text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
# Multilingual Transcription Pipeline
In this notebook, you will:
1. Deploy the trained model to a sagemaker endpoint
2. Create transcription pipeline that feeds model predictions into Amazon Transcribe
### Import libraries and load AWS credentials
```
!pip install -U sagemaker
import sagemaker
from sagemaker.estimator import Estimator
from sagemaker.serializers import JSONSerializer
from sagemaker.predictor import Predictor
import json
import tarfile
import os
import pandas as pd
import boto3
import time
role = sagemaker.get_execution_role()
sess = sagemaker.session.Session()
account_id = boto3.client('sts').get_caller_identity().get('Account')
region = boto3.session.Session().region_name
bucket = sess.default_bucket()
```
Insert your training job ID here
```
training_job_id = '<insert-training-job-id>'
```
### Deploy model
```
image_uri = f'{account_id}.dkr.ecr.{region}.amazonaws.com/spoken-language-detection'
model_path = f's3://{bucket}/models/{training_job_id}/output/model.tar.gz'
model = sagemaker.Model(
image_uri=image_uri,
model_data=model_path,
role=role
)
model.deploy(1, 'ml.m4.xlarge')
predictor = Predictor(model.endpoint_name, serializer=JSONSerializer())
```
### Detect language and run transcription jobs
Download sample audio clips from [Audio Lingua](audio-lingua.eu).
```
!mkdir samples
!wget -O samples/french-sample.wav https://audio-lingua.eu/spip.php?article7143
!wget -O samples/english-sample.wav https://audio-lingua.eu/spip.php?article6968
!wget -O samples/russian-sample.wav https://audio-lingua.eu/spip.php?article7109
!wget -O samples/spanish-sample.wav https://audio-lingua.eu/spip.php?article7103
!wget -O samples/italian-sample.wav https://audio-lingua.eu/spip.php?article7139
```
Show files to be transcribed from "samples" folder. Each file is a different language.
```
files = [os.path.join('samples', f) for f in os.listdir('samples')]
files
```
Loop through the files, upload each to s3, predict the language, and pass the predicted language and audio file to Transcribe. Go to Transcribe in the AWS console to see the transcription jobs. **Make sure that your role has access to Transcribe (attach AmazonTranscribeFullAccess policy to your role)**
```
lang_code_dict = {
'en' : 'en-US',
'es' : 'es-ES',
'it' : 'it-IT',
'fr' : 'fr-FR',
'ru' : 'ru-RU',
'de' : 'de-DE'
}
transcribe = boto3.client('transcribe')
for f in files:
s3_path = sess.upload_data(f, key_prefix='samples')
pred = predictor.predict([s3_path])
pred = json.loads(pred)[0]
print('Detected language : {}'.format(pred))
job_name = f.split('/')[-1].split('.wav')[0]
transcribe.start_transcription_job(
TranscriptionJobName=job_name,
Media={'MediaFileUri': s3_path},
MediaFormat='wav',
LanguageCode=lang_code_dict[pred]
)
```
Be sure to delete the endpoint after evaluation
```
predictor.delete_endpoint()
```
| github_jupyter |
### Import packages
```
library(data.table)
library(Matrix)
library(proxy)
library(Rtsne)
library(data.table)
library(irlba)
library(umap)
library(ggplot2)
```
### Preprocess
`bsub < count_reads_peaks.sh`
```
path = './count_reads_peaks_output/'
files <- list.files(path,pattern = "\\.txt$")
length(files)
#assuming tab separated values with a header
datalist = lapply(files, function(x)fread(paste0(path,x))$V4)
#assuming the same header/columns for all files
datafr = do.call("cbind", datalist)
dim(datafr)
df_regions = read.csv("../../input/combined.sorted.merged.bed",
sep = '\t',header=FALSE,stringsAsFactors=FALSE)
dim(df_regions)
peaknames = paste(df_regions$V1,df_regions$V2,df_regions$V3,sep = "_")
head(peaknames)
head(sapply(strsplit(files,'\\.'),'[', 1))
colnames(datafr) = sapply(strsplit(files,'\\.'),'[', 1)
rownames(datafr) = peaknames
head(datafr)
dim(datafr)
# saveRDS(datafr, file = './datafr.rds')
# datafr = readRDS('./datafr.rds')
run_pca <- function(mat,num_pcs=50,remove_first_PC=FALSE,scale=FALSE,center=FALSE){
set.seed(2019)
mat = as.matrix(mat)
SVD = irlba(mat, num_pcs, num_pcs,scale=scale,center=center)
sk_diag = matrix(0, nrow=num_pcs, ncol=num_pcs)
diag(sk_diag) = SVD$d
if(remove_first_PC){
sk_diag[1,1] = 0
SVD_vd = (sk_diag %*% t(SVD$v))[2:num_pcs,]
}else{
SVD_vd = sk_diag %*% t(SVD$v)
}
return(SVD_vd)
}
elbow_plot <- function(mat,num_pcs=50,scale=FALSE,center=FALSE,title='',width=3,height=3){
set.seed(2019)
mat = data.matrix(mat)
SVD = irlba(mat, num_pcs, num_pcs,scale=scale,center=center)
options(repr.plot.width=width, repr.plot.height=height)
df_plot = data.frame(PC=1:num_pcs, SD=SVD$d);
# print(SVD$d[1:num_pcs])
p <- ggplot(df_plot, aes(x = PC, y = SD)) +
geom_point(col="#cd5c5c",size = 1) +
ggtitle(title)
return(p)
}
filter_peaks <- function (datafr,cutoff = 0.01){
binary_mat = as.matrix((datafr > 0) + 0)
binary_mat = Matrix(binary_mat, sparse = TRUE)
num_cells_ncounted = Matrix::rowSums(binary_mat)
ncounts = binary_mat[num_cells_ncounted >= dim(binary_mat)[2]*cutoff,]
ncounts = ncounts[rowSums(ncounts) > 0,]
options(repr.plot.width=4, repr.plot.height=4)
hist(log10(num_cells_ncounted),main="No. of Cells Each Site is Observed In",breaks=50)
abline(v=log10(min(num_cells_ncounted[num_cells_ncounted >= dim(binary_mat)[2]*cutoff])),lwd=2,col="indianred")
# hist(log10(new_counts),main="Number of Sites Each Cell Uses",breaks=50)
datafr_filtered = datafr[rownames(ncounts),]
return(datafr_filtered)
}
```
### Obtain Feature Matrix
```
start_time <- Sys.time()
metadata <- read.table('../../input/metadata.tsv',
header = TRUE,
stringsAsFactors=FALSE,quote="",row.names=1)
datafr_filtered <- filter_peaks(datafr)
dim(datafr_filtered)
p_elbow_control <- elbow_plot(datafr_filtered,num_pcs = 100, title = 'PCA on the raw count')
p_elbow_control
fm_control = run_pca(datafr_filtered,num_pcs = 50)
dim(fm_control)
fm_control[1:3,1:3]
end_time <- Sys.time()
end_time - start_time
colnames(fm_control) = colnames(datafr)
rownames(fm_control) = paste('PC',1:dim(fm_control)[1])
dim(fm_control)
all(colnames(fm_control) == rownames(metadata))
saveRDS(fm_control, file = '../../output/feature_matrices/FM_Control_buenrostro2018.rds')
```
### Downstream Analysis
```
set.seed(0)
tsne_control = Rtsne(t(fm_control),pca=F)
library(RColorBrewer)
plot.tsne <- function(x, labels,
main="A tSNE visualization",n=20,
pad=0.1, cex=0.65, pch=19, add=FALSE, legend.suffix="",
cex.main=1, cex.legend=1) {
qual_col_pals = brewer.pal.info[brewer.pal.info$category == 'qual',]
col_vector = unlist(mapply(brewer.pal, qual_col_pals$maxcolors, rownames(qual_col_pals)))
layout = x
xylim = range(layout)
xylim = xylim + ((xylim[2]-xylim[1])*pad)*c(-0.5, 0.5)
if (!add) {
par(mar=c(0.2,0.7,1.2,0.7), ps=10)
plot(xylim, xylim, type="n", axes=F, frame=F)
rect(xylim[1], xylim[1], xylim[2], xylim[2], border="#aaaaaa", lwd=0.25)
}
points(layout[,1], layout[,2], col=col_vector[as.integer(labels)],
cex=cex, pch=pch)
mtext(side=3, main, cex=cex.main)
labels.u = unique(labels)
legend.pos = "topright"
legend.text = as.character(labels.u)
if (add) {
legend.pos = "bottomright"
legend.text = paste(as.character(labels.u), legend.suffix)
}
legend(legend.pos, legend=legend.text,
col=col_vector[as.integer(labels.u)],
bty="n", pch=pch, cex=cex.legend)
}
options(repr.plot.width=5, repr.plot.height=5)
plot.tsne(tsne_control$Y,as.factor(metadata[,'label']))
sessionInfo()
save.image(file = 'Control_buenrostro2018.RData')
```
| github_jupyter |
# Conjoint Analysis to understand Customer Preference
```
# Load Packages
from __future__ import division, print_function
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
sns.set()
import statsmodels.api as sm
import statsmodels.formula.api as smf
from patsy.contrasts import Sum
# Load data
conjoint_dat = pd.read_csv('conjoint_data-crisps.csv', delimiter = '\t')
conjoint_dat.head()
# set up sum contrasts for main effects coding as needed for conjoint analysis
# using C(effect,Sum) notation within main effects model specification
main_effects_model = 'ranking ~ C(flavor, Sum) + C(price, Sum)+ C(weight, Sum)+ C(fat_free, Sum)+ C(sodium_free, Sum)+ \
C(gluten_free, Sum)+ C(organic, Sum)+ C(multipack, Sum)'
main_effects_model_fit = smf.ols(main_effects_model, data = conjoint_dat).fit()
print(main_effects_model_fit.summary())
conjoint_attributes = ['flavor', 'price', 'weight', 'fat_free', 'sodium_free', 'gluten_free','organic', 'multipack']
# Build Part worth information one attribute at a time
level_name = []
part_worth = []
part_worth_range = []
important_levels = {}
end = 1 # Initialize index for coefficient in params
for item in conjoint_attributes:
nlevels = len(list(np.unique(conjoint_dat[item])))
level_name.append(list(np.unique(conjoint_dat[item])))
begin = end
end = begin + nlevels -1
new_part_worth = list(main_effects_model_fit.params[begin:end])
new_part_worth.append((-1)*sum(new_part_worth))
important_levels[item] = np.argmax(new_part_worth)
part_worth.append(new_part_worth)
part_worth_range.append(max(new_part_worth) - min(new_part_worth))
# next iteration
# Values in each of the attributes
level_name
[['cheese&onion', 'chicken_flavored', 'salt&vinegar', 'tomato_ketchup'],
['$1.00', '$2.00', '$3.00', '$4.00'],
['100g', '200g', '300g', '400g'],
['fat_free NO', 'fat_free YES'],
['low_sodium NO', 'low_sodium YES'],
['gluten_free NO', 'gluten_free YES'],
['organic NO', 'organic YES'],
['multipack NO', 'multipack YES']]
# Compute attribute relative importance values from ranges
attribute_importance = []
for item in part_worth_range:
attribute_importance.append(round(100*(item/sum(part_worth_range)),2))
# report conjoint measures to console
labels_pw = []
data_pw = []
part_worth_dict = {}
index = 0 # initialize for use in for-loop
for item in conjoint_attributes:
print('\nAttribute:', item)
print(' Importance:', attribute_importance[index])
print(' Level Part-Worths')
for level in range(len(level_name[index])):
print(' ',level_name[index][level], part_worth[index][level])
labels_pw.append(item+'_'+str(attribute_importance[index])+'__'+level_name[index][level]+
'_'+str(round(part_worth[index][level],3)))
data_pw.append(part_worth[index][level])
part_worth_dict[level_name[index][level]] = part_worth[index][level]
index = index + 1
plt.rcdefaults()
fig, ax = plt.subplots(figsize = (12,4))
# Example data
ax.bar(conjoint_attributes, attribute_importance,width = 0.5, align='center',color='#007acc')
ax.set_xlabel('Attributes')
ax.set_ylabel('Attribute Importance')
ax.set_title('Attribute Importance chart')
plt.tight_layout()
plt.show()
plt.rcdefaults()
fig, ax = plt.subplots()
# Example data
y_pos = np.arange(len(labels_pw))
ax.barh(y_pos, data_pw, height =0.1, align='center',color='#007acc')
plt.plot(data_pw, labels_pw, "o", markersize=5, color='#007acc', alpha=0.6)
#plt.hlines(y=labels, xmin=0, xmax=data, color='#007acc', alpha=0.2, linewidth=5)
ax.set_yticks(y_pos)
ax.set_yticklabels(labels_pw)
ax.invert_yaxis() # labels read top-to-bottom
ax.set_xlabel('Standardized Part-Worth ')
ax.set_title('Conjoint_spine chart')
ax.set_xlim(-12,12)
ax.axvline(0,)
plt.show()
conjoint_dat.columns[:-2]
# utility calculation
utility = []
for i in range(conjoint_dat.shape[0]):
score = part_worth_dict[conjoint_dat['flavor'][i]]+ part_worth_dict[conjoint_dat['price'][i]]+\
part_worth_dict[conjoint_dat['weight'][i]]+part_worth_dict[conjoint_dat['fat_free'][i]]+\
part_worth_dict[conjoint_dat['sodium_free'][i]]+part_worth_dict[conjoint_dat['gluten_free'][i]]+\
part_worth_dict[conjoint_dat['organic'][i]]+part_worth_dict[conjoint_dat['multipack'][i]]
utility.append(score)
conjoint_dat['utility'] = utility
plt.bar(range(len(utility)),utility)
plt.xlabel('row_number_of_entry')
plt.ylabel('Utility')
plt.title('Utility score for the given combinations')
# Max utility
print("The profile that has the highest utility score :",'\n', conjoint_dat.iloc[np.argmax(utility)])
# Optimal utility combination - This is the combination of max values from partworth levels
utility_optimal_solution = 0
i = 0
for key in important_levels.keys():
print("The preferred", key, "is", level_name[i][important_levels[key]])
utility_optimal_solution += part_worth_dict[level_name[i][important_levels[key]]]
i+=1
print('-------------------------------------------')
print("Utility_Score_of_the_optimal_combination is ", round(utility_optimal_solution,4))
print("The current highest utility score in the given combination", round(max(utility),4))
print("The new combination brings in additional",round(utility_optimal_solution-max(utility),3),"units of utility.")
# Predicted Market Share through logit model
total_utility=0
c= 0.8
for item in utility:
total_utility = total_utility + np.exp(c*item)
market_shares =[]
row_number = 0
for item in utility:
probability = np.exp(c*item)/total_utility
market_shares.append(probability)
print ('Market share of profile %s is %s ' % (row_number,probability*100))
row_number +=1
plt.figure(figsize = (12,4))
plt.bar(range(len(market_shares)),market_shares)
plt.xlabel('Profile_number')
plt.ylabel('Market Share')
plt.title('Predicted Market Share for the profiles')
```
| github_jupyter |
```
%matplotlib inline
import numpy as np
import tensorflow as tf
import matplotlib.pyplot as plt
from tensorflow.keras import models, layers
from sklearn.metrics import confusion_matrix
from tensorflow.keras.preprocessing.image import ImageDataGenerator
```
# Training for Eyes
```
import os
os.listdir("eye-data/train")
eye_class_names = ["closed", "open"]
train_path = "eye-data/train"
valid_path = "eye-data/valid"
test_path = "eye-data/test"
data_gen = ImageDataGenerator(preprocessing_function=tf.keras.applications.vgg16.preprocess_input)
train_data = ImageDataGenerator(preprocessing_function=tf.keras.applications.vgg16.preprocess_input, shear_range=0.2, zoom_range=0.2, horizontal_flip=True).flow_from_directory(directory=train_path, target_size=(100, 100), classes=eye_class_names, batch_size=5)
valid_data = data_gen.flow_from_directory(directory=valid_path, target_size=(100, 100), classes=eye_class_names, batch_size=5)
test_data = data_gen.flow_from_directory(directory=test_path, target_size=(100, 100), classes=eye_class_names, batch_size=5, shuffle=False)
imgs, labels = next(train_data)
for i in range(5):
plt.subplot(5, 1, i+1)
plt.axis("off")
plt.imshow(imgs[i], cmap=plt.cm.binary)
print(labels)
model = models.Sequential()
model.add(layers.Conv2D(filters=32, kernel_size=(3, 3), activation='relu', padding='same', input_shape=(100, 100, 3)))
model.add(layers.MaxPool2D(pool_size=(2, 2), strides=2))
model.add(layers.Conv2D(filters=64, kernel_size=(3, 3), activation='relu', padding='same'))
model.add(layers.MaxPool2D(pool_size=(2, 2), strides=2))
model.add(layers.Flatten())
model.add(layers.Dropout(0.2))
model.add(layers.Dense(units=32, activation='relu', kernel_regularizer='l2', bias_regularizer='l2'))
model.add(layers.Dropout(0.2))
model.add(layers.Dense(units=16, activation='relu', kernel_regularizer='l2', bias_regularizer='l2'))
model.add(layers.Dense(units=2, activation='softmax'))
model.summary()
K = tf.keras.backend
K.clear_session()
tf.random.set_seed(10)
early_stopping_cb = tf.keras.callbacks.EarlyStopping(patience=10) # stop if no improvement in 10 consecutive epochs
checkpoint_cb = tf.keras.callbacks.ModelCheckpoint("eye-model.h5", save_best_only=True) # save the best outcome so far
model.compile(loss='categorical_crossentropy', optimizer=tf.keras.optimizers.Adam(learning_rate=0.001), metrics=['accuracy'])
history = model.fit(x=train_data, validation_data=valid_data, epochs=100, verbose=2, callbacks=[early_stopping_cb, checkpoint_cb])
predictions = model.predict(x=test_data, verbose=0)
np.round(predictions)
accuracy = history.history['accuracy']
val_accuracy = history.history['val_accuracy']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(len(accuracy)) # Get number of epochs
# Plot training and validation accuracy per epoch
plt.plot(epochs, accuracy, 'r', label="Training Accuracy")
plt.plot(epochs, val_accuracy, 'b', label="Validation Accuracy")
plt.legend()
plt.title('Training and Validation Accuracy')
plt.show()
# Plot training and validation loss per epoch
plt.plot(epochs, loss, 'r', label="Training Loss")
plt.plot(epochs, val_loss, 'b', label="Validation Loss")
plt.legend()
plt.title('Training and Validation Loss')
plt.show()
cm = confusion_matrix(y_true=test_data.classes, y_pred=np.argmax(predictions, axis=-1))
def plot_confusion_matrix(cm, classes, normalize=False, title='Confusion matrix', cmap=plt.cm.winter):
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
print("Normalized confusion matrix")
else:
print('Confusion matrix, without normalization')
print(cm)
thresh = cm.max() / 2.
for i in range (cm.shape[0]):
for j in range (cm.shape[1]):
plt.text(j, i, cm[i, j],
horizontalalignment="center",
color="red" if cm[i, j] > thresh else "white")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
plot_confusion_matrix(cm=cm, classes=eye_class_names, title="Confusion Matrix")
# model.save("/content/drive/MyDrive/Colab Notebooks/eye-model.h5")
```
# Training for Yawns
```
yawn_class_names = ["no_yawn", "yawn"]
train_path = "yawn-data/train"
valid_path = "yawn-data/valid"
test_path = "yawn-data/test"
data_gen = ImageDataGenerator(preprocessing_function=tf.keras.applications.vgg16.preprocess_input)
train_data = ImageDataGenerator(preprocessing_function=tf.keras.applications.vgg16.preprocess_input, shear_range=0.2, zoom_range=0.2, horizontal_flip=True).flow_from_directory(directory=train_path, target_size=(400, 600), classes=yawn_class_names, batch_size=10)
valid_data = data_gen.flow_from_directory(directory=valid_path, target_size=(400, 600), classes=yawn_class_names, batch_size=10)
test_data = data_gen.flow_from_directory(directory=test_path, target_size=(400, 600), classes=yawn_class_names, batch_size=10, shuffle=False)
imgs, labels = next(train_data)
for i in range(5):
plt.subplot(5, 1, i+1)
plt.axis("off")
plt.imshow(imgs[i], cmap=plt.cm.binary)
print(labels)
model = models.Sequential()
model.add(layers.Conv2D(filters=16, kernel_size=(3, 3), activation='relu', padding='same', input_shape=(400, 600, 3)))
model.add(layers.MaxPool2D(pool_size=(2, 2), strides=2))
model.add(layers.Conv2D(filters=32, kernel_size=(3, 3), activation='relu', padding='same'))
model.add(layers.MaxPool2D(pool_size=(2, 2), strides=2))
model.add(layers.Conv2D(filters=32, kernel_size=(3, 3), activation='relu', padding='same'))
model.add(layers.MaxPool2D(pool_size=(2, 2), strides=2))
model.add(layers.Flatten())
# model.add(layers.Dropout(0.2))
model.add(layers.Dense(units=32, activation='relu', kernel_regularizer='l2', bias_regularizer='l2'))
model.add(layers.Dropout(0.2))
model.add(layers.Dense(units=16, activation='relu', kernel_regularizer='l2', bias_regularizer='l2'))
model.add(layers.Dense(units=2, activation='softmax'))
model.summary()
# tf.keras.utils.plot_model(model, "yawn_model.png", show_shapes=True)
K = tf.keras.backend
K.clear_session()
tf.random.set_seed(10)
early_stopping_cb = tf.keras.callbacks.EarlyStopping(patience=10) # stop if no improvement in 10 consecutive epochs
checkpoint_cb = tf.keras.callbacks.ModelCheckpoint("yawn-model.h5", save_best_only=True) # save the best outcome so far
model.compile(loss='categorical_crossentropy', optimizer=tf.keras.optimizers.Adam(learning_rate=0.001), metrics=['accuracy'])
history = model.fit(x=train_data, validation_data=valid_data, epochs=100, verbose=2, callbacks=[early_stopping_cb, checkpoint_cb])
predictions = model.predict(x=test_data, verbose=0)
np.round(predictions)
accuracy = history.history['accuracy']
val_accuracy = history.history['val_accuracy']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(len(accuracy)) # Get number of epochs
# Plot training and validation accuracy per epoch
plt.plot(epochs, accuracy, 'r', label="Training Accuracy")
plt.plot(epochs, val_accuracy, 'b', label="Validation Accuracy")
plt.legend()
plt.title('Training and Validation Accuracy')
plt.show()
# Plot training and validation loss per epoch
plt.plot(epochs, loss, 'r', label="Training Loss")
plt.plot(epochs, val_loss, 'b', label="Validation Loss")
plt.legend()
plt.title('Training and Validation Loss')
plt.show()
plot_confusion_matrix(cm=cm, classes=yawn_class_names, title="Confusion Matrix")
# model.save("/content/drive/MyDrive/Colab Notebooks/yawn-model.h5")
```
| github_jupyter |
# Bayesian optimisation implementation
> Walkthrough on implementing a modular bayesian optimization routine
- toc:true
- badges: true
- comments: true
- author: Pushkar G. Ghanekar
- categories: [python, machine-learning]
If $f$ (objective function) is cheap to evaluate we can sample various points and built a potential surface however, if the $f$ is expensive -- like in case of first-principles electronic structure calculations, it is important to minimize the number of $f$ calls and number of samples drawn from this evaluation. In that case, if an exact functional form for f is not available (that is, f behaves as a “black box”), what can we do?
Bayesian optimization proceeds by maintaining a probabilistic belief about $f$ and designing a so called **_acquisition function_** to determine where to evaluate the next function call. Bayesian optimization is particularly well-suited to global optimization problems where:
1. $f$ is an expensive black-box function
2. Analytical solution for the gradient of the function is difficult to evaluate
The idea is the find "global" minimum with least number of steps. Incorporating _prior_ beliefs about the underlying process and update the _prior_ with samples draw from the model to better estimate the _posterior_.
Model used for approximating the objective function is called the **_surrogate model_**.
Following are few links I have found useful in understanding the inner workings of the Bayesian opitmization and certain typical surrogate functions used in it:
* Good introductory write-up on Bayesian optimization [(Distill Blog)](https://distill.pub/2020/bayesian-optimization/)
* Nice lecture explaining the working of Gaussian Processes [here](https://www.youtube.com/watch?v=92-98SYOdlY&t=4827s)
### Surrogate model
A popular surrogate model applied for Bayesian optimization, although strictly not required, are Gaussian Processes (GPs). These are used to define a prior beliefs about the objective function. The GP posterior is cheap to evaluate and is used to propose points in the search space where sampling is likely to yield an improvement. Herein, we could substitute this for a ANNs or other surrogate models.
### Acquisition functions
Used to propose sampling points in the search space. Have to consider the trade-off between exploitation vs exploration.
* Exploitation == sampling where objective function value is high
* Exploration == where uncertainty is high
Both correspond to high `acquisition function` value. The goal is the maximize the acquisition value to determine next sampling point.
**Popular acquisition functions:**
1. Maximum probability of improvement
>PI involves sampling for points which improve on the current best objective function value. The point in the sample space with the highest probability of improvement, based on the value predicted by the surrogate function, is chosen as the next point for evaluating through the expensive method. However in this searching scheme we look only at the probability improvement and not the extent of improvement. This might lead it to get stuck in a local minima. Instead we can turn to the __Expected value__ of improvement wherein we consider the extent of improvement as well.
2. Expected improvement (EI)
3. Upper confidence bound (UCB)
### Optimization strategy
Following strategy is followed when optimizing using Bayesian optimization:
<ul>
<li>Find the next sampling point $\mathbf{x}_t$ by optimizing the acquisition function over a surrogate model (in this case a GP) fit over a distribution $\mathcal{D}_{1:t-1}$</li>
<li>Evaluate $f$ at $f(x_{t})$ i.e. sample $f(x_{t})$ from the $f$</li>
<li>Add the new point to the prior of the GP now $\mathcal{D}_{1:t} = ( \mathcal{D}_{1:t-1}, (x_{t},f(x_{t})) )$</li>
</ul>
### Expected improvement
Expected improvement is defined as:
$$\mathrm{EI}(\mathbf{x}) = \max((f(\mathbf{x}) - f(\mathbf{x}^+), 0))\tag{1}$$
where $f(\mathbf{x}^+)$ is the value of the best sample so far and $\mathbf{x}^+$ is the location of that sample i.e. $\mathbf{x}^+ = \mathrm{argmax}_{\mathbf{x}_i \in \mathbf{x}_{1:t}} f(\mathbf{x}_i)$. The expected improvement can be evaluated analytically under the GP model
Expected improvement can be evaluated analytically for a GP model.
Before diving in the technical details, I would like to acknowledge the work of Noah Wichrowski (JHU) in building the necessary ground work for the bayesian optimization routine implemented in this article.
### Implementation with Numpy and Scipy
```
import matplotlib.pyplot as plt
import numpy as np
%config InlineBackend.figure_format = 'retina'
# Plot matplotlib plots with white background:
%config InlineBackend.print_figure_kwargs={'facecolor' : "w"}
from sklearn.gaussian_process import GaussianProcessRegressor as GPR
from sklearn.gaussian_process import kernels
from Bayesian_optimization import plotting_utils, acquisition, objectives, opti
plot_params = {
'font.size' : 22,
'axes.titlesize' : 24,
'axes.labelsize' : 20,
'axes.labelweight' : 'bold',
'xtick.labelsize' : 16,
'ytick.labelsize' : 16,
}
plt.rcParams.update(plot_params)
```
## ## Import the acquisition functions implemented
```
EI = acquisition.ExpectedImprovement(delta = 0.01)
LCB = acquisition.LowerConfidenceBound(sigma = 1.96)
```
# A One-Dimensional Example
### Egg-carton objective function
```
objective = objectives.egg_carton
(low, high) = (0.0, 10.0)
domain = np.array([[low], [high]])
x_pts = np.linspace(low, high, 1000).reshape(-1, 1)
y_pts = objective(x_pts)
num_sample_points = 10
noise_ = 0.1
generator = np.random.default_rng(42)
x_sample = generator.uniform(low, high, size = (num_sample_points, 1))
y_sample = objective(x_sample, noise_)
fig, ax = plt.subplots(1,1, figsize=(8,3))
ax.plot(x_pts, y_pts, 'k-', linewidth=2.0, label='Ground truth')
ax.plot(x_sample, y_sample, 'ro', label='Noisy sampled points')
ax.set_xlabel(r"$x$", fontsize = 20)
ax.set_ylabel(r"$f(x)$", fontsize = 20)
ax.set_title("Initial setup", fontsize = 20)
ax.legend(fontsize = 15)
ax.grid(True)
```
## Fit a GPR model (surrogate function) to the sampled points
```
constant = kernels.ConstantKernel()
matern = kernels.Matern(nu = 2.5)
rbf = kernels.RBF()
gpr_model = GPR(kernel = constant*rbf, alpha = 1e-3, n_restarts_optimizer = 20, normalize_y = False, random_state = 42)
gpr_model.fit(x_sample, y_sample)
(mean_pred, stddev_pred) = gpr_model.predict(x_pts, return_std = True)
gpr_model.kernel_
```
## Plot the Initial Sample
```
(fig_ec, ax_ec) = plotting_utils.illustrate_1d_gpr(objective, gpr_model, x_pts, EI, LCB)
```
## Run a Few Iterations and Assess
```
pkwargs = {"num_sample": 10,
"num_improve": 5,
"generator": generator}
res_ec, _ = opti.bayesian_optimization(objective, gpr_model, LCB, domain, max_iter=10, noise=noise_, prop_kwargs = pkwargs)
gpr_model.fit(res_ec["X"], res_ec["y"]) # Incorporate final point into plots.
(fig_ec, ax_ec) = plotting_utils.illustrate_1d_gpr(objective, gpr_model, x_pts, EI, LCB, num_sample_points)
```
## Run a Few More Iterations
```
res_ec, _ = opti.bayesian_optimization(objective, gpr_model, LCB, domain, noise=noise_, prop_kwargs = pkwargs)
gpr_model.fit(res_ec["X"], res_ec["y"]) # Incorporate final point into plots.
(fig_ec, ax_ec) = plotting_utils.illustrate_1d_gpr(objective, gpr_model, x_pts, EI, LCB, num_sample_points)
```
In total the noisy estimation of the ground-truth is conducted on 30 additional points. It is evident from the plot that most of those points are near the x = (4,6) since that is the minimum value region for the function.
| github_jupyter |
# Reading data
```
import pandas as pd
import numpy as np
import seaborn as sns
import pickle
from sklearn.metrics import accuracy_score
from sklearn.utils import shuffle
from sklearn.neural_network import MLPClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.linear_model import LogisticRegressionCV
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import GridSearchCV, train_test_split, cross_val_score
data = pd.read_csv("../data/Emotion_Data_v2.csv")
emotions = {'neutral': 0,
'calm': 1,
'happy': 2,
'sad': 3,
'angry': 4,
'fearful': 5,
'disgust': 6,
'surprised': 7,
'boredom': 8}
data["Emotion"] = data["Emotion"].map(emotions)
names = data["File Name"]
data = data.drop("File Name", axis = 1)
data
y = data["Emotion"]
data = data.drop("Emotion", axis = 1)
X_train, X_test, y_train, y_test = train_test_split(data, y, test_size = 0.3)
X_train.shape, X_test.shape, y_train.shape, y_test.shape
```
#### Judging by the TSNE Random Forest and Logistic Regression may work nice. Also, let's check KNN
## KNN
```
knn = KNeighborsClassifier(n_neighbors = 103, n_jobs = -1)
# params = {
# "n_neighbors": np.arange(15, 151, 2)
# }
# knn = GridSearchCV(knn, params, n_jobs = -1, verbose = 3)
knn.fit(X_train, y_train)
# knn.best_params_
accuracy_score(knn.predict(X_test), y_test)
```
##### Not bad for KNN, but think that other models will work better
## Random Forest
```
rf = RandomForestClassifier(n_estimators = 201, oob_score = True, n_jobs = -1, random_state = 11, \
max_depth = 15, verbose = 3)
# params = {
# "n_estimators": [101, 201, 301],
# "max_depth": np.arange(12, 20),
# "min_samples_split": np.arange(5, 9)
# "min_samples_leaf": np.arange(3, 7),
# }
#
# rf = GridSearchCV(rf, params, n_jobs = -1, verbose = 3)
cross_val_score(rf, X_train, y_train)
rf.fit(X_train, y_train)
np.arange(5, 12)
# rf.best_params_
accuracy_score(rf.predict(X_test), y_test)
coefs = pd.DataFrame({"feature": X_train.columns,
"coef": rf.feature_importances_}).sort_values(by = "coef", ascending = False)
coefs
sns.scatterplot(x = coefs.index, y = coefs["coef"]);
```
##### We see kinda uniform distribution, but all coefs are near to 0. That's nice, but accuracy is not the best
## Logistic Regression without scaling
```
logit = LogisticRegressionCV(Cs = [10, 5, 1, 0.1, 0.01, 0.001], class_weight = "balanced", \
random_state = 11, verbose = 3, n_jobs = -1)
logit.fit(X_train, y_train)
logit.C_
accuracy_score(logit.predict(X_test), y_test)
coefs = pd.DataFrame({"feature": X_train.columns,
"coef": logit.coef_[0]}).sort_values(by = "coef", ascending = False)
coefs
sns.scatterplot(x = coefs.index, y = coefs["coef"]);
```
##### 0s. Need to use scaling
## Logistic Regression with scaling
```
from sklearn.preprocessing import StandardScaler
# logit = LogisticRegressionCV(class_weight = "balanced", random_state = 11, verbose = 3, n_jobs = -1)
# params = {
# "C": [10, 5, 1, 0.1, 0.01, 0.001]
# }
# logit = GridSearchCV(logit, params, n_jobs = -1, verbose = 3)
logit = LogisticRegressionCV(Cs = [10, 5, 1, 0.1, 0.01, 0.001], class_weight = "balanced", \
random_state = 11, verbose = 3, n_jobs = -1)
logit.fit(StandardScaler().fit_transform(X_train), y_train)
accuracy_score(logit.predict(StandardScaler().fit_transform(X_test)), y_test)
coefs = pd.DataFrame({"feature": X_train.columns,
"coef": logit.coef_[0]}).sort_values(by = "coef", ascending = False)
coefs
sns.scatterplot(x = coefs.index, y = coefs["coef"]);
```
##### Feature importance is lower, better to use Random Forest
# MLP
```
# net = MLPClassifier(hidden_layer_sizes = tuple([50] * 10), alpha = 0.01, learning_rate = "constant", \
# random_state = 11, verbose = 10)
# params = {
# "alpha": [0.0001, 0.001, 0.01, 0.1, 1]
# }
# net = GridSearchCV(net, params, n_jobs = -1, verbose = 10)
# net.fit(X_train, y_train)
# accuracy_score(net.predict(X_test), y_test)
# pd.DataFrame({"feature": X_train.columns,
# "coef": np.mean(net.best_estimator_.coefs_[0])})
```
# SVM
```
from sklearn.svm import SVC
scaler = StandardScaler()
X_train_scaled = scaler.fit_transform(X_train)
X_test = scaler.transform(X_test)
svc = SVC(verbose = 5, random_state = 11)
params = {"C": np.arange(8.5, 9.5, 0.1)}
svc = GridSearchCV(svc, params, n_jobs = -1)
svc.fit(X_train_scaled, y_train)
svc.best_params_
accuracy_score(svc.predict(X_test), y_test)
```
| github_jupyter |
```
#Import dependencies
import numpy as np
import cv2
import matplotlib.image as mpimg
import os
import pandas as pd
import matplotlib.pyplot as plt
# Image common params
i_height, i_width, i_channels = 66,200,3
img_shape = (i_height, i_width, i_channels)
# Dataset common params
data_dir ='./data/data/'
meta_file = data_dir +'driving_log.csv'
#!ls data/data/IMG/ | wc -l
# Load an image
def load_image(img_path):
"""
Load RGB image
"""
return mpimg.imread(os.path.join(data_dir, img_path.strip()))
# Test load_image
meta_df = pd.read_csv(meta_file)
meta_df.head()
def visualize_img(img):
"""
This is an utility to visualize a sample image
"""
plt.imshow(img)
plt.show()
# get a sample image path
image_path_center = meta_df['center'].values[0]
img = load_image(image_path_center)
print("Shape of the image: ", img.shape)
visualize_img(img)
image_path_left = meta_df['left'].values[0]
img = load_image(image_path_left)
visualize_img(img)
image_path_right = meta_df['right'].values[0]
img = load_image(image_path_right)
visualize_img(img)
```
### Data Preprocessing
```
# Crop images to extract required road sections and to remove sky from the road
def crop_image(in_img):
"""
This is used to cropping images
"""
return in_img[60:-25,:,:]
# test crop images
image_path_right = meta_df['right'].values[0]
img = load_image(image_path_right)
croped = crop_image(img)
#visualize
visualize_img(croped)
# resize the images
def resize_image(in_img):
"""
This is an utility function to resize images
"""
return cv2.resize(in_img, (i_width, i_height), cv2.INTER_AREA)
# Test resize images
resized = resize_image(croped)
#visualize
visualize_img(resized)
# convert RGB to YUV image
def convert_rgb2yuv(in_img):
"""
This is an utility function to convert RGB images to YUV.
This technique was intr by NVIDIA for their image pracessing pipeline
"""
return cv2.cvtColor(in_img, cv2.COLOR_RGB2YUV)
# Test RGB 2 YUV
yuv = convert_rgb2yuv(resized)
#visualize
visualize_img(yuv)
# Define image prepocess pipeline
def preprocess(img):
"""
This is a pipeline for image preprocesing
"""
img = crop_image(img)
img = resize_image(img)
img = convert_rgb2yuv(img)
return img
# Test image preprocessing pipeline
image_path_right = meta_df['right'].values[0]
img = load_image(image_path_right)
img = preprocess(img)
visualize_img(img)
```
### Image Data Augmentation
```
# Flip images
def random_flip(img, streeing_angle):
"""
"""
if np.random.rand() <0.5:
img = cv2.flip(img, 1)
streeing_angle = - streeing_angle
return img, streeing_angle
# Test for flipping
image_path_right = meta_df['right'].values[0]
img = load_image(image_path_right)
img = preprocess(img)
img , string_angle = random_flip(img, 0.1)
visualize_img(img)
# Translate images
def random_translate(img, streering_angle, range_x, range_y):
"""
"""
trans_x = range_x * (np.random.rand() - 0.5)
trans_y = range_y * (np.random.rand() - 0.5)
streering_angle += trans_x * 0.002
trans_m = np.float32([[1,0,trans_x], [0,1, trans_y]])
h,w = img.shape[:2]
img = cv2.warpAffine(img, trans_m, (w,h))
return img, streering_angle
# Test for translate
image_path_right = meta_df['right'].values[0]
img = load_image(image_path_right)
img = preprocess(img)
img , string_angle = random_translate(img, 0.1, 100, 10)
visualize_img(img)
# Add random shadow
def random_shadow(img):
"""
Generates and adds random shadow
"""
# (x1, y1) and (x2, y2) forms a line
# xm, ym gives all the locations of the image
x1, y1 = i_width * np.random.rand(), 0
x2, y2 = i_width * np.random.rand(), i_height
xm, ym = np.mgrid[0:i_height, 0:i_width]
# mathematically speaking, we want to set 1 below the line and zero otherwise
# Our coordinate is up side down. So, the above the line:
# (ym-y1)/(xm-x1) > (y2-y1)/(x2-x1)
# as x2 == x1 causes zero-division problem, we'll write it in the below form:
# (ym-y1)*(x2-x1) - (y2-y1)*(xm-x1) > 0
mask = np.zeros_like(img[:, :, 1])
mask[(ym - y1) * (x2 - x1) - (y2 - y1) * (xm - x1) > 0] = 1
# choose which side should have shadow and adjust saturation
cond = mask == np.random.randint(2)
s_ratio = np.random.uniform(low=0.2, high=0.5)
# adjust Saturation in HLS(Hue, Light, Saturation)
hls = cv2.cvtColor(img, cv2.COLOR_RGB2HLS)
hls[:, :, 1][cond] = hls[:, :, 1][cond] * s_ratio
return cv2.cvtColor(hls, cv2.COLOR_HLS2RGB)
# Test for shaow adgusments
image_path_right = meta_df['right'].values[0]
img = load_image(image_path_right)
img = preprocess(img)
img = random_shadow(img)
visualize_img(img)
# Adgust brightness randomly
def random_brightness(img):
"""
"""
# HSV (Hue, Saturation, Value) is also called HSB ('B' for Brightness).
hsv = cv2.cvtColor(img, cv2.COLOR_RGB2HSV)
ratio = 1.0 + 0.4 * (np.random.rand() - 0.5)
hsv[:,:,2] = hsv[:,:,2] * ratio
return cv2.cvtColor(hsv, cv2.COLOR_HSV2RGB)
# Test for shaow adgusments
image_path_right = meta_df['right'].values[0]
img = load_image(image_path_right)
img = preprocess(img)
img = random_brightness(img)
visualize_img(img)
# Image data augmentation
def augment(img, streering_angle, range_x =100, range_y=10):
"""
Augmenting images
"""
img, streering_angle = random_flip(img, streering_angle)
img, streering_angle = random_translate(img, streering_angle, range_x, range_y)
img = crop_image(img)
img = resize_image(img)
img = random_shadow(img)
img = random_brightness(img)
img = convert_rgb2yuv(img)
return img, streering_angle
# test image data augmentation
image_path_right = meta_df['right'].values[0]
img = load_image(image_path_right)
img, string_angle = augment(img, 0.2)
visualize_img(img)
# select random images
def select_random_image(center , left, right, streering_angle):
"""
"""
choise = np.random.randint(3)
if choise == 0:
return load_image(left), streering_angle + 0.2
elif choise == 1:
return load_image(right), streering_angle - 0.2
return load_image(center) , streering_angle
# Generate augmentation batch
def generate_batch(image_paths, streering_angles, batch_size, is_training):
"""
Generate training image give image paths and associated steering angles
"""
images = np.empty([batch_size, i_height, i_width, i_channels])
streerings = np.empty(batch_size)
while True:
i = 0
for idx in np.random.permutation(image_paths.shape[0]):
center, left, right = image_paths[idx]
steering_angle = streering_angles[idx]
if is_training and np.random.rand() < 0.6:
img, steering_angle = select_random_image(center, left, right, steering_angle)
# Augmenting
img, steering_angle = augment(img, steering_angle)
else:
img = load_image(center)
img = preprocess(img)
images[i] = img
streerings[i] = steering_angle
i += 1
if i == batch_size:
break
yield images, streerings
# test batch generates
images_paths = meta_df[['center', 'left', 'right']].values
streering_angles = meta_df['steering'].values
batch = generate_batch(images_paths,streering_angles, 10, True)
type(batch)
```
| github_jupyter |
# Quickstart
This notebook shows how to use PySTAC to read through the public Sentinel catalog and write a local version.
### Reading STAC
First, we want to hook into PySTAC to allow for reading of HTTP STAC items, as described in [the STAC_IO Concepts docs](concepts.html#using-stac-io).
__Note:__ this requires the [requests](https://requests.kennethreitz.org/en/master) library be installed.
```
from urllib.parse import urlparse
import requests
from pystac import STAC_IO
def requests_read_method(uri):
parsed = urlparse(uri)
if parsed.scheme.startswith('http'):
return requests.get(uri).text
else:
return STAC_IO.default_read_text_method(uri)
STAC_IO.read_text_method = requests_read_method
```
We can then read the STAC catalog located at the publicly available endpoint hosted by AWS:
```
from pystac import Catalog
cat = Catalog.from_file('https://sentinel-stac.s3.amazonaws.com/catalog.json')
```
There are a lot of items in this catalog; crawling through it all would take a significant amount of time. Here, we lean on the fact that [link resolution is lazy](concepts.html#lazy-resolution-of-stac-objects) and get to a catalog that contains items:
```
while len(cat.get_item_links()) == 0:
print('Crawling through {}'.format(cat))
cat = next(cat.get_children())
```
We can print some information about the catalog, including how many children it has:
```
print(cat.description)
print('Contains {} items.'.format(len(cat.get_item_links())))
```
Let's grab the first item, check out it's cloud cover, and start exploring the assets.
```
item = next(cat.get_items())
item.cloud_cover
for asset_key in item.assets:
asset = item.assets[asset_key]
print('{}: {} ({})'.format(asset_key, asset.href, asset.media_type))
```
We can use the `to_dict()` method to convert an Asset, or any PySTAC object, into a dictionary:
```
asset = item.assets['B03']
asset.to_dict()
```
Here the asset uses the band information associated with it's item:
```
asset.get_bands()[0].to_dict()
```
### Writing a STAC
Let's walk the catalog again, but this time create local clones of the STAC object, so we can end up with a copy that we can save off to the local file system.
```
import itertools
cat = Catalog.from_file('https://sentinel-stac.s3.amazonaws.com/catalog.json')
# Setup the root of our local STAC
local_root = cat.clone()
local_root.clear_children()
# Loop over catalogs and clone
curr_local_cat = local_root
while len(cat.get_item_links()) == 0:
print('Crawling through {}'.format(cat))
cat = next(cat.get_children())
local_cat = cat.clone()
local_cat.clear_children()
curr_local_cat.add_child(local_cat)
curr_local_cat = local_cat
# Clear the items from the last local catalog
curr_local_cat.clear_children()
curr_local_cat.clear_items()
# Take the first 5 items
items = itertools.islice(cat.get_items(), 5)
# Clone and add them to our local catalog
curr_local_cat.add_items([i.clone() for i in items])
```
Now that we have a smaller STAC, let's map over the items to reduce it even further by only including the thumbnail assets in our items:
```
def item_mapper(item):
thumbnail_asset = item.assets['thumbnail']
#
new_assets = { 'thumbnail': item.assets['thumbnail'] }
item.assets = new_assets
return item
local_root_2 = local_root.map_items(item_mapper)
```
We can now normalize our catalog and save it somewhere local:
```
!mkdir -p ./quickstart_stac
local_root_2.normalize_hrefs('./quickstart_stac')
from pystac import CatalogType
local_root_2.save(catalog_type=CatalogType.SELF_CONTAINED)
local_root_2.describe()
for item in local_root_2.get_all_items():
print('Item {}:')
print(' Assets: {}'.format(item.assets))
```
| github_jupyter |
# Descriptor Example: Attribute Validation
## LineItem Take #3: A Simple Descriptor
```
class Quantity:
def __init__(self, storage_name):
self.storage_name = storage_name
def __set__(self, instance, value):
if value > 0:
instance.__dict__[self.storage_name] = value
else:
raise ValueError('value must be > 0')
class LineItem:
weight = Quantity('weight')
price = Quantity('price')
def __init__(self, description, weight, price):
self.description = description
self.weight = weight
self.price = price
def subtotal(self):
return self.weight * self.price
truffle = LineItem('White truffle', 100, 0)
truffle = LineItem('White truffle', 100, 1)
truffle.__dict__
```
## LineItem Take #4: Automatic Storage Attribute Names
```
class Quantity:
__counter = 0
def __init__(self):
cls = self.__class__
prefix = cls.__name__
index = cls.__counter
self.storage_name = '_{}#{}'.format(prefix, index)
cls.__counter += 1
def __get__(self, instance, owner):
return getattr(instance, self.storage_name)
def __set__(self, instance, value):
if value > 0:
setattr(instance, self.storage_name, value)
else:
raise ValueError('value must be > 0')
class LineItem:
weight = Quantity()
price = Quantity()
def __init__(self, description, weight, price):
self.description = description
self.weight = weight
self.price = price
def subtotal(self):
return self.weight * self.price
coconuts = LineItem('Brazilian coconut', 20, 17.95)
coconuts.weight, coconuts.price
getattr(coconuts, '_Quantity#0'), getattr(coconuts, '_Quantity#1')
LineItem.weight
class Quantity:
__counter = 0
def __init__(self):
cls = self.__class__
prefix = cls.__name__
index = cls.__counter
self.storage_name = '_{}#{}'.format(prefix, index)
cls.__counter += 1
def __get__(self, instance, owner):
if instance is None:
return self
else:
return getattr(instance, self.storage_name)
def __set__(self, instance, value):
if value > 0:
setattr(instance, self.storage_name, value)
else:
raise ValueError('value must be > 0')
class LineItem:
weight = Quantity()
price = Quantity()
def __init__(self, description, weight, price):
self.description = description
self.weight = weight
self.price = price
def subtotal(self):
return self.weight * self.price
LineItem.price
br_nuts = LineItem('Brazil nuts', 10, 34.95)
br_nuts.price
```
## LineItem Take #5: A New Descriptor Type
```
import abc
class AutoStorage:
__counter = 0
def __init__(self):
cls = self.__class__
prefix = cls.__name__
index = cls.__counter
self.storage_name = '_{}#{}'.format(prefix, index)
cls.__counter += 1
def __get__(self, instance, owner):
if instance is None:
return self
else:
return getattr(instance, self.storage_name)
def __set__(self, instance, value):
setattr(instance, self.storage_name, value)
class Validated(abc.ABC, AutoStorage):
def __set__(self, instance, value):
value = self.validate(instance, value)
super().__set__(instance, value)
@abc.abstractmethod
def validate(self, instance, value):
"""return validated value or raise ValueError"""
class Quantity(Validated):
"""a number greater than zero"""
def validate(self, instance, value):
if value <= 0:
raise ValueError('value must be > 0')
return value
class NonBlank(Validated):
"""a string with at least one non-space character"""
def validate(self, instance, value):
value = value.strip()
if len(value) == 0:
raise ValueError('value cannot be empty or blank')
return value
class LineItem:
description = NonBlank()
weight = Quantity()
price = Quantity()
def __init__(self, description, weight, price):
self.description = description
self.weight = weight
self.price = price
def subtotal(self):
return self.weight * self.price
```
# Overriding Versus Nonoverriding Descriptors
```
### auxiliary functions for display only ###
def cls_name(obj_or_cls):
cls = type(obj_or_cls)
if cls is type:
cls = obj_or_cls
return cls.__name__.split('.')[-1]
def display(obj):
cls = type(obj)
if cls is type:
return '<class {}>'.format(obj.__name__)
elif cls in [type(None), int]:
return repr(obj)
else:
return '<{} object>'.format(cls_name(obj))
def print_args(name, *args):
pseudo_args = ', '.join(display(x) for x in args)
print('-> {}.__{}__({})'.format(cls_name(args[0]), name, pseudo_args))
### essential classes for this example ###
class Overriding:
"""a.k.a. data descriptor or enforded descriptor"""
def __get__(self, instance, owner):
print_args('get', self, instance, owner)
def __set__(self, instance, value):
print_args('set', self, instance, value)
class OverridingNoGet:
"""an overriding descriptor without ``__get__``"""
def __set__(self, instance, value):
print_args('set', self, instance, value)
class NonOverriding:
"""a.k.a. non-data or shadowable descriptor"""
def __get__(self, instance, owner):
print_args('get', self, instance, owner)
class Managed:
over = Overriding()
over_no_get = OverridingNoGet()
non_over = NonOverriding()
def spam(self):
print('-> Managed.spam({})'.format(display(self)))
```
## Overriding Descriptor
```
obj = Managed()
obj.over
Managed.over
obj.over = 7
obj.over
obj.__dict__['over'] = 8
vars(obj)
obj.over
```
## Overriding Descriptor Without __get__
```
obj.over_no_get
Managed.over_no_get
obj.over_no_get = 7
obj.over_no_get
obj.__dict__['over_no_get'] = 9
obj.over_no_get
obj.over_no_get = 7
obj.over_no_get
```
## Nonoverriding Descriptor
```
obj = Managed()
obj.non_over
obj.non_over = 7
obj.non_over
Managed.non_over
del obj.non_over
obj.non_over
```
## Overwriting a Descriptor in the Class
```
obj = Managed()
Managed.over = 1
Managed.over_no_get = 2
Managed.non_over = 3
obj.over, obj.over_no_get, obj.non_over
```
# Methods Are Descriptors
```
obj = Managed()
obj.spam
Managed.spam
obj.spam = 7
obj.spam
import collections
class Text(collections.UserString):
def __repr__(self):
return 'Text({!r})'.format(self.data)
def reverse(self):
return self[::-1]
word = Text('forward')
word
word.reverse()
Text.reverse(Text('backward'))
type(Text.reverse), type(word.reverse)
list(map(Text.reverse, ['repaid', (10, 20, 30), Text('stressed')]))
Text.reverse.__get__(word)
Text.reverse.__get__(None, Text)
word.reverse
word.reverse.__self__
word.reverse.__func__ is Text.reverse
```
# Descriptor docstring and Overriding Deletion
```
help(LineItem.weight)
help(LineItem)
```
| github_jupyter |
# How the PCA algorithm works
PCA makes several assumptions that are important to keep in mind. These include:
- high variance implies a high signal-to-noise ratio
- the data is standardized so that the variance is comparable across features
- linear transformations capture the relevant aspects of the data, and
- higher-order statistics beyond the first and second moment do not matter, which implies that the data has a normal distribution
The emphasis on the first and second moments align with standard risk/return metrics, but the normality assumption may conflict with the characteristics of market data.
The algorithm finds vectors to create a hyperplane of target dimensionality that minimizes the reconstruction error, measured as the sum of the squared distances of the data points to the plane. As illustrated above, this goal corresponds to finding a sequence of vectors that align with directions of maximum retained variance given the other components while ensuring all principal component are mutually orthogonal.
In practice, the algorithm solves the problem either by computing the eigenvectors of the covariance matrix or using the singular value decomposition.
## Imports & Settings
```
%matplotlib inline
import pandas as pd
import numpy as np
from numpy.linalg import inv, eig, svd
from numpy.random import uniform, randn, seed
from itertools import repeat
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
from sklearn.decomposition import PCA
from sklearn.datasets import load_digits
pd.options.display.float_format = '{:,.2f}'.format
seed(42)
def format3D(axis, labels=('x', 'y', 'z'), limits=None):
"""3D plot helper function to set labels, pane color, and axis limits"""
axis.set_xlabel('\n${}$'.format(labels[0]), linespacing=3)
axis.set_ylabel('\n${}$'.format(labels[1]), linespacing=3)
axis.set_zlabel('\n${}$'.format(labels[2]), linespacing=3)
transparent = (1.0, 1.0, 1.0, 0.0)
axis.w_xaxis.set_pane_color(transparent)
axis.w_yaxis.set_pane_color(transparent)
axis.w_zaxis.set_pane_color(transparent)
if limits:
axis.set_xlim(limits[0])
axis.set_ylim(limits[1])
axis.set_zlim(limits[2])
```
## Create a noisy 3D Ellipse
We illustrate the computation using a randomly generated three-dimensional ellipse with 100 data points.
```
n_points, noise = 100, 0.1
angles = uniform(low=-np.pi, high=np.pi, size=n_points)
x = 2 * np.cos(angles) + noise * randn(n_points)
y = np.sin(angles) + noise * randn(n_points)
theta = np.pi/4 # 45 degree rotation
rotation_matrix = np.array([[np.cos(theta), -np.sin(theta)],
[np.sin(theta), np.cos(theta)]])
rotated = np.column_stack((x, y)).dot(rotation_matrix)
x, y = rotated[:, 0], rotated[:, 1]
z = .2 * x + .2 * y + noise * randn(n_points)
data = np.vstack((x, y, z)).T
data.shape
```
### Plot the result
```
ax = plt.figure().gca(projection='3d')
ax.set_aspect('equal')
ax.scatter(x, y, z, s=25)
format3D(ax)
# plt.gcf().set_size_inches(12,12)
# plt.tight_layout();
```
## Principal Components using scikit-learn
The [sklearn.decomposition.PCA](http://scikit-learn.org/stable/modules/generated/sklearn.decomposition.PCA.html) implementation follows the standard API based on fit() and transform() methods that compute the desired number of principal components and project the data into the component space, respectively. The convenience method fit_transform() accomplishes this in a single step.
PCA offers three different algorithms that can be specified using the svd_solver parameter:
- full computes the exact SVD using the LAPACK solver provided by scipy,
- arpack runs a truncated version suitable for computing less than the full number of components.
- randomized uses a sampling-based algorithm that is more efficient when the data set has more than 500 observations and features, and the goal is to compute less than 80% of the components
- auto also randomized where most efficient, otherwise, it uses the full SVD.
Other key configuration parameters of the PCA object are:
- n_components: compute all principal components by passing None (the default), or limit the number to int. For svd_solver=full, there are two -additional options: a float in the interval [0, 1] computes the number of components required to retain the corresponding share of the variance in the data, and the option mle estimates the number of dimensions using maximum likelihood.
- whiten: if True, it standardizes the component vectors to unit variance that in some cases can be useful for use in a predictive model (default is False)
```
pca = PCA()
pca.fit(data)
C = pca.components_.T # columns = principal components
C
```
### First principal component
```
C[:, 0]
```
### Explained Variance
```
explained_variance = pca.explained_variance_
explained_variance
np.allclose(explained_variance/np.sum(explained_variance),
pca.explained_variance_ratio_)
```
### PCA to reduce dimensions
```
pca2 = PCA(n_components=2)
projected_data = pca2.fit_transform(data)
projected_data.shape
fig = plt.figure(figsize=plt.figaspect(0.5))
ax1 = fig.add_subplot(1, 2, 1, projection='3d')
ax1.set_aspect('equal')
ax1.scatter(x, y, z, s=15, c='k')
format3D(ax1)
ax2 = fig.add_subplot(1, 2, 2, projection='3d')
ax2.set_aspect('equal')
ax2.scatter(*projected_data.T, s=15, c='k')
format3D(ax1)
pca2.explained_variance_ratio_
```
## Principal Components from Covariance Matrix
We first compute the principal components using the square covariance matrix with the pairwise sample covariances for the features xi, xj, i, j = 1, ..., n as entries in row i and column j:
$$cov_{i,j}=\frac{\sum_{k=1}^{N}(x_{ik}−\bar{x_i})(x_{jk}−\bar{x_j})}{N−1}$$
Using `numpy`, we implement this as follows, where the pandas `DataFrame` data contains the 100 data points of the ellipse:
```
cov = np.cov(data.T) # each row represents a feature
cov.shape
```
### Eigendecomposition: Eigenvectors & Eigenvalues
- The Eigenvectors $w_i$ and Eigenvalues $\lambda_i$ for a square matrix $M$ are defined as follows:
$$M w_i = \lambda_i w_i$$
- This implies we can represent the matrix $M$ using Eigenvectors and Eigenvalues, where $W$ is a matrix that contains the Eigenvectors as column vectors, and $L$ is a matrix that contains the $\lambda_i$ as diagonal entries (and 0s otherwise):
$$M=WLW^{-1}$$
Next, we calculate the eigenvectors and eigenvalues of the covariance matrix. The eigenvectors contain the principal components (where the sign is arbitrary):
```
eigen_values, eigen_vectors = eig(cov)
```
### eigenvectors = principal components
```
eigen_vectors
```
We can compare the result with the result obtained from sklearn and find that they match in absolute terms:
```
np.allclose(np.abs(C), np.abs(eigen_vectors))
```
#### eigenvalues = explained variance
```
eigen_values
np.allclose(explained_variance, eigen_values)
```
#### Check that Eigendecomposition works
We can also verify the eigendecomposition, starting with the diagonal matrix L that contains the eigenvalues:
```
ev = np.zeros((3, 3))
np.fill_diagonal(ev, eigen_values)
ev # diagonal matrix
```
We find that the result does indeed hold:
```
decomposition = eigen_vectors.dot(ev).dot(inv(eigen_vectors))
np.allclose(cov, decomposition)
```
### Preferred: Singular Value Decomposition
Next, we'll look at the alternative computation using the Singular Value Decomposition (SVD). This algorithm is slower when the number of observations is greater than the number of features (the typical case) but yields better numerical stability, especially when some of the features are strongly correlated (often the reason to use PCA in the first place).
SVD generalizes the eigendecomposition that we just applied to the square and symmetric covariance matrix to the more general case of m x n rectangular matrices. It has the form shown at the center of the following figure. The diagonal values of Σ are the singular values, and the transpose of V* contains the principal components as column vectors.
#### Requires centering your data!
In this case, we need to make sure our data is centered with mean zero (the computation of the covariance above took care of this):
```
n_features = data.shape[1]
data_ = data - data.mean(axis=0)
```
We find that the decomposition does indeed reproduce the standardized data:
```
cov_manual = data_.T.dot(data_)/(len(data)-1)
np.allclose(cov_manual, cov)
U, s, Vt = svd(data_)
U.shape, s.shape, Vt.shape
# Convert s from vector to diagonal matrix
S = np.zeros_like(data_)
S[:n_features, :n_features] = np.diag(s)
S.shape
```
#### Show that the result indeed decomposes the original data
```
np.allclose(data_, U.dot(S).dot(Vt))
```
#### Confirm that $V^T$ contains the principal components
```
np.allclose(np.abs(C), np.abs(Vt.T))
```
### Visualize 2D Projection
```
pca = PCA(n_components=2)
data_2D = pca.fit_transform(data)
min_, max_ = data[:, :2].min(), data[:, :2].max()
xs, ys = np.meshgrid(np.linspace(min_,max_, n_points),
np.linspace(min_,max_, n_points))
normal_vector = np.cross(pca.components_[0], pca.components_[1])
d = -pca.mean_.dot(normal_vector)
zs = (-normal_vector[0] * xs - normal_vector[1] * ys - d) * 1 / normal_vector[2]
C = pca.components_.T.copy()
proj_matrix = C.dot(inv(C.T.dot(C))).dot(C.T)
C[:,0] *= 2
ax = plt.figure(figsize=(14,14)).gca(projection='3d')
ax.set_aspect('equal')
ax.plot_surface(xs, ys, zs, alpha=0.2, color='black',
linewidth=1, antialiased=True)
ax.scatter(x, y, z, c='k', s=25)
for i in range(n_points):
ax.plot(*zip(proj_matrix.dot(data[i]), data[i]),
color='k', lw=1)
origin = np.zeros((2, 3))
X, Y, Z, U, V, W = zip(*np.hstack((origin, C.T)))
ax.quiver(X, Y, Z, U, V, W, color='red')
format3D(ax, limits=list(repeat((min_, max_), 3)))
```
### 2D Representation
```
data_3D_inv = pca.inverse_transform(data_2D)
avg_error = np.mean(np.sum(np.square(data_3D_inv-data), axis=1))
fig = plt.figure()
ax = fig.add_subplot(111, aspect='equal',
xlabel='Principal Component 1',
ylabel='Principal Component 2',
title='Ellipse in 2D - Loss of Information: {:.2%}'.format(avg_error))
ax.scatter(data_2D[:, 0], data_2D[:, 1], color='k', s=15)
ax.arrow(0, 0, 0, .5, head_width=0.1, length_includes_head=True,
head_length=0.1, fc='k', ec='k')
ax.arrow(0, 0, 1, 0, head_width=0.1, length_includes_head=True,
head_length=0.1, fc='k', ec='k')
fig.tight_layout();
```
### How many Components to represent 64 dimensions?
```
n_classes = 10
digits = load_digits(n_class=n_classes)
X = digits.data
y = digits.target
n_samples, n_features = X.shape
n_samples, n_features
```
#### Evaluate the cumulative explained variance
```
pca = PCA(n_components=64).fit(X)
pd.Series(pca.explained_variance_ratio_).cumsum().plot()
plt.annotate('Elbow', xy=(15, .8), xycoords='data', xytext=(20, .65),
textcoords='data', horizontalalignment='center',
arrowprops=dict(color='k', lw=2, arrowstyle="->")
)
plt.axhline(.95, c='k', lw=1, ls='--');
```
### Automate generation of Components
```
pca = PCA(n_components=.95).fit(X)
pca.components_.shape
```
| github_jupyter |
# Weight Decay
:label:`sec_weight_decay`
Now that we have characterized the problem of overfitting,
we can introduce some standard techniques for regularizing models.
Recall that we can always mitigate overfitting
by going out and collecting more training data.
That can be costly, time consuming,
or entirely out of our control,
making it impossible in the short run.
For now, we can assume that we already have
as much high-quality data as our resources permit
and focus on regularization techniques.
Recall that in our
polynomial curve-fitting example
(:numref:`sec_model_selection`)
we could limit our model's capacity
simply by tweaking the degree
of the fitted polynomial.
Indeed, limiting the number of features
is a popular technique to avoid overfitting.
However, simply tossing aside features
can be too blunt an instrument for the job.
Sticking with the polynomial curve-fitting
example, consider what might happen
with high-dimensional inputs.
The natural extensions of polynomials
to multivariate data are called *monomials*,
which are simply products of powers of variables.
The degree of a monomial is the sum of the powers.
For example, $x_1^2 x_2$, and $x_3 x_5^2$
are both monomials of degree $3$.
Note that the number of terms with degree $d$
blows up rapidly as $d$ grows larger.
Given $k$ variables, the number of monomials
of degree $d$ is ${k - 1 + d} \choose {k - 1}$.
Even small changes in degree, say from $2$ to $3$,
dramatically increase the complexity of our model.
Thus we often need a more fine-grained tool
for adjusting function complexity.
## Squared Norm Regularization
*Weight decay* (commonly called *L2* regularization),
might be the most widely-used technique
for regularizing parametric machine learning models.
The technique is motivated by the basic intuition
that among all functions $f$,
the function $f = 0$
(assigning the value $0$ to all inputs)
is in some sense the *simplest*,
and that we can measure the complexity
of a function by its distance from zero.
But how precisely should we measure
the distance between a function and zero?
There is no single right answer.
In fact, entire branches of mathematics,
including parts of functional analysis
and the theory of Banach spaces,
are devoted to answering this issue.
One simple interpretation might be
to measure the complexity of a linear function
$f(\mathbf{x}) = \mathbf{w}^\top \mathbf{x}$
by some norm of its weight vector, e.g., $|| \mathbf{w} ||^2$.
The most common method for ensuring a small weight vector
is to add its norm as a penalty term
to the problem of minimizing the loss.
Thus we replace our original objective,
*minimize the prediction loss on the training labels*,
with new objective,
*minimize the sum of the prediction loss and the penalty term*.
Now, if our weight vector grows too large,
our learning algorithm might *focus*
on minimizing the weight norm $|| \mathbf{w} ||^2$
versus minimizing the training error.
That is exactly what we want.
To illustrate things in code,
let us revive our previous example
from :numref:`sec_linear_regression` for linear regression.
There, our loss was given by
$$l(\mathbf{w}, b) = \frac{1}{n}\sum_{i=1}^n \frac{1}{2}\left(\mathbf{w}^\top \mathbf{x}^{(i)} + b - y^{(i)}\right)^2.$$
Recall that $\mathbf{x}^{(i)}$ are the observations,
$y^{(i)}$ are labels, and $(\mathbf{w}, b)$
are the weight and bias parameters respectively.
To penalize the size of the weight vector,
we must somehow add $|| \mathbf{w} ||^2$ to the loss function,
but how should the model trade off the
standard loss for this new additive penalty?
In practice, we characterize this tradeoff
via the *regularization constant* $\lambda > 0$,
a non-negative hyperparameter
that we fit using validation data:
$$l(\mathbf{w}, b) + \frac{\lambda}{2} \|\mathbf{w}\|^2.$$
For $\lambda = 0$, we recover our original loss function.
For $\lambda > 0$, we restrict the size of $|| \mathbf{w} ||$.
The astute reader might wonder why we work with the squared
norm and not the standard norm (i.e., the Euclidean distance).
We do this for computational convenience.
By squaring the L2 norm, we remove the square root,
leaving the sum of squares of
each component of the weight vector.
This makes the derivative of the penalty easy to compute
(the sum of derivatives equals the derivative of the sum).
Moreover, you might ask why we work with the L2 norm
in the first place and not, say, the L1 norm.
In fact, other choices are valid and
popular throughout statistics.
While L2-regularized linear models constitute
the classic *ridge regression* algorithm,
L1-regularized linear regression
is a similarly fundamental model in statistics
(popularly known as *lasso regression*).
More generally, the $\ell_2$ is just one
among an infinite class of norms call p-norms,
many of which you might encounter in the future.
In general, for some number $p$,
the $\ell_p$ norm is defined as
$$\|\mathbf{w}\|_p^p := \sum_{i=1}^d |w_i|^p.$$
One reason to work with the L2 norm
is that it places and outsize penalty
on large components of the weight vector.
This biases our learning algorithm
towards models that distribute weight evenly
across a larger number of features.
In practice, this might make them more robust
to measurement error in a single variable.
By contrast, L1 penalties lead to models
that concentrate weight on a small set of features,
which may be desirable for other reasons.
The stochastic gradient descent updates
for L2-regularized regression follow:
$$
\begin{aligned}
\mathbf{w} & \leftarrow \left(1- \eta\lambda \right) \mathbf{w} - \frac{\eta}{|\mathcal{B}|} \sum_{i \in \mathcal{B}} \mathbf{x}^{(i)} \left(\mathbf{w}^\top \mathbf{x}^{(i)} + b - y^{(i)}\right),
\end{aligned}
$$
As before, we update $\mathbf{w}$ based on the amount
by which our estimate differs from the observation.
However, we also shrink the size of $\mathbf{w}$ towards $0$.
That is why the method is sometimes called "weight decay":
given the penalty term alone,
our optimization algorithm *decays*
the weight at each step of training.
In contrast to feature selection,
weight decay offers us a continuous mechanism
for adjusting the complexity of $f$.
Small values of $\lambda$ correspond
to unconstrained $\mathbf{w}$,
whereas large values of $\lambda$
constrain $\mathbf{w}$ considerably.
Whether we include a corresponding bias penalty $b^2$
can vary across implementations,
and may vary across layers of a neural network.
Often, we do not regularize the bias term
of a network's output layer.
## High-Dimensional Linear Regression
We can illustrate the benefits of
weight decay over feature selection
through a simple synthetic example.
First, we generate some data as before
$$y = 0.05 + \sum_{i = 1}^d 0.01 x_i + \epsilon \text{ where }
\epsilon \sim \mathcal{N}(0, 0.01).$$
choosing our label to be a linear function of our inputs,
corrupted by Gaussian noise with zero mean and variance 0.01.
To make the effects of overfitting pronounced,
we can increase the dimensionality of our problem to $d = 200$
and work with a small training set containing only 20 examples.
We will now import the relevant libraries for showing weight decay concept in action.
```
%mavenRepo snapshots https://oss.sonatype.org/content/repositories/snapshots/
%maven ai.djl:api:0.7.0-SNAPSHOT
%maven org.slf4j:slf4j-api:1.7.26
%maven org.slf4j:slf4j-simple:1.7.26
%maven ai.djl.mxnet:mxnet-engine:0.7.0-SNAPSHOT
%maven ai.djl.mxnet:mxnet-native-auto:1.7.0-b
%%loadFromPOM
<dependency>
<groupId>tech.tablesaw</groupId>
<artifactId>tablesaw-jsplot</artifactId>
<version>0.30.4</version>
</dependency>
%load ../utils/plot-utils.ipynb
%load ../utils/DataPoints.java
%load ../utils/Training.java
import ai.djl.*;
import ai.djl.engine.*;
import ai.djl.ndarray.*;
import ai.djl.ndarray.types.Shape;
import ai.djl.nn.*;
import ai.djl.nn.core.Linear;
import ai.djl.training.DefaultTrainingConfig;
import ai.djl.training.GradientCollector;
import ai.djl.training.Trainer;
import ai.djl.training.dataset.ArrayDataset;
import ai.djl.training.dataset.Batch;
import ai.djl.training.evaluator.Accuracy;
import ai.djl.training.listener.TrainingListener;
import ai.djl.training.loss.L2Loss;
import ai.djl.training.loss.Loss;
import ai.djl.training.optimizer.Optimizer;
import ai.djl.training.tracker.Tracker;
import org.apache.commons.lang3.ArrayUtils;
import tech.tablesaw.api.*;
import tech.tablesaw.plotly.api.*;
import tech.tablesaw.plotly.components.*;
import tech.tablesaw.plotly.Plot;
import tech.tablesaw.plotly.components.Figure;
int nTrain = 20;
int nTest = 100;
int numInputs = 200;
int batchSize = 5;
float trueB = 0.05f;
NDManager manager = NDManager.newBaseManager();
NDArray trueW = manager.ones(new Shape(numInputs, 1));
trueW = trueW.mul(0.01);
public ArrayDataset loadArray(NDArray features, NDArray labels, int batchSize, boolean shuffle) {
return new ArrayDataset.Builder()
.setData(features) // set the features
.optLabels(labels) // set the labels
.setSampling(batchSize, shuffle) // set the batch size and random sampling
.build();
}
DataPoints trainData = DataPoints.syntheticData(manager, trueW, trueB, nTrain);
ArrayDataset trainIter = loadArray(trainData.getX(), trainData.getY(), batchSize, true);
DataPoints testData = DataPoints.syntheticData(manager, trueW, trueB, nTest);
ArrayDataset testIter = loadArray(testData.getX(), testData.getY(), batchSize, false);
```
## Implementation from Scratch
Next, we will implement weight decay from scratch,
simply by adding the squared $\ell_2$ penalty
to the original target function.
### Initializing Model Parameters
First, we will define a function
to randomly initialize our model parameters
and run `attachGradient()` on each to allocate
memory for the gradients we will calculate.
```
public class InitParams{
private NDArray w;
private NDArray b;
private NDList l;
public NDArray getW(){
return this.w;
}
public NDArray getB(){
return this.b;
}
public InitParams(){
NDManager manager = NDManager.newBaseManager();
w = manager.randomNormal(0, 1.0f, new Shape(numInputs, 1), DataType.FLOAT32, Device.defaultDevice());
b = manager.zeros(new Shape(1));
w.attachGradient();
b.attachGradient();
}
}
```
### Defining $\ell_2$ Norm Penalty
Perhaps the most convenient way to implement this penalty
is to square all terms in place and sum them up.
We divide by $2$ by convention
(when we take the derivative of a quadratic function,
the $2$ and $1/2$ cancel out, ensuring that the expression
for the update looks nice and simple).
```
public NDArray l2Penalty(NDArray w){
return ((w.pow(2)).sum()).div(2);
}
Loss l2loss = Loss.l2Loss();
```
### Defining the Train and Test Functions
The following code fits a model on the training set
and evaluates it on the test set.
The linear network and the squared loss
have not changed since the previous chapter,
so we will just import them via `Training.linreg()` and `Training.squaredLoss()`.
The only change here is that our loss now includes the penalty term.
```
double[] trainLoss;
double[] testLoss;
double[] epochCount;
public void train(float lambd){
InitParams initParams = new InitParams();
NDList params = new NDList(initParams.getW(), initParams.getB());
int numEpochs = 100;
float lr = 0.003f;
trainLoss = new double[(numEpochs/5)];
testLoss = new double[(numEpochs/5)];
epochCount = new double[(numEpochs/5)];
for(int epoch = 1; epoch <= numEpochs; epoch++){
for(Batch batch : trainIter.getData(manager)){
NDArray X = batch.getData().head();
NDArray y = batch.getLabels().head();
NDArray w = params.get(0);
NDArray b = params.get(1);
try (GradientCollector gc = Engine.getInstance().newGradientCollector()) {
// Minibatch loss in X and y
NDArray l = Training.squaredLoss(Training.linreg(X, w, b), y).add(l2Penalty(w).mul(lambd));
gc.backward(l); // Compute gradient on l with respect to w and b
}
batch.close();
Training.sgd(params, lr, batchSize); // Update parameters using their gradient
}
if(epoch % 5 == 0){
NDArray testL = Training.squaredLoss(Training.linreg(testData.getX(), params.get(0), params.get(1)), testData.getY());
NDArray trainL = Training.squaredLoss(Training.linreg(trainData.getX(), params.get(0), params.get(1)), trainData.getY());
epochCount[epoch/5 - 1] = epoch;
trainLoss[epoch/5 -1] = trainL.mean().log10().getFloat();
testLoss[epoch/5 -1] = testL.mean().log10().getFloat();
}
}
System.out.println("l1 norm of w: " + params.get(0).abs().sum());
}
```
### Training without Regularization
We now run this code with `lambd = 0`,
disabling weight decay.
Note that we overfit badly,
decreasing the training error but not the
test error---a textook case of overfitting.
```
train(0f);
String[] lossLabel = new String[trainLoss.length + testLoss.length];
Arrays.fill(lossLabel, 0, testLoss.length, "test");
Arrays.fill(lossLabel, testLoss.length, trainLoss.length + testLoss.length, "train");
Table data = Table.create("Data").addColumns(
DoubleColumn.create("epochCount", ArrayUtils.addAll(epochCount, epochCount)),
DoubleColumn.create("loss", ArrayUtils.addAll(testLoss, trainLoss)),
StringColumn.create("lossLabel", lossLabel)
);
render(LinePlot.create("", data, "epochCount", "loss", "lossLabel"),"text/html");
```
### Using Weight Decay
Below, we run with substantial weight decay.
Note that the training error increases
but the test error decreases.
This is precisely the effect
we expect from regularization.
As an exercise, you might want to check
that the $\ell_2$ norm of the weights $\mathbf{w}$
has actually decreased.
```
// calling training with weight decay lambda = 3.0
train(3f);
String[] lossLabel = new String[trainLoss.length + testLoss.length];
Arrays.fill(lossLabel, 0, testLoss.length, "test");
Arrays.fill(lossLabel, testLoss.length, trainLoss.length + testLoss.length, "train");
Table data = Table.create("Data").addColumns(
DoubleColumn.create("epochCount", ArrayUtils.addAll(epochCount, epochCount)),
DoubleColumn.create("loss", ArrayUtils.addAll(testLoss, trainLoss)),
StringColumn.create("lossLabel", lossLabel)
);
render(LinePlot.create("", data, "epochCount", "loss", "lossLabel"),"text/html");
```
## Concise Implementation
Because weight decay is ubiquitous
in neural network optimization,
DJL makes it especially convenient,
integrating weight decay into the optimization algorithm itself
for easy use in combination with any loss function.
Moreover, this integration serves a computational benefit,
allowing implementation tricks to add weight decay to the algorithm,
without any additional computational overhead.
Since the weight decay portion of the update
depends only on the current value of each parameter,
and the optimizer must touch each parameter once anyway.
In the following code, we specify
the weight decay hyperparameter directly
through `wd` when instantiating our `Trainer`.
By default, DJL decays both
weights and biases simultaneously.
```
public void train_djl(float wd){
InitParams initParams = new InitParams();
NDList params = new NDList(initParams.getW(), initParams.getB());
int numEpochs = 100;
float lr = 0.003f;
trainLoss = new double[(numEpochs/5)];
testLoss = new double[(numEpochs/5)];
epochCount = new double[(numEpochs/5)];
Tracker lrt = Tracker.fixed(lr);
Optimizer sgd = Optimizer.sgd().setLearningRateTracker(lrt).build();
DefaultTrainingConfig config = new DefaultTrainingConfig(l2loss)
.optOptimizer(sgd) // Optimizer (loss function)
.addEvaluator(new Accuracy()) // Model Accuracy
.addEvaluator(l2loss)
.addTrainingListeners(TrainingListener.Defaults.logging()); // Logging
Model model = Model.newInstance("mlp");
SequentialBlock net = new SequentialBlock();
Linear linearBlock = Linear.builder().optBias(true).setUnits(1).build();
net.add(linearBlock);
model.setBlock(net);
Trainer trainer = model.newTrainer(config);
trainer.initialize(new Shape(batchSize, 2));
for(int epoch = 1; epoch <= numEpochs; epoch++){
for(Batch batch : trainer.iterateDataset(trainIter)){
NDArray X = batch.getData().head();
NDArray y = batch.getLabels().head();
NDArray w = params.get(0);
NDArray b = params.get(1);
try (GradientCollector gc = Engine.getInstance().newGradientCollector()) {
// Minibatch loss in X and y
NDArray l = Training.squaredLoss(Training.linreg(X, w, b), y).add(l2Penalty(w).mul(wd));
gc.backward(l); // Compute gradient on l with respect to w and b
}
batch.close();
Training.sgd(params, lr, batchSize); // Update parameters using their gradient
}
if(epoch % 5 == 0){
NDArray testL = Training.squaredLoss(Training.linreg(testData.getX(), params.get(0), params.get(1)), testData.getY());
NDArray trainL = Training.squaredLoss(Training.linreg(trainData.getX(), params.get(0), params.get(1)), trainData.getY());
epochCount[epoch/5 - 1] = epoch;
trainLoss[epoch/5 -1] = trainL.mean().log10().getFloat();
testLoss[epoch/5 -1] = testL.mean().log10().getFloat();
}
}
System.out.println("l1 norm of w: " + params.get(0).abs().sum());
}
```
The plots look identical to those when
we implemented weight decay from scratch.
However, they run appreciably faster
and are easier to implement,
a benefit that will become more
pronounced for large problems.
```
train_djl(0);
String[] lossLabel = new String[trainLoss.length + testLoss.length];
Arrays.fill(lossLabel, 0, testLoss.length, "test");
Arrays.fill(lossLabel, testLoss.length, trainLoss.length + testLoss.length, "train");
Table data = Table.create("Data").addColumns(
DoubleColumn.create("epochCount", ArrayUtils.addAll(epochCount, epochCount)),
DoubleColumn.create("loss", ArrayUtils.addAll(testLoss, trainLoss)),
StringColumn.create("lossLabel", lossLabel)
);
render(LinePlot.create("", data, "epochCount", "loss", "lossLabel"),"text/html");
```
So far, we only touched upon one notion of
what constitutes a simple *linear* function.
Moreover, what constitutes a simple *nonlinear* function
can be an even more complex question.
For instance, [Reproducing Kernel Hilbert Spaces (RKHS)](https://en.wikipedia.org/wiki/Reproducing_kernel_Hilbert_space)
allows one to apply tools introduced
for linear functions in a nonlinear context.
Unfortunately, RKHS-based algorithms
tend to scale purely to large, high-dimensional data.
In this book we will default to the simple heuristic
of applying weight decay on all layers of a deep network.
## Summary
* Regularization is a common method for dealing with overfitting. It adds a penalty term to the loss function on the training set to reduce the complexity of the learned model.
* One particular choice for keeping the model simple is weight decay using an $\ell_2$ penalty. This leads to weight decay in the update steps of the learning algorithm.
* DJL provides automatic weight decay functionality in the optimizer by setting the hyperparameter `wd`.
* You can have different optimizers within the same training loop, e.g., for different sets of parameters.
## Exercises
1. Experiment with the value of $\lambda$ in the estimation problem in this page. Plot training and test accuracy as a function of $\lambda$. What do you observe?
1. Use a validation set to find the optimal value of $\lambda$. Is it really the optimal value? Does this matter?
1. What would the update equations look like if instead of $\|\mathbf{w}\|^2$ we used $\sum_i |w_i|$ as our penalty of choice (this is called $\ell_1$ regularization).
1. We know that $\|\mathbf{w}\|^2 = \mathbf{w}^\top \mathbf{w}$. Can you find a similar equation for matrices (mathematicians call this the [Frobenius norm](https://en.wikipedia.org/wiki/Matrix_norm#Frobenius_norm))?
1. Review the relationship between training error and generalization error. In addition to weight decay, increased training, and the use of a model of suitable complexity, what other ways can you think of to deal with overfitting?
1. In Bayesian statistics we use the product of prior and likelihood to arrive at a posterior via $P(w \mid x) \propto P(x \mid w) P(w)$. How can you identify $P(w)$ with regularization?
| github_jupyter |
```
%cd ..
from hft.environment import sampler
import pandas as pd
import glob
import os
dst_dir = 'notebooks/volume-sampled-8kk'
if not os.path.exists(dst_dir):
volume = 8000000
samplerr = sampler.VolumeSampler('tests/resources/may1/orderbooks/0.csv.gz',
'tests/resources/may1/trades/0.csv.gz',
dst_dir, volume, target_symbol='XBTUSD', nrows=700000, max_workers=8)
samplerr.split_samples()
pairs = zip(glob.glob(f'{dst_dir}/orderbook_*'), glob.glob(f'{dst_dir}/trade_*'))
from hft.backtesting.strategy import CalmStrategy
from hft.backtesting.backtest import BacktestOnSample
from hft.backtesting.readers import OrderbookReader
def init_simulation(orderbook_file, trade_file):
reader = OrderbookReader(orderbook_file, trade_file, nrows=None, is_precomputed=True)
strategy = CalmStrategy(initial_balance=0.0)
backtest = BacktestOnSample(reader, strategy, delay=300)
backtest.run()
init_simulation('time-sampled/orderbook_0.csv.gz', 'time-sampled/trade_0.csv.gz')
```
## Traded volumes
```
from tqdm import tqdm
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import glob
import matplotlib.patches as mpatches
dst_dir = 'time-sampled-10min'
df = pd.read_csv(f'{dst_dir}/trade_0.csv.gz', header=None, names=['symbol', 'timestamp', 'millis', 'price', 'volume', 't', 'side'])
xbt_sell = df[(df.symbol == 'XBTUSD') & (df.side == 'Sell')]
xbt_buy = df[(df.symbol == 'XBTUSD') & (df.side == 'Buy') ]
xbt_sell.volume.sum(), len(xbt_sell)
storage = []
for trade_file in tqdm(glob.glob(f'{dst_dir}/trade_*')):
df = pd.read_csv(trade_file, header=None, names=['symbol', 'timestamp', 'millis', 'price', 'volume', 't', 'side'])
xbt = df[df.symbol == 'XBTUSD']
eth = df[df.symbol == 'ETHUSD']
xbt_sell = xbt[xbt.side == 'Sell']
xbt_buy = xbt[xbt.side == 'Buy']
eth_sell = eth[eth.side == 'Sell']
eth_buy = eth[eth.side == 'Buy']
values = tuple(map(lambda x: (x.volume.sum(), len(x)), [xbt_sell, xbt_buy, eth_sell, eth_buy]))
storage.append(values)
storage = np.array(storage).reshape((16776 // 8, 8))
df = pd.DataFrame(storage, columns=['xss', 'xst', 'xbs', 'xbt', 'ess', 'est', 'ebs', 'ebt'])
df['xs'] = df.xss + df.xbs
df['es'] = df.ess + df.ebs
df.describe()
fig=plt.figure(figsize=(16, 16))
columns = 2
rows = 2
labels = ['XBTUSD Sell volume per episode', 'XBTUSD Sell trades quantity per episode',
'XBTUSD Buy volume per episode', 'XBTUSD Buy trades quantity per episode']
for i, value, title in zip(range(1, columns*rows +1), [df.xss, df.xst, df.xbs, df.xbt], labels):
mean, std = value.mean(), value.std()
if i % 2 != 0:
mean /= 1e6
std /= 1e6
text = 'e6'
label = 'volume'
else:
text = ''
label = 'quantity'
patches = [
mpatches.Patch(color='red', label=f'mean: {mean:.2f}{text}'),
mpatches.Patch(color='blue', label=f'std: {std:.2f}{text}')
]
fig.add_subplot(rows, columns, i)
plt.scatter(range(0, len(value)), value, label=label)
plt.plot([0, len(value)], [value.mean(), value.mean()], c='red', label='mean value')
main_legend = plt.legend(loc=1)
plt.legend(handles=patches, loc=2)
plt.gca().add_artist(main_legend)
plt.title(title)
plt.show()
plt.scatter(range(0, len(storage)), df.xbs - df.xss, label='XBTUSD Sell quantity')
plt.legend()
```
| github_jupyter |
# In this notebook a Q learner with dyna will be trained and evaluated. The Q learner recommends when to buy or sell shares of one particular stock, and in which quantity (in fact it determines the desired fraction of shares in the total portfolio value).
```
# Basic imports
import os
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import datetime as dt
import scipy.optimize as spo
import sys
from time import time
from sklearn.metrics import r2_score, median_absolute_error
from multiprocessing import Pool
%matplotlib inline
%pylab inline
pylab.rcParams['figure.figsize'] = (20.0, 10.0)
%load_ext autoreload
%autoreload 2
sys.path.append('../../')
import recommender.simulator as sim
from utils.analysis import value_eval
from recommender.agent import Agent
from functools import partial
NUM_THREADS = 1
LOOKBACK = 252*2 + 28
STARTING_DAYS_AHEAD = 20
POSSIBLE_FRACTIONS = [0.0, 1.0]
DYNA = 20
# Get the data
SYMBOL = 'SPY'
total_data_train_df = pd.read_pickle('../../data/data_train_val_df.pkl').stack(level='feature')
data_train_df = total_data_train_df[SYMBOL].unstack()
total_data_test_df = pd.read_pickle('../../data/data_test_df.pkl').stack(level='feature')
data_test_df = total_data_test_df[SYMBOL].unstack()
if LOOKBACK == -1:
total_data_in_df = total_data_train_df
data_in_df = data_train_df
else:
data_in_df = data_train_df.iloc[-LOOKBACK:]
total_data_in_df = total_data_train_df.loc[data_in_df.index[0]:]
# Create many agents
index = np.arange(NUM_THREADS).tolist()
env, num_states, num_actions = sim.initialize_env(total_data_in_df,
SYMBOL,
starting_days_ahead=STARTING_DAYS_AHEAD,
possible_fractions=POSSIBLE_FRACTIONS)
agents = [Agent(num_states=num_states,
num_actions=num_actions,
random_actions_rate=0.98,
random_actions_decrease=0.999,
dyna_iterations=DYNA,
name='Agent_{}'.format(i)) for i in index]
def show_results(results_list, data_in_df, graph=False):
for values in results_list:
total_value = values.sum(axis=1)
print('Sharpe ratio: {}\nCum. Ret.: {}\nAVG_DRET: {}\nSTD_DRET: {}\nFinal value: {}'.format(*value_eval(pd.DataFrame(total_value))))
print('-'*100)
initial_date = total_value.index[0]
compare_results = data_in_df.loc[initial_date:, 'Close'].copy()
compare_results.name = SYMBOL
compare_results_df = pd.DataFrame(compare_results)
compare_results_df['portfolio'] = total_value
std_comp_df = compare_results_df / compare_results_df.iloc[0]
if graph:
plt.figure()
std_comp_df.plot()
```
## Let's show the symbols data, to see how good the recommender has to be.
```
print('Sharpe ratio: {}\nCum. Ret.: {}\nAVG_DRET: {}\nSTD_DRET: {}\nFinal value: {}'.format(*value_eval(pd.DataFrame(data_in_df['Close'].iloc[STARTING_DAYS_AHEAD:]))))
# Simulate (with new envs, each time)
n_epochs = 4
for i in range(n_epochs):
tic = time()
env.reset(STARTING_DAYS_AHEAD)
results_list = sim.simulate_period(total_data_in_df,
SYMBOL,
agents[0],
starting_days_ahead=STARTING_DAYS_AHEAD,
possible_fractions=POSSIBLE_FRACTIONS,
verbose=False,
other_env=env)
toc = time()
print('Epoch: {}'.format(i))
print('Elapsed time: {} seconds.'.format((toc-tic)))
print('Random Actions Rate: {}'.format(agents[0].random_actions_rate))
show_results([results_list], data_in_df)
env.reset(STARTING_DAYS_AHEAD)
results_list = sim.simulate_period(total_data_in_df,
SYMBOL, agents[0],
learn=False,
starting_days_ahead=STARTING_DAYS_AHEAD,
possible_fractions=POSSIBLE_FRACTIONS,
other_env=env)
show_results([results_list], data_in_df, graph=True)
```
## Let's run the trained agent, with the test set
### First a non-learning test: this scenario would be worse than what is possible (in fact, the q-learner can learn from past samples in the test set without compromising the causality).
```
TEST_DAYS_AHEAD = 20
env.set_test_data(total_data_test_df, TEST_DAYS_AHEAD)
tic = time()
results_list = sim.simulate_period(total_data_test_df,
SYMBOL,
agents[0],
learn=False,
starting_days_ahead=TEST_DAYS_AHEAD,
possible_fractions=POSSIBLE_FRACTIONS,
verbose=False,
other_env=env)
toc = time()
print('Epoch: {}'.format(i))
print('Elapsed time: {} seconds.'.format((toc-tic)))
print('Random Actions Rate: {}'.format(agents[0].random_actions_rate))
show_results([results_list], data_test_df, graph=True)
```
### And now a "realistic" test, in which the learner continues to learn from past samples in the test set (it even makes some random moves, though very few).
```
env.set_test_data(total_data_test_df, TEST_DAYS_AHEAD)
tic = time()
results_list = sim.simulate_period(total_data_test_df,
SYMBOL,
agents[0],
learn=True,
starting_days_ahead=TEST_DAYS_AHEAD,
possible_fractions=POSSIBLE_FRACTIONS,
verbose=False,
other_env=env)
toc = time()
print('Epoch: {}'.format(i))
print('Elapsed time: {} seconds.'.format((toc-tic)))
print('Random Actions Rate: {}'.format(agents[0].random_actions_rate))
show_results([results_list], data_test_df, graph=True)
```
## What are the metrics for "holding the position"?
```
print('Sharpe ratio: {}\nCum. Ret.: {}\nAVG_DRET: {}\nSTD_DRET: {}\nFinal value: {}'.format(*value_eval(pd.DataFrame(data_test_df['Close'].iloc[STARTING_DAYS_AHEAD:]))))
```
## Conclusion:
```
import pickle
with open('../../data/simple_q_learner_fast_learner_full_training.pkl', 'wb') as best_agent:
pickle.dump(agents[0], best_agent)
```
| github_jupyter |
# $\star\star$ Currently in development. This example doesn't work $\star\star$
Some parts of `pygsti` are works-in-progress. Here, we investigate how to do the task of "model selection" within GST, essentially answering the question "Can we do a better job of modeling the experiment by changing the assumptions within GST?".
## Testing variable-gateset-dimension GST with model selection
### Setup
```
from __future__ import print_function
import pygsti
from pygsti.construction import std1Q_XYI
#Load gateset and some string lists
gs_target = std1Q_XYI.gs_target
fiducialList = std1Q_XYI.fiducials
germList = std1Q_XYI.germs
specs = pygsti.construction.build_spam_specs(fiducialList)
expList = [1,2,4]
#Create some testing gate string lists
lgstList = pygsti.construction.list_lgst_gatestrings(specs, gs_target.gates.keys())
lsgstLists = [ lgstList[:] ]
for exp in expList:
gsList = pygsti.construction.create_gatestring_list(
"f0+germ*exp+f1", f0=fiducialList, f1=fiducialList,
germ=germList, exp=exp, order=['germ','f0','f1'])
lsgstLists.append( lsgstLists[-1] + gsList )
dsList = pygsti.remove_duplicates( lsgstLists[-1] )
#Test on fake data by depolarizing target set, increasing its dimension,
# and adding leakage to the gates into the new dimension.
gs_dataGen4 = gs_target.depolarize(gate_noise=0.1)
gs_dataGen5 = gs_dataGen4.increase_dimension(5)
leakGate = pygsti.construction.build_gate( [2,1],[('Q0',),('L0',)] , "LX(pi/4.0,0,2)","gm") # X(pi,Q0)*LX(pi,0,2)
gs_dataGen5['Gx'] = pygsti.objects.compose( gs_dataGen5['Gx'], leakGate, 'gm')
gs_dataGen5['Gy'] = pygsti.objects.compose( gs_dataGen5['Gy'], leakGate, 'gm')
print(gs_dataGen5.gates.keys())
#Some debugging...
#NOTE: with LX(pi,0,2) above, dim 5 test will choose a dimension 3 gateset, which may be sensible
# looking at the gate matrices in this case... but maybe LX(pi,...) is faulty?
#print(gs_dataGen4)
#print(gs_dataGen5)
#Jmx = GST.JOps.jamiolkowski_iso(gs_dataGen4['Gx'])
#Jmx = GST.JOps.jamiolkowski_iso(gs_dataGen5['Gx'],dimOrStateSpaceDims=[2,1])
#print("J = \n",Jmx)
#print("evals = ",eigvals(Jmx))
dsFake4 = pygsti.construction.generate_fake_data(gs_dataGen4, dsList, nSamples=1000000, sampleError="binomial", seed=1234)
dsFake5 = pygsti.construction.generate_fake_data(gs_dataGen5, dsList, nSamples=1000000, sampleError="binomial", seed=1234)
print("Number of gates = ",len(gs_target.gates.keys()))
print("Number of fiducials =",len(fiducialList))
print("Maximum length for a gate string in ds =",max(map(len,dsList)))
print("Number of LGST strings = ",len(lgstList))
print("Number of LSGST strings = ",map(len,lsgstLists))
```
### Test using dimension-4 fake data
```
#Run LGST to get an initial estimate for the gates in gs_target based on the data in ds
# NOTE: with nSamples less than 1M (100K, 10K, 1K) this routine will choose a higher-than-4 dimensional gateset
ds = dsFake4
gs_lgst4 = pygsti.do_lgst(ds, specs, targetGateset=gs_target, svdTruncateTo=4, verbosity=3)
gs_lgst6 = pygsti.do_lgst(ds, specs, targetGateset=gs_target, svdTruncateTo=6, verbosity=3)
#Print chi^2 of 4-dim and 6-dim estimates
chiSq4 = pygsti.chi2(ds, gs_lgst4, lgstList, minProbClipForWeighting=1e-4)
chiSq6 = pygsti.chi2(ds, gs_lgst6, lgstList, minProbClipForWeighting=1e-4)
print("LGST dim=4 chiSq = ", chiSq4)
print("LGST dim=6 chiSq = ", chiSq6)
# Least squares GST with model selection
gs_lsgst = pygsti.do_iterative_mc2gst_with_model_selection(ds, gs_lgst4, 1, lsgstLists, verbosity=2,
minProbClipForWeighting=1e-3, probClipInterval=(-1e5,1e5))
print(gs_lsgst)
```
### Test using dimension-5 fake data
```
#Run LGST to get an initial estimate for the gates in gs_target based on the data in ds
ds = dsFake5
gs_lgst4 = pygsti.do_lgst(ds, specs, targetGateset=gs_target, svdTruncateTo=4, verbosity=3)
gs_lgst6 = pygsti.do_lgst(ds, specs, targetGateset=gs_target, svdTruncateTo=6, verbosity=3)
#Print chi^2 of 4-dim and 6-dim estimates
chiSq4 = pygsti.chi2(ds, gs_lgst4, lgstList, minProbClipForWeighting=1e-2)
chiSq6 = pygsti.chi2(ds, gs_lgst6, lgstList, minProbClipForWeighting=1e-2)
print("LGST dim=4 chiSq = ", chiSq4)
print("LGST dim=6 chiSq = ", chiSq6)
# Least squares GST with model selection
gs_lsgst = pygsti.do_iterative_mc2gst_with_model_selection(ds, gs_lgst4, 1, lsgstLists, verbosity=2, minProbClipForWeighting=1e-3, probClipInterval=(-1e5,1e5), useFreqWeightedChiSq=False, regularizeFactor=1.0, check=False, check_jacobian=False)
print(gs_lsgst)
```
| github_jupyter |
# Fourier tranform time series
Fourier analysis is the study of the way general functions may be represented or approximated by sums of simpler trigonometric functions. By decomposing a function in this sum we can get informations about the dominant frequencies ( or the periods ) of these functions.
When we do a Fourier transform of a function of time, we decompose it into the frequencies that make it up.
$$ \hat{f}(\omega) = \int_{-\infty}^{\infty} f(t)\ e^{- 2\pi \omega i x}\,dt $$
### 1) Discrete Fourier Transform (DFT)
Discrete-time Fourier transform (DTFT) is a form of Fourier analysis that is applicable to uniformly-spaced samples of a continuous function. The term discrete-time refers to the fact that the transform operates on discrete data (samples) whose interval often has units of time.
$$ F(\omega) = \sum_{n=0}^{N-1} x[n] \,e^{-i \omega n} $$
### 1) Example:
```
# Importing Libraries
import matplotlib.pyplot as plt
import numpy as np
%matplotlib inline
# We create and plot some time-series
N = 400
fs = 400
t = np.linspace(-.5, .5, N)
f0 = 10
phi = np.pi/2
A = 1
def x(A,f0,t,phi):
return A * np.sin(2 * np.pi * f0 * t + phi)+np.cos((np.pi*f0+phi)/2)
x = x(A,f0,t,phi)
plt.plot(t, x)
plt.axis([-.5, .5, -1.8, 1.8])
plt.xlabel('time')
plt.ylabel('amplitude')
plt.show()
# We yse fft DFT function in numpy to decompose FT
DFT = np.fft.fft(x)
freq = np.fft.fftfreq(x.size)
plt.plot(freq, np.abs(DFT))
plt.xlabel('frequency')
plt.xlim(-0.13,0.13)
plt.show()
# Display only positive part
half = len(x) // 2
plt.plot(freq[:half], np.abs(DFT[:half]))
plt.xlabel('frequency')
plt.xlim(0,0.13)
plt.show()
```
### 2) Example
We are going to create some random distribution and estimate the frequencies
```
# 0 is the mean of the normal distribution you are choosing from
# .4 is the standard deviation of the normal distribution
# N is the number of elements you get in array noise
noise = np.random.normal(0,0.4,N)
plt.plot(t, noise)
plt.xlabel('time')
plt.ylabel('amplitude')
plt.show()
# Analyze the DFT (only positive half)
DFT = np.fft.fft(noise)
freq = np.fft.fftfreq(noise.size)
plt.plot(freq[:half], np.abs(DFT[:half]))
plt.show()
```
### 3) Example
```
f1 = 10
f2 = 2
phi_1 = np.pi/3
phi_2 = np.pi/4
B = 2
C = 3
D = 4
noise = D*np.random.normal(0,.4,N)
S = A*np.sin(2*np.pi*f0*t + phi) + B*np.sin(2*np.pi*f1*t + phi_1) + C*np.sin(2*np.pi*f2*t + phi_2) + noise
plt.plot(t, S)
plt.axis([-.5, .5, -10, 10])
plt.xlabel('time')
plt.ylabel('amplitude')
plt.show()
DFT = np.fft.fft(S)
freq = np.fft.fftfreq(S.size)
plt.plot(freq[:half], np.abs(DFT[:half]) ** 2)
plt.yscale("log")
plt.show()
```
### 4) Example: Sunspots
We are going to use the data of sunspots/year (after 1700AD) to estimate the frequency they occur.
```
# Load data
wolf = np.loadtxt("data/sunspots.txt")
year = 1700 + np.arange(len(wolf)) # years starting from 1700
plt.plot(year, wolf)
plt.xlabel("Year")
plt.ylabel("Wolf number")
plt.show()
# Function gets signal and gives freqs, power-spectrum and phase
def get_spectra(signal):
transform = np.fft.rfft(signal)
frequencies = np.fft.rfftfreq(signal.size)
power_spectrum = np.abs(transform) ** 2
phase_spectrum = np.angle(transform)
return frequencies, power_spectrum, phase_spectrum
freqs, power, phase = get_spectra(wolf)
# Make multiple plot
plt.figure(1,figsize=(13,10))
plt.subplot(221)
plt.plot(freqs, power, ".-")
plt.xlabel("Frequency ($yr^{-1}$)"); plt.ylabel("Power")
plt.yscale("log")
# plot power vs period for period < 25yr - ignore the constant term to avoid infinity
plt.subplot(222)
plt.plot(1.0 / freqs[1:], power[1:], ".-")
plt.axvline(11, color="r")
plt.xlim([0.0, 25.0])
plt.xlabel("Period (yr)"); plt.ylabel("Power")
plt.show()
```
| github_jupyter |
```
%matplotlib inline
%reload_ext autoreload
%autoreload 2
%config InlineBackend.figure_format = 'retina'
from rec_to_binaries.read_binaries import readTrodesExtractedDataFile
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
```
### .time directory
+ `.continuoustime.dat` lists the timestamp of every entry in the rec file, and include systime when that is enabled.
+ `.time.dat` lists the start and end time of the epoch
+ `.exporttime.log` is the log from the C function used to create `continuoustime.dat` and `time.dat` files. It also includes the arguments used to create the function.
```
readTrodesExtractedDataFile(
'../test_data/lotus/preprocessing/20190902/20190902_lotus_06_r3.time/20190902_lotus_06_r3.continuoustime.dat')
continuous_time = readTrodesExtractedDataFile(
'../test_data/lotus/preprocessing/20190902/20190902_lotus_06_r3.time/20190902_lotus_06_r3.continuoustime.dat')
sampling_rate = float(continuous_time['Clockrate'])
continuous_time = pd.DataFrame(continuous_time['data'])
seconds = continuous_time.systime / 1E9
ct = pd.to_datetime(continuous_time.systime.values)
ct
np.diff(ct) / np.timedelta64(1, 's')
(ct - pd.Timestamp("1970-01-01")) // pd.Timedelta('1ns'), continuous_time.systime.values
pd.Timestamp(continuous_time.systime.values[0], tz='US/Pacific')
plt.figure(figsize=(10, 8))
n_samples = 100
plt.plot(continuous_time.trodestime.values[:n_samples], seconds[:n_samples])
plt.plot(continuous_time.trodestime.values[:n_samples],
seconds[0] + np.arange(n_samples) / sampling_rate)
ideal = (seconds[0] + np.arange(continuous_time.systime.values.size) / sampling_rate)
plt.hist(np.abs(seconds - ideal) * 1000, bins=1000);
np.mean(np.diff(seconds) >= 2 / sampling_rate)
np.max(np.diff(seconds)) * 1000
np.diff(seconds) * 1000
plt.figure(figsize=(20, 4))
plt.boxplot(np.diff(seconds) * 1000, vert=False);
plt.axvline(1000 * 2 / sampling_rate, color='red', linestyle='--')
plt.xlabel('Difference between time samples (ms)')
np.mean(np.diff(continuous_time.trodestime) > 1)
np.max(np.diff(continuous_time.trodestime))
is_adjacent_sample = (np.diff(continuous_time.trodestime) == 1)
adjusted_sampling_rate = np.mean(np.diff(seconds))
pd.Timestamp(continuous_time.systime[0])
pd.Timedelta(1/30000, unit='s')
pd.Timestamp(continuous_time.systime[0]) + pd.Timedelta(1/30000, unit='s')
pd.period_range?
filename = '../test_data/lotus/preprocessing/20190902/20190902_lotus_06_r3.time/20190902_lotus_06_r3.continuoustime.dat'
with open(filename, 'rb') as f:
for line in f:
print(line)
print(line.decode('ascii').strip())
from rec_to_binaries.read_binaries import parse_dtype
filename = '../test_data/lotus/preprocessing/20190902/20190902_lotus_06_r3.time/20190902_lotus_06_r3.continuoustime.dat'
with open(filename, 'rb') as file:
# Check if first line is start of settings block
if file.readline().decode('ascii').strip() != '<Start settings>':
raise Exception("Settings format not supported")
fieldsText = {}
for line in file:
# Read through block of settings
line = line.decode('ascii').strip()
# filling in fields dict
if line != '<End settings>':
settings_name, setting = line.split(': ')
fieldsText[settings_name.lower()] = setting
# End of settings block, signal end of fields
else:
break
# Reads rest of file at once, using dtype format generated by parse_dtype()
dtype = parse_dtype(fieldsText['fields'])
fieldsText['data'] = np.fromfile(
file, dtype=dtype)
dtype
data_file = readTrodesExtractedDataFile(
'../test_data/lotus/preprocessing/20190902/20190902_lotus_06_r3.time/20190902_lotus_06_r3.continuoustime.dat')
data_file
def write_trodes_extracted_datafile(filename, data_file):
"""
Parameters
----------
data_file : dict
filename : str
"""
with open(filename, 'wb') as file:
file.write('<Start settings>\n'.encode())
for key, value in data_file.items():
if key != 'data':
line = f'{key}: {value}\n'.encode()
file.write(line)
file.write('<End settings>\n'.encode())
file.write(data_file['data'].tobytes())
write_trodes_extracted_datafile('test.dat', data_file)
readTrodesExtractedDataFile('test.dat')
from scipy.ndimage import label
continuous_time = readTrodesExtractedDataFile(
'../test_data/lotus/preprocessing/20190902/20190902_lotus_06_r3.time/20190902_lotus_06_r3.continuoustime.dat')
sampling_rate = float(continuous_time['Clockrate'])
continuous_time = pd.DataFrame(continuous_time['data'])
seconds = continuous_time.systime / 1E9
label(np.diff(continuous_time.trodestime.values))
fake_trodestime = continuous_time.trodestime.values
fake_trodestime[1:] = fake_trodestime[1:] + 1
np.allclose(fake_trodestime, continuous_time.trodestime.values)
fake_trodestime
label(fake_trodestime)
import os
import rec_to_binaries.trodes_data as td
data_dir = os.path.join(os.pardir, 'test_data')
animal = 'lotus'
animal_info = td.TrodesAnimalInfo(data_dir, animal)
extractor = td.ExtractRawTrodesData(animal_info)
raw_epochs_unionset = animal_info.get_raw_epochs_unionset()
if len(raw_epochs_unionset) == 0:
logger.warning('No epochs found!')
raw_dates = animal_info.get_raw_dates()
raw_dates, raw_epochs_unionset
from itertools import product
preprocessing_dir = animal_info.get_preprocessing_dir()
for date, epoch in product(raw_dates, raw_epochs_unionset):
preprocessing_dir = animal_info.get_preprocessing_dir()
task = os.path.basename(animal_info.get_raw_rec_path(date, epoch)[1]).split('.')[0].split('_')[-1]
time_folder = os.path.join(preprocessing_dir, date, f"{date}_{animal}_{epoch:02d}_{task}.time")
print(time_folder)
import glob
glob.glob(os.path.join(preprocessing_dir, '**', '*.continuoustime.dat'), recursive=True)
glob.glob?
df = animal_info.preproc_datatype_dirs
df[df.datatype == "time"].directory.values
test = np.concatenate((np.arange(10), np.arange(15, 20), np.arange(25, 30), np.arange(32, 40, 2)))
test
def label_time_chunks(time_index):
time_index = np.asarray(time_index)
is_gap = np.diff(time_index) > 1
is_gap = np.insert(is_gap, 0, False)
return np.cumsum(is_gap)
label_time_chunks(test)
continuous_time = continuous_time.assign(time_chunk_label=lambda df: label_time_chunks(df.trodestime))
continuous_time
systime = pd.to_datetime(continuous_time.systime)
systime
plt.plot(continuous_time.trodestime, systime.values)
plt.axhline(systime.mean(), color='black', linestyle='--')
midpoint = systime.mean()
estimated_sampling_rate = np.mean(np.diff(systime))
n_samples = len(systime)
n_samples // 2
midpoint + (np.arange(n_samples) * estimated_sampling_rate)
end_time = midpoint + estimated_sampling_rate * (n_samples - (n_samples // 2))
start_time = midpoint - estimated_sampling_rate * (n_samples // 2)
adjusted_timestamps = pd.date_range(start=start_time, end=end_time, periods=n_samples)
plt.plot(continuous_time.trodestime, systime.values)
plt.axhline(systime.mean(), color='black', linestyle='--')
plt.plot(continuous_time.trodestime, adjusted_timestamps)
time_slice = slice(10000, 10000 + 13)
plt.figure(figsize=(10, 10))
plt.plot(continuous_time.trodestime.values[time_slice], systime.values[time_slice])
plt.plot(continuous_time.trodestime.values[time_slice], adjusted_timestamps[time_slice])
adjusted_timestamps.astype(int).values
def adjust_timestamps(systime):
systime = pd.to_datetime(systime)
midpoint = systime.mean()
estimated_sampling_rate = np.mean(np.diff(systime))
n_samples = len(systime)
end_time = midpoint + estimated_sampling_rate * (n_samples - (n_samples // 2))
start_time = midpoint - estimated_sampling_rate * (n_samples // 2)
return pd.date_range(start=start_time, end=end_time, periods=n_samples).astype(int)
ct = continuous_time.assign(adjusted_systime=lambda df: adjust_timestamps(df.systime))
ct
def insert_new_data(data_file, df):
new_data_file = data_file.copy()
new_data_file['data'] = np.asarray(df.to_records(index=False))
new_data_file['Fields'] = ''.join(
[f'<{name} {dtype}>'
for name, (dtype, _) in new_data_file['data'].dtype.fields.items()])
return new_data_file
filename = '../test_data/lotus/preprocessing/20190902/20190902_lotus_06_r3.time/20190902_lotus_06_r3.continuoustime.dat'
data_file = readTrodesExtractedDataFile(filename)
new_data = (pd.DataFrame(data_file['data'])
.assign(time_chunk_label=lambda df: label_time_chunks(df.trodestime))
.assign(adjusted_systime=lambda df: adjust_timestamps(df.systime)))
new_data_file = insert_new_data(data_file, new_data)
# write_trodes_extracted_datafile(filename, new_data_file)
new_data_file
dtypes = np.asarray(ct.to_records(index=False)).dtype
dtypes
[f'<{name} {dtype}>' for name, dtype in dtypes.descr]
dtypes.descr[0][1]
data_file['data']
from scipy.stats import poisson, expon
plt.hist(expon.rvs(size=1000))
continuous_time
from sklearn.linear_model import LinearRegression
l = LinearRegression()
X = continuous_time.trodestime.values[:, np.newaxis]
y = continuous_time.systime / 1E9
l.fit(X, y)
predicted_time = l.predict(X)
plt.plot(continuous_time.trodestime.values, predicted_time)
plt.plot(continuous_time.trodestime.values, y)
time_slice = slice(-200, -1)
plt.plot(continuous_time.trodestime.values[time_slice ], predicted_time[time_slice ])
plt.plot(continuous_time.trodestime.values[time_slice ], y[time_slice ])
l.coef_
plt.plot(continuous_time.trodestime.values, l.predict(X))
from scipy.stats import linregress
x = continuous_time.trodestime.values.astype(float)
y = continuous_time.systime.values / 1E9
slope, intercept, r_value, p_value, std_err = linregress(x, y)
predicted_time = intercept + slope * x
plt.plot(x, y, label='original data')
plt.plot(x, predicted_time, label='fitted line')
time_slice = slice(3 * 52159456 // 4, (3 * 52159456 // 4) + 300)
plt.figure(figsize=(10, 10))
plt.plot(x[time_slice], y[time_slice], label='original data')
plt.plot(x[time_slice], predicted_time[time_slice], label='fitted line')
plt.figure(figsize=(10, 5))
plt.hist(y - predicted_time, bins=200);
plt.xlim((-0.001, 0.001))
plt.title('Residuals')
np.max(np.abs(y - predicted_time)) * 1000
np.diff(predicted_time) * 1E9
1 / 30000 * 1E9
x.shape
blah = (ct - pd.Timestamp("1970-01-01")) // pd.Timedelta('1s')
pd.Series(ct) // pd.Timedelta('1s')
from scipy.stats import linregress
def label_time_chunks(time_index):
time_index = np.asarray(time_index)
is_gap = np.diff(time_index) > 1
is_gap = np.insert(is_gap, 0, False)
return np.cumsum(is_gap)
def _regress_timestamps(trodestime, systime):
NANOSECONDS_TO_SECONDS = 1E9
# Convert
systime_seconds = np.asarray(systime).astype(
np.float64) / NANOSECONDS_TO_SECONDS
trodestime_index = np.asarray(trodestime).astype(np.float64)
slope, intercept, r_value, p_value, std_err = linregress(
trodestime_index, systime_seconds)
adjusted_timestamps = intercept + slope * trodestime_index
return (adjusted_timestamps * NANOSECONDS_TO_SECONDS).astype(np.int64)
def insert_new_data(data_file, df):
new_data_file = data_file.copy()
new_data_file['data'] = np.asarray(df.to_records(index=False))
new_data_file['Fields'] = ''.join(
[f'<{name} {dtype}>'
for name, (dtype, _) in new_data_file['data'].dtype.fields.items()])
return new_data_file
filename = '../test_data/lotus/preprocessing/20190902/20190902_lotus_06_r3.time/20190902_lotus_06_r3.continuoustime.dat'
data_file = readTrodesExtractedDataFile(filename)
new_data = (pd.DataFrame(data_file['data'])
.assign(time_chunk_label=lambda df: label_time_chunks(df.trodestime))
.assign(adjusted_systime=lambda df: _regress_timestamps(df.trodestime, df.systime)))
new_data_file = insert_new_data(data_file, new_data)
# write_trodes_extracted_datafile(filename, new_data_file)
new_data_file
continuous_time
```
| github_jupyter |
# Exemplo 11: Aprendizagem de Máquina Serial
## Identificação espécie de flores com Iris Dataset
---
Machine learning is the method of extracting knowledge from data. It is a field at the intersection of statistics, artificial intelligence, and computer science and is also used to predict analytics or statistic learning. They are called supervised learning algorithms because it is necessary to provide supervision to the algorithms in the form of the desired outputs for each example that they learn from. So, each line of dataset has the correct answer for the problem, in this casa, a correct classification. The Iris specie identification is a basic example of machine learning pratice. It is the "Hello world" in machine learning.
The Iris problem is so well understood and has intersting characteristics: (1) Attributes are numeric so you have to figure out how to load and handle data; (2) It is a classification problem, allowing you to practice with perhaps an easier type of supervised learning algorithm; (3) It is a multi-class classification problem (multi-nominal) that may require some specialized handling; (4) It only has 4 attributes and 150 rows, meaning it is small and easily fits into memory (and a screen or A4 page); and, (5) All of the numeric attributes are in the same units and the same scale, not requiring any special scaling or transforms to get started.
## Iris Plants Database
----
This is perhaps the best known database to be found in the pattern recognition literature. Fisher's paper is a classic in the field and is referenced frequently to this day. The data set contains 3 classes of 50 instances each, where each class refers to a type of iris plant. One class is linearly separable from the other 2; the latter are NOT linearly separable from each other.
### Predicted attribute:
Class of iris plant.
### Number of Instances:
150 (50 in each of three classes)
### Number of Attributes:
4 numeric, predictive attributes and the class
### Attribute Information:
1. sepal length in cm
2. sepal width in cm
3. petal length in cm
4. petal width in cm
5. class:
+ Iris Setosa
+ Iris Versicolour
+ Iris Virginica
### Missing Attribute Values:
None
### Class Distribution:
33.3% for each of 3 classes.
## Summary Statistics
-----
Attributes | Min | Max | Mean | SD | Class Correlation
-----------|-----|-----|------|----|------------
sepal length | 4.3 | 7.9 | 5.84 | 0.83 | 0.7826
sepal width | 2.0 | 4.4 | 3.05 | 0.43 | -0.4194
petal length | 1.0 | 6.9 | 3.76 | 1.76 | 0.9490
petal width | 0.1 | 2.5 | 1.20 | 0.76 | 0.9565
## References:
1. Fisher,R.A. "The use of multiple measurements in taxonomic problems" Annual Eugenics, 7, Part II, 179-188 (1936); also in "Contributions to Mathematical Statistics" (John Wiley, NY, 1950).
2. Duda,R.O., & Hart,P.E. (1973) Pattern Classification and Scene Analysis. John Wiley & Sons. ISBN 0-471-22361-1.
```
# Spark Lib
import findspark
findspark.init()
# Load libraries
import pyspark
from pyspark.sql import SparkSession
from pyspark.ml import Pipeline
from pyspark.ml.classification import DecisionTreeClassifier
from pyspark.ml.feature import StringIndexer, VectorIndexer
from pyspark.ml.evaluation import MulticlassClassificationEvaluator
from pyspark.mllib.util import MLUtils
from pyspark.ml.feature import StringIndexer, IndexToString
from pyspark.ml.feature import VectorAssembler, VectorIndexer
from pyspark.ml.classification import MultilayerPerceptronClassifier
from pyspark.ml.classification import NaiveBayes
from pyspark.ml.classification import LogisticRegression
from pyspark.ml.classification import LinearSVC, OneVsRest
from pyspark.ml.classification import RandomForestClassifier
from pyspark.ml.classification import DecisionTreeClassifier
from pyspark.ml.evaluation import MulticlassClassificationEvaluator
from pyspark.ml.linalg import Vectors
from pyspark.mllib.util import MLUtils
## SKLearn Lib
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.preprocessing import LabelEncoder
from sklearn.model_selection import train_test_split
from sklearn.neighbors import KNeighborsClassifier
from sklearn.metrics import confusion_matrix, accuracy_score
from sklearn.model_selection import cross_val_score
import time
start_time = time.time()
%matplotlib inline
```
## Configure parameters
```
# Path to dataset file
data_path='./data/'
# Sample of train and test dataset
train_sample = 0.7
test_sample = 0.3
```
## Read and show dataset
```
# Importing the dataset
dataset = pd.read_csv(data_path+"iris.data")
# Print dataset
dataset.head(5)
print("Number of itens per class")
dataset.groupby('class').size()
# Read features and calass
feature_columns = ['sepal length', 'sepal width', 'petal length','petal width']
X = dataset[feature_columns].values
y = dataset['class'].values
# SKLearn need all column as numbers. Tranform "class" column in number
le = LabelEncoder()
y = le.fit_transform(y)
# Split ramdomly the dataset into train and test group
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = test_sample, random_state = 0)
plt.figure()
sns.pairplot(dataset, hue = "class", height=3, markers=["o", "s", "D"])
plt.show()
```
## KNN using Scikit-learn
```
start_time_knn = time.time()
# Instantiate learning model (k = 3)
classifier = KNeighborsClassifier(n_neighbors=3)
# Fitting the model
classifier.fit(X_train, y_train)
# Predicting the Test set results
y_pred = classifier.predict(X_test)
accuracy_knn = accuracy_score(y_test, y_pred)*100
time_knn = time.time() - start_time_knn
print('KNN accuracy = ' + str(round(accuracy_knn, 1)) + ' %.')
print("K-Nearest Neighbors (KNN): accuracy = %3.1f %%" % accuracy_knn)
print("K-Nearest Neighbors (KNN): time = %3.3f s" % time_knn)
```
## Create Spark environment
```
# Create Spark Session
spark = SparkSession.builder \
.master("local[*]") \
.appName("MachineLearningIris") \
.getOrCreate()
```
## Reading Data
```
# Load Iris CSV dataset to Spark Dataframe
orig_data = spark.read.format("csv").options(sep=',',header='true',inferschema='true').\
load(data_path+"iris.data")
print("Original Dataframe read from CSV file")
#orig_data.dtypes
orig_data.show(5)
```
### Create Classifier Matrix
```
# ML libraries doesn't accept string column => everything should be numeric!
# create a numeric column "label" based on string column "class"
indexer = StringIndexer(inputCol="class", outputCol="label").fit(orig_data)
label_data = indexer.transform(orig_data)
# Save the inverse map from numeric "label" to string "class" to be used further in response
labelReverse = IndexToString().setInputCol("label")
# Show labeled dataframe with numeric lable
print("Dataframe with numeric lable")
label_data.show(5)
# Drop string column "class", no string column
label_data = label_data.drop("class")
# Most Machine Learning Lib inpute 2 columns: label (output) and feature (input)
# The label column is the result to train ML algorithm
# The feature column should join all parameters as a Vector
# Set the column names that is not part of features list
ignore = ['label']
# list will be all columns parts of features
list = [x for x in label_data.columns if x not in ignore]
# VectorAssembler mount the vector of features
assembler = VectorAssembler(
inputCols=list,
outputCol='features')
# Create final dataframe composed by label and a column of features vector
data = (assembler.transform(label_data).select("label","features"))
print("Final Dataframe suitable to classifier input format")
#data.printSchema()
data.show(5)
```
### Create Train and Test Dataset
```
# Split ramdomly the dataset into train and test group
# [0.7,0.3] => 70% for train and 30% for test
# [1.0,0.2] => 100% for train and 20% for test, not good, acuracy always 100%
# [0.1,0.02] => 10% for train and 2% for test, if big datasets
# 1234 is the random seed
(train, test) = data.randomSplit([train_sample, test_sample], 1234)
```
## Run Decision Tree
```
start_time_dt = time.time()
# impurity could be: entropy, gini'
trainer = DecisionTreeClassifier(featuresCol='features', labelCol='label', predictionCol='prediction', probabilityCol='probability',\
rawPredictionCol='rawPrediction', maxDepth=5, maxBins=32, minInstancesPerNode=1, minInfoGain=0.0,\
maxMemoryInMB=256, cacheNodeIds=False, checkpointInterval=10, impurity='gini', seed=None)
#trainer = LogisticRegression(maxIter=10, tol=1E-6, fitIntercept=True)
# train the model and get the result
model = trainer.fit(train)
result_dt = model.transform(test)
# compute accuracy on the test set against model
evaluator = MulticlassClassificationEvaluator(labelCol="label", predictionCol="prediction",\
metricName="accuracy")
accuracy_dt = evaluator.evaluate(result_dt) * 100
time_dt = time.time() - start_time_dt
print("Decision Tree: accuracy = %3.1f %%" % accuracy_dt)
print("Decision Tree: time = %3.3f s" % time_dt)
print("Decision Tree Final Result")
result_dt.show(5)
```
## Run Random Forest
```
start_time_rf = time.time()
trainer = RandomForestClassifier(featuresCol='features', labelCol='label', predictionCol='prediction', probabilityCol='probability',\
rawPredictionCol='rawPrediction', maxDepth=5, maxBins=32, minInstancesPerNode=1, minInfoGain=0.0,\
numTrees=50, featureSubsetStrategy='auto', seed=None, subsamplingRate=1.0,\
maxMemoryInMB=256, cacheNodeIds=False, checkpointInterval=10, impurity='gini')
# impurity could be: entropy, gini'
# numTrees= set the number of random trees to create
# train the model and get the result
model = trainer.fit(train)
result_rf = model.transform(test)
# compute accuracy on the test set against model
evaluator = MulticlassClassificationEvaluator(labelCol="label", predictionCol="prediction",\
metricName="accuracy")
accuracy_rf = evaluator.evaluate(result_rf) * 100
time_rf = time.time() - start_time_rf
print("Random Forest: accuracy = %3.1f %%" % accuracy_rf)
print("Random Forest: time = %3.3f s" % time_rf)
print("Decision Tree Final Result")
result_rf.show(5)
```
## Run Neural network Perceptron
```
start_time_pr = time.time()
# specify layers for the neural network
# parameter 1: input layer, should be the number of features
# parameter 2 and 3: the number os perceptron in two intermediate layers
# parameter 4: output layer should be the number os categories (labels)
layers = [4, 5, 5, 3]
# Create the trainer and set its parameters
# featuresCol=name_feature_column, labelCol=name_label_column
# maxIter=max_interaction, layers=list_number_perceptron
trainer = MultilayerPerceptronClassifier(featuresCol='features', labelCol='label',\
maxIter=100, layers=layers, blockSize=128, seed=1234)
# train the model and get the result
model = trainer.fit(train)
result_pr = model.transform(test)
print("Perceptron Final Result")
result_pr.show(5)
# compute accuracy on the test set against model
evaluator = MulticlassClassificationEvaluator(labelCol="label", predictionCol="prediction",\
metricName="accuracy")
accuracy_pr = evaluator.evaluate(result_pr) * 100
time_pr = time.time() - start_time_pr
print("Multilayer Perceptron: accuracy = %3.1f %%" % accuracy_pr)
print("Multilayer Perceptron: time = %3.3f s" % time_pr)
print("Perceptron final result with name of class")
labelReverse.transform(result_pr).show()
```
## Run Naive Bayes
```
start_time_nb = time.time()
# create the trainer and set its parameters
trainer = NaiveBayes(smoothing=1.0, modelType="multinomial")
#trainer = LogisticRegression(maxIter=10, tol=1E-6, fitIntercept=True)
#trainer = RandomForestClassifier(labelCol="indexedLabel", featuresCol="indexedFeatures", numTrees=10)
# train the model and get the result
model = trainer.fit(train)
result_nb = model.transform(test)
# compute accuracy on the test set against model
evaluator = MulticlassClassificationEvaluator(labelCol="label", predictionCol="prediction",\
metricName="accuracy")
accuracy_nb = evaluator.evaluate(result_nb) * 100
time_nb = time.time() - start_time_nb
print("Naive Bayes: accuracy = %3.1f %%" % accuracy_nb)
print("Naive Bayes: time = %3.3f s" % time_nb)
print("Naive Bayes Final Result")
result_nb.show()
print("Naive Bayes final result with name of class")
labelReverse.transform(result_nb).show()
```
## Run Suport Vector Machines (SVM)
```
start_time_svm = time.time()
# create the trainer and set its parameters
trainer = LinearSVC(featuresCol='features', labelCol='label',\
maxIter=100, regParam=0.1)
# LinearSVC classify ONLY in two classes
# To classify in more than 2 classes, the OneVsrest should be used
# Cloud use any kind of classifies
# instantiate the One Vs Rest Classifier.
ovr_trainer = OneVsRest(classifier=trainer)
# train the multiclass model.
model = ovr_trainer.fit(train)
# score the model on test data.
result_svm = model.transform(test)
# compute accuracy on the test set against model
evaluator = MulticlassClassificationEvaluator(labelCol="label", predictionCol="prediction",\
metricName="accuracy")
accuracy_svm = evaluator.evaluate(result_svm) * 100
time_svm = time.time() - start_time_svm
print("Suport Vector Machines (SVM): accuracy = %3.1f %%" % accuracy_svm)
print("Suport Vector Machines (SVM): time = %3.3f s" % time_svm)
print("Suport Vector Machines (SVM) Final Result")
result_svm.show()
```
## Sumary
```
print("==================================================================")
print("============= Compare Algorithm Acurancy and Time ===============")
print()
print(" Train sample = ",train_sample*100,"% Test sample = ",test_sample*100,"%")
print()
print("K-Nearest Neighbors (KNN): accuracy = %3.1f %% time = %3.3f s" % (accuracy_knn, time_knn))
print("Decision Tree: accuracy = %3.1f %% time = %3.3f s" % (accuracy_dt, time_dt))
print("Random Forest: accuracy = %3.1f %% time = %3.3f s" % (accuracy_rf, time_rf))
print("Multilayer Perceptron: accuracy = %3.1f %% time = %3.3f s" % (accuracy_pr, time_pr))
print("Naive Bayes: accuracy = %3.1f %% time = %3.3f s" % (accuracy_nb, time_nb))
print("Suport Vector Machines (SVM): accuracy = %3.1f %% time = %3.3f s" % (accuracy_svm, time_svm))
print("===================================================================")
spark.stop()
print("--- Execution time: %s seconds ---" % (time.time() - start_time))
```
| github_jupyter |
## Data Load / Read
- Explore
- Reads
- Frequency
- Percentage
- Use all
- Visualization
```
!head ../data/den1.fasta
!head ../data/den2.fasta
!head ../data/den3.fasta
!head ../data/den4.fasta
def readFASTA(inputfile):
"""Reads a sequence file and returns as string"""
with open(inputfile, "r") as seqfile:
# skip the name line
seq = seqfile.readline()
seq = seqfile.read()
seq = seq.replace("\n", "")
seq = seq.replace("\t", "")
return seq
# read
seq1 = readFASTA('../data/den1.fasta')
seq2 = readFASTA('../data/den2.fasta')
seq3 = readFASTA('../data/den3.fasta')
seq4 = readFASTA('../data/den4.fasta')
```
## Length
```
print("Length of DEN1: ", len(seq1))
print("Length of DEN2: ", len(seq2))
print("Length of DEN3: ", len(seq3))
print("Length of DEN4: ", len(seq4))
```
## Frequency
```
from collections import Counter
def basecount_fast(seq):
""""Count the frequencies of each bases in sequence including every letter"""
freqs = Counter(seq)
return freqs
print("Frequency of DEN1: ", basecount_fast(seq1))
print("Frequency of DEN2: ", basecount_fast(seq2))
print("Frequency of DEN3: ", basecount_fast(seq3))
print("Frequency of DEN4: ", basecount_fast(seq4))
def basecount(seq, useall=False, calfreqs_pc=False):
"""Count the frequencies of each bases in sequence including every letter"""
length = len(seq)
if calfreqs_pc:
freq_pc = {}
else:
base_counts = {}
if useall:
seqset = set(seq)
else:
seqset = ("A", "T", "C", "G")
for letter in seqset:
num = seq.count(letter)
if calfreqs_pc:
pc = num / length
freq_pc[letter] = pc
else:
base_counts[letter] = num
if calfreqs_pc:
return freq_pc
else:
return base_counts
print("Frequency of DEN1: ", basecount(seq1, useall=True))
print("Frequency of DEN2: ", basecount(seq2, useall=True))
print("Frequency of DEN3: ", basecount(seq3, useall=True))
print("Frequency of DEN4: ", basecount(seq4, useall=True))
print("Percentage of DEN1: ", basecount(seq1, calfreqs_pc=True))
print("Percentage of DEN2: ", basecount(seq2, calfreqs_pc=True))
print("Percentage of DEN3: ", basecount(seq3, calfreqs_pc=True))
print("Percentage of DEN4: ", basecount(seq4, calfreqs_pc=True))
f1 = basecount_fast(seq1)
f2 = basecount_fast(seq2)
f3 = basecount_fast(seq3)
f4 = basecount_fast(seq4)
import matplotlib.pyplot as plt
import seaborn as sns
plt.rcParams['figure.figsize'] = (8,6)
plt.rcParams['font.size'] = 14
plt.style.use('seaborn-whitegrid')
plt.bar(f1.keys(), f1.values())
plt.title("Nucleotide Frequency Distribution of DEN1")
plt.xlabel("Bases")
plt.ylabel("Frequency")
plt.tight_layout()
plt.show()
plt.bar(f2.keys(), f2.values())
plt.title("Nucleotide Frequency Distribution of DEN2")
plt.xlabel("Bases")
plt.ylabel("Frequency")
plt.tight_layout()
plt.show()
plt.bar(f3.keys(), f3.values())
plt.title("Nucleotide Frequency Distribution of DEN3")
plt.xlabel("Bases")
plt.ylabel("Frequency")
plt.tight_layout()
plt.show()
plt.bar(f4.keys(), f4.values())
plt.title("Nucleotide Frequency Distribution of DEN4")
plt.xlabel("Bases")
plt.ylabel("Frequency")
plt.tight_layout()
plt.show()
plt.pie(f1.values(), labels=f1.keys(), autopct='%1.1f%%', shadow=True)
plt.tight_layout()
plt.show()
plt.pie(f2.values(), labels=f2.keys(), autopct='%1.1f%%', shadow=True)
plt.tight_layout()
plt.show()
plt.pie(f3.values(), labels=f3.keys(), autopct='%1.1f%%', shadow=True)
plt.tight_layout()
plt.show()
plt.pie(f4.values(), labels=f4.keys(), autopct='%1.1f%%', shadow=True)
plt.tight_layout()
plt.show()
fig, ax = plt.subplots(nrows=2, ncols=2)
print(ax)
fig, ((ax1, ax2), (ax3, ax4)) = plt.subplots(nrows=2, ncols=2, sharex=True)
caption = "Figure: Nucleotide Frequency Distribution of Dengue"
# ax1
ax1.bar(f1.keys(), f1.values())
ax1.set_title("DEN1")
ax1.set_ylabel("Frequency")
# ax2
ax2.bar(f2.keys(), f2.values())
ax2.set_title("DEN2")
ax2.set_ylabel("Frequency")
# ax3
ax3.bar(f3.keys(), f3.values())
ax3.set_title("DEN3")
ax3.set_xlabel("Bases")
ax3.set_ylabel("Frequency")
# ax4
ax4.bar(f4.keys(), f4.values())
ax4.set_title("DEN4")
ax4.set_xlabel("Bases")
ax4.set_ylabel("Frequency")
# Caption
plt.figtext(0.5, 0.000001, caption, wrap=True, horizontalalignment='center', fontsize=18)
# layout
plt.tight_layout()
# plt.savefig('../output_figs/den_plot.png')
plt.show()
fig, ((ax1, ax2), (ax3, ax4)) = plt.subplots(nrows=2, ncols=2)
# ax1
ax1.bar(f1.keys(), f1.values())
ax1.set_title("DEN1")
ax1.set_ylabel("Frequency")
# ax2
ax2.pie(f1.values(), labels=f1.keys(), autopct='%1.1f%%', shadow=True)
ax2.set_title("DEN1")
# ax3
ax3.bar(f2.keys(), f2.values())
ax3.set_title("DEN2")
ax3.set_ylabel("Frequency")
# ax4
ax4.pie(f2.values(), labels=f2.keys(), autopct='%1.1f%%', shadow=True)
ax4.set_title("DEN2")
# layout
plt.tight_layout()
# plt.savefig('den.pdf')
plt.show()
def readFASTA(inputfile):
"""Reads a sequence file and returns as string"""
with open(inputfile, "r") as seqfile:
# skip the name line
seq = seqfile.readline()
seq = seqfile.read()
seq = seq.replace("\n", "")
seq = seq.replace("\t", "")
return seq
# read
seq1 = readFASTA('../data/den1.fasta')
seq2 = readFASTA('../data/den2.fasta')
seq3 = readFASTA('../data/den3.fasta')
seq4 = readFASTA('../data/den4.fasta')
len(seq1)
# den1
round((seq1.count("G")+seq1.count("C")) / len(seq1) * 100, 2)
len(seq2)
# den2
round((seq2.count("G")+seq2.count("C")) / len(seq2) * 100, 2)
# den3
round((seq3.count("G")+seq3.count("C")) / len(seq3) * 100, 2)
round((seq4.count("G")+seq4.count("C")) / len(seq4) * 100, 2)
def calculateGC(seq):
"""Take a sequence as input and calculate the GC %"""
gc = round((seq.count("G")+seq.count("C")) / len(seq) * 100, 2)
return gc
calculateGC(seq1)
def calculateAT(seq):
"""Take a sequence as input and calculate the AT %"""
at = round((seq.count("A")+seq.count("T")) / len(seq) * 100, 2)
return at
calculateAT(seq1)
# Sliding Window Analysis
len(seq1) / 10
calculateGC(seq1[0:2001])
calculateGC(seq1[2001:4001])
len(seq1)
sub = len(seq1) - 2000 + 1
range(len(seq1))
seq1[:10]
range(start=, stop=, step)
range(n) # n-1
range(n+1) # n
list(range(0, 10, 2 ))
seq1[0]
list(range(0, 10, 2))
list(range(0, 10735+1, 2000))
list(range(0, len(seq1)+1, 2000))
def subSeqGC(seq, windowsize=2000):
"""Returns sub-sequence GC Ration"""
res = []
for i in range(0, len(seq)-windowsize+1, windowsize):
subSeq = seq[i:i+windowsize]
gc = calculateGC(subSeq)
res.append(gc)
return res
subSeqGC(seq1)
len(subSeqGC(seq1))
subSeqGC(seq1, 500)
len(subSeqGC(seq1, 500))
import matplotlib.pyplot as plt
import seaborn as sns
plt.rcParams['figure.figsize'] = (8,6)
plt.rcParams['font.size'] = 14
plt.style.use('seaborn-whitegrid')
gc = subSeqGC(seq1, 300)
range(len(gc))
gc = subSeqGC(seq1, 300)
plt.plot(range(len(gc)), gc)
plt.title("Sub-sequence GC Distribution")
plt.xlabel("Base-pair Position")
plt.ylabel("% GC")
plt.tight_layout()
plt.savefig("../output_figs/sub_gc.pdf")
plt.show()
gc1 = subSeqGC(seq1, 300)
gc1
gc2 = subSeqGC(seq2, 300)
gc2
gc1 = subSeqGC(seq1, 300)
plt.plot(range(len(gc1)), gc1, 'ro', label="DEN1")
gc2 = subSeqGC(seq2, 300)
plt.plot(range(len(gc2)), gc2, 'bs', label="DEN2")
gc3 = subSeqGC(seq3, 300)
plt.plot(range(len(gc3)), gc3, color='purple', label="DEN3", linestyle='--')
gc4 = subSeqGC(seq4, 300)
plt.plot(range(len(gc4)), gc4, color='cyan', label="DEN4")
plt.title("Sub-sequence GC Distribution")
plt.xlabel("Base-pair Position")
plt.ylabel("% GC")
plt.tight_layout()
plt.legend()
# plt.savefig("../output_figs/sub_gc.pdf")
plt.show()
gc3 = subSeqGC(seq3, 300)
gc4 = subSeqGC(seq4, 300)
fig, ((ax1, ax2), (ax3, ax4)) = plt.subplots(nrows=2, ncols=2, sharex=True)
# ax1
gc1 = subSeqGC(seq1, 300)
ax1.plot(range(len(gc1)), gc1)
ax1.set_title("DEN1")
ax1.set_ylabel("% GC")
# ax2
gc2 = subSeqGC(seq2, 300)
ax2.plot(range(len(gc2)), gc2)
ax2.set_title("DEN2")
ax2.set_ylabel("% GC")
# ax3
gc3 = subSeqGC(seq3, 300)
ax3.plot(range(len(gc3)), gc3)
ax3.set_title("DEN3")
ax3.set_xlabel("Base-pair Position")
ax3.set_ylabel("% GC")
# ax4
gc4 = subSeqGC(seq4, 300)
ax4.plot(range(len(gc4)), gc4)
ax4.set_title("DEN4")
ax4.set_xlabel("Base-pair Position")
ax4.set_ylabel("% GC")
# layout
plt.tight_layout()
plt.savefig('../output_figs/den_plot.png')
plt.show()
def subSeqAT(seq, windowsize=2000):
"""Returns sub-sequence GC Ration"""
res = []
for i in range(0, len(seq)-windowsize+1, windowsize):
subSeq = seq[i:i+windowsize]
at = calculateAT(subSeq)
res.append(at)
return res
len(subSeqAT(seq1))
at = subSeqAT(seq1, 300)
plt.plot(range(len(at)), at)
plt.title("Sub-sequence AT Distribution")
plt.xlabel("Base-pair Position")
plt.ylabel("% AT")
plt.tight_layout()
# plt.savefig("../output_figs/sub_gc.pdf")
plt.show()
at1 = subSeqAT(seq1, 300)
plt.plot(range(len(at1)), at1, 'ro', label="DEN1")
at2 = subSeqAT(seq2, 300)
plt.plot(range(len(at2)), at2, 'bs', label="DEN2")
at3 = subSeqAT(seq3, 300)
plt.plot(range(len(at3)), at3, color='purple', label="DEN3", linestyle='--')
at4 = subSeqAT(seq4, 300)
plt.plot(range(len(at4)), at4, color='cyan', label="DEN4")
plt.title("Sub-sequence AT Distribution")
plt.xlabel("Base-pair Position")
plt.ylabel("% AT")
plt.tight_layout()
plt.legend()
plt.savefig("../output_figs/at_trend.pdf")
plt.show()
fig, ((ax1, ax2), (ax3, ax4)) = plt.subplots(nrows=2, ncols=2, sharex=True)
# ax1
at1 = subSeqAT(seq1, 300)
ax1.plot(range(len(at1)), at1)
ax1.set_title("DEN1")
ax1.set_ylabel("% AT")
# ax2
at2 = subSeqAT(seq2, 300)
ax2.plot(range(len(at2)), at2)
ax2.set_title("DEN2")
ax2.set_ylabel("% AT")
# ax3
at3 = subSeqAT(seq3, 300)
ax3.plot(range(len(at3)), at3)
ax3.set_title("DEN3")
ax3.set_xlabel("Base-pair Position")
ax3.set_ylabel("% AT")
# ax4
at4 = subSeqAT(seq4, 300)
ax4.plot(range(len(at4)), at4)
ax4.set_title("DEN4")
ax4.set_xlabel("Base-pair Position")
ax4.set_ylabel("% AT")
# layout
plt.tight_layout()
plt.savefig('../output_figs/den_plot.pdf')
plt.show()
```
## K-mer Analysis
```
A, T, C, G = Monomer
AA, TT, CC, CG, CT = Dimer
AAA, TTT, CTG = Codon=> Trimer
K-mer
k = 2 > AA, AT, TT
k = 3 > AAA
K = 4 > AAAA
range(start+1, stop, step)
list(range(10+1))
def buildKmers(seq, ksize):
# Store kmers
kmers = []
#
n_mers = len(seq) - ksize + 1
for i in range(n_mers):
kmer = seq[i:i+ksize]
kmers.append(kmer)
return kmers
seq = 'ATCGGTTAGGC'
len(seq)
len(seq) - 3 +1
# N errors
buildKmers('ATCGGTTAGGC',3)
def readFASTA(inputfile):
"""Reads a sequence file and returns as string"""
with open(inputfile, "r") as seqfile:
# skip the name line
seq = seqfile.readline()
seq = seqfile.read()
seq = seq.replace("\n", "")
seq = seq.replace("\t", "")
return seq
seq1 = readFASTA('../data/den1.fasta')
seq2 = readFASTA('../data/den2.fasta')
seq3 = readFASTA('../data/den3.fasta')
seq4 = readFASTA('../data/den4.fasta')
km1 = buildKmers(seq1, 2)
km2 = buildKmers(seq2, 2)
km3 = buildKmers(seq3, 2)
km4 = buildKmers(seq1, 2)
from collections import Counter
def kmerFrequency(kmers):
freq = Counter(kmers)
return freq
f1 = kmerFrequency(km1)
f1
import matplotlib.pyplot as plt
plt.bar(f1.keys(), f.values())
plt.show()
f2 = kmerFrequency(km2)
f2
plt.bar(f2.keys(), f2.values())
plt.show()
```
| github_jupyter |
# We're going to combine all components into a single model
```
import numpy as np
import matplotlib.pyplot as plt
import theano.tensor as tt
import lightkurve as lk
from astropy.units import cds
from astropy import units as u
import seaborn as sns
import corner
import pystan
import pandas as pd
import pickle
import glob
from astropy.io import ascii
import os
import sys
import pymc3 as pm
from pymc3.gp.util import plot_gp_dist
import arviz
import warnings
warnings.filterwarnings('ignore')
cpu = 'bear'
```
## Build the model
```
class model():
def __init__(self, f, n0_, n1_, n2_, deltanu_):
self.f = f
self.n0 = n0_
self.n1 = n1_
self.n2 = n2_
self.npts = len(f)
self.M = [len(n0_), len(n1_), len(n2_)]
self.deltanu = deltanu_
def epsilon(self, i):
eps = tt.zeros((3,3))
eps0 = tt.set_subtensor(eps[0][0], 1.)
eps1 = tt.set_subtensor(eps[1][0], tt.sqr(tt.cos(i)))
eps1 = tt.set_subtensor(eps1[1], 0.5 * tt.sqr(tt.sin(i)))
eps2 = tt.set_subtensor(eps[2][0], 0.25 * tt.sqr((3. * tt.sqr(tt.cos(i)) - 1.)))
eps2 = tt.set_subtensor(eps2[1], (3./8.) * tt.sqr(tt.sin(2*i)))
eps2 = tt.set_subtensor(eps2[2], (3./8.) * tt.sin(i)**4)
eps = tt.set_subtensor(eps[0], eps0)
eps = tt.set_subtensor(eps[1], eps1)
eps = tt.set_subtensor(eps[2], eps2)
return eps
def lor(self, freq, h, w):
return h / (1.0 + 4.0/tt.sqr(w)*tt.sqr((self.f - freq)))
def mode(self, l, freqs, hs, ws, eps, split=0):
for idx in range(self.M[l]):
for m in range(-l, l+1, 1):
self.modes += self.lor(freqs[idx] + (m*split),
hs[idx] * eps[l,abs(m)],
ws[idx])
def model(self, p, theano=True):
f0, f1, f2, g0, g1, g2, h0, h1, h2, split, i, phi = p
# Unpack background parameters
loga = phi[0]
logb = phi[1]
logc = phi[2]
logd = phi[3]
logj = phi[4]
logk = phi[5]
white = phi[6]
scale = phi[7]
nyq = phi[8]
# Calculate the modes
eps = self.epsilon(i)
self.modes = np.zeros(self.npts)
self.mode(0, f0, h0, g0, eps)
self.mode(1, f1, h1, g1, eps, split)
self.mode(2, f2, h2, g2, eps, split)
self.modes *= self.get_apodization(nyq)
#Calculate the background
self.back = self.get_background(loga, logb, logc, logd, logj, logk,
white, scale, nyq)
#Create the model
self.mod = self.modes + self.back
if theano:
return self.mod
else:
return self.mod.eval()
# Small separations are fractional
def asymptotic(self, n, numax, alpha, epsilon, d=0.):
nmax = (numax / self.deltanu) - epsilon
curve = (alpha/2.)*(n-nmax)*(n-nmax)
return (n + epsilon + d + curve) * self.deltanu
def f0(self, p):
numax, alpha, epsilon, d01, d02 = p
return self.asymptotic(self.n0, numax, alpha, epsilon, 0.)
def f1(self, p):
numax, alpha, epsilon, d01, d02 = p
return self.asymptotic(self.n1, numax, alpha, epsilon, d01)
def f2(self, p):
numax, alpha, epsilon, d01, d02 = p
return self.asymptotic(self.n2+1, numax, alpha, epsilon, -d02)
def gaussian(self, freq, numax, w, A):
return A * tt.exp(-0.5 * tt.sqr((freq - numax)) / tt.sqr(w))
def A0(self, f, p, theano=True):
numax, w, A, V1, V2 = p
height = self.gaussian(f, numax, w, A)
if theano:
return height
else:
return height.eval()
def A1(self, f, p, theano=True):
numax, w, A, V1, V2 = p
height = self.gaussian(f, numax, w, A)*V1
if theano:
return height
else:
return height.eval()
def A2(self, f, p, theano=True):
numax, w, A, V1, V2 = p
height = self.gaussian(f, numax, w, A)*V2
if theano:
return height
else:
return height.eval()
def harvey(self, a, b, c):
harvey = 0.9*tt.sqr(a)/b/(1.0 + tt.pow((self.f/b), c))
return harvey
def get_apodization(self, nyquist):
x = (np.pi * self.f) / (2 * nyquist)
return tt.sqr((tt.sin(x)/x))
def get_background(self, loga, logb, logc, logd, logj, logk, white, scale, nyq):
background = np.zeros(len(self.f))
background += self.get_apodization(nyq) * scale \
* (self.harvey(tt.pow(10, loga), tt.pow(10, logb), 4.) \
+ self.harvey(tt.pow(10, logc), tt.pow(10, logd), 4.) \
+ self.harvey(tt.pow(10, logj), tt.pow(10, logk), 2.))\
+ white
return background
```
### Build the range
```
nmodes = 4
nbase = 16
n0_ = np.arange(nmodes)+nbase
n1_ = np.copy(n0_)
n2_ = np.copy(n0_) - 1.
fs = .05
nyq = (0.5 * (1./58.6) * u.hertz).to(u.microhertz).value
ff = np.arange(fs, nyq, fs)
```
### Build the frequencies
```
deltanu_ = 60.
numax_= 1150.
alpha_ = 0.01
epsilon_ = 1.1
d01_ = deltanu_/2. / deltanu_
d02_ = 6. / deltanu_
mod = model(ff, n0_, n1_, n2_, deltanu_)
init_f = [numax_, alpha_, epsilon_, d01_, d02_]
f0_true = mod.f0(init_f)
f1_true = mod.f1(init_f)
f2_true = mod.f2(init_f)
sigma0_ = 1.5
sigma1_ = 2.0
sigma2_ = .5
f0_ = mod.f0(init_f) + np.random.randn(len(f0_true)) * sigma0_
f1_ = mod.f1(init_f) + np.random.randn(len(f1_true)) * sigma1_
f2_ = mod.f2(init_f) + np.random.randn(len(f2_true)) * sigma2_
lo = f2_.min() - .25*deltanu_
hi = f1_.max() + .25*deltanu_
sel = (ff > lo) & (ff < hi)
f = ff[sel]
```
### Reset model for new frequency range
```
mod = model(f, n0_, n1_, n2_, deltanu_)
```
### Build the linewidths
```
def kernel(n, rho, L):
return rho**2 * np.exp(-0.5 * np.subtract.outer(n,n)**2 / L**2)
m_ = .5
c_ = .5
rho_ = 0.1
L_ = 0.3
fs = np.concatenate((f0_, f1_, f2_))
fs -= fs.min()
nf = fs/fs.max()
mu_ = m_ * nf + c_
Sigma_ = kernel(nf, rho_, L_)
lng0_ = np.random.multivariate_normal(mu_, Sigma_)
widths = [np.exp(lng0_)[0:len(f0_)],
np.exp(lng0_)[len(f0_):len(f0_)+len(f1_)],
np.exp(lng0_)[len(f0_)+len(f1_):]]
nf_ = nf[:,None]
```
### Build the mode amplitudes
```
w_ = (0.25 * numax_)/2.355
V1_ = 1.2
V2_ = 0.7
A_ = 10.
init_h =[numax_, #numax
w_, #envelope width
A_, #envelope amplitude
V1_, #dipole visibility
V2_ #ocotopole visibility
]
sigmaA_ = .2
amps = [mod.A0(f0_, init_h, theano=False) + np.random.randn(len(f0_)) * sigmaA_,
mod.A1(f1_, init_h, theano=False) + np.random.randn(len(f0_)) * sigmaA_,
mod.A2(f2_, init_h, theano=False) + np.random.randn(len(f0_)) * sigmaA_]
```
### Build the background
```
labels=['loga','logb','logc','logd','logj','logk','white','scale','nyq']
phi_ = [ 1.6, 2.6, 1.6, 3.0, 1.7, 0.5, 0.4, 1., nyq]
phi_sigma = np.genfromtxt('phi_sigma.txt')
phi_cholesky = np.linalg.cholesky(phi_sigma)
```
### Construct the model
```
split_ = 1.
incl_ = np.pi/4.
init_m =[f0_, # l0 modes
f1_, # l1 modes
f2_, # l2 modes
widths[0], # l0 widths
widths[1], # l1 widths
widths[2], # l2 widths
amps[0]**2 * 2.0 / np.pi / widths[0] ,# l0 heights
amps[1]**2 * 2.0 / np.pi / widths[1] ,# l1 heights
amps[2]**2 * 2.0 / np.pi / widths[2] ,# l2 heights
split_, # splitting
incl_, # inclination angle
phi_ # background parameters
]
p = mod.model(init_m, theano=False)*np.random.chisquare(2., size=len(f))/2
with plt.style.context(lk.MPLSTYLE):
plt.plot(f, p)
plt.plot(f, mod.model(init_m, theano=False), lw=3)
plt.yscale('log')
plt.xscale('log')
if cpu == 'bear':
plt.savefig('data.png')
else: plt.show()
```
## First lets fit the background alone...
```
if cpu != 'bear':
pm_model = pm.Model()
with pm_model:
# Background treatment
phi = pm.MvNormal('phi', mu=phi_, chol=phi_cholesky, testval=phi_, shape=len(phi_))
# Construct the model
fit = mod.model([*init_m[:11], phi])
like = pm.Gamma('like', alpha=1, beta=1.0/fit, observed=p)
trace = pm.sample(target_accept=.99)
if cpu != 'bear':
pm.summary(trace)
if cpu != 'bear':
labels=['loga','logb','logc','logd','logj','logk',
'white','scale','nyq']
verbose=[r'$\log_{10}a$',r'$\log_{10}b$',
r'$\log_{10}c$',r'$\log_{10}d$',
r'$\log_{10}j$',r'$\log_{10}k$',
'white','scale',r'$\nu_{\rm nyq}$']
phichain = np.array([trace['phi'][:,idx] for idx in range(len(phi_))]).T
truth = phi_
corner.corner(phichain, truths=truth, show_titles=True, labels=verbose)
plt.show()
```
# Now lets try and fit this
## Building the initial guesses
```
init = {}
init['numax'] = numax_
init['alpha'] = alpha_
init['epsilon'] = epsilon_
init['d01'] = d01_
init['d02'] = d02_
init['sigma0'] = sigma0_
init['sigma1'] = sigma1_
init['sigma2'] = sigma2_
init['m'] = m_
init['c'] = c_
init['rho'] = rho_
init['L'] = L_
init['w'] = (0.25 * numax_)/2.355
init['A'] = np.sqrt(np.pi * np.nanmax(p) / 2.)
init['V1'] = V1_
init['V2'] = V2_
init['sigmaA'] = sigmaA_
init['a0'] = amps[0]
init['a1'] = amps[1]
init['a2'] = amps[2]
init['xsplit'] = split_ * np.sin(incl_)
init['cosi'] = np.cos(incl_)
```
## Building the model
```
nf_
pm_model = pm.Model()
with pm_model:
# Mode locations
numax = pm.Normal('numax', mu = init['numax'], sigma = 10., testval = init['numax'])
alpha = pm.Lognormal('alpha', mu = np.log(init['alpha']), sigma = 0.01, testval = init['alpha'])
epsilon = pm.Normal('epsilon', mu = init['epsilon'], sigma = 1., testval = init['epsilon'])
d01 = pm.Lognormal('d01', mu = np.log(init['d01']), sigma = 0.1, testval = init['d01'])
d02 = pm.Lognormal('d02', mu = np.log(init['d02']), sigma = 0.1, testval = init['d02'])
sigma0 = pm.HalfCauchy('sigma0', beta = 2., testval = init['sigma0'])
sigma1 = pm.HalfCauchy('sigma1', beta = 2., testval = init['sigma1'])
sigma2 = pm.HalfCauchy('sigma2', beta = 2., testval = init['sigma2'])
f0 = pm.Normal('f0', mu = mod.f0([numax, alpha, epsilon, d01, d02]), sigma = sigma0, shape=len(f0_))
f1 = pm.Normal('f1', mu = mod.f1([numax, alpha, epsilon, d01, d02]), sigma = sigma1, shape=len(f1_))
f2 = pm.Normal('f2', mu = mod.f2([numax, alpha, epsilon, d01, d02]), sigma = sigma2, shape=len(f2_))
# Mode Linewidths
m = pm.Normal('m', mu = init['m'], sigma = 1., testval = init['m'])
c = pm.Normal('c', mu = init['c'], sigma = 1., testval = init['c'])
rho = pm.Lognormal('rho', mu = np.log(init['rho']), sigma = 0.1, testval = init['rho'])
ls = pm.TruncatedNormal('ls', mu = np.log(init['L']), sigma = 0.1, lower=0., testval = init['L'])
mu = pm.gp.mean.Linear(coeffs = m, intercept = c)
cov = tt.sqr(rho) * pm.gp.cov.ExpQuad(1, ls = ls)
gp = pm.gp.Latent(cov_func = cov, mean_func = mu)
lng = gp.prior('lng', X = nf_)
g0 = pm.Deterministic('g0', tt.exp(lng)[0:len(f0_)])
g1 = pm.Deterministic('g1', tt.exp(lng)[len(f0_):len(f0_)+len(f1_)])
g2 = pm.Deterministic('g2', tt.exp(lng)[len(f0_)+len(f1_):])
# Mode Amplitude & Height
w = pm.Lognormal('w', mu = np.log(init['w']), sigma = 10., testval=init['w'])
A = pm.Lognormal('A', mu = np.log(init['A']), sigma = 1., testval=init['A'])
V1 = pm.Lognormal('V1', mu = np.log(init['V1']), sigma = 0.1, testval=init['V1'])
V2 = pm.Lognormal('V2', mu = np.log(init['V2']), sigma = 0.1, testval=init['V2'])
sigmaA = pm.HalfCauchy('sigmaA', beta = 1., testval = init['sigmaA'])
Da0 = pm.Normal('Da0', mu = 0., sigma = 1., shape=len(f0_))
Da1 = pm.Normal('Da1', mu = 0., sigma = 1., shape=len(f1_))
Da2 = pm.Normal('Da2', mu = 0., sigma = 1., shape=len(f2_))
a0 = pm.Deterministic('a0', sigmaA * Da0 + mod.A0(f0_, [numax, w, A, V1, V2]))
a1 = pm.Deterministic('a1', sigmaA * Da1 + mod.A1(f1_, [numax, w, A, V1, V2]))
a2 = pm.Deterministic('a2', sigmaA * Da2 + mod.A2(f2_, [numax, w, A, V1, V2]))
h0 = pm.Deterministic('h0', 2*tt.sqr(a0)/np.pi/g0)
h1 = pm.Deterministic('h1', 2*tt.sqr(a1)/np.pi/g1)
h2 = pm.Deterministic('h2', 2*tt.sqr(a2)/np.pi/g2)
# Mode splitting
xsplit = pm.HalfNormal('xsplit', sigma=2.0, testval = init['xsplit'])
cosi = pm.Uniform('cosi', 0., 1., testval = init['cosi'])
i = pm.Deterministic('i', tt.arccos(cosi))
split = pm.Deterministic('split', xsplit/tt.sin(i))
# Background treatment
phi = pm.MvNormal('phi', mu = phi_, chol = phi_cholesky, testval = phi_, shape = len(phi_))
# Construct model
fit = mod.model([f0, f1, f2, g0, g1, g2, h0, h1, h2, split, i, phi])
like = pm.Gamma('like', alpha = 1., beta = 1./fit, observed = p)
for RV in pm_model.basic_RVs:
print(RV.name, RV.logp(pm_model.test_point))
with pm_model:
trace = pm.sample(chains=4,
target_accept=.99,
start = init,
init = 'adapt_diag')
df = pm.backends.tracetab.trace_to_dataframe(trace)
df.to_csv('testchains.csv')
pm.summary(trace)
labels = ['numax','alpha','epsilon','d01','d02',
'split','i']
chain = np.array([trace[label] for label in labels])
truths = [numax_, alpha_, epsilon_, d01_, d02_,
split_, incl_]
corner.corner(chain.T, labels=labels, truths=truths, quantiles=[.16, .5, .84], truth_color='r',show_titles=True)
if cpu == 'bear':
plt.savefig('corner1.png')
else: plt.show()
labels = ['sigma0','sigma1','sigma2','w','A','V1','V2','sigmaA']
chain = np.array([trace[label] for label in labels])
truths = [sigma0_, sigma1_, sigma2_,w_, A_, V1_, V2_, 0.2]
corner.corner(chain.T, labels=labels, truths=truths, quantiles=[.16, .5, .84], truth_color='r',show_titles=True)
if cpu == 'bear':
plt.savefig('corner2.png')
else: plt.show()
with plt.style.context(lk.MPLSTYLE):
res_m = [np.median(trace[label], axis=0) for label in ['f0','f1','f2','g0','g1','g2',
'h0','h1','h2','split','i','phi']]
plt.plot(f, p)
plt.plot(f, mod.model(res_m, theano=False), lw=3)
if cpu == 'bear':
plt.savefig('modelfit.png')
else: plt.show()
fig, ax = plt.subplots()
res = [np.median(trace[label]) for label in ['numax', 'w', 'A', 'V1','V2']]
resls = [np.median(trace[label],axis=0) for label in ['a0','a1','a2']]
ax.plot(f0_, mod.A0(f0_, res,theano=False), label='0 Trend',lw=2, zorder=1)
ax.plot(f1_, mod.A1(f1_, res,theano=False), label='1 Trend',lw=2, zorder=1)
ax.plot(f2_, mod.A2(f2_, res,theano=False), label='2 Trend',lw=2, zorder=1)
ax.scatter(f0_, amps[0], marker='^',label='0 Errd', s=50, zorder=2)
ax.scatter(f1_, amps[1], marker='*',label='1 Errd', s=50, zorder=2)
ax.scatter(f2_, amps[2], marker='o',label='2 Errd', s=50, zorder=2)
ax.plot(f0_, mod.A0(f0_, init_h, theano=False), label='0 Pure',lw=2, zorder=1)
ax.plot(f1_, mod.A1(f1_, init_h, theano=False), label='1 Pure',lw=2, zorder=1)
ax.plot(f2_, mod.A2(f2_, init_h, theano=False), label='2 Pure',lw=2, zorder=1)
ax.scatter(f0_, resls[0], marker='^',label='0 mod', s=10, zorder=3)
ax.scatter(f1_, resls[1], marker='*',label='1 mod', s=10, zorder=3)
ax.scatter(f2_, resls[2], marker='o',label='2 mod', s=10, zorder=3)
ax.legend(loc='upper center', ncol=4, bbox_to_anchor=(0.5, 1.3))
ax.set_xlabel('Frequency')
ax.set_ylabel('Amplitude')
if cpu == 'bear':
plt.savefig('amplitudefit.png')
else: plt.show()
fig, ax = plt.subplots()
res = [np.median(trace[label]) for label in ['numax', 'alpha', 'epsilon','d01','d02']]
resls = [np.median(trace[label],axis=0) for label in ['f0','f1','f2']]
stdls = [np.std(trace[label],axis=0) for label in ['f0','f1','f2']]
ax.plot(mod.f0(res)%deltanu_, n0_, label='0 Trend',lw=2, zorder=1)
ax.plot(mod.f1(res)%deltanu_, n1_, label='1 Trend',lw=2, zorder=1)
ax.plot(mod.f2(res)%deltanu_, n2_, label='2 Trend',lw=2, zorder=1)
ax.scatter(f0_%deltanu_, n0_, marker='^',label='0 Truth (glitch)', s=50, zorder=2)
ax.scatter(f1_%deltanu_, n1_, marker='*',label='1 Truth (glitch)', s=50, zorder=2)
ax.scatter(f2_%deltanu_, n2_, marker='o',label='2 Truth (glitch)', s=50, zorder=2)
ax.plot(f0_true%deltanu_, n0_, alpha=.5, label='0 Truth (pure)', lw=2, zorder=1)
ax.plot(f1_true%deltanu_, n1_, alpha=.5, label='1 Truth (pure)', lw=2, zorder=1)
ax.plot(f2_true%deltanu_, n2_, alpha=.5, label='2 Truth (pure)', lw=2, zorder=1)
ax.scatter(resls[0]%deltanu_, n0_, marker='^',label='0 mod', s=10, zorder=3)
ax.scatter(resls[1]%deltanu_, n1_, marker='*',label='1 mod', s=10, zorder=3)
ax.scatter(resls[2]%deltanu_, n2_, marker='o',label='2 mod', s=10, zorder=3)
ax.set_xlabel(r'Frequency mod $\Delta\nu$')
ax.set_ylabel('Overtone order n')
ax.legend(loc='upper center', ncol=4, bbox_to_anchor=(0.5, 1.3))
if cpu == 'bear':
plt.savefig('frequencyfit.png')
else: plt.show()
nflin = np.linspace(nf.min(), nf.max(), 100)
fslin = np.linspace(fs.min(), fs.max(), 100)+f2_.min()
mulin = nflin * np.median(trace['m']) + np.median(trace['c'])
with pm_model:
f_pred = gp.conditional("f_pred", nflin[:,None])
expf_pred = pm.Deterministic('expf_pred', tt.exp(f_pred))
pred_samples = pm.sample_posterior_predictive(trace, vars=[expf_pred], samples=1000)
with plt.style.context(lk.MPLSTYLE):
fig, ax = plt.subplots()
plot_gp_dist(ax, pred_samples['expf_pred'], fslin, palette='viridis', fill_alpha=.05)
ax.plot(fslin, np.exp(mulin), label='Mean Trend', lw=2, ls='-.', alpha=.5, zorder=0)
ax.scatter(f0_, widths[0], label='truth', ec='k',s=50,zorder=5)
ax.scatter(f1_, widths[1], label='truth 1', ec='k',s=50,zorder=5)
ax.scatter(f2_, widths[2], label='truth 2', ec='k',s=50,zorder=5)
ax.scatter(f0_, np.median(trace['g0'],axis=0), marker='^', label='mod', s=10,zorder=5)
ax.scatter(f1_, np.median(trace['g1'],axis=0), marker='*', label='mod 1', s=10,zorder=5)
ax.scatter(f2_, np.median(trace['g2'],axis=0), marker='o', label='mod 2', s=10,zorder=5)
ax.errorbar(f0_, np.median(trace['g0'],axis=0), yerr=np.std(trace['g0'],axis=0), fmt='|', c='k', lw=3, alpha=.5)
ax.errorbar(f1_, np.median(trace['g1'],axis=0), yerr=np.std(trace['g1'],axis=0), fmt='|', c='k', lw=3, alpha=.5)
ax.errorbar(f2_, np.median(trace['g2'],axis=0), yerr=np.std(trace['g2'],axis=0), fmt='|', c='k', lw=3, alpha=.5)
ax.legend(loc='upper center', ncol=2, bbox_to_anchor=(0.5, 1.3))
if cpu == 'bear':
plt.savefig('widthfit.png')
else: plt.show()
labels=['loga','logb','logc','logd','logj','logk',
'white','scale','nyq']
verbose=[r'$\log_{10}a$',r'$\log_{10}b$',
r'$\log_{10}c$',r'$\log_{10}d$',
r'$\log_{10}j$',r'$\log_{10}k$',
'white','scale',r'$\nu_{\rm nyq}$']
phichain = np.array([trace['phi'][:,idx] for idx in range(len(phi_))]).T
truth = phi_
corner.corner(phichain, truths=truth, show_titles=True, labels=verbose)
if cpu == 'bear':
plt.savefig('backcorner.png')
else: plt.show()
residual = p/mod.model(res_m, theano=False)
sns.distplot(residual, label='Model')
sns.distplot(np.random.chisquare(2, size=10000)/2, label=r'Chi22')
plt.legend()
sys.exit()
```
| github_jupyter |
# Examining color selection at z~5
This notebook explores various parameters that affect optical color selection of quasars at z~5. The most prominent effect is the dependence of the selection efficiency on Lya equivalent width. Some discussion of this appears in McGreer et al. 2013,2017; in fact this notebook is used as the basis for the discussion in the appendix of McGreer et al. 2017.
First, the necessary imports from simqso to be able to generate arbitrary simulation grids and associated spectra.
```
%pylab inline
import matplotlib.gridspec as gridspec
from astropy.cosmology import WMAP9
from simqso.sqgrids import *
from simqso import sqbase
from simqso import hiforest
from simqso.sqrun import buildSpectraBySightLine,buildQsoSpectrum
from simqso import sqphoto
from simqso import sqmodels
random.seed(123)
```
Set the wavelength grid to cover the optical/IR range at low resolution.
```
# cover 3000A to 5um at R=500
wave = sqbase.fixed_R_dispersion(3000,5e4,500)
```
Build a set of IGM absorption sightlines based on the model given in McGreer et al. 2013.
```
forestModel = sqmodels.forestModels['McGreer+2013']
forestSpec = hiforest.IGMTransmissionGrid(wave,forestModel,100,zmax=5.3)
forestVar = HIAbsorptionVar(forestSpec)
```
## Selection dependence on Lya EW and UV continuum slope
This first simulation utilizes a 4-d grid where the last two dimensions are Lya equivalent width and continuum shape. The grid spans the luminosity and redshift ranges of the z~5 quasar sample from CFHTLS discussed in McGreer et al. 2017. The Lya EW is a grid of 5 fixed values, and the UV continuum spectral index is a grid of 6 fixed values.
```
lyaews = [0.,15,30,60,120]
uvslopes = [-1.5,-1.0,-0.6,-0.4,-0.2,0.0]
nperbin = 100
M = AbsMagVar(UniformSampler(-28,-23),restWave=1450)
z = RedshiftVar(UniformSampler(4.65,5.35))
lya = GaussianLineEqWidthVar(FixedSampler(lyaews),'LyaEW',1215.67,5.0)
uvslope = QsoSimVar(FixedSampler(uvslopes),'uvslope')
qsoGrid = QsoSimGrid([M,z,lya,uvslope],(10,7,5,6),nperbin,fixed_vars=['LyaEW','uvslope'],
cosmo=WMAP9,units='luminosity')
```
The far-UV slope is set to -1.5 (e.g., Lusso et al. 2016) with a breakpoint at 1100 Ang.
```
contVar = BrokenPowerLawContinuumVar([ConstSampler(-1.5),
FixedSampler(qsoGrid.uvslope)],[1100.])
qsoGrid.addVars([contVar,forestVar])
```
The spectra will be mapped to photometry that matches the characteristics of the CFHTLS Wide survey. The SDSS bandpasses are added for comparison.
```
qsoGrid.loadPhotoMap([('CFHT','CFHTLS_Wide'),('SDSS','Legacy')])
```
Now run the simulation and generate the spectra. This takes a while.
```
_ = buildSpectraBySightLine(wave,qsoGrid,verbose=5)
```
With the spectra in hand, convolve them with the survey bandpasses to get simulated photometry. Add photometric noise to get realistic photometry.
```
photoData = sqphoto.calcObsPhot(qsoGrid.synFlux,qsoGrid.photoMap)
qsoGrid.addData(photoData)
```
Now plot the "observed" photometry in r-i/i-z color space, which is the main selection cut. One panel is shown for each of the fixed Lya EW values. Note how decreasing the EW drives the colors away from the selection cuts (shown as black lines, solid line is "weak" criteria from McGreer et al. 2017, dashed line is "strict" criteria). The points are downsampled by a factor of 10 for clarity, and color-coded by the UV spectral index.
```
g_mags = qsoGrid.asGrid('obsMag') # last axis is ugriz
g_clrs = -diff(g_mags,axis=-1) # last axis is u-g,g-r,r-i,i-z
g_z = qsoGrid.asGrid('z')
g_a = qsoGrid.asGrid('uvslope')
fig = figure(figsize=(11,7))
for ew_i in range(5):
ax = subplot(2,3,ew_i+1)
pts = scatter(g_clrs[:,:,4-ew_i,:,::10,2],g_clrs[:,:,4-ew_i,:,::10,3],
c=g_a[:,:,4-ew_i,:,::10],s=7,cmap='coolwarm_r')
ax.set_title('EW = %d' % qsoGrid.gridCenters[2][4-ew_i])
ax.plot([1.3,1.3],[-0.5,0.55],c='k')
ax.plot([1.3,2.5],[0.55,0.55],c='k')
ri = linspace(1.3,1.7,2)
ax.plot(ri,polyval([0.875,0.15],ri-1.3),c='k',ls='--',lw=2)
ax.plot([1.7,2.5],[0.5,0.5],c='k',ls='--',lw=2)
ax.set_xlim(1.0,2.5)
ax.set_ylim(-0.35,0.9)
cbax = fig.add_axes([0.65,0.25,0.25,0.05])
cb = fig.colorbar(pts,cax=cbax,orientation='horizontal')
_ = cb.ax.set_xlabel('UV spectral index')
```
For comparison, here are the simulated colors (no noise added) in the SDSS bands. This is to show that with SDSS colors the effect is even more severe, and it is more difficult to select low-EW quasars.
```
g_mags = qsoGrid.asGrid('synMag') # last axis is ugriz
g_clrs = -diff(g_mags,axis=-1) # last axis is u-g,g-r,r-i,i-z
fig = figure(figsize=(11,7))
for ew_i in range(5):
ax = subplot(2,3,ew_i+1)
pts = scatter(g_clrs[:,:,4-ew_i,:,::10,2+5],g_clrs[:,:,4-ew_i,:,::10,3+5],
c=g_a[:,:,4-ew_i,:,::10],s=7,cmap='coolwarm_r')
ax.set_title('EW = %d' % qsoGrid.gridCenters[2][4-ew_i])
ax.plot([1.3,1.3],[-0.5,0.55],c='k')
ax.plot([1.3,2.5],[0.55,0.55],c='k')
ri = linspace(1.3,1.7,2)
ax.plot(ri,polyval([0.875,0.15],ri-1.3),c='k',ls='--',lw=2)
ax.plot([1.7,2.5],[0.5,0.5],c='k',ls='--',lw=2)
ax.set_xlim(1.0,2.5)
ax.set_ylim(-0.35,0.9)
cbax = fig.add_axes([0.65,0.25,0.25,0.05])
cb = fig.colorbar(pts,cax=cbax,orientation='horizontal')
_ = cb.ax.set_xlabel('UV spectral index')
```
Calculate the selection efficiency by applying the color criteria and determining the fraction of quasars passing those criteria within each grid cell.
```
mags = qsoGrid.asGrid('obsMag')
fluxes = qsoGrid.asGrid('obsFlux')
ferrs = qsoGrid.asGrid('obsFluxErr')
snr_u = fluxes[...,0]/ferrs[...,0]
snr_g = fluxes[...,1]/ferrs[...,1]
g_minus_r = mags[...,1] - mags[...,2]
r_minus_i = mags[...,2] - mags[...,3]
i_minus_z = mags[...,3] - mags[...,4]
# "weak" criteria from McGreer+17
weak_color_sel = ( (snr_u < 2.2) &
( (g_minus_r > 1.8) | (snr_g < 2.2) ) &
(r_minus_i>1.3) &
( (i_minus_z > -0.5) & (i_minus_z<0.55) ) )
# "strict" criteria
strict_color_sel = ( weak_color_sel &
(i_minus_z < 0.15 + 0.875*(r_minus_i-1.3)) &
(i_minus_z < 0.5) )
```
Display the efficiency by plotting absolute mag vs. redshift, split into panels corresponding to each of the fixed Lya EW and UV spectral index values. The color coding is the selected fraction.
```
gs = gridspec.GridSpec(5,6)
gs.update(wspace=0.07,hspace=0.25)
figure(figsize=(14,11))
for i in range(5):
for j in range(6):
ax = plt.subplot(gs[i,j])
if j>0: ax.yaxis.set_ticklabels([])
if i<4: ax.xaxis.set_ticklabels([])
fsel = weak_color_sel[:,:,-(i+1),j].sum(axis=-1) / float(nperbin)
imshow(fsel,vmin=0,vmax=1,extent=[4.7,5.3,-24,-29],
interpolation='nearest',aspect='auto',cmap=cm.hot_r)
title(r'EW=%d, $\alpha$=%.1f' %
(qsoGrid.gridCenters[2][-(i+1)],qsoGrid.gridCenters[3][j]),
size=12,weight='bold')
```
A line plot showing the variation of the efficiency with Lya EW at the nominal UV slope.
```
from matplotlib import ticker
import matplotlib
mpl.rcParams['mathtext.fontset'] = 'cm'
mpl.rcParams['mathtext.rm'] = 'serif'
M_i = 7
fig = figure(figsize=(7.8,3.0))
fig.subplots_adjust(0.07,0.14,0.99,0.90,0.28)
ax1 = subplot(121)
for color_sel,ls in zip([weak_color_sel,strict_color_sel],['-','--']):
fsel = color_sel[M_i,:,:,3].sum(axis=-1) / float(nperbin)
for i,lyaew in enumerate(lyaews):
lbl = '$%d$'%lyaew if ls=='-' else '_nolegend_'
ax1.plot(qsoGrid.gridCenters[1],fsel[:,i],lw=1.5,ls=ls,
color=cm.Blues(0.5+float(i)/len(lyaews)/2),
label=lbl)
handles, labels = ax1.get_legend_handles_labels()
legend(handles[::-1],labels[::-1],
handlelength=1,handletextpad=0.5,
borderpad=0,columnspacing=1,
loc=(0.02,0.01),
ncol=3,title=r'${\rm EW}_0({\rm Ly}\alpha)\ [\AA]$',
fontsize=10,frameon=False)#,bbox_to_anchor=(1,0.5))
ax1.set_title(r'$M_{1450}=%.1f,\ \ \alpha_\nu=%.1f$' %
(qsoGrid.gridCenters[0][M_i],uvslopes[3]),size=11)
# now by slope
ax2 = subplot(122)
for color_sel,ls in zip([weak_color_sel,strict_color_sel],['-','--']):
fsel = color_sel[M_i,:,2].sum(axis=-1) / float(nperbin)
for i,a_uv in enumerate(uvslopes):
lbl = r'$%.1f$'%a_uv if ls=='-' else '_nolegend_'
ax2.plot(qsoGrid.gridCenters[1],fsel[:,i],lw=1.5,ls=ls,
color=cm.plasma_r(0.5+float(i)/len(uvslopes)/2),
label=lbl)
legend(ncol=2,title=r'$\alpha_\nu$',
handlelength=1,handletextpad=0.5,
borderpad=0,columnspacing=1,
loc=(0.2,0.01),
fontsize=10,frameon=False)
ax2.set_title(r'$M_{1450}=%.1f,\ \ {\rm EW}_0({\rm Ly}\alpha)=%d$' %
(qsoGrid.gridCenters[0][M_i],lyaews[2]),size=11)
#
ax1.set_ylabel('$\mathrm{fraction\ selected}$',size=11)
for ax in [ax1,ax2]:
ax.set_xlabel('$z$',size=11)
ax.set_xlim(4.69,5.31)
ax.set_ylim(0,1.05)
for tick in ax.xaxis.get_major_ticks() + ax.yaxis.get_major_ticks():
tick.label1.set_fontsize(10)
ax.xaxis.set_major_locator(ticker.MultipleLocator(0.1))
ax.xaxis.set_minor_locator(ticker.MultipleLocator(0.05))
ax.yaxis.set_minor_locator(ticker.MultipleLocator(0.05))
ax.xaxis.set_major_formatter(ticker.FormatStrFormatter('$%.1f$'))
ax.yaxis.set_major_formatter(ticker.FormatStrFormatter('$%.1f$'))
savefig('lya_uvslope_sel.pdf')
```
## Selection dependence on Lya EW and dust extinction
This simulation utilizes a 4-d grid where the last two dimensions are Lya equivalent width and dust extinction.
```
M = AbsMagVar(UniformSampler(-29,-24),restWave=1450)
z = RedshiftVar(UniformSampler(4.7,5.3))
lya = GaussianLineEqWidthVar(FixedSampler([0.,15,60,120]),'LyaEW',1215.67,10.)
dust = SMCDustVar(UniformSampler(0,0.2))
qsoGrid = QsoSimGrid([M,z,lya,dust],(10,5,4,5),50,fixed_vars=['LyaEW','smcDustEBV'],
cosmo=WMAP9,units='luminosity')
```
In this case the full quasar model is turned on, including all emission lines and iron pseudo-continuum. This roughly follows what was used in Ross et al. 2013 for the "expdust" model.
```
# the "expdust" continuum model from Ross+13
contVar = BrokenPowerLawContinuumVar([GaussianSampler(-0.5,0.2),
GaussianSampler(-0.3,0.2)],
[1100.])
# generate lines using the Baldwin Effect emission line model from BOSS DR9
emLineVar = generateBEffEmissionLines(qsoGrid.absMag,ExcludeLines=['LyAn','LyAb'])
# the default iron template from Vestergaard & Wilkes 2001 was modified to fit BOSS spectra
fescales = [(0,1540,0.5),(1540,1680,2.0),(1680,1868,1.6),(1868,2140,1.0),(2140,3500,1.0)]
feVar = FeTemplateVar(VW01FeTemplateGrid(qsoGrid.z,wave,scales=fescales))
# Now add the features to the QSO grid
qsoGrid.addVars([contVar,emLineVar,feVar,forestVar])
qsoGrid.loadPhotoMap([('SDSS','Legacy'),('UKIRT','UKIDSS_LAS')])
# ready to generate spectra
_ = buildSpectraBySightLine(wave,qsoGrid,verbose=5)
g_mags = qsoGrid.asGrid('synMag')
g_z = qsoGrid.asGrid('z')
figure(figsize=(9,7))
for ew_i in range(4):
ax = subplot(2,2,ew_i+1)
scatter(g_mags[:,:,3-ew_i,0,::5,2]-g_mags[:,:,3-ew_i,0,::5,3],
g_mags[:,:,3-ew_i,0,::5,3]-g_mags[:,:,3-ew_i,0,::5,4],
c=g_z[:,:,3-ew_i,0,::5],s=7)
ax.set_title('EW = %d' % qsoGrid.gridCenters[2][3-ew_i])
ax.plot([1.2,1.2],[-0.5,0.13],c='m')
ax.plot([1.875,2.5],[0.55,0.55],c='m')
ri = linspace(1.2,1.875,2)
ax.plot(ri,polyval([0.625,0],ri-1),c='m')
ax.set_xlim(1.0,2.5)
ax.set_ylim(-0.5,1.3)
photoData = sqphoto.calcObsPhot(qsoGrid.synFlux,qsoGrid.photoMap)
qsoGrid.addData(photoData)
#g_mags = qsoGrid.asGrid('obsMag')
g_mags = qsoGrid.asGrid('synMag')
r_minus_i = g_mags[...,2] - g_mags[...,3]
i_minus_z = g_mags[...,3] - g_mags[...,4]
color_sel = ( (r_minus_i>1.2) &
(i_minus_z < 0.625*(r_minus_i-1)) &
(i_minus_z<0.55)
)
gs = gridspec.GridSpec(4,5)
gs.update(wspace=0.05,hspace=0.25)
figure(figsize=(14,8))
for i in range(4):
for j in range(5):
ax = plt.subplot(gs[i,j])
if j>0: ax.yaxis.set_ticklabels([])
if i<3: ax.xaxis.set_ticklabels([])
fsel = color_sel[:,:,-(i+1),j].sum(axis=-1) / 50.
imshow(fsel.T,vmin=0,vmax=1,extent=[4.7,5.3,-24,-29],
interpolation='nearest',aspect='auto',cmap=cm.hot_r)
title('EW=%d, E(B-V)=%.2f' %
(qsoGrid.gridCenters[2][-(i+1)],qsoGrid.gridCenters[3][j]),
size=12,weight='bold')
specFeatures = qsoGrid.getVars(SpectralFeatureVar)
ii = np.array([np.ravel_multi_index(t,qsoGrid.gridShape)
for t in [(0,0,3,0,0),(0,0,3,2,0),(0,0,3,4,0)]])
print ii
figure(figsize=(12,8))
for _i,i in enumerate(ii):
subplot(3,1,_i+1)
obj = qsoGrid.data[i]
sp = buildQsoSpectrum(wave,qsoGrid.cosmo,specFeatures,obj)
plot(wave,sp.f_lambda)
title('EW=%d, E(B-V)=%.2f' %
(obj['LyaEW'],obj['smcDustEBV']),size=12,weight='bold')
xlim(5000,1e4)
```
| github_jupyter |
```
# 定义读取文件的函数
def read_content(data_path):
"""
读取文件里的内容,并略去不能正确解码的行
"""
# 语料库使用GBK编码,对于不能编码的问题,选择略过
with open(data_path, "r", errors="ignore", encoding="GBK") as f:
raw_content = f.read()
content = ""
# 去掉其中的换行符
for i in raw_content.split("\n"):
content += i.strip()
return content
## 读取一个文件例子
print(read_content("./data/C3-Art/C3-Art0002.txt"))
# 使用pandas读取数据
import pandas as pd
from os import listdir
data_path = "./data"
category = ["C3-Art", "C11-Space", "C19-Computer", "C39-Sports"]
data = []
for i in category:
for j in listdir("%s/%s" % (data_path, i)):
content = read_content("%s/%s/%s" % (data_path, i, j))
data.append({"label": i, "content": content})
data = pd.DataFrame(data)
# 划分训练集和测试集
from sklearn.model_selection import train_test_split
train_data, test_data = train_test_split(data, test_size=0.3)
# 利用字典将文字转换为特征
from sklearn.feature_extraction.text import CountVectorizer
## 不进行中文分词,所以在字与字之间加入空格
train_data.loc[:, "content_simple_cut"] = train_data.apply(lambda x: " ".join(x["content"]), axis=1)
test_data.loc[:, "content_simple_cut"] = test_data.apply(lambda x: " ".join(x["content"]), axis=1)
## 利用CountVectorizer来构造字典,并进行特征提取
vect = CountVectorizer(token_pattern=r"(?u)\b\w+\b", binary=True)
X = vect.fit_transform(train_data["content_simple_cut"])
# 利用字典进行特征提取的展示
_test_string = ["我爱你你", "我不爱你"]
_test_string = [" ".join(i) for i in _test_string]
print(_test_string)
_binary_vect = CountVectorizer(token_pattern=r"(?u)\b\w+\b", binary=True)
print("binary vect:\n%s" % _binary_vect.fit_transform(_test_string).toarray())
_count_vect = CountVectorizer(token_pattern=r"(?u)\b\w+\b")
print("count vect:\n%s" % _count_vect.fit_transform(_test_string).toarray())
# 将文章的标签进行转换
from sklearn.preprocessing import LabelEncoder
le = LabelEncoder()
Y = le.fit_transform(train_data["label"])
# 使用伯努利模型对文本进行分类
from sklearn.naive_bayes import BernoulliNB
bernoulli_model = BernoulliNB()
bernoulli_model.fit(X, Y)
# 评估伯努利模型的分类效果
from sklearn.metrics import classification_report
print(classification_report(
le.transform(test_data["label"]),
bernoulli_model.predict(vect.transform(test_data["content_simple_cut"])),
target_names=le.classes_))
# 使用多项式模型对文本进行分类
from sklearn.naive_bayes import MultinomialNB
vect = CountVectorizer(token_pattern=r"(?u)\b\w+\b")
X = vect.fit_transform(train_data["content_simple_cut"])
multi_model = MultinomialNB()
multi_model.fit(X, Y)
print(classification_report(
le.transform(test_data["label"]),
multi_model.predict(vect.transform(test_data["content_simple_cut"])),
target_names=le.classes_))
# 使用TFIDF+多项式模型对数据建模
from sklearn.pipeline import Pipeline
from sklearn.feature_extraction.text import TfidfTransformer
## 使用pipeline来构造模型
pipe = Pipeline(
[("vect", CountVectorizer(token_pattern=r"(?u)\b\w+\b")),
("tfidf", TfidfTransformer(norm=None, sublinear_tf=True)),
("model", MultinomialNB())])
pipe.fit(train_data["content_simple_cut"], Y)
print(classification_report(
le.transform(test_data["label"]),
pipe.predict(test_data["content_simple_cut"]),
target_names=le.classes_))
# 使用第三方库jieba对文本进行分词,然后再进行分类
import jieba
print("/".join(jieba.cut("我爱北京天安门")))
print("/".join(jieba.cut("我爱北京天安门", cut_all=True)))
train_data.loc[:, "content_jieba_cut"] = train_data.apply(
lambda x: " ".join(jieba.cut(x["content"], cut_all=True)), axis=1)
test_data.loc[:, "content_jieba_cut"] = test_data.apply(
lambda x: " ".join(jieba.cut(x["content"], cut_all=True)), axis=1)
jieba_vect = CountVectorizer(token_pattern=r"(?u)\b\w+\b", binary=True)
X = jieba_vect.fit_transform(train_data["content_jieba_cut"])
bernoulli_model = BernoulliNB()
bernoulli_model.fit(X, Y)
print(classification_report(
le.transform(test_data["label"]),
bernoulli_model.predict(jieba_vect.transform(test_data["content_jieba_cut"])),
target_names=le.classes_))
```
| github_jupyter |
<table class="ee-notebook-buttons" align="left">
<td><a target="_blank" href="https://github.com/giswqs/earthengine-py-notebooks/tree/master/Datasets/Vectors/us_census_roads.ipynb"><img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" /> View source on GitHub</a></td>
<td><a target="_blank" href="https://nbviewer.jupyter.org/github/giswqs/earthengine-py-notebooks/blob/master/Datasets/Vectors/us_census_roads.ipynb"><img width=26px src="https://upload.wikimedia.org/wikipedia/commons/thumb/3/38/Jupyter_logo.svg/883px-Jupyter_logo.svg.png" />Notebook Viewer</a></td>
<td><a target="_blank" href="https://mybinder.org/v2/gh/giswqs/earthengine-py-notebooks/master?filepath=Datasets/Vectors/us_census_roads.ipynb"><img width=58px src="https://mybinder.org/static/images/logo_social.png" />Run in binder</a></td>
<td><a target="_blank" href="https://colab.research.google.com/github/giswqs/earthengine-py-notebooks/blob/master/Datasets/Vectors/us_census_roads.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" /> Run in Google Colab</a></td>
</table>
## Install Earth Engine API
Install the [Earth Engine Python API](https://developers.google.com/earth-engine/python_install) and [geehydro](https://github.com/giswqs/geehydro). The **geehydro** Python package builds on the [folium](https://github.com/python-visualization/folium) package and implements several methods for displaying Earth Engine data layers, such as `Map.addLayer()`, `Map.setCenter()`, `Map.centerObject()`, and `Map.setOptions()`.
The magic command `%%capture` can be used to hide output from a specific cell.
```
# %%capture
# !pip install earthengine-api
# !pip install geehydro
```
Import libraries
```
import ee
import folium
import geehydro
```
Authenticate and initialize Earth Engine API. You only need to authenticate the Earth Engine API once. Uncomment the line `ee.Authenticate()`
if you are running this notebook for this first time or if you are getting an authentication error.
```
# ee.Authenticate()
ee.Initialize()
```
## Create an interactive map
This step creates an interactive map using [folium](https://github.com/python-visualization/folium). The default basemap is the OpenStreetMap. Additional basemaps can be added using the `Map.setOptions()` function.
The optional basemaps can be `ROADMAP`, `SATELLITE`, `HYBRID`, `TERRAIN`, or `ESRI`.
```
Map = folium.Map(location=[40, -100], zoom_start=4)
Map.setOptions('HYBRID')
```
## Add Earth Engine Python script
```
fc = ee.FeatureCollection('TIGER/2016/Roads')
Map.setCenter(-73.9596, 40.7688, 12)
Map.addLayer(fc, {}, 'Census roads')
```
## Display Earth Engine data layers
```
Map.setControlVisibility(layerControl=True, fullscreenControl=True, latLngPopup=True)
Map
```
| github_jupyter |
```
import importlib.util
try:
import cirq
except ImportError:
print("installing cirq...")
!pip install --quiet cirq
print("installed cirq.")
try:
import quimb
except ImportError:
print("installing cirq-core[contrib]...")
!pip install --quiet cirq-core[contrib]
print("installed cirq-core[contrib].")
```
# Contract a Grid Circuit
Shallow circuits on a planar grid with low-weight observables permit easy contraction.
### Imports
```
import numpy as np
import networkx as nx
import cirq
import quimb
import quimb.tensor as qtn
from cirq.contrib.svg import SVGCircuit
import cirq.contrib.quimb as ccq
%matplotlib inline
from matplotlib import pyplot as plt
import seaborn as sns
sns.set_style('ticks')
plt.rc('axes', labelsize=16, titlesize=16)
plt.rc('xtick', labelsize=14)
plt.rc('ytick', labelsize=14)
plt.rc('legend', fontsize=14, title_fontsize=16)
# theme colors
QBLUE = '#1967d2'
QRED = '#ea4335ff'
QGOLD = '#fbbc05ff'
QGREEN = '#34a853ff'
QGOLD2 = '#ffca28'
QBLUE2 = '#1e88e5'
```
## Make an example circuit topology
We'll use entangling gates according to this topology and compute the value of an observable on the red nodes.
```
width = 3
height = 4
graph = nx.grid_2d_graph(width, height)
rs = np.random.RandomState(52)
nx.set_edge_attributes(graph, name='weight',
values={e: np.round(rs.uniform(), 2) for e in graph.edges})
zz_inds = ((width//2, (height//2-1)), (width//2, (height//2)))
nx.draw_networkx(graph,
pos={n:n for n in graph.nodes},
node_color=[QRED if node in zz_inds else QBLUE for node in graph.nodes])
```
### Circuit
```
qubits = [cirq.GridQubit(*n) for n in graph]
circuit = cirq.Circuit(
cirq.H.on_each(qubits),
ccq.get_grid_moments(graph),
cirq.Moment([cirq.rx(0.456).on_each(qubits)]),
)
SVGCircuit(circuit)
```
### Observable
```
ZZ = cirq.Z(cirq.GridQubit(*zz_inds[0])) * cirq.Z(cirq.GridQubit(*zz_inds[1]))
ZZ
```
### The contraction
The value of the observable is $\langle 0 | U^\dagger (ZZ) U |0 \rangle$.
```
tot_c = ccq.circuit_for_expectation_value(circuit, ZZ)
SVGCircuit(tot_c)
```
## We can simplify the circuit
By cancelling the "forwards" and "backwards" part of the circuit that are outside of the light-cone of the observable, we can reduce the number of gates to consider --- and sometimes the number of qubits involved at all. To see this in action, run the following cell and then keep re-running the following cell to watch gates disappear from the circuit.
```
compressed_c = tot_c.copy()
print(len(list(compressed_c.all_operations())), len(compressed_c.all_qubits()))
```
**(try re-running the following cell to watch the circuit get smaller)**
```
ccq.MergeNQubitGates(n_qubits=2).optimize_circuit(compressed_c)
ccq.MergeNQubitGates(n_qubits=1).optimize_circuit(compressed_c)
cirq.DropNegligible(tolerance=1e-6).optimize_circuit(compressed_c)
cirq.DropEmptyMoments().optimize_circuit(compressed_c)
print(len(list(compressed_c.all_operations())), len(compressed_c.all_qubits()))
SVGCircuit(compressed_c)
```
### Utility function to fully-simplify
We provide this utility function to fully simplify a circuit.
```
ccq.simplify_expectation_value_circuit(tot_c)
SVGCircuit(tot_c)
# simplification might eliminate qubits entirely for large graphs and
# shallow `p`, so re-get the current qubits.
qubits = sorted(tot_c.all_qubits())
print(len(qubits))
```
## Turn it into a Tensor Netowork
We explicitly "cap" the tensor network with `<0..0|` bras so the entire thing contracts to the expectation value $\langle 0 | U^\dagger (ZZ) U |0 \rangle$.
```
tensors, qubit_frontier, fix = ccq.circuit_to_tensors(
circuit=tot_c, qubits=qubits)
end_bras = [
qtn.Tensor(
data=quimb.up().squeeze(),
inds=(f'i{qubit_frontier[q]}_q{q}',),
tags={'Q0', 'bra0'}) for q in qubits
]
tn = qtn.TensorNetwork(tensors + end_bras)
tn.graph(color=['Q0', 'Q1', 'Q2'])
plt.show()
```
### `rank_simplify` effectively folds together 1- and 2-qubit gates
In practice, using this is faster than running the circuit optimizer to remove gates that cancel themselves, but please benchmark for your particular use case.
```
tn.rank_simplify(inplace=True)
tn.graph(color=['Q0', 'Q1', 'Q2'])
```
### The tensor contraction path tells us how expensive this will be
```
path_info = tn.contract(get='path-info')
path_info.opt_cost / int(3e9) # assuming 3gflop, in seconds
path_info.largest_intermediate * 128 / 8 / 1024 / 1024 / 1024 # gb
```
### Do the contraction
```
zz = tn.contract(inplace=True)
zz = np.real_if_close(zz)
print(zz)
```
## Big Circuit
```
width = 8
height = 8
graph = nx.grid_2d_graph(width, height)
rs = np.random.RandomState(52)
nx.set_edge_attributes(graph, name='weight',
values={e: np.round(rs.uniform(), 2) for e in graph.edges})
zz_inds = ((width//2, (height//2-1)), (width//2, (height//2)))
nx.draw_networkx(graph,
pos={n:n for n in graph.nodes},
node_color=[QRED if node in zz_inds else QBLUE for node in graph.nodes])
qubits = [cirq.GridQubit(*n) for n in graph]
circuit = cirq.Circuit(
cirq.H.on_each(qubits),
ccq.get_grid_moments(graph),
cirq.Moment([cirq.rx(0.456).on_each(qubits)]),
)
ZZ = cirq.Z(cirq.GridQubit(*zz_inds[0])) * cirq.Z(cirq.GridQubit(*zz_inds[1]))
ZZ
ccq.tensor_expectation_value(circuit, ZZ)
```
| github_jupyter |
```
import findspark
findspark.init('/home/ubuntu/spark-2.1.1-bin-hadoop2.7')
import pyspark
from pyspark.sql import SparkSession
spark = SparkSession.builder.appName('Project').getOrCreate()
import pandas as pd
import itertools
columns= []
table_rows =[]
Decision =''
decision_rows=[]
table_header =[]
table_content =[]
#ask the user to enter the file name and then the load the file data
def readFile():
is_Columns_found = False
column_count_loc=0
print('Enter FileName ')
FileName = input("> ")
print("Loading file {}".format(FileName))
file = open(FileName, 'r')
for row_Number, row in enumerate(file):
row_Item = row.split()
if(len(row_Item)==0):
continue
if(row_Item[0].startswith('<') or row_Item[0].startswith('!')):
continue
elif(is_Columns_found==False and row_Item[0].startswith('[')):
columns = row_Item[1:-2]
Decision=row_Item[-2]
table_header=row_Item[1:-1]
is_Columns_found=True
print(is_Columns_found)
else:
table_rows.append(row_Item[0:-1])
decision_rows.append(row_Item[-1])
row_Item.append(row_Number+1)
table_content.append(row_Item)
#table_content.append(row_Number+1)
column_count_loc+=1
print('Printing Attributes : ',columns)
print('Printing Decision : ',Decision)
return column_count_loc,columns,table_rows,Decision,decision_rows,table_header,table_content
#Calling Read file function
column_count,columns,table_rows,Decision,decision_rows,table_header,table_content=readFile()
from pyspark.sql.functions import monotonically_increasing_id
def convertTableToSparkDataframe():
df = spark.createDataFrame(table_content , table_header)
#res = df.withColumn("id", monotonically_increasing_id()+1)
print('Do you want to display the table (Y/N) ')
choice = input("> ")
if(choice=='Y' or choice == 'y'):
res.show()
return res
table_header.append("id")
res=convertTableToSparkDataframe()
table_content
table_header
res.show()
def is_numerical(num):
try:
float(num)
return True
except ValueError:
return False
def creatingBlocks(blocks,data_type,data_type_num,table_header,res):
for col in table_header:
new_df1=res.select(col).distinct()
b={}
count=0
if col in data_type_num.keys():
count=data_type_num[col]
for r in new_df1.head(new_df1.count()):
#print(r[0])
list1 = res.filter(res[col]==r[0]).select('id').collect()
list1_array =[int(i.id) for i in list1]
list1_array=list(set(list1_array))
b[r[0]]= list1_array
if(is_numerical(r[0])):
count+=len(list1_array)
data_type_num[col]=count
if col in blocks:
blocks[col].append(b)
else:
blocks[col]=b
print('Do you want to display the Blocks (Y/N) ')
choice = input("> ")
if(choice=='Y' or choice == 'y'):
print(blocks)
return blocks,data_type_num,data_type
data_type_num ={}
data_type = {}
blocks ={}
blocks,data_type_num,data_type=creatingBlocks(blocks,data_type,data_type_num,table_header,res)
def openFileToWrite(f_name):
temp_fileHandler= open('Log.txt','w')
temp_fileHandler.write('Starting file to write \n')
return temp_fileHandler
def writeBlockInFile(file,columns,blocks):
file.write('\n Dictionary\n')
for attr in columns:
file.write('Column/Attribute : ')
file.write(attr)
file.write('\n')
for val in blocks[attr]:
file.write(val)
file.write(': [ ')
tem=blocks[attr][val]
for v in tem:
file.write(str(v))
file.write(' ')
file.write(']\n')
def close_file(file):
file.write('\n ------ Closing File ------\n')
file.close()
#writing the block values in a file
file1=openFileToWrite('Block_Output.txt')
writeBlockInFile(file1,columns,blocks)
#
def processingDataForMissingValues():
list_missing=[]
for attr in columns:
list_missing=[]
#missing value *
if "*" in blocks[attr]:
list_missing = blocks[attr].pop('*',None)
if(len(list_missing)>0):
for val in blocks[attr]:
blocks[attr][val] = list(set(blocks[attr][val]).union(set(list_missing)))
#missing value -
if "-" in blocks[attr]:
list_missing = blocks[attr].pop('-',None)
if(len(list_missing)>0):
for num in list_missing:
#print(decision_rows[num])
for val in blocks[attr]:
if(len(list(set(set(blocks[Decision][decision_rows[num]]).intersection(set(set(blocks[attr][val]))))))>0):
Temp_arr=[]
Temp_arr.append(num)
blocks[attr][val] = list(set(blocks[attr][val]).union(set(Temp_arr)))
#missing value ?
if "?" in blocks[attr]:
blocks[attr].pop('?',None)
#print(blocks)
return blocks
blocks=processingDataForMissingValues()
#blocks
# writing the updated values in file
writeBlockInFile(file1,columns,blocks)
# remove the restriction
def ClassifyTableData():
column_count_limit = column_count*0.75
for col in table_header:
if(data_type_num[col]>column_count_limit):
data_type[col]="Numerical"
else:
data_type[col]="Discrete"
return data_type
data_type=ClassifyTableData()
print(data_type)
def add_Cases(formatedCut,colName,values):
if formatedCut in blocks[colName]:
blocks[colName][formatedCut]=list(set(blocks[colName][formatedCut]).union(set(values)))
else:
blocks[colName][formatedCut]=values
def convert_NumericalToConceptual():
for colName in columns:
if(data_type[colName]=='Numerical'):
tempArray = sorted(list(blocks[colName].keys()))
list_cutpoints = []
for cutpoint_i in range(len(tempArray)-1):
list_cutpoints.append((float(tempArray[cutpoint_i])+float(tempArray[cutpoint_i+1]))/2)
for val in tempArray:
for cut_point in list_cutpoints:
values = blocks[colName][val]
if(float(val)>cut_point):
formatedCut = " > {0:.2f} ".format(cut_point)
elif (float(val)<=cut_point):
formatedCut = " <= {0:.2f} ".format(cut_point)
add_Cases(formatedCut,colName,values)
for val in tempArray:
blocks[colName].pop(val,None)
convert_NumericalToConceptual()
#blocks
file1.write('\n Updated Dictionary converting Numerical Attributes to Conceptual\n')
writeBlockInFile(file1,columns,blocks)
from RuleClass import RuleClass
def print_rule(curRule,sr_no):
print('Rule ',sr_no)
#print(curRule.attributes, curRule.attributes_value)
#k=zip(curRule.attributes, curRule.attributes_value)
#print(list(k))
rule_str=''
is_first=True
for i in range(len(curRule.attributes)):
if(is_first==False):
rule_str = rule_str +' & '
rule_str=rule_str +'('+curRule.attributes[i]+', '+curRule.attributes_value[i]+")"
is_first=False
rule_str = rule_str +' --> ('+Decision+', '+curRule.Decision+")"
print(rule_str,',')
#print('Decision ',curRule.Decision)
#print('Decision Covered ',curRule.decision_cover)
#print('InCorrect Cover ',curRule.false_cover)
print('Strength: ',curRule.strength,',')
print('Specificity: ',curRule.specificity,',')
print('Support: ',curRule.support,',')
print('Conditional Probablity: ',curRule.conditionalProbablity,'.')
print('----------------------------------------------------------------------------------')
def print_rule_without_detials(curRule,sr_no):
print('Rule ',sr_no)
#print(curRule.attributes, curRule.attributes_value)
#k=zip(curRule.attributes, curRule.attributes_value)
#print(list(k))
rule_str=''
is_first=True
for i in range(len(curRule.attributes)):
if(is_first==False):
rule_str = rule_str +' & '
rule_str=rule_str +'('+curRule.attributes[i]+', '+curRule.attributes_value[i]+")"
is_first=False
rule_str = rule_str +' --> ('+Decision+', '+curRule.Decision+")"
print(rule_str)
#print('Decision ',curRule.Decision)
print('--------------------------------------------------------------------------------------------------')
min_rule_Length=1
max_rule_Length=1
min_Strength=5
min_prob=1
print('Default rule specification, (Please enter c or C, if you want to change any of the below condition)')
print('Minimum length of rule ',min_rule_Length)
print('Maximum length of rule ',max_rule_Length)
print('Minimum Strength of rule ',min_Strength)
print('Minimum Conditional Probablity ',min_prob)
choice = input("> ")
if(choice=='c' or choice == 'C'):
print('Enter minimum length of rule ',min_rule_Length)
min_rule_Length= input("> ")
print('Enter maximum length of rule ',max_rule_Length)
max_rule_Length= input("> ")
print('Enter minimum Strength of rule ',min_Strength)
min_Strength= input("> ")
print('Enter minimum Conditional Probablity ',min_prob)
min_rule_Length= input("> ")
possibleRules = []
rules = []
all_ID = res.select('id').collect()
all_ID =[int(i.id) for i in all_ID]
def compute_rules(min_rule_Length,max_rule_Length,min_Strength,min_prob):
temp_Decision=''
DecisionCovered = all_ID
#possible Rules
for j in range(min_rule_Length,max_rule_Length+1):
for idx, attributeSet in enumerate(itertools.combinations(columns, j)):
#print(idx)
print(attributeSet)
valueSet = [[val for val in blocks[attr].keys() if val not in ["*", "?", "-"]] for attr in attributeSet]
#print('Value Set ',valueSet)
for combination in itertools.product(*valueSet):
k=zip(attributeSet, combination)
#print(attributeSet,len(combination))
#print('Printing zip',list(k))
temp=res
temp_ID = all_ID
for ri in range(len(attributeSet)):
templist123= blocks[attributeSet[ri]][combination[ri]]
temp_ID=list(set(temp_ID).intersection(set(templist123)))
#temp = temp.filter(temp[attributeSet[ri]]==combination[ri])
#list_arr= temp.select('id').collect()
strength=len(temp_ID)
if(strength<min_Strength):
continue
#print(temp_ID)
max_covered =0
for key in blocks[Decision]:
DecisionCovered=list(set(temp_ID).intersection(set(blocks[Decision][key])))
if(max_covered<len(DecisionCovered)):
max_covered=len(DecisionCovered)
temp_Decision=key
DecisionCovered=list(set(temp_ID).intersection(set(blocks[Decision][temp_Decision])))
cond_prob=float(len(DecisionCovered))/float(strength)
if(cond_prob<min_prob ):
continue
inCorrectCover=list(set(temp_ID)-set(DecisionCovered))
#list1_array_temp =[int(i.id) for i in list_arr]
#print('Covered by Rule',list1_array_temp, temp_ID)
temp_rule=RuleClass(list(attributeSet),list(combination),temp_ID,temp_Decision,DecisionCovered,inCorrectCover)
possibleRules.append(temp_rule)
return possibleRules
possibleRules=compute_rules(min_rule_Length,max_rule_Length,min_Strength,min_prob)
def Display_Rules_SortStrength(possibleRules):
possibleRules = sorted(possibleRules, key=lambda each_rule: each_rule.strength,reverse=True)
#for each_rule in possibleRules:
#print_rule(each_rule)
return possibleRules
def Display_Rules_SortConditionalProbablity(possibleRules):
possibleRules = sorted(possibleRules, key=lambda each_rule: each_rule.strength,reverse=True)
possibleRules = sorted(possibleRules, key=lambda each_rule: each_rule.conditionalProbablity,reverse=True)
#for each_rule in possibleRules:
#if(float(min_prob)>float(each_rule.conditionalProbablity)):
#break
#print_rule(each_rule)
return possibleRules
print("Number of rules generated",len(possibleRules))
print('Enter \n 1. Sort rules by Conditional Probbality \n 2. Sort rules by Rule Strength \n By default 2 is selected')
choice = input("> ")
if(choice==1):
possibleRules1=Display_Rules_SortStrength(possibleRules)
else:
possibleRules1=Display_Rules_SortConditionalProbablity(possibleRules)
sr_no=1
for each_rule in possibleRules1:
print_rule_without_detials(each_rule,sr_no)
sr_no+=1
sr_no=0
for each_rule in possibleRules1:
sr_no=sr_no+1
print_rule(each_rule,sr_no)
out_fileHandler= open('output1.txt','w')
sr_no=0
for curRule in possibleRules1:
sr_no=sr_no+1
#out_fileHandler.write(str (sr_no))
#out_fileHandler.write('. ')
rule_str=''
is_first=True
for i in range(len(curRule.attributes)):
if(is_first==False):
rule_str = rule_str +' & '
rule_str=rule_str +'('+curRule.attributes[i]+', '+curRule.attributes_value[i]+")"
is_first=False
rule_str = rule_str +' --> ('+Decision+', '+curRule.Decision+")"
out_fileHandler.write(rule_str)
out_fileHandler.write('\n')
#print('Decision ',curRule.Decision)
#out_fileHandler.write('--------------------------------------------------------------------------------------------------')
#out_fileHandler.write('\n')
if(sr_no==1000):
break
out_fileHandler.write('\n ------ Closing File ------\n')
out_fileHandler.close()
sr_no
```
| github_jupyter |
# Introduction #
In this first exercise, you'll do a complete iteration of feature development: understand the dataset, create a baseline model, create a derived feature, and compare performance.
Run this next cell to set everything up!
```
# Setup feedback system
from learntools.core import binder
binder.bind(globals())
from learntools.feature_engineering_new.ex1 import *
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
from sklearn.model_selection import cross_val_score
from xgboost import XGBRegressor
# Set Matplotlib defaults
plt.style.use("seaborn-whitegrid")
plt.rc("figure", autolayout=True)
plt.rc(
"axes",
labelweight="bold",
labelsize="large",
titleweight="bold",
titlesize=14,
titlepad=10,
)
def score_dataset(X, y, model=XGBRegressor()):
# Label encoding for categoricals
for colname in X.select_dtypes(["category", "object"]):
X[colname], _ = X[colname].factorize()
# Metric for Housing competition is RMSLE (Root Mean Squared Log Error)
log_y = np.log(y)
score = cross_val_score(
model, X, log_y, cv=5, scoring="neg_mean_squared_error",
)
score = -1 * score.mean()
score = np.sqrt(score)
return score
```
You'll work with the **Ames Housing** dataset to predict house prices based on information like the number oof bedrooms, number of bathrooms, and the year it was built. You can find all of the datasets from this course [here](https://www.kaggle.com/ryanholbrook/fe-course-data).
Run the next code cell to load and preview the data.
```
df = pd.read_csv("../input/fe-course-data/ames.csv")
df.head()
```
Effective feature engineering makes use of prominent relationships in the dataset. Data visualization is one of the best ways to discover these relationships. Now you'll use Seaborn to discover some important things about the *Ames* data. (Check our our [Data Visualization](https://www.kaggle.com/learn/data-visualization) course, too!)
You can see the relationship a feature has to the target with a *scatterplot*. Take a look at scatterplots for `YearBuilt` and `MoSold` relative to `SalePrice`:
```
fig, axs = plt.subplots(1, 2, figsize=(10, 5), sharey=True)
axs[0] = sns.scatterplot(x="YearBuilt", y="SalePrice", data=df, ax=axs[0])
axs[1] = sns.scatterplot(x="MoSold", y="SalePrice", data=df, ax=axs[1])
```
# 1) Discover Relationships
Does there appear to be a significant relationship between either feature and the target? If so, does the relationship appear to be linear (best fit with a line)?
After you've thought about your answer, run the following cell for a solution.
```
# View the solution (Run this cell to receive credit!)
q_1.check()
```
-------------------------------------------------------------------------------
The number of bathrooms in a home is often important to prospective home-buyers. The data contains four such features:
- `FullBath`
- `HalfBath`
- `BsmtFullBath`
- `BsmtHalfBath`
# 2) Create a New Feature
Create a new feature `TotalBaths` that describes the *total* number of bathrooms in a home. There are a couple answers that might be reasonable. Can you think of a way that's better than just summing the features up?
```
X = df.copy()
y = X.pop('SalePrice')
# YOUR CODE HERE
X["TotalBaths"] = ____
# Check your answer
q_2.check()
# Lines below will give you a hint or solution code
#_COMMENT_IF(PROD)_
q_2.hint()
#_COMMENT_IF(PROD)_
q_2.solution()
#%%RM_IF(PROD)%%
X = df.copy()
y = X.pop('SalePrice')
X["TotalBaths"] = \
df.FullBath + df.HalfBath + \
df.BsmtFullBath + df.BsmtHalfBath
q_2.assert_check_failed()
#%%RM_IF(PROD)%%
X = df.copy()
y = X.pop('SalePrice')
X["TotalBaths"] = \
df.FullBath + df.HalfBath
q_2.assert_check_failed()
#%%RM_IF(PROD)%%
X = df.copy()
y = X.pop('SalePrice')
X["TotalBaths"] = \
df.BsmtFullBath + df.BsmtHalfBath
q_2.assert_check_failed()
#%%RM_IF(PROD)%%
X = df.copy()
y = X.pop('SalePrice')
# Solution 1: HalfBath with half the weight of FullBath
X["TotalBaths"] = \
df.FullBath + 0.5 * df.HalfBath + \
df.BsmtFullBath + 0.5 * df.BsmtHalfBath
# Solution 2: Same, but preserves int type
X["TotalBaths"] = \
2 * df.FullBath + df.HalfBath + \
2 * df.BsmtFullBath + df.BsmtHalfBath
q_2.assert_check_passed()
```
-------------------------------------------------------------------------------
Now compare the performance of XGBoost with and without the `TotalBaths` feature. (The `score_dataset` function performs 5-fold cross-validation with XGBoost using with the RMSLE metric, the same metric used in the *House Prices* competition.)
```
X_base = df.copy()
y_base = X_base.pop("SalePrice")
baseline_score = score_dataset(X_base, y_base)
new_score = score_dataset(X, y)
print(f"Score Without New Feature: {baseline_score:.4f} RMSLE")
print(f"Score With New Feature: {new_score:.4f} RMSLE")
```
# 3) Feature Selection
Based on the performance of XGBoost with and without the additional feature, would you decide to keep or discard the new feature? Or is there not enough information to decide?
After you've thought about you're answer, run the next cell for the solution.
```
# View the solution (Run this cell to receive credit!)
q_3.check()
```
# Iterating on Feature Sets #
You've just worked through a complete iteration of feature development: discovery, creation, validation, and selection. In most machine learning projects, you'll likely go through many such iterations before arriving at your final, best feature set.
In the next lesson, you'll learn about *feature utility*, a way of scoring features for their potential usefulness -- a big help when you're just getting started with a new dataset!
# Keep Going #
[**Discover useful features**](#$NEXT_NOTEBOOK_URL$) with mutual information.
| github_jupyter |
```
import pandas as pd
import json
import gzip
import os
import re as regex
import pickle
import progressbar
#pip install progressbar2
path = '../../../src/data/schemafiltereddata/TrainTestTables/'
#small_files_train = os.listdir(path+'Small/Train/')
small_files_train = [file for file in os.listdir(path+'Small/Train/') if file.endswith('.json.gz')]
#medium_files_train = [file for file in os.listdir(path+'Medium/Train/') if file.endswith('.json.gz')]
#large_files_train = [file for file in os.listdir(path+'Large/Train/') if file.endswith('.json.gz')]
#test_files = [file for file in os.listdir(path+'Test/') if file.endswith('.json.gz')]
#turl_input_path = os.path.join(product_path, 'TURL/input')
small_files_train
#medium_files_train
print(len(small_files_train))
#table_id = 'Product_kingmemorialpark.com_September2020.json.gz'
df = pd.read_json(os.path.join(path + 'Small/Train/Product_kingmemorialpark.com_September2020.json.gz'), compression='gzip', lines=True)
df_1 = pd.read_json(os.path.join(path + 'Small/Train/MusicRecording_bencrosbymusic.com_September2020.json.gz'), compression='gzip', lines=True)
df_1
file = 'MusicRecording_bencrosbymusic.com_September2020.json.gz'
category , rest = file.split('_', 1)
print(category)
page, other = file.split('_')
page
df #test for one table
j = 0
table_representation = [] #empty list for table
train_representation = []
table_representation.append(file)#append filename table id
#page, other = file.split('_', 2)
table_representation.append('kingmemorialpark.com')#append page title -> not relevant
table_representation.append('')#append wikipedia page id -> not given
category , rest = file.split('_', 1) #get category for file
table_representation.append(category)#append information about entity/section title
table_representation.append('')#append table caption -> not given
table_representation.append(list(df_1.columns)) #append headers
cells_representation = []#representation of all rows
#df_1_1 = df_1_1.drop(columns='row_id')
for col in df_1.columns: # cell representation of single column
cell_representation=[]
for i in range(len(df_1)):
cell_representation.append([[int(i),int(j)],[df_1.index[i],str(df_1[col][i])]])
j += 1
cells_representation.append(cell_representation) #append single cell representation to representation of all rows
table_representation.append(cells_representation) #append it to representation of whole table
col_types = [] # create empty list for col types
#category , file = file.split('_', 1)
for element in list(df_1.columns): #iterate over headers in tables
col_types = [[category + '.' + element] for element in list(df_1.columns)] #append category to header for col type annotation
table_representation.append(col_types) #append col type annotation
#train_representation.append(table_representation)
train_representation.append(table_representation)
train_representation
#train_representation
#train_file = open("train_representation.txt", "w")
#for element in train_representation:
# train_file.write(element + "\n")
#train_file.close()
turl_path = path = '../../../src/data/schemafiltereddata/TURL/input_old'
# write small train representation to txt
#with open(os.path.join(turl_path, 'small_train_representation.txt'), "wb") as fp: #Pickling - write list to txt file
# pickle.dump(train_representation, fp)
# write medium train representation to txt
with open(os.path.join(turl_path, 'test_input.txt'), "wb") as fp: #Pickling - write list to txt file
pickle.dump(train_representation, fp)
# write test representation to txt
#with open(os.path.join(turl_path, 'test_representation.txt'), "wb") as fp: #Pickling - write list to txt file
# pickle.dump(test_representation, fp)
with open(os.path.join(turl_path, 'medium_train_representation.txt'), "rb") as fp: # Unpickling
train_representation = pickle.load(fp)
path = '/work-ceph/bizer-tp2021/data_integration_using_deep_learning/src/data/data/CSV_files'
tables = pd.read_csv(path + '/all_files_cleaned.csv')
tables = tables.drop(columns={'Unnamed: 0'})
target_columns = tables.groupby('filename')['column_name'].apply(list).reset_index(name='listofcolumns')
target_columns.filename = target_columns.filename.str.split('_', expand = True)
target_columns.index = target_columns['filename']
target_columns
target_columns = target_columns.explode('listofcolumns')
target_columns
#create list of target columns types for type_vocab.txt file
target_type = [] # create emplyt list for target col types
for row in target_columns.itertuples(): # iterate over each rows in target_columns df
target_type.append(row.filename + '.' + row.listofcolumns)
target_type
len(target_type)
# write tagret col types to txt
with open(os.path.join(turl_path, 'type_vocab.txt'), "wb") as fp: #Pickling - write list to txt file
pickle.dump(target_type, fp)
with open(os.path.join(turl_path, 'type_vocab.txt'), "rb") as fp: # Unpickling
target_type = pickle.load(fp)
```
| github_jupyter |
```
# dependencies
import json
import os
from itertools import filterfalse
# zoltpy related dependencies
import zoltpy.util as zutil
from zoltpy.connection import ZoltarConnection
from zoltpy.quantile_io import json_io_dict_from_quantile_csv_file
from zoltpy.cdc_io import YYYY_MM_DD_DATE_FORMAT
from zoltpy.covid19 import COVID_TARGETS, covid19_row_validator, validate_quantile_csv_file, COVID_ADDL_REQ_COLS
# check current working directory
print(os.getcwd())
# hash db file path
HASH_DB_PATH = '../zoltar_scripts/validated_file_db.json'
# load hash database
with open(HASH_DB_PATH, 'r') as f:
hash_db = json.load(f)
print(len(hash_db))
# metadata fields to Zoltar params
MD_FIELDS_ZOLTAR_PARAMS = {
'team_name': 'team_name',
'model_name': 'name',
'model_abbr': 'abbreviation',
'model_contributors': 'contributors',
'website_url': 'home_url',
'license': 'license',
'team_model_designation': 'notes',
'methods': 'description',
'repo_url': 'aux_data_url',
'citation': 'citation',
'methods_long': 'methods'
}
# make sure that we have the required environment variables
is_have_zoltar_credentials = False
Z_USERNAME, Z_PASSWORD = map(os.environ.get, ['Z_USERNAME', 'Z_PASSWORD'])
if Z_USERNAME and Z_PASSWORD:
is_have_zoltar_credentials = True
print(f'Do we have Zoltar credentials? {"yes" if is_have_zoltar_credentials else "no"}')
# get connection to Zoltar
conn = ZoltarConnection()
try:
conn.authenticate('covid19hub', 'nL82*&dKMvX%')
except RuntimeError:
print(f'Connection to Zoltar not established; please try again')
print(f'Connection established with Zoltar')
# get all existing timezeros and models in the project
project_obj = [project for project in conn.projects if project.name == 'COVID-19 Forecasts'][0]
model_to_csvs_dict = { \
model.abbreviation: {f.source: f for f in model.forecasts} for model in project_obj.models \
}
# forecast directories path
FORECAST_DIRS_PATH = '../../data-processed/'
# get forecast directories
list_model_directories = filterfalse(
lambda m: not os.path.isdir(f'{FORECAST_DIRS_PATH}{m}'),
os.listdir(FORECAST_DIRS_PATH)
)
# get forecasts
for forecast_dir in list_model_directories:
print(f'current model: {forecast_dir}')
# m.split('.')[1] gets the extension
list_forecast_csvs = filterfalse(
lambda m: m.split('.')[-1] != 'csv',
os.listdir(f'{FORECAST_DIRS_PATH}{forecast_dir}')
)
# iterate through all csvs in forecast directory
# turn on the print statements for more info
for forecast_csv in list_forecast_csvs:
# print(f'checking if forecast {forecast_csv} was recorded...')
is_forecast_hash_not_in_table = forecast_csv not in hash_db
# print( \
# f'{forecast_csv} was not recorded.' \
# if is_forecast_hash_not_in_table \
# else f'{forecast_csv} was recorded.' \
# )
# print(f'checking if forecast {forecast_csv} is on Zoltar...')
is_forecast_on_zoltar = forecast_csv in model_to_csvs_dict[forecast_dir]
# print( \
# f'{forecast_csv} is on Zoltar.' \
# if is_forecast_on_zoltar \
# else f'{forecast_csv} is not on Zoltar; skipping.' \
# )
if is_forecast_hash_not_in_table and is_forecast_on_zoltar:
print(f'\tdeleting {forecast_csv} because its hash is not recorded but it is uploaded to Zoltar...')
job = model_to_csvs_dict[forecast_dir][forecast_csv].delete()
print(f'\tdelete job for {forecast_csv} enqueued')
pass
```
| github_jupyter |
### Necessary Imports
```
import numpy as np
import tensorflow as tf
from tensorflow.keras.utils import to_categorical, plot_model
from tensorflow.keras.models import Model
from tensorflow.keras.datasets import cifar10
from tensorflow.keras import Input
from tensorflow.keras.layers import Conv2D, BatchNormalization, Activation
from tensorflow.keras.regularizers import l2
from tensorflow.keras.layers import Flatten, add
from tensorflow.keras.layers import AveragePooling2D, Dense
from tensorflow.keras.optimizers import Adam
from tensorflow.keras.callbacks import ModelCheckpoint, LearningRateScheduler
from tensorflow.keras.callbacks import ReduceLROnPlateau
from tensorflow.keras.preprocessing.image import ImageDataGenerator
import os
import math
```
### Model Parameters
```
batch_size = 32
epochs = 10
data_augmentation = True
num_classes = 10
n = 3
depth = n * 6 + 2
subtract_pixel_mean = True
```
### Loadind Data
```
(X_train, y_train), (X_test, y_test) = cifar10.load_data()
X_train = X_train.reshape(50000, 32, 32, 3)
X_test = X_test.reshape(10000, 32, 32, 3)
X_train = X_train.astype('float32')
X_test = X_test.astype('float32')
if subtract_pixel_mean:
X_train_mean = np.mean(X_train, axis=0)
X_train -= X_train_mean
X_test -= X_train_mean
num_classes = len(np.unique(y_train))
y_train = to_categorical(y_train)
y_test = to_categorical(y_test)
input_shape = X_train.shape[1:]
X_train.shape
#y_train.shape
```
### Model Architecture
```
def resnet_layer(inputs,
kernel_size=3,
num_filters=16,
strides=1,
activation='relu',
batch_normalization=True,
conv_first=True):
"""2D Convolution-BatchNormalization-Activation Residual Block"""
conv = Conv2D(num_filters,
kernel_size=kernel_size,
strides=strides,
padding='same',
kernel_initializer='he_normal',
kernel_regularizer=l2())
x = inputs
if conv_first:
x = conv(x)
if batch_normalization:
x = BatchNormalization()(x)
if activation is not None:
x = Activation(activation)(x)
else:
if batch_normalization:
x = BatchNormalization()(x)
if activation is not None:
x = Activation(activation)(x)
x = conv(x)
return x
def resnet_v1(input_shape, depth, num_classes):
if (depth-2)%6 !=0:
raise ValueError('Depth shoutld be 6n+2: eg-20, 32, 44 etc')
# Model Definition
num_filters = 16
num_res_blocks = int((depth-2)/6)
inputs = Input(shape=input_shape)
x = resnet_layer(inputs=inputs)
# Instance the stack of residual units
for stack in range(3):
for res_block in range(num_res_blocks):
strides = 1
if stack>0 and res_block==0:
strides=2 # Downsampling
y = resnet_layer(inputs=x,
num_filters=num_filters,
strides=strides)
y = resnet_layer(inputs=y,
num_filters=num_filters,
activation=None)
if stack>0 and res_block==0:
# linear projection residual shortcut
# connection to match changed dims
x = resnet_layer(inputs=x,
num_filters=num_filters,
strides=strides,
activation=None,
batch_normalization=False)
x = add([x, y])
x = Activation('relu')(x)
num_filters *= 2
x = AveragePooling2D(pool_size=8)(x)
y = Flatten()(x)
outputs = Dense(num_classes,
activation='softmax',
kernel_initializer='he_normal')(y)
model = Model(inputs=inputs, outputs=outputs)
return model
model = resnet_v1(input_shape=input_shape, depth = depth, num_classes = num_classes)
model.summary()
```
### Model Compile
```
model.compile(loss='categorical_crossentropy',
optimizer=Adam(lr=0.0001),
metrics=['acc'])
#plot_model(model, to_file="resnet_v1.png", show_shapes=True)
```
### Model saving directory
```
save_dir = os.path.join(os.getcwd(), 'saved_moddels')
model_name = 'cifar_10_resnet_v1_model.{epoch:03d}.h5'
if not os.path.isdir(save_dir):
os.makedirs(save_dir)
filepath = os.path.join(save_dir, model_name)
```
### Training
```
checkpoint = ModelCheckpoint(filepath=filepath,
monitor='val_accuracy',
verbose=1,
save_best_only=True)
lr_reducer = ReduceLROnPlateau(factor=np.sqrt(0.1),
cooldown=0,
patience=5,
min_lr=0.5e-6)
callbacks = [checkpoint, lr_reducer]
if not data_augmentation:
print('Not using data augmentation.')
model.fit(X_train, y_train,
batch_size=batch_size,
epochs=epochs,
validation_data=(X_test, y_test),
shuffle=True,
callbacks=callbacks)
else:
print('Using real-time data augmentation.')
# this will do preprocessing and realtime data augmentation:
datagen = ImageDataGenerator(
# set input mean to 0 over the dataset
featurewise_center=False,
# set each sample mean to 0
samplewise_center=False,
# divide inputs by std of dataset
featurewise_std_normalization=False,
# divide each input by its std
samplewise_std_normalization=False,
# apply ZCA whitening
zca_whitening=False,
# randomly rotate images in the range (deg 0 to 180)
rotation_range=0,
# randomly shift images horizontally
width_shift_range=0.1,
# randomly shift images vertically
height_shift_range=0.1,
# randomly flip images
horizontal_flip=True,
# randomly flip images
vertical_flip=False)
# compute quantities required for featurewise normalization
# (std, mean, and principal components if ZCA whitening is applied).
datagen.fit(X_train)
steps_per_epoch = math.ceil(len(X_train) / batch_size)
# fit the model on the batches generated by datagen.flow().
model.fit(x=datagen.flow(X_train, y_train, batch_size=batch_size),
verbose=1,
epochs=epochs,
validation_data=(X_test, y_test),
steps_per_epoch=steps_per_epoch,
callbacks=callbacks)
```
| github_jupyter |
En el anterior capítulo ya teníamos [las maletas preparadas (instalación de nuestras herramientas)](https://pybonacci.org/2019/11/12/curso-de-creacion-de-guis-con-qt5-y-python-capitulo-00-instalacion/) y empezamos a leer [nuestra guía de viajes](https://pybonacci.org/2019/11/21/curso-de-creacion-de-guis-con-qt5-y-python-capitulo-01-qt-versiones-y-bindings/) para nuestro viaje por el mundo de las aplicaciones de escritorio usando Python y Qt. **En este capítulo voy a escribir el 'Hola, Mundo' de rigor para empezar a ver conceptos clave**.
## Índice:
* [Instalación de lo que vamos a necesitar](https://pybonacci.org/2019/11/12/curso-de-creacion-de-guis-con-qt-capitulo-00:-instalacion/).
* [Qt, versiones y diferencias](https://pybonacci.org/2019/11/21/curso-de-creacion-de-guis-con-qt5-y-python-capitulo-01-qt-versiones-y-bindings/).
* [Hola, Mundo](https://pybonacci.org/2019/11/26/curso-de-creacion-de-guis-con-qt5-y-python-capitulo-02-hola-mundo/) (este capítulo).
* [Módulos en Qt](https://pybonacci.org/2019/12/02/curso-de-creacion-de-guis-con-qt5-y-python-capitulo-03-modulos-qt/).
* [Añadimos icono a la ventana principal](https://pybonacci.org/2019/12/26/curso-de-creacion-de-guis-con-qt5-y-python-capitulo-04-icono-de-la-ventana/).
* [Tipos de ventana en un GUI](https://pybonacci.org/2020/01/31/curso-de-creacion-de-guis-con-qt-capitulo-05-ventanas-principales-diferencias/).
* [Ventana de inicio – SplashScreen](https://pybonacci.org/2020/02/26/curso-de-creacion-de-guis-con-qt-capitulo-06-splash-screen/).
* [Menu principal. Introducción](https://pybonacci.org/2020/03/18/curso-de-creacion-de-guis-con-qt-capitulo-07-menu/).
* [Mejorando algunas cosas vistas](https://pybonacci.org/2020/03/26/curso-de-creacion-de-guis-con-qt-capitulo-08-mejorando-lo-visto/).
* [Gestión de eventos o Acción y reacción](https://pybonacci.org/2020/03/27/curso-de-creacion-de-guis-con-qt-capitulo-09-signals-y-slots/).
* [Introducción a Designer](https://pybonacci.org/2020/04/14/curso-de-creacion-de-guis-con-qt-capitulo-10-introduccion-a-designer/).
* [Los Widgets vistos a través de Designer: Primera parte](https://pybonacci.org/2020/05/01/curso-de-creacion-de-guis-con-qt-capitulo-11-widgets-en-designer-i/).
* [Los Widgets vistos a través de Designer: Segunda parte](https://pybonacci.org/2020/05/02/curso-de-creacion-de-guis-con-qt-capitulo-12:-widgets-en-designer-(ii)/).
* [Los Widgets vistos a través de Designer: Tercera parte](https://pybonacci.org/2020/05/03/curso-de-creacion-de-guis-con-qt-capitulo-13-widgets-en-designer-iii/).
* [Los Widgets vistos a través de Designer: Cuarta parte](https://pybonacci.org/2020/05/04/curso-de-creacion-de-guis-con-qt-capitulo-14-widgets-en-designer-iv/).
* [Los Widgets vistos a través de Designer: Quinta parte](https://pybonacci.org/2020/05/05/curso-de-creacion-de-guis-con-qt-capitulo-15-widgets-en-designer-v/).
* [Los Widgets vistos a través de Designer: Sexta parte](https://pybonacci.org/2020/05/06/curso-de-creacion-de-guis-con-qt-capitulo-16:-widgets-en-designer-(vi)/).
* TBD… (lo actualizaré cuando tenga más claro los siguientes pasos).
**[Los materiales para este capítulo los podéis descargar de [aquí](https://github.com/kikocorreoso/pyboqt/tree/chapter02)]**
**[INSTALACIÓN] Si todavía no has pasado por el [inicio del curso, donde explico cómo poner a punto todo](https://pybonacci.org/2019/11/12/curso-de-creacion-de-guis-con-qt5-y-python-capitulo-00-instalacion/), ahora es un buen momento para hacerlo y después podrás seguir con esta nueva receta.**
**[AVISO] Cuando ejecute algo de código siempre voy a asumir que os encontráis en la carpeta correcta del repositorio y que tenéis activado el entorno virtual `pyboqt`. Todos los códigos estarán en la carpeta *apps* del repositorio. Por ejemplo, si voy a ejecutar un código del capítulo 02 (este capítulo) y os digo que lo ejecutéis desde una línea de comandos usando `python programa_a_ejecutar.py` siempre voy a asumir que la línea de comandos del terminal la tenéis localizada en la carpeta donde se encuentra el programa. Por ejemplo, si quiero ejecutar el programa *main.py* del capítulo 02 deberás estar en:**
```
cd /ruta/al/repo/pyboqt/apps/02-HolaMundo # En linux, MacOS
```
```
cd C:\ruta\al\repo\pyboqt\apps\02-HolaMundo # En windows
```
**Deberás tener el entorno virtual instalado y activado. Si aun no lo tienes instalado mira en los párrafos anteriores:**
```
conda activate pyboqt
```
**Y una vez que el terminal esté ahí podréis hacer:**
```
python main.py
```
**Para que se ejecute todo correcto.**
## Hola, Mundo
Vamos a escribir nuestro primer GUI con Python y Qt y vamos a aprender una serie de conceptos sobre los GUIs, Qt,... No os preocupéis si lo que vemos no tiene mucho sentido ahora mismo. Lo veremos más en detalle en próximos capítulos.
**[Descargo de responsabilidad]** Este GUI no va a hacer nada útil.
# El Código
```python
'''
Curso de creación de GUIs con Qt5 y Python
Author: Kiko Correoso
Website: pybonacci.org
Licencia: MIT
'''
import os
os.environ['QT_API'] = 'pyside2'
import sys
from qtpy.QtWidgets import QApplication, QWidget
if __name__ == '__main__':
app = QApplication(sys.argv)
w = QWidget()
w.resize(500, 300)
w.move(0, 0)
w.setWindowTitle('Hola, Mundo')
w.show()
sys.exit(app.exec_())
```
# Ejecución del código
Guardad el anterior código en un fichero con extensión *.py*, por ejemplo, *fichero.py*. en la carpeta que queráis. Si habéis descargado el material del repositorio tendréis el anterior código en el fichero "[/ruta/al/repo/pyboqt/apps/02-HolaMundo/main.py](https://github.com/kikocorreoso/pyboqt/blob/chapter02/apps/02-HolaMundo/main.py)". Abrid un terminal o Anaconda Prompt (Windows) y escribid:
`conda activate pyboqt`
y pulsad 'Intro' para que se active el entorno virtual donde tenemos todo instalado.
En este momento vamos a instalar qtpy. Igual ya está instalado pero por si acaso. Para ello hacemos:
```
conda install qtpy -c conda-forge
```
Ahora, podéis hacer lo siguiente:
`python /ruta/al/fichero.py # Linux o Mac`
o
`python C:\ruta\al\fichero.py # Windows`
y pulsad 'Intro'. Si todo funciona correctamente os debería de salir una ventanita como la siguiente:

Enhorabuena. Has creado tu primer GUI con Qt5.
# El código explicado
-----------------
```python
import os
os.environ['QT_API'] = 'pyside2'
import sys
```
Importamos la biblioteca `sys` (luego comento para qué la usamos) y establecemos la variable de entorno `QT_API` para usar PySide2 con ayuda de la biblioteca `os` (comenta las dos primeras líneas si prefieres usar PyQt5). De momento, esto no hace ninguna magia para mostrarnos GUIs en pantalla.
----------------------
```python
from qtpy.QtWidgets import QApplication, QWidget
```
Importamos `QApplication` y `QWidget` del módulo `QtWidgets` de PyQt5 o de PySide2.
- `QApplication`: Cualquier aplicación debe crear una instancia de esta clase. Más información abajo.
- `QWidget`: Es la clase base de todos los *widgets*. Los *widgets* vienen a ser las interfaces que permiten interactuar al usuario con el GUI (botones, líneas de texto, deslizadores,...).
----------------------
```python
if __name__ == '__main__':
```
[Lee aquí, DRY](https://stackoverflow.com/questions/419163/what-does-if-name-main-do#419185).
----------------------
```
app = QApplication(sys.argv)
```
Aquí es donde creamos una instancia de `QApplication`. Todo GUI hecho con PyQt5/PySide2 debe tener una instancia de `QApplication`. Esta instancia nos proporciona acceso a información global como la carpeta de la aplicación, la hoja de estilos usada, el tamaño de la pantalla en la que corre el GUI,...
Si hiciéramos un `print(dir(app))` obtendríamos lo siguiente:
```
['ApplicationFlags', 'ColorSpec', 'CustomColor', 'ManyColor', 'NormalColor', '__class__', '__delattr__', '__dict__', '__dir__', '__doc__', '__eq__', '__format__', '__ge__', '__getattribute__', '__gt__', '__hash__', '__init__', '__init_subclass__', '__le__', '__lt__', '__ne__', '__new__', '__reduce__', '__reduce_ex__', '__repr__', '__setattr__', '__sizeof__', '__str__', '__subclasshook__', 'aboutQt', 'aboutToQuit', 'activeModalWidget', 'activePopupWidget', 'activeWindow', 'addLibraryPath', 'alert', 'allWidgets', 'allWindows', 'applicationDirPath', 'applicationDisplayName', 'applicationFilePath', 'applicationName', 'applicationNameChanged', 'applicationPid', 'applicationState', 'applicationStateChanged', 'applicationVersion', 'applicationVersionChanged', 'arguments', 'autoSipEnabled', 'beep', 'blockSignals', 'changeOverrideCursor', 'childEvent', 'children', 'clipboard', 'closeAllWindows', 'closingDown', 'colorSpec', 'commitDataRequest', 'connect', 'connectNotify', 'cursorFlashTime', 'customEvent', 'deleteLater', 'desktop', 'desktopSettingsAware', 'destroyed', 'devicePixelRatio', 'disconnect', 'disconnectNotify', 'doubleClickInterval', 'dumpObjectInfo', 'dumpObjectTree', 'dynamicPropertyNames', 'emit', 'event', 'eventDispatcher', 'eventFilter', 'exec_', 'exit', 'findChild', 'findChildren', 'flush', 'focusChanged', 'focusObject', 'focusObjectChanged', 'focusWidget', 'focusWindow', 'focusWindowChanged', 'font', 'fontDatabaseChanged', 'fontMetrics', 'globalStrut', 'hasPendingEvents', 'inherits', 'installEventFilter', 'installTranslator', 'instance', 'isEffectEnabled', 'isFallbackSessionManagementEnabled', 'isLeftToRight', 'isQuitLockEnabled', 'isRightToLeft', 'isSavingSession', 'isSessionRestored', 'isSetuidAllowed', 'isSignalConnected', 'isWidgetType', 'isWindowType', 'keyboardInputInterval', 'keyboardModifiers', 'killTimer', 'lastWindowClosed', 'layoutDirection', 'layoutDirectionChanged', 'libraryPaths', 'metaObject', 'modalWindow', 'mouseButtons', 'moveToThread', 'notify', 'objectName', 'objectNameChanged', 'organizationDomain', 'organizationDomainChanged', 'organizationName', 'organizationNameChanged', 'overrideCursor', 'palette', 'paletteChanged', 'parent', 'platformName', 'postEvent', 'primaryScreen', 'primaryScreenChanged', 'processEvents', 'property', 'queryKeyboardModifiers', 'quit', 'quitOnLastWindowClosed', 'receivers', 'registerUserData', 'removeEventFilter', 'removeLibraryPath', 'removePostedEvents', 'removeTranslator', 'restoreOverrideCursor', 'saveStateRequest', 'screenAdded', 'screenRemoved', 'screens', 'sendEvent', 'sendPostedEvents', 'sender', 'senderSignalIndex', 'sessionId', 'sessionKey', 'setActiveWindow', 'setApplicationDisplayName', 'setApplicationName', 'setApplicationVersion', 'setAttribute', 'setAutoSipEnabled', 'setColorSpec', 'setCursorFlashTime', 'setDesktopSettingsAware', 'setDoubleClickInterval', 'setEffectEnabled', 'setEventDispatcher', 'setFallbackSessionManagementEnabled', 'setFont', 'setGlobalStrut', 'setKeyboardInputInterval', 'setLayoutDirection', 'setLibraryPaths', 'setObjectName', 'setOrganizationDomain', 'setOrganizationName', 'setOverrideCursor', 'setPalette', 'setParent', 'setProperty', 'setQuitLockEnabled', 'setQuitOnLastWindowClosed', 'setSetuidAllowed', 'setStartDragDistance', 'setStartDragTime', 'setStyle', 'setStyleSheet', 'setWheelScrollLines', 'setWindowIcon', 'signalsBlocked', 'startDragDistance', 'startDragTime', 'startTimer', 'startingUp', 'staticMetaObject', 'style', 'styleHints', 'styleSheet', 'sync', 'testAttribute', 'thread', 'timerEvent', 'topLevelAt', 'topLevelWidgets', 'topLevelWindows', 'tr', 'translate', 'wheelScrollLines', 'widgetAt', 'windowIcon']
```
Si os fijáis, al instanciar `QApplication` le pasamos `sys.argv` como argumento. Esto nos permite recoger los argumentos que pasamos a través de la línea de comandos. En el caso anterior lo único que le hemos pasado a Python es el nombre del programa (`python fichero.py`) y ese sería el único argumento de entrada y que podemos obtener mediante `sys.argv[0]` (`sys.argv` es una lista).
`QApplication` puede recibir una serie de argumentos desde la línea de comandos que le permiten definir su estado interno. Algunos ya vienen predefinidos desde C++ ([más información aquí](https://doc.qt.io/qt-5/qapplication.html#QApplication)). De momento nos vale con saber esto y ya veremos más sobre esto (o no) en otras recetas.
-----------------
```python
w = QWidget()
w.resize(500, 300)
w.move(0, 0)
w.setWindowTitle('Hola, Mundo')
w.show()
```
En el ejemplo creamos una instancia de `QWidget`, que es la ventana que vemos. A `QWidget` le podríamos pasar un objeto padre (un objeto al que pertenecería nuestra instancia `w`, luego veremos más en detalle qué significa esto). Como no le pasamos ningún padre esta será la ventana padre, ventana principal de nuestra aplicación o una ventana *top level window*. La instancia `w` dispone de varios métodos. En este caso usamos:
* `resize` para darle un tamaño a la ventana (500 x 300 píxeles),
* `move` para colocar la ventana en la posición que queremos de nuestra pantalla (arriba a la izquierda en el ejemplo),
* `setWindowTitle` nos permite poner un texto en la barra de arriba de la ventana y, por último, llamamos a
* `show` para que se muestre en pantalla ya que hasta ahora solo estaba en memoria.
------------------
```python
sys.exit(app.exec_())
```
Aquí llamamos al método `exec_` de la instancia de `QApplication`. Notad que el método lleva una barra baja al final para diferenciarlo del `exec` de Python y que no haya conflictos. Cuando llamamos al método `exec_` lo que estamos haciendo es iniciar el bucle de eventos o *event loop*.
¿Qué es el bucle de eventos?
Digamos que, desde el momento en que llamamos a `app.exec_()`, la aplicación inicia el bucle de eventos y su estado pasa a ser el de esperar/escuchar/estar pendiente de eventos/señales que vengan de la propia aplicación. Estos eventos/señales pueden ser cosas como un click en un botón, un cambio de estado de una celda de una tabla (cuyo *widget* en Qt es un `QTableWidget`), un *timeout* de un temporizador,..., y la reacción del GUI a estos mismos eventos.
El bucle de eventos estará 'escuchando' hasta que llamemos al método `exit` de la instancia `app`.
¿Por qué llamamos a `app.exec_()` dentro de `sys.exit`?
De esta forma nos aseguramos que el proceso finaliza correctamente o, si ha surgido un error, podemos devolver un código de error al sistema.
## El bucle de eventos en más detalle
Voy a dejar un esquema de cómo funciona, grosso modo, el bucle de eventos principal del GUI. Este esquema está adaptado (más bien copiado) del libro de Mark Summerfield [[1](#Referencias)]:

En la anterior imagen:
- [1] indica que llamamos al método `exec_` de la instancia de `QApplication`.
- [2] Lo anterior provoca que se inicie el bucle de eventos y que nuestra aplicación se ponga a 'escuchar'. Si no ocurre ningún evento sigue esperando. Pero si ocurre un evento reacciona.
- [3] Si el evento NO es llamar al método `exit` de la instancia de `QApplication` lo procesará como corresponda y volverá a [2] a seguir esperando por nuevos eventos.
- [4] Si el evento SÍ es llamar al método `exit` de la instancia de `QApplication` entonces cerrará la aplicación y devolverá el mensaje de estátus (normalmente `0` si ha ido todo correctamente).
# Recapitulando un poco el ciclo de vida de un GUI
- La aplicación será una instancia de `QApplication`.
- En el momento que se ha instanciado la aplicación podemos llamar y mostrar nuestras ventanas.
- Arrancamos el bucle de eventos (*event loop*) de la ventana principal y Qt empieza a gestionar los eventos relacionados con la aplicación.
- Finalizamos la aplicación cuando llamamos al método `exit` de la aplicación o cuando la cerramos, como en el caso del ejemplo, pulsando la 'X' de arriba a la derecha de la ventana o pulsando 'Alt+F4'. El hecho de pulsar es un evento que recibe el *event loop* y reacciona como le hayamos dicho. En este caso, la reacción es llamar al metodo `exit` de forma implícita vía el *widget*.
# Terminando
Esto ya se ha puesto más interesante y hemos visto como crear nuestra primera ventana. Han surgido conceptos nuevos que se han explicado muy brevemente. Muchos de ellos los veremos en los próximos capítulos.
Estad atentos al siguiente capítulo.
# Referencias
- Mark Summerfield, 2007: "Rapid GUI programming with Python and Qt: the definitive guide to PyQt programming". Prentice Hall.
| github_jupyter |
```
import sys
sys.path.append('../../../')
import warnings
warnings.filterwarnings('ignore')
import fpl_analytics
import plotly.express as px
import pandas as pd
fpl_data = fpl_analytics.fetch_data_url()
fpl_analytics.save_fpl(fpl_data)
fpl_data = fpl_analytics.load_fpl(23)
base_feature=fpl_analytics.BaseWeeklyFeature(fpl_data)
avg3_feature=fpl_analytics.Avg3WeeklyFeature(fpl_data)
opp_feature=fpl_analytics.OppWeeklyFeature(fpl_data)
hist_feature=fpl_analytics.HistWeeklyFeature(fpl_data)
sparse_feature=fpl_analytics.SparseFeature(fpl_data)
pos = ["Goalkeeper", "Defender", "Midfielder", "Forward"]
training_round = [2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22]
testing_round = [2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23]
pref_lags = [0]
prediction = {}
additional_feature_list = [
lambda p, r: opp_feature.extract(p, r),
lambda p, r: hist_feature.extract(p, r),
lambda p, r: avg3_feature.extract(p, r),
lambda p, r: sparse_feature.extract(p, r)
]
for p in pos:
X_base, y = base_feature.extract(p, training_round, min_minutes=15, x_only=False, lags=pref_lags)
y = pd.to_numeric(y)
X = X_base.apply(pd.to_numeric)
for fl in additional_feature_list:
X = X.join(fl(p, training_round).apply(pd.to_numeric))
features = [{i: base_feature.extract(p, [i], min_minutes=15, lags=pref_lags) for i in testing_round}]
for fl in additional_feature_list:
features.append({i: fl(p, [i]) for i in testing_round})
choices = {
"tweedie_regress": fpl_analytics.FplTweedieRegress(X.fillna(0.0), y,
powers=[1],
alphas=[0, 0.01, 0.05, 0.1, 0.2, 0.3, 0.4,
0.5, 0.6, 0.7, 0.8, 0.9, 1.0, 2.5,
5.0, 10.0, 20.0, 50.0, 100.0, 500.0, 1000.0, 5000.0],
max_iters=[100, 200, 500],
top=3),
"huber": fpl_analytics.FplHuberRegress(X.fillna(0.0), y,
top=3),
"lasso_regress":
fpl_analytics.FplLassoRegress(X.fillna(0.0), y,
alphas=[0, 0.01, 0.05, 0.1, 0.2, 0.3, 0.4,
0.5, 0.6, 0.7, 0.8, 0.9, 1.0, 2.5,
5.0, 10.0, 20.0, 50.0, 100.0, 500.0, 1000.0, 5000.0],
top=3),
"ridge_regress":
fpl_analytics.FplRidgeRegress(X.fillna(0.0), y,
alphas=[0, 0.01, 0.05, 0.1, 0.2, 0.3, 0.4,
0.5, 0.6, 0.7, 0.8, 0.9, 1.0, 2.5,
5.0, 10.0, 20.0, 50.0, 100.0, 500.0, 1000.0, 5000.0],
top=3),
"elastic_regress":
fpl_analytics.FplElasticNetRegress(X.fillna(0.0), y,
alphas=[0, 0.01, 0.05, 0.1, 0.2, 0.3, 0.4,
0.5, 0.6, 0.7, 0.8, 0.9, 1.0, 2.5,
5.0, 10.0, 20.0, 50.0, 100.0, 500.0, 1000.0, 5000.0],
l1_ratios=[0, 0.1, 0.2, 0.3, 0.4, 0.5,
0.6, 0.7, 0.8, 0.9, 1.0],
top=3),
"random_forest":
fpl_analytics.FplRondomForest(),
"adaboost":
fpl_analytics.FplAdaBoost(X.fillna(0.0),
y,
n_estimators=750,
top=3)
}
def top_func(ts):
n = len(ts)
total = (n) * (n+1) // 2
res = 0
for i in range(n):
res += (n-i) / total * ts.iloc[i]
return res
rf = fpl_analytics.FplCVBestModel(X.fillna(0.0), y, choices, top=3, subtop=2,
top_func=top_func, best_func=fpl_analytics.mean_func)
rf.fit(X.fillna(0.0), y)
prediction[p] = {"X": X.apply(pd.to_numeric), "y": y.apply(pd.to_numeric),
"features": features,
"model": rf}
test = 23
error_threshold_point = 50
result = {}
for p in pos:
X1 = prediction[p]["features"][0][test]
for i in range(1, len(prediction[p]["features"])):
X1 = X1.join(prediction[p]["features"][i][test])
res = prediction[p]["model"].run(X1.fillna(0))
result[p] = res
points = pd.concat(result.values(),
keys=result.keys(),
names=["position", "player"],
)
points = points[points<=15]
import numpy as np
n_ava_t = []
def plot_result(df, title, width=800, height=800, num=20):
df = df.sort_values(ascending=False)
display = df.head(num)
ava_p =[]
for i in display.index:
if np.all([(a not in i) for a in n_ava_t]):
ava_p.append(i)
display = display.loc[ava_p]
print (display)
display = display.to_frame().reset_index()
display.columns = ["name", "expected"]
fig = px.bar(display, x='name', y='expected',
width=width, height=height, title=f"{p}" )
fig.show()
p = "Goalkeeper"
plot_result(points.loc[p], f"{p} week {test}")
p = "Defender"
plot_result(points.loc[p], f"{p} week {test}")
p = "Midfielder"
plot_result(points.loc[p], f"{p} week {test}")
p = "Forward"
plot_result(points.loc[p], f"{p} week {test}")
points.to_pickle(f"points_predict_{test}.pkl")
```
| github_jupyter |
```
import re
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import descartes
import geopandas as gpd
from geopandas import GeoDataFrame
from shapely.geometry import Point, Polygon
from functools import partial
import pyproj
from shapely.ops import transform
pd.set_option('display.max_columns', 500)
pd.set_option('display.max_rows', 500)
def gen_points_inside_polygon(polygon, miles=0.25):
'''Takes a polygon and number of miles as input, creates a square grid around the polygon edges
and iterates in default quarter mile steps exhaustively across the grid, returns list of all the points
generated'''
#convert miles to degrees latitude/longitude
lat_increment = miles * 0.0145
lon_increment = miles * 0.02 #this is specifically for L.A. (varies based on lat)
#get the four corners of a grid of max and min lat/long values
min_long, min_lat, max_long, max_lat = polygon.bounds
#iterate over the grid one unit at a time, append lats and lons to lists:
longs, lats = [], []
#Latitudes: start one half unit away from min_lat:
lat = min_lat + (lat_increment / 2)
while lat < max_lat:
lats.append(lat)
lat += lat_increment
lats.append(max_lat)
#repeat for longitude; conversion is diff, 1 mile = 0.0145 degrees
#start one half unit above the min_lat
lon = min_long + (lon_increment / 2)
while lon < max_long:
longs.append(lon)
#increase .02 degrees (longitude conversion for Los Angeles)
lon += lon_increment
longs.append(max_long)
#iterate thru all latitude and longitude points, instantiate shapely Point objects
#and append them to a list:
points = []
for i in range(len(longs)):
for j in range(len(lats)):
points.append(Point(longs[i], lats[j]))
#iterate over the points and throw out ones outside the polygon:
points_inside_polygon = []
for i in range(len(points)):
if polygon.contains(points[i]):
points_inside_polygon.append(points[i])
#if there are no points inside the polygon (very small polygon), add center as a point:
if len(points_inside_polygon) == 0:
points_inside_polygon.append(polygon.centroid)
#return the list of Shapely points which are bound inside the polygon
return points_inside_polygon
def make_circle(lon, lat, miles=1):
km = miles * 1.60934
proj_wgs84 = pyproj.Proj(init='epsg:4326')
# Azimuthal equidistant projection
aeqd_proj = '+proj=aeqd +lat_0={lat} +lon_0={lon} +x_0=0 +y_0=0'
project = partial(
pyproj.transform,
pyproj.Proj(aeqd_proj.format(lat=lat, lon=lon)),
proj_wgs84)
buff = Point(0, 0).buffer(km * 1000)
return Polygon(transform(project, buff).exterior.coords[:])
def get_num_groceries_in_circle(df, circle):
min_long, min_lat, max_long, max_lat = circle.bounds
df = df[(df.LATITUDE >= min_lat)&(df.LATITUDE <= max_lat)&(df.LONGITUDE>=min_long)&(df.LONGITUDE<=max_long)]
num_groceries = 0
for i in range(len(df)):
if circle.contains(df.iloc[i]['LOCATION']):
num_groceries += 1
return num_groceries
def search_for_deserts(zip_df, markets_df):
#initialize master dict which will store dicts for each zip code:
return_dict = {}
#iterate over each row in zip code dataframe:
for i in range(len(zip_df)):
zip_code = zip_df.iloc[i].zip_code
nearby_grocery_counts = []
desert_list = []
#iterate over all the test_points for each zip code:
for point in zip_df.iloc[i].test_points:
lon, lat = point.x, point.y
#make the circle around the test_point:
circle = make_circle(lon, lat)
#get the num of groceries within the circle and add to list for that zip code:
num_nearby = get_num_groceries_in_circle(markets_df, circle)
nearby_grocery_counts.append(num_nearby)
#if there are no nearby groceries, add that point to list of food deserts:
if num_nearby == 0:
desert_list.append(point)
#record the num of food deserts found for each zip code
num_food_deserts = len(desert_list)
#store all the zip_code data as dict and add to master dict:
#some zip codes are broken into mulitple polygons: this makes sure the data for these zip codes are aggregated:
if zip_code in return_dict:
return_dict[zip_code]['nearby_grocery_counts'] += nearby_grocery_counts
return_dict[zip_code]['desert_list'] += desert_list
return_dict[zip_code]['num_food_deserts'] += num_food_deserts
else:
return_dict[zip_code] = {'nearby_grocery_counts': nearby_grocery_counts,
'desert_list': desert_list, 'num_food_deserts': num_food_deserts}
#finished all iteration, now return the master dict of dictionaries for each zip code:
return return_dict
def unpack_desert_search(desert_search_results):
#initialize dict w/ empty lists
unpacked = {'zip_code': [], 'nearby_grocery_counts': [], 'desert_list': [], 'num_food_deserts': []}
#first get the zip code:
for k, v in desert_search_results.items():
unpacked['zip_code'].append(k)
#now get the other key, value pairs for that zip code:
for key, val in v.items():
unpacked[key].append(val)
#convert the dict into pandas df:
unpacked_df = pd.DataFrame(unpacked)
#initialize empty list of deserts:
desert_list = []
#iterate over the dataframe desert_list column and extract each desert and append to list:
for i in range(len(unpacked_df)):
deserts = unpacked_df.iloc[i].desert_list
if len(deserts) > 0:
for desert in deserts:
desert_list.append(desert)
#drop the desert_list column from the df:
unpacked_df = unpacked_df.drop(columns=['desert_list'])
#iterate over the grocery number counts and convert it to avg nearby groceries for each zip code:
def get_avg(num_list):
return round(sum(num_list)/len(num_list),1)
unpacked_df.nearby_grocery_counts = unpacked_df.nearby_grocery_counts.apply(get_avg)
unpacked_df = unpacked_df.rename(columns={'nearby_grocery_counts': 'avg_num_nearby_groceries'})
#return the zip_code dataframe, and the list of all food deserts found in grid search:
return unpacked_df, desert_list
#load the new markets data
df = pd.read_csv('CA.csv').drop(columns=['Unnamed: 10'])
markets = df[df.County == 'LOS ANGELES ']
#same filter as prev markets data:
exclude_string = "bakery|smoke|fizz|news|ginseng|wealth|mochi|beauty|perfume|big lots|97 CENT|marshal|water|BOYS AND GIRLS CLUB|vitamin|beer|honeybaked|sporting|ross|cnn|office|carwash|airport|service station|LUIQUOR|almonds|farm|snack|sweet|cork|baby|muscle|yogurt|godiva|stub|diy|baskin|ice cream|liq|home depot|wine|spirit|eleven|car wash|staples|am pm|99 cent|best buy|forever 21|craft|7-11|duty|sugar|mobil|gift shop|rite aid|76|petrol|weight watchers|7 eleven|golden state|dollar tree|six flags|arco|dress for less|liquor|walgreens|edible|fuel|studios|jenny craig|macy's|candies|general nutrition center|wateria|shell|oil|7-eleven|99 cents|valero|chevron|24 HOUR FITNESS|98 CENTS|candy|party|am/pm|circle k|gnc|automotive|gas|cvs|pharmacy|daiso"
markets = markets[~markets['STORE_NAME'].str.contains(exclude_string, flags=re.IGNORECASE, regex=True) == True].reset_index(drop=True)
markets = markets.rename(columns={'ZIP5':'zip_code', 'longitude':'LONGITUDE', 'latitude':'LATITUDE'})
#set dummy location column, then iterate and fill w/ Shapely Points:
markets['LOCATION'] = 'one'
for i in range(len(markets)):
lat = markets.at[i, 'LATITUDE']
lon = markets.at[i, 'LONGITUDE']
markets.at[i, 'LOCATION'] = Point(lon, lat)
# Load the zip code polygon files
polygon_file = 'CAMS_ZIPCODE_STREET_SPECIFIC.shp'
polygons = gpd.read_file(polygon_file)
polygons = polygons.to_crs(epsg=4326)
#new df w/ only the columns we need:
zip_df = polygons[['Zip_Num', 'geometry']].rename(columns={'Zip_Num':'zip_code'})
#call the function and make a new column in our zip_df that holds a lit of points inside the polygon
zip_df['test_points'] = zip_df.geometry.apply(gen_points_inside_polygon)
#create new column that displays the num of points inside the polygon
zip_df['num_points'] = zip_df['test_points'].apply(len)
polygons = gpd.read_file(polygon_file)
polygons = polygons.to_crs(epsg=4326)
polygons = polygons[['Zip_Num', 'geometry']].rename(columns={'Zip_Num':'zip_code'})
polygons.head()
zip_df.head()
markets.head()
zip_df.iloc[0].geometry
zip_df.num_points.sum()
#execute the food desert grid search and unpack the results into df and list of deserts
full_search = search_for_deserts(zip_df, markets)
zip_code_groceries_df, all_deserts = unpack_desert_search(full_search)
#use the results to plot the deserts and grocery density by zip code:
#new df w/ only the columns we need:
polygons_df = zip_df[['zip_code', 'geometry']]
#convert list of desert points from grid search into df:
geo = {'geometry': all_deserts}
deserts_gdf = gpd.GeoDataFrame(geo)
# merge zip_deserts and polygons
grocery_density_df = polygons_df.merge(zip_code_groceries_df, left_on='zip_code', right_on='zip_code', how='outer')
# Plot the zip code polygons, and then overlay the food desert points:
ax = polygons_df.plot(figsize=(12, 12), color='#00cc66', edgecolor='white')
deserts_gdf.plot(color='#00004d', alpha=0.05, ax=ax)
ax.set_facecolor('#f0f0f5')
plt.grid(linewidth=0.2)
plt.title('Food Deserts of Los Angeles County', fontsize=20)
#plt.figtext(.5,.9,'Food Deserts of Los Angeles County', fontsize=20, ha='center')
plt.xlabel('Longitude', fontsize=15)
plt.ylabel('Latitude', fontsize=15)
ax.set_ylim([33.6,35])
fig, ax = plt.subplots(figsize = (12,12))
cmap = 'Spectral_r'
#Color bar is created below
sm = plt.cm.ScalarMappable(cmap=cmap, norm=plt.Normalize(vmin=0, vmax=75))
# Empty array for the data range
sm._A = []
# Add the colorbar to the figure
cbar = fig.colorbar(sm, fraction=.011)
ax.set_facecolor('#f0f0f5')
plt.title('Grocery Store Density (Per 1-mile Radius) in Los Angeles County',fontsize=20)
plt.xlabel('Longitude', fontsize=15)
plt.ylabel('Latitude',fontsize=15)
plt.grid(linewidth=0.2)
ax.set_ylim([33.6,35])
ax_zip = grocery_density_df.plot(column='avg_num_nearby_groceries', ax=ax, cmap=cmap, vmin=0, vmax =110)
income = pd.read_csv('income_clean.csv').drop(columns=['Community']).rename(columns={'Zip Code': 'zip_code'})
income_grocery_density = income.merge(grocery_density_df, on='zip_code', how='outer')
income_grocery_density['income_binned'] = pd.cut(income_grocery_density['Estimated Median Income'], [0, 15000, 25000, 50000, 1000000])
#income_grocery_density['income_binned'] = pd.qcut(income_grocery_density['Estimated Median Income'], 10)
income_grocery_density.head()
import seaborn as sns
plt.style.use('fivethirtyeight')
flatui = ["#9b59b6", "#3498db", "#95a5a6", "#34495e", "#2ecc71"]
fig, ax = plt.subplots(figsize=(8,6))
ax.set_facecolor('#f0f0f5')
plt.xticks(rotation=45)
plt.grid(linewidth=0.2)
plt.title('Grocery Density Relative to Median Income')
#plt.xlabel('AVG number of groceies within a 1-mile radius')
sns.catplot('income_binned', 'avg_num_nearby_groceries', data=income_grocery_density, kind='bar', ci=None, palette=sns.color_palette(flatui, desat=.5), ax=ax)
ax.set(xlabel='Median Household Income Range', ylabel='AVG number of groceies within a 1-mile radius')
```
| github_jupyter |
```
import utilities as utils
# This code is used to scale to processing numerous datasets
data_path_1: str = '../../../../Data/phase1/'
data_set_1: list = [ 'Darknet_reduced_features.csv' ]
data_set: list = data_set_1
file_path_1 = utils.get_file_path(data_path_1)
file_set: list = list(map(file_path_1, data_set_1))
current_job: int = 0
utils.data_set = data_set
utils.file_set = file_set
print(f'We will be cleaning {len(file_set)} files:')
utils.pretty(file_set)
```
## Label Analysis
Now we load the data and separate the dataset by label, giving us a traffic dataset and an application dataset. We also want to investigate how merging the Non-Tor and NonVNP labels together affects the clustering, so rename the samples under these labels as regular and produce a second traffic dataset with it.
```
dataset_1 : dict = utils.examine_dataset(1)
traffic_dataset_1 : dict = utils.package_data_for_inspection_with_label(utils.prune_dataset(dataset_1, ['Application Type']), 'Traffic_Dataset_1_Tor_VPN_Non_Tor_NonVPN')
traffic_dataset_2 : dict = utils.package_data_for_inspection_with_label(utils.rename_values_in_column(traffic_dataset_1, [('Traffic Type', {'Non-Tor': 'Regular', 'NonVPN': 'Regular'})]), 'Traffic_Dataset_2_Tor_VPN_Regular')
application_dataset_1: dict = utils.package_data_for_inspection_with_label(utils.prune_dataset(dataset_1, ['Traffic Type']), 'Application_Dataset_1')
```
Now, we train deep neural networks to classify the datasets based on the labeling schemes outlaid above. We will use the confusion matrices to make inferences about the clustering of the data.
```
traffic_dataset_1['Results'] = utils.run_deep_nn_experiment(
traffic_dataset_1['Dataset'],
traffic_dataset_1['File'],
'Traffic Type',
(100, 80)
)
traffic_dataset_2['Results'] = utils.run_deep_nn_experiment(
traffic_dataset_2['Dataset'],
traffic_dataset_2['File'],
'Traffic Type',
(100, 80)
)
application_dataset_1['Results'] = utils.run_deep_nn_experiment(
application_dataset_1['Dataset'],
application_dataset_1['File'],
'Application Type',
(100, 80)
)
```
# SMOTE Prototype
Here we load up our first SMOTE and train it on the whole traffic dataset. This way we have some data to play around with and can go about devising an augmentation strategy for the experiment process.
```
fake_df_traffic_1 = utils.create_and_visualize_smote(traffic_dataset_1['Dataset'], 'Traffic Type')
fake_df_traffic_2 = utils.create_and_visualize_smote(traffic_dataset_2['Dataset'], 'Traffic Type')
fake_df_application = utils.create_and_visualize_smote(application_dataset_1['Dataset'], 'Application Type')
fake_traffic_dataset_1: dict = utils.package_data_for_inspection_with_label(
utils.prune_dataset(utils.package_data_for_inspection(fake_df_traffic_1), ['Traffic Type'] ),
'Fake_Traffic_Dataset_1_Tor_VPN_Non_Tor_NonVPN'
)
fake_traffic_dataset_2: dict = utils.package_data_for_inspection_with_label(
utils.prune_dataset(utils.package_data_for_inspection(fake_df_traffic_2), ['Traffic Type'] ),
'Fake_Traffic_Dataset_2_Tor_VPN_Regualar'
)
fake_traffic_dataset_3: dict = utils.package_data_for_inspection_with_label(
utils.prune_dataset(utils.package_data_for_inspection(fake_df_application), ['Application Type'] ),
'Fake_Traffic_Dataset_3_Application_Dataset'
)
fake_traffic_dataset_1['Dataset'].to_csv('./synthetic/smote/smote_balanced_traffic_dataset_1_traffic_labels.csv', index=False)
fake_traffic_dataset_2['Dataset'].to_csv('./synthetic/smote/smote_balanced_traffic_dataset_2_traffic_labels.csv', index=False)
fake_traffic_dataset_3['Dataset'].to_csv('./synthetic/smote/smote_balanced_application_dataset_3_traffic_labels.csv', index=False)
print(f'Last Execution: {utils.datetime.datetime.now()}')
assert False, 'Nothing after this point is included in the study'
```
| github_jupyter |
```
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
from tqdm import tqdm
%matplotlib inline
from torch.utils.data import Dataset, DataLoader
import torch
import torchvision
import torch.nn as nn
import torch.optim as optim
from torch.nn import functional as F
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
print(device)
m = 20 # 5, 10, 20, 50, 100, 500
desired_num = 2000
tr_i = 0
tr_j = int(desired_num/2)
tr_k = desired_num
tr_i, tr_j, tr_k
```
# Generate dataset
```
np.random.seed(12)
y = np.random.randint(0,10,5000)
idx= []
for i in range(10):
print(i,sum(y==i))
idx.append(y==i)
x = np.zeros((5000,2))
np.random.seed(12)
x[idx[0],:] = np.random.multivariate_normal(mean = [5,5],cov=[[0.1,0],[0,0.1]],size=sum(idx[0]))
x[idx[1],:] = np.random.multivariate_normal(mean = [6,6],cov=[[0.1,0],[0,0.1]],size=sum(idx[1]))
x[idx[2],:] = np.random.multivariate_normal(mean = [5.5,6.5],cov=[[0.1,0],[0,0.1]],size=sum(idx[2]))
x[idx[3],:] = np.random.multivariate_normal(mean = [-1,0],cov=[[0.1,0],[0,0.1]],size=sum(idx[3]))
x[idx[4],:] = np.random.multivariate_normal(mean = [0,2],cov=[[0.1,0],[0,0.1]],size=sum(idx[4]))
x[idx[5],:] = np.random.multivariate_normal(mean = [1,0],cov=[[0.1,0],[0,0.1]],size=sum(idx[5]))
x[idx[6],:] = np.random.multivariate_normal(mean = [0,-1],cov=[[0.1,0],[0,0.1]],size=sum(idx[6]))
x[idx[7],:] = np.random.multivariate_normal(mean = [0,0],cov=[[0.1,0],[0,0.1]],size=sum(idx[7]))
x[idx[8],:] = np.random.multivariate_normal(mean = [-0.5,-0.5],cov=[[0.1,0],[0,0.1]],size=sum(idx[8]))
x[idx[9],:] = np.random.multivariate_normal(mean = [0.4,0.2],cov=[[0.1,0],[0,0.1]],size=sum(idx[9]))
x[idx[0]][0], x[idx[5]][5]
for i in range(10):
plt.scatter(x[idx[i],0],x[idx[i],1],label="class_"+str(i))
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
bg_idx = [ np.where(idx[3] == True)[0],
np.where(idx[4] == True)[0],
np.where(idx[5] == True)[0],
np.where(idx[6] == True)[0],
np.where(idx[7] == True)[0],
np.where(idx[8] == True)[0],
np.where(idx[9] == True)[0]]
bg_idx = np.concatenate(bg_idx, axis = 0)
bg_idx.shape
np.unique(bg_idx).shape
x = x - np.mean(x[bg_idx], axis = 0, keepdims = True)
np.mean(x[bg_idx], axis = 0, keepdims = True), np.mean(x, axis = 0, keepdims = True)
x = x/np.std(x[bg_idx], axis = 0, keepdims = True)
np.std(x[bg_idx], axis = 0, keepdims = True), np.std(x, axis = 0, keepdims = True)
for i in range(10):
plt.scatter(x[idx[i],0],x[idx[i],1],label="class_"+str(i))
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
foreground_classes = {'class_0','class_1', 'class_2'}
background_classes = {'class_3','class_4', 'class_5', 'class_6','class_7', 'class_8', 'class_9'}
fg_class = np.random.randint(0,3)
fg_idx = np.random.randint(0,m)
a = []
for i in range(m):
if i == fg_idx:
b = np.random.choice(np.where(idx[fg_class]==True)[0],size=1)
a.append(x[b])
print("foreground "+str(fg_class)+" present at " + str(fg_idx))
else:
bg_class = np.random.randint(3,10)
b = np.random.choice(np.where(idx[bg_class]==True)[0],size=1)
a.append(x[b])
print("background "+str(bg_class)+" present at " + str(i))
a = np.concatenate(a,axis=0)
print(a.shape)
print(fg_class , fg_idx)
np.reshape(a,(2*m,1))
mosaic_list_of_images =[]
mosaic_label = []
fore_idx=[]
for j in range(desired_num):
np.random.seed(j)
fg_class = np.random.randint(0,3)
fg_idx = np.random.randint(0,m)
a = []
for i in range(m):
if i == fg_idx:
b = np.random.choice(np.where(idx[fg_class]==True)[0],size=1)
a.append(x[b])
# print("foreground "+str(fg_class)+" present at " + str(fg_idx))
else:
bg_class = np.random.randint(3,10)
b = np.random.choice(np.where(idx[bg_class]==True)[0],size=1)
a.append(x[b])
# print("background "+str(bg_class)+" present at " + str(i))
a = np.concatenate(a,axis=0)
mosaic_list_of_images.append(np.reshape(a,(2*m,1)))
mosaic_label.append(fg_class)
fore_idx.append(fg_idx)
mosaic_list_of_images = np.concatenate(mosaic_list_of_images,axis=1).T
mosaic_list_of_images.shape
mosaic_list_of_images.shape, mosaic_list_of_images[0]
for j in range(m):
print(mosaic_list_of_images[0][2*j:2*j+2])
def create_avg_image_from_mosaic_dataset(mosaic_dataset,labels,foreground_index,dataset_number, m):
"""
mosaic_dataset : mosaic_dataset contains 9 images 32 x 32 each as 1 data point
labels : mosaic_dataset labels
foreground_index : contains list of indexes where foreground image is present so that using this we can take weighted average
dataset_number : will help us to tell what ratio of foreground image to be taken. for eg: if it is "j" then fg_image_ratio = j/9 , bg_image_ratio = (9-j)/8*9
"""
avg_image_dataset = []
cnt = 0
counter = np.zeros(m) #np.array([0,0,0,0,0,0,0,0,0])
for i in range(len(mosaic_dataset)):
img = torch.zeros([2], dtype=torch.float64)
np.random.seed(int(dataset_number*10000 + i))
give_pref = foreground_index[i] #np.random.randint(0,9)
# print("outside", give_pref,foreground_index[i])
for j in range(m):
if j == give_pref:
img = img + mosaic_dataset[i][2*j:2*j+2]*dataset_number/m #2 is data dim
else :
img = img + mosaic_dataset[i][2*j:2*j+2]*(m-dataset_number)/((m-1)*m)
if give_pref == foreground_index[i] :
# print("equal are", give_pref,foreground_index[i])
cnt += 1
counter[give_pref] += 1
else :
counter[give_pref] += 1
avg_image_dataset.append(img)
print("number of correct averaging happened for dataset "+str(dataset_number)+" is "+str(cnt))
print("the averaging are done as ", counter)
return avg_image_dataset , labels , foreground_index
avg_image_dataset_1 , labels_1, fg_index_1 = create_avg_image_from_mosaic_dataset(mosaic_list_of_images[0:tr_j], mosaic_label[0:tr_j], fore_idx[0:tr_j] , 1, m)
test_dataset , labels , fg_index = create_avg_image_from_mosaic_dataset(mosaic_list_of_images[tr_j : tr_k], mosaic_label[tr_j : tr_k], fore_idx[tr_j : tr_k] , m, m)
avg_image_dataset_1 = torch.stack(avg_image_dataset_1, axis = 0)
# avg_image_dataset_1 = (avg - torch.mean(avg, keepdims= True, axis = 0)) / torch.std(avg, keepdims= True, axis = 0)
# print(torch.mean(avg_image_dataset_1, keepdims= True, axis = 0))
# print(torch.std(avg_image_dataset_1, keepdims= True, axis = 0))
print("=="*40)
test_dataset = torch.stack(test_dataset, axis = 0)
# test_dataset = (avg - torch.mean(avg, keepdims= True, axis = 0)) / torch.std(avg, keepdims= True, axis = 0)
# print(torch.mean(test_dataset, keepdims= True, axis = 0))
# print(torch.std(test_dataset, keepdims= True, axis = 0))
print("=="*40)
x1 = (avg_image_dataset_1).numpy()
y1 = np.array(labels_1)
plt.scatter(x1[y1==0,0], x1[y1==0,1], label='class 0')
plt.scatter(x1[y1==1,0], x1[y1==1,1], label='class 1')
plt.scatter(x1[y1==2,0], x1[y1==2,1], label='class 2')
plt.legend()
plt.title("dataset4 CIN with alpha = 1/"+str(m))
x1 = (test_dataset).numpy() / m
y1 = np.array(labels)
plt.scatter(x1[y1==0,0], x1[y1==0,1], label='class 0')
plt.scatter(x1[y1==1,0], x1[y1==1,1], label='class 1')
plt.scatter(x1[y1==2,0], x1[y1==2,1], label='class 2')
plt.legend()
plt.title("test dataset4")
test_dataset[0:10]/m
test_dataset = test_dataset/m
test_dataset[0:10]
class MosaicDataset(Dataset):
"""MosaicDataset dataset."""
def __init__(self, mosaic_list_of_images, mosaic_label):
"""
Args:
csv_file (string): Path to the csv file with annotations.
root_dir (string): Directory with all the images.
transform (callable, optional): Optional transform to be applied
on a sample.
"""
self.mosaic = mosaic_list_of_images
self.label = mosaic_label
#self.fore_idx = fore_idx
def __len__(self):
return len(self.label)
def __getitem__(self, idx):
return self.mosaic[idx] , self.label[idx] #, self.fore_idx[idx]
avg_image_dataset_1[0].shape
avg_image_dataset_1[0]
batch = 200
traindata_1 = MosaicDataset(avg_image_dataset_1, labels_1 )
trainloader_1 = DataLoader( traindata_1 , batch_size= batch ,shuffle=True)
testdata_1 = MosaicDataset(avg_image_dataset_1, labels_1 )
testloader_1 = DataLoader( testdata_1 , batch_size= batch ,shuffle=False)
testdata_11 = MosaicDataset(test_dataset, labels )
testloader_11 = DataLoader( testdata_11 , batch_size= batch ,shuffle=False)
class Whatnet(nn.Module):
def __init__(self):
super(Whatnet,self).__init__()
self.linear1 = nn.Linear(2,3)
# self.linear2 = nn.Linear(50,10)
# self.linear3 = nn.Linear(10,3)
torch.nn.init.xavier_normal_(self.linear1.weight)
torch.nn.init.zeros_(self.linear1.bias)
def forward(self,x):
# x = F.relu(self.linear1(x))
# x = F.relu(self.linear2(x))
x = (self.linear1(x))
return x
def calculate_loss(dataloader,model,criter):
model.eval()
r_loss = 0
with torch.no_grad():
for i, data in enumerate(dataloader, 0):
inputs, labels = data
inputs, labels = inputs.to("cuda"),labels.to("cuda")
outputs = model(inputs)
loss = criter(outputs, labels)
r_loss += loss.item()
return r_loss/(i+1)
def test_all(number, testloader,net):
correct = 0
total = 0
out = []
pred = []
with torch.no_grad():
for data in testloader:
images, labels = data
images, labels = images.to("cuda"),labels.to("cuda")
out.append(labels.cpu().numpy())
outputs= net(images)
_, predicted = torch.max(outputs.data, 1)
pred.append(predicted.cpu().numpy())
total += labels.size(0)
correct += (predicted == labels).sum().item()
pred = np.concatenate(pred, axis = 0)
out = np.concatenate(out, axis = 0)
print("unique out: ", np.unique(out), "unique pred: ", np.unique(pred) )
print("correct: ", correct, "total ", total)
print('Accuracy of the network on the %d test dataset %d: %.2f %%' % (total, number , 100 * correct / total))
def train_all(trainloader, ds_number, testloader_list):
print("--"*40)
print("training on data set ", ds_number)
torch.manual_seed(12)
net = Whatnet().double()
net = net.to("cuda")
criterion_net = nn.CrossEntropyLoss()
optimizer_net = optim.Adam(net.parameters(), lr=0.001 ) #, momentum=0.9)
acti = []
loss_curi = []
epochs = 1000
running_loss = calculate_loss(trainloader,net,criterion_net)
loss_curi.append(running_loss)
print('epoch: [%d ] loss: %.3f' %(0,running_loss))
for epoch in range(epochs): # loop over the dataset multiple times
ep_lossi = []
running_loss = 0.0
net.train()
for i, data in enumerate(trainloader, 0):
# get the inputs
inputs, labels = data
inputs, labels = inputs.to("cuda"),labels.to("cuda")
# zero the parameter gradients
optimizer_net.zero_grad()
# forward + backward + optimize
outputs = net(inputs)
loss = criterion_net(outputs, labels)
# print statistics
running_loss += loss.item()
loss.backward()
optimizer_net.step()
running_loss = calculate_loss(trainloader,net,criterion_net)
if(epoch%200 == 0):
print('epoch: [%d] loss: %.3f' %(epoch + 1,running_loss))
loss_curi.append(running_loss) #loss per epoch
if running_loss<=0.05:
print('epoch: [%d] loss: %.3f' %(epoch + 1,running_loss))
break
print('Finished Training')
correct = 0
total = 0
with torch.no_grad():
for data in trainloader:
images, labels = data
images, labels = images.to("cuda"), labels.to("cuda")
outputs = net(images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the %d train images: %.2f %%' % (total, 100 * correct / total))
for i, j in enumerate(testloader_list):
test_all(i+1, j,net)
print("--"*40)
return loss_curi
train_loss_all=[]
testloader_list= [ testloader_1, testloader_11]
train_loss_all.append(train_all(trainloader_1, 1, testloader_list))
%matplotlib inline
for i,j in enumerate(train_loss_all):
plt.plot(j,label ="dataset "+str(i+1))
plt.xlabel("Epochs")
plt.ylabel("Training_loss")
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
```
| github_jupyter |
# Harmonizing Multi-Site Structural MRI Data Using ComBat
Team contributors: David MacDonald, and anyone else who is interested!
## Summary
This project involves using [ComBat](https://github.com/jfortin1/ComBatHarmonization), an open-source library for multi-site data harmonization, to remove site effects from subcortical volumetric data derived from the [ABIDE](http://fcon_1000.projects.nitrc.org/indi/abide/) dataset. If time permits, I would also like to explore systematic ways of comparing different site harmonization approaches.
# Project Definition
#### My Background
* Elementary school teacher from Ottawa, ON
* Currently Master's student in McGill's [CoBRA Lab](http://cobralab.ca)
* Some programming background (C++ and R)
#### Neuroscience Background
* Prior experience in a wet lab, doing molecular biology and immunohistochemistry
* Now studying Autism Spectrum Disorders and subcortical anatomy, using structural MRI
### Background
#### Project Background
* Working with large, multi-site dataset (ABIDE-Plus!)
* Investigating subcortical volumes and morphometry
* Thalamus, globus pallidus, caudate nucleus
* Dealing with site effects using meta-analytic techniques
# Project Definition
With this project, I would like to learn and apply two other techniques for inter-site data harmonization:
* Use the open source ComBat library, which was originally intended to mitigate batch effects in genetic studies, but has since been adapted to work with neuroimaging data.
* Use linear mixed-effect models.
I would also like to:
* Use the tools we have been learning about to make my project more transparent, easy to maintain, reproducible, and open to collaboration.
* If time permits, find a way to compare these three site harmonization techniques systematically.
### Tools
This project will make use of:
* git and github for version control, code sharing, project management, and collaboration
* Jupyter notebooks
* Python, including packages for linear regression and mixed models (numpy, statsmodels) and data manipulation (pandas)
* A virtualization technology, either conda or virtual environment, to improve reproducibility
* Python libraries for data visualization (matplotlib, Seaborn)
* [ComBat](https://github.com/Jfortin1/ComBatHarmonization): A linear regression tool that uses Empirical Bayes to model site effects
### Data
I will be using subcortical volumes, previously derived from the [ABIDE](http://fcon_1000.projects.nitrc.org/indi/abide/) dataset using the [MAGeT Brain](https://github.com/CobraLab/MAGeTbrain) pipeline.
 
### Deliverables
* github repository containing
* Analysis code in Jupyter notebooks and the subcortical volume data to be processed
* All visualizations for the linear mixed-models and ComBat harmonization implementations
* Virtual environment requirements.txt file, or conda environment.yaml file
* README.md file summarizing the background methodology, and results
* Link to final presentation slides
| github_jupyter |
## Experiment Setup
### Random seed / Colab / CUDA related
```
import time
import datetime
import os
import sys
import itertools
# Use Google Colab
use_colab = True
# Is this notebook running on Colab?
# If so, then google.colab package (github.com/googlecolab/colabtools)
# should be available in this environment
# Previous version used importlib, but we could do the same thing with
# just attempting to import google.colab
try:
from google.colab import drive
colab_available = True
except:
colab_available = False
if use_colab and colab_available:
drive.mount('/content/drive')
# If there's a package I need to install separately, do it here
!pip install pyro-ppl
# Install ronald_bdl package
!pip install git+https://github.com/ronaldseoh/ronald_bdl.git
# cd to the appropriate working directory under my Google Drive
%cd 'drive/My Drive/Colab Notebooks/bayesian-dl-experiments'
# List the directory contents
!ls
# IPython reloading magic
%load_ext autoreload
%autoreload 2
# Random seeds
# Based on https://pytorch.org/docs/stable/notes/randomness.html
random_seed = 682
```
### Third party libraries (NumPy, PyTorch, Pyro)
```
# Third party libraries import
import numpy as np
import torch
import pyro
import matplotlib.pyplot as plt
import tqdm
# Print version information
print("Python Version: " + sys.version)
print("NumPy Version: " + np.__version__)
print("PyTorch Version: " + torch.__version__)
print("Pyro Version: " + pyro.__version__)
# More imports...
from torch import nn, optim
from torch.utils.data import random_split, DataLoader, RandomSampler
import torchvision
import torchvision.transforms as transforms
from pyro.infer import SVI, Trace_ELBO, HMC, MCMC
# Import model and dataset classes from ronald_bdl
from ronald_bdl import models, datasets
from ronald_bdl.models import utils
# pyplot setting
%matplotlib inline
# torch.device / CUDA Setup
use_cuda = True
if use_cuda and torch.cuda.is_available():
torch_device = torch.device('cuda')
torch.backends.cudnn.deterministic = True
# Disable 'benchmark' mode
# Note: https://discuss.pytorch.org/t/what-does-torch-backends-cudnn-benchmark-do/5936
torch.backends.cudnn.benchmark = False
use_pin_memory = True # Faster Host to GPU copies with page-locked memory
# CUDA libraries version information
print("CUDA Version: " + str(torch.version.cuda))
print("cuDNN Version: " + str(torch.backends.cudnn.version()))
print("CUDA Device Name: " + str(torch.cuda.get_device_name()))
print("CUDA Capabilities: "+ str(torch.cuda.get_device_capability()))
else:
torch_device = torch.device('cpu')
use_pin_memory = False
```
### Toy dataset settings
```
"""
Toy dataset generation based on the experiment from
the Probabilstic Backpropagation paper (Hernandez-Lobato & Adams, 2015)
"""
# Random seed for toy dataset
dataset_toy_random_seed = 691
# Toy dataset size
dataset_toy_size = 20
random_x = True
# Toy dataset x distribution (uniform) parameters
dataset_toy_x_low = -4
dataset_toy_x_high = 4
# Toy dataset y noise distribution (normal noise)
dataset_toy_y_noise_mean = 0
dataset_toy_y_noise_std = 9
# Toy dataset additional noise
dataset_toy_more_noise_x_from=None
dataset_toy_more_noise_x_to=dataset_toy_x_high
dataset_toy_more_noise_y_from=50
dataset_toy_more_noise_y_to=100
```
### MC dropout parameters
```
"""
MC Dropout related
"""
# Dropout rate
dropout_rate = 0.1
# Length scale
length_scale = 0.01
# tau
tau = 0.25
# reg strength
reg_strength = utils.reg_strength(dropout_rate, length_scale, dataset_toy_size, tau)
print('reg_strength = ' + str(reg_strength))
# Epochs
n_epochs = 40000
# Optimizer learning rate
learning_rate = 0.001
# Loss function type
loss_function_type = 'log_gaussian_loss'
# Learn hetero noise?
learn_hetero_noise = True
# Number of test predictions (for each data point)
n_predictions = 1000
```
### Directory path to save results
```
# Test start time
test_start_time = datetime.datetime.today().strftime('%Y%m%d%H%M')
# Directory to store the results for this experiment
test_results_path = os.path.join(
'./test_results',
'comparison_toy',
(
test_start_time
+ '_' + str(n_epochs)
+ '_' + str(dropout_rate)
+ '_' + str(length_scale)
+ '_' + str(tau)
+ '_' + str(loss_function_type)
+ '_' + str(n_predictions))
)
# Create the directory if it doesn't exist
os.makedirs(test_results_path, exist_ok=True)
```
## Prepare data
### Get the data as a torch Dataset object
```
dataset = datasets.ToyDatasets(
random_seed=dataset_toy_random_seed,
n_samples=dataset_toy_size,
x_low=dataset_toy_x_low,
x_high=dataset_toy_x_high,
random_x=random_x,
y_noise_mean=dataset_toy_y_noise_mean,
y_noise_std=dataset_toy_y_noise_std,
more_noise_x_from=dataset_toy_more_noise_x_from,
more_noise_x_to=dataset_toy_more_noise_x_to,
more_noise_y_from=dataset_toy_more_noise_y_from,
more_noise_y_to=dataset_toy_more_noise_y_to,
)
# Get the test data
X_test = torch.linspace(dataset_toy_x_low-5, dataset_toy_x_high+5, 100).reshape(100, 1)
y_test = torch.pow(X_test, 3)
# Send it to the memory of the device to be used for training
X_test = X_test.to(torch_device)
y_test = y_test.to(torch_device)
# Save a copy of X_test at CPU for plotting
X_test_cpu = X_test.cpu()
# Print the size of the training set
print("dataset size = " + str((len(dataset), dataset.n_features)))
```
## Declare models
```
uncertainty_methods = {}
# MC (Bernoulli) dropout
do_mc_dropout = True
if do_mc_dropout:
uncertainty_methods['mc_dropout_relu'] = models.FCNet(
input_dim=dataset.n_features,
output_dim=dataset.n_targets,
hidden_dim=100,
n_hidden=0,
dropout_rate=dropout_rate,
dropout_type='bernoulli',
learn_hetero=learn_hetero_noise,
)
# uncertainty_methods['mc_dropout_tanh'] = models.FCNet(
# input_dim=dataset.n_features,
# output_dim=dataset.n_targets,
# hidden_dim=100,
# n_hidden=1,
# dropout_rate=dropout_rate,
# dropout_type='bernoulli',
# nonlinear_type='tanh',
# learn_hetero=learn_hetero_noise,
# )
# uncertainty_methods['mc_dropout_sigmoid'] = models.FCNet(
# input_dim=dataset.n_features,
# output_dim=dataset.n_targets,
# hidden_dim=100,
# n_hidden=0,
# dropout_rate=dropout_rate,
# dropout_type='bernoulli',
# nonlinear_type='sigmoid',
# learn_hetero=True,
# )
# Vanilla Variational Inference
do_vi = False
if do_vi:
uncertainty_methods['vi'] = models.FCNetPyro(
input_dim=dataset.n_features,
output_dim=dataset.n_targets,
hidden_dim=100,
n_hidden=0,
torch_device=torch_device,
)
# HMC
do_hmc = False
if do_hmc:
uncertainty_methods['hmc'] = models.FCNetPyro(
input_dim=dataset.n_features,
output_dim=dataset.n_targets,
hidden_dim=100,
n_hidden=0,
torch_device=torch_device,
)
```
## Train the models
### Train/test the model
```
for key, network in uncertainty_methods.items():
# Reset the random number generator for each method (to produce identical results)
torch.manual_seed(random_seed)
np.random.seed(random_seed)
pyro.set_rng_seed(random_seed)
# Print the method name
print("Now running " + str(key))
# Send the whole model to the selected torch.device
network.to(torch_device)
# Print the network structure
print(network)
print()
# Initialize training data loader
train_loader = DataLoader(dataset, batch_size=128)
"""
Optimizer Setup
"""
if isinstance(network, models.FCNet):
# Adam optimizer
# https://pytorch.org/docs/stable/optim.html?highlight=adam#torch.optim.Adam
# NOTE: Need to set L2 regularization from here
optimizer = torch.optim.Adam(
network.parameters(),
lr=learning_rate,
weight_decay=reg_strength, # L2 regularization
)
elif isinstance(network, models.FCNetPyro):
# Clear everything in Pyro parameter storage
# Looks like this parameter store would be globally shared across
# models in the same Python instance.
pyro.clear_param_store()
# Force all the PyTorch tensors to be CUDA tensors if available
if use_cuda and torch.cuda.is_available():
torch.set_default_tensor_type(torch.cuda.FloatTensor)
# Different optimizers for VI and HMC
if key == 'vi':
optimizer = pyro.optim.Adam({"lr": 0.01})
svi = SVI(network, network.guide, optimizer, loss=Trace_ELBO())
elif key == 'hmc':
kernel = HMC(network)
mcmc = MCMC(kernel, num_samples=10000)
"""
Training
"""
# Model to train mode
network.train()
# Record training start time (for this method)
tic = time.time()
if key == 'hmc':
# to() actually returns new Tensor; need to replace with that return
dataset.data_x = dataset.data_x.to(torch_device)
dataset.data_y = dataset.data_y.to(torch_device)
mcmc.run(dataset.data_x, dataset.data_y)
else:
progress_bar = tqdm.tqdm(range(n_epochs))
for epoch in progress_bar: # loop over the dataset multiple times
for i, data in enumerate(train_loader):
# get the inputs; data is a list of [inputs, labels]
inputs, targets = data
# Store the batch to torch_device's memory
# to() actually returns new Tensor; need to replace with that return
inputs = inputs.to(torch_device)
targets = targets.to(torch_device)
if isinstance(network, models.FCNet):
# zero the parameter gradients
optimizer.zero_grad()
# forward + backward + optimize
outputs, noises = network(inputs)
# Mean Squared Error for loss function to minimize
if loss_function_type == 'mse_loss':
total_loss = torch.nn.MSELoss()(outputs, targets)
elif loss_function_type == 'log_gaussian_loss':
total_loss = utils.log_gaussian_loss(outputs, targets, noises.exp())
total_loss.backward()
optimizer.step()
elif isinstance(network, models.FCNetPyro):
pyro.clear_param_store()
# calculate the loss and take a gradient step
total_loss = svi.step(inputs, targets)
progress_bar.set_description("loss: %5.4f" % total_loss)
# Record training end time
toc = time.time()
# Report the final loss
print("final loss = " + str(total_loss))
# Report the total training time
print("training time = " + str(toc - tic) + " seconds")
print()
```
## Plot the results
```
for key, network in uncertainty_methods.items():
"""
Testing
"""
# Model to eval mode
network.eval()
# Record testing start time (for this split)
tic_testing = time.time()
predictions_non_mc, _ = network(X_test)
predictions, mean, var, noises, metrics = network.predict_dist(
X_test, n_predictions,
y_test=y_test,
tau=torch.tensor(tau, dtype=torch.float))
# Record testing end time
toc_testing = time.time()
# Report the total testing time
print("testing time = " + str(toc_testing - tic_testing) + " seconds")
# These can now be stored in CPU memory for plotting
predictions_non_mc = predictions_non_mc.cpu()
predictions = predictions.cpu()
mean = mean.cpu()
var = var.cpu()
noises = noises.cpu()
total_uncertainty = var + noises
"""
Print results
"""
print()
print("Mean = " + str(mean.flatten()))
print("Variance = " + str(var.flatten()))
print("Noise = " + str(noises.flatten()))
# store additional metrics
if len(metrics) > 0:
for metric_key, value in metrics.items():
print(str(metric_key) + " = " + str(value))
print()
# Plot the uncertainty measured by each methods
plt.figure()
# Fix the scales of x-axis and y-axis
plt.xlim(dataset_toy_x_low-1, dataset_toy_x_high+1)
plot_ylim_low = dataset_toy_x_low ** 3 - 2 * dataset_toy_y_noise_std
plot_ylim_high = dataset_toy_x_high ** 3 + 2 * dataset_toy_y_noise_std
plt.ylim(plot_ylim_low, plot_ylim_high)
# Distributions around the predictions
plt.fill_between(
X_test_cpu.flatten(),
(mean.detach().numpy() - 2*torch.sqrt(total_uncertainty).detach().numpy()).flatten(),
(mean.detach().numpy() + 2*torch.sqrt(total_uncertainty).detach().numpy()).flatten(),
color='lightblue', alpha=0.6, label="Total predictive uncertainty")
# epistemic
plt.fill_between(
X_test_cpu.flatten(),
(mean.detach().numpy() - 2*torch.sqrt(var).detach().numpy()).flatten(),
(mean.detach().numpy() + 2*torch.sqrt(var).detach().numpy()).flatten(),
color='red', alpha=0.1, label="Epistemic uncertainty")
# Train data
plt.plot(dataset.data_x.cpu(), dataset.data_y.cpu(), 'or', label='Training data', alpha=0.3)
# Non-MC prediction
plt.plot(X_test_cpu, predictions_non_mc.detach().numpy(), 'green', label='non-MC prediction', alpha=0.9)
# MC prediction
plt.plot(X_test_cpu, mean.detach().numpy(), 'blue', label='mean prediction', alpha=0.7)
# Original data generating function without noise
plt.plot(X_test_cpu, X_test_cpu ** 3, 'gray', label='data generating function')
plt.legend(loc=2, prop={'size': 8})
plt.savefig(os.path.join(test_results_path, str(key) + '.png'), dpi=200)
plt.show()
```
| github_jupyter |
# Python Module: Affine Cipher Encryption
The affine cipher is more complicated than the shift cipher. This is good for encryption security, but makes the cipher even more cumbersome to implement by hand. In this Python module, you'll write a function that takes as inputs a plaintext message $P$, multiplicative key $m$, and additive key $k$, and outputs the ciphertext $C$ via the equation $C\equiv mP+k\mod{26}$. Then, you'll encrypt several messages with this function. Finally, you'll develop a new function that allows you to use a larger alphabet to affine-encipher messages, for example by adding characters such as !, ?, and *.
Let's begin by outlining the code necessary to encrypt a message using the affine cipher. Luckily, it only takes a small modification of your `ShiftEncrypt` code to encrypt using an affine cipher. Instead of adding $k$, our shift, and then reducing $\mod 26$, we're first multiplying by $m$, adding $k$, and reducing the result $\mod 26$.
The code to multiply 3 by 7 in Python is `3*7`. The code for the linear function $y=3x+7$ in Python is `y=3*x+7`.
If the plaintext letter in our code was 'n', which is 13 on our letter-to-number chart, write the code to encrypt 'n' using the affine cipher $C\equiv 3P+7\mod{26}$ in the chunk below. Test that, when you run the code, you get the number $20$, corresponding to the ciphertext letter U.
```
# output the ciphertext number corresponding to the plaintext number 13 under the affine cipher C = 3*P + 7 mod 26
Mod(3*13+7, 26)
```
Now, the plaintext letter that we're encrypting is not always going to be M, and our keys are not always going to be $m=3$ and $k=7$. Below, write the code to multiply the variable $m$ times the variable $P$, then add $k$. Finally, wrap this code in the parentheses after the word Mod(). **Don't run the code chunk below.**
Below, under the line `def AffineEncrypt(plaintext, m, k)`, copy-paste the code under the line `def ShiftEncrypt(plaintext, key)` from the `ShiftEncrypt` module (toward the very end, where you're defining the `ShiftEncrypt` function for the first time). Finally, change your code so instead of just adding $k$, we're multiplying by $m$ first and then adding $k$. Then run the chunk below (nothing should happen yet).
```
# Copy-paste your ShiftEncrypt function below and then make the designated changes.
def AffineEncrypt(plaintext, m, k):
alph = 'ABCDEFGHIJKLMNOPQRSTUVWXYZ'
ciphertext = ''
for plaintext_letter in plaintext:
plaintext_number = alph.find(plaintext_letter)
ciphertext_number = Mod(m*plaintext_number + k, 26)
ciphertext_letter = alph[ciphertext_number]
ciphertext += ciphertext_letter
print(ciphertext)
```
## Now it's your turn!
Use your affine encryption function to encrypt the following messages with their corresponding keys:
1. "GOGRIFFINS", $m = 5$, $k = 7$
2. "CRYPTOGRAPHYISAWESOME", $m = 17$, $k = 4$
3. "STOPHAMMERTIME", $m = 45$, $k = 32$.
4. "NANANANANANANANANABATMAN", $m = 13$, $k = 0$.
```
### Write your code for (1)-(4) here ###
```
Look at the way Sage enciphered "STOPHAMMERTIME" with multiplicative key 45 and additive key 32. Using modular arithmetic, rewrite the equation $C\equiv 45P+32\mod{26}$ to an equivalent equation in which each number is reduced mod 26. Write that equation below; you may use = instead of $\equiv$ for congruence. Test your equation by encrypting the letter "S" with it on a separate sheet of paper. Does the resulting ciphertext match that of the function?
write your answer here
What happens to the word "Batman" in (4)? Why?
(write your answer here)
Finally, define a function that uses a larger alphabet of your choice. For example, you could add the special characters ~!? to your alphabet. Be sure to adjust the modulus accordingly! Then, use this function to encrypt a message of your choice.
```
def AffineEncryptX(plaintext, m, k):
### Write your code here ###
```
| github_jupyter |
```
import onnx
from onnx.tools import update_model_dims
import numpy as np
import onnx.helper as helper
from onnx import shape_inference, TensorProto
import sys
import torch
import torchvision
from transformers import AutoTokenizer, AutoModel
from pathlib import Path
from transformers.convert_graph_to_onnx import convert
from transformers import AutoTokenizer, AutoModel
tokenizer = AutoTokenizer.from_pretrained("bert-base-uncased")
model = AutoModel.from_pretrained("bert-base-uncased")
inputs = tokenizer("Hello world!", return_tensors="pt")
outputs = model(**inputs)
print(outputs)
from transformers import AutoModelForSequenceClassification
model = AutoModelForSequenceClassification.from_pretrained("bert-base-cased", num_labels=2)
model = onnx.load("./models/bert-base/bert-base-cased.onnx")
oldnodes = [n for n in model.graph.node]
newnodes = oldnodes[0:125]
for n in model.graph.node:
model.graph.node.remove(n)
model.graph.node.extend(newnodes)
onnx.save(model, "./models_out/bert-1-v2.onnx")
graph = helper.GraphProto()
graph.node.extend(newnodes)
model_def = helper.make_model(graph, producer_name='onnx-example')
onnx.save(model_def, "./models_out/bert-1-v3.onnx")
graph_def = helper.make_graph(
['Mul_123'],
'Mul_123',
inputs=['A','B'], # 输入
outputs=['C'], # 输出
)
model = onnx.load("./models/vgg16.onnx")
for i in range(20):
print(model.graph.initializer[i].dims)
input_path = './models/vgg16.onnx'
output_path = './models_out/vgg16_out.onnx'
input_names = ['100']
output_names = ['121']
onnx.utils.extract_model(input_path, output_path, input_names, output_names)
input_path = './models/bert-base/bert-base-cased.onnx'
output_path = './models_out/bert_out.onnx'
# input_names = ['input_ids', 'attention_mask', 'token_type_ids']
input_names = ['235']
output_names = ['351']
onnx.utils.extract_model(input_path, output_path, input_names, output_names)
model = onnx.load("./models/bert-base/bert-base-cased.onnx")
onnx.checker.check_model(model)
class Extractor:
def __init__(self, model): # type: (ModelProto) -> None
self.model = onnx.shape_inference.infer_shapes(model)
self.graph = self.model.graph
self.wmap = self._build_name2obj_dict(self.graph.initializer)
self.vimap = self._build_name2obj_dict(self.graph.value_info)
@staticmethod
def _build_name2obj_dict(objs): # type: ignore
return {obj.name: obj for obj in objs}
def _collect_new_io_core(self, original_io, io_names_to_extract): # type: ignore
original_io_map = self._build_name2obj_dict(original_io)
original_io_names = set(original_io_map.keys())
s_io_names_to_extract = set(io_names_to_extract)
io_names_to_keep = s_io_names_to_extract & original_io_names
new_io_names_to_add = s_io_names_to_extract - original_io_names
new_io_tensors = []
for name in io_names_to_keep:
new_io_tensors.append(original_io_map[name])
for name in new_io_names_to_add:
# activation become input or output
new_io_tensors.append(self.vimap[name])
# adjust sequence
new_io_tensors_map = self._build_name2obj_dict(new_io_tensors)
return [new_io_tensors_map[name] for name in io_names_to_extract]
def _collect_new_inputs(self, names): # type: (List[Text]) -> List[ValueInfoProto]
return self._collect_new_io_core(self.graph.input, names) # type: ignore
def _collect_new_outputs(self, names): # type: (List[Text]) -> List[ValueInfoProto]
return self._collect_new_io_core(self.graph.output, names) # type: ignore
def _dfs_search_reachable_nodes(
self,
node_output_name, # type: Text
graph_input_names, # type: List[Text]
reachable_nodes, # type: List[NodeProto]
): # type: (...) -> None
if node_output_name in graph_input_names:
return
for node in self.graph.node:
if node in reachable_nodes:
continue
if node_output_name not in node.output:
continue
reachable_nodes.append(node)
for name in node.input:
self._dfs_search_reachable_nodes(name, graph_input_names, reachable_nodes)
def _collect_reachable_nodes(
self,
input_names, # type: List[Text]
output_names, # type: List[Text]
): # type: (...) -> List[NodeProto]
reachable_nodes = list() # type: ignore
for name in output_names:
self._dfs_search_reachable_nodes(name, input_names, reachable_nodes)
# needs to be topology sorted.
nodes = [n for n in self.graph.node if n in reachable_nodes]
return nodes
def _collect_reachable_tensors(
self,
nodes, # type: List[NodeProto]
): # type: (...) -> Tuple[List[TensorProto], List[ValueInfoProto]]
all_tensors_name = set()
for node in nodes:
for name in node.input:
all_tensors_name.add(name)
for name in node.output:
all_tensors_name.add(name)
initializer = [self.wmap[t] for t in self.wmap.keys() if t in all_tensors_name]
value_info = [self.vimap[t] for t in self.vimap.keys() if t in all_tensors_name]
assert(len(self.graph.sparse_initializer) == 0)
assert(len(self.graph.quantization_annotation) == 0)
return (initializer, value_info)
def _make_model(
self,
nodes, # type: List[NodeProto]
inputs, # type: List[ValueInfoProto]
outputs, # type: List[ValueInfoProto]
initializer, # type: List[TensorProto]
value_info # type: List[ValueInfoProto]
): # type: (...) -> ModelProto
name = 'Extracted from {' + self.graph.name + '}'
graph = onnx.helper.make_graph(nodes, name, inputs, outputs, initializer=initializer,
value_info=value_info)
meta = {
'ir_version': self.model.ir_version,
'opset_imports': self.model.opset_import,
'producer_name': 'onnx.utils.extract_model',
}
return onnx.helper.make_model(graph, **meta)
def extract_model(
self,
input_names, # type: List[Text]
output_names, # type: List[Text]
): # type: (...) -> ModelProto
inputs = self._collect_new_inputs(input_names)
outputs = self._collect_new_outputs(output_names)
nodes = self._collect_reachable_nodes(input_names, output_names)
initializer, value_info = self._collect_reachable_tensors(nodes)
model = self._make_model(nodes, inputs, outputs, initializer, value_info)
return model
input_path = './models/bert-base/bert-base-cased.onnx'
output_path = './models_out/bert_out.onnx'
input_names = ['235']
output_names = ['351']
onnx.checker.check_model(input_path)
model = onnx.load(input_path)
e = Extractor(model)
extracted = e.extract_model(input_names, output_names)
onnx.save(extracted, output_path)
extracted = onnx.shape_inference.infer_shapes(extracted)
onnx.save(extracted, './models_out/bert_out_2.onnx')
! mkdir ./models/bert-text-classification
from pathlib import Path
from transformers.convert_graph_to_onnx import convert
# Handles all the above steps for you
p = {"num_labels": 2}
convert(framework="pt", model="bert-base-cased", output=Path("./models/bert-text-classification/bert-cls.onnx"), opset=11, pipeline_name="text-classification", **p)
from transformers import AutoModelForSequenceClassification
model = AutoModelForSequenceClassification.from_pretrained("bert-base-cased", num_labels=2)
from transformers import TrainingArguments
training_args = TrainingArguments("test_trainer")
from datasets import load_dataset
from datasets import load_dataset
from datasets import list_datasets
datasets_list = list_datasets()
len(datasets_list)
raw_datasets = load_dataset("imdb")
from transformers import AutoTokenizer
tokenizer = AutoTokenizer.from_pretrained("bert-base-cased")
def tokenize_function(examples):
return tokenizer(examples["text"], padding="max_length", truncation=True)
tokenized_datasets = raw_datasets.map(tokenize_function, batched=True)
small_train_dataset = tokenized_datasets["train"].shuffle(seed=42).select(range(1000))
small_eval_dataset = tokenized_datasets["test"].shuffle(seed=42).select(range(1000))
full_train_dataset = tokenized_datasets["train"]
full_eval_dataset = tokenized_datasets["test"]
from transformers import AutoModelForSequenceClassification
model = AutoModelForSequenceClassification.from_pretrained("bert-base-cased", num_labels=2)
from transformers import TrainingArguments
training_args = TrainingArguments("test_trainer")
from transformers import Trainer
trainer = Trainer(
model=model, args=training_args, train_dataset=small_train_dataset, eval_dataset=small_eval_dataset
)
trainer.train()
```
| github_jupyter |
# Operator Overview
This notebook shows how to use the `Operator` class from the *Quantum Information* module of Qiskit to create custom matrix operators and custom unitary gates, and to evaluate the unitary matrix for a quantum circuit.
```
import numpy as np
from qiskit import QuantumCircuit, ClassicalRegister, QuantumRegister
from qiskit import execute, BasicAer
from qiskit.compiler import transpile
from qiskit.quantum_info.operators import Operator, Pauli
from qiskit.quantum_info import process_fidelity
from qiskit.extensions import RXGate, CnotGate, XGate
```
## Operator Class
The `Operator` class is used in Qiskit to represent matrix operators acting on a quantum system. It has several methods to build composite operators using tensor products of smaller operators, and to compose operators.
### Creating Operators
The easiest way to create an operator object is to initialize it with a matrix given as a list or a Numpy array. For example, to create a two-qubit Pauli-XX operator:
```
XX = Operator([[0, 0, 0, 1], [0, 0, 1, 0], [0, 1, 0, 0], [1, 0, 0, 0]])
XX
```
### Operator Properties
The operator object stores the underlying matrix, and the input and output dimension of subsystems.
* `data`: To access the underly Numpy array, we may use the `Operator.data` property.
* `dims`: To return the total input and output dimension of the operator, we may use the `Operator.dim` property. *Note: the output is returned as a tuple `(input_dim, output_dim)`, which is the reverse of the shape of the underlying matrix.*
```
XX.data
input_dim, output_dim = XX.dim
input_dim, output_dim
```
### Input and Output Dimensions
The operator class also keeps track of subsystem dimensions, which can be used for composing operators together. These can be accessed using the `input_dims` and `output_dims` functions.
For $2^N$ by $2^M$ operators, the input and output dimension will be automatically assumed to be M-qubit and N-qubit:
```
op = Operator(np.random.rand(2 ** 1, 2 ** 2))
print('Input dimensions:', op.input_dims())
print('Output dimensions:', op.output_dims())
```
If the input matrix is not divisible into qubit subsystems, then it will be stored as a single-qubit operator. For example, if we have a $6\times6$ matrix:
```
op = Operator(np.random.rand(6, 6))
print('Input dimensions:', op.input_dims())
print('Output dimensions:', op.output_dims())
```
The input and output dimension can also be manually specified when initializing a new operator:
```
# Force input dimension to be (4,) rather than (2, 2)
op = Operator(np.random.rand(2 ** 1, 2 ** 2), input_dims=[4])
print('Input dimensions:', op.input_dims())
print('Output dimensions:', op.output_dims())
# Specify system is a qubit and qutrit
op = Operator(np.random.rand(6, 6),
input_dims=[2, 3], output_dims=[2, 3])
print('Input dimensions:', op.input_dims())
print('Output dimensions:', op.output_dims())
```
We can also extract just the input or output dimensions of a subset of subsystems using the `input_dims` and `output_dims` functions:
```
print('Dimension of input system 0:', op.input_dims([0]))
print('Dimension of input system 1:', op.input_dims([1]))
```
## Converting classes to Operators
Several other classes in Qiskit can be directly converted to an `Operator` object using the operator initialization method. For example:
* `Pauli` objects
* `Gate` and `Instruction` objects
* `QuantumCircuits` objects
Note that the last point means we can use the `Operator` class as a unitary simulator to compute the final unitary matrix for a quantum circuit, without having to call a simulator backend. If the circuit contains any unsupported operations, an exception will be raised. Unsupported operations are: measure, reset, conditional operations, or a gate that does not have a matrix definition or decomposition in terms of gate with matrix definitions.
```
# Create an Operator from a Pauli object
pauliXX = Pauli(label='XX')
Operator(pauliXX)
# Create an Operator for a Gate object
Operator(CnotGate())
# Create an operator from a parameterized Gate object
Operator(RXGate(np.pi / 2))
# Create an operator from a QuantumCircuit object
circ = QuantumCircuit(10)
circ.h(0)
for j in range(1, 10):
circ.cx(j-1, j)
# Convert circuit to an operator by implicit unitary simulation
Operator(circ)
```
## Using Operators in circuits
Unitary `Operators` can be directly inserted into a `QuantumCircuit` using the `QuantumCircuit.append` method. This converts the `Operator` into a `UnitaryGate` object, which is added to the circuit.
If the operator is not unitary, an exception will be raised. This can be checked using the `Operator.is_unitary()` function, which will return `True` if the operator is unitary and `False` otherwise.
```
# Create an operator
XX = Operator(Pauli(label='XX'))
# Check unitary
print('Operatator is unitary:', XX.is_unitary())
# Add to a circuit
circ = QuantumCircuit(2, 2)
circ.append(XX, [0, 1])
circ.measure([0,1], [0,1])
print(circ)
backend = BasicAer.get_backend('qasm_simulator')
job = execute(circ, backend, basis_gates=['u1','u2','u3','cx'])
job.result().get_counts(0)
```
Note that in the above example we initialize the operator from a `Pauli` object. However, the `Pauli` object may also be directly inserted into the circuit itself and will be converted into a sequence of single-qubit Pauli gates:
```
# Add to a circuit
circ2 = QuantumCircuit(2, 2)
circ2.append(Pauli(label='XX'), [0, 1])
circ2.measure([0,1], [0,1])
print(circ2)
# Simulate
job2 = execute(circ2, backend)
job2.result().get_counts(0)
```
## Combining Operators
Operators my be combined using several methods.
### Tensor Product
Two operators $A$ and $B$ may be combined into a tensor product operator $A\otimes B$ using the `Operator.tensor` function. Note that if both A and B are single-qubit operators, then `A.tensor(B)` = $A\otimes B$ will have the subsystems indexed as matrix B on subsystem 0, and matrix $A$ on subsystem 1.
```
A = Operator(Pauli(label='X'))
B = Operator(Pauli(label='Z'))
A.tensor(B)
```
### Tensor Expansion
A closely related operation is `Operator.expand`, which acts like a tensor product but in the reverse order. Hence, for two operators $A$ and $B$ we have `A.expand(B)` = $B\otimes A$ where the subsystems indexed as matrix A on subsystem 0, and matrix $B$ on subsystem 1.
```
A = Operator(Pauli(label='X'))
B = Operator(Pauli(label='Z'))
A.expand(B)
```
### Composition
We can also compose two operators $A$ and $B$ to implement matrix multiplication using the `Operator.compose` method. We have that `A.compose(B)` returns the operator with matrix $B.A$:
```
A = Operator(Pauli(label='X'))
B = Operator(Pauli(label='Z'))
A.compose(B)
```
We can also compose in the reverse order by applying $B$ in front of $A$ using the `front` kwarg of `compose`: `A.compose(B, front=True)` = $A.B$:
```
A = Operator(Pauli(label='X'))
B = Operator(Pauli(label='Z'))
A.compose(B, front=True)
```
### Subsystem Composition
Note that the previous compose requires that the total output dimension of the first operator $A$ is equal to total input dimension of the composed operator $B$ (and similarly, the output dimension of $B$ must be equal to the input dimension of $A$ when composing with `front=True`).
We can also compose a smaller operator with a selection of subsystems on a larger operator using the `qargs` kwarg of `compose`, either with or without `front=True`. In this case, the relevant input and output dimensions of the subsystems being composed must match. *Note that the smaller operator must always be the argument of `compose` method.*
For example, to compose a two-qubit gate with a three-qubit Operator:
```
# Compose XZ with an 3-qubit identity operator
op = Operator(np.eye(2 ** 3))
XZ = Operator(Pauli(label='XZ'))
op.compose(XZ, qargs=[0, 2])
# Compose YX in front of the previous operator
op = Operator(np.eye(2 ** 3))
YX = Operator(Pauli(label='YX'))
op.compose(XZ, qargs=[0, 2], front=True)
```
### Linear combinations
Operators may also be combined using standard linear operators for addition, subtraction and scalar multiplication by complex numbers.
```
XX = Operator(Pauli(label='XX'))
YY = Operator(Pauli(label='YY'))
ZZ = Operator(Pauli(label='ZZ'))
op = 0.5 * (XX + YY - 3 * ZZ)
op
```
An important point is that while `tensor`, `expand` and `compose` will preserve the unitarity of unitary operators, linear combinations will not; hence, adding two unitary operators will, in general, result in a non-unitary operator:
```
op.is_unitary()
```
### Implicit Conversion to Operators
Note that for all the following methods, if the second object is not already an `Operator` object, it will be implicitly converted into one by the method. This means that matrices can be passed in directly without being explicitly converted to an `Operator` first. If the conversion is not possible, an exception will be raised.
```
# Compose with a matrix passed as a list
Operator(np.eye(2)).compose([[0, 1], [1, 0]])
```
## Comparison of Operators
Operators implement an equality method that can be used to check if two operators are approximately equal.
```
Operator(Pauli(label='X')) == Operator(XGate())
```
Note that this checks that each matrix element of the operators is approximately equal; two unitaries that differ by a global phase will not be considered equal:
```
Operator(XGate()) == np.exp(1j * 0.5) * Operator(XGate())
```
### Process Fidelity
We may also compare operators using the `process_fidelity` function from the *Quantum Information* module. This is an information theoretic quantity for how close two quantum channels are to each other, and in the case of unitary operators it does not depend on global phase.
```
# Two operators which differ only by phase
op_a = Operator(XGate())
op_b = np.exp(1j * 0.5) * Operator(XGate())
# Compute process fidelity
F = process_fidelity(op_a, op_b)
print('Process fidelity =', F)
```
Note that process fidelity is generally only a valid measure of closeness if the input operators are unitary (or CP in the case of quantum channels), and an exception will be raised if the inputs are not CP.
```
import qiskit.tools.jupyter
%qiskit_version_table
%qiskit_copyright
```
| github_jupyter |
```
library(caret, quiet=TRUE);
library(base64enc)
library(httr, quiet=TRUE)
```
# Build a Model
```
set.seed(1960)
create_model = function() {
model <- train(Species ~ ., data = iris, method = "rpart" , preProcess = c("invHyperbolicSine"))
return(model)
}
# dataset
model = create_model()
pred <- predict(model, as.matrix(iris[, -5]) , type="prob")
pred_labels <- predict(model, as.matrix(iris[, -5]) , type="raw")
sum(pred_labels != iris$Species)/length(pred_labels)
```
# SQL Code Generation
```
test_ws_sql_gen = function(mod) {
WS_URL = "https://sklearn2sql.herokuapp.com/model"
WS_URL = "http://localhost:1888/model"
model_serialized <- serialize(mod, NULL)
b64_data = base64encode(model_serialized)
data = list(Name = "caret_rpart_test_model", SerializedModel = b64_data , SQLDialect = "postgresql" , Mode="caret")
r = POST(WS_URL, body = data, encode = "json")
# print(r)
content = content(r)
# print(content)
lSQL = content$model$SQLGenrationResult[[1]]$SQL # content["model"]["SQLGenrationResult"][0]["SQL"]
return(lSQL);
}
lModelSQL = test_ws_sql_gen(model)
cat(lModelSQL)
```
# Execute the SQL Code
```
library(RODBC)
conn = odbcConnect("pgsql", uid="db", pwd="db", case="nochange")
odbcSetAutoCommit(conn , autoCommit = TRUE)
dataset = iris[,-5]
df_sql = as.data.frame(dataset)
names(df_sql) = sprintf("Feature_%d",0:(ncol(df_sql)-1))
df_sql$KEY = seq.int(nrow(dataset))
sqlDrop(conn , "INPUT_DATA" , errors = FALSE)
sqlSave(conn, df_sql, tablename = "INPUT_DATA", verbose = FALSE)
head(df_sql)
# colnames(df_sql)
# odbcGetInfo(conn)
# sqlTables(conn)
df_sql_out = sqlQuery(conn, lModelSQL)
head(df_sql_out)
```
# R Caret Rpart Output
```
pred_proba = predict(model, as.matrix(iris[,-5]), type = "prob")
df_r_out = data.frame(pred_proba)
names(df_r_out) = sprintf("Proba_%s",model$levels)
df_r_out$KEY = seq.int(nrow(dataset))
df_r_out$Score_setosa = NA
df_r_out$Score_versicolor = NA
df_r_out$Score_virginica = NA
df_r_out$LogProba_setosa = log(df_r_out$Proba_setosa)
df_r_out$LogProba_versicolor = log(df_r_out$Proba_versicolor)
df_r_out$LogProba_virginica = log(df_r_out$Proba_virginica)
df_r_out$Decision = predict(model, as.matrix(iris[,-5]), type = "raw")
df_r_out$DecisionProba = apply(pred_proba, 1, function(x) max(x))
head(df_r_out)
```
# Compare R and SQL output
```
df_merge = merge(x = df_r_out, y = df_sql_out, by = "KEY", all = TRUE, , suffixes = c("_R","_SQL"))
head(df_merge)
diffs_df = df_merge[df_merge$Decision_1 != df_merge$Decision_2,]
head(diffs_df)
stopifnot(nrow(diffs_df) == 0)
summary(df_sql_out)
summary(df_r_out)
prep = model$preProcess
prep
```
| github_jupyter |
# Deep Learning for Unstructured Data: Application to Image Search
#### Aivin V. Solatorio
e-mail: avsolatorio@gmail.com       github: <a href='https://github.com/avsolatorio/'>avsolatorio</a>       linkedin: <a href='https://linkedin.com/in/avsolatorio/'>avsolatorio</a>
# Configuration and loading of necessary modules
```
%matplotlib inline
import warnings
warnings.filterwarnings("ignore")
import os
os.environ["CUDA_VISIBLE_DEVICES"]="0"
from keras.utils import multi_gpu_model
import tensorflow as tf
from keras.backend.tensorflow_backend import set_session
config = tf.ConfigProto()
config.gpu_options.allow_growth = True
# config.gpu_options.per_process_gpu_memory_fraction = 0.9
set_session(tf.Session(config=config))
print tf.__version__
from PIL import Image
import pandas as pd
import pylab as plt
import numpy as np
import keras
import re
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
from keras.preprocessing.image import ImageDataGenerator
from IPython.display import SVG
from keras.utils.vis_utils import model_to_dot
from sklearn.metrics.pairwise import cosine_similarity
print keras.__version__
```
# Load dataset scraped from google images
- The dataset consists of google images of around 50 animals.
- Each animal set contains roughly 500 images.
- Data captured are the image, title, and caption for the image.

```
full_data_df = pd.read_hdf('../../data/full_data_common_animals.hdf', 'full_data_df')
full_data_df.head()
```
# Text data processing
```
sample_data = full_data_df[full_data_df.columns.drop(['image_link', 'image_type', 'record_id'])].loc[full_data_df.index]
sample_data = sample_data.drop_duplicates(['image'])
```
We apply a very simple tokenization procedure: Just use alphanumeric characters in the text data, i.e., captions and titles.
```
def process_raw_text(x, tokenize=True, filter_symbols=True):
x = x.lower()
x = x.replace('.jpg', '')
x = x.replace('.png', '')
if tokenize:
x = re.findall("[a-z0-9]+", x)
else:
x = re.sub("[^a-z0-9\ ]+", ' ', x)
return x
sample_data['title_tokens'] = sample_data['title'].map(process_raw_text)
sample_data['caption_tokens'] = sample_data['caption'].map(process_raw_text)
```
Let's apply Jaccard measure to know if the title is almost the same as the caption. We need to do this to avoid redundancy in the text data.
```
def jaccard_similarity(row):
intersection = len(set(row['title_tokens']).intersection(row['caption_tokens']))
union = len(set(row['title_tokens']).union(row['caption_tokens']))
similarity = 1.0 * intersection / union
return similarity
sample_data['jaccard_sim'] = sample_data.apply(jaccard_similarity, axis=1)
```
By using the similarity score, we select which text data are used. If the title and the caption are dissimilar based on a threshold, then we use both text (concatenated) as input.
```
def get_effective_sample_text(row, sim_thresh=0.5):
token_set = []
title_tokens = row['title_tokens']
caption_tokens = row['caption_tokens']
if row['jaccard_sim'] > sim_thresh:
if len(title_tokens) > len(caption_tokens):
token_set.extend(title_tokens)
else:
token_set.extend(caption_tokens)
else:
token_set.extend(title_tokens)
token_set.extend(caption_tokens)
return token_set
sample_data['tokens'] = sample_data.apply(get_effective_sample_text, axis=1)
sample_data = sample_data.drop(['caption', 'title', 'title_tokens', 'caption_tokens', 'jaccard_sim'], axis=1)
```
After taking the relevant text data, we retokenize them with a `Tokenizer` utility class from the keras preprocessing module.
Note that the `tokens` data is already a list of words, so we need to join each token together before fitting the tokenizer.
```
tokenizer = Tokenizer(num_words=10000, filters='', lower=True, split=' ', char_level=False, oov_token=None)
tokenizer.fit_on_texts(sample_data['tokens'].map(lambda x: ' '.join(x)))
```
We take the histogram of token length for each item in our dataset. With this, we can have an informed choice for our expected sequence length when standardizing the length of the text input.
```
pd.Series([len(i) for i in tokenizer.texts_to_sequences(
sample_data['tokens'].map(lambda x: ' '.join(x))
)]).hist(bins=range(0, 50, 5))
MAXLEN = 15
EMBEDDING_DIM = 300
```
In the snippet below, we perform standardization of the text data. The process involves truncation of text sequences longer than our specified `MAXLEN` as well as padding default values to sequences that are shorter than `MAXLEN`. This is required for convenience later in the training procedure.
```
sample_data['sequence'] = pad_sequences(
tokenizer.texts_to_sequences(
sample_data['tokens'].map(lambda x: ' '.join(x))
), maxlen=MAXLEN, padding='pre', truncating='post'
).tolist()
sample_data['empty_sequence'] = sample_data.sequence.map(lambda x: sum(x) == 0)
sample_data = sample_data[~sample_data.empty_sequence]
sample_data.shape
```
# How can we use the words in the text as input to our model?
## What are word vectors?
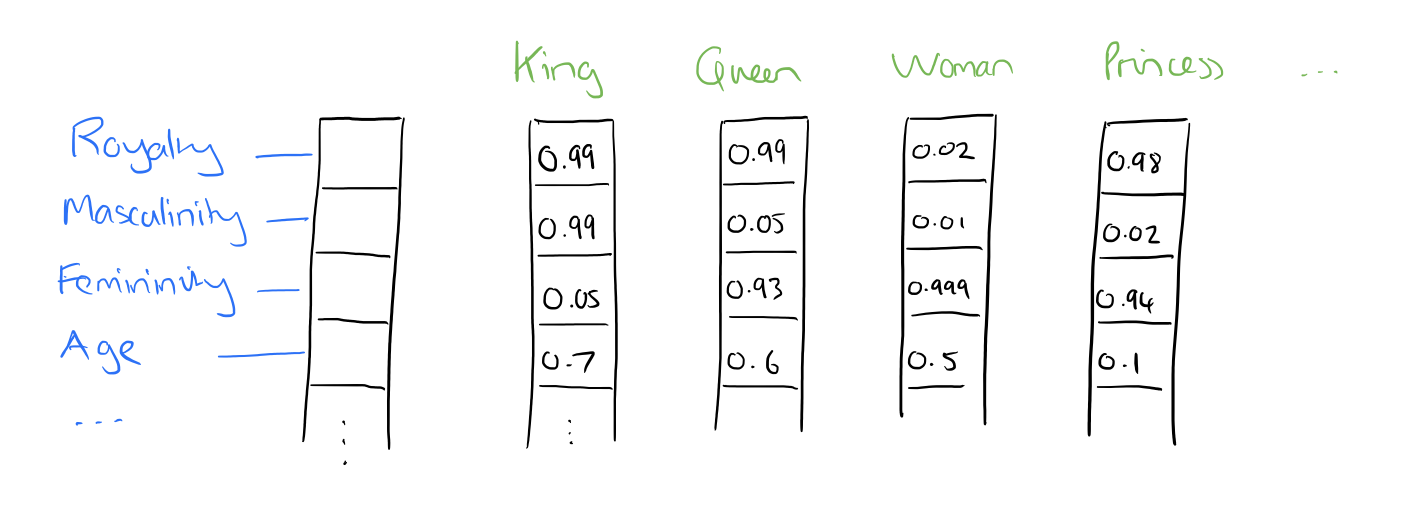
## Special properties of word vectors

## Learning a word vector representation
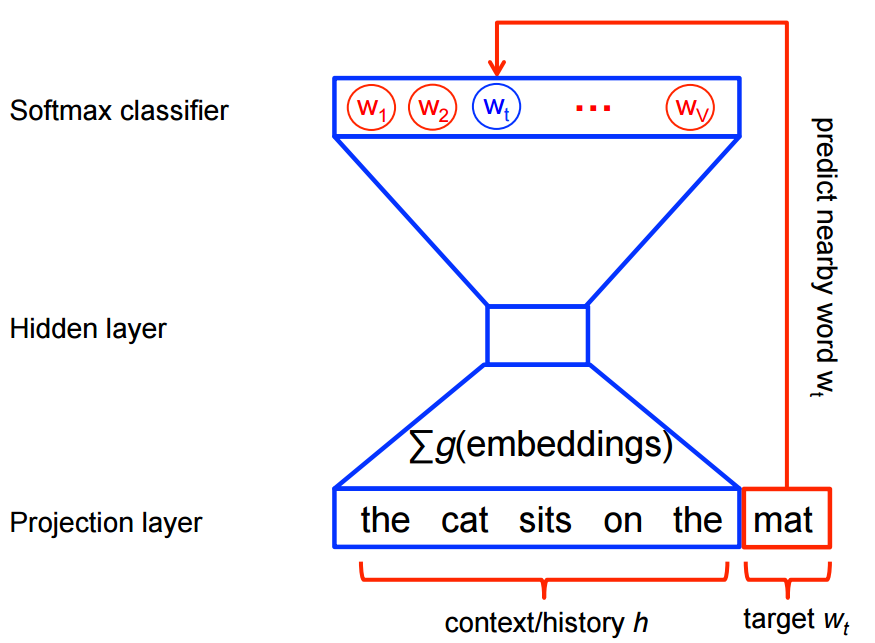
# Using pretrained word vectors
We can use the word vectors trained with large dataset as prior word vectors for our model. Google (word2vec), Facebook (fasttext), and Stanford (glove) have released some pretrained word vectors to the public domain.
```
%%time
def get_embedding_index(pre_trained_file, word_index):
# Expected format is space separated.
# First item is the word and succedding items are the elements of the vector
embedding_index = {}
with open(pre_trained_file) as fl:
for line in fl:
line = line.strip().split()
word = line[0]
vector = line[1:]
if word in word_index:
embedding_index[word] = np.array(vector, dtype=np.float32)
return embedding_index
embeddings_index = get_embedding_index(
pre_trained_file='/mnt/Datastore/WORK/pre-trained-models/glove.840B.300d.txt',
word_index=tokenizer.word_index
)
```
Let's build an embedding matrix containing each word vector for our vocabulary. Words not found in the pretrained model are set to random normal vectors. A default vector for padding is set to the zero vector.
```
%%time
# Keras reserves index 0 for masking
word_index = tokenizer.word_index
print('Found %s word vectors.' % len(embeddings_index))
embedding_matrix = np.random.randn(len(word_index) + 1, EMBEDDING_DIM)
embedding_matrix[0] = np.zeros(EMBEDDING_DIM)
for word, i in word_index.items():
embedding_vector = embeddings_index.get(word)
if embedding_vector is not None:
# words not found in embedding index will be all-zeros.
embedding_matrix[i] = embedding_vector
```
# Image data processing
```
from keras.layers import (
Input, GRU, Bidirectional, Dense, Conv2D, MaxPooling2D,
GlobalAveragePooling2D, Flatten, Dropout, LeakyReLU
)
from keras.applications import VGG16
```
Let's use a pretrained model as well for images to condition our image model. We chose VGG16 model without any special reason (other pretrained models can be used).
```
image_size = (150, 150, 3)
conv_base = VGG16(weights='imagenet',
include_top=False,
input_shape=image_size)
# Taken from sklearn and modified to add a `size` attribute
def imread(name, flatten=False, mode=None, size=None):
"""
Read an image from a file as an array.
Parameters
----------
name : str or file object
The file name or file object to be read.
flatten : bool, optional
If True, flattens the color layers into a single gray-scale layer.
mode : str, optional
Mode to convert image to, e.g. ``'RGB'``. See the Notes for more
details.
Returns
-------
imread : ndarray
The array obtained by reading the image.
Notes
-----
`imread` uses the Python Imaging Library (PIL) to read an image.
The following notes are from the PIL documentation.
`mode` can be one of the following strings:
* 'L' (8-bit pixels, black and white)
* 'P' (8-bit pixels, mapped to any other mode using a color palette)
* 'RGB' (3x8-bit pixels, true color)
* 'RGBA' (4x8-bit pixels, true color with transparency mask)
* 'CMYK' (4x8-bit pixels, color separation)
* 'YCbCr' (3x8-bit pixels, color video format)
* 'I' (32-bit signed integer pixels)
* 'F' (32-bit floating point pixels)
PIL also provides limited support for a few special modes, including
'LA' ('L' with alpha), 'RGBX' (true color with padding) and 'RGBa'
(true color with premultiplied alpha).
When translating a color image to black and white (mode 'L', 'I' or
'F'), the library uses the ITU-R 601-2 luma transform::
L = R * 299/1000 + G * 587/1000 + B * 114/1000
When `flatten` is True, the image is converted using mode 'F'.
When `mode` is not None and `flatten` is True, the image is first
converted according to `mode`, and the result is then flattened using
mode 'F'.
"""
im = Image.open(name)
if size is not None:
im = im.resize(size)
return fromimage(im, flatten=flatten, mode=mode)
def fromimage(im, flatten=False, mode=None):
"""
Return a copy of a PIL image as a numpy array.
Parameters
----------
im : PIL image
Input image.
flatten : bool
If true, convert the output to grey-scale.
mode : str, optional
Mode to convert image to, e.g. ``'RGB'``. See the Notes of the
`imread` docstring for more details.
Returns
-------
fromimage : ndarray
The different colour bands/channels are stored in the
third dimension, such that a grey-image is MxN, an
RGB-image MxNx3 and an RGBA-image MxNx4.
"""
if not Image.isImageType(im):
raise TypeError("Input is not a PIL image.")
if mode is not None:
if mode != im.mode:
im = im.convert(mode)
elif im.mode == 'P':
# Mode 'P' means there is an indexed "palette". If we leave the mode
# as 'P', then when we do `a = array(im)` below, `a` will be a 2-D
# containing the indices into the palette, and not a 3-D array
# containing the RGB or RGBA values.
if 'transparency' in im.info:
im = im.convert('RGBA')
else:
im = im.convert('RGB')
if flatten:
im = im.convert('F')
elif im.mode == '1':
# Workaround for crash in PIL. When im is 1-bit, the call array(im)
# can cause a seg. fault, or generate garbage. See
# https://github.com/scipy/scipy/issues/2138 and
# https://github.com/python-pillow/Pillow/issues/350.
#
# This converts im from a 1-bit image to an 8-bit image.
im = im.convert('L')
a = np.array(im)
return a
def recast_array_to_image(float_array):
uint_array = (float_array * 255 / np.max(float_array)).astype('uint8')
return Image.fromarray(uint_array)
image_cache = {}
def load_image(path, size=image_size[:-1], as_float=True):
if path not in image_cache:
im = imread(path, size=size)
if as_float:
im = im.astype(np.float32)
image_cache[path] = im
return image_cache[path]
```
We perform preprocessing of the images by standardizing their sizes and scaling the values from 0-255 to 0-1.
```
%%time
image_datagen = ImageDataGenerator(
rescale=1./255
)
sample_data['image_array'] = conv_base.predict(
image_datagen.flow(
np.stack(sample_data['image'].map(load_image)),
batch_size=sample_data.shape[0], shuffle=False
).next()
).tolist()
sample_data.head()
```
## Implementing a data generator for training
```
def pretrain_model_image_generator(image_data, batch_size):
image_datagen = ImageDataGenerator(
rescale=1./255
)
# Exploit image_datagen for dynamic perturbation of the data
image_data = image_datagen.flow(
image_data,
batch_size=len(image_data), shuffle=False
).next()
image_data = conv_base.predict(image_data)
for i in range(0, len(image_data) // batch_size):
yield image_data[i * batch_size: (i + 1) * batch_size]
def dataset_generator(dataset, batch_size=32, shuffle=True, negative_fold=1):
index = np.array(dataset.index.values) # Interestingly, data gets shuffled if we don't use np.array to create a new object!
full_batch_size = batch_size * (negative_fold + 1)
batch_index = []
while True:
if shuffle:
np.random.shuffle(index)
for i in index:
if len(batch_index) < batch_size:
batch_index.append(i)
else:
positive_df = dataset.loc[batch_index]
batch_image_data = np.stack(positive_df['image_array'])
positive_text_data = np.stack(positive_df['sequence'])
if negative_fold:
batch_image_data = np.vstack([batch_image_data] * (negative_fold + 1))
negative_index = np.random.choice(index, size=batch_size * negative_fold, replace=True)
negative_text_data = np.stack(dataset.loc[negative_index]['sequence'])
text_data = np.vstack([positive_text_data, negative_text_data])
else:
text_data = positive_text_data
label = np.zeros(full_batch_size)
label[:batch_size] = 1
ix = np.array(range(full_batch_size))
if shuffle:
ix = np.random.choice(ix, size=full_batch_size, replace=False)
batch_index = []
yield [text_data[ix], batch_image_data[ix]], label[ix]
# yield [text_data[ix], conv_base.predict(batch_image_data[ix])], label[ix]
%%time
data_gen = dataset_generator(sample_data, batch_size=32, shuffle=True, negative_fold=2)
q = data_gen.next()
```
# Building the model
The model consists of two parts
Text model
Image model
Our objective is to train these two models jointly such that they will share the same vector space.
## Text model
```
text_model_input = keras.layers.Input(shape=(MAXLEN,))
text_embedding = keras.layers.Embedding(
input_dim=len(word_index) + 1, output_dim=EMBEDDING_DIM,
weights=[embedding_matrix],
trainable=False,
mask_zero=True
)(text_model_input)
text_gru = Bidirectional(GRU(64, return_sequences=False))(text_embedding)
text_dense = Dense(300, activation='relu')(text_gru)
text_dense = Dropout(0.3)(text_dense)
text_model_output = Dense(200, activation='tanh')(text_dense)
TextModel = keras.models.Model(inputs=[text_model_input], outputs=[text_model_output])
# SVG(model_to_dot(TextModel).create(prog='dot', format='svg'))
```
## Image model
# Convolutional Network
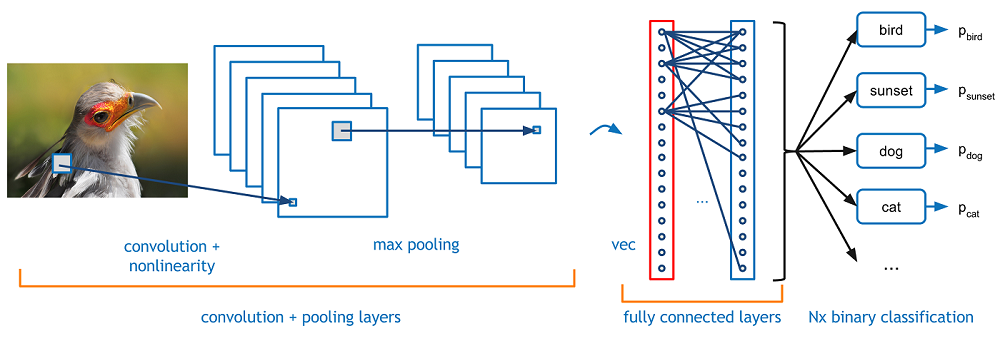
```
image_model_input = Input(shape=(4, 4, 512))
image_conv = Conv2D(64, 2)(image_model_input)
image_pool = MaxPooling2D(pool_size=(2, 2))(image_conv)
image_flatten = Flatten()(image_pool)
image_input_pool = MaxPooling2D(pool_size=(2, 2))(image_model_input)
image_input_flatten = Flatten()(image_input_pool)
full_image = keras.layers.concatenate([image_input_flatten, image_flatten])
image_model_output = Dense(200, activation='tanh')(full_image) # full_image_dense)
ImageModel = keras.models.Model(inputs=[image_model_input], outputs=[image_model_output])
# SVG(model_to_dot(ImageModel).create(prog='dot', format='svg'))
```
## Image Search model
```
image_search_ouput = keras.layers.merge([text_model_output, image_model_output], mode='cos')
image_search_ouput = keras.layers.Reshape((1,), input_shape=(1, 1))(image_search_ouput)
ImageSearchModel = keras.models.Model(inputs=[text_model_input, image_model_input], outputs=[image_search_ouput])
ImageSearchModel.compile(
optimizer='adam',
loss='binary_crossentropy'
)
# SVG(model_to_dot(ImageSearchModel).create(prog='dot', format='svg'))
ImageSearchModel.summary()
```
# Training the model
```
%%time
ImageSearchModel.fit_generator(data_gen, steps_per_epoch=full_data_df.shape[0] // 32, epochs=10)
%%time
ImageSearchModel.fit_generator(data_gen, steps_per_epoch=full_data_df.shape[0] // 32, epochs=100)
%%time
ImageSearchModel.fit_generator(data_gen, steps_per_epoch=full_data_df.shape[0] // 32, epochs=10)
```
# Transforming images to embedding vectors
```
%%time
image_embeddings = ImageModel.predict(np.stack(sample_data['image_array'].values))
image_embeddings.shape
```
# Making a text query
We process the query using the pipeline used in transforming our training data.
```
text_query = "the pool with dogs and cats"
seq = pad_sequences(
tokenizer.texts_to_sequences(
[' '.join(process_raw_text(text_query))]
), maxlen=MAXLEN, padding='pre', truncating='post'
)
query_vector = TextModel.predict(seq)
seq
```
We then compute the `cosine similarity` of the query vector against the image embedding vectors.
```
%%time
res = cosine_similarity(image_embeddings, query_vector)
top_ix = res[:, 0].argsort()[::-1][:10]
print(res[:, 0][top_ix])
print(top_ix)
sample_data.iloc[top_ix].head()
```
Retrieved images
```
plt.figure(figsize=(15, 5))
for ix, iloc in enumerate(top_ix[:5]):
im = Image.fromarray(
load_image(
sample_data.iloc[iloc]['image'],
size=None,
).astype(np.uint8)
)
plt.subplot(int('15{}'.format(ix + 1)))
plt.imshow(np.asarray(im))
plt.axis('off')
```
# Query by image
```
iloc = 30194
# iloc = 20194
im = Image.fromarray(
load_image(
sample_data.iloc[iloc]['image'],
size=None,
).astype(np.uint8)
)
im
image_query = ImageModel.predict(np.stack(sample_data.iloc[[iloc]].image_array.values))
res_im = cosine_similarity(image_embeddings, image_query)
top_im_ix = res_im[:, 0].argsort()[::-1][:10]
print(res_im[:, 0][top_im_ix])
print(top_im_ix)
sample_data.iloc[top_im_ix].head()
plt.figure(figsize=(15, 5))
for ix, iloc in enumerate(top_im_ix[:5]):
im = Image.fromarray(
load_image(
sample_data.iloc[iloc]['image'],
size=None,
).astype(np.uint8)
)
plt.subplot(int('15{}'.format(ix + 1)))
plt.imshow(np.asarray(im))
plt.axis('off')
from notebook.services.config import ConfigManager
cm = ConfigManager()
cm.update('livereveal', {
# 'width': 1024,
# 'height': 768,
'scroll': True,
})
```
| github_jupyter |
# Elementi di Programmazione
Un linguaggio di programmazione serve sia per istruire una macchina ad eseguire dei conti, che per organizzare le nostre idee su come quei conti devono essere eseguiti. Per questo, nella scelta di un linguaggio di programmazione, dobbiamo tener presente quali sono gli strumenti che vengono offerti dal linguaggio per formare idee più complesse partendo da poche idee semplici.
Ogni linguaggio di programmazione dovrebbe avere almeno tre caratteristiche per ottenere questo obiettivo:
1. Delle **espressioni primitive** che rappresentano le entità più semplici del linguaggio
2. Dei metodi per **combinare** gli elementi primitivi in elementi composti
3. Dei metodi per **astrarre** concetti primitivi in modo che gli elementi composti possano essere utilizzati a loro volta come elementi primitivi di entità ancora più complesse
In programmazione abbiamo due tipi di elementi: le PROCEDURE e i DATI.
In modo informale possiamo definire i **dati** come gli oggetti che vorremmo manipolare, e le **procedure** come la descrizione delle regole per manipolare i dati. Quindi, per quanto spiegato sopra, un linguaggio dovrebbe avere dei **dati primitivi** e delle **procedure primitive**, e dovrebbe avere dei metodi per combinare e astrarre i dati e le procedure.
## Dati e procedure numeriche
Iniziamo a vedere qualche semplice iterazione con l'interprete di Python: se digitiamo sulla tastiera una **espressione**, l'interprete risponde **valutando** tale espressione. Per esempio se digitiamo il numero 345 e poi premiamo la combinazione di tasti `shift + enter`, l'interprete valuta l'espressione che abbiamo appena scritto:
```
345
```
Semplici espressioni numeriche possono essere combinate usando delle procedure primitive che rappresentano l'applicazione di procedure a quei numeri. Per esempio:
```
339 + 6
345 - 6
2.7 / 12.1
345 - 12/6
```
Si noti come in questo caso, per queste semplici procedure numeriche che corrispondono agli operatori aritmetici, viene implicitamente usata una notazione chiamata **postfix**. Importando la libreria `operator` è possibile esprimere le stesse espressioni in notazione **prefix**:
```
# Importa tutte le procedure (funzioni) definite nel modulo "operator"
from operator import *
```
Si consiglia di leggere la **documentazione** della libreria `operator` sul sito di [python](https://docs.python.org/3.6/library/operator.html). Le funzioni principali che useremo in questo notebook sono:
* `add(a,b)` corrisponde ad `a+b`
* `sub(a,b)` corrisponde ad `a-b`
* `mul(a,b)` corrisponde ad `a*b`
* `truediv(a,b)` corrisponde ad `a/b`
Per esempio:
```
add(339, 6)
```
Uno dei vantaggi della notazione **prefix** è che rende sempre chiaro qual è l'operatore/procedura che deve essere svolta, applicandola a quali dati: `add` è il nome dell'operatore, mentre tra parentesi sono definiti i due dati numerici a cui deve essere applicata l'operazione.
```
sub(345, truediv(12, 6))
mul(add(2,3), (sub(add(2,2), add(3,2))))
```
Si noti come l'espressione precedente sarebbe più chiara se scritta come:
```
mul(
add(2, 3),
sub(
add(2, 2),
add(3, 2)
)
)
```
In questo caso l'interprete lavora in un ciclo chiamato "leggi-valuta-stampa": legge le espressioni composte e le espressioni primitive, le valuta nell'ordine in cui le trova, e stampa alla fine il risultato finale.
## Assegnazione di nomi ad oggetti
Un aspetto critico della programmazione è il modo in cui vengono assegnati i nomi agli oggetti computazionali.
Si dice che un **nome** identifica una **variabile** il cui valore è l'oggetto a cui viene associato. Per esempio:
```
a = 13
```
In questo caso abbiamo una variabile, che abbiamo chiamato `a`, e il cui **valore** è il numero 13. A questo punto possiamo usare la variabile `a` come un oggetto di tipo numerico:
```
3*a
add(a, add(a,a))
pi = 3.14159
raggio = 5
circonferenza = 2*pi*raggio
circonferenza
raggio = 10
circonferenza
```
In questo caso, l'interprete del linguaggio, ha prima valutato l'espressione `2*pi*raggio`, e dopo ha assegnato il **valore** ottenuto dalla valutazione dell'**espressione** alla variabile di nome `circonferenza`.
In questo caso, l'operatore di assegnamento `=` rappresenta il più semplice meccanismo di astrazione, perché permette di dare un **nome** al risultato di operazioni più complesse.
In pratica, qualsiasi programma viene costruito partendo dalla costruzione, passo passo, di oggetti computazionali via via più complessi.
L'uso di un interprete, che in modo incrementale valute le **espressioni** che li vengono passate, favorisce la definizione di tante piccole procedure, innestate l'uno nell'altra.
Dovrebbe essere chiaro a questo punto, che l'interprete deve mantenere una sorta di **MEMORIA** che tiene traccia di tutti gli assegnamenti di nomi a oggetti, chiamato il `global environment`. Per vedere quali sono i nomi memorizzati in memoria si usa il comando `who`:
```
who
```
## La valutazione di espressioni composte
Uno degli obiettivo di questo corso è di insegnare a pensare in maniera "*algoritmica*". Proviamo ad analizzare come l'interprete del linguaggio valuta operazioni composte come quelle viste prima. In pratica, la valutazione di operazioni composte avviene attraverso la procedura seguente:
1. Per valutare un'espressione composta:
1.1 Prima, valuta le sottoespressioni della espressioni composta
1.2 Applica la procedura indicata dalla sottoespressioni più a sinistra (l'operatore), agli argomenti che sono i valori della sottoespressione (gli operandi).
Si noti come questa procedura per valutare un'operazione composta, per prima cosa deve eseguire il processo di valutazione a ogni elemento dell'espressione composta. Quindi la regola di valutazione di un'espressione è intrinsecamente **RICORSIVA**, ovvero include come uno dei suoi passi la chiamata a se stessa.
NOTA: mostrare alla lavagna un *recursion tree* con l'espressione precedente.
## Definizione di procedure composte
Abbiamo identificato alcuni elementi che devono appartenere ad un linguaggio di programmazione:
1. I numeri e le operazioni aritmetiche sono **dati** e **procedure** primitive (in gergo, vengono chiamate *builtin*)
2. L'**annidamento di espressioni** composte offrono un meccanismo per comporre delle operazioni
3. Gli **assegnamenti** di nomi di variabili a valori offrono un livello di astrazione piuttosto limitato
Abbiamo quindi bisogno di un modo per poter **definire nuove procedure**, in modo che una nuova operazione possa essere definita in termini di composizione di operazioni più semplici.
Consideriamo per esempio una procedura di elevamento al quadrato.
```
quadrato = mul(3, 3)
quadrato
power = mul(x,x)
print(power)
x = 7
```
Per ottenere un livello di astrazione più alto abbiamo bisogno di un meccanismo (una sintassi del linguaggio) per definire nuove procedure (funzioni). La sintassi è la seguente:
```
def <Nome>(<parametri formali>):
<corpo della procedura>
```
Si noti come sia apparsa la prima *parole chiave riservata* del linguaggio: **def**. Inoltre, `<Nome>` è il nome che noi vogliamo associare alla procedura (funzione) che stiamo definendo, e i `<parametri formali>` (chiamati argomenti della procedura) sono le variabili che non appartengono direttamente al *working eniviroment* (ovvero alla MEMORIA dell'interprete), ma sono "visibili" solo internamente alla procedura in cui sono definite.
Se torniamo all'esempio delle definizione di una procedura per l'elevamento al quadrato, possiamo scrivere:
```
def Quadrato(numero):
return mul(numero, numero)
Quadrato
who
Quadrato(532)
quadrato(mul(3,2))
```
A questo punto possiamo anche definire nuove procedure in termini della procedura appena definita, definendo per esempio una nuova procedura chiamata `SommaDiQuadrati`:
```
def SommaQuadrati(x, y):
return add(Quadrato(x), Quadrato(y))
SommaQuadrati(4,3)
x
del x
```
**ESEMPIO**: considera la seguente espressione composta:
```
def F(a):
return SommaQuadrati(add(a, 1), mul(a, 2))
F(5)
```
**DOMANDA**: come è stata valutata questa procedura?
**DOMANDA**: con gli elementi del linguaggio sin qui introdotti, possiamo definire una funzione `ValoreAssoluto(x)`, ovvero
$|x| = x$ se $x \geq 0$, e $|x|=-x$ se $x < 0$?
**RISPOSTA**: No, il linguaggio non è abbastanza *espressivo*.
| github_jupyter |
# Expand With Interpolators
This notebook demonstrates the different interpolators available in SimpleITK available for image resampling. Their effect is demonstrated on the <a href="http://www.cs.cornell.edu/~srm/publications/Vis94-filters-abstract.html">Marschner-Lobb</a> image.
```
import matplotlib.pyplot as plt
%matplotlib inline
import SimpleITK as sitk
import numpy as np
import math
def myshow(img, title=None, margin=0.05):
if (img.GetDimension() == 3):
img = sitk.Tile( (img[img.GetSize()[0]//2,:,:],
img[:,img.GetSize()[1]//2,:],
img[:,:,img.GetSize()[2]//2]), [2,2])
aimg = sitk.GetArrayViewFromImage(img)
xsize,ysize = aimg.shape
dpi=80
# Make a figure big enough to accommodate an axis of xpixels by ypixels
# as well as the ticklabels, etc...
figsize = (1+margin)*ysize / dpi, (1+margin)*xsize / dpi
fig = plt.figure(figsize=figsize, dpi=dpi)
# Make the axis the right size...
ax = fig.add_axes([margin, margin, 1 - 2*margin, 1 - 2*margin])
t = ax.imshow(aimg)
if len(aimg.shape) == 2:
t.set_cmap("gray")
if(title):
plt.title(title)
plt.show()
def marschner_lobb(size=40, alpha=0.25, f_M=6.0):
img = sitk.PhysicalPointSource(sitk.sitkVectorFloat32, [size]*3, [-1]*3, [2.0/size]*3)
imgx = sitk.VectorIndexSelectionCast(img, 0)
imgy = sitk.VectorIndexSelectionCast(img, 1)
imgz = sitk.VectorIndexSelectionCast(img, 2)
del img
r = sitk.Sqrt(imgx**2 + imgy**2)
del imgx, imgy
pr = sitk.Cos((2.0*math.pi*f_M)*sitk.Cos((math.pi/2.0)*r))
return (1.0 - sitk.Sin((math.pi/2.0)*imgz) + alpha*(1.0+pr))/(2.0*(1.0+alpha))
myshow(marschner_lobb())
myshow(marschner_lobb(100))
ml = marschner_lobb()
ml = ml[:,:,ml.GetSize()[-1]//2]
myshow(sitk.Expand(ml, [5]*3, sitk.sitkNearestNeighbor), title="nearest neighbor")
myshow(sitk.Expand(ml, [5]*3, sitk.sitkLinear), title="linear")
myshow(sitk.Expand(ml, [5]*3, sitk.sitkBSpline), title="b-spline")
myshow(sitk.Expand(ml, [5]*3, sitk.sitkGaussian), title="Gaussian")
myshow(sitk.Expand(ml, [5]*3, sitk.sitkHammingWindowedSinc), title="Hamming windowed sinc")
myshow(sitk.Expand(ml, [5]*3, sitk.sitkBlackmanWindowedSinc), title="Blackman windowed sinc")
myshow(sitk.Expand(ml, [5]*3, sitk.sitkCosineWindowedSinc), title="Cosine windowed sinc")
myshow(sitk.Expand(ml, [5]*3, sitk.sitkWelchWindowedSinc), title="Welch windowed sinc")
myshow(sitk.Expand(ml, [5]*3, sitk.sitkLanczosWindowedSinc), title="Lanczos windowed sinc")
```
| github_jupyter |
# Generates the readout's coefficients using the output from the simulations created in the notebook 2DofArm_simulation-Main.ipnb
```
# Makes possible to show the output from matplotlib inline
%matplotlib inline
import matplotlib.pyplot as plt
# Makes the figures in the PNG format:
# For more information see %config InlineBackend
%config InlineBackend.figure_formats=set([u'png'])
plt.rcParams['figure.figsize'] = 5, 10
import numpy
import sys
from sklearn import linear_model
import save_load_file as slf
import membrane_lowpass_md
reload(sys.modules['membrane_lowpass_md']) # Makes sure the interpreter is going to reload the module
membrane_lowpass = membrane_lowpass_md.membrane_lowpass
STP_ON="" #means "NO STP"
# STP_ON="_STP_ON" #means "USING STP"
membrane_time_constant = 30E-3
using_spikes = 0 #controls if the output is based only on the spikes (1) or the membrane (0)
NofN = 600 # total number of neurons in the output
sim_step_time = 2E-3 #simulation step time (in seconds)
# Experiment identifier
sim_sets = ["set_A", "set_B", "set_C", "set_D"]
sim_set = sim_sets[0]
base_dir = "2DofArm_simulation_data"
total_trials = 20
number_of_trajectories = 4
# Reads the files with the input and output from the 4 trajectories
# Each trajectory was simulated 20 times using noise (at the input layer weights) to create slightly different inputs effects
output_spikes_simulation = []
for pos_i in xrange(1,number_of_trajectories+1):
temp_v = []
for run_i in range(1,total_trials+1):
simulated_values = slf.load_from_file_gz("./"+base_dir+"/"+sim_set+"/States_Torques_movement"+str(pos_i)+"_LSM_"+str(run_i)+STP_ON+".gzpickle")
# The format of simulated_values is (a list of tuples):
# [(
# current step, =>index 0
# current time (in ms), =>index 1
# numpy.array with input spikes/times, =>index 2
# list with the indices of the output spikes, =>index 3
# time the output spikes occured, =>index 4
# ), (...), ...]
# temp_v.append(numpy.array([\
# [simulated_values[i][3],\
# simulated_values[i][4]]\
# for i in range(len(simulated_values))],dtype=numpy.object))
temp_v.append([[simulated_values[i][3], simulated_values[i][4]] for i in range(len(simulated_values))])
output_spikes_simulation.append(temp_v)
del temp_v
# output_spikes_simulation[X] => all the simulated values for the trajectory X+1
# output_spikes_simulation[X][Y] => the simulated values for trial Y+1 of the trajectory X+1
# output_spikes_simulation[X][Y][Z] => the output spikes for the step Z of the simulated values for trial Y+1 of the trajectory X+1
# total_trajectories,total_trials,total_steps,temp = len(output_spikes_simulation),len(output_spikes_simulation[0]),len(output_spikes_simulation[0][0]),output_spikes_simulation[0][0].shape
# total_trajectories,total_trials,total_steps,temp
total_trajectories,total_trials,total_steps = len(output_spikes_simulation),len(output_spikes_simulation[0]),len(output_spikes_simulation[0][0])
total_trajectories,total_trials,total_steps
# Generates the data to be used with the linear regression
# The first index of the matrix is the trajectory
# Then the 20 versions are accessed in series
# E.g:
# linalg_matrix[0][0:250] => is the first experiment of the first trajectory
# linalg_matrix[0][250:250*2] => is the second experiment of the first trajectory
t_idx = 0 # 0=>first trajectory
e_idx = 0 # 0=>first experiment
# linalg_matrix=numpy.array([numpy.array([numpy.zeros(600) for i in range(250*20)]) for j in range(4)])
linalg_matrix = numpy.zeros((total_trajectories, total_trials*total_steps, NofN),dtype=numpy.float)
for t_idx in range(total_trajectories): # goes through all the trajectories
for e_idx in range(total_trials): # goes through all the trials
for i in range(total_steps): # goes through all the steps - the simulation step is 2ms and it goes from 0 to 498
temp_spikes=numpy.zeros(NofN)
if output_spikes_simulation[t_idx][e_idx][i][0] != []:
temp_spikes[numpy.array(output_spikes_simulation[t_idx][e_idx][i][0][0])]=1 # Writes "1" where each spike occured
if temp_spikes.any():
linalg_matrix[t_idx][i+total_steps*e_idx]=numpy.array(temp_spikes)
# Generates the FILTERED data to be used with the linear regression
# The first index of the matrix is the trajectory
# Then the 20 versions are accessed in series
# E.g:
# linalg_matrix_filtered[0][0:250] => is the first experiment of the first trajectory
# linalg_matrix_filtered[0][250:250*2] => is the second experiment of the first trajectory
t_idx = 0 # 0=>first trajectory
e_idx = 0 # 0=>first experiment
# linalg_matrix_filtered=numpy.array([numpy.array([numpy.zeros(600) for i in range(250*20)]) for j in range(4)])
linalg_matrix_filtered = numpy.zeros((total_trajectories, total_trials*total_steps, NofN),dtype=numpy.float)
for t_idx in range(total_trajectories): # goes through all the trajectories
for e_idx in range(total_trials): # goes through all the trials
m_v=membrane_lowpass(NofN,membrane_time_constant) # Initialize the membrane for each new trial
for i in range(total_steps): # goes through all the steps - the simulation step is 2ms and it goes from 0 to 498
if output_spikes_simulation[t_idx][e_idx][i][0] != []:
m_v.process_spikes(output_spikes_simulation[t_idx][e_idx][i][0][0],output_spikes_simulation[t_idx][e_idx][i][1][0]/1E3) #the /1E3 is to convert to seconds
linalg_matrix_filtered[t_idx][i+total_steps*e_idx]=m_v.check_values(i*sim_step_time) # Saves the membrane state at each time step
```
Now I need to read the CORRECT torque values to be used with the linear regression
```
torques = [] # Each line has the torques matrix of each trajectory
# torques[0][:,0] => returns an array with all the joint 1 torques of the trajectory 1
for i in xrange(1,number_of_trajectories+1):
# The data file below was created using the iPython notebook: "2DofArm_simulation_data_generator.ipynb"
s,t = slf.load_from_file_gz("./"+base_dir+"/"+sim_set+"/States_Torques_movement"+str(i)+".gzpickle")
torques.append(t)
torques = numpy.array(torques)
y_tau1_init=numpy.zeros((torques.shape[0],torques.shape[1]+1,1),dtype=numpy.float)
y_tau2_init=numpy.zeros((torques.shape[0],torques.shape[1]+1,1),dtype=numpy.float)
y_tau1_init[:,:total_steps-1,0]=torques[:,:,0]
y_tau2_init[:,:total_steps-1,0]=torques[:,:,1]
y_tau1=numpy.concatenate([y_tau1_init[j,:,0] for j in xrange(total_trajectories) for i in xrange(total_trials)])
y_tau2=numpy.concatenate([y_tau2_init[j,:,0] for j in xrange(total_trajectories) for i in xrange(total_trials)])
y_tau1.shape,y_tau2.shape
# # The 'if i < 250' is used because the torques list has 249 items and the LSM generates the last state without
# # having to read any inputs (it's the way the simulator works...)
# y_tau1=numpy.array([torques[j][:,0][:][i] if i < 250 else 0 for j in range(total_trajectories) for k in range(total_trials) for i in range(total_steps)])
# y_tau2=numpy.array([torques[j][:,1][:][i] if i < 250 else 0 for j in range(total_trajectories) for k in range(total_trials) for i in range(total_steps)])
# y_tau1.shape,y_tau2.shape
```
# Here I can choose to use the LOW-PASS (membrane) filtered values (according to Maass and Joshi) or using directly the spikes.
### If the variable X_matrix receives the linalg_matrix, it's using only the spikes.
### If the variable X_matrix receives the linalg_matrix_filtered, it's using the values after the membrane filter.
```
# X_matrix can be:
# linalg_matrix_filtered or linalg_matrix
if using_spikes:
X_matrix=linalg_matrix
else:
X_matrix=linalg_matrix_filtered
X_matrix.shape
# Prepare the matrix to be used with numpy linear regression:
X_reshaped=X_matrix.reshape(y_tau1.shape[0],NofN)
# Creates an empty matrix with an extra collumn with ones (numpy.linalg.lstsq demands this...)
X_reshaped=numpy.ones((X_reshaped.shape[0],X_reshaped.shape[1]+1))
# Writes the values in to the first 600 collumns
X_reshaped[:,:NofN]=X_matrix.reshape(y_tau1.shape[0],NofN)
X_reshaped.shape
X=X_reshaped
numpy.shape(X)[0]*numpy.shape(X)[1]
X.max(),X.min(),y_tau1.max(),y_tau1.min(),y_tau2.max(),y_tau2.min()
y_tau1_normal=numpy.array(y_tau1)
y_tau2_normal=numpy.array(y_tau2)
# Here I'm doing the equivalent to (t+1). Using this trick the readout is going to predict the next value.
# Because I know that the start and end torques are always zero, the numpy.roll function fits like a glove!
# y_tau1=numpy.roll(y_tau1,-1)
# y_tau2=numpy.roll(y_tau2,-1)
y_tau1=numpy.concatenate((y_tau1_normal[1:],[0]))
y_tau2=numpy.concatenate((y_tau2_normal[1:],[0]))
y_tau1_normal[10:13],y_tau1[10:12]
%%time
std_x = 0.1
std_y = 0.01
sklg = linear_model.LinearRegression(fit_intercept=True, normalize=True, copy_X=True, n_jobs=7)
# Using sklearn is a lot easier to try another linear model algorithm, I just have to change the above line.
NOISE_X = numpy.random.normal(loc=0,scale=std_x,size=X[:,:600].shape)
NOISE_Y = numpy.random.normal(loc=0,scale=std_y,size=y_tau1.shape)
sklg.fit(X[:,:600]+NOISE_X,y_tau1+NOISE_Y)
c_tau1=sklg.coef_
r_tau1=sklg.intercept_
NOISE_X = numpy.random.normal(loc=0,scale=std_x,size=X[:,:600].shape)
NOISE_Y = numpy.random.normal(loc=0,scale=std_y,size=y_tau2.shape)
sklg.fit(X[:,:600]+NOISE_X,y_tau2+NOISE_Y)
c_tau2=sklg.coef_
r_tau2=sklg.intercept_
```
#### Using sklearn I could try a lot of different readouts:
http://scikit-learn.org/stable/modules/classes.html#module-sklearn.linear_model
```
# This is the same linear regression least squares, but using numpy:
# %%time
# std_x = 0.1
# std_y = 0.01
# NOISE_X = numpy.random.normal(loc=0,scale=std_x,size=X.shape)
# NOISE_Y = numpy.random.normal(loc=0,scale=std_y,size=y_tau1.shape)
# linear_regression1 = numpy.linalg.lstsq(X+NOISE_X, y_tau1+NOISE_Y)
# c_tau1=linear_regression1[0][:-1] # Coefficients
# r_tau1=linear_regression1[0][-1] # Constant
# NOISE_X = numpy.random.normal(loc=0,scale=std_x,size=X.shape)
# NOISE_Y = numpy.random.normal(loc=0,scale=std_y,size=y_tau2.shape)
# linear_regression2 = numpy.linalg.lstsq(X+NOISE_X, y_tau2+NOISE_Y)
# c_tau2=linear_regression2[0][:-1] # Coefficients
# r_tau2=linear_regression2[0][-1] # Constant
# slf.save_to_file([c_tau1,r_tau1],"./"+base_dir+"/"+sim_set+"/coefficients_residues_tau1.pickle")
# slf.save_to_file([c_tau2,r_tau2],"./"+base_dir+"/"+sim_set+"/coefficients_residues_tau2.pickle")
y_tau1_calculated=X_matrix.reshape(y_tau1.shape[0],NofN).dot(c_tau1)+r_tau1
y_tau2_calculated=X_matrix.reshape(y_tau2.shape[0],NofN).dot(c_tau2)+r_tau2
# Plots the inputs and the outputs side-by-side
# %matplotlib inline
offset11 = y_tau1.shape[0]/number_of_trajectories
offset21 = y_tau2.shape[0]/number_of_trajectories
offset12 = y_tau1.shape[0]/number_of_trajectories/total_trials
offset22 = y_tau2.shape[0]/number_of_trajectories/total_trials
fig=plt.figure(figsize =(20,40));
for trajectory in xrange(number_of_trajectories):
# trajectory=0 # goes from 0 to 3
ymax=numpy.array([y_tau1[trajectory*(total_trials*251):251*(1+trajectory*(total_trials*251))].max(),y_tau2[trajectory*(total_trials*251):251*(1+trajectory*(total_trials*251))].max()]).max()
ymin=numpy.array([y_tau1[trajectory*(total_trials*251):251*(1+trajectory*(total_trials*251))].min(),y_tau2[trajectory*(total_trials*251):251*(1+trajectory*(total_trials*251))].min()]).min()
ymaxLSM=numpy.array([y_tau1_calculated[trajectory*(total_trials*251):251*(1+trajectory*(total_trials*251))].max(),y_tau2_calculated[trajectory*(total_trials*251):251*(1+trajectory*(total_trials*251))].max()]).max()
yminLSM=numpy.array([y_tau1_calculated[trajectory*(total_trials*251):251*(1+trajectory*(total_trials*251))].min(),y_tau2_calculated[trajectory*(total_trials*251):251*(1+trajectory*(total_trials*251))].min()]).min()
plt.subplot(number_of_trajectories,2,2*trajectory+1)
plt.plot(y_tau1[trajectory*(offset11):offset11*trajectory+offset12],'.-')
plt.plot(y_tau2[trajectory*(offset21):offset21*trajectory+offset22],'.-')
# plt.plot(y_tau1[trajectory*(offset11):offset12*(1+trajectory*(total_trials*251))][:251],'.-')
# plt.plot(y_tau2[trajectory*(offset21):offset22*(1+trajectory*(total_trials*251))][:251],'.-')
plt.ylim(ymin,ymax)
plt.title("Original analog torques - trajectory " + str(trajectory+1))
plt.subplot(number_of_trajectories,2,2*trajectory+2)
plt.plot(y_tau1_calculated.reshape(number_of_trajectories,total_trials,251)[trajectory,:,:].mean(axis=0),'b.-')
plt.plot(y_tau1_calculated.reshape(number_of_trajectories,total_trials,251)[trajectory,:,:].mean(axis=0)+y_tau1_calculated.reshape(number_of_trajectories,total_trials,251)[trajectory,:,:].std(axis=0),'b')
plt.plot(y_tau1_calculated.reshape(number_of_trajectories,total_trials,251)[trajectory,:,:].mean(axis=0)-y_tau1_calculated.reshape(number_of_trajectories,total_trials,251)[trajectory,:,:].std(axis=0),'b')
plt.plot(y_tau2_calculated.reshape(number_of_trajectories,total_trials,251)[trajectory,:,:].mean(axis=0),'g.-')
plt.plot(y_tau2_calculated.reshape(number_of_trajectories,total_trials,251)[trajectory,:,:].mean(axis=0)+y_tau2_calculated.reshape(number_of_trajectories,total_trials,251)[trajectory,:,:].std(axis=0),'g')
plt.plot(y_tau2_calculated.reshape(number_of_trajectories,total_trials,251)[trajectory,:,:].mean(axis=0)-y_tau2_calculated.reshape(number_of_trajectories,total_trials,251)[trajectory,:,:].std(axis=0),'g')
plt.ylim(yminLSM,ymaxLSM)
plt.title("LSM output (trials mean/std values) torques - trajectory " + str(trajectory+1))
# fig.subplots_adjust(bottom=0,hspace=.6) # Adjust the distance between subplots
plt.tight_layout()
plt.show()
plt.rcParams['figure.figsize'] = 20, 10
plt.plot(c_tau1,'v',label="Coefficients tau1")
plt.plot(c_tau2,'s',label="Coefficients tau2")
plt.title("Coefficients")
plt.legend()
plt.show()
# Independent terms
r_tau1,r_tau2
```
| github_jupyter |
```
import torch
from inception_model import InceptionSham
from torchvision import datasets, transforms
from torch.autograd import Variable
import torch.nn.functional as F
import numpy as np
import tqdm
def iterate_minibatches(inputs, targets, batchsize, shuffle=False):
assert len(inputs) == len(targets)
if shuffle:
indices = np.random.permutation(len(inputs))
for start_idx in trange(0, len(inputs) - batchsize + 1, batchsize):
if shuffle:
excerpt = indices[start_idx:start_idx + batchsize]
else:
excerpt = slice(start_idx, start_idx + batchsize)
yield inputs[excerpt], targets[excerpt]
def weights_init(m):
classname = m.__class__.__name__
if (classname.find('Conv')) != -1 and (classname.find('Basic') == -1):
m.weight.data.normal_(0,
0.02)
elif classname.find('BatchNorm') != -1:
m.weight.data.normal_(1,
0.02)
m.bias.data.fill_(0)
BATCH_SIZE = 30
N_EPOCH = 1
train_loader = torch.utils.data.DataLoader(
datasets.MNIST("../data", train=True, download=False, transform=transforms.ToTensor()),
batch_size=BATCH_SIZE,
shuffle=True)
test_loader = torch.utils.data.DataLoader(
datasets.MNIST("../data", train=False, download=False, transform=transforms.ToTensor()),
batch_size=BATCH_SIZE,
shuffle=True)
model = InceptionSham(num_classes=10, input_nc=1, dropout=0.5)
if torch.cuda.is_available():
model.cuda()
opt = torch.optim.Adam(model.parameters())
model.apply(weights_init)
print('inited')
def train(model, n_epoch):
model.train()
for epoch in range(n_epoch):
for i, (data, target) in tqdm.tqdm(enumerate(train_loader)):
if torch.cuda.is_available():
data = data.cuda()
target = target.cuda()
data = Variable(data)
target = Variable(target)
opt.zero_grad()
output = model(data)
loss = F.nll_loss(F.log_softmax(output, dim=1), target)
loss.backward()
opt.step()
def test(model):
model.eval()
accuracy = []
for i, (data, target) in enumerate(test_loader):
if torch.cuda.is_available():
data = data.cuda()
target = target.cuda()
data, target = Variable(data), Variable(target)
output = model(data).data.max(1)
accuracy.append(torch.sum(model(data).data.max(1)[1] == target.data) / BATCH_SIZE)
if i == 15:
break
return np.mean(accuracy)
train(model, n_epoch=1)
test(model)
inception_state = torch.load("../../../inception_sham_state.pth", map_location="cpu")
model.load_state_dict(inception_state)
test(model)
```
| github_jupyter |
Definition of post_pro() function
```
#
# Copyright (C) 2021 by Katja Gilly <katya@umh.es>
#
# This code is licensed under a Creative Commons Attribution 4.0 International License. (see LICENSE.txt for details)
#
# General Description - this notebook is used to extract data to compute cloud and edge delays from CloudSim logs
#
# It creates 7 types of output files per CloudSim simulation log that contain:
# - allocation information of virtualisation entities:
# - when allocations are optimal, results are exctracted to <res_alloc_OPT_TIMES_> files
# - otherwise, allocation information is collected in <res_fail_alloc_detail_> files
# - migration information of virtualisation entities:
# - when migrations are optimal, results are extracted to <res_migr_OPT_TIMES_> files
# - otherwise, migration information is collected in <res_fail_migr_detail_> files
# - when migration reattempts are successful, results are extracted to <res_reattempt_OK_migr_> files
# - when reattempts are unsuccessful, information is sent to <res_reattempt_FAIL_migr_> files
# - energy consumption information is extracted to <res_DCenergy_> files
#
# It also creates 2 files with global data of the whole set of simulations:
# - one about allocations (<total_resAllocHops.csv>) and
# - another about migrations (<total_resMigHops.csv>)
def post_pro(scenario, vehicles, VMs, path):
# parameters
# 1 = $scenario - input scenario format : "A B C D" (A=#MEChosts, B=hostType, C=OptGoal-allocations, D=OptGoal-migrations) i.e. "45 1 sc sc-mad",
# 2 = $vehicles - input number of vehicles
# 3 = $VMs - input number of VMs (if omnet simulation time is reduced from SUMO's, this value is different from $vehicles)
# 4 = $path - input files path
# two output files: resAllocHops.txt resMigHops.txt
# add only the lines that start with a number
(noHosts, hostType, ap, mp) = scenario.split()
print(noHosts, hostType, ap, mp)
# Log format: fogSUMOCloudSim_Log__117_1_sc_sc-mad_2509_initialPositioning-6900.txt_migrations-6900.txt
fileName = path + "fogSUMOCloudSim_Log__" + noHosts + "_" + hostType + "_" + ap + "_" + mp + "_" + VMs + "_initialPositioning-" + vehicles + ".txt_migrations-" + vehicles + ".txt.txt"
path = path + "post_pro/"
optMigrFileTimes = path + "res_migr_OPT_TIMES_" + noHosts + "_" + hostType + "_" + ap + "_" + mp + "_" + VMs + "_" + vehicles + ".txt"
reatFAILMigrFile = path + "res_reattempt_FAIL_migr_" + noHosts + "_" + hostType + "_" + ap + "_" + mp + "_" + VMs + "_" + vehicles + ".txt"
reatOKMigrFile = path + "res_reattempt_OK_migr_" + noHosts + "_" + hostType + "_" + ap + "_" + mp + "_" + VMs + "_" + vehicles + ".txt"
FailMigrFileDetail = path + "res_fail_migr_detail_" + noHosts + "_" + hostType + "_" + ap + "_" + mp + "_" + VMs + "_" + vehicles + ".txt"
allocOPTFileTimes = path + "res_alloc_OPT_TIMES_" + noHosts + "_" + hostType + "_" + ap + "_" + mp + "_" + VMs + "_" + vehicles + ".txt"
FailAllocFileDetail = path + "res_fail_alloc_detail_" + noHosts + "_" + hostType + "_" + ap + "_" + mp + "_" + VMs + "_" + vehicles + ".txt"
resDCenergy = path + "res_DCenergy_" + noHosts + "_" + hostType + "_" + ap + "_" + mp + "_" + VMs + "_" + vehicles + ".txt"
print(fileName)
# ENERGY: get energy data
!grep "Data center's energy" $fileName > temp
!echo "Time;W*sec" > $resDCenergy
!awk '{ print $1, $6 }' temp >> temp2
!cat temp2 >> $resDCenergy
!rm temp temp2
# ALLOCATIONS ANALYSIS: output files: resFileAllocHops.txt $allocOPTFileTimes
# get initial allocations
!grep "Initial placement" $fileName > res_alloc.txt
!echo "$noHosts;$hostType;$ap;$mp;$VMs;$vehicles" | tr -s '\r\n' ';' > resFileAllocHops.txt
# get the number of lines with optimal allocation
!grep -o 'optimal=true' res_alloc.txt | wc -l | tr -s '\r\n' ';' >> resFileAllocHops.txt
# get the number of lines with non optimal allocations
!grep -o 'optimal=false' res_alloc.txt | wc -l | tr -s '\r\n' ';' >> resFileAllocHops.txt
!echo "" >> resFileAllocHops.txt # new line
# check optimal allocations: time, car and community
!echo "Time Vm Comm" > $allocOPTFileTimes
!grep 'optimal=true' res_alloc.txt > temp
!awk '{print $1, $6, $16}' temp > temp2
!sed 's/\#//g' temp2 >> $allocOPTFileTimes
# check failed allocations
!echo "Time Vm Vm_resources Source_Comm Host Dest_Comm Dest_Comm_decision" > $FailAllocFileDetail
!grep 'optimal=false' res_alloc.txt > temp
!awk '{print $1, $6, $7, $10, $13, $16, $17}' temp > temp2
!sed 's/\#//g' temp2 >> $FailAllocFileDetail
!rm temp temp2 res_alloc.txt
# MIGRATIONS ANALYSIS: output files: resFileMigHops.txt $FailMigrFileDetail $optMigrFileTimes
# check reattempted migrations
!echo "Time Vm " > $reatOKMigrFile
!grep "reattempt migration: VM *" $fileName > temp
!awk '{printf ("%.2f %s\n",$1,$5)}' temp > temp2
!sed 's/\#//g' temp2 >> $reatOKMigrFile
# check optimal migrations
!grep -e "Optimal migration" $fileName > res_opt_migr_opt.txt
!grep -e "identical" $fileName > res_opt_migr_ident.txt
!echo "$noHosts;$hostType;$ap;$mp;$VMs;$vehicles" | tr -s '\r\n' ';' > resFileMigHops.txt
# check migrations failed
!echo "Time Vm " > $reatFAILMigrFile
!grep " reattempt migration failed" $fileName > temp
!awk '{printf ("%.2f %s\n",$1,$6)}' temp | sort -g | uniq > temp2
!sed 's/\#//g' temp2 >> $reatFAILMigrFile
# get the times of optimal migrations
!echo "Time Vm Source_Comm Host Dest_Comm " > $optMigrFileTimes
!awk '{print $1, $6, $9, $12, $15}' res_opt_migr_opt.txt > temp2
!awk '{print $1, $8, $13, $10, $13}' res_opt_migr_ident.txt >> temp2
!sed 's/\#//g' temp2 >> $optMigrFileTimes
# get the number of optimal migrations
!wc -l temp2 > temp
!awk '{print $1}' temp | tr -s '\r\n' ';' >> resFileMigHops.txt
# Non optimal migrations
!echo "Time Vm Source_Comm Dest_Comm" > $FailMigrFileDetail
!grep "Attempt to migrate failed" $fileName > temp
# round attempt times to 2 decimals and remove duplicates
!awk '{printf ("%.2f %s %s %s\n",$1,$7,$10,$13)}' temp | sort -g | uniq > temp2 #| uniq -u
!sed 's/\#//g' temp2 >> $FailMigrFileDetail
# get the number of non optimal migrations
!wc -l $FailMigrFileDetail > temp
!awk '{print $1}' temp | tr -s '\r\n' ';' >> resFileMigHops.txt
# get the number of succesful migration reattempts
!wc -l $reatOKMigrFile > temp
!awk '{print $1}' temp | tr -s '\r\n' ';' >> resFileMigHops.txt
# get the number of unsuccesful migration reattempts
!wc -l $reatFAILMigrFile > temp
!awk '{print $1}' temp | tr -s '\r\n' ';' >> resFileMigHops.txt
!echo "" >> resFileMigHops.txt # new line
!rm res_opt_migr_opt.txt res_opt_migr_ident.txt temp temp2
```
Input data : scenarios + vehicles
```
# parameters
# 1 = $scenario - input scenario format : "A B C D" (A=#MEChosts, B=hostType, C=OptGoal-allocations, D=OptGoal-migrations) i.e. "45 1 sc sc-mad",
# 2 = $vehicles - input number of vehicles
# 3 = $VMs - input number of VMs (if omnet simulation time is reduced from SUMO's, this value is different from $vehicles)
# 4 = $path - input files path
# two output files: resAllocHops.txt resMigHops.txt
# This code launches the first step of CloudSim log information extraction process, calls above function and creates 2 files with global data of the whole set of simulations:
# - one about allocations (<total_resAllocHops.csv>) and
# - another about migrations (<total_resMigHops.csv>)
# measuring execution time
!pip install ipython-autotime
%load_ext autotime
# define vehicles and scenarios
scenarios = ["45 1 sc sc-mad","63 1 sc sc-mad","81 1 sc sc-mad","99 1 sc sc-mad","117 1 sc sc-mad",
"45 0 sc sc-mad","63 0 sc sc-mad","81 0 sc sc-mad","99 0 sc sc-mad","117 0 sc sc-mad"]
vehicles = ['4956','4951','4955','5740','5734','5749','6910','6915','6923','8600','8620','8635']
VMs = ['4256','5074','5013','4966','5808','5858','6042','6187','7004','7455','7463','7486']
duration="10000"
!rm *.csv
#aBorrar2="res_*.txt"
path = "/content/drive/MyDrive/processed_logs/cloudsim_output/log_10000s/"
resFileAllocHops = path + "post_pro/" + "total_resAllocHops.csv"
resFileMigHops = path + "post_pro/" + "total_resMigHops.csv"
!echo "Hosts;Type;Ap;Mp;noVMs;noCars;Optimal;Non_optimal;Reatt_OK;Reatt_FAILED" > $resFileMigHops
!echo "Hosts;Type;Ap;Mp;noVMs;noCars;Optimal;Non_optimal" > $resFileAllocHops
for scen in scenarios:
i = 0
for vehi in vehicles:
!rm *.txt
post_pro(scen,vehi,VMs[i],path)
i = i+1
# update hops
!cat resFileAllocHops.txt >> $resFileAllocHops
!cat resFileMigHops.txt >> $resFileMigHops
```
| github_jupyter |
Imports.
```
import sys
sys.path.insert(0, 'utils/')
import datetime
import time
import os
import numpy as np
import matplotlib.pyplot as plt
import numba
from numba import njit
from scipy.integrate import quad
import tensorflow as tf
# because of tf version many warnings will occur
tf.compat.v1.logging.set_verbosity(tf.compat.v1.logging.ERROR)
print('TF version: {}'.format(tf.__version__))
print('GPU support enabled: {}'.format(tf.test.is_built_with_gpu_support()))
print('CUDA support enabled: {}'.format(tf.test.is_built_with_cuda()))
import deepxde as dde
```
Figures configuration for journal papers.
```
from plotting import latexconfig, figsize
latexconfig()
```
# Pennes Bioheat Transfer Equation
**Ref:** Analytical Study on Bioheat Transfer Problems with Spatial or Transient Heating on Skin Surface or Inside Biological Bodies; Zhong-Shan Deng, Jing Liu; DOI: 10.1115/1.1516810
The 1-D Pennes Bioheat Transfer Equation:
\begin{equation}
k \frac{d^2 T(x)}{dx^2} + \omega_b \cdot T_a - \omega_b \cdot T(x) + Q_m + Q_{ext} = 0
\end{equation}
$k$ is the thermal conductivity of the tissue; $\omega_b$ is the blood perfusion; $T_a$ is the arterial temperature and $T$ stands for the tissue temperature. $Q_m$ is the metabolic heat generation and $Q_{ext}$ is the heat generated from an external source.
Mixed boundary conditions are defined; Robins on the skin surface:
\begin{equation}
-k \frac{dT_0(x)}{dx}=h_0[T_f - T_0(x)] \hspace{10mm} x=0
\end{equation}
and Dirichlet at maximum heat penetration depth:
\begin{equation}
T_L(x) = T_c \hspace{10mm} x=L
\end{equation}
where $h_0$, $T_c$ and $T_f$ are the convection coefficient, the body core temperature and the atmospheric temperature, respectively.
****
In this study, time-invariant Pennes' bioheat transfer equation is solved using a novel neural network approach, where calculations were performed for no external source of heating, $Q_{ext} = 0$.
```
# consts
c_b = 4200
rho_b = 1e3
om_b = 5e-4
c = 4200
h_f = 100;
T_c = 37
L = 3e-2
# random variables
k = 0.5
w_b = c_b*rho_b*om_b
Q_m = 33800
h_0 = 10
T_f = 25
T_a = 37
rvs = [k, w_b, T_a, Q_m, h_0, T_f]
rvs = np.array(rvs).reshape(1, -1)
# solution domain
x = np.linspace(0, L, num=49)
```
## Analytical solution
```
def analyticSol(x):
A = np.sqrt(w_b/k)
lterm = (T_c - T_a - Q_m/w_b) * (A * np.cosh(A * x) +\
+ (h_0/k) * np.sinh(A * x)) / (A * np.cosh(A * L) + (h_0/k) * np.sinh(A * L))
rterm = h_0/k * (T_f - T_a - Q_m/w_b) * np.sinh(A * (L * np.ones(shape=x.shape) - x)) / ( A * np.cosh(A * L) + (h_0/k) * np.sinh(A * L))
return T_a + Q_m/w_b + lterm + rterm
T_anal = analyticSol(x)
fig = plt.figure(figsize=figsize(1))
ax = fig.add_subplot(111)
ax.plot(x, T_anal, 'r-', label='$T(x)$')
ax.set_xlabel('$x$ [m]')
ax.set_ylabel('$T$ [°C]')
plt.grid()
plt.legend(loc='best')
plt.show()
fig.savefig('{}.pdf'.format('figs/analytical-solution'), bbox_inches='tight')
```
## Monte Carlo Uncertainty Analysis
```
@njit
def monteCarloSim(rvs, coef_var=0.2, num_samples=1000, num_iters=1000):
sims_global = np.empty(shape=(num_iters, num_samples, rvs.shape[1]))
for i in range(num_iters):
xi = np.random.rand(num_samples, len(rvs)) # random values
sims = np.zeros(shape=xi.shape) # empty simulations matrix
rvs_min = rvs - coef_var*rvs
rvs_max = rvs + coef_var*rvs
sims = rvs_min + (rvs_max - rvs_min) * xi
sims_global[i, :, :] = sims
return sims_global
@njit
def monteCarloT(x, K, w_b, T_a, Q_m, h_0, T_f):
A = np.sqrt(w_b/K)
lterm = (T_c - T_a - Q_m/w_b) * (A * np.cosh(A * x) +\
+ (h_0/K) * np.sinh(A * x)) / (A * np.cosh(A * L) + (h_0/K) * np.sinh(A * L))
rterm = h_0/K * (T_f - T_a - Q_m/w_b) * np.sinh(A * (L * np.ones(shape=x.shape) - x)) / ( A * np.cosh(A * L) + (h_0/K) * np.sinh(A * L))
return T_a + Q_m/w_b + lterm + rterm
@njit
def monteCarloResample(x, mc):
num_iters, num_samples, n_rv = mc.shape
T_mc = np.empty(shape=(num_iters, num_samples, x.shape[0]))
for it in range(num_iters):
for sample in range(num_samples):
T_mc[it, sample, :] = monteCarloT(x, mc[it, sample, 0], mc[it, sample, 1], mc[it, sample, 2], mc[it, sample, 3], mc[it, sample, 4], mc[it, sample, 5])
return T_mc
num_samples = 100
num_iters = 100
coef_var = 0.2
start = time.time()
mc = monteCarloSim(rvs, coef_var, num_samples, num_iters)
T_mc = monteCarloResample(x, mc)
print('Simulation time: {}s'.format(time.time() - start))
# single sample
_sample_exmp = 0
T_mc0 = T_mc[_sample_exmp]
T_mc0_mean = np.mean(T_mc0, axis=0)
T_mc0_std = np.std(T_mc0, axis=0)
T_mc0_lower_bound = T_mc0_mean - T_mc0_std
T_mc0_upper_bound = T_mc0_mean + T_mc0_std
fig = plt.figure(figsize=figsize(1, 2))
ax = fig.add_subplot(211)
ax.plot(x, T_mc0_mean, 'r-', label='$<T(x)>$')
ax.plot(x, T_mc0_lower_bound, 'k--', label='$<T(x)> \pm \sigma_T$')
ax.plot(x, T_mc0_upper_bound, 'k--')
ax.fill_between(x, T_mc0_lower_bound, T_mc0_upper_bound, alpha=0.2)
ax.set_xlabel('$x$ [m]')
ax.set_ylabel('$<T>$ [°C]')
plt.legend(loc='best')
# plt.title('Single MC sample')
plt.grid()
# full simulation
T_mc_mean = np.mean(T_mc, axis=1)
T_mean = np.mean(T_mc_mean, axis=0)
T_std = np.std(T_mc_mean, axis=0)
T_lower_bound = T_mc_mean[np.argmin(T_mc_mean[:, 0], axis=0)]
T_upper_bound = T_mc_mean[np.argmax(T_mc_mean[:, 0], axis=0)]
ax = fig.add_subplot(212)
ax.plot(x, T_mean, 'r-', label='MCM ({} samples, {} epochs) mean'.format(num_samples, num_iters))
ax.plot(x, T_lower_bound, 'k--', label='MCM min')
ax.plot(x, T_upper_bound, 'k--', label='MCM max')
ax.fill_between(x, T_lower_bound, T_upper_bound, alpha=0.2)
ax.set_xlabel('$x$ [m]')
ax.set_ylabel('$<T>$ [°C]')
plt.legend(loc='best')
# plt.title('MC num. of epochs: {}'.format(num_iters))
plt.grid()
fig.savefig('{}-{}epochs-{}samples.pdf'.format('figs/mc-simulation', num_iters, num_samples), bbox_inches='tight')
plt.show()
# distribution of sample means
def ecdf(data):
"""Compute ECDF for a one-dimensional array of measurements."""
n = len(data)
x = np.sort(data)
y = np.arange(1, n+1) / n
return x, y
# distribution
_exmp = 0
x_cdf, y_cdf = ecdf(T_mc_mean[:, _exmp])
# theretical gaussian distribution based on the MC simualted data
samples = np.random.normal(np.mean(T_mc_mean[:, _exmp]), np.std(T_mc_mean[:, _exmp]), 10000)
x_theor, y_theor = ecdf(samples)
fig = plt.figure(figsize=figsize(1))
ax = fig.add_subplot(111)
ax.plot(x_cdf, y_cdf, marker='o', alpha=0.6, label='mean of samples on surface \n($x=0$)')
ax.plot(x_theor, y_theor, 'r-', linewidth=2, label='Gaussian fit')
ax.set_xlabel('$T$ [°C]')
ax.set_ylabel('CDF')
plt.grid()
plt.legend(loc='best')
fig.savefig('{}.pdf'.format('figs/mc-distribution'), bbox_inches='tight')
plt.show()
```
## Finite Element Numerical Solution
```
def assemble_l(lhss, lhs_glob):
if lhs_glob.shape[0]!=lhs_glob.shape[1]:
raise Exception("Global matrix is not square matrix!")
else:
for i, lhs in enumerate(lhss):
lhs_glob[i:i+lhs.shape[0], i:i+lhs.shape[1]] += lhs
return lhs_glob
def assemble_r(rhss, rhs_glob):
for i, rhs in enumerate(rhss):
rhs_glob[i:i+rhs.shape[0], ] += rhs
return rhs_glob
# domain
L = 3e-2
x = np.linspace(0, L, 100)
# fem
N = 20 # number of elements
# solution
lhs_glob = np.zeros((N+1, N+1))
T = np.zeros((N+1, 1))
rhs_glob = np.zeros((N+1, 1))
# base function combinations
n1 = lambda x, a, b: (w_b*T_a + Q_m) * (b-x)/(b-a)
n2 = lambda x, a, b: (w_b*T_a + Q_m) * (x-a)/(b-a)
nn11 = lambda x, a, b: w_b * ((b-x)/(b-a))**2
nn12 = lambda x, a, b: w_b * ((x-a)*(b-x))/((b-a)**2)
nn21 = lambda x, a, b: w_b * ((x-a)*(b-x))/((b-a)**2)
nn22 = lambda x, a, b: w_b * ((x-a)/(b-a))**2
nn_x11 = lambda x, a, b: k/((b-a)**2)
nn_x12 = lambda x, a, b: -k/((b-a)**2)
nn_x21 = lambda x, a, b: -k/((b-a)**2)
nn_x22 = lambda x, a, b: k/((b-a)**2)
a = 0
dx = L/N
b = dx
lhs = np.zeros((2, 2))
rhs = np.zeros((2, 1))
lhss = []
rhss = []
for i in range(N):
print('{}. element: integration domain [{}, {}]'.format(i+1, a, b))
lhs[0,0] = -(quad(nn11, a, b, args=(a, b))[0] + quad(nn_x11, a, b, args=(a, b))[0])
lhs[0,1] = -(quad(nn12, a, b, args=(a, b))[0] + quad(nn_x12, a, b, args=(a, b))[0])
lhs[1,0] = -(quad(nn21, a, b, args=(a, b))[0] + quad(nn_x21, a, b, args=(a, b))[0])
lhs[1,1] = -(quad(nn22, a, b, args=(a, b))[0] + quad(nn_x22, a, b, args=(a, b))[0])
lhss.append(lhs)
rhs[0, 0] = -(quad(n1, a, b, args=(a, b))[0])
rhs[1, 0] = -(quad(n2, a, b, args=(a, b))[0])
rhss.append(rhs)
a = a + dx
b = b + dx
# assemble in global matrices
lhs_glob = assemble_l(lhss, lhs_glob)
rhs_glob = assemble_r(rhss, rhs_glob)
# boundary conds
# DBC
T[-1, 0] = T_c
lhs_glob = lhs_glob[:-1, :-1]
rhs_glob = rhs_glob[:-1, ]
# RBC
lhs_glob[0, 0] -= h_0
rhs_glob[0, 0] -= h_0 * T_f
rhs_glob[-1, 0] -= T_c * lhs_glob[-1, -2]
# T = B \ A
T[:-1, ] = np.linalg.inv(lhs_glob) @ rhs_glob
x_anal = np.linspace(0, L, num=T_anal.shape[0])
x = np.linspace(0, L, num=T.shape[0])
fig = plt.figure(figsize=figsize(1))
ax = fig.add_subplot(111)
ax.axhline(y=T_c, color='black', linestyle=':', label='body/arterial temperature')
ax.axvline(x=x[np.where(np.max(T)==T)[0]], color='black', linestyle='-.', label='peak temperature depth')
ax.plot(x_anal, T_anal, linewidth=2, color='red', label='analytical')
ax.plot(x, T, linestyle='None', color='red', marker='o', markersize=7, label='FEM ({} elements)'.format(N))
ax.set_xlabel('$x$ [m]')
ax.set_ylabel('$T$ [°C]')
plt.grid()
plt.legend(loc='best')
# plt.title('{} finite elements'.format(N))
fig.savefig('{}-{}felems.pdf'.format('figs/FEM', N), bbox_inches='tight')
plt.show()
```
## Physics Informed Neural Network
Ref 1: [Data-driven Solutions of Nonlinear Partial Differential Equations, M. Raissi et al.](https://arxiv.org/abs/1711.10561)
Ref 2: [DeepXDE: A deep learning library for solving differential equations, Lu Lu et al.](https://arxiv.org/abs/1907.04502)
Data-driven solution of time-independent PDE using Physics-informed Neural Network approach. Instead of dealing with it numerically, the solution is transformed into optimization problem:
$$
k \cdot \frac{d^2 T}{d x^2} - w_b \cdot T(x) + w_b \cdot T_a + Q_m = 0
$$
$$ \Downarrow $$
$$
J(\theta) = [k \cdot \frac{d^2 \hat{T}}{d x^2} - w_b \cdot \hat T(x) + w_b \cdot T_a + Q_m]^2 + [\hat T(0) - T_0]^2 + [\hat T(L) - T_L]^2
$$
$$
\mbox{Optimization: } \fbox{ $\min_{\theta} J(\theta)$}
$$
where:
* $J(\theta)$ is the cost function;
* $\hat{T}(x)$ is the neural network's output for every input x and
* $T_0$ and $T_L$ are boundary conditions.
Neural network configuration:
* activation: tanh
* initializer: Glorot uniform
* architecture: input (1), 3 hidden (50), output (1)
* optimization: Adam
* metrics: l2 relative error
```
def pde(x, y):
dy_x = tf.gradients(y, x)[0]
dy_xx = tf.gradients(dy_x, x)[0]
return k*dy_xx - w_b*y + (w_b*T_a + Q_m)
def boundary_l(x, on_boundary):
return on_boundary and np.isclose(x[0], 0)
def boundary_r(x, on_boundary):
return on_boundary and np.isclose(x[0], L)
geom = dde.geometry.Interval(0, L)
#bc_l = dde.RobinBC(geom, lambda x, y: (h_0/k)*y - (h_0*T_f/k), boundary_l)
bc_l = dde.DirichletBC(geom, lambda x: analyticSol(x), boundary_l)
bc_r = dde.DirichletBC(geom, lambda x: analyticSol(x), boundary_r)
data = dde.data.PDE(geom, 1, pde, [bc_l, bc_r], 20, 2, func=analyticSol, num_test=100)
layer_size = [1] + [50] * 3 + [1]
activation = "tanh"
initializer = "Glorot uniform"
net = dde.maps.FNN(layer_size, activation, initializer)
net.outputs_modify(lambda x, y: y*10)
t = datetime.datetime.now().strftime('%Y%m%d-%H%M%S')
new_model_path = 'model/{}'.format(t)
if not os.path.exists(new_model_path):
os.makedirs(new_model_path)
model = dde.Model(data, net)
model.compile("adam", lr=0.001, metrics=["l2 relative error"], loss_weights=[1e-7, 1e-2, 1])
losshistory, train_state = model.train(epochs=10000, uncertainty=True)
dde.postprocessing.save_loss_history(losshistory, 'model/{}/loss.dat'.format(t))
dde.postprocessing.save_best_state(train_state, 'model/{}/train.dat'.format(t), 'model/{}/test.dat'.format(t))
X_train, y_train, X_test, y_test, best_y, best_ystd = train_state.packed_data()
y_dim = y_train.shape[1]
idx = np.argsort(X_test[:, 0])
X = X_test[idx, 0]
# regression plot
fig = plt.figure(figsize=figsize(1, nplots=1))
ax = fig.add_subplot(111)
for i in range(y_dim):
ax.plot(X_train[:, 0], y_train[:, i], 'ok', label='True points')
ax.plot(X, best_y[idx, i], "-r", label='Prediction')
ax.set_xlabel('$x$ [m]')
ax.set_ylabel('$T$ [°C]')
plt.grid()
plt.legend(loc='best')
fig.savefig('{}.pdf'.format('figs/regression-mean'), bbox_inches='tight')
plt.show()
fig = plt.figure(figsize=figsize(1, nplots=1))
ax = fig.add_subplot(111)
for i in range(y_dim):
#ax.plot(X_train[:, 0], y_train[:, i], 'ok', label='True points')
if best_ystd is not None:
ax.plot(X, best_y[idx, i] + 2 * best_ystd[idx, i], '-b', label='95\% confidence interval')
ax.plot(X, best_y[idx, i] - 2 * best_ystd[idx, i], '-b')
ax.plot(X, best_y[idx, i], "--r", label='Prediction')
ax.set_xlabel('$x$ [m]')
ax.set_ylabel('$T$ [°C]')
plt.grid()
plt.legend(loc='best')
fig.savefig('{}.pdf'.format('figs/regression-uncertainty'), bbox_inches='tight')
plt.show()
loss_train = np.sum(
np.array(losshistory.loss_train) * losshistory.loss_weights, axis=1
)
loss_test = np.sum(
np.array(losshistory.loss_test) * losshistory.loss_weights, axis=1
)
fig = plt.figure(figsize=figsize(1))
ax = fig.add_subplot(111)
ax.semilogy(losshistory.steps, loss_train, color='C0', label="Train loss")
ax.semilogy(losshistory.steps, loss_test, color='red', label="Test loss")
#for i in range(len(losshistory.metrics_test[0])):
#ax.semilogy(losshistory.steps, np.array(losshistory.metrics_test)[:, i], color='black', label="Test metric",)
ax.set_xlabel('Number of epochs')
ax.set_ylabel(r'$J(\theta)$')
plt.grid()
plt.legend(loc='best')
fig.savefig('{}.pdf'.format('figs/loss'), bbox_inches='tight')
plt.show()
# random variables
rvs = [k, w_b, T_a, Q_m, h_0]
rvs = np.array(rvs).reshape(1, -1)
num_samples = 5
num_iters = 5
rvs_data = monteCarloSim(rvs, coef_var=0.2, num_samples=num_samples, num_iters=num_iters)
rvs_data.shape
# train_state_ensemble = []
# for epoch in range(num_iters):
# train_state_full = []
# for sample in range(num_samples):
# k = rvs_data[epoch, sample, 0]
# w_b = rvs_data[epoch, sample, 1]
# T_a = rvs_data[epoch, sample, 2]
# Q_m = rvs_data[epoch, sample, 3]
# h_0 = rvs_data[epoch, sample, 4]
# geom = dde.geometry.Interval(0, L)
# bc_l = dde.DirichletBC(geom, lambda x: analyticSol(x), boundary_l)
# bc_r = dde.DirichletBC(geom, lambda x: analyticSol(x), boundary_r)
# data = dde.data.PDE(geom, 1, pde, [bc_l, bc_r], 50, 2, func=analyticSol, num_test=100)
# layer_size = [1] + [50] * 3 + [1]
# activation = "tanh"
# initializer = "Glorot uniform"
# net = dde.maps.FNN(layer_size, activation, initializer)
# net.outputs_modify(lambda x, y: y*10)
# model = dde.Model(data, net)
# model.compile("adam", lr=0.001, metrics=["l2 relative error"], loss_weights=[1e-7, 1e-2, 1])
# losshistory, train_state = model.train(epochs=10000, uncertainty=True, display_every=10000)
# train_state_full.append(train_state)
# train_state_ensemble.append(train_state_full)
# X_train = np.zeros(shape=(num_iters, num_samples, 54))
# y_train = np.zeros(shape=(num_iters, num_samples, 54))
# X_test = np.zeros(shape=(num_iters, num_samples, 100))
# y_test = np.zeros(shape=(num_iters, num_samples, 100))
# best_y = np.zeros(shape=(num_iters, num_samples, 100))
# best_ystd = np.zeros(shape=(num_iters, num_samples, 100))
# for i, train_state_full in enumerate(train_state_ensemble):
# for j, train_state in enumerate(train_state_full):
# unpacked = train_state.packed_data()
# X_train[i, j, :] = unpacked[0].ravel()
# y_train[i, j, :] = unpacked[1].ravel()
# X_test[i, j, :] = unpacked[2].ravel()
# y_test[i, j, :] = unpacked[3].ravel()
# best_y[i, j, :] = unpacked[4].ravel()
# best_ystd[i, j, :] = unpacked[5].ravel()
# np.save('data/X_train.npy', X_train)
# np.save('data/y_train.npy', y_train)
# np.save('data/X_test.npy', X_test)
# np.save('data/y_test.npy', y_test)
# np.save('data/best_y.npy', best_y)
# np.save('data/best_ystd.npy', best_ystd)
X_train = np.load('data/X_train.npy')
y_train = np.load('data/y_train.npy')
X_test = np.load('data/X_test.npy')
y_test = np.load('data/y_test.npy')
best_y = np.load('data/best_y.npy')
best_ystd = np.load('data/best_ystd.npy')
best_y_mc_mean = np.mean(best_y, axis=1)
best_y_mean = np.mean(best_y_mc_mean, axis=0)
best_y_std = np.std(best_y_mean, axis=0)
best_y_lower_bound = best_y_mc_mean[np.argmin(best_y_mc_mean[:, 0], axis=0)]
best_y_upper_bound = best_y_mc_mean[np.argmax(best_y_mc_mean[:, 0], axis=0)]
fig = plt.figure(figsize=figsize(1))
ax = fig.add_subplot(111)
ax.plot(X, best_y_mean, 'r-', label='MC Neural Network mean')
ax.plot(X, best_y_lower_bound, 'k--', label='MC lower bound')
ax.plot(X, best_y_upper_bound, 'k--', label='MC upper bound')
ax.fill_between(X, best_y_lower_bound, best_y_upper_bound, alpha=0.2)
ax.set_xlabel('$x$ [m]')
ax.set_ylabel('$<T>$ [°C]')
plt.legend(loc='best')
plt.grid()
fig.savefig('{}.pdf'.format('figs/uncertainty-mcnn-{}epochs{}samples'.format(num_iters, num_samples)), bbox_inches='tight')
plt.show()
```
| github_jupyter |
# Supplementary material
This notebook generates the figures from the supplementary section of *Discovery of Physics from Data: Universal Laws and Discrepancy Models.*
An interactive version of this notebook is available on [binder](https://mybinder.org/v2/gh/briandesilva/discovery-of-physics-from-data/master?filepath=code%2Fsupplementary_material.ipynb).
[](https://mybinder.org/v2/gh/briandesilva/discovery-of-physics-from-data/master?filepath=code%2Fsupplementary_material.ipynb)
```
from collections import OrderedDict
from cycler import cycler
import matplotlib
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import pysindy as ps
from scipy.integrate import odeint
import seaborn as sns
import warnings
import helpers
figheight = 6
figwidth = 7.5
title_font = 20
save_plots = False
norm = None # Passed to the ord argument of numpy.linalg.norm
```
## Sparse Identification of Nonlinear Dynamical systems
### A brief example
```
# Generate training data
def f(z, t):
return [
-0.1 * z[0] ** 3 + 2 * z[1] ** 3,
-2 * z[0] ** 3 - 0.1 * z[1] ** 3,
]
dt = 0.01
t = np.arange(0, 25, dt)
z0 = [2, 0]
z = odeint(f, z0, t)
train_inds = t < 5
z_train = z[train_inds]
t_train = t[train_inds]
# Fit the model
poly_order = 5
threshold = 0.05
model = ps.SINDy(
optimizer=ps.STLSQ(threshold=threshold),
feature_library=ps.PolynomialLibrary(degree=poly_order),
feature_names=["x", "y"],
)
model.fit(z_train, t=dt)
model.print()
pal = sns.color_palette("Set1", n_colors=4)
plot_kws = {"alpha": 0.7, "linewidth": 4}
# Simulate and plot the results
z_sim = model.simulate(z0, t)
fig, ax = plt.subplots(1, 1, figsize=(figwidth, figwidth))
ax.plot(z_train[:, 0], z_train[:, 1], color=pal[0], label="Training data", **plot_kws)
ax.plot(
z[~train_inds, 0], z[~train_inds, 1], color=pal[1], label="True model", **plot_kws
)
ax.plot(z_sim[:, 0], z_sim[:, 1], "k--", label="SINDy model", **plot_kws)
ax.legend(fontsize=12)
ax.set(xlabel="$x$", ylabel="$y$")
helpers.resize_fonts(ax)
if save_plots:
fname = "figures/nonlinear_oscillator"
plt.savefig(fname + ".pdf", bbox_inches="tight", pad_inches=0)
fig.show()
```
### Learning simplified equations of motion
```
timesteps = 50
grav = -9.8
drag = -0.5
h0 = 40
# Drag-free trajectory
h_dragfree, t = helpers.synthetic_ball_drop(grav, 0, h0=h0, timesteps=timesteps)
model_dragfree = ps.SINDy(optimizer=ps.STLSQ(threshold=0.1), feature_names=["v"])
v_dragfree = model_dragfree.differentiate(h_dragfree, t)
model_dragfree.fit(v_dragfree, t)
# Linear drag trajectory
h_drag, _ = helpers.synthetic_ball_drop(grav, drag, h0=h0, timesteps=timesteps)
model_drag = ps.SINDy(optimizer=ps.STLSQ(threshold=0.1), feature_names=["v"])
v_drag = model_drag.differentiate(h_drag, t)
model_drag.fit(v_drag, t)
print("Learned model (drag-free training data):")
model_dragfree.print()
print("\nLearned model (linear drag training data):")
model_drag.print()
# Reynolds number dependent drag trajectory
h_re_drag, t = helpers.re_dependent_synthetic_ball_drop(
0.067, accel=grav, mass=0.057, noise=0, timesteps=timesteps, h0=h0
)
model_re_drag = ps.SINDy(optimizer=ps.STLSQ(threshold=0.004), feature_names=["v"])
v_re_drag = model_re_drag.differentiate(h_re_drag, t)
model_re_drag.fit(v_re_drag, t)
print("Learned drag model (Re-dependent drag training data, small threshold):")
model_re_drag.print()
model_re_drag = ps.SINDy(optimizer=ps.STLSQ(threshold=0.1), feature_names=["v"])
model_re_drag.fit(v_re_drag, t)
print("\nLearned drag model (Re-dependent drag training data, large threshold):")
model_re_drag.print()
```
## Numerical differentiation
Plot test data
```
timesteps = 49
grav = -9.8
drag = -0.5
# True height, velocity, and acceleration
h, t = helpers.synthetic_ball_drop(grav, drag, h0=40, timesteps=timesteps)
v = (grav / drag) * (np.exp(drag * t) - 1)
a = grav * np.exp(drag * t)
with sns.axes_style("darkgrid"):
fig, ax = plt.subplots(1, 1, figsize=(10, 5))
noise_levels_plot = [1e-2, 1e-1, 1]
ax.plot(t, h, ":", label=0, alpha=0.7, linewidth=3.5)
for noise in noise_levels_plot:
ax.plot(
t, h + noise * np.random.randn(timesteps), label=noise, alpha=0.7, linewidth=2.5
)
ax.set(
title="Simulated trajectory with different noise levels",
xlabel="Time (s)",
ylabel="Height (m)",
)
ax.legend(title="Noise level", fontsize=12)
helpers.resize_fonts(ax)
# Save results
if save_plots:
fname = "figures/noisy_trajectory"
plt.savefig(fname + ".pdf", bbox_inches="tight", pad_inches=0)
fig.show()
```
### Comparison of differentiation methods
Plot error as a function of noise level for different numerical differentiation schemes
```
noise_levels_test = np.logspace(-3, 0, 200)
diff_methods = ["forward difference", "centered difference"]
v_error_unsmoothed = {diff_method: [] for diff_method in diff_methods}
a_error_unsmoothed = {diff_method: [] for diff_method in diff_methods}
v_error_smoothed = {diff_method: [] for diff_method in diff_methods}
a_error_smoothed = {diff_method: [] for diff_method in diff_methods}
for noise in noise_levels_test:
h_noisy = h + noise * np.random.randn(timesteps)
for diff_method in diff_methods:
# Unsmoothed results
v_approx = helpers.differentiate(h_noisy, t, diff_method=diff_method)
a_approx = helpers.differentiate(v_approx, t, diff_method=diff_method)
v_error_unsmoothed[diff_method].append(
helpers.relative_error(v, v_approx, ord=norm)
)
a_error_unsmoothed[diff_method].append(
helpers.relative_error(a, a_approx, ord=norm)
)
# Smoothed results
v_approx = helpers.differentiate(
h_noisy, t, diff_method=diff_method, smoother="savgol"
)
a_approx = helpers.differentiate(
v_approx, t, diff_method=diff_method, smoother="savgol"
)
v_error_smoothed[diff_method].append(
helpers.relative_error(v, v_approx, ord=norm)
)
a_error_smoothed[diff_method].append(
helpers.relative_error(a, a_approx, ord=norm)
)
with sns.axes_style("whitegrid"):
fig, axs = plt.subplots(1, 2, figsize=(2 * figwidth, figheight), sharey=True)
custom_cycle = cycler(color=sns.color_palette("Paired", n_colors=10))
axs[0].set_prop_cycle(custom_cycle)
axs[1].set_prop_cycle(custom_cycle)
linestyles = ["-", ":", "-.", "--"]
kwargs = {"linewidth": 2.5}
for k, diff_method in enumerate(diff_methods):
kwargs["linestyle"] = linestyles[k % len(diff_methods)]
axs[0].plot(
noise_levels_test,
v_error_unsmoothed[diff_method],
label="{} unsmoothed".format(diff_method),
**kwargs
)
axs[1].plot(
noise_levels_test,
a_error_unsmoothed[diff_method],
label="{} unsmoothed".format(diff_method),
**kwargs
)
axs[0].plot(
noise_levels_test,
v_error_smoothed[diff_method],
label="{} smoothed".format(diff_method),
**kwargs
)
axs[1].plot(
noise_levels_test,
a_error_smoothed[diff_method],
label="{} smoothed".format(diff_method),
**kwargs
)
axs[0].set(
title="Relative error in numerically computed velocity",
xlabel="Noise level",
ylabel="Relative error",
xscale="log",
yscale="log",
xlim=(min(noise_levels_test), max(noise_levels_test)),
)
axs[1].set(
title="Relative error in numerically computed acceleration",
xlabel="Noise level",
xscale="log",
yscale="log",
xlim=(min(noise_levels_test), max(noise_levels_test)),
)
axs[0].legend(
title="Differentiation method", loc="upper left", fontsize=12, title_fontsize=12
)
helpers.resize_fonts(axs, title=16, ticks=12)
fig.tight_layout()
# Save results
if save_plots:
fname = "figures/differentiation_method_comparison"
plt.savefig(fname + ".pdf", bbox_inches="tight", pad_inches=0)
fig.show()
```
### Varying smoothing window
```
window_lengths = np.arange(7, 36, 4)
v_error = {w: [] for w in window_lengths}
a_error = {w: [] for w in window_lengths}
for noise in noise_levels_test:
h_noisy = h + noise * np.random.randn(timesteps)
for window_length in window_lengths:
v_approx = helpers.differentiate(
h_noisy,
t,
diff_method="centered difference",
smoother="savgol",
window_length=window_length,
)
a_approx = helpers.differentiate(
v_approx,
t,
diff_method="centered difference",
smoother="savgol",
window_length=window_length,
)
v_error[window_length].append(helpers.relative_error(v, v_approx, ord=norm))
a_error[window_length].append(helpers.relative_error(a, a_approx, ord=norm))
with sns.axes_style("whitegrid"):
fig, axs = plt.subplots(1, 2, figsize=(2 * figwidth, figheight), sharey=True)
custom_cycle = cycler(color=sns.color_palette("GnBu", len(window_lengths)))
axs[0].set_prop_cycle(custom_cycle)
axs[1].set_prop_cycle(custom_cycle)
kwargs = {"linewidth": 1.5}
for window_length in window_lengths:
axs[0].plot(
noise_levels_test, v_error[window_length], label=window_length, **kwargs
)
axs[1].plot(
noise_levels_test, a_error[window_length], label=window_length, **kwargs
)
axs[0].set(
title="Relative error in numerically computed velocity",
xlabel="Noise level",
ylabel="Relative error",
xscale="log",
yscale="log",
xlim=(min(noise_levels_test), max(noise_levels_test)),
)
axs[1].set(
title="Relative error in numerically computed acceleration",
xlabel="Noise level",
xscale="log",
yscale="log",
xlim=(min(noise_levels_test), max(noise_levels_test)),
)
axs[0].legend(title="Window length", fontsize=12, title_fontsize=12)
helpers.resize_fonts(axs, title=16, ticks=12)
fig.tight_layout()
# Save results
if save_plots:
fname = "figures/differentiation_smoothing_window_comparison"
plt.savefig(fname + ".pdf", bbox_inches="tight", pad_inches=0)
fig.show()
```
### Difference between noisy and smoothed data
How much does the smoothed version of a vector differ from the original noisy version?
```
h_error = {w: [] for w in window_lengths}
for noise in noise_levels_test:
h_noisy = h + noise * np.random.randn(timesteps)
for window_length in window_lengths:
h_smoothed = helpers.smooth_data(
h_noisy, smoother="savgol", window_length=window_length
)
h_error[window_length].append(
helpers.relative_error(h_noisy, h_smoothed, ord=norm)
)
with sns.axes_style("whitegrid"):
fig, ax = plt.subplots(1, 1, figsize=(figwidth, figheight))
custom_cycle = cycler(color=sns.color_palette("GnBu", len(window_lengths)))
ax.set_prop_cycle(custom_cycle)
kwargs = {"linewidth": 1.5}
for window_length in window_lengths:
ax.plot(noise_levels_test, h_error[window_length], label=window_length, **kwargs)
ax.set(
title="Relative difference between noisy and smoothed heights",
xlabel="Noise level",
ylabel="Relative difference",
xscale="log",
yscale="log",
xlim=(min(noise_levels_test), max(noise_levels_test)),
)
ax.legend(title="Window length", fontsize=12, title_fontsize=12)
helpers.resize_fonts(ax, title=16, ticks=12)
fig.tight_layout()
# Save results
if save_plots:
fname = "figures/smoothing_error"
plt.savefig(fname + ".pdf", bbox_inches="tight", pad_inches=0)
fig.show()
```
### Estimating noise level in original data set
We can estimate the noise level in the original data set by assuming the noise is gaussian and checking the relative difference between the trajectories and their smoothed versions.
```
# Read in data
data_file = "../data/Ball_drops_data.xls"
bd_data = pd.read_excel(data_file, sheet_name=None)
# Remove bowling ball
del bd_data["Bowling Ball"]
ball_list = bd_data.keys()
n_balls = len(ball_list)
smoother = "savgol"
window_length = 35
h_diff = [{}, {}]
for drop in [1, 2]:
for ball in bd_data.keys():
ball_df = bd_data[ball].loc[bd_data[ball]["Drop #"] == drop]
h_data = ball_df["Height (m)"].values
t_data = ball_df["Time (s)"].values
ntc = helpers.need_to_clip(t_data)
if ntc:
h_data = h_data[:ntc]
t_data = t_data[:ntc]
h_smoothed = helpers.smooth_data(
h_data, smoother=smoother, window_length=window_length
)
h_diff[drop - 1][ball] = helpers.relative_error(h_data, h_smoothed, ord=norm)
dfs = [pd.DataFrame.from_dict(d, orient="index") for d in h_diff]
with sns.axes_style("whitegrid"):
fig, axs = plt.subplots(1, 2, figsize=(2 * figwidth, figheight))
custom_cycle = cycler(color=sns.color_palette("Set1", 2))
axs[0].set_prop_cycle(custom_cycle)
axs[1].set_prop_cycle(custom_cycle)
kwargs = {"alpha": 0.7, "markersize": 8}
for drop in [1, 2]:
df = dfs[drop - 1]
# Relative difference between smoothed and unsmoothed height
axs[0].plot(
np.arange(n_balls), df.values, "o", label="Drop {}".format(drop), **kwargs
)
# Plot approximate noise level for each ball trajectory
df["approx_noise"] = df.applymap(
lambda x: noise_levels_test[np.argmin(np.abs(np.array(h_error[35]) - x))]
)
axs[1].plot(
np.arange(n_balls),
df["approx_noise"].values,
"o",
label="Drop {}".format(drop),
**kwargs
)
axs[0].set(
title="Relative difference between smoothed and original height",
xticks=np.arange(n_balls),
xticklabels=dfs[0].index,
ylabel="Relative difference (m)",
)
axs[1].set(
title="Approximate noise in measured trajectories",
xticks=np.arange(n_balls),
xticklabels=dfs[0].index,
ylabel="Approximate noise",
)
for ax in axs:
ax.tick_params(axis="x", labelrotation=70)
ax.legend(loc="upper left", fontsize=12, title_fontsize=12)
helpers.resize_fonts(axs, title=16, ticks=12)
fig.tight_layout()
# Save results
if save_plots:
fname = "figures/noise_estimation"
plt.savefig(fname + ".pdf", bbox_inches="tight", pad_inches=0)
fig.show()
```
## Effect of varying sparsity parameter
```
# Plotting parameters
plot_kwargs = {
"linewidth": 2,
}
# Determine minimum number of timesteps
end_time_idx = 100
for drop in [1, 2]:
for ball in ball_list:
ball_df = bd_data[ball].loc[bd_data[ball]["Drop #"] == drop]
h = ball_df["Height (m)"].values
t = ball_df["Time (s)"].values
ntc = helpers.need_to_clip(t)
if ntc:
h = h[:ntc]
t = t[:ntc]
if len(t) < end_time_idx:
end_time_idx = len(t)
print("Shortened to ", len(t), "for ", ball)
```
### Unregularized
```
kwargs = {
"thresh": 0.05,
"diff_method": "centered_difference",
"smoother": "savgol",
"window_length": 35,
"remove_const": False,
"drag_coeff": None,
"library_type": "polynomial",
}
ball = "Tennis Ball"
threshs = [10, 2, 0.1, 0.005, 0.0045, 0.0035, 0.002]
for thresh in threshs:
kwargs["thresh"] = thresh
print("Threshold:", thresh)
for drop in [1, 2]:
ball_df = bd_data[ball].loc[bd_data[ball]["Drop #"] == drop]
h = ball_df["Height (m)"].values
t = ball_df["Time (s)"].values
ntc = helpers.need_to_clip(t)
if ntc:
h = h[:ntc]
t = t[:ntc]
sb = helpers.SINDyBall(**kwargs)
sb.fit(h, t)
sb.print_equation()
```
### Regularized
```
kwargs = {
"thresh": 1.5,
"group_sparse_method": "1-norm",
"normalize": False,
"plot_error": False,
"library_type": "polynomial",
"model_to_subtract": None,
"test_inds": [0, end_time_idx],
}
threshs = [70, 65, 2, 0.2, 0.14, 0.1, 0.05, 0.04, 0.02, 0.01, 0.005]
for thresh in threshs:
kwargs["thresh"] = thresh
print("Threshold:", thresh)
with warnings.catch_warnings(): # Suppres ODEintWarning
warnings.filterwarnings("ignore")
helpers.test_group_sparsity(bd_data, drops=[1], plot_drop=2, **kwargs)
helpers.test_group_sparsity(bd_data, drops=[2], plot_drop=1, **kwargs)
```
## Realistic falling ball simulations
```
data_file = "../data/Ball_drops_data.xls"
bd_data = pd.read_excel(data_file, sheet_name=None)
# Remove bowling ball
del bd_data["Bowling Ball"]
ball_list = list(bd_data.keys())
# Circumference was measured in cm
circumference = {
"Baseball": 22.25,
"Blue Basketball": 75.0,
"Green Basketball": 73.25,
"Volleyball": np.NaN,
"Golf Ball": 13.8,
"Tennis Ball": 20.75,
"Whiffle Ball 1": 22.8,
"Whiffle Ball 2": 22.8,
"Yellow Whiffle Ball": 29.0,
"Orange Whiffle Ball": 29.0,
}
# Convert circumference to radius (m)
radius = {key: value / (100 * 2 * np.pi) for key, value in circumference.items()}
# Weight was measured in oz
weight = {
"Baseball": 5,
"Blue Basketball": 18,
"Green Basketball": 16,
"Volleyball": np.NaN,
"Golf Ball": 1.6,
"Tennis Ball": 2,
"Whiffle Ball 1": 1,
"Whiffle Ball 2": 1,
"Yellow Whiffle Ball": 1.5,
"Orange Whiffle Ball": 1.5,
}
# Convert weight to mass (kg)
mass_dict = {key: 0.0283495 * value for key, value in weight.items()}
```
### Generate synthetic data set
Note that one can modify the simulations by setting the following parameters of `helpers.re_dependent_synthetic_ball_drop`.
* `diameter`: Diameter of the ball (m)
* `mass`: Mass of the ball (kg)
* `timesteps`: total number of time steps
* `dt`: the size of each time step (s)
* `air_density`: density of the air (kg / m$^3$)
```
# Generate representative example balls
balls = ["Golf Ball", "Tennis Ball", "Whiffle Ball 1", "Baseball", "Blue Basketball"]
diameters = [2 * radius[ball] for ball in balls]
masses = [mass_dict[ball] for ball in balls]
noise_levels = [0, 0.01, 0.1, 0.5, 1, 2]
seed = 100
np.random.seed(seed=seed)
synthetic_data = OrderedDict()
for k, dm in enumerate(zip(diameters, masses)):
diameter, mass = dm
ball_string = "Ball " + str(k + 1)
ball_df = pd.DataFrame(columns=["Time (s)", "Height (m)", "Drop #"])
for noise in noise_levels:
h, t = helpers.re_dependent_synthetic_ball_drop(
diameter, mass=mass, noise=noise
)
temp_df = pd.DataFrame.from_dict({"Time (s)": t, "Height (m)": h})
temp_df["Drop #"] = noise
ball_df = ball_df.append(temp_df, ignore_index=True)
synthetic_data[ball_string] = ball_df
```
### Collect coefficients when group sparsity is enforced
```
kwargs = {
"thresh": 1.0,
"group_sparse_method": "1-norm",
"plot_error": False,
"library_type": "polynomial",
"window_length": 35,
}
all_coefficient_list = []
for k in range(len(noise_levels)):
all_coefficient_list.append({ball: {"1": []} for ball in synthetic_data.keys()})
with warnings.catch_warnings(): # Suppres ODEintWarning
warnings.filterwarnings("ignore")
for k, noise in enumerate(noise_levels):
gs_error = helpers.test_group_sparsity(
synthetic_data,
drops=[noise_levels[k]],
plot_drop=noise_levels[k],
all_coefficient_dict=all_coefficient_list[k],
**kwargs
)
gs_dfs = [
pd.DataFrame.from_dict(cd, orient="index").applymap(lambda l: l[0]).transpose()
for cd in all_coefficient_list
]
```
### Collect coefficients when group sparsity is not enforced
```
library = {
"1": lambda x, v: 1,
"x": lambda x, v: x,
"v": lambda x, v: v,
"xv": lambda x, v: x * v,
"x^2": lambda x, v: x ** 2,
"v^2": lambda x, v: v ** 2,
"x^2v": lambda x, v: (x ** 2) * v,
"xv^2": lambda x, v: x * (v ** 2),
"x^3": lambda x, v: x ** 3,
"v^3": lambda x, v: v ** 3,
}
kwargs = {
"thresh": 0.15,
"diff_method": "centered_difference",
"smoother": "savgol",
"window_length": 35,
"remove_const": False,
"drag_coeff": None,
"library_type": "polynomial",
}
vanilla_dfs = []
for drop in noise_levels:
df = pd.DataFrame(index=library.keys())
for ball in synthetic_data.keys():
ball_df = synthetic_data[ball].loc[synthetic_data[ball]["Drop #"] == drop]
h = ball_df["Height (m)"].values
t = ball_df["Time (s)"].values
ntc = helpers.need_to_clip(t)
if ntc:
h = h[:ntc]
t = t[:ntc]
sb = helpers.SINDyBall(**kwargs)
sb.fit(h, t)
coeffs = sb.get_xi()
labels = sb.get_labels()
coeff_series = pd.Series(coeffs, index=labels, name=ball)
df[ball] = coeff_series
vanilla_dfs.append(df)
```
### Generate coefficient heatmaps
```
# Get rid of ticks
sns.set(style="white", rc={"axes.facecolor": (0, 0, 0, 0)})
# Generate the heatmaps and put in one figure
n_tests = len(noise_levels)
title_fontsize = 18
axes_fontsize = 14
cbar_tick_labels = [0, 0.1, 1, 9.8]
cbar_ticks = [np.cbrt(l) for l in cbar_tick_labels]
cbar_tick_labels = list(map(str, cbar_tick_labels))
tex_labels = np.array(["$" + s + "$" for s in gs_dfs[0].index])
heatmap_args = {
"yticklabels": tex_labels,
"vmin": 0,
"vmax": np.cbrt(10),
"cmap": sns.cubehelix_palette(20, dark=0.3, light=1),
"linewidths": 0.1,
"linecolor": "whitesmoke",
}
fig, axs = plt.subplots(2, n_tests, figsize=(10, 7), sharey=True, sharex=True)
cbar_ax = fig.add_axes([0.91, 0.3, 0.03, 0.5])
# This heatmap's entries define the colorbar
sns.heatmap(
np.cbrt(np.abs(vanilla_dfs[0])),
ax=axs[0, 0],
cbar_ax=cbar_ax,
cbar_kws={"ticks": cbar_ticks},
**heatmap_args,
)
for k in range(1, n_tests):
sns.heatmap(
np.cbrt(np.abs(vanilla_dfs[k])), ax=axs[0, k], cbar=False, **heatmap_args
)
for k in range(n_tests):
sns.heatmap(np.cbrt(np.abs(gs_dfs[k])), ax=axs[1, k], cbar=False, **heatmap_args)
for ax in axs.flatten():
ax.tick_params(axis="y", rotation=0)
# Annotate
for k in range(n_tests):
axs[0, k].set_title(f"$\eta$ = {noise_levels[k]}", fontsize=title_fontsize)
axs[0, 0].set_ylabel("Unregularized", fontsize=axes_fontsize, labelpad=10)
axs[1, 0].set_ylabel("With group sparsity", fontsize=axes_fontsize, labelpad=10)
cbar_ax.set_yticklabels(cbar_tick_labels)
fig.tight_layout(rect=[0, 0, 0.9, 1])
if save_plots:
fname = "figures/synthetic_heatmaps_realistic_drag"
plt.savefig(fname + ".pdf", bbox_inches="tight", pad_inches=0)
plt.show()
```
### Comparison of trajectories
```
sns.reset_defaults()
pal = sns.color_palette("Set1", n_colors=4)
plot_kws = {"alpha": 0.7, "linewidth": 3}
timesteps = 900
h0 = 4000
# Get linear drag trajectory
h_l_short, t_l_short = helpers.synthetic_ball_drop(-9.8, drag=-0.125, noise=0)
h_l_long, t_l_long = helpers.synthetic_ball_drop(
-9.8, drag=-0.125, noise=0, timesteps=timesteps, h0=h0
)
# Get modified linear drag trajectory
h_ml_short, t_ml_short = helpers.synthetic_ball_drop(-12.7, drag=-0.53, noise=0)
h_ml_long, t_ml_long = helpers.synthetic_ball_drop(
-12.7, drag=-0.53, noise=0, timesteps=timesteps, h0=h0
)
# Get Re-dependent drag trajectory
h_q_short, t_q_short = helpers.re_dependent_synthetic_ball_drop(
0.067, mass=0.057, noise=0
)
h_q_long, t_q_long = helpers.re_dependent_synthetic_ball_drop(
0.067, mass=0.057, noise=0, timesteps=timesteps, h0=h0
)
fig, axs = plt.subplots(1, 2, figsize=(8, 3))
axs[0].plot(t_q_short, h_q_short, label="Re-dependent drag", color=pal[1], **plot_kws)
axs[0].plot(t_l_short, h_l_short, "--", label="Linear drag", color=pal[0], **plot_kws)
axs[0].plot(
t_ml_short, h_ml_short, ":", label="Mod. linear drag", color="black", **plot_kws
)
axs[0].set(xlabel="Time (s)", ylabel="Height (m)", title="Short drop")
axs[1].plot(t_q_long, h_q_long, label="Re-dependent drag", color=pal[1], **plot_kws)
axs[1].plot(t_l_long, h_l_long, "--", label="Linear drag", color=pal[0], **plot_kws)
axs[1].plot(
t_ml_long, h_ml_long, ":", label="Mod. linear drag", color="black", **plot_kws
)
axs[1].set(xlabel="Time (s)", title="Long drop")
axs[0].legend()
axs[1].legend()
fig.tight_layout()
if save_plots:
fname = "figures/drag_comparison"
plt.savefig(fname + ".pdf", bbox_inches="tight", pad_inches=0)
fig.show()
```
| github_jupyter |
# Inference and Validation
Now that you have a trained network, you can use it for making predictions. This is typically called **inference**, a term borrowed from statistics. However, neural networks have a tendency to perform *too well* on the training data and aren't able to generalize to data that hasn't been seen before. This is called **overfitting** and it impairs inference performance. To test for overfitting while training, we measure the performance on data not in the training set called the **validation** set. We avoid overfitting through regularization such as dropout while monitoring the validation performance during training. In this notebook, I'll show you how to do this in PyTorch.
As usual, let's start by loading the dataset through torchvision. You'll learn more about torchvision and loading data in a later part. This time we'll be taking advantage of the test set which you can get by setting `train=False` here:
```python
testset = datasets.FashionMNIST('~/.pytorch/F_MNIST_data/', download=True, train=False, transform=transform)
```
The test set contains images just like the training set. Typically you'll see 10-20% of the original dataset held out for testing and validation with the rest being used for training.
```
import torch
from torchvision import datasets, transforms
print(torch.cuda.get_device_name(0))
# Define a transform to normalize the data
transform = transforms.Compose([transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
# Download and load the training data
trainset = datasets.FashionMNIST(
'~/.pytorch/F_MNIST_data/', download=True, train=True, transform=transform)
trainloader = torch.utils.data.DataLoader(
trainset, batch_size=64, shuffle=True)
# Download and load the test data
testset = datasets.FashionMNIST(
'~/.pytorch/F_MNIST_data/', download=True, train=False, transform=transform)
testloader = torch.utils.data.DataLoader(testset, batch_size=64, shuffle=True)
```
Here I'll create a model like normal, using the same one from my solution for part 4.
```
from torch import nn, optim
import torch.nn.functional as F
class Classifier(nn.Module):
def __init__(self):
super().__init__()
self.fc1 = nn.Linear(784, 256)
self.fc2 = nn.Linear(256, 128)
self.fc3 = nn.Linear(128, 64)
self.fc4 = nn.Linear(64, 10)
def forward(self, x):
# make sure input tensor is flattened
x = x.view(x.shape[0], -1)
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = F.relu(self.fc3(x))
x = F.log_softmax(self.fc4(x), dim=1)
return x
```
The goal of validation is to measure the model's performance on data that isn't part of the training set. Performance here is up to the developer to define though. Typically this is just accuracy, the percentage of classes the network predicted correctly. Other options are [precision and recall](https://en.wikipedia.org/wiki/Precision_and_recall#Definition_(classification_context)) and top-5 error rate. We'll focus on accuracy here. First I'll do a forward pass with one batch from the test set.
```
model = Classifier()
images, labels = next(iter(testloader))
# Get the class probabilities
ps = torch.exp(model(images))
# Make sure the shape is appropriate, we should get 10 class probabilities for 64 examples
print(ps.shape)
```
With the probabilities, we can get the most likely class using the `ps.topk` method. This returns the $k$ highest values. Since we just want the most likely class, we can use `ps.topk(1)`. This returns a tuple of the top-$k$ values and the top-$k$ indices. If the highest value is the fifth element, we'll get back 4 as the index.
```
top_p, top_class = ps.topk(1, dim=1)
# Look at the most likely classes for the first 10 examples
print(top_class[:10, :], top_p[:10, :])
```
Now we can check if the predicted classes match the labels. This is simple to do by equating `top_class` and `labels`, but we have to be careful of the shapes. Here `top_class` is a 2D tensor with shape `(64, 1)` while `labels` is 1D with shape `(64)`. To get the equality to work out the way we want, `top_class` and `labels` must have the same shape.
If we do
```python
equals = top_class == labels
```
`equals` will have shape `(64, 64)`, try it yourself. What it's doing is comparing the one element in each row of `top_class` with each element in `labels` which returns 64 True/False boolean values for each row.
```
equals = top_class == labels.view(*top_class.shape)
```
Now we need to calculate the percentage of correct predictions. `equals` has binary values, either 0 or 1. This means that if we just sum up all the values and divide by the number of values, we get the percentage of correct predictions. This is the same operation as taking the mean, so we can get the accuracy with a call to `torch.mean`. If only it was that simple. If you try `torch.mean(equals)`, you'll get an error
```
RuntimeError: mean is not implemented for type torch.ByteTensor
```
This happens because `equals` has type `torch.ByteTensor` but `torch.mean` isn't implement for tensors with that type. So we'll need to convert `equals` to a float tensor. Note that when we take `torch.mean` it returns a scalar tensor, to get the actual value as a float we'll need to do `accuracy.item()`.
```
accuracy = torch.mean(equals.type(torch.FloatTensor))
print(f'Accuracy: {accuracy.item()*100}%')
```
The network is untrained so it's making random guesses and we should see an accuracy around 10%. Now let's train our network and include our validation pass so we can measure how well the network is performing on the test set. Since we're not updating our parameters in the validation pass, we can speed up our code by turning off gradients using `torch.no_grad()`:
```python
# turn off gradients
with torch.no_grad():
# validation pass here
for images, labels in testloader:
...
```
>**Exercise:** Implement the validation loop below and print out the total accuracy after the loop. You can largely copy and paste the code from above, but I suggest typing it in because writing it out yourself is essential for building the skill. In general you'll always learn more by typing it rather than copy-pasting. You should be able to get an accuracy above 80%.
```
import numpy as np
model = Classifier()
model.cuda()
criterion = nn.NLLLoss()
optimizer = optim.Adam(model.parameters(), lr=0.003)
epochs = 30
steps = 0
train_losses, test_losses = [], []
for e in range(epochs):
running_loss = 0
for images, labels in trainloader:
images = images.cuda()
labels = labels.cuda()
optimizer.zero_grad()
log_ps = model(images)
loss = criterion(log_ps, labels)
loss.backward()
optimizer.step()
running_loss += loss.item()
else:
with torch.no_grad():
valid_acc = []
valid_loss = 0
for valid_images, valid_labels in testloader:
valid_images = valid_images.cuda()
valid_labels = valid_labels.cuda()
valid_log_ps = model.forward(valid_images)
valid_ps = torch.exp(valid_log_ps)
top_p, top_class = valid_ps.topk(1, dim=1)
equals = top_class == valid_labels.view(*top_class.shape)
accuracy = torch.mean(equals.type(torch.FloatTensor))
valid_acc.append(accuracy.item())
valid_loss += criterion(valid_log_ps, valid_labels).item()
valid_acc = np.array(valid_acc).mean()
# TODO: Implement the validation pass and print out the validation accuracy
print('+'*15 + '=' * 30 + '+'*15)
print(f'Epoch: {e} \t Train Loss: {running_loss / len(trainloader):.4f} Valid Loss : {valid_loss / len(testloader):.4f} Accuracy: {valid_acc*100:.2f}%')
```
## Overfitting
If we look at the training and validation losses as we train the network, we can see a phenomenon known as overfitting.
<img src='assets/overfitting.png' width=450px>
The network learns the training set better and better, resulting in lower training losses. However, it starts having problems generalizing to data outside the training set leading to the validation loss increasing. The ultimate goal of any deep learning model is to make predictions on new data, so we should strive to get the lowest validation loss possible. One option is to use the version of the model with the lowest validation loss, here the one around 8-10 training epochs. This strategy is called *early-stopping*. In practice, you'd save the model frequently as you're training then later choose the model with the lowest validation loss.
The most common method to reduce overfitting (outside of early-stopping) is *dropout*, where we randomly drop input units. This forces the network to share information between weights, increasing it's ability to generalize to new data. Adding dropout in PyTorch is straightforward using the [`nn.Dropout`](https://pytorch.org/docs/stable/nn.html#torch.nn.Dropout) module.
```python
class Classifier(nn.Module):
def __init__(self):
super().__init__()
self.fc1 = nn.Linear(784, 256)
self.fc2 = nn.Linear(256, 128)
self.fc3 = nn.Linear(128, 64)
self.fc4 = nn.Linear(64, 10)
# Dropout module with 0.2 drop probability
self.dropout = nn.Dropout(p=0.2)
def forward(self, x):
# make sure input tensor is flattened
x = x.view(x.shape[0], -1)
# Now with dropout
x = self.dropout(F.relu(self.fc1(x)))
x = self.dropout(F.relu(self.fc2(x)))
x = self.dropout(F.relu(self.fc3(x)))
# output so no dropout here
x = F.log_softmax(self.fc4(x), dim=1)
return x
```
During training we want to use dropout to prevent overfitting, but during inference we want to use the entire network. So, we need to turn off dropout during validation, testing, and whenever we're using the network to make predictions. To do this, you use `model.eval()`. This sets the model to evaluation mode where the dropout probability is 0. You can turn dropout back on by setting the model to train mode with `model.train()`. In general, the pattern for the validation loop will look like this, where you turn off gradients, set the model to evaluation mode, calculate the validation loss and metric, then set the model back to train mode.
```python
# turn off gradients
with torch.no_grad():
# set model to evaluation mode
model.eval()
# validation pass here
for images, labels in testloader:
...
# set model back to train mode
model.train()
```
> **Exercise:** Add dropout to your model and train it on Fashion-MNIST again. See if you can get a lower validation loss or higher accuracy.
```
# TODO: Define your model with dropout added
class Classifier(nn.Module):
def __init__(self):
super().__init__()
self.fc1 = nn.Linear(784, 256)
self.fc2 = nn.Linear(256, 128)
self.fc3 = nn.Linear(128, 64)
self.fc4 = nn.Linear(64, 10)
self.dropout = nn.Dropout(0.4)
def forward(self, x):
# make sure input tensor is flattened
x = x.view(x.shape[0], -1)
x = self.dropout(F.relu(self.fc1(x)))
x = self.dropout(F.relu(self.fc2(x)))
x = self.dropout(F.relu(self.fc3(x)))
x = F.log_softmax(self.fc4(x), dim=1)
return x
# TODO: Train your model with dropout, and monitor the training progress with the validation loss and accuracy
import numpy as np
model = Classifier()
model.cuda()
criterion = nn.NLLLoss()
optimizer = optim.Adam(model.parameters(), lr=0.003)
epochs = 30
steps = 0
train_losses, test_losses = [], []
for e in range(epochs):
running_loss = 0
train_acc = []
for images, labels in trainloader:
images = images.cuda()
labels = labels.cuda()
optimizer.zero_grad()
log_ps = model(images)
ps = torch.exp(log_ps)
top_p, top_class = ps.topk(1, dim=1)
equals = top_class == labels.view(*top_class.shape)
accuracy = torch.mean(equals.type(torch.FloatTensor))
train_acc.append(accuracy.item())
loss = criterion(log_ps, labels)
loss.backward()
optimizer.step()
running_loss += loss.item()
else:
train_acc = np.array(train_acc).mean()
with torch.no_grad():
model.eval()
valid_acc = []
valid_running_loss = 0
for valid_images, valid_labels in testloader:
valid_images = valid_images.cuda()
valid_labels = valid_labels.cuda()
valid_log_ps = model.forward(valid_images)
valid_ps = torch.exp(valid_log_ps)
top_p, top_class = valid_ps.topk(1, dim=1)
equals = top_class == valid_labels.view(*top_class.shape)
accuracy = torch.mean(equals.type(torch.FloatTensor))
valid_acc.append(accuracy.item())
valid_loss = criterion(valid_log_ps, valid_labels)
valid_running_loss += valid_loss.item()
valid_acc = np.array(valid_acc).mean()
# TODO: Implement the validation pass and print out the validation accuracy
print('+' * 15 + ' ' * 5 + f'Epoch: {e}' + ' ' * 5 + '+' * 15)
print(
f'Training: Loss = {running_loss / len(trainloader):.4f} Accuracy = {train_acc*100:.2f}%')
print(
f'Validation: Loss = {valid_running_loss / len(testloader):.4f} Accuracy = {valid_acc*100:.2f}%\n')
model.train()
```
## Inference
Now that the model is trained, we can use it for inference. We've done this before, but now we need to remember to set the model in inference mode with `model.eval()`. You'll also want to turn off autograd with the `torch.no_grad()` context.
```
# Import helper module (should be in the repo)
import helper
# Test out your network!
model.eval()
dataiter = iter(testloader)
images, labels = dataiter.next()
img = images[0]
# Convert 2D image to 1D vector
img = img.view(1, 784)
# Calculate the class probabilities (softmax) for img
with torch.no_grad():
output = model.forward(img.cuda())
ps = torch.exp(output).cpu()
# Plot the image and probabilities
helper.view_classify(img.view(1, 28, 28), ps, version='Fashion')
```
## Next Up!
In the next part, I'll show you how to save your trained models. In general, you won't want to train a model everytime you need it. Instead, you'll train once, save it, then load the model when you want to train more or use if for inference.
| github_jupyter |
<img src="https://raw.githubusercontent.com/Qiskit/qiskit-tutorials/master/images/qiskit-heading.png" alt="Note: In order for images to show up in this jupyter notebook you need to select File => Trusted Notebook" width="500 px" align="left">
# Implementation of Quantum Walks on Cycle Graph
This notebook is based on the paper of B L Douglas and J B Wang, "Efficient quantum circuit implementation of quantum walks", arXiv:0706.0304 [quant-ph].
***
### Contributors
Jordan Kemp (University of Chicago), Shin Nishio (Keio University), Ryosuke Satoh (Keio University), Desiree Vogt-Lee (University of Queensland), and Tanisha Bassan (The Knowledge Society)
### Qiskit Package Versions
```
import qiskit
qiskit.__qiskit_version__
```
## Introduction
There are many different types of quantum walks: a walker can walk on n-dimensional space or any limited graphs. First we will talk about the concept and dynamics of the quantum random walk and then show the implementation of a quantum walk on cycle graph.
## Random walk
A random walk is a dynamical path with a randomly evolving time system. The figure below shows a simple type of random walk.
<img src="../images/quantum_walk/random_walk.png" alt="Note: In order for images to show up in this jupyter notebook you need to select File => Trusted Notebook" width="500 px" align="center">
The dynamics can be regarded as a simple algorithm:
1. There is a $n$-dimensional space (in this case, one for simplicity) and a walker which starts at the point $x=0$
2. The walker then takes a step either forwards (towards $+x$) or backwards (towards $-x$)
In the second step, the choice is made randomly (eg. a coin-flip). We can call this the "Coin Operator".
For this system: $p+q = 1$.
## Quantum Walk
A quantum walk is the "quantum version" of a classical random walk. This means the coin function will be a Unitary gate ($U(2)$) which is non-random and reversible:
$$p+q = U \in U(2)$$
In this notebook, we use a Hadamard gate for executing the coin function since it puts our qubits in a state of superposition, allowing for the simulation of a coin based probability:
$$H=\frac{1}{\sqrt{2}}\left [{\begin{array}{rr}1 & 1 \\ 1 & -1\\ \end{array}}\right]$$
There are two kinds of random walks, continuous and discrete, and in this notebook we will use the discrete framework. In the discrete, unitary operations are made of coin and shift operators $U = SC$ which work in a state space.
It is represented by an arbitrary undirected graph $G(V,E)$ where $V = {v_1, v_2, ..v_n}$ as nodes on the graph and $E = {(v_x, v_y) , ( v_i, v_j) …}$ as edges that combine different nodes together.
The quantum walk extends into a position space where each node $v_i$ has a certain valency $d_i$ and is split into $d_i$ subnodes. The shifting operator then acts as $S (v_i, a_i) = (v_j, a_j)$ and with the coin operator, are unitary gates which combine the probability amplitudes with individual subnodes under each node.
A unitary of $v_i$ with valency $d_i$ can be represented as $(d_i \times d_i)$. The total state of system is defined by the Hilbert space
$$H = H_c + H_p$$
Where $H_C$ is the coin Hilbert space and $H_P$ is the position Hilbert space.
## The Coin Operator
The first operation in a quantum random walk is the coin operator. The operator works by performing an arbitrary unitary transformation in the coin space which creates a rotation similar to “coin-flip” in random walk. This is namely the Hadamard gate, which models a balanced unitary coin.
The coin register will continue interfering with its position state until it is measured, after all intermediate steps. The results are very different from random walks as it doesn’t converge to a Gaussian distribution, but rather evolves into an asymmetric probability distribution. This happens because the Hadamard coin operator treats each basis vectors, |↑> and |↓> , differently.
The rightwards path interferes more destructively as it is multiplied by -1, however, the leftwards path undergoes more constructive interference and the system tends to take steps towards the left. To reach symmetric results, both base vectors will start in a superposition of states (between |↑> and |↓>). Another way to reach symmetry is use a different coin operator which doesn’t bias the coin towards a certain base vector, such as the Y gate:
$$Y=\frac{1}{\sqrt{2}}\left [{\begin{array}{rr}1 & i \\ i & 1\\ \end{array}}\right]$$
## Quantum Walk on the Cycle Graph
The goal of this notebook is to conduct a quantum random walk on circular graph which can be efficiently and simply implemented on the quantum circuit. The graph has 8 nodes with 2 attached edges which act as the subnodes on the circuit.
<img src="../images/quantum_walk/8_white.jpg" alt="Note: In order for images to show up in this jupyter notebook you need to select File => Trusted Notebook" width="350 px" align="center">
The operations propagate systemically around the graph as each node is a seperate bit-string value in lexicographic order. For a 2n graph, n qubits are required to encode the problem and 1 ancilla qubit is required for the subnode (coin).
<img src="../images/quantum_walk/whole_circuit.jpg" alt="Note: In order for images to show up in this jupyter notebook you need to select File => Trusted Notebook" width="700 px" align="center">
The above circuit shows the whole process of the quantum walk on a cycle graph with $2^3$ nodes.
The gray rectangular frame outlines a set of coin operators and shift operators.
In this circuit, q[0] to q[2] represent the state (position) of quantum walker, and q[3] represents the coin operator.
In this style, a programmer can insert the initial position of walker as a 3-qubit state. For example, if the input is $110$, then the position is $6$ (see the earlier cycle graph).
The coin operator decides whether the walker proceeds clockwise or counterclockwise.
INC is a gate that increments the state of the walker which is equal to a clockwise rotation in the cycle graph.
DEC is gate that decrements the state of the walker which is equal to a counterclockwise rotation in cycle graph.
After repeatedly executing the coin operator and the shift operator, we can measure the qubits (excluding the ancilla coin qubit), and it is then possible to know the position of the walker.
## $n$-qubit Toffoli
The Toffoli gate is a CCNOT(CCX) gate. By using the Toffoli gate, X gates executed on Q2 if Q0 and Q1 is equal to 1.
In our quantum walk implementation, we need more connections to expand the quantum walk implementation.
<img src="../images/quantum_walk/toffoli.png" alt="Note: In order for images to show up in this jupyter notebook you need to select File => Trusted Notebook" align="center">
For example, CCX can be written as in the below circuit by using only the available gate sets of the IBMQ devices.
Therefore, for more than 4 qubits, we can implement many qubits of CX gate ("C$N$X gate") using this method. Reference shown [here]("https://journals.aps.org/pra/abstract/10.1103/PhysRevA.52.3457").
<img src="../images/quantum_walk/implement_toffoli.png" alt="Note: In order for images to show up in this jupyter notebook you need to select File => Trusted Notebook" align="center">
C$N$X can be represented using C($N-1$)X as shown.
```
def cnx(qc, *qubits):
if len(qubits) >= 3:
last = qubits[-1]
# A matrix: (made up of a and Y rotation, lemma4.3)
qc.crz(np.pi/2, qubits[-2], qubits[-1])
qc.cu3(np.pi/2, 0, 0, qubits[-2],qubits[-1])
# Control not gate
cnx(qc,*qubits[:-2],qubits[-1])
# B matrix (pposite angle)
qc.cu3(-np.pi/2, 0, 0, qubits[-2], qubits[-1])
# Control
cnx(qc,*qubits[:-2],qubits[-1])
# C matrix (final rotation)
qc.crz(-np.pi/2,qubits[-2],qubits[-1])
elif len(qubits)==3:
qc.ccx(*qubits)
elif len(qubits)==2:
qc.cx(*qubits)
```
We then need to decide the number of qubits $n$ to represent the walker's state (the whole circuit requires $n+1$ qubits).
```
import numpy as np
from qiskit import IBMQ, QuantumCircuit, ClassicalRegister, QuantumRegister, execute
from qiskit.tools.visualization import plot_histogram, plot_state_city
n=3
IBMQ.load_accounts()
IBMQ.backends()
```
We then need to execute the increment and decrement gate in order for the shift operator to walk, including the C$N$X gates which changes the position of the walker based on the coin operator.
```
def increment_gate(qwc, q, subnode):
cnx(qwc, subnode[0], q[2], q[1], q[0])
cnx(qwc, subnode[0], q[2], q[1])
cnx(qwc, subnode[0], q[2])
qwc.barrier()
return qwc
def decrement_gate(qwc, q, subnode):
qwc.x(subnode[0])
qwc.x(q[2])
qwc.x(q[1])
cnx(qwc, subnode[0], q[2], q[1], q[0])
qwc.x(q[1])
cnx(qwc, subnode[0], q[2], q[1])
qwc.x(q[2])
cnx(qwc, subnode[0], q[2])
qwc.x(subnode[0])
return qwc
def ibmsim(circ):
ibmqBE = IBMQ.get_backend('ibmq_qasm_simulator')
return execute(circ,ibmqBE, shots=1000).result().get_counts(circ)
```
Rerun the coin and shift operators for n number of steps (in this case 15).
```
qnodes = QuantumRegister(n,'qc')
qsubnodes = QuantumRegister(1,'qanc')
csubnodes = ClassicalRegister(1,'canc')
cnodes = ClassicalRegister(n,'cr')
qwc = QuantumCircuit(qnodes, qsubnodes, cnodes, csubnodes)
def runQWC(qwc, times):
for i in range(times):
qwc.h(qsubnodes[0])
increment_gate(qwc, qnodes, qsubnodes[0])
decrement_gate(qwc,qnodes,qsubnodes[0])
qwc.measure(qnodes, cnodes)
return qwc
import matplotlib as mpl
step = 1
qwc = runQWC(qwc, step)
qwc.draw(output="mpl")
result = ibmsim(qwc)
print(result)
def runQWC(qwc, times):
for i in range(times):
qwc.h(qsubnodes[0])
increment_gate(qwc, qnodes, qsubnodes[0])
decrement_gate(qwc,qnodes,qsubnodes[0])
qwc.measure(qnodes, cnodes)
return qwc
```
The first qubit which is always 0, is the coin qubit.
The second to fourth, is the position of the walker (binary).
The distribution can be seen using plot_histogram.
```
result = ibmsim(qwc)
plot_histogram(result)
```
## Results
The following animation is what the quantum walk looks like over its 19 iterations. The size of each node represents probability of the quantum walker existing in that position.
<img src="../images/quantum_walk/fast.gif" alt="Note: In order for images to show up in this jupyter notebook you need to select File => Trusted Notebook" align="center">
## Required Resources
In this algorithm, we needed $n+1$ qubits for a cycle graph with $2^n$ nodes. As you can see in the following graph, the time complexity increases linearly. This is the result of relation between execution time on 'qasm_simulator' and the number of steps.
<img src="../images/quantum_walk/executiontime.png" alt="Note: In order for images to show up in this jupyter notebook you need to select File => Trusted Notebook" align="center">
## Discussion about Future Work and Applications
_Expansion of number of nodes on graph_
The walk implemented in this project required 3 qubits for 8 nodes plus an additional 1 qubit for the coin operator. The total time for iterating through coin and shift operator was 16 seconds for 100 flips.
An example of a real world problem that can be solved using quantum random walks is the mapping of enzymes to understand their evolution when in contact with mutagens. This problem only requires 33 nodes which can be mapped out on 7 qubit circuit. This increase in qubits would increase the total time to 49 seconds for 100 flips. This is a scalable model which can continue to grow to map more complex graphs to problems.
The time complexity for the quantum simulator approximately follows $({\frac{m+1}{n+1}})^2$ if the number of nodes becomes $2^m$ from $2^n$. This value is based on number of qubits and is roughly estimated.
## Conclusion
In this notebook we showed the basics of the quantum random walk and its implementation on a cyclic quantum circuit.
The implemented algorithm requires $n+1$ qubits for any cycle graph with $2^n$ nodes.
| github_jupyter |
```
import pandas as pd
import numpy as np
import seaborn as sns
from matplotlib import pyplot as plt
from matplotlib.pyplot import figure
df = pd.read_csv('billboard_top_rap¬rap_albums_lyrics_moral_profanity_emotion_sentiment_scores_clean4.csv')
df = df.loc[:,~df.columns.str.match("Unnamed")]
df = df.rename(columns={"moral_nonmoral_ratio.1": "moral_nonmoral_ratio_vv"})
```
# Variables
Variables to use
- Morality: using eMFD, we have extracted the proability of the lyrics expressing either upholding or violating each of the five moral foundations
['care.virtue', 'fairness.virtue', 'loyalty.virtue',
'authority.virtue', 'sanctity.virtue', 'care.vice', 'fairness.vice',
'loyalty.vice', 'authority.vice', 'sanctity.vice', 'moral_nonmoral_ratio_vv']
- Sentiment: sentiment is calculated using both NRCLex and Vader
[NRC >> 'positive', 'negative']
[VADER >> 'vad_neg', 'vad_pos', 'vad_neu','vad_compound']
- Emotion: emotion is calculated using NRCLEx
['anger', 'anticipation', 'disgust', 'fear', 'joy','sadness', 'surprise', 'trust',]
- Profanity: proportion of profanity words in lyrics
['profanity_prop']
- year on billboard: the year the song was on billboard
['year_billboard']
- category: whether the song is a rap or not rap
['category']
-name of the artists
['artist']
-name of the album
['title_album']
```
foundations = ['care.virtue', 'fairness.virtue', 'loyalty.virtue',
'authority.virtue', 'sanctity.virtue', 'care.vice', 'fairness.vice',
'loyalty.vice', 'authority.vice', 'sanctity.vice', 'moral_nonmoral_ratio_vv']
probabilities = ['care_p','fairness_p','loyalty_p','authority_p','sanctity_p']
sentiment = ['positive', 'negative', 'vad_neg', 'vad_pos', 'vad_neu','vad_compound']
emotion = ['anger', 'anticipation', 'disgust', 'fear', 'joy','sadness', 'surprise', 'trust',]
#dummy coding category, rap is 1
df['category_d'] = pd.get_dummies(df['category']).rap
#computing individualizing and binding index
df['individualizing_index'] = df[[ 'care.virtue', 'fairness.virtue','care.vice', 'fairness.vice']].sum(axis=1)
df['binding_index'] = df[[ 'loyalty.virtue', 'authority.virtue','sanctity.virtue', 'loyalty.vice', 'authority.vice', 'sanctity.vice', ]].sum(axis=1)
# selecting rap songs to have the time frame
print(df.groupby(by='category').year_billboard.min())
df = df[(df.year_billboard >= 2015)]
```
# Factor Analysis
```
plt.figure(figsize=(16, 6))
# define the mask to set the values in the upper triangle to True
heatmap = sns.heatmap(df[sentiment + emotion].corr(), vmin=-1, vmax=1, annot=True, cmap='BrBG')
heatmap.set_title('Correlation of sentiment and emotion', fontdict={'fontsize':18}, pad=16);
from factor_analyzer.factor_analyzer import calculate_bartlett_sphericity
chi_square_value,p_value=calculate_bartlett_sphericity(df[sentiment+emotion])
print("Bartlett's test: chi-sqaur value", chi_square_value, "p_value:", p_value)
from factor_analyzer.factor_analyzer import calculate_kmo
kmo_all,kmo_model=calculate_kmo(df[sentiment+emotion])
print("Kaiser-Meyer-Olkin Test:", kmo_model)
if p_value < 0.05 and kmo_model > .6:
print("suitable for factor analysis")
from factor_analyzer import FactorAnalyzer
# Create factor analysis object and perform factor analysis
fa = FactorAnalyzer(rotation=None)
fa.fit(df[sentiment+emotion])
# Check Eigenvalues
ev, v = fa.get_eigenvalues()
# Create scree plot using matplotlib
plt.scatter(range(1,df[sentiment+emotion].shape[1]+1),ev)
plt.plot(range(1,df[sentiment+emotion].shape[1]+1),ev)
for a,b in (zip(range(1,df[sentiment+emotion].shape[1]+1),ev)):
plt.text(a, b, str(round(b, 2)))
plt.title('Scree Plot')
plt.xlabel('Factors')
plt.ylabel('Eigenvalue')
plt.grid()
plt.show()
factor_loadings = pd.DataFrame(fa.loadings_, columns = list(np.arange(0,fa.loadings_.shape[1])))
factor_loadings.index = sentiment+emotion
factor_loadings.sort_values(by=[0], ascending=False)
for column in df.columns:
if column in sentiment+emotion:
z = df[column].apply(lambda x: (x - df[column].mean())/df[column].std())
new_column = "z_"+ column
df[new_column] = z
df['negative_connotation'] = df[['z_negative', 'z_anger', 'z_sadness', 'z_fear', 'z_disgust', 'z_vad_neg']].mean(axis =1)
df['positive_connotation'] = df[['z_joy', 'z_positive', 'vad_pos', 'z_trust', 'z_anticipation']].mean(axis =1)
```
# Descriptive Statistics
```
df.groupby(by='category').category.count()
for group, group_df in df.groupby(by=['category']):
grouped = group_df.groupby(by='title_album').category.count()
print(group, 'mean number of tracks in an album:', grouped.mean(), 'std:', grouped.std(), 'min:', grouped.min(), 'max:', grouped.max())
plt.figure(figsize=(50, 20))
# define the mask to set the values in the upper triangle to True
heatmap = sns.heatmap(df[['category_d', 'year_billboard', 'negative_connotation', 'positive_connotation', 'profanity_prop', 'individualizing_index', 'binding_index'] + foundations].corr(), vmin=-1, vmax=1, annot=True, cmap='BrBG')
heatmap.set_title('Correlation of sentiment and emotion', fontdict={'fontsize':18}, pad=16);
rap_type=df["category"]
grouped = df.groupby(rap_type)
listby_not_rap = grouped.get_group("not_rap")
rap_type=df["category"]
grouped = df.groupby(rap_type)
listby_rap = grouped.get_group("rap")
```
### Profanity
Rap music includes more profanity than non rap music
```
# "Average Profanity Prop for Rap and Not_Rap Comparison from 2000-2021"
listby_not_rap_avg_profanity_prop=listby_not_rap.groupby(['year_billboard'],as_index=False).profanity_prop.mean()
listby_rap_avg_profanity_prop=listby_rap.groupby(['year_billboard'],as_index=False).profanity_prop.mean()
plt.figure(figsize=(10,6))
plt.xlabel("Years", size=14)
plt.ylabel("Average Profanity Prop", size=14)
plt.title("Average Proportion of Profanity in Lyrcis of Billboard Top Albums from 2000-2021")
plt.bar(listby_not_rap_avg_profanity_prop['year_billboard'], listby_not_rap_avg_profanity_prop['profanity_prop'], alpha=0.5, width = 1, label="Non-Rap Song")
plt.bar(listby_rap_avg_profanity_prop['year_billboard'], listby_rap_avg_profanity_prop['profanity_prop'], alpha=0.5, width = 1, label="Rap Song")
plt.legend(loc='upper right')
#plt.savefig("overlapping_histograms_with_matplotlib_Python.png")
plt.show()
```
## Morality
```
listby_not_rap_avg=listby_not_rap.groupby(['year_billboard'],as_index=False).moral_nonmoral_ratio_vv.mean()
listby_rap_avg=listby_rap.groupby(['year_billboard'],as_index=False).moral_nonmoral_ratio_vv.mean()
plt.figure(figsize=(10,6))
plt.xlabel("Years", size=14)
plt.ylabel("Average Moral to Nonmoral Propotion", size=14)
plt.title("Average Proportion of Moral to Nonmoral Words in Lyrcis of Billboard Top Albums from 2015-2021")
plt.plot(listby_not_rap_avg['year_billboard'], listby_not_rap_avg['moral_nonmoral_ratio_vv'], label = "Non-Rap")
plt.plot(listby_rap_avg['year_billboard'], listby_rap_avg['moral_nonmoral_ratio_vv'], label = "Rap")
plt.legend()
plt.show()
emfd_f = pd.melt(df[probabilities])
emfd_s = pd.melt(pd.DataFrame(df[probabilities+['category']]), id_vars='category')['category']
emfd_source = emfd_f.join(emfd_s, rsuffix='_source')
sns.set(font_scale=1.5)
sns.set_style('white')
sns.set_style("ticks")
fig, ax = plt.subplots(figsize=(10,7))
g = sns.boxplot(x='variable', y='value', hue='category', data = emfd_source, palette='husl')
sns.despine(offset=20, trim=False)
ax.legend(loc='upper left', bbox_to_anchor=(0.1, 1),
ncol=4, fancybox=False, shadow=True)
g.set_xticklabels(['Care','Fairness','Loyalty','Authority','Sanctity'])
g.set_ylim(bottom=0,top=0.2)
g.set_title('Mean eMFD Foundation Probabilties')
g.set_ylabel('Mean Word Scores')
g.set_xlabel('')
plt.tight_layout()
emfd_f = pd.melt(df[foundations])
emfd_s = pd.melt(pd.DataFrame(df[foundations+['category']]), id_vars='category')['category']
emfd_source = emfd_f.join(emfd_s, rsuffix='_source')
sns.set(font_scale=1.5)
sns.set_style('white')
sns.set_style("ticks")
fig, ax = plt.subplots(figsize=(10,7))
g = sns.boxplot(x='variable', y='value', hue='category', data = emfd_source, palette='husl')
sns.despine(offset=20, trim=False)
ax.legend(loc='upper left', bbox_to_anchor=(0.1, 1),
ncol=4, fancybox=False, shadow=True)
g.set_xticklabels(foundations, rotation = 90)
g.set_ylim(bottom=0,top=0.2)
g.set_title('Mean eMFD Word Scores')
g.set_ylabel('Mean Word Scores')
g.set_xlabel('')
plt.tight_layout()
```
# Connotation
```
# Sentiment
not_rap_positive=listby_not_rap.groupby(['year_billboard'],as_index=False).positive_connotation.mean()
rap_positive=listby_rap.groupby(['year_billboard'],as_index=False).positive_connotation.mean()
not_rap_negative=listby_not_rap.groupby(['year_billboard'],as_index=False).negative_connotation.mean()
rap_negative=listby_rap.groupby(['year_billboard'],as_index=False).negative_connotation.mean()
plt.figure(figsize=(10,6))
plt.xlabel("Years", size=14)
plt.ylabel("Average Sentiment Score", size=14)
plt.title("Average Sentiment Using NRC Lex in Lyrcis of Billboard Top Albums from 2000-2021")
plt.plot(not_rap_positive['year_billboard'], not_rap_positive['positive_connotation'], label = "Non-Rap Positive")
plt.plot(rap_positive['year_billboard'], rap_positive['positive_connotation'], label = "Rap Positive")
plt.plot(not_rap_negative['year_billboard'], not_rap_negative['negative_connotation'], label = "Non-Rap Negative")
plt.plot(rap_negative['year_billboard'], rap_negative['negative_connotation'], label = "Rap Negative")
plt.legend()
plt.show()
```
# Inferential Statistics
```
from scipy import stats
from math import sqrt
import statsmodels.api as sm
```
# Logistic Regression
```
X = df[['year_billboard','positive_connotation','negative_connotation', 'profanity_prop']+foundations]
Y= df['category_d']
# with statsmodels
X = sm.add_constant(X) # adding a constant
model = sm.OLS(Y, X).fit()
predictions = model.predict(X)
print_model = model.summary()
print(print_model)
# VIF dataframe
vif_data = pd.DataFrame()
vif_data["feature"] = X.columns
# calculating VIF for each feature
vif_data["VIF"] = [variance_inflation_factor(X.values, i)
for i in range(len(X.columns))]
print(vif_data)
```
| github_jupyter |
# Testing with [pytest](https://docs.pytest.org/en/latest/) - part 1
## Why to write tests?
* Who wants to perform manual testing?
* When you fix a bug or add a new feature, tests are a way to verify that you did not break anything on the way
* If you have clear requirements, you can have matching test(s) for each requirement
* You don't have to be afraid of refactoring
* Tests document your implementation - they show other people use cases of your implementation
* This list is endless...
## [Test-driven development](https://en.wikipedia.org/wiki/Test-driven_development) aka TDD
In short, the basic idea of TDD is to write tests before writing the actual implementation. Maybe the most significant benefit of the approach is that the developer focuses on writing tests which match with what the program should do. Whereas if the tests are written after the actual implementation, there is a high risk for rushing tests which just show green light for the already written logic.
Tests are first class citizens in modern, agile software development, which is why it's important to start thinking TDD early during your Python learning path.
The workflow of TDD can be summarized as follows:
1. Add a test case(s) for the change / feature / bug fix you are going to implement
2. Run all tests and check that the new one fails
3. Implement required changes
4. Run tests and verify that all pass
5. Refactor
### Running pytest inside notebooks
These are the steps required to run pytest inside Jupyter cells. You can copy the content of this cell to the top of your notebook which contains tests.
```
# Let's make sure pytest and ipytest packages are installed
# ipytest is required for running pytest inside Jupyter notebooks
import sys
!{sys.executable} -m pip install pytest
!{sys.executable} -m pip install ipytest
import pytest
import ipytest
ipytest.autoconfig()
# Filename has to be set explicitly for ipytest
__file__ = 'testing1.ipynb'
```
## `pytest` test cases
Let's consider we have a function called `sum_of_three_numbers` for which we want to write a test.
```
# This would be in your e.g. implementation.py
def sum_of_three_numbers(num1, num2, num3):
return num1 + num2 + num3
```
Pytest test cases are actually quite similar as you have already seen in the exercises. Most of the exercises are structured like pytest test cases by dividing each exercise into three cells:
1. Setup the variables used in the test
2. Your implementation
3. Verify that your implementation does what is wanted by using assertions
See the example test case below to see the similarities between the exercises and common structure of test cases.
```
%%run_pytest[clean]
# Mention this at the top of cells which contain test(s)
# This is only required for running pytest in Jupyter notebooks
# This would be in your test_implementation.py
def test_sum_of_three_numbers():
# 1. Setup the variables used in the test
num1 = 2
num2 = 3
num3 = 5
# 2. Call the functionality you want to test
result = sum_of_three_numbers(num1, num2, num3)
# 3. Verify that the outcome is expected
assert result == 10
```
Now go ahead and change the line `assert result == 10` such that the assertion fails to see the output of a failed test.
| github_jupyter |
### Select points
```
import numpy as np
import matplotlib.pyplot as plt
import osgeo.gdal as gdal
import os
from maxvol_cut import rect_maxvol_cut, f_no_cut, f_penal_2D
from tools import norm_data, add_coords, gen_input, extend_score, points_selection, f_no_cut, f_cut_eps, calc_score, good_points_brute_force, idx_to_idx
%matplotlib inline
# Set BIG figure
plt.rcParams["figure.figsize"] = (12, 8)
dr = '../local_tifs'
NUMBER_OF_POINTS=16
fl_names = list(filter(lambda fl: fl.endswith('.tif'), os.listdir(dr)))
files = list(map(lambda x: gdal.Open(os.path.join(dr, x)), fl_names[::-1]))
arrays = list(map(lambda x: x.ReadAsArray().flatten(), files))
shapes = [x.ReadAsArray().shape for x in files]
nodatas = list(map(lambda x: x.GetRasterBand(1).GetNoDataValue(), files))
names = list(map(lambda x: x.replace('.tif','').split('.')[0], fl_names))
dem_raw = gdal.Open('../10_3857.tif')
dem = dem_raw.ReadAsArray()
dem_flat = dem.flatten()
dem_nodata = dem_raw.GetRasterBand(1).GetNoDataValue()
init_dem_shape = dem.shape
#delete nodata
idx_nodata_0 = np.where(arrays[0] == nodatas[0])[0]
arrays_no_nodatas = np.zeros((len(arrays[0])-len(idx_nodata_0), len(arrays)))
idx_dem_nodata = np.where(dem_flat == dem_nodata)[0]
idx_dem = np.where(arrays[0] != nodatas[0])[0]
dem_no_nodata = np.delete(dem_flat, idx_nodata_0)
for i in range(len(arrays)):
idx_nodata = np.where(arrays[i] == nodatas[i])[0]
array = arrays[i].copy()
array[idx_nodata]=0
arrays_no_nodatas[:,i] = np.delete(array, idx_nodata_0)
data_arr = arrays_no_nodatas.copy()
# Prepare data
# U can normilize data, and/or add coords to it
mode = 1 # Change to 0, 1, 2 or 3
X, fn_X_embedded = gen_input(mode, data_arr, shapes, idx_dem)
X.shape
#function for distance between points
f_cut = lambda idx, i : f_cut_eps(idx, i, X=X, eps=0.3)
#function for distence from border
f_penal = f_penal_2D(X = X[:, -2], Y = X[:, -1], bnd = 0.3, level = 0.3)
#result of selection
#if result print "Failed", the number of chosing points will be smaller than NUMBER_OF_POINTS
#to fix it, reduce eps in f_cut and bnd, level in f_penal
result = points_selection(X, n_pnts = NUMBER_OF_POINTS, cut_fun = f_cut, penalty = None)
```
### Export coordinates
To export coordinates, you should know border values of your DEM (xmin, ymin, xmax, ymax).
They could be copied from QGIS, features of layer
```
xmin, ymin, xmax, ymax = [37.7928399298814384,51.9023655604198453,37.8064010466814366,51.9077426838598441]
dem_flat_img = dem_flat.copy()-np.min(dem_flat)
dem_flat_img[np.where(dem_flat == dem_nodata)] = float('NaN')
st = dem_flat_img.reshape(init_dem_shape)
xv = np.linspace(xmin,xmax, num=st.shape[1])
yv = np.linspace(ymax,ymin, num=st.shape[0])
coords = np.meshgrid(xv,yv)
mask = idx_dem
#select corresponding points by indecies
y_c,x_c = coords[0].flatten()[mask, None],coords[1].flatten()[mask, None]
y_idx, x_idx = y_c[result],x_c[result]
coord_idx = np.hstack((y_idx,x_idx))
np.savetxt('maxvol_result_'+str(NUMBER_OF_POINTS)+'.csv', coord_idx, delimiter=';')
```
| github_jupyter |
# Linear Regression with Robust Scaler
### Required Packages
```
import warnings
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import RobustScaler
from sklearn.linear_model import LinearRegression
from sklearn.model_selection import train_test_split
from sklearn.metrics import r2_score, mean_squared_error, mean_absolute_error
```
### Initialization
Path of CSV file
```
filepath = ""
```
Features list
```
#x values
features = []
```
Target feature for prediction
```
#y value
target = ''
```
### Data Fetching
We use Pandas, open-source and most popular library in data science domain for handling data.
```
df = pd.read_csv(filepath)
df.head()
```
### Feature Selections
It is the process of reducing the number of input variables when developing a predictive model. Used to reduce the number of input variables to both reduce the computational cost of modelling and, in some cases, to improve the performance of the model.
We will assign all the required input features to X and target/outcome to Y.
```
X = df[features]
Y = df[target]
```
### Data Preprocessing
Since the majority of the machine learning models in the Sklearn library doesn't handle string category data and Null value, we have to explicitly remove or replace null values. The below snippet have functions, which removes the null value if any exists. And convert the string classes data in the datasets by encoding them to integer classes.
```
def NullClearner(df):
if(isinstance(df, pd.Series) and (df.dtype in ['int16', 'int32', 'int64', 'float16', 'float32', 'float64'])):
df.fillna(df.mean(),inplace=True)
return df
elif(isinstance(df, pd.Series)):
df.fillna(df.mode()[0],inplace=True)
return df
else:return df
def EncodeX(df):
#Here drop_first means that drop one column out of the all one-hot encoded columns. This is to prevent dummy variable trap.
return pd.get_dummies(df, drop_first=True)
```
Calling the preprocessing functions on the feature and target set.
```
x=X.columns.to_list()
for i in x:
X[i]=NullClearner(X[i])
X=EncodeX(X)
Y=NullClearner(Y)
X.head()
```
### Correlation Map
In order to check the correlation between the features, we will plot a correlation matrix. It is effective in summarizing a large amount of data where the goal is to see patterns.
```
f,ax = plt.subplots(figsize=(18, 18))
matrix = np.triu(X.corr())
sns.heatmap(X.corr(), annot=True, linewidths=.5, fmt= '.1f',ax=ax, mask=matrix)
plt.show()
```
### Data Splitting
The train-test split is a procedure for evaluating the performance of an algorithm. The procedure involves taking a dataset and dividing it into two subsets. The first subset is utilized to fit/train the model. The second subset is used for prediction. The main motive is to estimate the performance of the model on new data.
```
x_train,x_test,y_train,y_test=train_test_split(X,Y,test_size=0.2,random_state=42)
```
### Model
Linear regression algorithm attempts to model the relationship between two variables by fitting a linear equation to observed data. One variable is considered to be an independent variable, and the other is considered to be a dependent variable.
LinearRegression fits a linear model with coefficients w = (w1, …, wp) to minimize the residual sum of squares between the observed targets in the dataset, and the targets predicted by the linear approximation.
#### Robust Scaler
Scale features using statistics that are robust to outliers.
This Scaler removes the median and scales the data according to the quantile range (defaults to IQR: Interquartile Range). The IQR is the range between the 1st quartile (25th quantile) and the 3rd quartile (75th quantile).
For more detail on RobustScaler refer [API](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.RobustScaler.html)
```
Input=[("robus",RobustScaler()),("model",LinearRegression())]
model = Pipeline(Input)
model.fit(x_train,y_train)
```
#### Model Accuracy
We will use the trained model to make a prediction on the test set.Then use the predicted value for measuring the accuracy of our model.
> **score**: The **score** function returns the coefficient of determination <code>R<sup>2</sup></code> of the prediction.
```
print("Accuracy score {:.2f} %\n".format(model.score(x_test,y_test)*100))
```
> **r2_score**: The **r2_score** function computes the percentage variablility explained by our model, either the fraction or the count of correct predictions.
> **mae**: The **mean abosolute error** function calculates the amount of total error(absolute average distance between the real data and the predicted data) by our model.
> **mse**: The **mean squared error** function squares the error(penalizes the model for large errors) by our model.
```
y_pred=model.predict(x_test)
print("R2 Score: {:.2f} %".format(r2_score(y_test,y_pred)*100))
print("Mean Absolute Error {:.2f}".format(mean_absolute_error(y_test,y_pred)))
print("Mean Squared Error {:.2f}".format(mean_squared_error(y_test,y_pred)))
```
#### Prediction Plot
First, we make use of a plot to plot the actual observations, with x_train on the x-axis and y_train on the y-axis. For the regression line, we will use x_train on the x-axis and then the predictions of the x_train observations on the y-axis.
```
plt.figure(figsize=(14,10))
plt.plot(range(20),y_test[0:20], color = "green")
plt.plot(range(20),model.predict(x_test[0:20]), color = "red")
plt.legend(["Actual","prediction"])
plt.title("Predicted vs True Value")
plt.xlabel("Record number")
plt.ylabel(target)
plt.show()
```
#### Creator: Yogesh Kumar Singh, Github: [Profile](https://github.com/mohityogesh44)
| github_jupyter |
## «Дистрибутивная семантика (word2vec, GloVe, AdaGram). WMD»
### Составление словарей для классификации по тональности
При классификации текстов или предложений по тональности необходимо использовать оценочные словари для предметной области, то есть, такие словари, в которых содержатся отрицательные и позитивные слова для какой-то предметной области. Идея подобных словарей основана на следующих наблюдениях: во-первых, для разных товаров используются разные оценочные слова (например бывает “захватывающая книга”, но не бывает “захватывающих лыж”), во-вторых, в контексте разных товаров одни и те же слова могут иметь разную окраску (слово “тормоз” в отзыве на велосипед имеет нейтральную окраску, в отзыве на компьютер – резко негативную, “пыль” в контексте пылесосов – нейтральную, в контексте кофемолок – положительную (“мелкий помол в пыль”)). Еще один пример: “теплое пиво” – это плохо, а “теплый свитер” – это хорошо.
Составление таких словарей в ручную – трудоемкий процесс, но, к счастью, его не сложно автоматизировать, если собрать достаточно большие корпуса отзывов. В этом домашнем задании вам предстоит попробовать реализовать один их подходов к составлению оценочных словарей, основанный на статье [Inducing Domain-Specific Sentiment Lexicons from Unlabeled Corpora](https://nlp.stanford.edu/pubs/hamilton2016inducing.pdf).
Данные для задания – уже знакомые вам отзывы на банки, собранные с нескольких сайтов Рунета. Отзывы могут быть как положительными (оценка 5), так и отрицательными (оценка 1).
---
- Разбейте всю коллекцию отзывов на предложения. Лемматизируйте все слова.
- Обучите по коллекции предложений word2vec
- Приведите несколько удачных и неудачных примеров решения стандартных текстов для word2vec:
- тест на определение ближайших слов
- тест на аналогии (мужчина – король : женщина – королева)
- тест на определение лишнего слова.
- Постройте несколько визуализаций:
- TSNE для топ-100 (или топ-500) слов и найдите осмысленные кластеры слов
- задайте координаты для нового пространства следующим образом: одна ось описывает отношение "плохо – хорошо", вторая – "медленно – быстро" и найдите координаты названий банков в этих координатах. Более формально: берем вектор слова "хорошо", вычитаем из него вектор слова "плохо", получаем новый вектор, который описывает разницу между хорошими и плохими словами. Берем вектор слова "сбербанк" и умножаем его на этот новый вектор – получаем координату по первой оси. Аналогично – для второй оси. Две координаты уже можно нарисовать на плоскости.
---
Ссылка на примеры визуализаций: https://towardsdatascience.com/game-of-thrones-word-embeddings-does-r-l-j-part-2-30290b1c0b4b
```
import json
import bz2
import regex
import pickle
import re
import os
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from scipy import sparse
from tqdm import tqdm
from collections import Counter
from pymorphy2 import MorphAnalyzer
from pymystem3 import Mystem
from gensim.models import Word2Vec
from sklearn.manifold import TSNE
from nltk.tokenize import sent_tokenize
from nltk.corpus import stopwords
from nltk import FreqDist
from pylab import rcParams
rcParams['figure.figsize'] = 10, 8
import warnings
warnings.filterwarnings('ignore')
import nltk
nltk.download('punkt')
nltk.download('stopwords')
%matplotlib inline
%pylab inline
%config InlineBackend.figure_format = 'retina'
```
#### Разбейте всю коллекцию отзывов на предложения. Лемматизируйте все слова.
```
responses = []
with bz2.BZ2File('/Users/aleksandr/Downloads/nlp-netology-master 2/4/banki_responses.json.bz2', 'r') as thefile:
for row in tqdm(thefile):
resp = json.loads(row)
if not resp['rating_not_checked'] and (len(resp['text'].split()) > 0):
responses.append(resp)
df = pd.DataFrame(responses)
df.head(3)
# разбивка текста по предложениям
sentenses = []
for text in tqdm(df.text):
for sent in sent_tokenize(text):
sentenses.append(sent)
# предобработка текста оставляем только буквы
regex = re.compile("[А-ЯЁа-яё]+")
def words_only(text, regex=regex):
try:
return " ".join(regex.findall(text)).replace('ё','е').replace('Ё','Е')
except:
return ""
# фильтруем текст по стопам
mystopwords = stopwords.words('russian') + ['это', 'наш' , 'тыс', 'млн', 'млрд', 'также', 'т', 'д','вы', 'г']
def remove_stopwords(text, mystopwords=mystopwords):
try:
return " ".join([token for token in text.split() if not token in mystopwords])
except:
return ""
# лематизируем текст
m = Mystem()
def lemmatize(text, mystem=m):
try:
return "".join(m.lemmatize(text)).strip()
except:
return " "
# удаляем лишние лемы по стопам
mystoplemmas = stopwords.words('russian') + [
'который','прошлый','сей', 'свой', 'наш', 'мочь', 'г', 'п', 'ом', 'ч',
'з', 'р', 'е', 'эдак', 'уэк', 'уос', 'ка', 'ибо', 'орс', 'кко', 'ки']
def remove_stoplemmas(text, mystopwords=mystoplemmas):
try:
return " ".join([token for token in text.split() if not token in mystopwords])
except:
return ""
# функция запуска предобработки
def preprocessing(text):
words = words_only(text)
no_stopwords = remove_stopwords(words)
lemmas = lemmatize(no_stopwords)
no_stoplemmas = remove_stoplemmas(lemmas)
return(no_stoplemmas)
# формируем разделенные предложения из лемов в новую переменную, для дальнейшей работы
sentenses_lemma=[]
for sent in tqdm(sentenses[:]):
sentenses_lemma.append(preprocessing(sent).split())
print(sentenses_lemma[:10])
```
#### Обучите по коллекции предложений word2vec
```
%time
model_w2v = Word2Vec(
sentenses_lemma,
size=300,
window=5,
min_count=20,
workers=12,
sg=True
)
```
#### Приведите несколько удачных и неудачных примеров решения стандартных текстов для word2vec:
- тест на определение ближайших слов
- тест на аналогии (мужчина – король : женщина – королева)
- тест на определение лишнего слова.
**тест на определение ближайших слов**
```
display(model_w2v.wv.most_similar('вклад'))
display(model_w2v.wv.most_similar('кредит'))
display(model_w2v.wv.most_similar('клиент'))
```
**тест на аналогии**
```
display(model_w2v.most_similar(positive=['рубль', 'доллар'], negative=['российский']))
display(model_w2v.most_similar(positive=['заемщик', 'вклад'], negative=['кредит']))
display(model_w2v.most_similar(positive=['кредит', 'автомобиль'], negative=['квартира']))
```
**тест на определение лишнего слова**
```
display(model_w2v.doesnt_match("деньги доллар кошка".split()))
display(model_w2v.doesnt_match("стол стул деньги".split()))
display(model_w2v.doesnt_match("клиент вкладчик банк".split()))
```
#### Постройте несколько визуализаций:
- TSNE для топ-100 (или топ-500) слов и найдите осмысленные кластеры слов
- задайте координаты для нового пространства следующим образом: одна ось описывает отношение "плохо – хорошо", вторая – "медленно – быстро" и найдите координаты названий банков в этих координатах.
Более формально: берем вектор слова "хорошо", вычитаем из него вектор слова "плохо", получаем новый вектор, который описывает разницу между хорошими и плохими словами. Берем вектор слова "сбербанк" и умножаем его на этот новый вектор – получаем координату по первой оси. Аналогично – для второй оси. Две координаты уже можно нарисовать на плоскости.
Ссылка на примеры визуализаций: https://towardsdatascience.com/game-of-thrones-word-embeddings-does-r-l-j-part-2-30290b1c0b4b
**TSNE для топ-100 (или топ-500) слов и найдите осмысленные кластеры слов**
```
fd = FreqDist()
top_words = []
for text in sentenses_lemma:
fd.update(text)
for i in fd.most_common(500):
top_words.append(i[0])
print(len(top_words), '\n')
print(top_words[:100])
top_words_vec = model_w2v[top_words]
tsne = TSNE(n_components=2, random_state=0)
top_words_tsne = tsne.fit_transform(top_words_vec)
# https://docs.bokeh.org/en/latest/
from bokeh.models import ColumnDataSource, LabelSet
from bokeh.plotting import figure, show, output_file
from bokeh.io import output_notebook
output_notebook()
p = figure(
tools="pan,wheel_zoom,reset,save",
toolbar_location="above",
title="word2vec T-SNE for most common words")
source = ColumnDataSource(
data=dict(
x1=top_words_tsne[:,0],
x2=top_words_tsne[:,1],
names=top_words))
p.scatter(
x="x1",
y="x2",
size=8,
source=source)
labels = LabelSet(
x="x1",
y="x2",
text="names",
y_offset=6,
text_font_size="6pt",
text_color="#555555",
source=source,
text_align='center')
p.add_layout(labels)
show(p)
```
**Посмотрим подробнее на скопления**
```
from IPython.display import Image
Image("Screenshot1.png")
Image("Screenshot2.png")
```
**_Получились довольно логичные скопления, выше выделены некоторые из них_**
#### Задайте координаты для нового пространства следующим образом: одна ось описывает отношение "плохо – хорошо", вторая – "медленно – быстро" и найдите координаты названий банков в этих координатах.
Более формально: берем вектор слова "хорошо", вычитаем из него вектор слова "плохо", получаем новый вектор, который описывает разницу между хорошими и плохими словами. Берем вектор слова "сбербанк" и умножаем его на этот новый вектор – получаем координату по первой оси. Аналогично – для второй оси. Две координаты уже можно нарисовать на плоскости.
```
xp = model_w2v['хороший'] - model_w2v['плохо']
bm = model_w2v['быстро'] - model_w2v['медленно']
data = df.copy()
data.head()
bank_name=data.bank_name.value_counts(normalize=1).index.tolist()[:15] # топ 15
bank_name=[preprocessing(bank.lower()).split()[0] for bank in bank_name ] # предобработка
bank_name
df_bank = pd.DataFrame(bank_name, columns=['bank_name'])
df_bank['X'] = df_bank.apply(
lambda row:np.dot(model_w2v[row['bank_name']], xp), axis=1)
df_bank['Y'] = df_bank.apply(
lambda row:np.dot(model_w2v[row['bank_name']], bm), axis=1)
df_bank.head(15)
fig = plt.figure(figsize=(12, 6))
ax = fig.add_subplot(1,1,1)
for i in range(len(df_bank)):
x, y, text = df_bank.iloc[i]['X'], df_bank.iloc[i]['Y'], df_bank.iloc[i]['bank_name']
ax.scatter(x, y)
ax.text(x, y, text)
ax.set_xlabel('плохо ---> хорошо')
ax.set_ylabel('медленно ---> быстро')
plt.show()
output_notebook()
p = figure(
tools="pan,wheel_zoom,reset,save",
toolbar_location="above",
title="word2vec T-SNE for most common words")
source = ColumnDataSource(
data=dict(
x1=df_bank.X,
x2=df_bank.Y,
names=df_bank.bank_name))
p.scatter(x="x1", y="x2", size=10, source=source)
labels = LabelSet(
x="x1",
y="x2",
text="names",
y_offset=6,
text_font_size="9pt",
text_color="#555555",
source=source,
text_align='center')
p.add_layout(labels)
p.xaxis.axis_label = 'плохо ---> хорошо'
p.yaxis.axis_label = 'медленно ---> быстро'
show(p)
```
#### Итого:
- Самый быстрый **"Ренессанс"**
- Самый медленный **"Связной"**
- Самый хороший **"Авангард"**
- Самый плохой **"Восточный"**
Стоит помниль, что данные не семплированы, т.е. у 'Сбера' м.б. 1000+ отзывов,\
а у 'Восточный' 100+, что влияет на конечный результат.
```
pass
```
| github_jupyter |
# Assignment 3 Ungraded Sections - Part 2: T5 SQuAD Model
Welcome to the part 2 of testing the models for this week's assignment. This time we will perform decoding using the T5 SQuAD model. In this notebook we'll perform Question Answering by providing a "Question", its "Context" and see how well we get the "Target" answer.
## Colab
Since this ungraded lab takes a lot of time to run on coursera, as an alternative we have a colab prepared for you.
[T5 SQuAD Model Colab](https://drive.google.com/file/d/1hc7PaXjuuMS0likb0etEHY0ryAzsqAZR/view?usp=sharing)
- If you run into a page that looks similar to the one below, with the option `Open with`, this would mean you need to download the `Colaboratory` app. You can do so by `Open with -> Connect more apps -> in the search bar write "Colaboratory" -> install`
<img src = "images/colab_help_1.png">
- After installation it should look like this. Click on `Open with Google Colaboratory`
<img src = "images/colab_help_2.png">
## Outline
- [Overview](#0)
- [Part 1: Resuming the assignment (T5 SQuAD Model)](#1)
- [Part 2: Fine-tuning on SQuAD](#2)
- [2.1 Loading in the data and preprocessing](#2.1)
- [2.2 Decoding from a fine-tuned model](#2.2)
<a name='0'></a>
### Overview
In this notebook you will:
* Implement the Bidirectional Encoder Representation from Transformer (BERT) loss.
* Use a pretrained version of the model you created in the assignment for inference.
<a name='1'></a>
# Part 1: Getting ready
Run the code cells below to import the necessary libraries and to define some functions which will be useful for decoding. The code and the functions are the same as the ones you previsouly ran on the graded assignment.
```
import string
import t5
import numpy as np
import trax
from trax.supervised import decoding
import textwrap
wrapper = textwrap.TextWrapper(width=70)
PAD, EOS, UNK = 0, 1, 2
def detokenize(np_array):
return trax.data.detokenize(
np_array,
vocab_type='sentencepiece',
vocab_file='sentencepiece.model',
vocab_dir='./models/')
def tokenize(s):
return next(trax.data.tokenize(
iter([s]),
vocab_type='sentencepiece',
vocab_file='sentencepiece.model',
vocab_dir='./models/'))
vocab_size = trax.data.vocab_size(
vocab_type='sentencepiece',
vocab_file='sentencepiece.model',
vocab_dir='./models/')
def get_sentinels(vocab_size, display=False):
sentinels = {}
for i, char in enumerate(reversed(string.ascii_letters), 1):
decoded_text = detokenize([vocab_size - i])
# Sentinels, ex: <Z> - <a>
sentinels[decoded_text] = f'<{char}>'
if display:
print(f'The sentinel is <{char}> and the decoded token is:', decoded_text)
return sentinels
sentinels = get_sentinels(vocab_size, display=False)
def pretty_decode(encoded_str_list, sentinels=sentinels):
# If already a string, just do the replacements.
if isinstance(encoded_str_list, (str, bytes)):
for token, char in sentinels.items():
encoded_str_list = encoded_str_list.replace(token, char)
return encoded_str_list
# We need to decode and then prettyfy it.
return pretty_decode(detokenize(encoded_str_list))
```
<a name='2'></a>
# Part 2: Fine-tuning on SQuAD
Now let's try to fine tune on SQuAD and see what becomes of the model.For this, we need to write a function that will create and process the SQuAD `tf.data.Dataset`. Below is how T5 pre-processes SQuAD dataset as a text2text example. Before we jump in, we will have to first load in the data.
<a name='2.1'></a>
### 2.1 Loading in the data and preprocessing
You first start by loading in the dataset. The text2text example for a SQuAD example looks like:
```json
{
'inputs': 'question: <question> context: <article>',
'targets': '<answer_0>',
}
```
The squad pre-processing function takes in the dataset and processes it using the sentencePiece vocabulary you have seen above. It generates the features from the vocab and encodes the string features. It takes on question, context, and answer, and returns "question: Q context: C" as input and "A" as target.
```
# Retrieve Question, C, A and return "question: Q context: C" as input and "A" as target.
def squad_preprocess_fn(dataset, mode='train'):
return t5.data.preprocessors.squad(dataset)
# train generator, this takes about 1 minute
train_generator_fn, eval_generator_fn = trax.data.tf_inputs.data_streams(
'squad/v1.1:3.0.0',
data_dir='./data/',
bare_preprocess_fn=squad_preprocess_fn,
input_name='inputs',
target_name='targets'
)
train_generator = train_generator_fn()
next(train_generator)
#print example from train_generator
(inp, out) = next(train_generator)
print(inp.decode('utf8').split('context:')[0])
print()
print('context:', inp.decode('utf8').split('context:')[1])
print()
print('target:', out.decode('utf8'))
```
<a name='2.2'></a>
### 2.2 Decoding from a fine-tuned model
You will now use an existing model that we trained for you. You will initialize, then load in your model, and then try with your own input.
```
# Initialize the model
model = trax.models.Transformer(
d_ff = 4096,
d_model = 1024,
max_len = 2048,
n_heads = 16,
dropout = 0.1,
input_vocab_size = 32000,
n_encoder_layers = 24,
n_decoder_layers = 24,
mode='predict') # Change to 'eval' for slow decoding.
# load in the model
# this will take a minute
shape11 = trax.shapes.ShapeDtype((1, 1), dtype=np.int32)
model.init_from_file('model_squad.pkl.gz',
weights_only=True, input_signature=(shape11, shape11))
# create inputs
# a simple example
# inputs = 'question: She asked him where is john? context: John was at the game'
# an extensive example
inputs = 'question: What are some of the colours of a rose? context: A rose is a woody perennial flowering plant of the genus Rosa, in the family Rosaceae, or the flower it bears.There are over three hundred species and tens of thousands of cultivars. They form a group of plants that can be erect shrubs, climbing, or trailing, with stems that are often armed with sharp prickles. Flowers vary in size and shape and are usually large and showy, in colours ranging from white through yellows and reds. Most species are native to Asia, with smaller numbers native to Europe, North America, and northwestern Africa. Species, cultivars and hybrids are all widely grown for their beauty and often are fragrant.'
# tokenizing the input so we could feed it for decoding
print(tokenize(inputs))
test_inputs = tokenize(inputs)
```
Run the cell below to decode.
### Note: This will take some time to run
```
# Temperature is a parameter for sampling.
# # * 0.0: same as argmax, always pick the most probable token
# # * 1.0: sampling from the distribution (can sometimes say random things)
# # * values inbetween can trade off diversity and quality, try it out!
output = decoding.autoregressive_sample(model, inputs=np.array(test_inputs)[None, :],
temperature=0.0, max_length=5)
print(wrapper.fill(pretty_decode(output[0])))
```
| github_jupyter |
In this notebook we are going to use functionality provided by the SimMLA (pronounced 'Sim-L-A') package to compute the excitation irradiance profile in an epi-illumination fluorescence microscope using a fly's eye condenser. Such a condenser is realized with a pair of microlens arrays (MLA's).
```
%pylab
import SimMLA.fftpack as simfft
import SimMLA.grids as grids
plt.style.use('dark_background')
plt.rcParams['image.cmap'] = 'plasma'
```
# Setup the MLA geometry
We'll start by defining the geometry of the MLA's.
+ Prior to the objective there are two MLA's with the same parameters.
+ They are spaced by one focal length such that the second MLA is in the focal plane of the first.
+ Each lenslet has the same focal length as all the other lenslets.
```
numLenslets = 11 # Must be odd; corresponds to the number of lenslets in one dimension
lensletSize = 500 # microns
focalLength = 13700 # microns
wavelength = 0.642 # microns
```
Now we'll establish a GridArray to represent the microlenses. Such an array is a coordinate system onto which the individual lenslets are mapped. The GridArray is primarily a convenience class because it manages all unit conversions when performing Fourier transforms.
The most important parameter here is the subgridSize. This is the number of discrete lattice sites into which a lenslet will be divided. The spatial sampling period for each lenslet will therefore be
$$\delta = \frac{p}{N_{sub}}$$
where $p$ is the MLA pitch in microns and $ N_{\text{sub}} $ is the subgrid size, i.e. the number of discrete lattice sites along one dimension of the MLA. For example, a MLA pitch of $p = 500 \, \mu m$ and a subgrid size of $N_{\text{sub}} = 51$ will result in a spatial sampling period of $\frac{500 \, \mu m}{51} \approx 9.8 \mu m$.
A good sampling rate should be smaller than the fastest oscillations in the amplitude and phase of the Fourier transform of the input field. This will probably mean that the best sampling rate is half the wavelength; unfortunately, this sampling rate will probably be too high for fast numerical computations, so we'll start with a more reasonable sampling rate.
```
subgridSize = 501 # Number of grid (or lattice) sites for a single lenslet
physicalSize = numLenslets * lensletSize # The full extent of the MLA
grid = grids.GridArray(numLenslets, subgridSize, physicalSize, wavelength, focalLength)
```
# Define the input field
Next, we'll define an input laser beam that will impinge upon the first MLA.
```
power = 100 # mW
beamStd = 1000 # microns
uIn = lambda x, y: np.sqrt(power) / (2 * np. pi * beamStd**2) * np.exp(-(x**2 + y**2) / 2 / beamStd**2)
plt.imshow(uIn(grid.px, grid.py),
extent = (grid.px.min(), grid.px.max(), grid.py.min(), grid.py.max()))
plt.xlabel('x-position, microns')
plt.ylabel('y-position, microns')
plt.show()
```
# Find the field immediately after the second MLA
The field immediately *after* the second MLA is a summation of the Fourier transforms of all the fields sampled by each lenslet in the first array. Each transform is centered around the axis of its corresponding lenslet. Therefore, we have to compute a Fourier transform for each lenslet separately and then shift the origin of its coordinate axes onto the axis for the corresponding lenslet.
The reason that there is no quadratic phase curvature preceding the transforms is because the second MLA acts as a field lens, effectively canceling the quadratic phase terms.
To compute this parallelized Fourier transform, we use SimMLA's fftSubgrid routine. It will return two lists of interpolated fields, one for the magnitude and one for the phase. We will then resample this field onto a new grid representing the coordinate axes immediately after the second MLA.
*The reason we interpolate the resulting fields is because the physical units that the grid is built on change when the Fourier transform is performed. To return results based on the input grid spacing would require downsampling anyway, so I figured it would be better to let the user define the new grid spacing for their needs.*
```
import importlib
importlib.reload(simfft)
# Compute the interpolated fields.
%time interpMag, interpPhase = simfft.fftSubgrid(uIn, grid)
```
Now that we have the interpolated fields, we'll define a new grid for sampling them. We'll give the grid the same physical extent as before, but we may choose to increase the spatial sampling rate slightly. We will also use the focal length of the objective because it will serve as the final Fourier transforming lens in the system.
*If we increase the sampling rate for the new grid, it won't necessarily fix any subsampling that occurred in the previous step. It will only provide better resolution at the possibly aliased results.*
```
%%time
fObj = 3300 # microns
newGridSize = subgridSize * numLenslets # microns
newGrid = grids.Grid(newGridSize, physicalSize, wavelength, fObj)
field = np.zeros((newGrid.gridSize, newGrid.gridSize))
# For each interpolated magnitude and phase corresponding to a lenslet
# 1) Compute the full complex field
# 2) Sum it with the other complex fields
for currMag, currPhase in zip(interpMag, interpPhase):
fieldMag = currMag(np.unique(newGrid.py), np.unique(newGrid.px))
fieldPhase = currPhase(np.unique(newGrid.py), np.unique(newGrid.px))
currField = fieldMag * np.exp(1j * fieldPhase)
field = field + currField
fig, (ax0, ax1) = plt.subplots(nrows = 1, ncols = 2, sharey = True)
ax0.imshow(np.abs(field),
interpolation = 'nearest',
extent = (newGrid.px.min(), newGrid.px.max(), newGrid.py.min(), newGrid.py.max()))
ax0.set_xlabel('x-position, microns')
ax0.set_ylabel('y-position, microns')
ax1.imshow(np.angle(field),
interpolation = 'nearest',
extent = (newGrid.px.min(), newGrid.px.max(), newGrid.py.min(), newGrid.py.max()))
ax1.set_xlabel('x-position, microns')
ax1.set_axes('equal')
plt.show()
plt.imshow(np.log10(np.abs(np.fft.fftshift(np.fft.fft2(field)))))
```
| github_jupyter |
## 1. Google Play Store apps and reviews
<p>Mobile apps are everywhere. They are easy to create and can be lucrative. Because of these two factors, more and more apps are being developed. In this notebook, we will do a comprehensive analysis of the Android app market by comparing over ten thousand apps in Google Play across different categories. We'll look for insights in the data to devise strategies to drive growth and retention.</p>
<p><img src="https://assets.datacamp.com/production/project_619/img/google_play_store.png" alt="Google Play logo"></p>
<p>Let's take a look at the data, which consists of two files:</p>
<ul>
<li><code>apps.csv</code>: contains all the details of the applications on Google Play. There are 13 features that describe a given app.</li>
<li><code>user_reviews.csv</code>: contains 100 reviews for each app, <a href="https://www.androidpolice.com/2019/01/21/google-play-stores-redesigned-ratings-and-reviews-section-lets-you-easily-filter-by-star-rating/">most helpful first</a>. The text in each review has been pre-processed and attributed with three new features: Sentiment (Positive, Negative or Neutral), Sentiment Polarity and Sentiment Subjectivity.</li>
</ul>
```
# Read in dataset
import pandas as pd
apps_with_duplicates = pd.read_csv("datasets/apps.csv")
# Drop duplicates
apps = apps_with_duplicates.drop_duplicates()
# Print the total number of apps
print('Total number of apps in the dataset = ', apps["App"].count())
# Have a look at a random sample of 5 rows
n = 5
apps.sample(n)
```
## 2. Data cleaning
<p>The three features that we will be working with most frequently henceforth are <code>Installs</code>, <code>Size</code>, and <code>Price</code>. A careful glance of the dataset reveals that some of these columns mandate data cleaning in order to be consumed by code we'll write later. Specifically, the presence of special characters (<code>, $ +</code>) and letters (<code>M k</code>) in the <code>Installs</code>, <code>Size</code>, and <code>Price</code> columns make their conversion to a numerical data type difficult. Let's clean by removing these and converting each column to a numeric type.</p>
```
# List of characters to remove
chars_to_remove = ["+",",","M","$"]
# List of column names to clean
cols_to_clean = ["Installs", "Size", "Price"]
# Loop for each column
for col in cols_to_clean:
# Replace each character with an empty string
for char in chars_to_remove:
apps[col] = apps[col].str.replace(char, '')
# Convert col to numeric
apps[col] = pd.to_numeric(apps[col])
```
## 3. Exploring app categories
<p>With more than 1 billion active users in 190 countries around the world, Google Play continues to be an important distribution platform to build a global audience. For businesses to get their apps in front of users, it's important to make them more quickly and easily discoverable on Google Play. To improve the overall search experience, Google has introduced the concept of grouping apps into categories.</p>
<p>This brings us to the following questions:</p>
<ul>
<li>Which category has the highest share of (active) apps in the market? </li>
<li>Is any specific category dominating the market?</li>
<li>Which categories have the fewest number of apps?</li>
</ul>
<p>We will see that there are <code>33</code> unique app categories present in our dataset. <em>Family</em> and <em>Game</em> apps have the highest market prevalence. Interestingly, <em>Tools</em>, <em>Business</em> and <em>Medical</em> apps are also at the top.</p>
```
import plotly
plotly.offline.init_notebook_mode(connected=True)
import plotly.graph_objs as go
# Print the total number of unique categories
num_categories = len(apps["Category"].unique())
print('Number of categories = ', num_categories)
# Count the number of apps in each 'Category' and sort them in descending order
num_apps_in_category = apps["Category"].value_counts().sort_values(ascending = False)
data = [go.Bar(
x = num_apps_in_category.index, # index = category name
y = num_apps_in_category.values, # value = count
)]
plotly.offline.iplot(data)
```
## 4. Distribution of app ratings
<p>After having witnessed the market share for each category of apps, let's see how all these apps perform on an average. App ratings (on a scale of 1 to 5) impact the discoverability, conversion of apps as well as the company's overall brand image. Ratings are a key performance indicator of an app.</p>
<p>From our research, we found that the average volume of ratings across all app categories is <code>4.17</code>. The histogram plot is skewed to the right indicating that the majority of the apps are highly rated with only a few exceptions in the low-rated apps.</p>
```
# Average rating of apps
avg_app_rating = apps["Rating"].mean()
print('Average app rating = ', avg_app_rating)
# Distribution of apps according to their ratings
data = [go.Histogram(
x = apps['Rating']
)]
# Vertical dashed line to indicate the average app rating
layout = {'shapes': [{
'type' :'line',
'x0': avg_app_rating,
'y0': 0,
'x1': avg_app_rating,
'y1': 1000,
'line': { 'dash': 'dashdot'}
}]
}
plotly.offline.iplot({'data': data, 'layout': layout})
```
## 5. Size and price of an app
<p>Let's now examine app size and app price. For size, if the mobile app is too large, it may be difficult and/or expensive for users to download. Lengthy download times could turn users off before they even experience your mobile app. Plus, each user's device has a finite amount of disk space. For price, some users expect their apps to be free or inexpensive. These problems compound if the developing world is part of your target market; especially due to internet speeds, earning power and exchange rates.</p>
<p>How can we effectively come up with strategies to size and price our app?</p>
<ul>
<li>Does the size of an app affect its rating? </li>
<li>Do users really care about system-heavy apps or do they prefer light-weighted apps? </li>
<li>Does the price of an app affect its rating? </li>
<li>Do users always prefer free apps over paid apps?</li>
</ul>
<p>We find that the majority of top rated apps (rating over 4) range from 2 MB to 20 MB. We also find that the vast majority of apps price themselves under \$10.</p>
```
%matplotlib inline
import seaborn as sns
sns.set_style("darkgrid")
import warnings
warnings.filterwarnings("ignore")
# Subset for categories with at least 250 apps
large_categories = apps.groupby("Category").filter(lambda x: len(x) >= 250).reset_index()
# Plot size vs. rating
plt1 = sns.jointplot(x = large_categories["Size"], y = large_categories["Rating"], kind = 'hex')
# Subset out apps whose type is 'Paid'
paid_apps = apps[apps['Type'] == "Paid"]
# Plot price vs. rating
plt2 = sns.jointplot(x = paid_apps["Price"], y = paid_apps["Rating"])
```
## 6. Relation between app category and app price
<p>So now comes the hard part. How are companies and developers supposed to make ends meet? What monetization strategies can companies use to maximize profit? The costs of apps are largely based on features, complexity, and platform.</p>
<p>There are many factors to consider when selecting the right pricing strategy for your mobile app. It is important to consider the willingness of your customer to pay for your app. A wrong price could break the deal before the download even happens. Potential customers could be turned off by what they perceive to be a shocking cost, or they might delete an app they’ve downloaded after receiving too many ads or simply not getting their money's worth.</p>
<p>Different categories demand different price ranges. Some apps that are simple and used daily, like the calculator app, should probably be kept free. However, it would make sense to charge for a highly-specialized medical app that diagnoses diabetic patients. Below, we see that <em>Medical and Family</em> apps are the most expensive. Some medical apps extend even up to \$80! All game apps are reasonably priced below \$20.</p>
```
import matplotlib.pyplot as plt
fig, ax = plt.subplots()
fig.set_size_inches(15, 8)
# Select a few popular app categories
popular_app_cats = apps[apps.Category.isin(['GAME', 'FAMILY', 'PHOTOGRAPHY',
'MEDICAL', 'TOOLS', 'FINANCE',
'LIFESTYLE','BUSINESS'])]
# Examine the price trend by plotting Price vs Category
ax = sns.stripplot(x = popular_app_cats["Price"], y = popular_app_cats["Category"], jitter=True, linewidth=1)
ax.set_title('App pricing trend across categories')
# Apps whose Price is greater than 200
apps_above_200 = popular_app_cats[['Category', 'App', 'Price']][popular_app_cats["Price"] > 200]
apps_above_200
```
## 7. Filter out "junk" apps
<p>It looks like a bunch of the really expensive apps are "junk" apps. That is, apps that don't really have a purpose. Some app developer may create an app called <em>I Am Rich Premium</em> or <em>most expensive app (H)</em> just for a joke or to test their app development skills. Some developers even do this with malicious intent and try to make money by hoping people accidentally click purchase on their app in the store.</p>
<p>Let's filter out these junk apps and re-do our visualization. The distribution of apps under \$20 becomes clearer.</p>
```
# Select apps priced below $100
apps_under_100 = popular_app_cats[popular_app_cats["Price"] < 100]
fig, ax = plt.subplots()
fig.set_size_inches(15, 8)
# Examine price vs category with the authentic apps
ax = sns.stripplot(x="Price", y="Category", data=apps_under_100,
jitter=True, linewidth=1)
ax.set_title('App pricing trend across categories after filtering for junk apps')
```
## 8. Popularity of paid apps vs free apps
<p>For apps in the Play Store today, there are five types of pricing strategies: free, freemium, paid, paymium, and subscription. Let's focus on free and paid apps only. Some characteristics of free apps are:</p>
<ul>
<li>Free to download.</li>
<li>Main source of income often comes from advertisements.</li>
<li>Often created by companies that have other products and the app serves as an extension of those products.</li>
<li>Can serve as a tool for customer retention, communication, and customer service.</li>
</ul>
<p>Some characteristics of paid apps are:</p>
<ul>
<li>Users are asked to pay once for the app to download and use it.</li>
<li>The user can't really get a feel for the app before buying it.</li>
</ul>
<p>Are paid apps installed as much as free apps? It turns out that paid apps have a relatively lower number of installs than free apps, though the difference is not as stark as I would have expected!</p>
```
trace0 = go.Box(
# Data for paid apps
y=apps[apps['Type'] == "Paid"]['Installs'],
name = 'Paid'
)
trace1 = go.Box(
# Data for free apps
y=apps[apps['Type'] == "Free"]['Installs'],
name = 'Free'
)
layout = go.Layout(
title = "Number of downloads of paid apps vs. free apps",
yaxis = dict(
type = 'log',
autorange = True
)
)
# Add trace0 and trace1 to a list for plotting
data = [trace0, trace1]
plotly.offline.iplot({'data': data, 'layout': layout})
```
## 9. Sentiment analysis of user reviews
<p>Mining user review data to determine how people feel about your product, brand, or service can be done using a technique called sentiment analysis. User reviews for apps can be analyzed to identify if the mood is positive, negative or neutral about that app. For example, positive words in an app review might include words such as 'amazing', 'friendly', 'good', 'great', and 'love'. Negative words might be words like 'malware', 'hate', 'problem', 'refund', and 'incompetent'.</p>
<p>By plotting sentiment polarity scores of user reviews for paid and free apps, we observe that free apps receive a lot of harsh comments, as indicated by the outliers on the negative y-axis. Reviews for paid apps appear never to be extremely negative. This may indicate something about app quality, i.e., paid apps being of higher quality than free apps on average. The median polarity score for paid apps is a little higher than free apps, thereby syncing with our previous observation.</p>
<p>In this notebook, we analyzed over ten thousand apps from the Google Play Store. We can use our findings to inform our decisions should we ever wish to create an app ourselves.</p>
```
# Load user_reviews.csv
reviews_df = pd.read_csv("datasets/user_reviews.csv")
# Join and merge the two dataframe
merged_df = pd.merge(apps, reviews_df, on = "App", how = "inner")
# Drop NA values from Sentiment and Translated_Review columns
merged_df = merged_df.dropna(subset=['Sentiment', 'Translated_Review'])
sns.set_style('ticks')
fig, ax = plt.subplots()
fig.set_size_inches(11, 8)
# User review sentiment polarity for paid vs. free apps
ax = sns.boxplot(x = "Type", y = "Sentiment_Polarity", data = merged_df)
ax.set_title('Sentiment Polarity Distribution')
```
| github_jupyter |
# Deliverable D4.10
## Evaluation report on the second prototype tool for the automatic semantic description of music samples
### Evaluation of "single event" algorithm
```
import os
import sys
sys.path.append(os.path.join(os.path.dirname(os.path.realpath("__file__")), os.pardir))
from ac_utils import freesound
import pprint
import random
import pymtg
freesound_client = freesound.FreesoundClient()
freesound_client.set_token('15dab96ed5a596aaba386b2bade17c8c5a5a68a2')
'''
results_pager = freesound_client.text_search(
query="",
filter="ac_single_event:true OR ac_single_event:false",
fields="id,previews,ac_analysis,name",
page_size=150,
sort_by='rating_desc',
group_by_pack=1,
)
all_results = []
for i in range(0, 50):
print('Page ', i + 1)
for sound in results_pager:
all_results.append(sound.as_dict())
results_pager = results_pager.next_page()
print(len(all_results))
import json
random.shuffle(all_results)
json.dump(all_results, open('freesound_sounds_data.json', 'w'))
'''
data = json.load(open('freesound_sounds_data.json', 'r'))
# Some util funcitons
def get_freesound_file_path(sound_id):
pymtg.io.unicode = str # hack to avoid problem incompatiblity in Python3
base_path = '/mtgdb/incoming/freesound/sounds/{0}/'.format(sound_id // 1000)
try:
return pymtg.io.get_filenames_in_dir(dir_name=base_path, keyword='{0}_*'.format(sound_id))[0][0]
except IndexError:
print('No audio file for id {0} can be found. Does this sound exist?'.format(sound_id))
return None
def get_freesound_embed_code(sound_id, size='small'):
width, height = {
'full_size': (920, 265),
'small': (375, 30),
}[size]
return '<iframe frameborder="0" scrolling="no" src="https://freesound.org/embed/sound/iframe/{0}/simple/{1}/" width="{2}" height="{3}"></iframe>'.format(sound_id, size, width, height)
def diplay_sound_widget(sid):
print('*', sid)
display(Audio(get_freesound_file_path(sid)))
from IPython.display import display, Markdown, HTML, Audio
from ipywidgets import interact_manual, interact
import random
import ipywidgets as widgets
N_PER_BATCH = 50
if False:
try:
already_in_gt = list(json.load(open('ground_truth.json')).keys())
except FileNotFoundError:
already_in_gt = list()
def on_change(change):
if change['type'] == 'change' and change['name'] == 'value':
try:
gt = json.load(open('ground_truth.json'))
except FileNotFoundError:
gt = {}
gt[change['owner']._sid] = change['new']
json.dump(gt, open('ground_truth.json', 'w'))
already_in_gt = list(gt.keys())
for sound in [s for s in data if str(s['id']) not in already_in_gt][:N_PER_BATCH]:
diplay_sound_widget(sound['id'])
buttons = widgets.RadioButtons(
options=['skip', 'no', 'yes'],
description='Is single event:'
)
buttons._sid = sound['id']
buttons.observe(on_change)
display(buttons)
print('\n\n')
gt = json.load(open('ground_truth.json'))
tp = [] # true positive (sound is single event and detected correctly) - YAY
fp = [] # false positive (sound detected as single_event but it is not) - OUCH
tn = [] # true negative (sound detected as not single_event and it is not) - YAY
fn = [] # true positive (sound detected as not single event but it is) - OUCH
data_dict = {str(item['id']):item for item in data}
for sound_id, sound_gt in gt.items():
if sound_gt == 'skip':
continue
sound = data_dict[sound_id]
if sound_gt == 'yes' and sound['ac_analysis']['ac_single_event'] == True:
tp.append(sound)
elif sound_gt == 'no' and sound['ac_analysis']['ac_single_event'] == True:
fp.append(sound)
elif sound_gt == 'yes' and sound['ac_analysis']['ac_single_event'] == False:
fn.append(sound)
elif sound_gt == 'no' and sound['ac_analysis']['ac_single_event'] == False:
tn.append(sound)
total = len(tp) + len(tn) + len(fp) + len(fn)
print('Size GT:', total)
print('TP:', len(tp)/total)
print('TN:', len(tn)/total)
print('FP:', len(fp)/total)
print('FN:', len(fn)/total)
print('Accuracy:', (len(tp) + len(tn))/total)
# Optimization of parameters for single event detection
# STEP1 : define imports and util functions
import json
import essentia
essentia.log.infoActive = False
essentia.log.warningActive = False
from essentia.standard import MonoLoader
import numpy as np
import pymtg
def estimate_number_of_events(audiofile, region_energy_thr=2, silence_thr_scale=4, group_regions_ms=100):
"""
Returns list of activity "onsets" for an audio signal based on its energy envelope.
This is more like "activity detecton" than "onset detection".
"""
def group_regions(regions, group_regions_ms):
"""
Group together regions which are very close in time (i.e. the end of a region is very close to the start of the following).
"""
if len(regions) <= 1:
grouped_regions = regions[:] # Don't do anything if only one region or no regions at all
else:
# Iterate over regions and mark which regions should be grouped with the following regions
to_group = []
for count, ((at0, at1, a_energy), (bt0, bt1, b_energy)) in enumerate(zip(regions[:-1], regions[1:])):
if bt0 - at1 < group_regions_ms / 1000:
to_group.append(1)
else:
to_group.append(0)
to_group.append(0) # Add 0 for the last one which will never be grouped with next (there is no "next region")
# Now generate the grouped list of regions based on the marked ones in 'to_group'
grouped_regions = []
i = 0
while i < len(to_group):
current_group_start = None
current_group_end = None
x = to_group[i]
if x == 1 and current_group_start is None:
# Start current grouping
current_group_start = i
while x == 1:
i += 1
x = to_group[i]
current_group_end = i
grouped_regions.append( (regions[current_group_start][0], regions[current_group_end][1], sum([z for x,y,z in regions[current_group_start:current_group_end+1]])))
current_group_start = None
current_group_end = None
else:
grouped_regions.append(regions[i])
i += 1
return grouped_regions
# Load audio file
sample_rate = 44100
audio_file = MonoLoader(filename=audiofile, sampleRate=sample_rate)
audio = audio_file.compute()
t = np.linspace(0, len(audio)/sample_rate, num=len(audio))
# Compute envelope and average signal energy
env_algo = essentia.standard.Envelope(
attackTime = 15,
releaseTime = 50,
)
envelope = env_algo(audio)
average_signal_energy = np.sum(np.array(envelope)**2)/len(envelope)
silence_thr = average_signal_energy * silence_thr_scale
# Get energy regions above threshold
# Implementation based on https://stackoverflow.com/questions/43258896/extract-subarrays-of-numpy-array-whose-values-are-above-a-threshold
mask = np.concatenate(([False], envelope > silence_thr, [False] ))
idx = np.flatnonzero(mask[1:] != mask[:-1])
idx -= 1 # Avoid index out of bounds (0-index)
regions = [(t[idx[i]], t[idx[i+1]], np.sum(envelope[idx[i]:idx[i+1]]**2)) for i in range(0, len(idx), 2)] # Energy is a list of tuples like (start_time, end_time, energy)
regions = [region for region in regions if region[2] > region_energy_thr] # Discard those below region_energy_thr
# Group detected regions that happen close together
regions = group_regions(regions, group_regions_ms)
return len(regions) # Return number of sound events detected
def get_freesound_file_path(sound_id):
pymtg.io.unicode = str # hack to avoid problem incompatiblity in Python3
base_path = '/mtgdb/incoming/freesound/sounds/{0}/'.format(sound_id // 1000)
try:
return pymtg.io.get_filenames_in_dir(dir_name=base_path, keyword='{0}_*'.format(sound_id))[0][0]
except IndexError:
print('No audio file for id {0} can be found. Does this sound exist?'.format(sound_id))
return None
def get_analysis_params_key(region_energy_thr, silence_thr_scale, group_regions_ms, max_duration):
return '{0}-{1}-{2}-{3}'.format(region_energy_thr, silence_thr_scale, group_regions_ms, max_duration)
# STEP 2: define variables
gt = {sound_id: sound_gt for sound_id, sound_gt in json.load(open('ground_truth.json')).items() if sound_gt != 'skip'}
analysis_results = {}
# STEP 3: run analysis
#analysis_results = json.load(open('analysis_results_0_8_0_8_0_200_1.json'))
wp = pymtg.processing.WorkParallelizer()
def analyze(sound_id, anlalysis_params_key, sound_path, region_energy_thr, silence_thr_scale, group_regions_ms, max_duration):
n_audio_events = estimate_number_of_events(sound_path, region_energy_thr, silence_thr_scale, group_regions_ms)
duration = len(MonoLoader(filename=sound_path, sampleRate=44100).compute())/44100
single_event = n_audio_events == 1
if single_event and max_duration is not None:
if duration > max_duration:
single_event = False
return (sound_id, anlalysis_params_key, single_event)
for (region_energy_thr, silence_thr_scale, group_regions_ms) in [(2, 4, 100), (0.5, 4.5, 50)]:
for max_duration in [11, 12, 13, 14, 15, 16]:#np.arange(5, 10, 1):
for sound_id, sound_gt in gt.items():
sound_path = get_freesound_file_path(int(sound_id))
anlalysis_params_key = get_analysis_params_key(region_energy_thr, silence_thr_scale, group_regions_ms, max_duration)
if anlalysis_params_key not in analysis_results[sound_id]:
wp.add_task(analyze, sound_id, anlalysis_params_key, sound_path, region_energy_thr, silence_thr_scale, group_regions_ms, max_duration)
wp.run(num_workers=4)
# Show errors after computing all tasks (if any)
if wp.num_tasks_failed > 0:
wp.show_errors()
# Store results in analysis dict
for t in wp.tasks_succeeded():
sound_id, anlalysis_params_key, result = t.result()
if sound_id not in analysis_results:
analysis_results[sound_id] = {}
analysis_results[sound_id][anlalysis_params_key] = result
# STEP 4: compute evaluation metrics
accuracies = list()
for anlalysis_params_key in list(analysis_results.values())[0].keys():
tp = [] # true positive (sound is single event and detected correctly) - YAY
fp = [] # false positive (sound detected as single_event but it is not) - OUCH
tn = [] # true negative (sound detected as not single_event and it is not) - YAY
fn = [] # true positive (sound detected as not single event but it is) - OUCH
for sound_id, sound_gt in gt.items():
result = analysis_results[sound_id][anlalysis_params_key]
if sound_gt == 'yes' and result == True:
tp.append(sound_id)
elif sound_gt == 'no' and result == True:
fp.append(sound_id)
elif sound_gt == 'yes' and result == False:
fn.append(sound_id)
elif sound_gt == 'no' and result == False:
tn.append(sound_id)
total = len(tp) + len(tn) + len(fp) + len(fn)
accuracies.append((anlalysis_params_key, (len(tp) + len(tn))/total))
if False:
print(anlalysis_params_key)
print('- Size GT:', total)
print('- TP:', len(tp)/total)
print('- TN:', len(tn)/total)
print('- FP:', len(fp)/total)
print('- FN:', len(fn)/total)
print('- Accuracy:', (len(tp) + len(tn))/total)
print('')
if False:
for sound_id in fp[0:5]:
diplay_sound_widget(int(sound_id))
print('\nSUMMARY:')
for name, accuracy in sorted(accuracies, key=lambda x: x[1], reverse=True)[0:40]:
print(name, '\t', accuracy)
```
# Info for deliverable:
Previous method:
2-4-100-None
- Size GT: 224
- TP: 0.24553571428571427
- TN: 0.39732142857142855
- FP: 0.26785714285714285
- FN: 0.08928571428571429
- Accuracy: 0.6428571428571429
Method with optimized parameters:
0.5-4.5-50-None
- Size GT: 224
- TP: 0.24107142857142858
- TN: 0.44642857142857145
- FP: 0.21875
- FN: 0.09375
- Accuracy: 0.6875
Method filtering by duration:
0.5-4.5-50-7
- Size GT: 224
- TP: 0.21875
- TN: 0.5982142857142857
- FP: 0.06696428571428571
- FN: 0.11607142857142858
- Accuracy: 0.8169642857142857
```
accuracies_with_filter = [] #[0.8080357142857143, 0.8080357142857143, 0.8169642857142857, 0.8125, 0.7991071428571429, 0.7901785714285714]
durations = [1, 2, 3, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14]
for duration in durations:
for acc in accuracies:
if acc[0] == '0.5-4.5-50-{0}'.format(duration):
accuracies_with_filter.append(acc[1])
break
import matplotlib.pyplot as plt
%matplotlib inline
plt.plot(durations, accuracies_with_filter, marker='o')
plt.title('single_event detection accuracy')
plt.xlabel('Druation filter (s)')
plt.axis([0.5, 14.5, 0.5, 1])
plt.grid(True)
plt.show()
```
| github_jupyter |
# GCP Dataflow Component Sample
A Kubeflow Pipeline component that prepares data by submitting an Apache Beam job (authored in Python) to Cloud Dataflow for execution. The Python Beam code is run with Cloud Dataflow Runner.
## Intended use
Use this component to run a Python Beam code to submit a Cloud Dataflow job as a step of a Kubeflow pipeline.
## Runtime arguments
Name | Description | Optional | Data type| Accepted values | Default |
:--- | :----------| :----------| :----------| :----------| :---------- |
python_file_path | The path to the Cloud Storage bucket or local directory containing the Python file to be run. | | GCSPath | | |
project_id | The ID of the Google Cloud Platform (GCP) project containing the Cloud Dataflow job.| | String | | |
region | The Google Cloud Platform (GCP) region to run the Cloud Dataflow job.| | String | | |
staging_dir | The path to the Cloud Storage directory where the staging files are stored. A random subdirectory will be created under the staging directory to keep the job information.This is done so that you can resume the job in case of failure. `staging_dir` is passed as the command line arguments (`staging_location` and `temp_location`) of the Beam code. | Yes | GCSPath | | None |
requirements_file_path | The path to the Cloud Storage bucket or local directory containing the pip requirements file. | Yes | GCSPath | | None |
args | The list of arguments to pass to the Python file. | No | List | A list of string arguments | None |
wait_interval | The number of seconds to wait between calls to get the status of the job. | Yes | Integer | | 30 |
## Input data schema
Before you use the component, the following files must be ready in a Cloud Storage bucket:
- A Beam Python code file.
- A `requirements.txt` file which includes a list of dependent packages.
The Beam Python code should follow the [Beam programming guide](https://beam.apache.org/documentation/programming-guide/) as well as the following additional requirements to be compatible with this component:
- It accepts the command line arguments `--project`, `--region`, `--temp_location`, `--staging_location`, which are [standard Dataflow Runner options](https://cloud.google.com/dataflow/docs/guides/specifying-exec-params#setting-other-cloud-pipeline-options).
- It enables `info logging` before the start of a Cloud Dataflow job in the Python code. This is important to allow the component to track the status and ID of the job that is created. For example, calling `logging.getLogger().setLevel(logging.INFO)` before any other code.
## Output
Name | Description
:--- | :----------
job_id | The id of the Cloud Dataflow job that is created.
## Cautions & requirements
To use the components, the following requirements must be met:
- Cloud Dataflow API is enabled.
- The component is running under a secret Kubeflow user service account in a Kubeflow Pipeline cluster. For example:
```
component_op(...)
```
The Kubeflow user service account is a member of:
- `roles/dataflow.developer` role of the project.
- `roles/storage.objectViewer` role of the Cloud Storage Objects `python_file_path` and `requirements_file_path`.
- `roles/storage.objectCreator` role of the Cloud Storage Object `staging_dir`.
## Detailed description
The component does several things during the execution:
- Downloads `python_file_path` and `requirements_file_path` to local files.
- Starts a subprocess to launch the Python program.
- Monitors the logs produced from the subprocess to extract the Cloud Dataflow job information.
- Stores the Cloud Dataflow job information in `staging_dir` so the job can be resumed in case of failure.
- Waits for the job to finish.
# Setup
```
project = 'Input your PROJECT ID'
region = 'Input GCP region' # For example, 'us-central1'
output = 'Input your GCS bucket name' # No ending slash
```
## Install Pipeline SDK
```
!python3 -m pip install 'kfp>=0.1.31' --quiet
```
## Load the component using KFP SDK
```
import kfp.deprecated.components as comp
dataflow_python_op = comp.load_component_from_url(
'https://raw.githubusercontent.com/kubeflow/pipelines/1.7.0-rc.3/components/gcp/dataflow/launch_python/component.yaml')
help(dataflow_python_op)
```
## Use the wordcount python sample
In this sample, we run a wordcount sample code in a Kubeflow Pipeline. The output will be stored in a Cloud Storage bucket. Here is the sample code:
```
!gsutil cat gs://ml-pipeline/sample-pipeline/word-count/wc.py
```
## Example pipeline that uses the component
```
import kfp.deprecated as kfp
from kfp.deprecated import dsl, Client
import json
@dsl.pipeline(
name='dataflow-launch-python-pipeline',
description='Dataflow launch python pipeline'
)
def pipeline(
python_file_path = 'gs://ml-pipeline/sample-pipeline/word-count/wc.py',
project_id = project,
region = region,
staging_dir = output,
requirements_file_path = 'gs://ml-pipeline/sample-pipeline/word-count/requirements.txt',
wait_interval = 30
):
dataflow_python_op(
python_file_path = python_file_path,
project_id = project_id,
region = region,
staging_dir = staging_dir,
requirements_file_path = requirements_file_path,
args = json.dumps(['--output', f'{staging_dir}/wc/wordcount.out']),
wait_interval = wait_interval)
```
## Submit the pipeline for execution
```
Client().create_run_from_pipeline_func(pipeline, arguments={})
```
#### Inspect the output
```
!gsutil cat $output/wc/wordcount.out
```
## References
* [Component python code](https://github.com/kubeflow/pipelines/blob/master/components/gcp/container/component_sdk/python/kfp_component/google/dataflow/_launch_python.py)
* [Component docker file](https://github.com/kubeflow/pipelines/blob/master/components/gcp/container/Dockerfile)
* [Sample notebook](https://github.com/kubeflow/pipelines/blob/master/components/gcp/dataflow/launch_python/sample.ipynb)
* [Dataflow Python Quickstart](https://cloud.google.com/dataflow/docs/quickstarts/quickstart-python)
| github_jupyter |
##### Copyright © 2019 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# TFX Estimator Component Tutorial
***A Component-by-Component Introduction to TensorFlow Extended (TFX)***
Note: We recommend running this tutorial in a Colab notebook, with no setup required! Just click "Run in Google Colab".
<div class="devsite-table-wrapper"><table class="tfo-notebook-buttons" align="left">
<td><a target="_blank" href="https://www.tensorflow.org/tfx/tutorials/tfx/components">
<img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a></td>
<td><a target="_blank" href="https://colab.research.google.com/github/tensorflow/tfx/blob/master/docs/tutorials/tfx/components.ipynb">
<img src="https://www.tensorflow.org/images/colab_logo_32px.png">Run in Google Colab</a></td>
<td><a target="_blank" href="https://github.com/tensorflow/tfx/tree/master/docs/tutorials/tfx/components.ipynb">
<img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png">View source on GitHub</a></td>
</table></div>
This Colab-based tutorial will interactively walk through each built-in component of TensorFlow Extended (TFX).
It covers every step in an end-to-end machine learning pipeline, from data ingestion to pushing a model to serving.
When you're done, the contents of this notebook can be automatically exported as TFX pipeline source code, which you can orchestrate with Apache Airflow and Apache Beam.
Note: This notebook and its associated APIs are **experimental** and are
in active development. Major changes in functionality, behavior, and
presentation are expected.
## Background
This notebook demonstrates how to use TFX in a Jupyter/Colab environment. Here, we walk through the Chicago Taxi example in an interactive notebook.
Working in an interactive notebook is a useful way to become familiar with the structure of a TFX pipeline. It's also useful when doing development of your own pipelines as a lightweight development environment, but you should be aware that there are differences in the way interactive notebooks are orchestrated, and how they access metadata artifacts.
### Orchestration
In a production deployment of TFX, you will use an orchestrator such as Apache Airflow, Kubeflow Pipelines, or Apache Beam to orchestrate a pre-defined pipeline graph of TFX components. In an interactive notebook, the notebook itself is the orchestrator, running each TFX component as you execute the notebook cells.
### Metadata
In a production deployment of TFX, you will access metadata through the ML Metadata (MLMD) API. MLMD stores metadata properties in a database such as MySQL or SQLite, and stores the metadata payloads in a persistent store such as on your filesystem. In an interactive notebook, both properties and payloads are stored in an ephemeral SQLite database in the `/tmp` directory on the Jupyter notebook or Colab server.
## Setup
First, we install and import the necessary packages, set up paths, and download data.
### Upgrade Pip
To avoid upgrading Pip in a system when running locally, check to make sure that we're running in Colab. Local systems can of course be upgraded separately.
```
try:
import colab
!pip install --upgrade pip
except:
pass
```
### Install TFX
**Note: In Google Colab, because of package updates, the first time you run this cell you must restart the runtime (Runtime > Restart runtime ...).**
```
# TODO(ccy): Fix need to use deprecated pip resolver.
!pip install -q -U --use-deprecated=legacy-resolver tfx
```
## Did you restart the runtime?
If you are using Google Colab, the first time that you run the cell above, you must restart the runtime (Runtime > Restart runtime ...). This is because of the way that Colab loads packages.
### Import packages
We import necessary packages, including standard TFX component classes.
```
import os
import pprint
import tempfile
import urllib
import absl
import tensorflow as tf
import tensorflow_model_analysis as tfma
tf.get_logger().propagate = False
pp = pprint.PrettyPrinter()
import tfx
from tfx.components import CsvExampleGen
from tfx.components import Evaluator
from tfx.components import ExampleValidator
from tfx.components import Pusher
from tfx.components import ResolverNode
from tfx.components import SchemaGen
from tfx.components import StatisticsGen
from tfx.components import Trainer
from tfx.components import Transform
from tfx.dsl.experimental import latest_blessed_model_resolver
from tfx.orchestration import metadata
from tfx.orchestration import pipeline
from tfx.orchestration.experimental.interactive.interactive_context import InteractiveContext
from tfx.proto import pusher_pb2
from tfx.proto import trainer_pb2
from tfx.proto.evaluator_pb2 import SingleSlicingSpec
from tfx.utils.dsl_utils import external_input
from tfx.types import Channel
from tfx.types.standard_artifacts import Model
from tfx.types.standard_artifacts import ModelBlessing
%load_ext tfx.orchestration.experimental.interactive.notebook_extensions.skip
```
Let's check the library versions.
```
print('TensorFlow version: {}'.format(tf.__version__))
# TODO(ccy): Revert to `tfx.__version__` after 0.27.0 release.
print('TFX version: {}'.format(__import__('tfx.version').__version__))
```
### Set up pipeline paths
```
# This is the root directory for your TFX pip package installation.
_tfx_root = tfx.__path__[0]
# This is the directory containing the TFX Chicago Taxi Pipeline example.
_taxi_root = os.path.join(_tfx_root, 'examples/chicago_taxi_pipeline')
# This is the path where your model will be pushed for serving.
_serving_model_dir = os.path.join(
tempfile.mkdtemp(), 'serving_model/taxi_simple')
# Set up logging.
absl.logging.set_verbosity(absl.logging.INFO)
```
### Download example data
We download the example dataset for use in our TFX pipeline.
The dataset we're using is the [Taxi Trips dataset](https://data.cityofchicago.org/Transportation/Taxi-Trips/wrvz-psew) released by the City of Chicago. The columns in this dataset are:
<table>
<tr><td>pickup_community_area</td><td>fare</td><td>trip_start_month</td></tr>
<tr><td>trip_start_hour</td><td>trip_start_day</td><td>trip_start_timestamp</td></tr>
<tr><td>pickup_latitude</td><td>pickup_longitude</td><td>dropoff_latitude</td></tr>
<tr><td>dropoff_longitude</td><td>trip_miles</td><td>pickup_census_tract</td></tr>
<tr><td>dropoff_census_tract</td><td>payment_type</td><td>company</td></tr>
<tr><td>trip_seconds</td><td>dropoff_community_area</td><td>tips</td></tr>
</table>
With this dataset, we will build a model that predicts the `tips` of a trip.
```
_data_root = tempfile.mkdtemp(prefix='tfx-data')
DATA_PATH = 'https://raw.githubusercontent.com/tensorflow/tfx/master/tfx/examples/chicago_taxi_pipeline/data/simple/data.csv'
_data_filepath = os.path.join(_data_root, "data.csv")
urllib.request.urlretrieve(DATA_PATH, _data_filepath)
```
Take a quick look at the CSV file.
```
!head {_data_filepath}
```
*Disclaimer: This site provides applications using data that has been modified for use from its original source, www.cityofchicago.org, the official website of the City of Chicago. The City of Chicago makes no claims as to the content, accuracy, timeliness, or completeness of any of the data provided at this site. The data provided at this site is subject to change at any time. It is understood that the data provided at this site is being used at one’s own risk.*
### Create the InteractiveContext
Last, we create an InteractiveContext, which will allow us to run TFX components interactively in this notebook.
```
# Here, we create an InteractiveContext using default parameters. This will
# use a temporary directory with an ephemeral ML Metadata database instance.
# To use your own pipeline root or database, the optional properties
# `pipeline_root` and `metadata_connection_config` may be passed to
# InteractiveContext. Calls to InteractiveContext are no-ops outside of the
# notebook.
context = InteractiveContext()
```
## Run TFX components interactively
In the cells that follow, we create TFX components one-by-one, run each of them, and visualize their output artifacts.
### ExampleGen
The `ExampleGen` component is usually at the start of a TFX pipeline. It will:
1. Split data into training and evaluation sets (by default, 2/3 training + 1/3 eval)
2. Convert data into the `tf.Example` format
3. Copy data into the `_tfx_root` directory for other components to access
`ExampleGen` takes as input the path to your data source. In our case, this is the `_data_root` path that contains the downloaded CSV.
Note: In this notebook, we can instantiate components one-by-one and run them with `InteractiveContext.run()`. By contrast, in a production setting, we would specify all the components upfront in a `Pipeline` to pass to the orchestrator (see the [Building a TFX Pipeline Guide](../tfx/guide/build_tfx_pipeline)).
```
example_gen = CsvExampleGen(input=external_input(_data_root))
context.run(example_gen)
```
Let's examine the output artifacts of `ExampleGen`. This component produces two artifacts, training examples and evaluation examples:
```
artifact = example_gen.outputs['examples'].get()[0]
print(artifact.split_names, artifact.uri)
```
We can also take a look at the first three training examples:
```
# Get the URI of the output artifact representing the training examples, which is a directory
train_uri = os.path.join(example_gen.outputs['examples'].get()[0].uri, 'train')
# Get the list of files in this directory (all compressed TFRecord files)
tfrecord_filenames = [os.path.join(train_uri, name)
for name in os.listdir(train_uri)]
# Create a `TFRecordDataset` to read these files
dataset = tf.data.TFRecordDataset(tfrecord_filenames, compression_type="GZIP")
# Iterate over the first 3 records and decode them.
for tfrecord in dataset.take(3):
serialized_example = tfrecord.numpy()
example = tf.train.Example()
example.ParseFromString(serialized_example)
pp.pprint(example)
```
Now that `ExampleGen` has finished ingesting the data, the next step is data analysis.
### StatisticsGen
The `StatisticsGen` component computes statistics over your dataset for data analysis, as well as for use in downstream components. It uses the [TensorFlow Data Validation](https://www.tensorflow.org/tfx/data_validation/get_started) library.
`StatisticsGen` takes as input the dataset we just ingested using `ExampleGen`.
```
statistics_gen = StatisticsGen(
examples=example_gen.outputs['examples'])
context.run(statistics_gen)
```
After `StatisticsGen` finishes running, we can visualize the outputted statistics. Try playing with the different plots!
```
context.show(statistics_gen.outputs['statistics'])
```
### SchemaGen
The `SchemaGen` component generates a schema based on your data statistics. (A schema defines the expected bounds, types, and properties of the features in your dataset.) It also uses the [TensorFlow Data Validation](https://www.tensorflow.org/tfx/data_validation/get_started) library.
`SchemaGen` will take as input the statistics that we generated with `StatisticsGen`, looking at the training split by default.
```
schema_gen = SchemaGen(
statistics=statistics_gen.outputs['statistics'],
infer_feature_shape=False)
context.run(schema_gen)
```
After `SchemaGen` finishes running, we can visualize the generated schema as a table.
```
context.show(schema_gen.outputs['schema'])
```
Each feature in your dataset shows up as a row in the schema table, alongside its properties. The schema also captures all the values that a categorical feature takes on, denoted as its domain.
To learn more about schemas, see [the SchemaGen documentation](https://www.tensorflow.org/tfx/guide/schemagen).
### ExampleValidator
The `ExampleValidator` component detects anomalies in your data, based on the expectations defined by the schema. It also uses the [TensorFlow Data Validation](https://www.tensorflow.org/tfx/data_validation/get_started) library.
`ExampleValidator` will take as input the statistics from `StatisticsGen`, and the schema from `SchemaGen`.
```
example_validator = ExampleValidator(
statistics=statistics_gen.outputs['statistics'],
schema=schema_gen.outputs['schema'])
context.run(example_validator)
```
After `ExampleValidator` finishes running, we can visualize the anomalies as a table.
```
context.show(example_validator.outputs['anomalies'])
```
In the anomalies table, we can see that there are no anomalies. This is what we'd expect, since this the first dataset that we've analyzed and the schema is tailored to it. You should review this schema -- anything unexpected means an anomaly in the data. Once reviewed, the schema can be used to guard future data, and anomalies produced here can be used to debug model performance, understand how your data evolves over time, and identify data errors.
### Transform
The `Transform` component performs feature engineering for both training and serving. It uses the [TensorFlow Transform](https://www.tensorflow.org/tfx/transform/get_started) library.
`Transform` will take as input the data from `ExampleGen`, the schema from `SchemaGen`, as well as a module that contains user-defined Transform code.
Let's see an example of user-defined Transform code below (for an introduction to the TensorFlow Transform APIs, [see the tutorial](https://www.tensorflow.org/tfx/tutorials/transform/simple)). First, we define a few constants for feature engineering:
Note: The `%%writefile` cell magic will save the contents of the cell as a `.py` file on disk. This allows the `Transform` component to load your code as a module.
```
_taxi_constants_module_file = 'taxi_constants.py'
%%writefile {_taxi_constants_module_file}
# Categorical features are assumed to each have a maximum value in the dataset.
MAX_CATEGORICAL_FEATURE_VALUES = [24, 31, 12]
CATEGORICAL_FEATURE_KEYS = [
'trip_start_hour', 'trip_start_day', 'trip_start_month',
'pickup_census_tract', 'dropoff_census_tract', 'pickup_community_area',
'dropoff_community_area'
]
DENSE_FLOAT_FEATURE_KEYS = ['trip_miles', 'fare', 'trip_seconds']
# Number of buckets used by tf.transform for encoding each feature.
FEATURE_BUCKET_COUNT = 10
BUCKET_FEATURE_KEYS = [
'pickup_latitude', 'pickup_longitude', 'dropoff_latitude',
'dropoff_longitude'
]
# Number of vocabulary terms used for encoding VOCAB_FEATURES by tf.transform
VOCAB_SIZE = 1000
# Count of out-of-vocab buckets in which unrecognized VOCAB_FEATURES are hashed.
OOV_SIZE = 10
VOCAB_FEATURE_KEYS = [
'payment_type',
'company',
]
# Keys
LABEL_KEY = 'tips'
FARE_KEY = 'fare'
def transformed_name(key):
return key + '_xf'
```
Next, we write a `preprocessing_fn` that takes in raw data as input, and returns transformed features that our model can train on:
```
_taxi_transform_module_file = 'taxi_transform.py'
%%writefile {_taxi_transform_module_file}
import tensorflow as tf
import tensorflow_transform as tft
import taxi_constants
_DENSE_FLOAT_FEATURE_KEYS = taxi_constants.DENSE_FLOAT_FEATURE_KEYS
_VOCAB_FEATURE_KEYS = taxi_constants.VOCAB_FEATURE_KEYS
_VOCAB_SIZE = taxi_constants.VOCAB_SIZE
_OOV_SIZE = taxi_constants.OOV_SIZE
_FEATURE_BUCKET_COUNT = taxi_constants.FEATURE_BUCKET_COUNT
_BUCKET_FEATURE_KEYS = taxi_constants.BUCKET_FEATURE_KEYS
_CATEGORICAL_FEATURE_KEYS = taxi_constants.CATEGORICAL_FEATURE_KEYS
_FARE_KEY = taxi_constants.FARE_KEY
_LABEL_KEY = taxi_constants.LABEL_KEY
_transformed_name = taxi_constants.transformed_name
def preprocessing_fn(inputs):
"""tf.transform's callback function for preprocessing inputs.
Args:
inputs: map from feature keys to raw not-yet-transformed features.
Returns:
Map from string feature key to transformed feature operations.
"""
outputs = {}
for key in _DENSE_FLOAT_FEATURE_KEYS:
# Preserve this feature as a dense float, setting nan's to the mean.
outputs[_transformed_name(key)] = tft.scale_to_z_score(
_fill_in_missing(inputs[key]))
for key in _VOCAB_FEATURE_KEYS:
# Build a vocabulary for this feature.
outputs[_transformed_name(key)] = tft.compute_and_apply_vocabulary(
_fill_in_missing(inputs[key]),
top_k=_VOCAB_SIZE,
num_oov_buckets=_OOV_SIZE)
for key in _BUCKET_FEATURE_KEYS:
outputs[_transformed_name(key)] = tft.bucketize(
_fill_in_missing(inputs[key]), _FEATURE_BUCKET_COUNT)
for key in _CATEGORICAL_FEATURE_KEYS:
outputs[_transformed_name(key)] = _fill_in_missing(inputs[key])
# Was this passenger a big tipper?
taxi_fare = _fill_in_missing(inputs[_FARE_KEY])
tips = _fill_in_missing(inputs[_LABEL_KEY])
outputs[_transformed_name(_LABEL_KEY)] = tf.where(
tf.math.is_nan(taxi_fare),
tf.cast(tf.zeros_like(taxi_fare), tf.int64),
# Test if the tip was > 20% of the fare.
tf.cast(
tf.greater(tips, tf.multiply(taxi_fare, tf.constant(0.2))), tf.int64))
return outputs
def _fill_in_missing(x):
"""Replace missing values in a SparseTensor.
Fills in missing values of `x` with '' or 0, and converts to a dense tensor.
Args:
x: A `SparseTensor` of rank 2. Its dense shape should have size at most 1
in the second dimension.
Returns:
A rank 1 tensor where missing values of `x` have been filled in.
"""
if not isinstance(x, tf.sparse.SparseTensor):
return x
default_value = '' if x.dtype == tf.string else 0
return tf.squeeze(
tf.sparse.to_dense(
tf.SparseTensor(x.indices, x.values, [x.dense_shape[0], 1]),
default_value),
axis=1)
```
Now, we pass in this feature engineering code to the `Transform` component and run it to transform your data.
```
transform = Transform(
examples=example_gen.outputs['examples'],
schema=schema_gen.outputs['schema'],
module_file=os.path.abspath(_taxi_transform_module_file))
context.run(transform)
```
Let's examine the output artifacts of `Transform`. This component produces two types of outputs:
* `transform_graph` is the graph that can perform the preprocessing operations (this graph will be included in the serving and evaluation models).
* `transformed_examples` represents the preprocessed training and evaluation data.
```
transform.outputs
```
Take a peek at the `transform_graph` artifact. It points to a directory containing three subdirectories.
```
train_uri = transform.outputs['transform_graph'].get()[0].uri
os.listdir(train_uri)
```
The `transformed_metadata` subdirectory contains the schema of the preprocessed data. The `transform_fn` subdirectory contains the actual preprocessing graph. The `metadata` subdirectory contains the schema of the original data.
We can also take a look at the first three transformed examples:
```
# Get the URI of the output artifact representing the transformed examples, which is a directory
train_uri = os.path.join(transform.outputs['transformed_examples'].get()[0].uri, 'train')
# Get the list of files in this directory (all compressed TFRecord files)
tfrecord_filenames = [os.path.join(train_uri, name)
for name in os.listdir(train_uri)]
# Create a `TFRecordDataset` to read these files
dataset = tf.data.TFRecordDataset(tfrecord_filenames, compression_type="GZIP")
# Iterate over the first 3 records and decode them.
for tfrecord in dataset.take(3):
serialized_example = tfrecord.numpy()
example = tf.train.Example()
example.ParseFromString(serialized_example)
pp.pprint(example)
```
After the `Transform` component has transformed your data into features, and the next step is to train a model.
### Trainer
The `Trainer` component will train a model that you define in TensorFlow (either using the Estimator API or the Keras API with [`model_to_estimator`](https://www.tensorflow.org/api_docs/python/tf/keras/estimator/model_to_estimator)).
`Trainer` takes as input the schema from `SchemaGen`, the transformed data and graph from `Transform`, training parameters, as well as a module that contains user-defined model code.
Let's see an example of user-defined model code below (for an introduction to the TensorFlow Estimator APIs, [see the tutorial](https://www.tensorflow.org/tutorials/estimator/premade)):
```
_taxi_trainer_module_file = 'taxi_trainer.py'
%%writefile {_taxi_trainer_module_file}
import tensorflow as tf
import tensorflow_model_analysis as tfma
import tensorflow_transform as tft
from tensorflow_transform.tf_metadata import schema_utils
from tfx_bsl.tfxio import dataset_options
import taxi_constants
_DENSE_FLOAT_FEATURE_KEYS = taxi_constants.DENSE_FLOAT_FEATURE_KEYS
_VOCAB_FEATURE_KEYS = taxi_constants.VOCAB_FEATURE_KEYS
_VOCAB_SIZE = taxi_constants.VOCAB_SIZE
_OOV_SIZE = taxi_constants.OOV_SIZE
_FEATURE_BUCKET_COUNT = taxi_constants.FEATURE_BUCKET_COUNT
_BUCKET_FEATURE_KEYS = taxi_constants.BUCKET_FEATURE_KEYS
_CATEGORICAL_FEATURE_KEYS = taxi_constants.CATEGORICAL_FEATURE_KEYS
_MAX_CATEGORICAL_FEATURE_VALUES = taxi_constants.MAX_CATEGORICAL_FEATURE_VALUES
_LABEL_KEY = taxi_constants.LABEL_KEY
_transformed_name = taxi_constants.transformed_name
def _transformed_names(keys):
return [_transformed_name(key) for key in keys]
# Tf.Transform considers these features as "raw"
def _get_raw_feature_spec(schema):
return schema_utils.schema_as_feature_spec(schema).feature_spec
def _build_estimator(config, hidden_units=None, warm_start_from=None):
"""Build an estimator for predicting the tipping behavior of taxi riders.
Args:
config: tf.estimator.RunConfig defining the runtime environment for the
estimator (including model_dir).
hidden_units: [int], the layer sizes of the DNN (input layer first)
warm_start_from: Optional directory to warm start from.
Returns:
A dict of the following:
- estimator: The estimator that will be used for training and eval.
- train_spec: Spec for training.
- eval_spec: Spec for eval.
- eval_input_receiver_fn: Input function for eval.
"""
real_valued_columns = [
tf.feature_column.numeric_column(key, shape=())
for key in _transformed_names(_DENSE_FLOAT_FEATURE_KEYS)
]
categorical_columns = [
tf.feature_column.categorical_column_with_identity(
key, num_buckets=_VOCAB_SIZE + _OOV_SIZE, default_value=0)
for key in _transformed_names(_VOCAB_FEATURE_KEYS)
]
categorical_columns += [
tf.feature_column.categorical_column_with_identity(
key, num_buckets=_FEATURE_BUCKET_COUNT, default_value=0)
for key in _transformed_names(_BUCKET_FEATURE_KEYS)
]
categorical_columns += [
tf.feature_column.categorical_column_with_identity( # pylint: disable=g-complex-comprehension
key,
num_buckets=num_buckets,
default_value=0) for key, num_buckets in zip(
_transformed_names(_CATEGORICAL_FEATURE_KEYS),
_MAX_CATEGORICAL_FEATURE_VALUES)
]
return tf.estimator.DNNLinearCombinedClassifier(
config=config,
linear_feature_columns=categorical_columns,
dnn_feature_columns=real_valued_columns,
dnn_hidden_units=hidden_units or [100, 70, 50, 25],
warm_start_from=warm_start_from)
def _example_serving_receiver_fn(tf_transform_graph, schema):
"""Build the serving in inputs.
Args:
tf_transform_graph: A TFTransformOutput.
schema: the schema of the input data.
Returns:
Tensorflow graph which parses examples, applying tf-transform to them.
"""
raw_feature_spec = _get_raw_feature_spec(schema)
raw_feature_spec.pop(_LABEL_KEY)
raw_input_fn = tf.estimator.export.build_parsing_serving_input_receiver_fn(
raw_feature_spec, default_batch_size=None)
serving_input_receiver = raw_input_fn()
transformed_features = tf_transform_graph.transform_raw_features(
serving_input_receiver.features)
return tf.estimator.export.ServingInputReceiver(
transformed_features, serving_input_receiver.receiver_tensors)
def _eval_input_receiver_fn(tf_transform_graph, schema):
"""Build everything needed for the tf-model-analysis to run the model.
Args:
tf_transform_graph: A TFTransformOutput.
schema: the schema of the input data.
Returns:
EvalInputReceiver function, which contains:
- Tensorflow graph which parses raw untransformed features, applies the
tf-transform preprocessing operators.
- Set of raw, untransformed features.
- Label against which predictions will be compared.
"""
# Notice that the inputs are raw features, not transformed features here.
raw_feature_spec = _get_raw_feature_spec(schema)
serialized_tf_example = tf.compat.v1.placeholder(
dtype=tf.string, shape=[None], name='input_example_tensor')
# Add a parse_example operator to the tensorflow graph, which will parse
# raw, untransformed, tf examples.
features = tf.io.parse_example(serialized_tf_example, raw_feature_spec)
# Now that we have our raw examples, process them through the tf-transform
# function computed during the preprocessing step.
transformed_features = tf_transform_graph.transform_raw_features(
features)
# The key name MUST be 'examples'.
receiver_tensors = {'examples': serialized_tf_example}
# NOTE: Model is driven by transformed features (since training works on the
# materialized output of TFT, but slicing will happen on raw features.
features.update(transformed_features)
return tfma.export.EvalInputReceiver(
features=features,
receiver_tensors=receiver_tensors,
labels=transformed_features[_transformed_name(_LABEL_KEY)])
def _input_fn(file_pattern, data_accessor, tf_transform_output, batch_size=200):
"""Generates features and label for tuning/training.
Args:
file_pattern: List of paths or patterns of input tfrecord files.
data_accessor: DataAccessor for converting input to RecordBatch.
tf_transform_output: A TFTransformOutput.
batch_size: representing the number of consecutive elements of returned
dataset to combine in a single batch
Returns:
A dataset that contains (features, indices) tuple where features is a
dictionary of Tensors, and indices is a single Tensor of label indices.
"""
return data_accessor.tf_dataset_factory(
file_pattern,
dataset_options.TensorFlowDatasetOptions(
batch_size=batch_size, label_key=_transformed_name(_LABEL_KEY)),
tf_transform_output.transformed_metadata.schema)
# TFX will call this function
def trainer_fn(trainer_fn_args, schema):
"""Build the estimator using the high level API.
Args:
trainer_fn_args: Holds args used to train the model as name/value pairs.
schema: Holds the schema of the training examples.
Returns:
A dict of the following:
- estimator: The estimator that will be used for training and eval.
- train_spec: Spec for training.
- eval_spec: Spec for eval.
- eval_input_receiver_fn: Input function for eval.
"""
# Number of nodes in the first layer of the DNN
first_dnn_layer_size = 100
num_dnn_layers = 4
dnn_decay_factor = 0.7
train_batch_size = 40
eval_batch_size = 40
tf_transform_graph = tft.TFTransformOutput(trainer_fn_args.transform_output)
train_input_fn = lambda: _input_fn( # pylint: disable=g-long-lambda
trainer_fn_args.train_files,
trainer_fn_args.data_accessor,
tf_transform_graph,
batch_size=train_batch_size)
eval_input_fn = lambda: _input_fn( # pylint: disable=g-long-lambda
trainer_fn_args.eval_files,
trainer_fn_args.data_accessor,
tf_transform_graph,
batch_size=eval_batch_size)
train_spec = tf.estimator.TrainSpec( # pylint: disable=g-long-lambda
train_input_fn,
max_steps=trainer_fn_args.train_steps)
serving_receiver_fn = lambda: _example_serving_receiver_fn( # pylint: disable=g-long-lambda
tf_transform_graph, schema)
exporter = tf.estimator.FinalExporter('chicago-taxi', serving_receiver_fn)
eval_spec = tf.estimator.EvalSpec(
eval_input_fn,
steps=trainer_fn_args.eval_steps,
exporters=[exporter],
name='chicago-taxi-eval')
run_config = tf.estimator.RunConfig(
save_checkpoints_steps=999, keep_checkpoint_max=1)
run_config = run_config.replace(model_dir=trainer_fn_args.serving_model_dir)
estimator = _build_estimator(
# Construct layers sizes with exponetial decay
hidden_units=[
max(2, int(first_dnn_layer_size * dnn_decay_factor**i))
for i in range(num_dnn_layers)
],
config=run_config,
warm_start_from=trainer_fn_args.base_model)
# Create an input receiver for TFMA processing
receiver_fn = lambda: _eval_input_receiver_fn( # pylint: disable=g-long-lambda
tf_transform_graph, schema)
return {
'estimator': estimator,
'train_spec': train_spec,
'eval_spec': eval_spec,
'eval_input_receiver_fn': receiver_fn
}
```
Now, we pass in this model code to the `Trainer` component and run it to train the model.
```
trainer = Trainer(
module_file=os.path.abspath(_taxi_trainer_module_file),
transformed_examples=transform.outputs['transformed_examples'],
schema=schema_gen.outputs['schema'],
transform_graph=transform.outputs['transform_graph'],
train_args=trainer_pb2.TrainArgs(num_steps=10000),
eval_args=trainer_pb2.EvalArgs(num_steps=5000))
context.run(trainer)
```
#### Analyze Training with TensorBoard
Optionally, we can connect TensorBoard to the Trainer to analyze our model's training curves.
```
# Get the URI of the output artifact representing the training logs, which is a directory
model_run_dir = trainer.outputs['model_run'].get()[0].uri
%load_ext tensorboard
%tensorboard --logdir {model_run_dir}
```
### Evaluator
The `Evaluator` component computes model performance metrics over the evaluation set. It uses the [TensorFlow Model Analysis](https://www.tensorflow.org/tfx/model_analysis/get_started) library. The `Evaluator` can also optionally validate that a newly trained model is better than the previous model. This is useful in a production pipeline setting where you may automatically train and validate a model every day. In this notebook, we only train one model, so the `Evaluator` automatically will label the model as "good".
`Evaluator` will take as input the data from `ExampleGen`, the trained model from `Trainer`, and slicing configuration. The slicing configuration allows you to slice your metrics on feature values (e.g. how does your model perform on taxi trips that start at 8am versus 8pm?). See an example of this configuration below:
```
eval_config = tfma.EvalConfig(
model_specs=[
# Using signature 'eval' implies the use of an EvalSavedModel. To use
# a serving model remove the signature to defaults to 'serving_default'
# and add a label_key.
tfma.ModelSpec(signature_name='eval')
],
metrics_specs=[
tfma.MetricsSpec(
# The metrics added here are in addition to those saved with the
# model (assuming either a keras model or EvalSavedModel is used).
# Any metrics added into the saved model (for example using
# model.compile(..., metrics=[...]), etc) will be computed
# automatically.
metrics=[
tfma.MetricConfig(class_name='ExampleCount')
],
# To add validation thresholds for metrics saved with the model,
# add them keyed by metric name to the thresholds map.
thresholds = {
'accuracy': tfma.MetricThreshold(
value_threshold=tfma.GenericValueThreshold(
lower_bound={'value': 0.5}),
change_threshold=tfma.GenericChangeThreshold(
direction=tfma.MetricDirection.HIGHER_IS_BETTER,
absolute={'value': -1e-10}))
}
)
],
slicing_specs=[
# An empty slice spec means the overall slice, i.e. the whole dataset.
tfma.SlicingSpec(),
# Data can be sliced along a feature column. In this case, data is
# sliced along feature column trip_start_hour.
tfma.SlicingSpec(feature_keys=['trip_start_hour'])
])
```
Next, we give this configuration to `Evaluator` and run it.
```
# Use TFMA to compute a evaluation statistics over features of a model and
# validate them against a baseline.
# The model resolver is only required if performing model validation in addition
# to evaluation. In this case we validate against the latest blessed model. If
# no model has been blessed before (as in this case) the evaluator will make our
# candidate the first blessed model.
model_resolver = ResolverNode(
instance_name='latest_blessed_model_resolver',
resolver_class=latest_blessed_model_resolver.LatestBlessedModelResolver,
model=Channel(type=Model),
model_blessing=Channel(type=ModelBlessing))
context.run(model_resolver)
evaluator = Evaluator(
examples=example_gen.outputs['examples'],
model=trainer.outputs['model'],
#baseline_model=model_resolver.outputs['model'],
# Change threshold will be ignored if there is no baseline (first run).
eval_config=eval_config)
context.run(evaluator)
```
Now let's examine the output artifacts of `Evaluator`.
```
evaluator.outputs
```
Using the `evaluation` output we can show the default visualization of global metrics on the entire evaluation set.
```
context.show(evaluator.outputs['evaluation'])
```
To see the visualization for sliced evaluation metrics, we can directly call the TensorFlow Model Analysis library.
```
import tensorflow_model_analysis as tfma
# Get the TFMA output result path and load the result.
PATH_TO_RESULT = evaluator.outputs['evaluation'].get()[0].uri
tfma_result = tfma.load_eval_result(PATH_TO_RESULT)
# Show data sliced along feature column trip_start_hour.
tfma.view.render_slicing_metrics(
tfma_result, slicing_column='trip_start_hour')
```
This visualization shows the same metrics, but computed at every feature value of `trip_start_hour` instead of on the entire evaluation set.
TensorFlow Model Analysis supports many other visualizations, such as Fairness Indicators and plotting a time series of model performance. To learn more, see [the tutorial](https://www.tensorflow.org/tfx/tutorials/model_analysis/tfma_basic).
Since we added thresholds to our config, validation output is also available. The precence of a `blessing` artifact indicates that our model passed validation. Since this is the first validation being performed the candidate is automatically blessed.
```
blessing_uri = evaluator.outputs.blessing.get()[0].uri
!ls -l {blessing_uri}
```
Now can also verify the success by loading the validation result record:
```
PATH_TO_RESULT = evaluator.outputs['evaluation'].get()[0].uri
print(tfma.load_validation_result(PATH_TO_RESULT))
```
### Pusher
The `Pusher` component is usually at the end of a TFX pipeline. It checks whether a model has passed validation, and if so, exports the model to `_serving_model_dir`.
```
pusher = Pusher(
model=trainer.outputs['model'],
model_blessing=evaluator.outputs['blessing'],
push_destination=pusher_pb2.PushDestination(
filesystem=pusher_pb2.PushDestination.Filesystem(
base_directory=_serving_model_dir)))
context.run(pusher)
```
Let's examine the output artifacts of `Pusher`.
```
pusher.outputs
```
In particular, the Pusher will export your model in the SavedModel format, which looks like this:
```
push_uri = pusher.outputs.model_push.get()[0].uri
model = tf.saved_model.load(push_uri)
for item in model.signatures.items():
pp.pprint(item)
```
We're finished our tour of built-in TFX components!
| github_jupyter |
<a href="https://colab.research.google.com/github/changhoonhahn/gqp_mc/blob/master/nb/training_desi_simpledust_speculator.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
from google.colab import drive
drive.mount('/content/drive')
%cd /content/drive/My\ Drive/speculator_fork
import os
import pickle
import numpy as np
import tensorflow as tf
import matplotlib.pyplot as plt
from speculator import SpectrumPCA
from speculator import Speculator
# read DESI wavelength
wave = np.load('wave_fsps.npy')
# theta = [b1, b2, b3, b4, g1, g2, tau, tage]
n_param = 8
n_wave = len(wave)
batches = '0_299'
n_pcas = 20
# load trained PCA basis object
print('training PCA bases')
PCABasis = SpectrumPCA(
n_parameters=n_param, # number of parameters
n_wavelengths=n_wave, # number of wavelength values
n_pcas=n_pcas, # number of pca coefficients to include in the basis
spectrum_filenames=None, # list of filenames containing the (un-normalized) log spectra for training the PCA
parameter_filenames=[], # list of filenames containing the corresponding parameter values
parameter_selection=None) # pass an optional function that takes in parameter vector(s) and returns True/False for any extra parameter cuts we want to impose on the training sample (eg we may want to restrict the parameter ranges)
PCABasis._load_from_file('DESI_simpledust.0_199.seed0.pca%i.hdf5' % n_pcas)
_training_theta = np.load('DESI_simpledust.%s.seed0.0_199pca%i_parameters.npy'% (batches, n_pcas))
_training_pca = np.load('DESI_simpledust.%s.seed0.0_199pca%i_pca.npy'% (batches, n_pcas))
training_theta = tf.convert_to_tensor(_training_theta.astype(np.float32))
training_pca = tf.convert_to_tensor(_training_pca.astype(np.float32))
print('training set size = %i' % training_pca.shape[0])
# train Speculator
speculator = Speculator(
n_parameters=n_param, # number of model parameters
wavelengths=wave, # array of wavelengths
pca_transform_matrix=PCABasis.pca_transform_matrix,
parameters_shift=PCABasis.parameters_shift,
parameters_scale=PCABasis.parameters_scale,
pca_shift=PCABasis.pca_shift,
pca_scale=PCABasis.pca_scale,
spectrum_shift=PCABasis.spectrum_shift,
spectrum_scale=PCABasis.spectrum_scale,
n_hidden=[256, 256, 256], # network architecture (list of hidden units per layer)
restore=False,
optimizer=tf.keras.optimizers.Adam()) # optimizer for model training
# cooling schedule
lr = [1e-3, 1e-4, 1e-5, 1e-6]
batch_size = [1000, 10000, 50000, 100000]#int(training_theta.shape[0])]
gradient_accumulation_steps = [1, 1, 1, 1] # split the largest batch size into 10 when computing gradients to avoid memory overflow
# early stopping set up
patience = 20
# train using cooling/heating schedule for lr/batch-size
for i in range(len(lr)):
print('learning rate = ' + str(lr[i]) + ', batch size = ' + str(batch_size[i]))
# set learning rate
speculator.optimizer.lr = lr[i]
n_training = training_theta.shape[0]
# create iterable dataset (given batch size)
training_data = tf.data.Dataset.from_tensor_slices((training_theta, training_pca)).shuffle(n_training).batch(batch_size[i])
# set up training loss
training_loss = [np.infty]
validation_loss = [np.infty]
best_loss = np.infty
early_stopping_counter = 0
# loop over epochs
while early_stopping_counter < patience:
# loop over batches
for theta, pca in training_data:
# training step: check whether to accumulate gradients or not (only worth doing this for very large batch sizes)
if gradient_accumulation_steps[i] == 1:
loss = speculator.training_step(theta, pca)
else:
loss = speculator.training_step_with_accumulated_gradients(theta, pca, accumulation_steps=gradient_accumulation_steps[i])
# compute validation loss at the end of the epoch
validation_loss.append(speculator.compute_loss(training_theta, training_pca).numpy())
# early stopping condition
if validation_loss[-1] < best_loss:
best_loss = validation_loss[-1]
early_stopping_counter = 0
else:
early_stopping_counter += 1
if early_stopping_counter >= patience:
speculator.update_emulator_parameters()
speculator.save('_DESI_simpledust_model.%s.pca%i.log' % (batches, n_pcas))
attributes = list([
list(speculator.W_),
list(speculator.b_),
list(speculator.alphas_),
list(speculator.betas_),
speculator.pca_transform_matrix_,
speculator.pca_shift_,
speculator.pca_scale_,
speculator.spectrum_shift_,
speculator.spectrum_scale_,
speculator.parameters_shift_,
speculator.parameters_scale_,
speculator.wavelengths])
# save attributes to file
f = open('DESI_simpledust_model.%s.pca%i.log.pkl' % (batches, n_pcas), 'wb')
pickle.dump(attributes, f)
f.close()
print('Validation loss = %s' % str(best_loss))
speculator = Speculator(restore=True, restore_filename='_DESI_simpledust_model.%s.pca%i.log' % (batches, n_pcas))
# read in training parameters and data
theta_test = np.load('DESI_simpledust.theta_test.npy')
logspectrum_test = np.load('DESI_simpledust.logspectrum_fsps_test.npy')
spectrum_test = 10**logspectrum_test
logspectrum_spec = speculator.log_spectrum(theta_test.astype(np.float32))
spectrum_spec = 10**logspectrum_spec
# figure comparing Speculator log spectrum to FSPS log spectrum
fig = plt.figure(figsize=(15,5))
sub = fig.add_subplot(111)
for ii, i in enumerate(np.random.choice(len(theta_test), size=5)):
sub.plot(wave, logspectrum_spec[i], c='C%i' % i, ls='-', label='Speculator')
sub.plot(wave, logspectrum_test[i], c='C%i' % i, ls=':', label='FSPS')
if ii == 0: sub.legend(loc='upper right', fontsize=20)
sub.set_xlabel('wavelength ($A$)', fontsize=25)
sub.set_xlim(3e3, 1e4)
sub.set_ylabel('log flux', fontsize=25)
fig.savefig('validate_desi_simpledust.%s.pca%i.0.png' % (batches, n_pcas), bbox_inches='tight')
# more quantitative accuracy test of the Speculator model
frac_dspectrum = (spectrum_spec - spectrum_test) / spectrum_test
frac_dspectrum_quantiles = np.nanquantile(frac_dspectrum,
[0.0005, 0.005, 0.025, 0.5, 0.975, 0.995, 0.9995], axis=0)
fig = plt.figure(figsize=(15,5))
sub = fig.add_subplot(111)
sub.fill_between(wave, frac_dspectrum_quantiles[0],
frac_dspectrum_quantiles[6], fc='C0', ec='none', alpha=0.1, label='99.9%')
sub.fill_between(wave, frac_dspectrum_quantiles[1],
frac_dspectrum_quantiles[5], fc='C0', ec='none', alpha=0.2, label='99%')
sub.fill_between(wave, frac_dspectrum_quantiles[2],
frac_dspectrum_quantiles[4], fc='C0', ec='none', alpha=0.3, label='95%')
sub.plot(wave, frac_dspectrum_quantiles[3], c='C0', ls='-')
sub.plot(wave, np.zeros(len(wave)), c='k', ls=':')
sub.legend(loc='upper right', fontsize=20)
sub.set_xlabel('wavelength ($A$)', fontsize=25)
sub.set_xlim(3e3, 1e4)
sub.set_ylabel(r'$(f_{\rm speculator} - f_{\rm fsps})/f_{\rm fsps}$', fontsize=25)
sub.set_ylim(-0.1, 0.1)
fig.savefig('validate_desi_simpledust.%s.pca%i.1.png' % (batches, n_pcas), bbox_inches='tight')
```
| github_jupyter |
# 1. EDA
Reference:
- [NanoMathias, Feature Engineering & Importance Testing](https://www.kaggle.com/nanomathias/feature-engineering-importance-testing)
- [NanoMathias, Bayesian Tuning of xgBoost & lightGBM | LB: 0.9769](https://www.kaggle.com/nanomathias/bayesian-tuning-of-xgboost-lightgbm-lb-0-9769)
- [gopisaran, Indepth EDA - Entire TalkingData dataset](https://www.kaggle.com/gopisaran/indepth-eda-entire-talkingdata-dataset)
## Run name
```
import time
project_name = 'TalkingdataAFD2018'
step_name = 'EDA'
time_str = time.strftime("%Y%m%d_%H%M%S", time.localtime())
run_name = '%s_%s_%s' % (project_name, step_name, time_str)
print('run_name: %s' % run_name)
t0 = time.time()
```
## Important params
```
date = 9
# test_n_rows = 18790469
test_n_rows = None
day_rows = {
6: {
'n_skiprows': 1,
'n_rows': 9308568
},
7: {
'n_skiprows': 1 + 9308568,
'n_rows': 59633310
},
8: {
'n_skiprows': 1 + 9308568 + 59633310,
'n_rows': 62945075
},
9: {
'n_skiprows': 1 + 9308568 + 59633310 + 62945075,
'n_rows': 53016937
}
}
n_skiprows = day_rows[date]['n_skiprows']
n_rows = day_rows[date]['n_rows']
```
## Import PKGs
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
from IPython.display import display
import os
import gc
import time
import random
import zipfile
import h5py
import pickle
import math
from PIL import Image
import shutil
from tqdm import tqdm
import multiprocessing
from multiprocessing import cpu_count
from sklearn.metrics import roc_auc_score
```
## Project folders
```
cwd = os.getcwd()
input_folder = os.path.join(cwd, 'input')
output_folder = os.path.join(cwd, 'output')
model_folder = os.path.join(cwd, 'model')
log_folder = os.path.join(cwd, 'log')
print('input_folder: \t\t\t%s' % input_folder)
print('output_folder: \t\t\t%s' % output_folder)
print('model_folder: \t\t\t%s' % model_folder)
print('log_folder: \t\t\t%s' % log_folder)
train_csv_file = os.path.join(input_folder, 'train.csv')
train_sample_csv_file = os.path.join(input_folder, 'train_sample.csv')
test_csv_file = os.path.join(input_folder, 'test.csv')
sample_submission_csv_file = os.path.join(input_folder, 'sample_submission.csv')
print('\ntrain_csv_file: \t\t%s' % train_csv_file)
print('train_sample_csv_file: \t\t%s' % train_sample_csv_file)
print('test_csv_file: \t\t\t%s' % test_csv_file)
print('sample_submission_csv_file: \t%s' % sample_submission_csv_file)
```
## Load data
```
%%time
train_csv = pd.read_csv(train_csv_file, skiprows=range(1, n_skiprows), nrows=n_rows, parse_dates=['click_time'])
test_csv = pd.read_csv(test_csv_file, nrows=test_n_rows, parse_dates=['click_time'])
sample_submission_csv = pd.read_csv(sample_submission_csv_file)
print('train_csv.shape: \t\t', train_csv.shape)
print('test_csv.shape: \t\t', test_csv.shape)
print('sample_submission_csv.shape: \t', sample_submission_csv.shape)
print('train_csv.dtypes: \n', train_csv.dtypes)
display(train_csv.head(2))
display(test_csv.head(2))
display(sample_submission_csv.head(2))
```
## Analyses
```
train_csv['day'] = train_csv['click_time'].dt.day.astype('uint8')
train_csv['hour'] = train_csv['click_time'].dt.hour.astype('uint8')
train_csv['minute'] = train_csv['click_time'].dt.minute.astype('uint8')
train_csv['second'] = train_csv['click_time'].dt.second.astype('uint8')
print('train_csv.shape: \t', train_csv.shape)
display(train_csv.head(2))
test_csv['day'] = test_csv['click_time'].dt.day.astype('uint8')
test_csv['hour'] = test_csv['click_time'].dt.hour.astype('uint8')
test_csv['minute'] = test_csv['click_time'].dt.minute.astype('uint8')
test_csv['second'] = test_csv['click_time'].dt.second.astype('uint8')
print('test_csv.shape: \t', test_csv.shape)
display(test_csv.head(2))
train_gp_day = train_csv.groupby(['day']).size()
print(train_gp_day.shape)
print(train_gp_day)
test_gp_day = test_csv.groupby(['day']).size()
print(test_gp_day.shape)
print(test_gp_day)
train_gp_day = train_csv.groupby(['day', 'hour']).size()
print(train_gp_day.shape)
print(train_gp_day)
train_gp_day.plot(kind='bar', figsize=(20, 20))
test_gp_day = test_csv.groupby(['day', 'hour']).size()
print(test_gp_day.shape)
print(test_gp_day)
test_gp_day.plot(kind='bar', figsize=(5, 5))
print(run_name)
print('Time cost: %.2f s' % (time.time() - t0))
print('Done!')
```
| github_jupyter |
```
import re
import sys
import spacy
import numpy as np
import json
import difflib
from num2words import num2words
#sys.path.insert(0,'/home/vishesh/TUM/Thesis/huggingface/neuralcoref')
#from neuralcoref import Coref
from random import randint
from os.path import join
from numpy import array
import string
import gensim
import time
import os
import multiprocessing
word2vec = gensim.models.Word2Vec
GLOVE_DIR = "/home/vishesh/TUM/Thesis/glove6B"
model = gensim.models.KeyedVectors.load_word2vec_format(join(GLOVE_DIR, 'glove.6B.50d.w2vformat.txt'), binary=False)
nlp = spacy.load('en')
def train_file_to_list(file):
train_list = []
for line in file:
train_list.append(line)
return train_list
def get_documents(train_file):
train_list = train_file_to_list(train_file)
document = []
part = []
sentence = ''
for i in range (len(train_list)):
if train_list[i] == '\n':
part.append(sentence)
sentence = ''
continue
cols = train_list[i].split()
if cols[0] == '#begin' or cols[0] == '#end':
if len(part) > 0:
document.append(part)
part = []
continue
else:
if cols[3] == '\'s' or cols[3] == '.' or cols[3] == ',' or cols[3] == '?':
sentence = sentence.strip() + cols[3] + ' '
else:
sentence += cols[3] + ' '
return document
def create_mention_cluster_list(cluster_start, start_pos, cluster_end, end_pos):
cluster_start_end_list = []
for start, pos in zip(cluster_start, start_pos):
cluster = [start, pos]
for i in range(len(cluster_end)):
if cluster_end[i] == start:
cluster.append(end_pos[i])
break
del cluster_end[i]
del end_pos[i]
cluster_start_end_list.append(cluster)
return cluster_start_end_list
def get_mention(train_list):
cluster_start = []
start_pos = []
cluster_end = []
end_pos = []
i = 1
for line in train_list:
if line == '\n' or line == '-':
i += 1
continue
part_number = line.split()[1]
coref_col = line.split()[-1]
for j in range (len(coref_col)):
if coref_col[j] == '(':
cluster_start.append((str(part_number) + '_' + re.findall(r'\d+', coref_col[j+1:])[0]))
start_pos.append(i)
if coref_col[j] == ')':
cluster_end.append((str(part_number)+ '_' + re.findall(r'\d+', coref_col[:j])[-1]))
end_pos.append(i)
i += 1
return cluster_start, start_pos, cluster_end, end_pos
def get_mention_words(train_file_as_list, pos1, pos2):
mention = ''
for line_no in range(pos1-1, pos2):
word = train_file_as_list[line_no].split()[3]
#if word == '\'s' or word == ',' or word == '.':
# mention = mention.strip() + word + ' '
#else:
mention += word + ' '
return mention.strip()
def get_preceding_words(list, pos):
word_part = list[pos-1].split()[1]
i = 2
num_words = 0
word = []
while(True):
if list[pos-i] != '\n':
if list[pos-i].split()[0] == '#begin' or list[pos-i].split()[0] == '#end':
break
part_no = list[pos-i].split()[1]
if part_no == word_part:
word.append(list[pos-i].split()[3])
num_words += 1
if num_words == 5:
break
i += 1
return word
def get_next_words(list, pos):
pos = pos-1
word_part = list[pos].split()[1]
i = 1
num_words = 0
word = []
while(True):
if list[pos+i] != '\n':
if list[pos+i].split()[0] == '#begin' or list[pos+i].split()[0] == '#end':
break
part_no = list[pos+i].split()[1]
if part_no == word_part:
word.append(list[pos+i].split()[3])
num_words += 1
if num_words == 5:
break
i += 1
return word
def mention_sentence(train_list, pos):
pos = pos-1
i = 1
start = 0
end = 0
while(True):
if train_list[pos-i] == '\n':
start = pos-i
break
if train_list[pos-i].split()[0] == '#begin':
start = pos-i
break
i += 1
start += 2
i = 1
while(True):
if train_list[pos+i] == '\n':
end = pos+i
break
i += 1
sentence = get_mention_words(train_list, start, end)
return sentence
def document_dictionary(train_file):
documents = get_documents(train_file)
doc_sent = ''
doc_no = 0
doc_dict = {}
for document in documents:
for part in document:
doc_sent += part
doc_dict[doc_no] = doc_sent
doc_sent = ''
doc_no += 1
return doc_dict
def get_mention_length(mention):
mention_words = mention.split()
mention_len = len(mention_words)
#len_in_words = num2words(mention_len)
return mention_len
# pronoun: [1, 0, 0, 0]
# proper: [0, 1, 0, 0]
# nominal(common noun): [0, 0, 1, 0]
# list: [0, 0, 0, 1]
def mention_type(doc, mention):
# pos 0: pronoun, pos 1: proper noun, pos 2: common noun
token_type = [0, 0, 0]
for token in doc:
if token.pos_ == 'PRON':
token_type[0] += 1
elif token.pos_ == 'PROPN':
token_type[1] += 1
elif token.pos_ == 'NOUN':
token_type[2] += 1
m = max(token_type)
a = [i for i, j in enumerate(token_type) if j == m]
is_dominant = m >= len(mention.split())/2
if is_dominant:
if a[0] == 0:
return np.array([1, 0, 0, 0])
if a[0] == 1:
return np.array([0, 1, 0, 0])
if a[0] == 2:
return np.array([0, 0, 1, 0])
else:
return np.array([0, 0, 0, 1])
def check_mention_contain(newlist):
for i in range(0, len(newlist)):
start = newlist[i]['mention_start']
end = newlist[i]['mention_end']
for j in range(0, len(newlist)):
c_start = newlist[j]['mention_start']
c_end = newlist[j]['mention_end']
if c_start == start and c_end == end:
continue
if c_start >= start and c_end <= end:
newlist[j]['contained'] = newlist[i]['id']
if c_start >= start and c_start <= end:
newlist[j]['overlap'] = newlist[i]['id']
for k in range(0, len(newlist)):
if 'contained' in newlist[k]:
continue
else:
newlist[k]['contained'] = False
if 'overlap' in newlist[k]:
continue
else:
newlist[k]['overlap'] = False
return newlist
def random_with_N_digits(n):
range_start = 10**(n-1)
range_end = (10**n)-1
return randint(range_start, range_end)
# not used
def get_all_mentions(train_file):
documents = get_documents(train_file)
each_doc = ''
mention_list = []
for docs in documents:
for d in docs:
each_doc += d
print (each_doc)
clusters = coref.one_shot_coref(utterances=each_doc)
mention_list.append(coref.get_mentions())
each_doc = ''
return mention_list
#not used
def get_all_mention_cluster(file_path, train_file):
mention_list = get_all_mentions(file_path)
train_list = train_file_to_list(train_file)
start_index = []
end_index = []
mention_cluster = []
for doc_num in range(0, len(mention_list)):
for men in mention_list[doc_num]:
for i in range(0, len(train_list)):
if train_list[i] != '\n' and train_list[i].split()[0] != '#begin' and train_list[i].split()[0] != '#end':
if train_list[i].split()[3] == str(men[0]) and train_list[i].split()[1] == str(doc_num):
len_mention = len(men)
flag = True
for j, k in zip(men, train_list[i:i+len_mention]):
if k != '\n' and k.split()[0] != '#begin' and k.split()[0] != '#end':
if str(j) != k.split()[3]:
flag = False
start = i+1
end = i+len_mention
for s, e in zip(start_index, end_index):
if s == start and e == end:
flag = False
if flag == True:
start_index.append(start)
end_index.append(end)
dummy_list = []
dummy_list.append(str(doc_num)+'_' + str(random_with_N_digits(10)))
dummy_list.append(start)
dummy_list.append(end)
mention_cluster.append(dummy_list)
break
return mention_cluster
def get_index(mention_info):
doc_count = '0'
count = 0
i = 0
mentions_in_each_doc = []
for m in mention_info:
if m['id'].split('_')[0] == doc_count:
count += 1
else:
mentions_in_each_doc.append(count)
doc_count = m['id'].split('_')[0]
count = 1
m['index'] = count
mentions_in_each_doc.append(count)
doc_count = '0'
for m in mention_info:
if m['id'].split('_')[0] == doc_count:
m['mention_position'] = m['index']/mentions_in_each_doc[i]
else:
doc_count = m['id'].split('_')[0]
i += 1
m['mention_position'] = m['index']/mentions_in_each_doc[i]
return mention_info
def train_dictionary(train_file):
mention_info = []
train_list = train_file_to_list(train_file)
cluster_start, start_pos, cluster_end, end_pos = get_mention(train_list)
mention_cluster = create_mention_cluster_list(cluster_start, start_pos, cluster_end, end_pos)
for m in mention_cluster:
mention_dict = {}
mention_words = get_mention_words(train_list, m[1], m[2])
doc = nlp(mention_words)
mention_dict['id'] = m[0]
mention_dict['mention_start'] = m[1]
mention_dict['mention_end'] = m[2]
mention_dict['mention'] = mention_words
mention_dict['first_word'] = mention_words.split()[0]
mention_dict['last_word'] = mention_words.split()[-1]
if mention_words.isdigit() or mention_words == 'its' or mention_words.lower() == 'that' or mention_words.lower() == 'this':
mention_dict['head_word'] = ''
else:
if len(list(doc.noun_chunks)) > 0:
mention_dict['head_word'] = list(doc.noun_chunks)[0].root.head.text
else:
mention_dict['head_word'] = ''
mention_dict['pre_words'] = get_preceding_words(train_list, m[1])
mention_dict['next_words'] = get_next_words(train_list, m[2])
mention_dict['mention_sentence'] = mention_sentence(train_list, m[1])
mention_dict['mention_type'] = mention_type(doc, mention_words).tolist()
mention_dict['mention_length'] = get_mention_length(mention_words)
mention_dict['speaker'] = train_list[m[1] - 1].split()[9]
mention_info.append(mention_dict)
mention_info = sorted(mention_info, key=lambda k: k['mention_start'])
mention_info = check_mention_contain(mention_info)
mention_info = get_index(mention_info)
return mention_info
def distance(a):
d = np.zeros((10))
d[a == 0, 0] = 1
d[a == 1, 1] = 1
d[a == 2, 2] = 1
d[a == 3, 3] = 1
d[a == 4, 4] = 1
d[(5 <= a) & (a < 8), 5] = 1
d[(8 <= a) & (a < 16), 6] = 1
d[(16 <= a) & (a < 32), 7] = 1
d[(a >= 32) & (a < 64), 8] = 1
d[a >= 64, 9] = 1
return d.tolist()
def get_mention_pairs(train_file):
mention_info = train_dictionary(train_file)
mention_pair_list = []
for i in range(1, len(mention_info)):
for j in range(0, i):
pair = []
if mention_info[i]['id'].split('_')[0] == mention_info[j]['id'].split('_')[0]:
pair.append(mention_info[i])
pair.append(mention_info[j])
if mention_info[i]['id'] == mention_info[j]['id']:
pair.append({'coref': 1})
else:
if j % 2 == 0 or j % 3 == 0 or j % 5 == 0 or j % 7 == 0 or j % 11 == 0:
continue
else:
pair.append({'coref': 0})
mention_pair_list.append(pair)
mention_pair_list = get_sentence_dist(mention_pair_list, train_file)
return mention_pair_list
def get_sentence_dist(mention_pair_list, train_file):
train_list = train_file_to_list(train_file)
for m in mention_pair_list:
count = 0
m1 = m[0]['mention_start']
m2 = m[1]['mention_start']
if m1 < m2:
for t in range(m1, m2+1):
if train_list[t] == '\n':
count += 1
seq=difflib.SequenceMatcher(None, m[0]['mention'],m[1]['mention'])
score = seq.ratio()
m.append({'sentence_dist_count': distance(count)})
m.append({'mention_dist_count': distance(m[0]['index'] - m[1]['index'])})
if m[1]['overlap'] == m[0]['id']:
m.append({'overlap': 1})
else:
m.append({'overlap': 0})
if m[1]['speaker'] == m[0]['speaker']:
m.append({'speaker': 1})
else:
m.append({'speaker': 0})
if m[1]['head_word'] == m[0]['head_word']:
m.append({'head_match': 1})
else:
m.append({'head_match': 0})
if m[1]['mention'] == m[0]['mention']:
m.append({'mention_exact_match': 1})
else:
m.append({'mention_exact_match': 0})
if score > 0.6:
m.append({'mention_partial_match': 1})
else:
m.append({'mention_partial_match': 0})
return mention_pair_list
#with open('/home/vishesh/TUM/Thesis/coref-json/trainfile1.json', 'w') as outfile:
# json.dump(pairs, outfile)
#with open('/home/vishesh/TUM/Thesis/coref-json/documents.json', 'w') as outfile:
# json.dump(doc_dict, outfile)
```
## Create input vector
```
def get_vector(word):
table = str.maketrans({key: None for key in string.punctuation})
word = word.lower()
if len(word) > 1:
word = word.translate(table)
try:
vec = model[word]
except:
vec = np.zeros((50, 1))
return vec.reshape((50, 1))
def get_average_vector(word_list):
sum = np.zeros((50, 1))
for i in range(0, len(word_list)):
sum += get_vector(word_list[i])
average_vector = sum/(i+1)
return average_vector
def calculate_docs_average(doc_dict):
doc_avg = []
for d in doc_dict:
doc_avg.append(get_average_vector(doc_dict[d].split()))
return doc_avg
def get_pair_features(feature_list):
# distance features
mention_dist = np.array(feature_list[4]['mention_dist_count']).reshape((10, 1))
s_dist = np.array(feature_list[3]['sentence_dist_count']).reshape((10, 1))
overlap = np.array(feature_list[5]['overlap']).reshape((1, 1))
# speaker feature
speaker = np.array(feature_list[6]['speaker']).reshape((1, 1))
# string matching features
head_match = np.array(feature_list[7]['head_match']).reshape((1, 1))
mention_exact_match = np.array(feature_list[8]['mention_exact_match']).reshape((1, 1))
mention_partial_match = np.array(feature_list[9]['mention_partial_match']).reshape((1, 1))
pair_features = np.concatenate((mention_dist, s_dist, overlap, speaker, head_match, \
mention_exact_match, mention_partial_match))
return pair_features
# p: previous, n: next, w: words, a: average, s: sentence
def get_mention_features(mention, doc_average):
features = []
#head_w = get_vector(mention['head_word'])
first_w = get_vector(mention['first_word'])
last_w = get_vector(mention['last_word'])
mention_length = np.array(mention['mention_length']).reshape((1,1))
mention_type = np.array(mention['mention_type']).reshape((4, 1))
mention_position = np.array(mention['mention_position']).reshape((1, 1))
if mention['contained'] == False:
mention_contain = np.zeros((1, 1))
else:
mention_contain = np.ones((1, 1))
if len(mention['pre_words']) > 0:
mention_p_w1 = get_vector(mention['pre_words'][0])
else:
mention_p_w1 = np.zeros((50, 1))
if len(mention['pre_words']) > 1:
mention_p_w2 = get_vector(mention['pre_words'][1])
else:
mention_p_w2 = np.zeros((50, 1))
if len(mention['next_words']) > 0:
mention_n_w1 = get_vector(mention['next_words'][0])
else:
mention_n_w1 = np.zeros((50, 1))
if len(mention['next_words']) > 1:
mention_n_w2 = get_vector(mention['next_words'][1])
else:
mention_n_w2 = np.zeros((50, 1))
if len(mention['pre_words']) > 0:
mention_p_w_a = get_average_vector(mention['pre_words'])
else:
mention_p_w_a = np.zeros((50, 1))
if len(mention['next_words']) > 0:
mention_n_w_a = get_average_vector(mention['next_words'])
else:
mention_n_w_a = np.zeros((50, 1))
mention_s_a = get_average_vector(mention['mention_sentence'].split())
doc_id = mention['id'].split('_')[0]
doc_avg = doc_average[int(doc_id)]
features = np.concatenate((first_w, last_w, mention_p_w1, mention_p_w2, mention_p_w_a, \
mention_n_w1, mention_n_w2, mention_n_w_a, mention_s_a, mention_length, \
mention_type, mention_position, mention_contain, doc_avg))
return features
def make_feature_input(pairs, doc_dict):
docs_avg = calculate_docs_average(doc_dict)
input_feature_list = []
i = 0
for m in pairs:
i += 1
input_feature_vector = []
mention_avg = get_average_vector(m[0]['mention'].split())
antecedent_avg = get_average_vector(m[1]['mention'].split())
mention_features = get_mention_features(m[0], docs_avg)
antecedent_features = get_mention_features(m[1], docs_avg)
pair_features = get_pair_features(m)
input_feature_vector.append(antecedent_avg)
input_feature_vector.append(antecedent_features)
input_feature_vector.append(mention_avg)
input_feature_vector.append(mention_features)
input_feature_vector.append(pair_features)
input_feature_list.append(input_feature_vector)
return input_feature_list
def make_input_vector(pairs, doc_dict):
feature_input = make_feature_input(pairs, doc_dict)
len_f_input = len(feature_input)
input_ = []
for f_input in feature_input:
con = np.concatenate((f_input[0], f_input[1], f_input[2], f_input[3], f_input[4]))
input_.append(con)
del con
return input_
def make_output_vector(pairs):
output = []
len_mentions = len(pairs)
for m in pairs:
output.append(m[2]['coref'])
output = np.array(output).reshape((len_mentions, 1))
return output
path_to_train_file = '/home/vishesh/TUM/Thesis/conll-formatted-ontonotes-5.0-12/conll-formatted-ontonotes-5.0-12/conll-formatted-ontonotes-5.0/data/train/data/english/annotations/bc/cctv/00/cctv_0001.gold_conll'
#train_file = open(path_to_train_file, 'r')
#doc_dict = document_dictionary(train_file)
list_of_conll_files = []
for path, subdirs, files in os.walk('/home/vishesh/TUM/Thesis/conll-formatted-ontonotes-5.0-12/conll-formatted-ontonotes-5.0-12/conll-formatted-ontonotes-5.0/data/train/data/english/annotations/'):
for name in files:
if name.endswith(".gold_conll"):
list_of_conll_files.append(os.path.join(path, name))
len(list_of_conll_files)
def train_network_data(path):
train_file = open(path, 'r')
doc_dict = document_dictionary(train_file)
train_file = open(path, 'r')
pairs = get_mention_pairs(train_file)
input_vector = make_input_vector(pairs, doc_dict)
output_vector = make_output_vector(pairs)
return input_vector, output_vector
#train_network_data(list_of_conll_files[432])
%%time
input_files_vector = []
output_files_vector = []
count = 0
for num, f in enumerate(list_of_conll_files[:5000]):
print ('file num: ' + str(num))
i_vector, o_vector = train_network_data(f)
if len(i_vector) > 0:
input_files_vector.append(i_vector)
output_files_vector.append(o_vector)
count += 1
print ('coref: ' + str(count))
ffnn_input = []
ffnn_output = []
for inp_vector, out_vector in zip(input_files_vector, output_files_vector):
for inp, out in zip(inp_vector, out_vector):
ffnn_input.append(inp)
ffnn_output.append(out)
len(ffnn_input)
np.save('/home/vishesh/TUM/Thesis/Coreference-Resolution/data/processed/ffnn_input', ffnn_input, allow_pickle=True, fix_imports=True)
np.save('/home/vishesh/TUM/Thesis/Coreference-Resolution/data/processed/ffnn_output', ffnn_output, allow_pickle=True, fix_imports=True)
```
| github_jupyter |
# DARP Benchmark Instances
Este notebook tem como objetivo estudar e analisar os dados extraidos do [Scopus](https://www.scopus.com/) sobre os três artigos que primeiramente propuseram as instâncias de benchmark para o DARP e que são mencionadas por Ho et al. (2018).
```
import pandas as pd
import re
import matplotlib.pyplot as plt
plt.style.use('dark_background')
% matplotlib inline
# Importar dados dos arquivos, e concatena-los
arquivos = ["darp/cordeau&laporte(2003)/cite_cordeau&laporte(2003).csv",
"darp/cordeau(2006)/cite_cordeau(2006).csv",
"darp/ropke_etal(2007)/cite_ropke_etal(2007).csv",
"darp/ropke&cordeau(2009)/cite_ropke&cordeau(2009).csv"]
colunas_relevantes = ['Authors', 'Title', 'Year', 'Source title', 'Author Keywords', 'Index Keywords',
'Abstract', 'Cited by', 'Document Type', 'Language of Original Document']
def importar_csv_com_fonte(arquivo):
df = (pd.read_csv(arquivo, usecols=colunas_relevantes)
.assign(CiteArticle = re.search("/cite_(.*).csv", arquivo).group(1))
)
return (df)
dados = (pd.concat(map(importar_csv_com_fonte, arquivos),)
.reset_index(drop=True)
)
# Verificar se todos os dados foram corretamente importados
dados.groupby("CiteArticle")["Title"].count()
# Deletar dados duplicados
dados.drop_duplicates(subset=colunas_relevantes, inplace=True)
```
Com os dados importados, procura-se filtrar os artigos que tratem de problemas dinâmicos. Para isso, será selecionado todo o artigo que mencionar a palavra "Dynamic" no título, resumo ou lista de palavras-chave
```
dados_dinamicos = (dados.assign(AllKeywords = lambda x: x["Author Keywords"].str.cat(x["Index Keywords"], na_rep=""))
.assign(DynamicTitle = lambda x: x.Title.str.contains("Dynamic"),
DynamicKeywords = lambda x: x.AllKeywords.str.contains("Dynamic"),
DynamicAbstract = lambda x: x.Abstract.str.contains("Dynamic"))
.assign(DynamicAny = lambda x: (x.DynamicTitle | x.DynamicKeywords | x.DynamicAbstract))
)
dados_dinamicos = dados_dinamicos.loc[dados_dinamicos["DynamicAny"] == True,:]
dados_dinamicos.shape
dados_dinamicos.groupby("Document Type")["Title"].count()
```
Com esse filtro aplicado, sobram 56 itens, dos quais 39 são artigos de revista, 13 são artigos de conferência. Lendo o resumo do livro, das revisões e da pesquisa, coclui-se que estes não são de interesse para análise, já quenão apresentam experimentos computacionais.
```
artigos_dinamicos = dados_dinamicos.loc[dados_dinamicos["Document Type"].isin(["Article", "Conference Paper"])]
artigos_dinamicos.groupby("Language of Original Document")["Title"].count()
```
Lendo o resumo dos artigos restantes, foi possível determinar quais tratam de DARP, ou PDPTW dinâmicos e quais executam experimentos computacionais. Todos os artigos que não cumprem essas exigências foram deletados da análise.
```
indices_para_dropar = [53, 58, 62, 63, 64, 72, 73, 78, 93, 118, 128, 133, 140, 144, 148, 160, 185, 192,
205, 226, 238, 281, 313, 319, 327, 358, 387, 407, 417, 459, 522, 562, 567, 593,
597, 615, 636, 644, 700]
indices_para_dropar.sort()
artigos_realmente_dinamicos = artigos_dinamicos.drop(labels=indices_para_dropar)
```
Através da leitura das seções dos artigos que denotam as características dos experimentos, observa-se quais as instâncias usadas. A informação observada está inserida na coluna `"Instances"` no `DataFrame` `artigos_dinamicos`.
```
artigos_instancias = artigos_realmente_dinamicos.assign(Instance = False)
artigos_instancias.loc[artigos_instancias["Title"].str.contains("Double-horizon based heuristics for"),
"Instance"] = "Self-made, Realistic"
artigos_instancias.loc[artigos_instancias["Title"].str.contains("Waiting strategies for the dynamic pickup and"),
"Instance"] = "Self-made, Realistic"
artigos_instancias.loc[artigos_instancias["Title"].str.contains("Dynamic transportation of patients in hospitals"),
"Instance"] = "Self-made, Real"
artigos_instancias.loc[artigos_instancias["Title"].str.contains("Metaheuristics for the dynamic stochastic di"),
"Instance"] = "Self-made, Realistic, Dynamization"
artigos_instancias.loc[artigos_instancias["Title"].str.contains("On dynamic pickup and delivery vehicle"),
"Instance"] = "Self-made, Realistic"
artigos_instancias.loc[artigos_instancias["Title"].str.contains("Large scale realtime ridesharing with"),
"Instance"] = "Self-made, Real"
artigos_instancias.loc[artigos_instancias["Title"].str.contains("A tabu search heuristic for the dynamic "),
"Instance"] = "Self-made, Real, Realistic"
artigos_instancias.loc[artigos_instancias["Title"].str.contains("A hybrid tabu search and constraint programming algorithm"),
"Instance"] = "Cordeau(2006), Dynamization"
artigos_instancias.loc[artigos_instancias["Title"].str.contains("Checking the feasibility of dial-a-ride instances using c"),
"Instance"] = "Cordeau(2006), Dynamization"
artigos_instancias.loc[artigos_instancias["Title"].str.contains("On dynamic demand responsive transport services with degree"),
"Instance"] = "Self-made"
artigos_instancias.loc[artigos_instancias["Title"].str.contains("Waiting strategies for the dynamic dial-a-ride problem"),
"Instance"] = "Self-made"
artigos_instancias.loc[artigos_instancias["Title"].str.contains("Maximizing the number of served requests in an online shared"),
"Instance"] = "Self-made"
artigos_instancias.loc[artigos_instancias["Title"].str.contains("Determination of robust solutions for the dynamic dial-a-ri"),
"Instance"] = "Self-made"
artigos_instancias.shape[0]
artigos_instancias.sort_values("Cited by", ascending=False)
```
Com isso, analizamos os métodos de dinamização de cada um dos três artigos que usam deste artifício.
```
artigos_instancias[artigos_instancias["Instance"].str.contains("Dynamization")]
```
| github_jupyter |
```
# Imports
import tensorflow as tf
import tensorflow_probability as tfp
import numpy as np
import time
tfd = tfp.distributions
# Start time measures
start_time = time.clock()
# Reset the graph
tf.reset_default_graph()
# Reproducibility
# Seed setting for reproducable research.
# Set numpy seed
np.random.seed(1234)
# Set graph-level seed
tf.set_random_seed(1234)
# Util functions
def tf_squared_norm(vector):
sum_of_squares = tf.reduce_sum(tf.square(vector))
return sum_of_squares
def np_squared_norm(vector):
sum_of_squares = np.sum(np.square(vector))
return sum_of_squares
# ## Distributions functions
#
# rnorm is defined using the variance (i.e sigma^2)
def rnorm(mean, var):
sd = tf.sqrt(var)
dist = tfd.Normal(loc= mean, scale= sd)
return dist.sample()
def rbeta(alpha, beta):
dist = tfd.Beta(alpha, beta)
return dist.sample()
def rinvchisq(df, scale): # scale factor = tau^2
dist = tfd.Chi2(df)
return (df * scale)/dist.sample()
def rbernoulli(p):
dist = tfd.Bernoulli(probs=p)
return dist.sample()
# Sampling functions
#
# sample mean
def sample_mu(N, Sigma2_e, Y, X, beta):
mean = tf.reduce_sum(tf.subtract(Y, tf.matmul(X, beta)))/N
var = Sigma2_e/N
return rnorm(mean, var)
# sample variance of beta
def sample_sigma2_b( beta, NZ, v0B, s0B):
df = v0B+NZ
scale = (tf_squared_norm(beta)+v0B*s0B) / df
return rinvchisq(df, scale)
# sample error variance of Y
def sample_sigma2_e( N, epsilon, v0E, s0E):
df = v0E + N
scale = (tf_squared_norm(epsilon) + v0E*s0E) / df
return rinvchisq(df, scale)
# sample mixture weight
def sample_w( M, NZ):
w=rbeta(1+NZ,1+M-NZ)
return w
## Simulate data
# Var(g) = 0.7
# Var(Beta_true) = Var(g) / M
# Var(error) = 1 - Var(g)
def build_toy_dataset(N, M, var_g):
sigma_b = np.sqrt(var_g/M)
sigma_e = np.sqrt(1 - var_g)
beta_true = np.random.normal(0, sigma_b , M)
x = sigma_b * np.random.randn(N, M)
y = np.dot(x, beta_true) + np.random.normal(0, sigma_e, N)
return x, y, beta_true
# Simulated data parameters
N = 10 # number of data points
M = 10 # number of features
var_g = 0.7 # M * var(Beta_true)
x, y, beta_true = build_toy_dataset(N, M, var_g)
X = tf.constant(x, shape=[N,M], dtype=tf.float32)
Y = tf.constant(y, shape = [N,1], dtype=tf.float32)
# Could be implemented:
# building datasets using TF API without numpy
# # Alternative simulated data
# beta_true = tf.constant(0.25, shape=[M,1], dtype=tf.float32)
# x = np.random.randn(N,M)
# X = tf.constant(x, dtype = tf.float32)
# Y = tf.matmul(X, beta_true) + (tf.random_normal([N,1]) * 0.375)
# Precomputations
sm = np.zeros(M)
for i in range(M):
sm[i] = np_squared_norm(x[:,i])
x[:,[0]]
# Parameters setup
#
# Distinction between constant and variables
# Variables: values might change between evaluation of the graph
# (if something changes within the graph, it should be a variable)
Emu = tf.Variable(0., dtype=tf.float32)
Ebeta = tf.Variable(tf.zeros([M,1]), dtype=tf.float32)
Ew = tf.Variable(0.)
epsilon = tf.Variable(Y)
NZ = tf.Variable(0.)
Sigma2_e = tf.Variable(tf_squared_norm(Y) / (N*0.5))
Sigma2_b = tf.Variable(rbeta(1.0,1.0))
vEmu = tf.ones([N,1])
colx = tf.placeholder(tf.float32, shape=(N,1))
# Alternatives parameterization of hyperpriors for variances
v0E = tf.constant(0.001)
v0B = tf.constant(0.001)
s0B = Sigma2_b.initialized_value() / 2
s0E = Sigma2_e.initialized_value() / 2
print_dict = {'Emu': Emu, 'Ew': Ew,
'NZ': NZ, 'Sigma2_e': Sigma2_e,
'Sigma2_b': Sigma2_b}
# Tensorboard graph
#writer = tf.summary.FileWriter('.')
#writer.add_graph(tf.get_default_graph())
# updates ops
# Emu_up = Emu.assign(sample_mu(N, Sigma2_e, Y, X, Ebeta_))
#sess.run(Cj.assign(tf.reduce_sum(tf.pow(X[:,marker],2)) + Sigma2_e/Sigma2_b)) #adjusted variance
#sess.run(rj.assign(tf.matmul(tf.reshape(X[:,marker], [1,N]),epsilon)[0][0])) # mean, tensordot instead of matmul ?
#sess.run(ratio.assign(tf.exp(-(tf.pow(rj,2))/(2*Cj*Sigma2_e))*tf.sqrt((Sigma2_b*Cj)/Sigma2_e)))
#sess.run(pij.assign(Ew/(Ew+ratio*(1-Ew))))
def cond_true():
return rnorm(rj/Cj,Sigma2_e/Cj)
def cond_false():
return 0.
# Number of Gibbs sampling iterations
num_iter = 30
with tf.Session() as sess:
# Initialize variable
sess.run(tf.global_variables_initializer())
# Begin Gibbs iterations
for i in range(num_iter):
time_in = time.clock()
# Print progress
print("Gibbs sampling iteration: ", i)
# Assign a random order of marker
index = np.random.permutation(M)
# Sample mu
sess.run(Emu.assign(sample_mu(N, Sigma2_e, Y, X, Ebeta))) # matmul here
# Reset NZ parameter
sess.run(NZ.assign(0.))
# Compute beta for each marker
#print("Current marker:", end=" ")
print("Current marker:")
for marker in index:
print(marker, end=" ", flush=True)
feed = x[:,marker].reshape(N,1)
sess.run(epsilon.assign_add(colx * Ebeta[marker]),
feed_dict={colx: feed})
#TODO have the assign op elsewhere and write below tf control dep
# of that assignment
Cj = sm[marker] + Sigma2_e/Sigma2_b
rj = tf.tensordot(tf.transpose(colx), epsilon, 1)[0]
ratio = tf.exp( - ( tf.square(rj) / ( 2*Cj*Sigma2_e ))) * tf.sqrt((Sigma2_b*Cj)/Sigma2_e)
pij = Ew / (Ew + ratio*(1-Ew))
# TODO: replace with tf.cond
sess.run(Ebeta[marker,0].assign(tf.cond(tf.not_equal(rbernoulli(pij)[0],0),cond_true, cond_false)),
feed_dict={colx: feed})
sess.run(tf.cond(tf.not_equal(Ebeta[marker,0], 0.),lambda: NZ.assign_add(1.), lambda: NZ.assign_add(0.)))
sess.run(epsilon.assign_sub(colx * Ebeta[marker,0]), feed_dict={colx: feed})
#for i in range(len(Ebeta)):
# print(Ebeta[i], "\t", ny[i])
#sess.run(NZ.assign(np.sum(ny)))
sess.run(Ew.assign(sample_w(M,NZ)))
#sess.run(Ebeta_.assign(Ebeta))
sess.run(epsilon.assign(Y-tf.matmul(X,Ebeta)-vEmu*Emu))
sess.run(Sigma2_b.assign(sample_sigma2_b(Ebeta,NZ,v0B,s0B)))
sess.run(Sigma2_e.assign(sample_sigma2_e(N,epsilon,v0E,s0E)))
# Print operations
print("\n")
print(sess.run(print_dict))
print(" ")
time_out = time.clock()
print('Time for the ', i, 'th iteration: ', time_out - time_in, ' seconds')
print(" ")
# ## Print results
print("Ebeta" + '\t'+ ' beta_true')
for i in range(M):
print(Ebeta[i], "\t", beta_true[i])
# ## Printe time
print('Time elapsed: ')
print(time.clock() - start_time, "seconds")
## ONE GIBBS ITERATION
# Set timer
time_in = time.clock()
# Assign a random order of marker
index = np.random.permutation(M)
# Sample mu
sess.run(Emu.assign(sample_mu(N, Sigma2_e, Y, X, Ebeta))) # matmul here
# Reset NZ parameter
sess.run(NZ.assign(0.))
# Compute beta for each marker
for marker in index:
print(marker, end=" ", flush=True)
feed = x[:,marker].reshape(N,1)
sess.run(epsilon.assign_add(colx * Ebeta[marker]),
feed_dict={colx: feed})
#TODO have the assign op elsewhere and write below tf control dep
# of that assignment
Cj = sm[marker] + Sigma2_e/Sigma2_b
rj = tf.tensordot(tf.transpose(colx), epsilon, 1)[0]
ratio = tf.exp( - ( tf.square(rj) / ( 2*Cj*Sigma2_e ))) * tf.sqrt((Sigma2_b*Cj)/Sigma2_e)
pij = Ew / (Ew + ratio*(1-Ew))
# TODO: replace with tf.cond
sess.run(Ebeta[marker,0].assign(tf.cond(tf.not_equal(rbernoulli(pij)[0],0),cond_true, cond_false)),
feed_dict={colx: feed})
sess.run(tf.cond(tf.not_equal(Ebeta[marker,0], 0.),lambda: NZ.assign_add(1.), lambda: NZ.assign_add(0.)))
sess.run(epsilon.assign_sub(colx * Ebeta[marker,0]), feed_dict={colx: feed})
#for i in range(len(Ebeta)):
# print(Ebeta[i], "\t", ny[i])
#sess.run(NZ.assign(np.sum(ny)))
sess.run(Ew.assign(sample_w(M,NZ)))
#sess.run(Ebeta_.assign(Ebeta))
sess.run(epsilon.assign(Y-tf.matmul(X,Ebeta)-vEmu*Emu))
sess.run(Sigma2_b.assign(sample_sigma2_b(Ebeta,NZ,v0B,s0B)))
sess.run(Sigma2_e.assign(sample_sigma2_e(N,epsilon,v0E,s0E)))
# Print operations
print("\n")
print(sess.run(print_dict))
print(" ")
time_out = time.clock()
print('Time for the ', i, 'th iteration: ', time_out - time_in, ' seconds')
print(" ")
```
| github_jupyter |
# Iris Training and Prediction with Sagemaker Scikit-learn
This tutorial shows you how to use [Scikit-learn](https://scikit-learn.org/stable/) with Sagemaker by utilizing the pre-built container. Scikit-learn is a popular Python machine learning framework. It includes a number of different algorithms for classification, regression, clustering, dimensionality reduction, and data/feature pre-processing.
The [sagemaker-python-sdk](https://github.com/aws/sagemaker-python-sdk) module makes it easy to take existing scikit-learn code, which we will show by training a model on the IRIS dataset and generating a set of predictions. For more information about the Scikit-learn container, see the [sagemaker-scikit-learn-containers](https://github.com/aws/sagemaker-scikit-learn-container) repository and the [sagemaker-python-sdk](https://github.com/aws/sagemaker-python-sdk) repository.
For more on Scikit-learn, please visit the Scikit-learn website: <http://scikit-learn.org/stable/>.
### Table of contents
* [Upload the data for training](#upload_data)
* [Create a Scikit-learn script to train with](#create_sklearn_script)
* [Create the SageMaker Scikit Estimator](#create_sklearn_estimator)
* [Train the SKLearn Estimator on the Iris data](#train_sklearn)
* [Using the trained model to make inference requests](#inferece)
* [Deploy the model](#deploy)
* [Choose some data and use it for a prediction](#prediction_request)
* [Endpoint cleanup](#endpoint_cleanup)
* [Batch Transform](#batch_transform)
* [Prepare Input Data](#prepare_input_data)
* [Run Transform Job](#run_transform_job)
* [Check Output Data](#check_output_data)
First, lets create our Sagemaker session and role, and create a S3 prefix to use for the notebook example.
This notebook has been tested using the Python 3 (Data Science) kernel
```
# S3 prefix
prefix = "Scikit-iris"
import sagemaker
from sagemaker import get_execution_role
sagemaker_session = sagemaker.Session()
# Get a SageMaker-compatible role used by this Notebook Instance.
role = get_execution_role()
```
## Upload the data for training <a class="anchor" id="upload_data"></a>
When training large models with huge amounts of data, you'll typically use big data tools, like Amazon Athena, AWS Glue, or Amazon EMR, to create your data in S3. For the purposes of this example, we're using a sample of the classic [Iris dataset](https://en.wikipedia.org/wiki/Iris_flower_data_set), which is included with Scikit-learn. We will load the dataset, write locally, then write the dataset to s3 to use.
```
import numpy as np
import os
from sklearn import datasets
# Load Iris dataset, then join labels and features
iris = datasets.load_iris()
joined_iris = np.insert(iris.data, 0, iris.target, axis=1)
# Create directory and write csv
os.makedirs("./data", exist_ok=True)
np.savetxt("./data/iris.csv", joined_iris, delimiter=",", fmt="%1.1f, %1.3f, %1.3f, %1.3f, %1.3f")
```
Once we have the data locally, we can use use the tools provided by the SageMaker Python SDK to upload the data to a default bucket.
```
WORK_DIRECTORY = "data"
train_input = sagemaker_session.upload_data(
WORK_DIRECTORY, key_prefix="{}/{}".format(prefix, WORK_DIRECTORY)
)
```
## Create a Scikit-learn script to train with <a class="anchor" id="create_sklearn_script"></a>
SageMaker can now run a scikit-learn script using the `SKLearn` estimator. When executed on SageMaker a number of helpful environment variables are available to access properties of the training environment, such as:
* `SM_MODEL_DIR`: A string representing the path to the directory to write model artifacts to. Any artifacts saved in this folder are uploaded to S3 for model hosting after the training job completes.
* `SM_OUTPUT_DIR`: A string representing the filesystem path to write output artifacts to. Output artifacts may include checkpoints, graphs, and other files to save, not including model artifacts. These artifacts are compressed and uploaded to S3 to the same S3 prefix as the model artifacts.
Supposing two input channels, 'train' and 'test', were used in the call to the `SKLearn` estimator's `fit()` method, the following environment variables will be set, following the format `SM_CHANNEL_[channel_name]`:
* `SM_CHANNEL_TRAIN`: A string representing the path to the directory containing data in the 'train' channel
* `SM_CHANNEL_TEST`: Same as above, but for the 'test' channel.
A typical training script loads data from the input channels, configures training with hyperparameters, trains a model, and saves a model to model_dir so that it can be hosted later. Hyperparameters are passed to your script as arguments and can be retrieved with an `argparse.ArgumentParser` instance. For example, the script that we will run in this notebook is the below:
```python
import argparse
import pandas as pd
import os
from sklearn import tree
from sklearn.externals import joblib
if __name__ == '__main__':
parser = argparse.ArgumentParser()
# Hyperparameters are described here. In this simple example we are just including one hyperparameter.
parser.add_argument('--max_leaf_nodes', type=int, default=-1)
# Sagemaker specific arguments. Defaults are set in the environment variables.
parser.add_argument('--output-data-dir', type=str, default=os.environ['SM_OUTPUT_DATA_DIR'])
parser.add_argument('--model-dir', type=str, default=os.environ['SM_MODEL_DIR'])
parser.add_argument('--train', type=str, default=os.environ['SM_CHANNEL_TRAIN'])
args = parser.parse_args()
# Take the set of files and read them all into a single pandas dataframe
input_files = [ os.path.join(args.train, file) for file in os.listdir(args.train) ]
if len(input_files) == 0:
raise ValueError(('There are no files in {}.\n' +
'This usually indicates that the channel ({}) was incorrectly specified,\n' +
'the data specification in S3 was incorrectly specified or the role specified\n' +
'does not have permission to access the data.').format(args.train, "train"))
raw_data = [ pd.read_csv(file, header=None, engine="python") for file in input_files ]
train_data = pd.concat(raw_data)
# labels are in the first column
train_y = train_data.ix[:,0]
train_X = train_data.ix[:,1:]
# Here we support a single hyperparameter, 'max_leaf_nodes'. Note that you can add as many
# as your training my require in the ArgumentParser above.
max_leaf_nodes = args.max_leaf_nodes
# Now use scikit-learn's decision tree classifier to train the model.
clf = tree.DecisionTreeClassifier(max_leaf_nodes=max_leaf_nodes)
clf = clf.fit(train_X, train_y)
# Print the coefficients of the trained classifier, and save the coefficients
joblib.dump(clf, os.path.join(args.model_dir, "model.joblib"))
def model_fn(model_dir):
"""Deserialized and return fitted model
Note that this should have the same name as the serialized model in the main method
"""
clf = joblib.load(os.path.join(model_dir, "model.joblib"))
return clf
```
Because the Scikit-learn container imports your training script, you should always put your training code in a main guard `(if __name__=='__main__':)` so that the container does not inadvertently run your training code at the wrong point in execution.
For more information about training environment variables, please visit https://github.com/aws/sagemaker-containers.
## Create SageMaker Scikit Estimator <a class="anchor" id="create_sklearn_estimator"></a>
To run our Scikit-learn training script on SageMaker, we construct a `sagemaker.sklearn.estimator.sklearn` estimator, which accepts several constructor arguments:
* __entry_point__: The path to the Python script SageMaker runs for training and prediction.
* __role__: Role ARN
* __train_instance_type__ *(optional)*: The type of SageMaker instances for training. __Note__: Because Scikit-learn does not natively support GPU training, Sagemaker Scikit-learn does not currently support training on GPU instance types.
* __sagemaker_session__ *(optional)*: The session used to train on Sagemaker.
* __hyperparameters__ *(optional)*: A dictionary passed to the train function as hyperparameters.
To see the code for the SKLearn Estimator, see here: https://github.com/aws/sagemaker-python-sdk/tree/master/src/sagemaker/sklearn
```
from sagemaker.sklearn.estimator import SKLearn
script_path = "scikit_learn_iris.py"
sklearn = SKLearn(
entry_point=script_path,
instance_type="ml.c5.xlarge",
role=role,
framework_version='0.23-1',
py_version='py3',
sagemaker_session=sagemaker_session,
hyperparameters={"max_leaf_nodes": 10},
)
```
## Train SKLearn Estimator on Iris data <a class="anchor" id="train_sklearn"></a>
Training is very simple, just call `fit` on the Estimator! This will start a SageMaker Training job that will download the data for us, invoke our scikit-learn code (in the provided script file), and save any model artifacts that the script creates.
```
sklearn.fit({"train": train_input})
```
## Using the trained model to make inference requests <a class="anchor" id="inference"></a>
### Deploy the model <a class="anchor" id="deploy"></a>
Deploying the model to SageMaker hosting just requires a `deploy` call on the fitted model. This call takes an instance count and instance type.
```
predictor = sklearn.deploy(initial_instance_count=1, instance_type="ml.m4.xlarge")
```
### Choose some data and use it for a prediction <a class="anchor" id="prediction_request"></a>
In order to do some predictions, we'll extract some of the data we used for training and do predictions against it. This is, of course, bad statistical practice, but a good way to see how the mechanism works.
```
import itertools
import pandas as pd
shape = pd.read_csv("data/iris.csv", header=None)
a = [50 * i for i in range(3)]
b = [40 + i for i in range(10)]
indices = [i + j for i, j in itertools.product(a, b)]
test_data = shape.iloc[indices[:-1]]
test_X = test_data.iloc[:, 1:]
test_y = test_data.iloc[:, 0]
```
Prediction is as easy as calling predict with the predictor we got back from deploy and the data we want to do predictions with. The output from the endpoint return an numerical representation of the classification prediction; in the original dataset, these are flower names, but in this example the labels are numerical. We can compare against the original label that we parsed.
```
print(predictor.predict(test_X.values))
print(test_y.values)
```
### Endpoint cleanup <a class="anchor" id="endpoint_cleanup"></a>
When you're done with the endpoint, you'll want to clean it up.
```
sagemaker_session.delete_endpoint(
endpoint_name=predictor.endpoint_name
)
```
## Batch Transform <a class="anchor" id="batch_transform"></a>
We can also use the trained model for asynchronous batch inference on S3 data using SageMaker Batch Transform.
```
# Define a SKLearn Transformer from the trained SKLearn Estimator
transformer = sklearn.transformer(instance_count=1, instance_type="ml.m4.xlarge")
```
### Prepare Input Data <a class="anchor" id="prepare_input_data"></a>
We will extract 10 random samples of 100 rows from the training data, then split the features (X) from the labels (Y). Then upload the input data to a given location in S3.
```
%%bash
# Randomly sample the iris dataset 10 times, then split X and Y
mkdir -p batch_data/XY batch_data/X batch_data/Y
for i in {0..9}; do
cat data/iris.csv | shuf -n 100 > batch_data/XY/iris_sample_${i}.csv
cat batch_data/XY/iris_sample_${i}.csv | cut -d',' -f2- > batch_data/X/iris_sample_X_${i}.csv
cat batch_data/XY/iris_sample_${i}.csv | cut -d',' -f1 > batch_data/Y/iris_sample_Y_${i}.csv
done
# Upload input data from local filesystem to S3
batch_input_s3 = sagemaker_session.upload_data("batch_data/X", key_prefix=prefix + "/batch_input")
```
### Run Transform Job <a class="anchor" id="run_transform_job"></a>
Using the Transformer, run a transform job on the S3 input data.
```
# Start a transform job and wait for it to finish
transformer.transform(batch_input_s3, content_type="text/csv")
print("Waiting for transform job: " + transformer.latest_transform_job.job_name)
transformer.wait()
```
### Check Output Data <a class="anchor" id="check_output_data"></a>
After the transform job has completed, download the output data from S3. For each file "f" in the input data, we have a corresponding file "f.out" containing the predicted labels from each input row. We can compare the predicted labels to the true labels saved earlier.
```
# Download the output data from S3 to local filesystem
batch_output = transformer.output_path
!mkdir -p batch_data/output
!aws s3 cp --recursive $batch_output/ batch_data/output/
# Head to see what the batch output looks like
!head batch_data/output/*
%%bash
# For each sample file, compare the predicted labels from batch output to the true labels
for i in {1..9}; do
diff -s batch_data/Y/iris_sample_Y_${i}.csv \
<(cat batch_data/output/iris_sample_X_${i}.csv.out | sed 's/[["]//g' | sed 's/, \|]/\n/g') \
| sed "s/\/dev\/fd\/63/batch_data\/output\/iris_sample_X_${i}.csv.out/"
done
```
| github_jupyter |
# Recommendations with IBM
In this notebook, you will be putting your recommendation skills to use on real data from the IBM Watson Studio platform.
You may either submit your notebook through the workspace here, or you may work from your local machine and submit through the next page. Either way assure that your code passes the project [RUBRIC](Need to update this). **Please save regularly.**
By following the table of contents, you will build out a number of different methods for making recommendations that can be used for different situations.
## Table of Contents
I. [Exploratory Data Analysis](#Exploratory-Data-Analysis)<br>
II. [Rank Based Recommendations](#Rank)<br>
III. [User-User Based Collaborative Filtering](#User-User)<br>
IV. [Content Based Recommendations (EXTRA - NOT REQUIRED)](#Content-Recs)<br>
V. [Matrix Factorization](#Matrix-Fact)<br>
VI. [Extras & Concluding](#conclusions)
At the end of the notebook, you will find directions for how to submit your work. Let's get started by importing the necessary libraries and reading in the data.
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import project_tests as t
import operator
import collections
%matplotlib inline
df = pd.read_csv('data/user-item-interactions.csv')
df_content = pd.read_csv('data/articles_community.csv')
del df['Unnamed: 0']
del df_content['Unnamed: 0']
# Show df to get an idea of the data
df.head()
#df.email.value_counts()
#df.title.value_counts()
# Show df_content to get an idea of the data
df_content.head()
```
### <a class="anchor" id="Exploratory-Data-Analysis">Part I : Exploratory Data Analysis</a>
Use the dictionary and cells below to provide some insight into the descriptive statistics of the data.
`1.` What is the distribution of how many articles a user interacts with in the dataset? Provide a visual and descriptive statistics to assist with giving a look at the number of times each user interacts with an article.
```
df.sort_values(['article_id'])
df.groupby(['title']).agg({'title': ['sum','min','count']}).head(5)
# Fill in the median and maximum number of user_article interactios below
median_val = 3# 50% of individuals interact with ____ number of articles or fewer.
max_views_by_user = 364# The maximum number of user-article interactions by any 1 user is ______.
```
`2.` Explore and remove duplicate articles from the **df_content** dataframe.
```
# Find and explore duplicate articles
df_content.sort_values(['doc_description'])
df_content.groupby(['doc_description']).agg({'article_id': ['sum','min','count']}).max()
df_new=df_content.drop_duplicates('doc_description')
df_new.groupby(['doc_description']).agg({'article_id': ['sum','min','count']}).max()
# Remove any rows that have the same article_id - only keep the first
df_new=df_content.drop_duplicates('article_id')
df_new.groupby(['article_id']).agg({'article_id': ['sum','min','count']}).max()
df_new.head()
```
`3.` Use the cells below to find:
**a.** The number of unique articles that have an interaction with a user.
**b.** The number of unique articles in the dataset (whether they have any interactions or not).<br>
**c.** The number of unique users in the dataset. (excluding null values)<br>
**d.** The number of user-article interactions in the dataset.
```
unique_articles = 714# The number of unique articles that have at least one interaction
total_articles = 1051# The number of unique articles on the IBM platform
unique_users = 5148# The number of unique users
user_article_interactions = 45993# The number of user-article interactions
```
`4.` Use the cells below to find the most viewed **article_id**, as well as how often it was viewed. After talking to the company leaders, the `email_mapper` function was deemed a reasonable way to map users to ids. There were a small number of null values, and it was find using other information that all of these null values likely belonged to a single user.
```
most_viewed_article_id = '1429.0'# The most viewed article in the dataset as a string with one value following the decimal
max_views = 937# The most viewed article in the dataset was viewed how many times?
## No need to change the code here - this will be helpful for later parts of the notebook
# Run this cell to map the user email to a user_id column and remove the email column
def email_mapper():
coded_dict = dict()
cter = 1
email_encoded = []
for val in df['email']:
if val not in coded_dict:
coded_dict[val] = cter
cter+=1
email_encoded.append(coded_dict[val])
return email_encoded
email_encoded = email_mapper()
del df['email']
df['user_id'] = email_encoded
# show header
df.head()
user_item = df.groupby(['user_id', 'article_id'])['user_id'].max().unstack()
user_item = user_item.notnull()
user_item = user_item.applymap(lambda x: 1 if x == True else x)
user_item = user_item.applymap(lambda x: 0 if x == False else x)
user_item.head()
#df.groupby(['article_id']).agg({'article_id': ['sum','min','count']}).transform(sum).sort('count')
## If you stored all your results in the variable names above,
## you shouldn't need to change anything in this cell
sol_1_dict = {
'`50% of individuals have _____ or fewer interactions.`': median_val,
'`The total number of user-article interactions in the dataset is ______.`': user_article_interactions,
'`The maximum number of user-article interactions by any 1 user is ______.`': max_views_by_user,
'`The most viewed article in the dataset was viewed _____ times.`': max_views,
'`The article_id of the most viewed article is ______.`': most_viewed_article_id,
'`The number of unique articles that have at least 1 rating ______.`': unique_articles,
'`The number of unique users in the dataset is ______`': unique_users,
'`The number of unique articles on the IBM platform`': total_articles
}
# Test your dictionary against the solution
t.sol_1_test(sol_1_dict)
```
### <a class="anchor" id="Rank">Part II: Rank-Based Recommendations</a>
Unlike in the earlier lessons, we don't actually have ratings for whether a user liked an article or not. We only know that a user has interacted with an article. In these cases, the popularity of an article can really only be based on how often an article was interacted with.
`1.` Fill in the function below to return the **n** top articles ordered with most interactions as the top. Test your function using the tests below.
```
def get_top_articles(n, df=df):
'''
INPUT:
n - (int) the number of top articles to return
df - (pandas dataframe) df as defined at the top of the notebook
OUTPUT:
top_articles - (list) A list of the top 'n' article titles
'''
# Your code here
top_articles = list(df.title.value_counts().sort_values(ascending=False).head(n).keys().astype(str))
return top_articles # Return the top article titles from df (not df_content)
def get_top_article_ids(n, df=df):
'''
INPUT:
n - (int) the number of top articles to return
df - (pandas dataframe) df as defined at the top of the notebook
OUTPUT:
top_articles - (list) A list of the top 'n' article titles
'''
# Your code here
top_articles = list(df.article_id.value_counts().sort_values(ascending=False).head(n).keys().astype(str))
return top_articles # Return the top article ids
print(get_top_articles(10))
print(get_top_article_ids(10))
# Test your function by returning the top 5, 10, and 20 articles
top_5 = get_top_articles(5)
top_10 = get_top_articles(10)
top_20 = get_top_articles(20)
# Test each of your three lists from above
t.sol_2_test(get_top_articles)
```
### <a class="anchor" id="User-User">Part III: User-User Based Collaborative Filtering</a>
`1.` Use the function below to reformat the **df** dataframe to be shaped with users as the rows and articles as the columns.
* Each **user** should only appear in each **row** once.
* Each **article** should only show up in one **column**.
* **If a user has interacted with an article, then place a 1 where the user-row meets for that article-column**. It does not matter how many times a user has interacted with the article, all entries where a user has interacted with an article should be a 1.
* **If a user has not interacted with an item, then place a zero where the user-row meets for that article-column**.
Use the tests to make sure the basic structure of your matrix matches what is expected by the solution.
```
# create the user-article matrix with 1's and 0's
def create_user_item_matrix(df):
'''
INPUT:
df - pandas dataframe with article_id, title, user_id columns
OUTPUT:
user_item - user item matrix
Description:
Return a matrix with user ids as rows and article ids on the columns with 1 values where a user interacted with
an article and a 0 otherwise
'''
# Fill in the function here
user_item = df.groupby(['user_id', 'article_id'])['user_id'].max().unstack()
user_item = user_item.notnull()
user_item = user_item.applymap(lambda x: 1 if x == True else x)
user_item = user_item.applymap(lambda x: 0 if x == False else x)
user_item.head()
return user_item # return the user_item matrix
#user_item = create_user_item_matrix(df)
user_item.head()
#print (np.asarray(user_item.loc[5,:].tolist()))
## Tests: You should just need to run this cell. Don't change the code.
assert user_item.shape[0] == 5149, "Oops! The number of users in the user-article matrix doesn't look right."
assert user_item.shape[1] == 714, "Oops! The number of articles in the user-article matrix doesn't look right."
assert user_item.sum(axis=1)[1] == 36, "Oops! The number of articles seen by user 1 doesn't look right."
print("You have passed our quick tests! Please proceed!")
```
`2.` Complete the function below which should take a user_id and provide an ordered list of the most similar users to that user (from most similar to least similar). The returned result should not contain the provided user_id, as we know that each user is similar to him/herself. Because the results for each user here are binary, it (perhaps) makes sense to compute similarity as the dot product of two users.
Use the tests to test your function.
```
def find_similar_users(user_id, user_item=user_item):
'''
INPUT:
user_id - (int) a user_id
user_item - (pandas dataframe) matrix of users by articles:
1's when a user has interacted with an article, 0 otherwise
OUTPUT:
similar_users - (list) an ordered list where the closest users (largest dot product users)
are listed first
Description:
Computes the similarity of every pair of users based on the dot product
Returns an ordered
'''
# compute similarity of each user to the provided user
#user_articles = user_item.loc[user_id][user_item.loc[user_id].eq(0) == False].index.values
user_articles = np.asarray(user_item.loc[user_id,:].tolist())
#print(user_articles)
n_users = user_item.shape[0]
articles_seen = dict()
corr_dic = dict()
# Set up a progress bar
# cnter = 0
#bar = progressbar.ProgressBar(maxval=n_users+1, widgets=[progressbar.Bar('=', '[', ']'), ' ', progressbar.Percentage()])
#bar.start()
for user1 in range(1, n_users+1):
# update progress bar
#cnter+=1
#bar.update(cnter)
if user1 == user_id:
continue
# assign list of movies to each user key
user1_articles = np.asarray(user_item.loc[user1,:].tolist())
articles_seen[user1] = user1_articles
corr_dic[user1] = np.dot(user_articles,user1_articles)
#corr_dic = sorted(corr_dic.items(), key=operator.itemgetter(1))
corr_dic = sorted(corr_dic.items(), key=operator.itemgetter(1),reverse=True)
#corr_dic = collections.OrderedDict(corr_dic)
#print (corr_dic[0][0])
most_similar_users = list()
for i in range(0,10):
most_similar_users.append(corr_dic[i][0])
#print(most_similar_users)
#bar.finish()
# sort by similarity
# create list of just the ids
# remove the own user's id
return most_similar_users # return a list of the users in order from most to least similar
# Do a spot check of your function
print("The 10 most similar users to user 1 are: {}".format(find_similar_users(1)[:10]))
print("The 5 most similar users to user 3933 are: {}".format(find_similar_users(3933)[:5]))
print("The 3 most similar users to user 46 are: {}".format(find_similar_users(46)[:3]))
```
`3.` Now that you have a function that provides the most similar users to each user, you will want to use these users to find articles you can recommend. Complete the functions below to return the articles you would recommend to each user.
```
def get_article_names(article_ids, df=df):
'''
INPUT:
article_ids - (list) a list of article ids
df - (pandas dataframe) df as defined at the top of the notebook
OUTPUT:
article_names - (list) a list of article names associated with the list of article ids
(this is identified by the title column)
'''
# Your code here
article_names = list(df[df.article_id.isin(article_ids)].title.unique())
return article_names # Return the article names associated with list of article ids
def get_user_articles(user_id, user_item=user_item):
'''
INPUT:
user_id - (int) a user id
user_item - (pandas dataframe) matrix of users by articles:
1's when a user has interacted with an article, 0 otherwise
OUTPUT:
article_ids - (list) a list of the article ids seen by the user
article_names - (list) a list of article names associated with the list of article ids
Description:
Provides a list of the article_ids and article titles that have been seen by a user
'''
# Your code here
article_ids = list(np.array(user_item.loc[user_id][user_item.loc[user_id]==1].index).astype(str))
article_names = get_article_names(article_ids)
return article_ids, article_names # return the ids and names
def user_user_recs(user_id, m=10):
'''
INPUT:
user_id - (int) a user id
m - (int) the number of recommendations you want for the user
OUTPUT:
recs - (list) a list of recommendations for the user
Description:
Loops through the users based on closeness to the input user_id
For each user - finds articles the user hasn't seen before and provides them as recs
Does this until m recommendations are found
Notes:
Users who are the same closeness are chosen arbitrarily as the 'next' user
For the user where the number of recommended articles starts below m
and ends exceeding m, the last items are chosen arbitrarily
'''
# Your code here
recs = []
article_ids, article_names = get_user_articles(user_id)
similar_users = find_similar_users(user_id)
for user in similar_users:
user_article_ids, user_article_names = get_user_articles(user)
#print (user_article_ids, user_article_names)
for id_ in user_article_ids:
if id_ not in article_ids:
recs.append(id_)
if len(recs) == m:
break
if len(recs) == m:
break
print ('rec ',len(recs) ,recs)
return recs # return your recommendations for this user_id
# Check Results
get_article_names(user_user_recs(1, 10)) # Return 10 recommendations for user 1
# Test your functions here - No need to change this code - just run this cell
assert set(get_article_names(['1024.0', '1176.0', '1305.0', '1314.0', '1422.0', '1427.0'])) == set(['using deep learning to reconstruct high-resolution audio', 'build a python app on the streaming analytics service', 'gosales transactions for naive bayes model', 'healthcare python streaming application demo', 'use r dataframes & ibm watson natural language understanding', 'use xgboost, scikit-learn & ibm watson machine learning apis']), "Oops! Your the get_article_names function doesn't work quite how we expect."
assert set(get_article_names(['1320.0', '232.0', '844.0'])) == set(['housing (2015): united states demographic measures','self-service data preparation with ibm data refinery','use the cloudant-spark connector in python notebook']), "Oops! Your the get_article_names function doesn't work quite how we expect."
assert set(get_user_articles(20)[0]) == set(['1320.0', '232.0', '844.0'])
assert set(get_user_articles(20)[1]) == set(['housing (2015): united states demographic measures', 'self-service data preparation with ibm data refinery','use the cloudant-spark connector in python notebook'])
assert set(get_user_articles(2)[0]) == set(['1024.0', '1176.0', '1305.0', '1314.0', '1422.0', '1427.0'])
assert set(get_user_articles(2)[1]) == set(['using deep learning to reconstruct high-resolution audio', 'build a python app on the streaming analytics service', 'gosales transactions for naive bayes model', 'healthcare python streaming application demo', 'use r dataframes & ibm watson natural language understanding', 'use xgboost, scikit-learn & ibm watson machine learning apis'])
print("If this is all you see, you passed all of our tests! Nice job!")
```
`4.` Now we are going to improve the consistency of the **user_user_recs** function from above.
* Instead of arbitrarily choosing when we obtain users who are all the same closeness to a given user - choose the users that have the most total article interactions before choosing those with fewer article interactions.
* Instead of arbitrarily choosing articles from the user where the number of recommended articles starts below m and ends exceeding m, choose articles with the articles with the most total interactions before choosing those with fewer total interactions. This ranking should be what would be obtained from the **top_articles** function you wrote earlier.
```
def get_top_sorted_users(user_id, df=df, user_item=user_item):
'''
INPUT:
user_id - (int)
df - (pandas dataframe) df as defined at the top of the notebook
user_item - (pandas dataframe) matrix of users by articles:
1's when a user has interacted with an article, 0 otherwise
OUTPUT:
neighbors_df - (pandas dataframe) a dataframe with:
neighbor_id - is a neighbor user_id
similarity - measure of the similarity of each user to the provided user_id
num_interactions - the number of articles viewed by the user - if a u
Other Details - sort the neighbors_df by the similarity and then by number of interactions where
highest of each is higher in the dataframe
'''
# Your code here
df_article_views = df.groupby('user_id').count().article_id
neighbors_df = pd.concat([user_item.loc[user_id].dot(user_item.T), df_article_views], axis=1 ).reset_index()
neighbors_df.rename(columns={"user_id": "neighbor_id",
user_id: "similarity", "article_id": "num_interactions"}, inplace=True)
neighbors_df.drop(neighbors_df[neighbors_df.neighbor_id==user_id].index, axis=0, inplace=True)
neighbors_df['similarity'] = neighbors_df['similarity'].astype('int')
neighbors_df['neighbor_id'] = neighbors_df['neighbor_id'].astype('int')
neighbors_df = neighbors_df.sort_values(by = ['similarity', 'num_interactions'], ascending = [False, False])
#print(neighbors_df)
return neighbors_df # Return the dataframe specified in the doc_string
#get_top_sorted_users(131, df=df, user_item=user_item)
def user_user_recs_part2(user_id, m=10):
'''
INPUT:
user_id - (int) a user id
m - (int) the number of recommendations you want for the user
OUTPUT:
recs - (list) a list of recommendations for the user by article id
rec_names - (list) a list of recommendations for the user by article title
Description:
Loops through the users based on closeness to the input user_id
For each user - finds articles the user hasn't seen before and provides them as recs
Does this until m recommendations are found
Notes:
* Choose the users that have the most total article interactions
before choosing those with fewer article interactions.
* Choose articles with the articles with the most total interactions
before choosing those with fewer total interactions.
'''
# Your code here
neighbors_df = get_top_sorted_users(user_id)
recs = []
for neighbor in neighbors_df.neighbor_id:
article_ids = list(np.array(user_item.loc[neighbor][user_item.loc[neighbor]==1].index).astype(str))
#article_ids = sort_articles_by_popularity(article_ids)
recs.extend([aid for aid in article_ids if aid not in recs])
if(len(recs) >= m):
break;
recs = recs[:m]
rec_names=get_article_names(recs)
return recs, rec_names
# Quick spot check - don't change this code - just use it to test your functions
rec_ids, rec_names = user_user_recs_part2(20, 10)
print("The top 10 recommendations for user 20 are the following article ids:")
print(rec_ids)
print()
print("The top 10 recommendations for user 20 are the following article names:")
print(rec_names)
```
`5.` Use your functions from above to correctly fill in the solutions to the dictionary below. Then test your dictionary against the solution. Provide the code you need to answer each following the comments below.
```
### Tests with a dictionary of results
user1_most_sim = get_top_sorted_users(1).iloc[0].neighbor_id# Find the user that is most similar to user 1
user131_10th_sim = get_top_sorted_users(131).iloc[9].neighbor_id# Find the 10th most similar user to user 131
print(user1_most_sim , user131_10th_sim)
## Dictionary Test Here
sol_5_dict = {
'The user that is most similar to user 1.': user1_most_sim,
'The user that is the 10th most similar to user 131': user131_10th_sim,
}
t.sol_5_test(sol_5_dict)
```
`6.` If we were given a new user, which of the above functions would you be able to use to make recommendations? Explain. Can you think of a better way we might make recommendations? Use the cell below to explain a better method for new users.
**Provide your response here.**
`7.` Using your existing functions, provide the top 10 recommended articles you would provide for the a new user below. You can test your function against our thoughts to make sure we are all on the same page with how we might make a recommendation.
```
new_user = '0.0'
# What would your recommendations be for this new user '0.0'? As a new user, they have no observed articles.
# Provide a list of the top 10 article ids you would give to
new_user_recs = get_top_article_ids(10) # Your recommendations here
assert set(new_user_recs) == set(['1314.0','1429.0','1293.0','1427.0','1162.0','1364.0','1304.0','1170.0','1431.0','1330.0']), "Oops! It makes sense that in this case we would want to recommend the most popular articles, because we don't know anything about these users."
print("That's right! Nice job!")
```
### <a class="anchor" id="Content-Recs">Part IV: Content Based Recommendations (EXTRA - NOT REQUIRED)</a>
Another method we might use to make recommendations is to perform a ranking of the highest ranked articles associated with some term. You might consider content to be the **doc_body**, **doc_description**, or **doc_full_name**. There isn't one way to create a content based recommendation, especially considering that each of these columns hold content related information.
`1.` Use the function body below to create a content based recommender. Since there isn't one right answer for this recommendation tactic, no test functions are provided. Feel free to change the function inputs if you decide you want to try a method that requires more input values. The input values are currently set with one idea in mind that you may use to make content based recommendations. One additional idea is that you might want to choose the most popular recommendations that meet your 'content criteria', but again, there is a lot of flexibility in how you might make these recommendations.
### This part is NOT REQUIRED to pass this project. However, you may choose to take this on as an extra way to show off your skills.
```
def make_content_recs():
'''
INPUT:
OUTPUT:
'''
```
`2.` Now that you have put together your content-based recommendation system, use the cell below to write a summary explaining how your content based recommender works. Do you see any possible improvements that could be made to your function? Is there anything novel about your content based recommender?
### This part is NOT REQUIRED to pass this project. However, you may choose to take this on as an extra way to show off your skills.
**Write an explanation of your content based recommendation system here.**
`3.` Use your content-recommendation system to make recommendations for the below scenarios based on the comments. Again no tests are provided here, because there isn't one right answer that could be used to find these content based recommendations.
### This part is NOT REQUIRED to pass this project. However, you may choose to take this on as an extra way to show off your skills.
```
# make recommendations for a brand new user
# make a recommendations for a user who only has interacted with article id '1427.0'
```
### <a class="anchor" id="Matrix-Fact">Part V: Matrix Factorization</a>
In this part of the notebook, you will build use matrix factorization to make article recommendations to the users on the IBM Watson Studio platform.
`1.` You should have already created a **user_item** matrix above in **question 1** of **Part III** above. This first question here will just require that you run the cells to get things set up for the rest of **Part V** of the notebook.
```
# Load the matrix here
user_item_matrix = pd.read_pickle('user_item_matrix.p')
# quick look at the matrix
user_item_matrix.head()
```
`2.` In this situation, you can use Singular Value Decomposition from [numpy](https://docs.scipy.org/doc/numpy-1.14.0/reference/generated/numpy.linalg.svd.html) on the user-item matrix. Use the cell to perfrom SVD, and explain why this is different than in the lesson.
```
# Perform SVD on the User-Item Matrix Here
u, s, vt = np.linalg.svd(user_item)# use the built in to get the three matrices
print(u.shape, s.shape, vt.shape, user_item.shape)
```
**Provide your response here.**
`3.` Now for the tricky part, how do we choose the number of latent features to use? Running the below cell, you can see that as the number of latent features increases, we obtain a lower error rate on making predictions for the 1 and 0 values in the user-item matrix. Run the cell below to get an idea of how the accuracy improves as we increase the number of latent features.
```
num_latent_feats = np.arange(10,700+10,20)
sum_errs = []
for k in num_latent_feats:
# restructure with k latent features
s_new, u_new, vt_new = np.diag(s[:k]), u[:, :k], vt[:k, :]
# take dot product
user_item_est = np.around(np.dot(np.dot(u_new, s_new), vt_new))
# compute error for each prediction to actual value
diffs = np.subtract(user_item, user_item_est)
# total errors and keep track of them
err = np.sum(np.sum(np.abs(diffs)))
sum_errs.append(err)
plt.plot(num_latent_feats, 1 - np.array(sum_errs)/df.shape[0]);
plt.xlabel('Number of Latent Features');
plt.ylabel('Accuracy');
plt.title('Accuracy vs. Number of Latent Features');
```
`4.` From the above, we can't really be sure how many features to use, because simply having a better way to predict the 1's and 0's of the matrix doesn't exactly give us an indication of if we are able to make good recommendations. Instead, we might split our dataset into a training and test set of data, as shown in the cell below.
Use the code from question 3 to understand the impact on accuracy of the training and test sets of data with different numbers of latent features. Using the split below:
* How many users can we make predictions for in the test set?
* How many users are we not able to make predictions for because of the cold start problem?
* How many articles can we make predictions for in the test set?
* How many articles are we not able to make predictions for because of the cold start problem?
```
df_train = df.head(40000)
df_test = df.tail(5993)
def create_test_and_train_user_item(df_train, df_test):
'''
INPUT:
df_train - training dataframe
df_test - test dataframe
OUTPUT:
user_item_train - a user-item matrix of the training dataframe
(unique users for each row and unique articles for each column)
user_item_test - a user-item matrix of the testing dataframe
(unique users for each row and unique articles for each column)
test_idx - all of the test user ids
test_arts - all of the test article ids
'''
# Your code here
user_item_train = create_user_item_matrix(df_train)
user_item_test = create_user_item_matrix(df_test)
test_idx = list(set(user_item_test.index))
test_arts = list(set(user_item_test.columns))
return user_item_train, user_item_test, test_idx, test_arts
#user_item_train, user_item_test, test_idx, test_arts = create_test_and_train_user_item(df_train, df_test)
#print ( '/n arts', test_arts)
user_item_train, user_item_test, test_idx, test_arts = create_test_and_train_user_item(df_train, df_test)
user_item_train.shape, user_item_test.shape, len(test_idx), len(test_arts)
# How many users can we make predictions for in the test set?
print(len(set(user_item_test.index) & set(user_item_train.index)))
# How many users in the test set are we not able to make predictions for because of the cold start problem?
print(len(set(user_item_test.index) - set(user_item_train.index)))
# How many articles can we make predictions for in the test set?
print(len(set(user_item_test.columns) & set(user_item_train.columns)))
# How many articles in the test set are we not able to make predictions for because of the cold start problem?
print(len(set(user_item_test.columns) - set(user_item_train.columns)))
# Replace the values in the dictionary below
a = 662
b = 574
c = 20
d = 0
sol_4_dict = {
'How many users can we make predictions for in the test set?': c,
'How many users in the test set are we not able to make predictions for because of the cold start problem?': a,
'How many articles can we make predictions for in the test set?': b,
'How many articles in the test set are we not able to make predictions for because of the cold start problem?': d
}
t.sol_4_test(sol_4_dict)
```
`5.` Now use the **user_item_train** dataset from above to find **U**, **S**, and **V** transpose using SVD. Then find the subset of rows in the **user_item_test** dataset that you can predict using this matrix decomposition with different numbers of latent features to see how many features makes sense to keep based on the accuracy on the test data. This will require combining what was done in questions `2` - `4`.
Use the cells below to explore how well SVD works towards making predictions for recommendations on the test data.
```
# fit SVD on the user_item_train matrix
u_train, s_train, vt_train = np.linalg.svd(user_item_train) # use the built in to get the three matrices
print(u_train.shape, s_train.shape, vt_train.shape, user_item_train.shape) # fit svd similar to above then use the cells below
# Use these cells to see how well you can use the training
# decomposition to predict on test data
u_test = u_train[user_item_train.index.isin(test_idx), :]
vt_test = vt_train[:, user_item_train.columns.isin(test_arts)]
# Similarly, find the predictable subset of users and articles
test_users = set(user_item_train.index) & set(user_item_test.index)
test_articles = set(user_item_train.columns) & set(user_item_test.columns)
user_item_test_predictable = user_item_test.loc[test_users, test_articles]
num_latent_feats = np.arange(10,700+10,20)
squared_errors_train = []
squared_errors_test = []
for k in num_latent_feats:
# restructure with k latent features
s_train_new, u_train_new, vt_train_new = np.diag(s_train[:k]), u_train[:, :k], vt_train[:k, :]
u_test_new, vt_test_new = u_test[:, :k], vt_test[:k, :]
# take dot product
user_item_train_pred = u_train_new.dot(s_train_new).dot(vt_train_new)
user_item_test_pred = u_test_new.dot(s_train_new).dot(vt_test_new)
# compute difference between predicted and actual value
diffs_train = np.subtract(user_item_train, user_item_train_pred)
diffs_test = np.subtract(user_item_test_predictable, user_item_test_pred)
# calculate squared errors
squared_errors_train.append((diffs_train**2).sum().sum())
squared_errors_test.append((diffs_test**2).sum().sum())
fig, ax1 = plt.subplots()
ax2 = ax1.twinx()
ax1.plot(num_latent_feats, 1 - np.array(squared_errors_train)/df.shape[0], label="Train accuracy")
ax2.plot(num_latent_feats, 1 - np.array(squared_errors_test)/df.shape[0], color='green', label="Test accuracy")
ax1.set_title('Accuracy vs. Number of Latent Features')
ax1.set_xlabel('Number of Latent Features')
ax1.set_ylabel('Train accuracy')
ax2.set_ylabel('Test accuracy', rotation=270, labelpad=12)
plt.show()
```
`6.` Use the cell below to comment on the results you found in the previous question. Given the circumstances of your results, discuss what you might do to determine if the recommendations you make with any of the above recommendation systems are an improvement to how users currently find articles?
**Your response here.**
<a id='conclusions'></a>
### Extras
Using your workbook, you could now save your recommendations for each user, develop a class to make new predictions and update your results, and make a flask app to deploy your results. These tasks are beyond what is required for this project. However, from what you learned in the lessons, you certainly capable of taking these tasks on to improve upon your work here!
## Conclusion
> Congratulations! You have reached the end of the Recommendations with IBM project!
> **Tip**: Once you are satisfied with your work here, check over your report to make sure that it is satisfies all the areas of the rubric (found on the project submission page at the end of the lesson). You should also probably remove all of the "Tips" like this one so that the presentation is as polished as possible.
## Directions to Submit
> Before you submit your project, you need to create a .html or .pdf version of this notebook in the workspace here. To do that, run the code cell below. If it worked correctly, you should get a return code of 0, and you should see the generated .html file in the workspace directory (click on the orange Jupyter icon in the upper left).
> Alternatively, you can download this report as .html via the **File** > **Download as** submenu, and then manually upload it into the workspace directory by clicking on the orange Jupyter icon in the upper left, then using the Upload button.
> Once you've done this, you can submit your project by clicking on the "Submit Project" button in the lower right here. This will create and submit a zip file with this .ipynb doc and the .html or .pdf version you created. Congratulations!
```
from subprocess import call
call(['python', '-m', 'nbconvert', 'Recommendations_with_IBM.ipynb'])
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import string
import datetime
import email
import re
from tqdm import tqdm
import logging
import nltk
from nltk.corpus import stopwords
import numpy as np
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import GridSearchCV
from sklearn.model_selection import cross_val_score
import torch
import transformers as ppb
import warnings
warnings.filterwarnings('ignore')
# Read the CSV File
raw_emails_df = pd.read_csv('giskard_dataset.csv', sep=';')
def get_text_from_email(msg):
'''
To get the content from email objects
'''
parts = []
for part in msg.walk():
if part.get_content_type() == 'text/plain':
parts.append( part.get_payload() )
return ''.join(parts)
# Parse the emails into a list email objects
messages = list(map(email.message_from_string, raw_emails_df['Message']))
raw_emails_df.drop('Message', axis=1, inplace=True)
# Get fields from parsed email objects
keys = messages[0].keys()
for key in keys:
raw_emails_df[key] = [doc[key] for doc in messages]
# Parse content from emails
raw_emails_df['Content'] = list(map(get_text_from_email, messages))
raw_emails_df
emails_df = raw_emails_df[['X-Folder','Subject','Content', 'Date','Target']]
emails_df = emails_df.rename(columns={'X-Folder': 'Folder'})
emails_df
# To Confirm there are no blank data
emails_df = emails_df.drop(emails_df.query('Date == "" | Subject == "" | Content == "" | Folder ==""').index)
emails_df.reset_index(drop=True, inplace=True) # To reset the index of the dropped rows
def clean_data(columns):
"""
To Clean Data from columns of dataframes
Removes unnecessary character, lowercases words
and performs stemming/lemitizing
"""
columns = re.sub("=\n", "", columns) # To remove the '=' at every EOL
#columns = re.sub(r"&", "", columns) # To remove the '&' in company names
stopwords = nltk.corpus.stopwords.words('english')
newstopwords = ['ect', 'hou', 'com', 'recipient', 'cc', 'na', 'ees', 'th', 'pm', 'pg']
stopwords.extend(newstopwords)
#data = re.sub(r"\S*https?:\S*", "", columns)
#data = re.sub('[^a-zA-Z]', ' ', columns) # removes all characters except a-z and A-Z
data = columns.lower()
data = nltk.word_tokenize(data)
data = [word for word in data if not word in (stopwords)]
# if clean == 'stem':
# ps = PorterStemmer()
# data = [ps.stem(word) for word in data if not word in (stopwords)] # stemming all words that are not stopwords
# elif clean == 'lem':
# lm = WordNetLemmatizer() # initialize lemmatizing
# data = [lm.lemmatize(word) for word in data if not word in (stopwords)] # lemmitizing all words that are not stopwords
# else:
# data = [word for word in data if not word in (stopwords)] # stemming all words that are not stopwords
data = ' '.join(data)
#data = ' '.join(i for i in data if i not in (string.punctuation))
return data
emails_df['Texttransformed'] = emails_df['Date'] + emails_df['Folder'] + " " + emails_df['Subject'] + " " + emails_df['Content']
emails_df['Texttransformed'] = emails_df.apply(lambda x: clean_data(x['Texttransformed']), axis=1)
# For DistilBERT:
model_class, tokenizer_class, pretrained_weights = (ppb.DistilBertModel, ppb.DistilBertTokenizer, 'distilbert-base-uncased')
## Want BERT instead of distilBERT? Uncomment the following line:
#model_class, tokenizer_class, pretrained_weights = (ppb.BertModel, ppb.BertTokenizer, 'bert-base-uncased')
# Load pretrained model/tokenizer
tokenizer = tokenizer_class.from_pretrained(pretrained_weights)
model = model_class.from_pretrained(pretrained_weights)
tokenized = emails_df['Texttransformed'].apply((lambda x: tokenizer.encode(x, add_special_tokens=True)))
tokenized
max_len = 0
for i in tokenized.values:
if len(i) > max_len:
max_len = len(i)
padded = np.array([i + [0]*(max_len-len(i)) for i in tokenized.values])
attention_mask = np.where(padded != 0, 1, 0)
attention_mask.shape
#x = torch.tensor(train).to(torch.int64)
input_ids = torch.tensor(padded).to(torch.int64)
attention_mask = torch.tensor(attention_mask)
with torch.no_grad():
last_hidden_states = model(input_ids, attention_mask=attention_mask)
```
| github_jupyter |
# Домашнее задание 2.
Продолжаем знакомиться с библиотекой `tensorflow`.
Задание выполнил(а): Подчезерцев Алексей
```
import tensorflow as tf
print(tf.__version__)
import numpy as np
import matplotlib.pyplot as plt
import time
%matplotlib inline
```
## Задание 1 -- tensorflow vs numpy (3 балла).
Сравните скорость работы функций над массивами в фреймворках tensorflow и numpy. Для этого реализуйте на нампае и тф'е следующее:
* Сумму квадратов диагональных элементов квадратной матрицы. Например для матрицы
$$
\begin{pmatrix}
1& 0& 5\\
-2& 8& 12\\
4& 1& -5
\end{pmatrix}
$$
такая сумма будет равна $1^2 + 8^2 + (-5)^2 = 90$.
* Угол между векторами в n-мерном пространстве. Напомним, что он вычисляется по формуле
$$
\arccos \cfrac{\left\langle x, y\right\rangle}{||x||\cdot ||y||}
$$
* Сумму элементов коммутатора квадратных матриц $A$ и $B$. Коммутатор матриц это матрица $C = AB - BA$.
Постройте графики зависимости времени выполнения операций от размера массивов (в логарифмическй шкале) для каждой задачи для tensorflow и numpy (три рисунка, по два графика на рисунок). Элементы матриц выбирайте случайным образом (через модуль tf.random и np.random соотвтетственно). Какой фреймворк оказывается быстрее? Как Вы думаете, почему?
Можете пользоваться образцом кода ниже.
**Замечание**. Графики должны быть опрятными! Подписывайте оси и единицы измерения, указывайте легенду. За неопрятные графики оценка за задание может быть снижена.
**Подсказка**. Функция time.time() возвращает время в секундах (с высокой точностью), прошедшее от 00:00 1 января 1970 года. Используйте её, чтобы посчитать, сколько длилось выполнение куска кода. Также вам могут пригодиться функции `tf.linalg.norm`, `tf.diag_part`, `tf.acos`, `tf.matmul`
```
def log_int_iterator(start, end, step):
i = start
last = None
while i <= end:
if int(i) != last:
last = int(i)
yield last
i *= step
MAX_VALUE = 1
MIN_VALUE = 0
hist_n = np.array([])
hist_t_np = np.array([])
hist_t_tf = np.array([])
for N in log_int_iterator(1, 10000, 1.1):
hist_n = np.append(hist_n, N)
np.random.seed(42)
x = np.random.random_sample(size=(N, N))
start = time.time()
_ = np.sum(x.diagonal() ** 2)
hist_t_np = np.append(hist_t_np, time.time() - start)
tf.reset_default_graph()
with tf.Session() as sess:
x = tf.random.uniform(shape=(N, N), minval=MIN_VALUE, maxval=MAX_VALUE, dtype=tf.float32, seed=42)
x_res = sess.run(x)
# нам не надо считать время генерации случайных чисел
x_placeholder = tf.placeholder(tf.float32, (N, N))
y = tf.math.reduce_sum(tf.square(tf.linalg.tensor_diag_part(x_placeholder)))
start = time.time()
sess.run(y, {x_placeholder:x_res})
hist_t_tf = np.append(hist_t_tf, time.time() - start)
hist_t_np = hist_t_np * 1000
hist_t_tf = hist_t_tf * 1000
plt.figure(figsize=[16, 7])
plt.title('Зависимость времени вычисления "суммы квадратов диагональных элементов квадратной матрицы" для numpy и tensorflow')
plt.scatter(hist_n, hist_t_np, c='r', label='numpy')
plt.scatter(hist_n, hist_t_tf, c='b', label='tensorflow')
plt.legend(loc='upper left')
plt.xscale("log")
plt.ylim([min(hist_t_tf.min(), hist_t_np.min()), max(hist_t_tf.max(), hist_t_np.max())])
plt.xlabel("Размер матрицы, элементы")
plt.ylabel("Время вычисления, мс")
plt.show()
MAX_VALUE = 1
MIN_VALUE = 0
hist_n = np.array([])
hist_t_np = np.array([])
hist_t_tf = np.array([])
for N in log_int_iterator(1, 1000, 1.1):
hist_n = np.append(hist_n, N)
np.random.seed(42)
x = np.random.random_sample(size=(N))
y = np.random.random_sample(size=(N))
start = time.time()
_ = np.arccos(np.dot(x, y)/(np.linalg.norm(x) * np.linalg.norm(y)))
hist_t_np = np.append(hist_t_np, time.time() - start)
tf.reset_default_graph()
with tf.Session() as sess:
x = tf.random.uniform(shape=(N,), minval=MIN_VALUE, maxval=MAX_VALUE, dtype=tf.float32, seed=42)
y = tf.random.uniform(shape=(N,), minval=MIN_VALUE, maxval=MAX_VALUE, dtype=tf.float32, seed=43)
x_res, y_res = sess.run([x, y])
x_placeholder = tf.placeholder(tf.float32, (N,))
y_placeholder = tf.placeholder(tf.float32, (N,))
res = tf.math.acos(tf.linalg.tensordot(x_placeholder, y_placeholder, 1)/(tf.linalg.norm(x_placeholder) * tf.linalg.norm(y_placeholder)))
start = time.time()
_ = sess.run(res, {x_placeholder:x_res, y_placeholder: y_res})
hist_t_tf = np.append(hist_t_tf, time.time() - start)
hist_t_np = hist_t_np * 1000
hist_t_tf = hist_t_tf * 1000
plt.figure(figsize=[16, 7])
plt.title('Зависимость времени вычисления "угла между векторами в n мерном пространстве" для numpy и tensorflow')
plt.scatter(hist_n, hist_t_np, c='r', label='numpy')
plt.scatter(hist_n, hist_t_tf, c='b', label='tensorflow')
plt.legend(loc='upper left')
plt.xscale("log")
plt.ylim([min(hist_t_tf.min(), hist_t_np.min()), max(hist_t_tf.max(), hist_t_np.max())])
plt.xlabel("Размер матрицы, элементы")
plt.ylabel("Время вычисления, мс")
plt.show()
MAX_VALUE = 1
MIN_VALUE = 0
hist_n = np.array([])
hist_t_np = np.array([])
hist_t_tf = np.array([])
for N in log_int_iterator(1, 5000, 1.1):
hist_n = np.append(hist_n, N)
np.random.seed(42)
A = np.random.random_sample(size=(N, N))
B = np.random.random_sample(size=(N, N))
start = time.time()
_ = np.sum(np.dot(A,B) - np.dot(B,A))
hist_t_np = np.append(hist_t_np, time.time() - start)
tf.reset_default_graph()
with tf.Session() as sess:
x = tf.random.uniform(shape=(N, N), minval=MIN_VALUE, maxval=MAX_VALUE, dtype=tf.float32, seed=42)
y = tf.random.uniform(shape=(N, N), minval=MIN_VALUE, maxval=MAX_VALUE, dtype=tf.float32, seed=43)
A,B = sess.run([x,y])
A_placeholder = tf.placeholder(tf.float32, (N,N))
B_placeholder = tf.placeholder(tf.float32, (N,N))
res = tf.math.reduce_sum(tf.matmul(A_placeholder,B_placeholder) - tf.matmul(B_placeholder,A_placeholder))
start = time.time()
_ = sess.run(res, {A_placeholder:A, B_placeholder:B})
hist_t_tf = np.append(hist_t_tf, time.time() - start)
hist_t_np = hist_t_np * 1000
hist_t_tf = hist_t_tf * 1000
plt.figure(figsize=[16, 7])
plt.title('Зависимость времени вычисления "сумма элементов коммутатора квадратных матриц" для numpy и tensorflow')
plt.scatter(hist_n, hist_t_np, c='r', label='numpy')
plt.scatter(hist_n, hist_t_tf, c='b', label='tensorflow')
plt.legend(loc='upper left')
plt.xscale("log")
plt.ylim([min(hist_t_tf.min(), hist_t_np.min()), max(hist_t_tf.max(), hist_t_np.max())])
plt.xlabel("Размер матрицы, элементы")
plt.ylabel("Время вычисления, мс")
plt.show()
```
Время выполнения кода на tensorflow несколько больше соотвествующего кода на numpy для задач поиска угла между векторами и суммы квадртов диагонали матриц, однако tensorflow быстрее находит сумму коммутатора квадратных матриц начиная примерно с $10^3$.
Стоит отметить, что необходимо правильно настроить вычисления на tensorflow $-$ сбрасывать сессии, графы высичлений, использовать placeholder'ы, использовать float вместо int, не учитывать время создания случайных величин. Без всего этого время вычисления может быть очень большим и не превликательным для дальнейшей работы.
## Задание 2 -- градиенты и оптимайзеры (3 балла).
Продолжим работать с датасетом MNIST с размером картинок 8х8.
```
from sklearn.datasets import load_digits
mnist = load_digits()
X, y = mnist.data, mnist.target
n_labels = len(np.unique(y))
```
Многие алгоритмы оптимизации имплементированы в `tensorflow`. В этом задании мы сравним их при одинаковых параметрах, а также переберём разные параметры для одного алгоритма.
**Задание 2.1** (1.5 балла). Исследуйте вклад параметра momentum в методу `tf.train.MomentumOptimizer`. Для этого для разных значений momentum постройте графики значения функции потерь от номера итерации. При каких значениях momentum алгоритм сходится быстрее? Используйте `learning_rate=0.01`.
**Замечание**. В этом задании используется многоклассовая логистическая регрессия. Не меняйте код модели в ячейке ниже.
```
tf.reset_default_graph()
w = tf.Variable(np.ones((X.shape[1], n_labels)), dtype="float32")
X_input = tf.placeholder("float32", (None, X.shape[1]))
y_input = tf.placeholder("int32", (None,))
predicted = tf.nn.softmax(X_input @ w)
loss = tf.losses.log_loss(tf.one_hot(y_input, depth=n_labels), predicted)
def train(X, y, train_op, batch_size=16):
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
n_batch_train = len(X) // batch_size
for epoch in range(1):
loss_history = []
for b in range(n_batch_train):
_, loss_ = sess.run([train_op, loss], feed_dict={X_input: X[b*batch_size:(b+1)*batch_size],
y_input: y[b*batch_size:(
b+1)*batch_size]
})
loss_history.append(loss_)
return loss_history
plt.figure(figsize=[16, 7])
for momentum in np.geomspace(0.8, 0.9, num=5):
train_op = tf.train.MomentumOptimizer(
learning_rate=0.01, momentum=momentum).minimize(loss)
hist = train(X, y, train_op)
plt.plot(hist, label="Momentum {:.5f}".format(momentum))
print("Momentum {:.5f}\tSteps {}\t loss {}".format(momentum, len(hist), hist[-1]))
plt.title("Зависимость функции потерь для метода Momentum при различных momentum")
plt.yscale("logit")
plt.legend(loc='upper left')
plt.xlabel("# iteration")
plt.ylabel("Loss")
plt.show()
```
Ваш ответ: 0.87389
**Задание 2.2** (0.5 баллa). Исследуйте вклад `learning_rate`. Для этого для разных значений `learning_rate` постройте графики значения функции потерь от номера итерации. При каких значениях длины шага градиентного спуска алгоритм сходится быстрее? Используйте параметр метод MomentumOptimizer с параметром, который вы считаете лучшим по итогам предыдущего задания.
```
MOMENTUM_BEST = 0.87389
plt.figure(figsize=[16, 7])
for learning_rate in np.geomspace(0.006, 0.015, num=10):
train_op = tf.train.MomentumOptimizer(
learning_rate=learning_rate, momentum=MOMENTUM_BEST).minimize(loss)
hist = train(X, y, train_op)
plt.plot(hist, label="learning_rate {:.5f}".format(learning_rate))
print("learning_rate {:.5f}\tSteps {}\t loss {}".format(learning_rate, len(hist), hist[-1]))
plt.title("Зависимость функции потерь для метода Momentum при различных learning_rate")
# plt.yscale("logit")
plt.legend(loc='upper left')
plt.xlabel("# iteration")
plt.ylabel("Loss")
plt.show()
```
Ваш ответ: 0.00814
**Задание 2.3** (0.5 балла) Проделайте то же, что и в пункте выше, но используйте в качестве базового алгоритма оптимизации `Adam` с дефолтными параметрами.
```
plt.figure(figsize=[16, 7])
for learning_rate in np.geomspace(0.00975, 0.01304, num=10):
train_op = tf.train.AdamOptimizer(learning_rate=learning_rate).minimize(loss)
hist = train(X, y, train_op)
plt.plot(hist, label="learning_rate {:.5f}".format(learning_rate))
print("learning_rate {:.5f}\tSteps {}\t loss {}".format(learning_rate, len(hist), hist[-1]))
plt.title("Зависимость функции потерь для метода Adam при различных learning_rate")
# plt.yscale("logit")
plt.legend(loc='upper left')
plt.xlabel("# iteration")
plt.ylabel("Loss")
plt.show()
```
Ваш ответ: 0.01109
**Задание 2.4** (0.5 балла) Сравните алгоритмы `Adam` и `Momentum` для данной задачи. Какой показывает себя лучше?
```
MOMENTUM_BEST = 0.87389
LEARNING_RATE_ADAM = 0.01109
LEARNING_RATE_MOMENTUM = 0.00814
plt.figure(figsize=[16, 7])
train_op = tf.train.MomentumOptimizer(learning_rate=LEARNING_RATE_MOMENTUM, momentum=MOMENTUM_BEST).minimize(loss)
hist = train(X, y, train_op)
plt.plot(hist, label="MomentumOptimizer")
print("MomentumOptimizer \tSteps {}\t loss {}".format(len(hist), hist[-1]))
train_op = tf.train.AdamOptimizer(learning_rate=LEARNING_RATE_ADAM).minimize(loss)
hist = train(X, y, train_op)
plt.plot(hist, label="Adam")
print("Adam \t\t\tSteps {}\t loss {}".format(len(hist), hist[-1]))
plt.title("Зависимость функции потерь для метода Adam и Momentum для найденных гиперпараметров")
plt.legend(loc='upper left')
plt.xlabel("# iteration")
plt.ylabel("Loss")
plt.show()
```
Качество обоих алгоритмов примерно одинаковое, но итоговый результат лучше у MomentumOptimizer
## Задание 3 -- наша первая нейросеть, часть 2 (4 балла).
В этом задании мы напишем нейросеть для работы с датасетом MNIST размера 28х28. Исользовать можно только полносвязные (dense) слои! Для этого мы "вытянем" картинки 28х28 в длинный вектор размера 784.
```
from mnist import load_dataset
X_train, y_train, X_test, y_test, _, _ = load_dataset()
X_train = X_train.reshape(len(X_train), -1)
X_test = X_test.reshape(len(X_test), -1)
for i in [228, 1437, 322, 420, 69]:
plt.title(y_train[i])
plt.imshow(X_train[i].reshape((28, 28)))
plt.show()
```
Подберите архитектуру и алгоритм оптимизации так, чтобы значение accuracy на тестовой выборке было не менее 97.5.
```
from sklearn.metrics import accuracy_score
tf.reset_default_graph()
EPOCHS = 5
def train_and_validate(X_train, y_train, X_test, y_test, train_op, batch_size=16):
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
n_batch_train = len(X_train) // batch_size
n_batch_test = len(X_test) // batch_size
for epoch in range(EPOCHS):
loss_history_train = []
for b in range(n_batch_train):
_, loss_ = sess.run([train_op, loss], feed_dict={X_input: X_train[b*batch_size:(b+1)*batch_size],
y_input: y_train[b*batch_size:(
b+1)*batch_size]
})
loss_history_train.append(loss_)
for epoch in range(1):
loss_history_test = []
prediction_history = []
for b in range(n_batch_test):
loss_, predicted_ = sess.run([loss, predicted], feed_dict={X_input: X_test[b*batch_size:(b+1)*batch_size],
y_input: y_test[b*batch_size:(
b+1)*batch_size]
})
loss_history_test.append(loss_)
prediction_history += predicted_.argmax(-1).tolist()
print("Test accuracy: ", accuracy_score(y_test, prediction_history))
return loss_history_train, loss_history_test
for i in log_int_iterator(64, 1024, 2):
tf.reset_default_graph()
X_input = tf.placeholder("float32", (None, 784)) # dim = [batch_size, 784]
y_input = tf.placeholder("int32", (None,)) # dim = [batch_size,]
# <define architecture as a function of X_input>
layer2 = tf.layers.dense(X_input, i, activation=tf.nn.relu)
# layer2 = tf.layers.dense(layer1, 256, activation=tf.nn.relu)
# <define 10-class outputs>
logits = tf.layers.dense(layer2, n_labels)
predicted = tf.nn.softmax(logits)
# <define log loss with one-hot vector of labels
loss = tf.nn.softmax_cross_entropy_with_logits_v2(tf.one_hot(y_input, depth=n_labels), logits)
# <define train operation here>
# train_op = tf.train.AdamOptimizer(learning_rate=LEARNING_RATE_ADAM).minimize(loss)
train_op = tf.train.AdamOptimizer().minimize(loss)
# train_op = tf.train.MomentumOptimizer(learning_rate=LEARNING_RATE_MOMENTUM, momentum=MOMENTUM_BEST).minimize(loss)
print("="*60, '\n', "Layer size:", i)
loss_history_train, loss_history_test = train_and_validate(
X_train, y_train, X_test, y_test, train_op)
```
Наилучшим образом показал себя слой из 1024 нейронов, но и необходимое качество было достигнуто и с 256.
Необходимое качество было достигнуто с помошью AdamOptimizer со стандартными параметрами, подобранные значения в задании 2 давали меньшее качество
Количество эпох было увеличено до 5 для дообучения модели и подбора точных весов, дальнейшее увеличение не дает заметного изменения качества модели.
| github_jupyter |
# How to remove K2 motion systematics with SFF?
You can use `lightkurve` to remove the motion from K2 data. Targets in K2 data move over multiple pixels during the exposure due to thruster firings. This can be corrected using the self flat fielding method (SFF), which you can read more about [here](04-replicate-vanderburg-2014-k2sff.html) On this page we will show a short tutorial demonstrating how you can apply the method on your light curves.
Let's download a K2 light curve of an exoplanet host star. Remember that for K2 data, we can use the same Kepler functions `KeplerTargetPixelFile` and `KeplerLightCurveFile` as they are the same instrument.
```
from lightkurve import search_lightcurvefile
lcf = search_lightcurvefile(247887989).download() # returns a KeplerLightCurveFile
lc = lcf.PDCSAP_FLUX # returns a KeplerLightCurve
#Remove nans and outliers
lc = lc.remove_nans().remove_outliers()
#Remove long term trends
lc = lc.flatten(window_length=401)
lc.scatter();
```
This light curve is of the object [K2-133b](https://exoplanetarchive.ipac.caltech.edu/cgi-bin/DisplayOverview/nph-DisplayOverview?objname=K2-133+b&type=CONFIRMED_PLANET), which has a period of 3.0712 days. Let's plot the folded version of it to see what it looks like.
```
lc.fold(period=3.0712).scatter();
```
We can see the hint of an exoplanet transit close to the center, but the motion of the spacecraft has made it difficult to make out above the noise. We can use the `correct` function on the lightcurve to remove this motion. An in-depth look into how the algorithm works can be found [here](http://lightkurve.keplerscience.org/tutorials/motion-correction/replicate-vanderburg-2014-k2sff.html). You can pass the following keywords to the `correct` function:
* `polyorder` : *int*
Degree of the polynomial which will be used to fit one centroid as a function of the other.
* `niters` : *int*
Number of iterations
* `bins` : *int*
Number of bins to be used to create the piece-wise interpolation of arclength vs flux correction.
* `windows` : *int*
Number of windows to subdivide the data. The SFF algorithm is run independently in each window.
This [tutorial](gully-tutorial) will teach you more about how to tune these parameters. For this problem, we will use the defaults, but increase the number of windows to 20.
```
corr_lc = lc.to_corrector("sff").correct(windows=20)
```
Now when we compare the two light curves we can see the clear signal from the exoplanet.
```
ax = lc.fold(period=3.0712).scatter(color='red', alpha=0.5, label='With Motion')
ax = corr_lc.fold(period=3.0712).scatter(ax=ax, color='blue', alpha=0.5, label='Motion Corrected');
```
| github_jupyter |
# Using the frame context in the TIMIT MLP model
This notebook is an extension of the MLP_TIMIT demo which takes a context of many frames at input to model the same output. So if we have a phoneme, say 'a', instead of just using one vector of 26 features to recognize it, we provide several frames of 26 features before and after the one we are looking at, in order to capture its context.
This technique helps greatly improve the the quality of the solution, but isn't as scalable as some other solutions. First of all, the greater the context, the more parameters we need to determine. The bigger the model, the more data is required to accurately appraise all the parameters.
One solution would be to use tied weights, rather then a classical dense layer in such a way that different frames (within the context) use the same set of weights, so the number of weights is kept constant even though we use a larger context.
Furthermore, the model assumes a context of a specific size. It would be nice if the size is unlimited. Again, this would probably make the model impractical if we use a standard dense layer, but could work with the tied weights technique.
Another way of looking at the tied weights solution that has an unlimited context is simply as an RNN. In fact, most implementions of BPTT (used to train RNN) simply unroll the training loop in time and treat the model as a simple MLP with tied weights. This works quite well, but has other issues that are solved using more advanced topologies (LSTM, GRU) which will be dicussed in other notebooks.
In this notebook, we will take an MLP which has an input context of 10 frames on the left and the right side of the analyzed frame. This is done in order to reproduce the resutls from the same paper and thesis as in the MLP_TIMIT notebook.
We begin with the same introductory code as in the previous notebook:
```
import os
os.environ['CUDA_VISIBLE_DEVICES']='1'
import numpy as np
from keras.models import Sequential
from keras.layers.core import Dense, Activation, Reshape
from keras.optimizers import Adam, SGD
from IPython.display import clear_output
from tqdm import *
```
## Loading the data
```
import sys
sys.path.append('../python')
from data import Corpus, History
train=Corpus('../data/TIMIT_train.hdf5',load_normalized=True,merge_utts=False)
dev=Corpus('../data/TIMIT_dev.hdf5',load_normalized=True,merge_utts=False)
test=Corpus('../data/TIMIT_test.hdf5',load_normalized=True,merge_utts=False)
tr_in,tr_out_dec=train.get()
dev_in,dev_out_dec=dev.get()
tst_in,tst_out_dec=test.get()
for u in range(tr_in.shape[0]):
tr_in[u]=tr_in[u][:,:26]
for u in range(dev_in.shape[0]):
dev_in[u]=dev_in[u][:,:26]
for u in range(tst_in.shape[0]):
tst_in[u]=tst_in[u][:,:26]
```
## Global training parameters
```
input_dim=tr_in[0].shape[1]
output_dim=61
hidden_num=250
epoch_num=1000
```
### 1-hot output
```
def dec2onehot(dec):
ret=[]
for u in dec:
assert np.all(u<output_dim)
num=u.shape[0]
r=np.zeros((num,output_dim))
r[range(0,num),u]=1
ret.append(r)
return np.array(ret)
tr_out=dec2onehot(tr_out_dec)
dev_out=dec2onehot(dev_out_dec)
tst_out=dec2onehot(tst_out_dec)
```
### Adding frame context
Here we add the frame context. The number 10 is taken from the paper thesis as: *symmetrical
time-windows from 0 to 10 frames*. Now I'm not 100% sure (and it's not explained anywhere), but I assume this means 10 frames on the left and 10 on the right (i.e. symmetrical), which gives 21 frames alltogether. It's written elsewhere that 0 means no context and uses one frame.
In Keras/Python we implement this in a slightly roundabout way: instead of duplicating the data explicitly, we merely make a 3D array that contains the references to the same data ranges in different cells. In other words, if we make an array where each utterance has a shape $(time\_steps, context*frame\_size)$, I think it would take more memory than by using the shape $(time\_steps,context,frame\_size)$, because in the latter case the same vector (located somewhere in the memory) can be reused in different cotenxts and time steps.
```
#adds context to data
ctx_fr=10
ctx_size=2*ctx_fr+1
def ctx(data):
ret=[]
for utt in data:
l=utt.shape[0]
ur=[]
for t in range(l):
f=[]
for s in range(t-ctx_fr,t+ctx_fr+1):
if(s<0):
s=0
if(s>=l):
s=l-1
f.append(utt[s,:])
ur.append(f)
ret.append(np.array(ur))
return np.array(ret)
tr_in=ctx(tr_in)
dev_in=ctx(dev_in)
tst_in=ctx(tst_in)
print tr_in.shape
print tr_in[0].shape
```
## Model definition
Since we have an input as a 3D shape, we use a Reshape layer at the start of the model to convert the input frames into a flat vector. Again, this is to save a little memory at the cost of time it takes to reshape the input. Not sure if its worth it or if it even works as intended (ie. saving memory).
Evertyhing else here is the same as with the standard MLP except for the learning rate which has to be lower in order to reproduce the same results as in the thesis.
```
model = Sequential()
model.add(Reshape(input_shape=(ctx_size,input_dim),target_shape=(ctx_size*input_dim,)))
model.add(Dense(output_dim=hidden_num))
model.add(Activation('sigmoid'))
model.add(Dense(output_dim=output_dim))
model.add(Activation('softmax'))
optimizer= SGD(lr=1e-3,momentum=0.9,nesterov=False)
loss='categorical_crossentropy'
metrics=['accuracy']
model.compile(loss=loss, optimizer=optimizer,metrics=['accuracy'])
```
## Training
```
from random import shuffle
tr_hist=History('Train')
dev_hist=History('Dev')
tst_hist=History('Test')
tr_it=range(tr_in.shape[0])
for e in range(epoch_num):
print 'Epoch #{}/{}'.format(e+1,epoch_num)
sys.stdout.flush()
shuffle(tr_it)
for u in tqdm(tr_it):
l,a=model.train_on_batch(tr_in[u],tr_out[u])
tr_hist.r.addLA(l,a,tr_out[u].shape[0])
clear_output()
tr_hist.log()
for u in range(dev_in.shape[0]):
l,a=model.test_on_batch(dev_in[u],dev_out[u])
dev_hist.r.addLA(l,a,dev_out[u].shape[0])
dev_hist.log()
for u in range(tst_in.shape[0]):
l,a=model.test_on_batch(tst_in[u],tst_out[u])
tst_hist.r.addLA(l,a,tst_out[u].shape[0])
tst_hist.log()
print 'Done!'
```
## Plotting progress
These can be handy for debugging. If you draw the graph with using different hyperparameters you can establish if it underfits (i.e. the values are still decreasing at the end of the training) or overfits (the minimum is reached earlier and dev/test values begin increasing as train continues to decrease). In this case, you can see hoe the graph changes with different learning rate values. It's impossible to achieve a single optimal value, but this one seems to be fairly good.
```
import matplotlib.pyplot as P
%matplotlib inline
fig,ax=P.subplots(2,sharex=True,figsize=(12,10))
ax[0].set_title('Loss')
ax[0].plot(tr_hist.loss,label='Train')
ax[0].plot(dev_hist.loss,label='Dev')
ax[0].plot(tst_hist.loss,label='Test')
ax[0].legend()
ax[0].set_ylim((0.8,2))
ax[1].set_title('PER %')
ax[1].plot(100*(1-np.array(tr_hist.acc)),label='Train')
ax[1].plot(100*(1-np.array(dev_hist.acc)),label='Dev')
ax[1].plot(100*(1-np.array(tst_hist.acc)),label='Test')
ax[1].legend()
ax[1].set_ylim((32,42))
```
## Final result
Here we reached the value from the thesis just fine, but we used a different learning rate. For some reason, the value from the thesis underfits by a great margin. Not sure if it's a mistake in the thesis or a consequence
```
print 'Min test PER: {:%}'.format(1-np.max(tst_hist.acc))
print 'Min dev PER epoch: #{}'.format((np.argmax(dev_hist.acc)+1))
print 'Test PER on min dev: {:%}'.format(1-tst_hist.acc[np.argmax(dev_hist.acc)])
```
Just as before we can check what epoch we reached the optimum.
```
wer=0.36999999
print 'Epoch where PER reached {:%}: #{}'.format(wer,np.where((1-np.array(tst_hist.acc))<wer)[0][0])
```
## Checking the accuracy calculation
When computing the final loss value, we simply measure the mean of the consecutive batch loss values, because we assume that weight updates are performed once per batch and the mean loss of the whole batch is used in the cross entropy to asses the model (just like in MSE).
With accuracy, however, it's not that simple as using the mean of all the batch accuracies. What we use instad is a weighted average where the weights are determined by the length of each batch/uterance. To make sure this is correct, I do a simple experiment here where I manually count the errors and sample amounts using the predict method. We can see that the values are identical, so using the weighted average is fine.
```
err=0
cnt=0
for u in range(tst_in.shape[0]):
p=model.predict_on_batch(tst_in[u])
c=np.argmax(p,axis=-1)
err+=np.sum(c!=tst_out_dec[u])
cnt+=tst_out[u].shape[0]
print 'Manual PER: {:%}'.format(err/float(cnt))
print 'PER using average: {:%}'.format(1-tst_hist.acc[-1])
```
| github_jupyter |
# This code assesses the outputs of VESIcal for the Iacono-Marziano model.
First, we compare VESIcal saturation pressure results to the calibration data for this model. Then, we compare VESIcal outputs to a number of the compositoins in the calibratoin dataset for a range of H$_2$O and CO$_2$ contents with those calculated from the web calculator hosted at http://calcul-isto.cnrs-orleans.fr/
```
import VESIcal as v
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from IPython.display import display, HTML
import pandas as pd
import matplotlib as mpl
import seaborn as sns
%matplotlib inline
plt.rcParams["font.family"] = 'arial'
plt.rcParams["mathtext.default"] = "regular"
plt.rcParams["mathtext.fontset"] = "dejavusans"
%matplotlib inline
sns.set(style="ticks", context="poster",rc={"grid.linewidth": 1,"xtick.major.width": 1,"ytick.major.width": 1, 'patch.edgecolor': 'black'})
plt.style.use("seaborn-colorblind")
#jtplot.style()
mpl.rcParams['patch.linewidth'] = 1
mpl.rcParams['axes.linewidth'] = 1 # set the value globally
myfile_cal = v.ExcelFile('Iacono_MixedH2O-CO2Dataset.xlsx', sheet_name=0, input_type='wtpercent')
myfile_web = v.ExcelFile('Iacono_MixedH2O-CO2Dataset.xlsx', sheet_name=1, input_type='wtpercent') # This sheet is the pressures calculated using the web calculator
data_cal = myfile_cal.data
data_cal.head()
data_web = myfile_web.data
data_web.head()
satPs_wtemps_Iacono_cal= myfile_cal.calculate_saturation_pressure(temperature="Temp", model='IaconoMarziano')
satPs_wtemps_Iacono_web= myfile_web.calculate_saturation_pressure(temperature="Temp", model='IaconoMarziano')
satPs_wtemps_Iacono_web.head()
X=data_cal['Press'].values.reshape(-1, 1)
Y=satPs_wtemps_Iacono_cal['SaturationP_bars_VESIcal'].values.reshape(-1, 1)
#mask = ~np.isnan(X) & ~np.isnan(Y)
#X_noNan=X[mask].values.reshape(-1, 1)
#Y_noNan=Y[mask].values.reshape(-1, 1)
from sklearn.linear_model import LinearRegression
from sklearn.metrics import r2_score
lr=LinearRegression()
lr.fit(X,Y)
Y_pred=lr.predict(X)
fig, ax1 = plt.subplots(figsize = (12,8)) # adjust dimensions of figure here
font = {'family': 'sans-serif',
'color': 'black',
'weight': 'normal',
'size': 20,
}
ax1.set_title('Calibration dataset experimental pressures vs. VESIcal saturation pressures',
fontdict= font, pad = 15)
plt.xlabel('Experimental Pressure (Calibration dataset)', fontdict=font, labelpad = 15)
plt.ylabel('P$_{Sat}$ VESIcal', fontdict=font, labelpad = 15)
plt.plot(X,Y_pred, color='red', linewidth=1)
plt.scatter(data_cal['Press'], satPs_wtemps_Iacono_cal['SaturationP_bars_VESIcal'], s=100, edgecolors='gray', facecolors='silver', marker='o', label = 'Eguchi')
#plt.plot([0, 12000], [0, 12000])
I='Intercept= ' + str(np.round(lr.intercept_, 3))[1:-1]
G='Gradient= ' + str(np.round(lr.coef_, 5))[2:-2]
R='R$^2$= ' + str(np.round(r2_score(Y, Y_pred), 5))
plt.text(1000, 8000, I, fontsize=15)
plt.text(1000, 7000, G, fontsize=15)
plt.text(1000, 6000, R, fontsize=15)
```
# Comparison of VESIcal saturation pressures with those from the web-app
```
X_web=data_web['App calculator P bar']
Y_web=satPs_wtemps_Iacono_web['SaturationP_bars_VESIcal']
mask = ~np.isnan(X_web) & ~np.isnan(Y_web)
X_noNan=X_web[mask].values.reshape(-1, 1)
Y_noNan=Y_web[mask].values.reshape(-1, 1)
lr=LinearRegression()
lr.fit(X_noNan,Y_noNan)
Y_pred_web=lr.predict(X_noNan)
fig, ax1 = plt.subplots(figsize = (12,8)) # adjust dimensions of figure here
font = {'family': 'sans-serif',
'color': 'black',
'weight': 'normal',
'size': 20,
}
ax1.set_title('Comparison of web app vs. VESIcal',
fontdict= font, pad = 15)
plt.xlabel('P$_{Sat}$ Web app', fontdict=font, labelpad = 15)
plt.ylabel('P$_{Sat}$ VESIcal', fontdict=font, labelpad = 15)
plt.plot(X_noNan,Y_pred_web, color='red', linewidth=1)
plt.scatter(X_noNan, Y_noNan, s=100, edgecolors='gray', facecolors='silver', marker='o', label = 'Eguchi')
#plt.plot([0, 12000], [0, 12000])
I='Intercept= ' + str(np.round(lr.intercept_, 3))[1:-1]
G='Gradient= ' + str(np.round(lr.coef_, 5))[2:-2]
R='R$^2$= ' + str(np.round(r2_score(Y_noNan, Y_pred_web), 5))
plt.text(1000, 8000, I, fontsize=15)
plt.text(1000, 7000, G, fontsize=15)
plt.text(1000, 6000, R, fontsize=15)
```
# Histogram showing the absolute offset between VESIcal and the Mac App in bars
```
fig, ax1 = plt.subplots(figsize = (12,8))
font = {'family': 'sans-serif',
'color': 'black',
'weight': 'normal',
'size': 20,
}
ax1.set_title('Difference between the Web App and VESIcal in bars',
fontdict= font, pad = 15)
plt.xlabel('P$_{Sat}$ (Web app) - P$_{Sat}$ (VESIcal), bars', fontdict=font, labelpad = 15)
plt.ylabel(' Number of measurements', fontdict=font, labelpad = 15)
plt.hist(X_noNan- Y_noNan, color='red', bins=int(180/5))
plt.plot([0, 0], [0, 12], color='k', linestyle='dashed')
ax1.set_ylim([0, 12])
```
# What does offset correlate with
```
TestVariable=data_web['CO2']
TestVariable=TestVariable[mask]
TestVariable2=data_web['H2O']
TestVariable2=TestVariable2[mask]
logical=TestVariable>0
fig, ax1 = plt.subplots(figsize = (12,8))
font = {'family': 'sans-serif',
'color': 'black',
'weight': 'normal',
'size': 20,
}
ax1.set_title('Difference between the Web App and VESIcal in bars',
fontdict= font, pad = 15)
plt.xlabel('CO2 (ppm)', fontdict=font, labelpad = 15)
plt.ylabel('100* App Calculator/VESIcal', fontdict=font, labelpad = 15)
plt.scatter(10**4*TestVariable, 100*X_noNan/Y_noNan, marker='o', c=TestVariable2)
#plt.scatter(10**4*TestVariable[logical], 100*X_noNan[logical]/Y_noNan[logical], color='blue')
cbar=plt.colorbar()
cbar.set_label('H$_2$O')
TestVariable2=data_web['H2O']
TestVariable2=TestVariable2[mask]
fig, ax1 = plt.subplots(figsize = (12,8))
font = {'family': 'sans-serif',
'color': 'black',
'weight': 'normal',
'size': 20,
}
ax1.set_title('Difference between the Web App and VESIcal in bars',
fontdict= font, pad = 15)
plt.xlabel('H2O', fontdict=font, labelpad = 15)
plt.ylabel('100* App Calculator/VESIcal', fontdict=font, labelpad = 15)
plt.scatter(TestVariable2, 100*X_noNan/Y_noNan, color='red')
logical=TestVariable2>0
plt.scatter(TestVariable2[logical], 100*X_noNan[logical]/Y_noNan[logical], color='blue')
```
# Histogram of Percentage Error
```
fig, ax1 = plt.subplots(figsize = (12,8)) # adjust dimensions of figure here
#ax1.set_xlim([35, 100]) # adjust x limits here if you want to focus on a specific part of compostional space
#ax1.set_ylim([0, 25]) # adjust y limits here
font = {'family': 'sans-serif',
'color': 'black',
'weight': 'normal',
'size': 20,
}
ax1.set_title('% Difference between the Mac App and VESIcal in bars',
fontdict= font, pad = 15)
plt.xlabel('100*(P$_{Sat}$ Web Calculator)/(P$_{Sat}$ VESIcal)', fontdict=font, labelpad = 15)
plt.ylabel(' Number of measurements', fontdict=font, labelpad = 15)
plt.hist(100.*X_noNan/Y_noNan, color='red', bins=int(180/5)) #
plt.plot([100, 100], [0, 22], color='k', linestyle='dashed')
ax1.set_ylim([0, 22])
myfile_web.save_excelfile(filename='IaconoTesting2.xlsx', calculations=[satPs_wtemps_Iacono_web], sheet_name=['web'])
```
# Dataset of 3 vectors of increasing CO2, one with no water, one with 0.5 wt%, the other with 1 T 1200x
```
myfile_test = v.ExcelFile('Iacono_MixedH2O-CO2Dataset.xlsx', sheet_name=2, input_type='wtpercent')
data_test = myfile_test.data
data_test.head()
satPs_wtemps_Iacono_test= myfile_test.calculate_saturation_pressure(temperature="Temp", model='IaconoMarziano')
satPs_wtemps_Iacono_test.head()
# X=Y comparison
fig, (ax1) = plt.subplots(1, 1, figsize = (17,8))
font = {'family': 'sans-serif',
'color': 'black',
'weight': 'normal',
'size': 20,
}
ax1.set_title('VESICAL',
fontdict= font, pad = 15)
ax2.set_title('WebApp',
fontdict= font, pad = 15)
ax1.set_xlabel('Saturation P (web)', fontdict=font, labelpad = 15)
ax1.set_ylabel(' Saturation P (VESIcal)', fontdict=font, labelpad = 15)
ax1.plot([0, 1300], [0, 1300], color='k', zorder=1, linewidth=0.5)
ax1.scatter(satPs_wtemps_Iacono_test['Press'].loc['NoH2O'], satPs_wtemps_Iacono_test['SaturationP_bars_VESIcal'].loc['NoH2O'], marker='o',s=20, facecolor='red', label='no water', zorder=5)
ax1.scatter(satPs_wtemps_Iacono_test['Press'].loc['H2O_05'], satPs_wtemps_Iacono_test['SaturationP_bars_VESIcal'].loc['H2O_05'], marker='o', s=20, facecolor='blue', label='0.5 wt% H2O', zorder=6)
ax1.scatter(satPs_wtemps_Iacono_test['Press'].loc['H2O_1'], satPs_wtemps_Iacono_test['SaturationP_bars_VESIcal'].loc['H2O_1'], marker='o', s=20, facecolor='gray', label='1 wt% H2O', zorder=7)
ax1.scatter(satPs_wtemps_Iacono_test['Press'].loc['H2O_15'], satPs_wtemps_Iacono_test['SaturationP_bars_VESIcal'].loc['H2O_15'], marker='o', s=20, facecolor='green', label='1.5 wt% H2O', zorder=8)
ax1.legend()
fig, (ax1, ax2) = plt.subplots(1, 2, figsize = (17,8))
font = {'family': 'sans-serif',
'color': 'black',
'weight': 'normal',
'size': 20,
}
ax1.set_title('VESICAL',
fontdict= font, pad = 15)
ax2.set_title('WebApp',
fontdict= font, pad = 15)
ax1.set_xlabel('CO2 (wt%)', fontdict=font, labelpad = 15)
ax1.set_ylabel(' Saturation P bars', fontdict=font, labelpad = 15)
ax2.set_xlabel('CO2 (wt%)', fontdict=font, labelpad = 15)
ax1.scatter(satPs_wtemps_Iacono_test['CO2'].loc['NoH2O'], satPs_wtemps_Iacono_test['SaturationP_bars_VESIcal'].loc['NoH2O'], marker='o',s=20, facecolor='red', label='no water')
ax1.scatter(satPs_wtemps_Iacono_test['CO2'].loc['H2O_05'], satPs_wtemps_Iacono_test['SaturationP_bars_VESIcal'].loc['H2O_05'], marker='o', s=20, facecolor='blue', label='0.5 wt% H2O')
ax1.scatter(satPs_wtemps_Iacono_test['CO2'].loc['H2O_1'], satPs_wtemps_Iacono_test['SaturationP_bars_VESIcal'].loc['H2O_1'], marker='o', s=20, facecolor='gray', label='1 wt% H2O')
ax1.scatter(satPs_wtemps_Iacono_test['CO2'].loc['H2O_15'], satPs_wtemps_Iacono_test['SaturationP_bars_VESIcal'].loc['H2O_15'], marker='o', s=20, facecolor='green', label='1.5 wt% H2O')
ax1.legend()
ax2.scatter(satPs_wtemps_Iacono_test['CO2'].loc['NoH2O'], satPs_wtemps_Iacono_test['Press'].loc['NoH2O'], marker='o',s=20, facecolor='red', label='no water')
ax2.scatter(satPs_wtemps_Iacono_test['CO2'].loc['H2O_05'], satPs_wtemps_Iacono_test['Press'].loc['H2O_05'], marker='o', s=20, facecolor='blue', label='0.5 wt% H2O')
ax2.scatter(satPs_wtemps_Iacono_test['CO2'].loc['H2O_1'], satPs_wtemps_Iacono_test['Press'].loc['H2O_1'], marker='o', s=20, facecolor='gray', label='1 wt% H2O')
ax2.scatter(satPs_wtemps_Iacono_test['CO2'].loc['H2O_15'], satPs_wtemps_Iacono_test['Press'].loc['H2O_15'], marker='o', s=20, facecolor='green', label='1.5 wt% H2O')
# Comparison of models
fig, (ax1) = plt.subplots(1, 1, figsize = (17,8))
font = {'family': 'sans-serif',
'color': 'black',
'weight': 'normal',
'size': 20,
}
ax1.set_title('VESICAL',
fontdict= font, pad = 15)
ax2.set_title('WebApp',
fontdict= font, pad = 15)
ax1.set_xlabel('CO2 (wt%)', fontdict=font, labelpad = 15)
ax1.set_ylabel('% Diff Saturation P bars VESICAl/app', fontdict=font, labelpad = 15)
ax2.set_xlabel('CO2 (wt%)', fontdict=font, labelpad = 15)
ax1.scatter(satPs_wtemps_Iacono_test['CO2'].loc['NoH2O'], 100.*satPs_wtemps_Iacono_test['SaturationP_bars_VESIcal'].loc['NoH2O']/satPs_wtemps_Iacono_test['Press'].loc['NoH2O'], marker='o',s=20, facecolor='red', label='no water')
ax1.scatter(satPs_wtemps_Iacono_test['CO2'].loc['H2O_05'], 100.*satPs_wtemps_Iacono_test['SaturationP_bars_VESIcal'].loc['H2O_05']/satPs_wtemps_Iacono_test['Press'].loc['H2O_05'], marker='o', s=20, facecolor='blue', label='0.5 wt% H2O')
ax1.scatter(satPs_wtemps_Iacono_test['CO2'].loc['H2O_1'], 100.*satPs_wtemps_Iacono_test['SaturationP_bars_VESIcal'].loc['H2O_1']/satPs_wtemps_Iacono_test['Press'].loc['H2O_1'], marker='o', s=20, facecolor='gray', label='1 wt% H2O')
ax1.scatter(satPs_wtemps_Iacono_test['CO2'].loc['H2O_15'], 100.*satPs_wtemps_Iacono_test['SaturationP_bars_VESIcal'].loc['H2O_15']/satPs_wtemps_Iacono_test['Press'].loc['H2O_15'], marker='o', s=20, facecolor='green', label='1.5 wt% H2O')
ax1.legend()
```
# 1100C
```
myfile_test2 = v.ExcelFile('Iacono_MixedH2O-CO2Dataset.xlsx', sheet_name=3, input_type='wtpercent')
data_test2 = myfile_test2.data
data_test2.head()
satPs_wtemps_Iacono_test2= myfile_test2.calculate_saturation_pressure(temperature="Temp", model='IaconoMarziano')
satPs_wtemps_Iacono_test2.head()
fig, (ax1) = plt.subplots(1, 1, figsize = (17,8))
font = {'family': 'sans-serif',
'color': 'black',
'weight': 'normal',
'size': 20,
}
ax1.set_title('VESICAL',
fontdict= font, pad = 15)
ax2.set_title('WebApp',
fontdict= font, pad = 15)
ax1.set_xlabel('Saturation P (web)', fontdict=font, labelpad = 15)
ax1.set_ylabel(' Saturation P (VESIcal)', fontdict=font, labelpad = 15)
ax1.plot([0, 1300], [0, 1300], color='k', zorder=1, linewidth=0.5)
ax1.scatter(satPs_wtemps_Iacono_test2['Press'].loc['NoH2O'], satPs_wtemps_Iacono_test2['SaturationP_bars_VESIcal'].loc['NoH2O'], marker='o',s=20, facecolor='red', label='no water', zorder=5)
ax1.scatter(satPs_wtemps_Iacono_test2['Press'].loc['H2O_05'], satPs_wtemps_Iacono_test2['SaturationP_bars_VESIcal'].loc['H2O_05'], marker='o', s=20, facecolor='blue', label='0.5 wt% H2O', zorder=6)
ax1.scatter(satPs_wtemps_Iacono_test2['Press'].loc['H2O_1'], satPs_wtemps_Iacono_test2['SaturationP_bars_VESIcal'].loc['H2O_1'], marker='o', s=20, facecolor='gray', label='1 wt% H2O', zorder=7)
ax1.scatter(satPs_wtemps_Iacono_test2['Press'].loc['H2O_15'], satPs_wtemps_Iacono_test2['SaturationP_bars_VESIcal'].loc['H2O_15'], marker='o', s=20, facecolor='green', label='1.5 wt% H2O', zorder=8)
ax1.legend()
# Comparison of models
fig, (ax1) = plt.subplots(1, 1, figsize = (17,8))
font = {'family': 'sans-serif',
'color': 'black',
'weight': 'normal',
'size': 20,
}
ax1.set_title('VESICAL',
fontdict= font, pad = 15)
ax2.set_title('WebApp',
fontdict= font, pad = 15)
ax1.set_xlabel('CO2 (wt%)', fontdict=font, labelpad = 15)
ax1.set_ylabel('% Diff Saturation P bars VESICAl/app', fontdict=font, labelpad = 15)
ax2.set_xlabel('CO2 (wt%)', fontdict=font, labelpad = 15)
ax1.scatter(satPs_wtemps_Iacono_test2['CO2'].loc['NoH2O'], 100.*satPs_wtemps_Iacono_test2['SaturationP_bars_VESIcal'].loc['NoH2O']/satPs_wtemps_Iacono_test2['Press'].loc['NoH2O'], marker='o',s=20, facecolor='red', label='no water')
ax1.scatter(satPs_wtemps_Iacono_test2['CO2'].loc['H2O_05'], 100.*satPs_wtemps_Iacono_test2['SaturationP_bars_VESIcal'].loc['H2O_05']/satPs_wtemps_Iacono_test2['Press'].loc['H2O_05'], marker='o', s=20, facecolor='blue', label='0.5 wt% H2O')
ax1.scatter(satPs_wtemps_Iacono_test2['CO2'].loc['H2O_1'], 100.*satPs_wtemps_Iacono_test2['SaturationP_bars_VESIcal'].loc['H2O_1']/satPs_wtemps_Iacono_test2['Press'].loc['H2O_1'], marker='o', s=20, facecolor='gray', label='1 wt% H2O')
#ax1.scatter(satPs_wtemps_Iacono_test2['CO2'].loc['H2O_15'], 100.*satPs_wtemps_Iacono_test2['SaturationP_bars_VESIcal'].loc['H2O_15']/satPs_wtemps_Iacono_test2['Press'].loc['H2O_15'], marker='o', s=20, facecolor='green', label='1.5 wt% H2O')
#ax1.scatter(satPs_wtemps_Iacono_test['CO2'].loc['NoH2O'], 100.*satPs_wtemps_Iacono_test['SaturationP_bars_VESIcal'].loc['NoH2O']/satPs_wtemps_Iacono_test['Press'].loc['NoH2O'], marker='^',s=20, facecolor='red', label='no water')
#ax1.scatter(satPs_wtemps_Iacono_test['CO2'].loc['H2O_05'], 100.*satPs_wtemps_Iacono_test['SaturationP_bars_VESIcal'].loc['H2O_05']/satPs_wtemps_Iacono_test['Press'].loc['H2O_05'], marker='^', s=20, facecolor='blue', label='0.5 wt% H2O')
#ax1.scatter(satPs_wtemps_Iacono_test['CO2'].loc['H2O_1'], 100.*satPs_wtemps_Iacono_test['SaturationP_bars_VESIcal'].loc['H2O_1']/satPs_wtemps_Iacono_test['Press'].loc['H2O_1'], marker='^', s=20, facecolor='gray', label='1 wt% H2O')
#ax1.scatter(satPs_wtemps_Iacono_test['CO2'].loc['H2O_15'], 100.*satPs_wtemps_Iacono_test['SaturationP_bars_VESIcal'].loc['H2O_15']/satPs_wtemps_Iacono_test['Press'].loc['H2O_15'], marker='^', s=20, facecolor='green', label='1.5 wt% H2O')
ax1.legend()
# Diff composition
myfile_test3 = v.ExcelFile('Iacono_MixedH2O-CO2Dataset.xlsx', sheet_name=4, input_type='wtpercent')
data_test3 = myfile_test3.data
data_test3.head()
satPs_wtemps_Iacono_test3= myfile_test3.calculate_saturation_pressure(temperature="Temp", model='IaconoMarziano')
satPs_wtemps_Iacono_test3.head()
# Comparison of models
fig, (ax1) = plt.subplots(1, 1, figsize = (17,8))
font = {'family': 'sans-serif',
'color': 'black',
'weight': 'normal',
'size': 20,
}
ax1.set_title('VESICAL',
fontdict= font, pad = 15)
ax2.set_title('WebApp',
fontdict= font, pad = 15)
ax1.set_xlabel('CO2 (wt%)', fontdict=font, labelpad = 15)
ax1.set_ylabel('% Diff Saturation P bars VESICAl/app', fontdict=font, labelpad = 15)
ax2.set_xlabel('CO2 (wt%)', fontdict=font, labelpad = 15)
ax1.scatter(satPs_wtemps_Iacono_test3['CO2'].loc['NoH2O'], 100.*satPs_wtemps_Iacono_test3['SaturationP_bars_VESIcal'].loc['NoH2O']/satPs_wtemps_Iacono_test3['Press'].loc['NoH2O'], marker='o',s=20, facecolor='red', label='no water')
ax1.scatter(satPs_wtemps_Iacono_test3['CO2'].loc['H2O_05'], 100.*satPs_wtemps_Iacono_test3['SaturationP_bars_VESIcal'].loc['H2O_05']/satPs_wtemps_Iacono_test3['Press'].loc['H2O_05'], marker='o', s=20, facecolor='blue', label='0.5 wt% H2O')
ax1.scatter(satPs_wtemps_Iacono_test3['CO2'].loc['H2O_1'], 100.*satPs_wtemps_Iacono_test3['SaturationP_bars_VESIcal'].loc['H2O_1']/satPs_wtemps_Iacono_test3['Press'].loc['H2O_1'], marker='o', s=20, facecolor='gray', label='1 wt% H2O')
#ax1.scatter(satPs_wtemps_Iacono_test3['CO2'].loc['H2O_15'], 100.*satPs_wtemps_Iacono_test3['SaturationP_bars_VESIcal'].loc['H2O_15']/satPs_wtemps_Iacono_test['Press'].loc['H2O_15'], marker='o', s=20, facecolor='green', label='1.5 wt% H2O')
ax1.legend()
```
| github_jupyter |
# music21: A Toolkit for Comupter-Aided Musicology
## Some examples to test basic music21 functionalities
This is a Jupyter notebook created by [@musicenfanthen](https://github.com/musicEnfanthen) and [@aWilsonandmore](https://github.com/aWilsonandmore) to work with some basic functionalities of music21 (http://web.mit.edu/music21/). For more information on Jupyter notebooks go to http://jupyter.org/.
To execute a block of code in this notebook, click in the cell and press `Shift+Enter`.
To get help on any music21 routine, click on it and press `Shift+Tab`.
### Imports and setup
To use music21 in this notebook and python, you have to import all (\*) routines from music21 at first with the following command.
<div class="alert alert-block alert-warning">
"You’ll probably get a few warnings that you’re missing some optional modules. That’s okay. If you get a warning that “no module named music21” then something probably went wrong above." (Source: http://web.mit.edu/music21/doc/usersGuide/usersGuide_01_installing.html)
</div>
```
from music21 import *
```
Probably you have to set manually the correct file path to an Application that is able to open MusicXML files (like MuseScore). To do so, use the `music21.environment` module to set an `musicxmlPath` key.
Make sure to change the string `path/to/your/musicXmlApplication` below to the correct file path (keep the quotation marks):
- on Mac e.g.: `/Applications/MuseScore 2.app/Contents/MacOS/mscore`
- or on Windows e.g.: `C:/Program Files (x86)/MuseScore 2/bin/MuseScore.exe`
and uncomment the line (remove the `#` at the begin of the line).
In the same way, you can also add a path to your lilypond installation, using
`env['lilypondPath']`:
- on Mac e.g.: `Applications/Lilypond.app`
- on Windows e.g.: `C:/Program Files (x86)/LilyPond/usr/bin/lilypond.exe`
Sometimes it's also necessary to adapt the `musescoreDirectPNGPath`. Check if it corresponds to your museScore path.
```
env = environment.Environment()
# env['musicxmlPath'] = 'path/to/your/musicXmlApplication'
# env['lilypondPath'] = 'path/to/your/lilypond'
# env['musescoreDirectPNGPath'] = 'path/to/your/museScore'
print('Environment settings:')
print('musicXML: ', env['musicxmlPath'])
print('musescore: ', env['musescoreDirectPNGPath'])
print('lilypond: ', env['lilypondPath'])
```
### Let's create some notes
One possible way to create notes in music21 is to use the `Note()`-Object (CAPITAL LETTER) within music21's `note`-subModule (small letter).
Let's use the twelve-tone row of Alban Berg's Violin Concerto (1935) as an example. Take care how the different octaves and accidentals are declared.
```
note1 = note.Note("G3") # declaration of first note
note2 = note.Note("B-3")
note3 = note.Note("D4")
note4 = note.Note("F#4")
note5 = note.Note("A4")
note6 = note.Note("C5")
note7 = note.Note("E5")
note8 = note.Note("G#5")
note9 = note.Note("B5")
note10 = note.Note("C#6")
note11 = note.Note("D#6")
note12 = note.Note("F6")
# combine the twelve notes in a row list
bergRow = [note1, note2, note3, note4, note5, note6, note7, note8, note9, note10, note11, note12]
bergRow # output of bergRow (by just using the name of the variable)
```
You can use `dir(MODULENAME)` to find out which objects any module contains at all (http://web.mit.edu/music21/doc/usersGuide/usersGuide_02_notes.html#usersguide-02-notes):
```
dir(note)
```
To iterate over every single item in a list, you can use a "FOR"-loop.
Syntax (indentation matters here!):
for ITEM in LIST:
do something with ITEM
...
```
for currentNote in bergRow: # for every note in bergRow list do...
currentNote.duration.type = 'whole' # ... declare duration of a whole note
print(currentNote.duration, currentNote.nameWithOctave) # ... output of note duration and name (using the print command)
```
### Create simple Streams
Streams are fundamental objects in music21. Almost everything (`Score`, `Parts`, `Voices`, `Measures` a.o.) is organized in terms of this abstract data structure. An empty stream is created by using the `Stream()`-Object (CAPITAL LETTER) within music21's `stream`-subModule (small letter).
```
bergStream = stream.Stream() # create empty stream
for currentNote in bergRow: # iterate over every note in bergRow and ...
bergStream.append(currentNote) # ... append current note to the stream
bergStream.show('text') # output of the stream (using the .show()-method with option 'text'; compare to output above)
```
You can get the length of a stream, what is the number of items in it, with `len(STREAM)`:
```
len(bergStream)
```
... or with just counting the Note-Elements (here you have to flatten the stream):
```
len(bergStream.flat.getElementsByClass(note.Note))
```
But let's have a look at the stream now. Calling the `.show()`-method without any option will display a graphical visualisation of any music object via the musicxmlApplication defined in the environment at the beginning of this notebook.
If you encounter problems here, make sure you have set the correct environment settings for `musicxmlPath` and `musescoreDirectPNGPath`.
```
bergStream.show()
```
You can also use further options to get the output as `pdf` or `png` via `lilypond`:
```
bergStream.show('lily.pdf')
bergStream.show('lily.png')
```
You could also use music21.tinyNotation, "a simple way of specifying single line melodies" (http://web.mit.edu/music21/doc/moduleReference/moduleTinyNotation.html), to define the notes of the row:
```
bergRowTiny = converter.parse("tinyNotation: G1 B- d f# a c' e' g'# b' c''# d''# f''")
bergRowTiny.show()
```
Our `bergRowTiny` is also a stream because the tinyNotation converter created it automatically. But keep aware of the slightly different structure:
```
bergRowTiny.show('text')
```
### Ok nice, but where is the analytical part?
music21 provides a large amount of build-in analytical tools. To start right away, just let's get the ambitus of the row in the stream using the `.analyze()`-method (http://web.mit.edu/music21/doc/moduleReference/moduleStream.html):
```
bergStream.analyze('ambitus')
```
But always keep a "thinking" eye on the results:
```
bergStream.analyze('key')
```
The twelve-tone row of Berg's Violin Concerto is special because of its two major triads, two minor triads and a part of the whole tone scale. Let's separate these elements into new `Chord()`-Objects (part of `chord`-submodule):
```
# declare some variables as Chord()-Objects
triad1 = chord.Chord()
triad2 = chord.Chord()
triad3 = chord.Chord()
triad4 = chord.Chord()
wtScale = chord.Chord()
# iterate over the first three notes in the stream
for currentNote in bergStream[0:3]:
triad1.add(currentNote) # add the currentNote to the Chord()
# ...
for currentNote in bergStream[2:5]:
triad2.add(currentNote)
# ...
for currentNote in bergStream[4:7]:
triad3.add(currentNote)
# ...
for currentNote in bergStream[6:9]:
triad4.add(currentNote)
# iterate over the last three notes in the stream
for currentNote in bergStream[8:12]:
wtScale.add(currentNote)
# output the 5 chords
triad1.show()
triad2.show()
triad3.show()
triad4.show()
wtScale.show()
```
You can recombine multiple Chords() within a new Chord()-Object:
```
fullChord = chord.Chord([triad1, triad2, triad3, triad4, wtScale])
fullChord.show()
```
You can also append the chords to a new Stream()-Object:
```
# create empty stream
chordsStream = stream.Stream()
# append all the triads to the stream
chordsStream.append(triad1);
chordsStream.append(triad2);
chordsStream.append(triad3);
chordsStream.append(triad4);
chordsStream.append(wtScale);
chordsStream.show()
```
And you can add some analytical descriptions to the objects using the `.addLyric()`-method and different attributes (e.g. `pitchedCommonName`, `intervalVector`, `primeForm`, `forteClass`) of the chords:
```
# iterate over every chord in the stream, and ...
for currentChord in chordsStream:
currentChord.addLyric(currentChord.pitchedCommonName) # ... add triad name
currentChord.addLyric(currentChord.intervalVector) # ... add interval vector
currentChord.addLyric(currentChord.primeForm) # ... add prime form
currentChord.addLyric(currentChord.forteClass) # ... add forte class
chordsStream.show()
```
Highlighting certain parts (e.g. all Forte classes "3-11A" = minor chord or "3-11B" = major chord) is also possible (http://web.mit.edu/music21/doc/usersGuide/usersGuide_10_examples1.html):
```
for currentChord in chordsStream.recurse().getElementsByClass('Chord'):
if currentChord.forteClass == '3-11A':
currentChord.style.color = 'red'
for x in currentChord.derivation.chain():
x.style.color = 'blue'
if currentChord.forteClass == '3-11B':
currentChord.style.color = 'blue'
for x in currentChord.derivation.chain():
x.style.color = 'blue'
chordsStream.show()
```
### Introducing music21 the serial module
Most (=all?) of the twelve tone rows by Schönberg, Berg and Webern are already incorporated into a dictionary list in music21. You get an sorted overview of the rows available in the dictionary with the following command: (http://web.mit.edu/music21/doc/moduleReference/moduleSerial.html)
```
sorted(list(serial.historicalDict))
```
For all these rows, music21 provides not only the pitches of the row, but some additional meta information. So let's see what we get with the 'RowBergViolinConcerto':
```
bergRowInternal = serial.getHistoricalRowByName('RowBergViolinConcerto')
print(type(bergRowInternal))
print(bergRowInternal.composer)
print(bergRowInternal.opus)
print(bergRowInternal.title)
print(bergRowInternal.row)
print(bergRowInternal.pitchClasses())
bergRowInternal.noteNames()
```
### Transformations
Using the serial modules' '.originalCenteredTransformation()'-method, you can retrieve transformational forms of a ToneRow()-Object. "Admissible transformations are ‘T’ (transposition), ‘I’ (inversion), ‘R’ (retrograde), and ‘RI’ (retrograde inversion)." (http://web.mit.edu/music21/doc/moduleReference/moduleSerial.html)
```
g = bergRowInternal.originalCenteredTransformation('T', 0)
u = bergRowInternal.originalCenteredTransformation('I', 0)
k = bergRowInternal.originalCenteredTransformation('R', 0)
ku = bergRowInternal.originalCenteredTransformation('RI', 0)
print('original:')
g.show()
print('inversion:')
u.show()
print('retrograde:')
k.show()
print('retrograde inversion:')
ku.show()
```
### 12-tone matrix
You can also easily get the 12-tone matrix of a twelve tone row:
```
bergMatrix1 = bergRowInternal.matrix()
print(bergMatrix1)
bergMatrix2 = serial.rowToMatrix(bergRowInternal.row)
print(bergMatrix2)
```
### Segmentation
One of the fundamental operations concerning the analysis of a twelve tone composition is segmentation. The following example provides a function, that iterates over a set of notes (`bergRowInternal`) and looks for every possible segment of a certain length (`segmentationSize`). Thus, per default, we iterate over every possible 3-tone segment of the Berg row.
```
segmentationList = {}
segmentationLength = 3 # here you can choose the length of the segments (try other values)
rangeEnd = 12 - segmentationLength + 1
# iterate over the whole tone row in (rangeEnd - 1) steps
for i in range(0,rangeEnd):
print('---')
# create an empty placeholder for the segment as a ToneRow()-Object
# at the position i in the segmentationList
segmentationList[i] = serial.ToneRow()
# fill up the segment with the corresponding notes
for currentNote in bergRowInternal[i:i+segmentationLength]:
segmentationList[i].append(currentNote)
print('Run ', i, ' completed.') # This is for control only.
segmentationList # output of the whole list
```
Now that we have every possible 3-tone segment of the Berg row, we can check if there are any triads in it:
```
# check for triads in the segmentation list
# make sure to use segmentLength = 3 above
# (for segmentLength = 4 you will get 7th and other tetra chords)
for i in segmentationList:
print('---')
print('RUN ', i)
outputString = ''
# get a list of the pitches of the current segment
currentPitchList = segmentationList[i].pitches
print(currentPitchList)
#use the pitchList as input for a chord
currentChord = chord.Chord(currentPitchList)
# check for minor triad (with highlighting)
# use forteClass 3-11A instead of 'isMinorTriad()'-method to catch enharmonic equivalents
if currentChord.forteClass == '3-11A':
currentChord.style.color = 'red'
outputString = 'MINOR TRIAD: '
# check for major triad (with highlighting)
# use forteClass 3-11B instead of 'isMajorTriad()'-method to catch enharmonic equivalents
if currentChord.forteClass == '3-11B':
currentChord.style.color = 'blue'
outputString = 'MAJOR TRIAD: '
currentChord.show()
outputString += currentChord.pitchedCommonName
print(outputString)
```
| github_jupyter |
# Creation of the annotation dataset
This notebook describes the steps involved in gathering, cleaning, and merging all manual annotations made by various analysist from various groups and using different software in a single large dataset.
## Data cleaning
Annotations are for each datasets are loaded, sorted, and re-writen into a parquet file. First, ecosound and the other linbraries need to be imported.
```
import sys
sys.path.append("..") # Adds higher directory to python modules path.
import os
from ecosound.core.annotation import Annotation
from ecosound.core.metadata import DeploymentInfo
```
### Dataset 1: DFO - Snake Island RCA-In
Definition of all the paths of all folders with the raw annotation and audio files for this deployment.
```
root_dir = r'C:\Users\xavier.mouy\Documents\PhD\Projects\Dectector\datasets\DFO_snake-island_rca-in_20181017'
deployment_file = r'deployment_info.csv'
annotation_dir = r'manual_annotations'
data_dir = r'audio_data'
```
Instantiate a DeploymentInfo object to handle metadata for the deployment, and create an empty deployment info file.
```
# Instantiate
Deployment = DeploymentInfo()
# write empty file to fill in (do once only)
Deployment.write_template(os.path.join(root_dir, deployment_file))
```
A csv file "deployment_info.csv" has now been created in the root_dir. It is empty and only has column headers, and includes teh following fiilds:
* audio_channel_number
* UTC_offset
* sampling_frequency (in Hz)
* bit_depth
* mooring_platform_name
* recorder_type
* recorder_SN
* hydrophone_model
* hydrophone_SN
* hydrophone_depth
* location_name
* location_lat
* location_lon
* location_water_depth
* deployment_ID
* deployment_date
* recovery_date
This file needs to be filled in by the user with teh appropriate deployment information. Once fileld in, the file can be loaded using the Deployment object:
```
# load deployment file
deployment_info = Deployment.read(os.path.join(root_dir, deployment_file))
deployment_info
```
Now we can load the manual annotations for this dataset. Here annotatiosn were performed with Raven:
```
# load all annotations
annot = Annotation()
annot.from_raven(os.path.join(root_dir, annotation_dir),
class_header='Class',
subclass_header='Sound type',
verbose=True)
```
Now we can fill in all the missing information in teh annotations field with the deployment information:
```
# Manually fill in missing information
annot.insert_values(software_version='1.5',
operator_name='Stephanie Archer',
audio_channel=deployment_info.audio_channel_number[0],
UTC_offset=deployment_info.UTC_offset[0],
audio_file_dir=os.path.join(root_dir, data_dir),
audio_sampling_frequency=deployment_info.sampling_frequency[0],
audio_bit_depth=deployment_info.bit_depth[0],
mooring_platform_name=deployment_info.mooring_platform_name[0],
recorder_type=deployment_info.recorder_type[0],
recorder_SN=deployment_info.recorder_SN[0],
hydrophone_model=deployment_info.hydrophone_model[0],
hydrophone_SN=deployment_info.hydrophone_SN[0],
hydrophone_depth=deployment_info.hydrophone_depth[0],
location_name = deployment_info.location_name[0],
location_lat = deployment_info.location_lat[0],
location_lon = deployment_info.location_lon[0],
location_water_depth = deployment_info.location_water_depth[0],
deployment_ID=deployment_info.deployment_ID[0],
)
```
Let's look at the different annotation labels that were used:
```
print(annot.get_labels_class())
```
It is clear that there are some inconsistencies in the label names (e.g. 'unknown', 'unkown'). Let's rename the class labels so everything has a consistent same name. We'll use teh following convention:
* 'FS' for fish
* 'UN' for unknown sound
* 'KW' for killer whale
* 'ANT' for anthropogenic sound
* 'HS' for harbor seal
```
annot.data['label_class'].replace(to_replace=['fish'], value='FS', inplace=True)
annot.data['label_class'].replace(to_replace=['fish?','unkown_invert','unknown_invert','fish?','unknown','unkown','whale?','?','sea lion?','mammal'], value='UN', inplace=True)
annot.data['label_class'].replace(to_replace=['airplane'], value='ANT', inplace=True)
annot.data['label_class'].dropna(axis=0, inplace=True)
```
Let's check that the class label are now all consistent.
```
print(annot.get_labels_class())
```
Now, having a look a summary of all the annotations available in this dataset.
```
# print summary (pivot table)
print(annot.summary())
```
Now that all the metadata (deployment information) are filled in the annotation fields and that all labels have been "cleaned up", we can save the dataset as a parquet file.
```
#annot.to_parquet(os.path.join(root_dir, 'Annotations_dataset_' + deployment_info.deployment_ID[0] + '.parquet'))
annot.to_netcdf(os.path.join(root_dir, 'Annotations_dataset_' + deployment_info.deployment_ID[0] + '.nc'))
```
The dataset can also be save as a Raven or PAMlab annotation file.
```
annot.to_pamlab(root_dir, outfile='Annotations_dataset_' + deployment_info.deployment_ID[0] +' annotations.log', single_file=True)
annot.to_raven(root_dir, outfile='Annotations_dataset_' + deployment_info.deployment_ID[0] +'.Table.1.selections.txt', single_file=True)
```
### Dataset 2: DFO - Snake Island RCA-Out
Now we can repeat the step above for all the other datasets:
```
root_dir = r'C:\Users\xavier.mouy\Documents\PhD\Projects\Dectector\datasets\DFO_snake-island_rca-out_20181015'
deployment_file = r'deployment_info.csv'
annotation_dir = r'manual_annotations'
data_dir = r'audio_data'
# Instantiate
Deployment = DeploymentInfo()
# write empty file to fill in (do once only)
#Deployment.write_template(os.path.join(root_dir, deployment_file))
# load deployment file
deployment_info = Deployment.read(os.path.join(root_dir, deployment_file))
# load all annotations
annot = Annotation()
annot.from_raven(os.path.join(root_dir, annotation_dir),
class_header='Class',
subclass_header='Sound type',
verbose=True)
# Manually fill in missing information
annot.insert_values(software_version='1.5',
operator_name='Stephanie Archer',
audio_channel=deployment_info.audio_channel_number[0],
UTC_offset=deployment_info.UTC_offset[0],
audio_file_dir=os.path.join(root_dir, data_dir),
audio_sampling_frequency=deployment_info.sampling_frequency[0],
audio_bit_depth=deployment_info.bit_depth[0],
mooring_platform_name=deployment_info.mooring_platform_name[0],
recorder_type=deployment_info.recorder_type[0],
recorder_SN=deployment_info.recorder_SN[0],
hydrophone_model=deployment_info.hydrophone_model[0],
hydrophone_SN=deployment_info.hydrophone_SN[0],
hydrophone_depth=deployment_info.hydrophone_depth[0],
location_name = deployment_info.location_name[0],
location_lat = deployment_info.location_lat[0],
location_lon = deployment_info.location_lon[0],
location_water_depth = deployment_info.location_water_depth[0],
deployment_ID=deployment_info.deployment_ID[0],
)
```
Some inconsistent class labels here as well:
```
print(annot.get_labels_class())
```
Fixing labels according to our naming convention:
```
annot.data['label_class'].replace(to_replace=['fish','fish?','fush'], value='FS', inplace=True)
annot.data['label_class'].replace(to_replace=['?'], value='UN', inplace=True)
annot.data['label_class'].dropna(axis=0, inplace=True)
print(annot.get_labels_class())
```
Summary:
```
# print summary (pivot table)
print(annot.summary())
```
Saving the cleaned up dataset:
```
# save as parquet file
annot.to_netcdf(os.path.join(root_dir, 'Annotations_dataset_' + deployment_info.deployment_ID[0] + '.nc'))
#annot.to_parquet(os.path.join(root_dir, 'Annotations_dataset_' + deployment_info.deployment_ID[0] + '.parquet'))
annot.to_pamlab(root_dir, outfile='Annotations_dataset_' + deployment_info.deployment_ID[0] +' annotations.log', single_file=True)
annot.to_raven(root_dir, outfile='Annotations_dataset_' + deployment_info.deployment_ID[0] +'.Table.1.selections.txt', single_file=True)
```
### Dataset 3: ONC - Delta Node 2014
Repeating the same steps as teh prvious dataset. The difference here is that the annotations were performed with PAMlab instead of Raven.
```
root_dir = r'C:\Users\xavier.mouy\Documents\PhD\Projects\Dectector\datasets\ONC_delta-node_2014'
deployment_file = r'deployment_info.csv'
annotation_dir = r'manual_annotations'
data_dir = r'audio_data'
# Instantiate
Deployment = DeploymentInfo()
# write empty file to fill in (do once only)
#Deployment.write_template(os.path.join(root_dir, deployment_file))
# # load deployment file
deployment_info = Deployment.read(os.path.join(root_dir, deployment_file))
# # load all annotations
annot = Annotation()
annot.from_pamlab(os.path.join(root_dir, annotation_dir), verbose=True)
# Mnaually fill in missing information
annot.insert_values(software_version='6.2.2',
operator_name='Xavier Mouy',
audio_channel=deployment_info.audio_channel_number[0],
UTC_offset=deployment_info.UTC_offset[0],
audio_file_dir=os.path.join(root_dir, data_dir),
audio_sampling_frequency=deployment_info.sampling_frequency[0],
audio_bit_depth=deployment_info.bit_depth[0],
mooring_platform_name=deployment_info.mooring_platform_name[0],
recorder_type=deployment_info.recorder_type[0],
recorder_SN=deployment_info.recorder_SN[0],
hydrophone_model=deployment_info.hydrophone_model[0],
hydrophone_SN=deployment_info.hydrophone_SN[0],
hydrophone_depth=deployment_info.hydrophone_depth[0],
location_name=deployment_info.location_name[0],
location_lat=deployment_info.location_lat[0],
location_lon=deployment_info.location_lon[0],
location_water_depth=deployment_info.location_water_depth[0],
deployment_ID=deployment_info.deployment_ID[0],
)
```
No inconsistent class labels this time:
```
print(annot.get_labels_class())
```
Summary:
```
# print summary (pivot table)
print(annot.summary())
```
Saving the cleaned up dataset:
```
# save as parquet file
annot.to_netcdf(os.path.join(root_dir, 'Annotations_dataset_' + deployment_info.deployment_ID[0] + '.nc'))
#annot.to_parquet(os.path.join(root_dir, 'Annotations_dataset_' + deployment_info.deployment_ID[0] + '.parquet'))
annot.to_pamlab(root_dir, outfile='Annotations_dataset_' + deployment_info.deployment_ID[0] +' annotations.log', single_file=True)
annot.to_raven(root_dir, outfile='Annotations_dataset_' + deployment_info.deployment_ID[0] +'.Table.1.selections.txt', single_file=True)
```
### Dataset 4: UVIC - Hornby Island
We can repeat the step above for all the other datasets:
```
root_dir = r'C:\Users\xavier.mouy\Documents\PhD\Projects\Dectector\datasets\UVIC_hornby-island_2019'
deployment_file = r'deployment_info.csv'
annotation_dir = r'manual_annotations'
data_dir = r'audio_data'
# Instantiate
Deployment = DeploymentInfo()
# write empty file to fill in (do once only)
#Deployment.write_template(os.path.join(root_dir, deployment_file))
# load deployment file
deployment_info = Deployment.read(os.path.join(root_dir, deployment_file))
# load all annotations
annot = Annotation()
annot.from_raven(os.path.join(root_dir, annotation_dir), verbose=True)
# Mnaually fill in missing information
annot.insert_values(software_version='1.5',
operator_name='Emie Woodburn',
audio_channel=deployment_info.audio_channel_number[0],
UTC_offset=deployment_info.UTC_offset[0],
audio_file_dir=os.path.join(root_dir, data_dir),
audio_sampling_frequency=deployment_info.sampling_frequency[0],
audio_bit_depth=deployment_info.bit_depth[0],
mooring_platform_name=deployment_info.mooring_platform_name[0],
recorder_type=deployment_info.recorder_type[0],
recorder_SN=deployment_info.recorder_SN[0],
hydrophone_model=deployment_info.hydrophone_model[0],
hydrophone_SN=deployment_info.hydrophone_SN[0],
hydrophone_depth=deployment_info.hydrophone_depth[0],
location_name = deployment_info.location_name[0],
location_lat = deployment_info.location_lat[0],
location_lon = deployment_info.location_lon[0],
location_water_depth = deployment_info.location_water_depth[0],
deployment_ID=deployment_info.deployment_ID[0],
)
```
Some inconsistent class labels :
```
print(annot.get_labels_class())
```
Fixing labels according to our naming convention:
```
annot.data['label_class'].replace(to_replace=['FSFS',' FS'], value='FS', inplace=True)
annot.data['label_class'].replace(to_replace=['KW '], value='KW', inplace=True)
annot.data['label_class'].replace(to_replace=['Seal','Seal\\'], value='HS', inplace=True)
annot.data['label_class'].replace(to_replace=['Unknown','Chirp',' ',' '], value='UN', inplace=True)
annot.data['label_class'].dropna(axis=0, inplace=True)
print(annot.get_labels_class())
```
Summary:
```
# print summary (pivot table)
print(annot.summary())
```
Saving the cleaned up dataset:
```
# save as parquet file
annot.to_netcdf(os.path.join(root_dir, 'Annotations_dataset_' + deployment_info.deployment_ID[0] + '.nc'))
#annot.to_parquet(os.path.join(root_dir, 'Annotations_dataset_' + deployment_info.deployment_ID[0] + '.parquet'))
annot.to_pamlab(root_dir, outfile='Annotations_dataset_' + deployment_info.deployment_ID[0] +' annotations.log', single_file=True)
annot.to_raven(root_dir, outfile='Annotations_dataset_' + deployment_info.deployment_ID[0] +'.Table.1.selections.txt', single_file=True)
```
### Dataset 5: UVIC - Mill Bay
We can repeat the step above for all the other datasets:
```
root_dir = r'C:\Users\xavier.mouy\Documents\PhD\Projects\Dectector\datasets\UVIC_mill-bay_2019'
deployment_file = r'deployment_info.csv'
annotation_dir = r'manual_annotations'
data_dir = r'audio_data'
# Instantiate
Deployment = DeploymentInfo()
# write empty file to fill in (do once only)
#Deployment.write_template(os.path.join(root_dir, deployment_file))
# load deployment file
deployment_info = Deployment.read(os.path.join(root_dir, deployment_file))
# load all annotations
annot = Annotation()
annot.from_raven(os.path.join(root_dir, annotation_dir),
class_header='Sound Type',
verbose=True)
# Mnaually fill in missing information
annot.insert_values(software_version='1.5',
operator_name='Courtney Evin',
audio_channel=deployment_info.audio_channel_number[0],
UTC_offset=deployment_info.UTC_offset[0],
audio_file_dir=os.path.join(root_dir, data_dir),
audio_sampling_frequency=deployment_info.sampling_frequency[0],
audio_bit_depth=deployment_info.bit_depth[0],
mooring_platform_name=deployment_info.mooring_platform_name[0],
recorder_type=deployment_info.recorder_type[0],
recorder_SN=deployment_info.recorder_SN[0],
hydrophone_model=deployment_info.hydrophone_model[0],
hydrophone_SN=deployment_info.hydrophone_SN[0],
hydrophone_depth=deployment_info.hydrophone_depth[0],
location_name = deployment_info.location_name[0],
location_lat = deployment_info.location_lat[0],
location_lon = deployment_info.location_lon[0],
location_water_depth = deployment_info.location_water_depth[0],
deployment_ID=deployment_info.deployment_ID[0],
)
```
Some inconsistent class labels :
```
print(annot.get_labels_class())
```
Fixing labels according to our naming convention:
```
annot.data['label_class'].replace(to_replace=['fs','F','SF'], value='FS', inplace=True)
annot.data['label_class'].replace(to_replace=['unknown-mammal?','unknown','unknown-invert'], value='UN', inplace=True)
annot.data['label_class'].dropna(axis=0, inplace=True)
print(annot.get_labels_class())
```
Summary:
```
# print summary (pivot table)
print(annot.summary())
```
Saving the cleaned up dataset:
```
# save as parquet file
annot.to_netcdf(os.path.join(root_dir, 'Annotations_dataset_' + deployment_info.deployment_ID[0] + '.nc'))
#annot.to_parquet(os.path.join(root_dir, 'Annotations_dataset_' + deployment_info.deployment_ID[0] + '.parquet'))
annot.to_pamlab(root_dir, outfile='Annotations_dataset_' + deployment_info.deployment_ID[0] +' annotations.log', single_file=True)
annot.to_raven(root_dir, outfile='Annotations_dataset_' + deployment_info.deployment_ID[0] +'.Table.1.selections.txt', single_file=True)
```
# Merging all datasets together
Now that all our datasets are cleaned up, we can merge them all in a single Master annotation dataset.
Defining the path of each dataset:
```
root_dir = r'C:\Users\xavier.mouy\Documents\PhD\Projects\Dectector\datasets'
dataset_files = ['UVIC_mill-bay_2019\Annotations_dataset_06-MILL.nc',
'UVIC_hornby-island_2019\Annotations_dataset_07-HI.nc',
'ONC_delta-node_2014\Annotations_dataset_ONC-Delta-2014.nc',
'DFO_snake-island_rca-in_20181017\Annotations_dataset_SI-RCAIn-20181017.nc',
'DFO_snake-island_rca-out_20181015\Annotations_dataset_SI-RCAOut-20181015.nc',
]
```
Looping through each dataset and merging in to a master dataset:
```
# # load all annotations
annot = Annotation()
for file in dataset_files:
tmp = Annotation()
tmp.from_netcdf(os.path.join(root_dir, file), verbose=True)
annot = annot + tmp
```
Now we can see a summary of all the annotatiosn we have:
```
# print summary (pivot table)
print(annot.summary())
```
We can also look at the contribution from each analyst:
```
print(annot.summary(rows='operator_name'))
```
Finally we can save our Master annotation dataset. It will be used for trainning and evealuation classification models.
```
#annot.to_parquet(os.path.join(root_dir, 'Master_annotations_dataset.parquet'))
annot.to_netcdf(os.path.join(root_dir, 'Master_annotations_dataset.nc'))
```
| github_jupyter |
```
import os
os.chdir('G:\\D\\Edureka\\Edureka - 24 June - Python\\Class 19 - 20\\')
import pandas as pd
import numpy as np
dataset = pd.read_csv('airline_passengers.csv')
dataset.plot()
dataset['Month'] = pd.to_datetime(dataset['Month'])
dataset
dataset.set_index('Month', inplace=True)
dataset.plot()
from statsmodels.tsa.seasonal import seasonal_decompose
decompostion = seasonal_decompose(dataset['Thousands of Passengers'], freq=12)
decompostion.plot()
from statsmodels.tsa.stattools import adfuller
adfuller(dataset['Thousands of Passengers'])
def adf_check(time_series):
result = adfuller(time_series)
print('Augmented Dickey Fuller Test')
labels =['ADF Test Statistic', 'P-value', '#lags', 'No of Obs']
for value, label in zip(result,labels):
print(label + ':' + str(value))
if result[1]<=0.05:
print('Strong evidence agaisnt null hypothesis and my time series is sationary')
else:
print('Weak evidence agaisnt null hypothesis and my time series is not stationary')
adf_check(dataset['Thousands of Passengers'])
dataset['Thousands of Passengers First Diff'] = dataset['Thousands of Passengers']-dataset['Thousands of Passengers'].shift(1)
adf_check(dataset['Thousands of Passengers First Diff'].dropna())
dataset['Thousands of Passengers second Diff'] = dataset['Thousands of Passengers First Diff']-dataset['Thousands of Passengers First Diff'].shift(1)
dataset
adf_check(dataset['Thousands of Passengers second Diff'].dropna())
#d=2
dataset['Seasonal Difference'] = dataset['Thousands of Passengers']-dataset['Thousands of Passengers'].shift(12)
adf_check(dataset['Seasonal Difference'].dropna())
#D=1
#AR(p=2)
#PACF (Partial Autocorrelation Function)
#Yt+1 = B1Yt + B2Yt-1 + e
#MA(q=2)
#ACF Autocorrealation Function
# Yt+1 = a1e1 + a2et-1 +... Mu + e
from statsmodels.graphics.tsaplots import plot_acf, plot_pacf
plot_pacf(dataset['Thousands of Passengers second Diff'].dropna(),lags=14)
#p=0
plot_acf(dataset['Thousands of Passengers second Diff'].dropna(), lags=14)
#q=0
plot_pacf(dataset['Seasonal Difference'].dropna(), lags=12)
#P=1
plot_acf(dataset['Seasonal Difference'].dropna(),lags=12)
#Q=2
#d = 2 , D = 1, p = 0 , P = 1, q = 0, Q = 2
from statsmodels.tsa.arima_model import ARIMA
import statsmodels.api as sm
model = sm.tsa.statespace.SARIMAX(dataset['Thousands of Passengers'], order=(0,2,0), seasonal_order=(1,2, 1, 12))
results = model.fit()
print(results.summary())
dataset['Forecast'] = results.predict(start = 130, end=144, dynamic=True)
dataset[['Thousands of Passengers', 'Forecast']].plot()
dataset
from pandas.tseries.offsets import DateOffset
future_dates = [dataset.index[-1]+ DateOffset(months=x) for x in range(0,24)]
future_dates
future_dates_df = pd.DataFrame(index=future_dates[1:], columns=dataset.columns)
future_dates_df
future_df = pd.concat([dataset,future_dates_df])
future_df
future_df['Forecast'] = results.predict(start=145, end=165, dynamic=True)
future_df[['Thousands of Passengers', 'Forecast']].plot()
future_df
Next Class 5PM to 8PM IST?
```
| github_jupyter |
Run the following command to install the Kubeflow Pipelines SDK. If you run this command in a Jupyter
notebook, restart the kernel after installing the SDK.
```
%pip install kfp --upgrade
# to install tekton compiler uncomment the line below
# %pip install kfp_tekton
```
Import Packages
```
import json
import time
import yaml
import kfp
import kfp.components as comp
import kfp.dsl as dsl
SPARK_COMPLETED_STATE = "COMPLETED"
SPARK_APPLICATION_KIND = "sparkapplications"
def get_spark_job_definition():
"""
Read Spark Operator job manifest file and return the corresponding dictionary and
add some randomness in the job name
:return: dictionary defining the spark job
"""
# Read manifest file
with open("spark-job.yaml", "r") as stream:
spark_job_manifest = yaml.safe_load(stream)
# Add epoch time in the job name
epoch = int(time.time())
spark_job_manifest["metadata"]["name"] = spark_job_manifest["metadata"]["name"].format(epoch=epoch)
return spark_job_manifest
def print_op(msg):
"""
Op to print a message.
"""
return dsl.ContainerOp(
name="Print message.",
image="alpine:3.6",
command=["echo", msg],
)
@dsl.graph_component # Graph component decorator is used to annotate recursive functions
def graph_component_spark_app_status(input_application_name):
k8s_get_op = comp.load_component_from_file("k8s-get-component.yaml")
check_spark_application_status_op = k8s_get_op(
name=input_application_name,
kind=SPARK_APPLICATION_KIND
)
# Remove cache
check_spark_application_status_op.execution_options.caching_strategy.max_cache_staleness = "P0D"
time.sleep(5)
with dsl.Condition(check_spark_application_status_op.outputs["applicationstate"] != SPARK_COMPLETED_STATE):
graph_component_spark_app_status(check_spark_application_status_op.outputs["name"])
@dsl.pipeline(
name="Spark Operator job pipeline",
description="Spark Operator job pipeline"
)
def spark_job_pipeline():
# Load spark job manifest
spark_job_definition = get_spark_job_definition()
# Load the kubernetes apply component
k8s_apply_op = comp.load_component_from_file("k8s-apply-component.yaml")
# Execute the apply command
spark_job_op = k8s_apply_op(object=json.dumps(spark_job_definition))
# Fetch spark job name
spark_job_name = spark_job_op.outputs["name"]
# Remove cache for the apply operator
spark_job_op.execution_options.caching_strategy.max_cache_staleness = "P0D"
spark_application_status_op = graph_component_spark_app_status(spark_job_op.outputs["name"])
spark_application_status_op.after(spark_job_op)
print_message = print_op(f"Job {spark_job_name} is completed.")
print_message.after(spark_application_status_op)
print_message.execution_options.caching_strategy.max_cache_staleness = "P0D"
```
### Compile and run your pipeline
After defining the pipeline in Python as described in the preceding section, use one of the following options to compile the pipeline and submit it to the Kubeflow Pipelines service.
#### Option 1: Compile and then upload in UI
1. Run the following to compile your pipeline and save it as `spark_job_pipeline.yaml`.
For Argo (Default)
```
# create piepline file for argo backend the default one if you use tekton use the block below
if __name__ == "__main__":
# Compile the pipeline
import kfp.compiler as compiler
import logging
logging.basicConfig(level=logging.INFO)
pipeline_func = spark_job_pipeline
pipeline_filename = pipeline_func.__name__ + ".yaml"
compiler.Compiler().compile(pipeline_func, pipeline_filename)
logging.info(f"Generated pipeline file: {pipeline_filename}.")
```
For Tekton
```
# uncomment the block below to create pipeline file for tekton
# if __name__ == '__main__':
# from kfp_tekton.compiler import TektonCompiler
# import logging
# logging.basicConfig(level=logging.INFO)
# pipeline_func = spark_job_pipeline
# pipeline_filename = pipeline_func.__name__ + ".yaml"
# TektonCompiler().compile(pipeline_func, pipeline_filename)
# logging.info(f"Generated pipeline file: {pipeline_filename}.")
```
2. Upload and run your `spark_job_pipeline.yaml` using the Kubeflow Pipelines user interface.
See the guide to [getting started with the UI][quickstart].
[quickstart]: https://www.kubeflow.org/docs/components/pipelines/overview/quickstart
#### Option 2: run the pipeline using Kubeflow Pipelines SDK client
1. Create an instance of the [`kfp.Client` class][kfp-client] following steps in [connecting to Kubeflow Pipelines using the SDK client][connect-api].
[kfp-client]: https://kubeflow-pipelines.readthedocs.io/en/latest/source/kfp.client.html#kfp.Client
[connect-api]: https://www.kubeflow.org/docs/components/pipelines/sdk/connect-api
```
client = kfp.Client() # change arguments accordingly
client.create_run_from_pipeline_func(
spark_job_pipeline)
```
| github_jupyter |
# Data preparation
```
import sqlite3
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import Cdf
import Pmf
# define global plot parameters
params = {'axes.labelsize' : 12, 'axes.titlesize' : 12,
'font.size' : 12, 'legend.fontsize' : 12,
'xtick.labelsize' : 12, 'ytick.labelsize' : 12}
plt.rcParams.update(params)
plt.rcParams.update({'figure.max_open_warning': 0})
# helper function to get CDFs for an array of dataframes for a specific column name
def get_cdfs(dfs, col):
cdfs = []
for df in dfs:
cdfs.append(Cdf.MakeCdfFromList(df[col]))
return cdfs
```
### Cache Identification based on heuristic (see paper)
Matching source and destination AS numbers indicate that request stayed within ISP network
```
# connect to database and get data
conn = sqlite3.connect('data/youtube-traceroute.db')
# cache for IPv4 only
v4_caches = pd.read_sql_query('select * \
from pair_medians_meta \
where src_asn_v4 = dst_asn_v4 and src_asn_v6 != dst_asn_v6', conn)
# cache for IPv6 only
v6_caches = pd.read_sql_query('select * \
from pair_medians_meta \
where src_asn_v4 != dst_asn_v4 and src_asn_v6 = dst_asn_v6', conn)
# caches for both
both_caches = pd.read_sql_query('select * \
from pair_medians_meta \
where src_asn_v4 = dst_asn_v4 and src_asn_v6 = dst_asn_v6', conn)
# no cache
no_caches = pd.read_sql_query('select * \
from pair_medians_meta \
where src_asn_v4 != dst_asn_v4 and src_asn_v6 != dst_asn_v6', conn)
conn.close()
```
## Putting lists of the four cases together for plotting
- IPv4 cache, no IPv6 cache
- IPv6 cache, no IPv4 cache
- caches for both
- no caches for either
```
# all lists will represent the four cases above and therefore have length 4
data = [v4_caches,
v6_caches,
both_caches,
no_caches]
titles = ['(v4 but no v6 cache)',
'(v6 but no v4 cache)',
'(v4 and v6 caches)',
'(no ISP caches)']
filenames = ['v4_cache',
'v6_cache',
'both_caches',
'no_cache']
# use helper function to get CDFs for each cases, and each metric
cdfs_ttl = get_cdfs(data, 'm_ttl_delta')
cdfs_rtt = get_cdfs(data, 'm_rtt_delta')
```
# Plotting
### Put all four cases into shared plots
- one plot for TTL
- one plot for RTT
```
markers = ['^',
's',
'o',
'd']
colors = ['blue',
'red',
'darkorchid',
'forestgreen']
labels = ['IPv4 only',
'IPv6 only',
'Both versions',
'Neither version']
## TTL
ttl_fig, ttl_ax = plt.subplots(figsize = (5, 2))
for i in range(4):
ttl_ax.plot(cdfs_ttl[i].xs, cdfs_ttl[i].ps,
label = '%s' % (labels[i]),
marker = markers[i],
linewidth = 0.5,
markersize = 3.5,
fillstyle = 'none',
color = colors[i])
# PLOT FORMATTING
ttl_ax.set_xlabel('TTL delta')
ttl_ax.set_ylabel('CDF')
ttl_ax.set_xlim([-15.3, 15.3])
ttl_ax.set_ylim([-0.05, 1.05])
ttl_ax.legend(loc = 'lower right', fontsize = 'small')
yticks = np.arange(0.0, 1.01, 0.2)
ttl_ax.set_yticks(yticks)
ttl_ax.grid(False)
ttl_ax.spines['right'].set_color('none')
ttl_ax.spines['top'].set_color('none')
ttl_ax.yaxis.set_ticks_position('left')
ttl_ax.xaxis.set_ticks_position('bottom')
ttl_ax.spines['bottom'].set_position(('axes', -0.02))
ttl_ax.spines['left'].set_position(('axes', -0.02))
## RTT
rtt_fig, rtt_ax = plt.subplots(figsize = (5, 2))
for i in range(4):
rtt_ax.plot(cdfs_rtt[i].xs, cdfs_rtt[i].ps,
label = '%s' % (labels[i]),
marker = markers[i],
linewidth = 0.5,
markersize = 2.5,
fillstyle = 'none',
color = colors[i])
# PLOT FORMATTING
# customize axes and grid appearance
rtt_ax.set_xlabel('RTT delta [ms]')
rtt_ax.set_xscale('symlog')
rtt_ax.set_ylabel('CDF')
rtt_ax.set_ylim([-0.05, 1.05])
rtt_ax.set_xlim([-1500, 1500])
rtt_ax.tick_params(axis = 'x', which = 'major', labelsize = 11)
rtt_ax.legend(loc = 'lower right', fontsize = 'small')
rtt_ax.set_yticks(yticks)
rtt_ax.grid(False)
rtt_ax.spines['right'].set_color('none')
rtt_ax.spines['top'].set_color('none')
rtt_ax.yaxis.set_ticks_position('left')
rtt_ax.xaxis.set_ticks_position('bottom')
rtt_ax.spines['bottom'].set_position(('axes', -0.02))
rtt_ax.spines['left'].set_position(('axes', -0.02))
from matplotlib.ticker import ScalarFormatter
rtt_ax.xaxis.set_major_formatter(ScalarFormatter())
# add arrows and text which indicate in which region IPv6 was better/worse than IPv4
ttl_ax.annotate('', xy = (0.4, 1.1),
xycoords = 'axes fraction',
xytext = (0.1, 1.1),
arrowprops = dict(arrowstyle = "<-"))
ttl_ax.annotate('', xy = (0.9, 1.1),
xycoords = 'axes fraction',
xytext = (0.6, 1.1),
arrowprops = dict(arrowstyle = '->'))
ttl_ax.text(-10.5, 1.2, 'IPv6 slower')
ttl_ax.text(4.5, 1.2, 'IPv6 faster')
rtt_ax.annotate('', xy = (0.4, 1.1),
xycoords = 'axes fraction',
xytext = (0.1, 1.1),
arrowprops = dict(arrowstyle = '<-'))
rtt_ax.annotate('', xy = (0.9, 1.1),
xycoords = 'axes fraction',
xytext = (0.6, 1.1),
arrowprops = dict(arrowstyle = '->'))
rtt_ax.text(-70, 1.2, 'IPv6 slower')
rtt_ax.text(2.5, 1.2, 'IPv6 faster')
# dotted vertical line to separate positive and negative delta region
ttl_ax.axvline(0, linestyle = 'dotted', color = 'black', linewidth = 0.5)
rtt_ax.axvline(0, linestyle = 'dotted', color = 'black', linewidth = 0.5)
# saving and showing plot
ttl_fig.savefig('plots/caches_ttl_comparison.pdf', bbox_inches = 'tight')
rtt_fig.savefig('plots/caches_rtt_comparison.pdf', bbox_inches = 'tight')
plt.show()
```
## Distributions of median deltas for all cases
```
# separate TTL CDFs out of list
v4, v6, both, no = cdfs_ttl
print('---- TTL v4 ----')
print('m_ttl_delta; cdf')
print('----------------')
for x, y in zip(v4.xs, v4.ps):
print('%.2f; %.5f' % (x, y))
print('\n--------------------------------------------------------\n')
print('---- TTL v6 ----')
print('m_ttl_delta; cdf')
print('----------------')
for x, y in zip(v6.xs, v6.ps):
print('%.2f; %.5f' % (x, y))
print('\n--------------------------------------------------------\n')
print('--- TTL both ---')
print('m_ttl_delta; cdf')
print('----------------')
for x, y in zip(both.xs, both.ps):
print('%.2f; %.5f' % (x, y))
print('\n--------------------------------------------------------\n')
print('--- TTL none ---')
print('m_ttl_delta; cdf')
print('----------------')
for x, y in zip(no.xs, no.ps):
print('%.2f; %.5f' % (x, y))
# separate RTT CDFs out of list
v4, v6, both, no = cdfs_rtt
print('---- RTT v4 ----')
print('m_rtt_delta; cdf')
print('----------------')
for x, y in list(zip(v4.xs, v4.ps))[0::10]:
print('%.2f; %.5f' % (x, y))
print('\n--------------------------------------------------------\n')
print('---- RTT v6 ----')
print('m_rtt_delta; cdf')
print('----------------')
for x, y in list(zip(v6.xs, v6.ps))[0::10]:
print('%.2f; %.5f' % (x, y))
print('\n--------------------------------------------------------\n')
print('--- RTT both ---')
print('m_rtt_delta; cdf')
print('----------------')
for x, y in list(zip(both.xs, both.ps))[0::10]:
print('%.2f; %.5f' % (x, y))
print('\n--------------------------------------------------------\n')
print('--- RTT none ---')
print('m_rtt_delta; cdf')
print('----------------')
for x, y in list(zip(no.xs, no.ps))[0::100]:
print('%.2f; %.5f' % (x, y))
```
| github_jupyter |
# Inference
Think Bayes, Second Edition
Copyright 2020 Allen B. Downey
License: [Attribution-NonCommercial-ShareAlike 4.0 International (CC BY-NC-SA 4.0)](https://creativecommons.org/licenses/by-nc-sa/4.0/)
```
# If we're running on Colab, install empiricaldist
# https://pypi.org/project/empiricaldist/
import sys
IN_COLAB = 'google.colab' in sys.modules
if IN_COLAB:
!pip install empiricaldist
# Get utils.py
import os
if not os.path.exists('utils.py'):
!wget https://github.com/AllenDowney/ThinkBayes2/raw/master/soln/utils.py
from utils import set_pyplot_params
set_pyplot_params()
```
Whenever people compare Bayesian inference with conventional approaches, one of the questions that comes up most often is something like, "What about p-values?"
And one of the most common examples is the comparison of two groups to see if there is a difference in their means.
In classical statistical inference, the usual tool for this scenario is a [Student's *t*-test](https://en.wikipedia.org/wiki/Student%27s_t-test), and the result is a [p-value](https://en.wikipedia.org/wiki/P-value).
This process is an example of [null hypothesis significance testing](https://en.wikipedia.org/wiki/Statistical_hypothesis_testing).
A Bayesian alternative is to compute the posterior distribution of the difference between the groups.
Then we can use that distribution to answer whatever questions we are interested in, including the most likely size of the difference, a credible interval that's likely to contain the true difference, the probability of superiority, or the probability that the difference exceeds some threshold.
To demonstrate this process, I'll solve a problem borrowed from a statistical textbook: evaluating the effect of an educational "treatment" compared to a control.
## Improving Reading Ability
We'll use data from a [Ph.D. dissertation in educational psychology](https://docs.lib.purdue.edu/dissertations/AAI8807671/) written in 1987, which was used as an example in a [statistics textbook](https://books.google.com/books/about/Introduction_to_the_practice_of_statisti.html?id=pGBNhajABlUC) from 1989 and published on [DASL](https://web.archive.org/web/20000603124754/http://lib.stat.cmu.edu/DASL/Datafiles/DRPScores.html), a web page that collects data stories.
Here's the description from DASL:
> An educator conducted an experiment to test whether new directed reading activities in the classroom will help elementary school pupils improve some aspects of their reading ability. She arranged for a third grade class of 21 students to follow these activities for an 8-week period. A control classroom of 23 third graders followed the same curriculum without the activities. At the end of the 8 weeks, all students took a Degree of Reading Power (DRP) test, which measures the aspects of reading ability that the treatment is designed to improve.
The [dataset is available here](https://web.archive.org/web/20000603124754/http://lib.stat.cmu.edu/DASL/Datafiles/DRPScores.html).
The following cell downloads the data.
```
import os
if not os.path.exists('drp_scores.csv'):
!wget https://github.com/AllenDowney/ThinkBayes2/raw/master/data/drp_scores.csv
```
I'll use Pandas to load the data into a `DataFrame`.
```
import pandas as pd
df = pd.read_csv('drp_scores.csv', skiprows=21, delimiter='\t')
df.head(3)
```
The `Treatment` column indicates whether each student was in the treated or control group.
The `Response` is their score on the test.
I'll use `groupby` to separate the data for the `Treated` and `Control` groups:
```
grouped = df.groupby('Treatment')
responses = {}
for name, group in grouped:
responses[name] = group['Response']
```
Here are CDFs of the scores for the two groups and summary statistics.
```
from empiricaldist import Cdf
from utils import decorate
for name, response in responses.items():
cdf = Cdf.from_seq(response)
cdf.plot(label=name)
decorate(xlabel='Score',
ylabel='CDF',
title='Distributions of test scores')
```
There is overlap between the distributions, but it looks like the scores are higher in the treated group.
The distribution of scores is not exactly normal for either group, but it is close enough that the normal model is a reasonable choice.
So I'll assume that in the entire population of students (not just the ones in the experiment), the distribution of scores is well modeled by a normal distribution with unknown mean and standard deviation.
I'll use `mu` and `sigma` to denote these unknown parameters,
and we'll do a Bayesian update to estimate what they are.
## Estimating Parameters
As always, we need a prior distribution for the parameters.
Since there are two parameters, it will be a joint distribution.
I'll construct it by choosing marginal distributions for each parameter and computing their outer product.
As a simple starting place, I'll assume that the prior distributions for `mu` and `sigma` are uniform.
The following function makes a `Pmf` object that represents a uniform distribution.
```
from empiricaldist import Pmf
def make_uniform(qs, name=None, **options):
"""Make a Pmf that represents a uniform distribution."""
pmf = Pmf(1.0, qs, **options)
pmf.normalize()
if name:
pmf.index.name = name
return pmf
```
`make_uniform` takes as parameters
* An array of quantities, `qs`, and
* A string, `name`, which is assigned to the index so it appears when we display the `Pmf`.
Here's the prior distribution for `mu`:
```
import numpy as np
qs = np.linspace(20, 80, num=101)
prior_mu = make_uniform(qs, name='mean')
```
I chose the lower and upper bounds by trial and error.
I'll explain how when we look at the posterior distribution.
Here's the prior distribution for `sigma`:
```
qs = np.linspace(5, 30, num=101)
prior_sigma = make_uniform(qs, name='std')
```
Now we can use `make_joint` to make the joint prior distribution.
```
from utils import make_joint
prior = make_joint(prior_mu, prior_sigma)
```
And we'll start by working with the data from the control group.
```
data = responses['Control']
data.shape
```
In the next section we'll compute the likelihood of this data for each pair of parameters in the prior distribution.
## Likelihood
We would like to know the probability of each score in the dataset for each hypothetical pair of values, `mu` and `sigma`.
I'll do that by making a 3-dimensional grid with values of `mu` on the first axis, values of `sigma` on the second axis, and the scores from the dataset on the third axis.
```
mu_mesh, sigma_mesh, data_mesh = np.meshgrid(
prior.columns, prior.index, data)
mu_mesh.shape
```
Now we can use `norm.pdf` to compute the probability density of each score for each hypothetical pair of parameters.
```
from scipy.stats import norm
densities = norm(mu_mesh, sigma_mesh).pdf(data_mesh)
densities.shape
```
The result is a 3-D array. To compute likelihoods, I'll multiply these densities along `axis=2`, which is the axis of the data:
```
likelihood = densities.prod(axis=2)
likelihood.shape
```
The result is a 2-D array that contains the likelihood of the entire dataset for each hypothetical pair of parameters.
We can use this array to update the prior, like this:
```
from utils import normalize
posterior = prior * likelihood
normalize(posterior)
posterior.shape
```
The result is a `DataFrame` that represents the joint posterior distribution.
The following function encapsulates these steps.
```
def update_norm(prior, data):
"""Update the prior based on data."""
mu_mesh, sigma_mesh, data_mesh = np.meshgrid(
prior.columns, prior.index, data)
densities = norm(mu_mesh, sigma_mesh).pdf(data_mesh)
likelihood = densities.prod(axis=2)
posterior = prior * likelihood
normalize(posterior)
return posterior
```
Here are the updates for the control and treatment groups:
```
data = responses['Control']
posterior_control = update_norm(prior, data)
data = responses['Treated']
posterior_treated = update_norm(prior, data)
```
And here's what they look like:
```
import matplotlib.pyplot as plt
from utils import plot_contour
plot_contour(posterior_control, cmap='Blues')
plt.text(49.5, 18, 'Control', color='C0')
cs = plot_contour(posterior_treated, cmap='Oranges')
plt.text(57, 12, 'Treated', color='C1')
decorate(xlabel='Mean (mu)',
ylabel='Standard deviation (sigma)',
title='Joint posterior distributions of mu and sigma')
```
Along the `x` axis, it looks like the mean score for the treated group is higher.
Along the `y` axis, it looks like the standard deviation for the treated group is lower.
If we think the treatment causes these differences, the data suggest that the treatment increases the mean of the scores and decreases their spread.
We can see these differences more clearly by looking at the marginal distributions for `mu` and `sigma`.
## Posterior Marginal Distributions
I'll use `marginal`, which we saw in <<_MarginalDistributions>>, to extract the posterior marginal distributions for the population means.
```
from utils import marginal
pmf_mean_control = marginal(posterior_control, 0)
pmf_mean_treated = marginal(posterior_treated, 0)
```
Here's what they look like:
```
pmf_mean_control.plot(label='Control')
pmf_mean_treated.plot(label='Treated')
decorate(xlabel='Population mean (mu)',
ylabel='PDF',
title='Posterior distributions of mu')
```
In both cases the posterior probabilities at the ends of the range are near zero, which means that the bounds we chose for the prior distribution are wide enough.
Comparing the marginal distributions for the two groups, it looks like the population mean in the treated group is higher.
We can use `prob_gt` to compute the probability of superiority:
```
Pmf.prob_gt(pmf_mean_treated, pmf_mean_control)
```
There is a 98% chance that the mean in the treated group is higher.
## Distribution of Differences
To quantify the magnitude of the difference between groups, we can use `sub_dist` to compute the distribution of the difference.
```
pmf_diff = Pmf.sub_dist(pmf_mean_treated, pmf_mean_control)
```
There are two things to be careful about when you use methods like `sub_dist`.
The first is that the result usually contains more elements than the original `Pmf`.
In this example, the original distributions have the same quantities, so the size increase is moderate.
```
len(pmf_mean_treated), len(pmf_mean_control), len(pmf_diff)
```
In the worst case, the size of the result can be the product of the sizes of the originals.
The other thing to be careful about is plotting the `Pmf`.
In this example, if we plot the distribution of differences, the result is pretty noisy.
```
pmf_diff.plot()
decorate(xlabel='Difference in population means',
ylabel='PDF',
title='Posterior distribution of difference in mu')
```
There are two ways to work around that limitation. One is to plot the CDF, which smooths out the noise:
```
cdf_diff = pmf_diff.make_cdf()
cdf_diff.plot()
decorate(xlabel='Difference in population means',
ylabel='CDF',
title='Posterior distribution of difference in mu')
```
The other option is to use kernel density estimation (KDE) to make a smooth approximation of the PDF on an equally-spaced grid, which is what this function does:
```
from scipy.stats import gaussian_kde
def kde_from_pmf(pmf, n=101):
"""Make a kernel density estimate for a PMF."""
kde = gaussian_kde(pmf.qs, weights=pmf.ps)
qs = np.linspace(pmf.qs.min(), pmf.qs.max(), n)
ps = kde.evaluate(qs)
pmf = Pmf(ps, qs)
pmf.normalize()
return pmf
```
`kde_from_pmf` takes as parameters a `Pmf` and the number of places to evaluate the KDE.
It uses `gaussian_kde`, which we saw in <<_KernelDensityEstimation>>, passing the probabilities from the `Pmf` as weights.
This makes the estimated densities higher where the probabilities in the `Pmf` are higher.
Here's what the kernel density estimate looks like for the `Pmf` of differences between the groups.
```
kde_diff = kde_from_pmf(pmf_diff)
kde_diff.plot()
decorate(xlabel='Difference in means',
ylabel='PDF',
title='Posterior distribution of difference in mu')
```
The mean of this distribution is almost 10 points on a test where the mean is around 45, so the effect of the treatment seems to be substantial.
```
pmf_diff.mean()
```
We can use `credible_interval` to compute a 90% credible interval.
```
pmf_diff.credible_interval(0.9)
```
Based on this interval, we are pretty sure the treatment improves test scores by 2 to 17 points.
## Using Summary Statistics
In this example the dataset is not very big, so it doesn't take too long to compute the probability of every score under every hypothesis.
But the result is a 3-D array; for larger datasets, it might be too big to compute practically.
Also, with larger datasets the likelihoods get very small, sometimes so small that we can't compute them with floating-point arithmetic.
That's because we are computing the probability of a particular dataset; the number of possible datasets is astronomically big, so the probability of any of them is very small.
An alternative is to compute a summary of the dataset and compute the likelihood of the summary.
For example, if we compute the mean and standard deviation of the data, we can compute the likelihood of those summary statistics under each hypothesis.
As an example, suppose we know that the actual mean of the population, $\mu$, is 42 and the actual standard deviation, $\sigma$, is 17.
```
mu = 42
sigma = 17
```
Now suppose we draw a sample from this distribution with sample size `n=20`, and compute the mean of the sample, which I'll call `m`, and the standard deviation of the sample, which I'll call `s`.
And suppose it turns out that:
```
n = 20
m = 41
s = 18
```
The summary statistics, `m` and `s`, are not too far from the parameters $\mu$ and $\sigma$, so it seems like they are not too unlikely.
To compute their likelihood, we will take advantage of three results from mathematical statistics:
* Given $\mu$ and $\sigma$, the distribution of `m` is normal with parameters $\mu$ and $\sigma/\sqrt{n}$;
* The distribution of $s$ is more complicated, but if we compute the transform $t = n s^2 / \sigma^2$, the distribution of $t$ is chi-squared with parameter $n-1$; and
* According to [Basu's theorem](https://en.wikipedia.org/wiki/Basu%27s_theorem), `m` and `s` are independent.
So let's compute the likelihood of `m` and `s` given $\mu$ and $\sigma$.
First I'll create a `norm` object that represents the distribution of `m`.
```
dist_m = norm(mu, sigma/np.sqrt(n))
```
This is the "sampling distribution of the mean".
We can use it to compute the likelihood of the observed value of `m`, which is 41.
```
like1 = dist_m.pdf(m)
like1
```
Now let's compute the likelihood of the observed value of `s`, which is 18.
First, we compute the transformed value `t`:
```
t = n * s**2 / sigma**2
t
```
Then we create a `chi2` object to represent the distribution of `t`:
```
from scipy.stats import chi2
dist_s = chi2(n-1)
```
Now we can compute the likelihood of `t`:
```
like2 = dist_s.pdf(t)
like2
```
Finally, because `m` and `s` are independent, their joint likelihood is the product of their likelihoods:
```
like = like1 * like2
like
```
Now we can compute the likelihood of the data for any values of $\mu$ and $\sigma$, which we'll use in the next section to do the update.
## Update With Summary Statistics
Now we're ready to do an update.
I'll compute summary statistics for the two groups.
```
summary = {}
for name, response in responses.items():
summary[name] = len(response), response.mean(), response.std()
summary
```
The result is a dictionary that maps from group name to a tuple that contains the sample size, `n`, the sample mean, `m`, and the sample standard deviation `s`, for each group.
I'll demonstrate the update with the summary statistics from the control group.
```
n, m, s = summary['Control']
```
I'll make a mesh with hypothetical values of `mu` on the `x` axis and values of `sigma` on the `y` axis.
```
mus, sigmas = np.meshgrid(prior.columns, prior.index)
mus.shape
```
Now we can compute the likelihood of seeing the sample mean, `m`, for each pair of parameters.
```
like1 = norm(mus, sigmas/np.sqrt(n)).pdf(m)
like1.shape
```
And we can compute the likelihood of the sample standard deviation, `s`, for each pair of parameters.
```
ts = n * s**2 / sigmas**2
like2 = chi2(n-1).pdf(ts)
like2.shape
```
Finally, we can do the update with both likelihoods:
```
posterior_control2 = prior * like1 * like2
normalize(posterior_control2)
```
To compute the posterior distribution for the treatment group, I'll put the previous steps in a function:
```
def update_norm_summary(prior, data):
"""Update a normal distribution using summary statistics."""
n, m, s = data
mu_mesh, sigma_mesh = np.meshgrid(prior.columns, prior.index)
like1 = norm(mu_mesh, sigma_mesh/np.sqrt(n)).pdf(m)
like2 = chi2(n-1).pdf(n * s**2 / sigma_mesh**2)
posterior = prior * like1 * like2
normalize(posterior)
return posterior
```
Here's the update for the treatment group:
```
data = summary['Treated']
posterior_treated2 = update_norm_summary(prior, data)
```
And here are the results.
```
plot_contour(posterior_control2, cmap='Blues')
plt.text(49.5, 18, 'Control', color='C0')
cs = plot_contour(posterior_treated2, cmap='Oranges')
plt.text(57, 12, 'Treated', color='C1')
decorate(xlabel='Mean (mu)',
ylabel='Standard deviation (sigma)',
title='Joint posterior distributions of mu and sigma')
```
Visually, these posterior joint distributions are similar to the ones we computed using the entire dataset, not just the summary statistics.
But they are not exactly the same, as we can see by comparing the marginal distributions.
## Comparing Marginals
Again, let's extract the marginal posterior distributions.
```
from utils import marginal
pmf_mean_control2 = marginal(posterior_control2, 0)
pmf_mean_treated2 = marginal(posterior_treated2, 0)
```
And compare them to results we got using the entire dataset (the dashed lines).
```
pmf_mean_control.plot(color='C5', linestyle='dashed')
pmf_mean_control2.plot(label='Control')
pmf_mean_treated.plot(color='C5', linestyle='dashed')
pmf_mean_treated2.plot(label='Treated')
decorate(xlabel='Population mean',
ylabel='PDF',
title='Posterior distributions of mu')
```
The posterior distributions based on summary statistics are similar to the posteriors we computed using the entire dataset, but in both cases they are shorter and a little wider.
That's because the update with summary statistics is based on the implicit assumption that the distribution of the data is normal.
But it's not; as a result, when we replace the dataset with the summary statistics, we lose some information about the true distribution of the data.
With less information, we are less certain about the parameters.
## Proof By Simulation
The update with summary statistics is based on theoretical distributions, and it seems to work, but I think it is useful to test theories like this, for a few reasons:
* It confirms that our understanding of the theory is correct,
* It confirms that the conditions where we apply the theory are conditions where the theory holds,
* It confirms that the implementation details are correct. For many distributions, there is more than one way to specify the parameters. If you use the wrong specification, this kind of testing will help you catch the error.
In this section I'll use simulations to show that the distribution of the sample mean and standard deviation is as I claimed.
But if you want to take my word for it, you can skip this section and the next.
Let's suppose that we know the actual mean and standard deviation of the population:
```
mu = 42
sigma = 17
```
I'll create a `norm` object to represent this distribution.
```
dist = norm(mu, sigma)
```
`norm` provides `rvs`, which generates random values from the distribution.
We can use it to simulate 1000 samples, each with sample size `n=20`.
```
n = 20
samples = dist.rvs((1000, n))
samples.shape
```
The result is an array with 1000 rows, each containing a sample or 20 simulated test scores.
If we compute the mean of each row, the result is an array that contains 1000 sample means; that is, each value is the mean of a sample with `n=20`.
```
sample_means = samples.mean(axis=1)
sample_means.shape
```
Now, let's compare the distribution of these means to `dist_m`.
I'll use `pmf_from_dist` to make a discrete approximation of `dist_m`:
```
def pmf_from_dist(dist, low, high):
"""Make a discrete approximation of a continuous distribution.
dist: SciPy dist object
low: low end of range
high: high end of range
returns: normalized Pmf
"""
qs = np.linspace(low, high, 101)
ps = dist.pdf(qs)
pmf = Pmf(ps, qs)
pmf.normalize()
return pmf
```
`pmf_from_dist` takes an object representing a continuous distribution, evaluates its probability density function at equally space points between `low` and `high`, and returns a normalized `Pmf` that approximates the distribution.
I'll use it to evaluate `dist_m` over a range of six standard deviations.
```
low = dist_m.mean() - dist_m.std() * 3
high = dist_m.mean() + dist_m.std() * 3
pmf_m = pmf_from_dist(dist_m, low, high)
```
Now let's compare this theoretical distribution to the means of the samples.
I'll use `kde_from_sample` to estimate their distribution and evaluate it in the same locations as `pmf_m`.
```
from utils import kde_from_sample
qs = pmf_m.qs
pmf_sample_means = kde_from_sample(sample_means, qs)
```
The following figure shows the two distributions.
```
pmf_m.plot(label='Theoretical distribution',
style=':', color='C5')
pmf_sample_means.plot(label='KDE of sample means')
decorate(xlabel='Mean score',
ylabel='PDF',
title='Distribution of the mean')
```
The theoretical distribution and the distribution of sample means are in accord.
## Checking Standard Deviation
Let's also check that the standard deviations follow the distribution we expect.
First I'll compute the standard deviation for each of the 1000 samples.
```
sample_stds = samples.std(axis=1)
sample_stds.shape
```
Now we'll compute the transformed values, $t = n s^2 / \sigma^2$.
```
transformed = n * sample_stds**2 / sigma**2
```
We expect the transformed values to follow a chi-square distribution with parameter $n-1$.
SciPy provides `chi2`, which we can use to represent this distribution.
```
from scipy.stats import chi2
dist_s = chi2(n-1)
```
We can use `pmf_from_dist` again to make a discrete approximation.
```
low = 0
high = dist_s.mean() + dist_s.std() * 4
pmf_s = pmf_from_dist(dist_s, low, high)
```
And we'll use `kde_from_sample` to estimate the distribution of the sample standard deviations.
```
qs = pmf_s.qs
pmf_sample_stds = kde_from_sample(transformed, qs)
```
Now we can compare the theoretical distribution to the distribution of the standard deviations.
```
pmf_s.plot(label='Theoretical distribution',
style=':', color='C5')
pmf_sample_stds.plot(label='KDE of sample std',
color='C1')
decorate(xlabel='Standard deviation of scores',
ylabel='PDF',
title='Distribution of standard deviation')
```
The distribution of transformed standard deviations agrees with the theoretical distribution.
Finally, to confirm that the sample means and standard deviations are independent, I'll compute their coefficient of correlation:
```
np.corrcoef(sample_means, sample_stds)[0][1]
```
Their correlation is near zero, which is consistent with their being independent.
So the simulations confirm the theoretical results we used to do the update with summary statistics.
We can also use `kdeplot` from Seaborn to see what their joint distribution looks like.
```
import seaborn as sns
sns.kdeplot(x=sample_means, y=sample_stds)
decorate(xlabel='Mean (mu)',
ylabel='Standard deviation (sigma)',
title='Joint distribution of mu and sigma')
```
It looks like the axes of the ellipses are aligned with the axes, which indicates that the variables are independent.
## Summary
In this chapter we used a joint distribution to represent prior probabilities for the parameters of a normal distribution, `mu` and `sigma`.
And we updated that distribution two ways: first using the entire dataset and the normal PDF; then using summary statistics, the normal PDF, and the chi-square PDF.
Using summary statistics is computationally more efficient, but it loses some information in the process.
Normal distributions appear in many domains, so the methods in this chapter are broadly applicable. The exercises at the end of the chapter will give you a chance to apply them.
## Exercises
**Exercise:** Looking again at the posterior joint distribution of `mu` and `sigma`, it seems like the standard deviation of the treated group might be lower; if so, that would suggest that the treatment is more effective for students with lower scores.
But before we speculate too much, we should estimate the size of the difference and see whether it might actually be 0.
Extract the marginal posterior distributions of `sigma` for the two groups.
What is the probability that the standard deviation is higher in the control group?
Compute the distribution of the difference in `sigma` between the two groups. What is the mean of this difference? What is the 90% credible interval?
```
# Solution goes here
# Solution goes here
# Solution goes here
# Solution goes here
# Solution goes here
# Solution goes here
# Solution goes here
```
## Exercise
An [effect size](http://en.wikipedia.org/wiki/Effect_size) is a statistic intended to quantify the magnitude of a phenomenon.
If the phenomenon is a difference in means between two groups, a common way to quantify it is Cohen's effect size, denoted $d$.
If the parameters for Group 1 are $(\mu_1, \sigma_1)$, and the
parameters for Group 2 are $(\mu_2, \sigma_2)$, Cohen's
effect size is
$ d = \frac{\mu_1 - \mu_2}{(\sigma_1 + \sigma_2)/2} $
Use the joint posterior distributions for the two groups to compute the posterior distribution for Cohen's effect size.
If we try enumerate all pairs from the two distributions, it takes too
long so we'll use random sampling.
The following function takes a joint posterior distribution and returns a sample of pairs.
It uses some features we have not seen yet, but you can ignore the details for now.
```
def sample_joint(joint, size):
"""Draw a sample from a joint distribution.
joint: DataFrame representing a joint distribution
size: sample size
"""
pmf = Pmf(joint.transpose().stack())
return pmf.choice(size)
```
Here's how we can use it to sample pairs from the posterior distributions for the two groups.
```
sample_treated = sample_joint(posterior_treated, 1000)
sample_treated.shape
sample_control = sample_joint(posterior_control, 1000)
sample_control.shape
```
The result is an array of tuples, where each tuple contains a possible pair of values for $\mu$ and $\sigma$.
Now you can loop through the samples, compute the Cohen effect size for each, and estimate the distribution of effect sizes.
```
# Solution goes here
# Solution goes here
# Solution goes here
# Solution goes here
# Solution goes here
# Solution goes here
```
**Exercise:** This exercise is inspired by [a question that appeared on Reddit](https://www.reddit.com/r/statistics/comments/hcvl2j/q_reverse_empirical_distribution_rule_question/).
An instructor announces the results of an exam like this, "The average score on this exam was 81. Out of 25 students, 5 got more than 90, and I am happy to report that no one failed (got less than 60)."
Based on this information, what do you think the standard deviation of scores was?
You can assume that the distribution of scores is approximately normal. And let's assume that the sample mean, 81, is actually the population mean, so we only have to estimate `sigma`.
Hint: To compute the probability of a score greater than 90, you can use `norm.sf`, which computes the survival function, also known as the complementary CDF, or `1 - cdf(x)`.
```
# Solution goes here
# Solution goes here
# Solution goes here
# Solution goes here
# Solution goes here
# Solution goes here
# Solution goes here
# Solution goes here
# Solution goes here
# Solution goes here
# Solution goes here
# Solution goes here
```
**Exercise:** I have a soft spot for crank science, so this exercise is about the [Variability Hypothesis](http://en.wikipedia.org/wiki/Variability_hypothesis), which
> "originated in the early nineteenth century with Johann Meckel, who argued that males have a greater range of ability than females, especially in intelligence. In other words, he believed that most geniuses and most mentally retarded people are men. Because he considered males to be the 'superior animal', Meckel concluded that females' lack of variation was a sign of inferiority."
I particularly like that last part because I suspect that if it turned out that women were _more_ variable, Meckel would have taken that as a sign of inferiority, too.
Nevertheless, the Variability Hypothesis suggests an exercise we can use to practice the methods in this chapter. Let's look at the distribution of heights for men and women in the U.S. and see who is more variable.
I used 2018 data from the CDC’s [Behavioral Risk Factor Surveillance System](https://www.cdc.gov/brfss/annual_data/annual_2018.html) (BRFSS), which includes self-reported heights from 154407 men and 254722 women.
Here's what I found:
* The average height for men is 178 cm; the average height for women is 163 cm. So men are taller on average; no surprise there.
* For men the standard deviation is 8.27 cm; for women it is 7.75 cm. So in absolute terms, men's heights are more variable.
But to compare variability between groups, it is more meaningful to use the [coefficient of variation](https://en.wikipedia.org/wiki/Coefficient_of_variation) (CV), which is the standard deviation divided by the mean. It is a dimensionless measure of variability relative to scale.
For men CV is 0.0465; for women it is 0.0475.
The coefficient of variation is higher for women, so this dataset provides evidence against the Variability Hypothesis. But we can use Bayesian methods to make that conclusion more precise.
Use these summary statistics to compute the posterior distribution of `mu` and `sigma` for the distributions of male and female height.
Use `Pmf.div_dist` to compute posterior distributions of CV.
Based on this dataset and the assumption that the distribution of height is normal, what is the probability that the coefficient of variation is higher for men?
What is the most likely ratio of the CVs and what is the 90% credible interval for that ratio?
Hint: Use different prior distributions for the two groups, and chose them so they cover all parameters with non-negligible probability.
Also, you might find this function helpful:
```
def get_posterior_cv(joint):
"""Get the posterior distribution of CV.
joint: joint distribution of mu and sigma
returns: Pmf representing the smoothed posterior distribution
"""
pmf_mu = marginal(joint, 0)
pmf_sigma = marginal(joint, 1)
pmf_cv = Pmf.div_dist(pmf_sigma, pmf_mu)
return kde_from_pmf(pmf_cv)
# Solution goes here
# Solution goes here
# Solution goes here
# Solution goes here
# Solution goes here
# Solution goes here
# Solution goes here
# Solution goes here
# Solution goes here
```
| github_jupyter |
## MESA
Mesa is agent-based modeling (or ABM) framework in Python. It enables users to quickly develop ABMs. It provides a variety of convenience components often used in ABMs, like different kinds of spaces within which agents can interact, different kinds of schedulers for controlling which agents in what order are making their moves, and basic support for dealing with the intrinisc stochastic nature of ABMs. MESA is ideally suited for learning agent-based modeling. It is less suited for developing large-scale computationally heavy ABMs. Given that MESA is a python library, and its focus on learning ABM, we have chosen to use MESA. The documentation of MESA can be found online: https://mesa.readthedocs.io/en/master/
Any Agent Based Model is typically composed of a collection of **Agent**, which are positioned in a **Space**. The Agents interact with each other typically based on how close they are in the Space, as well as sometimes with the Space itself. Agent Based Models are dynamic so they show some kind of pattern over time. To create this dynamic, Agent Based Models use **ticks**. A tick is like a timestep. In each tick, all agents are given the option to act. The order in which Agents can act is very important because Agent Based Models are prone to showing path dependence. Therefore, the order in which agents get to act is typcially handled by a **Scheduler**.
Let's build a very simple Agent Based Model, using MESA which contains these four components: Agents, Space, Schedulers, and ticks. The model we will build is a simple model of an economy. It consists of a collection of agent which move around in a gridded space. If, after having moved, an agent find itself in a gridcell occopied by other agents, the agent will tranfer one unit of wealth to a randomly selected other agent in the same grid cell.
To implement this model, we will extend the Agent and Model class provided by MESA. As space, we will use a MultiGrid. This is a gridded (think excel like) space where more than one agent can occupy the same cell. As scheduler, we will use the RandomActiviation. This means that at each tick, all agents in a random order get to make their move.
So we start with importing the classes from MESA that we need.
```
from mesa import Agent, Model
from mesa.time import RandomActivation
from mesa.space import MultiGrid
```
Next, we can extend the MESA Model class. For this, we have to implement the `__init__` and the `step` method. The `__init__` is the initial setup of the model. So we give our model a space (i.e., `self.grid = MultiGrid(width, height, True)`. We also give the model a scheduler (i.e., `self.schedule = RandomActivation(self)`. Last, we need to populate the grid with agents. We do this by first creating an Agend and next placing it at a randomly selected grid sell (i.e., the for loop). The step method is simple: we just invoke the step method on the scheduler (i.e., `self.schedule.step()`).
```
class MoneyModel(Model):
"""A model with some number of agents.
Parameters
N : int
the number of agents in the space
width : int
the width of the space
height : int
the height of the space
"""
def __init__(self, N, width, height):
self.num_agents = N
# the space within which agents move
self.grid = MultiGrid(width, height, True)
# the scheduler
self.schedule = RandomActivation(self)
# initialize the model by creating N agents
# positioned at a random location in the grid
for i in range(self.num_agents):
agent = MoneyAgent(i, self)
self.schedule.add(agent)
# Add the agent to a random grid cell
x = self.random.randrange(self.grid.width)
y = self.random.randrange(self.grid.height)
self.grid.place_agent(agent, (x, y))
def step(self):
self.schedule.step()
```
We also need to extend the Agent class that comes with MESA. In the simplest version, we would again only need to implement the `__init__` and the `step` method. However, in this case, we seperate the `step` method into 2 activities: moving to a randomly selected neighboring cell, and possibe giving money. We place both activities in their own methods to clearly seperate these two actions (i.e., we add a `move` and `give_money` method to the Agent class).
For the move method, we can use the grid of the model to get the neigboring cells, randomly pick one, and then inform the grid to move the agent to this randomly selected grid cell.
For the give_money method, we have to first check if the agent has any wealth to give. If not, nothing happens. Otherwise, we aks the grid for all agents occuping our current grid cell. If there is more than one agent, we randomly select one agent to give wealth to.
Question: look carefully at the give_money method. Do you notice anything odd in how this is currently implemented?
answer: the agent randomly select from the list of agents currently occupying the cell. This list includes also itself, so the agent might give wealth to itself.
```
class MoneyAgent(Agent):
""" An agent with fixed initial wealth."""
def __init__(self, unique_id, model):
super().__init__(unique_id, model)
self.wealth = 1
def move(self):
possible_steps = self.model.grid.get_neighborhood(
self.pos, moore=True, include_center=False)
new_position = self.random.choice(possible_steps)
self.model.grid.move_agent(self, new_position)
def give_money(self):
cellmates = self.model.grid.get_cell_list_contents([self.pos])
if len(cellmates) > 1:
other = self.random.choice(cellmates)
other.wealth += 1
self.wealth -= 1
def step(self):
self.move()
if self.wealth > 0:
self.give_money()
```
This completes the setup of the model. We can now instantiate the model, and run it for a number of ticks
```
model = MoneyModel(80, 10, 10)
n_ticks = 10
for _ in range(n_ticks):
model.step()
```
As a last step, we can visualize the model results. For example, we can show a histogram of the distribution of wealth over the agents like this
```
import matplotlib.pyplot as plt
import seaborn as sns
sns.set_style('white')
agent_wealth = [a.wealth for a in model.schedule.agents]
fig, ax = plt.subplots()
ax.hist(agent_wealth)
sns.despine()
plt.show()
```
We can also take a look at the grid itself like this:
```
fig, ax = plt.subplots()
agent_counts = np.zeros((model.grid.width, model.grid.height))
for cell in model.grid.coord_iter():
cell_content, x, y = cell
agent_count = len(cell_content)
agent_counts[x][y] = agent_count
img = ax.imshow(agent_counts, interpolation='nearest')
fig.colorbar(img, ax=ax)
plt.show()
```
Question: run the model a few times and replot the results. Do you notice anything strange and can you explain this?
Answer: There is quite some randomness in the model: where the original agents are located, where they move to, and to whom to give wealth. Everytime you rerun the model different random numbers are used and thus you get different results. This is a key and important difference with system dynamics models. ABMs are stochastic, while SD models are deterministic. In future assignments, we will look at this in more detail and also show how we can control the random numbers that are being generated so that the model becomes deterministic. This is desirable for debugging and reproducibility.
| github_jupyter |
<table class="ee-notebook-buttons" align="left">
<td><a target="_blank" href="https://github.com/giswqs/earthengine-py-notebooks/tree/master/Image/image_stats_by_band.ipynb"><img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" /> View source on GitHub</a></td>
<td><a target="_blank" href="https://nbviewer.jupyter.org/github/giswqs/earthengine-py-notebooks/blob/master/Image/image_stats_by_band.ipynb"><img width=26px src="https://upload.wikimedia.org/wikipedia/commons/thumb/3/38/Jupyter_logo.svg/883px-Jupyter_logo.svg.png" />Notebook Viewer</a></td>
<td><a target="_blank" href="https://mybinder.org/v2/gh/giswqs/earthengine-py-notebooks/master?filepath=Image/image_stats_by_band.ipynb"><img width=58px src="https://mybinder.org/static/images/logo_social.png" />Run in binder</a></td>
<td><a target="_blank" href="https://colab.research.google.com/github/giswqs/earthengine-py-notebooks/blob/master/Image/image_stats_by_band.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" /> Run in Google Colab</a></td>
</table>
## Install Earth Engine API
Install the [Earth Engine Python API](https://developers.google.com/earth-engine/python_install) and [geehydro](https://github.com/giswqs/geehydro). The **geehydro** Python package builds on the [folium](https://github.com/python-visualization/folium) package and implements several methods for displaying Earth Engine data layers, such as `Map.addLayer()`, `Map.setCenter()`, `Map.centerObject()`, and `Map.setOptions()`.
The following script checks if the geehydro package has been installed. If not, it will install geehydro, which automatically install its dependencies, including earthengine-api and folium.
```
import subprocess
try:
import geehydro
except ImportError:
print('geehydro package not installed. Installing ...')
subprocess.check_call(["python", '-m', 'pip', 'install', 'geehydro'])
```
Import libraries
```
import ee
import folium
import geehydro
```
Authenticate and initialize Earth Engine API. You only need to authenticate the Earth Engine API once.
```
try:
ee.Initialize()
except Exception as e:
ee.Authenticate()
ee.Initialize()
```
## Create an interactive map
This step creates an interactive map using [folium](https://github.com/python-visualization/folium). The default basemap is the OpenStreetMap. Additional basemaps can be added using the `Map.setOptions()` function.
The optional basemaps can be `ROADMAP`, `SATELLITE`, `HYBRID`, `TERRAIN`, or `ESRI`.
```
Map = folium.Map(location=[40, -100], zoom_start=4)
Map.setOptions('HYBRID')
```
## Add Earth Engine Python script
```
image = ee.Image('USDA/NAIP/DOQQ/m_3712213_sw_10_1_20140613')
Map.setCenter(-122.466123, 37.769833, 17)
Map.addLayer(image, {'bands': ['N', 'R','G']}, 'NAIP')
geometry = image.geometry()
means = image.reduceRegions(geometry, ee.Reducer.mean().forEachBand(image), 10)
print(means.getInfo())
```
## Display Earth Engine data layers
```
Map.setControlVisibility(layerControl=True, fullscreenControl=True, latLngPopup=True)
Map
```
| github_jupyter |
<h1>Table of Contents<span class="tocSkip"></span></h1>
<div class="toc"><ul class="toc-item"><li><span><a href="#The-mulearn.optimization-module" data-toc-modified-id="The-mulearn.optimization-module-1"><span class="toc-item-num">1 </span>The <code>mulearn.optimization</code> module</a></span></li></ul></div>
# The `mulearn.optimization` module
> The `mulearn.optimization` module contains the implementations of
the optimization processes behind fuzzy inference.
```
# default_exp optimization
# export
import numpy as np
import itertools as it
from warnings import warn
from collections.abc import Iterable
import logging
from gurobipy import GurobiError
import mulearn.kernel as kernel
logger = logging.getLogger(__name__)
try:
import gurobipy as gpy
gurobi_ok = True
except ModuleNotFoundError:
logger.warning('gurobi not available')
gurobi_ok = False
try:
import tensorflow as tf
tensorflow_ok = True
logging.getLogger('tensorflow').setLevel(logging.ERROR)
except ModuleNotFoundError:
logger.warning('tensorflow not available')
tensorflow_ok = False
try:
import tqdm
tqdm_ok = True
except ModuleNotFoundError:
logger.warning('tqdm not available')
tqdm_ok = False
#hide
from nbdev.showdoc import show_doc
#export
def solve_optimization_tensorflow(xs, mus,
c=1.0, k=kernel.GaussianKernel(),
init='fixed',
init_bound=0.1,
init_val=0.01,
n_iter=100,
optimizer=tf.optimizers.Adam(learning_rate=1e-4) \
if tensorflow_ok else None,
tracker=tqdm.trange if tqdm_ok else range,
penalization=10):
'''Builds and solves the constrained optimization problem on the basis
of the fuzzy learning procedure using the TensorFlow API.
- xs: iterable of objects
- mus: iterable of membership values for the objects in xs
- c: constant managing the trade-off in joint radius/error optimization
- k: kernel function to be used
- init: initialization method for weights (string)
- init_bound: absolute value of the extremes of the interval used
for random initialization of weights (float)
- init_val: value used for initializing weights to a constant (float)
- n_iter: number of iteration of the optimization process (int)
- optimizer: optiimzation algorithm to be used
- tracker: tool to graphically depict the optimization progress
Returns: a lists containing the optimal values for the independent
variables chis of the problem
Throws:
- ValueError if optimization fails or if tensorflow is not installed
'''
if not tensorflow_ok:
raise ValueError('tensorflow not available')
m = len(xs)
if type(init) == str and init == 'fixed':
chis = [tf.Variable(init_val, name=f'chi_{i}',
trainable=True, dtype=tf.float32)
for i in range(m)]
elif type(init) == str and init == 'random':
chis = [tf.Variable(ch, name=f'chi_{i}',
trainable=True, dtype=tf.float32)
for i, ch in enumerate(np.random.uniform(-init_bound,
init_bound, m))]
elif isinstance(init, Iterable):
chis = [tf.Variable(ch, name=f'chi_{i}',
trainable=True, dtype=tf.float32)
for i, ch in enumerate(init)]
else:
raise ValueError("init should either be set to 'fixed', "
"'random', or to a list of initial values.")
if type(k) is kernel.PrecomputedKernel:
gram = k.kernel_computations
else:
gram = np.array([[k.compute(x1, x2) for x1 in xs] for x2 in xs])
def obj():
kernels = tf.constant(gram, dtype='float32')
v = tf.tensordot(tf.linalg.matvec(kernels, chis), chis, axes=1)
v -= tf.tensordot(chis, [k.compute(x_i, x_i) for x_i in xs], axes=1)
v += penalization * tf.math.maximum(0, 1 - sum(chis))
v += penalization * tf.math.maximum(0, sum(chis) - 1)
if c < np.inf:
for ch, m in zip(chis, mus):
v += penalization * tf.math.maximum(0, ch - c*m)
v += penalization * tf.math.maximum(0, c*(1-m) - ch)
return v
for i in tracker(n_iter):
#old_chis = np.array([ch.numpy() for ch in chis])
optimizer.minimize(obj, var_list=chis)
#new_chis = np.array([ch.numpy() for ch in chis])
return [ch.numpy() for ch in chis]
xs = [1, 2, 5, 5.5, 7, 8, 9.5, 10]
mus = [1, 1, 1, 0.9, 0.4, 0.1, 0, 0]
solve_optimization_tensorflow(xs, mus, n_iter=1000, penalization=10000, tracker=range)
#export
def solve_optimization_gurobi(xs,
mus,
c=1.0,
k=kernel.GaussianKernel(),
time_limit=10*60,
adjustment=0):
'''Builds and solves the constrained optimization problem on the basis
of the fuzzy learning procedure using the gurobi API.
- `xs`: objects in training set (iterable).
- `mus`: membership values for the objects in xs (iterable).
- `c`: constant managing the trade-off in joint radius/error optimization
(float).
- `k`: kernel function to be used (kernel.Kernel).
- `time_limit`: time in seconds before stopping the optimization process
(int).
- `adjustment`: diagonal adjustment in order to deal with non PSD
matrices (float or 'auto' for automatic adjustment).
Return a lists containing the optimal values for the independent
variables chis of the problem.
Throws:
- ValueError if optimization fails or if gurobi is not installed
'''
if not gurobi_ok:
raise ValueError('gurobi not available')
m = len(xs)
model = gpy.Model('mulearn')
model.setParam('OutputFlag', 0)
model.setParam('TimeLimit', time_limit)
for i in range(m):
if c < np.inf:
model.addVar(name=f'chi_{i}', lb=-c*(1-mus[i]), ub=c*mus[i],
vtype=gpy.GRB.CONTINUOUS)
else:
model.addVar(name=f'chi_{i}', vtype=gpy.GRB.CONTINUOUS)
model.update()
chis = model.getVars()
obj = gpy.QuadExpr()
for i, j in it.product(range(m), range(m)):
obj.add(chis[i] * chis[j], k.compute(xs[i], xs[j]))
for i in range(m):
obj.add(-1 * chis[i] * k.compute(xs[i], xs[i]))
if adjustment and adjustment != 'auto':
for i in range(m):
obj.add(adjustment * chis[i] * chis[i])
model.setObjective(obj, gpy.GRB.MINIMIZE)
constEqual = gpy.LinExpr()
constEqual.add(sum(chis), 1.0)
model.addConstr(constEqual, gpy.GRB.EQUAL, 1)
try:
model.optimize()
except GurobiError as e:
if adjustment == 'auto':
s = e.message
a = float(s[s.find(' of ')+4:s.find(' would')])
logger.warning(f'non-diagonal Gram matrix, retrying with adjustment {a}')
for i in range(m):
obj.add(a * chis[i] * chis[i])
model.setObjective(obj, gpy.GRB.MINIMIZE)
model.optimize()
else:
raise e
if model.Status != gpy.GRB.OPTIMAL:
raise ValueError('optimal solution not found!')
return [ch.x for ch in chis]
try:
print(solve_optimization_gurobi(xs, mus))
except ValueError:
pass
#export
def solve_optimization(xs, mus, c=1.0, k=kernel.GaussianKernel(),
solve_strategy=solve_optimization_tensorflow,
**solve_strategy_args):
'''Builds and solves the constrained optimization problem on the basis
of the fuzzy learning procedure.
- xs: objects in training set (iterable).
- mus: membership values for the objects in xs (iterable).
- c: constant managing the trade-off in joint radius/error optimization
(float).
- k: kernel function to be used (kernel.Kernel).
- solve_strategy: algorithm to be used in order to numerically solve the
optimization problem.
- solve_strategy_args: optional parameters for the optimization
algorithm.
Return a lists containing the optimal values for the independent
variables chis of the problem
Throws:
- ValueError if c is non-positive or if xs and mus have different lengths
'''
if c <= 0:
raise ValueError('c should be positive')
mus = np.array(mus)
chis = solve_strategy(xs, mus, c, k, **solve_strategy_args)
chis_opt = [np.clip(ch, l, u) for ch,l,u in zip(chis, -c*(1-mus), c*mus)]
return chis_opt
solve_optimization(xs, mus)
try:
print(solve_optimization(xs, mus,
solve_strategy=solve_optimization_gurobi))
except ValueError:
pass
```
| github_jupyter |
## 使用 tf.keras 搭建神经网络-扩展
使用 “六步法” 搭建神经网络可以快速高效的完成简单的任务,特别是有着各种 API 的加持,更使得这些编码过程非常顺利。但是面对一些实际的任务场景,是否还可以有更多的思考?比如:
+ Q1: 如果有了自己本领域的数据与标签,如何给出 `x_train`、`y_train`、`x_test`、`y_test`?
+ Q2: 如何解决数据量过少,模型见识不足,导致训练出来的模型泛化力弱的问题?
+ Q3: 如果模型训练每次都从零开始,是件很不划算的事,如何改善呢?
+ Q4: 如何获取各层网络最优的参数并保存,并在下一次的使用中复现出模型?
+ Q5: 训练效果如何可视化?
+ Q6: 训练出来的模型如何应用?
以上只是一些简单的思考,但是针对以上问题,提出以下的解决方案,为之前的“六步法”进行一些扩展,使得其更具有实用性:
+ A1: 自制数据集,解决本领域应用
+ A2: 数据增强,扩充数据集
+ A3: 断点续训,存取最优模型
+ A4: 参数提取,把参数存入文本
+ A5: 可视化 acc 和 loss 曲线,查看训练效果
+ A6: 应用模型对新数据进行预测
接下来将在之前使用简单 “六步法” 实现手写数字识别的基础上进行改进
```
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
from PIL import Image
import os
```
### 1. 自制数据集,解决本领域应用
这一步,需要使用自定义格式的 MNIST 数据集,在目录 `./data/mnist_image_label` 下,具有:
+ `mnist_train_jpg_60000` 用于训练的 60000 张手写数字图片
+ `mnist_train_jpg_60000.txt` 训练数据的标签文件
+ `mnist_test_jpg_10000` 用于测试的 10000 张手写数字图片
+ `mnist_test_jpg_10000.txt` 测试数据的标签文件
对于标签文件的格式说明:`手写数字图片名称 标签`
```
2028_7.jpg 7
3933_2.jpg 2
8299_5.jpg 5
7726_4.jpg 4
9874_2.jpg 2
```
我们的工作就是把之前的 `mnist.load_data()` 给替换掉,使用自己写的函数得到这部分数据集的 `x_train`、`y_train`、`x_test`、`y_test`
```
# img_dir: 图片存放的目录
# label_txt: 标签文件
def gen_dataset(img_dir, label_txt):
# 使用只读打开标签文件
with open(label_txt, 'r') as f:
x, y = [], []
for line in f.readlines():
img_name, label = line.split()
# 读入图片
img = Image.open(img_dir + img_name)
# 图片变为 8 位宽度的灰度值
img = np.array(img.convert('L'))
# 数据归一化
img = img / 255
x.append(img)
y.append(label)
x = np.array(x)
y = np.array(y).astype(np.int64)
return x, y
```
这样我们只需要传入图片所在目录以及标签所在文件路径就可以获取到我们需要的数据了。那么是否还可以改进呢?首先要思考的时候,是否有必要在每次获取数据时都构造一遍数据集呢?很明显是没必要的。其实在每次构造完数据集后,都可以将数据集保存起来,当下次要使用的时候先检查是否已有保存好的数据集,如若没有,再去构造数据集。
```
# 定义变量
train_dir = './data/mnist_image_label/mnist_train_jpg_60000/'
train_label_txt = './data/mnist_image_label/mnist_train_jpg_60000.txt'
test_dir = './data/mnist_image_label/mnist_test_jpg_10000/'
test_label_txt = './data/mnist_image_label/mnist_test_jpg_10000.txt'
def load_data(x_train_path, y_train_path, x_test_path, y_test_path):
# 文件保存的位置应封装在函数内部,因为使用者并不关心文件存取细节
x_train_save_path = './data/mnist_image_label/mnist_x_train.npy'
y_train_save_path = './data/mnist_image_label/mnist_y_train.npy'
x_test_save_path = './data/mnist_image_label/mnist_x_test.npy'
y_test_save_path = './data/mnist_image_label/mnist_y_test.npy'
if os.path.exists(x_train_save_path) and os.path.exists(y_train_save_path) and os.path.exists(x_test_save_path) and os.path.exists(y_test_save_path):
# 数据集格式已保存则直接加载
x_train_save = np.load(x_train_save_path)
y_train = np.load(y_train_save_path)
x_test_save = np.load(x_test_save_path)
y_test = np.load(y_test_save_path)
x_train = np.reshape(x_train_save, (len(x_train_save), 28, 28))
x_test = np.reshape(x_test_save, (len(x_test_save), 28, 28))
return (x_train, y_train), (x_test, y_test)
else:
# 生成数据集
x_train, y_train = gen_dataset(x_train_path, y_train_path)
x_test, y_test = gen_dataset(x_test_path, y_test_path)
# 保存数据集
x_train_save = np.reshape(x_train, (len(x_train), -1))
x_test_save = np.reshape(x_test, (len(x_test), -1))
np.save(x_train_save_path, x_train_save)
np.save(y_train_save_path, y_train)
np.save(x_test_save_path, x_test_save)
np.save(y_test_save_path, y_test)
return (x_train, y_train), (x_test, y_test)
# 读取数据集
(x_train, y_train), (x_test, y_test) = load_data(train_dir, train_label_txt, test_dir, test_label_txt)
x_train.shape
y_train.shape
x_test.shape
y_test.shape
```
### 2. 数据增强,扩充数据集
数据增强,也就是增强数据量。对于图片数据增强,tf.keras 有属于自己的 API:`tf.keras.preprocessing.image.ImageDataGenerator()`,有以下几个参数:
+ 缩放系数:rescale=所有数据将乘以提供的值
+ 随机旋转:rotation_range=随机旋转角度数范围
+ 宽度偏移:width_shift_range=随机宽度偏移量
+ 高度偏移:height_shift_range=随机高度偏移量
+ 水平翻转:horizontal_flip=是否水平随机翻转
+ 随机缩放:zoom_range=随机缩放的范围 [1-n,1+n]
设定完图片增强方式后,再调用 `.fit(x_train)` 来增强训练数据集。**值得注意的是**,本次用于图像数据增强的训练集的维度要求为 4 维,即(数量,宽度,高度,通道数),所以在使用该函数之前,还需要对训练数据进行 reshape。
```
# 从 (60000, 28, 28) reshape 为 (60000, 28, 28, 1) 即 60000 张 28 行 28 列单通道数据
x_train = x_train.reshape(x_train.shape[0], 28, 28, 1)
image_gen_train = tf.keras.preprocessing.image.ImageDataGenerator(
rescale=1. / 1., # 如为图像,分母为255时,可归至0~1
rotation_range=45, # 随机45度旋转
width_shift_range=.15, # 宽度偏移
height_shift_range=.15, # 高度偏移
horizontal_flip=False, # 水平翻转
zoom_range=0.5 # 将图像随机缩放阈量50%
)
image_gen_train.fit(x_train)
```
以上就很简单的完成了数据增强的功能,并且在之后训练模型时可以直接进行应用,但与之前稍有差别,但是别担心,下面会继续讲解的。
根据之前的 “六步法”,到这里已经算是完成了前两个步骤,其实整体思路是没什么变化的,只是我们使用了自己的数据集并对数据集进行了增强。接下来的第三步逐步搭建网络模型和第四步配置训练方法跟之前一样。
```
model = tf.keras.models.Sequential([
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(128, activation='relu'),
tf.keras.layers.Dense(10, activation='softmax')
])
model.compile(optimizer='adam',
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=False),
metrics=['sparse_categorical_accuracy'])
```
### 3. 断点续训,存取最优模型
使用断点续训,可以存取模型。
要想读取模型,可以直接使用 TensorFlow 给出的 `load_weight(路径文件名)` 函数,告知文件存在哪里,就可以直接读取已有模型参数了。保存的模型文件后缀为 ckpt,在生成 ckpt 文件的时候会同步生成索引(index)表,因此可以判断是否已存在索引表,就可以知道是否已保存过模型参数了。
```
# 先定义文件存放路径
checkpoint_save_path = "./checkpoint/mnist.ckpt"
# 在生成 ckpt 文件的时候会同步生成索引(index)表,因此可以判断是否已存在索引表,就可以知道是否已保存过模型参数了
if os.path.exists(checkpoint_save_path + '.index'):
# 读取模型参数
model.load_weights(checkpoint_save_path)
```
如果要保存模型参数,可以使用 TensorFlow 给出的回调函数 `tf.keras.callbacks.ModelCheckpoint`,直接保存训练出来的模型参数。该函数有如下几个参数:
+ filepath:路径文件名,
+ save_weights_only:是否只保留模型参数
+ save_best_only:是否只保留最优结果
+ monitor='val_loss:被监测的数据,可选值为val_acc或val_loss
```
cp_callback = tf.keras.callbacks.ModelCheckpoint(filepath=checkpoint_save_path,
save_weights_only=True,
save_best_only=True)
```
设定完断点后,接下来就是“六步法”中的第四步训练模型。这一步在使用 `model.fit()` 的时候,根据情况会与之前稍有不同。
如果使用了数据增强,那么就需要将原来的 `model.fit(x_train, y_train, batch_size=32, ...)` 更改为 `model.fit(image_gen_train.flow(x_train, y_train, batch_size=32))`,也就是使用数据增强的 `.flow()` 函数,使用数据和标签生成每批次 32 个的增强数据。
如果使用了断点续训,就需要在训练模型的 `model.fit()` 中再增加一个参数值 `callbacks=[cp_callback]`,即 增加断点续训的回调。
还可以用一个变量 history 来保存 `model.fit` 的返回值,用于之后获取训练集和测试集的 acc 和 loss 曲线。
```
history = model.fit(image_gen_train.flow(x_train, y_train, batch_size=32), epochs=5, validation_data=(x_test, y_test),
validation_freq=1, callbacks=[cp_callback])
```
模型训练完成之后,会在当前目录下生成一个 checkpoint 文件夹,里面存放的就是模型的参数。当我们再次运行时,程序会加载之前保存好的模型参数,并且会在之前保存的模型的基础之上继续提升准确率。
模型训练完成之后,就可以执行第六步打印并查看网络结构了。
```
model.summary()
```
### 4. 参数提取,把参数存入文本
利用参数提取,可以将模型参数存入到文本中。利用 `model.model.trainable_variables()` 来返回当前模型中所有可训练参数,要想看到这些可训练参数,可以直接用 `print` 函数直接打印出来,但是如果直接使用这种方式来查看的话,中间会有很多数据被省略号替换掉。
可以使用 `np.set_printoptions` 来设置 `print` 函数的打印效果,有以下几个常用参数:
+ precision:小数点后按四舍五入保留几位,默认 8 位
+ threshold:打印范围在 threshold + 1 以内的信息或者打印全部信息但中间部分会用省略号代替(默认值为1000)
```
np.set_printoptions(threshold=np.inf) # 设置threshold为无穷大,可以打印全部信息
#print(model.trainable_variables)
# 把参数信息保存在文件里
with open('./data/mnist_image_label/weights.txt', 'w') as file:
for v in model.trainable_variables:
file.write(str(v.name) + '\n')
file.write(str(v.shape) + '\n')
file.write(str(v.numpy()) + '\n')
```
### 5. 可视化 acc 和 loss 曲线,查看训练效果
这一部分主要是把准确率上升、损失函数下降的过程可视化出来。这里就需要用到之前在第四步模型训练是的返回值 history。这个变量内部记录了一下几个值的变化:
+ loss:训练集 loss
+ val_loss:测试集 loss
+ sparse_categorical_accuracy:训练集准确率
+ val_sparse_categorical_accuracy:测试集准确率
```
# 先获取各值
acc = history.history['sparse_categorical_accuracy']
val_acc = history.history['val_sparse_categorical_accuracy']
loss = history.history['loss']
val_loss = history.history['val_loss']
# 绘制准确率 acc 曲线
plt.plot(acc, label='Training Accuracy')
plt.plot(val_acc, label='Validation Accuracy')
plt.title('Training and Validation Accuracy')
plt.legend()
plt.show()
# 绘制损失函数 loss 的曲线
plt.plot(loss, label='Training Loss')
plt.plot(val_loss, label='Validation Loss')
plt.title('Training and Validation Loss')
plt.legend()
plt.show()
```
### 6. 应用模型对新数据进行预测
经过上面的步骤,“六步法”得到了很好的补充,使用 tf.keras 实现神经网络的方法也更加的完善。但是要想让模型可用,还需要编写一套应用程序,让模型可用于对新数据的预测。
对于手写数字识别模型,那么就需要让训练好的模型可以对新的手写的数字进行识别。可以使用 `model.predict()` 函数来对新的输入特征进行预测,该函数有以下几个常用参数:
+ x:输入数据,Numpy 数组(或者 Numpy 数组的列表,如果模型有多个输出)
+ batch_size:整数,由于 GPU 的特性,batch_size 最好选用 8,16,32,64……,如果未指定,默认为 32
+ verbose: 日志显示模式,0 或 1
+ steps: 声明预测结束之前的总步数(批次样本),默认值 None
+ 返回:预测的 Numpy 数组(或数组列表)
要实现一个应用程序进行预测,大致分为三个步骤:
+ 复现模型
+ 加载参数
+ 预测结果
#### 6.1 复现模型
```
model = tf.keras.models.Sequential([
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(128, activation='relu'),
tf.keras.layers.Dense(10, activation='softmax')
])
```
#### 6.2 加载参数
```
model_save_path = './checkpoint/mnist.ckpt'
model.load_weights(model_save_path)
```
#### 6.3 预测结果
在 `./data/digits` 目录中,存放着 10 个新的手写数字图片 `0.png`、`1.png` …… `9.png`
这里要注意的是,预测数据,与训练用的数据格式不一致。
+ 训练模型时用的图片是 28 行 28 列的灰度图,而输入的预测数据则是任意尺寸的图片,因此需要先 resize 成 28 行 28 列的标准尺寸,并转换为灰度图;
+ 训练模型时用的数据集是黑底白字灰度图,而输入的预测数据则是白底黑字,因此需要让输入数据的每个像素点等于 255 减去当前像素值(相当于颜色取反);
+ 此外,输入的预测数据还需要完成归一化,即对图片数据除以 255。
只有经过这两个步骤,才可以让从没见过的数字图片满足神经网络模型对输入风格的要求,这个过程叫做**预处理**。预处理的方法有很多,对于手写数字识别,还可以让输入图片变为只有黑色和白色的高对比度图片,这样可以在保留图片有用信息的同时,滤去背景噪声,图片更干净。
由于神经网络训练时,都是按照 batch 送入网络的,所以输入数据进入 predict 函数之前还需要在前面添加 1 个维度,从 28 行 28 列的二维数据,变为 1 个 28 行 28 列的三维数据。
```
# 随机从选一个手写数字图片
rand_idx = np.random.randint(0, 10)
img_path = './data/digits/{}.png'.format(rand_idx)
im = plt.imread(img_path)
plt.imshow(im)
print(img_path)
plt.show()
# 获取图片
img = Image.open(img_path)
# resize 图像大小为 28x28
img = img.resize((28, 28), Image.ANTIALIAS)
# 转换为灰度图
img_arr = np.array(img.convert('L'))
# 颜色取反,原来是白底黑字,转换为黑底白字
img_arr = 255 - img_arr
# 归一化
img_arr = img_arr / 255.0
# 将 img_arr 的维度从 [28, 28] 转换为 [1, 28, 28]
x_predict = img_arr[tf.newaxis, ...]
# 预测,并得到所有值的概率
result = model.predict(x_predict)
# 得到最大概率值,即预测结果
pred = tf.argmax(result, axis=1)
tf.print('predict:', pred)
```
| github_jupyter |
```
%pylab inline
```
# Последовательное обнаружение разладок временных рядов
1. Всюду в рассматриваемых задачах
имеется две гипотезы $\mathbb{H}_0$ и $\mathbb{H}_1$
(иногда они обозначаются $\mathbb{H}_{\infty}$ и
$\mathbb{H}_0$, соответственно), причем каждая
из гипотез делает явные предположения о распределении
или его параметрах.
\item Критерий Неймана-Пирсона предписывает
принимать гипотезу исходя из значения величины
$$
L_n(X_1, \ldots, X_n) = \frac{f_0(X_1, \ldots, X_n)}{f_{\infty}(X_1, \ldots, X_n)},
$$
называемой отношением правдоподобия.
А именно, пусть $\varphi(X_1, \ldots, X_n)$~--
рандомизированное решающее правило, значение
которого равно вероятности принять
гипотезу $\mathbb{H}_1$. Тогда найдутся такие
константы $\lambda_a$ и $h_a$, что
$$
\varphi(X_1, \ldots, X_n) =
\begin{cases}
1, & L_n(X_1, \ldots, X_n) > h_a, \\
\lambda_a, & L_n(X_1, \ldots, X_n) = h_a, \\
0, & L_n(X_1, \ldots, X_n) < h_a,
\end{cases}
$$
является наиболее мощным
(т.е. с наименьшей вероятностью пропуска цели
или ошибки 2 рода $\beta(\varphi)$) тестом
среди тестов, вероятность ложной
тревоги $\alpha(\varphi)$
(ошибки 1 рода) которых не выше $a$.
2. Разладкой процесса $X_t$
называется ситуация, в которой траектория
процесса генерируется двумя (или
в общем случае несколькими) независимыми
вероятностными мерами $\mathrm{P}_{\infty}$
и $\mathrm{P}_{0}$, причем наблюдения имеют структуру
$$
X_n =
\begin{cases}
X^{\infty}_n, & \text{если } 1 \leqslant n < \theta, \\
X^{0}_n, & \text{если } n \geqslant \theta,
\end{cases}
$$
где $X^{\infty}_t$ ---
процесс, соответствующий мере $\mathrm{P}_{\infty}$,
и $X^{0}_t$ ---
процесс, соответствующий мере $\mathrm{P}_{0}$.
Момент $\theta \in [0, \infty]$
называется моментом разладки, причем
ситуация $\theta = 0$ соответствует тому, что
с самого начала идут наблюдения от
_разлаженного_ процесса $X^{0}$,
а ситуация $\theta = \infty$ заключается в том,
что разладка не появляется никогда.
Таким образом, траектория процесса $X$ выглядит
следующим образом:
$$
\underbrace{X^{\infty}_1, X^{\infty}_2, \ldots,
X^{\infty}_{\theta - 1}}
_{\text{мера } \mathrm{P}^{\infty}},
\underbrace{X^{0}_{\theta}, X^{0}_{\theta + 1}, \ldots}
_{\text{мера } \mathrm{P}^{0}}
$$
3. **Статистика кумулятивных сумм.**
- Вводятся статистики $\gamma_t$
и $T_t$
$$
\gamma_n = \sup\limits_{\theta \geqslant 0} \frac{f_{\theta}(X_1, \ldots, X_n)}{f_{\infty}(X_1, \ldots, X_n)}
\qquad \mbox{и} \qquad T_n = \log \gamma_n
$$
- Если случайные величины
$X_1, \ldots, X_n$ независимы, то
$$
\gamma_n = \max\Big\{1, \max\limits_{1 \leqslant \theta \leqslant n} \prod\limits_{k=\theta}^{n}
\frac{f_{0}(X_k)}{f_{\infty}(X_k)} \Big\}
$$
$$
T_n = \max\Big\{0, \max\limits_{1 \leqslant \theta \leqslant n} \sum\limits_{k=\theta}^{n}
\log \frac{f_{0}(X_k)}{f_{\infty}(X_k)} \Big\} =
\max\Big\{0, \max\limits_{1 \leqslant \theta \leqslant n} \sum\limits_{k=\theta}^{n}
\zeta_k \Big\}
$$
- Статистика $T_n$ обладает свойством $T_n = \max (0, T_{n-1} + \zeta_n)$
и называется статистикой кумулятивных сумм (CUmulative SUMs, CUSUM).
- Момент остановки
$$
\tau_{\mathrm{CUSUM}} = \inf \{n \geqslant 0: T_n \geqslant B\},
$$
построенный по статистике кумулятивных сумм,
оптимален (т.\,е.~обладает наименьшей задержкой
в обнаружении разладки) в~классе
$$
\mathcal{M}_T = \{\tau :
{\textstyle \mathrm{E}_{\infty}} \tau \geqslant T\}
$$
тех моментов остановки, для которых среднее время
до~ложной тревоги не меньше $T$.
4. **Статистика Ширяева-Робертса.**
- Вводится статистика
$$
R_n = \sum\limits_{\theta = 1}^{n} \frac{f_{\theta}(X_1, \ldots, X_n)}{f_{\infty}(X_1, \ldots, X_n)}
$$
- Если случайные величины $X_1, \ldots, X_n$ независимы, то
$$
R_n = \sum\limits_{\theta = 1}^{n} \prod\limits_{k=\theta}^{n}
\frac{f_{0}(X_k)}{f_{\infty}(X_k)} =
\sum\limits_{\theta = 1}^{n} \prod\limits_{k=\theta}^{n}
l_k.
$$
- Статистика $R_n$ обладает свойством $R_n = (1 + R_{n-1}) l_k$ и~называется статистикой Ширяева-Робертса (Shiryaev-Roberts, SR).
- Момент остановки
$$
\tau_{\mathrm{SR}} = \inf \{n \geqslant 0: R_n \geqslant B\},
$$
построенный по статистике Ширяева-Робертса,
оптимален (т.\,е.~обладает наименьшей задержкой
в обнаружении разладки) в~классе
$$
\mathcal{M}_T = \{\tau :
{\textstyle \mathrm{E}_{\infty}} \tau \geqslant T\}
$$
тех моментов остановки, для которых среднее время
до~ложной тревоги не меньше $T$.
## Часть 1. Временные ряды с известными характеристиками
В этой части задания вы исследуете временные ряды с разладками, модель которых вам полностью известна.
Данные $X_1, \ldots, X_n$ порождены нормальным $\mathcal{N}(0, 1)$ распределением до момента появления разладки, и нормальным $\mathcal{N}(\mu, 1)$ распределением после момента появления разладки. Момент разладки $\theta = 100$ -- неизвестная величина.
```
theta = 100
X = np.hstack((
np.random.normal(loc=0, scale=1, size=theta),
np.random.normal(loc=2, scale=1, size=100))
)
plot(X)
```
Заготовка для класса, реализующего статистику для обнаружения разладки, представлена ниже. Вам необходимо расширять этот класс.
```
class Stat(object):
def __init__(self, threshold, direction="unknown", init_stat=0.0):
self._direction = str(direction)
self._threshold = float(threshold)
self._stat = float(init_stat)
self._alarm = self._stat / self._threshold
@property
def direction(self):
return self._direction
@property
def stat(self):
return self._stat
@property
def alarm(self):
return self._alarm
@property
def threshold(self):
return self._threshold
def update(self, **kwargs):
# Statistics may use any of the following kwargs:
# ts - timestamp for the value
# value - original value
# mean - current estimated mean
# std - current estimated std
# adjusted_value - usually (value - mean) / std
# Statistics call this after updating '_stat'
self._alarm = self._stat / self._threshold
```
**Задание 1.** Вычислите отношение правдоподобия для элемента выборки $X_i$. Реализуйте процедуру подсчета отношения правдоподобия для этого элемента выборки в предположении, что данные нормальны.
```
### your code here
from scipy.stats import norm
# Вариант 1 заключается в том, что вычисляются правдоподобия выборки с помощью функций,
# предоставляемых библиотекой scipy.stats
def normal_likelihood(value, mean_0, mean_8, std):
return np.log(norm.pdf(value, mean_0, std) /
norm.pdf(value, mean_8, std))
# Вариант 2 заключается в том, что можно упростить выражение,
# вычислив правдоподобие вручную -- оно выражается формулой, реализованной в коде
def normal_likelihood_explicit(value, mean_0, mean_8, std):
return (value - (mean_0 + mean_8) / 2.) * (mean_0 - mean_8) / std ** 2
```
**Задание 2.** Постройте статистику кумулятивных сумм для обнаружения разладки в среднем значении процесса $X_n$. В какой момент $\tau$ следует подавать сигнал тревоги об обнаружении разладки?
```
### your code here
class Cusum(Stat):
def __init__(self, mean_0, mean_8, std,
threshold, direction="unknown", init_stat=0.0):
self.mean_0 = mean_0
self.mean_8 = mean_8
self.std = std
super(Cusum, self).__init__(threshold, direction, init_stat)
def update(self, value):
zeta_k = normal_likelihood(value, self.mean_0, self.mean_8,
self.std)
self._stat = max(0, self._stat + zeta_k)
super(Cusum, self).update()
stat_trajectory = []
mean_0 = 2.
mean_8 = 0.
std = 1.
cusum = Cusum(mean_0, mean_8, std, 100.)
for x_k in X:
cusum.update(x_k)
stat_trajectory.append(cusum._alarm)
plot(stat_trajectory)
```
**Задание 3.** Проведите моделирование Монте-Карло для подсчета средней величины задержки в обнаружении разладки для заданных данных.
_Подсказка:_ сгенерируйте $N$ независимых выборок, для каждой из них вычислите значения статистики кумулятивных сумм и момент подачи тревоги $\tau$. Полученные _задержки_ $\tau - \theta$ усредните.
```
### your code here
mean_0 = 1.
mean_8 = 0.
std = 1.
delays = []
for i in xrange(1000):
theta = 100
X = np.hstack((
np.random.normal(loc=mean_8, scale=std, size=theta),
np.random.normal(loc=mean_0, scale=std, size=100))
)
cusum = Cusum(mean_0, mean_8, std, 30.)
for k, x_k in enumerate(X):
cusum.update(x_k)
if cusum._alarm >= 1 and k >= theta:
break
delays.append(k - theta)
_ = hist(delays, bins=40, normed=1, histtype='step')
```
**Задание 4.** Проведите моделирование Монте-Карло для подсчета зависимости средней величины задержки в обнаружении разладки, которая появляется в момент времени $\theta = 0$, от значения порога срабатывания $h$. Отобразите полученную зависимость.
_Подсказка:_ для нескольких значений порога срабатывания $h$ сгенерируйте $N$ независимых выборок из распределения _с разладкой_ $\mathrm{P}_{0}$, для каждой из них вычислите значения статистики кумулятивных сумм и момент подачи тревоги $\tau$. Полученные _задержки_ $\tau - \theta$ усредните.
```
### your code here
mean_0 = 1.
mean_8 = 0.
std = 1.
thresholds = np.arange(10., 100., 5.)
delays = []
delays_std = []
for h in thresholds:
delays_for_h = []
for i in xrange(1000):
theta = 100
X = np.hstack((
np.random.normal(loc=mean_8, scale=std, size=theta),
np.random.normal(loc=mean_0, scale=std, size=100))
)
cusum = Cusum(mean_0, mean_8, std, h)
for k, x_k in enumerate(X):
cusum.update(x_k)
if cusum._alarm >= 1 and k >= theta:
break
delays_for_h.append(k - theta)
delays.append(np.mean(delays_for_h))
delays_std.append(np.std(delays_for_h))
errorbar(thresholds, delays, yerr=delays_std)
xlabel('threshold $h$')
ylabel('delay $\\tau - \\theta$')
grid('on')
```
## Часть 2. Временные ряды с неизвестными характеристиками
В этой части задания вам необходимо разработать процедуру обнаружения разладки для данных, распределение которых вам неизвестно. Вам задан набор данных `seminar.train_data`. Процедура чтения реализована для вас.
```
from datasets import read_simple_dataset
from itertools import izip
with open('seminar.train_data') as f:
data, changepoint, targets = read_simple_dataset(f, keep_target=True)
timestamps, values, changepoint = [], [], []
for (ts, value), cp_indicator in izip(data[1], targets[1]):
timestamps.append(ts)
values.append(value)
changepoint.append(cp_indicator)
plot(timestamps, values)
plot(timestamps, changepoint, 'r', linewidth=3)
```
**Задание 1.** Модифицируйте процедуру кумулятивных сумм для случая, когда среднее значение временного ряда после разладки неизвестно.
Для решения этой задачи заметим, что у нас, вообще говоря, нет информации о среднем значении выданного временного ряда. Поэтому разумным будет оценить это значение с помощью какой-нибудь простой процедуры, например, экспоненциально взвешенного скользящего среднего (см. Задание 2).
Кумулятивные суммы при этом должны принимать в качестве параметра уже не точное значение среднего после разладки, а относительное отклонение в среднем значении такое, что $\mu_0 = \mu_{\infty} + \Delta$.
```
### your code here
class AdjustedCusum(Stat):
def __init__(self, mean_diff,
threshold, direction="unknown", init_stat=0.0):
self.mean_diff = mean_diff
super(AdjustedCusum, self).__init__(threshold, direction, init_stat)
def update(self, value):
zeta_k = normal_likelihood(value, mean_diff, 0., 1.)
self._stat = max(0, self._stat + zeta_k)
super(AdjustedCusum, self).update()
theta = 100
X = np.hstack((
np.random.normal(loc=0, scale=1, size=theta),
np.random.normal(loc=2, scale=1, size=100))
)
mean = 0.
var = 1.
alpha = 0.05
beta = 0.005
mean_diff = 1.0
stat_trajectory = []
cusum = AdjustedCusum(mean_diff, 30.)
for k, x_k in enumerate(X):
cusum.update(x_k)
stat_trajectory.append(cusum._stat)
plot(stat_trajectory)
```
**Задание 2.** Реализуйте процедуру оценивания среднего значения временного ряда с помощью скользящего среднего. Постройте статистику кумулятивных сумм для обнаружения разладки в среднем значении процесса $X_n$. В какой момент $\tau$ следует подавать сигнал тревоги об обнаружении разладки?
```
### your code here
```
Разумным будет следующий алгоритм обнаружения разладки временного ряда $X_1, \ldots, X_n, \ldots$:
- В каждый новый момент времени $t$ по вновь поступившему наблюдению $X_t$ производится вычисление стандартизованного и центрированного значения $Z_t = (X_t - \hat{\mu_{t-1}}) / \hat{\sigma_{t-1}}$, где $\hat{\mu_{t-1}}$ и $\hat{\sigma}^2_{t-1}$ - оценки среднего и дисперсии с помощью экспоненциального сглаживания, полученная по наблюдениям до момента времени $t-1$ включительно
- Переоценить среднее значение новыми данными согласно $\hat{\mu}_{t} = (1 - \alpha) \hat{\mu}_{t - 1} + \alpha X_t$
- Переоценить дисперсию новыми данными согласно $\hat{\sigma}^2_{t} = (1 - \beta) \hat{\sigma}^2_{t - 1} + \beta (X_t - \hat{\mu}_{t})^2$
```
# Тут реализована модификация статистики взвешенного экспоненциального среднего.
class MeanExpNoDataException(Exception):
pass
class MeanExp(object):
def __init__(self, new_value_weight, load_function=median):
self._load_function = load_function
self._new_value_weight = new_value_weight
self.load([])
@property
def value(self):
if self._weights_sum <= 1:
raise MeanExpNoDataException('self._weights_sum <= 1')
return self._values_sum / self._weights_sum
def update(self, new_value, **kwargs):
self._values_sum = (1 - self._new_value_weight) * self._values_sum + new_value
self._weights_sum = (1 - self._new_value_weight) * self._weights_sum + 1.0
def load(self, old_values):
if old_values:
old_values = [value for ts, value in old_values]
mean = float(self._load_function(old_values))
self._weights_sum = min(float(len(old_values)), 1.0 / self._new_value_weight)
self._values_sum = mean * self._weights_sum
else:
self._values_sum = 0.0
self._weights_sum = 0.0
theta = 100
X = np.hstack((
np.random.normal(loc=0, scale=1, size=theta),
np.random.normal(loc=2, scale=1, size=100))
)
mean = 0.
var = 1.
alpha = 0.05
beta = 0.005
mean_diff = 1.0
stat_trajectory, mean_values, var_values = [], [], []
mean_exp = MeanExp(new_value_weight=alpha)
var_exp = MeanExp(new_value_weight=beta)
cusum = AdjustedCusum(mean_diff, 30.)
for k, x_k in enumerate(X):
try:
mean_estimate = mean_exp.value
except MeanExpNoDataException:
mean_estimate = 0.
try:
var_estimate = var_exp.value
except MeanExpNoDataException:
var_estimate = 1.
adjusted_value = (x_k - mean_estimate) / np.sqrt(var_estimate)
cusum.update(adjusted_value)
mean_exp.update(x_k)
diff_value = (x_k - mean) ** 2
var_exp.update(diff_value)
stat_trajectory.append(cusum._stat)
mean_values.append(mean_estimate)
var_values.append(np.sqrt(var_estimate))
plot(X)
plot(np.array(mean_values), 'k')
plot(np.array(mean_values) + np.sqrt(var_values), 'k')
plot(np.array(mean_values) - np.sqrt(var_values), 'k')
plot( (X - np.array(mean_values)) / np.sqrt(var_values))
plot(stat_trajectory)
mean = 0.
var = 1.
alpha = 0.01
beta = 0.001
mean_diff = 1.0
stat_trajectory, mean_values, var_values = [], [], []
timestamps, values, changepoint = [], [], []
mean_exp = MeanExp(new_value_weight=alpha)
var_exp = MeanExp(new_value_weight=beta)
cusum = AdjustedCusum(mean_diff, 30.)
for (ts, x_k), cp_indicator in izip(data[1], targets[1]):
timestamps.append(ts)
values.append(x_k)
try:
mean_estimate = mean_exp.value
except MeanExpNoDataException:
mean_estimate = 0.
try:
var_estimate = var_exp.value
except MeanExpNoDataException:
var_estimate = 1.
adjusted_value = (x_k - mean_estimate) / np.sqrt(var_estimate)
cusum.update(adjusted_value)
mean_exp.update(x_k)
diff_value = (x_k - mean) ** 2
var_exp.update(diff_value)
stat_trajectory.append(cusum._stat)
mean_values.append(mean_estimate)
var_values.append(np.sqrt(var_estimate))
plot(values)
plot(np.array(mean_values), 'k')
plot(np.array(mean_values) + np.sqrt(var_values), 'k')
plot(np.array(mean_values) - np.sqrt(var_values), 'k')
plot( (values - np.array(mean_values)) / np.sqrt(var_values))
plot(stat_trajectory)
```
**Задание 3.** Постройте статистику Ширяева-Робертса для обнаружения разладки в дисперсии временного ряда.
Данные $X_1, \ldots, X_n$ порождены нормальным $\mathcal{N}(0, \sigma^2_{\infty})$ распределением до момента появления разладки, и нормальным $\mathcal{N}(0, \sigma^2_{0})$ распределением после момента появления разладки. Момент разладки $\theta$ -- неизвестная величина.
```
### your code here
```
Формально, необходимо снова подсчитать отношение правдоподобия и получить аналитическую формулу. Но мы воспользуемся тем фактом, что, если $x \sim N(\mu, \sigma^2)$, то $\mathrm{E} (x - \mu)^2 = \sigma^2$, и обычной формулой для правдоподобия (предполагая _грубо_, что $(x - \mu)^2$ - нормальная величина, у которой надо искать разладку в среднем значении).
```
class AdjustedShiryaevRoberts(Stat):
def __init__(self, mean_diff, threshold, max_stat=float("+inf"), init_stat=0.0):
super(AdjustedShiryaevRoberts, self).__init__(threshold,
direction="up",
init_stat=init_stat)
self._mean_diff = mean_diff
self._max_stat = max_stat
def update(self, adjusted_value, **kwargs):
likelihood = np.exp(self._mean_diff * (adjusted_value - self._mean_diff / 2.))
self._stat = min(self._max_stat, (1. + self._stat) * likelihood)
Stat.update(self)
theta = 100
X = np.hstack((
np.random.normal(loc=0, scale=1, size=theta),
np.random.normal(loc=0, scale=3, size=100))
)
plot(X)
stat_trajectory = []
sigma_0 = 3.
sigma_8 = 1.
mean = 0.
sigma_diff = sigma_0 ** 2 - sigma_8 ** 2
sr = AdjustedShiryaevRoberts(sigma_diff, 1000., max_stat=1e6)
for x_k in X:
diff_value = (x_k - mean) ** 2
sr.update(diff_value)
stat_trajectory.append(sr._stat)
plot(stat_trajectory)
```
**Задание 4.** Модифицируйте процедуру обнаружения разладки с помощью статистики Ширяева-Робертса для обнаружения разладки в дисперсии временного ряда, заданного данными из файлов. Подсказка: постройте вначале процедуру оценивания значения дисперсии до момента разладки, а затем воспользуйтесь идеей из Задания 1.
```
with open('seminar_var.train_data') as f:
data_var, changepoint_var, targets_var = read_simple_dataset(f, keep_target=True)
timestamps, values, changepoint = [], [], []
for (ts, value), cp_indicator in izip(data_var[0], targets_var[0]):
timestamps.append(ts)
values.append(value)
changepoint.append(cp_indicator)
plot(timestamps, values)
plot(timestamps, changepoint, 'r', linewidth=3)
### your code here
```
Идея решения заключается в следующем:
- Разладка в дисперсии означает высокое значение разности оценки $\hat{\sigma}^2_{t}$ и значения $(X_t - \hat{\mu}_{t})^2$, поэтому именно эту разность $\hat{\sigma}^2_{t} - (X_t - \hat{\mu}_{t})^2$ мы будем подавать на вход процедуре Ширяева-Робертса (настроенной на поиск разладки "на увеличение")
Остальные шаги алгоритма полностью соответствуют случаю неизвестных моментов временного ряда:
- Переоценить среднее значение новыми данными согласно $\hat{\mu}_{t} = (1 - \alpha) \hat{\mu}_{t - 1} + \alpha X_t$
- Переоценить дисперсию новыми данными согласно $\hat{\sigma}^2_{t} = (1 - \beta) \hat{\sigma}^2_{t - 1} + \beta (X_t - \hat{\mu}_{t})^2$
```
alpha = 0.01
beta = 0.05
sigma_diff = 2.0
stat_trajectory, mean_values, var_values, diff_values = [], [], [], []
timestamps, values, changepoint = [], [], []
mean_exp = MeanExp(new_value_weight=alpha)
var_exp = MeanExp(new_value_weight=beta)
sr = AdjustedShiryaevRoberts(sigma_diff, 1000., max_stat=1e9)
for (ts, x_k), cp_indicator in izip(data_var[2], targets_var[2]):
timestamps.append(ts)
values.append(x_k)
try:
mean_estimate = mean_exp.value
except MeanExpNoDataException:
mean_estimate = 0.
try:
var_estimate = var_exp.value
except MeanExpNoDataException:
var_estimate = 1.
predicted_diff_value = (x_k - mean_estimate) ** 2
predicted_diff_mean = var_estimate
sr.update(predicted_diff_value - predicted_diff_mean)
diff_values.append(predicted_diff_value - predicted_diff_mean)
mean_exp.update(x_k)
diff_value = (x_k - mean_estimate) ** 2
var_exp.update(diff_value)
stat_trajectory.append(sr._stat)
mean_values.append(mean_estimate)
var_values.append(np.sqrt(var_estimate))
figure(figsize=(12, 6))
plot(values)
plot(np.array(mean_values), 'k')
plot(np.array(mean_values) + np.sqrt(var_values), 'k')
plot(np.array(mean_values) - np.sqrt(var_values), 'k')
figure(figsize=(12, 6))
plot(diff_values)
figure(figsize=(12, 6))
semilogy(stat_trajectory)
grid('on')
```
| github_jupyter |
#### Notes
* Use the citation network to get an idea of what papers you might want to add, especially since that'll give you a good idea of what's already linking the fields
* What am I going to DO with this dataset? Specifically problem attributes and strategies.
* Mathematica visuals will be great
* What other kinds of statistical analysis would be useful?
* It's not useful unless I can get other people to use it, so how am I going to do that?
* Pester people once I have preliminary results about how I can make this useful to them?
* Good Github repository and readme
* Edit Wikipedia articles?
* Citation tools: use a parser (anystyle or freecite) and then query web of science, scopus, or scifinder?
* Project defense: approved with a couple hrs of changes, vs like a week or two?
#### Code-Related Next Steps
* Put it in a github repository
* Look at connectivity on Wikipedia between network topics and graph topics
#### Notes on Crossref item properties:
* 'container-title' is the journal title, which I don't currently care about
```
# Not currently being used
import csv
from matplotlib import pyplot as plt
# For reading in bib.txt files
import re
import glob
# For getting the spreadsheet
import urllib
import gspread
import requests
# For using the .py database classes
import importlib
import paper
import database
import subject_assignment as subjects
import numpy as np
from collections import defaultdict
import time
from bs4 import BeautifulSoup, Comment, NavigableString
importlib.reload(database)
importlib.reload(paper)
db = database.Database()
db.initialize_parents("parents.txt")
#db.sync_parents()
db.rewrite_parents("parents.txt")
db.initialize_children()
#db.write_bads(<filename>)
#db.update_bads(<filename>)
db.rewrite_children()
db.update_bads("fixed_lastcallfornewguys.txt")
db.print_stats()
db.to_csv("final_database.csv")
importlib.reload(subjects)
subjects.clear_subjects(db)
journals = subjects.initialize_journals(db, verbose=True)
print()
topics = subjects.get_wordlists()
journals, journals_by_topic = subjects.tag_journals(db, journals, topics,
verbose=True)
print()
papers_by_topic = subjects.tag_papers(db, journals, topics, verbose=True)
with open('journal_titles.txt','w',errors='backslashreplace') as f:
for journal in journals:
is_1 = subjects.check_words(journal, topics['Biology'])
is_2 = subjects.check_words(journal, topics['Mathematics'])
is_3 = subjects.check_words(journal, topics['Computer Science'])
if is_1 == False and is_2 == False and is_3 == False:
f.write(str(journal)+'\n')
with open('all_journal_titles.txt','w',errors='backslashreplace') as f:
for journal in journals:
f.write(str(journal)+'\n')
num_with_word = dict()
for topic in topics:
topic_word = dict()
for word in topics[topic]:
is_word = set()
for p_hash, p in db.all_papers.items():
p_hasword = False
if p.container_title:
for thing in p.container_title:
if subjects.check_words(thing, [word]):
p_hasword = True
if p_hasword:
is_word.add(p_hash)
topic_word[word] = len(is_word.intersection(papers_by_topic[topic]))
num_with_word[topic] = topic_word
for topic in topics:
print(topic + ":")
for word in topics[topic]:
print("\t", num_with_word[topic][word], word)
importlib.reload(subjects)
subjects.clear_subjects(db)
journals = subjects.initialize_journals(db, verbose=True)
print()
topics = subjects.get_wordlists()
for topic in topics:
print(len(topics[topic]))
journals, journals_by_topic = subjects.tag_journals(db, journals, topics,
verbose=True)
is_big3 = set()
big3 = {'Algorithm','SIAM'}
for p_hash, p in db.all_papers.items():
p_inbig3 = False
if p.container_title:
for thing in p.container_title:
if subjects.check_words(thing, big3):
p_inbig3 = True
if p_inbig3:
is_big3.add(p_hash)
print()
papers_by_topic = subjects.tag_papers(db, journals, topics, verbose=True)
print()
print(len(is_big3))
for topic, topicpapers in papers_by_topic.items():
print(topic, len(is_big3.intersection(topicpapers)))
importlib.reload(subjects)
subjects.check_intersections(db, topics, papers_by_topic)
importlib.reload(subjects)
hashes = subjects.get_to_read_hashes()
outfile = "./ThesisClass/thesis_bibliography.txt"
for p_hash in hashes:
try:
p = db.all_papers[p_hash]
bib = requests.request('GET', 'http://dx.doi.org/' + p.DOI,
headers={'Accept':'application/x-bibtex'}, timeout=100)
if bib.text.find("<") < 0:
pass
else:
print("FAILURE:", p_hash)
except KeyError:
print("FAILED:", p_hash)
importlib.reload(subjects)
subjects.citation_network(db, 'subject_tagged_network.gml')
# SUBJECT IS A USELESS PIECE OF CROSSREF INFO
i = 0
with_subject = 0
without_subject = 0
subjects = set()
for p_hash, p in db.all_papers.items():
i += 1
if i > 10000:
break
"""
try:
print(p.item['subject'])
except KeyError:
pass
"""
if p.subject:
print("\n",p.title, p.subject)
subjects.update(set(p.subject))
with_subject += 1
else:
without_subject += 1
print(with_subject)
print(without_subject)
print(len(subjects))
import numpy as np
family = np.array([0.03,0.06,0.03,0.1,0.1,0.14,0.14,0.4])
comb = np.array([0.16,0.08,0.22,0.05,0.08,0.22,0.05,0.14])
trail = np.array([0.13,0.05,0.16,0.03,0.22,0.16,0.03,0.22])
helicopter = np.array([0.25,0.08,0.25,0.08,0.12,0.09,0.05,0.08])
u = np.array([100.0,70,55,50,20,15,10,0])
for action in [family, comb, trail, helicopter]:
print(np.dot(u,action))
A = np.arange(12).reshape(4,3)
shifted = A - np.max(A,axis=1).reshape(4,1)
indices = np.array([0,1,2,1])
print(A)
print(np.choose(indices, A.T))
print()
print(shifted)
print()
import itertools
import numpy as np
payoffs = {'CC':(3,3),'CD':(1,5),'DC':(5,1),'DD':(2,2)}
beta = 0.9 # discount factor
desired_reward = np.array([24.0,31.0])
for n in [2,3]:
scale = 1.0/(1.0-beta**n)
# Explore the 2^n possible strategies for a sequence of length n
for strategy in itertools.product(('CC','CD','DC','DD'), repeat=n):
# Construct the discounted payoffs for the infinite discounted strategy
payoff = np.zeros((2,n))
for i in range(n):
payoff[:,i] = payoffs[strategy[i]]
discount = np.array([beta**i for i in range(n)])
# This might make the strategy backwards with respect to player 1 vs.
# player 2, but it's simpler and an easy fix.
reward = np.sort(scale*np.dot(payoff, discount))
# Construct the enforceable and ">[24,31]" constraint
maximin_payoff = 2.0*scale*np.sum(discount)*np.ones(2)
is_enforceable = np.prod((reward >= maximin_payoff))
satisfies_constraint = np.prod((reward >= desired_reward))
# Report success if applicable
if is_enforceable and satisfies_constraint:
print("Strategy", strategy, ":", "Payoffs", np.round(reward,2))
```
| github_jupyter |
```
# Copyright 2019 Google LLC
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Purchase Prediction with AutoML Tables
<table align="left">
<td>
<a href="https://cloud.google.com/ml-engine/docs/tensorflow/getting-started-keras">
<img src="https://cloud.google.com/_static/images/cloud/icons/favicons/onecloud/super_cloud.png"
alt="Google Cloud logo" width="32px"> Read on cloud.google.com
</a>
</td>
<td>
<a href="">
<img src="https://cloud.google.com/ml-engine/images/colab-logo-32px.png" alt="Colab logo"> Run in Colab
</a>
</td>
<td>
<a href="">
<img src="https://cloud.google.com/ml-engine/images/github-logo-32px.png" alt="GitHub logo">
View on GitHub
</a>
</td>
</table>
## Overview
One of the most common use cases in Marketing is to predict the likelihood of conversion. Conversion could be defined by the marketer as taking a certain action like making a purchase, signing up for a free trial, subscribing to a newsletter, etc. Knowing the likelihood that a marketing lead or prospect will ‘convert’ can enable the marketer to target the lead with the right marketing campaign. This could take the form of remarketing, targeted email campaigns, online offers or other treatments.
Here we demonstrate how you can use Bigquery and AutoML Tables to build a supervised binary classification model for purchase prediction.
### Dataset
The model uses a real dataset from the [Google Merchandise store](https://www.googlemerchandisestore.com) consisting of Google Analytics web sessions.
The goal here is to predict the likelihood of a web visitor visiting the online Google Merchandise Store making a purchase on the website during that Google Analytics session. Past web interactions of the user on the store website in addition to information like browser details and geography are used to make this prediction.
This is framed as a binary classification model, to label a user during a session as either true (makes a purchase) or false (does not make a purchase). Dataset Details The dataset consists of a set of tables corresponding to Google Analytics sessions being tracked on the Google Merchandise Store. Each table is a single day of GA sessions. More details around the schema can be seen here.
You can access the data on BigQuery [here](https://support.google.com/analytics/answer/3437719?hl=en&ref_topic=3416089).
## Instructions
Here is a list of things to do with AutoML Tables:
* Set up your local development environment (optional)
* Set Project ID and Compute Region
* Authenticate your GCP account
* Import Python API SDK and create a Client instance,
* Create a dataset instance and import the data.
* Create a model instance and train the model.
* Evaluating the trained model.
* Deploy the model on the cloud for online predictions.
* Make online predictions.
* Undeploy the model
# 1. Before you begin
## Project setup
Follow the [AutoML Tables documentation](https://cloud.google.com/automl-tables/docs/) to:
* Create a Google Cloud Platform (GCP) project, replace "your-project" with your GCP project ID and set local development environment.
* Enable billing.
* Enable AutoML API.
* Enter your project ID in the cell below. Then run the cell to make sure the
**If you are using Colab or AI Platform Notebooks**, your environment already meets
all the requirements to run this notebook. You can skip this step from the AutoML Tables documentation
Cloud SDK uses the right project for all the commands in this notebook.
**Note**: Jupyter runs lines prefixed with `!` as shell commands, and it interpolates Python variables prefixed with `$` into these commands
```
PROJECT_ID = "<your-project>" # @param {type:"string"}
COMPUTE_REGION = "us-central1" # Currently only supported region.
! gcloud config set project $PROJECT_ID
```
---
This section runs initialization and authentication. It creates an authenticated session which is required for running any of the following sections.
## Authenticate your GCP account
**If you are using AI Platform Notebooks**, your environment is already
authenticated. Skip this step.
**If you are using Colab**, run the cell below and follow the instructions
when prompted to authenticate your account via oAuth.
**Otherwise**, follow these steps:
1. In the GCP Console, go to the [**Create service account key**
page](https://console.cloud.google.com/apis/credentials/serviceaccountkey).
2. From the **Service account** drop-down list, select **New service account**.
3. In the **Service account name** field, enter a name.
4. From the **Role** drop-down list, select
**AutoML > AutoML Admin** and
**Storage > Storage Object Admin**.
5. Click *Create*. A JSON file that contains your key downloads to your
local environment.
6. Enter the path to your service account key as the
`GOOGLE_APPLICATION_CREDENTIALS` variable in the cell below and run the cell.
```
import sys
# If you are running this notebook in Colab, run this cell and follow the
# instructions to authenticate your GCP account. This provides access to your
# Cloud Storage bucket and lets you submit training jobs and prediction
# requests.
if 'google.colab' in sys.modules:
from google.colab import files
keyfile_upload = files.upload()
keyfile = list(keyfile_upload.keys())[0]
%env GOOGLE_APPLICATION_CREDENTIALS $keyfile
# If you are running this notebook locally, replace the string below with the
# path to your service account key and run this cell to authenticate your GCP
# account.
else:
%env GOOGLE_APPLICATION_CREDENTIALS /path/to/service_account.json
```
## Install the client library
Run the following cell.
```
%pip install --quiet google-cloud-automl
```
### Import libraries and define constants
First, import Python libraries required for training,
The code example below demonstrates importing the AutoML Python API module into a python script.
```
# AutoML library
from google.cloud import automl_v1beta1 as automl
import google.cloud.automl_v1beta1.proto.data_types_pb2 as data_types
import matplotlib.pyplot as plt
```
Restart the kernel to allow automl_v1beta1 to be imported for Jupyter Notebooks.
```
from IPython.core.display import HTML
HTML("<script>Jupyter.notebook.kernel.restart()</script>")
```
### Create API Client to AutoML Service*
**If you are using AI Platform Notebooks**, or *Colab* environment is already
authenticated using GOOGLE_APPLICATION_CREDENTIALS. Run this step.
```
client = automl.TablesClient(project=PROJECT_ID, region=COMPUTE_REGION)
```
**If you are using Colab or Jupyter**, and you have defined a service account
follow the following steps to create the AutoML client
You can see a different way to create the API Clients using service account.
```
# from google.oauth2 import service_account
# credentials = service_account.Credentials.from_service_account_file('/path/to/service_account.json')
# client = automl.TablesClient(project=PROJECT_ID, region=COMPUTE_REGION, credentials=credentials)
```
---
## Storage setup
You also need to upload your data into [Google Cloud Storage](https://cloud.google.com/storage/) (GCS) or [BigQuery](https://cloud.google.com/bigquery/).
For example, to use GCS as your data source:
* [Create a GCS bucket](https://cloud.google.com/storage/docs/creating-buckets).
* Upload the training and batch prediction files.
---
# 3. Import, clean, transform and perform feature engineering on the training Data
### Create dataset in AutoML Tables
Select a dataset display name and pass your table source information to create a new dataset.
```
#@title Create dataset { vertical-output: true, output-height: 200 }
dataset_display_name = 'colab_trial1' #@param {type: 'string'}
dataset = client.create_dataset(dataset_display_name)
dataset
```
Create a bucket to store the training data in
```
#@title Create bucket to store data in { vertical-output: true, output-height: 200 }
bucket_name = '<your-bucket>' #@param {type: 'string'}
```
### Import Dependencies
```
!sudo pip install google-cloud-bigquery google-cloud-storage pandas pandas-gbq gcsfs oauth2client
import datetime
import pandas as pd
import gcsfs
from google.cloud import bigquery
from google.cloud import storage
client_bq = bigquery.Client(location='US', project=PROJECT_ID)
```
### Transformation and Feature Engineering Functions
The data cleaning and transformation step was by far the most involved. It includes a few sections that create an AutoML tables dataset, pull the Google merchandise store data from BigQuery, transform the data, and save it multiple times to csv files in google cloud storage.
The dataset that is made viewable in the AutoML Tables UI. It will eventually hold the training data after that training data is cleaned and transformed.
This dataset has only around 1% of its values with a positive label value of True i.e. cases when a transaction was made. This is a class imbalance problem. There are several ways to handle class imbalance. We chose to oversample the positive class by random over sampling. This resulted in an artificial increase in the sessions with the positive label of true transaction value.
There were also many columns with either all missing or all constant values. These columns would not add any signal to our model, so we dropped them.
There were also columns with NaN rather than 0 values. For instance, rather than having a count of 0, a column might have a null value. So we added code to change some of these null values to 0, specifically in our target column, in which null values were not allowed by AutoML Tables. However, AutoML Tables can handle null values for the features.
#### Feature Engineering
The dataset had rich information on customer location and behavior; however, it can be improved by performing feature engineering. Moreover, there was a concern about data leakage. The decision to do feature engineering, therefore, had two contributing motivations: remove data leakage without too much loss of useful data, and to improve the signal in our data.
##### Weekdays
The date seemed like a useful piece of information to include, as it could capture seasonal effects. Unfortunately, we only had one year of data, so seasonality on an annual scale would be difficult (read impossible) to incorporate. Fortunately, we could try and detect seasonal effects on a micro, with perhaps equally informative results. We ended up creating a new column of weekdays out of dates, to denote which day of the week the session was held on. This new feature turned out to have some useful predictive power, when added as a variable into our model.
##### Data Leakage
The marginal gain from adding a weekday feature, was overshadowed by the concern of data leakage in our training data. In the initial naive models we trained, we got outstanding results. So outstanding that we knew that something must be going on. As it turned out, quite a few features functioned as proxies for the feature we were trying to predict: meaning some of the features we conditioned on to build the model had an almost 1:1 correlation with the target feature. Intuitively, this made sense.
One feature that exhibited this behavior was the number of page views a customer made during a session. By conditioning on page views in a session, we could very reliably predict which customer sessions a purchase would be made in. At first this seems like the golden ticket, we can reliably predict whether or not a purchase is made! The catch: the full page view information can only be collected at the end of the session, by which point we would also have whether or not a transaction was made. Seen from this perspective, collecting page views at the same time as collecting the transaction information would make it pointless to predict the transaction information using the page views information, as we would already have both. One solution was to drop page views as a feature entirely. This would safely stop the data leakage, but we would lose some critically useful information. Another solution, (the one we ended up going with), was to track the page view information of all previous sessions for a given customer, and use it to inform the current session. This way, we could use the page view information, but only the information that we would have before the session even began. So we created a new column called previous_views, and populated it with the total count of all previous page views made by the customer in all previous sessions. We then deleted the page views feature, to stop the data leakage.
Our rationale for this change can be boiled down to the concise heuristic: only use the information that is available to us on the first click of the session. Applying this reasoning, we performed similar data engineering on other features which we found to be proxies for the label feature. We also refined our objective in the process: For a visit to the Google Merchandise store, what is the probability that a customer will make a purchase, and can we calculate this probability the moment the customer arrives? By clarifying the question, we both made the result more powerful/useful, and eliminated the data leakage that threatened to make the predictive power trivial.
```
def balanceTable(table):
#class count
count_class_false, count_class_true = table.totalTransactionRevenue.value_counts()
#divide by class
table_class_false = table[table["totalTransactionRevenue"] == False]
table_class_true = table[table["totalTransactionRevenue"] == True]
#random over-sampling
table_class_true_over = table_class_true.sample(count_class_false, replace = True)
table_test_over = pd.concat([table_class_false, table_class_true_over])
return table_test_over
def partitionTable(table, dt=20170500):
#the automl tables model could be training on future data and implicitly learning about past data in the testing
#dataset, this would cause data leakage. To prevent this, we are training only with the first 9 months of data (table1)
#and doing validation with the last three months of data (table2).
table1 = table[table["date"] <= dt]
table2 = table[table["date"] > dt]
return table1, table2
def N_updatePrevCount(table, new_column, old_column):
table = table.fillna(0)
table[new_column] = 1
table.sort_values(by=['fullVisitorId','date'])
table[new_column] = table.groupby(['fullVisitorId'])[old_column].apply(lambda x: x.cumsum())
table.drop([old_column], axis = 1, inplace = True)
return table
def N_updateDate(table):
table['weekday'] = 1
table['date'] = pd.to_datetime(table['date'].astype(str), format = '%Y%m%d')
table['weekday'] = table['date'].dt.dayofweek
return table
def change_transaction_values(table):
table['totalTransactionRevenue'] = table['totalTransactionRevenue'].fillna(0)
table['totalTransactionRevenue'] = table['totalTransactionRevenue'].apply(lambda x: x!=0)
return table
def saveTable(table, csv_file_name, bucket_name):
table.to_csv(csv_file_name, index = False)
storage_client = storage.Client()
bucket = storage_client.get_bucket(bucket_name)
blob = bucket.blob(csv_file_name)
blob.upload_from_filename(filename = csv_file_name)
```
### Import training data
You also have the option of just downloading the file, FULL.csv, [here](https://storage.cloud.google.com/cloud-ml-data/automl-tables/notebooks/trial_for_c4m/FULL.csv), instead of running the code below. Just be sure to move the file into the google cloud storage bucket you specified above.
```
#@title Input name of file to save data to { vertical-output: true, output-height: 200 }
query = """
SELECT
date,
device,
geoNetwork,
totals,
trafficSource,
fullVisitorId
FROM
`bigquery-public-data.google_analytics_sample.ga_sessions_*`
WHERE
_TABLE_SUFFIX BETWEEN FORMAT_DATE('%Y%m%d',DATE_SUB('2017-08-01', INTERVAL 366 DAY)) AND
FORMAT_DATE('%Y%m%d',DATE_SUB('2017-08-01', INTERVAL 1 DAY))
"""
df = client_bq.query(query).to_dataframe()
print(df.iloc[:3])
path_to_data_pre_transformation = "FULL.csv" #@param {type: 'string'}
saveTable(df, path_to_data_pre_transformation, bucket_name)
```
### Unnest the Data
```
query = """
SELECT
date,
device,
geoNetwork,
totals,
trafficSource,
fullVisitorId
FROM `bigquery-public-data.google_analytics_sample.ga_sessions_*`
WHERE
_TABLE_SUFFIX BETWEEN FORMAT_DATE('%Y%m%d',DATE_SUB('2017-08-01', INTERVAL 366 DAY)) AND
FORMAT_DATE('%Y%m%d',DATE_SUB('2017-08-01', INTERVAL 1 DAY))
"""
df = client_bq.query(query).to_dataframe()
#some transformations on the basic dataset
#@title Input the name of file to hold the unnested data to { vertical-output: true, output-height: 200 }
unnested_file_name = "FULL_unnested.csv" #@param {type: 'string'}
```
You also have the option of just downloading the file, FULL_unnested.csv, [here](https://storage.cloud.google.com/cloud-ml-data/automl-tables/notebooks/trial_for_c4m/FULL_unnested.csv), instead of running the code below. Just be sure to move the file into the google cloud storage bucket you specified above.
```
table = pd.read_csv("gs://"+bucket_name+"/"+unnested_file_name, low_memory=False)
column_names = ['device', 'geoNetwork','totals', 'trafficSource']
for name in column_names:
print(name)
table[name] = table[name].apply(lambda i: dict(eval(i)))
temp = table[name].apply(pd.Series)
table = pd.concat([table, temp], axis=1).drop(name, axis=1)
#need to drop a column
table.drop(['adwordsClickInfo'], axis = 1, inplace = True)
saveTable(table, unnested_file_name, bucket_name)
```
### Run the Transformations
```
table = pd.read_csv("gs://"+bucket_name+"/"+unnested_file_name, low_memory=False)
consts = ['transactionRevenue', 'transactions', 'adContent', 'browserSize', 'campaignCode',
'cityId', 'flashVersion', 'javaEnabled', 'language', 'latitude', 'longitude', 'mobileDeviceBranding',
'mobileDeviceInfo', 'mobileDeviceMarketingName','mobileDeviceModel','mobileInputSelector', 'networkLocation',
'operatingSystemVersion', 'screenColors', 'screenResolution', 'screenviews', 'sessionQualityDim', 'timeOnScreen',
'visits', 'uniqueScreenviews', 'browserVersion','referralPath','fullVisitorId', 'date']
table = N_updatePrevCount(table, 'previous_views', 'pageviews')
table = N_updatePrevCount(table, 'previous_hits', 'hits')
table = N_updatePrevCount(table, 'previous_timeOnSite', 'timeOnSite')
table = N_updatePrevCount(table, 'previous_Bounces', 'bounces')
table = change_transaction_values(table)
table1, table2 = partitionTable(table)
table1 = N_updateDate(table1)
table2 = N_updateDate(table2)
#validation_unnested_FULL.csv = the last 3 months of data
table1.drop(consts, axis = 1, inplace = True)
table2.drop(consts, axis = 1, inplace = True)
saveTable(table2,'validation_unnested_FULL.csv', bucket_name)
table1 = balanceTable(table1)
#training_unnested_FULL.csv = the first 9 months of data
saveTable(table1, 'training_unnested_balanced_FULL.csv', bucket_name)
#@title ... take the data source from GCS { vertical-output: true }
dataset_gcs_input_uris = ['gs://{}/training_unnested_balanced_FULL.csv'.format(bucket_name),] #@param
import_data_operation = client.import_data(
dataset=dataset,
gcs_input_uris=dataset_gcs_input_uris
)
print('Dataset import operation: {}'.format(import_data_operation))
# Synchronous check of operation status. Wait until import is done.
import_data_operation.result()
dataset = client.get_dataset(dataset_name=dataset.name)
dataset
```
---
# 4. Update dataset: assign a label column and enable nullable columns
AutoML Tables automatically detects your data column type. Depending on the type of your label column, AutoML Tables chooses to run a classification or regression model. If your label column contains only numerical values, but they represent categories, change your label column type to categorical by updating your schema.
```
# List table specs
list_table_specs_response = client.list_table_specs(dataset=dataset)
table_specs = [s for s in list_table_specs_response]
# List column specs
list_column_specs_response = client.list_column_specs(dataset=dataset)
column_specs = {s.display_name: s for s in list_column_specs_response}
# Print Features and data_type:
features = [(key, data_types.TypeCode.Name(value.data_type.type_code)) for key, value in column_specs.items()]
print('Feature list:\n')
for feature in features:
print(feature[0],':', feature[1])
# Table schema pie chart.
type_counts = {}
for column_spec in column_specs.values():
type_name = data_types.TypeCode.Name(column_spec.data_type.type_code)
type_counts[type_name] = type_counts.get(type_name, 0) + 1
plt.pie(x=type_counts.values(), labels=type_counts.keys(), autopct='%1.1f%%')
plt.axis('equal')
plt.show()
#@title Update a column: set to not nullable { vertical-output: true }
update_column_response = client.update_column_spec(
dataset=dataset,
column_spec_display_name='totalTransactionRevenue',
nullable=False,
)
update_column_response
```
**Tip:** You can use kwarg `type_code='CATEGORY'` in the preceding `update_column_spec(..)` call to convert the column data type from `FLOAT64` `to `CATEGORY`.
### Update dataset: assign a target column
```
#@title Update dataset { vertical-output: true }
update_dataset_response = client.set_target_column(
dataset=dataset,
column_spec_display_name='totalTransactionRevenue',
)
update_dataset_response
```
# 5. Creating a model
### Train a model
To create the datasets for training, testing and validation, we first had to consider what kind of data we were dealing with. The data we had keeps track of all customer sessions with the Google Merchandise store over a year. AutoML tables does its own training and testing, and delivers a quite nice UI to view the results in. For the training and testing dataset then, we simply used the over sampled, balanced dataset created by the transformations described above. But we first partitioned the dataset to include the first 9 months in one table and the last 3 in another. This allowed us to train and test with an entirely different dataset that what we used to validate.
Moreover, we held off on oversampling for the validation dataset, to not bias the data that we would ultimately use to judge the success of our model.
The decision to divide the sessions along time was made to avoid the model training on future data to predict past data. (This can be avoided with a datetime variable in the dataset and by toggling a button in the UI)
Training the model may take one hour or more. The following cell keeps running until the training is done. If your Colab times out, use `client.list_models()` to check whether your model has been created. Then use model name to continue to the next steps. Run the following command to retrieve your model. Replace `model_name` with its actual value.
model = client.get_model(model_name=model_name)
Note that we trained on the first 9 months of data and we validate using the last 3.
```
#@title Create model { vertical-output: true }
#this will create a model that can be access through the auto ml tables colab
model_display_name = 'trial_1' #@param {type:'string'}
create_model_response = client.create_model(
model_display_name,
dataset=dataset,
train_budget_milli_node_hours=1000,
)
print('Create model operation: {}'.format(create_model_response.operation))
# Wait until model training is done.
model = create_model_response.result()
model
```
---
# 6. Make a prediction
In this section, we take our validation data prediction results and plot the Precision Recall Curve and the ROC curve of both the false and true predictions.
There are two different prediction modes: online and batch. The following cell shows you how to make a batch prediction. Please replace the 'your_test_bucket' of the gcs_destination with your own bucket where the predictions results will be stored by AutoML Tables.
```
#@title Start batch prediction { vertical-output: true, output-height: 200 }
batch_predict_gcs_input_uris = ['gs://cloud-ml-data-tables/notebooks/validation_unnested_FULL.csv',] #@param
batch_predict_gcs_output_uri_prefix = 'gs://{}'.format(bucket_name) #@param {type:'string'}
batch_predict_response = client.batch_predict(
model=model,
gcs_input_uris=batch_predict_gcs_input_uris,
gcs_output_uri_prefix=batch_predict_gcs_output_uri_prefix,
)
print('Batch prediction operation: {}'.format(batch_predict_response.operation))
# Wait until batch prediction is done.
batch_predict_result = batch_predict_response.result()
batch_predict_response.metadata
```
# 7. Evaluate your prediction
The follow cell creates a Precision Recall Curve and a ROC curve for both the true and false classifications.
Fill in the batch_predict_results_location with the location of the results.csv file created in the previous "Make a prediction" step
```
import numpy as np
from sklearn import metrics
import matplotlib.pyplot as plt
def invert(x):
return 1-x
def switch_label(x):
return(not x)
batch_predict_results_location = 'gs://<your-test-bucket>/' #@param {type:'string'}
table = pd.read_csv(batch_predict_results_location +'/<resultid>/tables_1.csv')
y = table["totalTransactionRevenue"]
scores = table["totalTransactionRevenue_1.0_score"]
scores_invert = table['totalTransactionRevenue_0.0_score']
#code for ROC curve, for true values
fpr, tpr, thresholds = metrics.roc_curve(y, scores)
roc_auc = metrics.auc(fpr, tpr)
plt.figure()
lw = 2
plt.plot(fpr, tpr, color='darkorange',
lw=lw, label='ROC curve (area = %0.2f)' % roc_auc)
plt.plot([0, 1], [0, 1], color='navy', lw=lw, linestyle='--')
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('Receiver operating characteristic for True')
plt.legend(loc="lower right")
plt.show()
#code for ROC curve, for false values
plt.figure()
lw = 2
label_invert = y.apply(switch_label)
fpr, tpr, thresholds = metrics.roc_curve(label_invert, scores_invert)
plt.plot(fpr, tpr, color='darkorange',
lw=lw, label='ROC curve (area = %0.2f)' % roc_auc)
plt.plot([0, 1], [0, 1], color='navy', lw=lw, linestyle='--')
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('Receiver operating characteristic for False')
plt.legend(loc="lower right")
plt.show()
#code for PR curve, for true values
precision, recall, thresholds = metrics.precision_recall_curve(y, scores)
plt.figure()
lw = 2
plt.plot( recall, precision, color='darkorange',
lw=lw, label='Precision recall curve for True')
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('Recall')
plt.ylabel('Precision')
plt.title('Precision Recall Curve for True')
plt.legend(loc="lower right")
plt.show()
#code for PR curve, for false values
precision, recall, thresholds = metrics.precision_recall_curve(label_invert, scores_invert)
print(precision.shape)
print(recall.shape)
plt.figure()
lw = 2
plt.plot( recall, precision, color='darkorange',
label='Precision recall curve for False')
plt.xlim([0.0, 1.1])
plt.ylim([0.0, 1.1])
plt.xlabel('Recall')
plt.ylabel('Precision')
plt.title('Precision Recall Curve for False')
plt.legend(loc="lower right")
plt.show()
```
| github_jupyter |
# Hourly and 10-min buoy data collocation with Saildrone data for validation
This is the code that collocations the Saildrone and buoy data for validation
First there is some reampling of the Saildrone data to match to averaging done by buoys
Buoy sample 10 min averages with the time at the end of the average
```
import xarray as xr
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
from math import atan2, log, pi, sin, cos
import datetime as dt
```
Read in Saildrone data and resample to match buoy averaging
- 10 min = time at end of 10 min ave
- hour = last 8 minutes of hour included
```
def read_saildrone(filename_usv):
#filename_usv='F:/data/cruise_data/saildrone/baja-2018/saildrone-gen_4-baja_2018-EP-sd1002-ALL-1_min-v1.nc'
ds_usv=xr.open_dataset(filename_usv)
ds_usv.close()
ds_usv = ds_usv.sel(trajectory=1002)
ilen_usv=(len(ds_usv.latitude['obs']))
ds_usv=ds_usv.rename({'latitude':'lat','longitude':'lon'})
#used to create a whole new dataset to switch obs to time but found swap dim command
ds_usv=ds_usv.swap_dims({'obs':'time'})
#WS_height=int(ds_usv.UWND_MEAN.installed_height)
lon = ds_usv.lon
lat = ds_usv.lat
del ds_usv['lon']
del ds_usv['lat']
ds_usv['lon'] = ('time', lon)
ds_usv['lat'] = ('time', lat)
#usv is at 5 m, buoys are at 5m. leave alone rather than changing to 10m wind
ds_usv['VWND_MEAN'] = (ds_usv.VWND_MEAN*log(10./1e-4))/log(5.0/1e-4)
ds_usv['UWND_MEAN'] = (ds_usv.UWND_MEAN*log(10./1e-4))/log(5.0/1e-4)
#ds_usv['UWND_MEAN'] = (ds_usv.UWND_MEAN*log(10./1e-4))/log(WS_height/1e-4)
#ds_usv['VWND_MEAN'] = (ds_usv.VWND_MEAN*log(10./1e-4))/log(WS_height/1e-4)
#ds_usv['UWND_MEAN'] = (ds_usv.UWND_MEAN*log(10./1e-4))/log(WS_height/1e-4)
#ds_usv['VWND_MEAN'] = (ds_usv.VWND_MEAN*log(10./1e-4))/log(WS_height/1e-4)
#make 10 min averages with time at end of average like buoy data
ds_usv_10min = ds_usv.resample(time='10min', skipna=True, closed='right', label='right').mean('time')
#this calculates hourly data that only uses the last 8 minutes of each hour for the average
sub = ds_usv.copy(deep=True)
pt = pd.to_datetime(sub.time.data)
pmin=pt.minute
tem=sub.where(pmin>=52) #data average should be only use min 52 to :60
ds_usv_1hr = tem.resample(time='1h', skipna=True, label='right').mean()
return ds_usv_10min,ds_usv_1hr
```
Read in buoy data. There are redundant sensors so you have to check the quality of the values and use the best quality only
```
def get_buoy_data(ibuoy):
if ibuoy==0:
sbuoy_id,sdep=['46011','D5']
if ibuoy==1:
sbuoy_id,sdep=['46012','D5']
if ibuoy==2:
sbuoy_id,sdep=['46042','D4']
if ibuoy==3:
sbuoy_id,sdep=['46086','D5']
if ibuoy==4:
sbuoy_id,sdep=['46028','D6']
if ibuoy==5:
sbuoy_id,sdep=['46047','D6']
fname='F:/data/cruise_data/saildrone/baja-2018/buoy_data/NDBC_'+sbuoy_id+'_201804_'+sdep+'_v0010min_xrformat.nc'
xr_buoy10=xr.open_dataset(fname)
fname='F:/data/cruise_data/saildrone/baja-2018/buoy_data/NDBC_'+sbuoy_id+'_201805_'+sdep+'_v0010min_xrformat.nc'
tem=xr.open_dataset(fname)
xr_buoy10 = xr.concat([xr_buoy10, tem],'time')
fname='F:/data/cruise_data/saildrone/baja-2018/buoy_data/NDBC_'+sbuoy_id+'_201806_'+sdep+'_v0010min_xrformat.nc'
tem=xr.open_dataset(fname)
xr_buoy10 = xr.concat([xr_buoy10, tem],'time')
btime10=xr_buoy10.time
#read in hourly buoy data
fname='F:/data/cruise_data/saildrone/baja-2018/buoy_data/NDBC_'+sbuoy_id+'_201804_'+sdep+'_v00hrly_xrformat.nc'
xr_buoy=xr.open_dataset(fname)
fname='F:/data/cruise_data/saildrone/baja-2018/buoy_data/NDBC_'+sbuoy_id+'_201805_'+sdep+'_v00hrly_xrformat.nc'
tem=xr.open_dataset(fname)
xr_buoy = xr.concat([xr_buoy, tem],'time')
fname='F:/data/cruise_data/saildrone/baja-2018/buoy_data/NDBC_'+sbuoy_id+'_201806_'+sdep+'_v00hrly_xrformat.nc'
tem=xr.open_dataset(fname)
xr_buoy = xr.concat([xr_buoy, tem],'time')
time=xr_buoy.time
xr_buoy['air_pres1']=xr_buoy.air_pres1/100.
xr_buoy['air_pres2']=xr_buoy.air_pres2/100.
xr_buoy['air_pres_sea_level1']=xr_buoy.air_pres_sea_level1/100.
xr_buoy['air_pres_sea_level2']=xr_buoy.air_pres_sea_level2/100.
#caluclate the best SST using quality flags then use these to calculate u and v
xr_buoy['sst_best']=xr_buoy['sst1'].copy(deep=True)
xr_buoy['sstqc_best']=xr_buoy['sst1qc'].copy(deep=True)
xr_buoy['sst_best']=xr_buoy.sst2.where((xr_buoy.sst1qc!=0) & (xr_buoy.sst2qc==0),other=xr_buoy.sst_best)
xr_buoy['sstqc_best']=xr_buoy.sst2qc.where((xr_buoy.sst1qc!=0) & (xr_buoy.sst2qc==0),other=xr_buoy.sstqc_best)
#caluclate the best AIR using quality flags then use these to calculate u and v
xr_buoy['air_best']=xr_buoy['air1'].copy(deep=True)
xr_buoy['airqc_best']=xr_buoy['air1qc'].copy(deep=True)
xr_buoy['air_best']=xr_buoy.air2.where((xr_buoy.air1qc!=0) & (xr_buoy.air2qc==0),other=xr_buoy.air_best)
xr_buoy['airqc_best']=xr_buoy.air2qc.where((xr_buoy.air1qc!=0) & (xr_buoy.air2qc==0),other=xr_buoy.airqc_best)
#caluclate the best AIR using quality flags then use these to calculate u and v
xr_buoy['air_pres_best']=xr_buoy['air_pres1'].copy(deep=True)
xr_buoy['air_presqc_best']=xr_buoy['air_pres1qc'].copy(deep=True)
xr_buoy['air_pres_best']=xr_buoy.air_pres2.where((xr_buoy.air_pres1qc!=0) & (xr_buoy.air_pres2qc==0),other=xr_buoy.air_pres_best)
xr_buoy['air_presqc_best']=xr_buoy.air_pres2qc.where((xr_buoy.air_pres1qc!=0) & (xr_buoy.air_pres2qc==0),other=xr_buoy.air_presqc_best)
#caluclate the best AIR using quality flags then use these to calculate u and v
xr_buoy['air_pres_sea_level_best']=xr_buoy['air_pres_sea_level1'].copy(deep=True)
xr_buoy['air_pres_sea_levelqc_best']=xr_buoy['air_pres_sea_level1qc'].copy(deep=True)
xr_buoy['air_pres_sea_level_best']=xr_buoy.air_pres_sea_level2.where((xr_buoy.air_pres_sea_level1qc!=0) & (xr_buoy.air_pres_sea_level2qc==0),other=xr_buoy.air_pres_sea_level_best)
xr_buoy['air_pres_sea_levelqc_best']=xr_buoy.air_pres_sea_level2qc.where((xr_buoy.air_pres_sea_level1qc!=0) & (xr_buoy.air_pres_sea_level2qc==0),other=xr_buoy.air_pres_sea_levelqc_best)
#caluclate the best wind speed and dirction using quality flags then use these to calculate u and v
xr_buoy['wnd_best']=xr_buoy['wnd1'].copy(deep=True)
xr_buoy['wdir_best']=xr_buoy['wdir1'].copy(deep=True)
xr_buoy['wdirqc_best']=xr_buoy['wdir1qc'].copy(deep=True)
xr_buoy['wnd_best']=xr_buoy.wnd2.where((xr_buoy.wdir1qc!=0) & (xr_buoy.wdir2qc==0),other=xr_buoy.wnd_best)
xr_buoy['wnd_best']=xr_buoy.wnd3.where((xr_buoy.wdir1qc!=0) & (xr_buoy.wdir3qc==0),other=xr_buoy.wnd_best)
xr_buoy['wnd_best']=xr_buoy.wnd4.where((xr_buoy.wdir1qc!=0) & (xr_buoy.wdir4qc==0),other=xr_buoy.wnd_best)
xr_buoy['wdir_best']=xr_buoy.wdir2.where((xr_buoy.wdir1qc!=0) & (xr_buoy.wdir2qc==0),other=xr_buoy.wdir_best)
xr_buoy['wdir_best']=xr_buoy.wdir3.where((xr_buoy.wdir1qc!=0) & (xr_buoy.wdir3qc==0),other=xr_buoy.wdir_best)
xr_buoy['wdir_best']=xr_buoy.wdir4.where((xr_buoy.wdir1qc!=0) & (xr_buoy.wdir4qc==0),other=xr_buoy.wdir_best)
xr_buoy['wdirqc_best']=xr_buoy.wdir2qc.where((xr_buoy.wdir1qc!=0) & (xr_buoy.wdir2qc==0),other=xr_buoy.wdirqc_best)
xr_buoy['wdirqc_best']=xr_buoy.wdir3qc.where((xr_buoy.wdir1qc!=0) & (xr_buoy.wdir3qc==0),other=xr_buoy.wdirqc_best)
xr_buoy['wdirqc_best']=xr_buoy.wdir4qc.where((xr_buoy.wdir1qc!=0) & (xr_buoy.wdir4qc==0),other=xr_buoy.wdirqc_best)
#now caluclate the wind vectors & adjust from met convention
xr_buoy['wnd_best']=xr_buoy.wnd_best.where(xr_buoy.wdirqc_best==0)
#change to 10m wind
xr_buoy['wnd_best'] = (xr_buoy.wnd_best*log(10./1e-4))/log(5/1e-4)
xr_buoy['wdir_best']=xr_buoy.wdir_best.where(xr_buoy.wdirqc_best==0)
xr_buoy['uwnd_best']=-1*xr_buoy.wnd_best*np.sin((xr_buoy.wdir_best)*pi/180.) #met conv is deg clock from true north, wind from
xr_buoy['vwnd_best']=-1*xr_buoy.wnd_best*np.cos((xr_buoy.wdir_best)*pi/180.)
#caluclate the best wind speed and dirction using quality flags then use these to calculate u and v
xr_buoy10['wnd_best']=xr_buoy10['wnd1'].copy(deep=True)
xr_buoy10['wdir_best']=xr_buoy10['wdir1'].copy(deep=True)
xr_buoy10['wdirqc_best']=xr_buoy10['wdir1qc'].copy(deep=True)
xr_buoy10['wnd_best']=xr_buoy10.wnd2.where((xr_buoy10.wdir1qc!=0) & (xr_buoy10.wdir2qc==0),other=xr_buoy10.wnd_best)
xr_buoy10['wnd_best']=xr_buoy10.wnd3.where((xr_buoy10.wdir1qc!=0) & (xr_buoy10.wdir3qc==0),other=xr_buoy10.wnd_best)
xr_buoy10['wnd_best']=xr_buoy10.wnd4.where((xr_buoy10.wdir1qc!=0) & (xr_buoy10.wdir4qc==0),other=xr_buoy10.wnd_best)
xr_buoy10['wdir_best']=xr_buoy10.wdir2.where((xr_buoy10.wdir1qc!=0) & (xr_buoy10.wdir2qc==0),other=xr_buoy10.wdir_best)
xr_buoy10['wdir_best']=xr_buoy10.wdir3.where((xr_buoy10.wdir1qc!=0) & (xr_buoy10.wdir3qc==0),other=xr_buoy10.wdir_best)
xr_buoy10['wdir_best']=xr_buoy10.wdir4.where((xr_buoy10.wdir1qc!=0) & (xr_buoy10.wdir4qc==0),other=xr_buoy10.wdir_best)
xr_buoy10['wdirqc_best']=xr_buoy10.wdir2qc.where((xr_buoy10.wdir1qc!=0) & (xr_buoy10.wdir2qc==0),other=xr_buoy10.wdirqc_best)
xr_buoy10['wdirqc_best']=xr_buoy10.wdir3qc.where((xr_buoy10.wdir1qc!=0) & (xr_buoy10.wdir3qc==0),other=xr_buoy10.wdirqc_best)
xr_buoy10['wdirqc_best']=xr_buoy10.wdir4qc.where((xr_buoy10.wdir1qc!=0) & (xr_buoy10.wdir4qc==0),other=xr_buoy10.wdirqc_best)
#now caluclate the wind vectors & adjust from met convention
xr_buoy10['wnd_best']=xr_buoy10.wnd_best.where(xr_buoy10.wdirqc_best==0)
#change to 10m wind
xr_buoy10['wnd_best'] = (xr_buoy10.wnd_best*log(10./1e-4))/log(5/1e-4)
xr_buoy10['wdir_best']=xr_buoy10.wdir_best.where(xr_buoy10.wdirqc_best==0)
xr_buoy10['uwnd_best']=-1*xr_buoy10.wnd_best*np.sin((xr_buoy10.wdir_best)*pi/180.) #met conv is deg clock from true north, wind from
xr_buoy10['vwnd_best']=-1*xr_buoy10.wnd_best*np.cos((xr_buoy10.wdir_best)*pi/180.)
cond = xr_buoy.lat > 90
xr_buoy.lat[cond]=np.nan
cond = xr_buoy.lon > 90
xr_buoy.lon[cond]=np.nan
cond = xr_buoy10.lat > 90
xr_buoy10.lat[cond]=np.nan
cond = xr_buoy10.lon > 90
xr_buoy10.lon[cond]=np.nan
return xr_buoy,xr_buoy10,sbuoy_id
def get_buoy_data_all():
for ibuoy in range(0,6):
xr_buoy,xr_buoy10,sbuoy_id = get_buoy_data(ibuoy)
if ibuoy>0:
xr_buoy2 = xr.concat([xr_buoy2, xr_buoy], dim='time')
xr_buoy102 = xr.concat([xr_buoy102, xr_buoy10], dim='time')
sbuoy_id2 = sbuoy_id2+sbuoy_id
if ibuoy==0:
xr_buoy2 = xr_buoy.copy(deep=True)
xr_buoy102 = xr_buoy10.copy(deep=True)
sbuoy_id2 = sbuoy_id
return xr_buoy2,xr_buoy102,sbuoy_id2
#read in saildrone data and average to 10 min and 60 min
filename_usv='F:/data/cruise_data/saildrone/baja-2018/saildrone-gen_4-baja_2018-sd1002-20180411T180000-20180611T055959-1_minutes-v1.nc'
ds_usv_10min,ds_usv_1hr = read_saildrone(filename_usv)
#make figure showing problem at low winds
sv_data=np.zeros((7,16))
fig, (ax1, ax2) = plt.subplots(1,2,figsize=(10,4))
for ibuoy in range(0,6):
if ibuoy<=6:
xr_buoy,xr_buoy10,sbuoy_id = get_buoy_data(ibuoy)
else:
xr_buoy,xr_buoy10,sbuoy_id = get_buoy_data_all()
btime=xr_buoy.time
btime10=xr_buoy10.time
#interpolate USV onto buoy
ilen_usv=(len(ds_usv_10min.TEMP_CTD_MEAN['time']))
#filled=ds_usv.interpolate_na(dim='time') #fill nan by interpolating
ds_usv10=ds_usv_10min.interp(time=btime10,method='nearest')
ds_usvhr=ds_usv_1hr.interp(time=btime,method='nearest')
dwin=.05
dist=((ds_usvhr.lat-xr_buoy.lat)**2+(ds_usvhr.lon-xr_buoy.lon)**2)**.5
cond,cond2 = (dist < dwin),((dist < dwin) & np.isfinite(ds_usvhr.UWND_MEAN) & np.isfinite(xr_buoy.wnd_best))
subset_usv=ds_usvhr.where(cond,drop=True)
subset_buoy=xr_buoy.where(cond,drop=True) #[cond]
subset_usv_nonan=ds_usvhr.where(cond2,drop=True)
subset_buoy_nonan=xr_buoy.where(cond2,drop=True) #[cond]
dist=((ds_usv10.lat-xr_buoy10.lat)**2+(ds_usv10.lon-xr_buoy10.lon)**2)**.5
cond,cond2 = (dist < dwin),((dist < dwin) & np.isfinite(ds_usv10.UWND_MEAN) & np.isfinite(xr_buoy10.wnd_best))
# cond = (dist < dwin)
subset_usv10=ds_usv10.where(cond,drop=True)
subset_buoy10=xr_buoy10.where(cond,drop=True) #[cond]
subset_usv10_nonan=ds_usv10.where(cond2,drop=True)
subset_buoy10_nonan=xr_buoy10.where(cond2,drop=True) #[cond]
usv_spd = np.sqrt(subset_usv10_nonan.UWND_MEAN**2+subset_usv10_nonan.VWND_MEAN**2)
buoy_spd = np.sqrt(subset_buoy10_nonan.uwnd_best**2+subset_buoy10_nonan.vwnd_best**2)
usv_dir = np.arctan2(subset_usv10_nonan.VWND_MEAN,subset_usv10_nonan.UWND_MEAN)*180/pi
buoy_dir = np.arctan2(subset_buoy10_nonan.vwnd_best,subset_buoy10_nonan.uwnd_best)*180/pi
dif_spd,dif_dir = usv_spd - buoy_spd, usv_dir - buoy_dir
cond,cond2 = (dif_dir > 180),(dif_dir < -180)
cond,cond2 = (dif_dir > 180),(dif_dir < -180)
dif_dir[cond]-=360
dif_dir[cond2]+=360
sv_data[ibuoy,11:13]=[dif_spd.mean().data,dif_spd.std().data]
sv_data[ibuoy,13:16]=[dif_dir.mean().data,dif_dir.std().data,dif_spd.shape[0]]
if dif_spd.shape[0]<1:
continue
ax1.plot(usv_spd,dif_dir,'.')
ax1.legend({'46011','46012','46042','46086'})
ax1.set_xlabel('Saildrone wind speed (ms$^{-1}$)')
ax1.set_ylabel('Saildrone - buoy wind direction (deg)')
ax2.plot(usv_spd,dif_spd,'.')
#ax2.legend({'46011','46012','46042','46086'})
ax2.set_xlabel('Saildrone wind speed (ms$^{-1}$)')
ax2.set_ylabel('Saildrone - buoy wind speed (ms$^{-1}$)')
ax1.plot([-5,15],[0,0],'k')
ax2.plot([-5,15],[0,0],'k')
ax1.set_xlim([-0.5,13])
ax2.set_xlim([-0.5,13])
fig_fname='F:/data/cruise_data/saildrone/baja-2018/figs/buoy_figures/buoy_wdir_wspd_both_bias.png'
fig.savefig(fig_fname, transparent=False, format='png')
sv_data=np.zeros((7,7))
sv_data2=np.zeros((7,7))
sv_data3=np.zeros((7,7))
sv_data4=np.zeros((7,7))
sv_data5=np.zeros((7,7))
sv_data6=np.zeros((7,7))
sv_data7=np.zeros((7,7))
for ibuoy in range(0,7):
if ibuoy<=5:
xr_buoy,xr_buoy10,sbuoy_id = get_buoy_data(ibuoy)
else:
xr_buoy,xr_buoy10,sbuoy_id = get_buoy_data_all()
#print(xr_buoy.air_pres1[0:5])
#fname='F:/data/cruise_data/saildrone/baja-2018/buoy_data/NDBC_46011_201804_D5_v00hrly_xrformat.nc'
#xr_buoy=xr.open_dataset(fname)
btime=xr_buoy.time
btime10=xr_buoy10.time
#interpolate USV onto buoy
ilen_usv=(len(ds_usv_10min.TEMP_CTD_MEAN['time']))
#filled=ds_usv.interpolate_na(dim='time') #fill nan by interpolating
ds_usv10=ds_usv_10min.interp(time=btime10,method='nearest')
ds_usvhr=ds_usv_1hr.interp(time=btime,method='nearest')
ds_usv10['wspd']=np.sqrt(ds_usv10.UWND_MEAN**2+ds_usv10.VWND_MEAN**2)
ds_usvhr['wspd']=np.sqrt(ds_usvhr.UWND_MEAN**2+ds_usvhr.VWND_MEAN**2)
dwin=.05
bair=xr_buoy.air1
dist=((ds_usvhr.lat-xr_buoy.lat)**2+(ds_usvhr.lon-xr_buoy.lon)**2)**.5
cond,cond2 = (dist < dwin),((dist < dwin) & np.isfinite(ds_usvhr.UWND_MEAN) & np.isfinite(xr_buoy.wnd_best))
cond3 = ((dist < dwin) & np.isfinite(ds_usvhr.UWND_MEAN) & np.isfinite(xr_buoy.wnd_best) & (ds_usvhr.wspd>3.5))
subset_usv=ds_usvhr.where(cond,drop=True)
subset_buoy=xr_buoy.where(cond,drop=True) #[cond]
subset_usv_nonan=ds_usvhr.where(cond2,drop=True)
subset_buoy_nonan=xr_buoy.where(cond2,drop=True) #[cond]
subset_usv_nonan_nolow=ds_usvhr.where(cond3,drop=True)
subset_buoy_nonan_nolow=xr_buoy.where(cond3,drop=True) #[cond]
dist=((ds_usv10.lat-xr_buoy10.lat)**2+(ds_usv10.lon-xr_buoy10.lon)**2)**.5
cond,cond2 = (dist < dwin),((dist < dwin) & np.isfinite(ds_usv10.UWND_MEAN) & np.isfinite(xr_buoy10.wnd_best))
cond3 = ((dist < dwin) & np.isfinite(ds_usv10.UWND_MEAN) & np.isfinite(xr_buoy10.wnd_best) & (ds_usv10.wspd>3.5))
subset_usv10=ds_usv10.where(cond,drop=True)
subset_buoy10=xr_buoy10.where(cond,drop=True) #[cond]
subset_usv10_nonan=ds_usv10.where(cond2,drop=True)
subset_buoy10_nonan=xr_buoy10.where(cond2,drop=True) #[cond]
subset_usv10_nonan_nolow=ds_usv10.where(cond3,drop=True)
subset_buoy10_nonan_nolow=xr_buoy10.where(cond3,drop=True) #[cond]
print(sbuoy_id) #,subset_usv.shape)
sdif = (subset_usv.TEMP_CTD_MEAN-subset_buoy.sst_best+273.15).dropna('time')
sdifcor = np.corrcoef(subset_usv.TEMP_CTD_MEAN,subset_buoy.sst_best-273.15)[0,1]
std_robust = np.nanmedian(np.abs(sdif - np.nanmedian(sdif))) * 1.482602218505602
ilen = sdif.shape[0]
sv_data[ibuoy,0:7]=[sdif.mean().data,sdif.median().data,sdifcor,sdif.std().data,std_robust,
np.abs(sdif).mean().data,sdif.shape[0]]
sdif = (subset_usv.TEMP_AIR_MEAN-subset_buoy.air_best+273.15).dropna('time')
sdifcor = np.corrcoef(subset_usv.TEMP_AIR_MEAN,subset_buoy.air_best-273.15)[0,1]
std_robust = np.nanmedian(np.abs(sdif - np.nanmedian(sdif))) * 1.482602218505602
print('air:') #,(sdif).mean().data,(sdif).std().data,sdif.shape[0])
sv_data2[ibuoy,0:7]=[sdif.mean().data,sdif.median().data,sdifcor,sdif.std().data,std_robust,
np.abs(sdif).mean().data,sdif.shape[0]]
sdif = (subset_usv.BARO_PRES_MEAN-subset_buoy.air_pres_best).dropna('time')
sdifcor = np.corrcoef(subset_usv.BARO_PRES_MEAN,subset_buoy.air_pres_best)[0,1]
std_robust = np.nanmedian(np.abs(sdif - np.nanmedian(sdif))) * 1.482602218505602
sv_data3[ibuoy,0:7]=[sdif.mean().data,sdif.median().data,sdifcor,sdif.std().data,std_robust,
np.abs(sdif).mean().data,sdif.shape[0]]
usv_spd = np.sqrt(subset_usv_nonan.UWND_MEAN**2+subset_usv_nonan.VWND_MEAN**2)
buoy_spd = np.sqrt(subset_buoy_nonan.uwnd_best**2+subset_buoy_nonan.vwnd_best**2)
usv_dir = np.arctan2(subset_usv_nonan_nolow.VWND_MEAN,subset_usv_nonan_nolow.UWND_MEAN)*180/pi
buoy_dir = np.arctan2(subset_buoy_nonan_nolow.vwnd_best,subset_buoy_nonan_nolow.uwnd_best)*180/pi
dif_spd,dif_dir = usv_spd - buoy_spd, usv_dir - buoy_dir
cond,cond2 = (dif_dir > 180),(dif_dir < -180)
dif_dir[cond]-=360
dif_dir[cond2]+=360
print('wnd_spd:',dif_spd.mean().data,dif_spd.std().data)
print('wnd_dir:',dif_dir.mean().data,dif_dir.std().data)
sdifcor = np.corrcoef(usv_spd,buoy_spd)[0,1]
std_robust = np.nanmedian(np.abs(dif_spd - np.nanmedian(dif_spd))) * 1.482602218505602
sdif = dif_spd
sv_data4[ibuoy,0:7]=[sdif.mean().data,sdif.median().data,sdifcor,sdif.std().data,std_robust,
np.abs(sdif).mean().data,sdif.shape[0]]
sdifcor = np.corrcoef(usv_dir,buoy_dir)[0,1]
std_robust = np.nanmedian(np.abs(dif_dir - np.nanmedian(dif_dir))) * 1.482602218505602
sdif = dif_dir
sv_data5[ibuoy,0:7]=[sdif.mean().data,sdif.median().data,sdifcor,sdif.std().data,std_robust,
np.abs(sdif).mean().data,sdif.shape[0]]
usv_spd = np.sqrt(subset_usv10_nonan.UWND_MEAN**2+subset_usv10_nonan.VWND_MEAN**2)
buoy_spd = np.sqrt(subset_buoy10_nonan.uwnd_best**2+subset_buoy10_nonan.vwnd_best**2)
usv_dir = np.arctan2(subset_usv10_nonan_nolow.VWND_MEAN,subset_usv10_nonan_nolow.UWND_MEAN)*180/pi
buoy_dir = np.arctan2(subset_buoy10_nonan_nolow.vwnd_best,subset_buoy10_nonan_nolow.uwnd_best)*180/pi
dif_spd,dif_dir = usv_spd - buoy_spd, usv_dir - buoy_dir
cond,cond2 = (dif_dir > 180),(dif_dir < -180)
dif_dir[cond]-=360
dif_dir[cond2]+=360
print('wnd_spd:',dif_spd.mean().data,dif_spd.std().data,dif_spd.shape[0])
print('wnd_dir:',dif_dir.mean().data,dif_dir.std().data,dif_dir.shape[0])
sdifcor = np.corrcoef(usv_spd,buoy_spd)[0,1]
std_robust = np.nanmedian(np.abs(dif_spd - np.nanmedian(dif_spd))) * 1.482602218505602
sdif = dif_spd
sv_data6[ibuoy,0:7]=[sdif.mean().data,sdif.median().data,sdifcor,sdif.std().data,std_robust,
np.abs(sdif).mean().data,sdif.shape[0]]
sdifcor = np.corrcoef(usv_dir,buoy_dir)[0,1]
std_robust = np.nanmedian(np.abs(dif_dir - np.nanmedian(dif_dir))) * 1.482602218505602
sdif = dif_dir
sv_data7[ibuoy,0:7]=[sdif.mean().data,sdif.median().data,sdifcor,sdif.std().data,std_robust,
np.abs(sdif).mean().data,sdif.shape[0]]
sdif = (subset_usv.BARO_PRES_MEAN-subset_buoy.air_pres_best).dropna('time')
sdifcor = np.corrcoef(subset_usv.BARO_PRES_MEAN,subset_buoy.air_pres_best)[0,1]
std_robust = np.nanmedian(np.abs(sdif - np.nanmedian(sdif))) * 1.482602218505602
print([sdif.mean().data,sdif.median().data,sdifcor,sdif.std().data,std_robust,
np.abs(sdif).mean().data,sdif.shape[0]])
plt.plot(np.abs(sdif - np.nanmedian(sdif)))
print(np.nanmedian(np.abs(sdif - np.nanmedian(sdif))))
print(sdif.std().data)
#this is explaining why the STD is larger than the robust STD
#robust STD uses median which removes the spikes since there aren't many of them
#this shows that the pressure sensor sometimes goes back to the right observation! odd.
plt.plot(subset_usv.BARO_PRES_MEAN,'.')
plt.plot(subset_buoy.air_pres_best,'.')
plt.plot(ds_usv_10min.BARO_PRES_MEAN[1000:2000])
float_formatter = lambda x: "%.2f" % x
np.set_printoptions(formatter={'float_kind':float_formatter})
for i in range(7):
print(sv_data[i,:])
for i in range(0,7):
print(sv_data2[i,:])
for i in range(0,7):
print(sv_data3[i,:])
for i in range(0,7):
print(sv_data4[i,:])
for i in range(0,7):
print(sv_data5[i,:])
for i in range(0,7):
print(sv_data6[i,:])
for i in range(0,7):
print(sv_data7[i,:])
#plot buoy and saildrone data
fig, (ax1, ax2) = plt.subplots(1,2)
for ibuoy in range(0,6):
xr_buoy,xr_buoy10,sbuoy_id = get_buoy_data(ibuoy)
btime=xr_buoy.time
btime10=xr_buoy10.time
#interpolate USV onto buoy
ilen_usv=(len(ds_usv_10min.TEMP_CTD_MEAN['time']))
#filled=ds_usv.interpolate_na(dim='time') #fill nan by interpolating
ds_usv10=ds_usv_10min.interp(time=btime10,method='nearest')
ds_usvhr=ds_usv_1hr.interp(time=btime,method='nearest')
ds_usv10['wspd']=np.sqrt(ds_usv10.UWND_MEAN**2+ds_usv10.VWND_MEAN**2)
ds_usvhr['wspd']=np.sqrt(ds_usvhr.UWND_MEAN**2+ds_usvhr.VWND_MEAN**2)
#print(xr_buoy.air_pres1[0:5])
#fname='F:/data/cruise_data/saildrone/baja-2018/buoy_data/NDBC_46011_201804_D5_v00hrly_xrformat.nc'
#xr_buoy=xr.open_dataset(fname)
# cond = xr_buoy.lat > 90
# xr_buoy.lat[cond]=np.nan
## cond = xr_buoy.lon > 90
## xr_buoy.lon[cond]=np.nan
# blat=xr_buoy.lat.mean()
# blon=xr_buoy.lon.mean()
# btime=xr_buoy.time
#interpolate USV onto buoy
# ilen_usv=(len(ds_usv_1hr.TEMP_CTD_MEAN['time']))
#filled=ds_usv.interpolate_na(dim='time') #fill nan by interpolating
# ds_usv10=ds_usv_10min.interp(time=btime10,method='nearest')
# ds_usvhr=ds_usv_1hr.interp(time=btime,method='nearest')
fig, (ax1, ax2) = plt.subplots(1,2)
dwin=.05
dist=((ds_usvhr.lat-xr_buoy.lat)**2+(ds_usvhr.lon-xr_buoy.lon)**2)**.5
cond = (dist < dwin)
subset_usv=ds_usvhr.where(cond,drop=True)
subset_buoy=xr_buoy.where(cond,drop=True) #[cond]
dist=((ds_usv10.lat-xr_buoy10.lat)**2+(ds_usv10.lon-xr_buoy10.lon)**2)**.5
cond = (dist < dwin)
subset_usv10=ds_usv10.where(cond,drop=True)
subset_buoy10=xr_buoy10.where(cond,drop=True) #[cond]
ax1.scatter(subset_usv10.lon,subset_usv10.lat, c=subset_usv10.TEMP_CTD_MEAN,s=5,marker = "o")
ax1.scatter(subset_buoy.lon,subset_buoy.lat, c=subset_buoy.sst_best,s=5,marker="D")
ax1.quiver(subset_usv10.lon,subset_usv10.lat,subset_usv10.UWND_MEAN,subset_usv10.VWND_MEAN,color='k')
ax1.quiver(subset_buoy10.lon,subset_buoy10.lat,subset_buoy10.uwnd_best,subset_buoy10.vwnd_best,color='r')
ax1.legend(['USV','buoy'])
ymin, ymax = ax1.get_ylim()
xmin, xmax = ax1.get_xlim()
# print(xmin,xmax,ymin,ymax)
ax2.scatter(ds_usv.lon[0::60],ds_usv.lat[0::60], c=ds_usv.TEMP_CTD_MEAN[0::60],s=5,marker = "o")
ax2.scatter(xr_buoy.lon[0::60],xr_buoy.lat[0::60], c=xr_buoy.sst_best[0::60],s=5,marker="D")
ax2.quiver(ds_usv.lon[0::60],ds_usv.lat[0::60],ds_usv.UWND_MEAN[0::60],ds_usv.VWND_MEAN[0::60],color='k')
ax2.quiver(xr_buoy.lon[0::60],xr_buoy.lat[0::60],xr_buoy.uwnd_best[0::60],xr_buoy.vwnd_best[0::60],color='r')
ax2.legend(['USV','buoy'])
ax2.set_ylim(ymin,ymax)
ax2.set_xlim(xmin,xmax)
#ax2.plot(subset_usv10.time,subset_usv10.TEMP_CTD_MEAN-273.15,'.')
#ax2.plot(subset_buoy.time,subset_buoy.best_sst)
# ax2.set_xlim(tstart_buffer,tend_buffer)
# ax2.set_ylim(11,14.5)
# rotate and align the tick labels so they look better
fig.autofmt_xdate()
fig_fname='F:/data/cruise_data/saildrone/baja-2018/figs/buoy_figures/buoy_loc_image'+sbuoy_id+'_test.png'
fig.savefig(fig_fname, transparent=False, format='png')
plt.clf()
#plot buoy and saildrone data all on one figure
fig, ax0 = plt.subplots(1,6)
for ibuoy in range(0,6):
xr_buoy,xr_buoy10,sbuoy_id = get_buoy_data(ibuoy)
#print(xr_buoy.air_pres1[0:5])
#fname='F:/data/cruise_data/saildrone/baja-2018/buoy_data/NDBC_46011_201804_D5_v00hrly_xrformat.nc'
#xr_buoy=xr.open_dataset(fname)
cond = xr_buoy.lat > 90
xr_buoy.lat[cond]=np.nan
cond = xr_buoy.lon > 90
xr_buoy.lon[cond]=np.nan
blat=xr_buoy.lat.mean()
blon=xr_buoy.lon.mean()
btime=xr_buoy.time
dwin=.05
subset=ds_usv.copy(deep=True) #sel(time=slice(time_start1,time_end1))
lat_usv=subset.lat
lon_usv=subset.lon
time_usv=subset.time
sst_usv=subset.TEMP_CTD_MEAN
meets_condition = (lat_usv>blat-dwin) & (lat_usv<blat+dwin) & (lon_usv>blon-dwin) & (lon_usv<blon+dwin) #& time_usv<dt.datetime('2018-04-20')
if (lon_usv[meets_condition].shape[0]==0):
continue
ax1.scatter(lon_usv[meets_condition],lat_usv[meets_condition], c=sst_usv[meets_condition], s=5)
ax1.plot(xr_buoy.lon.sel(time=slice(tstart_buffer,tend_buffer)),xr_buoy.lat.sel(time=slice(tstart_buffer,tend_buffer)),'r.')
ax1.legend(['Buoy','USV'])
ax2.plot(subset.time[meets_condition],subset.TEMP_CTD_MEAN[meets_condition]-273.15,'.')
ax2.plot(xr_buoy.time,xr_buoy.sst1-273.15)
ax2.set_xlim(tstart_buffer,tend_buffer)
ax2.set_ylim(11,14.5)
# rotate and align the tick labels so they look better
fig.autofmt_xdate()
fig_fname='F:/data/cruise_data/saildrone/baja-2018/figs/buoy_figures/buoy_loc_image'+sbuoy_id+'_.png'
fig.savefig(fig_fname, transparent=False, format='png')
plt.clf()
dwin=.02
meets_condition = (lat_usv>blat-dwin) & (lat_usv<blat+dwin) & (lon_usv>blon-dwin) & (lon_usv<blon+dwin) #& time_usv<dt.datetime('2018-04-20')
fig, (ax1, ax2) = plt.subplots(1,2)
if (lon_usv[meets_condition].shape[0]==0):
continue
ax1.scatter(lon_usv[meets_condition],lat_usv[meets_condition], c=sst_usv[meets_condition], s=5)
ax1.plot(xr_buoy.lon.sel(time=slice(tstart_buffer,tend_buffer)),xr_buoy.lat.sel(time=slice(tstart_buffer,tend_buffer)),'r.')
ax1.legend(['Buoy','USV'])
ax2.plot(subset.time[meets_condition],subset.TEMP_CTD_MEAN[meets_condition]-273.15,'.')
ax2.plot(xr_buoy.time,xr_buoy.sst1-273.15)
ax2.set_xlim(tstart_buffer,tend_buffer)
ax2.set_ylim(11,14.5)
# rotate and align the tick labels so they look better
fig.autofmt_xdate()
fig_fname='F:/data/cruise_data/saildrone/baja-2018/figs/buoy_figures/buoy_loc_image'+sbuoy_id+'_02km.png'
fig.savefig(fig_fname, transparent=False, format='png')
plt.clf()
#now i only use part of buoy location that is for this time
dwin=.01
# blon_subset=xr_buoy.lon.sel(time=slice(time_start1,time_end1)).mean()
# blat_subset=xr_buoy.lat.sel(time=slice(time_start1,time_end1)).mean()
meets_condition = (lat_usv>blat-dwin) & (lat_usv<blat+dwin) & (lon_usv>blon-dwin) & (lon_usv<blon+dwin) #& time_usv<dt.datetime('2018-04-20')
# meets_condition = (lat_usv>blat_subset-dwin) & (lat_usv<blat_subset+dwin) & (lon_usv>blon_subset-dwin) & (lon_usv<blon_subset+dwin) #& time_usv<dt.datetime('2018-04-20')
fig, (ax1, ax2) = plt.subplots(1,2)
if (lon_usv[meets_condition].shape[0]==0):
continue
ax1.scatter(lon_usv[meets_condition],lat_usv[meets_condition], c=sst_usv[meets_condition], s=5)
ax1.plot(xr_buoy.lon.sel(time=slice(time_start1,time_end1)),xr_buoy.lat.sel(time=slice(time_start1,time_end1)),'r.')
ax1.legend(['Buoy','USV'])
ax2.plot(subset.time[meets_condition],subset.TEMP_CTD_MEAN[meets_condition]-273.15,'.')
ax2.plot(xr_buoy.time,xr_buoy.sst1-273.15)
ax2.set_xlim(time_start1,time_end1)
ax2.set_ylim(11,14.5)
# rotate and align the tick labels so they look better
fig.autofmt_xdate()
fig_fname='F:/data/cruise_data/saildrone/baja-2018/figs/buoy_figures/buoy_loc_image'+sbuoy_id+'_01km.png'
fig.savefig(fig_fname, transparent=False, format='png')
plt.clf()
#####old
filename_usv='F:/data/cruise_data/saildrone/baja-2018/saildrone-gen_4-baja_2018-sd1002-20180411T180000-20180611T055959-1_minutes-v1.nc'
#ds_usv_tem=xr.open_dataset(filename_usv)
ds_usv = read_saildrone(filename_usv)
ilen_usv=(len(ds_usv.TEMP_CTD_MEAN['time']))
filled=ds_usv.interpolate_na(dim='time') #fill nan by interpolating
#move lat and lon into variables so that they don't disappear when interpolating
lon = ds_usv.lon
lat = ds_usv.lat
del ds_usv['lon']
del ds_usv['lat']
ds_usv['lon'] = ('time', lon)
ds_usv['lat'] = ('time', lat)
ds_usv10=filled.interp(time=btime10)
ds_usvhr=filled.interp(time=btime)
#this take the average from min: 01 to :10 and assigns it :10
#use this for the wind comparisons since they are 10 min ave
subset = ds_usv.sel(time=slice('2018-04-10','2018-04-13'))
ds_usv_10min = ds_usv.resample(time='10min', skipna=True, closed='right', label='right').mean('time')
#this calculates hourly data that only uses the last 8 minutes of each hour for the average
#use this for the hourly data comparisons
sub = ds_usv.copy(deep=True)
pt = pd.to_datetime(sub.time.data)
pmin=pt.minute
tem=sub.where(pmin>=52) #data average should be only use min 52 to :60
ds_usv_1hr = tem.resample(time='1h', skipna=True, label='right').mean()
filename_usv='F:/data/cruise_data/saildrone/baja-2018/saildrone-gen_4-baja_2018-sd1002-20180411T180000-20180611T055959-1_minutes-v1.nc'
ds_usv = read_saildrone(filename_usv)
#ds_usv.close()
ilen_usv=(len(ds_usv.TEMP_CTD_MEAN['time']))
filled=ds_usv.interpolate_na(dim='time') #fill nan by interpolating
#ds_usv10=filled.interp(time=btime10)
#ds_usvhr=filled.interp(time=btime)
#ds_usv10=filled.resample(time='10Min').mean() #this code creates 10 averages
#ds_usvhr=filled.resample(time='1H').mean() #this code creates hourly averages
#this take the average from min: 01 to :10 and assigns it :10
ds_usv_10min = ds_usv.resample(time='10min', skipna=True, closed='right', label='right').mean('time')
#this calculates hourly data that only uses the last 8 minutes of each hour for the average
sub = ds_usv.copy(deep=True)
pt = pd.to_datetime(sub.time.data)
pmin=pt.minute
tem=sub.where(pmin>=52) #data average should be only use min 52 to :60
ds_usv_1hr = tem.resample(time='1h', skipna=True, label='right').mean()
#dates_usv64=ds_usv.time[0,:].values
#dates_usv=pd.to_datetime(dates_usv64, unit='ns') #new way to caluclate the time array
#ds=xr.open_dataset(filename_usv)
#ds = ds.sel(trajectory=1002)
#ds=ds.rename({'latitude':'lat','longitude':'lon'})
#ds=ds.swap_dims({'obs':'time'})
#print(ds)
#print(ds.time[0:10])
#print(xr_buoy.wnd_best[0:2])
for i in range(1,2):
print(i,xr_buoy.time[i].data,xr_buoy.wnd_best[i].data)
print('10min')
for i in range(0,15):
print(i,xr_buoy10.time[i].data,xr_buoy10.wnd_best[i].data)
print(xr_buoy10.wnd_best[6:12].mean().data)
subset = ds_usv.sel(time=slice('2018-04-10','2018-04-13'))
xftem=subset.UWND_MEAN
pt = pd.to_datetime(xftem.time.data)
pmin=pt.minute
tem=xftem[(pmin >=52)] #data average should be only use min 52 to :60
new_hr = tem.resample(time='1h').mean()
for i in range(0,1):
print(new_hr.time[i].data,new_hr[i].data)
print(xftem[52:60].mean().data)
#xftem=subset.UWND_MEAN
sub = subset.copy(deep=True)
pt = pd.to_datetime(sub.time.data)
pmin=pt.minute
#sub['mask']=pmin
tem=sub.where(pmin>=52) #data average should be only use min 52 to :60
new_hr = tem.resample(time='1h', label='right').mean()
#print(new_hr)
for i in range(0,1):
print(new_hr.time[i].data,new_hr.UWND_MEAN[i].data)
#print('min')
#for i in range(0,60):
# print(i,xftem.time[i].data,xftem[i].data)
print(subset.UWND_MEAN[52:60].mean().data)
#read in high resolution wind buoy data (10 minute)
sbuoy_id='46011'
sdep='D5'
fname='F:/data/cruise_data/saildrone/baja-2018/buoy_data/NDBC_'+sbuoy_id+'_201804_'+sdep+'_v0010min_xrformat.nc'
xr_buoy10=xr.open_dataset(fname)
fname='F:/data/cruise_data/saildrone/baja-2018/buoy_data/NDBC_'+sbuoy_id+'_201805_'+sdep+'_v0010min_xrformat.nc'
tem=xr.open_dataset(fname)
xr_buoy10 = xr.concat([xr_buoy10, tem],'time')
fname='F:/data/cruise_data/saildrone/baja-2018/buoy_data/NDBC_'+sbuoy_id+'_201806_'+sdep+'_v0010min_xrformat.nc'
tem=xr.open_dataset(fname)
xr_buoy10 = xr.concat([xr_buoy10, tem],'time')
btime10=xr_buoy10.time#read in hourly buoy data
fname='F:/data/cruise_data/saildrone/baja-2018/buoy_data/NDBC_'+sbuoy_id+'_201804_'+sdep+'_v00hrly_xrformat.nc'
xr_buoy=xr.open_dataset(fname)
fname='F:/data/cruise_data/saildrone/baja-2018/buoy_data/NDBC_'+sbuoy_id+'_201805_'+sdep+'_v00hrly_xrformat.nc'
tem=xr.open_dataset(fname)
xr_buoy = xr.concat([xr_buoy, tem],'time')
fname='F:/data/cruise_data/saildrone/baja-2018/buoy_data/NDBC_'+sbuoy_id+'_201806_'+sdep+'_v00hrly_xrformat.nc'
tem=xr.open_dataset(fname)
xr_buoy = xr.concat([xr_buoy, tem],'time')
time=xr_buoy.time
#fname='F:/data/cruise_data/saildrone/baja-2018/buoy_data/NDBC_46011_201804_D5_v00hrly_xrformat.nc'
#xr_buoy=xr.open_dataset(fname)
cond = xr_buoy.lat > 90
xr_buoy.lat[cond]=np.nan
cond = xr_buoy.lon > 90
xr_buoy.lon[cond]=np.nan
blat=xr_buoy.lat.mean()
blon=xr_buoy.lon.mean()
btime=xr_buoy.time
dwin=.02
meets_condition = (lat_usv>blat-dwin) & (lat_usv<blat+dwin) & (lon_usv>blon-dwin) & (lon_usv<blon+dwin) #& time_usv<dt.datetime('2018-04-20')
fig, (ax1, ax2) = plt.subplots(1,2)
if (lon_usv[meets_condition].shape[0]==0):
continue
ax1.scatter(lon_usv[meets_condition],lat_usv[meets_condition], c=sst_usv[meets_condition], s=5)
ax1.plot(xr_buoy.lon.sel(time=slice(tstart_buffer,tend_buffer)),xr_buoy.lat.sel(time=slice(tstart_buffer,tend_buffer)),'r.')
ax1.legend(['Buoy','USV'])
ax2.plot(subset.time[meets_condition],subset.TEMP_CTD_MEAN[meets_condition]-273.15,'.')
ax2.plot(xr_buoy.time,xr_buoy.sst1-273.15)
ax2.set_xlim(tstart_buffer,tend_buffer)
ax2.set_ylim(11,14.5)
# rotate and align the tick labels so they look better
fig.autofmt_xdate()
fig_fname='F:/data/cruise_data/saildrone/baja-2018/figs/buoy_figures/buoy_loc_image'+sbuoy_id+'_02km.png'
fig.savefig(fig_fname, transparent=False, format='png')
plt.clf()
#now i only use part of buoy location that is for this time
dwin=.01
# blon_subset=xr_buoy.lon.sel(time=slice(time_start1,time_end1)).mean()
# blat_subset=xr_buoy.lat.sel(time=slice(time_start1,time_end1)).mean()
meets_condition = (lat_usv>blat-dwin) & (lat_usv<blat+dwin) & (lon_usv>blon-dwin) & (lon_usv<blon+dwin) #& time_usv<dt.datetime('2018-04-20')
# meets_condition = (lat_usv>blat_subset-dwin) & (lat_usv<blat_subset+dwin) & (lon_usv>blon_subset-dwin) & (lon_usv<blon_subset+dwin) #& time_usv<dt.datetime('2018-04-20')
fig, (ax1, ax2) = plt.subplots(1,2)
if (lon_usv[meets_condition].shape[0]==0):
continue
ax1.scatter(lon_usv[meets_condition],lat_usv[meets_condition], c=sst_usv[meets_condition], s=5)
ax1.plot(xr_buoy.lon.sel(time=slice(time_start1,time_end1)),xr_buoy.lat.sel(time=slice(time_start1,time_end1)),'r.')
ax1.legend(['Buoy','USV'])
ax2.plot(subset.time[meets_condition],subset.TEMP_CTD_MEAN[meets_condition]-273.15,'.')
ax2.plot(xr_buoy.time,xr_buoy.sst1-273.15)
ax2.set_xlim(time_start1,time_end1)
ax2.set_ylim(11,14.5)
# rotate and align the tick labels so they look better
fig.autofmt_xdate()
fig_fname='F:/data/cruise_data/saildrone/baja-2018/figs/buoy_figures/buoy_loc_image'+sbuoy_id+'_01km.png'
fig.savefig(fig_fname, transparent=False, format='png')
plt.clf()
```
| github_jupyter |
<a href="https://colab.research.google.com/github/cstorm125/align_use/blob/master/sandbox.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Universal Sentence Encoder for Sentence Alignment
```
# #get europarl
# !wget https://www.dropbox.com/s/nr5h3xuqhlvj94m/europarl-v7.fr-en.en
# !wget https://www.dropbox.com/s/z730widwr5kw0zm/europarl-v7.fr-en.fr
# #get bucc2018
# !wget https://comparable.limsi.fr/bucc2018/bucc2018-fr-en.sample-gold.tar.bz2; tar -xf bucc2018-fr-en.sample-gold.tar.bz2; cd bucc2018/fr-en; mv * ../..
# #install dependencies
# !pip install tensorflow_text
!ls
import pandas as pd
import numpy as np
import tqdm
import tensorflow_hub as hub
import tensorflow_text
import tensorflow as tf #tensorflow 2.1.0
_model = hub.load('https://tfhub.dev/google/universal-sentence-encoder-multilingual/3')
class LSHasher:
def __init__(self,hash_size = 10, input_dim = 512):
self.hash_size = hash_size
self.input_dim = input_dim
self.projections = tf.random.normal((self.input_dim, self.hash_size))
def get_hash(self,inp):
res = inp@self.projections
res = tf.cast(res > 0,tf.int32).numpy().astype(str)
res = [''.join(i) for i in res]
return res
def get_alignment(all_emb,input_idx):
ref_idx = tf.constant(input_idx)[:,None]
sub_emb = tf.gather_nd(params=all_emb, indices=ref_idx)
x_ = tf.nn.l2_normalize(sub_emb,axis=1)
res = x_@tf.transpose(x_)
res = tf.linalg.set_diag(res, tf.constant(-2, dtype=float, shape=len(input_idx)))
max_idx = tf.argmax(res,0).numpy()
max_values = tf.reduce_max(res,0).numpy()
max_input_idx = [input_idx[i] for i in max_idx]
return max_input_idx, max_values
def get_alignment2(emb1,emb2):
emb1_ = tf.nn.l2_normalize(emb1,axis=1)
emb2_ = tf.nn.l2_normalize(emb2,axis=1)
res = emb1_@tf.transpose(emb2_)
max_idx = tf.argmax(res,1).numpy()
max_values = tf.reduce_max(res,1).numpy()
return max_idx, max_values
```
## Small Baseline with `europarl fr-en`
```
#read files
with open('europarl-v7.fr-en.fr','r') as f: fr = f.readlines()
with open('europarl-v7.fr-en.en','r') as f: en = f.readlines()
len(fr), len(en)
#create dataframe
df = pd.DataFrame({'fr':[i[:-1] for i in fr],'en':[i[:-1] for i in en]})
#deduplicate based on fr
df['rnk'] = df.groupby('fr').cumcount()
df = df[df.rnk==0].drop('rnk',1).reset_index(drop=True)
df.tail()
#randomize a subset of 1000 deduplicated sentences
rand_idx = np.random.choice(df.index, size=1000)
#get use
%time fr_emb = _model(df['fr'][rand_idx])
%time en_emb = _model(df['en'][rand_idx])
all_emb = tf.concat([fr_emb,en_emb],0)
all_emb.shape
#test with first 1000 rows of deduped fr-en europarl
input_idx = [i for i in range(2000)]
max_input_idx, max_values = get_alignment(all_emb,input_idx)
res_df = pd.DataFrame({'src_idx':input_idx, 'targ_idx':max_input_idx,'score':max_values})
res_df
#fr to en
(res_df.iloc[:1000,:].src_idx==res_df.iloc[:1000,:].targ_idx-1000).mean()
#en to fr
(res_df.iloc[1000:,:].src_idx==res_df.iloc[1000:,:].targ_idx+1000).mean()
```
## Larger Baseline with `bucc2018 fr-en`
```
!ls
fr = pd.read_csv('fr-en.sample.fr',sep='\t',header=None)
en = pd.read_csv('fr-en.sample.en',sep='\t',header=None)
for i in [fr,en]: i.columns = ['id','text']
gold = pd.read_csv('fr-en.sample.gold',sep='\t',header=None)
gold.columns=['fr_id','en_id']
fr.shape,en.shape,gold.shape
fr_embs = []
en_embs = []
bs = 10000
for i in range(fr.shape[0]//bs+1):
fr_embs.append(_model(fr.iloc[bs*i:bs*(i+1),1]))
for i in range(en.shape[0]//bs+1):
en_embs.append(_model(en.iloc[bs*i:bs*(i+1),1]))
fr_emb = tf.concat(fr_embs,0)
en_emb = tf.concat(en_embs,0)
fr_emb.shape, en_emb.shape
get_alignment2(fr_emb[:10000,:],fr_emb[:10000,:])
reses = []
for i in range(fr_emb.shape[0]//bs+1):
src_id = fr.iloc[bs*i:bs*(i+1),0]
for j in range(en_emb.shape[0]//bs+1):
dest_idx, score = get_alignment2(fr_emb[bs*i:bs*(i+1),:],en_emb[bs*j:bs*(j+1),:])
dest_id = en.iloc[dest_idx,0]
reses.append(pd.DataFrame({'src_id':list(src_id),'dest_id': list(dest_id),'score':score}))
res_df = pd.concat(reses,0).reset_index(drop=True)
res_df['rnk'] = res_df.sort_values('score',ascending=False).groupby('src_id').cumcount()
res_df = res_df[res_df.rnk==0].drop('rnk',1)
res_df.shape
#distribution of similarity score
res_df.score.hist()
#predict match only when similarity is above threshold
threses = [i/100 for i in range(5,100,5)]
for thres in threses:
res_joined = res_df[res_df.score>thres].merge(gold,how='left',left_on='src_id',right_on='fr_id')
precision = (res_joined.dest_id==res_joined.en_id).sum() / res_joined.shape[0]
recall = (res_joined.dest_id==res_joined.en_id).sum() / gold.shape[0]
print(f'''
threshold: {thres}
accuracy: {(res_joined.dest_id==res_joined.en_id).mean()}
f1: {2*((precision*recall)/(precision+recall))}
precision: {precision}
recall: {recall}
''')
#best thres around 0.7
thres=0.7
res_joined = res_df[res_df.score>thres].merge(gold,how='left',left_on='src_id',right_on='fr_id')
precision = (res_joined.dest_id==res_joined.en_id).sum() / res_joined.shape[0]
recall = (res_joined.dest_id==res_joined.en_id).sum() / gold.shape[0]
print(f'''
threshold: {thres}
accuracy: {(res_joined.dest_id==res_joined.en_id).mean()}
f1: {2*((precision*recall)/(precision+recall))}
precision: {precision}
recall: {recall}
''')
```
| github_jupyter |
```
import sys
import pandas as pd
import numpy as np
import scipy.stats as stats
import matplotlib.pyplot as plt
from sklearn.preprocessing import PowerTransformer, FunctionTransformer, QuantileTransformer
from sklearn.metrics import mean_squared_error
sys.path.append('../Scripts')
from Data_Processing import DataProcessing
import warnings
warnings.filterwarnings('ignore')
pd.set_option('display.max_columns', None)
df = DataProcessing('../Data/train.csv')
df
#df = df.loc[df['Lap_Time'] != 0]
y = df['Lap_Time']
X = df.drop(columns=['Lap_Time'])
obj_columns = list(X.select_dtypes(include=object).columns)
obj_columns.append('Lap_Improvement')
obj_columns.append('Lap_Number')
obj_columns.append('S1_Improvement')
obj_columns.append('S2_Improvement')
obj_columns.append('S3_Improvement')
num_columns = list(X.select_dtypes(include='number').columns)
num_columns.remove('Lap_Number')
num_columns.remove('Lap_Improvement')
num_columns.remove('S1_Improvement')
num_columns.remove('S2_Improvement')
num_columns.remove('S3_Improvement')
```
# Scalers
```
from sklearn.compose import ColumnTransformer
from sklearn.preprocessing import StandardScaler, OneHotEncoder, MinMaxScaler, OrdinalEncoder
import joblib
#pt = PowerTransformer(method='yeo-johnson')
#X[num_columns] = pt.fit_transform(X[num_columns])
X
column_transformer = ColumnTransformer(
[('num', MinMaxScaler(), num_columns),
('obj', OneHotEncoder(drop='first'), obj_columns)],
remainder='passthrough')
trans_X = column_transformer.fit_transform(X)
joblib.dump(column_transformer, '../Models/Column_Transformer_NN.pkl')
#joblib.dump(pt, '../Models/Power_Transformer.pkl')
#trans_X = trans_X.toarray()
y = np.asarray(y).astype(float)
scaler = StandardScaler()
y = scaler.fit_transform(y.reshape(-1,1))
joblib.dump(scaler, '../Models/NN_Y_Scaler.pkl')
```
# Train Test Split
```
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(trans_X, y, random_state=42, test_size=0.1, shuffle=True)
X_train.shape
```
# Neural Network
```
from tensorflow import keras
from keras.callbacks import ModelCheckpoint
from keras.models import load_model
from keras import backend as K
from keras.callbacks import EarlyStopping
from datetime import datetime
from tensorflow import keras
from tensorflow.keras.losses import MeanSquaredLogarithmicError
from sklearn.metrics import mean_squared_log_error
def root_mean_squared_log_error(y_true, y_pred):
return np.sqrt(mean_squared_log_error(y_true, y_pred))
mc = ModelCheckpoint(f'../Models/NN_model_test.h5', monitor='val_loss', mode='min', verbose=1, save_best_only=True)
early_stopping = EarlyStopping(
monitor='val_loss',
patience=10,
verbose=0,
mode="auto",
baseline=None,
restore_best_weights=True)
model = keras.Sequential([
keras.layers.Dense(100, activation='relu', input_dim=89),
keras.layers.LeakyReLU(500),
keras.layers.LeakyReLU(800),
keras.layers.LeakyReLU(200),
keras.layers.Dense(200, activation='relu'),
keras.layers.Dense(10, activation='relu'),
keras.layers.Dense(1, activation='relu')
])
opt = keras.optimizers.Adam(learning_rate=0.0001)
model.compile(optimizer=opt,
loss='mean_squared_logarithmic_error',
metrics=['mean_squared_logarithmic_error'])
history = model.fit(
X_train,
y_train,
batch_size=100,
epochs=5000,
validation_data=(X_test, y_test),
callbacks=[mc, early_stopping],
shuffle=True,
steps_per_epoch=3
)
results = pd.DataFrame()
y_pred = scaler.inverse_transform(model.predict(X_test))
results['Predicted'] = ((1 / y_pred) - 1).ravel()
y_actual = scaler.inverse_transform(y_test)
results['Actual'] = ((1 / y_actual) - 1).ravel()
results['Predicted'] = y_pred
results['Actual'] = y_actual
results['Difference'] = abs(results['Predicted'] - results['Actual'])
scaler.inverse_transform(model.predict(X_test))
results
results['Difference'].mean()
mean_squared_error(results['Actual'], results['Predicted'], squared=False)
root_mean_squared_log_error(results['Actual'], results['Predicted'])
df.hist(figsize=(20,20))
df['Time_Minutes'].hist(bins=50, figsize=(10,10))
df[df['S3'] < 1000].sort_values(by='S3', ascending=False)
log_cols = [
'Air_Temp',
'Lap_Number',
]
root_cols = [
'Elapsed',
'Wind_Speed',
'Track_Temp',
]
df
df
```
| github_jupyter |
<img src="results/latex.png" alt="logo" width="400px" style="float: right;">
# $\rm \textbf{Introduction to}$ ``pySYD``
```
# python imports
import pickle
import numpy as np
import matplotlib.pyplot as plt
from astropy.convolution import Box1DKernel, convolve
# pysyd-related imports
import pysyd
from pysyd import pipeline
from pysyd import utils
%matplotlib inline
```
## $\rm \textbf{Load in the star data}$
Main functions are called through ``pysyd.pipeline``. <br>
Let's load in data for our least-evolved example star: **1435467**
```
star = pipeline.load(star='1435467', CLI=False, verbose=True)
```
For now, we'll keep the verbose option set to ``True`` so that we can see
what the software is doing.
For this target, both the light curve and power
spectrum were provided and loaded in. In this scenario, the software will automatically
compute the oversampling factor (if applicable). This is reflected in the printed resolution,
which corresponds to the frequency resolution of the critically-sampled PS.
The ``star`` is an important moving part of the ``pySYD`` software, which is an instance of
the ``target.Target`` class. A new ``Target`` is created for each processed star, and contains
a whole range of information. For more details see our [API](https://pysyd.readthedocs.io/en/latest/target.html).
## $\rm \textbf{Don't know what}\,\, \nu_{max} \,\, \textbf{is for your star? Scratch that. Don't even know where to start?}$
$\rm \textbf{Don't worry, our first module can help with that!}$
```
star = utils.get_findex(star)
star.find_excess()
```
The first utility function merely sets up parameters and arrays to successfully run the module.
Let's take a look at the output figure.
```
with open('excess.pickle','rb') as file:
fig = pickle.load(file)
plt.show()
```
Ok, let's unpack.
<b><u>Top:</b></u><br>
The top left panel shows the time series data as a sanity check (but does not do any altering of the light curve). The top middle panel shows the power spectrum (PS) in log-log space. The PS is binned (red) and heavily smoothed (lime green) as a quick but efficient method to determine the approximate background contribution due to stellar granulation. The reason for this step is that the value of numax for some stars can vary greatly if these effects are not considered. The background-corrected power spectrum (BCPS) in the top right panel is the original (white) power spectrum divided by the heavily-smoothed (green) power spectrum.
<b><u>Bottom:</b></u><br>
The three panels show the frequency-resolved, collapsed autocorrelation function (ACF) computed from the BCPS using three different box (or bin) sizes. A Gaussian is fit to each ACF, where the ``trial`` yielding the highest SNR peak is adopted as our initial guess for $\nu_{\mathrm{max}}$. In this case, the measured values were all self-consistent but this will not always be the case (i.e. use your best judgement).
## $\rm \textbf{Now that we have an appoximate starting point for}\,\, \nu_{max}, \,\, \textbf{we can run the next module}$
(this routine does a majority of the heavy lifting)
```
# the conditional checks if there is an estimate for numax
# Note: numax is required for this module to run properly
if utils.check_fitbg(star):
star = utils.get_fitbg(star)
star.fit_global()
```
Note that this one provided a conditional to start. Let me explain.
Near-surface convection driving the acoustic oscillations is a stochastic process, which is why we see oscillation modes over a range of frequencies. Luckily, we know approximately how wide this region is based on ensemble analyses from <i>Kepler</i>. Using a scaling relation, we determine this width based on the value for $\nu_{\mathrm{max}}$. This region is then masked out in the beginning steps of the second module to more accurately determinie the stellar background contribution. <b> Therefore, the second routine <u><i>requires</i></u> an appoximate estimate for $\nu_{\mathrm{max}}$ (and this is where the conditional comes in).</b>
Luckily, we have three places one can provide/find such value:
1. By running the first module discussed above
2. In the 'info/star_info.csv' (this is described in more detail in our [documentation](https://pysyd.readthedocs.io/en/latest/overview.html))
3. Via command line i.e. ``--numax`` xxxx.x
```
with open('background.pickle','rb') as file:
fig = pickle.load(file)
plt.show()
```
<b><u>Top:</b></u><br>
So the first two figures should look somewhat familar from the first module (as they should because the light curve and power spectrum are exactly the same). In the second panel, we have a similar smoothed (red) power spectrum and then the binned (lime green) power spectrum. Like many other astrophysical phenomena, the stellar background consists of a combination of a single flat white-noise (horizontal blue dashed line) term and at least one red-noise component (curved blue dotted line(s)). The shaded power spectrum region delineated by the vertical orange dashed lines is exactly the power excess region we discussed above. The second and third panel are pretty similar, but the middle shows our initial guesses for these noise contributions and the right shows our final fit (i.e. the summed background total is the solid blue line). We use solar scaling relations for granulation timescales to estimate these initial guesses and as you can see, they do pretty well! Finally, the power excess is approximately Gaussian, shown by the dashed yellow line.
<b><u>Middle:</b></u><br>
The power spectrum is corrected by dividing out the best-fit background model, which means we are now ready to estimate the global asteroseismic parameters. We say the power excess is approximately Gaussian, but there is no physical reason to believe that the mode excitation is perfectly Gaussian. This is why we estimate $\nu_{\mathrm{max}}$ two ways: 1) using a heavily-smoothed power spectrum (white) and 2) by fitting a Gaussian (blue), which are both plotted together in the first panel. The middle panel shows the background-corrected PS (BCPS) centered on $\nu_{\mathrm{max}}$ and highlights some of the highest peaks (red). For acoustic oscillations that are equally spaced in frequency, the autocorrelation (ACF) is a powerful tool to quantitatively confirm the power excess. Therefore, the power in the middle figure is used to calculate the ACF shown in the third panel. Our peak corresponding to $\Delta\nu$ is highlighted, and we see harmonics at roughly half the spacing since the lowest order modes are separated by $\sim\Delta\nu/2$.
<b><u>Bottom:</b></u><br>
The first panel is zoomed in on the peak in the ACF closest to our expected $\Delta\nu$, which we fit with a Gaussian (green). Here, the red dashed line was our initial estimate based on a $\nu_{\mathrm{max}}$/$\Delta\nu$ scaling relation, so it was pretty close! Adopting the peak of the Gaussian as our value for $\Delta\nu$, we fold and stack the power spectrum in what is called an \`echelle diagram (which literally translates to ladder in French). This is shown in the middle panel, where we can see nice straight ridges corresponding to different oscillation modes. The right panel is similar to the middle plot but marganalizes over (or collapses) the y-axis, which is just another way to view this. Again, we see peaks in power that correspond to different oscillation modes.
## $\rm \textbf{So let's put this all together on another star.}$
```
star = pipeline.run(star='11618103', CLI=False)
```
Remember that by default, the verbose ouptut is ``False`` and is why we didn't see any output this time.
Let's take a look at the plots!
### $\rm \textbf{The first module:}$
```
with open('excess.pickle','rb') as file:
fig = pickle.load(file)
plt.show()
```
For this star, you might've noticed that $\nu_{\mathrm{max}}$ is lower by an order of magnitude. Additionally, the oscillations have much higher amplitudes!
Indeed these are two well-known facts for characteristics of solar-like oscillations:
1. The frequency corresponding to maximum power (or $\nu_{\mathrm{max}}$) scales linearly will the surface gravity (log$g$) of the star.
2. Mode amplitudes scale with the luminosity of the star.
### $\rm \textbf{The second module:}$
```
with open('background.pickle','rb') as file:
fig = pickle.load(file)
plt.show()
```
| github_jupyter |
# Image Classifier: Poison Ivy
Trains a model to classify a plant image as a type of poison ivy (or not).
Below we do the following:
1. Setup the training environment.
2. Load images of different poison ivy plants and look-alikes
3. Train an image classifier model.
3. Convert the model to CoreML and upload it to Skafos.
## Environment Setup
Below we ensure `CUDA 10` is installed and then use pip to install `turicreate`, `mxnet-cu100`, and `skafos` libraries.
```
# Confirm that you have CUDA 10
!nvcc --version
# Install libraries - you might need to restart the runtime after doing this
!pip install turicreate==5.4
# The wrong version of mxnet will be installed
!pip uninstall -y mxnet
# Install CUDA10-compatible version of mxnet
!pip install mxnet-cu100
# install Skafos python sdk
!pip install skafos
```
## Data Preparation and Model Training
The training data for this example are images of various plant species (some poisonous, some not), gathered and labeled by hand. One of the limitations of this space is having enough training data. Because our data is limited, if we were to try to make a more accurate model, we would need to collect and label more images for each class.
After unzipping and extracting the images, they are loaded into a Turi Create SFrame and labels are created for each image based on the path. The data is randomly split into train and test sets, where 80% of the data is used for training and 20% is used for model evaluation (if you desire). Training this model with a GPU is much faster than CPU time. By default, this runtime environment should be using a Python 3 GPU backend instance. Below, we tell Turicreate to use all available GPUs for processing.
```
# Import libraries and tell Turicreate to use all GPUs available - this may throw a warning
import urllib
import tarfile
import os
import coremltools
import turicreate as tc
tc.config.set_num_gpus(-1)
# Specify the data set download url
data_url = "https://s3.amazonaws.com/skafos.example.data/ImageClassifier/poisonPlants.tar.gz"
data_path = "poisonPlants.tar.gz"
# Pull the compressed data and extract it
retrieve = urllib.request.urlretrieve(data_url, data_path)
tar = tarfile.open(data_path)
tar.extractall()
tar.close()
# Load images - you can ignore various jpeg decode errors
data = tc.image_analysis.load_images('poisonPlants', with_path=True, ignore_failure=True)
# From the path-name, create a label column. This labels each image as the appropriate plant
data['label'] = data['path'].apply(lambda path: os.path.basename(os.path.dirname(path)))
# Make a train-test split
train_data, test_data = data.random_split(0.8)
# Train an image classification model - consider increasing max_iterations
model = tc.image_classifier.create(
dataset=train_data,
target='label',
model='resnet-50',
batch_size=4,
max_iterations=10
)
# Image Classification Training Docs:
# https://apple.github.io/turicreate/docs/api/generated/turicreate.image_classifier.create.html#turicreate.image_classifier.create
```
## Model Evaluation
```
# Let's see how the model performs on the hold out test data
predictions = model.predict(test_data)
accuracy = tc.evaluation.accuracy(test_data['label'], predictions)
print(f"Image classifier is {accuracy*100} % accurate on the testing dataset", flush=True)
```
## Model Export and Skafos Upload
- Convert the model to CoreML format so that it can run on an iOS device. Then deliver the model to your apps with **[Skafos](https://skafos.ai)**.
- If you don't already have an account, Sign Up for one **[here](https://dashboard.skafos.ai)**.
- Once you've signed up for an account, grab an API token from your account settings.
```
# Specify the CoreML model name
model_name = 'ImageClassifier'
coreml_model_name = model_name + '.mlmodel'
# Export the trained model to CoreML format
res = model.export_coreml(coreml_model_name)
import skafos
from skafos import models
import os
# Set your API Token first for repeated use
os.environ["SKAFOS_API_TOKEN"] = "<YOUR-SKAFOS-API-TOKEN>"
skafos.summary()
# You can retrieve this info with skafos.summary()
org_name = "<YOUR-SKAFOS-ORG-NAME>" # Example: "mike-gmail-com-467h2"
app_name = "<YOUR-SKAFOS-APP-NAME>" # Example: "ImageClassification-App"
model_name = "<YOUR-MODEL-NAME>" # Example: "ImageClassificationModel"
# Upload model version to Skafos
model_upload_result = models.upload_version(
files="ImageClassifier.mlmodel",
org_name=org_name,
app_name=app_name,
model_name=model_name
)
```
| github_jupyter |
```
import logging, scipy, dask, pandas, numpy, time, typing, asyncio
import dask.dataframe as ddf
import dask.bag as dbag
import dask.distributed as dask_distributed
from dask.distributed import Client
import distributed.diagnostics as daskdia
from sklearn.cluster import KMeans
from sklearn.externals import joblib
numpy.random.seed(201)
k = 100
representationThr = 3
# client = dask_distributed.Client()
# StreamZ Dask
client = await Client(processes=False)
# StreamZ Dask Async
# client = await Client(processes=False, asynchronous=True)
classes = [{'label': f'Class #{i}', 'mu': numpy.random.random() * 10, 'sigma': numpy.random.random() * 5} for i in range(5)]
def nextExample():
cl = classes[numpy.random.randint(0, len(classes))]
item = [float(numpy.random.normal(loc=cl['mu'], scale=cl['sigma'])) for i in range(40)]
return {'label': cl['label'], 'item': item,}
def limit(stream, limit):
msg = yield
for target_list in range(limit):
yield next(stream)
# %%timeit
# examples = dbag.from_sequence(limit(mkStream(), 10000))
# daskdia.progress(examples)
from dask.distributed import Client
from tornado.ioloop import IOLoop
import streamz
from streamz.dataframe import DataFrame as StreamzDataframe
exStream = streamz.Stream()
# exStream = streamz.Stream(asynchronous=True)
def aggOffile(accDF, example):
if example is None:
return accDF
return accDF.append(pandas.DataFrame({'label': [example['label']], 'item': [example['item']]}))
offlineAgg = exStream.accumulate(aggOffile, start=pandas.DataFrame({'label': [], 'item': []}), returns_state=False)
# sdf = StreamzDataframe(source,)
# # buffer(5)
# dfStream = source.map(lambda ex: pandas.dataframe(data=ex))
# dfStream.accumulate().map(lambda ex: pandas.dataframe(data=ex)).scatter().map(increment).gather().sink()
print(offlineAgg)
offlineAgg.sink(print)
exStream.emit(nextExample())
%%timeit
for i in range(10000):
exStream.emit(nextExample())
async def generate():
for i in range(10000):
await asyncio.sleep(10)
await exStream.emit(nextExample())
await exStream.emit(None)
# asyncio.get_loop().run(generate)
class Cluster():
def __init__(self, center=[]):
self.center = center
self.label = None
self.n = 0
self.lastExapleTMS = 0
self.sumDistance = 0
self.maxDistance = 0
def dist(self, item):
return scipy.spatial.distance.euclidean(self.center, item)
def addExample(self, item, distance=None):
self.n += 1
if example.timestamp > self.lastExapleTMS:
self.lastExapleTMS = example.timestamp
if distance == None:
distance = self.dist(item)
self.sumDistance += distance
if distance > self.maxDistance:
self.maxDistance = distance
def clustering(examples, label=None):
representationThr = 3
kmeans = KMeans( n_clusters = min(k, int(len(examples) / (3 * representationThr))) )
with joblib.parallel_backend('dask'):
kmeans.fit(examples)
return [Cluster(center=centroid, label=label) for centroid in kmeans.cluster_centers_]
def closestCluster(item, clusters):
return min([ (cl, cl.dist(item)) for cl in clusters ], key=lambda x: x[1])
def offline(label, group):
representationThr = 3
clusters = clustering(group, label)
for ex in group:
nearCl, dist = closestCluster(ex, clusters)
nearCl.addExample(ex)
return [cluster for cluster in clusters if cluster.n > representationThr]
def group(i, downStream=streamz.Stream()):
_, df = i
return [ downStream.emit( (l, df[g['item']]) ) for l, g in df.groupby('label') ]
def mapOffline(i):
label, group = i
return offline(label, group)
trainingTrigger = streamz.Stream()
groups = streamz.Stream()
L = []
def nnnn(i):
L.append(i)
trainingTrigger.zip(offlineAgg).sink(nnnn)
# grouped = trainingTrigger.zip(offlineAgg).map(group, downStream=groups)
# grouped.sink(lambda x: print('grouped {x!r} {t}'.format(x=x, t=type(x))))
# trained = groups.map(mapOffline)
# trained = groups.scatter().map(mapOffline).gather()
# trained.sink(lambda x: print('trained {x!r} {t}'.format(x=x, t=type(x))))
trainingTrigger.emit(True)
exStream.emit(nextExample())
trained.visualize(rankdir='LR')
# trainingTrigger.emit(True)
# trained.sink(print)
L
from streamz.dataframe import Random, DataFrame
%matplotlib inline
source = Random(freq='5ms', interval='100ms')
r = source.x.sum()
r.visualize()
sdf = (source - 0.5).cumsum()
sdf.tail()
p = (DataFrame({'raw': sdf.x,
'smooth': sdf.x.rolling('100ms').mean(),
'very-smooth': sdf.x.rolling('500ms').mean()})
.plot(width=700)
)
import streamz
def increment(x):
return x + 1
def decrement(x):
return x - 1
s = streamz.Stream()
a = s.map(increment).sink(print)
b = s.map(decrement).sink(print)
b.visualize(rankdir='LR')
```
| github_jupyter |
<a href="https://www.bigdatauniversity.com"><img src="https://ibm.box.com/shared/static/cw2c7r3o20w9zn8gkecaeyjhgw3xdgbj.png" width="400" align="center"></a>
<h1><center>Hierarchical Clustering</center></h1>
Welcome to Lab of Hierarchical Clustering with Python using Scipy and Scikit-learn package.
<h1>Table of contents</h1>
<div class="alert alert-block alert-info" style="margin-top: 20px">
<ol>
<li><a href="#hierarchical_agglomerative">Hierarchical Clustering - Agglomerative</a></li>
<ol>
<li><a href="#generating_data">Generating Random Data</a></li>
<li><a href="#agglomerative_clustering">Agglomerative Clustering</a></li>
<li><a href="#dendrogram">Dendrogram Associated for the Agglomerative Hierarchical Clustering</a></li>
</ol>
<li><a href="#clustering_vehicle_dataset">Clustering on the Vehicle Dataset</a></li>
<ol>
<li><a href="#data_cleaning">Data Cleaning</a></li>
<li><a href="#clustering_using_scipy">Clustering Using Scipy</a></li>
<li><a href="#clustering_using_skl">Clustering using scikit-learn</a></li>
</ol>
</ol>
</div>
<br>
<hr>
<h1 id="hierarchical_agglomerative">Hierarchical Clustering - Agglomerative</h1>
We will be looking at a clustering technique, which is <b>Agglomerative Hierarchical Clustering</b>. Remember that agglomerative is the bottom up approach. <br> <br>
In this lab, we will be looking at Agglomerative clustering, which is more popular than Divisive clustering. <br> <br>
We will also be using Complete Linkage as the Linkage Criteria. <br>
<b> <i> NOTE: You can also try using Average Linkage wherever Complete Linkage would be used to see the difference! </i> </b>
```
import numpy as np
import pandas as pd
from scipy import ndimage
from scipy.cluster import hierarchy
from scipy.spatial import distance_matrix
from matplotlib import pyplot as plt
from sklearn import manifold, datasets
from sklearn.cluster import AgglomerativeClustering
from sklearn.datasets.samples_generator import make_blobs
%matplotlib inline
```
<hr>
<h3 id="generating_data">Generating Random Data</h3>
We will be generating a set of data using the <b>make_blobs</b> class. <br> <br>
Input these parameters into make_blobs:
<ul>
<li> <b>n_samples</b>: The total number of points equally divided among clusters. </li>
<ul> <li> Choose a number from 10-1500 </li> </ul>
<li> <b>centers</b>: The number of centers to generate, or the fixed center locations. </li>
<ul> <li> Choose arrays of x,y coordinates for generating the centers. Have 1-10 centers (ex. centers=[[1,1], [2,5]]) </li> </ul>
<li> <b>cluster_std</b>: The standard deviation of the clusters. The larger the number, the further apart the clusters</li>
<ul> <li> Choose a number between 0.5-1.5 </li> </ul>
</ul> <br>
Save the result to <b>X1</b> and <b>y1</b>.
```
X1, y1 = make_blobs(n_samples=50, centers=[[4,4], [-2, -1], [1, 1], [10,4]], cluster_std=0.9)
```
Plot the scatter plot of the randomly generated data
```
plt.scatter(X1[:, 0], X1[:, 1], marker='o')
```
<hr>
<h3 id="agglomerative_clustering">Agglomerative Clustering</h3>
We will start by clustering the random data points we just created.
The <b> Agglomerative Clustering </b> class will require two inputs:
<ul>
<li> <b>n_clusters</b>: The number of clusters to form as well as the number of centroids to generate. </li>
<ul> <li> Value will be: 4 </li> </ul>
<li> <b>linkage</b>: Which linkage criterion to use. The linkage criterion determines which distance to use between sets of observation. The algorithm will merge the pairs of cluster that minimize this criterion. </li>
<ul>
<li> Value will be: 'complete' </li>
<li> <b>Note</b>: It is recommended you try everything with 'average' as well </li>
</ul>
</ul> <br>
Save the result to a variable called <b> agglom </b>
```
agglom = AgglomerativeClustering(n_clusters = 4, linkage = 'average')
```
Fit the model with <b> X2 </b> and <b> y2 </b> from the generated data above.
```
agglom.fit(X1,y1)
```
Run the following code to show the clustering! <br>
Remember to read the code and comments to gain more understanding on how the plotting works.
```
# Create a figure of size 6 inches by 4 inches.
plt.figure(figsize=(6,4))
# These two lines of code are used to scale the data points down,
# Or else the data points will be scattered very far apart.
# Create a minimum and maximum range of X1.
x_min, x_max = np.min(X1, axis=0), np.max(X1, axis=0)
# Get the average distance for X1.
X1 = (X1 - x_min) / (x_max - x_min)
# This loop displays all of the datapoints.
for i in range(X1.shape[0]):
# Replace the data points with their respective cluster value
# (ex. 0) and is color coded with a colormap (plt.cm.spectral)
plt.text(X1[i, 0], X1[i, 1], str(y1[i]),
color=plt.cm.nipy_spectral(agglom.labels_[i] / 10.),
fontdict={'weight': 'bold', 'size': 9})
# Remove the x ticks, y ticks, x and y axis
plt.xticks([])
plt.yticks([])
#plt.axis('off')
# Display the plot of the original data before clustering
plt.scatter(X1[:, 0], X1[:, 1], marker='.')
# Display the plot
plt.show()
```
<h3 id="dendrogram">Dendrogram Associated for the Agglomerative Hierarchical Clustering</h3>
Remember that a <b>distance matrix</b> contains the <b> distance from each point to every other point of a dataset </b>. <br>
Use the function <b> distance_matrix, </b> which requires <b>two inputs</b>. Use the Feature Matrix, <b> X2 </b> as both inputs and save the distance matrix to a variable called <b> dist_matrix </b> <br> <br>
Remember that the distance values are symmetric, with a diagonal of 0's. This is one way of making sure your matrix is correct. <br> (print out dist_matrix to make sure it's correct)
```
dist_matrix = distance_matrix(X1,X1)
print(dist_matrix)
```
Using the <b> linkage </b> class from hierarchy, pass in the parameters:
<ul>
<li> The distance matrix </li>
<li> 'complete' for complete linkage </li>
</ul> <br>
Save the result to a variable called <b> Z </b>
```
Z = hierarchy.linkage(dist_matrix, 'complete')
```
A Hierarchical clustering is typically visualized as a dendrogram as shown in the following cell. Each merge is represented by a horizontal line. The y-coordinate of the horizontal line is the similarity of the two clusters that were merged, where cities are viewed as singleton clusters.
By moving up from the bottom layer to the top node, a dendrogram allows us to reconstruct the history of merges that resulted in the depicted clustering.
Next, we will save the dendrogram to a variable called <b>dendro</b>. In doing this, the dendrogram will also be displayed.
Using the <b> dendrogram </b> class from hierarchy, pass in the parameter:
<ul> <li> Z </li> </ul>
```
dendro = hierarchy.dendrogram(Z)
```
## Practice
We used **complete** linkage for our case, change it to **average** linkage to see how the dendogram changes.
```
Z = hierarchy.linkage(dist_matrix, 'average')
dendro = hierarchy.dendrogram(Z)
```
Double-click **here** for the solution.
<!-- Your answer is below:
Z = hierarchy.linkage(dist_matrix, 'average')
dendro = hierarchy.dendrogram(Z)
-->
<hr>
<h1 id="clustering_vehicle_dataset">Clustering on Vehicle dataset</h1>
Imagine that an automobile manufacturer has developed prototypes for a new vehicle. Before introducing the new model into its range, the manufacturer wants to determine which existing vehicles on the market are most like the prototypes--that is, how vehicles can be grouped, which group is the most similar with the model, and therefore which models they will be competing against.
Our objective here, is to use clustering methods, to find the most distinctive clusters of vehicles. It will summarize the existing vehicles and help manufacturers to make decision about the supply of new models.
### Download data
To download the data, we will use **`!wget`** to download it from IBM Object Storage.
**Did you know?** When it comes to Machine Learning, you will likely be working with large datasets. As a business, where can you host your data? IBM is offering a unique opportunity for businesses, with 10 Tb of IBM Cloud Object Storage: [Sign up now for free](http://cocl.us/ML0101EN-IBM-Offer-CC)
```
!wget -O cars_clus.csv https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-ML0101EN-Coursera/labs/Data_files/cars_clus.csv
```
## Read data
lets read dataset to see what features the manufacturer has collected about the existing models.
```
filename = 'cars_clus.csv'
#Read csv
pdf = pd.read_csv(filename)
print ("Shape of dataset: ", pdf.shape)
pdf.head(5)
```
The feature sets include price in thousands (price), engine size (engine_s), horsepower (horsepow), wheelbase (wheelbas), width (width), length (length), curb weight (curb_wgt), fuel capacity (fuel_cap) and fuel efficiency (mpg).
<h2 id="data_cleaning">Data Cleaning</h2>
lets simply clear the dataset by dropping the rows that have null value:
```
print ("Shape of dataset before cleaning: ", pdf.size)
pdf[[ 'sales', 'resale', 'type', 'price', 'engine_s',
'horsepow', 'wheelbas', 'width', 'length', 'curb_wgt', 'fuel_cap',
'mpg', 'lnsales']] = pdf[['sales', 'resale', 'type', 'price', 'engine_s',
'horsepow', 'wheelbas', 'width', 'length', 'curb_wgt', 'fuel_cap',
'mpg', 'lnsales']].apply(pd.to_numeric, errors='coerce')
pdf = pdf.dropna()
pdf = pdf.reset_index(drop=True)
print ("Shape of dataset after cleaning: ", pdf.size)
pdf.head(5)
```
### Feature selection
Lets select our feature set:
```
featureset = pdf[['engine_s', 'horsepow', 'wheelbas', 'width', 'length', 'curb_wgt', 'fuel_cap', 'mpg']]
```
### Normalization
Now we can normalize the feature set. **MinMaxScaler** transforms features by scaling each feature to a given range. It is by default (0, 1). That is, this estimator scales and translates each feature individually such that it is between zero and one.
```
from sklearn.preprocessing import MinMaxScaler
x = featureset.values #returns a numpy array
min_max_scaler = MinMaxScaler()
feature_mtx = min_max_scaler.fit_transform(x)
feature_mtx [0:5]
```
<h2 id="clustering_using_scipy">Clustering using Scipy</h2>
In this part we use Scipy package to cluster the dataset:
First, we calculate the distance matrix.
```
import scipy
leng = feature_mtx.shape[0]
D = scipy.zeros([leng,leng])
for i in range(leng):
for j in range(leng):
D[i,j] = scipy.spatial.distance.euclidean(feature_mtx[i], feature_mtx[j])
```
In agglomerative clustering, at each iteration, the algorithm must update the distance matrix to reflect the distance of the newly formed cluster with the remaining clusters in the forest.
The following methods are supported in Scipy for calculating the distance between the newly formed cluster and each:
```
- single
- complete
- average
- weighted
- centroid
```
We use **complete** for our case, but feel free to change it to see how the results change.
```
import pylab
import scipy.cluster.hierarchy
Z = hierarchy.linkage(D, 'complete')
```
Essentially, Hierarchical clustering does not require a pre-specified number of clusters. However, in some applications we want a partition of disjoint clusters just as in flat clustering.
So you can use a cutting line:
```
from scipy.cluster.hierarchy import fcluster
max_d = 3
clusters = fcluster(Z, max_d, criterion='distance')
clusters
```
Also, you can determine the number of clusters directly:
```
from scipy.cluster.hierarchy import fcluster
k = 5
clusters = fcluster(Z, k, criterion='maxclust')
clusters
```
Now, plot the dendrogram:
```
fig = pylab.figure(figsize=(18,50))
def llf(id):
return '[%s %s %s]' % (pdf['manufact'][id], pdf['model'][id], int(float(pdf['type'][id])) )
dendro = hierarchy.dendrogram(Z, leaf_label_func=llf, leaf_rotation=0, leaf_font_size =12, orientation = 'right')
```
<h2 id="clustering_using_skl">Clustering using scikit-learn</h2>
Lets redo it again, but this time using scikit-learn package:
```
dist_matrix = distance_matrix(feature_mtx,feature_mtx)
print(dist_matrix)
```
Now, we can use the 'AgglomerativeClustering' function from scikit-learn library to cluster the dataset. The AgglomerativeClustering performs a hierarchical clustering using a bottom up approach. The linkage criteria determines the metric used for the merge strategy:
- Ward minimizes the sum of squared differences within all clusters. It is a variance-minimizing approach and in this sense is similar to the k-means objective function but tackled with an agglomerative hierarchical approach.
- Maximum or complete linkage minimizes the maximum distance between observations of pairs of clusters.
- Average linkage minimizes the average of the distances between all observations of pairs of clusters.
```
agglom = AgglomerativeClustering(n_clusters = 6, linkage = 'complete')
agglom.fit(feature_mtx)
agglom.labels_
```
And, we can add a new field to our dataframe to show the cluster of each row:
```
pdf['cluster_'] = agglom.labels_
pdf.head()
import matplotlib.cm as cm
n_clusters = max(agglom.labels_)+1
colors = cm.rainbow(np.linspace(0, 1, n_clusters))
cluster_labels = list(range(0, n_clusters))
# Create a figure of size 6 inches by 4 inches.
plt.figure(figsize=(16,14))
for color, label in zip(colors, cluster_labels):
subset = pdf[pdf.cluster_ == label]
for i in subset.index:
plt.text(subset.horsepow[i], subset.mpg[i],str(subset['model'][i]), rotation=25)
plt.scatter(subset.horsepow, subset.mpg, s= subset.price*10, c=color, label='cluster'+str(label),alpha=0.5)
# plt.scatter(subset.horsepow, subset.mpg)
plt.legend()
plt.title('Clusters')
plt.xlabel('horsepow')
plt.ylabel('mpg')
```
As you can see, we are seeing the distribution of each cluster using the scatter plot, but it is not very clear where is the centroid of each cluster. Moreover, there are 2 types of vehicles in our dataset, "truck" (value of 1 in the type column) and "car" (value of 1 in the type column). So, we use them to distinguish the classes, and summarize the cluster. First we count the number of cases in each group:
```
pdf.groupby(['cluster_','type'])['cluster_'].count()
```
Now we can look at the characteristics of each cluster:
```
agg_cars = pdf.groupby(['cluster_','type'])['horsepow','engine_s','mpg','price'].mean()
agg_cars
```
It is obvious that we have 3 main clusters with the majority of vehicles in those.
**Cars**:
- Cluster 1: with almost high mpg, and low in horsepower.
- Cluster 2: with good mpg and horsepower, but higher price than average.
- Cluster 3: with low mpg, high horsepower, highest price.
**Trucks**:
- Cluster 1: with almost highest mpg among trucks, and lowest in horsepower and price.
- Cluster 2: with almost low mpg and medium horsepower, but higher price than average.
- Cluster 3: with good mpg and horsepower, low price.
Please notice that we did not use **type** , and **price** of cars in the clustering process, but Hierarchical clustering could forge the clusters and discriminate them with quite high accuracy.
```
plt.figure(figsize=(16,10))
for color, label in zip(colors, cluster_labels):
subset = agg_cars.loc[(label,),]
for i in subset.index:
plt.text(subset.loc[i][0]+5, subset.loc[i][2], 'type='+str(int(i)) + ', price='+str(int(subset.loc[i][3]))+'k')
plt.scatter(subset.horsepow, subset.mpg, s=subset.price*20, c=color, label='cluster'+str(label))
plt.legend()
plt.title('Clusters')
plt.xlabel('horsepow')
plt.ylabel('mpg')
```
<h2>Want to learn more?</h2>
IBM SPSS Modeler is a comprehensive analytics platform that has many machine learning algorithms. It has been designed to bring predictive intelligence to decisions made by individuals, by groups, by systems – by your enterprise as a whole. A free trial is available through this course, available here: <a href="http://cocl.us/ML0101EN-SPSSModeler">SPSS Modeler</a>
Also, you can use Watson Studio to run these notebooks faster with bigger datasets. Watson Studio is IBM's leading cloud solution for data scientists, built by data scientists. With Jupyter notebooks, RStudio, Apache Spark and popular libraries pre-packaged in the cloud, Watson Studio enables data scientists to collaborate on their projects without having to install anything. Join the fast-growing community of Watson Studio users today with a free account at <a href="https://cocl.us/ML0101EN_DSX">Watson Studio</a>
<h3>Thanks for completing this lesson!</h3>
<h4>Author: <a href="https://ca.linkedin.com/in/saeedaghabozorgi">Saeed Aghabozorgi</a></h4>
<p><a href="https://ca.linkedin.com/in/saeedaghabozorgi">Saeed Aghabozorgi</a>, PhD is a Data Scientist in IBM with a track record of developing enterprise level applications that substantially increases clients’ ability to turn data into actionable knowledge. He is a researcher in data mining field and expert in developing advanced analytic methods like machine learning and statistical modelling on large datasets.</p>
| Date (YYYY-MM-DD) | Version | Changed By | Change Description |
| ----------------- | ------- | ---------- | --------------------- |
| 2020-08-04 | 0 | Nayef | Upload file to Gitlab |
| | | | |
<hr>
<p>Copyright © 2018 <a href="https://cocl.us/DX0108EN_CC">Cognitive Class</a>. This notebook and its source code are released under the terms of the <a href="https://bigdatauniversity.com/mit-license/">MIT License</a>.</p>
| github_jupyter |
# Autonomous driving - Car detection
Welcome to your week 3 programming assignment. You will learn about object detection using the very powerful YOLO model. Many of the ideas in this notebook are described in the two YOLO papers: [Redmon et al., 2016](https://arxiv.org/abs/1506.02640) and [Redmon and Farhadi, 2016](https://arxiv.org/abs/1612.08242).
**You will learn to**:
- Use object detection on a car detection dataset
- Deal with bounding boxes
## <font color='darkblue'>Updates</font>
#### If you were working on the notebook before this update...
* The current notebook is version "3a".
* You can find your original work saved in the notebook with the previous version name ("v3")
* To view the file directory, go to the menu "File->Open", and this will open a new tab that shows the file directory.
#### List of updates
* Clarified "YOLO" instructions preceding the code.
* Added details about anchor boxes.
* Added explanation of how score is calculated.
* `yolo_filter_boxes`: added additional hints. Clarify syntax for argmax and max.
* `iou`: clarify instructions for finding the intersection.
* `iou`: give variable names for all 8 box vertices, for clarity. Adds `width` and `height` variables for clarity.
* `iou`: add test cases to check handling of non-intersecting boxes, intersection at vertices, or intersection at edges.
* `yolo_non_max_suppression`: clarify syntax for tf.image.non_max_suppression and keras.gather.
* "convert output of the model to usable bounding box tensors": Provides a link to the definition of `yolo_head`.
* `predict`: hint on calling sess.run.
* Spelling, grammar, wording and formatting updates to improve clarity.
## Import libraries
Run the following cell to load the packages and dependencies that you will find useful as you build the object detector!
```
import argparse
import os
import matplotlib.pyplot as plt
from matplotlib.pyplot import imshow
import scipy.io
import scipy.misc
import numpy as np
import pandas as pd
import PIL
import tensorflow as tf
from keras import backend as K
from keras.layers import Input, Lambda, Conv2D
from keras.models import load_model, Model
from yolo_utils import read_classes, read_anchors, generate_colors, preprocess_image, draw_boxes, scale_boxes
from yad2k.models.keras_yolo import yolo_head, yolo_boxes_to_corners, preprocess_true_boxes, yolo_loss, yolo_body
%matplotlib inline
```
**Important Note**: As you can see, we import Keras's backend as K. This means that to use a Keras function in this notebook, you will need to write: `K.function(...)`.
## 1 - Problem Statement
You are working on a self-driving car. As a critical component of this project, you'd like to first build a car detection system. To collect data, you've mounted a camera to the hood (meaning the front) of the car, which takes pictures of the road ahead every few seconds while you drive around.
<center>
<video width="400" height="200" src="nb_images/road_video_compressed2.mp4" type="video/mp4" controls>
</video>
</center>
<caption><center> Pictures taken from a car-mounted camera while driving around Silicon Valley. <br> We thank [drive.ai](htps://www.drive.ai/) for providing this dataset.
</center></caption>
You've gathered all these images into a folder and have labelled them by drawing bounding boxes around every car you found. Here's an example of what your bounding boxes look like.
<img src="nb_images/box_label.png" style="width:500px;height:250;">
<caption><center> <u> **Figure 1** </u>: **Definition of a box**<br> </center></caption>
If you have 80 classes that you want the object detector to recognize, you can represent the class label $c$ either as an integer from 1 to 80, or as an 80-dimensional vector (with 80 numbers) one component of which is 1 and the rest of which are 0. The video lectures had used the latter representation; in this notebook, we will use both representations, depending on which is more convenient for a particular step.
In this exercise, you will learn how "You Only Look Once" (YOLO) performs object detection, and then apply it to car detection. Because the YOLO model is very computationally expensive to train, we will load pre-trained weights for you to use.
## 2 - YOLO
"You Only Look Once" (YOLO) is a popular algorithm because it achieves high accuracy while also being able to run in real-time. This algorithm "only looks once" at the image in the sense that it requires only one forward propagation pass through the network to make predictions. After non-max suppression, it then outputs recognized objects together with the bounding boxes.
### 2.1 - Model details
#### Inputs and outputs
- The **input** is a batch of images, and each image has the shape (m, 608, 608, 3)
- The **output** is a list of bounding boxes along with the recognized classes. Each bounding box is represented by 6 numbers $(p_c, b_x, b_y, b_h, b_w, c)$ as explained above. If you expand $c$ into an 80-dimensional vector, each bounding box is then represented by 85 numbers.
#### Anchor Boxes
* Anchor boxes are chosen by exploring the training data to choose reasonable height/width ratios that represent the different classes. For this assignment, 5 anchor boxes were chosen for you (to cover the 80 classes), and stored in the file './model_data/yolo_anchors.txt'
* The dimension for anchor boxes is the second to last dimension in the encoding: $(m, n_H,n_W,anchors,classes)$.
* The YOLO architecture is: IMAGE (m, 608, 608, 3) -> DEEP CNN -> ENCODING (m, 19, 19, 5, 85).
#### Encoding
Let's look in greater detail at what this encoding represents.
<img src="nb_images/architecture.png" style="width:700px;height:400;">
<caption><center> <u> **Figure 2** </u>: **Encoding architecture for YOLO**<br> </center></caption>
If the center/midpoint of an object falls into a grid cell, that grid cell is responsible for detecting that object.
Since we are using 5 anchor boxes, each of the 19 x19 cells thus encodes information about 5 boxes. Anchor boxes are defined only by their width and height.
For simplicity, we will flatten the last two last dimensions of the shape (19, 19, 5, 85) encoding. So the output of the Deep CNN is (19, 19, 425).
<img src="nb_images/flatten.png" style="width:700px;height:400;">
<caption><center> <u> **Figure 3** </u>: **Flattening the last two last dimensions**<br> </center></caption>
#### Class score
Now, for each box (of each cell) we will compute the following element-wise product and extract a probability that the box contains a certain class.
The class score is $score_{c,i} = p_{c} \times c_{i}$: the probability that there is an object $p_{c}$ times the probability that the object is a certain class $c_{i}$.
<img src="nb_images/probability_extraction.png" style="width:700px;height:400;">
<caption><center> <u> **Figure 4** </u>: **Find the class detected by each box**<br> </center></caption>
##### Example of figure 4
* In figure 4, let's say for box 1 (cell 1), the probability that an object exists is $p_{1}=0.60$. So there's a 60% chance that an object exists in box 1 (cell 1).
* The probability that the object is the class "category 3 (a car)" is $c_{3}=0.73$.
* The score for box 1 and for category "3" is $score_{1,3}=0.60 \times 0.73 = 0.44$.
* Let's say we calculate the score for all 80 classes in box 1, and find that the score for the car class (class 3) is the maximum. So we'll assign the score 0.44 and class "3" to this box "1".
#### Visualizing classes
Here's one way to visualize what YOLO is predicting on an image:
- For each of the 19x19 grid cells, find the maximum of the probability scores (taking a max across the 80 classes, one maximum for each of the 5 anchor boxes).
- Color that grid cell according to what object that grid cell considers the most likely.
Doing this results in this picture:
<img src="nb_images/proba_map.png" style="width:300px;height:300;">
<caption><center> <u> **Figure 5** </u>: Each one of the 19x19 grid cells is colored according to which class has the largest predicted probability in that cell.<br> </center></caption>
Note that this visualization isn't a core part of the YOLO algorithm itself for making predictions; it's just a nice way of visualizing an intermediate result of the algorithm.
#### Visualizing bounding boxes
Another way to visualize YOLO's output is to plot the bounding boxes that it outputs. Doing that results in a visualization like this:
<img src="nb_images/anchor_map.png" style="width:200px;height:200;">
<caption><center> <u> **Figure 6** </u>: Each cell gives you 5 boxes. In total, the model predicts: 19x19x5 = 1805 boxes just by looking once at the image (one forward pass through the network)! Different colors denote different classes. <br> </center></caption>
#### Non-Max suppression
In the figure above, we plotted only boxes for which the model had assigned a high probability, but this is still too many boxes. You'd like to reduce the algorithm's output to a much smaller number of detected objects.
To do so, you'll use **non-max suppression**. Specifically, you'll carry out these steps:
- Get rid of boxes with a low score (meaning, the box is not very confident about detecting a class; either due to the low probability of any object, or low probability of this particular class).
- Select only one box when several boxes overlap with each other and detect the same object.
### 2.2 - Filtering with a threshold on class scores
You are going to first apply a filter by thresholding. You would like to get rid of any box for which the class "score" is less than a chosen threshold.
The model gives you a total of 19x19x5x85 numbers, with each box described by 85 numbers. It is convenient to rearrange the (19,19,5,85) (or (19,19,425)) dimensional tensor into the following variables:
- `box_confidence`: tensor of shape $(19 \times 19, 5, 1)$ containing $p_c$ (confidence probability that there's some object) for each of the 5 boxes predicted in each of the 19x19 cells.
- `boxes`: tensor of shape $(19 \times 19, 5, 4)$ containing the midpoint and dimensions $(b_x, b_y, b_h, b_w)$ for each of the 5 boxes in each cell.
- `box_class_probs`: tensor of shape $(19 \times 19, 5, 80)$ containing the "class probabilities" $(c_1, c_2, ... c_{80})$ for each of the 80 classes for each of the 5 boxes per cell.
#### **Exercise**: Implement `yolo_filter_boxes()`.
1. Compute box scores by doing the elementwise product as described in Figure 4 ($p \times c$).
The following code may help you choose the right operator:
```python
a = np.random.randn(19*19, 5, 1)
b = np.random.randn(19*19, 5, 80)
c = a * b # shape of c will be (19*19, 5, 80)
```
This is an example of **broadcasting** (multiplying vectors of different sizes).
2. For each box, find:
- the index of the class with the maximum box score
- the corresponding box score
**Useful references**
* [Keras argmax](https://keras.io/backend/#argmax)
* [Keras max](https://keras.io/backend/#max)
**Additional Hints**
* For the `axis` parameter of `argmax` and `max`, if you want to select the **last** axis, one way to do so is to set `axis=-1`. This is similar to Python array indexing, where you can select the last position of an array using `arrayname[-1]`.
* Applying `max` normally collapses the axis for which the maximum is applied. `keepdims=False` is the default option, and allows that dimension to be removed. We don't need to keep the last dimension after applying the maximum here.
* Even though the documentation shows `keras.backend.argmax`, use `keras.argmax`. Similarly, use `keras.max`.
3. Create a mask by using a threshold. As a reminder: `([0.9, 0.3, 0.4, 0.5, 0.1] < 0.4)` returns: `[False, True, False, False, True]`. The mask should be True for the boxes you want to keep.
4. Use TensorFlow to apply the mask to `box_class_scores`, `boxes` and `box_classes` to filter out the boxes we don't want. You should be left with just the subset of boxes you want to keep.
**Useful reference**:
* [boolean mask](https://www.tensorflow.org/api_docs/python/tf/boolean_mask)
**Additional Hints**:
* For the `tf.boolean_mask`, we can keep the default `axis=None`.
**Reminder**: to call a Keras function, you should use `K.function(...)`.
```
# GRADED FUNCTION: yolo_filter_boxes
def yolo_filter_boxes(box_confidence, boxes, box_class_probs, threshold = .6):
"""Filters YOLO boxes by thresholding on object and class confidence.
Arguments:
box_confidence -- tensor of shape (19, 19, 5, 1)
boxes -- tensor of shape (19, 19, 5, 4)
box_class_probs -- tensor of shape (19, 19, 5, 80)
threshold -- real value, if [ highest class probability score < threshold], then get rid of the corresponding box
Returns:
scores -- tensor of shape (None,), containing the class probability score for selected boxes
boxes -- tensor of shape (None, 4), containing (b_x, b_y, b_h, b_w) coordinates of selected boxes
classes -- tensor of shape (None,), containing the index of the class detected by the selected boxes
Note: "None" is here because you don't know the exact number of selected boxes, as it depends on the threshold.
For example, the actual output size of scores would be (10,) if there are 10 boxes.
"""
# Step 1: Compute box scores
### START CODE HERE ### (≈ 1 line)
box_scores = box_confidence * box_class_probs
### END CODE HERE ###
# Step 2: Find the box_classes using the max box_scores, keep track of the corresponding score
### START CODE HERE ### (≈ 2 lines)
box_classes = K.argmax(box_scores, axis = -1)
box_class_scores = K.max(box_scores, axis = -1)
### END CODE HERE ###
# Step 3: Create a filtering mask based on "box_class_scores" by using "threshold". The mask should have the
# same dimension as box_class_scores, and be True for the boxes you want to keep (with probability >= threshold)
### START CODE HERE ### (≈ 1 line)
filtering_mask = ((box_class_scores) >= threshold)
### END CODE HERE ###
# Step 4: Apply the mask to box_class_scores, boxes and box_classes
### START CODE HERE ### (≈ 3 lines)
scores = tf.boolean_mask(box_class_scores, filtering_mask, name='boolean_mask')
boxes = tf.boolean_mask(boxes, filtering_mask, name='boolean_mask')
classes = tf.boolean_mask(box_classes, filtering_mask, name='boolean_mask')
### END CODE HERE ###
return scores, boxes, classes
with tf.Session() as test_a:
box_confidence = tf.random_normal([19, 19, 5, 1], mean=1, stddev=4, seed = 1)
boxes = tf.random_normal([19, 19, 5, 4], mean=1, stddev=4, seed = 1)
box_class_probs = tf.random_normal([19, 19, 5, 80], mean=1, stddev=4, seed = 1)
scores, boxes, classes = yolo_filter_boxes(box_confidence, boxes, box_class_probs, threshold = 0.5)
print("scores[2] = " + str(scores[2].eval()))
print("boxes[2] = " + str(boxes[2].eval()))
print("classes[2] = " + str(classes[2].eval()))
print("scores.shape = " + str(scores.shape))
print("boxes.shape = " + str(boxes.shape))
print("classes.shape = " + str(classes.shape))
```
**Expected Output**:
<table>
<tr>
<td>
**scores[2]**
</td>
<td>
10.7506
</td>
</tr>
<tr>
<td>
**boxes[2]**
</td>
<td>
[ 8.42653275 3.27136683 -0.5313437 -4.94137383]
</td>
</tr>
<tr>
<td>
**classes[2]**
</td>
<td>
7
</td>
</tr>
<tr>
<td>
**scores.shape**
</td>
<td>
(?,)
</td>
</tr>
<tr>
<td>
**boxes.shape**
</td>
<td>
(?, 4)
</td>
</tr>
<tr>
<td>
**classes.shape**
</td>
<td>
(?,)
</td>
</tr>
</table>
**Note** In the test for `yolo_filter_boxes`, we're using random numbers to test the function. In real data, the `box_class_probs` would contain non-zero values between 0 and 1 for the probabilities. The box coordinates in `boxes` would also be chosen so that lengths and heights are non-negative.
### 2.3 - Non-max suppression ###
Even after filtering by thresholding over the class scores, you still end up with a lot of overlapping boxes. A second filter for selecting the right boxes is called non-maximum suppression (NMS).
<img src="nb_images/non-max-suppression.png" style="width:500px;height:400;">
<caption><center> <u> **Figure 7** </u>: In this example, the model has predicted 3 cars, but it's actually 3 predictions of the same car. Running non-max suppression (NMS) will select only the most accurate (highest probability) of the 3 boxes. <br> </center></caption>
Non-max suppression uses the very important function called **"Intersection over Union"**, or IoU.
<img src="nb_images/iou.png" style="width:500px;height:400;">
<caption><center> <u> **Figure 8** </u>: Definition of "Intersection over Union". <br> </center></caption>
#### **Exercise**: Implement iou(). Some hints:
- In this code, we use the convention that (0,0) is the top-left corner of an image, (1,0) is the upper-right corner, and (1,1) is the lower-right corner. In other words, the (0,0) origin starts at the top left corner of the image. As x increases, we move to the right. As y increases, we move down.
- For this exercise, we define a box using its two corners: upper left $(x_1, y_1)$ and lower right $(x_2,y_2)$, instead of using the midpoint, height and width. (This makes it a bit easier to calculate the intersection).
- To calculate the area of a rectangle, multiply its height $(y_2 - y_1)$ by its width $(x_2 - x_1)$. (Since $(x_1,y_1)$ is the top left and $x_2,y_2$ are the bottom right, these differences should be non-negative.
- To find the **intersection** of the two boxes $(xi_{1}, yi_{1}, xi_{2}, yi_{2})$:
- Feel free to draw some examples on paper to clarify this conceptually.
- The top left corner of the intersection $(xi_{1}, yi_{1})$ is found by comparing the top left corners $(x_1, y_1)$ of the two boxes and finding a vertex that has an x-coordinate that is closer to the right, and y-coordinate that is closer to the bottom.
- The bottom right corner of the intersection $(xi_{2}, yi_{2})$ is found by comparing the bottom right corners $(x_2,y_2)$ of the two boxes and finding a vertex whose x-coordinate is closer to the left, and the y-coordinate that is closer to the top.
- The two boxes **may have no intersection**. You can detect this if the intersection coordinates you calculate end up being the top right and/or bottom left corners of an intersection box. Another way to think of this is if you calculate the height $(y_2 - y_1)$ or width $(x_2 - x_1)$ and find that at least one of these lengths is negative, then there is no intersection (intersection area is zero).
- The two boxes may intersect at the **edges or vertices**, in which case the intersection area is still zero. This happens when either the height or width (or both) of the calculated intersection is zero.
**Additional Hints**
- `xi1` = **max**imum of the x1 coordinates of the two boxes
- `yi1` = **max**imum of the y1 coordinates of the two boxes
- `xi2` = **min**imum of the x2 coordinates of the two boxes
- `yi2` = **min**imum of the y2 coordinates of the two boxes
- `inter_area` = You can use `max(height, 0)` and `max(width, 0)`
```
# GRADED FUNCTION: iou
def iou(box1, box2):
"""Implement the intersection over union (IoU) between box1 and box2
Arguments:
box1 -- first box, list object with coordinates (box1_x1, box1_y1, box1_x2, box_1_y2)
box2 -- second box, list object with coordinates (box2_x1, box2_y1, box2_x2, box2_y2)
"""
# Assign variable names to coordinates for clarity
(box1_x1, box1_y1, box1_x2, box1_y2) = box1
(box2_x1, box2_y1, box2_x2, box2_y2) = box2
# Calculate the (yi1, xi1, yi2, xi2) coordinates of the intersection of box1 and box2. Calculate its Area.
### START CODE HERE ### (≈ 7 lines)
xi1 = max(box1_x1,box2_x1)
yi1 = max(box1_y1,box2_y1)
xi2 = min(box1_x2,box2_x2)
yi2 = min(box1_y2,box2_y2)
inter_area = max(yi2 - yi1, 0) * max(xi2 - xi1, 0)
### END CODE HERE ###
# Calculate the Union area by using Formula: Union(A,B) = A + B - Inter(A,B)
### START CODE HERE ### (≈ 3 lines)
box1_area = (box1[3] - box1[1]) * (box1[2] - box1[0])
box2_area = (box2[3] - box2[1]) * (box2[2] - box2[0])
union_area = box1_area + box2_area - inter_area
### END CODE HERE ###
# compute the IoU
### START CODE HERE ### (≈ 1 line)
iou = inter_area / union_area
### END CODE HERE ###
return iou
## Test case 1: boxes intersect
box1 = (2, 1, 4, 3)
box2 = (1, 2, 3, 4)
print("iou for intersecting boxes = " + str(iou(box1, box2)))
## Test case 2: boxes do not intersect
box1 = (1,2,3,4)
box2 = (5,6,7,8)
print("iou for non-intersecting boxes = " + str(iou(box1,box2)))
## Test case 3: boxes intersect at vertices only
box1 = (1,1,2,2)
box2 = (2,2,3,3)
print("iou for boxes that only touch at vertices = " + str(iou(box1,box2)))
## Test case 4: boxes intersect at edge only
box1 = (1,1,3,3)
box2 = (2,3,3,4)
print("iou for boxes that only touch at edges = " + str(iou(box1,box2)))
```
**Expected Output**:
```
iou for intersecting boxes = 0.14285714285714285
iou for non-intersecting boxes = 0.0
iou for boxes that only touch at vertices = 0.0
iou for boxes that only touch at edges = 0.0
```
#### YOLO non-max suppression
You are now ready to implement non-max suppression. The key steps are:
1. Select the box that has the highest score.
2. Compute the overlap of this box with all other boxes, and remove boxes that overlap significantly (iou >= `iou_threshold`).
3. Go back to step 1 and iterate until there are no more boxes with a lower score than the currently selected box.
This will remove all boxes that have a large overlap with the selected boxes. Only the "best" boxes remain.
**Exercise**: Implement yolo_non_max_suppression() using TensorFlow. TensorFlow has two built-in functions that are used to implement non-max suppression (so you don't actually need to use your `iou()` implementation):
** Reference documentation **
- [tf.image.non_max_suppression()](https://www.tensorflow.org/api_docs/python/tf/image/non_max_suppression)
```
tf.image.non_max_suppression(
boxes,
scores,
max_output_size,
iou_threshold=0.5,
name=None
)
```
Note that in the version of tensorflow used here, there is no parameter `score_threshold` (it's shown in the documentation for the latest version) so trying to set this value will result in an error message: *got an unexpected keyword argument 'score_threshold.*
- [K.gather()](https://www.tensorflow.org/api_docs/python/tf/keras/backend/gather)
Even though the documentation shows `tf.keras.backend.gather()`, you can use `keras.gather()`.
```
keras.gather(
reference,
indices
)
```
```
# GRADED FUNCTION: yolo_non_max_suppression
def yolo_non_max_suppression(scores, boxes, classes, max_boxes = 10, iou_threshold = 0.5):
"""
Applies Non-max suppression (NMS) to set of boxes
Arguments:
scores -- tensor of shape (None,), output of yolo_filter_boxes()
boxes -- tensor of shape (None, 4), output of yolo_filter_boxes() that have been scaled to the image size (see later)
classes -- tensor of shape (None,), output of yolo_filter_boxes()
max_boxes -- integer, maximum number of predicted boxes you'd like
iou_threshold -- real value, "intersection over union" threshold used for NMS filtering
Returns:
scores -- tensor of shape (, None), predicted score for each box
boxes -- tensor of shape (4, None), predicted box coordinates
classes -- tensor of shape (, None), predicted class for each box
Note: The "None" dimension of the output tensors has obviously to be less than max_boxes. Note also that this
function will transpose the shapes of scores, boxes, classes. This is made for convenience.
"""
max_boxes_tensor = K.variable(max_boxes, dtype='int32') # tensor to be used in tf.image.non_max_suppression()
K.get_session().run(tf.variables_initializer([max_boxes_tensor])) # initialize variable max_boxes_tensor
nms_indices = tf.image.non_max_suppression(boxes = boxes, scores = scores, max_output_size = max_boxes, iou_threshold = iou_threshold)
### END CODE HERE ###
# Use K.gather() to select only nms_indices from scores, boxes and classes
### START CODE HERE ### (≈ 3 lines)
scores = K.gather(scores, nms_indices)
boxes = K.gather(boxes, nms_indices)
classes = K.gather(classes, nms_indices)
### END CODE HERE ###
return scores, boxes, classes
with tf.Session() as test_b:
scores = tf.random_normal([54,], mean=1, stddev=4, seed = 1)
boxes = tf.random_normal([54, 4], mean=1, stddev=4, seed = 1)
classes = tf.random_normal([54,], mean=1, stddev=4, seed = 1)
scores, boxes, classes = yolo_non_max_suppression(scores, boxes, classes)
print("scores[2] = " + str(scores[2].eval()))
print("boxes[2] = " + str(boxes[2].eval()))
print("classes[2] = " + str(classes[2].eval()))
print("scores.shape = " + str(scores.eval().shape))
print("boxes.shape = " + str(boxes.eval().shape))
print("classes.shape = " + str(classes.eval().shape))
```
**Expected Output**:
<table>
<tr>
<td>
**scores[2]**
</td>
<td>
6.9384
</td>
</tr>
<tr>
<td>
**boxes[2]**
</td>
<td>
[-5.299932 3.13798141 4.45036697 0.95942086]
</td>
</tr>
<tr>
<td>
**classes[2]**
</td>
<td>
-2.24527
</td>
</tr>
<tr>
<td>
**scores.shape**
</td>
<td>
(10,)
</td>
</tr>
<tr>
<td>
**boxes.shape**
</td>
<td>
(10, 4)
</td>
</tr>
<tr>
<td>
**classes.shape**
</td>
<td>
(10,)
</td>
</tr>
</table>
### 2.4 Wrapping up the filtering
It's time to implement a function taking the output of the deep CNN (the 19x19x5x85 dimensional encoding) and filtering through all the boxes using the functions you've just implemented.
**Exercise**: Implement `yolo_eval()` which takes the output of the YOLO encoding and filters the boxes using score threshold and NMS. There's just one last implementational detail you have to know. There're a few ways of representing boxes, such as via their corners or via their midpoint and height/width. YOLO converts between a few such formats at different times, using the following functions (which we have provided):
```python
boxes = yolo_boxes_to_corners(box_xy, box_wh)
```
which converts the yolo box coordinates (x,y,w,h) to box corners' coordinates (x1, y1, x2, y2) to fit the input of `yolo_filter_boxes`
```python
boxes = scale_boxes(boxes, image_shape)
```
YOLO's network was trained to run on 608x608 images. If you are testing this data on a different size image--for example, the car detection dataset had 720x1280 images--this step rescales the boxes so that they can be plotted on top of the original 720x1280 image.
Don't worry about these two functions; we'll show you where they need to be called.
```
# GRADED FUNCTION: yolo_eval
def yolo_eval(yolo_outputs, image_shape = (720., 1280.), max_boxes=10, score_threshold=.6, iou_threshold=.5):
"""
Converts the output of YOLO encoding (a lot of boxes) to your predicted boxes along with their scores, box coordinates and classes.
Arguments:
yolo_outputs -- output of the encoding model (for image_shape of (608, 608, 3)), contains 4 tensors:
box_confidence: tensor of shape (None, 19, 19, 5, 1)
box_xy: tensor of shape (None, 19, 19, 5, 2)
box_wh: tensor of shape (None, 19, 19, 5, 2)
box_class_probs: tensor of shape (None, 19, 19, 5, 80)
image_shape -- tensor of shape (2,) containing the input shape, in this notebook we use (608., 608.) (has to be float32 dtype)
max_boxes -- integer, maximum number of predicted boxes you'd like
score_threshold -- real value, if [ highest class probability score < threshold], then get rid of the corresponding box
iou_threshold -- real value, "intersection over union" threshold used for NMS filtering
Returns:
scores -- tensor of shape (None, ), predicted score for each box
boxes -- tensor of shape (None, 4), predicted box coordinates
classes -- tensor of shape (None,), predicted class for each box
"""
### START CODE HERE ###
# Retrieve outputs of the YOLO model (≈1 line)
box_confidence, box_xy, box_wh, box_class_probs = yolo_outputs
# Convert boxes to be ready for filtering functions (convert boxes box_xy and box_wh to corner coordinates)
boxes = yolo_boxes_to_corners(box_xy, box_wh)
# Use one of the functions you've implemented to perform Score-filtering with a threshold of score_threshold (≈1 line)
scores, boxes, classes = yolo_filter_boxes(box_confidence, boxes, box_class_probs, threshold = 0.5)
# Scale boxes back to original image shape.
boxes = scale_boxes(boxes, image_shape)
# Use one of the functions you've implemented to perform Non-max suppression with
# maximum number of boxes set to max_boxes and a threshold of iou_threshold (≈1 line)
scores, boxes, classes = yolo_non_max_suppression(scores, boxes, classes)
### END CODE HERE ###
return scores, boxes, classes
with tf.Session() as test_b:
yolo_outputs = (tf.random_normal([19, 19, 5, 1], mean=1, stddev=4, seed = 1),
tf.random_normal([19, 19, 5, 2], mean=1, stddev=4, seed = 1),
tf.random_normal([19, 19, 5, 2], mean=1, stddev=4, seed = 1),
tf.random_normal([19, 19, 5, 80], mean=1, stddev=4, seed = 1))
scores, boxes, classes = yolo_eval(yolo_outputs)
print("scores[2] = " + str(scores[2].eval()))
print("boxes[2] = " + str(boxes[2].eval()))
print("classes[2] = " + str(classes[2].eval()))
print("scores.shape = " + str(scores.eval().shape))
print("boxes.shape = " + str(boxes.eval().shape))
print("classes.shape = " + str(classes.eval().shape))
```
**Expected Output**:
<table>
<tr>
<td>
**scores[2]**
</td>
<td>
138.791
</td>
</tr>
<tr>
<td>
**boxes[2]**
</td>
<td>
[ 1292.32971191 -278.52166748 3876.98925781 -835.56494141]
</td>
</tr>
<tr>
<td>
**classes[2]**
</td>
<td>
54
</td>
</tr>
<tr>
<td>
**scores.shape**
</td>
<td>
(10,)
</td>
</tr>
<tr>
<td>
**boxes.shape**
</td>
<td>
(10, 4)
</td>
</tr>
<tr>
<td>
**classes.shape**
</td>
<td>
(10,)
</td>
</tr>
</table>
## Summary for YOLO:
- Input image (608, 608, 3)
- The input image goes through a CNN, resulting in a (19,19,5,85) dimensional output.
- After flattening the last two dimensions, the output is a volume of shape (19, 19, 425):
- Each cell in a 19x19 grid over the input image gives 425 numbers.
- 425 = 5 x 85 because each cell contains predictions for 5 boxes, corresponding to 5 anchor boxes, as seen in lecture.
- 85 = 5 + 80 where 5 is because $(p_c, b_x, b_y, b_h, b_w)$ has 5 numbers, and 80 is the number of classes we'd like to detect
- You then select only few boxes based on:
- Score-thresholding: throw away boxes that have detected a class with a score less than the threshold
- Non-max suppression: Compute the Intersection over Union and avoid selecting overlapping boxes
- This gives you YOLO's final output.
## 3 - Test YOLO pre-trained model on images
In this part, you are going to use a pre-trained model and test it on the car detection dataset. We'll need a session to execute the computation graph and evaluate the tensors.
```
sess = K.get_session()
```
### 3.1 - Defining classes, anchors and image shape.
* Recall that we are trying to detect 80 classes, and are using 5 anchor boxes.
* We have gathered the information on the 80 classes and 5 boxes in two files "coco_classes.txt" and "yolo_anchors.txt".
* We'll read class names and anchors from text files.
* The car detection dataset has 720x1280 images, which we've pre-processed into 608x608 images.
```
class_names = read_classes("model_data/coco_classes.txt")
anchors = read_anchors("model_data/yolo_anchors.txt")
image_shape = (720., 1280.)
```
### 3.2 - Loading a pre-trained model
* Training a YOLO model takes a very long time and requires a fairly large dataset of labelled bounding boxes for a large range of target classes.
* You are going to load an existing pre-trained Keras YOLO model stored in "yolo.h5".
* These weights come from the official YOLO website, and were converted using a function written by Allan Zelener. References are at the end of this notebook. Technically, these are the parameters from the "YOLOv2" model, but we will simply refer to it as "YOLO" in this notebook.
Run the cell below to load the model from this file.
```
yolo_model = load_model("model_data/yolo.h5")
```
This loads the weights of a trained YOLO model. Here's a summary of the layers your model contains.
```
yolo_model.summary()
```
**Note**: On some computers, you may see a warning message from Keras. Don't worry about it if you do--it is fine.
**Reminder**: this model converts a preprocessed batch of input images (shape: (m, 608, 608, 3)) into a tensor of shape (m, 19, 19, 5, 85) as explained in Figure (2).
### 3.3 - Convert output of the model to usable bounding box tensors
The output of `yolo_model` is a (m, 19, 19, 5, 85) tensor that needs to pass through non-trivial processing and conversion. The following cell does that for you.
If you are curious about how `yolo_head` is implemented, you can find the function definition in the file ['keras_yolo.py'](https://github.com/allanzelener/YAD2K/blob/master/yad2k/models/keras_yolo.py). The file is located in your workspace in this path 'yad2k/models/keras_yolo.py'.
```
yolo_outputs = yolo_head(yolo_model.output, anchors, len(class_names))
```
You added `yolo_outputs` to your graph. This set of 4 tensors is ready to be used as input by your `yolo_eval` function.
### 3.4 - Filtering boxes
`yolo_outputs` gave you all the predicted boxes of `yolo_model` in the correct format. You're now ready to perform filtering and select only the best boxes. Let's now call `yolo_eval`, which you had previously implemented, to do this.
```
scores, boxes, classes = yolo_eval(yolo_outputs, image_shape)
```
### 3.5 - Run the graph on an image
Let the fun begin. You have created a graph that can be summarized as follows:
1. <font color='purple'> yolo_model.input </font> is given to `yolo_model`. The model is used to compute the output <font color='purple'> yolo_model.output </font>
2. <font color='purple'> yolo_model.output </font> is processed by `yolo_head`. It gives you <font color='purple'> yolo_outputs </font>
3. <font color='purple'> yolo_outputs </font> goes through a filtering function, `yolo_eval`. It outputs your predictions: <font color='purple'> scores, boxes, classes </font>
**Exercise**: Implement predict() which runs the graph to test YOLO on an image.
You will need to run a TensorFlow session, to have it compute `scores, boxes, classes`.
The code below also uses the following function:
```python
image, image_data = preprocess_image("images/" + image_file, model_image_size = (608, 608))
```
which outputs:
- image: a python (PIL) representation of your image used for drawing boxes. You won't need to use it.
- image_data: a numpy-array representing the image. This will be the input to the CNN.
**Important note**: when a model uses BatchNorm (as is the case in YOLO), you will need to pass an additional placeholder in the feed_dict {K.learning_phase(): 0}.
#### Hint: Using the TensorFlow Session object
* Recall that above, we called `K.get_Session()` and saved the Session object in `sess`.
* To evaluate a list of tensors, we call `sess.run()` like this:
```
sess.run(fetches=[tensor1,tensor2,tensor3],
feed_dict={yolo_model.input: the_input_variable,
K.learning_phase():0
}
```
* Notice that the variables `scores, boxes, classes` are not passed into the `predict` function, but these are global variables that you will use within the `predict` function.
```
def predict(sess, image_file):
"""
Runs the graph stored in "sess" to predict boxes for "image_file". Prints and plots the predictions.
Arguments:
sess -- your tensorflow/Keras session containing the YOLO graph
image_file -- name of an image stored in the "images" folder.
Returns:
out_scores -- tensor of shape (None, ), scores of the predicted boxes
out_boxes -- tensor of shape (None, 4), coordinates of the predicted boxes
out_classes -- tensor of shape (None, ), class index of the predicted boxes
Note: "None" actually represents the number of predicted boxes, it varies between 0 and max_boxes.
"""
# Preprocess your image
image, image_data = preprocess_image("images/" + image_file, model_image_size = (608, 608))
# Run the session with the correct tensors and choose the correct placeholders in the feed_dict.
# You'll need to use feed_dict={yolo_model.input: ... , K.learning_phase(): 0})
### START CODE HERE ### (≈ 1 line)
out_scores, out_boxes, out_classes = sess.run([scores, boxes, classes], feed_dict={yolo_model.input: image_data, K.learning_phase(): 0})
### END CODE HERE ###
# Print predictions info
print('Found {} boxes for {}'.format(len(out_boxes), image_file))
# Generate colors for drawing bounding boxes.
colors = generate_colors(class_names)
# Draw bounding boxes on the image file
draw_boxes(image, out_scores, out_boxes, out_classes, class_names, colors)
# Save the predicted bounding box on the image
image.save(os.path.join("out", image_file), quality=90)
# Display the results in the notebook
output_image = scipy.misc.imread(os.path.join("out", image_file))
imshow(output_image)
return out_scores, out_boxes, out_classes
```
Run the following cell on the "test.jpg" image to verify that your function is correct.
```
out_scores, out_boxes, out_classes = predict(sess, "test.jpg")
```
**Expected Output**:
<table>
<tr>
<td>
**Found 7 boxes for test.jpg**
</td>
</tr>
<tr>
<td>
**car**
</td>
<td>
0.60 (925, 285) (1045, 374)
</td>
</tr>
<tr>
<td>
**car**
</td>
<td>
0.66 (706, 279) (786, 350)
</td>
</tr>
<tr>
<td>
**bus**
</td>
<td>
0.67 (5, 266) (220, 407)
</td>
</tr>
<tr>
<td>
**car**
</td>
<td>
0.70 (947, 324) (1280, 705)
</td>
</tr>
<tr>
<td>
**car**
</td>
<td>
0.74 (159, 303) (346, 440)
</td>
</tr>
<tr>
<td>
**car**
</td>
<td>
0.80 (761, 282) (942, 412)
</td>
</tr>
<tr>
<td>
**car**
</td>
<td>
0.89 (367, 300) (745, 648)
</td>
</tr>
</table>
The model you've just run is actually able to detect 80 different classes listed in "coco_classes.txt". To test the model on your own images:
1. Click on "File" in the upper bar of this notebook, then click "Open" to go on your Coursera Hub.
2. Add your image to this Jupyter Notebook's directory, in the "images" folder
3. Write your image's name in the cell above code
4. Run the code and see the output of the algorithm!
If you were to run your session in a for loop over all your images. Here's what you would get:
<center>
<video width="400" height="200" src="nb_images/pred_video_compressed2.mp4" type="video/mp4" controls>
</video>
</center>
<caption><center> Predictions of the YOLO model on pictures taken from a camera while driving around the Silicon Valley <br> Thanks [drive.ai](https://www.drive.ai/) for providing this dataset! </center></caption>
## <font color='darkblue'>What you should remember:
- YOLO is a state-of-the-art object detection model that is fast and accurate
- It runs an input image through a CNN which outputs a 19x19x5x85 dimensional volume.
- The encoding can be seen as a grid where each of the 19x19 cells contains information about 5 boxes.
- You filter through all the boxes using non-max suppression. Specifically:
- Score thresholding on the probability of detecting a class to keep only accurate (high probability) boxes
- Intersection over Union (IoU) thresholding to eliminate overlapping boxes
- Because training a YOLO model from randomly initialized weights is non-trivial and requires a large dataset as well as lot of computation, we used previously trained model parameters in this exercise. If you wish, you can also try fine-tuning the YOLO model with your own dataset, though this would be a fairly non-trivial exercise.
**References**: The ideas presented in this notebook came primarily from the two YOLO papers. The implementation here also took significant inspiration and used many components from Allan Zelener's GitHub repository. The pre-trained weights used in this exercise came from the official YOLO website.
- Joseph Redmon, Santosh Divvala, Ross Girshick, Ali Farhadi - [You Only Look Once: Unified, Real-Time Object Detection](https://arxiv.org/abs/1506.02640) (2015)
- Joseph Redmon, Ali Farhadi - [YOLO9000: Better, Faster, Stronger](https://arxiv.org/abs/1612.08242) (2016)
- Allan Zelener - [YAD2K: Yet Another Darknet 2 Keras](https://github.com/allanzelener/YAD2K)
- The official YOLO website (https://pjreddie.com/darknet/yolo/)
**Car detection dataset**:
<a rel="license" href="http://creativecommons.org/licenses/by/4.0/"><img alt="Creative Commons License" style="border-width:0" src="https://i.creativecommons.org/l/by/4.0/88x31.png" /></a><br /><span xmlns:dct="http://purl.org/dc/terms/" property="dct:title">The Drive.ai Sample Dataset</span> (provided by drive.ai) is licensed under a <a rel="license" href="http://creativecommons.org/licenses/by/4.0/">Creative Commons Attribution 4.0 International License</a>. We are grateful to Brody Huval, Chih Hu and Rahul Patel for providing this data.
| github_jupyter |
```
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import os
print(os.listdir("../input"))
import time
# import pytorch
import torch
import torch.nn as nn
import torch.nn.functional as F
from torch.optim import SGD,Adam,lr_scheduler
from torch.utils.data import random_split
import torchvision
from torchvision import transforms, datasets
from torch.utils.data import DataLoader
# define transformations for train
train_transform = transforms.Compose([
transforms.RandomHorizontalFlip(p=.40),
transforms.RandomRotation(30),
transforms.ToTensor(),
transforms.Normalize([0.5, 0.5, 0.5], [0.5, 0.5, 0.5])])
# define transformations for test
test_transform = transforms.Compose([
transforms.ToTensor(),
transforms.Normalize([0.5, 0.5, 0.5], [0.5, 0.5, 0.5])])
# define training dataloader
def get_training_dataloader(train_transform, batch_size=128, num_workers=0, shuffle=True):
""" return training dataloader
Args:
train_transform: transfroms for train dataset
path: path to cifar100 training python dataset
batch_size: dataloader batchsize
num_workers: dataloader num_works
shuffle: whether to shuffle
Returns: train_data_loader:torch dataloader object
"""
transform_train = train_transform
cifar10_training = torchvision.datasets.CIFAR10(root='.', train=True, download=True, transform=transform_train)
cifar10_training_loader = DataLoader(
cifar10_training, shuffle=shuffle, num_workers=num_workers, batch_size=batch_size)
return cifar10_training_loader
# define test dataloader
def get_testing_dataloader(test_transform, batch_size=128, num_workers=0, shuffle=True):
""" return training dataloader
Args:
test_transform: transforms for test dataset
path: path to cifar100 test python dataset
batch_size: dataloader batchsize
num_workers: dataloader num_works
shuffle: whether to shuffle
Returns: cifar100_test_loader:torch dataloader object
"""
transform_test = test_transform
cifar10_test = torchvision.datasets.CIFAR10(root='.', train=False, download=True, transform=transform_test)
cifar10_test_loader = DataLoader(
cifar10_test, shuffle=shuffle, num_workers=num_workers, batch_size=batch_size)
return cifar10_test_loader
# implement mish activation function
def f_mish(input, inplace = False):
'''
Applies the mish function element-wise:
mish(x) = x * tanh(softplus(x)) = x * tanh(ln(1 + exp(x)))
'''
return input * torch.tanh(F.softplus(input))
# implement class wrapper for mish activation function
class mish(nn.Module):
'''
Applies the mish function element-wise:
mish(x) = x * tanh(softplus(x)) = x * tanh(ln(1 + exp(x)))
Shape:
- Input: (N, *) where * means, any number of additional
dimensions
- Output: (N, *), same shape as the input
Examples:
>>> m = mish()
>>> input = torch.randn(2)
>>> output = m(input)
'''
def __init__(self, inplace = False):
'''
Init method.
'''
super().__init__()
self.inplace = inplace
def forward(self, input):
'''
Forward pass of the function.
'''
return f_mish(input, inplace = self.inplace)
# implement swish activation function
def f_swish(input, inplace = False):
'''
Applies the swish function element-wise:
swish(x) = x * sigmoid(x)
'''
return input * torch.sigmoid(input)
# implement class wrapper for swish activation function
class swish(nn.Module):
'''
Applies the swish function element-wise:
swish(x) = x * sigmoid(x)
Shape:
- Input: (N, *) where * means, any number of additional
dimensions
- Output: (N, *), same shape as the input
Examples:
>>> m = swish()
>>> input = torch.randn(2)
>>> output = m(input)
'''
def __init__(self, inplace = False):
'''
Init method.
'''
super().__init__()
self.inplace = inplace
def forward(self, input):
'''
Forward pass of the function.
'''
return f_swish(input, inplace = self.inplace)
class BasicConv2d(nn.Module):
def __init__(self, input_channels, output_channels, activation = 'relu', **kwargs):
super().__init__()
self.conv = nn.Conv2d(input_channels, output_channels, bias=False, **kwargs)
self.bn = nn.BatchNorm2d(output_channels)
if activation == 'relu':
self.relu = nn.ReLU(inplace=True)
if activation == 'swish':
self.relu = swish(inplace = True)
if activation == 'mish':
self.relu = mish(inplace = True)
def forward(self, x):
x = self.conv(x)
x = self.bn(x)
x = self.relu(x)
return x
#same naive inception module
class InceptionA(nn.Module):
def __init__(self, input_channels, pool_features, activation = 'relu'):
super().__init__()
self.branch1x1 = BasicConv2d(input_channels, 64, kernel_size=1, activation = activation)
self.branch5x5 = nn.Sequential(
BasicConv2d(input_channels, 48, kernel_size=1, activation = activation),
BasicConv2d(48, 64, kernel_size=5, padding=2, activation = activation)
)
self.branch3x3 = nn.Sequential(
BasicConv2d(input_channels, 64, kernel_size=1, activation = activation),
BasicConv2d(64, 96, kernel_size=3, padding=1, activation = activation),
BasicConv2d(96, 96, kernel_size=3, padding=1, activation = activation)
)
self.branchpool = nn.Sequential(
nn.AvgPool2d(kernel_size=3, stride=1, padding=1),
BasicConv2d(input_channels, pool_features, kernel_size=3, padding=1, activation = activation)
)
def forward(self, x):
#x -> 1x1(same)
branch1x1 = self.branch1x1(x)
#x -> 1x1 -> 5x5(same)
branch5x5 = self.branch5x5(x)
#branch5x5 = self.branch5x5_2(branch5x5)
#x -> 1x1 -> 3x3 -> 3x3(same)
branch3x3 = self.branch3x3(x)
#x -> pool -> 1x1(same)
branchpool = self.branchpool(x)
outputs = [branch1x1, branch5x5, branch3x3, branchpool]
return torch.cat(outputs, 1)
#downsample
#Factorization into smaller convolutions
class InceptionB(nn.Module):
def __init__(self, input_channels, activation = 'relu'):
super().__init__()
self.branch3x3 = BasicConv2d(input_channels, 384, kernel_size=3, stride=2, activation = activation)
self.branch3x3stack = nn.Sequential(
BasicConv2d(input_channels, 64, kernel_size=1, activation = activation),
BasicConv2d(64, 96, kernel_size=3, padding=1, activation = activation),
BasicConv2d(96, 96, kernel_size=3, stride=2, activation = activation)
)
self.branchpool = nn.MaxPool2d(kernel_size=3, stride=2)
def forward(self, x):
#x - > 3x3(downsample)
branch3x3 = self.branch3x3(x)
#x -> 3x3 -> 3x3(downsample)
branch3x3stack = self.branch3x3stack(x)
#x -> avgpool(downsample)
branchpool = self.branchpool(x)
#"""We can use two parallel stride 2 blocks: P and C. P is a pooling
#layer (either average or maximum pooling) the activation, both of
#them are stride 2 the filter banks of which are concatenated as in
#figure 10."""
outputs = [branch3x3, branch3x3stack, branchpool]
return torch.cat(outputs, 1)
#Factorizing Convolutions with Large Filter Size
class InceptionC(nn.Module):
def __init__(self, input_channels, channels_7x7, activation = 'relu'):
super().__init__()
self.branch1x1 = BasicConv2d(input_channels, 192, kernel_size=1, activation = activation)
c7 = channels_7x7
#In theory, we could go even further and argue that one can replace any n × n
#convolution by a 1 × n convolution followed by a n × 1 convolution and the
#computational cost saving increases dramatically as n grows (see figure 6).
self.branch7x7 = nn.Sequential(
BasicConv2d(input_channels, c7, kernel_size=1, activation = activation),
BasicConv2d(c7, c7, kernel_size=(7, 1), padding=(3, 0), activation = activation),
BasicConv2d(c7, 192, kernel_size=(1, 7), padding=(0, 3), activation = activation)
)
self.branch7x7stack = nn.Sequential(
BasicConv2d(input_channels, c7, kernel_size=1, activation = activation),
BasicConv2d(c7, c7, kernel_size=(7, 1), padding=(3, 0), activation = activation),
BasicConv2d(c7, c7, kernel_size=(1, 7), padding=(0, 3), activation = activation),
BasicConv2d(c7, c7, kernel_size=(7, 1), padding=(3, 0), activation = activation),
BasicConv2d(c7, 192, kernel_size=(1, 7), padding=(0, 3), activation = activation)
)
self.branch_pool = nn.Sequential(
nn.AvgPool2d(kernel_size=3, stride=1, padding=1),
BasicConv2d(input_channels, 192, kernel_size=1, activation = activation),
)
def forward(self, x):
#x -> 1x1(same)
branch1x1 = self.branch1x1(x)
#x -> 1layer 1*7 and 7*1 (same)
branch7x7 = self.branch7x7(x)
#x-> 2layer 1*7 and 7*1(same)
branch7x7stack = self.branch7x7stack(x)
#x-> avgpool (same)
branchpool = self.branch_pool(x)
outputs = [branch1x1, branch7x7, branch7x7stack, branchpool]
return torch.cat(outputs, 1)
class InceptionD(nn.Module):
def __init__(self, input_channels, activation = 'relu'):
super().__init__()
self.branch3x3 = nn.Sequential(
BasicConv2d(input_channels, 192, kernel_size=1, activation = activation),
BasicConv2d(192, 320, kernel_size=3, stride=2, activation = activation)
)
self.branch7x7 = nn.Sequential(
BasicConv2d(input_channels, 192, kernel_size=1, activation = activation),
BasicConv2d(192, 192, kernel_size=(1, 7), padding=(0, 3), activation = activation),
BasicConv2d(192, 192, kernel_size=(7, 1), padding=(3, 0), activation = activation),
BasicConv2d(192, 192, kernel_size=3, stride=2, activation = activation)
)
self.branchpool = nn.AvgPool2d(kernel_size=3, stride=2)
def forward(self, x):
#x -> 1x1 -> 3x3(downsample)
branch3x3 = self.branch3x3(x)
#x -> 1x1 -> 1x7 -> 7x1 -> 3x3 (downsample)
branch7x7 = self.branch7x7(x)
#x -> avgpool (downsample)
branchpool = self.branchpool(x)
outputs = [branch3x3, branch7x7, branchpool]
return torch.cat(outputs, 1)
#same
class InceptionE(nn.Module):
def __init__(self, input_channels, activation = 'relu'):
super().__init__()
self.branch1x1 = BasicConv2d(input_channels, 320, kernel_size=1, activation = activation)
self.branch3x3_1 = BasicConv2d(input_channels, 384, kernel_size=1, activation = activation)
self.branch3x3_2a = BasicConv2d(384, 384, kernel_size=(1, 3), padding=(0, 1), activation = activation)
self.branch3x3_2b = BasicConv2d(384, 384, kernel_size=(3, 1), padding=(1, 0), activation = activation)
self.branch3x3stack_1 = BasicConv2d(input_channels, 448, kernel_size=1, activation = activation)
self.branch3x3stack_2 = BasicConv2d(448, 384, kernel_size=3, padding=1, activation = activation)
self.branch3x3stack_3a = BasicConv2d(384, 384, kernel_size=(1, 3), padding=(0, 1), activation = activation)
self.branch3x3stack_3b = BasicConv2d(384, 384, kernel_size=(3, 1), padding=(1, 0), activation = activation)
self.branch_pool = nn.Sequential(
nn.AvgPool2d(kernel_size=3, stride=1, padding=1),
BasicConv2d(input_channels, 192, kernel_size=1, activation = activation)
)
def forward(self, x):
#x -> 1x1 (same)
branch1x1 = self.branch1x1(x)
# x -> 1x1 -> 3x1
# x -> 1x1 -> 1x3
# concatenate(3x1, 1x3)
#"""7. Inception modules with expanded the filter bank outputs.
#This architecture is used on the coarsest (8 × 8) grids to promote
#high dimensional representations, as suggested by principle
#2 of Section 2."""
branch3x3 = self.branch3x3_1(x)
branch3x3 = [
self.branch3x3_2a(branch3x3),
self.branch3x3_2b(branch3x3)
]
branch3x3 = torch.cat(branch3x3, 1)
# x -> 1x1 -> 3x3 -> 1x3
# x -> 1x1 -> 3x3 -> 3x1
#concatenate(1x3, 3x1)
branch3x3stack = self.branch3x3stack_1(x)
branch3x3stack = self.branch3x3stack_2(branch3x3stack)
branch3x3stack = [
self.branch3x3stack_3a(branch3x3stack),
self.branch3x3stack_3b(branch3x3stack)
]
branch3x3stack = torch.cat(branch3x3stack, 1)
branchpool = self.branch_pool(x)
outputs = [branch1x1, branch3x3, branch3x3stack, branchpool]
return torch.cat(outputs, 1)
class InceptionV3(nn.Module):
def __init__(self, num_classes=10, activation = 'relu'):
super().__init__()
self.Conv2d_1a_3x3 = BasicConv2d(3, 32, kernel_size=3, padding=1, activation = activation)
self.Conv2d_2a_3x3 = BasicConv2d(32, 32, kernel_size=3, padding=1, activation = activation)
self.Conv2d_2b_3x3 = BasicConv2d(32, 64, kernel_size=3, padding=1, activation = activation)
self.Conv2d_3b_1x1 = BasicConv2d(64, 80, kernel_size=1, activation = activation)
self.Conv2d_4a_3x3 = BasicConv2d(80, 192, kernel_size=3, activation = activation)
#naive inception module
self.Mixed_5b = InceptionA(192, pool_features=32, activation = activation)
self.Mixed_5c = InceptionA(256, pool_features=64, activation = activation)
self.Mixed_5d = InceptionA(288, pool_features=64, activation = activation)
#downsample
self.Mixed_6a = InceptionB(288, activation = activation)
self.Mixed_6b = InceptionC(768, channels_7x7=128, activation = activation)
self.Mixed_6c = InceptionC(768, channels_7x7=160, activation = activation)
self.Mixed_6d = InceptionC(768, channels_7x7=160, activation = activation)
self.Mixed_6e = InceptionC(768, channels_7x7=192, activation = activation)
#downsample
self.Mixed_7a = InceptionD(768, activation = activation)
self.Mixed_7b = InceptionE(1280, activation = activation)
self.Mixed_7c = InceptionE(2048, activation = activation)
#6*6 feature size
self.avgpool = nn.AdaptiveAvgPool2d((1, 1))
self.dropout = nn.Dropout2d()
self.linear = nn.Linear(2048, num_classes)
def forward(self, x):
#32 -> 30
x = self.Conv2d_1a_3x3(x)
x = self.Conv2d_2a_3x3(x)
x = self.Conv2d_2b_3x3(x)
x = self.Conv2d_3b_1x1(x)
x = self.Conv2d_4a_3x3(x)
#30 -> 30
x = self.Mixed_5b(x)
x = self.Mixed_5c(x)
x = self.Mixed_5d(x)
#30 -> 14
#Efficient Grid Size Reduction to avoid representation
#bottleneck
x = self.Mixed_6a(x)
#14 -> 14
#"""In practice, we have found that employing this factorization does not
#work well on early layers, but it gives very good results on medium
#grid-sizes (On m × m feature maps, where m ranges between 12 and 20).
#On that level, very good results can be achieved by using 1 × 7 convolutions
#followed by 7 × 1 convolutions."""
x = self.Mixed_6b(x)
x = self.Mixed_6c(x)
x = self.Mixed_6d(x)
x = self.Mixed_6e(x)
#14 -> 6
#Efficient Grid Size Reduction
x = self.Mixed_7a(x)
#6 -> 6
#We are using this solution only on the coarsest grid,
#since that is the place where producing high dimensional
#sparse representation is the most critical as the ratio of
#local processing (by 1 × 1 convolutions) is increased compared
#to the spatial aggregation."""
x = self.Mixed_7b(x)
x = self.Mixed_7c(x)
#6 -> 1
x = self.avgpool(x)
x = self.dropout(x)
x = x.view(x.size(0), -1)
x = self.linear(x)
return x
def inceptionv3(activation = 'relu'):
return InceptionV3(activation = activation)
trainloader = get_training_dataloader(train_transform)
testloader = get_testing_dataloader(test_transform)
epochs = 100
batch_size = 128
learning_rate = 0.001
device = torch.device('cuda:0' if torch.cuda.is_available() else "cpu")
device
model = inceptionv3(activation = 'mish')
# set loss function
criterion = nn.CrossEntropyLoss()
# set optimizer, only train the classifier parameters, feature parameters are frozen
optimizer = Adam(model.parameters(), lr=learning_rate)
train_stats = pd.DataFrame(columns = ['Epoch', 'Time per epoch', 'Avg time per step', 'Train loss', 'Train accuracy', 'Train top-3 accuracy','Test loss', 'Test accuracy', 'Test top-3 accuracy'])
#train the model
model.to(device)
steps = 0
running_loss = 0
for epoch in range(epochs):
since = time.time()
train_accuracy = 0
top3_train_accuracy = 0
for inputs, labels in trainloader:
steps += 1
# Move input and label tensors to the default device
inputs, labels = inputs.to(device), labels.to(device)
optimizer.zero_grad()
logps = model.forward(inputs)
loss = criterion(logps, labels)
loss.backward()
optimizer.step()
running_loss += loss.item()
# calculate train top-1 accuracy
ps = torch.exp(logps)
top_p, top_class = ps.topk(1, dim=1)
equals = top_class == labels.view(*top_class.shape)
train_accuracy += torch.mean(equals.type(torch.FloatTensor)).item()
# Calculate train top-3 accuracy
np_top3_class = ps.topk(3, dim=1)[1].cpu().numpy()
target_numpy = labels.cpu().numpy()
top3_train_accuracy += np.mean([1 if target_numpy[i] in np_top3_class[i] else 0 for i in range(0, len(target_numpy))])
time_elapsed = time.time() - since
test_loss = 0
test_accuracy = 0
top3_test_accuracy = 0
model.eval()
with torch.no_grad():
for inputs, labels in testloader:
inputs, labels = inputs.to(device), labels.to(device)
logps = model.forward(inputs)
batch_loss = criterion(logps, labels)
test_loss += batch_loss.item()
# Calculate test top-1 accuracy
ps = torch.exp(logps)
top_p, top_class = ps.topk(1, dim=1)
equals = top_class == labels.view(*top_class.shape)
test_accuracy += torch.mean(equals.type(torch.FloatTensor)).item()
# Calculate test top-3 accuracy
np_top3_class = ps.topk(3, dim=1)[1].cpu().numpy()
target_numpy = labels.cpu().numpy()
top3_test_accuracy += np.mean([1 if target_numpy[i] in np_top3_class[i] else 0 for i in range(0, len(target_numpy))])
print(f"Epoch {epoch+1}/{epochs}.. "
f"Time per epoch: {time_elapsed:.4f}.. "
f"Average time per step: {time_elapsed/len(trainloader):.4f}.. "
f"Train loss: {running_loss/len(trainloader):.4f}.. "
f"Train accuracy: {train_accuracy/len(trainloader):.4f}.. "
f"Top-3 train accuracy: {top3_train_accuracy/len(trainloader):.4f}.. "
f"Test loss: {test_loss/len(testloader):.4f}.. "
f"Test accuracy: {test_accuracy/len(testloader):.4f}.. "
f"Top-3 test accuracy: {top3_test_accuracy/len(testloader):.4f}")
train_stats = train_stats.append({'Epoch': epoch, 'Time per epoch':time_elapsed, 'Avg time per step': time_elapsed/len(trainloader), 'Train loss' : running_loss/len(trainloader), 'Train accuracy': train_accuracy/len(trainloader), 'Train top-3 accuracy':top3_train_accuracy/len(trainloader),'Test loss' : test_loss/len(testloader), 'Test accuracy': test_accuracy/len(testloader), 'Test top-3 accuracy':top3_test_accuracy/len(testloader)}, ignore_index=True)
running_loss = 0
model.train()
train_stats.to_csv('train_log_InceptionV3_Mish.csv')
```
| github_jupyter |
<a href="https://colab.research.google.com/github/eric-yyjau/cse252c_hw3-release/blob/master/hw3_questions.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# CSE252C: Homework 3
## Computing Resources
Please read the README file of this repository for the instructions
## Instructions
1. Attempt all questions.
2. Please comment all your code adequately.
3. Include all relevant information such as text answers, output images in notebook.
4. **Academic integrity:** The homework must be completed individually.
5. **Submission instructions:**
(a) Submit the notebook and its PDF version on Gradescope.
(b) Rename your submission files as Lastname_Firstname.ipynb and Lastname_Firstname.pdf.
(c) Correctly select pages for each answer on Gradescope to allow proper grading.
6. **Due date:** To be decided
## Q1: UNet for Image Segmentation
1. **Check the codes in `Segmentation`.** In this homework, we will provide the dataset loader, the evaluation code, the basic UNet structure and some useful functions. You will be asked to try different variations of network structure and decide the best training strategies to obtain good results. Like in previous homeworks, you are welcome to cite any open source codes that help you improve performance. The provided codes include:
1. `test.py`: The file for evaluation.
2. `dataLoader.py`: The file to load the data for training and testing.
3. `model.py`: The file for models. The residual block (`ResBlock`) and the code to load pretrained weights of `resnet18 loadPretrainedWeight`) are given. The basic encoder and decoder are also given as a reference.
4. `colormap.mat`: The color map used to visualize segmentation results.
5. `utils.py`: The file for two useful functions. The `computeAccuracy` function computes the unnormalized confusion matrix of each batch of labels. The `save\_label` function turns the label into an image using the given color map and saves the image at the assigned location. Also see `test.py` for how these two functions are being used.
6. `train.py`: An empty file where you will implement your training script.
2. **Implement the network structures.** You are required to implement 2 versions of UNet structure since the basic structure has already been given. In all three versions, the `resnet18` structure before average pooling and fully connected layer will be used as the building block for encoder. You are strongly recommended to use weights pretrained on ImageNet, which may have a major impact on the performance.
1. `Basic UNet`: The code is given as a reference. Please see `encoder` and `decoder` class in `model.py`. The `encoder` comes from `resnet18` and the decoder consists of transpose convolutional layers and bilinear interpolation layers so that the final output will be of the same size as the image. Skip links are added to help the network recover more details. Please do not change the encoder. However, you are free to change the decoder, while ensuring that the structure of your decoder across three versions of the networks are similar so that you can make a fair comparison of their performances.
2. `UNet with dilation`: We modify the encoder to a dilated `resnet18` as described in Section 2 of [1] (You are not required to consider degridding in Section 4 of [1] ). We set the stride of the last 4 residual blocks to be 1 so that the highest level feature maps will be $4\times 4$ times larger. To increase the receptive field, we set the dilation of residual blocks that are fourth and third from the end to be 2, while the dilation of the residual blocks that are first and second from the end are set to 4. The decoder should be modified accordingly. Implement your new encoder and decoder under class `encoderDilation` and `decoderDilation`. Ensure that for images of arbitrary shapes, the decoder will give segmentation outputs of the same shape. **[15 points]**
3. `UNet with dilation and pyramid pooling`: Based on the encoder-decoder structure with dilation, add pyramid pooling layer after the last residual block of encoder. Implement the pyramid pooling layer following [2]. Notice that after adding the pyramid layer, the number of channels of the output feature to the first transpose convolutional layer will change from 512 to 1024. Please implement your new encoder and decoder under classes `encoderSPP` and `decoderSPP`, respectively. **[15 points]**
3. **Implement training script and train the network.** Train your network using 1464 images from the training set of PASCAL VOC 2012. The dataset is on the server `/datasets/cse152-252-sp20-public/hw3_data/VOCdevkit`. If you are not familiar with training scripts, you can refer to `test.py` in this homework and `casia_train.py` in the previous homework. The structures of the training script are very similar. Please remember to output the training loss and training accuracy which may help you find the best hyper parameters. **[40 points]**
1. To accelerate the training speed, you can use the Pytorch multi-threaded data loader. **Important:** if you use multi-threaded data loader, remember to either randomly shuffle the data or change the random seeds after every epoch. Otherwise you will have severe overfitting issues because the data loader will always crop the same region of the image.
2. It is recommended to compute the prediction mIoU every epoch, since the curve of mIoU can be very different from the inverse of loss function. It may help you find the best training strategy.
3. To overcome over-fitting issues, you are encouraged to adopt more aggressive data augmentation methods, such as flipping the images or changing the intensity.
4. There are many things that may influence performance, such as learning rate, batch size and network structure of encoder and decoder. It might be hard to achieve state-of-the-art results. **The grading of the homework will not focus on the final mean IoU but more on analysis.** So don't be too worried if you cannot get very good performance. Just make sure that you describe what you observe and answer the questions succinctly.
4. **Answer the following questions:**
1. Describe the loss function you use to train the semantic segmentation network. If you change the structure of the decoder, describe your new network architecture. **[10 points]**
2. Describe your training details, such as: what kind of optimizer is used to train the network, what's the learning rate and the batch size, whether you decrease the learning rate as the training progresses, number of epochs required to train the network, or any other details that you find important for performance. Note that in order to compare the three network structures, learning details for them should be the same. **[10 points]**
3. Draw the loss curves of the three network structures in the same graph. **[10 points]**
4. Evaluate the trained models using the following commands. Draw a table to summarize quantitative performances of the 3 variations of the UNet structure. The table should include the mean IoU of 21 categories of objects and their average mean IoU. **[10 points]** :
1. `Basic UNet`: `python test.py`. The testing mean IoU of 21 categories of object are stored in `test/accuracy.npy`. You can add flags if necessary.
2. `UNet with dilation`: `python test.py --isDilation`. The testing mean IoU of 21 categories of objects are stored in `test_dilation/accuracy.npy`. You can add flags if necessary.
3. `UNet with dilation and pyramid pooling`: `python test.py --isSpp`. The testing mean IoU of 21 categories of object are stored in `test_spp/accuracy.npy`. You can add flags if necessary.
5. Make a figure for qualitative comparisons of the 3 methods, shown on 4 different input images. Please show the segmentation results for the same image but different networks so the differences can be compared. Briefly describe the results you obtain and any observations. **[10 points]**
6. Explain your observations in terms of: (i) what choices helped improve the accuracy and (ii) what other steps could have been tried to further improve accuracy? **[10 points]**
``Answer Q1.4.A here``
``Answer Q1.4.B here``
``Answer Q1.4.C here``
``Answer Q1.4.D here``
``Answer Q1.4.E here``
``Answer Q1.4.F here``
## Q2: SSD [3] Object Detection
1. **Check the codes in `Detection`.** The codes are modified from ``https://github.com/amdegroot/ssd.pytorch``. Run `eval.py` code to get the object detection average precision (AP) on the PASCAL VOC 2012 dataset. The model is already trained on the PASCAL VOC 2012 object detection dataset. Draw a table in your report summarizing the AP of all 20 object categories and their mean. **[10 points]**
``Answer Q2.3 here``
2. **Answer the following questions:**
1. Briefly explain how average precision is computed for PASCAL VOC 2012 dataset. Please check the code ($\mathtt{eval.py:~Line~163-191}$). In this homework, we use the Pascal VOC 2007 metric. **[10 points]**
2. Explain how SSD can be much faster compared to Faster RCNN [4]? **[10 points]**
3. Usually the number of negative bounding boxes (boxes without any object) is much larger than the number of positive bounding boxes. Explain how this imbalance is handled in SSD and Faster RCNN, respectively. **[10 points]**
``Answer Q2.2.A here``
``Answer Q2.2.B here``
``Answer Q2.2.C here``
3. Randomly pick up some images from the PASCAL VOC 2012 dataset and some from other sources. Visualize the bounding box prediction results and include a figure in your report. You can use the code in folder $\mathtt{demo}$ for visualization. **[10 points]**
``Answer Q2.3 here``
## References
1. Yu, Fisher, and Vladlen Koltun. "Multi-scale context aggregation by dilated convolutions." arXiv preprint arXiv:1511.07122 (2015).
2. Zhao, Hengshuang, et al. "Pyramid scene parsing network." Proceedings of the IEEE conference on computer vision and pattern recognition. 2017.
3. Liu, Wei, et al. "Ssd: Single shot multibox detector." European conference on computer vision. Springer, Cham, 2016.
4. Ren, Shaoqing, et al. "Faster r-cnn: Towards real-time object detection with region proposal networks." Advances in neural information processing systems. 2015.
| github_jupyter |
# Extracting Experiment Powers
This notebook aids in the extraction of powers from the power log files.
```
import os
POWER_DIR='/home/nsultana'
```
### Scanning through power log
Pre-processing the power log to grab (approximately) the time that is used for the experiments prevents numpy from having to parse days worth of data, and reduces waiting time
```
from io import StringIO
import pandas as pd
import numpy as np
import os
def to_datetime(t):
if isinstance(t, float):
t = pd.to_datetime(t * 1e9).tz_localize('UTC')
if isinstance(t, str):
t = pd.to_datetime(t)
return t
def open_at_time(fname, start_time, end_time=None, jump_bytes=64000):
''' Jumps backwards from the end of the file looking for the first matching timestamps.
File is readable from this location.
If end_time is present, reads the contents between start_time and end_time into a file-like buffer.
'''
# Must open in binary mode to enable .seek to arbitrary locations
f = open(fname, 'rb')
start_time, end_time = to_datetime(start_time), to_datetime(end_time)
end_loc = None
for i in range(1, 100000):
f.seek(-jump_bytes * i, os.SEEK_END)
firstline = f.readline()
line = f.readline().decode('utf-8')
strtime = ' '.join(line.split(' ')[:2])
curtime = pd.to_datetime(strtime)
if curtime <= start_time:
print("Found location in file: %s"%line)
f.seek(-len(line), 1)
if end_loc is not None:
rd_size = end_loc - f.tell()
print("Reading into buffer")
sio = StringIO(f.read(rd_size).decode('utf-8'))
print("Done.")
f.close()
return sio
else:
f2 = open(fname, 'r')
f2.seek(f.tell())
f.close()
return f2
if end_time is not None and curtime > end_time:
end_loc = f.tell()
print("Jumped 1000000 times and still didn't find it")
f.close()
return open(fname, 'r')
```
### Read the power log
Power log is reduced to only the time points at which the power changes.
It is cleaned and inserted into a pandas dataframe
```
import os
def read_power_log(num, start_time, end_time):
start_time, end_time = to_datetime(start_time), to_datetime(end_time)
fname = os.path.join(POWER_DIR, 'power_%d.out' % num)
f = open_at_time(fname, start_time, end_time)
# For some reason this is infinitely faster than using read_csv
# df = pd.read_csv(f, sep=' ', error_bad_lines=True, header=None,
# names=['date', 'time', 'device', 'ip', 'power'],
# dtype={'device': str, 'ip': str, 'power': str},
# date_parser=date_parser,
# parse_dates=[[0,1]], infer_datetime_format=True, engine='c',
# low_memory=False)
# Read into a numpy array
arr = np.loadtxt(f, delimiter=' ', dtype=object)
df = pd.DataFrame(arr, columns=['date', 'time', 'device', 'ip', 'power'])
date_time = df.date.astype(object) + ' ' + df.time.astype(object)
df['date_time'] = pd.to_datetime(date_time.astype(str))
del df['date']
del df['time']
# Discard any rows with missing power values
df.power = pd.to_numeric(df.power, errors='coerce')
df = df[~df.power.isna()]
# Error on power monitors sometimes reports very high power values
df = df[df.power < 10000]
# Filter based on the provided start time
df = df[df.date_time >= pd.to_datetime(start_time)]
if end_time is not None:
df = df[df.date_time <= pd.to_datetime(end_time)]
print("Read %d power values" % len(df))
# Reduce to only the time-points where the power changes
df = df[df.power.diff() != 0]
df = df.reset_index(drop=True)
return df
def read_all_power_logs(n, start_time, end_time=None):
''' Reads power logs numbered 0:(n-1) '''
out = []
for num in range(n):
out.append(read_power_log(num, start_time, end_time))
out = list(filter(lambda x: len(x) > 1, out))
return out
def read_experiments_powers(events_sets, experiments):
''' events_sets[group][experiment][trial][event]'''
print(experiments)
mintime = min([ev[experiments[0]][0][0]['time_'] for ev in events_sets])
maxtime = max([ev[experiments[-1]][-1][-1]['time_'] for ev in events_sets])
return read_all_power_logs(10, mintime, maxtime)
```
### Labeling powers during experiment
Using shremote logs, powers are combined into a single dataframe with an added `exp_time` column, which gives the amount of elapsed time since the start of the experiment
```
EXP_LEN = pd.Timedelta(50, 's')
def experiment_powers(events, dfs, buff = pd.Timedelta(15, 's')):
starts, ends = [], []
for exp_events in events:
# NB: Assumes the start of the experiment is the /second/ `tcpreplay`
replay_events = filter(lambda x: x['name_'] == 'tcpreplay', exp_events)
start = list(replay_events)[-1]['time_']
start = pd.to_datetime(start * 1e9).tz_localize('UTC') - buff
starts.append(start)
end = start + EXP_LEN + buff * 2
ends.append(end)
minstart = min(starts)
maxend = max(ends)
dfs_exp = []
for df in dfs:
df = df[(df.date_time > minstart) & (df.date_time < maxend)]
for i, (start, end) in enumerate(zip(starts, ends)):
dfi = df[(df.date_time > start) & (df.date_time < end)]
dfi = dfi[dfi.power < 1e3]
dfi = dfi.reset_index(drop=True)
dfi['exp_time'] = (dfi.date_time - start - buff).apply(lambda x: x.total_seconds())
dfi.date_time -= start
dfi["trial"] = i
dfs_exp.append(dfi)
return pd.concat(dfs_exp, ignore_index=True)
def all_experiment_powers(events, powers, buff = pd.Timedelta(15, 'seconds')):
all_dfs = []
for exp, events in events.items():
print("Read", exp)
df = experiment_powers(events, powers)
df['experiment'] = exp
all_dfs.append(df)
return pd.concat(all_dfs, ignore_index=True)
```
### Loading experiment logs
Need to read the timestamps associated with the experiments to load and label the right data
```
import json
EXPERIMENTS = ['baseline', 'hc', 'drop', 'fec', 'kv']
def load_experiment_events(exp, n, base_dir):
''' Loads experiments 1:n labeled `exp` in directory `base_dir` '''
events = []
for i in range(1, n+1):
exp_file = os.path.join(base_dir, exp, '%s_%d' % (exp, i), 'event_log.json')
with open(exp_file) as f:
events.append(json.load(f))
return events
def load_events(base_dir, n=10, experiments=EXPERIMENTS):
events = {}
for exp in experiments:
exp_events = load_experiment_events(exp, n, base_dir)
events[exp] = exp_events
return events
```
# Preprocess and save data
```
POWER_SAVE_DIR = '.'
def save_experiment_powers(base_dirs, n=10, experiments=EXPERIMENTS):
events = {}
for label, base_dir in base_dirs.items():
events[label] = load_events(base_dir, n, experiments)
powers = read_experiments_powers(events.values(), experiments)
exps_powers = {}
for label, exp_events in events.items():
exp_powers = all_experiment_powers(exp_events, powers)
exp_powers.to_pickle(os.path.join(POWER_SAVE_DIR, '%s.pickle' % label))
exps_powers[label] = exp_powers
return exps_powers
def load_experiment_powers(label):
return pd.read_pickle(os.path.join(POWER_SAVE_DIR, '%s.pickle' % label))
save_experiment_powers(dict(
fpga = 'data/e2e_output_500k_fpga_hc_11',
tofino = 'data/e2e_output_500k_tofino_hc_11'
));
save_experiment_powers(dict(
cpu = 'data/e2e_output_500k_cpu_hc_11'
), 10, ['baseline', 'hc']);
```
| github_jupyter |
Starting point for MNIST digit classification according to Jalalvand
```
import numpy as np
from sklearn.datasets import fetch_openml
from tqdm import tqdm
from sklearn.pipeline import FeatureUnion
from sklearn.cluster import MiniBatchKMeans
from sklearn.model_selection import train_test_split, ParameterGrid
from sklearn.base import clone
from sklearn.metrics import silhouette_score, mean_squared_error, accuracy_score, classification_report, confusion_matrix, ConfusionMatrixDisplay
from joblib import dump, load, Parallel, delayed
from pyrcn.echo_state_network import ESNClassifier
from pyrcn.linear_model import IncrementalRegression, FastIncrementalRegression
from pyrcn.base import InputToNode, NodeToNode
from matplotlib import pyplot as plt
plt.rcParams['image.cmap'] = 'jet'
plt.rcParams['pdf.fonttype'] = 42
plt.rcParams['ps.fonttype'] = 42
%matplotlib inline
```
Load the dataset
```
X, y = fetch_openml('mnist_784', version=1, return_X_y=True, as_frame=False)
```
Provide standard split in training and test. Further split training set for validation. Normalize to a range between [0, 1].
Reshape to obtain 28x28 images
```
X_train, X_test = X[:60000] / 255., X[60000:] / 255.
y = y.astype(int)
y_train, y_test = y[:60000], y[60000:]
X_train, X_val, y_train, y_val = train_test_split(X_train, y_train, stratify=y_train, test_size=0.2, random_state=42)
X_train = [X_train[k].reshape(28, 28).T for k in range(len(X_train))]
X_val = [X_val[k].reshape(28, 28).T for k in range(len(X_val))]
X_test = [X_test[k].reshape(28, 28).T for k in range(len(X_test))]
plt.figure()
plt.imshow(X_train[1])
X_train_bin = [X_train[k]]
```
Provide a basic ESN with Aza's parameters
```
base_input_to_nodes = InputToNode(hidden_layer_size=16000, activation='identity', k_in=10, input_scaling=0.4, bias_scaling=0.0)
base_nodes_to_nodes = NodeToNode(hidden_layer_size=16000, spectral_radius=0.4, leakage=0.5, bias_scaling=0.3, k_rec=10)
esn = ESNClassifier(input_to_nodes=[('default', base_input_to_nodes)],
nodes_to_nodes=[('default', base_nodes_to_nodes)],
regressor=FastIncrementalRegression(alpha=1e-3), random_state=10)
```
Train this ESN
```
try:
esn = load("rand_esn_16000_uni_fast.joblib")
except FileNotFoundError:
with tqdm(total=len(X_train)) as pbar:
for X, y in zip(X_train, y_train):
y = np.repeat(y, repeats=28, axis=0)
y[np.argwhere(X.sum(axis=1)==0)] = 10
esn.partial_fit(X=X, y=y, classes=np.arange(11), update_output_weights=False)
pbar.update(1)
with tqdm(total=len(X_val)) as pbar:
for X, y in zip(X_val[:-1], y_val[:-1]):
y = np.repeat(y, repeats=28, axis=0)
y[np.argwhere(X.sum(axis=1)==0)] = 10
esn.partial_fit(X=X, y=y, update_output_weights=False)
pbar.update(1)
X = X_val[-1]
y = np.repeat(y_val[-1], repeats=28, axis=0)
y[np.argwhere(X.sum(axis=1)==0)] = 10
esn.partial_fit(X=X, y=y, update_output_weights=True)
pbar.update(1)
dump(esn, "rand_esn_16000_uni_fast.joblib")
```
Compute final measurements
```
Y_true_train = []
Y_pred_train = []
with tqdm(total=len(X_train)) as pbar:
for X, y in zip(X_train, y_train):
y_pred = esn.predict_proba(X=X)[np.argwhere(X.sum(axis=1)!=0), :-1]
Y_true_train.append(y)
Y_pred_train.append(np.argmax(y_pred.sum(axis=0)))
pbar.update(1)
with tqdm(total=len(X_val)) as pbar:
for X, y in zip(X_val, y_val):
y_pred = esn.predict_proba(X=X)[np.argwhere(X.sum(axis=1)!=0), :-1]
Y_true_train.append(y)
Y_pred_train.append(np.argmax(y_pred.sum(axis=0)))
pbar.update(1)
Y_true_test = []
Y_pred_test = []
with tqdm(total=len(X_test)) as pbar:
for X, y in zip(X_test, y_test):
y_pred = esn.predict_proba(X=X)[np.argwhere(X.sum(axis=1)!=0), :-1]
Y_true_test.append(y)
Y_pred_test.append(np.argmax(y_pred.sum(axis=0)))
pbar.update(1)
```
Confusion matrices
```
cm = confusion_matrix(Y_true_train, Y_pred_train)
cm_display = ConfusionMatrixDisplay(cm, display_labels=[0, 1, 2, 3, 4, 5, 6, 7, 8, 9]).plot()
print("Classification training report for estimator %s:\n%s\n"
% (esn, classification_report(Y_true_train, Y_pred_train, digits=4)))
plt.show()
cm = confusion_matrix(Y_true_test, Y_pred_test)
cm_display = ConfusionMatrixDisplay(cm, display_labels=[0, 1, 2, 3, 4, 5, 6, 7, 8, 9]).plot()
print("Classification test report for estimator %s:\n%s\n"
% (esn, classification_report(Y_true_test, Y_pred_test, digits=4)))
plt.show()
```
Provide a basic ESN with Aza's parameters and train this ESN
```
kmeans = MiniBatchKMeans(n_clusters=8000, n_init=20, reassignment_ratio=0, max_no_improvement=50, init='k-means++', verbose=2, random_state=2)
kmeans.fit(X=np.concatenate(X_train + X_val))
base_input_to_nodes = InputToNode(hidden_layer_size=8000, activation='identity', k_in=10, input_scaling=0.5, bias_scaling=0.0, random_state=10)
base_nodes_to_nodes = NodeToNode(hidden_layer_size=8000, spectral_radius=0.8, leakage=0.2, bias_scaling=0.1, k_rec=10, random_state=10)
base_reg = FastIncrementalRegression(alpha=1e-3)
w_in = np.divide(kmeans.cluster_centers_, np.linalg.norm(kmeans.cluster_centers_, axis=1)[:, None])
# w_in = np.pad(np.divide(kmeans.cluster_centers_, np.linalg.norm(kmeans.cluster_centers_, axis=1)[:, None]), ((0, 1600), (0, 0)), mode='constant', constant_values=0)
base_input_to_nodes.fit(X=X_train[0])
base_input_to_nodes._input_weights = w_in.T
esn = ESNClassifier(input_to_nodes=[('default', base_input_to_nodes)],
nodes_to_nodes=[('default', base_nodes_to_nodes)],
regressor=base_reg)
esn._input_to_node =FeatureUnion(transformer_list=[('default', base_input_to_nodes)], n_jobs=None, transformer_weights=None).fit(X_train[0])
esn._input_to_node.transformer_list[0][1]._input_weights = w_in.T
try:
esn = load("kmeans_esn_8000_uni_fast.joblib")
except FileNotFoundError:
with tqdm(total=len(X_train)) as pbar:
for X, y in zip(X_train, y_train):
y = np.repeat(y, repeats=28, axis=0)
y[np.argwhere(X.sum(axis=1)==0)] = 10
esn.partial_fit(X=X, y=y, classes=np.arange(11))
pbar.update(1)
with tqdm(total=len(X_val)) as pbar:
for X, y in zip(X_val[:-1], y_val[:-1]):
y = np.repeat(y, repeats=28, axis=0)
y[np.argwhere(X.sum(axis=1)==0)] = 10
esn.partial_fit(X=X, y=y, classes=np.arange(11), update_output_weights=False)
pbar.update(1)
X = X_val[-1]
y = np.repeat(y_val[-1], repeats=28, axis=0)
y[np.argwhere(X.sum(axis=1)==0)] = 10
esn.partial_fit(X=X, y=y, update_output_weights=True)
pbar.update(1)
dump(esn, "kmeans_esn_8000_uni_fast.joblib")
```
Compute final measurements
```
Y_true_train = []
Y_pred_train = []
with tqdm(total=len(X_train)) as pbar:
for X, y in zip(X_train, y_train):
y_pred = esn.predict_proba(X=X)[np.argwhere(X.sum(axis=1)!=0), :-1]
Y_true_train.append(y)
Y_pred_train.append(np.argmax(y_pred.sum(axis=0)))
pbar.update(1)
with tqdm(total=len(X_val)) as pbar:
for X, y in zip(X_val, y_val):
y_pred = esn.predict_proba(X=X)[np.argwhere(X.sum(axis=1)!=0), :-1]
Y_true_train.append(y)
Y_pred_train.append(np.argmax(y_pred.sum(axis=0)))
pbar.update(1)
Y_true_test = []
Y_pred_test = []
with tqdm(total=len(X_test)) as pbar:
for X, y in zip(X_test, y_test):
y_pred = esn.predict_proba(X=X)[np.argwhere(X.sum(axis=1)!=0), :-1]
Y_true_test.append(y)
Y_pred_test.append(np.argmax(y_pred.sum(axis=0)))
pbar.update(1)
```
Confusion matrices
```
cm = confusion_matrix(Y_true_train, Y_pred_train)
cm_display = ConfusionMatrixDisplay(cm, display_labels=[0, 1, 2, 3, 4, 5, 6, 7, 8, 9]).plot()
print("Classification training report for estimator %s:\n%s\n"
% (esn, classification_report(Y_true_train, Y_pred_train, digits=4)))
plt.show()
cm = confusion_matrix(Y_true_test, Y_pred_test)
cm_display = ConfusionMatrixDisplay(cm, display_labels=[0, 1, 2, 3, 4, 5, 6, 7, 8, 9]).plot()
print("Classification test report for estimator %s:\n%s\n"
% (esn, classification_report(Y_true_test, Y_pred_test, digits=4)))
plt.show()
```
| github_jupyter |
```
# Copyright 2021 Google LLC
# Use of this source code is governed by an MIT-style
# license that can be found in the LICENSE file or at
# https://opensource.org/licenses/MIT.
# Notebook authors: Kevin P. Murphy (murphyk@gmail.com)
# and Mahmoud Soliman (mjs@aucegypt.edu)
# This notebook reproduces figures for chapter 13 from the book
# "Probabilistic Machine Learning: An Introduction"
# by Kevin Murphy (MIT Press, 2021).
# Book pdf is available from http://probml.ai
```
<a href="https://opensource.org/licenses/MIT" target="_parent"><img src="https://img.shields.io/github/license/probml/pyprobml"/></a>
<a href="https://colab.research.google.com/github/probml/pml-book/blob/main/pml1/figure_notebooks/chapter13_neural_networks_for_structured_data_figures.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
## Figure 13.1:<a name='13.1'></a> <a name='xor'></a>
(a) Illustration of the fact that the XOR function is not linearly separable, but can be separated by the two layer model using Heaviside activation functions. Adapted from Figure 10.6 of <a href='#Geron2019'>[Aur19]</a> .
Figure(s) generated by [xor_heaviside.py](https://github.com/probml/pyprobml/blob/master/scripts/xor_heaviside.py)
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
from deimport.deimport import deimport
print('finished!')
deimport(superimport)
%run xor_heaviside.py
```
## Figure 13.2:<a name='13.2'></a> <a name='activationFns2'></a>
(a) Illustration of how the sigmoid function is linear for inputs near 0, but saturates for large positive and negative inputs. Adapted from 11.1 of <a href='#Geron2019'>[Aur19]</a> . (b) Plots of some neural network activation functions.
Figure(s) generated by [activation_fun_plot.py](https://github.com/probml/pyprobml/blob/master/scripts/activation_fun_plot.py)
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
from deimport.deimport import deimport
print('finished!')
deimport(superimport)
%run activation_fun_plot.py
```
## Figure 13.3:<a name='13.3'></a> <a name='mlp-playground'></a>
An MLP with 2 hidden layers applied to a set of 2d points from 2 classes, shown in the top left corner. The visualizations associated with each hidden unit show the decision boundary at that part of the network. The final output is shown on the right. The input is $ \bm x \in \mathbb R ^2$, the first layer activations are $ \bm z _1 \in \mathbb R ^4$, the second layer activations are $ \bm z _2 \in \mathbb R ^2$, and the final logit is $a_3 \in \mathbb R $, which is converted to a probability using the sigmoid function. This is a screenshot from the interactive demo at http://playground.tensorflow.org
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
from deimport.deimport import deimport
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_13.3.png" width="256"/>
## Figure 13.4:<a name='13.4'></a> <a name='mlpMnist'></a>
Results of applying an MLP (with 2 hidden layers with 128 units and 1 output layer with 10 units) to some MNIST images (cherry picked to include some errors). Red is incorrect, blue is correct. (a) After 1 epoch of training. (b) After 2 epochs.
To reproduce this figure, click the open in colab button: <a href="https://colab.research.google.com/github/probml/probml-notebooks/blob/master/notebooks/mlp_mnist_tf.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
from deimport.deimport import deimport
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_13.4_A.png" width="256"/>
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_13.4_B.png" width="256"/>
## Figure 13.5:<a name='13.5'></a> <a name='twoHeaded'></a>
Illustration of an MLP with a shared ``backbone'' and two output ``heads'', one for predicting the mean and one for predicting the variance. From https://brendanhasz.github.io/2019/07/23/bayesian-density-net.html . Used with kind permission of Brendan Hasz
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
from deimport.deimport import deimport
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_13.5.png" width="256"/>
## Figure 13.6:<a name='13.6'></a> <a name='twoHeadedSineWaves'></a>
Illustration of predictions from an MLP fit using MLE to a 1d regression dataset with growing noise. (a) Output variance is input-dependent, as in \cref fig:twoHeaded . (b) Mean is computed using same model as in (a), but output variance is treated as a fixed parameter $\sigma ^2$, which is estimated by MLE after training, as in \cref sec:linregSigmaMLE .
To reproduce this figure, click the open in colab button: <a href="https://colab.research.google.com/github/probml/probml-notebooks/blob/master/notebooks/mlp_1d_regression_hetero_tfp.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
from deimport.deimport import deimport
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_13.6_A.png" width="256"/>
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_13.6_B.png" width="256"/>
## Figure 13.7:<a name='13.7'></a> <a name='reluPolytope2d'></a>
A decomposition of $\mathbb R ^2$ into a finite set of linear decision regions produced by an MLP with \ensuremath \mathrm ReLU \xspace activations with (a) one hidden layer of 25 hidden units and (b) two hidden layers. From Figure 1 of <a href='#Hein2019'>[MMJ19]</a> . Used with kind permission of Maksym Andriuschenko
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
from deimport.deimport import deimport
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_13.7_A.png" width="256"/>
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_13.7_B.png" width="256"/>
## Figure 13.8:<a name='13.8'></a> <a name='axons'></a>
Illustration of two neurons connected together in a ``circuit''. The output axon of the left neuron makes a synaptic connection with the dendrites of the cell on the right. Electrical charges, in the form of ion flows, allow the cells to communicate. From https://en.wikipedia.org/wiki/Neuron . Used with kind permission of Wikipedia author BruceBlaus
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
from deimport.deimport import deimport
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_13.8.png" width="256"/>
## Figure 13.9:<a name='13.9'></a> <a name='DNN-size-vs-time'></a>
Plot of neural network sizes over time. Models 1, 2, 3 and 4 correspond to the perceptron <a href='#Rosenblatt58'>[Ros58]</a> , the adaptive linear unit <a href='#Widrow1960'>[BH60]</a> the neocognitron <a href='#Fukushima1980'>[K80]</a> , and the first MLP trained by backprop <a href='#Rumelhart86'>[RHW86]</a> . Approximate number of neurons for some living organisms are shown on the right scale (the sponge has 0 neurons), based on https://en.wikipedia.org/wiki/List_of_animals_by_number_of_neurons . From Figure 1.11 of <a href='#GoodfellowBook'>[GBC16]</a> . Used with kind permission of Ian Goodfellow
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
from deimport.deimport import deimport
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_13.9.png" width="256"/>
## Figure 13.10:<a name='13.10'></a> <a name='feedforward-graph'></a>
A simple linear-chain feedforward model with 4 layers. Here $ \bm x $ is the input and $ \bm o $ is the output. From <a href='#Blondel2020'>[Mat20]</a>
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
from deimport.deimport import deimport
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_13.10.png" width="256"/>
## Figure 13.11:<a name='13.11'></a> <a name='computation-graph'></a>
An example of a computation graph with 2 (scalar) inputs and 1 (scalar) output. From <a href='#Blondel2020'>[Mat20]</a>
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
from deimport.deimport import deimport
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_13.11.png" width="256"/>
## Figure 13.12:<a name='13.12'></a> <a name='backwardsDiff'></a>
Notation for automatic differentiation at node $j$ in a computation graph. From <a href='#Blondel2020'>[Mat20]</a>
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
from deimport.deimport import deimport
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_13.12.png" width="256"/>
## Figure 13.13:<a name='13.13'></a> <a name='compGraphD2l'></a>
Computation graph for an MLP with input $ \bm x $, hidden layer $ \bm h $, output $ \bm o $, loss function $L=\ell ( \bm o ,y)$, an $\ell _2$ regularizer $s$ on the weights, and total loss $J=L+s$. From Figure 4.7.1 of <a href='#dive'>[Zha+20]</a> . Used with kind permission of Aston Zhang
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
from deimport.deimport import deimport
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_13.13.png" width="256"/>
## Figure 13.14:<a name='13.14'></a> <a name='activationWithGrad'></a>
(a) Some popular activation functions. (b) Plot of their gradients.
To reproduce this figure, click the open in colab button: <a href="https://colab.research.google.com/github/probml/probml-notebooks/blob/master/notebooks/activation_fun_deriv_torch.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
from deimport.deimport import deimport
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_13.14_A.png" width="256"/>
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_13.14_B.png" width="256"/>
## Figure 13.15:<a name='13.15'></a> <a name='residualVanishing'></a>
(a) Illustration of a residual block. (b) Illustration of why adding residual connections can help when training a very deep model. Adapted from Figure 14.16 of <a href='#Geron2019'>[Aur19]</a>
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
from deimport.deimport import deimport
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_13.15_A.png" width="256"/>
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_13.15_B.png" width="256"/>
## Figure 13.16:<a name='13.16'></a> <a name='multiGPU'></a>
Calculation of minibatch stochastic gradient using data parallelism and two GPUs. From Figure 12.5.2 of <a href='#dive'>[Zha+20]</a> . Used with kind permission of Aston Zhang
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
from deimport.deimport import deimport
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_13.16.png" width="256"/>
## Figure 13.17:<a name='13.17'></a> <a name='sparseNnet'></a>
(a) A deep but sparse neural network. The connections are pruned using $\ell _1$ regularization. At each level, nodes numbered 0 are clamped to 1, so their outgoing weights correspond to the offset/bias terms. (b) Predictions made by the model on the training set.
To reproduce this figure, click the open in colab button: <a href="https://colab.research.google.com/github/probml/probml-notebooks/blob/master/notebooks/sparse_mlp.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
from deimport.deimport import deimport
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_13.17_A.png" width="256"/>
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_13.17_B.png" width="256"/>
## Figure 13.18:<a name='13.18'></a> <a name='dropout'></a>
Illustration of dropout. (a) A standard neural net with 2 hidden layers. (b) An example of a thinned net produced by applying dropout with $p_0=0.5$. Units that have been dropped out are marked with an x. From Figure 1 of <a href='#Srivastava2014'>[Nit+14]</a> . Used with kind permission of Geoff Hinton
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
from deimport.deimport import deimport
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_13.18_A.png" width="256"/>
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_13.18_B.png" width="256"/>
## Figure 13.19:<a name='13.19'></a> <a name='flatMinima'></a>
Flat vs sharp minima. From Figures 1 and 2 of <a href='#Hochreiter1997'>[SJ97]</a> . Used with kind permission of Jürgen Schmidhuber
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
from deimport.deimport import deimport
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_13.19.png" width="256"/>
## Figure 13.20:<a name='13.20'></a> <a name='sgd-minima-unstable'></a>
Each curve shows how the loss varies across parameter values for a given minibatch. (a) A stable local minimum. (b) An unstable local minimum.
To reproduce this figure, click the open in colab button: <a href="https://colab.research.google.com/github/probml/probml-notebooks/blob/master/notebooks/sgd_minima_variance.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
from deimport.deimport import deimport
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_13.20_A.png" width="256"/>
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_13.20_B.png" width="256"/>
## Figure 13.21:<a name='13.21'></a> <a name='xorRBF'></a>
(a) xor truth table. (b) Fitting a linear logistic regression classifier using degree 10 polynomial expansion. (c) Same model, but using an RBF kernel with centroids specified by the 4 black crosses.
Figure(s) generated by [logregXorDemo.py](https://github.com/probml/pyprobml/blob/master/scripts/logregXorDemo.py)
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
from deimport.deimport import deimport
print('finished!')
deimport(superimport)
%run logregXorDemo.py
```
## Figure 13.22:<a name='13.22'></a> <a name='rbfDemo'></a>
Linear regression using 10 equally spaced RBF basis functions in 1d. Left column: fitted function. Middle column: basis functions evaluated on a grid. Right column: design matrix. Top to bottom we show different bandwidths for the kernel function: $\sigma =0.5, 10, 50$.
Figure(s) generated by [linregRbfDemo.py](https://github.com/probml/pyprobml/blob/master/scripts/linregRbfDemo.py)
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
from deimport.deimport import deimport
print('finished!')
deimport(superimport)
%run linregRbfDemo.py
```
## Figure 13.23:<a name='13.23'></a> <a name='mixexp'></a>
(a) Some data from a one-to-many function. (b) The responsibilities of each expert for the input domain. (c) Prediction of each expert. (d) Overeall prediction. Mean is red cross, mode is black square. Adapted from Figures 5.20 and 5.21 of <a href='#BishopBook'>[Bis06]</a> .
Figure(s) generated by [mixexpDemoOneToMany.py](https://github.com/probml/pyprobml/blob/master/scripts/mixexpDemoOneToMany.py)
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
from deimport.deimport import deimport
print('finished!')
deimport(superimport)
%run mixexpDemoOneToMany.py
```
## Figure 13.24:<a name='13.24'></a> <a name='deepMOE'></a>
Deep MOE with $m$ experts, represented as a neural network. From Figure 1 of <a href='#Chazan2017'>[SJS17]</a> . Used with kind permission of Jacob Goldberger
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
from deimport.deimport import deimport
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_13.24.png" width="256"/>
## Figure 13.25:<a name='13.25'></a> <a name='HMENN'></a>
A 2-level hierarchical mixture of experts as a neural network. The top gating network chooses between the left and right expert, shown by the large boxes; the left and right experts themselves choose between their left and right sub-experts
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
from deimport.deimport import deimport
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_13.25.png" width="256"/>
## References:
<a name='Geron2019'>[Aur19]</a> G. Aur'elien "Hands-On Machine Learning with Scikit-Learn and TensorFlow: Concepts, Tools, and Techniques for BuildingIntelligent Systems (2nd edition)". (2019).
<a name='Widrow1960'>[BH60]</a> W. Bernard and H. HoffMarcianE. "Adaptive Switching Circuits". (1960).
<a name='BishopBook'>[Bis06]</a> C. Bishop "Pattern recognition and machine learning". (2006).
<a name='GoodfellowBook'>[GBC16]</a> I. Goodfellow, Y. Bengio and A. Courville. "Deep Learning". (2016).
<a name='Fukushima1980'>[K80]</a> F. K "Neocognitron: a self organizing neural network model for amechanism of pattern recognition unaffected by shift in position". In: Biol. Cybern. (1980).
<a name='Hein2019'>[MMJ19]</a> H. Matthias, A. Maksym and B. Julian. "Why ReLU networks yield high-confidence predictions far awayfrom the training data and how to mitigate the problem". (2019).
<a name='Blondel2020'>[Mat20]</a> B. Mathieu "Automatic differentiation". (2020).
<a name='Srivastava2014'>[Nit+14]</a> S. Nitish, H. GeoffRey, K. Alex, S. Ilya and S. Ruslan. "Dropout: A Simple Way to Prevent Neural Networks from Over tting". In: jmlr (2014).
<a name='Rumelhart86'>[RHW86]</a> D. Rumelhart, G. Hinton and R. Williams. "Learning internal representations by error propagation". (1986).
<a name='Rosenblatt58'>[Ros58]</a> F. Rosenblatt "The Perceptron: A Probabilistic Model for Information Storage and Organization in the Brain". In: Psychological Review (1958).
<a name='Hochreiter1997'>[SJ97]</a> H. S and S. J. "Flat minima". In: Neural Comput. (1997).
<a name='Chazan2017'>[SJS17]</a> C. ShlomoE, G. Jacob and G. Sharon. "Speech Enhancement using a Deep Mixture of Experts". abs/1703.09302 (2017). arXiv: 1703.09302
<a name='dive'>[Zha+20]</a> A. Zhang, Z. Lipton, M. Li and A. Smola. "Dive into deep learning". (2020).
| github_jupyter |
```
import cv2 as cv
from glob import glob
import os
import numpy as np
from utils.poincare import calculate_singularities
from utils.segmentation import create_segmented_and_variance_images
from utils.normalization import normalize
from utils.gabor_filter import gabor_filter
from utils.frequency import ridge_freq
from utils import orientation
from utils.crossing_number import calculate_minutiaes
from utils.skeletonize import skeletonize
from MyMAS import MAS
def fingerprint_pipline(input_img):
block_size = 16
# pipe line picture re https://www.cse.iitk.ac.in/users/biometrics/pages/111.JPG
# normalization -> orientation -> frequency -> mask -> filtering
# normalization - removes the effects of sensor noise and finger pressure differences.
normalized_img = normalize(input_img.copy(), float(100), float(100))
# color threshold
# threshold_img = normalized_img
# _, threshold_im = cv.threshold(normalized_img,127,255,cv.THRESH_OTSU)
# cv.imshow('color_threshold', normalized_img); cv.waitKeyEx()
# ROI and normalisation
(segmented_img, normim, mask) = create_segmented_and_variance_images(normalized_img, block_size, 0.2)
# orientations
angles = orientation.calculate_angles(normalized_img, W=block_size, smoth=False)
orientation_img = orientation.visualize_angles(segmented_img, mask, angles, W=block_size)
# find the overall frequency of ridges in Wavelet Domain
freq = ridge_freq(normim, mask, angles, block_size, kernel_size=5, minWaveLength=5, maxWaveLength=15)
# create gabor filter and do the actual filtering
gabor_img = gabor_filter(normim, angles, freq)
# thinning oor skeletonize
thin_image = skeletonize(gabor_img)
# minutias
minutias, end, bif = calculate_minutiaes(thin_image)
# singularities
singularities_img, FingerType = calculate_singularities(thin_image, angles, 1, block_size, mask)
# visualize pipeline stage by stage
# output_imgs = [input_img, normalized_img, segmented_img, orientation_img, gabor_img, thin_image, minutias, singularities_img]
output_imgs = [input_img, minutias, singularities_img]
# for i in range(len(output_imgs)): # Lặp qua từng ảnh output
# if len(output_imgs[i].shape) == 2: # Nếu ảnh thứ i trong output là ảnh Gray(có 2 chiều)
# output_imgs[i] = cv.cvtColor(output_imgs[i], cv.COLOR_GRAY2RGB) # Chuyển thành ảnh RGB để concatenate với 2 ảnh cuối
# Chuyển list các ảnh thành ảnh 4x4
# results = np.concatenate([np.concatenate(output_imgs[:4], 1), np.concatenate(output_imgs[4:], 1)]).astype(np.uint8)
# return results
return output_imgs, end, bif, FingerType
def DISTANCE(U, V) -> float:
"""
"""
return np.sqrt((U[0] - V[0])**2 + (U[1] - V[1])**2)
def Points_distance(points1, points2) -> list:
n = points1.shape[0]
m = points2.shape[0]
Distance = np.zeros((n, m))
for i in range(n):
for j in range(m):
Distance[i, j] = DISTANCE(points1[i], points2[j])
return Distance
def Filter_Points(ListPoints, CenterPoint, size = (0,0)):
"""
"""
n = ListPoints.shape[0]
ResultPoints = []
for i in range(n):
distance = np.abs(CenterPoint - ListPoints[i])
if (distance[0] <= size[0]/2) and (distance[1] <= size[1]/2):
ResultPoints.append(ListPoints[i].tolist())
return np.array(ResultPoints)
def Transtation_Finger(Minutiae, Trans_Vector) -> np.ndarray:
"""
"""
return Minutiae + Trans_Vector
def MATCH(finger1, finger2, alpha=0.5, box_size=(0, 0)) -> bool:
"""
"""
# Get feature
center_point1, end_1, bif_1 = finger1[0], finger1[1], finger1[2]
center_point2, end_2, bif_2 = finger2[0], finger2[1], finger2[2]
translation_vector = center_point2 - center_point1
# Translation minutiaes
end_1 = Transtation_Finger(end_1, translation_vector)
bif_1 = Transtation_Finger(bif_1, translation_vector)
# Filter minutiaes
new_end_1 = Filter_Points(end_1, center_point2, size=box_size)
new_bif_1 = Filter_Points(bif_1, center_point2, size=box_size)
new_end_2 = Filter_Points(end_2, center_point2, size=box_size)
new_bif_2 = Filter_Points(bif_2, center_point2, size=box_size)
n1_end = new_end_1.shape[0]
n1_bif = new_bif_1.shape[0]
n2_end = new_end_2.shape[0]
n2_bif = new_bif_2.shape[0]
# Match ending points
Distance_end = Points_distance(new_end_1, new_end_2)
end_min_dist = Distance_end.min(axis=0)
end_min_index = Distance_end.argmin(axis=0)
# Match bifucation points
Distance_bif = Points_distance(new_bif_1, new_bif_2)
bif_min_dist = Distance_bif.min(axis=0)
bif_min_index = Distance_bif.argmin(axis=0)
# Check Matching
end_matched = np.where(end_min_dist <= 15)[0].shape[0]
bif_matched = np.where(bif_min_dist <= 15)[0].shape[0]
if (end_matched + bif_matched)/(n1_end + n1_bif) >= alpha:
return True
else:
return False
box_size = (150, 300)
# MATCH(images[0], images[2], box_size=(150, 300))
def Draw_Result():
"""
"""
pass
def FeaturesExtraction(finger):
"""
"""
imgresult, end, bif, fingertype = fingerprint_pipline(finger)
center_point = np.array(fingertype)[:,1:].astype(int).mean(axis=0)
end = np.array(end)[:,1:]
bif = np.array(bif)[:,1:]
return [center_point, end, bif]
# finger1 = FeaturesExtraction(images[0])
# finger2 = FeaturesExtraction(images[3])
# MATCH(finger1, finger2, box_size=(150,300))
def S_Generation(PIN, lenMask) -> str:
"""
"""
S = bin(PIN)[2:]
n = len(S)
k = lenMask//n
return S*k + S[:lenMask - n*k]
def Features2Msg(features):
"""
"""
center, end, bif = features[0].astype(int), features[1].astype(int), features[2].astype(int)
center_msg = "{:3d}{:3d}".format(center[0], center[1]).replace(' ', '0')
n = end.shape[0]
m = bif.shape[0]
end_msg = ''
for point in end:
end_msg += "{:3}{:3}".format(point[0], point[1]).replace(' ', '0')
bif_msg = ''
for point in bif:
bif_msg += "{:3}{:3}".format(point[0], point[1]).replace(' ', '0')
return "{}{:3}{:3}{}{}".format(center_msg, n, m, end_msg, bif_msg).replace(' ', '0')
#Features2Msg(finger1)
def EncryptFinger(finger, PIN, A, B, X, Code, lenMask, k, padWord):
"""
"""
# generation S
# S = S_Generation(PIN, lenMask)
S = b'10001101101011101000100011011010111010001110100011101000'
print('generation Mask successful!')
Msg = Features2Msg(finger)
print('generation Msg successful!')
cryptosystem = MAS(A, B, Code, X, S, k, padWord)
EMsg = cryptosystem.Encode(Msg)
print('Encode successful!')
return EMsg
def Emsg2Features(EMsg, PIN, A, B, X, Code, lenMask, k, padWord):
"""
"""
end = []
bif = []
S = b'10001101101011101000100011011010111010001110100011101000'
print('generation Mask successful!')
cryptosystem = MAS(A, B, Code, X, S, k, padWord)
Result = "".join(cryptosystem.Decode(EMsg))
CenterPoint = np.array([Result[:3], Result[3:6]]).astype(int)
n, m = int(Result[6:9]), int(Result[9: 12])
t = 12
for i in range(n):
end.append([int(Result[t: t+3]), int(Result[t+3: t+6])])
t += 6
for i in range(m):
bif.append([int(Result[t: t+3]), int(Result[t+3: t+6])])
t += 6
return [np.array(CenterPoint), np.array(end), np.array(bif)]
# PIN = 999999
# S = S_Generation(PIN, 56)
# print(S, '\n', len(S))
# Config Cryptosystem
PIN = 123456
lenMask = 56
# Example Initialization
A = ['0', '1', '2', '3', '4', '5', '6', '7', '8', '9']
B = ['a', 'b', 'c', 'd', 'e', 'f', 'g', 'h', 'i', 'j', 'k', 'l', 'm', 'n','o', 'p']
Code = {'a': b'1000',
'b': b'1110',
'c': b'0011',
'd': b'1111',
'e': b'1101',
'f': b'0010',
'g': b'1100',
'h': b'0101',
'i': b'1011',
'j': b'0000',
'k': b'1001',
'l': b'0111',
'm': b'0100',
'n': b'1010',
'o': b'0001',
'p': b'0110'}
# Language
X0 = ['a', 'cgh']
X1 = ['egm', 'nmc']
X2 = ['ig', 'fce']
X3 = ['jkd']
X4 = ['bea', 'mok']
X5 = ['fno', 'ihc']
X6 = ['cei']
X7 = ['demc', 'khm']
X8 = ['lbkh']
X9 = ['kog', 'dcef']
X = [X0, X1, X2, X3, X4, X5, X6, X7, X8, X9]
paddingWord = 'p'
k = 3
finger_path1 = './sample_inputs/101_1.tif'
image1 = cv.imread(finger_path1, 0)
finger_origin = FeaturesExtraction(image1)
EMsg = EncryptFinger(finger_origin, PIN, A, B, X, Code, lenMask, k, paddingWord)
finger_saved = Emsg2Features(EMsg, PIN, A, B, X, Code, lenMask, k, paddingWord)
finger_path2 = './sample_inputs/101_3.tif'
image2 = cv.imread(finger_path2, 0)
finger_match = FeaturesExtraction(image2)
MATCH(finger_saved, finger_match, box_size=(150, 300))
```
| github_jupyter |
# The Panorama Dataset
The dataset is from: https://www.kaggle.com/imdevskp/corona-virus-report
This shows the Coronavirus cases worldwide from 01/22/2020 to 07/27/2020. Below are some questions I came up with regarding it.
```
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
%matplotlib inline
data = pd.read_csv('covid.csv')
data.index = data['Date']
data.drop(columns = 'Date', inplace = True)
```
Here are the first ten days from the data:
```
data.head(10)
```
Observations:
- The comfirmed, active, and recovered cases increase steadily every day.
- The new cases and deaths are inconsistent.
- The deaths per 100 cases and recovered per 100 cases appear to be pretty consistent.
- The deaths per 100 individuals recovered are inconsistent.
- The number of countries steadily increase.
- Nothing is linear.
- Early stages of the pandemic.
Here are the last ten days from the data:
```
data.tail(10)
```
Observations:
- The confirmed, active, and recovered cases increase drastically every day.
- Each day numerous people die.
- New deaths are inconsistent.
- New recovered is inconsistent.
- Everything else is consistent.
- Seven months in the pandemic and the data has changed rapidly.
```
sns.displot(data['Confirmed'])
sns.displot(data['Deaths'])
sns.displot(data['Recovered'])
```
### What is the average for comfirmed, recovered, and active cases? What does this portray?
- The average comfirmed cases is 4406960 cases.
- The average recovered cases is 2066001 cases.
- The average active cases is 2110188 cases.
The indicates that there are more people infected than recovered.
```
data['Confirmed'].mean()
data['Recovered'].mean()
data['Active'].mean()
```
### When did the confirmed cases surpass ten million?
There were more than ten million confirmed cases starting 06-28-2020.
```
data[data['Confirmed'] > 10000000]
```
### When did we surpass 100 thousand deaths?
There were over 100,000 deaths on 04/09/2020
```
data[data['Deaths'] > 100000]
```
### What was the highest instance for deaths per 100 cases? When?
The highest instance were 7 deaths per 100 cases. There are quite a few dates.
```
data['Deaths / 100 Cases'].max()
data[data['Deaths / 100 Cases'] >= 7]
```
### In this dataset, the Coronavirus hit 187 countries. When did this occur?
On 05-12-2020, COVID-19 affected 187 countries.
```
data[data['No. of countries'] == 187]
```
### What is the death to recovered ratio?
The death to recovered ratio is 230770:2066001. About one death to ten recovered individuals.
```
data['Deaths'].mean()
data['Recovered'].mean()
```
## Some charts showing everything
```
data.plot()
data.plot(kind='bar')
sns.pairplot(data)
sns.heatmap(data.corr())
```
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.