text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
## Recap
Here's the code you've written so far.
```
# Code you have previously used to load data
import pandas as pd
from sklearn.metrics import mean_absolute_error
from sklearn.model_selection import train_test_split
from sklearn.tree import DecisionTreeRegressor
# Path of the file to read
iowa_file_path = '../input/home-data-for-ml-course/train.csv'
home_data = pd.read_csv(iowa_file_path)
# Create target object and call it y
y = home_data.SalePrice
# Create X
features = ['LotArea', 'YearBuilt', '1stFlrSF', '2ndFlrSF', 'FullBath', 'BedroomAbvGr', 'TotRmsAbvGrd']
X = home_data[features]
# Split into validation and training data
train_X, val_X, train_y, val_y = train_test_split(X, y, random_state=1)
# Specify Model
iowa_model = DecisionTreeRegressor(random_state=1)
# Fit Model
iowa_model.fit(train_X, train_y)
# Make validation predictions and calculate mean absolute error
val_predictions = iowa_model.predict(val_X)
val_mae = mean_absolute_error(val_predictions, val_y)
print("Validation MAE when not specifying max_leaf_nodes: {:,.0f}".format(val_mae))
# Using best value for max_leaf_nodes
iowa_model = DecisionTreeRegressor(max_leaf_nodes=100, random_state=1)
iowa_model.fit(train_X, train_y)
val_predictions = iowa_model.predict(val_X)
val_mae = mean_absolute_error(val_predictions, val_y)
print("Validation MAE for best value of max_leaf_nodes: {:,.0f}".format(val_mae))
# Set up code checking
from learntools.core import binder
binder.bind(globals())
from learntools.machine_learning.ex6 import *
print("\nSetup complete")
```
# Exercises
Data science isn't always this easy. But replacing the decision tree with a Random Forest is going to be an easy win.
## Step 1: Use a Random Forest
```
from sklearn.ensemble import RandomForestRegressor
# Define the model. Set random_state to 1
rf_model = RandomForestRegressor(random_state=1)
# fit your model
rf_model.fit(train_X, train_y)
# Calculate the mean absolute error of your Random Forest model on the validation data
rf_val_predictions = rf_model.predict(val_X)
rf_val_mae = mean_absolute_error(rf_val_predictions, val_y)
print("Validation MAE for Random Forest Model: {:,.0f}".format(rf_val_mae))
step_1.check()
# The lines below will show you a hint or the solution.
# step_1.hint()
step_1.solution()
```
# Think about Your Results
Under what circumstances might you prefer the Decision Tree to the Random Forest, even though the Random Forest generally gives more accurate predictions? Weigh in or follow the discussion in [this discussion thread](https://kaggle.com/learn-forum/----) **TODO: Add the link**
# Keep Going
So far, you have followed specific instructions at each step of your project. This helped learn key ideas and build your first model, but now you know enough to try things on your own.
Machine Learning competitions are a great way to try your own ideas and learn more as you independently navigate a machine learning project. Learn **[how to submit your work to a Kaggle competition](https://www.kaggle.com/dansbecker/submitting-from-a-kernel)**.
---
**[Course Home Page](https://www.kaggle.com/learn/machine-learning)**
| github_jupyter |
<a href="https://colab.research.google.com/github/SauravMaheshkar/trax/blob/SauravMaheshkar-example-1/examples/trax_data_Explained.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
#@title
# Copyright 2020 Google LLC.
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
# https://www.apache.org/licenses/LICENSE-2.0
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
## Install the Latest Version of Trax
!pip install --upgrade trax
```
Notebook Author: [@SauravMaheshkar](https://github.com/SauravMaheshkar)
# Introduction
```
import trax
```
# Serial Fn
In Trax, we use combinators to build input pipelines, much like building deep learning models. The `Serial` combinator applies layers serially using function composition and uses stack semantics to manage data.
Trax has the following definition for a `Serial` combinator.
> ```
def Serial(*fns):
def composed_fns(generator=None):
for f in fastmath.tree_flatten(fns):
generator = f(generator)
return generator
return composed_fns
```
The `Serial` function has the following structure:
* It takes as **input** arbitrary number of functions
* Convert the structure into lists
* Iterate through the list and apply the functions Serially
---
The [`fastmath.tree_flatten()`](https://github.com/google/trax/blob/c38a5b1e4c5cfe13d156b3fc0bfdb83554c8f799/trax/fastmath/numpy.py#L195) function, takes a tree as a input and returns a flattened list. This way we can use various generator functions like Tokenize and Shuffle, and apply them serially by '*iterating*' through the list.
Initially, we've defined `generator` to `None`. Thus, in the first iteration we have no input and thus the first step executes the first function in our tree structure. In the next iteration, the `generator` variable is updated to be the output of the next function in the list.
# Log Function
> ```
def Log(n_steps_per_example=1, only_shapes=True):
def log(stream):
counter = 0
for example in stream:
item_to_log = example
if only_shapes:
item_to_log = fastmath.nested_map(shapes.signature, example)
if counter % n_steps_per_example == 0:
logging.info(str(item_to_log))
print(item_to_log)
counter += 1
yield example
return log
Every Deep Learning Framework needs to have a logging component for efficient debugging.
`trax.data.Log` generator uses the `absl` package for logging. It uses a [`fastmath.nested_map`](https://github.com/google/trax/blob/c38a5b1e4c5cfe13d156b3fc0bfdb83554c8f799/trax/fastmath/numpy.py#L80) function that maps a certain function recursively inside a object. In the case depicted below, the function maps the `shapes.signature` recursively inside the input stream, thus giving us the shapes of the various objects in our stream.
--
The following two cells show the difference between when we set the `only_shapes` variable to `False`
```
data_pipeline = trax.data.Serial(
trax.data.TFDS('imdb_reviews', keys=('text', 'label'), train=True),
trax.data.Tokenize(vocab_dir='gs://trax-ml/vocabs/', vocab_file='en_8k.subword', keys=[0]),
trax.data.Log(only_shapes=False)
)
example = data_pipeline()
print(next(example))
data_pipeline = trax.data.Serial(
trax.data.TFDS('imdb_reviews', keys=('text', 'label'), train=True),
trax.data.Tokenize(vocab_dir='gs://trax-ml/vocabs/', vocab_file='en_8k.subword', keys=[0]),
trax.data.Log(only_shapes=True)
)
example = data_pipeline()
print(next(example))
```
# Shuffling our datasets
Trax offers two generator functions to add shuffle functionality in our input pipelines.
1. The `shuffle` function shuffles a given stream
2. The `Shuffle` function returns a shuffle function instead
## `shuffle`
> ```
def shuffle(samples, queue_size):
if queue_size < 1:
raise ValueError(f'Arg queue_size ({queue_size}) is less than 1.')
if queue_size == 1:
logging.warning('Queue size of 1 results in no shuffling.')
queue = []
try:
queue.append(next(samples))
i = np.random.randint(queue_size)
yield queue[i]
queue[i] = sample
except StopIteration:
logging.warning(
'Not enough samples (%d) to fill initial queue (size %d).',
len(queue), queue_size)
np.random.shuffle(queue)
for sample in queue:
yield sample
The `shuffle` function takes two inputs, the data stream and the queue size (minimum number of samples within which the shuffling takes place). Apart from the usual warnings, for negative and unity queue sizes, this generator function shuffles the given stream using [`np.random.randint()`](https://docs.python.org/3/library/random.html#random.randint) by randomly picks out integers using the `queue_size` as a range and then shuffle this new stream again using the [`np.random.shuffle()`](https://docs.python.org/3/library/random.html#random.shuffle)
```
sentence = ['Sed ut perspiciatis unde omnis iste natus error sit voluptatem accusantium doloremque laudantium, totam rem aperiam, eaque ipsa quae ab illo inventore veritatis et quasi architecto beatae vitae dicta sunt explicabo. Nemo enim ipsam voluptatem quia voluptas sit aspernatur aut odit aut fugit, sed quia consequuntur magni dolores eos qui ratione voluptatem sequi nesciunt. Neque porro quisquam est, qui dolorem ipsum quia dolor sit amet, consectetur, adipisci velit, sed quia non numquam eius modi tempora incidunt ut labore et dolore magnam aliquam quaerat voluptatem. Ut enim ad minima veniam, quis nostrum exercitationem ullam corporis suscipit laboriosam, nisi ut aliquid ex ea commodi consequatur? Quis autem vel eum iure reprehenderit qui in ea voluptate velit esse quam nihil molestiae consequatur, vel illum qui dolorem eum fugiat quo voluptas nulla pariatur?',
'But I must explain to you how all this mistaken idea of denouncing pleasure and praising pain was born and I will give you a complete account of the system, and expound the actual teachings of the great explorer of the truth, the master-builder of human happiness. No one rejects, dislikes, or avoids pleasure itself, because it is pleasure, but because those who do not know how to pursue pleasure rationally encounter consequences that are extremely painful. Nor again is there anyone who loves or pursues or desires to obtain pain of itself, because it is pain, but because occasionally circumstances occur in which toil and pain can procure him some great pleasure. To take a trivial example, which of us ever undertakes laborious physical exercise, except to obtain some advantage from it? But who has any right to find fault with a man who chooses to enjoy a pleasure that has no annoying consequences, or one who avoids a pain that produces no resultant pleasure?',
'Lorem ipsum dolor sit amet, consectetur adipiscing elit, sed do eiusmod tempor incididunt ut labore et dolore magna aliqua. Ut enim ad minim veniam, quis nostrud exercitation ullamco laboris nisi ut aliquip ex ea commodo consequat. Duis aute irure dolor in reprehenderit in voluptate velit esse cillum dolore eu fugiat nulla pariatur. Excepteur sint occaecat cupidatat non proident, sunt in culpa qui officia deserunt mollit anim id est laborum',
'At vero eos et accusamus et iusto odio dignissimos ducimus qui blanditiis praesentium voluptatum deleniti atque corrupti quos dolores et quas molestias excepturi sint occaecati cupiditate non provident, similique sunt in culpa qui officia deserunt mollitia animi, id est laborum et dolorum fuga. Et harum quidem rerum facilis est et expedita distinctio. Nam libero tempore, cum soluta nobis est eligendi optio cumque nihil impedit quo minus id quod maxime placeat facere possimus, omnis voluptas assumenda est, omnis dolor repellendus. Temporibus autem quibusdam et aut officiis debitis aut rerum necessitatibus saepe eveniet ut et voluptates repudiandae sint et molestiae non recusandae. Itaque earum rerum hic tenetur a sapiente delectus, ut aut reiciendis voluptatibus maiores alias consequatur aut perferendis doloribus asperiores repellat.']
def sample_generator(x):
for i in x:
yield i
example_shuffle = list(trax.data.inputs.shuffle(sample_generator(sentence), queue_size = 2))
example_shuffle
```
## `Shuffle`
> ```
def Shuffle(queue_size=1024):
return lambda g: shuffle(g, queue_size)
This function returns the aforementioned `shuffle` function and is mostly used in input pipelines.
# Batch Generators
## `batch`
This function, creates batches for the input generator function.
> ```
def batch(generator, batch_size):
if batch_size <= 0:
raise ValueError(f'Batch size must be positive, but is {batch_size}.')
buf = []
for example in generator:
buf.append(example)
if len(buf) == batch_size:
batched_example = tuple(np.stack(x) for x in zip(*buf))
yield batched_example
buf = []
It keeps adding objects from the generator into a list until the size becomes equal to the `batch_size` and then creates batches using the `np.stack()` function.
It also raises an error for non-positive batch_sizes.
## `Batch`
> ```
def Batch(batch_size):
return lambda g: batch(g, batch_size)
This Function returns the aforementioned `batch` function with given batch size.
# Pad to Maximum Dimensions
This function is used to pad a tuple of tensors to a joint dimension and return their batch.
For example, in this case a pair of tensors (1,2) and ( (3,4) , (5,6) ) is changed to (1,2,0) and ( (3,4) , (5,6) , 0)
```
import numpy as np
tensors = np.array([(1.,2.),
((3.,4.),(5.,6.))])
padded_tensors = trax.data.inputs.pad_to_max_dims(tensors=tensors, boundary=3)
padded_tensors
```
# Creating Buckets
For training Recurrent Neural Networks, with large vocabulary a method called Bucketing is usually applied.
The usual technique of using padding ensures that all occurences within a mini-batch are of the same length. But this reduces the inter-batch variability and intuitively puts similar sentences into the same batch therefore, reducing the overall robustness of the system.
Thus, we use Bucketing where multiple buckets are created depending on the length of the sentences and these occurences are assigned to buckets on the basis of which bucket corresponds to it's length. We need to ensure that the bucket sizes are large for adding some variablity to the system.
## `bucket_by_length`
> ```
def bucket_by_length(generator, length_fn, boundaries, batch_sizes,strict_pad_on_len=False):
buckets = [[] for _ in range(len(batch_sizes))]
boundaries = boundaries + [math.inf]
for example in generator:
length = length_fn(example)
bucket_idx = min([i for i, b in enumerate(boundaries) if length <= b])
buckets[bucket_idx].append(example)
if len(buckets[bucket_idx]) == batch_sizes[bucket_idx]:
batched = zip(*buckets[bucket_idx])
boundary = boundaries[bucket_idx]
boundary = None if boundary == math.inf else boundary
padded_batch = tuple(
pad_to_max_dims(x, boundary, strict_pad_on_len) for x in batched)
yield padded_batch
buckets[bucket_idx] = []
---
This function can be summarised as:
* Create buckets as per the lengths given in the `batch_sizes` array
* Assign sentences into buckets if their length matches the bucket size
* If padding is required, we use the `pad_to_max_dims` function
---
### Parameters
1. **generator:** The input generator function
2. **length_fn:** A custom length function for determing the length of functions, not necessarily `len()`
3. **boundaries:** A python list containing corresponding bucket boundaries
4. **batch_sizes:** A python list containing batch sizes
5. **strict_pad_on_len:** – A python boolean variable (`True` or `False`). If set to true then the function pads on the length dimension, where dim[0] is strictly a multiple of boundary.
## `BucketByLength`
> ```
def BucketByLength(boundaries, batch_sizes,length_keys=None, length_axis=0, strict_pad_on_len=False):
length_keys = length_keys or [0, 1]
length_fn = lambda x: _length_fn(x, length_axis, length_keys)
return lambda g: bucket_by_length(g, length_fn, boundaries, batch_sizes, strict_pad_on_len)
---
This function, is usually used inside input pipelines(*combinators*) and uses the afforementioned `bucket_by_length`. It applies a predefined `length_fn` which chooses the maximum shape on length_axis over length_keys.
It's use is illustrated below
```
data_pipeline = trax.data.Serial(
trax.data.TFDS('imdb_reviews', keys=('text', 'label'), train=True),
trax.data.Tokenize(vocab_dir='gs://trax-ml/vocabs/', vocab_file='en_8k.subword', keys=[0]),
trax.data.BucketByLength(boundaries=[32, 128, 512, 2048],
batch_sizes=[512, 128, 32, 8, 1],
length_keys=[0]),
trax.data.Log(only_shapes=True)
)
example = data_pipeline()
print(next(example))
```
# Filter by Length
> ```
def FilterByLength(max_length,length_keys=None, length_axis=0):
length_keys = length_keys or [0, 1]
length_fn = lambda x: _length_fn(x, length_axis, length_keys)
def filtered(gen):
for example in gen:
if length_fn(example) <= max_length:
yield example
return filtered
---
This function used the same predefined `length_fn` to only include those instances which are less than the given `max_length` parameter.
```
Filtered = trax.data.Serial(
trax.data.TFDS('imdb_reviews', keys=('text', 'label'), train=True),
trax.data.Tokenize(vocab_dir='gs://trax-ml/vocabs/', vocab_file='en_8k.subword', keys=[0]),
trax.data.BucketByLength(boundaries=[32, 128, 512, 2048],
batch_sizes=[512, 128, 32, 8, 1],
length_keys=[0]),
trax.data.FilterByLength(max_length=2048, length_keys=[0]),
trax.data.Log(only_shapes=True)
)
filtered_example = Filtered()
print(next(filtered_example))
```
# Adding Loss Weights
## `add_loss_weights`
> ```
def add_loss_weights(generator, id_to_mask=None):
for example in generator:
if len(example) > 3 or len(example) < 2:
assert id_to_mask is None, 'Cannot automatically mask this stream.'
yield example
else:
if len(example) == 2:
weights = np.ones_like(example[1]).astype(np.float32)
else:
weights = example[2].astype(np.float32)
mask = 1.0 - np.equal(example[1], id_to_mask).astype(np.float32)
weights *= mask
yield (example[0], example[1], weights)
---
This function essentially adds a loss mask (tensor of ones of the same shape) to the input stream.
**Masking** is essentially a way to tell sequence-processing layers that certain timesteps in an input are missing, and thus should be skipped when processing the data.
Thus, it adds 'weights' to the system.
---
### Parameters
1. **generator:** The input data generator
2. **id_to_mask:** The value with which to mask. Can be used as `<PAD>` in NLP.
```
train_generator = trax.data.inputs.add_loss_weights(
data_generator(batch_size, x_train, y_train,vocab['<PAD>'], True),
id_to_mask=vocab['<PAD>'])
```
For example, in this case I used the `add_loss_weights` function to add padding while implementing Named Entity Recogntion using the Reformer Architecture. You can read more about the project [here](https://www.kaggle.com/sauravmaheshkar/trax-ner-using-reformer).
## `AddLossWeights`
This function performs the afforementioned `add_loss_weights` to the data stream.
> ```
def AddLossWeights(id_to_mask=None):
return lambda g: add_loss_weights(g,id_to_mask=id_to_mask)
```
data_pipeline = trax.data.Serial(
trax.data.TFDS('imdb_reviews', keys=('text', 'label'), train=True),
trax.data.Tokenize(vocab_dir='gs://trax-ml/vocabs/', vocab_file='en_8k.subword', keys=[0]),
trax.data.Shuffle(),
trax.data.FilterByLength(max_length=2048, length_keys=[0]),
trax.data.BucketByLength(boundaries=[ 32, 128, 512, 2048],
batch_sizes=[512, 128, 32, 8, 1],
length_keys=[0]),
trax.data.AddLossWeights(),
trax.data.Log(only_shapes=True)
)
example = data_pipeline()
print(next(example))
```
| github_jupyter |
# Simple Waveform
following [this tutorial](https://www.pythonforengineers.com/audio-and-digital-signal-processingdsp-in-python/)
```
%matplotlib inline
import wave
import struct
import matplotlib.pyplot as plt
import numpy as np
import astropy.units as u
import kplr
import celerite
from celerite import terms
from scipy.ndimage import gaussian_filter1d
from astropy.io import fits
frequency = 440 * u.Hz
sampling_rate = 48000 * u.Hz
duration = 5 * u.s
num_samples = int(duration * sampling_rate)
# The sampling rate of the analog to digital convert
amplitude = 20000
file = "test.wav"
x = np.arange(0, num_samples)
sine_wave = np.sin(2 * np.pi * float(frequency/sampling_rate) * x)
@u.quantity_input(sampling_rate=u.Hz)
def write_wave(waveform, num_samples, sampling_rate, path='test.wav'):
nframes=num_samples
comptype = "NONE"
compname = "not compressed"
nchannels = 1
sampwidth = 2
fig, ax = plt.subplots()
ax.plot(np.arange(len(waveform))/sampling_rate.value, waveform)
with wave.open(path, 'w') as wav_file:
wav_file.setparams((nchannels, sampwidth, int(sampling_rate.value), nframes, comptype, compname))
frames = struct.pack(len(waveform)*'h', *(waveform*amplitude).astype(int).tolist())
wav_file.writeframes(frames)
return fig, ax
def light_curve(koi):
client = kplr.API()
# Find the target KOI.
koi = client.koi(koi + 0.01) #Kepler-17
# Get a list of light curve datasets.
lcs = koi.get_light_curves(short_cadence=False)
# Loop over the datasets and read in the data.
time, flux, ferr, quality = [], [], [], []
for lc in lcs:
with lc.open() as f:
# The lightcurve data are in the first FITS HDU.
hdu_data = f[1].data
t = hdu_data["time"]
f = hdu_data["sap_flux"]
not_nan = ~np.isnan(f) & ~np.isnan(t)
fit = np.polyval(np.polyfit(t[not_nan], f[not_nan], 2), t[not_nan])
time.append(t[not_nan])
flux.append(f[not_nan]/fit)
ferr.append(hdu_data["sap_flux_err"])
quality.append(hdu_data["sap_quality"])
return time, flux
def gp_interpolate(time, flux, N=10000):
kernel = terms.SHOTerm(log_S0=-5, log_omega0=np.log(2*np.pi*20), log_Q=1/np.sqrt(2))
gp = celerite.GP(kernel)
gp.compute(time)
from scipy.optimize import minimize
def neg_log_like(params, y, gp):
gp.set_parameter_vector(params)
return -gp.log_likelihood(y)
initial_params = gp.get_parameter_vector()
bounds = gp.get_parameter_bounds()
r = minimize(neg_log_like, initial_params, method="L-BFGS-B", args=(flux, gp)) # bounds=bounds,
gp.set_parameter_vector(r.x)
x = np.linspace(time.min(), time.max(), N)
predicted_flux = gp.predict(flux, x, return_cov=False)
return x, predicted_flux
write_wave(sine_wave, num_samples, sampling_rate)
```
# Trappist-1
```
d = fits.getdata('data/nPLDTrappist.fits')
time, flux = d['CADN'], d['FLUX']
flux /= flux.mean()
condition = (flux > 0.9) & (flux < 1.1) & (~np.isnan(flux)) & (~np.isnan(time))
fit = np.polyval(np.polyfit(time[condition] - time.mean(), flux[condition], 10), time[condition] - time.mean())
time = time[condition]
flux = flux[condition] - fit
flux = gaussian_filter1d(flux, 5)
plt.plot(time, flux)
x, predicted_flux = gp_interpolate(time[150:], flux[150:], N=8e4)
plt.plot(x, predicted_flux)
#flux = np.tile(flux / flux.max(), 100)
predicted_flux = np.tile(np.concatenate([predicted_flux, predicted_flux[::-1]]), 50)
#layers = 100*predicted_flux + np.tile(predicted_flux[::10], 10)# + np.tile(predicted_flux[::2], 2)
write_wave(100*predicted_flux[::2], len(predicted_flux[::2]), sampling_rate*2, path='trappist1.wav')
```
# Kepler-62
```
time, flux = light_curve(701)
time = np.concatenate(time)
flux = np.concatenate(flux)
flux -= flux.mean()
flux = gaussian_filter1d(flux, 10)
plt.plot(time, flux)
flux = np.tile(np.concatenate([flux, flux[::-1]]), 30)
write_wave(100*flux[::2], len(flux[::2]), sampling_rate*2, path='k62.wav')
```
# Kepler-296
```
time, flux = light_curve(1422)
time = np.concatenate(time)
flux = np.concatenate(flux)
flux -= flux.mean()
from scipy.ndimage import gaussian_filter1d
flux = gaussian_filter1d(flux, 10)
plt.plot(time, flux)
plt.plot(time, flux)
x, predicted_flux = gp_interpolate(time, flux)
plt.plot(x, predicted_flux)
flux = np.tile(np.concatenate([predicted_flux, predicted_flux[::-1]]), 50)
fig, ax = write_wave(40*flux, len(flux), sampling_rate, path='k296.wav')
# predicted_flux = np.tile(np.concatenate([predicted_flux, predicted_flux[::-1]]), 5)
# fig, ax = write_wave(100*predicted_flux, len(predicted_flux), sampling_rate*2, path='k296.wav')
```
# Kepler-442
```
time, flux = light_curve(4742)
time = np.concatenate(time)
flux = np.concatenate(flux)
flux -= flux.mean()
flux = gaussian_filter1d(flux, 10)
plt.plot(time, flux)
plt.plot(time, flux)
#x, predicted_flux = gp_interpolate(time, flux)
#plt.plot(x, predicted_flux)
flux = np.tile(np.concatenate([flux, flux[::-1]]), 30)
fig, ax = write_wave(100*flux[::2], len(flux[::2]), sampling_rate*2, path='k442.wav')
```
# Kepler-1229
```
time, flux = light_curve(2418)
time = np.concatenate(time)
flux = np.concatenate(flux)
flux -= flux.mean()
flux = gaussian_filter1d(flux, 40)
plt.plot(time, flux)
plt.plot(time, flux)
x, predicted_flux = gp_interpolate(time, flux)
plt.plot(x, predicted_flux)
flux = np.tile(np.concatenate([predicted_flux, predicted_flux[::-1]]), 30)
fig, ax = write_wave(100*flux, len(flux), sampling_rate, path='k1229.wav')
```
# Kepler-186
```
time, flux = light_curve(571)
time = np.concatenate(time)
flux = np.concatenate(flux)
flux -= flux.mean()
flux = gaussian_filter1d(flux, 10)
plt.plot(time, flux)
flux = np.tile(np.concatenate([flux, flux[::-1]]), 30)
fig, ax = write_wave(100*flux[::2], len(flux[::2]), sampling_rate*2, path='k186.wav')
```
| github_jupyter |
```
# Copyright 2020 Google LLC
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
import collections
import functools
import itertools
import os
import pathlib
import re
import textwrap
import matplotlib.pyplot as plt
import numpy as np
import datasets
import shutil
import tensorflow as tf
import transformers
import tqdm
# Prepare different HuggingFace objects that will definitely be needed.
kilt = datasets.load_dataset("kilt_tasks")
eli5 = {k.split("_")[0]: v for k, v in kilt.items() if "eli5" in k}
tokenizer = transformers.GPT2TokenizerFast.from_pretrained("gpt2-xl")
print(f"Dataset split keys: {list(eli5.keys())}")
# Extract the lengths of combined question and answer text, once tokenized with the GPT2 tokenizer.
def get_len(sample):
question = sample["input"].strip()
answer = min(sample["output"]["answer"], key=len).strip()
len_ = len(tokenizer(question + " " + answer)["input_ids"])
return {"len_":len_}
mapped = eli5["train"].map(get_len, num_proc=os.cpu_count())
# Count the number of entries of each length, then sort by length
counts = collections.Counter(mapped["len_"])
sorted_counts = sorted(counts.items(), key=lambda x: x[0])
# Compute the ratio of samples that are in a certain percentile of lengths, and compute how long retrievals
# would have to be for a certain fraction of the dataset to have access to that amount of retrievals
context_length = 1024
max_num_retrievals = 4
fractions = [.55, .6, .65, .7, .75, .775, .8, .825, .85, .875, .9, .925, .95, .975][::-1]
points = {}
max_pairs = {}
for top_fraction in fractions:
qty_accumulator = 0
cumulative = [
(count, qty_accumulator := qty / len(eli5["train"]) + qty_accumulator)
for count, qty in sorted_counts if qty_accumulator < top_fraction
]
x = [x[0] for x in cumulative]
y = [x[1] for x in cumulative]
points[top_fraction] = (x, y)
max_pair = max(cumulative, key=lambda pair: pair[1])
max_pairs[top_fraction] = max_pair[0]
print(f"{top_fraction:0.1%}: {max_pair[0]} bpe tokens or fewer")
for i in range(1, max_num_retrievals + 1):
print(f"({context_length} - {max_pair[0]}) / {i} = {(context_length - max_pair[0]) / i:0.0f}")
# Graph the results.
plt.figure(figsize=(10, 10));
ax = plt.gca()
ax.margins(tight=True)
ax.plot(*points[max(fractions)]);
plt.yticks(np.linspace(0, 1, 21))
percentages = max_pairs.keys()
lengths = max_pairs.values()
for x, y in zip(lengths, percentages):
ax.plot((x, x), (0, y), color="red")
ax.scatter(lengths,
percentages, color="red");
ax.scatter(lengths,
[0 for _ in lengths], color="red");
for y, x in max_pairs.items():
ax.text(x - 25, y + 0.025, f"{y:0.1%}", size=13)
ax.text(x - 10, -0.032, f"{x}", size=10)
```
| github_jupyter |
#### Dependencies
```
import numpy as np
import pandas as pd
import xarray as xr
# geo libs are optional
from shapely.geometry import Point
import geopandas as gpd
from geopandas import GeoDataFrame
# cd ../..
pwd # should be top-level, i.e. ~/*/generative-downscaling/
temp_1 = xr.open_dataset("./data/raw/temp/1406/2m_temperature_1991_1.40625deg.nc")
temp_5 = xr.open_dataset("./data/raw/temp/5625/2m_temperature_1991_5.625deg.nc")
```
#### Basic XArray
```
temp_1
temp_1.dims # 8760/365 = 24!
temp_5.dims
temp_1.coords
type(temp_1.t2m.data)
```
#### Basic Filtering
```
daily = temp_1.isel(time=(temp_1.time.dt.hour == 0))
daily.dims
daily_5 = temp_5.isel(time=(temp_5.time.dt.hour == 0))
daily.dims
```
## Visual Exploration
### Location & Consistency
#### Where is the Data Located?
```
def obs_by_geo(weatherbench: pd.DataFrame) -> GeoDataFrame:
""" Return total observations by coordinate """
obs_by_geo = weatherbench.groupby(["lat","lon"]).size()
obs_by_geo.name = "count"
obs_by_geo = obs_by_geo.reset_index()
gdf = GeoDataFrame(
obs_by_geo,
geometry=gpd.points_from_xy(obs_by_geo.lon - 180, obs_by_geo.lat)
)
return gdf
fine_cts = obs_by_geo(daily.to_dataframe())
coarse_cts = obs_by_geo(daily_5.to_dataframe())
world = gpd.read_file(gpd.datasets.get_path('naturalearth_lowres'))
ax = fine_cts.plot(
ax=world.plot(figsize=(15, 8), alpha=.33),
marker='o', color='red', markersize=.05,
label = "Fine Resolution"
);
coarse_cts.plot(
ax=ax,
marker='o', color='green', markersize=1,
label = "Coarse Resolution"
);
ax.set_title("WeatherBench Dataset Geographic Coverage (Full)", fontsize=20);
ax.set_xlabel("Longitude", fontsize=14);
ax.set_ylabel("Latitude", fontsize=14);
ax.legend(fontsize=12);
```
#### Americas Only
```
EPSILON = .0001
NA_LAT_FINE = (-5, 85+EPSILON) # epsilon to avoid non-inclusive right
NA_LON_FINE = (-137.5+180, -47.5+180+EPSILON)
NA_LAT_COARSE = (-10, 88+EPSILON) # Coarse should be wider for full coverage
NA_LON_COARSE = (-145+180, -45+180+EPSILON)
daily_na = daily.sel(lat=slice(*NA_LAT_FINE), lon=slice(*NA_LON_FINE))
daily_na_5 = daily_5.sel(lat=slice(*NA_LAT_COARSE), lon=slice(*NA_LON_COARSE))
fine_cts_americas = fine_cts[fine_cts['lat'].between(*NA_LAT_FINE) & fine_cts['lon'].between(*NA_LON_FINE)]
coarse_cts_americas = coarse_cts[coarse_cts['lat'].between(*NA_LAT_COARSE) & coarse_cts['lon'].between(*NA_LON_COARSE)]
americas = world[world['continent'].isin(["North America", "South America"])]
ax = fine_cts_americas.plot(
ax=americas.plot(figsize=(8,8), alpha=.33),
marker='o', color='red', markersize=.05,
label = "Fine Resolution"
);
coarse_cts_americas.plot(
ax=ax,
marker='o', color='green', markersize=1,
label = "Coarse Resolution"
);
ax.set_xlim(-160,-20);
ax.set_ylim(-30,90);
ax.set_title("WeatherBench Dataset Coverage (Americas Only)", fontsize=20);
ax.set_xlabel("Longitude", fontsize=14);
ax.set_ylabel("Latitude", fontsize=14);
ax.legend(fontsize=12);
```
### Temperature
```
t5_pdf = temp_5.to_dataframe()
# t5_pdf = t5_pdf[:100]
# coords = t5_pdf.reset_index()[['lat','lon']].drop_duplicates()
# coords[coords['lat'].between(40,50) & coords['lon'].between(70+180,80+180)]
coords_mtl_approx = (47.8125, 253.125) # lat, lon
mtl_1991 = t5_pdf.loc[coords_mtl_approx[0]].loc[coords_mtl_approx[1]]
mtl_1991_hi_lo = mtl_1991\
.groupby([mtl_1991.index.date])\
.agg([np.min, np.max])\
.rename(columns={'amin': 'low', 'amax': 'high'})
mtl_1991_hi_lo_c = mtl_1991_hi_lo - 273.15 # kelvin to celsius
mtl_1991_hi_lo_c.plot()
```
| github_jupyter |
# Video Codec Unit (VCU) Demo Example: STREAM_IN->DECODE ->DISPLAY
# Introduction
Video Codec Unit (VCU) in ZynqMP SOC is capable of encoding and decoding AVC/HEVC compressed video streams in real time.
This notebook example acts as Client pipeline in streaming use case. It needs to be run along with Server notebook (__vcu-demo-transcode-to-streamout. ipynb or vcu-demo-camera-encode-streamout. ipynb__). It receives encoded data over network, decode using VCU and render it on DP/HDMI Monitor.
# Implementation Details
<img src="pictures/block-diagram-streamin-decode.png" align="center" alt="Drawing" style="width: 600px; height: 200px"/>
This example requires two boards, board-1 is used for transcode and stream-out (as a server) and **board 2** is used for streaming-in and decode purpose (as a client) or VLC player on the host machine can be used as client instead of board-2 (More details regarding Test Setup for board-1 can be found in transcode → stream-out Example).
__Note:__ This notebook needs to be run along with "vcu-demo-transcode-to-streamout.ipynb" or "vcu-demo-camera-encode-streamout.ipynb". The configuration settings below are for Client-side pipeline.
### Board Setup
**Board 2 is used for streaming-in and decode purpose (as a client)**
1. Connect 4k DP/HDMI display to board.
2. Connect serial cable to monitor logs on serial console.
3. If Board is connected to private network, then export proxy settings in /home/root/.bashrc file on board as below,
- create/open a bashrc file using "vi ~/.bashrc"
- Insert below line to bashrc file
- export http_proxy="< private network proxy address >"
- export https_proxy="< private network proxy address >"
- Save and close bashrc file.
4. Connect two boards in the same network so that they can access each other using IP address.
5. Check server IP on server board.
- root@zcu106-zynqmp:~#ifconfig
6. Check client IP.
7. Check connectivity for board-1 & board-2.
- root@zcu106-zynqmp:~#ping <board-2's IP>
8. Run stream-in → Decode on board-2
Create test.sdp file on host with below content (Add separate line in test.sdp for each item below) and play test.sdp on host machine.
1. v=0 c=IN IP4 <Client machine IP address>
2. m=video 50000 RTP/AVP 96
3. a=rtpmap:96 H264/90000
4. a=framerate=30
Trouble-shoot for VLC player setup:
1. IP4 is client-IP address
2. H264/H265 is used based on received codec type on the client
3. Turn-off firewall in host machine if packets are not received to VLC.
```
from IPython.display import HTML
HTML('''<script>
code_show=true;
function code_toggle() {
if (code_show){
$('div.input').hide();
} else {
$('div.input').show();
}
code_show = !code_show
}
$( document ).ready(code_toggle);
</script>
<form action="javascript:code_toggle()"><input type="submit" value="Click here to toggle on/off the raw code."></form>''')
```
# Run the Demo
```
from ipywidgets import interact
import ipywidgets as widgets
from common import common_vcu_demo_streamin_decode_display
import os
from ipywidgets import HBox, VBox, Text, Layout
```
### Video
```
codec_type=widgets.RadioButtons(
options=['avc', 'hevc'],
description='Codec Type:',
disabled=False)
video_sink={'kmssink':['DP', 'HDMI'], 'fakevideosink':['none']}
def print_video_sink(VideoSink):
pass
def select_video_sink(VideoCodec):
display_type.options = video_sink[VideoCodec]
sink_name = widgets.RadioButtons(options=sorted(video_sink.keys(), key=lambda k: len(video_sink[k]), reverse=True), description='Video Sink:')
init = sink_name.value
display_type = widgets.RadioButtons(options=video_sink[init], description='Display:')
j = widgets.interactive(print_video_sink, VideoSink=display_type)
i = widgets.interactive(select_video_sink, VideoCodec=sink_name)
HBox([codec_type, i, j])
```
### Audio
```
audio_sink={'none':['none'], 'aac':['auto','alsasink','pulsesink'],'vorbis':['auto','alsasink','pulsesink']}
audio_src={'none':['none'], 'aac':['auto','alsasrc','pulsesrc'],'vorbis':['auto','alsasrc','pulsesrc']}
#val=sorted(audio_sink, key = lambda k: (-len(audio_sink[k]), k))
def print_audio_sink(AudioSink):
pass
def print_audio_src(AudioSrc):
pass
def select_audio_sink(AudioCodec):
audio_sinkW.options = audio_sink[AudioCodec]
audio_srcW.options = audio_src[AudioCodec]
audio_codecW = widgets.RadioButtons(options=sorted(audio_sink.keys(), key=lambda k: len(audio_sink[k])), description='Audio Codec:')
init = audio_codecW.value
audio_sinkW = widgets.RadioButtons(options=audio_sink[init], description='Audio Sink:')
audio_srcW = widgets.RadioButtons(options=audio_src[init], description='Audio Src:')
j = widgets.interactive(print_audio_sink, AudioSink=audio_sinkW)
i = widgets.interactive(select_audio_sink, AudioCodec=audio_codecW)
HBox([i, j])
```
### Advanced options:
```
kernel_recv_buffer_size=widgets.Text(value='',
placeholder='(optional) 16000000',
description='Kernel Recv Buf Size:',
style={'description_width': 'initial'},
#layout=Layout(width='33%', height='30px'),
disabled=False)
port_number=widgets.Text(value='',
placeholder='(optional) 50000, 42000',
description=r'Port No:',
#style={'description_width': 'initial'},
#
disabled=False)
#kernel_recv_buffer_size
HBox([kernel_recv_buffer_size, port_number])
entropy_buffers=widgets.Dropdown(
options=['2', '3', '4', '5', '6', '7', '8', '9', '10', '11', '12', '13', '14', '15'],
value='5',
description='Entropy Buffers Nos:',
style={'description_width': 'initial'},
disabled=False,)
show_fps=widgets.Checkbox(
value=False,
description='show-fps',
#style={'description_width': 'initial'},
disabled=False)
HBox([entropy_buffers, show_fps])
from IPython.display import clear_output
from IPython.display import Javascript
def run_all(ev):
display(Javascript('IPython.notebook.execute_cells_below()'))
def clear_op(event):
clear_output(wait=True)
return
button1 = widgets.Button(
description='Clear Output',
style= {'button_color':'lightgreen'},
#style= {'button_color':'lightgreen', 'description_width': 'initial'},
layout={'width': '300px'}
)
button2 = widgets.Button(
description='',
style= {'button_color':'white'},
#style= {'button_color':'lightgreen', 'description_width': 'initial'},
layout={'width': '38px'},
disabled=True
)
button1.on_click(run_all)
button1.on_click(clear_op)
def start_demo(event):
#clear_output(wait=True)
arg = common_vcu_demo_streamin_decode_display.cmd_line_args_generator(port_number.value, codec_type.value, audio_codecW.value, display_type.value, kernel_recv_buffer_size.value, sink_name.value, entropy_buffers.value, show_fps.value, audio_sinkW.value);
#sh vcu-demo-streamin-decode-display.sh $arg > logs.txt 2>&1
!sh vcu-demo-streamin-decode-display.sh $arg
return
button = widgets.Button(
description='click to start vcu-stream_in-decode-display demo',
style= {'button_color':'lightgreen'},
#style= {'button_color':'lightgreen', 'description_width': 'initial'},
layout={'width': '350px'}
)
button.on_click(start_demo)
HBox([button, button2, button1])
```
# References
[1] https://xilinx-wiki.atlassian.net/wiki/spaces/A/pages/18842546/Xilinx+Video+Codec+Unit
[2] https://www.xilinx.com/support.html#documentation (Refer to PG252)
| github_jupyter |
<table>
<tr align=left><td><img align=left src="https://i.creativecommons.org/l/by/4.0/88x31.png">
<td>Text provided under a Creative Commons Attribution license, CC-BY. All code is made available under the FSF-approved MIT license. (c) Kyle T. Mandli</td>
</table>
```
%matplotlib inline
from __future__ import print_function
import numpy
import matplotlib.pyplot as plt
```
# Convergence Results for Initial Value Problems
Convergence for IVPs is a bit different than with BVPs, we want in general
$$
\lim_{\Delta t \rightarrow 0} U^N = u(t_f)
$$
where $t_f$ is the final desired time and $N$ is the number of time steps needed to reach $t_f$ such that
$$
N \Delta t = t_f \quad \Rightarrow N = \frac{t_f}{\Delta t}.
$$
We need to be careful at this juncture however when we are talking about a convergent method. A method can be convergent for a particular set of equations and particular initial conditions but not others. Practically speaking we would like convergence results to apply to a reasonably large set of equations and initial conditions. With these considerations we have the following definition of convergence for IVPs.
If applying an $r$-step method to an ODE of the form
$$
u'(t) = f(t,u)
$$
with $f(t,u)$ Lipschitz continuous in $u$, and with any set of starting values satisfying
$$
\lim_{\Delta t\rightarrow 0} U^\nu(\Delta t) = u_0 \quad \text{for} \quad \nu = 0, 1, \ldots, r-1
$$
(i.e. the bootstrap startup for the multi-step method is consistent with the initial value as $\Delta t \rightarrow$), then the method is said to be *convergent* in the sense
$$
\lim_{\Delta t \rightarrow 0} U^N = u(t_f).
$$
As we saw previously for a method to be convergent it must be
- **consistent** - the local truncation error $\tau = \mathcal{O}(\Delta t^p)$ where $p > 0$ and
- **zero-stable** - a similar minimal form of stability implying that the sum total of the errors as $\Delta t \rightarrow 0$ is bounded and has the same order as $\tau$ which we know goes to zero as $\Delta t \rightarrow 0$.
## One-Step Method Convergence
Consider the simple linear problem
$$
\frac{\text{d}u}{\text{d}t} = \lambda u + g(t) \quad \text{with}\quad u(0) = u_0
$$
which we know has the solution
$$
u(t) = u_0 e^{\lambda (t - t_0)} + \int^t_{t_0} e^{\lambda (t - \tau)} g(\tau) d\tau.
$$
### Forward Euler on a Linear Problem
Applying Euler's method to this problem leads to
$$\begin{aligned}
U^{n+1} &= U^n + \Delta t\lambda U^n \\
&= (1 + \Delta t \lambda) U^n
\end{aligned}$$
We also know the local truncation error is
$$\begin{aligned}
\tau^n &= \left (\frac{u(t_{n+1}) - u(t_n)}{\Delta t} \right ) - \lambda u(t_n)\\
&= \left (u'(t_n) + \frac{1}{2} \Delta t u''(t_n) + \mathcal{O}(\Delta t^2) \right ) - u'(t_n) \\
&= \frac{1}{2} \Delta t u''(t_n) + \mathcal{O}(\Delta t^2)
\end{aligned}$$
Noting the original definition of $\tau^n$ we can rewrite the expression for the local truncation error as
$$
u(t_{n+1}) = (1 + \Delta t \lambda) u(t_n) + \Delta t \tau^n
$$
which in combination with the application of Euler's method leads to an expression for the global error
$$\begin{aligned}
E^{n+1} = U^{n+1} - u(t^{n+1}) &= (1 + \Delta t \lambda) U^n - (1 + \Delta t \lambda) u(t_n) - \Delta t \tau^n \\
&= (1+\Delta t \lambda) E^n - \Delta t \tau^n \\
\end{aligned}$$
Expanding this expression out backwards in time to $n=0$ leads to
$$
E^n = (1 + \Delta t \lambda) E^0 - \Delta t \sum^n_{i=1} (1 + \Delta t \lambda)^{n-i} \tau^{i - 1}.
$$
We can now see the importance of the term $(1 + \Delta t \lambda)$. We can bound this term by
$$
|1 + \Delta t \lambda| \leq e^{\Delta t \lambda}
$$
which then implies the term in the summation can be bounded by
$$
|1 + \Delta t \lambda|^{n - i} \leq e^{(n-i) \Delta t |\lambda|} \leq e^{n \Delta t |\lambda||} \leq e^{|\lambda| t_f}
$$
Using this expression in the expression for the global error we find
$$\begin{aligned}
E^n &= (1 + \Delta t \lambda) E^0 - \Delta t \sum^n_{i=1} (1 + \Delta t \lambda)^{n-i} \tau^{i - 1} \\
|E^n| &\leq e^{|\lambda| \Delta t} |E^0| - \Delta t \sum^n_{i=1} e^{|\lambda| t_f} |\tau^{i - 1}| \\
&\leq e^{|\lambda| t_f} \left(|E^0| - \Delta t \sum^n_{i=1} |\tau^{i - 1}|\right) \\
&\leq e^{|\lambda| t_f} \left(|E^0| - n \Delta t \max_{1 \leq i \leq n} |\tau^{i - 1}|\right)
\end{aligned}$$
In other words the global error is bounded by the original global error and the maximum one-step error made multiplied by the number of time steps taken. If $N = \frac{t_f}{\Delta t}$ as before and taking into account the local truncation error we can simplify this expression further to
$$
|E^n| \leq e^{|\lambda| t_f} \left[|E^0| + t_f \left(\frac{1}{2} \Delta t |u''| + \mathcal{O}(\Delta t^2)\right ) \right]
$$
If we assume that we have used the correct initial condition $u_0$ then $E_0 \rightarrow 0$ as $\Delta t \rightarrow 0$ and we see that the method is truly convergent as
$$
|E^n| \leq e^{|\lambda| t_f} t_f \left(\frac{1}{2} \Delta t |u''| + \mathcal{O}(\Delta t^2)\right ) = \mathcal{O}(\Delta t).
$$
### Relation to Stability for BVPs
We can see the relationship between the previous version of stability and the one outlined above. Try writing the forward Euler method as a linear system.
Forward Euler:
$$
A = \frac{1}{\Delta t} \begin{bmatrix}
1 \\
-(1 + \Delta t \lambda) & 1 \\
& -(1 + \Delta t \lambda) & 1 \\
& & -(1 + \Delta t \lambda) & 1 \\
& & & \ddots & \ddots \\
& & & & -(1 + \Delta t \lambda) & 1 \\
& & & & & -(1 + \Delta t \lambda) & 1
\end{bmatrix}
$$
with
$$
U = \begin{bmatrix} U^1 \\ U^2 \\ \vdots \\ U^N \end{bmatrix} ~~~~
F = \begin{bmatrix} (1 / \Delta t + \lambda) U^0 + g(t_0) \\ g(t_1) \\ \vdots \\ g(t_{N-1}) \end{bmatrix}
$$
Following our previous stability result and taking $\hat{U~}$ to be the vector obtained from the true solution ($\hat{U~}^i = u(t_i)$) we then have
$$
A U = F ~~~~~~ A \hat{U~} = F + \tau
$$
and therefore
$$
A (\hat{U~} - U) = \tau.
$$
Noting that $\hat{U~} - U = E$ we can then invert that matrix $A$ to find the relationship between the truncation error $\tau$ and the global error $E$. As before then we require that $A^{-1}$ is invertible (which is trivial in this case) and that $||A^{-1}|| < C$ in some norm. We can see this as
$$
A^{-1} = \Delta t \begin{bmatrix}
1 \\
(1 + \Delta t \lambda) & 1 \\
(1 + \Delta t \lambda)^2 & (1 + \Delta t \lambda) & 1 \\
(1 + \Delta t \lambda)^3 & (1 + \Delta t \lambda)^2 & (1 + \Delta t \lambda) & 1\\
\vdots & & & \ddots \\
(1 + \Delta t \lambda)^{N-1} & (1 + \Delta t \lambda)^{N-2} (1 + \Delta t \lambda)^{N-3} & \cdots & (1 + \Delta t \lambda) & 1
\end{bmatrix}
$$
whose infinity norm is
$$
||A^{-1}||_\infty = \Delta t \sum^N_{m=1} | (1 + \Delta t \lambda)^{N-M} |
$$
and therefore
$$
||A^{-1}||_\infty \leq \Delta t N e^{|\lambda| T} = T e^{|\lambda| T}.
$$
As $\Delta t \rightarrow 0$ this is bounded for **fixed T**.
### General One-Step Method Convergence
Consider the general one step method denoted by
$$
U^{n+1} = U^n + \Delta t \Phi(U^n, t_n, \Delta t).
$$
Assuming $\Phi$ is continuous in $t$ and $\Delta t$ and Lipschitz continous in $u$ with Lipschitz contsant $L$ (related to the Lipschitz constant of $f$). If the one-step method is consistent
$$
\Phi(u,t,0) = f(u,t)
$$
for all $u$, $t$, and $\Delta t$ and the local truncation error is
$$
\tau^n =\frac{u(t_{n+1}) - u(t_n)}{\Delta t} - \phi(u(t_n), t_n, \Delta t)
$$
then the one-step method is convergent.
Using the general approach we used for forward Euler we know that the true solution and $\tau$ are realted through
$$
u(t_{n+1}) = u(t_n) + \Delta t \Phi(u(t_n), t_n, \Delta t) + \Delta t \tau^n
$$
which subtracted from the approximate solution
$$
U^{n+1} = U^n + \Delta t \Phi(U^n, t_n, \Delta t)
$$
leads to
$$
E^{n+1} = E^n + \Delta t (\Phi(U^n, t_n, \Delta t) - \Phi(u(t_n), t_n, \Delta t)) - \Delta t \tau^n.
$$
Using the Lipschitz continuity of $\Phi$ we then have
$$
|E^{n+1}| \leq |E^n| + \Delta t L |E^n| + \Delta t |\tau^n|.
$$
which has the same form as we saw in the proof for forward Euler.
## Zero-Stability for Linear Multistep Methods
We can also make general statements for linear multistep methods although it is important to note that we have additional requirements for linear multistep methods so that they are convergent. As an example consider the method
$$
U^{n+2} - 3 U^{n+1} + 2 U^n = - \Delta t f(U^n)
$$
so that $\alpha_0 = 2$, $\alpha_1 = -3$, and $\alpha_2 = 1$ and $\beta_0 = -1$ with the rest equal to zero. Note that these coefficients satisfy our conditions for being consistent with a truncation error
$$
\tau^n = \frac{1}{\Delta t} (u(t_{n+2}) - 3 u(t_{n+1}) + 2 u(t_n) + \Delta t u'(t_n)) = \frac{5}{2} \Delta t u''(t_n) + \mathcal{O}(\Delta t^2).
$$
It turns out that although this method is consistent the global error does not converge in general!
$$
U^{n+2} - 3 U^{n+1} + 2 U^n = - \Delta t f(U^n)
$$
Consider the above method with the trivial ODE
$$
u'(t) = 0 \quad u(0) = 0
$$
so that we are left with the method
$$
U^{n+2} - 3 U^{n+1} + 2 U^n = 0.
$$
If we have exact values for $U^0$ and $U^1$ then this method would lead to $U^n = 0$. In general however we only have an approximation to $U^1$ so what happens then? We can solve the linear difference equation in terms of $U^0$ and $U^1$ to find
$$
U^n = 2 U^0 - U^1 + 2^n (U^1 - U^0).
$$
If we assume a error on the order of $\mathcal{O}(\Delta t)$ for $U^1$ this quickly leads to large values even for small $n$!
### Characteristic Polynomials and Linear Difference Equations
As an short aside, say we wanted to solve
$$\sum^r_{j=0} \alpha_j U^{n+j} = 0$$
given initial conditions $U^0, U^1, \ldots, U^{r-1}$ which has a solution in the general form $U^n = \xi^n$.
Plugging this into the equation we have
$$
\sum^r_{j=0} \alpha_j \xi^{n+j} = 0
$$
which simplifies to
$$
\sum^r_{j=0} \alpha_j \xi^j = 0
$$
by dividing by $\xi^n$.
If $\xi$ then is a root of the polynomial
$$
\rho(\xi) = \sum^r_{j=0} \alpha_j \xi^j
$$
then $\xi$ solves the equation.
Note that since these are linear methods that a linear combination of solutions is also a solution so the general form of a solution has the form
$$
U^n = c_1 \xi_1^n + c_2 \xi_2^n + \cdots + c_r \xi^n_r.
$$
Given initial values for $U^0, U^1, \ldots$ we can uniquely determine the $c_j$.
### General Zero-Stability Result for LMM
An $r$-step LMM is *zero-stable* if the roots of the characteristic polynomial $\rho(\xi)$ satisfy
$$
|\xi_j| \leq 1 \quad \quad \text{for} \quad j=1,2,3,\ldots,r
$$
if $\xi_j$ is not repeated and $|\xi_j| < 1$ for repeated roots.
#### Example
Consider the linear multistep method
$$
U^{n+2} - 2 U^{n+1} + U^n = \frac{\Delta t}{2} (f(U^{n+2}) - f(U^n)).
$$
Applying this to the ODE $u'(t) = 0$ leads to the difference equation
$$
U^{n+2} - 2 U^{n+1} + U^n = 0
$$
whose characteristic polynomial is
$$
\rho(\xi) = \xi^2 - 2 \xi + 1 = (\xi - 1)^2
$$
leading to the general solution
$$
U^n = c_1 + c_2 n.
$$
Here we see that given a $U^0$ and $U^1$ that the solution will still grow linearly with $n$ which will again lead to a divergent solution.
#### Example
Consider the linear multistep method
$$
U^{n+3} - 2 U^{n+2} + \frac{5}{4} U^{n+1} - \frac{1}{4} U^n = \frac{\Delta t}{4} f(U^n).
$$
Apply this to the ODE $u'(t) = 0$ and determine whether this method is zero-stable.
Applied to the ODE $u'(t) = 0$ we have the linear difference equation
$$
U^{n+3} - 2 U^{n+2} + \frac{5}{4} U^{n+1} - \frac{1}{4} U^n = 0
$$
leading to
$$
\rho(\xi) = \xi^3 - 2 \xi^2 + \frac{5}{4} \xi - \frac{1}{4} = 0
$$
whose solutions are
$$
\xi_1 = 1, \xi_2 = \xi_3 = 1 / 2
$$
with the general solution
$$
U^n = c_1 + c_2 \frac{1}{2^n} + c_3 n \frac{1}{2^n}
$$
which does converge due to the factor of $1 / 2^n$!
### Example: Adams Methods
The general form for all Adams methods take the form
$$
U^{n+r} = U^{n+r-1} + \Delta t \sum^r_{j=0} \beta_j f(U^{n+j})
$$
which has the characteristic polynomial (for the ODE $u'(t)=0$)
$$
\rho(\xi) = \xi^r - \xi^{r-1} = (\xi - 1) \xi^{r-1}
$$
leading to the roots $\xi_1 = 1$ and $\xi_2 = \xi_3 = \cdots = \xi_r = 0$ which satisfy the general zero-stability result and therefore all Adams methods are convergent.
## Absolute Stability
Although zero-stability guarantees stability it is much more difficult to work with in general as the limit $\Delta t \rightarrow 0$ can be difficult to compute. Instead we often consider a finite $\Delta t$ and examine if the method is stable for this particular choice of $\Delta t$. This has the practical upside that it will also tell us what particular $\Delta t$ will ensure that our method is indeed stable.
### Example
Consider the problem
$$u'(t) = \lambda (u - \cos t) - \sin t \quad \text{with} \quad u(0) = 1$$
whose exact solution is
$$u(t) = \cos t.$$
We can compute an estimate for what $\Delta t$ we need to use by examining the truncation error
$$\begin{aligned}
\tau &= \frac{1}{2} \Delta t u''(t) + \mathcal{O}(\Delta t^2) \\
&= -\frac{1}{2} \Delta t \cos t + \mathcal{O}(\Delta t^2)
\end{aligned}$$
and therefore
$$|E^n| \leq \Delta t \max_{0 \leq t \leq t_f} |\cos t| = \Delta t.$$
If we want a solution where $|E^n| < 10^{-3}$ then $\Delta t \approx 10^{-3}$. Turning to the application of Euler's method lets apply this to the case where $\lambda = -10$ and $\lambda = -2100$.
```
# Compare accuracy between Euler
f = lambda t, lam, u: lam * (u - numpy.cos(t)) - numpy.sin(t)
u_exact = lambda t: numpy.cos(t)
t_f = 2.0
num_steps = [2**n for n in range(4, 10)]
# num_steps = [2**n for n in range(15,20)]
delta_t = numpy.empty(len(num_steps))
error_10 = numpy.empty(len(num_steps))
error_2100 = numpy.empty(len(num_steps))
for (i, N) in enumerate(num_steps):
t = numpy.linspace(0, t_f, N)
delta_t[i] = t[1] - t[0]
# Compute Euler solution
U = numpy.empty(t.shape)
U[0] = 1.0
for (n, t_n) in enumerate(t[1:]):
U[n+1] = U[n] + delta_t[i] * f(t_n, -10.0, U[n])
error_10[i] = numpy.abs(U[-1] - u_exact(t_f)) / numpy.abs(u_exact(t_f))
U = numpy.empty(t.shape)
U[0] = 1.0
for (n, t_n) in enumerate(t[1:]):
U[n+1] = U[n] + delta_t[i] * f(t_n, -2100.0, U[n])
error_2100[i] = numpy.abs(U[-1] - u_exact(t_f)) / numpy.abs(u_exact(t_f))
# Plot error vs. delta_t
fig = plt.figure()
fig.set_figwidth(fig.get_figwidth() * 2)
axes = fig.add_subplot(1, 2, 1)
axes.loglog(delta_t, error_10, 'bo', label='Forward Euler')
order_C = lambda delta_x, error, order: numpy.exp(numpy.log(error) - order * numpy.log(delta_x))
axes.loglog(delta_t, order_C(delta_t[1], error_10[1], 1.0) * delta_t**1.0, 'r--', label="1st Order")
axes.legend(loc=4)
axes.set_title("Comparison of Errors")
axes.set_xlabel("$\Delta t$")
axes.set_ylabel("$|U(t_f) - u(t_f)|$")
axes = fig.add_subplot(1, 2, 2)
axes.loglog(delta_t, error_2100, 'bo', label='Forward Euler')
axes.loglog(delta_t, order_C(delta_t[1], error_2100[1], 1.0) * delta_t**1.0, 'r--', label="1st Order")
axes.set_title("Comparison of Errors")
axes.set_xlabel("$\Delta t$")
axes.set_ylabel("$|U(t_f) - u(t_f)|$")
plt.show()
```
So what went wrong with $\lambda = -2100$? The global error should go as
$$E^{n+1} = (1 + \Delta t \lambda) E^n - \Delta t T^n$$
If $\Delta t \approx 10^{-3}$ then for the case $\lambda = -10$ the previous global error is multiplied by
$$1 + 10^{-3} \cdot -10 = 0.99$$
which means the contribution from $E^n$ will slowly decrease as we take more time steps. For the other case we have
$$1 + 10^{-3} \cdot -2100 = -1.1$$
which means that for this $\Delta t$ the error made in previous time steps will grow! For this not to happen we would have to have $\Delta t < 1 / 2100$ which would lead to convergence again.
### Absolute Stability of the Forward Euler Method
Consider again the simple test problem $u'(t) = \lambda u$. We know from before that applying Euler's method to this problem leads to an update of the form
$$U_{n+1} = (1 + \Delta t \lambda) U_n.$$
As may have been clear from the last example, we know that if
$$|1 + \Delta t \lambda| \leq 1$$
that the method will be stable, this is called **absolute stability**. Note that the product of $\Delta t \lambda$ is what matters here and often we consider a **region of absolute stability** on the complex plain defined by the equation outlined where now $z = \Delta t \lambda$. This allows the values of $\lambda$ to be complex which can be an important case to consider, especially for systems of equations where the $\lambda$s are identified as the eigenvalues.
```
# Plot the region of absolute stability for Forward Euler
fig = plt.figure()
axes = fig.add_subplot(1, 1, 1)
t = numpy.linspace(0.0, 2.0 * numpy.pi, 100)
axes.fill(numpy.cos(t) - 1.0, numpy.sin(t), 'b')
axes.plot([-3, 3],[0.0, 0.0],'k--')
axes.plot([0.0, 0.0],[-3, 3],'k--')
axes.set_xlim((-3, 3.0))
axes.set_ylim((-3,3))
axes.set_aspect('equal')
axes.set_title("Absolute Stability Region for Forward Euler")
plt.show()
```
### General Stability Regions for Linear Multistep Methods
Going back to linear multistep methods and applying them in general to our test problem we have
$$
\sum^r_{j=0} \alpha_j U_{n+j} = \Delta t \sum^r_{j=0} \beta_j \lambda U_{n+j}
$$
which can be written as
$$
\sum^r_{j=0} (\alpha_j - \beta_j \Delta t \lambda) U_{n+j} = 0
$$
or using our notation of $z = \Delta t \lambda$ we have
$$
\sum^r_{j=0} (\alpha_j - \beta_j z) U_{n+j} = 0.
$$
This has a similar form to the linear difference equations considered above! Letting
$$
\rho(\xi) = \sum^r_{j=0} \alpha_j \xi^j
$$
and
$$
\sigma(\xi) = \sum^r_{j=0} \beta_j \xi^j
$$
we can write the expression above as
$$
\pi(\xi, z) = \rho(\xi) - z \sigma(\xi)
$$
called the **stability polynomial** of the the linear multi-step method.
It turns out that if the roots $\xi_i$ of this polynomial satisfy
$$
|\xi_i| \leq 1
$$
then the multi-step method is absolutely-stable. We then define the region of absolute stability as the values for $z$ for which this is true. This approach can also be applied to one-step methods.
### Example: Forward Euler's Method
Examining forward Euler's method we have
$$\begin{aligned}
0 &= U_{n+1} - U_n - \Delta t \lambda U_n \\
&= U_{n+1} - U_n (1 + \Delta t \lambda)\\
&= \xi - 1 (1 + z)\\
&=\pi(\xi, z)
\end{aligned}$$
whose root is $\xi = 1 + z$ and we have re-derived the stability region we had found before.
### Absolute Stability of the backward Euler Method
The backward version of Euler's method is defined as
$$
U_{n+1} = U_n + \Delta t f(t_{n+1}, U_{n+1}).
$$
Check to see if backward Euler is absolute stable.
If we again consider the test problem from before we find that
$$\begin{aligned}
0 &= U_{n+1} (1 - \Delta t \lambda) - U_n \\
&= \xi (1 - z) - 1
\end{aligned}$$
which has the root $\xi = \frac{1}{1 - z}$. We then have
$$
\left|\frac{1}{1-z}\right| \leq 1 \leftrightarrow |1 - z| \geq 1
$$
so in fact the stability region encompasses the entire complex plane except for a circle centered at $(1, 0)$ of radius 1 implying that the backward Euler method is in fact stable for any choice of $\Delta t$.
## Application to Stiff ODEs
Consider again the ODE we examined before
$$u'(t) = \lambda (u - \cos t) - \sin t$$
except this time with general initial condition $u(t_0) = \eta$. What happens to solutions that are slightly different from $\eta = 1$ or $t_0 = 0$? The general solution of the ODE is
$$u(t) = e^{\lambda (t - t_0)} (\eta - \cos t_0)) + \cos t$$.
```
# Plot "hairy" solutions to the ODE
u = lambda t_0, eta, lam, t: numpy.exp(lam * (t - t_0)) * (eta - numpy.cos(t_0)) + numpy.cos(t)
fig = plt.figure()
for lam in [-1, -10]:
fig = plt.figure()
axes = fig.add_subplot(1, 1, 1)
for eta in numpy.linspace(-1, 1, 10):
for t_0 in numpy.linspace(0.0, 9.0, 10):
t = numpy.linspace(t_0,10.0,100)
axes.plot(t, u(t_0, eta, lam, t),'b')
t = numpy.linspace(0.0,10.0,100)
axes.plot(t, numpy.cos(t), 'r', linewidth=5)
axes.set_title("Perturbed Solutions $\lambda = %s$" % lam)
axes.set_xlabel('$t$')
axes.set_ylabel('$u(t)$')
plt.show()
# Plot "inverse hairy" solutions to the ODE
u = lambda t_0, eta, lam, t: numpy.exp(lam * (t - t_0)) * (eta - numpy.cos(t_0)) + numpy.cos(t)
fig = plt.figure()
num_steps = 10
error = numpy.ones(num_steps) * 1.0
t_hat = numpy.linspace(0.0, 10.0, num_steps + 1)
t_whole = numpy.linspace(0.0, 10.0, 1000)
fig = plt.figure()
axes = fig.add_subplot(1, 1, 1)
eta = 1.0
lam = 0.1
for n in range(1,num_steps):
t = numpy.linspace(t_hat[n-1], t_hat[n], 100)
U = u(t_hat[n-1], eta, lam, t)
axes.plot(t, U, 'b')
axes.plot(t_whole, u(t_hat[n-1], eta, lam, t_whole),'b--')
axes.plot([t[-1], t[-1]], (U[-1], U[-1] + -1.0**n * error[n]), 'r')
eta = U[-1] + -1.0**n * error[n]
t = numpy.linspace(0.0, 10.0, 100)
axes.plot(t, numpy.cos(t), 'g')
axes.set_title("Perturbed Solutions $\lambda = %s$" % lam)
axes.set_xlabel('$t$')
axes.set_ylabel('$u(t)$')
axes.set_ylim((-10,10))
plt.show()
```
### Example: Chemical systems
Consider the transition of a chemical $A$ to a chemical $C$ through the process
$$A \overset{K_1}{\rightarrow} B \overset{K_2}{\rightarrow} C.$$
If we let
$$\vec{u} = \begin{bmatrix} [A] \\ [B] \\ [C] \end{bmatrix}$$
then we can model this simple chemical reaction with the system of ODEs
$$\frac{\text{d} \vec{u}}{\text{d} t} =
\begin{bmatrix}
-K_1 & 0 & 0 \\
K_1 & -K_2 & 0 \\
0 & K_2 & 0
\end{bmatrix} \vec{u}$$
The solution of this system is of the form
$$u_j(t) = c_{j1} e^{-K_1 t} + c_{j2}e^{-K_2 t} + c_{j3}$$
```
# Solve the chemical systems example
# Problem parameters
K_1 = 3
K_2 = 1
# K_1 = 30.0
# K_2 = 1.0
A = numpy.array([[-K_1, 0, 0], [K_1, -K_2, 0], [0, K_2, 0]])
f = lambda u: numpy.dot(A, u)
t = numpy.linspace(0.0, 8.0, 128)
delta_t = t[1] - t[0]
U = numpy.empty((t.shape[0], 3))
U[0, :] = [2.5, 5.0, 2.0]
for n in range(t.shape[0] - 1):
U[n+1, :] = U[n, :] + delta_t * f(U[n, :])
fig = plt.figure()
axes = fig.add_subplot(1, 1, 1)
axes.plot(t, U)
axes.set_title("Chemical System")
axes.set_xlabel("$t$")
axes.set_title("$[A], [B], [C]$")
axes.set_ylim((0.0, 10.))
plt.show()
```
### What is stiffness?
In general a **stiff** ODE is one where $u'(t) \ll f'(t, u)$. For systems of ODEs the **stiffness ratio**
$$\frac{\max_p |\lambda_p|}{\min_p |\lambda_p|}$$
can be used to characterize the stiffness of the system. In our last example this ratio was $K_1 / K_2$ if $K_1 > K_2$. As we increased this ratio we observed that the numerical method became unstable only a reduction in $\Delta t$ lead to stable solution again. For explicit time step methods this is problematic as the reduction of the time step for only one of the species leads to very expensive evaluations. For example, forward Euler has the stability criteria
$$|1 + \Delta t \lambda| < 1$$
where $\lambda$ will have to be the maximum eigenvalue of the system.
```
# Plot the region of absolute stability for Forward Euler
fig = plt.figure()
axes = fig.add_subplot(1, 1, 1)
t = numpy.linspace(0.0, 2.0 * numpy.pi, 100)
K_1 = 3.0
K_2 = 1.0
delta_t = 1.0
eigenvalues = [-K_1, -K_2]
axes.fill(numpy.cos(t) - 1.0, numpy.sin(t), color='lightgray')
for lam in eigenvalues:
print(lam * delta_t)
axes.plot(lam * delta_t, 0.0, 'ko')
axes.plot([-3, 3],[0.0, 0.0],'k--')
axes.plot([0.0, 0.0],[-3, 3],'k--')
# axes.set_xlim((-3, 1))
axes.set_ylim((-2,2))
axes.set_aspect('equal')
axes.set_title("Absolute Stability Region for Forward Euler")
plt.show()
```
### A-Stability
What if we could expand the absolute stability region to encompass more of the left-half plane or even better, all of it. A method that has this property is called **A-stable**. We have already seen one example of this with backward Euler which has a stability region of
$$|1 - z| \geq 1$$
which covers the full left-half plane.
It turns out that for linear multi-step methods a theorem by Dahlquist proves that there are no LMMs that satisfies the A-stability criterion beyond second order (trapezoidal rule). There are higher-order Runge-Kutta methods do however.
Perhaps this is too restrictive though. Often large eigenvalues for systems (for instance coming from a PDE discretization for the heat equation) lie completely on the real line. If the stability region can encompass as much of the real line as possible while leaving out the rest of the left-half plane we can possibly get a more efficient method. There are a number of methods that can be constructed that have this property but are higher-order.
```
# Plot the region of absolute stability for Backward Euler
fig = plt.figure()
axes = fig.add_subplot(1, 1, 1)
t = numpy.linspace(0.0, 2.0 * numpy.pi, 100)
K_1 = 3.0
K_2 = 1.0
delta_t = 1.0
eigenvalues = [-K_1, -K_2]
axes.set_facecolor('lightgray')
axes.fill(numpy.cos(t) + 1.0, numpy.sin(t), 'w')
for lam in eigenvalues:
print(lam * delta_t)
axes.plot(lam * delta_t, 0.0, 'ko')
axes.plot([-3, 3],[0.0, 0.0],'k--')
axes.plot([0.0, 0.0],[-3, 3],'k--')
# axes.set_xlim((-3, 1))
axes.set_ylim((-2,2))
axes.set_aspect('equal')
axes.set_title("Absolute Stability Region for Backward Euler")
plt.show()
```
### L-Stability
It turns out not all A-stable methods are alike. Consider the backward Euler method and the trapezoidal method. The stability polynomial for the trapezoidal method is
$$\begin{aligned}
0 &= U_{n+1} - U_n - \Delta t \frac{1}{2} (\lambda U_n + \lambda U_{n+1}) \\
&= U_{n+1}\left(1 - \frac{1}{2} \Delta t \lambda \right ) - U_n \left(1 + \frac{1}{2}\Delta t \lambda \right) \\
&= \left(\xi - \frac{1 + \frac{1}{2}z}{1 - \frac{1}{2} z}\right) \left(1 - \frac{1}{2} z \right )\\
\end{aligned}$$
which shows that it is A-stable. Lets apply both these methods to a problem we have seen before and see what happens.
```
# Compare accuracy between Euler
f = lambda t, lam, u: lam * (u - numpy.cos(t)) - numpy.sin(t)
u_exact = lambda t_0, eta, lam, t: numpy.exp(lam * (t - t_0)) * (eta - numpy.cos(t_0)) + numpy.cos(t)
t_0 = 0.0
eta = 1.5
lam = -1e6
num_steps = [10, 20, 40, 50]
delta_t = numpy.empty(len(num_steps))
error_euler = numpy.empty(len(num_steps))
error_trap = numpy.empty(len(num_steps))
for (i, N) in enumerate(num_steps):
t = numpy.linspace(0, t_f, N)
delta_t[i] = t[1] - t[0]
u = u_exact(t_0, eta, lam, t_f)
# Compute Euler solution
U_euler = numpy.empty(t.shape)
U_euler[0] = eta
for (n, t_n) in enumerate(t[1:]):
U_euler[n+1] = (U_euler[n] - lam * delta_t[i] * numpy.cos(t_n) - delta_t[i] * numpy.sin(t_n)) / (1.0 - lam * delta_t[i])
error_euler[i] = numpy.abs(U_euler[-1] - u) / numpy.abs(u)
# Compute using trapezoidal
U_trap = numpy.empty(t.shape)
U_trap[0] = eta
for (n, t_n) in enumerate(t[1:]):
U_trap[n+1] = (U_trap[n] + delta_t[i] * 0.5 * f(t_n, lam, U_trap[n]) - 0.5 * lam * delta_t[i] * numpy.cos(t_n) - 0.5 * delta_t[i] * numpy.sin(t_n)) / (1.0 - 0.5 * lam * delta_t[i])
error_trap[i] = numpy.abs(U_trap[-1] - u)
# Plot error vs. delta_t
fig = plt.figure()
fig.set_figwidth(fig.get_figwidth() * 2)
axes = fig.add_subplot(1, 2, 1)
axes.plot(t, U_euler, 'ro-')
axes.plot(t, u_exact(t_0, eta, lam, t),'k')
axes = fig.add_subplot(1, 2, 2)
axes.loglog(delta_t, error_euler, 'bo')
order_C = lambda delta_x, error, order: numpy.exp(numpy.log(error) - order * numpy.log(delta_x))
axes.loglog(delta_t, order_C(delta_t[1], error_euler[1], 1.0) * delta_t**1.0, 'r--', label="1st Order")
axes.loglog(delta_t, order_C(delta_t[1], error_euler[1], 2.0) * delta_t**2.0, 'b--', label="2nd Order")
axes.legend(loc=4)
axes.set_title("Comparison of Errors for Backwards Euler")
axes.set_xlabel("$\Delta t$")
axes.set_ylabel("$|U(t_f) - u(t_f)|$")
# Plots for trapezoid
fig = plt.figure()
fig.set_figwidth(fig.get_figwidth() * 2)
axes = fig.add_subplot(1, 2, 1)
axes.plot(t, U_trap, 'ro-')
axes.plot(t, u_exact(t_0, eta, lam, t),'k')
axes = fig.add_subplot(1, 2, 2)
axes.loglog(delta_t, error_trap, 'bo', label='Forward Euler')
order_C = lambda delta_x, error, order: numpy.exp(numpy.log(error) - order * numpy.log(delta_x))
axes.loglog(delta_t, order_C(delta_t[1], error_trap[1], 1.0) * delta_t**1.0, 'r--', label="1st Order")
axes.loglog(delta_t, order_C(delta_t[1], error_trap[1], 2.0) * delta_t**2.0, 'b--', label="2nd Order")
axes.legend(loc=4)
axes.set_title("Comparison of Errors for Trapezoidal Rule")
axes.set_xlabel("$\Delta t$")
axes.set_ylabel("$|U(t_f) - u(t_f)|$")
plt.show()
```
It turns out that if we look at a one-step method and define the following ratio
$$U_{n+1} = R(z) U_n$$
we can define another form of stability, called **L-stability**, where we require that the method is A-stable and that
$$\lim_{z \rightarrow \infty} |R(z)| = 0.$$
Backwards Euler is L-stable while the trapezoidal method is not.
## Backward Differencing Formulas
A class of LMM methods that are useful for stiff ODE problems are the backward difference formula (BDF) methods which have the form
$$\alpha_0 U_n + \alpha_1 U_{n+1} + \cdots + \alpha_r U_{n+r} = \Delta \beta_r f(U_{n+r})$$
These methods can be derived directly from backwards finite differences from the point $U_{n+r}$ and the rest of the points back in time. One can then derive r-step methods that are rth-order accurate this way. Some of the methods are
$$\begin{aligned}
r = 1:& & U_{n+1} - U_n = \Delta t f(U_{n+1}) \\
r = 2:& &3 U_{n+2} - 4 U_{n+1} + U_n = 2 \Delta t f(U_{n+1}) \\
r = 3:& &11U_{n+3} - 18U_{n+2} + 9U_{n+1} - 2 U_n = 6 \Delta t f(U_{n+3}) \\
r = 4:& &25 U_{n+4} - 48 U_{n+3} +36 U_{n+2} -16 U_{n+1} +3 U_n = 12 \Delta t f(U_{n+4})
\end{aligned}$$
## Plotting Stability Regions
If we think of the roots of the stability polynomial $\xi_j$ as complex numbers and write them in exponential form
$$
\xi_j = |\xi_j| e^{i \theta}.
$$
Here $|\xi_j|$ is the modulus (or magnitude) or the complex number and is defined as $|\xi_j| = x^2 + y^2$ where $\xi_j = x + i j$. If the $\xi_j$s are on the boundary of the absolute stability region then we know that $|\xi_j| = 1$. Using this in conjunction with the stability polynomial then leads to
$$
\rho(e^{i\theta}) - z \sigma(e^{i\theta}) = 0
$$
which solving for $z$ leads to
$$
z(\theta) = \frac{\rho(e^{i\theta})}{\sigma(e^{i\theta})}.
$$
As an example consider the Adams-Bashforth 2-stage method. The stability polynomial can be found as
$$\begin{aligned}
U_{n+2} &= U_{n+1} + \frac{\Delta t}{2} (-f(U_n) + 3 f(U_{n+1})) \\
\pi(\xi, z) &= U_{n+2} - U_{n+1} - \frac{\Delta t}{2} (-f(U_n) + 3 f(U_{n+1})) = 0 \\
&= U_{n+2} - U_{n+1} - \frac{1}{2} (\Delta t \lambda U_n - 3 \Delta t \lambda U_{n+1}) \\
&= 2 \xi^2 - 2 \xi + 3 z\xi - z \\
&= \rho(\xi, z) + z \sigma(\xi, z)
\end{aligned}$$
where
$$
\rho(\xi, z) = 2 ( \xi - 1) \xi ~~~ \text{and} ~~~ \sigma(\xi, z) = 3 \xi - 1
$$
so that
$$
z(\theta) = \frac{2 (\xi - 1) \xi}{3 \xi - 1}.
$$
This does not necessarily ensure that given a $\theta$ that $z(\theta)$ will lie on the absolute stability region's boundary. This can occur when $\xi_j = 1$ but to the left and right of the curve $\xi_j > 1$ and so therefore does not mark the boundary of the region. To determine whether a particular region outlined by this curve is inside or outside of the stability region we can evaluate all the roots of $\pi(\xi, z)$ at some $z$ inside of the region in question and see if then $\forall j, \xi_j < 1$.
For one-step methods this becomes easier, if we look at the ratio $R(z)$ we defined earlier as
$$
U_{n+1} = R(z) U_n
$$
in the case of the pth order Taylor series method applied to $u'(t) = \lambda u$ we get
$$\begin{aligned}
U_{n+1} &= U_n + \Delta t \lambda U_n + \frac{1}{2}\Delta t^2 \lambda^2 U_n + \cdots + \frac{1}{p!}\Delta t^p \lambda^p U_n \\
&=\left(1 + z + \frac{1}{2} z^2 + \frac{1}{6} z^3 + \cdots +\frac{1}{p!}z^p\right) U_n \Rightarrow \\
R(z) &= 1 + z + \frac{1}{2} z^2 + \frac{1}{6} z^3 + \cdots +\frac{1}{p!}z^p.
\end{aligned}$$
Setting $R(z) = e^{i\theta}$ could lead to a way for solving for the boundary but (where $|R(z)| = 1$) but this is very difficult to do in general. Instead if we plot the contours of $R(z)$ in the complex plain we can pick out the $R(z)=1$ contour and plot that.
```
theta = numpy.linspace(0.0, 2.0 * numpy.pi, 100)
# ==================================
# Forward euler
fig = plt.figure()
fig.set_figwidth(fig.get_figwidth() * 2.0)
fig.set_figheight(fig.get_figheight() * 2.0)
axes = fig.add_subplot(2, 2, 1)
x = numpy.linspace(-2, 2, 100)
y = numpy.linspace(-2, 2, 100)
X, Y = numpy.meshgrid(x, y)
Z = X + 1j * Y
# Evaluate which regions are actually in stability region
print("Forward Euler's method - Stability region tests")
z = -1 + 1j * 0
print(" Inside of circle: ", numpy.abs(1.0 + z))
z = -3 + 1j * 0
print(" Outside of circle: ", numpy.abs(1.0 + z))
axes.contour(X, Y, numpy.abs(1.0 + Z), levels=[1.0])
axes.plot(x, numpy.zeros(x.shape),'k')
axes.plot(numpy.zeros(y.shape), y,'k')
axes.set_aspect('equal')
axes.set_title("Forward Euler")
# ==================================
# Backwards Euler
axes = fig.add_subplot(2, 2, 2)
x = numpy.linspace(-2, 2, 100)
y = numpy.linspace(-2, 2, 100)
X, Y = numpy.meshgrid(x, y)
Z = X + 1j * Y
# Evaluate which regions are actually in stability region
print("Backward Euler's method - Stability region tests")
z = 1 + 1j * 0
print(" Inside of circle: ", numpy.abs(1.0 + z))
z = -1 + 1j * 0
print(" Outside of circle: ", numpy.abs(1.0 + z))
axes.contour(X, Y, numpy.abs(1.0 / (1.0 - Z)), levels=[1.0])
axes.plot(x, numpy.zeros(x.shape),'k')
axes.plot(numpy.zeros(y.shape), y,'k')
axes.set_aspect('equal')
axes.set_title("Backwards Euler")
# ==================================
# Taylor series method of order 4
axes = fig.add_subplot(2, 2, 3)
x = numpy.linspace(-5, 5, 100)
y = numpy.linspace(-5, 5, 100)
X, Y = numpy.meshgrid(x, y)
Z = X + 1j * Y
# Evaluate which regions are actually in stability region
print("Taylor series method of order 4 - Stability region tests")
z = -1 + 1j * 0
print(" Inside of strange region: ", numpy.abs(1.0 + z))
z = -4 + 1j * 0
print(" Outside of strange region: ", numpy.abs(1.0 + z))
axes.contour(X, Y, numpy.abs(1 + Z + 0.5 * Z**2 + 1.0/6.0 * Z**3 + 1.0 / 24.0 * Z**4), levels=[1.0])
axes.plot(x, numpy.zeros(x.shape),'k')
axes.plot(numpy.zeros(y.shape), y,'k')
axes.set_aspect('equal')
axes.set_title("4th Order Taylor Series")
# ==================================
# 2-step Adams-Bashforth
theta = numpy.linspace(0.0, 2.0 * numpy.pi, 1000)
xi = numpy.exp(1j * theta)
rho_2AB = lambda xi: (xi - 1.0) * xi
sigma_2AB = lambda xi: (3.0 * xi - 1.0) / 2.0
z_2AB = rho_2AB(xi) / sigma_2AB(xi)
z = rho_2AB(xi) / sigma_2AB(xi)
axes = fig.add_subplot(2, 2, 4)
axes.plot(z_2AB.real, z_2AB.imag)
axes.plot(x, numpy.zeros(x.shape),'k')
axes.plot(numpy.zeros(y.shape), y,'k')
axes.set_title("2-step Adams-Bashforth")
axes.set_aspect('equal')
axes.set_xlim([-2, 3])
axes.set_ylim([-2, 2])
plt.show()
```
| github_jupyter |
# Introduction #
Run this cell to set everything up!
```
# Setup feedback system
from learntools.core import binder
binder.bind(globals())
from learntools.time_series.ex1 import *
# Setup notebook
from pathlib import Path
from learntools.time_series.style import * # plot style settings
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sns
from sklearn.linear_model import LinearRegression
data_dir = Path('../input/ts-course-data/')
comp_dir = Path('../input/store-sales-time-series-forecasting')
book_sales = pd.read_csv(
data_dir / 'book_sales.csv',
index_col='Date',
parse_dates=['Date'],
).drop('Paperback', axis=1)
book_sales['Time'] = np.arange(len(book_sales.index))
book_sales['Lag_1'] = book_sales['Hardcover'].shift(1)
book_sales = book_sales.reindex(columns=['Hardcover', 'Time', 'Lag_1'])
ar = pd.read_csv(data_dir / 'ar.csv')
dtype = {
'store_nbr': 'category',
'family': 'category',
'sales': 'float32',
'onpromotion': 'uint64',
}
store_sales = pd.read_csv(
comp_dir / 'train.csv',
dtype=dtype,
parse_dates=['date'],
infer_datetime_format=True,
)
store_sales = store_sales.set_index('date').to_period('D')
store_sales = store_sales.set_index(['store_nbr', 'family'], append=True)
average_sales = store_sales.groupby('date').mean()['sales']
```
--------------------------------------------------------------------------------
One advantage linear regression has over more complicated algorithms is that the models it creates are *explainable* -- it's easy to interpret what contribution each feature makes to the predictions. In the model `target = weight * feature + bias`, the `weight` tells you by how much the `target` changes on average for each unit of change in the `feature`.
Run the next cell to see a linear regression on *Hardcover Sales*.
```
fig, ax = plt.subplots()
ax.plot('Time', 'Hardcover', data=book_sales, color='0.75')
ax = sns.regplot(x='Time', y='Hardcover', data=book_sales, ci=None, scatter_kws=dict(color='0.25'))
ax.set_title('Time Plot of Hardcover Sales');
```
# 1) Interpret linear regression with the time dummy
The linear regression line has an equation of (approximately) `Hardcover = 3.33 * Time + 150.5`. Over 6 days how much on average would you expect hardcover sales to change? After you've thought about it, run the next cell.
```
# View the solution (Run this line to receive credit!)
q_1.check()
# Uncomment the next line for a hint
#_COMMENT_IF(PROD)_
q_1.hint()
```
-------------------------------------------------------------------------------
Interpreting the regression coefficients can help us recognize serial dependence in a time plot. Consider the model `target = weight * lag_1 + error`, where `error` is random noise and `weight` is a number between -1 and 1. The `weight` in this case tells you how likely the next time step will have the same sign as the previous time step: a `weight` close to 1 means `target` will likely have the same sign as the previous step, while a `weight` close to -1 means `target` will likely have the opposite sign.
# 2) Interpret linear regression with a lag feature
Run the following cell to see two series generated according to the model just described.
```
fig, (ax1, ax2) = plt.subplots(2, 1, figsize=(11, 5.5), sharex=True)
ax1.plot(ar['ar1'])
ax1.set_title('Series 1')
ax2.plot(ar['ar2'])
ax2.set_title('Series 2');
```
One of these series has the equation `target = 0.95 * lag_1 + error` and the other has the equation `target = -0.95 * lag_1 + error`, differing only by the sign on the lag feature. Can you tell which equation goes with each series?
```
# View the solution (Run this cell to receive credit!)
q_2.check()
# Uncomment the next line for a hint
#_COMMENT_IF(PROD)_
q_2.hint()
```
-------------------------------------------------------------------------------
Now we'll get started with the *Store Sales - Time Series Forecasting* competition data. The entire dataset comprises almost 1800 series recording store sales across a variety of product families from 2013 into 2017. For this lesson, we'll just work with a single series (`average_sales`) of the average sales each day.
# 3) Fit a time-step feature
Complete the code below to create a linear regression model with a time-step feature on the series of average product sales. The target is in a column called `'sales'`.
```
from sklearn.linear_model import LinearRegression
df = average_sales.to_frame()
# YOUR CODE HERE: Create a time dummy
time = ____
#_UNCOMMENT_IF(PROD)_
#df['time'] = time
# YOUR CODE HERE: Create training data
X = ____ # features
y = ____ # target
# Train the model
#_UNCOMMENT_IF(PROD)_
#model = LinearRegression()
#_UNCOMMENT_IF(PROD)_
#model.fit(X, y)
# Store the fitted values as a time series with the same time index as
# the training data
#_UNCOMMENT_IF(PROD)_
#y_pred = pd.Series(model.predict(X), index=X.index)
# Check your answer
q_3.check()
# Lines below will give you a hint or solution code
#_COMMENT_IF(PROD)_
q_3.hint()
#_COMMENT_IF(PROD)_
q_3.solution()
#%%RM_IF(PROD)%%
from sklearn.linear_model import LinearRegression
df = average_sales.to_frame()
time = np.ones_like(df.index)
df['time'] = time
X = df.loc[:, ['time']]
y = df.loc[:, 'sales']
model = LinearRegression()
model.fit(X, y)
y_pred = pd.Series(model.predict(X), index=X.index)
q_3.assert_check_failed()
#%%RM_IF(PROD)%%
from sklearn.linear_model import LinearRegression
df = average_sales.to_frame()
time = np.arange(len(df.index))
df['time'] = time
X = df.loc[:, ['sales']]
y = df.loc[:, 'time']
model = LinearRegression()
model.fit(X, y)
y_pred = pd.Series(model.predict(X), index=X.index)
q_3.assert_check_failed()
#%%RM_IF(PROD)%%
from sklearn.linear_model import LinearRegression
df = average_sales.to_frame()
time = np.arange(len(df.index))
df['time'] = time
X = df.loc[:, ['time']]
y = df.loc[:, 'sales']
model = LinearRegression()
model.fit(X, y)
y_pred = pd.Series(model.predict(X), index=X.index)
q_3.assert_check_passed()
```
Run this cell if you'd like to see a plot of the result.
```
ax = y.plot(**plot_params, alpha=0.5)
ax = y_pred.plot(ax=ax, linewidth=3)
ax.set_title('Time Plot of Total Store Sales');
```
-------------------------------------------------------------------------------
# 4) Fit a lag feature to Store Sales
Complete the code below to create a linear regression model with a lag feature on the series of average product sales. The target is in a column of `df` called `'sales'`.
```
df = average_sales.to_frame()
# YOUR CODE HERE: Create a lag feature from the target 'sales'
lag_1 = ____
#_UNCOMMENT_IF(PROD)_
#df['lag_1'] = lag_1 # add to dataframe
#_UNCOMMENT_IF(PROD)_
#X = df.loc[:, ['lag_1']].dropna() # features
#_UNCOMMENT_IF(PROD)_
#y = df.loc[:, 'sales'] # target
#_UNCOMMENT_IF(PROD)_
#y, X = y.align(X, join='inner') # drop corresponding values in target
# YOUR CODE HERE: Create a LinearRegression instance and fit it to X and y.
model = ____
# YOUR CODE HERE: Create Store the fitted values as a time series with
# the same time index as the training data
y_pred = ____
# Check your answer
q_4.check()
# Lines below will give you a hint or solution code
q_4.hint()
q_4.solution()
#%%RM_IF(PROD)%%
df = average_sales.to_frame()
lag_1 = df['sales']
df['lag_1'] = lag_1
X = df.loc[:, ['lag_1']]
X.dropna(inplace=True) # drop missing values in the feature set
y = df.loc[:, 'sales'] # create the target
y, X = y.align(X, join='inner') # drop corresponding values in target
model = LinearRegression()
model.fit(X, y)
y_pred = pd.Series(model.predict(X), index=X.index)
q_4.assert_check_failed()
#%%RM_IF(PROD)%%
df = average_sales.to_frame()
lag_1 = df['sales'].shift(-1)
df['lag_1'] = lag_1
X = df.loc[:, ['lag_1']]
X.dropna(inplace=True) # drop missing values in the feature set
y = df.loc[:, 'sales'] # create the target
y, X = y.align(X, join='inner') # drop corresponding values in target
model = LinearRegression()
model.fit(X, y)
y_pred = pd.Series(model.predict(X), index=X.index)
q_4.assert_check_failed()
#%%RM_IF(PROD)%%
df = average_sales.to_frame()
lag_1 = df['sales'].shift(1)
df['lag_1'] = lag_1
X = df.loc[:, ['sales']]
X.dropna(inplace=True) # drop missing values in the feature set
y = df.loc[:, 'lag_1'] # create the target
y.dropna(inplace=True)
y, X = y.align(X, join='inner') # drop corresponding values in target
model = LinearRegression()
model.fit(X, y)
y_pred = pd.Series(model.predict(X), index=X.index)
q_4.assert_check_failed()
#%%RM_IF(PROD)%%
df = average_sales.to_frame()
lag_1 = df['sales'].shift(1)
df['lag_1'] = lag_1
X = df.loc[:, ['lag_1']]
X.dropna(inplace=True) # drop missing values in the feature set
y = df.loc[:, 'sales'] # create the target
y, X = y.align(X, join='inner') # drop corresponding values in target
model = LinearRegression()
model.fit(X, y)
y_pred = pd.Series(model.predict(X), index=X.index)
q_4.assert_check_passed()
```
Run the next cell if you'd like to see the result.
```
fig, ax = plt.subplots()
ax.plot(X['lag_1'], y, '.', color='0.25')
ax.plot(X['lag_1'], y_pred)
ax.set(aspect='equal', ylabel='sales', xlabel='lag_1', title='Lag Plot of Average Sales');
```
# Keep Going #
[**Model trend**](#$NEXT_NOTEBOOK_URL$) in time series with moving average plots and the time dummy.
| github_jupyter |
# Window Function on 2D clustering
steps are :
1. Transform the theory cell to the theory omega
2.1 Transfrom the window cell to the window omega
2.2 Compute the window omega directly with paircounts
3. Multiply the theory omega by the window omega
4. Transform back the window-convolved theory omega to cell
```
## import modules
%matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import healpy as hp
from scipy.optimize import curve_fit
from scipy.interpolate import InterpolatedUnivariateSpline as IUS
from scipy.special import eval_legendre as scipyLeg
from time import time
import logging
import sys
sys.path.insert(0, '/Users/rezaie/github/LSSutils')
from LSSutils import setup_logging
setup_logging('info')
plt.rc('font', size=20)#, family='serif')
def poly(x, *params):
ans = params[0]
for i in range(1, len(params)):
ans += np.power(x, i)*params[i]
return ans
def gauleg(ndeg, a=-1.0, b=1.0):
'''
Gauss-Legendre (default interval is [-1, 1])
'''
x, w = np.polynomial.legendre.leggauss(ndeg)
# Translate x values from the interval [-1, 1] to [a, b]
t = 0.5*(x + 1)*(b - a) + a
w *= 0.5*(b - a)
return t, w
def xi2cl(x, w, xi, nlmax):
'''
calculates Cell from omega
'''
cl = []#np.zeros(nlmax+1)
#m = np.arange(nlmax+1)
xiw = xi*w
for i in range(nlmax+1):
Pl = np.polynomial.Legendre.basis(i)(x)
#cl[i] = (xi * Pl * w).sum()
cl.append((xiw * Pl).sum())
cl = np.array(cl)
cl *= 2.*np.pi
return cl
def cl2xi(cell, costheta):
'''
calculates omega from Cell at Cos(theta)
'''
inv4pi = 0.07957747155 # 0.25/(np.pi)
ell = np.arange(cell.size)
coef = (2*ell+1) * cell
coef *= inv4pi
y = np.polynomial.legendre.legval(costheta, c=coef, tensor=False)
return y
def apply_window_v0(clth, theta, xiw, theory=True):
Xiw = IUS(np.cos(theta), xiw)
lmax = clth.size-1
x,w = np.polynomial.legendre.leggauss(lmax)
xith = cl2xi(clth, x)
xiw = Xiw(x)
if theory:
xif = xith * xiw
else:
xif = xith / xiw
clf = xi2cl(x, w, xif, lmax)
return clf, (x, xif)
class CellTheory:
logger = logging.getLogger('CellTheory')
def __init__(self,
cellfile='/Volumes/TimeMachine/data/mocks/fftloggz0.85redbias2.out',
lmax=10000):
self.l_mod, self.cl_mod = np.loadtxt(cellfile, usecols=(0,1), unpack=True)
self.lmax = int(self.l_mod.max())
self.lmin = int(self.l_mod.min())
self.logger.info(f'ellmin = {self.lmin}, C_ellmin = {self.cl_mod[0]}')
self.logger.info(f'ellmax = {self.lmax}, C_ellmax = {self.cl_mod[-1]}')
if self.lmin != 0.0:
self.logger.warning(f'insert 0.0 for ell=0')
self.cl_mod = np.insert(self.cl_mod, 0, 0.0)
if lmax > self.lmax:
self.logger.info(f'extend to {lmax}')
self._extend(lmax)
self.lmax = lmax
def convolve(self, XIW, lmax=512, ngauss=4*1024):
assert lmax <= self.lmax, 'lmax is too high'
t0 = time()
#--- convolve the theory with the window
cos_sep, weights = np.polynomial.legendre.leggauss(ngauss)
mid_sep = np.arccos(cos_sep)
xi_window = XIW(mid_sep)
xi_model = self.toxi(cos_sep)
t1 = time()
print(f'cl2xi : {t1-t0:6.1f} s')
xi_mod_wind = xi_model * xi_window
scalefactor = xi_model.sum()/xi_mod_wind.sum()
xi_mod_wind *= scalefactor
cl_mod_wind = xi2cl(cos_sep, weights, xi_mod_wind, lmax)
t2 = time()
print(f'xi2cl : {t2-t1:6.1f} s')
return {'ell':np.arange(lmax+1),
'Cell':self.cl_mod_ex[:lmax+1],
'Cellconv':cl_mod_wind,
'Xiwindow':xi_window,
'Ximodel':xi_model}
def _extend(self, lmax):
# fit
self.params,_ = curve_fit(self._lncl,
np.log(self.l_mod[-10:]),
np.log(self.cl_mod[-10:]),
p0=[1., 0.])
clmod_ext = list(self.cl_mod)
for ell in range(self.lmax, lmax+1):
clmod_ext.append(np.exp(self._lncl(np.log(ell), *self.params)))
self.cl_mod_ex = np.array(clmod_ext)
def _lncl(self, lnl, *params):
return params[0]+lnl*params[1]-(np.exp(lnl)/self.lmax-1.)
def toxi(self, cos_sep):
return cl2xi(self.cl_mod_ex, cos_sep)
def plot(self, ax=None, **kwargs):
if ax is None:
fig, ax = plt.subplots(figsize=(8, 6))
if hasattr(self, 'cl_mod_ex'):
ax.plot(self.cl_mod_ex, 'y-')
ax.plot(self.cl_mod, 'g--')
ax.set(xscale='log', xlabel=r'$\ell$', ylabel=r'C$_\ell$')
def load(self,
ximodel='/Volumes/TimeMachine/data/DR7/theta_omega_theory.txt'):
self.sep_rad, self.xi_mod = np.loadtxt(ximodel).T
self.dsep_rad = np.diff(self.sep_rad)
class XiWindow:
def __init__(self,
xifile='/Volumes/TimeMachine/data/DR7/dr7_window.cut.256.npy',
dof=8):
'''
'''
result = np.load(xifile, allow_pickle=True)
sep = result[0]
self.rr = result[1][0]
if sep[0] > sep[1]:
print('reordering arrays i.e., increasing')
sep = sep[::-1]
self.rr = self.rr[::-1]
self.mid_sep = 0.5*(sep[:-1]+sep[1:])
self.dsep = np.diff(sep)
# norm
self.rr_normed = self.rr / (np.sin(self.mid_sep)*self.dsep)
self.rr_normed /= self.rr_normed[0]
# fit
self._fit(dof)
norm = self(0.0)
# renormaliza
self.params = [p/norm for p in self.params]
self.rr_normed /= norm
def __call__(self, mid_sep):
return poly(mid_sep, *self.params)
def _fit(self, dof=8, nmodes_ext=50):
mid_sep_ext = list(self.mid_sep)
rr_ext = list(self.rr_normed)
for j in range(nmodes_ext):
# right end
mid_sep_ext.append(self.mid_sep[-1]+(j+1)*self.dsep[-1-j])
rr_ext.append(self.rr_normed[-1-j])
# left end
#mid_sep_ext.insert(0, xi_wind.mid_sep[0]-(j+1)*xi_wind.dsep[0])
#rr_ext.insert(0, xi_wind.rr_normed[j+1])
p0 = [0 for i in range(dof)]
self.params,_ = curve_fit(poly, mid_sep_ext, rr_ext, p0=p0)
frac = hp.read_map('/Volumes/TimeMachine/data/DR7/frac.hp.256.fits', verbose=False)
mask = hp.read_map('/Volumes/TimeMachine/data/DR7/mask.cut.hp.256.fits', verbose=False) > 0
print(frac[mask].sum())
frac = hp.read_map('/Volumes/TimeMachine/data/mocks/fracgood.hp256.fits', verbose=False)
mask = hp.read_map('/Volumes/TimeMachine/data/mocks/mask.cut.w.hp.256.fits', verbose=False) > 0
print(frac[mask].sum())
```
## Step 1: Transfor theory Cell to Omega
```
# read paircount
paircount = '/Volumes/TimeMachine/data/DR7/dr7_window.cut.256.npy'
XIW = XiWindow(paircount)
# --- theoretical C_ell ---
fftlog = '/Volumes/TimeMachine/data/mocks/fftlog.out'
CFFT = CellTheory(fftlog)
fftlog2 = '/Volumes/TimeMachine/data/mocks/fftloggz0.85redbias2.out'
CFFT2 = CellTheory(fftlog2)
clwind = CFFT.convolve(XIW)
clwind2 = CFFT2.convolve(XIW)
fig, ax = plt.subplots()
for cli in [clwind, clwind2]:
ax.plot(cli['Cellconv']/cli['Cell'], 'r--')
ax.set_xscale('log')
ax.tick_params(direction='in', which='both', axis='both', top=True, right=True)
ax.grid(True, ls=':', color='grey', alpha=0.3)
np.save('/Volumes/TimeMachine/data/DR7/results_referee/cl_window_model.npy',
{'rsd':clwind2, 'norsd':clwind})
# read paircount
paircount = '/Volumes/TimeMachine/data/mocks/mocks_window.cut.256.npy'
XIW = XiWindow(paircount)
# --- theoretical C_ell ---
fftlog = '/Volumes/TimeMachine/data/mocks/fftlog.out'
CFFT = CellTheory(fftlog)
fftlog2 = '/Volumes/TimeMachine/data/mocks/fftloggz0.85redbias2.out'
CFFT2 = CellTheory(fftlog2)
clwind = CFFT.convolve(XIW)
clwind2 = CFFT2.convolve(XIW)
fig, ax = plt.subplots()
for cli in [clwind, clwind2]:
ax.plot(cli['Cellconv']/cli['Cell'], 'r--')
ax.set_xscale('log')
ax.tick_params(direction='in', which='both', axis='both', top=True, right=True)
ax.grid(True, ls=':', color='grey', alpha=0.3)
```
## Step 2: Multiply theory omega by window omega
```
# # fftlog2 = '/Volumes/TimeMachine/data/mocks/fftloggz0.85redbias2.out'
# # ebosscl = '/Volumes/TimeMachine/data/mocks/eBOSSELG.cell.dat'
# cl_fft = CellTheory(fftlog)
# cl_fft2 = CellTheory(fftlog2)
# cl_eboss = CellTheory(ebosscl)
# fig, ax = plt.subplots(figsize=(12, 6))
# for cl_modi in [cl_fft, cl_fft2, cl_eboss]:
# ell = np.arange(cl_modi.cl_mod_ex.size)
# ax.plot(ell, cl_modi.cl_mod_ex)
# ax.set(xlabel=r'$\ell$', ylabel=r'C$_\ell$', xscale='log')
# ax.grid(True, ls=':', alpha=0.5, which='both')
# def plot(cl_mod):
# plt.plot(cl_mod.cl_mod_ex, 'y-',
# cl_mod.cl_mod, 'g--')
# return None
# plt.figure(figsize=(12, 8))
# # fig
# plot(cl_mod)
# plt.xscale('log')
# # sub fig
# plt.axes([0.55, 0.6, 0.3, 0.2])
# plot(cl_mod)
# plt.xlim(8.0e2, 1.2e3)
# plt.ylim(3.0e-8, 8.0e-8)
#--- data
# window xi x theory xi
# #xi_wind.smooth(saveto='/Volumes/TimeMachine/data/DR7/dr7_window_theta_omega.cut.256.txt')
# xi_wind.load(infile='/Volumes/TimeMachine/data/DR7/dr7_window_theta_omega.cut.256.txt', sf=0.6864554309873736)
# xi_wind.plot()
plt.plot(mid_sep_ext, rr_ext, color='grey')
# plt.plot(xi_wind.mid_sep, xi_wind.rr_normed, color='k')
plt.plot(xi_wind.mid_sep, poly(np.array(xi_wind.mid_sep), *params),
label='dof=%d'%dof, color='m', alpha=0.8, ls='--')
# plt.xscale('log')
plt.yscale('log')
plt.legend(bbox_to_anchor=(1., 1.))
plt.xlabel('theta [rad.]')
plt.ylabel('RR')
scalefactor
#
# plt.plot()
fig, ax = plt.subplots(ncols=2, figsize=(12, 4))
ax[0].plot(np.arange(513), cl_fft.cl_mod_ex[:513]/cl_mod_wind,
marker='.', c='k')
ax[0].legend(['Model/(ModelxWindow)'])
# ax[0].legend()
ax[0].set_xscale('log')
ax[0].grid(True, ls=':', color='grey')
ax[0].set_ylim(0.9, 1.1)
ax[0].set_xlim(0.6, 600)
ax[1].plot(cl_fft.cl_mod_ex*1.0e+6, 'grey')
ax[1].plot(cl_mod_wind*1.0e+6, 'r--')
ax[1].set_ylabel('Cellx1.e+6')
ax[1].set_xlim(0.6, 600)
ax[1].legend(['Model', 'ModelxWindow'])
ax[1].set_xscale('log')
#
# plt.plot()
fig, ax = plt.subplots(ncols=2, figsize=(12, 4))
ax[0].plot(np.arange(513), cl_mod.clmod[:513]/cl_mod_wind,
marker='.', c='k')
ax[0].legend(['Model/(ModelxWindow)'])
# ax[0].legend()
ax[0].set_xscale('log')
ax[0].grid(True, ls=':', color='grey')
ax[0].set_ylim(0.9, 1.1)
ax[0].set_xlim(0.6, 600)
ax[1].plot(cl_mod.cl_mod*1.0e+6, 'grey')
ax[1].plot(cl_mod_wind*1.0e+6, 'r--')
ax[1].set_ylabel('Cellx1.e+6')
ax[1].set_xlim(0.6, 600)
ax[1].legend(['Model', 'ModelxWindow'])
ax[1].set_xscale('log')
(cl_mod.clmod[512]/cl_mod_wind[512])
cl
cos_sep = np.cos(cl_mod.sep_rad)
xi_mod = cl_mod.xi_mod
xi_mod_intp = IUS(cos_sep[::-1], xi_mod[::-1])
xg, wg = np.polynomial.legendre.leggauss(20)
xi_mod_wind = xi_mod_intp(xg)*poly(np.arccos(xg), *params)
cl_mod_wind = xi2cl(xg, wg, xi_mod_wind, 2000)
# plot
ell = np.arange(2000)
plt.figure(figsize=(6, 4))
plt.plot(np.arange(cl_mod.cl_mod.size), cl_mod.cl_mod, 'k-',
ell, cl_mod_wind[ell], 'r-')
plt.loglog()
plt.legend(['model', 'model conv. window'], fontsize=12)
plt.xlabel(r'$\ell$')
plt.ylabel(r'C$_{\ell}$')
plt.show()
# ell = np.arange(1001)
# plt.figure(figsize=(6, 4))
# plt.scatter(ell, cl_mod.cl_mod[ell]/cl_mod_wind[ell], color='k', marker='.')
# plt.xlabel(r'$\ell$')
# plt.title(r'$C_{Model}/C_{{\rm Model conv. Window}}$')
# plt.xlim(0.8, 1200)
# # plt.ylim(5.7, 6.3)
# plt.ylim(0.9, 1.1)
# plt.xscale('log')
# plt.show()
#--- mocks
# window xi x theory xi
xi_wind = XiWindow('/Volumes/TimeMachine/data/mocks/mocks_window.cut.256.npy',
totalfrac=86669.4032)
#xi_wind.smooth()
# or
xi_wind.load('/Volumes/TimeMachine/data/mocks/mocks_window_theta_omega.cut.256.txt',
sf=0.3477201700151347)
xi_wind.plot()
def fix(x, y2):
yc = y2.copy()
m = x>100
xmin = x[m].min()
yc[m] *= np.exp(-(x[m]-xmin)**2/900)
return yc
fig, ax = plt.subplots(ncols=2, nrows=2, figsize=(12, 8))
ax = ax.flatten()
fig.subplots_adjust(hspace=0.3, wspace=0.3)
y3 =fix(x, y2)
ax[0].plot(x, y1, 'k-', x, y2, 'r--', x, y3, 'b:')
ax[0].set(xscale='log',ylabel='RR', xlabel = r'$\theta$ [deg]')
ax[1].plot(x, y1, 'k-', x, y2, 'r--', x, y3, 'b:')
ax[1].legend(['RR', 'RR (smoothed)'], fontsize=12, loc=3)
ax[1].set(yscale='log',ylabel='RR', xlabel = r'$\theta$ [deg]')
ax[2].plot(x, y2, 'r--', x, y3, 'b:')
ax[2].set(ylabel='RR', xlabel = r'$\theta$ [deg]')
ax[3].plot(x, y2, 'r--',x, y3, 'b:')
ax[3].set(xlim=(90, 200), ylim=(-0.1, 0.2),ylabel='RR', xlabel = r'$\theta$ [deg]')
plt.show()
plt.plot(x, y2, 'r-',
x, fix(x, y2), 'b--')
sep_rad = np.linspace(0, np.pi, 20*181, endpoint=True)
dsep_rad = np.diff(sep_rad)
cos_sep = np.cos(sep_rad)
xi_mod_intp = IUS(cos_sep[::-1], xi_mod[::-1])
xg, wg = np.polynomial.legendre.leggauss(5000)
xi_mod_wind = xi_mod_intp(xg)*poly(np.arccos(xg), *params)/sf
cl_mod_wind = xi2cl(xg, wg, xi_mod_wind, xg.size)
# plot
ell = np.arange(2000)
plt.figure(figsize=(6, 4))
plt.plot(np.arange(cl_mod.cl_mod.size), cl_mod.cl_mod, 'k-',
ell, cl_mod_wind[ell], 'r-')
plt.loglog()
plt.legend(['model', 'model conv. window'], fontsize=12)
plt.xlabel(r'$\ell$')
plt.ylabel(r'C$_{\ell}$')
plt.show()
ell = np.arange(1001)
plt.figure(figsize=(6, 4))
plt.scatter(ell, cl_mod.cl_mod[ell]/cl_mod_wind[ell], color='k', marker='.')
plt.xlabel(r'$\ell$')
plt.title(r'$C_{Model}/C_{{\rm Model conv. Window}}$')
plt.xlim(0.8, 1200)
# plt.ylim(5.7, 6.3)
plt.ylim(0.9, 1.1)
plt.xscale('log')
plt.show()
np.savetxt('/Volumes/TimeMachine/data/mocks/mocks_ell_cell_theory_window.cut.256.txt',
np.column_stack([ell, cl_mod.cl_mod[ell], cl_mod_wind[ell]]),
header='ell - Cell - Cell window conv.')
xi_mod_intp = IUS(cos_sep[::-1], xi_mod[::-1])
xg, wg = np.polynomial.legendre.leggauss(5000)
xi_mod_wind = xi_mod_intp(xg)*poly(np.arccos(xg), *params)/sf
cl_mod_wind = xi2cl(xg, wg, xi_mod_wind, xg.size)
# plot
ell = np.arange(2000)
plt.figure(figsize=(6, 4))
plt.plot(np.arange(cl_mod.cl_mod.size), cl_mod.cl_mod, 'k-',
ell, cl_mod_wind[ell], 'r-')
plt.loglog()
plt.legend(['model', 'model conv. window'], fontsize=12)
plt.xlabel(r'$\ell$')
plt.ylabel(r'C$_{\ell}$')
plt.show()
ell = np.arange(1001)
plt.figure(figsize=(6, 4))
plt.scatter(ell, cl_mod.cl_mod[ell]/cl_mod_wind[ell], color='k', marker='.')
plt.xlabel(r'$\ell$')
plt.title(r'$C_{Model}/C_{{\rm Model conv. Window}}$')
plt.xlim(0.8, 1200)
# plt.ylim(5.7, 6.3)
plt.ylim(0.9, 1.1)
plt.xscale('log')
plt.show()
np.savetxt('/Volumes/TimeMachine/data/DR7/dr7_ell_cell_theory_window.cut.256.txt',
np.column_stack([ell, cl_mod.cl_mod[ell], cl_mod_wind[ell]]),
header='ell - Cell - Cell window conv.')
cl_obs = np.load('/Volumes/TimeMachine/data/DR7/results/clustering/cl_nn_ab.npy',
allow_pickle=True).item()
frac = hp.read_map('/Volumes/TimeMachine/data/DR7/frac.hp.256.fits', verbose=False)
mask = hp.read_map('/Volumes/TimeMachine/data/DR7/mask.cut.hp.256.fits', verbose=False) > 0
frac[mask].sum()
mask1024 = hp.ud_grade(mask, nside_out=1024)
frac1024 = hp.ud_grade(frac, nside_out=1024)
hpfrac = hp.ma(frac1024)
hpfrac.mask=np.logical_not(mask1024>0)
hp.mollview(hpfrac.filled(), rot=-89, title='DR7')
cl_win = hp.anafast(hpfrac.filled(), lmax=1024)
plt.loglog(cl_win)
xi_win = cl2xi(cl_win, cos_sep)
plt.plot(np.degrees(sep_rad), xi_win/0.23623275756835938)
plt.xscale('log')
plt.ylim(0, 1)
cl_obs_wind,_ = apply_window_v0(cl_obs['auto']*0.23429479830970087, sep_rad[::-1], xi_win[::-1]/0.23429479830970087, theory=False)
plt.plot(cl_obs['auto'], 'k-',
cl_obs_wind, 'r--')
# plt.loglog()
# plt.ylim(1.e-9, 1.e-5)
mask.mean()
## uses Ashley's approach,
## does not allow multiple zero crossings
# import sys
# sys.path.append('/Users/rezaie/github/LSSanalysis')
# import legendre
# from time import time
# Leg = np.vectorize(legendre.legendre)
# x = np.linspace(-1.0, 1., 100, endpoint=True)
# c = np.zeros(5001)
# for j in [0, 1, 2]:
# t1 = time()
# y1 = scipyLeg(j, x)
# t2 = time()
# y2= Leg(j, x)
# t3 = time()
# plt.plot(x, y1, color='b', ls='-', alpha=0.5)
# plt.plot(x, y2, color='r', ls=':')
# print('scipy', t2-t1)
# print('ashley', t3-t2)
# t = np.linspace(0., 180., num=4*181, endpoint=True)
# cost = np.cos(np.deg2rad(t))
# Omega = np.zeros(cost.size)
# elmax = 10000
# clmod = cl_mod.run(np.arange(elmax+10))
# fig, ax = plt.subplots(nrows=3, figsize=(10, 6), sharey=True,
# sharex=True)
# ls = ['-', '-.', '--']
# j = 0
# #for cr_limit in [500]:
# # print(cr_limit)
# cr_limit=1.0e24
# for elmax_l in [10000]:#, 2000, 5000]:
# for i,cost_i in enumerate(cost):
# Omega[i] = 0.0
# el = 1
# domega = 0.0
# omega = 0.0
# cr = 0
# oldpl = 1
# while (cr <= cr_limit) & (el < elmax_l):
# pl = scipyLeg(el, cost_i)
# domega = (2*el+1)*pl*clmod[el]
# #print(pl, oldpl)
# #if (pl < 0) and (oldpl > 0):
# # cr += 1
# el += 1
# omega += domega
# oldpl = pl
# if i <3:print('i:{}, el:{}'.format(i, el))
# Omega[i] = omega
# Omega *= (0.25/np.pi)
# ax[j].plot(t, Omega/1.0e-5, ls=ls[j],
# label='Lmax=%d'%elmax_l, color='k')
# #label='# of zero crossing = %d'%cr_limit)
# ax[j].text(0.7, 0.1, 'Lmax=%d'%elmax_l, color='k', transform=ax[j].transAxes)
# ax[j].axhline(linestyle=':', color='grey')
# j += 1
# print('Sum Omega = {}'.format(sum(Omega*t)))
# # plt.ylim(-4.e-3, 6.e-3)
# ax[0].set(ylim=(-2.5, 1.2), title='Theoretical $\omega$')
# ax[2].set(xlabel=r'$\theta$ [deg]')
# ax[1].set(ylabel=r'$\omega$ [1e-5]')
cl_obs = np.load('/Volumes/TimeMachine/data/DR7/results/clustering/cl_nn_ab.npy',
allow_pickle=True).item()
cl_obs.keys()
cl_obs['sf']
cl_wobs,_ = apply_window_v0(cl_obs['auto']*0.23429479830970087, xi_th.sep, xi_th.xiw, theory=False)
cl_wmod,_ = apply_window_v0(cl_mod.cl_mod, xi_th.sep, xi_th.xiw, theory=True)
plt.plot(np.arange(cl_obs['auto'].size), cl_obs['auto'], marker='+', color='r', label='Observed', ls='None')
plt.plot(np.arange(cl_obs['auto'].size), cl_wobs*200, marker='x', color='b', label='Observed w window', ls='None')
plt.plot(np.arange(cl_mod.cl_mod.size), 4*cl_mod.cl_mod + 8.536537713893294e-07, 'k-', label='Model')
plt.plot(np.arange(cl_wmod.size), 4*cl_wmod/720 + 8.536537713893294e-07, 'k:', label='Model w window')
plt.ylim(1.e-6, 1.e-4)
plt.legend(bbox_to_anchor=(1., 1))
plt.title('DR7')
plt.xlabel(r'$\ell$')
plt.ylabel(r'C$_{\ell}$')
plt.loglog()
#plt.xlim(0.9, 1000)
# plt.yscale('log')
```
| github_jupyter |
## The Grid layout
The `GridBox` class is a special case of the `Box` widget.
The `Box` widget enables the entire CSS flexbox spec, enabling rich reactive layouts in the Jupyter notebook. It aims at providing an efficient way to lay out, align and distribute space among items in a container.
Again, the whole grid layout spec is exposed via the `layout` attribute of the container widget (`Box`) and the contained items. One may share the same `layout` attribute among all the contained items.
The following flexbox tutorial on the flexbox layout follows the lines of the article [A Complete Guide to Grid](https://css-tricks.com/snippets/css/complete-guide-grid/) by Chris House, and uses text and various images from the article [with permission](https://css-tricks.com/license/).
### Basics and browser support
To get started you have to define a container element as a grid with display: grid, set the column and row sizes with grid-template-rows, grid-template-columns, and grid_template_areas, and then place its child elements into the grid with grid-column and grid-row. Similarly to flexbox, the source order of the grid items doesn't matter. Your CSS can place them in any order, which makes it super easy to rearrange your grid with media queries. Imagine defining the layout of your entire page, and then completely rearranging it to accommodate a different screen width all with only a couple lines of CSS. Grid is one of the most powerful CSS modules ever introduced.
As of March 2017, most browsers shipped native, unprefixed support for CSS Grid: Chrome (including on Android), Firefox, Safari (including on iOS), and Opera. Internet Explorer 10 and 11 on the other hand support it, but it's an old implementation with an outdated syntax. The time to build with grid is now!
### Important terminology
Before diving into the concepts of Grid it's important to understand the terminology. Since the terms involved here are all kinda conceptually similar, it's easy to confuse them with one another if you don't first memorize their meanings defined by the Grid specification. But don't worry, there aren't many of them.
**Grid Container**
The element on which `display: grid` is applied. It's the direct parent of all the grid items. In this example container is the grid container.
```html
<div class="container">
<div class="item item-1"></div>
<div class="item item-2"></div>
<div class="item item-3"></div>
</div>
```
**Grid Item**
The children (e.g. direct descendants) of the grid container. Here the item elements are grid items, but sub-item isn't.
```html
<div class="container">
<div class="item"></div>
<div class="item">
<p class="sub-item"></p>
</div>
<div class="item"></div>
</div>
```
**Grid Line**
The dividing lines that make up the structure of the grid. They can be either vertical ("column grid lines") or horizontal ("row grid lines") and reside on either side of a row or column. Here the yellow line is an example of a column grid line.

**Grid Track**
The space between two adjacent grid lines. You can think of them like the columns or rows of the grid. Here's the grid track between the second and third row grid lines.

**Grid Cell**
The space between two adjacent row and two adjacent column grid lines. It's a single "unit" of the grid. Here's the grid cell between row grid lines 1 and 2, and column grid lines 2 and 3.

**Grid Area**
The total space surrounded by four grid lines. A grid area may be comprised of any number of grid cells. Here's the grid area between row grid lines 1 and 3, and column grid lines 1 and 3.

### Properties of the parent
**grid-template-rows, grid-template-colums**
Defines the columns and rows of the grid with a space-separated list of values. The values represent the track size, and the space between them represents the grid line.
Values:
- `<track-size>` - can be a length, a percentage, or a fraction of the free space in the grid (using the `fr` unit)
- `<line-name>` - an arbitrary name of your choosing
**grid-template-areas**
Defines a grid template by referencing the names of the grid areas which are specified with the grid-area property. Repeating the name of a grid area causes the content to span those cells. A period signifies an empty cell. The syntax itself provides a visualization of the structure of the grid.
Values:
- `<grid-area-name>` - the name of a grid area specified with `grid-area`
- `.` - a period signifies an empty grid cell
- `none` - no grid areas are defined
**grid-gap**
A shorthand for `grid-row-gap` and `grid-column-gap`
Values:
- `<grid-row-gap>`, `<grid-column-gap>` - length values
where `grid-row-gap` and `grid-column-gap` specify the sizes of the grid lines. You can think of it like setting the width of the gutters between the columns / rows.
- `<line-size>` - a length value
*Note: The `grid-` prefix will be removed and `grid-gap` renamed to `gap`. The unprefixed property is already supported in Chrome 68+, Safari 11.2 Release 50+ and Opera 54+.*
**align-items**
Aligns grid items along the block (column) axis (as opposed to justify-items which aligns along the inline (row) axis). This value applies to all grid items inside the container.
Values:
- `start` - aligns items to be flush with the start edge of their cell
- `end` - aligns items to be flush with the end edge of their cell
- `center` - aligns items in the center of their cell
- `stretch` - fills the whole height of the cell (this is the default)
**justify-items**
Aligns grid items along the inline (row) axis (as opposed to `align-items` which aligns along the block (column) axis). This value applies to all grid items inside the container.
Values:
- `start` - aligns items to be flush with the start edge of their cell
- `end` - aligns items to be flush with the end edge of their cell
- `center` - aligns items in the center of their cell
- `stretch` - fills the whole width of the cell (this is the default)
**align-content**
Sometimes the total size of your grid might be less than the size of its grid container. This could happen if all of your grid items are sized with non-flexible units like `px`. In this case you can set the alignment of the grid within the grid container. This property aligns the grid along the block (column) axis (as opposed to justify-content which aligns the grid along the inline (row) axis).
Values:
- `start` - aligns the grid to be flush with the start edge of the grid container
- `end` - aligns the grid to be flush with the end edge of the grid container
- `center` - aligns the grid in the center of the grid container
- `stretch` - resizes the grid items to allow the grid to fill the full height of the grid container
- `space-around` - places an even amount of space between each grid item, with half-sized spaces on the far ends
- `space-between` - places an even amount of space between each grid item, with no space at the far ends
- `space-evenly` - places an even amount of space between each grid item, including the far ends
**justify-content**
Sometimes the total size of your grid might be less than the size of its grid container. This could happen if all of your grid items are sized with non-flexible units like `px`. In this case you can set the alignment of the grid within the grid container. This property aligns the grid along the inline (row) axis (as opposed to align-content which aligns the grid along the block (column) axis).
Values:
- `start` - aligns the grid to be flush with the start edge of the grid container
- `end` - aligns the grid to be flush with the end edge of the grid container
- `center` - aligns the grid in the center of the grid container
- `stretch` - resizes the grid items to allow the grid to fill the full width of the grid container
- `space-around` - places an even amount of space between each grid item, with half-sized spaces on the far ends
- `space-between` - places an even amount of space between each grid item, with no space at the far ends
- `space-evenly` - places an even amount of space between each grid item, including the far ends
**grid-auto-columns, grid-auto-rows**
Specifies the size of any auto-generated grid tracks (aka implicit grid tracks). Implicit tracks get created when there are more grid items than cells in the grid or when a grid item is placed outside of the explicit grid. (see The Difference Between Explicit and Implicit Grids)
Values:
- `<track-size>` - can be a length, a percentage, or a fraction of the free space in the grid (using the `fr` unit)
### Properties of the items
*Note: `float`, `display: inline-block`, `display: table-cell`, `vertical-align` and `column-??` properties have no effect on a grid item.*
**grid-column, grid-row**
Determines a grid item's location within the grid by referring to specific grid lines. `grid-column-start`/`grid-row-start` is the line where the item begins, and `grid-column-end`/`grid-row-end` is the line where the item ends.
Values:
- `<line>` - can be a number to refer to a numbered grid line, or a name to refer to a named grid line
- `span <number>` - the item will span across the provided number of grid tracks
- `span <name>` - the item will span across until it hits the next line with the provided name
- `auto` - indicates auto-placement, an automatic span, or a default span of one
```css
.item {
grid-column: <number> | <name> | span <number> | span <name> | auto /
<number> | <name> | span <number> | span <name> | auto
grid-row: <number> | <name> | span <number> | span <name> | auto /
<number> | <name> | span <number> | span <name> | auto
}
```
Examples:
```css
.item-a {
grid-column: 2 / five;
grid-row: row1-start / 3;
}
```

```css
.item-b {
grid-column: 1 / span col4-start;
grid-row: 2 / span 2;
}
```

If no `grid-column` / `grid-row` is declared, the item will span 1 track by default.
Items can overlap each other. You can use `z-index` to control their stacking order.
**grid-area**
Gives an item a name so that it can be referenced by a template created with the `grid-template-areas` property. Alternatively, this property can be used as an even shorter shorthand for `grid-row-start` + `grid-column-start` + `grid-row-end` + `grid-column-end`.
Values:
- `<name>` - a name of your choosing
- `<row-start> / <column-start> / <row-end> / <column-end>` - can be numbers or named lines
```css
.item {
grid-area: <name> | <row-start> / <column-start> / <row-end> / <column-end>;
}
```
Examples:
As a way to assign a name to the item:
```css
.item-d {
grid-area: header
}
```
As the short-shorthand for `grid-row-start` + `grid-column-start` + `grid-row-end` + `grid-column-end`:
```css
.item-d {
grid-area: 1 / col4-start / last-line / 6
}
```

**justify-self**
Aligns a grid item inside a cell along the inline (row) axis (as opposed to `align-self` which aligns along the block (column) axis). This value applies to a grid item inside a single cell.
Values:
- `start` - aligns the grid item to be flush with the start edge of the cell
- `end` - aligns the grid item to be flush with the end edge of the cell
- `center` - aligns the grid item in the center of the cell
- `stretch` - fills the whole width of the cell (this is the default)
```css
.item {
justify-self: start | end | center | stretch;
}
```
Examples:
```css
.item-a {
justify-self: start;
}
```

```css
.item-a {
justify-self: end;
}
```

```css
.item-a {
justify-self: center;
}
```

```css
.item-a {
justify-self: stretch;
}
```

To set alignment for *all* the items in a grid, this behavior can also be set on the grid container via the `justify-items` property.
```
from ipywidgets import Button, GridBox, Layout, ButtonStyle
```
Placing items by name:
```
header = Button(description='Header',
layout=Layout(width='auto', grid_area='header'),
style=ButtonStyle(button_color='lightblue'))
main = Button(description='Main',
layout=Layout(width='auto', grid_area='main'),
style=ButtonStyle(button_color='moccasin'))
sidebar = Button(description='Sidebar',
layout=Layout(width='auto', grid_area='sidebar'),
style=ButtonStyle(button_color='salmon'))
footer = Button(description='Footer',
layout=Layout(width='auto', grid_area='footer'),
style=ButtonStyle(button_color='olive'))
GridBox(children=[header, main, sidebar, footer],
layout=Layout(
width='50%',
grid_template_rows='auto auto auto',
grid_template_columns='25% 25% 25% 25%',
grid_template_areas='''
"header header header header"
"main main . sidebar "
"footer footer footer footer"
''')
)
```
Setting up row and column template and gap
```
GridBox(children=[Button(layout=Layout(width='auto', height='auto'),
style=ButtonStyle(button_color='darkseagreen')) for i in range(9)
],
layout=Layout(
width='50%',
grid_template_columns='100px 50px 100px',
grid_template_rows='80px auto 80px',
grid_gap='5px 10px')
)
```
| github_jupyter |
# Iterative Quantum Phase Estimation Algorithm
The goal of this tutorial is to understand how the Iterative Phase Estimation (IPE) algorithm works, why would we use the IPE algorithm instead of the QPE (Quantum Phase Estimation) algorithm and how to build it with Qiskit using the same circuit exploiting reset gate and the `c_if` method that allows to apply gates conditioned by the values stored in a classical register, resulting from previous measurements.
**References**
- [Section 2 of Lab 4: Iterative Phase Estimation (IPE) Algorithm](https://qiskit.org/textbook/ch-labs/Lab04_IterativePhaseEstimation.html#2-iterative-phase-estimation-ipe-algorithm)
- [Ch.3.6 Quantum Phase Estimation](https://qiskit.org/textbook/ch-algorithms/quantum-phase-estimation.html)
```
from qiskit import QuantumCircuit, ClassicalRegister, QuantumRegister, transpile, Aer
from qiskit.tools.visualization import plot_histogram
from math import pi
import matplotlib.pyplot as plt
```
# Conditioned gates: the c_if method
Before starting the IPE algorithm, we will give a brief tutorial about the Qiskit conditional method, c_if, as it goes into building the IPE circuit.
`c_if` is a function (actually a method of the gate class) to perform conditioned operations based on the value stored previously in a classical register. With this feature you can apply gates after a measurement in the same circuit conditioned by the measurement outcome.
For example, the following code will execute the $X$ gate if the value of the classical register is $0$.
```
q = QuantumRegister(1,'q')
c = ClassicalRegister(1,'c')
qc = QuantumCircuit(q, c)
qc.h(0)
qc.measure(0,0)
qc.x(0).c_if(c, 0)
qc.draw(output='mpl')
```
We highlight that the method c_if expects as the first argument a whole classical register, not a single classical bit (or a list of classical bits), and as the second argument a value in decimal representation (a non-negative integer), not the value of a single bit, 0, or 1 (or a list/string of binary digits).
Let's make another example. Consider that we want to perform a bit flip on the third qubit after the measurements in the following circuit, when the results of the measurement of $q_0$ and $q_1$ are both $1$.
```
q = QuantumRegister(3,'q')
c = ClassicalRegister(3,'c')
qc = QuantumCircuit(q, c)
qc.h(q[0])
qc.h(q[1])
qc.h(q[2])
qc.barrier()
qc.measure(q,c)
qc.draw('mpl')
```
We want to apply the $X$ gate, only if both the results of the measurement of $q_0$ and $q_1$ are $1$. We can do this using the c_if method, conditioning the application of $X$ depending on the value passed as argument to c_if.
We will have to encode the value to pass to the c_if method such that it will check the values 011 and 111 (in binary representation), since it does not matter what $q_2$ is measured as.
The 2 integer values in decimal representation:
<img src="../../images/binary.png" width="50%" >
We can check the solutions using the bin() method in python (the prefix `0b` indicates the binary format).
```
print(bin(3))
print(bin(7))
```
So we have to apply $X$ to $q_2$ using c_if two times, one for each value corresponding to 011 and 111.
```
q = QuantumRegister(3,'q')
c = ClassicalRegister(3,'c')
qc = QuantumCircuit(q, c)
qc.h(0)
qc.h(1)
qc.h(2)
qc.barrier()
qc.measure(q,c)
qc.x(2).c_if(c, 3) # for the 011 case
qc.x(2).c_if(c, 7) # for the 111 case
qc.draw(output='mpl')
```
# IPE
The motivation for using the IPE algorithm is that QPE algorithm works fine for short depth circuits but when the circuit starts to grow, it doesn't work properly due to gate noise and decoherence times.
The detailed explanation of how the algorithm works can be found in [Iterative Phase Estimation (IPE) Algorithm](https://qiskit.org/textbook/ch-labs/Lab04_IterativePhaseEstimation.html#2-iterative-phase-estimation-ipe-algorithm). To understand QPE in depth, you can see also [Ch.3.6 Quantum Phase Estimation](https://qiskit.org/textbook/ch-algorithms/quantum-phase-estimation.html).
## IPE example with a 1-qubit gate for $U$
We want to apply the IPE algorithm to estimate the phase for a 1-qubit operator $U$. For example, here we use the $S$-gate.
Let's apply the IPE algorithm to estimate the phase for $S$-gate.
Its matrix is
$$ S =
\begin{bmatrix}
1 & 0\\
0 & e^\frac{i\pi}{2}\\
\end{bmatrix}$$
That is, the $S$-gate adds a phase $\pi/2$ to the state $|1\rangle$, leaving unchanged the phase of the state $|0\rangle$
$$ S|1\rangle = e^\frac{i\pi}{2}|1\rangle $$
In the following, we will use the notation and terms used in [Section 2 of lab 4](https://qiskit.org/textbook/ch-labs/Lab04_IterativePhaseEstimation.html#2-iterative-phase-estimation-ipe-algorithm).
Let's consider to estimate the phase $\phi=\frac{\pi}{2}$ for the eigenstate $|1\rangle$, we should find $\varphi=\frac{1}{4}$ (where $\phi = 2 \pi \varphi$). Therefore to estimate the phase we need exactly 2 phase bits, i.e. $m=2$, since $1/2^2=1/4$. So $\varphi=0.\varphi_1\varphi_2$.
Remember from the theory that for the IPE algorithm, $m$ is also the number of iterations, so we need only $2$ iterations or steps.
First, we initialize the circuit. IPE works with only 1 auxiliary qubit, instead of $m$ counting qubits of the QPE algorithm. Therefore, we need 2 qubits, 1 auxiliary qubit and 1 for the eigenstate of $U$-gate, and a classical register of 2 bits, for the phase bits $\varphi_1$, $\varphi_2$.
```
nq = 2
m = 2
q = QuantumRegister(nq,'q')
c = ClassicalRegister(m,'c')
qc_S = QuantumCircuit(q,c)
```
### First step
Now we build the quantum circuit for the first step, that is, the first iteration of the algorithm, to estimate the least significant phase bit $\varphi_m$, in this case $\varphi_2$. For the first step we have 3 sub-steps:
- initialization
- application of the Controlled-$U$ gates
- measure of the auxiliary qubit in x-basis
#### Initialization
The initialization consists of application the Hadamard gate to the auxiliary qubit and the preparation of the eigenstate $|1\rangle$.
```
qc_S.h(0)
qc_S.x(1)
qc_S.draw('mpl')
```
#### Application of the Controlled-$U$ gates
Then we have to apply $2^t$ times the Controlled-$U$ operators (see also in the docs [Two qubit gates](https://qiskit.org/documentation/tutorials/circuits/3_summary_of_quantum_operations.html#Two-qubit-gates)), that, in this example, is the Controlled-$S$ gate ($CS$ for short).
To implement $CS$ in the circuit, since $S$ is a phase gate, we can use the controlled phase gate $\text{CP}(\theta)$, with $\theta=\pi/2$.
```
cu_circ = QuantumCircuit(2)
cu_circ.cp(pi/2,0,1)
cu_circ.draw('mpl')
```
Let's apply $2^t$ times $\text{CP}(\pi/2)$. Since for the first step $t=m-1$, and $m=2$, we have $2^t=2$.
```
for _ in range(2**(m-1)):
qc_S.cp(pi/2,0,1)
qc_S.draw('mpl')
```
#### Measure in x-basis
Finally, we perform the measurement of the auxiliary qubit in x-basis. So we will define a function to perform the x_measure and then apply it.
```
def x_measurement(qc, qubit, cbit):
"""Measure 'qubit' in the X-basis, and store the result in 'cbit'"""
qc.h(qubit)
qc.measure(qubit, cbit)
```
In this way we obtain the phase bit $\varphi_2$ and store it in the classical bit $c_0$.
```
x_measurement(qc_S, q[0], c[0])
qc_S.draw('mpl')
```
### Subsequent steps (2nd step)
Now we build the quantum circuit for the other remaining steps, in this example, only the second one.
In these steps we have 4 sub-steps: the 3 sub-steps as in the first step and, in the middle, the additional step of the phase correction
- initialization with reset
- phase correction
- application of the Control-$U$ gates
- measure of the auxiliary qubit in x-basis
#### Initialization with reset
As we want to perform an iterative algorithm in the same circuit, we need to reset the auxiliary qubit $q0$ after the measument gate and initialize it again as before to recycle the qubit.
```
qc_S.reset(0)
qc_S.h(0)
qc_S.draw('mpl')
```
#### Phase correction (for step 2)
As seen in the theory, in order to extract the phase bit $\varphi_{1}$, we perform a phase correction of $-\pi\varphi_2/2$.
Of course, we need to apply the phase correction in the circuit only if the phase bit $\varphi_2=1$, i.e. we have to apply the phase correction of $-\pi/2$ only if the classical bit $c_0$ is 1.
So, after the reset we apply the phase gate $P(\theta)$ with phase $\theta=-\pi/2$ conditioned by the classical bit $c_0$ ($=\varphi_2$) using the `c_if` method.
So as we saw in the first part of this tutorial, we have to use the `c_if` method with a value of 1, as $1_{10} = 001_{2}$ (the subscripts $_{10}$ and $_2$ indicate the decimal and binary representations).
```
qc_S.p(-pi/2,0).c_if(c,1)
qc_S.draw('mpl')
```
#### Application of the Control-$U$ gates and x-measurement (for step 2)
We apply the $CU$ operations as we did in the first step. For the second step we have $t=m-2$, hence $2^t=1$. So we apply $\text{CP}(\pi/2)$ once. And then we perform the x-measurment of the qubit $q_0$, storing the result, the phase bit $\varphi_1$, in the bit $c_1$ of classical register.
```
## 2^t c-U operations (with t=m-2)
for _ in range(2**(m-2)):
qc_S.cp(pi/2,0,1)
x_measurement(qc_S, q[0], c[1])
```
Et voilà, we have our final circuit
```
qc_S.draw('mpl')
```
Let's execute the circuit with the `qasm_simulator`, the simulator without noise that run locally.
```
sim = Aer.get_backend('qasm_simulator')
count0 = sim.run(transpile(qc_S, sim)).result().get_counts()
key_new = [str(int(key,2)/2**m) for key in list(count0.keys())]
count1 = dict(zip(key_new, count0.values()))
fig, ax = plt.subplots(1,2)
plot_histogram(count0, ax=ax[0])
plot_histogram(count1, ax=ax[1])
plt.tight_layout()
```
In the picture we have the same histograms but on the left we have on the x-axis the string with phase bits $\varphi_1$, $\varphi_2$ and on the right the actual phase $\varphi$ in decimal representation.
As we expected we have found $\varphi=\frac{1}{4}=0.25$ with a $100\%$ probability.
## IPE example with a 2-qubit gate
Now, we want to apply the IPE algorithm to estimate the phase for a 2-qubit gate $U$. For this example, let's consider the controlled version of the $T$ gate, i.e. the gate $U=\textrm{Controlled-}T$ (that from now we will express more compactly with $CT$). Its matrix is
$$ CT =
\begin{bmatrix}
1 & 0 & 0 & 0\\
0 & 1 & 0 & 0\\
0 & 0 & 1 & 0\\
0 & 0 & 0 & e^\frac{i\pi}{4}\\
\end{bmatrix} $$
That is, the $CT$ gate adds a phase $\pi/4$ to the state $|11\rangle$, leaving unchanged the phase of the other computational basis states $|00\rangle$, $|01\rangle$, $|10\rangle$.
Let's consider to estimate the phase $\phi=\pi/4$ for the eigenstate $|11\rangle$, we should find $\varphi=1/8$, since $\phi = 2 \pi \varphi$. Therefore to estimate the phase we need exactly 3 classical bits, i.e. $m=3$, since $1/2^3=1/8$. So $\varphi=0.\varphi_1\varphi_2\varphi_3$.
As done with the example for the 1-qubit $U$ operator we will go through the same steps but this time we will have $3$ steps since $m=3$, and we will not repeat all the explanations. So for details see the above example for 1-qubit $U$ gate.
First, we initialize the circuit with 3 qubits, 1 for the auxiliary qubit and 2 for the 2-qubit gate, and 3 classical bits to store the phase bits $\varphi_1$, $\varphi_2$, $\varphi_3$.
```
nq = 3 # number of qubits
m = 3 # number of classical bits
q = QuantumRegister(nq,'q')
c = ClassicalRegister(m,'c')
qc = QuantumCircuit(q,c)
```
### First step
Now we build the quantum circuit for the first step, to estimate the least significant phase bit $\varphi_m=\varphi_3$.
#### Initialization
We initialize the auxiliary qubit and the other qubits with the eigenstate $|11\rangle$.
```
qc.h(0)
qc.x([1,2])
qc.draw('mpl')
```
#### Application of the Controlled-$U$ gates
Then we have to apply multiple times the $CU$ operator, that, in this example, is the Controlled-$CT$ gate ($CCT$ for short).
To implement $CCT$ in the circuit, since $T$ is a phase gate, we can use the multi-controlled phase gate $\text{MCP}(\theta)$, with $\theta=\pi/4$.
```
cu_circ = QuantumCircuit(nq)
cu_circ.mcp(pi/4,[0,1],2)
cu_circ.draw('mpl')
```
Let's apply $2^t$ times $\text{MCP}(\pi/4)$. Since for the first step $t=m-1$ and $m=3$, we have $2^t=4$.
```
for _ in range(2**(m-1)):
qc.mcp(pi/4,[0,1],2)
qc.draw('mpl')
```
#### Measure in x-basis
Finally, we perform the measurenment of the auxiliary qubit in x-basis.
We can use the `x_measurement` function defined above in the example for 1-qubit gate. In this way we have obtained the phase bit $\varphi_3$ and stored it in the classical bit $c_0$.
```
x_measurement(qc, q[0], c[0])
qc.draw('mpl')
```
### Subsequent steps (2nd, 3rd)
Now we build the quantum circuit for the other remaining steps, the second and the third ones.
As said in the first example, in these steps we have the additional sub-step of the phase correction.
#### Initialization with reset
```
qc.reset(0)
qc.h(0)
qc.draw('mpl')
```
#### Phase correction (for step 2)
In order to extract the phase bit $\varphi_{2}$, we perform a phase correction of $-\pi\varphi_3/2$.
So, after the reset we apply the phase gate $P(\theta)$ with phase $\theta=-\pi/2$ conditioned by the classical bit $c_0$ ($=\varphi_3$).
```
qc.p(-pi/2,0).c_if(c,1)
qc.draw('mpl')
```
#### Application of the Control-$U$ gates and x-measurement (for step 2)
We apply the $CU$ operations as we did in the first step. For the second step we have $t=m-2$, hence $2^t=2$. So we apply $\text{MCP}(\pi/4)$ $2$ times. And then we perform the x-measurement of the qubit $q_0$, storing the phase bit $\varphi_2$ in the bit $c_1$.
```
for _ in range(2**(m-2)):
qc.mcp(pi/4,[0,1],2)
x_measurement(qc, q[0], c[1])
qc.draw('mpl')
```
#### All substeps of the 3rd step
For the 3rd and last step, we perform the reset and initialization of the auxiliary qubit as done in the second step.
Then at the 3rd step we have to perform the phase correction of $-2\pi 0.0\varphi_{2}\varphi_{3}= -2\pi \left(\frac{\varphi_2}{4}+\frac{\varphi_3}{8}\right)=-\frac{\varphi_2\pi}{2}-\frac{ \varphi_3\pi}{4}$, thus we have to apply 2 conditioned phase corrections, one conditioned by $\varphi_3$ ($=c_0$) and the other by $\varphi_2$($=c_1$). To do this we have to apply the following:
- gate $P(-\pi/4)$ conditioned by $c_0=1$, that is, by $c=001$ (c_if with value $1$)
- gate $P(-\pi/2)$ conditioned by $c_1=1$, that is, the gate is applied when $c=010$ (c_if with values $2$)
- gate $P(-3\pi/4)$ conditioned by $c_1=1$ and $c_0=1$ that is, the gate is applied when $c=011$ (c_if with values $3$)
Next, the $CU$ operations: we apply $2^t$ times the $\text{MCP}(\pi/4)$ gate and since at the 3rd step $t=m-3=0$, we apply the gate only once.
```
# initialization of qubit q0
qc.reset(0)
qc.h(0)
# phase correction
qc.p(-pi/4,0).c_if(c,1)
qc.p(-pi/2,0).c_if(c,2)
qc.p(-3*pi/2,0).c_if(c,3)
# c-U operations
for _ in range(2**(m-3)):
qc.mcp(pi/4,[0,1],2)
# X measurement
qc.h(0)
qc.measure(0,2)
qc.draw('mpl')
```
Now, we execute the circuit with the simulator without noise.
```
count0 = sim.run(transpile(qc, sim)).result().get_counts()
key_new = [str(int(key,2)/2**m) for key in list(count0.keys())]
count1 = dict(zip(key_new, count0.values()))
fig, ax = plt.subplots(1,2)
plot_histogram(count0, ax=ax[0])
plot_histogram(count1, ax=ax[1])
plt.tight_layout()
```
We have obtained $100\%$ probability to find $\varphi=0.125$, that is, $1/8$, as expected.
```
import qiskit.tools.jupyter
%qiskit_version_table
%qiskit_copyright
```
| github_jupyter |
### Today:
* Supervised Learning
* Linear Regression
* Model
* Cost Function
* Optimization
* Gradient Descent
* Feature Scaling
* Polynomial Regression
* Model
* Overfitting and Underfitting
* Regularization
### Resources:
* Supervised Learning: https://mcerovic.github.io/notes/SupervisedLearning/index.html
* Linear Regression: https://mcerovic.github.io/notes/LinearRegression/index.html
* Gradient Descent: https://mcerovic.github.io/notes/GradientDescent/index.html
* Feature Scaling: http://sebastianraschka.com/Articles/2014_about_feature_scaling.html#about-standardization
# Linear regression
```
# Import necessary libraries
import numpy as np
import tensorflow as tf
import matplotlib.pyplot as plt
%matplotlib inline
# Load dataset as numpy array
y, x = np.loadtxt('../../data/02_LinearRegression/house_price.csv', delimiter=',', unpack=True)
n_samples = len(x)
# Normalize data
x = (x - np.mean(x)) / np.std(x)
y = (y - np.mean(y)) / np.std(y)
# Graphical preview
fig, ax = plt.subplots()
ax.set_xlabel('Size')
ax.set_ylabel('Price')
ax.scatter(x, y, edgecolors='k', label='Real house price')
ax.grid(True, color='gray', linestyle='dashed')
```
## Model
```
X = tf.placeholder(tf.float32, name='X')
Y = tf.placeholder(tf.float32, name='Y')
w = tf.Variable(0.0, name='weights')
b = tf.Variable(0.0, name='bias')
Y_predicted = tf.add(tf.multiply(X, w), b)
```
## Cost function
```
cost = tf.reduce_mean(tf.square(Y - Y_predicted), name='cost')
```
## Optimization
```
optimizer = tf.train.GradientDescentOptimizer(learning_rate=0.001).minimize(cost)
```
## Train
```
with tf.Session() as sess:
# Initialize the necessary variables, in this case, w and b
sess.run(tf.global_variables_initializer())
# writer = tf.summary.FileWriter('./graphs/linear_reg', sess.graph)
# Train the model in 50 epochs
for i in range(50):
total_cost = 0
# Session runs train_op and fetch values of loss
for sample in range(n_samples):
# Session looks at all trainable variables that loss depends on and update them
_, l = sess.run([optimizer, cost], feed_dict={X: x[sample], Y:y[sample]})
total_cost += l
# Print epoch and loss
print('Epoch {0}: {1}'.format(i, total_cost / n_samples))
# Close the writer when you're done using it
# writer.close()
# Output the values of w and b
w, b = sess.run([w, b])
print('W: %f, b: %f' % (w, b))
print('Cost: %f' % sess.run(cost, feed_dict={X: x, Y: y}))
# Append hypothesis that we found on the plot
ax.plot(x, x * w + b, color='r', label='Predicted house price')
ax.legend()
fig
# Predict at point 0.5
print(0.5 * w + b)
```
| github_jupyter |
# Mask R-CNN - Train on Shapes Dataset
### Notes from implementation
This notebook shows how to train Mask R-CNN on your own dataset. To keep things simple we use a synthetic dataset of shapes (squares, triangles, and circles) which enables fast training. You'd still need a GPU, though, because the network backbone is a Resnet101, which would be too slow to train on a CPU. On a GPU, you can start to get okay-ish results in a few minutes, and good results in less than an hour.
The code of the *Shapes* dataset is included below. It generates images on the fly, so it doesn't require downloading any data. And it can generate images of any size, so we pick a small image size to train faster.
```
%matplotlib inline
%load_ext autoreload
%autoreload 2
import os
import sys
import random
import math
import re
import time
import numpy as np
import cv2
import matplotlib
import matplotlib.pyplot as plt
sys.path.append('../')
from mrcnn.config import Config
import mrcnn.model as modellib
import mrcnn.visualize as visualize
from mrcnn.model import log
import mrcnn.shapes as shapes
from mrcnn.dataset import Dataset
# Root directory of the project
ROOT_DIR = os.getcwd()
MODEL_PATH = 'E:\Models'
# Directory to save logs and trained model
MODEL_DIR = os.path.join(MODEL_PATH, "mrcnn_logs")
# Path to COCO trained weights
COCO_MODEL_PATH = os.path.join(MODEL_PATH, "mask_rcnn_coco.h5")
RESNET_MODEL_PATH = os.path.join(MODEL_PATH, "resnet50_weights_tf_dim_ordering_tf_kernels_notop.h5")
import tensorflow as tf
import keras
import pprint
import keras.backend as KB
print("Tensorflow Version: {} Keras Version : {} ".format(tf.__version__,keras.__version__))
pp = pprint.PrettyPrinter(indent=2, width=100)
config = shapes.ShapesConfig()
config.BATCH_SIZE = 2 #Batch size is 2 (# GPUs * images/GPU).
config.IMAGES_PER_GPU = 2
config.STEPS_PER_EPOCH = 7
# config.IMAGES_PER_GPU = 1
config.display()
```
## Configurations
```
# from keras import backend as KB
# if 'tensorflow' == KB.backend():
# import tensorflow as tf
# from keras.backend.tensorflow_backend import set_session
# # tfconfig = tf.ConfigProto(
# # gpu_options = tf.GPUOptions(per_process_gpu_memory_fraction=0.5),
# # device_count = {'GPU': 1}
# # )
# tfconfig = tf.ConfigProto()
# tfconfig.gpu_options.allow_growth=True
# tfconfig.gpu_options.visible_device_list = "0"
# tfconfig.gpu_options.per_process_gpu_memory_fraction=0.5
# tf_sess = tf.Session(config=tfconfig)
# set_session(tf_sess)
```
## Notebook Preferences
```
def get_ax(rows=1, cols=1, size=8):
"""Return a Matplotlib Axes array to be used in
all visualizations in the notebook. Provide a
central point to control graph sizes.
Change the default size attribute to control the size
of rendered images
"""
_, ax = plt.subplots(rows, cols, figsize=(size*cols, size*rows))
return ax
```
## Dataset
Create a synthetic dataset
Extend the Dataset class and add a method to load the shapes dataset, `load_shapes()`, and override the following methods:
* load_image()
* load_mask()
* image_reference()
```
# Training dataset
# generate 500 shapes
dataset_train = shapes.ShapesDataset()
dataset_train.load_shapes(500, config.IMAGE_SHAPE[0], config.IMAGE_SHAPE[1])
dataset_train.prepare()
# Validation dataset
dataset_val = shapes.ShapesDataset()
dataset_val.load_shapes(50, config.IMAGE_SHAPE[0], config.IMAGE_SHAPE[1])
dataset_val.prepare()
# Load and display random samples
# image_ids = np.random.choice(dataset_train.image_ids, 3)
# for image_id in [3]:
# image = dataset_train.load_image(image_id)
# mask, class_ids = dataset_train.load_mask(image_id)
# visualize.display_top_masks(image, mask, class_ids, dataset_train.class_names)
```
## Create Model
```
# import importlib
# importlib.reload(model)
# Create model in training mode
# MODEL_DIR = os.path.join(MODEL_PATH, "mrcnn_logs")
import gc
# del history
try :
del model
except:
pass
gc.collect()
model = modellib.MaskRCNN(mode="training", config=config, model_dir=MODEL_DIR)
print(MODEL_PATH)
print(COCO_MODEL_PATH)
print(RESNET_MODEL_PATH)
print(MODEL_DIR)
print(model.find_last())
# Which weights to start with?
init_with = "last" # imagenet, coco, or last
if init_with == "imagenet":
# loc=model.load_weights(model.get_imagenet_weights(), by_name=True)
loc=model.load_weights(RESNET_MODEL_PATH, by_name=True)
elif init_with == "coco":
# Load weights trained on MS COCO, but skip layers that
# are different due to the different number of classes
# See README for instructions to download the COCO weights
loc=model.load_weights(COCO_MODEL_PATH, by_name=True,
exclude=["mrcnn_class_logits", "mrcnn_bbox_fc",
"mrcnn_bbox", "mrcnn_mask"])
elif init_with == "last":
# Load the last model you trained and continue training
loc= model.load_weights(model.find_last()[1], by_name=True)
print('Load weights complete')
# for i in range(len(model.keras_model.layers)):
# print(i, ' Name of layer: ', model.keras_model.layers[i].name)
# sess = tf.InteractiveSession()
# model.keras_model.layers[229].output.eval()
```
## Training
Train in two stages:
1. Only the heads. Here we're freezing all the backbone layers and training only the randomly initialized layers (i.e. the ones that we didn't use pre-trained weights from MS COCO). To train only the head layers, pass `layers='heads'` to the `train()` function.
2. Fine-tune all layers. For this simple example it's not necessary, but we're including it to show the process. Simply pass `layers="all` to train all layers.
### Training head using Keras.model.fit_generator()
```
model.config.display()
# Train the head branches
# Passing layers="heads" freezes all layers except the head
# layers. You can also pass a regular expression to select
# which layers to train by name pattern.
model.train(dataset_train, dataset_val,
learning_rate=config.LEARNING_RATE,
epochs=69,
layers='heads')
```
### Training heads using train_on_batch()
We need to use this method for the time being as the fit generator does not have provide EASY access to the output in Keras call backs. By training in batches, we pass a batch through the network, pick up the generated RoI detections and bounding boxes and generate our semantic / gaussian tensors ...
```
model.train_in_batches(dataset_train, dataset_val,
learning_rate=config.LEARNING_RATE,
epochs_to_run = 5,
layers='heads')
```
#### Some network information
```
from mrcnn.draft import show_modelstuff
model.compile(model.config.LEARNING_RATE, model.config.LEARNING_MOMENTUM)
show_modelstuff(model.keras_model)
# def get_layer_output(model, model_input,output_layer, training_flag = True):
# _my_input = model_input
# for name,inp in zip(model.input_names, model_input):
# print(' Input Name: ({:24}) \t Input shape: {}'.format(name, inp.shape))
# _mrcnn_class = KB.function(model.input , model.output)
# # [model.keras_model.layers[output_layer].output])
# output = _mrcnn_class(_my_input)
# for name,out in zip (model.output_names,output):
# print(' Output Name: ({:24}) \t Output shape: {}'.format(name, out.shape))
# return output
from scipy.stats import multivariate_normal
import numpy as np
def bbox_gaussian( bbox, Zin ):
"""
receive a bounding box, and generate a gaussian distribution centered on the bounding box and with a
covariance matrix based on the width and height of the bounding box/.
Inputs :
--------
bbox : (index, class_id, class_prob, y1, x1, y2, x2)
bbox : (index, class_id, class_prob, cx, cy, width, height)
Returns:
--------
bbox_g grid mesh [image_height, image width] covering the distribution
"""
print(bbox.shape)
width = bbox[6] - bbox[4]
height = bbox[5] - bbox[3]
cx = bbox[4] + ( width / 2.0)
cy = bbox[3] + ( height / 2.0)
# cx, cy, width, height = bbox[3:]
print('center is ({},{}) width: {} height: {} '.format(cx, cy, width, height))
# srtd_cpb_2 = np.column_stack((srtd_cpb[:, 0:2], cx,cy, width, height ))
X = np.arange(0, 128, 1)
Y = np.arange(0, 128, 1)
X, Y = np.meshgrid(X, Y)
pos = np.empty(X.shape+(2,)) # concatinate shape of x to make ( x.rows, x.cols, 2)
pos[:,:,0] = X;
pos[:,:,1] = Y;
rv = multivariate_normal([cx,cy],[[12,0.0] , [0.0,19]])
Zout = rv.pdf(pos)
Zout += Zin
return Zout
from matplotlib import pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
from matplotlib import cm
from matplotlib.ticker import LinearLocator, FormatStrFormatter
def plot_gaussian( Z ):
fig = plt.figure()
ax = fig.gca(projection='3d')
X = np.arange(0, 128, 1)
Y = np.arange(0, 128, 1)
X, Y = np.meshgrid(X, Y)
pos = np.empty(X.shape+(2,)) # concatinate shape of x to make ( x.rows, x.cols, 2)
pos[:,:,0] = X;
pos[:,:,1] = Y;
surf = ax.plot_surface(X, Y, Z,cmap=cm.coolwarm, linewidth=0, antialiased=False)
# # Customize the z axis.
ax.set_zlim(0.0 , 0.05)
ax.set_ylim(0,130)
ax.set_xlim(0,130)
ax.set_xlabel(' X axis')
ax.set_ylabel(' Y axis')
ax.invert_yaxis()
ax.view_init(elev=140, azim=-88)
# ax.zaxis.set_major_locator(LinearLocator(10))
# ax.zaxis.set_major_formatter(FormatStrFormatter('%.02f'))
# Add a color bar which maps values to colors.
fig.colorbar(surf, shrink=0.5, aspect=5)
plt.show()
def stack_tensor(model):
pred_cpb_all = np.empty((0,8))
for i in range(1,model.config.NUM_CLASSES):
if pred_cls_cnt[i] > 0:
pred_cpb_all = np.vstack((pred_cpb_all, pred_cpb[i,0:pred_cls_cnt[i]] ))
from mrcnn.datagen import data_generator, load_image_gt
np.set_printoptions(linewidth=100)
train_generator = data_generator(dataset_train, model.config, shuffle=True,
batch_size=model.config.BATCH_SIZE,
augment = False)
val_generator = data_generator(dataset_val, model.config, shuffle=True,
batch_size=model.config.BATCH_SIZE,
augment=False)
# train_batch_x, train_batch_y = next(train_generator)
mm = model.keras_model
```
### Display shape loaded
```
sample_x, sample_y = next(train_generator)
sample_x, sample_y = next(train_generator)
imgmeta_idx= mm.input_names.index('input_image_meta')
img_meta = sample_x[imgmeta_idx]
image_id = img_meta[0,0]
print(' image id is :',img_meta[:,0])
image = dataset_train.load_image(image_id)
mask, class_ids = dataset_train.load_mask(image_id)
print(mask.shape, class_ids.shape)
visualize.display_top_masks(image, mask, class_ids, dataset_train.class_names)
print(' image id is :',img_meta[:,1])
image_id = img_meta[1,0]
image = dataset_train.load_image(image_id)
mask, class_ids = dataset_train.load_mask(image_id)
print(mask.shape, class_ids.shape)
visualize.display_top_masks(image, mask, class_ids, dataset_train.class_names)
# mask_0 = mask[:,:,:]
# print(mask_0.shape)
# print(mask[:,:,0])
# print('\n\n\n')
# print(mask[:,:,1])
# print('\n\n\n')
# print(mask[:,:,2])
# print('\n\n\n')
from mrcnn.utils import minimize_mask, extract_bboxes,resize_mask
import scipy
# img = 1
# image_1 = sample_x[0][img]
# class_ids_1 = sample_x[4][img,0:objs].astype(int)
# bbox_1 = sample_x[5][img,0:objs]
# mask_1 = sample_x[6][img,:,:,0:objs]
# objs = np.count_nonzero(sample_x[4][img])
# print(class_ids_1, objs)
# print(objs)
# # mask_1 = minimize_mask(sampl,sample_x[6][0,:,:,0:objs],model.config.MINI_MASK_SHAPE)
# print(mask_1.shape, bbox_1.shape)
# print(mask_1[:,:,0])
# print('\n\n\n')
# print(mask_1[:,:,1])
# print('\n\n\n')
# print(mask_1[:,:,2])
# i = 0
# mask is 128 x 128 x num_masks
# m = mask[:,:,1]
# m_str = np.array2string(m.astype('int'))
# print(m.shape, m.size)
# print(m_str)
# y1, x1, y2, x2 = bbox_1[0][:4]
# print(y1,':', y2, x1,':', x2, y2-y1, x2-x1 )
# mm = m[y1:y2, x1:x2]
# mask_1 is 56x56
# m1 = mask_1[:,:,0]
# m1_str = np.array2string(m1.astype('int'))
# print(m1.shape, m1.size)
# print(m1_str)
# m_str = np.array2string(m[0:50,:].astype(int))
# print(m.shape)
# print(m_str)
# print('\n\n\n')
# m2 = scipy.misc.imresize(m.astype(float), (56,56), interp='bilinear')
# m2_mask = np.where(m2 >= 128, 1, 0)
# m2_str = np.array2string(m2.astype('int'))
# print(m2.size)
# print(m2_str)
# mini_mask = np.zeros((56,56, objs), dtype=bool)
# mini_mask[:, :, i] = np.where(m2 >= 128, 1, 0)
# print(np.array2string(np.where(m >= 128, 1, 0)[:,:,0]))
# m[25:60,30:60]
# class_ids_1 = sample_x[4][0]
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:100% !important; }</style>"))
```
| github_jupyter |
# Text Processing
## Capturing Text Data
### Plain Text
```
import os
# Read in a plain text file
with open(os.path.join("data", "hieroglyph.txt"), "r") as f:
text = f.read()
print(text)
```
### Tabular Data
```
import pandas as pd
# Extract text column from a dataframe
df = pd.read_csv(os.path.join("data", "news.csv"))
df.head()[['publisher', 'title']]
# Convert text column to lowercase
df['title'] = df['title'].str.lower()
df.head()[['publisher', 'title']]
```
### Online Resource
```
import requests
import json
# Fetch data from a REST API
r = requests.get(
"https://quotes.rest/qod.json")
res = r.json()
print(json.dumps(res, indent=4))
# Extract relevant object and field
q = res["contents"]["quotes"][0]
print(q["quote"], "\n--", q["author"])
```
## Cleaning
```
import requests
# Fetch a web page
r = requests.get("https://news.ycombinator.com")
print(r.text)
import re
# Remove HTML tags using RegEx
pattern = re.compile(r'<.*?>') # tags look like <...>
print(pattern.sub('', r.text)) # replace them with blank
from bs4 import BeautifulSoup
# Remove HTML tags using Beautiful Soup library
soup = BeautifulSoup(r.text, "html5lib")
print(soup.get_text())
# Find all articles
summaries = soup.find_all("tr", class_="athing")
summaries[0]
# Extract title
summaries[0].find("a", class_="storylink").get_text().strip()
# Find all articles, extract titles
articles = []
summaries = soup.find_all("tr", class_="athing")
for summary in summaries:
title = summary.find("a", class_="storylink").get_text().strip()
articles.append((title))
print(len(articles), "Article summaries found. Sample:")
print(articles[0])
```
## Normalization
### Case Normalization
```
# Sample text
text = "The first time you see The Second Renaissance it may look boring. Look at it at least twice and definitely watch part 2. It will change your view of the matrix. Are the human people the ones who started the war ? Is AI a bad thing ?"
print(text)
# Convert to lowercase
text = text.lower()
print(text)
```
### Punctuation Removal
```
import re
# Remove punctuation characters
text = re.sub(r"[^a-zA-Z0-9]", " ", text)
print(text)
```
## Tokenization
```
# Split text into tokens (words)
words = text.split()
print(words)
```
### NLTK: Natural Language ToolKit
```
import os
import nltk
nltk.data.path.append(os.path.join(os.getcwd(), "nltk_data"))
# Another sample text
text = "Dr. Smith graduated from the University of Washington. He later started an analytics firm called Lux, which catered to enterprise customers."
print(text)
from nltk.tokenize import word_tokenize
# Split text into words using NLTK
words = word_tokenize(text)
print(words)
from nltk.tokenize import sent_tokenize
# Split text into sentences
sentences = sent_tokenize(text)
print(sentences)
# List stop words
from nltk.corpus import stopwords
print(stopwords.words("english"))
# Reset text
text = "The first time you see The Second Renaissance it may look boring. Look at it at least twice and definitely watch part 2. It will change your view of the matrix. Are the human people the ones who started the war ? Is AI a bad thing ?"
# Normalize it
text = re.sub(r"[^a-zA-Z0-9]", " ", text.lower())
# Tokenize it
words = text.split()
print(words)
# Remove stop words
words = [w for w in words if w not in stopwords.words("english")]
print(words)
```
### Sentence Parsing
```
import nltk
# Define a custom grammar
my_grammar = nltk.CFG.fromstring("""
S -> NP VP
PP -> P NP
NP -> Det N | Det N PP | 'I'
VP -> V NP | VP PP
Det -> 'an' | 'my'
N -> 'elephant' | 'pajamas'
V -> 'shot'
P -> 'in'
""")
parser = nltk.ChartParser(my_grammar)
# Parse a sentence
sentence = word_tokenize("I shot an elephant in my pajamas")
for tree in parser.parse(sentence):
print(tree)
```
## Stemming & Lemmatization
### Stemming
```
from nltk.stem.porter import PorterStemmer
# Reduce words to their stems
stemmed = [PorterStemmer().stem(w) for w in words]
print(stemmed)
```
### Lemmatization
```
from nltk.stem.wordnet import WordNetLemmatizer
# Reduce words to their root form
lemmed = [WordNetLemmatizer().lemmatize(w) for w in words]
print(lemmed)
# Lemmatize verbs by specifying pos
lemmed = [WordNetLemmatizer().lemmatize(w, pos='v') for w in lemmed]
print(lemmed)
```
| github_jupyter |
> Note: If you came here trying to figure out how to create simulated X-ray photons and observations,
you should go [here](http://hea-www.cfa.harvard.edu/~jzuhone/pyxsim/) instead.
This functionality provides the ability to create metallicity-dependent X-ray luminosity, emissivity, and photon emissivity fields for a given photon energy range. This works by interpolating from emission tables created from the photoionization code [Cloudy](https://www.nublado.org/) or the collisional ionization database [AtomDB](http://www.atomdb.org). These can be downloaded from https://yt-project.org/data from the command line like so:
`# Put the data in a directory you specify`
`yt download cloudy_emissivity_v2.h5 /path/to/data`
`# Put the data in the location set by "supp_data_dir"`
`yt download apec_emissivity_v2.h5 supp_data_dir`
The data path can be a directory on disk, or it can be "supp_data_dir", which will download the data to the directory specified by the `"supp_data_dir"` yt configuration entry. It is easiest to put these files in the directory from which you will be running yt or `"supp_data_dir"`, but see the note below about putting them in alternate locations.
Emission fields can be made for any energy interval between 0.1 keV and 100 keV, and will always be created for luminosity $(\rm{erg~s^{-1}})$, emissivity $\rm{(erg~s^{-1}~cm^{-3})}$, and photon emissivity $\rm{(photons~s^{-1}~cm^{-3})}$. The only required arguments are the
dataset object, and the minimum and maximum energies of the energy band. However, typically one needs to decide what will be used for the metallicity. This can either be a floating-point value representing a spatially constant metallicity, or a prescription for a metallicity field, e.g. `("gas", "metallicity")`. For this first example, where the dataset has no metallicity field, we'll just assume $Z = 0.3~Z_\odot$ everywhere:
```
import yt
ds = yt.load("GasSloshing/sloshing_nomag2_hdf5_plt_cnt_0150")
xray_fields = yt.add_xray_emissivity_field(ds, 0.5, 7.0, table_type='apec', metallicity=0.3)
```
> Note: If you place the HDF5 emissivity tables in a location other than the current working directory or the location
specified by the "supp_data_dir" configuration value, you will need to specify it in the call to
`add_xray_emissivity_field`:
`xray_fields = yt.add_xray_emissivity_field(ds, 0.5, 7.0, data_dir="/path/to/data", table_type='apec', metallicity=0.3)`
Having made the fields, one can see which fields were made:
```
print (xray_fields)
```
The luminosity field is useful for summing up in regions like this:
```
sp = ds.sphere("c", (2.0, "Mpc"))
print (sp.quantities.total_quantity(("gas","xray_luminosity_0.5_7.0_keV")))
```
Whereas the emissivity fields may be useful in derived fields or for plotting:
```
slc = yt.SlicePlot(ds, 'z', [('gas', 'xray_emissivity_0.5_7.0_keV'),
('gas', 'xray_photon_emissivity_0.5_7.0_keV')],
width=(0.75, "Mpc"))
slc.show()
```
The emissivity and the luminosity fields take the values one would see in the frame of the source. However, if one wishes to make projections of the X-ray emission from a cosmologically distant object, the energy band will be redshifted. For this case, one can supply a `redshift` parameter and a `Cosmology` object (either from the dataset or one made on your own) to compute X-ray intensity fields along with the emissivity and luminosity fields.
This example shows how to do that, Where we also use a spatially dependent metallicity field and the Cloudy tables instead of the APEC tables we used previously:
```
ds2 = yt.load("D9p_500/10MpcBox_HartGal_csf_a0.500.d")
# In this case, use the redshift and cosmology from the dataset,
# but in theory you could put in something different
xray_fields2 = yt.add_xray_emissivity_field(ds2, 0.5, 2.0, redshift=ds2.current_redshift, cosmology=ds2.cosmology,
metallicity=("gas", "metallicity"), table_type='cloudy')
```
Now, one can see that two new fields have been added, corresponding to X-ray intensity / surface brightness when projected:
```
print (xray_fields2)
```
Note also that the energy range now corresponds to the *observer* frame, whereas in the source frame the energy range is between `emin*(1+redshift)` and `emax*(1+redshift)`. Let's zoom in on a galaxy and make a projection of the energy intensity field:
```
prj = yt.ProjectionPlot(ds2, "x", ("gas","xray_intensity_0.5_2.0_keV"),
center="max", width=(40, "kpc"))
prj.set_zlim("xray_intensity_0.5_2.0_keV", 1.0e-32, 5.0e-24)
prj.show()
```
> Warning: The X-ray fields depend on the number density of hydrogen atoms, given by the yt field
`H_nuclei_density`. In the case of the APEC model, this assumes that all of the hydrogen in your
dataset is ionized, whereas in the Cloudy model the ionization level is taken into account. If
this field is not defined (either in the dataset or by the user), it will be constructed using
abundance information from your dataset. Finally, if your dataset contains no abundance information,
a primordial hydrogen mass fraction (X = 0.76) will be assumed.
Finally, if you want to place the source at a local, non-cosmological distance, you can forego the `redshift` and `cosmology` arguments and supply a `dist` argument instead, which is either a `(value, unit)` tuple or a `YTQuantity`. Note that here the redshift is assumed to be zero.
```
xray_fields3 = yt.add_xray_emissivity_field(ds2, 0.5, 2.0, dist=(1.0,"Mpc"), metallicity=("gas", "metallicity"),
table_type='cloudy')
prj = yt.ProjectionPlot(ds2, "x", ("gas", "xray_photon_intensity_0.5_2.0_keV"),
center="max", width=(40, "kpc"))
prj.set_zlim("xray_photon_intensity_0.5_2.0_keV", 1.0e-24, 5.0e-16)
prj.show()
```
| github_jupyter |
# H2O.ai GPU Edition Machine Learning $-$ Multi-GPU GLM Demo
### In this demo, we will train 4000 regularized linear regression models (aka Generalized Linear Models or GLMs) on the U.S. Census dataset, with the goal to predict the earned income of a person, given approximately 10000 features such as gender, age, occupation, zip code, etc.
### The dataset is about 2GB in memory (50k rows, 10k cols, single-precision floating-point values), so it easily fits onto the GPU memory.
### By using multiple GPUs, we are able to speed up this process significantly, and can train about 40 models per second (on a DGX-1 with 8 GPUs) vs 1 model per second on dual-Xeon server.
### Import Dependencies
```
## First time only: Install dependencies
#!pip install https://s3.amazonaws.com/h2o-beta-release/goai/h2o4gpu-0.0.2-py2.py3-none-any.whl
#!pip install Cython pandas seaborn psutil feather_format
#!pip install -e "git+https://github.com/fbcotter/py3nvml#egg=py3nvml"
## Now restart the kernel to get py3nvml to work
%reset -f
%matplotlib inline
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
```
### Import Data Frame and create raw X and y arrays
```
import os, sys, time
import feather
file = os.path.join(os.getcwd(), "ipums.feather")
if not os.path.exists(file):
!wget https://s3.amazonaws.com/h2o-public-test-data/h2o4gpu/open_data/ipums.feather
t0 = time.time()
df = feather.read_dataframe(file)
t1 = time.time()
print("Time to read data via feather: %r" % (t1-t0))
## We predict the last column "INCEARN" - Income earned
target = df.columns[-1]
cols = [c for c in df.columns if c != target]
df[target].hist(bins=50)
import numpy as np
X = np.array(df.ix[:,cols], order='f').astype('float32')
y = np.array(df[target].values, dtype='float32')
print(X.shape)
print(y.shape)
```
### Split the dataset into Training (80%) and Validation (20%)
```
intercept = True
validFraction=0.2
standardize = 0
lambda_min_ratio = 1e-7
if standardize:
print ("implement standardization transformer")
exit()
# Setup Train/validation Set Split
morig = X.shape[0]
norig = X.shape[1]
fortran = X.flags.f_contiguous
print("fortran order=%d" % (fortran))
print("Original data rows=%d cols=%d" % (morig,norig))
# Do train/valid split
HO=int(validFraction*morig)
H=morig-HO
print("Training rows=%d" % (H))
print("Vaidation rows=%d" % (HO))
trainX = np.copy(X[0:H,:])
trainY = np.copy(y[0:H])
validX = np.copy(X[H:morig,:])
validY = np.copy(y[H:morig])
trainW = np.copy(trainY)*0.0 + 1.0 # constant unity weight
mTrain = trainX.shape[0]
mvalid = validX.shape[0]
# if using upload_data and fit_ptr, then have to create own intercept as last column in trainX and validX
if intercept:
trainX = np.hstack([trainX, np.ones((trainX.shape[0],1),dtype=trainX.dtype)])
validX = np.hstack([validX, np.ones((validX.shape[0],1),dtype=validX.dtype)])
n = trainX.shape[1]
print("New data rows=%d cols=%d" % (mTrain,n))
```
### Define some helper methods for plotting and running the algorithm
```
import seaborn as sns
sns.set_style("whitegrid")
def new_alpha(row_fold):
if row_fold == 0:
return -0.025
elif row_fold == 1:
return -0.05
elif row_fold == 3:
return 0.025
elif row_fold == 4:
return 0.05
else: return 0
def plot_cpu_perf(axis, cpu_labels, cpu_snapshot):
axis.cla()
axis.grid(False)
axis.set_ylim([0,100])
axis.set_ylabel('Percent', labelpad=2, fontsize = 14)
axis.bar(cpu_labels, cpu_snapshot, color='dodgerblue', edgecolor='none')
axis.set_title('CPU Utilization', fontsize = 16)
def plot_gpu_perf(axis, gpu_labels, gpu_snapshot):
axis.cla()
axis.grid(False)
axis.set_ylim([0,100])
axis.set_xticks(gpu_labels)
axis.set_ylabel('Percent', labelpad=2, fontsize = 14)
axis.bar(gpu_labels, gpu_snapshot, width =0.5, color = 'limegreen',align='center', edgecolor='none')
axis.set_title('GPU Utilization', fontsize = 16)
def plot_glm_results(axis, results, best_rmse, cb):
axis.cla()
axis.set_xscale('log')
axis.set_xlim([1e2, 1e9])
axis.set_ylim([-0.12, 1.12])
axis.set_yticks([x/7. for x in range(0,8)])
axis.set_ylabel('Parameter 1: '+r'$\alpha$', fontsize = 16)
axis.set_xlabel('Parameter 2: '+r'$\lambda$', fontsize = 16)
num_models = min(4000,int(4000*results.shape[0]/2570))
axis.set_title('Elastic Net Models Trained and Evaluated: ' + str(num_models), fontsize = 16)
try:
from matplotlib.colors import ListedColormap
cm = ListedColormap(sns.color_palette("RdYlGn", 10).as_hex())
cf = axis.scatter(results['lambda'], results['alpha_prime'], c=results['rel_acc'],
cmap=cm, vmin=0, vmax=1, s=60, lw=0)
axis.plot(best_rmse['lambda'],best_rmse['alpha_prime'], 'o',
ms=15, mec='k', mfc='none', mew=2)
if not cb:
cb = pl.colorbar(cf, ax=axis)
cb.set_label('Relative Validation Accuracy', rotation=270,
labelpad=18, fontsize = 16)
cb.update_normal(cf)
except:
#print("plot_glm_results exception -- no frame")
pass
def RunAnimation(arg):
import os, sys, time
import subprocess
import psutil
import pylab as pl
from IPython import display
import matplotlib.gridspec as gridspec
import seaborn as sns
import pandas as pd
import numpy as np
print("RunAnimation")
sys.stdout.flush()
deviceCount = arg
# Need this only for animation of GPU usage to be consistent with
#from py3nvml.py3nvml import *
import py3nvml
maxNGPUS = int(subprocess.check_output("nvidia-smi -L | wc -l", shell=True))
print("\nNumber of GPUS:", maxNGPUS)
py3nvml.py3nvml.nvmlInit()
total_deviceCount = py3nvml.py3nvml.nvmlDeviceGetCount()
if deviceCount == -1:
deviceCount = total_deviceCount
#for i in range(deviceCount):
# handle = nvmlDeviceGetHandleByIndex(i)
# print("Device {}: {}".format(i, nvmlDeviceGetName(handle)))
#print ("Driver Version:", nvmlSystemGetDriverVersion())
print("Animation deviceCount=%d" % (deviceCount))
file = os.getcwd() + "/error.txt"
print("opening %s" % (file))
fig = pl.figure(figsize = (9,9))
pl.rcParams['xtick.labelsize'] = 14
pl.rcParams['ytick.labelsize'] = 14
gs = gridspec.GridSpec(3, 2, wspace=0.3, hspace=0.4)
ax1 = pl.subplot(gs[0,-2])
ax2 = pl.subplot(gs[0,1])
ax3 = pl.subplot(gs[1:,:])
fig.suptitle('H2O.ai Machine Learning $-$ Generalized Linear Modeling', size=18)
pl.gcf().subplots_adjust(bottom=0.2)
#cb = False
from matplotlib.colors import ListedColormap
cm = ListedColormap(sns.color_palette("RdYlGn", 10).as_hex())
cc = ax3.scatter([0.001, 0.001], [0,0], c =[0,1], cmap = cm)
cb = pl.colorbar(cc, ax=ax3)
os.system("mkdir -p images")
i=0
while(True):
#try:
#print("In try i=%d" % i)
#sys.stdout.flush()
#cpu
snapshot = psutil.cpu_percent(percpu=True)
cpu_labels = range(1,len(snapshot)+1)
plot_cpu_perf(ax1, cpu_labels, snapshot)
#gpu
gpu_snapshot = []
gpu_labels = list(range(1,deviceCount+1))
import py3nvml
for j in range(deviceCount):
handle = py3nvml.py3nvml.nvmlDeviceGetHandleByIndex(j)
util = py3nvml.py3nvml.nvmlDeviceGetUtilizationRates(handle)
gpu_snapshot.append(util.gpu)
gpu_snapshot = gpu_snapshot
plot_gpu_perf(ax2, gpu_labels, gpu_snapshot)
res = pd.read_csv(file, sep="\s+",header=None,names=['time','pass','fold','a','i','alpha','lambda','trainrmse','ivalidrmse','validrmse'])
res['rel_acc'] = ((42665- res['validrmse'])/(42665-31000))
res['alpha_prime'] = res['alpha'] + res['fold'].apply(lambda x: new_alpha(x))
best = res.ix[res['rel_acc']==np.max(res['rel_acc']),:]
plot_glm_results(ax3, res, best.tail(1), cb)
# flag for colorbar to avoid redrawing
#cb = True
# Add footnotes
footnote_text = "*U.S. Census dataset (predict Income): 45k rows, 10k cols\nParameters: 5-fold cross-validation, " + r'$\alpha = \{\frac{i}{7},i=0\ldots7\}$' + ", "\
'full $\lambda$-' + "search"
#pl.figtext(.05, -.04, footnote_text, fontsize = 14,)
pl.annotate(footnote_text, (0,0), (-30, -50), fontsize = 12,
xycoords='axes fraction', textcoords='offset points', va='top')
#update the graphics
display.display(pl.gcf())
display.clear_output(wait=True)
time.sleep(0.01)
#save the images
saveimage=0
if saveimage:
file_name = './images/glm_run_%04d.png' % (i,)
pl.savefig(file_name, dpi=200)
i=i+1
#except KeyboardInterrupt:
# break
#except:
# #print("Could not Create Frame")
# pass
def RunH2Oaiglm(arg):
import h2o4gpu as h2o4gpu
intercept, lambda_min_ratio, nFolds, n_alphas, n_lambdas, n_gpus = arg
print("Begin Setting up Solver")
os.system("rm -f error.txt ; touch error.txt ; rm -f varimp.txt ; touch varimp.txt") ## for visualization
enet = h2o4gpu.ElasticNetH2O(n_gpus=n_gpus, fit_intercept=intercept, lambda_min_ratio=lambda_min_ratio, n_lambdas=n_lambdas, n_folds=n_folds, n_alphas=n_alphas)
print("End Setting up Solver")
## First, get backend pointers
sourceDev=0
t0 = time.time()
a,b,c,d,e = enet.prepare_and_upload_data(trainX, trainY, validX, validY, trainW, source_dev=sourceDev)
t1 = time.time()
print("Time to ingest data: %r" % (t1-t0))
## Solve
if 1==1:
print("Solving")
t0 = time.time()
order='c' if fortran else 'r'
double_precision=0 # Not used
store_full_path=0
enet.fit_ptr(mTrain, n, mvalid, double_precision, order, a, b, c, d, e, source_dev=sourceDev)
t1 = time.time()
print("Done Solving")
print("Time to train H2O AI GLM: %r" % (t1-t0))
```
### Train 4000 Elastic Net Models (5-fold cross-validation, 8 $\alpha$ values, 100 $\lambda$ values)
```
import subprocess
import concurrent.futures
from concurrent.futures import ProcessPoolExecutor
lambda_min_ratio=1E-9
n_folds=5
n_alphas=8
n_lambdas=100
n_gpus=-1 # -1 means use all GPUs
arg = intercept, lambda_min_ratio, n_folds, n_alphas, n_lambdas, n_gpus
from threading import Thread
background_thread = Thread(target=RunH2Oaiglm, args=(arg,))
background_thread.start()
#futures = []
#Executor = ProcessPoolExecutor(max_workers=1)
#futures.append(Executor.submit(RunH2Oaiglm, arg)) ## run in separate process
RunAnimation(n_gpus)
#concurrent.futures.wait(futures)
```
| github_jupyter |
# Predict heart failure with Watson Machine Learning

This notebook contains steps and code to create a predictive model to predict heart failure and then deploy that model to Watson Machine Learning so it can be used in an application.
## Learning Goals
The learning goals of this notebook are:
* Load a CSV file into the Object Storage service linked to your Watson Studio
* Create an Apache Spark machine learning model
* Train and evaluate a model
* Persist a model in a Watson Machine Learning repository
## 1. Setup
Before you use the sample code in this notebook, you must perform the following setup tasks (also mentioned in the course "Analyzing and Predicting Heart Failure on IBM Cloud"):
* Create a Watson Machine Learning service instance (a free plan is offered) and associate it with your project
* Upload heart failure data to the Object Store service that is part of Watson Studio
We'll be using a few libraries for this exercise:
1. [Machine learning and AI in Watson Studio](https://dataplatform.cloud.ibm.com/docs/content/wsj/analyze-data/wml-ai.html): Client library to work with the Watson Machine Learning service on IBM Cloud.
1. [Pixiedust](https://github.com/pixiedust/pixiedust): Python Helper library for Jupyter Notebooks
1. [ibmos2spark](https://github.com/ibm-watson-data-lab/ibmos2spark): Facilitates Data I/O between Spark and IBM Object Storage services
```
!pip install -U ibm-watson-machine-learning
!pip install --upgrade pixiedust
```
Next we will try to import the sparksession just to see if everything is ok. If all is good, then you should see no errors raised after executing the cell.
```
try:
from pyspark.sql import SparkSession
except:
print('Error: Spark runtime is missing. If you are using Watson Studio change the notebook runtime to Spark.')
raise
```
## 2. Load and explore data
In this section you will load the data as an Apache Spark DataFrame and perform a basic exploration. Load the data to the Spark DataFrame from your associated Object Storage instance.
> **IMPORTANT**: Follow the lab instructions to insert an Apache Spark DataFrame in the cell below.
> **IMPORTANT**: Ensure the DataFrame is named `df_data`.
> **IMPORTANT**: Add `.option('inferSchema','True')\` to the inserted code.
```
.option('inferSchema','True')\
```
Explore the loaded data by using the following Apache® Spark DataFrame methods:
* `df_data.printSchema` to print the data schema
* `df_data.describe()` to print the top twenty records
* `df_data.count()` to count all records
```
df_data.printSchema()
```
As you can see, the data contains ten fields. The HEARTFAILURE field is the one we would like to predict (label).
```
df_data.show()
df_data.describe().show()
df_data.count()
```
As you can see, the data set contains 10800 records.
## 3. Interactive Visualizations w/PixieDust
```
import pixiedust
```
### Simple visualization using bar charts
With PixieDust's `display()` method you can visually explore the loaded data using built-in charts, such as, bar charts, line charts, scatter plots, or maps.
To explore a data set: choose the desired chart type from the drop down, configure chart options, configure display options.
```
display(df_data)
```
## 4. Create a Spark machine learning model
In this section you will learn how to prepare data, create and train a Spark machine learning model.
### 4.1 Prepare data
In this subsection you will split your data into: train and test data sets.
```
split_data = df_data.randomSplit([0.8, 0.20], 24)
train_data = split_data[0]
test_data = split_data[1]
print("Number of training records: " + str(train_data.count()))
print("Number of testing records : " + str(test_data.count()))
```
As you can see our data has been successfully split into two data sets:
* The train data set, which is the largest group, is used for training.
* The test data set will be used for model evaluation and is used to test the assumptions of the model.
### 4.2 Create pipeline and train a model
In this section you will create a Spark machine learning pipeline and then train the model. In the first step you need to import the Spark machine learning packages that will be needed in the subsequent steps. A sequence of data processing is called a _data pipeline_. Each step in the pipeline processes the data and passes the result to the next step in the pipeline, this allows you to transform and fit your model with the raw input data.
```
from pyspark.ml.feature import StringIndexer, IndexToString, VectorAssembler
from pyspark.ml.classification import RandomForestClassifier
from pyspark.ml.evaluation import MulticlassClassificationEvaluator
from pyspark.ml import Pipeline, Model
```
In the following step, convert all the string fields to numeric ones by using the StringIndexer transformer.
```
stringIndexer_label = StringIndexer(inputCol="HEARTFAILURE", outputCol="label").fit(df_data)
stringIndexer_sex = StringIndexer(inputCol="SEX", outputCol="SEX_IX")
stringIndexer_famhist = StringIndexer(inputCol="FAMILYHISTORY", outputCol="FAMILYHISTORY_IX")
stringIndexer_smoker = StringIndexer(inputCol="SMOKERLAST5YRS", outputCol="SMOKERLAST5YRS_IX")
```
In the following step, create a feature vector by combining all features together.
```
vectorAssembler_features = VectorAssembler(inputCols=["AVGHEARTBEATSPERMIN","PALPITATIONSPERDAY","CHOLESTEROL","BMI","AGE","SEX_IX","FAMILYHISTORY_IX","SMOKERLAST5YRS_IX","EXERCISEMINPERWEEK"], outputCol="features")
```
Next, define estimators you want to use for classification. Random Forest is used in the following example.
```
rf = RandomForestClassifier(labelCol="label", featuresCol="features")
```
Finally, indexed labels back to original labels.
```
labelConverter = IndexToString(inputCol="prediction", outputCol="predictedLabel", labels=stringIndexer_label.labels)
transform_df_pipeline = Pipeline(stages=[stringIndexer_label, stringIndexer_sex, stringIndexer_famhist, stringIndexer_smoker, vectorAssembler_features])
transformed_df = transform_df_pipeline.fit(df_data).transform(df_data)
transformed_df.show()
```
Let's build the pipeline now. A pipeline consists of transformers and an estimator.
```
pipeline_rf = Pipeline(stages=[stringIndexer_label, stringIndexer_sex, stringIndexer_famhist, stringIndexer_smoker, vectorAssembler_features, rf, labelConverter])
```
Now, you can train your Random Forest model by using the previously defined **pipeline** and **training data**.
```
model_rf = pipeline_rf.fit(train_data)
```
You can check your **model accuracy** now. To evaluate the model, use **test data**.
```
predictions = model_rf.transform(test_data)
evaluatorRF = MulticlassClassificationEvaluator(labelCol="label", predictionCol="prediction", metricName="accuracy")
accuracy = evaluatorRF.evaluate(predictions)
print("Accuracy = %g" % accuracy)
print("Test Error = %g" % (1.0 - accuracy))
```
You can tune your model now to achieve better accuracy. For simplicity of this example tuning section is omitted.
## 5. Persist model
In this section you will learn how to store your pipeline and model in Watson Machine Learning repository by using Python client libraries that we installed earlier
> **IMPORTANT**: Update the `wml_credentials` variable below. Replace the value (Replace me) for apikey with the APIKEY that you copied earlier within the course, as for the URL, replace the value with one of these based on what location your machine learning service is based in:
Dallas - "https://us-south.ml.cloud.ibm.com"
London - "https://eu-gb.ml.cloud.ibm.com"
Frankfurt - "https://eu-de.ml.cloud.ibm.com"
Tokyo - "https://jp-tok.ml.cloud.ibm.com"
```
from ibm_watson_machine_learning import APIClient
wml_credentials = {
"url": "(Replace Me)",
"apikey":"(Replace Me)"
}
client = APIClient(wml_credentials)
```
Just to test we print the client version.
```
print(client.version)
```
> **IMPORTANT**: Update the `space_uid` variable below. Replace the value (Replace me) with the Space UID that you copied earlier within the course. You can also get your space id by running this code: client.spaces.list(limit=5)
```
client.spaces.list(limit=5)
space_uid = "(Replace Me)"
```
Now we set it as our default space
```
client.set.default_space(space_uid)
```
Now we specify some software sepecifications and create model artifact (abstraction layer) for the model to run properly. It has already been filled for you.
```
# Model Metadata
software_spec_uid = client.software_specifications.get_uid_by_name('spark-mllib_2.4-py37')
model_props={
client.repository.ModelMetaNames.NAME: "Heart failure",
client.repository.ModelMetaNames.SPACE_UID: space_uid,
client.repository.ModelMetaNames.SOFTWARE_SPEC_UID: software_spec_uid,
client.repository.ModelMetaNames.TYPE: "mllib_2.4"
}
published_model = client.repository.store_model(model=model_rf, pipeline=pipeline_rf, meta_props=model_props, training_data=train_data)
```
## 5.1 Save pipeline and model
In this subsection you will learn how to save pipeline and model artifacts to your Watson Machine Learning instance.
Let's try to print the ID of the published model just to double check everything.
```
published_model_ID = client.repository.get_model_uid(published_model)
print("Model Id: " + str(published_model_ID))
```
## 5.2 Load model to verify that it was saved correctly
You can load your model to make sure that it was saved correctly.
```
loaded_model = client.repository.load(published_model_ID)
print(loaded_model)
```
Call model against test data to verify that it has been loaded correctly. Examine top 3 results
```
test_predictions = loaded_model.transform(test_data)
test_predictions.select('probability', 'predictedLabel').show(n=3, truncate=False)
```
## <font color=green>Congratulations</font>, you've sucessfully created a predictive model and saved it in the Watson Machine Learning service.
That's about it for the notebook. Please make sure you save your work and then switch back to the course to see how to deploy and integrate this model with your web app.
***
| github_jupyter |
```
import SimPEG as simpeg
from SimPEG import NSEM
from glob import glob
import numpy as np, sys, matplotlib.pyplot as plt
# sys.path.append('/home/gudni/Dropbox/Work/UBCwork/SyntheticModels/SynGeothermalStructures/ThesisModels')
# import synhelpFunc
def convergeCurves(resList):
its = np.array([res['iter'] for res in resList]).T
ind = np.argsort(its)
phid = np.array([res['phi_d'] for res in resList]).T
try:
phim = np.array([res['phi_m'] for res in resList]).T
except:
phim = np.array([res['phi_ms'] for res in resList]).T + np.array([res['phi_mx'] for res in resList]).T + np.array([res['phi_my'] for res in resList]).T + np.array([res['phi_mz'] for res in resList]).T
x = np.arange(len(its))
fig, ax1 = plt.subplots()
ax1.semilogy(x,phid[ind],'bx--')
ax1.set_ylabel('phi_d', color='b')
plt.hlines(len(resList[0]['dpred'])*.75,0,len(x),colors='g',linestyles='-.')
for tl in ax1.get_yticklabels():
tl.set_color('b')
ax2 = ax1.twinx()
ax2.semilogy(x,phim[ind],'rx--',)
ax2.set_ylabel('phi_m', color='r')
for tl in ax2.get_yticklabels():
tl.set_color('r')
plt.show()
def tikanovCurve(resList):
its = np.array([res['iter'] for res in resList]).T
ind = np.argsort(its)
phid = np.array([res['phi_d'] for res in resList]).T
phim = np.array([res['phi_m'] for res in resList]).T
x = np.arange(len(its))
fig, ax1 = plt.subplots()
ax1.loglog(phim[ind],phid[ind],'bx--')
ax1.set_ylabel('phi_d')
ax1.set_xlabel('phi_m')
plt.hlines(len(resList[0]['dpred'])*.75,np.min(phim),np.max(phim),colors='g',linestyles='-.')
plt.show()
def allconvergeCurves(resList):
its = np.array([res['iter'] for res in resList]).T
ind = np.argsort(its)
phid = np.array([res['phi_d'] for res in resList]).T
phim = np.array([res['phi_m'] for res in resList]).T
phims = np.array([res['phi_ms'] for res in resList]).T
phimx = np.array([res['phi_mx'] for res in resList]).T
phimy = np.array([res['phi_my'] for res in resList]).T
phimz = np.array([res['phi_mz'] for res in resList]).T
x = np.arange(len(its))
fig, ax1 = plt.subplots()
ax1.semilogy(x,phid[ind],'bx--',label='phid')
ax1.set_ylabel('phi_d', color='b')
plt.hlines(len(resList[0]['dpred'])*.75,0,len(x),colors='g',linestyles='-.')
for tl in ax1.get_yticklabels():
tl.set_color('b')
ax1.semilogy(x,phim[ind],'gx--',label='phim')
ax1.semilogy(x,phims[ind],'y,--',label='phims')
ax1.semilogy(x,phimx[ind],'r.--',label='phimx')
ax1.semilogy(x,phimy[ind],'r+--',label='phimy')
ax1.semilogy(x,phimz[ind],'r*--',label='phimz')
plt.legend()
plt.show()
def loadInversionMakeVTRFiles(dirStr,mesh,mapping):
temp = [np.load(f) for f in glob(dirStr+'/*Inversion*.npz')]
iterResults = [i if len(i.keys()) > 1 else i['arr_0'].tolist() for i in temp ]
# Make the vtk models
for it in iterResults:
mesh.writeVTK(dirStr+'/recoveredMod_{:s}_it{:.0f}.vtr'.format(dirStr,int(it['iter'])),{'S/m':mapping*it['m']})
return iterResults
# Load the model
mesh, modDict = simpeg.Mesh.TensorMesh.readVTK('nsmesh_GKRCoarseHKPK1.vtr')
sigma = modDict['S/m']
# Make the mapping
active = sigma != 1e-8
actMap = simpeg.Maps.InjectActiveCells(mesh, active, np.log(1e-8), nC=mesh.nC)
mappingExpAct = simpeg.Maps.ExpMap(mesh) * actMap
# Load the data
drecAll = np.load('MTdataStArr_nsmesh_GKRHKPK1.npy')
```
### run1
```
run1Files = loadInversionMakeVTRFiles('run1',mesh,mappingExpAct)
ls run1
%matplotlib inline
convergeCurves(run1Files)
[res['iter'] for res in run1Files]
%matplotlib qt
sys.path.append('/home/gudni/Dropbox/code/python/MTview/')
import interactivePlotFunctions as iPf
finData = NSEM.Data(survey,run1Files[np.argmax([it['iter'] for it in run1Files])]['dpred']).toRecArray('Complex')
%matplotlib notebook
iPf.MTinteractiveMap([dUse.toRecArray('Complex'),finData])
```
### run2
```
run2Files = loadInversionMakeVTRFiles('run2',mesh,mappingExpAct)
%matplotlib inline
convergeCurves(run2Files)
finData = NSEM.Data(survey,run2Files[0]['dpred']).toRecArray('Complex')
```
| github_jupyter |
# Blood Fractionation Method Analysis
First, we configure several variables we are using during this demo: project id, S3 profile and VM path to save data files from the S3 bucket:
```
from cdispyutils.hmac4 import get_auth
import requests, json, re
import pandas as pd
project = ''
profile = ''
path = 'files/'
```
### Get Authorization using your keys from the BPA API: data.bloodpac.org/identity
These keys must be loaded in your VM's .secrets file. In the case of an access error, you may need a new keypair.
```
with open('/home/ubuntu/.secrets','r') as f:
secrets = json.load(f)
auth = get_auth(secrets['access_key'], secrets['secret_key'], 'submission')
```
### Querying data from graphQL
** Find how many samples in commons have linked quantification assays:|**
```
# Setting 'gql' to the BloodPAC API graphQL url:
api='http://kubenode.internal.io:30006/v0/submission/bpa/'
gql = 'http://kubenode.internal.io:30006/v0/submission/graphql/'
data = {'query': """ {
_sample_count(with_path_to:{type:"quantification_assay"})
} """};
resp = requests.post(gql,auth=auth,json=data)
scount = re.search(".*_sample_count\": (\d+)",str(resp.text)).group(1)
print resp.text
scount
```
** Another way to do the same thing and get the samples' submitter_id and fractionation_method: **
```
data = {'query': """ {
sample(first:0,with_path_to:{type:"quantification_assay"}){
submitter_id
blood_fractionation_method
}
} """};
resp = requests.post(gql,auth=auth,json=data)
samps = re.findall(".*submitter_id\": \"(.+)\"",str(resp.text))
methods = re.findall('.*blood_fractionation_method\": \"(.+)\"',str(resp.text))
z = zip(samps,methods)
z = list(set(z))
len(z)
print "There are "+str(len(set(methods)))+" different blood fractionation methods for "+str(len(set(samps)))+" samples with links to quantification assays."
# List the Sample IDs and their blood fractionation method
zdf = pd.DataFrame(data=z)
zdf.columns = ['Sample ID', 'Blood Fractionation Method']
zdf
```
** Find the 'blood_fractionation_method" that yields the highest 'molecular_concentration':|**
** You will see that the following query in this cell returns: "Request Timeout" because it is returning too many results: **
```
data = {'query': """{
quantification_assay(first:0,
not: {molecular_concentration:0}
order_by_desc:"molecular_concentration") {
molecular_concentration
project_id
analytes(first:0){
aliquots(first:0){
samples(first:0){
submitter_id
blood_fractionation_method
}
}
}
}
} """};
resp = requests.post(gql, auth=auth, json=data)
print resp.text
```
** Using pagination with graphQL: **
We can use pagination (using combination of "offset" and "first") inside a loop to get all the data.
When no data is returned, the API response is: `"quantification_assay": []`, which is when we break out of the loop.
When this string is matched, the length of r is "1" and our loops terminates.
If any data is returned, there is no match, and len(r) is "0".
```
response = "" # this string will hold all the results
offset=0 # start at the first result, with no offset
r = () # r is a list that will contain the result of the regular expression that checks whether data is returned
while len(r) != 1:
q = """{
quantification_assay(first:100,offset:"""+str(offset)+""",
not: {molecular_concentration:0}
order_by_desc:"molecular_concentration"){
molecular_concentration
project_id
analytes(first:0){
aliquots(first:0){
samples(first:0){
submitter_id
blood_fractionation_method
}
}
}
}
} """
data = {'query':q}
resp = requests.post(gql, auth=auth, json=data)
response += str(resp.text) # concatenate the returned data
r = re.findall(".*quantification_assay\": \[\]",str(resp.text)) #this is where we check whether data was returned
offset+=100 #increase the offset to get the next 100 records
```
** Using REGEX to extract our data from the response: **
```
methods = re.findall('.*blood_fractionation_method\": \"(.+)\"',response)
concs = re.findall(".*molecular_concentration\": (.+),",response)
c = map(float,concs)
ids = re.findall(".*submitter_id\": \"(.+)\"",response)
z = zip(methods,c)
assays = dict(zip(ids,z ))
#a = pd.DataFrame(data=assays)
a = pd.Series(assays, name='Method, Molecular Concentration')
a.index.name = 'Sample ID'
a.reset_index()
print("There are " + str(len(set(ids))) + " samples with " +
str(len(set(methods))) + " different blood fractionation methods for " +
str(len(concs)) + " different quantification assays with non-zero molecular concentrations.")
```
### 2) Plot the distribution of molecular concentrations
```
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.mlab as mlab
#plt.hist(numpy.log(c))
plt.hist(c)
plt.show()
print("mean: "+str(np.mean(c))+", median: "+str(np.median(c))+ ", max: "+str(max(c))+", min: "+str(min(c))+".")
# the histogram of the data
plt.subplot(211)
n, bins, patches = plt.hist(c, 2000, normed=1, facecolor='green', alpha=0.75)
p = plt.hist(c, 100, normed=1, facecolor='green', alpha=0.75)
# add a 'best fit' line
mu = np.mean(c)
sigma = np.std(c)
y = mlab.normpdf( bins, mu, sigma)
l = plt.plot(bins, y, 'r--', linewidth=1)
plt.xlabel('DNA Concentration')
plt.ylabel('Frequency')
plt.title(r'$\mathrm{Histogram\ of\ [DNA]}$')
plt.axis([0, 850, 0, 0.05])
plt.grid(True)
plt.subplot(212)
p = plt.hist(c, 2000, normed=1, facecolor='green', alpha=0.75)
plt.axis([0, 10, 0, 0.2])
plt.grid(True)
plt.show(p)
```
# Question: How many samples have blood_fractionation_method and molecular concentration? Can you export a table with 2 columns for these two fields?
```
q = """{
sample(first:0,
with_path_to:[
{type:"quantification_assay"},
{type:"analyte",analyte_type:"DNA"}]) {
blood_fractionation_method
submitter_id
project_id
aliquots(first:0,with_path_to:[{type:"quantification_assay"},{type:"analyte",analyte_type:"DNA"}]) {
analytes(first:0,analyte_type:"DNA") {
quantification_assays(first:0) {
molecular_concentration
}
}
}
}
}"""
data = {'query':q}
resp = requests.post(gql, auth=auth, json=data)
jresp = json.loads(resp.text)
#jresp
mc = []
for i in jresp["data"]["sample"]:
subid = i["submitter_id"]
projid = i["project_id"]
bfm = i["blood_fractionation_method"]
dnac = []
for j in i["aliquots"]:
for k in j["analytes"]:
for l in k["quantification_assays"]:
dnac.append(l["molecular_concentration"])
line = str(subid)+"\t"+str(projid)+"\t"+str(bfm)+"\t"+str(dnac)
# print line
mcs = [subid,projid,bfm,dnac]
mc.append(mcs)
print "There are "+str(len(mc))+" samples with 'blood_fractionation_method' and 'molecular concentration' and links to DNA analytes."
m = pd.DataFrame(data=mc)
m.columns = ['Sample ID', 'Project ID', 'Blood Fractionation Method', 'Molecular Concentration']
m
```
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
from kyle.sampling.fake_clf import DirichletFC
from kyle.evaluation import EvalStats, compute_accuracy, compute_ECE, compute_expected_max
from kyle.transformations import *
import numpy as np
import matplotlib.pyplot as plt
import logging
logging.basicConfig(level=logging.INFO)
n_samples = 100000
```
# Dirichlet fake classifiers
Add explanation about the model and integrals
## Computing properties with integrals
The asymptotic values for ECE and accuracy can be computed through (numerical or analytical)
integration.
```
n_classes = 3
alpha=[0.2, 0.3, 0.4]
dirichlet_fc = DirichletFC(n_classes, alpha=alpha)
print("mostly underestimating all classes (starting at 1/n_classes) with PowerLawSimplexAut")
transform = PowerLawSimplexAut(np.array([2, 2, 2]))
dirichlet_fc.set_simplex_automorphism(transform)
eval_stats = EvalStats(*dirichlet_fc.get_sample_arrays(n_samples))
print(f"Accuracy is {eval_stats.accuracy()}")
print(f"ECE is {eval_stats.expected_calibration_error()}")
ece_approx = - eval_stats.expected_confidence() + eval_stats.accuracy()
print(f"{ece_approx=}")
eval_stats.plot_reliability_curves([0, 1, EvalStats.TOP_CLASS_LABEL], display_weights=True)
theoretical_acc = compute_accuracy(dirichlet_fc)[0]
theoretical_ece = compute_ECE(dirichlet_fc)[0]
print(f"{theoretical_acc=} , {theoretical_ece=}")
print("mostly overestimating all classes (starting at 1/n_classes) with PowerLawSimplexAut")
print("Note the variance and the resulting sensitivity to binning")
transform = PowerLawSimplexAut(np.array([0.3, 0.1, 0.2]))
dirichlet_fc.set_simplex_automorphism(transform)
eval_stats = EvalStats(*dirichlet_fc.get_sample_arrays(n_samples), bins=500)
print(f"Accuracy is {eval_stats.accuracy()}")
print(f"ECE is {eval_stats.expected_calibration_error()}")
ece_approx = eval_stats.expected_confidence() - eval_stats.accuracy()
print(f"{ece_approx=}")
eval_stats.plot_reliability_curves([0, 1, EvalStats.TOP_CLASS_LABEL], display_weights=True)
# theoretical_acc = compute_accuracy(dirichlet_fc)[0]
# theoretical_ece = compute_ECE(dirichlet_fc)[0]
# print(f"{theoretical_acc=} , {theoretical_ece=}")
print("Overestimating predictions with MaxComponent")
def overestimating_max(x: np.ndarray):
x = x.copy()
mask = x > 1/2
x[mask] = x[mask] - (1/4 - (1-x[mask])**2)
return x
transform = MaxComponentSimplexAut(overestimating_max)
dirichlet_fc.set_simplex_automorphism(transform)
eval_stats = EvalStats(*dirichlet_fc.get_sample_arrays(n_samples))
print(f"Accuracy is {eval_stats.accuracy()}")
print(f"ECE is {eval_stats.expected_calibration_error()}")
eval_stats.plot_reliability_curves([0, 1, EvalStats.TOP_CLASS_LABEL], display_weights=True)
# Integrals converge pretty slowly, this takes time
# theoretical_acc = compute_accuracy(dirichlet_fc, opts={"limit": 75})[0]
# theoretical_ece = compute_ECE(dirichlet_fc, opts={"limit": 75})[0]
# print(f"{theoretical_acc=} , {theoretical_ece=}")
```
# Analytical results
For top-class overconfident classifiers we have
$ECE_i = \int_{A_i} \ (c_i - h_i(\vec c)) \cdot p(\vec c)$
$acc_i = \int_{A_i} \ h_i(\vec c) \cdot p(\vec c)$
In many relevant regimes, the DirichletFC can be approximately regarded as sufficiently confident.
This means we can approximate ECE and accuracy as:
$ECE_i \ \lessapprox \ \int_{\tilde A_i} \ (c_i - h_i(\vec c)) \cdot p(\vec c)$
$acc_i \ \lessapprox \ \int_{\tilde A_i} \ h_i(\vec c) \cdot p(\vec c)$
We can explicitly calculate the first part of the ECE:
$ \int_{\tilde A_i} \ c_i \cdot p(\vec c) = \frac{\alpha_i}{\alpha_0}
\left(1 - (\alpha_0-\alpha_i) \ \beta(1/2;\ \alpha_i + 1, \alpha_0-\alpha_i) \ \binom{\alpha_0}{\alpha_i} \right)$
As expected, when $\alpha_i \rightarrow \alpha_0$, this expression goes to one
The second part depends on the simplex automorphism $h$.
We can sort of compute it for the RestrictedPowerAut and for some MaxComponentSimplexAut.
However, both transforms seem to be rather on the pathological side of things...
```
print("mostly underestimating first two classes with RestrictedPowerSimplexAut")
transform = RestrictedPowerSimplexAut(np.array([2, 4]))
dirichlet_fc.set_simplex_automorphism(transform)
eval_stats = EvalStats(*dirichlet_fc.get_sample_arrays(n_samples))
print(f"Accuracy is {eval_stats.accuracy()}")
print(f"ECE is {eval_stats.expected_calibration_error()}")
print("Theoretical approximation of ECE")
print(eval_stats.expected_confidence() - eval_stats.accuracy())
eval_stats.plot_reliability_curves([0, 1, 2, EvalStats.TOP_CLASS_LABEL], display_weights=True)
# theoretical_acc = compute_accuracy(dirichlet_fc)[0]
# theoretical_ece = compute_ECE(dirichlet_fc)[0]
# print(f"{theoretical_acc=} , {theoretical_ece=}")
print(f"""
NOTE: here the ECE completely fails to converge to it's true, continuous value.
This is probably due to the binning-variance, see plots below with 500 bins.
The sharp peak in weights at the end certainly does not help convergence either.
""")
eval_stats.set_bins(500)
eval_stats.plot_reliability_curves([EvalStats.TOP_CLASS_LABEL], display_weights=True)
```
## The Calibration Game
Below are potential 5-classes classifiers that we will use in the calibration game.
They all have roughly the same accuracy but very differing ECEs, corresponding to
different difficulty settings for the game.
```
n_classes = 5
n_samples = 500000
print("hardest setting: accuracy 80, ECE 18")
exponents = np.array([0.05, 0.4, 0.1, 0.2, 0.1]) * 2/3
alpha = np.ones(5) * 1/150
# exponents = np.ones(5) * 1/5
# alpha = np.ones(5) * 1/45
dirichlet_fc = DirichletFC(n_classes, alpha=alpha)
transform = PowerLawSimplexAut(exponents)
dirichlet_fc.set_simplex_automorphism(transform)
eval_stats = EvalStats(*dirichlet_fc.get_sample_arrays(n_samples), bins=200)
print(f"Accuracy is {eval_stats.accuracy()}")
print(f"ECE is {eval_stats.expected_calibration_error()}")
eval_stats.plot_reliability_curves([0, eval_stats.TOP_CLASS_LABEL], display_weights=True)
print("medium setting: accuracy 80, ECE 10")
exponents = np.array([0.5, 1, 1, 1, 0.5]) * 1/1.8
alpha = np.array([0.5, 2, 3, 4, 5]) * 1/65
n_samples = 300000
n_classes = 5
dirichlet_fc = DirichletFC(n_classes, alpha=alpha)
transform = PowerLawSimplexAut(exponents)
dirichlet_fc.set_simplex_automorphism(transform)
eval_stats = EvalStats(*dirichlet_fc.get_sample_arrays(n_samples), bins=200)
print(f"Accuracy is {eval_stats.accuracy()}")
print(f"ECE is {eval_stats.expected_calibration_error()}")
eval_stats.plot_reliability_curves([4, eval_stats.TOP_CLASS_LABEL], display_weights=True)
print("mostly underestimating all classes (starting at 1/n_classes)")
# accuracy 80, ECE 0
alpha = np.array([1, 2, 3, 2, 3]) * 1/19
n_samples = 300000
n_classes = 5
dirichlet_fc = DirichletFC(n_classes, alpha=alpha)
eval_stats = EvalStats(*dirichlet_fc.get_sample_arrays(n_samples))
print(f"Accuracy is {eval_stats.accuracy()}")
print(f"ECE is {eval_stats.expected_calibration_error()}")
eval_stats.plot_reliability_curves([4, eval_stats.TOP_CLASS_LABEL], display_weights=True)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_01_5_python_functional.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# T81-558: Applications of Deep Neural Networks
**Module 1: Python Preliminaries**
* Instructor: [Jeff Heaton](https://sites.wustl.edu/jeffheaton/), McKelvey School of Engineering, [Washington University in St. Louis](https://engineering.wustl.edu/Programs/Pages/default.aspx)
* For more information visit the [class website](https://sites.wustl.edu/jeffheaton/t81-558/).
# Module 1 Material
* Part 1.1: Course Overview [[Video]](https://www.youtube.com/watch?v=Rqq-UnVXtMg&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_01_1_overview.ipynb)
* Part 1.2: Introduction to Python [[Video]](https://www.youtube.com/watch?v=czq5d53vKvo&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_01_2_intro_python.ipynb)
* Part 1.3: Python Lists, Dictionaries, Sets and JSON [[Video]](https://www.youtube.com/watch?v=kcGx2I5akSs&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_01_3_python_collections.ipynb)
* Part 1.4: File Handling [[Video]](https://www.youtube.com/watch?v=FSuSLCMgCZc&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_01_4_python_files.ipynb)
* **Part 1.5: Functions, Lambdas, and Map/Reduce** [[Video]](https://www.youtube.com/watch?v=jQH1ZCSj6Ng&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_01_5_python_functional.ipynb)
# Google CoLab Instructions
The following code ensures that Google CoLab is running the correct version of TensorFlow.
```
try:
from google.colab import drive
%tensorflow_version 2.x
COLAB = True
print("Note: using Google CoLab")
except:
print("Note: not using Google CoLab")
COLAB = False
```
# Part 1.5: Functions, Lambdas, and Map/Reduce
Functions, **lambdas**, and **map/reduce** can allow you to process your data in advanced ways. We will introduce these techniques here and expand on them in the next module, which will discuss Pandas.
Function parameters can be named or unnamed in Python. Default values can also be used. Consider the following function.
```
def say_hello(speaker, person_to_greet, greeting = "Hello"):
print(f'{greeting} {person_to_greet}, this is {speaker}.')
say_hello('Jeff', "John")
say_hello('Jeff', "John", "Goodbye")
say_hello(speaker='Jeff', person_to_greet="John", greeting = "Goodbye")
```
A function is a way to capture code that is commonly executed. Consider the following function that can be used to trim white space from a string capitalize the first letter.
```
def process_string(str):
t = str.strip()
return t[0].upper()+t[1:]
```
This function can now be called quite easily.
```
str = process_string(" hello ")
print(f'"{str}"')
```
Python's **map** is a very useful function that is provided in many different programming languages. The **map** function takes a **list** and applies a function to each member of the **list** and returns a second **list** that is the same size as the first.
```
l = [' apple ', 'pear ', 'orange', 'pine apple ']
list(map(process_string, l))
```
### Map
The **map** function is very similar to the Python **comprehension** that we previously explored. The following **comprehension** accomplishes the same task as the previous call to **map**.
```
l = [' apple ', 'pear ', 'orange', 'pine apple ']
l2 = [process_string(x) for x in l]
print(l2)
```
The choice of using a **map** function or **comprehension** is up to the programmer. I tend to prefer **map** since it is so common in other programming languages.
### Filter
While a **map function** always creates a new **list** of the same size as the original, the **filter** function creates a potentially smaller **list**.
```
def greater_than_five(x):
return x>5
l = [ 1, 10, 20, 3, -2, 0]
l2 = list(filter(greater_than_five, l))
print(l2)
```
### Lambda
It might seem somewhat tedious to have to create an entire function just to check to see if a value is greater than 5. A **lambda** saves you this effort. A lambda is essentially an unnamed function.
```
l = [ 1, 10, 20, 3, -2, 0]
l2 = list(filter(lambda x: x>5, l))
print(l2)
```
### Reduce
Finally, we will make use of **reduce**. Like **filter** and **map** the **reduce** function also works on a **list**. However, the result of the **reduce** is a single value. Consider if you wanted to sum the **values** of a **list**. The sum is implemented by a **lambda**.
```
from functools import reduce
l = [ 1, 10, 20, 3, -2, 0]
result = reduce(lambda x,y: x+y,l)
print(result)
```
| github_jupyter |
```
from __future__ import print_function
import os
import pandas as pd
%matplotlib inline
from matplotlib import pyplot as plt
#Set current dir and work relative to it
os.chdir('D:/Practical Time Series')
#Read dataset into a pandas.DataFrame
beer_df = pd.read_csv('datasets/quarterly-beer-production-in-aus-March 1956-June 1994.csv')
beer_df.index = beer_df['Quarter']
#Display shape of the dataset
print('Shape of the dataframe:', beer_df.shape)
#Show top 10 rows
beer_df.head(10)
#Rename the 2nd column
beer_df.rename(columns={'Quarterly beer production in Australia: megalitres. March 1956 ? June 1994':
'Beer_Prod'
},
inplace=True
)
#Remove missing values
missing = (pd.isnull(beer_df['Quarter'])) | (pd.isnull(beer_df['Beer_Prod']))
print('Number of rows with at least one missing values:', missing.sum())
beer_df = beer_df.loc[~missing, :]
print('Shape after removing missing values:', beer_df.shape)
#In order to remove seasonal patterns let us calculate 2X4 quarter moving average
MA4 = beer_df['Beer_Prod'].rolling(window=4).mean()
TwoXMA4 = MA4.rolling(window=2).mean()
TwoXMA4 = TwoXMA4.loc[~pd.isnull(TwoXMA4)]
#Let's plot the original time series and the seasonal moving averages
fig = plt.figure(figsize=(5.5, 5.5))
ax = fig.add_subplot(1,1,1)
beer_df['Beer_Prod'].plot(ax=ax, color='b', linestyle='-')
TwoXMA4.plot(ax=ax, color='r', linestyle='-')
plt.xticks(rotation=60)
ax.set_title('Quaterly Beer Production between in Australia and 2X4 quarter MA')
plt.savefig('plots/ch2/B07887_02_16.png', format='png', dpi=300)
#Let's compute the residuals after removing the trend
residuals = beer_df['Beer_Prod']-TwoXMA4
residuals = residuals.loc[~pd.isnull(residuals)]
#Let's plot the residuals
fig = plt.figure(figsize=(5.5, 5.5))
ax = fig.add_subplot(1,1,1)
residuals.plot(ax=ax, color='b', linestyle='-')
plt.xticks(rotation=60)
ax.set_title('Residuals in Quaterly Beer Production time series')
plt.savefig('plots/ch2/B07887_02_17.png', format='png', dpi=300)
from pandas.plotting import autocorrelation_plot
#Let's plot the autocorrelation function of the residuals
fig = plt.figure(figsize=(5.5, 5.5))
ax = fig.add_subplot(1,1,1)
autocorrelation_plot(residuals, ax=ax)
ax.set_title('ACF of Residuals in Quaterly Beer Production time series')
plt.savefig('plots/ch2/B07887_02_18.png', format='png', dpi=300)
autocorrelation_plot(residuals)
#Let's compute quarterly differecing to remove quaterly seasonality
residuals_qtr_diff = residuals.diff(4)
#Remove null values
residuals_qtr_diff = residuals_qtr_diff.loc[~pd.isnull(residuals_qtr_diff)]
#Let's plot the autocorrelation function of the residuals
fig = plt.figure(figsize=(5.5, 5.5))
ax = fig.add_subplot(1,1,1)
autocorrelation_plot(residuals_qtr_diff, ax=ax)
ax.set_title('ACF of Quaterly Differenced Residuals')
plt.savefig('plots/ch2/B07887_02_19.png', format='png', dpi=300)
```
| github_jupyter |
```
# Dependencies
from bs4 import BeautifulSoup as bs
import requests
import os
from splinter import Browser
import pandas as pd
from webdriver_manager.chrome import ChromeDriverManager
executable_path = {'executable_path': ChromeDriverManager().install()}
browser = Browser('chrome', **executable_path, headless=False)
url = 'https://mars.nasa.gov/news/'
response = requests.get(url)
soup = bs(response.text, 'html.parser')
#print(soup.prettify())
```
## News Titles and paragraph
```
latest_title = soup.find('div', class_="content_title").get_text()
latest_para = soup.find('div', class_="rollover_description_inner").get_text()
print("Title: {}".format(latest_title))
print("Description: {}".format(latest_para))
```
# JPL Mars Space Images - Featured Image
```
#Use splinter to navigate the site
#browser.visit('https://www.jpl.nasa.gov/spaceimages/?search=&category=Mars')
url="https://www.jpl.nasa.gov/images?search=&category=Mars"
response =requests.get(url)
soup = bs(response.text, 'html.parser')
#print(soup.prettify())
#find the image url for the current Featured Mars Image
image = soup.find('div', class_='sm:object-cover object-cover')
for featured in image:
featured_image_url = featured.get('data-src')
print(f"featured_image_url: {featured_image_url}")
```
# Mars Facts
```
#Visit the Mars Facts webpage
mars_facts_url = "https://space-facts.com/mars/"
table = pd.read_html(mars_facts_url)
table[0]
#use Pandas to scrape the table containing facts about the planet including Diameter, Mass, etc.
#type(table)
mars_df = pd.DataFrame(table[0])
mars_df .columns = ["Facts", "Value"]
mars_df .set_index(["Facts"])
mars_df
#Use Pandas to convert the data to a HTML table string.
converted_to_html = mars_df.to_html()
print(converted_to_html)
```
# Mars Hemispheres
```
#Visit the USGS Astrogeology site
url_hem = ('https://astrogeology.usgs.gov/search/results?q=hemisphere+enhanced&k1=target&v1=Mars')
browser.visit(url_hem)
import time
html = browser.html
soup = bs(html, 'html.parser')
#print(soup.prettify())
mars_hemisphere=[]
# loop through the four tags and load the data to the dictionary
links = browser.find_by_css("a.product-item h3")
for i in range (len(links)):
link_dict= {}
browser.find_by_css("a.product-item h3")[i].click()
sample_list = browser.links.find_by_text('Sample').first
link_dict['img_url'] = sample_list['href']
link_dict['title'] = browser.find_by_css("h2.title").text
mars_hemisphere.append(link_dict)
browser.back()
mars_hemisphere
```
| github_jupyter |
<a href="https://colab.research.google.com/github/NeuromatchAcademy/course-content/blob/mrk-W2D1/tutorials/W2D1_BayesianStatistics/student/W2D1_Tutorial4.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# NMA 2020 W2D1 -- (Bonus) Tutorial 4: Bayesian Decision Theory & Cost functions
__Content creators:__ Vincent Valton, Konrad Kording, with help from Matthew Krause
__Content reviewers:__ Matthew Krause, Jesse Livezey, Karolina Stosio, Saeed Salehi
# Tutorial Objectives
*This tutorial is optional! Please do not feel pressured to finish it!*
In the previous tutorials, we investigated the posterior, which describes beliefs based on a combination of current evidence and prior experience. This tutorial focuses on Bayesian Decision Theory, which combines the posterior with **cost functions** that allow us to quantify the potential impact of making a decision or choosing an action based on that posterior. Cost functions are therefore critical for turning probabilities into actions!
In Tutorial 3, we used the mean of the posterior $p(x | \tilde x)$ as a proxy for the response $\hat x$ for the participants. What prompted us to use the mean of the posterior as a **decision rule**? In this tutorial we will see how different common decision rules such as the choosing the mean, median or mode of the posterior distribution correspond to minimizing different cost functions.
In this tutorial, you will
1. Implement three commonly-used cost functions: mean-squared error, absolute error, and zero-one loss
2. Discover the concept of expected loss, and
3. Choose optimal locations on the posterior that minimize these cost functions. You will verify that it these locations can be found analytically as well as empirically.
```
#@title Video 1: Introduction
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id='BV1Tv411q77s', width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
video
```
---
Please execute the cell below to initialize the notebook environment
---
### Setup
```
# Imports
import numpy as np
import matplotlib.pyplot as plt
#@title Figure Settings
import ipywidgets as widgets
plt.style.use("https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/nma.mplstyle")
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
# @title Helper Functions
def my_gaussian(x_points, mu, sigma):
"""Returns un-normalized Gaussian estimated at points `x_points`
DO NOT EDIT THIS FUNCTION !!!
Args :
x_points (numpy array of floats) - points at which the gaussian is evaluated
mu (scalar) - mean of the Gaussian
sigma (scalar) - std of the gaussian
Returns:
(numpy array of floats): un-normalized Gaussian (i.e. without constant) evaluated at `x`
"""
return np.exp(-(x_points-mu)**2/(2*sigma**2))
def visualize_loss_functions(mse=None, abse=None, zero_one=None):
"""Visualize loss functions
Args:
- mse (func) that returns mean-squared error
- abse: (func) that returns absolute_error
- zero_one: (func) that returns zero-one loss
All functions should be of the form f(x, x_hats). See Exercise #1.
Returns:
None
"""
x = np.arange(-3, 3.25, 0.25)
fig, ax = plt.subplots(1)
if mse is not None:
ax.plot(x, mse(0, x), linewidth=2, label="Mean Squared Error")
if abse is not None:
ax.plot(x, abse(0, x), linewidth=2, label="Absolute Error")
if zero_one_loss is not None:
ax.plot(x, zero_one_loss(0, x), linewidth=2, label="Zero-One Loss")
ax.set_ylabel('Cost')
ax.set_xlabel('Predicted Value ($\hat{x}$)')
ax.set_title("Loss when the true value $x$=0")
ax.legend()
plt.show()
def moments_myfunc(x_points, function):
"""Returns the mean, median and mode of an arbitrary function
DO NOT EDIT THIS FUNCTION !!!
Args :
x_points (numpy array of floats) - x-axis values
function (numpy array of floats) - y-axis values of the function evaluated at `x_points`
Returns:
(tuple of 3 scalars): mean, median, mode
"""
# Calc mode of an arbitrary function
mode = x_points[np.argmax(function)]
# Calc mean of an arbitrary function
mean = np.sum(x_points * function)
# Calc median of an arbitrary function
cdf_function = np.zeros_like(x_points)
accumulator = 0
for i in np.arange(x.shape[0]):
accumulator = accumulator + posterior[i]
cdf_function[i] = accumulator
idx = np.argmin(np.abs(cdf_function - 0.5))
median = x_points[idx]
return mean, median, mode
def loss_plot(x, loss, min_loss, loss_label, show=False, ax=None):
if not ax:
fig, ax = plt.subplots()
ax.plot(x, loss, '-r', linewidth=2, label=loss_label)
ax.axvline(min_loss, ls='dashed', color='red', label='Minimum')
ax.set_ylabel('Expected Loss')
ax.set_xlabel('Orientation (Degrees)')
ax.legend()
if show:
plt.show()
def loss_plot_subfigures(x,
MSEloss, min_MSEloss, loss_MSElabel,
ABSEloss, min_ABSEloss, loss_ABSElabel,
ZeroOneloss, min_01loss, loss_01label):
fig_w, fig_h = plt.rcParams.get('figure.figsize')
fig, ax = plt.subplots(nrows=2, ncols=3, figsize=(fig_w*2, fig_h*2), sharex=True)
ax[0, 0].plot(x, MSEloss, '-r', linewidth=2, label=loss_MSElabel)
ax[0, 0].axvline(min_MSEloss, ls='dashed', color='red', label='Minimum')
ax[0, 0].set_ylabel('Expected Loss')
ax[0, 0].set_xlabel('Orientation (Degrees)')
ax[0, 0].set_title("Mean Squared Error")
ax[0, 0].legend()
pmoments_plot(x, posterior, ax=ax[1,0])
ax[0, 1].plot(x, ABSEloss, '-b', linewidth=2, label=loss_ABSElabel)
ax[0, 1].axvline(min_ABSEloss, ls='dashdot', color='blue', label='Minimum')
ax[0, 1].set_ylabel('Expected Loss')
ax[0, 1].set_xlabel('Orientation (Degrees)')
ax[0, 1].set_title("Absolute Error")
ax[0, 1].legend()
pmoments_plot(x, posterior, ax=ax[1,1])
ax[0, 2].plot(x, ZeroOneloss, '-g', linewidth=2, label=loss_01label)
ax[0, 2].axvline(min_01loss, ls='dotted', color='green', label='Minimum')
ax[0, 2].set_ylabel('Expected Loss')
ax[0, 2].set_xlabel('Orientation (Degrees)')
ax[0, 2].set_title("0-1 Loss")
ax[0, 2].legend()
pmoments_plot(x, posterior, ax=ax[1,2])
plt.show()
def pmoments_plot(x, posterior,
prior=None, likelihood=None, show=False, ax=None):
if not ax:
fig, ax = plt.subplots()
if prior:
ax.plot(x, prior, '-r', linewidth=2, label='Prior')
if likelihood:
ax.plot(x, likelihood, '-b', linewidth=2, label='Likelihood')
ax.plot(x, posterior, '-g', linewidth=4, label='Posterior')
mean, median, mode = moments_myfunc(x, posterior)
ax.axvline(mean, ls='dashed', color='red', label='Mean')
ax.axvline(median, ls='dashdot', color='blue', label='Median')
ax.axvline(mode, ls='dotted', color='green', label='Mode')
ax.set_ylabel('Probability')
ax.set_xlabel('Orientation (Degrees)')
ax.legend()
if show:
plt.show()
def generate_example_pdfs():
"""Generate example probability distributions as in T2"""
x=np.arange(-5, 5, 0.01)
prior_mean = 0
prior_sigma1 = .5
prior_sigma2 = 3
prior1 = my_gaussian(x, prior_mean, prior_sigma1)
prior2 = my_gaussian(x, prior_mean, prior_sigma2)
alpha = 0.05
prior_combined = (1-alpha) * prior1 + (alpha * prior2)
prior_combined = prior_combined / np.sum(prior_combined)
likelihood_mean = -2.7
likelihood_sigma = 1
likelihood = my_gaussian(x, likelihood_mean, likelihood_sigma)
likelihood = likelihood / np.sum(likelihood)
posterior = prior_combined * likelihood
posterior = posterior / np.sum(posterior)
return x, prior_combined, likelihood, posterior
def plot_posterior_components(x, prior, likelihood, posterior):
with plt.xkcd():
fig = plt.figure()
plt.plot(x, prior, '-r', linewidth=2, label='Prior')
plt.plot(x, likelihood, '-b', linewidth=2, label='Likelihood')
plt.plot(x, posterior, '-g', linewidth=4, label='Posterior')
plt.legend()
plt.title('Sample Output')
plt.show()
```
### The Posterior Distribution
This notebook will use a model similar to the puppet & puppeteer sound experiment developed in Tutorial 2, but with different probabilities for $p_{common}$, $p_{independent}$, $\sigma_{common}$ and $\sigma_{independent}$. Specifically, our model will consist of these components, combined according to Bayes' rule:
$$
\begin{eqnarray}
\textrm{Prior} &=& \begin{cases} \mathcal{N_{common}}(0, 0.5) & 95\% \textrm{ weight}\\
\mathcal{N_{independent}}(0, 3.0) & 5\% \textrm{ weight} \\
\end{cases}\\\\
\textrm{Likelihood} &= &\mathcal{N}(-2.7, 1.0)
\end{eqnarray}
$$
We will use this posterior as an an example through this notebook. Please run the cell below to import and plot the model. You do not need to edit anything. These parameter values were deliberately chosen for illustration purposes: there is nothing intrinsically special about them, but they make several of the exercises easier.
```
x, prior, likelihood, posterior = generate_example_pdfs()
plot_posterior_components(x, prior, likelihood, posterior)
```
# Section 1: The Cost Functions
Next, we will implement the cost functions.
A cost function determines the "cost" (or penalty) of estimating $\hat{x}$ when the true or correct quantity is really $x$ (this is essentially the cost of the error between the true stimulus value: $x$ and our estimate: $\hat x$ -- Note that the error can be defined in different ways):
$$\begin{eqnarray}
\textrm{Mean Squared Error} &=& (x - \hat{x})^2 \\
\textrm{Absolute Error} &=& \big|x - \hat{x}\big| \\
\textrm{Zero-One Loss} &=& \begin{cases}
0,& \text{if } x = \hat{x} \\
1, & \text{otherwise}
\end{cases}
\end{eqnarray}
$$
In the cell below, fill in the body of these cost function. Each function should take one single value for $x$ (the true stimulus value : $x$) and one or more possible value estimates: $\hat{x}$.
Return an array containing the costs associated with predicting $\hat{x}$ when the true value is $x$. Once you have written all three functions, uncomment the final line to visulize your results.
_Hint:_ These functions are easy to write (1 line each!) but be sure *all* three functions return arrays of `np.float` rather than another data type.
```
def mse(x, x_hats):
"""Mean-squared error cost function
Args:
x (scalar): One true value of $x$
x_hats (scalar or ndarray): Estimate of x
Returns:
same shape/type as x_hats): MSE costs associated with
predicting x_hats instead of x$
"""
##############################################################################
# Complete the MSE cost function
#
### Comment out the line below to test your function
raise NotImplementedError("You need to complete the MSE cost function!")
##############################################################################
my_mse = ...
return my_mse
def abs_err(x, x_hats):
"""Absolute error cost function
Args:
x (scalar): One true value of $x$
x_hats (scalar or ndarray): Estimate of x
Returns:
(same shape/type as x_hats): absolute error costs associated with
predicting x_hats instead of x$
"""
##############################################################################
# Complete the absolute error cost function
#
### Comment out the line below to test your function
raise NotImplementedError("You need to complete the absolute error function!")
##############################################################################
my_abs_err = ...
return my_abs_err
def zero_one_loss(x, x_hats):
"""Zero-One loss cost function
Args:
x (scalar): One true value of $x$
x_hats (scalar or ndarray): Estimate of x
Returns:
(same shape/type as x_hats) of the 0-1 Loss costs associated with predicting x_hat instead of x
"""
##############################################################################
# Complete the zero-one loss cost function
#
### Comment out the line below to test your function
raise NotImplementedError("You need to complete the 0-1 loss cost function!")
##############################################################################
my_zero_one_loss = ...
return my_zero_one_loss
## When you are done with the functions above, uncomment the line below to
## visualize them
# visualize_loss_functions(mse, abs_err, zero_one_loss)
```
[*Click for solution*](https://github.com/erlichlab/course-content/tree/master//tutorials/W2D1_BayesianStatistics/solutions/W2D1_Tutorial4_Solution_0c5a484f.py)
*Example output:*
<img alt='Solution hint' align='left' width=416 height=272 src=https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/tutorials/W2D1_BayesianStatistics/static/W2D1_Tutorial4_Solution_0c5a484f_2.png>
# Section 2: Expected Loss
```
#@title Video 2: Expected Loss
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id='BV1av411q7iK', width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
video
```
A posterior distribution tells us about the confidence or credibility we assign to different choices. A cost function describes the penalty we incur when choosing an incorrect option. These concepts can be combined into an *expected loss* function. Expected loss is defined as:
$$
\begin{eqnarray}
\mathbb{E}[\text{Loss} | \hat{x}] = \int L[\hat{x},x] \odot p(x|\tilde{x}) dx
\end{eqnarray}
$$
where $L[ \hat{x}, x]$ is the loss function, $p(x|\tilde{x})$ is the posterior, and $\odot$ represents the [Hadamard Product](https://en.wikipedia.org/wiki/Hadamard_product_(matrices)) (i.e., elementwise multiplication), and $\mathbb{E}[\text{Loss} | \hat{x}]$ is the expected loss.
In this exercise, we will calculate the expected loss for the: means-squared error, the absolute error, and the zero-one loss over our bimodal posterior $p(x | \tilde x)$.
**Suggestions:**
* We already pre-completed the code (commented-out) to calculate the mean-squared error, absolute error, and zero-one loss between $x$ and an estimate $\hat x$ using the functions you created in exercise 1
* Calculate the expected loss ($\mathbb{E}[MSE Loss]$) using your posterior (imported above as `posterior`) & each of the loss functions described above (MSELoss, ABSELoss, and Zero-oneLoss).
* Find the x position that minimizes the expected loss for each cost function and plot them using the `loss_plot` function provided (commented-out)
## Exercise 2: Finding the expected loss empirically via integration
```
def expected_loss_calculation(x, posterior):
ExpectedLoss_MSE = np.zeros_like(x)
ExpectedLoss_ABSE = np.zeros_like(x)
ExpectedLoss_01 = np.zeros_like(x)
for idx in np.arange(x.shape[0]):
estimate = x[idx]
###################################################################
## Insert code below to find the expected loss under each loss function
##
## remove the raise when the function is complete
raise NotImplementedError("Calculate the expected loss over all x values!")
###################################################################
MSELoss = mse(estimate, x)
ExpectedLoss_MSE[idx] = ...
ABSELoss = abs_err(estimate, x)
ExpectedLoss_ABSE[idx] = ...
ZeroOneLoss = zero_one_loss(estimate, x)
ExpectedLoss_01[idx] = ...
###################################################################
## Now, find the `x` location that minimizes expected loss
##
## remove the raise when the function is complete
raise NotImplementedError("Finish the Expected Loss calculation")
###################################################################
min_MSE = ...
min_ABSE = ...
min_01 = ...
return (ExpectedLoss_MSE, ExpectedLoss_ABSE, ExpectedLoss_01,
min_MSE, min_ABSE, min_01)
## Uncomment the lines below to plot the expected loss as a function of the estimates
#ExpectedLoss_MSE, ExpectedLoss_ABSE, ExpectedLoss_01, min_MSE, min_ABSE, min_01 = expected_loss_calculation(x, posterior)
#loss_plot(x, ExpectedLoss_MSE, min_MSE, f"Mean Squared Error = {min_MSE:.2f}")
#loss_plot(x, ExpectedLoss_ABSE, min_ABSE, f"Absolute Error = {min_ABSE:.2f}")
#loss_plot(x, ExpectedLoss_01, min_01, f"Zero-One Error = {min_01:.2f}")
```
[*Click for solution*](https://github.com/erlichlab/course-content/tree/master//tutorials/W2D1_BayesianStatistics/solutions/W2D1_Tutorial4_Solution_ee8aff37.py)
*Example output:*
<img alt='Solution hint' align='left' width=424 height=280 src=https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/tutorials/W2D1_BayesianStatistics/static/W2D1_Tutorial4_Solution_ee8aff37_0.png>
<img alt='Solution hint' align='left' width=424 height=280 src=https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/tutorials/W2D1_BayesianStatistics/static/W2D1_Tutorial4_Solution_ee8aff37_1.png>
<img alt='Solution hint' align='left' width=424 height=280 src=https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/tutorials/W2D1_BayesianStatistics/static/W2D1_Tutorial4_Solution_ee8aff37_2.png>
# Section 3: Analytical Solutions
```
#@title Video 3: Analytical Solutions
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id='BV1aa4y1a7Ex', width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
video
```
In the previous exercise, we found the minimum expected loss via brute-force: we searched over all possible values of $x$ and found the one that minimized each of our loss functions. This is feasible for our small toy example, but can quickly become intractable.
Fortunately, the three loss functions examined in this tutorial have are minimized at specific points on the posterior, corresponding to the itss mean, median, and mode. To verify this property, we have replotted the loss functions from Exercise 2 below, with the posterior on the same scale beneath. The mean, median, and mode are marked on the posterior.
Which loss form corresponds to each summary statistics?
```
loss_plot_subfigures(x,
ExpectedLoss_MSE, min_MSE, f"Mean Squared Error = {min_MSE:.2f}",
ExpectedLoss_ABSE, min_ABSE, f"Absolute Error = {min_ABSE:.2f}",
ExpectedLoss_01, min_01, f"Zero-One Error = {min_01:.2f}")
```
[*Click for solution*](https://github.com/erlichlab/course-content/tree/master//tutorials/W2D1_BayesianStatistics/solutions/W2D1_Tutorial4_Solution_0716b099.py)
# Section 4: Conclusion
```
#@title Video 4: Outro
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id='BV1kh411o7cu', width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
video
```
In this tutorial, we learned about three kinds of cost functions: mean-squared error, absolute error, and zero-one loss. We used expected loss to quantify the results of making a decision, and showed that optimizing under different cost functions led us to choose different locations on the posterior. Finally, we found that these optimal locations can be identified analytically, sparing us from a brute-force search.
Here are some additional questions to ponder:
* Suppose your professor offered to grade your work with a zero-one loss or mean square error.
* When might you choose each?
* Which would be easier to learn from?
* All of the loss functions we considered are symmetrical. Are there situations where an asymmetrical loss function might make sense? How about a negative one?
| github_jupyter |
```
import math
import json
import pandas as pd
import numpy as np
from ete3 import Tree
from Bio import SeqIO
from Bio.Seq import Seq
from Bio.SeqRecord import SeqRecord
import matplotlib.pyplot as plt
import seaborn as sns
def get_numdate(input_file_branch_lengths, node_name):
with open(input_file_branch_lengths, "r") as bl_handle:
branch_lengths = json.load(bl_handle)
for node, v in branch_lengths['nodes'].items():
if node == node_name:
return v['numdate']
def get_tmrca(input_file_branch_lengths, tree, strains):
tree_strains = []
for leaf in tree:
if leaf.name in strains:
tree_strains.append(leaf.name)
common_ancestor = tree.get_common_ancestor(tree_strains)
common_ancestor_numdate = get_numdate(input_file_branch_lengths, common_ancestor.name)
times_to_common_ancestor = []
for leaf in tree_strains:
leaf_numdate = get_numdate(input_file_branch_lengths, leaf)
pairwise_distance = leaf_numdate - common_ancestor_numdate
times_to_common_ancestor.append(pairwise_distance)
tmrca = sum(times_to_common_ancestor)/len(times_to_common_ancestor)
return tmrca
def separate_clades(cov, gene):
if cov == 'hku1':
clade_file = '../'+str(cov)+'/results/clades_full.json'
else:
if gene =='spike' or gene == 's1' or gene == 's2':
clade_file = '../'+str(cov)+'/results/clades_spike.json'
else:
clade_file = '../'+str(cov)+'/results/clades_full.json'
clade_lists = []
with open(clade_file, "r") as clade_handle:
clades = json.load(clade_handle)
for node, v in clades['nodes'].items():
if 'NODE' not in node:
clade_lists.append({'clade':v['clade_membership'],
'strain':node})
clade_df = pd.DataFrame(clade_lists)
return clade_df
def tmrca_by_clade(cov, gene, window, clade, min_seqs, year_max=None, year_min=None):
input_file_alignment = '../'+str(cov)+'/results/aligned_'+str(cov)+'_'+str(gene)+'.fasta'
input_file_branch_lengths = '../'+str(cov)+'/results/branch_lengths_'+str(cov)+'_'+str(gene)+'.json'
input_tree = Tree('../'+str(cov)+'/results/tree_'+str(cov)+'_'+str(gene)+'.nwk', format=1)
metafile = '../'+str(cov)+'/results/metadata_'+str(cov)+'_'+str(gene)+'.tsv'
#Subset data based on time windows
meta = pd.read_csv(metafile, sep = '\t')
meta.drop(meta[meta['date']=='?'].index, inplace=True)
meta.dropna(subset=['date'], inplace=True)
meta['year'] = meta['date'].str[:4].astype('int')
if year_max:
meta.drop(meta[meta['year']>year_max].index, inplace=True)
if year_min:
meta.drop(meta[meta['year']<year_min].index, inplace=True)
date_range = meta['year'].max() - meta['year'].min()
if clade!= None:
clade_df = separate_clades(cov, gene)
meta = meta.merge(clade_df, on='strain')
meta.drop(meta[meta['clade']!=clade].index, inplace=True)
#Group viruses by time windows
virus_time_subset = {}
if window == 'all':
years = str(meta['year'].min()) + '-' + str(meta['year'].max())
virus_time_subset[years] = meta['strain'].tolist()
else:
date_window_start = meta['year'].min()
date_window_end = meta['year'].min() + window
while date_window_end <= meta['year'].max():
years = str(date_window_start) + '-' + str(date_window_end)
strains = meta[(meta['year']>=date_window_start) & (meta['year']<date_window_end)]['strain'].tolist()
virus_time_subset[years] = strains
#sliding window
date_window_end += 1
date_window_start += 1
#initiate lists to record all time windows
year_windows = []
window_midpoint = []
seqs_in_window = []
tmrcas = []
for years, subset_viruses in virus_time_subset.items():
#don't use windows with fewer than min_seqs
if len(subset_viruses) >= min_seqs:
year_windows.append(years)
window_start = int(years[0:4])
window_end = int(years[-4:])
window_midpoint.append((window_start + window_end)/2)
seqs_in_window.append(len(subset_viruses))
tmrca = get_tmrca(input_file_branch_lengths, input_tree, subset_viruses)
tmrcas.append(tmrca)
return year_windows, window_midpoint, seqs_in_window, tmrcas
def plot_tmrca(cov, genes, window, clade, min_seqs, year_max=None, year_min=None):
data_to_plot = []
for gene in genes:
(year_windows, window_midpoint,seqs_in_window, tmrcas) = tmrca_by_clade(cov, gene, window,
clade, min_seqs,
year_max, year_min)
for year in range(len(window_midpoint)):
data_to_plot.append({'cov_clade': str(cov)+str(clade), 'gene': gene,
'year': window_midpoint[year],
'tmrca': tmrcas[year]})
df_to_plot = pd.DataFrame(data_to_plot)
color_map = {'oc43A': '#CB4335', 'oc43B':'#FF9A00', '229eNone': '#2E86C1',
'nl63A': '#009888', 'nl63B': '#87C735', 'hku1A': '#7c5295', 'hku1B': '#b491c8'}
g = sns.FacetGrid(df_to_plot, col='gene', col_wrap=2, hue='cov_clade', height=6, aspect=1,
palette=color_map, sharey=True, sharex=False)
g.map(sns.pointplot, 'year', 'tmrca', ci=None)
plot_tmrca('oc43', ['s1', 's2', 'spike', 'replicase1ab'], 2, 'A', 3, year_max=None, year_min=None)
plot_tmrca('229e', ['s1', 's2', 'spike', 'replicase1ab'], 3, None, 3, year_max=None, year_min=None)
plot_tmrca('nl63', ['s1', 's2', 'spike', 'replicase1ab'], 3, 'B', 3, year_max=None, year_min=None)
plot_tmrca('oc43', ['s1', 's2', 'spike', 'replicase1ab'], 3, 'A', 3, year_max=None, year_min=None)
plot_tmrca('oc43', ['s1', 's2', 'spike', 'replicase1ab'], 5, 'A', 3, year_max=None, year_min=None)
```
| github_jupyter |
```
import re
import pandas as pd
import numpy as np
import nltk
from nltk.corpus import stopwords
from nltk.stem import SnowballStemmer
from nltk.stem import WordNetLemmatizer
import spacy
from spacy.tokenizer import Tokenizer
import torch
import torch.nn as nn
from torchtext import data, legacy
from torchtext.vocab import Vectors, GloVe
import tqdm
nltk.download('stopwords')
nltk.download('wordnet')
train = pd.read_csv('../train.csv')
test = pd.read_csv('../test.csv')
smaple = pd.read_csv('../sample_submission.csv')
print(train.shape)
print(test.shape)
```
# Exploration
```
train.head()
test.head()
```
# Data Cleaning
```
print(train.info())
print("=================")
print(test.info())
train.drop(['location', 'keyword'], axis=1, inplace=True)
test.drop(['location', 'keyword'], axis=1, inplace=True)
train.shape, test.shape
for i in range(50):
print(train['text'][i])
print("------------")
puncts = [',', '.', '"', ':', ')', '(', '-', '!', '?', '|', ';', "'", '$', '&', '/',
'[', ']', '>', '%', '=', '#', '*', '+', '\\', '•', '~', '@', '£', '·', '_',
'{', '}', '©', '^', '®', '`', '<', '→', '°', '€', '™', '›', '♥', '←', '×',
'§', '″', '′', 'Â', '█', '½', 'à', '…', '“', '★', '”', '–', '●', 'â', '►', '−',
'¢', '²', '¬', '░', '¶', '↑', '±', '¿', '▾', '═', '¦', '║', '―', '¥', '▓', '—',
'‹', '─', '▒', ':', '¼', '⊕', '▼', '▪', '†', '■', '’', '▀', '¨', '▄', '♫', '☆',
'é', '¯', '♦', '¤', '▲', 'è', '¸', '¾', 'Ã', '⋅', '‘', '∞', '∙', ')', '↓', '、',
'│', '(', '»', ',', '♪', '╩', '╚', '³', '・', '╦', '╣', '╔', '╗', '▬', '❤', 'ï',
'Ø', '¹', '≤', '‡', '√', ]
def clean_text(x):
x = str(x)
for punct in puncts:
x = x.replace(punct, f' {punct} ')
return x
def clean_numbers(x):
x = re.sub('[0-9]{5, }', '#####', x)
x = re.sub('[0-9]{4}', '####', x)
x = re.sub('[0-9]{3}', '###', x)
x = re.sub('[0-9]{2}', '##', x)
return x
mispell_dict = {"aren't": "are not", "can't": "cannot",
"couldn't": "could not", "didn't": "did not",
"doesn't": "does not", "don't": "do not",
"hadn't": "had not", "hasn't": "has not",
"haven't": "have not", "he'd": "he would",
"he'll": "he will", "he's": "he is",
"i'd": "I would", "i'd": "I had", "i'll":
"I will", "i'm" : "I am", "isn't": "is not",
"it's": "it is", "it'll": "it will",
"i've" : "I have", "let's": "let us",
"mightn't": "might not", "mustn't": "must not",
"shan't" : "shall not", "she'd": "she would",
"she'll": "she will", "she's": "she is",
"shouldn't": "should not", "that's": "that is",
"there's": "there is","they'd": "they would",
"they'll": "they will", "they're": "they are",
"they've": "they have", "we'd": "we would",
"we're": "we are", "weren't": "were not",
"we've": "we have", "what'll": "what will",
"what're": "what are", "what's": "what is",
"what've": "what have", "where's": "where is",
"who'd": "who would", "who'll": "who will",
"who're": "who are", "who's": "who is",
"who've": "who have", "won't": "will not",
"wouldn't" : "would not", "you'd": "you would",
"you'll": "you will", "you're": "you are",
"you've": "you have", "'re": " are",
"wasn't": "was not", "we'll": " will",
"didn't": "did not", "tryin'": "trying",
"colour": "color", "centre": "center",
"didnt": "did not", "doesnt": "does not",
"isnt": "is not", "shouldnt": "should not",
"favourite": "favorite", "travelling": "traveling",
"counselling": "counseling", "theatre": "theater",
"cancelled": "canceled", "labour": "labor",
"organisation": "organization", "wwii": "world war 2",
"citicise": "criticize", "instagram": "social medium",
"whatsapp": "social medium", "snapchat": "social medium"}
def get_mispell(mispell_dict):
mispell_re = re.compile('(%s)' % '|'.join(mispell_dict.keys()))
return mispell_dict, mispell_re
def replace_typical_misspell(text):
def replace(match):
return mispellings[match.group(0)]
return mispellings_re.sub(replace, text)
mispellings, mispellings_re = get_mispell(mispell_dict)
mispellings, mispellings_re
def remove_emoji(sentence):
emoji_pattern = re.compile("["
u"\U0001F600-\U0001F64F" # emoticons
u"\U0001F300-\U0001F5FF" # symbols & pictographs
u"\U0001F680-\U0001F6FF" # transport & map symbols
u"\U0001F1E0-\U0001F1FF" # flags (iOS)
u"\U00002702-\U000027B0"
u"\U000024C2-\U0001F251"
"]+", flags=re.UNICODE)
return emoji_pattern.sub(r'', sentence)
def remove_stopwords(sentence):
words = sentence.split()
words = [word for word in words if word not in stopwords.words('english')]
return ' '.join(words)
stemmer = SnowballStemmer('english')
wl = WordNetLemmatizer()
def lemmatize_words(sentence):
words = sentence.split()
words = [wl.lemmatize(word) for word in words]
return ' '.join(words)
# Clean the text
train["text"] = train["text"].apply(lambda x: clean_text(x.lower()))
test["text"] = test["text"].apply(lambda x: clean_text(x.lower()))
# Clean numbers
train["text"] = train["text"].apply(lambda x: clean_numbers(x))
test["text"] = test["text"].apply(lambda x: clean_numbers(x))
# Clean spellings
train["text"] = train["text"].apply(lambda x: replace_typical_misspell(x))
test["text"] = test["text"].apply(lambda x: replace_typical_misspell(x))
# Clear emojis
train["text"] = train["text"].apply(lambda x: remove_emoji(x))
test["text"] = test["text"].apply(lambda x: remove_emoji(x))
# Stopwords
train["text"] = train["text"].apply(lambda x: remove_stopwords(x))
test["text"] = test["text"].apply(lambda x: remove_stopwords(x))
# Lemmatization
train["text"] = train["text"].apply(lambda x: lemmatize_words(x))
test["text"] = test["text"].apply(lambda x: lemmatize_words(x))
for i in range(50):
print(train['text'][i])
print("------------")
print(train.head())
print("\n================================================================\n")
print(test.head())
def get_iterator(dataset, batch_size, train=True,
shuffle=True, repeat=False):
device = torch.device('cuda:0' if torch.cuda.is_available()
else 'cpu')
dataset_iter = legacy.data.Iterator(
dataset, batch_size=batch_size, device=device,
train=train, shuffle=shuffle, repeat=repeat,
sort=False
)
return dataset_iter
def prepare_csv(df_train, df_test, seed=27, val_ratio=0.3):
idx = np.arange(df_train.shape[0])
np.random.seed(seed)
np.random.shuffle(idx)
val_size = int(len(idx) * val_ratio)
if not os.path.exists('cache'):
os.makedirs('cache')
df_train.iloc[idx[val_size:], :][['id', 'target', 'text']].to_csv(
'cache/dataset_train.csv', index=False
)
df_train.iloc[idx[:val_size], :][['id', 'target', 'text']].to_csv(
'cache/dataset_val.csv', index=False
)
df_test[['id', 'text']].to_csv('cache/dataset_test.csv',
index=False)
import logging
from copy import deepcopy
import random
import os
SEED=12
LOGGER = logging.getLogger('tweets_dataset')
def get_dataset(fix_length=100, lower=False, vectors=None):
if vectors is not None:
lower=True
LOGGER.debug('Preparing CSV files...')
prepare_csv(train, test)
TEXT = legacy.data.Field(sequential=True,
lower=True,
include_lengths=True,
batch_first=True,
fix_length=25)
LABEL = legacy.data.Field(use_vocab=True,
sequential=False,
dtype=torch.float16)
ID = legacy.data.Field(use_vocab=False,
sequential=False,
dtype=torch.float16)
LOGGER.debug('Reading train csv files...')
train_temp, val_temp = legacy.data.TabularDataset.splits(
path='cache/', format='csv', skip_header=True,
train='dataset_train.csv', validation='dataset_val.csv',
fields=[
('id', ID),
('target', LABEL),
('text', TEXT)
]
)
LOGGER.debug('Reading test csv file...')
test_temp = legacy.data.TabularDataset(
path='cache/dataset_test.csv', format='csv',
skip_header=True,
fields=[
('id', ID),
('text', TEXT)
]
)
LOGGER.debug('Building vocabulary...')
TEXT.build_vocab(
train_temp, val_temp, test_temp,
max_size=20000,
min_freq=10,
vectors=GloVe(name='6B', dim=300) # We use it for getting vocabulary of words
)
LABEL.build_vocab(
train_temp
)
ID.build_vocab(
train_temp, val_temp, test_temp
)
word_embeddings = TEXT.vocab.vectors
vocab_size = len(TEXT.vocab)
train_iter = get_iterator(train_temp, batch_size=32,
train=True, shuffle=True,
repeat=False)
val_iter = get_iterator(val_temp, batch_size=32,
train=True, shuffle=True,
repeat=False)
test_iter = get_iterator(test_temp, batch_size=32,
train=False, shuffle=False,
repeat=False)
LOGGER.debug('Done preparing the datasets')
return TEXT, vocab_size, word_embeddings, train_iter, val_iter, test_iter
TEXT, vocab_size, word_embeddings, train_iter, val_iter, test_iter = get_dataset()
class LSTMClassifier(torch.nn.Module):
def __init__(self, vocab_size, output_size, embedding_dim, hidden_dim, n_layers, weights):
super(LSTMClassifier, self).__init__()
self.output_size = output_size
self.n_layers = n_layers
self.hidden_dim = hidden_dim
self.word_embeddings = torch.nn.Embedding(vocab_size,
embedding_dim)
self.word_embeddings.weight.data.copy_(weights)
self.dropout_1 = torch.nn.Dropout(0.3)
self.lstm = torch.nn.LSTM(embedding_dim,
hidden_dim,
n_layers,
dropout=0.3,
batch_first=True)
self.dropout_2 = torch.nn.Dropout(0.3)
self.label_layer = torch.nn.Linear(hidden_dim, output_size)
self.act = torch.nn.Sigmoid()
def forward(self, x, hidden):
batch_size = x.size(0)
x = self.word_embeddings(x)
x = self.dropout_1(x)
lstm_out, hidden = self.lstm(x, hidden)
lstm_out = lstm_out.contiguous().view(-1, self.hidden_dim)
out = self.dropout_2(lstm_out)
out = self.label_layer(out)
out = out.view(batch_size, -1, self.output_size)
out = out[:, -1, :]
out = self.act(out)
return out, hidden
def init_hidden(self, batch_size):
weight = next(self.parameters()).data
hidden = (weight.new(self.n_layers, batch_size, self.hidden_dim).zero_().to(device),
weight.new(self.n_layers, batch_size, self.hidden_dim).zero_().to(device))
return hidden
def train_model(model, train_iter, val_iter, optim, loss, num_epochs, batch_size=32):
h = model.init_hidden(batch_size)
clip = 5
val_loss_min = np.Inf
total_train_epoch_loss = list()
total_train_epoch_acc = list()
total_val_epoch_loss = list()
total_val_epoch_acc = list()
device = torch.device('cuda:0' if torch.cuda.is_available()
else 'cpu')
for epoch in range(num_epochs):
model.train()
train_epoch_loss = list()
train_epoch_acc = list()
val_epoch_loss = list()
val_epoch_acc = list()
for idx, batch in enumerate(train_iter):
h = tuple([e.data for e in h])
text = batch.text[0]
target = batch.target
target = target - 1
target = target.type(torch.LongTensor)
text = text.to(device)
target = target.to(device)
optim.zero_grad()
if text.size()[0] is not batch_size:
continue
prediction, h = model(text, h)
loss_train = loss(prediction.squeeze(), target)
loss_train.backward()
num_corrects = (torch.max(prediction, 1)[1].
view(target.size()).data == target.data).float().sum()
acc = 100.0 * num_corrects / len(batch)
train_epoch_loss.append(loss_train.item())
train_epoch_acc.append(acc.item())
torch.nn.utils.clip_grad_norm_(model.parameters(), 5)
optim.step()
print(f'Train Epoch: {epoch}, Training Loss: {np.mean(train_epoch_loss):.4f}, Training Accuracy: {np.mean(train_epoch_acc): .2f}%')
model.eval()
with torch.no_grad():
for idx, batch in enumerate(val_iter):
val_h = tuple([e.data for e in h])
text = batch.text[0]
target = batch.target
target = target - 1
target = target.type(torch.LongTensor)
text = text.to(device)
target = target.to(device)
if text.size()[0] is not batch_size:
continue
prediction, h = model(text, h)
loss_val = loss(prediction.squeeze(), target)
num_corrects = (torch.max(prediction, 1)[1].
view(target.size()).data == target.data).float().sum()
acc = 100.0 * num_corrects / len(batch)
val_epoch_loss.append(loss_val.item())
val_epoch_acc.append(acc.item())
print(f'Vadlidation Epoch: {epoch}, Training Loss: {np.mean(val_epoch_loss):.4f}, Training Accuracy: {np.mean(val_epoch_acc): .2f}%')
if np.mean(val_epoch_loss) <= val_loss_min:
# torch.save(model.state_dict(), 'state_dict.pth')
print('Validation loss decreased ({:.6f} --> {:.6f})'.
format(val_loss_min, np.mean(val_epoch_loss)))
val_loss_min = np.mean(val_epoch_loss)
total_train_epoch_loss.append(np.mean(train_epoch_loss))
total_train_epoch_acc.append(np.mean(train_epoch_acc))
total_val_epoch_loss.append(np.mean(val_epoch_loss))
total_val_epoch_acc.append(np.mean(val_epoch_acc))
return (total_train_epoch_loss, total_train_epoch_acc,
total_val_epoch_loss, total_val_epoch_acc)
lr = 1e-4
batch_size = 32
output_size = 2
hidden_size = 128
embedding_length = 300
model = LSTMClassifier(vocab_size=vocab_size,
output_size=output_size,
embedding_dim=embedding_length,
hidden_dim=hidden_size,
n_layers=2,
weights=word_embeddings
)
device = torch.device('cuda:0' if torch.cuda.is_available()
else 'cpu')
model.to(device)
optim = torch.optim.Adam(model.parameters(), lr=lr)
loss = torch.nn.CrossEntropyLoss()
train_loss, train_acc, val_loss, val_acc = train_model(model=model,
train_iter=train_iter,
val_iter=val_iter,
optim=optim,
loss=loss,
num_epochs=20,
batch_size=batch_size)
results_target = list()
with torch.no_grad():
for batch in test_iter:
for text, idx in zip(batch.text[0], batch.id):
text = text.unsqueeze(0)
res, _ = model(text, hidden=None)
target = np.round(res.cpu().numpy())
results_target.append(target[0][1])
smaple['target'] = list(map(int, results_target))
smaple.to_csv('submission.csv', index=False)
```
| github_jupyter |
[link to live lecture](https://colab.research.google.com/drive/1udp7HIM_Zpvp-LY8vgcPixC8Z6Eza-EH)
## Review from Last lecture
1. Variables
2. Variable Expressions
3. Logic
Save a copy to your drive, and follow along with the lecture!
Jumping right in with Conditional Statements
# Lecture 2 Conditions & Loops
By the end of this lecture, you should feel comfortable with the following
- Writing conditional code blocks (`if`, `ifel` and `else`)
- Begin writing your own while loop and understand the key terms **continue**, **pass**, and **break**
- Confidently using Colab Notebooks
### 2.00 Conditionals
Let's tackle this one part at a time. What does it mean to be a condition? Really, all an `if` is checking is whether the conditional evaluates to `True` or `False`. If the condition is true, then the body of the `if` statement is executed. If the condition is false, the `if` block is skipped. Intuitively, true and false are concepts that make perfect sense to us. But we should take the time to clearly define them in a programming context here.
`True` and `False` are what we call booleans in logic (`bool` for short), and what Python calls them. They are a special variable type with many potential uses; mainly they are used as a way to put a label on the truth of a statement. There are two specifically reserved words for bools in Python, `True` and `False`. Note that these begin with capital letters.
```python
In [1]: type(True)
Out[1]: bool
In [2]: type(False)
Out[2]: bool
```
```
```
In addition, a wide variety of statements can evaluate to booleans. The ones that we will focus on today are the equalities, *equal to* and *not equal to*, and the inequalities, *less than*, *greater than*, *less than or equal to* and *greater than or equal to*. These comparisons are available in Python via `==`, `!=`, `<`, `>`, `<=` and `>=`, respectively.
```python
In [1]: 1 == 2
Out[1]: False
In [2]: 1 != 2
Out[2]: True
In [3]: 1 < 2
Out[3]: True
In [4]: 1 > 2
Out[4]: False
In [5]: 1 <= 2
Out[5]: True
In [6]: 1 >= 2
Out[6]: False
```
```
# Elliott
x = 50
if x >= 25*2:
print('X is greater than or equal to 25*2')
```
### Using the If
Now that we understand conditionals, let's talk about how we can use them with variables to make our programs dynamic. Consider the following code block.
```python
if x > 5:
x += 10
print(x)
```
**Note**: The print function simply pipes the value passed to it to the console.
### 2.1 Your turn
In your own notebook, play around with conditional statements!
```
```
In the above code we don't need to know what the value of `x` is, but we can say that if it's greater than 5, it will come out of the code block 10 greater than before the `if` statement.
From what we know so far, this functionality isn't super useful. So let's quickly go over a way that we can make our Python more flexible. Until now, we've had to hard code any variable or value that we want to use in our program. Python has a built in way to accept input from the user of a program. Let's examine this now. Consider that the following code was stored in a file named `print_number.py`.
```python
x = input('Please enter a number: ')
print(x)
```
If we then ran the script from IPython, we would would see:
```python
In [1]: run print_number.py
Please enter a number:
```
**Note**: The `input()` function accepts character input from the keyboard, printing the message it is passed as a prompt.
We can then type a number followed by enter, and the script will print that number.
```python
In [1]: run print_number.py
Please enter a number: 3
3
```
**Note**: `input()` halts the execution of your script, so nothing will happen until you type something and press enter.
Now that we have a way to get arbitrary input from the user of our program, we can begin to see the full power of the `if`. Let's combine the last two code blocks from above, and say we stored it in a script named `print_number_with_if.py`.
```python
x = int(input('Please enter a number: '))
if x > 5:
x += 10
print(x)
```
**Note**: `input()` actually interprets the input as strings, so we have to manually tell Python to treat the number we pass as an integer with `int()`. We'll talk about strings more next week.
If we then ran the script from IPython as above, let's look at two ways we could interact with it.
```python
In [1]: run print_number_with_if.py
Please enter a number: 3
3
In [2]: run print_number_with_if.py
Please enter a number: 8
18
```
Notice that the first time we run `print_number_with_if` and give it 3, it acts just like `print_number`. However, the second time, when we give it 8, it adds 10 and prints 18. Why did it do this? Because 8 is greater than 5, so our program added 10 to it before it was printed.
This may seem like a trivial example, and therefore, not very exciting. Let me assure you, though, that what you have just learned is amazingly powerful! So congratulations!
### 2.11 Your turn
Using the input command, create an **logic** statement that executes based on the value that you input.
```
```
### 2.12 Building on the If
Ok, so, the `if` is cool. But it seems like, due to it's structure, there are only so many things you can do with it. Let's summarize this with what's known as a *flow diagram*.
You can see that there are two branches created by the `if` statement, one when the condition is true, and the other when it is false. In the former case, the conditional code is executed, and in the latter, the conditional code is ignored. But what if we wanted to check more than one thing (i.e. have more than two branches in our flow diagram)?
Python gives us two ways to do this. One by offering other conditionals, `elif` and `else`, and the other by allowing us to combine conditions with logicals `and`, `or` and `not`.
#### 2.13 Elif and Else
In addition to the `if`, Python provides us with two other statements to build out those logical trees, the `elif` and the `else`. The `elif` is just like the `if` - it accepts a condition to check the truth of and has an indented code block that is executed when that condition evaluates to `True`. The `else` is similar, but it doesn't accept a condition. Instead, it mainly acts as a catch all for any other situation that you don't need to cover with your `if`s and `elif`s. Note that there can only be a single `if` and up to a single `else`, but any number of `elif`s in an `if`-`elif`-`else` block. Let's take a closer look at this in the following code block that we'll store in `if_elif_else.py`.
```python
x = int(input('Please enter a number: '))
if x < 0:
print('You entered a negative number.')
elif x > 0:
print('You entered a positive number.')
else:
print('You entered the number 0.')
```
### 2.14 Your Turn!
Using this, write a code block that prints out if the inputted number is greater than 0, equal to 0 or negative. Your code shoule use `if`, `elif`, and an `else` statement
```
b = 15
if b>10:
print("best day ever")
if b<10:
print ("terrible day")
else:
print("decent day")
```
Running the program and passing a number when prompted will cause the conditions to be checked and result in the following output.
```python
In [1]: run if_elif_else.py
Please enter a number: 10
You entered a positive number.
In [2]: run if_elif_else.py
Please enter a number: -10
You entered a negative number.
In [3]: run if_elif_else.py
Please enter a number: 0
You entered the number 0.
```
Let's specifically talk about how the `if`-`elif`-`else` statements work. The programmers of Python designed these statements so that they would execute highly efficiently. They achieved this by making it so that when Python is going through your `if`-`elif`-`else` statements and encounters a condition that evaluates to `True`, it will execute the corresponding conditional code block and then skip to the line directly following the last conditional block. Let's examine this in the following code saved again in `if_elif_else.py`.
```python
x = int(input('Please enter a number: '))
if x > 5:
print('You entered a number bigger than 5.')
elif x > 0:
print('You entered a positive number.')
elif x < 0:
print('You entered a negative number.')
else:
print('You entered the number 0.')
```
Running this program produces slightly unexpected results. But, they will soon make perfect sense, and knowing what is going on will allow you full control over the flow of your programs.
```python
In [1]: run if_elif_else.py
Please enter a number: 5
You entered a positive number.
In [2]: run if_elif_else.py
Please enter a number: 6
You entered a number bigger than 5.
```
In the first example we got something unsurprising. The only condition that evaluates to true when `x` is 5 is the second one. However, the second example yields only 'You entered a number bigger than 5.', even though 6 is greater than 0. This shows that only one of the conditional blocks in an `if`-`elif`-`else` statement will ever be evaluated, and once this happens the rest are skipped.
**Note**: The `else` part of the statement is actually optional. If it is not included, then we'd notice that at most one of the conditional blocks in an `if`-`elif` statement will be evaluated.
## 2.2 And, Or and Not
There are plenty of times when we want to execute some specific code when more than one condition is true. Check out the following code snippet.
```python
if x > 5:
if x < 10:
print(x)
```
We can see that what this **nested** `if` statement is checking for are numbers that lie in the interval (5, 10), and if it finds one it prints it. We can intuitively guess that there is a better way to check for this condition. And there is!!!
Python gives us full access to what are known as boolean operations. The ones that we will use most often are `and`, `or` and `not`. Both `and` and `or` take two conditions as inputs, while `not` takes only a single condition. They all return a single boolean, with `and` requiring both conditions to be `True` to return `True`, and the `or` requiring only one of the conditions to be `True` to return `True`. The `not` switches the truth of the input condition. These operations are derived from formal logic, and you can find a full discussion of their intricacies found [here](https://en.wikipedia.org/wiki/Truth_table).
### 2.21 Your Turn
Play around with the and statements
Try to make the longest statement positive that will still return `True`
```
# Elliott
True and True or False
```
What this means is that we now have a natural way to combine conditions. The previously nested `if` statement can now be written as a simple `if x > 5 and x < 10`. We can also chain other interesting conditionals together.
```python
if x > 10 or x < 5:
print(x)
if not (x <= 10 and x >= 5):
print(x)
```
Notice how the first `if` in the above code snippet uses an `or`, printing `x` if it is greater than 10 or less than 5. Inherently this statement is also saying that it will print `x` if `x` is not between 5 and 10, which is expressed in the second `if` statement. This illustrates an important point - there is always more than one way to accomplish the same thing in programming.
## 2.30 Looping
We are now prepared to learn about another extremely powerful programming construct. Everything that we learned in the last section on logic is part of an idea called **control flow**. Flow refers to the order in which statements in your program are executed. Controlling this flow can be done in many ways; so far we have learned about `if`-`elif`-`else` statements, but there are a number of others.
One thing that we find in programming is that we want to do something over and over (and over), possibly under the same circumstances each time, but frequently under slightly different circumstances each time. With the tools that we currently possess, we have to write out a line of Python for each time that we want to do that something. Let's go through a more concrete example.
Consider that you are asked to write a program to calculate the sum of the numbers between 1 and 8 (without the use of any built-in Python functions). We could write an extremely simple program to do this for us.
```python
sum_1_8 = 1 + 2 + 3 + 4 + 5 + 6 + 7 + 8
print(sum_1_8)
```
While this definitely works, there are a couple of things I want to draw your attention to (which will become themes about how to analyze how well code is written). First, what happens if we want to add the numbers 1 through 9 together? Not that hard, just add 9 to `sum_1_8` you say. Ok, fine. What if you want to add 2 through 9 together? Now we could take `sum_1_8`, add 9 and subtract off 1. And that works, but it involves some thinking to make this new idea work with the existing code that we have written.
Instead of having all of these **hard coded** values in our definition of `sum_1_8`, we could instead **abstract** away part of our problem. What is this abstraction? In programming, we talk about abstraction when we want to refer to an idea whose implementation is more general and/or hidden from us. In the above example, we see exactly what we're doing to sum the numbers 1 through 8. This isn't abstracted at all. So how are we to solve this problem more abstractly?
This is a question that you will frequently be faced with; how do you do something... in code? A good strategy to solve these problems is to approach the problem from a high level (i.e. in plain English, no code).
So let's do that with our coding problem above. We were asked to add together the numbers 1 through 8. This can be thought of as given a starting number, 1, and then adding on the next number, 2, to get 3. Then we can repeat this process, taking the next number, 3, and adding it on, giving us 6. We could then continue this process until we reach the final number, 8, and then stop. (This is inherently what we were doing in that single line of Python when we said `1 + 2 + 3 + 4 + 5 + 6 + 7 + 8`, but that implementation is what we call **brittle** - it only works for that specific case and breaks whenever we want to do something even slightly different.)
### 2.31 While Loops
Notice how in our high level description of the problem solution, we kept saying "and then". This repetitious language brings us to our next control flow tool, loops. There are two types of loops in Python, but today we're going to focus on `while` loops. `while` loops are an amazing tool which simply allow us to have a predefined chunk of code which we tell Python we want to run over and over under certain conditions.
So what are these conditions? They are in fact the conditions we learned about in the logic section (i.e. any expression that is evaluated to a boolean). So how does this work with `while`? Let's take a look at the structure of a `while` statement.
```python
while condition:
while_block_statement
```
```
# Elliott
x = 0
while x<10:
print(x)
x = x+1
```

As with `if`, a `while` statement has a condition; unlike the `if`, the while block will execute over and over again as long as the condition is `True`. This is where we get the name `while` loop from - as long as the condition evaluates to `True`, we will execute the code inside the `while` block, looping over it. The `while` condition is checked each time before the `while` statement block is executed.
Let's look at how we can harness this new structure to solve our previous problem. Take a look at the following code.
```python
total, x = 0, 1
while x <= 8:
total += x
x += 1
print(total)
```
Let's break down this code to see what is going on. On the first line, we declare a couple of variables (here you see the Python syntax used to do multiple assignments in a single line), `total` and `x`. `total` is the variable that we are going to aggregate our sum into, and `x` is the first number that we start our adding at. The next line declares the start of our newly learned `while` block. It's condition is x <= 8, and naturally reads as: "while x is less than or equal to 8", do stuff in the block. The block then says we are to add the current value of `x` to total, then add one to `x`.
### 2.32 More Control Flow
#### Continue
So what if we want even more control over how the body of our loop is executed? Let's motivate this idea with a problem. Say we want to add all the numbers from 1 to 8... but not 5. Again, we could solve this with our current solution, and then subtract off 5. But, again, that takes a lot of manipulation. Instead, we can use the main structure of our current loop and add in a new condition with an `if` and use a new tool to interrupt our program's flow.
Enter `continue`. What `continue` does is simply tell Python that it should skip the rest of the body of the `while` block, and jump (`continue`) to the next iteration of the loop. Let's take a look at `continue` in action.
```python
total, x = 0, 1
while x <= 8:
if x == 5:
x += 1
continue
total += x
x += 1
print(total, x)
print(total)
```
In this updated program we can see that we will, at each iteration of the loop, check to see if the current value that we're about to add on to `total` is 5. If it isn't, we go on with our aggregation of `total`. If `x` is 5, we add one to `x` (do you see why we need to do this?), and skip adding `x` to total by executing a `continue`, jumping immediately to the next iteration of the loop. Let's see how this would look in the loop table.
| After loop # | total | x | x <= 8 | x == 5 |
| ------------- |:---------:|:-----:|:----------:|:-----------:|
| 1 | 1 | 2 | True | False |
| 2 | 3 | 3 | True | False |
| 3 | 6 | 4 | True | False |
| 4 | 10 | 5 | True | True |
| 5 | 10 | 6 | True | False |
| 6 | 16 | 7 | True | False |
| 7 | 23 | 8 | True | False |
| 8 | 31 | 9 | False | False |
During the fourth iteration of the loop, when `x` is 5, we see that `total` does not get 5 added to it. Therefore, the final answer is, as we'd expect, 31.
### 2.33 Break Statements
In addition to the continue, we have another, more aggressive, method to control the flow of our programs - `break`. Where `continue` allowed us to skip the rest of the loop's code block and jump directly to the next iteration of the loop, `break` allows us to manually leave the loop entirely.
Time for an illustrative example. Consider trying to write a program that adds the numbers 1 to 8, but only up to 25. If the sum exceeds 25, the total is set to 25 and the message, "The sum exceeded the max value of 25." is printed. We could certainly complete this task with the tools that we already possess, but `break` is better suited to meet the needs of this situation. Let's take a look at what this implementation would look like.
```python
total, x = 0, 1
while x <= 8:
if total > 25:
total = 25
print('The sum exceeded the max value of 25.')
break
total += x
x += 1
print(total)
```
At this point I'm confident that you are tired of looking at tables of values, but let's do this one last time for consistency under the above program specifications.
| After loop # | total | x | x <= 8 | total > 25 |
| ------------- |:---------:|:-----:|:----------:|:-----------:|
| 1 | 1 | 2 | True | False |
| 2 | 3 | 3 | True | False |
| 3 | 6 | 4 | True | False |
| 4 | 10 | 5 | True | False |
| 5 | 15 | 6 | True | False |
| 6 | 21 | 7 | True | False |
| 7 | 28 | 8 | True | True |
At this point `total` is set to 25 and the message "The sum exceeded the max value of 25." is printed. The loop is exited and then 25 (the value of `total` now) is printed to the screen.
### 2.34 Pass
There's one more statement that allows us control over our programs - `pass`. All `pass` does is tell Python to do nothing. Because of this, it is rarely used for control flow, since the same result could be achieved by doing nothing. Instead, it is frequently offered as a place holder, since Python will complain about empty code blocks.
So while you're building up the skeleton of a program, `pass` can be useful as a method to get the framework written up without focusing on implementation. To illustrate...
```python
if x < 0:
pass
elif x > 0:
pass
else:
print('x is the value of 0.')
```
In the above example, we have set it up so that if `x` is 0, then our program tells us so. Otherwise, we know that we're going to do something specific when `x` is positive and something different when `x` is negative. We have used pass to suggest that we either haven't figured those things out yet, or simply haven't implemented them.
```
```
| github_jupyter |
<a href="https://colab.research.google.com/github/purple221/Hack-Cambridge/blob/main/Untitled0.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
#Description: This progrma detects covid-19 based off of data.
#import libraries
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
#loading data
from google.colab import files
uploaded = files.upload()
#Store the data into a variable
dataFrame = pd.read_csv('COVID19-data.csv')
#Print the first seven rows of data
dataFrame.head(7)
#Get the shape of the data
dataFrame.shape
```
```
#Visualize the count
sns.countplot(dataFrame['Gender'])
sns.countplot(dataFrame['Age group'])
#Viewing some basic statistic
dataFrame.describe()
#remove the columns that do not contribute
dataFrame = dataFrame.drop('Case identifier number', axis=1)
dataFrame = dataFrame.drop('Region', axis=1)
dataFrame = dataFrame.drop('Onset year of symptoms', axis=1)
dataFrame = dataFrame.drop('Episode week', axis=1)
dataFrame = dataFrame.drop('Episode week group', axis=1)
dataFrame = dataFrame.drop('Episode year', axis=1)
dataFrame = dataFrame.drop('Gender', axis=1)
dataFrame = dataFrame.drop('Occupation', axis=1)
dataFrame = dataFrame.drop('Onset week of symptoms', axis=1)
dataFrame = dataFrame.drop('Asymptomatic', axis=1)
dataFrame = dataFrame.drop('Age group', axis=1)
dataFrame = dataFrame.drop('Hospital status', axis=1)
dataFrame = dataFrame.drop('Recovered', axis=1)
dataFrame = dataFrame.drop('Recovery week', axis=1)
dataFrame = dataFrame.drop('Recovery year', axis=1)
dataFrame = dataFrame.drop('Death', axis=1)
dataFrame = dataFrame.drop('Transmission', axis=1)
#Get the correlation of the columns
dataFrame.corr()
import matplotlib.pyplot as plt
plt.figure(figsize=(7,7))
sns.heatmap(dataFrame.corr(),annot=False,fmt='.0%')
#Split the data into feature data and target data
X = dataFrame.iloc[:, :-1].values
Y=dataFrame.iloc[:, -1].values
#Split the data again, into 75% training data set and 25% testing data set
from sklearn.model_selection import train_test_split
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size = 0.25, random_state = 1)
#Feature Scaling
#Scale the values in the data to be values between 0 and 1 inclusive
from sklearn.preprocessing import StandardScaler
sc = StandardScaler()
X_train = sc.fit_transform(X_train)
X_test = sc.transform(X_test)
# use Random Forest Classifier
from sklearn.ensemble import RandomForestClassifier
forest = RandomForestClassifier(n_estimators = 10, criterion = 'entropy', random_state = 1)
forest.fit(X_train, Y_train)
```
| github_jupyter |
# Experimenation
Having two comparable models, we want to answer simple question - **is model A or B doing better?** It turns out that answering this simple question is quite complex.
```
# youtube
from IPython.display import YouTubeVideo
YouTubeVideo('-kjRL-Q-KBc', start=55, width=600, height=400)
```
## Clinical Trials
<img src="https://ichef.bbci.co.uk/news/660/cpsprodpb/D597/production/_91097645_jameslind.jpg" />
[James Lind and Scurvy: The First Clinical Trial in History?](https://www.bbvaopenmind.com/en/science/leading-figures/james-lind-and-scurvy-the-first-clinical-trial-in-history)
## Online Controlled Experiments
Online controlled experiments, A/B tests or simply experiments, are widely used by data-driven companies to evaluate the impact of change and are the only data-driven approach to prove causality.
<img src="./imlp3_files/oce.png" width="600" />
Sample of real users, not WEIRD (Western, Educated, Industrialized, Rich, and Democratic).
Experiment execution:
1. Users are randomly assigned to one of the two variants: Control (A) or Treatment (B).
2. Control is usually existing system while Treatment is the system with the new feature $X$ to test.
3. Users' interactions with the system are recorded.
4. We calculate metrics from recorded data.
If the experiment has been designed and executed correctly, the only thing consistently different between the two variants is feature $X$. All external factors are eliminated by being evenly distributed among Control and Treatment. We can hypothesize that any difference in metrics between the two variants can be attributed to either feature $X$ or to due to a change resulted from random assigment of users to variants. The later is ruled out (probalistically) using statistical tests. This establishes a causal relationship between the feature change $X$ and changes in user behavior. In S. Gupta, R. Kohavi, et al. [Top Challenges from the first Practical Online Controlled
Experiments Summit](https://exp-platform.com/Documents/2019-FirstPracticalOnlineControlledExperimentsSummit_SIGKDDExplorations.pdf)
1. [Statistics at Heart](#intuition)
1. [Null Hypothesis Significance Testing and Sampling Distributions](#null)
1. [Bayes Analysis](#bayes)
1. [Decision Rules](#decision)
1. [Probabilistic Results](#pbb)
1. [Rules of Thumb](#rules)
1. [Top Challenges](#not)
1. [Resources](#resources)
## Why A/B Tests?
Because assessing the value of novel ideas is hard.
### Correlation and Causation
Bloodletting effect of having calming effects on patients let doctors practice it for 2000 years.
<img src="https://upload.wikimedia.org/wikipedia/commons/b/bc/BloodlettingPhoto.jpg" width="300" />
[Wikipedia](https://en.wikipedia.org/wiki/Bloodletting)
### Novelty
1. [Semmelweis Reflex](https://en.wikipedia.org/wiki/Semmelweis_reflex).
2. It is hard to subject your idea to the metric and receive negative feedback.
<img src="https://upload.wikimedia.org/wikipedia/commons/thumb/f/f8/Ignaz_Semmelweis_1860.jpg/440px-Ignaz_Semmelweis_1860.jpg" width="300" />
### Learnings
1. *To have a great idea, have a lot of them.* (Thomas Edison)
1. *If you have to kiss a lot of frogs, to find a prince, find more frogs and kiss them faster and faster.* (Mike Moran, Do it Wrong Quickly)
<table><tr>
<td><a href="https://en.wikipedia.org/wiki/Thomas_Edison"><img src="https://upload.wikimedia.org/wikipedia/commons/thumb/9/9d/Thomas_Edison2.jpg/440px-Thomas_Edison2.jpg"/ width="300"></a></td>
<td><a href="https://www.amazon.com/Do-Wrong-Quickly-Changes-Marketing/dp/0132255960"><img src="https://images-na.ssl-images-amazon.com/images/I/51WczH0uCPL._SX375_BO1,204,203,200_.jpg" width="300"/></a></td>
</tr></table>
### HiPPO

*If we have data, let's look at data. If all we have are opinions, let's go with mine.* (Jim Barksdale - former CEO of Netscape). [Link](https://exp-platform.com/hippo/).
```
# import some basic libraries to do analysis and ML
import pandas as pd
import numpy as np
import sklearn as sk
import matplotlib as mpl
import matplotlib.pyplot as plt
import matplotlib.pylab as pylab
import scipy.stats as st
import seaborn as sns
%matplotlib inline
import warnings
warnings.filterwarnings('ignore')
# Visualization details
mpl.style.use('ggplot')
sns.set_style('white')
pylab.rcParams['figure.figsize'] = 12,8
```
<a id="intuition"></a>
## Statistics at Heart
Experiments and their evaluation is a work with uncertainity that requires some mathematics so we can talk about it precisly :)
Let's use an experiment measuring click through rate CTR (defined as clicks/views) of some button on some website. Control variant A (current default) uses green color, threatment variant B (challenger) uses red button. We ran randomized experiment on users of our website with following results
### What Variant to Choose?
| Variant | Views | Clicks | CTR | Relative Difference |
| ------- | -----:| ------:| ---:| -------------------:|
| A | 1,000 | 100 | 10% | --- |
| B | 1,000 | 110 | 11% | +10% |
Before we decided to run such test, we agreed on stopping it when both variants reach 1,000 views. We can't run this experiment any longer because it takes too long to get that many views. Our question to answer is if variant B is better than variant A. This is all information we have and we have to determine if we trust that B is better than A.
We work with a sample of 1,000 views per variant and want to reason about whole populations of views (users, ...) in A and B. CTR of 10% (11%) are the most likely estimates of population CTR given the data we have. But the real situation is little bit more complicated. True population CTRs can be anywhere around our estimates.
Probability distribution of true population CTRs.
```
# estimations
N = 100000
a = st.beta(100, 1000 - 100)
b = st.beta(110, 1000 - 110)
a_rvs = a.rvs(N)
b_rvs = b.rvs(N)
fig, ax = plt.subplots(figsize=(7, 5))
cs = sns.color_palette("husl", 2)
sns.kdeplot(a_rvs, bw='silverman', legend=True, ax=ax, label='A', linewidth=4, color=cs[0]);
ax.axvline(np.mean(a_rvs), 1., 0., c=cs[0], linestyle='dashed', alpha=.4, label='A Mean', linewidth=3)
sns.kdeplot(b_rvs, bw='silverman', legend=True, ax=ax, label='B', linewidth=4, color=cs[1]);
ax.axvline(np.mean(b_rvs), 1., 0., c=cs[1], linestyle='dashed', alpha=.4, label='B Mean', linewidth=3)
ax.set_title('Estimation of CTR')
ax.set_xlabel('CTR');
ax.set_ylabel('Density')
ax.legend()
ax.grid();
```
It can easily happen that true CTR of variant B is below true CTR of variant A despite measuring otherwise.
```
# where could true population means be?
fig, ax = plt.subplots(figsize=(7, 5))
cs = sns.color_palette("husl", 2)
sns.kdeplot(a_rvs, bw='silverman', legend=True, ax=ax, label='A', linewidth=4, color=cs[0]);
ax.axvline(np.mean(a_rvs), 1., 0., c=cs[0], linestyle='dashed', alpha=.4, label='A Mean', linewidth=3)
sns.kdeplot(b_rvs, bw='silverman', legend=True, ax=ax, label='B', linewidth=4, color=cs[1]);
ax.axvline(np.mean(b_rvs), 1., 0., c=cs[1], linestyle='dashed', alpha=.4, label='B Mean', linewidth=3)
ax.set_title('Estimation of CTR')
ax.set_xlabel('CTR');
ax.set_ylabel('Density')
ax.legend()
ax.grid();
ax.arrow(0.11, 10, 0, -10, length_includes_head=True, width=.005, head_length=2, head_width=.01, overhang=0, color=cs[0], shape='full', zorder=10);
ax.arrow(0.09, 10, 0, -10, length_includes_head=True, width=.005, head_length=2, head_width=.01, overhang=0, color=cs[1], shape='full', zorder=10);
```
What to do, do having more data help in this case? Yes, having more data makes our estimate more accurate. So it is very unlikely that CTR of variant A to be close to 0.11.
```
# effect of more samples in the experiment
N = 100000
a = st.beta(100, 1000 - 100)
b = st.beta(1000, 10000 - 1000)
a_rvs = a.rvs(N)
b_rvs = b.rvs(N)
fig, ax = plt.subplots(figsize=(7, 5))
cs = sns.color_palette("husl", 2)
sns.kdeplot(a_rvs, bw='silverman', legend=True, ax=ax, label='A with 100 clicks', linewidth=4, color=cs[0]);
ax.axvline(np.mean(a_rvs), 1., 0., c=cs[0], linestyle='dashed', alpha=.4, label='A Mean', linewidth=3)
sns.kdeplot(b_rvs, bw='silverman', legend=True, ax=ax, label='A with 1,000 clicks', linewidth=4, color=cs[1]);
ax.axvline(np.mean(b_rvs), 1., 0., c=cs[1], linestyle='dashed', alpha=.4, label='B Mean', linewidth=3)
ax.set_title('Estimation of CTR')
ax.set_xlabel('CTR');
ax.set_ylabel('Density')
ax.legend()
ax.grid();
ax.arrow(0.11, 30, 0, -30, length_includes_head=True, width=.005, head_length=5, head_width=.01, overhang=0, color=cs[1], shape='full', zorder=10);
```
<a id="null"></a>
## Null Hypothesis Significance Testing
*Null hypothesis significance testing* is common approach to deal with this uncertainties.
*NHST* decides if a null hypothesis can be rejected given obtained data.
Examples of null and alternative hypothesis:
1. $H_0$: A coin is fair.
1. Meaning we can expect equal number of heads and tails from $N$ tosses of the coin.
1. $H_1$: A coin is biased towards heads or tails.
1. $H_0$: CTRs of variants A and B are equal.
1. $H_1$: There's difference between CTR in A and B.
### Sampling Distribution
```
# sampling distributions
import scipy.stats as st
x1 = st.norm.rvs(size=1000)
y1 = st.norm.rvs(size=1000)
x2 = st.norm.rvs(scale=2, size=1000)
y2 = st.norm.rvs(scale=2, size=1000)
fig, ax = plt.subplots(figsize=(6, 6));
# ax = axx[0]
# ax.scatter(x1, y1, marker='o', alpha=.4);
# ax.scatter([1.5], [0], marker='o', color='g', lw=10)
# c = plt.Circle((0, 0), radius=1.5, color='g', fill=False, lw=3);
# ax.add_artist(c);
# ax.arrow(1.5, 0., 1.0, 0., width=.05, head_width=.2, overhang=0.1, color='k');
# ax.arrow(2.5, 0., -0.7, 0., width=.05, head_width=.2, overhang=0.1, color='k', shape='full');
# ax.arrow(1.5, 2., 0, -1.6, width=.05, head_width=.2, overhang=0.1, color='k', shape='full');
# ax.text(2, .3, 'Extreme values', fontsize=18);
# ax.text(1.5, 2, 'Actual Outcome', fontsize=18);
# ax.set_xlim((-7, 7));
# ax.set_ylim((-7, 7));
# ax.set_title('Sampling Distribution for Null Hypothesis H');
# ax.grid()
ax.scatter(x2, y2, marker='o', alpha=.4);
ax.scatter([2.5], [0], marker='o', color='r', lw=10)
c = plt.Circle((0, 0), radius=2.5, color='r', fill=False, lw=3);
ax.add_artist(c);
ax.arrow(2.5, 0., 3., 0., width=.05, head_width=.3, overhang=0.1, color='k', shape='full');
ax.arrow(4.5, 0., -1.6, 0., width=.05, head_width=.3, overhang=0.1, color='k', shape='full');
ax.arrow(2.5, 2., 0, -1.6, width=.05, head_width=.3, overhang=0.1, color='k', shape='full');
ax.text(2.5, 2, 'Actual Outcome', fontsize=18);
ax.text(2.7, .3, 'Extreme values', fontsize=18);
ax.set_xlim((-7, 8.5));
ax.set_ylim((-7, 8.5));
ax.set_title('Sampling Distribution for Null Hypothesis H');
ax.grid()
```
Let's assume our null hypothesis is valid. If we ran the same experiment many times, we get many point estimates $\hat{\mu}$ of population metric mean $\mu$ (eg. CTR). These estimates are normally distributed because of a Central limit theorem and because we assume the null hypothesis is true they are centered around population $\mu$. This distribution is called *sampling distribution* and it is **only imaginative**, we do not construct it from the data, it comes from the assumption about the null hypothesis.
We usually run only 1 experiment giving us point estimate $\hat{\mu}$ of population mean $\mu$. If $\hat{\mu}$ lies out of where eg. 95% of samples from the imaginative sampling distribution would lay, we say that we collected evidence that shows tha under 95% significance level ($0.95 = 1-\alpha$), we can reject the null hypothesis as being valid.
Note that this does not say much about a probability of the null hypothesis to be true or false. It can only reject the null hypothesis.

[STAT 414 / 415, Probability Theory and Mathematical Statistics](https://newonlinecourses.science.psu.edu/stat414/node/196/)
### p-Value
*p-Value* is the probability of getting point estimates $\hat{\mu}_i$ from running multiple experiments equally or more extreme than the one point estimate $\hat{\mu}$ we've got from our real experiment under *sampling distribution* based on our null hypothesis. If p-value is below 5%, we reject the null hypothesis under 95% significance level.
Note that p-value does not talk about probability of where we can expect $\mu$. It talks about probability of getting such or more extreme result.
Let's illustrate p-value under some null hypothesis and stopping criteria. Most of all imaginative experiment results falls around the mean of sampling distribution (from central limit theorem). Some fall beyond actual outcome. p-value is proportion of the cloud at actual outcome and beyond.
### p-Value Depends on How we Perform the Test
Many of the following examples are taken from very well written book **John Kruschke** [Doing Bayesian Data Analysis](https://www.amazon.com/dp/0124058884).
Test if a coin is a fair coin ($\theta = \frac{1}{2}$) or not given a sample consisting of $N$ tosses with $z$ heads.
```
# simulation
z = 7
N = 24
theta = 0.5
b = st.binom(N, theta)
zs = np.arange(0., N + 1)
ns = np.arange(z, 1000.)
proba_n = b.pmf(zs)
left_tail_n = np.sum(b.pmf(zs[zs <= 7]))
proba_z = z/ns * st.binom.pmf(z, ns, theta)
left_tail_z = 1 - np.sum(z / ns[ns < N] * st.binom.pmf(z, ns[ns < N], theta))
fig, axx = plt.subplots(nrows=1, ncols=2, figsize=(17, 6));
fig.suptitle('p-Value Depends on How we Perform the Test', fontsize=18)
ax = axx[0]
ax.vlines(zs/N, 0, proba_n, colors='b', linestyles='-', lw=3);
ax.set_ylim((-0.005, 0.2));
ax.set_xlabel('Sample Proportion z/N');
ax.set_ylabel('p(z/N)');
ax.grid();
ax.set_title('Sampling Distribution - Fixed N');
ax = axx[1]
ax.vlines(z/ns, 0, proba_z, colors='b', linestyles='-', lw=3);
ax.set_ylim((-0.005, 0.15));
ax.set_xlabel('Sample Proportion z/N');
ax.set_ylabel('p(z/N)');
ax.grid()
ax.set_title('Sampling Distribution - Fixed z');
# simulation
fig, axx = plt.subplots(nrows=1, ncols=2, figsize=(17, 6));
fig.suptitle('p-Value Depends on How we Perform the Test', fontsize=18)
ax = axx[0]
ax.vlines(zs/N, 0, proba_n, colors='b', linestyles='-', lw=3);
ax.plot([float(z)/N], [0.], marker='o', color='k', markersize=10);
ax.text(.1, .14, 'N = %0.0f\nz = %0.0f\np = %0.3f' % (N, z, left_tail_n), fontsize=18)
ax.set_ylim((-0.005, 0.2));
ax.set_xlabel('Sample Proportion z/N');
ax.set_ylabel('p(z/N)');
ax.grid();
ax.set_title('Sampling Distribution - Fixed N');
ax.arrow(float(z)/N, 0., -float(z)/N, 0., width=.001, head_width=.008, overhang=0.2, color='k');
ax = axx[1]
ax.vlines(z/ns, 0, proba_z, colors='b', linestyles='-', lw=3);
ax.plot([float(z)/N], [0.], marker='o', color='k', markersize=10);
plt.text(.1, .1, 'N = %0.0f\nz = %0.0f\np = %0.5f' % (N, z, left_tail_z), fontsize=18)
ax.set_ylim((-0.005, 0.15));
ax.set_xlabel('Sample Proportion z/N');
ax.set_ylabel('p(z/N)');
ax.grid()
ax.set_title('Sampling Distribution - Fixed z');
ax.arrow(float(z)/N, 0., -float(z)/N, 0., width=.001, head_width=.006, overhang=0.2, color='k');
```
### Quiz
You've run an A/B test. Your A/B testing software has given you a p-value of $0.03$. Which of the following is true? (Several or none of the statements may be correct.)
1. You have disproved the null hypothesis (that is, there is no difference between the variations).
1. The probability of the null hypothesis being true is 0.03.
1. You have proved your experimental hypothesis (that the variation is better than the control).
1. The probability of the variation being better than control is 97%.
[Chris Stucchio, Bayesian A/B Testing at VWO](https://www.chrisstucchio.com/pubs/slides/gilt_bayesian_ab_2015/slides.html#7)
All above statements are wrong!
### NHST Pros
1. Simplicity.
2. Binary thinking.
3. Studied well, tools and knowledge available.
### NHST Cons
1. Sad faces of experimenters when p-value >= 5% and we don't know if $H_0$ or $H_1$ is valid (this happens 90% of times!).
1. Needs statisticians to explain and overcome many misleading pitfalls when not running 100% correctly.
1. Unintuitive reasoning based on sampling distributions.
## Bayesian Analysis
Bayesian analysis solves all our cons of NHST.
### How Do We Update Our Belief in Things?
1. We assume our own belief for an event to happen (eg. someone gets sick) from colloquial/anecdotal knowledge.
1. When we have more evidence pointing toward (away from) the event, we increase (decrease) our beliefe for an event to happen.
<center><i>Probability that an event has happened given a set of evidence for it <br />is equal to the probability of the evidence being caused by the event multiplied by the probability of the event itself.</i></center>
$$P(\text{patient has disease given positive test result)}$$
$$\text{"is"}$$
$$P(\text{test result positive AND patient has disease})$$
$$\text{"is"}$$
$$P(\text{test result positive given patient has disease}) \times P(\text{disease})$$
$$P(d\,|\,t)\ \text{"is"}\ P(t\,|\,d) \times P(d)$$
### Example
1. Patient has tested positive for disease $d$.
2. The test used has hit rate (recall) of 99% - probability of testing positive $t$ given patient has a disease.
3. The test used has false alarm rate (fpr) of 5% - probability of testing positive $t$ given patient does not have a disease.
4. The disease is rare striking only 1 out of 1,000 people.
What do you think is the probability of the patient having the disease when his test was positive?
### Example
1. Patient has tested positive for disease $d$.
2. The test used has accuracy of 99% - probability of testing positive $t$ given patient has disease $d$
3. The test used has false alarm rate of 5% - probability of testing positive $t$ given patient does not have a disease.
4. The disease is rare striking only 1 out of 1,000 people.
The probability that the patient actually has the disease (given he has tested positive) is:
$$ P(d\,|\,t) = \frac{P(t\,|\,d) \times P(d)}{P(t)} $$
$$ P(d\,|\,t) = \frac{P(t\,|\,d) \times P(d)}{P(t)} $$
1. $P(t\,|\,d) = 0.99$
1. $P(d) = 0.001$
1. $P(t)$ overall probability of test returning a positive value is $P(t) = P(t\,|\,d)\,P(d) + P(t\,|\,\neg d)\,P(\neg d) = 0.05$
1. $P(t\,|\,\neg d) = 0.05$
$$ P(d\,|\,t) = \frac{0.99 \cdot 0.001}{0.99 \cdot 0.001 + 0.05 \cdot (1-0.001)} = 0.019 $$
### Different (Better) Setting?
But the patient went to see a doctor, the prior of 1 out of 1,000 is not a reality. Doctor's data show that 1 patient has the disease out of 100 of those who enter the office.
$$ P(d\,|\,t) = \frac{0.99 \times 0.01}{0.05} = 0.198 $$
### Bayesian Inference
```
# prior, likelihood, posterior
colors = ["windows blue", "amber", "greyish", "faded green", "dusty purple", "pink", "brown", "red", "light blue", "green"]
colors = sns.xkcd_palette(colors)
N = 50
z = 35
theta = z/N
rv = st.binom(N, theta)
mu = rv.mean()
a, b = 20, 20
prior = st.beta(a, b)
post = st.beta(z+a, N-z+b)
ci = post.interval(0.95)
thetas = np.linspace(0, 1, 200)
plt.figure(figsize=(20, 7))
plt.subplot(131)
plt.plot(thetas, prior.pdf(thetas), label='Prior', c=colors[1], lw=3)
plt.fill_between(thetas, 0, prior.pdf(thetas), color=colors[1]);
plt.xlabel(r'$\theta$', fontsize=14)
plt.ylabel(r'$P(\theta) = beta(\theta\ |\ a, b)$', fontsize=16)
plt.legend();
plt.grid()
plt.ylim((-0.5, 9));
plt.title('Prior');
plt.subplot(132)
plt.plot(thetas, N*st.binom(N, thetas).pmf(z), label='Likelihood', c=colors[0], lw=3)
plt.fill_between(thetas, N*st.binom(N, thetas).pmf(z), color=colors[0]);
plt.xlabel(r'$\theta$', fontsize=14)
plt.ylabel(r'$P(z,N\ |\ \theta)$', fontsize=16)
plt.legend();
plt.grid()
plt.ylim((-0.5, 9));
plt.title('Likelihood');
plt.subplot(133)
plt.plot(thetas, prior.pdf(thetas), label='Prior', c=colors[1], lw=3, dashes=[2, 2])
plt.plot(thetas, N*st.binom(N, thetas).pmf(z), label='Likelihood', c=colors[0], lw=3,dashes=[2, 2])
plt.plot(thetas, post.pdf(thetas), label='Posterior', c=colors[2], lw=3)
plt.fill_between(thetas, 0, post.pdf(thetas), color=colors[2]);
# plt.axvline((z+a-1)/(N+a+b-2), c=colors[2], linestyle='dashed', alpha=0.8, label='MAP', lw=3)
# plt.axvline(mu/N, c=colors[0], linestyle='dashed', alpha=0.8, label='MLE', lw=3)
plt.xlim((0, 1));
plt.ylim((-0.5, 11));
plt.axhline(post.pdf(ci[0]), ci[0], ci[1], c='black', label='95% HDI', lw=3);
plt.axvline(ci[0], 0.5/11.5, (post.pdf(ci[0])+0.5)/11.5, c='black', linestyle='dotted', lw=3);
plt.axvline(ci[1], 0.5/11.5, (post.pdf(ci[0])+0.5)/11.5, c='black', linestyle='dotted', lw=3);
plt.xlabel(r'$\theta$', fontsize=14)
plt.ylabel(r'$P(\theta\ |\ z,N) = beta(\theta\ |\ a+z, b+N-z)$', fontsize=16)
plt.legend();
plt.ylim((-0.5, 9));
plt.grid()
plt.title('Posterior');
```
### Bayes Rule
$$ P(\theta\,|\,D) = \frac{P(D\,|\,\theta)\times P(\theta)}{P(D)} $$
where $\theta$ is our estimation (eg. CTR), $D$ data we measured ($z$ heads from $N$ tosses).
$$ \text{Posterior} = \frac{\text{Likelihood} \times \text{Prior}}{\text{Total Evidence}} $$
### Impact of Prior to Posterior
```
# prior to posterior
N = 30
z = 25
theta = z/N
rv = st.binom(N, theta)
thetas = np.linspace(0, 1, 200)
fig, axx = plt.subplots(nrows=3, ncols=3, figsize=(15, 8));
fig.suptitle('Updating a Beta Prior Distribution', fontsize=18);
fig.tight_layout(rect=[0, 0.03, 1, 0.92])
fig.subplots_adjust(hspace=0.5, wspace=0.25)
colors = ["windows blue", "amber", "greyish", "faded green", "dusty purple", "pink", "brown", "red", "light blue", "green"]
colors = sns.xkcd_palette(colors)
def plot_column(a, b, z, N, thetas, axx, y_lim_top=13):
prior = st.beta(a, b)
post = st.beta(z+a, N-z+b)
ci = post.interval(0.95)
ax = axx[0]
ax.plot(thetas, prior.pdf(thetas), c=colors[1]);
ax.set_xlabel(r'$\theta$', fontsize=16);
ax.set_ylabel(r'$beta(%d, %d)$' % (a, b), fontsize=14);
ax.set_title(r'Prior $beta(%d, %d)$' % (a, b));
ax.set_ylim(bottom=-0.3);
ax.grid()
if y_lim_top is not None:
ax.set_ylim(top=y_lim_top);
ax.fill_between(thetas, 0, prior.pdf(thetas), color=colors[1]);
ax = axx[1]
pmf = N*st.binom(N, thetas).pmf(z)
ax.plot(thetas, pmf, label='Likelihood', c=colors[0]);
ax.set_xlabel(r'$\theta$', fontsize=16);
ax.set_ylabel(r'$P(n,z\ |\ \theta)$', fontsize=16);
ax.set_title(r'Likelihood $binomial(%d, %d, \theta)$' % (z, N));
ax.set_ylim(bottom=-0.3);
if y_lim_top is not None:
ax.set_ylim(top=y_lim_top);
ax.grid()
ax.fill_between(thetas, 0, pmf, color=colors[0]);
ax = axx[2]
ax.plot(thetas, post.pdf(thetas), label='Posterior', color=colors[2]);
ax.fill_between(thetas, 0, post.pdf(thetas), color=colors[2]);
ax.axhline(0, ci[0], ci[1], c='black', label='95% CI');
# ax.text(ci[0]-0.1, .5, '%0.3f' % ci[0], fontsize=12);
# ax.text(ci[1], .5, '%0.3f' % ci[1], fontsize=12);
ax.text(ci[0]+0.05, .3, '95%% HDI' % ci[0], fontsize=12);
ax.set_xlabel(r'$\theta$', fontsize=16);
ax.set_ylabel(r'$beta(%d, %d)$' % (z+a, N-z+b), fontsize=14);
ax.set_title(r'Posterior $beta(%d + %d, %d + %d)$' % (z, a, N-z, b));
ax.set_ylim(bottom=-1);
ax.grid()
if y_lim_top is not None:
ax.set_ylim(top=y_lim_top);
plot_column(1, 1, z, N, thetas, axx[:][0])
plot_column(18, 6, z, N, thetas, axx[:][1])
plot_column(100, 100, z, N, thetas, axx[:][2])
```
### Bayes Rule for Conversions
1. Estimate conversion rate $\theta$ from $N$ impressions of an online ad that recorded $z$ clicks.
2. Or estimate the bias $\theta$ of a coin given a sample consisting of $N$ tosses with $z$ heads.
$$P(\theta\,|\,z, N) = \frac{P(z, N\,|\,\theta)\ \text{beta}(a, b)}{P(z, N)} = \text{beta}(a+z, b+N-z)$$
Where
* $P(z, N\,|\,\theta) = P(z, N\,|\,\theta) = \binom{N}{z}\theta^z(1-\theta)^{N-z}$ is binomial likelihood function of getting test results for different $\theta$.
* $P(\theta) = beta(\theta\,|\,a, b)$ is our prior belief about where $\theta$ is before running experiment.
* $P(\theta\,|\,z,N)$ is posterior distribution aka our prior believe updated by the evidence from the experiment results of $z/N$ of where we can expect $\theta$ to lie.
* $P(\theta\,|\,z,N) = \text{beta}(\theta\,|\,a + z, b + N - z)$
$$P(\theta\,|\,z, N) = \text{beta}(a+z, b+N-z)$$
Where
* $\text{beta}$ is beta distribution
* $a$ is prior number of successes
* $b$ is prior number of failures
* $N$ number of observations in the test
* $z$ number of successes (conversions) in the test
### Beta Distribution
```
# beta distributions
plots = 5
thetas = np.linspace(0, 1, 200)
fig, axx = plt.subplots(nrows=plots, ncols=plots, figsize=(15, 8));
fig.tight_layout(rect=[0, 0.03, 1, 0.95])
fig.subplots_adjust(hspace=0.9, wspace=0.4)
colors = ["windows blue", "amber", "greyish", "faded green", "dusty purple", "pink", "brown", "red", "light blue", "green"]
colors = sns.xkcd_palette(colors)
for a in range(plots):
for b in range(plots):
aa = a+1
bb = b+1
beta = st.beta(aa, bb)
ax = axx[a][b]
pst = beta.pdf(thetas)
ax.plot(thetas, pst, c=colors[1])
ax.fill_between(thetas, 0, pst, color=colors[1]);
ax.set_xlabel(r'$\theta$', fontsize=16);
# ax.set_ylabel(r'$p(\theta|a=%d, b=%d)$' % (aa, bb), fontsize=14);
ax.set_title(r'$beta(%d, %d)$' % (aa, bb));
ax.grid()
```
### Is Bayesian Analysis More Informative?
1. NHST does not work with priors while BA makes them explicit and part of the reasoning process.
1. NHST uses sampling distribution for inference while BA uses posterior distribution.
* Sampling distribution tells us probabilities of possible data given a particular (null) hypothesis.
* Posterior distribution tells us credibilities of possible hypothesis (values of $\theta$) given the data.
Good discussion of these properties from frequentists point of view could be found in:
1. [G. Georgiev, The Google Optimize Statistical Engine and Approach](http://blog.analytics-toolkit.com/2018/google-optimize-statistical-significance-statistical-engine/)
2. [G. Georgiev, 5 Reasons to Go Bayesian in AB Testing – Debunked](http://blog.analytics-toolkit.com/2017/5-reasons-bayesian-ab-testing-debunked/)
<a id='decision'></a>
## Decision Rules
1. NHST - Reject null hypothesis if p-value $\lt 5\%$.
1. Bayes
* Reject null hypothesis if 95% HDI is outside of *Region of Practical Equivalence (ROPE)* defined around null value.
* Accept null hypothesis if 95% HDI is completely inside of Region of Practical Equivalence.
### Decision Rules and False Positives
NHST has 100% false positive rate when doing sequentional testing stopped when p-value is below 5% with enough patience. Reasons:
1. NHST only rejects null hypothesis (cannot accept one).
1. There is non zero probability to get long stretch of extreme values that move p-value below 5%.
### Simulation of Decision Making
There are simulated data of 2000 flips with p-value calculated after every flip below. The first set is for case when null hypothesis of $\theta = 0.5$ is true. The second set is for case when null hypothesis is not true.
Graph with *$z/N$* shows proportion of $z$ in $N$ flips after every flip. Graph with p-value shows p-value calculated for cumulated data after every flip.
Graph with 95% HDI shows 95% HDI intervals after every flip as vertical line. Decision rule based on accepting null hypothesis when 95% HDI interval is completely within ROPE (dashed lines for $\theta = 0.45$ and $\theta = 0.55$) is much stable than NHST decision rule although it has some false positives when not enough data are available.
```
# null hypothesis is true
theta = 0.5
N = 2000
ns = np.arange(1, N+1)
zs = np.array(st.bernoulli.rvs(theta, size=N))
zs = np.cumsum(zs)
# np.save('zs_nh_true.npy', zs, allow_pickle=False, fix_imports=False)
zs = np.load('./imlp3_files/zs_nh_true.npy')
theta = 0.5
p_values = [np.sum(st.binom(n, theta).pmf(np.arange(0, zs[n-1] + 1))) for n in ns]
posts = st.beta(1 + zs, 1 + ns - zs)
cs = posts.interval(.95)
fig, axx = plt.subplots(nrows=3, ncols=1, figsize=(17, 10));
fig.suptitle(r'Sequentional Testing, Null Hypothesis is True - $\theta = 0.5$', fontsize=18);
fig.tight_layout(rect=[0, 0.03, 1, 0.95])
fig.subplots_adjust(hspace=0.3, wspace=0.25)
colors = ["windows blue", "amber", "greyish", "faded green", "dusty purple", "pink", "brown", "red", "light blue", "green"]
colors = sns.xkcd_palette(colors)
ax = axx[0]
ax.plot(ns, zs/ns, lw=7);
ax.axhline(0.5, ls='dashed', c='gray', lw=2)
ax.grid()
ax.set_ylabel(r'$z/N$')
ax.set_ylim((-0.05, 1.05))
ax = axx[1]
ax.scatter(ns, p_values, c=['b' if p_values[n] < 0.05 else 'k' for n in range(0, N)], lw=3);
ax.axhline(0.05, ls='dashed', c='gray', lw=2)
ax.grid()
ax.set_ylabel('p-value')
ax.set_ylim(bottom=-0.05)
ax = axx[2]
ax.vlines(ns, cs[0], cs[1], colors=['b' if cs[0][n] >= 0.45 and cs[1][n] <= 0.55 else 'k' for n in range(0, N)], linestyles='-')
ax.axhline(0.45, ls='dashed', c='gray', lw=2)
ax.axhline(0.55, ls='dashed', c='gray', lw=2)
ax.grid()
ax.set_ylim((-0.05, 1.05))
ax.set_ylabel('Accepting 95% HDI')
ax.set_xlabel('Flips');
```
#### Null Hypothesis is True
1. p-value < 5% based decision rule shows long stretches of p-value < 5% when if we stopped the test there, it would falsly reject null hypothesis. This example shows that even around flip 2000 it would falsly reject true null hypothesis.
2. 95% HDI decision rule shows much stable behavior starting fairy stably accepting true null hypothesis around step 1300.
```
# null hypothesis is false
theta = 0.7
zs1 = np.array(st.bernoulli.rvs(theta, size=N))
zs1 = np.cumsum(zs1)
# np.save('zsa.npy', zs1, allow_pickle=False, fix_imports=False)
zs1 = np.load('./imlp3_files/zs_null_false.npy')
theta = 0.7
p_values = [np.sum(st.binom(n, theta).pmf(np.arange(0, zs1[n-1] + 1))) for n in ns]
posts = st.beta(1 + zs1, 1 + ns - zs1)
cs = posts.interval(.95)
fig, axx = plt.subplots(nrows=3, ncols=1, figsize=(17, 10));
fig.suptitle(r'Sequentional Testing, Null Hypothesis is False - $\theta = %0.2f$' % theta, fontsize=18);
fig.tight_layout(rect=[0, 0.03, 1, 0.95])
fig.subplots_adjust(hspace=0.3, wspace=0.25)
colors = ["windows blue", "amber", "greyish", "faded green", "dusty purple", "pink", "brown", "red", "light blue", "green"]
colors = sns.xkcd_palette(colors)
ax = axx[0]
ax.plot(ns, zs1/ns, lw=7);
ax.axhline(0.5, ls='dashed', c='gray', lw=2)
ax.set_ylabel(r'$z/N$')
ax.grid()
ax.set_ylim((-0.05, 1.05))
ax = axx[1]
ax.scatter(ns, p_values, c=['b' if p_values[n] < 0.05 else 'k' for n in range(0, N)], lw=3);
ax.axhline(0.05, ls='dashed', c='gray', lw=2)
ax.set_ylabel('p-value')
ax.grid()
ax.set_ylim(bottom=-0.05)
ax = axx[2]
ax.vlines(ns, cs[0], cs[1], colors=['b' if cs[0][n] > 0.55 else 'k' for n in range(0, N)], linestyles='-')
ax.axhline(0.45, ls='dashed', c='gray', lw=2)
ax.axhline(0.55, ls='dashed', c='gray', lw=2)
ax.set_ylim((-0.05, 1.05))
ax.set_ylabel('Rejecting 95% HDI')
ax.grid()
ax.set_xlabel('Flips');
```
#### Null Hypothesis is False
1. p-value < 5% based decision rule starts correctly rejecting null hypothesis after step 1300. There's also stretch around 1800 steps when it stops rejecting null hypothesis.
2. 95% HDI decision rule starts rejecting null hypothesis much sooner and shows stable behavior.
<a id="pbb"></a>
## Probabilistic Results
We measure experiment results (conversion rates) and calculate their posterior distributions. We sample many (>= 1,000,000) samples from these posterior distributions and calculate following properties of these samples per variant.
1. Improvement
2. Probability to Beat Baseline
3. Probability to Be the Best
```
N = 10000 # samples per variant in experiment
S = 10000 # number of samples taken from posterior distribution
theta_a = 0.1 # simulated conversion rates of three variants
theta_b = 0.11
theta_c = 0.12
ns = np.arange(1, N+1)
zs_a = np.cumsum(np.array(st.bernoulli.rvs(theta_a, size=N))) # cumulated number of conversion
zs_b = np.cumsum(np.array(st.bernoulli.rvs(theta_b, size=N)))
zs_c = np.cumsum(np.array(st.bernoulli.rvs(theta_c, size=N)))
posts_a = st.beta(1 + zs_a, 1 + ns - zs_a) # for each measurement, take its beta distribution, we have N betas per variant
posts_b = st.beta(1 + zs_b, 1 + ns - zs_b)
posts_c = st.beta(1 + zs_c, 1 + ns - zs_c)
posts_a_rvs = posts_a.rvs(size=(S, N)) # sample S samples from each of N betas
posts_b_rvs = posts_b.rvs(size=(S, N))
posts_c_rvs = posts_c.rvs(size=(S, N))
improvement = np.median(((posts_c_rvs - posts_a_rvs) / posts_a_rvs), axis=0)
beat_baseline = np.sum(np.where(posts_c_rvs > posts_a_rvs, 1, 0), axis=0) / S
pbb = np.argmax(np.array((posts_a_rvs, posts_b_rvs, posts_c_rvs)), axis=0) # who won in 3x S samples in every N samples
pbb_a = np.sum(np.where(pbb == 0, 1, 0), axis=0) / S
pbb_b = np.sum(np.where(pbb == 1, 1, 0), axis=0) / S
pbb_c = np.sum(np.where(pbb == 2, 1, 0), axis=0) / S
```
### Improvement
Median improvement that we can expect over the baseline if we full scale the variant.
$$ \text{Improvement of C over A} = \text{median}\left(\frac{\text{Sample C} - \text{Sample A}}{\text{Sample A}}\right) $$
```
# improvement of C against A
f = 50
fig = plt.figure(figsize=(12, 3));
plt.plot(ns[f:], improvement[f:], lw=6, c=colors[0]);
plt.grid();
plt.title('Improvement of C against A');
plt.xlabel('Samples in Experiment');
plt.ylabel('Improvement');
plt.ylim((-.3, 1));
```
### Probability to Beat Baseline
Number of times treatment variant sample beats control (baseline) variant sample.
$$ \text{Probability of variant C beating A} = \frac{\text{Number of times variant C sample is greater than variant A sample}}{\text{Total number of samples}} $$
```
# probability of C beating baseline A
fig = plt.figure(figsize=(12, 6));
plt.plot(ns, beat_baseline, lw=6, c=colors[0]);
plt.axhline(0.95, ls='dashed', c='gray', lw=3)
plt.grid();
plt.ylim((-0.1, 1.1))
plt.title('Probability of C Beating Baseline A');
plt.xlabel('Samples in Experiment');
plt.ylabel('Probability of Beating Baseline');
```
### Probability to Be the Best
Number of times variant sample beats all other variants samples.
$$ \text{Probability of variant C being the best} = \frac{\text{Number of times variant C sample is greater than other variant samples}}{\text{Total number of samples}} $$
```
# probability of being best
fig = plt.figure(figsize=(12, 6));
plt.axhline(0.95, ls='dashed', c='gray', lw=3)
plt.plot(ns[f:], pbb_a[f:], lw=6, label='A', c=colors[1]);
plt.plot(ns[f:], pbb_b[f:], lw=6, label='B', c=colors[2]);
plt.plot(ns[f:], pbb_c[f:], lw=6, label='C', c=colors[0]);
plt.grid();
plt.legend();
plt.title('Probability of Being Best');
plt.xlabel('Samples in Experiment');
plt.ylabel('Probability of Being Best');
```
### AB Test Dashboard

<a id='resources'></a>
## Resources
1. **Prem S. Mann** Introductory Statistics 7nd Edition
2. **R. Kohavi, A. Deng** [Seven Rules of Thumb for Website Experimenters](https://exp-platform.com/Documents/2014%20experimentersRulesOfThumb.pdf)
3. **John Kruschke** [Doing Bayesian Data Analysis](https://www.amazon.com/dp/0124058884)
4. **S. Gupta, R. Kohavi et al.** [Top Challenges from the first Practical Online Controlled Experiments Summit](https://exp-platform.com/Documents/2019-FirstPracticalOnlineControlledExperimentsSummit_SIGKDDExplorations.pdf)
5. **H. Hohnhold, D. O'Brien, D. Tang** [Focusing on the Long-term: It's Good for Users and Business](https://static.googleusercontent.com/media/research.google.com/en//pubs/archive/43887.pdf)
5. [ExP Experimentation Platform](https://exp-platform.com/)
6. [Computational Statistics in Python](https://people.duke.edu/~ccc14/sta-663/MCMC.html)
| github_jupyter |
```
import pandas as pd
import numpy as np
import os
import time
import tensorflow as tf
import tensorflow_hub as hub
import zipfile
from html2text import HTML2Text
from tqdm import tqdm
import re
from sklearn.metrics import pairwise_distances
from sklearn.preprocessing import normalize
import matplotlib.pyplot as plt
import seaborn as sns
pd.set_option('display.max_rows', None)
pd.set_option('display.max_columns', None)
pd.set_option('display.width', None)
pd.set_option('display.max_colwidth', None)
```
### Set global Parameters
Set your parameters here:
data_path: In this path put the data you have downloaded with YouTube Data Tools.
output_path: Tghe files generated in this notebook will be saved here.
url_dict: URLs to models on Tensorflow hub are saved here. Other models are available there.
model_type: Define which model you would like to use. Choose one from url_dict
new_embeddings: If this is true, new embeddings will be generated and saved at output_path. Otherwise, embeddings are loaded from Disc.
```
data_path = './data/videoinfo_xnudgOC9D5Y_2020_11_24-21_12_03_comments.tab'
output_path = "./output/"
new_embeddings = True
url_dict = {
'Transformer' : "https://tfhub.dev/google/universal-sentence-encoder-large/5",
'DAN' : "https://tfhub.dev/google/universal-sentence-encoder/4",
'Transformer_Multilingual': "https://tfhub.dev/google/universal-sentence-encoder-multilingual-large/3"
}
model_type = 'Transformer' #@param ['DAN','Transformer','Transformer_Multilingual']
try:
os.mkdir(output_path)
except OSError:
print ("Creation of the directory %s failed" % output_path)
else:
print ("Successfully created the directory %s " % output_path)
if new_embeddings:
data = pd.read_csv(data_path,sep='\t',header=(0))
data.head()
if new_embeddings:
data = data.dropna(subset=['text', 'authorName']) # drop rows with no content
data=data.drop(['id', 'replyCount','likeCount','authorChannelUrl','authorChannelId','isReplyTo','isReplyToName'],axis=1) # drop unused columns
data.head()
if new_embeddings:
# Remove HTML tags
tqdm.pandas()
h = HTML2Text()
h.ignore_links = True
data['cleaned'] = data['text'].progress_apply(lambda x: h.handle(x))
print( "Removed HTML Tags.")
# Remove links
http_link_pattern = r'http\S+'
bitly_link_pattern = r'bit.ly/\S+'
data['cleaned'] = data['cleaned'].str.replace(http_link_pattern, '')
data['cleaned'] = data['cleaned'].str.replace(bitly_link_pattern, '')
print( "Removed Links.")
# Remove user names
keep_names = ["earth", "Tide", "Geologist", "A Person", "Titanic", "adventure", "Sun", "The United States Of America"] # user names we want to keep
user_names = [name for name in data['authorName'].unique() if (len(name)> 3 and name not in keep_names)]
data['cleaned'] = data['cleaned'].str.replace('|'.join(map(re.escape, user_names)), '')
print( "Removed user names.")
if new_embeddings:
data.to_pickle(output_path+'data_preprocessed'+'.pkl')
else:
data = pd.read_pickle(output_path+'data_preprocessed'+'.pkl')
data.head()
hub_url = url_dict[model_type]
if new_embeddings:
print("Loading model. This will take some time...")
embed = hub.load(hub_url)
# Produce embeddings of your documents.
if new_embeddings:
for k,g in data.groupby(np.arange(len(data))//200):
if k == 0:
embeddings = embed(g['cleaned'])
else:
embeddings_new = embed(g['cleaned'])
embeddings = tf.concat(values=[embeddings,embeddings_new],axis = 0)
print(k , end =" ")
print("The embeddings vector is of fixed length {}".format(embeddings.shape[1]))
np.save(output_path+'/embeddings'+model_type+'.npy', embeddings, allow_pickle=True, fix_imports=True)
else:
embeddings = np.load(output_path+'/embeddings'+model_type+'.npy', mmap_mode=None, allow_pickle=False, fix_imports=True, encoding='ASCII')
embeddings.shape
```
### Calculate Similarity Matrix with angular distance
'Following Cer et al. (2018), we first compute the sentence embeddings u, v for an STS sentence pair, and then score the sentence pair similarity based on the angular distance between the two embedding vectors d = − arccos (uv/||u|| ||v||).'
```
from sklearn.metrics.pairwise import cosine_similarity
def cos_sim(input_vectors):
similarity = cosine_similarity(input_vectors)
return similarity
cosine_similarity_matrix = cos_sim(np.array(embeddings))
print(cosine_similarity_matrix)
np.shape(cosine_similarity_matrix)
import seaborn as sns
def plot_similarity(labels, features, rotation):
corr = np.inner(features, features)
sns.set(font_scale=1.2)
g = sns.heatmap(
corr,
xticklabels=labels,
yticklabels=labels,
vmin=0,
vmax=1,
cmap="YlOrRd")
g.set_xticklabels(labels, rotation=rotation)
g.set_title("Semantic Textual Similarity")
num_samples = 5
off_set = 100
plot_similarity(data.iloc[off_set:off_set+num_samples]['cleaned'], embeddings[off_set:off_set+num_samples], 90)
# Define which comment to analyze
comment_index = 3
comment = data["cleaned"][comment_index]
comment_list = data["cleaned"].tolist()
print(comment)
# Print similar comments.
def get_top_similar(sentence, sentence_list, similarity_matrix, topN):
# find the index of sentence in list
index = sentence_list.index(sentence)
# get the corresponding row in similarity matrix
similarity_row = np.array(similarity_matrix[index, :])
# get the indices of top similar
indices = similarity_row.argsort()[-topN:][::-1]
return [sentence_list[i] for i in indices]
for i, value in enumerate(get_top_similar(comment, comment_list, cosine_similarity_matrix, 20)):
print("Top similar comment {}: {}".format(i+1, value))
```
| github_jupyter |
# Thread
Python threading allows you to have different parts of your program run concurrently and can simplify your design.
```
import logging # To create logs
from time import sleep
from threading import Thread
# Thread function
def thread_function(name):
logging.info("Thread %s: starting", name)
time.sleep(2)
logging.info("Thread %s: finishing", name)
# Main thread, all lines of code specified under its scope
# will be considered to be executed in main thread.
if __name__ == "__main__":
format = "%(asctime)s: %(message)s"
logging.basicConfig(format=format, level=logging.INFO, datefmt="%H:%M:%S")
logging.info("Main : before starting a thread")
# Creating instance of thread
x = Thread(target=thread_function, args=(1,))
logging.info("Main : before running the thread")
# Start the thread
x.start()
logging.info("Main : wait for the thread to finish")
x.join()
logging.info("Main : All operations are done")
print("Main - Started")
t = Thread(target = thread_function, args = ("New",))
print("Main - Before starting thread")
t.start()
print("Main - Waiting for thread to finish")
print("Main - Done")
print("Main - Started")
t = Thread(target = thread_function, args = ("New",))
print("Main - Before starting thread")
t.start()
print("Main - Waiting for thread to finish")
t.join()
print("Main - Done")
# Multiple Threading
if __name__ == "__main__":
format = "%(asctime)s: %(message)s"
logging.basicConfig(format=format, level=logging.INFO, datefmt="%H:%M:%S")
threads = list()
for i in range(5):
logging.info("Main : before starting a thread %d",i)
x = Thread(target=thread_function, args=(i,))
threads.append(x)
#Start the thread
x.start()
logging.info("Main : before running the thread")
print(threads)
for index,thread in enumerate(threads):
logging.info("Main thread : before joining the thread %d ",index)
thread.join()
logging.info("Main : thread %d done",index)
#Another multi thread example
threads = []
for index in range(3):
print("Main - Creating and started thread "+ str(index))
t = Thread(target = thread_function, args = (index,))
threads.append(t)
t.start()
for index, thread in enumerate(threads):
print("Main - Before joining thread "+ str(index))
thread.join()
print("Main - After joining thread "+ str(index))
```
### Daemon Threads
In computer science, a daemon is a process that runs in the background.
Python threading has a more specific meaning for daemon. A daemon thread will shut down immediately when the program exits. One way to think about these definitions is to consider the daemon thread a thread that runs in the background without worrying about shutting it down.
If a program is running Threads that are not daemons, then the program will wait for those threads to complete before it terminates. Threads that are daemons, however, are just killed wherever they are when the program is exiting.
```
if __name__ == "__main__":
format = "%(asctime)s: %(message)s"
logging.basicConfig(format=format, level=logging.INFO, datefmt="%H:%M:%S")
logging.info("Main : before starting a thread")
# Creating instance of thread
x = Thread(target=thread_function, args=(1,), daemon=True)
logging.info("Main : before running the thread")
# Start the thread
x.start()
logging.info("Main : wait for the thread to finish")
x.join()
logging.info("Main : All operations are done")
```
### ThreadPoolExecutor
Group Threads together. consider as a box where you add threads to. this will manage all your threads.
```
from time import sleep
from threading import Thread
from concurrent.futures import ThreadPoolExecutor
def thread_function(name):
print("Thread {0} started".format(name))
sleep(2)
print("Thread {0} ended".format(name))
with ThreadPoolExecutor(max_workers = 3) as executor:
executor.map(thread_function,range(6))
print("Done")
with ThreadPoolExecutor(max_workers = 3) as executor:
executor.submit(thread_function,1)
executor.submit(thread_function,2)
executor.submit(thread_function,3)
executor.submit(thread_function,4)
executor.submit(thread_function,5)
executor.submit(thread_function,6)
print("Done")
```
### Lock
mutex = mutual exclusion. We allow only one thread to read modify write code at a time
lock is like key to open door
basic functions: aquire() & .release()
```
from time import sleep
from threading import Thread, Lock
from concurrent.futures import ThreadPoolExecutor
class Counter:
def __init__(self):
self.value = 0
def update(self,name):
print("Update started on Thread :" + str(name)+" with self.value = " + str(self.value))
val = self.value
val +=1
self.value = val
print("Update ended on Thread :" + str(name) +" with self.value = " + str(self.value))
counter = Counter()
with ThreadPoolExecutor(max_workers = 3) as executor:
for index in range(3):
executor.submit(counter.update, index)
print("Done")
class CounterwithRaceCondition:
def __init__(self):
self.value = 0
def update(self,name):
print("Update started on Thread :" + str(name)+" with self.value = " + str(self.value))
val = self.value
val +=1
sleep(1)
self.value = val
print("Update ended on Thread :" + str(name) +" with self.value = " + str(self.value))
counter = CounterwithRaceCondition()
with ThreadPoolExecutor(max_workers = 3) as executor:
for index in range(3):
executor.submit(counter.update, index)
print("Done")
class CounterwithLock:
def __init__(self):
self.value = 0
self._lock = Lock()
def update(self,name):
print("Update started on Thread :" + str(name)+" with self.value = " + str(self.value))
with self._lock:
val = self.value
val +=1
sleep(1)
self.value = val
print("Update ended on Thread :" + str(name) +" with self.value = " + str(self.value))
counter = CounterwithLock()
with ThreadPoolExecutor(max_workers = 3) as executor:
for index in range(3):
executor.submit(counter.update, index)
print("Done")
class CounterwithLockAcquire:
def __init__(self):
self.value = 0
self._lock = Lock()
def update(self,name):
print("Update started on Thread :" + str(name)+" with self.value = " + str(self.value))
self._lock.acquire()
val = self.value
val +=1
sleep(1)
self.value = val
self._lock.release()
print("Update ended on Thread :" + str(name) +" with self.value = " + str(self.value))
counter = CounterwithLockAcquire()
with ThreadPoolExecutor(max_workers = 3) as executor:
for index in range(3):
executor.submit(counter.update, index)
print("Done")
```
| github_jupyter |
<center>
<img src="https://gitlab.com/ibm/skills-network/courses/placeholder101/-/raw/master/labs/module%201/images/IDSNlogo.png" width="300" alt="cognitiveclass.ai logo" />
</center>
# **SpaceX Falcon 9 First Stage Landing Prediction**
## Assignment: Exploring and Preparing Data
Estimated time needed: **70** minutes
In this assignment, we will predict if the Falcon 9 first stage will land successfully. SpaceX advertises Falcon 9 rocket launches on its website with a cost of 62 million dollars; other providers cost upward of 165 million dollars each, much of the savings is due to the fact that SpaceX can reuse the first stage.
In this lab, you will perform Exploratory Data Analysis and Feature Engineering.
Falcon 9 first stage will land successfully

Several examples of an unsuccessful landing are shown here:

Most unsuccessful landings are planned. Space X performs a controlled landing in the oceans.
## Objectives
Perform exploratory Data Analysis and Feature Engineering using `Pandas` and `Matplotlib`
* Exploratory Data Analysis
* Preparing Data Feature Engineering
***
### Import Libraries and Define Auxiliary Functions
We will import the following libraries the lab
```
# andas is a software library written for the Python programming language for data manipulation and analysis.
import pandas as pd
#NumPy is a library for the Python programming language, adding support for large, multi-dimensional arrays and matrices, along with a large collection of high-level mathematical functions to operate on these arrays
import numpy as np
# Matplotlib is a plotting library for python and pyplot gives us a MatLab like plotting framework. We will use this in our plotter function to plot data.
import matplotlib.pyplot as plt
#Seaborn is a Python data visualization library based on matplotlib. It provides a high-level interface for drawing attractive and informative statistical graphics
import seaborn as sns
```
## Exploratory Data Analysis
First, let's read the SpaceX dataset into a Pandas dataframe and print its summary
```
df=pd.read_csv("https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBM-DS0321EN-SkillsNetwork/datasets/dataset_part_2.csv")
# If you were unable to complete the previous lab correctly you can uncomment and load this csv
# df = pd.read_csv('https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-DS0701EN-SkillsNetwork/api/dataset_part_2.csv')
df.head(5)
```
First, let's try to see how the `FlightNumber` (indicating the continuous launch attempts.) and `Payload` variables would affect the launch outcome.
We can plot out the <code>FlightNumber</code> vs. <code>PayloadMass</code>and overlay the outcome of the launch. We see that as the flight number increases, the first stage is more likely to land successfully. The payload mass is also important; it seems the more massive the payload, the less likely the first stage will return.
```
sns.catplot(y="PayloadMass", x="FlightNumber", hue="Class", data=df, aspect = 5)
plt.xlabel("Flight Number",fontsize=20)
plt.ylabel("Pay load Mass (kg)",fontsize=20)
plt.show()
```
We see that different launch sites have different success rates. <code>CCAFS LC-40</code>, has a success rate of 60 %, while <code>KSC LC-39A</code> and <code>VAFB SLC 4E</code> has a success rate of 77%.
Next, let's drill down to each site visualize its detailed launch records.
### TASK 1: Visualize the relationship between Flight Number and Launch Site
Use the function <code>catplot</code> to plot <code>FlightNumber</code> vs <code>LaunchSite</code>, set the parameter <code>x</code> parameter to <code>FlightNumber</code>,set the <code>y</code> to <code>Launch Site</code> and set the parameter <code>hue</code> to <code>'class'</code>
```
# Plot a scatter point chart with x axis to be Flight Number and y axis to be the launch site, and hue to be the class value
sns.catplot(y='LaunchSite', x='FlightNumber', hue='Class', data=df)
plt.xlabel('Flight Number')
plt.ylabel('Launch Site')
plt.show()
```
Now try to explain the patterns you found in the Flight Number vs. Launch Site scatter point plots.
### TASK 2: Visualize the relationship between Payload and Launch Site
We also want to observe if there is any relationship between launch sites and their payload mass.
```
# Plot a scatter point chart with x axis to be Pay Load Mass (kg) and y axis to be the launch site, and hue to be the class value
sns.catplot(y='LaunchSite', x='PayloadMass', hue='Class', data=df)
plt.xlabel('Pay Load Mass (kg)')
plt.ylabel('Launch Site')
plt.show()
```
Now if you observe Payload Vs. Launch Site scatter point chart you will find for the VAFB-SLC launchsite there are no rockets launched for heavypayload mass(greater than 10000).
### TASK 3: Visualize the relationship between success rate of each orbit type
Next, we want to visually check if there are any relationship between success rate and orbit type.
Let's create a `bar chart` for the sucess rate of each orbit
```
# HINT use groupby method on Orbit column and get the mean of Class column
df_sorted = df.groupby('Orbit').mean()['Class'].reset_index().sort_values(['Class'], ascending=True)
fig, ax = plt.subplots()
ax.barh(df_sorted.Orbit, df_sorted.Class * 100)
plt.xlabel('Success Rate (%)')
plt.ylabel('Orbit Type')
plt.show()
```
Analyze the ploted bar chart try to find which orbits have high sucess rate.
### TASK 4: Visualize the relationship between FlightNumber and Orbit type
For each orbit, we want to see if there is any relationship between FlightNumber and Orbit type.
```
# Plot a scatter point chart with x axis to be FlightNumber and y axis to be the Orbit, and hue to be the class value
sns.catplot(x='FlightNumber', y='Orbit', hue='Class', data=df)
plt.xlabel('Fligh Number')
plt.ylabel('Orbit Type')
plt.show()
```
You should see that in the LEO orbit the Success appears related to the number of flights; on the other hand, there seems to be no relationship between flight number when in GTO orbit.
### TASK 5: Visualize the relationship between Payload and Orbit type
Similarly, we can plot the Payload vs. Orbit scatter point charts to reveal the relationship between Payload and Orbit type
```
# Plot a scatter point chart with x axis to be Payload and y axis to be the Orbit, and hue to be the class value
sns.catplot(x='PayloadMass', y='Orbit', hue='Class', data=df)
plt.xlabel('Pay load Mass (kg)')
plt.ylabel('Orbit Type')
plt.show()
```
With heavy payloads the successful landing or positive landing rate are more for Polar,LEO and ISS.
However for GTO we cannot distinguish this well as both positive landing rate and negative landing(unsuccessful mission) are both there here.
### TASK 6: Visualize the launch success yearly trend
You can plot a line chart with x axis to be <code>Year</code> and y axis to be average success rate, to get the average launch success trend.
The function will help you get the year from the date:
```
# A function to Extract years from the date
year=[]
def Extract_year(date):
for i in df["Date"]:
year.append(i.split("-")[0])
return year
# Plot a line chart with x axis to be the extracted year and y axis to be the success rate
df.groupby(Extract_year(df['Date'])).mean()['Class'].plot(kind='line')
plt.xlabel('Year')
plt.ylabel('Success Rate')
plt.show()
```
you can observe that the sucess rate since 2013 kept increasing till 2020
## Features Engineering
By now, you should obtain some preliminary insights about how each important variable would affect the success rate, we will select the features that will be used in success prediction in the future module.
```
features = df[['FlightNumber', 'PayloadMass', 'Orbit', 'LaunchSite', 'Flights', 'GridFins', 'Reused', 'Legs', 'LandingPad', 'Block', 'ReusedCount', 'Serial']]
features.head()
```
### TASK 7: Create dummy variables to categorical columns
Use the function <code>get_dummies</code> and <code>features</code> dataframe to apply OneHotEncoder to the column <code>Orbits</code>, <code>LaunchSite</code>, <code>LandingPad</code>, and <code>Serial</code>. Assign the value to the variable <code>features_one_hot</code>, display the results using the method head. Your result dataframe must include all features including the encoded ones.
```
# HINT: Use get_dummies() function on the categorical columns
features_one_hot = pd.get_dummies(features, columns=['Orbit', 'LaunchSite', 'LandingPad', 'Serial'])
features_one_hot.head()
```
### TASK 8: Cast all numeric columns to `float64`
Now that our <code>features_one_hot</code> dataframe only contains numbers cast the entire dataframe to variable type <code>float64</code>
```
# HINT: use astype function
features_one_hot.astype('float64')
```
We can now export it to a <b>CSV</b> for the next section,but to make the answers consistent, in the next lab we will provide data in a pre-selected date range.
<code>features_one_hot.to_csv('dataset_part\_3.csv', index=False)</code>
## Authors
<a href="https://www.linkedin.com/in/joseph-s-50398b136/?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDS0321ENSkillsNetwork26802033-2021-01-01">Joseph Santarcangelo</a> has a PhD in Electrical Engineering, his research focused on using machine learning, signal processing, and computer vision to determine how videos impact human cognition. Joseph has been working for IBM since he completed his PhD.
<a href="https://www.linkedin.com/in/nayefaboutayoun/?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDS0321ENSkillsNetwork26802033-2021-01-01">Nayef Abou Tayoun</a> is a Data Scientist at IBM and pursuing a Master of Management in Artificial intelligence degree at Queen's University.
## Change Log
| Date (YYYY-MM-DD) | Version | Changed By | Change Description |
| ----------------- | ------- | ------------- | ----------------------- |
| 2021-10-12 | 1.1 | Lakshmi Holla | Modified markdown |
| 2020-09-20 | 1.0 | Joseph | Modified Multiple Areas |
| 2020-11-10 | 1.1 | Nayef | updating the input data |
Copyright © 2020 IBM Corporation. All rights reserved.
| github_jupyter |
```
%matplotlib inline
import numpy as np
from sklearn.calibration import calibration_curve
from matplotlib import pyplot as plt
import pickle
import torch
from torch import nn
from torch.autograd import Variable
from model import Model
import utils
np.random.seed(42)
```
# Training Models
Run the following commands:
`python train_model.py --gpu_id 0 --mode dropout --model_id 0 --dprob 0.0`
`python train_model.py --gpu_id 0 --mode dropout --model_id 1 --dprob 0.1`
`python train_model.py --gpu_id 0 --mode dropout --model_id 2 --dprob 0.2`
`python train_model.py --gpu_id 0 --mode dropout --model_id 3 --dprob 0.3`
`python train_model.py --gpu_id 0 --mode dropout --model_id 4 --dprob 0.4`
`python train_model.py --gpu_id 0 --mode concrete --model_id 5 --dr 0.0001`
`python train_model.py --gpu_id 0 --mode concrete --model_id 6 --dr 1.0`
`python train_model.py --gpu_id 0 --mode concrete --model_id 7 --dr 100.0`
# Dropout Rate Convergence
```
plt.figure(figsize=(15, 3))
for index in range(5):
plt.subplot(1, 5, index + 1)
for training_data in cdropout_training_datas:
convergence = [x[5][index] for x in training_data]
plt.plot(convergence)
plt.title('L = {}'.format(index))
plt.xlabel('#epoch')
plt.ylim([0, 0.5])
plt.ylabel('p')
plt.tight_layout()
plt.show()
```
# Learning Curves
```
plt.figure(figsize=(15, 6))
plt.subplot(1, 2, 1)
for model, training_data, dropout_prob, color in zip(dropout_models, dropout_training_datas,
dropout_probs, ['r', 'g', 'b', 'c', 'm']):
train_scores = [x[2] for x in training_data]
val_scores = [x[4] for x in training_data]
plt.plot(np.arange(20, 100), train_scores[20:], color, label='train, p={}'.format(dropout_prob))
plt.plot(np.arange(20, 100), val_scores[20:], color + '--', label='val, p={}'.format(dropout_prob))
plt.legend()
plt.xlabel('#epoch')
plt.ylabel('Avg. Hamming distance')
plt.title('Dropout')
plt.subplot(1, 2, 2)
for model, training_data, dr, color in zip(cdropout_models, cdropout_training_datas,
cdropout_regs, ['r', 'g', 'b']):
train_scores = [x[2] for x in training_data]
val_scores = [x[4] for x in training_data]
plt.plot(np.arange(20, 100), train_scores[20:], color, label='train, dr={}'.format(dr))
plt.plot(np.arange(20, 100), val_scores[20:], color + '--', label='val, dr={}'.format(dr))
plt.legend()
plt.xlabel('#epoch')
plt.ylabel('Avg. Hamming distance')
plt.title('Concrete Dropout')
plt.show()
plt.figure(figsize=(15, 6))
plt.subplot(1, 2, 1)
for training_data, dropout_prob, color in zip(dropout_training_datas,
dropout_probs, ['r', 'g', 'b', 'c', 'm']):
train_scores = [x[2] for x in training_data]
val_scores = [x[4] for x in training_data]
dropout_execution_times = [x[0] for x in training_data]
plt.plot(dropout_execution_times[20:], train_scores[20:], color,
label='train, p={}'.format(dropout_prob))
plt.plot(dropout_execution_times[20:], val_scores[20:], color + '--',
label='val, p={}'.format(dropout_prob))
plt.legend()
plt.xlabel('min.')
plt.ylabel('Avg. Hamming distance')
plt.title('Dropout')
plt.subplot(1, 2, 2)
for training_data, dr, color in zip(cdropout_models, cdropout_training_datas,
cdropout_regs, ['r', 'g', 'b']):
train_scores = [x[2] for x in training_data]
val_scores = [x[4] for x in training_data]
dropout_execution_times = [x[0] for x in training_data]
plt.plot(dropout_execution_times[20:], train_scores[20:], color,
label='train, dr={}'.format(dr))
plt.plot(dropout_execution_times[20:], val_scores[20:], color + '--',
label='val, dr={}'.format(dr))
plt.legend()
plt.xlabel('min.')
plt.ylabel('Avg. Hamming distance')
plt.title('Concrete Dropout')
plt.show()
```
| github_jupyter |
```
# Copyright 2021 Google LLC
# Use of this source code is governed by an MIT-style
# license that can be found in the LICENSE file or at
# https://opensource.org/licenses/MIT.
# Author(s): Kevin P. Murphy (murphyk@gmail.com) and Mahmoud Soliman (mjs@aucegypt.edu)
```
<a href="https://opensource.org/licenses/MIT" target="_parent"><img src="https://img.shields.io/github/license/probml/pyprobml"/></a>
<a href="https://colab.research.google.com/github/probml/pyprobml/blob/master/notebooks/figures//AppendixE_figures.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Cloning the pyprobml repo
```
!git clone https://github.com/probml/pyprobml
%cd pyprobml/scripts
```
# Installing required software (This may take few minutes)
```
!apt-get install octave -qq > /dev/null
!apt-get install liboctave-dev -qq > /dev/null
%%capture
%load_ext autoreload
%autoreload 2
DISCLAIMER = 'WARNING : Editing in VM - changes lost after reboot!!'
from google.colab import files
def interactive_script(script, i=True):
if i:
s = open(script).read()
if not s.split('\n', 1)[0]=="## "+DISCLAIMER:
open(script, 'w').write(
f'## {DISCLAIMER}\n' + '#' * (len(DISCLAIMER) + 3) + '\n\n' + s)
files.view(script)
%run $script
else:
%run $script
def show_image(img_path):
from google.colab.patches import cv2_imshow
import cv2
img = cv2.imread(img_path, cv2.IMREAD_UNCHANGED)
img=cv2.resize(img,(600,600))
cv2_imshow(img)
```
## Figure E.1:<a name='E.1'></a> <a name='bootstrapDemoBer'></a>
Bootstrap (top row) vs Bayes (bottom row). The $N$ data cases were generated from $\mathrm Ber (\theta =0.7)$. Left column: $N=10$. Right column: $N=100$. (a-b) A bootstrap approximation to the sampling distribution of the MLE for a Bernoulli distribution. We show the histogram derived from $B=10,000$ bootstrap samples. (c-d) Histogram of 10,000 samples from the posterior distribution using a uniform prior.
Figure(s) generated by [bootstrapDemoBer.m](https://github.com/probml/pmtk3/blob/master/demos/bootstrapDemoBer.m)
```
!octave -W bootstrapDemoBer.m >> _
```
## Figure E.2:<a name='E.2'></a> <a name='samplingDistGaussShrinkage'></a>
Left: Sampling distribution of the MAP estimate (equivalent to the posterior mean) under a $\mathcal N (\theta _0=0,\sigma ^2/\kappa _0)$ prior with different prior strengths $\kappa _0$. (If we set $\kappa =0$, the MAP estimate reduces to the MLE.) The data is $n=5$ samples drawn from $\mathcal N (\theta ^*=1,\sigma ^2=1)$. Right: MSE relative to that of the MLE versus sample size. Adapted from Figure 5.6 of <a href='#Hoff09'>[Hof09]</a> .
Figure(s) generated by [samplingDistGaussShrinkage.m](https://github.com/probml/pmtk3/blob/master/demos/samplingDistGaussShrinkage.m)
```
!octave -W samplingDistGaussShrinkage.m >> _
```
## Figure E.3:<a name='E.3'></a> <a name='biasVarianceLinReg'></a>
Illustration of bias-variance tradeoff for ridge regression. We generate 100 data sets from the true function, shown in solid green. Left: we plot the regularized fit for 20 different data sets. We use linear regression with a Gaussian RBF expansion, with 25 centers evenly spread over the $[0,1]$ interval. Right: we plot the average of the fits, averaged over all 100 datasets. Top row: strongly regularized: we see that the individual fits are similar to each other (low variance), but the average is far from the truth (high bias). Bottom row: lightly regularized: we see that the individual fits are quite different from each other (high variance), but the average is close to the truth (low bias). Adapted from <a href='#BishopBook'>[Bis06]</a> Figure 3.5.
Figure(s) generated by [biasVarModelComplexity3.m](https://github.com/probml/pmtk3/blob/master/demos/biasVarModelComplexity3.m)
```
!octave -W biasVarModelComplexity3.m >> _
```
## Figure E.4:<a name='E.4'></a> <a name='fig:biasVarianceCartoon'></a>
Cartoon illustration of the bias variance tradeoff. From http://scott.fortmann-roe.com/docs/BiasVariance.html . Used with kind permission of Scott Fortmann-Roe.
```
show_image("/content/pyprobml/notebooks/figures/images/biasVarCartoon.png")
```
## Figure E.5:<a name='E.5'></a> <a name='riskFnGauss'></a>
Risk functions for estimating the mean of a Gaussian. Each curve represents $R( \theta _i(\cdot ),\theta ^*)$ plotted vs $\theta ^*$, where $i$ indexes the estimator. Each estimator is applied to $N$ samples from $\mathcal N (\theta ^*,\sigma ^2=1)$. The dark blue horizontal line is the sample mean (MLE); the red line horizontal line is the sample median; the black curved line is the estimator $ \theta =\theta _0=0$; the green curved line is the posterior mean when $\kappa =1$; the light blue curved line is the posterior mean when $\kappa =5$. (a) $N=5$ samples. (b) $N=20$ samples. Adapted from Figure B.1 of <a href='#Bernardo94'>[BS94]</a> .
Figure(s) generated by [riskFnGauss.m](https://github.com/probml/pmtk3/blob/master/demos/riskFnGauss.m)
```
!octave -W riskFnGauss.m >> _
```
## Figure E.6:<a name='E.6'></a> <a name='minimaxRisk'></a>
Risk functions for two decision procedures, $\pi _1$ and $\pi _2$. Since $\pi _1$ has lower worst case risk, it is the minimax estimator, even though $\pi _2$ has lower risk for most values of $\theta $. Thus minimax estimators are overly conservative.
```
show_image("/content/pyprobml/notebooks/figures/images/{minimaxRiskCurvesCropped}.png")
```
## Figure E.7:<a name='E.7'></a> <a name='fig:powerCurves'></a>
(a) Illustration of the Neyman-Pearson hypothesis testing paradigm.
Figure(s) generated by [neymanPearson2.m](https://github.com/probml/pmtk3/blob/master/demos/neymanPearson2.m) [twoPowerCurves.m](https://github.com/probml/pmtk3/blob/master/demos/twoPowerCurves.m)
```
!octave -W neymanPearson2.m >> _
!octave -W twoPowerCurves.m >> _
```
## References:
<a name='Bernardo94'>[BS94]</a> J. Bernardo and A. Smith. "Bayesian Theory". (1994).
<a name='BishopBook'>[Bis06]</a> C. Bishop "Pattern recognition and machine learning". (2006).
<a name='Hoff09'>[Hof09]</a> P. Hoff "A First Course in Bayesian Statistical Methods". (2009).
| github_jupyter |
# Credit Risk Classification
Credit risk poses a classification problem that’s inherently imbalanced. This is because healthy loans easily outnumber risky loans. In this Challenge, you’ll use various techniques to train and evaluate models with imbalanced classes. You’ll use a dataset of historical lending activity from a peer-to-peer lending services company to build a model that can identify the creditworthiness of borrowers.
## Instructions:
This challenge consists of the following subsections:
* Split the Data into Training and Testing Sets
* Create a Logistic Regression Model with the Original Data
* Predict a Logistic Regression Model with Resampled Training Data
### Split the Data into Training and Testing Sets
Open the starter code notebook and then use it to complete the following steps.
1. Read the `lending_data.csv` data from the `Resources` folder into a Pandas DataFrame.
2. Create the labels set (`y`) from the “loan_status” column, and then create the features (`X`) DataFrame from the remaining columns.
> **Note** A value of `0` in the “loan_status” column means that the loan is healthy. A value of `1` means that the loan has a high risk of defaulting.
3. Check the balance of the labels variable (`y`) by using the `value_counts` function.
4. Split the data into training and testing datasets by using `train_test_split`.
### Create a Logistic Regression Model with the Original Data
Employ your knowledge of logistic regression to complete the following steps:
1. Fit a logistic regression model by using the training data (`X_train` and `y_train`).
2. Save the predictions on the testing data labels by using the testing feature data (`X_test`) and the fitted model.
3. Evaluate the model’s performance by doing the following:
* Calculate the accuracy score of the model.
* Generate a confusion matrix.
* Print the classification report.
4. Answer the following question: How well does the logistic regression model predict both the `0` (healthy loan) and `1` (high-risk loan) labels?
### Predict a Logistic Regression Model with Resampled Training Data
Did you notice the small number of high-risk loan labels? Perhaps, a model that uses resampled data will perform better. You’ll thus resample the training data and then reevaluate the model. Specifically, you’ll use `RandomOverSampler`.
To do so, complete the following steps:
1. Use the `RandomOverSampler` module from the imbalanced-learn library to resample the data. Be sure to confirm that the labels have an equal number of data points.
2. Use the `LogisticRegression` classifier and the resampled data to fit the model and make predictions.
3. Evaluate the model’s performance by doing the following:
* Calculate the accuracy score of the model.
* Generate a confusion matrix.
* Print the classification report.
4. Answer the following question: How well does the logistic regression model, fit with oversampled data, predict both the `0` (healthy loan) and `1` (high-risk loan) labels?
### Write a Credit Risk Analysis Report
For this section, you’ll write a brief report that includes a summary and an analysis of the performance of both machine learning models that you used in this challenge. You should write this report as the `README.md` file included in your GitHub repository.
Structure your report by using the report template that `Starter_Code.zip` includes, and make sure that it contains the following:
1. An overview of the analysis: Explain the purpose of this analysis.
2. The results: Using bulleted lists, describe the balanced accuracy scores and the precision and recall scores of both machine learning models.
3. A summary: Summarize the results from the machine learning models. Compare the two versions of the dataset predictions. Include your recommendation for the model to use, if any, on the original vs. the resampled data. If you don’t recommend either model, justify your reasoning.
```
# Import the modules
import numpy as np
import pandas as pd
from pathlib import Path
from sklearn.metrics import balanced_accuracy_score
from sklearn.metrics import confusion_matrix
from imblearn.metrics import classification_report_imbalanced
from sklearn.metrics import classification_report
import warnings
warnings.filterwarnings('ignore')
```
---
## Split the Data into Training and Testing Sets
### Step 1: Read the `lending_data.csv` data from the `Resources` folder into a Pandas DataFrame.
```
# Read the CSV file from the Resources folder into a Pandas DataFrame
lending_df = pd.read_csv(
Path("Resources/lending_data.csv")
)
# Review the DataFrame
lending_df.head()
```
### Step 2: Create the labels set (`y`) from the “loan_status” column, and then create the features (`X`) DataFrame from the remaining columns.
```
# Separate the data into labels and features
#Review 12.3.4
# Separate the y variable, the labels
y = lending_df["loan_status"]
# Separate the X variable, the features
X = lending_df.drop(columns=['loan_status'])
# Review the y variable Series
y.head()
# Review the X variable DataFrame
X.head()
```
### Step 3: Check the balance of the labels variable (`y`) by using the `value_counts` function.
```
# Check the balance of our target values
y.value_counts()
```
### Step 4: Split the data into training and testing datasets by using `train_test_split`.
```
# Import the train_test_learn module
from sklearn.model_selection import train_test_split
# Split the data using train_test_split
# Assign a random_state of 1 to the function
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=1)
```
---
## Create a Logistic Regression Model with the Original Data
### Step 1: Fit a logistic regression model by using the training data (`X_train` and `y_train`).
```
# Import the LogisticRegression module from SKLearn
from sklearn.linear_model import LogisticRegression
# Instantiate the Logistic Regression model
# Assign a random_state parameter of 1 to the model
logistic_regression_model = LogisticRegression(random_state=1)
# Fit the model using training data
lr_model = logistic_regression_model.fit(X_train, y_train)
lr_model
```
### Step 2: Save the predictions on the testing data labels by using the testing feature data (`X_test`) and the fitted model.
```
# Make a prediction using the testing data
#ASK HERE: USE LOGISTIC OR Lr_modul?
testing_predictions = logistic_regression_model.predict(X_test)
```
### Step 3: Evaluate the model’s performance by doing the following:
* Calculate the accuracy score of the model.
* Generate a confusion matrix.
* Print the classification report.
```
# Print the balanced_accuracy score of the model
baso = balanced_accuracy_score(y_test, testing_predictions)
print(baso)
# Generate a confusion matrix for the model
test_matrix = confusion_matrix(y_test, testing_predictions)
print(test_matrix)
# Print the classification report for the model
testing_report = classification_report_imbalanced(y_test, testing_predictions)
print(testing_report)
```
### Step 4: Answer the following question.
**Question:** How well does the logistic regression model predict both the `0` (healthy loan) and `1` (high-risk loan) labels?
**Answer:** The logistic regression model predicts well on booth "0" and "1" labels. We can observe that from the classification report that the precision for "0" is 100% and the precision for "1" is 85%. The recall is also pretty good for "0" as 99% is correct, and "1" as 91% correct. F1-score is also high for both healthy and high-risk loan, which demonstrates that the model performs well. The testing data score is very high, even though it can be misleading.
---
## Predict a Logistic Regression Model with Resampled Training Data
### Step 1: Use the `RandomOverSampler` module from the imbalanced-learn library to resample the data. Be sure to confirm that the labels have an equal number of data points.
```
# Import the RandomOverSampler module form imbalanced-learn
from imblearn.over_sampling import RandomOverSampler
# Instantiate the random oversampler model
# # Assign a random_state parameter of 1 to the model
#Review 12.4.4
random_oversampler = RandomOverSampler(random_state=1)
# Fit the original training data to the random_oversampler model
X_resampled, y_resampled = random_oversampler.fit_resample(X_train, y_train)
# Count the distinct values of the resampled labels data
y_resampled.value_counts()
```
### Step 2: Use the `LogisticRegression` classifier and the resampled data to fit the model and make predictions.
```
# Instantiate the Logistic Regression model
# Assign a random_state parameter of 1 to the model
model = LogisticRegression(random_state=1)
# Fit the model using the resampled training data
lr_resampled_model = model.fit(X_resampled, y_resampled)
lr_resampled_model
# Make a prediction using the testing data
y_resampled_pred = lr_resampled_model.predict(X_test)
```
### Step 3: Evaluate the model’s performance by doing the following:
* Calculate the accuracy score of the model.
* Generate a confusion matrix.
* Print the classification report.
```
# Print the balanced_accuracy score of the model
basr = balanced_accuracy_score(y_test, y_resampled_pred)
print(basr)
# Generate a confusion matrix for the model
confusion_matrix(y_test, y_resampled_pred)
# Print the classification report for the model
print(classification_report_imbalanced(y_test, y_resampled_pred))
```
### Step 4: Answer the following question
```
print(classification_report(y_test, testing_predictions))
print ("-----------------------------------------------------------------------------------------------------")
print(classification_report(y_test, y_resampled_pred))
```
**Question:** How well does the logistic regression model, fit with oversampled data, predict both the `0` (healthy loan) and `1` (high-risk loan) labels?
**Answer:** The logistic regression model fitted with the oversampled data predicts very well on both healthy and high-risk loan labels. We can observe from the classification report that the precision for "0" is 100% and the precision for "1" is 84%. The recall is also pretty good for "0" as 99% is correct, and "1" as 99% correct. F1-score is also high for both healthy and high-risk loan, which demonstrates that the model performs well. The testing data score is very high, even though it can be misleading.
Comparing with the logistic regression model fitted with test data, the logistic regression model fitted with oversampled data predicts better. From the prints above, we can find that the model with oversampled data performs better than the model fitted with test data by a higher correctness in recall (0.99 > 0.91) and F1 score (0.91 > 0.88) on "1" (high-risk loan). Even though the model fitted with oversampled data underperforms slightly on precision (0.84 < 0.85) than the model fitted with test data, the model fitted with oversampled data also overall outperforms than the model fitted with test data.
Since we are more interested in the prediction on "1" (high-risk loan with label), the logistic regression model fitted with oversampled data probably a better choice.
| github_jupyter |
# Introduction to Python
This notebook is primarily focused on introducing the specifics of using Python in an interactive environment such as Cloud Datalab. It is not intended to provide a complete tutorial to Python as a language. If you're completely new to Python, no problem! Python is quite straightforward, and there are lots of resources. The interactive step-by-step material at [Codecademy](https://www.codecademy.com/tracks/python) might be of interest.
To get started, below is a code cell that contains a statement of Python. You can run it by pressing `Shift+Enter` or clicking the the `Run` toolbar command with the cell selected.
```
print "Hello World"
```
You can edit the cell above and re-execute it to iterate over it. You can also add additional code cells to enter new blocks of code.
```
import sys
number = 10
def square(n):
return n * n
```
The cell above created a variable named `number` and a function named `square`, and placed them into the _global namespace_. It also imported the `sys` module into the same namespace. This global namespace is shared across all the cells in the notebook.
As a result, the following cell should be able to access them (as well as modify them).
```
print 'The number is currently %d' % number
number = 11
sys.stderr.write('And now it is %d' % number)
square(number)
```
By now you've probably noticed a few interesting things about code cells.
* Upon execution, their results are shown inline, in the notebook, after the code that produced the results. These results are included into the saved notebook. Results include outputs of print statements (text that might have been written out to stdout as well as stderr), and the final result of the cell.
* Some code cells do not have any visible output.
* Visually, code cells have a distinguishing border on the left. This border is washed out when the notebook is first loaded, indicating a cell has not been run yet, and then has a filled blue border to indicate it has been run.
## Getting Help
Python APIs are usually accompanied by documentation. You can use `?` to invoke help on a class or a method. For example, execute the cells below:
```
str?
g = globals()
g.get?
```
When run, these cells produce docstrings content that is displayed in the help pane within the sidebar.
Additionally, the code cells also provide auto-suggest. For example, press `Shift+Tab` after the '.' to see a list of members callable on the `g` variable that was just declared.
```
# Intentionally, incomplete for purposes of auto-suggest demo, rather than running unmodified.
g.
```
Function signature help is also available. For example, press `Tab` in between the empty parens below.
```
str()
```
Note that help in Python is based on the interpreter being able to resolve the type of the expression you are invoking help on.
Sometimes, you may not have executed code yet, but can get help by invoking it directly on the class or method you're interested in, rather than the variable itself. Try this.
```
import datetime
datetime.datetime?
```
# Python Libraries
Cloud Datalab includes the standard python library and a set of libraries that you can easily import. Most of the libraries were installed using pip, the Python package manager.
```
%%bash
pip list
```
If you have suggestions for additional packages to include, please do submit feedback, so it may be considered for inclusion in a future version.
## Installing a Python Library
You can use `pip` to install your own python libraries.
Keep in mind, that this will install the library within the virtual machine instance being used for Cloud Datalab, and will become available to all notebooks and all users sharing the same instance.
It is also temporary. If the virtual machine instance is recreated, you will need to reinstall the library.
The example below, installs [scrapy](http://scrapy.org/), a library that helps in scraping web content.
```
%%bash
apt-get install -y -q libxslt-dev libxml2-dev
pip install -q scrapy
```
| github_jupyter |
# Using Tensorflow DALI plugin: using various readers
### Overview
This example shows how different readers could be used to interact with Tensorflow. It shows how flexible DALI is.
The following readers are used in this example:
- MXNetReader
- CaffeReader
- FileReader
- TFRecordReader
For details on how to use them please see other [examples](../../index.rst).
Let us start with defining some global constants
`DALI_EXTRA_PATH` environment variable should point to the place where data from [DALI extra repository](https://github.com/NVIDIA/DALI_extra) is downloaded. Please make sure that the proper release tag is checked out.
```
import os.path
test_data_root = os.environ['DALI_EXTRA_PATH']
# MXNet RecordIO
db_folder = os.path.join(test_data_root, 'db', 'recordio/')
# Caffe LMDB
lmdb_folder = os.path.join(test_data_root, 'db', 'lmdb')
# image dir with plain jpeg files
image_dir = "../../data/images"
# TFRecord
tfrecord = os.path.join(test_data_root, 'db', 'tfrecord', 'train')
tfrecord_idx = "idx_files/train.idx"
tfrecord2idx_script = "tfrecord2idx"
N = 8 # number of GPUs
BATCH_SIZE = 128 # batch size per GPU
ITERATIONS = 32
IMAGE_SIZE = 3
```
Create idx file by calling `tfrecord2idx` script
```
from subprocess import call
import os.path
if not os.path.exists("idx_files"):
os.mkdir("idx_files")
if not os.path.isfile(tfrecord_idx):
call([tfrecord2idx_script, tfrecord, tfrecord_idx])
```
Let us define:
- common part of pipeline, other pipelines will inherit it
```
from nvidia.dali.pipeline import Pipeline
import nvidia.dali.ops as ops
import nvidia.dali.types as types
class CommonPipeline(Pipeline):
def __init__(self, batch_size, num_threads, device_id):
super(CommonPipeline, self).__init__(batch_size, num_threads, device_id)
self.decode = ops.ImageDecoder(device = "mixed", output_type = types.RGB)
self.resize = ops.Resize(device = "gpu",
interp_type = types.INTERP_LINEAR)
self.cmn = ops.CropMirrorNormalize(device = "gpu",
dtype = types.FLOAT,
crop = (227, 227),
mean = [128., 128., 128.],
std = [1., 1., 1.])
self.uniform = ops.random.Uniform(range = (0.0, 1.0))
self.resize_rng = ops.random.Uniform(range = (256, 480))
def base_define_graph(self, inputs, labels):
images = self.decode(inputs)
images = self.resize(images, resize_shorter = self.resize_rng())
output = self.cmn(images, crop_pos_x = self.uniform(),
crop_pos_y = self.uniform())
return (output, labels.gpu())
```
- MXNetReaderPipeline
```
from nvidia.dali.pipeline import Pipeline
import nvidia.dali.ops as ops
import nvidia.dali.types as types
class MXNetReaderPipeline(CommonPipeline):
def __init__(self, batch_size, num_threads, device_id, num_gpus):
super(MXNetReaderPipeline, self).__init__(batch_size, num_threads, device_id)
self.input = ops.MXNetReader(path = [db_folder+"train.rec"], index_path=[db_folder+"train.idx"],
random_shuffle = True, shard_id = device_id, num_shards = num_gpus)
def define_graph(self):
images, labels = self.input(name="Reader")
return self.base_define_graph(images, labels)
```
- CaffeReadPipeline
```
class CaffeReadPipeline(CommonPipeline):
def __init__(self, batch_size, num_threads, device_id, num_gpus):
super(CaffeReadPipeline, self).__init__(batch_size, num_threads, device_id)
self.input = ops.CaffeReader(path = lmdb_folder,
random_shuffle = True, shard_id = device_id, num_shards = num_gpus)
def define_graph(self):
images, labels = self.input()
return self.base_define_graph(images, labels)
```
- FileReadPipeline
```
class FileReadPipeline(CommonPipeline):
def __init__(self, batch_size, num_threads, device_id, num_gpus):
super(FileReadPipeline, self).__init__(batch_size, num_threads, device_id)
self.input = ops.FileReader(file_root = image_dir)
def define_graph(self):
images, labels = self.input()
return self.base_define_graph(images, labels)
```
- TFRecordPipeline
```
import nvidia.dali.tfrecord as tfrec
class TFRecordPipeline(CommonPipeline):
def __init__(self, batch_size, num_threads, device_id, num_gpus):
super(TFRecordPipeline, self).__init__(batch_size, num_threads, device_id)
self.input = ops.TFRecordReader(path = tfrecord,
index_path = tfrecord_idx,
features = {"image/encoded" : tfrec.FixedLenFeature((), tfrec.string, ""),
"image/class/label": tfrec.FixedLenFeature([1], tfrec.int64, -1)
})
def define_graph(self):
inputs = self.input()
images = inputs["image/encoded"]
labels = inputs["image/class/label"]
return self.base_define_graph(images, labels)
```
Now let us create function which builds pipeline on demand:
```
import tensorflow as tf
import nvidia.dali.plugin.tf as dali_tf
try:
from tensorflow.compat.v1 import GPUOptions
from tensorflow.compat.v1 import ConfigProto
from tensorflow.compat.v1 import Session
from tensorflow.compat.v1 import placeholder
except:
# Older TF versions don't have compat.v1 layer
from tensorflow import GPUOptions
from tensorflow import ConfigProto
from tensorflow import Session
from tensorflow import placeholder
try:
tf.compat.v1.disable_eager_execution()
except:
pass
def get_batch_test_dali(batch_size, pipe_type):
pipe_name, label_type, _ = pipe_type
pipes = [pipe_name(batch_size=batch_size, num_threads=2, device_id = device_id, num_gpus = N) for device_id in range(N)]
daliop = dali_tf.DALIIterator()
images = []
labels = []
for d in range(N):
with tf.device('/gpu:%i' % d):
image, label = daliop(pipeline = pipes[d],
shapes = [(BATCH_SIZE, 3, 227, 227), ()],
dtypes = [tf.int32, label_type],
device_id = d)
images.append(image)
labels.append(label)
return [images, labels]
```
At the end let us test if all pipelines have been correctly built and run with TF session
```
import numpy as np
pipe_types = [[MXNetReaderPipeline, tf.float32, (0, 999)],
[CaffeReadPipeline, tf.int32, (0, 999)],
[FileReadPipeline, tf.int32, (0, 1)],
[TFRecordPipeline, tf.int64, (1, 1000)]]
for pipe_name in pipe_types:
print ("RUN: " + pipe_name[0].__name__)
test_batch = get_batch_test_dali(BATCH_SIZE, pipe_name)
x = placeholder(tf.float32, shape=[BATCH_SIZE, IMAGE_SIZE, IMAGE_SIZE, 3], name='x')
gpu_options = GPUOptions(per_process_gpu_memory_fraction=0.8)
config = ConfigProto(gpu_options=gpu_options)
with Session(config=config) as sess:
for i in range(ITERATIONS):
imgs, labels = sess.run(test_batch)
# Testing correctness of labels
for label in labels:
## labels need to be integers
assert(np.equal(np.mod(label, 1), 0).all())
## labels need to be in range pipe_name[2]
assert((label >= pipe_name[2][0]).all())
assert((label <= pipe_name[2][1]).all())
print("OK : " + pipe_name[0].__name__)
```
| github_jupyter |
```
import pandas as pd
import numpy as np
from fbprophet import Prophet
import matplotlib.pyplot as plt
import pickle
%matplotlib inline
df_train = pd.read_csv('./../../train/KPI/train_7c189dd36f048a6c.csv')
df_test = pd.read_csv('./../../test/KPI/test_7c189dd36f048a6c.csv')
df_train = df_train.drop(['KPI ID','label'],axis=1)
df_test = df_test.drop(['KPI ID'],axis=1)
df_train['ds'] = df_train['timestamp']
df_train['y'] = df_train['value']
df_test['ds'] = df_test['timestamp']
df_test = df_test.drop(['timestamp'], axis=1)
df_test['y'] = df_test['value']
df_test = df_test.drop(['value'], axis=1)
df_train = df_train.drop(['timestamp','value'],axis=1)
df_test
model = Prophet()
model.fit(df_test)
pickle.dump(model, open('./../../PICKLE/fb_test_'+ KPI + '.pkl',"wb"))
forecast = model.predict(df_test)
forecast.yhat_upper.max()
fig1 = model.plot(forecast)
```
## Plot
```
forecast = forecast.set_index(['ds'])
dataset = pd.DataFrame(forecast['yhat'], index=forecast.index)
df_test['yhat'] = dataset['yhat']
df_test
df_test.head(40*1440).plot(kind='line',figsize=(14,8))
# plt.plot(dataset['yhat'],alpha=.5)
sup = df_test.head(40*1440)
pred = np.where( abs(sup.yhat - sup.value) > 900, 1, 0)
sup['pred'] = pred
sup.groupby(sup.pred).count()
sup['value'].plot(kind='line',figsize=(14,8))
plt.plot(sup['pred']*500,alpha=.5)
# pickle.dump(model, open('./../../PICKLE/fb_7c189dd36f048a6c.pkl',"wb"))
# model2 = pickle.loads(save_model)
# model2.plot(forecast);
```
## Generic
```
input_dir_train = './../../train/KPI/train_'
input_dir_test = './../../test/KPI/test_'
KPI_arr = pd.read_csv('./../../KPI/KPI.csv').values
for KPI in KPI_arr:
# df_train = pd.read_csv(input_dir_train + KPI[0] + '.csv')
df_test = pd.read_csv(input_dir_test + KPI[0] + '.csv')
# Format datasets
# df_train = df_train.drop(['KPI ID','label'],axis=1)
df_test = df_test.drop(['KPI ID'],axis=1)
# df_train['ds'] = df_train['timestamp']
# df_train['y'] = df_train['value']
# df_train = df_train.drop(['timestamp','value'],axis=1)
df_test['ds'] = df_test['timestamp']
df_test['y'] = df_test['value']
df_test = df_test.drop(['timestamp','value'], axis=1)
# Train with Train Data
# model = Prophet()
# model.fit(df_train)
# pickle.dump(model, open('./../../PICKLE/fb_train_'+ KPI[0] + '.pkl',"wb"))
# Train with Test Data
model = Prophet()
model.fit(df_test)
pickle.dump(model, open('./../../PICKLE/fb_test_'+ KPI[0] + '.pkl',"wb"))
# forecast = model.predict(df_test)
```
## Single Model
```
df_test
input_dir_train = './../../train/KPI/train_'
input_dir_test = './../../test/KPI/test_'
KPI = ['1c35dbf57f55f5e4']
# KPI_arr = pd.read_csv('./../../KPI/KPI.csv').values
# df_train = pd.read_csv(input_dir_train + KPI[0] + '.csv')
df_test = pd.read_csv(input_dir_test + KPI[0] + '.csv')
print(df_test.shape)
df_test = df_test[ df_test['value'] < 2100]
df_test = df_test[ df_test['value'] > 1000]
print(df_test.shape)
# Format datasets
# df_train = df_train.drop(['KPI ID','label'],axis=1)
df_test = df_test.drop(['KPI ID'],axis=1)
df_test['ds'] = df_test['timestamp']
df_test['y'] = df_test['value']
df_test = df_test.drop(['timestamp','value'], axis=1)
# Train with Train Data
# model = Prophet()
# model.fit(df_train)
# pickle.dump(model, open('./../../PICKLE/fb_train_'+ KPI[0] + '.pkl',"wb"))
# Train with Test Data
model = Prophet()
model.fit(df_test)
pickle.dump(model, open('./../../PICKLE/fb_test_'+ KPI[0] + '.pkl',"wb"))
input_dir_test = './../../test/KPI/test_'
df_test = pd.read_csv(input_dir_test + '1c35dbf57f55f5e4' + '.csv')
df_test
```
| github_jupyter |
## Informal Introduction
Recursion Relation is also know as **Recursive Formula**. Informal definition of recursion relation is the way to define series that each item defined by the previous items. But, to construct a complete recursion series, we need to define initial terms as precondition at first place. By its construction, it is easier to define rather than to answer what value of n in the series.
* Fibonacci recurrence relation is:<br>
$$F(0) = 0, F(1) = 1, F(n) = F(n-1) + F(n-2) \ \text{if} \ n \geq 2$$<br>
Formula for \\(F(n)\\) which involve only n is:<br>
$$F(n) = \bigl(\frac{\Phi^n - (-\phi)^n)}{\sqrt5}\bigr) \ \text{where} \ \Phi = \frac{(\sqrt5+1)}{2} \text{and} \ \phi=\frac{(\sqrt5-1)}{2}$$
<br><br>
* Find unique **derangements** of \\(n\\) people sit in \\(\{s_i, s_j, ..., s_n\}\\) where \\(i \neq j\\).
This problem has direct factorial formula (defined in Taylor series):<br>
$$D(n) = n!(\frac{1}{2!} - \frac{1}{3!} + \frac{1}{4!} - ... \frac{(-1)^n}{n!}), \ if \ n > 2$$<br>
The recursion formula is:<br>
$$D(n) = (n - 1) (D(n-1) + D(n-2))$$<br>
Formula for \\(D(n)\\) is:<br>
$$D(n) = n D(n-1) + (-1)^n$$
Further introductory can be explored in [Recurrence Relations & Generating Functions by R. Knott](http://www.maths.surrey.ac.uk/hosted-sites/R.Knott/Fibonacci/LRGF.html)
## Geometric Series
For \\(r \neq 1\\), the sum of the first n of geometric series is:
$$a+ar+ar^{2}+ar^{3}+ \cdots +ar^{n-1}=\sum_{k=0}^{n-1}ar^{k}=a\left({\frac{1-r^{n}}{1-r}}\right)$$
**PROOF**: *a* is initial term and *r* is common ratio.
$$\eqalign{
s&=a+ar+ar^{2}+ar^{3}+\cdots +ar^{n-1},\\
rs&=ar+ar^{2}+ar^{3}+ar^{4}+\cdots +ar^{n},\\
s-rs&=a-ar^{n},\\s(1-r)&=a(1-r^{n}),
}$$
So:
$${\displaystyle s=a\left({\frac {1-r^{n}}{1-r}}\right)\quad {\text{(if }}r\neq 1{\text{)}}.}$$
As *n* goes to infinity, the absolute value of *r* must be less than one for the series to converge. The sum then becomes:
$$a+ar+ar^{2}+ar^{3}+ar^{4}+\cdots =\sum _{k=0}^{\infty }ar^{k}={\frac {a}{1-r}},{\text{ for }}|r|<1.$$
When *a = 1*, this can be simplified to:
$$1\,+\,r\,+\,r^{2}\,+\,r^{3}\,+\,\cdots \;=\;{\frac {1}{1-r}},$$
the left-hand side being a geometric series with common ratio *r*.
The formula also holds for complex *r*, with the corresponding restriction, the modulus of *r* is strictly less than one.
**Proof of Convergence**
We can prove that the geometric series converges using the sum formula for a geometric progression:
$${\displaystyle {\begin{aligned}1+r+r^{2}+r^{3}+\cdots \ &=\lim _{n\rightarrow \infty }\left(1+r+r^{2}+\cdots +r^{n}\right)\\&=\lim _{n\rightarrow \infty }{\frac {1-r^{n+1}}{1-r}}.\end{aligned}}}$$
Since \\((1 + r + r^2 + ... + r^n)(1−r) = 1−r^{n+1}\\) and \\(r^{n+1} \rightarrow 0 \ \text{for} \ |r| < 1\\)
Convergence of geometric series can also be demonstrated by rewriting the series as an equivalent telescoping series. Consider the function,
$$g(K)={\frac {r^{K}}{1-r}}$$
Note that
$${\displaystyle 1=g(0)-g(1)\ ,\ r=g(1)-g(2)\ ,\ r^{2}=g(2)-g(3)\ ,\ldots }$$
Thus,
$${\displaystyle S=1+r+r^{2}+r^{3}+\cdots =(g(0)-g(1))+(g(1)-g(2))+(g(2)-g(3))+\cdots .}$$
If,
$$|r| < 1$$
then
$$g(K)\longrightarrow 0{\text{ as }}K\to \infty$$
so *S* converges to:
$$g(0)={\frac {1}{1-r}}$$
## Binomial Theorem (Factorial Method)
Let \\(n\\) be a positive integer, and \\(x\\) and \\(y\\) real numbers (or complex numbers, or polynomials). The coefficient of \\(x^ky^{n-k}\\), in \\(k^{th}\\) term in the expansion of \\((x+y)^n\\), is equal to \\({n \choose k}\\), where:
$${n \choose k} = \frac{n!}{(n-k)!k!}$$
So the **Binomial Expansion** is:
$$(x+y)^n = \sum_{r=0}^n {n \choose r} x^{n-r} y^r = \sum_{r = 0}^n {n \choose r} x^r y^{n-r}$$
**PROOF**:
$$
\begin{align}
(x+y)^n &= (x+y)(x+y)^{n-1} \\
&= (x+y)\bigg(\binom{n-1}{0} x^{n-1} + \binom{n-1}{1} x^{n-2}y + \cdots + \binom{n-1}{n-1}y^{n-1}\bigg) \\
&= x^n + \left( \binom{n-1}{0} + \binom{n-1}{1} \right) x^{n-1}y + \left( \binom{n-1}{1} + \binom{n-1}{2} \right) x^{n-2}y^2 \phantom{=} + \cdots + \left(\binom{n-1}{n-2} + \binom{n-1}{n-1} \right) xy^{n-1} + y^n \\
\end{align}
$$
and now Pascal's identity applies:
$$\binom{n-1}{k-1}+\binom{n-1}{k} = \binom{n}{k}$$
So the right side simplifies to:
$$x^n + \binom{n}{1} x^{n-1}y + \binom{n}{2} x^{n-2}y^2 + \cdots + \binom{n}{n-1}xy^{n-1} + y^n$$
## First Order Recurrences
The simplest type of recurrence reduce immediately to a product. The recurrence:
$$a_n = x_na_{n-1} \qquad \text{for} \ n>0 \ \text{with} \ a_0 = 1$$
equivalent to:
$$a_n = \prod_{1 \leq k \leq n} x_k$$
**Theorem 2.1 (First Order linear Recurrences)**
The recurrence
$$a_n = x_na_{n-1} + y_n \qquad \text{for} \ n>0 \ \text{with} \ a_0 = 0$$
has the explicit solution:
$$a_n = y_n + \sum_{1 \leq j < n}y_jx_{j+1}x_{j+2}...x_n$$
**PROOF**. Dividing both sides by \\(x_nx_{n-1}...x_1\\) and iterating, we have:
$$
\eqalign{
a_n &= x_nx_{n-1}...x_1 \sum_{1 \leq j \leq n}\frac{y_j}{x_jx_{j-1}...x_1}\\
&= y_n + \sum_{1 \leq j < n}y_jx_{j+1}x_{j+2}...x_n
}
$$
## Nonlinear First-Order Recurrences
**Simple convergence**. One convincing reason to calculate initial values is that many recurrences with a complicated appearance simply converge to a constant. For example:
$$a_n = \frac{a}{(1+a_{n-1})} \qquad for \ n>0 \ with \ a_0 = 1$$
This is a so-called continued fraction equation, which is discussed in §2.5. By calculating initial values, we can guess that the recurrence converges to a constant:

Each iteration increases the number of significant digits available by a constant number of digits (about half a digit). is is known as simple convergence. If we assume that the recurrence does converge to a constant, we know that the constant must satisfy \\(\alpha = \frac{1}{(1+\alpha)}\\), or \\(1 - \alpha - \alpha^2 = 0\\), which leads to the solution \\(\alpha = \frac{(\sqrt{5}-1)}{2} \approx .6180334\\).
## Higher-Order Recurrences
Recurrences where the right-hand side of the equation for \\(a_n\\) is a linear combination of \\(a_{n-1}, a_{n-2},\\) and so on.
For example:
$$a_n = 3a_{n-1} - 2a_{n-2} \qquad for \ n>1 \ with \ a_0 = 0 \ and \ a_1=1$$
Solution:
$$
\eqalign{
a_n - a_{n-1} &= 2(a_{n-1} - 2_{n-2})\\
&= 2^{n-1}\\
a_n &= 2^{n}-1\\
}
$$
Another example:
$$a_n = 5a_{n-1} - 6a_{n-1} \qquad for \ n>1 \ with \ a_0 = 0 \ and \ a_1=1$$
Has solution:
$$
\eqalign{
a_n - 3a_{n-1} &= 2(a_{n-1} - 3_{n-2})\\
&= 2^{n-1}\\
a_n &= 3^n - 2^n
}
$$
**Theorem 2.2 (Linear recurrences with constant coefficients)**
$$a_n = x_1a_{n-1}+x_2a_{n-2}+...+x_ta_{n-t} \qquad for \ n \geq \ t$$
Can be expressed as a linear combination (with coefficients depending on the initial conditions \\(a_0,a_1,...,a_{t-1}\\)):
$$q(z) \equiv z^t - x_1z^{t-1} - x_2z^{t-2} - ... - x_t$$
**PROOF**. For \\(a_n = \beta^n\\), where \\(\beta\\) is a root of the "characteristic polynomial":
$$\beta^n = x_1\beta^{n-1} + x_2\beta^{n-2} + ... + x_t\beta^{n-t} \qquad for \ n \geq t$$
or, equivalently,
$$\beta^{n-t}q(\beta) = 0$$
That is, \\(\beta^n\\) is a solution to the recurrence for any root \\(\beta\\) of the characteristic polynomial.
**Finding the coefficients**. Develop theorem 2.2 to create a system of simultaneously equations that can be solved to yield the constants in the linear combination. For example:
$$a_n = 5a_{n-1} - 6a_{n-2} \qquad for \ n \geq 2 \ with a_0 = 0 \ and \ a_1 = 1$$
The characteristic equation is \\(z^2 - 5z + 6 = (z-3)(z-2)\\), so:
$$a_n = c_03^n + c_12^n$$
Matching this formula against the values at \\(n = 0\\) and \\(n = 1\\), we have:
$$a_0 = 0 = c_0 + c_1$$
$$a_1 = 1 = 3c_0 + 2c_1$$
The solution to these simultaneous equations is \\(c_0 = 1\\) and \\(c_1 = -1\\), so \\(a_n = 3^n - 2^n\\)
**Degenerate cases**. When the coefficients turn out to be zero and/or some roots have the same modulus, the result can be somewhat counterintuitive. For example:
$$a_n = 2a_{n-1} - a_{n-2} \qquad for \ n \geq 2 \ with \ q_0 = 1 \ and \ a_1 = 2$$
The characteristic equation is \\(z^2 - 2z + 1 = (z-1)^2\\), so:
$$a_n = c_01^n + c_1n1^n$$
We have:
$$a_0 = 1 = c_0$$
$$a_1 = 2= c_0 + c_1$$
gives \\(c_0 = c_1 = 1\\), so \\(a_n = n + 1\\).
**Fibonacci numbers**. \\({0, 1, 1, 2, 3, 4, 8, 13, ...}\\) is defined by prototypical second-order recurrence:
$$F_n = F_{n-1} + F_{n-2} \qquad for \ n>1 \ with \ F_0 = 0 \ and \ F_1 = 1$$
roots of \\(u^2-u-1 = \phi = (1+\sqrt(5)/2 = 1.61803..\\) and \\(\hat{\phi} = (1-\sqrt(5)/2 = -.61803...)\\), Theorem 2.2 says that the solution is:
$$F_N = c_0\phi^N+c_1\hat{\phi}^N$$
for some constants \\(c_0\\) and \\(c1\\):
$$F_0 = 0 = c_0 + c_1$$
$$F_1 = 1 = c_0\phi + c_1\hat{\phi}$$
yield the solution:
$$F_N = \frac{1}{\sqrt{(5)}}(\phi^N - \hat{\phi}^N)$$
**Nonconstant coefficients**. If the coefficients are not constants, then more advanced technique are needed because Theorem 2.2 does not apply. For example:
$$\eqalign{
a_n &= na_{n-1} + n(n-1)a_{n-2} \qquad for \ n>1 \ with \ a_1 = 1 \ and \ a_0 = 0\\
\frac{a_n}{n!} &= \frac{na_{n-1} + n(n-1)a_{n-2}}{n!}\\
a_n &= n!F_n\\
}$$
## Method for Solving Recurrences
**Change of Variables**
For example:
$$a_n = \sqrt{a_{n-1}a_{n-2}} \qquad for \ n>1 \ with \ a_0 = 1 \ and \ a_1 = 2$$
Let \\(b_n = lga_n\\):
$$b_n = \frac{1}{2}(b_{n-1} + b_{n-2}) \qquad for \ n>1 \ with \ b_0 = 0 \ and \ b_1 = 1$$
<br><br>
Another example, for recurrence to continued fractions:
$$a_n = \frac{1}{(1+a_{n-1})} \qquad for \ n>0 \ with \ a_0 = 1$$
Iterating this recurrence gives the sequence:
$$\eqalign{
a_0 &= 1\\
a_1 &= \frac{1}{1+1} = \frac{1}{2}\\
a_2 &= \frac{1}{1 + \frac{1}{1+1}} = \frac{1}{1+\frac{1}{2}} = \frac{2}{3}\\
a_3 &= \frac{1}{1+\frac{1}{1 + \frac{1}{1+1}}} = \frac{1}{1+\frac{2}{3}} = \frac{3}{5}
}$$
That form, reveals the Fibonacci numbers. Let \\(a_n = b_{n-1} / b_n\\):
$$\frac{b_{n-1}}{b_n} = 1 / (1 + \frac{b_{n-2}}{b_{n-1}}) \qquad for \ n > 1 \ with \ b_0 = b_1 = 1$$
Dividng both sides by \\(b_{n-1}\\) gives:
$$\frac{1}{b_n} = \frac{1}{b_{n-1} + b_{n-2}} \qquad for \ n > 1 \ with \ b_0 = b_1 = 1$$
**Repertoire**
$$a_n = (n-1)a_{n-1}-na_{n-2}+n+1 \qquad for \ n>1 \ with \ a_0 = a_1 = 1$$
Introducing a quantity \\(f(n)\\) to the right-hand side:
$$a_n = (n-1)a_{n-1}-na_{n-2}+f(n)$$
We arrive at the table:
| \\(a_n\\) | \\(f(n)\\) |
|-------------|--------------|
| 1 | 2 |
| \\(n\\) | n - 1 |
| \\(n^2\\) | n + 1 |
Subtracting the first row form the third gives the result that means that:
$$a_n = n^2 - 1 \qquad when \qquad f(n) = n-1\\
initial \ condition \ a_0 = -1 \ and \ a_1 = 0$$
Now, we have two (linearly independent) solutions for \\(f(n) = n - 1\\), which we combine to get the right initial values, yielding the result:
$$a_n = n^2 - n + 1$$
**Bootstapping**
Given Fibonacci recurrence:
$$a_n = a_{n-1} + a_{n-2} \qquad for \ n > 1 \ with a_0 = 0 \ and a_1 = 1$$
Note, that \\(a_n\\) is increasing. Therefore, \\(a_{n-1} > a_{n-2}\\) and \\(a_n > 2a_{n-2}\\). Iterating this inequality imply the lower bound:
$$a_n > 2^{n/2}$$
On the other hand, \\(a_{n-2} > a_{n-1}\\) and \\(a_n < 2a_{n-1}\\). Iterating this inequality imply the upper bound:
$$a_n < 2^{n}$$
"guessing" a solution of the form \\(a_n \sim c_0\alpha^n\\), with \\(\sqrt(2) < \alpha < 2\\)
**Perturbation**
$$a_{n+1} = 2a_n + \frac{a_{n-1}}{n^2} \qquad for \ n>1 \ with \ a_0=1 \ and \ a_1 = 2$$
Eliminate \\(\frac{1}{n^2}\\) because makes only a small constribution to recurrence, so:
$$a_{n+1} \approx 2a_n$$
This a growth of the rough form \\(a_n \approx 2^n\\) is anticipated. To make this preciese, consider the simpler sequence:
$$b_{n+1} = 2b_n \qquad for \ n>0 \ with \ b_0=1$$
(so that \\(b_n = 2^n\\)) and compare the two recurrence by forming the ratio:
$$\rho_n = \frac{a_n}{b_n} = \frac{a_n}{2^n}$$
From the recurrences, we have:
$$\rho_{n+1} = \rho_n + \frac{1}{4n^2} \rho_{n-1} \qquad for \ n>0 \ with \ \rho_0=1$$
Clearly, the \\(\rho_n\\) are increasing. To prove they tend to a constant, note that:
$$\rho_{n+1} \leq \rho_n (1+\frac{1}{4n^2}) \qquad for \ n \geq 1 \ so \ that \qquad \rho_{n+1} \leq \Pi_{k=1}^n (1+\frac{1}{4k^2})$$
But the infinite product corresponding to the right-hand side converges monotonically to:
$$\alpha_0 = \Pi_{k=1}^\infty (1 + \frac{1}{4k^2}) = 1.46505...$$
This, \\(\rho_n\\) is bounded from above by \\(\alpha_0\\) and, as it is increasing, it must converge to a constant. We have thus proved that:
$$a_n \sim \alpha \cdot 2^n$$
for some constant \\(\alpha \le 1.46505...\\)
## Binary Divide-and-Conquer Recurrences and Binary Numbers
The number of comparisons used by mergesort is given by the solution to the recurrence:
$$C_N=C_{\lfloor{N/2}\rfloor}+C_{\lceil{N/2}\rceil}+N \qquad for \ N > 1 \ with \ C_1=0 \qquad (4)$$

<br>

**Binary Search**
The number comparisons used during an unsuccessful search with binary search in a table of size N in the worst case is equal to the number of bits in the binary representation of N.
$$B_N = B_{\lfloor{N/2}\rfloor}+1 \qquad for \ N \geq 2 \ with \ B_1 = 1$$
Which exact solution is:
$$B_N = \lfloor{lgN}\rfloor + 1$$
**Exact solution of mergesort recurrence**
The mergesort recurrence (4) is easily solved by differencing: if \\(D_N\\) is defined to be \\(C_{N+1} - C_N\\), the \\(D_N\\) satisfied the recurrence:
$$D_N = D_{\lfloor{N/2}\rfloor}+1 \qquad for \ N \geq 2 \ with \ D_1 = 2$$
which iterates to:
$$D_N = {\lfloor{lgN}\rfloor}+2$$
and, therefore,
$$C_N = N - 1 + \sum_{1 \leq k < N} ({\lfloor{lgk}\rfloor}+1)$$

**Theorem 2.4 (Mergesort)**. The number of comparison used by mergesort is equal to \\(N - 1\\) plus the number of bits in the binary representations of all the numbers less than \\(N\\). Both quantities are described by the recurrence:
$$C_N=C_{\lfloor{N/2}\rfloor}+C_{\lceil{N/2}\rceil}+N \qquad for \ N \geq 2 \ with \ C_1=0$$
Which has the exact solution \\(C_N = N{\lfloor{lgN}\rfloor} + 2N - 2^{{\lfloor{lgN}\rfloor} + 1}\\).
**PROOF**
$$
\eqalign{
C_N &= (N-1) + (N-1) + (N-2) + (N-4) + ... + (N - 2^{{\lfloor{lgN}\rfloor} + 1})\\
&= (N-1) + N({\lfloor{lgN}\rfloor} + 1) - (1 + 2 + 4 + ... + 2^{{\lfloor{lgN}\rfloor} + 1})\\
&= N{\lfloor{lgN}\rfloor} + 2N - 2^{{\lfloor{lgN}\rfloor} + 1}
}
$$
## General Divide-and-Conquer Recurrences
A variety of "devide-and-conquer" recurrence qrise that depend on the number and relative size of subproblems, the extent to which they overlap, and the cost of recombining them for the solution.
In pursuit of general solution, we start with the recursive formula:
$$\alpha(x) = \alpha a (x/\beta) + f(x) \qquad for \ x>1 \ with \ a(x) = 0 \ for \ x \leq 1$$
defining a function over the positive real numbers. In essence, this corresponds to a divide-and-conquer algorithm that divides a problem of size x into \\(\alpha\\) subproblems of size \\(x/\beta\\) and recombines them at a cost of \\(f(x)\\).
For example, consider the case where \\(f(x)=x\\) and we restrict ourselces to the integers \\(N=\beta^n\\). We have:
$$\alpha_{\beta n} = \alpha a_{\beta n - 1} + \beta^n \qquad for \ n > 0 \ with \ a_1 = 0$$
Dividing both sides by \\(\alpha^n\\) and iterating, we have the solution:
$$\alpha_{\beta n} = \alpha^n \sum_{1 \leq j \leq n} \left(\frac{\beta}{\alpha}\right)^j$$
**Theorem 2.5 (Divide-and-conquer functions)**. If the function \\(a(x)\\) satisfies the recurrence:
$$a(x) = \alpha a (x/\beta) + x \qquad for \ x > 1 \ with \ a(x) = 0 \ for \ x \leq 1$$
then
$$
\eqalign{
if \ \alpha < \beta &\qquad a(x) \sim \frac{\beta}{\beta-\alpha}x\\
if \ \alpha = \beta &\qquad a(x) \sim xlog_{\beta}x\\
if \ \alpha > \beta &\qquad a(x) \sim \frac{\alpha}{\alpha-\beta}\left(\frac{\beta}{\alpha}\right)^{{log_{\beta}\alpha}} x^{log_{\beta}\alpha}
}
$$

## Master Theorem
### Divide-and-conquer algorithms
Suppose that an algorithm attacks a problem of size \\(N\\) by:
* Dividing into \\(\alpha\\) parts of size about \\(N/\beta\\)
* Solving recursively
* Combining solutions with extra cost \\(\theta(N^{\gamma}(log N)^{\delta})\\)
1. Mergesort (\\(\alpha = 2, \ \beta = 2, \ \gamma = 1, \ \delta = 0\\))
<br>
$$C_N = 2C_{N/2}+N$$
<br><br>
2. Batcher network (\\(\alpha = 2, \ \beta = 2, \ \gamma = 1, \ \delta = 1\\))
<br>
$$C_N = 2C_{N/2} + NlgN$$
<br><br>
3. Karatsuba multiplication (\\(\alpha = 3, \ \beta = 2, \ \gamma = 1, \ \delta = 0\\))
<br>
$$C_N = 3C_{N/2}+N$$
<br><br>
4. Strasses matrix multiply (\\(\alpha = 7, \ \beta = 2, \ \gamma = 1, \ \delta = 0\\))
<br>
$$C_N = 7C_{N/2}+N$$
<br><br>
| github_jupyter |
<font face="微軟正黑體">
## Build DataFrame
- [Different ways to create Pandas Dataframe](https://www.geeksforgeeks.org/different-ways-to-create-pandas-dataframe/)
```
import pandas as pd
import numpy as np
import json
# Method 1
data = {
'weekday': ['Sun', 'Sun', 'Mon', 'Mon'],
'city': ['Austin', 'Dallas', 'Austin', 'Dallas'],
'visitor': [139, 237, 326, 456]
}
visitors_1 = pd.DataFrame(data)
print(visitors_1)
# Method 2
cities = ['Austin', 'Dallas', 'Austin', 'Dallas']
weekdays = ['Sun', 'Sun', 'Mon', 'Mon']
visitors = [139, 237, 326, 456]
list_labels = ['city', 'weekday', 'visitor']
list_cols = [cities, weekdays, visitors]
zipped = list(zip(list_labels, list_cols))
visitors_2 = pd.DataFrame(dict(zipped))
print(visitors_2)
```
<font face="微軟正黑體">
## 一個簡單例子
假設你想知道如果利用 pandas 計算上述資料中,每個 weekday 的平均 visitor 數量,
通過 google 你找到了 https://stackoverflow.com/questions/30482071/how-to-calculate-mean-values-grouped-on-another-column-in-pandas
想要測試的時候就可以用 visitors_1 這個只有 4 筆資料的資料集來測試程式碼
```
data = {
'weekday': ['Sun', 'Mon', 'Mon', 'Mon'],
'city': ['Austin', 'Dallas', 'Austin', 'Dallas'],
'visitor1': [139, 237, 326, 456],
'visitor2': [233, 444, 315, 789]
}
visitors_3 = pd.DataFrame(data)
groupby_weekday = visitors_3.groupby(by="weekday")
print(type(groupby_weekday))
print("--- groupby weekday - size ---\n{}\n".format(groupby_weekday.size()))
print("--- groupby weekday - groups ---\n{}\n".format(groupby_weekday.groups))
print("--- groupby weekday - mean ---\n{}\n".format(groupby_weekday.mean().astype(int)))
print("--- groupby weekday - sum ---\n{}\n".format(groupby_weekday.sum()))
print("--- groupby weekday - visitor1 mean ---\n{}\n".format(groupby_weekday["visitor1"].mean()))
```
<font face="微軟正黑體">
## 練習時間
在小量的資料上,我們用眼睛就可以看得出來程式碼是否有跑出我們理想中的結果
請嘗試想像一個你需要的資料結構 (裡面的值可以是隨機的),然後用上述的方法把它變成 pandas DataFrame
#### Ex: 想像一個 dataframe 有兩個欄位,一個是國家,一個是人口,求人口數最多的國家
### Hints: [隨機產生數值](https://blog.csdn.net/christianashannon/article/details/78867204)
```
# Build countries DataFrame
countries = ["Taiwan", "United States", "Thailand"]
population = np.random.randint(low=10E6, high=10E7, size=3)
columns_label = ["countries", "population"]
# countries_df = pd.DataFrame(dict(list(zip(columns_label, [countries, population]))))
countries_df = pd.DataFrame(list(zip(countries, population)), columns=columns_label)
# show DataFrame
print(countries_df)
print()
# Get the most population country
print(countries_df.iloc[countries_df["population"].idxmax()])
```
| github_jupyter |
<a href="https://colab.research.google.com/github/mrdbourke/tensorflow-deep-learning/blob/main/07_food_vision_milestone_project_1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Milestone Project 1: 🍔👁 Food Vision Big™
In the previous notebook ([transfer learning part 3: scaling up](https://github.com/mrdbourke/tensorflow-deep-learning/blob/main/06_transfer_learning_in_tensorflow_part_3_scaling_up.ipynb)) we built Food Vision mini: a transfer learning model which beat the original results of the [Food101 paper](https://data.vision.ee.ethz.ch/cvl/datasets_extra/food-101/) with only 10% of the data.
But you might be wondering, what would happen if we used all the data?
Well, that's what we're going to find out in this notebook!
We're going to be building Food Vision Big™, using all of the data from the Food101 dataset.
Yep. All 75,750 training images and 25,250 testing images.
And guess what...
This time **we've got the goal of beating [DeepFood](https://www.researchgate.net/publication/304163308_DeepFood_Deep_Learning-Based_Food_Image_Recognition_for_Computer-Aided_Dietary_Assessment)**, a 2016 paper which used a Convolutional Neural Network trained for 2-3 days to achieve 77.4% top-1 accuracy.
> 🔑 **Note:** **Top-1 accuracy** means "accuracy for the top softmax activation value output by the model" (because softmax outputs a value for every class, but top-1 means only the highest one is evaluated). **Top-5 accuracy** means "accuracy for the top 5 softmax activation values output by the model", in other words, did the true label appear in the top 5 activation values? Top-5 accuracy scores are usually noticeably higher than top-1.
| | 🍔👁 Food Vision Big™ | 🍔👁 Food Vision mini |
|-----|-----|-----|
| Dataset source | TensorFlow Datasets | Preprocessed download from Kaggle |
| Train data | 75,750 images | 7,575 images |
| Test data | 25,250 images | 25,250 images |
| Mixed precision | Yes | No |
| Data loading | Performanant tf.data API | TensorFlow pre-built function |
| Target results | 77.4% top-1 accuracy (beat [DeepFood paper](https://arxiv.org/abs/1606.05675)) | 50.76% top-1 accuracy (beat [Food101 paper](https://data.vision.ee.ethz.ch/cvl/datasets_extra/food-101/static/bossard_eccv14_food-101.pdf)) |
*Table comparing difference between Food Vision Big (this notebook) versus Food Vision mini (previous notebook).*
Alongside attempting to beat the DeepFood paper, we're going to learn about two methods to significantly improve the speed of our model training:
1. Prefetching
2. Mixed precision training
But more on these later.
## What we're going to cover
* Using TensorFlow Datasets to download and explore data
* Creating preprocessing function for our data
* Batching & preparing datasets for modelling (**making our datasets run fast**)
* Creating modelling callbacks
* Setting up **mixed precision training**
* Building a feature extraction model (see [transfer learning part 1: feature extraction](https://github.com/mrdbourke/tensorflow-deep-learning/blob/main/04_transfer_learning_in_tensorflow_part_1_feature_extraction.ipynb))
* Fine-tuning the feature extraction model (see [transfer learning part 2: fine-tuning](https://github.com/mrdbourke/tensorflow-deep-learning/blob/main/05_transfer_learning_in_tensorflow_part_2_fine_tuning.ipynb))
* Viewing training results on TensorBoard
## How you should approach this notebook
You can read through the descriptions and the code (it should all run, except for the cells which error on purpose), but there's a better option.
Write all of the code yourself.
Yes. I'm serious. Create a new notebook, and rewrite each line by yourself. Investigate it, see if you can break it, why does it break?
You don't have to write the text descriptions but writing the code yourself is a great way to get hands-on experience.
Don't worry if you make mistakes, we all do. The way to get better and make less mistakes is to write more code.
> 📖 **Resource:** See the full set of course materials on GitHub: https://github.com/mrdbourke/tensorflow-deep-learning
## Check GPU
For this notebook, we're going to be doing something different.
We're going to be using mixed precision training.
Mixed precision training was introduced in [TensorFlow 2.4.0](https://blog.tensorflow.org/2020/12/whats-new-in-tensorflow-24.html) (a very new feature at the time of writing).
What does **mixed precision training** do?
Mixed precision training uses a combination of single precision (float32) and half-precision (float16) data types to speed up model training (up 3x on modern GPUs).
We'll talk about this more later on but in the meantime you can read the [TensorFlow documentation on mixed precision](https://www.tensorflow.org/guide/mixed_precision) for more details.
For now, before we can move forward if we want to use mixed precision training, we need to make sure the GPU powering our Google Colab instance (if you're using Google Colab) is compatible.
For mixed precision training to work, you need access to a GPU with a compute compability score of 7.0+.
Google Colab offers P100, K80 and T4 GPUs, however, **the P100 and K80 aren't compatible with mixed precision training**.
Therefore before we proceed we need to make sure we have **access to a Tesla T4 GPU in our Google Colab instance**.
If you're not using Google Colab, you can find a list of various [Nvidia GPU compute capabilities on Nvidia's developer website](https://developer.nvidia.com/cuda-gpus#compute).
> 🔑 **Note:** If you run the cell below and see a P100 or K80, try going to Runtime -> Factory Reset Runtime (note: this will remove any saved variables and data from your Colab instance) and then retry to get a T4.
```
# If using Google Colab, this should output "Tesla T4" otherwise,
# you won't be able to use mixed precision training
!nvidia-smi -L
```
Since mixed precision training was introduced in TensorFlow 2.4.0, make sure you've got at least TensorFlow 2.4.0+.
```
# Check TensorFlow version (should be 2.4.0+)
import tensorflow as tf
print(tf.__version__)
```
## Get helper functions
We've created a series of helper functions throughout the previous notebooks in the course. Instead of rewriting them (tedious), we'll import the [`helper_functions.py`](https://github.com/mrdbourke/tensorflow-deep-learning/blob/main/extras/helper_functions.py) file from the GitHub repo.
```
# Get helper functions file
!wget https://raw.githubusercontent.com/mrdbourke/tensorflow-deep-learning/main/extras/helper_functions.py
# Import series of helper functions for the notebook (we've created/used these in previous notebooks)
from helper_functions import create_tensorboard_callback, plot_loss_curves, compare_historys
```
## Use TensorFlow Datasets to Download Data
In previous notebooks, we've downloaded our food images (from the [Food101 dataset](https://www.kaggle.com/dansbecker/food-101/home)) from Google Storage.
And this is a typical workflow you'd use if you're working on your own datasets.
However, there's another way to get datasets ready to use with TensorFlow.
For many of the most popular datasets in the machine learning world (often referred to and used as benchmarks), you can access them through [TensorFlow Datasets (TFDS)](https://www.tensorflow.org/datasets/overview).
What is **TensorFlow Datasets**?
A place for prepared and ready-to-use machine learning datasets.
Why use TensorFlow Datasets?
* Load data already in Tensors
* Practice on well established datasets
* Experiment with different data loading techniques (like we're going to use in this notebook)
* Experiment with new TensorFlow features quickly (such as mixed precision training)
Why *not* use TensorFlow Datasets?
* The datasets are static (they don't change, like your real-world datasets would)
* Might not be suited for your particular problem (but great for experimenting)
To begin using TensorFlow Datasets we can import it under the alias `tfds`.
```
# Get TensorFlow Datasets
import tensorflow_datasets as tfds
```
To find all of the available datasets in TensorFlow Datasets, you can use the `list_builders()` method.
After doing so, we can check to see if the one we're after (`"food101"`) is present.
```
# List available datasets
datasets_list = tfds.list_builders() # get all available datasets in TFDS
print("food101" in datasets_list) # is the dataset we're after available?
```
Beautiful! It looks like the dataset we're after is available (note there are plenty more available but we're on Food101).
To get access to the Food101 dataset from the TFDS, we can use the [`tfds.load()`](https://www.tensorflow.org/datasets/api_docs/python/tfds/load) method.
In particular, we'll have to pass it a few parameters to let it know what we're after:
* `name` (str) : the target dataset (e.g. `"food101"`)
* `split` (list, optional) : what splits of the dataset we're after (e.g. `["train", "validation"]`)
* the `split` parameter is quite tricky. See [the documentation for more](https://github.com/tensorflow/datasets/blob/master/docs/splits.md).
* `shuffle_files` (bool) : whether or not to shuffle the files on download, defaults to `False`
* `as_supervised` (bool) : `True` to download data samples in tuple format (`(data, label)`) or `False` for dictionary format
* `with_info` (bool) : `True` to download dataset metadata (labels, number of samples, etc)
> 🔑 **Note:** Calling the `tfds.load()` method will start to download a target dataset to disk if the `download=True` parameter is set (default). This dataset could be 100GB+, so make sure you have space.
```
# Load in the data (takes about 5-6 minutes in Google Colab)
(train_data, test_data), ds_info = tfds.load(name="food101", # target dataset to get from TFDS
split=["train", "validation"], # what splits of data should we get? note: not all datasets have train, valid, test
shuffle_files=True, # shuffle files on download?
as_supervised=True, # download data in tuple format (sample, label), e.g. (image, label)
with_info=True) # include dataset metadata? if so, tfds.load() returns tuple (data, ds_info)
```
Wonderful! After a few minutes of downloading, we've now got access to entire Food101 dataset (in tensor format) ready for modelling.
Now let's get a little information from our dataset, starting with the class names.
Getting class names from a TensorFlow Datasets dataset requires downloading the "`dataset_info`" variable (by using the `as_supervised=True` parameter in the `tfds.load()` method, **note:** this will only work for supervised datasets in TFDS).
We can access the class names of a particular dataset using the `dataset_info.features` attribute and accessing `names` attribute of the `"label"` key.
```
# Features of Food101 TFDS
ds_info.features
# Get class names
class_names = ds_info.features["label"].names
class_names[:10]
```
### Exploring the Food101 data from TensorFlow Datasets
Now we've downloaded the Food101 dataset from TensorFlow Datasets, how about we do what any good data explorer should?
In other words, "visualize, visualize, visualize".
Let's find out a few details about our dataset:
* The shape of our input data (image tensors)
* The datatype of our input data
* What the labels of our input data look like (e.g. one-hot encoded versus label-encoded)
* Do the labels match up with the class names?
To do, let's take one sample off the training data (using the [`.take()` method](https://www.tensorflow.org/api_docs/python/tf/data/Dataset#take)) and explore it.
```
# Take one sample off the training data
train_one_sample = train_data.take(1) # samples are in format (image_tensor, label)
```
Because we used the `as_supervised=True` parameter in our `tfds.load()` method above, data samples come in the tuple format structure `(data, label)` or in our case `(image_tensor, label)`.
```
# What does one sample of our training data look like?
train_one_sample
```
Let's loop through our single training sample and get some info from the `image_tensor` and `label`.
```
# Output info about our training sample
for image, label in train_one_sample:
print(f"""
Image shape: {image.shape}
Image dtype: {image.dtype}
Target class from Food101 (tensor form): {label}
Class name (str form): {class_names[label.numpy()]}
""")
```
Because we set the `shuffle_files=True` parameter in our `tfds.load()` method above, running the cell above a few times will give a different result each time.
Checking these you might notice some of the images have different shapes, for example `(512, 342, 3)` and `(512, 512, 3)` (height, width, color_channels).
Let's see what one of the image tensors from TFDS's Food101 dataset looks like.
```
# What does an image tensor from TFDS's Food101 look like?
image
# What are the min and max values?
tf.reduce_min(image), tf.reduce_max(image)
```
Alright looks like our image tensors have values of between 0 & 255 (standard red, green, blue colour values) and the values are of data type `unit8`.
We might have to preprocess these before passing them to a neural network. But we'll handle this later.
In the meantime, let's see if we can plot an image sample.
### Plot an image from TensorFlow Datasets
We've seen our image tensors in tensor format, now let's really adhere to our motto.
"Visualize, visualize, visualize!"
Let's plot one of the image samples using [`matplotlib.pyplot.imshow()`](https://matplotlib.org/stable/api/_as_gen/matplotlib.pyplot.imshow.html) and set the title to target class name.
```
# Plot an image tensor
import matplotlib.pyplot as plt
plt.imshow(image)
plt.title(class_names[label.numpy()]) # add title to image by indexing on class_names list
plt.axis(False);
```
Delicious!
Okay, looks like the Food101 data we've got from TFDS is similar to the datasets we've been using in previous notebooks.
Now let's preprocess it and get it ready for use with a neural network.
## Create preprocessing functions for our data
In previous notebooks, when our images were in folder format we used the method [`tf.keras.preprocessing.image_dataset_from_directory()`](https://www.tensorflow.org/api_docs/python/tf/keras/preprocessing/image_dataset_from_directory) to load them in.
Doing this meant our data was loaded into a format ready to be used with our models.
However, since we've downloaded the data from TensorFlow Datasets, there are a couple of preprocessing steps we have to take before it's ready to model.
More specifically, our data is currently:
* In `uint8` data type
* Comprised of all differnet sized tensors (different sized images)
* Not scaled (the pixel values are between 0 & 255)
Whereas, models like data to be:
* In `float32` data type
* Have all of the same size tensors (batches require all tensors have the same shape, e.g. `(224, 224, 3)`)
* Scaled (values between 0 & 1), also called normalized
To take care of these, we'll create a `preprocess_img()` function which:
* Resizes an input image tensor to a specified size using [`tf.image.resize()`](https://www.tensorflow.org/api_docs/python/tf/image/resize)
* Converts an input image tensor's current datatype to `tf.float32` using [`tf.cast()`](https://www.tensorflow.org/api_docs/python/tf/cast)
> 🔑 **Note:** Pretrained EfficientNetBX models in [`tf.keras.applications.efficientnet`](https://www.tensorflow.org/api_docs/python/tf/keras/applications/efficientnet) (what we're going to be using) have rescaling built-in. But for many other model architectures you'll want to rescale your data (e.g. get its values between 0 & 1). This could be incorporated inside your "`preprocess_img()`" function (like the one below) or within your model as a [`tf.keras.layers.experimental.preprocessing.Rescaling`](https://www.tensorflow.org/api_docs/python/tf/keras/layers/experimental/preprocessing/Rescaling) layer.
```
# Make a function for preprocessing images
def preprocess_img(image, label, img_shape=224):
"""
Converts image datatype from 'uint8' -> 'float32' and reshapes image to
[img_shape, img_shape, color_channels]
"""
image = tf.image.resize(image, [img_shape, img_shape]) # reshape to img_shape
return tf.cast(image, tf.float32), label # return (float32_image, label) tuple
```
Our `preprocess_img()` function above takes image and label as input (even though it does nothing to the label) because our dataset is currently in the tuple structure `(image, label)`.
Let's try our function out on a target image.
```
# Preprocess a single sample image and check the outputs
preprocessed_img = preprocess_img(image, label)[0]
print(f"Image before preprocessing:\n {image[:2]}...,\nShape: {image.shape},\nDatatype: {image.dtype}\n")
print(f"Image after preprocessing:\n {preprocessed_img[:2]}...,\nShape: {preprocessed_img.shape},\nDatatype: {preprocessed_img.dtype}")
```
Excellent! Looks like our `preprocess_img()` function is working as expected.
The input image gets converted from `uint8` to `float32` and gets reshaped from its current shape to `(224, 224, 3)`.
How does it look?
```
# We can still plot our preprocessed image as long as we
# divide by 255 (for matplotlib capatibility)
plt.imshow(preprocessed_img/255.)
plt.title(class_names[label])
plt.axis(False);
```
All this food visualization is making me hungry. How about we start preparing to model it?
## Batch & prepare datasets
Before we can model our data, we have to turn it into batches.
Why?
Because computing on batches is memory efficient.
We turn our data from 101,000 image tensors and labels (train and test combined) into batches of 32 image and label pairs, thus enabling it to fit into the memory of our GPU.
To do this in effective way, we're going to be leveraging a number of methods from the [`tf.data` API](https://www.tensorflow.org/api_docs/python/tf/data).
> 📖 **Resource:** For loading data in the most performant way possible, see the TensorFlow documentation on [Better performance with the tf.data API](https://www.tensorflow.org/guide/data_performance).
Specifically, we're going to be using:
* [`map()`](https://www.tensorflow.org/api_docs/python/tf/data/Dataset#map) - maps a predefined function to a target dataset (e.g. `preprocess_img()` to our image tensors)
* [`shuffle()`](https://www.tensorflow.org/api_docs/python/tf/data/Dataset#shuffle) - randomly shuffles the elements of a target dataset up `buffer_size` (ideally, the `buffer_size` is equal to the size of the dataset, however, this may have implications on memory)
* [`batch()`](https://www.tensorflow.org/api_docs/python/tf/data/Dataset#batch) - turns elements of a target dataset into batches (size defined by parameter `batch_size`)
* [`prefetch()`](https://www.tensorflow.org/api_docs/python/tf/data/Dataset#prefetch) - prepares subsequent batches of data whilst other batches of data are being computed on (improves data loading speed but costs memory)
* Extra: [`cache()`](https://www.tensorflow.org/api_docs/python/tf/data/Dataset#cache) - caches (saves them for later) elements in a target dataset, saving loading time (will only work if your dataset is small enough to fit in memory, standard Colab instances only have 12GB of memory)
Things to note:
- Can't batch tensors of different shapes (e.g. different image sizes, need to reshape images first, hence our `preprocess_img()` function)
- `shuffle()` keeps a buffer of the number you pass it images shuffled, ideally this number would be all of the samples in your training set, however, if your training set is large, this buffer might not fit in memory (a fairly large number like 1000 or 10000 is usually suffice for shuffling)
- For methods with the `num_parallel_calls` parameter available (such as `map()`), setting it to`num_parallel_calls=tf.data.AUTOTUNE` will parallelize preprocessing and significantly improve speed
- Can't use `cache()` unless your dataset can fit in memory
Woah, the above is alot. But once we've coded below, it'll start to make sense.
We're going to through things in the following order:
```
Original dataset (e.g. train_data) -> map() -> shuffle() -> batch() -> prefetch() -> PrefetchDataset
```
This is like saying,
> "Hey, map this preprocessing function across our training dataset, then shuffle a number of elements before batching them together and make sure you prepare new batches (prefetch) whilst the model is looking through the current batch".
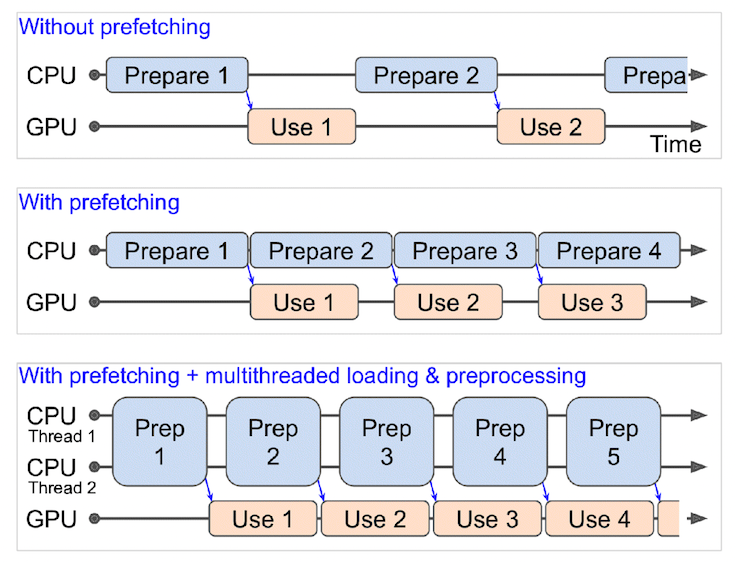
*What happens when you use prefetching (faster) versus what happens when you don't use prefetching (slower). **Source:** Page 422 of [Hands-On Machine Learning with Scikit-Learn, Keras & TensorFlow Book by Aurélien Géron](https://www.oreilly.com/library/view/hands-on-machine-learning/9781492032632/).*
```
# Map preprocessing function to training data (and paralellize)
train_data = train_data.map(map_func=preprocess_img, num_parallel_calls=tf.data.AUTOTUNE)
# Shuffle train_data and turn it into batches and prefetch it (load it faster)
train_data = train_data.shuffle(buffer_size=1000).batch(batch_size=32).prefetch(buffer_size=tf.data.AUTOTUNE)
# Map prepreprocessing function to test data
test_data = test_data.map(preprocess_img, num_parallel_calls=tf.data.AUTOTUNE)
# Turn test data into batches (don't need to shuffle)
test_data = test_data.batch(32).prefetch(tf.data.AUTOTUNE)
```
And now let's check out what our prepared datasets look like.
```
train_data, test_data
```
Excellent! Looks like our data is now in tuples of `(image, label)` with datatypes of `(tf.float32, tf.int64)`, just what our model is after.
> 🔑 **Note:** You can get away without calling the `prefetch()` method on the end of your datasets, however, you'd probably see significantly slower data loading speeds when building a model. So most of your dataset input pipelines should end with a call to [`prefecth()`](https://www.tensorflow.org/api_docs/python/tf/data/Dataset#prefetch).
Onward.
## Create modelling callbacks
Since we're going to be training on a large amount of data and training could take a long time, it's a good idea to set up some modelling callbacks so we be sure of things like our model's training logs being tracked and our model being checkpointed (saved) after various training milestones.
To do each of these we'll use the following callbacks:
* [`tf.keras.callbacks.TensorBoard()`](https://www.tensorflow.org/api_docs/python/tf/keras/callbacks/TensorBoard) - allows us to keep track of our model's training history so we can inspect it later (**note:** we've created this callback before have imported it from `helper_functions.py` as `create_tensorboard_callback()`)
* [`tf.keras.callbacks.ModelCheckpoint()`](https://www.tensorflow.org/api_docs/python/tf/keras/callbacks/ModelCheckpoint) - saves our model's progress at various intervals so we can load it and reuse it later without having to retrain it
* Checkpointing is also helpful so we can start fine-tuning our model at a particular epoch and revert back to a previous state if fine-tuning offers no benefits
```
# Create TensorBoard callback (already have "create_tensorboard_callback()" from a previous notebook)
from helper_functions import create_tensorboard_callback
# Create ModelCheckpoint callback to save model's progress
checkpoint_path = "model_checkpoints/cp.ckpt" # saving weights requires ".ckpt" extension
model_checkpoint = tf.keras.callbacks.ModelCheckpoint(checkpoint_path,
montior="val_acc", # save the model weights with best validation accuracy
save_best_only=True, # only save the best weights
save_weights_only=True, # only save model weights (not whole model)
verbose=0) # don't print out whether or not model is being saved
```
## Setup mixed precision training
We touched on mixed precision training above.
However, we didn't quite explain it.
Normally, tensors in TensorFlow default to the float32 datatype (unless otherwise specified).
In computer science, float32 is also known as [single-precision floating-point format](https://en.wikipedia.org/wiki/Single-precision_floating-point_format). The 32 means it usually occupies 32 bits in computer memory.
Your GPU has a limited memory, therefore it can only handle a number of float32 tensors at the same time.
This is where mixed precision training comes in.
Mixed precision training involves using a mix of float16 and float32 tensors to make better use of your GPU's memory.
Can you guess what float16 means?
Well, if you thought since float32 meant single-precision floating-point, you might've guessed float16 means [half-precision floating-point format](https://en.wikipedia.org/wiki/Half-precision_floating-point_format). And if you did, you're right! And if not, no trouble, now you know.
For tensors in float16 format, each element occupies 16 bits in computer memory.
So, where does this leave us?
As mentioned before, when using mixed precision training, your model will make use of float32 and float16 data types to use less memory where possible and in turn run faster (using less memory per tensor means more tensors can be computed on simultaneously).
As a result, using mixed precision training can improve your performance on modern GPUs (those with a compute capability score of 7.0+) by up to 3x.
For a more detailed explanation, I encourage you to read through the [TensorFlow mixed precision guide](https://www.tensorflow.org/guide/mixed_precision) (I'd highly recommend at least checking out the summary).

*Because mixed precision training uses a combination of float32 and float16 data types, you may see up to a 3x speedup on modern GPUs.*
> 🔑 **Note:** If your GPU doesn't have a score of over 7.0+ (e.g. P100 in Colab), mixed precision won't work (see: ["Supported Hardware"](https://www.tensorflow.org/guide/mixed_precision#supported_hardware) in the mixed precision guide for more).
> 📖 **Resource:** If you'd like to learn more about precision in computer science (the detail to which a numerical quantity is expressed by a computer), see the [Wikipedia page](https://en.wikipedia.org/wiki/Precision_(computer_science)) (and accompanying resources).
Okay, enough talk, let's see how we can turn on mixed precision training in TensorFlow.
The beautiful thing is, the [`tensorflow.keras.mixed_precision`](https://www.tensorflow.org/api_docs/python/tf/keras/mixed_precision/) API has made it very easy for us to get started.
First, we'll import the API and then use the [`set_global_policy()`](https://www.tensorflow.org/api_docs/python/tf/keras/mixed_precision/set_global_policy) method to set the *dtype policy* to `"mixed_float16"`.
```
# Turn on mixed precision training
from tensorflow.keras import mixed_precision
mixed_precision.set_global_policy(policy="mixed_float16") # set global policy to mixed precision
```
Nice! As long as the GPU you're using has a compute capability of 7.0+ the cell above should run without error.
Now we can check the global dtype policy (the policy which will be used by layers in our model) using the [`mixed_precision.global_policy()`](https://www.tensorflow.org/api_docs/python/tf/keras/mixed_precision/global_policy) method.
```
mixed_precision.global_policy() # should output "mixed_float16"
```
Great, since the global dtype policy is now `"mixed_float16"` our model will automatically take advantage of float16 variables where possible and in turn speed up training.
## Build feature extraction model
Callbacks: ready to roll.
Mixed precision: turned on.
Let's build a model.
Because our dataset is quite large, we're going to move towards fine-tuning an existing pretrained model (EfficienetNetB0).
But before we get into fine-tuning, let's set up a feature-extraction model.
Recall, the typical order for using transfer learning is:
1. Build a feature extraction model (replace the top few layers of a pretrained model)
2. Train for a few epochs with lower layers frozen
3. Fine-tune if necessary with multiple layers unfrozen
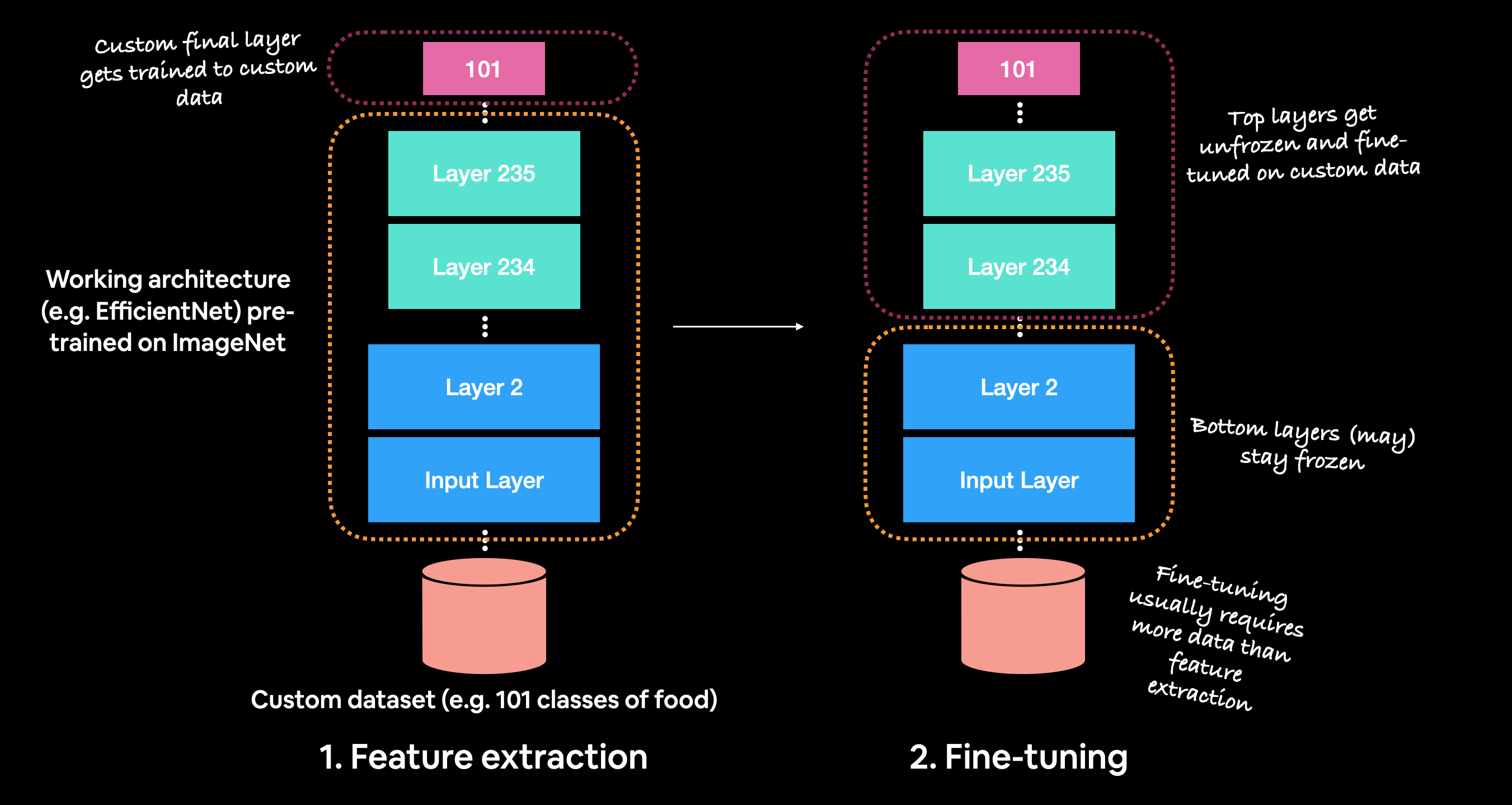
*Before fine-tuning, it's best practice to train a feature extraction model with custom top layers.*
To build the feature extraction model (covered in [Transfer Learning in TensorFlow Part 1: Feature extraction](https://github.com/mrdbourke/tensorflow-deep-learning/blob/main/04_transfer_learning_in_tensorflow_part_1_feature_extraction.ipynb)), we'll:
* Use `EfficientNetB0` from [`tf.keras.applications`](https://www.tensorflow.org/api_docs/python/tf/keras/applications) pre-trained on ImageNet as our base model
* We'll download this without the top layers using `include_top=False` parameter so we can create our own output layers
* Freeze the base model layers so we can use the pre-learned patterns the base model has found on ImageNet
* Put together the input, base model, pooling and output layers in a [Functional model](https://keras.io/guides/functional_api/)
* Compile the Functional model using the Adam optimizer and [sparse categorical crossentropy](https://www.tensorflow.org/api_docs/python/tf/keras/losses/SparseCategoricalCrossentropy) as the loss function (since our labels **aren't** one-hot encoded)
* Fit the model for 3 epochs using the TensorBoard and ModelCheckpoint callbacks
> 🔑 **Note:** Since we're using mixed precision training, our model needs a separate output layer with a hard-coded `dtype=float32`, for example, `layers.Activation("softmax", dtype=tf.float32)`. This ensures the outputs of our model are returned back to the float32 data type which is more numerically stable than the float16 datatype (important for loss calculations). See the ["Building the model"](https://www.tensorflow.org/guide/mixed_precision#building_the_model) section in the TensorFlow mixed precision guide for more.

*Turning mixed precision on in TensorFlow with 3 lines of code.*
```
from tensorflow.keras import layers
from tensorflow.keras.layers.experimental import preprocessing
# Create base model
input_shape = (224, 224, 3)
base_model = tf.keras.applications.EfficientNetB0(include_top=False)
base_model.trainable = False # freeze base model layers
# Create Functional model
inputs = layers.Input(shape=input_shape, name="input_layer")
# Note: EfficientNetBX models have rescaling built-in but if your model didn't you could have a layer like below
# x = preprocessing.Rescaling(1./255)(x)
x = base_model(inputs, training=False) # set base_model to inference mode only
x = layers.GlobalAveragePooling2D(name="pooling_layer")(x)
x = layers.Dense(len(class_names))(x) # want one output neuron per class
# Separate activation of output layer so we can output float32 activations
outputs = layers.Activation("softmax", dtype=tf.float32, name="softmax_float32")(x)
model = tf.keras.Model(inputs, outputs)
# Compile the model
model.compile(loss="sparse_categorical_crossentropy", # Use sparse_categorical_crossentropy when labels are *not* one-hot
optimizer=tf.keras.optimizers.Adam(),
metrics=["accuracy"])
# Check out our model
model.summary()
```
## Checking layer dtype policies (are we using mixed precision?)
Model ready to go!
Before we said the mixed precision API will automatically change our layers' dtype policy's to whatever the global dtype policy is (in our case it's `"mixed_float16"`).
We can check this by iterating through our model's layers and printing layer attributes such as `dtype` and `dtype_policy`.
```
# Check the dtype_policy attributes of layers in our model
for layer in model.layers:
print(layer.name, layer.trainable, layer.dtype, layer.dtype_policy) # Check the dtype policy of layers
```
Going through the above we see:
* `layer.name` (str) : a layer's human-readable name, can be defined by the `name` parameter on construction
* `layer.trainable` (bool) : whether or not a layer is trainable (all of our layers are trainable except the efficientnetb0 layer since we set it's `trainable` attribute to `False`
* `layer.dtype` : the data type a layer stores its variables in
* `layer.dtype_policy` : the data type a layer computes in
> 🔑 **Note:** A layer can have a dtype of `float32` and a dtype policy of `"mixed_float16"` because it stores its variables (weights & biases) in `float32` (more numerically stable), however it computes in `float16` (faster).
We can also check the same details for our model's base model.
```
# Check the layers in the base model and see what dtype policy they're using
for layer in model.layers[1].layers[:20]: # only check the first 20 layers to save output space
print(layer.name, layer.trainable, layer.dtype, layer.dtype_policy)
```
> 🔑 **Note:** The mixed precision API automatically causes layers which can benefit from using the `"mixed_float16"` dtype policy to use it. It also prevents layers which shouldn't use it from using it (e.g. the normalization layer at the start of the base model).
## Fit the feature extraction model
Now that's one good looking model. Let's fit it to our data shall we?
Three epochs should be enough for our top layers to adjust their weights enough to our food image data.
To save time per epoch, we'll also only validate on 15% of the test data.
```
# Fit the model with callbacks
history_101_food_classes_feature_extract = model.fit(train_data,
epochs=3,
steps_per_epoch=len(train_data),
validation_data=test_data,
validation_steps=int(0.15 * len(test_data)),
callbacks=[create_tensorboard_callback("training_logs",
"efficientnetb0_101_classes_all_data_feature_extract"),
model_checkpoint])
```
Nice, looks like our feature extraction model is performing pretty well. How about we evaluate it on the whole test dataset?
```
# Evaluate model (unsaved version) on whole test dataset
results_feature_extract_model = model.evaluate(test_data)
results_feature_extract_model
```
And since we used the `ModelCheckpoint` callback, we've got a saved version of our model in the `model_checkpoints` directory.
Let's load it in and make sure it performs just as well.
## Load and evaluate checkpoint weights
We can load in and evaluate our model's checkpoints by:
1. Cloning our model using [`tf.keras.models.clone_model()`](https://www.tensorflow.org/api_docs/python/tf/keras/models/clone_model) to make a copy of our feature extraction model with reset weights.
2. Calling the `load_weights()` method on our cloned model passing it the path to where our checkpointed weights are stored.
3. Calling `evaluate()` on the cloned model with loaded weights.
A reminder, checkpoints are helpful for when you perform an experiment such as fine-tuning your model. In the case you fine-tune your feature extraction model and find it doesn't offer any improvements, you can always revert back to the checkpointed version of your model.
```
# Clone the model we created (this resets all weights)
cloned_model = tf.keras.models.clone_model(model)
cloned_model.summary()
# Where are our checkpoints stored?
checkpoint_path
# Load checkpointed weights into cloned_model
cloned_model.load_weights(checkpoint_path)
```
Each time you make a change to your model (including loading weights), you have to recompile.
```
# Compile cloned_model (with same parameters as original model)
cloned_model.compile(loss="sparse_categorical_crossentropy",
optimizer=tf.keras.optimizers.Adam(),
metrics=["accuracy"])
# Evalaute cloned model with loaded weights (should be same score as trained model)
results_cloned_model_with_loaded_weights = cloned_model.evaluate(test_data)
```
Our cloned model with loaded weight's results should be very close to the feature extraction model's results (if the cell below errors, something went wrong).
```
# Loaded checkpoint weights should return very similar results to checkpoint weights prior to saving
import numpy as np
assert np.isclose(results_feature_extract_model, results_cloned_model_with_loaded_weights).all() # check if all elements in array are close
```
Cloning the model preserves `dtype_policy`'s of layers (but doesn't preserve weights) so if we wanted to continue fine-tuning with the cloned model, we could and it would still use the mixed precision dtype policy.
```
# Check the layers in the base model and see what dtype policy they're using
for layer in cloned_model.layers[1].layers[:20]: # check only the first 20 layers to save space
print(layer.name, layer.trainable, layer.dtype, layer.dtype_policy)
```
## Save the whole model to file
We can also save the whole model using the [`save()`](https://www.tensorflow.org/api_docs/python/tf/keras/Model#save) method.
Since our model is quite large, you might want to save it to Google Drive (if you're using Google Colab) so you can load it in for use later.
> 🔑 **Note:** Saving to Google Drive requires mounting Google Drive (go to Files -> Mount Drive).
```
# ## Saving model to Google Drive (optional)
# # Create save path to drive
# save_dir = "drive/MyDrive/tensorflow_course/food_vision/07_efficientnetb0_feature_extract_model_mixed_precision/"
# # os.makedirs(save_dir) # Make directory if it doesn't exist
# # Save model
# model.save(save_dir)
```
We can also save it directly to our Google Colab instance.
> 🔑 **Note:** Google Colab storage is ephemeral and your model will delete itself (along with any other saved files) when the Colab session expires.
```
# Save model locally (if you're using Google Colab, your saved model will Colab instance terminates)
save_dir = "07_efficientnetb0_feature_extract_model_mixed_precision"
model.save(save_dir)
```
And again, we can check whether or not our model saved correctly by loading it in and evaluating it.
```
# Load model previously saved above
loaded_saved_model = tf.keras.models.load_model(save_dir)
```
Loading a `SavedModel` also retains all of the underlying layers `dtype_policy` (we want them to be `"mixed_float16"`).
```
# Check the layers in the base model and see what dtype policy they're using
for layer in loaded_saved_model.layers[1].layers[:20]: # check only the first 20 layers to save output space
print(layer.name, layer.trainable, layer.dtype, layer.dtype_policy)
# Check loaded model performance (this should be the same as results_feature_extract_model)
results_loaded_saved_model = loaded_saved_model.evaluate(test_data)
results_loaded_saved_model
# The loaded model's results should equal (or at least be very close) to the model's results prior to saving
# Note: this will only work if you've instatiated results variables
import numpy as np
assert np.isclose(results_feature_extract_model, results_loaded_saved_model).all()
```
That's what we want! Our loaded model performing as it should.
> 🔑 **Note:** We spent a fair bit of time making sure our model saved correctly because training on a lot of data can be time-consuming, so we want to make sure we don't have to continaully train from scratch.
## Preparing our model's layers for fine-tuning
Our feature-extraction model is showing some great promise after three epochs. But since we've got so much data, it's probably worthwhile that we see what results we can get with fine-tuning (fine-tuning usually works best when you've got quite a large amount of data).
Remember our goal of beating the [DeepFood paper](https://arxiv.org/pdf/1606.05675.pdf)?
They were able to achieve 77.4% top-1 accuracy on Food101 over 2-3 days of training.
Do you think fine-tuning will get us there?
Let's find out.
To start, let's load in our saved model.
> 🔑 **Note:** It's worth remembering a traditional workflow for fine-tuning is to freeze a pre-trained base model and then train only the output layers for a few iterations so their weights can be updated inline with your custom data (feature extraction). And then unfreeze a number or all of the layers in the base model and continue training until the model stops improving.
Like all good cooking shows, I've saved a model I prepared earlier (the feature extraction model from above) to Google Storage.
We can download it to make sure we're using the same model going forward.
```
# Download the saved model from Google Storage
!wget https://storage.googleapis.com/ztm_tf_course/food_vision/07_efficientnetb0_feature_extract_model_mixed_precision.zip
# Unzip the SavedModel downloaded from Google Stroage
!mkdir downloaded_gs_model # create new dir to store downloaded feature extraction model
!unzip 07_efficientnetb0_feature_extract_model_mixed_precision.zip -d downloaded_gs_model
# Load and evaluate downloaded GS model
loaded_gs_model = tf.keras.models.load_model("/content/downloaded_gs_model/07_efficientnetb0_feature_extract_model_mixed_precision")
# Get a summary of our downloaded model
loaded_gs_model.summary()
```
And now let's make sure our loaded model is performing as expected.
```
# How does the loaded model perform?
results_loaded_gs_model = loaded_gs_model.evaluate(test_data)
results_loaded_gs_model
```
Great, our loaded model is performing as expected.
When we first created our model, we froze all of the layers in the base model by setting `base_model.trainable=False` but since we've loaded in our model from file, let's check whether or not the layers are trainable or not.
```
# Are any of the layers in our model frozen?
for layer in loaded_gs_model.layers:
layer.trainable = True # set all layers to trainable
print(layer.name, layer.trainable, layer.dtype, layer.dtype_policy) # make sure loaded model is using mixed precision dtype_policy ("mixed_float16")
```
Alright, it seems like each layer in our loaded model is trainable. But what if we got a little deeper and inspected each of the layers in our base model?
> 🤔 **Question:** *Which layer in the loaded model is our base model?*
Before saving the Functional model to file, we created it with five layers (layers below are 0-indexed):
0. The input layer
1. The pre-trained base model layer (`tf.keras.applications.EfficientNetB0`)
2. The pooling layer
3. The fully-connected (dense) layer
4. The output softmax activation (with float32 dtype)
Therefore to inspect our base model layer, we can access the `layers` attribute of the layer at index 1 in our model.
```
# Check the layers in the base model and see what dtype policy they're using
for layer in loaded_gs_model.layers[1].layers[:20]:
print(layer.name, layer.trainable, layer.dtype, layer.dtype_policy)
```
Wonderful, it looks like each layer in our base model is trainable (unfrozen) and every layer which should be using the dtype policy `"mixed_policy16"` is using it.
Since we've got so much data (750 images x 101 training classes = 75750 training images), let's keep all of our base model's layers unfrozen.
> 🔑 **Note:** If you've got a small amount of data (less than 100 images per class), you may want to only unfreeze and fine-tune a small number of layers in the base model at a time. Otherwise, you risk overfitting.
## A couple more callbacks
We're about to start fine-tuning a deep learning model with over 200 layers using over 100,000 (75k+ training, 25K+ testing) images, which means our model's training time is probably going to be much longer than before.
> 🤔 **Question:** *How long does training take?*
It could be a couple of hours or in the case of the [DeepFood paper](https://arxiv.org/pdf/1606.05675.pdf) (the baseline we're trying to beat), their best performing model took 2-3 days of training time.
You will really only know how long it'll take once you start training.
> 🤔 **Question:** *When do you stop training?*
Ideally, when your model stops improving. But again, due to the nature of deep learning, it can be hard to know when exactly a model will stop improving.
Luckily, there's a solution: the [`EarlyStopping` callback](https://www.tensorflow.org/api_docs/python/tf/keras/callbacks/EarlyStopping).
The `EarlyStopping` callback monitors a specified model performance metric (e.g. `val_loss`) and when it stops improving for a specified number of epochs, automatically stops training.
Using the `EarlyStopping` callback combined with the `ModelCheckpoint` callback saving the best performing model automatically, we could keep our model training for an unlimited number of epochs until it stops improving.
Let's set both of these up to monitor our model's `val_loss`.
```
# Setup EarlyStopping callback to stop training if model's val_loss doesn't improve for 3 epochs
early_stopping = tf.keras.callbacks.EarlyStopping(monitor="val_loss", # watch the val loss metric
patience=3) # if val loss decreases for 3 epochs in a row, stop training
# Create ModelCheckpoint callback to save best model during fine-tuning
checkpoint_path = "fine_tune_checkpoints/"
model_checkpoint = tf.keras.callbacks.ModelCheckpoint(checkpoint_path,
save_best_only=True,
monitor="val_loss")
```
Woohoo! Fine-tuning callbacks ready.
If you're planning on training large models, the `ModelCheckpoint` and `EarlyStopping` are two callbacks you'll want to become very familiar with.
We're almost ready to start fine-tuning our model but there's one more callback we're going to implement: [`ReduceLROnPlateau`](https://www.tensorflow.org/api_docs/python/tf/keras/callbacks/ReduceLROnPlateau).
Remember how the learning rate is the most important model hyperparameter you can tune? (if not, treat this as a reminder).
Well, the `ReduceLROnPlateau` callback helps to tune the learning rate for you.
Like the `ModelCheckpoint` and `EarlyStopping` callbacks, the `ReduceLROnPlateau` callback montiors a specified metric and when that metric stops improving, it reduces the learning rate by a specified factor (e.g. divides the learning rate by 10).
> 🤔 **Question:** *Why lower the learning rate?*
Imagine having a coin at the back of the couch and you're trying to grab with your fingers.
Now think of the learning rate as the size of the movements your hand makes towards the coin.
The closer you get, the smaller you want your hand movements to be, otherwise the coin will be lost.
Our model's ideal performance is the equivalent of grabbing the coin. So as training goes on and our model gets closer and closer to it's ideal performance (also called **convergence**), we want the amount it learns to be less and less.
To do this we'll create an instance of the `ReduceLROnPlateau` callback to monitor the validation loss just like the `EarlyStopping` callback.
Once the validation loss stops improving for two or more epochs, we'll reduce the learning rate by a factor of 5 (e.g. `0.001` to `0.0002`).
And to make sure the learning rate doesn't get too low (and potentially result in our model learning nothing), we'll set the minimum learning rate to `1e-7`.
```
# Creating learning rate reduction callback
reduce_lr = tf.keras.callbacks.ReduceLROnPlateau(monitor="val_loss",
factor=0.2, # multiply the learning rate by 0.2 (reduce by 5x)
patience=2,
verbose=1, # print out when learning rate goes down
min_lr=1e-7)
```
Learning rate reduction ready to go!
Now before we start training, we've got to recompile our model.
We'll use sparse categorical crossentropy as the loss and since we're fine-tuning, we'll use a 10x lower learning rate than the Adam optimizers default (`1e-4` instead of `1e-3`).
```
# Compile the model
loaded_gs_model.compile(loss="sparse_categorical_crossentropy", # sparse_categorical_crossentropy for labels that are *not* one-hot
optimizer=tf.keras.optimizers.Adam(0.0001), # 10x lower learning rate than the default
metrics=["accuracy"])
```
Okay, model compiled.
Now let's fit it on all of the data.
We'll set it up to run for up to 100 epochs.
Since we're going to be using the `EarlyStopping` callback, it might stop before reaching 100 epochs.
> 🔑 **Note:** Running the cell below will set the model up to fine-tune all of the pre-trained weights in the base model on all of the Food101 data. Doing so with **unoptimized** data pipelines and **without** mixed precision training will take a fairly long time per epoch depending on what type of GPU you're using (about 15-20 minutes on Colab GPUs). But don't worry, **the code we've written above will ensure it runs much faster** (more like 4-5 minutes per epoch).
```
# Start to fine-tune (all layers)
history_101_food_classes_all_data_fine_tune = loaded_gs_model.fit(train_data,
epochs=100, # fine-tune for a maximum of 100 epochs
steps_per_epoch=len(train_data),
validation_data=test_data,
validation_steps=int(0.15 * len(test_data)), # validation during training on 15% of test data
callbacks=[create_tensorboard_callback("training_logs", "efficientb0_101_classes_all_data_fine_tuning"), # track the model training logs
model_checkpoint, # save only the best model during training
early_stopping, # stop model after X epochs of no improvements
reduce_lr]) # reduce the learning rate after X epochs of no improvements
```
> 🔑 **Note:** If you didn't use mixed precision or use techniques such as [`prefetch()`](https://www.tensorflow.org/api_docs/python/tf/data/Dataset#prefetch) in the *Batch & prepare datasets* section, your model fine-tuning probably takes up to 2.5-3x longer per epoch (see the output below for an example).
| | Prefetch and mixed precision | No prefetch and no mixed precision |
|-----|-----|-----|
| Time per epoch | ~280-300s | ~1127-1397s |
*Results from fine-tuning 🍔👁 Food Vision Big™ on Food101 dataset using an EfficienetNetB0 backbone using a Google Colab Tesla T4 GPU.*
```
Saving TensorBoard log files to: training_logs/efficientB0_101_classes_all_data_fine_tuning/20200928-013008
Epoch 1/100
2368/2368 [==============================] - 1397s 590ms/step - loss: 1.2068 - accuracy: 0.6820 - val_loss: 1.1623 - val_accuracy: 0.6894
Epoch 2/100
2368/2368 [==============================] - 1193s 504ms/step - loss: 0.9459 - accuracy: 0.7444 - val_loss: 1.1549 - val_accuracy: 0.6872
Epoch 3/100
2368/2368 [==============================] - 1143s 482ms/step - loss: 0.7848 - accuracy: 0.7838 - val_loss: 1.0402 - val_accuracy: 0.7142
Epoch 4/100
2368/2368 [==============================] - 1127s 476ms/step - loss: 0.6599 - accuracy: 0.8149 - val_loss: 0.9599 - val_accuracy: 0.7373
```
*Example fine-tuning time for non-prefetched data as well as non-mixed precision training (~2.5-3x longer per epoch).*
Let's make sure we save our model before we start evaluating it.
```
# # Save model to Google Drive (optional)
# loaded_gs_model.save("/content/drive/MyDrive/tensorflow_course/food_vision/07_efficientnetb0_fine_tuned_101_classes_mixed_precision/")
# Save model locally (note: if you're using Google Colab and you save your model locally, it will be deleted when your Google Colab session ends)
loaded_gs_model.save("07_efficientnetb0_fine_tuned_101_classes_mixed_precision")
```
Looks like our model has gained a few performance points from fine-tuning, let's evaluate on the whole test dataset and see if managed to beat the [DeepFood paper's](https://arxiv.org/abs/1606.05675) result of 77.4% accuracy.
```
# Evaluate mixed precision trained loaded model
results_loaded_gs_model_fine_tuned = loaded_gs_model.evaluate(test_data)
results_loaded_gs_model_fine_tuned
```
Woohoo!!!! It looks like our model beat the results mentioned in the DeepFood paper for Food101 (DeepFood's 77.4% top-1 accuracy versus our ~79% top-1 accuracy).
## Download fine-tuned model from Google Storage
As mentioned before, training models can take a significant amount of time.
And again, like any good cooking show, here's something we prepared earlier...
It's a fine-tuned model exactly like the one we trained above but it's saved to Google Storage so it can be accessed, imported and evaluated.
```
# Download and evaluate fine-tuned model from Google Storage
!wget https://storage.googleapis.com/ztm_tf_course/food_vision/07_efficientnetb0_fine_tuned_101_classes_mixed_precision.zip
```
The downloaded model comes in zip format (`.zip`) so we'll unzip it into the Google Colab instance.
```
# Unzip fine-tuned model
!mkdir downloaded_fine_tuned_gs_model # create separate directory for fine-tuned model downloaded from Google Storage
!unzip /content/07_efficientnetb0_fine_tuned_101_classes_mixed_precision -d downloaded_fine_tuned_gs_model
```
Now we can load it using the [`tf.keras.models.load_model()`](https://www.tensorflow.org/tutorials/keras/save_and_load) method and get a summary (it should be the exact same as the model we created above).
```
# Load in fine-tuned model from Google Storage and evaluate
loaded_fine_tuned_gs_model = tf.keras.models.load_model("/content/downloaded_fine_tuned_gs_model/07_efficientnetb0_fine_tuned_101_classes_mixed_precision")
# Get a model summary (same model architecture as above)
loaded_fine_tuned_gs_model.summary()
```
Finally, we can evaluate our model on the test data (this requires the `test_data` variable to be loaded.
```
# Note: Even if you're loading in the model from Google Storage, you will still need to load the test_data variable for this cell to work
results_downloaded_fine_tuned_gs_model = loaded_fine_tuned_gs_model.evaluate(test_data)
results_downloaded_fine_tuned_gs_model
```
Excellent! Our saved model is performing as expected (better results than the DeepFood paper!).
Congratulations! You should be excited! You just trained a computer vision model with competitive performance to a research paper and in far less time (our model took ~20 minutes to train versus DeepFood's quoted 2-3 days).
In other words, you brought Food Vision life!
If you really wanted to step things up, you could try using the [`EfficientNetB4`](https://www.tensorflow.org/api_docs/python/tf/keras/applications/EfficientNetB4) model (a larger version of `EfficientNetB0`). At at the time of writing, the EfficientNet family has the [state of the art classification results](https://paperswithcode.com/sota/fine-grained-image-classification-on-food-101) on the Food101 dataset.
> 📖 **Resource:** To see which models are currently performing the best on a given dataset or problem type as well as the latest trending machine learning research, be sure to check out [paperswithcode.com](http://paperswithcode.com/) and [sotabench.com](https://sotabench.com/).
## View training results on TensorBoard
Since we tracked our model's fine-tuning training logs using the `TensorBoard` callback, let's upload them and inspect them on TensorBoard.dev.
```
!tensorboard dev upload --logdir ./training_logs \
--name "Fine-tuning EfficientNetB0 on all Food101 Data" \
--description "Training results for fine-tuning EfficientNetB0 on Food101 Data with learning rate 0.0001" \
--one_shot
View experiment: https://tensorboard.dev/experiment/2KINdYxgSgW2bUg7dIvevw/
```
Viewing at our [model's training curves on TensorBoard.dev](https://tensorboard.dev/experiment/2KINdYxgSgW2bUg7dIvevw/), it looks like our fine-tuning model gains boost in performance but starts to overfit as training goes on.
See the training curves on TensorBoard.dev here: https://tensorboard.dev/experiment/2KINdYxgSgW2bUg7dIvevw/
To fix this, in future experiments, we might try things like:
* A different iteration of `EfficientNet` (e.g. `EfficientNetB4` instead of `EfficientNetB0`).
* Unfreezing less layers of the base model and training them rather than unfreezing the whole base model in one go.
```
# View past TensorBoard experiments
!tensorboard dev list
# Delete past TensorBoard experiments
# !tensorboard dev delete --experiment_id YOUR_EXPERIMENT_ID
# Example
!tensorboard dev delete --experiment_id OAE6KXizQZKQxDiqI3cnUQ
```
## 🛠 Exercises
1. Use the same evaluation techniques on the large-scale Food Vision model as you did in the previous notebook ([Transfer Learning Part 3: Scaling up](https://github.com/mrdbourke/tensorflow-deep-learning/blob/main/06_transfer_learning_in_tensorflow_part_3_scaling_up.ipynb)). More specifically, it would be good to see:
* A confusion matrix between all of the model's predictions and true labels.
* A graph showing the f1-scores of each class.
* A visualization of the model making predictions on various images and comparing the predictions to the ground truth.
* For example, plot a sample image from the test dataset and have the title of the plot show the prediction, the prediction probability and the ground truth label.
2. Take 3 of your own photos of food and use the Food Vision model to make predictions on them. How does it go? Share your images/predictions with the other students.
3. Retrain the model (feature extraction and fine-tuning) we trained in this notebook, except this time use [`EfficientNetB4`](https://www.tensorflow.org/api_docs/python/tf/keras/applications/EfficientNetB4) as the base model instead of `EfficientNetB0`. Do you notice an improvement in performance? Does it take longer to train? Are there any tradeoffs to consider?
4. Name one important benefit of mixed precision training, how does this benefit take place?
## 📖 Extra-curriculum
* Read up on learning rate scheduling and the [learning rate scheduler callback](https://www.tensorflow.org/api_docs/python/tf/keras/callbacks/LearningRateScheduler). What is it? And how might it be helpful to this project?
* Read up on TensorFlow data loaders ([improving TensorFlow data loading performance](https://www.tensorflow.org/guide/data_performance)). Is there anything we've missed? What methods you keep in mind whenever loading data in TensorFlow? Hint: check the summary at the bottom of the page for a great round up of ideas.
* Read up on the documentation for [TensorFlow mixed precision training](https://www.tensorflow.org/guide/mixed_precision). What are the important things to keep in mind when using mixed precision training?
| github_jupyter |
# Using the SDFG API in DaCe
In this tutorial, we will create an SDFG manually using the SDFG API. This interface gives full control over the representation, and it is also the one used for developing new transformations and other graph manipulation.
The code we will write executes a stencil in a sequence (without boundary conditions). In SDFG terms, it is a sequential for-loop (state machine) of tasklets nested in maps.
```
import dace
import numpy as np
```
We begin by defining the temporal and spatial dimensions as symbols:
```
T = dace.symbol('T')
N = dace.symbol('N')
```
Creating an SDFG requires giving it a name (which will be used in compilation to create the library files and function names):
```
sdfg = dace.SDFG('jacobi2d')
```
Then, we need to define the set of data descriptors used throughout the Data nodes in the SDFG. Since we use a double-buffering approach, we define a 2D array `A` and a 2D array `tmp`. `tmp` is transient, which means it is not an input/output of the SDFG, and can thus participate in transformations.
```
sdfg.add_array('A', shape=[N, N], dtype=dace.float32)
sdfg.add_transient('tmp', shape=[N, N], dtype=dace.float32)
```
Next, we construct a state, which will contain the main computational part:
```
state = sdfg.add_state()
sdfg
```
Now the SDFG contains only one empty state. We will create the contents of the main state, which is two stencils, `A->tmp` and `tmp->A`. Since the code is equivalent, we define a function once and call it twice:
```
def mainstate(state, src_node, dst_node):
# Creates Map (entry and exit nodes), Tasklet node, and connects the three
tasklet, map_entry, map_exit = state.add_mapped_tasklet(
'%s_to_%s' % (src_node.data, dst_node.data), # name
dict(i='1:N-1', j='1:N-1'), # map range
dict(inp=dace.Memlet.simple(src_node.data, # input memlets
'i-1:i+2, j-1:j+2')),
''' # code
out = 0.2 * (inp[0,1] + inp[1,0] + inp[1,1] + # (5-point Jacobi)
inp[1,2] + inp[2,1])
''',
dict(out=dace.Memlet.simple(dst_node.data, 'i,j')) # output memlets
)
#######################
# Add external connections from map to arrays
# Add input path (src->entry) with the overall memory accessed
# NOTE: This can be inferred automatically by the system
# using external_edges=True in `add_mapped_tasklet`
# or using the `propagate_edge` function.
state.add_edge(
src_node, None,
map_entry, None,
memlet=dace.Memlet.simple(src_node.data, '0:N, 0:N'))
# Add output path (exit->dst)
state.add_edge(
map_exit, None,
dst_node, None,
memlet=dace.Memlet.simple(dst_node.data, '1:N-1, 1:N-1'))
```
We add and connect the read, access (read/write), and write nodes for the main state, as well as the code:
```
A_in = state.add_read('A')
tmp = state.add_access('tmp')
A_out = state.add_write('A')
sdfg
mainstate(state, A_in, tmp)
mainstate(state, tmp, A_out)
sdfg
```
Notice the boxes inside the tasklet nodes, connected to the edges. These are **connectors**, the way to identify the edge's behavior. A connector has a type and shape, just like arrays, and is also used to create unique paths through map scopes.
In the above case, it is clear that the edges leading to and from the map entry/exit nodes form a path. However, when multiple edges are involved, it may be ambiguous. To uniquely identify paths, scope entry/exit nodes can have input connectors that begin with `IN_` and output connectors that begin with `OUT_`. As a convenience function, an SDFG can try to fill its scope connectors on its own (using the data names on the memlets):
```
sdfg.fill_scope_connectors()
sdfg
```
This single-state SDFG is now valid, as no exceptions are raised below:
```
sdfg.validate()
```
However, it only runs for two time-steps, and will produce incorrect results, due to the boundaries of `tmp`. We thus need to define a starting state that sets `tmp` to zero, and a looping state machine. Initialization state is defined below:
```
bstate = sdfg.add_state('begin') # States can be named
# We use the convenience parameter external_edges to add the tmp array node and connectors
bstate.add_mapped_tasklet('init_tmp', dict(i='0:N', j='0:N'), {}, # no inputs
'out = 0', dict(out=dace.Memlet.simple('tmp', 'i,j')),
external_edges=True)
sdfg
```
The loop will be defined in the same manner as C for-loops: A `guard` state, which jumps into the loop as long as it is in range; an `end` (empty) state; and the `loop` state (currently our main state), which jumps back to the guard and increments the iteration variable. Notice that adding edges on the SDFG (as opposed to adding them in states) requires a different edge object type: `InterstateEdge`.
```
guard = sdfg.add_state('guard')
endstate = sdfg.add_state('endstate')
# State connection (control flow)
# Note: dataflow (arrays) CAN affect control flow assignments and conditions,
# but not the other way around (you cannot change an interstate variable
# inside a state). The following code works as well:
#sdfg.add_edge(state0, guard, dace.InterstateEdge(assigments=dict('k', 'A[0]')))
# Loop initialization (k=0)
sdfg.add_edge(bstate, guard, dace.InterstateEdge(assignments=dict(k='0')))
# Loop condition (k < T / k >= T)
sdfg.add_edge(guard, state, dace.InterstateEdge('k < T'))
sdfg.add_edge(guard, endstate, dace.InterstateEdge('k >= T'))
# Loop incrementation (k++)
sdfg.add_edge(
state, guard, dace.InterstateEdge(assignments=dict(k='k+1')))
# Display resulting SDFG
sdfg
```
And the SDFG is complete. Now all that is left is to execute it and validate the results:
```
from scipy import ndimage
# Symbol values
N = 24
T = 5
# Arrays
inp = np.zeros(shape=(N, N), dtype=np.float32)
inp[1:N-1, 1:N-1] = np.random.rand(N-2, N-2).astype(np.float32)
expected = np.copy(inp[1:N-1, 1:N-1])
kernel = np.array([[0, 0.2, 0], [0.2, 0.2, 0.2], [0, 0.2, 0]], dtype=np.float32)
# Evaluate expected result
for k in range(T * 2):
expected = ndimage.convolve(
expected, kernel, mode='constant', cval=0.0)
sdfg(A=inp, N=N, T=T)
print('Difference:', np.linalg.norm(expected - inp[1:N-1, 1:N-1]))
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import nltk
from collections import Counter
from sklearn.metrics import log_loss
from scipy.optimize import minimize
import multiprocessing
import difflib
import time
import gc
import xgboost as xgb
from sklearn.cross_validation import train_test_split
from sklearn.feature_extraction.text import TfidfVectorizer, CountVectorizer
from xgb_utils import *
def get_train():
keras_q1 = np.load('../features/q1train_spacylemmat_fullclean_170len_treetrunc.npy')
keras_q2 = np.load('../features/q2train_spacylemmat_fullclean_170len_treetrunc.npy')
xgb_feats = pd.read_csv('../../data/features/the_1owl/owl_train.csv')
abhishek_feats = pd.read_csv('../../data/features/abhishek/train_features.csv',
encoding = 'ISO-8859-1').iloc[:, 2:]
text_feats = pd.read_csv('../../data/features/other_features/text_features_train.csv',
encoding = 'ISO-8859-1')
img_feats = pd.read_csv('../../data/features/other_features/img_features_train.csv')
srk_feats = pd.read_csv('../../data/features/srk/SRK_grams_features_train.csv')
turkewitz_feats = pd.read_csv('../../data/features/lemmat_spacy_features/train_turkewitz_features.csv')
turkewitz_feats = turkewitz_feats[['q1_freq', 'q2_freq']]
xgb_feats.drop(['z_len1', 'z_len2', 'z_word_len1', 'z_word_len2'], axis = 1, inplace = True)
y_train = xgb_feats['is_duplicate']
xgb_feats = xgb_feats.iloc[:, 8:]
X_train2 = np.concatenate([keras_q1, keras_q2, xgb_feats, abhishek_feats, text_feats, img_feats,
turkewitz_feats], axis = 1)
for i in range(X_train2.shape[1]):
if np.sum(X_train2[:, i] == y_train.values) == X_train2.shape[0]:
print('LEAK FOUND')
X_train2 = X_train2.astype('float32')
X_train2 = pd.DataFrame(X_train2)
X_train2['is_duplicate'] = y_train
print('Training data shape:', X_train2.shape)
return X_train2, y_train
def get_test():
keras_q1 = np.load('../features/q1train_spacylemmat_fullclean_170len_treetrunc.npy')
keras_q2 = np.load('../features/q2train_spacylemmat_fullclean_170len_treetrunc.npy')
xgb_feats = pd.read_csv('../../data/features/the_1owl/owl_test.csv')
abhishek_feats = pd.read_csv('../../data/features/abhishek/test_features.csv',
encoding = 'ISO-8859-1').iloc[:, 2:]
text_feats = pd.read_csv('../../data/features/other_features/text_features_test.csv',
encoding = 'ISO-8859-1')
img_feats = pd.read_csv('../../data/features/other_features/img_features_test.csv')
srk_feats = pd.read_csv('../../data/features/srk/SRK_grams_features_test.csv')
turkewitz_feats = pd.read_csv('../../data/features/lemmat_spacy_features/test_turkewitz_features.csv')
turkewitz_feats = turkewitz_feats[['q1_freq', 'q2_freq']]
xgb_feats.drop(['z_len1', 'z_len2', 'z_word_len1', 'z_word_len2'], axis = 1, inplace = True)
xgb_feats = xgb_feats.iloc[:, 5:]
X_test2 = np.concatenate([keras_q1, keras_q2, xgb_feats, abhishek_feats, text_feats, img_feats,
turkewitz_feats], axis = 1)
X_test2 = X_test2.astype('float32')
X_test2 = pd.DataFrame(X_test2)
print('Test data shape:', X_test2.shape)
return X_test2
def predict_test(model_name):
print('Predicting on test set.')
X_test = get_test()
gbm = xgb.Booster(model_file = 'saved_models/XGB/{}.txt'.format(model_name))
test_preds = gbm.predict(xgb.DMatrix(X_test))
sub_src = '/media/w/1c392724-ecf3-4615-8f3c-79368ec36380/DS Projects/Kaggle/Quora/submissions/'
sample_sub = pd.read_csv(sub_src + 'sample_submission.csv')
sample_sub['is_duplicate'] = test_preds
sample_sub.is_duplicate = sample_sub.is_duplicate.apply(transform)
sample_sub.to_csv(sub_src + '{}.csv'.format(model_name), index = False)
return
def train_xgb(cv = False):
t = time.time()
params = {
'seed': 1337,
'colsample_bytree': 0.48,
'silent': 1,
'subsample': 0.74,
'eta': 0.05,
'objective': 'binary:logistic',
'eval_metric': 'logloss',
'max_depth': 12,
'min_child_weight': 20,
'nthread': 8,
'tree_method': 'hist',
#'updater': 'grow_gpu',
}
X_train, y_train = get_train()
X_train = X_train.astype('float32')
X_train.drop(['is_duplicate'], axis = 1, inplace = True)
if cv:
dtrain = xgb.DMatrix(X_train, y_train)
hist = xgb.cv(params, dtrain, num_boost_round = 100000, nfold = 5,
stratified = True, early_stopping_rounds = 350, verbose_eval = 250,
seed = 1337)
del X_train, y_train
gc.collect()
print('Time it took to train in CV manner:', time.time() - t)
return hist
else:
X_tr, X_val, y_tr, y_val = train_test_split(X_train, y_train, stratify = y_train,
test_size = 0.2, random_state = 111)
del X_train, y_train
gc.collect()
dtrain = xgb.DMatrix(X_tr, label = y_tr)
dval = xgb.DMatrix(X_val, label = y_val)
watchlist = [(dtrain, 'train'), (dval, 'valid')]
print('Start training...')
gbm = xgb.train(params, dtrain, 100000, watchlist,
early_stopping_rounds = 350, verbose_eval = 250)
print('Start predicting...')
val_pred = gbm.predict(xgb.DMatrix(X_val), ntree_limit=gbm.best_ntree_limit)
score = log_loss(y_val, val_pred)
print('Final score:', score, '\n', 'Time it took to train and predict:', time.time() - t)
del X_tr, X_val, y_tr, y_val
gc.collect()
return gbm
def run_xgb(model_name, train = True, test = False, cv = False):
if cv:
gbm_hist = train_xgb(True)
return gbm_hist
if train:
gbm = train_xgb()
gbm.save_model('saved_models/XGB/{}.txt'.format(model_name))
if test:
predict_test('{}'.format(model_name))
return gbm
gbm = run_xgb('XGB_firstBO_turkewitz_Qspacyencode', train = True, test = True)
predict_test('XGB_firstBO_turkewitz_Qspacyencode')
```
| github_jupyter |


# Bitmapped Graphics
## Background image
Loading an image is a simple process and involves only one line of code.
Example:
`pygame.image.load("saturn_family1.jpg")`
Normally We need a variable set equal to what the `load()` command returns.
In the next version of our load command, we create a new variable named background_image. See below for version two:
`background_image = pygame.image.load("saturn_family1.jpg").convert()`
Finally, the image needs to be converted to a format Pygame can more easily work with.
To do that, we append `.convert()` to the command to call the convert function.
The function .convert() is a method in the Image class.
Loading the image should be done before the main program loop
To display the image use the `blit` command.
This “blits” the image bits to the screen. We've already used this command once before when displaying text
## Moving an image
Suppose you need an image to be used as player, to be moved by the mouse.
The image can be downloaded from Internet, or you can find a .gif or .png that you like with a white or black background.
Don't use a .jpg.
To load the image we need the same type of command that we used with the background image
```python
player_image = pygame.image.load("player.png").convert()
# Get the current mouse position. This returns the position
# as a list of two numbers.
player_position = pygame.mouse.get_pos()
x = player_position[0]
y = player_position[1]
# Copy image to screen:
screen.blit(player_image, [x, y])
```
## Setting proper background
All images are rectangular.
How do we show only the part of the image we want?
he way to get around this is to tell the program to make one color “transparent” and not display.
For example, if it is black:
`player_image.set_colorkey(BLACK)`
This will work for most files ending in .gif and .png.
This does not work well for most .jpg files because of their compression.
# Sounds
Like images, sounds must be loaded before they are used.
This should be done once sometime before the main program loop.
Uncompressed sound files usually end in **.wav**.
These files are larger than other formats because no algorithm has been run on them to make them smaller.
There is also the ever popular **.mp3** format, although that format has patents that can make it undesirable for certain applications.
Another format that is free to use is the OGG Vorbis format that ends in **.ogg**.
With free tools like **Audacity** is possible to convert between formats.
## Associated to events
```python
click_sound = pygame.mixer.Sound("laser5.ogg")
for event in pygame.event.get():
if event.type == pygame.QUIT:
done = True
elif event.type == pygame.MOUSEBUTTONDOWN:
click_sound.play()
```
## In background
Loop repeated
```python
bgk_sound = pygame.mixer.Sound("song.ogg")
bgk_sound.play(-1)
```
# Example
See `04_bitmapped_sounds.py`
# Exercises
## Test
http://programarcadegames.com/quiz/quiz.php?file=bitmapped_graphics&lang=en
## Assignments
Create a graphics based program. You can start a new program, or continue with a prior lab.
# Bibliography
Program Arcade Games With Python And Pygame, ch. 11
http://programarcadegames.com/index.php
| github_jupyter |
<a href="https://colab.research.google.com/github/NeuromatchAcademy/course-content/blob/master/tutorials/W1D3_ModelFitting/W1D3_Tutorial6.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Neuromatch Academy: Week 1, Day 3, Tutorial 6
# Model Selection: Cross-validation
**Content creators**: Pierre-Étienne Fiquet, Anqi Wu, Alex Hyafil with help from Ella Batty
**Content reviewers**: Lina Teichmann, Patrick Mineault, Michael Waskom
---
#Tutorial Objectives
This is Tutorial 6 of a series on fitting models to data. We start with simple linear regression, using least squares optimization (Tutorial 1) and Maximum Likelihood Estimation (Tutorial 2). We will use bootstrapping to build confidence intervals around the inferred linear model parameters (Tutorial 3). We'll finish our exploration of regression models by generalizing to multiple linear regression and polynomial regression (Tutorial 4). We end by learning how to choose between these various models. We discuss the bias-variance trade-off (Tutorial 5) and Cross Validation for model selection (Tutorial 6).
Tutorial objectives:
* Implement cross-validation and use it to compare polynomial regression model
---
# Setup
```
import numpy as np
import matplotlib.pyplot as plt
from sklearn.model_selection import KFold
#@title Figure Settings
%config InlineBackend.figure_format = 'retina'
plt.style.use("https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/nma.mplstyle")
#@title Helper functions
def ordinary_least_squares(x, y):
"""Ordinary least squares estimator for linear regression.
Args:
x (ndarray): design matrix of shape (n_samples, n_regressors)
y (ndarray): vector of measurements of shape (n_samples)
Returns:
ndarray: estimated parameter values of shape (n_regressors)
"""
return np.linalg.inv(x.T @ x) @ x.T @ y
def make_design_matrix(x, order):
"""Create the design matrix of inputs for use in polynomial regression
Args:
x (ndarray): input vector of shape (n_samples)
order (scalar): polynomial regression order
Returns:
ndarray: design matrix for polynomial regression of shape (samples, order+1)
"""
# Broadcast to shape (n x 1)
if x.ndim == 1:
x = x[:, None]
#if x has more than one feature, we don't want multiple columns of ones so we assign
# x^0 here
design_matrix = np.ones((x.shape[0], 1))
# Loop through rest of degrees and stack columns
for degree in range(1, order + 1):
design_matrix = np.hstack((design_matrix, x**degree))
return design_matrix
def solve_poly_reg(x, y, max_order):
"""Fit a polynomial regression model for each order 0 through max_order.
Args:
x (ndarray): input vector of shape (n_samples)
y (ndarray): vector of measurements of shape (n_samples)
max_order (scalar): max order for polynomial fits
Returns:
dict: fitted weights for each polynomial model (dict key is order)
"""
# Create a dictionary with polynomial order as keys, and np array of theta
# (weights) as the values
theta_hats = {}
# Loop over polynomial orders from 0 through max_order
for order in range(max_order + 1):
X = make_design_matrix(x, order)
this_theta = ordinary_least_squares(X, y)
theta_hats[order] = this_theta
return theta_hats
def evaluate_poly_reg(x, y, theta_hats, max_order):
""" Evaluates MSE of polynomial regression models on data
Args:
x (ndarray): input vector of shape (n_samples)
y (ndarray): vector of measurements of shape (n_samples)
theta_hat (dict): fitted weights for each polynomial model (dict key is order)
max_order (scalar): max order of polynomial fit
Returns
(ndarray): mean squared error for each order, shape (max_order)
"""
mse = np.zeros((max_order + 1))
for order in range(0, max_order + 1):
X_design = make_design_matrix(x, order)
y_hat = np.dot(X_design, theta_hats[order])
residuals = y - y_hat
mse[order] = np.mean(residuals ** 2)
return mse
```
---
# Section 1: Cross-validation
```
#@title Video 1: Cross-Validation
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id='BV1mt4y1Q7C4', width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
video
```
We now have multiple choices for which model to use for a given problem: we could use linear regression, order 2 polynomial regression, order 3 polynomial regression, etc. As we saw in Tutorial 5, different models will have different quality of predictions, both on the training data and on the test data.
A commonly used method for model selection is to asks how well the model predicts new data that it hasn't seen yet. But we don't want to use test data to do this, otherwise that would mean using it during the training process! One approach is to use another kind of held-out data which we call **validation data**: we do not fit the model with this data but we use it to select our best model.
We often have a limited amount of data though (especially in neuroscience), so we do not want to further reduce our potential training data by reassigning some as validation. Luckily, we can use **k-fold cross-validation**! In k-fold cross validation, we divide up the training data into k subsets (that are called *folds*, see diagram below), train our model on the first k-1 folds, and then compute error on the last held-out fold. We can then repeat this process k times, once on each k-1 folds of the data. Each of these k instances (which are called *splits*, see diagram below) excludes a different fold from fitting. We then average the error of each of the k trained models on its held-out subset - this is the final measure of performance which we can use to do model selection.
To make this explicit, let's say we have 1000 samples of training data and choose 4-fold cross-validation. Samples 0 - 250 would be subset 1, samples 250 - 500 subset 2, samples 500 - 750 subset 3, and samples 750-1000 subset 4. First, we train an order 3 polynomial regression on subsets 1, 2, 3 and evaluate on subset 4. Next, we train an order 3 polynomial model on subsets 1, 2, 4 and evalute on subset 3. We continue until we have 4 instances of a trained order 3 polynomial regression model, each with a different subset as held-out data, and average the held-out error from each instance.
We can now compare the error of different models to pick a model that generalizes well to held-out data. We can choose the measure of prediction quality to report error on the held-out subsets to suit our purposes. We will use MSE here but we could also use log likelihood of the data and so on.
As a final step, it is common to retrain this model on all of the training data (without subset divisions) to get our final model that we will evaluate on test data. This approach allows us to evaluate the quality of predictions on new data without sacrificing any of our precious training data.
Note that the held-out subsets are called either validation or test subsets. There is not a consensus and may depend on the exact use of k-fold cross validation. Sometimes people use k-fold cross validation to choose between different models/parameters to then apply to held-out test data and sometimes people report the averaged error on the held-out subsets as the model performance. If you are doing the former (using k-fold cross validation for model selection), you must report performance on held-out test data! In this text/code, we will refer to them as validation subsets to differentiate from our completely held-out test data.
These steps are summarized in this diagram from Scikit-learn (https://scikit-learn.org/stable/modules/cross_validation.html)
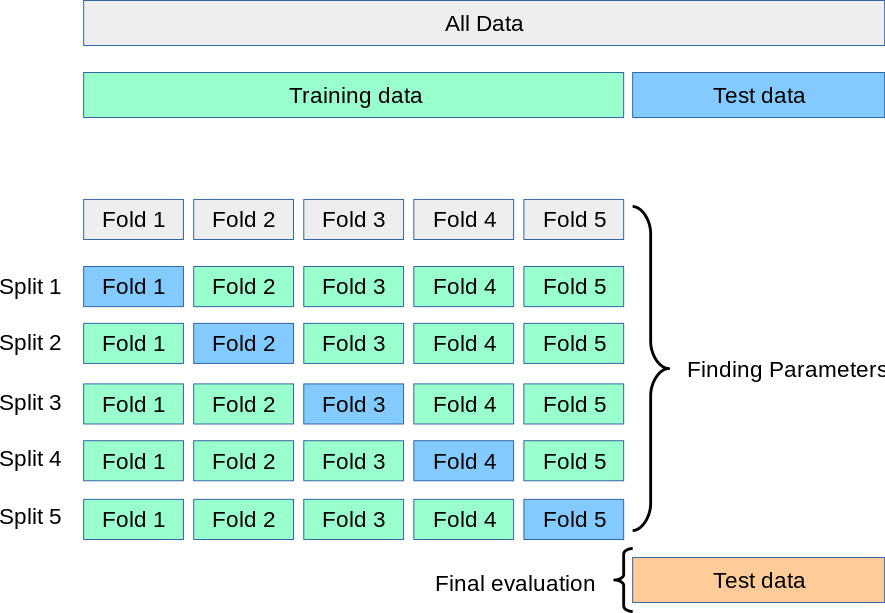
Importantly, we need to be very careful when dividing the data into subsets. The held-out subset should not be used in any way to fit the model. We should not do any preprocessing (e.g. normalization) before we divide into subsets or the held-out subset could influence the training subsets. A lot of false-positives in cross-validation come from wrongly dividing.
An important consideration in the choice of model selection method are the relevant biases. If we just fit using MSE on training data, we will generally find that fits get better as we add more parameters because the model will overfit the data, as we saw in Tutorial 5. When using cross-validation, the bias is the other way around. Models with more parameters are more affected by variance so cross-validation will generally prefer models with fewer parameters.
We will again simulate some train and test data and fit polynomial regression models
```
#@title
#@markdown Execute this cell to simulate data and fit polynomial regression models
### Generate training data
np.random.seed(0)
n_train_samples = 50
x_train = np.random.uniform(-2, 2.5, n_train_samples) # sample from a uniform distribution over [-2, 2.5)
noise = np.random.randn(n_train_samples) # sample from a standard normal distribution
y_train = x_train**2 - x_train - 2 + noise
### Generate testing data
n_test_samples = 20
x_test = np.random.uniform(-3, 3, n_test_samples) # sample from a uniform distribution over [-2, 2.5)
noise = np.random.randn(n_test_samples) # sample from a standard normal distribution
y_test = x_test**2 - x_test - 2 + noise
### Fit polynomial regression models
max_order = 5
theta_hats = solve_poly_reg(x_train, y_train, max_order)
```
## Exercise 1: Implement cross-validation
Given our set of models to evaluate (polynomial regression models with orders 0 through 5), we will use cross-validation to determine which model has the best predictions on new data according to MSE.
In this code, we split the data into 10 subsets using `Kfold` (from `sklearn.model_selection`). `KFold` handles cross-validation subset splitting and train/val assignments. In particular, the `Kfold.split` method returns an iterator which we can loop through. On each loop, this iterator assigns a different subset as validation and returns new training and validation indices with which to split the data.
We will loop through the 10 train/validation splits and fit several different polynomial regression models (with different orders) for each split. You will need to use the `solve_poly_reg` method from Tutorial 4 and `evaluate_poly_reg` from Tutorial 5 (already implemented in this notebook).
We will visualize the validation MSE over 10 splits of the data for each polynomial order using box plots.
```
def cross_validate(x_train, y_train, max_order, n_splits):
""" Compute MSE for k-fold validation for each order polynomial
Args:
x_train (ndarray): training data input vector of shape (n_samples)
y_train (ndarray): training vector of measurements of shape (n_samples)
max_order (scalar): max order of polynomial fit
n_split (scalar): number of folds for k-fold validation
Return:
ndarray: MSE over splits for each model order, shape (n_splits, max_order + 1)
"""
# Initialize the split method
kfold_iterator = KFold(n_splits)
# Initialize np array mse values for all models for each split
mse_all = np.zeros((n_splits, max_order + 1))
for i_split, (train_indices, val_indices) in enumerate(kfold_iterator.split(x_train)):
# Split up the overall training data into cross-validation training and validation sets
x_cv_train = x_train[train_indices]
y_cv_train = y_train[train_indices]
x_cv_val = x_train[val_indices]
y_cv_val = y_train[val_indices]
#############################################################################
## TODO for students: Fill in missing ... in code below to choose which data
## to fit to and compute MSE for
# Fill out function and remove
raise NotImplementedError("Student exercise: implement cross-validation")
#############################################################################
# Fit models
theta_hats = ...
# Compute MSE
mse_this_split = ...
mse_all[i_split] = mse_this_split
return mse_all
# Uncomment below to test function
max_order = 5
n_splits = 10
plt.figure()
#mse_all = cross_validate(x_train, y_train, max_order, n_splits)
#plt.boxplot(mse_all, labels=np.arange(0, max_order + 1))
plt.xlabel('Polynomial Order')
plt.ylabel('Validation MSE')
plt.title(f'Validation MSE over {n_splits} splits of the data');
# to_remove solution
def cross_validate(x_train, y_train, max_order, n_splits):
""" Compute MSE for k-fold validation for each order polynomial
Args:
x_train (ndarray): training data input vector of shape (n_samples)
y_train (ndarray): training vector of measurements of shape (n_samples)
max_order (scalar): max order of polynomial fit
n_split (scalar): number of folds for k-fold validation
Return:
ndarray: MSE over splits for each model order, shape (n_splits, max_order + 1)
"""
# Initialize the split method
kfold_iterator = KFold(n_splits)
# Initialize np array mse values for all models for each split
mse_all = np.zeros((n_splits, max_order + 1))
for i_split, (train_indices, val_indices) in enumerate(kfold_iterator.split(x_train)):
# Split up the overall training data into cross-validation training and validation sets
x_cv_train = x_train[train_indices]
y_cv_train = y_train[train_indices]
x_cv_val = x_train[val_indices]
y_cv_val = y_train[val_indices]
# Fit models
theta_hats = solve_poly_reg(x_cv_train, y_cv_train, max_order)
# Compute MSE
mse_this_split = evaluate_poly_reg(x_cv_val, y_cv_val, theta_hats, max_order)
mse_all[i_split] = mse_this_split
return mse_all
max_order = 5
n_splits = 10
with plt.xkcd():
plt.figure()
mse_all = cross_validate(x_train, y_train, max_order, n_splits)
plt.boxplot(mse_all, labels=np.arange(0, max_order + 1))
plt.xlabel('Polynomial Order')
plt.ylabel('Validation MSE')
plt.title(f'Validation MSE over {n_splits} splits of the data');
```
Which polynomial order do you think is a better model of the data?
---
# Summary
We need to use model selection methods to determine the best model to use for a given problem.
Cross-validation focuses on how well the model predicts new data.
---
# Appendix
## Akaike's Information Criterion (AIC)
In order to choose the best model for a given problem, we can ask how likely the data is under a given model. We want to choose a model that assigns high probability to the data. A commonly used method for model selection that uses this approach is **Akaike’s Information Criterion (AIC)**.
Essentially, AIC estimates how much information would be lost if the model predictions were used instead of the true data (the relative information value of the model). We compute the AIC for each model and choose the model with the lowest AIC. Note that AIC only tells us relative qualities, not absolute - we do not know from AIC how good our model is independent of others.
AIC strives for a good tradeoff between overfitting and underfitting by taking into account the complexity of the model and the information lost. AIC is calculated as:
$$ AIC = 2K - 2 log(L)$$
where K is the number of parameters in your model and L is the likelihood that the model could have produced the output data.
Now we know what AIC is, we want to use it to pick between our polynomial regression models. We haven't been thinking in terms of likelihoods though - so how will we calculate L?
As we saw in Tutorial 2, there is a link between mean squared error and the likelihood estimates for linear regression models that we can take advantage of.
*Derivation time!*
We start with our formula for AIC from above:
$$ AIC = 2k - 2 log L $$
For a model with normal errors, we can use the log likelihood of the normal distribution:
$$ \log L = -\frac{n}{2} \log(2 \pi) -\frac{n}{2}log(\sigma^2) - \sum_i^n \frac{1}{2 \sigma^2} (y_i - \tilde y_i)^2$$
We can drop the first and last terms as both are constants and we're only assessing relative information with AIC. Once we drop those terms and incorporate into the AIC formula we get:
$$AIC = 2k + nlog(\sigma^2)$$
We can replace $\sigma^2$ with the computation for variance (the sum of squared errors divided by number of samples). Thus, we end up with the following formula for AIC for linear and polynomial regression:
$$ AIC = 2K + n log(\frac{SSE}{n})$$
where k is the number of parameters, n is the number of samples, and SSE is the summed squared error.
### Bonus Exercise: Compute AIC
```
AIC = np.zeros((max_order + 1))
for order in range(0, max_order + 1):
# Compute predictions for this model
X_design = make_design_matrix(x_train, order)
y_hat = np.dot(X_design, theta_hats[order])
#####################################################################################################
## TODO for students: Compute AIC for this order polynomial regression model
# 1) Compute sum of squared errors given prediction y_hat and y_train (SSE in formula above)
# 2) Identify number of parameters in this model (K in formula above)
# 3) Compute AIC (call this_AIC) according to formula above
#####################################################################################################
# Compute SSE
#residuals = ...
#sse = ...
# Get K
# K = len(theta_hats[order])
# Compute AIC
# AIC[order] = ...
plt.bar(range(max_order + 1), AIC);
plt.ylabel('AIC')
plt.xlabel('polynomial order')
plt.title('comparing polynomial fits')
plt.show()
# to_remove solution
AIC = np.zeros((max_order + 1))
for order in range(0, max_order + 1):
# Compute predictions for this model
X_design = make_design_matrix(x_train, order)
y_hat = np.dot(X_design, theta_hats[order])
# Compute SSE
residuals = y_train - y_hat
sse = np.sum(residuals ** 2)
# Get K
K = len(theta_hats[order])
# Compute AIC
AIC[order] = 2*K + n_train_samples * np.log(sse/n_train_samples)
with plt.xkcd():
plt.bar(range(max_order + 1), AIC);
plt.ylabel('AIC')
plt.xlabel('polynomial order')
plt.title('comparing polynomial fits')
plt.show()
```
Which model would we choose based on AIC?
| github_jupyter |
```
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# Plot parameters
sns.set()
%pylab inline
pylab.rcParams['figure.figsize'] = (4, 4)
plt.rcParams['xtick.major.size'] = 0
plt.rcParams['ytick.major.size'] = 0
# Avoid inaccurate floating values (for inverse matrices in dot product for instance)
# See https://stackoverflow.com/questions/24537791/numpy-matrix-inversion-rounding-errors
np.set_printoptions(suppress=True)
%%html
<style>
.pquote {
text-align: left;
margin: 40px 0 40px auto;
width: 70%;
font-size: 1.5em;
font-style: italic;
display: block;
line-height: 1.3em;
color: #5a75a7;
font-weight: 600;
border-left: 5px solid rgba(90, 117, 167, .1);
padding-left: 6px;
}
.notes {
font-style: italic;
display: block;
margin: 40px 10%;
}
img + em {
text-align: center;
display: block;
color: gray;
font-size: 0.9em;
font-weight: 600;
}
</style>
```
$$
\newcommand\bs[1]{\boldsymbol{#1}}
\newcommand\norm[1]{\left\lVert#1\right\rVert}
$$
# Introduction
In this lesson, we'll introduce ourselves to an important concept for machine learning and deep learning: the norm. Norms are what we generally use to evaluate the error of our models. For instance, it is used to calculate the error between the output of a neural network and what is expected (the actual label or value). You can think of the norm as the length of a vector. It is a function that maps a vector to a positive value. Different functions can be used and we will see few examples.
# 2.5 Norms
#### Definition
Given vectors $x$ and $y$ of length one, which are simply scalars $x$ and $y$, the most natural notion of distance between $x$ and $y$ is obtained from the absolute value. Therefore we define the distance to be $\lvert{x − y}\rvert$. We can therefore define a distance function for vectors that has similar properties.
A function $\lvert\lvert \cdot \rvert\rvert \colon \mathbb{R}^{n} \rightarrow
\mathbb{R}$ is called a vector norm if it has the following properties:
>$\lvert\lvert \bs{x} \rvert\rvert \ge 0 \text{ for any vector } \bs{x} \in \mathbb R^{n}, \text{ and } \lvert\lvert \bs{x} \rvert\rvert = 0 \text{ if and only if } \bs{x} = 0$
>$\lvert\lvert \alpha \bs{x} \rvert\rvert = \lvert \alpha \rvert \lvert\lvert \bs{x} \rvert\rvert \text{ for any vector } \bs{x} \in \mathbb R^{n} \text{ and any } \alpha \in \mathbb R$
>$\lvert\lvert \bs{x} + \bs{y} \rvert\rvert \le \lvert\lvert \bs{x} \rvert\rvert + \lvert\lvert \bs{y} \rvert\rvert \text { for any vectors } \bs{x},\bs{y} \in \mathbb R^{n}$
That last property is called the [triangle inequality](https://en.wikipedia.org/wiki/Triangle_inequality). It should be noted that when $n = 1$, the absolute value function is a vector norm.
#### A note on notation
>Norms are usually represented with two horizontal bars: $\norm{\bs{x}}$
>When we say that $x \in \mathbb R$, we mean that $x$ is a (one-dimensional) scalar that happens to be a real number. For example, we might have $x=1$ or $x=-12$.
>On the other hand, when we say that $\vec x \in \mathbb R^2$, we mean that $\vec x$ is a two-dimensional vector whose two components are both real numbers. In other words, $\vec x$ is an ordered pair in the Cartesian plane that has the form $(x_1, x_2)$, where $x_1,x_2 \in \mathbb R$. For example, we might have $\vec x = (-1, 7)$, or $\vec x = (\pi, 2.54)$.
>When we define a function $\lvert\lvert \cdot \rvert\rvert \colon \mathbb R^2 \to \mathbb R$, we mean that the function $\lvert\lvert \cdot \rvert\rvert$ maps each ordered pair (which contains two numbers as input) to a single number (as output). For example, we could define such a mapping by:
$$\lvert\lvert (x_1, x_2) \rvert\rvert = 2x_1 + 3x_2$$
>In this case, the function $\lvert\lvert \cdot \rvert\rvert$ would map vector $\vec x = (-1, 7)$ to $2(-1) + 3(7) = 19$.
# The triangle inequity
In plain English, the norm of the sum of some vectors is less than or equal to the sum of the norms of these vectors.
$$
\norm{\bs{x}+\bs{y}} \leq \norm{\bs{x}}+\norm{\bs{y}}
$$
### Example 1.
$$
\bs{x}=
\begin{bmatrix}
1 & 6
\end{bmatrix}
$$
and
$$
\bs{y}=
\begin{bmatrix}
4 & 2
\end{bmatrix}
$$
$$
\norm{\bs{x}+\bs{y}} = \sqrt{(1+4)^2+(6+2)^2} = \sqrt{89} \approx 9.43
$$
$$
\norm{\bs{x}}+\norm{\bs{y}} = \sqrt{1^2+6^2}+\sqrt{4^2+2^2} = \sqrt{37}+\sqrt{20} \approx 10.55
$$
Let's check these results:
```
x = np.array([1, 6])
x
y = np.array([4, 2])
y
x+y
np.linalg.norm(x+y)
np.linalg.norm(x)+np.linalg.norm(y)
```
Geometrically, this simply means that the shortest path between two points is a line.
```
x = [0,0,1,6]
y = [0,0,4,2]
x_bis = [1,6,y[2],y[3]]
w = [0,0,5,8]
plt.quiver([x[0], x_bis[0], w[0]],
[x[1], x_bis[1], w[1]],
[x[2], x_bis[2], w[2]],
[x[3], x_bis[3], w[3]],
angles='xy', scale_units='xy', scale=1, color=sns.color_palette())
# plt.rc('text', usetex=True)
plt.xlim(-2, 6)
plt.ylim(-2, 9)
plt.axvline(x=0, color='grey')
plt.axhline(y=0, color='grey')
plt.text(-1, 3.5, r'$||\vec{x}||$', color=sns.color_palette()[0], size=20)
plt.text(2.5, 7.5, r'$||\vec{y}||$', color=sns.color_palette()[1], size=20)
plt.text(2, 2, r'$||\vec{x}+\vec{y}||$', color=sns.color_palette()[2], size=20)
```
# P Norms
Once you get deep into machine learning algorithms, you'll eventually be faced with choosing between the $L^1$-norm or the $L^2$-norm for your model's **loss function**, which tries to minimize the difference between predicted and true values when estimating parameters.
If you want to make an informed decision about your loss functions, then you need to become familiar with the **vector norms**.
The most commonly used vector norms belong to the family of $p$-norms, or $\ell_{p}$-norms, which are defined by:
$$\left\| x \right\| _p = \left( |x_1|^p + |x_2|^p + \dotsb + |x_n|^p \right) ^{1/p}$$
which can be more concisely expressed with the formula:
$$\norm{\bs{x}}_p=(\sum_{i}^{n}|\bs{x}_i|^p)^{1/p}$$
In plain English, this is how you calculate the $p$-norm of a vector:
1. Calculate the absolute value of each element
2. Raise each of those absolute values to the power $p$
3. Sum all of these powered absolute values
4. Raise this result to the power $\frac{1}{p}$
Let's make this a bit more concrete by looking at some widely used $p$-norms.
# The $L^0$ norm
Raise any positive number to the $0^{th}$ power and you get $1$. And since $0^{0}$ is undefined, the $L^0$ norm gives the number of non-zero elements within a vector.
>Technically speaking, the $L^0$ norm isn't really a norm. If you look at calculation #4 above, you'll see that when $p=0$, you can't perform $\frac{1}{p}$ since you can't divide by $0$. The real reason for the name $L^0$ is that it is the limit as $p\rightarrow0$ of the $L^p$ norm:
$$\norm{\bs{x}}_0 = \lim_{p\rightarrow0} \sum^{n}_{k-1}|\bs{x}_k|^p$$
In short, The $L^0$ norm is the number of non-zero elements in a vector.
# The $L^1$ norm
When $p=1$, you have the $L^1$ norm, which is simply the sum of the absolute values:
$$
\norm{\bs{x}}_1=\sum_{i} |\bs{x}_i|
$$
The $L^1$-norm is also known as [least absolute deviations (LAD)](https://en.wikipedia.org/wiki/Least_absolute_deviations). This norm is used to minimize the sum of the absolute differences $S$ between the target value $Y_i$ and the estimated values $f(x_i)$:
$$S = \sum^{n}_{i=1}|y_i-f(x_i)|$$
# The Euclidean norm ($L^2$ norm)
When $p=2$, you have what's called the Euclidean norm. Why is it called the Euclidean norm? Because the $L^2$ norm gives the distance from the origin to the point $X$, a consequence of the Pythagorean theorem.
$$
\norm{\bs{x}}_2=(\sum_i \bs{x}_i^2)^{1/2}\Leftrightarrow \sqrt{\sum_i \bs{x}_i^2}
$$
The $L^2$-norm is also known as least squares and "SRSS", which is an acronym for the square root of the sum of squares.
In machine learning, it is used to minimuze the sum of the square of the differences $S$ between the target value $Y_i$ and the estimated values $f(x_i)$:
$$S = \sum^{n}_{i=1}(y_i-f(x_i))^2$$
Let's see an example of this norm:
### Example 2.
Graphically, the Euclidean norm corresponds to the length of the vector from the origin to the point obtained by linear combination (like applying Pythagorean theorem).
$$
\bs{u}=
\begin{bmatrix}
3 \\\\
4
\end{bmatrix}
$$
$$
\begin{align*}
\norm{\bs{u}}_2 &=\sqrt{|3|^2+|4|^2}\\\\
&=\sqrt{25}\\\\
&=5
\end{align*}
$$
So the $L^2$ norm is $5$.
The $L^2$ norm can be calculated with the `linalg.norm` function from numpy. We can check the result:
```
np.linalg.norm([3, 4])
```
Here is the graphical representation of the vectors:
```
u = [0,0,3,4]
plt.quiver([u[0]],
[u[1]],
[u[2]],
[u[3]],
angles='xy', scale_units='xy', scale=1)
plt.xlim(-2, 4)
plt.ylim(-2, 5)
plt.axvline(x=0, color='grey')
plt.axhline(y=0, color='grey')
plt.annotate('', xy = (3.2, 0), xytext = (3.2, 4),
arrowprops=dict(edgecolor='black', arrowstyle = '<->'))
plt.annotate('', xy = (0, -0.2), xytext = (3, -0.2),
arrowprops=dict(edgecolor='black', arrowstyle = '<->'))
plt.text(1, 2.5, r'$\vec{u}$', size=18)
plt.text(3.3, 2, r'$\vec{u}_y$', size=18)
plt.text(1.5, -1, r'$\vec{u}_x$', size=18)
```
In this case, the vector is in a 2-dimensional space, so it's easy to visualize.
But you can also apply this norm to higher dimensional spaces.
$$
u=
\begin{bmatrix}
u_1\\\\
u_2\\\\
\cdots \\\\
u_n
\end{bmatrix}
$$
$$
||u||_2 = \sqrt{u_1^2+u_2^2+\cdots+u_n^2}
$$
# The squared Euclidean norm (squared $L^2$ norm)
$$
\sum_i|\bs{x}_i|^2
$$
The squared $L^2$ norm is convenient because it removes the square root and we end up with the simple sum of every squared values of the vector.
The squared Euclidean norm is widely used in machine learning partly because it can be calculated with the vector operation $\bs{x}^\text{T}\bs{x}$. Operations like this yield performance gains: see [here](https://softwareengineering.stackexchange.com/questions/312445/why-does-expressing-calculations-as-matrix-multiplications-make-them-faster) and [here](https://www.quora.com/What-makes-vector-operations-faster-than-for-loops) for more details.
### Example 3.
$$
\bs{x}=
\begin{bmatrix}
2 \\\\
5 \\\\
3 \\\\
3
\end{bmatrix}
$$
$$
\bs{x}^\text{T}=
\begin{bmatrix}
2 & 5 & 3 & 3
\end{bmatrix}
$$
$$
\begin{align*}
\bs{x}^\text{T}\bs{x}&=
\begin{bmatrix}
2 & 5 & 3 & 3
\end{bmatrix} \times
\begin{bmatrix}
2 \\\\
5 \\\\
3 \\\\
3
\end{bmatrix}\\\\
&= 2\times 2 + 5\times 5 + 3\times 3 + 3\times 3= 47
\end{align*}
$$
```
x = np.array([[2], [5], [3], [3]])
x
euclideanNorm = x.T.dot(x)
euclideanNorm
np.linalg.norm(x)**2
```
They're the same!
## Derivative of the squared $L^2$ norm
Another advantage of the squared $L^2$ norm is that its partial derivative is easily computed:
$$
u=
\begin{bmatrix}
u_1\\\\
u_2\\\\
\cdots \\\\
u_n
\end{bmatrix}
$$
$$
\norm{u}_2 = u_1^2+u_2^2+\cdots+u_n^2
$$
$$
\begin{cases}
\dfrac{d\norm{u}_2}{du_1} = 2u_1\\\\
\dfrac{d\norm{u}_2}{du_2} = 2u_2\\\\
\cdots\\\\
\dfrac{d\norm{u}_2}{du_n} = 2u_n
\end{cases}
$$
## Derivative of the $L^2$ norm
In the case of the $L^2$ norm, the derivative is more complicated and takes every elements of the vector into account:
$$
\norm{u}_2 = \sqrt{(u_1^2+u_2^2+\cdots+u_n^2)} = (u_1^2+u_2^2+\cdots+u_n^2)^{\frac{1}{2}}
$$
$$
\begin{align*}
\dfrac{d\norm{u}_2}{du_1} &=
\dfrac{1}{2}(u_1^2+u_2^2+\cdots+u_n^2)^{\frac{1}{2}-1}\cdot
\dfrac{d}{du_1}(u_1^2+u_2^2+\cdots+u_n^2)\\\\
&=\dfrac{1}{2}(u_1^2+u_2^2+\cdots+u_n^2)^{-\frac{1}{2}}\cdot
\dfrac{d}{du_1}(u_1^2+u_2^2+\cdots+u_n^2)\\\\
&=\dfrac{1}{2}\cdot\dfrac{1}{(u_1^2+u_2^2+\cdots+u_n^2)^{\frac{1}{2}}}\cdot
\dfrac{d}{du_1}(u_1^2+u_2^2+\cdots+u_n^2)\\\\
&=\dfrac{1}{2}\cdot\dfrac{1}{(u_1^2+u_2^2+\cdots+u_n^2)^{\frac{1}{2}}}\cdot
2\cdot u_1\\\\
&=\dfrac{u_1}{\sqrt{(u_1^2+u_2^2+\cdots+u_n^2)}}\\\\
\end{align*}
$$
$$
\begin{cases}
\dfrac{d\norm{u}_2}{du_1} = \dfrac{u_1}{\sqrt{(u_1^2+u_2^2+\cdots+u_n^2)}}\\\\
\dfrac{d\norm{u}_2}{du_2} = \dfrac{u_2}{\sqrt{(u_1^2+u_2^2+\cdots+u_n^2)}}\\\\
\cdots\\\\
\dfrac{d\norm{u}_2}{du_n} = \dfrac{u_n}{\sqrt{(u_1^2+u_2^2+\cdots+u_n^2)}}\\\\
\end{cases}
$$
>### Why is it important to easily and quickly calculate these partial derivatives?
> calculus, you use derivatives to understand how functions change over time. In machine learning, you can apply this concept to find models with parameters that minimize prediction error.
>#### Partial derivatives
>In functions with 2 or more variables, the partial derivative is the derivative of one variable with respect to the others. If we change $x$, but hold all other variables constant, how does $f(x,z)$ change? That’s one partial derivative. The next variable is $z$. If we change $z$ but hold $x$ constant, how does $f(x,z)$ change? Once calculate all of the partial derivatices, you store them in something called a gradient, which represents the full derivative of the multivariable function.
>#### Gradients
>A gradient is a vector that stores the partial derivatives of multivariable functions. It helps us calculate the slope at a specific point on a curve for functions with multiple independent variables. In order to calculate this more complex slope, we need to isolate each variable to determine how it impacts the output on its own. To do this we iterate through each of the variables and calculate the derivative of the function after holding all other variables constant. Each iteration produces a partial derivative which we store in the gradient.
>#### Gradient Descent
>Gradient descent is the iterative optimization algorithm for finding the minimum of a function. In machine learning, this means we're finding a model that minimizes the error of our predictions.
One problem of the squared $L^2$ norm is that it hardly discriminates between 0 and small values because the increase of the function is slow.
We can see this by graphically comparing the squared $L^2$ norm with the $L^2$ norm. The $z$-axis corresponds to the norm and the $x$- and $y$-axis correspond to two parameters. The same thing is true with more than 2 dimensions but it would be hard to visualize that!
$L^2$ norm:
<img src="images/l2-norm.png" width="500" alt="Representation of the L2 norm" title="The L2 norm">
<em>The L2 norm</em>
Squared $L^2$ norm:
<img src="images/squared-l2-norm.png" width="500" alt="Representation of the squared L2 norm" title="The squared L2 norm">
<em>The squared L2 norm</em>
$L^1$ norm:
<img src="images/L1-norm.png" alt="Representation of the L1 norm" title="The L1 norm" width="500">
<em>The L1 norm</em>
These plots were made with the help of this [website](https://academo.org/demos/3d-surface-plotter/).
# The max norm
The $L^\infty$ norm corresponds to the absolute value of the greatest element of the vector.
$$
\norm{\bs{x}}_\infty = \max\limits_i|x_i|
$$
The length of a vector can be calculated using the max norm.
Max norm is also used as a regularization in machine learning, such as on neural network weights, called max norm regularization.
```
a = array([1, 2, 3,10,99])
a
maxnorm = norm(a, inf)
print(maxnorm)
```
# Matrix norms: the Frobenius norm
The Frobenius norm is equivalent to taking the $L^2$ norm of a matrix after flattening it.
$$
\norm{\bs{A}}_F=\sqrt{\sum_{i,j}A^2_{i,j}}
$$
>#### Regularization
This norm is most often used as for regularization, which is the process of introducing additional information in order to solve an [ill-posed problem](https://en.wikipedia.org/wiki/Well-posed_problem) or to prevent [overfitting](https://en.wikipedia.org/wiki/Overfitting). One particular use of regularization is in the field of classification. Empirical learning of classifiers (learning from a finite data set) is always an underdetermined problem, because in general we are trying to infer a function of any $x$ given only some examples $x_{1},x_{2},\dots x_{n}$. Thus, in classification problems, a regularization term (or regularizer) $R(f)$ can be added to a loss function.
Let's see how we can peform this operation with numpy:
```
A = np.array([[1, 2], [6, 4], [3, 2]])
A
np.linalg.norm(A)
```
# Finding the angle between two vectors.
Suppose we are interested in finding the angle between two given vectors, $\bs{x}$ and $\bs{y}$.
Suppose we have two $n$-dimensional vectors $\bs{x}$ and $\bs{y}$ as shown below:
$$x=\left( \begin{array}{c} x_1 \\ x_2 \\ x_3 \\ \vdots \\x_n \end{array} \right) \textrm{ and } y=\left( \begin{array}{c} y_1 \\ y_2 \\ y_3 \\ \vdots \\ y_n \end{array} \right)$$
In a previous lesson, we defined the dot product like this:
$$x \cdot y = x_1 y_1 + x_2 y_2 + x_3 y_3 + \cdots + x_n y_n$$
We also noted that the dot product is commutative:
$$\begin{array}{rcl}
x \cdot y & = & x_1 y_1 + x_2 y_2 \cdots + x_n y_n \\
& = & y_1 x_1 + y_2 x_2 \cdots + y_n x_n \\
& = & y \cdot x
\end{array}$$
We also showed that the dot product distributes over vector sums:
$$\begin{array}{rcl}
x \cdot (y + z) & = & x_1 (y_1 + z_1) + x_2 (y_2 + z_2) + \cdots x_n (y_n + z_n)\\
& = & x_1 y_1 + x_1 z_1 + x_2 y_2 + x_2 z_2 + \cdots + x_n y_n + x_n z_n\\
& = & (x_1 y_1 + x_2 y_2 \cdots + x_n y_n) + (x_1 z_1 + x_2 z_2 \cdots + x_n z_n)\\
& = & x \cdot y + x \cdot z
\end{array}$$
We also noted that for any scalar $k$, we have:
$$\begin{array}{rcl}
(kx) \cdot y & = & kx_1 y_1 + kx_2 y_2 + \cdots + kx_n y_n\\
& = & k(x_1 y_1 + x_2 y_2 \cdots + x_n y_n)\\
& = & k(x \cdot y)
\end{array}$$
We will use these three properties in our calculation below.
But first, recall how we defined $L^2$-norm of a vector above:
$$||x||_2 = \sqrt{x_1^2 + x_2^2 + \cdots + x_n^2}$$
The $L^2$-norm is meant to be geometrically interpreted as the length of the vector, or equivalently, the distance between the points $(0,0,...,0)$ and $(x_1,x_2,...,x_n)$.
Interestingly, note how the $L^2$-norm can be written in a much shorter way by invoking the dot product:
$$||x||_2 = \sqrt{x \cdot x}$$
Now, armed with the ideas of the dot product and the norm of a vector, suppose we are interested in finding the angle between two given vectors, $\bs{x}$ and $\bs{y}$.
Although you probably don't remember the [Law of Cosines](https://en.wikipedia.org/wiki/Law_of_cosines) from your high school geometry class, we're going to use it since it is a generalization on the Pythagorean Theorem and gives us the relationship between the side lengths of an arbitrary triangle. Specifically, if a triangle has side lengths $a$, $b$, and $c$, then:
$$a^2 + b^2 - 2ab\cos \theta = c^2$$
where $\theta$ is the angle between the sides of length $a$ and $b$.
As an example, consider the triangle that can be formed from the vectors $\bs{x}$, $\bs{y}$, and $\bs{x−y}$:
<img src="images/triangle.png" height=250 width =250 img>
Applying the Law of Cosines to this triangle, we have:
$$||x||_2 +||y||_2 - 2||x||_2 \, ||y||_2\cos \theta = ||x-y||_2$$
But this implies, using our observations about the dot product made above, that:
$$\begin{array}{rcl}
(x \cdot x) + (y \cdot y) - 2||x||_2 \, ||y||_2\cos \theta & = & (x-y) \cdot (x-y)\\
& = & x \cdot (x-y) - y \cdot (x-y)\\
& = & (x \cdot x) - (x \cdot y) - (y \cdot x) + (y \cdot y)\\
& = & (x \cdot x) - (x \cdot y) - (x \cdot y) + (y \cdot y)\\
& = & (x \cdot x) - 2(x \cdot y) + (y \cdot y)\\
\end{array}$$
Subtracting the common $(\bs{x}\cdot\bs{x})$ and $(\bs{y}\cdot\bs{y})$ from both sides, we find:
$$- 2||x||_2 \, ||y||_2\cos \theta = - 2(x \cdot y)$$
Solving for $\cos \theta$ tells us:
$$\cos \theta = \frac{x \cdot y}{||x||_2 \, ||y||_2}$$
Rearranging, we get:
$$x \cdot y = \norm{\bs{x}}_2\norm{\bs{y}}_2\cos\theta$$
Finally, the dot product can be expressed as
$$x \cdot y = x^\text{T}y$$
And if we substitute in $x^\text{T}y$, we get:
$$
x^\text{T}y = \norm{x}_2\cdot\norm{y}_2\cos\theta
$$
Which shows how the dot product can be expressed as the product of $L^2$ norms and $\cos\theta$.
### Example 4.
That was a whole bunch of math, so let's try to visualize this with a simple example.
$$
\bs{x}=
\begin{bmatrix}
0 \\\\
2
\end{bmatrix}
$$
and
$$
\bs{y}=
\begin{bmatrix}
2 \\\\
2
\end{bmatrix}
$$
```
x = [0,0,0,2]
y = [0,0,2,2]
plt.xlim(-2, 4)
plt.ylim(-2, 5)
plt.axvline(x=0, color='grey', zorder=0)
plt.axhline(y=0, color='grey', zorder=0)
plt.quiver([x[0], y[0]],
[x[1], y[1]],
[x[2], y[2]],
[x[3], y[3]],
angles='xy', scale_units='xy', scale=1)
plt.text(-0.5, 1, r'$\vec{x}$', size=18)
plt.text(1.5, 0.5, r'$\vec{y}$', size=18)
plt.show()
plt.close()
```
We took this example for its simplicity. As we can see, the angle $\theta$ is equal to 45°.
$$
\bs{x^\text{T}y}=
\begin{bmatrix}
0 & 2
\end{bmatrix} \cdot
\begin{bmatrix}
2 \\\\
2
\end{bmatrix} =
0\times2+2\times2 = 4
$$
and
$$
\norm{\bs{x}}_2=\sqrt{0^2+2^2}=\sqrt{4}=2
$$
$$
\norm{\bs{y}}_2=\sqrt{2^2+2^2}=\sqrt{8}
$$
$$
2\times\sqrt{8}\times cos(45)=4
$$
Here are the operations using numpy:
```
# Note: np.cos take the angle in radian
np.cos(np.deg2rad(45))*2*np.sqrt(8)
```
# References
The proof for the dot products, norms, and angles between vectors came from [here](http://www.oxfordmathcenter.com/drupal7/node/168).
This lesson requried a lot of mathematical notation. A nice cheat sheet for these symbols can be found [here](https://artofproblemsolving.com/wiki/index.php/LaTeX:Symbols#Dots).
| github_jupyter |

[](https://colab.research.google.com/github/JohnSnowLabs/spark-nlp-workshop/blob/master/tutorials/streamlit_notebooks/healthcare/NER_ANATOMY.ipynb)
# **Detect anatomical references**
To run this yourself, you will need to upload your license keys to the notebook. Just Run The Cell Below in order to do that. Also You can open the file explorer on the left side of the screen and upload `license_keys.json` to the folder that opens.
Otherwise, you can look at the example outputs at the bottom of the notebook.
## 1. Colab Setup
Import license keys
```
import os
import json
from google.colab import files
license_keys = files.upload()
with open(list(license_keys.keys())[0]) as f:
license_keys = json.load(f)
sparknlp_version = license_keys["PUBLIC_VERSION"]
jsl_version = license_keys["JSL_VERSION"]
print ('SparkNLP Version:', sparknlp_version)
print ('SparkNLP-JSL Version:', jsl_version)
```
Install dependencies
```
%%capture
for k,v in license_keys.items():
%set_env $k=$v
!wget https://raw.githubusercontent.com/JohnSnowLabs/spark-nlp-workshop/master/jsl_colab_setup.sh
!bash jsl_colab_setup.sh
# Install Spark NLP Display for visualization
!pip install --ignore-installed spark-nlp-display
```
Import dependencies into Python and start the Spark session
```
import pandas as pd
from pyspark.ml import Pipeline
from pyspark.sql import SparkSession
import pyspark.sql.functions as F
import sparknlp
from sparknlp.annotator import *
from sparknlp_jsl.annotator import *
from sparknlp.base import *
import sparknlp_jsl
spark = sparknlp_jsl.start(license_keys['SECRET'])
# manually start session
'''
builder = SparkSession.builder \
.appName('Spark NLP Licensed') \
.master('local[*]') \
.config('spark.driver.memory', '16G') \
.config('spark.serializer', 'org.apache.spark.serializer.KryoSerializer') \
.config('spark.kryoserializer.buffer.max', '2000M') \
.config('spark.jars.packages', 'com.johnsnowlabs.nlp:spark-nlp_2.11:' +sparknlp.version()) \
.config('spark.jars', f'https://pypi.johnsnowlabs.com/{secret}/spark-nlp-jsl-{jsl_version}.jar')
'''
```
## 2. Select the NER model and construct the pipeline
Select the NER model - Anatomy models: **ner_anatomy**
For more details: https://github.com/JohnSnowLabs/spark-nlp-models#pretrained-models---spark-nlp-for-healthcare
```
# Change this to the model you want to use and re-run the cells below.
# Anatomy models: ner_anatomy
MODEL_NAME = "ner_anatomy"
```
Create the pipeline
```
document_assembler = DocumentAssembler() \
.setInputCol('text')\
.setOutputCol('document')
sentence_detector = SentenceDetector() \
.setInputCols(['document'])\
.setOutputCol('sentence')
tokenizer = Tokenizer()\
.setInputCols(['sentence']) \
.setOutputCol('token')
word_embeddings = WordEmbeddingsModel.pretrained('embeddings_clinical', 'en', 'clinical/models') \
.setInputCols(['sentence', 'token']) \
.setOutputCol('embeddings')
clinical_ner = MedicalNerModel.pretrained(MODEL_NAME, "en", "clinical/models") \
.setInputCols(["sentence", "token", "embeddings"])\
.setOutputCol("ner")
ner_converter = NerConverter()\
.setInputCols(['sentence', 'token', 'ner']) \
.setOutputCol('ner_chunk')
nlp_pipeline = Pipeline(stages=[
document_assembler,
sentence_detector,
tokenizer,
word_embeddings,
clinical_ner,
ner_converter])
```
## 3. Create example inputs
```
# Enter examples as strings in this array
input_list = [
"""This is an 11-year-old female who comes in for two different things. 1. She was seen by the allergist. No allergies present, so she stopped her Allegra, but she is still real congested and does a lot of snorting. They do not notice a lot of snoring at night though, but she seems to be always like that. 2. On her right great toe, she has got some redness and erythema. Her skin is kind of peeling a little bit, but it has been like that for about a week and a half now.
General: Well-developed female, in no acute distress, afebrile.
HEENT: Sclerae and conjunctivae clear. Extraocular muscles intact. TMs clear. Nares patent. A little bit of swelling of the turbinates on the left. Oropharynx is essentially clear. Mucous membranes are moist.
Neck: No lymphadenopathy.
Chest: Clear.
Abdomen: Positive bowel sounds and soft.
Dermatologic: She has got redness along the lateral portion of her right great toe, but no bleeding or oozing. Some dryness of her skin. Her toenails themselves are very short and even on her left foot and her left great toe the toenails are very short."""
]
```
## 4. Use the pipeline to create outputs
```
empty_df = spark.createDataFrame([['']]).toDF('text')
pipeline_model = nlp_pipeline.fit(empty_df)
df = spark.createDataFrame(pd.DataFrame({'text': input_list}))
result = pipeline_model.transform(df)
```
## 5. Visualize results
```
from sparknlp_display import NerVisualizer
NerVisualizer().display(
result = result.collect()[0],
label_col = 'ner_chunk',
document_col = 'document'
)
```
| github_jupyter |
# **Introduction to Competitive Programming**
---
Date and Time: 8th July 2019 Monday 5-7pm
Venue: Matthews Bldg RM232
Handlers: Payton Yao (Canva), Kathrina Ondap (Google)
Coordinator: Luke Sy
Repository: https://github.com/ieeeunswsb/cpworkshop
```
print("Welcome to IEEE UNSW student branch's introduction to competitive programming!")
print("We'll be having Payton and Kathy (and to some extent Luke) to guide us through this workshop!")
from urllib.request import urlretrieve
import os
def download(url, file):
if not os.path.isfile(file):
print("Download file... " + file + " ...")
urlretrieve(url,file)
print("File downloaded")
Repo = 'https://raw.githubusercontent.com/ieeeunswsb/cpworkshop/master/'
Files = ['sample_data/s01-foregone.txt',
'sample_data/s01-maxpathsum1.txt',
'sample_data/s01-maxpathsum2.txt',
'sample_data/s01-gorosort.txt',
'sample_data/s01-mutual-friend-zone.txt',
'sample_data/s01-cryptopangrams.txt']
for i in Files:
download(Repo+i, i)
```
## Problem 01: Foregone Solution (GCJ2019 Qualification)
Link: https://codingcompetitions.withgoogle.com/codejam/round/0000000000051705/0000000000088231?fbclid=IwAR2hq-J3PzLTiDXzKInwkT8CdnhJZjpdAnWH70qcojFxugkDm4HlguJRtQs
```
import sys
# comment the first line (read file) and uncomment the 2nd line (read stdin)
# when submitting your answer
with open('sample_data/s01-foregone.txt', 'r') as f:
# with sys.stdin as f:
T = int(f.readline())
# put your code here
print("Answer")
```
## Problem 02: Max Sum Path I and II (Project Euler 18 and 67)
Link1: https://projecteuler.net/problem=18
Link2: https://projecteuler.net/problem=67
```
import sys
with open('sample_data/s01-maxpathsum2.txt', 'r') as f:
# N = 15 # max sum path I
N = 100 # max sum path II
# put your code here
print("Answer")
```
## Problem 03: Gorosort (GCJ 2011 Qualification)
Link: https://code.google.com/codejam/contest/dashboard?c=975485&fbclid=IwAR1nckjG1Wpmddyb0xb1tUaeLi9hnOgVq-uY-J9P4dL-Cg9QlUUZxEmV5S0#s=p3
```
import sys
# comment the first line (read file) and uncomment the 2nd line (read stdin)
# when submitting your answer
with open('sample_data/s01-gorosort.txt', 'r') as f:
# with sys.stdin as f:
T = int(f.readline())
# put your code here
print("Answer")
```
## Problem 04: Mutual Friend Zone
Link: https://www.hackerrank.com/contests/noi-ph-practice-page/challenges/mutual-friendzone-not-hacked?fbclid=IwAR0Q8J0CzHcDeAfGpENaQZBjOsMvpLwbiFlT5jCajKImhqxsBkV4uH-qxQ0
```
import sys
with open('sample_data/s01-mutual-friend-zone.txt') as f:
# with sys.stdin as f:
T = int(f.readline())
print(T)
# put your code here
print("Answer")
```
## (EXTRA) Problem 05: Cryptopangrams
In case you find problem 1-4 TOO easy
Link: https://codingcompetitions.withgoogle.com/codejam/round/0000000000051705/000000000008830b?fbclid=IwAR3dyzboFg7sMafZ5uvGulwGLL7ow00_zr9l3z9jQcDickWXkzi_8v66wyE
```
import sys
from math import gcd
with open('sample_data/s01-cryptopangrams.txt') as f:
# with sys.stdin as f:
T = int(f.readline())
# put your code here
print("Answer")
```
In case you find problem 1-5 TOO easy, call the attention of Payton or Luke in the workshop.
```
```
| github_jupyter |
# Lesson 2
In the screencast for this lesson I go through a few scenarios for time series. This notebook contains the code for that with a few little extras! :)
# Setup
```
!pip install -U tf-nightly-2.0-preview
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
from tensorflow import keras
def plot_series(time, series, format="-", start=0, end=None, label=None):
plt.plot(time[start:end], series[start:end], format, label=label)
plt.xlabel("Time")
plt.ylabel("Value")
if label:
plt.legend(fontsize=14)
plt.grid(True)
```
# Trend and Seasonality
```
def trend(time, slope=0):
return slope * time
```
Let's create a time series that just trends upward:
```
time = np.arange(4 * 365 + 1)
baseline = 10
series = trend(time, 0.1)
plt.figure(figsize=(10, 6))
plot_series(time, series)
plt.show()
```
Now let's generate a time series with a seasonal pattern:
```
def seasonal_pattern(season_time):
"""Just an arbitrary pattern, you can change it if you wish"""
return np.where(season_time < 0.4,
np.cos(season_time * 2 * np.pi),
1 / np.exp(3 * season_time))
def seasonality(time, period, amplitude=1, phase=0):
"""Repeats the same pattern at each period"""
season_time = ((time + phase) % period) / period
return amplitude * seasonal_pattern(season_time)
baseline = 10
amplitude = 40
series = seasonality(time, period=365, amplitude=amplitude)
plt.figure(figsize=(10, 6))
plot_series(time, series)
plt.show()
```
Now let's create a time series with both trend and seasonality:
```
slope = 0.05
series = baseline + trend(time, slope) + seasonality(time, period=365, amplitude=amplitude)
plt.figure(figsize=(10, 6))
plot_series(time, series)
plt.show()
```
# Noise
In practice few real-life time series have such a smooth signal. They usually have some noise, and the signal-to-noise ratio can sometimes be very low. Let's generate some white noise:
```
def white_noise(time, noise_level=1, seed=None):
rnd = np.random.RandomState(seed)
return rnd.randn(len(time)) * noise_level
noise_level = 5
noise = white_noise(time, noise_level, seed=42)
plt.figure(figsize=(10, 6))
plot_series(time, noise)
plt.show()
```
Now let's add this white noise to the time series:
```
series += noise
plt.figure(figsize=(10, 6))
plot_series(time, series)
plt.show()
```
All right, this looks realistic enough for now. Let's try to forecast it. We will split it into two periods: the training period and the validation period (in many cases, you would also want to have a test period). The split will be at time step 1000.
```
split_time = 1000
time_train = time[:split_time]
x_train = series[:split_time]
time_valid = time[split_time:]
x_valid = series[split_time:]
def autocorrelation(time, amplitude, seed=None):
rnd = np.random.RandomState(seed)
φ1 = 0.5
φ2 = -0.1
ar = rnd.randn(len(time) + 50)
ar[:50] = 100
for step in range(50, len(time) + 50):
ar[step] += φ1 * ar[step - 50]
ar[step] += φ2 * ar[step - 33]
return ar[50:] * amplitude
def autocorrelation(time, amplitude, seed=None):
rnd = np.random.RandomState(seed)
φ = 0.8
ar = rnd.randn(len(time) + 1)
for step in range(1, len(time) + 1):
ar[step] += φ * ar[step - 1]
return ar[1:] * amplitude
series = autocorrelation(time, 10, seed=42)
plot_series(time[:200], series[:200])
plt.show()
series = autocorrelation(time, 10, seed=42) + trend(time, 2)
plot_series(time[:200], series[:200])
plt.show()
series = autocorrelation(time, 10, seed=42) + seasonality(time, period=50, amplitude=150) + trend(time, 2)
plot_series(time[:200], series[:200])
plt.show()
series = autocorrelation(time, 10, seed=42) + seasonality(time, period=50, amplitude=150) + trend(time, 2)
series2 = autocorrelation(time, 5, seed=42) + seasonality(time, period=50, amplitude=2) + trend(time, -1) + 550
series[200:] = series2[200:]
#series += noise(time, 30)
plot_series(time[:300], series[:300])
plt.show()
def impulses(time, num_impulses, amplitude=1, seed=None):
rnd = np.random.RandomState(seed)
impulse_indices = rnd.randint(len(time), size=10)
series = np.zeros(len(time))
for index in impulse_indices:
series[index] += rnd.rand() * amplitude
return series
series = impulses(time, 10, seed=42)
plot_series(time, series)
plt.show()
def autocorrelation(source, φs):
ar = source.copy()
max_lag = len(φs)
for step, value in enumerate(source):
for lag, φ in φs.items():
if step - lag > 0:
ar[step] += φ * ar[step - lag]
return ar
signal = impulses(time, 10, seed=42)
series = autocorrelation(signal, {1: 0.99})
plot_series(time, series)
plt.plot(time, signal, "k-")
plt.show()
signal = impulses(time, 10, seed=42)
series = autocorrelation(signal, {1: 0.70, 50: 0.2})
plot_series(time, series)
plt.plot(time, signal, "k-")
plt.show()
series_diff1 = series[1:] - series[:-1]
plot_series(time[1:], series_diff1)
from pandas.plotting import autocorrelation_plot
autocorrelation_plot(series)
from statsmodels.tsa.arima_model import ARIMA
model = ARIMA(series, order=(5, 1, 0))
model_fit = model.fit(disp=0)
print(model_fit.summary())
df = pd.read_csv("sunspots.csv", parse_dates=["Date"], index_col="Date")
series = df["Monthly Mean Total Sunspot Number"].asfreq("1M")
series.head()
series.plot(figsize=(12, 5))
series["1995-01-01":].plot()
series.diff(1).plot()
plt.axis([0, 100, -50, 50])
from pandas.plotting import autocorrelation_plot
autocorrelation_plot(series)
autocorrelation_plot(series.diff(1)[1:])
autocorrelation_plot(series.diff(1)[1:].diff(11 * 12)[11*12+1:])
plt.axis([0, 500, -0.1, 0.1])
autocorrelation_plot(series.diff(1)[1:])
plt.axis([0, 50, -0.1, 0.1])
116.7 - 104.3
[series.autocorr(lag) for lag in range(1, 50)]
pd.read_csv(filepath_or_buffer, sep=',', delimiter=None, header='infer', names=None, index_col=None, usecols=None, squeeze=False, prefix=None, mangle_dupe_cols=True, dtype=None, engine=None, converters=None, true_values=None, false_values=None, skipinitialspace=False, skiprows=None, skipfooter=0, nrows=None, na_values=None, keep_default_na=True, na_filter=True, verbose=False, skip_blank_lines=True, parse_dates=False, infer_datetime_format=False, keep_date_col=False, date_parser=None, dayfirst=False, iterator=False, chunksize=None, compression='infer', thousands=None, decimal=b'.', lineterminator=None, quotechar='"', quoting=0, doublequote=True, escapechar=None, comment=None, encoding=None, dialect=None, tupleize_cols=None, error_bad_lines=True, warn_bad_lines=True, delim_whitespace=False, low_memory=True, memory_map=False, float_precision=None)
Read a comma-separated values (csv) file into DataFrame.
from pandas.plotting import autocorrelation_plot
series_diff = series
for lag in range(50):
series_diff = series_diff[1:] - series_diff[:-1]
autocorrelation_plot(series_diff)
import pandas as pd
series_diff1 = pd.Series(series[1:] - series[:-1])
autocorrs = [series_diff1.autocorr(lag) for lag in range(1, 60)]
plt.plot(autocorrs)
plt.show()
```
| github_jupyter |
```
```

Cross-validation is a technique in which we train our model using the subset of the data-set and then evaluate using the complementary subset of the data-set.
steps:
1. You reserve a sample data set.
2. Train the model using the remaining part of the dataset.
3. Use the reserve sample of the test (validation) set. This will help you in gauging the effectiveness of your model’s performance. If your model delivers a positive result on validation data, go ahead with the current model
**Holdout Validation Approach - Train and Test Set Split**
we perform training on the 50% of the given data-set and rest 50% is used for the testing purpose. The major drawback of this method is that we perform training on the 50% of the dataset, it may possible that the remaining 50% of the data contains some important information which we are leaving while training our model i.e higher bias
```
from sklearn.datasets import load_iris
from sklearn.model_selection import train_test_split
from sklearn.neighbors import KNeighborsClassifier
from sklearn import metrics
import pandas as pd
data = load_iris()
df = pd.DataFrame(data.data, columns=data.feature_names)
df['target'] = pd.Series(data.target)
df.head()
X = df.iloc[:,:-1]
Y = df.iloc[:,-1]
# use train/test split with different random_state values
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=6)
train.shape, validation.shape
# check classification accuracy of KNN with K=5
knn = KNeighborsClassifier(n_neighbors=5)
knn.fit(X_train, y_train)
y_pred = knn.predict(X_test)
metrics.accuracy_score(y_test, y_pred)
```
**Leave one out cross validation (LOOCV)**
In this approach, we reserve only one data point from the available dataset, and train the model on the rest of the data. This process iterates for each data point.
* We make use of all data points, hence the bias will be low.
* We repeat the cross validation process n times (where n is number of data points) which results in a higher execution time
* This approach leads to higher variation in testing model effectiveness because we test against one data point
```
# Import necessary modules
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_squared_error
from math import sqrt
from sklearn import model_selection
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import KFold
from sklearn.model_selection import LeaveOneOut
from sklearn.model_selection import LeavePOut
from sklearn.model_selection import ShuffleSplit
from sklearn.model_selection import StratifiedKFold
loocv = LeaveOneOut()
loocv.get_n_splits(X)
# printing the training and validation data
#for train_index, test_index in loocv.split(X):
#print(train_index,test_index)
knn_loocv = KNeighborsClassifier(n_neighbors=5)
results_loocv = model_selection.cross_val_score(knn_loocv, X, Y, cv=loocv)
print("Accuracy: %.2f%%" % (results_loocv.mean()*100.0))
```
**K-fold cross validation**
1. Randomly split your entire dataset into k”folds”
2. For each k-fold in your dataset, build your model on k – 1 folds of the dataset. Then, test the model to check the effectiveness for kth fold
Record the error you see on each of the predictions
3. Repeat this until each of the k-folds has served as the test set
4. The average of your k recorded errors is called the cross-validation error and will serve as your performance metric for the model.
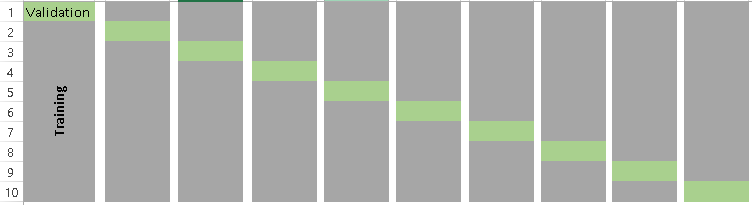

```
#defining number of folds for model
from sklearn.model_selection import KFold
kf = KFold(n_splits=5, shuffle=False)
kf
for fold, (train_idx, val_idx) in enumerate(kf.split(X,Y)):
print(len(train_idx), len(val_idx))
model_kfold = KNeighborsClassifier(n_neighbors=5)
results_kfold = model_selection.cross_val_score(model_kfold,X,Y, cv=kf)
print("Accuracy: %.2f%%" % (results_kfold.mean()*100.0))
```
This runs K times faster than Leave One Out cross-validation because K-fold cross-validation repeats the train/test split K-times.
**Stratified k-fold cross validation**
Stratification is the process of rearranging the data so as to ensure that each fold is a good representative of the whole. For example, in a binary classification problem where each class comprises of 50% of the data, it is best to arrange the data such that in every fold, each class comprises of about half the instances.

```
from sklearn.model_selection import StratifiedKFold
skf = StratifiedKFold(n_splits=5, random_state=None)
model_skfold = KNeighborsClassifier(n_neighbors=5)
results_kfold = model_selection.cross_val_score(model_skfold,X,Y, cv=skf)
print("Accuracy: %.2f%%" % (results_kfold.mean()*100.0))
```
**RepeatedKFold**
This is where the k-fold cross-validation procedure is repeated n times, where importantly, the data sample is shuffled prior to each repetition, which results in a different split of the sample.
```
from sklearn.model_selection import RepeatedKFold
rkf = RepeatedKFold(n_splits=5, n_repeats=10, random_state=None)
model_rkfold = KNeighborsClassifier(n_neighbors=5)
results_kfold = model_selection.cross_val_score(model_rkfold,X,Y, cv=rkf)
print("Accuracy: %.2f%%" % (results_kfold.mean()*100.0))
```
**Cross Validation for time series**
Splitting a time-series dataset randomly does not work because the time section of your data will be messed up. For a time series forecasting problem, we perform cross validation in the following manner.
Folds for time series cross valdiation are created in a forward chaining fashion
Suppose we have a time series for yearly consumer demand for a product during a period of n years. The folds would be created like:
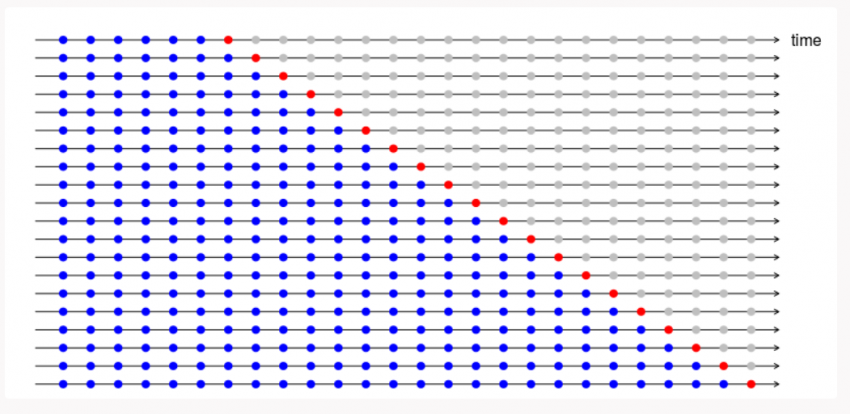
fold 1: training [1], test [2]
fold 2: training [1 2], test [3]
fold 3: training [1 2 3], test [4]
fold 4: training [1 2 3 4], test [5]
fold 5: training [1 2 3 4 5], test [6]
.
.
.
fold n: training [1 2 3 ….. n-1], test [n]
**Underfitting – High bias and low variance**
Techniques to reduce underfitting :
1. Increase model complexity
2. Increase number of features, performing feature engineering
3. Remove noise from the data.
4. Increase the number of epochs or increase the duration of training to get better resul
**Techniques to reduce overfitting :**
1. Increase training data.
2. Reduce model complexity.
3. Early stopping during the training phase (have an eye over the loss over the training period as soon as loss begins to increase stop training).
4. Ridge Regularization and Lasso Regularization
5. Use dropout for neural networks to tackle overfitting.
```
```
| github_jupyter |
[](https://www.pythonista.io)
# Álgebra lineal con *Numpy*.
El componente más poderoso de *Numpy* es su capacida de realizar operaciones con arreglos, y un caso particular de ellos son las matrices numéricas.
```
import numpy as np
```
## Producto punto entre dos matrices.
La función ```np.dot()```permite realizar las operaciones de producto punto entre dos matrices compatibles.
```
np.dot(<arreglo_1>,(<arreglo_2>)
```
**Ejemplo:**
* La siguiente celda creará al arreglo con nombre ```arreglo_1 ``` de forma ```(3, 2)```.
```
arreglo_1 = np.array([[1, 2],
[3, 4],
[5, 6]])
arreglo_1
```
* La siguiente celda creará al arreglo con nombre ```arreglo_2 ``` de forma ```(2, 4)```.
```
arreglo_2 = np.array([[11, 12, 13, 14],
[15, 16, 17, 18]])
arreglo_2
```
* La siguiente celda realizará la operación de producto punto entre ```arreglo_1``` y ```arreglo_2```, regresando una matriz de la forma ```(3, 4)```.
```
np.dot(arreglo_1, arreglo_2)
```
* El signo ```@``` es reconocido por *Numpy* como el operador de producto punto.
```
arreglo_1 @ arreglo_2
```
## Producto cruz entre dos matrices.
* La siguiente celda creará un arreglo de una dimensión, de forma ```(1,2)``` al que se le llamará ```vector_1```.
```
vector_1 = np.array([1, 2])
vector_1.shape
```
* La siguiente celda creará un arreglo de una dimensión, de forma ```(1,2)``` al que se le llamará ```vector_2```.
```
vector_2 = np.array([11, 12])
vector_2.shape
```
* La siguiente celda ejecutará la función ```np.cross()``` con ```vector_1``` y ```vector_2```.
```
np.cross(vector_1, vector_2)
vector_3 = np.array([[1, 2, 3]])
vector_4 = np.array([11, 12, 13])
np.cross(vector_3, vector_4)
np.cross(vector_3, vector_4).shape
```
## El paquete ```numpy.linalg```.
La biblioteca especialziada en operaciones de algebra lineal de *Numpy* es ```numpy.linalg```.
El estudio de todas las funciones contenodas en este paquete están fuera de los alcances de este curso, pero se ejemplificarán las funciones.
* ```np.linalg.det()```
* ```np.linalg.solve()```
* ```np.linalg.inv()```
https://numpy.org/doc/stable/reference/routines.linalg.html
```
import numpy.linalg
```
### Cálculo del determinate de una matriz mediante ```numpy.linalg.det()```.
**Ejemplo:**
* Se calculará el determinante de la matriz:
$$ \det\begin{vmatrix}0&1&2\\3&4&5\\6&7&8\end{vmatrix}$$
* El cálculo del determinante es el siguiente:
$$ ((0 * 4 * 8) + (1 * 5 * 6) + (2 * 3 * 7)) - ((6 * 4 * 2) + (7 * 5 * 0) + (8 * 3* 1)) = 0$$
```
matriz = np.arange(9).reshape(3, 3)
matriz
numpy.linalg.det(matriz)
```
* Se calculará el determinante de la matriz:
$$ \det\begin{vmatrix}1&1&2\\3&4&5\\6&7&8\end{vmatrix}$$
* El cálculo del determinante es el siguiente:
$$ ((1 * 4 * 8) + (1 * 5 * 6) + (2 * 3 * 7)) - ((6 * 4 * 2) + (7 * 5 * 1) + (8 * 3* 1)) = -3$$
```
matriz = np.array([[1, 1, 2],
[3, 4, 5],
[6, 7, 8]])
numpy.linalg.det(matriz)
```
### Soluciones de ecuaciones lineales con la función ```np.linalg.solve()```.
Un sistema de ecuaciones lineales coresponde un conjunto de ecuaciones de la forma:
$$
a_{11}x_1 + a_{12}x_2 + \cdots a_{1n}x_n = y_1 \\
a_{21}x_1 + a_{22}x_2 + \cdots a_{2n}x_n = y_2\\
\vdots\\
a_{m1}x_1 + a_{m2}x_2 + \cdots a_{mn}x_n = y_m
$$
Lo cual puede ser expresado de forma matricial.
$$
\begin{bmatrix}a_{11}\\a_{21}\\ \vdots\\ a_{m1}\end{bmatrix}x_1 + \begin{bmatrix}a_{12}\\a_{22}\\ \vdots\\ a_{m2}\end{bmatrix}x_2 + \cdots \begin{bmatrix}a_{m1}\\a_{m2}\\ \vdots\\ a_{mn}\end{bmatrix}x_n = \begin{bmatrix}y_{1}\\y_{2}\\ \vdots\\ y_{m}\end{bmatrix}
$$
Existen múltiples métodos para calcular los valores $x_1, x_2 \cdots x_n$ que cumplan con el sistema siempre que $m = n$.
Numpy cuenta con la función *np.linalg.solve()*, la cual puede calcular la solución de un sistema de ecuaciones lineales al expresarse como un par de matrices de la siguiente foma:
$$
\begin{bmatrix}a_{11}&a_{12}&\cdots&a_{1n}\\a_{21}&a_{22}&\cdots&a_{2n}\\ \vdots\\ a_{n1}&a_{n2}&\cdots&a_{nn}\end{bmatrix}= \begin{bmatrix}y_{1}\\y_{2}\\ \vdots\\ y_{n}\end{bmatrix}
$$
La función ```numpy.linagl.solve()``` permite resolver sistemas de ecuaciones lineales ingresando un arreglo de dimensiones ```(n, n)``` como primer argumente y otro con dimensión ```(n)``` como segundo argumento.
**Ejemplo:**
* Para resolver el sistema de ecuaciones:
$$
2x_1 + 5x_2 - 3x_3 = 22.2 \\
11x_1 - 4x_2 + 22x_3 = 11.6 \\
54x_1 + 1x_2 + 19x_3 = -40.1 \\
$$
* Se realiza lo siguiente:
```
a = np.array([[2, 5, -3],
[11, -4, 22],
[54, 1, 19]])
a.shape
y = np.array([22.2, 11.6, -40.1])
y.shape
np.linalg.solve(a, y)
```
## Matriz inversa.
```
np.linalg.inv(a)
np.linalg.inv(a).dot(y)
```
<p style="text-align: center"><a rel="license" href="http://creativecommons.org/licenses/by/4.0/"><img alt="Licencia Creative Commons" style="border-width:0" src="https://i.creativecommons.org/l/by/4.0/80x15.png" /></a><br />Esta obra está bajo una <a rel="license" href="http://creativecommons.org/licenses/by/4.0/">Licencia Creative Commons Atribución 4.0 Internacional</a>.</p>
<p style="text-align: center">© José Luis Chiquete Valdivieso. 2021.</p>
| github_jupyter |
# Python Primer Exercises (with Solutions)
## Lesson 1 - Variables, Expressions, and Functions
----
### Exercise: Types
Consider the following code:
```python
var1 = 1
var2 = 1.
var3 = "one"
var4 = "3.14159"
```
What is the type of `var1`?
`int`
What is the type of `var2`?
`float`
What is the type of `var3`?
`str`
What is the type of `var4`?
`str`
### Exercise: Find the type
Determine the type of the variable `mystery` using a built-in function
```python
mystery = {"a": 1, "b": 2}
```
```
mystery = {"a": 1, "b": 2}
type(mystery)
```
### Exercise: Converting types
Given this:
```python
var2 = 1.
var4 = "1.0"
```
1. Explain in simple terms what `float(var4)` does.
1. Explain in simple terms what `str(var2)` does.
```
var2 = 1.
var4 = "1.0"
## float(var4) turns var4 from a str to a float
float(var4)
## str(var2) turns var2 from a float to a str
str(var2)
```
### Exercise: Imports
Fill in the blanks so that the two programs below run without errors.
```python
import __
print("The current directory is {}".format(os.getcwd()))
import ____
print("The square root of 2 is {}".format(math.sqrt(2)))
```
```
import os
print("The current directory is {}".format(os.getcwd()))
import math
print("The square root of 2 is {}".format(math.sqrt(2)))
```
## Lesson 2: Lists and Strings
----
### Exercise: Slicing
What does the following print:
```python
material = 'carbon'
print('material[1:3] is:', material[1:3])
```
In words, explain what does `material[low:high]` do? (where low and high are arbitrary numbers)
```
material = 'carbon'
print('material[1:3] is:', material[1:3])
## it returns a slice of element from position low to position high (starting from 0), e.g.
material[2:4]
```
In words, explain what does `material[low:]` do? (where low is an arbitrary number)
```
## it returns everything in element starting from position low, e.g.
material[2:]
```
In words, explain what does `material[:high]` do? (where low is an arbitrary number)
```
## it returns everything in element before position high, e.g.
material[:2]
```
What does `material[:]` do?
```
## it returns the entire string
material[:]
```
What about `material[::2]`? `material[::-1]`?
```
## material[::2] returns every second character of material
material[::2]
## material[:-1] returns every character, in reverse order
material[::-1]
```
### Exercise: Fill in the blanks
Fill in the blanks so that the program below produces the output shown.
```python
values = ____
values.____(1)
values.____(3)
values.____(5)
print('first time:', values)
values = values[____]
print('second time:', values)
```
---
```
first time: [1, 3, 5]
second time: [3, 5]
```
```
values = []
values.append(1)
values.append(3)
values.append(5)
print('first time:', values)
values = values[1:]
print('second time:', values)
```
### Exercise: From strings to lists and back
Given this:
```python
print('string to list:', list('tin'))
print('list to string:', '-'.join(['g', 'o', 'l', 'd']))
```
---
```
['t', 'i', 'n']
'g-o-l-d'
```
1. Explain in simple terms what `list('some string')` does.
```
## list('some string') returns a list, where each element of the list is one character of the string (including spaces), e.g.
list('some string')
```
2. What does `' <=> '.join(['x', 'y'])` generate?
```
## '.'.join(list) returns the string of every element in a list, separated by the str that precedes 'join'
' <=> '.join(['x','y','z'])
```
### Exercise: Sort and Sorted
What do these two programs print? In simple terms, explain the difference between `sorted(letters)` and `letters.sort()`.
```python
## Program A
letters = list('gold')
result = sorted(letters)
print('letters is', letters, 'and result is', result)
```
---
```python
## Program B
letters = list('gold')
result = letters.sort()
print('letters is', letters, 'and result is', result)
```
`sorted(letters)` returns a new list containing the elements of `letters` sorted alphebetically
```
## Program A
letters = list('gold')
result = sorted(letters)
print('letters is', letters, 'and result is', result)
```
`letters.sort()` sorts `letters` alphabetically and doesn't return anyting (`result` is `None`)
```
## Program B
letters = list('gold')
result = letters.sort()
print('letters is', letters, 'and result is', result)
```
## Lesson 3: For loops
### Exercise: write a for loop that computes the sum of all of the squares from 2 to 11.
```
s = 0
for i in range(10):
s += (i + 2)**2
print(s)
```
### Exercise: write a for loop that prints each item in the list `friends`, but stops when it encounters the third value
`friends = ["Rachel", "Monica", "Chandler", "Ross", "Joey"]`
```
friends = ["Rachel", "Monica", "Chandler", "Ross", "Joey"]
for i, friend in enumerate(friends):
if i == 3:
break
else:
print(friend)
```
### Exercise: Explain in a few words what happens when you call `zip` on two iterables that have different lengths
`zip` will stop when it reaches the end of the shorter iterable
```
s1 = 'Jim'
s2 = 'Spock'
for a, b in zip(s1,s2):
print (a,b)
```
### Exercise (bonus): Write a Python program to construct the following pattern, using a nested for loop.
<dd>* </dd>
<dd>* * </dd>
<dd>* * * </dd>
<dd>* * * * </dd>
<dd>* * * * * </dd>
<dd>* * * * </dd>
<dd>* * * </dd>
<dd>* * </dd>
<dd>* </dd>
Hint: by default the `print` function inserts a newline character at the end of every line it prints. To suppress this, set the `end` argument to "", like `print("something", end="")`
Hint: you can iterate through a range of numbers in reverse order by passing a negative number as a third argument to `range`, e.g. `range(0,10,-1)`
```
n=5
for i in range(n): # rows
for j in range(i): # columns
print ('* ', end="")
print('')
for i in range(n,0,-1):
for j in range(i):
print('* ', end="")
print('')
```
| github_jupyter |
# `pandas` Part 7: Combining Datasets with `concat()`
# Learning Objectives
## By the end of this tutorial you will be able to:
1. Combine DataFrames and/or Series with `concat()`
2. Understand a multi-index
3. Reset an index with `reset_index()`
4. Perform descriptive analytics on a combined DataFrame
## Files Needed for this lesson:
>- `CAvideos.csv`
>- `GBvideos.csv`
>- Download this csv files from Canvas prior to the lesson
>- C:\\Users\\mimc2537\\OneDrive - UCB-O365\\python\\pandas\\
## The general steps to working with pandas:
1. import pandas as pd
2. Create or load data into a pandas DataFrame or Series
3. Reading data with `pd.read_`
>- Excel files: `pd.read_excel('fileName.xlsx')`
>- Csv files: `pd.read_csv('fileName.csv')`
>- Note: if the file you want to read into your notebook is not in the same folder you can do one of two things:
>>- Move the file you want to read into the same folder/directory as the notebook
>>- Type out the full path into the read function
4. After steps 1-3 you will want to check out your DataFrame
>- Use `shape` to see how many records and columns are in your DataFrame
>- Use `head()` to show the first 5-10 records in your DataFrame
# Introduction Notes on Combining Data Using `pandas`
1. Being able to combine data from multiple sources is a critical skill for analytics professionals
2. We will learn the `pandas` way of combining data but there are similarities here to SQL
3. Why combine data with `pandas` if you can do the same thing in SQL?
>- The answer to this depends on the project
>- Some projects may be completed more efficiently all with `pandas` so you wouldn't necessarily need SQL
>- For some projects incorporating SQL into our python code makes sense
>- In a an analytics job, you will likely use both python and SQL to get the job done!
# Initial set-up steps
1. import modules and check working directory
2. Read data in
3. Check the data
# Step 2 Read Data Into a DataFrame with `read_csv()`
>- file names:
>>- `CAvideos.csv`
>>- `GBvideos.csv`
### Check how many rows and columns are in our DataFrames
### Check a couple of rows of data in one of the new DataFrames
## Check the datatypes
# Combining DataFrames
>- The three common ways to combine datastest in pandas is with `concat()`, `join()`, and `merge()`
>- `concat()` will take two DataFrames or Series and append them together
>>- This is basically taking DataFrames and stacking their data on top of each other into one DataFrame
>>- For `concat()` you need the columns/fields in both DataFrames to the be the same
>- `join()` "links" DataFrames together based on a common field/column between the two
>- `merge()` also links DataFrames together based on common field/columns but with different syntax.
>>- We will cover the most basic join in this class
>>- A more in depth study of joins is provided in SQL focused courses
>>- Pandas join reference for further study: https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.join.html
# Using the YouTube DataFrames to practice combining data with pandas
>- The YouTube datasets store data on various YouTube trending statistics
>- Our example datasets show several months of data and daily trending YouTube videos.
>- For more information and for other YouTube datasets see the following link:
>>- https://www.kaggle.com/datasnaek/youtube-new
### First, creating a new DataFrame that appends the Canadian and British YouTube DataFrames
#### Some notes on the previous code
>- Line 1: We define a new DataFrame named `CanUK` which is defined as the concatenation,`concat()`, of two datasets
>>- Dataset 1 = canadian_youtube
>>- Dataset 2 = uk_youtube
>>- The `concat()` function takes the two (or more if applicable) DataFrames and "stacks" them on top of each
>- Line 2: We use `keys` option to define a multi-index (aka hierarchical index)
>>- Because our datasets represent YouTube videos from different countries we pass the abbreviated names of those countries as a list to `keys`
>>- Enter the keys names in order they appear in line 1 (e.g., 'can' first, 'uk' second)
>- Line 3: We use the `names` option to label our index columns from line 2
>>- Without the `names` option we would not have anything above our index columns
### Check the index for any dataframe using `DataFrame.index`
>- Note how `concat()` uses the rowid's for each country's dataset versus continuing the count
### Take a look at our new DataFrame
#### Did using `concat()` work to append the two DataFrames together?
>- Check the shape of your new DataFrame
>- Compare the number of records to each one individually
>>- canadian_youtube = 40881 records
>>- uk_youtube = 38916 records
>>- 40881 + 38916 = 79797 total records
#### `reset_index`:
##### Note: You can reset a an index with `reset_index`
>- This can be useful for some situations
>- For a multi-index you can pass the `level` option and specify what index you want to reset
>- Note: To make the change to our current DataFrame we would need to use the option, `inplace=True`
# Now some descriptive analytics
### What channels have the most trending videos?
### What are the quantitative descriptive statistics for TheEllenShow?
##### Alternatively, you can use `loc[]` to peform the filtering operation
>- The use of `where()` or `loc[]` depends on the question/purpose or sometimes just personal preference
### What were the total YouTube videos, total views, likes and dislikes for TheEllenShow?
>- Using the agg() function to calculate specific aggregations on different columns
## What are the totals for TheEllenShow's top 5 most viewed videos?
>- Only include the title names as part of the output (not channel or any other categorical fields)
>- Include total views, likes, dislikes, and comment count in the output
# Some Notes on the Previous Example
>- Our pandas code in the previous example is similar to SQL in the following ways
1. `loc[CanUk.channel_title == 'TheEllenShow',` is SQL equivalent to `WHERE channel_title = 'TheEllenShow'`
2. `['title','views','likes','dislikes','comment_count']` is SQL equivalent to:
`SELECT title, sum(views),sum(likes),sum(dislikes),sum(comment_count)`
3. `groupby(['title`]) is SQL equivalent to GROUP BY title
4. Now in pandas we enter the aggregation after the `groupby()`, in this example `sum()`
>>- In SQL we write the aggregation in the SELECT statement
## In future lessons we will continue to learn how pandas and SQL relate
| github_jupyter |
```
import panel as pn
pn.extension()
```
Panel objects are built on top of [Param](https://param.pyviz.org), which allows Panel objects to declare parameters that a user can flexibly manipulate to control the displayed output. In addition to parameters specific to each component and class of components, all components define a shared set of parameters to control the size and style of the rendered views.
## Styling Components
#### ``css_classes``
The ``css_classes`` parameter allows associating a Panel component with one or more CSS classes. CSS styles can be embedded in raw form or by referencing an external .css file by providing each to the panel extension using the ``raw_css`` and ``css_files`` arguments; both should be supplied as lists. Outside the notebook or if we want to add some CSS in an external module or library, we can simply append to the ``pn.config.raw_css`` and ``pn.config.js_files`` config parameters.
To demonstrate this usage, let us define a CSS class called ``panel-widget-box`` which we will give a background and a nice border:
```
css = '''
.bk.panel-widget-box {
background: #f0f0f0;
border-radius: 5px;
border: 1px black solid;
}
'''
pn.extension(raw_css=[css])
```
Now that we have defined and loaded the CSS we will create a ``Column`` with the widget-box CSS class:
```
pn.Column(
pn.widgets.FloatSlider(name='Number', margin=(10, 5, 5, 10)),
pn.widgets.Select(name='Fruit', options=['Apple', 'Orange', 'Pear'], margin=(0, 5, 5, 10)),
pn.widgets.Button(name='Run', margin=(5, 10, 10, 10)),
css_classes=['panel-widget-box'])
```
#### ``background``
In case we simply want to give the component a background we can define one as a hex string:
```
pn.pane.HTML(background='#f307eb', width=100, height=100)
```
#### `loading`
All components also have a `loading` parameter which indicates that they are currently processing some event. Setting the parameter will display the global `loading_spinner` on top of the component. To configure the loading spinner you can set the:
* `pn.config.loading_spinner`: The style of the global loading indicator, e.g. 'arcs', 'bars', 'dots', 'petals'.
* `pn.config.loading_color`: The color of the global loading indicator as a hex color, e.g. #6a6a6a
In the notebook these should be configured before loading the `pn.extension` and may in fact be set using `pn.extension(loading_spinner='dots', loading_color='#00aa41')`.
```
pn.pane.HTML(background='#00aa41', width=100, height=100, loading=True)
```
#### ``style``
Certain components, specifically markup-related panes, expose a ``style`` parameter that allows defining CSS styles applying to the HTML container of the pane's contents, e.g. the ``Markdown`` pane:
```
pn.pane.Markdown('### A serif Markdown heading', style={'font-family': "serif"})
```
#### ``visible``
All components provide a `visible` parameter which allows toggling whether the component should be visible or not. Below we display a set of components and provide some widgets to toggle the `visible` property on or off:
```
a = pn.pane.HTML(width=60, height=60, background='green')
b = pn.pane.HTML(width=60, height=60, background='blue', visible=False)
c = pn.pane.HTML(width=60, height=60, background='red')
layout = pn.Row(a, b, c)
controls = pn.Row(*(c.controls(['visible'])[1] for c in layout))
for c in controls:
c.width = 50
pn.Column(controls, layout)
```
## Component Size and Layout
The size of components and their spacing is also controlled through a set of parameters shared by all components.
#### ``margin``
The ``margin`` parameter can be used to create space around an element defined as the number of pixels at the (top, right, bottom, and left). The ``margin`` can be defined in one of three ways:
margin=25
top, bottom, left, and right margins are 25px
margin=(25, 50)
top and bottom margins are 25px
right and left margins are 50px
margin=(25, 50, 75, 100)
top margin is 25px
right margin is 50px
bottom margin is 75px
left margin is 100px
```
pn.Row(
pn.Column(pn.widgets.Button(name='Run', margin=25), background='#f0f0f0'),
pn.Column(pn.widgets.Button(name='Run', margin=(25, 50)), background='#f0f0f0'),
pn.Column(pn.widgets.Button(name='Run', margin=(25, 50, 75, 100)), background='#f0f0f0'))
```
## ``align``
The `align` parameter controls how components align vertically and horizontally. It supports 'start', 'center', and 'end' values and can be set for both horizontal and vertical directions at once or for each separately by passing in a tuple of the form `(horizontal, vertical)`.
One common use-case where alignment is important is when placing multiple items with different heights in a `Row`:
```
pn.Row(pn.widgets.IntSlider(name='Test'), pn.widgets.IntSlider(align='end'))
```
In a grid you may also have to specify the horizontal and vertical alignment separately to achieve the layout you are after:
```
pn.GridBox(
pn.widgets.IntSlider(name='Test'), pn.widgets.IntSlider(align='end'),
pn.widgets.TextInput(name='Test', width=150), pn.widgets.TextInput(width=150, align=('start', 'end')),
ncols=2)
```
### Absolute sizing using ``width`` and ``height``
By default all components use either auto-sizing or absolute sizing. Panels will generally take up as much space as the components within them, and text or image-based panes will adjust to the size of their contents. To set a fixed size on a component, it is usually sufficient to set a width or height, but in certain cases setting ``sizing_mode='fixed'`` explicitly may also be required.
```
pn.Row(
pn.pane.Markdown('ABCDE', background='#f0f0f0', width=200, height=200),
pn.pane.PNG('https://upload.wikimedia.org/wikipedia/commons/4/47/PNG_transparency_demonstration_1.png', width=200),
pn.widgets.FloatSlider(width=200))
```
#### Plots
Unlike other components, the size of a plot is usually determined by the underlying plotting library, so it may be necessary to ensure that you set the size and aspect when declaring the plot.
### Responsive sizing
By default, panel objects will use a fixed size if one is provided or adapt to the size of the content. However most panel objects also support reactive sizing which adjusts depending on the size of the viewport. These responsive sizing modes can be controlled using the ``sizing_mode`` parameter.
#### ``sizing_mode``
* **"fixed"**: Component is not responsive. It will retain its original width and height regardless of any subsequent browser window resize events.
This is usually the default behavior and simply respects the provided width and height.
* **"stretch_width"**: Component will responsively resize to stretch to the available width, without maintaining any aspect ratio. The height of the component depends on the type of the component and may be fixed or fit to component's contents.
To demonstrate this behavior we create a Row with a fixed height and responsive width to fill:
```
pn.Row(
pn.pane.Str(background='#f0f0f0', height=100, sizing_mode='stretch_width'),
width_policy='max', height=200
)
```
* **"stretch_height"**: Component will responsively resize to stretch to the available height, without maintaining any aspect ratio. The width of the component depends on the type of the component and may be fixed or fit to component's contents.
To demonstrate the filling behavior in a document we declare a Column with a fixed height for the component to fill:
```
pn.Column(
pn.pane.Str(background='#f0f0f0', sizing_mode='stretch_height', width=200),
height=200
)
```
* **"stretch_both"**: Component is completely responsive, independently in width and height, and will occupy all the available horizontal and vertical space, even if this changes the aspect ratio of the component.
To demonstrate this behavior we will declare a Column with a fixed height and responsive width for the component to fill:
```
pn.Column(
pn.pane.Str(background='#f0f0f0', sizing_mode='stretch_both'),
height=200, width_policy='max'
)
```
* **"scale_height"**: Component will responsively resize to stretch to the available height, while maintaining the original or provided aspect ratio.
* **"scale_width"**: Component will responsively resize to stretch to the available width, while maintaining the original or provided aspect ratio.
* **"scale_both"**: Component will responsively resize to both the available width and height, while maintaining the original or provided aspect ratio.
```
pn.Column(
pn.pane.PNG('https://upload.wikimedia.org/wikipedia/commons/4/47/PNG_transparency_demonstration_1.png', sizing_mode='scale_both'),
height=400, width=500, background='#3f3f3f')
```
### Spacers
Spacers are a very versatile component which makes it easy to put fixed or responsive spacing between objects. Like all other components spacers support both absolute and responsive sizing modes:
```
pn.Row(1, pn.Spacer(width=200), 2, pn.Spacer(width=100), 3, pn.Spacer(width=50), 4, pn.Spacer(width=25), 5)
```
``VSpacer``and ``HSpacer`` provide responsive vertical and horizontal spacing, respectively. Using these components we can space objects equidistantly in a layout and allow the empty space to shrink when the browser is resized.
```
pn.Row(
pn.layout.HSpacer(), '* Item 1\n* Item2', pn.layout.HSpacer(), '1. First\n2. Second', pn.layout.HSpacer()
)
```
| github_jupyter |
<h1 style="text-align:center">Fourier Analysis using SymPy</h1>
<h3 style="text-align:center"> MCHE 485: Mechanical Vibrations</h3>
<p style="text-align:center">Dr. Joshua Vaughan <br>
<a href="mailto:joshua.vaughan@louisiana.edu">joshua.vaughan@louisiana.edu</a><br>
http://www.ucs.louisiana.edu/~jev9637/ </p>
This notebook will look at the Fourier Analysis of periodic functions. In this notebook, we'll use [SymPy](http://sympy.org/en/index.html), "a Python library for symbolic mathematics."
We can write any periodic function as an infinite sum of sines and cosines:
$ \quad f(t) = \sum_{n=0}^{\infty}a_n\cos(n\omega_0t) + \sum_{n=1}^{\infty}b_n\sin(n\omega_0t) $
where
$ \quad a_n = \frac{\omega_0}{\pi}\int_0^{\frac{2\pi}{\omega_0}}f(t)\cos(n\omega_0t)dt $,
$ \quad b_n = \frac{\omega_0}{\pi}\int_0^{\frac{2\pi}{\omega_0}}f(t)\sin(n\omega_0t)dt $,
and
$ \quad a_0 = \frac{\omega_0}{2\pi}\int_0^{\frac{2\pi}{\omega_0}}f(t)dt $
For more information on this process, you can see the lectures at the [class website](http://www.ucs.louisiana.edu/~jev9637/MCHE485.html).
By changing the number of terms we use to approximate the original function we can approach its shape, as shown in Figure 1.
<p style="text-align:center">
<img src="http://shared.crawlab.org/SquareWave_Fourier_Approx.gif" alt="Fourier Analysis" width=50%/></a><br>
<strong> Figure 1: Increasing the Number of Terms in the Approximation </strong>
</p>
The remainder of this notebook will focus on determining the $a_n$ and $b_n$ terms in the Fourier Expansion.
<br>
<br>
```
# import Sympy and start "pretty printing"
import sympy
sympy.init_printing()
# Define the sympy symbolic variables we'll need
t, w0, tau_0 = sympy.symbols(['t', 'omega_0', 'tau_0'], real=True, positive=True)
#--------- Input your function to examine here --------
# Use the sympy Piecewise function to define the square wave - This matches the one in the Figure 1 above.
y = 2 + sympy.Piecewise((1, t < sympy.pi/w0), (-1, t > sympy.pi/w0))
# Use the sympy Piecewise function to define the triangle wave
# First define F0
# F0 = sympy.symbols('F0')
# y = sympy.Piecewise((F0/2*t, t < sympy.pi/w0), (-(F0/2)*t + 2*F0, t >= sympy.pi/w0))
# Use the sympy Piecewise function to define a trapezoid function
# y = sympy.Piecewise((3*F0*w0/(2*sympy.pi)*t, t < (2*sympy.pi/(3*w0))), (F0, t < (4*sympy.pi/(3*w0))),
# (-3*F0*w0/(2*sympy.pi)*t + 3*F0, t > (4*sympy.pi/(3*w0))))
# define the number of terms to use in the approximation
num_terms = 7
# get the a0 term
a0 = w0 / (2*sympy.pi) * sympy.integrate(y, (t, 0, 2*sympy.pi/w0))
# Define matrices of 0s to fill the the an and bn terms
a = sympy.zeros(1, num_terms)
b = sympy.zeros(1, num_terms)
# cycle through the 1 to num_terms Fourier coefficients (a_n and b_n)
for n in range(num_terms):
integral_cos = y * sympy.cos((n+1)*w0*t) # define the integral "interior"
a[n] = w0 / sympy.pi * sympy.integrate(integral_cos, (t, 0, 2*sympy.pi/w0)) # solve for a_n
integral_sin = y * sympy.sin((n+1)*w0*t) # define the integral "interior"
b[n] = w0 / sympy.pi * sympy.integrate(integral_sin, (t, 0, 2*sympy.pi/w0)) # solve for b_n
# Simplify and display a0
sympy.simplify(a0)
# Simplify and display the an terms
sympy.simplify(a)
# Simplify and diplay the bn terms
sympy.simplify(b)
```
<hr style="border: 0px;
height: 1px;
text-align: center;
background: #333;
background-image: -webkit-linear-gradient(left, #ccc, #333, #ccc);
background-image: -moz-linear-gradient(left, #ccc, #333, #ccc);
background-image: -ms-linear-gradient(left, #ccc, #333, #ccc);
background-image: -o-linear-gradient(left, #ccc, #333, #ccc);">
#### Licenses
Code is licensed under a 3-clause BSD style license. See the licenses/LICENSE.md file.
Other content is provided under a [Creative Commons Attribution-NonCommercial 4.0 International License](http://creativecommons.org/licenses/by-nc/4.0/), CC-BY-NC 4.0.
```
# This cell will just improve the styling of the notebook
from IPython.core.display import HTML
import urllib.request
response = urllib.request.urlopen("https://cl.ly/1B1y452Z1d35")
HTML(response.read().decode("utf-8"))
```
| github_jupyter |
# Lab4: Gradient Blending
This project explores gradient-domain processing, a simple technique with a broad set of applications including blending, tone-mapping, and non-photorealistic rendering. This specific project explores seamless image compositing via "Poisson blending".
The primary goal of this assignment is to seamlessly blend an object or texture from a source image into a target image. The simplest method would be to just copy and paste the pixels from one image directly into the other (and this is eactly what the starter code does). Unfortunately, this will create very noticeable seams, even if the backgrounds are similar. How can we get rid of these seams without doing too much perceptual damage to the source region?
The insight is that people are more sensitive to gradients than absolute image intensities. So we can set up the problem as finding values for the output pixels that maximally preserve the gradient of the source region without changing any of the background pixels. Note that we are making a deliberate decision here to ignore the overall intensity! We will add an object into an image by reintegrating from (modified) gradients and forgetting whatever absolute intensity it started at.
## Simple 1d Examples
Let's start with a simple case where instead of copying in new gradients we only want to fill in a missing region of an image and keep the gradients as smooth (close to zero) as possible. To simplify things further, let's start with a one dimensional signal instead of a two dimensional image.
Here is our signal t and a mask M specifying which "pixels" are missing.

```python
t = [5 4 0 0 0 0 2 4];
M = [0 0 1 1 1 1 0 0];
M = logical(M);
```
We can formulate our objective as a least squares problem. Given the intensity values of t, we want to solve for new intensity values v under the mask M such that

Here i is a coordinate (1d or 2d) for each pixel under mask M. Each j is a neighbor of i. Each summation guides the gradient (the local pixel differences) in all directions to be close to 0. In the first summation, the gradient is between two unknown pixels, and the second summation handles the border situation where one pixel is unknown and one pixel is known (outside the mask M). Minimizing this equation could be called a Poisson fill.
For this example let's define neighborhood to be the pixel to your left (you could define neighborhood to be all surrounding pixels. In 2d, you would at least need to consider vertical and horizontal neighbors). The least squares solution to the following system of equations satisfies the formula above.
```python
v(1) - t(2) = 0; %left border
v(2) - v(1) = 0;
v(3) - v(2) = 0;
v(4) - v(3) = 0;
t(7) - v(4) = 0; %right border
```
Note that the coordinates don't directly correspond between v and t — v(1), the first unknown pixel, sits on top of t(3). You could formulate it differently if you choose. Plugging in known values of t we get
```python
v(1) - 4 = 0;
v(2) - v(1) = 0;
v(3) - v(2) = 0;
v(4) - v(3) = 0;
2 - v(4) = 0;
```
Now let's convert this to matrix form and solve it
```python
A = [ 1 0 0 0; ...
-1 1 0 0; ...
0 -1 1 0; ...
0 0 -1 1; ...
0 0 0 -1];
b = [4; 0; 0; 0; -2];
```

As it turns out, in the 1d case, the Poisson fill is simply a linear interpolation between the boundary values. But in 2d the Poisson fill exhibits more complexity. Now instead of just doing a fill, let's try to seamlessly blend content from one 1d signal into another. We'll fill the missing values in t using the correspondig values in s:
```python
s = [5 6 7 2 4 5 7 3];
```

Now our objective changes — instead of trying to minimize the gradients, we want the gradients to match another set of gradients (those in s). We can write this as follows:

We minimize this by finding the least squares solution to this system of equations:
```python
v(1) - t(2) = s(3) - s(2);
v(2) - v(1) = s(4) - s(3);
v(3) - v(2) = s(5) - s(4);
v(4) - v(3) = s(6) - s(5);
t(7) - v(4) = s(7) - s(6);
```
After plugging in known values from t and s this becomes:
```python
v(1) - 4 = 1;
v(2) - v(1) = -5;
v(3) - v(2) = 2;
v(4) - v(3) = 1;
2 - v(4) = 2;
```
Finally, in matrix form for Matlab
```python
A = [ 1 0 0 0; ...
-1 1 0 0; ...
0 -1 1 0; ...
0 0 -1 1; ...
0 0 0 -1];
b = [5; -5; 2; 1; 0];
```
If we solve Ax=b and combine the values of x with t, we get:

## Task 1: 1D Blending (10)
Simply implement the example above and plot the results for linear interpolation and using the gradients from S. Note that the matrix A is the same!!
```
%matplotlib inline
from pylab import *
import numpy as np
t = np.array([5., 4., 0., 0., 0., 0., 2., 4.])
mask = np.array([1, 1, 0, 0, 0, 0, 1, 1])
s = np.array([5., 6., 7., 2., 4., 5., 7., 3.]);
poisson_filling = [ 5., 4., 3.6, 3.2, 2.8, 2.4, 2., 4. ]
plot (poisson_filling)
gradient_composite = [ 5., 4., 4.4, -1.2, 0.2, 0.6, 2., 4. ]
plot (gradient_composite)
mSize=np.count_nonzero(mask)
size=len(t)-mSize
A=np.eye(size + 1, size) - np.eye(size + 1, size, k = -1)
b=zeros(size+1)
offset=1
b[0]+=t[offset]
b[-1]-=t[offset+mSize+1]
lin=np.linalg.lstsq(A,b)
poisson=t
poisson[offset+1:offset+mSize+1] = lin[0]
plot (poisson)
b2=s[offset+1:offset+mSize+2] - s[offset:offset+mSize+1]
b2[0]+=t[offset]
b2[-1]-=t[offset+mSize+1]
gr=np.linalg.lstsq(A,b2)
gradient=t
gradient[offset+1:offset+mSize+1] = gr[0]
plot (gradient)
```
## Poisson reconstruction in 2D
Notice that in our quest to preserve gradients without regard for intensity we might have gone too far — our signal now has negative values. The same thing can happen in the image domain, so you'll want to watch for that and at the very least clamp values back to the valid range. When working with images, the basic idea is the same as above, except that each pixel has at least two neighbors (left and top) and possibly four neighbors. Either formulation will work.
For example, in a 2d image using a 4-connected neighborhood, our equations above imply that for a single pixel in v, at coordinate (i,j) which is fully under the mask you would have the following equations:
```python
v(i,j) - v(i-1, j) = s(i,j) - s(i-1, j)
v(i,j) - v(i+1, j) = s(i,j) - s(i+1, j)
v(i,j) - v(i, j-1) = s(i,j) - s(i, j-1)
v(i,j) - v(i, j+1) = s(i,j) - s(i, j+1)
```
In this case we have many equations for each unknown. It may be simpler to combine these equations such that there is one equation for each pixel, as this can make the mapping between rows in your matrix A and pixels in your images easier. Adding the four equations above we get:
```python
4*v(i,j) - v(i-1, j) - v(i+1, j) - v(i, j-1) - v(i, j+1) = 4*s(i,j) - s(i-1, j) - s(i+1, j) - s(i, j-1) - s(i, j+1)
```
Note that the right hand of this equation is known and coincides with the discrete Laplacian that we computed in previous labs!
## Task 2: 2D Poisson Reconstruction (20)
!BE AWARE! This will be slow...
For this task we will take a simple image, compute it's laplacian and reconstruct from this laplacian the original image. This will be very slow, so only use it for the small test image.
* For color images, process each color channel independently (hint: matrix A won't change, so don't go through the computational expense of rebuilding it for each color channel).
```
%matplotlib inline
from pylab import *
import imageio as imio
import numpy as np
import scipy.ndimage as nd
import scipy.sparse
im = nd.imread("data/test.png")[:,:,:3]/255.
def Laplacian(im):
laplacian_kernel = np.array([[0, -1, 0], [-1, 4, -1], [0, -1, 0]])
#{‘reflect’,’constant’,’nearest’,’mirror’, ‘wrap’}
conv = nd.convolve(im, laplacian_kernel,mode='constant')
return conv
def add_border(im):
border_im = np.zeros((im.shape[0]+2,im.shape[1]+2,im.shape[2]))
border_im[1:-1,1:-1] = im
return border_im
def buildSystemNaive(im_shape):
A = np.identity(np.prod(im_shape))
size=im_shape[0]
#complete the values of A
A=A*4.
print len(A)
A=A - np.eye(size*size, k=1) - np.eye(size**2, k = -1)
A=A - np.eye(size*size, k=-size) - np.eye(size**2, k = size)
print A
return A
im = add_border(im)
# this is done once for all three channels
A = buildSystemNaive(im[:,:,0].shape)
b = Laplacian(im[:,:,0])
red = np.linalg.lstsq(A, b.ravel())[0]
red = np.clip(red.reshape(b.shape),0.0,1.0)
b2 = Laplacian(im[:,:,1])
green = np.linalg.lstsq(A, b2.ravel())[0]
green = np.clip(green.reshape(b2.shape),0.0,1.0)
b3 = Laplacian(im[:,:,2])
blue = np.linalg.lstsq(A, b3.ravel())[0]
blue = np.clip(blue.reshape(b3.shape),0.0,1.0)
result =np.dstack((red,green,blue))
# The result should be the original image
imshow(result)
import scipy.sparse as sp
# imshow(im)
imshow((result-im)*10**12)
x=sp.csr_matrix(A)
x[0,0]=20
# print x
```
## Task 3: 2D Poisson Reconstruction Sparse (30)
The linear system of equations (and thus the matrix A) becomes enormous. But A is also very sparse because each equation only relates a pixel to some number of its immediate neighbors.
A needs at least as many rows and columns as there are pixels in the masked region (in our example the hole image).
If the mask covers 100,000 pixels, this implies a matrix with at least 100,000,000,000 entries. Don't try that.
So we will use sparse matrix and a least squares solver for sparse matrices. You can use whatever matrix format you want for indexing, but when returning A, transform it to one of the compressed formats, A = A.tocsr() or A.tocsc(), that improves the performance of the solvers by a lot.
I tested scipy sparse. https://docs.scipy.org/doc/scipy-0.15.1/reference/sparse.html
You have lmrs and lmqr but they are quite slow. The iterative solvers are much faster. http://www.scipy-lectures.org/advanced/scipy_sparse/solvers.html Any of them should give you a fair performance. In my case the congugate gradient seemed to be the fastest.
You can try with a bigger image now. Do not forget to add the border so the image is properly reconstructed.
Now building the matrix A is probably the slowest part of the process if you did it with loops.
You can use %%time at the beginning of the cell to measure its time of execution
```
%matplotlib inline
from pylab import *
import numpy as np
import scipy.ndimage as nd
import scipy.sparse as sp
# im = nd.imread("data/test.png")[:,:,:3]/255.
im = nd.imread("data/bear2.png")[:,:,:3]/255.
def Laplacian(im):
laplacian_kernel = np.array([[0, -1, 0], [-1, 4, -1], [0, -1, 0]])
#{‘reflect’,’constant’,’nearest’,’mirror’, ‘wrap’}
conv = nd.convolve(im, laplacian_kernel,mode='constant')
return conv
def add_border(im):
border_im = np.zeros((im.shape[0]+2,im.shape[1]+2,im.shape[2]))
border_im[1:-1,1:-1] = im
return border_im
im = add_border(im)
imshow(im)
def buildSystemSparse(im_shape):
sizey,sizex=im_shape
A = sp.eye(sizex*sizey, format="csr")
A=A*4.
A=A - sp.eye(sizex*sizey, k=1) - sp.eye(sizex*sizey, k = -1)
A=A - sp.eye(sizex*sizey, k=-sizex) - sp.eye(sizex*sizey, k = sizex)
# print A
return A
# this is done once for all three channels
A = buildSystemSparse(im[:,:,0].shape)
b = Laplacian(im[:,:,0])
red = sp.linalg.cgs(A, b.ravel())[0]
red = np.clip(red.reshape(b.shape),0.0,1.0)
b2 = Laplacian(im[:,:,1])
green = sp.linalg.cgs(A, b2.ravel())[0]
green = np.clip(green.reshape(b2.shape),0.0,1.0)
b3 = Laplacian(im[:,:,2])
blue = sp.linalg.cgs(A, b3.ravel())[0]
blue = np.clip(blue.reshape(b3.shape),0.0,1.0)
result =np.dstack((red,green,blue))
# The result should be the original image
imshow(result)
```
## Task 4: 2D Poisson Compositing (40)
This formulation is similar to equation 8 in [Pérez, et al.](http://cs.brown.edu/courses/cs129/asgn/proj2/resources/PoissonImageEditing.pdf) You can read more of that paper, especially the "Discrete Poisson Solver", if you want more guidance.
Esssentially you need to build A and b so it takes care of the boundary conditions using the mask. You can do it when you iterate to create the matrix A, or you can operate with the mask and the Laplacian to do it directly on with the images. In the second case, you probably have problems if your mask or composite does not have a border.
The test data has a standard easy boundary, you don't even have to move the image in order to do the composite.
The equations (A and b) for the pixels inside the pasted image (fg) will be the same as before, however the pixels at the boundary change.
If we look at the equation in Task 1 for 2D image
4*v(i,j) - v(i-1, j) - v(i+1, j) - v(i, j-1) - v(i, j+1) = 4*s(i,j) - s(i-1, j) - s(i+1, j) - s(i, j-1) - s(i, j+1)
Now some of the neighbours in the left side (unkowns) are actually know (values from the background image at the boundary. So these values need to be added to the right side.
When constructing A, you also want to have the mask into account, so you only have equations for the values that you want to compute. In convenience you can create the matrix of the size of the mask and leave the rows for the pixels not interested to 0. Of course also for the values of b.
For this task you can do a clean solution ignoring special cases of boundaries as follow.
Pasted | Poisson
- | -
 | 
HINT: You can do this using the mask (and its inverse) and the laplacian operator with the fg and bg
```
%matplotlib inline
from pylab import *
import imageio as imio
import numpy as np
import scipy.ndimage as nd
import scipy.sparse
import subprocess
import sys
from IPython.display import clear_output
mask = imio.imread("data/bear2_mask.png")[:,:,1]/255
mask_inv = 1.-mask
fg =imio.imread("data/bear2.png")[:,:,:3]/255.
bg =imio.imread("data/waterpool.png")[:,:,:3]/255.
off = [0,0]
def buildSystemSparse(im_shape,mask):
sizey,sizex=im_shape
A = sp.eye(sizex*sizey, format="csr")
A=A*4.
A=A - sp.eye(sizex*sizey, k=1) - sp.eye(sizex*sizey, k = -1)
A=A - sp.eye(sizex*sizey, k=-sizex) - sp.eye(sizex*sizey, k = sizex)
mask_r=mask.reshape(1,-1)
print mask_r.shape
i=0
last = 0
for row in A:
row=sp.csr_matrix(np.array(row.todense()) * np.array(mask_r))
A[i]=row
if(int(i*100./(sizey*sizex))>last):
last=i*100./(sizey*sizex)
clear_output()
sys.stdout.write(str(int(last)+1)+"%\n")
sys.stdout.flush()
i+=1
return A
A = buildSystemSparse(fg[:,:,0].shape,mask_inv)
b1 = Laplacian(fg[:,:,0])*mask_inv - Laplacian(bg[:160,:293,0]*mask)*mask_inv
print "x"
red = sp.linalg.cgs(A, b1.ravel())[0]
red = np.clip(red.reshape(b1.shape),0.0,1.0)
b2 = Laplacian(fg[:,:,1])*mask_inv - Laplacian(bg[:160,:293,1]*mask)*mask_inv
print "x"
green = sp.linalg.cgs(A, b2.ravel())[0]
green = np.clip(green.reshape(b2.shape),0.0,1.0)
b3 = Laplacian(fg[:,:,2])*mask_inv - Laplacian(bg[:160,:293,2]*mask)*mask_inv
print "x"
blue = sp.linalg.cgs(A, b3.ravel())[0]
blue = np.clip(blue.reshape(b3.shape),0.0,1.0)
result =np.dstack((red,green,blue))
print "x"
imshow(result)
fg=result
bg[off[0]:fg.shape[0]+off[0],off[1]:fg.shape[1]+off[1]] = bg[off[0]:fg.shape[0]+off[0],off[1]:fg.shape[1]+off[1]]*dstack((mask,mask,mask))+ fg*dstack((mask_inv,mask_inv,mask_inv))
imshow(bg)
imio.imwrite("data/pasted.png",bg)
```
## Extras : Improve compositing and test other images
Allow for moving the composite on other places and with your images, paste several things... This is easy to do, maybe tedious, but the results can be really fun.
## Extras : Other editings with gradients.
Gradient editing is very powerful, in the [paper](http://cs.brown.edu/courses/cs129/asgn/proj2/resources/PoissonImageEditing.pdf) you have some examples. Some ideas...
* Mixing Gradients to allow transparent images or images with holes. Instead of trying to adhere to the gradients of the source image, at each masked pixel use the largest gradient available in either the source or target. You can also try taking the average of gradients in the source and target.
* Color2Gray: Sometimes, in converting a color image to grayscale (e.g., when printing to a laser printer), we lose the important contrast information, making the image difficult to understand. For example, compare the color version of the image on right with its grayscale version produced by rgb2gray(). Can you do better than rgb2gray? Gradient-domain processing provides one avenue: create a gray image that has similar intensity to the rgb2gray output but has similar contrast to the original RGB image. This is an example of a tone-mapping problem, conceptually similar to that of converting HDR images to RGB displays. To get credit for this, show the grayscale image that you produce (the numbers should be easily readable). Hint: Try converting the image to HSV space and looking at the gradients in each channel. Then, approach it as a mixed gradients problem where you also want to preserve the grayscale intensity.
Color | Gray
- | -
 | 
* Automatically shifting the offset of the source image to decrease the difference of the images in the composite area.
* Implement and compare other blending techniques, such as Laplacian pyramid blending.
* Perform the blending on video frames instead of still images.
* Try other applications of gradient domain editing such as non-photorealistic rendering, edge enhancement, and texture transfer.
This Lab is an adaptation of this one http://cs.brown.edu/courses/cs129/asgn/proj2/
| github_jupyter |
# Realization of Non-Recursive Filters
*This jupyter notebook is part of a [collection of notebooks](../index.ipynb) on various topics of Digital Signal Processing. Please direct questions and suggestions to [Sascha.Spors@uni-rostock.de](mailto:Sascha.Spors@uni-rostock.de).*
## Fast Convolution
The straightforward convolution of two finite-length signals $x[k]$ and $h[k]$ is a numerically complex task. This has led to the development of various techniques with considerably lower complexity. The basic concept of the *fast convolution* is to exploit the correspondence between the convolution and the scalar multiplication in the frequency domain.
### Convolution of Finite-Length Signals
The convolution of a causal signal $x_L[k]$ of length $L$ with a causal impulse response $h_N[k]$ of length $N$ is given as
\begin{equation}
y[k] = x_L[k] * h_N[k] = \sum_{\kappa = 0}^{L-1} x_L[\kappa] \; h_N[k - \kappa] = \sum_{\kappa = 0}^{N-1} h_N[\kappa] \; x_L[k - \kappa]
\end{equation}
where $x_L[k] = 0$ for $k<0 \wedge k \geq L$ and $h_N[k] = 0$ for $k<0 \wedge k \geq N$. The resulting signal $y[k]$ is of finite length $M = N+L-1$. The computation of $y[k]$ for $k=0,1, \dots, M-1$ requires $M \cdot N$ multiplications and $M \cdot (N-1)$ additions. The computational complexity of the convolution is consequently [in the order of](https://en.wikipedia.org/wiki/Big_O_notation) $\mathcal{O}(M \cdot N)$. Discrete-time Fourier transformation (DTFT) of above relation yields
\begin{equation}
Y(e^{j \Omega}) = X_L(e^{j \Omega}) \cdot H_N(e^{j \Omega})
\end{equation}
Discarding the effort of transformation, the computationally complex convolution is replaced by a scalar multiplication with respect to the frequency $\Omega$. However, $\Omega$ is a continuous frequency variable which limits the numerical evaluation of this scalar multiplication. In practice, the DTFT is replaced by the discrete Fourier transformation (DFT). Two aspects have to be considered before a straightforward application of the DFT
1. The DFTs $X_L[\mu]$ and $H_N[\mu]$ are of length $L$ and $N$ respectively and cannot be multiplied straightforwardly
2. For $N = L$, the multiplication of the two spectra $X_L[\mu]$ and $H_L[\mu]$ would result in the [periodic/circular convolution](https://en.wikipedia.org/wiki/Circular_convolution) $x_L[k] \circledast h_L[k]$ due to the periodicity of the DFT. Since we aim at realizing the linear convolution $x_L[k] * h_N[k]$ with the DFT, special care has to be taken to avoid cyclic effects.
### Linear Convolution by Periodic Convolution
The periodic convolution of the two signals $x_L[k]$ and $h_N[k]$ is defined as
\begin{equation}
x_L[k] \circledast h_N[k] = \sum_{\kappa=0}^{M-1} \tilde{x}_M[k - \kappa] \; \tilde{h}_M[\kappa]
\end{equation}
where the periodic continuations $\tilde{x}_M[k]$ of $x_L[k]$ and $\tilde{h}_M[k]$ of $h_N[k]$ with period $M$ are given as
\begin{align}
\tilde{x}_M[k] &= \sum_{m = -\infty}^{\infty} x_L[m \cdot M + k] \\
\tilde{h}_M[k] &= \sum_{m = -\infty}^{\infty} h_N[m \cdot M + k]
\end{align}
The result of the circular convolution has a periodicity of $M$.
To compute the linear convolution by the periodic convolution one has to take care that the result of the linear convolution fits into one period of the periodic convolution. Hence, the periodicity has to be chosen as $M \geq N+L-1$. This can be achieved by zero-padding of $x_L[k]$ and $h_N[k]$ to a total length of $M$
\begin{align}
x_M[k] &= \begin{cases}
x_L[k] & \mathrm{for} \; k=0, 1, \dots, L-1 \\
0 & \mathrm{for} \; k=L, L+1, \dots, M-1
\end{cases}
\\
h_M[k] &= \begin{cases}
h_N[k] & \mathrm{for} \; k=0, 1, \dots, N-1 \\
0 & \mathrm{for} \; k=N, N+1, \dots, M-1
\end{cases}
\end{align}
This results in the desired equality of linear and periodic convolution
\begin{equation}
x_L[k] * h_N[k] = x_M[k] \circledast h_M[k]
\end{equation}
for $k = 0,1,\dots, M-1$ with $M = N+L-1$.
#### Example - Linear by periodic convolution
The following example computes the linear, periodic and linear by periodic convolution of a rectangular signal $x[k] = \text{rect}_L[k]$ of length $L$ with a triangular signal $h[k] = \Lambda_N[k]$ of length $N$.
```
%matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import scipy.signal as sig
L = 32 # length of signal x[k]
N = 16 # length of signal h[k]
M = 16 # periodicity of periodic convolution
def periodic_summation(x, N):
"Zero-padding to length N or periodic summation with period N."
M = len(x)
rows = int(np.ceil(M/N))
if (M < int(N*rows)):
x = np.pad(x, (0, int(N*rows-M)), 'constant')
x = np.reshape(x, (rows, N))
return np.sum(x, axis=0)
def periodic_convolve(x, y, P):
"Periodic convolution of two signals x and y with period P."
x = periodic_summation(x, P)
h = periodic_summation(y, P)
return np.array([np.dot(np.roll(x[::-1], k+1), h) for k in range(P)], float)
# generate signals
x = np.ones(L)
h = sig.triang(N)
# linear convolution
y1 = np.convolve(x, h, 'full')
# periodic convolution
y2 = periodic_convolve(x, h, M)
# linear convolution via periodic convolution
xp = np.append(x, np.zeros(N-1))
hp = np.append(h, np.zeros(L-1))
y3 = periodic_convolve(xp, hp, L+N-1)
# plot results
def plot_signal(x):
plt.figure(figsize = (10, 3))
plt.stem(x)
plt.xlabel(r'$k$')
plt.ylabel(r'$y[k]$')
plt.axis([0, N+L, 0, 1.1*x.max()])
plot_signal(x)
plt.title('Signal $x[k]$')
plot_signal(y1)
plt.title('Linear convolution')
plot_signal(y2)
plt.title('Periodic convolution with period M = %d' %M)
plot_signal(y3)
plt.title('Linear convolution by periodic convolution');
```
**Exercise**
* Change the lengths `L`, `N` and `M` and check how the results for the different convolutions change
### The Fast Convolution
Using the above derived equality of the linear and periodic convolution one can express the linear convolution $y[k] = x_L[k] * h_N[k]$ by the DFT as
\begin{equation}
y[k] = \text{IDFT}_M \{ \; \text{DFT}_M\{ x_M[k] \} \cdot \text{DFT}_M\{ h_M[k] \} \; \}
\end{equation}
This operation requires three DFTs of length $M$ and $M$ complex multiplications. On first sight this does not seem to be an improvement, since one DFT/IDFT requires $M^2$ complex multiplications and $M \cdot (M-1)$ complex additions. The overall numerical complexity is hence in the order of $\mathcal{O}(M^2)$. The DFT can be realized efficiently by the [fast Fourier transformation](https://en.wikipedia.org/wiki/Fast_Fourier_transform) (FFT), which lowers the computational complexity to $\mathcal{O}(M \log_2 M)$. The resulting algorithm is known as *fast convolution* due to its computational efficiency.
The fast convolution algorithm is composed of the following steps
1. Zero-padding of the two input signals $x_L[k]$ and $h_N[k]$ to at least a total length of $M \geq N+L-1$
2. Computation of the DFTs $X[\mu]$ and $H[\mu]$ using a FFT of length $M$
3. Multiplication of the spectra $Y[\mu] = X[\mu] \cdot H[\mu]$
4. Inverse DFT of $Y[\mu]$ using an inverse FFT of length $M$
The overall complexity depends on the particular implementation of the FFT. Many FFTs are most efficient for lengths which are a power of two. It therefore can make sense, in terms of computational complexity, to choose $M$ as a power of two instead of the shortest possible length $N+L-1$. For real valued signals $x[k] \in \mathbb{R}$ and $h[k] \in \mathbb{R}$ the computational complexity can be reduced significantly by using a real valued FFT.
#### Example - Fast convolution
The implementation of the fast convolution algorithm is straightforward. Most implementations of the FFT include zero-padding to a given length $M$, e.g in `numpy` by `numpy.fft.fft(x, M)`. In the following example an implementation of the fast convolution is shown. For illustration the convolution of a rectangular signal $x[k] = \text{rect}_L[k]$ of length $L$ with a triangular signal $h[k] = \Lambda_N[k]$ of length $N$ is considered.
```
L = 16 # length of signal x[k]
N = 16 # length of signal h[k]
M = N+L-1
# generate signals
x = np.ones(L)
h = sig.triang(N)
# linear convolution
y1 = np.convolve(x, h, 'full')
# fast convolution
y2 = np.fft.ifft(np.fft.fft(x, M) * np.fft.fft(h, M))
plt.figure(figsize=(10, 6))
plt.subplot(211)
plt.stem(y1)
plt.xlabel(r'$k$')
plt.ylabel(r'$y[k] = x_L[k] * h_N[k]$')
plt.title('Result of linear convolution')
plt.subplot(212)
plt.stem(y1)
plt.xlabel(r'$k$')
plt.ylabel(r'$y[k] = x_L[k] * h_N[k]$')
plt.title('Result of fast convolution')
plt.tight_layout()
```
#### Example - Numerical complexity
It was already argued that the numerical complexity of the fast convolution is considerably lower due to the usage of the FFT. The gain with respect to the convolution is evaluated in the following. In order to measure the execution times for both algorithms the `timeit` module is used. The algorithms are evaluated for the convolution of two random signals $x_L[k]$ and $h_N[k]$ of length $L=N=2^n$ for $n=0, 1, \dots, 16$.
```
import timeit
n = np.arange(17) # lengths = 2**n to evaluate
reps = 20 # number of repetitions for timeit
gain = np.zeros(len(n))
for N in n:
length = 2**N
# setup environment for timeit
tsetup = 'import numpy as np; from numpy.fft import rfft, irfft; \
x=np.random.randn(%d); h=np.random.randn(%d)' % (length, length)
# direct convolution
tc = timeit.timeit('np.convolve(x, x, mode="full")', setup=tsetup, number=reps)
# fast convolution
tf = timeit.timeit('irfft(rfft(x, %d) * rfft(h, %d))' % (2*length, 2*length), setup=tsetup, number=reps)
# speedup by using the fast convolution
gain[N] = tc/tf
# show the results
plt.figure(figsize = (15, 10))
plt.barh(n, gain, log=True)
plt.plot([1, 1], [-1, n[-1]+1], 'r-')
plt.yticks(n, 2**n)
plt.xlabel('Gain of fast convolution')
plt.ylabel('Length of signals')
plt.title('Comparison between direct/fast convolution')
plt.grid()
```
**Exercise**
* When is the fast convolution more efficient/faster than a direct convolution?
* Why is it slower below a given signal length?
* Is the trend of the gain as expected by the numerical complexity of the FFT?
Solution: The gain in execution time of a fast convolution over a direct implementation of the the convolution for different signal lengths depends heavily on the particular implementation and hardware used. The fast convolution in this example is faster for two signals having a length equal or larger than 1024 samples. Discarding the outliers and short lengths, the overall trend in the gain is approximately logarithmic as predicted above.
**Copyright**
This notebook is provided as [Open Educational Resource](https://en.wikipedia.org/wiki/Open_educational_resources). Feel free to use the notebook for your own purposes. The text is licensed under [Creative Commons Attribution 4.0](https://creativecommons.org/licenses/by/4.0/), the code of the IPython examples under the [MIT license](https://opensource.org/licenses/MIT). Please attribute the work as follows: *Sascha Spors, Digital Signal Processing - Lecture notes featuring computational examples, 2016-2018*.
| github_jupyter |
```
# This cell is added by sphinx-gallery
!pip install mrsimulator --quiet
%matplotlib inline
import mrsimulator
print(f'You are using mrsimulator v{mrsimulator.__version__}')
```
# Wollastonite, ²⁹Si (I=1/2)
²⁹Si (I=1/2) spinning sideband simulation.
Wollastonite is a high-temperature calcium-silicate,
$\beta−\text{Ca}_3\text{Si}_3\text{O}_9$, with three distinct
$^{29}\text{Si}$ sites. The $^{29}\text{Si}$ tensor parameters
were obtained from Hansen `et al.` [#f1]_
```
import matplotlib.pyplot as plt
from mrsimulator import Simulator, SpinSystem, Site
from mrsimulator.methods import BlochDecaySpectrum
from mrsimulator import signal_processing as sp
```
**Step 1:** Create the sites.
```
S29_1 = Site(
isotope="29Si",
isotropic_chemical_shift=-89.0, # in ppm
shielding_symmetric={"zeta": 59.8, "eta": 0.62}, # zeta in ppm
)
S29_2 = Site(
isotope="29Si",
isotropic_chemical_shift=-89.5, # in ppm
shielding_symmetric={"zeta": 52.1, "eta": 0.68}, # zeta in ppm
)
S29_3 = Site(
isotope="29Si",
isotropic_chemical_shift=-87.8, # in ppm
shielding_symmetric={"zeta": 69.4, "eta": 0.60}, # zeta in ppm
)
sites = [S29_1, S29_2, S29_3] # all sites
```
**Step 2:** Create the spin systems from these sites. Again, we create three
single-site spin systems for better performance.
```
spin_systems = [SpinSystem(sites=[s]) for s in sites]
```
**Step 3:** Create a Bloch decay spectrum method.
```
method = BlochDecaySpectrum(
channels=["29Si"],
magnetic_flux_density=14.1, # in T
rotor_frequency=1500, # in Hz
spectral_dimensions=[
{
"count": 2048,
"spectral_width": 25000, # in Hz
"reference_offset": -10000, # in Hz
"label": r"$^{29}$Si resonances",
}
],
)
# A graphical representation of the method object.
plt.figure(figsize=(4, 2))
method.plot()
plt.show()
```
**Step 4:** Create the Simulator object and add the method and spin system objects.
```
sim = Simulator()
sim.spin_systems += spin_systems # add the spin systems
sim.methods += [method] # add the method
```
**Step 5:** Simulate the spectrum.
```
sim.run()
# The plot of the simulation before signal processing.
plt.figure(figsize=(4.25, 3.0))
ax = plt.subplot(projection="csdm")
ax.plot(sim.methods[0].simulation.real, color="black", linewidth=1)
ax.invert_xaxis()
plt.tight_layout()
plt.show()
```
**Step 6:** Add post-simulation signal processing.
```
processor = sp.SignalProcessor(
operations=[sp.IFFT(), sp.apodization.Exponential(FWHM="70 Hz"), sp.FFT()]
)
processed_data = processor.apply_operations(data=sim.methods[0].simulation)
# The plot of the simulation after signal processing.
plt.figure(figsize=(4.25, 3.0))
ax = plt.subplot(projection="csdm")
ax.plot(processed_data.real, color="black", linewidth=1)
ax.invert_xaxis()
plt.tight_layout()
plt.show()
```
.. [#f1] Hansen, M. R., Jakobsen, H. J., Skibsted, J., $^{29}\text{Si}$
Chemical Shift Anisotropies in Calcium Silicates from High-Field
$^{29}\text{Si}$ MAS NMR Spectroscopy, Inorg. Chem. 2003,
**42**, *7*, 2368-2377.
`DOI: 10.1021/ic020647f <https://doi.org/10.1021/ic020647f>`_
| github_jupyter |
# Ingesting Data Into The Cloud
In this section, we will describe a typical scenario in which an application writes data into an Amazon S3 Data Lake and the data needs to be accessed by both the data science / machine learning team, as well as the business intelligence / data analyst team as shown in the figure below.
<img src="img/ingest_overview.png" width="80%" align="left">
As a **data scientist or machine learning engineer**, you want to have access to all of the raw data, and be able to quickly explore it. We will show you how to leverage **Amazon Athena** as an interactive query service to analyze data in Amazon S3 using standard SQL, without moving the data.
* In the first step, we will register the TSV data in our S3 bucket with Athena, and then run some ad-hoc queries on the dataset.
* We will also show how you can easily convert the TSV data into the more query-optimized, columnar file format Apache Parquet.
Your **business intelligence team and data analysts** might also want to have a subset of the data in a data warehouse which they can then transform, and query with their standard SQL clients to create reports and visualize trends. We will show you how to leverage **Amazon Redshift**, a fully managed data warehouse service, to
* insert TSV data into Amazon Redshift, but also be able to combine the data warehouse queries with the data that’s still in our S3 data lake via **Amazon Redshift Spectrum**.
* You can also use Amazon Redshift’s data lake export functionality to unload data back into our S3 data lake in Parquet file format.
# Amazon Customer Reviews Dataset
https://s3.amazonaws.com/amazon-reviews-pds/readme.html
### Dataset Columns:
- `marketplace`: 2-letter country code (in this case all "US").
- `customer_id`: Random identifier that can be used to aggregate reviews written by a single author.
- `review_id`: A unique ID for the review.
- `product_id`: The Amazon Standard Identification Number (ASIN). `http://www.amazon.com/dp/<ASIN>` links to the product's detail page.
- `product_parent`: The parent of that ASIN. Multiple ASINs (color or format variations of the same product) can roll up into a single parent.
- `product_title`: Title description of the product.
- `product_category`: Broad product category that can be used to group reviews (in this case digital videos).
- `star_rating`: The review's rating (1 to 5 stars).
- `helpful_votes`: Number of helpful votes for the review.
- `total_votes`: Number of total votes the review received.
- `vine`: Was the review written as part of the [Vine](https://www.amazon.com/gp/vine/help) program?
- `verified_purchase`: Was the review from a verified purchase?
- `review_headline`: The title of the review itself.
- `review_body`: The text of the review.
- `review_date`: The date the review was written.
# Release Resources
```
%%html
<p><b>Shutting down your kernel for this notebook to release resources.</b></p>
<button class="sm-command-button" data-commandlinker-command="kernelmenu:shutdown" style="display:none;">Shutdown Kernel</button>
<script>
try {
els = document.getElementsByClassName("sm-command-button");
els[0].click();
}
catch(err) {
// NoOp
}
</script>
%%javascript
try {
Jupyter.notebook.save_checkpoint();
Jupyter.notebook.session.delete();
}
catch(err) {
// NoOp
}
```
| github_jupyter |
<table class="ee-notebook-buttons" align="left">
<td><a target="_blank" href="https://github.com/giswqs/earthengine-py-notebooks/tree/master/NAIP/from_name.ipynb"><img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" /> View source on GitHub</a></td>
<td><a target="_blank" href="https://nbviewer.jupyter.org/github/giswqs/earthengine-py-notebooks/blob/master/NAIP/from_name.ipynb"><img width=26px src="https://upload.wikimedia.org/wikipedia/commons/thumb/3/38/Jupyter_logo.svg/883px-Jupyter_logo.svg.png" />Notebook Viewer</a></td>
<td><a target="_blank" href="https://mybinder.org/v2/gh/giswqs/earthengine-py-notebooks/master?filepath=NAIP/from_name.ipynb"><img width=58px src="https://mybinder.org/static/images/logo_social.png" />Run in binder</a></td>
<td><a target="_blank" href="https://colab.research.google.com/github/giswqs/earthengine-py-notebooks/blob/master/NAIP/from_name.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" /> Run in Google Colab</a></td>
</table>
## Install Earth Engine API
Install the [Earth Engine Python API](https://developers.google.com/earth-engine/python_install) and [geehydro](https://github.com/giswqs/geehydro). The **geehydro** Python package builds on the [folium](https://github.com/python-visualization/folium) package and implements several methods for displaying Earth Engine data layers, such as `Map.addLayer()`, `Map.setCenter()`, `Map.centerObject()`, and `Map.setOptions()`.
The magic command `%%capture` can be used to hide output from a specific cell.
```
# %%capture
# !pip install earthengine-api
# !pip install geehydro
```
Import libraries
```
import ee
import folium
import geehydro
```
Authenticate and initialize Earth Engine API. You only need to authenticate the Earth Engine API once. Uncomment the line `ee.Authenticate()`
if you are running this notebook for this first time or if you are getting an authentication error.
```
# ee.Authenticate()
ee.Initialize()
```
## Create an interactive map
This step creates an interactive map using [folium](https://github.com/python-visualization/folium). The default basemap is the OpenStreetMap. Additional basemaps can be added using the `Map.setOptions()` function.
The optional basemaps can be `ROADMAP`, `SATELLITE`, `HYBRID`, `TERRAIN`, or `ESRI`.
```
Map = folium.Map(location=[40, -100], zoom_start=4)
Map.setOptions('HYBRID')
```
## Add Earth Engine Python script
```
image = ee.Image('USDA/NAIP/DOQQ/m_3712213_sw_10_1_20140613')
Map.setCenter(-122.466123, 37.769833, 17)
Map.addLayer(image, {'bands': ['N', 'R','G']}, 'NAIP')
```
## Display Earth Engine data layers
```
Map.setControlVisibility(layerControl=True, fullscreenControl=True, latLngPopup=True)
Map
```
| github_jupyter |
# Bloom filters in Ethereum
### Requirements
Python3
$ pip install ethereum
$ pip install rlp
$ pip install jupyter
$ pip install eth_abi
```
# Install a pip package in the current Jupyter kernel
import sys
!{sys.executable} -m pip install ethereum
!{sys.executable} -m pip install ethereum
!{sys.executable} -m pip install eth_abi
!{sys.executable} -m pip install rlp
# import abi
from ethereum import bloom, utils
import eth_abi
import rlp
```
We define a few functions that will be useful for us, the first one decoding an integer from a hexadecimal and the second one encoding a hex from an int
We can visualize an "empty" bloom filter as a bitmap of all 0s of length 512
```
# A "normal" bloom filter start, a bitmap with all 0s
log_bloom = utils.encode_hex(utils.zpad(utils.int_to_big_endian(0), 256))
print(log_bloom)
print(len(log_bloom))
```
Now we add an element to the bloom filter, we will add an address to it
```
encoded_address = utils.decode_hex(utils.remove_0x_head('0x864Be2775d392787D5fa37ee1DB45FE0b1B3D1FC'))
# Add an address ot the bloom filter
b = bloom.bloom_insert(0, encoded_address)
# let's see what we have in hex
print(bloom.b64(b))
# Turn it into a hex string
print(utils.encode_hex(bloom.b64(b)))
```
We see that we have 3 entries in the logbloom. How are they determined?
We take the hash of the address (keccack256), take the low order 9 bits of the first three double bytes (2 hex numbers) of the hash digest. The numbers that come out are the positions of the bits.
Example:
bloom(0f572e5295c57f15886f9b263e2f6d2d6c7b5ec6)
sha3: bd2b01afcd27800b54d2179edc49e2bffde5078bb6d0b204694169b1643fb108
first double-bytes: bd2b, 01af, cd27 -- which leads to bits in bloom --> 299, 431, 295
We can check very easily if the address that we passed in is in the set
```
print(bloom.bloom_query(b, encoded_address))
```
Let's try another one that is not in the set
```
other_address = encoded_address = utils.decode_hex(utils.remove_0x_head('0x0f572e5295c57f15886f9b263e2f6d2d6c7b5ec6'))
print(bloom.bloom_query(b, encoded_address))
new_b = bloom.bloom_insert(b, 1);
print(bloom.bloom_query(new_b, '1'))
print(bloom.bloom_query(new_b, '2'))
```
What does bloom query do? Takes in the value and adds it to a new 0 bloom and compares the 2 blooms
```
bloom2 = bloom.bloom_insert(0, '2')
print((new_b & bloom2) == bloom2)
bloom3 = bloom.bloom_insert(0, '1')
print((new_b & bloom3) == bloom3)
```
Now let's look at a bloom filter in action for receipts in ethereum. We first use the abi to be able to encode the values. The log entry consists of one topic and a data field. We notice that we have 2 indexed inputs: the to and from address, while the value is not.
```
abi = {
"anonymous": "false",
"inputs": [
{
"indexed": True,
"name": "from",
"type": "address"
},
{
"indexed": True,
"name": "to",
"type": "address"
},
{
"indexed": False,
"name": "value",
"type": "uint256"
}
],
"name": "Transfer",
"type": "event"
}
```
We add a real world example transaction receipt, one where a user transferred tokens from one address to the other.
```
example_receipt = {
"jsonrpc": "2.0",
"id": 1,
"result": {
"blockHash": "0xc57ff9020f066420198584aafc2944c8abaac1038e56a3f5a347bbd199111956",
"blockNumber": "0x54c0fb",
"contractAddress": "null",
"cumulativeGasUsed": "0x684a0c",
"from": "0xcc56dcc36d43341c074f0fc06aec3211cd8f8f44",
"gasUsed": "0x92c3",
"logs": [{
"address": "0xea38eaa3c86c8f9b751533ba2e562deb9acded40",
"topics": ["0xddf252ad1be2c89b69c2b068fc378daa952ba7f163c4a11628f55a4df523b3ef", "0x000000000000000000000000cc56dcc36d43341c074f0fc06aec3211cd8f8f44", "0x0000000000000000000000001c4b70a3968436b9a0a9cf5205c787eb81bb558c"],
"data": "0x000000000000000000000000000000000000000000000cb28be99bb554a80000",
"blockNumber": "0x54c0fb",
"transactionHash": "0x865edf70c0e4b9860a6fe3af62f095ad7f9d3d881ab5ab4dfe3cf8fcead8c843",
"transactionIndex": "0x3b",
"blockHash": "0xc57ff9020f066420198584aafc2944c8abaac1038e56a3f5a347bbd199111956",
"logIndex": "0x9e",
"removed": "false"
}],
"logsBloom": "0x00000000000000000000000000000000000000000000000000000000000000100000008000000000000000000100000000000000000000000000000000000000000000000000000000000008001000000000000000000000000000000000000000000000000000000000000000000000200000000000000000000010000000000000000000002000000000000000000000000000000000000000000000000000000000000000000000000000000000000002000000000000000000000000000000000002000000000000000000000000000000000000000000000000004000000000000000000000000080000000000000000000000000000000000000000000",
"status": "0x1",
"to": "0xea38eaa3c86c8f9b751533ba2e562deb9acded40",
"transactionHash": "0x865edf70c0e4b9860a6fe3af62f095ad7f9d3d881ab5ab4dfe3cf8fcead8c843",
"transactionIndex": "0x3b"
}
}
```
Under "topics", we see that we have an array with 3 different elements, one is a hash, and others seems to be encoded data
```
topics = example_receipt["result"]["logs"][0]["topics"]
print(topics)
```
The first field corresponds to the hash ot the event
```
event_hash = '0x' + utils.encode_hex(utils.sha3('Transfer(address,address,uint256)'))
print(event_hash == topics[0])
```
Now let's retrieve the values that are indexed and not indexed from the abi
```
types = [i['type'] for i in abi['inputs'] if not i['indexed']]
print(types)
```
The unindexed type is the value field and it's actual value is located in the data field of the receipt
```
logs = example_receipt["result"]["logs"]
values = eth_abi.decode_abi(types, logs[0]["data"])
print(values)
```
Now we do the same with the indexed event variables, but now their value is stored in the topics array that we saw earlier
```
# indexed ones
indexed_types = [i['type'] for i in abi['inputs'] if i['indexed']]
indexed_names = [i['name'] for i in abi['inputs'] if i['indexed']]
indexed_values = [eth_abi.decode_single(t, v) for t, v in zip(indexed_types, logs[0]['topics'][1:])]
print(indexed_names, indexed_values)
```
We then bring it all together into the originial event that happened:
```
event_info = {
"from": indexed_values[0],
"to": indexed_values[1],
"value": values[0]
}
print(event_info)
# https://hur.st/bloomfilter/?n=3&p=1.0E-9&m=256&k=
```
| github_jupyter |
# Built-in Data Structures
We have seen Python's simple types: ``int``, ``float``, ``complex``, ``bool``, ``str``, etc.
Python also has several built-in compound types, which act as containers for other types.
These compound types are:
| Type Name | Example |Description |
|-----------|---------------------------|---------------------------------|
| ``list`` | ``[1, 2, 3]`` | ordered collection |
| ``tuple`` | ``(1, 2, 3)`` | immutable ordered collection |
| ``dict`` | ``{'a':1, 'b':2, 'c':3}`` | unordered (key,value) mapping |
| ``set`` | ``{1, 2, 3}`` | unordered collection |
As you can see, round, square, and curly brackets have distinct meanings when it comes to the type of collection produced.
We'll take a quick tour of these data structures here.
## Lists
Lists are the basic *ordered* and *mutable* data collection type in Python.
They can be defined with comma-separated values between square brackets; for example, here is a list of the first several prime numbers
```
L = [2, 3, 5, 7]
```
Lists have a number of useful properties and methods available to them.
Here we'll take a quick look at some of the more common and useful ones:
```
# Length of a list
len(L)
# Append a value to the end
L.append(11)
L
# Addition concatenates lists
L + [13, 17, 19]
# sort() method sorts in-place
L = [2, 5, 1, 6, 3, 4]
L.sort()
L
```
In addition there are many more built-in list methods; they are well-covered in Python's [online documentation](https://docs.python.org/3/tutorial/datastructures.html).
While we've been demonstrating lists containing values of a single type, one of the powerful features of Python's compound objects is that they can contain objects of *any* type, or even a mix of types. For example:
```
L = [1, 'two', 3.14, [0, 3, 5]]
```
This flexibility is a consequence of Python's dynamic type system.
Creating such a mixed sequence in a statically-typed language like C can be much more of a headache!
We see that lists can even contain other lists as elements.
Such type flexibility is an essential piece of what makes Python code relatively quick and easy to write.
So far we've been considering manipulations of lists as a whole; another essential piece is the accessing of individual elements.
This is done in Python via *indexing* and *slicing*, which we'll explore next.
### List Indexing and Slicing
Python provides access to elements in compound types through *indexing* for single elements, and *slicing* for multiple elements.
As we'll see, both are indicated by a square-bracket syntax.
Suppose we return to our list of the first several primes:
```
L = [2, 3, 5, 7, 11]
```
Python uses *zero-based* indexing, so we can access the first and second element in using the following syntax:
```
L[0]
L[1]
```
Elements at the end of the list can be accessed with negative numbers, starting from -1:
```
L[-1]
L[-2]
```
You can visualize this indexing scheme this way:

Here values in the list are represented by large numbers in the squares; list indices are represented by small numbers above and below.
In this case ``L[2]`` returns ``5``, because that is the next value at index ``2``.
Where *indexing* is a means of fetching a single value from the list, *slicing* is a means of accessing multiple values in sub-lists.
It uses a colon to indicate the start point (inclusive) and end point (non-inclusive) of the sub-array.
For example, to get the first three elements of the list, we can write
```
L[0:3]
```
Notice where ``0`` and ``3`` lie in the above diagram, and how the slice takes just the values between the indices.
If we leave out the first index, ``0`` is assumed, so we can equivalently write
```
L[:3]
```
Similarly, if we leave out the last index, it defaults to the length of the list.
Thus the last three elements can be accessed as follows
```
L[-3:]
```
Finally, it is possible to specify a third integer which represents the step size; for example, to select every second element of the list, we can write:
```
L[::2] # equivalent to L[0:len(L):2]
```
A particularly useful version of this is to specify a negative step, which will reverse the array:
```
L[::-1]
```
Both indexing and slicing can be used to set elements as well as access them.
The syntax is as you would expect:
```
L[0] = 100
print(L)
L[1:3] = [55, 56]
print(L)
```
A very similar slicing syntax is also used in many data science oriented packages, including NumPy and Pandas (mentioned in the introduction).
Now that we have seen Python lists and how to access elements in ordered compound types, let's take a look at the other three standard compound data types mentioned above.
## Tuples
Tuples are in many ways similar to lists, but they are defined with parentheses rather than square brackets:
```
t = (1, 2, 3)
```
They can also be defined without any brackets at all:
```
t = 1, 2, 3
print(t)
```
Like the lists discussed above, tuples have a length, and individual elements can be extracted using square-bracket indexing:
```
len(t)
t[0]
```
The main distinguishing feature of tuples is that they are **immutable**: this means that once they are created, their size and contents cannot be changed:
```
t[1] = 4
t.append(4)
```
Tuples are often used in a Python program; a particularly common case is in functions which have multiple return values.
For example, the ``as_integer_ratio()`` method of floating-point objects returns a numerator and a denominator; this dual return value comes in the form of a tuple:
```
x = 0.125
x.as_integer_ratio()
```
These multiple return values can be individually assigned as follows:
```
numerator, denominator = x.as_integer_ratio()
print(numerator / denominator)
```
The indexing and slicing logic covered above for lists works for tuples as well, along with a host of other methods.
Refer to the online Python documentation for a more complete list of these.
## Dictionaries
Dictionaries are extremely flexible mappings of keys to values, and form the basis of much of Python's internal implementation.
They can be created via a comma-separated list of ``key:value`` pairs within curly braces:
```
numbers = {'one':1, 'two':2, 'three':3}
```
Items are accessed and set via the indexing syntax used for lists and tuples, except here the index is not a zero-based order but valid key in the dictionary:
```
# Access a value via the key
numbers['two']
```
New items can be added to the dictionary using indexing as well:
```
# Set a new key, value pair
numbers['ninety'] = 90
print(numbers)
```
Keep in mind that dictionaries do not maintain any sense of order for the input parameters; this is by design.
This lack of ordering allows dictionaries to be implemented very efficiently, so that random element access is very fast, regardless of the size of the dictionary (if you're curious how this works, read about the concept of a *hash table*).
The [python documentation](https://docs.python.org/3/library/stdtypes.html) has a complete list of the methods available for dictionaries.
## Sets
The fourth basic collection is the set, which contains unordered collections of unique items.
They are defined much like lists and tuples, except they use the curly brackets of dictionaries:
```
primes = {2, 3, 5, 7}
odds = {1, 3, 5, 7, 9}
```
If you're familiar with the mathematics of sets, you'll be familiar with operations like the union, intersection, difference, symmetric difference, and others.
Python's sets have all of these operations built-in, via methods or operators.
For each, we'll show the two equivalent methods:
```
# union: items appearing in either
primes | odds # with an operator
primes.union(odds) # equivalently with a method
# intersection: items appearing in both
primes & odds # with an operator
primes.intersection(odds) # equivalently with a method
# difference: items in primes but not in odds
primes - odds # with an operator
primes.difference(odds) # equivalently with a method
# symmetric difference: items appearing in only one set
primes ^ odds # with an operator
primes.symmetric_difference(odds) # equivalently with a method
```
Many more set methods and operations are available.
You've probably already guessed what I'll say next: refer to Python's [online documentation](https://docs.python.org/3/library/stdtypes.html) for a complete reference.
## More Specialized Data Structures
Python contains several other data structures which you might find useful; these can generally be found in the built-in ``collections`` module.
The collections module is fully-documented in Python's online documentation, and you can read more about the various objects available there.
In particular, I've found the following very useful on occasion:
- ``collections.namedtuple``: like a tuple, but each value has a name
- ``collections.defaultdict``: like a dictionary, but unspecified keys have a user-specified default value
- ``collections.OrderedDict``: like a dictionary, but the order of keys is maintained
Once you've seen the standard built-in collection types, the use of these extended functionalities is very intuitive, and I'd suggest reading about their use.
| github_jupyter |
This is a `bqplot` recreation of Mike Bostock's [Wealth of Nations](https://bost.ocks.org/mike/nations/). This was also done by [Gapminder](http://www.gapminder.org/world/#$majorMode=chart$is;shi=t;ly=2003;lb=f;il=t;fs=11;al=30;stl=t;st=t;nsl=t;se=t$wst;tts=C$ts;sp=5.59290322580644;ti=2013$zpv;v=0$inc_x;mmid=XCOORDS;iid=phAwcNAVuyj1jiMAkmq1iMg;by=ind$inc_y;mmid=YCOORDS;iid=phAwcNAVuyj2tPLxKvvnNPA;by=ind$inc_s;uniValue=8.21;iid=phAwcNAVuyj0XOoBL_n5tAQ;by=ind$inc_c;uniValue=255;gid=CATID0;by=grp$map_x;scale=log;dataMin=194;dataMax=96846$map_y;scale=lin;dataMin=23;dataMax=86$map_s;sma=49;smi=2.65$cd;bd=0$inds=;modified=60). It is originally based on a TED Talk by [Hans Rosling](http://www.ted.com/talks/hans_rosling_shows_the_best_stats_you_ve_ever_seen).
```
import pandas as pd
import numpy as np
import os
from bqplot import (
LogScale, LinearScale, OrdinalColorScale, ColorAxis,
Axis, Scatter, Lines, CATEGORY10, Label, Figure, Tooltip
)
from ipywidgets import HBox, VBox, IntSlider, Play, jslink
initial_year = 1800
```
#### Cleaning and Formatting JSON Data
```
data = pd.read_json(os.path.abspath('./nations.json'))
def clean_data(data):
for column in ['income', 'lifeExpectancy', 'population']:
data = data.drop(data[data[column].apply(len) <= 4].index)
return data
def extrap_interp(data):
data = np.array(data)
x_range = np.arange(1800, 2009, 1.)
y_range = np.interp(x_range, data[:, 0], data[:, 1])
return y_range
def extrap_data(data):
for column in ['income', 'lifeExpectancy', 'population']:
data[column] = data[column].apply(extrap_interp)
return data
data = clean_data(data)
data = extrap_data(data)
income_min, income_max = np.min(data['income'].apply(np.min)), np.max(data['income'].apply(np.max))
life_exp_min, life_exp_max = np.min(data['lifeExpectancy'].apply(np.min)), np.max(data['lifeExpectancy'].apply(np.max))
pop_min, pop_max = np.min(data['population'].apply(np.min)), np.max(data['population'].apply(np.max))
def get_data(year):
year_index = year - 1800
income = data['income'].apply(lambda x: x[year_index])
life_exp = data['lifeExpectancy'].apply(lambda x: x[year_index])
pop = data['population'].apply(lambda x: x[year_index])
return income, life_exp, pop
```
#### Creating the Tooltip to display the required fields
`bqplot`'s native `Tooltip` allows us to simply display the data fields we require on a mouse-interaction.
```
tt = Tooltip(fields=['name', 'x', 'y'], labels=['Country Name', 'Income per Capita', 'Life Expectancy'])
```
#### Creating the Label to display the year
Staying true to the `d3` recreation of the talk, we place a `Label` widget in the bottom-right of the `Figure` (it inherits the `Figure` co-ordinates when no scale is passed to it). With `enable_move` set to `True`, the `Label` can be dragged around.
```
year_label = Label(x=[0.75], y=[0.10], font_size=52, font_weight='bolder', colors=['orange'],
text=[str(initial_year)], enable_move=True)
```
#### Defining Axes and Scales
The inherent skewness of the income data favors the use of a `LogScale`. Also, since the color coding by regions does not follow an ordering, we use the `OrdinalColorScale`.
```
x_sc = LogScale(min=income_min, max=income_max)
y_sc = LinearScale(min=life_exp_min, max=life_exp_max)
c_sc = OrdinalColorScale(domain=data['region'].unique().tolist(), colors=CATEGORY10[:6])
size_sc = LinearScale(min=pop_min, max=pop_max)
ax_y = Axis(label='Life Expectancy', scale=y_sc, orientation='vertical', side='left', grid_lines='solid')
ax_x = Axis(label='Income per Capita', scale=x_sc, grid_lines='solid')
```
#### Creating the Scatter Mark with the appropriate size and color parameters passed
To generate the appropriate graph, we need to pass the population of the country to the `size` attribute and its region to the `color` attribute.
```
# Start with the first year's data
cap_income, life_exp, pop = get_data(initial_year)
wealth_scat = Scatter(x=cap_income, y=life_exp, color=data['region'], size=pop,
names=data['name'], display_names=False,
scales={'x': x_sc, 'y': y_sc, 'color': c_sc, 'size': size_sc},
default_size=4112, tooltip=tt, animate=True, stroke='Black',
unhovered_style={'opacity': 0.5})
nation_line = Lines(x=data['income'][0], y=data['lifeExpectancy'][0], colors=['Gray'],
scales={'x': x_sc, 'y': y_sc}, visible=False)
```
#### Creating the Figure
```
time_interval = 10
fig = Figure(marks=[wealth_scat, year_label, nation_line], axes=[ax_x, ax_y],
title='Health and Wealth of Nations', animation_duration=time_interval)
```
#### Using a Slider to allow the user to change the year and a button for animation
Here we see how we can seamlessly integrate `bqplot` into the jupyter widget infrastructure.
```
year_slider = IntSlider(min=1800, max=2008, step=1, description='Year', value=initial_year)
```
When the `hovered_point` of the `Scatter` plot is changed (i.e. when the user hovers over a different element), the entire path of that country is displayed by making the `Lines` object visible and setting it's `x` and `y` attributes.
```
def hover_changed(change):
if change.new is not None:
nation_line.x = data['income'][change.new + 1]
nation_line.y = data['lifeExpectancy'][change.new + 1]
nation_line.visible = True
else:
nation_line.visible = False
wealth_scat.observe(hover_changed, 'hovered_point')
```
On the slider value `callback` (a function that is triggered everytime the `value` of the slider is changed) we change the `x`, `y` and `size` co-ordinates of the `Scatter`. We also update the `text` of the `Label` to reflect the current year.
```
def year_changed(change):
wealth_scat.x, wealth_scat.y, wealth_scat.size = get_data(year_slider.value)
year_label.text = [str(year_slider.value)]
year_slider.observe(year_changed, 'value')
```
#### Add an animation button
```
play_button = Play(min=1800, max=2008, interval=time_interval)
jslink((play_button, 'value'), (year_slider, 'value'))
```
#### Displaying the GUI
```
VBox([HBox([play_button, year_slider]), fig])
```
| github_jupyter |
# K-Nearest Neighbor Classifier with MinMaxScaler
This Code template is for the Classification task using a simple KNeighborsClassifier based on the K-Nearest Neighbors algorithm and feature rescaling technique MinMaxScaler in a pipeline.
### **Required Packages**
```
import warnings
import numpy as np
import pandas as pd
import seaborn as se
import matplotlib.pyplot as plt
from sklearn.pipeline import make_pipeline
from sklearn.model_selection import train_test_split
from sklearn.neighbors import KNeighborsClassifier
from imblearn.over_sampling import RandomOverSampler
from sklearn.preprocessing import LabelEncoder,MinMaxScaler
from sklearn.metrics import classification_report,plot_confusion_matrix
warnings.filterwarnings('ignore')
```
### **Initialization**
Filepath of CSV file
```
file_path= ""
```
List of features which are required for model training .
```
features = []
```
Target feature for prediction.
```
target = ''
```
### **Dataset Overview**
Pandas is an open-source, BSD-licensed library providing high-performance, easy-to-use data manipulation and data analysis tools.
We will use panda's library to read the CSV file using its storage path.And we use the head function to display the initial row or entry.
```
df=pd.read_csv(file_path)
df.head()
```
### **Dataset Information**
Print a concise summary of a DataFrame.
We will use info() method to print the information about the DataFrame including the index dtype and columns, non-null values and memory usage.
```
df.info()
```
### **Dataset Describe**
Generate descriptive statistics.
Descriptive statistics include those that summarize the central tendency, dispersion and shape of a dataset’s distribution, excluding NaN values.
We will analyzes both numeric and object series, as well as DataFrame column sets of mixed data types.
```
df.describe()
```
### **Feature Selection**
It is the process of reducing the number of input variables when developing a predictive model. Used to reduce the number of input variables to both reduce the computational cost of modelling and, in some cases, to improve the performance of the model.
We will assign all the required input features to X and target/outcome to Y.
```
X=df[features]
Y=df[target]
```
### **Data Preprocessing**
Since we do not know what is the number of Null values in each column.So,we print the columns arranged in descreasnig orde
```
print(df.isnull().sum().sort_values(ascending=False))
```
Since the majority of the machine learning models in the Sklearn library doesn't handle string category data and Null value, we have to explicitly remove or replace null values. The below snippet have functions, which removes the null value if any exists. And convert the string classes data in the datasets by encoding them to integer classes.
```
def NullClearner(df):
if(isinstance(df, pd.Series) and (df.dtype in ["float64","int64"])):
df.fillna(df.mean(),inplace=True)
return df
elif(isinstance(df, pd.Series)):
df.fillna(df.mode()[0],inplace=True)
return df
else:return df
def EncodeX(df):
return pd.get_dummies(df)
def EncodeY(df):
if len(df.unique())<=2:
return df
else:
un_EncodedT=np.sort(pd.unique(df), axis=-1, kind='mergesort')
df=LabelEncoder().fit_transform(df)
EncodedT=[xi for xi in range(len(un_EncodedT))]
print("Encoded Target: {} to {}".format(un_EncodedT,EncodedT))
return df
```
Calling preprocessing functions on the feature and target set.
```
x=X.columns.to_list()
for i in x:
X[i]=NullClearner(X[i])
X=EncodeX(X)
Y=EncodeY(NullClearner(Y))
X.head()
```
#### **Correlation Map**
In order to check the correlation between the features, we will plot a correlation matrix. It is effective in summarizing a large amount of data where the goal is to see patterns.
```
f,ax = plt.subplots(figsize=(18, 18))
matrix = np.triu(X.corr())
se.heatmap(X.corr(), annot=True, linewidths=.5, fmt= '.1f',ax=ax, mask=matrix)
plt.show()
```
#### **Distribution of Target Variable**
```
plt.figure(figsize = (10,6))
se.countplot(Y)
```
## **Data Splitting**
The train-test split is a procedure for evaluating the performance of an algorithm. The procedure involves taking a dataset and dividing it into two subsets. The first subset is utilized to fit/train the model. The second subset is used for prediction. The main motive is to estimate the performance of the model on new data.
```
x_train,x_test,y_train,y_test=train_test_split(X,Y,test_size=0.2,random_state=78)
```
#### **Handling Target Imbalance**
The challenge of working with imbalanced datasets is that most machine learning techniques will ignore, and in turn have poor performance on, the minority class, although typically it is performance on the minority class that is most important.
One approach to addressing imbalanced datasets is to oversample the minority class. The simplest approach involves duplicating examples in the minority class.We will perform overspampling using imblearn library.
```
x_train,y_train = RandomOverSampler(random_state=78).fit_resample(x_train, y_train)
```
### **Model**
KNN is one of the easiest Machine Learning algorithms based on Supervised Machine Learning technique. The algorithm stores all the available data and classifies a new data point based on the similarity. It assumes the similarity between the new data and data and put the new case into the category that is most similar to the available categories.KNN algorithm at the training phase just stores the dataset and when it gets new data, then it classifies that data into a category that is much similar to the available data.
**Model Tuning Parameters**
> * **n_neighbors** -> Number of neighbors to use by default for kneighbors queries.
> * **weights** -> weight function used in prediction. {uniform,distance}
> * **algorithm**-> Algorithm used to compute the nearest neighbors. {‘auto’, ‘ball_tree’, ‘kd_tree’, ‘brute’}
> * **p** -> Power parameter for the Minkowski metric. When p = 1, this is equivalent to using manhattan_distance (l1), and euclidean_distance (l2) for p = 2. For arbitrary p, minkowski_distance (l_p) is used.
> * **leaf_size** -> Leaf size passed to BallTree or KDTree. This can affect the speed of the construction and query, as well as the memory required to store the tree. The optimal value depends on the nature of the problem.
### **MinMaxScaler**
As Standardization of a dataset is a common requirement for many machine learning estimators,we will use MinMaxScaler for that purpose.
It transforms features by scaling each feature to a given range.
This estimator scales and translates each feature individually such that it is in the given range on the training set.
MinMaxScaler scales all the data features in the range [0, 1] or else in the range [-1, 1] if there are negative values in the dataset. This scaling compresses all the inliers in the narrow range [0, 0.005].
In the presence of outliers
```
model=make_pipeline(MinMaxScaler(),KNeighborsClassifier(n_jobs=-1))
model.fit(x_train,y_train)
```
#### **Model Accuracy**
score() method return the mean accuracy on the given test data and labels.
In multi-label classification, this is the subset accuracy which is a harsh metric since you require for each sample that each label set be correctly predicted.
```
print("Accuracy score {:.2f} %\n".format(model.score(x_test,y_test)*100))
```
#### **Confusion Matrix**
A confusion matrix is utilized to understand the performance of the classification model or algorithm in machine learning for a given test set where results are known.
```
plot_confusion_matrix(model,x_test,y_test,cmap=plt.cm.Reds)
```
#### **Classification Report**
A Classification report is used to measure the quality of predictions from a classification algorithm. How many predictions are True, how many are False.
where:
> * Precision:- Accuracy of positive predictions.
> * Recall:- Fraction of positives that were correctly identified.
> * f1-score:- percent of positive predictions were correct
> * support:- Support is the number of actual occurrences of the class in the specified dataset.
```
print(classification_report(y_test,model.predict(x_test)))
```
##### Creator:Prateek Kumar ,Github [Profile](https://github.com/pdpandey26)
| github_jupyter |
# Predict BMI
This script shows a real world example using BPt to study the relationship between BMI and the brain. The data used in this notebook cannot be made public as it is from the ABCD Study, which requires a data use agreement in order to use the data.
This notebook covers a number of different topics:
- Preparing Data
- Evaluating a single pipeline
- Considering different options for how to use a test set
- Introduce and use the Evaluate input option
```
import pandas as pd
import BPt as bp
import numpy as np
# Don't show sklearn convergence warnings
from warnings import simplefilter
from sklearn.exceptions import ConvergenceWarning
simplefilter("ignore", category=ConvergenceWarning)
# Display tables up to five decimals
pd.options.display.float_format = "{:,.5f}".format
```
## Preparing Data
We will first load in the underlying dataset for this project which has been saved as a csv. It contains multi-modal change in ROI data from two timepoints of the ABCD Study (difference from follow up and baseline).
This saved dataset doesn't include the real family ids, but an interesting piece of the ABCD study derived data is that there are a number of subjects from the same family. We will handle that in this example (granted with a fake family structure which we will generate below) by ensuring that for any cross-validation split, members of the same family stay in the same training or testing fold.
```
data = pd.read_excel('data/structure_base.xlsx')
list(data)[:10]
```
This dataset contains a number of columns we don't need. We will use the next cell to both group variables of interest together, and then select only the relvant columns to keep.
```
# Our target variable
targets = ['b_bmi']
# Columns with different traditional 'co-variates'.
covars = ['b_sex', 'b_demo_highest_education_categories','b_race_ethnicity_categories',
'b_agemos', 'b_mri_info_deviceserialnumber']
# Let's also note which of these are categorical
cat_covars = ['b_mri_info_deviceserialnumber',
'b_demo_highest_education_categories',
'b_race_ethnicity_categories',
'b_sex']
# These variables are any which we might want to use
# but not directly as input features! E.g., we
# might want to use them to inform choice of cross-validation.
non_input = ['b_rel_family_id']
# The different imaging features
thick = [d for d in list(data) if 'thick' in d]
area = [d for d in list(data) if 'smri_area_cort' in d]
subcort = [d for d in list(data) if 'smri_vol' in d]
dti_fa = [d for d in list(data) if 'dmri_dti_full_fa_' in d]
dti_md = [d for d in list(data) if 'dmri_dti_full_md_' in d]
brain = thick + area + subcort + dti_fa + dti_md
# All to keep
to_keep = brain + targets + covars + non_input
data = data[to_keep]
data.shape
```
Now let's convert from a pandas DataFrame to a BPt Dataset.
```
data = bp.Dataset(data)
# This is optional, to print some extra statements.
data.verbose = 1
data.shape
```
Next, we perform some actions specific to the Dataset class. These include specifying which columns are 'target' and 'non input', with any we don't set to one these roles treated as the default role, 'data'.
```
# Set's targets to target role
data = data.set_role(targets, 'target')
# Set non input to non input role
data = data.set_role(non_input, 'non input')
# Drop any missing values in the target variable
data = data.drop_subjects_by_nan(scope='target')
# We can optional add the categories we made as scopes!
data.add_scope(covars, 'covars', inplace=True)
data.add_scope(cat_covars, 'cat covars', inplace=True)
data.add_scope(thick, 'thick', inplace=True)
data.add_scope(area, 'area', inplace=True)
data.add_scope(subcort, 'subcort', inplace=True)
data.add_scope(dti_fa, 'dti_fa', inplace=True)
data.add_scope(dti_md, 'dti_md', inplace=True)
data.add_scope(brain, 'brain', inplace=True )
# Drop all NaN from any column
# Though BPt can generally handle NaN data fine,
# it makes certain pieces easier for this example as we don't have to worry
# about imputation.
data = data.dropna()
# Just show the first few rows
data.head()
```
The scopes we defined are nice, as it lets use check columns, or compose different scopes together. For example
we can check the scope we set 'cat covars' as composed with another variable as:
```
data.get_cols([cat_covars, 'b_rel_family_id'])
```
These are notably the columns we want to make sure are categorical, so lets ordinalize them, then plot.
```
data = data.ordinalize([cat_covars, 'b_rel_family_id'])
# Then plot just the categorical variables
data.plot(scope='category', subjects='all', decode_values=True)
```
Let's plot the target variables as well.
```
data.plot('target')
```
Okay, we note that there are some extreme outliers.
```
data = data.filter_outliers_by_std(n_std=10, scope='target')
data['target'].max()
data.plot('target')
```
Okay this maximum seem much more reasonable. Let's also assume that there may be some extreme values present in the input data as well, and that these represet corrupted data that we therefore want to drop.
```
# Repeat it twice, to deal with outliers at multiple scales
data = data.filter_outliers_by_std(n_std=10, scope='float')
data = data.filter_outliers_by_std(n_std=10, scope='float')
```
Next, we consider splitting up out data with a global train and test split. This can be useful in some instances. Note that we also define a cv strategy which says to perform the train test split keeping members of the same family in the same fold.
```
# Let's say we want to keep family members in the same train or test split
cv_strategy = bp.CVStrategy(groups='b_rel_family_id')
# Test split
data = data.set_test_split(.2, random_state=5, cv_strategy=cv_strategy)
data
```
## Single Training Set Evaluation
Our Dataset is now fully prepared. We can now define and evaluate a machine learning pipeline.
We will start by considering a pipeline which a few steps, these are:
1. Winsorize just the brain data.
2. Perform Robust Scaling on any 'float' type columns, 'brain' or 'covars'
3. One Hot Encode any categorical features
4. Fit an Elastic-Net Regression with nested random hyper-parameter search
Let's use one other feature of the toolbox, that is a custom cross-validation strategy. This is the same idea that we used when defining the train-test split, but now we want it to apply both during the evaluation and during the splits made when evaluating hyper-parameters.
```
#1
w_scaler = bp.Scaler('winsorize', quantile_range=(2, 98), scope='brain')
#2
s_scaler = bp.Scaler('standard', scope='float')
#3
ohe = bp.Transformer('one hot encoder', scope='category')
#4
param_search=bp.ParamSearch('RandomSearch', n_iter=60,
cv=bp.CV(splits=3, cv_strategy=cv_strategy))
elastic = bp.Model('elastic', params=1,
param_search=param_search)
# Now we can actually defined the pipeline
pipe = bp.Pipeline(steps=[w_scaler, s_scaler, ohe, elastic])
pipe
```
Let's say we want to use a 5-fold CV to evaluate this model on just the training set. We can first define the cv, the same as for the param_search above, but this time with splits=5.
```
cv=bp.CV(splits=5, n_repeats=1, cv_strategy=cv_strategy)
cv
```
And we will make a problem_spec to store some common params. In this case the random state for reproducibility of results and the number of jobs to use.
```
# Use problem spec just to store n jobs and random state
ps = bp.ProblemSpec(n_jobs=8, random_state=10)
ps
```
Now we are ready to evaluate this pipeline, let's check an example using the function evaluate.
We will set just one specific parameter to start, which is a scope of 'brain' to say
we just want to run a model with just the brain features (i.e., not the co-variates)
```
evaluator = bp.evaluate(pipeline=pipe,
dataset=data,
problem_spec=ps,
scope='brain',
cv=cv)
evaluator
```
Wait, what subset of subjects did we use? We want to make sure we are using just the training set of subjects at this stage. We just used default for the subjects, which in this case, i.e., when their is a test set defined, uses the training set as the subjects when passed to evaluate. We can confirm this by checking the intersection of the first train and validation fold with the test subjects
```
evaluator.train_subjects[0].intersection(data.test_subjects), evaluator.val_subjects[0].intersection(data.test_subjects)
```
As we can see, they are empty. We could also specify subjects='train' above to make sure this behavior occured.
## How to employ Test Set?
Next, we will discuss briefly two methods of evaluating on the test set.
1. There is a notion in some of the literature that the test set should be used to evaluate an existing estimator. In this case, that would consist of selecting from the 5 trained models we evaluated on the train set, and testing the one which performed best.
2. On the otherhand, we could instead re-train an estimator on the full training set and then test that on the test set. This is actually the reccomended strategy, but its still worth comparing both below.
Let's first do strategy 1.
```
# Get best model
best_model_ind = np.argmax(evaluator.scores['explained_variance'])
best_model = evaluator.estimators[best_model_ind]
# Get the correct test data - setting the problem spec
# as the problem spec saved when running this evaluator
X_test, y_test = data.get_Xy(problem_spec=evaluator.ps,
subjects='test')
# Score
score = best_model.score(X_test, y_test)
print(score)
```
Next, lets compare that with re-training our model on the full training set, then testing.
```
evaluator = bp.evaluate(pipeline=pipe,
dataset=data,
problem_spec=ps,
scope='brain', # Same as before
cv='test')
evaluator
```
Note that in this case, when cv='test' then all of the subjects will be used. If we wanted to make this behavior explicit we couldpass subjects='all'.
We can see that the results difer, though not hugely. One explanation is that at these sample sizes we still get boosts in performance from sample size, which means we expect our final model trained on the full and larger training set to do better. Another contributing factor is that by selecting from the 5-folds the model which does 'best' we may actually be choosing an overfit model which happened to do better on its corresponding validation set. We therefore reccomend the latter strategy.
In fact, we can formalize this intution, and test it on just the training set. We can do this because really the first method is just a type of meta-estimator. What it does is just in an internal nesting step, trains five models and selects the best one. We can formalize this as a custom model and compare it below.
We will also change the pipeline being compared a little bit for simplicity, replacing the elastic net we made via BPt with the ElasticNetCV object from scikit-learn.
```
from sklearn.model_selection import cross_validate
from sklearn.linear_model import ElasticNetCV
from sklearn.base import BaseEstimator
class BestOfModel(BaseEstimator):
def __init__(self, estimator, cv=5):
self.estimator = estimator
self.cv = cv
def fit(self, X, y):
scores = cross_validate(self.estimator, X, y, cv=self.cv,
return_estimator=True)
best_ind = np.argmax(scores['test_score'])
self.estimator_ = scores['estimator'][best_ind]
def predict(self, X):
return self.estimator_.predict(X)
# The new elastic net cv to use
new_elastic = ElasticNetCV()
# The best of model
best_of_model = bp.Model(
BestOfModel(estimator=new_elastic, cv=5))
# The static model
static_model = bp.Model(new_elastic)
# Use the same initial steps as before, but now with the best of model and the static models
best_of_pipe = bp.Pipeline(steps=[w_scaler, s_scaler, ohe, best_of_model])
static_pipe = bp.Pipeline(steps=[w_scaler, s_scaler, ohe, static_model])
```
Let's try the best of model first, evaluating it on the training set with 5-fold CV.
Note: This is going to be a bit slow as each time a model is trained in one of the 5-folds it needs to internally train 5 models. So it should take roughly 5x longer to evaluate then it's static counterpart.
```
evaluator = bp.evaluate(pipeline=best_of_pipe,
dataset=data,
problem_spec=ps,
scope='brain',
cv=cv)
evaluator
```
Next, let's run its corresponding 'static' counterpart.
```
evaluator = bp.evaluate(pipeline=static_pipe,
dataset=data,
problem_spec=ps,
scope='brain',
cv=cv)
evaluator
```
These results confirm the intution, namely that it is better to re-train using as many subjects as possible rather than chose a 'best model' from an internal CV.
## Compare & Evaluate
Let's now try out a newer feature the input class Compare. Let's define as our options to compare using different subsets of the input data.
```
compare_scopes = bp.Compare(['covars', 'brain', 'all',
'thick', 'area', 'subcort',
'dti_fa', 'dti_md'])
evaluators = bp.evaluate(pipeline=pipe,
scope=compare_scopes,
dataset=data,
problem_spec=ps,
cv=cv)
# Look at a summary of the results - note this option is only avaliable after a call to evaluate with
# Compare's has been made!
evaluators.summary()
```
We can also try another method avaliable which will run a pairwise model ttest comparison between all options
```
evaluators.pairwise_t_stats(metric='explained_variance')
```
We can also explicitly compare two evaluators
```
e1 = evaluators['scope=covars']
e2 = evaluators['scope=brain']
# Look at function docstring
print(' ', e1.compare.__doc__)
print()
e1.compare(e2)
```
Okay so it looks like it ran all the different combinations, but how do we look at each indivudal evaluator / the full results?? There are few different ways to index the object storing the different evaluators, but its essentially a dictionary. We can index one run as follows:
```
evaluators['scope=covars']
# Note, with get_fis, if mean=True it will return
# the mean and only non-null, and non-zero features directly.
evaluators['scope=covars'].get_fis(mean=True)
```
Warning: This doesn't mean that this mean features were selected! Remember this is the average over training 5 models, so it means that if a feature shows up it was selected at least once in one of the models!
```
# These are the different options
evaluators.keys()
```
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
%env CUDA_VISIBLE_DEVICES=0
import os, sys, time
sys.path.insert(0, '..')
import lib
import math
import numpy as np
from copy import deepcopy
import torch, torch.nn as nn
import torch.nn.functional as F
import matplotlib.pyplot as plt
%matplotlib inline
plt.style.use('seaborn-darkgrid')
plt.rcParams['pdf.fonttype'] = 42
plt.rcParams['ps.fonttype'] = 42
# For reproducibility
import random
seed = random.randint(0, 2 ** 32 - 1)
random.seed(seed)
np.random.seed(seed)
torch.manual_seed(seed)
torch.backends.cudnn.deterministic = True
torch.backends.cudnn.benchmark = False
print(seed)
```
# Setting
```
model_type = 'fixup_resnet'
# Dataset
data_dir = './data'
train_batch_size = 128
valid_batch_size = 128
test_batch_size = 64
num_workers = 3
pin_memory = True
num_classes = 100
device = 'cuda' if torch.cuda.is_available() else 'cpu'
loss_function = F.cross_entropy
# DIMAML initializers params
num_quantiles = 100
# MAML
max_steps = 3000
inner_loop_steps = 200
loss_kwargs={'reduction':'mean'}
first_val_step = 40
loss_interval = 40
assert (inner_loop_steps - first_val_step) % loss_interval == 0
validation_steps = int((inner_loop_steps - first_val_step) / loss_interval + 1)
# Optimizer
learning_rate=0.1
inner_optimizer_type='momentum'
inner_optimizer_kwargs = dict(
lr=learning_rate, momentum=0.9,
nesterov=True, weight_decay=0.0005
)
# Meta optimizer
meta_betas = (0.9, 0.997)
meta_learning_rate = 0.001
meta_grad_clip = 10.
checkpoint_steps = 3
recovery_step = None
kwargs = dict(
first_valid_step=first_val_step,
valid_loss_interval=loss_interval,
loss_kwargs=loss_kwargs,
)
exp_name = f"PLIF_FixupResNet18_CIFAR100_{inner_optimizer_type}"
exp_name += f"_steps{inner_loop_steps}_interval{loss_interval}"
exp_name += f"_tr_bs{train_batch_size}_val_bs{valid_batch_size}_seed_{seed}"
print("Experiment name: ", exp_name)
logs_path = "./logs/{}".format(exp_name)
assert recovery_step is not None or not os.path.exists(logs_path)
# !rm -rf {logs_path}
```
## Prepare CIFAR100
```
from torchvision import transforms, datasets
from torch.utils.data.sampler import SubsetRandomSampler
from torch.utils.data import TensorDataset, DataLoader
train_transform = transforms.Compose([
transforms.RandomCrop(32, padding=4),
transforms.RandomHorizontalFlip(),
transforms.ToTensor(),
transforms.Normalize((0.5071, 0.4865, 0.4409), (0.2673, 0.2564, 0.2762)),
])
eval_transform = transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.5071, 0.4865, 0.4409), (0.2673, 0.2564, 0.2762)),
])
train_dataset = datasets.CIFAR100(root=data_dir, train=True, download=True, transform=train_transform)
valid_dataset = datasets.CIFAR100(root=data_dir, train=True, download=True, transform=eval_transform)
test_set = datasets.CIFAR100(root=data_dir, train=False, download=True, transform=eval_transform)
num_train = len(train_dataset)
indices = list(range(num_train))
split = 40000
np.random.shuffle(indices)
train_idx, valid_idx = indices[:split], indices[split:]
train_sampler = SubsetRandomSampler(train_idx)
valid_sampler = SubsetRandomSampler(valid_idx)
train_loader = torch.utils.data.DataLoader(
train_dataset, batch_size=train_batch_size, sampler=train_sampler,
num_workers=num_workers, pin_memory=pin_memory,
)
valid_loader = torch.utils.data.DataLoader(
valid_dataset, batch_size=valid_batch_size, sampler=valid_sampler,
num_workers=num_workers, pin_memory=pin_memory,
)
test_loader = torch.utils.data.DataLoader(
test_set, batch_size=test_batch_size, shuffle=False,
num_workers=num_workers, pin_memory=pin_memory
)
```
## Create the model and meta-optimizer
```
optimizer = lib.make_inner_optimizer(inner_optimizer_type, **inner_optimizer_kwargs)
model = lib.models.fixup_resnet.FixupResNet18(num_classes=num_classes)
maml = lib.PLIF_MAML(model, model_type, optimizer=optimizer,
checkpoint_steps=checkpoint_steps,
loss_function=loss_function,
num_quantiles=num_quantiles
).to(device)
```
## Trainer
```
def samples_batches(dataloader, num_batches):
x_batches, y_batches = [], []
for batch_i, (x_batch, y_batch) in enumerate(dataloader):
if batch_i >= num_batches: break
x_batches.append(x_batch)
y_batches.append(y_batch)
return x_batches, y_batches
class TrainerResNet(lib.Trainer):
def train_on_batch(self, train_loader, valid_loader, prefix='train/', **kwargs):
""" Performs a single gradient update and reports metrics """
# Sample train and val batches
x_batches, y_batches = samples_batches(train_loader, inner_loop_steps)
x_val_batches, y_val_batches = samples_batches(valid_loader, validation_steps)
# Perform a meta training step
self.meta_optimizer.zero_grad()
with lib.training_mode(self.maml, is_train=True):
self.maml.resample_parameters()
updated_model, train_loss_history, valid_loss_history, *etc = \
self.maml.forward(x_batches, y_batches, x_val_batches, y_val_batches, **kwargs)
train_loss = torch.cat(train_loss_history).mean()
valid_loss = torch.cat(valid_loss_history).mean() if len(valid_loss_history) > 0 else torch.zeros(1)
valid_loss.backward()
# Check gradients
grad_norm = lib.utils.total_norm_frobenius(self.maml.initializers.parameters())
self.writer.add_scalar(prefix + "grad_norm", grad_norm, self.total_steps)
bad_grad = not math.isfinite(grad_norm)
if not bad_grad and self.meta_grad_clip:
nn.utils.clip_grad_norm_(list(self.maml.initializers.parameters()), self.meta_grad_clip)
else:
print("Fix bad grad. Loss {} | Grad {}".format(train_loss.item(), grad_norm))
for param in self.maml.initializers.parameters():
param.grad = torch.where(torch.isfinite(param.grad),
param.grad, torch.zeros_like(param.grad))
self.meta_optimizer.step()
return self.record(train_loss=train_loss.item(),
valid_loss=valid_loss.item(), prefix=prefix)
def evaluate_metrics(self, train_loader, test_loader, prefix='val/', **kwargs):
""" Predicts and evaluates metrics over the entire dataset """
torch.cuda.empty_cache()
print('Baseline')
self.maml.resample_parameters(initializers=self.maml.untrained_initializers, is_final=True)
base_model = deepcopy(self.maml.model)
base_train_loss_history, base_test_loss_history, base_test_error_history = \
eval_model(base_model, train_loader, test_loader, epochs=1, device=self.device)
print('Ours')
self.maml.resample_parameters(is_final=True)
maml_model = deepcopy(self.maml.model)
maml_train_loss_history, maml_test_loss_history, maml_test_error_history = \
eval_model(maml_model, train_loader, test_loader, epochs=1, device=self.device)
lib.utils.resnet_draw_plots(base_train_loss_history, base_test_loss_history,
base_test_error_history, maml_train_loss_history,
maml_test_loss_history, maml_test_error_history)
self.writer.add_scalar(prefix + "train_AUC", sum(maml_train_loss_history), self.total_steps)
self.writer.add_scalar(prefix + "test_AUC", sum(maml_test_loss_history), self.total_steps)
self.writer.add_scalar(prefix + "test_loss", maml_test_loss_history[-1], self.total_steps)
self.writer.add_scalar(prefix + "test_cls_error", maml_test_error_history[-1], self.total_steps)
########################
# Generate Train Batch #
########################
def generate_train_batches(train_loader, batches_in_epoch=150):
x_batches, y_batches = [], []
for batch_i, (x_batch, y_batch) in enumerate(train_loader):
if batch_i >= batches_in_epoch: break
x_batches.append(x_batch)
y_batches.append(y_batch)
assert len(x_batches) == len(y_batches) == batches_in_epoch
local_x = torch.cat(x_batches, dim=0)
local_y = torch.cat(y_batches, dim=0)
local_dataset = TensorDataset(local_x, local_y)
local_dataloader = DataLoader(local_dataset, batch_size=train_batch_size,
shuffle=True, num_workers=num_workers)
return local_dataloader
##################
# Eval functions #
##################
def adjust_learning_rate(optimizer, epoch, milestones=[30, 50]):
"""decrease the learning rate at 30 and 50 epoch"""
lr = learning_rate
if epoch >= milestones[0]:
lr /= 10
if epoch >= milestones[1]:
lr /= 10
for param_group in optimizer.param_groups:
if param_group['initial_lr'] == learning_rate:
param_group['lr'] = lr
else:
if epoch < milestones[0]:
param_group['lr'] = param_group['initial_lr']
elif epoch < milestones[1]:
param_group['lr'] = param_group['initial_lr'] / 10.
else:
param_group['lr'] = param_group['initial_lr'] / 100.
return lr
@torch.no_grad()
def compute_test_loss(model, loss_function, test_loader, device='cuda'):
model.eval()
test_loss, cls_error = 0., 0.
for x_test, y_test in test_loader:
x_test, y_test = x_test.to(device), y_test.to(device)
preds = model(x_test)
test_loss += loss_function(preds, y_test) * x_test.shape[0]
cls_error += 1. * (y_test != preds.argmax(axis=-1)).sum()
test_loss /= len(test_loader.dataset)
cls_error /= len(test_loader.dataset)
model.train()
return test_loss.item(), cls_error.item()
def eval_model(model, train_loader, test_loader, epochs=3, test_loss_interval=40, device='cuda'):
optimizer = lib.optimizers.make_eval_inner_optimizer(
maml, model, inner_optimizer_type,
**inner_optimizer_kwargs
)
for param_group in optimizer.param_groups:
param_group['initial_lr'] = learning_rate
# Train loop
train_loss_history = []
test_loss_history = []
test_error_history = []
training_mode = model.training
total_iters = 0
for epoch in range(epochs):
model.train()
lr = adjust_learning_rate(optimizer, epoch)
for i, (x_batch, y_batch) in enumerate(train_loader):
optimizer.zero_grad()
preds = model(x_batch.to(device))
loss = loss_function(preds, y_batch.to(device))
loss.backward()
grad_norm = nn.utils.clip_grad_norm_(model.parameters(), 4)
optimizer.step()
if (total_iters == 0) or (total_iters + 1) % test_loss_interval == 0:
train_loss_history.append(loss.item())
model.eval()
test_loss, test_error = compute_test_loss(model, loss_function, test_loader, device=device)
print("Epoch {} | Train Loss {:.4f} | Test Loss {:.4f} | Classification Error {:.4f}"\
.format(epoch, loss.item(), test_loss, test_error))
test_loss_history.append(test_loss)
test_error_history.append(test_error)
model.train()
total_iters += 1
model.train(training_mode)
return train_loss_history, test_loss_history, test_error_history
train_loss_history = []
valid_loss_history = []
trainer = TrainerResNet(maml, meta_lr=meta_learning_rate,
meta_betas=meta_betas, meta_grad_clip=meta_grad_clip,
exp_name=exp_name, recovery_step=recovery_step)
```
## Training
```
from IPython.display import clear_output
t0 = time.time()
while trainer.total_steps <= max_steps:
lib.free_memory()
metrics = trainer.train_on_batch(
train_loader, valid_loader, **kwargs
)
train_loss = metrics['train_loss']
train_loss_history.append(train_loss)## Training
valid_loss = metrics['valid_loss']
valid_loss_history.append(valid_loss)
if trainer.total_steps % 10 == 0:
clear_output(True)
print("Step: %d | Time: %f | Train Loss %.5f | Valid loss %.5f"
% (trainer.total_steps, time.time()-t0, train_loss, valid_loss))
plt.figure(figsize=[16, 5])
plt.subplot(1,2,1)
plt.title('Train Loss over time')
plt.plot(lib.utils.moving_average(train_loss_history, span=50))
plt.scatter(range(len(train_loss_history)), train_loss_history, alpha=0.1)
plt.subplot(1,2,2)
plt.title('Valid Loss over time')
plt.plot(lib.utils.moving_average(valid_loss_history, span=50))
plt.scatter(range(len(valid_loss_history)), valid_loss_history, alpha=0.1)
plt.show()
local_train_loader = generate_train_batches(train_loader, inner_loop_steps)
trainer.evaluate_metrics(local_train_loader, test_loader, test_interval=20)
lib.utils.resnet_visualize_quantile_functions(maml)
t0 = time.time()
if trainer.total_steps % 100 == 0:
trainer.save_model()
trainer.total_steps += 1
lib.utils.resnet_visualize_quantile_functions(maml)
```
# Evaluation
```
seed = random.randint(0, 2 ** 32 - 1)
random.seed(seed)
np.random.seed(seed)
torch.manual_seed(seed)
print(seed)
def gradient_quotient(loss, params, eps=1e-5):
grad = torch.autograd.grad(loss, params, retain_graph=True, create_graph=True)
prod = torch.autograd.grad(sum([(g**2).sum() / 2 for g in grad]),
params, retain_graph=True, create_graph=True)
out = sum([((g - p) / (g + eps * (2*(g >= 0).float() - 1).detach()) - 1).abs().sum()
for g, p in zip(grad, prod)])
return out / sum([p.data.nelement() for p in params])
def metainit(model, criterion, x_size, y_size, lr=0.1, momentum=0.9, steps=200, eps=1e-5):
model.eval()
params = [p for p in model.parameters()
if p.requires_grad and len(p.size()) >= 2 and
math.isfinite(p.std().item()) and p.std().item() > 0]
memory = [0] * len(params)
for i in range(steps):
input = torch.Tensor(*x_size).normal_(0, 1).cuda()
target = torch.randint(0, y_size, (x_size[0],)).cuda()
loss = criterion(model(input), target)
gq = gradient_quotient(loss, list(model.parameters()), eps)
grad = torch.autograd.grad(gq, params)
for j, (p, g_all) in enumerate(zip(params, grad)):
norm = p.data.norm().item()
g = torch.sign((p.data * g_all).sum() / norm)
memory[j] = momentum * memory[j] - lr * g.item()
new_norm = norm + memory[j]
p.data.mul_(new_norm / (norm + eps))
print("%d/GQ = %.2f" % (i, gq.item()))
def genOrthgonal(dim):
a = torch.zeros((dim, dim)).normal_(0, 1)
q, r = torch.qr(a)
d = torch.diag(r, 0).sign()
diag_size = d.size(0)
d_exp = d.view(1, diag_size).expand(diag_size, diag_size)
q.mul_(d_exp)
return q
def makeDeltaOrthogonal(weights, gain):
rows = weights.size(0)
cols = weights.size(1)
if rows < cols:
print("In_filters should not be greater than out_filters.")
weights.data.fill_(0)
dim = max(rows, cols)
q = genOrthgonal(dim)
mid1 = weights.size(2) // 2
mid2 = weights.size(3) // 2
with torch.no_grad():
weights[:, :, mid1, mid2] = q[:weights.size(0), :weights.size(1)]
weights.mul_(gain)
```
## Eval TinyImageNet
```
data_dir = 'data/tiny-imagenet-200/'
num_workers = {'train': 0, 'val': 0,'test': 0}
data_transforms = {
'train': transforms.Compose([
transforms.ColorJitter(brightness=0.1, contrast=0.1, saturation=0.1),
transforms.RandomRotation(20),
transforms.RandomHorizontalFlip(0.5),
transforms.ToTensor(),
transforms.Normalize([0.4802, 0.4481, 0.3975], [0.2302, 0.2265, 0.2262]),
]),
'val': transforms.Compose([
transforms.ToTensor(),
transforms.Normalize([0.4802, 0.4481, 0.3975], [0.2302, 0.2265, 0.2262]),
]),
'test': transforms.Compose([
transforms.ToTensor(),
transforms.Normalize([0.4802, 0.4481, 0.3975], [0.2302, 0.2265, 0.2262]),
])
}
image_datasets = {x: datasets.ImageFolder(os.path.join(data_dir, x), data_transforms[x])
for x in ['train', 'val','test']}
dataloaders = {x: torch.utils.data.DataLoader(image_datasets[x], batch_size=128, shuffle=True, num_workers=num_workers[x])
for x in ['train', 'val', 'test']}
dataset_sizes = {x: len(image_datasets[x]) for x in ['train', 'val', 'test']}
ti_batches_in_epoch = len(dataloaders['train'])
assert ti_batches_in_epoch == 782
num_reruns = 10
reruns_base_test_loss_history = []
reruns_base_test_error_history = []
reruns_metainit_test_loss_history = []
reruns_metainit_test_error_history = []
reruns_maml_test_loss_history = []
reruns_maml_test_error_history = []
reruns_deltaorthogonal_test_loss_history = []
reruns_deltaorthogonal_test_error_history = []
for i in range(num_reruns):
print(f"Rerun {i}")
print("Baseline")
maml.resample_parameters(initializers=maml.untrained_initializers, is_final=True)
base_model = deepcopy(maml.model)
base_model.fc = nn.Linear(in_features=512, out_features=200, bias=True).to(device)
nn.init.constant_(base_model.fc.weight, 0)
nn.init.constant_(base_model.fc.bias, 0)
base_train_loss_history, base_test_loss_history, base_test_error_history = \
eval_model(base_model, dataloaders['train'], dataloaders['test'],
epochs=70, test_loss_interval=ti_batches_in_epoch, device=device)
reruns_base_test_loss_history.append(base_test_loss_history)
reruns_base_test_error_history.append(base_test_error_history)
print("DIMAML")
maml.resample_parameters(is_final=True)
maml_model = deepcopy(maml.model)
maml_model.fc = nn.Linear(in_features=512, out_features=200, bias=True).to(device)
nn.init.constant_(maml_model.fc.weight, 0)
nn.init.constant_(maml_model.fc.bias, 0)
maml_train_loss_history, maml_test_loss_history, maml_test_error_history = \
eval_model(maml_model, dataloaders['train'], dataloaders['test'],
epochs=70, test_loss_interval=ti_batches_in_epoch, device=device)
reruns_maml_test_loss_history.append(maml_test_loss_history)
reruns_maml_test_error_history.append(maml_test_error_history)
print("MetaInit")
batch_x, _ = next(iter(dataloaders['train']))
batch_x = batch_x[:64]
metainit_model = lib.models.FixupResNet18(num_classes=200, default_init=True).to(device)
metainit(metainit_model, loss_function, batch_x.shape, 200)
metainit_train_loss_history, metainit_test_loss_history, metainit_test_error_history = \
eval_model(metainit_model, dataloaders['train'], dataloaders['test'],
epochs=70, test_loss_interval=ti_batches_in_epoch, device=device)
reruns_metainit_test_loss_history.append(metainit_test_loss_history)
reruns_metainit_test_error_history.append(metainit_test_error_history)
print("DeltaOrthogonal")
deltaorthogonal_model = lib.models.FixupResNet18(num_classes=200).to(device)
for param in deltaorthogonal_model.parameters():
if len(param.size()) >= 4:
makeDeltaOrthogonal(param, nn.init.calculate_gain('leaky_relu'))
deltaorthogonal_train_loss_history, deltaorthogonal_test_loss_history, deltaorthogonal_test_error_history = \
eval_model(deltaorthogonal_model, dataloaders['train'], dataloaders['test'],
epochs=70, test_loss_interval=ti_batches_in_epoch, device=device)
reruns_deltaorthogonal_test_loss_history.append(deltaorthogonal_test_loss_history)
reruns_deltaorthogonal_test_error_history.append(deltaorthogonal_test_error_history)
base_mean = np.array(reruns_base_test_error_history).mean(0)
base_std = np.array(reruns_base_test_error_history).std(0, ddof=1)
maml_mean = np.array(reruns_maml_test_error_history).mean(0)
maml_std = np.array(reruns_maml_test_error_history).std(0, ddof=1)
metainit_mean = np.array(reruns_metainit_test_error_history).mean(0)
metainit_std = np.array(reruns_metainit_test_error_history).std(0, ddof=1)
deltaorthogonal_mean = np.array(reruns_deltaorthogonal_test_error_history).mean(0)
deltaorthogonal_std = np.array(reruns_deltaorthogonal_test_error_history).std(0, ddof=1)
```
| github_jupyter |
# Package Usage Guide
This notebook contains all the steps you need to follow to get the most out of this package. This notebook also contains and briefly explains the available modules, classes and methods in the package.
## Objective
As health data is private information and cannot be shared freely, the limitation on how much can be learnt from the limited freely available data is quite evident. The HealthGAN neural network in this generates synthetic dataset from the original dataset which can be shared without impairing privacy.
The package supplements the GAN with preprocessing and evaluation metrics so the package can be used as needed.
## Using the package
Lets dive and see how the package can be used.
```
# Remove warnings occuring due to use of older package versions
import warnings
warnings.filterwarnings("ignore")
```
### Processing
The first step is to have a training file and a testing file. We will consider the case that we have the training file *train.csv* and testing file as *test.csv* inside the folder *data_files*.
We will use the **processing** module to create the **Encoder()** class which encodes the training ang testing files into SDV files which the GAN accepts using **encode_train()** and **encode_test()** functions respectively.
```
from synthetic_data.generators.processing import Encoder
en = Encoder()
```
The **encode_train()** method expects the training file and returns the SDV file along with **limits**, **min_max** and **cols** files which are used for encoding and decoding.
```
en.encode_train("data/train.csv")
```
The **encode_test()** method expects the testing file as first argument and the original training file as the second argument. One must note that the training file must be encoded before the testing file is encoded.
```
en.encode_test("data/test.csv", "data/train.csv")
```
These will generate the SDV files inside the *data_files* folder which can now be used for training our model.
### Using HealthGAN
Now, the files are ready to be used by the HealthGAN, so we import it and simply call the **train()** method on the **HealthGAN** class. The GAN expects SDV converted files, thus we should pass the appropriate files generated by the encoder above (same names with suffix *_sdv*).
```
from synthetic_data.generators.gan import HealthGAN
gan = HealthGAN(train_file = "data/train_sdv.csv",
test_file = "data/test_sdv.csv",
base_nodes = 64,
critic_iters = 5,
num_epochs = 10000)
gan.train()
```
The GAN produces the model values and 10 synthetic data files which are all saved in the folder *gen_data*.
## Evaluation
The package provides several different types of evaluation metrics: **Adversarial accuracy**, **Divergence score**, **Discrepancy score**, **PCA plot**, **6 subplot PCA plot** and **6 subplot TSNE plot**.
```
from synthetic_data.metrics.scores import Scores
from synthetic_data.metrics.plots import LossPlot, ComponentPlots
```
Here, we'll consider the name of various generated synthetic files as *synth_* followed by a unique number, and the log file will be *log.pkl*
#### Adversarial accuracy, divergence and discrepancy scores
```
scores = Scores(train_file = "data/train_sdv.csv",
test_file = "data/test_sdv.csv",
synthetic_files = ["gen_data/synth_0.csv"])
scores.calculate_accuracy()
scores.compute_divergence()
scores.compute_discrepancy()
```
#### Plots
```
lossPlot = LossPlot(log_file = "gen_data/log.pkl")
lossPlot.plot()
componentPlots = ComponentPlots()
componentPlots.pca_plot(real_data = "data/train_sdv.csv",
synthetic_data = "gen_data/synth_0.csv")
componentPlots.combined_pca(real_data = "data/train_sdv.csv",
synthetic_datas = ["gen_data/synth_0.csv",
"gen_data/synth_1.csv",
"gen_data/synth_2.csv",
"gen_data/synth_3.csv",
"gen_data/synth_4.csv",
"gen_data/synth_5.csv"],
names = ["Data1", "Data2", "Data3", "Data4", "Data5", "Data6"])
componentPlots.combined_tsne(real_data = "data/train_sdv.csv",
synthetic_datas = ["gen_data/synth_0.csv",
"gen_data/synth_1.csv",
"gen_data/synth_2.csv",
"gen_data/synth_3.csv",
"gen_data/synth_4.csv",
"gen_data/synth_5.csv"],
names = ["Data1", "Data2", "Data3", "Data4", "Data5", "Data6"])
```
For each of these plots, the images are saved inside *gen_data/plots* folder.
| github_jupyter |
Copyright Amazon.com, Inc. or its affiliates. All Rights Reserved.
SPDX-License-Identifier: Apache-2.0

# Link Prediction - Introduction
In this Notebook we are going to examine the process of using Amazon Neptune ML feature to perform link prediction in a property graph.
**Note:** Link prediction models are computationally complex, so this notebook takes ~2-3 hours hour to complete
Neptune ML is the ability within Amazon Neptune to create and manage machine learning models using [Amazon SageMaker](https://aws.amazon.com/sagemaker/) and [Deep Graph Library](https://www.dgl.ai/) as use these models within a query to predict elements in the graph. For this notebook we are going to focus on how to predict edges between a pair of vertices in a graph using a type of task known as **link prediction**.
(https://docs.aws.amazon.com/neptune/latest/userguide/machine-learning.html#machine-learning-overview)
To accomplish this in Neptune ML we use a four step process:
1. **Load Data** - Data is loaded into a Neptune cluster using any of the normal methods such as the via the Gremlin drivers or using the Neptune Bulk Loader.
2. **Export Data** - A service call is made specifying the machine learning model type and model configuration parameters. The data and model configuration parameters are then exported from a Neptune cluster to an S3 bucket.
3. **Model Training** - A set of service calls are made to pre-process the exported data, train the machine learning model, and then generate an Amazon SageMaker endpoint that exposes the model.
4. **Run Queries** - The final step is to use this inference endpoint within our Gremlin queries to infer data using the machine learning model.

For this notebook we'll use the [MovieLens 100k dataset](https://grouplens.org/datasets/movielens/100k/) provided by [GroupLens Research](https://grouplens.org/datasets/movielens/). This dataset consists of movies, users, and ratings of those movies by users.

Each of these vertices have a set of edge associated with them but for this notebook we are interested in the `rated` edge between a `user` vertex and a `movie` vertex. To demonstrate link prediction we'll walk through constructing a Neptune ML model from scratch. We'll then demonstrate how to use that model to infer `rated` edges within Gremlin traversals.
## Checking that we are ready to run Neptune ML
This notebook assumes that you have already setup and enabled a cluster for Neptune ML. If you have not then please follow the directions located here: [Neptune ML Overview](https://docs.aws.amazon.com/neptune/latest/userguide/machine-learning.html#machine-learning-overview)
Run the code below to check that your cluster is configured to run Neptune ML.
```
import neptune_ml_utils as neptune_ml
neptune_ml.check_ml_enabled()
```
If the check above did not say that this cluster is ready to run Neptune ML jobs then please check the configuration of your cluster.
# Load the data
The first step in building a Neptune ML model is to load data into the Neptune cluster. Loading data for Neptune ML follows the standard process of ingesting data into Amazon Neptune, for this example we'll be using the Bulk Loader.
For this notebook we have written a script that automates the process of downloading the data from the MovieLens websites and formatting it to load into Neptune. To begin specify an S3 bucket URI (must be located in the same region as the cluster) and then run the cell below, which takes ~1 minute:
```
s3_bucket_uri="s3://<INSERT S3 BUCKET>"
response = neptune_ml.prepare_movielens_data(s3_bucket_uri)
```
This process only takes a few minutes and once it has completed you can load the data using the `%load` command in the cell below.
```
%load -s {response} -f csv -p OVERSUBSCRIBE
```
## Check to make sure the data is loaded
Once the cell has completes execution, the data has now been loaded into the cluster. Assuming the cluster was previously empty we can verify the data loaded correctly by running the traversals below to see the count of vertices by label:
```
%%gremlin
g.V().groupCount().by(label).unfold()
```
If our vertices loaded correctly then the output is:
* 1682 movies
* 943 users
* 19 genres
* 100000 rating
To check that our edges loaded correctly we check the edge counts:
```
%%gremlin
g.E().groupCount().by(label).unfold()
```
If our edges loaded correctly then the output is:
* 100000 rated
* 2893 included_in
* 100000 wrote
* 100000 about
## Preparing for Export
With our data validated let's remove some `rated` edges so that we can build a model that predicts these missing connections. In a normal scenario, the data you would like to predict is most likely missing from the data being loaded so removing these values prior to building our machine learning model simulates that situation.
Specifically, let's remove the `rated` edgesfor `user_1`, to provide us with a few candidate vertices to run our link prediction tasks. Let's start by taking a look at what `rated` edges currently exist.
```
%%gremlin
g.V('user_1').outE('rated')
```
Now let's remove these edges to simulate them missing from our data.
```
%%gremlin
g.V('user_1').outE('rated').drop()
```
Checking our data again we see that the edges have now been removed.
```
%%gremlin
g.V('user_1').outE('rated')
```
# Export the data and model configuration
**Note:** If you created your cluster using the CloudFormation script provided in the documentation then the Neptune Export has been installed and configured for you. If you did not then you need to ensure it is configured as described here: [Neptune Export Service](https://docs.aws.amazon.com/neptune/latest/userguide/machine-learning-data-export-service.html#machine-learning-data-export-service-run-export).
With the data loaded we are ready to export the data and configuration which will be used to train the ML model.
The export process is triggered by a call to the [Neptune Export service endpoint](https://docs.aws.amazon.com/neptune/latest/userguide/machine-learning-data-export-service.html). This call contains a configuration object which specifies the type of machine learning model to build, in this example node classification, as well as any feature configurations required.
**Note** The used in this notebook specifies only a minimal set of configuration options meaning that our model's predictions are not as accurate as they could be. The parameters included in this configuration are one of a couple of sets of options available to the end user to tune the model and optimize the accuracy of the resulting predictions.
The configuration options provided to the export service are broken into two main sections, selecting the target and configuring features.
## Selecting Targets for Link Prediction
In the first section, selecting the target, we specify what type of machine learning task will be run. To run a link prediction mdoel do not specify any `targets` in the `additionalParams` value. Unlike node classification or node regression, link prediction can be used to predict any edge type that exists in the graph between any two vertices. Becasue of this, there is no need to define a target set of values.
## Configuring Features to Train the Model
The second section of the configuration, configuring features, is where we specify details about the types of data stored in our graph and how the machine learning model should interpret that data. In machine learning, each property is known as a feature and these features are used by the model to make predictions.
In machine learning, each property is known as a feature and these features are used by the model to make predictions. When data is exported from Neptune all properties of all vertices are included. Each property is treated as a separate feature for the ML model. Neptune ML does its best to infer the correct type of feature for a property, in many cases, the accuracy of the model can be improved by specifying information about the property used for a feature. By default Neptune ML puts features into one of two categories:
* If the feature represents a numerical property (float, double, int) then it is treated as a `numerical` feature type. In this feature type data is represented as a continuous set of numbers. In our example, the `age` of a `user` would best be represented as a numerical feature as the age of a user is best represented as a continuous set of values.
* All other property types are represented as `category` features. In this feature type, each unique value of data is represented as a unique value in the set of classifications used by the model. In our MovieLens example the `occupation` of a `user` would represent a good example of a `category` feature as we want to group users that all have the same job.
If all of the properties fit into these two feature types then no configuration changes are needed at the time of export. However, in many scenarios these defaults are not always the best choice. In these cases, additional configuration options should be specified to better define how the property should be represented as a feature.
One common feature that needs additional configuration is numerical data, and specifically properties of numerical data that represent chunks or groups of items instead of a continuous stream.
Let's say that instead of wanting `age` to be represented as a set of continuous values we want to represent it as a set of discrete buckets of values (e.g. 18-25, 26-24, 35-44, etc.). In this scenario we want to specify some additional attributes of that feature to bucket this attribute into certain known sets. We can achieve this by specifying this feature as a `numerical_bucket`. This feature type takes a range of expected values, as well as a number of buckets, and groups data into buckets during the training process.
Another common feature that needs additional attributes are text features such as names, titles, or descriptions. While Neptune ML will treat these as categorical features by default the reality of these features is that they will likely be unique for each vertex. For example, since the `title` property of a `movie` vertex does not fit into a category grouping our model would be better served by representing this type of feature as a `word2vec` feature. A `word2vec` feature uses techniques from natural language processing to create a vector of data that represents a string of text.
In our export example below we have specified that the `title` property of our `movie` should be exported and trained as a `word2vec` feature and that our `age` field should range from 0-100 and that data should be bucketed into 10 distinct groups.
**Important:** The example below is an example of a minimal amount of the features of the model configuration parameters and will not create the most accurate model possible. Additional options are available for tuning this configuration to produce an optimal model are described here: [Neptune Export Process Parameters](https://docs.aws.amazon.com/neptune/latest/userguide/machine-learning-data-export-parameters.html)
Running the cell below we set the export configuration and run the export process. Neptune export is capable of automatically creating a clone of the cluster by setting `cloneCluster=True` which takes about 20 minutes to complete and will incur additional costs while the cloned cluster is running. Exporting from the existing cluster takes about 5 minutes but requires that the `neptune_query_timeout` parameter in the [parameter group](https://docs.aws.amazon.com/neptune/latest/userguide/parameters.html) is set to a large enough value (>72000) to prevent timeout errors.
```
export_params={
"command": "export-pg",
"params": { "endpoint": neptune_ml.get_host(),
"profile": "neptune_ml",
"cloneCluster": False
},
"outputS3Path": f'{s3_bucket_uri}/neptune-export',
"additionalParams": {
"neptune_ml": {
"features": [
{
"node": "movie",
"property": "title",
"type": "word2vec"
},
{
"node": "user",
"property": "age",
"type": "bucket_numerical",
"range" : [1, 100],
"num_buckets": 10
}
]
}
},
"jobSize": "medium"}
%%neptune_ml export start --export-url {neptune_ml.get_export_service_host()} --export-iam --wait --store-to export_results
${export_params}
```
# ML Data Processing, Model Training, and Endpoint Creation
Once the export job is completed we are now ready to train our machine learning model and create the inference endpoint. Training our Neptune ML model requires three steps. The cells below configure a minimal set of parameters required to run a model training.
The first step (data processing) processes the exported graph dataset using standard feature preprocessing techniques to prepare it for use by DGL. This step performs functions such as feature normalization for numeric data and encoding text features using word2vec. At the conclusion of this step the dataset is formatted for model training.
This step is implemented using a SageMaker Processing Job and data artifacts are stored in a pre-specified S3 location once the job is complete.
Additional options and configuration parameters for the data processing job can be found using the links below:
* [Data Processing](https://docs.aws.amazon.com/neptune/latest/userguide/machine-learning-on-graphs-processing.html)
* [dataprocessing command](https://docs.aws.amazon.com/neptune/latest/userguide/machine-learning-api-dataprocessing.html)
**Note** Each time you run this you must also provide a unique identifier for the training run under the `training_job_name`. If you attempt to use a name that has already been used then it will result in an error.
```
# The training_job_name can be set to a unique value below, otherwise one will be auto generated
training_job_name=neptune_ml.get_training_job_name('link-prediction')
processing_params = f"""
--config-file-name training-job-configuration.json
--job-id {training_job_name}
--s3-input-uri {export_results['outputS3Uri']}
--s3-processed-uri {str(s3_bucket_uri)}/preloading """
%neptune_ml dataprocessing start --wait --store-to processing_results {processing_params}
```
The second step (model training) trains the ML model that will be used for predictions. The model training is done in two stages. The first stage uses a SageMaker Processing job to generate a model training strategy. A model training strategy is a configuration set that specifies what type of model and model hyperparameter ranges will be used for the model training. Once the first stage is complete, the SageMaker Processing job launches a SageMaker Hyperparameter tuning job. The SageMaker Hyperparameter tuning job runs a pre-specified number of model training job trials on the processed data, and stores the model artifacts generated by the training in the output S3 location. Once all the training jobs are complete, the Hyperparameter tuning job also notes the training job that produced the best performing model.
Additional options and configuration parameters for the data processing job can be found using the links below:
* [Model Training](https://docs.aws.amazon.com/neptune/latest/userguide/machine-learning-on-graphs-model-training.html)
* [modeltraining command](https://docs.aws.amazon.com/neptune/latest/userguide/machine-learning-api-modeltraining.html)
**Note** Link prediction is a computationally complex model to generate so training this model will take 2-3 hours
```
training_params=f"""
--job-id {training_job_name}
--data-processing-id {training_job_name}
--instance-type ml.p3.2xlarge
--s3-output-uri {str(s3_bucket_uri)}/training """
%neptune_ml training start --wait --store-to training_results {training_params}
```
The final step is to create the inference endpoint which is an Amazon SageMaker endpoint instance that is launched with the model artifacts produced by the best training job. This endpoint will be used by our graph queries to return the model predictions for the inputs in the request. The endpoint once created stays active until it is manually deleted. Each model is tied to a single endpoint.
Additional options and configuration parameters for the data processing job can be found using the links below:
* [Inference Endpoint](https://docs.aws.amazon.com/neptune/latest/userguide/machine-learning-on-graphs-inference-endpoint.html)
* [endpoint command](https://docs.aws.amazon.com/neptune/latest/userguide/machine-learning-api-endpoints.html)
**Note** The endpoint creation process takes ~5-10 minutes
```
endpoint_params=f"""
--job-id {training_job_name}
--model-job-id {training_job_name}"""
%neptune_ml endpoint create --wait --store-to endpoint_results {endpoint_params}
```
Once this has completed we get the endpoint name for our newly created inference endpoint. The cell below will print out the endpoint name which will be used in the configuration of Neptune ML in your Gremlin query.
```
print(endpoint_results['endpoint']['name'])
```
# Querying using Gremlin
Now that we have built our inference endpoint let's see how we can use this to query our graph to find out what `rated` edges will connect `user_1` to movies.
Link prediction is the ability to infer edges between two vertices in our graph. Unlike node classification and node regression, link prediction can infer any of the edge labels that existed in our graph when the model was created. In our model this means we could infer the probability that a `wrote`, `about`, `rated`, or `included_in` edge exists between any two vertices. However for this example we are going to focus on inferring the `rated` edges between the `user` and `movie` vertices.
To demonstrate this let's predict the `rated` edges exist for `user_1`.
A standard Gremlin query to find this information would look like the cell below:
```
%%gremlin
g.V('user_1').out('rated').hasLabel('movie').valueMap()
```
As we can see, their are currently not any `rated` edges for `user_1` so let's see how we can go about predicting them.
First, as with node classification and regression we need to specify the endpoint we are going to use. (`with("Neptune#ml.endpoint","<INSERT ENDPOINT NAME>"`).
Next, when we ask for the link within our query we use the `out()` step to predict the target vertex or the `in()` step to predict the source vertex. For each of these steps we need to specify the type of model being used with a with() step (with("Neptune#ml.prediction")).
One additional requirement for link prediction is that we need to use the `hasLabel('labelname')` step after the `out()` or `in()` step which specifies the type of vertex to predict.
Putting these items together we get the query below, which will find the movies that` user_1` is likely to rate.
```
%%gremlin
g.with("Neptune#ml.endpoint","<INSERT ENDPOINT NAME>").
V('user_1').out('rated').with("Neptune#ml.prediction").hasLabel('movie').valueMap()
```
Great, we can now see that we are getting edges returned from our prediction that do not exist within the graph data. In the example above we predicted the target vertex but we can also use the same mechanism to predict the source vertex. In the example below we find the three `user` vertices most likely to create a `rated` edge for the movie *Toy Story*.
```
%%gremlin
g.with("Neptune#ml.endpoint","<INSERT ENDPOINT NAME>").
with("Neptune#ml.limit",3).
V().has('title', 'Toy Story (1995)').
in('rated').with("Neptune#ml.prediction").hasLabel('user').valueMap()
```
As with the other example models we have shown here these link prediction values can be combined with other Gremlin traversal steps to create more complex traversals, such as returning the path from `user_1` across the inferred `rated` edge to the `movie` vertex to retrieve the `genre` vertices that it is included in.
```
%%gremlin
g.with("Neptune#ml.endpoint","<INSERT ENDPOINT NAME>").
V('user_1').out('rated').with("Neptune#ml.prediction").hasLabel('movie').out('included_in').path()
```
From the examples we have shown here you can begin to see how the ability to infer unknown connections within a graph starts to enable many interesting and unique use cases within Amazon Neptune.
# Cleaning Up
Now that you have completed this walkthrough you have created a Sagemaker endpoint which is currently running and will incur the standard charges. If you are done trying out Neptune ML and would like to avoid thes recurring costs, run the cell below to delete the inference endpoint.
```
neptune_ml.delete_endpoint(training_job_name)
```
In addition to the inference endpoint the CloudFormation script that you used has setup several additional resources. If you are finished then we suggest you delete the CloudFormation stack to avoid any recurring charges. For instructions, see Deleting a Stack on the AWS CloudFormation Console. Be sure to delete the root stack (the stack you created earlier). Deleting the root stack deletes any nested stacks.
| github_jupyter |
# Contents of the Repo , New York Taxi LFS
```
# Getting Dendencies
import pandas as pd
import csv
import numpy as np
from datetime import datetime
import datetime as dt
from datetime import timedelta # it is a spanned based time .
from datetime import datetime, date, time
from datetime import date
import matplotlib.pyplot as plt
import os, json, requests, pickle
from scipy.stats import skew
from shapely.geometry import Point,Polygon,MultiPoint,MultiPolygon
from scipy.stats import ttest_ind, f_oneway, lognorm, levy, skew, chisquare
#import scipy.stats as st
from sklearn.preprocessing import normalize, scale
from tabulate import tabulate #pretty print of tables. source: http://txt.arboreus.com/2013/03/13/pretty-print-tables-in-python.html
from shapely.geometry import Point,Polygon,MultiPoint
%matplotlib inline
import warnings
warnings.filterwarnings('ignore')
! pip install shapely
! pip install shapely[vectorized]
!pip install git-lfs
from scipy.stats import chisquare
data= pd.read_csv('Data_Sptember.csv')
data.head(10)
# # data
# # Firs Step . Download Dataset and Print out the size.
# if os.path.exists('Data_Sptember.csv'):# check if the Data is present and load it .
# # else: # Download Data Jan if not available on desktop .
# url = "https://s3.amazonaws.com/nyc-tlc/trip+data/green_tripdata_2019-01.csv"
# data= pd.read_url('url')
# data= Data.to_csv(url.split('/')[-1]
# define the figure with 2 subplots
fig,ax = plt.subplots(1,2,figsize = (15,4))
# Histogram of Number of Trip distance
data.Fare_amount.hist(bins=30 , ax = ax[0])
ax[0].set_xlabel('fare_amount (per mile)')
ax[0].set_ylabel('Count')
ax[0].set_yscale('log')
ax[0].set_title('A. Histogram of fare_amount With outliers included')
# Create a vector to contain Trip Distance
v = data.Trip_distance
# # Exclude any data point that is further than 4 standard deviations of median point
# and plot hist with 40 bin
v[~((v-v.median()).abs()>4* v.std())].hist(bins=40, ax= ax[1])
ax[1].set_xlabel('fare_amount (per mile)')
ax[1].set_ylabel('Trip Count')
ax[1].set_title(' g . Histogram of fare_amount without outliers')
# Apply a lognormal fit . use the mean of Trip Distancce as the scale parameter
Scatter,loc,mean = lognorm.fit(data.Trip_distance.values,
scale = data.Trip_distance.mean(),
loc = 0 )
pdf_fitted = lognorm.pdf(np.arange(0,14,.1), Scatter , loc, mean)
ax[1].plot(np.arange(0,14,.1), 500000 * pdf_fitted, 'r')
ax[1].legend (['Data', 'lognorm fit '])
# # # create a vector to contain Trip Distance
v = data.Trip_distance
# # exclude any data point located further than 3 standard deviations of the median point and
# # plot the histogram with 30 bins
# apply a lognormal fit. Use the mean of trip distance as the scale parameter
scatter,loc,mean = lognorm.fit(data.Trip_distance.values,
scale=data.Trip_distance.mean(),
loc=0)
pdf_fitted = lognorm.pdf(np.arange(0,12,.1),scatter,loc,mean)
ax[1].plot(np.arange(0,12,.1),600000 * pdf_fitted,'r')
ax[1].legend(['Lognormal_Fit', 'Data'])
# # export the figure
plt.savefig('Histograms.jpeg',format='jpeg')
plt.show()
```
The Trip Distance is asymmetrically distributed.It is skewed to the right and it has a median smaller than its mean and both smaller than the standard deviation. The skewness is due to the fact that the variable has a lower boundary of 0.The distance can't be negative. https://www.itl.nist.gov/div898/handbook/eda/section3/eda3669.htm
To the left is plotted the distribution of the entire raw set of Trip distance. To the right, outliers have been removed before plotting. Outliers are defined as any point located further than 3 standard deviations from the mean
The hypothesis: The trips are not random. If there were random, we would have a (symmetric) Gaussian distribution. The non-zero autocorrelation may be related the fact that people taking ride are pushed by a common cause, for instance, people rushing to work.
# We are going to examine if the time of the day has any impact on the trip distance.
```
data_median = data['Trip_distance'].median()
data_median
# First, convert pickup and drop off datetime variable in their specific righ format
data['lpep_pickup_dt'] = data.lpep_pickup_datetime.apply(lambda x:dt.datetime.strptime(x,"%m/%d/%y %H:%M"))
data['lpep_dropoff_dt'] = data.Lpep_dropoff_datetime.apply(lambda x:dt.datetime.strptime(x,"%m/%d/%y %H:%M"))
# Second, create a variable for pickup hours
data['Pickup_hour'] = data.lpep_pickup_dt.apply(lambda x:x.hour)
# # Mean and Median of trip distance by pickup hour
# # I will generate the table but also generate a plot for a better visualization
fig,ax = plt.subplots(1,figsize=(9,5)) # prepare fig to plot mean and median values
# use a pivot table to aggregate Trip_distance by hour
table1 = data.pivot_table(index='Pickup_hour',values='Trip_distance',aggfunc=('mean','median')).reset_index()
# # # rename columns
table1.columns =['Hour','Mean_distance','Average_distance']
table1[['Mean_distance','Average_distance']].plot(ax=ax)
plt.ylabel('Metric (miles)')
plt.xlabel('Hours after midnight')
plt.title('Distribution of trip distance by pickup hour')
plt.xticks(np.arange(0,30,6)+0.35,range(0,30,6))
plt.xlim([0,23])
plt.savefig('Question3_1.jpeg',format='jpeg')
plt.show()
print ('-----Trip distance by hour of the day-----\n')
print (tabulate(table1.values.tolist(),["Hour","Mean distance","Median distance", "Maximum distance"]))
# First, convert pickup and drop off datetime variable in their specific righ format
data['lpep_pickup_dt'] = data.lpep_pickup_datetime.apply(lambda x:dt.datetime.strptime(x,"%m/%d/%y %H:%M"))
data['lpep_dropoff_dt'] = data.Lpep_dropoff_datetime.apply(lambda x:dt.datetime.strptime(x,"%m/%d/%y %H:%M"))
# Second, create a variable for pickup hours
data['dropoff_hour'] = data.lpep_dropoff_dt.apply(lambda x:x.hour)
# # Mean and Median of trip distance by pickup hour
# # I will generate the table but also generate a plot for a better visualization
fig,ax = plt.subplots(1,figsize=(9,5)) # prepare fig to plot mean and median values
# use a pivot table to aggregate Trip_distance by hour
table1 = data.pivot_table(index='dropoff_hour',values='Trip_distance',aggfunc=('mean','median')).reset_index()
# # # rename columns
table1.columns =['Hour','Mean_distance','Average_distance']
table1[['Mean_distance','Average_distance']].plot(ax=ax)
plt.ylabel('Metric (miles)')
plt.xlabel('Hours after midnight')
plt.title('Distribution of trip distance by dropoff hour')
plt.xticks(np.arange(0,30,6)+0.35,range(0,30,6))
plt.xlim([0,23])
plt.savefig('Question3_1.jpeg',format='jpeg')
plt.show()
print ('-----Trip distance by hour of the day-----\n')
print (tabulate(table1.values.tolist(),["Hour","Mean distance","Median distance", "Maximum distance"]))
# Selecting airpor trips, finding the average of fare per trip, total amount charged.
airports_trips = data[(data.RateCodeID== 1) | (data.RateCodeID == 3)]
print("Number of Trips to/ from NYC airports :" ,airports_trips.shape[0])
print("Average of Fare (calculated by meter) of trips to / from NYC airports: $ ",airports_trips.Fare_amount.mean(),"per trip")
print("Average Total Amount charged bef ore the trips to / from NYC airports: $ ",airports_trips.Total_amount.mean(),"per trip")
print("Maximum of Fare (calculated by meter) of trips to / from NYC airports: $ ",airports_trips.Fare_amount.max(),"per trip")
```
Now that we have seen the number and mean fare of airport trips,
let's take a closer look at how trips are distributed by trip distances and hour of the day
```
##### Create a vector to contain Trip Distance .
v1 = airports_trips.Trip_distance
v2 = data.loc[~data.index.isin (v1.index),'Trip_distance'] # non- airport trips
# remove outliers :
# Exclude any data point located further than 2 standard deviations of the median point and histogram with 30 bins
v1 = v1[~((v1-v1.mean()).abs()>2*v1.std())]
v2 = v2[~((v2-v2.mean ()).abs()>2*v2.std())]
# # Define Bins bounries.
bins = np.histogram(v1, normed = True)[1]
h1 = np.histogram(v1, bins = bins , normed = True)
h2 = np.histogram(v2 , bins = bins , normed =True)
# # plots distribution of trips distance normilized among groups
fig,ax = plt.subplots(1,2,figsize = (15,5))
w =.4 *(bins[1] - bins[0])
ax[0].bar(bins[:-1], h1[0], alpha = 1 , width = w, color = 'y')
ax[0].bar(bins[:-1] + w, h2[0], alpha = 1 , width = w, color = 'orange')
ax[0].legend(['Airport Trip','Non-airport trips'] ,loc = 'best', title = 'Group')
ax[0].set_xlabel('Trip Distance(miles)')
ax[0].set_ylabel('Group Normalized Trips Count')
ax[0].set_title('A. Trip Distance Distribution')
airports_trips.Pickup_hour.value_counts(normalize=True).sort_index().plot(ax=ax[1])
data.loc[~data.index.isin(v1.index),'Pickup_hour'].value_counts(normalize=True).sort_index().plot(ax=ax[1])
ax[1].set_xlabel('Hours after midnight')
ax[1].set_ylabel('Group normalized trips count')
ax[1].set_title('B. Hourly distribution of trips')
ax[1].legend(['Airport trips','Non-airport trips'],loc='best',title='group')
plt.savefig('Question3_2.jpeg',format='jpeg')
plt.show()
```
# Predictive Model
In this section , I will take you through my analysis towards building a model to predict the percentage trip.
1. Let's build a derived variable for tips for a percentage of the total fare.
Before we proceed with this , some cleaning are necessary . Since the initial charge for NYC green is $2.5 ,
any transaction with a small total amount is invalid , thus it to be dropped.
```
data.head()
data = data[(data.Total_amount >= 2.5)] # Cleaning
data["Tip_percentage"] = 100 * data.Total_amount / data.Total_amount
print("Summary: Tip_percentage \n", data.Tip_percentage.describe())
```
2 . Similarly to the comparison between trips to /from airports with the rest of trips , it is worthy to spend more
time and check wether trips originating from Upper Manhattan have different percentage tip.
To identify trips originating from upper Manhattan :
* From google map , collect latitude and longitude data of atleast 12 points that approximately define
the bounding box of upper Manhattan .
*Create a polygon using shapely.geometry.Polygon :[https://pypi.org/project/Shapely/]
* Check if the polygon contain a location defined by (latitude, longitude)
```
# import library
from shapely.geometry import Point,Polygon,MultiPoint
# data points that define the bounding box of the Upper Manhattan
U_manhattan = [(40.796937, -73.949503),(40.787945, -73.955822),(40.782772, -73.943575),
(40.794715, -73.929801),(40.811261, -73.934153),(40.835371, -73.934515),
(40.868910, -73.911145),(40.872719, -73.910765),(40.878252, -73.926350),
(40.850557, -73.947262),(40.836225, -73.949899),(40.806050, -73.971255)]
print(U_manhattan)
# Point of origin
poi = Polygon(U_manhattan)
# poi
# create a function to check if a location is located inside Upper Manhattan
def is_within_bbox(loc,poi=poi):
"""
This function returns 1 if a location loc(lat,lon) is located inside a polygon of interest poi
loc: tuple, (latitude, longitude)
poi: shapely.geometry.Polygon, polygon of interest
"""
return 1*(Point(loc).within(poi))
tic = dt.datetime.now()
print(tic)
# # Create a new variable to check if a trip originated in Upper Manhattan
data['U_manhattan'] = data[['Pickup_latitude','Pickup_longitude']].apply(lambda r:is_within_bbox((r[0],r[1])),axis=1)
print("Processing Time :", dt.datetime.now() - tic)
v1 = data[(data.U_manhattan== 0) & (data.Tip_amount>0)].Tip_amount
v2 = data[(data.U_manhattan==1) & (data.Tip_amount>0)].Tip_amount
# generate bins and histogram values
bins = np.histogram(v1,bins=10)[1]
h1 = np.histogram(v1,bins=bins)
h2 = np.histogram(v2,bins=bins)
# generate the plot
# First suplot: visualize all data with outliers
fig,ax = plt.subplots(1,1,figsize=(10,5))
w = .4*(bins[1]-bins[0])
ax.bar(bins[:-1],h1[0],width=w,color='r')
ax.bar(bins[:-1]+w,h2[0],width=w,color='g')
ax.set_yscale('log')
ax.set_xlabel('Tip amount (%)')
ax.set_ylabel('Count')
ax.set_title('Tip')
ax.legend(['Non-Manhattan','Manhattah'],title='origin')
plt.savefig('Origin_Tip.jpeg' , format ='jpeg')
plt.show()
print ('t-test results:', ttest_ind(v1,v2,equal_var=False))
```
Then two distribution look the same however the t-test results in zero p-value to imply tha the groups are different
at 95 % level of comfidence
# The Model
# Summary
The initial dataset contained 1048575 transactions with 21 time-series, categorical and numerical variables. In order to build the final model, four phases were followed (1) data cleaning, (2) feature engineering (3) exploratory data analysis and (4) model creation
The cleaning consisted in drop zero variance variables (Ehail_fee), replacing invalid with the most frequent values in each categorical variable whereas the median was used for continuous numerical variables. Invalid values could be missing values or values not allowed for specific variables as per the dictionary of variables. In this part, variables were also converted in their appropriate format such datetime.
The feature engineering part created 10 new variables derived from pickup and dropoff locations and timestamps, trip distance.
During the exploration, each variable was carefully analyzed and compared to other variables and eventually the target variable, Percentage tip. All numerical variables were found to follow lognormal or power law distributions althouth there was found no linear relationship between numerical and the target variable. An interesting insight was uncovered in the distribution of the percentage tip. It was found that only 40% of the transactions paid tip. And 99.99% of these payments were done by credit cards. This inspired me to build the predictive model in two stages (1) classification model to find out weither a transaction will pay tip and (2) regression model to find the percentage of the tip only if the transaction was classified as a tipper. Another insight was that the most frequent percentage is 18% which corresponds to the usual restaurant gratuity rate.
With lack of linear relationship between independent and depend variables, the predictive model was built on top of the random forest regression and gradient boosting classifier algorithms implemented in sklearn after routines to optimize best parameters. A usable script to make predictions as attached to this notebook and available in the same directory.
Note: The code to make predictions is provided in the same directory as tip_predictor.py and the instructions are in the recommendation part of this section.
# Data Cleaning
```
# define a function to clean a loaded dataset
def clean_data(adata):
"""
This function cleans the input dataframe adata:
. drop Ehail_fee [99% transactions are NaNs]
. compute missing values in Trip_type
. replace invalid data by most frequent value for RateCodeID and Extra
. encode categorical to numeric
. rename pickup and dropff time variables (for later use)
input:
adata: pandas.dataframe
output:
pandas.dataframe
"""
## make a copy of the input
data = adata.copy()
## drop Ehail_fee: 99% of its values are NaNs
if 'Ehail_fee' in data.columns:
data.drop('Ehail_fee',axis=1,inplace=True)
## replace missing values in Trip_type with the most frequent value which is going to be 1
data['Trip_type '] = data['Trip_type '].replace(np.NaN,1)
## replace all values that are not allowed as per the variable dictionary with the most frequent allowable value
# remove negative values from Total amound and Fare_amount
print ("Negative values found and replaced by their abs")
print ("Total_amount", 100*data[data.Total_amount<10].shape[0]/float(data.shape[0]),"%")
print ("Fare_amount", 100*data[data.Fare_amount<10].shape[0]/float(data.shape[0]),"%")
print ("Improvement_surcharge", 100*data[data.improvement_surcharge<0].shape[0]/float(data.shape[0]),"%")
print ("Tip_amount", 100*data[data.Tip_amount<10].shape[0]/float(data.shape[0]),"%")
print ("Tolls_amount", 100*data[data.Tolls_amount<10].shape[0]/float(data.shape[0]),"%")
print ("MTA_tax", 100*data[data.MTA_tax<10].shape[0]/float(data.shape[0]),"%")
data.Total_amount = data.Total_amount.abs()
data.Fare_amount = data.Fare_amount.abs()
data.improvement_surcharge = data.improvement_surcharge.abs()
data.Tip_amount = data.Tip_amount.abs()
data.Tolls_amount = data.Tolls_amount.abs()
data.MTA_tax = data.MTA_tax.abs()
# # RateCodeID
indices_oi = data[~((data.RateCodeID>=1) & (data.RateCodeID<=6))].index
data.loc[indices_oi, 'RateCodeID'] = 2 # 2 = Cash payment was identified as the common method
print (round(100*len(indices_oi)/float(data.shape[0]),2),"% of values in RateCodeID were invalid.--> Replaced by the most frequent 2")
# # Extra
indices_oi = data[~((data.Extra==0) | (data.Extra==0.5) | (data.Extra==1))].index
data.loc[indices_oi, 'Extra'] = 0 # 0 was identified as the most frequent value
print (round(100*len(indices_oi)/float(data.shape[0]),2),"% of values in Extra were invalid.--> Replaced by the most frequent 0")
# # Total_amount: the minimum charge is 2.5, so I will replace every thing less than 2.5 by the median 11.76 (pre-obtained in analysis)
indices_oi = data[(data.Total_amount<2.5)].index
data.loc[indices_oi,'Total_amount'] = 11.76
print (round(100*len(indices_oi)/float(data.shape[0]),2),"% of values in total amount worth <$2.5.--> Replaced by the median 1.76")
# encode categorical to numeric (I avoid to use dummy to keep dataset small)
if data.Store_and_fwd_flag.dtype.name != 'int64':
data['Store_and_fwd_flag'] = (data.Store_and_fwd_flag=='Y')*1
# rename time stamp variables and convert them to the right format
print ("renaming variables...")
data.rename(columns={'lpep_pickup_datetime':'Pickup_dt','Lpep_dropoff_datetime':'Dropoff_dt'},inplace=True)
# Converting time stamp to the right format .
print("converting timestamps variables to right format ...")
try:
data['Pickup_dt'] = data.Pickup_dt.apply(lambda x :dt.datetime.strptime(x,"%m/%d/%y %H:%M"))
except TypeError:
print("Value of timestamp")
try:
data['Dropoff_dt'] = data.Dropoff_dt.apply(lambda x :dt.datetime.strptime(x, "%m/%d/%y %H:%M"))
except TypeError:
print("Value not in timestamp")
print ("Done cleaning")
return data
# Run code to clean the data
data = clean_data(data)
# data = clean_data(data)
```
# Feature Engineering
In this step, I intuitively created new varibles derived from current variables.
*Time variables: Week, Month_day(Day of month), Week_day (Day of week), Hour (hour of day), Shift_type (shift period of the day) and Trip_duration.The were created under the hypothesis that people may be willing to tip depending on the week days or time of the day. For instance, people are more friendly and less stressful to easily tip over the weekend. They were derived from pickup time
*.Trip directions: Direction_NS (is the cab moving Northt to South?) and Direction_EW (is the cab moving East to West). These are components of the two main directions, horizontal and vertical. The hypothesis is that the traffic may be different in different directions and it may affect the riders enthousiasm to tipping. They were derived from pickup and dropoff coordinates
*.Speed: this the ratio of Trip_distance to Trip_duration. At this level, all entries with speeds higher than 240 mph were dropped since this is the typical highest speed for cars commonly used as taxi in addition to the fact that the speed limit in NYC is 50 mph. An alternative filter threshold would be the highest posted speed limit in NYC but it might be sometimes violated.
*.With_tip: This is to identify transactions with tips or not. This variable was created after discovering that 60% of transactions have 0 tip.
*****.As seen that using the mean of trips from Manhattan is different from the mean from other boroughs, this variable can be considered as well in the model building. A further and deep analysis, would be to create a variable of the origin and destination of each trip. This was tried but it was computationally excessive to my system. Here, coming from Manhattan or not, is the only variable to be used.
```
# Function to run the feature engineering
def engineer_features(data):
"""
This function create new variables based on present variables in the dataset adata. It creates:
. Week: int {1,2,3,4,5}, Week a transaction was done
. Week_day: int [0-6], day of the week a transaction was done
. Month_day: int [0-30], day of the month a transaction was done
. Hour: int [0-23], hour the day a transaction was done
. Shift type: int {1=(7am to 3pm), 2=(3pm to 11pm) and 3=(11pm to 7am)}, shift of the day
. Speed_mph: float, speed of the trip
. Tip_percentage: float, target variable
. With_tip: int {0,1}, 1 = transaction with tip, 0 transction without tip
input:
adata: pandas.dataframe
output:
pandas.dataframe
"""
# make copy of the original dataset
data = data.copy()
# derive time variablesntag
print("deriving time variables...")
# reference week "First week of Septemeber in 2015"
# date(2003, 12, 29).isocalendar()
ref_week = dt.datetime(2015,9,1).isocalendar()[1]
data['Week'] = data.Pickup_dt.apply(lambda x:x.isocalendar()[1]- ref_week + 1 )
data['Week_day'] = data.Pickup_dt.apply(lambda x:isocalendar()[2])
data['Month_day'] = data.Pickup_dt.apply(lambda x:x.day)
data.rename(columns ={'Pick_hour':'Hour'} , inplace =True)
data['Hour'] = data.Pickup_dt.apply(lambda x:x.hour)
# Create shift variable: 1= (7am to 3 pm) , 2 =(3pm to 11 pm) , 3 = (11 pm to 7 am )
data['Shift_type']=np.NAN
data.loc[data[(data.Hour >=7) & (data.Hour<15)].index,'Shift_type'] = 1
data.loc[data[(data.Hour >=15) & (data.Hour<23)].index,'Shift_type'] = 2
data.loc[data[data.Shift_type.isnull()].index,'Shift_type'] = 3
# Trip Duration
print("Deriving Trip_duration ....")
data['Trip_duration'] = ((data.Dropoff_dt - data.Pickup_dt).apply(lambda x:x.total_seconds()/60.))
print("Deriving direction variables...")
# Create direction variables Direction_North_South (NS)
# This is 2 if Taxi moving frm North to South , 1 in the opposite direction and = 0 otherwise.
data['Direction_NS'] = (data.Pickup_latitude > data.Dropoff_latitude)* 1+1
indices = data[(data.Pickup_latitude == data.Dropoff_latitude) & (data.Pickup_latitude !=0)].index
try:
data.loc[indeces, 'Direction_NS'] = 0
except UnboundLocalError:
print('local variable indeces referenced before assignment')
# Create direction variables of Direction_EW(East to West)
data["Direction_Ew"] = (data.Pickup_longitude > data.Dropoff_longitude) * 1+1
indeces = data[(data.Pickup_longitude ==data.Dropoff_longitude) & (data.Pickup_longitude !=0)].index
data.loc[indeces,'Direction_EW'] = 0
# Create a variable for speed
print("deriving Speed. make sure to check for possible NANS and inf vals...")
data['Speed_mph'] = data.Trip_distance / (data.Trip_duration / 60)
# replace all NANs values and values > 240mph by values samples from a random distribution of mean 12.9 and standard deviation
# 6.8 mph. These Values were extracted from the distribution
indeces_oi = data[(data.Speed_mph.isnull()) / (data.Speed_mph>240)].index
data.loc[indeces_oi, 'Speed_mph'] = np.abs(np.random.normal(loc=12.9, scale =6.8, size= len(indices_oi)))
print("Greate we are done with Feature Engineering ! :-")
# Create a new a variable to check if the trip originated in Upper Manhattan
print("Checking where the trip originated")
data[U_manhattan] = data[['Pickup_latitude','Pickup_longitude']]. apply(lambda r: is_within_bbox((r[0], r[1])), axis=1)
# create a tip percentage variable
data['Tip_percentage'] = 100*data.Tip_amount / data.Total_amount
# Create with tip variable
data['With_tip'] =(data.Tip_percentage > 0)*1
return data
# collected bounding box points
umanhattan = [(40.796937, -73.949503),(40.787945, -73.955822),(40.782772, -73.943575),
(40.794715, -73.929801),(40.811261, -73.934153),(40.835371, -73.934515),
(40.868910, -73.911145),(40.872719, -73.910765),(40.878252, -73.926350),
(40.850557, -73.947262),(40.836225, -73.949899),(40.806050, -73.971255)]
poi = Polygon(umanhattan)
# create a function to check if a location is located inside Upper Manhattan
def is_within_bbox(loc,poi=poi):
"""
This function checks if a location loc with lat and lon is located within the polygon of interest
input:
loc: tuple, (latitude, longitude)
poi: shapely.geometry.Polygon, polygon of interest
# """
return 1*(Point(loc).within(poi))
# return the code to create a new features on the dataset
# There is a bug here that I am trying to fix . error message : Isocalendar not defined .
# Excepected the size to change after feature engineering
print("Size before feature engineering:", data.shape)
# data = engineer_features(data)
print("Size after feature engineering:",data.shape)
# Uncomment to check for data validity.
data.describe().T
```
# Exploratory Data Analysis
This was the key phase of the analysis .
* A look at the distribution of the target
variable , Tip_percentage showed that 60 % of all transaction did not give tip(See Figure below , left).
A second tip at 18 % corresponds to the usual NYC customary
gratuity rate which fluctuates between 18 % and 25 % (See Figure below , right).
Based on the information , the model ca be bult in two steps .
* First Step: Create classification model to predict wether tip will be given or not . Here a new variable " with_tip" of 1 (if there is tip ) and 0 (otherwise) was created.
* Second Create regression model for transaction with non-zero tip.
The two distributions look the same however the t-test results in a zero p-value to imply that the two groups are different at 92% level of confidence.
```
data["Tip_percentage"] = 100 * data.Tip_amount/ data.Total_amount
print(data["Tip_percentage"])
# Code to compare the two tips_percentage indentifed groups
## code to compare the two Tip_percentage identified groups
# split data in the two groups
data1 = data[data.Tip_percentage >0]
data2 = data[data.Tip_percentage ==0]
fig ,ax = plt.subplots(1,2,figsize =(14,4))
data.Tip_percentage.hist(bins=20, normed = True , ax=ax[0] , color = 'g')
ax[0].set_xlabel('Tip(%)')
ax[0].set_title('Distribution of fare_amount (%) - all transaction')
data1.Tip_percentage.hist(bins =20, normed = True, ax=ax[1])
ax[1].set_xlabel('Tip(%)')
ax[1].set_title('Distribution of the tip (%) - transaction with tips')
ax[1].set_ylabel('Group normed count')
plt.show()
```
Next , each variable distribution and its relationship with the Tip
percentage were explored. Few functions were implimented to quickly explore those variables.
```
df = data
df.head()
# Functions For Exploratory Data Analysis
def visualize_contious(df,label,method={'type':'histogram','bins':20},outlier ='on'):
"""
The function to quickly visualize continuos variables
df:pandas.dataFrame
label:str, name of variable to be plotted . it should be present in df.columns
method:dict, contains info of the type of plot to generate . It can be histogram or boxplot
[-Not yet developped]
outlier:{'on','off'}, set it off you need to cut off outliers .
Outliers are all those points
Located at 3 standard deviations further from the mean.
"""
# Create Vector of variable of interest
v = df[label]
# Define mean and standard deviation
m = v.mean()
s = v.std()
# set up fig for plotting
fig, ax = plt.subplots (1, 2, figsize=(14,4))
ax[0].set_title('Distribution of' + label)
ax[1].set_title('Tip % by '+ label)
# Let`s set the condition of if statemment
if outlier == 'off': # remove outliers accondingly and update titles
v = v[(v-m) < 3 * s]
ax[0].set_title('Distribution of' + label + '(no outliers)')
ax[1].set_title('Tip % by' + label+'(no outliers)')
if method['type'] == 'histogram': # plot the histogram
v.hist(bins = method['bins'], ax=ax[0])
if method['type'] == 'boxplot' : # plot the box plot
df.loc[v.index].boxplot(label, ax =ax[0])
ax[1].plot(v,data.loc[v.index].Tip_percentage ,'.',alpha = 0.4)
ax[0].set_xlabel(label)
ax[1].set_xlabel(label)
ax[0].set_ylabel('Count')
ax[1].set_ylabel('Tip (%)')
plt.show()
def visualize_categories(df, catName ,chart_type = 'Histogram',ylimit =[None,None]):
"""
This Functions Helps to quickly visualize categorical variables .
this functions calls other functions generate_boxplot and generate_histogram
df:pandas.DataFrame
chart_type : {histogram, boxplot}, choose which type of chart to plot
ylim:tuple, list.Valid if chart_type is histogram
"""
print(catName)
cats = sorted(pd.unique(df[catName]))
if chart_type == 'boxplot': # generate boxplot
generate_boxplot(df, catName , ylimit)
elif chart_type == 'histogram': # generate histogram
generate_histogram(df, catName)
else:
pass
# calculate test Stastics
groups = df[[catName,'Tip_percentage']].groupby(catName).groups # create groups
tips = df.Tip_percentage
if len(cats) <= 'catName': # if there are only two groups use t-test
print(ttest_ind(tips[groups[cats[0]]], tips[groups[cats[1]]]))
else: # otherwise the command to be evaluated
cmd = "f_oneway("
for cat in cats :
cmd +="tips[groups["+str(cat)+"]],"
cmd = cmd[:-1]+")"
print("one way anova test:" , eval(cmd)) # Evaluate the command and print
print("frequency of categories (%):\n", df[catName].value_counts(normalize = True)*100)
def test_classification(df, label, y1=[0,50]):
"""
This Function test if the means of the two groups with_tip and without tip are different
at 95 % of confidence level.
It will also generate a box plot of the viariable by tipping groups
label:str ,label to test
y1:tuple or list (default = [0,50]) , y limits on the ylabel of the boxplot
df:pandas.DataFrame(deafult =data)
Example:run <visualize_continous(data,'Fare_amount', outlier ='on ') >
"""
if len(pd.unique(df[label])) == 2: # check if the variable is categorical with only two
# categories and run the test .
vals = pd.unique(df[label])
gp1 =df[df.With_tip == 0][label].value_counts().sort_index()
gp2 =df[df.With_tip == 1][label].value_counts().sort_index()
print("t-test if ", label , "can be used to distinguish transanctionwith tip and without tip")
print("chisquare(gp1,gp2)")
elif len(pd.unique(df[label])) >= 10 : # otherwise run the t-test
df.boxplot(label, by='With_tip')
plt.ylim(y)
plt.show
print("t-test if, label , can be used to distinguish transaction with tip and without " )
print("results:", ttest_ind(df[df.WIth_tip == 0][label].value, df[df.With_tip ==1][label].values , False))
else:
passs
def generate_boxplot(df,catName,ylimit):
"""
generate boxplot of tip percentage by variable "catName" with ylim set ylimit
df:pandas.Dataframe
catName :str
ylimit :tuple , list
"""
df.boxplot('Tip_percentage', by=catName)
plt.title('Tip % by ' +catName )
plt.title('')
plt.ylabel('Tip (%)')
if ylimit != [None, None]:
plt.ylim(ylimit)
plt.show()
def generate_histogram(df, catName):
"""
generate histogram of tip percentage by varible "CatName" with ylimit set ylimit
df: pandas.dataFrame
catName : str
ylimit : tuple , list
"""
cats = sorted(pd.unique(df[catName]))
colors =plt,cm.jet(np.linspace(0,1,len(cats)))
hx = np.array(map(lambda x : round(x, 1 ), np.histogram(df.Tip_percentage,bins=20)[1]))
fig , ax = plt.subplots(1,1,figsize=(15, 4))
for i ,cat in enumerate (cats):
vals =df [df[catName] == cat ].Tip_percentage
h=np.histogram(vals, bins =hx)
w =0.9*(hx [:1]+ w*i, h[0], colors == colors[i], width == w)
plt.legend(cats)
plt.ysccale('log')
plt.title ('Distribution of Tip by' + catName )
plt.xlabel ('Tip (%)')
plt.show()
```
Starting with continuous variables, two main insights were discovered:
A lognormal-like or power law distribution of the Fare amount and a non-linear function of the tip percentage as function
of the total amount. The tip percentage decreases as the fare amount increases but converges around 20%.
The density of scattered points implies that there is a high frequency of smaller tipps at low Fare_amount.
Can we say that people restrain themselves to tipping more money as the cost of the ride becomes more and more expensive?
Or since the fare grows with the length of the trip and trip duration, can we say that riders get bored and
don't appreciate the service they are getting? Many questsions can be
explored at this point.
```
# Example of exploration of the Fare_amount using the implemented:
visualize_contious(df , 'Fare_amount', outlier ='on')
try:
test_classification(df,'Fare_amount',[10,25])
except KeyError :
print("raised With_tip while passing the value ")
except NameError:
print('Name y function is not defined ')
plt.savefig('DistribFAmount.png', format ='png')
plt.show()
# """
# This Function test if the means of the two groups with_tip and without tip are different
# at 95 % of confidence level.
# It will also generate a box plot of the viariable by tipping groups
# label:str ,label to test
# y1:tuple or list (default = [0,50]) , y limits on the ylabel of the boxplot
# df:pandas.DataFrame(deafult =data)
# Example:run <visualize_continous(data,'Fare_amount', outlier ='on ') >
# """
def test_classification(df,label, yl=[0,50]):
# check if the variable is categorical with only two categories and run the test .
if len(pd.unique(df[label])) == 2:
vals = pd.unique(df[label])
gp1 = df[df.with_tip== 0][label].value_counts().sort_index()
gp2 = df[df.with_tip == 1][label].value_counts().sort_index()
print("t-test if ",label,"can be used to distinguish transanction with tip and without tip")
print("chisquare(gp1,gp2)")
elif len(pd.unique(df[label])) >= 10 : # otherwise run the t-test
df.boxplot(label, by='Fare_amount')
plt.ylim(y)
plt.show
print("t-test if, label , can be used to distinguish transaction with tip and without " )
print("results:", ttest_ind(df[df.WIth_tip == 0][label].value, df[df.With_tip ==1][label].values , False))
else:
passs
try:
test_classification(data,'Fare_amount',[0,25])
except NameError :
# continue
print("Fare_amount")
```
A negative t-test value and null p-value imply that the means of Total_amount are significantly different in the group of transactions with tips compared to the group with no tip. Therefore,
this variable would used to train the classification model.
Using the same function, a plot of the tip percentage as function of trip duration showed a cluster of points at duration time greater than 1350 min (22 hours).
These points look like outliers since it doesn't make sense to have a trip of 22 hours within NYC. Probably, tourists can!
The following code was used to analyze the cluser with trip duration greater than 1350 min
```
data = data.rename(columns ={'Lpep_dropoff_datetime':'Dropoff_dt','lpep_pickup_datetime':'Pickup_dt'})
data.head()
data['Trip_Duration'] = ((data.Dropoff_dt - data.Pickup_dt).apply(lambda x:x.total_seconds()/60))
data['Trip_Duration']
# Code to generate the heat map to uncover hidden information in the cluster
# We will first source NYC boroughs shape files,
# then create polygons and check to which polygon does each of the point of the cluster begongs
# download geojson of NYC boroughs
nyc_boros = json.loads(requests.get("https://raw.githubusercontent.com/dwillis/nyc-maps/master/boroughs.geojson").content)
# parse boros into Multipolygons
boros = {}
for f in nyc_boros['features']:
name = f['properties']['BoroName']
code = f['properties']['BoroCode']
polygons = []
for p in f['geometry']['coordinates']:
polygons.append(Polygon(p[0]))
boros[code] = {'name':name,'polygon':MultiPolygon(polygons=polygons)}
# creae function to assign each coordinates point to its borough
def find_borough(lat,lon):
"""
return the borough of a location given its latitude and longitude
lat: float, latitude
lon: float, longitude
"""
boro = 0 # initialize borough as 0
for k,v in boros.items(): # update boro to the right key corresponding to the parent polygon
if v['polygon'].contains(Point(lon,lat)):
boro = k
break # break the loop once the borough is found
return [boro]
## Analyse the cluster now
# create data frame of boroughs
df = data[data.Trip_Duration >= 1300]
orig_dest = []
for v in df[['Pickup_latitude','Pickup_longitude','Dropoff_latitude','Dropoff_longitude']].values:
orig_dest.append((find_borough(v[0],v[1])[0],find_borough(v[2],v[3])[0]))
df2 = pd.DataFrame(orig_dest)
## creae pivot table for the heat map plot
df2['val']= 1 # dummy variable
# try:
mat_cluster1 = df2.pivot_table(index =0, columns=1 ,values ='val',aggfunc = 'count')
# except KeyError:
# pass
# # generate the map
fig,ax= plt.subplots(1,2,figsize=(15,4))
im = ax[0].imshow(mat_cluster1)
ax[0].set_ylabel('From')
ax[0].set_xlabel('To')
ax[0].set_xticklabels(['','other', 'Manhattan','Bronx','Brooklyn','Queens'], rotation= 'vertical')
ax[0].set_yticklabels(['','other','Manhattan','Bronx','Brooklyn','Queens'])
ax[0].set_title('Cluster of rides with duration > 1300 min')
fig.colorbar(im,ax=ax[0])
h = df.Pickup_hour.value_counts(normalize=True)
plt.bar(h.index,h.values,width =.4,color='y')
h = data1.Pickup_hour.value_counts(normalize=True)
ax[1].bar(h.index +.4,h.values,width=.5,color='b')
ax[1].set_title('Hourly traffic: All rides vs cluster rides')
ax[1].legend(['cluster','all'],loc='best')
ax[1].set_xlabel('Hour')
ax[1].set_xticks(np.arange(25)+.4,range(25))
ax[1].set_ylabel('Normalized Count')
plt.savefig('duration_cluster.jpeg',format='jpeg')
plt.show()
```
The heat map color represents the number of trips between two given
bouroghs. We can see that the majority of the trips are intra-boughs.
There is a great number of trips from brooklyn to manhattan whereas
there is no staten Island trip that takes more than 1300 minutes.
are there specific hours for these events?
Unfortunetely , the distribution on the right shows that the cluster
behaves the same as the rest of the traffic.
Finally , correlation heatmap was used to find which independent variables
are correlated to each other . The following code provides the construction
of the correlation heatmap.
```
continous_variables = ['Total_amount','Fare_amount','Trip_distance','Tolls_amount', 'Tip_percentage']
cor_mat = data1[continous_variables].corr()
# fig , ax = plt.subplots(1,1,figsize =[10,10])
# fig,ax = plt.subplots(1,2 figsize =[6,6])
plt.imshow(cor_mat )
plt.xticks(range(len(continous_variables )), continous_variables,rotation = 'vertical')
plt.yticks(range(len(continous_variables )),continous_variables)
plt.colorbar()
plt.title('Correration Between Continous_Variables')
plt.show()
# data1
```
A further analysis of all continuous variables revealed
similar lognormal and non-linearlity behaviors.
Since there is no linear relationship between the Tip percentage and variables, random forest algorithm will be considered to build
the regression part of the model
As far as categorical variables concerned, the function
visualize_categories was used to explore each variable as
it was done for
continuous numerical variables (See demostration below)
# Building A Model
As explained in the previous section , this model will be a combination
of rules from two models (1) the classification model to classify a transaction
into a tipper (=1) or not (=0) and (2) regression model to estimate the percentage
of the tip.
```
#import scikit learn libraries
from sklearn import cross_validation , metrics # MODEL OPTIMIZATION AND VALUATION TOOLS
from sklearn.grid_search import GridSearchCV # PERFORM GRID SEARCH
# define a function that help to train model and perform cv
def modelfit(alg, dtrain, predictos ,target,scoring_method, performCV=True, printFeatureImportance =True , CV_folds =5):
"""
This function train the model given as 'alg' which stannd by algorithm by performing cross-validation .
It works on both regression and classification alg:sklearn model
dtrain: pandas.DataFrame , training set
predictors : list,labels to be used in the model training process . They should be in the columns name of dtrain
target: str, target variable
scoring_method :str , method to be used by in the cross -validation to valuate the model
performCV: bool, perform Cv or not
printFeatureImportance: bool, plot histogram of features importance or not
cv_folds: int, degree of cross-validation
"""
# train the algorithm on data
alg.fit(dtrain[predictors],dtrain[target])
# predict on train set :
dtrain_predictions = alg.predict(dtrain[predictors])
if scoring_method =='roc_auc':
dtrain_predprob= alg.predict_proba(dtrain[predictors])[:,1]
# peform cross-validation
if performCV:
cv_score = cross_validation.cross_val_score(alg, dtrain[predictors],dtrain[target] ,
cv=cv_folds, scoring =scoring_method)
# print model report
print ("\nModel report:")
if scoring_method =='roc_auc':
print ("Accuracy:", metrics.accuracy_score(dtrain[target].values, dtrain_predictions))
print ("AUC Score(Train):", metrics.roc_auc_score(dtrain[target], dtrain_predprob))
if (scoring_method == 'mean_squared_error'):
print ("Accuracy:", metrics.mean_squared_error(dtrain[target].values, dtrain_predictions))
if performCV:
print ("CV Score - Mean: %.7g | std :%.7g| Min:%.7 |Max:%.7g:"%(np.mean(cv_score),np.std(cv_score),np.min(cv_score),np.max(cv_score)))
#print feature importance
if printFeatureImportance:
if dir(alg)[0] == '_Booster': #runs only if alg is xgboost
feat_imp = pd.Series(alg.booster().get_fscore()).sort_values(ascending=False)
else:
feat_imp = pd.Series(alg.feature_importances_,predictors).sort_values(ascending=False)
feat_imp.plot(kind='bar',title='Feature Importances')
plt.ylabel('Feature Importe Score')
# print("I am here")
plt.show()
# optimize n_estimator through grid search
def optimize_num_trees(alg,param_test,scoring_method,train,predictors,target):
"""
This functions is used to tune paremeters of a predictive algorithm
alg: sklearn model,
param_test: dict, parameters to be tuned
scoring_method: str, method to be used by the cross-validation to valuate the model
train: pandas.DataFrame, training data
predictors: list, labels to be used in the model training process. They should be in the column names of dtrain
target: str, target variable
"""
gsearch = GridSearchCV(estimator=alg, param_grid = param_test, scoring=scoring_method,n_jobs=2,iid=False,cv=5)
gsearch.fit(train[predictors],train[target])
return gsearch
# plot optimization results
def plot_opt_results(alg):
cv_results = []
for i in range(len(param_test['n_estimators'])):
cv_results.append((alg.grid_scores_[i][1],alg.grid_scores_[i][0]['n_estimators']))
cv_results = pd.DataFrame(cv_results)
plt.plot(cv_results[1],cv_results[0])
plt.xlabel('# trees')
plt.ylabel('score')
plt.title('optimization report')
```
Classification Model
while I spent time on feature exploration, engineering and discovering that the Payment_type was a strong variable (99.99% of all transactions with tips were paid with credit cards) to differentiate transactions with tip from those without tip, a model based on the logistic regression classifier algorithm was optimized and gave an accuracy score of 0.94.
However, there was away to in crease accuracy score using the GradientBoostingClassifier : The idea behind "gradient boosting" is to take a weak hypothesis or weak learning algorithm and make a series of tweaks to it that will improve the strength of the hypothesis/learner using scikit learn which improved our acuracy score to be 0.96. Starting with the GradientBoostinClassier model (default paremeters), the number of trees was optimized through a grid search (see function 'optimize_num_trees').
References: https://stackabuse.com/gradient-boosting-classifiers-in-python-with-scikit-learn/
*-- Key points --*:
- Sample size for training and optimization was chosen as 100000. This is surely a small sample size compared to the available data but the optimization was stable and good enough with 5 folds cross-validation
- Only the number of trees were optimized as they are the controlling key of boosting model accuracy. Other parameters were not optimized since the improvement yield was too small compared to the computation time and cost
- ROC-AUC (Area under the curve of receiver operating characteristic) was used as a model validation metric
*-- Results --*:
- optimized number of trees: 130
- optimized variables: ['Payment_type','Total_amount','Trip_duration','Speed_mph','MTA_tax','Extra','Hour','Direction_NS', 'Direction_EW','U_manhattan']
- roc-auc on a different test sample: 0.9636
The following code will cover in depth optimization process:
```
# data
# Optimization and Traing of classifier
from sklearn.ensemble import GradientBoostingClassifier
print ("Optimazing classifier...")
train = data.copy () #Make a copy of the train set
# I will only use a small size to carry on my traing and 5 folds cross-validation since my dataset is too big.
train = train.loc[np.random.choice(train.index, size=100000 , replace =False)]
target = 'With_tip' # set target variable - it will be used later during the optimization
tic = dt.datetime.now()
# intiate the training
# will start with candidate that we identified ealier during the EDA for predictors
predictors = data[['Payment_type','Total_amount','Trip_Duration','Speed_mph','MTA_tax',
'Extra','Hour','Direction_NS', 'Direction_EW','U_manhattan']]
# optimize n_estimetor through grid search
parameter_test = {'n_estimators':range (30,151,20)} # define range over whicjh numbert of trees is to be optimezed
# Initialize classification model
model_cls = GradientBoostingClassifier(
learning_rate =0.1, #use default
min_samples_split =2 , # use default
max_features = 'auto',
subsample = 0.8,
random_state =10)
# try < 1 to decrease variance anad increse biase
# Bias is the assumptions made by the model to make the target function easier to approximate.
# Variance is the amount that the estimate of the target function will change given different training data
# get the results of the search grid
gs_cls = optimize_num_trees(models_cls, param_test, 'roc_auc',train, predictors , target)
print ("gs_cls:", gs_cls.grid_scores_, gs_cls.best_params_,gs_cls.best_socre_)
# cross validate the best model with optimized number of estimators
modelfit(gs_cls.best_estimator_,train,predictors,target,'roc_auc')
# save the best estimator on disk as pickle for a later use
with open('my_classifier.pkl','wb') as fid:
pickle.dump(gs_cls.best_estimator_,fid)
fid.close()
print("Processing time:", dt.datetime.now()-tic)
```
| github_jupyter |
```
import pickle
import pandas as pd
from sklearn.manifold import TSNE
import matplotlib.pyplot as plt
import config
import os
import data_utils
from adjustText import adjust_text
experiment_name = 'test_h1_record'
file_name = '20190302201128_init_all_correct.pickle'
# Read the pickle file. ===============================================
file_path = os.path.join(config.dir_h1_logs(), experiment_name, file_name)
with open(file_path, 'rb') as f:
data = pickle.load(f)
print('Read {}'.format(file_path))
# t-SNE training =======================================================
# t-SNE hyperparmeter setting
tsne = TSNE(n_components=2, perplexity=5.0, n_iter=100000, init='pca')
# t-SNE fitting
X_2d = tsne.fit_transform(data['h1'])
# Create DataFrame containing data to plot. ============================
# Create annotation texts.
n_examples = data['h1'].shape[0]
op_texts = list()
for i in range(n_examples):
op_text = '[{carries}] {str_operation}'.format(
carries=data['carry'][i],
str_operation=data_utils.np_io2str_op(data['input'][i], data['output'][i], data['operator'])
)
op_texts.append(op_text)
# Create DataFrame
df = pd.DataFrame({
'x':X_2d[:,0],
'y':X_2d[:,1],
'carries':data['carry'],
'op_text':op_texts
})
# Plot section ==========================================================
# [Plot 1] Carry scatter plot ===========================================
# figure configuartion
plt.figure(figsize=(10,10))
pad = 2
plt.xlim(df['x'].min() - pad, df['x'].max() + pad)
plt.ylim(df['y'].min() - pad, df['y'].max() + pad)
plt.title('h1 visualization using the t-SNE method\n{}'.format(file_path))
# scatter plot
for carries in df['carries'].unique():
df_carries = df[df['carries'] == carries]
plt.plot(df_carries['x'], df_carries['y'], 'o', label=carries)
plt.legend(title='Carries')
plt.show()
# [Plot 2] Plot with annotations ===========================================
# figure configuartion
plt.figure(figsize=(70,70))
pad = 2
plt.xlim(df['x'].min() - pad, df['x'].max() + pad)
plt.ylim(df['y'].min() - pad, df['y'].max() + pad)
plt.title('h1 visualization using the t-SNE method with annotations\n{}'.format(file_path))
# scatter plot
for carries in df['carries'].unique():
df_carries = df[df['carries'] == carries]
plt.plot(df_carries['x'], df_carries['y'], 'o', label=carries)
# tag labels
texts = list()
for i in range(len(df)):
text = plt.text(df['x'][i], df['y'][i], df['op_text'][i], ha='center', va='center')
texts.append(text)
adjust_text(texts, arrowprops=dict(arrowstyle='-', color='red'))
plt.legend(title='Carries')
plt.show()
```
| github_jupyter |
# 005_lists
[Source](https://github.com/iArunava/Python-TheNoTheoryGuide/)
```
# Simple Lists
names = ["Jennifer", "Python", "Scarlett"]
nums = [1, 2, 3, 4, 5]
chars = ['A', 'q', 'E', 'z', 'Y']
print (names)
print (nums)
print (chars)
# Can have multiple data types in one list
rand_list = ["Jennifer", "Python", "refinneJ", 'J', '9', 9, 12.90, "Who"]
print (rand_list)
# Accessing elements in a list
# O-indexed
print (names[2])
print (rand_list[3])
print (names[0] + " " + rand_list[2].title())
# Negetive indexes: Access elements from the end of the list without knowing the size of the list
print (rand_list[-1]) # Returns the last element of the list [1st from the end]
print (rand_list[-2]) # Returns the 2nd last element
# and so on..
# Now here's a question.
print (rand_list[-1] + " is " + names[2] + "?")
print ("A) " + rand_list[0] + "'s sister\tB) " + names[0] + "'s Friend\nC) Not Related to " + rand_list[-8] + "\tD) Nice question but I don't know")
# Modifying elements in a list
str_list = ["Scarlett", "is", "a", "nice", 'girl', '!']
print (str_list)
str_list[0] = "Jennifer"
print (str_list)
# Adding elements to a list
# Use append() to add elements to the end of the list
str_list.append ('She is 21.')
print (str_list)
# So, you can build lists like this
my_list = []
my_list.append ("myname")
my_list.append ("myage")
my_list.append ("myaddress")
my_list.append ("myphn")
my_list.append ("is")
my_list.append (1234567890)
print (my_list)
# Insert elements at specific positions of the list
# insert(index, element)
my_list.insert (0, "Mr/Miss/Mrs")
print (my_list)
my_list.insert(4, "mybday")
print (my_list)
# Using '-1' to insert at the end doesn't work and inserts element at the 2nd last position.
my_list = ['A', 'B', 'C', 'D']
my_list.insert (-1, 'E')
print (my_list)
# Using '-2' inserts at 3rd last position
# In general, use '-n' to insert at 'n+1'th position from end.
my_list = ['A', 'B', 'C', 'D']
my_list.insert (-2, 'E')
print (my_list)
# Insert elements at the end
l1 = ['A', 'B', 'C', 'D']
l2 = ['A', 'B', 'C', 'D']
l1.append('E')
l2.insert(len(my_list), 'E')
print (l1)
print (l2)
# Length of the list
l1 = ['A', 'B', 'C', 'D', 'E']
print (len(l1))
# # Removing elements from list
# del can remove any element from list as long as you know its index
l1 = ['A', 'B', 'C', 'D', 'E']
print (l1)
del l1[0]
print (l1)
del l1[-1]
print (l1)
# pop() can remove the last element from list when used without any arguments
l1 = ['A', 'B', 'C', 'D', 'E']
# pop() returns the last element, so c would store the popped element
c = l1.pop()
print (l1)
print (c)
# pop(n) -> Removes the element at index 'n' and returns it
l1 = ['A', 'B', 'C', 'D', 'E']
# Removes the element at 0 position and returns it
c = l1.pop(0)
print (l1)
print (c)
# Works as expected with negetive indexes
c = l1.pop(-1)
print (l1)
print (c)
# Removing an item by value
# remove() only removes the first occurence of the value that is specified.
q1 = ["Seriously, ", "what", "happened", "to", "Jennifer", "and", "Jennifer", "?"]
print (q1)
q1.remove ("Jennifer")
print (q1)
n1 = "and"
q1.remove(n1)
print (q1)
# Sorting a list
# sort() -> sorts list in increasing or decreasing order, *permantantly*
# Sorts in alphabetical order
l1 = ['E', 'D', 'C', 'B', 'A']
l1.sort()
print (l1)
# Sorts in increasing order
l2 = [2, 200, 16, 4, 1, 0, 9.45, 45.67, 90, 12.01, 12.02]
l2.sort()
print (l2)
# Reverse sorts alphabetical order
l1 = ['E', 'D', 'C', 'B', 'A']
l1.sort(reverse=True)
print (l1)
# Sorts in decreasing order
l2 = [2, 200, 16, 4, 1, 0, 9.45, 45.67, 90, 12.01, 12.02]
l2.sort(reverse=True)
print (l2)
# sorted() -> Sorts list in increasing or decreasing order, *temporarily*
# Sorts in increasing order
l2 = [2, 200, 16, 4, 1, 0, 9.45, 45.67, 90, 12.01, 12.02]
print (l2)
print (sorted(l2))
print (l2)
# Sorts in decreasing order
l2 = [2, 200, 16, 4, 1, 0, 9.45, 45.67, 90, 12.01, 12.02]
print (l2)
print (sorted(l2, reverse=True))
print (l2)
# Reverse list
l1 = ['E', 'D', 'C', 'B', 'A']
l1.reverse()
print (l1)
# Looping Through a list using for
l1 = ["Scarlett", "is", "now", "back", "from", "her first", "Python", "lesson."]
# Do notice the indentations
for each_word in l1:
print (each_word)
# Looping through a list using while
l1 = ["Scarlett", "is", "in", "love", "with", "Python"]
i = 0
while i is not len(l1):
print (l1[i])
i += 1
# Numerical lists
# Note: range(n, m) will loop over numbers from n to m-1
l1 = ['A', 'B', 'C', 'D', 'E']
print ("Guess how much Scarlett scored in her first lesson out of 5:")
for val in range(1, 6):
print (l1[val-1] + ") " + str(val))
# Using range() to make a list of numbers
num_list = list(range(1, 6))
print (num_list)
# Use range() to skip values at intervals
# range (num_to_start_from, num_to_end_at+1, interval)
l1 = list(range(10, 51, 5))
print (l1)
# Operations with list of numbers with -> min() max() sum()
l1 = [2, 3, 4, 45, 1, 5, 6, 3, 1, 23, 14]
print ("Sum: " + str(sum(l1)))
print ("Max: " + str(max(l1)))
print ("Min: " + str(min(l1)))
# List Comprehensions
# Simple
l1 = [i for i in range(20, 30, 1)]
l2 = [i+1 for i in range(20, 30, 1)]
l3 = [[i, i**2] for i in range(2, 12, 3)]
print (l1)
print (l2)
print (l3)
# A few more list comprehension examples
equi_list_1 = [[x, y, z] for x in range(1, 3) for y in range(3, 6) for z in range(6, 9)]
print (equi_list_1)
# The above list comprehension is equivalent of the following code
equi_list_2 = []
for x in range(1, 3):
for y in range(3, 6):
for z in range(6, 9):
equi_list_2.append([x, y, z])
print (equi_list_2)
# Proof of equivalence (Do execute the above two blocks of code before running this)
print (equi_list_1 == equi_list_2)
# List Comprehension with conditionals
l1 = [x if x%5==0 else "blank" for x in range(20)]
print (l1)
# One more list comprehension with conditionals
l1 = ["Jennifer", "met", "Scarlett", "in", "Python", "lessons", "they", "take."]
l2 = [[str(x) + ") " + y] for x in range(len(l1)) for y in l1 if l1[x] == y]
print (l2)
# Slicing a list
l1 = ["Jennifer", "is", "now", "friends", "with", "Scarlett"]
# [start_index : end_index+1]
print("[2:5] --> " + str(l1[2:5]))
print("[:4] --> " + str(l1[:4])) # everthing before 4th index [excluding the 4th]
print("[2:] --> " + str(l1[2:])) # everything from 2nd index [including the 2nd]
print("[:] --> " + str(l1[:])) # every element in the list
# Some more slicing
l1 = ["Jennifer", "and", "Scarlett", "now", "Pythonistas", "!"]
print ("[-2:] --> " + str(l1[-2:]))
print ("[:-3] --> " + str(l1[:-3]))
print ("[-5:-2] --> " + str(l1[-5:-2]))
print ("[-4:-6] --> " + str(l1[-4:-6]))
# Looping through a slice
l1 = ["Pythonistas", "rock", "!!!", "XD"]
for w in l1[-4:-1]:
print (w.upper())
# Copying a list
l1 = ["We", "should", "use", "[:]", "to", "copy", "the", "whole", "list"]
l2 = l1[:]
print(l2)
# Proof that the above two lists are different
l2.append(". Using [:] ensures the two lists are different")
print (l1)
print (l2)
# What happens if we directly assign one list to the other instead of using slices
l1 = ["Jennifer", "now", "wonders", "what", "happens", "if", "we", "directly", "assign."]
l2 = l1
l2.append("Both variables point to the same list")
print (l1)
print (l2)
```
| github_jupyter |
<h1>Table of Contents<span class="tocSkip"></span></h1>
<div class="toc" style="margin-top: 1em;"><ul class="toc-item"><li><span><a href="#Chart-object" data-toc-modified-id="Chart-object-1"><span class="toc-item-num">1 </span>Chart object</a></span></li><li><span><a href="#Adding-chart-labels" data-toc-modified-id="Adding-chart-labels-2"><span class="toc-item-num">2 </span>Adding chart labels</a></span></li><li><span><a href="#Getting-help" data-toc-modified-id="Getting-help-3"><span class="toc-item-num">3 </span>Getting help</a></span></li><li><span><a href="#Callouts" data-toc-modified-id="Callouts-4"><span class="toc-item-num">4 </span>Callouts</a></span></li><li><span><a href="#Axes" data-toc-modified-id="Axes-5"><span class="toc-item-num">5 </span>Axes</a></span></li><li><span><a href="#Method-chaining" data-toc-modified-id="Method-chaining-6"><span class="toc-item-num">6 </span>Method chaining</a></span></li><li><span><a href="#Plotting" data-toc-modified-id="Plotting-7"><span class="toc-item-num">7 </span>Plotting</a></span><ul class="toc-item"><li><span><a href="#Input-data-format" data-toc-modified-id="Input-data-format-7.1"><span class="toc-item-num">7.1 </span>Input data format</a></span></li><li><span><a href="#Pivoted-data:-INVALID" data-toc-modified-id="Pivoted-data:-INVALID-7.2"><span class="toc-item-num">7.2 </span>Pivoted data: INVALID</a></span><ul class="toc-item"><li><span><a href="#Melting-pivoted-data:-VALID" data-toc-modified-id="Melting-pivoted-data:-VALID-7.2.1"><span class="toc-item-num">7.2.1 </span>Melting pivoted data: VALID</a></span></li></ul></li><li><span><a href="#Pandas-series:-INVALID" data-toc-modified-id="Pandas-series:-INVALID-7.3"><span class="toc-item-num">7.3 </span>Pandas series: INVALID</a></span></li><li><span><a href="#Pandas-index:-INVALID" data-toc-modified-id="Pandas-index:-INVALID-7.4"><span class="toc-item-num">7.4 </span>Pandas index: INVALID</a></span></li><li><span><a href="#Pandas-DataFrame:-VALID" data-toc-modified-id="Pandas-DataFrame:-VALID-7.5"><span class="toc-item-num">7.5 </span>Pandas DataFrame: VALID</a></span></li></ul></li><li><span><a href="#Axis-types" data-toc-modified-id="Axis-types-8"><span class="toc-item-num">8 </span>Axis types</a></span></li><li><span><a href="#Vertical-Bar-plot" data-toc-modified-id="Vertical-Bar-plot-9"><span class="toc-item-num">9 </span>Vertical Bar plot</a></span></li><li><span><a href="#Examples" data-toc-modified-id="Examples-10"><span class="toc-item-num">10 </span>Examples</a></span></li><li><span><a href="#Bar-plot---Horizontal-vs.-Vertical" data-toc-modified-id="Bar-plot---Horizontal-vs.-Vertical-11"><span class="toc-item-num">11 </span>Bar plot - Horizontal vs. Vertical</a></span></li><li><span><a href="#Grouped-bar-plot" data-toc-modified-id="Grouped-bar-plot-12"><span class="toc-item-num">12 </span>Grouped bar plot</a></span></li><li><span><a href="#show('html')-vs.-show('png')" data-toc-modified-id="show('html')-vs.-show('png')-13"><span class="toc-item-num">13 </span>show('html') vs. show('png')</a></span></li><li><span><a href="#Color-palette-types" data-toc-modified-id="Color-palette-types-14"><span class="toc-item-num">14 </span>Color palette types</a></span></li><li><span><a href="#Color-palettes" data-toc-modified-id="Color-palettes-15"><span class="toc-item-num">15 </span>Color palettes</a></span></li><li><span><a href="#Layouts" data-toc-modified-id="Layouts-16"><span class="toc-item-num">16 </span>Layouts</a></span></li><li><span><a href="#Advanced-usage-with-Bokeh" data-toc-modified-id="Advanced-usage-with-Bokeh-17"><span class="toc-item-num">17 </span>Advanced usage with Bokeh</a></span></li></ul></div>
```
# Copyright (c) 2017-2018 Spotify AB
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
import chartify
import pandas as pd
def print_public_methods(obj):
print('Methods:')
print('\n'.join([x for x in dir(obj) if not x.startswith('_')]))
```
# Chart object
- Run the cell below to instantiate a chart and assign to to a variable
```
ch = chartify.Chart()
```
- Use .show() to render the chart.
```
ch.show()
```
- Note that the chart is blank at this point.
- The default labels provide directions for how to override their values.
# Adding chart labels
- __Your turn__: Add labels to the following chart. Look at the default values for instruction.
- Title
- Subtitle
- Source
- X label
- Y label
```
ch = chartify.Chart()
# Add code here to overwrite the labels
ch.show()
```
# Getting help
- From within a jupyter notebook you can see the available attributes of the chart object by pressing "tab"
- Select the space just after the "." character below and hit tab.
```
ch = chartify.Chart()
ch.
```
- You can also use "?" to pull up documentation for objects and methods.
- Run the cell below to pull up the chartify.Chart documentation
```
chartify.Chart?
```
- This can also be accomplished by pressing "shift + tab".
- Press "shift + tab" twice to see the expanded documentation.
- Try it with the next cell.
```
chartify.Chart
```
# Callouts
- The chart object has a callout object (ch.callout) that contains methods for adding callouts to the chart.
- Callouts can be used to add text, lines, or shaded areas to annotate parts of your chart.
- __Your Turn:__ Fill in the code below to add a text callout that says "hi" at coordinate (10, 10)
- Look up the documentation for ch.callout.text if you need help
```
ch = chartify.Chart()
ch.callout.text(# Fill in the code here)
ch.show()
```
- Use tab below to see what callouts are available.
```
ch.callout.
```
- You should see this list of callouts:
```
# List of available callouts:
print_public_methods(ch.callout)
```
# Axes
- The axes object contains methods for setting or getting axis properties.
```
# Avaiable axes methods:
print_public_methods(ch.axes)
```
- __Your turn__: modify the chart below so the xaxis range goes from 0 to 100
```
ch = chartify.Chart()
ch.callout.text('hi', 10, 10)
# Add code here to modify the xrange to (0, 100)
ch.show()
```
# Method chaining
- Chart methods can be chained by wrapping the statments in parentheses. See the example below:
```
(chartify.Chart(blank_labels=True)
.callout.text('hi', 10, 10)
.axes.set_xaxis_range(0, 100)
.show()
)
```
# Plotting
## Input data format
Chartify expects the input data to be:
- Tidy (Each variable has its own column, each row corresponds to an observation)
- In the columns of a Pandas DataFrame.
Below we'll explore some examples of valid and invalid input data
- Run this cell to generate an example dataset
```
data = chartify.examples.example_data()
data.head()
```
## Pivoted data: INVALID
- Pivoted data is not Tidy (note the `country` dimension has an observation in each column)
```
pivoted_data = pd.pivot_table(data, columns='country', values='quantity', index='fruit', aggfunc='sum')
pivoted_data
```
### Melting pivoted data: VALID
- You can use pandas.melt to convert pivoted data into the tidy data format.
- The output of SQL queries with `groupby` produces output in tidy format.
```
value_columns = pivoted_data.columns
melted_data = pd.melt(pivoted_data.reset_index(), # Need to reset the index to put "fruit" into a column.
id_vars='fruit',
value_vars=value_columns)
melted_data.head()
```
## Pandas series: INVALID
- Data in a pandas Series must be converted to a DataFrame for use with Chartify.
```
data.groupby(['country'])['quantity'].sum()
```
## Pandas index: INVALID
- The output below is a pandas DataFrame, but the country dimension is in the Index.
```
data.groupby(['country'])[['quantity']].sum()
```
## Pandas DataFrame: VALID
- The code below produces a valid pandas DataFrame for use with Chartify.
- Notice how the country dimension is now in a column.
```
chart_data = data.groupby(['country'])['quantity'].sum().reset_index()
chart_data
```
# Axis types
- Specify the x_axis_type and y_axis_type parameters when instantiating the chart object.
- Both are set to `linear` by default.
- Look at the chart object documentation to see the list of available options for x_axis_type and y_axis_type
```
chartify.Chart?
```
- __The Chart axis types influence the plots that are available__
- Look at how the plot methods change based on the axis types:
```
ch = chartify.Chart(x_axis_type='datetime',
y_axis_type='linear')
# List of available callouts:
print_public_methods(ch.plot)
ch = chartify.Chart(x_axis_type='categorical',
y_axis_type='linear')
# List of available plots:
print_public_methods(ch.plot)
```
- __Your turn__: Create a chart with 'density' y and 'linear' x axis types. What type of plots are available?
```
ch = chartify.Chart(# Your code goes here)
```
# Vertical Bar plot
- __Your turn__: Create a bar plot based on the dataframe below.
```
bar_data = (data.groupby('country')[['quantity']].sum()
.reset_index()
)
bar_data
# Implement the bar plot here.
# Set the appropriate x_axis_type otherwise the bar method won't be available.
# Look at the bar documentation to figure out how to pass in the parameters.
# If you get stuck move on to the next section for hints.
ch = chartify.Chart(# Your code goes here)
```
# Examples
- Chartify includes many examples. They're a good starting point if you're trying to create a chart that you're unfamiliar with.
```
# List of available examples
print_public_methods(chartify.examples)
```
- Run the appropriate method to see examples and the corresponding code that generates the example.
```
chartify.examples.plot_bar()
```
# Bar plot - Horizontal vs. Vertical
- Copy your bar plot here, but make it horizantal instead of vertical. Look to the example above if you get stuck.
# Grouped bar plot
- __Your Turn__: Create a grouped bar plot with the data below.
```
grouped_bar_data = (data.groupby(['country', 'fruit'])[['quantity']].sum()
.reset_index()
)
grouped_bar_data
# Implement the grouped bar plot here.
# Look at the example for help if you get stuck.
```
# show('html') vs. show('png')
- Chartify charts can be rendered as either "HTML" or "PNG" (HTML is the default)
- HTML output:
- Is faster and good for iteration.
- Can be saved as an image by screenshotting, or by clicking the icon on the top right.
- Will _NOT_ show up in Jupyter notebooks when uploaded to GHE.
- PNG output:
- Is slower and better for the finished product.
- Can be copy and pasted directly from the jupyter notebook (Right click on the image)
- Will show up in Jupyter notebooks when uploaded to GHE.
- See the difference by running the two cells below
```
(chartify.Chart(blank_labels=True)
.set_title("HTML output")
.set_subtitle("Faster, but will not show up in GHE")
.show()
)
(chartify.Chart(blank_labels=True)
.set_title("PNG output")
.set_subtitle("Slower, but will show up in GHE. Right click to copy + paste.")
.show('png')
)
```
# Color palette types
- Chartify includes 4 different color palette types: `categorical`, `accent`, `sequential`, `diverging`.
- Note the differences in the examples below
```
chartify.examples.style_color_palette_categorical()
chartify.examples.style_color_palette_accent()
chartify.examples.style_color_palette_diverging()
chartify.examples.style_color_palette_sequential()
```
# Color palettes
- Chartify includes a set of pre-defined color palettes:
```
chartify.color_palettes
```
- Use .show() to see the colors associated with each:
```
chartify.color_palettes.show()
```
- Assign the color palettes with `.set_color_palette`
```
ch = chartify.Chart(x_axis_type='categorical',
blank_labels=True)
ch.style.set_color_palette('categorical', 'Dark2')
ch.plot.bar(data_frame=grouped_bar_data,
categorical_columns=['fruit', 'country'],
numeric_column='quantity',
color_column='fruit')
ch.show()
```
- Color palette objects include methods for manipulation. See the examples below:
```
dark2 = chartify.color_palettes['Dark2']
dark2.show()
```
- Sort
```
sorted_dark2 = dark2.sort_by_hue()
sorted_dark2.show()
```
- Expand
```
dark2.expand_palette(20).show()
```
- Shift
```
shifted_dark2 = dark2.shift_palette('white', percent=20)
shifted_dark2.show()
- Assign the shifted color palette to a chart:
ch = chartify.Chart(x_axis_type='categorical',
blank_labels=True)
ch.style.set_color_palette('categorical', shifted_dark2)
ch.plot.bar(data_frame=grouped_bar_data,
categorical_columns=['fruit', 'country'],
numeric_column='quantity',
color_column='fruit')
ch.show()
```
# Layouts
- Chartify layouts are tailored toward use in slides.
- Notice how the output changes for each of the slide layout options below:
```
layout_options = ['slide_100%', 'slide_75%', 'slide_50%', 'slide_25%']
for option in layout_options:
ch = chartify.Chart(layout=option, blank_labels=True, x_axis_type='categorical')
ch.set_title('Layout: {}'.format(option))
ch.plot.bar(data_frame=grouped_bar_data,
categorical_columns=['fruit', 'country'],
numeric_column='quantity',
color_column='fruit')
ch.show()
```
# Advanced usage with Bokeh
- Chartify is built on top of another visualization package called [Bokeh](http://bokeh.pydata.org/en/latest/)
- The example below shows how you can access the Bokeh [figure](https://bokeh.pydata.org/en/latest/docs/reference/plotting.html#bokeh.plotting.figure.Figure) from a Chartify chart object.
```
ch = chartify.Chart(blank_labels=True, x_axis_type='categorical')
ch.plot.bar(data_frame=grouped_bar_data,
categorical_columns=['fruit', 'country'],
numeric_column='quantity',
color_column='fruit')
ch.figure
```
- The following example shows how you can modify attributes not exposed in Chartify by accessing the Bokeh figure. See [Bokeh](http://bokeh.pydata.org/en/latest/) documentation for more details.
```
ch.figure.xaxis.axis_label_text_font_size = '30pt'
ch.figure.xaxis.axis_label_text_color = 'red'
ch.figure.height = 400
ch.axes.set_xaxis_label('A large xaxis label')
ch.show()
```
| github_jupyter |
# Welcome to 101 Exercises for Python Fundamentals
Solving these exercises will help make you a better programmer. Solve them in order, because each solution builds scaffolding, working code, and knowledge you can use on future problems. Read the directions carefully, and have fun!
> "Learning to program takes a little bit of study and a *lot* of practice" - Luis Montealegre
## Getting Started
1. Go to https://colab.research.google.com/github/ryanorsinger/101-exercises/blob/main/101-exercises.ipynb
2. To save your work to your Google Drive, go to File then "Save Copy in Drive".
3. Your own work will now appear in your Google Drive account!
If you need a fresh, blank copy of this document, go to https://colab.research.google.com/github/ryanorsinger/101-exercises/blob/main/101-exercises.ipynb and save a fresh copy in your Google Drive.
## Orientation
- This code notebook is composed of cells. Each cell is either text or Python code.
- To run a cell of code, click the "play button" icon to the left of the cell or click on the cell and press "Shift+Enter" on your keyboard. This will execute the Python code contained in the cell. Executing a cell that defines a variable is important before executing or authoring a cell that depends on that previously created variable assignment.
- Each *assert* line is both an example and a test that tests for the presence and functionality of the instructed exercise.
## Outline
- Each cell starts with a problem statement that describes the exercise to complete.
- Underneath each problem statement, learners will need to write code to produce an answer.
- The **assert** lines test to see that your code solves the problem appropriately.
- Do not alter or delete the assertion tests, since those are providing automated testing.
- Many exercises will rely on previous solutions to be correctly completed
- The `print("Exercise is complete")` line will only run if your solution passes the assertion test(s)
- Be sure to create programmatic solutions that will work for all inputs:
- For example, calling the `is_even(2)` returns `True`, but your function should work for all even numbers, both positive and negative.
## Guidance
- Get Python to do the work for you. For example, if the exercise instructs you to reverse a list of numbers, your job is to find the
- Save often by clicking the blue "Save" button.
- If you need to clear the output or reset the notebook, go to "Run" then "Restart Session" to clear up any error messages.
- Do not move or alter the lines of code that contain the `assert` statements. Those are what run your solution and test its actual output vs. expected outputs.
- Seek to understand the problem before trying to solve it. Can you explain the problem to someone else in English? Can you explain the solution in English?
- Slow down and read any error messages you encounter. Error messages provide insight into how to resolve the error. When in doubt, put your exact error into a search engine and look for results that reference an identical or similar problem.
## Get Python To Do The Work For You
One of the main jobs of a programming language is to help people solve problems programatically, so we don't have to do so much by hand. For example, it's easy for a person to manually reverse the list `[1, 2, 3]`, but imagine reversing a list of a million things or sorting a list of even a hundred things. When we write programmatic solutions in code, we are providing instructions to the computer to do a task. Computers follow the letter of the code, not the intent, and do exactly what they are told to do. In this way, Python can reverse a list of 3 numbers or 100 numbers or ten million numbers with the same instructions. Repetition is a key idea behind programming languages.
This means that your task with these exercises is to determine a sequence of steps that solve the problem and then find the Python code that will run those instructions. If you're sorting or reversing things by hand, you're not doing it right!
## How To Discover How To Do Something in Python
1. The first step is to make sure you know what the problem is asking.
2. The second step is to determine, in English (or your first spoken language), what steps you need to take.
3. Use a search engine to look for code examples to identical or similar problems.
One of the best ways to discover how to do things in Python is to use a search engine. Go to your favorite search engine and search for "how to reverse a list in Python" or "how to sort a list in Python". That's how both learners and professionals find answers and examples all the time. Search for what you want and add "in Python" and you'll get lots of code examples. Searching for "How to sum a list of numbers in Python" is a very effective way to discover exactly how to do that task.
### Learning to Program and Code
- You can make a new blank cell for Python code at any time in this document.
- If you want more freedom to explore learning Python in a blank notebook, go here https://colab.research.google.com/#create=true and make yourself a blank, new notebook.
- Programming is an intellectual activity of designing a solution. "Coding" means turning your programmatic solution into code w/ all the right syntax and parts of the programming language.
- Expect to make mistakes and adopt the attitude that **the error message provides the information you need to proceed**. You will put lots of error messages into search engines to learn this craft!
- Because computers have zero ability to read in between the lines or "catch the drift" or know what you mean, code only does what it is told to do.
- Code doesn't do what you *want* it to do, code does what you've told it to do.
- Before writing any code, figure out how you would solve the problem in spoken language to describe the sequence of steps in the solution.
- Think about your solution in English (or your natural language). It's **critical** to solve the problem in your natural language before trying to get a programming language to do the work.
## Troubleshooting
- If this entire document shows "Name Error" for many cells, it means you should read the "Getting Started" instructions above to make your own copy.
- Be sure to commit your work to make save points, as you go.
- If you load this page and you see your code but not the results of the code, be sure to run each cell (shift + Enter makes this quick)
- "Name Error" means that you need to assign a variable or define the function as instructed.
- "Assertion Error" means that your provided solution does not match the correct answer.
- "Type Error" means that your data type provided is not accurate
- If your kernel freezes, click on "Run" then select "Restart Session"
- If you require additional troubleshooting assistance, click on "Help" and then "Docs" to access documentation for this platform.
- If you have discoverd a bug or typo, please triple check your spelling then create a new issue at [https://github.com/ryanorsinger/101-exercises/issues](https://github.com/ryanorsinger/101-exercises/issues) to notify the author.
## What to do when you don't know what to do next
- When the exercise asks you to reverse an list, the way forward is to search for "How to reverse a list in Python" in your favorite search engine.
- When the exercise asks you to check if a number is even, the way forward is to search for "how to check if a number is even in Python".
- When the exercise has you calculate the area of a circle, the way forward is to search for "how to calculate the area of a circle in Python" or "How to get pi in Python".
> The pattern for finding what you need in JavaScript is to rely very heavily on search engine searches so you can find examples of working code and discussions about code that speak to your questions.
```
# Example problem:
# Uncomment the line below and run this cell.
# The hashtag "#" character in a line of Python code is the comment character.
# doing_python_right_now = True
# The lines below will test your answer. If you see an error, then it means that your answer is incorrect or incomplete.
assert doing_python_right_now == True, "If you see a NameError, it means that the variable is not created and assigned a value. An 'Assertion Error' means that the value of the variable is incorrect."
print("Exercise 0 is correct") # This line will print if your solution passes the assertion above.
# Exercise 1
# On the line below, create a variable named on_mars_right_now and assign it the boolean value of False
assert on_mars_right_now == False, "If you see a Name Error, be sure to create the variable and assign it a value."
print("Exercise 1 is correct.")
# Exercise 2
# Create a variable named fruits and assign it a list of fruits containing the following fruit names as strings:
# mango, banana, guava, kiwi, and strawberry.
assert fruits == ["mango", "banana", "guava", "kiwi", "strawberry"], "If you see an Assert Error, ensure the variable contains all the strings in the provided order"
print("Exercise 2 is correct.")
# Exercise 3
# Create a variable named vegetables and assign it a list of fruits containing the following vegetable names as strings:
# eggplant, broccoli, carrot, cauliflower, and zucchini
assert vegetables == ["eggplant", "broccoli", "carrot", "cauliflower", "zucchini"], "Ensure the variable contains all the strings in the provided order"
print("Exercise 3 is correct.")
# Exercise 4
# Create a variable named numbers and assign it a list of numbers, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10
assert numbers == [1, 2, 3, 4, 5, 6, 7, 8, 9, 10], "Ensure the variable contains the numbers 1-10 in order."
print("Exercise 4 is correct.")
```
## List Operations
**Hint** Recommend finding and using built-in Python functionality whenever possible.
```
# Exercise 5
# Given the following assigment of the list of fruits, add "tomato" to the end of the list.
fruits = ["mango", "banana", "guava", "kiwi", "strawberry"]
assert fruits == ["mango", "banana", "guava", "kiwi", "strawberry", "tomato"], "Ensure the variable contains all the strings in the right order"
print("Exercise 5 is correct")
# Exercise 6
# Given the following assignment of the vegetables list, add "tomato" to the end of the list.
vegetables = ["eggplant", "broccoli", "carrot", "cauliflower", "zucchini"]
assert vegetables == ["eggplant", "broccoli", "carrot", "cauliflower", "zucchini", "tomato"], "Ensure the variable contains all the strings in the provided order"
print("Exercise 6 is correct")
# Exercise 7
# Given the list of numbers defined below, reverse the list of numbers that you created above.
numbers = [1, 2, 3, 4, 5, 6, 7, 8, 9, 10]
assert numbers == [10, 9, 8, 7, 6, 5, 4, 3, 2, 1], "Assert Error means that the answer is incorrect."
print("Exercise 7 is correct.")
# Exercise 8
# Sort the vegetables in alphabetical order
assert vegetables == ['broccoli', 'carrot', 'cauliflower', 'eggplant', 'tomato', 'zucchini']
print("Exercise 8 is correct.")
# Exercise 9
# Write the code necessary to sort the fruits in reverse alphabetical order
assert fruits == ['tomato', 'strawberry', 'mango', 'kiwi', 'guava', 'banana']
print("Exercise 9 is correct.")
# Exercise 10
# Write the code necessary to produce a single list that holds all fruits then all vegetables in the order as they were sorted above.
assert fruits_and_veggies == ['tomato', 'strawberry', 'mango', 'kiwi', 'guava', 'banana', 'broccoli', 'carrot', 'cauliflower', 'eggplant', 'tomato', 'zucchini']
print("Exercise 10 is correct")
```
## Basic Functions
**Hint** Be sure to `return` values from your function definitions. The assert statements will call your function(s) for you.
```
# Run this cell in order to generate some numbers to use in our functions after this.
import random
positive_even_number = random.randrange(2, 101, 2)
negative_even_number = random.randrange(-100, -1, 2)
positive_odd_number = random.randrange(1, 100, 2)
negative_odd_number = random.randrange(-101, 0, 2)
print("We now have some random numbers available for future exercises.")
print("The random positive even number is", positive_even_number)
print("The random positive odd nubmer is", positive_odd_number)
print("The random negative even number", negative_even_number)
print("The random negative odd number", negative_odd_number)
# Example function defintion:
# Write a say_hello function that adds the string "Hello, " to the beginning and "!" to the end of any given input.
def say_hello(name):
return "Hello, " + name + "!"
assert say_hello("Jane") == "Hello, Jane!", "Double check the inputs and data types"
assert say_hello("Pat") == "Hello, Pat!", "Double check the inputs and data types"
assert say_hello("Astrud") == "Hello, Astrud!", "Double check the inputs and data types"
print("The example function definition ran appropriately")
# Another example function definition:
# This plus_two function takes in a variable and adds 2 to it.
def plus_two(number):
return number + 2
assert plus_two(3) == 5
assert plus_two(0) == 2
assert plus_two(-2) == 0
print("The plus_two assertions executed appropriately... The second function definition example executed appropriately.")
# Exercise 11
# Write a function definition for a function named add_one that takes in a number and returns that number plus one.
assert add_one(2) == 3, "Ensure that the function is defined, named properly, and returns the correct value"
assert add_one(0) == 1, "Zero plus one is one."
assert add_one(positive_even_number) == positive_even_number + 1, "Ensure that the function is defined, named properly, and returns the correct value"
assert add_one(negative_odd_number) == negative_odd_number + 1, "Ensure that the function is defined, named properly, and returns the correct value"
print("Exercise 11 is correct.")
# Exercise 12
# Write a function definition named is_positive that takes in a number and returns True or False if that number is positive.
assert is_positive(positive_odd_number) == True, "Ensure that the function is defined, named properly, and returns the correct value"
assert is_positive(positive_even_number) == True, "Ensure that the function is defined, named properly, and returns the correct value"
assert is_positive(negative_odd_number) == False, "Ensure that the function is defined, named properly, and returns the correct value"
assert is_positive(negative_even_number) == False, "Ensure that the function is defined, named properly, and returns the correct value"
assert is_positive(0) == False, "Zero is not a positive number."
print("Exercise 12 is correct.")
# Exercise 13
# Write a function definition named is_negative that takes in a number and returns True or False if that number is negative.
assert is_negative(positive_odd_number) == False, "Ensure that the function is defined, named properly, and returns the correct value"
assert is_negative(positive_even_number) == False, "Ensure that the function is defined, named properly, and returns the correct value"
assert is_negative(negative_odd_number) == True, "Ensure that the function is defined, named properly, and returns the correct value"
assert is_negative(negative_even_number) == True, "Ensure that the function is defined, named properly, and returns the correct value"
assert is_negative(0) == False, "Zero is not a negative number."
print("Exercise 13 is correct.")
# Exercise 14
# Write a function definition named is_odd that takes in a number and returns True or False if that number is odd.
assert is_odd(positive_odd_number) == True, "Ensure that the function is defined, named properly, and returns the correct value"
assert is_odd(positive_even_number) == False, "Ensure that the function is defined, named properly, and returns the correct value"
assert is_odd(negative_odd_number) == True, "Ensure that the function is defined, named properly, and returns the correct value"
assert is_odd(negative_even_number) == False, "Ensure that the function is defined, named properly, and returns the correct value"
print("Exercise 14 is correct.")
# Exercise 15
# Write a function definition named is_even that takes in a number and returns True or False if that number is even.
assert is_even(2) == True, "Ensure that the function is defined, named properly, and returns the correct value"
assert is_even(positive_odd_number) == False, "Ensure that the function is defined, named properly, and returns the correct value"
assert is_even(positive_even_number) == True, "Ensure that the function is defined, named properly, and returns the correct value"
assert is_even(negative_odd_number) == False, "Ensure that the function is defined, named properly, and returns the correct value"
assert is_even(negative_even_number) == True, "Ensure that the function is defined, named properly, and returns the correct value"
print("Exercise 15 is correct.")
# Exercise 16
# Write a function definition named identity that takes in any argument and returns that argument's value. Don't overthink this one!
assert identity(fruits) == fruits, "Ensure that the function is defined, named properly, and returns the correct value"
assert identity(vegetables) == vegetables, "Ensure that the function is defined, named properly, and returns the correct value"
assert identity(positive_odd_number) == positive_odd_number, "Ensure that the function is defined, named properly, and returns the correct value"
assert identity(positive_even_number) == positive_even_number, "Ensure that the function is defined, named properly, and returns the correct value"
assert identity(negative_odd_number) == negative_odd_number, "Ensure that the function is defined, named properly, and returns the correct value"
assert identity(negative_even_number) == negative_even_number, "Ensure that the function is defined, named properly, and returns the correct value"
print("Exercise 16 is correct.")
# Exercise 17
# Write a function definition named is_positive_odd that takes in a number and returns True or False if the value is both greater than zero and odd
assert is_positive_odd(3) == True, "Double check your syntax and logic"
assert is_positive_odd(positive_odd_number) == True, "Double check your syntax and logic"
assert is_positive_odd(positive_even_number) == False, "Double check your syntax and logic"
assert is_positive_odd(negative_odd_number) == False, "Double check your syntax and logic"
assert is_positive_odd(negative_even_number) == False, "Double check your syntax and logic"
print("Exercise 17 is correct.")
# Exercise 18
# Write a function definition named is_positive_even that takes in a number and returns True or False if the value is both greater than zero and even
assert is_positive_even(4) == True, "Double check your syntax and logic"
assert is_positive_even(positive_odd_number) == False, "Double check your syntax and logic"
assert is_positive_even(positive_even_number) == True, "Double check your syntax and logic"
assert is_positive_even(negative_odd_number) == False, "Double check your syntax and logic"
assert is_positive_even(negative_even_number) == False, "Double check your syntax and logic"
print("Exercise 18 is correct.")
# Exercise 19
# Write a function definition named is_negative_odd that takes in a number and returns True or False if the value is both less than zero and odd.
assert is_negative_odd(-3) == True, "Double check your syntax and logic"
assert is_negative_odd(positive_odd_number) == False, "Double check your syntax and logic"
assert is_negative_odd(positive_even_number) == False, "Double check your syntax and logic"
assert is_negative_odd(negative_odd_number) == True, "Double check your syntax and logic"
assert is_negative_odd(negative_even_number) == False, "Double check your syntax and logic"
print("Exercise 19 is correct.")
# Exercise 20
# Write a function definition named is_negative_even that takes in a number and returns True or False if the value is both less than zero and even.
assert is_negative_even(-4) == True, "Double check your syntax and logic"
assert is_negative_even(positive_odd_number) == False, "Double check your syntax and logic"
assert is_negative_even(positive_even_number) == False, "Double check your syntax and logic"
assert is_negative_even(negative_odd_number) == False, "Double check your syntax and logic"
assert is_negative_even(negative_even_number) == True, "Double check your syntax and logic"
print("Exercise 20 is correct.")
# Exercise 21
# Write a function definition named half that takes in a number and returns half the provided number.
assert half(4) == 2
assert half(5) == 2.5
assert half(positive_odd_number) == positive_odd_number / 2
assert half(positive_even_number) == positive_even_number / 2
assert half(negative_odd_number) == negative_odd_number / 2
assert half(negative_even_number) == negative_even_number / 2
print("Exercise 21 is correct.")
# Exercise 22
# Write a function definition named double that takes in a number and returns double the provided number.
assert double(4) == 8
assert double(5) == 10
assert double(positive_odd_number) == positive_odd_number * 2
assert double(positive_even_number) == positive_even_number * 2
assert double(negative_odd_number) == negative_odd_number * 2
assert double(negative_even_number) == negative_even_number * 2
print("Exercise 22 is correct.")
# Exercise 23
# Write a function definition named triple that takes in a number and returns triple the provided number.
assert triple(4) == 12
assert triple(5) == 15
assert triple(positive_odd_number) == positive_odd_number * 3
assert triple(positive_even_number) == positive_even_number * 3
assert triple(negative_odd_number) == negative_odd_number * 3
assert triple(negative_even_number) == negative_even_number * 3
print("Exercise 23 is correct.")
# Exercise 24
# Write a function definition named reverse_sign that takes in a number and returns the provided number but with the sign reversed.
assert reverse_sign(4) == -4
assert reverse_sign(-5) == 5
assert reverse_sign(positive_odd_number) == positive_odd_number * -1
assert reverse_sign(positive_even_number) == positive_even_number * -1
assert reverse_sign(negative_odd_number) == negative_odd_number * -1
assert reverse_sign(negative_even_number) == negative_even_number * -1
print("Exercise 24 is correct.")
# Exercise 25
# Write a function definition named absolute_value that takes in a number and returns the absolute value of the provided number
assert absolute_value(4) == 4
assert absolute_value(-5) == 5
assert absolute_value(positive_odd_number) == positive_odd_number
assert absolute_value(positive_even_number) == positive_even_number
assert absolute_value(negative_odd_number) == negative_odd_number * -1
assert absolute_value(negative_even_number) == negative_even_number * -1
print("Exercise 25 is correct.")
# Exercise 26
# Write a function definition named is_multiple_of_three that takes in a number and returns True or False if the number is evenly divisible by 3.
assert is_multiple_of_three(3) == True
assert is_multiple_of_three(15) == True
assert is_multiple_of_three(9) == True
assert is_multiple_of_three(4) == False
assert is_multiple_of_three(10) == False
print("Exercise 26 is correct.")
# Exercise 27
# Write a function definition named is_multiple_of_five that takes in a number and returns True or False if the number is evenly divisible by 5.
assert is_multiple_of_five(3) == False
assert is_multiple_of_five(15) == True
assert is_multiple_of_five(9) == False
assert is_multiple_of_five(4) == False
assert is_multiple_of_five(10) == True
print("Exercise 27 is correct.")
# Exercise 28
# Write a function definition named is_multiple_of_both_three_and_five that takes in a number and returns True or False if the number is evenly divisible by both 3 and 5.
assert is_multiple_of_both_three_and_five(15) == True
assert is_multiple_of_both_three_and_five(45) == True
assert is_multiple_of_both_three_and_five(3) == False
assert is_multiple_of_both_three_and_five(9) == False
assert is_multiple_of_both_three_and_five(4) == False
print("Exercise 28 is correct.")
# Exercise 29
# Write a function definition named square that takes in a number and returns the number times itself.
assert square(3) == 9
assert square(2) == 4
assert square(9) == 81
assert square(positive_odd_number) == positive_odd_number * positive_odd_number
print("Exercise 29 is correct.")
# Exercise 30
# Write a function definition named add that takes in two numbers and returns the sum.
assert add(3, 2) == 5
assert add(10, -2) == 8
assert add(5, 7) == 12
print("Exercise 30 is correct.")
# Exercise 31
# Write a function definition named cube that takes in a number and returns the number times itself, times itself.
assert cube(3) == 27
assert cube(2) == 8
assert cube(5) == 125
assert cube(positive_odd_number) == positive_odd_number * positive_odd_number * positive_odd_number
print("Exercise 31 is correct.")
# Exercise 32
# Write a function definition named square_root that takes in a number and returns the square root of the provided number
assert square_root(4) == 2.0
assert square_root(64) == 8.0
assert square_root(81) == 9.0
print("Exercise 32 is correct.")
# Exercise 33
# Write a function definition named subtract that takes in two numbers and returns the first minus the second argument.
assert subtract(8, 6) == 2
assert subtract(27, 4) == 23
assert subtract(12, 2) == 10
print("Exercise 33 is correct.")
# Exercise 34
# Write a function definition named multiply that takes in two numbers and returns the first times the second argument.
assert multiply(2, 1) == 2
assert multiply(3, 5) == 15
assert multiply(5, 2) == 10
print("Exercise 34 is correct.")
# Exercise 35
# Write a function definition named divide that takes in two numbers and returns the first argument divided by the second argument.
assert divide(27, 9) == 3
assert divide(15, 3) == 5
assert divide(5, 2) == 2.5
assert divide(10, 2) == 5
print("Exercise 35 is correct.")
# Exercise 36
# Write a function definition named quotient that takes in two numbers and returns only the quotient from dividing the first argument by the second argument.
assert quotient(27, 9) == 3
assert quotient(5, 2) == 2
assert quotient(10, 3) == 3
print("Exercise 36 is correct.")
# Exercise 37
# Write a function definition named remainder that takes in two numbers and returns the remainder of first argument divided by the second argument.
assert remainder(3, 3) == 0
assert remainder(5, 2) == 1
assert remainder(7, 5) == 2
print("Exercise 37 is correct.")
# Exercise 38
# Write a function definition named sum_of_squares that takes in two numbers, squares each number, then returns the sum of both squares.
assert sum_of_squares(3, 2) == 13
assert sum_of_squares(5, 2) == 29
assert sum_of_squares(2, 4) == 20
print("Exercise 38 is correct.")
# Exercise 39
# Write a function definition named times_two_plus_three that takes in a number, multiplies it by two, adds 3 and returns the result.
assert times_two_plus_three(0) == 3
assert times_two_plus_three(1) == 5
assert times_two_plus_three(2) == 7
assert times_two_plus_three(3) == 9
assert times_two_plus_three(5) == 13
print("Exercise 39 is correct.")
# Exercise 40
# Write a function definition named area_of_rectangle that takes in two numbers and returns the product.
assert area_of_rectangle(1, 3) == 3
assert area_of_rectangle(5, 2) == 10
assert area_of_rectangle(2, 7) == 14
assert area_of_rectangle(5.3, 10.3) == 54.59
print("Exercise 40 is correct.")
import math
# Exercise 41
# Write a function definition named area_of_circle that takes in a number representing a circle's radius and returns the area of the circl
assert area_of_circle(3) == 28.274333882308138
assert area_of_circle(5) == 78.53981633974483
assert area_of_circle(7) == 153.93804002589985
print("Exercise 41 is correct.")
import math
# Exercise 42
# Write a function definition named circumference that takes in a number representing a circle's radius and returns the circumference.
assert circumference(3) == 18.84955592153876
assert circumference(5) == 31.41592653589793
assert circumference(7) == 43.982297150257104
print("Exercise 42 is correct.")
```
## Functions working with strings
If you need some guidance working with the next few problems, recommend reading through [this example code](https://gist.github.com/ryanorsinger/f758599c886549e7615ec43488ae514c)
```
# Exercise 43
# Write a function definition named is_vowel that takes in value and returns True if the value is a, e, i, o, u in upper or lower case.
assert is_vowel("a") == True
assert is_vowel("e") == True
assert is_vowel("i") == True
assert is_vowel("o") == True
assert is_vowel("u") == True
assert is_vowel("A") == True
assert is_vowel("E") == True
assert is_vowel("I") == True
assert is_vowel("O") == True
assert is_vowel("U") == True
assert is_vowel("banana") == False
assert is_vowel("Q") == False
assert is_vowel("y") == False
assert is_vowel("aei") == False
assert is_vowel("iou") == False
print("Exercise 43 is correct.")
# Exercise 44
# Write a function definition named has_vowels that takes in value and returns True if the string contains any vowels.
assert has_vowels("banana") == True
assert has_vowels("ubuntu") == True
assert has_vowels("QQQQ") == False
assert has_vowels("wyrd") == False
print("Exercise 44 is correct.")
# Exercise 45
# Write a function definition named count_vowels that takes in value and returns the count of the nubmer of vowels in a sequence.
assert count_vowels("banana") == 3
assert count_vowels("ubuntu") == 3
assert count_vowels("mango") == 2
assert count_vowels("QQQQ") == 0
assert count_vowels("wyrd") == 0
print("Exercise 45 is correct.")
# Exercise 46
# Write a function definition named remove_vowels that takes in string and returns the string without any vowels
assert remove_vowels("banana") == "bnn"
assert remove_vowels("ubuntu") == "bnt"
assert remove_vowels("mango") == "mng"
assert remove_vowels("QQQQ") == "QQQQ"
print("Exercise 46 is correct.")
# Exercise 47
# Write a function definition named starts_with_vowel that takes in string and True if the string starts with a vowel
assert starts_with_vowel("ubuntu") == True
assert starts_with_vowel("banana") == False
assert starts_with_vowel("mango") == False
print("Exercise 47 is correct.")
# Exercise 48
# Write a function definition named ends_with_vowel that takes in string and True if the string ends with a vowel
assert ends_with_vowel("ubuntu") == True
assert ends_with_vowel("banana") == True
assert ends_with_vowel("mango") == True
assert ends_with_vowel("spinach") == False
print("Exercise 48 is correct.")
# Exercise 49
# Write a function definition named starts_and_ends_with_vowel that takes in string and returns True if the string starts and ends with a vowel
assert starts_and_ends_with_vowel("ubuntu") == True
assert starts_and_ends_with_vowel("banana") == False
assert starts_and_ends_with_vowel("mango") == False
print("Exercise 49 is correct.")
```
## Accessing List Elements
```
# Exercise 50
# Write a function definition named first that takes in sequence and returns the first value of that sequence.
assert first("ubuntu") == "u"
assert first([1, 2, 3]) == 1
assert first(["python", "is", "awesome"]) == "python"
print("Exercise 50 is correct.")
# Exercise 51
# Write a function definition named second that takes in sequence and returns the second value of that sequence.
assert second("ubuntu") == "b"
assert second([1, 2, 3]) == 2
assert second(["python", "is", "awesome"]) == "is"
print("Exercise 51 is correct.")
# Exercise 52
# Write a function definition named third that takes in sequence and returns the third value of that sequence.
assert third("ubuntu") == "u"
assert third([1, 2, 3]) == 3
assert third(["python", "is", "awesome"]) == "awesome"
print("Exercise 52 is correct.")
# Exercise 53
# Write a function definition named forth that takes in sequence and returns the forth value of that sequence.
assert forth("ubuntu") == "n"
assert forth([1, 2, 3, 4]) == 4
assert forth(["python", "is", "awesome", "right?"]) == "right?"
print("Exercise 53 is correct.")
# Exercise 54
# Write a function definition named last that takes in sequence and returns the last value of that sequence.
assert last("ubuntu") == "u"
assert last([1, 2, 3, 4]) == 4
assert last(["python", "is", "awesome"]) == "awesome"
assert last(["kiwi", "mango", "guava"]) == "guava"
print("Exercise 54 is correct.")
# Exercise 55
# Write a function definition named second_to_last that takes in sequence and returns the second to last value of that sequence.
assert second_to_last("ubuntu") == "t"
assert second_to_last([1, 2, 3, 4]) == 3
assert second_to_last(["python", "is", "awesome"]) == "is"
assert second_to_last(["kiwi", "mango", "guava"]) == "mango"
print("Exercise 55 is correct.")
# Exercise 56
# Write a function definition named third_to_last that takes in sequence and returns the third to last value of that sequence.
assert third_to_last("ubuntu") == "n"
assert third_to_last([1, 2, 3, 4]) == 2
assert third_to_last(["python", "is", "awesome"]) == "python"
assert third_to_last(["strawberry", "kiwi", "mango", "guava"]) == "kiwi"
print("Exercise 56 is correct.")
# Exercise 57
# Write a function definition named first_and_second that takes in sequence and returns the first and second value of that sequence as a list
assert first_and_second([1, 2, 3, 4]) == [1, 2]
assert first_and_second(["python", "is", "awesome"]) == ["python", "is"]
assert first_and_second(["strawberry", "kiwi", "mango", "guava"]) == ["strawberry", "kiwi"]
print("Exercise 57 is correct.")
# Exercise 58
# Write a function definition named first_and_last that takes in sequence and returns the first and last value of that sequence as a list
assert first_and_last([1, 2, 3, 4]) == [1, 4]
assert first_and_last(["python", "is", "awesome"]) == ["python", "awesome"]
assert first_and_last(["strawberry", "kiwi", "mango", "guava"]) == ["strawberry", "guava"]
print("Exercise 58 is correct.")
# Exercise 59
# Write a function definition named first_to_last that takes in sequence and returns the sequence with the first value moved to the end of the sequence.
assert first_to_last([1, 2, 3, 4]) == [2, 3, 4, 1]
assert first_to_last(["python", "is", "awesome"]) == ["is", "awesome", "python"]
assert first_to_last(["strawberry", "kiwi", "mango", "guava"]) == ["kiwi", "mango", "guava", "strawberry"]
print("Exercise 59 is correct.")
```
## Functions to describe data
```
# Exercise 60
# Write a function definition named sum_all that takes in sequence of numbers and returns all the numbers added together.
assert sum_all([1, 2, 3, 4]) == 10
assert sum_all([3, 3, 3]) == 9
assert sum_all([0, 5, 6]) == 11
print("Exercise 60 is correct.")
# Exercise 61
# Write a function definition named mean that takes in sequence of numbers and returns the average value
assert mean([1, 2, 3, 4]) == 2.5
assert mean([3, 3, 3]) == 3
assert mean([1, 5, 6]) == 4
print("Exercise 61 is correct.")
# Exercise 62
# Write a function definition named median that takes in sequence of numbers and returns the average value
assert median([1, 2, 3, 4, 5]) == 3.0
assert median([1, 2, 3]) == 2.0
assert median([1, 5, 6]) == 5.0
assert median([1, 2, 5, 6]) == 3.5
print("Exercise 62 is correct.")
# Exercise 63
# Write a function definition named mode that takes in sequence of numbers and returns the most commonly occuring value
assert mode([1, 2, 2, 3, 4]) == 2
assert mode([1, 1, 2, 3]) == 1
assert mode([2, 2, 3, 3, 3]) == 3
print("Exercise 63 is correct.")
# Exercise 64
# Write a function definition named product_of_all that takes in sequence of numbers and returns the product of multiplying all the numbers together
assert product_of_all([1, 2, 3]) == 6
assert product_of_all([3, 4, 5]) == 60
assert product_of_all([2, 2, 3, 0]) == 0
print("Exercise 64 is correct.")
```
## Applying functions to lists
```
# Exercise 65
# Write a function definition named get_highest_number that takes in sequence of numbers and returns the largest number.
assert get_highest_number([1, 2, 3]) == 3
assert get_highest_number([-5, -4, -3, -2, -1, 1, 2, 3, 4, 5]) == 5
assert get_highest_number([-5, -3, 1]) == 1
print("Exercise 65 is correct.")
# Exercise 66
# Write a function definition named get_smallest_number that takes in sequence of numbers and returns the smallest number.
assert get_smallest_number([1, 3, 2]) == 1
assert get_smallest_number([5, -5, -4, -3, -2, -1, 1, 2, 3, 4]) == -5
assert get_smallest_number([-4, -3, 1, -10]) == -10
print("Exercise 66 is correct.")
# Exercise 67
# Write a function definition named only_odd_numbers that takes in sequence of numbers and returns the odd numbers in a list.
assert only_odd_numbers([1, 2, 3]) == [1, 3]
assert only_odd_numbers([-5, -4, -3, -2, -1, 1, 2, 3, 4, 5]) == [-5, -3, -1, 1, 3, 5]
assert only_odd_numbers([-4, -3, 1]) == [-3, 1]
assert only_odd_numbers([2, 2, 2, 2, 2]) == []
print("Exercise 67 is correct.")
# Exercise 68
# Write a function definition named only_even_numbers that takes in sequence of numbers and returns the even numbers in a list.
assert only_even_numbers([1, 2, 3]) == [2]
assert only_even_numbers([-5, -4, -3, -2, -1, 1, 2, 3, 4, 5]) == [-4, -2, 2, 4]
assert only_even_numbers([-4, -3, 1]) == [-4]
assert only_even_numbers([1, 1, 1, 1, 1, 1]) == []
print("Exercise 68 is correct.")
# Exercise 69
# Write a function definition named only_positive_numbers that takes in sequence of numbers and returns the positive numbers in a list.
assert only_positive_numbers([1, 2, 3]) == [1, 2, 3]
assert only_positive_numbers([-5, -4, -3, -2, -1, 1, 2, 3, 4, 5]) == [1, 2, 3, 4, 5]
assert only_positive_numbers([-4, -3, 1]) == [1]
print("Exercise 69 is correct.")
# Exercise 70
# Write a function definition named only_negative_numbers that takes in sequence of numbers and returns the negative numbers in a list.
assert only_negative_numbers([1, 2, 3]) == []
assert only_negative_numbers([-5, -4, -3, -2, -1, 1, 2, 3, 4, 5]) == [-5, -4, -3, -2, -1]
assert only_negative_numbers([-4, -3, 1]) == [-4, -3]
print("Exercise 70 is correct.")
# Exercise 71
# Write a function definition named has_evens that takes in sequence of numbers and returns True if there are any even numbers in the sequence
assert has_evens([1, 2, 3]) == True
assert has_evens([2, 5, 6]) == True
assert has_evens([3, 3, 3]) == False
assert has_evens([]) == False
print("Exercise 71 is correct.")
# Exercise 72
# Write a function definition named count_evens that takes in sequence of numbers and returns the number of even numbers
assert count_evens([1, 2, 3]) == 1
assert count_evens([2, 5, 6]) == 2
assert count_evens([3, 3, 3]) == 0
assert count_evens([5, 6, 7, 8] ) == 2
print("Exercise 72 is correct.")
# Exercise 73
# Write a function definition named has_odds that takes in sequence of numbers and returns True if there are any odd numbers in the sequence
assert has_odds([1, 2, 3]) == True
assert has_odds([2, 5, 6]) == True
assert has_odds([3, 3, 3]) == True
assert has_odds([2, 4, 6]) == False
print("Exercise 73 is correct.")
# Exercise 74
# Write a function definition named count_odds that takes in sequence of numbers and returns True if there are any odd numbers in the sequence
assert count_odds([1, 2, 3]) == 2
assert count_odds([2, 5, 6]) == 1
assert count_odds([3, 3, 3]) == 3
assert count_odds([2, 4, 6]) == 0
print("Exercise 74 is correct.")
# Exercise 75
# Write a function definition named count_negatives that takes in sequence of numbers and returns a count of the number of negative numbers
assert count_negatives([1, -2, 3]) == 1
assert count_negatives([2, -5, -6]) == 2
assert count_negatives([3, 3, 3]) == 0
print("Exercise 75 is correct.")
# Exercise 76
# Write a function definition named count_positives that takes in sequence of numbers and returns a count of the number of positive numbers
assert count_positives([1, -2, 3]) == 2
assert count_positives([2, -5, -6]) == 1
assert count_positives([3, 3, 3]) == 3
assert count_positives([-2, -1, -5]) == 0
print("Exercise 76 is correct.")
# Exercise 77
# Write a function definition named only_positive_evens that takes in sequence of numbers and returns a list containing all the positive evens from the sequence
assert only_positive_evens([1, -2, 3]) == []
assert only_positive_evens([2, -5, -6]) == [2]
assert only_positive_evens([3, 3, 4, 6]) == [4, 6]
assert only_positive_evens([2, 3, 4, -1, -5]) == [2, 4]
print("Exercise 77 is correct.")
# Exercise 78
# Write a function definition named only_positive_odds that takes in sequence of numbers and returns a list containing all the positive odd numbers from the sequence
assert only_positive_odds([1, -2, 3]) == [1, 3]
assert only_positive_odds([2, -5, -6]) == []
assert only_positive_odds([3, 3, 4, 6]) == [3, 3]
assert only_positive_odds([2, 3, 4, -1, -5]) == [3]
print("Exercise 78 is correct.")
# Exercise 79
# Write a function definition named only_negative_evens that takes in sequence of numbers and returns a list containing all the negative even numbers from the sequence
assert only_negative_evens([1, -2, 3]) == [-2]
assert only_negative_evens([2, -5, -6]) == [-6]
assert only_negative_evens([3, 3, 4, 6]) == []
assert only_negative_evens([-2, 3, 4, -1, -4]) == [-2, -4]
print("Exercise 79 is correct.")
# Exercise 80
# Write a function definition named only_negative_odds that takes in sequence of numbers and returns a list containing all the negative odd numbers from the sequence
assert only_negative_odds([1, -2, 3]) == []
assert only_negative_odds([2, -5, -6]) == [-5]
assert only_negative_odds([3, 3, 4, 6]) == []
assert only_negative_odds([2, -3, 4, -1, -4]) == [-3, -1]
print("Exercise 80 is correct.")
# Exercise 81
# Write a function definition named shortest_string that takes in a list of strings and returns the shortest string in the list.
assert shortest_string(["kiwi", "mango", "strawberry"]) == "kiwi"
assert shortest_string(["hello", "everybody"]) == "hello"
assert shortest_string(["mary", "had", "a", "little", "lamb"]) == "a"
print("Exercise 81 is correct.")
# Exercise 82
# Write a function definition named longest_string that takes in sequence of strings and returns the longest string in the list.
assert longest_string(["kiwi", "mango", "strawberry"]) == "strawberry"
assert longest_string(["hello", "everybody"]) == "everybody"
assert longest_string(["mary", "had", "a", "little", "lamb"]) == "little"
print("Exercise 82 is correct.")
```
## Working with sets
**Hint** Take a look at the `set` function in Python, the `set` data type, and built-in `set` methods.
```
# Example set function usage
print(set("kiwi"))
print(set([1, 2, 2, 3, 3, 3, 4, 4, 4, 4]))
# Exercise 83
# Write a function definition named get_unique_values that takes in a list and returns a set with only the unique values from that list.
assert get_unique_values(["ant", "ant", "mosquito", "mosquito", "ladybug"]) == {"ant", "mosquito", "ladybug"}
assert get_unique_values(["b", "a", "n", "a", "n", "a", "s"]) == {"b", "a", "n", "s"}
assert get_unique_values(["mary", "had", "a", "little", "lamb", "little", "lamb", "little", "lamb"]) == {"mary", "had", "a", "little", "lamb"}
print("Exercise 83 is correct.")
# Exercise 84
# Write a function definition named get_unique_values_from_two_lists that takes two lists and returns a single set with only the unique values
assert get_unique_values_from_two_lists([5, 1, 2, 3], [3, 4, 5, 5]) == {1, 2, 3, 4, 5}
assert get_unique_values_from_two_lists([1, 1], [2, 2, 3]) == {1, 2, 3}
assert get_unique_values_from_two_lists(["tomato", "mango", "kiwi"], ["eggplant", "tomato", "broccoli"]) == {"tomato", "mango", "kiwi", "eggplant", "broccoli"}
print("Exercise 84 is correct.")
# Exercise 85
# Write a function definition named get_values_in_common that takes two lists and returns a single set with the values that each list has in common
assert get_values_in_common([5, 1, 2, 3], [3, 4, 5, 5]) == {3, 5}
assert get_values_in_common([1, 2], [2, 2, 3]) == {2}
assert get_values_in_common(["tomato", "mango", "kiwi"], ["eggplant", "tomato", "broccoli"]) == {"tomato"}
print("Exercise 85 is correct.")
# Exercise 86
# Write a function definition named get_values_not_in_common that takes two lists and returns a single set with the values that each list does not have in common
assert get_values_not_in_common([5, 1, 2, 3], [3, 4, 5, 5]) == {1, 2, 4}
assert get_values_not_in_common([1, 1], [2, 2, 3]) == {1, 2, 3}
assert get_values_not_in_common(["tomato", "mango", "kiwi"], ["eggplant", "tomato", "broccoli"]) == {"mango", "kiwi", "eggplant", "broccoli"}
print("Exercise 86 is correct.")
```
## Working with Dictionaries
```
# Run this cell in order to have these two dictionary variables defined.
tukey_paper = {
"title": "The Future of Data Analysis",
"author": "John W. Tukey",
"link": "https://projecteuclid.org/euclid.aoms/1177704711",
"year_published": 1962
}
thomas_paper = {
"title": "A mathematical model of glutathione metabolism",
"author": "Rachel Thomas",
"link": "https://www.ncbi.nlm.nih.gov/pubmed/18442411",
"year_published": 2008
}
# Exercise 87
# Write a function named get_paper_title that takes in a dictionary and returns the title property
assert get_paper_title(tukey_paper) == "The Future of Data Analysis"
assert get_paper_title(thomas_paper) == "A mathematical model of glutathione metabolism"
print("Exercise 87 is correct.")
# Exercise 88
# Write a function named get_year_published that takes in a dictionary and returns the value behind the "year_published" key.
assert get_year_published(tukey_paper) == 1962
assert get_year_published(thomas_paper) == 2008
print("Exercise 88 is correct.")
# Run this code to create data for the next two questions
book = {
"title": "Genetic Algorithms and Machine Learning for Programmers",
"price": 36.99,
"author": "Frances Buontempo"
}
# Exercise 89
# Write a function named get_price that takes in a dictionary and returns the price
assert get_price(book) == 36.99
print("Exercise 89 is complete.")
# Exercise 90
# Write a function named get_book_author that takes in a dictionary (the above declared book variable) and returns the author's name
assert get_book_author(book) == "Frances Buontempo"
print("Exercise 90 is complete.")
```
## Working with Lists of Dictionaries
**Hint** If you need an example of lists of dictionaries, see
- [Getting Started With a List of Dictionaries](https://colab.research.google.com/github/ryanorsinger/list_of_dictionaries/blob/main/getting_started.ipynb)
- [Practice Exercises for List of Dictionaries](https://colab.research.google.com/github/ryanorsinger/list_of_dictionaries/blob/main/exercises.ipynb)
- [Companion Video](https://www.youtube.com/watch?v=pPdEahZgv8U)
```
# Run this cell in order to have some setup data for the next exercises
books = [
{
"title": "Genetic Algorithms and Machine Learning for Programmers",
"price": 36.99,
"author": "Frances Buontempo"
},
{
"title": "The Visual Display of Quantitative Information",
"price": 38.00,
"author": "Edward Tufte"
},
{
"title": "Practical Object-Oriented Design",
"author": "Sandi Metz",
"price": 30.47
},
{
"title": "Weapons of Math Destruction",
"author": "Cathy O'Neil",
"price": 17.44
}
]
# Exercise 91
# Write a function named get_number_of_books that takes in a list of objects and returns the number of dictionaries in that list.
assert get_number_of_books(books) == 4
print("Exercise 91 is complete.")
# Exercise 92
# Write a function named total_of_book_prices that takes in a list of dictionaries and returns the sum total of all the book prices added together
assert total_of_book_prices(books) == 122.9
print("Exercise 92 is complete.")
# Exercise 93
# Write a function named get_average_book_price that takes in a list of dictionaries and returns the average book price.
assert get_average_book_price(books) == 30.725
print("Exercise 93 is complete.")
# Exercise 94
# Write a function called highest_price_book that takes in the above defined list of dictionaries "books" and returns the dictionary containing the title, price, and author of the book with the highest priced book.
# Hint: Much like sometimes start functions with a variable set to zero, you may want to create a dictionary with the price set to zero to compare to each dictionary's price in the list
assert highest_price_book(books) == {
"title": "The Visual Display of Quantitative Information",
"price": 38.00,
"author": "Edward Tufte"
}
print("Exercise 94 is complete")
# Exercise 95
# Write a function called lowest_priced_book that takes in the above defined list of dictionaries "books" and returns the dictionary containing the title, price, and author of the book with the lowest priced book.
# Hint: Much like sometimes start functions with a variable set to zero or float('inf'), you may want to create a dictionary with the price set to float('inf') to compare to each dictionary in the list
assert lowest_price_book(books) == {
"title": "Weapons of Math Destruction",
"author": "Cathy O'Neil",
"price": 17.44
}
print("Exercise 95 is complete.")
shopping_cart = {
"tax": .08,
"items": [
{
"title": "orange juice",
"price": 3.99,
"quantity": 1
},
{
"title": "rice",
"price": 1.99,
"quantity": 3
},
{
"title": "beans",
"price": 0.99,
"quantity": 3
},
{
"title": "chili sauce",
"price": 2.99,
"quantity": 1
},
{
"title": "chocolate",
"price": 0.75,
"quantity": 9
}
]
}
# Exercise 96
# Write a function named get_tax_rate that takes in the above shopping cart as input and returns the tax rate.
# Hint: How do you access a key's value on a dictionary? The tax rate is one key of the entire shopping_cart dictionary.
assert get_tax_rate(shopping_cart) == .08
print("Exercise 96 is complete")
# Exercise 97
# Write a function named number_of_item_types that takes in the shopping cart as input and returns the number of unique item types in the shopping cart.
# We're not yet using the quantity of each item, but rather focusing on determining how many different types of items are in the cart.
assert number_of_item_types(shopping_cart) == 5
print("Exercise 97 is complete.")
# Exercise 98
# Write a function named total_number_of_items that takes in the shopping cart as input and returns the total number all item quantities.
# This should return the sum of all of the quantities from each item type
assert total_number_of_items(shopping_cart) == 17
print("Exercise 98 is complete.")
# Exercise 99
# Write a function named get_average_item_price that takes in the shopping cart as an input and returns the average of all the item prices.
# Hint - This should determine the total price divided by the number of types of items. This does not account for each item type's quantity.
assert get_average_item_price(shopping_cart) == 2.1420000000000003
print("Exercise 99 is complete.")
# Exercise 100
# Write a function named get_average_spent_per_item that takes in the shopping cart and returns the average of summing each item's quanties times that item's price.
# Hint: You may need to set an initial total price and total total quantity to zero, then sum up and divide that total price by the total quantity
assert get_average_spent_per_item(shopping_cart) == 1.333529411764706
print("Exercise 100 is complete.")
# Exercise 101
# Write a function named most_spent_on_item that takes in the shopping cart as input and returns the dictionary associated with the item that has the highest price*quantity.
# Be sure to do this as programmatically as possible.
# Hint: Similarly to how we sometimes begin a function with setting a variable to zero, we need a starting place:
# Hint: Consider creating a variable that is a dictionary with the keys "price" and "quantity" both set to 0. You can then compare each item's price and quantity total to the one from "most"
assert most_spent_on_item(shopping_cart) == {
"title": "chocolate",
"price": 0.75,
"quantity": 9
}
print("Exercise 101 is complete.")
```
Created by [Ryan Orsinger](https://ryanorsinger.com)
Source code on [https://github.com/ryanorsinger/101-exercises](https://github.com/ryanorsinger/101-exercises)
| github_jupyter |
```
%matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import time
import h5py
import keras
import pandas as pd
import math
import joblib
from IPython.display import display
```
# Load Models
```
generator_path = "saved_models/generator.hdf5"
generator = keras.models.load_model(generator_path)
# Need to re-include loss functions so model knows how to load them
def kl_loss(x, x_decoded_mean):
kl_loss = - 0.5 * K.sum(1. + z_log_var - K.square(z_mean) - K.exp(z_log_var), axis=-1)
return K.mean(kl_loss)
def logx_loss(x, x_decoded_mean):
loss = ( 0.5 * math.log(2 * math.pi)
+ 0.5 * K.log(_x_decoded_var + var_epsilon)
+ 0.5 * K.square(x - x_decoded_mean) / (_x_decoded_var + var_epsilon))
loss = K.sum(loss, axis=-1)
return K.mean(loss)
def vae_loss(x, x_decoded_mean):
return logx_loss(x, x_decoded_mean) + kl_loss(x, x_decoded_mean)
vae_path = "saved_models/vae.hdf5"
vae = keras.models.load_model(vae_path, custom_objects={'vae_loss': vae_loss}, compile=False)
pca_model_file = 'saved_models/pca_500.pkl'
pca = joblib.load(pca_model_file)
n_components = pca.n_components_
pca
```
# Sample Some Random Images
```
digit_size = 32
img_rows, img_cols, img_chns = 32, 32, 3
latent_dim = 512
```
Using the generator network (decoder), we'll sample some from our isotropic standard Gaussians and generate a grid of 10x10 images.
```
n = 10
figure = np.zeros((digit_size * n, digit_size * n, img_chns))
for i in range(n):
for j in range(n):
z_sample = np.random.normal(size=latent_dim).reshape(1, latent_dim)
x_decoded = generator.predict(z_sample, batch_size=1)
x_recon = np.clip(pca.inverse_transform(x_decoded), 0., .999)
x_recon *= 255.
digit = x_recon.reshape(digit_size, digit_size, img_chns)
d_x = i * digit_size
d_y = j * digit_size
figure[d_x:d_x + digit_size, d_y:d_y + digit_size] = digit.astype(int)
plt.figure(figsize=(10, 10))
plt.imshow(figure)
plt.show()
```
Definitely a quite blurry but definitely some good images that came out. There are a few that are much clearer than others. VAEs are known to produce blurry images compared to GANs.
# VAE Analogies
For analogies, we'll take one of the images from our original sample and then pump it through the entire VAE pipeline. Under the hood, the network should estimate the latent mean and variance for that sample, and then use the generator network to decode it into a "similar" looking image that sampled from our shifted/scaled latent Gaussian random variable.
```
f = h5py.File("../../data/svhn/svhn_format_2.hdf5", "r")
X_dataset, y_dataset = list(f.items())
# Use a nice round number divisible by our batch size
N=630000
X_dataset, y_dataset = np.moveaxis(X_dataset[1][:N], 1, 3), y_dataset[1][:N]
print ("Full Dataset - DType X=%s, y=%s" % (X_dataset.dtype, y_dataset.dtype))
print ("Full Dataset - Shape X=%s, y=%s" % (X_dataset.shape, y_dataset.shape))
def random_image(digit):
assert 0 <= digit <= 9
while True:
index = np.random.randint(0, len(X_dataset))
if y_dataset[index] == digit:
return X_dataset[index]
batch_size = 1000
figure = np.zeros((digit_size * 10, digit_size * 10, img_chns))
for i in range(0, 10):
image = random_image(i)
# Create batch to send through our network
X_sample = np.stack([image] * batch_size)
batch = pca.transform(X_sample.reshape(batch_size, -1) / 255.)
x_decoded = vae.predict(batch, batch_size=batch_size)
# Convert back to RGB format
x_recon = np.clip(pca.inverse_transform(x_decoded), 0.0, .999)
x_recon *= 255.
analogue = x_recon.reshape(batch_size, digit_size, digit_size, img_chns)
# First column is our original imge
d_x = i * digit_size
d_y = 0
figure[d_x:d_x + digit_size, d_y:d_y + digit_size] = image.astype(int)
last = [image]
# Rest of the columns are analogies, attempting to select ones that are different
for j in range(1, 10):
d_x = i * digit_size
d_y = j * digit_size
best_val = 0
best_img = None
for k in range(j * 100, (j + 1) * 100):
val = np.mean([np.abs(l - analogue[k]).mean() for l in last])
if val >= best_val:
best_val = val
best_img = analogue[k]
figure[d_x:d_x + digit_size, d_y:d_y + digit_size] = best_img.astype(int)
last.append(best_img)
plt.figure(figsize=(10, 10))
plt.imshow(figure)
plt.show()
```
Analogies are actually pretty interesting. I was hoping for rows of all the same number, not quite that but close! There is of course similarities in terms of colour and style. The generated images are much blurrier, probably because the model hasn't fit the data that well.
| github_jupyter |
## Training metrics
*Metrics* for training fastai models are simply functions that take `input` and `target` tensors, and return some metric of interest for training. You can write your own metrics by defining a function of that type, and passing it to [`Learner`](/basic_train.html#Learner) in the [`metrics`](/metrics.html#metrics) parameter, or use one of the following pre-defined functions.
```
from fastai.gen_doc.nbdoc import *
from fastai.basics import *
```
## Predefined metrics:
```
show_doc(accuracy)
jekyll_warn("This metric is intended for classification of objects belonging to a single class.")
show_doc(accuracy_thresh)
```
Prediction are compared to `thresh` after `sigmoid` is maybe applied. Then we count the numbers that match the targets.
```
jekyll_note("This function is intended for one-hot-encoded targets (often in a multiclassification problem).")
show_doc(top_k_accuracy)
show_doc(dice)
show_doc(error_rate)
show_doc(mean_squared_error)
show_doc(mean_absolute_error)
show_doc(mean_squared_logarithmic_error)
show_doc(exp_rmspe)
show_doc(root_mean_squared_error)
show_doc(fbeta)
```
`beta` determines the value of the fbeta applied, `eps` is there for numeric stability. If `sigmoid=True`, a sigmoid is applied to the predictions before comparing them to `thresh` then to the targets. See the [F1 score wikipedia page](https://en.wikipedia.org/wiki/F1_score) for details on the fbeta score.
```
jekyll_note("This function is intended for one-hot-encoded targets (often in a multiclassification problem).")
show_doc(explained_variance)
show_doc(r2_score)
```
The following metrics are classes, don't forget to instantiate them when you pass them to a [`Learner`](/basic_train.html#Learner).
```
show_doc(RMSE, title_level=3)
show_doc(ExpRMSPE, title_level=3)
show_doc(Precision, title_level=3)
show_doc(Recall, title_level=3)
show_doc(FBeta, title_level=3)
show_doc(R2Score, title_level=3)
show_doc(ExplainedVariance, title_level=3)
show_doc(MatthewsCorreff, title_level=3)
```
Ref.: https://github.com/scikit-learn/scikit-learn/blob/bac89c2/sklearn/metrics/classification.py
```
show_doc(KappaScore, title_level=3)
```
Ref.: https://github.com/scikit-learn/scikit-learn/blob/bac89c2/sklearn/metrics/classification.py
[`KappaScore`](/metrics.html#KappaScore) supports linear and quadratic weights on the off-diagonal cells in the [`ConfusionMatrix`](/metrics.html#ConfusionMatrix), in addition to the default unweighted calculation treating all misclassifications as equally weighted. Leaving [`KappaScore`](/metrics.html#KappaScore)'s `weights` attribute as `None` returns the unweighted Kappa score. Updating `weights` to "linear" means off-diagonal ConfusionMatrix elements are weighted in linear proportion to their distance from the diagonal; "quadratic" means weights are squared proportional to their distance from the diagonal.
Specify linear or quadratic weights, if using, by first creating an instance of the metric and then updating the `weights` attribute, similar to as follows:
```
kappa = KappaScore()
kappa.weights = "quadratic"
learn = cnn_learner(data, model, metrics=[error_rate, kappa])
```
```
show_doc(ConfusionMatrix, title_level=3)
show_doc(MultiLabelFbeta, title_level=3)
```
[`MultiLabelFbeta`](/metrics.html#MultiLabelFbeta) implements mutlilabel classification fbeta score similar to [scikit-learn's](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.f1_score.html) as a [`LearnerCallback`](/basic_train.html#LearnerCallback). Average options: ["micro", "macro", "weighted", "none"]. Intended to use with one-hot encoded targets with 1s and 0s.
## Creating your own metric
Creating a new metric can be as simple as creating a new function. If your metric is an average over the total number of elements in your dataset, just write the function that will compute it on a batch (taking `pred` and `targ` as arguments). It will then be automatically averaged over the batches (taking their different sizes into account).
Sometimes metrics aren't simple averages however. If we take the example of precision for instance, we have to divide the number of true positives by the number of predictions we made for that class. This isn't an average over the number of elements we have in the dataset, we only consider those where we made a positive prediction for a specific thing. Computing the precision for each batch, then averaging them will yield to a result that may be close to the real value, but won't be it exactly (and it really depends on how you deal with special case of 0 positive predictions).
This why in fastai, every metric is implemented as a callback. If you pass a regular function, the library transforms it to a proper callback called `AverageCallback`. The callback metrics are only called during the validation phase, and only for the following events:
- <code>on_epoch_begin</code> (for initialization)
- <code>on_batch_begin</code> (if we need to have a look at the input/target and maybe modify them)
- <code>on_batch_end</code> (to analyze the last results and update our computation)
- <code>on_epoch_end</code>(to wrap up the final result that should be added to `last_metrics`)
As an example, the following code is the exact implementation of the [`AverageMetric`](/callback.html#AverageMetric) callback that transforms a function like [`accuracy`](/metrics.html#accuracy) into a metric callback.
```
class AverageMetric(Callback):
"Wrap a `func` in a callback for metrics computation."
def __init__(self, func):
# If it's a partial, use func.func
name = getattr(func,'func',func).__name__
self.func, self.name = func, name
def on_epoch_begin(self, **kwargs):
"Set the inner value to 0."
self.val, self.count = 0.,0
def on_batch_end(self, last_output, last_target, **kwargs):
"Update metric computation with `last_output` and `last_target`."
if not is_listy(last_target): last_target=[last_target]
self.count += last_target[0].size(0)
val = self.func(last_output, *last_target)
self.val += last_target[0].size(0) * val.detach().cpu()
def on_epoch_end(self, last_metrics, **kwargs):
"Set the final result in `last_metrics`."
return add_metrics(last_metrics, self.val/self.count)
```
Here [`add_metrics`](/torch_core.html#add_metrics) is a convenience function that will return the proper dictionary for us:
```python
{'last_metrics': last_metrics + [self.val/self.count]}
```
And here is another example that properly computes the precision for a given class.
```
class Precision(Callback):
def on_epoch_begin(self, **kwargs):
self.correct, self.total = 0, 0
def on_batch_end(self, last_output, last_target, **kwargs):
preds = last_output.argmax(1)
self.correct += ((preds==0) * (last_target==0)).float().sum()
self.total += (preds==0).float().sum()
def on_epoch_end(self, last_metrics, **kwargs):
return add_metrics(last_metrics, self.correct/self.total)
```
The following custom callback class example measures peak RAM usage during each epoch:
```
import tracemalloc
class TraceMallocMetric(Callback):
def __init__(self):
super().__init__()
self.name = "peak RAM"
def on_epoch_begin(self, **kwargs):
tracemalloc.start()
def on_epoch_end(self, last_metrics, **kwargs):
current, peak = tracemalloc.get_traced_memory()
tracemalloc.stop()
return add_metrics(last_metrics, torch.tensor(peak))
```
To deploy it, you need to pass an instance of this custom metric in the [`metrics`](/metrics.html#metrics) argument:
```
learn = cnn_learner(data, model, metrics=[accuracy, TraceMallocMetric()])
learn.fit_one_cycle(3, max_lr=1e-2)
```
And then the output changes to:
```
Total time: 00:54
epoch train_loss valid_loss accuracy peak RAM
1 0.333352 0.084342 0.973800 2395541.000000
2 0.096196 0.038386 0.988300 2342145.000000
3 0.048722 0.029234 0.990200 2342680.000000
```
As mentioner earlier, using the [`metrics`](/metrics.html#metrics) argument with a custom metrics class is limited in the number of phases of the callback system it can access, it can only return one numerical value and as you can see its output is hardcoded to have 6 points of precision in the output, even if the number is an int.
To overcome these limitations callback classes should be used instead.
For example, the following class:
* uses phases not available for the metric classes
* it reports 3 columns, instead of just one
* its column report ints, instead of floats
```
import tracemalloc
class TraceMallocMultiColMetric(LearnerCallback):
_order=-20 # Needs to run before the recorder
def __init__(self, learn):
super().__init__(learn)
self.train_max = 0
def on_train_begin(self, **kwargs):
self.learn.recorder.add_metric_names(['used', 'max_used', 'peak'])
def on_batch_end(self, train, **kwargs):
# track max memory usage during the train phase
if train:
current, peak = tracemalloc.get_traced_memory()
self.train_max = max(self.train_max, current)
def on_epoch_begin(self, **kwargs):
tracemalloc.start()
def on_epoch_end(self, last_metrics, **kwargs):
current, peak = tracemalloc.get_traced_memory()
tracemalloc.stop()
return add_metrics(last_metrics, [current, self.train_max, peak])
```
Note, that it subclasses [`LearnerCallback`](/basic_train.html#LearnerCallback) and not [`Callback`](/callback.html#Callback), since the former provides extra features not available in the latter.
Also `_order=-20` is crucial - without it the custom columns will not be added - it tells the callback system to run this callback before the recorder system.
To deploy it, you need to pass the name of the class (not an instance!) of the class in the `callback_fns` argument. This is because the `learn` object doesn't exist yet, and it's required to instantiate `TraceMallocMultiColMetric`. The system will do it for us automatically as soon as the learn object has been created.
```
learn = cnn_learner(data, model, metrics=[accuracy], callback_fns=TraceMallocMultiColMetric)
learn.fit_one_cycle(3, max_lr=1e-2)
```
And then the output changes to:
```
Total time: 00:53
epoch train_loss valid_loss accuracy used max_used peak
1 0.321233 0.068252 0.978600 156504 2408404 2419891
2 0.093551 0.032776 0.988500 79343 2408404 2348085
3 0.047178 0.025307 0.992100 79568 2408404 2342754
```
Another way to do the same is by using `learn.callbacks.append`, and this time we need to instantiate `TraceMallocMultiColMetric` with `learn` object which we now have, as it is called after the latter was created:
```
learn = cnn_learner(data, model, metrics=[accuracy])
learn.callbacks.append(TraceMallocMultiColMetric(learn))
learn.fit_one_cycle(3, max_lr=1e-2)
```
Configuring the custom metrics in the `learn` object sets them to run in all future [`fit`](/basic_train.html#fit)-family calls. However, if you'd like to configure it for just one call, you can configure it directly inside [`fit`](/basic_train.html#fit) or [`fit_one_cycle`](/train.html#fit_one_cycle):
```
learn = cnn_learner(data, model, metrics=[accuracy])
learn.fit_one_cycle(3, max_lr=1e-2, callbacks=TraceMallocMultiColMetric(learn))
```
And to stress the differences:
* the `callback_fns` argument expects a classname or a list of those
* the [`callbacks`](/callbacks.html#callbacks) argument expects an instance of a class or a list of those
* `learn.callbacks.append` expects a single instance of a class
For more examples, look inside fastai codebase and its test suite, search for classes that subclass either [`Callback`](/callback.html#Callback), [`LearnerCallback`](/basic_train.html#LearnerCallback) and subclasses of those two.
Finally, while the above examples all add to the metrics, it's not a requirement. A callback can do anything it wants and it is not required to add its outcomes to the metrics printout.
## Undocumented Methods - Methods moved below this line will intentionally be hidden
```
show_doc(FBeta.on_batch_end)
show_doc(FBeta.on_epoch_begin)
show_doc(FBeta.on_epoch_end)
show_doc(mae)
show_doc(msle)
show_doc(mse)
show_doc(rmse)
show_doc(Precision.on_epoch_end)
show_doc(FBeta.on_train_end)
show_doc(KappaScore.on_epoch_end)
show_doc(MatthewsCorreff.on_epoch_end)
show_doc(FBeta.on_train_begin)
show_doc(RMSE.on_epoch_end)
show_doc(ConfusionMatrix.on_train_begin)
show_doc(ConfusionMatrix.on_batch_end)
show_doc(ConfusionMatrix.on_epoch_end)
show_doc(Recall.on_epoch_end)
show_doc(ExplainedVariance.on_epoch_end)
show_doc(ExpRMSPE.on_epoch_end)
show_doc(ConfusionMatrix.on_epoch_begin)
show_doc(R2Score.on_epoch_end)
```
## New Methods - Please document or move to the undocumented section
| github_jupyter |
# Open exploration
In this second tuturial, I will showcase how to use the ema_workbench for performing open exploration. This tuturial will continue with the same example as used in the previos tuturial.
## some background
In exploratory modeling, we are interested in understanding how regions in the uncertainty space and/or the decision space map to the whole outcome space, or partitions thereof. There are two general approaches for investigating this mapping. The first one is through systematic sampling of the uncertainty or decision space. This is sometimes also known as open exploration. The second one is to search through the space in a directed manner using some type of optimization approach. This is sometimes also known as directed search.
The workbench support both open exploration and directed search. Both can be applied to investigate the mapping of the uncertainty space and/or the decision space to the outcome space. In most applications, search is used for finding promising mappings from the decision space to the outcome space, while exploration is used to stress test these mappings under a whole range of possible resolutions to the various uncertainties. This need not be the case however. Optimization can be used to discover the worst possible scenario, while sampling can be used to get insight into the sensitivity of outcomes to the various decision levers.
## open exploration
To showcase the open exploration functionality, let's start with a basic example using the Direct Policy Search (DPS) version of the lake problem [(Quinn et al 2017)](https://doi.org/10.1016/j.envsoft.2017.02.017). This is the same model as we used in the general introduction. Note that for convenience, I have moved the code for the model to a module called dps_lake_model.py, which I import here for further use.
We are going to simultaneously sample over uncertainties and decision levers. We are going to generate 1000 scenarios and 5 policies, and see how they jointly affect the outcomes. A *scenario* is understood as a point in the uncertainty space, while a *policy* is a point in the decision space. The combination of a scenario and a policy is called *experiment*. The uncertainty space is spanned by uncertainties, while the decision space is spanned by levers.
Both uncertainties and levers are instances of *RealParameter* (a continuous range), *IntegerParameter* (a range of integers), or *CategoricalParameter* (an unorder set of things). By default, the workbench will use Latin Hypercube sampling for generating both the scenarios and the policies. Each policy will be always evaluated over all scenarios (i.e. a full factorial over scenarios and policies).
```
from ema_workbench import (RealParameter, ScalarOutcome, Constant,
Model)
from dps_lake_model import lake_model
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
model = Model('lakeproblem', function=lake_model)
#specify uncertainties
model.uncertainties = [RealParameter('b', 0.1, 0.45),
RealParameter('q', 2.0, 4.5),
RealParameter('mean', 0.01, 0.05),
RealParameter('stdev', 0.001, 0.005),
RealParameter('delta', 0.93, 0.99)]
# set levers
model.levers = [RealParameter("c1", -2, 2),
RealParameter("c2", -2, 2),
RealParameter("r1", 0, 2),
RealParameter("r2", 0, 2),
RealParameter("w1", 0, 1)]
#specify outcomes
model.outcomes = [ScalarOutcome('max_P'),
ScalarOutcome('utility'),
ScalarOutcome('inertia'),
ScalarOutcome('reliability')]
# override some of the defaults of the model
model.constants = [Constant('alpha', 0.41),
Constant('nsamples', 150),
Constant('myears', 100)]
from ema_workbench import (MultiprocessingEvaluator, ema_logging,
perform_experiments)
ema_logging.log_to_stderr(ema_logging.INFO)
with MultiprocessingEvaluator(model, n_processes=7) as evaluator:
experiments, outcomes = evaluator.perform_experiments(scenarios=1000, policies=5)
```
### Visual analysis
Having generated these results, the next step is to analyze them and see what we can learn from the results. The workbench comes with a variety of techniques for this analysis. A simple first step is to make a few quick visualizations of the results. The workbench has convenience functions for this, but it also possible to create your own visualizations using the scientific Python stack.
```
from ema_workbench.analysis import pairs_plotting
fig, axes = pairs_plotting.pairs_scatter(experiments, outcomes, group_by='policy',
legend=False)
fig.set_size_inches(8,8)
plt.show()
```
Often, it is convenient to separate the process of performing the experiments from the analysis. To make this possible, the workbench offers convenience functions for storing results to disc and loading them from disc. The workbench will store the results in a tarbal with .csv files and separate metadata files. This is a convenient format that has proven sufficient over the years.
```python
from ema_workbench import save_results
save_results(results, '1000 scenarios 5 policies.tar.gz')
from ema_workbench import load_results
results = load_results('1000 scenarios 5 policies.tar.gz')
```
## advanced analysis
In addition to visual analysis, the workbench comes with a variety of techniques to perform a more in-depth analysis of the results. In addition, other analyses can simply be performed by utilizing the scientific python stack. The workbench comes with
* [Scenario Discovery](https://waterprogramming.wordpress.com/2015/08/05/scenario-discovery-in-python/), a model driven approach to scenario development
* [Feature Scoring](https:/doi.org/10.1016/j.envsoft.2018.06.011), a poor man's alternative to global sensitivity analysis
* [Dimensional stacking](https://www.onepetro.org/conference-paper/SPE-174774-MS), a quick visual approach drawing on feature scoring to enable scenario discovery. This approach has received limited attention in the literature. The implementation in the workbench replaces the rule mining approach with a feature scoring approach.
* [Regional sensitivity analysis](https://doi.org/10.1016/j.envsoft.2016.02.008)
### Scenario Discovery
A detailed discussion on scenario discovery can be found in an [earlier blogpost](https://waterprogramming.wordpress.com/2015/08/05/scenario-discovery-in-python/). For completeness, I provide a code snippet here. Compared to the previous blog post, there is one small change. The library mpld3 is currently not being maintained and broken on Python 3.5. and higher. To still utilize the interactive exploration of the trade offs within the notebook, one could use the interactive back-end [(% matplotlib notebook)](http://ipython.readthedocs.io/en/stable/interactive/magics.html#magic-matplotlib).
```
from ema_workbench.analysis import prim
x = experiments
y = outcomes['max_P'] <0.8
prim_alg = prim.Prim(x, y, threshold=0.8)
box1 = prim_alg.find_box()
box1.show_tradeoff()
plt.show()
```
We can inspect any of the points on the trade off curve using the inspect method. As shown, we can show the results either in a table format or in a visual format.
```
box1.inspect(43)
box1.inspect(43, style='graph')
plt.show()
box1.show_pairs_scatter(43)
plt.show()
```
### feature scoring
Feature scoring is a family of techniques often used in machine learning to identify the most relevant features to include in a model. This is similar to one of the use cases for global sensitivity analysis, namely factor prioritisation. The main advantage of feature scoring techniques is that they impose no specific constraints on the experimental design, while they can handle real valued, integer valued, and categorical valued parameters. The workbench supports multiple techniques, the most useful of which generally is extra trees [(Geurts et al. 2006)](https://link.springer.com/article/10.1007/s10994-006-6226-1).
For this example, we run feature scoring for each outcome of interest. We can also run it for a specific outcome if desired. Similarly, we can choose if we want to run in regression mode or classification mode. The later is applicable if the outcome is a categorical variable and the results should be interpreted similar to regional sensitivity analysis results. For more details, see the documentation.
```
from ema_workbench.analysis import feature_scoring
x = experiments
y = outcomes
fs = feature_scoring.get_feature_scores_all(x, y)
sns.heatmap(fs, cmap='viridis', annot=True)
plt.show()
```
From the results, we see that max_P is primarily influenced by b, while utility is driven by delta, for inertia and reliability the situation is a little bit less clear cut.
The foregoing feature scoring was using the raw values of the outcomes. However, in scenario discovery applications, we are typically dealing with a binary clasification. This might produce slightly different results as demonstrated below
```
x = experiments
y = outcomes['max_P'] <0.8
fs, alg = feature_scoring.get_ex_feature_scores(x, y, mode=feature_scoring.CLASSIFICATION)
fs.sort_values(ascending=False, by=1)
```
Here we ran extra trees feature scoring on a binary vector for *max_P*. the *b* parameter is still important, similar to in the previous case, but the introduction of the binary classifiaction now also highlights some addtional parameters as being potentially relevant.
### dimensional stacking
Dimensional stacking was suggested as a more visual approach to scenario discovery. It involves two steps: identifying the most important uncertainties that affect system behavior, and creating a pivot table using the most influential uncertainties. In order to do this, we first need, as in scenario discovery, specify the outcomes that are of interest. The creating of the pivot table involves binning the uncertainties. More details can be found in [Suzuki et al. (2015)](https://www.onepetro.org/conference-paper/SPE-174774-MS) or by looking through the code in the workbench. Compared to Suzuki et al, the workbench uses feature scoring for determining the most influential uncertainties. The code is set up in a modular way so other approaches to global sensitivity analysis can easily be used as well if so desired.
```
from ema_workbench.analysis import dimensional_stacking
x = experiments
y = outcomes['max_P'] <0.8
dimensional_stacking.create_pivot_plot(x,y, 2, nbins=3)
plt.show()
```
We can see from this visual that if B is high, while Q is high, we have a high concentration of cases where pollution stays below 0.8. The mean and stdev have some limited additional influence. By playing around with an alternative number of bins, or different number of layers, patterns can be coarsened or refined.
### regional sensitivity analysis
A fourth approach for supporting scenario discovery is to perform a regional sensitivity analysis. The workbench implements a visual approach based on plotting the empirical CDF given a classification vector. Please look at section 3.4 in [Pianosi et al (2016)](http://www.sciencedirect.com/science/article/pii/S1364815216300287#sec3.4) for more details.
```
from ema_workbench.analysis import regional_sa
from numpy.lib import recfunctions as rf
sns.set_style('white')
# model is the same across experiments
x = experiments.copy()
x = x.drop('model', axis=1)
y = outcomes['max_P'] < 0.8
fig = regional_sa.plot_cdfs(x,y)
sns.despine()
plt.show()
```
The above results clearly show that both B and Q are important. to a lesser extend, the mean also is relevant.
## More advanced sampling techniques
The workbench can also be used for more advanced sampling techniques. To achieve this, it relies on [SALib](http://salib.readthedocs.io/en/latest/). On the workbench side, the only change is to specify the sampler we want to use. Next, we can use SALib directly to perform the analysis. To help with this, the workbench provides a convenience function for generating the problem dict which SALib provides. The example below focusses on performing SOBOL on the uncertainties, but we could do the exact same thing with the levers instead. The only changes required would be to set `lever_sampling` instead of `uncertainty_sampling`, and get the SALib problem dict based on the levers.
```
from SALib.analyze import sobol
from ema_workbench.em_framework.salib_samplers import get_SALib_problem
with MultiprocessingEvaluator(model) as evaluator:
sa_results = evaluator.perform_experiments(scenarios=1000,
uncertainty_sampling='sobol')
experiments, outcomes = sa_results
problem = get_SALib_problem(model.uncertainties)
Si = sobol.analyze(problem, outcomes['max_P'],
calc_second_order=True, print_to_console=False)
```
We have now completed the sobol analysis and have calculated the metrics. What remains is to visualize the metrics. Which can be done as shown below, focussing on St and S1. The error bars indicate the confidence intervals.
```
scores_filtered = {k:Si[k] for k in ['ST','ST_conf','S1','S1_conf']}
Si_df = pd.DataFrame(scores_filtered, index=problem['names'])
sns.set_style('white')
fig, ax = plt.subplots(1)
indices = Si_df[['S1','ST']]
err = Si_df[['S1_conf','ST_conf']]
indices.plot.bar(yerr=err.values.T,ax=ax)
fig.set_size_inches(8,6)
fig.subplots_adjust(bottom=0.3)
plt.show()
```
| github_jupyter |
# Task 4: Time-dependent general Hartree-Fock solver
Having constructed and solved the general Hartree-Fock ground state problem, we now set out to shine a monochromatic dipole laser on the system.
This means appending a time-dependent operator to the Hamiltonian.
The operator describes a semi-classical electric field in the dipole approximation in the length gauge.
For a more thorough discussion of this operator check out chapters 2 - 2.4 in {cite}`joachain2012atoms`.
The operator will in the one-dimensional case be described by
\begin{align}
\hat{h}_I(t) = -f(t) \hat{d},
\end{align}
where $\hat{d} \equiv q\hat{x}$ is the dipole moment operator with $q = -1$ the electron charge, and $f(t)$ the time-dependent laser field.
From {cite}`zanghellini_2004` we have $f(t) = \mathcal{E}_0 \sin(\omega t)$, (note that $\omega$ is not necessarily the same as the harmonic oscillator well frequency) which describes a monochromatic laser field that is always active.
## Time-dependent Hartree-Fock
In the time-dependent Hartree-Fock method we use an ansatz for the many-body wave function as
\begin{align}
| \Psi(t) \rangle = | \Phi(t) \rangle = | \phi_1(t) \phi_2(t) \dots \phi_n(t) \rangle,
\end{align}
where the time-dependence is kept in the molecular orbitals.
```{note}
Add derivation of the time-dependent Hartree-Fock equations.
```
The time-evolution of the molecular orbitals is described by the time-dependent Hartree-Fock equation
\begin{align}
i \frac{\text{d}}{\text{d} t} | \phi_p(t) \rangle
= \hat{f}(t) | \phi_p(t) \rangle,
\end{align}
where $\hat{f}(t)$ is the time-dependent Fock operator and we have set $\hbar = 1$.
We have chosen to expand the time-independent molecular orbitals in a known basis of atomic orbitals (the harmonic oscillator eigenfunctions).
Here as well we choose to expand our time-dependent molecular orbitals in a time-independent basis of atomic orbitals, and let the time-evolution occur in the coefficients.
That is,
\begin{align}
| \phi_p(t) \rangle
= \sum_{\mu = 1}^{l} C_{\mu p}(t) | \psi_{\mu} \rangle,
\end{align}
where $\{\psi_{\mu}\}_{\mu = 1}^{l}$ is a time-independent atomic orbital basis which we assume to be orthonormal.
Inserting this expansion into the time-dependent Hartree-Fock equations and left-projecting with $\langle \psi_{\mu}|$ we get
\begin{gather}
i \frac{\text{d}}{\text{d} t} \sum_{\nu = 1}^{l} C_{\nu p}(t) | \psi_{\nu} \rangle
= \hat{f}(t) \sum_{\nu = 1}^{l} C_{\nu p}(t) | \psi_{\nu} \rangle
\\
\implies
i \sum_{\nu = 1}^{l} \dot{C}_{\nu p}(t)
\langle \psi_{\mu} | \psi_{\nu} \rangle
= i \dot{C}_{\mu p}(t)
= \langle \psi_{\mu} | \hat{f}(t) | \psi_{\nu} \rangle C_{\nu p}(t)
= f_{\mu \nu}(t) C_{\nu p}(t),
\end{gather}
where we now need to find the matrix elements of the time-dependent Fock operator in the atomic orbital basis.
As the laser field interaction operator $\hat{h}_I(t)$ is a one-body operator, the time-dependent Fock operator will need to add this term.
| github_jupyter |
<a href="https://colab.research.google.com/github/annasajkh/Face-Generator/blob/main/Conv_AutoEncoder.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
from google.colab import drive
drive.mount('/content/drive')
import os
os.environ["KAGGLE_CONFIG_DIR"] = "/content/drive/MyDrive/kaggle"
!kaggle datasets download -d greatgamedota/ffhq-face-data-set
!unzip ffhq-face-data-set.zip
import glob
from PIL import Image
import numpy as np
imgs = []
count = 0
for file in glob.glob("thumbnails128x128/*"):
imgs.append(np.array(Image.open(file).resize((64,64))) / 255)
count += 1
if count == 20_000:
break
np.save("face_dataset.npy", np.array(imgs))
%cp drive/MyDrive/face_dataset.npy /content/
%cp drive/MyDrive/models/face_model.pkl /content/
from tqdm import tqdm
from torch.nn.modules.linear import Linear
import torch
import torch.nn as nn
from torch.optim import Adam
from tqdm import tqdm
import numpy as np
dataset = torch.from_numpy(np.load("face_dataset.npy")).float()
print(len(dataset))
if torch.cuda.is_available():
dataset = dataset.cuda()
dataset = dataset.view(len(dataset), 3, 64, 64)
batch_size = 32
class AutoEncoder(nn.Module):
def __init__(self):
super(AutoEncoder, self).__init__()
self.encoder = nn.Sequential(
nn.Conv2d(3, 120, kernel_size=3, stride=2, padding=1),
nn.LeakyReLU(True),
nn.Conv2d(120, 160, kernel_size=3, stride=2, padding=1),
nn.BatchNorm2d(160),
nn.LeakyReLU(True),
nn.Conv2d(160, 200, kernel_size=3, stride=2, padding=0),
nn.LeakyReLU(True),
nn.Flatten()
)
self.fc1 = nn.Sequential(
nn.Linear(9800, 600),
nn.LeakyReLU(True)
)
self.fc2 = nn.Sequential(
nn.Linear(300, 600),
nn.LeakyReLU(True),
nn.Linear(600, 9800),
nn.LeakyReLU(True)
)
self.decoder = nn.Sequential(
nn.ConvTranspose2d(200, 160, kernel_size=3, stride=2, output_padding=0),
nn.BatchNorm2d(160),
nn.ReLU(True),
nn.ConvTranspose2d(160, 120, kernel_size=5, stride=2, padding=1, output_padding=1),
nn.BatchNorm2d(120),
nn.ReLU(True),
nn.ConvTranspose2d(120, 3, kernel_size=3, stride=2, padding=1, output_padding=1),
nn.Sigmoid()
)
def reparameterize(self, mu, log_var):
"""
:param mu: mean from the encoder's latent space
:param log_var: log variance from the encoder's latent space
"""
std = torch.exp(0.5*log_var) # standard deviation
eps = torch.randn_like(std) # `randn_like` as we need the same size
sample = mu + (eps * std) # sampling as if coming from the input space
return sample
def forward(self, x):
#code from https://debuggercafe.com/getting-started-with-variational-autoencoder-using-pytorch/
x = self.encode(x).view(-1, 2, 300)
# get `mu` and `log_var`
mu = x[:, 0, :] # the first feature values as mean
log_var = x[:, 1, :] # the other feature values as variance
# get the latent vector through reparameterization
z = self.reparameterize(mu, log_var)
x = self.decode(z)
return x, mu, log_var
def encode(self, x):
x = self.encoder(x)
x = self.fc1(x)
return x
def decode(self, x):
x = self.fc2(x)
x = x.view(x.shape[0], 200, 7, 7)
x = self.decoder(x)
return x
#code from https://debuggercafe.com/getting-started-with-variational-autoencoder-using-pytorch/
def final_loss(bce_loss, mu, logvar):
"""
This function will add the reconstruction loss (BCELoss) and the
KL-Divergence.
KL-Divergence = 0.5 * sum(1 + log(sigma^2) - mu^2 - sigma^2)
:param bce_loss: recontruction loss
:param mu: the mean from the latent vector
:param logvar: log variance from the latent vector
"""
BCE = bce_loss
KLD = -0.5 * torch.sum(1 + logvar - mu.pow(2) - logvar.exp())
return BCE + KLD
model = AutoEncoder()
model.load_state_dict(torch.load("face_model.pkl"))
optimizer = Adam(model.parameters(), lr=3e-4)
loss_function = nn.BCELoss(reduction="sum")
if torch.cuda.is_available():
model = model.cuda()
loss_function = loss_function.cuda()
model = AutoEncoder()
optimizer = Adam(model.parameters(), lr=3e-4)
loss_function = nn.BCELoss(reduction="sum")
if torch.cuda.is_available():
model = model.cuda()
loss_function = loss_function.cuda()
model.train()
epoch = 10_000
for e in tqdm(range(epoch)):
running_loss = 0.0
for i in range(1, int(len(dataset) / batch_size)):
data = dataset[(i-1)*batch_size:i*batch_size]
optimizer.zero_grad()
reconstruction, mu, logvar = model(data)
bce_loss = loss_function(reconstruction, data)
loss = final_loss(bce_loss, mu, logvar)
running_loss += loss.item()
loss.backward()
optimizer.step()
train_loss = running_loss/len(dataset)
print(f"Epoch: {e}\nLoss: {train_loss}")
torch.save(model.state_dict(), "drive/MyDrive/models/face_model.pkl")
print(len(dataset))
import matplotlib.pyplot as plt
model.eval()
plt.imshow(dataset[0].cpu().reshape(64, 64, 3))
reconstruction, mu, logvar = model(dataset[0].view(1, 3, 64, 64))
plt.imshow(reconstruction.cpu().detach().reshape(64, 64, 3))
```
| github_jupyter |
# Training Neural Networks
The network we built in the previous part isn't so smart, it doesn't know anything about our handwritten digits. Neural networks with non-linear activations work like universal function approximators. There is some function that maps your input to the output. For example, images of handwritten digits to class probabilities. The power of neural networks is that we can train them to approximate this function, and basically any function given enough data and compute time.
<img src="assets/function_approx.png" width=500px>
At first the network is naive, it doesn't know the function mapping the inputs to the outputs. We train the network by showing it examples of real data, then adjusting the network parameters such that it approximates this function.
To find these parameters, we need to know how poorly the network is predicting the real outputs. For this we calculate a **loss function** (also called the cost), a measure of our prediction error. For example, the mean squared loss is often used in regression and binary classification problems
$$
\large \ell = \frac{1}{2n}\sum_i^n{\left(y_i - \hat{y}_i\right)^2}
$$
where $n$ is the number of training examples, $y_i$ are the true labels, and $\hat{y}_i$ are the predicted labels.
By minimizing this loss with respect to the network parameters, we can find configurations where the loss is at a minimum and the network is able to predict the correct labels with high accuracy. We find this minimum using a process called **gradient descent**. The gradient is the slope of the loss function and points in the direction of fastest change. To get to the minimum in the least amount of time, we then want to follow the gradient (downwards). You can think of this like descending a mountain by following the steepest slope to the base.
<img src='assets/gradient_descent.png' width=350px>
## Backpropagation
For single layer networks, gradient descent is straightforward to implement. However, it's more complicated for deeper, multilayer neural networks like the one we've built. Complicated enough that it took about 30 years before researchers figured out how to train multilayer networks.
Training multilayer networks is done through **backpropagation** which is really just an application of the chain rule from calculus. It's easiest to understand if we convert a two layer network into a graph representation.
<img src='assets/backprop_diagram.png' width=550px>
In the forward pass through the network, our data and operations go from bottom to top here. We pass the input $x$ through a linear transformation $L_1$ with weights $W_1$ and biases $b_1$. The output then goes through the sigmoid operation $S$ and another linear transformation $L_2$. Finally we calculate the loss $\ell$. We use the loss as a measure of how bad the network's predictions are. The goal then is to adjust the weights and biases to minimize the loss.
To train the weights with gradient descent, we propagate the gradient of the loss backwards through the network. Each operation has some gradient between the inputs and outputs. As we send the gradients backwards, we multiply the incoming gradient with the gradient for the operation. Mathematically, this is really just calculating the gradient of the loss with respect to the weights using the chain rule.
$$
\large \frac{\partial \ell}{\partial W_1} = \frac{\partial L_1}{\partial W_1} \frac{\partial S}{\partial L_1} \frac{\partial L_2}{\partial S} \frac{\partial \ell}{\partial L_2}
$$
**Note:** I'm glossing over a few details here that require some knowledge of vector calculus, but they aren't necessary to understand what's going on.
We update our weights using this gradient with some learning rate $\alpha$.
$$
\large W^\prime_1 = W_1 - \alpha \frac{\partial \ell}{\partial W_1}
$$
The learning rate $\alpha$ is set such that the weight update steps are small enough that the iterative method settles in a minimum.
## Losses in PyTorch
Let's start by seeing how we calculate the loss with PyTorch. Through the `nn` module, PyTorch provides losses such as the cross-entropy loss (`nn.CrossEntropyLoss`). You'll usually see the loss assigned to `criterion`. As noted in the last part, with a classification problem such as MNIST, we're using the softmax function to predict class probabilities. With a softmax output, you want to use cross-entropy as the loss. To actually calculate the loss, you first define the criterion then pass in the output of your network and the correct labels.
Something really important to note here. Looking at [the documentation for `nn.CrossEntropyLoss`](https://pytorch.org/docs/stable/nn.html#torch.nn.CrossEntropyLoss),
> This criterion combines `nn.LogSoftmax()` and `nn.NLLLoss()` in one single class.
>
> The input is expected to contain scores for each class.
This means we need to pass in the raw output of our network into the loss, not the output of the softmax function. This raw output is usually called the *logits* or *scores*. We use the logits because softmax gives you probabilities which will often be very close to zero or one but floating-point numbers can't accurately represent values near zero or one ([read more here](https://docs.python.org/3/tutorial/floatingpoint.html)). It's usually best to avoid doing calculations with probabilities, typically we use log-probabilities.
```
# The MNIST datasets are hosted on yann.lecun.com that has moved under CloudFlare protection
# Run this script to enable the datasets download
# Reference: https://github.com/pytorch/vision/issues/1938
from six.moves import urllib
opener = urllib.request.build_opener()
opener.addheaders = [('User-agent', 'Mozilla/5.0')]
urllib.request.install_opener(opener)
import torch
from torch import nn
import torch.nn.functional as F
from torchvision import datasets, transforms
# Define a transform to normalize the data
transform = transforms.Compose([transforms.ToTensor(),
transforms.Normalize((0.5,), (0.5,)),
])
# Download and load the training data
trainset = datasets.MNIST('~/.pytorch/MNIST_data/', download=True, train=True, transform=transform)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=64, shuffle=True)
# Build a feed-forward network
model = nn.Sequential(nn.Linear(784, 128),
nn.ReLU(),
nn.Linear(128, 64),
nn.ReLU(),
nn.Linear(64, 10))
# Define the loss
criterion = nn.CrossEntropyLoss()
# Get our data
images, labels = next(iter(trainloader))
# Flatten images
images = images.view(images.shape[0], -1)
# Forward pass, get our logits
logits = model(images)
# Calculate the loss with the logits and the labels
loss = criterion(logits, labels)
print(loss)
```
In my experience it's more convenient to build the model with a log-softmax output using `nn.LogSoftmax` or `F.log_softmax` ([documentation](https://pytorch.org/docs/stable/nn.html#torch.nn.LogSoftmax)). Then you can get the actual probabilites by taking the exponential `torch.exp(output)`. With a log-softmax output, you want to use the negative log likelihood loss, `nn.NLLLoss` ([documentation](https://pytorch.org/docs/stable/nn.html#torch.nn.NLLLoss)).
>**Exercise:** Build a model that returns the log-softmax as the output and calculate the loss using the negative log likelihood loss.
```
## Solution
# Build a feed-forward network
model = nn.Sequential(nn.Linear(784, 128),
nn.ReLU(),
nn.Linear(128, 64),
nn.ReLU(),
nn.Linear(64, 10),
nn.LogSoftmax(dim=1))
# Define the loss
criterion = nn.NLLLoss()
# Get our data
images, labels = next(iter(trainloader))
# Flatten images
images = images.view(images.shape[0], -1)
# Forward pass, get our log-probabilities
logps = model(images)
# Calculate the loss with the logps and the labels
loss = criterion(logps, labels)
print(loss)
```
## Autograd
Now that we know how to calculate a loss, how do we use it to perform backpropagation? Torch provides a module, `autograd`, for automatically calculating the gradients of tensors. We can use it to calculate the gradients of all our parameters with respect to the loss. Autograd works by keeping track of operations performed on tensors, then going backwards through those operations, calculating gradients along the way. To make sure PyTorch keeps track of operations on a tensor and calculates the gradients, you need to set `requires_grad = True` on a tensor. You can do this at creation with the `requires_grad` keyword, or at any time with `x.requires_grad_(True)`.
You can turn off gradients for a block of code with the `torch.no_grad()` content:
```python
x = torch.zeros(1, requires_grad=True)
>>> with torch.no_grad():
... y = x * 2
>>> y.requires_grad
False
```
Also, you can turn on or off gradients altogether with `torch.set_grad_enabled(True|False)`.
The gradients are computed with respect to some variable `z` with `z.backward()`. This does a backward pass through the operations that created `z`.
```
x = torch.randn(2,2, requires_grad=True)
print(x)
y = x**2
print(y)
```
Below we can see the operation that created `y`, a power operation `PowBackward0`.
```
## grad_fn shows the function that generated this variable
print(y.grad_fn)
```
The autograd module keeps track of these operations and knows how to calculate the gradient for each one. In this way, it's able to calculate the gradients for a chain of operations, with respect to any one tensor. Let's reduce the tensor `y` to a scalar value, the mean.
```
z = y.mean()
print(z)
```
You can check the gradients for `x` and `y` but they are empty currently.
```
print(x.grad)
```
To calculate the gradients, you need to run the `.backward` method on a Variable, `z` for example. This will calculate the gradient for `z` with respect to `x`
$$
\frac{\partial z}{\partial x} = \frac{\partial}{\partial x}\left[\frac{1}{n}\sum_i^n x_i^2\right] = \frac{x}{2}
$$
```
z.backward()
print(x.grad)
print(x/2)
```
These gradients calculations are particularly useful for neural networks. For training we need the gradients of the weights with respect to the cost. With PyTorch, we run data forward through the network to calculate the loss, then, go backwards to calculate the gradients with respect to the loss. Once we have the gradients we can make a gradient descent step.
## Loss and Autograd together
When we create a network with PyTorch, all of the parameters are initialized with `requires_grad = True`. This means that when we calculate the loss and call `loss.backward()`, the gradients for the parameters are calculated. These gradients are used to update the weights with gradient descent. Below you can see an example of calculating the gradients using a backwards pass.
```
# Build a feed-forward network
model = nn.Sequential(nn.Linear(784, 128),
nn.ReLU(),
nn.Linear(128, 64),
nn.ReLU(),
nn.Linear(64, 10),
nn.LogSoftmax(dim=1))
criterion = nn.NLLLoss()
images, labels = next(iter(trainloader))
images = images.view(images.shape[0], -1)
logps = model(images)
loss = criterion(logps, labels)
print('Before backward pass: \n', model[0].weight.grad)
loss.backward()
print('After backward pass: \n', model[0].weight.grad)
```
## Training the network!
There's one last piece we need to start training, an optimizer that we'll use to update the weights with the gradients. We get these from PyTorch's [`optim` package](https://pytorch.org/docs/stable/optim.html). For example we can use stochastic gradient descent with `optim.SGD`. You can see how to define an optimizer below.
```
from torch import optim
# Optimizers require the parameters to optimize and a learning rate
optimizer = optim.SGD(model.parameters(), lr=0.01)
```
Now we know how to use all the individual parts so it's time to see how they work together. Let's consider just one learning step before looping through all the data. The general process with PyTorch:
* Make a forward pass through the network
* Use the network output to calculate the loss
* Perform a backward pass through the network with `loss.backward()` to calculate the gradients
* Take a step with the optimizer to update the weights
Below I'll go through one training step and print out the weights and gradients so you can see how it changes. Note that I have a line of code `optimizer.zero_grad()`. When you do multiple backwards passes with the same parameters, the gradients are accumulated. This means that you need to zero the gradients on each training pass or you'll retain gradients from previous training batches.
```
print('Initial weights - ', model[0].weight)
images, labels = next(iter(trainloader))
images.resize_(64, 784)
# Clear the gradients, do this because gradients are accumulated
optimizer.zero_grad()
# Forward pass, then backward pass, then update weights
output = model(images)
loss = criterion(output, labels)
loss.backward()
print('Gradient -', model[0].weight.grad)
# Take an update step and few the new weights
optimizer.step()
print('Updated weights - ', model[0].weight)
```
### Training for real
Now we'll put this algorithm into a loop so we can go through all the images. Some nomenclature, one pass through the entire dataset is called an *epoch*. So here we're going to loop through `trainloader` to get our training batches. For each batch, we'll doing a training pass where we calculate the loss, do a backwards pass, and update the weights.
> **Exercise: ** Implement the training pass for our network. If you implemented it correctly, you should see the training loss drop with each epoch.
```
model = nn.Sequential(nn.Linear(784, 128),
nn.ReLU(),
nn.Linear(128, 64),
nn.ReLU(),
nn.Linear(64, 10),
nn.LogSoftmax(dim=1))
criterion = nn.NLLLoss()
optimizer = optim.SGD(model.parameters(), lr=0.003)
epochs = 5
for e in range(epochs):
running_loss = 0
for images, labels in trainloader:
# Flatten MNIST images into a 784 long vector
images = images.view(images.shape[0], -1)
# TODO: Training pass
optimizer.zero_grad()
output = model(images)
loss = criterion(output, labels)
loss.backward()
optimizer.step()
running_loss += loss.item()
else:
print(f"Training loss: {running_loss/len(trainloader)}")
```
With the network trained, we can check out it's predictions.
```
%matplotlib inline
import helper
images, labels = next(iter(trainloader))
img = images[0].view(1, 784)
# Turn off gradients to speed up this part
with torch.no_grad():
logps = model(img)
# Output of the network are log-probabilities, need to take exponential for probabilities
ps = torch.exp(logps)
helper.view_classify(img.view(1, 28, 28), ps)
```
Now our network is brilliant. It can accurately predict the digits in our images. Next up you'll write the code for training a neural network on a more complex dataset.
| github_jupyter |
```
!pip install keras
!pip install lightgbm
import pandas as pd
import numpy as np
train=pd.read_csv("train_all.csv")
test=pd.read_csv("test_all.csv")
all=pd.concat([train,test],ignore_index=True)
item=all["user_items"].values.tolist()
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.feature_extraction.text import TfidfTransformer
docs = item
#该类会将文本中的词语转换为词频矩阵,矩阵元素a[i][j] 表示j词在i类文本下的词频
vectorizer = CountVectorizer(max_features=15)
#该类会统计每个词语的tf-idf权值
tf_idf_transformer = TfidfTransformer()
#将文本转为词频矩阵并计算tf-idf
tf_idf = tf_idf_transformer.fit_transform(vectorizer.fit_transform(docs))
#将tf-idf矩阵抽取出来,元素a[i][j]表示j词在i类文本中的tf-idf权重
docs_weight = tf_idf.toarray()
train_list=docs_weight[0:900000]
test_list=docs_weight[900000:1900000]
# a=pd.DataFrame(embedding_list)
# train.drop(["user_items"],axis=1,inplace=True)
# train_em=pd.concat([train,a],axis=1)
# train_em.to_csv("train_em.csv")
import lightgbm as lgb
from sklearn import datasets
from sklearn.model_selection import train_test_split
target=np.array(train["age"])#年龄建模
X_train,X_test,y_train,y_test=train_test_split(train_list,target,test_size=0.03)
import numpy as np
train_data=lgb.Dataset(X_train,label=y_train)
validation_data=lgb.Dataset(X_test,label=y_test)
params={
'learning_rate':0.1,
'lambda_l1':0.1,
'lambda_l2':0.2,
'max_depth':4,
'objective':'multiclass',
'num_class':13, #lightgbm.basic.LightGBMError: b‘Number of classes should be specified and greater than 1 for multiclass training‘
}
clf=lgb.train(params,train_data,valid_sets=[validation_data])
clf.save_model('age_model.txt')
from sklearn.metrics import roc_auc_score,accuracy_score
y_pred=clf.predict(X_test)
y_pred=[list(x).index(max(x)) for x in y_pred]
# print(y_pred)
print(accuracy_score(y_test,y_pred))
import lightgbm as lgb
from sklearn import datasets
from sklearn.model_selection import train_test_split
target=np.array(train["gender"])#性别建模
X_train,X_test,y_train,y_test=train_test_split(train_list,target,test_size=0.04)
import numpy as np
train_data=lgb.Dataset(X_train,label=y_train)
validation_data=lgb.Dataset(X_test,label=y_test)
params={
'learning_rate':0.1,
'lambda_l1':0.1,
'lambda_l2':0.2,
'max_depth':4,
'objective':'multiclass',
'num_class':4, #lightgbm.basic.LightGBMError: b‘Number of classes should be specified and greater than 1 for multiclass training‘
}
clf=lgb.train(params,train_data,valid_sets=[validation_data])
clf.save_model('gender_model.txt')
from sklearn.metrics import roc_auc_score,accuracy_score
y_pred=clf.predict(X_test)
y_pred=[list(x).index(max(x)) for x in y_pred]
# print(y_pred)
print(accuracy_score(y_test,y_pred))
import lightgbm as lgb
# 模型加载
gbm = lgb.Booster(model_file='age_model.txt')
# 模型预测
age_pred = gbm.predict(test_list)
age_pred=[list(x).index(max(x)) for x in age_pred]
import lightgbm as lgb
# 模型加载
gbm = lgb.Booster(model_file='gender_model.txt')
# 模型预测
gender_pred = gbm.predict(test_list, num_iteration=gbm.best_iteration)
gender_pred=[list(x).index(max(x)) for x in gender_pred]
submission=pd.DataFrame(test["user_id"])
submission["predicted_age"]=age_pred
submission["predicted_gender"]=gender_pred
submission.to_csv("submission_tf-idf.csv",index=False)
submission
```
| github_jupyter |
```
import pandas as pd
from sklearn.metrics import classification_report
!ls
train = pd.read_csv('../Post Processing/data/postproc_train.csv')
val = pd.read_csv('../Post Processing/data/postproc_val.csv')
test = pd.read_csv('../Post Processing/data/postproc_test.csv')
test_gt = pd.read_csv('../../data/english_test_with_labels.csv')
val_gt = pd.read_csv('../../data/Constraint_Val.csv')
def post_proc(row):
if (row['domain_real']>row['domain_fake']):
return 0
elif (row['domain_real']<row['domain_fake']):
return 1
else:
# if (row['username_real']>row['username_fake']) & (row['username_real']>0.88):
# return 0
# elif (row['username_real']<row['username_fake']) & (row['username_fake']>0.88):
# return 1
# else:
if row['class1_pred']>row['class0_pred']:
return 1
elif row['class1_pred']<row['class0_pred']:
return 0
def post_proc1(row):
if row['class1_pred']>row['class0_pred']:
return 1
elif row['class1_pred']<row['class0_pred']:
return 0
train['final_pred'] = train.apply(lambda x: post_proc(x), 1)
print(classification_report(train['label'], train['final_pred']))
val['final_pred'] = val.apply(lambda x: post_proc(x), 1)
print(classification_report(val['label'], val['final_pred']))
from sklearn.metrics import f1_score,accuracy_score,precision_score,recall_score
print('f1_score : ',f1_score(val['label'], val['final_pred'],average='micro'))
print('precision_score : ',precision_score(val['label'], val['final_pred'],average='micro'))
print('recall_score : ',recall_score(val['label'], val['final_pred'],average='micro'))
test['final_pred'] = test.apply(lambda x: post_proc(x), 1)
print(classification_report(test['label'], test['final_pred']))
from sklearn.metrics import f1_score,accuracy_score,precision_score,recall_score
print('f1_score : ',f1_score(test['label'], test['final_pred'],average='micro'))
print('precision_score : ',precision_score(test['label'], test['final_pred'],average='micro'))
print('recall_score : ',recall_score(test['label'], test['final_pred'],average='micro'))
```
## Get False Pred samples
```
val_false_pred = val[val.final_pred!=val.label]
pd.merge(val_false_pred, val_gt, left_index=True, right_index=True)
pd.merge(val_false_pred, val_gt, left_index=True, right_index=True).to_csv('../Post Processing/results/val_false_pred_var_1_1.csv')
test_false_pred = test[test.final_pred!=test.label]
pd.merge(test_false_pred, test_gt, left_index=True, right_index=True)
pd.merge(test_false_pred, test_gt, left_index=True, right_index=True).to_csv('../Post Processing/results/test_false_pred_var_1_1.csv')
```
| github_jupyter |
# Score AI: NLP Essay Scorer
## Why Score AI
Teachers remain stretched thin and under-supported. Automating some of the work they do will help them focus their time and energy on the most important aspects of their work -- building relationships with students that allow them to draw out their best selves. Scoring essays is one of the most time-intensive tasks that a teacher has to do, and the delay in feedback makes it harder for the student to use and internalize that feedback.
## Challenges
- There is a lack of labelled essay data
- There are no straightforward scoring metrics for essay organization
- There are vastly different writing styles and essay lengths
- Good writing transcends formulaic approaches
## Responses
- Scraped data
- Used background knowledge about essay grading to develop a scoring metric
- Chose to only score argumentative essays to start and made scoring length-independent
- Suggest use of scorer only for non-expert writers
## Scraping
Of the dozen or so essay sites that I investigated for scraping, only 2 of them freely offered the entire text of essays without a login. I used a scrapy script within the terminal to run spiders over those two sites to collect essays on about 40 different search terms for popular essay topics and save the necessary information in json files. The scrapy scripts are located in the same Github repo as this project.
In order to get a ground truth for topic modelling, I used the spiders to scrape tags from the essay pages. Unfortunately, many of the essay tags turned out to be blank or meaningless.
```
!pip install pyLDAvis
import pandas as pd
import spacy
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sns
import csv
import sklearn
import re
import nltk
import pickle
import gensim
import gensim.corpora as corpora
import pyLDAvis
import pyLDAvis.gensim
from sklearn.preprocessing import StandardScaler, MinMaxScaler
from gensim.models import CoherenceModel, LsiModel, LdaModel
from string import punctuation
from itertools import starmap, combinations
%matplotlib inline
!python -m spacy download en_core_web_md
# Read in essay data from both sites, concatenate into one data frame
sm_df = pd.read_json('studymoose.json', orient='records')
phd_df = pd.read_json('phdessay.json', orient='records')
raw_df = pd.concat([sm_df, phd_df], ignore_index=True)
len(raw_df)
```
## Data Cleaning and Preprocessing
The essay texts are not in good shape to be topic modeled and tokenized. Some paragraphs are just newlines, some sentences are too short to be meaningful. Some sentences are citations/references. Essays are too long or contain too many one-sentence paragraphs. I will be removing these all from the dataset because they are interspersed randomly through the data and will not too greatly affect the makeup of the corpus.
```
# Find text that only newline characters
error_df = raw_df[raw_df['Text'].str.contains('\\n', regex=True)]
error_df = error_df[error_df['Text'].str.len() < 50]
drop_indices = error_df.index.tolist()
# Drop newlines
raw_df.drop(drop_indices, axis=0, inplace=True)
raw_df.reset_index(drop=True, inplace=True)
raw_df.tail()
# Split each paragraph into sentences
sentences = raw_df.Text.str.split(r'[.!?] ', expand=True)
# Create a sentence-level data frame by joining the sentences to the raw_df
sent_df = raw_df.drop('Text', axis=1).join(
sentences.stack().reset_index(drop=True, level=1).rename('Text'))
sent_df.head()
# Remove lines that are too short
short_df = sent_df[sent_df['Text'].str.len() < 15]
short_indices = short_df.index.tolist()
sent_df.drop(short_indices, axis=0, inplace=True)
# Reset index
sent_df.reset_index(drop=True, inplace=True)
sent_df.tail()
# Remove lines that have too much punctuation (90% citations)
ref_indices = []
for i in range(len(sent_df)):
text = sent_df.iloc[i, 4]
punct = [c for c in text if c in punctuation]
ratio = len(punct) / len(text)
if ratio > 0.1:
ref_indices.append(i)
sent_df.drop(ref_indices, axis=0, inplace=True)
len(sent_df)
# Count the sentences
sent_df['Sentences'] = sent_df.groupby(
['URL', 'Paragraph'])['Text'].transform('count')
sent_df.head()
# Remove essays that are too long
long_essays = sent_df[sent_df['Paragraph']>=66]['URL'].unique().tolist()
sent_df = sent_df[~sent_df['URL'].isin(long_essays)]
len(sent_df)
# Number each essay
essays = sent_df.URL.unique()
essay_dict = dict(zip(essays, range(len(essays))))
# Create an Essay column with the number of each essay
sent_df['Essay'] = sent_df['URL'].replace(essay_dict)
sent_df.reset_index(drop= True, inplace=True)
sent_df.tail()
# Find essays that have too many one sentence paragraphs
choppy_essays = []
for i, grp in sent_df.groupby('Essay'):
short_paras = grp[grp['Sentences']==1]
paras = grp['Paragraph'].unique().tolist()
if len(short_paras) / len(paras) > 0.5:
choppy_essays.append(i)
len(choppy_essays)
# Remove essays that have too many one sentence paragraphs
choppy_indices = sent_df[sent_df['Essay'].isin(choppy_essays)].index.tolist()
sent_df.drop(choppy_indices, inplace=True)
len(sent_df)
# Remove remaining one sentence paragraphs
one_liners = sent_df[sent_df['Sentences']==1].index.tolist()
sent_df.drop(one_liners, inplace=True)
len(sent_df)
# Number each essay
essays = sent_df.URL.unique()
essay_dict = dict(zip(essays, range(len(essays))))
# Create an Essay column with the number of each essay
sent_df['Essay'] = sent_df['URL'].replace(essay_dict)
sent_df.reset_index(drop= True, inplace=True)
sent_df.tail()
# Find the paragraphs per essay and sentences per paragraph
maxima = sent_df.groupby('Essay')['Paragraph'].max()
sents_per_para = sent_df.Sentences.value_counts().to_dict()
sents_per_para = {k: int(v/k) for k, v in sents_per_para.items()}
s_counts = [v for v in sents_per_para.values()]
```
## Distributions of Essay Length
From the plots above, it is easy to see that there is no standard size for essays in terms of the number of paragraphs in the essay or the number of sentences in a paragraph. After exploring the distribution of paragraphs that are one sentence long and the essays that are highly composed of one sentence paragraphs, I decided to remove them from the data set. I believe I can do so without significantly impacting the performance of my model.
Removal of one sentence paragraphs and essays with a high percentage of on e sentence paragraphs in this model stems from the data collection process. I believe many of the one sentence paragraph to be a result of delineating paragraphs based on the html paragraph tag. In future iterations of this model where essay are collected from users, one sentence paragraphs will be scored at a z-score of -3.
```
# Visualize distributions of paragraphs and sentences per para
sns.set_style('darkgrid')
plt.figure(figsize=(15, 5))
plt.subplot(121)
sns.distplot(maxima,
kde=False,
norm_hist=False)
plt.title('Distribution of Paragraphs per Essay')
plt.xlabel('Number of paragraphs')
plt.subplot(122)
sns.barplot([k for k in sents_per_para.keys()],
s_counts,
color='green')
plt.title('Distribution of Sentences per Paragraph')
plt.xlabel('Sentences per paragraph')
plt.show()
print('Paragraphs per Essay:\nMean -- {}\nMedian -- {}\n\n'.format(
maxima.mean(), maxima.median()))
s_list = []
for i in range (1, 33):
if i in sents_per_para.keys():
for j in range(sents_per_para[i]):
s_list.append(i)
s_mean = np.mean(s_list)
mid = len(s_list)//2
s_median = (s_list[mid] + s_list[mid-1]) / 2
print('Sentences per Para:\nMean -- {}\nMedian -- {}\n\n'.format(
s_mean, s_median))
```
## Tokenizing and Topic Modeling
### Spacy Tokens
Since the model depends on the cosine similarity of sentence vectors, I needed to use one of the larger spacy tokenizers. I needed part of speech tags to get only semantically substantive words for topic modeling, but I could disable the dependency parser and named entity recognizer to cut down on memory usage and processing time.
### Topic Modeling
I used the gensim package for topic modeling predominantly for its CoherenceModel implementation that would help me visualize how well the topic models describe the corpus.
```
# Load spacy model for tokenizing
try:
with open('essays.pickle', 'rb') as fp:
tokenized_sents = pickle.load(fp)
print('Pretrained Model loaded!')
except:
nlp = spacy.load(
'en_core_web_md',
disable=['parser', 'ner'])
# Run NLP to get tokens
tokenized_sents = []
i = 0
for sent in sent_df.Text.tolist():
i += 1
if i % 5000 == 0:
print('Processing {}'.format(i))
doc = nlp(sent)
# Get lemmas for each word
tokenized_sents.append(doc)
with open('essays.pickle', 'wb') as fp:
pickle.dump(tokenized_sents, fp)
print('New model trained!')
# Set sent_df Text column to tokenized sentences
sent_df['Text'] = tokenized_sents
sent_df.head()
# Group texts by essay to send essays through as documents for the corpus
essays = sent_df.groupby('Essay')['Text'].apply(lambda x: list(x))
# Extract only substantive words for topic modeling
essay_lemmas = []
for essay in essays:
essay_lemmas.append([x.lemma_ for e in essay for x in e
if not x.is_punct and
x.pos_ in ['PRON', 'NOUN', 'VERB', 'ADJ'] and
x.lemma_ not in ['be', 'have', 'do']])
# Create Dictionary
id2word = corpora.Dictionary([essay_lemmas[0]])
# Add the rest of the essays to the dictionary
for i in range(1, len(essays)):
try:
id2word.add_documents([essay_lemmas[i]])
except:
print('Error: {}'.format(i))
if i % 500 == 0:
print('Processed line {}'.format(i))
# Create Corpus
texts = []
for essay in essay_lemmas:
texts.append([e for e in essay])
# Term Document Frequency
corpus = []
for i in range(len(texts)):
try:
corpus.append(id2word.doc2bow(texts[i]))
except:
print('Error: {}'.format(i))
# View
print(corpus[:1])
def get_lda_model(num_topics):
'''Trains an LDA model and outputs model and list of information
to be used for model info data frame that will be useful for
selecting best topic model'''
model_type = 'LDA'
model_name = 'checkpoints/lda' + str(num_topics) + '.model'
try:
return LdaModel.load(model_name), [model_type, num_topics]
except:
print('Training new LDA {} Model'.format(num_topics))
# Build LDA model
lda_model = LdaModel(corpus=corpus,
id2word=id2word,
num_topics=num_topics,
update_every=1,
chunksize=100,
passes=10,
alpha='auto',
per_word_topics=True)
# Save model
lda_model.save(model_name)
return lda_model, [model_type, num_topics]
# Get LDA Models
lda12_model, lda12_info = get_lda_model(num_topics=12)
lda16_model, lda16_info = get_lda_model(num_topics=16)
lda20_model, lda20_info = get_lda_model(num_topics=20)
lda22_model, lda22_info = get_lda_model(num_topics=22)
lda24_model, lda24_info = get_lda_model(num_topics=24)
lda26_model, lda26_info = get_lda_model(num_topics=26)
lda28_model, lda28_info = get_lda_model(num_topics=28)
def get_lsa_model(num_topics):
'''Trains an LSA model and outputs model and list of information
to be used for model info data frame that will be useful for
selecting best topic model'''
model_type = 'LSA'
model_name = 'checkpoints/lsa' + str(num_topics) + '.model'
try:
return LsiModel.load(model_name), [model_type, num_topics]
except:
print('Training new LSA {} Model'.format(num_topics))
# Build LSA Model
lsa_model = LsiModel(corpus=corpus,
id2word=id2word,
num_topics=num_topics,
chunksize=100)
lsa_model.save(model_name)
return lsa_model, [model_type, num_topics]
# Get LSA Models
lsa12_model, lsa12_info = get_lsa_model(num_topics=12)
lsa16_model, lsa16_info = get_lsa_model(num_topics=16)
lsa20_model, lsa20_info = get_lsa_model(num_topics=20)
lsa22_model, lsa22_info = get_lsa_model(num_topics=22)
lsa24_model, lsa24_info = get_lsa_model(num_topics=24)
lsa26_model, lsa26_info = get_lsa_model(num_topics=26)
lsa28_model, lsa28_info = get_lsa_model(num_topics=28)
def get_model_info(df, topic_model, model_type, n_topics, corpus):
'''Add LDA and LSA model information to a data frame
in order to compare, plot, and select the best model'''
print('Getting model info for {} {}'.format(model_type, n_topics))
# Add model info to dataframe
model = model_type
n = n_topics
topics = topic_model.print_topics(num_topics=n_topics)
if model == 'LDA':
perplexity = topic_model.log_perplexity(corpus)
else:
perplexity = 'N/A'
coherence_model = CoherenceModel(model=topic_model,
texts=texts,
dictionary=id2word,
coherence='c_v')
coherence = coherence_model.get_coherence()
return pd.concat([df,
pd.DataFrame.from_dict({'Model Type': model,
'Num_Topics': n,
'Topics': topics,
'Perplexity': perplexity,
'Coherence': coherence})],
ignore_index=True)
# Retrieve model_info data frame or create a new one
try:
model_info = pd.read_csv('model_info.csv')
except:
models = [[lda12_model, lda12_info],
[lda16_model, lda16_info],
[lda20_model, lda20_info],
[lda22_model, lda22_info],
[lda24_model, lda24_info],
[lda26_model, lda26_info],
[lda28_model, lda28_info],
[lsa12_model, lsa12_info],
[lsa16_model, lsa16_info],
[lsa20_model, lsa20_info],
[lsa22_model, lsa22_info],
[lsa24_model, lsa24_info],
[lsa26_model, lsa26_info],
[lsa28_model, lsa28_info]]
model_info = pd.DataFrame(columns=['Model Type',
'Num_Topics',
'Topics',
'Perplexity',
'Coherence'])
for model in models:
model_info = get_model_info(model_info,
model[0],
model[1][0],
model[1][1],
corpus=corpus)
model_info.to_csv('model_info.csv')
model_info.head()
```
## Comparing Topic Models
LDA will achieve different results with each model because of the random weight initiation, but through 10 different versions of these LDA models, there was always a peak at 16, a relatively similar peak at 22 and an upward trend from 26 to 28 -- all hovering around 0.50 coherence. While the LSA models do not perform as well, they seem to confirm a peak at 16.
Prior to having run any topic models, my mentor and I discussed how many topics we might expect. There were 40 search terms, but many of them could be lumped into similar topics or were even synonyms to try to capture more essays about the same topic. We determined that there were about 16 topics and were surprised to see that intuition confirmed by the topic models. Though other numbers of topics for LDA may produce similar or better coherences, we decided to go with the consensus of two different modeling functions and human expertise to use the LDA16 model going forward.
```
# Compare topic models
plt.figure(figsize=(10, 5))
sns.lineplot(x='Num_Topics',
y='Coherence',
hue='Model Type',
data=model_info)
plt.title('Model Coherence by Number of Topics')
plt.show()
```
## Topics Worth Keeping
Though I tried dropping parts of speech and modal verbs that would not contribute to determining the topic, the model still generated a few topics based on keywords too generic to provide a meaningful topic. For this reason, I will refrain from assigning topics 1, 7, 8, or 13 as the dominant topic for any essay.
```
# Visualize the topics
pyLDAvis.enable_notebook()
vis = pyLDAvis.gensim.prepare(lda16_model, corpus, id2word)
vis
# Code attributed to:
# https://www.machinelearningplus.com/nlp/topic-modeling-gensim-python/
def format_topics_sentences(ldamodel=lda16_model, corpus=corpus, texts=texts):
# Init output
df = pd.DataFrame()
# Get main topic in each document
for i, row in enumerate(ldamodel[corpus[:-1]]):
row = sorted(row[0], key=lambda x: (x[1]), reverse=True)
# Get the Dominant topic,
# Percent Contribution and Keywords for each document
for j, (topic_num, prop_topic) in enumerate(row):
# Prevent dominant topics 7 or 8, they're meaningless
if j in [0, 1, 2, 3, 4] and topic_num not in [1.0, 7.0, 8.0, 13.0]:
wp = ldamodel.show_topic(topic_num)
topic_keywords = ", ".join([word for word, prop in wp])
df = df.append(
pd.Series([int(topic_num),
round(prop_topic,4),
topic_keywords]),
ignore_index=True)
break
else:
continue
df.columns = ['Dominant_Topic',
'Percent_Contrib',
'Topic_Keywords']
# Add original text to the end of the output
contents = pd.Series(texts)
df = pd.concat([df, contents], axis=1)
return(df)
topics_df = format_topics_sentences(ldamodel=lda16_model,
corpus=corpus,
texts=texts)
# Format
topics_df.reset_index(drop= True, inplace=True)
# Set column names
topics_df.columns = ['Dominant_Topic',
'Topic_Percent_Contrib',
'Keywords',
'Text']
# Create Essay number column
topics_df['Essay'] = sent_df.Essay.unique().tolist()
# Show
topics_df.head(10)
pd.options.display.max_rows = 100
# Code attributed to:
# https://www.machinelearningplus.com/nlp/topic-modeling-gensim-python/
# Group top 5 essays under each topic
sorted_topics = pd.DataFrame()
grouped = topics_df.groupby('Dominant_Topic')
for i, grp in grouped:
sorted_topics = pd.concat([sorted_topics,
grp.sort_values(['Topic_Percent_Contrib'],
ascending=[0]).head(5)],
axis=0)
# Reset Index
sorted_topics.reset_index(drop=True, inplace=True)
# Format
sorted_topics.columns = ['Topic_Num',
'Topic_Percent_Contrib',
'Keywords',
'Text',
'Essay']
# Show
sorted_topics
```
## Dominant Topic Distribution
The dominant topics are relatively well distributed once the more generic topics are removed. No one topic dominates the essay and a cursory look confirms the distribution I would expect based on the number of essays I was able to scrape for each topic.
```
# Number of Documents for Each Topic
topic_counts = topics_df.groupby(
'Keywords')['Dominant_Topic'].count()
topic_counts
# Isolate tags for tokenization and topic modeling
tags = sent_df.groupby('Essay')['Tags'].apply(lambda x: np.unique(x)[0])
# Tokenize tags and prepare for topic model
tokenized_tags =[]
nlp = spacy.load('en')
i = 0
for tag in tags:
i += 1
if i % 500 == 0:
print('Processing {}'.format(i))
tag = ' '.join(tag)
doc = nlp(tag)
# Get lemmas for each word
tokenized_tags.append([token.lemma_ for token in doc if not token.is_punct])
tag_corpus = []
for i in range(len(tokenized_tags)):
try:
tag_corpus.append(id2word.doc2bow(tokenized_tags[i]))
except:
print('Error: {}'.format(i))
# Get topics for tags
tags_df = format_topics_sentences(ldamodel=lda16_model,
corpus=tag_corpus,
texts=tokenized_tags)
tags_df.columns = ['Tag_Topic',
'Tag_Percent_Contrib',
'Tag_Keywords',
'Tags']
tags_df.head()
# Add tags and topics to the essay-topic data frame
topics_df = topics_df.join(tags_df)
topics_df.head()
```
## Topic Accuracy
The topic accuracy rose from about 25% to around 38% when the generic topics were pruned. Still, I believe the main thing holding back the accuracy of the topics to be the fact that my labels (the tags scraped from the websites) were not that descriptive of the essay topics to begin with. Given more time, I would find a way to include the search term with the output of the scraper so I could use that as the label instead of the scraped tags. For now, I will show examples of essays and tags where the topic labels match and examples where they don't match to try to illustrate the issue of poor labels over poor topic models.
```
# Determine the percentage of essay topics that match their tag topic
matches = np.where(
topics_df['Dominant_Topic'] == topics_df['Tag_Topic'])[0]
l = len(matches)
print('Tag topic to essay topic accuracy: {}'.format(l/len(topics_df)))
# Separate matches and non-matches
match_df = topics_df.loc[matches]
nonmatch_df = topics_df[~topics_df.index.isin(matches)]
# Show a random sample of matches
match_df.sample(25)
# Show a random sample of non-matches
nonmatch_df.sample(25)
# Read in the paragraph organization scores or generate them
try:
score_df = pd.read_csv('score_df.csv')
except:
def get_similarity(x, y):
# Get cosine similarity between sentence vectors
return x.similarity(y)
# Establish a column for paragraph level scores
sent_df['Para_score'] = 0
for i, grp in sent_df.groupby(['Essay', 'Paragraph']):
idx = grp.index
if i[0] % 500 == 0:
print('Processing {}'.format(i))
# Set the paragraph level score as the mean of the cosine
# similarity scores of all combinations of sentences within
# a paragraph
sent_df.loc[idx, 'Para_score'] = np.mean([x for x in starmap(
get_similarity, combinations(grp['Text'], 2))])
if i[0] % 500 == 0:
print('Para_score {} = {}'.format(idx, sent_df.loc[idx, 'Para_score']))
sent_df.to_csv('score_df.csv')
score_df = sent_df.copy()
score_df.head()
```
## Paragraph Scores Distribution
The scores based on the average cosine similarity of sentence vectors within the paragraph tends to be a relatively normal distribution close to one with a long left tail. For this reason, I left all words in the sentence rather than reducing it to semantically-relevant lemmas as in the topic model. When the sentences are reduced in that way, it shifts the distribution even closer to one and increases the kurtosis.
The negative skew seems to come at least in part from paragraphs containing citations/references, headings, websites, or other text that would not contribute to the body of a paragraph. This issue stems again from the data collection method and these are the pieces of text I was unable to filter out in the earlier cleaning and preprocessing.
## Essay Scores Distribution
The distribution of essay scores is similar to that of paragraph scores while clipping some of the tail on the left. This lets me know that the poor paragraph scores were not all concentrated in a few essays.
```
# Plot the distribution of Paragraph scores
plt.figure(figsize=(10,5))
sns.distplot(score_df['Para_score'])
plt.title('Distribution of paragraph-level organization scores')
plt.show()
print('Paragraph Score Min: {}'.format(score_df['Para_score'].min()))
print('Paragraph Score Max: {}'.format(score_df['Para_score'].max()))
print('Paragraph Score Mean: {}'.format(score_df['Para_score'].mean()))
# Explore paragraphs that comprise the long tail
score_df[score_df['Para_score'] < 0.6]
# Initialize an essay-level organization score column
score_df['Essay_score'] = 0
for i, grp in score_df.groupby('Essay'):
idx = grp.index
if i % 500 == 0:
print('Processing {}'.format(i))
# Set essay score to the mean of paragraph-level organization scores
score_df.loc[idx, 'Essay_score'] = np.mean(
[x for x in grp['Para_score'].unique()])
score_df.head()
# Plot the distribution of Essay scores
plt.figure(figsize=(10,5))
sns.distplot(score_df['Essay_score'])
plt.title('Distribution of essay-level organization scores')
plt.show()
print('Essay Score Min: {}'.format(score_df['Essay_score'].min()))
print('Essay Score Max: {}'.format(score_df['Essay_score'].max()))
print('Essay Score Mean: {}'.format(score_df['Essay_score'].mean()))
# Scale Essay scores based on their z-score
scaler = StandardScaler()
score_df['z_score'] = scaler.fit_transform(
np.array(score_df['Essay_score']).reshape(-1,1))
print(score_df['z_score'].min())
print(score_df['z_score'].max())
# Cut of the tails of the Essay z-scores by assigning them to 3 or -3
score_df['z_score'] = score_df['z_score'].apply(lambda x: x if x <= 3 else 3)
score_df['z_score'] = score_df['z_score'].apply(lambda x: x if x > -3 else -3)
print(score_df['z_score'].min())
print(score_df['z_score'].max())
# Add 3 to all z-scores to get rid of negative values
score_df['z_score'] += 3
print(score_df['z_score'].min())
print(score_df['z_score'].max())
# Add essay topic coherence scores to the data frame
on_topic_dict = dict(zip(
topics_df['Essay'], topics_df['Topic_Percent_Contrib']))
score_df['Topic_score'] = score_df['Essay'].replace(on_topic_dict)
score_df.head()
# Drop the one essay with null values
score_df.dropna(how='any', axis=0, inplace=True)
# Scale essay topic coherence scores between 0 and 1
mmscaler = MinMaxScaler()
score_df['on_topic'] = mmscaler.fit_transform(
np.array(score_df['Topic_score']).reshape(-1,1))
# Get a final organization score by mutliplying the essay org z-score
# by the essay topic coherence score
score_df['Org_score'] = score_df['z_score'] * score_df['on_topic']
score_df.head()
```
## Final Organization Score
The topic model is not perfect, so the distribution of final organization scores (comprised of essay-level org z-scores with the tails clipped and made positive times topic contrib scores scaled between 0 and 1) did not end up on a 0 - 6 scale as I had hoped. However, it seemed to me there was a way to divide the distribution of organization scores to give it the types of grades teachers might give to their students on for essay organization.
- 0 -- This writer cannot demonstrate having learned about organization
-1 -- This writer demonstrates minimal organization -- the essay is divided into paragraphs, some remain on topic
- 2 -- This writer is approaching sufficient organization -- many paragraphs remain on topic
- 3 -- This writer is proficient at organization -- nearly all paragraphs are on topic and contribute to the thesis
- 4 -- This writer is has advanced organization -- all paragraphs remain on topic and contribute to the thesis
As with the topic model accuracy, I will have to pull out examples to qualitatively determine whether these labels are accurate.
```
# Plot the distribution of Organization scores
m = score_df['Org_score'].mean()
maximum = score_df['Org_score'].max()
minimum = score_df['Org_score'].min()
std = score_df['Org_score'].std()
plt.figure(figsize=(15,7))
sns.distplot(score_df['Org_score'])
# Assign teacher interpretable scores -- "Grade on a curve"
plt.axvspan(minimum, (m-(1.5*std)), facecolor='r', alpha=0.4, label='0')
plt.axvspan((m-(1.5*std)), (m-std), facecolor='orange', alpha=0.4, label='1')
plt.axvspan((m-std), (m+std), facecolor='y', alpha=0.4, label='2')
plt.axvspan((m+std), (m+(3*std)), facecolor='g', alpha=0.4, label='3')
plt.axvspan((m+(3*std)), maximum, facecolor='lime', alpha=0.4, label='4')
plt.legend(title='Grades')
plt.title('Distribution of final organization scores')
plt.show()
print('Minimum: {}'.format(minimum))
print('Maximum: {}'.format(maximum))
print('Mean: {}'.format(m))
print('Standard Deviation: {}'.format(std))
# Sample best, average, and worst essays
expert = score_df[score_df['Org_score']>3]
average = score_df[score_df['Org_score']<1.1]
average = average[average['Org_score']>0.8]
worst = score_df[score_df['Org_score']<0.25]
# Show a random sample of matches
ex_sample = expert.sample(25)['URL'].tolist()
av_sample = average.sample(25)['URL'].tolist()
w_sample = worst.sample(25)['URL'].tolist()
print('Expert essays:\n')
for url in ex_sample:
print(url + '\n')
print('\n\nAverage essays:\n')
for url in av_sample:
print(url + '\n')
print('\n\nBad essays:\n')
for url in w_sample:
print(url + '\n')
```
## Conclusions
This is not a fantastic model, but it is a start. Some of the challenges that I faced in getting good results from this model stem from the lack of meaningful labels generated by experts, poor paragraph delineations from the method of data collection, and the inability to remove all citations/references and other non-semantically-relevant text.
For future iterations, I will address these challenges by collecting labeled essays from teachers through an online form, asking that they upload only the text of the essay and using code to try to confirm that, and if possible have them denote the thesis statement as the topic against which to compare on-topic coherence.
| github_jupyter |
```
# @hidden_cell
# The project token is an authorization token that is used to access project resources like data sources, connections, and used by platform APIs.
from project_lib import Project
project = Project(project_id='d25a9164-8f3f-44da-b8ee-e1fe2ad1101e', project_access_token='p-44854a4d967f4b81993b4199ac3553ed54c3c78e')
pc = project.project_context
```
# MARATONA BEHIND THE CODE 2020
## DESAFIO 6 - LIT
<hr>
## Installing Libs
```
!pip install scikit-learn --upgrade
!pip install xgboost --upgrade
!pip install imblearn --upgrade
```
<hr>
## Download dos conjuntos de dados em formato .csv
```
import pandas as pd
!wget --no-check-certificate --content-disposition https://raw.githubusercontent.com/vanderlei-test/dataset-3/master/training_dataset.csv
df_training_dataset = pd.read_csv(r'training_dataset.csv')
df_training_dataset.tail()
```
Sobre o arquivo "training_dataset.csv", temos algumas informações gerais sobre os usuários da plataforma:
**id**
**graduacao**
**universidade**
**profissao**
**organizacao**
**pretende_fazer_cursos_lit**
**interesse_mba_lit**
**importante_ter_certificado**
**horas_semanais_estudo**
**como_conheceu_lit**
**total_modulos**
**modulos_iniciados**
**modulos_finalizados**
**certificados**
**categoria**
```
df_training_dataset.info()
df_training_dataset.nunique()
```
<hr>
## Detalhamento do desafio: classificação multiclasse
Este é um desafio cujo objetivo de negócio é a segmentação dos usuários de uma plataforma de ensino. Para tal, podemos utilizar duas abordagens: aprendizado de máquina supervisionado (classificação) ou não-supervisionado (clustering). Neste desafio será aplicada a classificação, pois é disponível um dataset já com "labels", ou em outras palavras, já com exemplos de dados juntamente com a variável alvo.
Na biblioteca scikit-learn temos diversos algoritmos para classificação. O participante é livre para utilizar o framework que desejar para completar esse desafio.
Neste notebook será mostrado um exeplo de uso do algoritmo "Decision Tree" para classificar parte dos estudantes em seis diferentes perfís.
# Atenção!
A coluna-alvo neste desafio é a coluna ``categoria``
<hr>
## Pre-processando o dataset antes do treinamento
### Removendo todas as linhas que possuem algum valor nulos em determinadas colunas
Usando o método Pandas **DataFrame.dropna()** você pode remover todas as linhas nulas do dataset.
Docs: https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.dropna.html
```
# Exibindo os dados ausentes do conjunto de dados antes da primeira transformação (df)
print("Valores nulos no df_training_dataset antes da transformação DropNA: \n\n{}\n".format(df_training_dataset.isnull().sum(axis = 0)))
# Aplicando a função para deletar todas as linhas com valor NaN na coluna ``certificados'' e ``total_modulos'':
df_training_dataset = df_training_dataset.dropna(axis='index', how='any', subset=['certificados', 'total_modulos'])
# Exibindo os dados ausentes do conjunto de dados após a primeira transformação (df)
print("Valores nulos no df_training_dataset após a transformação DropNA: \n\n{}\n".format(df_training_dataset.isnull().sum(axis = 0)))
```
### Processando valores NaN com o SimpleImputer do sklearn
Para os valores NaN, usaremos a substituição pela constante 0 como **exemplo**.
Você pode escolher a estratégia que achar melhor para tratar os valores nulos :)
Docs: https://scikit-learn.org/stable/modules/generated/sklearn.impute.SimpleImputer.html?highlight=simpleimputer#sklearn.impute.SimpleImputer
```
from sklearn.impute import SimpleImputer
import numpy as np
impute_zeros = SimpleImputer(
missing_values=np.nan,
strategy='constant',
fill_value=0,
verbose=0,
copy=True
)
# Exibindo os dados ausentes do conjunto de dados antes da primeira transformação (df)
print("Valores nulos no df_training_dataset antes da transformação SimpleImputer: \n\n{}\n".format(df_training_dataset.isnull().sum(axis = 0)))
# Aplicando a transformação ``SimpleImputer`` no conjunto de dados base
impute_zeros.fit(X=df_training_dataset)
# Reconstruindo um Pandas DataFrame com os resultados
df_training_dataset_imputed = pd.DataFrame.from_records(
data=impute_zeros.transform(
X=df_training_dataset
),
columns=df_training_dataset.columns
)
# Exibindo os dados ausentes do conjunto de dados após a primeira transformação (df)
print("Valores nulos no df_training_dataset após a transformação SimpleImputer: \n\n{}\n".format(df_training_dataset_imputed.isnull().sum(axis = 0)))
```
### Eliminando colunas indesejadas
Vamos **demonstrar** abaixo como usar o método **DataFrame.drop()**.
Docs: https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.drop.html
```
df_training_dataset_imputed.tail()
df_training_dataset_rmcolumns = df_training_dataset_imputed.drop(columns=['id', 'graduacao', 'universidade', 'organizacao', 'como_conheceu_lit'], inplace=False)
df_training_dataset_rmcolumns.tail()
```
# Atenção!
As colunas removidas acima são apenas para fim de exemplo, você pode usar as colunas que quiser e inclusive criar novas colunas com dados que achar importantes!
### Tratamento de de variáveis categóricas
Como mencionado antes, os computadores não são bons com variáveis "categóricas" (ou strings).
Dado uma coluna com variável categórica, o que podemos realizar é a codificação dessa coluna em múltiplas colunas contendo variáveis binárias. Esse processo é chamado de "one-hot-encoding" ou "dummy encoding". Se você não é familiarizado com esses termos, você pode pesquisar mais sobre isso na internet :)
```
# Tratando variáveis categóricas com o método Pandas ``get_dummies()''
df_training = pd.get_dummies(df_training_dataset_rmcolumns, columns=['profissao'])
df_training.tail()
```
# Atenção!
A coluna **categoria** deve ser mantida como uma string. Você não precisa processar/codificar a variável-alvo.
<hr>
## Treinando um classificador com base em uma árvore de decisão
### Selecionando FEATURES e definindo a variável TARGET
```
df_training.columns
features = df_training[
[
'pretende_fazer_cursos_lit', 'interesse_mba_lit',
'importante_ter_certificado', 'horas_semanais_estudo', 'total_modulos',
'modulos_iniciados', 'modulos_finalizados', 'certificados',
'profissao_0', 'profissao_Advogado', 'profissao_Analista',
'profissao_Analista Senior', 'profissao_Assessor',
'profissao_Coordenador', 'profissao_Diretor', 'profissao_Engenheiro',
'profissao_Gerente', 'profissao_Outros', 'profissao_SEM EXPERIÊNCIA',
'profissao_Supervisor', 'profissao_Sócio/Dono/Proprietário'
]
]
target = df_training['categoria'] ## NÃO TROQUE O NOME DA VARIÁVEL TARGET.
```
### Dividindo nosso conjunto de dados em conjuntos de treinamento e teste
```
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(features, target, test_size=0.2, random_state=133)
```
### Treinando uma árvore de decisão
```
# Método para creacion de modelos basados en arbol de desición
from sklearn.ensemble import GradientBoostingClassifier
dtc = GradientBoostingClassifier().fit(X_train, y_train)
```
### Fazendo previsões na amostra de teste
```
y_pred = dtc.predict(X_test)
print(y_pred)
```
### Analisando a qualidade do modelo através da matriz de confusão
```
import matplotlib.pyplot as plt
import numpy as np
import itertools
def plot_confusion_matrix(cm, target_names, title='Confusion matrix', cmap=None, normalize=True):
accuracy = np.trace(cm) / float(np.sum(cm))
misclass = 1 - accuracy
if cmap is None:
cmap = plt.get_cmap('Blues')
plt.figure(figsize=(8, 6))
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
if target_names is not None:
tick_marks = np.arange(len(target_names))
plt.xticks(tick_marks, target_names, rotation=45)
plt.yticks(tick_marks, target_names)
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
thresh = cm.max() / 1.5 if normalize else cm.max() / 2
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
if normalize:
plt.text(j, i, "{:0.4f}".format(cm[i, j]),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
else:
plt.text(j, i, "{:,}".format(cm[i, j]),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label\naccuracy={:0.4f}; misclass={:0.4f}'.format(accuracy, misclass))
plt.show()
from sklearn.metrics import confusion_matrix
plot_confusion_matrix(confusion_matrix(y_test, y_pred), ['parfil1', 'perfil2', 'perfil3', 'perfil4', 'perfil5', 'perfil6'])
```
<hr>
## Scoring dos dados necessários para entregar a solução
Como entrega da sua solução, esperamos os resultados classificados no seguinte dataset chamado "to_be_scored.csv":
### Download da "folha de respostas"
```
!wget --no-check-certificate --content-disposition https://raw.githubusercontent.com/vanderlei-test/dataset-3/master/to_be_scored.csv
df_to_be_scored = pd.read_csv(r'to_be_scored.csv')
df_to_be_scored.tail()
```
# Atenção!
O dataframe ``to_be_scored`` é a sua "folha de respostas". Note que a coluna "categoria" não existe nessa amostra, que não pode ser então utilizada para treino de modelos de aprendizado supervisionado.
```
df_to_be_scored.info()
```
<hr>
# Atenção!
# Para poder aplicar seu modelo e classificar a folha de respostas, você precisa primeiro aplicar as mesmas transformações com colunas que você aplicou no dataset de treino.
# Não remova ou adicione linhas na folha de respostas.
# Não altere a ordem das linhas na folha de respostas.
# Ao final, as 1000 entradas devem estar classificadas, com os valores previstos em uma coluna chamada "target"
<hr>
Na célula abaixo, repetimos rapidamente os mesmos passos de pré-processamento usados no exemplo dado com árvore de decisão
```
# 1 - Removendo linhas com valores NaN em "certificados" e "total_modulos"
df_to_be_scored_1 = df_to_be_scored.dropna(axis='index', how='any', subset=['certificados', 'total_modulos'])
# 2 - Inputando zeros nos valores faltantes
impute_zeros.fit(X=df_to_be_scored_1)
df_to_be_scored_2 = pd.DataFrame.from_records(
data=impute_zeros.transform(
X=df_to_be_scored_1
),
columns=df_to_be_scored_1.columns
)
# 3 - Remoção de colunas
df_to_be_scored_3 = df_to_be_scored_2.drop(columns=['id', 'graduacao', 'universidade', 'organizacao', 'como_conheceu_lit'], inplace=False)
# 4 - Encoding com "dummy variables"
df_to_be_scored_4 = pd.get_dummies(df_to_be_scored_3, columns=['profissao'])
df_to_be_scored_4.tail()
```
<hr>
Pode ser verificado abaixo que as colunas da folha de resposta agora são idênticas às que foram usadas para treinar o modelo:
```
df_training[
[
'pretende_fazer_cursos_lit', 'interesse_mba_lit',
'importante_ter_certificado', 'horas_semanais_estudo', 'total_modulos',
'modulos_iniciados', 'modulos_finalizados', 'certificados',
'profissao_0', 'profissao_Advogado', 'profissao_Analista',
'profissao_Analista Senior', 'profissao_Assessor',
'profissao_Coordenador', 'profissao_Diretor', 'profissao_Engenheiro',
'profissao_Gerente', 'profissao_Outros', 'profissao_SEM EXPERIÊNCIA',
'profissao_Supervisor', 'profissao_Sócio/Dono/Proprietário'
]
].columns
df_to_be_scored_4.columns
```
# Atenção
Para todas colunas que não existirem no "df_to_be_scored", você pode usar a técnica abaixo para adicioná-las:
```
df_to_be_scored_4['profissao_0'] = 0
y_pred = dtc.predict(df_to_be_scored_4)
df_to_be_scored_4['target'] = y_pred
df_to_be_scored_4.tail()
```
### Salvando a folha de respostas como um arquivo .csv para ser submetido
```
project.save_data(file_name="results.csv", data=df_to_be_scored_4.to_csv(index=False))
```
# Atenção
# A execução da célula acima irá criar um novo "data asset" no seu projeto no Watson Studio. Você precisará realizar o download deste arquivo juntamente com este notebook e criar um arquivo zip com os arquivos **results.csv** e **notebook.ipynb** para submissão. (os arquivos devem estar nomeados desta forma)
<hr>
## Parabéns!
Se você já está satisfeito com a sua solução, vá até a página abaixo e envie os arquivos necessários para submissão.
# https://lit.maratona.dev
| github_jupyter |
# Costa Rican Household Poverty Level Prediction
## Primary Objective
<br>
The target of our analysis would be to predict poverty on a household level as defined in the dataset. Given that dataset is on an individual level in terms of the datapoints, however we will only include the head of the household to stick with our plan of conducting the analysis on a household basis. The following would be the target variable values:
1 = extreme poverty <br>
2 = moderate poverty <br>
3 = vulnerable households <br>
4 = non vulnerable households <br>
<br>
## Special Features
<br>
Out of the 143 features, following features will be treated little differently in our analysis:
<br>
*Id*: a unique identifier for each individual, this should not be a feature that we use!<br>
<br>
*idhogar*: a unique identifier for each household. This variable is not a feature, but will be used to group individuals by household as all individuals in a household will have the same identifier.<br>
<br>
*parentesco1*: indicates if this person is the head of the household.<br>
<br>
*Target*: the label, which should be equal for all members in a household <br>
<br>
## Scoring Metric
<br>
Ultimately we want to build a machine learning model that can predict the integer poverty level of a household. Our predictions will be assessed by the Macro F1 Score.
<br>
## Secondary Objective
<br>
1. Feature Engineering <br>
2. Select a model by comparison <br>
3. Optimise the choice for the model <br>
4. Understand the outputs of the model <br>
5. Draw insights and breakdown the analysis and prediction <br>
<br>
## Data Cleansing
<br>
Let's first go through our dataset and look for errors and missing values and see how we can fix it in this section.
```
# Data manipulation
import pandas as pd
import numpy as np
# Visualization
import matplotlib.pyplot as plt
import seaborn as sns
# Set a few plotting defaults
%matplotlib inline
pd.options.display.max_columns = 150
# Read in data
train = pd.read_csv('train.csv')
test = pd.read_csv('test.csv')
train.head()
test['Target'] = np.nan
data = train.append(test, ignore_index = True)
# Heads of household
heads = data.loc[data['parentesco1'] == 1].copy()
# Labels for training
train_labels = data.loc[(data['Target'].notnull()) & (data['parentesco1'] == 1), ['Target', 'idhogar']]
# Value counts of target
label_counts = train_labels['Target'].value_counts().sort_index()
test.head()
unique_values = train.groupby('idhogar')['Target'].apply(lambda x: x.nunique() == 1)
different_households = unique_values[unique_values != True]
print('There are {} households where the family members do not all have the same target.'.format(len(different_households)))
train[train['idhogar'] == different_households.index[0]][['idhogar', 'parentesco1', 'Target']]
```
As described in the initial background, we concluded that we would analyse the head of each household to predict which poverty level the household belongs in (i.e. parentesco = 1). So here members of the same households would belong to the same target variables which is 3 in our case.
### Families without a head of the household
Now let's check the families that don't have a head in our dataset
```
check_leader = train.groupby('idhogar')['parentesco1'].sum()
#Check for head
no_head = train.loc[train['idhogar'].isin(check_leader[check_leader == 0].index), :]
print('There are {} households without a head.'.format(no_head['idhogar'].nunique()))
```
And check for households with no heads but have different labels.
```
no_head_equal = no_head.groupby('idhogar')['Target'].apply(lambda x: x.nunique() == 1)
print('{} Households with no head have different labels.'.format(sum(no_head_equal == False)))
```
Meaning that there is no household which has no head and the family members have different labels.
### Applying Correct Labels
So given that we identified households which do not have a head however their family members have the same label. SO let's fix it by making sure that every family member in the same family has a common target variable.
```
for each_household in different_households.index:
#find the correct label
true_target = int(train[(train['idhogar'] == each_household) & (train['parentesco1'] == 1.0)]['Target'])
#assign the correct label for each member
train.loc[train['idhogar'] == each_household, 'Target'] = true_target
unique_values = train.groupby('idhogar')['Target'].apply(lambda x: x.nunique() == 1)
different_households = unique_values[unique_values != True]
print('There are {} households where the family members do not all have the same target.'.format(len(different_households)))
```
### Finding Feaures with Missing Values and filling them up
Now we need to carefully go through each variable and see how we can replace the missing values with the values we want to fill.
```
missing_variables = pd.DataFrame(data.isnull().sum()).rename(columns = {0: 'total'})
missing_variables['percent'] = missing_variables['total']/len(data)
missing_variables.sort_values('percent', ascending = False).head(10).drop('Target')
```
Let's start with fillinf missing values in v2a1 which is essentially the Number of Tablets in a households
```
heads.groupby('v18q')['v18q1'].apply(lambda x: x.isnull().sum())
```
Now all the families that do not own a tablet have a NaN as value. So we can assign this NaN as 0 instead!
```
data['v18q1'] = data['v18q1'].fillna(0)
```
Second, we got v2a1 which is Monthly Rent Payment. We have following categorical variable values:
tipovivi1, =1 own and fully paid house <br>
tipovivi2, "=1 own, paying in installments" <br>
tipovivi3, =1 rented <br>
tipovivi4, =1 precarious <br>
tipovivi5, "=1 other(assigned, borrowed)" <br>
<br>
So now for the households that owned and have a missing monthly rent payment, we can set the value of the rent payment to zero. For the other homes, we can leave the missing values to be imputed but we'll add a flag (Boolean) column indicating that these households had missing values.
```
# Fill in households that own the house with 0 rent payment
data.loc[(data['tipovivi1'] == 1), 'v2a1'] = 0
# Create missing rent payment column
data['v2a1-missing'] = data['v2a1'].isnull()
data['v2a1-missing'].value_counts()
```
Now we can move to the next column which is **rez_esc**(years behind in school).<br>
<br>
So it maybe possible that certain datapoints with null value in this column could have the case of none of the family members having any schooling at all. Let's see what are the ages of the family members for which we have null values.
```
data.loc[data['rez_esc'].notnull()]['age'].describe()
```
We can learn that this variable is only defined for individuals between 7 and 19. Anyone older or younger might not have any years of schooling so this can be set to zero. For the rest we shall add a boolean flag.
```
# If individual is over 19 or younger than 7 and missing years behind, set it to 0
data.loc[((data['age'] > 19) | (data['age'] < 7)) & (data['rez_esc'].isnull()), 'rez_esc'] = 0
# Add a flag for those between 7 and 19 with a missing value
data['rez_esc-missing'] = data['rez_esc'].isnull()
#Setting the maximum value to 5
data.loc[data['rez_esc'] > 5, 'rez_esc'] = 5
```
| github_jupyter |
# Lecture 30: Modifying a LeNet for CIFAR
```
%matplotlib inline
import torch
import torch.nn as nn
import torch.nn.functional as F
import numpy as np
from torchvision import transforms,datasets
import torch.optim as optim
from torch.autograd import Variable
import matplotlib.pyplot as plt
import torchvision
import copy
import time
```
## Load data:
```
apply_transform = transforms.Compose([transforms.Resize(32),transforms.ToTensor()])
BatchSize = 100
trainset = datasets.CIFAR10(root='./CIFAR10', train=True, download=True, transform=apply_transform)
trainLoader = torch.utils.data.DataLoader(trainset, batch_size=BatchSize,
shuffle=True, num_workers=4) # Creating dataloader
testset = datasets.CIFAR10(root='./CIFAR10', train=False, download=True, transform=apply_transform)
testLoader = torch.utils.data.DataLoader(testset, batch_size=BatchSize,
shuffle=False, num_workers=4) # Creating dataloader
# Size of train and test datasets
print('No. of samples in train set: '+str(len(trainLoader.dataset)))
print('No. of samples in test set: '+str(len(testLoader.dataset)))
```
## Define network architecture
```
class LeNet(nn.Module):
def __init__(self):
super(LeNet, self).__init__()
self.conv1 = nn.Conv2d(3, 6, 5)
self.pool = nn.MaxPool2d(2, 2)
self.conv2 = nn.Conv2d(6, 16, 5)
self.fc1 = nn.Linear(16 * 5 * 5, 120)
self.fc2 = nn.Linear(120, 84)
self.fc3 = nn.Linear(84, 10)
def forward(self, x):
x = self.pool(F.relu(self.conv1(x)))
x = self.pool(F.relu(self.conv2(x)))
x = x.view(-1, 16 * 5 * 5)
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = self.fc3(x)
return F.log_softmax(x)
net = LeNet()
print(net)
# Check availability of GPU
use_gpu = torch.cuda.is_available()
if use_gpu:
print('GPU is available!')
net = net.cuda()
```
## Define loss function and optimizer
```
criterion = nn.NLLLoss() # Negative Log-likelihood
optimizer = optim.Adam(net.parameters(), lr=1e-4) # Adam
```
## Train the network
```
iterations = 10
trainLoss = []
testAcc = []
start = time.time()
for epoch in range(iterations):
epochStart = time.time()
runningLoss = 0
net.train(True) # For training
for data in trainLoader:
inputs,labels = data
# Wrap them in Variable
if use_gpu:
inputs, labels = Variable(inputs.cuda()), \
Variable(labels.cuda())
else:
inputs, labels = Variable(inputs), Variable(labels)
# Initialize gradients to zero
optimizer.zero_grad()
# Feed-forward input data through the network
outputs = net(inputs)
# Compute loss/error
loss = criterion(outputs, labels)
# Backpropagate loss and compute gradients
loss.backward()
# Update the network parameters
optimizer.step()
# Accumulate loss per batch
runningLoss += loss.data[0]
avgTrainLoss = runningLoss/50000.0
trainLoss.append(avgTrainLoss)
# Evaluating performance on test set for each epoch
net.train(False) # For testing [Affects batch-norm and dropout layers (if any)]
running_correct = 0
for data in testLoader:
inputs,labels = data
# Wrap them in Variable
if use_gpu:
inputs = Variable(inputs.cuda())
outputs = net(inputs)
_, predicted = torch.max(outputs.data, 1)
predicted = predicted.cpu()
else:
inputs = Variable(inputs)
outputs = net(inputs)
_, predicted = torch.max(outputs.data, 1)
running_correct += (predicted == labels).sum()
avgTestAcc = running_correct/10000.0
testAcc.append(avgTestAcc)
# Plotting training loss vs Epochs
fig1 = plt.figure(1)
plt.plot(range(epoch+1),trainLoss,'r-',label='train')
if epoch==0:
plt.legend(loc='upper left')
plt.xlabel('Epochs')
plt.ylabel('Training loss')
# Plotting testing accuracy vs Epochs
fig2 = plt.figure(2)
plt.plot(range(epoch+1),testAcc,'g-',label='test')
if epoch==0:
plt.legend(loc='upper left')
plt.xlabel('Epochs')
plt.ylabel('Testing accuracy')
epochEnd = time.time()-epochStart
print('Iteration: {:.0f} /{:.0f} ; Training Loss: {:.6f} ; Testing Acc: {:.3f} ; Time consumed: {:.0f}m {:.0f}s '\
.format(epoch + 1,iterations,avgTrainLoss,avgTestAcc*100,epochEnd//60,epochEnd%60))
end = time.time()-start
print('Training completed in {:.0f}m {:.0f}s'.format(end//60,end%60))
```
| github_jupyter |
## Data Preparation
Once the dataset has been profiled to gain a better understanding of its dimensionality, its quality, and the content and distribution of values in its features (columns), the next activity is to address any defects that were identified to prepare the data for exploratory data analysis (EDA) and for training and evaluating machine learning models. Depending on which defects were identified, activities may involve removing duplicate observations and deciding whether to exclude observations (rows) or features (columns) that contain missing or corrupt data, or to impute new values into those tuples.
#### Import Libraries
```
import warnings
warnings.filterwarnings("ignore")
import os
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.compose import ColumnTransformer
from sklearn.ensemble import ExtraTreesRegressor
from sklearn.experimental import enable_iterative_imputer
from sklearn.impute import SimpleImputer, IterativeImputer, MissingIndicator
from sklearn.linear_model import BayesianRidge
from sklearn.neighbors import KNeighborsRegressor
from sklearn.pipeline import Pipeline
from sklearn.tree import DecisionTreeRegressor
```
#### Import the Data
```
data_dir = os.path.join(os.getcwd(), 'Data')
source_data_file = 'titanic.csv';
data_file = os.path.join(data_dir, source_data_file)
df = pd.read_csv(data_file, header=0, index_col=0)
# Ensure the index values are: seed=1, increment=1
df.reset_index(drop=True, inplace=True)
print(f"Shape: {df.shape[0]} Observations x {df.shape[1]} Features")
```
### 1.0. Removing Duplicate Observations
While performing the initial profile it was determined that there were 2 more observations (rows) then there were unique values in the **name** feature. We then identified two passengers (Kate Connolly and James Kelly) with duplicate records. In each case, the first observation contained fewer missing values (NaN) so they should be kept with the second instances being excluded. For the sake of training a model this really shouldn't have any appreciable impact, but it is useful to know how to de-duplicate a dataset in situations where there could possibly be a negative impact. Also, since each passenger's *name* is certain to have no logical correlation to whether or not they survived, once the data has been de-duplicated it will be safe to exclude the **name** feature.
```
df.drop_duplicates(subset='name', keep='first', inplace=True)
df.reset_index(drop=True, inplace=True)
df.drop(['name'], axis=1, inplace=True)
print(f"Shape: {df.shape[0]} Observations x {df.shape[1]} Features")
```
### 2.0. Modifying the Data Type of Categorical Features Having Numerical Values
While performing the initial profile, some features having numerical data types were detected that contain values which may also be considered categorical (e.g., survived, sibsp, parch). While **sibsp** and **parch** are, in fact, summable numerical values their discrete nature and limited range indicates that they may influence the *Target* more if treated as categorical values. Here those features will be converted from their present numerical types to the *Object* type to indicate their categorical nature. What's more, a new feature (survived_desc) will be created wherein descriptive labels will be mapped to the Target variable's (survived) existing numerical values. This new column will be used to enhance some visualizations that will be created while conducting exploratory data analysis (EDA).
```
df['survived_desc'] = df.survived.map({0 : 'perished', 1 : 'survived'})
df[['survived','sibsp','parch']] = df[['survived','sibsp','parch']].astype('object')
df.dtypes
```
### 3.0. Complete-Case Analysis *(Removing Missing Values)*
Complete Case Analysis (CCA), also called "list-wise deletion" of cases, involves analyzing only those observations where all of the variables in the data set contain useful data. Subsequently, complete-case analysis consists of discarding observations where values in any of the variables are missing. Implementing CCA is prudent for the sake of avoiding any future doubt with regards to either the validity of any correlations that might be observed among the features, or the efficacy of any models that might be trained using the sample. However, in situations where the remaining sample would be too small to effectively train and test machine learning models there may be no other choice, besides sourcing additional data, than to impute missing values.
#### 3.1. Quantify Missing Values
```
df.isnull().sum().sort_values(ascending=True)
```
#### 3.2. Experiment with Dropping All Observations Containing Missing Values (CCA)
```
df_cca = df.dropna()
print(f"Shape: {df_cca.shape[0]} Observations x {df_cca.shape[1]} Features")
```
#### 3.3. Experiment with Dropping Observations by Subsetting Features
```
df_cca = df.dropna(subset=['survived','sex','sibsp','parch','ticket'
,'fare','embarked','age','home.dest'
#,'home.dest','boat','cabin','body'
])
print(f"Shape: {df_cca.shape[0]} Observations x {df_cca.shape[1]} Features")
```
Our experimentation has revealed that there are so many missing values that removing them all would either leave too few observations, or exclude potentially influential features. In this situation it may be necessary to experiment with various methods for imputing missing values.
### 4.0. Imputing Missing Values
The following sections demonstrate how to impute missing values (e.g., NaN or NULL) using two different approaches: using Pandas, and using Scikit-Learn's SimpleImputer. The advantage of using the SimpleImputer is that is can be used to build Scikit-Learn Pipelines which both streamline and formalize the process into reusable processes called *pipelines*.
The approach implemented to impute missing values is driven by the nature of the data the feature contains. The first thing to consider is whether the feature contains numerical or categorical data. If the feature is numerical then the following techniques may be most appropriate:
- Imputing with the Mean or Median
- Imputing with the Mode
- Imputing with an Arbitrary Number
- Imputing with a Value at the End of the Distribution
- Imputing with a Random Sample
- Imputing with MICE (Multivariate Imputation by Chained Equations)
If the feature is categorical then the following techniques may be more appropriate:
- Imputing with a Custom Category
- Imputing with the Most Frequent Category
- Imputing with MICE (Multivariate Imputation by Chained Equations)
- Imputing with a Missing-Value Indicator
#### Separate Numerical and Categorical Features
```
numerical_cols = [col for col in df.columns if df.dtypes[col] != 'O']
categorical_cols = [col for col in df.columns if col not in numerical_cols]
print(numerical_cols)
print(categorical_cols)
```
### 4.1. Imputing Numerical Values
#### 4.1.1. Impute Missing Values with the Median or the Mean
When imputing numerical variables a reasonable first approach would be to estimate missing values using either the **mean** of the remaining non-null observations, if the variable is reasonable parametric, or with the **median** of those remaining values if the variable contains outliers that would apply too much leverage against its appropriate mean.
##### Using Pandas
```
df_pd = df.copy() # Make a copy of the dataframe.
for col in numerical_cols:
median = df_pd[col].median()
df_pd[col] = df_pd[col].fillna(median)
df_pd[numerical_cols].isnull().sum()
```
##### Using Scikit-Learn
```
df_sk = df.copy() # Make a copy of the dataframe
imputer = SimpleImputer(strategy='median')
imputer.fit(df_sk[numerical_cols])
imputer.statistics_
```
#### 4.1.2. Impute Missing Values with the Mode
Mode imputation consists of replacing all occurrences of missing values (NA) within a variable by the mode; i.e., the most frequently occuring value.
##### Using Pandas
```
df_pd = df.copy() # Make a copy of the dataframe.
for col in numerical_cols:
mode = df_pd[col].mode()[0]
df_pd[col] = df_pd[col].fillna(mode)
df_pd[numerical_cols].isnull().sum()
```
##### Using Scikit-Learn
```
df_sk = df.copy() # Make a copy of the dataframe
imputer = SimpleImputer(strategy='most_frequent')
imputer.fit(df_sk[numerical_cols])
imputer.statistics_
```
#### 4.1.3. Impute Missing Values with an Arbitrary Number
##### First, inspect the maximum value per feature to ensure the arbitrary number doesn't overlap existing values
```
df[numerical_cols].max()
df_pd = df.copy() # Make a copy of the dataframe.
for col in numerical_cols:
df_pd[col].fillna(999, inplace=True)
df_pd[numerical_cols].isnull().sum()
```
##### Using Scikit-Learn
```
df_sk = df.copy() # Make a copy of the dataframe
imputer = SimpleImputer(strategy='constant', fill_value=999)
imputer.fit(df_sk[numerical_cols])
imputer.statistics_
```
#### 4.1.4. Imputing with a Value at the End of the Distribution
Here we will replace missing values by a value at the end of the distribution, estimated with the Gaussian approximation or the inter-quantal range proximity rule, using Pandas.
```
df_pd = df.copy() # Make a copy of the dataframe.
for col in numerical_cols:
IQR = df_pd[col].quantile(0.75) - df_pd[col].quantile(0.25)
value = df_pd[col].quantile(0.75) + 1.25 * IQR
df_pd[col] = df_pd[col].fillna(value)
df_pd[numerical_cols].isnull().sum()
```
#### 4.1.5. Impute Missing Values with a Random Sample
```
df_pd = df.copy() # Make a copy of the dataframe.
for col in numerical_cols:
number_missing_values = df_pd[col].isnull().sum()
random_sample = df_pd[col].dropna().sample(number_missing_values, replace=True, random_state=0)
random_sample.index = df_pd[df_pd[col].isnull()].index
df_pd.loc[df_pd[col].isnull(), col] = random_sample
df_pd[numerical_cols].isnull().sum()
```
#### 4.1.6. Impute Missing Values with MICE (Multivariate Imputation by Chained Equations) Using Scikit-Learn
The imputation techniques implemented so far have been **univariate** imputations; i.e., the values are either statitically assigned, or are estimated (calculated) using the non-null values present in the specified variable. Conversely, **multivariate** imputation techniques estimate new values taking into account the values present in all the variables (features) in the dataset. Multivariate Imputation by Chained Equations (MICE) is an imputation technique that models each variable with missing values as a function of the remaining variables and uses that estimate for imputation. Each variable with missing data can be modeled based on the remaining variable using any one of many estimators (e.g., Bayes, k-nearest neighbors, decision trees, random forests, linear regression).
##### Using a Bayesian Ridge estimator
```
df_sk = df.copy() # Make a copy of the dataframe.
bayes = IterativeImputer(estimator=BayesianRidge(), max_iter=10, random_state=0)
bayes.fit(df_sk[numerical_cols])
df_bayes = bayes.transform(df_sk[numerical_cols])
```
##### Use a K-Nearest Neighbors (KNN) estimator
```
knn = IterativeImputer(estimator=KNeighborsRegressor(n_neighbors=5), max_iter=10, random_state=0)
knn.fit(df_sk[numerical_cols])
df_knn = knn.transform(df_sk[numerical_cols])
```
##### Use a Decision Tree Regressor estimator
```
dtr = IterativeImputer(estimator=DecisionTreeRegressor(max_features='sqrt', random_state=0), max_iter=10, random_state=0)
dtr.fit(df_sk[numerical_cols])
df_dtr = dtr.transform(df_sk[numerical_cols])
```
##### Use an Extra Trees Regressor estimator
```
etr = IterativeImputer(estimator=ExtraTreesRegressor(n_estimators=10, random_state=0), max_iter=10, random_state=0)
etr.fit(df_sk[numerical_cols])
df_etr = etr.transform(df_sk[numerical_cols])
```
##### Plot to Compare the Performance of Each estimator
```
df_bayes = pd.DataFrame(df_bayes, columns=numerical_cols)
df_knn = pd.DataFrame(df_knn, columns=numerical_cols)
df_dtr = pd.DataFrame(df_dtr, columns=numerical_cols)
df_etr = pd.DataFrame(df_etr, columns=numerical_cols)
fig = plt.figure()
ax = fig.add_subplot(111)
df.age.plot(kind='kde', ax=ax, color='black')
df_bayes.age.plot(kind='kde', ax=ax, color='red')
df_knn.age.plot(kind='kde', ax=ax, color='green')
df_dtr.age.plot(kind='kde', ax=ax, color='blue')
df_etr.age.plot(kind='kde', ax=ax, color='orange')
lines, labels = ax.get_legend_handles_labels()
labels = ['Original','Bayes','KNN','Trees','Forest']
ax.legend(lines, labels, loc='best')
plt.show()
```
This comparison clearly illustrates that the Bayes algorithm outperforms the others; therefore, it should be used for any MICE imputations against this dataset.
### 4.2. Imputing Categorical Values
##### Inspect the percentage of missing values in each categorical variable
```
df[categorical_cols].isnull().mean()
```
##### Exclude categorical variables having no missing values
```
del categorical_cols[0:5] # survived, sex, sibsp, parch, ticket
del categorical_cols[-1] # survived_desc
categorical_cols
```
#### 4.2.1. Impute Missing Values with a Custom Category
When handling missing values in Categorical features, it is customary to create an additional category by imputing the value 'Unknown', or 'Missing' into those tuples where NULL or NaN values occur.
##### Using Pandas
```
df_pd = df.copy()
for col in categorical_cols:
df_pd[col].fillna('Unknown', inplace=True)
df_pd[categorical_cols].isnull().sum()
```
##### Using Scikit-Learn
```
df_sk = df.copy()
imputer = SimpleImputer(strategy='constant', fill_value='Unknown')
imputer.fit(df_sk[categorical_cols])
imputer.statistics_
```
#### 4.2.2. Impute Missing Values with the Most Frequently Category
Most frequent category imputation consists of replacing all occurrences of missing values within a variable with the most frequently occuring value (mode).
##### Using Pandas
```
df_pd = df.copy()
for col in categorical_cols:
mode = df_pd[col].mode()[0]
df_pd[col] = df_pd[col].fillna(mode)
df_pd[categorical_cols].isnull().sum()
```
##### Using Scikit-Learn
```
df_sk = df.copy()
imputer = SimpleImputer(strategy='most_frequent')
imputer.fit(df_sk[numerical_cols])
imputer.statistics_
```
#### 4.2.3. Impute Missing Values with a Missing-Value Indicator
##### Using Pandas
```
df_pd = df.copy()
for col in categorical_cols:
df_pd[col+'_NA'] = np.where(df_pd[col].isnull(), 1, 0)
df_pd.head(2)
```
##### Using Scikit-Learn
```
df_sk = df.copy()
indicator = MissingIndicator(error_on_new=True, features='missing-only')
indicator.fit(df_sk)
indicator.features_
```
### 5.0. Building an Imputation Pipeline
The previous sections have demonstrated how to implement numerous imputation methods, first using Pandas to convey the conceptual meaning of each technique, and then using various Scikit-Learn imputers that abstract the implementation details. When productionalizing a machine learning model as a service it should be assumed that new data will be submitted to that service in an unprepared condition; i.e., very similar to the 'raw' dataset provided for training the machine learning model. Therefore, that new 'dirty' data must be prepared exactly as it was during the development process. The easiest method for ensuring this is to codify each transformation (e.g., imputation, datatype change, variable-name change) in a reusable construct that accurately represents each step in the exact order as it was implemented when the training dataset was prepared. Fortunately, Scikit-Learn (and other ML frameworks) have such a construct: the Pipeline. In this section, constructing a data preparation pipeline will be demonstrated.
#### 5.1. Specify Imputation Techniques Per Feature:
- Since **age** is consistantly between 21 & 39 with some outliers, Median may be appropriate.
- Since **fare** is likely related to other features like Cabin, MICE may be appropriate.
- Since **body** weight is likely related to other features like Sex and Age, MICE may be appropriate
- Since not everyone aboard made it to a life**boat**, a custom category like 'unknown' is appropriate.
- Since **cabin, embarked** and **home.dest**ination are categorical, most frequent values may be appropriate.
- Since it's advantageous to keep track of which tuples have been imputed, missing indicator columns should be created.
```
num_cols_median = ['age']
num_cols_mice = ['fare','body']
cat_cols_mode = ['cabin','embarked','home.dest']
cat_cols_custom = ['boat']
cat_cols_missing = ['cabin','embarked','boat','home.dest']
```
##### 5.2. Instantiate each imputer in its own Pipeline
```
imp_num_median = Pipeline(steps=[
('imputer', SimpleImputer(strategy='median'))
])
imp_num_mice = Pipeline(steps=[
('imputer', IterativeImputer(estimator=BayesianRidge(), max_iter=10, random_state=0))
])
imp_cat_mode = Pipeline(steps=[
('imputer', SimpleImputer(strategy='most_frequent'))
])
imp_cat_custom = Pipeline(steps=[
('imputer', SimpleImputer(strategy='constant', fill_value='unknown'))
])
imp_cat_missing = Pipeline(steps=[
('imputer', MissingIndicator(error_on_new=True, features='missing-only'))
])
```
#### 5.3. Align features with their corresponding Imputers in a Column Transformer
```
preprocessor = ColumnTransformer(transformers=[('num_median', imp_num_median, num_cols_median),
('num_mice', imp_num_mice, num_cols_mice),
('cat_mode', imp_cat_mode, cat_cols_mode),
('cat_custom', imp_cat_custom, cat_cols_custom),
('cat_missing', imp_cat_missing, cat_cols_missing)
], remainder='passthrough')
```
#### 5.4. Fit the preprocessor and impute the data
```
preprocessor.fit(df)
df_imputed = preprocessor.transform(df)
columns = ['age','fare','body','cabin','embarked','home.dest','boat'
,'cabin_NA','embarked_NA','boat_NA','home.dest_NA'
,'survived','sex','sibsp','parch','ticket','survived_desc'
]
df_prepared = pd.DataFrame(df_imputed, columns=columns)
df_prepared.isnull().sum()
df_prepared.head(2)
```
### 6.0. Export the Prepared Data for Further Exploration
```
prepared_titanic_file = os.path.join(data_dir, 'titanic-eda.csv')
df_prepared.to_csv(prepared_titanic_file, index=True)
```
| github_jupyter |
# Lecture 7: Gambler's Ruin & Random Variables
## Stat 110, Prof. Joe Blitzstein, Harvard University
----
## Gambler's Ruin
Two gamblers $A$ and $B$ are successively playing a game until one wins all the money and the other is ruined (goes bankrupt). There is a sequence of rounds, with a one dollar bet each time. The rounds are independent events. Let $p = P(\text{A wins a certain round})$ and the inverse is $q = 1 - p$, by convention.
_What is the probability that $A$ wins the entire game?_
Some clarifications:
* there is a total of $N$ dollars in this closed system game (no other money comes into play)
* $A$ starts with $i$ dollars, $B$ starts with $N-i$ dollars
But where do we begin to solve this problem?
### Random Walk
A [random walk](https://en.wikipedia.org/wiki/Random_walk) between two points on a number line is very similar to the Gambler's Ruin.

How many rounds could a game last? Is it possible for the game to continue on to infinity?
Well, notice how this has a very nice __recursive nature__. If $A$ loses a round, the game can be seen as starting anew at $i-1$, and if he wins, the game would start anew at $i+1$. It is the same problem, but with a different starting condition.
### Strategy
Conditioning on the _first step_ is called __first step analysis__.
Let $P_i = P(\text{A wins the entire game|A starts with i dollars})$. Then from the Law of Total Probability, we have:
\begin{align}
P_i &= p P_{i+1} + q P_{i-1} \text{, } & &\text{where }1 \lt i \lt N-1 \\
& & & P_0 = 0 \\
& & & P_N = 1 \\
\end{align}
See how this is a recursive equation? This is called a [__difference equation__](http://mathworld.wolfram.com/DifferenceEquation.html), which is a discrete analog of a differential equation.
### Solving the Difference Equation
\begin{align}
P_i &= p P_{i+1} + q P_{i-1} & & \\
\\
\\
P_i &= x^i & &\text{see what happens when we guess with a power} \\
\Rightarrow x^i &= p x^{i+1} + q x^{i-1} \\
0 &= p x^2 - x + q & &\text{factoring out } x^{i-1} \text{, we are left with a quadratic}\\
\\
x &= \frac{1 \pm \sqrt{1-4pq}}{2p} & &\text{solving with the quadratic formula} \\
&= \frac{1 \pm \sqrt{(2p-1)^2}}{2p} & &\text{since }1-4pq = 1-4p(1-p) = 4p^2 - 4p - 1 = (2p -1)^2 \\
&= \frac{1 \pm (2p-1)}{2p} \\
&\in \left\{1, \frac{q}{p} \right\} \\
\\
\\
P_i &= A(1)^i + B\left(\frac{q}{p}\right)^i & &\text{if } p \neq q~~~~ \text{(general solution for difference equation)} \\
\Rightarrow B &= -A & &\text{from }P_0 = 1\\
\Rightarrow 1 &= A(1)^N + B\left(\frac{q}{p}\right)^N & &\text{from } P_N = 1\\
&= A(1)^N - A\left(\frac{q}{p}\right)^N \\
&= A\left(1-\frac{q}{p}\right)^N \\
\\
\\
\therefore P_i &=
\begin{cases}
\frac{1-\left(\frac{q}{p}\right)^i}{1-\left(\frac{q}{p}\right)^N} & \quad \text{ if } p \neq q \\
\frac{i}{N} & \quad \text{ if } p = q \\
\end{cases}
\end{align}
### Example calculations of $P_i$ over a range of $N$
Assuming an unfair game where $p=0.49$, $q=0.51$:
```
import math
def gamblers_ruin(i, p, q, N):
if math.isclose(p,q):
return i/N
else:
return ((1 - (q/p)**i)) / (1 - (q/p)**N)
p = 0.49
q = 1.0 - p
N = 20
i = N/2
print("With N={} and p={}, probability that A wins all is {:.2f}".format(N, p, gamblers_ruin(i, p, q, N)))
N = 100
i = N/2
print("With N={} and p={}, probability that A wins all is {:.2f}".format(N, p, gamblers_ruin(i, p, q, N)))
N = 200
i = N/2
print("With N={} and p={}, probability that A wins all is {:.2f}".format(N, p, gamblers_ruin(i, p, q, N)))
```
And assuming a fair game where $p = q = 0.5$:
```
p = 0.5
q = 1.0 - p
N = 20
i = N/2
print("With N={} and p={}, probability that A wins all is {:.2f}".format(N, p, gamblers_ruin(i, p, q, N)))
N = 100
i = N/2
print("With N={} and p={}, probability that A wins all is {:.2f}".format(N, p, gamblers_ruin(i, p, q, N)))
N = 200
i = N/2
print("With N={} and p={}, probability that A wins all is {:.2f}".format(N, p, gamblers_ruin(i, p, q, N)))
```
#### Could the game ever continue forever on to infinity?
Recall that we have the following solution to the difference equation for the Gambler's Ruin game:
\begin{align}
P_i &=
\begin{cases}
\frac{1-\left(\frac{q}{p}\right)^i}{1-\left(\frac{q}{p}\right)^N} & \quad \text{ if } p \neq q \\
\frac{i}{N} & \quad \text{ if } p = q \\
\end{cases}
\end{align}
The only time you'd think the game could continue on to infinity is when $p=q$. But
\begin{align}
P(\Omega) &= 1\\
&= P(\text{A wins all}) + P(\text{B wins all}) \\
&= P_i + P_{N-i} \\
&= \frac{i}{N} + \frac{N-i}{N}
\end{align}
The above implies that aside from the case where $A$ wins all, and the case where $B$ wins all, there is no other event in $\Omega$ to consider, hence the game can never continue on to infinity without either side winning.
This also means that unless $p=q$, you __will__ lose your money, and the only question is how fast will you lose it.
----
# Random Variables
Consider these statements:
\begin{align}
x + 2 &= 9 \\
x &= 7
\end{align}
_What is a variable?_
* variable $x$ is a symbol that we use as a substitute for an arbitrary _constant_ value.
_What is a __random__ variable?_
* This is not a _variable_, but a __function from the sample space $S$ to $\mathbb{R}$__.
* It is a "summary" of an aspect of the experiment (this is where the randomness comes from)
Here are a few of the most useful _discrete random variables_.
----
## Bernoulli Distribution
### Description
A probability distribution of a random variable that takes the value 1 in the case of a success with probability $p$; or takes the value 0 in case of a failure with probability $1-p$.
A most common example would be a coin toss, where heads might be considered a success with probability $p=0.5$ if the coin is a fair.
A random variable $x$ has the Bernoulli distribution if
- $x \in \{0, 1\}$
- $P(x=1) = p$
- $P(x=0) = 1-p$
### Notation
$X \sim \operatorname{Bern}(p)$
### Parameters
$0 < p < 1 \text{, } p \in \mathbb{R}$
### Probability mass function
The probability mass function $P(x)$ over possible values $x$
\begin{align}
P(x) =
\begin{cases}
1-p, &\text{ if } x = 0 \\
p, &\text{ if } x = 1 \\
\end{cases} \\
\end{align}
### Expected value
\begin{align}
\mathbb{E}(X) &= 1 P(X=1) + 0 P(X=0) \\
&= p
\end{align}
### Special case: Indicator random variables (r.v.)
\begin{align}
&X =
\begin{cases}
1, &\text{ if event A occurs} \\
0, &\text{ otherwise} \\
\end{cases} \\
\\
\\
\Rightarrow &\mathbb{E}(X) = P(A)
\end{align}
## Binomial Distribution
### Description
The distribution of the number of successes in $n$ independent Bernoulli trials $\operatorname{Bern}(p)$, where the chance of success $p$ is the same for all trials $n$.
Another case might be a string of indicator random variables.
### Notation
$X \sim \operatorname{Bin}(n, p)$
### Parameters
- $n \in \mathbb{N}$
- $p \in [0,1]$
```
import matplotlib
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.ticker import (MultipleLocator, FormatStrFormatter,
AutoMinorLocator)
from scipy.stats import binom
%matplotlib inline
plt.xkcd()
_, ax = plt.subplots(figsize=(12,8))
# a few Binomial parameters n and p
pop_sizes = [240, 120, 60, 24]
p_values = [0.2, 0.3, 0.4, 0.8]
params = list(zip(pop_sizes, p_values))
# colorblind-safe, qualitative color scheme
colors = ['#a6cee3','#1f78b4','#b2df8a','#33a02c']
for i,(n,p) in enumerate(params):
x = np.arange(binom.ppf(0.01, n, p), binom.ppf(0.99, n, p))
y = binom.pmf(x, n, p)
ax.plot(x, y, 'o', ms=8, color=colors[i], label='n={}, p={}'.format(n,p))
ax.vlines(x, 0, y, color=colors[i], alpha=0.3)
# legend styling
legend = ax.legend()
for label in legend.get_texts():
label.set_fontsize('large')
for label in legend.get_lines():
label.set_linewidth(1.5)
# y-axis
ax.set_ylim([0.0, 0.23])
ax.set_ylabel(r'$P(x=k)$')
# x-axis
ax.set_xlim([10, 65])
ax.set_xlabel('# of successes k out of n Bernoulli trials')
# x-axis tick formatting
majorLocator = MultipleLocator(5)
majorFormatter = FormatStrFormatter('%d')
minorLocator = MultipleLocator(1)
ax.xaxis.set_major_locator(majorLocator)
ax.xaxis.set_major_formatter(majorFormatter)
ax.grid(color='grey', linestyle='-', linewidth=0.3)
plt.suptitle(r'Binomial PMF: $P(x=k) = \binom{n}{k} p^k (1-p)^{n-k}$')
plt.show()
```
### Probability mass function
\begin{align}
P(x=k) &= \binom{n}{k} p^k (1-p)^{n-k}
\end{align}
### Expected value
\begin{align}
\mathbb{E}(X) = np
\end{align}
## In parting...
Now think about this true statement as we move on to Lecture 3:
\begin{align}
X &\sim \operatorname{Bin}(n,p) \text{, } Y \sim \operatorname{Bin}(m,p) \\
\rightarrow X+Y &\sim \operatorname{Bin}(n+m, p)
\end{align}
----
## Appendix A: Solving $P_i$ when $p=q$ using l'Hopital's Rule
To solve for for the case where $p = q$, let $x = \frac{q}{p}$.
\begin{align}
lim_{x \rightarrow 1}{\frac{1-x^i}{1-x^N}} &= lim_{x\rightarrow1}{\frac{ix^{i-1}}{Nx^{N-1}}} &\text{ by l'Hopital's Rule} \\
&= \frac{i}{N}
\end{align}
----
View [Lecture 7: Gambler's Ruin and Random Variables | Statistics 110](http://bit.ly/2PmMbdV) on YouTube.
| github_jupyter |
##### Copyright 2018 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
#@title MIT License
#
# Copyright (c) 2017 François Chollet
#
# Permission is hereby granted, free of charge, to any person obtaining a
# copy of this software and associated documentation files (the "Software"),
# to deal in the Software without restriction, including without limitation
# the rights to use, copy, modify, merge, publish, distribute, sublicense,
# and/or sell copies of the Software, and to permit persons to whom the
# Software is furnished to do so, subject to the following conditions:
#
# The above copyright notice and this permission notice shall be included in
# all copies or substantial portions of the Software.
#
# THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR
# IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY,
# FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL
# THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER
# LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING
# FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER
# DEALINGS IN THE SOFTWARE.
```
# 과대적합과 과소적합
<table class="tfo-notebook-buttons" align="left">
<td> <a target="_blank" href="https://www.tensorflow.org/tutorials/keras/overfit_and_underfit"><img src="https://www.tensorflow.org/images/tf_logo_32px.png">TensorFlow.org에서 보기</a> </td>
<td><a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/ko/tutorials/keras/overfit_and_underfit.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png">Google Colab에서 실행</a></td>
<td><a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/ko/tutorials/keras/overfit_and_underfit.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png">GitHub에서 소스 보기</a></td>
<td><a href="https://storage.googleapis.com/tensorflow_docs/docs-l10n/site/ko/tutorials/keras/overfit_and_underfit.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png">노트북 다운로드</a></td>
</table>
지금까지 그랬듯이 이 예제의 코드도 `tf.keras` API를 사용합니다. 텐서플로 [케라스 가이드](https://www.tensorflow.org/guide/keras)에서 `tf.keras` API에 대해 더 많은 정보를 얻을 수 있습니다.
앞서 영화 리뷰 분류와 주택 가격 예측의 두 예제에서 일정 에포크 동안 훈련하면 검증 세트에서 모델 성능이 최고점에 도달한 다음 감소하기 시작한 것을 보았습니다.
다른 말로 하면, 모델이 훈련 세트에 *과대적합*(overfitting)된 것입니다. 과대적합을 다루는 방법은 꼭 배워야 합니다. *훈련 세트*에서 높은 성능을 얻을 수 있지만 진짜 원하는 것은 *테스트 세트*(또는 이전에 본 적 없는 데이터)에 잘 일반화되는 모델입니다.
과대적합의 반대는 *과소적합*(underfitting)입니다. 과소적합은 테스트 세트의 성능이 향상될 여지가 아직 있을 때 일어납니다. 발생하는 원인은 여러가지입니다. 모델이 너무 단순하거나, 규제가 너무 많거나, 그냥 단순히 충분히 오래 훈련하지 않는 경우입니다. 즉 네트워크가 훈련 세트에서 적절한 패턴을 학습하지 못했다는 뜻입니다.
모델을 너무 오래 훈련하면 과대적합되기 시작하고 테스트 세트에서 일반화되지 못하는 패턴을 훈련 세트에서 학습합니다. 과대적합과 과소적합 사이에서 균형을 잡아야 합니다. 이를 위해 적절한 에포크 횟수동안 모델을 훈련하는 방법을 배워보겠습니다.
과대적합을 막는 가장 좋은 방법은 더 많은 훈련 데이터를 사용하는 것입니다. 많은 데이터에서 훈련한 모델은 자연적으로 일반화 성능이 더 좋습니다. 데이터를 더 준비할 수 없을 때 그다음으로 가장 좋은 방법은 규제(regularization)와 같은 기법을 사용하는 것입니다. 모델이 저장할 수 있는 정보의 양과 종류에 제약을 부과하는 방법입니다. 네트워크가 소수의 패턴만 기억할 수 있다면 최적화 과정 동안 일반화 가능성이 높은 가장 중요한 패턴에 촛점을 맞출 것입니다.
이 노트북에서 널리 사용되는 두 가지 규제 기법인 가중치 규제와 드롭아웃(dropout)을 알아 보겠습니다. 이런 기법을 사용하여 IMDB 영화 리뷰 분류 모델의 성능을 향상시켜 보죠.
이 노트북에서는 몇 가지 일반적인 정규화 기술을 살펴보고 분류 모델을 개선하는 데 사용할 것입니다.
## 설정
시작하기 전에 필요한 패키지를 가져옵니다.
```
import tensorflow as tf
from tensorflow import keras
import numpy as np
import matplotlib.pyplot as plt
print(tf.__version__)
!pip install git+https://github.com/tensorflow/docs
import tensorflow_docs as tfdocs
import tensorflow_docs.modeling
import tensorflow_docs.plots
from IPython import display
from matplotlib import pyplot as plt
import numpy as np
import pathlib
import shutil
import tempfile
logdir = pathlib.Path(tempfile.mkdtemp())/"tensorboard_logs"
shutil.rmtree(logdir, ignore_errors=True)
```
## IMDB 데이터셋 다운로드
이 튜토리얼의 목표는 입자 물리학을 수행하는 것이 아니므로 데이터 세트의 세부 사항에 집착하지 마세요. 여기에는 각각 28개의 특성과 이진 클래스 레이블이 있는 11,000,000개의 예제가 포함되어 있습니다.
```
gz = tf.keras.utils.get_file('HIGGS.csv.gz', 'http://mlphysics.ics.uci.edu/data/higgs/HIGGS.csv.gz')
FEATURES = 28
```
`tf.data.experimental.CsvDataset` 클래스는 중간 압축 해제 단계 없이 gzip 파일에서 직접 csv 레코드를 읽는 데 사용할 수 있습니다.
```
ds = tf.data.experimental.CsvDataset(gz,[float(),]*(FEATURES+1), compression_type="GZIP")
```
해당 csv 판독기 클래스는 각 레코드에 대한 스칼라 목록을 반환합니다. 다음 함수는 해당 스칼라 목록을 (feature_vector, label) 쌍으로 다시 압축합니다.
```
def pack_row(*row):
label = row[0]
features = tf.stack(row[1:],1)
return features, label
```
TensorFlow는 대규모 데이터 배치에서 작업할 때 가장 효율적입니다.
따라서 각 행을 개별적으로 다시 압축하는 대신 10000개 예제의 배치를 취하고 각 배치에 `pack_row` 함수를 적용한 다음 배치를 다시 개별 레코드로 분할하는 새로운 `Dataset`를 만듭니다.
```
packed_ds = ds.batch(10000).map(pack_row).unbatch()
```
이 새로운 `packed_ds`의 일부 레코드를 살펴보세요.
특성이 완벽하게 정규화되지는 않았지만 이 튜토리얼에서는 이것으로 충분합니다.
```
for features,label in packed_ds.batch(1000).take(1):
print(features[0])
plt.hist(features.numpy().flatten(), bins = 101)
```
이 튜토리얼을 비교적 짧게 유지하기 위해 처음 1000개의 샘플만 검증에 사용하고 다음 10,000개는 훈련에 사용합니다.
```
N_VALIDATION = int(1e3)
N_TRAIN = int(1e4)
BUFFER_SIZE = int(1e4)
BATCH_SIZE = 500
STEPS_PER_EPOCH = N_TRAIN//BATCH_SIZE
```
`Dataset.skip` 및 `Dataset.take` 메서드를 사용하면 이를 쉽게 수행할 수 있습니다.
동시에 `Dataset.cache` 메서드를 사용하여 로더가 각 epoch에서 파일의 데이터를 다시 읽을 필요가 없도록 합니다.
```
validate_ds = packed_ds.take(N_VALIDATION).cache()
train_ds = packed_ds.skip(N_VALIDATION).take(N_TRAIN).cache()
train_ds
```
이러한 데이터세트는 개별 예제를 반환합니다. `.batch` 메서드를 사용하여 훈련에 적합한 크기의 배치를 생성합니다. 또한 배치 처리하기 전에 훈련 세트에 대해 `.shuffle` 및 `.repeat`를 수행하는 것도 잊지 마세요.
```
validate_ds = validate_ds.batch(BATCH_SIZE)
train_ds = train_ds.shuffle(BUFFER_SIZE).repeat().batch(BATCH_SIZE)
```
## 과대적합 예제
과대적합을 막는 가장 간단한 방법은 모델의 규모를 축소하는 것입니다. 즉, 모델에 있는 학습 가능한 파라미터의 수를 줄입니다(모델 파라미터는 층(layer)의 개수와 층의 유닛(unit) 개수에 의해 결정됩니다). 딥러닝에서는 모델의 학습 가능한 파라미터의 수를 종종 모델의 "용량"이라고 말합니다. 직관적으로 생각해 보면 많은 파라미터를 가진 모델이 더 많은 "기억 용량"을 가집니다. 이런 모델은 훈련 샘플과 타깃 사이를 일반화 능력이 없는 딕셔너리와 같은 매핑으로 완벽하게 학습할 수 있습니다. 하지만 이전에 본 적 없는 데이터에서 예측을 할 땐 쓸모가 없을 것입니다.
항상 기억해야 할 점은 딥러닝 모델이 훈련 세트에는 학습이 잘 되는 경향이 있지만 진짜 해결할 문제는 학습이 아니라 일반화라는 것입니다.
반면에 네트워크의 기억 용량이 부족하다면 이런 매핑을 쉽게 학습할 수 없을 것입니다. 손실을 최소화하기 위해서는 예측 성능이 더 많은 압축된 표현을 학습해야 합니다. 또한 너무 작은 모델을 만들면 훈련 데이터를 학습하기 어렵울 것입니다. "너무 많은 용량"과 "충분하지 않은 용량" 사이의 균형을 잡아야 합니다.
안타깝지만 어떤 모델의 (층의 개수나 뉴런 개수에 해당하는) 적절한 크기나 구조를 결정하는 마법같은 공식은 없습니다. 여러 가지 다른 구조를 사용해 실험을 해봐야만 합니다.
알맞은 모델의 크기를 찾으려면 비교적 적은 수의 층과 파라미터로 시작해서 검증 손실이 감소할 때까지 새로운 층을 추가하거나 층의 크기를 늘리는 것이 좋습니다. 영화 리뷰 분류 네트워크를 사용해 이를 실험해 보죠.
`Dense` 층만 사용하는 간단한 기준 모델을 만들고 작은 규모의 버전와 큰 버전의 모델을 만들어 비교하겠습니다.
`layers.Dense`만 사용하는 간단한 기준 모델로 시작한 다음 더 큰 버전을 만들고 서로 비교합니다.
### 기준 모델 만들기
훈련하는 동안 학습률을 점진적으로 낮추면 많은 모델이 더 잘 훈련됩니다. 시간 경과에 따른 학습률을 줄이려면 `optimizers.schedules`를 사용하세요.
```
lr_schedule = tf.keras.optimizers.schedules.InverseTimeDecay(
0.001,
decay_steps=STEPS_PER_EPOCH*1000,
decay_rate=1,
staircase=False)
def get_optimizer():
return tf.keras.optimizers.Adam(lr_schedule)
```
위의 코드는 1000 epoch에서 학습률을 기본 학습률의 1/2, 2000 epoch에서는 1/3 등 쌍곡선 방식으로 줄이도록 `schedules.InverseTimeDecay`를 설정합니다.
```
step = np.linspace(0,100000)
lr = lr_schedule(step)
plt.figure(figsize = (8,6))
plt.plot(step/STEPS_PER_EPOCH, lr)
plt.ylim([0,max(plt.ylim())])
plt.xlabel('Epoch')
_ = plt.ylabel('Learning Rate')
```
이 튜토리얼의 각 모델은 동일한 훈련 구성을 사용합니다. 따라서 콜백 목록부터 시작하여 재사용 가능한 방식으로 설정하세요.
이 튜토리얼의 훈련은 다수의 짧은 epoch 동안 실행됩니다. 로깅 노이즈를 줄이기 위해 각 epoch에 대해 단순히 `.`을 인쇄하고 100개의 epoch마다 전체 메트릭을 인쇄하는 `tfdocs.EpochDots`를 사용합니다.
Next include `callbacks.EarlyStopping` to avoid long and unnecessary training times. Note that this callback is set to monitor the `val_binary_crossentropy`, not the `val_loss`. This difference will be important later.
`callbacks.TensorBoard`를 사용하여 훈련에 대한 TensorBoard 로그를 생성합니다.
```
def get_callbacks(name):
return [
tfdocs.modeling.EpochDots(),
tf.keras.callbacks.EarlyStopping(monitor='val_binary_crossentropy', patience=200),
tf.keras.callbacks.TensorBoard(logdir/name),
]
```
마찬가지로 각 모델은 동일한 `Model.compile` 및 `Model.fit` 설정을 사용합니다.
```
def compile_and_fit(model, name, optimizer=None, max_epochs=10000):
if optimizer is None:
optimizer = get_optimizer()
model.compile(optimizer=optimizer,
loss=tf.keras.losses.BinaryCrossentropy(from_logits=True),
metrics=[
tf.keras.losses.BinaryCrossentropy(
from_logits=True, name='binary_crossentropy'),
'accuracy'])
model.summary()
history = model.fit(
train_ds,
steps_per_epoch = STEPS_PER_EPOCH,
epochs=max_epochs,
validation_data=validate_ds,
callbacks=get_callbacks(name),
verbose=0)
return history
```
### 미소 모델
모델 훈련으로 시작합니다.
```
tiny_model = tf.keras.Sequential([
layers.Dense(16, activation='elu', input_shape=(FEATURES,)),
layers.Dense(1)
])
size_histories = {}
size_histories['Tiny'] = compile_and_fit(tiny_model, 'sizes/Tiny')
```
이제 모델이 어떻게 작동했는지 확인합니다.
```
plotter = tfdocs.plots.HistoryPlotter(metric = 'binary_crossentropy', smoothing_std=10)
plotter.plot(size_histories)
plt.ylim([0.5, 0.7])
```
### 작은 모델
작은 모델의 성능을 능가할 수 있는지 확인하기 위해 일부 큰 모델을 점진적으로 훈련합니다.
각각 16개 단위가 있는 두 개의 숨겨진 레이어를 사용해 봅니다.
```
baseline_model = keras.Sequential([
# `.summary` 메서드 때문에 `input_shape`가 필요합니다
keras.layers.Dense(16, activation='relu', input_shape=(NUM_WORDS,)),
keras.layers.Dense(16, activation='relu'),
keras.layers.Dense(1, activation='sigmoid')
])
baseline_model.compile(optimizer='adam',
loss='binary_crossentropy',
metrics=['accuracy', 'binary_crossentropy'])
baseline_model.summary()
baseline_history = baseline_model.fit(train_data,
train_labels,
epochs=20,
batch_size=512,
validation_data=(test_data, test_labels),
verbose=2)
```
### 작은 모델 만들기
앞서 만든 기준 모델과 비교하기 위해 적은 수의 은닉 유닛을 가진 모델을 만들어 보죠:
```
smaller_model = keras.Sequential([
keras.layers.Dense(4, activation='relu', input_shape=(NUM_WORDS,)),
keras.layers.Dense(4, activation='relu'),
keras.layers.Dense(1, activation='sigmoid')
])
smaller_model.compile(optimizer='adam',
loss='binary_crossentropy',
metrics=['accuracy', 'binary_crossentropy'])
smaller_model.summary()
```
같은 데이터를 사용해 이 모델을 훈련합니다:
```
smaller_history = smaller_model.fit(train_data,
train_labels,
epochs=20,
batch_size=512,
validation_data=(test_data, test_labels),
verbose=2)
```
### 큰 모델 만들기
아주 큰 모델을 만들어 얼마나 빠르게 과대적합이 시작되는지 알아 볼 수 있습니다. 이 문제에 필요한 것보다 훨씬 더 큰 용량을 가진 네트워크를 추가해서 비교해 보죠:
```
bigger_model = keras.models.Sequential([
keras.layers.Dense(512, activation='relu', input_shape=(NUM_WORDS,)),
keras.layers.Dense(512, activation='relu'),
keras.layers.Dense(1, activation='sigmoid')
])
bigger_model.compile(optimizer='adam',
loss='binary_crossentropy',
metrics=['accuracy','binary_crossentropy'])
bigger_model.summary()
```
역시 같은 데이터를 사용해 모델을 훈련합니다:
```
bigger_history = bigger_model.fit(train_data, train_labels,
epochs=20,
batch_size=512,
validation_data=(test_data, test_labels),
verbose=2)
```
### 훈련 손실과 검증 손실 그래프 그리기
실선은 훈련 손실이고 점선은 검증 손실입니다(낮은 검증 손실이 더 좋은 모델입니다). 여기서는 작은 네트워크가 기준 모델보다 더 늦게 과대적합이 시작되었습니다(즉 에포크 4가 아니라 6에서 시작됩니다). 또한 과대적합이 시작되고 훨씬 천천히 성능이 감소합니다.
더 큰 모델을 빌드하면 더 많은 파워가 제공되지만 이 파워가 어떤 이유로 제한되지 않으면 훈련 세트에 쉽게 과대적합될 수 있습니다.
이 예에서는 일반적으로 `"Tiny"` 모델만 과대적합을 완전히 피하고 더 큰 각 모델은 데이터를 더 빠르게 과대적합합니다. `"large"` 모델의 경우 이것이 너무 심각해져서 실제로 어떤 상황이 벌어지는지 보려면 플롯을 로그 스케일로 전환해야 합니다.
검증 메트릭을 플롯하고 이를 훈련 메트릭과 비교하면 이것이 분명해집니다.
- 약간의 차이가 있는 것이 정상입니다.
- 두 메트릭이 같은 방향으로 움직이면 모든 것이 정상입니다.
- 훈련 메트릭이 계속 개선되는 동안 검증 메트릭이 정체되기 시작하면 과대적합에 가까워진 것입니다.
- 검증 메트릭이 잘못된 방향으로 가고 있다면 모델이 확실하게 과대적합된 것입니다.
```
def plot_history(histories, key='binary_crossentropy'):
plt.figure(figsize=(16,10))
for name, history in histories:
val = plt.plot(history.epoch, history.history['val_'+key],
'--', label=name.title()+' Val')
plt.plot(history.epoch, history.history[key], color=val[0].get_color(),
label=name.title()+' Train')
plt.xlabel('Epochs')
plt.ylabel(key.replace('_',' ').title())
plt.legend()
plt.xlim([0,max(history.epoch)])
plot_history([('baseline', baseline_history),
('smaller', smaller_history),
('bigger', bigger_history)])
```
참고: 위의 모든 훈련 실행은 모델이 개선되지 않는다는 것이 분명해지면 훈련을 종료하도록 `callbacks.EarlyStopping`을 사용했습니다.
### TensorBoard에서 보기
이러한 모델은 모두 훈련 중에 TensorBoard 로그를 작성했습니다.
노트북 내에 내장된 TensorBoard 뷰어를 엽니다.
```
#docs_infra: no_execute
# Load the TensorBoard notebook extension
%load_ext tensorboard
# Open an embedded TensorBoard viewer
%tensorboard --logdir {logdir}/sizes
```
[TensorBoard.dev](https://tensorboard.dev/)에서 이 노트북의 [이전 실행 결과](https://tensorboard.dev/experiment/vW7jmmF9TmKmy3rbheMQpw/#scalars&_smoothingWeight=0.97)를 볼 수 있습니다.
TensorBoard.dev는 ML 실험을 호스팅 및 추적하고 모든 사람과 공유하기 위한 관리 환경입니다.
편의를 위해 `<iframe>`에도 포함시켰습니다.
```
display.IFrame(
src="https://tensorboard.dev/experiment/vW7jmmF9TmKmy3rbheMQpw/#scalars&_smoothingWeight=0.97",
width="100%", height="800px")
```
TensorBoard 결과를 공유하려면 다음을 코드 셀에 복사하여 [TensorBoard.dev](https://tensorboard.dev/)에 로그를 업로드할 수 있습니다.
참고: 이 단계에는 Google 계정이 필요합니다.
```
!tensorboard dev upload --logdir {logdir}/sizes
```
주의: 이 명령은 종료되지 않으며, 장기 실험 결과를 지속적으로 업로드하도록 설계되었습니다. 데이터가 업로드되면 노트북 도구의 "실행 중단" 옵션을 사용하여 이를 중지해야 합니다.
## 과대적합을 방지하기 위한 전략
이 섹션의 내용을 시작하기 전에 위의 `"Tiny"` 모델에서 훈련 로그를 복사하여 비교 기준으로 사용합니다.
```
shutil.rmtree(logdir/'regularizers/Tiny', ignore_errors=True)
shutil.copytree(logdir/'sizes/Tiny', logdir/'regularizers/Tiny')
regularizer_histories = {}
regularizer_histories['Tiny'] = size_histories['Tiny']
```
### 가중치를 규제하기
아마도 오캄의 면도날(Occam's Razor) 이론을 들어 보았을 것입니다. 어떤 것을 설명하는 두 가지 방법이 있다면 더 정확한 설명은 최소한의 가정이 필요한 가장 "간단한" 설명일 것입니다. 이는 신경망으로 학습되는 모델에도 적용됩니다. 훈련 데이터와 네트워크 구조가 주어졌을 때 이 데이터를 설명할 수 있는 가중치의 조합(즉, 가능한 모델)은 많습니다. 간단한 모델은 복잡한 것보다 과대적합되는 경향이 작을 것입니다.
여기서 "간단한 모델"은 모델 파라미터의 분포를 봤을 때 엔트로피(entropy)가 작은 모델입니다(또는 앞 절에서 보았듯이 적은 파라미터를 가진 모델입니다). 따라서 과대적합을 완화시키는 일반적인 방법은 가중치가 작은 값을 가지도록 네트워크의 복잡도에 제약을 가하는 것입니다. 이는 가중치 값의 분포를 좀 더 균일하게 만들어 줍니다. 이를 "가중치 규제"(weight regularization)라고 부릅니다. 네트워크의 손실 함수에 큰 가중치에 해당하는 비용을 추가합니다. 이 비용은 두 가지 형태가 있습니다:
- [L1 규제](https://developers.google.com/machine-learning/glossary/#L1_regularization)는 가중치의 절댓값에 비례하는 비용이 추가됩니다(즉, 가중치의 "L1 노름(norm)"을 추가합니다).
- [L2 규제](https://developers.google.com/machine-learning/glossary/#L2_regularization)는 가중치의 제곱에 비례하는 비용이 추가됩니다(즉, 가중치의 "L2 노름"의 제곱을 추가합니다). 신경망에서는 L2 규제를 가중치 감쇠(weight decay)라고도 부릅니다. 이름이 다르지만 혼돈하지 마세요. 가중치 감쇠는 수학적으로 L2 규제와 동일합니다.
L1 정규화는 가중치를 정확히 0으로 푸시하여 희소 모델을 유도합니다. L2 정규화는 가중치 매개변수를 희소하게 만들지 않고 여기에 페널티를 부여하는데, 작은 가중치에 대해서는 페널티가 0이 되기 때문입니다(L2가 더 일반적인 이유 중 하나).
`tf.keras`에서는 가중치 규제 객체를 층의 키워드 매개변수에 전달하여 가중치에 규제를 추가합니다. L2 가중치 규제를 추가해 보죠.
```
l2_model = keras.models.Sequential([
keras.layers.Dense(16, kernel_regularizer=keras.regularizers.l2(0.001),
activation='relu', input_shape=(NUM_WORDS,)),
keras.layers.Dense(16, kernel_regularizer=keras.regularizers.l2(0.001),
activation='relu'),
keras.layers.Dense(1, activation='sigmoid')
])
l2_model.compile(optimizer='adam',
loss='binary_crossentropy',
metrics=['accuracy', 'binary_crossentropy'])
l2_model_history = l2_model.fit(train_data, train_labels,
epochs=20,
batch_size=512,
validation_data=(test_data, test_labels),
verbose=2)
```
`l2(0.001)`는 네트워크의 전체 손실에 층에 있는 가중치 행렬의 모든 값이 `0.001 * weight_coefficient_value**2`만큼 더해진다는 의미입니다. 이런 페널티(penalty)는 훈련할 때만 추가됩니다. 따라서 테스트 단계보다 훈련 단계에서 네트워크 손실이 훨씬 더 클 것입니다.
L2 규제의 효과를 확인해 보죠:
따라서 `L2` 정규화 패널티가 있는 이 동일한 `"Large"` 모델의 성능이 훨씬 더 좋습니다.
```
plot_history([('baseline', baseline_history),
('l2', l2_model_history)])
```
결과에서 보듯이 모델 파라미터의 개수는 같지만 L2 규제를 적용한 모델이 기본 모델보다 과대적합에 훨씬 잘 견디고 있습니다.
#### 더 많은 정보
이러한 종류의 정규화에 대해 주목해야 할 두 가지 중요한 사항이 있습니다.
**첫 번째:** 자체 훈련 루프를 작성하는 경우 모델에 정규화 손실을 요청해야 합니다.
```
result = l2_model(features)
regularization_loss=tf.add_n(l2_model.losses)
```
**두 번째:** 이 구현은 모델의 손실에 가중치 패널티를 추가한 다음 표준 최적화 절차를 적용하는 식으로 작동합니다.
대신 원시 손실에 대해서만 옵티마이저를 실행한 다음, 계산된 단계를 적용하는 동안 옵티마이저가 약간의 가중치 감소를 적용하는 두 번째 접근 방식이 있습니다. 이 "분리된 가중치 감소"는 `optimizers.FTRL` 및 `optimizers.AdamW`와 같은 옵티마이저에서 볼 수 있습니다.
### 드롭아웃 추가하기
드롭아웃(dropout)은 신경망에서 가장 효과적이고 널리 사용하는 규제 기법 중 하나입니다. 토론토(Toronto) 대학의 힌튼(Hinton)과 그의 제자들이 개발했습니다. 드롭아웃을 층에 적용하면 훈련하는 동안 층의 출력 특성을 랜덤하게 끕니다(즉, 0으로 만듭니다). 훈련하는 동안 어떤 입력 샘플에 대해 [0.2, 0.5, 1.3, 0.8, 1.1] 벡터를 출력하는 층이 있다고 가정해 보죠. 드롭아웃을 적용하면 이 벡터에서 몇 개의 원소가 랜덤하게 0이 됩니다. 예를 들면, [0, 0.5, 1.3, 0, 1.1]가 됩니다. "드롭아웃 비율"은 0이 되는 특성의 비율입니다. 보통 0.2에서 0.5 사이를 사용합니다. 테스트 단계에서는 어떤 유닛도 드롭아웃하지 않습니다. 훈련 단계보다 더 많은 유닛이 활성화되기 때문에 균형을 맞추기 위해 층의 출력 값을 드롭아웃 비율만큼 줄입니다.
`tf.keras`에서는 `Dropout` 층을 이용해 네트워크에 드롭아웃을 추가할 수 있습니다. 이 층은 바로 이전 층의 출력에 드롭아웃을 적용합니다.
IMDB 네트워크에 두 개의 `Dropout` 층을 추가하여 과대적합이 얼마나 감소하는지 알아 보겠습니다:
"드롭아웃 비율"은 0이 되는 특성의 비율로, 일반적으로 0.2에서 0.5 사이로 설정됩니다. 테스트 시간에는 어떤 유닛도 드롭아웃되지 않고 대신 레이어의 출력 값이 드롭아웃 비율과 동일한 계수만큼 축소되는데, 이는 훈련 시간에 더 많은 유닛이 활성화된다는 사실과 균형을 맞추기 위해서입니다.
`tf.keras`에서 Dropout 레이어를 통해 네트워크에 드롭아웃을 도입할 수 있습니다. 이 레이어는 직전 레이어의 출력에 적용됩니다.
네트워크에 두 개의 드롭아웃 레이어를 추가하여 과대적합을 줄이는 효과가 얼마나 되는지 살펴보겠습니다.
```
dpt_model = keras.models.Sequential([
keras.layers.Dense(16, activation='relu', input_shape=(NUM_WORDS,)),
keras.layers.Dropout(0.5),
keras.layers.Dense(16, activation='relu'),
keras.layers.Dropout(0.5),
keras.layers.Dense(1, activation='sigmoid')
])
dpt_model.compile(optimizer='adam',
loss='binary_crossentropy',
metrics=['accuracy','binary_crossentropy'])
dpt_model_history = dpt_model.fit(train_data, train_labels,
epochs=20,
batch_size=512,
validation_data=(test_data, test_labels),
verbose=2)
plot_history([('baseline', baseline_history),
('dropout', dpt_model_history)])
```
이 플롯으로부터 이러한 정규화 접근 방식 모두 `"Large"` 모델의 동작을 개선한다는 것이 분명합니다. 그러나 여전히 `"Tiny"` 기준을 넘어서지는 못합니다.
다음으로, 둘 다 함께 시도하고 더 나은지 확인합니다.
### L2 + 드롭아웃 결합
```
combined_model = tf.keras.Sequential([
layers.Dense(512, kernel_regularizer=regularizers.l2(0.0001),
activation='elu', input_shape=(FEATURES,)),
layers.Dropout(0.5),
layers.Dense(512, kernel_regularizer=regularizers.l2(0.0001),
activation='elu'),
layers.Dropout(0.5),
layers.Dense(512, kernel_regularizer=regularizers.l2(0.0001),
activation='elu'),
layers.Dropout(0.5),
layers.Dense(512, kernel_regularizer=regularizers.l2(0.0001),
activation='elu'),
layers.Dropout(0.5),
layers.Dense(1)
])
regularizer_histories['combined'] = compile_and_fit(combined_model, "regularizers/combined")
plotter.plot(regularizer_histories)
plt.ylim([0.5, 0.7])
```
`"Combined"` 정규화가 있는 이 모델은 분명히 지금까지 최고의 모델입니다.
### TensorBoard에서 보기
이러한 모델은 TensorBoard 로그도 기록했습니다.
노트북 내에 내장된 Tensorboard 뷰어를 열려면 코드 셀에 다음 내용을 복사하세요.
```
%tensorboard --logdir {logdir}/regularizers
```
[TensorDoard.dev](https://tensorboard.dev/experiment/fGInKDo8TXes1z7HQku9mw/#scalars&_smoothingWeight=0.97)에서 이 노트북의 [이전 실행 결과](https://tensorboard.dev/)를 볼 수 있습니다.
편의를 위해 `<iframe>`에도 포함시켰습니다.
```
display.IFrame(
src="https://tensorboard.dev/experiment/fGInKDo8TXes1z7HQku9mw/#scalars&_smoothingWeight=0.97",
width = "100%",
height="800px")
```
다음과 함께 업로드되었습니다.
```
!tensorboard dev upload --logdir {logdir}/regularizers
```
## 결론
요약하자면 신경망에서 과대적합을 방지하는 가장 일반적인 방법은 다음과 같습니다.
- 더 많은 훈련 데이터를 얻음
- 네트워크 용량을 줄임
- 가중치 정규화를 추가함
- 드롭아웃을 추가함
이 가이드에서 다루지 않는 두 가지 중요한 접근 방식은 다음과 같습니다.
- 데이터 증강
- 배치 정규화
각 방법은 그 자체로 도움이 될 수 있지만 이를 결합하여 더 큰 효과를 거둘 수 있는 경우가 종종 있다는 점을 기억하기 바랍니다.
| github_jupyter |
# Jupyter Notebook Markdown
1. Headings & List
2. Embedded Code
3. LaTeX Equations
4. Attach Local/Internet Files
5. Tables
Resource: https://jupyter-notebook.readthedocs.io/en/stable/examples/Notebook/Working%20With%20Markdown%20Cells.html
## 1. Heading & List
### Headings:
Heading of a markdown is going to help your audiences focus on the content of a markdown block. We can add headings by starting a line with one (or multiple) ```#``` followed by a space, as in the following example:
# Heading 1
## Heading 1.1
### Heading 1.1.1
#### Heading 1.1.1.1
### List:
It's often the case that we want to list out the content in a markdown block. We can build nested itemized or enumerated lists:
- First Item
- First Item Sublist
- Sub-Item
- Second Item - Not Sublist - Not Sublist
- Third Item
- Fourth Item
Here is another list:
1. This is the first item
1. This is the first sub-item of the first item
2. This is the second sub-tiem of the first item
2. This is the second item
3. This is the last item in the list
### Text Style
1. *There are two ways to type italic text*
- *Use a single asterisk* | ```*text*```
- OR *Use a single underscore* | ```_text_```
2. **There are two ways to type bold text**
- **Use double asterisks** | ```**test**```
- OR **Use double underscores** | ```__text__```
## 2. Embedded Code
We can embed code meant for illustration instead of execution in Python:
``` Python
def f(x):
"""Function f(x) = x^2"""
return x**2
```
We can also write code for other languages:
``` r
f = function(x){
print(x**2)
}
```
## 3. LaTex Equations
We can include mathematical expression both inline and displayed. Inline expression can be added by surrounding the latex code with a single ```$```. Expressions on their own line are surrounded by double ```$$```.
This is the equation I want to show inline with this text: $\sum{(x_i - \bar{x})^2}$.
This is the equation I want to display by itself.
$$\hat{\sigma}^2 = \frac{\sum{(x_i - \bar{x})^2}}{n-1}$$
Alternatively, you can also surround the expression with HTML tags ```\begin{equation}``` and ```\end{equation}```:
``` html
\begin{equation}
\hat{\sigma}^2 = \frac{\sum{(x_i - \bar{x})^2}}{n-1}
\end{equation}
```
## 4. Attach Local / Internet Files
If we have local files in the Notebook directory, we can refer to these files in Markdown cells directly:
``` md

```
Alternatively, we can also use the HTML tag:
``` md
<img src="./images/python.jpg" />
```

We can also reference external file and attach a file to a markdown cell:
```md

```

## Table
It's often case that we want to display some high-level data or description in a tabular format, which we can easily create table in a markdown cell like this:
``` md
| Column 1 | Column 2 | Column 3 |
| :--- | ---: | :---: |
| Norman | Lo | Economics |
| Tokyo | Japan | Location |
| MJ | Basketball | Sport |
```
| Column 1 | Column 2 | Column 3 |
| :--- | ---: | :---: |
| Norman | Lo | Economics |
| Tokyo | Japan | Location |
| MJ | Basketball | Sport |
| github_jupyter |
Load the Data:
- Column 1: Subject ID
- Column 2: Video ID
- Column 3: Attention (Proprietary measure of mental focus)
- Column 4: Mediation (Proprietary measure of calmness)
- Column 5: Raw (Raw EEG signal)
- Column 6: Delta (1-3 Hz of power spectrum)
- Column 7: Theta (4-7 Hz of power spectrum)
- Column 8: Alpha 1 (Lower 8-11 Hz of power spectrum)
- Column 9: Alpha 2 (Higher 8-11 Hz of power spectrum)
- Column 10: Beta 1 (Lower 12-29 Hz of power spectrum)
- Column 11: Beta 2 (Higher 12-29 Hz of power spectrum)
- Column 12: Gamma 1 (Lower 30-100 Hz of power spectrum)
- Column 13: Gamma 2 (Higher 30-100 Hz of power spectrum)
- Column 14: predefined label (whether the subject is expected to be confused)
- Column 15: user-defined label (whether the subject is actually confused)
```
from pandas import read_csv
import matplotlib.pyplot as plt
from matplotlib import pyplot as plt
import numpy as np
np.random.seed = 3
dataset = read_csv('EEGdata.csv', engine = 'python', skipfooter=2)
dataset = dataset.values.astype('float32')
```
#### First we only consider the EGG data:
```
Leave_One_Out = 9
print(dataset.shape)
Train = dataset [(dataset[:,0]!=Leave_One_Out)]
print(Train.shape)
Test = dataset [(dataset[:,0]==Leave_One_Out)]
print(Test.shape)
# Manual Padding:
VideoID = list(set(dataset[:,1]))
SubjectID = list(set(dataset[:,0]))
import numpy as np
A=0
for i in range(len(SubjectID)):
for j in range(len(VideoID)):
Xtemp=dataset[(dataset[:,0]==SubjectID[i]) & (dataset[:,1]==VideoID[j])]
A = max(len(Xtemp[:,14]),A)
print(A)
TrainNumber = 0
Xtrain = {}
ytrain = {}
VideoID = list(set(Train[:,1]))
SubjectID = list(set(Train[:,0]))
for i in range(len(SubjectID)):
for j in range(len(VideoID)):
Xtemp=Train[(Train[:,0]==SubjectID[i]) & (Train[:,1]==VideoID[j])]
Xtrain[TrainNumber]=np.concatenate([(A-len(Xtemp[:,4]))*[0], Xtemp[:,4]],axis=0)
ytrain[TrainNumber]= Xtemp[:,14].mean()
TrainNumber+=1
Xtest = {}
ytest = {}
TestNumber = 0
VideoID = list(set(Test[:,1]))
for i in range(len(VideoID)):
Xtemp=Test[Test[:,1]==VideoID[i]]
Xtest[TestNumber]=np.concatenate([(A-len(Xtemp[:,4]))*[0], Xtemp[:,4]],axis=0)
ytest[TestNumber]= Xtemp[:,14].mean()
TestNumber+=1
```
### LSTM
```
from random import random
from random import randint
from numpy import array
from numpy import zeros
from keras.models import Sequential
from keras.layers import LSTM
from keras.layers import Dense
from keras.layers import Dropout
from keras.layers import Flatten
from keras.layers import TimeDistributed
# configure problem
length = A # length of the signal
NS = 1 #Number of the signal
model = Sequential()
model.add(LSTM(30, input_shape=(length, NS), dropout=0.3))
#model.add(LSTM(30, input_shape=(length, NS),return_sequences=True, dropout=0.3))
#model.add(LSTM(20, dropout=0.3))
model.add(Dense(1, activation='sigmoid'))
model.compile(loss='binary_crossentropy', optimizer='adam', metrics=['acc'])
print(model.summary())
for i in range(len(Xtrain)):
X = np.array(Xtrain[i]).reshape(1,A,1)
y = np.array(int(ytrain[i])).reshape(1,1)
history = model.fit(X, y, epochs=1, verbose = 0)
#odel.reset_states()
# evaluate model
correct = 0
for i in range(len(Xtest)):
X = np.array(Xtest[i]).reshape(1,A,1)
y = np.array(int(ytest[i])).reshape(1,1)
loss, acc = model.evaluate(X, y, verbose=0)
print('Probability: %f, acc: %f' % (model.predict(X), acc*100))
yhat = 1 if model.predict(X)>=0.5 else 0
if yhat == y:
correct += 1
print('Accuracy: %f %%' % ((correct/len(Xtest))*100.0))
```
### ConvNet-LSTM
```
# define the model
from keras.layers.convolutional import Conv1D
from keras.layers.convolutional import MaxPooling1D
from random import random
from random import randint
from numpy import array
from numpy import zeros
from keras.models import Sequential
from keras.layers import LSTM
from keras.layers import Dense
from keras.layers import Flatten
from keras.layers import TimeDistributed
from keras.layers import Bidirectional
model2 = Sequential()
model2.add(TimeDistributed(Conv1D(filters=20, kernel_size=100, padding='same', activation='relu'), input_shape=(1,A,1)))
model2.add(TimeDistributed(MaxPooling1D(pool_size=5)))
#model2.add(TimeDistributed(Conv1D(filters=100, kernel_size=100, padding='same', activation='relu')))
#model2.add(TimeDistributed(MaxPooling1D(pool_size=2)))
model2.add(TimeDistributed(Flatten()))
#model2.add(LSTM(50, return_sequences=True, dropout=0.3))
model2.add(LSTM(100, return_sequences=True, dropout=0.3))
model2.add(LSTM(10, dropout=0.3))
model2.add(Dense(1, activation='sigmoid'))
model2.compile(loss='binary_crossentropy', optimizer='adam', metrics=['acc'])
print(model2.summary())
for i in range(len(Xtrain)):
X = np.array(Xtrain[i]).reshape(1,1,A,1)
y = np.array(int(ytrain[i])).reshape(1,1)
model2.fit(X, y, epochs=1, batch_size=1, verbose = 0)
model2.reset_states()
correct = 0
for i in range(len(Xtest)):
X = np.array(Xtest[i]).reshape(1,1,A,1)
y = np.array(int(ytest[i])).reshape(1,1)
loss, acc = model2.evaluate(X, y, verbose=0)
print('loss: %f, acc: %f' % (loss, acc*100))
yhat = 1 if model2.predict(X)>=0.5 else 0
if yhat == y:
correct += 1
print('Accuracy: %f %%' % ((correct/len(Xtest))*100.0))
```
### Bidirectional LSTM
```
# define the model
from keras.layers.convolutional import Conv1D
from keras.layers.convolutional import MaxPooling1D
from random import random
from random import randint
from numpy import array
from numpy import zeros
from keras.models import Sequential
from keras.layers import LSTM
from keras.layers import Dense
from keras.layers import Flatten
from keras.layers import TimeDistributed
from keras.layers import Bidirectional
model3 = Sequential()
model3.add(Bidirectional(LSTM(10, return_sequences=True), input_shape=(A, 1)))
model3.add(LSTM(20, dropout=0.3))
model3.add(Dense(1, activation='sigmoid'))
model3.compile(loss='binary_crossentropy', optimizer='adam', metrics=['acc'])
print(model3.summary())
for i in range(len(Xtrain)):
X = np.array(Xtrain[i]).reshape(1, A, 1)
y = np.array(int(ytrain[i])).reshape(1,1)
model3.fit(X, y, epochs=1, verbose = 0)
model3.reset_states()
correct = 0
for i in range(len(Xtest)):
X = np.array(Xtest[i]).reshape(1,A,1)
y = np.array(int(ytest[i])).reshape(1,1)
loss, acc = model3.evaluate(X, y, verbose=0)
print('loss: %f, acc: %f' % (loss, acc*100))
yhat = 1 if model3.predict(X)>=0.5 else 0
if yhat == y:
correct += 1
print('Accuracy: %f %%' % ((correct/len(Xtest))*100.0))
```
#### Multivariate LSTM
To consider all the signals, first we need to normalize features (between 0-1):
```
from sklearn.preprocessing import MinMaxScaler
from keras.layers.convolutional import Conv1D
from keras.layers.convolutional import MaxPooling1D
from random import random
from random import randint
from numpy import array
from numpy import zeros
from keras.models import Sequential
from keras.layers import LSTM
from keras.layers import Dense
from keras.layers import Flatten
from keras.layers import TimeDistributed
from keras.layers import Bidirectional
def NormSignal(S, I):
# normalize features
S=S.reshape(-1, 1)
if I not in [0, 1, 13, 14]:
scaler = MinMaxScaler(feature_range=(0, 1))
scaled = scaler.fit_transform(S)
scaled = scaled
else:
scaled = S
return scaled.reshape(-1).tolist()
from pandas import read_csv
import matplotlib.pyplot as plt
from matplotlib import pyplot as plt
import numpy as np
np.random.seed = 7
dataset = read_csv('EEGdata.csv', engine = 'python', skipfooter=2)
dataset = dataset.values.astype('float32')
NormDataG = [NormSignal(dataset[:,i], i) for i in range(0,15)]
NormDataG = np.array(NormDataG)
NormDataG = NormDataG.T
```
##### The input shape will be 144 time steps with 11 features, 100 training sets
```
VideoID = list(set(NormDataG[:,1]))
SubjectID = list(set(NormDataG[:,0]))
import numpy as np
A=0 # length of signal
for i in range(len(SubjectID)):
for j in range(len(VideoID)):
Xtemp=NormDataG[(NormDataG[:,0]==SubjectID[i]) & (NormDataG[:,1]==VideoID[j])]
A = max(len(Xtemp[:,14]),A)
print(A)
# configure problem
length = A # length of the signal
NS = 11 #Number of the signal
model4 = Sequential()
model4.add(LSTM(10, return_sequences=True, input_shape=(length, NS), dropout=0.3))
model4.add(LSTM(20, dropout=0.3))
model4.add(Dense(1, activation='sigmoid'))
model4.compile(loss='binary_crossentropy', optimizer='adam', metrics=['acc'])
print(model4.summary())
Leave_One_Out = 1
Train = NormDataG [(NormDataG[:,0]!=Leave_One_Out)]
Test = NormDataG [(NormDataG[:,0]==Leave_One_Out)]
# Manual Padding to fixed size:
k=0
VideoID = list(set(Train[:,1]))
SubjectID = list(set(Train[:,0]))
for i in range(len(SubjectID)):
for j in range(len(VideoID)):
Xtemp=Train[(Train[:,0]==SubjectID[i]) & (Train[:,1]==VideoID[j])]
z = np.zeros((A-Xtemp.shape[0], 11), dtype=Train.dtype)
Xt=np.concatenate((Xtemp[:,2:13], z), axis=0)
Xt = Xt.reshape(1, A, -1)
yt= Xtemp[:,14].mean().reshape(1, -1)
if k!=0:
Xtrain = np.vstack((Xtrain,Xt))
ytrain = np.vstack((ytrain,yt))
else:
Xtrain=Xt
ytrain=yt
k=1
k=0
VideoID = list(set(Test[:,1]))
for i in range(len(VideoID)):
Xtemp=Test[Test[:,1]==VideoID[i]]
z = np.zeros((A-Xtemp.shape[0], 11), dtype=Train.dtype)
Xt=np.concatenate((Xtemp[:,2:13], z), axis=0)
Xt = Xt.reshape(1, A, -1)
yt= Xtemp[:,14].mean().reshape(1, -1)
if k!=0:
Xtest = np.vstack((Xtest,Xt))
ytest = np.vstack((ytest,yt))
else:
Xtest=Xt
ytest=yt
k=1
print(Xtrain.shape, Xtest.shape, ytraintest)
Xtrain.shape
AAA=np.array([2,2,3])
BBB=[2,20,3]
CCC=AAA.T
print(CCC)
correct = 0
for i in range(len(Xtest)):
X = np.array(Xtest[i]).reshape(1,A,11)
y = np.array(int(ytest[i])).reshape(1,1)
loss, acc = model4.evaluate(X, y, verbose=0)
print('Probability: %f, acc: %f' % ( model4.predict(X), acc*100))
yhat = 1 if model4.predict(X)>=0.5 else 0
if yhat == y:
correct += 1
print('Accuracy: %f %%' % ((correct/len(Xtest))*100.0))
```
## Bidirectional Multovariate LSTM
```
# define the model
from keras.layers.convolutional import Conv1D
from keras.layers.convolutional import MaxPooling1D
from random import random
from random import randint
from numpy import array
from numpy import zeros
from keras.models import Sequential
from keras.layers import LSTM
from keras.layers import Dense
from keras.layers import Flatten
from keras.layers import TimeDistributed
from keras.layers import Bidirectional
model3 = Sequential()
model3.add(Bidirectional(LSTM(10, return_sequences=True), input_shape=(A, 11)))
model3.add(LSTM(20, dropout=0.3))
model3.add(Dense(1, activation='sigmoid'))
model3.compile(loss='binary_crossentropy', optimizer='adam', metrics=['acc'])
print(model3.summary())
BB=NormDataG[NormDataG[:,0]==9]
plt.plot(BB[1100:,2])
plt.show()
```
| github_jupyter |
```
import math
import pandas_datareader as web
import numpy as np
import pandas as pd
from sklearn.preprocessing import MinMaxScaler
from keras.models import Sequential
from keras.layers import Dense, LSTM
import datetime
import os
import plotly.graph_objects as go
import plotly.express as px
from keras.models import Sequential, load_model
from keras.layers import LSTM, Dense, Dropout
def process(comp,n):
company=comp
s=datetime.date(2010,1,1)
e=datetime.date.today() - datetime.timedelta(1)
df = web.DataReader(company, data_source='yahoo', start=s, end=e)
fig = go.Figure(data=[go.Candlestick(x=df.index,
open=df['Open'],
high=df['High'],
low=df['Low'],
close=df['Close'],
name='ohlc' )])
fig.update_layout(xaxis_rangeslider_visible=False,
title=company,
xaxis_title="Date",
yaxis_title="Price",)
print(fig.show)
return fig,df
def closeT(df):
#Create a new dataframe with only the 'Close column
data = df.filter(['Close'])
print(data)
#Convert the dataframe to a numpy array
dataset = data.values
print(dataset)
#Get the number of rows to train the model on
training_data_len = math.ceil( len(dataset) * .8 )
scaler = MinMaxScaler(feature_range=(0,1))
scaled_data = scaler.fit_transform(dataset)
#Create the training data set
#Create the scaled training data set
train_data = scaled_data[0:training_data_len , :]
#Split the data into x_train and y_train data sets
x_train = []
y_train = []
for i in range(60, len(train_data)):
x_train.append(train_data[i-60:i, 0])
y_train.append(train_data[i, 0])
if i<= 61:
print(x_train)
print(y_train)
print()
#Convert the x_train and y_train to numpy arrays
x_train, y_train = np.array(x_train), np.array(y_train)
#Reshape the data
x_train = np.reshape(x_train, (x_train.shape[0], x_train.shape[1], 1))
#Build the LSTM model
model = Sequential()
model.add(LSTM(50, return_sequences=True, input_shape= (x_train.shape[1], 1)))
model.add(LSTM(50, return_sequences= False))
model.add(Dense(25))
model.add(Dense(1))
#Compile the model
model.compile(optimizer='adam', loss='mean_squared_error')
if (not os.path.exists(f'stock_prediction-{ticker}.h5')):
model.fit(x_train, y_train, batch_size=1, epochs=1)
model.save(f'stock_prediction-{ticker}.h5')
def closeT(company,df):
from keras.models import load_model
from keras import backend as K
#Create a new dataframe with only the 'Close column
data = df.filter(['Close'])
#print(data)
#Convert the dataframe to a numpy array
dataset = data.values
#print(dataset)
#Get the number of rows to train the model on
training_data_len = math.ceil( len(dataset) * .8 )
scaler = MinMaxScaler(feature_range=(0,1))
scaled_data = scaler.fit_transform(dataset)
#Create the training data set
#Create the scaled training data set
train_data = scaled_data[0:training_data_len , :]
#Split the data into x_train and y_train data sets
x_train = []
y_train = []
for i in range(60, len(train_data)):
x_train.append(train_data[i-60:i, 0])
y_train.append(train_data[i, 0])
if i<= 61:
print(x_train)
print(y_train)
print()
#Convert the x_train and y_train to numpy arrays
x_train, y_train = np.array(x_train), np.array(y_train)
#Reshape the data
x_train = np.reshape(x_train, (x_train.shape[0], x_train.shape[1], 1))
#Build the LSTM model
model = Sequential()
model.add(LSTM(50, return_sequences=True, input_shape= (x_train.shape[1], 1)))
model.add(LSTM(50, return_sequences= False))
model.add(Dense(25))
model.add(Dense(1))
#Compile the model
model.compile(optimizer='adam', loss='mean_squared_error')
if (not os.path.exists(f'stock_prediction-{company}.h5')):
model.fit(x_train, y_train, batch_size=1, epochs=1)
model.save(f'stock_prediction-{company}.h5')
#Create the testing data set
test_data = scaled_data[training_data_len - 60: , :]
#Create the data sets x_test and y_test
x_test = []
y_test = dataset[training_data_len:, :]
for i in range(60, len(test_data)):
x_test.append(test_data[i-60:i, 0])
#Convert the data to a numpy array
x_test = np.array(x_test)
#Reshape the data
x_test = np.reshape(x_test, (x_test.shape[0], x_test.shape[1], 1 ))
#Get the models predicted price values
K.clear_session()
model1 = load_model(f'stock_prediction-{company}.h5')
predictions = model1.predict(x_test)
predictions = scaler.inverse_transform(predictions)
#Get the root mean squared error (RMSE)
rmse=np.sqrt(np.mean(((predictions- y_test)**2)))
print(rmse)#Plot the data
train = data[:training_data_len]
valid = data[training_data_len:]
valid['Predictions'] = predictions
#Visualize the data
fig = go.Figure()
# Create and style traces
fig.add_trace(go.Scatter(x=df.index,y=df['Close'], name='close' ))
fig.add_trace(go.Scatter(x=valid.index,y=valid['Predictions'],name='Close Predictions'))
#print(fig.show)
#print(valid)
return valid,fig,scaler,model1
def predict(company,n,scaler,model):
e=datetime.date.today()
apple_quote = web.DataReader(company, data_source='yahoo', start='2015-01-01', end=e)
#Create a new dataframe
apple_quote.reset_index(inplace=True)
#print(apple_quote)
new_df = pd.DataFrame(apple_quote[['Date','Close']])
#print(new_df)
#Get teh last 60 day closing price values and convert the dataframe to an array
for z in range(0,n):
new_df1=new_df.filter(['Close'])
tomorrow=new_df['Date'].tail(1)
#print(tomorrow)
for date in tomorrow:
d1=date
#print(tomorrow)
#print(d1)
if(d1.weekday()==4):
dd=3;
else:
dd=1;
last_60_days = new_df1[-60:].values
#print("last_60_days",last_60_days)
#Scale the data to be values between 0 and 1
last_60_days_scaled = scaler.transform(last_60_days)
#Create an empty list
X_test = []
#Append teh past 60 days
X_test.append(last_60_days_scaled)
#Convert the X_test data set to a numpy array
X_test = np.array(X_test)
#Reshape the data
X_test = np.reshape(X_test, (X_test.shape[0], X_test.shape[1], 1))
#Get the predicted scaled price
pred_price = model.predict(X_test)
#undo the scaling
pred_price = scaler.inverse_transform(pred_price)
#print(pred_price)
for i in pred_price:
price=i
for j in i:
p1=j
new_row = {"Date":d1 + datetime.timedelta(days=dd),"Close":p1}
new_df=new_df.append(new_row,ignore_index=True)
new_df2=new_df.tail(n)
#print(new_df)
fig = px.line(new_df, x='Date', y='Close')
#print(fig.show)
return new_df2,fig
from plotly.offline import plot
import json
import plotly
from flask import Flask, flash, redirect, render_template, request, url_for,session
app = Flask(__name__)
@app.route('/')
def forms():
return render_template('/index.html')
@app.route('/index.html')
def index():
return render_template('/index.html')
@app.route('/predict.html')
def cindex():
return render_template('/predict.html')
@app.route('/proof.html')
def pindex():
return render_template('proof.html')
@app.route('/result.html', methods = ['POST', 'GET'])
def result():
#if request.method == 'POST':
company=request.form["company"]
n1=request.form["period"]
n=int(n1)
fig,df=process(company,n)
df1=df.tail(92)
fig_json = json.dumps(fig, cls=plotly.utils.PlotlyJSONEncoder)
valid,fig1,scaler,model=closeT(company,df)
fig_json1 = json.dumps(fig1, cls=plotly.utils.PlotlyJSONEncoder)
new_df2,fig2=predict(company,n,scaler,model)
fig_json2 = json.dumps(fig2, cls=plotly.utils.PlotlyJSONEncoder)
return render_template("/result.html",comp=company,period=n,table_df=[df1.to_html(table_id="myTable",classes=None,border=0,header="true")],tables=[new_df2.to_html(header="true",table_id="myTable1",border=0,index=False)],plot=fig_json,plot1=fig_json1,plot2=fig_json2)
if __name__ == "__main__":
app.run(debug = True,use_reloader=False)
```
| github_jupyter |
# IPython Module Tutorial
This notebook introduces the `utilipy.ipython.imports` module
## Functions:
The main function of this module is `run_imports`, which combines the functionality of both `import_from_file` as well as offers a variety of prepared import options, accessible by keyword, or from an external file by providing the file path. The details are provided below.
<br>
### run_imports()
This function imports prepared files of imports using ipython magic. These files come in two varieties: custom files or files included in utilipy.
for custom files, see `import_from_file`. As just a simple reference guide, there are two arguments `*files`, which are the filepaths for the import files, and `relative` which specifies (by either a boolean or a boolean list of the same length as `*files`) whether the filepaths are relative or absolute paths.
### Prepared Import Files
These can be imported by keyword=True
| Keyword | Function | Description |
| ------- | -------- | ---------- |
| base | base_imports() | a set of standard imports<br> Base: os, sys, time, pdb, warnings, <br>$\qquad$ numpy -> np, scipy, <br>$\qquad$ tqdm_notebook -> tqdm <br>Logging: .LogFile<br>Misc: ObjDict <br>IPython: display, Latex, Markdown, set_trace, <br>$\qquad$ printmd, printMD, printltx, printLaTeX, <br>$\qquad$ configure_matplotlib, <br>$\qquad$ set_autoreload, aimport, <br>$\qquad$ run_imports, import_from_file |
| extended | extended_imports() | some less used standard imports|
| matplotlib | matplotlib_imports() | matplotlib imports |
| astropy | astropy_imports() | astropy related imports |
| galpy | galpy_imports() | galpy related imports |
<br>
if both astropy and matplotlib are imported here, then `run_imports` sets the matplotlib style to `astropy_mpl_style`, made by `astropy`.
<br><br><br>
### import_from_file()
## Custom Import Files
Making your own import file is trivial since it conforms exactly to the standard python import paradigm. Since the imports have been specified in a file separate from the main script, this is equivalent to doing `from custom_imports import *`, where `custom_imports.py` is the file of imports.
Since `import *` does not provide information about what is being imported, it is usefule to include this information in a `print` statement in the import file. An example `custom_imports.py` file, conforming to the standard file format adopted in `utilipy`, is shown below.
When importing from a custom file with `run_imports` or `import_from_file`, it is important to specify whether the file path is absolute or relative to the current jupyter notebook. This can be controlled using the `relative` keyword.
Example `custom_imports.py` file:
```
#!/usr/bin/env python
# -*- coding: utf-8 -*-
# ----------------------------------------------------------------------------
#
# TITLE :
#
# ----------------------------------------------------------------------------
### Docstring and Metadata
"""
"""
__author__ = ""
##############################################################################
### IMPORTS
# +---------------------------------------------------------------------------+
# First Set of Imports
import this
from that import theother
# +---------------------------------------------------------------------------+
# Second Set of Imports
import lorem
from ipsum import dolor
##############################################################################
### Printing Information
print("""custom_imports:
__information about custom imports here__
""")
##############################################################################
### END
```
#
<br><br>
- - -
<br><br>
## Examples
First we import the `imports` module from `utilipy.ipython`.
For most cases this is unnecessary as both `run_imports` and `import_from_file` are accessible directly from `utilipy.ipython`.
```
from utilipy.ipython import imports
```
### run_imports()
```
imports.run_imports(base=True)
```
testing numpy has actually been imported
```
if 'np' in locals():
print('success loading numpy')
```
as an example of the extended imports
```
imports.run_imports('../utilipy/imports/extend_imports.py',
relative='False')
```
<br><br>
- - -
<span style='font-size:40px;font-weight:650'>
END
</span>
| github_jupyter |
<a href="https://colab.research.google.com/github/parekhakhil/pyImageSearch/blob/main/902_keras_regression_cnns.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Keras, Regression, and CNNs
This notebook is associated with the [Keras, Regression, and CNNs](https://www.pyimagesearch.com/2019/01/28/keras-regression-and-cnns/) blog post published on 2019-01-28.
Only the code for the blog post is here. Most codeblocks have a 1:1 relationship with what you find in the blog post with two exceptions: (1) Python classes are not separate files as they are typically organized with PyImageSearch projects, and (2) Command Line Argument parsing is replaced with an `args` dictionary that you can manipulate as needed.
We recommend that you execute (press ▶️) the code block-by-block, as-is, before adjusting parameters and `args` inputs. Once you've verified that the code is working, you are welcome to hack with it and learn from manipulating inputs, settings, and parameters. For more information on using Jupyter and Colab, please refer to these resources:
* [Jupyter Notebook User Interface](https://jupyter-notebook.readthedocs.io/en/stable/notebook.html#notebook-user-interface)
* [Overview of Google Colaboratory Features](https://colab.research.google.com/notebooks/basic_features_overview.ipynb)
Happy hacking!
<hr>
### Download the code zip file
```
!wget https://s3-us-west-2.amazonaws.com/static.pyimagesearch.com/keras-regression-cnns/keras-regression-cnns.zip
!unzip -qq keras-regression-cnns.zip
%cd keras-regression-cnns
```
### Downloading the House Prices Dataset
```
!git clone https://github.com/emanhamed/Houses-dataset
```
## Blog Post Code
### Import Packages
```
# import the necessary packages
from sklearn.preprocessing import LabelBinarizer
from sklearn.preprocessing import MinMaxScaler
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import BatchNormalization
from tensorflow.keras.layers import Conv2D
from tensorflow.keras.layers import MaxPooling2D
from tensorflow.keras.layers import Activation
from tensorflow.keras.layers import Dropout
from tensorflow.keras.layers import Dense
from tensorflow.keras.layers import Flatten
from tensorflow.keras.layers import Input
from tensorflow.keras.models import Model
from tensorflow.keras.optimizers import Adam
from sklearn.model_selection import train_test_split
import pandas as pd
import numpy as np
import argparse
import locale
import glob
import cv2
import os
```
### Loading the house prices image dataset
```
def load_house_attributes(inputPath):
# initialize the list of column names in the CSV file and then
# load it using Pandas
cols = ["bedrooms", "bathrooms", "area", "zipcode", "price"]
df = pd.read_csv(inputPath, sep=" ", header=None, names=cols)
# determine (1) the unique zip codes and (2) the number of data
# points with each zip code
zipcodes = df["zipcode"].value_counts().keys().tolist()
counts = df["zipcode"].value_counts().tolist()
# loop over each of the unique zip codes and their corresponding
# count
for (zipcode, count) in zip(zipcodes, counts):
# the zip code counts for our housing dataset is *extremely*
# unbalanced (some only having 1 or 2 houses per zip code)
# so let's sanitize our data by removing any houses with less
# than 25 houses per zip code
if count < 25:
idxs = df[df["zipcode"] == zipcode].index
df.drop(idxs, inplace=True)
# return the data frame
return df
def load_house_images(df, inputPath):
# initialize our images array (i.e., the house images themselves)
images = []
# loop over the indexes of the houses
for i in df.index.values:
# find the four images for the house and sort the file paths,
# ensuring the four are always in the *same order*
basePath = os.path.sep.join([inputPath, "{}_*".format(i + 1)])
housePaths = sorted(list(glob.glob(basePath)))
# initialize our list of input images along with the output image
# after *combining* the four input images
inputImages = []
outputImage = np.zeros((64, 64, 3), dtype="uint8")
# loop over the input house paths
for housePath in housePaths:
# load the input image, resize it to be 32 32, and then
# update the list of input images
image = cv2.imread(housePath)
image = cv2.resize(image, (32, 32))
inputImages.append(image)
# tile the four input images in the output image such the first
# image goes in the top-right corner, the second image in the
# top-left corner, the third image in the bottom-right corner,
# and the final image in the bottom-left corner
outputImage[0:32, 0:32] = inputImages[0]
outputImage[0:32, 32:64] = inputImages[1]
outputImage[32:64, 32:64] = inputImages[2]
outputImage[32:64, 0:32] = inputImages[3]
# add the tiled image to our set of images the network will be
# trained on
images.append(outputImage)
# return our set of images
return np.array(images)
```
### Using Keras to implement a CNN for regression
```
def create_cnn(width, height, depth, filters=(16, 32, 64), regress=False):
# initialize the input shape and channel dimension, assuming
# TensorFlow/channels-last ordering
inputShape = (height, width, depth)
chanDim = -1
# define the model input
inputs = Input(shape=inputShape)
# loop over the number of filters
for (i, f) in enumerate(filters):
# if this is the first CONV layer then set the input
# appropriately
if i == 0:
x = inputs
# CONV => RELU => BN => POOL
x = Conv2D(f, (3, 3), padding="same")(x)
x = Activation("relu")(x)
x = BatchNormalization(axis=chanDim)(x)
x = MaxPooling2D(pool_size=(2, 2))(x)
# flatten the volume, then FC => RELU => BN => DROPOUT
x = Flatten()(x)
x = Dense(16)(x)
x = Activation("relu")(x)
x = BatchNormalization(axis=chanDim)(x)
x = Dropout(0.5)(x)
# apply another FC layer, this one to match the number of nodes
# coming out of the MLP
x = Dense(4)(x)
x = Activation("relu")(x)
# check to see if the regression node should be added
if regress:
x = Dense(1, activation="linear")(x)
# construct the CNN
model = Model(inputs, x)
# return the CNN
return model
```
### Implementing the regression training script
```
# # construct the argument parser and parse the arguments
# ap = argparse.ArgumentParser()
# ap.add_argument("-d", "--dataset", type=str, required=True,
# help="path to input dataset of house images")
# args = vars(ap.parse_args())
# since we are using Jupyter Notebooks we can replace our argument
# parsing code with *hard coded* arguments and values
args = {
"dataset": "Houses-dataset/Houses Dataset/"
}
# construct the path to the input .txt file that contains information
# on each house in the dataset and then load the dataset
print("[INFO] loading house attributes...")
inputPath = os.path.sep.join([args["dataset"], "HousesInfo.txt"])
df = load_house_attributes(inputPath)
# load the house images and then scale the pixel intensities to the
# range [0, 1]
print("[INFO] loading house images...")
images = load_house_images(df, args["dataset"])
images = images / 255.0
# partition the data into training and testing splits using 75% of
# the data for training and the remaining 25% for testing
split = train_test_split(df, images, test_size=0.25, random_state=42)
(trainAttrX, testAttrX, trainImagesX, testImagesX) = split
# find the largest house price in the training set and use it to
# scale our house prices to the range [0, 1] (will lead to better
# training and convergence)
maxPrice = trainAttrX["price"].max()
trainY = trainAttrX["price"] / maxPrice
testY = testAttrX["price"] / maxPrice
# create our Convolutional Neural Network and then compile the model
# using mean absolute percentage error as our loss, implying that we
# seek to minimize the absolute percentage difference between our
# price *predictions* and the *actual prices*
model = create_cnn(64, 64, 3, regress=True)
opt = Adam(lr=1e-3, decay=1e-3 / 200)
model.compile(loss="mean_absolute_percentage_error", optimizer=opt)
# train the model
print("[INFO] training model...")
model.fit(x=trainImagesX, y=trainY,
validation_data=(testImagesX, testY),
epochs=200, batch_size=8)
# make predictions on the testing data
print("[INFO] predicting house prices...")
preds = model.predict(testImagesX)
# compute the difference between the *predicted* house prices and the
# *actual* house prices, then compute the percentage difference and
# the absolute percentage difference
diff = preds.flatten() - testY
percentDiff = (diff / testY) * 100
absPercentDiff = np.abs(percentDiff)
# compute the mean and standard deviation of the absolute percentage
# difference
mean = np.mean(absPercentDiff)
std = np.std(absPercentDiff)
# finally, show some statistics on our model
locale.setlocale(locale.LC_ALL, "en_US.UTF-8")
print("[INFO] avg. house price: {}, std house price: {}".format(
locale.currency(df["price"].mean(), grouping=True),
locale.currency(df["price"].std(), grouping=True)))
print("[INFO] mean: {:.2f}%, std: {:.2f}%".format(mean, std))
```
For a detailed walkthrough of the concepts and code, be sure to refer to the full tutorial, [*Keras, Regression, and CNNs*](https://www.pyimagesearch.com/2019/01/28/keras-regression-and-cnns/) published on 2019-01-28.
| github_jupyter |
# Swiss VAT Numbers
## Introduction
The function `clean_ch_vat()` cleans a column containing Swiss VAT number (VAT) strings, and standardizes them in a given format. The function `validate_ch_vat()` validates either a single VAT strings, a column of VAT strings or a DataFrame of VAT strings, returning `True` if the value is valid, and `False` otherwise.
VAT strings can be converted to the following formats via the `output_format` parameter:
* `compact`: only number strings without any seperators or whitespace, like "CHE107787577IVA'"
* `standard`: VAT strings with proper whitespace in the proper places, like "CHE-107.787.577 IVA"
Invalid parsing is handled with the `errors` parameter:
* `coerce` (default): invalid parsing will be set to NaN
* `ignore`: invalid parsing will return the input
* `raise`: invalid parsing will raise an exception
The following sections demonstrate the functionality of `clean_ch_vat()` and `validate_ch_vat()`.
### An example dataset containing VAT strings
```
import pandas as pd
import numpy as np
df = pd.DataFrame(
{
"vat": [
'CHE107787577IVA',
'CHE-107.787.578 IVA',
"51824753556",
"51 824 753 556",
"hello",
np.nan,
"NULL"
],
"address": [
"123 Pine Ave.",
"main st",
"1234 west main heights 57033",
"apt 1 789 s maple rd manhattan",
"robie house, 789 north main street",
"(staples center) 1111 S Figueroa St, Los Angeles",
"hello",
]
}
)
df
```
## 1. Default `clean_ch_vat`
By default, `clean_ch_vat` will clean vat strings and output them in the standard format with proper separators.
```
from dataprep.clean import clean_ch_vat
clean_ch_vat(df, column = "vat")
```
## 2. Output formats
This section demonstrates the output parameter.
### `standard` (default)
```
clean_ch_vat(df, column = "vat", output_format="standard")
```
### `compact`
```
clean_ch_vat(df, column = "vat", output_format="compact")
```
## 3. `inplace` parameter
This deletes the given column from the returned DataFrame.
A new column containing cleaned VAT strings is added with a title in the format `"{original title}_clean"`.
```
clean_ch_vat(df, column="vat", inplace=True)
```
## 4. `errors` parameter
### `coerce` (default)
```
clean_ch_vat(df, "vat", errors="coerce")
```
### `ignore`
```
clean_ch_vat(df, "vat", errors="ignore")
```
## 4. `validate_ch_vat()`
`validate_ch_vat()` returns `True` when the input is a valid VAT. Otherwise it returns `False`.
The input of `validate_ch_vat()` can be a string, a Pandas DataSeries, a Dask DataSeries, a Pandas DataFrame and a dask DataFrame.
When the input is a string, a Pandas DataSeries or a Dask DataSeries, user doesn't need to specify a column name to be validated.
When the input is a Pandas DataFrame or a dask DataFrame, user can both specify or not specify a column name to be validated. If user specify the column name, `validate_ch_vat()` only returns the validation result for the specified column. If user doesn't specify the column name, `validate_ch_vat()` returns the validation result for the whole DataFrame.
```
from dataprep.clean import validate_ch_vat
print(validate_ch_vat("CHE107787577IVA"))
print(validate_ch_vat("CHE-107.787.578 IVA"))
print(validate_ch_vat("51824753556"))
print(validate_ch_vat("51 824 753 556"))
print(validate_ch_vat("hello"))
print(validate_ch_vat(np.nan))
print(validate_ch_vat("NULL"))
```
### Series
```
validate_ch_vat(df["vat"])
```
### DataFrame + Specify Column
```
validate_ch_vat(df, column="vat")
```
### Only DataFrame
```
validate_ch_vat(df)
```
| github_jupyter |
## Practice: BiLSTM for PoS Tagging
*This notebook is based on [open-source implementation](https://github.com/bentrevett/pytorch-pos-tagging) of PoS Tagging in PyTorch.*
### Introduction
In this series we'll be building a machine learning model that produces an output for every element in an input sequence, using PyTorch and TorchText. Specifically, we will be inputting a sequence of text and the model will output a part-of-speech (PoS) tag for each token in the input text. This can also be used for named entity recognition (NER), where the output for each token will be what type of entity, if any, the token is.
In this notebook, we'll be implementing a multi-layer bi-directional LSTM (BiLSTM) to predict PoS tags using the Universal Dependencies English Web Treebank (UDPOS) dataset.
### Preparing Data
First, let's import the necessary Python modules.
```
import torch
import torch.nn as nn
import torch.optim as optim
from torchtext import data
from torchtext import datasets
import spacy
import numpy as np
import time
import random
```
Next, we'll set the random seeds for reproducability.
```
SEED = 1234
random.seed(SEED)
np.random.seed(SEED)
torch.manual_seed(SEED)
torch.backends.cudnn.deterministic = True
```
One of the key parts of TorchText is the `Field`. The `Field` handles how your dataset is processed.
Our `TEXT` field handles how the text that we need to tag is dealt with. All we do here is set `lower = True` which lowercases all of the text.
Next we'll define the `Fields` for the tags. This dataset actually has two different sets of tags, [universal dependency (UD) tags](https://universaldependencies.org/u/pos/) and [Penn Treebank (PTB) tags](https://www.sketchengine.eu/penn-treebank-tagset/). We'll only train our model on the UD tags, but will load the PTB tags to show how they could be used instead.
`UD_TAGS` handles how the UD tags should be handled. Our `TEXT` vocabulary - which we'll build later - will have *unknown* tokens in it, i.e. tokens that are not within our vocabulary. However, we won't have unknown tags as we are dealing with a finite set of possible tags. TorchText `Fields` initialize a default unknown token, `<unk>`, which we remove by setting `unk_token = None`.
`PTB_TAGS` does the same as `UD_TAGS`, but handles the PTB tags instead.
```
TEXT = data.Field(lower = True)
UD_TAGS = data.Field(unk_token = None)
PTB_TAGS = data.Field(unk_token = None)
```
We then define `fields`, which handles passing our fields to the dataset.
Note that order matters, if you only wanted to load the PTB tags your field would be:
```
fields = (("text", TEXT), (None, None), ("ptbtags", PTB_TAGS))
```
Where `None` tells TorchText to not load those tags.
```
fields = (("text", TEXT), ("udtags", UD_TAGS), ("ptbtags", PTB_TAGS))
```
Next, we load the UDPOS dataset using our defined fields.
```
train_data, valid_data, test_data = datasets.UDPOS.splits(fields)
```
We can check how many examples are in each section of the dataset by checking their length.
```
print(f"Number of training examples: {len(train_data)}")
print(f"Number of validation examples: {len(valid_data)}")
print(f"Number of testing examples: {len(test_data)}")
```
Let's print out an example:
```
print(vars(train_data.examples[0]))
```
We can also view the text and tags separately:
```
print(vars(train_data.examples[0])['text'])
print(vars(train_data.examples[0])['udtags'])
print(vars(train_data.examples[0])['ptbtags'])
```
Next, we'll build the vocabulary - a mapping of tokens to integers.
We want some unknown tokens within our dataset in order to replicate how this model would be used in real life, so we set the `min_freq` to 2 which means only tokens that appear twice in the training set will be added to the vocabulary and the rest will be replaced by `<unk>` tokens.
We also load the [GloVe](https://nlp.stanford.edu/projects/glove/) pre-trained token embeddings. Specifically, the 100-dimensional embeddings that have been trained on 6 billion tokens. Using pre-trained embeddings usually leads to improved performance - although admittedly the dataset used in this tutorial is too small to take advantage of the pre-trained embeddings.
`unk_init` is used to initialize the token embeddings which are not in the pre-trained embedding vocabulary. By default this sets those embeddings to zeros, however it is better to not have them all initialized to the same value, so we initialize them from a Normal/Gaussian distribution.
These pre-trained vectors are now loaded into our vocabulary and we will initialize our model with these values later.
```
MIN_FREQ = 2
TEXT.build_vocab(train_data,
min_freq = MIN_FREQ,
vectors = "glove.6B.100d",
unk_init = torch.Tensor.normal_)
UD_TAGS.build_vocab(train_data)
PTB_TAGS.build_vocab(train_data)
```
We can check how many tokens and tags are in our vocabulary by getting their length:
```
print(f"Unique tokens in TEXT vocabulary: {len(TEXT.vocab)}")
print(f"Unique tokens in UD_TAG vocabulary: {len(UD_TAGS.vocab)}")
print(f"Unique tokens in PTB_TAG vocabulary: {len(PTB_TAGS.vocab)}")
```
Exploring the vocabulary, we can check the most common tokens within our texts:
```
print(TEXT.vocab.freqs.most_common(20))
```
We can see the vocabularies for both of our tags:
```
print(UD_TAGS.vocab.itos)
print(PTB_TAGS.vocab.itos)
```
We can also see how many of each tag are in our vocabulary:
```
print(UD_TAGS.vocab.freqs.most_common())
print(PTB_TAGS.vocab.freqs.most_common())
```
We can also view how common each of the tags are within the training set:
```
def tag_percentage(tag_counts):
total_count = sum([count for tag, count in tag_counts])
tag_counts_percentages = [(tag, count, count/total_count) for tag, count in tag_counts]
return tag_counts_percentages
print("Tag\t\tCount\t\tPercentage\n")
for tag, count, percent in tag_percentage(UD_TAGS.vocab.freqs.most_common()):
print(f"{tag}\t\t{count}\t\t{percent*100:4.1f}%")
print("Tag\t\tCount\t\tPercentage\n")
for tag, count, percent in tag_percentage(PTB_TAGS.vocab.freqs.most_common()):
print(f"{tag}\t\t{count}\t\t{percent*100:4.1f}%")
```
The final part of data preparation is handling the iterator.
This will be iterated over to return batches of data to process. Here, we set the batch size and the `device` - which is used to place the batches of tensors on our GPU, if we have one.
```
BATCH_SIZE = 128
device = torch.device('cuda:0' if torch.cuda.is_available() else 'cpu')
train_iterator, valid_iterator, test_iterator = data.BucketIterator.splits(
(train_data, valid_data, test_data),
batch_size = BATCH_SIZE,
device = device)
```
## Building the Model
Next up, we define our model - a multi-layer bi-directional LSTM. The image below shows a simplified version of the model with only one LSTM layer and omitting the LSTM's cell state for clarity.

The model takes in a sequence of tokens, $X = \{x_1, x_2,...,x_T\}$, passes them through an embedding layer, $e$, to get the token embeddings, $e(X) = \{e(x_1), e(x_2), ..., e(x_T)\}$.
These embeddings are processed - one per time-step - by the forward and backward LSTMs. The forward LSTM processes the sequence from left-to-right, whilst the backward LSTM processes the sequence right-to-left, i.e. the first input to the forward LSTM is $x_1$ and the first input to the backward LSTM is $x_T$.
The LSTMs also take in the the hidden, $h$, and cell, $c$, states from the previous time-step
$$h^{\rightarrow}_t = \text{LSTM}^{\rightarrow}(e(x^{\rightarrow}_t), h^{\rightarrow}_{t-1}, c^{\rightarrow}_{t-1})$$
$$h^{\leftarrow}_t=\text{LSTM}^{\leftarrow}(e(x^{\leftarrow}_t), h^{\leftarrow}_{t-1}, c^{\leftarrow}_{t-1})$$
After the whole sequence has been processed, the hidden and cell states are then passed to the next layer of the LSTM.
The initial hidden and cell states, $h_0$ and $c_0$, for each direction and layer are initialized to a tensor full of zeros.
We then concatenate both the forward and backward hidden states from the final layer of the LSTM, $H = \{h_1, h_2, ... h_T\}$, where $h_1 = [h^{\rightarrow}_1;h^{\leftarrow}_T]$, $h_2 = [h^{\rightarrow}_2;h^{\leftarrow}_{T-1}]$, etc. and pass them through a linear layer, $f$, which is used to make the prediction of which tag applies to this token, $\hat{y}_t = f(h_t)$.
When training the model, we will compare our predicted tags, $\hat{Y}$ against the actual tags, $Y$, to calculate a loss, the gradients w.r.t. that loss, and then update our parameters.
We implement the model detailed above in the `BiLSTMPOSTagger` class.
`nn.Embedding` is an embedding layer and the input dimension should be the size of the input (text) vocabulary. We tell it what the index of the padding token is so it does not update the padding token's embedding entry.
`nn.LSTM` is the LSTM. We apply dropout as regularization between the layers, if we are using more than one.
`nn.Linear` defines the linear layer to make predictions using the LSTM outputs. We double the size of the input if we are using a bi-directional LSTM. The output dimensions should be the size of the tag vocabulary.
We also define a dropout layer with `nn.Dropout`, which we use in the `forward` method to apply dropout to the embeddings and the outputs of the final layer of the LSTM.
```
class BiLSTMPOSTagger(nn.Module):
def __init__(self,
input_dim,
embedding_dim,
hidden_dim,
output_dim,
n_layers,
bidirectional,
dropout,
pad_idx):
super().__init__()
self.embedding = nn.Embedding(input_dim, embedding_dim, padding_idx = pad_idx)
self.lstm = nn.LSTM(embedding_dim,
hidden_dim,
num_layers = n_layers,
bidirectional = bidirectional,
dropout = dropout if n_layers > 1 else 0)
self.fc = nn.Linear(hidden_dim * 2 if bidirectional else hidden_dim, output_dim)
self.dropout = nn.Dropout(dropout)
def forward(self, text):
#text = [sent len, batch size]
#pass text through embedding layer
embedded = self.dropout(self.embedding(text))
#embedded = [sent len, batch size, emb dim]
#pass embeddings into LSTM
outputs, (hidden, cell) = self.lstm(embedded)
#outputs holds the backward and forward hidden states in the final layer
#hidden and cell are the backward and forward hidden and cell states at the final time-step
#output = [sent len, batch size, hid dim * n directions]
#hidden/cell = [n layers * n directions, batch size, hid dim]
#we use our outputs to make a prediction of what the tag should be
predictions = self.fc(self.dropout(outputs))
#predictions = [sent len, batch size, output dim]
return predictions
```
## Training the Model
Next, we instantiate the model. We need to ensure the embedding dimensions matches that of the GloVe embeddings we loaded earlier.
The rest of the hyperparmeters have been chosen as sensible defaults, though there may be a combination that performs better on this model and dataset.
The input and output dimensions are taken directly from the lengths of the respective vocabularies. The padding index is obtained using the vocabulary and the `Field` of the text.
```
INPUT_DIM = len(TEXT.vocab)
EMBEDDING_DIM = 100
HIDDEN_DIM = 128
OUTPUT_DIM = len(UD_TAGS.vocab)
N_LAYERS = 2
BIDIRECTIONAL = True
DROPOUT = 0.25
PAD_IDX = TEXT.vocab.stoi[TEXT.pad_token]
model = BiLSTMPOSTagger(INPUT_DIM,
EMBEDDING_DIM,
HIDDEN_DIM,
OUTPUT_DIM,
N_LAYERS,
BIDIRECTIONAL,
DROPOUT,
PAD_IDX)
```
We initialize the weights from a simple Normal distribution. Again, there may be a better initialization scheme for this model and dataset.
```
def init_weights(m):
for name, param in m.named_parameters():
nn.init.normal_(param.data, mean = 0, std = 0.1)
model.apply(init_weights)
```
Next, a small function to tell us how many parameters are in our model. Useful for comparing different models.
```
def count_parameters(model):
return sum(p.numel() for p in model.parameters() if p.requires_grad)
print(f'The model has {count_parameters(model):,} trainable parameters')
```
We'll now initialize our model's embedding layer with the pre-trained embedding values we loaded earlier.
This is done by getting them from the vocab's `.vectors` attribute and then performing a `.copy` to overwrite the embedding layer's current weights.
```
pretrained_embeddings = TEXT.vocab.vectors
print(pretrained_embeddings.shape)
model.embedding.weight.data.copy_(pretrained_embeddings)
```
It's common to initialize the embedding of the pad token to all zeros. This, along with setting the `padding_idx` in the model's embedding layer, means that the embedding should always output a tensor full of zeros when a pad token is input.
```
model.embedding.weight.data[PAD_IDX] = torch.zeros(EMBEDDING_DIM)
print(model.embedding.weight.data)
```
We then define our optimizer, used to update our parameters w.r.t. their gradients. We use Adam with the default learning rate.
```
optimizer = optim.Adam(model.parameters())
```
Next, we define our loss function, cross-entropy loss.
Even though we have no `<unk>` tokens within our tag vocab, we still have `<pad>` tokens. This is because all sentences within a batch need to be the same size. However, we don't want to calculate the loss when the target is a `<pad>` token as we aren't training our model to recognize padding tokens.
We handle this by setting the `ignore_index` in our loss function to the index of the padding token in our tag vocabulary.
```
TAG_PAD_IDX = UD_TAGS.vocab.stoi[UD_TAGS.pad_token]
criterion = nn.CrossEntropyLoss(ignore_index = TAG_PAD_IDX)
```
We then place our model and loss function on our GPU, if we have one.
```
model = model.to(device)
criterion = criterion.to(device)
```
We will be using the loss value between our predicted and actual tags to train the network, but ideally we'd like a more interpretable way to see how well our model is doing - accuracy.
The issue is that we don't want to calculate accuracy over the `<pad>` tokens as we aren't interested in predicting them.
The function below only calculates accuracy over non-padded tokens. `non_pad_elements` is a tensor containing the indices of the non-pad tokens within an input batch. We then compare the predictions of those elements with the labels to get a count of how many predictions were correct. We then divide this by the number of non-pad elements to get our accuracy value over the batch.
```
def categorical_accuracy(preds, y, tag_pad_idx):
"""
Returns accuracy per batch, i.e. if you get 8/10 right, this returns 0.8, NOT 8
"""
max_preds = preds.argmax(dim = 1, keepdim = True) # get the index of the max probability
non_pad_elements = (y != tag_pad_idx).nonzero()
correct = max_preds[non_pad_elements].squeeze(1).eq(y[non_pad_elements])
return correct.sum() / torch.FloatTensor([y[non_pad_elements].shape[0]])
```
Next is the function that handles training our model.
We first set the model to `train` mode to turn on dropout/batch-norm/etc. (if used). Then we iterate over our iterator, which returns a batch of examples.
For each batch:
- we zero the gradients over the parameters from the last gradient calculation
- insert the batch of text into the model to get predictions
- as PyTorch loss functions cannot handle 3-dimensional predictions we reshape our predictions
- calculate the loss and accuracy between the predicted tags and actual tags
- call `backward` to calculate the gradients of the parameters w.r.t. the loss
- take an optimizer `step` to update the parameters
- add to the running total of loss and accuracy
```
def train(model, iterator, optimizer, criterion, tag_pad_idx):
epoch_loss = 0
epoch_acc = 0
model.train()
for batch in iterator:
text = batch.text
tags = batch.udtags
optimizer.zero_grad()
#text = [sent len, batch size]
predictions = model(text)
#predictions = [sent len, batch size, output dim]
#tags = [sent len, batch size]
predictions = predictions.view(-1, predictions.shape[-1])
tags = tags.view(-1)
#predictions = [sent len * batch size, output dim]
#tags = [sent len * batch size]
loss = criterion(predictions, tags)
acc = categorical_accuracy(predictions, tags, tag_pad_idx)
loss.backward()
optimizer.step()
epoch_loss += loss.item()
epoch_acc += acc.item()
return epoch_loss / len(iterator), epoch_acc / len(iterator)
```
The `evaluate` function is similar to the `train` function, except with changes made so we don't update the model's parameters.
`model.eval()` is used to put the model in evaluation mode, so dropout/batch-norm/etc. are turned off.
The iteration loop is also wrapped in `torch.no_grad` to ensure we don't calculate any gradients. We also don't need to call `optimizer.zero_grad()` and `optimizer.step()`.
```
def evaluate(model, iterator, criterion, tag_pad_idx):
epoch_loss = 0
epoch_acc = 0
model.eval()
with torch.no_grad():
for batch in iterator:
text = batch.text
tags = batch.udtags
predictions = model(text)
predictions = predictions.view(-1, predictions.shape[-1])
tags = tags.view(-1)
loss = criterion(predictions, tags)
acc = categorical_accuracy(predictions, tags, tag_pad_idx)
epoch_loss += loss.item()
epoch_acc += acc.item()
return epoch_loss / len(iterator), epoch_acc / len(iterator)
```
Next, we have a small function that tells us how long an epoch takes.
```
def epoch_time(start_time, end_time):
elapsed_time = end_time - start_time
elapsed_mins = int(elapsed_time / 60)
elapsed_secs = int(elapsed_time - (elapsed_mins * 60))
return elapsed_mins, elapsed_secs
```
Finally, we train our model!
After each epoch we check if our model has achieved the best validation loss so far. If it has then we save the parameters of this model and we will use these "best" parameters to calculate performance over our test set.
```
N_EPOCHS = 15
best_valid_loss = float('inf')
for epoch in range(N_EPOCHS):
start_time = time.time()
train_loss, train_acc = train(model, train_iterator, optimizer, criterion, TAG_PAD_IDX)
valid_loss, valid_acc = evaluate(model, valid_iterator, criterion, TAG_PAD_IDX)
end_time = time.time()
epoch_mins, epoch_secs = epoch_time(start_time, end_time)
if valid_loss < best_valid_loss:
best_valid_loss = valid_loss
torch.save(model.state_dict(), 'tut1-model.pt')
print(f'Epoch: {epoch+1:02} | Epoch Time: {epoch_mins}m {epoch_secs}s')
print(f'\tTrain Loss: {train_loss:.3f} | Train Acc: {train_acc*100:.2f}%')
print(f'\t Val. Loss: {valid_loss:.3f} | Val. Acc: {valid_acc*100:.2f}%')
```
We then load our "best" parameters and evaluate performance on the test set.
```
model.load_state_dict(torch.load('tut1-model.pt'))
test_loss, test_acc = evaluate(model, test_iterator, criterion, TAG_PAD_IDX)
print(f'Test Loss: {test_loss:.3f} | Test Acc: {test_acc*100:.2f}%')
```
## Inference
88% accuracy looks pretty good, but let's see our model tag some actual sentences.
We define a `tag_sentence` function that will:
- put the model into evaluation mode
- tokenize the sentence with spaCy if it is not a list
- lowercase the tokens if the `Field` did
- numericalize the tokens using the vocabulary
- find out which tokens are not in the vocabulary, i.e. are `<unk>` tokens
- convert the numericalized tokens into a tensor and add a batch dimension
- feed the tensor into the model
- get the predictions over the sentence
- convert the predictions into readable tags
As well as returning the tokens and tags, it also returns which tokens were `<unk>` tokens.
```
def tag_sentence(model, device, sentence, text_field, tag_field):
model.eval()
if isinstance(sentence, str):
nlp = spacy.load('en')
tokens = [token.text for token in nlp(sentence)]
else:
tokens = [token for token in sentence]
if text_field.lower:
tokens = [t.lower() for t in tokens]
numericalized_tokens = [text_field.vocab.stoi[t] for t in tokens]
unk_idx = text_field.vocab.stoi[text_field.unk_token]
unks = [t for t, n in zip(tokens, numericalized_tokens) if n == unk_idx]
token_tensor = torch.LongTensor(numericalized_tokens)
token_tensor = token_tensor.unsqueeze(-1).to(device)
predictions = model(token_tensor)
top_predictions = predictions.argmax(-1)
predicted_tags = [tag_field.vocab.itos[t.item()] for t in top_predictions]
return tokens, predicted_tags, unks
```
We'll get an already tokenized example from the training set and test our model's performance.
```
example_index = 1
sentence = vars(train_data.examples[example_index])['text']
actual_tags = vars(train_data.examples[example_index])['udtags']
print(sentence)
```
We can then use our `tag_sentence` function to get the tags. Notice how the tokens referring to subject of the sentence, the "respected cleric", are both `<unk>` tokens!
```
tokens, pred_tags, unks = tag_sentence(model,
device,
sentence,
TEXT,
UD_TAGS)
print(unks)
```
We can then check how well it did. Surprisingly, it got every token correct, including the two that were unknown tokens!
```
print("Pred. Tag\tActual Tag\tCorrect?\tToken\n")
for token, pred_tag, actual_tag in zip(tokens, pred_tags, actual_tags):
correct = '✔' if pred_tag == actual_tag else '✘'
print(f"{pred_tag}\t\t{actual_tag}\t\t{correct}\t\t{token}")
```
Let's now make up our own sentence and see how well the model does.
Our example sentence below has every token within the model's vocabulary.
```
sentence = 'The Queen will deliver a speech about the conflict in North Korea at 1pm tomorrow.'
tokens, tags, unks = tag_sentence(model,
device,
sentence,
TEXT,
UD_TAGS)
print(unks)
```
Looking at the sentence it seems like it gave sensible tags to every token!
```
print("Pred. Tag\tToken\n")
for token, tag in zip(tokens, tags):
print(f"{tag}\t\t{token}")
```
We've now seen how to implement PoS tagging with PyTorch and TorchText!
The BiLSTM isn't a state-of-the-art model, in terms of performance, but is a strong baseline for PoS tasks and is a good tool to have in your arsenal.
### Going deeper
What if we could combine word-level and char-level approaches?

Actually, we can. Let's use LSTM or GRU to generate embedding for every word on char-level.
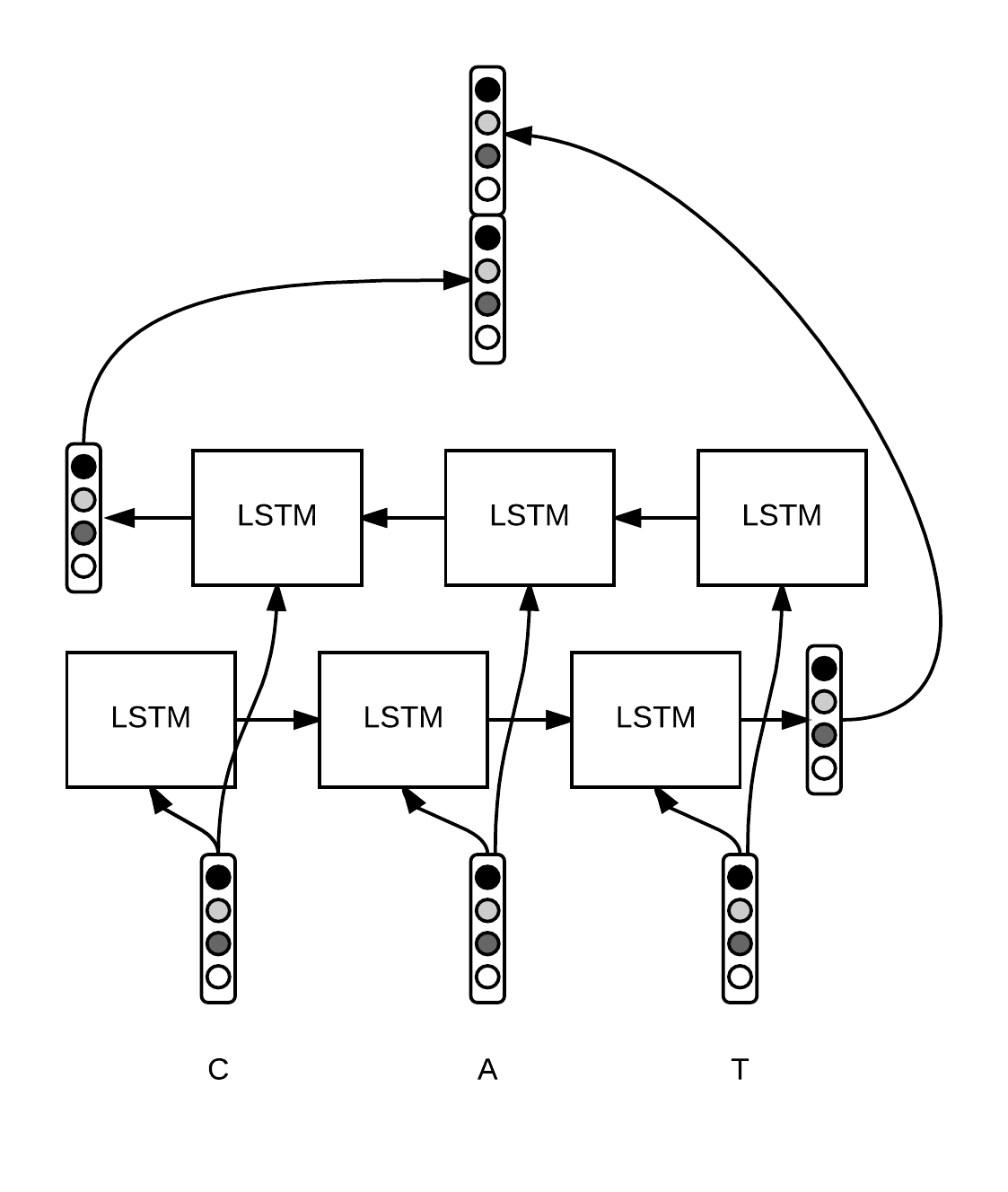
*Image source: https://guillaumegenthial.github.io/sequence-tagging-with-tensorflow.html*
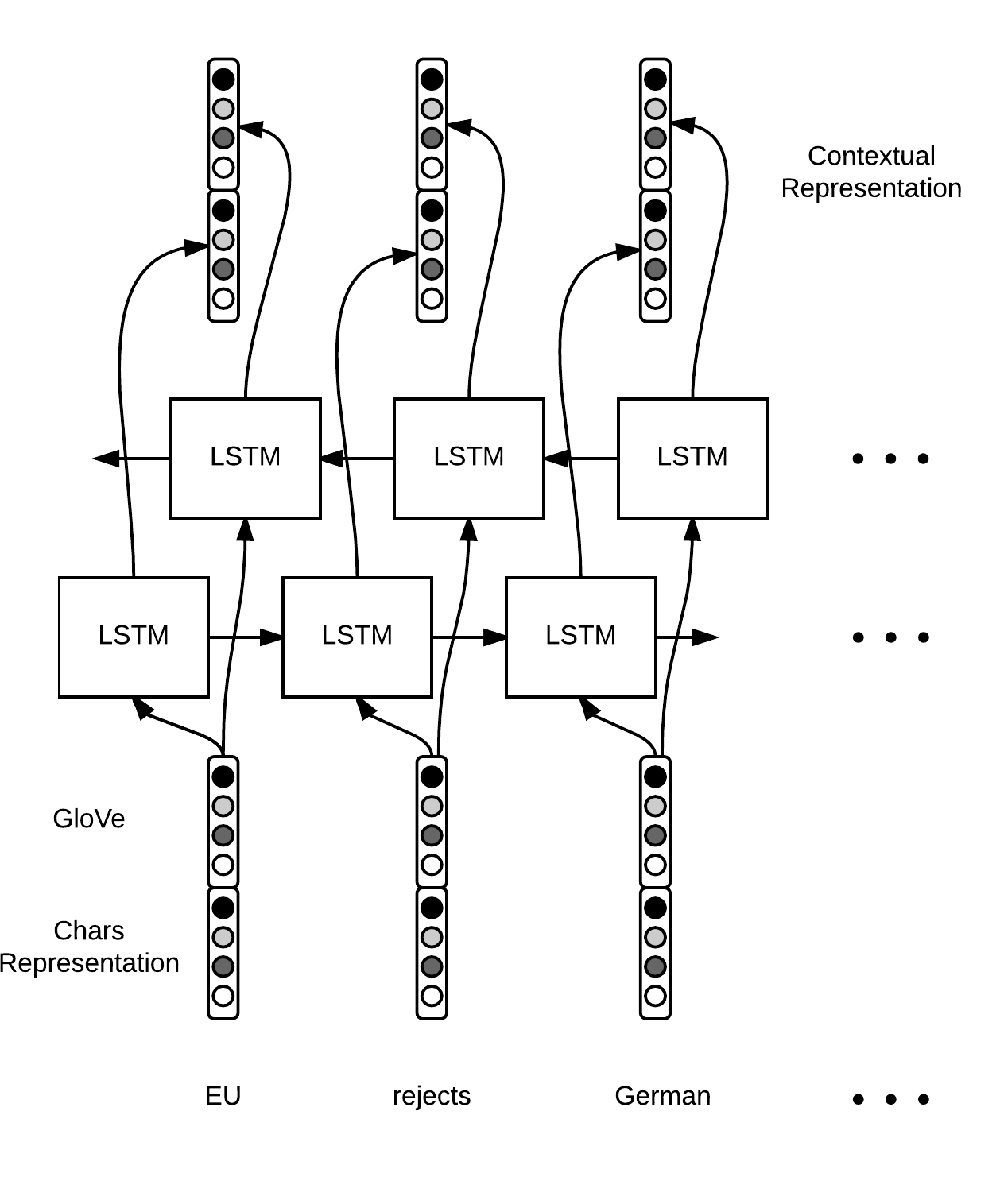
*Image source: https://guillaumegenthial.github.io/sequence-tagging-with-tensorflow.html*
To do that we need to make few adjustments to the code above
```
# Now lets try both word and character embeddings
WORD = data.Field(lower = True)
UD_TAG = data.Field(unk_token = None)
PTB_TAG = data.Field(unk_token = None)
# We'll use NestedField to tokenize each word into list of chars
CHAR_NESTING = data.Field(tokenize=list, init_token="<bos>", eos_token="<eos>")
CHAR = data.NestedField(CHAR_NESTING)#, init_token="<bos>", eos_token="<eos>")
fields = [(('word', 'char'), (WORD, CHAR)), ('udtag', UD_TAG), ('ptbtag', PTB_TAG)]
train_data, valid_data, test_data = datasets.UDPOS.splits(fields)
# train, val, test = datasets.UDPOS.splits(fields=fields)
print(train_data.fields)
print(len(train_data))
print(vars(train_data[0]))
WORD.build_vocab(
train_data,
min_freq = MIN_FREQ,
vectors="glove.6B.100d",
unk_init = torch.Tensor.normal_
)
CHAR.build_vocab(train_data)
UD_TAG.build_vocab(train_data)
PTB_TAG.build_vocab(train_data)
print(f"Unique tokens in WORD vocabulary: {len(WORD.vocab)}")
print(f"Unique tokens in CHAR vocabulary: {len(CHAR.vocab)}")
print(f"Unique tokens in UD_TAG vocabulary: {len(UD_TAG.vocab)}")
print(f"Unique tokens in PTB_TAG vocabulary: {len(PTB_TAG.vocab)}")
BATCH_SIZE = 64
device = torch.device('cuda:0' if torch.cuda.is_available() else 'cpu')
train_iterator, valid_iterator, test_iterator = data.BucketIterator.splits(
(train_data, valid_data, test_data),
batch_size = BATCH_SIZE,
device = device)
batch = next(iter(train_iterator))
text = batch.word
chars = batch.char
tags = batch.udtag
class BiLSTMPOSTaggerWithChars(nn.Module):
def __init__(self,
word_input_dim,
word_embedding_dim,
char_input_dim,
char_embedding_dim,
char_hidden_dim,
hidden_dim,
output_dim,
n_layers,
bidirectional,
dropout,
pad_idx):
super().__init__()
self.char_embedding = # YOUR CODE HERE
self.char_gru = # YOUR CODE HERE
self.word_embedding = nn.Embedding(word_input_dim, word_embedding_dim, padding_idx = pad_idx)
self.lstm = nn.LSTM(word_embedding_dim + # YOUR CODE HERE,
hidden_dim,
num_layers = n_layers,
bidirectional = bidirectional,
dropout = dropout if n_layers > 1 else 0)
self.fc = nn.Linear(hidden_dim * 2 if bidirectional else hidden_dim, output_dim)
self.dropout = nn.Dropout(dropout)
def forward(self, text, chars):
#text = [sent len, batch size]
#pass text through embedding layer
embedded = self.dropout(self.word_embedding(text))
#embedded = [sent len, batch size, emb dim]
chars_embedded = # YOUR CODE HERE
hid_from_chars = # YOUR CODE HERE
embedded_with_chars = torch.cat([embedded, hid_from_chars], dim=2)
#pass embeddings into LSTM
outputs, (hidden, cell) = self.lstm(embedded_with_chars)
# outputs, (hidden, cell) = self.lstm(hid)
#outputs holds the backward and forward hidden states in the final layer
#hidden and cell are the backward and forward hidden and cell states at the final time-step
#output = [sent len, batch size, hid dim * n directions]
#hidden/cell = [n layers * n directions, batch size, hid dim]
#we use our outputs to make a prediction of what the tag should be
predictions = self.fc(self.dropout(outputs))
#predictions = [sent len, batch size, output dim]
return predictions
INPUT_DIM = len(WORD.vocab)
EMBEDDING_DIM = 100
HIDDEN_DIM = 160
CHAR_INPUT_DIM = 112
CHAR_EMBEDDING_DIM = 30
CHAR_HIDDEN_DIM = 30
OUTPUT_DIM = len(UD_TAGS.vocab)
N_LAYERS = 2
BIDIRECTIONAL = True
DROPOUT = 0.25
PAD_IDX = TEXT.vocab.stoi[TEXT.pad_token]
model = BiLSTMPOSTaggerWithChars(
INPUT_DIM,
EMBEDDING_DIM,
CHAR_INPUT_DIM,
CHAR_EMBEDDING_DIM,
CHAR_HIDDEN_DIM,
HIDDEN_DIM,
OUTPUT_DIM,
N_LAYERS,
BIDIRECTIONAL,
DROPOUT,
PAD_IDX
)
```
**Congratulations, you've got LSTM which relies on GRU output on each step.**
Now we need only to train it. Same actions, very small adjustments.
```
def init_weights(m):
for name, param in m.named_parameters():
nn.init.normal_(param.data, mean = 0, std = 0.1)
model.apply(init_weights)
def count_parameters(model):
return sum(p.numel() for p in model.parameters() if p.requires_grad)
print(f'The model has {count_parameters(model):,} trainable parameters')
pretrained_embeddings = TEXT.vocab.vectors
print(pretrained_embeddings.shape)
model.word_embedding.weight.data.copy_(pretrained_embeddings)
model.word_embedding.weight.data[PAD_IDX] = torch.zeros(EMBEDDING_DIM)
print(model.word_embedding.weight.data)
optimizer = optim.Adam(model.parameters())
TAG_PAD_IDX = UD_TAGS.vocab.stoi[UD_TAGS.pad_token]
criterion = nn.CrossEntropyLoss(ignore_index = TAG_PAD_IDX)
model = model.to(device)
criterion = criterion.to(device)
def train(model, iterator, optimizer, criterion, tag_pad_idx):
epoch_loss = 0
epoch_acc = 0
model.train()
for batch in iterator:
text = batch.word
chars = batch.char
tags = batch.udtag
optimizer.zero_grad()
#text = [sent len, batch size]
predictions = model(text, chars)
#predictions = [sent len, batch size, output dim]
#tags = [sent len, batch size]
predictions = predictions.view(-1, predictions.shape[-1])
tags = tags.view(-1)
#predictions = [sent len * batch size, output dim]
#tags = [sent len * batch size]
loss = criterion(predictions, tags)
acc = categorical_accuracy(predictions, tags, tag_pad_idx)
loss.backward()
optimizer.step()
epoch_loss += loss.item()
epoch_acc += acc.item()
return epoch_loss / len(iterator), epoch_acc / len(iterator)
def evaluate(model, iterator, criterion, tag_pad_idx):
epoch_loss = 0
epoch_acc = 0
model.eval()
with torch.no_grad():
for batch in iterator:
text = batch.word
chars = batch.char
tags = batch.udtag
predictions = model(text, chars)
predictions = predictions.view(-1, predictions.shape[-1])
tags = tags.view(-1)
loss = criterion(predictions, tags)
acc = categorical_accuracy(predictions, tags, tag_pad_idx)
epoch_loss += loss.item()
epoch_acc += acc.item()
return epoch_loss / len(iterator), epoch_acc / len(iterator)
N_EPOCHS = 15
best_valid_loss = float('inf')
for epoch in range(N_EPOCHS):
start_time = time.time()
train_loss, train_acc = train(model, train_iterator, optimizer, criterion, TAG_PAD_IDX)
valid_loss, valid_acc = evaluate(model, valid_iterator, criterion, TAG_PAD_IDX)
end_time = time.time()
epoch_mins, epoch_secs = epoch_time(start_time, end_time)
if valid_loss < best_valid_loss:
best_valid_loss = valid_loss
torch.save(model.state_dict(), 'tut2-model.pt')
print(f'Epoch: {epoch+1:02} | Epoch Time: {epoch_mins}m {epoch_secs}s')
print(f'\tTrain Loss: {train_loss:.3f} | Train Acc: {train_acc*100:.2f}%')
print(f'\t Val. Loss: {valid_loss:.3f} | Val. Acc: {valid_acc*100:.2f}%')
# Let's take a look at the model from the last epoch
test_loss, test_acc = evaluate(model, test_iterator, criterion, TAG_PAD_IDX)
print(f'Test Loss: {test_loss:.3f} | Test Acc: {test_acc*100:.2f}%')
# And at the best checkpoint (based on validation score)
model.load_state_dict(torch.load('tut2-model.pt'))
test_loss, test_acc = evaluate(model, test_iterator, criterion, TAG_PAD_IDX)
print(f'Test Loss: {test_loss:.3f} | Test Acc: {test_acc*100:.2f}%')
```
| github_jupyter |
```
import pandera
import fugue
import pandas as pd
import numpy as np
import pyspark
from great_expectations.dataset.sparkdf_dataset import SparkDFDataset
from pyspark.sql import SparkSession
spark = SparkSession \
.builder \
.getOrCreate()
a = np.random.randint(1, 100, 1000)
b = np.random.randint(1, 100, 1000)
test1 = pd.DataFrame({'a':a, 'b':b})
test2 = test1 + 100
test1['partition'] = 'a'
test2['partition'] = 'b'
test = test1.append(test2)
spark_test = spark.createDataFrame(test)
spark_test = spark_test.withColumnRenamed("a","col1")
sparkdf = SparkDFDataset(spark_test)
sparkdf.expect_column_values_to_be_between("col1",
min_value=0,
max_value=9,
mostly=0.95,
result_format="SUMMARY")
# # schema: *, filled:double
# def fillna(df:Iterable[Dict[str,Any]], value:float=0) -> Iterable[Dict[str,Any]]:
# for row in df:
# row["filled"] = (row["value"] or value)
# yield row
# with FugueWorkflow(SparkExecutionEngine) as dag:
# df = dag.load("file.parquet").transform(fillna)
import pandas as pd
df = pd.DataFrame({'State': ['FL','FL','FL','CA','CA','CA'],
'City': ['Tampa', 'Orlando', 'Miami', 'Oakland', 'San Francisco', 'San Jose'],
'Price': [8, 12, 10, 16, 20, 16]})
df.head()
import pandera as pa
price_check = pa.DataFrameSchema({
"Price": pa.Column(pa.Int, pa.Check.in_range(min_value=5,max_value=20)),
})
# schema: *
def price_validation(df:pd.DataFrame) -> pd.DataFrame:
price_check.validate(df)
return df
price_validation(df)
from fugue import FugueWorkflow
from fugue_spark import SparkExecutionEngine
with FugueWorkflow(SparkExecutionEngine) as dag:
df = dag.df(df).transform(price_validation)
df.show()
import pandera as pa
from pandera import Column, Check, DataFrameSchema
price_check = pa.DataFrameSchema({
"price": Column(pa.Float, Check.in_range(min_value=5,max_value=10)),
})
price_check.validate(df)
# schema: *
def price_validation(df:pd.DataFrame) -> pd.DataFrame:
price_check.validate(df)
return df
with FugueWorkflow(SparkExecutionEngine) as dag:
df = df.transform(price_validation)
from great_expectations.dataset.sparkdf_dataset import SparkDFDataset
sparkdf = SparkDFDataset(sparkdf)
sparkdf.expect_column_values_to_be_between("col1",
min_value=0,
max_value=95,
mostly=0.95,
result_format="SUMMARY")
import pandera as pa
from pandera import Column, Check, DataFrameSchema
from fugue import FugueWorkflow
from fugue_spark import SparkExecutionEngine
price_check_FL = pa.DataFrameSchema({
"price": Column(pa.Float, Check.in_range(min_value=7,max_value=13)),
})
price_check_CA = pa.DataFrameSchema({
"price": Column(pa.Float, Check.in_range(min_value=15,max_value=11)),
})
price_checks = {'CA': price_check_CA, 'FL': price_check_FL}
# schema: *
def price_validation(df:pd.DataFrame) -> pd.DataFrame:
location = df['location'].iloc[0]
check = price_checks[location]
check.validate(df)
return df
with FugueWorkflow(SparkExecutionEngine) as dag:
df = dag.df(df).partition(by=["location"]).transform(price_validation)
df.show()
import pandera as pa
from pandera import Column, Check, DataFrameSchema
schema_test1 = pa.DataFrameSchema({
"a": Column(pa.Int, Check.is_b(100)),
})
schema_test2 = pa.DataFrameSchema({
"a": Column(pa.Int, Check.greater_than(99))
})
partition_schema = {"a": schema_test1, "b": schema_test2}
# schema: *
def validator(df:pd.DataFrame) -> pd.DataFrame:
partition = df['partition'].iloc[0]
schema = partition_schema[partition]
schema.validate(df)
return df
from fugue import FugueWorkflow
from fugue_spark import SparkExecutionEngine
with FugueWorkflow(SparkExecutionEngine) as dag:
df = dag.df(test)
df = df.partition(by=["partition"]).transform(validator)
df.show(5)
```
| github_jupyter |
```
import math
import tensorflow as tf
from tensorflow.python.data import Dataset
from sklearn import metrics
import numpy as np
import pandas as pd
# Setup pandas environment
pd.options.display.max_rows = 10
pd.options.display.float_format = '{:.1f}'.format
# Prepare the dataset
california_housing_dataframe = pd.read_csv("https://download.mlcc.google.com/mledu-datasets/california_housing_train.csv", sep=",")
# Shuffle the dataset for better training performance
california_housing_dataframe = california_housing_dataframe.reindex(
np.random.permutation(california_housing_dataframe.index))
california_housing_dataframe["median_house_value"] /= 1000.0
california_housing_dataframe
"""Define Features, Configure Feature Columns and Target
Args:
Define the input feature: total_rooms.
Configure a numeric feature column for total_rooms.
Configure a target or label
Return:
feature_column: total_rooms
target: median_house_value
"""
my_feature = california_housing_dataframe[["total_rooms"]]
feature_columns = [tf.feature_column.numeric_column("total_rooms")]
targets = california_housing_dataframe["median_house_value"]
# Configure the LinearRegressor
# Use gradient descent as the optimizer for training the model.
my_optimizer = tf.train.GradientDescentOptimizer(learning_rate=0.0000001)
my_optimizer = tf.contrib.estimator.clip_gradients_by_norm(my_optimizer, 5.0)
# Configure the linear regression model with our feature columns and optimizer.
# Set a learning rate of 0.0000001 for Gradient Descent.
linear_regressor = tf.estimator.LinearRegressor(
feature_columns=feature_columns,
optimizer=my_optimizer
)
# Defince input function
def input_fn(features, targets, batch_size=1, shuffle=True, num_epochs=None):
"""Trains a linear regression model of one feature.
Args:
features: pandas DataFrame of features
targets: pandas DataFrame of targets
batch_size: Size of batches to be passed to the model
shuffle: True or False. Whether to shuffle the data.
num_epochs: Number of epochs for which data should be repeated. None = repeat indefinitely
Returns:
Tuple of (features, labels) for next data batch
"""
# Convert pandas data into a dict of np arrays.
features = {key:np.array(value) for key,value in dict(features).items()}
# Construct a dataset, and configure batching/repeating.
ds = Dataset.from_tensor_slices((features,targets)) # warning: 2GB limit
ds = ds.batch(batch_size).repeat(num_epochs)
# Shuffle the data, if specified.
if shuffle:
ds = ds.shuffle(buffer_size=10000)
# Return the next batch of data.
features, labels = ds.make_one_shot_iterator().get_next()
return features, labels
_ = linear_regressor.train(
input_fn = lambda: input_fn(my_feature, targets),
steps=100
)
"""Make predictions
Create an input function for predictions.
Note: Since we're making just one prediction for each example, we don't
need to repeat or shuffle the data here.
"""
prediction_input_fn = lambda: input_fn(my_feature, targets, num_epochs=1, shuffle=False)
# Call predict() on the linear_regressor to make predictions.
predictions = linear_regressor.predict(input_fn=prediction_input_fn)
# Format predictions as a NumPy array, so we can calculate error metrics.
predictions = np.array([item['predictions'][0] for item in predictions])
# Print Mean Squared Error and Root Mean Squared Error.
mean_squared_error = metrics.mean_squared_error(predictions, targets)
root_mean_squared_error = math.sqrt(mean_squared_error)
print("Mean Squared Error (on training data): %0.3f" % mean_squared_error)
print("Root Mean Squared Error (on training data): %0.3f" % root_mean_squared_error)
```
| github_jupyter |
<a href="https://qworld.net" target="_blank" align="left"><img src="../qworld/images/header.jpg" align="left"></a>
$ \newcommand{\bra}[1]{\langle #1|} $
$ \newcommand{\ket}[1]{|#1\rangle} $
$ \newcommand{\braket}[2]{\langle #1|#2\rangle} $
$ \newcommand{\dot}[2]{ #1 \cdot #2} $
$ \newcommand{\biginner}[2]{\left\langle #1,#2\right\rangle} $
$ \newcommand{\mymatrix}[2]{\left( \begin{array}{#1} #2\end{array} \right)} $
$ \newcommand{\myvector}[1]{\mymatrix{c}{#1}} $
$ \newcommand{\myrvector}[1]{\mymatrix{r}{#1}} $
$ \newcommand{\mypar}[1]{\left( #1 \right)} $
$ \newcommand{\mybigpar}[1]{ \Big( #1 \Big)} $
$ \newcommand{\sqrttwo}{\frac{1}{\sqrt{2}}} $
$ \newcommand{\dsqrttwo}{\dfrac{1}{\sqrt{2}}} $
$ \newcommand{\onehalf}{\frac{1}{2}} $
$ \newcommand{\donehalf}{\dfrac{1}{2}} $
$ \newcommand{\hadamard}{ \mymatrix{rr}{ \sqrttwo & \sqrttwo \\ \sqrttwo & -\sqrttwo }} $
$ \newcommand{\vzero}{\myvector{1\\0}} $
$ \newcommand{\vone}{\myvector{0\\1}} $
$ \newcommand{\stateplus}{\myvector{ \sqrttwo \\ \sqrttwo } } $
$ \newcommand{\stateminus}{ \myrvector{ \sqrttwo \\ -\sqrttwo } } $
$ \newcommand{\myarray}[2]{ \begin{array}{#1}#2\end{array}} $
$ \newcommand{\X}{ \mymatrix{cc}{0 & 1 \\ 1 & 0} } $
$ \newcommand{\I}{ \mymatrix{rr}{1 & 0 \\ 0 & 1} } $
$ \newcommand{\Z}{ \mymatrix{rr}{1 & 0 \\ 0 & -1} } $
$ \newcommand{\Htwo}{ \mymatrix{rrrr}{ \frac{1}{2} & \frac{1}{2} & \frac{1}{2} & \frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & \frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} & \frac{1}{2} } } $
$ \newcommand{\CNOT}{ \mymatrix{cccc}{1 & 0 & 0 & 0 \\ 0 & 1 & 0 & 0 \\ 0 & 0 & 0 & 1 \\ 0 & 0 & 1 & 0} } $
$ \newcommand{\norm}[1]{ \left\lVert #1 \right\rVert } $
$ \newcommand{\pstate}[1]{ \lceil \mspace{-1mu} #1 \mspace{-1.5mu} \rfloor } $
$ \newcommand{\greenbit}[1] {\mathbf{{\color{green}#1}}} $
$ \newcommand{\bluebit}[1] {\mathbf{{\color{blue}#1}}} $
$ \newcommand{\redbit}[1] {\mathbf{{\color{red}#1}}} $
$ \newcommand{\brownbit}[1] {\mathbf{{\color{brown}#1}}} $
$ \newcommand{\blackbit}[1] {\mathbf{{\color{black}#1}}} $
<font style="font-size:28px;" align="left"><b> <font color="blue"> Solutions for </font>Two Qubits </b></font>
<br>
_prepared by Abuzer Yakaryilmaz_
<br><br>
<a id="task3"></a>
<h3> Task 3 </h3>
We define a quantum circuit with two qubits: $ q_0 $ and $ q_1 $. They are tensored as $ q_1 \otimes q_0 $ in Qiskit.
We apply the Hadamard operator to $q_1$.
```
from qiskit import QuantumCircuit
qc = QuantumCircuit(2)
qc.h(1)
display(qc.draw(output='mpl',reverse_bits=True))
```
Then, the quantum operator applied to both qubits will be $ H \otimes I $.
Read the quantum operator of the above circuit by using 'unitary_simulator' and then verify that it is $ H \otimes I $.
<h3> Solution </h3>
$ H \otimes I = \hadamard \otimes \I = \mymatrix{c|c}{ \sqrttwo \I & \sqrttwo \I \\ \hline \sqrttwo \I & -\sqrttwo \I } = \mymatrix{rr|rr} { \sqrttwo & 0 & \sqrttwo & 0 \\ 0 & \sqrttwo & 0 & \sqrttwo \\ \hline \sqrttwo & 0 & -\sqrttwo & 0 \\ 0 & \sqrttwo & 0 & -\sqrttwo } $
```
from qiskit import execute, Aer
job = execute(qc, Aer.get_backend('unitary_simulator'),shots=1,optimization_level=0)
current_unitary = job.result().get_unitary(qc, decimals=3)
for row in current_unitary:
column = ""
for entry in row:
column = column + str(round(entry.real,3)) + " "
print(column)
```
<a id="task5"></a>
<h3> Task 5 </h3>
Create a quantum curcuit with $ n=5 $ qubits.
Set each qubit to $ \ket{1} $.
Repeat 4 times:
<ul>
<li>Randomly pick a pair of qubits, and apply cx-gate (CNOT operator) on the pair.</li>
</ul>
Draw your circuit, and execute your program 100 times.
Verify your measurement results by checking the diagram of the circuit.
<h3> Solution </h3>
```
# import all necessary objects and methods for quantum circuits
from qiskit import QuantumRegister, ClassicalRegister, QuantumCircuit, execute, Aer
# import randrange for random choices
from random import randrange
n = 5
m = 4
states_of_qubits = [] # we trace the state of each qubit also by ourselves
q = QuantumRegister(n,"q") # quantum register with n qubits
c = ClassicalRegister(n,"c") # classical register with n bits
qc = QuantumCircuit(q,c) # quantum circuit with quantum and classical registers
# set each qubit to |1>
for i in range(n):
qc.x(q[i]) # apply x-gate (NOT operator)
states_of_qubits.append(1) # the state of each qubit is set to 1
# randomly pick m pairs of qubits
for i in range(m):
controller_qubit = randrange(n)
target_qubit = randrange(n)
# controller and target qubits should be different
while controller_qubit == target_qubit: # if they are the same, we pick the target_qubit again
target_qubit = randrange(n)
# print our picked qubits
print("the indices of the controller and target qubits are",controller_qubit,target_qubit)
# apply cx-gate (CNOT operator)
qc.cx(q[controller_qubit],q[target_qubit])
# we also trace the results
if states_of_qubits[controller_qubit] == 1: # if the value of the controller qubit is 1,
states_of_qubits[target_qubit] = 1 - states_of_qubits[target_qubit] # then flips the value of the target qubit
# remark that 1-x gives the negation of x
# measure the quantum register
qc.barrier()
qc.measure(q,c)
# draw the circuit in reading order
display(qc.draw(output='mpl',reverse_bits=True))
# execute the circuit 100 times in the local simulator
job = execute(qc,Aer.get_backend('qasm_simulator'),shots=100)
counts = job.result().get_counts(qc)
print("the measurument result is",counts)
our_result=""
for state in states_of_qubits:
our_result = str(state) + our_result
print("our result is",our_result)
```
<a id="task6"></a>
<h3>Task 6</h3>
Our task is to learn the behavior of the following quantum circuit by doing experiments.
Our circuit has two qubits: $ q_0 $ and $ q_1 $. They are tensored as $ q_1 \otimes q_0 $ in Qiskit.
<ul>
<li> Apply Hadamard to the both qubits.
<li> Apply CNOT($q_1$,$q_0$).
<li> Apply Hadamard to the both qubits.
<li> Measure the circuit.
</ul>
Iteratively initialize the qubits to $ \ket{00} $, $ \ket{01} $, $ \ket{10} $, and $ \ket{11} $.
Execute your program 100 times for each iteration, and then check the outcomes for each iteration.
Observe that the overall circuit implements CNOT($q_0$,$q_1$).
<h3> Solution </h3>
```
# import all necessary objects and methods for quantum circuits
from qiskit import QuantumRegister, ClassicalRegister, QuantumCircuit, execute, Aer
all_inputs=['00','01','10','11']
for input in all_inputs:
q = QuantumRegister(2,"q") # quantum register with 2 qubits
c = ClassicalRegister(2,"c") # classical register with 2 bits
qc = QuantumCircuit(q,c) # quantum circuit with quantum and classical registers
# initialize the inputs w.r.t the reading order of Qiskit
if input[0]=='1':
qc.x(q[1]) # set the state of the up qubit to |1>
if input[1]=='1':
qc.x(q[0]) # set the state of the down qubit to |1>
# apply h-gate to both qubits
qc.h(q[0])
qc.h(q[1])
# apply cx(up-qubit,down-qubit)
qc.cx(q[1],q[0])
# apply h-gate to both qubits
qc.h(q[0])
qc.h(q[1])
# measure both qubits
qc.barrier()
qc.measure(q,c)
# draw the circuit w.r.t the reading order of Qiskit
display(qc.draw(output='mpl',reverse_bits=True))
# execute the circuit 100 times in the local simulator
job = execute(qc,Aer.get_backend('qasm_simulator'),shots=100)
counts = job.result().get_counts(qc)
print(input,"is mapped to",counts)
```
<a id="task7"></a>
<h3>Task 7</h3>
Our task is to learn the behavior of the following quantum circuit by doing experiments.
Our circuit has two qubits: $ q_0 $ and $ q_1 $. They are tensored as $ q_1 \otimes q_0 $ in Qiskit.
<ul>
<li> Apply CNOT($q_1$,$q_0$).
<li> Apply CNOT($q_0$,$q_1$).
<li> Apply CNOT($q_1$,$q_0$).
</ul>
Iteratively initialize the qubits to $ \ket{00} $, $ \ket{01} $, $ \ket{10} $, and $ \ket{11} $.
Execute your program 100 times for each iteration, and then check the outcomes for each iteration.
Observe that the overall circuit swaps the values of the first and second qubits:
<ul>
<li> $\ket{00} \rightarrow \ket{00} $ </li>
<li> $\ket{01} \rightarrow \ket{10} $ </li>
<li> $\ket{10} \rightarrow \ket{01} $ </li>
<li> $\ket{11} \rightarrow \ket{11} $ </li>
</ul>
<h3> Solution </h3>
```
# import all necessary objects and methods for quantum circuits
from qiskit import QuantumRegister, ClassicalRegister, QuantumCircuit, execute, Aer
all_inputs=['00','01','10','11']
for input in all_inputs:
q = QuantumRegister(2,"q") # quantum register with 2 qubits
c = ClassicalRegister(2,"c") # classical register with 2 bits
qc = QuantumCircuit(q,c) # quantum circuit with quantum and classical registers
#initialize the inputs w.r.t the reading order of Qiskit
if input[0]=='1':
qc.x(q[1]) # set the state of the up qubit to |1>
if input[1]=='1':
qc.x(q[0]) # set the state of the down qubit to |1>
# apply cx(up-qubit,down-qubit)
qc.cx(q[1],q[0])
# apply cx(down-qubit,up-qubit)
qc.cx(q[0],q[1])
# apply cx(up-qubit,down-qubit)
qc.cx(q[1],q[0])
# measure both qubits
qc.barrier()
qc.measure(q,c)
# draw the circuit w.r.t the reading order of Qiskit
qc.draw(output='mpl',reverse_bits=True)
# execute the circuit 100 times in the local simulator
job = execute(qc,Aer.get_backend('qasm_simulator'),shots=100)
counts = job.result().get_counts(qc)
print(input,"is mapped to",counts)
```
| github_jupyter |
```
%matplotlib inline
```
# Tutorial 5: Colors and colorbars
This tutorial demonstrates how to configure the colorbar(s) with ``surfplot``.
## Layer color maps and colorbars
The color map can be specified for each added plotting layer using the `cmap`
parameter of :func:`~surfplot.plotting.Plot.add_layer`, along with the
associated ``matplotlib`` colorbar drawn if specified. The colobar can be
turned off by `cbar=False`. The range of the colormap is specified with the
`color_range` parameter, which takes a tuple of (`minimum`, `maximum`) values.
If no color range is specified (the default, i.e. `None`), then the color range
is computed automically based on the minimum and maximum of the data.
Let's get started by setting up a plot with surface shading added as well.
Following the first initial steps of
`sphx_glr_auto_examples_plot_tutorial_01.py` :
```
from neuromaps.datasets import fetch_fslr
from surfplot import Plot
surfaces = fetch_fslr()
lh, rh = surfaces['inflated']
p = Plot(lh, rh)
sulc_lh, sulc_rh = surfaces['sulc']
p.add_layer({'left': sulc_lh, 'right': sulc_rh}, cmap='binary_r', cbar=False)
```
Now let's add a plotting layer with a colorbar using the example data. The
`cmap` parameter accepts any named `matplotlib colormap`_, or a
`colormap object`_. This means that ``surfplot`` can work with pretty much
any colormap, including those from `seaborn`_ and `cmasher`_, for example.
```
from surfplot.datasets import load_example_data
# default mode network associations
default = load_example_data(join=True)
p.add_layer(default, cmap='GnBu_r', cbar_label='Default mode')
fig = p.build()
fig.show()
```
`cbar_label` added a text label to the colorbar. Although not necessary in
cases where a single layer/colorbar is shown, it can be useful when adding
multiple layers. To demonstrate that, let's add another layer using the
`frontoparietal` network associations from
:func:`~surfplot.datasets.load_example_data`:
```
fronto = load_example_data('frontoparietal', join=True)
p.add_layer(fronto, cmap='YlOrBr_r', cbar_label='Frontoparietal')
fig = p.build()
fig.show()
```
The order of the colorbars is always based on the order of the layers, where
the outermost colorbar is the last (i.e. uppermost) plotting layer. Of
course, more layers and colorbars can lead to busy-looking figure, so be sure
not to overdo it.
## cbar_kws
Once all layers have been added, the positioning and style can be adjusted
using the `cbar_kws` parameter in :func:`~surfplot.plotting.Plot.build`,
which are keyword arguments for :func:`surfplot.plotting.Plot._add_colorbars`.
Each one is briefly described below (see :func:`~surfplot.plotting.Plot._add_colorbars`
for more detail):
1. `location`: The location, relative to the surface plot
2. `label_direction`: Angle to draw label for colorbars
3. `n_ticks`: Number of ticks to include on colorbar
4. `decimals`: Number of decimals to show for colorbal tick values
5. `fontsize`: Font size for colorbar labels and tick labels
6. `draw_border`: Draw ticks and black border around colorbar
7. `outer_labels_only`: Show tick labels for only the outermost colorbar
8. `aspect`: Ratio of long to short dimensions
9. `pad`: Space that separates each colorbar
10. `shrink`: Fraction by which to multiply the size of the colorbar
11. `fraction`: Fraction of original axes to use for colorbar
Let's plot colorbars on the right, which will generate vertical colorbars
instead of horizontal colorbars. We'll also add some style changes for a
cleaner look:
```
kws = {'location': 'right', 'label_direction': 45, 'decimals': 1,
'fontsize': 8, 'n_ticks': 2, 'shrink': .15, 'aspect': 8,
'draw_border': False}
fig = p.build(cbar_kws=kws)
fig.show()
# sphinx_gallery_thumbnail_number = 3
```
Be sure to check out `sphx_glr_auto_examples_examples_plot_example_01.py`
for another example of colorbar styling.
| github_jupyter |
# OMF.v2 Block Model Storage
**Authors:** Rowan Cockett, Franklin Koch <br>
**Company:** Seequent <br>
**Date:** March 3, 2019
## Overview
The proposal below defines a storage algorithm for all sub block model formats in OMF.v2.
The storage & access algorithm is based on [sparse matrix/array storage](https://en.wikipedia.org/wiki/Sparse_matrix#Storing_a_sparse_matrix) in linear algebra.
The algorithm for the _Compressed Block Index_ format is largely similar between the various block model formats supported by OMF.v2:
* _Regular Block Model_: No aditional storage information necessary.
* _Tensor Block Model_: No aditional storage information necessary.
* _Regular Sub Block Model_: Single storage array required to record sub-blocking and provide indexing into attribute arrays.
* _Octree Sub Block Model_: Storage array required as well as storage for each Z-Order Curve per octree (discussed in detail below).
* _Arbitrary Sub Block Model_: Storage array required as well as storage of sub-block centroids and sizes.
## Summary
* The compression format for a _Regular Sub Block Model_ scales with parent block count rather than sub block count.
* Storing an _Octree Sub Block Model_ is 12 times more efficient than an _Arbitrary Sub Block Model_ for the same structure. For example, an _Octree Sub Block Model_ with 10M sub-blocks would save 3.52 GB of space.
* Attributes for all sub-block types are stored on-disk in contiguous chunks per parent block, allowing for easy memory mapping of attributes, if necessary.
```
import cbi
import cbi_plot
import z_order_utils
import numpy as np
%matplotlib inline
```
# Compressed Block Index
The _Compressed Block Index_ format (or `cbi` in code) is a monotonically increasing integer array, which starts at 0 (`cbi[0] := 0`) and ends at the total number of blocks `cbi[i * j * k] := num_blocks`. For the n-th parent block, `cbi[n+1] - cbi[n]` will always be the number of sub-blocks per parent (`prod(sub_block_count)` for a _Regular Sub Block Model_). This can be used to determine if the n-th parent block is sub-blocked (i.e. `is_sub_blocked[n]`), as well as the index into any attribute array to retrive all of the attribute data for that parent block. That is, `attribute[cbi[n] : cbi[n+1]]` will always return the attributes for the n-th parent block, regardless of if the parent block is sub-blocked or not. The `cbi` indexing is also useful for the Octree and Arbitrary Sub Block Models, allowing additional topology information about the tree structure or arbitrary sub-blocks, respectively, to be stored in a single array.
The _Compressed Block Index_ format means the total size for storage is a fixed length `UInt32` array plus a small amount of metadata (i.e. nine extra numbers, name, description, etc.). That is, this compression format **scales with the parent block count** rather than the sub-block count. All other information can be derived from the `cbi` array (e.g. `is_sub_blocked` as a boolean and all indexing into the attribute arrays). **Note:** `cbi` could instead use Int64, for the index depending on the upper limit required for number of blocks.
The technique is to be used as underlying storage for _Regular Sub Block Model_, _Octree Sub Block Model_, and _Arbitrary Sub Block Model_. This index is not required for _Tensor Block Model_ or _Regular Block Model_; however, it could be used as an optional property to have null-blocks (e.g. above the topography) that would decrease the storage of all array attributes. In this case, `cbi[n] == cbi[n+1]`.
**Note** - For the final implementation, we may store _compressed block count_, so like `[1, 1, 32, 1]` instead of `[0, 1, 2, 34, 35]`, and _compressed block index_ is a computed sum. This has slight performance advantages, i.e. refining a parent block into sub-blocks only is O(1) rather than O(n), and storage size advantages, since we can likely use `UInt16`, constrained by number of sub-blocks per parent, not total sub-blocks.
# All Block Models
All block models have been decided to be defined inside of a rotated coordinate frame. The implementation of this orientation uses three `axis` vectors (named `axis_u`, `axis_v` and `axis_w`) and a `corner` that defines a bounding box in the project coordinate reference system. These axis vectors must be orthogonal but are not opinionated about "handed-ness". The implementation is explicitly not (a) rotation matrices, which may have skew and or (b) defined as three rotations, which may be applied in other orders (e.g. `ZYX` vs `YXZ`) and not be consistent. The unwrapping of attributes and the `ijk` block index is relative to these axis, respectively, in the rotated coordinate frame. By convention, the `axis` vectors are normalized since their length does not have meaning. Total size of the block model is determined by summing parent block sizes on each dimension. However, it is not absolutely necessary for normalized lengths to be enforced by OMF.
**Stored Properties**
* `name` - Name of the block model
* `description` - Description of the block model
* `attributes` - list of standard [OMF.v1 attributes](https://omf.readthedocs.io/en/stable/content/data.html#scalardata)
* `axis_u` (Vector3) Orientation of the i-axis in the project CRS
* `axis_v` (Vector3) Orientation of the j-axis in the project CRS
* `axis_w` (Vector3) Orientation of the k-axis in the project CRS
* `corner` (Vector3) Minimum x/y/z in the project CRS
* `location` - String representation of where attributes are defiend on the block model. Either `"parent_blocks"` or `"sub_blocks"` (if sub blocks are present in the block model class). This could be extended to `"faces"`, `"edges"`, and `"Nodes"` for Regular and Tensor Block Models
**Attributes**
All block models are stored with flat attribute arrays, allowing for efficient storage and access, as well as adhering to existing standards set out for all other Elements in the OMF.v1 format. The standard counting is _column major ordering_, following "Fortran" style indexing -- in `numpy` (Python) this uses `array.flatten(order='F')` where array is the 3D attribute array. To be explicit, inside a for-loop the `i` index always moves the fastest:
```python
count = regular_block_model.block_count
index = 0
for k in range(count[2]):
for j in range(count[1]):
for i in range(count[0]):
print(index, (i, j, k))
index += 1
```
# Regular Block Model
**Stored Properties**
* `block_size`: a Vector3 (Float) that describes how large each block is
* `block_count`: a Vector3 (Int) that describes how many blocks in each dimension
**Note**
For the final implementation we will use property names `size_blocks`/`size_parent_blocks`/`size_sub_blocks` and equivalently `num_*` - this enables slightly easier discoverability of properties across different element types.
```
rbm = cbi.RegularBlockModel()
rbm.block_size = [1.5, 2.5, 10.]
rbm.block_count = [3, 2, 1]
rbm.validate()
cbi_plot.plot_rbm(rbm)
```
# Tensor Block Model
**Stored Properties**
* `tensor_u`: a Float64 array of spacings along `axis_u`
* `tensor_v`: a Float64 array of spacings along `axis_v`
* `tensor_w`: a Float64 array of spacings along `axis_w`
**Note:** `block_size[0]` for the i-th block is `tensor_u[i]` and `block_count[0]` is `len(tensor_u)`. Counting for attributes is the same as _Regular Block Model_.
```
tbm = cbi.TensorBlockModel()
tbm.tensor_u = [2.5, 1.0, 1.0]
tbm.tensor_v = [3.5, 1.5]
tbm.tensor_w = [10.0]
tbm.validate()
print("block_count:", tbm.block_count)
print("num_blocks:", tbm.num_blocks)
cbi_plot.plot_tbm(tbm)
```
# Regular Sub Block Model
The `RegularSubBlockModel` requires storage of information to store the parent and sub block counts as well as the parent block sizes. Attribute ordering for sub-blocks within each parent block is also _column-major ordering_.
**Stored Properties**
* `parent_block_size`: a Vector3 (Float) that describes how large each parent block is
* `parent_block_count`: a Vector3 (Int) that describes how many parent blocks in each dimension
* `sub_block_count`: a Vector3 (Int) that describes how many sub blocks in each dimension are contained within each parent block
* `compressed_block_index`: a UInt32 array of length (`i * j * k + 1`) that defines the sub block topology
```
rsbm = cbi.RegularSubBlockModel()
rsbm.parent_block_size = [1.5, 2.5, 10.]
rsbm.parent_block_count = [3, 2, 1]
rsbm.sub_block_count = [2, 2, 2]
rsbm.validate()
print("cbi:", rsbm.compressed_block_index)
print("num_blocks:", rsbm.num_blocks)
print("is_sub_blocked:", rsbm.is_sub_blocked)
print("sub_block_size:", rsbm.sub_block_size)
cbi_plot.plot_rsbm(rsbm)
rsbm.refine((0, 1, 0))
print("cbi:", rsbm.compressed_block_index)
print("num_blocks:", rsbm.num_blocks)
print("is_sub_blocked:", rsbm.is_sub_blocked)
print("sub_block_size:", rsbm.sub_block_size)
cbi_plot.plot_rsbm(rsbm)
```
# Octree Sub Block Model
The _Octree Sub Block Model_ is a "forest" of individual octrees, with the "root" of every octree positioined at the center of each parent block within a _Regular Block Model_. Each octree is stored as a [Linear Octree](https://en.wikipedia.org/wiki/Linear_octree) with the space-filling curve chosen to be a [Z-Order Curve](https://en.wikipedia.org/wiki/Z-order_curve) (also known as a Morton curve). The Z-Order curve was chosen based on the efficient properties of bit-interleaving to produce a sorted integer array that defines both the attribute ordering and the topology of the sub blocks; this has been used successfully in HPC algorithms for "forests of octrees" (e.g. [Parallel Forset of Octree's](https://epubs.siam.org/doi/abs/10.1137/100791634), [PDF](http://p4est.github.io/papers/BursteddeWilcoxGhattas11.pdf)). Note, that the maximum level necessary for each octree must be decided upon in OMF.v2, the industry standard is up to eight refinements, and that has been proposed. The `level` information is stored in this integer through a left-shift binary operation (i.e. `(z_order << 3) + level`). For efficient access to the attributes, the _Compressed Block Index_ is also stored.
**Stored Properties**
* `parent_block_size`: a Vector3 (Float64) that describes how large each parent block is
* `parent_block_count`: a Vector3 (Int16) that describes how many parent blocks in each dimension
* `compressed_block_index`: a UInt32 array of length (`i * j * k + 1`) that defines delineation between octrees in the forest
* `z_order_curves`: a UInt32 array of length `num_blocks` containing the Z-Order curves for all octrees. Unrefined parents have z-order curve of `0`
See first three functions of [discretize tree mesh](https://github.com/simpeg/discretize/blob/1721a8626682cf7df0083f8401fff9d0c643b999/discretize/TreeUtils.pyx) for an implementation of z-order curve.
```
osbm = cbi.OctreeSubBlockModel()
osbm.parent_block_size = [1.5, 2.5, 10.]
osbm.parent_block_count = [3, 2, 1]
osbm.validate();
print('cbi: ', osbm.compressed_block_index)
print('z_order_curves: ', osbm.z_order_curves)
print('num_blocks: ', osbm.num_blocks)
cbi_plot.plot_osbm(osbm)
# This part needs work in the implementation for a high level wrapper
osbm._refine_child((0, 1, 0), 0)
osbm._refine_child((0, 1, 0), 1)
print('cbi: ', osbm.compressed_block_index)
print('z_order_curves: ', osbm.z_order_curves)
print('num_blocks: ', osbm.num_blocks)
cbi_plot.plot_osbm(osbm)
```
## Octree Pointers and Level
A Z-Order curve is used to encode each octree into a linear array. The below example shows visually how the pointer and level information is encoded into a single 32 bit integer. The key pieces are to decide how many levels are possible within each tree. Choosing the current industry standard of 8 levels, allows for 256 sub-blocks in each dimension. This can accomodate 16.7 million sub-blocks within each parent block. Note that the actual block model may have many more blocks than those in a single parent block.
The `pointer` of an octree sub-block has an `ijk` index, which is the sub-block corner relative to the parent block corner, the max dimension of each is 256. There is also a `level` that corresponds to the level of the octree -- 0 corresponds to the largest block size (i.e. same as the parent block) and 7 corresponds to the smallest block size.
The sub-blocks must be refined as an octree. That is, the root block has `level=0` and `width=256`, and can be refined into 8 children - each with `level=1` and `width=128`.
```
osbm = cbi.OctreeSubBlockModel()
osbm.parent_block_size = [1.5, 2.5, 10.]
osbm.parent_block_count = [3, 2, 1]
print('Refine the (0, 0, 0) parent block.')
children = osbm._refine_child((0, 0, 0), 0)
print('The children are:')
print(children)
print('Refine the (0, 0, 0) parent block, sub-block (0, 0, 0, 1).')
children = osbm._refine_child((0, 0, 0), 1)
print('The children are:')
print(children)
```
## Linear Octree Encoding
The encoding into a linear octree is done through bit-interleaving of each location integer. This produces a [Z-Order Curve](https://en.wikipedia.org/wiki/Z-order_curve), which is a space filling curve - it guarantees a unique index for each block, and has the nice property that blocks close together are stored close together in the attribute arrays.
<center><img src="zordercurve.png" style="width:250px"><br>Visualization of the space filling Z-Order Curve</center>
```
pointer = [0, 128, 0]
level = 1
ind = z_order_utils.get_index(pointer, level)
pnt, lvl = z_order_utils.get_pointer(ind)
# assert that you get back what you put in:
assert (pointer == pnt) & (level == lvl)
print(ind)
print(pnt, lvl)
```
The actual encoding is completed through bit-interleaving of the three ijk-indices and then adding the level via left-shifting the integer. This is visualized in text as:
```
z_order_utils._print_example(pointer, level);
```
### Octree Storage Summary
The overall storage format reduces to two arrays, (1) `csb` has length equal to the number of parent blocks; (2) `z_order_curves` has length equal to the total number of sub-blocks. This parallels standard storage formats for sparse matrices as well as standard octree storage formats. The outcome is a storage format that is compact and allows for efficient access of, for example, all sub-blocks in a parent block. The contiguous data access allows for memory-mapped arrays, among other efficiencies. The format is also **twelve times more efficient** than the equivalent storage of an _Arbtrary Sub Block Model_ (one UInt32 vs six Float64 arrays). For example, a 10M cell block model **saves 3.52 GB of space** stored in this format. The format also enforces consistency on the indexing of the attributes. These efficiencies, as well as classic algorithms possible for searching octree, can be taken advantage of in vendor applications both for visualization and for evaluation of other attributes.
# Arbitrary Sub Block Model
The _Arbitrary Sub Block Model_ is the most flexible and also least efficient storage format. The format allows for storage of arbitrary blocks that are contained within the parent block. The _Arbitrary Sub Block Model_ does not enforce that sub-blocks fill the entire space of the parent block.
**Stored Properties**
* `parent_block_size`: a Vector3 (Float64) that describes how large each parent block is
* `parent_block_count`: a Vector3 (Int16) that describes how many parent blocks in each dimension
* `compressed_block_index`: a UInt32 array of length (`i * j * k + 1`) that defines the sub block count
* `sub_block_centroids`: a Float64 array containing the sub block centroids for all parent blocks - there are no assumptions about how the sub-blocks are ordered within each parent block
* `sub_block_sizes`: a Float64 array containing the sub block sizes for all parent blocks
**Centroids and Sizes**
These are stored as a two Float64 arrays as `[x_1, y_1, z_1, x_2, y_2, z_2, ...]` to ensure centroids can easily be accessed through the `cbi` indexing as well as memory mapped per parent block. The sizes and centroids are **normalized** within the parent block, that is, `0 < centroid < ` and `0 < size <= 1`. This has 2 advantages: (1) it's easy to tell if values are outside the parent block, and (2) given a large offset, this may allow smaller storage size.
**Parent blocks without sub blocks**
Since the `cbi` is used to index into sub block centroid/size arrays, non-sub-blocked parents require an entry in these arrays. Likely this is centroid `[.5, .5, .5]` and size `[1, 1, 1]`.
**Question: Centroid vs. Corner**
Should we store the `corner` instead to be consistent with the orientation of the block model storage? Storing the corner means three less operations per centroid for checking if it is contained by the parent (although one more for access, as centroid seems to be the industry standard). We could even store opposing corners, instead of corner and size, which would enable exact comparisons to determine adjacent sub blocks.
There is no _storage_ advantage to corners/corners vs. corners/sizes vs. centroids/sizes, especially if these are all normalized. Corners/sizes gives the most API consistency, since we store block model corner and block size. Regardless of which we store, all these properties should be exposed in the client libraries.
```
asbm = cbi.ArbitrarySubBlockModel()
asbm.parent_block_size = [1.5, 2.5, 10.]
asbm.parent_block_count = [3, 2, 1]
asbm.validate();
print('cbi: ', asbm.compressed_block_index)
print('num_blocks: ', asbm.num_blocks)
print('num_parent_blocks: ', asbm.num_parent_blocks)
# Nothing to plot to start with
def add_parent_block(asbm, ijk):
"""Nothing special about these, they are just sub-blocks."""
pbs = np.array(asbm.parent_block_size)
half = pbs / 2.0
offset = half + pbs * ijk
asbm._add_sub_blocks(ijk, offset, half*2)
# Something special for the first ones
asbm._add_sub_blocks(
(0, 0, 0), [0.75, 1.25, 2.5], [1.5, 2.5, 5.]
)
asbm._add_sub_blocks(
(0, 0, 0), [0.375, 1.25, 7.5], [0.75, 2.5, 5.]
)
asbm._add_sub_blocks(
(0, 0, 0), [1.175, 1.25, 7.5], [0.75, 2.5, 5.]
)
add_parent_block(asbm, (1, 0, 0))
add_parent_block(asbm, (2, 0, 0))
add_parent_block(asbm, (0, 1, 0))
add_parent_block(asbm, (1, 1, 0))
add_parent_block(asbm, (2, 1, 0))
print('cbi: ', asbm.compressed_block_index)
print('num_blocks: ', asbm.num_blocks)
print('num_parent_blocks: ', asbm.num_parent_blocks)
cbi_plot.plot_asbm(asbm)
```
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.