIR}/test_itn.txt'
! echo "on march second twenty twenty" > $DATA_DIR/test.txt
! python $NEMO_TOOLS_PATH/run_predict.py --input=$INPUT_FILE --output=$OUTPUT_FILE
# check that the raw text was indeed converted to the written form
! cat $OUTPUT_FILE
```
## Run evaluation
[Google Text normalization dataset](https://www.kaggle.com/richardwilliamsproat/text-normalization-for-english-russian-and-polish) consists of 1.1 billion words of English text from Wikipedia, divided across 100 files. The normalized text is obtained with [The Kestrel TTS text normalization system](https://www.cambridge.org/core/journals/natural-language-engineering/article/abs/kestrel-tts-text-normalization-system/F0C18A3F596B75D83B75C479E23795DA)).
Although a large fraction of this dataset can be reused for ITN by swapping input with output, the dataset is not bijective.
For example: `1,000 -> one thousand`, `1000 -> one thousand`, `3:00pm -> three p m`, `3 pm -> three p m` are valid data samples for normalization but the inverse does not hold for ITN.
We used regex rules to disambiguate samples where possible, see `nemo_text_processing/inverse_text_normalization/clean_eval_data.py`.
To run evaluation, the input file should follow the Google Text normalization dataset format. That is, every line of the file needs to have the format `<semiotic class>\t<unnormalized text>\t<self>` if it's trivial class or `<semiotic class>\t<unnormalized text>\t<normalized text>` in case of a semiotic class.
Example evaluation run:
`python run_evaluate.py \
--input=./en_with_types/output-00001-of-00100 \
[--cat CATEGORY] \
[--filter]`
Use `--cat` to specify a `CATEGORY` to run evaluation on (all other categories are going to be excluded from evaluation). With the option `--filter`, the provided data will be cleaned to avoid disambiguates (use `clean_eval_data.py` to clean up the data upfront).
```
eval_text = """PLAIN\ton\t<self>
DATE\t22 july 2012\tthe twenty second of july twenty twelve
PLAIN\tthey\t<self>
PLAIN\tworked\t<self>
PLAIN\tuntil\t<self>
TIME\t12:00\ttwelve o'clock
<eos>\t<eos>
"""
INPUT_FILE_EVAL = f'{DATA_DIR}/test_eval.txt'
with open(INPUT_FILE_EVAL, 'w') as f:
f.write(eval_text)
! cat $INPUT_FILE_EVAL
! python $NEMO_TOOLS_PATH/run_evaluate.py --input=$INPUT_FILE_EVAL
```
`run_evaluate.py` call will output both **sentence level** and **token level** accuracies.
For our example, the expected output is the following:
```
Loading training data: data_dir/test_eval.txt
Sentence level evaluation...
- Data: 1 sentences
100% 1/1 [00:00<00:00, 58.42it/s]
- Denormalized. Evaluating...
- Accuracy: 1.0
Token level evaluation...
- Token type: PLAIN
- Data: 4 tokens
100% 4/4 [00:00<00:00, 504.73it/s]
- Denormalized. Evaluating...
- Accuracy: 1.0
- Token type: DATE
- Data: 1 tokens
100% 1/1 [00:00<00:00, 118.95it/s]
- Denormalized. Evaluating...
- Accuracy: 1.0
- Token type: TIME
- Data: 1 tokens
100% 1/1 [00:00<00:00, 230.44it/s]
- Denormalized. Evaluating...
- Accuracy: 1.0
- Accuracy: 1.0
- Total: 6
Class | Num Tokens | Denormalization
sent level | 1 | 1.0
PLAIN | 4 | 1.0
DATE | 1 | 1.0
CARDINAL | 0 | 0
LETTERS | 0 | 0
VERBATIM | 0 | 0
MEASURE | 0 | 0
DECIMAL | 0 | 0
ORDINAL | 0 | 0
DIGIT | 0 | 0
MONEY | 0 | 0
TELEPHONE | 0 | 0
ELECTRONIC | 0 | 0
FRACTION | 0 | 0
TIME | 1 | 1.0
ADDRESS | 0 | 0
```
# C++ deployment
The instructions on how to export `Pynini` grammars and to run them with `Sparrowhawk`, could be found at [NeMo/tools/text_processing_deployment](https://github.com/NVIDIA/NeMo/tree/main/tools/text_processing_deployment).
# WFST and Common Pynini Operations
Finite-state acceptor (or FSA) is a finite state automaton that has a finite number of states and no output. FSA either accepts (when the matching patter is found) or rejects a string (no match is found).
```
print([byte for byte in bytes('fst', 'utf-8')])
# create an acceptor from a string
pynini.accep('fst')
```
Here `0` - is a start note, `1` and `2` are the accept nodes, while `3` is a finite state.
By default (token_type="byte", `Pynini` interprets the string as a sequence of bytes, assigning one byte per arc.
A finite state transducer (FST) not only matches the pattern but also produces output according to the defined transitions.
```
# create an FST
pynini.cross('fst', 'FST')
```
Pynini supports the following operations:
- `closure` - Computes concatenative closure.
- `compose` - Constructively composes two FSTs.
- `concat` - Computes the concatenation (product) of two FSTs.
- `difference` - Constructively computes the difference of two FSTs.
- `invert` - Inverts the FST's transduction.
- `optimize` - Performs a generic optimization of the FST.
- `project` - Converts the FST to an acceptor using input or output labels.
- `shortestpath` - Construct an FST containing the shortest path(s) in the input FST.
- `union`- Computes the union (sum) of two or more FSTs.
The list of most commonly used `Pynini` operations could be found [https://github.com/kylebgorman/pynini/blob/master/CHEATSHEET](https://github.com/kylebgorman/pynini/blob/master/CHEATSHEET).
Pynini examples could be found at [https://github.com/kylebgorman/pynini/tree/master/pynini/examples](https://github.com/kylebgorman/pynini/tree/master/pynini/examples).
Use `help()` to explore the functionality. For example:
```
help(pynini.union)
```
# NeMo ITN API
NeMo ITN defines the following APIs that are called in sequence:
- `find_tags() + select_tag()` - creates a linear automaton from the input string and composes it with the final classification WFST, which transduces numbers and inserts semantic tags.
- `parse()` - parses the tagged string into a list of key-value items representing the different semiotic tokens.
- `generate_permutations()` - takes the parsed tokens and generates string serializations with different reorderings of the key-value items. This is important since WFSTs can only process input linearly, but the word order can change from spoken to written form (e.g., `three dollars -> $3`).
- `find_verbalizer() + select_verbalizer` - takes the intermediate string representation and composes it with the final verbalization WFST, which removes the tags and returns the written form.

# References and Further Reading:
- [Zhang, Yang, Bakhturina, Evelina, Gorman, Kyle and Ginsburg, Boris. "NeMo Inverse Text Normalization: From Development To Production." (2021)](https://arxiv.org/abs/2104.05055)
- [Ebden, Peter, and Richard Sproat. "The Kestrel TTS text normalization system." Natural Language Engineering 21.3 (2015): 333.](https://www.cambridge.org/core/journals/natural-language-engineering/article/abs/kestrel-tts-text-normalization-system/F0C18A3F596B75D83B75C479E23795DA)
- [Gorman, Kyle. "Pynini: A Python library for weighted finite-state grammar compilation." Proceedings of the SIGFSM Workshop on Statistical NLP and Weighted Automata. 2016.](https://www.aclweb.org/anthology/W16-2409.pdf)
- [Mohri, Mehryar, Fernando Pereira, and Michael Riley. "Weighted finite-state transducers in speech recognition." Computer Speech & Language 16.1 (2002): 69-88.](https://cs.nyu.edu/~mohri/postscript/csl01.pdf)
| github_jupyter |
<a href="https://colab.research.google.com/github/lustraka/Data_Analysis_Workouts/blob/main/Analyse_Twitter_Data/wrangle_report.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
## WeRateDogs Data Wrangle Report
This report briefly describes wrangling efforts focused on data from an archive of WeRateDogs Twitter account. **WeRateDogs** Twitter account rates people's dogs with a humorous comment about the dog. The account was started in 2015 by college student Matt Nelson, and has received international media attention both for its popularity and for the attention drawn to social media copyright law (see [Wikipedia](https://en.wikipedia.org/wiki/WeRateDogs)).
## Gather Data
Data is gathered from three sources:
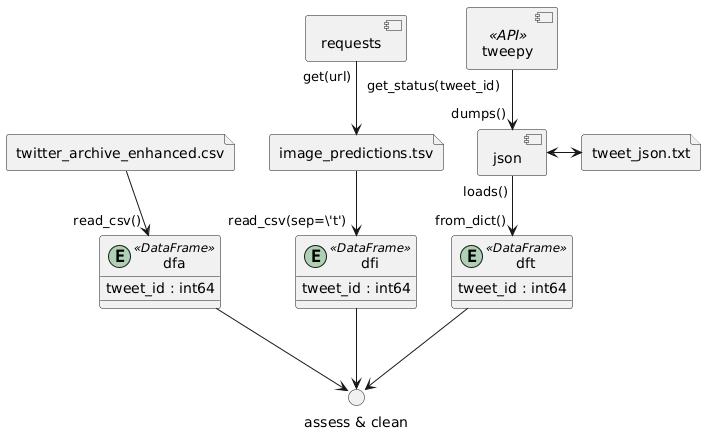
### Enhanced Twitter Archive
The WeRateDogs Twitter archive contains basic tweet data for all tweets from the November 15th, 2015 to the August 17th, 2017. The each tweet's text had been used to extract rating, dog name, and dog "stage" (i.e. doggo, floofer, pupper, and puppo) to enhance this archive.
The enhanced archive is stored in the `twitter_archive_enhanced.csv` file and read directly by `pd.read_csv()` method to the `dfa` DataFrame.
### Image Prediction File
Every image in the WeRateDogs Twitter archive had been run through a machine learning algorithm to classify breeds of dogs. The `image_prediction.tsv` file contains top three predictions alongside each tweet ID, image URL, and the image number that corresponded to the most confident prediction.
The image prediction file is first downloaded using the `requests` library and then loaded with `pd.read_csv()` method to the `dfi` DataFrame.
### Twitter API
IDs of tweets from the Twitter archive was used to access missing valuable data, namely the number of likes (*favorite_count*) and the number of retweets (*retweet_count*), via Twitter's API. Before we could ran our API querying code, we needed to set up our own Twitter application. We used the `tweepy` library to create an API object for gathering tweet data:
```python
import tweepy
consumer_key = 'hidden'
consumer_secret = 'hidden'
access_token = 'hidden'
access_secret = 'hidden'
auth = tweepy.OAuthHandler(consumer_key, consumer_secret)
auth.set_access_token(access_token, access_secret)
api = tweepy.API(auth, wait_on_rate_limit=True, wait_on_rate_limit_notify=True)
```
After quering each tweet ID, we wrote its JSON data to the `tweet_json.txt` file with each tweet's JSON data on its own line. We then read this file, line by line, to create the `dft` DataFrame with required data extracted.
## Assess Data
Key assumptions of assessing data are as follows:
* We only want original ratings (no retweets or replies) that have images. Though there are 5000+ tweets in the dataset, not all are dog ratings and some are retweets.
* Assessing and cleaning the entire dataset completely would require a lot of time. Therefore, we will assess and clean 8 quality issues and 3 tidiness issues in this dataset.
* The fact that the rating numerators are greater than the denominators does not need to be cleaned. This [unique rating system](http://knowyourmeme.com/memes/theyre-good-dogs-brent) is a big part of the popularity of WeRateDogs.
* We will gather the additional tweet data only for tweets in the *twitter_archive_enhanced.csv* dataset.
With these assumptions in mind we assessed the input data and identified these issues:
**Quality issues**
- Q1. Replies are not original tweets.
- Q2. Retweets are not original tweets.
- Q3. Some tweets don't have any image
- Q4. Some ratings are incorrectly identified
- Q5. Some ratings are missing
- Q6. Names starting with lowercase are incorrect
- Q7. Names with value None are incorrect
- Q8. Column timestamp has the dtype object (string)
**Tidiness issues**
- T1. Dogs' stages (doggo, pupper, puppo, floofer) as columns
- T2. Multiple image predictions in one row
- T3. Data in multiple datasets
Concerning quality, Q1, Q2, Q3 and Q5 are **completeness** issues in a sense that there is data out of the scope of this project. Q8 is a **validity** issue. Q4, Q6, and Q7 are **accuracy** issues.
## Clean Data
Before cleaning we made copies of input data:
```python
dfa_clean = dfa.copy() # archive
dfi_clean = dfi.copy() # image predictions
dft_clean = dft.copy() # data from Twitter API
```
Here is a toolbox for cleaning efforts:
Issue | Code | Test
--- | --- | ---
Q1 | Filter rows out by df.isna(); Drop cols by df.drop() | df.info()
Q2 | ditto | ditto
Q3 | Filter rows out by df.apply() | df.apply()
Q4 | Update values by df.at[] | df.loc[]
Q5 | Drop rows by df.drop() | df.index.intersection()
Q6 | Replace values by df.apply() | re.findall()
Q7 | ditto | df.query()
Q8 | Convert dtype by pd.to_datetime() | df.dtype
T1 | Derive a new col by df.apply(func, axis=1) | df.value_counts(), df.info()
T2 | ditto | df.info()
T3 | Merge datasets by df.merge() | df.info(), df.head()
The next diagram displays transformations used to provide a tidy clean dataset:
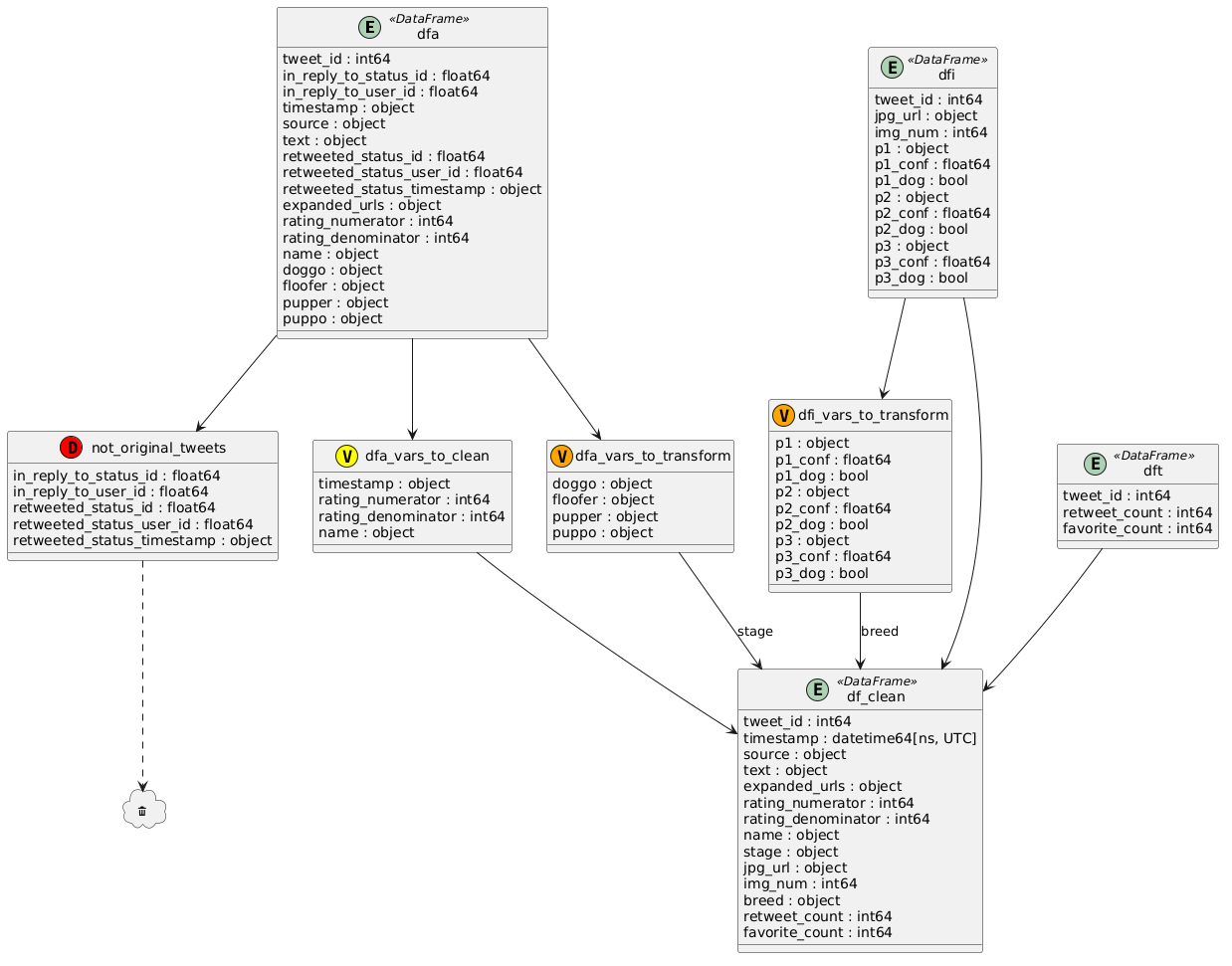
## Store Data
Even if a Jupyter notebook ensures repeatability of the wrangling process, it is recommended to save the cleaned dataset for further processing. It is beneficial especially in light of the amount of time needed for quering the Twitter API.
We saved the cleaned dataset to both a CSV file and a SQlite database using this code:
```python
# Store the dataframe for further processing in CSV format
with open('twitter_archive_master.csv', 'w') as file:
df_clean.to_csv(file)
# Store the dataframe for further processing in a database
from sqlalchemy import create_engine
# Create SQLAlchemy engine and empty database
engine = create_engine('sqlite:///weratedogsdata_clean.db')
# Store dataframes in database
df_clean.to_sql('df_clean', engine, index=False)
```
To restore data use this code (provided that the archive had been uploaded to a GitHub repository):
```python
# Restore dataframe for further processing
import requests
from sqlalchemy import create_engine
# Upload the database from GitHub
url_db = 'https://github.com/lustraka/Data_Analysis_Workouts/blob/main/Analyse_Twitter_Data/weratedogsdata_clean.db?raw=true'
r = requests.get(url_db)
with open('weratedogsdata_clean.db', 'wb') as file:
file.write(r.content)
# Create SQLAlchemy engine and connect to the database
engine = create_engine('sqlite:///weratedogsdata_clean.db')
# Read dataframes from SQlite database
df_clean = pd.read_sql('SELECT * FROM df_clean', engine)
df_clean.shape
```
## Result: WeRateDogs Master Data Set
The description of the resulting dataset is as follows:
| # | Variable | Non-Null | Nunique | Dtype | Notes |
|---|----------|----------|---------|-------|-------|
| 0 | tweet_id | 1657 | 1657 | int64 | The Tweet's unique identifier .|
| 1 | timestamp | 1657 | 1657 | datetime64[ns, UTC] | Time when this Tweet was created. |
| 2 | source | 1657 | 3 | object | Utility used to post the Tweet. |
| 3 | text | 1657 | 1657 | object | The actual text of the status update. |
| 4 | expanded_urls | 1657 | 1657 | object | The URLs of the Tweet's photos. |
| 5 | rating_numerator | 1657 | 26 | int64 | The rating numerator extracted from the text. |
| 6 | rating_denominator | 1657 | 10 | int64 | The rating denominator extracted from the text. |
| 7 | name | 1657 | 831 | object | The dog's name extracted from the text. |
| 8 | stage | 1657 | 5 | object | The dog's stage extracted from the text.|
| 9 | jpg_url | 1657 | 1657 | object | The URL of the image used to classify the breed of dog. |
| 10 | img_num | 1657 | 4 | int64 | The image number that corresponded to the most confident prediction. |
| 11 | breed | 1657 | 113 | object | The most confident classification of the breed of dog predicted from the image. |
| 12 | retweet_count | 1657 | 1352 | int64 | Number of times this Tweet has been retweeted. |
| 13 | favorite_count | 1657 | 1561 | int64 | Indicates approximately how many times this Tweet has been liked by Twitter users. |
## Caveats
- The cleaned dataset has 1657 observations starting at the November 15th, 2015 when the WeRateDogs Twitter account was launched and ending at the August 17th, 2017 when the archive was exported.
- Variables *rating_numerator, rating_denominator, name,* and *stage* was extracted from the tweet's text. The rating is a part of a humorous aspect of the content. There is hardly any value in analysing these variables.
- The variable *breed* is inferred from the image using machine learning algorithm. We can use this variable keeping in mind that there can be some inaccuracies.
- The variables *favorite_count*, and *retweet_count* reflects the preferences of Twitter users. We can use these variables keeping in mind they come from a non-random sample of human population.
| github_jupyter |
```
import numpy as np
import pandas as pd
data_path = "dataset/winequality-red.csv"
wine_df = pd.read_csv(data_path)
wine_df.head()
#print the shape
wine_df.shape
wine_df.count()
wine_df.describe()
wine_df.info()
#check for missing values
wine_df.isna().sum()
#draw box for all columns plot for checking distribution and outliers
import matplotlib.pyplot as plt
import seaborn as sns
for i in wine_df.columns:
sns.boxplot(x=wine_df[i])
plt.show()
#plotting the heatmap
plt.figure(figsize=(20,10))
sns.heatmap(wine_df.corr(),annot=True)
plt.show()
#create a copy for visualization
wine_df_copy = wine_df.copy()
#plotting counts of category
wine_df_copy.quality = wine_df_copy.quality.map({0:'0',1:'1',2:'2',3:'3',4:'4',5:'5',6:'6',7:'7',8:'8',9:'9',10:'10'})
sns.countplot(x="quality", data=wine_df_copy)
#draw kde plot
wine_df_copy['fixed acidity'].plot.kde()
wine_df_copy['volatile acidity'].plot.kde()
#compare quality with other columns
sns.boxplot(x=wine_df_copy['quality'], y=wine_df_copy['fixed acidity'])
plt.title("fixed acidity vs quality")
sns.boxplot(x=wine_df_copy['quality'], y=wine_df_copy['volatile acidity'])
plt.title("volatile acidity vs quality")
# dividing data into input and output
X = wine_df.drop('quality',axis=1)
y = wine_df['quality']
X.head()
y.head()
#defining the keras model
#importing the required library
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers
#defining the sequential model for binary classification
model = keras.Sequential()
model.add(layers.Dense(11,activation='relu',input_dim=11))
model.add(layers.Dense(5,activation='relu'))
model.add(layers.Dense(10,activation='softmax'))
#compile and fit the model
model.compile(loss='sparse_categorical_crossentropy', metrics=['accuracy'])
history = model.fit(X, y, validation_split=0.22, batch_size=100, epochs=20)
model.summary()
#plot loss vs epochs
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('no of epochs')
plt.legend(['train','test'],loc='upper left')
plt.show()
#plot accuracy vs epochs
plt.plot(history.history['accuracy'])
plt.plot(history.history['val_accuracy'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('no of epochs')
plt.legend(['train','test'],loc='upper left')
plt.show()
import numpy as np
from numpy import array
#input for 11 features
Xnew = np.array([[7.9,0.6,0.06,1.6,0.069,15.0,59.0,0.9964,3.3,0.46,9.4]])
#Xnew = np.array([[8.3,0.655,0.12,2.3,0.083,15.0,113.0,0.9966,3.17,0.66,9.8]])
#convert into numpy array
Xnew = np.array(Xnew, dtype=np.float64)
#making the prediction
Ynew = model.predict(Xnew)
print(Ynew)
max_index_row = np.argmax(Ynew, axis=1)
print(max_index_row)
```
| github_jupyter |
### Data Mining and Machine Learning
### Feature Selection in Supervised Classification
### Edgar Acuna
#### September 2021
##### Modules required: feature_selection from scikit-learn, feature selection from Arizona State University , skrebate, Orange and mRMR
##### Datasets: Diabetes, Bupa, Breast-Cancer-Wisconsin, vehicle, Landsat, Loan and Heart-Cleveland
```
import warnings
warnings.filterwarnings("ignore")
import pandas as pd
import numpy as np
import math as m
from sklearn.model_selection import train_test_split
from sklearn.feature_selection import SelectKBest
from sklearn.feature_selection import chi2, SelectPercentile, f_classif, mutual_info_classif
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
```
#### Example 1: Diabetes dataset
```
url= "http://academic.uprm.edu/eacuna/diabetes.dat"
names = ['preg', 'plas', 'pres', 'skin', 'test', 'mass', 'pedi', 'age', 'class']
data = pd.read_table(url, names=names,header=None)
y=data['class']
X=data.iloc[:,0:8]
```
### I. Feature Selection by Visualization using boxplots
```
print(data.shape)
data.head()
#normalizando los datos
data1=data.drop('class',axis=1)
data1_norm=(data1 - data1.min()) / (data1.max() - data1.min())
data1_norm['class']=data['class']
plt.figure(figsize=(10,50))
df_long = pd.melt(data1_norm, "class", var_name="a", value_name="c")
sns.factorplot("a", hue="class", y="c", data=df_long, kind="box")
```
Comment: From the boxplots we can see that features: plas, age,mass, preg,and pedi have boxplot not so overlaped
## II-Feature selection using statistical measures
### II-1 Using the Chi-Square test
It is desirable to have discretized attributes and two classes. I do not recommend
to use this method because the Chi-Square test is approximated and it is not robust.
```
y=data['class']
X=data.iloc[:,0:8]
# Funcion auxiliar para discretizar cualquier columna de un dataframe
def disc_col_ew(df,str,k,out):
df1=df[str]
bins=np.linspace(df1.min(), df1.max(),k)
if out=="num":
df1=pd.cut(df1,bins=bins,include_lowest=True, right=True,labels=False)
else:
bins[0]=float('-inf')
bins[k-1]=float('inf')
df1=pd.cut(df1,bins=bins,include_lowest=True, right=True)
return df1
# funcion auxiliar para determinar el numero optimo de intervalos segun la formula de scott
def nclass_scott(x):
h=3.5*(np.var(x,ddof=1)**.5)*len(x)**(-.3333)
intervals=m.ceil((max(x)-min(x))/h)
return int(intervals)
#Funcion para discretizar todas las colunmnas de un dataframe
def disc_ew(df,out):
name=df.columns.tolist()
disc=pd.DataFrame()
for name in df.columns.tolist():
k=nclass_scott(df[name])
disc[name]=disc_col_ew(df,name,k,out)
return disc
#discretizando las columnas de la matriz predictora X de diabetes
diab_disc=disc_ew(X,out="num")
#Choosing the best three features with Chi-square and scikit-learn
y1=y.to_numpy()
X1=diab_disc.to_numpy()
test = SelectKBest(score_func=chi2, k=3)
fit = test.fit(X1, y1)
# summarize scores
np.set_printoptions(precision=3)
print(fit.scores_)
#Ranking the features according to their scores
idx = np.argsort(fit.scores_)
print(X.columns[idx[::-1]])
#Printing the subset containing the best three fetaures
features= fit.transform(X1)
print(features)
```
Comentario: Las tres mejores variables con la prueba de Chi-square son plas, age y preg por tener el Chi-Square mas alto.
### II-2 Using the F test to find the best features
Either scikit-learn or the ASU library can be used
```
# Selecion de Features usando los p-values de la F-test como score
# Aqui usamos scikit-learn y se selecciona el 30% de deatures con el mas alto score
selector = SelectPercentile(f_classif, percentile=30)
fit=selector.fit(X, y)
scores = -np.log10(selector.pvalues_)
scores /= scores.max()
print(scores)
features= fit.transform(X)
# Imprime los datos de las tres mejores features
print(features)
```
Comentario: Las tres, mejores variables con la prueba de F son plas,mass y age
```
import numpy as np
from sklearn.feature_selection import f_classif
def f_score(X, y):
"""
This function implements the anova f_value feature selection (existing method for classification in scikit-learn),
where f_score = sum((ni/(c-1))*(mean_i - mean)^2)/((1/(n - c))*sum((ni-1)*std_i^2))
Input
-----
X: {numpy array}, shape (n_samples, n_features)
input data
y : {numpy array},shape (n_samples,)
input class labels
Output
------
F: {numpy array}, shape (n_features,)
f-score for each feature
"""
F, pval = f_classif(X, y)
return F
def feature_ranking(F):
"""
Rank features in descending order according to f-score, the higher the f-score, the more important the feature is
"""
idx = np.argsort(F)
return idx[::-1]
scoref =f_score(X, y)
idx = feature_ranking(scoref)
print(scoref,idx)
```
Comentario: las tres variables mas importantes con la prueba de F son: plas,mass y age
## III Uaing Information measures
### Usando Mutual Information (Entropia)
Using the scikit-learn module
```
# Univariate feature selection with Mutual Information
scores = mutual_info_classif(X,y)
scores /= scores.max()
print(scores)
#Ranking the features according to their scores
idx = np.argsort(scores)
print(X.columns[idx[::-1]])
```
Comentario: Las tres variables usando el criterio de Mutual Information son: plas, mass y age
## IV-Using ReliefF to select the best features
#### IV-1 Usando el modulo skrebate
```
from skrebate import ReliefF
df=data.drop('class',axis=1)
#Normalizando las predictoras
df_norm=(df - df.min()) / (df.max() - df.min())
features, labels = df_norm.values, data['class'].values
features
fs = ReliefF(n_neighbors=10)
fs.fit(features, labels)
for feature_name, feature_score in zip(data.columns,fs.feature_importances_):
print (feature_name, feature_score)
yscore=fs.feature_importances_
oyscore=-np.sort(-yscore)
print(oyscore)
import matplotlib.pyplot as plt
x=np.arange(0,8)
plt.plot(x,oyscore,'k-',lw=3)
```
Comentario: las tres variables mas importantes con el ReliefF son: plas,mass y skin
#### IV-2 Usando el Relief de Orange
```
import Orange
df = Orange.data.Table("https://academic.uprm.edu/eacuna/diabetes.tab")
#ReliefF usando 10 vecinos mas cercanos y una muestrd m=100 para updating de los pesos
print('Feature scores for best features (scored individually):')
meas =Orange.preprocess.score.ReliefF(n_iterations=400,k_nearest=10)
scores = meas(df)
for attr, score in zip(df.domain.attributes, scores):
print('%.3f' % score, attr.name)
#Ranking the features according to their scores
idx = np.argsort(scores)
print(X.columns[idx[::-1]])
```
The best features are: plas, mass and skin.
### Example 2: Bupa dataset
```
#load data
url= "http://academic.uprm.edu/eacuna/bupa.dat"
names = ['mcv', 'alkphos', 'sgpt', 'aspar', 'gammagt', 'drinks', 'class']
data = pd.read_table(url, names=names,header=None)
y=data['class']
X=data.iloc[:,0:6]
y1=y.to_numpy()
X1=X.to_numpy()
features, labels = X.values, data['class'].values
```
#### Using skrebate
```
df=data.drop('class',axis=1)
#Normalizando las predictoras
df_norm=(df - df.min()) / (df.max() - df.min())
features, labels = df_norm.values, data['class'].values
df.info()
fs = ReliefF(n_neighbors=10)
fs.fit(features, labels)
for feature_name, feature_score in zip(data.columns,fs.feature_importances_):
print (feature_name, feature_score)
```
El relielf de skrebate recomienda gammagt, aspar and sgpt
### Using the ReliefF from Orange
```
import Orange
df = Orange.data.Table("https://datasets.biolab.si/core/bupa.tab")
#ReliefF usando 10 vecinos mas cercanos y una muestra de n=300 para updating de los pesos
print('Feature scores for best ten features (scored individually):')
meas =Orange.preprocess.score.ReliefF(n_iterations=300,k_nearest=10)
scores = meas(df)
for attr, score in zip(df.domain.attributes, scores):
print('%.4f' % score, attr.name)
```
### Example 3: Breast-cancer-Wisconsin
```
names = [
'ID','Clump', 'Unif_Cell_Size', 'Unif_Cell_Shape', 'Marginal_Adh', 'Single_Cell_Size', 'Bare_Nuclei', 'Bland_Chromatine', 'Normal_Nucleoi', 'Mitoses','class']
data = pd.read_table("https://academic.uprm.edu/eacuna/breast-cancer-wisconsin.data", names=names, header=None,sep=",",na_values="?")
data.info()
```
### Using skrebate
```
#Dropping the ID column
df=data.drop('ID',axis=1)
#Dropping the class column
df1=df.drop('class',axis=1)
#Normalizando las predictoras
df_norm=(df1 - df1.min()) / (df1.max() - df1.min())
features, labels = df_norm.values, df['class'].values
fs = ReliefF(n_neighbors=10)
fs.fit(features, labels)
for feature_name, feature_score in zip(df1.columns,fs.feature_importances_):
print (feature_name, feature_score)
yscore=fs.feature_importances_
oyscore=-np.sort(-yscore)
print(oyscore)
x=np.arange(0,9)
plt.plot(x,oyscore,'k-',lw=3)
```
### Using Relief from Orange
```
import Orange
df = Orange.data.Table("https://academic.uprm.edu/eacuna/breast-cancer-wisconsin.tab")
#ReliefF usando 10 vecinos mas cercanos y una muestrd m=400 para updating de los pesos
print('Feature scores for best ten features (scored individually):')
meas =Orange.preprocess.score.ReliefF(n_iterations=500,k_nearest=1)
scores = meas(df)
for attr, score in zip(df.domain.attributes, scores):
print('%.3f' % score, attr.name)
```
Comment:The best attributes are:6(Bare Nuclei),1 (Clump Thickness),7 (Bland Chromatin), 8 (Normal. nucleoli)
## V. Relief for multiclass problem
### Example 4: The vehicle dataset
```
df1=pd.read_csv("http://academic.uprm.edu/eacuna/vehicle.csv")
df1.info()
#Convirtiendo en matriz la tabla de predictoras y la columna de clases
y=df1['Class']
X=df1.iloc[:,0:18]
y1=y.values
X1=X.values
#Normalizando las predictoras
X_norm=(X - X.min()) / (X.max() - X.min())
features, labels = X_norm.values, y1
```
### Using skrebate
```
fs = ReliefF(n_neighbors=10)
fs.fit(features, labels)
for feature_name, feature_score in zip(X.columns,fs.feature_importances_):
print (feature_name, feature_score)
idx = np.argsort(fs.feature_importances_)
print(X.columns[idx[::-1]])
yscore=fs.feature_importances_
oyscore=-np.sort(-yscore)
print(oyscore)
x=np.arange(0,18)
plt.plot(x,oyscore,'k-',lw=3)
```
#### The best features are: 'ELONGATEDNESS', 'SCATTER_RATIO', 'SCALED_VARIANCE_MINOR', 'HOLLOWS_RATIO', 'PR.AXIS_RECTANGULARITY', 'DISTANCE_CIRCULARITY', 'MAX.LENGTH_RECTANGULARITY', 'SCALED_VARIANCE_MAJOR'
### Example 5: Landsat Dataset
```
names=['a1','a2','a3','a4','a5','a6','a7','a8','a9','a10','a11','a12','a13','a14','a15','a16','a17','a18','a19','a20','a21','a22','a23','a24','a25','a26','a27','a38','a29','a30','a31','a32','a33','a34','a35','a36','class']
url='http://academic.uprm.edu/eacuna/landsat.txt'
data = pd.read_csv(url,names=names, header=None,delim_whitespace=True)
data.head()
y=data['class']
X=data.iloc[:,0:36]
y1=y.to_numpy()
X1=X.to_numpy()
features, labels = X.values, data['class'].values
df=data.drop('class',axis=1)
#Normalizando las predictoras
df_norm=(df - df.min()) / (df.max() - df.min())
features, labels = df_norm.values, data['class'].values
df.shape
fs = ReliefF(n_neighbors=10)
fs.fit(features, labels)
for feature_name, feature_score in zip(data.columns,fs.feature_importances_):
print (feature_name, feature_score)
idx = np.argsort(fs.feature_importances_)
print(df.columns[idx[::-1]])
```
Comment:The best attributes are:a13,a25,a21,a17,a33,a29,a9,a5,a1,a30
### Using Relief from Orange
```
#Usando el Relief de Orange
import Orange
df = Orange.data.Table("https://academic.uprm.edu/eacuna/landsat.tab")
print('Feature scores (scored individually):')
#ReliefF usando 10 vecinos mas cercanos y una muestrd m=3000 para updating de los pesos
meas =Orange.preprocess.score.ReliefF(n_iterations=3000,k_nearest=10)
scores = meas(df)
for attr, score in zip(df.domain.attributes, scores):
print('%.3f' % score, attr.name)
idx = np.argsort(scores)
print(idx[::-1])
```
Comment:The best attributes are:a13,a25,a17,a21,a33,a29,a1,a5,a9
## VI. Applying Relief to a dataset with mixed type of attributes
#### VI-1 The Loan Dataset
```
import Orange
df = Orange.data.Table("https://academic.uprm.edu/eacuna/loan.tab")
df
meas =Orange.preprocess.score.ReliefF(n_iterations=2000,k_nearest=10)
scores = meas(df)
for attr, score in zip(df.domain.attributes, scores):
print('%.3f' % score, attr.name)
```
### VI-2 The Heart-Cleveland Dataset
```
#Usando el Relief de Orange
import Orange
df = Orange.data.Table("https://academic.uprm.edu/eacuna/heart_disease.tab")
print('Feature scores (scored individually):')
#ReliefF usando 10 vecinos mas cercanos y una muestrd m=200 ara updating de los pesos
meas =Orange.preprocess.score.ReliefF(n_iterations=200,k_nearest=10)
scores = meas(df)
for attr, score in zip(df.domain.attributes, scores):
print('%.3f' % score, attr.name)
idx = np.argsort(scores)
print(idx[::-1])
```
The best features are: chest pain, thal, major vessels colored, gender
### VII. Feature selection using Minumum Redudancy and Maximum Relevancy (mRMR)
#### mRMR applied to landsat
```
import pandas as pd
from mrmr import mrmr_classif
# Chhosing teh top ten best features
mrmr = mrmr_classif(X,y, K = 36)
print(mrmr)
```
| github_jupyter |
```
from google.colab import drive
drive.mount('/content/drive')
```
### Dependencies
```
!pip install keras-rectified-adam
!pip install segmentation-models
import os
import cv2
import math
import random
import shutil
import warnings
import numpy as np
import pandas as pd
import seaborn as sns
import multiprocessing as mp
import albumentations as albu
from keras_radam import RAdam
import matplotlib.pyplot as plt
import segmentation_models as sm
from tensorflow import set_random_seed
from sklearn.model_selection import train_test_split
from keras import optimizers
from keras import backend as K
from keras.utils import Sequence
from keras.losses import binary_crossentropy
from keras.preprocessing.image import ImageDataGenerator
from keras.callbacks import EarlyStopping, ReduceLROnPlateau, ModelCheckpoint, Callback
def seed_everything(seed=0):
random.seed(seed)
os.environ['PYTHONHASHSEED'] = str(seed)
np.random.seed(seed)
set_random_seed(seed)
seed = 0
seed_everything(seed)
warnings.filterwarnings("ignore")
base_path = '/content/drive/My Drive/Colab Notebooks/[Kaggle] Understanding Clouds from Satellite Images/Data/'
model_base_path = '/content/drive/My Drive/Colab Notebooks/[Kaggle] Understanding Clouds from Satellite Images/Models/files/'
submission_base_path = '/content/drive/My Drive/Colab Notebooks/[Kaggle] Understanding Clouds from Satellite Images/Data/submissions/'
train_path = base_path + 'train.csv'
test_path = base_path + 'sample_submission.csv'
hold_out_set_path = base_path + 'hold-out.csv'
train_images_dest_path = base_path + 'train_images/'
validation_images_dest_path = base_path + 'validation_images/'
test_images_dest_path = base_path + 'test_images/'
```
### Load data
```
train = pd.read_csv(train_path)
submission = pd.read_csv(test_path)
hold_out_set = pd.read_csv(hold_out_set_path)
X_train = hold_out_set[hold_out_set['set'] == 'train']
X_val = hold_out_set[hold_out_set['set'] == 'validation']
print('Compete set samples:', len(train))
print('Train samples: ', len(X_train))
print('Validation samples: ', len(X_val))
print('Test samples:', len(submission))
# Preprocecss data
train['image'] = train['Image_Label'].apply(lambda x: x.split('_')[0])
submission['image'] = submission['Image_Label'].apply(lambda x: x.split('_')[0])
test = pd.DataFrame(submission['image'].unique(), columns=['image'])
test['set'] = 'test'
display(X_train.head())
```
# Model parameters
```
BACKBONE = 'densenet121'
BATCH_SIZE = 8
EPOCHS = 20
LEARNING_RATE = 3e-4
HEIGHT = 256
WIDTH = 384
CHANNELS = 3
N_CLASSES = 4
ES_PATIENCE = 5
RLROP_PATIENCE = 3
DECAY_DROP = 0.5
model_name = 'uNet_%s_%sx%s' % (BACKBONE, HEIGHT, WIDTH)
model_path = model_base_path + '%s.h5' % (model_name)
submission_path = submission_base_path + '%s_submission.csv' % (model_name)
submission_post_path = submission_base_path + '%s_submission_post.csv' % (model_name)
preprocessing = sm.backbones.get_preprocessing(BACKBONE)
augmentation = albu.Compose([albu.HorizontalFlip(p=0.5),
albu.VerticalFlip(p=0.5),
albu.GridDistortion(p=0.5),
albu.ShiftScaleRotate(scale_limit=0.5, rotate_limit=0,
shift_limit=0.1, border_mode=0, p=0.5),
albu.OpticalDistortion(p=0.5, distort_limit=2, shift_limit=0.5)
])
```
### Auxiliary functions
```
#@title
def np_resize(img, input_shape):
height, width = input_shape
return cv2.resize(img, (width, height))
def mask2rle(img):
pixels= img.T.flatten()
pixels = np.concatenate([[0], pixels, [0]])
runs = np.where(pixels[1:] != pixels[:-1])[0] + 1
runs[1::2] -= runs[::2]
return ' '.join(str(x) for x in runs)
def build_rles(masks, reshape=None):
width, height, depth = masks.shape
rles = []
for i in range(depth):
mask = masks[:, :, i]
if reshape:
mask = mask.astype(np.float32)
mask = np_resize(mask, reshape).astype(np.int64)
rle = mask2rle(mask)
rles.append(rle)
return rles
def build_masks(rles, input_shape, reshape=None):
depth = len(rles)
if reshape is None:
masks = np.zeros((*input_shape, depth))
else:
masks = np.zeros((*reshape, depth))
for i, rle in enumerate(rles):
if type(rle) is str:
if reshape is None:
masks[:, :, i] = rle2mask(rle, input_shape)
else:
mask = rle2mask(rle, input_shape)
reshaped_mask = np_resize(mask, reshape)
masks[:, :, i] = reshaped_mask
return masks
def rle2mask(rle, input_shape):
width, height = input_shape[:2]
mask = np.zeros( width*height ).astype(np.uint8)
array = np.asarray([int(x) for x in rle.split()])
starts = array[0::2]
lengths = array[1::2]
current_position = 0
for index, start in enumerate(starts):
mask[int(start):int(start+lengths[index])] = 1
current_position += lengths[index]
return mask.reshape(height, width).T
def dice_coefficient(y_true, y_pred):
y_true = np.asarray(y_true).astype(np.bool)
y_pred = np.asarray(y_pred).astype(np.bool)
intersection = np.logical_and(y_true, y_pred)
return (2. * intersection.sum()) / (y_true.sum() + y_pred.sum())
def dice_coef(y_true, y_pred, smooth=1):
y_true_f = K.flatten(y_true)
y_pred_f = K.flatten(y_pred)
intersection = K.sum(y_true_f * y_pred_f)
return (2. * intersection + smooth) / (K.sum(y_true_f) + K.sum(y_pred_f) + smooth)
def post_process(probability, threshold=0.5, min_size=10000):
mask = cv2.threshold(probability, threshold, 1, cv2.THRESH_BINARY)[1]
num_component, component = cv2.connectedComponents(mask.astype(np.uint8))
predictions = np.zeros(probability.shape, np.float32)
for c in range(1, num_component):
p = (component == c)
if p.sum() > min_size:
predictions[p] = 1
return predictions
def get_metrics(model, df, df_images_dest_path, tresholds, min_mask_sizes, set_name='Complete set'):
class_names = ['Fish', 'Flower', 'Gravel', 'Sugar']
metrics = []
for class_name in class_names:
metrics.append([class_name, 0, 0])
metrics_df = pd.DataFrame(metrics, columns=['Class', 'Dice', 'Dice Post'])
for i in range(0, df.shape[0], 500):
batch_idx = list(range(i, min(df.shape[0], i + 500)))
batch_set = df[batch_idx[0]: batch_idx[-1]+1]
ratio = len(batch_set) / len(df)
generator = DataGenerator(
directory=df_images_dest_path,
dataframe=batch_set,
target_df=train,
batch_size=len(batch_set),
target_size=(HEIGHT, WIDTH),
n_channels=CHANNELS,
n_classes=N_CLASSES,
preprocessing=preprocessing,
seed=seed,
mode='fit',
shuffle=False)
x, y = generator.__getitem__(0)
preds = model.predict(x)
for class_index in range(N_CLASSES):
class_score = []
class_score_post = []
mask_class = y[..., class_index]
pred_class = preds[..., class_index]
for index in range(len(batch_idx)):
sample_mask = mask_class[index, ]
sample_pred = pred_class[index, ]
sample_pred_post = post_process(sample_pred, threshold=tresholds[class_index], min_size=min_mask_sizes[class_index])
if (sample_mask.sum() == 0) & (sample_pred.sum() == 0):
dice_score = 1.
else:
dice_score = dice_coefficient(sample_pred, sample_mask)
if (sample_mask.sum() == 0) & (sample_pred_post.sum() == 0):
dice_score_post = 1.
else:
dice_score_post = dice_coefficient(sample_pred_post, sample_mask)
class_score.append(dice_score)
class_score_post.append(dice_score_post)
metrics_df.loc[metrics_df['Class'] == class_names[class_index], 'Dice'] += np.mean(class_score) * ratio
metrics_df.loc[metrics_df['Class'] == class_names[class_index], 'Dice Post'] += np.mean(class_score_post) * ratio
metrics_df = metrics_df.append({'Class':set_name, 'Dice':np.mean(metrics_df['Dice'].values), 'Dice Post':np.mean(metrics_df['Dice Post'].values)}, ignore_index=True).set_index('Class')
return metrics_df
def plot_metrics(history):
fig, axes = plt.subplots(4, 1, sharex='col', figsize=(22, 14))
axes = axes.flatten()
axes[0].plot(history['loss'], label='Train loss')
axes[0].plot(history['val_loss'], label='Validation loss')
axes[0].legend(loc='best')
axes[0].set_title('Loss')
axes[1].plot(history['iou_score'], label='Train IOU Score')
axes[1].plot(history['val_iou_score'], label='Validation IOU Score')
axes[1].legend(loc='best')
axes[1].set_title('IOU Score')
axes[2].plot(history['dice_coef'], label='Train Dice coefficient')
axes[2].plot(history['val_dice_coef'], label='Validation Dice coefficient')
axes[2].legend(loc='best')
axes[2].set_title('Dice coefficient')
axes[3].plot(history['score'], label='Train F-Score')
axes[3].plot(history['val_score'], label='Validation F-Score')
axes[3].legend(loc='best')
axes[3].set_title('F-Score')
plt.xlabel('Epochs')
sns.despine()
plt.show()
def cosine_decay_with_warmup(global_step,
learning_rate_base,
total_steps,
warmup_learning_rate=0.0,
warmup_steps=0,
hold_base_rate_steps=0):
"""
Cosine decay schedule with warm up period.
In this schedule, the learning rate grows linearly from warmup_learning_rate
to learning_rate_base for warmup_steps, then transitions to a cosine decay
schedule.
:param global_step {int}: global step.
:param learning_rate_base {float}: base learning rate.
:param total_steps {int}: total number of training steps.
:param warmup_learning_rate {float}: initial learning rate for warm up. (default: {0.0}).
:param warmup_steps {int}: number of warmup steps. (default: {0}).
:param hold_base_rate_steps {int}: Optional number of steps to hold base learning rate before decaying. (default: {0}).
:param global_step {int}: global step.
:Returns : a float representing learning rate.
:Raises ValueError: if warmup_learning_rate is larger than learning_rate_base, or if warmup_steps is larger than total_steps.
"""
if total_steps < warmup_steps:
raise ValueError('total_steps must be larger or equal to warmup_steps.')
learning_rate = 0.5 * learning_rate_base * (1 + np.cos(
np.pi *
(global_step - warmup_steps - hold_base_rate_steps
) / float(total_steps - warmup_steps - hold_base_rate_steps)))
if hold_base_rate_steps > 0:
learning_rate = np.where(global_step > warmup_steps + hold_base_rate_steps,
learning_rate, learning_rate_base)
if warmup_steps > 0:
if learning_rate_base < warmup_learning_rate:
raise ValueE32rror('learning_rate_base must be larger or equal to warmup_learning_rate.')
slope = (learning_rate_base - warmup_learning_rate) / warmup_steps
warmup_rate = slope * global_step + warmup_learning_rate
learning_rate = np.where(global_step < warmup_steps, warmup_rate,
learning_rate)
return np.where(global_step > total_steps, 0.0, learning_rate)
class WarmUpCosineDecayScheduler(Callback):
"""Cosine decay with warmup learning rate scheduler"""
def __init__(self,
learning_rate_base,
total_steps,
global_step_init=0,
warmup_learning_rate=0.0,
warmup_steps=0,
hold_base_rate_steps=0,
verbose=0):
"""
Constructor for cosine decay with warmup learning rate scheduler.
:param learning_rate_base {float}: base learning rate.
:param total_steps {int}: total number of training steps.
:param global_step_init {int}: initial global step, e.g. from previous checkpoint.
:param warmup_learning_rate {float}: initial learning rate for warm up. (default: {0.0}).
:param warmup_steps {int}: number of warmup steps. (default: {0}).
:param hold_base_rate_steps {int}: Optional number of steps to hold base learning rate before decaying. (default: {0}).
:param verbose {int}: quiet, 1: update messages. (default: {0}).
"""
super(WarmUpCosineDecayScheduler, self).__init__()
self.learning_rate_base = learning_rate_base
self.total_steps = total_steps
self.global_step = global_step_init
self.warmup_learning_rate = warmup_learning_rate
self.warmup_steps = warmup_steps
self.hold_base_rate_steps = hold_base_rate_steps
self.verbose = verbose
self.learning_rates = []
def on_batch_end(self, batch, logs=None):
self.global_step = self.global_step + 1
lr = K.get_value(self.model.optimizer.lr)
self.learning_rates.append(lr)
def on_batch_begin(self, batch, logs=None):
lr = cosine_decay_with_warmup(global_step=self.global_step,
learning_rate_base=self.learning_rate_base,
total_steps=self.total_steps,
warmup_learning_rate=self.warmup_learning_rate,
warmup_steps=self.warmup_steps,
hold_base_rate_steps=self.hold_base_rate_steps)
K.set_value(self.model.optimizer.lr, lr)
if self.verbose > 0:
print('\nBatch %02d: setting learning rate to %s.' % (self.global_step + 1, lr))
```
### Data generator
```
#@title
class DataGenerator(Sequence):
def __init__(self, dataframe, target_df=None, mode='fit', directory=train_images_dest_path,
batch_size=BATCH_SIZE, n_channels=CHANNELS, target_size=(HEIGHT, WIDTH),
n_classes=N_CLASSES, seed=seed, shuffle=True, preprocessing=None, augmentation=None):
self.batch_size = batch_size
self.dataframe = dataframe
self.mode = mode
self.directory = directory
self.target_df = target_df
self.target_size = target_size
self.n_channels = n_channels
self.n_classes = n_classes
self.shuffle = shuffle
self.augmentation = augmentation
self.preprocessing = preprocessing
self.seed = seed
self.mask_shape = (1400, 2100)
self.list_IDs = self.dataframe.index
if self.seed is not None:
np.random.seed(self.seed)
self.on_epoch_end()
def __len__(self):
return len(self.list_IDs) // self.batch_size
def __getitem__(self, index):
indexes = self.indexes[index*self.batch_size:(index+1)*self.batch_size]
list_IDs_batch = [self.list_IDs[k] for k in indexes]
X = self.__generate_X(list_IDs_batch)
if self.mode == 'fit':
Y = self.__generate_Y(list_IDs_batch)
if self.augmentation:
X, Y = self.__augment_batch(X, Y)
return X, Y
elif self.mode == 'predict':
return X
def on_epoch_end(self):
self.indexes = np.arange(len(self.list_IDs))
if self.shuffle == True:
np.random.shuffle(self.indexes)
def __generate_X(self, list_IDs_batch):
X = np.empty((self.batch_size, *self.target_size, self.n_channels))
for i, ID in enumerate(list_IDs_batch):
img_name = self.dataframe['image'].loc[ID]
img_path = self.directory + img_name
img = cv2.imread(img_path)
img = cv2.cvtColor(img, cv2.COLOR_BGR2RGB)
if self.preprocessing:
img = self.preprocessing(img)
X[i,] = img
return X
def __generate_Y(self, list_IDs_batch):
Y = np.empty((self.batch_size, *self.target_size, self.n_classes), dtype=int)
for i, ID in enumerate(list_IDs_batch):
img_name = self.dataframe['image'].loc[ID]
image_df = self.target_df[self.target_df['image'] == img_name]
rles = image_df['EncodedPixels'].values
masks = build_masks(rles, input_shape=self.mask_shape, reshape=self.target_size)
Y[i, ] = masks
return Y
def __augment_batch(self, X_batch, Y_batch):
for i in range(X_batch.shape[0]):
X_batch[i, ], Y_batch[i, ] = self.__random_transform(X_batch[i, ], Y_batch[i, ])
return X_batch, Y_batch
def __random_transform(self, X, Y):
composed = self.augmentation(image=X, mask=Y)
X_aug = composed['image']
Y_aug = composed['mask']
return X_aug, Y_aug
train_generator = DataGenerator(
directory=train_images_dest_path,
dataframe=X_train,
target_df=train,
batch_size=BATCH_SIZE,
target_size=(HEIGHT, WIDTH),
n_channels=CHANNELS,
n_classes=N_CLASSES,
preprocessing=preprocessing,
augmentation=augmentation,
seed=seed)
valid_generator = DataGenerator(
directory=validation_images_dest_path,
dataframe=X_val,
target_df=train,
batch_size=BATCH_SIZE,
target_size=(HEIGHT, WIDTH),
n_channels=CHANNELS,
n_classes=N_CLASSES,
preprocessing=preprocessing,
seed=seed)
```
# Model
```
model = sm.Unet(backbone_name=BACKBONE,
encoder_weights='imagenet',
classes=N_CLASSES,
activation='sigmoid',
input_shape=(HEIGHT, WIDTH, CHANNELS))
checkpoint = ModelCheckpoint(model_path, monitor='val_loss', mode='min', save_best_only=True, save_weights_only=True)
es = EarlyStopping(monitor='val_loss', mode='min', patience=ES_PATIENCE, restore_best_weights=True, verbose=1)
rlrop = ReduceLROnPlateau(monitor='val_loss', mode='min', patience=RLROP_PATIENCE, factor=DECAY_DROP, min_lr=1e-6, verbose=1)
metric_list = [dice_coef, sm.metrics.iou_score, sm.metrics.f1_score]
callback_list = [checkpoint, es, rlrop]
optimizer = RAdam(learning_rate=LEARNING_RATE, warmup_proportion=0.1)
model.compile(optimizer=optimizer, loss=sm.losses.bce_dice_loss, metrics=metric_list)
model.summary()
model.load_weights(model_path)
STEP_SIZE_TRAIN = len(X_train)//BATCH_SIZE
STEP_SIZE_VALID = len(X_val)//BATCH_SIZE
history = model.fit_generator(generator=train_generator,
steps_per_epoch=STEP_SIZE_TRAIN,
validation_data=valid_generator,
validation_steps=STEP_SIZE_VALID,
callbacks=callback_list,
epochs=EPOCHS,
verbose=1).history
```
## Model loss graph
```
#@title
plot_metrics(history)
```
# Threshold and mask size tunning
```
#@title
class_names = ['Fish ', 'Flower', 'Gravel', 'Sugar ']
mask_grid = [0, 500, 1000, 5000, 7500, 10000, 15000]
threshold_grid = np.arange(.5, 1, .05)
metrics = []
for class_index in range(N_CLASSES):
for threshold in threshold_grid:
for mask_size in mask_grid:
metrics.append([class_index, threshold, mask_size, 0])
metrics_df = pd.DataFrame(metrics, columns=['Class', 'Threshold', 'Mask size', 'Dice'])
for i in range(0, X_val.shape[0], 500):
batch_idx = list(range(i, min(X_val.shape[0], i + 500)))
batch_set = X_val[batch_idx[0]: batch_idx[-1]+1]
ratio = len(batch_set) / len(X_val)
generator = DataGenerator(
directory=validation_images_dest_path,
dataframe=batch_set,
target_df=train,
batch_size=len(batch_set),
target_size=(HEIGHT, WIDTH),
n_channels=CHANNELS,
n_classes=N_CLASSES,
preprocessing=preprocessing,
seed=seed,
mode='fit',
shuffle=False)
x, y = generator.__getitem__(0)
preds = model.predict(x)
for class_index in range(N_CLASSES):
class_score = []
label_class = y[..., class_index]
pred_class = preds[..., class_index]
for threshold in threshold_grid:
for mask_size in mask_grid:
mask_score = []
for index in range(len(batch_idx)):
label_mask = label_class[index, ]
pred_mask = pred_class[index, ]
pred_mask = post_process(pred_mask, threshold=threshold, min_size=mask_size)
dice_score = dice_coefficient(pred_mask, label_mask)
if (pred_mask.sum() == 0) & (label_mask.sum() == 0):
dice_score = 1.
mask_score.append(dice_score)
metrics_df.loc[(metrics_df['Class'] == class_index) & (metrics_df['Threshold'] == threshold) &
(metrics_df['Mask size'] == mask_size), 'Dice'] += np.mean(mask_score) * ratio
metrics_df_0 = metrics_df[metrics_df['Class'] == 0]
metrics_df_1 = metrics_df[metrics_df['Class'] == 1]
metrics_df_2 = metrics_df[metrics_df['Class'] == 2]
metrics_df_3 = metrics_df[metrics_df['Class'] == 3]
optimal_values_0 = metrics_df_0.loc[metrics_df_0['Dice'].idxmax()].values
optimal_values_1 = metrics_df_1.loc[metrics_df_1['Dice'].idxmax()].values
optimal_values_2 = metrics_df_2.loc[metrics_df_2['Dice'].idxmax()].values
optimal_values_3 = metrics_df_3.loc[metrics_df_3['Dice'].idxmax()].values
best_tresholds = [optimal_values_0[1], optimal_values_1[1], optimal_values_2[1], optimal_values_3[1]]
best_masks = [optimal_values_0[2], optimal_values_1[2], optimal_values_2[2], optimal_values_3[2]]
best_dices = [optimal_values_0[3], optimal_values_1[3], optimal_values_2[3], optimal_values_3[3]]
for index, name in enumerate(class_names):
print('%s treshold=%.2f mask size=%d Dice=%.3f' % (name, best_tresholds[index], best_masks[index], best_dices[index]))
```
# Model evaluation
```
#@title
train_metrics = get_metrics(model, X_train, train_images_dest_path, best_tresholds, best_masks, 'Train')
display(train_metrics)
validation_metrics = get_metrics(model, X_val, validation_images_dest_path, best_tresholds, best_masks, 'Validation')
display(validation_metrics)
```
# Apply model to test set
```
#@title
test_df = []
for i in range(0, test.shape[0], 500):
batch_idx = list(range(i, min(test.shape[0], i + 500)))
batch_set = test[batch_idx[0]: batch_idx[-1]+1]
test_generator = DataGenerator(
directory=test_images_dest_path,
dataframe=batch_set,
target_df=submission,
batch_size=1,
target_size=(HEIGHT, WIDTH),
n_channels=CHANNELS,
n_classes=N_CLASSES,
preprocessing=preprocessing,
seed=seed,
mode='predict',
shuffle=False)
preds = model.predict_generator(test_generator)
for index, b in enumerate(batch_idx):
filename = test['image'].iloc[b]
image_df = submission[submission['image'] == filename].copy()
pred_masks = preds[index, ].round().astype(int)
pred_rles = build_rles(pred_masks, reshape=(350, 525))
image_df['EncodedPixels'] = pred_rles
### Post procecssing
pred_masks_post = preds[index, ].astype('float32')
for class_index in range(N_CLASSES):
pred_mask = pred_masks_post[...,class_index]
pred_mask = post_process(pred_mask, threshold=best_tresholds[class_index], min_size=best_masks[class_index])
pred_masks_post[...,class_index] = pred_mask
pred_rles_post = build_rles(pred_masks_post, reshape=(350, 525))
image_df['EncodedPixels_post'] = pred_rles_post
###
test_df.append(image_df)
sub_df = pd.concat(test_df)
```
### Regular submission
```
#@title
submission_df = sub_df[['Image_Label' ,'EncodedPixels']]
submission_df.to_csv(submission_path, index=False)
display(submission_df.head())
```
### Submission with post processing
```
#@title
submission_df_post = sub_df[['Image_Label' ,'EncodedPixels_post']]
submission_df_post.columns = ['Image_Label' ,'EncodedPixels']
submission_df_post.to_csv(submission_post_path, index=False)
display(submission_df_post.head())
```
| github_jupyter |
# SageMaker Data Wrangler Job Notebook
This notebook uses the Data Wrangler .flow file to submit a SageMaker Data Wrangler Job
with the following steps:
* Push Data Wrangler .flow file to S3
* Parse the .flow file inputs, and create the argument dictionary to submit to a boto client
* Submit the ProcessingJob arguments and wait for Job completion
Optionally, the notebook also gives an example of starting a SageMaker XGBoost TrainingJob using
the newly processed data.
```
# SageMaker Python SDK version 2.x is required
import pkg_resources
import subprocess
import sys
original_version = pkg_resources.get_distribution("sagemaker").version
_ = subprocess.check_call(
[sys.executable, "-m", "pip", "install", "sagemaker==2.20.0"]
)
import json
import os
import time
import uuid
import boto3
import sagemaker
```
## Parameters
The following lists configurable parameters that are used throughout this notebook.
```
# S3 bucket for saving processing job outputs
# Feel free to specify a different bucket here if you wish.
sess = sagemaker.Session()
bucket = sess.default_bucket()
prefix = "data_wrangler_flows"
flow_id = f"{time.strftime('%d-%H-%M-%S', time.gmtime())}-{str(uuid.uuid4())[:8]}"
flow_name = f"flow-{flow_id}"
flow_uri = f"s3://{bucket}/{prefix}/{flow_name}.flow"
flow_file_name = "customers.flow"
iam_role = sagemaker.get_execution_role()
container_uri = "663277389841.dkr.ecr.us-east-1.amazonaws.com/sagemaker-data-wrangler-container:1.x"
# Processing Job Resources Configurations
instance_count = 1
instance_type = "ml.m5.4xlarge"
# Network Isolation mode; default is off
enable_network_isolation = False
# Processing Job Path URI Information
output_prefix = f"export-{flow_name}/output"
output_path = f"s3://{bucket}/{output_prefix}"
output_name = "dc0ba3db-3a12-49ef-8b39-a7b7867e295b.default"
processing_job_name = f"data-wrangler-flow-processing-{flow_id}"
processing_dir = "/opt/ml/processing"
# Modify the variable below to specify the content type to be used for writing each output
# Currently supported options are 'CSV' or 'PARQUET', and default to 'CSV'
output_content_type = "CSV"
# URL to use for sagemaker client.
# If this is None, boto will automatically construct the appropriate URL to use
# when communicating with sagemaker.
sagemaker_endpoint_url = None
```
## Push Flow to S3
Use the following cell to upload the Data Wrangler .flow file to Amazon S3 so that
it can be used as an input to the processing job.
```
# Load .flow file
with open(flow_file_name) as f:
flow = json.load(f)
# Upload to S3
s3_client = boto3.client("s3")
s3_client.upload_file(flow_file_name, bucket, f"{prefix}/{flow_name}.flow")
print(f"Data Wrangler Flow notebook uploaded to {flow_uri}")
```
## Create Processing Job arguments
This notebook submits a Processing Job using the Sagmaker Python SDK. Below, utility methods are
defined for creating Processing Job Inputs for the following sources: S3, Athena, and Redshift.
```
from sagemaker.processing import ProcessingInput, ProcessingOutput
from sagemaker.dataset_definition.inputs import AthenaDatasetDefinition, DatasetDefinition, RedshiftDatasetDefinition
def create_flow_notebook_processing_input(base_dir, flow_s3_uri):
return ProcessingInput(
source=flow_s3_uri,
destination=f"{base_dir}/flow",
input_name="flow",
s3_data_type="S3Prefix",
s3_input_mode="File",
s3_data_distribution_type="FullyReplicated",
)
def create_s3_processing_input(s3_dataset_definition, name, base_dir):
return ProcessingInput(
source=s3_dataset_definition['s3ExecutionContext']['s3Uri'],
destination=f"{base_dir}/{name}",
input_name=name,
s3_data_type="S3Prefix",
s3_input_mode="File",
s3_data_distribution_type="FullyReplicated",
)
def create_athena_processing_input(athena_dataset_defintion, name, base_dir):
return ProcessingInput(
input_name=name,
dataset_definition=DatasetDefinition(
local_path=f"{base_dir}/{name}",
data_distribution_type="FullyReplicated",
athena_dataset_definition=AthenaDatasetDefinition(
catalog=athena_dataset_defintion["catalogName"],
database=athena_dataset_defintion["databaseName"],
query_string=athena_dataset_defintion["queryString"],
output_s3_uri=athena_dataset_defintion["s3OutputLocation"] + f"{name}/",
output_format=athena_dataset_defintion["outputFormat"].upper()
)
)
)
def create_redshift_processing_input(redshift_dataset_defintion, name, base_dir):
return ProcessingInput(
input_name=name,
dataset_definition=DatasetDefinition(
local_path=f"{base_dir}/{name}",
data_distribution_type="FullyReplicated",
redshift_dataset_definition=RedshiftDatasetDefinition(
cluster_id=redshift_dataset_defintion["clusterIdentifier"],
database=redshift_dataset_defintion["database"],
db_user=redshift_dataset_defintion["dbUser"],
query_string=redshift_dataset_defintion["queryString"],
cluster_role_arn=redshift_dataset_defintion["unloadIamRole"],
output_s3_uri=redshift_dataset_defintion["s3OutputLocation"] + f"{name}/",
output_format=redshift_dataset_defintion["outputFormat"].upper()
)
)
)
def create_processing_inputs(processing_dir, flow, flow_uri):
"""Helper function for creating processing inputs
:param flow: loaded data wrangler flow notebook
:param flow_uri: S3 URI of the data wrangler flow notebook
"""
processing_inputs = []
flow_processing_input = create_flow_notebook_processing_input(processing_dir, flow_uri)
processing_inputs.append(flow_processing_input)
for node in flow["nodes"]:
if "dataset_definition" in node["parameters"]:
data_def = node["parameters"]["dataset_definition"]
name = data_def["name"]
source_type = data_def["datasetSourceType"]
if source_type == "S3":
processing_inputs.append(create_s3_processing_input(data_def, name, processing_dir))
elif source_type == "Athena":
processing_inputs.append(create_athena_processing_input(data_def, name, processing_dir))
elif source_type == "Redshift":
processing_inputs.append(create_redshift_processing_input(data_def, name, processing_dir))
else:
raise ValueError(f"{source_type} is not supported for Data Wrangler Processing.")
return processing_inputs
def create_processing_output(output_name, output_path, processing_dir):
return ProcessingOutput(
output_name=output_name,
source=os.path.join(processing_dir, "output"),
destination=output_path,
s3_upload_mode="EndOfJob"
)
def create_container_arguments(output_name, output_content_type):
output_config = {
output_name: {
"content_type": output_content_type
}
}
return [f"--output-config '{json.dumps(output_config)}'"]
```
## Start ProcessingJob
Now, the Processing Job is submitted using the Processor from the Sagemaker SDK.
Logs are turned off, but can be turned on for debugging purposes.
```
from sagemaker.processing import Processor
from sagemaker.network import NetworkConfig
processor = Processor(
role=iam_role,
image_uri=container_uri,
instance_count=instance_count,
instance_type=instance_type,
network_config=NetworkConfig(enable_network_isolation=enable_network_isolation),
sagemaker_session=sess
)
processor.run(
inputs=create_processing_inputs(processing_dir, flow, flow_uri),
outputs=[create_processing_output(output_name, output_path, processing_dir)],
arguments=create_container_arguments(output_name, output_content_type),
wait=True,
logs=False,
job_name=processing_job_name
)
```
## Kick off SageMaker Training Job (Optional)
Data Wrangler is a SageMaker tool for processing data to be used for Machine Learning. Now that
the data has been processed, users will want to train a model using the data. The following shows
an example of doing so using a popular algorithm XGBoost.
It is important to note that the following XGBoost objective ['binary', 'regression',
'multiclass'], hyperparameters, or content_type may not be suitable for the output data, and will
require changes to train a proper model. Furthermore, for CSV training, the algorithm assumes that
the target variable is in the first column. For more information on SageMaker XGBoost, please see
https://docs.aws.amazon.com/sagemaker/latest/dg/xgboost.html.
### Find Training Data path
The below demonstrates how to recursively search the output directory to find the data location.
```
s3_client = boto3.client("s3")
list_response = s3_client.list_objects_v2(Bucket=bucket, Prefix=output_prefix)
training_path = None
for content in list_response["Contents"]:
if "_SUCCESS" not in content["Key"]:
training_path = content["Key"]
print(training_path)
```
Next, the Training Job hyperparameters are set. For more information on XGBoost Hyperparameters,
see https://xgboost.readthedocs.io/en/latest/parameter.html.
```
region = boto3.Session().region_name
container = sagemaker.image_uris.retrieve("xgboost", region, "1.2-1")
hyperparameters = {
"max_depth":"5",
"objective": "reg:squarederror",
"num_round": "10",
}
train_content_type = (
"application/x-parquet" if output_content_type.upper() == "PARQUET"
else "text/csv"
)
train_input = sagemaker.inputs.TrainingInput(
s3_data=f"s3://{bucket}/{training_path}",
content_type=train_content_type,
)
```
The TrainingJob configurations are set using the SageMaker Python SDK Estimator, and which is fit
using the training data from the ProcessingJob that was run earlier.
```
estimator = sagemaker.estimator.Estimator(
container,
iam_role,
hyperparameters=hyperparameters,
instance_count=1,
instance_type="ml.m5.2xlarge",
)
estimator.fit({"train": train_input})
```
### Cleanup
Uncomment the following code cell to revert the SageMaker Python SDK to the original version used
before running this notebook. This notebook upgrades the SageMaker Python SDK to 2.x, which may
cause other example notebooks to break. To learn more about the changes introduced in the
SageMaker Python SDK 2.x update, see
[Use Version 2.x of the SageMaker Python SDK.](https://sagemaker.readthedocs.io/en/stable/v2.html).
```
# _ = subprocess.check_call(
# [sys.executable, "-m", "pip", "install", f"sagemaker=={original_version}"]
# )
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import csv
```
### Explore the DisGeNet dataset, find the disease of interest and get the list of human genes involved.
```
PATH = "./disgenet/"
filename = "./disgenet/curated_gene_disease_associations.tsv"
df = pd.read_csv(filename, sep = '\t')
df.head()
target = df.loc[df['diseaseName'] == 'Malignant mesothelioma']
target
```
### The file was already saved
```
t = pd.read_csv(PATH+"malignant_mesothelioma_curated_genes.tsv", sep = '\t')
t = t.drop('Unnamed: 0', axis = 1)
t.head()
curated = pd.read_csv("./disgenet/browser_source_genes_summary_CURATED.tsv", sep = '\t')
curated
#now let's save the gene symbols, entrez ID and names in arrays
geneSymbol=[]
geneID=[]
geneFullName = []
for i in range(len(curated)):
geneSymbol.append(curated['Gene'][i])
geneID.append(curated['Gene_id'][i])
geneFullName.append(curated['Gene_Full_Name'][i])
dict_Gene_GeneID = dict(zip(geneSymbol, geneID)) #NO
dict_Gene_GeneName = dict(zip(geneSymbol, geneFullName))
#we check on HGNC to see if we need to change genes name
#dict_Gene_GeneName
# printing original list
# using join()
# avoiding printing last comma
print("The formatted output is : ")
print(', '.join(geneSymbol))
# deleted quotes to pass the list into site
#https://www.genenames.org/tools/multi-symbol-checker/
```
### All symbols were approved on HGNC DB https://www.genenames.org/tools/multi-symbol-checker/
Now we need to retrieve the Uniprot AC map
```
uniprot=pd.read_csv('uniprot/uniprot-FINAL.tab',sep="\t")
uniprot
uniprotAC=[]
for i in range(len(uniprot)):
uniprotAC.append(uniprot['Entry'][i])
function=[]
for i in range(len(uniprot)):
function.append(uniprot['Function [CC]'][i])
geneName=[]
for i in range(len(uniprot)):
geneName.append(uniprot['Protein names'][i])
def returnFirstName(Namelist):
return Namelist.split()[0]
#costruisci la lista dei simboli e verifica la correttezza con la lista.
#Una volta accertata costruisci la lista uniprot scorrendo nuovamente la lista dei nomi in uniprot
geneUniprot = []
for i in range(len(uniprot)):
geneUniprot.append(returnFirstName(uniprot['Gene names'][i]))
uniprot['Gene names'][i] = returnFirstName(uniprot['Gene names'][i])
count = 0
geneMatched = []
for i in range(len(geneUniprot)):
if geneUniprot[i] in geneSymbol:
geneMatched.append(geneUniprot[i])
print(geneMatched)
geneNotMatched = []
for i in range(len(geneSymbol)):
if geneSymbol[i] not in geneMatched:
geneNotMatched.append(geneSymbol[i])
print(geneNotMatched)
#costruire l'indice
import math
#truncate description inside function when reaching .
def truncate_description(s):
s = s[s.find(":")+2:s.find(".")]
return s
new_function = []
for idx,name in enumerate(function):
if name == '' or (isinstance(name, float) and math.isnan(name)):
new_function.append("No description available for this gene")
else:
new_function.append(truncate_description(function[idx]))
#BUILD corrispondent geneID list and geneName list looking for keys in the dictionary
#usage : geneMatched
geneUniprotID = []
for key in dict_Gene_GeneID:
if key in geneMatched:
geneUniprotID.append(dict_Gene_GeneID[key])
geneUniprotName = []
for key in dict_Gene_GeneName:
if key in geneMatched:
geneUniprotName.append(dict_Gene_GeneName[key])
#geneUniprotID
results = pd.DataFrame(list(zip(geneUniprot,geneUniprotName,geneUniprotID,uniprotAC,new_function)), columns=['Symbol','Name','ID','UniprotAC', 'function'])
results.to_csv('mesothelioma-curated-genes.csv')
results.head()
results
```
### Exercise 1.2
For each seed gene, collect all binary protein interactions from two different PPI sources:
* Biogrid Human, latest release available
* IID Integrated Interactions Database (experimental data only, all tissues, unless stated otherwise in further instruction)
Note: once you got the list of the proteins interacting with at least one seed gene, you must
also retrieve and include in your interactome the interactions among these non-seed
proteins
```
#open biogrid DB
biogrid=pd.read_csv('BIOGRID-ALL-3.5.179.tab2.txt', sep='\t')
#select only human genes
biogrid_human=biogrid.loc[(biogrid['Organism Interactor A']==9606) & (biogrid['Organism Interactor B']==9606)]
biogrid_human
# look for the genes which interacts with at least one seed genes
biogrid_seed_genes = biogrid_human.loc[(biogrid_human['Official Symbol Interactor A'].isin(geneSymbol)) | (biogrid_human['Official Symbol Interactor B'].isin(geneSymbol))]
biogrid_seed_genes
```
### from our seed interaction let's search non-seeds genes that interacts with at least one seed gene
```
#Interactor A is not a seed gene but Interactor B is a seed gene
non_seed_df_A = biogrid_seed_genes.loc[~(biogrid_seed_genes['Official Symbol Interactor A'].isin(geneSymbol))]
seed_B = non_seed_df_A.loc[(non_seed_df_A ['Official Symbol Interactor B'].isin(geneSymbol))]
# interactor B is not a seed genes but Interactor A is a seed gene
non_seed_df_B = biogrid_seed_genes.loc[~(biogrid_seed_genes['Official Symbol Interactor B'].isin(geneSymbol))]
# Reset Index, otherwise they don't work with list
non_seed_df_A=non_seed_df_A.reset_index(drop=True)
non_seed_df_B=non_seed_df_B.reset_index(drop=True)
#build a list with only non seed genes that interacts with at least one seed gene
non_seed_list=[]
for i in range(0, len(non_seed_df_A)):
non_seed_list.append(non_seed_df_A['Official Symbol Interactor A'][i])
for i in range(0, len(non_seed_df_B)):
non_seed_list.append(non_seed_df_B['Official Symbol Interactor B'][i])
#drop duplicates
non_seed_list = list(dict.fromkeys(non_seed_list))
# Verify Correctness
for i in range(0, len(geneSymbol)):
for j in range(0, len(non_seed_list)):
if(geneSymbol[i] == non_seed_list[j]):
print("Something is wrong")
print("Ok")
print(len(non_seed_list))
#now search for non seed interactions from the human DB
#Return positions of non-seed genes that interacts with a non seed gene but both interacts with at least one seed gene
biogrid_human = biogrid_human.reset_index(drop=True)
#create a list of index of the original matrix with non seed genes interacting each others
list_of_idx=[]
#fill the list
for i in range(len(biogrid_human)):
if biogrid_human['Official Symbol Interactor A'][i] in non_seed_list and biogrid_human['Official Symbol Interactor B'][i] in non_seed_list:
list_of_idx.append(i)
#biogrid_seed_genes
biogrid_non_seed = biogrid_human.loc[list_of_idx]
biogrid_non_seed = biogrid_non_seed.reset_index(drop=True)
biogrid_non_seed
#make a table in which interactor 1 is seed and interactor 2 can be seed or non-seed
biogrid_seed_df = biogrid_seed_genes.loc[(biogrid_seed_genes['Official Symbol Interactor A'].isin(geneSymbol))]
interactome = pd.concat([biogrid_seed_df, biogrid_non_seed], ignore_index=True)
interactome.to_csv("interactome-biogrid.txt", sep='\t')
interactome = pd.read_csv("interactome-biogrid.txt", sep= '\t')
interactome = interactome.drop(['Unnamed: 0'], axis = 1)
interactome
iid = pd.read_csv('iid.txt', sep='\t')
iid = iid.drop(['Unnamed: 0'], axis = 1)
iid
#SAME PROCEDURE as above
iid_seed_genes = iid.loc[(iid['symbol1'].isin(geneSymbol)) | (iid['symbol2'].isin(geneSymbol))]
non_seed_1 = iid_seed_genes.loc[~(iid_seed_genes['symbol1'].isin(geneSymbol))]
# interactor 1 is not a seed genes and interacts with a seed gene
#B is a seed gene
non_seed_2 = iid_seed_genes.loc[~(iid_seed_genes['symbol2'].isin(geneSymbol))]
non_seed_1=non_seed_1.reset_index(drop=True)
non_seed_2=non_seed_2.reset_index(drop=True)
#build a list with only non seed genes that interacts with at least one seed gene
non_seed=[]
#non_seed_df_A = biogrid_seed_genes.loc[~(biogrid_seed_genes['Official Symbol Interactor A'].isin(geneSymbol))]
for i in range(0, len(non_seed_1)):
non_seed.append(non_seed_1['symbol1'][i])
for i in range(0, len(non_seed_2)):
non_seed.append(non_seed_2['symbol2'][i])
#drop duplicates
non_seed = list(dict.fromkeys(non_seed))
len(non_seed)
iid=iid.reset_index(drop=True)
#non_seed_df_A = biogrid_seed_genes.loc[~(biogrid_seed_genes['Official Symbol Interactor A'].isin(geneSymbol))]
list_of_idx=[]
#fill the list
for i in range(len(iid_seed_genes)):
if iid['symbol1'][i] in non_seed and iid['symbol2'][i] in non_seed:
list_of_idx.append(i)
iid_non_seed = iid.loc[list_of_idx]
iid_non_seed = iid_non_seed.reset_index(drop=True)
iid_seed_df = iid_seed_genes.loc[(iid_seed_genes['symbol1'].isin(geneSymbol))]
interactome2 = pd.concat([iid_seed_df, iid_non_seed], ignore_index=True)
interactome2 = pd.read_csv("interactome-iid.txt", sep='\t')
interactome2 = interactome2.drop(['Unnamed: 0'], axis = 1)
interactome2
```
### Summarize the main results in a table reporting:
* no. of seed genes found in each different DBs (some seed genes may be missing in the DBs);
* total no. of interacting proteins, including seed genes, for each DB;
* total no. of interactions found in each DB.
```
def find_unique_genes(dataframe, column_name):
genes_found = []
for index in range(len(dataframe)):
if dataframe[column_name][index] not in genes_found:
genes_found.append(dataframe[column_name][index])
return genes_found
# Genes from Biogrid
seed_B = non_seed_df_A.loc[(non_seed_df_A ['Official Symbol Interactor B'].isin(geneSymbol))]
seed_A = non_seed_df_B.loc[(non_seed_df_B ['Official Symbol Interactor A'].isin(geneSymbol))]
la = find_unique_genes(seed_B, 'Official Symbol Interactor B')
lb = find_unique_genes(seed_A, 'Official Symbol Interactor A')
l_tot = la + lb
#drop duplicates
total_genes = list(dict.fromkeys(l_tot))
print("seed genes in seed list:", len(geneSymbol), "seed genes found in BIOGRID: ",len(total_genes))
missing_gene = []
for index in range(len(geneSymbol)):
if geneSymbol[index] not in total_genes:
missing_gene.append(geneSymbol[index])
print("Genes missing in Biogrid\n", missing_gene)
# Genes from IID
seed1 = non_seed_2.loc[(non_seed_2['symbol1'].isin(geneSymbol))]
seed2 = non_seed_1.loc[(non_seed_1['symbol2'].isin(geneSymbol))]
# interactor 1 is not a seed genes and interacts with a seed gene
#B is a seed gene
l1 = find_unique_genes(seed1, 'symbol1')
l2 = find_unique_genes(seed2, 'symbol2')
l = l1+l2
#drop duplicates
i_genes = list(dict.fromkeys(l))
iid_missing = []
for index in range(len(geneSymbol)):
if geneSymbol[index] not in i_genes:
iid_missing.append(geneSymbol[index])
print("seed genes in seed list:", len(geneSymbol), "seed genes found in IID: ",len(i_genes))
print("Genes missing from IID DB \n", iid_missing)
```
total no. of interacting proteins, including seed genes, for each DB;
```
# total no. of interacting proteins, including seed genes, for BIOGRID;
interactors= []
for index in range(0, len(biogrid_human)):
if biogrid_human['Official Symbol Interactor A'][index] not in interactors:
interactors.append(biogrid_human['Official Symbol Interactor A'][index])
if biogrid_human['Official Symbol Interactor B'][index] not in interactors:
interactors.append(biogrid_human['Official Symbol Interactor B'][index])
len(interactors)
# total no. of interacting proteins, including seed genes, for IID;
def count_interactors(dataframe, column1, column2):
interactors= []
for index in range(len(dataframe)):
if dataframe[column1][index] not in interactors:
interactors.append(dataframe[column1][index])
if dataframe[column2][index] not in interactors:
interactors.append(dataframe[column2][index])
return len(interactors)
iid_len = count_interactors(iid, 'symbol1', 'symbol2')
print(iid_len)
len(iid)
```
total no. of interactions found in each DB.
```
# NUMBER OF INTERACTIONS FOR BIOGRID
countbio = biogrid_human['#BioGRID Interaction ID'].nunique()
print(countbio)
#NUMBER OF INTERACTIONS FOR IID
uniquesyms = []
for index in range(0, len(iid)):
sym1 = iid['symbol1'][index]
sym2 = iid['symbol2'][index]
uniquesyms.append(sym1+sym2)
#drop duplicates
uniquesyms = list(dict.fromkeys(uniquesyms))
print(len(uniquesyms))
```
### Build and store three tables:
* seed genes interactome: interactions that involve seed genes only, from all DBs, in the format:
interactor A gene symbol, interactor B gene symbol, interactor A Uniprot AC, interactor B
Uniprot AC, database source
* union interactome: all proteins interacting with at least one seed gene, from all DBs, same format as above.
* intersection interactome: all proteins interacting with at least one seed gene confirmed by both DBs, in the format: interactor A gene symbol, interactor B gene symbol, interactor A Uniprot AC, interactor B Uniprot AC
Always check that interactors are both human (i.e. organism ID is always 9606, Homo
Sapiens)
```
biogrid_human = biogrid_human.reset_index(drop=True)
uniprot_human = pd.read_csv("./uniprot/HUMAN_9606_idmapping.dat", sep = '\t')
uniprot_human
#save a list of all the symbols in order to search their uniprot
sym_to_fix=[]
sym_to_fix.extend(biogrid_human['Official Symbol Interactor A'])
sym_to_fix.extend(biogrid_human['Official Symbol Interactor B'])
#remove duplicates
sym_to_fix=list(set(sym_to_fix))
# using join()
# avoiding printing last comma
print("The formatted output is : ")
print(', '.join(sym_to_fix))
##print in order to search on uniprot.com
#upload the uniprot fixing file
unigene = pd.read_csv("./uniprot/geneid-uniprot.tab", sep = '\t')
unigene
#create a dictionary that maps symbol with its uniprot
unigene=unigene.rename(columns={"yourlist:M202001096746803381A1F0E0DB47453E0216320D5745D00": "symbol"})
unigene=pd.Series(unigene.Entry.values, index=unigene.symbol).to_dict()
biogrid_human['UniprotAC interactor A']= biogrid_human['Official Symbol Interactor A'].map(unigene)
biogrid_human['UniprotAC interactor B']= biogrid_human['Official Symbol Interactor B'].map(unigene)
biogrid_human.to_csv("biogrid_human_ext.tsv", sep = '\t')
biogrid_human.columns
iid_human = pd.read_csv('iid.txt', sep = '\t')
iid_human.drop(['Unnamed: 0'], axis = 1)
#SEED GENES INTERACTOME
def build_first_table(biogrid_human, iid_human):
db1 = 'Biogrid Human'
db2 = 'Integrated Interactions Database experimental data'
t = pd.DataFrame(columns=['interactorA', 'interactorB',
'interactorA_Uniprot_AC', 'interactorB_Uniprot_AC', 'db_source'])
for i in range(len(biogrid_human)):
sa = biogrid_human['Official Symbol Interactor A'][i]
sb = biogrid_human['Official Symbol Interactor B'][i]
uniprota = biogrid_human['UniprotAC interactor A'][i]
uniprotb = biogrid_human['UniprotAC interactor B'][i]
if sa in geneSymbol and sb in geneSymbol:
t = t.append({'interactorA':sa, 'interactorB':sb,
'interactorA_Uniprot_AC':uniprota, 'interactorB_Uniprot_AC':uniprotb, 'db_source': db1}
, ignore_index=True)
for i in range(len(iid_human)):
sa = iid_human['symbol1'][i]
sb = iid_human['symbol2'][i]
uniprota = iid_human['uniprot1'][i]
uniprotb = iid_human['uniprot2'][i]
if sa in geneSymbol and sb in geneSymbol:
t = t.append({'interactorA':sa, 'interactorB':sb,
'interactorA_Uniprot_AC':uniprota, 'interactorB_Uniprot_AC':uniprotb, 'db_source': db2}
, ignore_index=True)
t.to_csv("seed_genes_interactome.tsv", sep = '\t')
def build_union_interactome(biogrid_human, iid_human):
db1 = 'Biogrid Human'
db2 = 'Integrated Interactions Database experimental data'
t = pd.DataFrame(columns=['interactorA', 'interactorB',
'interactorA_Uniprot_AC', 'interactorB_Uniprot_AC', 'db_source'])
for i in range(len(biogrid_human)):
sa = biogrid_human['Official Symbol Interactor A'][i]
sb = biogrid_human['Official Symbol Interactor B'][i]
uniprota = biogrid_human['UniprotAC interactor A'][i]
uniprotb = biogrid_human['UniprotAC interactor B'][i]
if sa in geneSymbol or sb in geneSymbol:
t = t.append({'interactorA':sa, 'interactorB':sb,
'interactorA_Uniprot_AC':uniprota, 'interactorB_Uniprot_AC':uniprotb, 'db_source': db1}
, ignore_index=True)
for i in range(len(iid_human)):
sa = iid_human['symbol1'][i]
sb = iid_human['symbol2'][i]
uniprota = iid_human['uniprot1'][i]
uniprotb = iid_human['uniprot2'][i]
if sa in geneSymbol or sb in geneSymbol:
t = t.append({'interactorA':sa, 'interactorB':sb,
'interactorA_Uniprot_AC':uniprota, 'interactorB_Uniprot_AC':uniprotb, 'db_source': db2}
, ignore_index=True)
t.to_csv("union_interactome.tsv", sep = '\t')
# consider also non/seed-non/seed interactions
def build_nonseed_dataframes(biogrid_human, iid_human, union_human):
non_seed_df_A = union_human.loc[~(union_human['interactorA'].isin(geneSymbol))]
non_seed_df_B = union_human.loc[~(union_human['interactorB'].isin(geneSymbol))]
non_seed_df_A=non_seed_df_A.reset_index(drop=True)
non_seed_df_B=non_seed_df_B.reset_index(drop=True)
#build a list with only non seed genes that interacts with at least one seed gene
non_seed_list=[]
for i in range(0, len(non_seed_df_A)):
non_seed_list.append(non_seed_df_A['interactorA'][i])
for i in range(0, len(non_seed_df_B)):
non_seed_list.append(non_seed_df_B['interactorB'][i])
#drop duplicates
non_seed_list = list(dict.fromkeys(non_seed_list))
list_of_idx=[]
#fill the list
for i in range(len(biogrid_human)):
if biogrid_human['Official Symbol Interactor A'][i] in non_seed_list and biogrid_human['Official Symbol Interactor B'][i] in non_seed_list:
list_of_idx.append(i)
biogrid_non_seed = biogrid_human.loc[list_of_idx]
biogrid_non_seed = biogrid_non_seed.reset_index(drop=True)
biogrid_non_seed.to_csv("biogrid_union_non_seed.tsv", sep = '\t')
#same procedure for iid
list_of_idx=[]
#fill the list
for i in range(len(iid_human)):
if iid_human['symbol1'][i] in non_seed_list and iid_human['symbol2'][i] in non_seed_list:
list_of_idx.append(i)
iid_non_seed = iid_human.loc[list_of_idx]
iid_non_seed = iid_non_seed.reset_index(drop=True)
iid_non_seed.to_csv("iid_union_non_seed.tsv", sep = '\t')
def merge_non_seed_dataframes(biogrid_human, iid_human, union_human):
db1 = 'Biogrid Human'
db2 = 'Integrated Interactions Database experimental data'
union_dict = {}
for i in range(len(iid_human)):
sa = iid_human['symbol1'][i]
sb = iid_human['symbol2'][i]
uniprota = iid_human['uniprot1'][i]
uniprotb = iid_human['uniprot2'][i]
'''union_human = union_human.append({'interactorA':sa, 'interactorB':sb,
'interactorA_Uniprot_AC':uniprota, 'interactorB_Uniprot_AC':uniprotb, 'db_source': db2}
, ignore_index=True)'''
union_dict[i] = {"interactorA": sa, 'interactorB':sb,
'interactorA_Uniprot_AC':uniprota, 'interactorB_Uniprot_AC':uniprotb, 'db_source': db2}
df = pd.DataFrame.from_dict(union_dict, "index")
new = pd.concat([union_human, df], ignore_index=True)
print("done with iid", len(union_human))
union_dict = {}
for i in range(len(biogrid_human)):
sa = biogrid_human['Official Symbol Interactor A'][i]
sb = biogrid_human['Official Symbol Interactor B'][i]
uniprota = biogrid_human['UniprotAC interactor A'][i]
uniprotb = biogrid_human['UniprotAC interactor B'][i]
'''union_human = union_human.append({'interactorA':sa, 'interactorB':sb,
'interactorA_Uniprot_AC':uniprota, 'interactorB_Uniprot_AC':uniprotb, 'db_source': db1}
, ignore_index=True)'''
union_dict[i] = {"interactorA": sa, 'interactorB':sb,
'interactorA_Uniprot_AC':uniprota, 'interactorB_Uniprot_AC':uniprotb, 'db_source': db1}
df = pd.DataFrame.from_dict(union_dict, "index")
new = pd.concat([new, df], ignore_index=True)
# important to set the 'orient' parameter to "index" to make the keys as rows
# drop duplicates for interactorA and interactorB
new = new.drop_duplicates(subset = ['interactorA', 'interactorB'], keep='first')
new.to_csv("union_interactome_extended.tsv", sep = '\t')
biogrid_human=biogrid_human.reset_index(drop=True)
iid_human=iid_human.reset_index(drop=True)
build_union_interactome(biogrid_human, iid_human)
build_first_table(biogrid_human, iid_human)
interactome_seed = pd.read_csv("seed_genes_interactome.tsv", sep = '\t')
interactome_seed.drop(['Unnamed: 0'], axis = 1)
union_human = pd.read_csv("union_interactome.tsv", sep = '\t')
biogrid_human=biogrid_human.reset_index(drop=True)
iid_human=iid_human.reset_index(drop=True)
union_human=union_human.reset_index(drop=True)
build_nonseed_dataframes(biogrid_human, iid_human, union_human)
nsbio = pd.read_csv("biogrid_union_non_seed.tsv", sep = '\t')
nsiid = pd.read_csv("iid_union_non_seed.tsv", sep = '\t')
nsiid = nsiid.reset_index(drop=True)
nsbio = nsbio.reset_index(drop=True)
merge_non_seed_dataframes(nsbio, nsiid, union_human)
union_ext = pd.read_csv("union_interactome_extended.tsv", sep = '\t')
#intersection interactome: all proteins interacting with at least one seed gene confirmed by both DBs
def build_intersection_interactome(biogrid_human, iid_human):
db1 = 'Biogrid Human'
db2 = 'Integrated Interactions Database experimental data'
union = pd.read_csv("union_interactome.tsv", sep = '\t')
union_biogrid = union.loc[(union['db_source'] == 'Biogrid Human')]
union_biogrid = union_biogrid.drop(['Unnamed: 0', 'db_source'], axis = 1)
union_iid = union.loc[(union['db_source'] == 'Integrated Interactions Database experimental data')]
union_iid = union_iid.drop(['Unnamed: 0', 'db_source'], axis = 1)
intersect.dropna(inplace=True)
intersect = pd.merge(union_biogrid, union_iid)
intersect.to_csv("intersection_interactome.tsv", sep = '\t')
biogrid_human=biogrid_human.reset_index(drop=True)
iid_human=iid_human.reset_index(drop=True)
build_intersection_interactome(biogrid_human, iid_human)
intersection_interactome = pd.read_csv("intersection_interactome.tsv", sep = '\t')
intersection_interactome
approvedsym = pd.read_csv("approved-symbols.csv", sep = ',')
approvedsym
unionint = pd.read_csv("union_interactome_extended.tsv", sep = '\t')
unionint = unionint.drop(['Unnamed: 0'], axis = 1)
sym_to_fix=[]
sym_to_fix.extend(unionint['interactorA'])
sym_to_fix.extend(unionint['interactorB'])
#remove duplicates
sym_to_fix=list(set(sym_to_fix))
# using join()
# avoiding printing last comma
#print(', '.join(sym_to_fix))
# deleted quotes to pass the list into site
#https://www.genenames.org/tools/multi-symbol-checker/
def substring_after(s, delim):
return s.split(delim)
new = []
for s in sym_to_fix:
if(" " not in s):
c = substring_after(s, ";")
for character in c:
new.append(character)
new
sym_to_fix=list(set(new))
# using join()
# avoiding printing last comma
print(', '.join(sym_to_fix))
a = pd.read_csv("./hgnc/unionintext_symbols.csv")
for sym in a['Input']:
print(sym)
```
# Enrichr Analysis
```
kegg_union = pd.read_table("enrichr/kegg_human/union/KEGG_2019_Human_table.txt")
kegg_union = kegg_union[:10]
kegg_union.to_csv("enrichr/kegg_human/union/KEGG_2019_Human_table.txt")
go_bp_union = pd.read_table("enrichr/ontologies/union/GO_Biological_Process_2018_table.txt")
go_bp_union = go_bp_union[:10]
go_bp_union.to_csv("enrichr/ontologies/union/GO_Biological_Process_2018_table.txt")
go_mf_union = pd.read_table("enrichr/ontologies/union/GO_Molecular_Function_2018_table.txt")
go_mf_union = go_mf_union[:10]
go_mf_union.to_csv("enrichr/ontologies/union/GO_Molecular_Function_2018_table.txt")
go_cc_union = pd.read_csv("enrichr/ontologies/union/GO_Cellular_Component_2018_table.csv")
go_cc_union = go_cc_union[:10]
go_cc_union.to_csv("enrichr/ontologies/union/GO_Cellular_Component_2018_table.csv")
go_cc_union
kegg_union
listgene=list(set(geneSymbol))
for gene in listgene:
print(gene)
kegg_seed = pd.read_table("enrichr/kegg_human/seed/KEGG_2019_Human_table.txt")
kegg_seed = kegg_seed[:10]
kegg_seed.to_csv("enrichr/kegg_human/seed/KEGG_2019_Human_table.txt")
go_bp_seed = pd.read_table("enrichr/ontologies/seed/GO_Biological_Process_2018_table.txt")
go_bp_seed = go_bp_seed[:10]
go_bp_seed.to_csv("enrichr/ontologies/seed/GO_Biological_Process_2018_table.txt")
go_mf_seed = pd.read_table("enrichr/ontologies/seed/GO_Molecular_Function_2018_table.txt")
go_mf_seed = go_mf_seed[:10]
go_mf_seed.to_csv("enrichr/ontologies/seed/GO_Molecular_Function_2018_table.txt")
go_mf_seed
```
| github_jupyter |
<table class="ee-notebook-buttons" align="left">
<td><a target="_blank" href="https://github.com/giswqs/earthengine-py-notebooks/tree/master/Image/conditional_operations.ipynb"><img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" /> View source on GitHub</a></td>
<td><a target="_blank" href="https://nbviewer.jupyter.org/github/giswqs/earthengine-py-notebooks/blob/master/Image/conditional_operations.ipynb"><img width=26px src="https://upload.wikimedia.org/wikipedia/commons/thumb/3/38/Jupyter_logo.svg/883px-Jupyter_logo.svg.png" />Notebook Viewer</a></td>
<td><a target="_blank" href="https://mybinder.org/v2/gh/giswqs/earthengine-py-notebooks/master?filepath=Image/conditional_operations.ipynb"><img width=58px src="https://mybinder.org/static/images/logo_social.png" />Run in binder</a></td>
<td><a target="_blank" href="https://colab.research.google.com/github/giswqs/earthengine-py-notebooks/blob/master/Image/conditional_operations.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" /> Run in Google Colab</a></td>
</table>
## Install Earth Engine API and geemap
Install the [Earth Engine Python API](https://developers.google.com/earth-engine/python_install) and [geemap](https://github.com/giswqs/geemap). The **geemap** Python package is built upon the [ipyleaflet](https://github.com/jupyter-widgets/ipyleaflet) and [folium](https://github.com/python-visualization/folium) packages and implements several methods for interacting with Earth Engine data layers, such as `Map.addLayer()`, `Map.setCenter()`, and `Map.centerObject()`.
The following script checks if the geemap package has been installed. If not, it will install geemap, which automatically installs its [dependencies](https://github.com/giswqs/geemap#dependencies), including earthengine-api, folium, and ipyleaflet.
**Important note**: A key difference between folium and ipyleaflet is that ipyleaflet is built upon ipywidgets and allows bidirectional communication between the front-end and the backend enabling the use of the map to capture user input, while folium is meant for displaying static data only ([source](https://blog.jupyter.org/interactive-gis-in-jupyter-with-ipyleaflet-52f9657fa7a)). Note that [Google Colab](https://colab.research.google.com/) currently does not support ipyleaflet ([source](https://github.com/googlecolab/colabtools/issues/60#issuecomment-596225619)). Therefore, if you are using geemap with Google Colab, you should use [`import geemap.eefolium`](https://github.com/giswqs/geemap/blob/master/geemap/eefolium.py). If you are using geemap with [binder](https://mybinder.org/) or a local Jupyter notebook server, you can use [`import geemap`](https://github.com/giswqs/geemap/blob/master/geemap/geemap.py), which provides more functionalities for capturing user input (e.g., mouse-clicking and moving).
```
# Installs geemap package
import subprocess
try:
import geemap
except ImportError:
print('geemap package not installed. Installing ...')
subprocess.check_call(["python", '-m', 'pip', 'install', 'geemap'])
# Checks whether this notebook is running on Google Colab
try:
import google.colab
import geemap.eefolium as emap
except:
import geemap as emap
# Authenticates and initializes Earth Engine
import ee
try:
ee.Initialize()
except Exception as e:
ee.Authenticate()
ee.Initialize()
```
## Create an interactive map
The default basemap is `Google Satellite`. [Additional basemaps](https://github.com/giswqs/geemap/blob/master/geemap/geemap.py#L13) can be added using the `Map.add_basemap()` function.
```
Map = emap.Map(center=[40,-100], zoom=4)
Map.add_basemap('ROADMAP') # Add Google Map
Map
```
## Add Earth Engine Python script
```
# Add Earth Engine dataset
# Load a Landsat 8 image.
image = ee.Image('LANDSAT/LC08/C01/T1_TOA/LC08_044034_20140318')
# Create NDVI and NDWI spectral indices.
ndvi = image.normalizedDifference(['B5', 'B4'])
ndwi = image.normalizedDifference(['B3', 'B5'])
# Create a binary layer using logical operations.
bare = ndvi.lt(0.2).And(ndwi.lt(0))
# Mask and display the binary layer.
Map.setCenter(-122.3578, 37.7726, 12)
Map.setOptions('satellite')
Map.addLayer(bare.updateMask(bare), {}, 'bare')
```
## Display Earth Engine data layers
```
Map.addLayerControl() # This line is not needed for ipyleaflet-based Map.
Map
```
| github_jupyter |
# CHAPTER 3
# Section 1
## Playing with PyTorch tensors
```
import torch
data = torch.ones(3)
data
data[0], data[1]
float(data[0]), float(data[1])
data[1] = 4 # mutable
data
a = torch.tensor([4, 5, 6, 7])
a
a.dtype
float(a[2])
p = torch.tensor([[4, 2], [4, 5.6], [1.3, 13]], dtype=torch.float64)
p
p.shape
p[0]
p[0, 0], p[0, 1]
p[:, 0] # all rows, first column
p[None].shape
# example image
img = torch.rand(3, 28, 28) * 255 # channels x rows x columns
img
w = torch.tensor([0.2126, 0.7152, 0.0722])
batch = torch.rand(2, 3, 28, 28) * 255 # 2 is the number of examples
batch.shape
img.mean(-3).shape, batch.mean(-3).shape
w_un = w.unsqueeze(-1).unsqueeze(-1)
w_un.shape
img_weights = w_un * img
batch_weights = w_un * batch
img_weights.shape, batch_weights.shape
%matplotlib inline
import matplotlib
import numpy as np
import matplotlib.pyplot as plt
img_gray = img_weights.sum(-3)
batch_gray = batch_weights.sum(-3)
img_gray.shape, batch_gray.shape
# trying to plot the images
from pylab import rcParams
rcParams['figure.figsize'] = 50, 50
img_n = img.resize(28, 28, 3)
batch_n = batch.resize(28, 28, 3, 2)
plt.subplot(2, 3, 1)
plt.title("Image color", fontsize=30)
fig = plt.imshow(img_n)
plt.subplot(2, 3, 2)
plt.title("Batch 1 color", fontsize=30)
plt.imshow(batch_n[..., 0])
plt.subplot(2, 3, 3)
plt.title("Batch 2 color", fontsize=30)
plt.imshow(batch_n[..., 1])
plt.subplot(2, 3, 4)
plt.title("Image grayscale", fontsize=30)
fig = plt.imshow(img_gray)
batch_n = batch_gray.resize(28, 28, 2)
plt.subplot(2, 3, 5)
plt.title("Batch 1 grayscale", fontsize=30)
plt.imshow(batch_n[..., 0])
plt.subplot(2, 3, 6)
plt.title("Batch 2 grayscale", fontsize=30)
plt.imshow(batch_n[..., 1])
plt.show()
# different dtypes in pytorch
torch.int8, torch.int16, torch.int32, torch.int64
torch.float, torch.float16, torch.float32, torch.float64
torch.bool
# TO CHANGE THE DTYPE TWO METHODS ARE THERE
print(p.dtype)
print(p.int().dtype)
print(p.to(torch.int64).dtype)
# trying to understand the underlying storage working of tensors
print(p)
p.storage() # storage is where the data is stored in a 1D array (always)
# regardless of the tensor's dimension
st = p.storage()
st[3] = 13123123123
# this will change the actual tensor too
print(st)
p
# methods with a trailing _ changes the tensor being operated on
# instead of creating a new copy with the change
p.zero_()
p
p = torch.rand(2, 4).to(dtype=torch.float32)
p
p.t() # shorthand transpose function
p.is_contiguous()
# tensor to numpy
n = p.numpy()
n
# numpy to tensor
f = torch.from_numpy(n)
f
# saving tensors
torch.save(p, '../Chapter_3/p.t')
# reading tensors from file
p_n = torch.load('../Chapter_3/p.t')
p_n
```
| github_jupyter |
# Learning Objectives
By the end of this class, you will be able to...
- Compute probability density functions and cumulative density functions
- Use the `scipy.stats` package to compute the Survival Value or CDF Value for a known distribution
```
import numpy as np
import pandas as pd
df = pd.read_csv('../Pandas/titanic.csv')
```
## Probability Distribution Function (PDF)
First let's review the difference between discrete and continuous random variables:
- **Discrete:** takes on a finite or countable number of values.
- **Continuous:** takes on an infinite number of values
Because continuous random variables can take on an infinite number of values, we can't say with certantiy what value the variable will be at any point, so we have to instead provide an **interval, or range** of values that the variable could be.
### An Example
For example, what's the probability that New York City get's 4 inches of snow on December 17th? 3.99999 and 4.0001 inches don't count, it has to be _exactly_ 4. It would be impossible to say with exact certainty given there are infinite decimals! However, if instead of looking for exactly 4, we looked at the range of values between 3.9 and 4.1, we could compute the probability for that range! This is where PDFs come in to help us calculate this!
### Graphing PDFs
When we graph a PDF, it has a similar pattern to a histogram. The main difference is that instead of the y-axis corresponding to values, it shows the percent probability of the x-axis values. This is known as _normalizing the values_.
## Activity: Reminder of how to Plot Histograms
To start, let's plot the histogram for the Age of people on the Titanic. To do this, we'll use the [seaborn](https://seaborn.pydata.org/) library
## Activity (Remind Histogram): Plot the Histogram of Age for Titanic Dataset
```
import seaborn as sns
# create a list of Age values not including N/A values
ls_age = df['Age'].dropna()
# Now plot the data in this list into a histogram!
sns.distplot(ls_age, hist=True, kde=False, bins=16)
```
- Lets now plot the PDF of the same data
```
import seaborn as sns
# Notice only the KDE parameter is different!
# What does kde stand for? https://seaborn.pydata.org/generated/seaborn.distplot.html
sns.distplot(df['Age'].dropna(), hist=True, kde=True, bins=16)
```
## Activity: Proving our PDF
We know that the y-axis shows the percent probability of the x-axis values. But where do the y-axis numbers come from? For example, for Ages 20-25, why is the y-value around 0.030?
Let's prove it!
```
# custom histogram function,
# Same as calling sns.distplot(ls_age, hist=True, kde=False, bins=16),
# but instead of plotting, returns a dictionary with a RANGE as the key (20.315, 25.28875),
# and TOTAL NUMBER OF VALUES IN THAT RANGE (122) as values
def custom_hist(ls, interval):
hist_ls_dict = dict()
# min value in the range of values
min_ls = np.min(ls)
# max value in the range of values
max_ls = np.max(ls)
print(max_ls)
# distance between each bin
I = ((max_ls - min_ls) / interval)
print(I)
# Create the dictionary
for j in range(interval):
# print((min_ls + j*I, min_ls + (j+1) *I))
# print(np.sum(((min_ls + j*I) <=ls) & (ls <= (min_ls + (j+1) *I))))
hist_ls_dict[(min_ls + j*I, min_ls + (j+1) *I)]= np.sum(((min_ls + j*I) <=ls) & (ls <= (min_ls + (j+1) *I)))
return hist_ls_dict
print(custom_hist(df['Age'].dropna().values, 16))
```
Why is our `I` (the distance between each bin) not exactly 5? 80/16 = 5
- Because our range doesn't start at 0! Our range starts at 0.42. `(80-0.42)/16 = 4.97375`
```
# Remember the two print statements in custom_hist!
hist_dict = custom_hist(df['Age'].dropna().values, 16)
# number of values in the histogram
sum(hist_dict.values())
```
We now know the following information:
- The number of people whose ages fall between 20-25: 122
- The total number of values in our dataset: 714
- The distance between each bin: 4.97375
```
122/714/4.97375
```
## Activity: What percent of passengers are younger than 40?
```
How_many_younger_40 = df[df['Age'] <= 40]
pr_below_40 = len(How_many_younger_40)/len(df['Age'].dropna())
print(pr_below_40)
```
Only 3 lines of code! Now imagine you **only** had the PDF graph, and not access to the data.
We'd have to calcualte the area of the graph in the range (0.42, 40), which is a very large area, and would be difficult to compute.
## It is not easy to calculate this percentage from PDF as we should compute the area
## Cumulative Density Function (CDF)
- In the above example, we could not easily obtain the percentage from a PDF, although it is possible.
- This is much easier if we use a **CDF**. A CDF calculates the probability that a random variable is less than a threshold value
- Let's learn CDF by example: given an array of numbers (our random variable) and a threshold value as input:
1. Find the minimum value in the array
1. Set the threshold to be the minimum value of the array
1. For a given array of numbers and a given threshold, count all of the elements in the array that are less than the threshold, and divide that count by the length of the array
1. Repeat step three, increasing the threshold by one, until you go through step three where threshold is equal to the maximum value in the array
```
ls_age = df['Age'].dropna().values
def calculate_cdf(x, threshold):
return np.sum(x <= threshold)
# Create an array cdf_age where each value is the cdf of the age for each threshold
cdf_age = []
#Find the minimum value in the array
min_val = np.min(ls_age)
# Set the threshold to be the minimum value of the array
threshold = np.min(ls_age)
# For a given array of numbers and a given threshold,
for r in range(int(np.min(ls_age)), int(np.max(ls_age))):
# count all of the elements in the array that are less than the threshold,
less_than_threshold = calculate_cdf(ls_age, r)
# and divide that count by the length of the array
age_value = less_than_threshold/len(ls_age)
cdf_age.append(age_value)
# Repeat, increasing the threshold by one,
# until the threshold is equal to the maximum value in the array
import matplotlib.pyplot as plt
plt.plot(range(int(np.min(ls_age)), int(np.max(ls_age))), cdf_age)
plt.grid()
```
## Use Seaborn or Matplotlib to plot CDF of Age
```
sns.distplot(df['Age'].dropna(), hist_kws=dict(cumulative=True), kde_kws=dict(cumulative=True))
df['Age'].dropna().hist(cumulative=True, density=True)
```
## More about PDF
Sometimes we'll have more than one data sets we want to graph. For example, if we want to show the age of both male and female passengers.
PDFs can be displayed an alternative way when we have 2 or more lists, as seen in the example below:
```
sns.violinplot(x="Sex", y="Age", data=df)
```
If you look at it sideways (tilt your head), it looks similar to the first PDF we saw!
## Normal Distribution
When we plot histograms or PDFs from an array, we may occasionally get graphs that have a _perfect_ bell shape curve. Histograms that have this shape have **normal distributions!**
These graphs are helpful because their shape allows us to easily compute the area!
An example of a PDF with a normal distibution is given below:
```
import numpy as np
import seaborn as sns
# Generate 1000 samples with 60 as its mean and 10 as its std
a = np.random.normal(60, 10, 1000)
sns.distplot(a, hist=True, kde=True, bins=20)
```
**Note that the _entire area is always equal to 1 for any PDF._**
## Activity:
The DS 1.1 instructor just finished grading the final exam. He is reporting that the mean was 60 (with the possible score range from 0 to 100) with standard deviation of 10.
What is the probability that students got more than 70? Assume that the instructor plotted the PDF of the data and it was _normal_
**Hint:** If we can obtain area of the graph for everything less than 70, that's the same as getting the CDF at 70! How would this be useful?
**Hint:** look into this [library from the scipy.stats documentation](https://docs.scipy.org/doc/scipy/reference/generated/scipy.stats.norm.html#scipy.stats.norm). It may be helpful :)
```
from scipy.stats import norm
1 - norm.cdf(70, loc=60, scale=10)
# or
print(norm.sf(70, loc=60, scale=10))
```
This area we just calculated is called the _survival_
## Normal Distribution Properties:
When the data is Normally distributed:
- 68% of the data is captured within one standard deviation from the mean.
- 95% of the data is captured within two standard deviations from the mean.
- 99.7% of the data is captured within three standard deviations from the mean.
<br><img src="http://www.oswego.edu/~srp/stats/images/normal_34.gif" /><br>
## Activity:
Let's prove that these normal distribution properties are true. Show that about 68% of the values in our previous DS final exam example are in the [50, 70] range
```
norm.cdf(70, loc=60, scale=10) - norm.cdf(50, loc=60, scale=10)
```
## Scaling the Normal Ditribution
We can take any list of values which are normally distribute and apply simple arithmatic on the list such that the mean is 0 and the standard deviation is 1 for our new transformed list. We can do this through following the steps below:
For each of the values in the list, we will need to _subtract the mean of list from the value, and divide this difference by the standard deviation_
Let's walk through this using the DS final exam data and normally distributing it:
```
# Our list of exam scores with mean = 60, and std = 10
exam_scores = np.random.normal(60, 10, 1000)
normalized_scores = []
for score in exam_scores:
# subtract the mean of list from the value,
# and divide this difference by the standard deviation
normal_score = (score - 60)/10
normalized_scores.append(normal_score)
sns.distplot(normalized_scores, hist=True, kde=True, bins=20)
np.mean(normalized_scores)
np.std(normalized_scores)
```
## Z-Distribution
**Z-distribution is another name for standard Normal distribution!** This is what we just calculated in the last example!
Following the process as outlined above will allow you to calculate the z-distribution of a data set!
| github_jupyter |
# Qiskit Visualizations
```
from qiskit import *
from qiskit.visualization import plot_histogram
from qiskit.tools.monitor import job_monitor
```
## Plot histogram <a name='histogram'></a>
To visualize the data from a quantum circuit run on a real device or `qasm_simulator` we have made a simple function
`plot_histogram(data)`
As an example we make a 2-qubit Bell state
```
# quantum circuit to make a Bell state
bell = QuantumCircuit(2, 2)
bell.h(0)
bell.cx(0, 1)
meas = QuantumCircuit(2, 2)
meas.measure([0,1], [0,1])
# execute the quantum circuit
backend = BasicAer.get_backend('qasm_simulator') # the device to run on
circ = bell + meas
result = execute(circ, backend, shots=1000).result()
counts = result.get_counts(circ)
print(counts)
plot_histogram(counts)
```
### Options when plotting a histogram
The `plot_histogram()` has a few options to adjust the output graph. The first option is the `legend` kwarg. This is used to provide a label for the executions. It takes a list of strings use to label each execution's results. This is mostly useful when plotting multiple execution results in the same histogram. The `sort` kwarg is used to adjust the order the bars in the histogram are rendered. It can be set to either ascending order with `asc` or descending order with `dsc`. The `number_to_keep` kwarg takes an integer for the number of terms to show, the rest are grouped together in a single bar called rest. You can adjust the color of the bars with the `color` kwarg which either takes a string or a list of strings for the colors to use for the bars for each execution. You can adjust whether labels are printed above the bars or not with the `bar_labels` kwarg. The last option available is the `figsize` kwarg which takes a tuple of the size in inches to make the output figure.
```
# Execute 2-qubit Bell state again
second_result = execute(circ, backend, shots=1000).result()
second_counts = second_result.get_counts(circ)
# Plot results with legend
legend = ['First execution', 'Second execution']
plot_histogram([counts, second_counts], legend=legend)
plot_histogram([counts, second_counts], legend=legend, sort='desc', figsize=(15,12),
color=['orange', 'black'], bar_labels=False)
```
### Using the output from plot_histogram()
When using the plot_histogram() function it returns a `matplotlib.Figure` for the rendered visualization. Jupyter notebooks understand this return type and render it for us in this tutorial, but when running outside of Jupyter you do not have this feature automatically. However, the `matplotlib.Figure` class natively has methods to both display and save the visualization. You can call `.show()` on the returned object from `plot_histogram()` to open the image in a new window (assuming your configured matplotlib backend is interactive). Or alternatively you can call `.savefig('out.png')` to save the figure to `out.png`. The `savefig()` method takes a path so you can adjust the location and filename where you're saving the output.
## Plot State <a name='state'></a>
In many situations you want to see the state of a quantum computer. This could be for debugging. Here we assume you have this state (either from simulation or state tomography) and the goal is to visualize the quantum state. This requires exponential resources, so we advise to only view the state of small quantum systems. There are several functions for generating different types of visualization of a quantum state
```
plot_state_city(quantum_state)
plot_state_paulivec(quantum_state)
plot_state_qsphere(quantum_state)
plot_state_hinton(quantum_state)
plot_bloch_multivector(quantum_state)
```
A quantum state is either a state matrix $\rho$ (Hermitian matrix) or statevector $|\psi\rangle$ (complex vector). The state matrix is related to the statevector by
$$\rho = |\psi\rangle\langle \psi|,$$
and is more general as it can represent mixed states (positive sum of statevectors)
$$\rho = \sum_k p_k |\psi_k\rangle\langle \psi_k |.$$
The visualizations generated by the functions are:
- `'plot_state_city'`: The standard view for quantum states where the real and imaginary (imag) parts of the state matrix are plotted like a city.
- `'plot_state_qsphere'`: The Qiskit unique view of a quantum state where the amplitude and phase of the state vector are plotted in a spherical ball. The amplitude is the thickness of the arrow and the phase is the color. For mixed states it will show different `'qsphere'` for each component.
- `'plot_state_paulivec'`: The representation of the state matrix using Pauli operators as the basis $\rho=\sum_{q=0}^{d^2-1}p_jP_j/d$.
- `'plot_state_hinton'`: Same as `'city'` but where the size of the element represents the value of the matrix element.
- `'plot_bloch_multivector'`: The projection of the quantum state onto the single qubit space and plotting on a bloch sphere.
```
from qiskit.visualization import plot_state_city, plot_bloch_multivector
from qiskit.visualization import plot_state_paulivec, plot_state_hinton
from qiskit.visualization import plot_state_qsphere
# execute the quantum circuit
backend = BasicAer.get_backend('statevector_simulator') # the device to run on
result = execute(bell, backend).result()
psi = result.get_statevector(bell)
plot_state_city(psi)
plot_state_hinton(psi)
plot_state_qsphere(psi)
plot_state_paulivec(psi)
plot_bloch_multivector(psi)
```
Here we see that there is no information about the quantum state in the single qubit space as all vectors are zero.
### Options when using state plotting functions
The various functions for plotting quantum states provide a number of options to adjust how the plots are rendered. Which options are available depends on the function being used.
**plot_state_city()** options
- **title** (str): a string that represents the plot title
- **figsize** (tuple): figure size in inches (width, height).
- **color** (list): a list of len=2 giving colors for real and imaginary components of matrix elements.
```
plot_state_city(psi, title="My City", color=['black', 'orange'])
```
**plot_state_hinton()** options
- **title** (str): a string that represents the plot title
- **figsize** (tuple): figure size in inches (width, height).
```
plot_state_hinton(psi, title="My Hinton")
```
**plot_state_paulivec()** options
- **title** (str): a string that represents the plot title
- **figsize** (tuple): figure size in inches (width, height).
- **color** (list or str): color of the expectation value bars.
```
plot_state_paulivec(psi, title="My Paulivec", color=['purple', 'orange', 'green'])
```
**plot_state_qsphere()** options
- **figsize** (tuple): figure size in inches (width, height).
**plot_bloch_multivector()** options
- **title** (str): a string that represents the plot title
- **figsize** (tuple): figure size in inches (width, height).
```
plot_bloch_multivector(psi, title="My Bloch Spheres")
```
### Using the output from state plotting functions
When using any of the state plotting functions it returns a `matplotlib.Figure` for the rendered visualization. Jupyter notebooks understand this return type and render it for us in this tutorial, but when running outside of Jupyter you do not have this feature automatically. However, the `matplotlib.Figure` class natively has methods to both display and save the visualization. You can call `.show()` on the returned object to open the image in a new window (assuming your configured matplotlib backend is interactive). Or alternatively you can call `.savefig('out.png')` to save the figure to `out.png` in the current working directory. The `savefig()` method takes a path so you can adjust the location and filename where you're saving the output.
## Interactive State Plots for Jupyter Notebooks <a name='interstate'></a>
Just like with `plot_histogram()` there is a second set of functions for each of the functions to plot the quantum state. These functions have the same name but with a prepended `i`:
```
iplot_state_city(quantum_state)
iplot_state_paulivec(quantum_state)
iplot_state_qsphere(quantum_state)
iplot_state_hinton(quantum_state)
iplot_bloch_multivector(quantum_state)
```
These functions are made using an externally hosted JS library for use in Jupyter notebooks. The interactive plot can only be used if you're running inside a Jupyter notebook and only if you have external connectivity to the host with the JS library. If you use these functions outside of a Jupyter notebook it will fail.
```
from qiskit.tools.visualization import iplot_state_paulivec
# Generate an interactive pauli vector plot
iplot_state_paulivec(psi)
```
## Plot Bloch Vector <a name='bloch'></a>
A standard way of plotting a quantum system is using the Bloch vector. This only works for a single qubit and takes as input the Bloch vector.
The Bloch vector is defined as $[x = \mathrm{Tr}[X \rho], y = \mathrm{Tr}[Y \rho], z = \mathrm{Tr}[Z \rho]]$, where $X$, $Y$, and $Z$ are the Pauli operators for a single qubit and $\rho$ is the state matrix.
```
from qiskit.visualization import plot_bloch_vector
plot_bloch_vector([0,1,0])
```
### Options for plot_bloch_vector()
- **title** (str): a string that represents the plot title
- **figsize** (tuple): Figure size in inches (width, height).
```
plot_bloch_vector([0,1,0], title='My Bloch Sphere')
```
### Adjusting the output from plot_bloch_vector()
When using the `plot_bloch_vector` function it returns a `matplotlib.Figure` for the rendered visualization. Jupyter notebooks understand this return type and render it for us in this tutorial, but when running outside of Jupyter you do not have this feature automatically. However, the `matplotlib.Figure` class natively has methods to both display and save the visualization. You can call `.show()` on the returned object to open the image in a new window (assuming your configured matplotlib backend is interactive). Or alternatively you can call `.savefig('out.png')` to save the figure to `out.png` in the current working directory. The `savefig()` method takes a path so you can adjust the location and filename where you're saving the output.
```
import qiskit.tools.jupyter
%qiskit_version_table
%qiskit_copyright
```
| github_jupyter |
Problem Statement Predicting Coupon Redemption XYZ Credit Card company regularly helps it’s merchants understand their data better and take key business decisions accurately by providing machine learning and analytics consulting. ABC is an established Brick & Mortar retailer that frequently conducts marketing campaigns for its diverse product range. As a merchant of XYZ, they have sought XYZ to assist them in their discount marketing process using the power of machine learning. Can you wear the AmExpert hat and help out ABC?
Discount marketing and coupon usage are very widely used promotional techniques to attract new customers and to retain & reinforce loyalty of existing customers. The measurement of a consumer’s propensity towards coupon usage and the prediction of the redemption behaviour are crucial parameters in assessing the effectiveness of a marketing campaign.
ABC’s promotions are shared across various channels including email, notifications, etc. A number of these campaigns include coupon discounts that are offered for a specific product/range of products. The retailer would like the ability to predict whether customers redeem the coupons received across channels, which will enable the retailer’s marketing team to accurately design coupon construct, and develop more precise and targeted marketing strategies.
The data available in this problem contains the following information, including the details of a sample of campaigns and coupons used in previous campaigns -
User Demographic Details Campaign and coupon Details Product details Previous transactions Based on previous transaction & performance data from the last 18 campaigns, predict the probability for the next 10 campaigns in the test set for each coupon and customer combination, whether the customer will redeem the coupon or not?
Dataset Description Here is the schema for the different data tables available. The detailed data dictionary is provided next.
You are provided with the following files in train.zip:
train.csv: Train data containing the coupons offered to the given customers under the 18 campaigns
Variable Definition id Unique id for coupon customer impression campaign_id Unique id for a discount campaign coupon_id Unique id for a discount coupon customer_id Unique id for a customer redemption_status (target) (0 - Coupon not redeemed, 1 - Coupon redeemed) campaign_data.csv: Campaign information for each of the 28 campaigns
Variable Definition campaign_id Unique id for a discount campaign campaign_type Anonymised Campaign Type (X/Y) start_date Campaign Start Date end_date Campaign End Date coupon_item_mapping.csv: Mapping of coupon and items valid for discount under that coupon
Variable Definition coupon_id Unique id for a discount coupon (no order) item_id Unique id for items for which given coupon is valid (no order) customer_demographics.csv: Customer demographic information for some customers
Variable Definition customer_id Unique id for a customer age_range Age range of customer family in years marital_status Married/Single rented 0 - not rented accommodation, 1 - rented accommodation family_size Number of family members no_of_children Number of children in the family income_bracket Label Encoded Income Bracket (Higher income corresponds to higher number) customer_transaction_data.csv: Transaction data for all customers for duration of campaigns in the train data
Variable Definition date Date of Transaction customer_id Unique id for a customer item_id Unique id for item quantity quantity of item bought selling_price Sales value of the transaction other_discount Discount from other sources such as manufacturer coupon/loyalty card coupon_discount Discount availed from retailer coupon item_data.csv: Item information for each item sold by the retailer
Variable Definition item_id Unique id for item brand Unique id for item brand brand_type Brand Type (local/Established) category Item Category test.csv: Contains the coupon customer combination for which redemption status is to be predicted
Variable Definition id Unique id for coupon customer impression campaign_id Unique id for a discount campaign coupon_id Unique id for a discount coupon customer_id Unique id for a customer *Campaign, coupon and customer data for test set is also contained in train.zip
sample_submission.csv: This file contains the format in which you have to submit your predictions.
To summarise the entire process:
Customers receive coupons under various campaigns and may choose to redeem it. They can redeem the given coupon for any valid product for that coupon as per coupon item mapping within the duration between campaign start date and end date Next, the customer will redeem the coupon for an item at the retailer store and that will reflect in the transaction table in the column coupon_discount.
```
'''
https://www.kaggle.com/bharath901/amexpert-2019/data#
'''
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
pd.set_option('display.max_columns', None)
pd.set_option('display.max_rows', None)
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import classification_report, precision_score, recall_score, f1_score, accuracy_score
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import confusion_matrix
from sklearn.feature_selection import RFE
import csv
```
Import Function to various files
```
'''
Defining a Class to import the Data into a pandas DataFrame for analysis
- Method to storing the Data into a DataFrame
- Method to extracting Information of the Data to understand datatype associated with each column
- Method to describing the Data
- Method to understanding Null values distribution
- Method to understanding Unique values distribution
'''
class import_data():
'''
Method to extract and store the data as pandas dataframe
'''
def __init__(self,path):
self.raw_data = pd.read_csv(path)
display (self.raw_data.head(10))
'''
Method to extract information about the data and display
'''
def get_info(self):
display (self.raw_data.info())
'''
Method to describe the data
'''
def get_describe(self):
display (self.raw_data.describe())
'''
Mehtod to understand Null values distribution
'''
def null_value(self):
col_null = pd.DataFrame(self.raw_data.isnull().sum()).reset_index()
col_null.columns = ['DataColumns','NullCount']
col_null['NullCount_Pct'] = round((col_null['NullCount']/self.raw_data.shape[0])*100,2)
display (col_null)
'''
Method to understand Unique values distribution
'''
def unique_value(self):
col_uniq = pd.DataFrame(self.raw_data.nunique()).reset_index()
col_uniq.columns = ['DataColumns','UniqCount']
col_uniq_cnt = pd.DataFrame(self.raw_data.count(axis=0)).reset_index()
col_uniq_cnt.columns = ['DataColumns','UniqCount']
col_uniq['UniqCount_Pct'] = round((col_uniq['UniqCount']/col_uniq_cnt['UniqCount'])*100,2)
display (col_uniq)
'''
Method to return the dataset as dataframe
'''
def return_data(self):
base_loan_data = self.raw_data
return (base_loan_data)
'''
Evaluation and Analysis starts here for train.csv
'''
#path = str(input('Enter the path to load the dataset:'))
path = './data/train.csv'
print ('='*100)
data = import_data(path)
#data.get_info()
#data.null_value()
#data.unique_value()
#data.get_describe()
train_data = data.return_data()
'''
Evaluation and Analysis starts here for Campaign Data
'''
#path = str(input('Enter the path to load the dataset:'))
path = './data/campaign_data.csv'
print ('='*100)
data = import_data(path)
#data.get_info()
#data.null_value()
#data.unique_value()
#data.get_describe()
campaign_data = data.return_data()
'''
Evaluation and Analysis starts here for Coupon Data
'''
#path = str(input('Enter the path to load the dataset:'))
path = './data/coupon_item_mapping.csv'
print ('='*100)
data = import_data(path)
#data.get_info()
#data.null_value()
#data.unique_value()
#data.get_describe()
coupon_data = data.return_data()
'''
Evaluation and Analysis starts here for Item Data
'''
#path = str(input('Enter the path to load the dataset:'))
path = './data/item_data.csv'
print ('='*100)
data = import_data(path)
#data.get_info()
#data.null_value()
#data.unique_value()
#data.get_describe()
item_data = data.return_data()
'''
Evaluation and Analysis starts here for Customer Demographic
'''
#path = str(input('Enter the path to load the dataset:'))
path = './data/customer_demographics.csv'
print ('='*100)
data = import_data(path)
#data.get_info()
#data.null_value()
#data.unique_value()
#data.get_describe()
cust_demo_data = data.return_data()
'''
Evaluation and Analysis starts here for Customer Transaction
'''
#path = str(input('Enter the path to load the dataset:'))
path = './data/customer_transaction_data.csv'
print ('='*100)
data = import_data(path)
#data.get_info()
#data.null_value()
#data.unique_value()
#data.get_describe()
cust_tran_data = data.return_data()
'''
Function to write the experiment result to csv file
'''
file_write_cnt = 1
# writing the baseline results to csv file
def write_file(F1,F2,F3,F4,F5,F6,F7,F8,F9,F10,F11):
# field names
fields = ['Expt No','Outlier Treatment','Skewness Treatment','Null Treatment',
'No of Features','Feature Selected','Model Used','Precision for 1',
'Recall for 1','Accuracy','Comment']
# data in the file
rows = [[F1,F2,F3,F4,F5,F6,F7,F8,F9,F10,F11]]
# name of csv file
filename = "./data/score_dashboard.csv"
if int(F1) == 1:
# writing to csv file
with open(filename, 'w') as csvfile:
# creating a csv writer object
csvwriter = csv.writer(csvfile)
# writing the fields
csvwriter.writerow(fields)
# writing the data rows
csvwriter.writerows(rows)
if int(F1) > 1:
# writing to csv file
with open(filename, 'a') as csvfile:
# creating a csv writer object
csvwriter = csv.writer(csvfile)
# writing the data rows
csvwriter.writerows(rows)
'''
Function to convert the date into quarter
'''
def date_q(date,split):
if split == '/':
"""
Convert Date to Quarter when separated with /
"""
qdate = date.strip().split('/')[1:]
qdate1 = qdate[0]
if qdate1 in ['01','02','03']:
return (str('Q1' + '-' + qdate[1]))
if qdate1 in ['04','05','06']:
return (str('Q2' + '-' + qdate[1]))
if qdate1 in ['07','08','09']:
return (str('Q3' + '-' + qdate[1]))
if qdate1 in ['10','11','12']:
return (str('Q4' + '-' + qdate[1]))
if split == '-':
"""
Convert Date to Quarter when separated with -
"""
qdate = date.strip().split('-')[0:2]
qdate1 = qdate[1]
qdate2 = str(qdate[0])
if qdate1 in ['01','02','03']:
return (str('Q1' + '-' + qdate2[2:]))
if qdate1 in ['04','05','06']:
return (str('Q2' + '-' + qdate2[2:]))
if qdate1 in ['07','08','09']:
return (str('Q3' + '-' + qdate2[2:]))
if qdate1 in ['10','11','12']:
return (str('Q4' + '-' + qdate2[2:]))
'''
Function to aggregate Customer Transaction Data
'''
def tran_summation(column):
cust_tran_data_expt['tot_'+column] = pd.DataFrame(cust_tran_data_expt.groupby(['customer_id','item_id','coupon_id'])[column].transform('sum'))
cust_tran_data_expt.drop([column],axis=1,inplace=True)
def tran_summation_1(column):
cust_tran_data_expt['tot_'+column] = pd.DataFrame(cust_tran_data_expt.groupby(['customer_id','item_id','coupon_id','tran_date_q'])[column].transform('sum'))
cust_tran_data_expt.drop([column],axis=1,inplace=True)
def tran_summation_2(column):
cust_tran_data_expt['tot_'+column] = pd.DataFrame(cust_tran_data_expt.groupby(['customer_id','coupon_id'])[column].transform('sum'))
cust_tran_data_expt.drop([column],axis=1,inplace=True)
'''
Function to label encode
'''
def label_encode(column):
train_data_merge[column] = train_data_merge[column].astype('category').cat.codes
'''
Function to convert Categorical column to Integer using Coupon Redemption percentage
'''
def cat_percent(column):
train_data_merge[column+'_redeem_sum'] = pd.DataFrame(train_data_merge.groupby([column])['redemption_status'].transform('sum'))
train_data_merge[column+'_redeem_count'] = pd.DataFrame(train_data_merge.groupby([column])['redemption_status'].transform('count'))
train_data_merge[column+'_redeem_percent'] = pd.DataFrame(train_data_merge[column+'_redeem_sum']*100/train_data_merge[column+'_redeem_count'])
train_data_merge.drop(column,axis=1,inplace=True)
train_data_merge.drop([column+'_redeem_sum'],axis=1,inplace=True)
train_data_merge.drop([column+'_redeem_count'],axis=1,inplace=True)
```
# To define baseline model with basic data preprocessing.
```
cust_demo_data_expt = cust_demo_data.copy()
cust_demo_data_expt['marital_status'].fillna('Unspecified',inplace=True)
cust_demo_data_expt['no_of_children'].fillna(0,inplace=True)
cust_demo_data_expt['age_range'].replace(['18-25','26-35','36-45','46-55','56-70','70+'],[18,26,36,46,56,70],inplace=True)
cust_demo_data_expt['family_size'].replace('5+',5,inplace=True)
cust_demo_data_expt['no_of_children'].replace('3+',3,inplace=True)
cust_tran_data_expt = cust_tran_data.copy()
cust_tran_data_expt = pd.merge(cust_tran_data_expt,coupon_data,how='inner',on='item_id')
cust_tran_data_expt.drop('date',axis=1,inplace=True)
for column in ['quantity','coupon_discount','other_discount','selling_price']:
tran_summation(column)
cust_tran_data_expt.drop_duplicates(subset=['customer_id','item_id','coupon_id'], keep='first', inplace=True)
train_data_merge = pd.merge(train_data,cust_tran_data_expt,how='inner',on=['customer_id','coupon_id'])
train_data_merge = pd.merge(train_data_merge,cust_demo_data_expt,how='left',on='customer_id')
train_data_merge = pd.merge(train_data_merge,item_data,how='left',on='item_id')
train_data_merge.drop('marital_status',axis=1,inplace=True)
train_data_merge.fillna({'age_range':0,'rented':0,'family_size':0,'no_of_children':0,'income_bracket':0},inplace=True)
train_data_merge['family_size'].astype('int8')
train_data_merge['no_of_children'].astype('int8')
train_data_merge = pd.get_dummies(train_data_merge, columns=['brand_type','category'], drop_first=False)
X = train_data_merge.drop('redemption_status', axis=1)
y = train_data_merge['redemption_status']
X_train,X_test,y_train,y_test = train_test_split(X,y,train_size=0.7,random_state=7)
# defining the model
classifier = LogisticRegression(solver='lbfgs',max_iter=10000)
# fitting the model
classifier.fit(X_train,y_train)
# predicting test result with model
y_pred = classifier.predict(X_test)
# Creating Classification report for Logistic Regression Baseline model
print ("Classification Report for Baseline Logistic Regression")
print(classification_report(y_test,y_pred))
report = pd.DataFrame(classification_report(y_test,y_pred,output_dict=True)).transpose()
write_file(file_write_cnt,'No','No','Yes',len(X.columns),list(X.columns),'Logistic Regresssion',report['precision'][1],report['recall'][1],report['support']['accuracy'],'Baseline Model')
file_write_cnt = file_write_cnt + 1
# defining the model
classifier = RandomForestClassifier(n_estimators=100)
# fitting the model
classifier.fit(X_train,y_train)
# predicting test result with model
y_pred = classifier.predict(X_test)
# Creating Classification report for RandomForest Classifier Baseline model
print ("Classification Report for Baseline RandomForest Classifier")
print(classification_report(y_test,y_pred))
report = pd.DataFrame(classification_report(y_test,y_pred,output_dict=True)).transpose()
write_file(file_write_cnt,'No','No','Yes',len(X.columns),X.columns,'Random Forest Classifier',report['precision'][1],report['recall'][1],report['support']['accuracy'],'Baseline Model')
file_write_cnt += 1
```
# One Hot Encoding
```
del train_data_merge
train_data_merge = pd.merge(train_data,cust_tran_data_expt,how='inner',on=['customer_id','coupon_id'])
train_data_merge = pd.merge(train_data_merge,cust_demo_data,how='left',on='customer_id')
train_data_merge = pd.merge(train_data_merge,item_data,how='left',on='item_id')
train_data_merge['no_of_children'].fillna(0,inplace=True)
train_data_merge.fillna({'marital_status':'Unspecified','rented':'Unspecified','family_size':'Unspecified','age_range':'Unspecified'},inplace=True)
train_data_merge['income_bracket'].fillna(train_data_merge['income_bracket'].mean(),inplace=True)
train_data_merge.drop(['id'],axis=1,inplace=True)
train_data_merge['no_of_children'].replace('3+',3,inplace=True)
train_data_merge['no_of_children'].astype('int')
train_data_merge = pd.get_dummies(train_data_merge, columns=['age_range','marital_status','rented','family_size','brand_type','category'], drop_first=False)
X = train_data_merge.drop('redemption_status', axis=1)
y = train_data_merge['redemption_status']
X_train,X_test,y_train,y_test = train_test_split(X,y,train_size=0.7,random_state=7)
# defining the model
classifier = RandomForestClassifier(n_estimators=100)
# fitting the model
classifier.fit(X_train,y_train)
# predicting test result with model
y_pred = classifier.predict(X_test)
# Creating Classification report for RandomForest Classifier Baseline model
print ("Classification Report for RandomForest Classifier")
print(classification_report(y_test,y_pred))
report = pd.DataFrame(classification_report(y_test,y_pred,output_dict=True)).transpose()
write_file(file_write_cnt,'No','No','Yes',len(X.columns),X.columns,'Random Forest Classfier',report['precision'][1],report['recall'][1],report['support']['accuracy'],'with OHE')
file_write_cnt += 1
```
# Label Encoding
```
del train_data_merge
train_data_merge = pd.merge(train_data,cust_tran_data_expt,how='inner',on=['customer_id','coupon_id'])
train_data_merge = pd.merge(train_data_merge,cust_demo_data,how='left',on='customer_id')
train_data_merge = pd.merge(train_data_merge,item_data,how='left',on='item_id')
train_data_merge['no_of_children'].fillna(0,inplace=True)
train_data_merge.fillna({'marital_status':'Unspecified','rented':'Unspecified','family_size':'Unspecified','age_range':'Unspecified'},inplace=True)
train_data_merge['income_bracket'].fillna(train_data_merge['income_bracket'].mean(),inplace=True)
train_data_merge['no_of_children'].replace('3+',3,inplace=True)
train_data_merge['no_of_children'].astype('int')
train_data_merge.drop(['id'],axis=1,inplace=True)
for column in ['marital_status','rented','family_size','age_range','brand_type','category']:
label_encode(column)
X = train_data_merge.drop('redemption_status', axis=1)
y = train_data_merge['redemption_status']
X_train,X_test,y_train,y_test = train_test_split(X,y,train_size=0.7,random_state=7)
# defining the model
classifier = RandomForestClassifier(n_estimators=100)
# fitting the model
classifier.fit(X_train,y_train)
# predicting test result with model
y_pred = classifier.predict(X_test)
# Creating Classification report for RandomForest Classifier Baseline model
print ("Classification Report for RandomForest Classifier")
print(classification_report(y_test,y_pred))
report = pd.DataFrame(classification_report(y_test,y_pred,output_dict=True)).transpose()
write_file(file_write_cnt,'No','No','Yes',len(X.columns),X.columns,'Random Forest Classifier',report['precision'][1],report['recall'][1],report['support']['accuracy'],'with Label Encoding')
file_write_cnt += 1
```
# Feature Engineering with Treatment of Campaign Date, Transaction Date and using Coupon Redemption percentage as a value to convert categorical columns to integers
```
del train_data_merge
del cust_tran_data_expt
campaign_data_expt = campaign_data.copy()
campaign_data_expt['start_date_q'] = campaign_data_expt['start_date'].map(lambda x: date_q(x,'/'))
campaign_data_expt['end_date_q'] = campaign_data_expt['end_date'].map(lambda x: date_q(x,'/'))
campaign_data_expt.drop(['start_date','end_date'],axis=1,inplace=True)
cust_tran_data_expt = cust_tran_data.copy()
cust_tran_data_expt = pd.merge(cust_tran_data_expt,coupon_data,how='inner',on='item_id')
cust_tran_data_expt['tran_date_q'] = cust_tran_data_expt['date'].map(lambda x: date_q(x,'-'))
cust_tran_data_expt.drop('date',axis=1,inplace=True)
for column in ['quantity','coupon_discount','other_discount','selling_price']:
tran_summation_1(column)
cust_tran_data_expt.drop_duplicates(subset=['customer_id','item_id','coupon_id','tran_date_q'], keep='first', inplace=True)
train_data_merge = pd.merge(train_data,cust_tran_data_expt,how='inner',on=['customer_id','coupon_id'])
train_data_merge = pd.merge(train_data_merge,cust_demo_data,how='left',on='customer_id')
train_data_merge = pd.merge(train_data_merge,item_data,how='left',on='item_id')
train_data_merge = pd.merge(train_data_merge,campaign_data_expt,how='left',on='campaign_id')
train_data_merge['no_of_children'].fillna(0,inplace=True)
train_data_merge.fillna({'marital_status':'Unspecified','rented':'Unspecified','family_size':'Unspecified','age_range':'Unspecified','income_bracket':'Unspecified'},inplace=True)
train_data_merge.drop('id',axis=1,inplace=True)
train_data_merge['no_of_children'].replace('3+',3,inplace=True)
for column in ['customer_id','coupon_id','item_id','campaign_id','no_of_children','marital_status','rented','family_size','age_range','income_bracket','start_date_q','end_date_q','tran_date_q','brand','category']:
cat_percent(column)
train_data_merge = pd.get_dummies(train_data_merge, columns=['campaign_type','brand_type'], drop_first=False)
X = train_data_merge.drop('redemption_status', axis=1)
y = train_data_merge['redemption_status']
feature_sel = [5,10,15,23]
rforc = RandomForestClassifier(n_estimators=100)
for i in feature_sel:
rfe = RFE(rforc, i)
rfe.fit(X, y)
# Selecting columns
sel_cols = []
for a, b, c in zip(rfe.support_, rfe.ranking_, X.columns):
if b == 1:
sel_cols.append(c)
print ('Number of features selected are ::',i)
print ('Columns Selected are ::',sel_cols)
# Creating new DataFrame with selected columns only as X
X_sel = X[sel_cols]
# Split data in to train and test
X_sel_train, X_sel_test, y_sel_train, y_sel_test = train_test_split(X_sel, y, train_size=0.7, random_state=7)
# Fit and Predict the model using selected number of features
grid={"n_estimators":[100]}
rforc_cv = GridSearchCV(rforc,grid,cv=10)
rforc_cv.fit(X_sel_train, y_sel_train)
rforc_pred = rforc_cv.predict(X_sel_test)
# Classification Report
print(classification_report(y_sel_test,rforc_pred))
report = pd.DataFrame(classification_report(y_sel_test,rforc_pred,output_dict=True)).transpose()
write_file(file_write_cnt,'No','No','Yes',len(X_sel.columns),X_sel.columns,'Random Forest Classifier',report['precision'][1],report['recall'][1],report['support']['accuracy'],'Treating Date and Label Encoding with RFE')
file_write_cnt += 1
```
| github_jupyter |
# PyDMD
### Tutorial 3: Multiresolution DMD: different time scales
In this tutorial we will show the possibilities of the multiresolution dynamic modes decomposition (mrDMD) with respect to the classical DMD. We follow a wonderful blog post written by Robert Taylor [available here](http://www.pyrunner.com/weblog/2016/08/05/mrdmd-python/). We did not use his implementation of the mrDMD but only the sample data and the structure of the tutorial. You can find a mathematical reference for the mrDMD by Kutz et al. [here](http://epubs.siam.org/doi/pdf/10.1137/15M1023543).
For the advanced settings of the DMD base class please refer to [this tutorial](https://github.com/mathLab/PyDMD/blob/master/tutorials/tutorial-2-adv-dmd.ipynb).
First of all we just import the MrDMD and DMD classes from the pydmd package, we set matplotlib for the notebook and we import numpy.
```
%matplotlib inline
import matplotlib.pyplot as plt
from pydmd import MrDMD
from pydmd import DMD
import numpy as np
```
The code below generates a spatio-temporal example dataset. The data can be thought of as 80 locations or signals (the x-axis) being sampled 1600 times at a constant rate in time (the t-axis). It contains many features at varying time scales, like oscillating sines and cosines, one-time events, and random noise.
```
def create_sample_data():
x = np.linspace(-10, 10, 80)
t = np.linspace(0, 20, 1600)
Xm, Tm = np.meshgrid(x, t)
D = np.exp(-np.power(Xm/2, 2)) * np.exp(0.8j * Tm)
D += np.sin(0.9 * Xm) * np.exp(1j * Tm)
D += np.cos(1.1 * Xm) * np.exp(2j * Tm)
D += 0.6 * np.sin(1.2 * Xm) * np.exp(3j * Tm)
D += 0.6 * np.cos(1.3 * Xm) * np.exp(4j * Tm)
D += 0.2 * np.sin(2.0 * Xm) * np.exp(6j * Tm)
D += 0.2 * np.cos(2.1 * Xm) * np.exp(8j * Tm)
D += 0.1 * np.sin(5.7 * Xm) * np.exp(10j * Tm)
D += 0.1 * np.cos(5.9 * Xm) * np.exp(12j * Tm)
D += 0.1 * np.random.randn(*Xm.shape)
D += 0.03 * np.random.randn(*Xm.shape)
D += 5 * np.exp(-np.power((Xm+5)/5, 2)) * np.exp(-np.power((Tm-5)/5, 2))
D[:800,40:] += 2
D[200:600,50:70] -= 3
D[800:,:40] -= 2
D[1000:1400,10:30] += 3
D[1000:1080,50:70] += 2
D[1160:1240,50:70] += 2
D[1320:1400,50:70] += 2
return D.T
```
Here we have an auxiliary function that we will use to plot the data.
```
def make_plot(X, x=None, y=None, figsize=(12, 8), title=''):
"""
Plot of the data X
"""
plt.figure(figsize=figsize)
plt.title(title)
X = np.real(X)
CS = plt.pcolor(x, y, X)
cbar = plt.colorbar(CS)
plt.xlabel('Space')
plt.ylabel('Time')
plt.show()
```
Let us start by creating the dataset and plot the data in order to have a first idea of the problem.
```
sample_data = create_sample_data()
x = np.linspace(-10, 10, 80)
t = np.linspace(0, 20, 1600)
make_plot(sample_data.T, x=x, y=t)
```
First we apply the classical DMD without the svd rank truncation, and then we try to reconstruct the data. You can clearly see that all the transient time events are missing.
```
first_dmd = DMD(svd_rank=-1)
first_dmd.fit(X=sample_data)
make_plot(first_dmd.reconstructed_data.T, x=x, y=t)
```
Now we do the same but using the mrDMD instead. The result is remarkable even with the svd rank truncation (experiment changing the input parameters).
```
sub_dmd = DMD(svd_rank=-1)
dmd = MrDMD(sub_dmd, max_level=7, max_cycles=1)
dmd.fit(X=sample_data)
make_plot(dmd.reconstructed_data.T, x=x, y=t)
```
Ok, pretty amazing eh? Let us have a look at the eigenvalues in order to better understand the mrDMD. Without truncation we have 80 eigenvalues.
```
print('The number of eigenvalues is {}'.format(dmd.eigs.shape[0]))
dmd.plot_eigs(show_axes=True, show_unit_circle=True, figsize=(8, 8))
```
It is also possible to plot only specific eigenvalues, given the level and the node. If the node is not provided all the eigenvalues of that level will be plotted.
```
dmd.plot_eigs(show_axes=True, show_unit_circle=True, figsize=(8, 8), level=3, node=0)
```
The idea is to extract the slow modes at each iteration, where a slow mode is a mode with a relative low frequency. This just means that the mode changes somewhat slowly as the system evolves in time. Thus the mrDMD is able to catch different time events.
The general mrDMD algorithm is as follows:
1. Compute DMD for available data.
2. Determine fast and slow modes.
3. Find the best DMD approximation to the available data constructed from the slow modes only.
4. Subtract off the slow-mode approximation from the available data.
5. Split the available data in half.
6. Repeat the procedure for the first half of data (including this step).
7. Repeat the procedure for the second half of data (including this step).
Let us have a look at the modes for the first two levels and the corresponding time evolution. At the first level we have two very slow modes, while at the second one there are 5 modes.
```
pmodes = dmd.partial_modes(level=0)
fig = plt.plot(x, pmodes.real)
pdyna = dmd.partial_dynamics(level=0)
fig = plt.plot(t, pdyna.real.T)
```
Notice the discontinuities in the time evolution where the data were split.
```
pdyna = dmd.partial_dynamics(level=1)
print('The number of modes in the level number 1 is {}'.format(pdyna.shape[0]))
fig = plt.plot(t, pdyna.real.T)
```
Now we recreate the original data by adding levels together. For each level, starting with the first (note that the starting index is 0), we construct an approximation of the data.
```
pdata = dmd.partial_reconstructed_data(level=0)
make_plot(pdata.T, x=x, y=t, title='level 0', figsize=(7.5, 5))
```
Then, we sequentially add them all together, one on top of another. It is interesting to see how the original data has been broken into features of finer and finer resolution.
```
for i in range(1, 7):
pdata += dmd.partial_reconstructed_data(level=i)
make_plot(pdata.T, x=x, y=t, title='levels 0-' + str(i), figsize=(7.5, 5))
```
The multiresolution DMD has been employed in many different fields of study due to its versatility. Feel free to share with us your applications!
| github_jupyter |
```
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from torch.utils.data import DataLoader, Dataset
from sklearn.preprocessing import LabelEncoder, OneHotEncoder, StandardScaler
import pandas as pd
import numpy as np
import pickle
import random
import json
import gc
import os
import re
from imblearn.over_sampling import SMOTE
from datetime import datetime
import time
import matplotlib.pyplot as plt
import seaborn as sns
import warnings
warnings.simplefilter('ignore')
def seed_everything(seed=42):
print('Setting Random Seed')
random.seed(seed)
os.environ['PYTHONHASHSEED'] = str(seed)
np.random.seed(seed)
torch.manual_seed(seed)
torch.cuda.manual_seed(seed)
torch.backends.cudnn.deterministic = True
TARGET = 'service_canceled'
fts_continuous = ['customer_age_appl', 'time_start_process', 'operator_count', 'previous_customer_count']
fts_categorical = ['date', 'branch_name', 'customer_gender', 'customer_city', 'service_name_organization',
'service_name', 'service_name_2']
print('Categorical Features', fts_categorical)
print('Continuous Features', fts_continuous)
print('Categorical Feature Count', len(fts_categorical))
print('Continuous Feature Count', len(fts_continuous))
```
## Configuration
```
PATH = "./"
SAVE_PATH = "./outputs/"
os.makedirs(SAVE_PATH, exist_ok=True)
run_key = 'DAE_TRIAL' #due to long run time prefer to split into sections of 600 epochs
DAE_CFG = {'debug': False,
'batch_size': 384,
'init_lr': 0.0003,
'lr_decay': 0.998, # rate of decrease of learning rate
'noise_decay': 0.999, # rate of decrease of noise level
'max_epochs': 2000,
'save_freq': 100,
'hidden_size': 1024, # hidden_size == embed_dim * num_subspaces
'num_subspaces': 8,
'embed_dim': 128,
'num_heads': 8,
'dropout': 0,
'feedforward_dim': 512,
'emphasis': 0.75, #weighing of loss to 'corrupted' data points - i tried varying over time, did not show immediate improvement
'task_weights': [len(fts_categorical), len(fts_continuous)], #weighting for categorical vs continuous
'mask_loss_weight': 2, #weighting of mask prediction vs prediction of reconstructed original data values
'prob_categorical': 0.2, #probability of noise in categoricals
'prob_continuous': 0.1, #probability of noise in continuous
'random_state': 2021,
'run_key': run_key
}
if DAE_CFG['debug']:
DAE_CFG['max_epochs'] = 10
with open(SAVE_PATH + f"{DAE_CFG['run_key']}_DAE_CFG.pickle", 'wb') as f:
pickle.dump(DAE_CFG, f)
with open(SAVE_PATH + f"{DAE_CFG['run_key']}_DAE_CFG.json", 'w') as f:
json.dump(DAE_CFG, f)
```
## Check Noise and Learning Rate
```
tracking_df = pd.DataFrame(index=range(DAE_CFG['max_epochs']),
columns=['loss', 'lr', 'run_code', 'time', 'elapsed', 'noise_categorical', 'noise_continuous'],
data=0.0)
tracking_df['lr'] = DAE_CFG['init_lr'] * (DAE_CFG['lr_decay']**tracking_df.index)
tracking_df['noise_categorical'] = DAE_CFG['prob_categorical'] * (DAE_CFG['noise_decay']**tracking_df.index)
tracking_df['noise_continuous'] = DAE_CFG['prob_continuous'] * (DAE_CFG['noise_decay']**tracking_df.index)
tracking_df['run_code'] = DAE_CFG['run_key']
sns.set(font_scale=1.4)
fig,axes=plt.subplots(nrows=1,ncols=3,figsize=(18,6))
axes[0].plot(tracking_df.index, tracking_df['lr'], color='Blue')
axes[0].set_ylim(0,)
axes[0].set_title('Learning Rate')
axes[0].set_xlabel('Epochs')
axes[1].plot(tracking_df.index, tracking_df['noise_categorical'], color='Red')
axes[1].set_ylim(0,1)
axes[1].set_title('Categorical Noise Prob')
axes[2].plot(tracking_df.index, tracking_df['noise_continuous'], color='Red')
axes[2].set_ylim(0,1)
axes[2].set_title('Continuous Noise Prob')
plt.tight_layout()
```
# Functions to Get Data
```
def feats_engineering(train, test):
unique_counts = list()
all_df = pd.concat([train, test]).reset_index(drop=True)
all_df['customer_age_appl'].fillna(all_df['customer_age_appl'].mode()[0], inplace=True)
all_df['time_start_process'].fillna(all_df['time_start_process'].mode()[0], inplace=True)
all_df['customer_age_appl'] = all_df['customer_age_appl'].apply(lambda x: (int(x.split('-')[0])+int(x.split('-')[1]))/2)
all_df['time_start_process'] = all_df['time_start_process'].apply(lambda x: int(x[:2]))
for col in fts_categorical:
unique_counts.append(all_df[col].nunique())
df_train = all_df[:train.shape[0]]
df_test = all_df[train.shape[0]:].reset_index(drop=True)
return df_train, df_test, unique_counts
def get_data():
train = pd.read_csv(PATH + "queue_dataset_train_small_sample.csv")
test = pd.read_csv(PATH + "queue_dataset_test.csv")
train_data, test_data, unique_counts = feats_engineering(train, test)
#combine train and test data vertically
X_nums = np.vstack([
train_data.loc[:, fts_continuous].to_numpy(),
test_data.loc[:, fts_continuous].to_numpy()
])
X_nums = (X_nums - X_nums.mean(0)) / X_nums.std(0) #normalize
#stack the categorical data
X_cat = np.vstack([
train_data.loc[:, fts_categorical].to_numpy(),
test_data.loc[:, fts_categorical].to_numpy()
])
#encode the categoricals
encoder = OneHotEncoder(sparse=False)
X_cat = encoder.fit_transform(X_cat)
#join the categorical and continuous data horizontally
X = np.hstack([X_cat, X_nums])
y = train_data[TARGET].to_numpy().reshape(-1, 1)
return X, y, X_cat.shape[1], X_nums.shape[1], unique_counts #this lets us know how many categorical and continuous features there are
class SingleDataset(Dataset):
def __init__(self, x, is_sparse=False):
self.x = x.astype('float32')
self.is_sparse = is_sparse
def __len__(self):
return self.x.shape[0]
def __getitem__(self, index):
x = self.x[index]
if self.is_sparse: x = x.toarray().squeeze()
return x
```
# Losses
```
bce_logits = torch.nn.functional.binary_cross_entropy_with_logits
mse = torch.nn.functional.mse_loss
```
# AutoEncoder
```
#torch docs
#embed_dim – total dimension of the model.
#num_heads – parallel attention heads.
#dropout – a Dropout layer on attn_output_weights. Default: 0.0.
#bias – add bias as module parameter. Default: True.
#add_bias_kv – add bias to the key and value sequences at dim=0.
#add_zero_attn – add a new batch of zeros to the key and value sequences at dim=1.
#kdim – total number of features in key. Default: None.
#vdim – total number of features in value. Default: None.
class TransformerEncoder(torch.nn.Module):
def __init__(self, embed_dim, num_heads, dropout, feedforward_dim):
super().__init__()
self.attn = torch.nn.MultiheadAttention(embed_dim, num_heads, dropout=dropout)
self.linear_1 = torch.nn.Linear(embed_dim, feedforward_dim)
self.linear_2 = torch.nn.Linear(feedforward_dim, embed_dim)
self.layernorm_1 = torch.nn.LayerNorm(embed_dim)
self.layernorm_2 = torch.nn.LayerNorm(embed_dim)
def forward(self, x_in):
attn_out, _ = self.attn(x_in, x_in, x_in)
x = self.layernorm_1(x_in + attn_out)
ff_out = self.linear_2(torch.nn.functional.relu(self.linear_1(x)))
x = self.layernorm_2(x + ff_out)
return x
class TransformerAutoEncoder(torch.nn.Module):
def __init__(
self,
num_inputs,
n_cats,
n_nums,
hidden_size=1024,
num_subspaces=8,
embed_dim=128,
num_heads=8,
dropout=0,
feedforward_dim=512,
emphasis=.75,
task_weights=[len(fts_categorical), len(fts_continuous)],
mask_loss_weight=2,
):
super().__init__()
assert hidden_size == embed_dim * num_subspaces
self.n_cats = n_cats
self.n_nums = n_nums
self.num_subspaces = num_subspaces
self.num_heads = num_heads
self.embed_dim = embed_dim
self.emphasis = emphasis
self.task_weights = np.array(task_weights) / sum(task_weights)
self.mask_loss_weight = mask_loss_weight
self.excite = torch.nn.Linear(in_features=num_inputs, out_features=hidden_size)
self.encoder_1 = TransformerEncoder(embed_dim, num_heads, dropout, feedforward_dim)
self.encoder_2 = TransformerEncoder(embed_dim, num_heads, dropout, feedforward_dim)
self.encoder_3 = TransformerEncoder(embed_dim, num_heads, dropout, feedforward_dim)
self.mask_predictor = torch.nn.Linear(in_features=hidden_size, out_features=num_inputs)
self.reconstructor = torch.nn.Linear(in_features=hidden_size + num_inputs, out_features=num_inputs)
def divide(self, x):
batch_size = x.shape[0]
x = x.reshape((batch_size, self.num_subspaces, self.embed_dim)).permute((1, 0, 2))
return x
def combine(self, x):
batch_size = x.shape[1]
x = x.permute((1, 0, 2)).reshape((batch_size, -1))
return x
def forward(self, x):
x = torch.nn.functional.relu(self.excite(x))
x = self.divide(x)
x1 = self.encoder_1(x)
x2 = self.encoder_2(x1)
x3 = self.encoder_3(x2)
x = self.combine(x3)
predicted_mask = self.mask_predictor(x)
reconstruction = self.reconstructor(torch.cat([x, predicted_mask], dim=1))
return (x1, x2, x3), (reconstruction, predicted_mask)
def split(self, t):
return torch.split(t, [self.n_cats, self.n_nums], dim=1)
def feature(self, x):
attn_outs, _ = self.forward(x)
return torch.cat([self.combine(x) for x in attn_outs], dim=1) #the feature is the data extracted for use in inference
def loss(self, x, y, mask, reduction='mean'):
_, (reconstruction, predicted_mask) = self.forward(x)
x_cats, x_nums = self.split(reconstruction)
y_cats, y_nums = self.split(y)
#weights are detemined by the emphasis - which is currently heavier weights for corrupted data (mask = 1)
w_cats, w_nums = self.split(mask * self.emphasis + (1 - mask) * (1 - self.emphasis))
#BCE loss for reconstructed vs actual categoricals
cat_loss = self.task_weights[0] * torch.mul(w_cats, bce_logits(x_cats, y_cats, reduction='none'))
#mse loss for reconstructed vs actual continuous
num_loss = self.task_weights[1] * torch.mul(w_nums, mse(x_nums, y_nums, reduction='none'))
#BCE+MSE = reconstruction loss
reconstruction_loss = torch.cat([cat_loss, num_loss], dim=1) if reduction == 'none' else cat_loss.mean() + num_loss.mean()
#mask loss = how well the model predicts which values are corrupted - can use BCE as this is 0/1
mask_loss = self.mask_loss_weight * bce_logits(predicted_mask, mask, reduction=reduction)
return reconstruction_loss + mask_loss if reduction == 'mean' else [reconstruction_loss, mask_loss]
```
# Noise Masker
```
class SwapNoiseMasker(object):
def __init__(self, probas, decay_rate):
self.probas = torch.from_numpy(np.array(probas))
self.decay_rate = decay_rate
def apply(self, X, epoch_number):
epoch_probas = self.probas * (self.decay_rate ** epoch_number)
#generates Y/N for swap / dont swap
should_swap = torch.bernoulli(epoch_probas.to(X.device) * torch.ones((X.shape)).to(X.device))
#switches data where swap = Y
corrupted_X = torch.where(should_swap == 1, X[torch.randperm(X.shape[0])], X)
#mask is whether data has been changed or not
#nb for one hot categorical data, presumably quite often mask != shouldswap, as 1 is swapped for 1 or 0 swapped for 0
mask = (corrupted_X != X).float()
return corrupted_X, mask
```
# Prepare Data
```
# get data
X, Y, n_cats, n_nums, unique_counts = get_data()
seed_everything(DAE_CFG['random_state'])
train_dl = DataLoader(
dataset=SingleDataset(X),
batch_size=DAE_CFG['batch_size'],
shuffle=True,
pin_memory=True,
drop_last=True
)
print(X.shape, Y.shape, n_cats, n_nums)
print(unique_counts)
```
# Define Column Noise Probabilities
```
#repeats of probabilities for one hot encoding which creates new columns for categoricals
repeats = [x for x in unique_counts] + [1 for x in range(len(fts_continuous))]
#probabilities for original columns
probas = [DAE_CFG['prob_categorical'] for x in range(len(fts_categorical))] + [DAE_CFG['prob_continuous'] for x in range(len(fts_continuous))]
#expand these to the one hot columns
swap_probas = sum([[p] * r for p, r in zip(probas, repeats)], [])
print('length', len(swap_probas))
print('examples', swap_probas[0:10], swap_probas[-len(fts_continuous):])
```
# Prepare DAE Model
```
# setup model
model_params = dict(
hidden_size=DAE_CFG['hidden_size'],
num_subspaces=DAE_CFG['num_subspaces'],
embed_dim=DAE_CFG['embed_dim'],
num_heads=DAE_CFG['num_heads'],
dropout=DAE_CFG['dropout'],
feedforward_dim=DAE_CFG['feedforward_dim'],
emphasis=DAE_CFG['emphasis'],
mask_loss_weight=DAE_CFG['mask_loss_weight']
)
dae = TransformerAutoEncoder(
num_inputs=X.shape[1],
n_cats=n_cats,
n_nums=n_nums,
**model_params
).cuda()
model_checkpoint = 'model_checkpoint.pth'
optimizer = torch.optim.Adam(dae.parameters(), lr=DAE_CFG['init_lr'])
scheduler = torch.optim.lr_scheduler.ExponentialLR(optimizer, gamma=DAE_CFG['lr_decay'])
```
# Training DAE Model
```
class AverageMeter(object):
def __init__(self):
self.reset()
def reset(self):
self.val, self.avg, self.sum, self.count = 0, 0, 0, 0
def update(self, val, n=1):
self.val = val
self.sum += val * n
self.count += n
self.avg = self.sum / self.count
noise_maker = SwapNoiseMasker(swap_probas, DAE_CFG['noise_decay'])
s0 = time.time()
for epoch in range(DAE_CFG['max_epochs']):
t0 = time.time()
dae.train()
meter = AverageMeter()
for i, x in enumerate(train_dl):
x = x.cuda()
x_corrputed, mask = noise_maker.apply(x, epoch) #added epoch to allow noise level to decrease over time
optimizer.zero_grad()
loss = dae.loss(x_corrputed, x, mask)
loss.backward()
optimizer.step()
meter.update(loss.detach().cpu().numpy())
delta1 = (time.time() - s0)
delta2 = (time.time() - t0)
scheduler.step()
remain = (DAE_CFG['max_epochs'] - (epoch+1)) * delta2
print(f"\r epoch {epoch:5d} - loss {meter.avg:.6f} - {delta2:4.2f} sec per epoch - {delta1:6.2f} sec elapsed - {remain:6.2f} sec remaining", end="")
model_checkpoint = f'model_checkpoint_{epoch}.pth'
tracking_df.loc[epoch, 'loss'] = meter.avg
tracking_df.loc[epoch, 'time'] = round(delta2, 2)
tracking_df.loc[epoch, 'elapsed'] = round(delta1, 2)
if epoch%DAE_CFG['save_freq']==0:
## print('Saving to checkpoint')
#as i have flat noise level across all columns, i just print the noise average
## print('average noise level', np.array(swap_probas).mean()*(DAE_CFG['noise_decay']**epoch))
model_checkpoint = SAVE_PATH + f"{DAE_CFG['run_key']}_model_checkpoint_{epoch}.pth"
torch.save({
"optimizer": optimizer.state_dict(),
"scheduler": scheduler.state_dict(),
"model": dae.state_dict()
}, model_checkpoint
)
model_checkpoint = SAVE_PATH + f"{DAE_CFG['run_key']}_model_checkpoint_final.pth"
torch.save({
"optimizer": optimizer.state_dict(),
"scheduler": scheduler.state_dict(),
"model": dae.state_dict()
}, model_checkpoint
)
tracking_df.to_csv(f"{DAE_CFG['run_key']}_tracking_loss.csv")
# extract features
dl = DataLoader(dataset=SingleDataset(X), batch_size=DAE_CFG['batch_size'], shuffle=False, pin_memory=True, drop_last=False)
features = []
dae.eval()
with torch.no_grad():
for x in dl:
features.append(dae.feature(x.cuda()).detach().cpu().numpy())
features = np.vstack(features)
display(features.shape)
np.save(SAVE_PATH + f"{DAE_CFG['run_key']}_dae_features_epoch{DAE_CFG['max_epochs']}.npy", features)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/DemonFlexCouncil/DDSP-48kHz-Stereo/blob/master/ddsp/colab/ddsp_train_and_timbre_transfer_48kHz_stereo.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
##### Copyright 2020 Google LLC.
Licensed under the Apache License, Version 2.0 (the "License");
```
# Copyright 2020 Google LLC. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ==============================================================================
```
# Train & Timbre Transfer--DDSP Autoencoder on GPU--48kHz/Stereo
Made by [Google Magenta](https://magenta.tensorflow.org/)--altered by [Demon Flex Council](https://demonflexcouncil.wixsite.com/demonflexcouncil)
This notebook demonstrates how to install the DDSP library and train it for synthesis based on your own data using command-line scripts. If run inside of Colaboratory, it will automatically use a free or Pro Google Cloud GPU, depending on your membership level.
<img src="https://storage.googleapis.com/ddsp/additive_diagram/ddsp_autoencoder.png" alt="DDSP Autoencoder figure" width="700">
**Note that bash commands are prefixed with a `!` inside of Colaboratory, but you would leave them out if running directly in a terminal.**
**A Little Background**
A producer friend of mine turned me on to Magenta’s DDSP, and I’m glad he did. In my mind it represents the way forward for AI music. Finally we have a glimpse inside the black box, with access to musical parameters as well as neural net hyperparameters. And DDSP leverages decades of studio knowledge by utilizing traditional processors like synthesizers and effects. One can envision a time when DDSP-like elements will sit at the heart of production DAWs.
According to Magenta’s paper, this algorithm was intended as proof of concept, but I wanted to bend it more towards a tool for producers. I bumped the sample rate up to 48kHz and made it stereo. I also introduced a variable render length so you can feed it loops or phrases. However, there are limits to this parameter. The total number of samples in your render length (number of seconds * 48000) must be evenly divisible by 800. In practice, this means using round-numbered or highly-divisible tempos (105, 96, 90, 72, 50…) or using material that does not depend on tempo.
Also note that longer render times may require a smaller batch size, which is currently set at 8 for a 4-second render. This may diminish audio quality, so use shorter render times if at all possible.
You can train with or without latent vectors, z(t), for the audio. There is a tradeoff here. No latent vectors allows for more pronounced shifts in the “Modify Conditioning” section, but the rendered audio sounds cloudier. Then again, sometimes cloudier is better. The default mode is latent vectors.
The dataset and audio primer files must be WAVE format, stereo, and 48kHz. Most DAWs and audio editors have a 48kHz export option, including the free [Audacity](https://www.audacityteam.org/). There appears to be a lower limit on the total size of the dataset, somewhere around 20MB. Anything lower than that and the TFRecord maker will create blank records (0 bytes). Also, Colaboratory may throw memory errors if it encounters large single audio files—cut the file into smaller pieces if this happens.
## **Step 1**--Install Dependencies
First we install the required dependencies with `pip` (takes about 5 minutes).
```
!pip install tensorflow==2.2
!pip install mir_eval
!pip install apache_beam
!pip install crepe
!pip install pydub
!pip3 install ffmpeg-normalize
import os
import re
import glob
import tensorflow as tf
```
## **Step 2**--Confirm you are running Tensorflow version 2.2.0.
This is the only version which will work with this notebook. If you see any other version than 2.2.0 below, factory restart your runtime (in the "Runtime" menu) and run Step 1 again.
```
!pip show tensorflow
```
## **Step 3**--Login and mount your Google Drive
This will require an authentication code. You should then be able to see your Drive in the file browser on the left panel--make sure you've clicked the folder icon on the far left side of your Internet browser.
```
from google.colab import drive
drive.mount('/content/drive', force_remount=True)
```
## **Step 4**--Set render length
Determines the length of audio slices for training and resynthesis. Decimals are OK.
```
RENDER_SECONDS = 4.0#@param {type:"number", min:1, max:10}
RENDER_SAMPLES = int(RENDER_SECONDS * 48000)
if ((RENDER_SAMPLES % 800) != 0):
raise ValueError("Number of samples at 48kHz must be divisble by 800.")
```
## **Step 5**--Latent vectors mode
Uncheck the box to train without z(t).
```
LATENT_VECTORS = True #@param{type:"boolean"}
```
## **Step 6**--Set your audio directory on Drive and get DDSP repository from Github
Find a folder on Drive where you want to upload audio files and store checkpoints. Then right-click on the folder and select "Copy path". Enter the path below.
```
DRIVE_DIR = "/content/drive/My Drive/test" #@param {type:"string"}
if LATENT_VECTORS:
!git clone https://github.com/DemonFlexCouncil/DDSP-48kHz-Stereo.git
else:
!git clone https://github.com/DemonFlexCouncil/DDSP-48kHz-Stereo-NoZ.git
AUDIO_DIR = '/content/data/audio'
!mkdir -p $AUDIO_DIR
AUDIO_FILEPATTERN = AUDIO_DIR + '/*'
AUDIO_INPUT_DIR = DRIVE_DIR + '/audio_input'
AUDIO_OUTPUT_DIR = DRIVE_DIR + '/audio_output'
CKPT_OUTPUT_DIR = DRIVE_DIR + '/ckpt'
SAVE_DIR = os.path.join(DRIVE_DIR, 'model')
%cd $DRIVE_DIR
!mkdir -p audio_input audio_output ckpt data model primers
```
## **Step 7**--Upload your audio files to Drive and create a TFRecord dataset
Put all of your training audio files in the "audio_input" directory inside the directory you set as DRIVE_DIR. The algorithm typically works well with audio from a single acoustic environment.
Preprocessing involves inferring the fundamental frequency (or "pitch") with [CREPE](http://github.com/marl/crepe), and computing the loudness. These features will then be stored in a sharded [TFRecord](https://www.tensorflow.org/tutorials/load_data/tfrecord) file for easier loading. Depending on the amount of input audio, this process usually takes a few minutes.
```
audio_files = glob.glob(os.path.join(AUDIO_INPUT_DIR, '*.wav'))
for fname in audio_files:
target_name = os.path.join(AUDIO_DIR,
os.path.basename(fname).replace(' ', '_'))
print('Copying {} to {}'.format(fname, target_name))
!cp "$fname" $target_name
TRAIN_TFRECORD = '/content/data/train.tfrecord'
TRAIN_TFRECORD_FILEPATTERN = TRAIN_TFRECORD + '*'
drive_data_dir = os.path.join(DRIVE_DIR, 'data')
drive_dataset_files = glob.glob(drive_data_dir + '/*')
# Make a new dataset.
if not glob.glob(AUDIO_FILEPATTERN):
raise ValueError('No audio files found. Please use the previous cell to '
'upload.')
if LATENT_VECTORS:
!python /content/DDSP-48kHz-Stereo/ddsp/training/data_preparation/prepare_tfrecord.py \
--input_audio_filepatterns=$AUDIO_FILEPATTERN \
--output_tfrecord_path=$TRAIN_TFRECORD \
--num_shards=10 \
--example_secs=$RENDER_SECONDS \
--alsologtostderr
else:
!python /content/DDSP-48kHz-Stereo-NoZ/ddsp/training/data_preparation/prepare_tfrecord.py \
--input_audio_filepatterns=$AUDIO_FILEPATTERN \
--output_tfrecord_path=$TRAIN_TFRECORD \
--num_shards=10 \
--example_secs=$RENDER_SECONDS \
--alsologtostderr
TRAIN_TFRECORD_DIR = DRIVE_DIR + '/data'
TRAIN_TFRECORD_DIR = TRAIN_TFRECORD_DIR.replace("My Drive", "My\ Drive")
!cp $TRAIN_TFRECORD_FILEPATTERN $TRAIN_TFRECORD_DIR
```
## **Step 8**--Save dataset statistics for timbre transfer
Quantile normalization helps match loudness of timbre transfer inputs to the
loudness of the dataset, so let's calculate it here and save in a pickle file.
```
if LATENT_VECTORS:
%cd /content/DDSP-48kHz-Stereo/ddsp/
else:
%cd /content/DDSP-48kHz-Stereo-NoZ/ddsp/
from colab import colab_utils
from training import data
TRAIN_TFRECORD = '/content/data/train.tfrecord'
TRAIN_TFRECORD_FILEPATTERN = TRAIN_TFRECORD + '*'
data_provider = data.TFRecordProvider(TRAIN_TFRECORD_FILEPATTERN, example_secs=RENDER_SECONDS)
dataset = data_provider.get_dataset(shuffle=False)
PICKLE_FILE_PATH = os.path.join(SAVE_DIR, 'dataset_statistics.pkl')
colab_utils.save_dataset_statistics(data_provider, PICKLE_FILE_PATH)
```
## **Step 9**--Train model
DDSP was designed to model a single instrument, but I've had more interesting results training it on sparse multi-timbral material. In this case, the neural network will attempt to model all timbres, but will likely associate certain timbres with different pitch and loudness conditions.
Note that [gin configuration](https://github.com/google/gin-config) files specify parameters for the both the model architecture (solo_instrument.gin) and the dataset (tfrecord.gin). These parameters can be overriden in the run script below (!python ddsp/ddsp_run.py).
### Training Notes:
* Models typically perform well when the loss drops to the range of ~7.0-8.5.
* Depending on the dataset this can take anywhere from 30k-90k training steps usually.
* The default is set to 90k, but you can stop training at any time (select "Interrupt execution" from the "Runtime" menu).
* On the Colaboratory Pro GPU, training takes about 3-9 hours. Free GPUs may be slower.
* By default, checkpoints will be saved every 300 steps with a maximum of 10 checkpoints.
* Feel free to adjust these numbers depending on the frequency of saves you would like and the space on your drive.
* If your Colaboratory runtime has stopped, re-run steps 1 through 9 to resume training from your most recent checkpoint.
```
if LATENT_VECTORS:
%cd /content/DDSP-48kHz-Stereo
else:
%cd /content/DDSP-48kHz-Stereo-NoZ
TRAIN_TFRECORD = '/content/data/train.tfrecord'
TRAIN_TFRECORD_FILEPATTERN = TRAIN_TFRECORD + '*'
!python ddsp/ddsp_run.py \
--mode=train \
--alsologtostderr \
--save_dir="$SAVE_DIR" \
--gin_file=models/solo_instrument.gin \
--gin_file=datasets/tfrecord.gin \
--gin_param="TFRecordProvider.file_pattern='$TRAIN_TFRECORD_FILEPATTERN'" \
--gin_param="TFRecordProvider.example_secs=$RENDER_SECONDS" \
--gin_param="Autoencoder.n_samples=$RENDER_SAMPLES" \
--gin_param="batch_size=8" \
--gin_param="train_util.train.num_steps=90000" \
--gin_param="train_util.train.steps_per_save=300" \
--gin_param="trainers.Trainer.checkpoints_to_keep=10"
```
## **Step 10**--Timbre transfer imports
Now it's time to render the final audio file with the aid of an audio primer file for timbre transfer. We'll start with some basic imports.
```
if LATENT_VECTORS:
%cd /content/DDSP-48kHz-Stereo/ddsp
else:
%cd /content/DDSP-48kHz-Stereo-NoZ/ddsp
# Ignore a bunch of deprecation warnings
import warnings
warnings.filterwarnings("ignore")
import copy
import time
import pydub
import gin
import crepe
import librosa
import matplotlib.pyplot as plt
import numpy as np
import pickle
import tensorflow as tf
import tensorflow_datasets as tfds
import core
import spectral_ops
from training import metrics
from training import models
from colab import colab_utils
from colab.colab_utils import (auto_tune, detect_notes, fit_quantile_transform, get_tuning_factor, download, play, record, specplot, upload, DEFAULT_SAMPLE_RATE)
from google.colab import files
# Helper Functions
sample_rate = 48000
print('Done!')
```
## **Step 11**--Process audio primer
The key to transcending the sonic bounds of the dataset is the audio primer file. This file will graft its frequency and loudness information onto the rendered audio file, sort of like a vocoder. Then you can use the sliders in the "Modify Conditioning" section to further alter the rendered file.
Put your audio primer files in the "primers" directory inside the directory you set as DRIVE_DIR. Input the file name of the primer you want to use on the line below.
```
PRIMER_DIR = DRIVE_DIR + '/primers/'
PRIMER_FILE = "OTO16S48a.wav" #@param {type:"string"}
# Check for .wav extension
match = re.search(r'.wav', PRIMER_FILE)
if match:
print ('')
else:
PRIMER_FILE = PRIMER_FILE + ".wav"
PATH_TO_PRIMER = PRIMER_DIR + PRIMER_FILE
from scipy.io.wavfile import read as read_audio
from scipy.io.wavfile import write as write_audio
primer_sample_rate, audio = read_audio(PATH_TO_PRIMER)
# Setup the session.
spectral_ops.reset_crepe()
# Compute features.
start_time = time.time()
audio_features = metrics.compute_audio_features(audio)
audio_features['loudness_dbM'] = audio_features['loudness_dbM'].astype(np.float32)
audio_features['loudness_dbL'] = audio_features['loudness_dbL'].astype(np.float32)
audio_features['loudness_dbR'] = audio_features['loudness_dbR'].astype(np.float32)
audio_features_mod = None
print('Audio features took %.1f seconds' % (time.time() - start_time))
```
## **Step 12**--Load most recent checkpoint
```
# Copy most recent checkpoint to "ckpt" folder
%cd $DRIVE_DIR/ckpt/
!rm *
CHECKPOINT_ZIP = 'ckpt.zip'
latest_checkpoint_fname = os.path.basename(tf.train.latest_checkpoint(SAVE_DIR)) + '*'
!cd "$SAVE_DIR"
!cd "$SAVE_DIR" && zip $CHECKPOINT_ZIP $latest_checkpoint_fname* operative_config-0.gin dataset_statistics.pkl
!cp "$SAVE_DIR/$CHECKPOINT_ZIP" "$DRIVE_DIR/ckpt/"
!unzip -o "$CHECKPOINT_ZIP"
!rm "$CHECKPOINT_ZIP"
%cd $SAVE_DIR
!rm "$CHECKPOINT_ZIP"
model_dir = DRIVE_DIR + '/ckpt/'
gin_file = os.path.join(model_dir, 'operative_config-0.gin')
# Load the dataset statistics.
DATASET_STATS = None
dataset_stats_file = os.path.join(model_dir, 'dataset_statistics.pkl')
print(f'Loading dataset statistics from {dataset_stats_file}')
try:
if tf.io.gfile.exists(dataset_stats_file):
with tf.io.gfile.GFile(dataset_stats_file, 'rb') as f:
DATASET_STATS = pickle.load(f)
except Exception as err:
print('Loading dataset statistics from pickle failed: {}.'.format(err))
# Parse gin config,
with gin.unlock_config():
gin.parse_config_file(gin_file, skip_unknown=True)
# Assumes only one checkpoint in the folder, 'ckpt-[iter]`.
ckpt_files = [f for f in tf.io.gfile.listdir(model_dir) if 'ckpt' in f]
ckpt_name = ckpt_files[0].split('.')[0]
ckpt = os.path.join(model_dir, ckpt_name)
# Ensure dimensions and sampling rates are equal
time_steps_train = gin.query_parameter('DefaultPreprocessor.time_steps')
n_samples_train = RENDER_SAMPLES
hop_size = int(n_samples_train / time_steps_train)
time_steps = int(audio_features['audioL'].shape[1] / hop_size)
n_samples = time_steps * hop_size
# Trim all input vectors to correct lengths
for key in ['f0_hzM', 'f0_hzL', 'f0_hzR', 'f0_confidenceM', 'f0_confidenceL', 'f0_confidenceR']:
audio_features[key] = audio_features[key][:time_steps]
for key in ['loudness_dbM', 'loudness_dbL', 'loudness_dbR']:
audio_features[key] = audio_features[key][:, :time_steps]
audio_features['audioM'] = audio_features['audioM'][:, :n_samples]
audio_features['audioL'] = audio_features['audioL'][:, :n_samples]
audio_features['audioR'] = audio_features['audioR'][:, :n_samples]
# Set up the model just to predict audio given new conditioning
model = models.Autoencoder()
model.restore(ckpt)
# Build model by running a batch through it.
start_time = time.time()
_ = model(audio_features, training=False)
print('Restoring model took %.1f seconds' % (time.time() - start_time))
```
## **Step 13**--Modify Conditioning (optional)
These models were not explicitly trained to perform timbre transfer, so they may sound unnatural if the incoming loudness and frequencies are very different then the training data (which will always be somewhat true).
```
#@markdown ## Note Detection
#@markdown You can leave this at 1.0 for most cases
threshold = 1 #@param {type:"slider", min: 0.0, max:2.0, step:0.01}
#@markdown ## Automatic
ADJUST = True #@param{type:"boolean"}
#@markdown Quiet parts without notes detected (dB)
quiet = 30 #@param {type:"slider", min: 0, max:60, step:1}
#@markdown Force pitch to nearest note (amount)
autotune = 0 #@param {type:"slider", min: 0.0, max:1.0, step:0.1}
#@markdown ## Manual
#@markdown Shift the pitch (octaves)
pitch_shift = 0 #@param {type:"slider", min:-2, max:2, step:1}
#@markdown Adjsut the overall loudness (dB)
loudness_shift = 0 #@param {type:"slider", min:-20, max:20, step:1}
audio_features_mod = {k: v.copy() for k, v in audio_features.items()}
## Helper functions.
def shift_ld(audio_features, ld_shiftL=0.0, ld_shiftR=0.0):
"""Shift loudness by a number of ocatves."""
audio_features['loudness_dbL'] += ld_shiftL
audio_features['loudness_dbR'] += ld_shiftR
return audio_features
def shift_f0(audio_features, pitch_shiftL=0.0, pitch_shiftR=0.0):
"""Shift f0 by a number of ocatves."""
audio_features['f0_hzL'] *= 2.0 ** (pitch_shiftL)
audio_features['f0_hzL'] = np.clip(audio_features['f0_hzL'],
0.0,
librosa.midi_to_hz(110.0))
audio_features['f0_hzR'] *= 2.0 ** (pitch_shiftR)
audio_features['f0_hzR'] = np.clip(audio_features['f0_hzR'],
0.0,
librosa.midi_to_hz(110.0))
return audio_features
mask_on = None
if ADJUST and DATASET_STATS is not None:
# Detect sections that are "on".
mask_onL, note_on_valueL = detect_notes(audio_features['loudness_dbL'],
audio_features['f0_confidenceL'],
threshold)
mask_onR, note_on_valueR = detect_notes(audio_features['loudness_dbR'],
audio_features['f0_confidenceR'],
threshold)
if np.any(mask_onL):
# Shift the pitch register.
target_mean_pitchL = DATASET_STATS['mean_pitchL']
target_mean_pitchR = DATASET_STATS['mean_pitchR']
pitchL = core.hz_to_midi(audio_features['f0_hzL'])
pitchR = core.hz_to_midi(audio_features['f0_hzR'])
pitchL = np.expand_dims(pitchL, axis=0)
pitchR = np.expand_dims(pitchR, axis=0)
mean_pitchL = np.mean(pitchL[mask_onL])
mean_pitchR = np.mean(pitchR[mask_onR])
p_diffL = target_mean_pitchL - mean_pitchL
p_diffR = target_mean_pitchR - mean_pitchR
p_diff_octaveL = p_diffL / 12.0
p_diff_octaveR = p_diffR / 12.0
round_fnL = np.floor if p_diff_octaveL > 1.5 else np.ceil
round_fnR = np.floor if p_diff_octaveR > 1.5 else np.ceil
p_diff_octaveL = round_fnL(p_diff_octaveL)
p_diff_octaveR = round_fnR(p_diff_octaveR)
audio_features_mod = shift_f0(audio_features_mod, p_diff_octaveL, p_diff_octaveR)
# Quantile shift the note_on parts.
_, loudness_normL = colab_utils.fit_quantile_transform(
audio_features['loudness_dbL'],
mask_onL,
inv_quantile=DATASET_STATS['quantile_transformL'])
# Quantile shift the note_on parts.
_, loudness_normR = colab_utils.fit_quantile_transform(
audio_features['loudness_dbR'],
mask_onR,
inv_quantile=DATASET_STATS['quantile_transformR'])
# Turn down the note_off parts.
mask_offL = np.logical_not(mask_onL)
mask_offR = np.logical_not(mask_onR)
loudness_normL = np.squeeze(loudness_normL)
loudness_normR = np.squeeze(loudness_normR)
loudness_normL[np.squeeze(mask_offL)] -= quiet * (1.0 - note_on_valueL[mask_offL])
loudness_normR[np.squeeze(mask_offR)] -= quiet * (1.0 - note_on_valueR[mask_offR])
loudness_normL = np.reshape(loudness_normL, audio_features['loudness_dbL'].shape)
loudness_normR = np.reshape(loudness_normR, audio_features['loudness_dbR'].shape)
audio_features_mod['loudness_dbL'] = loudness_normL
audio_features_mod['loudness_dbR'] = loudness_normR
# Auto-tune.
if autotune:
f0_midiL = np.array(core.hz_to_midi(audio_features_mod['f0_hzL']))
f0_midiR = np.array(core.hz_to_midi(audio_features_mod['f0_hzR']))
tuning_factorL = get_tuning_factor(f0_midiL, audio_features_mod['f0_confidenceL'], np.squeeze(mask_onL))
tuning_factorR = get_tuning_factor(f0_midiR, audio_features_mod['f0_confidenceR'], np.squeeze(mask_onR))
f0_midi_atL = auto_tune(f0_midiL, tuning_factorL, np.squeeze(mask_onL), amount=autotune)
f0_midi_atR = auto_tune(f0_midiR, tuning_factorR, np.squeeze(mask_onR), amount=autotune)
audio_features_mod['f0_hzL'] = core.midi_to_hz(f0_midi_atL)
audio_features_mod['f0_hzR'] = core.midi_to_hz(f0_midi_atR)
else:
print('\nSkipping auto-adjust (no notes detected or ADJUST box empty).')
else:
print('\nSkipping auto-adujst (box not checked or no dataset statistics found).')
# Manual Shifts.
audio_features_mod = shift_ld(audio_features_mod, loudness_shift, loudness_shift)
audio_features_mod = shift_f0(audio_features_mod, pitch_shift, pitch_shift)
TRIM = -15
# Plot Features.
has_maskL = int(mask_onL is not None)
n_plots = 3 if has_maskL else 2
figL, axesL = plt.subplots(nrows=n_plots,
ncols=1,
sharex=True,
figsize=(2*n_plots, 8))
if has_maskL:
ax = axesL[0]
ax.plot(np.ones_like(np.squeeze(mask_onL)[:TRIM]) * threshold, 'k:')
ax.plot(np.squeeze(note_on_valueL)[:TRIM])
ax.plot(np.squeeze(mask_onL)[:TRIM])
ax.set_ylabel('Note-on Mask--Left')
ax.set_xlabel('Time step [frame]--Left')
ax.legend(['Threshold', 'Likelihood','Mask'])
ax = axesL[0 + has_maskL]
ax.plot(np.squeeze(audio_features['loudness_dbL'])[:TRIM])
ax.plot(np.squeeze(audio_features_mod['loudness_dbL'])[:TRIM])
ax.set_ylabel('loudness_db--Left')
ax.legend(['Original','Adjusted'])
ax = axesL[1 + has_maskL]
ax.plot(librosa.hz_to_midi(np.squeeze(audio_features['f0_hzL'])[:TRIM]))
ax.plot(librosa.hz_to_midi(np.squeeze(audio_features_mod['f0_hzL'])[:TRIM]))
ax.set_ylabel('f0 [midi]--Left')
_ = ax.legend(['Original','Adjusted'])
has_maskR = int(mask_onR is not None)
n_plots = 3 if has_maskR else 2
figR, axesR = plt.subplots(nrows=n_plots,
ncols=1,
sharex=True,
figsize=(2*n_plots, 8))
if has_maskR:
ax = axesR[0]
ax.plot(np.ones_like(np.squeeze(mask_onR)[:TRIM]) * threshold, 'k:')
ax.plot(np.squeeze(note_on_valueR)[:TRIM])
ax.plot(np.squeeze(mask_onR)[:TRIM])
ax.set_ylabel('Note-on Mask--Right')
ax.set_xlabel('Time step [frame]--Right')
ax.legend(['Threshold', 'Likelihood','Mask'])
ax = axesR[0 + has_maskR]
ax.plot(np.squeeze(audio_features['loudness_dbR'])[:TRIM])
ax.plot(np.squeeze(audio_features_mod['loudness_dbR'])[:TRIM])
ax.set_ylabel('loudness_db--Right')
ax.legend(['Original','Adjusted'])
ax = axesR[1 + has_maskR]
ax.plot(librosa.hz_to_midi(np.squeeze(audio_features['f0_hzR'])[:TRIM]))
ax.plot(librosa.hz_to_midi(np.squeeze(audio_features_mod['f0_hzR'])[:TRIM]))
ax.set_ylabel('f0 [midi]--Right')
_ = ax.legend(['Original','Adjusted'])
```
## **Step 14**--Render audio
After running this cell, your final rendered file should be downloaded automatically. If not, look for it in the "audio_output/normalized" directory inside the directory you set as DRIVE_DIR. There are also unnormalized stereo and mono files in the "audio_output" directory.
```
%cd $AUDIO_OUTPUT_DIR
!mkdir -p normalized
!rm normalized/*
af = audio_features if audio_features_mod is None else audio_features_mod
# Run a batch of predictions.
start_time = time.time()
audio_genM, audio_genL, audio_genR = model(af, training=False)
print('Prediction took %.1f seconds' % (time.time() - start_time))
audio_genL = np.expand_dims(np.squeeze(audio_genL.numpy()), axis=1)
audio_genR = np.expand_dims(np.squeeze(audio_genR.numpy()), axis=1)
audio_genS = np.concatenate((audio_genL, audio_genR), axis=1)
audio_genM = np.expand_dims(np.squeeze(audio_genM.numpy()), axis=1)
write_audio("renderS.wav", 48000, audio_genS)
write_audio("renderM.wav", 48000, audio_genM)
!ffmpeg-normalize renderS.wav -o normalized/render.wav -t -15 -ar 48000
colab_utils.download("normalized/render.wav")
```
## **Step 15** (optional)--Download your model for later use
```
%cd $CKPT_OUTPUT_DIR
!zip -r checkpoint.zip *
colab_utils.download('checkpoint.zip')
!rm checkpoint.zip
```
| github_jupyter |
<a href="https://colab.research.google.com/github/ryanleeallred/DS-Unit-1-Sprint-3-Linear-Algebra/blob/master/module1-vectors-and-matrices/LS_DS_131_Vectors_and_Matrices.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
Lambda School Data Science
*Unit 1, Sprint 3, Module 1*
---
# Vectors and Matrices
- Student can illustrate why we care about linear algebra in the scope of data science.
- Student can graph vectors, identify their dimensionality, calculate their length (norm), and take the dot product of two vectors.
- Student can identify the dimensionality of matrices, multiply them, identify when matrix multiplication is a legal operation, and transpose a matrix.
- Student can identify special types of square matrices including the identity matrix, as well as find the determinant and inverse of a matrix.
- Student can use NumPy to perform basic Linear Algebra operations with Python.
# [Why Linear Algebra?](#why-linear-algebra)
Student can illustrate why we care about linear algebra in the scope of data science.
## Overview
Data Science, Machine Learning, and Artificial intelligence is all about getting computers to do things for us better, cheaper, and faster than we could do them ourselves.
How do we do that? Computers are good at doing small repetitive tasks (like arithmetic). if we tell them what small repetitive tasks to do in the right order then sometimes all of those combined behaviors will result in something that looks like a human's behavior (or at least the decisions/output look like something a human might decide to do/create).
<img alt="Le Comte de Belamy - GAN Art" src="https://obvious-art.com/assets/img/comtedorures.jpg" width='300'>
[Le Comte de Belamy](https://obvious-art.com/le-comte-de-belamy.htm)
The set of instructions that we give to a computer to complete certain tasks is called an **algorithm**. The better that we can organize the set of instructions, the faster that computers can do them. The method that we use to organize and store our set of instructions so that the computer can do them super fast is called a **data structure**. The practice of optimizing the organization of our data structures so that they run really fast and efficiently is called **computer science**. (This is why we will have a unit dedicated solely to computer science in a few months). Data Scientists should care how fast computers can process their sets of instructions (algorithms).
## Follow Along
Here's a simple data structure, in Python it's known as a **list**. It's one of the simplest ways that we can store things (data) and maintain their order. When giving instructions to a computer, it's important that the computer knows in what order to execute them.
```
selfDrivingCarInstructions = ["open door",
"sit on seat",
"put key in ignition",
"turn key to the right until it stops",
"push brake pedal",
"change gear to 'Drive'",
"release brake pedal",
"push gas pedal",
'''turn wheel to navigate streets with thousands of small rules and
exeptions to rules all while avoiding collision with other
objects/humans/cars, obeying traffic laws, not running out of fuel and
getting there in a timely manner''',
"close door"]
# We'll have self-driving cars next week for sure. NBD
```
# Maintaining the order of our sets of ordered instruction-sets
Here's another data structure we can make by putting lists inside of lists, this is called a two-dimensional list. Sometimes it is also known as a two-dimensional array or --if you put some extra methods on it-- a dataframe. As you can see things are starting to get a little bit more complicated.
```
holdMyData = [
[1,2,3],
[4,5,6],
[7,8,9]
]
# Disregard the quality of these bad instructions
```
## Linear Algebra - organize and execute big calculations/operations really fast
So why linear algebra? Because the mathematical principles behinds **vectors** and **matrices** (lists and 2D lists) will help us understand how we can tell computers how to do an insane number of calculations in a very short amount of time.
Remember when we said that computers are really good at doing small and repetitive tasks very quickly?
## I Give You... Matrix Multiplication:
<img src="https://2000thingswpf.files.wordpress.com/2013/04/794-002.png?w=630" width="400">
If you mess up any of those multiplications or additions you're up a creek.
## I Give You... Finding the Determinant of a Matrix: (an introductory linear algebra topic)
## 2x2 Matrix
<img src="http://cdn.virtualnerd.com/tutorials/Alg2_04_01_0017/assets/Alg2_04_01_0017_D_01_16.png" width="400">
Just use the formula!</center>
## 3x3 Matrix
<img src="https://www.thecalculator.co/includes/forms/assets/img/Matrix%20determinant%203x3%20formula.jpg" width='400'>
Just calculate the determinant of 3 different 2x2 matrices and multiply them by 3 other numbers and add it all up.
## 4x4 Matrix
<img src="https://semath.info/img/inverse_cofactor_ex4_02.jpg" width='400'>
Just calculate 3 diferent 3x3 matrix determinants which will require the calculating of 9 different 2x2 matrix determinants, multiply them all by the right numbers and add them all up. And if you mess up any of those multiplications or additions you're up a creek.
## 5x5 Matrix!
## ...
## ...
Just kidding, any linear algebra professor who assigns the hand calculation of a 5x5 matrix determinant (or larger) is a sadist. This is what computers were invented for! Why risk so much hand calculation in order to do something that computers **never** make a mistake at?
By the way, when was the last time that you worked with a dataframe that was 4 rows x 4 columns or smaller?
Quick, find the determinant of this 42837x42837 dataframe by hand!
### Common Applications of Linear Algebra in Data Science:
### Vectors: Rows, Columns, lists, arrays
### Matrices: tables, spreadsheets, dataframes
### Linear Regression: (You might remember from the intro course)
<img src="https://jeffycyang.github.io/content/images/2015/11/Screen-Shot-2015-11-28-at-7-35-39-PM.png" width="400">
```
# Linear Regression Example
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
# Read CSV
df = pd.read_csv('https://raw.githubusercontent.com/ryanleeallred/datasets/master/Ice_Cream_Sales.csv')
# Create Column of 1s
df['Ones'] = np.ones(11)
# Format X and Y Matrices
X = df[['Ones', 'Farenheit']].values
Y = df['Dollars'].values.reshape(-1, 1)
# Calculate Beta Values
beta = np.matmul(np.linalg.inv(np.matmul(np.transpose(X), X)), np.matmul(np.transpose(X), Y))
print(beta)
# Assign Beta Values to Variables
beta_0 = beta[0,0]
beta_1 = beta[1,0]
# Plot points with line of best fit
plt.scatter(df['Farenheit'], df['Dollars'])
axes = plt.gca()
x_vals = np.array(axes.get_xlim())
y_vals = beta_0 + beta_1 * x_vals
plt.plot(x_vals, y_vals, '-', color='b')
plt.title('Ice Cream Sales Regression Line')
plt.xlabel('Farenheit')
plt.ylabel('Dollars')
plt.show()
```
### Dimensionality Reduction Techniques: Principle Component Analysis (PCA) and Singular Value Decomposition (SVD)
Take a giant dataset and distill it down to its important parts. (typically as a pre-processing step for creating visualizations or putting into other models.)
<img src="https://machine-learning-course.readthedocs.io/en/latest/_images/pca4.png" width="400">
### Deep Learning: Convolutional Neural Networks, (Image Recognition)
"Convolving" is the process of passing a filter/kernel (small matrix) over the pixels of an image, multiplying them together, and using the result to create a new matrix. The resulting matrix will be a new image that has been modified by the filter to emphasize certain qualities of an image. This is entirely a linear algebra-based process. A convolutional neural network learns the filters that help it best identify certain aspects of images and thereby classify immages more accurately.
<img src="https://hackernoon.com/hn-images/1*ZCjPUFrB6eHPRi4eyP6aaA.gif">
```
!pip install imageio
# Convolution in action
import imageio
import matplotlib.pyplot as plt
import numpy as np
import scipy.ndimage as nd
from skimage.exposure import rescale_intensity
from skimage import color
img = imageio.imread('https://www.dropbox.com/s/dv3vtiqy439pzag/all_the_things.png?raw=1')
plt.axis('off')
plt.imshow(img);
# Convert I to grayscale, so it will be MxNx1 instead of MxNx4
grayscale = color.rgb2gray(img)
plt.axis('off')
plt.imshow(grayscale, cmap=plt.cm.gray);
laplacian = np.array([[0,0,1,0,0],
[0,0,2,0,0],
[1,2,-16,2,1],
[0,0,2,0,0],
[0,0,1,0,0]])
laplacian_image = nd.convolve(grayscale, laplacian)
plt.axis('off')
plt.imshow(laplacian_image, cmap=plt.cm.gray);
sobel_x = np.array([
[-1,0,1],
[-2,0,2],
[-1,0,1]
])
sobel_x_image = nd.convolve(grayscale, sobel_x)
plt.axis('off')
plt.imshow(sobel_x_image, cmap=plt.cm.gray);
sobel_y = np.array([
[1,2,1],
[0,0,0],
[-1,-2,-1]
])
sobel_y_image = nd.convolve(grayscale, sobel_y)
plt.axis('off')
plt.imshow(sobel_y_image, cmap=plt.cm.gray);
```
## Are we going to learn to do Linear Algebra by hand?
Let me quote your seventh grade math teacher:
> "ArE yOu GoInG To CaRrY a CaLcUlAtOr ArOuNd wItH yOu EvErYwHeRe YoU gO???"
Of course you're going to carry a calculator around everywhere, so mostly **NO**, we're not going to do a lot of hand calculating. We're going to try and refrain from calculating things by hand unless it is absolutely necessary in order to understand and implement the concepts.
We're not trying to re-invent the wheel.
We're learning how to **use** the wheel.
## Challenge
I hope that this teaser helps you be excited about how studying Linear Algebra will help us build the intuition and skills to unlock some of the most exciting and valuable tools and techniques in machine learning. Over the next two modules we are going start building up to the concept of PCA which as you'll see will require a firm understanding of many important linear algebra intuitions.
# [Salars and Vectors](#scalars-and-vectors)
Student can graph vectors, identify their dimensionality, calculate their length (norm), and take the dot product of two vectors.
## Overview
Scalars and vectors are the basic building blocks of linear algebra. Scalars are just numbers stored as a variable, and when vectors or matrices are multiplied by them they will be scaled according to the size of the number. Vectors are a lot like lists of values in Python or like rows/columns in a dataframe many of the principles and intuitions that we learn here can be tied back to our datasets. Understanding vectors is important because matrices can be thought of as a set of vectors just like a dataframe can be thought of as a group of columns or rows.
## Follow Along
### Scalars
A single number. Variables representing scalars are typically written in lower case.
Scalars can be whole numbers or decimals.
\begin{align}
a = 2
\qquad
b = 4.815162342
\end{align}
They can be positive, negative, 0 or any other real number.
\begin{align}
c = -6.022\mathrm{e}{+23}
\qquad
d = \pi
\end{align}
```
import math
import matplotlib.pyplot as plt
import numpy as np
# Start with a simple vector
blue = [.5, .5]
# Then multiply it by a scalar
green = np.multiply(2, blue)
red = np.multiply(math.pi, blue)
orange = np.multiply(-0.5, blue)
# Plot the Scaled Vectors
plt.arrow(0,0, red[0], red[1],head_width=.05, head_length=0.05, color ='red')
plt.arrow(0,0, green[0], green[1],head_width=.05, head_length=0.05, color ='green')
plt.arrow(0,0, blue[0], blue[1],head_width=.05, head_length=0.05, color ='blue')
plt.arrow(0,0, orange[0], orange[1],head_width=.05, head_length=0.05, color ='orange')
plt.xlim(-1,2)
plt.ylim(-1,2)
plt.title("Scaled Vectors")
plt.show()
```
### Vectors:
A vector of dimension *n* is an **ordered** collection of *n* elements, which are called **components** (Note, the components of a vector are **not** referred to as "scalars"). Vector notation variables are commonly written as a bold-faced lowercase letters or italicized non-bold-faced lowercase characters with an arrow (→) above the letters:
Written: $\vec{v}$
Examples:
\begin{align}
\vec{a} =
\begin{bmatrix}
1\\
2
\end{bmatrix}
\qquad
\vec{b} =
\begin{bmatrix}
-1\\
0\\
2
\end{bmatrix}
\qquad
\vec{c} =
\begin{bmatrix}
4.5
\end{bmatrix}
\qquad
\vec{d} =
\begin{bmatrix}
Pl\\
a\\
b\\
\frac{2}{3}
\end{bmatrix}
\end{align}
The above vectors have dimensions 2, 3, 1, and 4 respectively.
Why do the vectors below only have two components?
```
# Vector Examples
yellow = [.5, .5]
red = [.2, .1]
blue = [.1, .3]
# Coordinate Pairs for where the arrowheads are
# yellow = (.5, .5)
# red = (.2, .1)
# blue = (.1, .3)
plt.arrow(0, 0, .5, .5, head_width=.02, head_length=0.01, color = 'y')
plt.arrow(0, 0, .2, .1, head_width=.02, head_length=0.01, color = 'r')
plt.arrow(0, 0, .1, .3, head_width=.02, head_length=0.01, color = 'b')
plt.title('Vector Examples')
plt.show()
```
In domains such as physics it is emphasized that vectors have two properties: direction and magnitude. It's rare that we talk about them in that sense in Data Science unless we're specifically in a physics context. We just note that the length of the vector is equal to the number of dimensions of the vector.
What happens if we add a third component to each of our vectors?
```
from mpl_toolkits.mplot3d import Axes3D
import numpy as np
yellow = [.5, .5, .5]
red = [.2, .1, .0]
blue = [.1, .3, .3]
vectors = np.array([[0, 0, 0, .5, .5, .5],
[0, 0, 0, .2, .1, .0],
[0, 0, 0, .1, .3, .3]])
X, Y, Z, U, V, W = zip(*vectors)
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
ax.quiver(X, Y, Z, U, V, W, length=1)
ax.set_xlim([0, 1])
ax.set_ylim([0, 1])
ax.set_zlim([0, 1])
ax.set_xlabel('X')
ax.set_ylabel('Y')
ax.set_zlabel('Z')
plt.show()
```
### Norm of a Vector (Magnitude or length)
The *Norm* or *Magnitude* of a vector is nothing more than the **length** of the vector. Since a vector is just a line (essentially) if you treat it as the hypotenuse of a triangle you could use the pythagorean theorem to find the equation for the norm of a vector. We're essentially just generalizing the equation for the hypotenuse of a triangle that results from the pythagorean theorem to n dimensional space.
We denote the norm of a vector by wrapping it in double pipes (like double absolute value signs)
\begin{align}
||v|| =
\sqrt{v_{1}^2 + v_{2}^2 + \ldots + v_{n}^2}
\\
\vec{a} =
\begin{bmatrix}
3 & 7 & 2 & 4
\end{bmatrix}
\\
||a|| = \sqrt{3^2 + 7^2 + 2^2 + 4^2} \\
||a|| = \sqrt{9 + 49 + 4 + 16} \\
||a|| = \sqrt{78}
\end{align}
The Norm is the square root of the sum of the squared elements of a vector.
Properties of the Norm:
The norm is always positive or zero $||x|| \geq 0$
The norm is only equal to zero if all of the elements of the vector are zero.
The Triangle Inequality: $|| x + y ||\leq ||x|| + ||y||$
### The Equation for the norm of a vector is just Pythagorean Theorem extended to more than two dimensions.
$a^2 + b^2 = c^2$
a = one side of a right triangle
b = the other side
c = the "hypotenuse" of a right triangle (We consider the vector that we're looking at to be the hypotenuse of a triangle
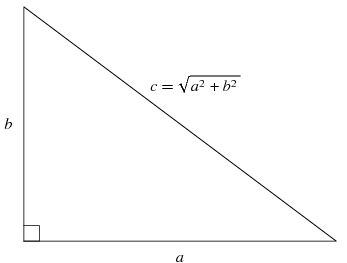
$c = \sqrt{a^2 + b^2}$
### Dot Product
The dot product of two vectors $\vec{a}$ and $\vec{b}$ is a scalar quantity that is equal to the sum of pair-wise products of the components of vectors a and b. An example will make this make much more sense:
\begin{align} \vec{a} \cdot \vec{b} = (a_{1} \times b_{1}) + (a_{2} \times b_{2}) + \ldots + ( a_{n} \times b_{n}) \end{align}
Example:
\begin{align}
\vec{a} =
\begin{bmatrix}
3 & 7 & 2 & 4
\end{bmatrix}
\qquad
\vec{b} =
\begin{bmatrix}
4 & 1 & 12 & 6
\end{bmatrix}
\end{align}
The dot product of two vectors would be:
\begin{align}
a \cdot b = (3)(4) + (7)(1) + (2)(12) + (4)(6) \\
= 12 + 7 + 24 + 24 \\
= 67
\end{align}
The dot product is commutative: $ a \cdot b = b \cdot a$
The dot product is distributive: $a \cdot (b + c) = a \cdot b + a \cdot c$
Two vectors must have the same number of components in order for the dot product to exist. If they lengths differ the dot product is undefined.
## Challenge
Being able to calculate the length of a vector is vital as a vector's length is one of the key attributes that defines it. Likewise taking the dot product of two vectors is a process that you **need** to be comfortable with as it is involved in many more complex linear algebra processes. You will practice both of these concepts as well as graphing vectors on your assignment this afternoon.
# [Matrices](#matrices)
Student can identify the dimensionality of matrices, multiply them, identify when matrix multiplication is a legal operation, and transpose a matrix.
A **matrix** is a rectangular grid of numbers arranged in rows and columns. Variables that represent matrices are typically written as capital letters (boldfaced as well if you want to be super formal).
\begin{align}
A =
\begin{bmatrix}
1 & 2 & 3\\
4 & 5 & 6\\
7 & 8 & 9
\end{bmatrix}
\qquad
B = \begin{bmatrix}
1 & 2 & 3\\
4 & 5 & 6
\end{bmatrix}
\end{align}
You'll notice that we arrange our dataframes in a similar grid-like pattern meaning that anything we learn about matrices also applies to the dataframes that we work with in Pandas
## Overview
A **matrix** is a rectangular grid of numbers arranged in rows and columns. Variables that represent matrices are typically written as capital letters (boldfaced as well if you want to be super formal).
\begin{align}
A =
\begin{bmatrix}
1 & 2 & 3\\
4 & 5 & 6\\
7 & 8 & 9
\end{bmatrix}
\qquad
B = \begin{bmatrix}
1 & 2 & 3\\
4 & 5 & 6
\end{bmatrix}
\end{align}
You'll notice that we arrange our dataframes in a similar grid-like pattern meaning that anything we learn about matrices also applies to the dataframes that we work with in Pandas
```
import pandas as pd
df = pd.DataFrame({'a': [1,2,3], 'b': [4,5,6]})
print(df.shape)
df.head()
```
## Follow Along
### Dimensionality
The number of rows and columns that a matrix has is called its **dimension**.
When listing the dimension of a matrix we always list rows first and then columns.
The dimension of matrix A is 3x3. (Note: This is read "Three by Three", the 'x' isn't a multiplication sign.)
What is the Dimension of Matrix B?
### Matrix Equality
In order for two Matrices to be equal the following conditions must be true:
1) They must have the same dimensions.
2) Corresponding elements must be equal.
\begin{align}
\begin{bmatrix}
1 & 4\\
2 & 5\\
3 & 6
\end{bmatrix}
\neq
\begin{bmatrix}
1 & 2 & 3\\
4 & 5 & 6
\end{bmatrix}
\end{align}
### Matrix Multiplication
You can multipy any two matrices where the number of columns of the first matrix is equal to the number of rows of the second matrix.
The unused dimensions of the factor matrices tell you what the dimensions of the product matrix will be.
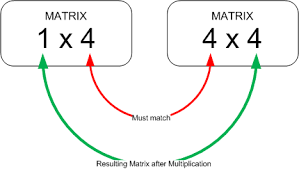
There is no commutative property of matrix multiplication (you can't switch the order of the matrices and always get the same result).
Matrix multiplication is best understood in terms of the dot product. Remember:
\begin{align} \vec{a} \cdot \vec{b} = (a_{1} \times b_{1}) + (a_{2} \times b_{2}) + \ldots + ( a_{n} \times b_{n}) \end{align}
To multiply to matrices together, we will take the dot product of each row of the first matrix with each column of the second matrix. The position of the resulting entries will correspond to the row number and column number of the row and column vector that were used to find that scalar. Lets look at an example to make this more clear.

\begin{align}
\begin{bmatrix}
1 & 2 & 3 \\
4 & 5 & 6
\end{bmatrix}
\begin{bmatrix}
7 & 8 \\
9 & 10 \\
11 & 12
\end{bmatrix}
=
\begin{bmatrix}
(1)(7)+(2)(9)+(3)(11) & (1)(8)+(2)(10)+(3)(12)\\
(4)(7)+(5)(9)+(6)(11) & (4)(8)+(5)(10)+(6)(12)
\end{bmatrix}
=
\begin{bmatrix}
(7)+(18)+(33) & (8)+(20)+(36)\\
(28)+(45)+(66) & (32)+(50)+(72)
\end{bmatrix}
=
\begin{bmatrix}
58 & 64\\
139 & 154
\end{bmatrix}
\end{align}
## Transpose
A transposed matrix is one whose rows are the columns of the original and whose columns are the rows of the original.
Common notation for the transpose of a matrix is to have a capital $T$ superscript or a tick mark:
\begin{align}
B^{T}
\qquad
B^{\prime}
\end{align}
The first is read "B transpose" the second is sometimes read as "B prime" but can also be read as "B transpose".
The transpose of any matrix can be found easily by fixing the elements on the main diagonal and flipping the placement of all other elements across that diagonal.

\begin{align}
B =
\begin{bmatrix}
1 & 2 & 3 \\
4 & 5 & 6
\end{bmatrix}
\qquad
B^{T} =
\begin{bmatrix}
1 & 4 \\
2 & 5 \\
3 & 6
\end{bmatrix}
\end{align}
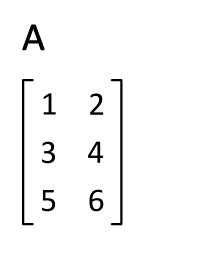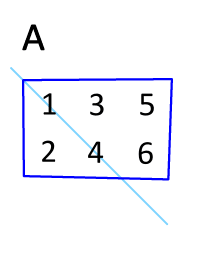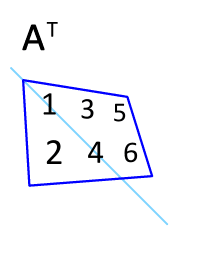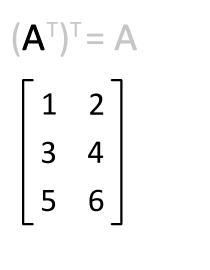
```
df
# Transposing a dataframe switches its rows and columns
df.T
```
## Challenge
Throughout the week we will be learning how the princples of mathematical matrices apply to our datasets. We'll be focusing mostly on math topics and techniques but as the week goes on we will be making more and more connections back to how we think about and work with our datasets.
# [Square Matrices](#square-matrices)
Student can identify special types of square matrices including the identity matrix, as well as find the determinant and inverse of a matrix.
## Overview
In a traditional linear algebra class after the first few weeks you would deal almost exclusively with square matrices. They have very nice properties that their lopsided sisters and brothers just don't possess.
A square matrix is any matrix that has the same number of rows as columns:
\begin{align}
A =
\begin{bmatrix}
a_{1,1}
\end{bmatrix}
\qquad
B =
\begin{bmatrix}
b_{1,1} & b_{1,2} \\
b_{2,1} & b_{2,2}
\end{bmatrix}
\qquad
C =
\begin{bmatrix}
c_{1,1} & c_{1,2} & c_{1,3} \\
c_{2,1} & c_{2,2} & c_{2,3} \\
c_{3,1} & c_{3,2} & c_{3,3}
\end{bmatrix}
\end{align}
## Follow Along
### Special Kinds of Square Matrices
**Diagonal:** Values on the main diagonal, zeroes everywhere else.
\begin{align}
A =
\begin{bmatrix}
a_{1,1} & 0 & 0 \\
0 & a_{2,2} & 0 \\
0 & 0 & a_{3,3}
\end{bmatrix}
\end{align}
**Upper Triangular:** Values on and above the main diagonal, zeroes everywhere else.
\begin{align}
B =
\begin{bmatrix}
b_{1,1} & b_{1,2} & b_{1,3} \\
0 & b_{2,2} & b_{2,3} \\
0 & 0 & b_{3,3}
\end{bmatrix}
\end{align}
**Lower Triangular:** Values on and below the main diagonal, zeroes everywhere else.
\begin{align}
C =
\begin{bmatrix}
c_{1,1} & 0 & 0 \\
c_{2,1} & c_{2,2} & 0 \\
c_{3,1} & c_{3,2} & c_{3,3}
\end{bmatrix}
\end{align}
**Identity Matrix:** A diagonal matrix with ones on the main diagonal and zeroes everywhere else. The product of the any square matrix and the identity matrix is the original square matrix $AI == A$. Also, any matrix multiplied by its inverse will give the identity matrix as its product. $AA^{-1} = I$
\begin{align}
D =
\begin{bmatrix}
1
\end{bmatrix}
\qquad
E =
\begin{bmatrix}
1 & 0 \\
0 & 1
\end{bmatrix}
\qquad
F =
\begin{bmatrix}
1 & 0 & 0 \\
0 & 1 & 0 \\
0 & 0 & 1
\end{bmatrix}
\end{align}
**Symmetric:** The numbers above the main diagonal are mirrored below/across the main diagonal.
\begin{align}
G =
\begin{bmatrix}
1 & 4 & 5 \\
4 & 2 & 6 \\
5 & 6 & 3
\end{bmatrix}
\end{align}
### Determinant
The determinant is a property that all square matrices possess and is denoted $det(A)$ or using pipes (absolute value symbols) $|A|$
The equation given for finding the determinant of a 2x2 matrix is as follows:
\begin{align}
A = \begin{bmatrix}
a & b \\
c & d
\end{bmatrix}
\qquad
|A| = ad-bc
\end{align}
The determinant of larger square matrices recursively by finding the determinats of the smaller matrics that make up the large matrix.
For example:
<center><img src="https://wikimedia.org/api/rest_v1/media/math/render/svg/14f2f2a449d6d152ee71261e47551aa0a31c801e" width=500></center>
The above equation is **very** similar to the equation that we use to find the cross-product of a 3x3 matrix.
### Inverse
There are multiple methods that we could use to find the inverse of a matrix by hand. I would suggest you explore those methods --if this content isn't already overwhelming enough. The inverse is like the reciprocal of the matrix that was used to generate it. Just like $\frac{1}{8}$ is the reciprocal of 8, $A^{-1}$ acts like the reciprocal of $A$. The equation for finding the determinant of a 2x2 matrix is as follows:
\begin{align}
A = \begin{bmatrix}
a & b \\
c & d
\end{bmatrix}
\qquad
A^{-1} = \frac{1}{ad-bc}\begin{bmatrix}
d & -b\\
-c & a
\end{bmatrix}
\end{align}
### What happens if we multiply a matrix by its inverse?
The product of a matrix multiplied by its inverse is the identity matrix of the same dimensions as the original matrix. There is no concept of "matrix division" in linear algebra, but multiplying a matrix by its inverse is very similar since $8\times\frac{1}{8} = 1$.
\begin{align}
A^{-1}A = I
\end{align}
### Not all matrices are invertible
Matrices that are not square are not invertible.
A matrix is invertible if and only if its determinant is non-zero. You'll notice that the fraction on the left side of the matrix is $\frac{1}{det(A)}$.
As you know, dividing anything by 0 leads to an undefined quotient. Therefore, if the determinant of a matrix is 0, then the entire inverse becomes undefined.
### What leads to a 0 determinant?
A square matrix that has a determinant of 0 is known as a "singular" matrix. One thing that can lead to a matrix having a determinant of 0 is if two rows or columns in the matrix are perfectly collinear. Another way of saying this is that the determinant will be zero if the rows or columns of a matrix are not linearly dependent.
One of the most common ways that a matrix can end up having rows that are linearly dependent is if one column a multiple of another column. Lets look at an example:
\begin{align}
C =\begin{bmatrix}
1 & 5 & 2 \\
2 & 7 & 4 \\
3 & 2 & 6
\end{bmatrix}
\end{align}
Look at the columns of the above matrix, column 3 is exactly double column 1. (could be any multiple or fraction) Think about if you had some measure in a dataset of distance in miles, but then you also wanted to convert its units to feet, so you create another column and multiply the mile measure by 5,280 (Thanks Imperial System). But then you forget to drop one of the columns so you end up with two columns that are linearly dependent which causes the determinant of your dataframe to be 0 and will cause certain algorithms to fail. We'll go deeper into this concept next week (this can cause problems with linear regression) so just know that matrices that have columns that are a multiple or fraction of another column will cause the determinant of that matrix to be 0.
## Challenge
Square Matrices have many important properties like the inverse and determinant that we will be using in the future. For more details about the implications of a square matrix having an inverse you can try googling the "Invertible Matrix Theorem" mastering and proving the different partso of this theorem is one of the main focuses of traditional linear algebra education.
# [Doing Linear Algebra with NumPy](#linear-algebra-NumPy)
Student can use NumPy to perform basic Linear Algebra operations with Python.
## Overview
NumPy is at the core of many types of scientific computing and is the main tool that we will use to perform Linear Algebra operations with Python.
We have talked about how vectors are a lot like Python Lists, however we will mostly use NumPy arrays instead of Python lists as the main data structure. Lets demonstrate some of the main differences betwen Python lists and NumPy Arrays.
```
### What will happen when I add the two Python lists together?
# Lets create two 1-Dimensional NumPy arrays and add them together?
# What do you think the result will be?
# Array Broadcasting
# Doing some arithmetic operation to vectors "element-wise"
```
What we have just demonstrated is one of the primary benefits of NumPy and it is called "Array Broadcasting" This means that arithmetic operations happen "element-wise"
```
```
## Follow Along
Lets calculate a dot product. What do we need to do? We need to multiply two vectors "element-wise" and them sum up all of those products
```
# Find the dot product of vectos a and b
# Dot Product
```
We can also use provided linear algebra functions from the [np.linalg](https://docs.scipy.org/doc/numpy-1.15.1/reference/routines.linalg.html) documentation in order to perform Linear Algebra processes.
```
```
### 1-Dimensional vs 2-Dimensional NumPy Arrays
One dimensional NumPy arrays can be used to represent vectors but cannot be used to differentiate between row and column vectors.
\begin{align}
\text{column vector} = \begin{bmatrix}1 \\ 2 \\ 3\end{bmatrix}
\end{align}
\begin{align}
\text{row vector} = \begin{bmatrix} 1 & 2 & 3\end{bmatrix}
\end{align}
Notice that one dimensional arrays do not have a second element listed for their shape. so that there is no way to tell if they represent column vectors or row vectors
```
# Notice how only the number of items in the array is listed
# And if we swap rows and columns the shape does not change
```
### If I explicitly want a row of column vector then I need to use a 2D NumPy Array
```
```
### Only use NumPy Arrans, DO NOT use the NumPy Matrix class.
NumPy has a matrix class which can be used for representing matrices. However, I don't want you to use that datatype because it will be removed from NumPy in the future. However, I want you to be aware that this other datatype exists so that you don't get confused when you look at stack overflow posts or other tutorials that use the matrix datatype. Keep an eye out for resources that use it and just know that they will soon be outdated. Anything that can be done with the matrix class can also be done with simple NumPY arrays.
```
# Do not work with this datatype
# This will be removed from NumPy in the future
# Matrix as a NumPy Array
# ndarrays
```
## Challenge
In your afternoon assignment you will perform many different linear algebra functions using NumPy. Many of which will be done easily by using these [Helpful NumPy Linear Algebra Functions](https://docs.scipy.org/doc/numpy-1.15.1/reference/routines.linalg.html) found in the NumPy documentation.
# Review
Lets good at working with vectors and matrices with NumPy!
Remember that you'll need many functions from [the documentation](https://docs.scipy.org/doc/numpy-1.15.1/reference/routines.linalg.html) in order to complete the assignment
Remember to use 2D NumPy arrays to represent row/column vectors as well as matrices and please share any helpful resources that you find in the cohort channel.
| github_jupyter |
### Practice: Voice Command Recognition

Today you're finally gonna deal with speech! We'll walk you through all the main steps of speech processing pipeline and you'll get to write your own voice recognition system. It's gonna be fun! _(they said)_
```
from IPython.display import display, Audio
import matplotlib.pyplot as plt
%matplotlib inline
import numpy as np
import librosa
!wget https://github.com/yandexdataschool/nlp_course/raw/2019/week10_asr/sample1.wav -O sample1.wav
!wget https://github.com/yandexdataschool/nlp_course/raw/2019/week10_asr/sample2.wav -O sample2.wav
display(Audio("sample1.wav"))
display(Audio("sample2.wav"))
```
Consider an audio sample above. The reason you can hear a sound is because your speakers/headphones cause oscillations of air that reach your ears. There's a part of your inner ear called [Cochlea](https://en.wikipedia.org/wiki/Cochlea) that looks like a spiral where every spot is receptive to a specific range of sound frequency.

The sound itself can be recorded as a collection of __amplitudes__ of these oscillations over time:
```
amplitudes, sample_rate = librosa.core.load("./sample1.wav")
print("Length: {} seconds at sample rate {}".format(amplitudes.shape[0] / sample_rate, sample_rate))
plt.figure(figsize=[16, 4])
plt.title("First 10^4 out of {} amplitudes".format(len(amplitudes)))
plt.plot(amplitudes[:10000]);
```
### Task 1: Mel-Spectrogram (3 points)
As you can see, amplitudes follow a periodic patterns with different frequencies. However, it is very difficult to process these amplitudes directly because there's so many of them! A typical WAV file contains 22050 amplitudes per second, which is already way above a typical sequence length for other NLP applications. Hence, we need to compress this information to something manageable.
A typical solution is to use __spectrogram:__ instead of saving thousands of amplitudes, we can perform Fourier transformation to find which periodics are prevalent at each point in time. More formally, a spectrogram applies [Short-Time Fourier Transform (STFT)](https://en.wikipedia.org/wiki/Short-time_Fourier_transform) to small overlapping windows of the amplitude time-series:
<img src="https://www.researchgate.net/profile/Phillip_Lobel/publication/267827408/figure/fig2/AS:295457826852866@1447454043380/Spectrograms-and-Oscillograms-This-is-an-oscillogram-and-spectrogram-of-the-boatwhistle.png" width="480px">
However, this spectrogram may have extraordinarily large numbers that can break down neural networks. Therefore the standard approach is to convert spectrogram into a __mel-spectrogram__ by changing frequencies to [Mel-frequency spectrum(https://en.wikipedia.org/wiki/Mel-frequency_cepstrum)].
Hence, the algorithm to compute spectrogram of amplitudes $y$ becomes:
1. Compute Short-Time Fourier Transform (STFT): apply fourier transform to overlapping windows
2. Build a spectrogram: $S_{ij} = abs(STFT(y)_{ij}^2)$
3. Convert spectrogram to a Mel basis
By far the hardest pard of this is computing STFT, so let's focus on that first:
1.
```
# Some helpers:
# 1. slice time-series into overlapping windows
def slice_into_frames(amplitudes, window_length, hop_length):
return librosa.core.spectrum.util.frame(
np.pad(amplitudes, int(window_length // 2), mode='reflect'),
frame_length=window_length, hop_length=hop_length)
# output shape: [window_length, num_windows]
dummy_amps = amplitudes[2048: 6144]
dummy_frames = slice_into_frames(dummy_amps, 2048, 512)
plt.figure(figsize=[16, 4])
plt.subplot(121, title='Whole audio sequence', ylim=[-3, 3])
plt.plot(dummy_amps)
plt.subplot(122, title='Overlapping frames', yticks=[])
for i, frame in enumerate(dummy_frames.T):
plt.plot(frame + 10 - i);
# 2. Weights for window transform. Before performing FFT you can scale amplitudes by a set of weights
# The weights we're gonna use are large in the middle of the window and small on the sides
dummy_window_length = 3000
dummy_weights_window = librosa.core.spectrum.get_window('hann', dummy_window_length, fftbins=True)
plt.plot(dummy_weights_window); plt.plot([1500, 1500], [0, 1.1], label='window center'); plt.legend()
# 3. Fast Fourier Transform in Numpy. Note: this function can process several inputs at once (mind the axis!)
dummy_fft = np.fft.rfft(dummy_amps[:3000, None] * dummy_weights_window[:, None], axis=0) # complex[sequence_length, num_sequences]
plt.plot(np.real(dummy_fft)[:, 0])
```
Okay, now it's time to combine everything into a __S__hort-__T__ime __F__ourier __T__ransform
```
def get_STFT(amplitudes, window_length, hop_length):
""" Compute short-time Fourier Transform """
# slice amplitudes into overlapping frames [window_length, num_frames]
frames = <YOUR CODE>
# get weights for fourier transform, float[window_length]
fft_weights = <YOUR CODE>
# apply fourier transfrorm to frames scaled by weights
stft = <YOUR CODE>
return stft
def get_melspectrogram(amplitudes, sample_rate=22050, n_mels=128,
window_length=2048, hop_length=512, fmin=1, fmax=8192):
"""
Implement mel-spectrogram as described above.
:param amplitudes: float [num_amplitudes], time-series of sound amplitude, same as above
:param sample rate: num amplitudes per second
:param n_mels: spectrogram channels
:param window_length: length of a patch to which you apply FFT
:param hop_length: interval between consecutive windows
:param f_min: minimal frequency
:param f_max: maximal frequency
:returns: mel-spetrogram [n_mels, duration]
"""
# Step I: compute Short-Time Fourier Transform
stft = <YOUR CODE>
assert stft.shape == (window_length // 2 + 1, len(amplitudes) // 512 + 1)
# Step II: convert stft to a spectrogram
spectrogram = <YOUR CODE>
# Step III: convert spectrogram into Mel basis (multiplying by transformation matrix)
mel_basis = librosa.filters.mel(sample_rate, n_fft=window_length,
n_mels=n_mels, fmin=fmin, fmax=fmax)
# ^-- matrix [n_mels, window_length / 2 + 1]
mel_spectrogram = <TOUR CODE>
assert mel_spectrogram.shape == (n_mels, len(amplitudes) // 512 + 1)
return mel_spectrogram
amplitudes1, _ = librosa.core.load("./sample1.wav")
amplitudes2, _ = librosa.core.load("./sample2.wav")
ref1 = librosa.feature.melspectrogram(amplitudes1, sr=sample_rate, n_mels=128, fmin=1, fmax=8192)
ref2 = librosa.feature.melspectrogram(amplitudes2, sr=sample_rate, n_mels=128, fmin=1, fmax=8192)
assert np.allclose(get_melspectrogram(amplitudes1), ref1, rtol=1e-4, atol=1e-4)
assert np.allclose(get_melspectrogram(amplitudes2), ref2, rtol=1e-4, atol=1e-4)
plt.figure(figsize=[16, 4])
plt.subplot(1, 2, 1)
plt.title("That's no moon - it's a space station!"); plt.xlabel("Time"); plt.ylabel("Frequency")
plt.imshow(get_melspectrogram(amplitudes1))
plt.subplot(1, 2, 2)
plt.title("Help me, Obi Wan Kenobi. You're my only hope."); plt.xlabel("Time"); plt.ylabel("Frequency")
plt.imshow(get_melspectrogram(amplitudes2));
# note that the second spectrogram has higher mean frequency corresponding to the difference in gender
```
### Speech Commands Dataset
We're now going to train a classifier to recognize voice. More specifically, we'll use the [Speech Commands Dataset] that contains around 30 different words with a few thousand voice records each. Naturally, we'll preprocess each training sample using the `get_melspectrogram` you designed.
```
import os
datadir = "speech_commands"
!wget http://download.tensorflow.org/data/speech_commands_v0.01.tar.gz -O speech_commands_v0.01.tar.gz
# alternative url: https://www.dropbox.com/s/j95n278g48bcbta/speech_commands_v0.01.tar.gz?dl=1
!mkdir {datadir} && tar -C {datadir} -xvzf speech_commands_v0.01.tar.gz 1> log
samples_by_target = {
cls: [os.path.join(datadir, cls, name) for name in os.listdir("./speech_commands/{}".format(cls))]
for cls in os.listdir(datadir)
if os.path.isdir(os.path.join(datadir, cls))
}
print('Classes:', ', '.join(sorted(samples_by_target.keys())[1:]))
```
Let's begin by training a model to recognize direcions. Imagine a mobile robot that you can control via speech. You say "Go right" and it obeys. Or it doesn't - depending on how you trained it :)
```
from sklearn.model_selection import train_test_split
from itertools import chain
from tqdm import tqdm
import joblib as jl
classes = ("left", "right", "up", "down", "stop")
def preprocess_sample(filepath, max_length=150):
amplitudes, sr = librosa.core.load(filepath)
spectrogram = get_melspectrogram(amplitudes, sample_rate=sr)[:, :max_length]
spectrogram = np.pad(spectrogram, [[0, 0], [0, max(0, max_length - spectrogram.shape[1])]], mode='constant')
target = classes.index(filepath.split(os.sep)[-2])
return np.float32(spectrogram), np.int64(target)
all_files = chain(*(samples_by_target[cls] for cls in classes))
spectrograms_and_targets = jl.Parallel(n_jobs=-1)(tqdm(list(map(jl.delayed(preprocess_sample), all_files))))
X, y = map(np.stack, zip(*spectrograms_and_targets))
X = X.transpose([0, 2, 1]) # to [batch, time, channels]
X_train, X_val, y_train, y_val = train_test_split(X, y, random_state=42)
```
__Train a model:__ finally, lets' build and train a classifier neural network. You can use _any_ library you like. If in doubt, consult the model & training tips below.
```
import keras, keras.layers as L
# Build, compile and train a model. If you're out of ideas, see hints below
# Use any framework you want, the stub below is optional
model = keras.models.Sequential([
L.InputLayer(input_shape=(None, 128)), # (batch), time, channels
<YOUR CODE>
])
# Compile and train
<YOUR CODE>
accuracy = np.mean(model.predict(X_val).argmax(-1) == y_val)
print("Final Accuracy:", accuracy)
assert accuracy >= 0.85, "Your model is not good enough. Yet."
print("Well done!")
```
__Training tips:__ here's what you can try:
* __Layers:__ 1d or 2d convolutions, perhaps with some batch normalization in between;
* __Architecture:__ VGG-like, residual, highway, densely-connected - you name it :)
* __Batch size matters:__ smaller batches usually train slower but better. Try to find the one that suits you best.
* __Data augmentation:__ add background noise, faster/slower, change pitch;
* __Average checkpoints:__ you can make model more stable with [this simple technique (arxiv)](https://arxiv.org/abs/1803.05407)
* __For full scale stage:__ make sure you're not losing too much data due to max_length in the pre-processing stage!
These are just recommendations. As long as your model works, you're not required to follow them.
### Main quest: full scale recognition (4+ points)
Your final task is to train a full-scale voice command spotter and apply it to a video:
1. Build the dataset with all 30+ classes (directions, digits, names, etc.)
* __Optional:__ include a special "noise" class that contains random unrelated sounds
* You can download youtube videos with [`youtube-dl`](https://ytdl-org.github.io/youtube-dl/index.html) library.
2. Train a model on this full dataset. Kudos for tuning its accuracy :)
3. Apply it to a audio/video of your choice to spot the occurences of each keyword
* Here's one [video about primes](https://www.youtube.com/watch?v=EK32jo7i5LQ) that you can try. It should be full of numbers :)
* There are multiple ways you can analyze the performance of your network, e.g. plot probabilities predicted for every time-step. Chances are you'll discover something useful about how to improve your model :)
As usual, please briefly describe what you did in a short informal report.
```
<A WHOLE LOT OF YOUR CODE>
```
| github_jupyter |
# Python: The numpy library
**Goal**: manipulate matrices or multidimensional arrays with the numpy package!
## Introduction to numpy
Numpy is one of the fundamental package for scientific computing in Python. The numpy library (http://www.numpy.org/) allows you to perform numerical calculations with Python. It introduces an easy management of arrays of numbers. To use numpy, you must first import the numpy package with the following instruction ``` import numpy ```.
```
import numpy as np
```
## Arrays with numpy
In this section, we will discover the numpy tables. So, the main data structure in numpy is the ``` ndarray ```. Arrays can be created with ``` numpy.array() ```. Square brackets are used to delimit lists of elements in arrays. In summary, a vector represents a list and a matrix represents a list of lists.
```
vector = np.array([1, 2, 3, 4, 5])
print(vector)
matrix = np.array([[1.0, 2.0, 3.0], [4.0, 5.0, 6.0], [7.0, 8.0, 9.0]])
print(matrix)
```
## Size of an array
When manipulating matrices (matrix multiplication for example), it is very useful to know the ``` size of the array ```. Otherwise, you may get errors. So to do this, we have the array property ``` ndarray.shape ```. For vectors, shape will return a tuple with an element. On the other hand, the size of a matrix has two elements, the number of rows and the number of columns.
```
vector.shape
matrix.shape
```
## Read dataset using numpy
To read a dataset directly with the numpy library, we use the numpy ``` genfromtxt() ``` function.
```
world_alcohol = np.genfromtxt("world_alcohol.csv", delimiter=",")
world_alcohol
print(type(world_alcohol))
```
We can clearly see that there is a lot of nan in the result. In reality, they are not nan, we will see later how to correct this problem. In fact, when we talk about matrices, we must have only numbers in the dataset. However numpy can accept many types of data, not only numbers. But the particularity of a numpy array is that all the values of the different columns must have the same data type by default. The default format type used by the genformtext function is float, see the documentation: https://numpy.org/doc/stable/reference/generated/numpy.genfromtxt.html.
## Data types in numpy
Here we will see the main data types in numpy which have slight differences with the Python data types. We will also see the particularity with numpy arrays. The data types accepted by numpy are: ```bool```, ```integer```, ```float``` and ```string```. To know the data type of an array, we use the ```dtype``` method of the numpy ```ndarray object```.
```
vector.dtype
matrix.dtype
world_alcohol.dtype
```
## Display data correctly
We have seen previously that our dataset displays nan on some columns. In this section, we will see how to display the data correctly. As a reminder, ```nan``` means ```not a number```, it is a ```data type``` that is used to say that the ```value is missing```. There is another type of data that is ```na (not available)``` for items that are not available. So you have to know that ```nan``` and ```na``` are data types.
```
world_alcohol
```
To display the correct values, we use the ```dtype=U75``` parameter of the ```genfromtxt``` function. This is to force numpy to read each value in the ```U75 (Unicode 75 bit)``` format.
```
world_alcohol = np.genfromtxt("world_alcohol.csv", delimiter=",", dtype="U75")
world_alcohol
```
We notice that our dataset has a header. To remove it, we use the ```skip_header``` parameter which we will set to a ```1``` to skip the first row.
```
world_alcohol = np.genfromtxt("world_alcohol.csv", delimiter=",", dtype="U75", skip_header=1)
world_alcohol
```
## Extract a value from a numpy array
The extraction of an element from a numpy array (vectors) is done in a similar way to a list. For matrices, indexing is similar as for lists of list.
```
# example for vector
vector, vector[0]
# example for matrice
matrix, matrix[0,0]
```
### Training
In this section, we will try to answer the following questions:
* assign the number of liters of wine drunk by an Ivorian in 1985 to the variable value_ivory_1985, this corresponds to the data in the 3rd line
* assign the name of the country in the 2nd line to the variable second_country_name
```
world_alcohol
# number of liters of wine drunk by an Ivorian in 1985
value_ivory_1985 = world_alcohol[2,4]
print(value_ivory_1985)
# name of the country in the 2nd line
second_country_name = world_alcohol[1,2]
print(second_country_name)
```
## Extract a vector of values from a numpy array
The extraction of a vector from a numpy array is done in the same way as the extraction of an element. It consists in a kind of **slicing** from a list. However, the extraction of a matrix is a bit more complex. You can extract a row or column vector, or in a more advanced way, extract a subset of a matrix.
```
# example for vector
vector
sub_vector = vector[0:3]
sub_vector
# example for matrix
matrix
# column extraction
first_column = matrix[:,0]
first_column
second_column = matrix[:,1]
second_column
third_column = matrix[:,2]
third_column
# row extraction
first_row = matrix[0,:]
first_row
second_row = matrix[1,:]
second_row
third_row = matrix[2,:]
third_row
```
### Training
In this section, we will try to answer the following questions:
* assign all the 3rd column of world_alcohol to the countries variable
* assign the whole 5th column of world_alcohol to the variable alcohol_consumption
```
countries = world_alcohol[:,2]
countries
alcohol_consumption = world_alcohol[:,4]
alcohol_consumption
```
## Extract an array of values from a numpy array
In this section we will see how to extract an array of values (matrix or sub-matrix) from a numpy array of values (matrix). Extraction is done in the same way just by double slicing the rows and columns of a matrix.
```
# example
matrix
first_sub_matrix = matrix[:2,:2]
first_sub_matrix
second_sub_matrix = matrix[:2,1:3]
second_sub_matrix
third_sub_matrix = matrix[1:3,:2]
third_sub_matrix
fourth_sub_matrix = matrix[1:3,1:3]
fourth_sub_matrix
```
These extractions are interesting when the determinant of a matrix has to be calculated.
### Training
In this section, we will try to answer the following questions:
* assign all lines of the first 2 columns of world_acohol to the variable first_two_columns
* assign the first 10 rows of the first column of world_alcohol to the variable first_ten_years
* assign the first 10 rows of all world_alcohol columns to the variable first_ten_rows
* assign the first 20 rows of the world_alcohol index 1 and 2 columns to the variable first_twenty_regions
```
first_two_colomns = world_alcohol[:,:2]
first_two_colomns
first_ten_years = world_alcohol[:10,0]
first_ten_years
first_ten_rows = world_alcohol[:10,:]
first_ten_rows
first_twenty_regions = world_alcohol[:20,1:3]
first_twenty_regions
```
| github_jupyter |
```
from cforest.forest import CausalForest
# %load simulate.py
"""
This module provides functions for the simulation of data which is used in the
monte carlo simulation.
"""
import numpy as np
def simulate(
nobs,
nfeatures,
coefficients,
error_var,
seed,
propensity_score=None,
alpha=0.8,
):
"""Simulate data with heterogenous treatment effects.
Simulate outcomes y (length *nobs*), features X (*nobs* x *nfeatures*) and
treatment status treatment_status (length *nobs*) using the potential
outcome framework from Neyman and Rubin.
We simulate the data by imposing a linear model with two relevant
features plus a treatment effect. However, we return a design matrix with
*nfeatures* from which only the two used in the simulation are relevant.
The model looks the following:
y(0)_i = coef[0] + X_1i coef[1] + + X_2i coef[2] + error_i,
y(1)_i = coef[0] + X_1i coef[1] + + X_2i coef[2] + treatment_i + error_i,
where coef[0], coef[1] and coef[2] denote the intercept and slopes
respectively, and treatment_i = treatment(X_i) denotes the heterogenous
treatment effect, which is solely dependent on the location of the
individual in the feature space of the first two dimensions, i.e. as with
the linear model it only depends on X_1i and X_2i; at last y(0)_i, y(1)_i
denote the potential outcomes of individual i.
Args:
nobs (int): positive integer denoting the number of observations.
nfeatures (int): positiv integer denoting the number of features. Must
be greater than or equal to 2.
coefficients: Coefficients for the linear model, first value denotes
the intercept, second and third the slope for the second and third
feature. Must be convertable to a np.ndarray.
error_var (float): positive float denoting the error variance.
seed (int): seed for the random number generator.
propensity_score (np.array): array containing propensity scores, must
be of length *nobs*. If None, will be set to 0.5 for each
individual.
alpha (float): positive parameter influencing the shape of the
function. Default is 0.8.
Returns:
X (np.array): [nobs x nfeatures] numpy array with simulated features.
t (np.array): [nobs] boolean numpy array containing treatment
status of individuals.
y (np.array): [nobs] numpy array containing "observed" outcomes.
Raises:
ValueError, if dimensions mismatch or data type of inputs is incorrect.
"""
np.random.seed(seed)
# Assert input values
if not float.is_integer(float(nobs)):
raise ValueError("Argument *nobs* is not an integer.")
if not float.is_integer(float(nfeatures)):
raise ValueError("Argument *nfeatures* is not an integer.")
coefficients = np.array(coefficients)
if nfeatures < 2:
raise ValueError(
"Argument *nfeatures* must be greater or equal than" "two"
)
if len(coefficients) != 3:
raise ValueError("Argument *coefficients* needs to be of length 3.")
# Simulate treatment status
if propensity_score is None:
treatment_status = np.random.binomial(1, 0.5, nobs)
else:
if len(propensity_score) != nobs:
raise ValueError(
"Dimensions of argument *propensity_score* do not"
"match with *nobs*."
)
treatment_status = np.random.binomial(1, propensity_score, nobs)
error = np.random.normal(0, np.sqrt(error_var), nobs)
X = np.random.uniform(-15, 15, (nobs, nfeatures))
y = _compute_outcome(X, coefficients, treatment_status, error, alpha)
t = np.array(treatment_status, dtype=bool)
return X, t, y
def _compute_outcome(X, coefficients, treatment_status, error, alpha):
"""Compute observed potential outcome.
Simulates potential outcomes and returns outcome corresponding to the
treatment status.
Args:
X (np.array): design array containing features.
coefficients (np.array): coefficient array containing intercept and
two slope parameter.
treatment_status (np.array): array containing the treatment status
error (np.array): array containing the error terms for the linear model
alpha (float): positive parameter influencing the shape of the function
Returns:
y (np.array): the observed potential outcomes.
>>> import numpy as np
>>> X = np.array([[1, 2, 3], [4, 5, 6], [7, 8, 9]])
>>> coefficients = np.array([0, 1, -1])
>>> treatment_status = np.array([True, False, True])
>>> error = np.array([-0.1, 0, 0.1])
>>> alpha = 0.2
>>> _compute_outcome(X, coefficients, treatment_status, error, alpha)
array([-0.78932461, -1.0, -0.24908566])
"""
#baseline_model = coefficients[0] + np.dot(X[:, :2], coefficients[1:])
baseline_model = 0
y0 = baseline_model + error
treat_effect = true_treatment_effect(X[:, 0], X[:, 1], alpha)
y1 = y0 + treat_effect
y = (1 - treatment_status) * y0 + treatment_status * y1
return y
def true_treatment_effect(x, y, alpha=0.8, scale=5):
"""Compute individual treatment effects.
Computes individual treatment effect conditional on features *X* using
parameter *alpha* to determine the smoothness of the conditional
treatment function and *scale* to determine the scaling.
Args:
X (np.array): array with n rows and 2 columns depicting two features
for n individuals.
alpha (float): positive parameter influencing the shape of the
function. Defaults to 0.8.
scale (float): positive parameter determining the scaling of the
function. Defaults to 5. With scale=x the range of the function is
[0, x].
Returns:
result (np.array): array of length n containing individual treatment
effects.
>>> import numpy as np
>>> X = np.array([[0.5, 0.75], [0.25, 1]])
>>> alpha = 0.2
>>> _true_treatment_effect(X, alpha)
array([0.26475042, 0.26442005])
"""
denominatorx = 1 + np.exp(-alpha * (x - 1 / 3))
fractionx = 1 / denominatorx
denominatory = 1 + np.exp(-alpha * (y - 1 / 3))
fractiony = 1 / denominatory
result = scale * fractionx * fractiony
return result
from sklearn.model_selection import train_test_split
simparams = {
'nobs': 10000,
'nfeatures': 2,
'coefficients': [0.5, 0, 0],
'error_var': 0.1,
'seed': 1,
'alpha': 0.975
}
X, t, y = simulate(**simparams)
cf = CausalForest(
num_trees=50,
split_ratio=0.5,
min_leaf=5,
max_depth=20,
#use_transformed_outcomes=True,
num_workers=4,
seed_counter=1,
)
cf.fit(X, t, y)
XX = np.array([
[0.5, 0.75],
[0.25, 1],
[0.9, 0.9],
[0.1, 0.1]
])
cf.predict(XX)
```
| github_jupyter |
## Correlation between the Rank of Explanations ${\alpha}_i\phi(\mathbf{x}_i)^T\phi(\mathbf{x}_t)$ and term ${\alpha}_i\phi(\mathbf{x}_i)$ for RPS-$l_2$ and RPS-LJE
Figure 2
```
import torch
import numpy as np
from matplotlib import pyplot as plt
path = '../saved_models/base'
file = np.load('{}/model/saved_outputs.npz'.format(path))
intermediate_train = file['intermediate_train']
intermediate_test = file['intermediate_test']
labels_train = file['labels_train']
labels_test = file['labels_test']
pred_train = file['pred_train'].squeeze()
pred_test = file['pred_test'].squeeze()
weight_matrix_rep = np.load('{}/calculated_weights/representer_weight_matrix.npz'.format(path), allow_pickle=True)['weight_matrix']
weight_matrix_influence = np.load('{}/calculated_weights/influence_weight_matrix.npz'.format(path), allow_pickle=True)['weight_matrix'].squeeze()
jaccobian_test = np.load('{}/calculated_weights/influence_weight_matrix.npz'.format(path), allow_pickle=True)['jaccobian_test']
weight_matrix_ours = np.load('{}/calculated_weights/ours_weight_matrix_with_lr_0.01.npz'.format(path), allow_pickle=True)['weight_matrix'].squeeze()
alpha_fi_rep = weight_matrix_rep*intermediate_train
def get_all_influence(test_point):
representer_weight = weight_matrix_rep[:,0] * np.dot(intermediate_train,
intermediate_test[test_point,:])
ours_weight = np.dot(weight_matrix_ours,
intermediate_test[test_point,:])
return representer_weight, ours_weight
test_points = np.random.randint(0,2000, size=(500))
x_rep = []
x_ours = []
y = []
for test_point in test_points:
representer_weight, ours_weight = get_all_influence(test_point)
order_rep = np.abs(representer_weight).argsort()
order_ours = np.abs(ours_weight).argsort()
for i in range(10000):
y.append(i)
x_rep.append(np.linalg.norm(alpha_fi_rep[order_rep[i]]))
x_ours.append(np.linalg.norm(weight_matrix_ours[order_ours[i]]))
y = np.stack(y)
x_rep = np.stack(x_rep)
x_ours = np.stack(x_ours)
y_name = 'rank of $|\\alpha_ik(\mathbf{x}_i,\mathbf{x}_j)|$'
x_name = 'vector length of $\\alpha_i \phi(\mathbf{x}_i)$'
print(y_name)
import pandas as pd
df_rep = pd.DataFrame({y_name:y, x_name:x_rep})
df_ours = pd.DataFrame({y_name:y, x_name:x_ours})
# df_rep.to_csv('Importance_vs_weight_rep.csv')
# df_ours.to_csv('Importance_vs_weight_ours.csv')
# df_rep = pd.read_csv('Importance_vs_weight_rep.csv', index_col=0)
# df_ours = pd.read_csv('Importance_vs_weight_ours.csv', index_col=0)
plt.figure(figsize=(5,2.5))
plt.plot(x_name, y_name, data=df_rep,
marker='o',markersize=5, linestyle='none')
plt.xlabel(x_name)
plt.ylabel(y_name, labelpad=0)
plt.ticklabel_format(style='sci', axis='y', scilimits=(0,4))
plt.savefig('figs/order_vs_weight_rep.png', dpi=300, bbox_inches='tight')
plt.show()
plt.figure(figsize=(5,2.5))
plt.plot(x_name, y_name, data=df_ours,
marker='o',markersize=5, linestyle='none')
plt.xlabel(x_name)
plt.ylabel(y_name, labelpad=0)
plt.ticklabel_format(style='sci', axis='y', scilimits=(0,4))
plt.savefig('figs/order_vs_weight_ours.png', dpi=300, bbox_inches='tight')
plt.show()
```
| github_jupyter |
```
import os, sys
root_dir, _ = os.path.split(os.getcwd())
script_dir = os.path.join(root_dir, 'scripts')
sys.path.append(script_dir)
import tensorflow as tf
from tensorflow.keras import layers
import numpy as np
import tensorflow_probability as tfp
from hparams import hparams
from custom_layers_integer import Inv1x1ConvPermute, WaveNetIntegerBlock
from custom_layers_integer import DiscreteLogisticMixParametersWaveNet, FactorOutLayer
from training_utils import log_sum_exp
class WaveGlowInteger(tf.keras.Model):
"""
Waveglow implementation using the Invertible1x1Conv custom layer and
the WaveNet custom block
"""
def __init__(self, hparams, **kwargs):
super(WaveGlowInteger, self).__init__(dtype=hparams['ftype'], **kwargs)
assert(hparams['n_group'] % 2 == 0)
self.n_flows = hparams['n_flows']
self.n_group = hparams['n_group']
self.n_early_every = hparams['n_early_every']
self.n_early_size = hparams['n_early_size']
self.upsampling_size = hparams['upsampling_size']
self.hidden_channels = hparams['hidden_channels']
self.hparams = hparams
self.normalisation = hparams['train_batch_size'] * hparams['segment_length']
self.batch_size = hparams['train_batch_size']
self.seed = hparams['seed']
# Added for IDF
self.seeds = tf.random.uniform([self.n_flows], maxval=pow(2,16), seed=self.seed, dtype=tf.int32)
self.n_logistic_in_mixture = hparams["n_logistic_in_mixture"]
self.blocks = int(self.n_flows / self.n_early_every)
self.n_factorized_channels = self.n_early_size
self.xdim = hparams['xdim']
self.zdim = hparams['zdim']
self.compressing = False
self.last_log_shift = hparams['last_log_scale']
self.waveNetIntegerBlocks = []
self.inv1x1ConvPermuteLayers = []
# Added for IDF
self.factorOutLayers = []
self.discreteLogisticMixParametersNets = []
n_fourth = self.n_group // 4
n_remaining_channels = self.n_group
block = 0
for index in range(self.n_flows):
if ((index % self.n_early_every == 0) and (index > 0)):
n_fourth -= self.n_early_size // 4
n_remaining_channels -= self.n_early_size
self.factorOutLayers.append(
FactorOutLayer(
n_remaining_channels = n_remaining_channels,
n_early_size = self.n_early_size))
self.discreteLogisticMixParametersNets.append(
DiscreteLogisticMixParametersWaveNet(
n_factorized_channels=self.n_early_size,
n_logistic_in_mixture=self.n_logistic_in_mixture,
n_channels=hparams['n_channels'],
n_layers=hparams['n_layers'],
kernel_size=hparams['kernel_size'],
dtype=hparams['ftype']))
block += 1
self.inv1x1ConvPermuteLayers.append(
Inv1x1ConvPermute(
filters=n_remaining_channels,
seed=self.seeds[index],
dtype=hparams['ftype'],
name="newInv1x1conv_{}".format(index)))
self.waveNetIntegerBlocks.append(
WaveNetIntegerBlock(n_in_channels=n_fourth,
n_channels=hparams['n_channels'],
n_layers=hparams['n_layers'],
kernel_size=hparams['kernel_size'],
dtype=hparams['ftype'],
name="waveNetIntegerBlock_{}".format(index)))
self.n_remaining_channels = n_remaining_channels
self.n_blocks = block
def call(self, inputs, training=None):
"""
Evaluate model against inputs
"""
audio = inputs['wav']
audio = layers.Reshape(
target_shape = [self.hparams["segment_length"] // self.n_group,
self.n_group],
dtype=self.dtype) (audio)
output_audio = []
output_means = []
output_log_scales = []
output_logit = []
n_remaining_channels = self.n_group
block = 0
for index in range(self.n_flows):
if ((index % self.n_early_every == 0) and (index > 0)):
n_remaining_channels -= self.n_early_size
audio, output_chunk = self.factorOutLayers[block](audio, forward=True)
logit, means, log_scales = self.discreteLogisticMixParametersNets[block](audio, training=training)
output_audio.append(output_chunk)
output_logit.append(logit)
output_means.append(means)
output_log_scales.append(log_scales)
block += 1
audio = self.inv1x1ConvPermuteLayers[index](audio)
audio = self.waveNetIntegerBlocks[index](audio,
forward=True,
training=training)
# Last factored out audio will be encoded as discrete logistic
# The parameters are fixed and no mixture. To implement clean loss
# easier to generate mix of the same discrete logistic
audio = tf.reshape(audio, [audio.shape[0], audio.shape[1] * audio.shape[2], 1])
last_means = tf.zeros(audio.shape[:-1] + [self.n_logistic_in_mixture])
last_log_scales = tf.zeros(audio.shape[:-1] + [self.n_logistic_in_mixture]) + self.last_log_shift
last_logit = tf.concat([tf.ones(audio.shape),
tf.zeros(audio.shape[:-1] + [self.n_logistic_in_mixture - 1])],
axis=2)
# Append last outputs
output_audio.append(audio)
output_logit.append(last_logit)
output_means.append(last_means)
output_log_scales.append(last_log_scales)
# Concatenate outputs
output_means = tf.concat(output_means, axis=1)
output_logit = tf.concat(output_logit, axis=1)
output_log_scales = tf.concat(output_log_scales, axis=1)
output_audio = tf.concat(output_audio, axis=1)
return (output_audio, output_logit, output_means, output_log_scales)
def generate(self, signal, block):
n_factorized_channels = ((self.n_group - self.n_remaining_channels) // self.n_early_every)
target_shape = [self.batch_size,
(self.hparams["segment_length"] // self.n_group),
self.n_remaining_channels + (self.n_blocks - block) * n_factorized_channels]
audio = tf.reshape(signal, target_shape)
for index in list(reversed(range(self.n_flows)))[(self.n_blocks - block) * self.n_early_every:(self.n_blocks - block + 1) * self.n_early_every]:
audio = self.waveNetIntegerBlocks[index](audio, forward=False)
audio = self.inv1x1ConvPermuteLayers[index](audio, forward=False)
if block - 1 >= 0:
logits, means, log_scales = self.discreteLogisticMixParametersNets[block-1](audio)
else:
means = tf.zeros([self.batch_size, self.hparams["segment_length"] // self.n_remaining_channels, self.n_logistic_in_mixture])
log_scales = tf.zeros_like(means) + self.last_log_shift
logits = tf.concat([tf.ones([self.batch_size, self.hparams["segment_length"] // self.n_remaining_channels, 1]),
tf.zeros([self.batch_size, self.hparams["segment_length"] // self.n_remaining_channels, self.n_logistic_in_mixture - 1])],
axis=2)
target_shape = [self.batch_size,-1,1]
audio = tf.reshape(audio, target_shape)
means_shape = [self.batch_size,
(self.hparams["segment_length"] // self.n_group) * n_factorized_channels,
1]
return audio, (logits, means, log_scales)
def infer_craystack(self, inputs, training=None):
audio, logits, means, log_scales = self.call(inputs=inputs, training=training)
audio = [(tf.squeeze(x, axis=-1).numpy() * pow(2, 15)) + pow(2, 15) for x in tf.split(audio, self.n_blocks + 1, axis=1)]
all_params = [(tf.transpose(x, [0, 2, 1]).numpy().astype(np.float32),
tf.transpose(y, [0, 2, 1]).numpy().astype(np.float32),
tf.transpose(z, [0, 2, 1]).numpy().astype(np.float32)) for x,y,z in zip(tf.split(logits, self.n_blocks + 1, axis=1),
tf.split(means, self.n_blocks + 1, axis=1),
tf.split(log_scales, self.n_blocks + 1, axis=1))]
return audio, all_params
def generate_craystack(self, x=None, z=None, block=4):
if block == 4:
means = tf.zeros([self.batch_size, np.prod(self.zdim), self.hparams["n_logistic_in_mixture"]])
log_scales = tf.zeros_like(means) + self.last_log_shift
logits = tf.concat([tf.ones([self.batch_size, np.prod(self.zdim), 1]),
tf.zeros([self.batch_size, np.prod(self.zdim), self.hparams["n_logistic_in_mixture"] - 1])], axis=2)
else:
x = None if x is None else x
z = tf.convert_to_tensor((z - pow(2, 15)) / pow(2, 15) , dtype=tf.float32)
signal = z if x is None else tf.concat([tf.reshape(z, shape=[self.batch_size,
self.hparams['segment_length'] // self.n_group,
self.n_remaining_channels]),
tf.reshape(x, shape=[self.batch_size,
self.hparams['segment_length'] // self.n_group,
(self.n_blocks - block) * self.n_remaining_channels])],
axis=2)
x, (logits, means, log_scales) = self.generate(signal=signal, block=block)
logits = tf.transpose(logits, [0,2,1])
means = tf.transpose(means, [0,2,1])
log_scales = tf.transpose(log_scales, [0,2,1])
return x, (logits.numpy().astype(np.float32), means.numpy().astype(np.float32), log_scales.numpy().astype(np.float32))
def select_means_and_scales(self, logits, means, log_scales, output_shape):
argmax = tf.argmax(tf.nn.softmax(logits), axis=2)
selector = tf.one_hot(argmax,
depth=logits.shape[2],
dtype=tf.float32)
scales = tf.reduce_sum(tf.math.exp(log_scales) * selector, axis=2)
means = tf.reduce_sum(means * selector, axis=2)
return tf.reshape(means, output_shape), tf.reshape(scales, output_shape)
def set_compression(self):
self.compressing = True
self.batch_size = self.hparams['compress_batch_size']
def set_training(self):
self.compressing = False
self.batch_size = self.hparams['train_batch_size']
def get_config(self):
config = super(WaveGlow, self).get_config()
config.update(hparams = hparams)
return config
def sample_from_discretized_mix_logistic(self, logits, means, log_scales, log_scale_min=-8.):
'''
Args:
logits, means, log_scales: Tensor, [batch_size, time_length, n_logistic_in_mixture]
Returns:
Tensor: sample in range of [-1, 1]
Adapted from pixelcnn++
'''
# sample mixture indicator from softmax
temp = tf.random.uniform(logits.shape, minval=1e-5, maxval=1. - 1e-5)
temp = logits - tf.math.log(-tf.math.log(temp))
argmax = tf.math.argmax(temp, -1)
# [batch_size, time_length] -> [batch_size, time_length, nr_mix]
one_hot = tf.one_hot(argmax,
depth=self.n_logistic_in_mixture,
dtype=tf.float32)
# select logistic parameters
means = tf.reduce_sum(means * one_hot, axis=-1)
log_scales = tf.maximum(tf.reduce_sum(
log_scales * one_hot, axis=-1), log_scale_min)
# sample from logistic & clip to interval
# we don't actually round to the nearest 8-bit value when sampling
u = tf.random.uniform(means.shape, minval=1e-5, maxval=1. - 1e-5)
x = means + tf.math.exp(log_scales) * (tf.math.log(u) - tf.math.log(1 - u))
return tf.minimum(tf.maximum(x, -1. + 1e-5), 1. - 1e-5)
def total_loss(self, outputs, num_classes=65536, log_scale_min=-7.0):
'''
Discretized mix of logistic distributions loss.
Note that it is assumed that input is scaled to [-1, 1]
Adapted from pixelcnn++
'''
# audio[batch_size, time_length, 1] + unpack parameters: [batch_size, time_length, num_mixtures]
output_audio, logit_probs, means, log_scales = outputs
# output_audio [batch_size, time_length, 1] -> [batch_size, time_length, num_mixtures]
y = output_audio * tf.ones(shape=[1, 1, self.n_logistic_in_mixture], dtype=self.dtype)
centered_y = y - means
inv_stdv = tf.math.exp(-log_scales)
plus_in = inv_stdv * (centered_y + 1. / (num_classes - 1))
cdf_plus = tf.nn.sigmoid(plus_in)
min_in = inv_stdv * (centered_y - 1. / (num_classes - 1))
cdf_min = tf.nn.sigmoid(min_in)
log_cdf_plus = plus_in - tf.nn.softplus(plus_in) # log probability for edge case of -32768 (before scaling)
log_one_minus_cdf_min = - tf.nn.softplus(min_in) # log probability for edge case of 32767 (before scaling)
#probability for all other cases
cdf_delta = cdf_plus - cdf_min
mid_in = inv_stdv * centered_y
#log probability in the center of the bin, to be used in extreme cases
log_pdf_mid = mid_in - log_scales - 2. * tf.nn.softplus(mid_in)
log_probs = tf.where(y < -0.999, log_cdf_plus,
tf.where(y > 0.999, log_one_minus_cdf_min,
tf.where(cdf_delta > 1e-5,
tf.math.log(tf.maximum(cdf_delta, 1e-12)),
log_pdf_mid - np.log((num_classes - 1) / 2))))
log_probs = log_probs + tf.nn.log_softmax(logit_probs, axis=-1)
logistic_loss = -tf.reduce_sum(log_sum_exp(log_probs)) / (y.shape[0] * y.shape[1] * y.shape[2])
tf.summary.scalar(name='total_loss',
data=(logistic_loss))
for block, log_scales in enumerate(tf.split(log_scales, 4, axis=1)[:-1]):
tf.summary.scalar(name=f'mean_log_scales_block{block}',
data=tf.reduce_mean(log_scales),
step=tf.summary.experimental.get_step())
tf.summary.scalar(name=f'max_log_scales_block{block}',
data=tf.reduce_max(log_scales),
step=tf.summary.experimental.get_step())
tf.summary.scalar(name=f'min_log_scales_block{block}',
data=tf.reduce_min(log_scales),
step=tf.summary.experimental.get_step())
return logistic_loss
```
| github_jupyter |
## Ensemble Voting
Ensemble methods enable combining multiple model scores into a single score to create a robust generalized model. Ensemble voting is one of the simplest ways of combining the predictions from multiple machine learning algorithms. It works by first creating multiple standalone models from your training dataset. A Voting Classifier can then be used to wrap your models and average the predictions of the sub-models when asked to make predictions for new data.
The predictions of the sub-models can be weighted, but specifying the weights for classifiers manually or even heuristically is difficult. More advanced methods can learn how to best weight the predictions from submodels, but this is called stacking (stacked aggregation) and is currently not provided in scikit-learn.
```
import warnings
warnings.filterwarnings("ignore")
import matplotlib.pyplot as plt
%matplotlib inline
import pandas as pd
import numpy as np
# set seed for reproducability
np.random.seed(2017)
import statsmodels.api as sm
from sklearn import metrics
from sklearn.linear_model import LogisticRegression
from sklearn.naive_bayes import GaussianNB
from sklearn.ensemble import RandomForestClassifier
from sklearn.svm import SVC
from sklearn.neighbors import KNeighborsClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import AdaBoostClassifier
from sklearn.ensemble import BaggingClassifier
from sklearn.ensemble import GradientBoostingClassifier
# currently its available as part of mlxtend and not sklearn
from mlxtend.classifier import EnsembleVoteClassifier
from sklearn import cross_validation
from sklearn import metrics
from sklearn.cross_validation import train_test_split
# read the data in
df = pd.read_csv("Data/Diabetes.csv")
X = df.ix[:,:8] # independent variables
y = df['class'] # dependent variables
# evaluate the model by splitting into train and test sets
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=2017)
LR = LogisticRegression(random_state=2017)
RF = RandomForestClassifier(n_estimators = 100, random_state=2017)
SVM = SVC(random_state=0, probability=True)
KNC = KNeighborsClassifier()
DTC = DecisionTreeClassifier()
ABC = AdaBoostClassifier(n_estimators = 100)
BC = BaggingClassifier(n_estimators = 100)
GBC = GradientBoostingClassifier(n_estimators = 100)
clfs = []
print('5-fold cross validation:\n')
for clf, label in zip([LR, RF, SVM, KNC, DTC, ABC, BC, GBC],
['Logistic Regression',
'Random Forest',
'Support Vector Machine',
'KNeighbors',
'Decision Tree',
'Ada Boost',
'Bagging',
'Gradient Boosting']):
scores = cross_validation.cross_val_score(clf, X_train, y_train, cv=5, scoring='accuracy')
print("Train CV Accuracy: %0.2f (+/- %0.2f) [%s]" % (scores.mean(), scores.std(), label))
md = clf.fit(X, y)
clfs.append(md)
print("Test Accuracy: %0.2f " % (metrics.accuracy_score(clf.predict(X_test), y_test)))
```
From above benchmarking we see that 'Logistic Regression', 'Random Forest', 'Bagging', Ada/Gradient Boosting algorithms are giving better accuracy compared to other models. Let's combine non-similar models such as Logistic regression (base model), Random Forest (bagging model) and Gradient boosting (boosting model) to create a robust generalized model.
```
# re-building considering only the best performing non-similar models
clfs = []
print('5-fold cross validation:\n')
for clf, label in zip([LR, RF, GBC],
['Logistic Regression',
'Random Forest',
'Gradient Boosting']):
scores = cross_validation.cross_val_score(clf, X_train, y_train, cv=5, scoring='accuracy')
print("Train CV Accuracy: %0.2f (+/- %0.2f) [%s]" % (scores.mean(), scores.std(), label))
md = clf.fit(X, y)
clfs.append(md)
print("Test Accuracy: %0.2f " % (metrics.accuracy_score(clf.predict(X_test), y_test)))
```
#### Hard voting vs Soft voting
Majority voting is also known as hard voting. The argmax of the sum of predicted probabilities is known as soft voting. Parameteer 'weights' can be used to assign specific weightage to classifiers. The predicted class probabilities for each classifier are multiplied by the classifier weight, and averaged. Then the final class label is derived from the highest average probability class label.
Assume we assign equal weight of 1 to all classifiers (see below table). Based on soft voting, the predicted class label is 1, as it has the highest average probability.
Note: Some classifiers of scikit-learn do not support the predict_proba method
```
from IPython.display import Image
Image(filename='../Chapter 4 Figures/soft voting.png', width=500)
# ### Ensemble Voting
clfs = []
print('5-fold cross validation:\n')
ECH = EnsembleVoteClassifier(clfs=[LR, RF, GBC], voting='hard')
ECS = EnsembleVoteClassifier(clfs=[LR, RF, GBC], voting='soft', weights=[1,1,1])
for clf, label in zip([ECH, ECS],
['Ensemble Hard Voting',
'Ensemble Soft Voting']):
scores = cross_validation.cross_val_score(clf, X_train, y_train, cv=5, scoring='accuracy')
print("Train CV Accuracy: %0.2f (+/- %0.2f) [%s]" % (scores.mean(), scores.std(), label))
md = clf.fit(X, y)
clfs.append(md)
print("Test Accuracy: %0.2f " % (metrics.accuracy_score(clf.predict(X_test), y_test)))
```
| github_jupyter |
# A machine learning decision tree approach
The iMeta algorithm is essentially a decision tree algorithm, where the variables and threshold for the decisions at each step are manually specified based on human analysis. The simplest way to apply machine learning techniques to the problem would be to use a similar structure to iMeta, which is a decision tree, but use standard ML training techiniques to learn the parameters such as what thresholds to use and how many branches/leaves to have in the tree for the best results.
```
import os
import sys
import pathlib
import functools
import pandas
import numpy
import matplotlib
import matplotlib.pyplot
import warnings
warnings.filterwarnings('ignore')
import ipywidgets
import time
import sklearn
import sklearn.model_selection
import sklearn.preprocessing
import sklearn.tree
import sklearn.metrics
root_repo_dir = pathlib.Path().absolute().parent
sys.path = [os.path.join(root_repo_dir)] + sys.path
import xbt.dataset
from xbt.dataset import XbtDataset, UNKNOWN_STR, cat_output_formatter, check_value_found
from xbt.imeta import imeta_classification, XBT_MAX_DEPTH
# Set up some site specific parameters for the notebook
try:
environment = os.environ['XBT_ENV_NAME']
except KeyError:
environment = 'pangeo'
root_data_dirs = {
'MO_scitools': '/data/users/shaddad/xbt-data/',
'pangeo': '/data/misc/xbt-data/',
}
env_date_ranges = {
'MO_scitools': (1966,2015),
'pangeo': (1966,2015)
}
# Set up some dataset specific parameters
root_data_dir = root_data_dirs[environment]
year_range = env_date_ranges[environment]
cv_metric_names = ['f1_weighted','precision_weighted','recall_weighted']
input_feature_names = ['country','max_depth', 'year', 'lat', 'lon']
input_dir_name = 'csv_with_imeta'
exp_out_dir_name = 'experiment_outputs'
experiment_name = 'nb_single_decisionTree_country'
classifier_class = sklearn.tree.DecisionTreeClassifier
classifier_name = 'decision_tree'
suffix='countryAndLatLon'
classifier_opts = {'max_depth': 20,
'min_samples_leaf': 1,
'criterion': 'gini'
}
xbt_input_dir = os.path.join(root_data_dir, input_dir_name)
xbt_output_dir = os.path.join(root_data_dir, exp_out_dir_name, experiment_name)
# create the output for this experiment if it doesn't exist
if not os.path.isdir(xbt_output_dir):
os.makedirs(xbt_output_dir)
print(f'outputting to {xbt_output_dir}')
output_fname_template = 'xbt_output_{exp_name}_{subset}.csv'
result_fname_template = 'xbt_metrics_{classifier}_{suffix}.csv'
%%time
xbt_full_dataset = XbtDataset(xbt_input_dir, year_range)
```
## Data preparation
We are only testing on the labelled data, to be able to evluate performance. The XbtDataset class has filtered out some bad data including profiles with maximum depths less that 0.0 or greater than 2000.0. There were also some profiles with bad date entries, which have been excluded for now.
```
%%time
xbt_labelled_raw = xbt_full_dataset.filter_obs({'labelled': 'labelled'})
xbt_labelled_raw.shape
xbt_labelled = xbt_labelled_raw.filter_obs({'country': 'UNKNOWN'}, mode='exclude').filter_obs({'cruise_number': '0'}, mode='exclude', check_type='exact')
xbt_labelled.shape
_ = xbt_labelled.get_ml_dataset(return_data = False)
_ = xbt_labelled.filter_features(['instrument','model','manufacturer']).encode_target(return_data = False)
%%time
unseen_cruise_numbers = xbt_labelled.sample_feature_values('cruise_number', fraction=0.1)
%%time
xbt_unseen = xbt_labelled.filter_obs({'cruise_number': unseen_cruise_numbers}, mode='include', check_type='in_filter_set')
xbt_working = xbt_labelled.filter_obs({'cruise_number': unseen_cruise_numbers}, mode='exclude', check_type='in_filter_set')
imeta_classes = xbt_labelled.xbt_df.apply(imeta_classification, axis=1)
imeta_model = imeta_classes.apply(lambda t1: t1[0])
imeta_manufacturer = imeta_classes.apply(lambda t1: t1[1])
imeta_instrument = imeta_classes.apply(lambda t1: f'XBT: {t1[0]} ({t1[1]})')
```
We are currently training and evaulating separately for model and manufacturer. We will also need to train and evaulate together as this is ultimately what is wanted (a combined probe model and manufacturer field).
We are using the default 80/20 split in scikit-learn for now. Further work will need to do proper cross validation where several different splits are randomly selected to verify our results are not an artifact of the randomly chosen split.
```
%%time
xbt_train_all, xbt_test_all = xbt_working.train_test_split(refresh=True, features=['instrument', 'year'])
X_train_all = xbt_train_all.filter_features(input_feature_names).get_ml_dataset()[0]
X_test_all = xbt_test_all.filter_features(input_feature_names).get_ml_dataset()[0]
X_unseen_all = xbt_unseen.filter_features(input_feature_names).get_ml_dataset()[0]
y_instr_train_all = xbt_train_all.filter_features(['instrument']).get_ml_dataset()[0]
y_instr_test_all = xbt_test_all.filter_features(['instrument']).get_ml_dataset()[0]
y_instr_unseen_all = xbt_unseen.filter_features(['instrument']).get_ml_dataset()[0]
```
## Training the classifier
We are using the scikit-learn classifier as the closest analogue to the structure of the iMeta algorithm. This tree can have many more nodes and leaves than iMeta though. it is quick to train and evaluate so it is a useful starting point for setting up the ML processing pipelines, as all the scikit-learn classifiers have a common interface.
For the model and manufacturer, we train a Decision ree Classifier, then use it to predict values for the train and test sets. We then calculate the accuracy metrics for each for the whole dataset.
I am using precision, recall and F1 as fairly standard ML metrics of accuracy. Recall is what has been used in the two previous papers (Palmer et. al, Leahy and Llopis et al) so that is the focus. Support is a useful to see what proportion of the profiles in the dataset belong to each of the different classes.
```
metrics_per_class_all = {}
metrics_avg_all = {}
clf_dt_instr1 = classifier_class(**classifier_opts)
clf_dt_instr1.fit(X_train_all,y_instr_train_all)
metrics_per_class_all['instrument'] = list(xbt_labelled._feature_encoders['instrument'].classes_)
y_res_train_instr_all = clf_dt_instr1.predict(X_train_all)
metrics1 = sklearn.metrics.precision_recall_fscore_support(y_instr_train_all, y_res_train_instr_all, labels=list(range(0,len(metrics_per_class_all['instrument']))))
metrics_per_class_all.update( {
'precision_instr_train': metrics1[0],
'recall_instr_train': metrics1[1],
'f1_instr_train': metrics1[2],
'support_instr_train': metrics1[3],
})
metrics_avg_all.update({
'precision_instr_train' : sum(metrics1[0] * metrics1[3])/ sum(metrics1[3]),
'recall_instr_train' : sum(metrics1[1] * metrics1[3])/ sum(metrics1[3]),
'f1_instr_train' : sum(metrics1[2] * metrics1[3])/ sum(metrics1[3]),
})
y_res_test_instr_all = clf_dt_instr1.predict(X_test_all)
metrics1 = sklearn.metrics.precision_recall_fscore_support(y_instr_test_all, y_res_test_instr_all, labels=list(range(0,len(metrics_per_class_all['instrument']))))
metrics_per_class_all.update( {
'precision_instr_test': metrics1[0],
'recall_instr_test': metrics1[1],
'f1_instr_test': metrics1[2],
'support_instr_test': metrics1[3],
})
metrics_avg_all.update({
'precision_instr_test' : sum(metrics1[0] * metrics1[3])/ sum(metrics1[3]),
'recall_instr_test' : sum(metrics1[1] * metrics1[3])/ sum(metrics1[3]),
'f1_instr_test' : sum(metrics1[2] * metrics1[3])/ sum(metrics1[3]),
})
y_res_unseen_instr_all = clf_dt_instr1.predict(X_unseen_all)
metrics1 = sklearn.metrics.precision_recall_fscore_support(y_instr_unseen_all, y_res_unseen_instr_all, labels=list(range(0,len(metrics_per_class_all['instrument']))))
metrics_per_class_all.update( {
'precision_instr_unseen': metrics1[0],
'recall_instr_unseen': metrics1[1],
'f1_instr_unseen': metrics1[2],
'support_instr_unseen': metrics1[3],
})
metrics_avg_all.update({
'precision_instr_unseen' : sum(metrics1[0] * metrics1[3])/ sum(metrics1[3]),
'recall_instr_unseen' : sum(metrics1[1] * metrics1[3])/ sum(metrics1[3]),
'f1_instr_unseen' : sum(metrics1[2] * metrics1[3])/ sum(metrics1[3]),
})
df_metrics_per_class_instr = pandas.DataFrame.from_dict({k1:v1 for k1,v1 in metrics_per_class_all.items() if 'instr' in k1})
df_metrics_avg = pandas.DataFrame.from_dict({
'target': ['instrument_train','instrument_test', 'instrument_unseen'],
'precision': [v1 for k1,v1 in metrics_avg_all.items() if 'precision' in k1],
'recall': [v1 for k1,v1 in metrics_avg_all.items() if 'recall' in k1],
'f1': [v1 for k1,v1 in metrics_avg_all.items() if 'f1' in k1],
})
df_metrics_avg
```
# Classification result plots
The plots below show the results for the whole XBT dataset. We see that the DT classifier performs well on the training data, but does not seem to generalise well. This especially true, as one would expect, for classes with very little support in the training dataset.
```
fig_results_all_dt = matplotlib.pyplot.figure('xbt_results_all_dt', figsize=(24,8))
axis_instr_metrics = fig_results_all_dt.add_subplot(121)
_ = df_metrics_per_class_instr.plot.bar(x='instrument', y=['recall_instr_train','recall_instr_test','recall_instr_unseen'],ax=axis_instr_metrics)
axis_instr_support = fig_results_all_dt.add_subplot(122)
_ = df_metrics_per_class_instr.plot.bar(x='instrument',y=['support_instr_train', 'support_instr_test', 'support_instr_unseen'], ax=axis_instr_support)
df_metrics_avg.plot.bar(figsize=(18,12), x='target', y='recall')
```
## Classification results
The contents of the XBT dataset varies over the time period, so previous papers have looked at classification accuracy (recall) year by year to evaluate how performance varies with different distribution of probe types.
To do this we apply the classifier to the train and test data for each year separetly and calculate the metrics year by year.
```
def score_year(xbt_df, year, clf, input_features, target_feature):
X_year = xbt_df.filter_obs({'year': year}, ).filter_features(input_features).get_ml_dataset()[0]
y_year = xbt_df.filter_obs({'year': year} ).filter_features([target_feature]).get_ml_dataset()[0]
y_res_year = clf.predict(X_year)
metric_year = sklearn.metrics.precision_recall_fscore_support(
y_year, y_res_year, average='micro')
return metric_year
eval_progress = ipywidgets.IntProgress(min=env_date_ranges[environment][0],
max= env_date_ranges[environment][1],
description='Evaluating',
bar_style='info')
eval_progress
results_by_year = {}
for year in range(env_date_ranges[environment][0],env_date_ranges[environment][1]):
results_by_year[year] = {
'metric_train_instr' : score_year(xbt_train_all, year, clf_dt_instr1, input_feature_names, 'instrument'),
'metric_test_instr' : score_year(xbt_test_all, year, clf_dt_instr1, input_feature_names, 'instrument'),
'metric_unseen_instr' : score_year(xbt_unseen, year, clf_dt_instr1, input_feature_names, 'instrument'),
}
eval_progress.value = year
recall_by_year = pandas.DataFrame.from_dict({
'year': list(results_by_year.keys()),
'recall_train_instr' : [m1['metric_train_instr'][1] for y1,m1 in results_by_year.items()],
'recall_test_instr' : [m1['metric_test_instr'][1] for y1,m1 in results_by_year.items()],
'recall_unseen_instr' : [m1['metric_unseen_instr'][1] for y1,m1 in results_by_year.items()],
})
instr_encoder = xbt_labelled._feature_encoders['instrument']
eval_progress.value = env_date_ranges[environment][0]
imeta_results = []
for year in range(env_date_ranges[environment][0],env_date_ranges[environment][1]):
y_imeta_instr = instr_encoder.transform(pandas.DataFrame(imeta_instrument[xbt_labelled.xbt_df.year == year]))
xbt_instr1 = instr_encoder.transform(pandas.DataFrame(xbt_labelled.xbt_df[xbt_labelled.xbt_df.year == year].instrument))
(im_pr_instr, im_rec_instr, im_f1_instr, im_sup_instr) = sklearn.metrics.precision_recall_fscore_support(xbt_instr1, y_imeta_instr,average='micro')
imeta_results += [{'year': year,
'imeta_instr_recall': im_rec_instr,
'imeta_instr_precision': im_pr_instr,
}]
eval_progress.value = year
imeta_res_df = pandas.DataFrame.from_records(imeta_results)
results_df = pandas.merge(recall_by_year, imeta_res_df).merge(
pandas.DataFrame.from_dict({
'year': xbt_labelled['year'].value_counts(sort=False).index,
'num_samples': xbt_labelled['year'].value_counts(sort=False).values,
}))
fig_model_recall_results = matplotlib.pyplot.figure('xbt_model_recall', figsize=(12,12))
ax_instr_recall_results = fig_model_recall_results.add_subplot(111, title='XBT instrument recall results')
_ = results_df.plot.line(x='year',y=['recall_train_instr','recall_test_instr', 'recall_unseen_instr', 'imeta_instr_recall'], ax=ax_instr_recall_results)
results_df['improvement_instr'] = results_df.apply(lambda r1: ((r1['recall_test_instr'] / r1['imeta_instr_recall'])-1)*100.0 , axis=1)
fig_num_samples_per_year = matplotlib.pyplot.figure('fig_num_samples_per_year', figsize=(16,16))
ax_num_samples = fig_num_samples_per_year.add_subplot(121, title='number of samples per year')
_ = results_df.plot.line(ax=ax_num_samples, x='year',y=['num_samples'],c='purple' )
ax_num_samples = fig_num_samples_per_year.add_subplot(122, title='improvement instrument per year')
_ = results_df.plot.line(ax=ax_num_samples, x='year',y=['improvement_instr'], c='green' )
```
| github_jupyter |
# Training the Adaboost classifier with HOG&LBP features
```
import scipy
import numpy as np
from scipy import interp
from sklearn import preprocessing, cross_validation, neighbors,datasets, svm
from sklearn import datasets
from sklearn.preprocessing import StandardScaler
from sklearn.neural_network import MLPClassifier
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
from sklearn.metrics import roc_curve, auc
from sklearn.datasets import make_hastie_10_2
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.model_selection import cross_val_score
from sklearn.ensemble import AdaBoostClassifier
import matplotlib.pyplot as plt
from sklearn.metrics import cohen_kappa_score
fileName = 'HOGLBP'
x = scipy.io.loadmat(fileName)
cc = [x][0]
HOGLBP = cc['HOGLBP']
HOGLBP = np.asarray(HOGLBP)
print "HOGLBP dataset loaded"
fileName = 'LBPlabels'
x = scipy.io.loadmat(fileName)
cc = [x][0]
LBPlabels = cc['WW']
LBPlabels = np.asarray(LBPlabels)
print 'labels loading completed'
c, r = LBPlabels.shape
LBPlabels = LBPlabels.reshape(c,)
X_train, X_test, y_train, y_test = cross_validation.train_test_split(HOGLBP, LBPlabels, test_size=0.3, random_state=0)
#scaling
scaler = StandardScaler()
#scaler = preprocessing.MinMaxScaler()
# Fit only on training data
scaler.fit(X_train)
X_train = scaler.transform(X_train)
# apply same transformation to test data
X_test = scaler.transform(X_test)
clf = AdaBoostClassifier(n_estimators=500)
scores = cross_val_score(clf, X_train, y_train)
scores.mean()
clf.fit(X_train, y_train)
Accuracy = clf.score(X_train, y_train)
print "Accuracy in the training data: ", Accuracy*100, "%"
accuracy = clf.score(X_test, y_test)
print "Accuracy in the test data", accuracy*100, "%"
y_pred = clf.predict(X_train)
print '\nTraining classification report\n', classification_report(y_train, y_pred)
print "\n Confusion matrix of training \n", confusion_matrix(y_train, y_pred)
y_pred = clf.predict(X_test)
print '\nTesting classification report\n', classification_report(y_test, y_pred)
print "\nConfusion matrix of the testing\n", confusion_matrix(y_test, y_pred)
probas = clf.fit(X_train, y_train).predict_proba(X_test)
fpr, tpr, thresholds = roc_curve(y_test, probas[:, 1])
mean_tpr = 0.0
mean_fpr = np.linspace(0, 1, 100)
mean_tpr += interp(mean_fpr, fpr, tpr)
mean_tpr[0] = 0.0
roc_auc = auc(fpr, tpr)
print "\nArea Under the ROC curve: ", roc_auc
meanTP = 0
for t in tpr:
meanTP += t
print "Mean True Positive rate (testing): ", meanTP/len(tpr)
meanFP = 0
for t in fpr:
meanFP += t
print "Mean False Positive rate (testing): ", meanFP/len(fpr)
plt.title('Receiver Operating Characteristic')
plt.plot(fpr, tpr, 'b', label = 'AUC = %0.2f' % roc_auc)
plt.legend(loc = 'lower right')
plt.plot([0, 1], [0, 1],'r--')
plt.xlim([0, 1])
plt.ylim([0, 1])
plt.ylabel('True Positive Rate')
plt.xlabel('False Positive Rate')
plt.show()
```
# Persisting the model
```
from sklearn.externals import joblib
joblib.dump(clf, 'AdaBoostTBModel.pkl')
print "AdaBoostTBModel Generated"
#X is the sample you want to classify (HOG &LBP features of an x-ray)
#Image size used in the training is 100*100pixels
#You can pass one image or a numpy array of more than one
#to use the model for testing uncomment the following lines:
#clf = joblib.load('AdaBoostTBModel.pkl')
#clf.predict(X)
```
| github_jupyter |
# A short demo of PyArrow & Neo4j 😀
<img src="https://arrow.apache.org/docs/_static/arrow.png" style="width:200px;">
## Our Dependencies
Nothing special really...the usual cast of characters with the addition of `pyarrow`
```
%pip install pyarrow pandas scikit-learn matplotlib seaborn
%matplotlib inline
```
### Imports and our Integration
We'll set up our imports next.
One special import is [neo4j_arrow](https://gist.github.com/voutilad/ac1107a383affcdfdaaf4a08d1f14df4), the client wrapper to simplify talking to the server-side `Neo4j-Arrow` service. It's like **100 lines of Python** and uses the `PyArrow` framework...no Neo4j code!
> Server-side it exists as a database plugin. If you have access, the code is here: https://github.com/neo4j-field/neo4j-arrow
```
# Get our DS imports ready!
from sklearn.decomposition import PCA
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
# To time stuff...
import time
# And our neo4j integration!
import neo4j_arrow
```
---
## Connecting our Neo4jArrow Client
Very simple. We provide access credentials (username, password) and then provide the location of the server in a tuple of `(host, port)`.
`Neo4jArrow` uses Neo4j's built-in authorization framework. All our calls to the server are authenticated like any other Neo4j client.
```
client = neo4j_arrow.Neo4jArrow('neo4j', 'password',
('voutila-arrow-test', 9999),
tls=True, verify_tls=False)
```
### Discovering Available Actions
Arrow Flight uses an RPC concept. In short, clients can perform _Actions_ sending optional payload data to the server with each action. Clients can also consume or put _streams_ to/from the server.
Let's discover our available actions!
```
actions = client.list_actions()
for action in actions:
print(action)
```
Each of these actions can be called by an Apache Arrow client, regardless if it's PyArrow or the Arrow R package or Arrow for Rust!
---
## Working with Cypher Jobs
The way I've architected `Neo4jArrow` is designed around submitting "jobs" that construct streams. Let's submit some Cypher!
> Note: Cypher jobs are orders of magnitude _slower_ than GDS jobs...but still faster than
> using a Neo4j Python driver by an order of magnitude!
```
cypher = """
UNWIND range(1, $rows) AS row
RETURN row, [_ IN range(1, $dimension) | rand() ] AS n;
"""
params = {
"rows": 1_000_000,
"dimension": 256
}
print(f"Submitting cypher with params:\n{cypher}\n{params}")
ticket = client.cypher(cypher, params=params)
print(f"Got a ticket: {ticket}")
```
### Waiting for our Results (Optional)
Each job results in a _ticket_. Clients use the ticket to check on job status or request a stream of the results.
Let's wait until our Cypher is producing results and our stream is ready for consumption. This little helper function just polls the `jobStatus` _Action_ waiting for our job to be in a "producing" state.
```
print(f'>> Polling for status on ticket {ticket}...')
ready = client.wait_for_job(ticket, timeout=35)
if not ready:
raise Exception('something is wrong...did you submit a job?')
else:
print('>> Stream is Ready!')
```
### Consuming our Results
Clients consume streams by presenting their ticket. They bet back a PyArrow stream reader and have some options to how they consume the stream:
1. They can iterate over batches in the stream and process them incrementally.
2. They can consume the entire stream into a PyArrow Table
3. They can consume the entire stream immediately into a Pandas data frame
```
# Let's get a dataframe!
print('>> Reading the result of our Cypher job into a dataframe. Please wait...')
start = time.time()
table = client.stream(ticket).read_all()
delta = round(time.time() - start, 1)
print(f'>> Read our stream entirely into a PyArrow table in {delta} seconds!')
print(table)
megs = table.nbytes >> 20
print(f"Table is approximately {megs:,} MiB")
```
#### Let's Work with Pandas and Scikit-Learn!
```
# We'll convert our series of embedding arrays into a NumPy matrix
print(">> Converting to a Pandas DataFrame...")
df = table.select(['n'])[0].to_pandas()
print(">> Building a NumPy matrix...")
m = np.matrix(df.tolist())
# Then let's do dimensional reduction so we can plot our vectors
pca = PCA(n_components=2)
print(">> Fitting PCA transform...please wait!")
pc = pca.fit_transform(m)
print('>> Before, our data looked like :')
print(table.select(['n'])[0][0:1])
print('>> Now we have a matrix like:')
print(pc)
```
Let's plot!
```
pc_df = pd.DataFrame(data=pc, columns=['PC1', 'PC2'])
sns.lmplot( x="PC1", y="PC2",
data=pc_df,
fit_reg=False)
```
---
## Now for the fun stuff: Direct GDS Integration!
We just played with Cypher, which is fine...but what about working with _even more data_ from things like GDS?
Our traditional methods using the Python driver would absolutely choke here...but with PyArrow, I can stay in my comfy little Python world and still get data _fast_.
### Submitting a GDS Job
We'll submit a GDS job that reads directly from the in-memory graph. In this case, it's analagous to something like:
```
CALL gds.graph.streamNodeProperties('random', ['fastRp'])
```
Suppose we already have a random graph with node embeddings built via something like:
```
CALL gds.beta.graph.generate('random', toInteger(5e6), 5);
CALL gds.fastRP.mutate('random', {
embeddingDimension: 256,
mutateProperty: 'fastRp'
});
```
> Note that this follows the same general flow as before: submit job, get ticket, get stream
```
# Submit our GDS job to retrieve some node embeddings from a graph projection
ticket = client.gds_nodes('random', properties=['fastRp'])
client.wait_for_job(ticket, timeout=15)
print('>> Reading the result of our GDS job...''')
start = time.time()
# Retrieve and consume the stream into a PyArrow Table
table = client.stream(ticket).read_all()
delta = round(time.time() - start, 2)
gigs, rows = round((table.nbytes >> 20) / 1024.0, 2), table.num_rows
print(f'>> Took {delta:,}s to consume stream into a PyArrow table.')
print(f'>> Table has {rows:,} rows, represents ~{gigs:,} GiB.')
# And convert to Pandas!
start = time.time()
df = table.to_pandas()
delta = round(time.time() - start, 2)
print(f'>> Built a Pandas DataFrame in {delta:,} seconds!')
print(df)
```
Now we'll do our little data conversion and PCA...
```
# Select out just our embedding vectors and convert it to a numpy matrix
m = np.matrix(df['fastRp'].tolist())
# Then let's do dimensional reduction so we can plot our vectors in a lower dimension
print('>> Performing dimensional reduction...please wait!')
pca = PCA(n_components=3)
print('>> Fitting transform...')
pc = pca.fit_transform(m)
print('>> Our new 3-dimensional vectors look like:')
print(pc)
```
And plot!
```
pc_df = pd.DataFrame(data=pc, columns=['PC1', 'PC2', 'PC3'])
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
ax.scatter(pc_df['PC1'], pc_df['PC2'], pc_df['PC3'])
plt.show()
```
---
## Towards the Future 🚀
Other languages supported by Apache Arrow:
* R
* Matlab
* Julia
* and more!
Some ponderings:
* can we do better bulk updates/writes by moving data via Arrow to the server?
* how about replicating an entire in-memory graph to another Neo4j system?
* if we ditch the whole 2-step process (get ticket, get stream) how simple can our DX be?
| github_jupyter |
# Part 8 - Introduction to Plans
### Context
We introduce here an object which is crucial to scale to industrial Federated Learning: the Plan. It reduces dramatically the bandwidth usage, allows asynchronous schemes and give more autonomy to remote devices. The original concept of plan can be found in the paper [Towards Federated Learning at Scale: System Design](https://arxiv.org/pdf/1902.01046.pdf), but it has been adapted to our needs in the PySyft library.
A Plan is intended to store a sequence of torch operations, just like a function, but it allows to send this sequence of operations to remote workers and to keep a reference to it. This way, to compute remotely this sequence of $n$ operations on some remote input referenced through pointers, instead of sending $n$ messages you need now to send a single message with the references of the plan and the pointers. You can also provide tensors with your function (that we call _state tensors_) to have extended functionalities. Plans can be seen either like a function that you can send, or like a class which can also be sent and executed remotely. Hence, for high level users, the notion of plan disappears and is replaced by a magic feature which allow to send to remote workers arbitrary functions containing sequential torch functions.
One thing to notice is that the class of functions that you can transform into plans is currently limited to sequences of hooked torch operations exclusively. This excludes in particular logical structures like `if`, `for` and `while` statements, even if we are working to have workarounds soon. _To be completely precise, you can use these but the logical path you take (first `if` to False and 5 loops in `for` for example) in the first computation of your plan will be the one kept for all the next computations, which we want to avoid in the majority of cases._
Authors:
- Théo Ryffel - Twitter [@theoryffel](https://twitter.com/theoryffel) - GitHub: [@LaRiffle](https://github.com/LaRiffle)
- Bobby Wagner - Twitter [@bobbyawagner](https://twitter.com/bobbyawagner) - GitHub: [@robert-wagner](https://github.com/robert-wagner)
- Marianne Monteiro - Twitter [@hereismari](https://twitter.com/hereismari) - GitHub: [@mari-linhares](https://github.com/mari-linhares)
### Imports and model specifications
First let's make the official imports.
```
import torch
import torch.nn as nn
import torch.nn.functional as F
```
And than those specific to PySyft, with one important note: **the local worker should not be a client worker.** *Non client workers can store objects and we need this ability to run a plan.*
```
import syft as sy # import the Pysyft library
hook = sy.TorchHook(torch) # hook PyTorch ie add extra functionalities
# IMPORTANT: Local worker should not be a client worker
hook.local_worker.is_client_worker = False
server = hook.local_worker
```
We define remote workers or _devices_, to be consistent with the notions provided in the reference article.
We provide them with some data.
```
x11 = torch.tensor([-1, 2.]).tag('input_data')
x12 = torch.tensor([1, -2.]).tag('input_data2')
x21 = torch.tensor([-1, 2.]).tag('input_data')
x22 = torch.tensor([1, -2.]).tag('input_data2')
device_1 = sy.VirtualWorker(hook, id="device_1", data=(x11, x12))
device_2 = sy.VirtualWorker(hook, id="device_2", data=(x21, x22))
devices = device_1, device_2
```
### Basic example
Let's define a function that we want to transform into a plan. To do so, it's as simple as adding a decorator above the function definition!
```
@sy.func2plan()
def plan_double_abs(x):
x = x + x
x = torch.abs(x)
return x
```
Let's check, yes we have now a plan!
```
plan_double_abs
```
To use a plan, you need two things: to build the plan (_ie register the sequence of operations present in the function_) and to send it to a worker / device. Fortunately you can do this very easily!
#### Building a plan
To build a plan you just need to call it on some data.
Let's first get a reference to some remote data: a request is sent over the network and a reference pointer is returned.
```
pointer_to_data = device_1.search('input_data')[0]
pointer_to_data
```
If we tell the plan it must be executed remotely on the device`location:device_1`... we'll get an error because the plan was not built yet.
```
plan_double_abs.is_built
# Sending non-built Plan will fail
try:
plan_double_abs.send(device_1)
except RuntimeError as error:
print(error)
```
To build a plan you just need to call `build` on the plan and pass the arguments needed to execute the plan (a.k.a some data). When a plan is built all the commands are executed sequentially by the local worker, and are catched by the plan and stored in its `actions` attribute!
```
plan_double_abs.build(torch.tensor([1., -2.]))
plan_double_abs.is_built
```
If we try to send the plan now it works!
```
# This cell is executed successfully
pointer_plan = plan_double_abs.send(device_1)
pointer_plan
```
As with then tensors, we get a pointer to the object sent. Here it is simply called a `PointerPlan`.
One important thing to remember is that when a plan is built we pre-set ahead of computation the id(s) where the result(s) should be stored. This will allow to send commands asynchronously, to already have a reference to a virtual result and to continue local computations without waiting for the remote result to be computed. One major application is when you require computation of a batch on device_1 and don't want to wait for this computation to end to launch another batch computation on device_2.
#### Running a Plan remotely
We can now remotely run the plan by calling the pointer to the plan with a pointer to some data. This issues a command to run this plan remotely, so that the predefined location of the output of the plan now contains the result (remember we pre-set location of result ahead of computation). This also requires a single communication round.
The result is simply a pointer, just like when you call an usual hooked torch function!
```
pointer_to_result = pointer_plan(pointer_to_data)
print(pointer_to_result)
```
And you can simply ask the value back.
```
pointer_to_result.get()
```
### Towards a concrete example
But what we want to do is to apply Plan to deep and federated learning, right? So let's look to a slightly more complicated example, using neural networks as you might be willing to use them.
Note that we are now transforming a class into a plan. To do so, we inherit our class from sy.Plan (instead of inheriting from nn.Module).
```
class Net(sy.Plan):
def __init__(self):
super(Net, self).__init__()
self.fc1 = nn.Linear(2, 3)
self.fc2 = nn.Linear(3, 2)
def forward(self, x):
x = F.relu(self.fc1(x))
x = self.fc2(x)
return F.log_softmax(x, dim=0)
net = Net()
net
```
Let's build the plan using some mock data.
```
net.build(torch.tensor([1., 2.]))
```
We now send the plan to a remote worker
```
pointer_to_net = net.send(device_1)
pointer_to_net
```
Let's retrieve some remote data
```
pointer_to_data = device_1.search('input_data')[0]
```
Then, the syntax is just like normal remote sequential execution, that is, just like local execution. But compared to classic remote execution, there is a single communication round for each execution.
```
pointer_to_result = pointer_to_net(pointer_to_data)
pointer_to_result
```
And we get the result as usual!
```
pointer_to_result.get()
```
Et voilà! We have seen how to dramatically reduce the communication between the local worker (or server) and the remote devices!
### Switch between workers
One major feature that we want to have is to use the same plan for several workers, that we would change depending on the remote batch of data we are considering.
In particular, we don't want to rebuild the plan each time we change of worker. Let's see how we do this, using the previous example with our small network.
```
class Net(sy.Plan):
def __init__(self):
super(Net, self).__init__()
self.fc1 = nn.Linear(2, 3)
self.fc2 = nn.Linear(3, 2)
def forward(self, x):
x = F.relu(self.fc1(x))
x = self.fc2(x)
return F.log_softmax(x, dim=0)
net = Net()
# Build plan
net.build(torch.tensor([1., 2.]))
```
Here are the main steps we just executed
```
pointer_to_net_1 = net.send(device_1)
pointer_to_data = device_1.search('input_data')[0]
pointer_to_result = pointer_to_net_1(pointer_to_data)
pointer_to_result.get()
```
And actually you can build other PointerPlans from the same plan, so the syntax is the same to run remotely a plan on another device
```
pointer_to_net_2 = net.send(device_2)
pointer_to_data = device_2.search('input_data')[0]
pointer_to_result = pointer_to_net_2(pointer_to_data)
pointer_to_result.get()
```
> Note: Currently, with Plan classes, you can only use a single method and you have to name it "forward".
### Automatically building plans that are functions
For functions (`@` `sy.func2plan`) we can automatically build the plan with no need to explicitly calling `build`, actually in the moment of creation the plan is already built.
To get this functionality the only thing you need to change when creating a plan is setting an argument to the decorator called `args_shape` which should be a list containing the shapes of each argument.
```
@sy.func2plan(args_shape=[(-1, 1)])
def plan_double_abs(x):
x = x + x
x = torch.abs(x)
return x
plan_double_abs.is_built
```
The `args_shape` parameter is used internally to create mock tensors with the given shape which are used to build the plan.
```
@sy.func2plan(args_shape=[(1, 2), (-1, 2)])
def plan_sum_abs(x, y):
s = x + y
return torch.abs(s)
plan_sum_abs.is_built
```
You can also provide state elements to functions!
```
@sy.func2plan(args_shape=[(1,)], state=(torch.tensor([1]), ))
def plan_abs(x, state):
bias, = state.read()
x = x.abs()
return x + bias
pointer_plan = plan_abs.send(device_1)
x_ptr = torch.tensor([-1, 0]).send(device_1)
p = pointer_plan(x_ptr)
p.get()
```
To learn more about this, you can discover how we use Plans with Protocols in Tutorial Part 8 bis!
### Star PySyft on GitHub
The easiest way to help our community is just by starring the repositories! This helps raise awareness of the cool tools we're building.
- [Star PySyft](https://github.com/OpenMined/PySyft)
### Pick our tutorials on GitHub!
We made really nice tutorials to get a better understanding of what Federated and Privacy-Preserving Learning should look like and how we are building the bricks for this to happen.
- [Checkout the PySyft tutorials](https://github.com/OpenMined/PySyft/tree/master/examples/tutorials)
### Join our Slack!
The best way to keep up to date on the latest advancements is to join our community!
- [Join slack.openmined.org](http://slack.openmined.org)
### Join a Code Project!
The best way to contribute to our community is to become a code contributor! If you want to start "one off" mini-projects, you can go to PySyft GitHub Issues page and search for issues marked `Good First Issue`.
- [Good First Issue Tickets](https://github.com/OpenMined/PySyft/issues?q=is%3Aopen+is%3Aissue+label%3A%22good+first+issue%22)
### Donate
If you don't have time to contribute to our codebase, but would still like to lend support, you can also become a Backer on our Open Collective. All donations go toward our web hosting and other community expenses such as hackathons and meetups!
- [Donate through OpenMined's Open Collective Page](https://opencollective.com/openmined)
| github_jupyter |
<a href="https://colab.research.google.com/github/tensorflow/tpu/blob/master/tools/colab/keras_mnist_tpu.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
##### Copyright 2018 The TensorFlow Hub Authors.
Licensed under the Apache License, Version 2.0 (the "License");
```
# Copyright 2018 The TensorFlow Hub Authors. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ==============================================================================
```
## MNIST on TPU or GPU using tf.Keras and tf.data.Dataset
<table><tr><td><img valign="middle" src="https://raw.githubusercontent.com/GoogleCloudPlatform/tensorflow-without-a-phd/master/tensorflow-rl-pong/images/keras-tensorflow-tpu300px.png" width="300" alt="Keras+Tensorflow+Cloud TPU"></td></tr></table>
## Overview
This sample trains an "MNIST" handwritten digit
recognition model on a GPU or TPU backend using a Keras
model. Data are handled using the tf.data.Datset API. This is
a very simple sample provided for educational purposes. Do
not expect outstanding TPU performance on a dataset as
small as MNIST.
This notebook is hosted on GitHub. To view it in its original repository, after opening the notebook, select **File > View on GitHub**.
## Learning objectives
In this notebook, you will learn how to:
* Authenticate in Colab to access Google Cloud Storage (GSC)
* Format and prepare a dataset using tf.data.Dataset
* Create convolutional and dense layers using tf.keras.Sequential
* Build a Keras classifier with softmax, cross-entropy, and the adam optimizer
* Run training and validation in Keras using Cloud TPU
* Export a model for serving from ML Engine
* Deploy a trained model to ML Engine
* Test predictions on a deployed model
## Instructions
<h3><a href="https://cloud.google.com/gpu/"><img valign="middle" src="https://raw.githubusercontent.com/GoogleCloudPlatform/tensorflow-without-a-phd/master/tensorflow-rl-pong/images/gpu-hexagon.png" width="50"></a> Train on GPU or TPU <a href="https://cloud.google.com/tpu/"><img valign="middle" src="https://raw.githubusercontent.com/GoogleCloudPlatform/tensorflow-without-a-phd/master/tensorflow-rl-pong/images/tpu-hexagon.png" width="50"></a></h3>
1. On the main menu, click Runtime and select **Change runtime type**. Set "TPU" as the hardware accelerator.
1. Click Runtime again and select **Runtime > Run All** (Watch out: the "Colab-only auth for this notebook and the TPU" cell requires user input).
<h3><a href="https://cloud.google.com/ml-engine/"><img valign="middle" src="https://raw.githubusercontent.com/GoogleCloudPlatform/tensorflow-without-a-phd/master/tensorflow-rl-pong/images/mlengine-hexagon.png" width="50"></a> Deploy to Cloud Machine Learning (ML) Engine</h3>
* Near the end of this notebook you can deploy your trained model to ML Engine for a serverless, autoscaled, REST API experience. You will need a GCP project and a GCS bucket for this last part.
1. Create a Cloud Storage bucket at http://console.cloud.google.com/storage.
1. Fill the BUCKET parameter in the "Configuration" section below.
1. You can now run the cells under "Deploy the trained model to ML Engine"
TPUs are located in Google Cloud, for optimal performance, they read data directly from Google Cloud Storage (GCS)
## Data, model, and training
```
BATCH_SIZE = 128
training_images_file = 'gs://mnist-public/train-images-idx3-ubyte'
training_labels_file = 'gs://mnist-public/train-labels-idx1-ubyte'
validation_images_file = 'gs://mnist-public/t10k-images-idx3-ubyte'
validation_labels_file = 'gs://mnist-public/t10k-labels-idx1-ubyte'
```
### Imports
```
import os, re, math, json, shutil, pprint
import PIL.Image, PIL.ImageFont, PIL.ImageDraw
import numpy as np
import tensorflow as tf
from matplotlib import pyplot as plt
from tensorflow.python.platform import tf_logging
print("Tensorflow version " + tf.__version__)
```
### Colab-only auth for this notebook and the TPU
```
# backend identification
IS_COLAB_BACKEND = 'COLAB_GPU' in os.environ # this is always set on Colab, the value is 0 or 1 depending on GPU presence
HAS_COLAB_TPU = 'COLAB_TPU_ADDR' in os.environ
HAS_CTPU_TPU = 'TPU_NAME' in os.environ
HAS_MANUAL_TPU = False
# Uncomment the following line to work around the case when TPU_NAME is not set. Please set your vm name manually.
# HAS_MANUAL_TPU, MANUAL_VM_NAME = True, 'MY_VM_NAME'
USE_TPU = HAS_COLAB_TPU or HAS_CTPU_TPU or HAS_MANUAL_TPU
# Auth on Colab (little wrinkle: without auth, Colab will be extremely slow in accessing data from a GCS bucket, even public).
if IS_COLAB_BACKEND:
from google.colab import auth
auth.authenticate_user()
# Also propagate the Colab Auth to TPU so that it can access your GCS buckets, even private ones.
if IS_COLAB_BACKEND and HAS_COLAB_TPU:
with tf.Session('grpc://{}'.format(os.environ['COLAB_TPU_ADDR'])) as sess:
with open('/content/adc.json', 'r') as f:
auth_info = json.load(f) # Upload the credentials to TPU.
tf.contrib.cloud.configure_gcs(sess, credentials=auth_info)
# find the TPU
if IS_COLAB_BACKEND and HAS_COLAB_TPU:
TPU_ADDRESS = 'grpc://{}'.format(os.environ['COLAB_TPU_ADDR'])
elif HAS_CTPU_TPU:
TPU_ADDRESS = os.environ['TPU_NAME']
elif HAS_MANUAL_TPU:
TPU_ADDRESS = MANUAL_VM_NAME
if USE_TPU:
print('Using TPU:', TPU_ADDRESS)
#@title visualization utilities [RUN ME]
"""
This cell contains helper functions used for visualization
and downloads only. You can skip reading it. There is very
little useful Keras/Tensorflow code here.
"""
# Matplotlib config
plt.rc('image', cmap='gray_r')
plt.rc('grid', linewidth=0)
plt.rc('xtick', top=False, bottom=False, labelsize='large')
plt.rc('ytick', left=False, right=False, labelsize='large')
plt.rc('axes', facecolor='F8F8F8', titlesize="large", edgecolor='white')
plt.rc('text', color='a8151a')
plt.rc('figure', facecolor='F0F0F0')# Matplotlib fonts
MATPLOTLIB_FONT_DIR = os.path.join(os.path.dirname(plt.__file__), "mpl-data/fonts/ttf")
# pull a batch from the datasets. This code is not very nice, it gets much better in eager mode (TODO)
def dataset_to_numpy_util(training_dataset, validation_dataset, N):
# get one batch from each: 10000 validation digits, N training digits
unbatched_train_ds = training_dataset.apply(tf.data.experimental.unbatch())
v_images, v_labels = validation_dataset.make_one_shot_iterator().get_next()
t_images, t_labels = unbatched_train_ds.batch(N).make_one_shot_iterator().get_next()
# Run once, get one batch. Session.run returns numpy results
with tf.Session() as ses:
(validation_digits, validation_labels,
training_digits, training_labels) = ses.run([v_images, v_labels, t_images, t_labels])
# these were one-hot encoded in the dataset
validation_labels = np.argmax(validation_labels, axis=1)
training_labels = np.argmax(training_labels, axis=1)
return (training_digits, training_labels,
validation_digits, validation_labels)
# create digits from local fonts for testing
def create_digits_from_local_fonts(n):
font_labels = []
img = PIL.Image.new('LA', (28*n, 28), color = (0,255)) # format 'LA': black in channel 0, alpha in channel 1
font1 = PIL.ImageFont.truetype(os.path.join(MATPLOTLIB_FONT_DIR, 'DejaVuSansMono-Oblique.ttf'), 25)
font2 = PIL.ImageFont.truetype(os.path.join(MATPLOTLIB_FONT_DIR, 'STIXGeneral.ttf'), 25)
d = PIL.ImageDraw.Draw(img)
for i in range(n):
font_labels.append(i%10)
d.text((7+i*28,0 if i<10 else -4), str(i%10), fill=(255,255), font=font1 if i<10 else font2)
font_digits = np.array(img.getdata(), np.float32)[:,0] / 255.0 # black in channel 0, alpha in channel 1 (discarded)
font_digits = np.reshape(np.stack(np.split(np.reshape(font_digits, [28, 28*n]), n, axis=1), axis=0), [n, 28*28])
return font_digits, font_labels
# utility to display a row of digits with their predictions
def display_digits(digits, predictions, labels, title, n):
plt.figure(figsize=(13,3))
digits = np.reshape(digits, [n, 28, 28])
digits = np.swapaxes(digits, 0, 1)
digits = np.reshape(digits, [28, 28*n])
plt.yticks([])
plt.xticks([28*x+14 for x in range(n)], predictions)
for i,t in enumerate(plt.gca().xaxis.get_ticklabels()):
if predictions[i] != labels[i]: t.set_color('red') # bad predictions in red
plt.imshow(digits)
plt.grid(None)
plt.title(title)
# utility to display multiple rows of digits, sorted by unrecognized/recognized status
def display_top_unrecognized(digits, predictions, labels, n, lines):
idx = np.argsort(predictions==labels) # sort order: unrecognized first
for i in range(lines):
display_digits(digits[idx][i*n:(i+1)*n], predictions[idx][i*n:(i+1)*n], labels[idx][i*n:(i+1)*n],
"{} sample validation digits out of {} with bad predictions in red and sorted first".format(n*lines, len(digits)) if i==0 else "", n)
# utility to display training and validation curves
def display_training_curves(training, validation, title, subplot):
if subplot%10==1: # set up the subplots on the first call
plt.subplots(figsize=(10,10), facecolor='#F0F0F0')
plt.tight_layout()
ax = plt.subplot(subplot)
ax.grid(linewidth=1, color='white')
ax.plot(training)
ax.plot(validation)
ax.set_title('model '+ title)
ax.set_ylabel(title)
ax.set_xlabel('epoch')
ax.legend(['train', 'valid.'])
```
### tf.data.Dataset: parse files and prepare training and validation datasets
Please read the [best practices for building](https://www.tensorflow.org/guide/performance/datasets) input pipelines with tf.data.Dataset
```
def read_label(tf_bytestring):
label = tf.decode_raw(tf_bytestring, tf.uint8)
label = tf.reshape(label, [])
label = tf.one_hot(label, 10)
return label
def read_image(tf_bytestring):
image = tf.decode_raw(tf_bytestring, tf.uint8)
image = tf.cast(image, tf.float32)/256.0
image = tf.reshape(image, [28*28])
return image
def load_dataset(image_file, label_file):
imagedataset = tf.data.FixedLengthRecordDataset(image_file, 28*28, header_bytes=16)
imagedataset = imagedataset.map(read_image, num_parallel_calls=16)
labelsdataset = tf.data.FixedLengthRecordDataset(label_file, 1, header_bytes=8)
labelsdataset = labelsdataset.map(read_label, num_parallel_calls=16)
dataset = tf.data.Dataset.zip((imagedataset, labelsdataset))
return dataset
def get_training_dataset(image_file, label_file, batch_size):
dataset = load_dataset(image_file, label_file)
dataset = dataset.cache() # this small dataset can be entirely cached in RAM, for TPU this is important to get good performance from such a small dataset
dataset = dataset.shuffle(5000, reshuffle_each_iteration=True)
dataset = dataset.repeat() # Mandatory for Keras for now
dataset = dataset.batch(batch_size, drop_remainder=True) # drop_remainder is important on TPU, batch size must be fixed
dataset = dataset.prefetch(10) # fetch next batches while training on the current one
return dataset
def get_validation_dataset(image_file, label_file):
dataset = load_dataset(image_file, label_file)
dataset = dataset.cache() # this small dataset can be entirely cached in RAM, for TPU this is important to get good performance from such a small dataset
dataset = dataset.batch(10000, drop_remainder=True) # 10000 items in eval dataset, all in one batch
dataset = dataset.repeat() # Mandatory for Keras for now
return dataset
# instantiate the datasets
training_dataset = get_training_dataset(training_images_file, training_labels_file, BATCH_SIZE)
validation_dataset = get_validation_dataset(validation_images_file, validation_labels_file)
# For TPU, we will need a function that returns the dataset
training_input_fn = lambda: get_training_dataset(training_images_file, training_labels_file, BATCH_SIZE)
validation_input_fn = lambda: get_validation_dataset(validation_images_file, validation_labels_file)
```
### Let's have a look at the data
```
N = 24
(training_digits, training_labels,
validation_digits, validation_labels) = dataset_to_numpy_util(training_dataset, validation_dataset, N)
display_digits(training_digits, training_labels, training_labels, "training digits and their labels", N)
display_digits(validation_digits[:N], validation_labels[:N], validation_labels[:N], "validation digits and their labels", N)
font_digits, font_labels = create_digits_from_local_fonts(N)
```
### Keras model: 3 convolutional layers, 2 dense layers
If you are not sure what cross-entropy, dropout, softmax or batch-normalization mean, head here for a crash-course: [Tensorflow and deep learning without a PhD](https://github.com/GoogleCloudPlatform/tensorflow-without-a-phd/#featured-code-sample)
```
# This model trains to 99.4% sometimes 99.5% accuracy in 10 epochs (with a batch size of 32)
l = tf.keras.layers
model = tf.keras.Sequential(
[
l.Reshape(input_shape=(28*28,), target_shape=(28, 28, 1)),
l.Conv2D(filters=6, kernel_size=3, padding='same', use_bias=False), # no bias necessary before batch norm
l.BatchNormalization(scale=False, center=True), # no batch norm scaling necessary before "relu"
l.Activation('relu'), # activation after batch norm
l.Conv2D(filters=12, kernel_size=6, padding='same', use_bias=False, strides=2),
l.BatchNormalization(scale=False, center=True),
l.Activation('relu'),
l.Conv2D(filters=24, kernel_size=6, padding='same', use_bias=False, strides=2),
l.BatchNormalization(scale=False, center=True),
l.Activation('relu'),
l.Flatten(),
l.Dense(200, use_bias=False),
l.BatchNormalization(scale=False, center=True),
l.Activation('relu'),
l.Dropout(0.5), # Dropout on dense layer only
l.Dense(10, activation='softmax')
])
model.compile(optimizer='adam', # learning rate will be set by LearningRateScheduler
loss='categorical_crossentropy',
metrics=['accuracy'])
# print model layers
model.summary()
# set up learning rate decay
lr_decay = tf.keras.callbacks.LearningRateScheduler(lambda epoch: 0.0001 + 0.02 * math.pow(0.5, 1+epoch), verbose=True)
```
### Train and validate the model
```
EPOCHS = 10
steps_per_epoch = 60000//BATCH_SIZE # 60,000 items in this dataset
trained_model = model
# Counting steps and batches on TPU: the tpu.keras_to_tpu_model API regards the batch size of the input dataset
# as the per-core batch size. The effective batch size is 8x more because Cloud TPUs have 8 cores. It increments
# the step by +8 everytime a global batch (8 per-core batches) is processed. Therefore batch size and steps_per_epoch
# settings can stay as they are for TPU training. The training will just go faster.
# Warning: this might change in the final version of the Keras/TPU API.
if USE_TPU:
TPU_WORKER = 'grpc://' + os.environ['COLAB_TPU_ADDR']
# On a VM created using "ctpu up", you can use TPUClusterResolver(os.environ('TPU_NAME')) # The TPU name is usually the same as the name of your VM, use that if for some reason the TPU_NAME environment variable is not set
strategy = tf.contrib.tpu.TPUDistributionStrategy(tf.contrib.cluster_resolver.TPUClusterResolver(TPU_WORKER))
trained_model = tf.contrib.tpu.keras_to_tpu_model(model, strategy=strategy)
# Work in progress: reading directly from dataset object not yet implemented
# for Keras/TPU. Keras/TPU needs a function that returns a dataset
history = trained_model.fit(training_input_fn, steps_per_epoch=steps_per_epoch, epochs=EPOCHS,
validation_data=validation_input_fn, validation_steps=1, callbacks=[lr_decay])
else:
history = trained_model.fit(training_dataset, steps_per_epoch=steps_per_epoch, epochs=EPOCHS,
validation_data=validation_dataset, validation_steps=1, callbacks=[lr_decay])
```
### Visualize training and validation curves
```
print(history.history.keys())
display_training_curves(history.history['acc'], history.history['val_acc'], 'accuracy', 211)
display_training_curves(history.history['loss'], history.history['val_loss'], 'loss', 212)
```
### Visualize predictions
```
# recognize digits from local fonts
probabilities = trained_model.predict(font_digits, steps=1)
predicted_labels = np.argmax(probabilities, axis=1)
display_digits(font_digits, predicted_labels, font_labels, "predictions from local fonts (bad predictions in red)", N)
# recognize validation digits
probabilities = trained_model.predict(validation_digits, steps=1)
predicted_labels = np.argmax(probabilities, axis=1)
display_top_unrecognized(validation_digits, predicted_labels, validation_labels, N, 7)
```
## Deploy the trained model to ML Engine
Push your trained model to production on ML Engine for a serverless, autoscaled, REST API experience.
You will need a GCS bucket and a GCP project for this.
Models deployed on ML Engine autoscale to zero if not used. There will be no ML Engine charges after you are done testing.
Google Cloud Storage incurs charges. Empty the bucket after deployment if you want to avoid these. Once the model is deployed, the bucket is not useful anymore.
### Configuration
```
PROJECT = "" #@param {type:"string"}
BUCKET = "gs://" #@param {type:"string", default:"jddj"}
NEW_MODEL = True #@param {type:"boolean"}
MODEL_NAME = "colabmnist" #@param {type:"string"}
MODEL_VERSION = "v0" #@param {type:"string"}
assert PROJECT, 'For this part, you need a GCP project. Head to http://console.cloud.google.com/ and create one.'
assert re.search(r'gs://.+', BUCKET), 'For this part, you need a GCS bucket. Head to http://console.cloud.google.com/storage and create one.'
```
### Export the model for serving from ML Engine
```
class ServingInput(tf.keras.layers.Layer):
# the important detail in this boilerplate code is "trainable=False"
def __init__(self, name, dtype, batch_input_shape=None):
super(ServingInput, self).__init__(trainable=False, name=name, dtype=dtype, batch_input_shape=batch_input_shape)
def get_config(self):
return {'batch_input_shape': self._batch_input_shape, 'dtype': self.dtype, 'name': self.name }
def call(self, inputs):
# When the deployed model is called through its REST API,
# the JSON payload is parsed automatically, transformed into
# a tensor and passed to this input layer. You can perform
# additional transformations, such as decoding JPEGs for example,
# before sending the data to your model. However, you can only
# use tf.xxxx operations.
return inputs
# little wrinkle: must copy the model from TPU to CPU manually. This is a temporary workaround.
tf_logging.set_verbosity(tf_logging.INFO)
restored_model = model
restored_model.set_weights(trained_model.get_weights()) # this copied the weights from TPU, does nothing on GPU
tf_logging.set_verbosity(tf_logging.WARN)
# add the serving input layer
serving_model = tf.keras.Sequential()
serving_model.add(ServingInput('serving', tf.float32, (None, 28*28)))
serving_model.add(restored_model)
export_path = tf.contrib.saved_model.save_keras_model(serving_model, os.path.join(BUCKET, 'keras_export')) # export he model to your bucket
export_path = export_path.decode('utf-8')
print("Model exported to: ", export_path)
```
### Deploy the model
This uses the command-line interface. You can do the same thing through the ML Engine UI at https://console.cloud.google.com/mlengine/models
```
# Create the model
if NEW_MODEL:
!gcloud ml-engine models create {MODEL_NAME} --project={PROJECT} --regions=us-central1
# Create a version of this model (you can add --async at the end of the line to make this call non blocking)
# Additional config flags are available: https://cloud.google.com/ml-engine/reference/rest/v1/projects.models.versions
# You can also deploy a model that is stored locally by providing a --staging-bucket=... parameter
!echo "Deployment takes a couple of minutes. You can watch your deployment here: https://console.cloud.google.com/mlengine/models/{MODEL_NAME}"
!gcloud ml-engine versions create {MODEL_VERSION} --model={MODEL_NAME} --origin={export_path} --project={PROJECT} --runtime-version=1.10
```
### Test the deployed model
Your model is now available as a REST API. Let us try to call it. The cells below use the "gcloud ml-engine"
command line tool but any tool that can send a JSON payload to a REST endpoint will work.
```
# prepare digits to send to online prediction endpoint
digits = np.concatenate((font_digits, validation_digits[:100-N]))
labels = np.concatenate((font_labels, validation_labels[:100-N]))
with open("digits.json", "w") as f:
for digit in digits:
# the format for ML Engine online predictions is: one JSON object per line
data = json.dumps({"serving_input": digit.tolist()}) # "serving_input" because the ServingInput layer was named "serving". Keras appends "_input"
print(data, file=f)
# Request online predictions from deployed model (REST API) using the "gcloud ml-engine" command line.
predictions = !gcloud ml-engine predict --model={MODEL_NAME} --json-instances digits.json --project={PROJECT} --version {MODEL_VERSION}
print(predictions)
probabilities = np.stack([json.loads(p) for p in predictions[1:]]) # first line is the name of the input layer: drop it, parse the rest
predictions = np.argmax(probabilities, axis=1)
display_top_unrecognized(digits, predictions, labels, N, 100//N)
```
## What's next
* Learn about [Cloud TPUs](https://cloud.google.com/tpu/docs) that Google designed and optimized specifically to speed up and scale up ML workloads for training and inference and to enable ML engineers and researchers to iterate more quickly.
* Explore the range of [Cloud TPU tutorials and Colabs](https://cloud.google.com/tpu/docs/tutorials) to find other examples that can be used when implementing your ML project.
On Google Cloud Platform, in addition to GPUs and TPUs available on pre-configured [deep learning VMs](https://cloud.google.com/deep-learning-vm/), you will find [AutoML](https://cloud.google.com/automl/)*(beta)* for training custom models without writing code and [Cloud ML Engine](https://cloud.google.com/ml-engine/docs/) which will allows you to run parallel trainings and hyperparameter tuning of your custom models on powerful distributed hardware.
## License
---
author: Martin Gorner<br>
twitter: @martin_gorner
---
Copyright 2018 Google LLC
Licensed under the Apache License, Version 2.0 (the "License");
you may not use this file except in compliance with the License.
You may obtain a copy of the License at
http://www.apache.org/licenses/LICENSE-2.0
Unless required by applicable law or agreed to in writing, software
distributed under the License is distributed on an "AS IS" BASIS,
WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
See the License for the specific language governing permissions and
limitations under the License.
---
This is not an official Google product but sample code provided for an educational purpose
| github_jupyter |
```
%matplotlib inline
```
# Simple 1D Kernel Density Estimation
This example uses the :class:`sklearn.neighbors.KernelDensity` class to
demonstrate the principles of Kernel Density Estimation in one dimension.
The first plot shows one of the problems with using histograms to visualize
the density of points in 1D. Intuitively, a histogram can be thought of as a
scheme in which a unit "block" is stacked above each point on a regular grid.
As the top two panels show, however, the choice of gridding for these blocks
can lead to wildly divergent ideas about the underlying shape of the density
distribution. If we instead center each block on the point it represents, we
get the estimate shown in the bottom left panel. This is a kernel density
estimation with a "top hat" kernel. This idea can be generalized to other
kernel shapes: the bottom-right panel of the first figure shows a Gaussian
kernel density estimate over the same distribution.
Scikit-learn implements efficient kernel density estimation using either
a Ball Tree or KD Tree structure, through the
:class:`sklearn.neighbors.KernelDensity` estimator. The available kernels
are shown in the second figure of this example.
The third figure compares kernel density estimates for a distribution of 100
samples in 1 dimension. Though this example uses 1D distributions, kernel
density estimation is easily and efficiently extensible to higher dimensions
as well.
```
# Author: Jake Vanderplas <jakevdp@cs.washington.edu>
#
import numpy as np
import matplotlib.pyplot as plt
from scipy.stats import norm
from sklearn.neighbors import KernelDensity
#----------------------------------------------------------------------
# Plot the progression of histograms to kernels
np.random.seed(1)
N = 20
X = np.concatenate((np.random.normal(0, 1, int(0.3 * N)),
np.random.normal(5, 1, int(0.7 * N))))[:, np.newaxis]
X_plot = np.linspace(-5, 10, 1000)[:, np.newaxis]
bins = np.linspace(-5, 10, 10)
fig, ax = plt.subplots(2, 2, sharex=True, sharey=True)
fig.subplots_adjust(hspace=0.05, wspace=0.05)
# histogram 1
ax[0, 0].hist(X[:, 0], bins=bins, fc='#AAAAFF', normed=True)
ax[0, 0].text(-3.5, 0.31, "Histogram")
# histogram 2
ax[0, 1].hist(X[:, 0], bins=bins + 0.75, fc='#AAAAFF', normed=True)
ax[0, 1].text(-3.5, 0.31, "Histogram, bins shifted")
# tophat KDE
kde = KernelDensity(kernel='tophat', bandwidth=0.75).fit(X)
log_dens = kde.score_samples(X_plot)
ax[1, 0].fill(X_plot[:, 0], np.exp(log_dens), fc='#AAAAFF')
ax[1, 0].text(-3.5, 0.31, "Tophat Kernel Density")
# Gaussian KDE
kde = KernelDensity(kernel='gaussian', bandwidth=0.75).fit(X)
log_dens = kde.score_samples(X_plot)
ax[1, 1].fill(X_plot[:, 0], np.exp(log_dens), fc='#AAAAFF')
ax[1, 1].text(-3.5, 0.31, "Gaussian Kernel Density")
for axi in ax.ravel():
axi.plot(X[:, 0], np.zeros(X.shape[0]) - 0.01, '+k')
axi.set_xlim(-4, 9)
axi.set_ylim(-0.02, 0.34)
for axi in ax[:, 0]:
axi.set_ylabel('Normalized Density')
for axi in ax[1, :]:
axi.set_xlabel('x')
#----------------------------------------------------------------------
# Plot all available kernels
X_plot = np.linspace(-6, 6, 1000)[:, None]
X_src = np.zeros((1, 1))
fig, ax = plt.subplots(2, 3, sharex=True, sharey=True)
fig.subplots_adjust(left=0.05, right=0.95, hspace=0.05, wspace=0.05)
def format_func(x, loc):
if x == 0:
return '0'
elif x == 1:
return 'h'
elif x == -1:
return '-h'
else:
return '%ih' % x
for i, kernel in enumerate(['gaussian', 'tophat', 'epanechnikov',
'exponential', 'linear', 'cosine']):
axi = ax.ravel()[i]
log_dens = KernelDensity(kernel=kernel).fit(X_src).score_samples(X_plot)
axi.fill(X_plot[:, 0], np.exp(log_dens), '-k', fc='#AAAAFF')
axi.text(-2.6, 0.95, kernel)
axi.xaxis.set_major_formatter(plt.FuncFormatter(format_func))
axi.xaxis.set_major_locator(plt.MultipleLocator(1))
axi.yaxis.set_major_locator(plt.NullLocator())
axi.set_ylim(0, 1.05)
axi.set_xlim(-2.9, 2.9)
ax[0, 1].set_title('Available Kernels')
#----------------------------------------------------------------------
# Plot a 1D density example
N = 100
np.random.seed(1)
X = np.concatenate((np.random.normal(0, 1, int(0.3 * N)),
np.random.normal(5, 1, int(0.7 * N))))[:, np.newaxis]
X_plot = np.linspace(-5, 10, 1000)[:, np.newaxis]
true_dens = (0.3 * norm(0, 1).pdf(X_plot[:, 0])
+ 0.7 * norm(5, 1).pdf(X_plot[:, 0]))
fig, ax = plt.subplots()
ax.fill(X_plot[:, 0], true_dens, fc='black', alpha=0.2,
label='input distribution')
for kernel in ['gaussian', 'tophat', 'epanechnikov']:
kde = KernelDensity(kernel=kernel, bandwidth=0.5).fit(X)
log_dens = kde.score_samples(X_plot)
ax.plot(X_plot[:, 0], np.exp(log_dens), '-',
label="kernel = '{0}'".format(kernel))
ax.text(6, 0.38, "N={0} points".format(N))
ax.legend(loc='upper left')
ax.plot(X[:, 0], -0.005 - 0.01 * np.random.random(X.shape[0]), '+k')
ax.set_xlim(-4, 9)
ax.set_ylim(-0.02, 0.4)
plt.show()
```
| github_jupyter |
```
import qcportal as ptl
import pandas as pd
import datetime
import time
from management import *
from lifecycle import DataSet
# connect without auth
# read only
client = ptl.FractalClient()
```
```
client
dataset = "OpenFF Gen 2 Torsion Set 1 Roche 2"
ds = client.get_collection("TorsionDriveDataset", dataset)
ds.list_specifications()
results = defaultdict(dict)
for spec in ds.list_specifications().index.tolist():
tdrs = get_torsiondrives(ds, spec, client)
for status in ['COMPLETE', 'RUNNING', 'ERROR']:
results[spec][status] = len(
[tdr for tdr in tdrs if tdr.status == status])
df_tdr = pd.DataFrame(results).transpose()
df_tdr.index.name = 'specification'
df_tdr
print(df_tdr.to_markdown())
results = defaultdict(dict)
for spec in ds.list_specifications().index.tolist():
opts = merge(
get_torsiondrive_optimizations(ds, spec, client))
for status in ['COMPLETE', 'INCOMPLETE', 'ERROR']:
results[spec][status] = len(
[opt for opt in opts if opt.status == status])
df_tdr_opt = pd.DataFrame(results).transpose()
df_tdr_opt.index.name = 'specification'
df_tdr_opt
from datetime import datetime
datehr = datetime.utcnow().strftime("%Y-%m-%d %H:%M UTC")
datehr
dataset_type = 'TorsionDriveDataset'
dataset_name = 'OpenFF Gen 2 Torsion Set 1 Roche 2'
meta = {'**Dataset Name**': dataset_name,
'**Dataset Type**': dataset_type,
'**UTC Date**': datehr}
meta = pd.DataFrame(pd.Series(meta, name=""))
comment = f"""
## Error Cycling Report
{meta.to_markdown()}
### `TorsionDriveRecord` current status
{df_tdr.to_markdown()}
### `OptimizationRecord` current status
{df_tdr_opt.to_markdown()}
"""
print(comment)
```
## Error Cycling Report
| --- | --- |
| :------------- | -----------: |
| **Dataset Type** | TorsionDriveDataset |
| **Dataset Name** | OpenFF Gen 2 Torsion Set 1 Roche 2 |
| **UTC Date** | 2020-07-09 23hr UTC |
### `TorsionDriveRecord` current status
| specification | COMPLETE | ERROR | RUNNING |
|:----------------|-----------:|--------:|----------:|
| default | 136 | 1 | 5 |
### `OptimizationRecord` current status
| specification | COMPLETE | ERROR | INCOMPLETE |
|:----------------|-----------:|--------:|-------------:|
| default | 6966 | 4 | 5 |
## DataSet testing
```
from github import Github
from getpass import getpass
ghapi = Github(getpass())
dst = DataSet('../submissions/2020-03-23-OpenFF-Gen-2-Torsion-Set-1-Roche-2', )
```
We'll begin our live testing here.
```
comment = f"""
## Error Cycling Report
{meta.to_markdown()}
### `TorsionDriveRecord` current status
{df_tdr.to_markdown()}
### `OptimizationRecord` current status
{df_tdr_opt.to_markdown()}
"""
print("\n".join([substr.strip() for substr in comment.split('\n')]))
```
| github_jupyter |
# Developing an AI application
Going forward, AI algorithms will be incorporated into more and more everyday applications. For example, you might want to include an image classifier in a smart phone app. To do this, you'd use a deep learning model trained on hundreds of thousands of images as part of the overall application architecture. A large part of software development in the future will be using these types of models as common parts of applications.
In this project, you'll train an image classifier to recognize different species of flowers. You can imagine using something like this in a phone app that tells you the name of the flower your camera is looking at. In practice you'd train this classifier, then export it for use in your application. We'll be using [this dataset](http://www.robots.ox.ac.uk/~vgg/data/flowers/102/index.html) of 102 flower categories, you can see a few examples below.
<img src='assets/Flowers.png' width=500px>
The project is broken down into multiple steps:
* Load and preprocess the image dataset
* Train the image classifier on your dataset
* Use the trained classifier to predict image content
We'll lead you through each part which you'll implement in Python.
When you've completed this project, you'll have an application that can be trained on any set of labeled images. Here your network will be learning about flowers and end up as a command line application. But, what you do with your new skills depends on your imagination and effort in building a dataset. For example, imagine an app where you take a picture of a car, it tells you what the make and model is, then looks up information about it. Go build your own dataset and make something new.
First up is importing the packages you'll need. It's good practice to keep all the imports at the beginning of your code. As you work through this notebook and find you need to import a package, make sure to add the import up here.
Please make sure if you are running this notebook in the workspace that you have chosen GPU rather than CPU mode.
```
# Imports here
import numpy as np
from torchvision import datasets,transforms
import torch
import os
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import pylab
from PIL import Image
from sklearn import preprocessing
from PIL import Image, ImageOps
from torch import nn
from torch import optim
import torch.nn.functional as F
import torchvision
import json
import seaborn as sb
import matplotlib.pyplot as plt
import pandas as pd
from collections import OrderedDict
import time
import copy
import operator
print(torch.__version__)
%matplotlib inline
```
## Load the data
Here you'll use `torchvision` to load the data ([documentation](http://pytorch.org/docs/0.3.0/torchvision/index.html)). The data should be included alongside this notebook, otherwise you can [download it here](https://s3.amazonaws.com/content.udacity-data.com/nd089/flower_data.tar.gz). The dataset is split into three parts, training, validation, and testing. For the training, you'll want to apply transformations such as random scaling, cropping, and flipping. This will help the network generalize leading to better performance. You'll also need to make sure the input data is resized to 224x224 pixels as required by the pre-trained networks.
The validation and testing sets are used to measure the model's performance on data it hasn't seen yet. For this you don't want any scaling or rotation transformations, but you'll need to resize then crop the images to the appropriate size.
The pre-trained networks you'll use were trained on the ImageNet dataset where each color channel was normalized separately. For all three sets you'll need to normalize the means and standard deviations of the images to what the network expects. For the means, it's `[0.485, 0.456, 0.406]` and for the standard deviations `[0.229, 0.224, 0.225]`, calculated from the ImageNet images. These values will shift each color channel to be centered at 0 and range from -1 to 1.
```
data_dir = 'flowers'
train_dir = data_dir + '/train'
valid_dir = data_dir + '/valid'
test_dir = data_dir + '/test'
# TODO: Define your transforms for the training, validation, and testing sets
train_transforms = transforms.Compose([transforms.RandomRotation(30),
transforms.RandomResizedCrop(224),
transforms.RandomHorizontalFlip(),
transforms.ToTensor(),
transforms.Normalize([0.485, 0.456, 0.406],
[0.229, 0.224, 0.225])])
verify_transforms = transforms.Compose([transforms.Resize(256),
transforms.CenterCrop(224),
transforms.ToTensor(),
transforms.Normalize([0.485, 0.456, 0.406],
[0.229, 0.224, 0.225])])
# TODO: Load the datasets with ImageFolder
train_datasets = datasets.ImageFolder(train_dir, transform=train_transforms)
valid_datasets = datasets.ImageFolder(valid_dir, transform=verify_transforms)
test_datasets = datasets.ImageFolder(test_dir, transform=verify_transforms)
# TODO: Using the image datasets and the trainforms, define the dataloaders
trainloader = torch.utils.data.DataLoader(train_datasets, batch_size=64, shuffle=True)
validloader = torch.utils.data.DataLoader(valid_datasets, batch_size=32)
testloader = torch.utils.data.DataLoader(test_datasets, batch_size=32)
# See some examples
# print(train_datasets.samples)
# print(trainloader.sampler.data_source.samples[5000])
# Run this to test your data loader
'''
images, labels = next(iter(testloader))
def show(img):
npimg = img.numpy()
plt.imshow(np.transpose(npimg, (1,2,0)), interpolation='nearest')
print(images[0].shape)
print((images[0]+1)/2)
show((images[0]+1)/2)'''
```
### Label mapping
You'll also need to load in a mapping from category label to category name. You can find this in the file `cat_to_name.json`. It's a JSON object which you can read in with the [`json` module](https://docs.python.org/2/library/json.html). This will give you a dictionary mapping the integer encoded categories to the actual names of the flowers.
```
with open('cat_to_name.json', 'r') as f:
cat_to_name = json.load(f)
newdict={int(i):j for i,j in cat_to_name.items()}
#print(newdict.keys())
sorted_newdict = sorted(newdict.items(), key=operator.itemgetter(0))
#print(sorted_newdict)
#for key, value in sorted(cat_to_name.iteritems(), key = lambda(k,v): (v,k)):
# print "%s: %s" % (key, value)
```
# Building and training the classifier
Now that the data is ready, it's time to build and train the classifier. As usual, you should use one of the pretrained models from `torchvision.models` to get the image features. Build and train a new feed-forward classifier using those features.
We're going to leave this part up to you. Refer to [the rubric](https://review.udacity.com/#!/rubrics/1663/view) for guidance on successfully completing this section. Things you'll need to do:
* Load a [pre-trained network](http://pytorch.org/docs/master/torchvision/models.html) (If you need a starting point, the VGG networks work great and are straightforward to use)
* Define a new, untrained feed-forward network as a classifier, using ReLU activations and dropout
* Train the classifier layers using backpropagation using the pre-trained network to get the features
* Track the loss and accuracy on the validation set to determine the best hyperparameters
We've left a cell open for you below, but use as many as you need. Our advice is to break the problem up into smaller parts you can run separately. Check that each part is doing what you expect, then move on to the next. You'll likely find that as you work through each part, you'll need to go back and modify your previous code. This is totally normal!
When training make sure you're updating only the weights of the feed-forward network. You should be able to get the validation accuracy above 70% if you build everything right. Make sure to try different hyperparameters (learning rate, units in the classifier, epochs, etc) to find the best model. Save those hyperparameters to use as default values in the next part of the project.
One last important tip if you're using the workspace to run your code: To avoid having your workspace disconnect during the long-running tasks in this notebook, please read in the earlier page in this lesson called Intro to GPU Workspaces about Keeping Your Session Active. You'll want to include code from the workspace_utils.py module.
```
# TODO: Build and train your network
model = torchvision.models.vgg16(pretrained=True)
dropout=0.1
# Freeze parameters so we don't backprop through them
for param in model.parameters():
param.requires_grad = False
classifier = nn.Sequential(OrderedDict([
('fc1', nn.Linear(25088, 4096)),
('relu1', nn.ReLU()),
('drop1',nn.Dropout(p=dropout)),
('fc2', nn.Linear(4096, 1024)),
('drop2',nn.Dropout(p=dropout)),
('fc3', nn.Linear(1024, 256)),
('drop3',nn.Dropout(p=dropout)),
('fc4', nn.Linear(256, 102)),
('output', nn.LogSoftmax(dim=1))
]))
model.classifier = classifier
model
#define training var
learning_rate=0.001
criterion = nn.NLLLoss()
optimizer = optim.Adam(model.classifier.parameters(), lr=learning_rate)
scheduler = optim.lr_scheduler.StepLR(optimizer, step_size=4, gamma=0.1)
epochs = 25
print_every=50
# method for validation
def validation(model, validloader, criterion):
valid_loss = 0
accuracy = 0
for images, labels in validloader:
images, labels = images.to('cuda'), labels.to('cuda')
output = model.forward(images)
valid_loss += criterion(output, labels).item()
ps = torch.exp(output)
equality = (labels.data == ps.max(dim=1)[1])
accuracy += equality.type(torch.FloatTensor).mean()
return valid_loss, accuracy
## Train the network
steps=0
model.to('cuda')
for e in range(epochs):
running_loss = 0
model.train()
for images, labels in trainloader:
steps += 1
images, labels = images.to('cuda'), labels.to('cuda')
optimizer.zero_grad()
# Forward and backward passes
outputs = model.forward(images)
loss = criterion(outputs, labels)
loss.backward()
optimizer.step()
running_loss += loss.item()
if steps % print_every == 0:
# change nn to eval mode for inference
model.eval()
# turn off gradients for validation
with torch.no_grad():
valid_loss, valid_accuracy = validation(model, validloader, criterion)
print("Epoch: {}/{}... ".format(e+1, epochs),
"Train Loss: {:.4f}".format(running_loss/print_every),
"Validation Loss: {:.4f}".format(valid_loss/len(validloader)),
"Validation Accuracy: {:.4f}".format(valid_accuracy/len(validloader)))
running_loss = 0
```
def do_deep_learning(model, trainloader, epochs, print_every, criterion, optimizer, device='cpu'):
epochs = epochs
print_every = print_every
steps = 0
# change to cuda
model.to('cuda')
for e in range(epochs):
running_loss = 0
for ii, (inputs, labels) in enumerate(trainloader):
steps += 1
inputs, labels = inputs.to('cuda'), labels.to('cuda')
optimizer.zero_grad()
# Forward and backward passes
outputs = model.forward(inputs)
loss = criterion(outputs, labels)
loss.backward()
optimizer.step()
running_loss += loss.item()
if steps % print_every == 0:
print("Epoch: {}/{}... ".format(e+1, epochs),
"Loss: {:.4f}".format(running_loss/print_every))
running_loss = 0
```
#do_deep_learning(model, trainloader, 3, 40, criterion, optimizer, 'gpu')
#model=train_model(model, criterion, optimizer, scheduler,num_epochs=epochs, device='cuda')
```
## Testing your network
It's good practice to test your trained network on test data, images the network has never seen either in training or validation. This will give you a good estimate for the model's performance on completely new images. Run the test images through the network and measure the accuracy, the same way you did validation. You should be able to reach around 70% accuracy on the test set if the model has been trained well.
```
# TODO: Do validation on the test set
def check_accuracy_on_test(testloader):
correct = 0
total = 0
# model.to('cpu')
model.to('cuda')
# torch.set_default_tensor_type('torch.cuda.FloatTensor')
with torch.no_grad():
for data in testloader:
images, labels = data
outputs = model(images.cuda())
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted.cuda() == labels.cuda()).sum().item()
print('Accuracy of the network on the 10000 test images: %d %%' % (100 * correct / total))
check_accuracy_on_test(testloader)
```
## Save the checkpoint
Now that your network is trained, save the model so you can load it later for making predictions. You probably want to save other things such as the mapping of classes to indices which you get from one of the image datasets: `image_datasets['train'].class_to_idx`. You can attach this to the model as an attribute which makes inference easier later on.
```model.class_to_idx = image_datasets['train'].class_to_idx```
Remember that you'll want to completely rebuild the model later so you can use it for inference. Make sure to include any information you need in the checkpoint. If you want to load the model and keep training, you'll want to save the number of epochs as well as the optimizer state, `optimizer.state_dict`. You'll likely want to use this trained model in the next part of the project, so best to save it now.
```
# TODO: Save the checkpoint
model.class_to_idx = train_datasets.class_to_idx
checkpoint = {'classifier': model.classifier,
'state_dict': model.state_dict(),
'arch':'vgg16',
'class_to_idx': model.class_to_idx}
torch.save(checkpoint, 'checkpoint.pth')
print(model.class_to_idx)
```
## Loading the checkpoint
At this point it's good to write a function that can load a checkpoint and rebuild the model. That way you can come back to this project and keep working on it without having to retrain the network.
```
# TODO: Write a function that loads a checkpoint and rebuilds the model
def load_checkpoint(filepath):
checkpoint=torch.load(filepath)
model=choosemodel(checkpoint['arch'])
model.classifier = checkpoint['classifier']
model.load_state_dict(checkpoint['state_dict'])
model.class_to_idx = checkpoint['class_to_idx']
#newdict=checkpoint['cat_to_name_dict']
#return {'model':model,'newdict':newdict}
return model
import torchvision.models as models
def choosemodel(arch):
if arch=='restnet18':
return models.resnet18(pretrained=True)
elif arch=='alexnet':
return models.alexnet(pretrained=True)
elif arch == 'vgg16':
return models.vgg16(pretrained=True)
elif arch == 'squeezenet':
return models.squeezenet1_0(pretrained=True)
elif arch == 'densenet':
return models.densenet161(pretrained=True)
elif arch == 'inception':
return models.inception_v3(pretrained=True)
else:
return models.vgg16(pretrained=True)
#### load model and newdict
model = load_checkpoint('saved_classifier/checkpoint.pth')#['model','newdict']
print('model class to idx is ',model.class_to_idx)
```
# Inference for classification
Now you'll write a function to use a trained network for inference. That is, you'll pass an image into the network and predict the class of the flower in the image. Write a function called `predict` that takes an image and a model, then returns the top $K$ most likely classes along with the probabilities. It should look like
```python
probs, classes = predict(image_path, model)
print(probs)
print(classes)
> [ 0.01558163 0.01541934 0.01452626 0.01443549 0.01407339]
> ['70', '3', '45', '62', '55']
```
First you'll need to handle processing the input image such that it can be used in your network.
## Image Preprocessing
You'll want to use `PIL` to load the image ([documentation](https://pillow.readthedocs.io/en/latest/reference/Image.html)). It's best to write a function that preprocesses the image so it can be used as input for the model. This function should process the images in the same manner used for training.
First, resize the images where the shortest side is 256 pixels, keeping the aspect ratio. This can be done with the [`thumbnail`](http://pillow.readthedocs.io/en/3.1.x/reference/Image.html#PIL.Image.Image.thumbnail) or [`resize`](http://pillow.readthedocs.io/en/3.1.x/reference/Image.html#PIL.Image.Image.thumbnail) methods. Then you'll need to crop out the center 224x224 portion of the image.
Color channels of images are typically encoded as integers 0-255, but the model expected floats 0-1. You'll need to convert the values. It's easiest with a Numpy array, which you can get from a PIL image like so `np_image = np.array(pil_image)`.
As before, the network expects the images to be normalized in a specific way. For the means, it's `[0.485, 0.456, 0.406]` and for the standard deviations `[0.229, 0.224, 0.225]`. You'll want to subtract the means from each color channel, then divide by the standard deviation.
And finally, PyTorch expects the color channel to be the first dimension but it's the third dimension in the PIL image and Numpy array. You can reorder dimensions using [`ndarray.transpose`](https://docs.scipy.org/doc/numpy-1.13.0/reference/generated/numpy.ndarray.transpose.html). The color channel needs to be first and retain the order of the other two dimensions.
```
def process_image(image):
''' Scales, crops, and normalizes a PIL image for a PyTorch model,
returns an Numpy array
'''
size=(256,256)
pilim=ImageOps.fit(image, size, Image.ANTIALIAS)
width, height = pilim.size # Get dimensions
new_width=224
new_height=224
left = (width - new_width)/2
top = (height - new_height)/2
right = (width + new_width)/2
bottom = (height + new_height)/2
croppedIm=pilim.crop((left, top, right, bottom))
numpyim= np.array(croppedIm)/255
mean = numpyim.mean(axis=(0,1))
std= numpyim.std(axis=(0,1))
nmean = np.array([0.485, 0.456, 0.406])
nstd = np.array([0.229, 0.224, 0.225])
#numpyim = ((numpyim - mean)/std)*nstd+nmean
numpyim = (numpyim - mean)/std
return numpyim.transpose((2,0,1))
# TODO: Process a PIL image for use in a PyTorch model
```
To check your work, the function below converts a PyTorch tensor and displays it in the notebook. If your `process_image` function works, running the output through this function should return the original image (except for the cropped out portions).
```
def imshow(image,df, ax=None, title=None):
if ax is None:
a4_dims = (4.5, 8.27)
fig, (ax,ax1) = plt.subplots(2,1,figsize=a4_dims)
#ax1 = plt.subplots(1,2)
#else:
#fig, axs = plt.subplots(1,2)
#ax1=sb.barplot(data=df,x='category',y='probability')
# PyTorch tensors assume the color channel is the first dimension
# but matplotlib assumes is the third dimension
image = image.transpose((1, 2, 0))
# Undo preprocessing
mean = np.array([0.485, 0.456, 0.406])
std = np.array([0.229, 0.224, 0.225])
image = std * image + mean
# Image needs to be clipped between 0 and 1 or it looks like noise when displayed
image = np.clip(image, 0, 1)
ax.imshow(image)
#sb.set(rc={'figure.figsize':(11.7,8.27)})
sb.barplot(data=df,y='category',x='probability',ax=ax1)
#print(resultdic)
ax.axes.get_xaxis().set_visible(False)
ax.axes.get_yaxis().set_visible(False)
ax.set_title(df['category'].iloc[0])
return ax
```
## Class Prediction
Once you can get images in the correct format, it's time to write a function for making predictions with your model. A common practice is to predict the top 5 or so (usually called top-$K$) most probable classes. You'll want to calculate the class probabilities then find the $K$ largest values.
To get the top $K$ largest values in a tensor use [`x.topk(k)`](http://pytorch.org/docs/master/torch.html#torch.topk). This method returns both the highest `k` probabilities and the indices of those probabilities corresponding to the classes. You need to convert from these indices to the actual class labels using `class_to_idx` which hopefully you added to the model or from an `ImageFolder` you used to load the data ([see here](#Save-the-checkpoint)). Make sure to invert the dictionary so you get a mapping from index to class as well.
Again, this method should take a path to an image and a model checkpoint, then return the probabilities and classes.
```python
probs, classes = predict(image_path, model)
print(probs)
print(classes)
> [ 0.01558163 0.01541934 0.01452626 0.01443549 0.01407339]
> ['70', '3', '45', '62', '55']
```
```
def predict(image_path, model, topk=5):
''' Predict the class (or classes) of an image using a trained deep learning model.
'''
'''
data_transforms = transforms.Compose([transforms.RandomRotation(30),
transforms.RandomResizedCrop(244),
transforms.RandomHorizontalFlip(),
transforms.ToTensor(),
transforms.Normalize([0.5, 0.5, 0.5],
[0.5,0.5,0.5])])
image_datasets = datasets.ImageFolder(image_path, transform=data_transforms)
'''
im=Image.open(image_path)
im=process_image(im)
#image=Image.open(image_path, mode='r')
#print(image)
#image=process_image(image_datasets)
imagetensor=torch.from_numpy(im).float()
#print(imagetensor.shape)
# Add the extra select dimension for unsqueezing
unsqueezedtensor=imagetensor.unsqueeze_(0)
print(unsqueezedtensor.shape)
#get output
model.to('cuda')
try:
output=model(unsqueezedtensor.cuda())
except:
output=model(unsqueezedtensor)
# TODO: Implement the code to predict the class from an image file
idx_to_class = {val: key for key, val in model.class_to_idx.items()}
probability_output,categorynumber=torch.topk(output.data,5,1)
probability=torch.exp(probability_output)
resultdic={'category':[],'probability':[]}
for i,j in zip(categorynumber[0],probability[0]):
category_id=int(idx_to_class[i.item()])
resultdic['category'].append(newdict[category_id])
resultdic['probability'].append(j.item())
df=pd.DataFrame(resultdic)
imshow(im,df)
resultdic['probs'] = df.pop('probability')
resultdic['classes'] = df.pop('category')
return resultdic
```
## Sanity Checking
Now that you can use a trained model for predictions, check to make sure it makes sense. Even if the testing accuracy is high, it's always good to check that there aren't obvious bugs. Use `matplotlib` to plot the probabilities for the top 5 classes as a bar graph, along with the input image. It should look like this:
<img src='assets/inference_example.png' width=300px>
You can convert from the class integer encoding to actual flower names with the `cat_to_name.json` file (should have been loaded earlier in the notebook). To show a PyTorch tensor as an image, use the `imshow` function defined above.
```
# TODO: Display an image along with the top 5 classes
def getimagepath(root_path):
for root,dirs,filenames in os.walk(root_path):
for filename in filenames:
yield(os.path.join(root,filename))
def display(image_path, model, topk=5):
result=predict(image_path,model,topk)
print(result)
#print(newdict[categorynumber.numpy()[0]],probability.numpy()[0])
#print(newdict[categorynumber.item()],probability.item())
image_path=getimagepath('flowers')
# display('flowers/valid/1/image_06739.jpg',model)
display(next(image_path),model)
newdict[1]
```
| github_jupyter |
# Determinental Point Process (DPP)
This notebook demonstrates how to draw random samples from FiniteDPP process defined by a square exponential similarity kernel with different values of lenght_scale and prefactor.
For each combination of lenght_scale and prefactor, we draw a random sample and compute the average and the standard deviation in the number of objects.
```
#!pip install -U neptune-notebooks
%matplotlib inline
import matplotlib.pyplot as plt
import torch
import numpy
from genus.util_ml import Grid_DPP
from genus.util_vis import show_batch
grid_size = 8
lenght_scales = [20, 10, 5, 2, 1]
weights = [0.05, 0.1, 1, 5, 10]
with torch.no_grad():
fig, ax = plt.subplots(ncols=len(weights), nrows=len(lenght_scales), figsize=(8,8))
for row, l in enumerate(lenght_scales):
for col, w in enumerate(weights):
DPP = Grid_DPP(length_scale=l, weight=w)
value = DPP.sample(size=torch.Size([grid_size, grid_size])) # draw a random samples on a grid'
n_sampled = value.sum(dim=(-1,-2)).item()
logp = DPP.log_prob(value=value)
n_av = DPP.n_mean.item()
n_stddev = DPP.n_stddev.item()
print("n_sampled={0:3d}, n_expected={1:.3f} +/- {2:.3f}, l={3}, w={4}".format(n_sampled, n_av, n_stddev, l, w))
ax[row,col].axis('off')
ax[row,col].imshow(value, cmap='gray')
ax[row,col].set_title("l={0},w={1}".format(l,w))
fig.tight_layout()
fig.savefig("./DPP_samples.png")
```
# Choose one setting
You can change the value of lenght_scale and weight till you find the combination which generates the expected number of samples
```
grid_size = 8
DPP = Grid_DPP(length_scale=5, weight=0.3)
value = DPP.sample(size=torch.Size([grid_size, grid_size]))
print("n_avg ={0:.3f}, n_stdev={1:.3f}".format(DPP.n_mean.item(), DPP.n_variance.item()))
```
# Draw many samples for that setting
Now we draw many random sample and compute the empirical density.
Note that due to the DPP repulsion the particle density is enhanced close to the boundaries
```
n_samples=5000
value = DPP.sample(size=torch.Size([n_samples, grid_size, grid_size]))
print("configurations.shape ->",value.shape)
```
Visualize few samples
```
show_batch(value[:20].unsqueeze(-3).float(), n_col=5, pad_value=0.5, n_padding=1,
title="Few random samples", figsize=(12,6))
```
Visualize empirical density and object number distribution
```
print("configurations.shape ->",value.shape)
fig, axes = plt.subplots(ncols=4, nrows=1, figsize=(16, 4))
axes[0].set_title("Average density")
density = axes[0].imshow(value.float().mean(dim=-3))
axes[0].grid(color='white')
axes[0].set_xticks(ticks=torch.arange(value.shape[-1])-0.5)
axes[0].set_yticks(ticks=torch.arange(value.shape[-2])-0.5)
axes[0].set_xticklabels(labels="")
axes[0].set_yticklabels(labels="")
fig.colorbar(density, ax=axes[0])
axes[1].set_title("One random sample")
axes[1].imshow(value[0])
axes[1].grid()
axes[1].set_xticks(ticks=torch.arange(value.shape[-1])-0.5)
axes[1].set_yticks(ticks=torch.arange(value.shape[-2])-0.5)
axes[1].set_xticklabels(labels="")
axes[1].set_yticklabels(labels="")
axes[2].set_title("Another random sample")
axes[2].imshow(value[1])
axes[2].grid()
axes[2].set_xticks(ticks=torch.arange(value.shape[-1])-0.5)
axes[2].set_yticks(ticks=torch.arange(value.shape[-2])-0.5)
axes[2].set_xticklabels(labels="")
axes[2].set_yticklabels(labels="")
# Make Histogram of the number of particles samples from DPP
n_particles = value.sum(dim=(-1,-2))
counts = torch.bincount(n_particles).float()
counts /= counts.sum()
n_avg = DPP.n_mean.item()
n_var = DPP.n_variance.item()
n_stddev = DPP.n_stddev.item()
n_min = n_particles.min().item()
n_max = n_particles.max().item()
print("n_avg={0:.3f}, n_std={1:.3f}".format(n_avg,n_stddev))
print("n_min={0}, n_max={1}".format(n_min,n_max))
x = torch.linspace(n_min,n_max,100)
y = torch.exp(-(x-n_avg)**2/(2*n_var)) / numpy.sqrt(2*numpy.pi*n_var)
axes[3].set_title("Histogram number of particles")
_ = axes[3].bar(torch.arange(counts.shape[0]), counts)
_ = axes[3].plot(x,y,label="Gaussian Fit")
_ = axes[3].set_xticks(ticks=torch.arange(n_max))
_ = axes[3].set_xlabel("number of particles")
_ = axes[3].legend(loc='upper right')
```
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.