text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
```
import numpy as np
from matplotlib import pyplot as plt
import sys
sys.path.append('/global/u1/s/sfschen/Python/velocileptors/')
from velocileptors.LPT.lpt_rsd_fftw import LPT_RSD
from classy import Class
z = 0.61
omega_b = 0.02242
omega_c = 0.11933
lnAs = 3.047
h = 0.6766
ns = 0.9665
nnu = 1
nur = 2.033
mnu = 0.06037735849
fac = 1.0
pkparams = {
'output': 'mPk',
'P_k_max_h/Mpc': 20.,
'z_pk': '0.0,10',
'A_s': fac * np.exp(lnAs)*1e-10,
'n_s': ns,
'h': h,
'N_ur': nur,
'N_ncdm': nnu,
'm_ncdm': mnu,
'tau_reio': 0.0568,
'omega_b': omega_b,
'omega_cdm': omega_c}
import time
t1 = time.time()
pkclass = Class()
pkclass.set(pkparams)
pkclass.compute()
speed_of_light = 2.99792458e5
Hz = pkclass.Hubble(z) * speed_of_light / h # this H(z) in units km/s/(Mpc/h) = 100 * E(z)
chiz = pkclass.angular_distance(z) * (1.+z) * h # this is the comoving radius in units of Mpc/h
fnu = pkclass.Omega_nu/pkclass.Omega_m()
f = pkclass.scale_independent_growth_factor_f(z) * (1 - 0.6*fnu)
ki = np.logspace(-3.0,1.0,200)
pi = np.array( [pkclass.pk_cb(k*h, z ) * h**3 for k in ki] )
sigma8 = pkclass.sigma8()
sigma8
# Reference Cosmology:
Omega_M = 0.31
fb = 0.1571
pkparams = {
'output': 'mPk',
'P_k_max_h/Mpc': 20.,
'z_pk': '0.0,10',
'A_s': np.exp(3.040)*1e-10,
'n_s': ns,
'h': h,
'N_ur': 3.046,
'N_ncdm': 0,#1,
#'m_ncdm': 0,
'tau_reio': 0.0568,
'omega_b': h**2 * fb * Omega_M,
'omega_cdm': h**2 * (1-fb) * Omega_M}
import time
t1 = time.time()
pkclass = Class()
pkclass.set(pkparams)
pkclass.compute()
Hz_fid = pkclass.Hubble(z) * speed_of_light / h # this H(z) in units km/s/(Mpc/h) = 100 * E(z)
chiz_fid = pkclass.angular_distance(z) * (1.+z) * h # this is the comoving radius in units of Mpc/h
apar, aperp = Hz_fid / Hz, chiz / chiz_fid
# Now make the PT object:
modPT = LPT_RSD(ki, pi, kIR=0.2,\
cutoff=10, extrap_min = -4, extrap_max = 3, N = 2000, threads=1, jn=5)
modPT.make_pltable(f, kmin=1e-3, kmax=0.5, nk = 200, apar=apar, aperp=aperp, ngauss=4)
# Now set up the power spectrum model
from scipy.interpolate import InterpolatedUnivariateSpline as Spline
kth = np.arange(0, 0.4, 0.001)
kth_c = kth + 0.001/2
kobs = np.arange(0,0.4,0.01)
kobs_c = kobs + 0.01/2
def pkell(pars):
b1, b2, bs, alpha0, alpha2, sn2 = pars
b3 = 0
sn0 = 0
alpha4 = 0
sn4 = 0
bvec = [b1,b2,bs,b3,alpha0,alpha2,alpha4,0,1e3*sn0,1e3*sn2,1e3*sn4]
kv, p0, p2, p4 = modPT.combine_bias_terms_pkell(bvec)
p0th, p2th, p4th = Spline(kv,p0,ext=3)(kth_c), Spline(kv,p2,ext=3)(kth_c), Spline(kv,p4,ext=3)(kth_c)
return p0th, p2th, p4th
# Load matrices and such things:
from pktools import*
filename = '../boss_data_renorm/pk/ps1D_BOSS_DR12_NGC_z3_COMPnbar_TSC_700_700_700_400_renorm.dat'
dats = read_power(filename)
dats['sig0']
plt.figure(figsize=(10,5))
kdat = dats['k_center']
p0dat = dats['pk0']
err0 = dats['sig0']
p2dat = dats['pk2']
err2 = dats['sig2']
plt.errorbar(kdat, kdat * p0dat, yerr=kdat * err0, fmt='o')
plt.errorbar(kdat, kdat * p2dat, yerr=kdat * err2, fmt='o')
plt.xlim(0,0.2)
M = np.loadtxt('../boss_data_renorm/matrices/M_BOSS_DR12_NGC_z3_V6C_1_1_1_1_1_1200_2000.matrix.gz')
W = np.loadtxt('../boss_data_renorm/matrices/W_BOSS_DR12_NGC_z3_V6C_1_1_1_1_1_10_200_2000_averaged_v1.matrix.gz')
# Now define function with the whole thing:
def pkobs(bvec):
p0th, p2th, p4th = pkell(bvec)
pk_model_vector = np.concatenate((p0th,p2th,p4th))
# wide angle
expanded_model = np.matmul(M, pk_model_vector )
# Convolve with window (true) −> (conv) see eq. 2.18
convolved_model = np.matmul(W, expanded_model )
return convolved_model
# Trial run:
thy = pkobs([1,0,0,0,0,0])
p0thy = thy[:40]
p2thy = thy[80:120]
plt.plot(kobs, kobs * p0thy)
plt.plot(kobs, kobs * p2thy)
# Now set up the covariance
cov = np.loadtxt('../boss_data_renorm/covariances/C_2048_BOSS_DR12_NGC_z3_V6C_1_1_1_1_1_10_200_200_prerecon.matrix.gz')
err0 = np.diag(cov)[:40]**0.5
err2 = np.diag(cov)[80:120]**0.5
# Set the fit ranges
kbins, pkvec = dict_to_vec(dats)
krange = (kobs_c > 0.02) * (kobs_c < 0.2)
kexcl = kobs_c < 0
fit_range = np.concatenate( (krange, kexcl, krange, kexcl, kexcl))
fit_data = pkvec[fit_range]
Cinv = np.linalg.inv( cov[np.ix_(fit_range, fit_range)] )
plt.imshow( np.log(np.linalg.inv(Cinv)) )
def chi2(bvec):
thy = pkobs(bvec)[fit_range]
diff = thy - fit_data
return np.dot(diff ,np.dot(Cinv ,diff ))
def diff_pk(bvec):
thy = pkobs(bvec)[fit_range]
diff = thy - fit_data
return diff
err0
plt.plot(kobs_c, err0/p0dat, c='C0')
plt.plot(kobs_c, -err0/p0dat, c='C0')
plt.plot(kobs_c, err2/p2dat, c='C1')
plt.plot(kobs_c, -err2/p2dat, c='C1')
plt.ylim(-0.1,0.1)
plt.xlim(0,0.2)
plt.errorbar(kobs_c, kobs_c * p2dat, yerr= kobs_c * err2, fmt='o', c='C1')
plt.xlim(0,0.2)
plt.ylim(200,400)
# optimize
from scipy.optimize import minimize
res = minimize(chi2, [1,0,0,0,0,0] )
plt.figure(figsize=(10,5))
thy = pkobs(res.x)
p0thy = thy[0:40]
p2thy = thy[80:120]
plt.plot(kobs_c, kobs_c * p0thy)
plt.plot(kobs_c, kobs_c * p2thy)
plt.errorbar(kobs_c, kobs_c * p0dat, yerr=kobs_c * err0, fmt='o', c='C0')
plt.errorbar(kobs_c, kobs_c * p2dat, yerr= kobs_c * err2, fmt='o', c='C1')
plt.text(0.18, 1750, 'NGC z3')
plt.text(0.18, 1600, r'$\sigma_8 = $ %.2f' %(sigma8))
plt.text(0.18, 1450, r'$\chi^2 = $ %.1f' %(res.fun))
plt.xlabel('k [h/Mpc]')
plt.ylabel(r'$k P_\ell$ [(Mpc/h)$^2$]')
plt.xlim(0,0.2)
plt.ylim(-100,2200)
plt.figure(figsize=(10,5))
thy = pkobs(res.x)
p0thy = thy[0:40]
p2thy = thy[80:120]
#plt.plot(kobs_c, kobs_c * p0thy)
#plt.plot(kobs_c, kobs_c * p2thy)
plt.errorbar(kobs_c, kobs_c * (p0dat - p0thy), yerr=kobs_c * err0, fmt='o', c='C0')
plt.errorbar(kobs_c, kobs_c * (p2dat - p2thy), yerr= kobs_c * err2, fmt='o', c='C1')
#plt.text(0.18, 1750, 'NGC z3')
#plt.text(0.18, 1600, r'$\sigma_8 = $ %.2f' %(sigma8))
#plt.text(0.18, 1450, r'$\chi^2 = $ %.1f' %(res.fun))
plt.xlabel('k [h/Mpc]')
plt.ylabel(r'$ \Delta k P_\ell$ [(Mpc/h)$^2$]')
plt.xlim(0,0.2)
plt.ylim(-200,200)
# Look at a bunch of values of sigma8:
facs = [0.6, 0.8, 1.0, 1.2]
#facs = np.arange(0.6, 1.2, 0.025)
chi2s = []
p0ths = []
p2ths = []
for fac in facs:
print(fac)
kvec = np.concatenate( ([0.0005,], np.logspace(np.log10(0.0015),np.log10(0.025),10, endpoint=True), np.arange(0.03,0.51,0.01)) )
modPT = LPT_RSD(ki, fac * pi, kIR=0.2,\
cutoff=10, extrap_min = -4, extrap_max = 3, N = 2000, threads=1, jn=5)
#modPT.make_pltable(f, kmin=1e-3, kmax=0.5, nk = 200, apar=apar, aperp=aperp, ngauss=4)
modPT.make_pltable(f, kv=kvec, apar=apar, aperp=aperp, ngauss=4)
res = minimize(chi2, [1,0,0,0,0,0], options={'gtol': 1e-08,} )
print(res.x)
thy = pkobs(res.x)
p0ths += [thy[0:40], ]
p2ths += [thy[80:120], ]
chi2s += [res.fun]
plt.figure(figsize=(10,5))
thy = pkobs(res.x)
p0thy = thy[0:40]
p2thy = thy[80:120]
for ii, fac in enumerate(facs):
print(sigma8)
plt.plot(kobs_c, kobs_c * p0ths[ii], 'C'+str(ii), label=r'$\sigma_8,\, \chi^2 = $%.2f, %.1f' %(sigma8*np.sqrt(fac),chi2s[ii]))
plt.plot(kobs_c, kobs_c * p2ths[ii], 'C'+str(ii))
plt.errorbar(kobs_c, kobs_c * p0dat, yerr=kobs_c * err0, fmt='o', c='k')
plt.errorbar(kobs_c, kobs_c * p2dat, yerr= kobs_c * err2, fmt='o', c='k')
#plt.text(0.18, 1750, 'NGC z3')
#plt.text(0.18, 1600, r'$\sigma_8 = $ %.2f' %(sigma8))
#plt.text(0.18, 1450, r'$\chi^2 = $ %.1f' %(res.fun))
plt.xlabel('k [h/Mpc]')
plt.ylabel(r'$k P_\ell$ [(Mpc/h)$^2$]')
plt.legend(ncol=2)
plt.xlim(0,0.2)
plt.ylim(-100,2000)
plt.figure(figsize=(10,5))
thy = pkobs(res.x)
p0thy = thy[0:40]
p2thy = thy[80:120]
for ii, fac in enumerate(facs):
#plt.plot(kobs_c, kobs_c * p0ths[ii], 'C'+str(ii), label=r'$\sigma_8,\, \chi^2 = $%.2f, %.1f' %(sigma8*np.sqrt(fac),chi2s[ii]))
#plt.plot(kobs_c, kobs_c * p2ths[ii], 'C'+str(ii))
plt.errorbar(kobs_c, kobs_c**2 * (p0dat - p0ths[ii]), yerr=kobs_c**2 * err0, fmt='o', c='C'+str(ii), label=r'$\sigma_8,\, \chi^2 = $%.2f, %.1f' %(sigma8*np.sqrt(fac),chi2s[ii]))
plt.errorbar(kobs_c, kobs_c**2 * (p2dat - p2ths[ii]), yerr= kobs_c**2 * err2, fmt='^', c='C'+str(ii))
#plt.text(0.18, 1750, 'NGC z3')
#plt.text(0.18, 1600, r'$\sigma_8 = $ %.2f' %(sigma8))
#plt.text(0.18, 1450, r'$\chi^2 = $ %.1f' %(res.fun))
plt.xlabel('k [h/Mpc]')
plt.ylabel(r'$k^2 \Delta P_\ell$ [(Mpc/h)$^2$]')
plt.legend(loc='lower left', ncol=2)
plt.xlim(0,0.2)
plt.ylim(-20,20)
SMALL_SIZE = 8
MEDIUM_SIZE = 10
BIGGER_SIZE = 10
plt.rc('font', size=MEDIUM_SIZE) # controls default text sizes
plt.rc('axes', titlesize=BIGGER_SIZE) # fontsize of the axes title
plt.rc('axes', labelsize=BIGGER_SIZE) # fontsize of the x and y labels
plt.rc('xtick', labelsize=BIGGER_SIZE) # fontsize of the tick labels
plt.rc('ytick', labelsize=BIGGER_SIZE) # fontsize of the tick labels
plt.rc('legend', fontsize=SMALL_SIZE) # legend fontsize
plt.rc('figure', titlesize=BIGGER_SIZE) # fontsize of the figure title
# Joint Plot
fig = plt.figure(figsize=(8,6))
plt.subplot(2,1,1)
for ii, fac in enumerate(facs):
print(sigma8)
plt.plot(kobs_c, kobs_c * p0ths[ii], 'C'+str(ii), )
plt.plot(kobs_c, kobs_c * p2ths[ii], 'C'+str(ii))
plt.errorbar(kobs_c, kobs_c * p0dat, yerr=kobs_c * err0, fmt='o', c='k', label=r'$\ell = 0$')
plt.errorbar(kobs_c, kobs_c * p2dat, yerr= kobs_c * err2, fmt='^', c='k', label=r'$\ell = 2$')
#plt.text(0.18, 1750, 'NGC z3')
#plt.text(0.18, 1600, r'$\sigma_8 = $ %.2f' %(sigma8))
#plt.text(0.18, 1450, r'$\chi^2 = $ %.1f' %(res.fun))
#plt.xlabel('k [h/Mpc]')
plt.ylabel(r'$k P_\ell$ [(Mpc/h)$^2$]')
plt.legend(loc='upper right',ncol=3)
plt.xlim(0,0.2)
plt.ylim(-100,2200)
plt.text(0.0915, 100, 'NGCz3')
plt.xticks([])
plt.subplot(2,1,2)
for ii, fac in enumerate(facs):
#plt.plot(kobs_c, kobs_c * p0ths[ii], 'C'+str(ii), label=r'$\sigma_8,\, \chi^2 = $%.2f, %.1f' %(sigma8*np.sqrt(fac),chi2s[ii]))
#plt.plot(kobs_c, kobs_c * p2ths[ii], 'C'+str(ii))
plt.errorbar(kobs_c, kobs_c**2 * (p0dat - p0ths[ii]), yerr=kobs_c**2 * err0, fmt='o', c='C'+str(ii))
plt.errorbar(kobs_c, kobs_c**2 * (p2dat - p2ths[ii]), yerr= kobs_c**2 * err2, fmt='^', c='C'+str(ii))
plt.plot(-1000,1000, 'C'+str(ii), label=r'$\sigma_8,\, \chi^2 = $%.2f, %.1f' %(sigma8*np.sqrt(fac),chi2s[ii]))
#plt.text(0.18, 1750, 'NGC z3')
#plt.text(0.18, 1600, r'$\sigma_8 = $ %.2f' %(sigma8))
#plt.text(0.18, 1450, r'$\chi^2 = $ %.1f' %(res.fun))
plt.plot([0,1],[0,0],'k--')
plt.xlabel('k [h/Mpc]')
plt.ylabel(r'$k^2 \Delta P_\ell$ [(Mpc/h)$^2$]')
#plt.legend(loc='lower left', ncol=2)
plt.legend(loc='lower center', ncol=2)
plt.xlim(0,0.2)
plt.ylim(-22,22)
plt.tight_layout()
plt.savefig('Figures/sigma8_scan_planck_lcdm_ngc_z3.pdf')
plt.plot(facs**0.5 * sigma8, chi2s, 'o-')
chi2(res.x)
chi2(res.x)
a, b, c = pkell(res.x)
d, e, g = pkell(res.x)
plt.semilogx(kth_c, b/e-1, '.')
plt.ylim(-0.01,0.01)
t2-t1
# Testing for number of k bins in modPT
# Now make th
#make PT object:
import time
t1 = time.time()
modPT = LPT_RSD(ki, pi, kIR=0.2,\
cutoff=10, extrap_min = -4, extrap_max = 3, N = 2000, threads=1, jn=5)
modPT.make_pltable(f, kmin=1e-3, kmax=0.5, nk = 200, apar=apar, aperp=aperp, ngauss=4)
#modPT.make_pltable(f, kv=kth_c, apar=apar, aperp=aperp, ngauss=4)
t2 = time.time()
print(t2-t1)
# Leaner
#kvec = np.concatenate( (np.logspace(-3,np.log10(0.025),10, endpoint=True), np.arange(0.03,0.45,0.01)) )
t1 = time.time()
modPT = LPT_RSD(ki, pi, kIR=0.2,\
cutoff=10, extrap_min = -4, extrap_max = 3, N = 2000, threads=1, jn=5)
modPT.make_pltable(f, kv=kvec, apar=apar, aperp=aperp, ngauss=4)
t2 =time.time()
print(t2-t1)
kvec.shape
def pkell(pars,PT):
b1, b2, bs, alpha0, alpha2, sn2 = pars
b3 = 0
sn0 = 0
alpha4 = 0
sn4 = 0
bvec = [b1,b2,bs,b3,alpha0,alpha2,alpha4,0,1e3*sn0,1e3*sn2,1e3*sn4]
kv, p0, p2, p4 = PT.combine_bias_terms_pkell(bvec)
p0th, p2th, p4th = Spline(kv,p0,ext=3)(kth_c), Spline(kv,p2,ext=3)(kth_c), Spline(kv,p4,ext=3)(kth_c)
return p0th, p2th, p4th
p0fid, p2fid, p4fid = pkell(res.x, modPT)
p01, p21, p41 = pkell(res.x, modPT1)
plt.plot(kth_c, p2fid/p21-1)
plt.ylim(-0.01,0.01)
#kv = np.logspace(-3,np.log10(0.5),50)
kvec = np.concatenate( ([0.0005,], np.logspace(np.log10(0.0015),np.log10(0.025),10, endpoint=True), np.arange(0.03,0.51,0.01)) )
plt.plot(kvec, kvec * np.interp(kvec, ki,pi), '.')
np.arange(0.06,0.5,0.01).shape
kvec.shape
```
| github_jupyter |
```
%matplotlib inline
import pandas as pd
import os
import networkx as nx
import matplotlib.pyplot as plt
import caselawnet
filepath = '/media/sf_VBox_Shared/CaseLaw/2018-01-29-lido/derived/'
links_sub = pd.read_csv(os.path.join(filepath, 'subnetwork', 'case-to-case-links.csv'))
cases_sub = pd.read_csv(os.path.join(filepath, 'subnetwork', 'case-nodes-sub.csv'))
links_sub_ext = pd.read_csv(os.path.join(filepath, 'subnetwork', 'case-to-case-links-ext.csv'))
cases_sub_ext = pd.read_csv(os.path.join(filepath, 'subnetwork', 'case-nodes-ext.csv'))
links_sub_hr = pd.read_csv(os.path.join(filepath, 'subnetwork', 'case-to-case-links-hr.csv'))
print(len(cases_sub), len(links_sub))
print(len(cases_sub_ext), len(links_sub_ext))
ecli_links_sub = pd.DataFrame({'source': links_sub.source.map(lambda s: s.split('/')[-1]), 'target': links_sub.target.map(lambda s: s.split('/')[-1])})
ecli_links_sub.to_csv('/media/sf_VBox_Shared/CaseLaw/2018-01-29-lido/derived/subnetwork/article/ecli_links.csv', index=False)
ecli_links_sub_ext = pd.DataFrame({'source': links_sub_ext.source.map(lambda s: s.split('/')[-1]), 'target': links_sub_ext.target.map(lambda s: s.split('/')[-1])})
ecli_links_sub_ext.to_csv('/media/sf_VBox_Shared/CaseLaw/2018-01-29-lido/derived/subnetwork/article/ecli_links_extended.csv', index=False)
ecli_links_sub_hr = pd.DataFrame({'source': links_sub_hr.source.map(lambda s: s.split('/')[-1]), 'target': links_sub_hr.target.map(lambda s: s.split('/')[-1])})
ecli_links_sub_hr.to_csv('/media/sf_VBox_Shared/CaseLaw/2018-01-29-lido/derived/subnetwork/article/ecli_links_hr.csv', index=False)
graph_sub = nx.from_pandas_edgelist(ecli_links_sub)
graph_sub_gcc = graph_sub.subgraph(sorted(nx.connected_components(graph_sub), key=len)[-1])
ecli_links_sub_gcc = pd.DataFrame([(s, t) for s,t in graph_sub_gcc.edges()], columns=['source', 'target'])
ecli_links_sub_gcc.to_csv('/media/sf_VBox_Shared/CaseLaw/2018-01-29-lido/derived/subnetwork/article/ecli_links_gcc.csv', index=False)
graph_sub_ext = nx.from_pandas_edgelist(ecli_links_sub_ext)
graph_sub_ext_gcc = graph_sub.subgraph(sorted(nx.connected_components(graph_sub_ext), key=len)[-1])
ecli_links_sub_ext_gcc = pd.DataFrame([(s, t) for s,t in graph_sub_ext_gcc.edges()], columns=['source', 'target'])
ecli_links_sub_ext_gcc.to_csv('/media/sf_VBox_Shared/CaseLaw/2018-01-29-lido/derived/subnetwork/article/ecli_links_ext_gcc.csv', index=False)
```
## Remove PHR cases
```
fp_out = '/media/sf_VBox_Shared/CaseLaw/2018-01-29-lido/derived/subnetwork/article/no_phr/'
def remove_phr(df):
return df[(df.source.map(lambda s: s.split(':')[2])!='PHR') & (df.target.map(lambda s: s.split(':')[2])!='PHR')]
ecli_links_sub_nophr = remove_phr(ecli_links_sub)
ecli_links_sub_nophr.to_csv(os.path.join(fp_out, 'ecli_links.csv'), index=False)
ecli_links_sub_ext_nophr = remove_phr(ecli_links_sub_ext)
ecli_links_sub_ext_nophr.to_csv(os.path.join(fp_out, 'ecli_links_extended.csv'), index=False)
graph_sub_nophr = nx.from_pandas_edgelist(ecli_links_sub_nophr)
graph_sub_npphr_gcc = graph_sub_nophr.subgraph(sorted(nx.connected_components(graph_sub_nophr), key=len)[-1])
ecli_links_sub_gcc = pd.DataFrame([(s, t) for s,t in graph_sub_npphr_gcc.edges()], columns=['source', 'target'])
ecli_links_sub_gcc.to_csv(os.path.join(fp_out, 'ecli_links_gcc.csv'), index=False)
graph_sub_nophr = nx.from_pandas_edgelist(ecli_links_sub_ext_nophr)
graph_sub_npphr_gcc = graph_sub_nophr.subgraph(sorted(nx.connected_components(graph_sub_nophr), key=len)[-1])
ecli_links_sub_gcc = pd.DataFrame([(s, t) for s,t in graph_sub_npphr_gcc.edges()], columns=['source', 'target'])
ecli_links_sub_gcc.to_csv(os.path.join(fp_out, 'ecli_links_ext_gcc.csv'), index=False)
```
| github_jupyter |
# Benchmarker Analysis
Analysis of tng-sdk-benchmark's behavior for 5GTANGO D6.3.
```
%matplotlib inline
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
import matplotlib
import numpy as np
sns.set(font_scale=1.3, style="ticks")
def select_and_rename(df, mapping):
"""
Helper: Selects columns of df using the keys
of the mapping dict.
It renames the columns to the values of the
mappings dict.
"""
# select subset of columns
dff = df[list(mapping.keys())]
# rename
for k, v in mapping.items():
#print("Renaming: {} -> {}".format(k, v))
dff.rename(columns={k: v}, inplace=True)
#print(dff.head())
return dff
def cleanup(df):
"""
Cleanup of df data.
Dataset specific.
"""
def _replace(df, column, str1, str2):
if column in df:
df[column] = df[column].str.replace(str1, str2)
def _to_num(df, column):
if column in df:
df[column] = pd.to_numeric(df[column])
_replace(df, "flow_size", "tcpreplay -i data -tK --loop 40000 --preload-pcap /pcaps/smallFlows.pcap", "0")
_replace(df, "flow_size", "tcpreplay -i data -tK --loop 40000 --preload-pcap /pcaps/bigFlows.pcap", "1")
_to_num(df, "flow_size")
_replace(df, "ruleset", "./start.sh small_ruleset", "1")
_replace(df, "ruleset", "./start.sh big_ruleset", "2")
_replace(df, "ruleset", "./start.sh", "0")
_to_num(df, "ruleset")
_replace(df, "req_size", "ab -c 1 -t 60 -n 99999999 -e /tngbench_share/ab_dist.csv -s 60 -k -i http://20.0.0.254:8888/", "0")
_replace(df, "req_size", "ab -c 1 -t 60 -n 99999999 -e /tngbench_share/ab_dist.csv -s 60 -k http://20.0.0.254:8888/bunny.mp4", "1")
_replace(df, "req_size", "ab -c 1 -t 60 -n 99999999 -e /tngbench_share/ab_dist.csv -s 60 -k -i -X 20.0.0.254:3128 http://40.0.0.254:80/", "0")
_replace(df, "req_size", "ab -c 1 -t 60 -n 99999999 -e /tngbench_share/ab_dist.csv -s 60 -k -X 20.0.0.254:3128 http://40.0.0.254:80/bunny.mp4", "1")
_to_num(df, "req_size")
_replace(df, "req_type", "malaria publish -t -n 20000 -H 20.0.0.254 -q 1 --json /tngbench_share/malaria.json", "0")
_replace(df, "req_type", "malaria publish -t -n 20000 -H 20.0.0.254 -q 2 --json /tngbench_share/malaria.json", "1")
_replace(df, "req_type", "malaria publish -s 10 -n 20000 -H 20.0.0.254 --json /tngbench_share/malaria.json", "2")
_replace(df, "req_type", "malaria publish -s 10000 -n 20000 -H 20.0.0.254 --json /tngbench_share/malaria.json", "3")
_to_num(df, "req_type")
```
## Data
```
df_sec01 = pd.read_csv("/home/manuel/sndzoo/ds_nfv_sec01/data/csv_experiments.csv")
df_sec02 = pd.read_csv("/home/manuel/sndzoo/ds_nfv_sec02/data/csv_experiments.csv")
df_sec03 = pd.read_csv("/home/manuel/sndzoo/ds_nfv_sec03/data/csv_experiments.csv")
df_web01 = pd.read_csv("/home/manuel/sndzoo/ds_nfv_web01/data/csv_experiments.csv")
df_web02 = pd.read_csv("/home/manuel/sndzoo/ds_nfv_web02/data/csv_experiments.csv")
df_web03 = pd.read_csv("/home/manuel/sndzoo/ds_nfv_web03/data/csv_experiments.csv")
df_iot01 = pd.read_csv("/home/manuel/sndzoo/ds_nfv_iot01/data/csv_experiments.csv")
df_iot02 = pd.read_csv("/home/manuel/sndzoo/ds_nfv_iot02/data/csv_experiments.csv")
# do renaming and selection
map_sec01 = {
"run_id": "run_id",
"experiment_name": "ex_name",
"experiment_start": "ex_start",
"experiment_stop": "ex_stop",
"param__header__all__config_id": "conf_id",
"param__header__all__repetition": "repetition",
"param__header__all__time_limit": "time_limit",
"param__header__all__time_warmup": "time_warmup",
"param__func__mp.input__cmd_start": "flow_size",
"param__func__de.upb.ids-suricata.0.1__cmd_start": "ruleset",
"param__func__de.upb.ids-suricata.0.1__cpu_bw": "cpu_bw",
"param__func__de.upb.ids-suricata.0.1__mem_max": "memory",
#"metric__vnf0.vdu01.0__suricata_bytes": "ids_bytes",
#"metric__vnf0.vdu01.0__suricata_packets": "ids_pkts",
#"metric__vnf0.vdu01.0__suricata_dropped": "ids_drop",
#"metric__vnf0.vdu01.0__suricata_drops": "ids_drops",
"metric__vnf0.vdu01.0__stat__input__rx_bytes": "if_rx_bytes",
#"metric__vnf0.vdu01.0__stat__input__rx_dropped": "if_in_rx_dropped",
#"metric__vnf0.vdu01.0__stat__input__rx_errors": "if_in_rx_errors",
#"metric__vnf0.vdu01.0__stat__input__rx_packets": "if_in_rx_packets",
#"metric__vnf0.vdu01.0__stat__input__tx_bytes": "if_in_tx_byte",
#"metric__vnf0.vdu01.0__stat__input__tx_dropped": "if_in_tx_dropped",
#"metric__vnf0.vdu01.0__stat__input__tx_errors": "if_in_tx_errors",
#"metric__vnf0.vdu01.0__stat__input__tx_packets": "if_in_tx_packets",
}
map_sec02 = {
"experiment_name": "ex_name",
"experiment_start": "ex_start",
"experiment_stop": "ex_stop",
"param__header__all__config_id": "conf_id",
"param__header__all__repetition": "repetition",
"param__header__all__time_limit": "time_limit",
"param__header__all__time_warmup": "time_warmup",
"param__func__mp.input__cmd_start": "flow_size",
"param__func__de.upb.ids-snort2.0.1__cmd_start": "ruleset",
"param__func__de.upb.ids-snort2.0.1__cpu_bw": "cpu_bw",
"param__func__de.upb.ids-snort2.0.1__mem_max": "memory",
#"metric__vnf0.vdu01.0__snort_bytes": "ids_bytes",
#"metric__vnf0.vdu01.0__snort_packets": "ids_pkts",
#"metric__vnf0.vdu01.0__snort_dropped": "ids_drop",
#"metric__vnf0.vdu01.0__snort_drops": "ids_drops",
"metric__vnf0.vdu01.0__stat__input__rx_bytes": "if_rx_bytes",
#"metric__vnf0.vdu01.0__stat__input__rx_dropped": "if_in_rx_dropped",
#"metric__vnf0.vdu01.0__stat__input__rx_errors": "if_in_rx_errors",
#"metric__vnf0.vdu01.0__stat__input__rx_packets": "if_in_rx_packets",
#"metric__vnf0.vdu01.0__stat__input__tx_bytes": "if_in_tx_byte",
#"metric__vnf0.vdu01.0__stat__input__tx_dropped": "if_in_tx_dropped",
#"metric__vnf0.vdu01.0__stat__input__tx_errors": "if_in_tx_errors",
#"metric__vnf0.vdu01.0__stat__input__tx_packets": "if_in_tx_packets",
}
map_sec03 = {
"experiment_name": "ex_name",
"experiment_start": "ex_start",
"experiment_stop": "ex_stop",
"param__header__all__config_id": "conf_id",
"param__header__all__repetition": "repetition",
"param__header__all__time_limit": "time_limit",
"param__header__all__time_warmup": "time_warmup",
"param__func__mp.input__cmd_start": "flow_size",
"param__func__de.upb.ids-snort3.0.1__cmd_start": "ruleset",
"param__func__de.upb.ids-snort3.0.1__cpu_bw": "cpu_bw",
"param__func__de.upb.ids-snort3.0.1__mem_max": "memory",
#"metric__vnf0.vdu01.0__snort3_total_allow": "ids_allow",
#"metric__vnf0.vdu01.0__snort3_total_analyzed": "ids_anlyzd",
#"metric__vnf0.vdu01.0__snort3_total_received": "ids_recv",
#"metric__vnf0.vdu01.0__snort3_total_outstanding": "ids_outstanding",
#"metric__vnf0.vdu01.0__snort3_total_dropped": "ids_drop",
"metric__vnf0.vdu01.0__stat__input__rx_bytes": "if_rx_bytes",
#"metric__vnf0.vdu01.0__stat__input__rx_dropped": "if_in_rx_dropped",
#"metric__vnf0.vdu01.0__stat__input__rx_errors": "if_in_rx_errors",
#"metric__vnf0.vdu01.0__stat__input__rx_packets": "if_in_rx_packets",
#"metric__vnf0.vdu01.0__stat__input__tx_bytes": "if_in_tx_byte",
#"metric__vnf0.vdu01.0__stat__input__tx_dropped": "if_in_tx_dropped",
#"metric__vnf0.vdu01.0__stat__input__tx_errors": "if_in_tx_errors",
#"metric__vnf0.vdu01.0__stat__input__tx_packets": "if_in_tx_packets",
}
map_web01 = {
"experiment_name": "ex_name",
"experiment_start": "ex_start",
"experiment_stop": "ex_stop",
"param__header__all__config_id": "conf_id",
"param__header__all__repetition": "repetition",
"param__header__all__time_limit": "time_limit",
"param__header__all__time_warmup": "time_warmup",
"param__func__mp.input__cmd_start": "req_size",
"param__func__de.upb.lb-nginx.0.1__cpu_bw": "cpu_bw",
"param__func__de.upb.lb-nginx.0.1__mem_max": "memory",
"metric__mp.input.vdu01.0__ab_completed_requests": "req_compl",
#"metric__mp.input.vdu01.0__ab_concurrent_lvl": "req_concurrent",
#"metric__mp.input.vdu01.0__ab_failed_requests": "req_failed",
#"metric__mp.input.vdu01.0__ab_html_transfer_byte": "req_html_bytes",
#"metric__mp.input.vdu01.0__ab_mean_time_per_request": "req_time_mean",
#"metric__mp.input.vdu01.0__ab_request_per_second": "req_per_sec",
#"metric__mp.input.vdu01.0__ab_time_used_s": "req_time_used",
#"metric__mp.input.vdu01.0__ab_total_transfer_byte": "transf_bytes",
#"metric__mp.input.vdu01.0__ab_transfer_rate_kbyte_per_second": "req_transf_rate",
"metric__vnf0.vdu01.0__stat__input__rx_bytes": "if_rx_bytes",
#"metric__vnf0.vdu01.0__stat__input__rx_dropped": "if_in_rx_dropped",
#"metric__vnf0.vdu01.0__stat__input__rx_errors": "if_in_rx_errors",
#"metric__vnf0.vdu01.0__stat__input__rx_packets": "if_in_rx_packets",
#"metric__vnf0.vdu01.0__stat__input__tx_bytes": "if_tx_bytes",
#"metric__vnf0.vdu01.0__stat__input__tx_dropped": "if_in_tx_dropped",
#"metric__vnf0.vdu01.0__stat__input__tx_errors": "if_in_tx_errors",
#"metric__vnf0.vdu01.0__stat__input__tx_packets": "if_in_tx_packets",
}
map_web02 = {
"experiment_name": "ex_name",
"experiment_start": "ex_start",
"experiment_stop": "ex_stop",
"param__header__all__config_id": "conf_id",
"param__header__all__repetition": "repetition",
"param__header__all__time_limit": "time_limit",
"param__header__all__time_warmup": "time_warmup",
"param__func__mp.input__cmd_start": "req_size",
"param__func__de.upb.lb-haproxy.0.1__cpu_bw": "cpu_bw",
"param__func__de.upb.lb-haproxy.0.1__mem_max": "memory",
"metric__mp.input.vdu01.0__ab_completed_requests": "req_compl",
#"metric__mp.input.vdu01.0__ab_concurrent_lvl": "req_concurrent",
#"metric__mp.input.vdu01.0__ab_failed_requests": "req_failed",
#"metric__mp.input.vdu01.0__ab_html_transfer_byte": "req_html_bytes",
#"metric__mp.input.vdu01.0__ab_mean_time_per_request": "req_time_mean",
#"metric__mp.input.vdu01.0__ab_request_per_second": "req_per_sec",
#"metric__mp.input.vdu01.0__ab_time_used_s": "req_time_used",
#"metric__mp.input.vdu01.0__ab_total_transfer_byte": "transf_bytes",
#"metric__mp.input.vdu01.0__ab_transfer_rate_kbyte_per_second": "req_transf_rate",
"metric__vnf0.vdu01.0__stat__input__rx_bytes": "if_rx_bytes",
#"metric__vnf0.vdu01.0__stat__input__rx_dropped": "if_in_rx_dropped",
#"metric__vnf0.vdu01.0__stat__input__rx_errors": "if_in_rx_errors",
#"metric__vnf0.vdu01.0__stat__input__rx_packets": "if_in_rx_packets",
#"metric__vnf0.vdu01.0__stat__input__tx_bytes": "if_tx_bytes",
#"metric__vnf0.vdu01.0__stat__input__tx_dropped": "if_in_tx_dropped",
#"metric__vnf0.vdu01.0__stat__input__tx_errors": "if_in_tx_errors",
#"metric__vnf0.vdu01.0__stat__input__tx_packets": "if_in_tx_packets",
}
map_web03 = {
"experiment_name": "ex_name",
"experiment_start": "ex_start",
"experiment_stop": "ex_stop",
"param__header__all__config_id": "conf_id",
"param__header__all__repetition": "repetition",
"param__header__all__time_limit": "time_limit",
"param__header__all__time_warmup": "time_warmup",
"param__func__mp.input__cmd_start": "req_size",
"param__func__de.upb.px-squid.0.1__cpu_bw": "cpu_bw",
"param__func__de.upb.px-squid.0.1__mem_max": "memory",
"metric__mp.input.vdu01.0__ab_completed_requests": "req_compl",
#"metric__mp.input.vdu01.0__ab_concurrent_lvl": "req_concurrent",
#"metric__mp.input.vdu01.0__ab_failed_requests": "req_failed",
#"metric__mp.input.vdu01.0__ab_html_transfer_byte": "req_html_bytes",
#"metric__mp.input.vdu01.0__ab_mean_time_per_request": "req_time_mean",
#"metric__mp.input.vdu01.0__ab_request_per_second": "req_per_sec",
#"metric__mp.input.vdu01.0__ab_time_used_s": "req_time_used",
#"metric__mp.input.vdu01.0__ab_total_transfer_byte": "transf_bytes",
#"metric__mp.input.vdu01.0__ab_transfer_rate_kbyte_per_second": "req_transf_rate",
"metric__vnf0.vdu01.0__stat__input__rx_bytes": "if_rx_bytes",
#"metric__vnf0.vdu01.0__stat__input__rx_dropped": "if_in_rx_dropped",
#"metric__vnf0.vdu01.0__stat__input__rx_errors": "if_in_rx_errors",
#"metric__vnf0.vdu01.0__stat__input__rx_packets": "if_in_rx_packets",
#"metric__vnf0.vdu01.0__stat__input__tx_bytes": "if_tx_bytes",
#"metric__vnf0.vdu01.0__stat__input__tx_dropped": "if_in_tx_dropped",
#"metric__vnf0.vdu01.0__stat__input__tx_errors": "if_in_tx_errors",
#"metric__vnf0.vdu01.0__stat__input__tx_packets": "if_in_tx_packets",
}
map_iot01 = {
"experiment_name": "ex_name",
"experiment_start": "ex_start",
"experiment_stop": "ex_stop",
"param__header__all__config_id": "conf_id",
"param__header__all__repetition": "repetition",
"param__header__all__time_limit": "time_limit",
"param__header__all__time_warmup": "time_warmup",
"param__func__mp.input__cmd_start": "req_type",
"param__func__de.upb.broker-mosquitto.0.1__cpu_bw": "cpu_bw",
"param__func__de.upb.broker-mosquitto.0.1__mem_max": "memory",
#"metric__mp.input.vdu01.0__malaria_clientid": "mal_id",
#"metric__mp.input.vdu01.0__malaria_count_ok": "mal_count_ok",
#"metric__mp.input.vdu01.0__malaria_count_total": "mal_count_total",
#"metric__mp.input.vdu01.0__malaria_msgs_per_sec": "msg_per_sec",
#"metric__mp.input.vdu01.0__malaria_rate_ok": "mal_rate_ok",
#"metric__mp.input.vdu01.0__malaria_time_max": "mal_time_max",
#"metric__mp.input.vdu01.0__malaria_time_mean": "msg_t_mean",
#"metric__mp.input.vdu01.0__malaria_time_min": "mal_time_min",
#"metric__mp.input.vdu01.0__malaria_time_stddev": "msg_t_std",
#"metric__mp.input.vdu01.0__malaria_time_total": "mal_time_total",
#"metric__mp.output.vdu01.0__malaria_client_count": "mal_ccount",
#"metric__mp.output.vdu01.0__malaria_clientid": "mal_cid2",
#"metric__mp.output.vdu01.0__malaria_flight_time_max": "mal_ft_max",
#"metric__mp.output.vdu01.0__malaria_flight_time_mean": "mal_ft_mean",
#"metric__mp.output.vdu01.0__malaria_flight_time_min": "mal_ft_min",
#"metric__mp.output.vdu01.0__malaria_flight_time_stddev": "mal_ft_stddev",
#"metric__mp.output.vdu01.0__malaria_ms_per_msg": "mal_ms_per_msg",
#"metric__mp.output.vdu01.0__malaria_msg_count": "mal_out_msg_count",
#"metric__mp.output.vdu01.0__malaria_msg_duplicates": "mal_out_msg_dup",
#"metric__mp.output.vdu01.0__malaria_msg_per_sec": "mal_out_msgs_per_sec",
#"metric__mp.output.vdu01.0__malaria_test_complete": "mal_test_complete",
#"metric__mp.output.vdu01.0__malaria_time_total": "mal_out_t_total",
"metric__vnf0.vdu01.0__stat__input__rx_bytes": "if_rx_bytes",
#"metric__vnf0.vdu01.0__stat__input__rx_dropped": "if_in_rx_dropped",
#"metric__vnf0.vdu01.0__stat__input__rx_errors": "if_in_rx_errors",
#"metric__vnf0.vdu01.0__stat__input__rx_packets": "if_in_rx_packets",
#"metric__vnf0.vdu01.0__stat__input__tx_bytes": "if_tx_bytes",
#"metric__vnf0.vdu01.0__stat__input__tx_dropped": "if_in_tx_dropped",
#"metric__vnf0.vdu01.0__stat__input__tx_errors": "if_in_tx_errors",
#"metric__vnf0.vdu01.0__stat__input__tx_packets": "if_in_tx_packets",
}
map_iot02 = {
"experiment_name": "ex_name",
"experiment_start": "ex_start",
"experiment_stop": "ex_stop",
"param__header__all__config_id": "conf_id",
"param__header__all__repetition": "repetition",
"param__header__all__time_limit": "time_limit",
#"param__header__all__time_warmup": "time_warmup",
#"param__func__mp.input__cmd_start": "req_type",
#"param__func__de.upb.broker-emqx.0.1__cpu_bw": "cpu_bw",
#"param__func__de.upb.broker-emqx.0.1__mem_max": "memory",
#"metric__mp.input.vdu01.0__malaria_clientid": "mal_id",
#"metric__mp.input.vdu01.0__malaria_count_ok": "mal_count_ok",
#"metric__mp.input.vdu01.0__malaria_count_total": "mal_count_total",
#"metric__mp.input.vdu01.0__malaria_msgs_per_sec": "msg_per_sec",
#"metric__mp.input.vdu01.0__malaria_rate_ok": "mal_rate_ok",
#"metric__mp.input.vdu01.0__malaria_time_max": "mal_time_max",
#"metric__mp.input.vdu01.0__malaria_time_mean": "msg_t_mean",
#"metric__mp.input.vdu01.0__malaria_time_min": "mal_time_min",
#"metric__mp.input.vdu01.0__malaria_time_stddev": "msg_t_std",
#"metric__mp.input.vdu01.0__malaria_time_total": "mal_time_total",
#"metric__mp.output.vdu01.0__malaria_client_count": "mal_ccount",
#"metric__mp.output.vdu01.0__malaria_clientid": "mal_cid2",
#"metric__mp.output.vdu01.0__malaria_flight_time_max": "mal_ft_max",
#"metric__mp.output.vdu01.0__malaria_flight_time_mean": "mal_ft_mean",
#"metric__mp.output.vdu01.0__malaria_flight_time_min": "mal_ft_min",
#"metric__mp.output.vdu01.0__malaria_flight_time_stddev": "mal_ft_stddev",
#"metric__mp.output.vdu01.0__malaria_ms_per_msg": "mal_ms_per_msg",
#"metric__mp.output.vdu01.0__malaria_msg_count": "mal_out_msg_count",
#"metric__mp.output.vdu01.0__malaria_msg_duplicates": "mal_out_msg_dup",
#"metric__mp.output.vdu01.0__malaria_msg_per_sec": "mal_out_msgs_per_sec",
#"metric__mp.output.vdu01.0__malaria_test_complete": "mal_test_complete",
#"metric__mp.output.vdu01.0__malaria_time_total": "mal_out_t_total",
"metric__vnf0.vdu01.0__stat__input__rx_bytes": "if_rx_bytes",
#"metric__vnf0.vdu01.0__stat__input__rx_dropped": "if_in_rx_dropped",
#"metric__vnf0.vdu01.0__stat__input__rx_errors": "if_in_rx_errors",
#"metric__vnf0.vdu01.0__stat__input__rx_packets": "if_in_rx_packets",
#"metric__vnf0.vdu01.0__stat__input__tx_bytes": "if_tx_bytes",
#"metric__vnf0.vdu01.0__stat__input__tx_dropped": "if_in_tx_dropped",
#"metric__vnf0.vdu01.0__stat__input__tx_errors": "if_in_tx_errors",
#"metric__vnf0.vdu01.0__stat__input__tx_packets": "if_in_tx_packets",
}
# add additional data
df_sec01["vnf"] = "suricata"
df_sec02["vnf"] = "snort2"
df_sec03["vnf"] = "snort3"
df_web01["vnf"] = "nginx"
df_web02["vnf"] = "haproxy"
df_web03["vnf"] = "squid"
df_iot01["vnf"] = "mosquitto"
df_iot02["vnf"] = "emqx"
# cleanup data sets
dfs_raw = [df_sec01, df_sec02, df_sec03, df_web01, df_web02, df_web03, df_iot01, df_iot02]
map_list = [map_sec01, map_sec02, map_sec03, map_web01, map_web02, map_web03, map_iot01, map_iot02]
dfs = list() # clean data frames
for (df, m) in zip(dfs_raw, map_list):
tmp = select_and_rename(df.copy(), m)
cleanup(tmp)
dfs.append(tmp)
dfs[0].info()
dfs[0]["ex_start"] = pd.to_datetime(dfs[0]["ex_start"], errors='coerce')
dfs[0]["ex_stop"] = pd.to_datetime(dfs[0]["ex_stop"], errors='coerce')
dfs[0]["td_measure"] = dfs[0]["ex_stop"] - dfs[0]["ex_start"]
dfs[0]["td_measure"] = dfs[0]["td_measure"]/np.timedelta64(1,'s')
dfs[0]["delta_s"] = dfs[0]["time_limit"] - dfs[0]["td_measure"]
dfs[0].info()
#dfs[0].describe()
dfs[0]
g = sns.scatterplot(data=dfs[0], x="run_id", y="td_measure", linewidth=0, alpha=0.5)
g.set_ylim(120.0, 120.15)
g.set(xlabel="Experiment run ID", ylabel="Measurement time [s]")
plt.tight_layout()
plt.savefig("bench_roundtime.png", dpi=300)
```
## Experiment Runtime
```
rtdata = list()
rtdata.append({"name": "SEC01", "runtime": 4266})
rtdata.append({"name": "SEC02", "runtime": 4352})
rtdata.append({"name": "SEC03", "runtime": 2145})
rtdata.append({"name": "WEB01", "runtime": 4223})
rtdata.append({"name": "WEB02", "runtime": 4213})
rtdata.append({"name": "WEB03", "runtime": 4232})
rtdata.append({"name": "IOT01", "runtime": 4298})
rtdata.append({"name": "IOT02", "runtime": 6949})
rtdf = pd.DataFrame(rtdata)
rtdf
g = sns.barplot(data=rtdf, x="name", y="runtime", color="gray")
for item in g.get_xticklabels():
item.set_rotation(45)
g.set(xlabel="Experiment", ylabel="Runtime [min]")
plt.tight_layout()
plt.savefig("bench_experiment_runtime_total.png", dpi=300)
```
| github_jupyter |
This notebook __demonstrates the use of [`nnetsauce`](https://github.com/thierrymoudiki/nnetsauce)'s Adaboost classifier__ on two popular (and public) datasets. `nnetsauce`'s implementation of this algorithm has __some specificities__, as it will be shown in the sequel of this notebook. It is worth noting that the __current implementation is 100% in Python__.
We start by installing the package's development version from Github (use the command line):
```
pip install git+https://github.com/thierrymoudiki/nnetsauce.git
```
Next, we __import the packages necessary for the job__, along with `nnetsauce` (namely `numpy` and `sklearn`, nothing weird!):
```
import nnetsauce as ns
import numpy as np
from sklearn.datasets import load_breast_cancer, load_wine, load_iris
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split
from sklearn import metrics
```
Our __first example__ is based on `wisconsin breast cancer` dataset from [UCI (University of California at Irvine) repository](http://archive.ics.uci.edu/ml/index.php), and available in `sklearn`. More details about the content of these datasets can be found [here](http://archive.ics.uci.edu/ml/datasets/Breast+Cancer+Wisconsin+%28Diagnostic%29) and [here](http://archive.ics.uci.edu/ml/datasets/Wine).
`wisconsin breast cancer` dataset is splitted into a __training set__ (for training the model to pattern recognition) and __test set__ (for model validation):
```
# Import dataset from sklearn
breast_cancer = load_breast_cancer()
X = breast_cancer.data
y = breast_cancer.target
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2,
random_state=123)
```
The first version of Adaboost that we apply is __`SAMME.R`__, also known as Real Adaboost. `SAMME` stands for Stagewise Additive Modeling using a Multi-class Exponential loss function, and [`nnetsauce`](https://github.com/thierrymoudiki/nnetsauce)'s implementation of this algorithm has some __specificities__:
- The base learners are quasi-randomized (__deterministic__) networks
- At each boosting iteration, a fraction of the datasets' rows or columns can be randomly chosen to increase diversity of the ensemble
```
# SAMME.R
# base learner
clf = LogisticRegression(solver='liblinear', multi_class = 'ovr',
random_state=123)
# nnetsauce's Adaboost
fit_obj = ns.AdaBoostClassifier(clf,
n_hidden_features=11,
direct_link=True,
n_estimators=250, learning_rate=0.01126343,
col_sample=0.72684326, row_sample=0.86429443,
dropout=0.63078613, n_clusters=2,
type_clust="gmm",
verbose=1, seed = 123,
method="SAMME.R")
```
The base learner, `clf`, is a logistic regression model. __But it could be anything__, including decision trees. `fit_obj` is a `nnetsauce` object that augments `clf` with a hidden layer, and typically makes its predictions nonlinear.
`n_hidden_features` is the number of nodes in the hidden layer, and `dropout` randomly drops some of these nodes at each boosting iteration. `col_sample` and `row_sample` specify the __fraction of columns and rows__ chosen for fitting the base learner at each iteration. With `n_clusters`, the data can be clustered into homogeneous groups before model training.
__`nnetsauce`'s Adaboost can now be fitted__; `250` iterations are used:
```
# Fitting the model to training set
fit_obj.fit(X_train, y_train)
# Obtain model's accuracy on test set
print(fit_obj.score(X_test, y_test))
```
With the following graph, we can __visualize how well our data have been classified__ by `nnetsauce`'s Adaboost.
```
import matplotlib.pyplot as plt
import seaborn as sns; sns.set()
from sklearn.metrics import confusion_matrix
preds = fit_obj.predict(X_test)
mat = confusion_matrix(y_test, preds)
sns.heatmap(mat.T, square=True, annot=True, fmt='d', cbar=False)
plt.xlabel('true label')
plt.ylabel('predicted label');
```
`1` denotes a malignant tumor, and `0`, its absence. For the 3 (out of 114) patients remaining missclassified, it could be interesting to change the model `sample_weight`s, in order to give them more weight in the learning procedure. Then, see how well the result evolves; depending on which decision we consider being the worst (or best). But note that:
- __The model will never be perfect__ (plus, the labels are based on human-eyed labelling ;) ).
- Patients are not labelled. _Label_ is just a generic term in classification, for all types of classification models and data.
Our __second example__ is based on `wine` dataset from [UCI repository](http://archive.ics.uci.edu/ml/index.php). This dataset contains information about wines' quality, depending on their characteristics. `SAMME` is now used instead of `SAMME.R`. This second algorithm seems to require more iterations to converge than `SAMME.R` (but you, tell me from your experience!):
```
# load dataset
wine = load_wine()
Z = wine.data
t = wine.target
np.random.seed(123)
Z_train, Z_test, y_train, y_test = train_test_split(Z, t, test_size=0.2)
# SAMME
clf = LogisticRegression(solver='liblinear', multi_class = 'ovr',
random_state=123)
fit_obj = ns.AdaBoostClassifier(clf,
n_hidden_features=np.int(8.21154785e+01),
direct_link=True,
n_estimators=1000, learning_rate=2.96252441e-02,
col_sample=4.22766113e-01, row_sample=7.87268066e-01,
dropout=1.56909180e-01, n_clusters=3,
type_clust="gmm",
verbose=1, seed = 123,
method="SAMME")
# Fitting the model to training set
fit_obj.fit(Z_train, y_train)
```
After fitting the model, we can obtain some statistics (`accuracy`, `precision`, `recall`, `f1-score`; every `nnetsauce` model is 100% `sklearn`-compatible) about it's quality:
```
preds = fit_obj.predict(Z_test)
print(metrics.classification_report(preds, y_test))
```
| github_jupyter |
```
import tensorflow as tf
import numpy as np
import os
import sys
slim = tf.contrib.slim
from nets import ssd_vgg_300, np_methods
# Input placeholder.
net_shape = (299, 299)
data_format = 'NHWC'
with tf.variable_scope('ssd'):
inputs = tf.placeholder(tf.float32, (None, 299, 299, 3), "input")
# Evaluation pre-processing: resize to SSD net shape.
# Define the SSD model.
reuse = True if 'ssd_net' in locals() else None
ssd_net = ssd_vgg_300.SSDNet()
with slim.arg_scope(ssd_net.arg_scope(data_format=data_format)):
predictions, localisations, _, _ = ssd_net.net(inputs, is_training=False, reuse=reuse)
# Restore SSD model.
ckpt_filename = 'checkpoints/ssd_300_vgg.ckpt/ssd_300_vgg.ckpt'
ssd_session = tf.Session();
ssd_session.run(tf.global_variables_initializer())
saver = tf.train.import_meta_graph('checkpoints/ssd_300_vgg.ckpt/ssd_300_vgg.ckpt.index')
saver.restore(ssd_session, ckpt_filename)
ssd_anchors = ssd_net.anchors(net_shape)
print("restored")
print(inputs)
with tf.variable_scope('ssd/priors'):
feat_shape = tf.placeholder(tf.int32, (1), name="input_shape")
feat_float = tf.placeholder(tf.float32, (1), name="input_div")
offset=0.5
ranged = tf.range(feat_shape[0])
ones = tf.fill([feat_shape[0],feat_shape[0]], 1)
by = tf.cast(tf.multiply(ones, ranged), tf.float32)
bx = tf.cast(tf.multiply(ones, tf.reshape(ranged, (-1, 1))), tf.float32)
y_d = tf.divide((by + offset), feat_float)
y = tf.expand_dims(y_d, 2, name="y")
x_d = tf.divide((bx + offset), feat_float)
x = tf.expand_dims(x_d, 2, name="x")
with tf.Session() as sess:
x_out, y_out = sess.run((x, y), feed_dict={feat_shape: [38], feat_float: [37.5]})
```
w and heights are always the same for our standard shape
[
[[ 0.07023411 0.10281222 0.04966302 0.09932604],
[ 0.07023411 0.10281222 0.09932604 0.04966302]],
[[ 0.15050167 0.22323005 0.10642076 0.21284151 0.08689218 0.26067653],
[ 0.15050167 0.22323005 0.21284151 0.10642076 0.26067653 0.08689218]],
[[ 0.33110368 0.41161588 0.23412566 0.46825132 0.19116279 0.57348841],
[ 0.33110368 0.41161588 0.46825132 0.23412566 0.57348841 0.19116279]],
[[ 0.5117057 0.59519559 0.36183056 0.72366112 0.2954334 0.88630027],
[ 0.5117057 0.59519559 0.72366112 0.36183056 0.88630027 0.2954334]],
[[ 0.69230771 0.77738154 0.48953545 0.9790709],
[ 0.69230771 0.77738154 0.9790709 0.48953545]],
[[ 0.87290972 0.95896852 0.61724037 1.23448074],
[ 0.87290972 0.95896852 1.23448074 0.61724037]]
]
```
"""
we are passed x,y points and a selection of widths and heights
"""
with tf.variable_scope('ssd/select'):
l_feed = tf.placeholder(tf.float32, [None, None, None, None, 4], name="localizations")
p_feed = tf.placeholder(tf.float32, [None, None, None, None, 21], name="predictions")
d_pred = p_feed[:, :, :, :, 1:]
d_conditions = tf.greater(d_pred, 0.5)
d_chosen = tf.where(condition=d_conditions)
c_index = d_chosen[:,:-1]
x_feed = tf.placeholder(tf.float32, [None, None, None], name="x")
y_feed = tf.placeholder(tf.float32, [None, None, None], name="y")
h_feed = tf.placeholder(tf.float32, [None], name="h")
w_feed = tf.placeholder(tf.float32, [None], name="w")
box_shape = tf.shape(l_feed)
box_reshape = [-1, box_shape[-2], box_shape[-1]]
box_feat_localizations = tf.reshape(l_feed, box_reshape)
box_yref = tf.reshape(y_feed, [-1, 1])
box_xref = tf.reshape(x_feed, [-1, 1])
box_dx = box_feat_localizations[:, :, 0] * w_feed * 0.1 + box_xref
box_dy = box_feat_localizations[:, :, 1] * h_feed * 0.1 + box_yref
box_w = w_feed * tf.exp(box_feat_localizations[:, :, 2] * 0.2)
box_h = h_feed * tf.exp(box_feat_localizations[:, :, 3] * 0.2)
box_ymin = box_dy - box_h / 2.
box_xmin = box_dx - box_w / 2.
box_xmax = box_dy + box_h / 2.
box_ymax = box_dx + box_w / 2.
box_stack = tf.stack([box_ymin, box_xmin, box_xmax, box_ymax], axis=1)
box_transpose = tf.transpose(box_stack, [0,2,1])
box_gather_reshape = tf.reshape(box_transpose, box_shape, name="reshaping")
classes_selected = tf.cast(tf.transpose(d_chosen)[-1]+1, tf.float32)
classes_expand = tf.expand_dims(classes_selected, 1)
box_gather = tf.gather_nd(box_gather_reshape, c_index)
p_gather = tf.expand_dims(tf.gather_nd(d_pred, d_chosen), 1)
s_out = tf.concat([box_gather, p_gather, classes_expand], axis=1, name="output")
```
# Basic image input
get a local image and expand it to a 4d tensor
```
image_path = os.path.join('images/', 'street_smaller.jpg')
mean = tf.constant([123, 117, 104], dtype=tf.float32)
with tf.variable_scope('image'):
image_data = tf.gfile.FastGFile(image_path, 'rb').read()
#we want to use decode_image here but it's buggy
decoded = tf.image.decode_jpeg(image_data, channels=None)
normed = tf.divide(tf.cast(decoded, tf.float32), 255.0)
batched = tf.expand_dims(normed, 0)
resized_image = tf.image.resize_bilinear(batched, [299, 299])
standard_size = resized_image
graph_norm = standard_size * 255.0 - mean
with tf.Session() as image_session:
raw_image, file_image, plot_image = image_session.run((decoded, graph_norm, standard_size), feed_dict={})
# Main image processing routine.
predictions_net, localizations_net = ssd_session.run([predictions, localisations],
feed_dict={'ssd/input:0': file_image})
l_bboxes = []
for i in range(6):
box_feed = {l_feed: localizations_net[i], p_feed: predictions_net[i], \
y_feed: ssd_anchors[i][0], x_feed: ssd_anchors[i][1], \
h_feed: ssd_anchors[i][2], w_feed: ssd_anchors[i][3]}
bboxes = ssd_session.run([s_out], feed_dict=box_feed)
l_bboxes.append(bboxes[0])
bboxes = np.concatenate(l_bboxes, 0)
# implement these in frontend
# rclasses, rscores, rbboxes = np_methods.bboxes_sort(rclasses, rscores, rbboxes, top_k=400)
# rclasses, rscores, rbboxes = np_methods.bboxes_nms(rclasses, rscores, rbboxes, nms_threshold=nms_threshold)
print(predictions)
print(localisations)
print(bboxes)
from simple_heatmap import create_nms
create_nms()
with tf.variable_scope('gather'):
gather_indices = tf.placeholder(tf.int32, [None], name='indices')
gather_values = tf.placeholder(tf.float32, [None, 6], name='values')
gathered = tf.gather(gather_values, gather_indices, name='output')
nms_feed={'nms/bounds:0': bboxes, 'nms/threshold:0': [.8]}
pick = ssd_session.run(('nms/output:0'), feed_dict=nms_feed)
if bboxes.size>0 and pick.size>0:
gather_feed={'gather/indices:0': pick, 'gather/values:0': bboxes}
boxes = ssd_session.run(('gather/output:0'), feed_dict=gather_feed)
print(boxes)
import matplotlib.pyplot as plt
import matplotlib.patches as patches
import matplotlib.image as mpimg
fig, ax = plt.subplots(1)
show_image = np.reshape(plot_image, (299,299,3))
ax.imshow(raw_image)
print(raw_image.shape)
height = raw_image.shape[0]
width = raw_image.shape[1]
for box in boxes:
# Create a Rectangle patch
x = box[1] * width
y = box[0] * height
w = (box[3]-box[1]) * width
h = (box[2]-box[0]) * height
rect = patches.Rectangle((x,y),w,h,linewidth=3,edgecolor='r',facecolor='none')
# Add the patch to the Axes
ax.add_patch(rect)
plt.show()
from tensorflow.python.framework import graph_util
from tensorflow.python.training import saver as saver_lib
from tensorflow.core.protobuf import saver_pb2
checkpoint_prefix = os.path.join("checkpoints", "saved_checkpoint")
checkpoint_state_name = "checkpoint_state"
input_graph_name = "input_ssd_graph.pb"
output_graph_name = "ssd.pb"
input_graph_path = os.path.join("checkpoints", input_graph_name)
saver = saver_lib.Saver(write_version=saver_pb2.SaverDef.V2)
checkpoint_path = saver.save(
ssd_session,
checkpoint_prefix,
global_step=0,
latest_filename=checkpoint_state_name)
graph_def = ssd_session.graph.as_graph_def()
from tensorflow.python.lib.io import file_io
file_io.atomic_write_string_to_file(input_graph_path, str(graph_def))
print("wroteIt")
from tensorflow.python.tools import freeze_graph
input_saver_def_path = ""
input_binary = False
output_node_names = "ssd_300_vgg/softmax/Reshape_1,"+\
"ssd_300_vgg/softmax_1/Reshape_1,"+\
"ssd_300_vgg/softmax_2/Reshape_1,"+\
"ssd_300_vgg/softmax_3/Reshape_1,"+\
"ssd_300_vgg/softmax_4/Reshape_1,"+\
"ssd_300_vgg/softmax_5/Reshape_1,"+\
"ssd_300_vgg/block4_box/Reshape,"+\
"ssd_300_vgg/block7_box/Reshape,"+\
"ssd_300_vgg/block8_box/Reshape,"+\
"ssd_300_vgg/block9_box/Reshape,"+\
"ssd_300_vgg/block10_box/Reshape,"+\
"ssd_300_vgg/block11_box/Reshape,"+\
"ssd/priors/x,"+\
"ssd/priors/y,"+\
"gather/output,"+\
"nms/output,"+\
"ssd/select/output"
restore_op_name = "save/restore_all"
filename_tensor_name = "save/Const:0"
output_graph_path = os.path.join("data", output_graph_name)
clear_devices = False
freeze_graph.freeze_graph(input_graph_path, input_saver_def_path,
input_binary, checkpoint_path, output_node_names,
restore_op_name, filename_tensor_name,
output_graph_path, clear_devices, "")
```
| github_jupyter |
# Validation
## Splitting data
```
import numpy as np
X = np.array([
[1, 2],
[3, 4],
[5, 6],
[7, 8],
[9, 9],
[7, 7]
])
y = np.array([0, 0, 0, 1, 1, 1])
groups = np.array([0, 1, 2, 2, 1, 0])
```
### Group k-fold
```
from sklearn.model_selection import GroupKFold
gkf = GroupKFold(n_splits=3)
for train_index, test_index in gkf.split(X, y, groups):
print(f'TRAIN: {train_index}, TEST: {test_index}')
```
### Group shuffle split
```
from sklearn.model_selection import GroupShuffleSplit
gss = GroupShuffleSplit(n_splits=3, random_state=37)
for train_index, test_index in gss.split(X, y, groups):
print(f'TRAIN: {train_index}, TEST: {test_index}')
```
### Leave one group out
```
from sklearn.model_selection import LeaveOneGroupOut
logo = LeaveOneGroupOut()
for train_index, test_index in logo.split(X, y, groups):
print(f'TRAIN: {train_index}, TEST: {test_index}')
```
### Leave p-groups out
```
from sklearn.model_selection import LeavePGroupsOut
lpgo = LeavePGroupsOut(n_groups=2)
for train_index, test_index in lpgo.split(X, y, groups):
print(f'TRAIN: {train_index}, TEST: {test_index}')
```
### Leave one out
```
from sklearn.model_selection import LeaveOneOut
loo = LeaveOneOut()
for train_index, test_index in loo.split(X, y):
print(f'TRAIN: {train_index}, TEST: {test_index}')
```
### Leave p-out
```
from sklearn.model_selection import LeavePOut
lpo = LeavePOut(p=3)
for train_index, test_index in lpo.split(X, y):
print(f'TRAIN: {train_index}, TEST: {test_index}')
```
### K-fold
```
from sklearn.model_selection import KFold
kf = KFold(n_splits=2, random_state=37)
for train_index, test_index in kf.split(X, y):
print(f'TRAIN: {train_index}, TEST: {test_index}')
```
### Stratified k-fold
```
from sklearn.model_selection import StratifiedKFold
skf = StratifiedKFold(n_splits=2, random_state=37)
for train_index, test_index in skf.split(X, y):
print(f'TRAIN: {train_index}, TEST: {test_index}')
```
### Shuffle split
```
from sklearn.model_selection import ShuffleSplit
ss = ShuffleSplit(n_splits=2, random_state=37)
for train_index, test_index in ss.split(X, y):
print(f'TRAIN: {train_index}, TEST: {test_index}')
```
### Stratified shuffle split
```
from sklearn.model_selection import StratifiedShuffleSplit
sss = ShuffleSplit(n_splits=2, random_state=37)
for train_index, test_index in sss.split(X, y):
print(f'TRAIN: {train_index}, TEST: {test_index}')
```
## K-fold cross validation example
### Data
```
import numpy as np
from random import randint
from sklearn.datasets import make_classification
from sklearn.model_selection import StratifiedShuffleSplit
from collections import namedtuple
def get_data(n_features=20, n_samples=2000, n_missing=100):
def generate_coordinates(m, n):
seen = set()
x, y = randint(0, m - 1), randint(0, n - 1)
while True:
seen.add((x, y))
yield (x, y)
x, y = randint(0, m - 1), randint(0, n - 1)
while (x, y) in seen:
x, y = randint(0, m - 1), randint(0, n - 1)
def make_missing(X):
coords = generate_coordinates(n_samples, n_features)
for _ in range(n_missing):
i, j = next(coords)
X[i][j] = np.nan
X, y = make_classification(**{
'n_samples': n_samples,
'n_features': n_features,
'n_informative': 2,
'n_redundant': 2,
'n_repeated': 0,
'n_classes': 2,
'n_clusters_per_class': 2,
'random_state': 37
})
make_missing(X)
return X, y
np.random.seed(37)
X, y = get_data()
```
### Pipeline
```
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.decomposition import PCA
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import RandomForestClassifier
from sklearn.experimental import enable_iterative_imputer
from sklearn.impute import IterativeImputer
from sklearn.metrics import roc_auc_score, average_precision_score
def get_rf_pipeline():
imputer = IterativeImputer(missing_values=np.nan, random_state=37)
scaler = StandardScaler()
pca = PCA(n_components=3, random_state=37)
rf = RandomForestClassifier(n_estimators=100)
pipeline = Pipeline([
('imputer', imputer),
('scaler', scaler),
('pca', pca),
('rf', rf)
])
return pipeline
def get_lr_pipeline():
imputer = IterativeImputer(missing_values=np.nan, random_state=37)
scaler = StandardScaler()
lr = LogisticRegression(penalty='l1', solver='liblinear')
pipeline = Pipeline([
('imputer', imputer),
('scaler', scaler),
('lr', lr)
])
return pipeline
```
### Validation
```
import pandas as pd
def do_validation(train_index, test_index, X, y):
X_train, X_test = X[train_index], X[test_index]
y_train, y_test = y[train_index], y[test_index]
rf = get_rf_pipeline()
rf.fit(X_train, y_train)
y_preds = rf.predict_proba(X_test)[:,1]
rf_roc, rf_pr = roc_auc_score(y_test, y_preds), average_precision_score(y_test, y_preds)
lr = get_lr_pipeline()
lr.fit(X_train, y_train)
y_preds = lr.predict_proba(X_test)[:,1]
lr_roc, lr_pr = roc_auc_score(y_test, y_preds), average_precision_score(y_test, y_preds)
return rf_roc, lr_roc, rf_pr, lr_pr
kf = KFold(n_splits=10, random_state=37)
results = [do_validation(train_index, test_index, X, y)
for train_index, test_index in kf.split(X, y)]
df = pd.DataFrame(results, columns=['rf_roc', 'lr_roc', 'rf_pr', 'lr_pr'])
df
df.mean()
```
| github_jupyter |
## 内容概要
- 模型评估的目的及一般评估流程
- 分类准确率的用处及其限制
- 混淆矩阵(confusion matrix)是如何表示一个分类器的性能
- 混淆矩阵中的度量是如何计算的
- 通过改变分类阈值来调整分类器性能
- ROC曲线的用处
- 曲线下面积(Area Under the Curve, AUC)与分类准确率的不同
## 1. 回顾
模型评估可以用于在不同的模型类型、调节参数、特征组合中选择适合的模型,所以我们需要一个模型评估的流程来估计训练得到的模型对于非样本数据的泛化能力,并且还需要恰当的模型评估度量手段来衡量模型的性能表现。
对于模型评估流程而言,之前介绍了K折交叉验证的方法,针对模型评估度量方法,回归问题可以采用平均绝对误差(Mean Absolute Error)、均方误差(Mean Squared Error)、均方根误差(Root Mean Squared Error),而分类问题可以采用分类准确率和这篇文章中介绍的度量方法。
## 2. 分类准确率(Classification accuracy)
这里我们使用Pima Indians Diabetes dataset,其中包含健康数据和糖尿病状态数据,一共有768个病人的数据。
```
# read the data into a Pandas DataFrame
import pandas as pd
url = 'https://archive.ics.uci.edu/ml/machine-learning-databases/pima-indians-diabetes/pima-indians-diabetes.data'
col_names = ['pregnant', 'glucose', 'bp', 'skin', 'insulin', 'bmi', 'pedigree', 'age', 'label']
pima = pd.read_csv(url, header=None, names=col_names)
# print the first 5 rows of data
pima.head()
```
上面表格中的label一列,1表示该病人有糖尿病,0表示该病人没有糖尿病
```
# define X and y
feature_cols = ['pregnant', 'insulin', 'bmi', 'age']
X = pima[feature_cols]
y = pima.label
# split X and y into training and testing sets
from sklearn.cross_validation import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=0)
# train a logistic regression model on the training set
from sklearn.linear_model import LogisticRegression
logreg = LogisticRegression()
logreg.fit(X_train, y_train)
# make class predictions for the testing set
y_pred_class = logreg.predict(X_test)
# calculate accuracy
from sklearn import metrics
print metrics.accuracy_score(y_test, y_pred_class)
```
**分类准确率**分数是指所有分类正确的百分比。
**空准确率(null accuracy)**是指当模型总是预测比例较高的类别,那么其正确的比例是多少
```
# examine the class distribution of the testing set (using a Pandas Series method)
y_test.value_counts()
# calculate the percentage of ones
y_test.mean()
# calculate the percentage of zeros
1 - y_test.mean()
# calculate null accuracy(for binary classification problems coded as 0/1)
max(y_test.mean(), 1-y_test.mean())
```
我们看到空准确率是68%,而分类准确率是69%,这说明该分类准确率并不是很好的模型度量方法,**分类准确率的一个缺点是其不能表现任何有关测试数据的潜在分布。**
```
# calculate null accuracy (for multi-class classification problems)
y_test.value_counts().head(1) / len(y_test)
```
比较真实和预测的类别响应值:
```
# print the first 25 true and predicted responses
print "True:", y_test.values[0:25]
print "Pred:", y_pred_class[0:25]
```
从上面真实值和预测值的比较中可以看出,当正确的类别是0时,预测的类别基本都是0;当正确的类别是1时,预测的类别大都不是1。换句话说,该训练的模型大都在比例较高的那项类别的预测中预测正确,而在另外一中类别的预测中预测失败,而我们没法从分类准确率这项指标中发现这个问题。
分类准确率这一衡量分类器的标准比较容易理解,但是**它不能告诉你响应值的潜在分布,并且它也不能告诉你分类器犯错的类型**。接下来介绍的混淆矩阵可以识别这个问题。
## 3. 混淆矩阵
```
# IMPORTANT: first argument is true values, second argument is predicted values
print metrics.confusion_matrix(y_test, y_pred_class)
```

- 真阳性(True Positive,TP):指被分类器正确分类的正例数据
- 真阴性(True Negative,TN):指被分类器正确分类的负例数据
- 假阳性(False Positive,FP):被错误地标记为正例数据的负例数据
- 假阴性(False Negative,FN):被错误地标记为负例数据的正例数据
```
# save confusion matrix and slice into four pieces
confusion = metrics.confusion_matrix(y_test, y_pred_class)
TP = confusion[1, 1]
TN = confusion[0, 0]
FP = confusion[0, 1]
FN = confusion[1, 0]
print "TP:", TP
print "TN:", TN
print "FP:", FP
print "FN:", FN
```
## 4. 基于混淆矩阵的评估度量
**准确率、识别率(Classification Accuracy)**:分类器正确分类的比例
```
print (TP+TN) / float(TP+TN+FN+FP)
print metrics.accuracy_score(y_test, y_pred_class)
```
**错误率、误分类率(Classification Error)**:分类器误分类的比例
```
print (FP+FN) / float(TP+TN+FN+FP)
print 1-metrics.accuracy_score(y_test, y_pred_class)
```
考虑**类不平衡问题**,其中感兴趣的主类是稀少的。即数据集的分布反映负类显著地占多数,而正类占少数。故面对这种问题,需要其他的度量,评估分类器正确地识别正例数据的情况和正确地识别负例数据的情况。
**灵敏性(Sensitivity),也称为真正例识别率、召回率(Recall)**:正确识别的正例数据在实际正例数据中的百分比
```
print TP / float(TP+FN)
recall = metrics.recall_score(y_test, y_pred_class)
print metrics.recall_score(y_test, y_pred_class)
```
**特效性(Specificity),也称为真负例率**:正确识别的负例数据在实际负例数据中的百分比
```
print TN / float(TN+FP)
```
**假阳率(False Positive Rate)**:实际值是负例数据,预测错误的百分比
```
print FP / float(TN+FP)
specificity = TN / float(TN+FP)
print 1 - specificity
```
**精度(Precision)**:看做精确性的度量,即标记为正类的数据实际为正例的百分比
```
print TP / float(TP+FP)
precision = metrics.precision_score(y_test, y_pred_class)
print precision
```
**F度量(又称为F1分数或F分数)**,是使用精度和召回率的方法组合到一个度量上
$$F = \frac{2*precision*recall}{precision+recall}$$
$$F_{\beta} = \frac{(1+{\beta}^2)*precision*recall}{{\beta}^2*precision+recall}$$
$F$度量是精度和召回率的调和均值,它赋予精度和召回率相等的权重。
$F_{\beta}$度量是精度和召回率的加权度量,它赋予召回率权重是赋予精度的$\beta$倍。
```
print (2*precision*recall) / (precision+recall)
print metrics.f1_score(y_test, y_pred_class)
```
**总结**
混淆矩阵赋予一个分类器性能表现更全面的认识,同时它通过计算各种分类度量,指导你进行模型选择。
使用什么度量取决于具体的业务要求:
- 垃圾邮件过滤器:优先优化**精度**或者**特效性**,因为该应用对假阳性(非垃圾邮件被放进垃圾邮件箱)的要求高于对假阴性(垃圾邮件被放进正常的收件箱)的要求
- 欺诈交易检测器:优先优化**灵敏度**,因为该应用对假阴性(欺诈行为未被检测)的要求高于假阳性(正常交易被认为是欺诈)的要求
## 5. 调整分类的阈值
```
# print the first 10 predicted responses
logreg.predict(X_test)[0:10]
y_test.values[0:10]
# print the first 10 predicted probabilities of class membership
logreg.predict_proba(X_test)[0:10, :]
```
上面的输出中,第一列显示的是预测值为0的百分比,第二列显示的是预测值为1的百分比。
```
# print the first 10 predicted probabilities for class 1
logreg.predict_proba(X_test)[0:10, 1]
```
我们看到,预测为1的和实际的类别号差别很大,所以这里有50%作为分类的阈值显然不太合理。于是我们将所有预测类别为1的百分比数据用直方图的方式形象地表示出来,然后尝试重新设置阈值。
```
# store the predicted probabilities for class 1
y_pred_prob = logreg.predict_proba(X_test)[:, 1]
# allow plots to appear in the notebook
%matplotlib inline
import matplotlib.pyplot as plt
# histogram of predicted probabilities
plt.hist(y_pred_prob, bins=8)
plt.xlim(0, 1)
plt.title('Histogram of predicted probabilities')
plt.xlabel('Predicted probability of diabetes')
plt.ylabel('Frequency')
```
我们发现在20%-30%之间的数高达45%,故以50%作为分类阈值时,只有很少的一部分数据会被认为是类别为1的情况。我们可以将阈值调小,以改变分类器的**灵敏度和特效性**。
```
# predict diabetes if the predicted probability is greater than 0.3
from sklearn.preprocessing import binarize
y_pred_class = binarize(y_pred_prob, 0.3)[0]
# print the first 10 predicted probabilities
y_pred_prob[0:10]
# print the first 10 predicted classes with the lower threshold
y_pred_class[0:10]
y_test.values[0:10]
```
从上面两组数据对比来看,效果确实改善不少
```
# previous confusion matrix (default threshold of 0.5)
print confusion
# new confusion matrix (threshold of 0.3)
print metrics.confusion_matrix(y_test, y_pred_class)
# sensitivity has increased (used to be 0.24)
print 46 / float(46 + 16)
print metrics.recall_score(y_test, y_pred_class)
# specificity has decreased (used to be 0.91)
print 80 / float(80 + 50)
```
**总结:**
- 0.5作为阈值时默认的情况
- 调节阈值可以改变灵敏性和特效性
- 灵敏性和特效性是一对相反作用的指标
- 该阈值的调节是作为改善分类性能的最后一步,应更多去关注分类器的选择或构建更好的分类器
## 6. ROC曲线和AUC
ROC曲线指受试者工作特征曲线/接收器操作特性(receiver operating characteristic,ROC)曲线, 是反映灵敏性和特效性连续变量的综合指标,是用构图法揭示敏感性和特异性的相互关系,它通过将连续变量设定出多个不同的临界值,从而计算出一系列敏感性和特异性。
ROC曲线是根据一系列不同的二分类方式(分界值或决定阈),以真正例率(也就是灵敏度)(True Positive Rate,TPR)为纵坐标,假正例率(1-特效性)(False Positive Rate,FPR)为横坐标绘制的曲线。
**ROC观察模型正确地识别正例的比例与模型错误地把负例数据识别成正例的比例之间的权衡。TPR的增加以FPR的增加为代价。ROC曲线下的面积是模型准确率的度量。**
```
# IMPORTANT: first argument is true values, second argument is predicted probabilities
fpr, tpr, thresholds = metrics.roc_curve(y_test, y_pred_prob)
plt.plot(fpr, tpr)
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.0])
plt.title('ROC curve for diabetes classifier')
plt.xlabel('False Positive Rate (1 - Specificity)')
plt.ylabel('True Positive Rate (Sensitivity)')
plt.grid(True)
```
ROC曲线上的每一个点对应于一个threshold,对于一个分类器,每个threshold下会有一个TPR和FPR。
比如Threshold最大时,TP=FP=0,对应于原点;Threshold最小时,TN=FN=0,对应于右上角的点(1,1)
正如上面所述,TPR的增加以FPR的增加为代价,所以ROC曲线可以帮助我们选择一个可以平衡灵敏性和特效性的阈值。通过ROC曲线我们没法看到响应阈值的对应关系,所以我们用下面的函数来查看。
```
# define a function that accepts a threshold and prints sensitivity and specificity
def evaluate_threshold(threshold):
print 'Sensitivity:', tpr[thresholds > threshold][-1]
print 'Specificity:', 1 - fpr[thresholds > threshold][-1]
evaluate_threshold(0.5)
evaluate_threshold(0.3)
```
AUC(Area Under Curve)被定义为ROC曲线下的面积,也可以认为是ROC曲线下面积占单位面积的比例,显然这个面积的数值不会大于1。又由于ROC曲线一般都处于y=x这条直线的上方,所以AUC的取值范围在0.5和1之间。
**对应AUC更大的分类器效果更好。**所以AUC是衡量分类器性能的一个很好的度量,并且它不像分类准确率那样,在类别比例差别很大的情况下,依然是很好的度量手段。在欺诈交易检测中,由于欺诈案例是很小的一部分,这时分类准确率就不再是一个良好的度量,而可以使用AUC来度量。
```
# IMPORTANT: first argument is true values, second argument is predicted probabilities
print metrics.roc_auc_score(y_test, y_pred_prob)
# calculate cross-validated AUC
from sklearn.cross_validation import cross_val_score
cross_val_score(logreg, X, y, cv=10, scoring='roc_auc').mean()
```
## 参考资料
- scikit-learn documentation: [Model evaluation](http://scikit-learn.org/stable/modules/model_evaluation.html)
- [ROC曲线-阈值评价标准](http://blog.csdn.net/abcjennifer/article/details/7359370)
| github_jupyter |
# This notebook provides the functionality to build, train, and test a CNN for predicting mosquito age, grouped age, species, and status.
## Structure:
* Import packages to be used.
* Load mosquito data.
* Define fucntions for plotting, visualisation, and logging.
* Define a function to build the CNN.
* Define a function to train the CNN.
* Main section to organise data, define the CNN, and call the building and training of the CNN.
```
import pylab as pl
import datetime
import pandas as pd
import itertools
from itertools import cycle
import pickle
import random as rn
import os
from time import time
from tqdm import tqdm
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
from matplotlib.lines import Line2D
from sklearn.preprocessing import MultiLabelBinarizer
from sklearn.model_selection import train_test_split, KFold
from sklearn.metrics import confusion_matrix
from sklearn.preprocessing import normalize, StandardScaler
from sklearn.utils import resample
import tensorflow as tf
import keras
from keras.models import Sequential, Model
from keras import layers, metrics
from keras.layers import Input
from keras.layers.merge import Concatenate
from keras.layers.core import Dense, Dropout, Activation, Flatten
from keras.layers.normalization import BatchNormalization
from keras.layers.convolutional import Conv1D, MaxPooling1D
from keras.models import model_from_json, load_model
from keras.regularizers import *
from keras.callbacks import CSVLogger
from keras import backend as K
# rand_seed = np.random.randint(low=0, high=100)
rand_seed = 16
print(rand_seed)
os.environ['PYTHONHASHSEED'] = '0'
## The below is necessary for starting Numpy generated random numbers in a well-defined initial state.
np.random.seed(42)
## The below is necessary for starting core Python generated random numbers in a well-defined state.
rn.seed(12345)
## Force TensorFlow to use single thread.
## Multiple threads are a potential source of
## non-reproducible results.
## For further details, see: https://stackoverflow.com/questions/42022950/which-seeds-have-to-be-set-where-to-realize-100-reproducibility-of-training-res
# session_conf = tf.ConfigProto(device_count = {'GPU':0}, intra_op_parallelism_threads=4) #session_conf = tf.ConfigProto(intra_op_parallelism_threads=1, inter_op_parallelism_threads=1)
# session_conf = tf.ConfigProto(device_count = {'GPU':0}) #session_conf = tf.ConfigProto(intra_op_parallelism_threads=1, inter_op_parallelism_threads=1)
#session_conf.gpu_options.per_process_gpu_memory_fraction = 0.5
## The below tf.set_random_seed() will make random number generation
## in the TensorFlow backend have a well-defined initial state.
## For further details, see: https://www.tensorflow.org/api_docs/python/tf/set_random_seed
tf.set_random_seed(1234)
gpu_options = tf.GPUOptions(per_process_gpu_memory_fraction=0.35)
sess = tf.Session(graph=tf.get_default_graph(), config=tf.ConfigProto(gpu_options=gpu_options))
K.set_session(sess)
```
## Function used to create a new folder for the CNN outputs.
Useful to stop forgetting to name a new folder when trying out a new model varient and overwriting a days training.
```
def build_folder(fold, to_build = False):
if not os.path.isdir(fold):
if to_build == True:
os.mkdir(fold)
else:
print('Directory does not exists, not creating directory!')
else:
if to_build == True:
raise NameError('Directory already exists, cannot be created!')
```
## Function for plotting confusion matrcies
This normalizes the confusion matrix and ensures neat plotting for all outputs.
```
def plot_confusion_matrix(cm, classes, output, save_path, model_name, fold,
normalize=True,
title='Confusion matrix',
cmap=plt.cm.Blues,
printout=False):
font = {'weight' : 'normal',
'size' : 18}
matplotlib.rc('font', **font)
"""
This function prints and plots the confusion matrix.
Normalization can be applied by setting `normalize=True`.
"""
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
if printout:
print("Normalized confusion matrix")
else:
if printout:
print('Confusion matrix, without normalization')
if printout:
print(cm)
plt.figure(figsize=(8,8))
plt.imshow(cm, interpolation='nearest', cmap=cmap, vmin=0, vmax=1) # np.max(np.sum(cm, axis=1)))
# plt.title([title+' - '+model_name])
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
fmt = '.2f' if normalize else 'd'
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, format(cm[i, j], fmt),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.tight_layout(pad=2)
# plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
plt.savefig((save_path+"Confusion_Matrix_"+model_name+"_"+fold+"_"+output[1:]+".png"))
plt.close()
```
## Function used for visualizing outputs
This splits the output data into the four categories before plotting the confusion matricies.
```
## for visualizing losses and metrics once the neural network fold is trained
def visualize(histories, save_path, model_name, fold, classes, outputs, predicted, true):
# Sort out predictions and true labels
for label_predictions_arr, label_true_arr, classes, outputs in zip(predicted, true, classes, outputs):
classes_pred = np.argmax(label_predictions_arr, axis=-1)
classes_true = np.argmax(label_true_arr, axis=-1)
cnf_matrix = confusion_matrix(classes_true, classes_pred)
plot_confusion_matrix(cnf_matrix, classes, outputs, save_path, model_name, fold)
```
## Data logging
```
## for logging data associated with the model
def log_data(log, name, fold, save_path):
f = open((save_path+name+'_'+str(fold)+'_log.txt'), 'w')
np.savetxt(f, log)
f.close()
```
## Fucntion for graphing the training data
This fucntion creates tidy graphs of loss and accuracy as the models are training.
```
def graph_history(history, model_name, model_ver_num, fold, save_path):
font = {'weight' : 'normal',
'size' : 18}
matplotlib.rc('font', **font)
#not_validation = list(filter(lambda x: x[0:3] != "val", history.history.keys()))
# print('history.history.keys : {}'.format(history.history.keys()))
filtered = filter(lambda x: x[0:3] != "val", history.history.keys())
not_validation = list(filtered)
for i in not_validation:
plt.figure(figsize=(15,7))
# plt.title(i+"/ "+"val_"+i)
plt.plot(history.history[i], label=i)
plt.plot(history.history["val_"+i], label="val_"+i)
plt.legend()
plt.xlabel("epoch")
plt.ylabel(i)
plt.savefig(save_path +model_name+"_"+str(model_ver_num)+"_"+str(fold)+"_"+i)
plt.close()
```
## funciton to create the CNN
This function takes as an input a list of dictionaries. Each element in the list is a new hidden layer in the model. For each layer the dictionary defines the layer to be used.
### Available options are:
Convolutional Layer:
* type = 'c'
* filter = optional number of filters
* kernel = optional size of the filters
* stride = optional size of stride to take between filters
* pooling = optional width of the max pooling
* {'type':'c', 'filter':16, 'kernel':5, 'stride':1, 'pooling':2}
dense layer:
* type = 'd'
* width = option width of the layer
* {'type':'d', 'width':500}
```
def create_models(model_shape, input_layer_dim):
regConst = 0.02
sgd = keras.optimizers.SGD(lr=0.003, decay=1e-5, momentum=0.9, nesterov=True, clipnorm=1.)
cce = 'categorical_crossentropy'
input_vec = Input(name='input', shape=(input_layer_dim,1))
for i, layerwidth in zip(range(len(model_shape)),model_shape):
if i == 0:
if model_shape[i]['type'] == 'c':
xd = Conv1D(name=('Conv'+str(i+1)), filters=model_shape[i]['filter'],
kernel_size = model_shape[i]['kernel'], strides = model_shape[i]['stride'],
activation = 'relu',
kernel_regularizer=l2(regConst),
kernel_initializer='he_normal')(input_vec)
xd = BatchNormalization(name=('batchnorm_'+str(i+1)))(xd)
xd = MaxPooling1D(pool_size=(model_shape[i]['pooling']))(xd)
elif model_shape[i]['type'] == 'd':
xd = Dense(name=('d'+str(i+1)), units=model_shape[i]['width'], activation='relu',
kernel_regularizer=l2(regConst),
kernel_initializer='he_normal')(input_vec)
xd = BatchNormalization(name=('batchnorm_'+str(i+1)))(xd)
xd = Dropout(name=('dout'+str(i+1)), rate=0.5)(xd)
else:
if model_shape[i]['type'] == 'c':
xd = Conv1D(name=('Conv'+str(i+1)), filters=model_shape[i]['filter'],
kernel_size = model_shape[i]['kernel'], strides = model_shape[i]['stride'],
activation = 'relu',
kernel_regularizer=l2(regConst),
kernel_initializer='he_normal')(xd)
xd = BatchNormalization(name=('batchnorm_'+str(i+1)))(xd)
xd = MaxPooling1D(pool_size=(model_shape[i]['pooling']))(xd)
elif model_shape[i]['type'] == 'd':
if model_shape[i-1]['type'] == 'c':
xd = Flatten()(xd)
xd = Dropout(name=('dout'+str(i+1)), rate=0.5)(xd)
xd = Dense(name=('d'+str(i+1)), units=model_shape[i]['width'], activation='relu',
kernel_regularizer=l2(regConst),
kernel_initializer='he_normal')(xd)
xd = BatchNormalization(name=('batchnorm_'+str(i+1)))(xd)
# xAge = Dense(name = 'age', units = 17,
# activation = 'softmax',
# kernel_regularizer = l2(regConst),
# kernel_initializer = 'he_normal')(xd)
xAgeGroup = Dense(name = 'age_group', units = 3,
activation = 'softmax',
kernel_regularizer = l2(regConst),
kernel_initializer = 'he_normal')(xd)
xSpecies = Dense(name ='species', units = 3,
activation = 'softmax',
kernel_regularizer = l2(regConst),
kernel_initializer = 'he_normal')(xd)
outputs = []
# for i in ['xAge', 'xAgeGroup', 'xSpecies']:
for i in ['xAgeGroup', 'xSpecies']:
outputs.append(locals()[i])
model = Model(inputs = input_vec, outputs = outputs)
model.compile(loss=cce, metrics=['acc'],
optimizer=sgd)
# model.summary()
return model
```
## Function to train the model
This function will split the data into training and validation and call the create models function. This fucntion returns the model and training history.
```
def train_models(model_to_test, save_path, SelectFreqs=False):
model_shape = model_to_test["model_shape"][0]
model_name = model_to_test["model_name"][0]
model_ver_num = model_to_test["model_ver_num"][0]
fold = model_to_test["fold"][0]
label = model_to_test["labels"][0]
features = model_to_test["features"][0]
classes = model_to_test["classes"][0]
outputs = model_to_test["outputs"][0]
compile_loss = model_to_test["compile_loss"][0]
compile_metrics = model_to_test["compile_metrics"][0]
## Kfold training
seed = rand_seed
kfold = KFold(n_splits=10, shuffle=True, random_state=seed)
model_ver_num = 0
cv_scores = []
best_score = 0
for train_index, val_index in kfold.split(features):
print('Fold {} Running'.format(model_ver_num))
X_train, X_val = features[train_index], features[val_index]
y_train, y_val = list(map(lambda y:y[train_index], label)), list(map(lambda y:y[val_index], label))
model = create_models(model_shape, input_layer_dim)
if model_ver_num == 0:
model.summary()
history = model.fit(x = X_train,
y = y_train,
batch_size = 128*16,
verbose = 0,
epochs = 8000,
validation_data = (X_val, y_val),
callbacks = [keras.callbacks.EarlyStopping(monitor='val_loss',
patience=400, verbose=0, mode='auto'),
CSVLogger(save_path+model_name+"_"+str(model_ver_num)+'.csv', append=True, separator=';')])
scores = model.evaluate(X_val, y_val)
if (scores[3] + scores[4]) > best_score:
out_model = model
out_history = history
model_ver_num += 1
# # Clear the Keras session, otherwise it will keep adding new
# # models to the same TensorFlow graph each time we create
# # a model with a different set of hyper-parameters.
# K.clear_session()
# # Delete the Keras model with these hyper-parameters from memory.
# del model
out_model.save((save_path+model_name+"_"+'Model.h5'))
graph_history(out_history, model_name, 0, 0, save_path)
return out_model, out_history
```
## Train Tanzania + Burkino Faso field + Tanzania semi-field
## Test Burkino Fasa semi-field
## Load the data
The data file is created using Loco Mosquito:
https://github.com/magonji/MIMI-project/blob/master/Loco%20mosquito%204.0.ipynb
### The data file has headings: Species - Status - RearCnd - Age - Country- Frequencies
```
df = pd.read_csv("/home/josh/Documents/Mosquito_Project/New_Data/Data/MIMIdata_update_19_02/mosquitoes_country_LM_5_0.dat", '\t')
df.head(10)
RearCnd_counts = df.groupby('RearCnd').size()
df['AgeGroup'] = 0
df['AgeGroup'] = np.where(df['Age']>10, 2, np.where(df['Age']>4, 1, 0))
df_vf = df[df['RearCnd']=='VF']
df_vf = df_vf[df_vf['Status']=='UN']
df = df[df['RearCnd']!='VF']
df = df[df['Status']!='UN']
df_l = df[df['RearCnd']=='TL']
df_l_g = df_l[df_l['Country']=='S']
df_l_g_a = df_l_g[df_l_g['Species']=='AA']
age_counts = df_l_g_a.groupby('AgeGroup').size()
df_l_g_g = df_l_g[df_l_g['Species']=='AG']
age_counts = df_l_g_g.groupby('AgeGroup').size()
df_l_g_c = df_l_g[df_l_g['Species']=='AC']
age_counts = df_l_g_c.groupby('AgeGroup').size()
df_l_t = df_l[df_l['Country']=='T']
df_l_t_a = df_l_t[df_l_t['Species']=='AA']
age_counts = df_l_t_a.groupby('AgeGroup').size()
df_l_t_g = df_l_t[df_l_t['Species']=='AG']
age_counts = df_l_t_g.groupby('AgeGroup').size()
df_l_b = df_l[df_l['Country']=='B']
df_l_b_g = df_l_b[df_l_b['Species']=='AG']
age_counts = df_l_b_g.groupby('AgeGroup').size()
df_l_b_c = df_l_b[df_l_b['Species']=='AC']
age_counts = df_l_b_c.groupby('AgeGroup').size()
df_f = df[df['RearCnd']=='TF']
df_f_t = df_f[df_f['Country']=='T']
df_f_t_a = df_f_t[df_f_t['Species']=='AA']
age_counts = df_f_t_a.groupby('AgeGroup').size()
# df_f_t_g = df_f_t[df_f_t['Species']=='AG'] #There isn't any
df_f_b = df_f[df_f['Country']=='B']
df_f_b_g = df_f_b[df_f_b['Species']=='AG']
age_counts = df_f_b_g.groupby('AgeGroup').size()
df_f_b_c = df_f_b[df_f_b['Species']=='AC']
age_counts = df_f_b_c.groupby('AgeGroup').size()
df_vf_t = df_vf[df_vf['Country']=='T']
df_vf_t_a = df_vf_t[df_vf_t['Species']=='AA']
age_counts = df_vf_t_a.groupby('AgeGroup').size()
print(f'Size of df_vf_t_a {age_counts}')
df_vf_t_g = df_vf_t[df_vf_t['Species']=='AG']
age_counts = df_vf_t_g.groupby('AgeGroup').size()
print(f'Size of df_vf_t_g {age_counts}')
df_vf_b = df_vf[df_vf['Country']=='B']
df_vf_b_g = df_vf_b[df_vf_b['Species']=='AG']
age_counts = df_vf_b_g.groupby('AgeGroup').size()
print(f'Size of df_vf_b_g {age_counts}')
df_vf_b_c = df_vf_b[df_vf_b['Species']=='AC']
age_counts = df_vf_b_c.groupby('AgeGroup').size()
print(f'Size of df_vf_b_c {age_counts}')
# size_inc = 400
# for age in range(3):
# df_temp = df_l_t_a[df_l_t_a['AgeGroup']==age]
# size_df_temp = np.arange(len(df_temp))
# np.random.seed(42)
# np.random.shuffle(size_df_temp)
# index_df_temp_inc = size_df_temp[:size_inc]
# index_df_temp_not_inc = size_df_temp[size_inc:]
# if age == 0:
# df_train = df_temp.iloc[index_df_temp_inc]
# # df_test = df_temp.iloc[index_df_temp_not_inc]
# else:
# df_train = pd.concat([df_train, df_temp.iloc[index_df_temp_inc]])
# # df_test = pd.concat([df_test, df_temp.iloc[index_df_temp_not_inc]])
# for age in range(3):
# df_temp = df_l_t_g[df_l_t_g['AgeGroup']==age]
# size_df_temp = np.arange(len(df_temp))
# np.random.seed(42)
# np.random.shuffle(size_df_temp)
# index_df_temp_inc = size_df_temp[:size_inc]
# index_df_temp_not_inc = size_df_temp[size_inc:]
# df_train = pd.concat([df_train, df_temp.iloc[index_df_temp_inc]])
# # df_test = pd.concat([df_test, df_temp.iloc[index_df_temp_not_inc]])
# size_inc = 400
# for age in range(3):
# df_temp = df_l_b_g[df_l_b_g['AgeGroup']==age]
# size_df_temp = np.arange(len(df_temp))
# np.random.seed(42)
# np.random.shuffle(size_df_temp)
# index_df_temp_inc = size_df_temp[:size_inc]
# index_df_temp_not_inc = size_df_temp[size_inc:]
# df_train = pd.concat([df_train, df_temp.iloc[index_df_temp_inc]])
# # df_test = pd.concat([df_test, df_temp.iloc[index_df_temp_not_inc]])
# for age in range(3):
# df_temp = df_l_b_c[df_l_b_c['AgeGroup']==age]
# size_df_temp = np.arange(len(df_temp))
# np.random.seed(42)
# np.random.shuffle(size_df_temp)
# index_df_temp_inc = size_df_temp[:size_inc]
# index_df_temp_not_inc = size_df_temp[size_inc:]
# df_train = pd.concat([df_train, df_temp.iloc[index_df_temp_inc]])
# # df_test = pd.concat([df_test, df_temp.iloc[index_df_temp_not_inc]])
size_inc = 300 # 50
for age in range(3):
df_temp = df_f_t_a[df_f_t_a['AgeGroup']==age]
size_df_temp = np.arange(len(df_temp))
np.random.seed(42)
np.random.shuffle(size_df_temp)
index_df_temp_inc = size_df_temp[:size_inc]
index_df_temp_not_inc = size_df_temp[size_inc:]
if age == 0:
df_train = df_temp.iloc[index_df_temp_inc]
# df_test = df_temp.iloc[index_df_temp_not_inc]
else:
df_train = pd.concat([df_train, df_temp.iloc[index_df_temp_inc]])
# df_test = pd.concat([df_test, df_temp.iloc[index_df_temp_not_inc]])
for age in range(3):
df_temp = df_f_b_g[df_f_b_g['AgeGroup']==age]
size_df_temp = np.arange(len(df_temp))
np.random.seed(42)
np.random.shuffle(size_df_temp)
index_df_temp_inc = size_df_temp[:size_inc]
index_df_temp_not_inc = size_df_temp[size_inc:]
df_train = pd.concat([df_train, df_temp.iloc[index_df_temp_inc]])
# df_test = pd.concat([df_test, df_temp.iloc[index_df_temp_not_inc]])
for age in range(3):
df_temp = df_f_b_c[df_f_b_c['AgeGroup']==age]
size_df_temp = np.arange(len(df_temp))
np.random.seed(42)
np.random.shuffle(size_df_temp)
index_df_temp_inc = size_df_temp[:size_inc]
index_df_temp_not_inc = size_df_temp[size_inc:]
df_train = pd.concat([df_train, df_temp.iloc[index_df_temp_inc]])
# df_test = pd.concat([df_test, df_temp.iloc[index_df_temp_not_inc]])
size_inc = 200
for age in range(3):
df_temp = df_vf_t_a[df_vf_t_a['AgeGroup']==age]
size_df_temp = np.arange(len(df_temp))
np.random.seed(42)
np.random.shuffle(size_df_temp)
index_df_temp_inc = size_df_temp[:size_inc]
index_df_temp_not_inc = size_df_temp[size_inc:]
df_train = pd.concat([df_train, df_temp.iloc[index_df_temp_inc]])
# df_test = pd.concat([df_test, df_temp.iloc[index_df_temp_not_inc]])
for age in range(3):
df_temp = df_vf_t_g[df_vf_t_g['AgeGroup']==age]
size_df_temp = np.arange(len(df_temp))
np.random.seed(42)
np.random.shuffle(size_df_temp)
index_df_temp_inc = size_df_temp[:size_inc]
index_df_temp_not_inc = size_df_temp[size_inc:]
df_train = pd.concat([df_train, df_temp.iloc[index_df_temp_inc]])
# df_test = pd.concat([df_test, df_temp.iloc[index_df_temp_not_inc]])
size_inc = 0
for age in range(3):
df_temp = df_vf_b_g[df_vf_b_g['AgeGroup']==age]
size_df_temp = np.arange(len(df_temp))
np.random.seed(42)
np.random.shuffle(size_df_temp)
index_df_temp_inc = size_df_temp[:size_inc]
index_df_temp_not_inc = size_df_temp[size_inc:]
if age == 0:
# df_train = df_temp.iloc[index_df_temp_inc]
df_test = df_temp.iloc[index_df_temp_not_inc]
else:
# df_train = pd.concat([df_train, df_temp.iloc[index_df_temp_inc]])
df_test = pd.concat([df_test, df_temp.iloc[index_df_temp_not_inc]])
for age in range(3):
df_temp = df_vf_b_c[df_vf_b_c['AgeGroup']==age]
size_df_temp = np.arange(len(df_temp))
np.random.seed(42)
np.random.shuffle(size_df_temp)
index_df_temp_inc = size_df_temp[:size_inc]
index_df_temp_not_inc = size_df_temp[size_inc:]
# df_train = pd.concat([df_train, df_temp.iloc[index_df_temp_inc]])
df_test = pd.concat([df_test, df_temp.iloc[index_df_temp_not_inc]])
X = df_train.iloc[:,6:-1]
y_age = df_train["Age"]
y_age_groups = df_train["AgeGroup"]
y_species = df_train["Species"]
y_status = df_train["Status"]
X = np.asarray(X)
y_age = np.asarray(y_age)
y_age_groups = np.asarray(y_age_groups)
y_species = np.asarray(y_species)
y_status = np.asarray(y_status)
print('shape of X : {}'.format(X.shape))
print('shape of y age : {}'.format(y_age.shape))
print('shape of y age groups : {}'.format(y_age_groups.shape))
print('shape of y species : {}'.format(y_species.shape))
print('shape of y status : {}'.format(y_status.shape))
X_vf = df_test.iloc[:,6:-1]
y_age_vf = df_test["Age"]
y_age_groups_vf = df_test["AgeGroup"]
y_species_vf = df_test["Species"]
y_status_vf = df_test["Status"]
X_vf = np.asarray(X_vf)
y_age_vf = np.asarray(y_age_vf)
y_age_groups_vf = np.asarray(y_age_groups_vf)
y_species_vf = np.asarray(y_species_vf)
y_status_vf = np.asarray(y_status_vf)
print('shape of X vf : {}'.format(X_vf.shape))
print('shape of y age vf : {}'.format(y_age_vf.shape))
print('shape of y age groups vf : {}'.format(y_age_groups_vf.shape))
print('shape of y species vf : {}'.format(y_species_vf.shape))
print('shape of y status vf : {}'.format(y_status_vf.shape))
```
## Main section
Functionality:
* Oganises the data into a format of lists of data, classes, labels.
* Define the CNN to be built.
* Define the KFold validation to be used.
* Build a folder to output data into.
* Standardize and oragnise data into training/testing.
* Call the model training.
* Organize outputs and call visualization for plotting and graphing.
```
input_layer_dim = len(Xf[0])
y_age_groups_list_l = [[age] for age in y_age_groups]
y_species_list_l = [[species] for species in y_species]
age_groups_l = MultiLabelBinarizer().fit_transform(np.array(y_age_groups_list_l))
age_group_classes = ["1-4", "5-10", "11-17"]
species_l = MultiLabelBinarizer().fit_transform(np.array(y_species_list_l))
species_classes = list(np.unique(y_species_list_l))
y_age_groups_list_vf = [[age] for age in y_age_groups_vf]
y_species_list_vf = [[species] for species in y_species_vf]
age_groups_vf = MultiLabelBinarizer().fit_transform(np.array(y_age_groups_list_vf))
species_vf = MultiLabelBinarizer().fit_transform(np.array(y_species_list_vf))
outdir = "Results_Paper/"
build_folder(outdir, False)
SelectFreqs = False
## Labels default - all classification
labels_default_vf, labels_default_l, classes_default, outputs_default = [age_groups_vf, species_vf], [age_groups_l, species_l], [age_group_classes, species_classes], ['xAgeGroup', 'xSpecies']
## Declare and train the model
model_size = [{'type':'c', 'filter':16, 'kernel':8, 'stride':1, 'pooling':1},
{'type':'c', 'filter':16, 'kernel':8, 'stride':2, 'pooling':1},
{'type':'c', 'filter':16, 'kernel':3, 'stride':1, 'pooling':1},
{'type':'c', 'filter':16, 'kernel':6, 'stride':2, 'pooling':1},
{'type':'c', 'filter':16, 'kernel':5, 'stride':1, 'pooling':2},
{'type':'d', 'width':500}]
## Name the model
model_name = 'Baseline_CNN'
histories = []
fold = 1
train_model = True
## Name a folder for the outputs to go into
savedir = (outdir+"Trian_TCField_No_Bobo/")
build_folder(savedir, True)
start_time = time()
save_predicted = []
save_true = []
## Scale train, test
scl = StandardScaler()
features_scl = scl.fit(X=np.vstack((X,X_vf)))
features_l = features_scl.transform(X=X)
features_vf = features_scl.transform(X=X_vf)
## Split into training / testing
# test_splits = train_test_split(features_f, *(labels_default_f), test_size=0.1, shuffle=True, random_state=rand_seed)
## Pack up data
X_train = features_l
X_test = features_vf
y_train = labels_default_l
y_test = labels_default_vf
if not SelectFreqs:
X_train = np.expand_dims(X_train, axis=2)
X_test = np.expand_dims(X_test, axis=2)
model_to_test = {
"model_shape" : [model_size], # defines the hidden layers of the model
"model_name" : [model_name],
"input_layer_dim" : [input_layer_dim], # size of input layer
"model_ver_num" : [0],
"fold" : [fold], # kf.split number on
"labels" : [y_train],
"features" : [X_train],
"classes" : [classes_default],
"outputs" : [outputs_default],
"compile_loss": [{'age': 'categorical_crossentropy'}],
"compile_metrics" :[{'age': 'accuracy'}]
}
## Call function to train all the models from the dictionary
model, history = train_models(model_to_test, savedir, SelectFreqs=SelectFreqs)
histories.append(history)
predicted_labels = list([] for i in range(len(y_train)))
true_labels = list([] for i in range(len(y_train)))
y_predicted = model.predict(X_test)
predicted_labels = [x+[y] for x,y in zip(predicted_labels,y_predicted)]
true_labels = [x+[y] for x,y in zip(true_labels,y_test)]
predicted_labels = [predicted_labels[i][0].tolist() for i in range(len(predicted_labels))]
true_labels = [true_labels[i][0].tolist() for i in range(len(true_labels))]
for pred, tru in zip(predicted_labels, true_labels):
save_predicted.append(pred)
save_true.append(tru)
## Visualize the results
visualize(histories, savedir, model_name, str(fold), classes_default, outputs_default, predicted_labels, true_labels)
# log_data(test_index, 'test_index', fold, savedir)
# Clear the Keras session, otherwise it will keep adding new
# models to the same TensorFlow graph each time we create
# a model with a different set of hyper-parameters.
K.clear_session()
# Delete the Keras model with these hyper-parameters from memory.
del model
# visualize(1, savedir, model_name, "Averaged", classes_default, outputs_default, save_predicted, save_true)
end_time = time()
print('Run time : {} s'.format(end_time-start_time))
print('Run time : {} m'.format((end_time-start_time)/60))
print('Run time : {} h'.format((end_time-start_time)/3600))
```
## Train Tanzania + Burkino Faso field + Burkino Faso semi-field
## Test Burkino Faso semi-field
```
df = pd.read_csv("/home/josh/Documents/Mosquito_Project/New_Data/Data/MIMIdata_update_19_02/mosquitoes_country_LM_5_0.dat", '\t')
df.head(10)
RearCnd_counts = df.groupby('RearCnd').size()
df['AgeGroup'] = 0
df['AgeGroup'] = np.where(df['Age']>10, 2, np.where(df['Age']>4, 1, 0))
df_vf = df[df['RearCnd']=='VF']
df_vf = df_vf[df_vf['Status']=='UN']
df = df[df['RearCnd']!='VF']
df = df[df['Status']!='UN']
df_l = df[df['RearCnd']=='TL']
df_l_g = df_l[df_l['Country']=='S']
df_l_g_a = df_l_g[df_l_g['Species']=='AA']
age_counts = df_l_g_a.groupby('AgeGroup').size()
df_l_g_g = df_l_g[df_l_g['Species']=='AG']
age_counts = df_l_g_g.groupby('AgeGroup').size()
df_l_g_c = df_l_g[df_l_g['Species']=='AC']
age_counts = df_l_g_c.groupby('AgeGroup').size()
df_l_t = df_l[df_l['Country']=='T']
df_l_t_a = df_l_t[df_l_t['Species']=='AA']
age_counts = df_l_t_a.groupby('AgeGroup').size()
df_l_t_g = df_l_t[df_l_t['Species']=='AG']
age_counts = df_l_t_g.groupby('AgeGroup').size()
df_l_b = df_l[df_l['Country']=='B']
df_l_b_g = df_l_b[df_l_b['Species']=='AG']
age_counts = df_l_b_g.groupby('AgeGroup').size()
df_l_b_c = df_l_b[df_l_b['Species']=='AC']
age_counts = df_l_b_c.groupby('AgeGroup').size()
df_f = df[df['RearCnd']=='TF']
df_f_t = df_f[df_f['Country']=='T']
df_f_t_a = df_f_t[df_f_t['Species']=='AA']
age_counts = df_f_t_a.groupby('AgeGroup').size()
# df_f_t_g = df_f_t[df_f_t['Species']=='AG'] #There isn't any
df_f_b = df_f[df_f['Country']=='B']
df_f_b_g = df_f_b[df_f_b['Species']=='AG']
age_counts = df_f_b_g.groupby('AgeGroup').size()
df_f_b_c = df_f_b[df_f_b['Species']=='AC']
age_counts = df_f_b_c.groupby('AgeGroup').size()
df_vf_t = df_vf[df_vf['Country']=='T']
df_vf_t_a = df_vf_t[df_vf_t['Species']=='AA']
age_counts = df_vf_t_a.groupby('AgeGroup').size()
print(f'Size of df_vf_t_a {age_counts}')
df_vf_t_g = df_vf_t[df_vf_t['Species']=='AG']
age_counts = df_vf_t_g.groupby('AgeGroup').size()
print(f'Size of df_vf_t_g {age_counts}')
df_vf_b = df_vf[df_vf['Country']=='B']
df_vf_b_g = df_vf_b[df_vf_b['Species']=='AG']
age_counts = df_vf_b_g.groupby('AgeGroup').size()
print(f'Size of df_vf_b_g {age_counts}')
df_vf_b_c = df_vf_b[df_vf_b['Species']=='AC']
age_counts = df_vf_b_c.groupby('AgeGroup').size()
print(f'Size of df_vf_b_c {age_counts}')
# size_inc = 400
# for age in range(3):
# df_temp = df_l_t_a[df_l_t_a['AgeGroup']==age]
# size_df_temp = np.arange(len(df_temp))
# np.random.seed(42)
# np.random.shuffle(size_df_temp)
# index_df_temp_inc = size_df_temp[:size_inc]
# index_df_temp_not_inc = size_df_temp[size_inc:]
# if age == 0:
# df_train = df_temp.iloc[index_df_temp_inc]
# # df_test = df_temp.iloc[index_df_temp_not_inc]
# else:
# df_train = pd.concat([df_train, df_temp.iloc[index_df_temp_inc]])
# # df_test = pd.concat([df_test, df_temp.iloc[index_df_temp_not_inc]])
# for age in range(3):
# df_temp = df_l_t_g[df_l_t_g['AgeGroup']==age]
# size_df_temp = np.arange(len(df_temp))
# np.random.seed(42)
# np.random.shuffle(size_df_temp)
# index_df_temp_inc = size_df_temp[:size_inc]
# index_df_temp_not_inc = size_df_temp[size_inc:]
# df_train = pd.concat([df_train, df_temp.iloc[index_df_temp_inc]])
# # df_test = pd.concat([df_test, df_temp.iloc[index_df_temp_not_inc]])
# size_inc = 400
# for age in range(3):
# df_temp = df_l_b_g[df_l_b_g['AgeGroup']==age]
# size_df_temp = np.arange(len(df_temp))
# np.random.seed(42)
# np.random.shuffle(size_df_temp)
# index_df_temp_inc = size_df_temp[:size_inc]
# index_df_temp_not_inc = size_df_temp[size_inc:]
# df_train = pd.concat([df_train, df_temp.iloc[index_df_temp_inc]])
# # df_test = pd.concat([df_test, df_temp.iloc[index_df_temp_not_inc]])
# for age in range(3):
# df_temp = df_l_b_c[df_l_b_c['AgeGroup']==age]
# size_df_temp = np.arange(len(df_temp))
# np.random.seed(42)
# np.random.shuffle(size_df_temp)
# index_df_temp_inc = size_df_temp[:size_inc]
# index_df_temp_not_inc = size_df_temp[size_inc:]
# df_train = pd.concat([df_train, df_temp.iloc[index_df_temp_inc]])
# # df_test = pd.concat([df_test, df_temp.iloc[index_df_temp_not_inc]])
size_inc = 300 # 50
for age in range(3):
df_temp = df_f_t_a[df_f_t_a['AgeGroup']==age]
size_df_temp = np.arange(len(df_temp))
np.random.seed(42)
np.random.shuffle(size_df_temp)
index_df_temp_inc = size_df_temp[:size_inc]
index_df_temp_not_inc = size_df_temp[size_inc:]
if age == 0:
df_train = df_temp.iloc[index_df_temp_inc]
# df_test = df_temp.iloc[index_df_temp_not_inc]
else:
df_train = pd.concat([df_train, df_temp.iloc[index_df_temp_inc]])
# df_test = pd.concat([df_test, df_temp.iloc[index_df_temp_not_inc]])
for age in range(3):
df_temp = df_f_b_g[df_f_b_g['AgeGroup']==age]
size_df_temp = np.arange(len(df_temp))
np.random.seed(42)
np.random.shuffle(size_df_temp)
index_df_temp_inc = size_df_temp[:size_inc]
index_df_temp_not_inc = size_df_temp[size_inc:]
df_train = pd.concat([df_train, df_temp.iloc[index_df_temp_inc]])
# df_test = pd.concat([df_test, df_temp.iloc[index_df_temp_not_inc]])
for age in range(3):
df_temp = df_f_b_c[df_f_b_c['AgeGroup']==age]
size_df_temp = np.arange(len(df_temp))
np.random.seed(42)
np.random.shuffle(size_df_temp)
index_df_temp_inc = size_df_temp[:size_inc]
index_df_temp_not_inc = size_df_temp[size_inc:]
df_train = pd.concat([df_train, df_temp.iloc[index_df_temp_inc]])
# df_test = pd.concat([df_test, df_temp.iloc[index_df_temp_not_inc]])
size_inc = 0
for age in range(3):
df_temp = df_vf_t_a[df_vf_t_a['AgeGroup']==age]
size_df_temp = np.arange(len(df_temp))
np.random.seed(42)
np.random.shuffle(size_df_temp)
index_df_temp_inc = size_df_temp[:size_inc]
index_df_temp_not_inc = size_df_temp[size_inc:]
if age == 0:
# df_train = df_temp.iloc[index_df_temp_inc]
df_test = df_temp.iloc[index_df_temp_not_inc]
else:
# df_train = pd.concat([df_train, df_temp.iloc[index_df_temp_inc]])
df_test = pd.concat([df_test, df_temp.iloc[index_df_temp_not_inc]])
for age in range(3):
df_temp = df_vf_t_g[df_vf_t_g['AgeGroup']==age]
size_df_temp = np.arange(len(df_temp))
np.random.seed(42)
np.random.shuffle(size_df_temp)
index_df_temp_inc = size_df_temp[:size_inc]
index_df_temp_not_inc = size_df_temp[size_inc:]
# df_train = pd.concat([df_train, df_temp.iloc[index_df_temp_inc]])
df_test = pd.concat([df_test, df_temp.iloc[index_df_temp_not_inc]])
size_inc = 50
for age in range(3):
df_temp = df_vf_b_g[df_vf_b_g['AgeGroup']==age]
size_df_temp = np.arange(len(df_temp))
np.random.seed(42)
np.random.shuffle(size_df_temp)
index_df_temp_inc = size_df_temp[:size_inc]
index_df_temp_not_inc = size_df_temp[size_inc:]
df_train = pd.concat([df_train, df_temp.iloc[index_df_temp_inc]])
# df_test = pd.concat([df_test, df_temp.iloc[index_df_temp_not_inc]])
for age in range(3):
df_temp = df_vf_b_c[df_vf_b_c['AgeGroup']==age]
size_df_temp = np.arange(len(df_temp))
np.random.seed(42)
np.random.shuffle(size_df_temp)
index_df_temp_inc = size_df_temp[:size_inc]
index_df_temp_not_inc = size_df_temp[size_inc:]
df_train = pd.concat([df_train, df_temp.iloc[index_df_temp_inc]])
# df_test = pd.concat([df_test, df_temp.iloc[index_df_temp_not_inc]])
X = df_train.iloc[:,6:-1]
y_age = df_train["Age"]
y_age_groups = df_train["AgeGroup"]
y_species = df_train["Species"]
y_status = df_train["Status"]
X = np.asarray(X)
y_age = np.asarray(y_age)
y_age_groups = np.asarray(y_age_groups)
y_species = np.asarray(y_species)
y_status = np.asarray(y_status)
print('shape of X : {}'.format(X.shape))
print('shape of y age : {}'.format(y_age.shape))
print('shape of y age groups : {}'.format(y_age_groups.shape))
print('shape of y species : {}'.format(y_species.shape))
print('shape of y status : {}'.format(y_status.shape))
X_vf = df_test.iloc[:,6:-1]
y_age_vf = df_test["Age"]
y_age_groups_vf = df_test["AgeGroup"]
y_species_vf = df_test["Species"]
y_status_vf = df_test["Status"]
X_vf = np.asarray(X_vf)
y_age_vf = np.asarray(y_age_vf)
y_age_groups_vf = np.asarray(y_age_groups_vf)
y_species_vf = np.asarray(y_species_vf)
y_status_vf = np.asarray(y_status_vf)
print('shape of X vf : {}'.format(X_vf.shape))
print('shape of y age vf : {}'.format(y_age_vf.shape))
print('shape of y age groups vf : {}'.format(y_age_groups_vf.shape))
print('shape of y species vf : {}'.format(y_species_vf.shape))
print('shape of y status vf : {}'.format(y_status_vf.shape))
input_layer_dim = len(Xf[0])
y_age_groups_list_l = [[age] for age in y_age_groups]
y_species_list_l = [[species] for species in y_species]
age_groups_l = MultiLabelBinarizer().fit_transform(np.array(y_age_groups_list_l))
age_group_classes = ["1-4", "5-10", "11-17"]
species_l = MultiLabelBinarizer().fit_transform(np.array(y_species_list_l))
species_classes = list(np.unique(y_species_list_l))
y_age_groups_list_vf = [[age] for age in y_age_groups_vf]
y_species_list_vf = [[species] for species in y_species_vf]
age_groups_vf = MultiLabelBinarizer().fit_transform(np.array(y_age_groups_list_vf))
species_vf = MultiLabelBinarizer().fit_transform(np.array(y_species_list_vf))
outdir = "Results_Paper/"
build_folder(outdir, False)
SelectFreqs = False
## Labels default - all classification
labels_default_vf, labels_default_l, classes_default, outputs_default = [age_groups_vf, species_vf], [age_groups_l, species_l], [age_group_classes, species_classes], ['xAgeGroup', 'xSpecies']
## Declare and train the model
model_size = [{'type':'c', 'filter':16, 'kernel':8, 'stride':1, 'pooling':1},
{'type':'c', 'filter':16, 'kernel':8, 'stride':2, 'pooling':1},
{'type':'c', 'filter':16, 'kernel':3, 'stride':1, 'pooling':1},
{'type':'c', 'filter':16, 'kernel':6, 'stride':2, 'pooling':1},
{'type':'c', 'filter':16, 'kernel':5, 'stride':1, 'pooling':2},
{'type':'d', 'width':500}]
## Name the model
model_name = 'Baseline_CNN'
histories = []
fold = 1
train_model = True
## Name a folder for the outputs to go into
savedir = (outdir+"Trian_TCField_No_Ifakara/")
build_folder(savedir, True)
start_time = time()
save_predicted = []
save_true = []
## Scale train, test
scl = StandardScaler()
features_scl = scl.fit(X=np.vstack((X,X_vf)))
features_l = features_scl.transform(X=X)
features_vf = features_scl.transform(X=X_vf)
## Split into training / testing
# test_splits = train_test_split(features_f, *(labels_default_f), test_size=0.1, shuffle=True, random_state=rand_seed)
## Pack up data
X_train = features_l
X_test = features_vf
y_train = labels_default_l
y_test = labels_default_vf
if not SelectFreqs:
X_train = np.expand_dims(X_train, axis=2)
X_test = np.expand_dims(X_test, axis=2)
model_to_test = {
"model_shape" : [model_size], # defines the hidden layers of the model
"model_name" : [model_name],
"input_layer_dim" : [input_layer_dim], # size of input layer
"model_ver_num" : [0],
"fold" : [fold], # kf.split number on
"labels" : [y_train],
"features" : [X_train],
"classes" : [classes_default],
"outputs" : [outputs_default],
"compile_loss": [{'age': 'categorical_crossentropy'}],
"compile_metrics" :[{'age': 'accuracy'}]
}
## Call function to train all the models from the dictionary
model, history = train_models(model_to_test, savedir, SelectFreqs=SelectFreqs)
histories.append(history)
predicted_labels = list([] for i in range(len(y_train)))
true_labels = list([] for i in range(len(y_train)))
y_predicted = model.predict(X_test)
predicted_labels = [x+[y] for x,y in zip(predicted_labels,y_predicted)]
true_labels = [x+[y] for x,y in zip(true_labels,y_test)]
predicted_labels = [predicted_labels[i][0].tolist() for i in range(len(predicted_labels))]
true_labels = [true_labels[i][0].tolist() for i in range(len(true_labels))]
for pred, tru in zip(predicted_labels, true_labels):
save_predicted.append(pred)
save_true.append(tru)
## Visualize the results
visualize(histories, savedir, model_name, str(fold), classes_default, outputs_default, predicted_labels, true_labels)
# log_data(test_index, 'test_index', fold, savedir)
# Clear the Keras session, otherwise it will keep adding new
# models to the same TensorFlow graph each time we create
# a model with a different set of hyper-parameters.
K.clear_session()
# Delete the Keras model with these hyper-parameters from memory.
del model
# visualize(1, savedir, model_name, "Averaged", classes_default, outputs_default, save_predicted, save_true)
end_time = time()
print('Run time : {} s'.format(end_time-start_time))
print('Run time : {} m'.format((end_time-start_time)/60))
print('Run time : {} h'.format((end_time-start_time)/3600))
```
## Testing Phase
### Process testing data
```
def test_data_extract(RearCnd, Country, Species):
df_RearCnd = df_test[df_test['RearCnd']==RearCnd]
df_Country = df_RearCnd[df_RearCnd['Country']==Country]
df_Species = df_Country[df_Country['Species']==Species]
X_test_extract = df_Species.iloc[:,6:-1]
y_age_test_extract = df_Species["Age"]
y_age_groups_test_extract = df_Species["AgeGroup"]
y_species_test_extract = df_Species["Species"]
y_status_test_extract = df_Species["Status"]
print('shape of X_vf : {}'.format(X_test_extract.shape))
print('shape of y_age_vf : {}'.format(y_age_test_extract.shape))
print('shape of y_age_groups_test_extract : {}'.format(y_age_groups_test_extract.shape))
print('shape of y y_species_vf : {}'.format(y_species_test_extract.shape))
print('shape of y y_status_vf : {}'.format(y_status_test_extract.shape))
X_test_extract = np.asarray(X_test_extract)
y_age_test_extract = np.asarray(y_age_test_extract)
y_age_groups_test_extract = np.asarray(y_age_groups_test_extract)
y_species_test_extract = np.asarray(y_species_test_extract)
y_status_test_extract = np.asarray(y_status_test_extract)
# y_age_groups = np.where((y_age_test_extract<=4), 0, 0)
# y_age_groups = np.where((y_age_test_extract>=5) & (y_age_test_extract<=10), 1, y_age_groups)
# y_age_groups = np.where((y_age_test_extract>=11), 2, y_age_groups)
return X_test_extract, y_age_test_extract, y_age_groups_test_extract, y_species_test_extract, y_status_test_extract
def test_data_format(y_age_groups, y_species):
y_age_groups_list = [[age] for age in y_age_groups]
y_species_list = [[species] for species in y_species]
age_groups = MultiLabelBinarizer().fit([(0,), (1,), (2,)])
age_groups = age_groups.transform(np.array(y_age_groups_list))
age_group_classes = ["1-4", "5-10", "11-17"]
species = MultiLabelBinarizer().fit([set(['AA']), set(['AC']), set(['AG'])])
species = species.transform(np.array(y_species_list))
species_classes = list(np.unique(y_species_list))
# labels_default, classes_default, outputs_default = [age_groups, species], [age_group_classes, species_classes], ['xAgeGroup', 'xSpecies']
labels_default, classes_default, outputs_default = [species], [species_classes], ['xSpecies']
return labels_default, classes_default, outputs_default
## Set up folders
outdir = "output_data_update_19_02/"
build_folder(outdir, False)
loaddir = (outdir+"Train_Field/Trian_Species_Only/")
for i in range(0,10):
savedir = (outdir+"Train_Field/Trian_Species_Only/Testing/model"+str(i+1)+"/")
build_folder(savedir, True)
## Load model
model = load_model((loaddir+"Baseline_CNN_0_"+str(i+1)+"_Model.h5"))
print('Model loaded successfully')
## Set up testing data choice
RearCnd = 'TL'
Country = 'T'
Species = 'AA'
for RearCnd in ['TL', 'TF', 'VF']:
for Country, Species in [['T', 'AA'], ['T', 'AG'], ['B', 'AG'], ['B', 'AC']]:
## Extract data
X_test, y_age_test, y_age_groups_test, y_species_test, y_status_test = test_data_extract(RearCnd, Country, Species)
if len(y_age_test) == 0:
pass
else:
## Format data
labels_default, classes_default, outputs_default = test_data_format(y_age_groups_test, y_species_test)
model_name = 'Testing_'+RearCnd+'_'+Country+'_'+Species
## Scale train, test
scl = StandardScaler()
scl_fit = scl.fit(X=X)
features = scl_fit.transform(X=X)
features_test = scl_fit.transform(X=X_test)
## Split data into test and train
X_test = features_test[:]
y_test = list(map(lambda y:y[:], labels_default))
X_test = np.expand_dims(X_test, axis=2)
## Prediction
predicted_labels = list([] for i in range(len(y_test)))
true_labels = list([] for i in range(len(y_test)))
y_predicted = model.predict(X_test)
predicted_labels = [x+[y] for x,y in zip(predicted_labels,y_predicted)]
true_labels = [x+[y] for x,y in zip(true_labels,y_test)]
predicted_labels = [predicted_labels[i][0].tolist() for i in range(len(predicted_labels))]
true_labels = [true_labels[i][0].tolist() for i in range(len(true_labels))]
species_classes_pred_temp = np.unique(np.argmax(y_predicted[1], axis=-1))
species_classes_pred = []
for spec in species_classes_pred_temp:
if spec == 0:
species_classes_pred.append('AA')
elif spec == 1:
species_classes_pred.append('AC')
elif spec == 2:
species_classes_pred.append('AG')
species_classes_pred = np.array(species_classes_pred)
classes_pred = [classes_default[0], species_classes_pred]
## Visualize the results
visualize(histories, savedir, model_name, str(0), classes_pred, outputs_default, predicted_labels, true_labels)
print('Testing of {} - {} - {} completed'.format(RearCnd, Country, Species))
```
## Sensitivity Plots
```
## Functions for Z-score and sensitivity for input-output
def generate_sensitivity_Z_score(model, layer_name, age, size=1625):
layer_output = model.get_layer(layer_name).output
if layer_name == 'age':
df_1 = df[df['Age']==age]
loss = layer_output[:, age-1]
elif layer_name == 'species':
df_1 = df[df['Species']==age]
if age == 'AA':
loss = layer_output[:, 0]
elif age == 'AC':
loss = layer_output[:, 1]
elif age == 'AG':
loss = layer_output[:, 2]
elif layer_name == 'age_group':
df1 = df_train[df_train['AgeGroup']==age]
loss = layer_output[:, age]
X = df1.iloc[:,6:-1]
X = np.asarray(X)
grads = K.gradients(loss, model.input)[0]
iterate = K.function([model.input], [loss, grads])
gradients = []
for i in range(len(X)):
input_img_data = X[i]
input_img_data = np.expand_dims(input_img_data, axis=0)
input_img_data = np.expand_dims(input_img_data, axis=2)
loss_value, grads_value = iterate([input_img_data])
gradients.append(np.squeeze(np.abs(grads_value)))
sensitivity = 1/len(gradients) * np.sum(gradients, axis=0)
return sensitivity/np.linalg.norm(sensitivity)
def sensitivites_for_age(age):
outdir = "Results/"
loaddir = (outdir+"Trian_Lab_Field/")
sensitivities = []
for count in tqdm(range(10)):
model = load_model((loaddir+"Baseline_CNN_0_"+str(count+1)+"_Model.h5"))
# model.summary()
for layer in model.layers:
layer.trainable = False
sensitivity = generate_sensitivity_Z_score(model, 'age_group', age)
sensitivities.append(sensitivity)
del model
return sensitivities
## Generates outputs of Z-score and sensitivty for input-ouput
outdir = "Results/"
savedir = (outdir+"Trian_Lab_Field/Sensitivity/")
build_folder(savedir, False)
sensitivities_save = []
for age in tqdm(range(0,3)):
sensitivities = sensitivites_for_age(age)
# print(sensitivities)
# print(sensitivities.shape)
sensitivities_save.append(sensitivities)
sensitivities = [sensitivities_save[0][i] + sensitivities_save[1][i] + sensitivities_save[2][i] for i in range(10)]
Z_scores = []
m_signals = []
for sens1 in range(10):
for sens2 in range(10):
s_signal = (sensitivities[sens1] + sensitivities[sens2]) / np.sqrt(2)
mean_b = np.mean(s_signal)
sigma_b = np.std(s_signal)
for sig in s_signal:
Z_b = (sig-mean_b)/sigma_b
Z_scores.append(Z_b)
m_signals.append(sig)
fig = plt.figure()
plt.scatter(m_signals, Z_scores)
poly_index = 3
plt.plot(np.unique(m_signals), np.poly1d(np.polyfit(m_signals, Z_scores, poly_index))(np.unique(m_signals)), color='k', linewidth=3)
index_95 = (np.where(np.logical_and(np.poly1d(np.polyfit(m_signals, Z_scores, poly_index))(np.unique(m_signals)) < 1.7, np.poly1d(np.polyfit(m_signals, Z_scores, poly_index))(np.unique(m_signals)) > 1.6)))
index_95 = index_95[0][int(len(index_95)/2)]
y_value = (np.poly1d(np.polyfit(m_signals, Z_scores, poly_index))(np.unique(m_signals))[index_95])
x_value = (np.unique(m_signals)[index_95])
plt.plot([0, x_value], [y_value, y_value], 'k--')
plt.plot([x_value, x_value], [-4, y_value], 'k--')
# plt.xlim([0, 0.2])
plt.ylim([-4, 6])
plt.xlabel('Signal value')
plt.ylabel('Z-score')
plt.title(('Z-score Calculation - Age '+str(age)))
plt.tight_layout()
plt.savefig((savedir+'Z_Score_Grouped_Age_'+str(age)+'.png'))
## Start of individual age Sensitivity plots
# print(sensitivities)
font = {'family' : 'normal',
'weight' : 'normal',
'size' : 14}
matplotlib.rc('font', **font)
fig = plt.figure(figsize=(8,4))
ax = fig.add_subplot(1,1,1)
sens_vals = np.sum(np.array(sensitivities), axis=0)/10
print(sens_vals.shape)
l1 = plt.plot(np.arange(3800, 550, -2), np.squeeze(sensitivities[0]), c='b')
# l1 = plt.plot(np.arange(3800, 550, -2), np.squeeze(sens_vals), c='b')
l3 = plt.plot([3800, 550], [x_value, x_value], 'k--')
ax.set_xlim(3800, 550)
ax.set_ylim(0, 0.3)
ax.set_xlabel('Wavenumber $cm^{-1}$', fontsize=18)
ax.set_ylabel('Sensitivity', fontsize=18)
# ax.set_title(('Sensitivity map'))
for mol in [3400, 3276, 2923, 2859, 1901, 1746, 1636, 1539, 1457, 1307, 1154, 1076, 1027, 880]:
l2 = plt.plot([mol, mol], [0, 0.3], 'k', linewidth=1)
ax2 = ax.twiny()
new_tick_loc = [3400, 3276, 2923, 2859, 1901, 1746, 1636, 1539, 1457, 1307, 1154, 1076, 1027, 880]
ax2.set_xlim(ax.get_xlim())
ax2.set_xticks(new_tick_loc)
ax2.set_xticklabels(['O-H', 'N-H', 'C-H_2', 'C-H_2', '', 'C=O', 'C=O', 'O=C-N', 'C-CH_3', 'C-N',
'C-O-C', 'C-O', 'C-O', '', 'C-C'])
plt.setp(ax2.get_xticklabels(), rotation=90)
# custom_points = [Line2D([3800], [0], color = 'b', label='Sensitivity'),
# Line2D([3800], [0], color = 'k', lw=1, label='Previously used wavenumbers'),
# Line2D([3800], [0], color = 'k', linestyle = '--', label='95% confidence interval')]
# ax.legend(custom_points, ['Sensitivity', 'Molecular Bands', '95% confidence interval'], loc='center left', bbox_to_anchor=(1,0.5))
plt.tight_layout()
plt.savefig((savedir+'Sensitivity_Map_Grouped_Age.png'))
## Start of individual age Sensitivity plots
# print(sensitivities)
font = {'family' : 'normal',
'weight' : 'normal',
'size' : 14}
matplotlib.rc('font', **font)
fig = plt.figure(figsize=(8,4))
ax = fig.add_subplot(1,1,1)
sens_vals = np.sum(np.array(sensitivities), axis=0)/10
print(sens_vals.shape)
l1 = plt.plot(np.arange(3800, 550, -2), np.squeeze(sensitivities[0]), c='b')
# l1 = plt.plot(np.arange(3800, 550, -2), np.squeeze(sens_vals), c='b')
l3 = plt.plot([3800, 550], [x_value, x_value], 'k--')
ax.set_xlim(3800, 550)
ax.set_ylim(0, 0.3)
ax.set_xlabel('Wavenumber $cm^{-1}$', fontsize=18)
ax.set_ylabel('Sensitivity', fontsize=18)
# ax.set_title(('Sensitivity map'))
for mol in [3400, 3276, 2923, 2859, 1901, 1746, 1636, 1539, 1457, 1307, 1154, 1076, 1027, 880]:
l2 = plt.plot([mol, mol], [0, 0.3], 'k', linewidth=1)
ax2 = ax.twiny()
new_tick_loc = [3400, 3276, 2923, 2859, 1901, 1746, 1636, 1539, 1457, 1307, 1154, 1076, 1027, 880]
ax2.set_xlim(ax.get_xlim())
ax2.set_xticks(new_tick_loc)
ax2.set_xticklabels(['O-H', 'N-H', 'C-H_2', 'C-H_2', '', 'C=O', 'C=O', 'O=C-N', 'C-CH_3', 'C-N',
'C-O-C', 'C-O', 'C-O', '', 'C-C'])
plt.setp(ax2.get_xticklabels(), rotation=90)
# custom_points = [Line2D([3800], [0], color = 'b', label='Sensitivity'),
# Line2D([3800], [0], color = 'k', lw=1, label='Previously used wavenumbers'),
# Line2D([3800], [0], color = 'k', linestyle = '--', label='95% confidence interval')]
# ax.legend(custom_points, ['Sensitivity', 'Molecular Bands', '95% confidence interval'], loc='center left', bbox_to_anchor=(1,0.5))
plt.tight_layout()
plt.savefig((savedir+'Sensitivity_Map_Grouped_Age.png'))
```
| github_jupyter |
<!-- COLOCAR O TREINAMENTO NUMA FUNCAO E FAZER UM GRID SEARCH -->
```
# !pip3.7 install ray==1.3.0
# ray.shutdown()
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import gym
import ray
from ray.rllib.agents.ppo import PPOTrainer, DEFAULT_CONFIG
from ray.tune.logger import pretty_print
# devo diminuir o tempo usado, no caso, o x
ray.init()
import random
import ta
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.preprocessing import MinMaxScaler
import tensortrade.env.default as default
from tensortrade.feed.core import Stream, DataFeed, NameSpace
from tensortrade.oms.exchanges import Exchange
from tensortrade.oms.services.execution.simulated import execute_order
from tensortrade.oms.instruments import USD, BTC, ETH
from tensortrade.oms.wallets import Wallet, Portfolio
from tensortrade.agents import A2CAgent
import tensortrade.stochastic as sp
from tensortrade.oms.instruments import Instrument
from tensortrade.env.default.actions import SimpleOrders, ManagedRiskOrders
from collections import OrderedDict
from tensortrade.oms.orders.criteria import Stop, StopDirection
from tensortrade.env.default.actions import ManagedRiskOrders
from tensortrade.env.default.rewards import RiskAdjustedReturns, TensorTradeRewardScheme
from scipy.signal import savgol_filter
from tensortrade.env.default.actions import TensorTradeActionScheme
import numpy as np
import pandas as pd
import os
dfs = []
for dirname, _, filenames in os.walk('archive/'):
for filename in filenames:
print(os.path.join(dirname, filename))
df = pd.read_json(os.path.join(dirname, filename))
if not df.empty and len(df) >= 51197:
df.columns = ["unix", "open", "high", "low", "close", "volume"]
df.drop(columns=["unix"], inplace=True)
dfs.append(df)
# mudei aqui para tail
else:
print("empty or small")
from tensortrade.env.default.rewards import PBR
from gym.spaces import Space, Discrete
from tensortrade.oms.orders import (
Broker,
Order,
OrderListener,
OrderSpec,
proportion_order,
risk_managed_order,
TradeSide,
TradeType
)
from tensortrade.oms.orders.criteria import StopDirection, Stop
class BSH(TensorTradeActionScheme):
"""A simple discrete action scheme where the only options are to buy, sell,
or hold.
Parameters
----------
cash : `Wallet`
The wallet to hold funds in the base intrument.
asset : `Wallet`
The wallet to hold funds in the quote instrument.
"""
registered_name = "bsh"
def __init__(self, cash: 'Wallet', asset: 'Wallet'):
super().__init__()
self.cash = cash
self.asset = asset
self.listeners = []
self.action = 0
stopLoss = Stop(StopDirection('down'), 0.2)
# sem criteria e com self.default stop, + durations = nada mudou
# sem nada = parou no 600 mas grafico foi bem parecido
# incluindo criteria - bem diferente, apesar de negativo max 1200
# incluindo criteria sem take - conclui quase no 1000, max 1400
# incluindo criteria sem take - 16 neuronios - fracasso, max 1000, cabou em 500, tudo antes era 64,
# incluindo criteria sem take - 128 neuronios - 1000, max = 2500
# incluindo criteria sem take - 256 neuronios - 2000, max = 2600
# incluindo criteria com take 0.05 - 256 neuronios - take 0.05 - 800-max 1300
# incluindo criteria com 3 take 0.05 e 0.01 e 01 - 256 neuronios - bosta
# incluindo criteria com 1 take 0.05 e 0.01 e 01 - 256 neuronios - bosta
# incluindo criteria sem take stoploss 0.3 - 256 neuronios ruim mas parou
# incluindo criteria sem take stoploss 0.3 - 256 neuronios 25k primeiros, ruim
# take = Stop(StopDirection('up'), 0.05)
criteria = [stopLoss]
criteria = self.default('criteria', criteria)
self.criteria = criteria if isinstance(criteria, list) else [criteria]
self.historic_awnser = []
# posso mudar pra market o tipo de ordem, ai para com essa porra
# acho q o correto eh usar uma criteria pra stoploss e take
# tensortrade.org/en/latest/search.html?q=criteria&check_keywords=yes&area=default
# https://github.com/tensortrade-org/tensortrade/blob/4a1ea95cf92a38d704056aac33b6b1d1e5416d2e/tensortrade/env/default/actions.py#L175-L211
# stop = [0.2]
# take = [0.05]
# self.stop = self.default('stop', stop)
# self.take = self.default('take', take)
#com e sem deu msm bosta
# durations = 200
# durations = self.default('durations', durations)
# self.durations = durations if isinstance(durations, list) else [durations]
# https://github.com/tensortrade-org/tensortrade/blob/4a1ea95cf92a38d704056aac33b6b1d1e5416d2e/tensortrade/oms/orders/order.py#L332
@property
def action_space(self):
return Discrete(2)
def attach(self, listener):
self.listeners += [listener]
return self
def get_orders(self, action: int, portfolio: 'Portfolio') -> 'Order':
order = None
if abs(action - self.action) > 0:
src = self.cash if self.action == 0 else self.asset
tgt = self.asset if self.action == 0 else self.cash
if src.balance == 0: # We need to check, regardless of the proposed order, if we have balance in 'src'
return [] # Otherwise just return an empty order list
order = proportion_order(portfolio, src, tgt, 0.1)
self.action = action
for listener in self.listeners:
listener.on_action(action)
self.historic_awnser.append(action)
return [order]
def reset(self):
super().reset()
self.action = 0
class SimpleProfit(TensorTradeRewardScheme):
"""A simple reward scheme that rewards the agent for incremental increases
in net worth.
Parameters
----------
window_size : int
The size of the look back window for computing the reward.
Attributes
----------
window_size : int
The size of the look back window for computing the reward.
"""
def __init__(self, window_size: int = 1):
self._window_size = self.default('window_size', window_size)
def get_reward(self, portfolio: 'Portfolio') -> float:
"""Rewards the agent for incremental increases in net worth over a
sliding window.
Parameters
----------
portfolio : `Portfolio`
The portfolio being used by the environment.
Returns
-------
float
The cumulative percentage change in net worth over the previous
`window_size` time steps.
"""
net_worths = [nw['net_worth'] for nw in portfolio.performance.values()]
returns = [(b - a) / a for a, b in zip(net_worths[::1], net_worths[1::1])]
returns = np.array([x + 1 for x in returns[-self._window_size:]]).cumprod() - 1
return 0 if len(returns) < 1 else returns[-1]
numCoins = 1
def fetchTaFeatures(data):
data = ta.add_all_ta_features(data, 'open', 'high', 'low', 'close', 'volume', fillna=True)
data.columns = [name.lower() for name in data.columns]
return data
def createEnv(config):
coins = ["coin{}".format(x) for x in range(numCoins)]
bitfinex_streams = []
with NameSpace("bitfinex"):
for coin in coins:
coinColumns = filter(lambda name: name.startswith(coin), config["data"].columns)
bitfinex_streams += [
Stream.source(list(config["data"][c]), dtype="float").rename(c) for c in coinColumns
]
feed = DataFeed(bitfinex_streams)
streams = []
for coin in coins:
streams.append(Stream.source(list(data[coin+":"+"close"]), dtype="float").rename("USD-"+coin))
streams = tuple(streams)
bitstamp = Exchange("bitfinex", service=execute_order)(
Stream.source(list(config["data"]["coin0:close"]), dtype="float").rename("USD-TTC0"),
# Stream.source(list(config["data"]["coin1:close"]), dtype="float").rename("USD-TTC1"),
# Stream.source(list(config["data"]["coin2:close"]), dtype="float").rename("USD-TTC2"),
# Stream.source(list(config["data"]["coin3:close"]), dtype="float").rename("USD-TTC3"),
# Stream.source(list(data["coin4:close"]), dtype="float").rename("USD-TTC4"),
# Stream.source(list(data["coin5:close"]), dtype="float").rename("USD-TTC5"),
# Stream.source(list(data["coin6:close"]), dtype="float").rename("USD-TTC6"),
# Stream.source(list(data["coin7:close"]), dtype="float").rename("USD-TTC7"),
# Stream.source(list(data["coin8:close"]), dtype="float").rename("USD-TTC8"),
# Stream.source(list(data["coin9:close"]), dtype="float").rename("USD-TTC9"),
# Stream.source(list(data["coin10:close"]), dtype="float").rename("USD-TTC10"),
# Stream.source(list(data["coin11:close"]), dtype="float").rename("USD-TTC11"),
# Stream.source(list(data["coin12:close"]), dtype="float").rename("USD-TTC12"),
# Stream.source(list(data["coin13:close"]), dtype="float").rename("USD-TTC13"),
# Stream.source(list(data["coin14:close"]), dtype="float").rename("USD-TTC14"),
# Stream.source(list(data["coin15:close"]), dtype="float").rename("USD-TTC15"),
# Stream.source(list(data["coin16:close"]), dtype="float").rename("USD-TTC16"),
# Stream.source(list(data["coin17:close"]), dtype="float").rename("USD-TTC17"),
# Stream.source(list(data["coin18:close"]), dtype="float").rename("USD-TTC18"),
)
instruments = []
assets = []
for i, coin in enumerate(coins):
instrument = Instrument(f"TTC{i}", 8, f"TensorTrade Coin{i}")
instruments.append(instrument)
assets.append(Wallet(bitstamp, 0 * instrument))
cash = Wallet(bitstamp, 1000 * USD)
portfolio = Portfolio(USD, [cash] + assets)
# https://tensortradex.readthedocs.io/en/latest/api/tensortrade.actions.managed_risk_orders.html
reward = RiskAdjustedReturns(return_algorithm = "sortino", window_size=1)
# reward = SimpleProfit()
# take=[0.05, 0.1, 0.04, 0.01, 0.02]
# action_scheme = ManagedRiskOrders(stop=[0.2], trade_sizes=[4])
# criar minha classe aqui, nao vale a pena usar essa bsh sem modificar
action_scheme = BSH(cash=cash, asset=assets[0])
env = default.create(
feed=feed,
portfolio=portfolio,
action_scheme=action_scheme,
reward_scheme=reward,
window_size=config["window_size"]
)
return env
from scipy.signal import savgol_filter
coins = ["coin{}".format(x) for x in range(numCoins)]
dfFinal = []
for i, df in enumerate(dfs[:numCoins]):
coin = coins[i]
for column in ["close", "open", "high", "low", "volume"]:
df[f"diff_{column}"] = df[f"{column}"].apply(np.log).diff().dropna()
df[f"soft_{column}"] = savgol_filter(df[column], 35, 2)
# Usar feature selection?
ta.add_all_ta_features(
df,
**{k: k for k in ['open', 'high', 'low', 'close', 'volume']})
# testar ta normal e ta no soft
# normal piorou
dfFinal.append(df.add_prefix(f"{coin}:"))
data = pd.concat(dfFinal, axis=1)
data = data.reset_index(drop=True)
data.replace([np.inf, -np.inf], np.nan, inplace=True)
# ray.shutdown()
# from sklearn.preprocessing import MinMaxScaler
# scaler = MinMaxScaler(feature_range=(0.1, 1.1))
# norm_data = pd.DataFrame(scaler.fit_transform(data), columns=data.columns)
# import joblib
# norm_data = pd.read_csv("fake_data_norm.csv")
from ray.rllib.agents.ppo import PPOTrainer, DEFAULT_CONFIG
config = DEFAULT_CONFIG.copy()
config['model']["use_lstm"] = True
# config['num_cpus_per_worker'] = 0 # This avoids running out of resources in the notebook environment when this cell is re-executed
# config['train_batch_size'] = 30
# config["sgd_minibatch_size"] = 10
# config['num_workers'] = 4
# config['num_sgd_iter'] = 30
# config['sgd_minibatch_size'] = 30
config['model']['lstm_use_prev_reward'] = True
config['model']['lstm_use_prev_action'] = True
cellsize = 256
config['model']['lstm_cell_size'] = cellsize
# '': 16,
# window=15
# cell = 16
# prev action = true
# reward = true
config
norm_data = data
# ray.shutdown()
# ray.init()
from ray.tune.registry import register_env
# config["env_config"]["data"]=norm_data
register_env("TradingEnv", createEnv)
config["env_config"]["data"]=norm_data
config["env_config"]["window_size"]=15
config['vf_clip_param'] = 6000000
agent = PPOTrainer(config, 'TradingEnv')
# # Tunar
# config["env_config"]["window_size"]=10
# config["model"]["max_seq_len"] = 20
# config['model']['fcnet_hiddens'] = [128, 32]
# config['train_batch_size'] = 30
for i in range(150):
result = agent.train()
# print(pretty_print(result))
checkpoint_path = agent.save()
print(checkpoint_path)
trained_config = config.copy()
test_agent = PPOTrainer(trained_config, 'TradingEnv')
test_agent.restore(checkpoint_path)
config_env = {}
config_env["data"] = norm_data
config_env["window_size"] = 15
env = createEnv(config_env)
obs = env.reset()
done = False
cumulative_reward = 0
policy = test_agent.get_policy()
state=[np.zeros(cellsize, np.float32),
np.zeros(cellsize, np.float32)]
actions=np.zeros(2*16, np.float32).reshape(2,16)
rewards=np.zeros(16, np.float32)
action = np.zeros(env.action_space.shape, dtype=np.float32)
reward = 0
actionsA = []
episode_reward =0
i = 0
while not done:
# action, state, logits = policy.compute_single_action(obs, state)
# state, reward, done, _ = env.step(action)
# cumulative_reward += reward
action, state, logits = test_agent.compute_action(obs, state,prev_action=action,prev_reward=reward)
obs, reward, done, info = env.step(action)
actionsA.append(action)
actions[:,:-1] = actions[:,1:]
actions[:, -1] = action
rewards[:-1] = rewards[1:]
rewards[-1] = reward
episode_reward += reward
print(episode_reward)
print(actionsA)
orDict = OrderedDict()
for k in env.action_scheme.portfolio.performance.keys():
orDict[k] = env.action_scheme.portfolio.performance[k]["net_worth"]
pd.DataFrame().from_dict(orDict, orient='index').plot()
plt.show()
# ray.shutdown()
fig, axs = plt.subplots(1, 2, figsize=(15, 5))
fig.suptitle("Performance")
# axs[0].plot(np.arange(len(norm_data["close"])), norm_data["close"], label="price")
# axs[0].set_title("Trading Chart")
performance_df = pd.DataFrame().from_dict(env.action_scheme.portfolio.performance, orient='index')
performance_df.plot(ax=axs[1])
axs[1].set_title("Net Worth")
plt.show()
env.observer.history.rows
```
Tune
PS: Pesado em disco e memoria
```
# ray.shutdown()
# from ray import tune
# import ray
# # ray.init()
# from ray.tune.registry import register_env
# register_env("TradingEnv", createEnv)
# from ray.tune.registry import register_env
# # register_env("TradingEnv", createEnv)
# config = DEFAULT_CONFIG.copy()
# config["env_config"]["data"]=norm_data
# config["env_config"]["window_size"]=tune.grid_search([10, 15, 30])
# config["env"] = "TradingEnv"
# # agent = PPOTrainer(config, 'TradingEnv')
# config['model']["use_lstm"] = True
# config['num_workers'] = 1
# config["log_level"] = "DEBUG"
# config["model"]["lstm_cell_size"] = tune.grid_search([16, 32, 62, 128, 256])
# # config["model"]["max_seq_len"] = tune.grid_search([5, 10, 16, 32, 62])
# config["model"]["lstm_use_prev_action"] = tune.grid_search([True, False])
# config["model"]["lstm_use_prev_reward"] = tune.grid_search([True, False])
# import os
# os.environ["TUNE_MAX_PENDING_TRIALS_PG"] = "1"
# # config['model']["max_seq_len"] = 32
# # config['model']["lstm_cell_size"]= 256
# from ray.tune.registry import register_env
# register_env("TradingEnv", createEnv)
# analysis = tune.run(
# "PPO",
# # num_samples=1,
# stop={
# # "episode_reward_mean": 500,
# "training_iteration": 30
# },
# metric="episode_reward_mean",
# mode="max",
# config=config,
# checkpoint_at_end=True,
# # resume=True ,
# )
# config
# import ray.rllib.agents.ppo as ppo
# # # Get checkpoint
# checkpoints = analysis.get_trial_checkpoints_paths(
# trial=analysis.get_best_trial("episode_reward_mean", mode="max"),
# metric="episode_reward_mean"
# )
# checkpoint_path = checkpoints[0][0]
# # Restore agent
# agent = ppo.PPOTrainer(
# env="TradingEnv",
# config=config
# )
# agent.restore(checkpoint_path)
# env = createEnv({
# "window_size": 10,
# "data": norm_data
# })
# config_env = {}
# config_env["data"] = norm_data
# config_env["window_size"] = 10
# env = createEnv(config_env)
# obs = env.reset()
# done = False
# cumulative_reward = 0
# # https://github.com/ray-project/ray/issues/9220
# # ver cell_size
# policy = agent.get_policy()
# cellsize=256
# state=[np.zeros(cellsize, np.float32),
# np.zeros(cellsize, np.float32)]
# # state = policy.get_initial_state()
# # , prev_action=0, prev_reward=0
# actions=np.zeros(2*16, np.float32).reshape(2,16)
# rewards=np.zeros(16, np.float32)
# episode_reward =0
# while not done:
# action, state, logits = agent.compute_action(obs, state)
# obs, reward, done, info = env.step(action)
# actions[:,:-1] = actions[:,1:]
# actions[:, -1] = action
# rewards[:-1] = rewards[1:]
# rewards[-1] = reward
# episode_reward += reward
# fig, axs = plt.subplots(1, 2, figsize=(15, 10))
# fig.suptitle("Performance")
# axs[0].plot(np.arange(len(data["coin0:close"])), data["coin0:close"], label="price")
# axs[0].set_title("Trading Chart")
# performance_df = pd.DataFrame().from_dict(env.action_scheme.portfolio.performance, orient='index')
# performance_df.plot(ax=axs[1])
# axs[1].set_title("Net Worth")
# plt.show()
# orDict = OrderedDict()
# for k in env.action_scheme.portfolio.performance.keys():
# orDict[k] = env.action_scheme.portfolio.performance[k]["net_worth"]
# pd.DataFrame().from_dict(orDict, orient='index').plot()
# plt.show()
# config
```
| github_jupyter |
```
%pylab inline
import pandas as pd
import h5py
from tqdm import tqdm
store = pd.HDFStore('store.h5')
df= pd.read_table('/staging/as/skchoudh/re-ribo-analysis/hg38/SRP010679/ribocop_results_Feb2019_longest/SRX118287_translating_ORFs.tsv')
df.head()
h5py_obj.close()
row
import h5py
h5py_obj = h5py.File ('/staging/as/skchoudh/re-ribo-analysis/hg38/SRP010679/ribocop_results_Feb2019_longest/SRX118287_translating_ORFs.h5', 'w')
total_rows = df.shape[0]
with tqdm(total=total_rows) as pbar:
for idx, row in df.iterrows():
orf_id = row['ORF_ID']
profile = eval(row['profile'])
read_count = row['read_count']
phase_score = row['phase_score']
valid_codons = row['valid_codons']
subgroup = h5py_obj.create_group(orf_id)
dset = subgroup.create_dataset(
"profile",
(len(profile),),
dtype=np.dtype("int64"),
compression="gzip",
compression_opts=9,
)
dset[...] = profile
dset = subgroup.create_dataset(
"read_count",
(1,),
dtype=np.dtype("int64"),
compression="gzip",
compression_opts=9,
)
dset[...] = [read_count]
dset = subgroup.create_dataset(
"valid_codons",
(1,),
dtype=np.dtype("int64"),
compression="gzip",
compression_opts=9,
)
dset[...] = [valid_codons]
dset = subgroup.create_dataset(
"phase_score",
(1,),
dtype=np.dtype("int64"),
compression="gzip",
compression_opts=9,
)
dset[...] = [phase_score]
pbar.update()
class HDFParser(object):
def __init__(self, filepath):
self.filepath = filepath
self.h5py_obj = h5py.File(filepath, "r")
chrom_names = self.h5py_obj["chrom_names"]
chrom_sizes = self.h5py_obj["chrom_sizes"]
chrom_lengths = dict(
[
(chrom.replace("_neg", "").replace("_pos", ""), int(size))
for chrom, size in zip(chrom_names, chrom_sizes)
]
)
chrom_lengths = OrderedDict(sorted(chrom_lengths.items()))
self.chromosome_lengths = chrom_lengths
def close(self):
self.h5py_obj.close()
def __enter__(self):
return self
def __exit__(self, exc_type, exc_value, traceback):
self.h5py_obj.close()
def get_coverage(
self,
region=(None, None, None, None),
fragment_length=None,
orientation="5prime",
outprefix=None,
):
"""Get coverage for a selected region.
Since the region is user provided, we assume the user is aware if it is strand
specfic or not
Paramters
---------
region: tuple
(chromosome, start[0-based], end[1-based], strand(+/-))
fragment_length: list or string
list of fragment lengths to use
Example: [20, 21, 22] or 'all'
"""
chrom, start, stop, strand = region
start = int(start)
stop = int(stop)
assert chrom is not None, "chromosome not set"
assert start is not None, "start position is not set"
assert stop is not None, "end position not set"
assert strand is not None, "strand is not set"
assert start < stop, "start should be < stop"
assert fragment_length is not None, "fragment_length not set"
if isinstance(fragment_length, int):
fragment_length = [fragment_length]
if fragment_length == "all":
h5py_fragments = self.h5py_obj["read_lengths"]
fragment_length = h5py_fragments
fragment_length = list([str(x) for x in fragment_length])
coverages_normalized = pd.DataFrame()
coverages = pd.DataFrame()
position_counter = Counter()
for l in fragment_length:
if strand == "-":
chrom_name = "{}__neg".format(chrom)
elif strand == "+":
chrom_name = "{}__pos".format(chrom)
else:
raise ValueError("strand ill-defined: {}".format(strand))
root_obj = self.h5py_obj["fragments"][l][orientation]
if chrom_name not in list(root_obj.keys()):
# This chromosome does not exist in the
# key value store
# So should returns zeros all way
coverage = pd.Series(
[0] * (stop - start), index=list(range(start, stop))
)
else:
chrom_obj = root_obj[chrom_name]
counts_series = pd.Series(
list(chrom_obj["counts"]), index=list(chrom_obj["positions"])
)
# print(counts_series)
coverage = counts_series.get(list(range(start, stop)))
if coverage is None:
coverage = pd.Series(
[0] * (stop - start), index=list(range(start, stop))
)
coverage = coverage.fillna(0)
# Mean is taken by summing the rows
coverage_mean = coverage.mean(axis=0, skipna=True)
# to normalize
# we divide the sum by vector obtained in previous
# such that each column gets divided
coverage_normalized = coverage.divide(coverage_mean).fillna(0)
coverages = coverages.join(
pd.DataFrame(coverage, columns=[str(l)]), how="outer"
)
coverages_normalized = coverages_normalized.join(
pd.DataFrame(coverage_normalized, columns=[str(l)]), how="outer"
)
position_counter += Counter(coverage.index.tolist())
position_counter = pd.Series(Counter(position_counter)).sort_index()
coverage_sum = coverages.sum(axis=1)
coverage_normalized_sum = coverages_normalized.sum(axis=1)
coverage_sum_div = coverage_sum.divide(position_counter, axis="index")
coverage_normalized_sum_div = coverage_normalized_sum.divide(
position_counter, axis="index"
)
if outprefix:
mkdir_p(os.path.dirname(outprefix))
coverage_sum_div.to_csv("{}_raw.tsv", index=False, header=True, sep="\t")
coverage_normalized_sum_div.to_csv(
"{}_normalized.tsv", index=False, header=True, sep="\t"
)
coverages.index.name = "start"
coverages = coverages.reset_index()
coverages_normalized.index.name = "start"
coverages_normalized = coverages_normalized.reset_index()
return (
coverages,
coverages_normalized,
coverage_sum_div,
coverage_normalized_sum_div,
)
def get_read_length_dist(self):
read_lengths = list([int(x) for x in self.h5py_obj["read_lengths"]])
read_counts = list(self.h5py_obj["read_lengths_counts"])
return pd.Series(read_counts, index=read_lengths).sort_index()
def get_query_alignment_length_dist(self):
read_lengths = list([int(x) for x in self.h5py_obj["query_alignment_lengths"]])
read_counts = list(self.h5py_obj["query_alignment_lengths_counts"])
return pd.Series(read_counts, index=read_lengths).sort_index()
```
| github_jupyter |
<a href="https://colab.research.google.com/github/anadiedrichs/time-series-analysis/blob/master/proyecto_forma_de_onda_2019.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Laboratorio #1 Proyecto forma de onda - Unidad análisis de series temporales - Teleinformática IoT - 2019
**Docente: Ing. Ana Laura Diedrichs**
Consultas: lunes 19 hs en sala consulta de sistemas.
Contacto por otros horarios de consulta o dudas:
* Email: ana.diedrichs@frm.utn.edu.ar
* Telegram @anadiedrichs
## Objetivos del laboratorio:
* Analizar una serie temporal para un problema específico
* Fomentar el trabajo en clase, discusión y aportes de soluciones entre los participantes
* Ser una entrada en calor o introducción del uso del entorno colab.research.google.com y librerías Python como pandas
## Pre-requisitos o pre-condiciones
* Tener una cuenta google (gmail)
* Tener instalado el navegador google chrome
* Contar con conectividad a Internet
## Entrega y uso del laboratorio
**USO**
* Antes que cualquier cosa, **cree una copia de este notebook: Click en *File*, luego *Save a Copy in Drive***
* Renombre el archivo con el siguiente formato: APELLIDO_NOMBRE_LEGAJO_titulonotebook.ipynb
Ejemplo: DIEDRICHS_ANA_99999_proyecto_forma_de_onda_2019.ipynb
* Use el notebook, complete las actividades y consignas que se elija.
* Este laboratorio es una actividad individual.
* Se fomenta el intercambio de opiniones en clase y exposición.
**ENTREGA**
* Una vez finalizado el laboratorio, complete [el formulario de entrega](https://forms.gle/zUnbfN79YEEeJFz7A) indicando
* Apellido
* Nombre
* Nro Legajo
* Carrera
* Actividad (cuál laboratorio o actividad)
* link de su notebook. El mismo se obtiene si realiza click en *Share* (esquina superior derecha) y luego en *Get shareable link*
No se aceptarán otras formas de entrega distintas a la mencionada.
Fecha límite de entrega: 17 de octubre de 2019.
## Intro
Durante el año 2018 era la primera vez que se dictaba esta unidad en la materia. En la primer clase realicé la pregunta sobre si les interesaba algunos datos en particular para analizar o algún dataset. Un alumno nos acercó su inquietud y compartió el dataset que es el que vamos a usar en este laboratorio.
El dataset contiene mediciones de una toma de corriente trifásica.
Vea más sobre corrienta alterna en https://es.wikipedia.org/wiki/Corriente_alterna
Notará que las señales son senoidales.
## Carga del dataset
### por pasos: chusmeando con la CLI primero, luego importar con pandas
```
!wget https://github.com/anadiedrichs/time-series-analysis/raw/master/datasets/forma_de_onda/forma_de_onda.csv
!ls -lh
```
Notamos que forma_de_onda.csv pesa unos 47 Megabytes.
Chusmeo las primeras líneas
```
!head forma_de_onda.csv
```
Observe como están separados los campos de información (por coma), el formato de la fecha y la hora.
Importamos usando la funcion read_csv, puede demorar un poco en ejecutarse.
```
import pandas as pd
from pandas import Series
data = pd.read_csv('forma_de_onda.csv',header=0, parse_dates=[0],squeeze=True) #index_col=0,
```
index_col es 0 : toma la primer columna como indice por defecto
header es 0, la primer fila es usada como cabecera
parse_dates es True, intenta convertir los tipos de datos a tipo DateTime
La option squeeze=True regresa tipo Series
Mostramos las primeras 10 líneas del dataset
```
data.head(10)
type(data) #el tipo de variable que es data, es un DataFrame
```
## Accediendo a pedacitos del dataset
Veo desde la fila 1 a la 20, las columnas 2 a la 7
```
data.iloc[1:20,2:7]
```
Grafico un rango del dataset
```
data.iloc[1:100,2:7].plot()
```
## Propiedades del dataset
Cuántas filas y columnas tiene
```
data.shape
```
Son las mismas columnas que miramos ejecutando *head*
```
data.head(10)
```
¿Lo siguiente muestra número de filas o columnas?
```
data.shape[0]
```
¿Lo siguiente muestra número de filas o columnas?
```
data.shape[1]
```
Podemos observar de cada DataFrame algunas características estadísticas usando *describe()*
```
data.describe()
```
## Valores perdidos
Chusmeamos el dataset nuevamente
```
data.iloc[50:60,0:5]
data.iloc[1:100,2:7].plot()
```
### [Actividad]
Habrá notado que hay valores pedidos en la serie.
Determine si reconstruye la señal compleando los valores perdidos e indicados como NaN o los ignora. Justifique
SU RESPUESTA AQUI
```
```
## [actividad] Intervalos de muestreo
¿Cada cuánto tenemos una medición? ¿Qué nos puede decir sobre el intervalo de muestreo?
**SU RESPUESTA AQUI**
## [actividad] Análisis de la señal
¿Son todas las ondas "perfectamente" senoidales?
¿Por qué cree que alguna no?
**SU RESPUESTA AQUI**
### [actividad] Calcule y grafique la FFT de la señal
ESCRIBA SUS COMENTARIOS AQUI
```
```
¿qué concluye al ver este gráfico?
SU RESPUESTA AQUI
## [actividad] Calcule la integral de la señal
¿Qué significado tiene calcular la integral de un tramo de la señal? Comente con sus colegas, discuta o busque información.
| github_jupyter |
HMA1 Class
==========
In this guide we will go through a series of steps that will let you
discover functionalities of the `HMA1` class.
What is HMA1?
-------------
The `sdv.relational.HMA1` class implements what is called a
*Hierarchical Modeling Algorithm* which is an algorithm that allows to
recursively walk through a relational dataset and apply tabular models
across all the tables in a way that lets the models learn how all the
fields from all the tables are related.
Let's now discover how to use the `HMA1` class.
Quick Usage
-----------
We will start by loading and exploring one of our demo datasets.
```
from sdv.demo import sample_relational_demo
metadata, tables = sample_relational_demo(size=30)
```
This will return two objects:
1. A `Metadata` object with all the information that **SDV** needs to
know about the dataset.
```
metadata
metadata.visualize()
```
For more details about how to build the `Metadata` for your own dataset,
please refer to the [relational_metadata](relational_metadata.ipynb)
Guide.
2. A dictionary containing three `pandas.DataFrames` with the tables
described in the metadata object.
```
from sdv.utils import display_tables
display_tables(tables)
for name, table in tables.items():
print(name, table.shape)
```
Let us now use the `HMA1` class to learn this data to be ready to sample
synthetic data about new users. In order to do this you will need to:
- Import the `sdv.relational.HMA1` class and create an instance of it
passing the `metadata` that we just loaded.
- Call its `fit` method passing the `tables` dict.
```
from sdv.relational import HMA1
model = HMA1(metadata)
model.fit(tables)
```
<div class="alert alert-info">
**Note**
During the previous steps SDV walked through all the tables in the
dataset following the relationships specified by the metadata, learned
each table using a [gaussian_copula](gaussian_copula.ipynb) and
then augmented the parent tables using the copula parameters before
learning them. By doing this, each copula model was able to learn how
the child table rows were related to their parent tables.
</div>
### Generate synthetic data from the model
Once the training process has finished you are ready to generate new
synthetic data by calling the `sample_all` method from your model.
```
new_data = model.sample()
```
This will return a dictionary of tables identical to the one which the
model was fitted on, but filled with new data which resembles the
original one.
```
display_tables(new_data)
for name, table in new_data.items():
print(name, table.shape)
```
### Save and Load the model
In many scenarios it will be convenient to generate synthetic versions
of your data directly in systems that do not have access to the original
data source. For example, if you may want to generate testing data on
the fly inside a testing environment that does not have access to your
production database. In these scenarios, fitting the model with real
data every time that you need to generate new data is feasible, so you
will need to fit a model in your production environment, save the fitted
model into a file, send this file to the testing environment and then
load it there to be able to `sample` from it.
Let's see how this process works.
#### Save and share the model
Once you have fitted the model, all you need to do is call its `save`
method passing the name of the file in which you want to save the model.
Note that the extension of the filename is not relevant, but we will be
using the `.pkl` extension to highlight that the serialization protocol
used is [pickle](https://docs.python.org/3/library/pickle.html).
```
model.save('my_model.pkl')
```
This will have created a file called `my_model.pkl` in the same
directory in which you are running SDV.
<div class="alert alert-info">
**Important**
If you inspect the generated file you will notice that its size is much
smaller than the size of the data that you used to generate it. This is
because the serialized model contains **no information about the
original data**, other than the parameters it needs to generate
synthetic versions of it. This means that you can safely share this
`my_model.pkl` file without the risc of disclosing any of your real
data!
</div>
#### Load the model and generate new data
The file you just generated can be send over to the system where the
synthetic data will be generated. Once it is there, you can load it
using the `HMA1.load` method, and then you are ready to sample new data
from the loaded instance:
```
loaded = HMA1.load('my_model.pkl')
new_data = loaded.sample()
new_data.keys()
```
<div class="alert alert-warning">
**Warning**
Notice that the system where the model is loaded needs to also have
`sdv` installed, otherwise it will not be able to load the model and use
it.
</div>
### How to control the number of rows?
In the steps above we did not tell the model at any moment how many rows
we wanted to sample, so it produced as many rows as there were in the
original dataset.
If you want to produce a different number of rows you can pass it as the
`num_rows` argument and it will produce the indicated number of rows:
```
model.sample(num_rows=5)
```
<div class="alert alert-info">
**Note**
Notice that the root table `users` has the indicated number of rows but
some of the other tables do not. This is because the number of rows from
the child tables is sampled based on the values form the parent table,
which means that only the root table of the dataset is affected by the
passed `num_rows` argument.
</div>
### Can I sample a subset of the tables?
In some occasions you will not be interested in generating rows for the
entire dataset and would rather generate data for only one table and its
children.
To do this you can simply pass the name of table that you want to
sample.
For example, pass the name `sessions` to the `sample` method, the model
will only generate data for the `sessions` table and its child table,
`transactions`.
```
model.sample('sessions', num_rows=5)
```
If you want to further restrict the sampling process to only one table
and also skip its child tables, you can add the argument
`sample_children=False`.
For example, you can sample data from the table `users` only without
producing any rows for the tables `sessions` and `transactions`.
```
model.sample('users', num_rows=5, sample_children=False)
```
<div class="alert alert-info">
**Note**
In this case, since we are only producing a single table, the output is
given directly as a `pandas.DataFrame` instead of a dictionary.
</div>
| github_jupyter |
# Week 9
## In-Class Activity Workbook and Homework
## Learning Objectives
### In this notebook you will learn about and practice:
1. Section 1: <a id='Section 1'></a>[Section 1: Reading and examining files with pandas](#Section-1)
2. Section 2: <a id='Section 2'></a>[Section 2: Selecting Data with pandas](#Section-2)
3. Section 3: <a id='Section 3'></a>[Section 3: More practice selecting data](#Section-3)
4. Section 4: <a id='Section 4'></a>[Section 4: Conditional Selection](#Section-4)
### Additional Sources
>- Check out the `pandas` cheat sheets provided by Data Camp and posted on Canvas
>>- https://www.datacamp.com/community/blog/python-pandas-cheat-sheet
# Section 1
## What is the `pandas` module?
>- pandas is a flexible data analysis library built within the C language and is one of the fastest ways of getting from zero to answer
>- `pandas` is the go to tool for most business analysts and scientists working in python and learning to proficient in `pandas` will do wonders to your productivity and look great on your resume
>- Some say `pandas` is basically Excel on steroids
>- `pandas` can be thought of as a mix of Python and SQL so if you know SQL working with `pandas` may come easier to you but knowing SQL is not a prerequisite for working in `pandas`
### Some of the useful ways in which you can use the `pandas` module include:
1. Transforming tabular data into python to work with
2. Cleaning and filtering data, whether it's missing or incomplete
3. Feature engineer new columns that can be applied in your analysis
4. Calculating statistics that answer questions (mean, median, max, min, etc)
5. Finding correlations between columns
6. Visualizing data with matplotlib
## Reading and Writing Files with the Python `pandas` Module
### Read csv or Excel files
>- csv files: `pd.read_csv('fileName.csv')`
>- Excel files: `pd.read_excel('fileName.xlsx')`
>- Multiple sheets from the same Excel file:
>>- `xlsx = pd.ExcelFile('file.xls')` # reads in the entire workbook
>>- `df1 = pd.read_excel(xlsx, 'Sheet1')` # reads in sheet you specify
>>- `df2 = pd.read_excel(xlsx, 'Sheet2')`
### Write csv or Excel files
>- csv files: `pd.to_csv('YourDataFrame.csv')`
>- Excel files: `pd.to_excel('YourDataFrame.xlsx')`
# Section 1
## Reading Files and Initial Data Examination with `pandas`
## Creating a `stu` DataFrame
>- Complete the following steps to practice reading in a csv file
>>- Note: You should download the `students.csv` and `students100.xlsx` files from Canvas and save it in the same director/folder that you have this notebook saved in
1. Import the pandas module and alias it `pd`
### Step 1: Check your working directory and make sure you have the `students.csv` and `students100.xlxs` files there
>- Note: There are several ways to do this
### Step 2: import the `pandas` module and alias it `pd`
### Step 3: Read the `students.csv` file into a pandas dataframe named, `stu`
>- Look at the first five records to make sure `stu` is imported correctly
#### Loading a CSV file
function: `pd.read_csv()`
[Docu read_csv](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.read_csv.html?highlight=read_excel#pandas.read_excel)
#### Now, set the index column of `stu` to the studentID column
>- Note: make sure to make this change in-place
>- There are several ways to do this. Below are couple of options
1. Use the `index_col` option when reading in the file
2. Look up the `set_index` method and apply it to `stu`
#### Look at the first five records after you have set the index
#### Show the last five records of `stu`
#### Show a tuple of the number of columns and rows in `stu`
#### Show the number rows another way
#### Show the columns in `stu`
#### Show the datatypes that are in `stu`
# Section 2
## Accessing Data using `pandas`
#### Access all the records of the `firstName` Column Only
>- Try doing this three different ways
#### Show all the data for rows 3 to 10
#### Show all the data in the last row
#### Show the first 3 records of the third column
>- Try doing this in three different ways
#### Show the last 3 records of the first and last columns
>- Try doing this in three different ways
# Section 3
## More practice and notes on selecting data
>- `iloc` - index based selection
>- `loc` - label based selection
### Additional Source for DataFrame Methods
[Docu DF Methods](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.html)
## `.iloc(arg1, arg2)`
accesses data by row and column (i stands for integer)
`arg1`: row number
`arg2`: column number
first row has index 0 <br>
first column has index 0 <br>
index column is always shown (not included in column count)
### First, let's remind ourselves what the `stu` DataFrame looks like
>- Show the first five records of the `stu` DataFrame
### Using `iloc()`, show the first ten records of the first 2 columns of `stu`
### Using `iloc()` show rows 2-10 for the last two columns of `stu`
### Using `iloc()` show rows 85 to the end of the DataFrame for all columns
### Using `iloc()`, show columns 2 to 4 (lastName to Points) for all records
### Using `iloc()`, show rows 20 to 25, columns 1 to 2 (firstname and lastname)
### Using `iloc()`, show rows 1,5,7 and columns 1,4 (firstName and Points)
## `.loc`, label-based selection
>- Use index values to show specific rows
>- Use column labels to show specific columns
### Using `loc` show students with student number 75-80
### Using `loc` show the `birthdate` of studentIds of 10-15
### Using `loc`, show the birthdate and points for student ids between 3 and 8
### Using `loc`, show the last name and points for student ids of 1, 50, and 100
# Section 4
## Conditional Selection with `loc`
### Using `loc` show the students with point values greater than 90
### Using `loc` show students with points between 50 and 60
### Show students with points either less than 10 or greater than 90
### Find students with last name 'Holmes'
### Find students whose birth date is in 1989
>- Hint: apply `.str.endswith()` when accessing the `birthdate` column
>>- Check the data type of the birthdate field to understand why we can use a string method on a date
### Find students whose first name starts with 'A'
### Find students whose `lastName` starts with 'C' that have 90 or more `Points`
### Find students whose `lastName` starts with 'C' or 'B' with 80 or more `Points`
| github_jupyter |
[Table of Contents](http://nbviewer.ipython.org/github/rlabbe/Kalman-and-Bayesian-Filters-in-Python/blob/master/table_of_contents.ipynb)
# FilterPy Source
For your convienence I have loaded several of FilterPy's core algorithms into this appendix.
## KalmanFilter
```
# %load https://raw.githubusercontent.com/rlabbe/filterpy/master/filterpy/kalman/kalman_filter.py
"""Copyright 2014 Roger R Labbe Jr.
filterpy library.
http://github.com/rlabbe/filterpy
Documentation at:
https://filterpy.readthedocs.org
Supporting book at:
https://github.com/rlabbe/Kalman-and-Bayesian-Filters-in-Python
This is licensed under an MIT license. See the readme.MD file
for more information.
"""
from __future__ import (absolute_import, division, print_function,
unicode_literals)
import numpy as np
import scipy.linalg as linalg
from numpy import dot, zeros, eye, isscalar
from filterpy.common import setter, setter_1d, setter_scalar, dot3
class KalmanFilter(object):
""" Implements a Kalman filter. You are responsible for setting the
various state variables to reasonable values; the defaults will
not give you a functional filter.
You will have to set the following attributes after constructing this
object for the filter to perform properly. Please note that there are
various checks in place to ensure that you have made everything the
'correct' size. However, it is possible to provide incorrectly sized
arrays such that the linear algebra can not perform an operation.
It can also fail silently - you can end up with matrices of a size that
allows the linear algebra to work, but are the wrong shape for the problem
you are trying to solve.
**Attributes**
x : numpy.array(dim_x, 1)
state estimate vector
P : numpy.array(dim_x, dim_x)
covariance estimate matrix
R : numpy.array(dim_z, dim_z)
measurement noise matrix
Q : numpy.array(dim_x, dim_x)
process noise matrix
F : numpy.array()
State Transition matrix
H : numpy.array(dim_x, dim_x)
You may read the following attributes.
**Readable Attributes**
y : numpy.array
Residual of the update step.
K : numpy.array(dim_x, dim_z)
Kalman gain of the update step
S : numpy.array
Systen uncertaintly projected to measurement space
"""
def __init__(self, dim_x, dim_z, dim_u=0):
""" Create a Kalman filter. You are responsible for setting the
various state variables to reasonable values; the defaults below will
not give you a functional filter.
**Parameters**
dim_x : int
Number of state variables for the Kalman filter. For example, if
you are tracking the position and velocity of an object in two
dimensions, dim_x would be 4.
This is used to set the default size of P, Q, and u
dim_z : int
Number of of measurement inputs. For example, if the sensor
provides you with position in (x,y), dim_z would be 2.
dim_u : int (optional)
size of the control input, if it is being used.
Default value of 0 indicates it is not used.
"""
assert dim_x > 0
assert dim_z > 0
assert dim_u >= 0
self.dim_x = dim_x
self.dim_z = dim_z
self.dim_u = dim_u
self._x = zeros((dim_x,1)) # state
self._P = eye(dim_x) # uncertainty covariance
self._Q = eye(dim_x) # process uncertainty
self._B = 0 # control transition matrix
self._F = 0 # state transition matrix
self.H = 0 # Measurement function
self.R = eye(dim_z) # state uncertainty
self._alpha_sq = 1. # fading memory control
# gain and residual are computed during the innovation step. We
# save them so that in case you want to inspect them for various
# purposes
self._K = 0 # kalman gain
self._y = zeros((dim_z, 1))
self._S = 0 # system uncertainty in measurement space
# identity matrix. Do not alter this.
self._I = np.eye(dim_x)
def update(self, z, R=None, H=None):
"""
Add a new measurement (z) to the kalman filter. If z is None, nothing
is changed.
**Parameters**
z : np.array
measurement for this update.
R : np.array, scalar, or None
Optionally provide R to override the measurement noise for this
one call, otherwise self.R will be used.
"""
if z is None:
return
if R is None:
R = self.R
elif isscalar(R):
R = eye(self.dim_z) * R
# rename for readability and a tiny extra bit of speed
if H is None:
H = self.H
P = self._P
x = self._x
# y = z - Hx
# error (residual) between measurement and prediction
self._y = z - dot(H, x)
# S = HPH' + R
# project system uncertainty into measurement space
S = dot3(H, P, H.T) + R
# K = PH'inv(S)
# map system uncertainty into kalman gain
K = dot3(P, H.T, linalg.inv(S))
# x = x + Ky
# predict new x with residual scaled by the kalman gain
self._x = x + dot(K, self._y)
# P = (I-KH)P(I-KH)' + KRK'
I_KH = self._I - dot(K, H)
self._P = dot3(I_KH, P, I_KH.T) + dot3(K, R, K.T)
self._S = S
self._K = K
def test_matrix_dimensions(self):
""" Performs a series of asserts to check that the size of everything
is what it should be. This can help you debug problems in your design.
This is only a test; you do not need to use it while filtering.
However, to use you will want to perform at least one predict() and
one update() before calling; some bad designs will cause the shapes
of x and P to change in a silent and bad way. For example, if you
pass in a badly dimensioned z into update that can cause x to be
misshapen."""
assert self._x.shape == (self.dim_x, 1), \
"Shape of x must be ({},{}), but is {}".format(
self.dim_x, 1, self._x.shape)
assert self._P.shape == (self.dim_x, self.dim_x), \
"Shape of P must be ({},{}), but is {}".format(
self.dim_x, self.dim_x, self._P.shape)
assert self._Q.shape == (self.dim_x, self.dim_x), \
"Shape of P must be ({},{}), but is {}".format(
self.dim_x, self.dim_x, self._P.shape)
def predict(self, u=0):
""" Predict next position.
**Parameters**
u : np.array
Optional control vector. If non-zero, it is multiplied by B
to create the control input into the system.
"""
# x = Fx + Bu
self._x = dot(self._F, self.x) + dot(self._B, u)
# P = FPF' + Q
self._P = self._alpha_sq * dot3(self._F, self._P, self._F.T) + self._Q
def batch_filter(self, zs, Rs=None, update_first=False):
""" Batch processes a sequences of measurements.
**Parameters**
zs : list-like
list of measurements at each time step `self.dt` Missing
measurements must be represented by 'None'.
Rs : list-like, optional
optional list of values to use for the measurement error
covariance; a value of None in any position will cause the filter
to use `self.R` for that time step.
update_first : bool, optional,
controls whether the order of operations is update followed by
predict, or predict followed by update. Default is predict->update.
**Returns**
means: np.array((n,dim_x,1))
array of the state for each time step after the update. Each entry
is an np.array. In other words `means[k,:]` is the state at step
`k`.
covariance: np.array((n,dim_x,dim_x))
array of the covariances for each time step after the update.
In other words `covariance[k,:,:]` is the covariance at step `k`.
means_predictions: np.array((n,dim_x,1))
array of the state for each time step after the predictions. Each
entry is an np.array. In other words `means[k,:]` is the state at
step `k`.
covariance_predictions: np.array((n,dim_x,dim_x))
array of the covariances for each time step after the prediction.
In other words `covariance[k,:,:]` is the covariance at step `k`.
"""
try:
z = zs[0]
except:
assert not isscalar(zs), 'zs must be list-like'
if self.dim_z == 1:
assert isscalar(z) or (z.ndim==1 and len(z) == 1), \
'zs must be a list of scalars or 1D, 1 element arrays'
else:
assert len(z) == self.dim_z, 'each element in zs must be a'
'1D array of length {}'.format(self.dim_z)
n = np.size(zs,0)
if Rs is None:
Rs = [None]*n
# mean estimates from Kalman Filter
if self.x.ndim == 1:
means = zeros((n, self.dim_x))
means_p = zeros((n, self.dim_x))
else:
means = zeros((n, self.dim_x, 1))
means_p = zeros((n, self.dim_x, 1))
# state covariances from Kalman Filter
covariances = zeros((n, self.dim_x, self.dim_x))
covariances_p = zeros((n, self.dim_x, self.dim_x))
if update_first:
for i, (z, r) in enumerate(zip(zs, Rs)):
self.update(z, r)
means[i,:] = self._x
covariances[i,:,:] = self._P
self.predict()
means_p[i,:] = self._x
covariances_p[i,:,:] = self._P
else:
for i, (z, r) in enumerate(zip(zs, Rs)):
self.predict()
means_p[i,:] = self._x
covariances_p[i,:,:] = self._P
self.update(z, r)
means[i,:] = self._x
covariances[i,:,:] = self._P
return (means, covariances, means_p, covariances_p)
def rts_smoother(self, Xs, Ps, Qs=None):
""" Runs the Rauch-Tung-Striebal Kalman smoother on a set of
means and covariances computed by a Kalman filter. The usual input
would come from the output of `KalmanFilter.batch_filter()`.
**Parameters**
Xs : numpy.array
array of the means (state variable x) of the output of a Kalman
filter.
Ps : numpy.array
array of the covariances of the output of a kalman filter.
Q : list-like collection of numpy.array, optional
Process noise of the Kalman filter at each time step. Optional,
if not provided the filter's self.Q will be used
**Returns**
'x' : numpy.ndarray
smoothed means
'P' : numpy.ndarray
smoothed state covariances
'K' : numpy.ndarray
smoother gain at each step
**Example**::
zs = [t + random.randn()*4 for t in range (40)]
(mu, cov, _, _) = kalman.batch_filter(zs)
(x, P, K) = rts_smoother(mu, cov, fk.F, fk.Q)
"""
assert len(Xs) == len(Ps)
shape = Xs.shape
n = shape[0]
dim_x = shape[1]
F = self._F
if not Qs:
Qs = [self.Q] * n
# smoother gain
K = zeros((n,dim_x,dim_x))
x, P = Xs.copy(), Ps.copy()
for k in range(n-2,-1,-1):
P_pred = dot3(F, P[k], F.T) + Qs[k]
K[k] = dot3(P[k], F.T, linalg.inv(P_pred))
x[k] += dot (K[k], x[k+1] - dot(F, x[k]))
P[k] += dot3 (K[k], P[k+1] - P_pred, K[k].T)
return (x, P, K)
def get_prediction(self, u=0):
""" Predicts the next state of the filter and returns it. Does not
alter the state of the filter.
**Parameters**
u : np.array
optional control input
**Returns**
(x, P)
State vector and covariance array of the prediction.
"""
x = dot(self._F, self._x) + dot(self._B, u)
P = self._alpha_sq * dot3(self._F, self._P, self._F.T) + self._Q
return (x, P)
def residual_of(self, z):
""" returns the residual for the given measurement (z). Does not alter
the state of the filter.
"""
return z - dot(self.H, self._x)
def measurement_of_state(self, x):
""" Helper function that converts a state into a measurement.
**Parameters**
x : np.array
kalman state vector
**Returns**
z : np.array
measurement corresponding to the given state
"""
return dot(self.H, x)
@property
def alpha(self):
""" Fading memory setting. 1.0 gives the normal Kalman filter, and
values slightly larger than 1.0 (such as 1.02) give a fading
memory effect - previous measurements have less influence on the
filter's estimates. This formulation of the Fading memory filter
(there are many) is due to Dan Simon [1].
** References **
[1] Dan Simon. "Optimal State Estimation." John Wiley & Sons.
p. 208-212. (2006)
"""
return self._alpha_sq**.5
@alpha.setter
def alpha(self, value):
assert np.isscalar(value)
assert value > 0
self._alpha_sq = value**2
@property
def Q(self):
""" Process uncertainty"""
return self._Q
@Q.setter
def Q(self, value):
self._Q = setter_scalar(value, self.dim_x)
@property
def P(self):
""" covariance matrix"""
return self._P
@P.setter
def P(self, value):
self._P = setter_scalar(value, self.dim_x)
@property
def F(self):
""" state transition matrix"""
return self._F
@F.setter
def F(self, value):
self._F = setter(value, self.dim_x, self.dim_x)
@property
def B(self):
""" control transition matrix"""
return self._B
@B.setter
def B(self, value):
""" control transition matrix"""
self._B = setter (value, self.dim_x, self.dim_u)
@property
def x(self):
""" filter state vector."""
return self._x
@x.setter
def x(self, value):
self._x = setter_1d(value, self.dim_x)
@property
def K(self):
""" Kalman gain """
return self._K
@property
def y(self):
""" measurement residual (innovation) """
return self._y
@property
def S(self):
""" system uncertainty in measurement space """
return self._S
```
## ExtendedKalmanFilter
```
# %load https://raw.githubusercontent.com/rlabbe/filterpy/master/filterpy/kalman/EKF.py
"""Copyright 2014 Roger R Labbe Jr.
filterpy library.
http://github.com/rlabbe/filterpy
Documentation at:
https://filterpy.readthedocs.org
Supporting book at:
https://github.com/rlabbe/Kalman-and-Bayesian-Filters-in-Python
This is licensed under an MIT license. See the readme.MD file
for more information.
"""
from __future__ import (absolute_import, division, print_function,
unicode_literals)
import numpy as np
import scipy.linalg as linalg
from numpy import dot, zeros, eye
from filterpy.common import setter, setter_1d, setter_scalar, dot3
class ExtendedKalmanFilter(object):
def __init__(self, dim_x, dim_z, dim_u=0):
""" Extended Kalman filter. You are responsible for setting the
various state variables to reasonable values; the defaults below will
not give you a functional filter.
**Parameters**
dim_x : int
Number of state variables for the Kalman filter. For example, if
you are tracking the position and velocity of an object in two
dimensions, dim_x would be 4.
This is used to set the default size of P, Q, and u
dim_z : int
Number of of measurement inputs. For example, if the sensor
provides you with position in (x,y), dim_z would be 2.
"""
self.dim_x = dim_x
self.dim_z = dim_z
self._x = zeros((dim_x,1)) # state
self._P = eye(dim_x) # uncertainty covariance
self._B = 0 # control transition matrix
self._F = 0 # state transition matrix
self._R = eye(dim_z) # state uncertainty
self._Q = eye(dim_x) # process uncertainty
self._y = zeros((dim_z, 1))
# identity matrix. Do not alter this.
self._I = np.eye(dim_x)
def predict_update(self, z, HJacobian, Hx, u=0):
""" Performs the predict/update innovation of the extended Kalman
filter.
**Parameters**
z : np.array
measurement for this step.
If `None`, only predict step is perfomed.
HJacobian : function
function which computes the Jacobian of the H matrix (measurement
function). Takes state variable (self.x) as input, returns H.
Hx : function
function which takes a state variable and returns the measurement
that would correspond to that state.
u : np.array or scalar
optional control vector input to the filter.
"""
if np.isscalar(z) and self.dim_z == 1:
z = np.asarray([z], float)
F = self._F
B = self._B
P = self._P
Q = self._Q
R = self._R
x = self._x
H = HJacobian(x)
# predict step
x = dot(F, x) + dot(B, u)
P = dot3(F, P, F.T) + Q
# update step
S = dot3(H, P, H.T) + R
K = dot3(P, H.T, linalg.inv (S))
self._x = x + dot(K, (z - Hx(x)))
I_KH = self._I - dot(K, H)
self._P = dot3(I_KH, P, I_KH.T) + dot3(K, R, K.T)
def update(self, z, HJacobian, Hx, R=None):
""" Performs the update innovation of the extended Kalman filter.
**Parameters**
z : np.array
measurement for this step.
If `None`, only predict step is perfomed.
HJacobian : function
function which computes the Jacobian of the H matrix (measurement
function). Takes state variable (self.x) as input, returns H.
Hx : function
function which takes a state variable and returns the measurement
that would correspond to that state.
"""
P = self._P
if R is None:
R = self._R
elif np.isscalar(R):
R = eye(self.dim_z) * R
if np.isscalar(z) and self.dim_z == 1:
z = np.asarray([z], float)
x = self._x
H = HJacobian(x)
S = dot3(H, P, H.T) + R
K = dot3(P, H.T, linalg.inv (S))
y = z - Hx(x)
self._x = x + dot(K, y)
I_KH = self._I - dot(K, H)
self._P = dot3(I_KH, P, I_KH.T) + dot3(K, R, K.T)
def predict_x(self, u=0):
""" predicts the next state of X. If you need to
compute the next state yourself, override this function. You would
need to do this, for example, if the usual Taylor expansion to
generate F is not providing accurate results for you. """
self._x = dot(self._F, self._x) + dot(self._B, u)
def predict(self, u=0):
""" Predict next position.
**Parameters**
u : np.array
Optional control vector. If non-zero, it is multiplied by B
to create the control input into the system.
"""
self.predict_x(u)
self._P = dot3(self._F, self._P, self._F.T) + self._Q
@property
def Q(self):
""" Process uncertainty"""
return self._Q
@Q.setter
def Q(self, value):
self._Q = setter_scalar(value, self.dim_x)
@property
def P(self):
""" covariance matrix"""
return self._P
@P.setter
def P(self, value):
self._P = setter_scalar(value, self.dim_x)
@property
def R(self):
""" measurement uncertainty"""
return self._R
@R.setter
def R(self, value):
self._R = setter_scalar(value, self.dim_z)
@property
def F(self):
return self._F
@F.setter
def F(self, value):
self._F = setter(value, self.dim_x, self.dim_x)
@property
def B(self):
return self._B
@B.setter
def B(self, value):
""" control transition matrix"""
self._B = setter(value, self.dim_x, self.dim_u)
@property
def x(self):
return self._x
@x.setter
def x(self, value):
self._x = setter_1d(value, self.dim_x)
@property
def K(self):
""" Kalman gain """
return self._K
@property
def y(self):
""" measurement residual (innovation) """
return self._y
@property
def S(self):
""" system uncertainty in measurement space """
return self._S
```
## UnscentedKalmanFilter
```
# %load https://raw.githubusercontent.com/rlabbe/filterpy/master/filterpy/kalman/UKF.py
"""Copyright 2014 Roger R Labbe Jr.
filterpy library.
http://github.com/rlabbe/filterpy
Documentation at:
https://filterpy.readthedocs.org
Supporting book at:
https://github.com/rlabbe/Kalman-and-Bayesian-Filters-in-Python
This is licensed under an MIT license. See the readme.MD file
for more information.
"""
# pylint bug - warns about numpy functions which do in fact exist.
# pylint: disable=E1101
#I like aligning equal signs for readability of math
# pylint: disable=C0326
from __future__ import (absolute_import, division, print_function,
unicode_literals)
from numpy.linalg import inv, cholesky
import numpy as np
from numpy import asarray, eye, zeros, dot, isscalar, outer
from filterpy.common import dot3
class UnscentedKalmanFilter(object):
# pylint: disable=too-many-instance-attributes
# pylint: disable=C0103
""" Implements the Unscented Kalman filter (UKF) as defined by Simon J.
Julier and Jeffery K. Uhlmann [1]. Succintly, the UKF selects a set of
sigma points and weights inside the covariance matrix of the filter's
state. These points are transformed through the nonlinear process being
filtered, and are rebuilt into a mean and covariance by computed the
weighted mean and expected value of the transformed points. Read the paper;
it is excellent. My book "Kalman and Bayesian Filters in Python" [2]
explains the algorithm, develops this code, and provides examples of the
filter in use.
You will have to set the following attributes after constructing this
object for the filter to perform properly.
**Attributes**
x : numpy.array(dim_x)
state estimate vector
P : numpy.array(dim_x, dim_x)
covariance estimate matrix
R : numpy.array(dim_z, dim_z)
measurement noise matrix
Q : numpy.array(dim_x, dim_x)
process noise matrix
You may read the following attributes.
**Readable Attributes**
Pxz : numpy.aray(dim_x, dim_z)
Cross variance of x and z computed during update() call.
**References**
.. [1] Julier, Simon J.; Uhlmann, Jeffrey "A New Extension of the Kalman
Filter to Nonlinear Systems". Proc. SPIE 3068, Signal Processing,
Sensor Fusion, and Target Recognition VI, 182 (July 28, 1997)
.. [2] Labbe, Roger R. "Kalman and Bayesian Filters in Python"
https://github.com/rlabbe/Kalman-and-Bayesian-Filters-in-Python
"""
def __init__(self, dim_x, dim_z, dt, hx, fx, kappa=0.):
""" Create a Kalman filter. You are responsible for setting the
various state variables to reasonable values; the defaults below will
not give you a functional filter.
**Parameters**
dim_x : int
Number of state variables for the filter. For example, if
you are tracking the position and velocity of an object in two
dimensions, dim_x would be 4.
dim_z : int
Number of of measurement inputs. For example, if the sensor
provides you with position in (x,y), dim_z would be 2.
dt : float
Time between steps in seconds.
hx : function(x)
Measurement function. Converts state vector x into a measurement
vector of shape (dim_z).
fx : function(x,dt)
function that returns the state x transformed by the
state transistion function. dt is the time step in seconds.
kappa : float, default=0.
Scaling factor that can reduce high order errors. kappa=0 gives
the standard unscented filter. According to [1], if you set
kappa to 3-dim_x for a Gaussian x you will minimize the fourth
order errors in x and P.
**References**
[1] S. Julier, J. Uhlmann, and H. Durrant-Whyte. "A new method for
the nonlinear transformation of means and covariances in filters
and estimators," IEEE Transactions on Automatic Control, 45(3),
pp. 477-482 (March 2000).
"""
self.Q = eye(dim_x)
self.R = eye(dim_z)
self.x = zeros(dim_x)
self.P = eye(dim_x)
self._dim_x = dim_x
self._dim_z = dim_z
self._dt = dt
self._num_sigmas = 2*dim_x + 1
self.kappa = kappa
self.hx = hx
self.fx = fx
# weights for the sigma points
self.W = self.weights(dim_x, kappa)
# sigma points transformed through f(x) and h(x)
# variables for efficiency so we don't recreate every update
self.sigmas_f = zeros((self._num_sigmas, self._dim_x))
def update(self, z, R=None, residual=np.subtract, UT=None):
""" Update the UKF with the given measurements. On return,
self.x and self.P contain the new mean and covariance of the filter.
**Parameters**
z : numpy.array of shape (dim_z)
measurement vector
R : numpy.array((dim_z, dim_z)), optional
Measurement noise. If provided, overrides self.R for
this function call.
residual : function (z, z2), optional
Optional function that computes the residual (difference) between
the two measurement vectors. If you do not provide this, then the
built in minus operator will be used. You will normally want to use
the built in unless your residual computation is nonlinear (for
example, if they are angles)
UT : function(sigmas, Wm, Wc, noise_cov), optional
Optional function to compute the unscented transform for the sigma
points passed through hx. Typically the default function will
work, but if for example you are using angles the default method
of computing means and residuals will not work, and you will have
to define how to compute it.
"""
if isscalar(z):
dim_z = 1
else:
dim_z = len(z)
if R is None:
R = self.R
elif np.isscalar(R):
R = eye(self._dim_z) * R
# rename for readability
sigmas_f = self.sigmas_f
sigmas_h = zeros((self._num_sigmas, dim_z))
if UT is None:
UT = unscented_transform
# transform sigma points into measurement space
for i in range(self._num_sigmas):
sigmas_h[i] = self.hx(sigmas_f[i])
# mean and covariance of prediction passed through inscented transform
zp, Pz = UT(sigmas_h, self.W, self.W, R)
# compute cross variance of the state and the measurements
'''self.Pxz = zeros((self._dim_x, dim_z))
for i in range(self._num_sigmas):
self.Pxz += self.W[i] * np.outer(sigmas_f[i] - self.x,
residual(sigmas_h[i], zp))'''
# this is the unreadable but fast implementation of the
# commented out loop above
yh = sigmas_f - self.x[np.newaxis, :]
yz = residual(sigmas_h, zp[np.newaxis, :])
self.Pxz = yh.T.dot(np.diag(self.W)).dot(yz)
K = dot(self.Pxz, inv(Pz)) # Kalman gain
y = residual(z, zp)
self.x = self.x + dot(K, y)
self.P = self.P - dot3(K, Pz, K.T)
def predict(self, dt=None):
""" Performs the predict step of the UKF. On return, self.xp and
self.Pp contain the predicted state (xp) and covariance (Pp). 'p'
stands for prediction.
**Parameters**
dt : double, optional
If specified, the time step to be used for this prediction.
self._dt is used if this is not provided.
Important: this MUST be called before update() is called for the
first time.
"""
if dt is None:
dt = self._dt
# calculate sigma points for given mean and covariance
sigmas = self.sigma_points(self.x, self.P, self.kappa)
for i in range(self._num_sigmas):
self.sigmas_f[i] = self.fx(sigmas[i], dt)
self.x, self.P = unscented_transform(
self.sigmas_f, self.W, self.W, self.Q)
def batch_filter(self, zs, Rs=None, residual=np.subtract, UT=None):
""" Performs the UKF filter over the list of measurement in `zs`.
**Parameters**
zs : list-like
list of measurements at each time step `self._dt` Missing
measurements must be represented by 'None'.
Rs : list-like, optional
optional list of values to use for the measurement error
covariance; a value of None in any position will cause the filter
to use `self.R` for that time step.
residual : function (z, z2), optional
Optional function that computes the residual (difference) between
the two measurement vectors. If you do not provide this, then the
built in minus operator will be used. You will normally want to use
the built in unless your residual computation is nonlinear (for
example, if they are angles)
UT : function(sigmas, Wm, Wc, noise_cov), optional
Optional function to compute the unscented transform for the sigma
points passed through hx. Typically the default function will
work, but if for example you are using angles the default method
of computing means and residuals will not work, and you will have
to define how to compute it.
**Returns**
means: np.array((n,dim_x,1))
array of the state for each time step after the update. Each entry
is an np.array. In other words `means[k,:]` is the state at step
`k`.
covariance: np.array((n,dim_x,dim_x))
array of the covariances for each time step after the update.
In other words `covariance[k,:,:]` is the covariance at step `k`.
"""
try:
z = zs[0]
except:
assert not isscalar(zs), 'zs must be list-like'
if self._dim_z == 1:
assert isscalar(z) or (z.ndim==1 and len(z) == 1), \
'zs must be a list of scalars or 1D, 1 element arrays'
else:
assert len(z) == self._dim_z, 'each element in zs must be a' \
'1D array of length {}'.format(self._dim_z)
n = np.size(zs,0)
if Rs is None:
Rs = [None]*n
# mean estimates from Kalman Filter
if self.x.ndim == 1:
means = zeros((n, self._dim_x))
else:
means = zeros((n, self._dim_x, 1))
# state covariances from Kalman Filter
covariances = zeros((n, self._dim_x, self._dim_x))
for i, (z, r) in enumerate(zip(zs, Rs)):
self.predict()
self.update(z, r)
means[i,:] = self.x
covariances[i,:,:] = self.P
return (means, covariances)
def rts_smoother(self, Xs, Ps, Qs=None, dt=None):
""" Runs the Rauch-Tung-Striebal Kalman smoother on a set of
means and covariances computed by the UKF. The usual input
would come from the output of `batch_filter()`.
**Parameters**
Xs : numpy.array
array of the means (state variable x) of the output of a Kalman
filter.
Ps : numpy.array
array of the covariances of the output of a kalman filter.
Q : list-like collection of numpy.array, optional
Process noise of the Kalman filter at each time step. Optional,
if not provided the filter's self.Q will be used
dt : optional, float or array-like of float
If provided, specifies the time step of each step of the filter.
If float, then the same time step is used for all steps. If
an array, then each element k contains the time at step k.
Units are seconds.
**Returns**
'x' : numpy.ndarray
smoothed means
'P' : numpy.ndarray
smoothed state covariances
'K' : numpy.ndarray
smoother gain at each step
**Example**::
zs = [t + random.randn()*4 for t in range (40)]
(mu, cov, _, _) = kalman.batch_filter(zs)
(x, P, K) = rts_smoother(mu, cov, fk.F, fk.Q)
"""
assert len(Xs) == len(Ps)
n, dim_x = Xs.shape
if dt is None:
dt = [self._dt] * n
elif isscalar(dt):
dt = [dt] * n
if Qs is None:
Qs = [self.Q] * n
# smoother gain
Ks = zeros((n,dim_x,dim_x))
num_sigmas = 2*dim_x + 1
xs, ps = Xs.copy(), Ps.copy()
sigmas_f = zeros((num_sigmas, dim_x))
for k in range(n-2,-1,-1):
# create sigma points from state estimate, pass through state func
sigmas = self.sigma_points(xs[k], ps[k], self.kappa)
for i in range(num_sigmas):
sigmas_f[i] = self.fx(sigmas[i], dt[k])
# compute backwards prior state and covariance
xb = dot(self.W, sigmas_f)
Pb = 0
x = Xs[k]
for i in range(num_sigmas):
y = sigmas_f[i] - x
Pb += self.W[i] * outer(y, y)
Pb += Qs[k]
# compute cross variance
Pxb = 0
for i in range(num_sigmas):
z = sigmas[i] - Xs[k]
y = sigmas_f[i] - xb
Pxb += self.W[i] * outer(z, y)
# compute gain
K = dot(Pxb, inv(Pb))
# update the smoothed estimates
xs[k] += dot (K, xs[k+1] - xb)
ps[k] += dot3(K, ps[k+1] - Pb, K.T)
Ks[k] = K
return (xs, ps, Ks)
@staticmethod
def weights(n, kappa):
""" Computes the weights for an unscented Kalman filter. See
__init__() for meaning of parameters.
"""
assert n > 0, "n must be greater than 0, it's value is {}".format(n)
k = .5 / (n+kappa)
W = np.full(2*n+1, k)
W[0] = kappa / (n+kappa)
return W
@staticmethod
def sigma_points(x, P, kappa):
""" Computes the sigma points for an unscented Kalman filter
given the mean (x) and covariance(P) of the filter.
kappa is an arbitrary constant. Returns sigma points.
Works with both scalar and array inputs:
sigma_points (5, 9, 2) # mean 5, covariance 9
sigma_points ([5, 2], 9*eye(2), 2) # means 5 and 2, covariance 9I
**Parameters**
X An array-like object of the means of length n
Can be a scalar if 1D.
examples: 1, [1,2], np.array([1,2])
P : scalar, or np.array
Covariance of the filter. If scalar, is treated as eye(n)*P.
kappa : float
Scaling factor.
**Returns**
sigmas : np.array, of size (n, 2n+1)
2D array of sigma points. Each column contains all of
the sigmas for one dimension in the problem space. They
are ordered as:
.. math::
sigmas[0] = x \n
sigmas[1..n] = x + [\sqrt{(n+\kappa)P}]_k \n
sigmas[n+1..2n] = x - [\sqrt{(n+\kappa)P}]_k
"""
if np.isscalar(x):
x = asarray([x])
n = np.size(x) # dimension of problem
if np.isscalar(P):
P = eye(n)*P
sigmas = zeros((2*n+1, n))
# implements U'*U = (n+kappa)*P. Returns lower triangular matrix.
# Take transpose so we can access with U[i]
U = cholesky((n+kappa)*P).T
#U = sqrtm((n+kappa)*P).T
sigmas[0] = x
sigmas[1:n+1] = x + U
sigmas[n+1:2*n+2] = x - U
return sigmas
def unscented_transform(Sigmas, Wm, Wc, noise_cov):
""" Computes unscented transform of a set of sigma points and weights.
returns the mean and covariance in a tuple.
"""
kmax, n = Sigmas.shape
# new mean is just the sum of the sigmas * weight
x = dot(Wm, Sigmas) # dot = \Sigma^n_1 (W[k]*Xi[k])
# new covariance is the sum of the outer product of the residuals
# times the weights
'''P = zeros((n, n))
for k in range(kmax):
y = Sigmas[k] - x
P += Wc[k] * np.outer(y, y)'''
# this is the fast way to do the commented out code above
y = Sigmas - x[np.newaxis,:]
P = y.T.dot(np.diag(Wc)).dot(y)
if noise_cov is not None:
P += noise_cov
return (x, P)
```
| github_jupyter |
# 3. LightGBM
**Start from the most basic features, and try to improve step by step.**
Kaggle score: 0.9307
## Run name
```
import time
project_name = 'TalkingdataAFD2018'
step_name = 'LightGBM'
time_str = time.strftime("%Y%m%d_%H%M%S", time.localtime())
run_name = '%s_%s_%s' % (project_name, step_name, time_str)
print('run_name: %s' % run_name)
t0 = time.time()
```
## Important params
```
date = 6
print('date: ', date)
# test_n_rows = 18790469
test_n_rows = None
day_rows = {
0: {
'n_skiprows': 1,
'n_rows': 10 * 10000
},
6: {
'n_skiprows': 1,
'n_rows': 9308568
},
7: {
'n_skiprows': 1 + 9308568,
'n_rows': 59633310
},
8: {
'n_skiprows': 1 + 9308568 + 59633310,
'n_rows': 62945075
},
9: {
'n_skiprows': 1 + 9308568 + 59633310 + 62945075,
'n_rows': 53016937
}
}
n_skiprows = day_rows[date]['n_skiprows']
n_rows = day_rows[date]['n_rows']
```
## Import PKGs
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
from IPython.display import display
import os
import gc
import time
import random
import zipfile
import h5py
import pickle
import math
from PIL import Image
import shutil
from tqdm import tqdm
import multiprocessing
from multiprocessing import cpu_count
from sklearn.model_selection import train_test_split, GridSearchCV
from sklearn.metrics import accuracy_score
random_num = np.random.randint(10000)
print('random_num: %s' % random_num)
```
## Project folders
```
cwd = os.getcwd()
input_folder = os.path.join(cwd, 'input')
output_folder = os.path.join(cwd, 'output')
model_folder = os.path.join(cwd, 'model')
log_folder = os.path.join(cwd, 'log')
print('input_folder: \t\t\t%s' % input_folder)
print('output_folder: \t\t\t%s' % output_folder)
print('model_folder: \t\t\t%s' % model_folder)
print('log_folder: \t\t\t%s' % log_folder)
train_csv_file = os.path.join(input_folder, 'train.csv')
train_sample_csv_file = os.path.join(input_folder, 'train_sample.csv')
test_csv_file = os.path.join(input_folder, 'test.csv')
sample_submission_csv_file = os.path.join(input_folder, 'sample_submission.csv')
print('\ntrain_csv_file: \t\t%s' % train_csv_file)
print('train_sample_csv_file: \t\t%s' % train_sample_csv_file)
print('test_csv_file: \t\t\t%s' % test_csv_file)
print('sample_submission_csv_file: \t%s' % sample_submission_csv_file)
```
## Load data
```
%%time
train_csv = pd.read_csv(train_csv_file, skiprows=range(1, n_skiprows), nrows=n_rows, parse_dates=['click_time'])
test_csv = pd.read_csv(test_csv_file, nrows=test_n_rows, parse_dates=['click_time'])
sample_submission_csv = pd.read_csv(sample_submission_csv_file)
print('train_csv.shape: \t\t', train_csv.shape)
print('test_csv.shape: \t\t', test_csv.shape)
print('sample_submission_csv.shape: \t', sample_submission_csv.shape)
print('train_csv.dtypes: \n', train_csv.dtypes)
display(train_csv.head(2))
display(test_csv.head(2))
display(sample_submission_csv.head(2))
y_data = train_csv['is_attributed']
train_csv.drop(['is_attributed'], axis=1, inplace=True)
display(y_data.head())
```
## Features
```
train_csv['day'] = train_csv['click_time'].dt.day.astype('uint8')
train_csv['hour'] = train_csv['click_time'].dt.hour.astype('uint8')
train_csv['minute'] = train_csv['click_time'].dt.minute.astype('uint8')
train_csv['second'] = train_csv['click_time'].dt.second.astype('uint8')
print('train_csv.shape: \t', train_csv.shape)
display(train_csv.head(2))
test_csv['day'] = test_csv['click_time'].dt.day.astype('uint8')
test_csv['hour'] = test_csv['click_time'].dt.hour.astype('uint8')
test_csv['minute'] = test_csv['click_time'].dt.minute.astype('uint8')
test_csv['second'] = test_csv['click_time'].dt.second.astype('uint8')
print('test_csv.shape: \t', test_csv.shape)
display(test_csv.head(2))
```
## Prepare data
```
train_useless_features = ['click_time', 'attributed_time']
train_csv.drop(train_useless_features, axis=1, inplace=True)
test_useless_features = ['click_time', 'click_id']
test_csv.drop(test_useless_features, axis=1, inplace=True)
display(train_csv.head())
display(test_csv.head())
x_train, x_val, y_train, y_val = train_test_split(train_csv, y_data, test_size=0.01, random_state=2017)
x_test = test_csv
print(x_train.shape)
print(y_train.shape)
print(x_val.shape)
print(y_val.shape)
print(x_test.shape)
```
## Train
```
import lightgbm as lgb
from sklearn.metrics import roc_auc_score
lgb_train = lgb.Dataset(x_train, label=y_train)
lgb_val = lgb.Dataset(x_val, label=y_val, reference=lgb_train)
# LightGBM parameters
params = {
'task': 'train',
'num_boost_round': 200,
'early_stopping_rounds': 10,
'boosting_type': 'gbdt', # (default="gbdt")
'num_leaves': 300, # (default=31)
'max_depth': -1, # (default=-1)
'learning_rate': 0.1, # (default=0.1)
'n_estimators': 500, # (default=10)
'max_bin': 30, # (default=255)
'subsample_for_bin': 100*10000, # (default=50000)
'objective': 'binary', # (default=None)
'min_split_gain': 0., # (default=0.)
'min_child_weight': 1e-3, # (default=1e-3)
'min_child_samples': 10, # (default=20)
'subsample': 0.7, # (default=1.)
# 'subsample_freq': 1, # (default=1)
'colsample_bytree': 0.9, # (default=1.)
'reg_alpha': 0., # (default=0.)
'reg_lambda': 0., # (default=0.)
'random_state': random_num, # (default=None)
'n_jobs': -1, # (default=-1)
'silent': False, # (default=True)
'metric': ['auc', 'binary_logloss'],
}
print('params: ', params)
# train
gbm = lgb.train(
params,
train_set=lgb_train,
valid_sets=lgb_val
)
print('*' * 80)
y_train_proba = gbm.predict(x_train, num_iteration=gbm.best_iteration)
y_train_pred = (y_train_proba>=0.5).astype(int)
acc_train = accuracy_score(y_train, y_train_pred)
roc_train = roc_auc_score(y_train, y_train_proba)
print('acc_train: %.4f \t roc_train: %.4f' % (acc_train, roc_train))
y_val_proba = gbm.predict(x_val, num_iteration=gbm.best_iteration)
y_val_pred = (y_val_proba>=0.5).astype(int)
acc_val = accuracy_score(y_val, y_val_pred)
roc_val = roc_auc_score(y_val, y_val_proba)
print('acc_val: %.4f \t roc_val: %.4f' % (acc_val, roc_val))
```
## Predict
```
run_name_acc = run_name + '_' + str(int(roc_val*10000)).zfill(4)
print(run_name_acc)
y_test_proba = gbm.predict(x_test, num_iteration=gbm.best_iteration)
print(y_test_proba.shape)
print(y_test_proba[:20])
def save_proba(y_train_proba, y_train, y_val_proba, y_val, y_test_proba, click_ids, file_name):
print(click_ids[:5])
if os.path.exists(file_name):
os.remove(file_name)
print('File removed: \t%s' % file_name)
with h5py.File(file_name) as h:
h.create_dataset('y_train_proba', data=y_train_proba)
h.create_dataset('y_train', data=y_train)
h.create_dataset('y_val_proba', data=y_val_proba)
h.create_dataset('y_val', data=y_val)
h.create_dataset('y_test_proba', data=y_test_proba)
h.create_dataset('click_ids', data=click_ids)
print('File saved: \t%s' % file_name)
def load_proba(file_name):
with h5py.File(file_name, 'r') as h:
y_train_proba = np.array(h['y_train_proba'])
y_train = np.array(h['y_train'])
y_val_proba = np.array(h['y_val_proba'])
y_val = np.array(h['y_val'])
y_test_proba = np.array(h['y_test_proba'])
click_ids = np.array(h['click_ids'])
print('File loaded: \t%s' % file_name)
print(click_ids[:5])
return y_train_proba, y_train, y_val_proba, y_val, y_test_proba, click_ids
y_proba_file = os.path.join(model_folder, 'proba_%s.p' % run_name_acc)
save_proba(y_train_proba, y_train, y_val_proba, y_val, y_test_proba, np.array(sample_submission_csv['click_id']), y_proba_file)
y_train_proba, y_train, y_val_proba, y_val, y_test_proba, click_ids = load_proba(y_proba_file)
print(y_train_proba.shape)
print(y_train.shape)
print(y_val_proba.shape)
print(y_val.shape)
print(y_test_proba.shape)
print(len(click_ids))
%%time
submission_csv_file = os.path.join(output_folder, 'pred_%s.csv' % run_name_acc)
print(submission_csv_file)
submission_csv = pd.DataFrame({ 'click_id': click_ids , 'is_attributed': y_test_proba })
submission_csv.to_csv(submission_csv_file, index = False)
print('Time cost: %.2f s' % (time.time() - t0))
print('random_num: ', random_num)
print('date: ', date)
print(run_name_acc)
print('Done!')
```
| github_jupyter |
```
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
```
<div style="padding: 6px 12px 18px 12px; background: #eeffee; border: 2px solid #88aa88; border-radius: 4px;">
<h2>Preface: Installing Devito (do not include in manuscipt)</h2>
<p>This tutorial and the coming second part are based on Devito version 3.1.0. It requires the installation of the full software with examples, not only the code generation API. To install:</p>
<pre style="background: #eeffee;">
git clone -b v3.1.0 https://github.com/opesci/devito
cd devito
conda env create -f environment.yml
source activate devito
pip install -e .
</pre>
<p>That final dot is important, don't miss it out!</p>
<h3>Useful links</h3>
<ul>
<li><a href="http://www.opesci.org/">Devito documentation</a></li>
<li><a href="https://github.com/opesci/Devito">Devito source code and examples</a></li>
<li><a href="https://github.com/opesci/Devito/examples/seismic/tutorials">Tutorial notebooks with latest Devito/master</a></li>
</ul>
</div>
### Geophysics tutorial
# Full-waveform inversion 2: adjoint modeling
Mathias Louboutin<sup>1</sup>\*, Philipp Witte<sup>1</sup>, Michael Lange<sup>2</sup>, Navjot Kukreja<sup>2</sup>, Fabio Luporini<sup>2</sup>, Gerard Gorman<sup>2</sup>, and Felix J. Herrmann<sup>1,3</sup>
<sup>1</sup> Seismic Laboratory for Imaging and Modeling (SLIM), The University of British Columbia
<sup>2</sup> Imperial College London, London, UK
<sup>3</sup> now at Georgia Institute of Technology, USA
Corresponding author: mloubout@eoas.ubc.ca
## Introduction
This tutorial is the second part of a three part tutorial series on full-waveform inversion (FWI), in which we provide a step by step walk through of setting up forward and adjoint wave equation solvers and an optimization framework for inversion. In Part 1 (Louboutin et al., 2017), we showed how to use [Devito](http://www.opesci.org/devito-public) to set up and solve acoustic wave equations with (impulsive) seismic sources and sample wavefields at the receiver locations to forward model shot records. In the second part of this tutorial series, we will discuss how to set up and solve adjoint wave equations with Devito and from that, how we can calculate gradients and function values of the FWI objective function.
The gradient of FWI is most commonly computed via the adjoint state method, by cross-correlating forward and adjoint wavefields and summing the contributions over all time steps (Plessix, 2006). Calculating the gradient for one source location consists of three steps:
* Solve the forward wave equation to create a shot record. The time varying wavefield must be stored for use in step 3; techniques such as subsampling can be used to reduce the storage requirements.
* Compute the data residual (or misfit) between the predicted and observed data.
* Solve the corresponding discrete adjoint model using the data residual as the source. Within the adjoint (reverse) time loop, cross correlate the second time derivative of the adjoint wavefield with the forward wavefield. These cross correlations are summed to form the gradient.
We start with the definition and derivation of the adjoint wave equation and its Devito stencil and then show how to compute the gradient of the conventional least squares FWI misfit function. As usual, this tutorial is accompanied by all the code you need to reproduce the figures. Go to [github.com/seg/tutorials-2018](https://github.com/seg/tutorials-2018) and follow the links.
## A simple experiment
To demonstrate the gradient computation in the simplest possible way, we perform a small seismic transmission experiment with a circular imaging phantom, i.e. a constant velocity model with a circular high velocity inclusion in its centre, as shown in Figure 1. For a transmission experiment, we place 21 seismic sources on the left-hand side of the model and 101 receivers on the right-hand side.
We will use the forward propagator from part 1 to independently model the 21 "observed" shot records using the true model. As the initial model for our gradient calculation, we use a constant velocity model with the same velocity as the true model, but without the circular velocity perturbation. We will then model the 21 predicted shot records for the initial model, calculate the data residual and gradient for each shot, and sum them to obtain the full gradient.
> _++ Figure 1 is generated later in the manuscript ++_
>
> **Figure 1:** (a) The velocity model, with sources and receivers arranged vertically. (b) The initial estimate. (c) The difference between the model and the initial estimate.
## The adjoint wave equation
Adjoint wave equations are a main component in seismic inversion algorithms and are required for computing gradients of both linear and non-linear objective functions. To ensure stability of the adjoint modeling scheme and the expected convergence of inversion algorithms, it is very important that the adjoint wave equation is in fact the adjoint (transpose) of the forward wave equation. The derivation of the adjoint wave equation in the acoustic case is simple, as it is self-adjoint if we ignore the absorbing boundaries for the moment. However, in the general case, discrete wave equations do not have this property (such as the coupled anisotropic TTI wave equation (Zhang et al., 2011)) and require correct derivations of their adjoints. We concentrate here, as in part 1, on the acoustic case and follow an optimize-discretize approach, which means we write out the adjoint wave equation for the continuous case first and then discretize it, using finite difference operators of the same order as for the forward equation. With the variables defined as in part 1 and the data residual $\delta d(x,y,t; x_r, y_r)$, located at $x_r, y_r$ (receiver locations) as the adjoint source, the continuous adjoint wave equation is given by:
$$
m(x,y) \frac{\mathrm{d}^2 v(t,x,y)}{\mathrm{d}t^2}\ -\ \Delta v(t,x,y)\ -\ \eta(x,y) \frac{\mathrm{d} v(t,x,y)}{\mathrm{d}t}\ \ =\ \ \delta d(t,x,y;x_r, y_r)
$$
The adjoint acoustic wave equation is equivalent to the forward equation with the exception of the damping term $\eta(x,y) \mathrm{d}v(t,x,y)/\mathrm{d}t$, which contains a first time derivative and therefore has a change of sign in its adjoint. (A second derivative matrix is the same as its transpose, whereas a first derivative matrix is equal to its negative transpose and vice versa.)
Following the pattern of part 1, we first define the discrete adjoint wavefield $\mathbf{v}$ as a Devito `TimeFunction` object. For reasons we'll explain later, we do not need to save the adjoint wavefield:
```
# NOT FOR MANUSCRIPT
from examples.seismic import Model, demo_model, plot_velocity, plot_perturbation
shape = (101, 101) # Number of grid point (nx, nz)
spacing = (10., 10.) # Grid spacing in m. The domain size is now 1km by 1km
origin = (0., 0.) # Need origin to define relative source and receiver locations
model = demo_model('circle-isotropic',
vp=3.0,
vp_background=2.5,
origin=origin,
shape=shape,
spacing=spacing,
nbpml=40)
# For the manuscript, we'll re-form the model using the Vp field from
# this newly created model.
vp = model.vp
model0 = demo_model('circle-isotropic',
vp=2.5,
vp_background=2.5,
origin=origin,
shape=shape,
spacing=spacing,
nbpml=40)
t0 = 0. # Simulation starts a t=0
tn = 1000. # Simulation last 1 second (1000 ms)
dt = model.critical_dt # Time step from model grid spacing
nt = int(1 + (tn-t0) / dt) # Discrete time axis length
time = np.linspace(t0, tn, nt) # Discrete modelling time
# NOT FOR MANUSCRIPT
from devito import TimeFunction
v = TimeFunction(name="v", grid=model.grid,
time_order=2, space_order=4,
save=False)
```
Now symbolically set up the PDE:
```
pde = model.m * v.dt2 - v.laplace - model.damp * v.dt
```
As before, we then define a stencil:
```
# NOT FOR MANUSCRIPT
from devito import Eq
from sympy import solve
stencil_v = Eq(v.backward, solve(pde, v.backward)[0])
```
Just as for the forward wave equation, `stencil_v` defines the update for the adjoint wavefield of a single time step. The only difference is that, while the forward modeling propagator goes forward in time, the adjoint propagator goes backwards in time, since the initial time conditions for the forward propagator turn into final time conditions for the adjoint propagator. As for the forward stencil, we can write out the corresponding discrete expression for the update of the adjoint wavefield:
$$
\mathbf{v}[\text{time}-\text{dt}] = 2\mathbf{v}[\text{time}] - \mathbf{v}[\text{time}+\text{dt}] + \frac{\text{dt}^2}{\mathbf{m}}\Delta \mathbf{v}[\text{time}], \quad \text{time} = n_{t-1} \cdots 1
$$
with $\text{dt}$ being the time stepping interval. Once again, this expression does not contain any (adjoint) source terms so far, which will be defined as a separate `SparseFunction` object. Since the source term for the adjoint wave equation is the difference between an observed and modeled shot record, we first define an (empty) shot record `residual` with ``101`` receivers and coordinates defined in `rec_coords`. We then set the data field `rec.data` of our shot record to be the data residual between the observed data `d_obs` and the predicted data `d_pred`. The symbolic residual source expression `res_term` for our adjoint wave equation is then obtained by *injecting* the data residual into the modeling scheme (`residual.inject`). Since we solve the time-stepping loop backwards in time, the `res_term` is used to update the previous adjoint wavefield `v.backward`, rather than the next wavefield. As in the forward modeling example, the source is scaled by $\mathrm{dt}^2/\mathbf{m}$. In Python, we have:
```
# NOT FOR MANUSCRIPT
from examples.seismic import Receiver
nshots = 21 # Number of shots to create gradient from
nreceivers = 101 # Number of receiver locations per shot
# Recs are distributed across model, at depth of 20 m.
z_extent, _ = model.domain_size
z_locations = np.linspace(0, z_extent, num=nreceivers)
rec_coords = np.array([(980, z) for z in z_locations])
# NOT FOR MANUSCRIPT
from examples.seismic import PointSource
residual = PointSource(name='residual', ntime=nt,
grid=model.grid, coordinates=rec_coords)
res_term = residual.inject(field=v.backward,
expr=residual * dt**2 / model.m,
offset=model.nbpml)
# NOT FOR MANUSCRIPT
rec = Receiver(name='rec', npoint=nreceivers, ntime=nt,
grid=model.grid, coordinates=rec_coords)
# NOT FOR MANUSCRIPT
from examples.seismic import RickerSource
# At first, we want only a single shot.
# Src is 5% across model, at depth of 500 m.
z_locations = np.linspace(0, z_extent, num=nshots)
src_coords = np.array([(z_extent/50, z) for z in z_locations])
# NOT FOR MANUSCRIPT
f0 = 0.010 # kHz, peak frequency.
src = RickerSource(name='src', grid=model.grid, f0=f0,
time=time, coordinates=src_coords[nshots//2])
# NOT FOR MANUSCRIPT
plt.plot(src.time, src.data)
plt.xlabel("Time (ms)")
plt.ylabel("Amplitude")
plt.show()
# NOT FOR MANUSCRIPT
# Generates Figure 1
from matplotlib.gridspec import GridSpec
# Set up figure, grid, and parameters.
fig = plt.figure(figsize=(8, 12))
gs = GridSpec(3, 2)
ax0 = fig.add_subplot(gs[:2, :2])
ax1 = fig.add_subplot(gs[2, 0])
ax2 = fig.add_subplot(gs[2, 1])
extent = [model.origin[0], model.origin[0] + 1e-3 * model.shape[0] * model.spacing[0],
model.origin[1] + 1e-3*model.shape[1] * model.spacing[1], model.origin[1]]
model_param = dict(vmin=2.5, vmax=3.0, cmap="GnBu", aspect=1, extent=extent)
diff_param = dict(vmin=-1, vmax=0, cmap="GnBu", aspect=1, extent=extent)
# Part (a)
im = ax0.imshow(np.transpose(model.vp), **model_param)
ax0.scatter(*rec_coords.T/1000, lw=0, c='green', s=8)
ax0.scatter(*src_coords.T/1000, lw=0, c='red',s=24)
ax0.set_ylabel('Depth (km)', fontsize=14)
ax0.text(0.5, 0.08, "model.vp", ha="center", color='k', size=18)
ax0.text(0.5, 0.5, "3000 m/s", ha="center", va='center', color='w', size=14)
ax0.text(0.8, 0.5, "2500 m/s", ha="center", va='center', color='navy', size=14)
ax0.text(0.04, 0.04, "sources", ha="left", color='r', size=12)
ax0.text(0.96, 0.04, "receivers", ha="right", color='green', size=12)
ax0.set_ylim(1, 0)
plt.setp(ax0.get_xticklabels(), fontsize=12)
plt.setp(ax0.get_yticklabels(), fontsize=12)
# Part (b)
im = ax1.imshow(np.transpose(model0.vp), **model_param)
ax1.set_xlabel('X position (km)', fontsize=14)
ax1.set_yticklabels([0, '', 0.5, '', 1])
ax1.text(0.5, 0.08, "model0.vp", ha="center", color='k', size=14)
ax1.set_ylabel('Depth (km)', fontsize=14)
plt.setp(ax1.get_xticklabels(), fontsize=12)
plt.setp(ax1.get_yticklabels(), fontsize=12)
# Part (c)
im = ax2.imshow(np.transpose(model0.vp - model.vp), **diff_param)
ax2.set_xlabel('X position (km)', fontsize=14)
ax2.set_yticklabels([])
ax2.text(0.5, 0.08, "model0.vp – model.vp", ha="center", color='w', size=14)
ax2.text(0.5, 0.5, "–500 m/s", ha="center", va='center', color='w', size=12)
ax2.text(0.85, 0.5, "0 m/s", ha="center", va='center', color='w', size=12)
plt.setp(ax2.get_xticklabels(), fontsize=12)
plt.setp(ax2.get_yticklabels(), fontsize=12)
plt.savefig("../Figures/Figure_1.pdf")
plt.savefig("../Figures/Figure_1.png", dpi=400)
plt.show()
```
In this demonstration, there is no real data. Instead we will generate the 'observed' data via forward modeling with the true model `model`. The synthetic data is generated from the initial model `model0`. The resulting data, and their difference, are shown in Figure 2.
```
# NOT FOR MANUSCRIPT
from examples.seismic.acoustic import AcousticWaveSolver
solver = AcousticWaveSolver(model, src, rec, space_order=4)
# Compute 'real' data with forward operator.
obs , _, _ = solver.forward(src=src, m=model.m)
# NOT FOR MANUSCRIPT
# Compute initial data with forward operator.
pred, u0, _ = solver.forward(src=src, m=model0.m, save=True)
# NOT FOR MANUSCRIPT
fig = plt.figure(figsize=(15, 5))
extent = [model.origin[0], # Horizontal min
model.origin[0] + 1e-3 * model.shape[0] * model.spacing[0],
tn/1000, # Vertical min (bottom)
t0/1000] # Vertical max (top)
ma = np.percentile(obs.data, 99.5)
params = dict(vmin=-ma, vmax=ma, cmap="Greys", aspect=1, extent=extent)
text_params = dict(ha="center", color='w', size=16)
ax0 = fig.add_subplot(131)
im = plt.imshow(obs.data, **params)
ax0.set_ylabel('Time (s)', fontsize=16)
ax0.text(0.5, 0.08, "obs", **text_params)
plt.xticks(fontsize=10)
plt.yticks(fontsize=10)
ax1 = fig.add_subplot(132)
im = plt.imshow(pred.data, **params)
ax1.set_xlabel('Z position (km)', fontsize=16)
ax1.set_yticklabels([])
ax1.text(0.5, 0.08, "pred", **text_params)
plt.xticks(fontsize=10)
plt.yticks(fontsize=10)
ax2 = fig.add_subplot(133)
im = plt.imshow(pred.data - obs.data, **params)
ax2.set_yticklabels([])
ax2.text(0.5, 0.08, "pred – obs", **text_params)
plt.xticks(fontsize=10)
plt.yticks(fontsize=10)
plt.savefig("../Figures/Figure2.pdf")
plt.savefig("../Figures/Figure2.png", dpi=400)
plt.show()
```
> **Figure 2.** Shot records for a shot at a Z position of 0.5 km. (a) The observed data, using the known model with the high velocity disc, contains a perturbation not present in (b) the 'predicted' data, using the initial estimate of the model, which contais no disc. (c) The residual.
Finally, we create the full propagator by adding the residual source expression to our previously defined stencil and set the flag `time_axis=Backward`, to specify that the propagator runs in backwards in time:
```
# NOT FOR MANUSCRIPT
from devito import Operator, Backward
op_adj = Operator([stencil_v] + res_term, time_axis=Backward)
```
In contrast to forward modeling, we do not record any measurements at the surface since we are only interested in the adjoint wavefield itself. The full script for setting up the adjoint wave equation, including an animation of the adjoint wavefield is available in **`adjoint_modeling.ipynb`**.
## Computing the FWI gradient
The goal of FWI is to estimate a discrete parametrization of the subsurface by minimizing the misfit between the observed shot records of a seismic survey and numerically modeled shot records. The predicted shot records are obtained by solving an individual wave equation per shot location and depend on the parametrization $\mathbf{m}$ of our wave propagator. The most common function for measuring the data misfit between the observed and modeled data is the $\ell_2$ norm, which leads to the following objective function (Lions (1971), Tarantola (1984)):
$$
\mathop{\hbox{minimize}}_{\mathbf{m}} \hspace{.2cm} f(\mathbf{m})= \sum_{i=1}^{n_\mathrm{s}} \frac{1}{2} \left\lVert \mathbf{d}^\mathrm{pred}_i (\mathbf{m}, \mathbf{q}_i) - \mathbf{d}_i^\mathrm{obs} \right\rVert_2^2,
$$
where the index $i$ runs over the total number of shots $n_\mathrm{s}$ and the model parameters are the squared slowness. Optimization problems of this form are called nonlinear least-squares problems, since the predicted data modeled with the forward modeling propagator (`op_fwd()` in part 1) depends nonlinearly on the unknown parameters $\mathbf{m}$. The full derivation of the FWI gradient using the adjoint state method is outside the scope of this tutorial, but conceptually we obtain the gradient by applying the chain rule and taking the partial derivative of the inverse wave equation $\mathbf{A}(\mathbf{m})^{-1}$ with respect to $\mathbf{m}$, which yields the following expression (Plessix, 2006, Virieux and Operto, 2009):
$$
\nabla f (\mathbf{m})= - \sum_{i=1}^{n_\mathrm{s}} \sum_{\text{time}=1}^{n_t} \mathbf{u}[\text{time}]\odot \ddot{\mathbf{v}}[\text{time}].
$$
The inner sum $\text{time}=1,...,n_t$ runs over the number of computational time steps $n_t$ and $\ddot{\mathbf{v}}$ denotes the second temporal derivative of the adjoint wavefield $\mathbf{v}$. Computing the gradient of Equation 3, therefore corresponds to performing the point-wise multiplication (denoted by the symbol $\odot$) of the forward wavefields with the second time derivative of the adjoint wavefield and summing over all time steps.
To avoid the need to store the adjoint wavefield, the FWI gradient is calculated in the reverse time-loop while solving the adjoint wave equation. To compute the gradient $\mathbf{g}$ for the current time step $\mathbf{v}[\text{time}]$:
$$
\mathbf{g} = \mathbf{g} - \frac{\mathbf{v}[\text{time-dt}] - 2\mathbf{v}[\text{time}] + \mathbf{v}[\text{time+dt}]}{\mathrm{dt}^2} \odot \mathbf{u}[\text{time}], \quad \text{time}=1 \cdots n_{t-1}
$$
The second time derivative of the adjoint wavefield is computed with a second order finite-difference stencil and uses the three adjoint wavefields that are kept in memory during the adjoint time loop (Equation 2).
In Devito we define the gradient as a `Function` since the gradient is computed as the sum over all time steps and therefore has no time dependence:
```
# NOT FOR MANUSCRIPT
from devito import TimeFunction, Function
# NOT FOR MANUSCRIPT
# This is the same u as in Part 1.
u = TimeFunction(name="u", grid=model.grid,
time_order=2, space_order=4,
save=True, time_dim=nt)
grad = Function(name="grad", grid=model.grid)
```
The update for the gradient as defined in Equations 4 and 5 is then:
```
grad_update = Eq(grad, grad - u * v.dt2)
```
Now we must add the gradient update expression to the adjoint propagator `op_grad`. This yields a single symbolic expression with update instructions for both the adjoint wavefield and the gradient:
```
op_grad = Operator([stencil_v] + res_term + [grad_update],
time_axis=Backward)
```
Solving the adjoint wave equation by running the following now computes the FWI gradient for a single source. Its value is stored in `grad.data`.
```
op_grad(u=u0, v=v, m=model0.m,
residual=pred.data-obs.data,
time=nt, dt=dt)
# NOT FOR MANUSCIPT
plt.figure(figsize=(8,8))
plt.imshow(np.transpose(grad.data[40:-40,40:-40]), extent=extent, cmap='RdBu', vmin=-1e3, vmax=1e3)
plt.colorbar(shrink=0.75)
plt.show()
# NOT FOR MANUSCRIPT
tmp = grad.copy()
```
Now we can iterate over all the shot locations, running the same sequence of commands each time.
```
# NOT FOR MANUSCRIPT
from devito import configuration
configuration['log_level'] = 'WARNING'
# Create the symbols.
u0 = TimeFunction(name='u0', grid=model.grid, time_order=2, space_order=4, save=True, time_dim=nt)
u = TimeFunction(name='u', grid=model.grid, time_order=2, space_order=4, save=True, time_dim=nt)
v = TimeFunction(name='v', grid=model.grid, time_order=2, space_order=4, save=False)
# Define the wave equation, but with a negated damping term
eqn = model.m * v.dt2 - v.laplace - model.damp * v.dt
# Use SymPy to rearrange the equation into a stencil expression.
stencil = Eq(v.backward, solve(eqn, v.backward)[0])
# Define the residual injection.
residual = PointSource(name='residual', ntime=nt, coordinates=rec_coords, grid=model.grid)
res_term = residual.inject(field=v.backward, expr=residual * dt**2 / model.m, offset=model.nbpml)
# Correlate u and v for the current time step and add it to the gradient.
grad = Function(name='grad', grid=model.grid, dtype=model.m.dtype)
grad_update = Eq(grad, grad - u * v.dt2)
# Compose the operator.
op_grad2 = Operator([stencil] + res_term + [grad_update], time_axis=Backward)
# Iterate over the shots.
for i in range(nshots):
print("Source {} of {}".format(i, nshots))
# Opdate source location.
src.coordinates.data[0, :] = src_coords[i]
# Generate data from true model and current model estimate.
obs, _, _ = solver.forward(src=src, m=model.m)
pred, _, _ = solver.forward(src=src, m=model0.m, u=u0, save=True)
# Compute the gradient from the residual.
v = TimeFunction(name='v', grid=model.grid, time_order=2, space_order=4)
residual.data[:] = pred.data - obs.data
op_grad2(u=u0, v=v, m=model0.m, residual=residual, grad=grad, dt=dt)
configuration['log_level'] = 'INFO'
# NOT FOR MANUSCRIPT
plt.figure(figsize=(8,8))
plt.imshow(np.transpose(grad.data)[40:-40, 40:-40], extent=extent, vmin=-1e4, vmax=1e4, cmap='RdBu')
plt.colorbar(shrink=0.75)
plt.show()
# NOT FOR MANUSCRIPT
# Generates Figure 3
from plot_utils import add_subplot_axes
fig = plt.figure(figsize=(12,6))
ax0 = fig.add_subplot(121)
ma = 1e3
im = ax0.imshow(np.transpose(tmp.data[40:-40,40:-40]), extent=extent, cmap='RdBu', vmin=-ma, vmax=ma)
cax = add_subplot_axes(ax0, [0.3, 0.075, 0.4, 0.02])
fig.colorbar(im, cax=cax, orientation='horizontal')
cax.text(0, 1, "{:.0f} ".format(-ma), ha='right', va='top', size=10)
cax.text(1, 1, " {:.0f}".format(ma), ha='left', va='top', size=10)
cax.set_axis_off()
ax0.text(0.5, 0.05, "Shot @ 500 m", ha='center', va='center', size=16)
ax0.set_xlabel('X position (m)', fontsize=16)
ax0.set_ylabel('Z position (m)', fontsize=16)
plt.setp(ax0.get_xticklabels(), fontsize=12)
plt.setp(ax0.get_yticklabels(), fontsize=12)
ax1 = fig.add_subplot(122)
ma = 1e4
im = ax1.imshow(np.transpose(grad.data[40:-40,40:-40]), extent=extent, cmap='RdBu', vmin=-ma, vmax=ma)
cax = add_subplot_axes(ax1, [0.3, 0.075, 0.4, 0.02])
fig.colorbar(im, cax=cax, orientation='horizontal')
cax.text(0, 1, "{:.0f} ".format(-ma), ha='right', va='top', size=10)
cax.text(1, 1, " {:.0f}".format(ma), ha='left', va='top', size=10)
cax.set_axis_off()
ax1.text(0.5, 0.05, "All {} shots".format(nshots), ha='center', va='center', size=16)
ax1.set_xlabel('X position (m)', fontsize=16)
ax1.set_yticklabels([])
plt.setp(ax1.get_xticklabels(), fontsize=12)
plt.setp(ax1.get_yticklabels(), fontsize=12)
plt.savefig("../Figures/Figure3.pdf")
plt.savefig("../Figures/Figure3.png", dpi=400)
plt.show()
```
> **Figure 3.** Gradient plots for (a) a single shot at 0.5 km and (b) the sum of all shots.
This gradient can then be used for a simple gradient descent optimization loop, as illustrated at the end of the notebook `adjoint_modeling.ipynb`. After each update, a new gradient is computed for the new velocity model until sufficient decrease of the objective or chosen number of iteration is reached. A detailed treatment of optimization and more advanced algorithms will be described in the third and final part of this tutorial series.
<div style="padding: 6px 12px 18px 12px; background: #eeffee; border: 2px solid #88aa88; border-radius: 4px;">
<h3>Verification (do not include in manuscipt)</h3>
<p>The next step of the adjoint modeling and gradient part is verification with unit testing, i.e. we ensure that the adjoints and gradients are implemented correctly. Incorrect adjoints can lead to unpredictable behaviour during and inversion and in the worst case cause slower convergence or convergence to wrong solutions.</p>
<p>Since our forward-adjoint wave equation solvers correspond to forward-adjoint pairs, we need to ensure that the adjoint defined dot test holds within machine precision (see **`tests/test_adjointA.py`** for the dot test). Furthermore, we verify the correct implementation of the FWI gradient by ensuring that using the gradient leads to first order convergence. The gradient test can be found in **`tests/test_gradient.py`**.</p>
</div>
## Conclusions
We need the gradient of the FWI objective function in order to find the optimal solution. It is computed by solving adjoint wave equations and summing the point-wise product of forward and adjoint wavefields over all time steps. Using Devito, the adjoint wave equation is set up in a similar fashion as the forward wave equation, with the main difference being the (adjoint) source, which is the residual between the observed and predicted shot records.
With the ability to model shot records and compute gradients of the FWI objective function, we are ready to demonstrate how to set up more gradient-based algorithms for FWI in Part 3 next month.
## Acknowledgments
This research was carried out as part of the SINBAD II project with the support of the member organizations of the SINBAD Consortium. This work was financially supported in part by EPSRC grant EP/L000407/1 and the Imperial College London Intel Parallel Computing Centre.
## References
[1] Michael Lange, Navjot Kukreja, Mathias Louboutin, Fabio Luporini, Felippe Vieira Zacarias, Vincenzo Pandolfo, Paulius Velesko, Paulius Kazakas, and Gerard Gorman. Devito: Towards a generic finite difference DSL using symbolic python. In 6th Workshop on Python for High-Performance and Scientific Computing, pages 67–75, 11 2016. doi: 10.1109/PyHPC.2016.9.
[2] J. L. Lions. Optimal control of systems governed by partial differential equations. Springer-Verlag Berlin Heidelberg, 1st edition, 1971. ISBN 978-3-642-65026-0.
[3] Mathias Louboutin, Philipp A. Witte, Michael Lange, Navjot Kukreja, Fabio Luporini, Gerard Gorman, and Felix J. Herrmann. Full-waveform inversion - part 1: forward modeling. Submitted to The Leading Edge for the tutorial section on October 30, 2017., 2017.
[4] Aaron Meurer, Christopher P. Smith, Mateusz Paprocki, Ondřej Čertík, Sergey B. Kirpichev, Matthew Rocklin, AMiT Kumar, Sergiu Ivanov, Jason K. Moore, Sartaj Singh, Thilina Rathnayake, Sean Vig, Brian E. Granger, Richard P. Muller, Francesco Bonazzi, Harsh Gupta, Shivam Vats, Fredrik Johansson, Fabian Pedregosa, Matthew J. Curry, Andy R. Terrel, Štěpán Roučka, Ashutosh Saboo, Isuru Fernando, Sumith Kulal, Robert Cimrman, and Anthony Scopatz. Sympy: symbolic computing in python. Peer J Computer Science, 3:e103, January 2017. ISSN 2376-5992. doi: 10.7717/peerj-cs.103. URL https: //doi.org/10.7717/peerj-cs.103.
[5] R.-E. Plessix. A review of the adjoint-state method for computing the gradient of a functional with geophysical applications. Geophysical Journal International, 167(2):495, 2006. doi: 10.1111/j.1365-246X.2006.02978.x. URL +http://dx.doi.org/10.1111/j.1365-246X.2006.02978.x
[6] Albert Tarantola. Inversion of seismic reflection data in the acoustic approximation. GEOPHYSICS, 49(8): 1259–1266, 1984. doi: 10.1190/1.1441754. URL https://doi.org/10.1190/1.1441754
[7] J. Virieux and S. Operto. An overview of full-waveform inversion in exploration geophysics. GEOPHYSICS, 74 (5):WCC1–WCC26, 2009. doi: 10.1190/1.3238367. URL http://library.seg.org/doi/abs/10.1190/1.3238367
[8] Yu Zhang, Houzhu Zhang, and Guanquan Zhang. A stable tti reverse time migration and its implementation. GEOPHYSICS, 76(3):WA3–WA11, 2011. doi: 10.1190/1.3554411. URL https://doi.org/10.1190/1.3554411.
<hr>
© 2017 The authors — licensed CC-BY-SA
| github_jupyter |
#### New to Plotly?
Plotly's Python library is free and open source! [Get started](https://plot.ly/python/getting-started/) by downloading the client and [reading the primer](https://plot.ly/python/getting-started/).
<br>You can set up Plotly to work in [online](https://plot.ly/python/getting-started/#initialization-for-online-plotting) or [offline](https://plot.ly/python/getting-started/#initialization-for-offline-plotting) mode, or in [jupyter notebooks](https://plot.ly/python/getting-started/#start-plotting-online).
<br>We also have a quick-reference [cheatsheet](https://images.plot.ly/plotly-documentation/images/python_cheat_sheet.pdf) (new!) to help you get started!
#### Version Check
Note: 2D Histograms are available in version <b>1.9.12+</b><br>
Run `pip install plotly --upgrade` to update your Plotly version
```
import plotly
plotly.__version__
```
### 2D Histogram with Slider Control ###
Add slider controls to 2d histograms with the [postMessage API](https://github.com/plotly/postMessage-API).
See the [code on JSFiddle](https://jsfiddle.net/plotlygraphs/y9sdy76h/4/).
Watch [the 5 second video](https://raw.githubusercontent.com/plotly/documentation/gh-pages/all_static/images
/flight_conflicts.gif) of how it works.
```
from IPython.core.display import display,HTML
display(HTML('<iframe height=600 width=950 src="https://jsfiddle.net/plotlygraphs/y9sdy76h/4/embedded/result,js,html/"></iframe>'))
```
### 2D Histogram of a Bivariate Normal Distribution ###
```
import plotly.plotly as py
import plotly.graph_objs as go
import numpy as np
x = np.random.randn(500)
y = np.random.randn(500)+1
data = [
go.Histogram2d(
x=x,
y=y
)
]
py.iplot(data)
```
### 2D Histogram Binning and Styling Options ###
```
import plotly.plotly as py
import plotly.graph_objs as go
import numpy as np
x = np.random.randn(500)
y = np.random.randn(500)+1
data = [
go.Histogram2d(x=x, y=y, histnorm='probability',
autobinx=False,
xbins=dict(start=-3, end=3, size=0.1),
autobiny=False,
ybins=dict(start=-2.5, end=4, size=0.1),
colorscale=[[0, 'rgb(12,51,131)'], [0.25, 'rgb(10,136,186)'], [0.5, 'rgb(242,211,56)'], [0.75, 'rgb(242,143,56)'], [1, 'rgb(217,30,30)']]
)
]
py.iplot(data)
```
### 2D Histogram Overlaid with a Scatter Chart ###
```
import plotly.plotly as py
import plotly.graph_objs as go
import numpy as np
x0 = np.random.randn(100)/5. + 0.5 # 5. enforces float division
y0 = np.random.randn(100)/5. + 0.5
x1 = np.random.rand(50)
y1 = np.random.rand(50) + 1.0
x = np.concatenate([x0, x1])
y = np.concatenate([y0, y1])
trace1 = go.Scatter(
x=x0,
y=y0,
mode='markers',
showlegend=False,
marker=dict(
symbol='x',
opacity=0.7,
color='white',
size=8,
line=dict(width=1),
)
)
trace2 = go.Scatter(
x=x1,
y=y1,
mode='markers',
showlegend=False,
marker=dict(
symbol='circle',
opacity=0.7,
color='white',
size=8,
line=dict(width=1),
)
)
trace3 = go.Histogram2d(
x=x,
y=y,
colorscale='YlGnBu',
zmax=10,
nbinsx=14,
nbinsy=14,
zauto=False,
)
layout = go.Layout(
xaxis=dict( ticks='', showgrid=False, zeroline=False, nticks=20 ),
yaxis=dict( ticks='', showgrid=False, zeroline=False, nticks=20 ),
autosize=False,
height=550,
width=550,
hovermode='closest',
)
data = [trace1, trace2, trace3]
fig = go.Figure(data=data, layout=layout)
py.iplot(fig)
```
#### Reference
See https://plot.ly/python/reference/#histogram2d for more information and chart attribute options!
```
from IPython.display import display, HTML
display(HTML('<link href="//fonts.googleapis.com/css?family=Open+Sans:600,400,300,200|Inconsolata|Ubuntu+Mono:400,700rel="stylesheet" type="text/css" />'))
display(HTML('<link rel="stylesheet" type="text/csshref="http://help.plot.ly/documentation/all_static/css/ipython-notebook-custom.css">'))
! pip install git+https://github.com/plotly/publisher.git --upgrade
import publisher
publisher.publish(
'2d-histograms.ipynb', 'python/2D-Histogram/', 'Python 2D Histograms | plotly',
'How to make 2D Histograms in Python with Plotly.',
title = 'Python 2D Histograms | plotly',
name = '2D Histograms',
has_thumbnail='true', thumbnail='thumbnail/histogram2d.jpg',
language='python', display_as='statistical', order=6,
ipynb= '~notebook_demo/24')
```
| github_jupyter |
Copyright (c) 2015-2017 [Sebastian Raschka](sebastianraschka.com)
https://github.com/rasbt/python-machine-learning-book
[MIT License](https://github.com/rasbt/python-machine-learning-book/blob/master/LICENSE.txt)
# Python Machine Learning - Code Examples
# Chapter 11 - Working with Unlabeled Data – Clustering Analysis
Note that the optional watermark extension is a small IPython notebook plugin that I developed to make the code reproducible. You can just skip the following line(s).
```
%load_ext watermark
%watermark -a 'Sebastian Raschka' -u -d -v -p numpy,pandas,matplotlib,scipy,sklearn
```
*The use of `watermark` is optional. You can install this IPython extension via "`pip install watermark`". For more information, please see: https://github.com/rasbt/watermark.*
<br>
<br>
### Overview
- [Grouping objects by similarity using k-means](#Grouping-objects-by-similarity-using-k-means)
- [K-means++](#K-means++)
- [Hard versus soft clustering](#Hard-versus-soft-clustering)
- [Using the elbow method to find the optimal number of clusters](#Using-the-elbow-method-to-find-the-optimal-number-of-clusters)
- [Quantifying the quality of clustering via silhouette plots](#Quantifying-the-quality-of-clustering-via-silhouette-plots)
- [Organizing clusters as a hierarchical tree](#Organizing-clusters-as-a-hierarchical-tree)
- [Performing hierarchical clustering on a distance matrix](#Performing-hierarchical-clustering-on-a-distance-matrix)
- [Attaching dendrograms to a heat map](#Attaching-dendrograms-to-a-heat-map)
- [Applying agglomerative clustering via scikit-learn](#Applying-agglomerative-clustering-via-scikit-learn)
- [Locating regions of high density via DBSCAN](#Locating-regions-of-high-density-via-DBSCAN)
- [Summary](#Summary)
<br>
<br>
```
from IPython.display import Image
%matplotlib inline
```
# Grouping objects by similarity using k-means
```
from sklearn.datasets import make_blobs
X, y = make_blobs(n_samples=150,
n_features=2,
centers=3,
cluster_std=0.5,
shuffle=True,
random_state=0)
import matplotlib.pyplot as plt
plt.scatter(X[:, 0], X[:, 1], c='white', marker='o', s=50)
plt.grid()
plt.tight_layout()
#plt.savefig('./figures/spheres.png', dpi=300)
plt.show()
from sklearn.cluster import KMeans
km = KMeans(n_clusters=3,
init='random',
n_init=10,
max_iter=300,
tol=1e-04,
random_state=0)
y_km = km.fit_predict(X)
plt.scatter(X[y_km == 0, 0],
X[y_km == 0, 1],
s=50,
c='lightgreen',
marker='s',
label='cluster 1')
plt.scatter(X[y_km == 1, 0],
X[y_km == 1, 1],
s=50,
c='orange',
marker='o',
label='cluster 2')
plt.scatter(X[y_km == 2, 0],
X[y_km == 2, 1],
s=50,
c='lightblue',
marker='v',
label='cluster 3')
plt.scatter(km.cluster_centers_[:, 0],
km.cluster_centers_[:, 1],
s=250,
marker='*',
c='red',
label='centroids')
plt.legend()
plt.grid()
plt.tight_layout()
#plt.savefig('./figures/centroids.png', dpi=300)
plt.show()
```
<br>
## K-means++
...
## Hard versus soft clustering
...
## Using the elbow method to find the optimal number of clusters
```
print('Distortion: %.2f' % km.inertia_)
distortions = []
for i in range(1, 11):
km = KMeans(n_clusters=i,
init='k-means++',
n_init=10,
max_iter=300,
random_state=0)
km.fit(X)
distortions.append(km.inertia_)
plt.plot(range(1, 11), distortions, marker='o')
plt.xlabel('Number of clusters')
plt.ylabel('Distortion')
plt.tight_layout()
#plt.savefig('./figures/elbow.png', dpi=300)
plt.show()
```
<br>
## Quantifying the quality of clustering via silhouette plots
```
import numpy as np
from matplotlib import cm
from sklearn.metrics import silhouette_samples
km = KMeans(n_clusters=3,
init='k-means++',
n_init=10,
max_iter=300,
tol=1e-04,
random_state=0)
y_km = km.fit_predict(X)
cluster_labels = np.unique(y_km)
n_clusters = cluster_labels.shape[0]
silhouette_vals = silhouette_samples(X, y_km, metric='euclidean')
y_ax_lower, y_ax_upper = 0, 0
yticks = []
for i, c in enumerate(cluster_labels):
c_silhouette_vals = silhouette_vals[y_km == c]
c_silhouette_vals.sort()
y_ax_upper += len(c_silhouette_vals)
color = cm.jet(float(i) / n_clusters)
plt.barh(range(y_ax_lower, y_ax_upper), c_silhouette_vals, height=1.0,
edgecolor='none', color=color)
yticks.append((y_ax_lower + y_ax_upper) / 2.)
y_ax_lower += len(c_silhouette_vals)
silhouette_avg = np.mean(silhouette_vals)
plt.axvline(silhouette_avg, color="red", linestyle="--")
plt.yticks(yticks, cluster_labels + 1)
plt.ylabel('Cluster')
plt.xlabel('Silhouette coefficient')
plt.tight_layout()
# plt.savefig('./figures/silhouette.png', dpi=300)
plt.show()
```
Comparison to "bad" clustering:
```
km = KMeans(n_clusters=2,
init='k-means++',
n_init=10,
max_iter=300,
tol=1e-04,
random_state=0)
y_km = km.fit_predict(X)
plt.scatter(X[y_km == 0, 0],
X[y_km == 0, 1],
s=50,
c='lightgreen',
marker='s',
label='cluster 1')
plt.scatter(X[y_km == 1, 0],
X[y_km == 1, 1],
s=50,
c='orange',
marker='o',
label='cluster 2')
plt.scatter(km.cluster_centers_[:, 0], km.cluster_centers_[:, 1],
s=250, marker='*', c='red', label='centroids')
plt.legend()
plt.grid()
plt.tight_layout()
#plt.savefig('./figures/centroids_bad.png', dpi=300)
plt.show()
cluster_labels = np.unique(y_km)
n_clusters = cluster_labels.shape[0]
silhouette_vals = silhouette_samples(X, y_km, metric='euclidean')
y_ax_lower, y_ax_upper = 0, 0
yticks = []
for i, c in enumerate(cluster_labels):
c_silhouette_vals = silhouette_vals[y_km == c]
c_silhouette_vals.sort()
y_ax_upper += len(c_silhouette_vals)
color = cm.jet(float(i) / n_clusters)
plt.barh(range(y_ax_lower, y_ax_upper), c_silhouette_vals, height=1.0,
edgecolor='none', color=color)
yticks.append((y_ax_lower + y_ax_upper) / 2.)
y_ax_lower += len(c_silhouette_vals)
silhouette_avg = np.mean(silhouette_vals)
plt.axvline(silhouette_avg, color="red", linestyle="--")
plt.yticks(yticks, cluster_labels + 1)
plt.ylabel('Cluster')
plt.xlabel('Silhouette coefficient')
plt.tight_layout()
# plt.savefig('./figures/silhouette_bad.png', dpi=300)
plt.show()
```
<br>
<br>
# Organizing clusters as a hierarchical tree
```
Image(filename='./images/11_05.png', width=400)
import pandas as pd
import numpy as np
np.random.seed(123)
variables = ['X', 'Y', 'Z']
labels = ['ID_0', 'ID_1', 'ID_2', 'ID_3', 'ID_4']
X = np.random.random_sample([5, 3])*10
df = pd.DataFrame(X, columns=variables, index=labels)
df
```
<br>
## Performing hierarchical clustering on a distance matrix
```
from scipy.spatial.distance import pdist, squareform
row_dist = pd.DataFrame(squareform(pdist(df, metric='euclidean')),
columns=labels,
index=labels)
row_dist
```
We can either pass a condensed distance matrix (upper triangular) from the `pdist` function, or we can pass the "original" data array and define the `metric='euclidean'` argument in `linkage`. However, we should not pass the squareform distance matrix, which would yield different distance values although the overall clustering could be the same.
```
# 1. incorrect approach: Squareform distance matrix
from scipy.cluster.hierarchy import linkage
row_clusters = linkage(row_dist, method='complete', metric='euclidean')
pd.DataFrame(row_clusters,
columns=['row label 1', 'row label 2',
'distance', 'no. of items in clust.'],
index=['cluster %d' % (i + 1)
for i in range(row_clusters.shape[0])])
# 2. correct approach: Condensed distance matrix
row_clusters = linkage(pdist(df, metric='euclidean'), method='complete')
pd.DataFrame(row_clusters,
columns=['row label 1', 'row label 2',
'distance', 'no. of items in clust.'],
index=['cluster %d' % (i + 1)
for i in range(row_clusters.shape[0])])
# 3. correct approach: Input sample matrix
row_clusters = linkage(df.values, method='complete', metric='euclidean')
pd.DataFrame(row_clusters,
columns=['row label 1', 'row label 2',
'distance', 'no. of items in clust.'],
index=['cluster %d' % (i + 1)
for i in range(row_clusters.shape[0])])
from scipy.cluster.hierarchy import dendrogram
# make dendrogram black (part 1/2)
# from scipy.cluster.hierarchy import set_link_color_palette
# set_link_color_palette(['black'])
row_dendr = dendrogram(row_clusters,
labels=labels,
# make dendrogram black (part 2/2)
# color_threshold=np.inf
)
plt.tight_layout()
plt.ylabel('Euclidean distance')
#plt.savefig('./figures/dendrogram.png', dpi=300,
# bbox_inches='tight')
plt.show()
```
<br>
## Attaching dendrograms to a heat map
```
# plot row dendrogram
fig = plt.figure(figsize=(8, 8), facecolor='white')
axd = fig.add_axes([0.09, 0.1, 0.2, 0.6])
# note: for matplotlib < v1.5.1, please use orientation='right'
row_dendr = dendrogram(row_clusters, orientation='left')
# reorder data with respect to clustering
df_rowclust = df.iloc[row_dendr['leaves'][::-1]]
axd.set_xticks([])
axd.set_yticks([])
# remove axes spines from dendrogram
for i in axd.spines.values():
i.set_visible(False)
# plot heatmap
axm = fig.add_axes([0.23, 0.1, 0.6, 0.6]) # x-pos, y-pos, width, height
cax = axm.matshow(df_rowclust, interpolation='nearest', cmap='hot_r')
fig.colorbar(cax)
axm.set_xticklabels([''] + list(df_rowclust.columns))
axm.set_yticklabels([''] + list(df_rowclust.index))
# plt.savefig('./figures/heatmap.png', dpi=300)
plt.show()
```
<br>
## Applying agglomerative clustering via scikit-learn
```
from sklearn.cluster import AgglomerativeClustering
ac = AgglomerativeClustering(n_clusters=2,
affinity='euclidean',
linkage='complete')
labels = ac.fit_predict(X)
print('Cluster labels: %s' % labels)
```
<br>
<br>
# Locating regions of high density via DBSCAN
```
Image(filename='./images/11_11.png', width=500)
from sklearn.datasets import make_moons
X, y = make_moons(n_samples=200, noise=0.05, random_state=0)
plt.scatter(X[:, 0], X[:, 1])
plt.tight_layout()
# plt.savefig('./figures/moons.png', dpi=300)
plt.show()
```
K-means and hierarchical clustering:
```
f, (ax1, ax2) = plt.subplots(1, 2, figsize=(8, 3))
km = KMeans(n_clusters=2, random_state=0)
y_km = km.fit_predict(X)
ax1.scatter(X[y_km == 0, 0], X[y_km == 0, 1],
c='lightblue', marker='o', s=40, label='cluster 1')
ax1.scatter(X[y_km == 1, 0], X[y_km == 1, 1],
c='red', marker='s', s=40, label='cluster 2')
ax1.set_title('K-means clustering')
ac = AgglomerativeClustering(n_clusters=2,
affinity='euclidean',
linkage='complete')
y_ac = ac.fit_predict(X)
ax2.scatter(X[y_ac == 0, 0], X[y_ac == 0, 1], c='lightblue',
marker='o', s=40, label='cluster 1')
ax2.scatter(X[y_ac == 1, 0], X[y_ac == 1, 1], c='red',
marker='s', s=40, label='cluster 2')
ax2.set_title('Agglomerative clustering')
plt.legend()
plt.tight_layout()
#plt.savefig('./figures/kmeans_and_ac.png', dpi=300)
plt.show()
```
Density-based clustering:
```
from sklearn.cluster import DBSCAN
db = DBSCAN(eps=0.2, min_samples=5, metric='euclidean')
y_db = db.fit_predict(X)
plt.scatter(X[y_db == 0, 0], X[y_db == 0, 1],
c='lightblue', marker='o', s=40,
label='cluster 1')
plt.scatter(X[y_db == 1, 0], X[y_db == 1, 1],
c='red', marker='s', s=40,
label='cluster 2')
plt.legend()
plt.tight_layout()
#plt.savefig('./figures/moons_dbscan.png', dpi=300)
plt.show()
```
<br>
<br>
# Summary
...
| github_jupyter |
```
import numpy as np
import pandas as pd
from sklearn.metrics import classification_report
from sklearn.model_selection import train_test_split
```
# Shallow Neural Network
```
class ShallowNeuralNetwork:
def __init__(self, n_h=4, type='binary', h_func_type='relu', random_state=-1):
# #hidden_units
self.n_h = n_h
self.type = type
self.h_func_type = h_func_type
self.random_state = random_state
self.functions = {'sigmoid': self.sigmoid,
'relu': self.relu,
'tanh': np.tanh}
self.derivative = {'sigmoid': self.sigmoidDerivative,
'relu': self.reluDerivative,
'tanh': self.tanhDerivative}
def parametersInitialization(self, n_x, n_y):
if self.random_state != -1:
np.random.seed(self.random_state)
# shape(#hidden_units, #features)
W1 = np.random.randn(self.n_h, n_x) * 0.01
# shape(#hidden_units, 1)
b1 = np.zeros((self.n_h, 1))
# shape(#output_unit, #hidden_units)
W2 = np.random.randn(n_y, self.n_h) * 0.01
# shape(#output_unit, 1)
b2 = np.zeros((n_y, 1))
self.parameters = {'W1': W1,
'b1': b1,
'W2': W2,
'b2': b2}
def retrieveParameters(self):
return self.parameters['W1'], self.parameters['b1'], self.parameters['W2'], self.parameters['b2']
def retrieveCache(self):
return self.cache['Z1'], self.cache['A1'], self.cache['Z2'], self.cache['A2']
def retrieveGrads(self):
return self.grads['dW1'], self.grads['db1'], self.grads['dW2'], self.grads['db2']
def sigmoid(self, z):
return (1 / (1 + np.exp(-z)))
def relu(self, z):
return np.maximum(0, z)
def softmax(self, z):
return (np.exp(z) / np.sum(np.exp(z), axis=0))
def sigmoidDerivative(self, Z):
A = self.sigmoid(Z)
return (A * (1 - A))
def reluDerivative(self, Z):
A = np.where(Z >= 0., 1., 0.)
return A
def tanhDerivative(self, Z):
A = np.tanh(Z)
return (1 - A**2)
def forward(self, X):
W1, b1, W2, b2 = self.retrieveParameters()
# shape(#hidden_units, #samples)
Z1 = np.dot(W1, X) + b1
# shape(#hidden_units, #samples)
A1 = self.functions[self.h_func_type](Z1)
# shape(#output_unit, #samples)
Z2 = np.dot(W2, A1) + b2
if self.type == 'multi':
# shape(#output_unit, #samples)
A2 = self.softmax(Z2)
else:
# shape(#output_unit, #samples)
A2 = self.sigmoid(Z2)
self.cache= {'Z1': Z1,
'A1': A1,
'Z2': Z2,
'A2': A2}
return A2
def binary_crossEntropy(self, Y_hat, Y):
# Y_hat : shape(#output_unit, #samples)
# Y : shape(#output_unit, #samples)
m = Y.shape[1]
# shape(#output_unit, #samples)
loss = - (Y * np.log(Y_hat) + (1 - Y) * np.log(1 - Y_hat))
# scalar
cost = np.sum(loss) / m
return cost
def crossEntropy(self, Y_hat, Y):
# Y_hat : shape(#output_unit, #samples)
# Y : shape(#output_unit, #samples)
m = Y.shape[1]
# shape(#samples, )
loss = - (np.sum(np.log(Y_hat) * (Y), axis=0))
# scalar
cost = np.sum(loss) / m
return cost
def backward(self, X, Y):
m = X.shape[1]
W1, b1, W2, b2 = self.retrieveParameters()
Z1, A1, Z2, A2 = self.retrieveCache()
# shape(#output_unit, #samples)
dZ2 = A2 - Y
# shape(#output_unit, #hidden_units)
dW2 = np.dot(dZ2, A1.T) / m
# shape(#output_unit, 1)
db2 = np.sum(dZ2, axis=1, keepdims=True) / m
# shape(#hidden_units, #samples)
dZ1 = (np.dot(W2.T, dZ2)) * (self.derivative[self.h_func_type](Z1))
# shape(#hidden_units, #features)
dW1 = np.dot(dZ1, X.T) / m
# shape(#hidden_units, 1)
db1 = np.sum(dZ1, axis=1, keepdims=True) / m
self.grads = {'dW1': dW1,
'db1': db1,
'dW2': dW2,
'db2': db2}
def updateParameters(self, alpha):
W1, b1, W2, b2 = self.retrieveParameters()
dW1, db1, dW2, db2 = self.retrieveGrads()
# shape(#hidden_units, #features)
W1 = W1 - alpha * dW1
# shape(#hidden_units, 1)
b1 = b1 - alpha * db1
# shape(#output_unit, #hidden_units)
W2 = W2 - alpha * dW2
# shape(#output_unit, 1)
b2 = b2 - alpha * db2
self.parameters = {"W1": W1,
"b1": b1,
"W2": W2,
"b2": b2}
def train(self, X, Y, alpha, epochs, print_cost=False):
# X : shape(#features, #samples)
# Y : shape(1, #samples)
# #features, #output_unit
n_x, n_y = X.shape[0], Y.shape[0]
self.parametersInitialization(n_x, n_y)
for i in range(epochs):
# shape(1, #output_unit)
A2 = self.forward(X)
if self.type == 'multi':
# scalar
cost = self.crossEntropy(A2, Y)
else:
# scalar
cost = self.binary_crossEntropy(A2, Y)
self.backward(X, Y)
self.updateParameters(alpha)
if print_cost and i % (epochs // 10) == 0:
print ("Cost after iteration %i : %f" %(i, cost))
def predict(self, X):
W1, b1, W2, b2 = self.retrieveParameters()
A2 = self.forward(X)
if self.type == 'multi':
Y_pred = A2.argmax(axis=0)
else:
Y_pred = np.where(A2 > 0.5, 1., 0.)
return Y_pred
```
# Load Dataset
```
def loadIrisBinary(path, size=0.2, random_state=0):
df = pd.read_csv(path)
df = df.sample(frac=1, random_state=random_state)
df.Species.replace(('Iris-setosa', 'Iris-versicolor'), (0., 1.), inplace=True)
X_train, X_val, Y_train, Y_val = train_test_split(df.drop(['Species'], axis=1),
df.Species,
test_size=size,
random_state=random_state)
X_train, X_val = X_train.values.T, X_val.values.T
Y_train, Y_val = Y_train.values.reshape(1, -1), Y_val.values.reshape(1, -1)
return X_train, Y_train, X_val, Y_val
def loadIrisMulti(path, size=0.2, random_state=0):
df = pd.read_csv(path)
df.Species.replace(('Iris-setosa', 'Iris-versicolor', 'Iris-virginica'), (0, 1, 2), inplace=True)
df = df.sample(frac=1, random_state=random_state)
X_train, X_val, Y_train, Y_val = train_test_split(df.drop(['Species'], axis=1),
df.Species,
test_size=size,
random_state=random_state)
X_train, X_val = X_train.values.T, X_val.values.T
Y_train, Y_val = Y_train.values, Y_val.values
Y_train = ((np.arange(np.max(Y_train) + 1) == Y_train[:, None]).astype(float)).T
return X_train, Y_train, X_val, Y_val
```
# Training
### Iris Binary
```
X_train, Y_train, X_val, Y_val = loadIrisBinary('data/Iris_binary.csv', size=0.1)
model = ShallowNeuralNetwork(n_h=4, type='binary', h_func_type='tanh', random_state=0)
model.train(X_train, Y_train, 0.1, 100, print_cost=True)
Y_pred = model.predict(X_val)
print(classification_report(Y_val.flatten(), Y_pred.flatten()))
```
### Iris Multiclass
```
X_train, Y_train, X_val, Y_val = loadIrisMulti('data/Iris.csv')
model = ShallowNeuralNetwork(n_h=4, type='multi', h_func_type='tanh', random_state=0)
model.train(X_train, Y_train, 0.1, 150, print_cost=True)
Y_pred = model.predict(X_val)
print(classification_report(Y_val, Y_pred))
```
| github_jupyter |
```
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import numpy as np
#from pandas_profiling import ProfileReport
trans_2018 = pd.read_excel('../data/Transaction Data.xlsx', sheet_name= 'Transactions18')
trans_2019 = pd.read_excel('../data/Transaction Data.xlsx', sheet_name= 'Transactions19')
X_cleaned_classfn = pd.read_csv('../data/X_cleaned_classfn_based_on_rfe.csv')
trans_2019 = trans_2019.rename(columns={'sales_12M': 'sales_2019', 'new_Fund_added_12M':'new_fund_2019'})
full_df = pd.merge(trans_2018,trans_2019, on = 'CONTACT_ID' )
full_df.head()
X_cleaned_classfn.head()
firms_a = pd.read_excel('../data/Firm Information.xlsx', sheet_name= 'Asset fund summary', skiprows= 1)
firms_rep_summary = pd.read_excel('../data/Firm Information.xlsx', sheet_name= 'Rep summary')
firms_rep_summary = firms_rep_summary.rename({'Contact ID':'CONTACT_ID'}, axis = 1)
firms_rep_summary.head()
full_df = pd.merge(full_df, firms_rep_summary, on = 'CONTACT_ID')
missing_diagrams = {
'heatmap' : True , 'dendrogram' : True, 'matrix' : True, 'bar' : True ,
}
profile = ProfileReport(full_df, title ='Nuveen Profile report', missing_diagrams = missing_diagrams, minimal = True)
profile.to_file(output_file = "nuveen_profiling_initial.html")
sns.boxplot(full_df['no_of_sales_12M_1'])
full_df['no_of_sales_12M_1'].fillna(0, inplace = True)
select_columns = [i for i in full_df.columns if i.lower().startswith('no_of')]
select_columns
for i in full_df.columns:
if i.lower().startswith('no_of'):
full_df[i].fillna(0, inplace = True)
full_df[select_columns].describe()
full_df['AUM'].fillna(0, inplace = True)
for i in full_df.columns:
if i.lower().startswith('aum'):
#print(full_df[i].count()) #contains the count of elements in each column
full_df[i].fillna(0, inplace = True)
full_df.info()
full_df['AUM'].describe()
full_df['AUM'] = full_df['AUM'].apply(lambda x: x if x >=0 else 0)
full_df['AUM'].describe()
for i in full_df.columns:
if i.startswith('aum'):
full_df.drop(i, axis= 1, inplace = True)
full_df.info()
for i in full_df.columns:
if i.startswith('refresh_date'):
full_df.drop(i, axis= 1, inplace = True)
full_df.info()
for i in full_df.columns:
if i.lower().startswith('sales'):
full_df[i] = full_df[i].apply(lambda x: x if x >=0 else 0)
full_df['sales_12M'].describe()
for i in full_df.columns:
if i.lower().startswith('new'):
full_df[i] = full_df[i].apply(lambda x: x if x >=0 else 0)
full_df['new_Fund_added_12M'].describe()
plt.figure(figsize = (20,15))
sns.heatmap(full_df.corr(), annot = True)
plt.scatter(full_df['sales_12M'], full_df['sales_2019'])
full_df.info()
full_df['redemption_curr'].head()
for i in full_df.columns:
if i.lower().startswith('redemption'):
full_df[i].fillna(0, inplace = True)
full_df['redemption_12M'].head()
full_df.info()
full_df.to_csv('../data/cleaned_eda_data.csv')
```
| github_jupyter |
# Télécharger en block les données de WaPOR
Dans ce notebook, nous utiliserons le module Python local pour automatiser les étapes de lecture des métadonnées de l'ensemble de données, demander l'URL de téléchargement, télécharger l'ensemble de données raster, prétraiter l'ensemble de données raster en utilisant les informations des métadonnées (telles que le facteur de conversion, l'unité). **Les données téléchargées par ce script seront automatiquement corrigées par le facteur de conversion**.
Exécutez la cellule de code ci-dessous pour importer les packages et les modules nécessaires. Il vous sera demandé de fournir la clé API WaPOR (voir [Start here](0_Start_here.ipynb) la page Démarrer ici pour savoir comment obtenir la clé API). Le module **WaPOR** module gardera votre clé en mémoire et l'enregistrera dans le dossier du module. Si vous souhaitez utiliser une nouvelle clé API, vous devez supprimer ce fichier
**'modules\WaPOR\wapor_api_token.pickle'**
```
import os
import glob
import sys
import shapefile
import matplotlib.pyplot as plt
folder=r"..\..\modules"
sys.path.append(folder) #ajouter un dossier avec des modules locaux aux chemins du système
import WaPOR #importer les modules locaux de 'WaPOR'
```
## Dataset code
Le module **WaPOR** contient les functions *download_dekadal*, *download_monthly*, and *download_yearly* qui téléchargent par lots les données WaPOR pour une résolution temporelle décadaire, mensuelle et annuelle. Pour séparer la couche de données à télécharger, ces fonctions nécessitent le code des données, la coordonnée de la boîte de délimentation, le niveau de données et la date de début et de fin de la période. Le code pour chaque ensemble de données est disponible dans le tableau ci-dessous:
| code | Name |
| :---------: |:-------------:|
| LCC | Classification de couverture du sol |
| PCP | Précipitation |
| RET | Evapotranspiration de référence |
| NPP | Production primaire nette |
| AETI | Evapotranspiration réelle et Interception |
| E | Evaporation |
| I | Interception |
| T | Transpiration |
| PHE | Phénologie |
| GBWP | Productivité brute de l’eau de la biomasse |
Utilisez le code de données sélectionné au lieu de **RET** pour la valeur du paramètre **data** dans les cellules de code ci-dessous.
## Exemple de fonctions du module WaPOR
Les exemples de cellules de code ci-dessous montrent comment le module WaPOR peut être utilisé pour télécharger par lots les données WaPOR: évapotranspiration de référence décadaire, mensuelle et annuelle d'une étendue. L'étendue est définie par la délimitation du fichier de formes du bassin Awash. Les raster seront collectés et prétraités à l'unité correcte mm / période (mm / décade, mm / mois ou mm / an respectivement), et sauvegardées dans le [output folder](data)
### Définir l'étendue du téléchargement et le dossier de sortie
Changer les valeurs xmin, ymin, xmax, ymax à l'étendue du téléchargement.
Si nécessaire changer le chemin d´accès *output_dir* pour l´adresse du dossier où vous voulez sauvegarder les données.
**Remarque sur la syntaxe** : Pour les scripts de ces blocs-notes, les fins de la plupart des fichiers d'entrée et de sortie contiennent "_fh". fh signifie "file handle" et fait référence à un seul fichier. Vous pouvez également voir "_fhs", qui signifie "file handle series". Il s'agit d'une série ou d'un groupe de fichiers.
```
#définir l'étendue à télécharger à partir d'un fichier de formes:
shape_fh=r".\data\Awash_shapefile.shp"
shape=shapefile.Reader(shape_fh)
xmin,ymin,xmax,ymax=shape.bbox
# définir l'étendue à télécharger par coordonnées:
# xmin,ymin,xmax,ymax=(32.65692516077674,-25.16412729789142,32.90420244933813,-25.01450956754679)
output_dir=r'.\data' # dossier pour enregistrer les données
```
### Télécharger les données journalières
Notez que les données journalières ne sont disponibles que pour l'évapotranspiration et les précipitations de référence de niveau 1
```
WaPOR.download_daily(output_dir,
data='RET',
Startdate='2009-01-01',
Enddate='2009-01-31',
latlim=[ymin-0.5, ymax+0.5],
lonlim=[xmin-0.5, xmax+0.5],
level=1,
)
```
### Télécharger les données décadaires
**Notez que:** l'unité de l'ensemble de données de la décade sera convertie en quantité/décade au lieu de la quantité journalière moyenne/jour
```
WaPOR.download_dekadal(output_dir,
data='RET',
Startdate='2009-01-01',
Enddate='2009-12-31',
latlim=[ymin-0.5, ymax+0.5],
lonlim=[xmin-0.5, xmax+0.5],
level=1,
)
```
### Télécharger les données mensuelles
```
WaPOR.download_monthly(output_dir,
data='RET',
Startdate='2009-01-01',
Enddate='2009-12-31',
latlim=[ymin-0.5, ymax+0.5],
lonlim=[xmin-0.5, xmax+0.5],
level=1,
)
```
### Télécharger les données annuelles
```
WaPOR.download_yearly(output_dir,
data='RET',
Startdate='2009-01-01',
Enddate='2009-12-31',
latlim=[ymin-0.5, ymax+0.5],
lonlim=[xmin-0.5, xmax+0.5],
level=1,
)
```
### Télécharger les données saisonnières
**Notez que**: les données saisonnières ne sont disponibles que pour certaines couches (par exemple, la productivité brute de l'eau de la biomasse, la phénologie)
```
WaPOR.download_seasonal(output_dir,
data='GBWP',
Startdate='2009-01-01',
Enddate='2010-12-31',
latlim=[ymin-0.5, ymax+0.5],
lonlim=[xmin-0.5, xmax+0.5],
level=2,
)
WaPOR.download_seasonal(output_dir,
data='PHE',
Startdate='2009-01-01',
Enddate='2010-12-31',
latlim=[ymin-0.5, ymax+0.5],
lonlim=[xmin-0.5, xmax+0.5],
level=2,
)
```
# Exercice
Télécharger les données mensuelles de niveau 1 des précipitations, de l'évapotranspiration et de l'interception réelles et de la classification annuelle de la couverture terrestre de 2009 à 2010 pour la zone délimitée par le fichier de formes ".\data\Awash_shapefile.shp"
Utilisez les fonctions du module **WaPOR** en suivant les exemples donnés ci-dessus.
```
'''
Ecrivez votre code ici
'''
```
| github_jupyter |
##### Copyright 2019 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/examples/blob/master/courses/udacity_intro_to_tensorflow_for_deep_learning/l07c01_saving_and_loading_models.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/examples/blob/master/courses/udacity_intro_to_tensorflow_for_deep_learning/l07c01_saving_and_loading_models.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
</table>
# Saving and Loading Models
In this tutorial we will learn how we can take a trained model, save it, and then load it back to keep training it or use it to perform inference. In particular, we will use transfer learning to train a classifier to classify images of cats and dogs, just like we did in the previous lesson. We will then take our trained model and save it as an HDF5 file, which is the format used by Keras. We will then load this model, use it to perform predictions, and then continue to train the model. Finally, we will save our trained model as a TensorFlow SavedModel and then we will download it to a local disk, so that it can later be used for deployment in different platforms.
## Concepts that will be covered in this Colab
1. Saving models in HDF5 format for Keras
2. Saving models in the TensorFlow SavedModel format
3. Loading models
4. Download models to Local Disk
Before starting this Colab, you should reset the Colab environment by selecting `Runtime -> Reset all runtimes...` from menu above.
# Imports
In this Colab we will use the TensorFlow 2.0 Beta version.
```
try:
# Use the %tensorflow_version magic if in colab.
%tensorflow_version 2.x
except Exception:
!pip install -U "tensorflow-gpu==2.0.0rc0"
!pip install -U tensorflow_hub
!pip install -U tensorflow_datasets
```
Some normal imports we've seen before.
```
from __future__ import absolute_import, division, print_function, unicode_literals
import time
import numpy as np
import matplotlib.pylab as plt
import tensorflow as tf
import tensorflow_hub as hub
import tensorflow_datasets as tfds
tfds.disable_progress_bar()
from tensorflow.keras import layers
```
# Part 1: Load the Cats vs. Dogs Dataset
We will use TensorFlow Datasets to load the Dogs vs Cats dataset.
```
(train_examples, validation_examples), info = tfds.load(
'cats_vs_dogs',
split=['train[:80%]', 'train[80%:]'],
with_info=True,
as_supervised=True,
)
```
The images in the Dogs vs. Cats dataset are not all the same size. So, we need to reformat all images to the resolution expected by MobileNet (224, 224)
```
def format_image(image, label):
# `hub` image modules exepct their data normalized to the [0,1] range.
image = tf.image.resize(image, (IMAGE_RES, IMAGE_RES))/255.0
return image, label
num_examples = info.splits['train'].num_examples
BATCH_SIZE = 32
IMAGE_RES = 224
train_batches = train_examples.cache().shuffle(num_examples//4).map(format_image).batch(BATCH_SIZE).prefetch(1)
validation_batches = validation_examples.cache().map(format_image).batch(BATCH_SIZE).prefetch(1)
```
# Part 2: Transfer Learning with TensorFlow Hub
We will now use TensorFlow Hub to do Transfer Learning.
```
URL = "https://tfhub.dev/google/tf2-preview/mobilenet_v2/feature_vector/4"
feature_extractor = hub.KerasLayer(URL,
input_shape=(IMAGE_RES, IMAGE_RES,3))
```
Freeze the variables in the feature extractor layer, so that the training only modifies the final classifier layer.
```
feature_extractor.trainable = False
```
## Attach a classification head
Now wrap the hub layer in a `tf.keras.Sequential` model, and add a new classification layer.
```
model = tf.keras.Sequential([
feature_extractor,
layers.Dense(2)
])
model.summary()
```
## Train the model
We now train this model like any other, by first calling `compile` followed by `fit`.
```
model.compile(
optimizer='adam',
loss=tf.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=['accuracy'])
EPOCHS = 3
history = model.fit(train_batches,
epochs=EPOCHS,
validation_data=validation_batches)
```
## Check the predictions
Get the ordered list of class names.
```
class_names = np.array(info.features['label'].names)
class_names
```
Run an image batch through the model and convert the indices to class names.
```
image_batch, label_batch = next(iter(train_batches.take(1)))
image_batch = image_batch.numpy()
label_batch = label_batch.numpy()
predicted_batch = model.predict(image_batch)
predicted_batch = tf.squeeze(predicted_batch).numpy()
predicted_ids = np.argmax(predicted_batch, axis=-1)
predicted_class_names = class_names[predicted_ids]
predicted_class_names
```
Let's look at the true labels and predicted ones.
```
print("Labels: ", label_batch)
print("Predicted labels: ", predicted_ids)
plt.figure(figsize=(10,9))
for n in range(30):
plt.subplot(6,5,n+1)
plt.imshow(image_batch[n])
color = "blue" if predicted_ids[n] == label_batch[n] else "red"
plt.title(predicted_class_names[n].title(), color=color)
plt.axis('off')
_ = plt.suptitle("Model predictions (blue: correct, red: incorrect)")
```
# Part 3: Save as Keras `.h5` model
Now that we've trained the model, we can save it as an HDF5 file, which is the format used by Keras. Our HDF5 file will have the extension '.h5', and it's name will correpond to the current time stamp.
```
t = time.time()
export_path_keras = "./{}.h5".format(int(t))
print(export_path_keras)
model.save(export_path_keras)
!ls
```
You can later recreate the same model from this file, even if you no longer have access to the code that created the model.
This file includes:
- The model's architecture
- The model's weight values (which were learned during training)
- The model's training config (what you passed to `compile`), if any
- The optimizer and its state, if any (this enables you to restart training where you left off)
# Part 4: Load the Keras `.h5` Model
We will now load the model we just saved into a new model called `reloaded`. We will need to provide the file path and the `custom_objects` parameter. This parameter tells keras how to load the `hub.KerasLayer` from the `feature_extractor` we used for transfer learning.
```
reloaded = tf.keras.models.load_model(
export_path_keras,
# `custom_objects` tells keras how to load a `hub.KerasLayer`
custom_objects={'KerasLayer': hub.KerasLayer})
reloaded.summary()
```
We can check that the reloaded model and the previous model give the same result
```
result_batch = model.predict(image_batch)
reloaded_result_batch = reloaded.predict(image_batch)
```
The difference in output should be zero:
```
(abs(result_batch - reloaded_result_batch)).max()
```
As we can see, the reult is 0.0, which indicates that both models made the same predictions on the same batch of images.
# Keep Training
Besides making predictions, we can also take our `reloaded` model and keep training it. To do this, you can just train the `reloaded` as usual, using the `.fit` method.
```
EPOCHS = 3
history = reloaded.fit(train_batches,
epochs=EPOCHS,
validation_data=validation_batches)
```
# Part 5: Export as SavedModel
You can also export a whole model to the TensorFlow SavedModel format. SavedModel is a standalone serialization format for Tensorflow objects, supported by TensorFlow serving as well as TensorFlow implementations other than Python. A SavedModel contains a complete TensorFlow program, including weights and computation. It does not require the original model building code to run, which makes it useful for sharing or deploying (with TFLite, TensorFlow.js, TensorFlow Serving, or TFHub).
The SavedModel files that were created contain:
* A TensorFlow checkpoint containing the model weights.
* A SavedModel proto containing the underlying Tensorflow graph. Separate graphs are saved for prediction (serving), train, and evaluation. If the model wasn't compiled before, then only the inference graph gets exported.
* The model's architecture config, if available.
Let's save our original `model` as a TensorFlow SavedModel. To do this we will use the `tf.saved_model.save()` function. This functions takes in the model we want to save and the path to the folder where we want to save our model.
This function will create a folder where you will find an `assets` folder, a `variables` folder, and the `saved_model.pb` file.
```
t = time.time()
export_path_sm = "./{}".format(int(t))
print(export_path_sm)
tf.saved_model.save(model, export_path_sm)
!ls {export_path_sm}
```
# Part 6: Load SavedModel
Now, let's load our SavedModel and use it to make predictions. We use the `tf.saved_model.load()` function to load our SavedModels. The object returned by `tf.saved_model.load` is 100% independent of the code that created it.
```
reloaded_sm = tf.saved_model.load(export_path_sm)
```
Now, let's use the `reloaded_sm` (reloaded SavedModel) to make predictions on a batch of images.
```
reload_sm_result_batch = reloaded_sm(image_batch, training=False).numpy()
```
We can check that the reloaded SavedModel and the previous model give the same result.
```
(abs(result_batch - reload_sm_result_batch)).max()
```
As we can see, the result is 0.0, which indicates that both models made the same predictions on the same batch of images.
# Part 7: Loading the SavedModel as a Keras Model
The object returned by `tf.saved_model.load` is not a Keras object (i.e. doesn't have `.fit`, `.predict`, `.summary`, etc. methods). Therefore, you can't simply take your `reloaded_sm` model and keep training it by running `.fit`. To be able to get back a full keras model from the Tensorflow SavedModel format we must use the `tf.keras.models.load_model` function. This function will work the same as before, except now we pass the path to the folder containing our SavedModel.
```
t = time.time()
export_path_sm = "./{}".format(int(t))
print(export_path_sm)
tf.saved_model.save(model, export_path_sm)
reload_sm_keras = tf.keras.models.load_model(
export_path_sm,
custom_objects={'KerasLayer': hub.KerasLayer})
reload_sm_keras.summary()
```
Now, let's use the `reloaded_sm)keras` (reloaded Keras model from our SavedModel) to make predictions on a batch of images.
```
result_batch = model.predict(image_batch)
reload_sm_keras_result_batch = reload_sm_keras.predict(image_batch)
```
We can check that the reloaded Keras model and the previous model give the same result.
```
(abs(result_batch - reload_sm_keras_result_batch)).max()
```
# Part 8: Download your model
You can download the SavedModel to your local disk by creating a zip file. We wil use the `-r` (recursice) option to zip all subfolders.
```
!zip -r model.zip {export_path_sm}
```
The zip file is saved in the current working directory. You can see what the current working directory is by running:
```
!ls
```
Once the file is zipped, you can download it to your local disk.
```
try:
from google.colab import files
files.download('./model.zip')
except ImportError:
pass
```
The `files.download` command will search for files in your current working directory. If the file you want to download is in a directory other than the current working directory, you have to include the path to the directory where the file is located.
| github_jupyter |
```
import json
import numpy as np
import pandas as pd
from sklearn.model_selection import train_test_split
from textwiser import TextWiser, Embedding, Transformation
```
## Goodreads Preprocessing
This notebook contains the preprocessing necessary to run the Goodreads experiments. Before running the code, download the young adult subset of Goodreads dataset [here](https://sites.google.com/eng.ucsd.edu/ucsdbookgraph/home).
```
# Input
books_json = 'goodreads_books_young_adult.json'
interactions_json = 'goodreads_interactions_young_adult.json'
reviews_json = 'goodreads_reviews_young_adult.json'
# Output
responses_csv = 'responses.csv.gz'
user_features_csv = 'user_features.csv.gz'
```
We first get the relevent columns of the interactions. We binarize the interactions where any rating greater than or equal to 4 as a positive response, anything else is a negative response.
```
with open(interactions_json) as fp:
interactions = []
for line in fp:
interactions.append(json.loads(line))
interactions = pd.DataFrame(interactions)
interactions = interactions[interactions['is_read']][['user_id', 'book_id', 'rating']]
interactions.head()
interactions['rating'].hist()
interactions['rating'].describe()
interactions['response'] = interactions['rating'] >= 4
interactions.drop('rating', axis=1, inplace=True)
interactions.head()
```
We then look at the reviews and limit the userbase to the subset where the users have reviews _and_ interactions.
```
with open(reviews_json) as fp:
reviews = []
for line in fp:
reviews.append(json.loads(line))
reviews = pd.DataFrame(reviews)
reviews.head()
user_ids = np.intersect1d(interactions['user_id'].unique(), reviews['user_id'].unique())
print(user_ids.shape[0])
interactions = interactions[interactions['user_id'].isin(user_ids)]
reviews = reviews[reviews['user_id'].isin(user_ids)]
interactions.to_csv(responses_csv, index=False)
```
We then go through the books dataset, calculating the most popular shelves.
```
with open(books_json) as fp:
books = []
for line in fp:
books.append(json.loads(line))
books = pd.DataFrame(books)
books.head()
books['popular_shelves']
from collections import Counter
from collections import ChainMap
shelves = sum(books['popular_shelves'].apply(lambda x: Counter(ChainMap(*[{t['name']: int(t['count'])} for t in x]))), Counter())
shelves
```
We use the top 100 most common shelves as genres. We create binary features for all books based on whether a book contains one of the top 100 shelves.
In order to weed-out the false positives, we only look at the top 5 shelves for each book.
```
shelf_names = [t[0] for t in shelves.most_common(100)]
shelves_per_book = books['popular_shelves'].str[:5].apply(lambda x: [t['name'] for t in x])
shelves_per_book
contains_shelf = pd.concat([books['book_id']] + [shelves_per_book.apply(lambda x: shelf_name in x) for shelf_name in shelf_names], axis=1)
contains_shelf.columns = ['book_id'] + shelf_names
contains_shelf.head()
```
Since we don't have any user features, we construct user features by going through the user reviews and featurizing them using TfIdf + NMF combination. We use the [TextWiser library](https://github.com/fidelity/textwiser) to generate the review features.
```
np.random.seed(42)
tw = TextWiser(Embedding.TfIdf(), Transformation.NMF(n_components=30))
review_vectors = tw.fit_transform(reviews['review_text'].values)
feat_cols = [f'feat_{i}' for i in range(review_vectors.shape[1])]
reviews = pd.concat([
reviews.reset_index(drop=True),
pd.DataFrame(review_vectors, columns=feat_cols),
], axis=1)
reviews.loc[reviews['rating'] == 0, 'rating'] = 1
```
The first set of user features are the aggregated review vectors over all the books the user has consumed.
```
user_features = reviews.groupby('user_id').apply(lambda x: pd.Series(np.mean(x[feat_cols], axis=0), index=feat_cols))
user_features = user_features.reset_index()
user_features
```
We also use create a train/test split based on the users.
```
train_inds, test_inds = train_test_split(np.arange(user_features.shape[0]), test_size=0.3, random_state=40)
user_features.loc[train_inds, 'set'] = 'train'
user_features.loc[test_inds, 'set'] = 'test'
user_features.head()
user_features.to_csv(user_features_csv, index=False)
interactions = pd.read_csv(responses_csv, dtype={'book_id': str})
interactions.head()
```
The second set of features we have for users are whether the user has read any book from a given genre/shelf.
```
user_features = pd.read_csv(user_features_csv)
user_genre = interactions.join(contains_shelf.set_index('book_id'), on='book_id').groupby('user_id').max()
user_genre.drop(columns=['book_id', 'response'], inplace=True)
user_genre.head()
user_genre.shape
user_features = user_features.join(user_genre, on='user_id')
user_features.head()
user_features.to_csv(user_features_csv, index=False)
```
At the end, we have two data files:
1. User Features, which contains the averaged TfIdf-NMF vectors of all books the user has read, and whether the user has read a book that is in one of the top 100 shelves. This data also contains a column that indicates whether a user is in the test set or in the train set.
2. Response matrix, which contains user/item/binary response tuples.
| github_jupyter |
# Build multiclass classifiers with Amazon SageMaker linear learner
Amazon SageMaker is a fully managed service for scalable training and hosting of machine learning models. We're adding multiclass classification support to the linear learner algorithm in Amazon SageMaker. Linear learner already provides convenient APIs for linear models such as logistic regression for ad click prediction, fraud detection, or other classification problems, and linear regression for forecasting sales, predicting delivery times, or other problems where you want to predict a numerical value. If you haven't worked with linear learner before, you might want to start with the [documentation](https://docs.aws.amazon.com/sagemaker/latest/dg/linear-learner.html) or our previous [example notebook](https://github.com/awslabs/amazon-sagemaker-examples/blob/master/scientific_details_of_algorithms/linear_learner_class_weights_loss_functions/linear_learner_class_weights_loss_functions.ipynb) on this algorithm. If it's your first time working with Amazon SageMaker, you can get started [here](https://aws.amazon.com/about-aws/whats-new/2017/11/introducing-amazon-sagemaker/).
In this example notebook we'll cover three aspects of training a multiclass classifier with linear learner:
1. Training a multiclass classifier
1. Multiclass classification metrics
1. Training with balanced class weights
## Training a multiclass classifier
Multiclass classification is a machine learning task where the outputs are known to be in a finite set of labels. For example, we might classify emails by assigning each one a label from the set *inbox, work, shopping, spam*. Or we might try to predict what a customer will buy from the set *shirt, mug, bumper_sticker, no_purchase*. If we have a dataset where each example has numerical features and a known categorical label, we can train a multiclass classifier.
### Related problems: binary, multiclass, and multilabel
Multiclass classification is related to two other machine learning tasks, binary classification and the multilabel problem. Binary classification is already supported by linear learner, and multiclass classification is available with linear learner starting today, but multilabel support is not yet available from linear learner.
If there are only two possible labels in your dataset, then you have a binary classification problem. Examples include predicting whether a transaction will be fraudulent or not based on transaction and customer data, or detecting whether a person is smiling or not based on features extracted from a photo. For each example in your dataset, one of the possible labels is correct and the other is incorrect. The person is smiling or not smiling.
If there are more than two possible labels in your dataset, then you have a multiclass classification problem. For example, predicting whether a transaction will be fraudulent, cancelled, returned, or completed as usual. Or detecting whether a person in a photo is smiling, frowning, surprised, or frightened. There are multiple possible labels, but only one is correct at a time.
If there are multiple labels, and a single training example can have more than one correct label, then you have a multilabel problem. For example, tagging an image with tags from a known set. An image of a dog catching a Frisbee at the park might be labeled as *outdoors*, *dog*, and *park*. For any given image, those three labels could all be true, or all be false, or any combination. Although we haven't added support for multilabel problems yet, there are a couple of ways you can solve a multilabel problem with linear learner today. You can train a separate binary classifier for each label. Or you can train a multiclass classifier and predict not only the top class, but the top k classes, or all classes with probability scores above some threshold.
Linear learner uses a softmax loss function to train multiclass classifiers. The algorithm learns a set of weights for each class, and predicts a probability for each class. We might want to use these probabilities directly, for example if we're classifying emails as *inbox, work, shopping, spam* and we have a policy to flag as spam only if the class probability is over 99.99%. But in many multiclass classification use cases, we'll simply take the class with highest probability as the predicted label.
### Hands-on example: predicting forest cover type
As an example of multiclass prediction, let's take a look at the [Covertype dataset](https://archive.ics.uci.edu/ml/datasets/covertype) (copyright Jock A. Blackard and Colorado State University). The dataset contains information collected by the US Geological Survey and the US Forest Service about wilderness areas in northern Colorado. The features are measurements like soil type, elevation, and distance to water, and the labels encode the type of trees - the forest cover type - for each location. The machine learning task is to predict the cover type in a given location using the features. We'll download and explore the dataset, then train a multiclass classifier with linear learner using the Python SDK.
```
# import data science and visualization libraries
%matplotlib inline
from sklearn.model_selection import train_test_split
import pandas as pd
import numpy as np
import seaborn as sns
# download the raw data and unzip
!wget https://archive.ics.uci.edu/ml/machine-learning-databases/covtype/covtype.data.gz
!gunzip covtype.data.gz
# read the csv and extract features and labels
covtype = pd.read_csv("covtype.data", delimiter=",", dtype="float32").values
covtype_features, covtype_labels = covtype[:, :54], covtype[:, 54]
# transform labels to 0 index
covtype_labels -= 1
# shuffle and split into train and test sets
np.random.seed(0)
train_features, test_features, train_labels, test_labels = train_test_split(
covtype_features, covtype_labels, test_size=0.2
)
# further split the test set into validation and test sets
val_features, test_features, val_labels, test_labels = train_test_split(
test_features, test_labels, test_size=0.5
)
```
Note that we transformed the labels to a zero index rather than an index starting from one. That step is important, since linear learner requires the class labels to be in the range \[0, k-1\], where k is the number of labels. Amazon SageMaker algorithms expect the `dtype` of all feature and label values to be `float32`. Also note that we shuffled the order of examples in the training set. We used the `train_test_split` method from `numpy`, which shuffles the rows by default. That's important for algorithms trained using stochastic gradient descent. Linear learner, as well as most deep learning algorithms, use stochastic gradient descent for optimization. Shuffle your training examples, unless your data have some natural ordering which needs to be preserved, such as a forecasting problem where the training examples should all have time stamps earlier than the test examples.
We split the data into training, validation, and test sets with an 80/10/10 ratio. Using a validation set will improve training, since linear learner uses the validation data to stop training once overfitting is detected. That means shorter training times and more accurate predictions. We can also provide a test set to linear learner. The test set will not affect the final model, but algorithm logs will contain metrics from the final model's performance on the test set. Later on in this example notebook, we'll also use the test set locally to dive a little bit deeper on model performance.
### Exploring the data
Let's take a look at the mix of class labels present in training data. We'll add meaningful category names using the mapping provided in the [dataset documentation](https://archive.ics.uci.edu/ml/machine-learning-databases/covtype/covtype.info).
```
# assign label names and count label frequencies
label_map = {
0: "Spruce/Fir",
1: "Lodgepole Pine",
2: "Ponderosa Pine",
3: "Cottonwood/Willow",
4: "Aspen",
5: "Douglas-fir",
6: "Krummholz",
}
label_counts = (
pd.DataFrame(data=train_labels)[0]
.map(label_map)
.value_counts(sort=False)
.sort_index(ascending=False)
)
label_counts.plot(kind="barh", color="tomato", title="Label Counts")
```
We can see that some forest cover types are much more common than others. Lodgepole Pine and Spruce/Fir are both well represented. Some labels, such as Cottonwood/Willow, are extremely rare. Later in this example notebook, we'll see how to fine-tune the algorithm depending on how important these rare categories are for our use case. But first we'll train with the defaults for the best all-around model.
### Training a classifier using the Amazon SageMaker Python SDK
We'll use the high-level estimator class `LinearLearner` to instantiate our training job and inference endpoint. For an example using the Python SDK's generic `Estimator` class, take a look at this previous [example notebook](https://github.com/awslabs/amazon-sagemaker-examples/blob/master/scientific_details_of_algorithms/linear_learner_class_weights_loss_functions/linear_learner_class_weights_loss_functions.ipynb). The generic Python SDK estimator offers some more control options, but the high-level estimator is more succinct and has some advantages. One is that we don't need to specify the location of the algorithm container we want to use for training. It will pick up the latest version of the linear learner algorithm. Another advantage is that some code errors will be surfaced before a training cluster is spun up, rather than after. For example, if we try to pass `n_classes=7` instead of the correct `num_classes=7`, then the high-level estimator will fail immediately, but the generic Python SDK estimator will spin up a cluster before failing.
```
import sagemaker
from sagemaker.amazon.amazon_estimator import RecordSet
import boto3
# instantiate the LinearLearner estimator object
multiclass_estimator = sagemaker.LinearLearner(
role=sagemaker.get_execution_role(),
train_instance_count=1,
train_instance_type="ml.m4.xlarge",
predictor_type="multiclass_classifier",
num_classes=7,
)
```
Linear learner accepts training data in protobuf or csv content types, and accepts inference requests in protobuf, csv, or json content types. Training data have features and ground-truth labels, while the data in an inference request has only features. In a production pipeline, we recommend converting the data to the Amazon SageMaker protobuf format and storing it in S3. However, to get up and running quickly, we provide a convenience method `record_set` for converting and uploading when the dataset is small enough to fit in local memory. It accepts `numpy` arrays like the ones we already have, so we'll use it here. The `RecordSet` object will keep track of the temporary S3 location of our data.
```
# wrap data in RecordSet objects
train_records = multiclass_estimator.record_set(train_features, train_labels, channel="train")
val_records = multiclass_estimator.record_set(val_features, val_labels, channel="validation")
test_records = multiclass_estimator.record_set(test_features, test_labels, channel="test")
# start a training job
multiclass_estimator.fit([train_records, val_records, test_records])
```
## Multiclass classification metrics
Now that we have a trained model, we want to make predictions and evaluate model performance on our test set. For that we'll need to deploy a model hosting endpoint to accept inference requests using the estimator API:
```
# deploy a model hosting endpoint
multiclass_predictor = multiclass_estimator.deploy(
initial_instance_count=1, instance_type="ml.m4.xlarge"
)
```
We'll add a convenience function for parsing predictions and evaluating model metrics. It will feed test features to the endpoint and receive predicted test labels. To evaluate the models we create, we'll capture predicted test labels and compare them to actuals using some common multiclass classification metrics. As mentioned earlier, we're extracting the `predicted_label` from each response payload. That's the class with the highest predicted probability. We'll get one class label per example. To get a vector of seven probabilities for each example (the predicted probability for each class) , we would extract the `score` from the response payload. Details of linear learner's response format are in the [documentation](https://docs.aws.amazon.com/sagemaker/latest/dg/LL-in-formats.html).
```
def evaluate_metrics(predictor, test_features, test_labels):
"""
Evaluate a model on a test set using the given prediction endpoint. Display classification metrics.
"""
# split the test dataset into 100 batches and evaluate using prediction endpoint
prediction_batches = [predictor.predict(batch) for batch in np.array_split(test_features, 100)]
# parse protobuf responses to extract predicted labels
extract_label = lambda x: x.label["predicted_label"].float32_tensor.values
test_preds = np.concatenate(
[np.array([extract_label(x) for x in batch]) for batch in prediction_batches]
)
test_preds = test_preds.reshape((-1,))
# calculate accuracy
accuracy = (test_preds == test_labels).sum() / test_labels.shape[0]
# calculate recall for each class
recall_per_class, classes = [], []
for target_label in np.unique(test_labels):
recall_numerator = np.logical_and(
test_preds == target_label, test_labels == target_label
).sum()
recall_denominator = (test_labels == target_label).sum()
recall_per_class.append(recall_numerator / recall_denominator)
classes.append(label_map[target_label])
recall = pd.DataFrame({"recall": recall_per_class, "class_label": classes})
recall.sort_values("class_label", ascending=False, inplace=True)
# calculate confusion matrix
label_mapper = np.vectorize(lambda x: label_map[x])
confusion_matrix = pd.crosstab(
label_mapper(test_labels),
label_mapper(test_preds),
rownames=["Actuals"],
colnames=["Predictions"],
normalize="index",
)
# display results
sns.heatmap(confusion_matrix, annot=True, fmt=".2f", cmap="YlGnBu").set_title(
"Confusion Matrix"
)
ax = recall.plot(
kind="barh", x="class_label", y="recall", color="steelblue", title="Recall", legend=False
)
ax.set_ylabel("")
print("Accuracy: {:.3f}".format(accuracy))
# evaluate metrics of the model trained with default hyperparameters
evaluate_metrics(multiclass_predictor, test_features, test_labels)
```
The first metric reported is accuracy. Accuracy for multiclass classification means the same thing as it does for binary classification: the percent of predicted labels which match ground-truth labels. Our model predicts the right type of forest cover over 72% of the time.
Next we see the confusion matrix and a plot of class recall for each label. Recall is a binary classification metric which is also useful in the multiclass setting. It measures the model's accuracy when the true label belongs to the first class, the second class, and so on. If we average the recall values across all classes, we get a metric called *macro recall*, which you can find reported in the algorithm logs. You'll also find *macro precision* and *macro f-score*, which are constructed the same way.
The recall achieved by our model varies widely among the classes. Recall is high for the most common labels, but is very poor for the rarer labels like Aspen or Cottonwood/Willow. Our predictions are right most of the time, but when the true cover type is a rare one like Aspen or Cottonwood/Willow, our model tends to predict wrong.
A confusion matrix is a tool for visualizing the performance of a multiclass model. It has entries for all possible combinations of correct and incorrect predictions, and shows how often each one was made by our model. It has been row-normalized: each row sums to one, so that entries along the diagonal correspond to recall. For example, the first row shows that when the true label is Aspen, the model predicts correctly only 1% of the time, and incorrectly predicts Lodgepole Pine 95% of the time. The second row shows that when the true forest cover type is Cottonwood/Willow, the model has 27% recall, and incorrectly predicts Ponderosa Pine 65% of the time. If our model had 100% accuracy, and therefore 100% recall in every class, then all of the predictions would fall along the diagonal of the confusion matrix.
It's normal that the model performs poorly on very rare classes. It doesn't have much data to learn about them, and it was optimized for global performance. By default, linear learner uses the softmax loss function, which optimizes the likelihood of a multinomial distribution. It's similar in principle to optimizing global accuracy.
But what if one of the rare class labels is especially important to our use case? For example, maybe we're predicting customer outcomes, and one of the potential outcomes is a dissatisfied customer. Hopefully that's a rare outcome, but it might be one that's especially important to predict and act on quickly. In that case, we might be able to sacrifice a bit of overall accuracy in exchange for much improved recall on rare classes. Let's see how.
## Training with balanced class weights
Class weights alter the loss function optimized by the linear learner algorithm. They put more weight on rarer classes so that the importance of each class is equal. Without class weights, each example in the training set is treated equally. If 80% of those examples have labels from one overrepresented class, that class will get 80% of the attention during model training. With balanced class weights, each class has the same amount of influence during training.
With balanced class weights turned on, linear learner will count label frequencies in your training set. This is done efficiently using a sample of the training set. The weights will be the inverses of the frequencies. A label that's present in 1/3 of the sampled training examples will get a weight of 3, and a rare label that's present in only 0.001% of the examples will get a weight of 100,000. A label that's not present at all in the sampled training examples will get a weight of 1,000,000 by default. To turn on class weights, use the `balance_multiclass_weights` hyperparameter:
```
# instantiate the LinearLearner estimator object
balanced_multiclass_estimator = sagemaker.LinearLearner(
role=sagemaker.get_execution_role(),
train_instance_count=1,
train_instance_type="ml.m4.xlarge",
predictor_type="multiclass_classifier",
num_classes=7,
balance_multiclass_weights=True,
)
# start a training job
balanced_multiclass_estimator.fit([train_records, val_records, test_records])
# deploy a model hosting endpoint
balanced_multiclass_predictor = balanced_multiclass_estimator.deploy(
initial_instance_count=1, instance_type="ml.m4.xlarge"
)
# evaluate metrics of the model trained with balanced class weights
evaluate_metrics(balanced_multiclass_predictor, test_features, test_labels)
```
The difference made by class weights is immediately clear from the confusion matrix. The predictions now line up nicely along the diagonal of the matrix, meaning predicted labels match actual labels. Recall for the rare Aspen class was only 1%, but now recall for every class is above 50%. That's a huge improvement in our ability to predict rare labels correctly.
But remember that the confusion matrix has each row normalized to sum to 1. Visually, we've given each class equal weight in our diagnostic tool. That emphasizes the gains we've made in rare classes, but it de-emphasizes the price we'll pay in terms of predicting more common classes. Recall for the most common class, Lodgepole Pine, has gone from 81% to 52%. For that reason, overall accuracy also decreased from 72% to 59%. To decide whether to use balanced class weights for your application, consider the business impact of making errors in common cases and how it compares to the impact of making errors in rare cases.
#### Deleting the hosting endpoints
Finally, we'll delete the hosting endpoints. The machines used for training spin down automatically, but the hosting endpoints remain active until you shut them down.
```
# delete endpoints
multiclass_predictor.delete_endpoint()
balanced_multiclass_predictor.delete_endpoint()
```
## Conclusion
In this example notebook, we introduced the new multiclass classification feature of the Amazon SageMaker linear learner algorithm. We showed how to fit a multiclass model using the convenient high-level estimator API, and how to evaluate and interpret model metrics. We also showed how to achieve higher recall for rare classes using linear learner's automatic class weights calculation. Try Amazon SageMaker and linear learner on your classification problems today!
| github_jupyter |
# Evaluation of Diagnostic Models
Welcome to the second assignment of course 1. In this assignment, we will be working with the results of the X-ray classification model we developed in the previous assignment. In order to make the data processing a bit more manageable, we will be working with a subset of our training, and validation datasets. We will also use our manually labeled test dataset of 420 X-rays.
As a reminder, our dataset contains X-rays from 14 different conditions diagnosable from an X-ray. We'll evaluate our performance on each of these classes using the classification metrics we learned in lecture.
## Outline
Click on these links to jump to a particular section of this assignment!
- [1. Packages](#1)
- [2. Overview](#2)
- [3. Metrics](#3)
- [3.1 True Positives, False Positives, True Negatives, and False Negatives](#3-1)
- [3.2 Accuracy](#3-2)
- [3.3 Prevalence](#3-3)
- [3.4 Sensitivity and Specificity](#3-4)
- [3.5 PPV and NPV](#3-5)
- [3.6 ROC Curve](#3-6)
- [4. Confidence Intervals](#4)
- [5. Precision-Recall Curve](#5)
- [6. F1 Score](#6)
- [7. Calibration](#7)
**By the end of this assignment you will learn about:**
1. Accuracy
1. Prevalence
1. Specificity & Sensitivity
1. PPV and NPV
1. ROC curve and AUCROC (c-statistic)
1. Confidence Intervals
<a name='1'></a>
## 1. Packages
In this assignment, we'll make use of the following packages:
- [numpy](https://docs.scipy.org/doc/numpy/) is a popular library for scientific computing
- [matplotlib](https://matplotlib.org/3.1.1/contents.html) is a plotting library compatible with numpy
- [pandas](https://pandas.pydata.org/docs/) is what we'll use to manipulate our data
- [sklearn](https://scikit-learn.org/stable/index.html) will be used to measure the performance of our model
Run the next cell to import all the necessary packages as well as custom util functions.
```
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import util
```
<a name='2'></a>
## 2. Overview
We'll go through our evaluation metrics in the following order.
- Metrics
- TP, TN, FP, FN
- Accuracy
- Prevalence
- Sensitivity and Specificity
- PPV and NPV
- AUC
- Confidence Intervals
Let's take a quick peek at our dataset. The data is stored in two CSV files called `train_preds.csv` and `valid_preds.csv`. We have precomputed the model outputs for our test cases. We'll work with these predictions and the true class labels throughout the assignment.
```
train_results = pd.read_csv("train_preds.csv")
valid_results = pd.read_csv("valid_preds.csv")
# the labels in our dataset
class_labels = ['Cardiomegaly',
'Emphysema',
'Effusion',
'Hernia',
'Infiltration',
'Mass',
'Nodule',
'Atelectasis',
'Pneumothorax',
'Pleural_Thickening',
'Pneumonia',
'Fibrosis',
'Edema',
'Consolidation']
# the labels for prediction values in our dataset
pred_labels = [l + "_pred" for l in class_labels]
train_results.head()
valid_results.head()
print(pred_labels)
```
Extract the labels (y) and the predictions (pred).
```
y = valid_results[class_labels].values # shape => 1000, 14
pred = valid_results[pred_labels].values # shape => 1000, 14
# let us see what is stored in y and pred
# Mental note: Imagine 'y' and 'pred' as the ground truth
# and the model's prediction for 14 classes as you move
# across each row in the dataframe/table
print("y : " , y[10,:])
print("\npred : ", pred[10,:])
```
Run the next cell to view them side by side.
```
# let's take a peek at our dataset
# First 14 columns are 'y' and
# last 14 columns are 'pred'
valid_results[np.concatenate([class_labels, pred_labels])].head()
```
To further understand our dataset details, here's a histogram of the number of samples for each label in the validation dataset:
```
plt.xticks(rotation=90)
plt.bar(x = class_labels, height= y.sum(axis=0));
```
It seem like our dataset has an imbalanced population of samples. Specifically, our dataset has a small number of patients diagnosed with a `Hernia`.
<a name='3'></a>
## 3 Metrics
<a name='3-1'></a>
### 3.1 True Positives, False Positives, True Negatives, and False Negatives
The most basic statistics to compute from the model predictions are the true positives, true negatives, false positives, and false negatives.
As the name suggests
- true positive (TP): The model classifies the example as positive, and the actual label also positive.
- false positive (FP): The model classifies the example as positive, **but** the actual label is negative.
- true negative (TN): The model classifies the example as negative, and the actual label is also negative.
- false negative (FN): The model classifies the example as negative, **but** the label is actually positive.
We will count the number of TP, FP, TN and FN in the given data. All of our metrics can be built off of these four statistics.
Recall that the model outputs real numbers between 0 and 1.
* To compute binary class predictions, we need to convert these to either 0 or 1.
* We'll do this using a threshold value $th$.
* Any model outputs above $th$ are set to 1, and below $th$ are set to 0.
**All of our metrics (except for AUC at the end) will depend on the choice of this threshold.**
Fill in the functions to compute the TP, FP, TN, and FN for a given threshold below.
The first one has been done for you.
```
# UNQ_C1 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
def true_positives(y, pred, th=0.5):
"""
Count true positives.
Args:
y (np.array): ground truth, size (n_examples)
pred (np.array): model output, size (n_examples)
th (float): cutoff value for positive prediction from model
Returns:
TP (int): true positives
"""
TP = 0
# get thresholded predictions
thresholded_preds = pred >= th
# compute TP
TP = np.sum((y == 1) & (thresholded_preds == 1))
return TP
def true_negatives(y, pred, th=0.5):
"""
Count true negatives.
Args:
y (np.array): ground truth, size (n_examples)
pred (np.array): model output, size (n_examples)
th (float): cutoff value for positive prediction from model
Returns:
TN (int): true negatives
"""
TN = 0
# get thresholded predictions
thresholded_preds = pred >= th
### START CODE HERE (REPLACE INSTANCES OF 'None' with your code) ###
# compute TN
TN = np.sum((y == 0) & (thresholded_preds == 0))
### END CODE HERE ###
return TN
def false_positives(y, pred, th=0.5):
"""
Count false positives.
Args:
y (np.array): ground truth, size (n_examples)
pred (np.array): model output, size (n_examples)
th (float): cutoff value for positive prediction from model
Returns:
FP (int): false positives
"""
FP = 0
# get thresholded predictions
thresholded_preds = pred >= th
### START CODE HERE (REPLACE INSTANCES OF 'None' with your code) ###
# compute FP
FP = np.sum((y == 0) & (thresholded_preds == 1))
### END CODE HERE ###
return FP
def false_negatives(y, pred, th=0.5):
"""
Count false positives.
Args:
y (np.array): ground truth, size (n_examples)
pred (np.array): model output, size (n_examples)
th (float): cutoff value for positive prediction from model
Returns:
FN (int): false negatives
"""
FN = 0
# get thresholded predictions
thresholded_preds = pred >= th
### START CODE HERE (REPLACE INSTANCES OF 'None' with your code) ###
# compute FN
FN = np.sum((y == 1) & (thresholded_preds == 0))
### END CODE HERE ###
return FN
# Note: we must explicity import 'display' in order for the autograder to compile the submitted code
# Even though we could use this function without importing it, keep this import in order to allow the grader to work
from IPython.display import display
# Test
df = pd.DataFrame({'y_test': [1,1,0,0,0,0,0,0,0,1,1,1,1,1],
'preds_test': [0.8,0.7,0.4,0.3,0.2,0.5,0.6,0.7,0.8,0.1,0.2,0.3,0.4,0],
'category': ['TP','TP','TN','TN','TN','FP','FP','FP','FP','FN','FN','FN','FN','FN']
})
display(df)
#y_test = np.array([1, 0, 0, 1, 1])
y_test = df['y_test']
#preds_test = np.array([0.8, 0.8, 0.4, 0.6, 0.3])
preds_test = df['preds_test']
threshold = 0.5
print(f"threshold: {threshold}\n")
print(f"""Our functions calcualted:
TP: {true_positives(y_test, preds_test, threshold)}
TN: {true_negatives(y_test, preds_test, threshold)}
FP: {false_positives(y_test, preds_test, threshold)}
FN: {false_negatives(y_test, preds_test, threshold)}
""")
print("Expected results")
print(f"There are {sum(df['category'] == 'TP')} TP")
print(f"There are {sum(df['category'] == 'TN')} TN")
print(f"There are {sum(df['category'] == 'FP')} FP")
print(f"There are {sum(df['category'] == 'FN')} FN")
```
Run the next cell to see a summary of evaluative metrics for the model predictions for each class.
```
util.get_performance_metrics(y, pred, class_labels)
```
Right now it only has TP, TN, FP, FN. Throughout this assignment we'll fill in all the other metrics to learn more about our model performance.
<a name='3-2'></a>
### 3.2 Accuracy
Let's use a threshold of .5 for the probability cutoff for our predictions for all classes and calculate our model's accuracy as we would normally do in a machine learning problem.
$$accuracy = \frac{\text{true positives} + \text{true negatives}}{\text{true positives} + \text{true negatives} + \text{false positives} + \text{false negatives}}$$
Use this formula to compute accuracy below:
<details>
<summary>
<font size="3" color="darkgreen"><b>Hints</b></font>
</summary>
<p>
<ul>
<li>Remember to set the value for the threshold when calling the functions.</li>
</ul>
</p>
```
# UNQ_C2 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
def get_accuracy(y, pred, th=0.5):
"""
Compute accuracy of predictions at threshold.
Args:
y (np.array): ground truth, size (n_examples)
pred (np.array): model output, size (n_examples)
th (float): cutoff value for positive prediction from model
Returns:
accuracy (float): accuracy of predictions at threshold
"""
accuracy = 0.0
### START CODE HERE (REPLACE INSTANCES OF 'None' with your code) ###
# get TP, FP, TN, FN using our previously defined functions
TP = true_positives(y, pred, th)
FP = false_positives(y, pred, th)
TN = true_negatives(y, pred, th)
FN = false_negatives(y, pred, th)
# Compute accuracy using TP, FP, TN, FN
accuracy = (TP + TN) / (TP+ TN + FP + FN)
### END CODE HERE ###
return accuracy
# Test
print("Test case:")
y_test = np.array([1, 0, 0, 1, 1])
print('test labels: {y_test}')
preds_test = np.array([0.8, 0.8, 0.4, 0.6, 0.3])
print(f'test predictions: {preds_test}')
threshold = 0.5
print(f"threshold: {threshold}")
print(f"computed accuracy: {get_accuracy(y_test, preds_test, threshold)}")
```
#### Expected output:
```Python
test labels: {y_test}
test predictions: [0.8 0.8 0.4 0.6 0.3]
threshold: 0.5
computed accuracy: 0.6
```
Run the next cell to see the accuracy of the model output for each class, as well as the number of true positives, true negatives, false positives, and false negatives.
```
util.get_performance_metrics(y, pred, class_labels, acc=get_accuracy)
```
If we were to judge our model's performance based on the accuracy metric, we would say that our model is not very accurate for detecting the `Infiltration` cases (accuracy of 0.657) but pretty accurate for detecting `Emphysema` (accuracy of 0.889).
**But is that really the case?...**
Let's imagine a model that simply predicts that **none** of the patients have `Emphysema`, regardless of patient's measurements. Let's calculate the accuracy for such a model.
```
get_accuracy(valid_results["Emphysema"].values, np.zeros(len(valid_results)))
```
As you can see above, such a model would be 97% accurate! Even better than our deep learning based model.
But is this really a good model? Wouldn't this model be wrong 100% of the time if the patient actually had this condition?
In the following sections, we will address this concern with more advanced model measures - **sensitivity and specificity** - that evaluate how well the model predicts positives for patients with the condition and negatives for cases that actually do not have the condition.
<a name='3-3'></a>
### 3.3 Prevalence
Another important concept is **prevalence**.
* In a medical context, prevalence is the proportion of people in the population who have the disease (or condition, etc).
* In machine learning terms, this is the proportion of positive examples. The expression for prevalence is:
$$prevalence = \frac{1}{N} \sum_{i} y_i$$
where $y_i = 1$ when the example is 'positive' (has the disease).
Let's measure prevalence for each disease:
<details>
<summary>
<font size="3" color="darkgreen"><b>Hints</b></font>
</summary>
<p>
<ul>
<li>
You can use <a href="https://docs.scipy.org/doc/numpy/reference/generated/numpy.mean.html" > np.mean </a> to calculate the formula.</li>
<li>Actually, the automatic grader is expecting numpy.mean, so please use it instead of using an equally valid but different way of calculating the prevalence. =) </li>
</ul>
</p>
```
# UNQ_C3 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
def get_prevalence(y):
"""
Compute accuracy of predictions at threshold.
Args:
y (np.array): ground truth, size (n_examples)
Returns:
prevalence (float): prevalence of positive cases
"""
prevalence = 0.0
### START CODE HERE (REPLACE INSTANCES OF 'None' with your code) ###
prevalence = np.mean(y==1)
### END CODE HERE ###
return prevalence
# Test
print("Test case:\n")
y_test = np.array([1, 0, 0, 1, 1, 0, 0, 0, 0, 1])
print(f'test labels: {y_test}')
print(f"computed prevalence: {get_prevalence(y_test)}")
util.get_performance_metrics(y, pred, class_labels, acc=get_accuracy, prevalence=get_prevalence)
```
`Hernia` has a prevalence 0.002, which is the rarest among the studied conditions in our dataset.
<a name='3-4'></a>
### 3.4 Sensitivity and Specificity
<img src="sens_spec.png" width="30%">
Sensitivity and specificity are two of the most prominent numbers that are used to measure diagnostics tests.
- Sensitivity is the probability that our test outputs positive given that the case is actually positive.
- Specificity is the probability that the test outputs negative given that the case is actually negative.
We can phrase this easily in terms of true positives, true negatives, false positives, and false negatives:
$$sensitivity = \frac{\text{true positives}}{\text{true positives} + \text{false negatives}}$$
$$specificity = \frac{\text{true negatives}}{\text{true negatives} + \text{false positives}}$$
Let's calculate sensitivity and specificity for our model:
```
# UNQ_C4 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
def get_sensitivity(y, pred, th=0.5):
"""
Compute sensitivity of predictions at threshold.
Args:
y (np.array): ground truth, size (n_examples)
pred (np.array): model output, size (n_examples)
th (float): cutoff value for positive prediction from model
Returns:
sensitivity (float): probability that our test outputs positive given that the case is actually positive
"""
sensitivity = 0.0
### START CODE HERE (REPLACE INSTANCES OF 'None' with your code) ###
# get TP and FN using our previously defined functions
TP = true_positives(y, pred, th)
FN = false_negatives(y, pred, th)
# use TP and FN to compute sensitivity
sensitivity = TP / (TP + FN)
### END CODE HERE ###
return sensitivity
def get_specificity(y, pred, th=0.5):
"""
Compute specificity of predictions at threshold.
Args:
y (np.array): ground truth, size (n_examples)
pred (np.array): model output, size (n_examples)
th (float): cutoff value for positive prediction from model
Returns:
specificity (float): probability that the test outputs negative given that the case is actually negative
"""
specificity = 0.0
### START CODE HERE (REPLACE INSTANCES OF 'None' with your code) ###
# get TN and FP using our previously defined functions
TN = true_negatives(y, pred, th)
FP = false_positives(y, pred, th)
# use TN and FP to compute specificity
specificity = TN / (TN + FP)
### END CODE HERE ###
return specificity
# Test
print("Test case")
y_test = np.array([1, 0, 0, 1, 1])
print(f'test labels: {y_test}\n')
preds_test = np.array([0.8, 0.8, 0.4, 0.6, 0.3])
print(f'test predictions: {preds_test}\n')
threshold = 0.5
print(f"threshold: {threshold}\n")
print(f"computed sensitivity: {get_sensitivity(y_test, preds_test, threshold):.2f}")
print(f"computed specificity: {get_specificity(y_test, preds_test, threshold):.2f}")
```
#### Expected output:
```Python
Test case
test labels: [1 0 0 1 1]
test predictions: [0.8 0.8 0.4 0.6 0.3]
threshold: 0.5
computed sensitivity: 0.67
computed specificity: 0.50
```
```
util.get_performance_metrics(y, pred, class_labels, acc=get_accuracy, prevalence=get_prevalence,
sens=get_sensitivity, spec=get_specificity)
```
Note that specificity and sensitivity do not depend on the prevalence of the positive class in the dataset.
* This is because the statistics are only computed within people of the same class
* Sensitivity only considers output on people in the positive class
* Similarly, specificity only considers output on people in the negative class.
<a name='3-5'></a>
### 3.5 PPV and NPV
Diagnostically, however, sensitivity and specificity are not helpful. Sensitivity, for example, tells us the probability our test outputs positive given that the person already has the condition. Here, we are conditioning on the thing we would like to find out (whether the patient has the condition)!
What would be more helpful is the probability that the person has the disease given that our test outputs positive. That brings us to positive predictive value (PPV) and negative predictive value (NPV).
- Positive predictive value (PPV) is the probability that subjects with a positive screening test truly have the disease.
- Negative predictive value (NPV) is the probability that subjects with a negative screening test truly don't have the disease.
Again, we can formulate these in terms of true positives, true negatives, false positives, and false negatives:
$$PPV = \frac{\text{true positives}}{\text{true positives} + \text{false positives}}$$
$$NPV = \frac{\text{true negatives}}{\text{true negatives} + \text{false negatives}}$$
NOTE: FOR COMPARISON PURPOSE
Sensitivity and specificity are two of the most prominent numbers that are used to measure diagnostics tests.
- Sensitivity is the probability that our test outputs positive given that the case is actually positive.
- Specificity is the probability that the test outputs negative given that the case is actually negative.
Let's calculate PPV & NPV for our model:
```
# UNQ_C5 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
def get_ppv(y, pred, th=0.5):
"""
Compute PPV of predictions at threshold.
Args:
y (np.array): ground truth, size (n_examples)
pred (np.array): model output, size (n_examples)
th (float): cutoff value for positive prediction from model
Returns:
PPV (float): positive predictive value of predictions at threshold
"""
PPV = 0.0
### START CODE HERE (REPLACE INSTANCES OF 'None' with your code) ###
# get TP and FP using our previously defined functions
TP = true_positives(y, pred, th)
FP = false_positives(y, pred, th)
# use TP and FP to compute PPV
PPV = TP / (TP + FP)
### END CODE HERE ###
return PPV
def get_npv(y, pred, th=0.5):
"""
Compute NPV of predictions at threshold.
Args:
y (np.array): ground truth, size (n_examples)
pred (np.array): model output, size (n_examples)
th (float): cutoff value for positive prediction from model
Returns:
NPV (float): negative predictive value of predictions at threshold
"""
NPV = 0.0
### START CODE HERE (REPLACE INSTANCES OF 'None' with your code) ###
# get TN and FN using our previously defined functions
TN = true_negatives(y, pred, th)
FN = false_negatives(y, pred, th)
# use TN and FN to compute NPV
NPV = TN / (TN + FN)
### END CODE HERE ###
return NPV
# Test
print("Test case:\n")
y_test = np.array([1, 0, 0, 1, 1])
print(f'test labels: {y_test}')
preds_test = np.array([0.8, 0.8, 0.4, 0.6, 0.3])
print(f'test predictions: {preds_test}\n')
threshold = 0.5
print(f"threshold: {threshold}\n")
print(f"computed ppv: {get_ppv(y_test, preds_test, threshold):.2f}")
print(f"computed npv: {get_npv(y_test, preds_test, threshold):.2f}")
```
#### Expected output:
```Python
Test case:
test labels: [1 0 0 1 1]
test predictions: [0.8 0.8 0.4 0.6 0.3]
threshold: 0.5
computed ppv: 0.67
computed npv: 0.50
```
```
util.get_performance_metrics(y, pred, class_labels, acc=get_accuracy, prevalence=get_prevalence,
sens=get_sensitivity, spec=get_specificity, ppv=get_ppv, npv=get_npv)
```
Notice that despite having very high sensitivity and accuracy, the PPV of the predictions could still be very low.
This is the case with `Edema`, for example.
* The sensitivity for `Edema` is 0.75.
* However, given that the model predicted positive, the probability that a person has Edema (its PPV) is only 0.066!
<a name='3-6'></a>
### 3.6 ROC Curve
So far we have been operating under the assumption that our model's prediction of `0.5` and above should be treated as positive and otherwise it should be treated as negative. This however was a rather arbitrary choice. One way to see this, is to look at a very informative visualization called the receiver operating characteristic (ROC) curve.
The ROC curve is created by plotting the true positive rate (TPR) against the false positive rate (FPR) at various threshold settings. The ideal point is at the top left, with a true positive rate of 1 and a false positive rate of 0. The various points on the curve are generated by gradually changing the threshold.
Let's look at this curve for our model:
```
util.get_curve(y, pred, class_labels)
```
The area under the ROC curve is also called AUCROC or C-statistic and is a measure of goodness of fit. In medical literature this number also gives the probability that a randomly selected patient who experienced a condition had a higher risk score than a patient who had not experienced the event. This summarizes the model output across all thresholds, and provides a good sense of the discriminative power of a given model.
Let's use the `sklearn` metric function of `roc_auc_score` to add this score to our metrics table.
```
from sklearn.metrics import roc_auc_score
util.get_performance_metrics(y, pred, class_labels, acc=get_accuracy, prevalence=get_prevalence,
sens=get_sensitivity, spec=get_specificity, ppv=get_ppv, npv=get_npv, auc=roc_auc_score)
```
<a name='4'></a>
## 4. Confidence Intervals
Of course our dataset is only a sample of the real world, and our calculated values for all above metrics is an estimate of the real world values. It would be good to quantify this uncertainty due to the sampling of our dataset. We'll do this through the use of confidence intervals. A 95\% confidence interval for an estimate $\hat{s}$ of a parameter $s$ is an interval $I = (a, b)$ such that 95\% of the time when the experiment is run, the true value $s$ is contained in $I$. More concretely, if we were to run the experiment many times, then the fraction of those experiments for which $I$ contains the true parameter would tend towards 95\%.
While some estimates come with methods for computing the confidence interval analytically, more complicated statistics, such as the AUC for example, are difficult. For these we can use a method called the *bootstrap*. The bootstrap estimates the uncertainty by resampling the dataset with replacement. For each resampling $i$, we will get a new estimate, $\hat{s}_i$. We can then estimate the distribution of $\hat{s}$ by using the distribution of $\hat{s}_i$ for our bootstrap samples.
In the code below, we create bootstrap samples and compute sample AUCs from those samples. Note that we use stratified random sampling (sampling from the positive and negative classes separately) to make sure that members of each class are represented.
```
def bootstrap_auc(y, pred, classes, bootstraps = 100, fold_size = 1000):
statistics = np.zeros((len(classes), bootstraps))
for c in range(len(classes)):
df = pd.DataFrame(columns=['y', 'pred'])
df.loc[:, 'y'] = y[:, c]
df.loc[:, 'pred'] = pred[:, c]
# get positive examples for stratified sampling
df_pos = df[df.y == 1]
df_neg = df[df.y == 0]
prevalence = len(df_pos) / len(df)
for i in range(bootstraps):
# stratified sampling of positive and negative examples
pos_sample = df_pos.sample(n = int(fold_size * prevalence), replace=True)
neg_sample = df_neg.sample(n = int(fold_size * (1-prevalence)), replace=True)
y_sample = np.concatenate([pos_sample.y.values, neg_sample.y.values])
pred_sample = np.concatenate([pos_sample.pred.values, neg_sample.pred.values])
score = roc_auc_score(y_sample, pred_sample)
statistics[c][i] = score
return statistics
statistics = bootstrap_auc(y, pred, class_labels)
```
Now we can compute confidence intervals from the sample statistics that we computed.
```
util.print_confidence_intervals(class_labels, statistics)
```
As you can see, our confidence intervals are much wider for some classes than for others. Hernia, for example, has an interval around (0.30 - 0.98), indicating that we can't be certain it is better than chance (at 0.5).
<a name='5'></a>
## 5. Precision-Recall Curve
Precision-Recall is a useful measure of success of prediction when the classes are very imbalanced.
In information retrieval
- Precision is a measure of result relevancy and that is equivalent to our previously defined PPV.
- Recall is a measure of how many truly relevant results are returned and that is equivalent to our previously defined sensitivity measure.
The precision-recall curve (PRC) shows the trade-off between precision and recall for different thresholds. A high area under the curve represents both high recall and high precision, where high precision relates to a low false positive rate, and high recall relates to a low false negative rate.
High scores for both show that the classifier is returning accurate results (high precision), as well as returning a majority of all positive results (high recall).
Run the following cell to generate a PRC:
```
util.get_curve(y, pred, class_labels, curve='prc')
```
<a name='6'></a>
## 6. F1 Score
F1 score is the harmonic mean of the precision and recall, where an F1 score reaches its best value at 1 (perfect precision and recall) and worst at 0.
Again, we can simply use `sklearn`'s utility metric function of `f1_score` to add this measure to our performance table.
```
from sklearn.metrics import f1_score
util.get_performance_metrics(y, pred, class_labels, acc=get_accuracy, prevalence=get_prevalence,
sens=get_sensitivity, spec=get_specificity, ppv=get_ppv, npv=get_npv, auc=roc_auc_score,f1=f1_score)
```
<a name='7'></a>
## 7. Calibration
When performing classification we often want not only to predict the class label, but also obtain a probability of each label. This probability would ideally give us some kind of confidence on the prediction. In order to observe how our model's generated probabilities are aligned with the real probabilities, we can plot what's called a *calibration curve*.
In order to generate a calibration plot, we first bucketize our predictions to a fixed number of separate bins (e.g. 5) between 0 and 1. We then calculate a point for each bin: the x-value for each point is the mean for the probability that our model has assigned to these points and the y-value for each point fraction of true positives in that bin. We then plot these points in a linear plot. A well-calibrated model has a calibration curve that almost aligns with the y=x line.
The `sklearn` library has a utility `calibration_curve` for generating a calibration plot. Let's use it and take a look at our model's calibration:
```
from sklearn.calibration import calibration_curve
def plot_calibration_curve(y, pred):
plt.figure(figsize=(20, 20))
for i in range(len(class_labels)):
plt.subplot(4, 4, i + 1)
fraction_of_positives, mean_predicted_value = calibration_curve(y[:,i], pred[:,i], n_bins=20)
plt.plot([0, 1], [0, 1], linestyle='--')
plt.plot(mean_predicted_value, fraction_of_positives, marker='.')
plt.xlabel("Predicted Value")
plt.ylabel("Fraction of Positives")
plt.title(class_labels[i])
plt.tight_layout()
plt.show()
plot_calibration_curve(y, pred)
```
As the above plots show, for most predictions our model's calibration plot does not resemble a well calibrated plot. How can we fix that?...
Thankfully, there is a very useful method called [Platt scaling](https://en.wikipedia.org/wiki/Platt_scaling) which works by fitting a logistic regression model to our model's scores. To build this model, we will be using the training portion of our dataset to generate the linear model and then will use the model to calibrate the predictions for our test portion.
```
from sklearn.linear_model import LogisticRegression as LR
y_train = train_results[class_labels].values
pred_train = train_results[pred_labels].values
pred_calibrated = np.zeros_like(pred)
for i in range(len(class_labels)):
lr = LR(solver='liblinear', max_iter=10000)
lr.fit(pred_train[:, i].reshape(-1, 1), y_train[:, i])
pred_calibrated[:, i] = lr.predict_proba(pred[:, i].reshape(-1, 1))[:,1]
plot_calibration_curve(y[:,], pred_calibrated)
```
# That's it!
Congratulations! That was a lot of metrics to get familiarized with.
We hope that you feel a lot more confident in your understanding of medical diagnostic evaluation and test your models correctly in your future work :)
| github_jupyter |
```
%matplotlib inline
```
# Superimposed fundamental solitons
This examples demonstrates the generation of two-frequency soliton molecules,
using the forward model for the analytic signal [1,2], in `py-fmas` implemented
as :class:`FMAS`.
In particular, this example shows how soliton molecules are generated from two
initially superimposed fundamental solitons at distinctly different frequencies
[3]. The exmample reproduces the propagation scenario shown in Fig. S10 of the
supplementary material to [3].
References:
[1] Sh. Amiranashvili, A. Demircan, Hamiltonian structure of
propagation equations for ultrashort optical pulses, Phys. Rev. E 10
(2010) 013812, http://dx.doi.org/10.1103/PhysRevA.82.013812.
[2] Sh. Amiranashvili, A. Demircan, Ultrashort Optical Pulse Propagation in
terms of Analytic Signal, Adv. Opt. Tech. 2011 (2011) 989515,
http://dx.doi.org/10.1155/2011/989515.
[3] O. Melchert, S. Willms, S. Bose, A. Yulin, B. Roth, F. Mitschke, U.
Morgner, I. Babushkin, A. Demircan, Soliton Molecules with Two Frequencies,
Phys. Rev. Lett. 123 (2019) 243905,
https://doi.org/10.1103/PhysRevLett.123.243905.
.. codeauthor:: Oliver Melchert <melchert@iqo.uni-hannover.de>
```
import fmas
import numpy as np
from fmas.grid import Grid
from fmas.models import FMAS
from fmas.solver import IFM_RK4IP
from fmas.analytic_signal import AS
from fmas.tools import change_reference_frame, plot_evolution
from fmas.propagation_constant import PropConst
def define_beta_fun():
r"""Custom refractive index.
"""
p = np.poly1d((9.653881, -39.738626, 16.8848987, -2.745456)[::-1])
q = np.poly1d((1.000000, -9.496406, 4.2206250, -0.703437)[::-1])
n_idx = lambda w: p(w)/q(w) # (-)
c0 = 0.29979 # (micron/fs)
return lambda w: n_idx(w)*w/c0 # (1/micron)
def main():
t_max = 2000. # (fs)
t_num = 2**14 # (-)
z_max = 0.06e6 # (micron)
z_num = 25000 # (-)
z_skip = 50 # (-)
chi = 1.0 # (micron^2/W)
c0 = 0.29979 # (micron/fs)
# -- PROPAGATION CONSTANT
beta_fun = define_beta_fun()
pc = PropConst(beta_fun)
# -- INITIALIZE DATA-STRUCTURES AND ALGORITHMS
grid = Grid( t_max = t_max, t_num = t_num, z_max = z_max, z_num = z_num)
model = FMAS(w=grid.w, beta_w = beta_fun(grid.w), chi = chi )
solver = IFM_RK4IP( model.Lw, model.Nw, user_action = model.claw)
# -- PREPARE INITIAL CONDITION AND RUN SIMULATION
w01, t01, A01 = 1.178, 30.0, 0.0248892 # (rad/fs), (fs), (sqrt(W))
w02, t02, A02 = 2.909, 30.0, 0.0136676 # (rad/fs), (fs), (sqrt(W))
A_0t_fun = lambda t, A0, t0, w0: np.real(A0/np.cosh(t/t0)*np.exp(1j*w0*t))
E_0t = A_0t_fun(grid.t, A01, t01, w01) + A_0t_fun(grid.t, A02, t02, w02)
solver.set_initial_condition( grid.w, AS(E_0t).w_rep)
solver.propagate( z_range = z_max, n_steps = z_num, n_skip = z_skip)
# -- SHOW RESULTS IN MOVING FRAME OF REFERENCE
v0 = 0.0749641870819 # (micron/fs)
utz = change_reference_frame(solver.w, solver.z, solver.uwz, v0)
plot_evolution( solver.z, grid.t, utz, t_lim=(-100,150), w_lim=(0.3,3.8))
if __name__=='__main__':
main()
```
| github_jupyter |
# Candlestick Up/Down-gap side-by-side white lines
https://www.investopedia.com/terms/u/updown-gap-sidebyside-white-lines.asp
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import talib
import warnings
warnings.filterwarnings("ignore")
# yahoo finance is used to fetch data
import yfinance as yf
yf.pdr_override()
# input
symbol = 'AAPL'
start = '2018-01-01'
end = '2021-10-11'
# Read data
df = yf.download(symbol,start,end)
# View Columns
df.head()
```
## Candlestick with Up/Down-gap side-by-side white lines
```
from matplotlib import dates as mdates
import datetime as dt
dfc = df.copy()
dfc['VolumePositive'] = dfc['Open'] < dfc['Adj Close']
#dfc = dfc.dropna()
dfc = dfc.reset_index()
dfc['Date'] = pd.to_datetime(dfc['Date'])
dfc['Date'] = dfc['Date'].apply(mdates.date2num)
dfc.head()
from mplfinance.original_flavor import candlestick_ohlc
fig = plt.figure(figsize=(14,10))
ax = plt.subplot(2, 1, 1)
candlestick_ohlc(ax,dfc.values, width=0.5, colorup='g', colordown='r', alpha=1.0)
ax.xaxis_date()
ax.xaxis.set_major_formatter(mdates.DateFormatter('%d-%m-%Y'))
ax.grid(True, which='both')
ax.minorticks_on()
axv = ax.twinx()
colors = dfc.VolumePositive.map({True: 'g', False: 'r'})
axv.bar(dfc.Date, dfc['Volume'], color=colors, alpha=0.4)
axv.axes.yaxis.set_ticklabels([])
axv.set_ylim(0, 3*df.Volume.max())
ax.set_title('Stock '+ symbol +' Closing Price')
ax.set_ylabel('Price')
gap_side_by_side_white_lines = talib.CDLGAPSIDESIDEWHITE(df['Open'], df['High'], df['Low'], df['Close'])
gap_side_by_side_white_lines = gap_side_by_side_white_lines[gap_side_by_side_white_lines != 0]
df['gap_side_by_side_white_lines'] = talib.CDLGAPSIDESIDEWHITE(df['Open'], df['High'], df['Low'], df['Close'])
df.loc[df['gap_side_by_side_white_lines'] !=0]
df['Adj Close'].loc[df['gap_side_by_side_white_lines'] !=0]
df['Adj Close'].loc[df['gap_side_by_side_white_lines'] !=0].index
gap_side_by_side_white_lines
gap_side_by_side_white_lines.index
df
fig = plt.figure(figsize=(20,16))
ax = plt.subplot(2, 1, 1)
candlestick_ohlc(ax,dfc.values, width=0.5, colorup='g', colordown='r', alpha=1.0)
ax.xaxis_date()
ax.xaxis.set_major_formatter(mdates.DateFormatter('%d-%m-%Y'))
ax.grid(True, which='both')
ax.minorticks_on()
axv = ax.twinx()
ax.plot_date(df['Adj Close'].loc[df['gap_side_by_side_white_lines'] !=0].index, df['Adj Close'].loc[df['gap_side_by_side_white_lines'] !=0],
'or', # marker style 'o', color 'g'
fillstyle='none', # circle is not filled (with color)
ms=10.0)
colors = dfc.VolumePositive.map({True: 'g', False: 'r'})
axv.bar(dfc.Date, dfc['Volume'], color=colors, alpha=0.4)
axv.axes.yaxis.set_ticklabels([])
axv.set_ylim(0, 3*df.Volume.max())
ax.set_title('Stock '+ symbol +' Closing Price')
ax.set_ylabel('Price')
```
## Plot Certain dates
```
df = df['2019-11-01':'2019-12-01']
dfc = df.copy()
dfc['VolumePositive'] = dfc['Open'] < dfc['Adj Close']
#dfc = dfc.dropna()
dfc = dfc.reset_index()
dfc['Date'] = pd.to_datetime(dfc['Date'])
dfc['Date'] = dfc['Date'].apply(mdates.date2num)
dfc.head()
fig = plt.figure(figsize=(20,16))
ax = plt.subplot(2, 1, 1)
#ax.set_facecolor('lightyellow')
candlestick_ohlc(ax,dfc.values, width=0.5, colorup='g', colordown='r', alpha=1.0)
ax.xaxis_date()
ax.xaxis.set_major_formatter(mdates.DateFormatter('%d-%m-%Y'))
#ax.grid(True, which='both')
#ax.minorticks_on()
axv = ax.twinx()
ax.plot_date(df['Adj Close'].loc[df['gap_side_by_side_white_lines'] !=0].index, df['Adj Close'].loc[df['gap_side_by_side_white_lines'] !=0],
'3', # marker style 'o', color 'g'
fillstyle='none', # circle is not filled (with color)
ms=30.0)
colors = dfc.VolumePositive.map({True: 'g', False: 'r'})
axv.bar(dfc.Date, dfc['Volume'], color=colors, alpha=0.4)
axv.axes.yaxis.set_ticklabels([])
axv.set_ylim(0, 3*df.Volume.max())
ax.set_title('Stock '+ symbol +' Closing Price')
ax.set_ylabel('Price')
```
# Highlight Candlestick
```
from matplotlib.dates import date2num
from datetime import datetime
fig = plt.figure(figsize=(20,16))
ax = plt.subplot(2, 1, 1)
candlestick_ohlc(ax,dfc.values, width=0.5, colorup='g', colordown='r', alpha=1.0)
ax.xaxis_date()
ax.xaxis.set_major_formatter(mdates.DateFormatter('%d-%m-%Y'))
#ax.grid(True, which='both')
#ax.minorticks_on()
axv = ax.twinx()
ax.axvspan(date2num(datetime(2019,11,4)), date2num(datetime(2019,11,5)),
label="Up Gap Side-by-Side White Lines",color="green", alpha=0.3)
ax.axvspan(date2num(datetime(2019,11,6)), date2num(datetime(2019,11,7)),
label="Down Gap Side-by-Side White Lines",color="red", alpha=0.3)
ax.axvspan(date2num(datetime(2019,11,8)), date2num(datetime(2019,11,9)),
label="Up Gap Side-by-Side White Lines",color="green", alpha=0.3)
ax.legend(loc='upper center')
colors = dfc.VolumePositive.map({True: 'g', False: 'r'})
axv.bar(dfc.Date, dfc['Volume'], color=colors, alpha=0.4)
axv.axes.yaxis.set_ticklabels([])
axv.set_ylim(0, 3*df.Volume.max())
ax.set_title('Stock '+ symbol +' Closing Price')
ax.set_ylabel('Price')
```
| github_jupyter |
# MNIST handwritten digits anomaly detection
In this notebook, we'll test some anomaly detection methods to detect outliers within MNIST digits data using scikit-learn.
First, the needed imports.
```
%matplotlib inline
from pml_utils import get_mnist, show_anomalies
import numpy as np
from sklearn import __version__
from sklearn.ensemble import IsolationForest
from sklearn.neighbors import LocalOutlierFactor
import matplotlib.pyplot as plt
import seaborn as sns
sns.set()
from distutils.version import LooseVersion as LV
assert(LV(__version__) >= LV("0.20")), "Version >= 0.20 of sklearn is required."
```
Then we load the MNIST data. First time it downloads the data, which can take a while.
To speed up the computations, let's use only 10000 digits in this notebook.
```
X_train, y_train, X_test, y_test = get_mnist('MNIST')
X = X_train[:10000]
y = y_train[:10000]
print()
print('MNIST data loaded:')
print('X:', X.shape)
print('y:', y.shape)
```
Let us then create some outliers in our data. We
* invert all pixels of one sample
* shuffle all pixels of one sample, and
* add salt-and-pepper noise to 10% of pixels of one sample.
You can also continue creating more outliers in a similar fashion.
```
X[9999,:]=255-X[9999,:]
np.random.shuffle(X[9998,:])
for i in np.random.randint(0, X.shape[1], int(X.shape[1]*0.1)):
X[9997,i] = 0.0 if np.random.rand()<0.5 else 255.0
```
Let's have a look at our outliers:
```
n_outliers = 3
pltsize = 5
plt.figure(figsize=(n_outliers*pltsize, pltsize))
for i in range(n_outliers):
plt.subplot(1,10,i+1)
plt.axis('off')
plt.imshow(X[9999-i,:].reshape(28,28), cmap="gray")
```
## Isolation forest
[Isolation forest](https://scikit-learn.org/stable/modules/generated/sklearn.ensemble.IsolationForest.html#sklearn.ensemble.IsolationForest) is an outlier detection method based on using random forests. The idea is to isolate data items by random features and splits. Outliers are easier to isolate, so they tend to produce shorter paths on average.
We specify the number of trees as `n_estimators` and the assumed proportion of outliers in the data set as `if_contamination`.
```
%%time
n_estimators = 100
if_contamination = 0.001
if_model = IsolationForest(n_estimators=n_estimators,
contamination=if_contamination, behaviour='new')
if_pred = if_model.fit(X).predict(X)
print('Number of anomalies:', np.sum(if_pred==-1))
```
We use a function `show_anomalies` to take a look at the found outliers.
```
show_anomalies(if_pred, X)
```
## Local outlier factor
[Local outlier factor](https://scikit-learn.org/stable/modules/generated/sklearn.neighbors.LocalOutlierFactor.html#sklearn.neighbors.LocalOutlierFactor) is another method for outlier detection. It is based on k-nearest neighbors and computes the local density of data points with respect to their neighbors. Outliers have substantially lower local density than inliers.
We specify the number of neighbors considered as `n_neighbors` and the assumed proportion of outliers in the data set as `lof_contamination`.
```
%%time
n_neighbors = 20
lof_contamination = 0.001
lof_model = LocalOutlierFactor(n_neighbors=n_neighbors,
contamination=lof_contamination)
lof_pred = lof_model.fit_predict(X)
print('Number of anomalies:', np.sum(lof_pred==-1))
show_anomalies(lof_pred, X)
```
## Experiments
Experiment with different parameters for [isolation forest](https://scikit-learn.org/stable/modules/generated/sklearn.ensemble.IsolationForest.html#sklearn.ensemble.IsolationForest) and [local outlier factor](https://scikit-learn.org/stable/modules/generated/sklearn.neighbors.LocalOutlierFactor.html#sklearn.neighbors.LocalOutlierFactor). Are the algorithms able to find all the generated outliers?
You can also create more outliers in a similar fashion.
| github_jupyter |
# Finite Volume Discretisation
In this notebook, we explain the discretisation process that converts an expression tree, representing a model, to a linear algebra tree that can be evaluated by the solvers.
We use Finite Volumes as an example of a spatial method, since it is the default spatial method for most PyBaMM models. This is a good spatial method for battery problems as it is conservative: for lithium-ion battery models, we can be sure that the total amount of lithium in the system is constant. For more details on the Finite Volume method, see [Randall Leveque's book](https://books.google.co.uk/books/about/Finite_Volume_Methods_for_Hyperbolic_Pro.html?id=QazcnD7GUoUC&printsec=frontcover&source=kp_read_button&redir_esc=y#v=onepage&q&f=false).
This notebook is structured as follows:
1. **Setting up a discretisation**. Overview of the parameters that are passed to the discretisation
2. **Discretisations and spatial methods**. Operations that are common to most spatial methods:
- Discretising a spatial variable (e.g. $x$)
- Discretising a variable (e.g. concentration)
3. **Example: Finite Volume operators**. Finite Volume implementation of some useful operators:
- Gradient operator
- Divergence operator
- Integral operator
4. **Example: Discretising a simple model**. Setting up and solving a simple model, using Finite Volumes as the spatial method
To find out how to implement a new spatial method, see the [tutorial](https://pybamm.readthedocs.io/en/latest/tutorials/add-spatial-method.html) in the API docs.
## Setting up a Discretisation
We first import `pybamm` and some useful other modules, and change our working directory to the root of the `PyBaMM` folder:
```
%pip install pybamm -q # install PyBaMM if it is not installed
import pybamm
import numpy as np
import os
import matplotlib.pyplot as plt
from pprint import pprint
os.chdir(pybamm.__path__[0]+'/..')
```
To set up a discretisation, we must create a geometry, mesh this geometry, and then create the discretisation with the appropriate spatial method(s). The easiest way to create a geometry is to the inbuilt battery geometry:
```
parameter_values = pybamm.ParameterValues(
values={
"Negative electrode thickness [m]": 0.3,
"Separator thickness [m]": 0.2,
"Positive electrode thickness [m]": 0.3,
}
)
geometry = pybamm.battery_geometry()
parameter_values.process_geometry(geometry)
```
We then use this geometry to create a mesh, which for this example consists of uniform 1D submeshes
```
submesh_types = {
"negative electrode": pybamm.Uniform1DSubMesh,
"separator": pybamm.Uniform1DSubMesh,
"positive electrode": pybamm.Uniform1DSubMesh,
"negative particle": pybamm.Uniform1DSubMesh,
"positive particle": pybamm.Uniform1DSubMesh,
"current collector": pybamm.SubMesh0D,
}
var = pybamm.standard_spatial_vars
var_pts = {var.x_n: 15, var.x_s: 10, var.x_p: 15, var.r_n: 10, var.r_p: 10}
mesh = pybamm.Mesh(geometry, submesh_types, var_pts)
```
Finally, we can use the mesh to create a discretisation, using Finite Volumes as the spatial method for this example
```
spatial_methods = {
"macroscale": pybamm.FiniteVolume(),
"negative particle": pybamm.FiniteVolume(),
"positive particle": pybamm.FiniteVolume(),
}
disc = pybamm.Discretisation(mesh, spatial_methods)
```
## Discretisations and Spatial Methods
### Spatial Variables
Spatial variables, such as $x$ and $r$, are converted to `pybamm.Vector` nodes
```
# Set up
macroscale = ["negative electrode", "separator", "positive electrode"]
x_var = pybamm.SpatialVariable("x", domain=macroscale)
r_var = pybamm.SpatialVariable("r", domain=["negative particle"])
# Discretise
x_disc = disc.process_symbol(x_var)
r_disc = disc.process_symbol(r_var)
print("x_disc is a {}".format(type(x_disc)))
print("r_disc is a {}".format(type(r_disc)))
# Evaluate
x = x_disc.evaluate()
r = r_disc.evaluate()
f, (ax1, ax2) = plt.subplots(1, 2, figsize=(13,4))
ax1.plot(x, "*")
ax1.set_xlabel("index")
ax1.set_ylabel(r"$x$")
ax2.plot(r, "*")
ax2.set_xlabel("index")
ax2.set_ylabel(r"$r$")
plt.tight_layout()
plt.show()
```
We define `y_macroscale`, `y_microscale` and `y_scalar` for evaluation and visualisation of results below
```
y_macroscale = x ** 3 / 3
y_microscale = np.cos(r)
y_scalar = np.array([[5]])
y = np.concatenate([y_macroscale, y_microscale, y_scalar])
```
### Variables
In this notebook, we will work with three variables `u`, `v`, `w`.
```
u = pybamm.Variable("u", domain=macroscale) # u is a variable in the macroscale (e.g. electrolyte potential)
v = pybamm.Variable("v", domain=["negative particle"]) # v is a variable in the negative particle (e.g. particle concentration)
w = pybamm.Variable("w") # w is a variable without a domain (e.g. time, average concentration)
variables = [u,v,w]
```
Before discretising, trying to evaluate the variables raises a `NotImplementedError`:
```
try:
u.evaluate()
except NotImplementedError as e:
print(e)
```
For any spatial method, a `pybamm.Variable` gets converted to a `pybamm.StateVector` which, when evaluated, takes the appropriate slice of the input vector `y`.
```
# Pass the list of variables to the discretisation to calculate the slices to be used (order matters here!)
disc.set_variable_slices(variables)
# Discretise the variables
u_disc = disc.process_symbol(u)
v_disc = disc.process_symbol(v)
w_disc = disc.process_symbol(w)
# Print the outcome
print("Discretised u is the StateVector {}".format(u_disc))
print("Discretised v is the StateVector {}".format(v_disc))
print("Discretised w is the StateVector {}".format(w_disc))
```
Since the variables have been passed to `disc` in the order `[u,v,w]`, they each read the appropriate part of `y` when evaluated:
```
x_fine = np.linspace(x[0], x[-1], 1000)
r_fine = np.linspace(r[0], r[-1], 1000)
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(13,4))
ax1.plot(x_fine, x_fine**3/3, x, u_disc.evaluate(y=y), "o")
ax1.set_xlabel("x")
ax1.legend(["x^3/3", "u"], loc="best")
ax2.plot(r_fine, np.cos(r_fine), r, v_disc.evaluate(y=y), "o")
ax2.set_xlabel("r")
ax2.legend(["cos(r)", "v"], loc="best")
plt.tight_layout()
plt.show()
print("w = {}".format(w_disc.evaluate(y=y)))
```
## Finite Volume Operators
### Gradient operator
The gradient operator is converted to a Matrix-StateVector multiplication. In 1D, the gradient operator is equivalent to $\partial/\partial x$ on the macroscale and $\partial/\partial r$ on the microscale. In Finite Volumes, we take the gradient of an object on nodes (shape (n,)), which returns an object on the edges (shape (n-1,)).
```
grad_u = pybamm.grad(u)
grad_u_disc = disc.process_symbol(grad_u)
grad_u_disc.render()
```
The Matrix in `grad_u_disc` is the standard `[-1,1]` sparse matrix, divided by the step sizes `dx`:
```
macro_mesh = mesh.combine_submeshes(*macroscale)
print("gradient matrix is:\n")
print("1/dx *\n{}".format(macro_mesh.d_nodes[:,np.newaxis] * grad_u_disc.children[0].entries.toarray()))
```
When evaluated with `y_macroscale=x**3/3`, `grad_u_disc` is equal to `x**2` as expected:
```
x_edge = macro_mesh.edges[1:-1] # note that grad_u_disc is evaluated on the node edges
fig, ax = plt.subplots()
ax.plot(x_fine, x_fine**2, x_edge, grad_u_disc.evaluate(y=y), "o")
ax.set_xlabel("x")
legend = ax.legend(["x^2", "grad(u).evaluate(y=x**3/3)"], loc="best")
plt.show()
```
Similary, we can create, discretise and evaluate the gradient of `v`, which is a variable in the negative particles. Note that the syntax for doing this is identical: we do not need to explicitly specify that we want the gradient in `r`, since this is inferred from the `domain` of `v`.
```
v.domain
grad_v = pybamm.grad(v)
grad_v_disc = disc.process_symbol(grad_v)
print("grad(v) tree is:\n")
grad_v_disc.render()
micro_mesh = mesh["negative particle"]
print("\n gradient matrix is:\n")
print("1/dr *\n{}".format(micro_mesh.d_nodes[:,np.newaxis] * grad_v_disc.children[0].entries.toarray()))
r_edge = micro_mesh.edges[1:-1] # note that grad_u_disc is evaluated on the node edges
fig, ax = plt.subplots()
ax.plot(r_fine, -np.sin(r_fine), r_edge, grad_v_disc.evaluate(y=y), "o")
ax.set_xlabel("x")
legend = ax.legend(["-sin(r)", "grad(v).evaluate(y=cos(r))"], loc="best")
plt.show()
```
#### Boundary conditions
If the discretisation is provided with boundary conditions, appropriate ghost nodes are concatenated onto the variable, and a larger gradient matrix is used. The ghost nodes are chosen based on the value of the first/last node in the variable and the boundary condition.
For a Dirichlet boundary condition $u=a$ on the left-hand boundary, we set the value of the left ghost node to be equal to
$$2*a-u[0],$$
where $u[0]$ is the value of $u$ in the left-most cell in the domain. Similarly, for a Dirichlet condition $u=b$ on the right-hand boundary, we set the right ghost node to be
$$2*b-u[-1].$$
Note also that the size of the gradient matrix is now (41,42) instead of (39,40), to account for the presence of boundary conditions in the State Vector.
```
disc.bcs = {u.id: {"left": (pybamm.Scalar(1), "Dirichlet"), "right": (pybamm.Scalar(2), "Dirichlet")}}
grad_u_disc = disc.process_symbol(grad_u)
print("The gradient object is:")
(grad_u_disc.render())
u_eval = grad_u_disc.children[1].evaluate(y=y)
print("The value of u on the left-hand boundary is {}".format((u_eval[0] + u_eval[1]) / 2))
print("The value of u on the right-hand boundary is {}".format((u_eval[-2] + u_eval[-1]) / 2))
```
For a Neumann boundary condition $\partial u/\partial x=c$ on the left-hand boundary, we set the value of the left ghost node to be
$$u[0] - c * dx,$$
where $dx$ is the step size at the left-hand boundary. For a Neumann boundary condition $\partial u/\partial x=d$ on the right-hand boundary, we set the value of the right ghost node to be
$$u[-1] + d * dx.$$
```
disc.bcs = {u.id: {"left": (pybamm.Scalar(3), "Neumann"), "right": (pybamm.Scalar(4), "Neumann")}}
grad_u_disc = disc.process_symbol(grad_u)
print("The gradient object is:")
(grad_u_disc.render())
grad_u_eval = grad_u_disc.evaluate(y=y)
print("The gradient on the left-hand boundary is {}".format(grad_u_eval[0]))
print("The gradient of u on the right-hand boundary is {}".format(grad_u_eval[-1]))
```
We can mix the types of the boundary conditions:
```
disc.bcs = {u.id: {"left": (pybamm.Scalar(5), "Dirichlet"), "right": (pybamm.Scalar(6), "Neumann")}}
grad_u_disc = disc.process_symbol(grad_u)
print("The gradient object is:")
(grad_u_disc.render())
grad_u_eval = grad_u_disc.evaluate(y=y)
u_eval = grad_u_disc.children[1].evaluate(y=y)
print("The value of u on the left-hand boundary is {}".format((u_eval[0] + u_eval[1])/2))
print("The gradient on the right-hand boundary is {}".format(grad_u_eval[-1]))
```
Robin boundary conditions can be implemented by specifying a Neumann condition where the flux depends on the variable.
### Divergence operator
Before computing the Divergence operator, we set up Neumann boundary conditions. The behaviour with Dirichlet boundary conditions is very similar.
```
disc.bcs = {u.id: {"left": (pybamm.Scalar(-1), "Neumann"), "right": (pybamm.Scalar(1), "Neumann")}}
```
Now we can process `div(grad(u))`, converting it to a Matrix-Vector multiplication, plus a vector for the boundary conditions. Since we have Neumann boundary conditions, the divergence of an object of size (n+1,) has size (n,), and so div(grad) of an object of size (n,) has size (n,)
```
div_grad_u = pybamm.div(grad_u)
div_grad_u_disc = disc.process_symbol(div_grad_u)
div_grad_u_disc.render()
```
The div(grad) matrix is automatically simplified to the well-known `[1,-2,1]` matrix (divided by the square of the distance between the edges), except in the first and last rows for boundary conditions
```
print("div(grad) matrix is:\n")
print("1/dx^2 * \n{}".format(
macro_mesh.d_edges[:,np.newaxis]**2 * div_grad_u_disc.left.left.entries.toarray()
))
```
### Integral operator
Finally, we can define an integral operator, which integrates the variable across the domain specified by the integration variable.
```
int_u = pybamm.Integral(u, x_var)
int_u_disc = disc.process_symbol(int_u)
print("int(u) = {} is approximately equal to 1/12, {}".format(int_u_disc.evaluate(y=y), 1/12))
# We divide v by r to evaluate the integral more easily
int_v_over_r2 = pybamm.Integral(v/r_var**2, r_var)
int_v_over_r2_disc = disc.process_symbol(int_v_over_r2)
print("int(v/r^2) = {} is approximately equal to 4 * pi * sin(1), {}".format(
int_v_over_r2_disc.evaluate(y=y), 4 * np.pi * np.sin(1))
)
```
The integral operators are also Matrix-Vector multiplications
```
print("int(u):\n")
int_u_disc.render()
print("\nint(v):\n")
int_v_over_r2_disc.render()
int_u_disc.children[0].evaluate() / macro_mesh.d_edges
int_v_over_r2_disc.children[0].evaluate() / micro_mesh.d_edges
```
## Discretising a model
We can now discretise a whole model. We create, and discretise, a simple model for the concentration in the electrolyte and the concentration in the particles, and discretise it with a single command:
```
disc.process_model(model)
```
```
model = pybamm.BaseModel()
c_e = pybamm.Variable("electrolyte concentration", domain=macroscale)
N_e = pybamm.grad(c_e)
c_s = pybamm.Variable("particle concentration", domain=["negative particle"])
N_s = pybamm.grad(c_s)
model.rhs = {c_e: pybamm.div(N_e) - 5, c_s: pybamm.div(N_s)}
model.boundary_conditions = {
c_e: {"left": (np.cos(0), "Neumann"), "right": (np.cos(10), "Neumann")},
c_s: {"left": (0, "Neumann"), "right": (-1, "Neumann")},
}
model.initial_conditions = {c_e: 1 + 0.1 * pybamm.sin(10*x_var), c_s: 1}
# Create a new discretisation and process model
disc2 = pybamm.Discretisation(mesh, spatial_methods)
disc2.process_model(model);
```
The initial conditions are discretised to vectors, and an array of concatenated initial conditions is created.
```
c_e_0 = model.initial_conditions[c_e].evaluate()
c_s_0 = model.initial_conditions[c_s].evaluate()
y0 = model.concatenated_initial_conditions.evaluate()
fig, (ax1, ax2, ax3) = plt.subplots(1, 3, figsize=(13,4))
ax1.plot(x_fine, 1 + 0.1*np.sin(10*x_fine), x, c_e_0, "o")
ax1.set_xlabel("x")
ax1.legend(["1+0.1*sin(10*x)", "c_e_0"], loc="best")
ax2.plot(x_fine, np.ones_like(r_fine), r, c_s_0, "o")
ax2.set_xlabel("r")
ax2.legend(["1", "c_s_0"], loc="best")
ax3.plot(y0,"*")
ax3.set_xlabel("index")
ax3.set_ylabel("y0")
plt.tight_layout()
plt.show()
```
The discretised rhs can be evaluated, for example at `0,y0`:
```
rhs_c_e = model.rhs[c_e].evaluate(0, y0)
rhs_c_s = model.rhs[c_s].evaluate(0, y0)
rhs = model.concatenated_rhs.evaluate(0, y0)
fig, (ax1, ax2, ax3) = plt.subplots(1, 3, figsize=(13,4))
ax1.plot(x_fine, -10*np.sin(10*x_fine) - 5, x, rhs_c_e, "o")
ax1.set_xlabel("x")
ax1.set_ylabel("rhs_c_e")
ax1.legend(["1+0.1*sin(10*x)", "c_e_0"], loc="best")
ax2.plot(r, rhs_c_s, "o")
ax2.set_xlabel("r")
ax2.set_ylabel("rhs_c_s")
ax3.plot(rhs,"*")
ax3.set_xlabel("index")
ax3.set_ylabel("rhs")
plt.tight_layout()
plt.show()
```
The function `model.concatenated_rhs` is then passed to the solver to solve the model, with initial conditions `model.concatenated_initial_conditions`.
## Upwinding and downwinding
If a system is advection-dominated (Peclet number greater than around 40), then it is important to use upwinding (if velocity is positive) or downwinding (if velocity is negative) to obtain accurate results. To see this, consider the following model (without upwinding)
```
model = pybamm.BaseModel()
model.length_scales = {
"negative electrode": pybamm.Scalar(1),
"separator": pybamm.Scalar(1),
"positive electrode": pybamm.Scalar(1)
}
# Define concentration and velocity
c = pybamm.Variable("c", domain=["negative electrode", "separator", "positive electrode"])
v = pybamm.PrimaryBroadcastToEdges(1, ["negative electrode", "separator", "positive electrode"])
model.rhs = {c: -pybamm.div(c * v) + 1}
model.initial_conditions = {c: 0}
model.boundary_conditions = {c: {"left": (0, "Dirichlet")}}
model.variables = {"c": c}
def solve_and_plot(model):
model_disc = disc.process_model(model, inplace=False)
t_eval = [0,100]
solution = pybamm.CasadiSolver().solve(model_disc, t_eval)
# plot
plot = pybamm.QuickPlot(solution,["c"],spatial_unit="m")
plot.dynamic_plot()
solve_and_plot(model)
```
The concentration grows indefinitely, which is clearly an incorrect solution. Instead, we can use upwinding:
```
model.rhs = {c: -pybamm.div(pybamm.upwind(c) * v) + 1}
solve_and_plot(model)
```
This gives the expected linear steady state from 0 to 1. Similarly, if the velocity is negative, downwinding gives accurate results
```
model.rhs = {c: -pybamm.div(pybamm.downwind(c) * (-v)) + 1}
model.boundary_conditions = {c: {"right": (0, "Dirichlet")}}
solve_and_plot(model)
```
## More advanced concepts
Since this notebook is only an introduction to the discretisation, we have not covered everything. More advanced concepts, such as the ones below, can be explored by looking into the [API docs](https://pybamm.readthedocs.io/en/latest/source/spatial_methods/finite_volume.html).
- Gradient and divergence of microscale variables in the P2D model
- Indefinite integral
If you would like detailed examples of these operations, please [create an issue](https://github.com/pybamm-team/PyBaMM/blob/develop/CONTRIBUTING.md#a-before-you-begin) and we will be happy to help.
## References
The relevant papers for this notebook are:
```
pybamm.print_citations()
```
| github_jupyter |
# Extracting waterbodies from Sentinel-2 <img align="right" src="../Supplementary_data/dea_logo.jpg">
* [**Sign up to the DEA Sandbox**](https://docs.dea.ga.gov.au/setup/sandbox.html) to run this notebook interactively from a browser
* **Compatibility:** Notebook currently compatible with both the `NCI` and `DEA Sandbox` environments
* **Products used:**
[s2a_ard_granule](https://explorer.sandbox.dea.ga.gov.au/s2a_ard_granule),
[s2b_ard_granule](https://explorer.sandbox.dea.ga.gov.au/s2b_ard_granule),
[ga_ls5t_ard_3](https://explorer.sandbox.dea.ga.gov.au/ga_ls5t_ard_3),
[ga_ls7e_ard_3](https://explorer.sandbox.dea.ga.gov.au/ga_ls7e_ard_3),
[ga_ls8c_ard_3](https://explorer.sandbox.dea.ga.gov.au/ga_ls8c_ard_3),
[wofs_albers](https://explorer.sandbox.dea.ga.gov.au/wofs_albers)
## Background
[DEA Waterbodies](https://www.ga.gov.au/dea/products/dea-waterbodies) is a time-series data product that summarises the surface area of open waterbodies in Australia using [Water Observations from Space (WOfS)](https://www.ga.gov.au/dea/products/wofs).
WOfS classifies Landsat pixels into wet and dry. Landsat data have a resolution of 25m$^2$, but it would be really nice if we could instead use Sentinel 2 data with its 10m$^2$ resolution.
This would help distinguish neighbouring waterbodies that are blurred together in Landsat (and hence in DEA Waterbodies).
Sentinel 2 does not yet have WOfS.
One alternative to WOfS that we _could_ evaluate for Sentinel 2 is the [Modified Normalised Difference Water Idex (MNDWI)](http://doi.org/10.1080/01431160600589179), which can be calculated directly from surface reflectance.
MNDWI greater than zero is indicative of water. It is not as accurate as WOfS, but easier to obtain.
## Description
This notebook compares DEA Waterbodies derived from WOfS to an analogous product derived from MNDWI.
There are two main factors we can vary: how to derive our polygons, and how to derive our time series.
Both polygons and time series can be generated from Landsat WOfS, from Landsat MNDWI, or from Sentinel 2 MNDWI.
We can choose to combine any polygon method and any time series method.
This gives us a grid of possible evaluations:
| | Landsat WOfS polygons | Landsat MNDWI polygons | Sentinel 2 MNDWI polygons |
|-|---------------|------------------------|---------------------------|
|**Landsat WOfS time series** | DEA Waterbodies | Polygon proxy quality | Good differentation for merged waterbodies |
|**Landsat MNDWI time series** | Time series proxy quality | Total proxy quality | - |
|**Sentinel 2 MNDWI time series** | Sentinel 2 proxy quality | - | Highest resolution |
The upper left corner is the existing DEA Waterbodies.
Polygons derived from Sentinel 2 would be great for differentiating neighbouring waterbodies, even with lower resolution surface area data.
Landsat WOfS polygons with MNDWI time series informs us how well MNDWI approximates the surface area time series.
Similarly, Landsat MNDWI polygons with WOfS time series informs us how well the MNDWI polygons approximate those we obtain from WOfS.
The highest attainable resolution is deriving both parts of the product from Sentinel 2.
***
## Getting started
Edit the analysis parameters and run all the cells in this notebook to analyse a region of Australia.
### Load packages
Import Python packages that are used for the analysis.
```
import sys
import datacube
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import xarray as xr
from datacube.storage import masking
import datacube.virtual as virtual
import geopandas as gpd
sys.path.append("../Scripts")
from dea_plotting import display_map
from dea_spatialtools import xr_rasterize, xr_vectorize
from dea_dask import create_local_dask_cluster
from dea_datahandling import wofs_fuser
```
### Connect to the datacube
Connect to the datacube so we can access DEA data.
The `app` parameter is a unique name for the analysis which is based on the notebook file name.
```
dc = datacube.Datacube(app="Sentinel_2_waterbodies")
```
### Set up Dask
Set up a local Dask server for parallelism and lazy loading.
```
create_local_dask_cluster()
```
### Analysis parameters
Choose which area of Australia to analyse:
```
# Lake Boort and Lake Lyndger
xlim = (143.70141, 143.76180)
ylim = (-36.14517, -36.09688)
```
### Check the analysis area
Display a map of the area we plan to analyse.
```
display_map(x=xlim, y=ylim)
```
## Build a virtual product for the MNDWI calculation
The [MNDWI](DOI:10.1080/01431160600589179) is given by
$$
\frac{\mathrm{green} - \mathrm{SWIR}}{\mathrm{green} + \mathrm{SWIR}}.
$$
We'll build a virtual product that calculates this without having to load all of the channels ourselves.
A virtual product is just like a regular DEA product, except it has some transformations applied before we see it.
We can define one with YAML that applies cloud masking and calculates the MNDWI.
> **Note:** For more information about virtual products, refer to the [Virtual products notebook](../Frequently_used_code/Virtual_products.ipynb).
```
cat_yaml = """
products:
ls8_MNDWI:
recipe:
transform: expressions
output:
MNDWI:
formula: (green - swir1) / (green + swir1)
dtype: float32
input:
transform: apply_mask
mask_measurement_name: fmask
input:
transform: expressions
output:
fmask:
formula: (fmask == 1) | (fmask == 5)
nodata: False
green: green
swir1: swir1
input:
product: ga_ls8c_ard_3
measurements: [green, swir1, fmask]
output_crs: EPSG:3577
resolution: [-30, 30]
resampling:
fmask: nearest
'*': average
s2a_MNDWI:
recipe:
transform: expressions
output:
MNDWI:
formula: (nbart_green - nbart_swir_2) / (nbart_green + nbart_swir_2)
dtype: float32
input:
transform: apply_mask
mask_measurement_name: fmask
input:
transform: expressions
output:
fmask:
formula: (fmask == 1) | (fmask == 5)
nodata: False
nbart_green: nbart_green
nbart_swir_2: nbart_swir_2
input:
collate:
- product: s2b_ard_granule
measurements: [nbart_green, nbart_swir_2, fmask]
output_crs: EPSG:3577
resolution: [-10, 10]
resampling:
fmask: nearest
'*': average
- product: s2a_ard_granule
measurements: [nbart_green, nbart_swir_2, fmask]
output_crs: EPSG:3577
resolution: [-10, 10]
resampling:
fmask: nearest
'*': average
wofs_masked:
recipe:
transform: expressions
output:
water:
formula: water
nodata: -1
input:
transform: apply_mask
mask_measurement_name: mask
input:
transform: expressions
output:
mask:
formula: (water == 0) | (water == 128)
nodata: False
water:
formula: water
nodata: -1
input:
product: wofs_albers
measurements: [water]
output_crs: EPSG:3577
resolution: [-25, 25]
"""
```
Then we can convert the YAML into a catalogue containing our custom virtual products, which we can use to load data just like a datacube:
```
cat = virtual.catalog_from_yaml(cat_yaml)
```
Now we can use this catalogue to load our MNDWI virtual products for Sentinel 2 and Landsat 8.
Use dask to avoid loading everything into memory at once if the data are too big, or otherwise use `.load` here to load the data immediately.
> **Note:** This step can take **several minutes to load**.
```
# Set up spatio-temporal query used to load data for both products
query = {
"x": xlim,
"y": ylim,
"time": ("2016-01", "2018-12"),
"group_by": "solar_day",
"dask_chunks": {"x": 3000, "y": 3000, "time": 1},
}
# Load Sentinel-2 MNDWI data
s2a_mndwi = cat["s2a_MNDWI"].load(dc, **query).load()
# Load Landsat 8 MNDWI data
ls8_mndwi = cat["ls8_MNDWI"].load(dc, **query).load()
```
Finally, grab WOfS for the same time period as our Landsat observations.
```
# Load WOfS data for the same spatio-temporal query
wofs = cat["wofs_masked"].load(dc, fuse_func=wofs_fuser, **query).load()
# Set nodata to `NaN`, and wet pixels (value 128) to 1
wofs = masking.mask_invalid_data(wofs)
wofs = xr.where(wofs == 128, 1, wofs)
```
## Calculate an all-time average
In analogy to WOfS all-time summary, calculate an all-time average for water derived from MNDWI. First, let's threshold each MNDWI frame so we can find the water. What would be a good threshold? This may vary depending on the satellite.
```
# Find all the dates common to both datasets so we can visualise them
# with a fair comparison.
common_dates = sorted(
set(list(s2a_mndwi.time.astype("datetime64[D]").values))
& set(list(ls8_mndwi.time.astype("datetime64[D]").values))
)
# Define MNDWI water index thresholds
s2_threshold = 0
ls8_threshold = 0
# Plot a single date for both Sentinel 2 and Landsat 8 data
fig, axs = plt.subplots(1, 2, figsize=(10, 5))
(s2a_mndwi.MNDWI.sel(time=common_dates[3], method="nearest") >
s2_threshold).plot.imshow(ax=axs[0])
(ls8_mndwi.MNDWI.sel(time=common_dates[3], method="nearest") >
ls8_threshold).plot.imshow(ax=axs[1])
axs[0].set_title("Sentinel 2 > threshold")
axs[1].set_title("LS8 > threshold")
```
A variable threshold like [local Otsu thresholding](https://scikit-image.org/docs/stable/api/skimage.filters.html?highlight=otsu#skimage.filters.threshold_otsu) might be useful in future.
Next, summarise these thresholded scenes and also generate a summary for WOfS by calculating how often each pixel is wet in the data.
Here we make the summaries and plot them.
We have chosen some thresholds for Sentinel 2 and Landsat 8 that seem to get the best results and plotted these as contours - in reality we'd have no manual oversight on this, so this is something of a best-case attempt.
To match the methodology of DEA Waterbodies, we need two thresholds: one for identifying waterbodies and one for finding their maximum extent.
In WOfS (DEA Waterbodies) these are 5% and 10% respectively.
```
# Threshold MNDWI by the MNDWI threshold defined above, then compute
# a summary frequency layer by taking the mean along the time dimension
s2a_mndwi_summary = (s2a_mndwi >= s2_threshold).MNDWI.mean(dim='time')
ls8_mndwi_summary = (ls8_mndwi >= ls8_threshold).MNDWI.mean(dim='time')
wofs_summary = wofs.water.mean(axis=0)
# Summary layer thresholds
s2a_thresholds = [0.03, 0.08]
ls8_thresholds = [0.05, 0.10]
wofs_thresholds = [0.05, 0.10]
fig, axs = plt.subplots(1, 3, figsize=(15, 5))
colours = ["black", "white"]
s2a_mndwi_summary.plot.imshow(ax=axs[0])
ls8_mndwi_summary.plot.imshow(ax=axs[1])
s2a_mndwi_summary.plot.contour(levels=s2a_thresholds, ax=axs[0], colors=colours)
ls8_mndwi_summary.plot.contour(levels=ls8_thresholds, ax=axs[1], colors=colours)
wofs_summary.plot.contour(levels=wofs_thresholds, ax=axs[2], colors=colours)
wofs_summary.plot.imshow(ax=axs[2])
axs[0].set_title("Sentinel 2 MNDWI average")
axs[1].set_title("Landsat 8 MNDWI average")
axs[2].set_title("WOfS water frequency")
```
## Generating polygons
We can generate polygons by judicious choice of threshold for each of these.
The existing method for DEA Waterbodies is to generate polygons at the 10% inundation frequency level as well as the 5% level.
Then, use the 10% polygons to detect waterbodies and the 5% waterbodies to get the maximum extent.
We will use the thresholds from above, which are different for Sentinel 2 and Landsat.
```
def mask_to_polygons(mask):
gdf = xr_vectorize(mask, crs="EPSG:3577")
# Vectorised polygons will have their attribute column
# set based on the pixel values of the mask.
# This is a boolean mask, so we can just check
# for attribute == 1.
return gdf[gdf.attribute == 1]
# Mask Sentinel 2 summary by both thresholds and convert to polygons
s2a_lower_wbs = mask_to_polygons(s2a_mndwi_summary >= s2a_thresholds[0])
s2a_upper_wbs = mask_to_polygons(s2a_mndwi_summary >= s2a_thresholds[1])
# Mask Landsat 8 summary by both thresholds and convert to polygons
ls8_lower_wbs = mask_to_polygons(ls8_mndwi_summary >= ls8_thresholds[0])
ls8_upper_wbs = mask_to_polygons(ls8_mndwi_summary >= ls8_thresholds[1])
# Mask WOfS summary by both thresholds and convert to polygons
wofs_lower_wbs = mask_to_polygons(wofs_summary >= wofs_thresholds[0])
wofs_upper_wbs = mask_to_polygons(wofs_summary >= wofs_thresholds[1])
```
Use the lower boundaries, but only if they contain an upper polygon.
```
s2a_wbs = s2a_lower_wbs.loc[
gpd.sjoin(s2a_lower_wbs, s2a_upper_wbs, how="right").index_left
]
ls8_wbs = ls8_lower_wbs.loc[
gpd.sjoin(ls8_lower_wbs, ls8_upper_wbs, how="right").index_left
]
wofs_wbs = wofs_lower_wbs.loc[
gpd.sjoin(wofs_lower_wbs, wofs_upper_wbs, how="right").index_left
]
```
Let's discard everything less than 5 Landsat pixels in area: 4500 m$^2$ for Landsat Collection 3. This matches the methodology of DEA Waterbodies.
```
# Set an area threshold in square metres
area_threshold = 4500
# Apply this threshold to remove smaller waterbodies
s2a_wbs = s2a_wbs[s2a_wbs.area >= area_threshold]
ls8_wbs = ls8_wbs[ls8_wbs.area >= area_threshold]
wofs_wbs = wofs_wbs[wofs_wbs.area >= area_threshold]
# Plot the resulting waterbodies for Sentinel 2, Landsat 8 and WOfS
fig, axs = plt.subplots(1, 3, figsize=(15, 5))
for wbs, ax, t in zip(
[s2a_wbs, ls8_wbs, wofs_wbs], axs, ["Sentinel 2 MNDWI", "Landsat 8 MNDWI", "WOfS"]
):
wbs.plot(ax=ax)
ax.set_title(t)
```
We end up with a few polygons that are duplicated or overlapping. Combine these by doing a unary union and then split back into polygons again.
```
s2a_wbs = gpd.GeoDataFrame(
geometry=[poly for poly in s2a_wbs.unary_union], crs="EPSG:3577"
)
ls8_wbs = gpd.GeoDataFrame(
geometry=[poly for poly in ls8_wbs.unary_union], crs="EPSG:3577"
)
wofs_wbs = gpd.GeoDataFrame(
geometry=[poly for poly in wofs_wbs.unary_union], crs="EPSG:3577"
)
```
Finally give everything a new ID. We can use this later to generate our masks.
```
s2a_wbs["ID"] = np.arange(1, len(s2a_wbs) + 1).astype(int)
ls8_wbs["ID"] = np.arange(1, len(ls8_wbs) + 1).astype(int)
wofs_wbs["ID"] = np.arange(1, len(wofs_wbs) + 1).astype(int)
```
Set the ID to be the index so we can query by it. Don't drop the ID so we can use it as a column for later rasterising.
```
s2a_wbs.set_index("ID", drop=False, inplace=True)
ls8_wbs.set_index("ID", drop=False, inplace=True)
wofs_wbs.set_index("ID", drop=False, inplace=True)
```
## Surface area time series comparison
Let's choose a waterbody and compare time series derived from different methods.
Select a polygon in the Sentinel 2 waterbodies:
```
s2a_wb = s2a_wbs.iloc[0]
s2a_wb.geometry
```
Then find its equivalent in Landsat MNDWI and WOfS polygons by finding the polygon with the largest intersection.
```
ls8_wb = ls8_wbs.iloc[ls8_wbs.intersection(s2a_wb.geometry).area.argmax()]
wofs_wb = wofs_wbs.iloc[wofs_wbs.intersection(s2a_wb.geometry).area.argmax()]
ls8_wb.geometry
wofs_wb.geometry
```
Calculate a mask that matches the waterbodies.
There are nine masks, which corresponds to three different polygon sets and three different time series.
```
polygon_sets = {
"wofs": wofs_wbs,
"ls8": ls8_wbs,
"s2a": s2a_wbs,
}
ts_dataarrays = {
"wofs": wofs.water,
"ls8": ls8_mndwi.MNDWI >= ls8_threshold,
"s2a": s2a_mndwi.MNDWI >= s2_threshold,
}
ids = {
"wofs": wofs_wb.ID,
"ls8": ls8_wb.ID,
"s2a": s2a_wb.ID,
}
area_per_px = {
"wofs": 25 ** 2,
"ls8": 30 ** 2,
"s2a": 10 ** 2,
}
masks = {} # (polygons, time series)
for poly_name, poly_set in polygon_sets.items():
for ts_name, ts_dataarray in ts_dataarrays.items():
mask = xr_rasterize(poly_set, ts_dataarray.isel(time=0), attribute_col="ID")
masks[poly_name, ts_name] = mask
```
Each mask has pixels set to the ID value of the polygon that contains those pixels.
```
masks["ls8", "ls8"].plot(cmap="rainbow", add_colorbar=False)
```
Use the masks to extract pixel values for each time.
We also want to extract how many pixels are invalid, so we can figure out which days have good observations.
```
ts_wet = {}
ts_invalid = {}
for ts_name, ts_dataarray in ts_dataarrays.items():
for poly_name, poly_set in polygon_sets.items():
mask = masks[poly_name, ts_name] == ids[poly_name]
ts_wet[poly_name, ts_name] = (
ts_dataarray.where(mask).sum(axis=(1, 2)) * area_per_px[ts_name]
).load()
ts_invalid[poly_name, ts_name] = (
ts_dataarray.isnull().where(mask).sum(axis=(1, 2)) * area_per_px[ts_name]
).load()
```
DEA Waterbodies considers an observation valid if there is at least 90% valid pixels.
```
ts_ok = {}
for (poly_name, ts_name), invalid_px in ts_invalid.items():
max_extent = polygon_sets[poly_name].area.loc[ids[poly_name]]
ts_ok[poly_name, ts_name] = ts_invalid[poly_name, ts_name] < 0.1 * max_extent
```
We can plot all of these time series now.
This step may take a while if you didn't preload the data, as it's where Dask has to finally load it all.
```
fig, axs = plt.subplots(3, 3, figsize=(15, 15))
# Subplot label padding.
padding = 10 # pt
# Our datasets have different date ranges, so set these manually for display.
wofs_xlimits = (wofs.time.values.min(), wofs.time.values.max())
xticks = np.arange(np.datetime64("2016", "Y"), np.datetime64("2020", "Y"))
for x, poly_name in enumerate(polygon_sets):
for y, ts_name in enumerate(ts_dataarrays):
ax = axs[y, x]
# https://stackoverflow.com/questions/25812255/row-and-column-headers-in-matplotlibs-subplots
if x == 0:
# Label the vertical axis of subplots.
ax.annotate(
s=ts_name + " time series",
xy=(-0.1, 0.5),
xycoords="axes fraction",
textcoords="offset points",
size="large",
ha="center",
va="center",
xytext=(-padding, 0),
rotation="vertical",
)
if y == 0:
# Label the horizontal axis of subplots.
ax.annotate(
s=poly_name + " polygons",
xy=(0.5, 1),
xycoords="axes fraction",
textcoords="offset points",
size="large",
ha="center",
va="center",
xytext=(0, padding),
rotation="horizontal",
)
# Plot valid dates of the time series.
ok = ts_ok[poly_name, ts_name]
ts = ts_wet[poly_name, ts_name]
ax.plot(ts.time[ok], ts[ok], marker="x", linestyle="None")
ax.set_xlim(wofs_xlimits)
ax.set_xticks(xticks)
ax.set_xticklabels(xticks)
```
The MNDWI series are noticably noisier, with lots of no-water or low-water observations that are wet in WOfS.
However, the general pattern matches WOfS, and in particular seems to bound it below.
Maybe we could consider MNDWI a lower bound for wetness?
We can also estimate a mean-squared difference between each time series and the WOfS time series with WOfS polygons (= DEA Waterbodies).
This assumes Gaussian error, but for sufficiently large numbers of pixels the binomial distribution we actually have should be well-approximated by a Gaussian.
By calculating such a difference, we can summarise the discrepancies between the above plots as a single number per plot.
```
# Make a pandas series of valid data for comparison.
# We resample to daily to avoid time rounding issues.
ok = ts_ok["wofs", "wofs"]
ts = ts_wet["wofs", "wofs"]
comparison_series = (
pd.Series(ts[ok], index=ts.time[ok].values, name="comparison").resample("D").mean()
)
differences = {}
for poly_name in polygon_sets:
for ts_name in ts_dataarrays:
# Make a pandas series of valid data.
ok = ts_ok[poly_name, ts_name]
ts = ts_wet[poly_name, ts_name]
series = (
pd.Series(ts[ok], index=ts.time[ok].values, name="test")
.resample("D")
.mean()
)
# Join this with the comparison series to have a unified time index.
joint_series = pd.merge(
series, comparison_series, left_index=True, right_index=True, how="outer"
)
# Interpolate and compute the difference.
joint_series.interpolate(inplace=True)
difference = joint_series.test - joint_series.comparison
# The mean-squared error is the difference we seek.
differences[poly_name, ts_name] = (difference ** 2).mean()
```
Now we can look at these differences as a table.
```
fig = plt.figure(figsize=(8, 8))
grid = np.zeros((3, 3))
for x, poly_name in enumerate(sorted(polygon_sets)):
for y, ts_name in enumerate(sorted(ts_dataarrays)):
grid[x, y] = differences[poly_name, ts_name]
plt.pcolor(grid, cmap="Reds")
plt.colorbar(label="Difference")
plt.xticks(np.arange(3) + 0.5, [i + " polygons" for i in sorted(polygon_sets)])
plt.yticks(np.arange(3) + 0.5, [i + " time series" for i in sorted(ts_dataarrays)]);
```
The polygons dominate the differences between the time series.
This shows that the polygons are the most important change here between the datasets, dwarfing the impact of the low-water observations.
To see these more clearly, we can normalise the difference by the kind of polygons before we plot:
```
fig = plt.figure(figsize=(8, 8))
plt.pcolor(grid / grid.max(axis=0), cmap="Reds")
plt.colorbar(label="Normalised difference")
plt.xticks(np.arange(3) + 0.5, [i + " polygons" for i in sorted(polygon_sets)])
plt.yticks(np.arange(3) + 0.5, [i + " time series" for i in sorted(ts_dataarrays)]);
```
S2 and LS8 MNDWI are equally good when applied to non-WOfS polygons.
S2 time series deviate appreciably when applied to WOfS polygons, though.
This makes sense because S2 will include lots of dry areas considered part of the waterbody by the much larger WOfS pixels.
***
## Additional information
**License:** The code in this notebook is licensed under the [Apache License, Version 2.0](https://www.apache.org/licenses/LICENSE-2.0).
Digital Earth Australia data is licensed under the [Creative Commons by Attribution 4.0](https://creativecommons.org/licenses/by/4.0/) license.
**Contact:** If you need assistance, please post a question on the [Open Data Cube Slack channel](http://slack.opendatacube.org/) or on the [GIS Stack Exchange](https://gis.stackexchange.com/questions/ask?tags=open-data-cube) using the `open-data-cube` tag (you can view previously asked questions [here](https://gis.stackexchange.com/questions/tagged/open-data-cube)).
If you would like to report an issue with this notebook, you can file one on [Github](https://github.com/GeoscienceAustralia/dea-notebooks).
**Last modified:** October 2020
**Compatible datacube version:**
```
print(datacube.__version__)
```
## Tags
Browse all available tags on the DEA User Guide's [Tags Index](https://docs.dea.ga.gov.au/genindex.html)
| github_jupyter |
# Importing Libraries
```
import sys
sys.path.append('..')
from letcon.src.utils.utils import save_artifacts
from letcon.src.data_loader import data_ml
from letcon.src.model import model_ml
import requests
import pandas as pd
```
# Loading Data
```
TrainData = pd.read_csv('data/wine-quality/winequality-white-updated.csv')
```
# Custom Data Processing Library
## Creating config for our data processing library
```
data_config= {'train_data' : TrainData,
'test_data' : None,
'unique_id' : None,
'time_id' : None,
'x_vars' : [
'fixed_acidity',
'volatile_acidity',
'citric_acid',
'residual_sugar',
'chlorides',
'free_sulfur_dioxide',
'total_sulfur_dioxide',
'density',
'pH',
'sulphates',
'alcohol',
],
'cat_vars' : [],
'y_var' : 'quality',
'stratify' : 'quality',
'test_size' : 0.20,
'use_full_dataset' : True,
'encoding_style' : 'label_encoding',
'impute_missing' : 'mean',
'capping_vars' : {},
'task' : 'regression',
'random_state' : 42,
}
```
## Initializing data processing object
```
data = data_ml.DataLoader(config=data_config)
```
# Custom Modelling Library
## Creating config for our modelling library
```
modelling_config = {
'data': data.get_data(),
'model_type' : 'simple',
'model_name' : 'xgboost',
'model_inputs' : {},
'scoring_function' : 'rmse',
'model_initial_params' : {'verbose':True,
'boosting': 'gbtree',
'tree_method': 'exact',
'n_estimators': 127,
'max_depth': 9,
'reg_alpha': 10,
'reg_lambda': 22,
'min_child_weight': 1,
'gamma': 1,
'learning_rate': 0.4901527567844427,
},
'hyperparmeter_tuning' : {'enable_tuning' : True,
'optimizer' : 'optuna',
'optimizer_params' : {'fixed' : {'n_trials' : 5},
'varying' : {'boosting' : ['gbtree', 'gblinear'],
'tree_method' : ['exact','approx','hist'],
'n_estimators' : {'min' : 50, 'max' :200},
'max_depth' : {'min' : 2, 'max' : 10},
'reg_alpha' : {'min' : 0, 'max' : 10},
'reg_lambda' : {'min' : 0, 'max' : 25},
'min_child_weight' : {'min' : 0, 'max' : 1},
'gamma' : {'min' : 0, 'max' : 50},
'learning_rate' : {'min' : 0.3, 'max' : 0.5}
}
},
},
'shap_analysis' : {'enable_shap' : True,
'use_explainer' : 'tree',
},
'task' : 'regression',
'random_state' : 42,
}
```
## Initializing modelling object
```
model = model_ml.Model(config=modelling_config)
model.create_model()
```
# Saving Data and Model Artifacts
## Initializing objects for the class which will be used for prediction
```
data_pipeline_object = data_ml.ProcessPredictionData(config = data.get_data_artifacts())
trained_model_object = model_ml.PredictOnNewData(config = model.get_model_artifacts())
```
## Saving the objects in a readable pickle format
```
save_artifacts(data_object=data_pipeline_object,
model_object=trained_model_object)
```
# Let's Turn ON the Uvicorn Server
## Creating packets of request which will be used for inference
```
to_predict_dict = {"pH": 0.38,
"chlorides": 0.53,
"volatile_acidity": 2.0,
"citric_acid": 157,
"alcohol": 3.0,
"total_sulfur_dioxide": 0,
"density": 0,
"residual_sugar": 0.0,
"fixed_acidity": 0.0,
"sulphates" : 0.0,
"free_sulfur_dioxide" : 1.0}
# to_predict_dict = {"pH": 3.00,
# "chlorides": 0.045,
# "volatile_acidity": 0.27,
# "citric_acid": 0.36,
# "alcohol": 8.8,
# "total_sulfur_dioxide": 170.0,
# "density": 1.0010,
# "residual_sugar": 20.7,
# "fixed_acidity": 7.0,
# "sulphates" : 0.45,
# "free_sulfur_dioxide" : 45.0}
url = 'http://127.0.0.1:8000/predict'
r = requests.post(url, json=to_predict_dict)
print(r.json())
```
| github_jupyter |
# Regression with Amazon SageMaker XGBoost algorithm
_**Single machine training for regression with Amazon SageMaker XGBoost algorithm**_
---
---
## Contents
1. [Introduction](#Introduction)
2. [Setup](#Setup)
1. [Fetching the dataset](#Fetching-the-dataset)
2. [Data Ingestion](#Data-ingestion)
3. [Training the XGBoost model](#Training-the-XGBoost-model)
1. [Plotting evaluation metrics](#Plotting-evaluation-metrics)
4. [Set up hosting for the model](#Set-up-hosting-for-the-model)
1. [Import model into hosting](#Import-model-into-hosting)
2. [Create endpoint configuration](#Create-endpoint-configuration)
3. [Create endpoint](#Create-endpoint)
5. [Validate the model for use](#Validate-the-model-for-use)
---
## Introduction
This notebook demonstrates the use of Amazon SageMaker’s implementation of the XGBoost algorithm to train and host a regression model. We use the [Abalone data](https://www.csie.ntu.edu.tw/~cjlin/libsvmtools/datasets/regression.html) originally from the [UCI data repository](https://archive.ics.uci.edu/ml/datasets/abalone). More details about the original dataset can be found [here](https://archive.ics.uci.edu/ml/machine-learning-databases/abalone/abalone.names). In the libsvm converted [version](https://www.csie.ntu.edu.tw/~cjlin/libsvmtools/datasets/regression.html), the nominal feature (Male/Female/Infant) has been converted into a real valued feature. Age of abalone is to be predicted from eight physical measurements.
---
## Setup
This notebook was created and tested on an ml.m4.4xlarge notebook instance.
Let's start by specifying:
1. The S3 bucket and prefix that you want to use for training and model data. This should be within the same region as the Notebook Instance, training, and hosting.
1. The IAM role arn used to give training and hosting access to your data. See the documentation for how to create these. Note, if more than one role is required for notebook instances, training, and/or hosting, please replace the boto regexp with a the appropriate full IAM role arn string(s).
```
%%time
import os
import boto3
import re
from sagemaker import get_execution_role
role = get_execution_role()
region = boto3.Session().region_name
bucket='<bucket-name>' # put your s3 bucket name here, and create s3 bucket
prefix = 'sagemaker/DEMO-xgboost-regression'
# customize to your bucket where you have stored the data
bucket_path = 'https://s3-{}.amazonaws.com/{}'.format(region,bucket)
```
### Fetching the dataset
Following methods split the data into train/test/validation datasets and upload files to S3.
```
%%time
import io
import boto3
import random
def data_split(FILE_DATA, FILE_TRAIN, FILE_VALIDATION, FILE_TEST, PERCENT_TRAIN, PERCENT_VALIDATION, PERCENT_TEST):
data = [l for l in open(FILE_DATA, 'r')]
train_file = open(FILE_TRAIN, 'w')
valid_file = open(FILE_VALIDATION, 'w')
tests_file = open(FILE_TEST, 'w')
num_of_data = len(data)
num_train = int((PERCENT_TRAIN/100.0)*num_of_data)
num_valid = int((PERCENT_VALIDATION/100.0)*num_of_data)
num_tests = int((PERCENT_TEST/100.0)*num_of_data)
data_fractions = [num_train, num_valid, num_tests]
split_data = [[],[],[]]
rand_data_ind = 0
for split_ind, fraction in enumerate(data_fractions):
for i in range(fraction):
rand_data_ind = random.randint(0, len(data)-1)
split_data[split_ind].append(data[rand_data_ind])
data.pop(rand_data_ind)
for l in split_data[0]:
train_file.write(l)
for l in split_data[1]:
valid_file.write(l)
for l in split_data[2]:
tests_file.write(l)
train_file.close()
valid_file.close()
tests_file.close()
def write_to_s3(fobj, bucket, key):
return boto3.Session().resource('s3').Bucket(bucket).Object(key).upload_fileobj(fobj)
def upload_to_s3(bucket, channel, filename):
fobj=open(filename, 'rb')
key = prefix+'/'+channel
url = 's3://{}/{}/{}'.format(bucket, key, filename)
print('Writing to {}'.format(url))
write_to_s3(fobj, bucket, key)
```
### Data ingestion
Next, we read the dataset from the existing repository into memory, for preprocessing prior to training. This processing could be done *in situ* by Amazon Athena, Apache Spark in Amazon EMR, Amazon Redshift, etc., assuming the dataset is present in the appropriate location. Then, the next step would be to transfer the data to S3 for use in training. For small datasets, such as this one, reading into memory isn't onerous, though it would be for larger datasets.
```
%%time
import urllib.request
# Load the dataset
FILE_DATA = 'abalone'
urllib.request.urlretrieve("https://www.csie.ntu.edu.tw/~cjlin/libsvmtools/datasets/regression/abalone", FILE_DATA)
#split the downloaded data into train/test/validation files
FILE_TRAIN = 'abalone.train'
FILE_VALIDATION = 'abalone.validation'
FILE_TEST = 'abalone.test'
PERCENT_TRAIN = 70
PERCENT_VALIDATION = 15
PERCENT_TEST = 15
data_split(FILE_DATA, FILE_TRAIN, FILE_VALIDATION, FILE_TEST, PERCENT_TRAIN, PERCENT_VALIDATION, PERCENT_TEST)
#upload the files to the S3 bucket
upload_to_s3(bucket, 'train', FILE_TRAIN)
upload_to_s3(bucket, 'validation', FILE_VALIDATION)
upload_to_s3(bucket, 'test', FILE_TEST)
```
## Training the XGBoost model
After setting training parameters, we kick off training, and poll for status until training is completed, which in this example, takes between 5 and 6 minutes.
```
from sagemaker.amazon.amazon_estimator import get_image_uri
container = get_image_uri(boto3.Session().region_name, 'xgboost')
%%time
import boto3
from time import gmtime, strftime
job_name = 'DEMO-xgboost-regression-' + strftime("%Y-%m-%d-%H-%M-%S", gmtime())
print("Training job", job_name)
#Ensure that the training and validation data folders generated above are reflected in the "InputDataConfig" parameter below.
create_training_params = \
{
"AlgorithmSpecification": {
"TrainingImage": container,
"TrainingInputMode": "File"
},
"RoleArn": role,
"OutputDataConfig": {
"S3OutputPath": bucket_path + "/" + prefix + "/single-xgboost"
},
"ResourceConfig": {
"InstanceCount": 1,
"InstanceType": "ml.m4.4xlarge",
"VolumeSizeInGB": 5
},
"TrainingJobName": job_name,
"HyperParameters": {
"max_depth":"5",
"eta":"0.2",
"gamma":"4",
"min_child_weight":"6",
"subsample":"0.7",
"silent":"0",
"objective":"reg:linear",
"num_round":"50"
},
"StoppingCondition": {
"MaxRuntimeInSeconds": 3600
},
"InputDataConfig": [
{
"ChannelName": "train",
"DataSource": {
"S3DataSource": {
"S3DataType": "S3Prefix",
"S3Uri": bucket_path + "/" + prefix + '/train',
"S3DataDistributionType": "FullyReplicated"
}
},
"ContentType": "libsvm",
"CompressionType": "None"
},
{
"ChannelName": "validation",
"DataSource": {
"S3DataSource": {
"S3DataType": "S3Prefix",
"S3Uri": bucket_path + "/" + prefix + '/validation',
"S3DataDistributionType": "FullyReplicated"
}
},
"ContentType": "libsvm",
"CompressionType": "None"
}
]
}
client = boto3.client('sagemaker')
client.create_training_job(**create_training_params)
import time
status = client.describe_training_job(TrainingJobName=job_name)['TrainingJobStatus']
print(status)
while status !='Completed' and status!='Failed':
time.sleep(60)
status = client.describe_training_job(TrainingJobName=job_name)['TrainingJobStatus']
print(status)
```
Note that the "validation" channel has been initialized too. The SageMaker XGBoost algorithm actually calculates RMSE and writes it to the CloudWatch logs on the data passed to the "validation" channel.
### Plotting evaluation metrics
Evaluation metrics for the completed training job are available in CloudWatch. We can pull the area under curve metric for the validation data set and plot it to see the performance of the model over time.
```
%matplotlib inline
from sagemaker.analytics import TrainingJobAnalytics
metric_name = 'validation:rmse'
metrics_dataframe = TrainingJobAnalytics(training_job_name=job_name, metric_names=[metric_name]).dataframe()
plt = metrics_dataframe.plot(kind='line', figsize=(12,5), x='timestamp', y='value', style='b.', legend=False)
plt.set_ylabel(metric_name);
```
## Set up hosting for the model
In order to set up hosting, we have to import the model from training to hosting.
### Import model into hosting
Register the model with hosting. This allows the flexibility of importing models trained elsewhere.
```
%%time
import boto3
from time import gmtime, strftime
model_name=job_name + '-model'
print(model_name)
info = client.describe_training_job(TrainingJobName=job_name)
model_data = info['ModelArtifacts']['S3ModelArtifacts']
print(model_data)
primary_container = {
'Image': container,
'ModelDataUrl': model_data
}
create_model_response = client.create_model(
ModelName = model_name,
ExecutionRoleArn = role,
PrimaryContainer = primary_container)
print(create_model_response['ModelArn'])
```
### Create endpoint configuration
SageMaker supports configuring REST endpoints in hosting with multiple models, e.g. for A/B testing purposes. In order to support this, customers create an endpoint configuration, that describes the distribution of traffic across the models, whether split, shadowed, or sampled in some way. In addition, the endpoint configuration describes the instance type required for model deployment.
```
from time import gmtime, strftime
endpoint_config_name = 'DEMO-XGBoostEndpointConfig-' + strftime("%Y-%m-%d-%H-%M-%S", gmtime())
print(endpoint_config_name)
create_endpoint_config_response = client.create_endpoint_config(
EndpointConfigName = endpoint_config_name,
ProductionVariants=[{
'InstanceType':'ml.m4.xlarge',
'InitialVariantWeight':1,
'InitialInstanceCount':1,
'ModelName':model_name,
'VariantName':'AllTraffic'}])
print("Endpoint Config Arn: " + create_endpoint_config_response['EndpointConfigArn'])
```
### Create endpoint
Lastly, the customer creates the endpoint that serves up the model, through specifying the name and configuration defined above. The end result is an endpoint that can be validated and incorporated into production applications. This takes 9-11 minutes to complete.
```
%%time
import time
endpoint_name = 'DEMO-XGBoostEndpoint-' + strftime("%Y-%m-%d-%H-%M-%S", gmtime())
print(endpoint_name)
create_endpoint_response = client.create_endpoint(
EndpointName=endpoint_name,
EndpointConfigName=endpoint_config_name)
print(create_endpoint_response['EndpointArn'])
resp = client.describe_endpoint(EndpointName=endpoint_name)
status = resp['EndpointStatus']
print("Status: " + status)
while status=='Creating':
time.sleep(60)
resp = client.describe_endpoint(EndpointName=endpoint_name)
status = resp['EndpointStatus']
print("Status: " + status)
print("Arn: " + resp['EndpointArn'])
print("Status: " + status)
```
## Validate the model for use
Finally, the customer can now validate the model for use. They can obtain the endpoint from the client library using the result from previous operations, and generate classifications from the trained model using that endpoint.
```
runtime_client = boto3.client('runtime.sagemaker')
```
Start with a single prediction.
```
!head -1 abalone.test > abalone.single.test
%%time
import json
from itertools import islice
import math
import struct
file_name = 'abalone.single.test' #customize to your test file
with open(file_name, 'r') as f:
payload = f.read().strip()
response = runtime_client.invoke_endpoint(EndpointName=endpoint_name,
ContentType='text/x-libsvm',
Body=payload)
result = response['Body'].read()
result = result.decode("utf-8")
result = result.split(',')
result = [math.ceil(float(i)) for i in result]
label = payload.strip(' ').split()[0]
print ('Label: ',label,'\nPrediction: ', result[0])
```
OK, a single prediction works. Let's do a whole batch to see how good is the predictions accuracy.
```
import sys
import math
def do_predict(data, endpoint_name, content_type):
payload = '\n'.join(data)
response = runtime_client.invoke_endpoint(EndpointName=endpoint_name,
ContentType=content_type,
Body=payload)
result = response['Body'].read()
result = result.decode("utf-8")
result = result.split(',')
preds = [float((num)) for num in result]
preds = [math.ceil(num) for num in preds]
return preds
def batch_predict(data, batch_size, endpoint_name, content_type):
items = len(data)
arrs = []
for offset in range(0, items, batch_size):
if offset+batch_size < items:
results = do_predict(data[offset:(offset+batch_size)], endpoint_name, content_type)
arrs.extend(results)
else:
arrs.extend(do_predict(data[offset:items], endpoint_name, content_type))
sys.stdout.write('.')
return(arrs)
```
The following helps us calculate the Median Absolute Percent Error (MdAPE) on the batch dataset.
```
%%time
import json
import numpy as np
with open(FILE_TEST, 'r') as f:
payload = f.read().strip()
labels = [int(line.split(' ')[0]) for line in payload.split('\n')]
test_data = [line for line in payload.split('\n')]
preds = batch_predict(test_data, 100, endpoint_name, 'text/x-libsvm')
print('\n Median Absolute Percent Error (MdAPE) = ', np.median(np.abs(np.array(labels) - np.array(preds)) / np.array(labels)))
```
### Delete Endpoint
Once you are done using the endpoint, you can use the following to delete it.
```
client.delete_endpoint(EndpointName=endpoint_name)
```
| github_jupyter |
# Session 3: Unsupervised and Supervised Learning
<p class="lead">
Assignment: Build Unsupervised and Supervised Networks
</p>
<p class="lead">
Parag K. Mital<br />
<a href="https://www.kadenze.com/courses/creative-applications-of-deep-learning-with-tensorflow/info">Creative Applications of Deep Learning w/ Tensorflow</a><br />
<a href="https://www.kadenze.com/partners/kadenze-academy">Kadenze Academy</a><br />
<a href="https://twitter.com/hashtag/CADL">#CADL</a>
</p>
<a name="learning-goals"></a>
# Learning Goals
* Learn how to build an autoencoder
* Learn how to explore latent/hidden representations of an autoencoder.
* Learn how to build a classification network using softmax and onehot encoding
# Outline
<!-- MarkdownTOC autolink=true autoanchor=true bracket=round -->
- [Assignment Synopsis](#assignment-synopsis)
- [Part One - Autoencoders](#part-one---autoencoders)
- [Instructions](#instructions)
- [Code](#code)
- [Visualize the Embedding](#visualize-the-embedding)
- [Reorganize to Grid](#reorganize-to-grid)
- [2D Latent Manifold](#2d-latent-manifold)
- [Part Two - General Autoencoder Framework](#part-two---general-autoencoder-framework)
- [Instructions](#instructions-1)
- [Part Three - Deep Audio Classification Network](#part-three---deep-audio-classification-network)
- [Instructions](#instructions-2)
- [Preparing the Data](#preparing-the-data)
- [Creating the Network](#creating-the-network)
- [Assignment Submission](#assignment-submission)
- [Coming Up](#coming-up)
<!-- /MarkdownTOC -->
This next section will just make sure you have the right version of python and the libraries that we'll be using. Don't change the code here but make sure you "run" it (use "shift+enter")!
```
# First check the Python version
import sys
if sys.version_info < (3,4):
print('You are running an older version of Python!\n\n' \
'You should consider updating to Python 3.4.0 or ' \
'higher as the libraries built for this course ' \
'have only been tested in Python 3.4 and higher.\n')
print('Try installing the Python 3.5 version of anaconda '
'and then restart `jupyter notebook`:\n' \
'https://www.continuum.io/downloads\n\n')
# Now get necessary libraries
try:
import os
import numpy as np
import matplotlib.pyplot as plt
from skimage.transform import resize
from skimage import data
from scipy.misc import imresize
import IPython.display as ipyd
except ImportError:
print('You are missing some packages! ' \
'We will try installing them before continuing!')
!pip install "numpy>=1.11.0" "matplotlib>=1.5.1" "scikit-image>=0.11.3" "scikit-learn>=0.17" "scipy>=0.17.0"
import os
import numpy as np
import matplotlib.pyplot as plt
from skimage.transform import resize
from skimage import data
from scipy.misc import imresize
import IPython.display as ipyd
print('Done!')
# Import Tensorflow
try:
import tensorflow as tf
except ImportError:
print("You do not have tensorflow installed!")
print("Follow the instructions on the following link")
print("to install tensorflow before continuing:")
print("")
print("https://github.com/pkmital/CADL#installation-preliminaries")
# This cell includes the provided libraries from the zip file
# and a library for displaying images from ipython, which
# we will use to display the gif
try:
from libs import utils, gif, datasets, dataset_utils, vae, dft
except ImportError:
print("Make sure you have started notebook in the same directory" +
" as the provided zip file which includes the 'libs' folder" +
" and the file 'utils.py' inside of it. You will NOT be able"
" to complete this assignment unless you restart jupyter"
" notebook inside the directory created by extracting"
" the zip file or cloning the github repo.")
# We'll tell matplotlib to inline any drawn figures like so:
%matplotlib inline
plt.style.use('ggplot')
# Bit of formatting because I don't like the default inline code style:
from IPython.core.display import HTML
HTML("""<style> .rendered_html code {
padding: 2px 4px;
color: #c7254e;
background-color: #f9f2f4;
border-radius: 4px;
} </style>""")
```
<a name="assignment-synopsis"></a>
# Assignment Synopsis
In the last session we created our first neural network. We saw that in order to create a neural network, we needed to define a cost function which would allow gradient descent to optimize all the parameters in our network. We also saw how neural networks become much more expressive by introducing series of linearities followed by non-linearities, or activation functions. We then explored a fun application of neural networks using regression to learn to paint color values given x, y positions. This allowed us to build up a sort of painterly like version of an image.
In this session, we'll see how to construct a few more types of neural networks. First, we'll explore a generative network called autoencoders. This network can be extended in a variety of ways to include convolution, denoising, or a variational layer. In Part Two, you'll then use a general autoencoder framework to encode your own list of images. In Part three, we'll then explore a discriminative network used for classification, and see how this can be used for audio classification of music or speech.
One main difference between these two networks are the data that we'll use to train them. In the first case, we will only work with "unlabeled" data and perform unsupervised learning. An example would be a collection of images, just like the one you created for assignment 1. Contrast this with "labeled" data which allows us to make use of supervised learning. For instance, we're given both images, and some other data about those images such as some text describing what object is in the image. This allows us to optimize a network where we model a distribution over the images given that it should be labeled as something. This is often a *much* simpler distribution to train, but with the expense of it being much harder to collect.
One of the major directions of future research will be in how to better make use of unlabeled data and unsupervised learning methods.
<a name="part-one---autoencoders"></a>
# Part One - Autoencoders
<a name="instructions"></a>
## Instructions
Work with a dataset of images and train an autoencoder. You can work with the same dataset from assignment 1, or try a larger dataset. But be careful with the image sizes, and make sure to keep it relatively small (e.g. < 200 x 200 px).
Recall from the lecture that autoencoders are great at "compressing" information. The network's construction and cost function are just like what we've done in the last session. The network is composed of a series of matrix multiplications and nonlinearities. The only difference is the output of the network has exactly the same shape as what is input. This allows us to train the network by saying that the output of the network needs to be just like the input to it, so that it tries to "compress" all the information in that video.
Autoencoders have some great potential for creative applications, as they allow us to compress a dataset of information and even *generate* new data from that encoding. We'll see exactly how to do this with a basic autoencoder, and then you'll be asked to explore some of the extensions to produce your own encodings.
<a name="code"></a>
## Code
We'll now go through the process of building an autoencoder just like in the lecture. First, let's load some data. You can use the first 100 images of the Celeb Net, your own dataset, or anything else approximately under 1,000 images. Make sure you resize the images so that they are <= 200x200 pixels, otherwise the training will be *very* slow, and the montages we create will be too large.
<h3><font color='red'>TODO! COMPLETE THIS SECTION!</font></h3>
```
# See how this works w/ Celeb Images or try your own dataset instead:
imgs = ...
# Then convert the list of images to a 4d array (e.g. use np.array to convert a list to a 4d array):
Xs = ...
print(Xs.shape)
assert(Xs.ndim == 4 and Xs.shape[1] <= 250 and Xs.shape[2] <= 250)
```
We'll now make use of something I've written to help us store this data. It provides some interfaces for generating "batches" of data, as well as splitting the data into training, validation, and testing sets. To use it, we pass in the data and optionally its labels. If we don't have labels, we just pass in the data. In the second half of this notebook, we'll explore using a dataset's labels as well.
```
ds = datasets.Dataset(Xs)
# ds = datasets.CIFAR10(flatten=False)
```
It allows us to easily find the mean:
```
mean_img = ds.mean().astype(np.uint8)
plt.imshow(mean_img)
```
Or the deviation:
```
std_img = ds.std()
plt.imshow(std_img)
print(std_img.shape)
```
Recall we can calculate the mean of the standard deviation across each color channel:
```
std_img = np.mean(std_img, axis=2).astype(np.uint8)
plt.imshow(std_img)
```
All the input data we gave as input to our `Datasets` object, previously stored in `Xs` is now stored in a variable as part of our `ds` Datasets object, `X`:
```
plt.imshow(ds.X[0])
print(ds.X.shape)
```
It takes a parameter, `split` at the time of creation, which allows us to create train/valid/test sets. By default, this is set to `[1.0, 0.0, 0.0]`, which means to take all the data in the train set, and nothing in the validation and testing sets. We can access "batch generators" of each of these sets by saying: `ds.train.next_batch`. A generator is a really powerful way of handling iteration in Python. If you are unfamiliar with the idea of generators, I recommend reading up a little bit on it, e.g. here: http://intermediatepythonista.com/python-generators - think of it as a for loop, but as a function. It returns one iteration of the loop each time you call it.
This generator will automatically handle the randomization of the dataset. Let's try looping over the dataset using the batch generator:
```
for (X, y) in ds.train.next_batch(batch_size=10):
print(X.shape)
```
This returns `X` and `y` as a tuple. Since we're not using labels, we'll just ignore this. The `next_batch` method takes a parameter, `batch_size`, which we'll set appropriately to our batch size. Notice it runs for exactly 10 iterations to iterate over our 100 examples, then the loop exits. The order in which it iterates over the 100 examples is randomized each time you iterate.
Write two functions to preprocess (normalize) any given image, and to unprocess it, i.e. unnormalize it by removing the normalization. The `preprocess` function should perform exactly the task you learned to do in assignment 1: subtract the mean, then divide by the standard deviation. The `deprocess` function should take the preprocessed image and undo the preprocessing steps. Recall that the `ds` object contains the `mean` and `std` functions for access the mean and standarad deviation. We'll be using the `preprocess` and `deprocess` functions on the input and outputs of the network. Note, we could use Tensorflow to do this instead of numpy, but for sake of clarity, I'm keeping this separate from the Tensorflow graph.
<h3><font color='red'>TODO! COMPLETE THIS SECTION!</font></h3>
```
# Write a function to preprocess/normalize an image, given its dataset object
# (which stores the mean and standard deviation!)
def preprocess(img, ds):
norm_img = (img - ...) / ...
return norm_img
# Write a function to undo the normalization of an image, given its dataset object
# (which stores the mean and standard deviation!)
def deprocess(norm_img, ds):
img = norm_img * ... + ...
return img
```
We're going to now work on creating an autoencoder. To start, we'll only use linear connections, like in the last assignment. This means, we need a 2-dimensional input: Batch Size x Number of Features. We currently have a 4-dimensional input: Batch Size x Height x Width x Channels. We'll have to calculate the number of features we have to help construct the Tensorflow Graph for our autoencoder neural network. Then, when we are ready to train the network, we'll reshape our 4-dimensional dataset into a 2-dimensional one when feeding the input of the network. Optionally, we could create a `tf.reshape` as the first operation of the network, so that we can still pass in our 4-dimensional array, and the Tensorflow graph would reshape it for us. We'll try the former method, by reshaping manually, and then you can explore the latter method, of handling 4-dimensional inputs on your own.
<h3><font color='red'>TODO! COMPLETE THIS SECTION!</font></h3>
```
# Calculate the number of features in your image.
# This is the total number of pixels, or (height x width x channels).
n_features = ...
print(n_features)
```
Let's create a list of how many neurons we want in each layer. This should be for just one half of the network, the encoder only. It should start large, then get smaller and smaller. We're also going to try an encode our dataset to an inner layer of just 2 values. So from our number of features, we'll go all the way down to expressing that image by just 2 values. Try the values I've put here for the celeb dataset, then explore your own values:
<h3><font color='red'>TODO! COMPLETE THIS SECTION!</font></h3>
```
encoder_dimensions = [2048, 512, 128, 2]
```
Now create a placeholder just like in the last session in the tensorflow graph that will be able to get any number (None) of `n_features` inputs.
<h3><font color='red'>TODO! COMPLETE THIS SECTION!</font></h3>
```
X = tf.placeholder(...
assert(X.get_shape().as_list() == [None, n_features])
```
Now complete the function `encode` below. This takes as input our input placeholder, `X`, our list of `dimensions`, and an `activation` function, e.g. `tf.nn.relu` or `tf.nn.tanh`, to apply to each layer's output, and creates a series of fully connected layers. This works just like in the last session! We multiply our input, add a bias, then apply a non-linearity. Instead of having 20 neurons in each layer, we're going to use our `dimensions` list to tell us how many neurons we want in each layer.
One important difference is that we're going to also store every weight matrix we create! This is so that we can use the same weight matrices when we go to build our decoder. This is a *very* powerful concept that creeps up in a few different neural network architectures called weight sharing. Weight sharing isn't necessary to do of course, but can speed up training and offer a different set of features depending on your dataset. Explore trying both. We'll also see how another form of weight sharing works in convolutional networks.
<h3><font color='red'>TODO! COMPLETE THIS SECTION!</font></h3>
```
def encode(X, dimensions, activation=tf.nn.tanh):
# We're going to keep every matrix we create so let's create a list to hold them all
Ws = []
# We'll create a for loop to create each layer:
for layer_i, n_output in enumerate(dimensions):
# TODO: just like in the last session,
# we'll use a variable scope to help encapsulate our variables
# This will simply prefix all the variables made in this scope
# with the name we give it. Make sure it is a unique name
# for each layer, e.g., 'encoder/layer1', 'encoder/layer2', or
# 'encoder/1', 'encoder/2',...
with tf.variable_scope(...)
# TODO: Create a weight matrix which will increasingly reduce
# down the amount of information in the input by performing
# a matrix multiplication. You can use the utils.linear function.
h, W = ...
# Finally we'll store the weight matrix.
# We need to keep track of all
# the weight matrices we've used in our encoder
# so that we can build the decoder using the
# same weight matrices.
Ws.append(W)
# Replace X with the current layer's output, so we can
# use it in the next layer.
X = h
z = X
return Ws, z
# Then call the function
Ws, z = encode(X, encoder_dimensions)
# And just some checks to make sure you've done it right.
assert(z.get_shape().as_list() == [None, 2])
assert(len(Ws) == len(encoder_dimensions))
```
Let's take a look at the graph:
```
[op.name for op in tf.get_default_graph().get_operations()]
```
So we've created a few layers, encoding our input `X` all the way down to 2 values in the tensor `z`. We do this by multiplying our input `X` by a set of matrices shaped as:
```
[W_i.get_shape().as_list() for W_i in Ws]
```
Resulting in a layer which is shaped as:
```
z.get_shape().as_list()
```
## Building the Decoder
Here is a helpful animation on what the matrix "transpose" operation does:
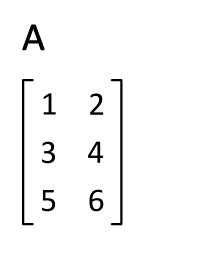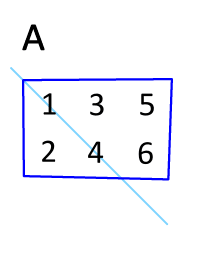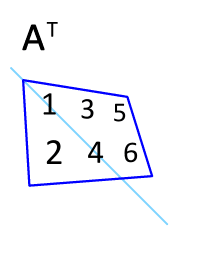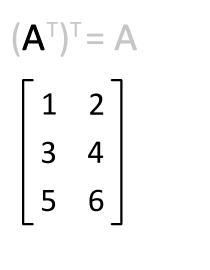
Basically what is happening is rows becomes columns, and vice-versa. We're going to use our existing weight matrices but transpose them so that we can go in the opposite direction. In order to build our decoder, we'll have to do the opposite of what we've just done, multiplying `z` by the transpose of our weight matrices, to get back to a reconstructed version of `X`. First, we'll reverse the order of our weight matrics, and then append to the list of dimensions the final output layer's shape to match our input:
```
# We'll first reverse the order of our weight matrices
decoder_Ws = Ws[::-1]
# then reverse the order of our dimensions
# appending the last layers number of inputs.
decoder_dimensions = encoder_dimensions[::-1][1:] + [n_features]
print(decoder_dimensions)
assert(decoder_dimensions[-1] == n_features)
```
Now we'll build the decoder. I've shown you how to do this. Read through the code to fully understand what it is doing:
```
def decode(z, dimensions, Ws, activation=tf.nn.tanh):
current_input = z
for layer_i, n_output in enumerate(dimensions):
# we'll use a variable scope again to help encapsulate our variables
# This will simply prefix all the variables made in this scope
# with the name we give it.
with tf.variable_scope("decoder/layer/{}".format(layer_i)):
# Now we'll grab the weight matrix we created before and transpose it
# So a 3072 x 784 matrix would become 784 x 3072
# or a 256 x 64 matrix, would become 64 x 256
W = tf.transpose(Ws[layer_i])
# Now we'll multiply our input by our transposed W matrix
h = tf.matmul(current_input, W)
# And then use a relu activation function on its output
current_input = activation(h)
# We'll also replace n_input with the current n_output, so that on the
# next iteration, our new number inputs will be correct.
n_input = n_output
Y = current_input
return Y
Y = decode(z, decoder_dimensions, decoder_Ws)
```
Let's take a look at the new operations we've just added. They will all be prefixed by "decoder" so we can use list comprehension to help us with this:
```
[op.name for op in tf.get_default_graph().get_operations()
if op.name.startswith('decoder')]
```
And let's take a look at the output of the autoencoder:
```
Y.get_shape().as_list()
```
Great! So we should have a synthesized version of our input placeholder, `X`, inside of `Y`. This `Y` is the result of many matrix multiplications, first a series of multiplications in our encoder all the way down to 2 dimensions, and then back to the original dimensions through our decoder. Let's now create a pixel-to-pixel measure of error. This should measure the difference in our synthesized output, `Y`, and our input, `X`. You can use the $l_1$ or $l_2$ norm, just like in assignment 2. If you don't remember, go back to homework 2 where we calculated the cost function and try the same idea here.
<h3><font color='red'>TODO! COMPLETE THIS SECTION!</font></h3>
```
# Calculate some measure of loss, e.g. the pixel to pixel absolute difference or squared difference
loss = ...
# Now sum over every pixel and then calculate the mean over the batch dimension (just like session 2!)
# hint, use tf.reduce_mean and tf.reduce_sum
cost = ...
```
Now for the standard training code. We'll pass our `cost` to an optimizer, and then use mini batch gradient descent to optimize our network's parameters. We just have to be careful to make sure we're preprocessing our input and feed it in the right shape, a 2-dimensional matrix of [batch_size, n_features] in dimensions.
<h3><font color='red'>TODO! COMPLETE THIS SECTION!</font></h3>
```
learning_rate = ...
optimizer = tf.train.AdamOptimizer...
```
Below is the training code for our autoencoder. Please go through each line of code to make sure you understand what is happening, and fill in the missing pieces. This will take awhile. On my machine, it takes about 15 minutes. If you're impatient, you can "Interrupt" the kernel by going to the Kernel menu above, and continue with the notebook. Though, the longer you leave this to train, the better the result will be.
What I really want you to notice is what the network learns to encode first, based on what it is able to reconstruct. It won't able to reconstruct everything. At first, it will just be the mean image. Then, other major changes in the dataset. For the first 100 images of celeb net, this seems to be the background: white, blue, black backgrounds. From this basic interpretation, you can reason that the autoencoder has learned a representation of the backgrounds, and is able to encode that knowledge of the background in its inner most layer of just two values. It then goes on to represent the major variations in skin tone and hair. Then perhaps some facial features such as lips. So the features it is able to encode tend to be the major things at first, then the smaller things.
<h3><font color='red'>TODO! COMPLETE THIS SECTION!</font></h3>
```
# (TODO) Create a tensorflow session and initialize all of our weights:
sess = ...
sess.run(tf.initialize_all_variables())
# Some parameters for training
batch_size = 100
n_epochs = 31
step = 10
# We'll try to reconstruct the same first 100 images and show how
# The network does over the course of training.
examples = ds.X[:100]
# We have to preprocess the images before feeding them to the network.
# I'll do this once here, so we don't have to do it every iteration.
test_examples = preprocess(examples, ds).reshape(-1, n_features)
# If we want to just visualize them, we can create a montage.
test_images = utils.montage(examples).astype(np.uint8)
# Store images so we can make a gif
gifs = []
# Now for our training:
for epoch_i in range(n_epochs):
# Keep track of the cost
this_cost = 0
# Iterate over the entire dataset in batches
for batch_X, _ in ds.train.next_batch(batch_size=batch_size):
# (TODO) Preprocess and reshape our current batch, batch_X:
this_batch = preprocess(..., ds).reshape(-1, n_features)
# Compute the cost, and run the optimizer.
this_cost += sess.run([cost, optimizer], feed_dict={X: this_batch})[0]
# Average cost of this epoch
avg_cost = this_cost / ds.X.shape[0] / batch_size
print(epoch_i, avg_cost)
# Let's also try to see how the network currently reconstructs the input.
# We'll draw the reconstruction every `step` iterations.
if epoch_i % step == 0:
# (TODO) Ask for the output of the network, Y, and give it our test examples
recon = sess.run(...
# Resize the 2d to the 4d representation:
rsz = recon.reshape(examples.shape)
# We have to unprocess the image now, removing the normalization
unnorm_img = deprocess(rsz, ds)
# Clip to avoid saturation
clipped = np.clip(unnorm_img, 0, 255)
# And we can create a montage of the reconstruction
recon = utils.montage(clipped).astype(np.uint8)
# Store for gif
gifs.append(recon)
fig, axs = plt.subplots(1, 2, figsize=(10, 10))
axs[0].imshow(test_images)
axs[0].set_title('Original')
axs[1].imshow(recon)
axs[1].set_title('Synthesis')
fig.canvas.draw()
plt.show()
```
Let's take a look a the final reconstruction:
```
fig, axs = plt.subplots(1, 2, figsize=(10, 10))
axs[0].imshow(test_images)
axs[0].set_title('Original')
axs[1].imshow(recon)
axs[1].set_title('Synthesis')
fig.canvas.draw()
plt.show()
plt.imsave(arr=test_images, fname='test.png')
plt.imsave(arr=recon, fname='recon.png')
```
<a name="visualize-the-embedding"></a>
## Visualize the Embedding
Let's now try visualizing our dataset's inner most layer's activations. Since these are already 2-dimensional, we can use the values of this layer to position any input image in a 2-dimensional space. We hope to find similar looking images closer together.
We'll first ask for the inner most layer's activations when given our example images. This will run our images through the network, half way, stopping at the end of the encoder part of the network.
```
zs = sess.run(z, feed_dict={X:test_examples})
```
Recall that this layer has 2 neurons:
```
zs.shape
```
Let's see what the activations look like for our 100 images as a scatter plot.
```
plt.scatter(zs[:, 0], zs[:, 1])
```
If you view this plot over time, and let the process train longer, you will see something similar to the visualization here on the right: https://vimeo.com/155061675 - the manifold is able to express more and more possible ideas, or put another way, it is able to encode more data. As it grows more expressive, with more data, and longer training, or deeper networks, it will fill in more of the space, and have different modes expressing different clusters of the data. With just 100 examples of our dataset, this is *very* small to try to model with such a deep network. In any case, the techniques we've learned up to now apply in exactly the same way, even if we had 1k, 100k, or even many millions of images.
Let's try to see how this minimal example, with just 100 images, and just 100 epochs looks when we use this embedding to sort our dataset, just like we tried to do in the 1st assignment, but now with our autoencoders embedding.
<a name="reorganize-to-grid"></a>
## Reorganize to Grid
We'll use these points to try to find an assignment to a grid. This is a well-known problem known as the "assignment problem": https://en.wikipedia.org/wiki/Assignment_problem - This is unrelated to the applications we're investigating in this course, but I thought it would be a fun extra to show you how to do. What we're going to do is take our scatter plot above, and find the best way to stretch and scale it so that each point is placed in a grid. We try to do this in a way that keeps nearby points close together when they are reassigned in their grid.
```
n_images = 100
idxs = np.linspace(np.min(zs) * 2.0, np.max(zs) * 2.0,
int(np.ceil(np.sqrt(n_images))))
xs, ys = np.meshgrid(idxs, idxs)
grid = np.dstack((ys, xs)).reshape(-1, 2)[:n_images,:]
fig, axs = plt.subplots(1,2,figsize=(8,3))
axs[0].scatter(zs[:, 0], zs[:, 1],
edgecolors='none', marker='o', s=2)
axs[0].set_title('Autoencoder Embedding')
axs[1].scatter(grid[:,0], grid[:,1],
edgecolors='none', marker='o', s=2)
axs[1].set_title('Ideal Grid')
```
To do this, we can use scipy and an algorithm for solving this assignment problem known as the hungarian algorithm. With a few points, this algorithm runs pretty fast. But be careful if you have many more points, e.g. > 1000, as it is not a very efficient algorithm!
```
from scipy.spatial.distance import cdist
cost = cdist(grid[:, :], zs[:, :], 'sqeuclidean')
from scipy.optimize._hungarian import linear_sum_assignment
indexes = linear_sum_assignment(cost)
```
The result tells us the matching indexes from our autoencoder embedding of 2 dimensions, to our idealized grid:
```
indexes
plt.figure(figsize=(5, 5))
for i in range(len(zs)):
plt.plot([zs[indexes[1][i], 0], grid[i, 0]],
[zs[indexes[1][i], 1], grid[i, 1]], 'r')
plt.xlim([-3, 3])
plt.ylim([-3, 3])
```
In other words, this algorithm has just found the best arrangement of our previous `zs` as a grid. We can now plot our images using the order of our assignment problem to see what it looks like:
```
examples_sorted = []
for i in indexes[1]:
examples_sorted.append(examples[i])
plt.figure(figsize=(15, 15))
img = utils.montage(np.array(examples_sorted)).astype(np.uint8)
plt.imshow(img,
interpolation='nearest')
plt.imsave(arr=img, fname='sorted.png')
```
<a name="2d-latent-manifold"></a>
## 2D Latent Manifold
We'll now explore the inner most layer of the network. Recall we go from the number of image features (the number of pixels), down to 2 values using successive matrix multiplications, back to the number of image features through more matrix multiplications. These inner 2 values are enough to represent our entire dataset (+ some loss, depending on how well we did). Let's explore how the decoder, the second half of the network, operates, from just these two values. We'll bypass the input placeholder, X, and the entire encoder network, and start from Z. Let's first get some data which will sample Z in 2 dimensions from -1 to 1. Then we'll feed these values through the decoder network to have our synthesized images.
```
# This is a quick way to do what we could have done as
# a nested for loop:
zs = np.meshgrid(np.linspace(-1, 1, 10),
np.linspace(-1, 1, 10))
# Now we have 100 x 2 values of every possible position
# in a 2D grid from -1 to 1:
zs = np.c_[zs[0].ravel(), zs[1].ravel()]
```
Now calculate the reconstructed images using our new zs. You'll want to start from the beginning of the decoder! That is the `z` variable! Then calculate the `Y` given our synthetic values for `z` stored in `zs`.
<h3><font color='red'>TODO! COMPLETE THIS SECTION!</font></h3>
```
recon = sess.run(Y, feed_dict={...})
# reshape the result to an image:
rsz = recon.reshape(examples.shape)
# Deprocess the result, unnormalizing it
unnorm_img = deprocess(rsz, ds)
# clip to avoid saturation
clipped = np.clip(unnorm_img, 0, 255)
# Create a montage
img_i = utils.montage(clipped).astype(np.uint8)
```
And now we can plot the reconstructed montage representing our latent space:
```
plt.figure(figsize=(15, 15))
plt.imshow(img_i)
plt.imsave(arr=img_i, fname='manifold.png')
```
<a name="part-two---general-autoencoder-framework"></a>
# Part Two - General Autoencoder Framework
There are a number of extensions we can explore w/ an autoencoder. I've provided a module under the libs folder, `vae.py`, which you will need to explore for Part Two. It has a function, `VAE`, to create an autoencoder, optionally with Convolution, Denoising, and/or Variational Layers. Please read through the documentation and try to understand the different parameters.
```
help(vae.VAE)
```
Included in the `vae.py` module is the `train_vae` function. This will take a list of file paths, and train an autoencoder with the provided options. This will spit out a bunch of images of the reconstruction and latent manifold created by the encoder/variational encoder. Feel free to read through the code, as it is documented.
```
help(vae.train_vae)
```
I've also included three examples of how to use the `VAE(...)` and `train_vae(...)` functions. First look at the one using MNIST. Then look at the other two: one using the Celeb Dataset; and lastly one which will download Sita Sings the Blues, rip the frames, and train a Variational Autoencoder on it. This last one requires `ffmpeg` be installed (e.g. for OSX users, `brew install ffmpeg`, Linux users, `sudo apt-get ffmpeg-dev`, or else: https://ffmpeg.org/download.html). The Celeb and Sita Sings the Blues training require us to use an image pipeline, which I've mentioned briefly during the lecture. This does many things for us: it loads data from disk in batches, decodes the data as an image, resizes/crops the image, and uses a multithreaded graph to handle it all. It is *very* efficient and is the way to go when handling large image datasets.
The MNIST training does not use this. Instead, the entire dataset is loaded into the CPU memory, and then fed in minibatches to the graph using Python/Numpy. This is far less efficient, but will not be an issue for such a small dataset, e.g. 70k examples of 28x28 pixels = ~1.6 MB of data, easily fits into memory (in fact, it would really be better to use a Tensorflow variable with this entire dataset defined). When you consider the Celeb Net, you have 200k examples of 218x178x3 pixels = ~700 MB of data. That's just for the dataset. When you factor in everything required for the network and its weights, then you are pushing it. Basically this image pipeline will handle loading the data from disk, rather than storing it in memory.
<a name="instructions-1"></a>
## Instructions
You'll now try to train your own autoencoder using this framework. You'll need to get a directory full of 'jpg' files. You'll then use the VAE framework and the `vae.train_vae` function to train a variational autoencoder on your own dataset. This accepts a list of files, and will output images of the training in the same directory. These are named "test_xs.png" as well as many images named prefixed by "manifold" and "reconstruction" for each iteration of the training. After you are happy with your training, you will need to create a forum post with the "test_xs.png" and the very last manifold and reconstruction image created to demonstrate how the variational autoencoder worked for your dataset. You'll likely need a lot more than 100 images for this to be successful.
Note that this will also create "checkpoints" which save the model! If you change the model, and already have a checkpoint by the same name, it will try to load the previous model and will fail. Be sure to remove the old checkpoint or specify a new name for `ckpt_name`! The default parameters shown below are what I have used for the celeb net dataset which has over 200k images. You will definitely want to use a smaller model if you do not have this many images! Explore!
<h3><font color='red'>TODO! COMPLETE THIS SECTION!</font></h3>
```
# Get a list of jpg file (Only JPG works!)
files = [os.path.join(some_dir, file_i) for file_i in os.listdir(some_dir) if file_i.endswith('.jpg')]
# Train it! Change these parameters!
vae.train_vae(files,
input_shape,
learning_rate=0.0001,
batch_size=100,
n_epochs=50,
n_examples=10,
crop_shape=[64, 64, 3],
crop_factor=0.8,
n_filters=[100, 100, 100, 100],
n_hidden=256,
n_code=50,
convolutional=True,
variational=True,
filter_sizes=[3, 3, 3, 3],
dropout=True,
keep_prob=0.8,
activation=tf.nn.relu,
img_step=100,
save_step=100,
ckpt_name="vae.ckpt")
```
<a name="part-three---deep-audio-classification-network"></a>
# Part Three - Deep Audio Classification Network
<a name="instructions-2"></a>
## Instructions
In this last section, we'll explore using a regression network, one that predicts continuous outputs, to perform classification, a model capable of predicting discrete outputs. We'll explore the use of one-hot encodings and using a softmax layer to convert our regression outputs to a probability which we can use for classification. In the lecture, we saw how this works for the MNIST dataset, a dataset of 28 x 28 pixel handwritten digits labeled from 0 - 9. We converted our 28 x 28 pixels into a vector of 784 values, and used a fully connected network to output 10 values, the one hot encoding of our 0 - 9 labels.
In addition to the lecture material, I find these two links very helpful to try to understand classification w/ neural networks:
https://colah.github.io/posts/2014-03-NN-Manifolds-Topology/
https://cs.stanford.edu/people/karpathy/convnetjs//demo/classify2d.html
The GTZAN Music and Speech dataset has 64 music and 64 speech files, each 30 seconds long, and each at a sample rate of 22050 Hz, meaning there are 22050 samplings of the audio signal per second. What we're going to do is use all of this data to build a classification network capable of knowing whether something is music or speech. So we will have audio as input, and a probability of 2 possible values, music and speech, as output. This is *very* similar to the MNIST network. We just have to decide on how to represent our input data, prepare the data and its labels, build batch generators for our data, create the network, and train it. We'll make use of the `libs/datasets.py` module to help with some of this.
<a name="preparing-the-data"></a>
## Preparing the Data
Let's first download the GTZAN music and speech dataset. I've included a helper function to do this.
```
dst = 'gtzan_music_speech'
if not os.path.exists(dst):
dataset_utils.gtzan_music_speech_download(dst)
```
Inside the `dst` directory, we now have folders for music and speech. Let's get the list of all the wav files for music and speech:
```
# Get the full path to the directory
music_dir = os.path.join(os.path.join(dst, 'music_speech'), 'music_wav')
# Now use list comprehension to combine the path of the directory with any wave files
music = [os.path.join(music_dir, file_i)
for file_i in os.listdir(music_dir)
if file_i.endswith('.wav')]
# Similarly, for the speech folder:
speech_dir = os.path.join(os.path.join(dst, 'music_speech'), 'speech_wav')
speech = [os.path.join(speech_dir, file_i)
for file_i in os.listdir(speech_dir)
if file_i.endswith('.wav')]
# Let's see all the file names
print(music, speech)
```
We now need to load each file. We can use the `scipy.io.wavefile` module to load the audio as a signal.
Audio can be represented in a few ways, including as floating point or short byte data (16-bit data). This dataset is the latter and so can range from -32768 to +32767. We'll use the function I've provided in the utils module to load and convert an audio signal to a -1.0 to 1.0 floating point datatype by dividing by the maximum absolute value. Let's try this with just one of the files we have:
```
file_i = music[0]
s = utils.load_audio(file_i)
plt.plot(s)
```
Now, instead of using the raw audio signal, we're going to use the [Discrete Fourier Transform](https://en.wikipedia.org/wiki/Discrete_Fourier_transform) to represent our audio as matched filters of different sinuoids. Unfortunately, this is a class on Tensorflow and I can't get into Digital Signal Processing basics. If you want to know more about this topic, I highly encourage you to take this course taught by the legendary Perry Cook and Julius Smith: https://www.kadenze.com/courses/physics-based-sound-synthesis-for-games-and-interactive-systems/info - there is no one better to teach this content, and in fact, I myself learned DSP from Perry Cook almost 10 years ago.
After taking the DFT, this will return our signal as real and imaginary components, a polar complex value representation which we will convert to a cartesian representation capable of saying what magnitudes and phases are in our signal.
```
# Parameters for our dft transform. Sorry we can't go into the
# details of this in this course. Please look into DSP texts or the
# course by Perry Cook linked above if you are unfamiliar with this.
fft_size = 512
hop_size = 256
re, im = dft.dft_np(s, hop_size=256, fft_size=512)
mag, phs = dft.ztoc(re, im)
print(mag.shape)
plt.imshow(mag)
```
What we're seeing are the features of the audio (in columns) over time (in rows). We can see this a bit better by taking the logarithm of the magnitudes converting it to a psuedo-decibel scale. This is more similar to the logarithmic perception of loudness we have. Let's visualize this below, and I'll transpose the matrix just for display purposes:
```
plt.figure(figsize=(10, 4))
plt.imshow(np.log(mag.T))
plt.xlabel('Time')
plt.ylabel('Frequency Bin')
```
We could just take just a single row (or column in the second plot of the magnitudes just above, as we transposed it in that plot) as an input to a neural network. However, that just represents about an 80th of a second of audio data, and is not nearly enough data to say whether something is music or speech. We'll need to use more than a single row to get a decent length of time. One way to do this is to use a sliding 2D window from the top of the image down to the bottom of the image (or left to right). Let's start by specifying how large our sliding window is.
```
# The sample rate from our audio is 22050 Hz.
sr = 22050
# We can calculate how many hops there are in a second
# which will tell us how many frames of magnitudes
# we have per second
n_frames_per_second = sr // hop_size
# We want 500 milliseconds of audio in our window
n_frames = n_frames_per_second // 2
# And we'll move our window by 250 ms at a time
frame_hops = n_frames_per_second // 4
# We'll therefore have this many sliding windows:
n_hops = (len(mag) - n_frames) // frame_hops
```
Now we can collect all the sliding windows into a list of `Xs` and label them based on being music as `0` or speech as `1` into a collection of `ys`.
```
Xs = []
ys = []
for hop_i in range(n_hops):
# Creating our sliding window
frames = mag[(hop_i * frame_hops):(hop_i * frame_hops + n_frames)]
# Store them with a new 3rd axis and as a logarithmic scale
# We'll ensure that we aren't taking a log of 0 just by adding
# a small value, also known as epsilon.
Xs.append(np.log(np.abs(frames[..., np.newaxis]) + 1e-10))
# And then store the label
ys.append(0)
```
The code below will perform this for us, as well as create the inputs and outputs to our classification network by specifying 0s for the music dataset and 1s for the speech dataset. Let's just take a look at the first sliding window, and see it's label:
```
plt.imshow(Xs[0][..., 0])
plt.title('label:{}'.format(ys[1]))
```
Since this was the first audio file of the music dataset, we've set it to a label of 0. And now the second one, which should have 50% overlap with the previous one, and still a label of 0:
```
plt.imshow(Xs[1][..., 0])
plt.title('label:{}'.format(ys[1]))
```
So hopefully you can see that the window is sliding down 250 milliseconds at a time, and since our window is 500 ms long, or half a second, it has 50% new content at the bottom. Let's do this for every audio file now:
<h3><font color='red'>TODO! COMPLETE THIS SECTION!</font></h3>
```
# Store every magnitude frame and its label of being music: 0 or speech: 1
Xs, ys = [], []
# Let's start with the music files
for i in music:
# Load the ith file:
s = utils.load_audio(i)
# Now take the dft of it (take a DSP course!):
re, im = dft.dft_np(s, fft_size=fft_size, hop_size=hop_size)
# And convert the complex representation to magnitudes/phases (take a DSP course!):
mag, phs = dft.ztoc(re, im)
# This is how many sliding windows we have:
n_hops = (len(mag) - n_frames) // frame_hops
# Let's extract them all:
for hop_i in range(n_hops):
# Get the current sliding window
frames = mag[(hop_i * frame_hops):(hop_i * frame_hops + n_frames)]
# We'll take the log magnitudes, as this is a nicer representation:
this_X = np.log(np.abs(frames[..., np.newaxis]) + 1e-10)
# And store it:
Xs.append(this_X)
# And be sure that we store the correct label of this observation:
ys.append(0)
# Now do the same thing with speech (TODO)!
for i in speech:
# Load the ith file:
s = ...
# Now take the dft of it (take a DSP course!):
re, im = ...
# And convert the complex representation to magnitudes/phases (take a DSP course!):
mag, phs = ...
# This is how many sliding windows we have:
n_hops = (len(mag) - n_frames) // frame_hops
# Let's extract them all:
for hop_i in range(n_hops):
# Get the current sliding window
frames = mag[(hop_i * frame_hops):(hop_i * frame_hops + n_frames)]
# We'll take the log magnitudes, as this is a nicer representation:
this_X = np.log(np.abs(frames[..., np.newaxis]) + 1e-10)
# And store it:
Xs.append(this_X)
# Make sure we use the right label (TODO!)!
ys.append...
# Convert them to an array:
Xs = np.array(Xs)
ys = np.array(ys)
print(Xs.shape, ys.shape)
# Just to make sure you've done it right. If you've changed any of the
# parameters of the dft/hop size, then this will fail. If that's what you
# wanted to do, then don't worry about this assertion.
assert(Xs.shape == (15360, 43, 256, 1) and ys.shape == (15360,))
```
Just to confirm it's doing the same as above, let's plot the first magnitude matrix:
```
plt.imshow(Xs[0][..., 0])
plt.title('label:{}'.format(ys[0]))
```
Let's describe the shape of our input to the network:
```
n_observations, n_height, n_width, n_channels = Xs.shape
```
We'll now use the `Dataset` object I've provided for you under `libs/datasets.py`. This will accept the `Xs`, `ys`, a list defining our dataset split into training, validation, and testing proportions, and a parameter `one_hot` stating whether we want our `ys` to be converted to a one hot vector or not.
<h3><font color='red'>TODO! COMPLETE THIS SECTION!</font></h3>
```
ds = datasets.Dataset(Xs=..., ys=..., split=[0.8, 0.1, 0.1], one_hot=True)
```
Let's take a look at the batch generator this object provides. We can all any of the splits, the `train`, `valid`, or `test` splits as properties of the object. And each split provides a `next_batch` method which gives us a batch generator. We should have specified that we wanted `one_hot=True` to have our batch generator return our ys with 2 features, one for each possible class.
```
Xs_i, ys_i = next(ds.train.next_batch())
# Notice the shape this returns. This will become the shape of our input and output of the network:
print(Xs_i.shape, ys_i.shape)
assert(ys_i.shape == (100, 2))
```
Let's take a look at the first element of the randomized batch:
```
plt.imshow(Xs_i[0, :, :, 0])
plt.title('label:{}'.format(ys_i[0]))
```
And the second one:
```
plt.imshow(Xs_i[1, :, :, 0])
plt.title('label:{}'.format(ys_i[1]))
```
So we have a randomized order in minibatches generated for us, and the `ys` are represented as a one-hot vector with each class, music and speech, encoded as a 0 or 1. Since the `next_batch` method is a generator, we can use it in a loop until it is exhausted to run through our entire dataset in mini-batches.
<a name="creating-the-network"></a>
## Creating the Network
Let's now create the neural network. Recall our input `X` is 4-dimensional, with the same shape that we've just seen as returned from our batch generator above. We're going to create a deep convolutional neural network with a few layers of convolution and 2 finals layers which are fully connected. The very last layer must have only 2 neurons corresponding to our one-hot vector of `ys`, so that we can properly measure the cross-entropy (just like we did with MNIST and our 10 element one-hot encoding of the digit label). First let's create our placeholders:
<h3><font color='red'>TODO! COMPLETE THIS SECTION!</font></h3>
```
tf.reset_default_graph()
# Create the input to the network. This is a 4-dimensional tensor!
# Recall that we are using sliding windows of our magnitudes (TODO):
X = tf.placeholder(name='X', shape=..., dtype=tf.float32)
# Create the output to the network. This is our one hot encoding of 2 possible values (TODO)!
Y = tf.placeholder(name='Y', shape=..., dtype=tf.float32)
```
Let's now create our deep convolutional network. Start by first creating the convolutional layers. Try different numbers of layers, different numbers of filters per layer, different activation functions, and varying the parameters to get the best training/validation score when training below. Try first using a kernel size of `3` and a stride of `1`. You can use the `utils.conv2d` function to help you create the convolution.
<h3><font color='red'>TODO! COMPLETE THIS SECTION!</font></h3>
```
# TODO: Explore different numbers of layers, and sizes of the network
n_filters = [9, 9, 9, 9]
# Now let's loop over our n_filters and create the deep convolutional neural network
H = X
for layer_i, n_filters_i in enumerate(n_filters):
# Let's use the helper function to create our connection to the next layer:
# TODO: explore changing the parameters here:
H, W = utils.conv2d(
H, n_filters_i, k_h=3, k_w=3, d_h=2, d_w=2,
name=str(layer_i))
# And use a nonlinearity
# TODO: explore changing the activation here:
H = tf.nn.relu(H)
# Just to check what's happening:
print(H.get_shape().as_list())
```
We'll now connect our last convolutional layer to a fully connected layer of 100 neurons. This is essentially combining the spatial information, thus losing the spatial information. You can use the `utils.linear` function to do this, which will internally also reshape the 4-d tensor to a 2-d tensor so that it can be connected to a fully-connected layer (i.e. perform a matrix multiplication).
<h3><font color='red'>TODO! COMPLETE THIS SECTION!</font></h3>
```
# Connect the last convolutional layer to a fully connected network (TODO)!
fc, W = utils.linear(H, ...
# And another fully connceted network, now with just 2 outputs, the number of outputs that our
# one hot encoding has (TODO)!
Y_pred, W = utils.linear(fc, ...
```
We'll now create our cost. Unlike the MNIST network, we're going to use a binary cross entropy as we only have 2 possible classes. You can use the `utils.binary_cross_entropy` function to help you with this. Remember, the final cost measure the average loss of your batches.
```
loss = utils.binary_cross_entropy(Y_pred, Y)
cost = tf.reduce_mean(tf.reduce_sum(loss, 1))
```
Just like in MNIST, we'll now also create a measure of accuracy by finding the prediction of our network. This is just for us to monitor the training and is not used to optimize the weights of the network! Look back to the MNIST network in the lecture if you are unsure of how this works (it is exactly the same):
<h3><font color='red'>TODO! COMPLETE THIS SECTION!</font></h3>
```
predicted_y = tf.argmax(...
actual_y = tf.argmax(...
correct_prediction = tf.equal(...
accuracy = tf.reduce_mean(...
```
We'll now create an optimizer and train our network:
<h3><font color='red'>TODO! COMPLETE THIS SECTION!</font></h3>
```
learning_rate = ...
optimizer = tf.train.AdamOptimizer(...
```
Now we're ready to train. This is a pretty simple dataset for a deep convolutional network. As a result, I've included code which demonstrates how to monitor validation performance. A validation set is data that the network has never seen, and is not used for optimizing the weights of the network. We use validation to better understand how well the performance of a network "generalizes" to unseen data.
You can easily run the risk of [overfitting](https://en.wikipedia.org/wiki/Overfitting) to the training set of this problem. Overfitting simply means that the number of parameters in our model are so high that we are not generalizing our model, and instead trying to model each individual point, rather than the general cause of the data. This is a very common problem that can be addressed by using less parameters, or enforcing regularization techniques which we didn't have a chance to cover (dropout, batch norm, l2, augmenting the dataset, and others).
For this dataset, if you notice that your validation set is performing worse than your training set, then you know you have overfit! You should be able to easily get 97+% on the validation set within < 10 epochs. If you've got great training performance, but poor validation performance, then you likely have "overfit" to the training dataset, and are unable to generalize to the validation set. Try varying the network definition, number of filters/layers until you get 97+% on your validation set!
<h3><font color='red'>TODO! COMPLETE THIS SECTION!</font></h3>
```
# Explore these parameters: (TODO)
n_epochs = 10
batch_size = 200
# Create a session and init!
sess = tf.Session()
sess.run(tf.initialize_all_variables())
# Now iterate over our dataset n_epoch times
for epoch_i in range(n_epochs):
print('Epoch: ', epoch_i)
# Train
this_accuracy = 0
its = 0
# Do our mini batches:
for Xs_i, ys_i in ds.train.next_batch(batch_size):
# Note here: we are running the optimizer so
# that the network parameters train!
this_accuracy += sess.run([accuracy, optimizer], feed_dict={
X:Xs_i, Y:ys_i})[0]
its += 1
print(this_accuracy / its)
print('Training accuracy: ', this_accuracy / its)
# Validation (see how the network does on unseen data).
this_accuracy = 0
its = 0
# Do our mini batches:
for Xs_i, ys_i in ds.valid.next_batch(batch_size):
# Note here: we are NOT running the optimizer!
# we only measure the accuracy!
this_accuracy += sess.run(accuracy, feed_dict={
X:Xs_i, Y:ys_i})
its += 1
print('Validation accuracy: ', this_accuracy / its)
```
Let's try to inspect how the network is accomplishing this task, just like we did with the MNIST network. First, let's see what the names of our operations in our network are.
```
g = tf.get_default_graph()
[op.name for op in g.get_operations()]
```
Now let's visualize the `W` tensor's weights for the first layer using the utils function `montage_filters`, just like we did for the MNIST dataset during the lecture. Recall from the lecture that this is another great way to inspect the performance of your network. If many of the filters look uniform, then you know the network is either under or overperforming. What you want to see are filters that look like they are responding to information such as edges or corners.
<h3><font color='red'>TODO! COMPLETE THIS SECTION!</font></h3>
```
g = tf.get_default_graph()
W = ...
assert(W.dtype == np.float32)
m = montage_filters(W)
plt.figure(figsize=(5, 5))
plt.imshow(m)
plt.imsave(arr=m, fname='audio.png')
```
We can also look at every layer's filters using a loop:
```
g = tf.get_default_graph()
for layer_i in range(len(n_filters)):
W = sess.run(g.get_tensor_by_name('{}/W:0'.format(layer_i)))
plt.figure(figsize=(5, 5))
plt.imshow(montage_filters(W))
plt.title('Layer {}\'s Learned Convolution Kernels'.format(layer_i))
```
In the next session, we'll learn some much more powerful methods of inspecting such networks.
<a name="assignment-submission"></a>
# Assignment Submission
After you've completed the notebook, create a zip file of the current directory using the code below. This code will make sure you have included this completed ipython notebook and the following files named exactly as:
<pre>
session-3/
session-3.ipynb
test.png
recon.png
sorted.png
manifold.png
test_xs.png
audio.png
</pre>
You'll then submit this zip file for your third assignment on Kadenze for "Assignment 3: Build Unsupervised and Supervised Networks"! Remember to post Part Two to the Forum to receive full credit! If you have any questions, remember to reach out on the forums and connect with your peers or with me.
To get assessed, you'll need to be a premium student! This will allow you to build an online portfolio of all of your work and receive grades. If you aren't already enrolled as a student, register now at http://www.kadenze.com/ and join the [#CADL](https://twitter.com/hashtag/CADL) community to see what your peers are doing! https://www.kadenze.com/courses/creative-applications-of-deep-learning-with-tensorflow/info
Also, if you share any of the GIFs on Facebook/Twitter/Instagram/etc..., be sure to use the #CADL hashtag so that other students can find your work!
```
utils.build_submission('session-3.zip',
('test.png',
'recon.png',
'sorted.png',
'manifold.png',
'test_xs.png',
'audio.png',
'session-3.ipynb'))
```
<a name="coming-up"></a>
# Coming Up
In session 4, we'll start to interrogate pre-trained Deep Convolutional Networks trained to recognize 1000 possible object labels. Along the way, we'll see how by inspecting the network, we can perform some very interesting image synthesis techniques which led to the Deep Dream viral craze. We'll also see how to separate the content and style of an image and use this for generative artistic stylization! In Session 5, we'll explore a few other powerful methods of generative synthesis, including Generative Adversarial Networks, Variational Autoencoding Generative Adversarial Networks, and Recurrent Neural Networks.
| github_jupyter |
```
import numpy as np
from data_generator import AudioGenerator
from keras import backend as K
from utils import int_sequence_to_text
from IPython.display import Audio
def get_predictions(index, partition, input_to_softmax, model_path):
""" Print a model's decoded predictions
Params:
index (int): The example you would like to visualize
partition (str): One of 'train' or 'validation'
input_to_softmax (Model): The acoustic model
model_path (str): Path to saved acoustic model's weights
"""
# load the train and test data
data_gen = AudioGenerator()
data_gen.load_train_data()
data_gen.load_validation_data()
# obtain the true transcription and the audio features
if partition == 'validation':
transcr = data_gen.valid_texts[index]
audio_path = data_gen.valid_audio_paths[index]
data_point = data_gen.normalize(data_gen.featurize(audio_path))
elif partition == 'train':
transcr = data_gen.train_texts[index]
audio_path = data_gen.train_audio_paths[index]
data_point = data_gen.normalize(data_gen.featurize(audio_path))
else:
raise Exception('Invalid partition! Must be "train" or "validation"')
# obtain and decode the acoustic model's predictions
input_to_softmax.load_weights(model_path)
prediction = input_to_softmax.predict(np.expand_dims(data_point, axis=0))
output_length = [input_to_softmax.output_length(data_point.shape[0])]
pred_ints = (K.eval(K.ctc_decode(
prediction, output_length)[0][0])+1).flatten().tolist()
# play the audio file, and display the true and predicted transcriptions
print('-'*80)
Audio(audio_path)
print('True transcription:\n' + '\n' + transcr)
print('-'*80)
print('Predicted transcription:\n' + '\n' + ''.join(int_sequence_to_text(pred_ints)))
print('-'*80)
from keras import backend as K
from keras.models import Model
from keras.layers import (BatchNormalization, Conv1D, Dense, Input,
TimeDistributed, Activation, Bidirectional, SimpleRNN, GRU, LSTM)
def simple_rnn_model(input_dim, output_dim=29):
""" Build a recurrent network for speech
"""
# Main acoustic input
input_data = Input(name='the_input', shape=(None, input_dim))
# Add recurrent layer
simp_rnn = GRU(output_dim, return_sequences=True,
implementation=2, name='rnn')(input_data)
# Add softmax activation layer
y_pred = Activation('softmax', name='softmax')(simp_rnn)
# Specify the model
model = Model(inputs=input_data, outputs=y_pred)
model.output_length = lambda x: x
print(model.summary())
return model
model_0 = simple_rnn_model(input_dim=161) # change to 13 if you would like to use MFCC features
get_predictions(index=0,
partition='train',
input_to_softmax=model_0,
model_path='results/model_0.h5')
```
| github_jupyter |
# CLEAN & PREPROCESS TEXT DATA
## 1. Import libraries
**Import neccessary packages and modules**
```
import time
import os
t = time.time()
import json
import string
import random
import math
import random
import pandas as pd
import numpy as np
from sklearn.preprocessing import LabelBinarizer
import tensorflow as tf
import matplotlib
from matplotlib import pyplot as plt
%matplotlib inline
```
**Import nlp packages and modules**
```
import nltk
# nltk.download()
import nltk, re, time
from nltk.corpus import stopwords
from nltk.stem.snowball import SnowballStemmer
```
## 2. Load and Inspect Data
**Set directory**
```
input_dir = "../input/"
```
**Load train and test data**
```
train_data = pd.read_csv(input_dir+'train.csv')
test_data = pd.read_csv(input_dir+'test.csv')
```
**Inspect train and test data**
```
train_data.head(20)
print("Shape of train data:", train_data.shape)
test_data.head(20)
print("Shape of test data:", test_data.shape)
```
## 3. Preprocess the text data
```
# load list of stopwords
sw = set(stopwords.words("english"))
# load teh snowball stemmer
stemmer = SnowballStemmer("english")
# translator object to replace punctuations with space
translator = str.maketrans(string.punctuation, ' '*len(string.punctuation))
print(sw)
```
**Function for preprocessing text**
```
def clean_text(text):
"""
A function for preprocessing text
"""
text = str(text)
# replacing the punctuations with no space,which in effect deletes the punctuation marks
text = text.translate(translator)
# remove stop word
text = [word.lower() for word in text.split() if word.lower() not in sw]
text = " ".join(text)
# stemming
text = [stemmer.stem(word) for word in text.split()]
text = " ".join(text)
# Clean the text
text = re.sub(r"<br />", " ", text)
text = re.sub(r"[^a-z]", " ", text)
text = re.sub(r" ", " ", text) # Remove any extra spaces
text = re.sub(r" ", " ", text)
return(text)
```
**Clean train and test data**
```
t1 = time.time()
train_data['comment_text'] = train_data['comment_text'].apply(clean_text)
print("Finished cleaning the train set.", "Time needed:", time.time()-t1,"sec")
t2 = time.time()
test_data['comment_text'] = test_data['comment_text'].apply(clean_text)
print("Finished cleaning the test set.", "Time needed:", time.time()-t2,"sec")
```
**Inspect the cleaned train and test data**
```
train_data.head(10)
test_data.head(10)
```
## 4. Create columns for length of the data
**A function for finding the length of text**
```
def find_length(text):
"""
A function to find the length
"""
text = str(text)
return len(text.split())
```
**Create the column of text length in train and test data**
```
train_data['length'] = train_data['comment_text'].apply(find_length)
train_data.head(10)
test_data['length'] = test_data['comment_text'].apply(find_length)
test_data.head(10)
```
## 5. Save the modified train and test data
**Save directory**
```
save_dir = "../input/"
```
**Save the modified train and test data to the designated directory**
```
train_data.to_csv(save_dir+"modified_train_data.csv",header=True, index=False)
test_data.to_csv(save_dir+"modified_test_data.csv",header=True, index=False)
```
**Inpect the saved train and test csv**
```
pd.read_csv(save_dir+"modified_train_data.csv").head(20)
pd.read_csv(save_dir+"modified_train_data.csv").head(20)
```
| github_jupyter |
# Step 2: Feature Engineering
According to [Wikipedia, Feature engineering](https://en.wikipedia.org/wiki/Feature_engineering) is the process of using domain knowledge of the data to create features that make machine learning algorithms work. Feature engineering is fundamental to the application of machine learning, and is both difficult and expensive.
This Feature engineering notebook will load the data sets created in the **Data Ingestion** notebook (`1_data_ingestion.ipynb`) from an Azure storage container and combine them to create a single data set of features (variables) that can be used to infer a machines's health condition over time. The notebook steps through several feature engineering and labeling methods to create this data set for use in our predictive maintenance machine learning solution.
**Note:** This notebook will take aound 6 minutes to execute all cells on a Standard_DS13_v2 cluster.
```
## Setup our environment by importing required libraries
import time
import os
import glob
# For creating some preliminary EDA plots.
import matplotlib.pyplot as plt
import seaborn as sns
import pandas as pd
plt.style.use('ggplot')
import datetime
import pyspark.sql.functions as F
from pyspark.sql.functions import (col, unix_timestamp,
round, datediff, to_date)
from pyspark.sql.window import Window
from pyspark.sql.types import DoubleType
from pyspark.ml import Pipeline
from pyspark.ml.feature import OneHotEncoder,StringIndexer
from pyspark.sql import SparkSession
# Time the notebook execution.
# This will only make sense if you "Run all cells"
tic = time.time()
```
## Load raw data from databricks workspace storage
In the **Data Ingestion** notebook (`1_data_ingestion.ipynb`), we downloaded, converted and stored the following data sets:
* **Machines**: Features differentiating each machine. For example age and model.
* **Error**: The log of non-critical errors. These errors may still indicate an impending component failure.
* **Maint**: Machine maintenance history detailing component replacement or regular maintenance activities withe the date of replacement.
* **Telemetry**: The operating conditions of a machine e.g. data collected from sensors.
* **Failure**: The failure history of a machine or component within the machine.
We first load these files...
```
parquet_files_names = {'machines':'machines_files.parquet','maint':'maint_files.parquet',
'errors': 'errors_files.parquet','telemetry':'telemetry_files.parquet',
'failures':'failure_files.parquet', 'features':'featureengineering_files.parquet'}
target_dir = "dbfs:/dataset/"
```
### Machines data set
Now, we load the machines data set
```
# Read in the data
machines = spark.read.parquet(os.path.join(target_dir, parquet_files_names['machines']))
print(machines.count())
display(machines.limit(5))
```
### Errors data set
Load the errors data set.
```
errors = spark.read.parquet(os.path.join(target_dir, parquet_files_names['errors']))
print(errors.count())
display(errors.limit(5))
```
### Maintenance data set
Load the maintenance data set.
```
maint = spark.read.parquet(os.path.join(target_dir, parquet_files_names['maint']))
print(maint.count())
display(maint.limit(5))
```
### Telemetry
Load the telemetry data set.
```
telemetry = spark.read.parquet(os.path.join(target_dir, parquet_files_names['telemetry']))
print(telemetry.count())
display(telemetry.limit(5))
```
### Failures data set
Load the failures data set.
```
failures = spark.read.parquet(os.path.join(target_dir, parquet_files_names['failures']))
print(failures.count())
display(failures.limit(5))
```
## Feature engineering
Our feature engineering will combine the different data sources together to create a single data set of features (variables) that can be used to infer a machines's health condition over time. The ultimate goal is to generate a single record for each time unit within each asset. The record combines features and labels to be fed into the machine learning algorithm.
Predictive maintenance take historical data, marked with a timestamp, to predict current health of a component and the probability of failure within some future window of time. These problems can be characterised as a _classification method_ involving _time series_ data. Time series, since we want to use historical observations to predict what will happen in the future. Classification, because we classify the future as having a probability of failure.
### Lag features
There are many ways of creating features from the time series data. We start by dividing the duration of data collection into time units where each record belongs to a single point in time for each asset. The measurement unit for is in fact arbitrary. Time can be in seconds, minutes, hours, days, or months, or it can be measured in cycles, miles or transactions. The measurement choice is typically specific to the use case domain.
Additionally, the time unit does not have to be the same as the frequency of data collection. For example, if temperature values were being collected every 10 seconds, picking a time unit of 10 seconds for analysis may inflate the number of examples without providing any additional information if the temperature changes slowly. A better strategy may be to average the temperature over a longer time horizon which might better capture variations that contribute to the target outcome.
Once we set the frequency of observations, we want to look for trends within measures, over time, in order to predict performance degradation, which we would like to connect to how likely a component will fail. We create features for these trends within each record using time lags over previous observations to check for these performance changes. The lag window size $W$ is a hyper parameter that we can optimize. The following figures indicate a _rolling aggregate window_ strategy for averaging a measure $t_i$ over a window $W = 3$ previous observations.
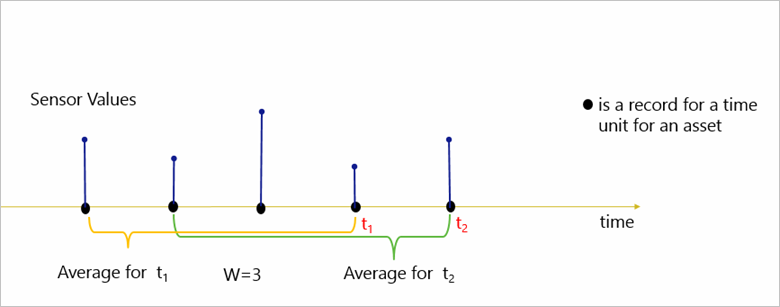
We are note constrained to averages, we can roll aggregates over counts, average, the standard deviation, outliers based on standard deviations, CUSUM measures, minimum and maximum values for the window.
We could also use a tumbling window approach, if we were interested in a different time window measure than the frequncy of the observations. For example, we might have obersvations evert 6 or 12 hours, but want to create features aligned on a day or week basis.
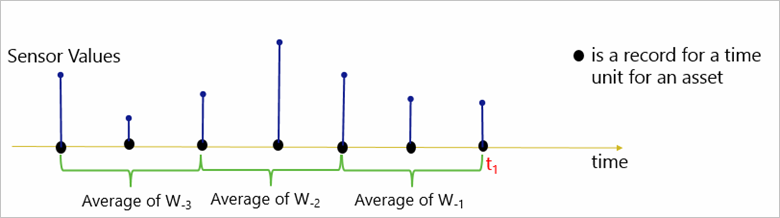
In the following sections, we will build our features using only a rolling strategy to demonstrate the process. We align our data, and then build features along those normalized observations times. We start with the telemetry data.
## Telemetry features
Because the telemetry data set is the largest time series data we have, we start feature engineering here. The telemetry data has 8761000 hourly observations for out 1000 machines. We can improve the model performance by aligning our data by aggregating average sensor measures on a tumbling 12 hour window. In this case we replace the raw data with the tumbling window data, reducing the sensor data to 731000 observations. This will directly reduce the computaton time required to do the feature engineering, labeling and modeling required for our solution.
Once we have the reduced data, we set up our lag features by compute rolling aggregate measures such as mean, standard deviation, minimum, maximum, etc. to represent the short term history of the telemetry over time.
The following code blocks alignes the data on 12 hour observations and calculates a rolling mean and standard deviation of the telemetry data over the last 12, 24 and 36 hour lags.
```
# rolling mean and standard deviation
# Temporary storage for rolling means
tel_mean = telemetry
# Which features are we interested in telemetry data set
rolling_features = ['volt','rotate', 'pressure', 'vibration']
# n hours = n * 3600 seconds
time_val = 12 * 3600
# Choose the time_val hour timestamps to align the data
# dt_truncated looks at the column named "datetime" in the current data set.
# remember that Spark is lazy... this doesn't execute until it is in a withColumn statement.
dt_truncated = ((round(unix_timestamp(col("datetime")) / time_val) * time_val).cast("timestamp"))
# We choose windows for our rolling windows 12hrs, 24 hrs and 36 hrs
lags = [12, 24, 36]
# align the data
for lag_n in lags:
wSpec = Window.partitionBy('machineID').orderBy('datetime').rowsBetween(1-lag_n, 0)
for col_name in rolling_features:
tel_mean = tel_mean.withColumn(col_name+'_rollingmean_'+str(lag_n),
F.avg(col(col_name)).over(wSpec))
tel_mean = tel_mean.withColumn(col_name+'_rollingstd_'+str(lag_n),
F.stddev(col(col_name)).over(wSpec))
# Calculate lag values...
telemetry_feat = (tel_mean.withColumn("dt_truncated", dt_truncated)
.drop('volt', 'rotate', 'pressure', 'vibration')
.fillna(0)
.groupBy("machineID","dt_truncated")
.agg(F.mean('volt_rollingmean_12').alias('volt_rollingmean_12'),
F.mean('rotate_rollingmean_12').alias('rotate_rollingmean_12'),
F.mean('pressure_rollingmean_12').alias('pressure_rollingmean_12'),
F.mean('vibration_rollingmean_12').alias('vibration_rollingmean_12'),
F.mean('volt_rollingmean_24').alias('volt_rollingmean_24'),
F.mean('rotate_rollingmean_24').alias('rotate_rollingmean_24'),
F.mean('pressure_rollingmean_24').alias('pressure_rollingmean_24'),
F.mean('vibration_rollingmean_24').alias('vibration_rollingmean_24'),
F.mean('volt_rollingmean_36').alias('volt_rollingmean_36'),
F.mean('vibration_rollingmean_36').alias('vibration_rollingmean_36'),
F.mean('rotate_rollingmean_36').alias('rotate_rollingmean_36'),
F.mean('pressure_rollingmean_36').alias('pressure_rollingmean_36'),
F.stddev('volt_rollingstd_12').alias('volt_rollingstd_12'),
F.stddev('rotate_rollingstd_12').alias('rotate_rollingstd_12'),
F.stddev('pressure_rollingstd_12').alias('pressure_rollingstd_12'),
F.stddev('vibration_rollingstd_12').alias('vibration_rollingstd_12'),
F.stddev('volt_rollingstd_24').alias('volt_rollingstd_24'),
F.stddev('rotate_rollingstd_24').alias('rotate_rollingstd_24'),
F.stddev('pressure_rollingstd_24').alias('pressure_rollingstd_24'),
F.stddev('vibration_rollingstd_24').alias('vibration_rollingstd_24'),
F.stddev('volt_rollingstd_36').alias('volt_rollingstd_36'),
F.stddev('rotate_rollingstd_36').alias('rotate_rollingstd_36'),
F.stddev('pressure_rollingstd_36').alias('pressure_rollingstd_36'),
F.stddev('vibration_rollingstd_36').alias('vibration_rollingstd_36'), ))
print(telemetry_feat.count())
telemetry_feat.where((col("machineID") == 1)).limit(5).toPandas()
```
## Errors features
Like telemetry data, errors come with timestamps. An important difference is that the error IDs are categorical values and should not be averaged over time intervals like the telemetry measurements. Instead, we count the number of errors of each type within a lag window.
Again, we align the error counts data by tumbling over the 12 hour window using a join with telemetry data.
```
# create a column for each errorID
error_ind = (errors.groupBy("machineID","datetime","errorID").pivot('errorID')
.agg(F.count('machineID').alias('dummy')).drop('errorID').fillna(0)
.groupBy("machineID","datetime")
.agg(F.sum('error1').alias('error1sum'),
F.sum('error2').alias('error2sum'),
F.sum('error3').alias('error3sum'),
F.sum('error4').alias('error4sum'),
F.sum('error5').alias('error5sum')))
# join the telemetry data with errors
error_count = (telemetry.join(error_ind,
((telemetry['machineID'] == error_ind['machineID'])
& (telemetry['datetime'] == error_ind['datetime'])), "left")
.drop('volt', 'rotate', 'pressure', 'vibration')
.drop(error_ind.machineID).drop(error_ind.datetime)
.fillna(0))
error_features = ['error1sum','error2sum', 'error3sum', 'error4sum', 'error5sum']
wSpec = Window.partitionBy('machineID').orderBy('datetime').rowsBetween(1-24, 0)
for col_name in error_features:
# We're only interested in the erros in the previous 24 hours.
error_count = error_count.withColumn(col_name+'_rollingmean_24',
F.avg(col(col_name)).over(wSpec))
error_feat = (error_count.withColumn("dt_truncated", dt_truncated)
.drop('error1sum', 'error2sum', 'error3sum', 'error4sum', 'error5sum').fillna(0)
.groupBy("machineID","dt_truncated")
.agg(F.mean('error1sum_rollingmean_24').alias('error1sum_rollingmean_24'),
F.mean('error2sum_rollingmean_24').alias('error2sum_rollingmean_24'),
F.mean('error3sum_rollingmean_24').alias('error3sum_rollingmean_24'),
F.mean('error4sum_rollingmean_24').alias('error4sum_rollingmean_24'),
F.mean('error5sum_rollingmean_24').alias('error5sum_rollingmean_24')))
print(error_feat.count())
display(error_feat.limit(5))
```
## Days since last replacement from maintenance
A crucial data set in this example is the use of maintenance records, which contain the information regarding component replacement. Possible features from this data set can be the number of replacements of each component over time or to calculate how long it has been since a component has been replaced. Replacement time is expected to correlate better with component failures since the longer a component is used, the more degradation would be expected.
As a side note, creating lagging features from maintenance data is not straight forward. This type of ad-hoc feature engineering is very common in predictive maintenance as domain knowledge plays a crucial role in understanding the predictors of a failure problem. In the following code blocks, the days since last component replacement are calculated for each component from the maintenance data. We start by counting the component replacements for the set of machines.
```
# create a column for each component replacement
maint_replace = (maint.groupBy("machineID","datetime","comp").pivot('comp')
.agg(F.count('machineID').alias('dummy')).fillna(0)
.groupBy("machineID","datetime")
.agg(F.sum('comp1').alias('comp1sum'),
F.sum('comp2').alias('comp2sum'),
F.sum('comp3').alias('comp3sum'),
F.sum('comp4').alias('comp4sum')))
maint_replace = maint_replace.withColumnRenamed('datetime','datetime_maint')
print(maint_replace.count())
maint_replace.limit(5).toPandas()
```
Replacement features are then created by tracking the number of days between each component replacement. We'll repeat these calculations for each of the four components and join them together into a maintenance feature table.
First component number 1 (`comp1`):
```
# We want to align the component information on telemetry features timestamps.
telemetry_times = (telemetry_feat.select(telemetry_feat.machineID, telemetry_feat.dt_truncated)
.withColumnRenamed('dt_truncated','datetime_tel'))
# Grab component 1 records
maint_comp1 = (maint_replace.where(col("comp1sum") == '1').withColumnRenamed('datetime','datetime_maint')
.drop('comp2sum', 'comp3sum', 'comp4sum'))
# Within each machine, get the last replacement date for each timepoint
maint_tel_comp1 = (telemetry_times.join(maint_comp1,
((telemetry_times ['machineID']== maint_comp1['machineID'])
& (telemetry_times ['datetime_tel'] > maint_comp1['datetime_maint'])
& ( maint_comp1['comp1sum'] == '1')))
.drop(maint_comp1.machineID))
# Calculate the number of days between replacements
comp1 = (maint_tel_comp1.withColumn("sincelastcomp1",
datediff(maint_tel_comp1.datetime_tel, maint_tel_comp1.datetime_maint))
.drop(maint_tel_comp1.datetime_maint).drop(maint_tel_comp1.comp1sum))
print(comp1.count())
comp1.filter(comp1.machineID == '625').orderBy(comp1.datetime_tel).limit(5).toPandas()
```
Then component 2 (`comp2`):
```
# Grab component 2 records
maint_comp2 = (maint_replace.where(col("comp2sum") == '1').withColumnRenamed('datetime','datetime_maint')
.drop('comp1sum', 'comp3sum', 'comp4sum'))
# Within each machine, get the last replacement date for each timepoint
maint_tel_comp2 = (telemetry_times.join(maint_comp2,
((telemetry_times ['machineID']== maint_comp2['machineID'])
& (telemetry_times ['datetime_tel'] > maint_comp2['datetime_maint'])
& ( maint_comp2['comp2sum'] == '1')))
.drop(maint_comp2.machineID))
# Calculate the number of days between replacements
comp2 = (maint_tel_comp2.withColumn("sincelastcomp2",
datediff(maint_tel_comp2.datetime_tel, maint_tel_comp2.datetime_maint))
.drop(maint_tel_comp2.datetime_maint).drop(maint_tel_comp2.comp2sum))
print(comp2.count())
comp2.filter(comp2.machineID == '625').orderBy(comp2.datetime_tel).limit(5).toPandas()
```
Then component 3 (`comp3`):
```
# Grab component 3 records
maint_comp3 = (maint_replace.where(col("comp3sum") == '1').withColumnRenamed('datetime','datetime_maint')
.drop('comp1sum', 'comp2sum', 'comp4sum'))
# Within each machine, get the last replacement date for each timepoint
maint_tel_comp3 = (telemetry_times.join(maint_comp3, ((telemetry_times ['machineID']==maint_comp3['machineID'])
& (telemetry_times ['datetime_tel'] > maint_comp3['datetime_maint'])
& ( maint_comp3['comp3sum'] == '1')))
.drop(maint_comp3.machineID))
# Calculate the number of days between replacements
comp3 = (maint_tel_comp3.withColumn("sincelastcomp3",
datediff(maint_tel_comp3.datetime_tel, maint_tel_comp3.datetime_maint))
.drop(maint_tel_comp3.datetime_maint).drop(maint_tel_comp3.comp3sum))
print(comp3.count())
comp3.filter(comp3.machineID == '625').orderBy(comp3.datetime_tel).limit(5).toPandas()
```
and component 4 (`comp4`):
```
# Grab component 4 records
maint_comp4 = (maint_replace.where(col("comp4sum") == '1').withColumnRenamed('datetime','datetime_maint')
.drop('comp1sum', 'comp2sum', 'comp3sum'))
# Within each machine, get the last replacement date for each timepoint
maint_tel_comp4 = telemetry_times.join(maint_comp4, ((telemetry_times['machineID']==maint_comp4['machineID'])
& (telemetry_times['datetime_tel'] > maint_comp4['datetime_maint'])
& (maint_comp4['comp4sum'] == '1'))).drop(maint_comp4.machineID)
# Calculate the number of days between replacements
comp4 = (maint_tel_comp4.withColumn("sincelastcomp4",
datediff(maint_tel_comp4.datetime_tel, maint_tel_comp4.datetime_maint))
.drop(maint_tel_comp4.datetime_maint).drop(maint_tel_comp4.comp4sum))
print(comp4.count())
comp4.filter(comp4.machineID == '625').orderBy(comp4.datetime_tel).limit(5).toPandas()
```
Now, we join the four component replacement tables together. Once joined, align the data by tumbling the average across 12 hour observation windows.
```
# Join component 3 and 4
comp3_4 = (comp3.join(comp4, ((comp3['machineID'] == comp4['machineID'])
& (comp3['datetime_tel'] == comp4['datetime_tel'])), "left")
.drop(comp4.machineID).drop(comp4.datetime_tel))
# Join component 2 to 3 and 4
comp2_3_4 = (comp2.join(comp3_4, ((comp2['machineID'] == comp3_4['machineID'])
& (comp2['datetime_tel'] == comp3_4['datetime_tel'])), "left")
.drop(comp3_4.machineID).drop(comp3_4.datetime_tel))
# Join component 1 to 2, 3 and 4
comps_feat = (comp1.join(comp2_3_4, ((comp1['machineID'] == comp2_3_4['machineID'])
& (comp1['datetime_tel'] == comp2_3_4['datetime_tel'])), "left")
.drop(comp2_3_4.machineID).drop(comp2_3_4.datetime_tel)
.groupBy("machineID", "datetime_tel")
.agg(F.max('sincelastcomp1').alias('sincelastcomp1'),
F.max('sincelastcomp2').alias('sincelastcomp2'),
F.max('sincelastcomp3').alias('sincelastcomp3'),
F.max('sincelastcomp4').alias('sincelastcomp4'))
.fillna(0))
# Choose the time_val hour timestamps to align the data
dt_truncated = ((round(unix_timestamp(col("datetime_tel")) / time_val) * time_val).cast("timestamp"))
# Collect data
maint_feat = (comps_feat.withColumn("dt_truncated", dt_truncated)
.groupBy("machineID","dt_truncated")
.agg(F.mean('sincelastcomp1').alias('comp1sum'),
F.mean('sincelastcomp2').alias('comp2sum'),
F.mean('sincelastcomp3').alias('comp3sum'),
F.mean('sincelastcomp4').alias('comp4sum')))
print(maint_feat.count())
maint_feat.limit(5).toPandas()
```
## Machine features
The machine features capture specifics of the individuals. These can be used without further modification since it include descriptive information about the type of each machine and its age (number of years in service). If the age information had been recorded as a "first use date" for each machine, a transformation would have been necessary to turn those into a numeric values indicating the years in service.
We do need to create a set of dummy features, a set of boolean variables, to indicate the model of the machine. This can either be done manually, or using a _one-hot encoding_ step. We use the one-hot encoding for demonstration purposes.
```
# one hot encoding of the variable model, basically creates a set of dummy boolean variables
catVarNames = ['model']
sIndexers = [StringIndexer(inputCol=x, outputCol=x + '_indexed') for x in catVarNames]
machines_cat = Pipeline(stages=sIndexers).fit(machines).transform(machines)
# one-hot encode
ohEncoders = [OneHotEncoder(inputCol=x + '_indexed', outputCol=x + '_encoded')
for x in catVarNames]
ohPipelineModel = Pipeline(stages=ohEncoders).fit(machines_cat)
machines_cat = ohPipelineModel.transform(machines_cat)
drop_list = [col_n for col_n in machines_cat.columns if 'indexed' in col_n]
machines_feat = machines_cat.select([column for column in machines_cat.columns if column not in drop_list])
print(machines_feat.count())
display(machines_feat.limit(5))
```
## Merging feature data
Next, we merge the telemetry, maintenance, machine and error feature data sets into a large feature data set. Since most of the data has already been aligned on the 12 hour observation period, we can merge with a simple join strategy.
```
# join error features with component maintenance features
error_maint = (error_feat.join(maint_feat,
((error_feat['machineID'] == maint_feat['machineID'])
& (error_feat['dt_truncated'] == maint_feat['dt_truncated'])), "left")
.drop(maint_feat.machineID).drop(maint_feat.dt_truncated))
# now join that with machines features
error_maint_feat = (error_maint.join(machines_feat,
((error_maint['machineID'] == machines_feat['machineID'])), "left")
.drop(machines_feat.machineID))
# Clean up some unecessary columns
error_maint_feat = error_maint_feat.select([c for c in error_maint_feat.columns if c not in
{'error1sum', 'error2sum', 'error3sum', 'error4sum', 'error5sum'}])
# join telemetry with error/maint/machine features to create final feature matrix
final_feat = (telemetry_feat.join(error_maint_feat,
((telemetry_feat['machineID'] == error_maint_feat['machineID'])
& (telemetry_feat['dt_truncated'] == error_maint_feat['dt_truncated'])), "left")
.drop(error_maint_feat.machineID).drop(error_maint_feat.dt_truncated))
print(final_feat.count())
final_feat.filter(final_feat.machineID == '625').orderBy(final_feat.dt_truncated).limit(5).toPandas()
```
# Label construction
Predictive maintenance is supervised learning. To train a model to predict failures requires examples of failures, and the time series of observations leading up to those failures. Additionally, the model needs examples of periods of healthy operation in order to discern the difference between the two states. The classification between these states is typically a boolean label (healthy vs failed).
Once we have the healthy vs. failure states, the predictive maintenance approach is only useful if the method will give some advanced warning of an impending failure. To accomplish this _prior warning_ criteria, we slightly modify the label definition from a _failure event_ which occurs at a specific moment in time, to a longer window of _failure event occurs within this window_. The window length is defined by the business criteria. Is knowing a failure will occur within 12 hours, enough time to prevent the failure from happening? Is 24 hours, or 2 weeks? The ability of the model to accurately predict an impending failure is dependent sizing this window. If the failure signal is short, longer windows will not help, and can actually degrade, the potential performance.
To acheive the redefinition of failure to _about to fail_, we over label failure events, labeling all observations within the failure warning window as failed. The prediction problem then becomes estimating the probability of failure within this window.
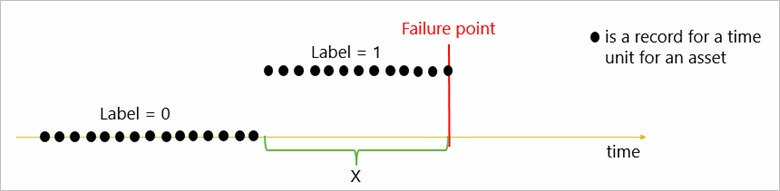
For this example scenerio, we estimate the probability that a machine will fail in the near future due to a failure of a certain component. More specifically, the goal is to compute the probability that a machine will fail in the next 7 days due to a component failure (component 1, 2, 3, or 4).
Below, a categorical failure feature is created to serve as the label. All records within a 24 hour window before a failure of component 1 have failure="comp1", and so on for components 2, 3, and 4; all records not within 7 days of a component failure have failure="none".
The first step is to align the failure data to the feature observation time points (every 12 hours).
```
dt_truncated = ((round(unix_timestamp(col("datetime")) / time_val) * time_val).cast("timestamp"))
fail_diff = (failures.withColumn("dt_truncated", dt_truncated)
.drop(failures.datetime))
print(fail_diff.count())
display(fail_diff.limit(5))
```
Next, we convert the labels from text to numeric values. In the end, this will transform the problem from boolean of 'healthy'/'impending failure' to a multiclass 'healthy'/'component `n` impending failure'.
```
# map the failure data to final feature matrix
labeled_features = (final_feat.join(fail_diff,
((final_feat['machineID'] == fail_diff['machineID'])
& (final_feat['dt_truncated'] == fail_diff['dt_truncated'])), "left")
.drop(fail_diff.machineID).drop(fail_diff.dt_truncated)
.withColumn('failure', F.when(col('failure') == "comp1", 1.0).otherwise(col('failure')))
.withColumn('failure', F.when(col('failure') == "comp2", 2.0).otherwise(col('failure')))
.withColumn('failure', F.when(col('failure') == "comp3", 3.0).otherwise(col('failure')))
.withColumn('failure', F.when(col('failure') == "comp4", 4.0).otherwise(col('failure'))))
labeled_features = (labeled_features.withColumn("failure",
labeled_features.failure.cast(DoubleType()))
.fillna(0))
print(labeled_features.count())
labeled_features.limit(5).toPandas()
```
To verify we have assigned the component failure records correctly, we count the failure classes within the feature data.
```
# To get the frequency of each component failure
lf_count = labeled_features.groupBy('failure').count().collect()
display(lf_count)
```
To now, we have labels as _failure events_. To convert to _impending failure_, we over label over the previous 7 days as _failed_.
```
# lag values to manually backfill label (bfill =7)
my_window = Window.partitionBy('machineID').orderBy(labeled_features.dt_truncated.desc())
# Create the previous 7 days
labeled_features = (labeled_features.withColumn("prev_value1",
F.lag(labeled_features.failure).
over(my_window)).fillna(0))
labeled_features = (labeled_features.withColumn("prev_value2",
F.lag(labeled_features.prev_value1).
over(my_window)).fillna(0))
labeled_features = (labeled_features.withColumn("prev_value3",
F.lag(labeled_features.prev_value2).
over(my_window)).fillna(0))
labeled_features = (labeled_features.withColumn("prev_value4",
F.lag(labeled_features.prev_value3).
over(my_window)).fillna(0))
labeled_features = (labeled_features.withColumn("prev_value5",
F.lag(labeled_features.prev_value4).
over(my_window)).fillna(0))
labeled_features = (labeled_features.withColumn("prev_value6",
F.lag(labeled_features.prev_value5).
over(my_window)).fillna(0))
labeled_features = (labeled_features.withColumn("prev_value7",
F.lag(labeled_features.prev_value6).
over(my_window)).fillna(0))
# Create a label features
labeled_features = (labeled_features.withColumn('label', labeled_features.failure +
labeled_features.prev_value1 +
labeled_features.prev_value2 +
labeled_features.prev_value3 +
labeled_features.prev_value4 +
labeled_features.prev_value5 +
labeled_features.prev_value6 +
labeled_features.prev_value7))
# Restrict the label to be on the range of 0:4, and remove extra columns
labeled_features = (labeled_features.withColumn('label_e', F.when(col('label') > 4, 4.0)
.otherwise(col('label')))
.drop(labeled_features.prev_value1).drop(labeled_features.prev_value2)
.drop(labeled_features.prev_value3).drop(labeled_features.prev_value4)
.drop(labeled_features.prev_value5).drop(labeled_features.prev_value6)
.drop(labeled_features.prev_value7).drop(labeled_features.label))
print(labeled_features.count())
labeled_features.limit(5).toPandas()
```
To verify the label construction, we plot the label distribuiton of a sample of four machines over the data set life time. We expect the labels to cluster for each component, since there are 7 day windows of "fail". We have omitted the healthy labels, as they are uninformative.
We see each of the four machines have multiple failures over the course of the dataset. Each labeled failure includes the date of failure and the previous seven days, all are marked with the number indicating the component that failed.
The goal of the model will be to predict when a failure will occur and which component will fail simultaneously. This will be a multiclass classification problem, though we could pivot the data to individually predict binary failure of a component instead of a machine.
```
plt_data = (labeled_features.filter(labeled_features.label_e > 0)
.where(col("machineID").isin({"65", "558", "222", "965"}))
.select(labeled_features.machineID, labeled_features.label_e)).toPandas()
fig, ax = plt.subplots(figsize = (14,7))
sns.barplot(plt_data['machineID'], plt_data['label_e'], hue = plt_data['label_e'], alpha=0.8).set_title('Label distribution per machine')
ax.set_ylabel(" Label Count")
ax.set_xlabel("Machine ID")
display(ax.figure)
```
## Write the feature data to storage
Write the final labeled feature data as parquet file to cluster storage.
```
# Write labeled feature data to storage
labeled_features.write.mode('overwrite').parquet(os.path.join(target_dir, parquet_files_names['features']))
toc = time.time()
print("Full run took %.2f minutes" % ((toc - tic)/60))
```
# Conclusion
The next step is to build and compare machine learning models using the feature data set we have just created. The `3_model_building.ipynb` notebook works through building a Decision Tree Classifier and a Random Forest Classifier using this data set.
| github_jupyter |
# A Hydrofunctions Tutorial
This guide will step you through the basics of using hydrofunctions. Read more in our [User's Guide](https://hydrofunctions.readthedocs.io), or visit us on [GitHub](https://github.com/mroberge/hydrofunctions)!
## Installation
The first step before using hydrofunctions is to get it installed on your system. For scientific computing, we highly recommend using the free, open-source [Anaconda](https://www.anaconda.com/download/) distribution to load and manage all of your Python tools and packages. Once you have downloaded and installed Anaconda, or if you already have Python set up on your computer, your next step is to use the pip tool from your operating system's command line to download hydrofunctions.
In Linux:
`$ pip install hydrofunctions`
In Windows:
`C:\MyPythonWorkspace\> pip install hydrofunctions`
If you have any difficulties, visit our [Installation](https://hydrofunctions.readthedocs.io/en/master/installation.html) page in the User's Guide.
## Getting started in Python
From here on out, we will assume that you have installed hydrofunctions and you are working at a Python command prompt,
perhaps in ipython or in a Jupyter notebook.
```
# The first step is to import hydrofunctions so that we can use it here.
import hydrofunctions as hf
# This second line allows us to automatically plot diagrams in this notebook.
%matplotlib inline
```
### Get data for a USGS streamflow gage
The USGS runs an amazing web service called the [National Water Information System](https://waterdata.usgs.gov/nwis). Our
first task is to download daily mean discharge data for a stream called Herring Run. Set the start date and the end date for
our download, and use the site number for Herring Run ('01585200') to specify which stream gage we want to collect data from. Once we request the data, it will be saved to a file. If the file is already present, we'll just use that instead of requesting it from the NWIS.
You can visit the [NWIS](https://waterdata.usgs.gov/nwis) website or use [hydrocloud.org](https://mroberge.github.io/HydroCloud/) to find the site number for a stream gage near you.
```
start = '2017-06-01'
end = '2017-07-14'
herring = hf.NWIS('01585200', 'dv', start, end, file='herring_july.parquet')
herring # This last command will print out a description of what we have.
```
## Viewing our data
There are several ways to view our data. Try herring.json() or better still, use a [Pandas](https://pandas.pydata.org/) dataframe:
```
herring.df()
```
Pandas' dataframes give you access to hundreds of useful methods, such as .describe() and .plot():
```
herring.df().describe()
herring.df().plot()
```
## Multiple sites, other parameters
It's possible to load data from several different sites at the same time, and you aren't limited to just stream discharge.
Requests can use lists of sites:
```
sites = ['380616075380701','394008077005601']
```
The NWIS can deliver data as daily mean values ('dv') or as instantaneous values ('iv')
that can get collected as often as every five minutes!
```
service = 'iv'
```
Depending on the site, the USGS collects groundwater levels ('72019'), stage ('00065'), precipitation, and more!
```
pcode = '72019'
```
Now we'll create a new dataset called 'groundwater' using the values we set up above.
Since one of the parameters gets collected every 30 minutes, and the other gets collected every 15 minutes, Hydrofunctions will interpolate values for every 15 minutes for every parameter we've requested. These interpolated values will be marked with a special `hf.interpolate` flag in the qualifiers column.
```
groundwater = hf.NWIS(sites, service, '2018-01-01', '2018-01-31', parameterCd=pcode, file='groundwater.parquet')
groundwater
```
Calculate the mean for every data column:
```
groundwater.df().mean()
```
View the data in a specially styled graph!
```
groundwater.df().plot(marker='o', mfc='white', ms=4, mec='black', color='black')
```
## Learning more
hydrofunctions comes with a variety of built-in help functions that you can access from the command line, in addition to our online [User's Guide](https://hydrofunctions.readthedocs.io).
Jupyter Notebooks provide additional helpful shortcuts, such as code completion. This will list all of the available
methods for an object just by hitting <TAB> like this: `herring.<TAB>` this is equivalent to using dir(herring) to list
all of the methods available to you.
Typing help() or dir() for different objects allows you to access additional information.
`help(hf.NWIS)` is equivalent to just using a question mark like this: `?hf.NWIS`
```
help(hf.NWIS)
```
## Advanced techniques
### Download data for a large number of sites
```
sites = ['07227500', '07228000', '07235000', '07295500', '07297910', '07298500', '07299540',
'07299670', '07299890', '07300000', '07301300', '07301410', '07308200', '07308500', '07311600',
'07311630', '07311700', '07311782', '07311783', '07311800', '07311900', '07312100', '07312200',
'07312500', '07312700', '07314500', '07314900', '07315200', '07315500', '07342465', '07342480',
'07342500', '07343000', '07343200', '07343500', '07344210', '07344500', '07346000']
mult = hf.NWIS(sites, "dv", "2018-01-01", "2018-01-31", file='mult.parquet')
print('No. sites: {}'.format(len(sites)))
```
This will calculate the mean value for each site.
```
mult.df().mean()
```
Plot just the discharge data for one site in the list:
```
mult.df('07228000', 'discharge').plot()
```
#### List some of the data available to you in a dataframe.
```
mult
```
#### Create a table of discharge data
`.head()` only show the first five
`.tail()` only show the last five
```
mult.df('discharge').head()
```
### Download all streamflow data for the state of Virginia
```
# Use this carefully! You can easily request more data than you will know what to do with.
start = "2017-01-01"
end = "2017-12-31"
param = '00060'
virginia = hf.NWIS(None, "dv", start, end, parameterCd=param, stateCd='va', file='virginia.parquet')
# Calculate the mean for each site.
virginia.df('discharge').mean()
```
#### Plot all streamflow data for the state of Virginia
```
# There are so many sites that we can't read them all!
virginia.df('q').plot(legend=None)
```
### Download all streamflow data for Fairfax and Prince William counties in the state of Virginia
```
start = "2017-01-01"
end = "2017-12-31"
county = hf.NWIS(None, "dv", start, end, parameterCd='00060', countyCd=['51059', '51061'], file='PG.parquet')
county.df('data').head()
```
#### Plot all streamflow data for Fairfax and Prince William counties in the state of Virginia
```
county.df('data').plot()
```
## Thanks for using hydrofunctions!
We would love to hear your comments and [suggestions](https://github.com/mroberge/hydrofunctions/issues)!
| github_jupyter |
```
import open3d as o3d
import numpy as np
import re
import os
import sys
# monkey patches visualization and provides helpers to load geometries
sys.path.append('..')
import open3d_tutorial as o3dtut
# change to True if you want to interact with the visualization windows
o3dtut.interactive = not "CI" in os.environ
# if running on Travis CI, the number of iterations is reduced
is_ci = "CI" in os.environ
```
# Color Map Optimization
Consider color mapping to the geometry reconstructed from depth cameras. As color and depth frames are not perfectly aligned, the texture mapping using color images is subject to results in blurred color map. Open3D provides color map optimization method proposed by [\[Zhou2014\]](../reference.html#zhou2014). The following script shows an example of color map optimization.
## Input
This code below reads color and depth image pairs and makes `rgbd_image`. Note that `convert_rgb_to_intensity` flag is `False`. This is to preserve 8-bit color channels instead of using single channel float type image.
It is always good practice to visualize the RGBD image before applying it to the color map optimization. The `debug_mode` switch can be set to `True` to visualize the RGBD image.
```
def sorted_alphanum(file_list_ordered):
convert = lambda text: int(text) if text.isdigit() else text
alphanum_key = lambda key: [convert(c) for c in re.split('([0-9]+)', key)]
return sorted(file_list_ordered, key=alphanum_key)
def get_file_list(path, extension=None):
if extension is None:
file_list = [
path + f for f in os.listdir(path) if os.path.isfile(join(path, f))
]
else:
file_list = [
path + f
for f in os.listdir(path)
if os.path.isfile(os.path.join(path, f)) and
os.path.splitext(f)[1] == extension
]
file_list = sorted_alphanum(file_list)
return file_list
path = o3dtut.download_fountain_dataset()
debug_mode = False
rgbd_images = []
depth_image_path = get_file_list(os.path.join(path, "depth/"), extension=".png")
color_image_path = get_file_list(os.path.join(path, "image/"), extension=".jpg")
assert (len(depth_image_path) == len(color_image_path))
for i in range(len(depth_image_path)):
depth = o3d.io.read_image(os.path.join(depth_image_path[i]))
color = o3d.io.read_image(os.path.join(color_image_path[i]))
rgbd_image = o3d.geometry.RGBDImage.create_from_color_and_depth(
color, depth, convert_rgb_to_intensity=False)
if debug_mode:
pcd = o3d.geometry.PointCloud.create_from_rgbd_image(
rgbd_image,
o3d.camera.PinholeCameraIntrinsic(
o3d.camera.PinholeCameraIntrinsicParameters.PrimeSenseDefault))
o3d.visualization.draw_geometries([pcd])
rgbd_images.append(rgbd_image)
```
The code below reads a camera trajectory and a mesh.
```
camera = o3d.io.read_pinhole_camera_trajectory(
os.path.join(path, "scene/key.log"))
mesh = o3d.io.read_triangle_mesh(os.path.join(path, "scene", "integrated.ply"))
```
To visualize how the camera poses are not good for color mapping, this code intentionally sets the iteration number to 0, which means no optimization. `color_map_optimization` paints a mesh using corresponding RGBD images and camera poses. Without optimization, the texture map is blurred.
```
# Before full optimization, let's just visualize texture map
# with given geometry, RGBD images, and camera poses.
option = o3d.pipelines.color_map.ColorMapOptimizationOption()
option.maximum_iteration = 0
with o3d.utility.VerbosityContextManager(
o3d.utility.VerbosityLevel.Debug) as cm:
o3d.pipelines.color_map.color_map_optimization(mesh, rgbd_images, camera,
option)
o3d.visualization.draw_geometries([mesh],
zoom=0.5399,
front=[0.0665, -0.1107, -0.9916],
lookat=[0.7353, 0.6537, 1.0521],
up=[0.0136, -0.9936, 0.1118])
```
## Rigid Optimization
The next step is to optimize camera poses to get a sharp color map.
The code below sets `maximum_iteration = 300` for actual iterations.
```
# Optimize texture and save the mesh as texture_mapped.ply
# This is implementation of following paper
# Q.-Y. Zhou and V. Koltun,
# Color Map Optimization for 3D Reconstruction with Consumer Depth Cameras,
# SIGGRAPH 2014
option.maximum_iteration = 100 if is_ci else 300
option.non_rigid_camera_coordinate = False
with o3d.utility.VerbosityContextManager(
o3d.utility.VerbosityLevel.Debug) as cm:
o3d.pipelines.color_map.color_map_optimization(mesh, rgbd_images, camera,
option)
o3d.visualization.draw_geometries([mesh],
zoom=0.5399,
front=[0.0665, -0.1107, -0.9916],
lookat=[0.7353, 0.6537, 1.0521],
up=[0.0136, -0.9936, 0.1118])
```
The residual error implies inconsistency of image intensities. Lower residual leads to a better color map quality. By default, `ColorMapOptimizationOption` enables rigid optimization. It optimizes 6-dimentional pose of every cameras.
## Non-rigid Optimization
For better alignment quality, there is an option for non-rigid optimization. To enable this option, simply set `option.non_rigid_camera_coordinate` to `True` before calling `color_map_optimization`. Besides 6-dimentional camera poses, non-rigid optimization even considers local image warping represented by anchor points. This adds even more flexibility and leads to an even higher quality color mapping. The residual error is smaller than the case of rigid optimization.
```
option.maximum_iteration = 100 if is_ci else 300
option.non_rigid_camera_coordinate = True
with o3d.utility.VerbosityContextManager(
o3d.utility.VerbosityLevel.Debug) as cm:
o3d.pipelines.color_map.color_map_optimization(mesh, rgbd_images, camera,
option)
o3d.visualization.draw_geometries([mesh],
zoom=0.5399,
front=[0.0665, -0.1107, -0.9916],
lookat=[0.7353, 0.6537, 1.0521],
up=[0.0136, -0.9936, 0.1118])
```
| github_jupyter |
<a href="https://colab.research.google.com/github/NeuromatchAcademy/course-content/blob/W2D1-postcourse-bugfix/tutorials/W2D1_BayesianStatistics/student/W2D1_Tutorial4.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# NMA 2020 W2D1 -- (Bonus) Tutorial 4: Bayesian Decision Theory & Cost functions
__Content creators:__ Vincent Valton, Konrad Kording, with help from Matthew Krause
__Content reviewers:__ Matthew Krause, Jesse Livezey, Karolina Stosio, Saeed Salehi
# Tutorial Objectives
*This tutorial is optional! Please do not feel pressured to finish it!*
In the previous tutorials, we investigated the posterior, which describes beliefs based on a combination of current evidence and prior experience. This tutorial focuses on Bayesian Decision Theory, which combines the posterior with **cost functions** that allow us to quantify the potential impact of making a decision or choosing an action based on that posterior. Cost functions are therefore critical for turning probabilities into actions!
In Tutorial 3, we used the mean of the posterior $p(x | \tilde x)$ as a proxy for the response $\hat x$ for the participants. What prompted us to use the mean of the posterior as a **decision rule**? In this tutorial we will see how different common decision rules such as the choosing the mean, median or mode of the posterior distribution correspond to minimizing different cost functions.
In this tutorial, you will
1. Implement three commonly-used cost functions: mean-squared error, absolute error, and zero-one loss
2. Discover the concept of expected loss, and
3. Choose optimal locations on the posterior that minimize these cost functions. You will verify that it these locations can be found analytically as well as empirically.
```
#@title Video 1: Introduction
from IPython.display import YouTubeVideo
video = YouTubeVideo(id='z2DF4H_sa-k', width=854, height=480, fs=1)
print("Video available at https://youtube.com/watch?v=" + video.id)
video
```
---
Please execute the cell below to initialize the notebook environment
---
### Setup
```
# Imports
import numpy as np
import matplotlib.pyplot as plt
#@title Figure Settings
import ipywidgets as widgets
plt.style.use("https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/nma.mplstyle")
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
# @title Helper Functions
def my_gaussian(x_points, mu, sigma):
"""Returns un-normalized Gaussian estimated at points `x_points`
DO NOT EDIT THIS FUNCTION !!!
Args :
x_points (numpy array of floats) - points at which the gaussian is evaluated
mu (scalar) - mean of the Gaussian
sigma (scalar) - std of the gaussian
Returns:
(numpy array of floats): un-normalized Gaussian (i.e. without constant) evaluated at `x`
"""
return np.exp(-(x_points-mu)**2/(2*sigma**2))
def visualize_loss_functions(mse=None, abse=None, zero_one=None):
"""Visualize loss functions
Args:
- mse (func) that returns mean-squared error
- abse: (func) that returns absolute_error
- zero_one: (func) that returns zero-one loss
All functions should be of the form f(x, x_hats). See Exercise #1.
Returns:
None
"""
x = np.arange(-3, 3.25, 0.25)
fig, ax = plt.subplots(1)
if mse is not None:
ax.plot(x, mse(0, x), linewidth=2, label="Mean Squared Error")
if abse is not None:
ax.plot(x, abse(0, x), linewidth=2, label="Absolute Error")
if zero_one_loss is not None:
ax.plot(x, zero_one_loss(0, x), linewidth=2, label="Zero-One Loss")
ax.set_ylabel('Cost')
ax.set_xlabel('Predicted Value ($\hat{x}$)')
ax.set_title("Loss when the true value $x$=0")
ax.legend()
plt.show()
def moments_myfunc(x_points, function):
"""Returns the mean, median and mode of an arbitrary function
DO NOT EDIT THIS FUNCTION !!!
Args :
x_points (numpy array of floats) - x-axis values
function (numpy array of floats) - y-axis values of the function evaluated at `x_points`
Returns:
(tuple of 3 scalars): mean, median, mode
"""
# Calc mode of an arbitrary function
mode = x_points[np.argmax(function)]
# Calc mean of an arbitrary function
mean = np.sum(x_points * function)
# Calc median of an arbitrary function
cdf_function = np.zeros_like(x_points)
accumulator = 0
for i in np.arange(x.shape[0]):
accumulator = accumulator + posterior[i]
cdf_function[i] = accumulator
idx = np.argmin(np.abs(cdf_function - 0.5))
median = x_points[idx]
return mean, median, mode
def loss_plot(x, loss, min_loss, loss_label, show=False, ax=None):
if not ax:
fig, ax = plt.subplots()
ax.plot(x, loss, '-C1', linewidth=2, label=loss_label)
ax.axvline(min_loss, ls='dashed', color='C1', label='Minimum')
ax.set_ylabel('Expected Loss')
ax.set_xlabel('Orientation (Degrees)')
ax.legend()
if show:
plt.show()
def loss_plot_subfigures(x,
MSEloss, min_MSEloss, loss_MSElabel,
ABSEloss, min_ABSEloss, loss_ABSElabel,
ZeroOneloss, min_01loss, loss_01label):
fig_w, fig_h = plt.rcParams.get('figure.figsize')
fig, ax = plt.subplots(nrows=2, ncols=3, figsize=(fig_w*2, fig_h*2), sharex=True)
ax[0, 0].plot(x, MSEloss, '-C1', linewidth=2, label=loss_MSElabel)
ax[0, 0].axvline(min_MSEloss, ls='dashed', color='C1', label='Minimum')
ax[0, 0].set_ylabel('Expected Loss')
ax[0, 0].set_xlabel('Orientation (Degrees)')
ax[0, 0].set_title("Mean Squared Error")
ax[0, 0].legend()
pmoments_plot(x, posterior, ax=ax[1,0])
ax[0, 1].plot(x, ABSEloss, '-C0', linewidth=2, label=loss_ABSElabel)
ax[0, 1].axvline(min_ABSEloss, ls='dashdot', color='C0', label='Minimum')
ax[0, 1].set_ylabel('Expected Loss')
ax[0, 1].set_xlabel('Orientation (Degrees)')
ax[0, 1].set_title("Absolute Error")
ax[0, 1].legend()
pmoments_plot(x, posterior, ax=ax[1,1])
ax[0, 2].plot(x, ZeroOneloss, '-C2', linewidth=2, label=loss_01label)
ax[0, 2].axvline(min_01loss, ls='dotted', color='C2', label='Minimum')
ax[0, 2].set_ylabel('Expected Loss')
ax[0, 2].set_xlabel('Orientation (Degrees)')
ax[0, 2].set_title("0-1 Loss")
ax[0, 2].legend()
pmoments_plot(x, posterior, ax=ax[1,2])
plt.show()
def pmoments_plot(x, posterior,
prior=None, likelihood=None, show=False, ax=None):
if not ax:
fig, ax = plt.subplots()
if prior:
ax.plot(x, prior, '-C1', linewidth=2, label='Prior')
if likelihood:
ax.plot(x, likelihood, '-C0', linewidth=2, label='Likelihood')
ax.plot(x, posterior, '-C2', linewidth=4, label='Posterior')
mean, median, mode = moments_myfunc(x, posterior)
ax.axvline(mean, ls='dashed', color='C1', label='Mean')
ax.axvline(median, ls='dashdot', color='C0', label='Median')
ax.axvline(mode, ls='dotted', color='C2', label='Mode')
ax.set_ylabel('Probability')
ax.set_xlabel('Orientation (Degrees)')
ax.legend()
if show:
plt.show()
def generate_example_pdfs():
"""Generate example probability distributions as in T2"""
x=np.arange(-5, 5, 0.01)
prior_mean = 0
prior_sigma1 = .5
prior_sigma2 = 3
prior1 = my_gaussian(x, prior_mean, prior_sigma1)
prior2 = my_gaussian(x, prior_mean, prior_sigma2)
alpha = 0.05
prior_combined = (1-alpha) * prior1 + (alpha * prior2)
prior_combined = prior_combined / np.sum(prior_combined)
likelihood_mean = -2.7
likelihood_sigma = 1
likelihood = my_gaussian(x, likelihood_mean, likelihood_sigma)
likelihood = likelihood / np.sum(likelihood)
posterior = prior_combined * likelihood
posterior = posterior / np.sum(posterior)
return x, prior_combined, likelihood, posterior
def plot_posterior_components(x, prior, likelihood, posterior):
with plt.xkcd():
fig = plt.figure()
plt.plot(x, prior, '-C1', linewidth=2, label='Prior')
plt.plot(x, likelihood, '-C0', linewidth=2, label='Likelihood')
plt.plot(x, posterior, '-C2', linewidth=4, label='Posterior')
plt.legend()
plt.title('Sample Output')
plt.show()
```
### The Posterior Distribution
This notebook will use a model similar to the puppet & puppeteer sound experiment developed in Tutorial 2, but with different probabilities for $p_{common}$, $p_{independent}$, $\sigma_{common}$ and $\sigma_{independent}$. Specifically, our model will consist of these components, combined according to Bayes' rule:
$$
\begin{eqnarray}
\textrm{Prior} &=& \begin{cases} \mathcal{N_{common}}(0, 0.5) & 95\% \textrm{ weight}\\
\mathcal{N_{independent}}(0, 3.0) & 5\% \textrm{ weight} \\
\end{cases}\\\\
\textrm{Likelihood} &= &\mathcal{N}(-2.7, 1.0)
\end{eqnarray}
$$
We will use this posterior as an an example through this notebook. Please run the cell below to import and plot the model. You do not need to edit anything. These parameter values were deliberately chosen for illustration purposes: there is nothing intrinsically special about them, but they make several of the exercises easier.
```
x, prior, likelihood, posterior = generate_example_pdfs()
plot_posterior_components(x, prior, likelihood, posterior)
```
# Section 1: The Cost Functions
Next, we will implement the cost functions.
A cost function determines the "cost" (or penalty) of estimating $\hat{x}$ when the true or correct quantity is really $x$ (this is essentially the cost of the error between the true stimulus value: $x$ and our estimate: $\hat x$ -- Note that the error can be defined in different ways):
$$\begin{eqnarray}
\textrm{Mean Squared Error} &=& (x - \hat{x})^2 \\
\textrm{Absolute Error} &=& \big|x - \hat{x}\big| \\
\textrm{Zero-One Loss} &=& \begin{cases}
0,& \text{if } x = \hat{x} \\
1, & \text{otherwise}
\end{cases}
\end{eqnarray}
$$
In the cell below, fill in the body of these cost function. Each function should take one single value for $x$ (the true stimulus value : $x$) and one or more possible value estimates: $\hat{x}$.
Return an array containing the costs associated with predicting $\hat{x}$ when the true value is $x$. Once you have written all three functions, uncomment the final line to visulize your results.
_Hint:_ These functions are easy to write (1 line each!) but be sure *all* three functions return arrays of `np.float` rather than another data type.
## Exercise 1: Implement the cost functions
```
def mse(x, x_hats):
"""Mean-squared error cost function
Args:
x (scalar): One true value of $x$
x_hats (scalar or ndarray): Estimate of x
Returns:
same shape/type as x_hats): MSE costs associated with
predicting x_hats instead of x$
"""
##############################################################################
# Complete the MSE cost function
#
### Comment out the line below to test your function
raise NotImplementedError("You need to complete the MSE cost function!")
##############################################################################
my_mse = ...
return my_mse
def abs_err(x, x_hats):
"""Absolute error cost function
Args:
x (scalar): One true value of $x$
x_hats (scalar or ndarray): Estimate of x
Returns:
(same shape/type as x_hats): absolute error costs associated with
predicting x_hats instead of x$
"""
##############################################################################
# Complete the absolute error cost function
#
### Comment out the line below to test your function
raise NotImplementedError("You need to complete the absolute error function!")
##############################################################################
my_abs_err = ...
return my_abs_err
def zero_one_loss(x, x_hats):
"""Zero-One loss cost function
Args:
x (scalar): One true value of $x$
x_hats (scalar or ndarray): Estimate of x
Returns:
(same shape/type as x_hats) of the 0-1 Loss costs associated with predicting x_hat instead of x
"""
##############################################################################
# Complete the zero-one loss cost function
#
### Comment out the line below to test your function
raise NotImplementedError("You need to complete the 0-1 loss cost function!")
##############################################################################
my_zero_one_loss = ...
return my_zero_one_loss
## When you are done with the functions above, uncomment the line below to
## visualize them
# visualize_loss_functions(mse, abs_err, zero_one_loss)
```
[*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W2D1_BayesianStatistics/solutions/W2D1_Tutorial4_Solution_8da2f3c2.py)
*Example output:*
<img alt='Solution hint' align='left' width=416 height=272 src=https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/tutorials/W2D1_BayesianStatistics/static/W2D1_Tutorial4_Solution_8da2f3c2_0.png>
# Section 2: Expected Loss
```
#@title Video 2: Expected Loss
from IPython.display import YouTubeVideo
video = YouTubeVideo(id='FTBpCfylV_Y', width=854, height=480, fs=1)
print("Video available at https://youtube.com/watch?v=" + video.id)
video
```
A posterior distribution tells us about the confidence or credibility we assign to different choices. A cost function describes the penalty we incur when choosing an incorrect option. These concepts can be combined into an *expected loss* function. Expected loss is defined as:
$$
\begin{eqnarray}
\mathbb{E}[\text{Loss} | \hat{x}] = \int L[\hat{x},x] \odot p(x|\tilde{x}) dx
\end{eqnarray}
$$
where $L[ \hat{x}, x]$ is the loss function, $p(x|\tilde{x})$ is the posterior, and $\odot$ represents the [Hadamard Product](https://en.wikipedia.org/wiki/Hadamard_product_(matrices)) (i.e., elementwise multiplication), and $\mathbb{E}[\text{Loss} | \hat{x}]$ is the expected loss.
In this exercise, we will calculate the expected loss for the: means-squared error, the absolute error, and the zero-one loss over our bimodal posterior $p(x | \tilde x)$.
**Suggestions:**
* We already pre-completed the code (commented-out) to calculate the mean-squared error, absolute error, and zero-one loss between $x$ and an estimate $\hat x$ using the functions you created in exercise 1
* Calculate the expected loss ($\mathbb{E}[MSE Loss]$) using your posterior (imported above as `posterior`) & each of the loss functions described above (MSELoss, ABSELoss, and Zero-oneLoss).
* Find the x position that minimizes the expected loss for each cost function and plot them using the `loss_plot` function provided (commented-out)
## Exercise 2: Finding the expected loss empirically via integration
```
def expected_loss_calculation(x, posterior):
ExpectedLoss_MSE = np.zeros_like(x)
ExpectedLoss_ABSE = np.zeros_like(x)
ExpectedLoss_01 = np.zeros_like(x)
for idx in np.arange(x.shape[0]):
estimate = x[idx]
###################################################################
## Insert code below to find the expected loss under each loss function
##
## remove the raise when the function is complete
raise NotImplementedError("Calculate the expected loss over all x values!")
###################################################################
MSELoss = mse(estimate, x)
ExpectedLoss_MSE[idx] = ...
ABSELoss = abs_err(estimate, x)
ExpectedLoss_ABSE[idx] = ...
ZeroOneLoss = zero_one_loss(estimate, x)
ExpectedLoss_01[idx] = ...
###################################################################
## Now, find the `x` location that minimizes expected loss
##
## remove the raise when the function is complete
raise NotImplementedError("Finish the Expected Loss calculation")
###################################################################
min_MSE = ...
min_ABSE = ...
min_01 = ...
return (ExpectedLoss_MSE, ExpectedLoss_ABSE, ExpectedLoss_01,
min_MSE, min_ABSE, min_01)
## Uncomment the lines below to plot the expected loss as a function of the estimates
#ExpectedLoss_MSE, ExpectedLoss_ABSE, ExpectedLoss_01, min_MSE, min_ABSE, min_01 = expected_loss_calculation(x, posterior)
#loss_plot(x, ExpectedLoss_MSE, min_MSE, f"Mean Squared Error = {min_MSE:.2f}")
#loss_plot(x, ExpectedLoss_ABSE, min_ABSE, f"Absolute Error = {min_ABSE:.2f}")
#loss_plot(x, ExpectedLoss_01, min_01, f"Zero-One Error = {min_01:.2f}")
```
[*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W2D1_BayesianStatistics/solutions/W2D1_Tutorial4_Solution_3a9250ef.py)
*Example output:*
<img alt='Solution hint' align='left' width=424 height=280 src=https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/tutorials/W2D1_BayesianStatistics/static/W2D1_Tutorial4_Solution_3a9250ef_0.png>
<img alt='Solution hint' align='left' width=424 height=280 src=https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/tutorials/W2D1_BayesianStatistics/static/W2D1_Tutorial4_Solution_3a9250ef_1.png>
<img alt='Solution hint' align='left' width=424 height=280 src=https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/tutorials/W2D1_BayesianStatistics/static/W2D1_Tutorial4_Solution_3a9250ef_2.png>
# Section 3: Analytical Solutions
```
#@title Video 3: Analytical Solutions
from IPython.display import YouTubeVideo
video = YouTubeVideo(id='wmDD51N9rs0', width=854, height=480, fs=1)
print("Video available at https://youtube.com/watch?v=" + video.id)
video
```
In the previous exercise, we found the minimum expected loss via brute-force: we searched over all possible values of $x$ and found the one that minimized each of our loss functions. This is feasible for our small toy example, but can quickly become intractable.
Fortunately, the three loss functions examined in this tutorial have are minimized at specific points on the posterior, corresponding to the itss mean, median, and mode. To verify this property, we have replotted the loss functions from Exercise 2 below, with the posterior on the same scale beneath. The mean, median, and mode are marked on the posterior.
Which loss form corresponds to each summary statistics?
```
loss_plot_subfigures(x,
ExpectedLoss_MSE, min_MSE, f"Mean Squared Error = {min_MSE:.2f}",
ExpectedLoss_ABSE, min_ABSE, f"Absolute Error = {min_ABSE:.2f}",
ExpectedLoss_01, min_01, f"Zero-One Error = {min_01:.2f}")
```
[*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W2D1_BayesianStatistics/solutions/W2D1_Tutorial4_Solution_8cdbd46a.py)
# Section 4: Conclusion
```
#@title Video 4: Outro
from IPython.display import YouTubeVideo
video = YouTubeVideo(id='3nTvamDVx2s', width=854, height=480, fs=1)
print("Video available at https://youtube.com/watch?v=" + video.id)
video
```
In this tutorial, we learned about three kinds of cost functions: mean-squared error, absolute error, and zero-one loss. We used expected loss to quantify the results of making a decision, and showed that optimizing under different cost functions led us to choose different locations on the posterior. Finally, we found that these optimal locations can be identified analytically, sparing us from a brute-force search.
Here are some additional questions to ponder:
* Suppose your professor offered to grade your work with a zero-one loss or mean square error.
* When might you choose each?
* Which would be easier to learn from?
* All of the loss functions we considered are symmetrical. Are there situations where an asymmetrical loss function might make sense? How about a negative one?
| github_jupyter |
##### Copyright 2019 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# 基于注意力的神经机器翻译
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.google.cn/tutorials/text/nmt_with_attention">
<img src="https://www.tensorflow.org/images/tf_logo_32px.png" />
在 TensorFlow.org 上查看</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/zh-cn/tutorials/text/nmt_with_attention.ipynb">
<img src="https://www.tensorflow.org/images/colab_logo_32px.png" />
在 Google Colab 运行</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/zh-cn/tutorials/text/nmt_with_attention.ipynb">
<img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />
在 GitHub 上查看源代码</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/docs-l10n/site/zh-cn/tutorials/text/nmt_with_attention.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />下载此 notebook</a>
</td>
</table>
Note: 我们的 TensorFlow 社区翻译了这些文档。因为社区翻译是尽力而为, 所以无法保证它们是最准确的,并且反映了最新的
[官方英文文档](https://www.tensorflow.org/?hl=en)。如果您有改进此翻译的建议, 请提交 pull request 到
[tensorflow/docs](https://github.com/tensorflow/docs) GitHub 仓库。要志愿地撰写或者审核译文,请加入
[docs-zh-cn@tensorflow.org Google Group](https://groups.google.com/a/tensorflow.org/forum/#!forum/docs-zh-cn)。
此笔记本训练一个将西班牙语翻译为英语的序列到序列(sequence to sequence,简写为 seq2seq)模型。此例子难度较高,需要对序列到序列模型的知识有一定了解。
训练完此笔记本中的模型后,你将能够输入一个西班牙语句子,例如 *"¿todavia estan en casa?"*,并返回其英语翻译 *"are you still at home?"*
对于一个简单的例子来说,翻译质量令人满意。但是更有趣的可能是生成的注意力图:它显示在翻译过程中,输入句子的哪些部分受到了模型的注意。
<img src="https://tensorflow.org/images/spanish-english.png" alt="spanish-english attention plot">
请注意:运行这个例子用一个 P100 GPU 需要花大约 10 分钟。
```
import tensorflow as tf
import matplotlib.pyplot as plt
import matplotlib.ticker as ticker
from sklearn.model_selection import train_test_split
import unicodedata
import re
import numpy as np
import os
import io
import time
```
## 下载和准备数据集
我们将使用 http://www.manythings.org/anki/ 提供的一个语言数据集。这个数据集包含如下格式的语言翻译对:
```
May I borrow this book? ¿Puedo tomar prestado este libro?
```
这个数据集中有很多种语言可供选择。我们将使用英语 - 西班牙语数据集。为方便使用,我们在谷歌云上提供了此数据集的一份副本。但是你也可以自己下载副本。下载完数据集后,我们将采取下列步骤准备数据:
1. 给每个句子添加一个 *开始* 和一个 *结束* 标记(token)。
2. 删除特殊字符以清理句子。
3. 创建一个单词索引和一个反向单词索引(即一个从单词映射至 id 的词典和一个从 id 映射至单词的词典)。
4. 将每个句子填充(pad)到最大长度。
```
# 下载文件
path_to_zip = tf.keras.utils.get_file(
'spa-eng.zip', origin='http://storage.googleapis.com/download.tensorflow.org/data/spa-eng.zip',
extract=True)
path_to_file = os.path.dirname(path_to_zip)+"/spa-eng/spa.txt"
# 将 unicode 文件转换为 ascii
def unicode_to_ascii(s):
return ''.join(c for c in unicodedata.normalize('NFD', s)
if unicodedata.category(c) != 'Mn')
def preprocess_sentence(w):
w = unicode_to_ascii(w.lower().strip())
# 在单词与跟在其后的标点符号之间插入一个空格
# 例如: "he is a boy." => "he is a boy ."
# 参考:https://stackoverflow.com/questions/3645931/python-padding-punctuation-with-white-spaces-keeping-punctuation
w = re.sub(r"([?.!,¿])", r" \1 ", w)
w = re.sub(r'[" "]+', " ", w)
# 除了 (a-z, A-Z, ".", "?", "!", ","),将所有字符替换为空格
w = re.sub(r"[^a-zA-Z?.!,¿]+", " ", w)
w = w.rstrip().strip()
# 给句子加上开始和结束标记
# 以便模型知道何时开始和结束预测
w = '<start> ' + w + ' <end>'
return w
en_sentence = u"May I borrow this book?"
sp_sentence = u"¿Puedo tomar prestado este libro?"
print(preprocess_sentence(en_sentence))
print(preprocess_sentence(sp_sentence).encode('utf-8'))
# 1. 去除重音符号
# 2. 清理句子
# 3. 返回这样格式的单词对:[ENGLISH, SPANISH]
def create_dataset(path, num_examples):
lines = io.open(path, encoding='UTF-8').read().strip().split('\n')
word_pairs = [[preprocess_sentence(w) for w in l.split('\t')] for l in lines[:num_examples]]
return zip(*word_pairs)
en, sp = create_dataset(path_to_file, None)
print(en[-1])
print(sp[-1])
def max_length(tensor):
return max(len(t) for t in tensor)
def tokenize(lang):
lang_tokenizer = tf.keras.preprocessing.text.Tokenizer(
filters='')
lang_tokenizer.fit_on_texts(lang)
tensor = lang_tokenizer.texts_to_sequences(lang)
tensor = tf.keras.preprocessing.sequence.pad_sequences(tensor,
padding='post')
return tensor, lang_tokenizer
def load_dataset(path, num_examples=None):
# 创建清理过的输入输出对
targ_lang, inp_lang = create_dataset(path, num_examples)
input_tensor, inp_lang_tokenizer = tokenize(inp_lang)
target_tensor, targ_lang_tokenizer = tokenize(targ_lang)
return input_tensor, target_tensor, inp_lang_tokenizer, targ_lang_tokenizer
```
### 限制数据集的大小以加快实验速度(可选)
在超过 10 万个句子的完整数据集上训练需要很长时间。为了更快地训练,我们可以将数据集的大小限制为 3 万个句子(当然,翻译质量也会随着数据的减少而降低):
```
# 尝试实验不同大小的数据集
num_examples = 30000
input_tensor, target_tensor, inp_lang, targ_lang = load_dataset(path_to_file, num_examples)
# 计算目标张量的最大长度 (max_length)
max_length_targ, max_length_inp = max_length(target_tensor), max_length(input_tensor)
# 采用 80 - 20 的比例切分训练集和验证集
input_tensor_train, input_tensor_val, target_tensor_train, target_tensor_val = train_test_split(input_tensor, target_tensor, test_size=0.2)
# 显示长度
print(len(input_tensor_train), len(target_tensor_train), len(input_tensor_val), len(target_tensor_val))
def convert(lang, tensor):
for t in tensor:
if t!=0:
print ("%d ----> %s" % (t, lang.index_word[t]))
print ("Input Language; index to word mapping")
convert(inp_lang, input_tensor_train[0])
print ()
print ("Target Language; index to word mapping")
convert(targ_lang, target_tensor_train[0])
```
### 创建一个 tf.data 数据集
```
BUFFER_SIZE = len(input_tensor_train)
BATCH_SIZE = 64
steps_per_epoch = len(input_tensor_train)//BATCH_SIZE
embedding_dim = 256
units = 1024
vocab_inp_size = len(inp_lang.word_index)+1
vocab_tar_size = len(targ_lang.word_index)+1
dataset = tf.data.Dataset.from_tensor_slices((input_tensor_train, target_tensor_train)).shuffle(BUFFER_SIZE)
dataset = dataset.batch(BATCH_SIZE, drop_remainder=True)
example_input_batch, example_target_batch = next(iter(dataset))
example_input_batch.shape, example_target_batch.shape
```
## 编写编码器 (encoder) 和解码器 (decoder) 模型
实现一个基于注意力的编码器 - 解码器模型。关于这种模型,你可以阅读 TensorFlow 的 [神经机器翻译 (序列到序列) 教程](https://github.com/tensorflow/nmt)。本示例采用一组更新的 API。此笔记本实现了上述序列到序列教程中的 [注意力方程式](https://github.com/tensorflow/nmt#background-on-the-attention-mechanism)。下图显示了注意力机制为每个输入单词分配一个权重,然后解码器将这个权重用于预测句子中的下一个单词。下图和公式是 [Luong 的论文](https://arxiv.org/abs/1508.04025v5)中注意力机制的一个例子。
<img src="https://www.tensorflow.org/images/seq2seq/attention_mechanism.jpg" width="500" alt="attention mechanism">
输入经过编码器模型,编码器模型为我们提供形状为 *(批大小,最大长度,隐藏层大小)* 的编码器输出和形状为 *(批大小,隐藏层大小)* 的编码器隐藏层状态。
下面是所实现的方程式:
<img src="https://www.tensorflow.org/images/seq2seq/attention_equation_0.jpg" alt="attention equation 0" width="800">
<img src="https://www.tensorflow.org/images/seq2seq/attention_equation_1.jpg" alt="attention equation 1" width="800">
本教程的编码器采用 [Bahdanau 注意力](https://arxiv.org/pdf/1409.0473.pdf)。在用简化形式编写之前,让我们先决定符号:
* FC = 完全连接(密集)层
* EO = 编码器输出
* H = 隐藏层状态
* X = 解码器输入
以及伪代码:
* `score = FC(tanh(FC(EO) + FC(H)))`
* `attention weights = softmax(score, axis = 1)`。 Softmax 默认被应用于最后一个轴,但是这里我们想将它应用于 *第一个轴*, 因为分数 (score) 的形状是 *(批大小,最大长度,隐藏层大小)*。最大长度 (`max_length`) 是我们的输入的长度。因为我们想为每个输入分配一个权重,所以 softmax 应该用在这个轴上。
* `context vector = sum(attention weights * EO, axis = 1)`。选择第一个轴的原因同上。
* `embedding output` = 解码器输入 X 通过一个嵌入层。
* `merged vector = concat(embedding output, context vector)`
* 此合并后的向量随后被传送到 GRU
每个步骤中所有向量的形状已在代码的注释中阐明:
```
class Encoder(tf.keras.Model):
def __init__(self, vocab_size, embedding_dim, enc_units, batch_sz):
super(Encoder, self).__init__()
self.batch_sz = batch_sz
self.enc_units = enc_units
self.embedding = tf.keras.layers.Embedding(vocab_size, embedding_dim)
self.gru = tf.keras.layers.GRU(self.enc_units,
return_sequences=True,
return_state=True,
recurrent_initializer='glorot_uniform')
def call(self, x, hidden):
x = self.embedding(x)
output, state = self.gru(x, initial_state = hidden)
return output, state
def initialize_hidden_state(self):
return tf.zeros((self.batch_sz, self.enc_units))
encoder = Encoder(vocab_inp_size, embedding_dim, units, BATCH_SIZE)
# 样本输入
sample_hidden = encoder.initialize_hidden_state()
sample_output, sample_hidden = encoder(example_input_batch, sample_hidden)
print ('Encoder output shape: (batch size, sequence length, units) {}'.format(sample_output.shape))
print ('Encoder Hidden state shape: (batch size, units) {}'.format(sample_hidden.shape))
class BahdanauAttention(tf.keras.layers.Layer):
def __init__(self, units):
super(BahdanauAttention, self).__init__()
self.W1 = tf.keras.layers.Dense(units)
self.W2 = tf.keras.layers.Dense(units)
self.V = tf.keras.layers.Dense(1)
def call(self, query, values):
# 隐藏层的形状 == (批大小,隐藏层大小)
# hidden_with_time_axis 的形状 == (批大小,1,隐藏层大小)
# 这样做是为了执行加法以计算分数
hidden_with_time_axis = tf.expand_dims(query, 1)
# 分数的形状 == (批大小,最大长度,1)
# 我们在最后一个轴上得到 1, 因为我们把分数应用于 self.V
# 在应用 self.V 之前,张量的形状是(批大小,最大长度,单位)
score = self.V(tf.nn.tanh(
self.W1(values) + self.W2(hidden_with_time_axis)))
# 注意力权重 (attention_weights) 的形状 == (批大小,最大长度,1)
attention_weights = tf.nn.softmax(score, axis=1)
# 上下文向量 (context_vector) 求和之后的形状 == (批大小,隐藏层大小)
context_vector = attention_weights * values
context_vector = tf.reduce_sum(context_vector, axis=1)
return context_vector, attention_weights
attention_layer = BahdanauAttention(10)
attention_result, attention_weights = attention_layer(sample_hidden, sample_output)
print("Attention result shape: (batch size, units) {}".format(attention_result.shape))
print("Attention weights shape: (batch_size, sequence_length, 1) {}".format(attention_weights.shape))
class Decoder(tf.keras.Model):
def __init__(self, vocab_size, embedding_dim, dec_units, batch_sz):
super(Decoder, self).__init__()
self.batch_sz = batch_sz
self.dec_units = dec_units
self.embedding = tf.keras.layers.Embedding(vocab_size, embedding_dim)
self.gru = tf.keras.layers.GRU(self.dec_units,
return_sequences=True,
return_state=True,
recurrent_initializer='glorot_uniform')
self.fc = tf.keras.layers.Dense(vocab_size)
# 用于注意力
self.attention = BahdanauAttention(self.dec_units)
def call(self, x, hidden, enc_output):
# 编码器输出 (enc_output) 的形状 == (批大小,最大长度,隐藏层大小)
context_vector, attention_weights = self.attention(hidden, enc_output)
# x 在通过嵌入层后的形状 == (批大小,1,嵌入维度)
x = self.embedding(x)
# x 在拼接 (concatenation) 后的形状 == (批大小,1,嵌入维度 + 隐藏层大小)
x = tf.concat([tf.expand_dims(context_vector, 1), x], axis=-1)
# 将合并后的向量传送到 GRU
output, state = self.gru(x)
# 输出的形状 == (批大小 * 1,隐藏层大小)
output = tf.reshape(output, (-1, output.shape[2]))
# 输出的形状 == (批大小,vocab)
x = self.fc(output)
return x, state, attention_weights
decoder = Decoder(vocab_tar_size, embedding_dim, units, BATCH_SIZE)
sample_decoder_output, _, _ = decoder(tf.random.uniform((64, 1)),
sample_hidden, sample_output)
print ('Decoder output shape: (batch_size, vocab size) {}'.format(sample_decoder_output.shape))
```
## 定义优化器和损失函数
```
optimizer = tf.keras.optimizers.Adam()
loss_object = tf.keras.losses.SparseCategoricalCrossentropy(
from_logits=True, reduction='none')
def loss_function(real, pred):
mask = tf.math.logical_not(tf.math.equal(real, 0))
loss_ = loss_object(real, pred)
mask = tf.cast(mask, dtype=loss_.dtype)
loss_ *= mask
return tf.reduce_mean(loss_)
```
## 检查点(基于对象保存)
```
checkpoint_dir = './training_checkpoints'
checkpoint_prefix = os.path.join(checkpoint_dir, "ckpt")
checkpoint = tf.train.Checkpoint(optimizer=optimizer,
encoder=encoder,
decoder=decoder)
```
## 训练
1. 将 *输入* 传送至 *编码器*,编码器返回 *编码器输出* 和 *编码器隐藏层状态*。
2. 将编码器输出、编码器隐藏层状态和解码器输入(即 *开始标记*)传送至解码器。
3. 解码器返回 *预测* 和 *解码器隐藏层状态*。
4. 解码器隐藏层状态被传送回模型,预测被用于计算损失。
5. 使用 *教师强制 (teacher forcing)* 决定解码器的下一个输入。
6. *教师强制* 是将 *目标词* 作为 *下一个输入* 传送至解码器的技术。
7. 最后一步是计算梯度,并将其应用于优化器和反向传播。
```
@tf.function
def train_step(inp, targ, enc_hidden):
loss = 0
with tf.GradientTape() as tape:
enc_output, enc_hidden = encoder(inp, enc_hidden)
dec_hidden = enc_hidden
dec_input = tf.expand_dims([targ_lang.word_index['<start>']] * BATCH_SIZE, 1)
# 教师强制 - 将目标词作为下一个输入
for t in range(1, targ.shape[1]):
# 将编码器输出 (enc_output) 传送至解码器
predictions, dec_hidden, _ = decoder(dec_input, dec_hidden, enc_output)
loss += loss_function(targ[:, t], predictions)
# 使用教师强制
dec_input = tf.expand_dims(targ[:, t], 1)
batch_loss = (loss / int(targ.shape[1]))
variables = encoder.trainable_variables + decoder.trainable_variables
gradients = tape.gradient(loss, variables)
optimizer.apply_gradients(zip(gradients, variables))
return batch_loss
EPOCHS = 10
for epoch in range(EPOCHS):
start = time.time()
enc_hidden = encoder.initialize_hidden_state()
total_loss = 0
for (batch, (inp, targ)) in enumerate(dataset.take(steps_per_epoch)):
batch_loss = train_step(inp, targ, enc_hidden)
total_loss += batch_loss
if batch % 100 == 0:
print('Epoch {} Batch {} Loss {:.4f}'.format(epoch + 1,
batch,
batch_loss.numpy()))
# 每 2 个周期(epoch),保存(检查点)一次模型
if (epoch + 1) % 2 == 0:
checkpoint.save(file_prefix = checkpoint_prefix)
print('Epoch {} Loss {:.4f}'.format(epoch + 1,
total_loss / steps_per_epoch))
print('Time taken for 1 epoch {} sec\n'.format(time.time() - start))
```
## 翻译
* 评估函数类似于训练循环,不同之处在于在这里我们不使用 *教师强制*。每个时间步的解码器输入是其先前的预测、隐藏层状态和编码器输出。
* 当模型预测 *结束标记* 时停止预测。
* 存储 *每个时间步的注意力权重*。
请注意:对于一个输入,编码器输出仅计算一次。
```
def evaluate(sentence):
attention_plot = np.zeros((max_length_targ, max_length_inp))
sentence = preprocess_sentence(sentence)
inputs = [inp_lang.word_index[i] for i in sentence.split(' ')]
inputs = tf.keras.preprocessing.sequence.pad_sequences([inputs],
maxlen=max_length_inp,
padding='post')
inputs = tf.convert_to_tensor(inputs)
result = ''
hidden = [tf.zeros((1, units))]
enc_out, enc_hidden = encoder(inputs, hidden)
dec_hidden = enc_hidden
dec_input = tf.expand_dims([targ_lang.word_index['<start>']], 0)
for t in range(max_length_targ):
predictions, dec_hidden, attention_weights = decoder(dec_input,
dec_hidden,
enc_out)
# 存储注意力权重以便后面制图
attention_weights = tf.reshape(attention_weights, (-1, ))
attention_plot[t] = attention_weights.numpy()
predicted_id = tf.argmax(predictions[0]).numpy()
result += targ_lang.index_word[predicted_id] + ' '
if targ_lang.index_word[predicted_id] == '<end>':
return result, sentence, attention_plot
# 预测的 ID 被输送回模型
dec_input = tf.expand_dims([predicted_id], 0)
return result, sentence, attention_plot
# 注意力权重制图函数
def plot_attention(attention, sentence, predicted_sentence):
fig = plt.figure(figsize=(10,10))
ax = fig.add_subplot(1, 1, 1)
ax.matshow(attention, cmap='viridis')
fontdict = {'fontsize': 14}
ax.set_xticklabels([''] + sentence, fontdict=fontdict, rotation=90)
ax.set_yticklabels([''] + predicted_sentence, fontdict=fontdict)
ax.xaxis.set_major_locator(ticker.MultipleLocator(1))
ax.yaxis.set_major_locator(ticker.MultipleLocator(1))
plt.show()
def translate(sentence):
result, sentence, attention_plot = evaluate(sentence)
print('Input: %s' % (sentence))
print('Predicted translation: {}'.format(result))
attention_plot = attention_plot[:len(result.split(' ')), :len(sentence.split(' '))]
plot_attention(attention_plot, sentence.split(' '), result.split(' '))
```
## 恢复最新的检查点并验证
```
# 恢复检查点目录 (checkpoint_dir) 中最新的检查点
checkpoint.restore(tf.train.latest_checkpoint(checkpoint_dir))
translate(u'hace mucho frio aqui.')
translate(u'esta es mi vida.')
translate(u'¿todavia estan en casa?')
# 错误的翻译
translate(u'trata de averiguarlo.')
```
## 下一步
* [下载一个不同的数据集](http://www.manythings.org/anki/)实验翻译,例如英语到德语或者英语到法语。
* 实验在更大的数据集上训练,或者增加训练周期。
| github_jupyter |
```
import foolbox
import numpy as np
import torchvision.models as models
# instantiate model (supports PyTorch, Keras, TensorFlow (Graph and Eager), MXNet and many more)
model = models.resnet18(pretrained=True).eval()
preprocessing = dict(mean=[0.485, 0.456, 0.406], std=[0.229, 0.224, 0.225], axis=-3)
fmodel = foolbox.models.PyTorchModel(model, bounds=(0, 1), num_classes=1000, preprocessing=preprocessing)
!wget https://gist.githubusercontent.com/yrevar/942d3a0ac09ec9e5eb3a/raw/238f720ff059c1f82f368259d1ca4ffa5dd8f9f5/imagenet1000_clsidx_to_labels.txt
import json
name = 'imagenet1000_clsidx_to_labels.txt'
with open(name,'r') as f:
txt = f.readlines()
text = list(map(lambda x:x.replace("'","\""),txt))
labels_text = text
# get a batch of images and labels and print the accuracy
images, labels = foolbox.utils.samples(dataset='imagenet', batchsize=16, data_format='channels_first', bounds=(0, 1))
print(np.mean(fmodel.forward(images).argmax(axis=-1) == labels))
# -> 0.9375
len(labels[:1])
# apply the attack
attack = foolbox.attacks.SaliencyMapAttack(fmodel)
adversarials = attack(images[:1], labels[:1])
# if the i'th image is misclassfied without a perturbation, then adversarials[i] will be the same as images[i]
# if the attack fails to find an adversarial for the i'th image, then adversarials[i] will all be np.nan
# Foolbox guarantees that all returned adversarials are in fact in adversarials
print(np.mean(fmodel.forward(adversarials).argmax(axis=-1) == labels))
# -> 0.0
# ---
# In rare cases, it can happen that attacks return adversarials that are so close to the decision boundary,
# that they actually might end up on the other (correct) side if you pass them through the model again like
# above to get the adversarial class. This is because models are not numerically deterministic (on GPU, some
# operations such as `sum` are non-deterministic by default) and indepedent between samples (an input might
# be classified differently depending on the other inputs in the same batch).
# You can always get the actual adversarial class that was observed for that sample by Foolbox by
# passing `unpack=False` to get the actual `Adversarial` objects:
attack = foolbox.attacks.SaliencyMapAttack(fmodel, distance=foolbox.distances.Linf)
adversarials = attack(images, labels, unpack=False)
adversarial_classes = np.asarray([a.adversarial_class for a in adversarials])
print(labels)
print(adversarial_classes)
print(np.mean(adversarial_classes == labels)) # will always be 0.0
labels
def transform_shape(img):
shape = img[0].shape
res = [[0 for i in range(shape[0])] for j in range(shape[1])]
for i in range(shape[0]):
for j in range(shape[1]):
res[i][j] = [img[0][i][j], img[1][i][j], img[2][i][j]]
return np.array(res)
adv = adversarials[8]
perturbed, original = adv.perturbed, adv.unperturbed
perturbed = transform_shape(perturbed)
original = transform_shape(original)
import matplotlib.pyplot as plt
plt.figure('FGSM')
plt.subplot(1,3,1)
plt.title(labels_text[adv.adversarial_class])
plt.imshow(perturbed)
plt.subplot(1,3,2)
plt.title(labels_text[adv.original_class])
plt.imshow(original)
plt.subplot(1,3,3)
difference = perturbed - original
maxi, mini = np.max(difference), np.min(difference)
difference = (difference - mini) / (maxi- mini)
#print(np.max(difference), np.min(difference))
plt.imshow(difference)
plt.savefig('FGSM')
plt.show()
ls
!pwd
# The `Adversarial` objects also provide a `distance` attribute. Note that the distances
# can be 0 (misclassified without perturbation) and inf (attack failed).
distances = np.asarray([a.distance.value for a in adversarials])
print("{:.1e}, {:.1e}, {:.1e}".format(distances.min(), np.median(distances), distances.max()))
print("{} of {} attacks failed".format(sum(adv.distance.value == np.inf for adv in adversarials), len(adversarials)))
print("{} of {} inputs misclassified without perturbation".format(sum(adv.distance.value == 0 for adv in adversarials), len(adversarials)))
!git add -A
```
| github_jupyter |
# GOOGLE PLAY STORE - Transforming Raw to Clean Data
_______________________
* Consideration: source data was scraped from the web
## Objectives:
* Create a cleaned up version of the Google Play Store Source Data by filtering:
- Games with no reviews
- Duplicates
- Converting all ratings, reviews, installs, and price to uniform types and formats by column
* Subsequently, make sure there's no duplicate app names or double counting / aggegration; organize by apps, and remove exact duplicates, and or take the higher of the two
* Final Product should be a cleaned gps source data we'll use to create charts with
```
# Import Dependencies
import os
import csv
import numpy as np
import pandas as pd
# Set up to load googleplaystore.csv into the 'chamber'
# This is the bullet
playstore_csvpath= os.path.join(".","googleplaystore.csv")
# Load the bullet into the "gun" aka Pandas
with open(playstore_csvpath, newline= "") as csvfile:
csv_reader= csv.reader(csvfile, delimiter=",")
# Pull the trigger and covert the original CSV to a dataframe, and print the DF
gps_sourcedata_df = pd.read_csv("./googleplaystore.csv")
# Run to see count
gps_sourcedata_df.count()
# To see count and type
#type(gps_sourcedata_df.head())
# Identify Columns we want to remove and keep
gps_sourcedata_df
# Sort by Reviews, and drop any cells with missing information to make all columns equal
#gps_sourcedata_df = gps_sourcedata_df.sort_values(by= ["Reviews"], ascending=True).dropna(how="any")
gps_sourcedata_df = gps_sourcedata_df.sort_values(by= ["Reviews"], ascending=False)
gps_sourcedata_df
gps_sourcedata_df.count()
```
### Only run the ".drop function once. if you have to restart the kernel, unhash it and run it.
### if you try to run it twice, it will say an error because nothign is there to drop
#### it may take a few times. run the 'gps_sourcedata_df' to view it a few times to make sure.
```
# create a list to drop unwanted columns and store into a new dataframe
#to_drop =['Android Ver', 'Current Ver', 'Size']
#gps_sourcedata_df.drop(to_drop, inplace=True, axis=1)
gps_sourcedata_df
gps_sourcedata_df['Reviews'].value_counts().head()
#gps_sourcedata_df['Reviews'].describe()
# Sort the file by
gps_sourcedata_df = gps_sourcedata_df.sort_values(['Reviews'], ascending=False)
#gps_sourcedata_df.max()
gps_sourcedata_df
gps_sourcedata_df.info()
#create a list to drop unwanted columns and store into a new dataframe
#only run the drop once
to_drop =['Android Ver', 'Current Ver', 'Size']
gps_sourcedata_df = gps_sourcedata_df.drop(columns=to_drop)
#gps_cleaning_df
#gps_sourcedata_df.count()
gps_sourcedata_df
#gps_sourcedata_df['App'].value_counts()
gps_sourcedata_df['App'].unique()
# Clean up installs - can use replace or .map function to remove '+' signs
# Khaled said once next time just remove commas in between text
gps_sourcedata_df['Installs'] = gps_sourcedata_df['Installs'].map(lambda x: str(x)[:-1])
gps_sourcedata_df.info()
# Remove commas in betwen numbers in installs, and convert to int64
gps_sourcedata_df = gps_sourcedata_df.dropna(how="any")
gps_sourcedata_df['Installs'] = [x.replace(",","") for x in gps_sourcedata_df['Installs']]
# Remove commas in betwen numbers in Reviews, and convert to int64
gps_sourcedata_df['Reviews'] = [x.replace(",","") for x in gps_sourcedata_df['Reviews']]
gps_sourcedata_df['Reviews'].astype(np.int64).head()
# Remove $ in betwen numbers in Reviews, and convert to float64
gps_sourcedata_df['Price'] = [x.replace("$","") for x in gps_sourcedata_df['Price']]
#gps_sourcedata_df['Price'] = gps_sourcedata_df['Price'].astype(np.float64)
gps_sourcedata_df
# Sort Ratings - right way below:
gps_sourcedata_df = gps_sourcedata_df.sort_values(['Rating'], ascending=False)
gps_sourcedata_df.head()
#gps_sourcedata_df = gps_sourcedata_df.sort_values(['Reviews'], ascending=False)
# this is wrong: gps_sourcedata_df['Rating'].sort_values(gps_sourcedata_df['Rating'], ascending=False)
#gps_sourcedata_df.sort_values(['Category'], ascending=True)
category_list = np.sort(gps_sourcedata_df['Category'].unique())
# for x in category_list:
# print(x)
#gps_sourcedata_df = gps_sourcedata_df.sort_values(['Content Rating'], ascending=False)
#gps_sourcedata_df.sort_values(['Content Rating'], ascending=True)
category_list = np.sort(gps_sourcedata_df['Content Rating'].unique())
for x in category_list:
print(x)
#gps_sourcedata_df.head()
gps_sourcedata_df['Content Rating'] = [x.replace("Everyone 10+","Everyone") for x in gps_sourcedata_df['Content Rating']]
gps_sourcedata_df['Content Rating'].head()
gps_sourcedata_df = gps_sourcedata_df.sort_values(['Category'], ascending=True)
x_list = gps_sourcedata_df['Category'].unique()
for x in x_list:
print(x)
gps_sourcedata_df.head()
gps_sourcedata_df['Reviews'] = gps_sourcedata_df['Reviews'].astype(np.int64)
gps_sourcedata_df = gps_sourcedata_df.sort_values(['Reviews'], ascending=False)
gps_filterdata_df = gps_sourcedata_df.drop_duplicates(['App']).sort_values(['Reviews'], ascending=False)
gps_filterdata_df
gps_filterdata_df.describe()
gps_filterdata_df.head()
top_quartile = np.percentile(gps_filterdata_df['Reviews'], 75)
top_quartile
# for notes. don't use.
# top_quartile = int((gps_filterdata_df['Reviews'].max()*.5))
top_quartile_data_df = gps_filterdata_df.loc[gps_filterdata_df['Reviews'] > top_quartile]
top_quartile_data_df.head(20)
top_quartile_data_df['Category'].unique()
top_quartile_data_df = top_quartile_data_df.sort_values(['Category'], ascending=True)
x_list = top_quartile_data_df['Category'].unique()
for x in x_list:
print(x)
```
### Clean up Cateogry Columns to set up pie and bar charts:
```
# top_quartile_data_df["Category"] = [x.replace("FINANCE", "Business") for x in top_quartile_data_df["Category"]]
# top_quartile_data_df["Category"] = [x.replace("LIBRARIES_AND_DEMO", "Education") for x in top_quartile_data_df["Category"]]
# top_quartile_data_df["Category"] = [x.replace("GAMES", "Games") for x in top_quartile_data_df["Category"]]
# top_quartile_data_df["Category"] = [x.replace("COMICS", "Games") for x in top_quartile_data_df["Category"]]
# top_quartile_data_df["Category"] = [x.replace("HEALTH_AND_FITNESS", "Health and Fitness") for x in top_quartile_data_df["Category"]]
# top_quartile_data_df["Category"] = [x.replace("MEDICAL", "Health and Fitness") for x in top_quartile_data_df["Category"]]
# top_quartile_data_df["Category"] = [x.replace("DATING","Life Stlye") for x in top_quartile_data_df["Category"]]
# top_quartile_data_df["Category"] = [x.replace("BEAUTY", "Life Style") for x in top_quartile_data_df["Category"]]
# top_quartile_data_df["Category"] = [x.replace("PARENTING", "Life Style") for x in top_quartile_data_df["Category"]]
# top_quartile_data_df["Category"] = [x.replace("LIFE_STYLE", "Life Style") for x in top_quartile_data_df["Category"]]
# top_quartile_data_df["Category"] = [x.replace("BOOKS_AND_REFERENCES", "Productivity") for x in top_quartile_data_df["Category"]]
# top_quartile_data_df["Category"] = [x.replace("TRAVEL_AND_LOCAL", "Productivity") for x in top_quartile_data_df["Category"]]
# top_quartile_data_df["Category"] = [x.replace("AUTO_AND_VEHICLE", "Productivity") for x in top_quartile_data_df["Category"]]
# top_quartile_data_df["Category"] = [x.replace("PRODUCTIVITY", "Productivity") for x in top_quartile_data_df["Category"]]
# top_quartile_data_df["Category"] = [x.replace("FOOD_AND_DRINK", "Social Networking") for x in top_quartile_data_df["Category"]]
# top_quartile_data_df["Category"] = [x.replace("FAMILY", "Social Networking") for x in top_quartile_data_df["Category"]]
# top_quartile_data_df["Category"] = [x.replace("ENTERTAINMENT", "Social Networking") for x in top_quartile_data_df["Category"]]
# top_quartile_data_df["Category"] = [x.replace("COMMUNITCATION", "Social Networking") for x in top_quartile_data_df["Category"]]
# top_quartile_data_df["Category"] = [x.replace("NEWS_AND_MAGAZING", "Social Networking") for x in top_quartile_data_df["Category"]]
# top_quartile_data_df["Category"] = [x.replace("PERSONALIZATION", "Social Networking") for x in top_quartile_data_df["Category"]]
# top_quartile_data_df["Category"] = [x.replace("SOCIAL", "Social Networking") for x in top_quartile_data_df["Category"]]
# top_quartile_data_df["Category"] = [x.replace("LIFE_STYLE", "Social Networking") for x in top_quartile_data_df["Category"]]
# top_quartile_data_df["Category"] = [x.replace("SPORTS", "Sports") for x in top_quartile_data_df["Category"]]
# top_quartile_data_df["Category"] = [x.replace("VIDEO_PLAYERS", "Utility") for x in top_quartile_data_df["Category"]]
# top_quartile_data_df["Category"] = [x.replace("MAPS_AND_NAVIGATION", "Utility") for x in top_quartile_data_df["Category"]]
# top_quartile_data_df["Category"] = [x.replace("TOOLS", "Utility") for x in top_quartile_data_df["Category"]]
# top_quartile_data_df["Category"] = [x.replace("TRAVEL_AND_LOCATION", "Utility") for x in top_quartile_data_df["Category"]]
# top_quartile_data_df["Category"] = [x.replace("WEATHER", "Weather") for x in top_quartile_data_df["Category"]]
top_quartile_data_df
top_quartile_data_df["Category"] = [x.replace("Productivity", "Productivity") for x in top_quartile_data_df["Category"]]
top_quartile_data_df
plt.figure(2, figsize=(6,8))
values_list = [urb_fares, rural_fares, suburb_fares]
labels = ['Urban', 'Rural', 'Suburban']
colors = ['lightcoral', 'gold', 'lightskyblue']
explode = (0.1, 0, 0)
for city in fare_city_list:
plt.pie(fare_city_list, labels=labels, colors=colors, explode=explode, autopct="%1.1f%%", shadow=True, startangle=270)
plt.axis('equal')
plt.title('% of Total Fares by City Type')
plt.show()
plt.savefig('./Pyber_TotalFares_CityType.png')
# slicing pulling imporant data out strainging our data to condense it to juice
data_file_pd.sort_values( by= ["Installs","Rating"], ascending=False).dropna(how="any")
# get top 20 apps
business
education
games
health_fitness
lifestyle
photo_video
productivity
social Networking
sports
travel
utility
weather
plt.figure(1, figsize=(6,6))
fare_city_list = [urb_fares, rural_fares, suburb_fares] # rename: with all categories
labels = ['Urban', 'Rural', 'Suburban'] # list of all category names
colors = ['lightcoral', 'gold', 'lightskyblue']
explode = (0.1, 0, 0)
for cat in category_list:
plt.pie(fare_city_list, labels=labels, colors=colors, explode=explode, autopct="%1.1f%%", shadow=True, startangle=270)
plt.axis('equal')
plt.title('% of Total Fares by City Type')
plt.show()
plt.savefig('./Pyber_TotalFares_CityType.png')
data_file_pd.drop_duplicates(["App"], keep="first")
data_file_pd.sort_values( by= ["Installs","Rating"], ascending=False).dropna(how="any")
data_file_pd.sort_values( by= ["Installs","Rating"], ascending=False).dropna(how="any")
data_file_pd.drop_duplicates(["App"], keep="first")
```
Top 20 for installs and ratings : ez /
remove the + field when data scrubbing so we can chart issues
list of 4 apps to check success. /
factor in categories on how we can compare the two
get rod of duplicates
sory by app name and where app name == true ommitt data /
remove rows and append findings to a clean DF thats sorted
investigate unique instances and removal /
| github_jupyter |
# Now, we can start a new training job
We'll send a zip file called **trainingjob.zip**, with the following structure:
- trainingjob.json (Sagemaker training job descriptor)
- assets/deploy-model-prd.yml (Cloudformation for deploying our model into Production)
- assets/deploy-model-dev.yml (Cloudformation for deploying our model into Development)
## Let's start by defining the hyperparameters for both algorithms
```
hyperparameters = {
"logistic_max_iter": 100,
"logistic_solver": "lbfgs",
"random_forest_max_depth": 10,
"random_forest_n_jobs": 5,
"random_forest_verbose": 1
}
```
## Then, let's create the trainingjob descriptor
```
import time
import sagemaker
import boto3
sts_client = boto3.client("sts")
model_prefix='iris-model'
account_id = sts_client.get_caller_identity()["Account"]
region = boto3.session.Session().region_name
training_image = '{}.dkr.ecr.{}.amazonaws.com/{}:latest'.format(account_id, region, model_prefix)
roleArn = "arn:aws:iam::{}:role/MLOps".format(account_id)
timestamp = time.strftime('-%Y-%m-%d-%H-%M-%S', time.gmtime())
job_name = model_prefix + timestamp
sagemaker_session = sagemaker.Session()
training_params = {}
# Here we set the reference for the Image Classification Docker image, stored on ECR (https://aws.amazon.com/pt/ecr/)
training_params["AlgorithmSpecification"] = {
"TrainingImage": training_image,
"TrainingInputMode": "File"
}
# The IAM role with all the permissions given to Sagemaker
training_params["RoleArn"] = roleArn
# Here Sagemaker will store the final trained model
training_params["OutputDataConfig"] = {
"S3OutputPath": 's3://{}/{}'.format(sagemaker_session.default_bucket(), model_prefix)
}
# This is the config of the instance that will execute the training
training_params["ResourceConfig"] = {
"InstanceCount": 1,
"InstanceType": "ml.m4.xlarge",
"VolumeSizeInGB": 30
}
# The job name. You'll see this name in the Jobs section of the Sagemaker's console
training_params["TrainingJobName"] = job_name
for i in hyperparameters:
hyperparameters[i] = str(hyperparameters[i])
# Here you will configure the hyperparameters used for training your model.
training_params["HyperParameters"] = hyperparameters
# Training timeout
training_params["StoppingCondition"] = {
"MaxRuntimeInSeconds": 360000
}
# The algorithm currently only supports fullyreplicated model (where data is copied onto each machine)
training_params["InputDataConfig"] = []
# Please notice that we're using application/x-recordio for both
# training and validation datasets, given our dataset is formated in RecordIO
# Here we set training dataset
# Training data should be inside a subdirectory called "train"
training_params["InputDataConfig"].append({
"ChannelName": "training",
"DataSource": {
"S3DataSource": {
"S3DataType": "S3Prefix",
"S3Uri": 's3://{}/{}/input'.format(sagemaker_session.default_bucket(), model_prefix),
"S3DataDistributionType": "FullyReplicated"
}
},
"ContentType": "text/csv",
"CompressionType": "None"
})
training_params["Tags"] = []
```
## Before we start the training process, we need to upload our dataset to S3
```
import sagemaker
# Get the current Sagemaker session
sagemaker_session = sagemaker.Session()
default_bucket = sagemaker_session.default_bucket()
role = sagemaker.get_execution_role()
!mkdir -p input/data/training
import pandas as pd
import numpy as np
from sklearn import datasets
iris = datasets.load_iris()
dataset = np.insert(iris.data, 0, iris.target,axis=1)
pd = pd.DataFrame(data=dataset, columns=['iris_id'] + iris.feature_names)
pd.to_csv('input/data/training/iris.csv', header=None, index=False, sep=',', encoding='utf-8')
data_location = sagemaker_session.upload_data(path='input/data/training', key_prefix='iris-model/input')
```
## Alright! Now it's time to start the training process
```
import boto3
import io
import zipfile
import json
s3 = boto3.client('s3')
sts_client = boto3.client("sts")
session = boto3.session.Session()
account_id = sts_client.get_caller_identity()["Account"]
region = session.region_name
bucket_name = "mlops-%s-%s" % (region, account_id)
key_name = "training_jobs/iris_model/trainingjob.zip"
zip_buffer = io.BytesIO()
with zipfile.ZipFile(zip_buffer, 'a') as zf:
zf.writestr('trainingjob.json', json.dumps(training_params))
zf.writestr('assets/deploy-model-prd.yml', open('../../assets/deploy-model-prd.yml', 'r').read())
zf.writestr('assets/deploy-model-dev.yml', open('../../assets/deploy-model-dev.yml', 'r').read())
zip_buffer.seek(0)
s3.put_object(Bucket=bucket_name, Key=key_name, Body=bytearray(zip_buffer.read()))
```
### Ok, now open the AWS console in another tab and go to the CodePipeline console to see the status of our building pipeline
> Finally, click here [NOTEBOOK](04_Check%20Progress%20and%20Test%20the%20endpoint.ipynb) to see the progress and test your endpoint
| github_jupyter |
# Using Pypbomb
Let's go over some quick examples of how you might use pypbomb to design a detonation tube for your research.
```
from itertools import product
import warnings
import cantera as ct
import numpy as np
import pandas as pd
import pint
import seaborn as sns
from matplotlib import pyplot as plt
from matplotlib.ticker import MaxNLocator
from pypbomb import Tube, Flange, DDT, Window, Bolt
ureg = pint.UnitRegistry()
quant = ureg.Quantity
sns.set_context("notebook")
sns.set_style("white")
```
## Determine tube size and operating limits
First let us pick a mixture of stoichiometric propane/air. For this example we will use ``gri30.cti`` as the mechanism of choice for our Cantera calculations.
```
fuel = "C3H8"
oxidizer = "O2:1 N2:3.76"
material = "316L"
mechanism = "gri30.cti"
gas = ct.Solution(mechanism)
gas.set_equivalence_ratio(1, fuel, oxidizer)
```
Next, let's consider 316L since it's a commonly used stainless steel.
```
material = "316L"
```
At this point we should probably figure out which schedules are available across all of the potential pipe sizes that we'd like to consider.
```
potential_sizes = ["1", "3", "4", "6"]
common_sizes = set(Tube.get_available_pipe_schedules(potential_sizes[0]))
for size in potential_sizes[1:]:
common_sizes.intersection_update(
set(Tube.get_available_pipe_schedules(size))
)
common_sizes
```
Given these options, let's choose schedules 40, 80, 120, and XXH for consideration.
```
potential_schedules = ["40", "80", "XXH"]
```
Let's also look at a range of operating temperatures, in case we need to preheat our tube.
```
initial_temperatures = quant(
np.linspace(20, 400, 6),
"degC"
)
```
Now let's figure out what we can do with each combination of pipe size, pipe schedule, and initial temperature. The steps we will use for each combination are:
1. look up the tube dimensions,
2. look up the maximum allowable stress,
3. calculate the corresponding maximum pressure,
4. look up the elastic modulus, density, and Poisson ratio of our tube material, and
5. calculate the maximum safe initial pressure that we can test at.
Note that we are setting ``multiprocessing=False``, since multiprocessing can cause unexpected misbehavior when run from within a jupyter notebook.
```
calculate_results = True
results_file = "tube_size_results.h5"
combinations = list(
product(
potential_schedules,
potential_sizes,
initial_temperatures
)
)
if calculate_results:
results = pd.DataFrame(
columns=[
"schedule",
"size",
"max initial pressure (psi)",
"initial temperature (F)",
"tube_temp",
"max_pressure",
"DLF",
]
)
for i, (schedule, size, initial_temperature) in enumerate(combinations):
dims = Tube.get_dimensions(
size,
schedule,
unit_registry=ureg
)
max_stress = Tube.calculate_max_stress(
initial_temperature,
material,
welded=False,
unit_registry=ureg
)
max_pressure = Tube.calculate_max_pressure(
dims["inner_diameter"],
dims["outer_diameter"],
max_stress
)
elastic_modulus = Tube.get_elastic_modulus(material, ureg)
density = Tube.get_density(material, ureg)
poisson = Tube.get_poisson(material)
initial_pressure, dlf = Tube.calculate_max_initial_pressure(
dims["inner_diameter"],
dims["outer_diameter"],
initial_temperature,
gas.mole_fraction_dict(),
mechanism,
max_pressure.to("Pa"),
elastic_modulus,
density,
poisson,
use_multiprocessing=False,
return_dlf=True
)
current_results = pd.Series(dtype="object")
current_results["schedule"] = schedule
current_results["size"] = size
current_results["max initial pressure (psi)"] = \
initial_pressure.to("psi").magnitude
current_results["initial temperature (F)"] = \
initial_temperature.to("degF").magnitude
current_results["tube_temp"] = initial_temperature
current_results["max_pressure"] = max_pressure
current_results["inner_diameter"] = dims["inner_diameter"]
current_results["DLF"] = dlf
results = pd.concat(
(results, current_results.to_frame().T),
ignore_index=True
)
float_keys = [
"max initial pressure (psi)",
"initial temperature (F)",
"DLF"
]
results[float_keys] = results[float_keys].astype(float)
with warnings.catch_warnings():
warnings.simplefilter("ignore")
with pd.HDFStore(results_file, "w") as store:
store.put("data", results)
else:
with pd.HDFStore(results_file, "r") as store:
results = store.data
sns.relplot(
x="initial temperature (F)",
y="max initial pressure (psi)",
col="size",
style="schedule",
kind="line",
data=results,
aspect=0.5
)
sns.despine()
```
Here we have operating limits for our mixture over varying initial temperatures across three schedules and four pipe sizes. At NPS 3 something interesting happens: schedule 80 pipe has a lower initial pressure than schedule 40 pipe. This is caused by the dynamic load factor, as shown below.
```
sns.catplot(
x="schedule",
y="DLF",
col="size",
kind="bar",
data=results,
aspect=0.5
);
```
Let us select NPS 6 schedule 80 pipe for the main section of our detonation tube, in order to maximize the size of the viewing windows and allow for a wider range of cell sizes to be captured by our camera.
```
nps_main = "6"
schedule_main = "80"
results_main = results[
(results["size"] == nps_main) &
(results["schedule"] == schedule_main)
].copy()
results_original = results_main[
["max initial pressure (psi)",
"initial temperature (F)"]].astype(float)
results_main.drop(
["max initial pressure (psi)",
"initial temperature (F)"],
axis=1,
inplace=True
)
results_main
```
With operational limits for the main detonation section out of the way, let's consider a smaller, parallel tube with an internal fan to facilitate reactant mixing. Let's also try to use schedule 40, since it is cheaper and more readily available than schedule 80. An important thing to check here is that the mixing section can handle at least as much pressure as the main section.
```
nps_mix = "1"
schedule_mix = "40"
results_mix = results[
(results["size"] == nps_mix) &
(results["schedule"] == schedule_mix)
].drop(
["max initial pressure (psi)",
"initial temperature (F)",
"inner_diameter"],
axis=1
)
results_mix["max_main_pressure"] = results_main["max_pressure"].values
results_mix["safe"] = (results_mix["max_pressure"] >
results_mix["max_main_pressure"])
if all(results_mix["safe"]):
print("Mix tube is safe :)")
else:
print("Mix tube is unsafe :(")
results_mix
```
## Select proper flanges
Now we need to know what class of flanges each of our sections require. For this we will use the maximum system pressure, which is the pressure behind the reflected detonation wave at the maximum allowable initial pressure.
### Main section
```
results_main["flange class"] = results_main.apply(
lambda x: Flange.get_class(
x["max_pressure"],
x["tube_temp"],
material,
ureg
),
axis=1
)
results_main["flange class"].max()
```
### Mixing section
```
results_mix["flange class"] = results_mix.apply(
lambda x: Flange.get_class(
x["max_main_pressure"],
x["tube_temp"],
material,
ureg
),
axis=1
)
results_mix["flange class"].max()
```
## Determine window dimensions and bolt pattern
### Window Dimensions
Since our main tube is NPS-6 schedule 80 pipe, the inner diameter is 5.76 inches. Therefore, let's design a viewing section with a visible window height of 5.75 inches. First, let's figure out how thick a fused quartz window needs to be in order to have a safety factor of 4. Also, let's try to constrain ourselves to keeping the window thickness under 1 inch if we can.
```
window_height = quant(5.75, "in")
window_lengths = np.linspace(0.25, 7, 100)
window_thicknesses = Window.minimum_thickness(
length=window_height,
width=quant(window_lengths, "in"),
safety_factor=4,
pressure=results_main["max_pressure"].max(),
rupture_modulus=(197.9, "MPa"),
unit_registry=ureg
).to("in").magnitude
max_desired_thickness = 1 # inch
plt.plot(window_lengths, window_thicknesses, "k")
plt.fill_between(
window_lengths[window_thicknesses <= max_desired_thickness],
window_thicknesses[window_thicknesses <= max_desired_thickness],
0,
color="g",
alpha=0.25,
zorder=-1
)
plt.xlim([window_lengths.min(), window_lengths.max()])
plt.ylim([0, plt.ylim()[1]])
plt.xlabel("Window length (in)")
plt.ylabel("Window thickness (in)")
plt.title(
"Window minimum thickness vs. length\n"
"Max length {:3.2f} in at {:3.2f} in. thick".format(
window_lengths[window_thicknesses <= max_desired_thickness].max(),
max_desired_thickness
)
)
sns.despine()
```
let's go with a 2.25 inch wide window (due to camera limitations), and stick to 1 inch thick. what's the safety factor?
```
window_length = quant(2.25, "in")
Window.safety_factor(
window_length,
window_height,
quant(1, "in"),
pressure=results_main["max_pressure"].max(),
rupture_modulus=quant(197.9, "MPa"),
)
```
Nice.
### Bolt pattern
Now let's see how many 1/4-28 grade 8 bolts we need to clamp our window in place if we can tap them 1/2 inch into the viewing section plate.
```
num_bolts = np.array(range(1, 25))
bolt_safety_factors = Bolt.calculate_safety_factors(
max_pressure=results_main["max_pressure"].max(),
window_area = window_length * window_height,
num_bolts=num_bolts,
thread_size="1/4-28",
thread_class="2",
bolt_max_tensile=(150, "kpsi"), # grade 8
plate_max_tensile=(485, "MPa"), # 316L,
engagement_length=(0.5, "in"),
unit_registry=ureg
)
fig, ax = plt.subplots()
ax.plot(
num_bolts,
bolt_safety_factors["bolt"],
"k--",
label="bolt"
)
ax.plot(
num_bolts,
bolt_safety_factors["plate"],
"k-.",
label="plate"
)
ax.xaxis.set_major_locator(MaxNLocator(integer=True))
ax.legend()
ax.set_ylim([0, ax.get_ylim()[1]])
ax.set_xlim([num_bolts.min()-1, num_bolts.max()+1])
ax.set_xlabel("Number of bolts")
ax.set_ylabel("Safety factor")
ax.set_title("Bolt and plate safety factors vs. number of bolts")
ax.fill_between(
ax.get_xlim(),
2,
zorder=-1,
color="r",
alpha=0.25,
)
sns.despine()
```
For a safety factor of 2.5, we will need a minimum of 12 bolts. Another important thing to pay attention to: The bolt safety factor is less than the plate safety factor. This is desirable; bolts are much cheaper to replace than a detonation tube, particularly when the part of the tube in question is machined from a very large and expensive piece of stainless steel!
## Estimate DDT length
Now let's estimate how long it will take a deflagration ignited at one end of the tube to transition into a detonation. The `calculate_blockage_diameter` and `calculate_blockage_ratio` functions included in `DDT` are for a Shchelkin spiral. This is not the only blockage pattern that can be used, however if you want to use an arbitrary blockage you will have to handle blockage ratio calculations on your own. That being said, the `DDT.calculate_run_up` function accepts arguments of blockage ratio and tube diameter, meaning that its only geometric assumption is a circular chamber cross-section.
```
main_id = results_main["inner_diameter"].iloc[0]
target_blockage_diameter = DDT.calculate_blockage_diameter(
main_id,
0.45,
unit_registry=ureg
)
print("Target blockage diameter: {:3.2f}".format(target_blockage_diameter))
blockage_actual = DDT.calculate_blockage_ratio(
main_id,
(0.75, "in"),
unit_registry=ureg
)
print("Actual blockage ratio: {:4.1f}%".format(blockage_actual*100))
runup = DDT.calculate_run_up(
blockage_actual,
main_id,
(70, "degF"),
(1, "atm"),
gas.mole_fraction_dict(),
mechanism,
unit_registry=ureg
)
print("Runup distance: {:1.2f}".format(runup.to("millifurlong")))
```
Of course, since this package makes use of `pint`, you may use whatever ridiculous units you want for inputs and outputs.
## Find safe operation limits for a new mixture
Finally, since the dynamic load factor is a function of wave speed as well as the tube's geometry and material properties, the operational limits of your tube may change from mixture to mixture. Let's take a look at what would happen to our safe operation limits for this tube if we decided to pack it with hydrogen and oxygen instead of propane and air.
```
results_file = "second_mixture_results.h5"
calculate_results = True
new_fuel = "H2"
new_oxidizer = "O2"
gas2 = ct.Solution(mechanism)
gas2.set_equivalence_ratio(1, new_fuel, new_oxidizer)
if calculate_results:
results = pd.DataFrame(
columns=[
"max initial pressure (psi)",
"initial temperature (F)",
]
)
for i, initial_temperature in enumerate(initial_temperatures):
dims = Tube.get_dimensions(
nps_main,
schedule_main,
unit_registry=ureg
)
max_stress = Tube.calculate_max_stress(
initial_temperature,
material,
welded=False,
unit_registry=ureg
)
max_pressure = Tube.calculate_max_pressure(
dims["inner_diameter"],
dims["outer_diameter"],
max_stress
)
elastic_modulus = Tube.get_elastic_modulus(material, ureg)
density = Tube.get_density(material, ureg)
poisson = Tube.get_poisson(material)
initial_pressure = Tube.calculate_max_initial_pressure(
dims["inner_diameter"],
dims["outer_diameter"],
initial_temperature,
gas2.mole_fraction_dict(),
mechanism,
max_pressure.to("Pa"),
elastic_modulus,
density,
poisson,
use_multiprocessing=False
)
current_results = pd.Series(dtype="object")
current_results["max initial pressure (psi)"] = \
initial_pressure.to("psi").magnitude
current_results["initial temperature (F)"] = \
initial_temperature.to("degF").magnitude
results = pd.concat(
(results, current_results.to_frame().T),
ignore_index=True
)
with warnings.catch_warnings():
warnings.simplefilter("ignore")
with pd.HDFStore(results_file, "w") as store:
store.put("data", results)
else:
with pd.HDFStore(results_file, "r") as store:
results = store.data
results_original["mixture"] = "original"
results["mixture"] = "new"
sns.lineplot(
x="initial temperature (F)",
y="max initial pressure (psi)",
data=pd.concat((results_original, results)),
style="mixture",
color="k",
)
plt.title("Mixture comparison\nMax safe initial pressure vs. Temperature")
sns.despine()
```
| github_jupyter |
<center>
<img src="https://gitlab.com/ibm/skills-network/courses/placeholder101/-/raw/master/labs/module%201/images/IDSNlogo.png" width="300" alt="cognitiveclass.ai logo" />
</center>
# **Exploratory Data Analysis Lab**
Estimated time needed: **30** minutes
In this module you get to work with the cleaned dataset from the previous module.
In this assignment you will perform the task of exploratory data analysis.
You will find out the distribution of data, presence of outliers and also determine the correlation between different columns in the dataset.
## Objectives
In this lab you will perform the following:
* Identify the distribution of data in the dataset.
* Identify outliers in the dataset.
* Remove outliers from the dataset.
* Identify correlation between features in the dataset.
***
## Hands on Lab
Import the pandas module.
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from scipy.stats import norm
%matplotlib inline
```
Load the dataset into a dataframe.
```
df = pd.read_csv("m2_survey_data.csv")
df.head()
df.shape
```
## Distribution
### Determine how the data is distributed
The column `ConvertedComp` contains Salary converted to annual USD salaries using the exchange rate on 2019-02-01.
This assumes 12 working months and 50 working weeks.
Plot the distribution curve for the column `ConvertedComp`.
```
plt.figure(figsize=(10,5))
sns.distplot(a=df["ConvertedComp"],bins=20,hist=False)
plt.show()
```
Plot the histogram for the column `ConvertedComp`.
```
plt.figure(figsize=(10,5))
sns.distplot(a=df["ConvertedComp"],bins=20,kde=False)
plt.show()
```
What is the median of the column `ConvertedComp`?
```
df["ConvertedComp"].median()
```
How many responders identified themselves only as a **Man**?
```
df["Gender"].value_counts()
```
Find out the median ConvertedComp of responders identified themselves only as a **Woman**?
```
woman = df[df["Gender"] == "Woman"]
woman["ConvertedComp"].median()
```
Give the five number summary for the column `Age`?
```
df["Age"].describe()
```
**Double click here for hint**.
<!--
min,q1,median,q3,max of a column are its five number summary.
-->
Plot a histogram of the column `Age`.
```
plt.figure(figsize=(10,5))
sns.distplot(a=df["Age"],bins=20,kde=False)
plt.show()
plt.figure(figsize=(10,5))
sns.distplot(a=df["Respondent"],bins=20,kde=False)
plt.show()
```
## Outliers
### Finding outliers
Find out if outliers exist in the column `ConvertedComp` using a box plot?
```
plt.figure(figsize=(10,5))
sns.boxplot(x=df.ConvertedComp, data=df)
plt.show()
plt.figure(figsize=(10,5))
sns.boxplot(x=df.Age, data=df)
plt.show()
```
Find out the Inter Quartile Range for the column `ConvertedComp`.
```
df["ConvertedComp"].describe()
1.000000e+05 - 2.686800e+04
```
Find out the upper and lower bounds.
```
Q1 = df["ConvertedComp"].quantile(0.25)
Q3 = df["ConvertedComp"].quantile(0.75)
IQR = Q3 - Q1
print(IQR)
```
Identify how many outliers are there in the `ConvertedComp` column.
```
outliers = (df["ConvertedComp"] < (Q1 - 1.5 * IQR)) | (df["ConvertedComp"] > (Q3 + 1.5 * IQR))
outliers.value_counts()
```
Create a new dataframe by removing the outliers from the `ConvertedComp` column.
```
less = (df["ConvertedComp"] < (Q1 - 1.5 * IQR))
less.value_counts()
more = (df["ConvertedComp"] > (Q3 + 1.5 * IQR))
more.value_counts()
convertedcomp_out = df[~(df["ConvertedComp"] > (Q3 + 1.5 * IQR))]
convertedcomp_out.head()
convertedcomp_out["ConvertedComp"].median()
convertedcomp_out["ConvertedComp"].mean()
```
## Correlation
### Finding correlation
Find the correlation between `Age` and all other numerical columns.
```
df.corr()
```
## Authors
Ramesh Sannareddy
### Other Contributors
Rav Ahuja
## Change Log
| Date (YYYY-MM-DD) | Version | Changed By | Change Description |
| ----------------- | ------- | ----------------- | ---------------------------------- |
| 2020-10-17 | 0.1 | Ramesh Sannareddy | Created initial version of the lab |
Copyright © 2020 IBM Corporation. This notebook and its source code are released under the terms of the [MIT License](https://cognitiveclass.ai/mit-license?utm_medium=Exinfluencer\&utm_source=Exinfluencer\&utm_content=000026UJ\&utm_term=10006555\&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDA0321ENSkillsNetwork21426264-2021-01-01\&cm_mmc=Email_Newsletter-\_-Developer_Ed%2BTech-\_-WW_WW-\_-SkillsNetwork-Courses-IBM-DA0321EN-SkillsNetwork-21426264\&cm_mmca1=000026UJ\&cm_mmca2=10006555\&cm_mmca3=M12345678\&cvosrc=email.Newsletter.M12345678\&cvo_campaign=000026UJ).
| github_jupyter |
```
import pennylane as qml
from pennylane import numpy as np
from pennylane.templates import RandomLayers
import tensorflow as tf
from tensorflow import keras
import matplotlib.pyplot as plt
from qiskit import *
n_epochs = 30 # Number of optimization epochs
n_layers = 1 # Number of random layers
n_train = 50 # Size of the train dataset
n_test = 30 # Size of the test dataset
SAVE_PATH = "quanvolution/" # Data saving folder
PREPROCESS = True # If False, skip quantum processing and load data from SAVE_PATH
# np.random.seed(0) # Seed for NumPy random number generator
tf.random.set_seed(0) # Seed for TensorFlow random number generator
mnist_dataset = keras.datasets.mnist
(train_images, train_labels), (test_images, test_labels) = mnist_dataset.load_data()
# Reduce dataset size
train_images = train_images[:n_train]
train_labels = train_labels[:n_train]
test_images = test_images[:n_test]
test_labels = test_labels[:n_test]
# Normalize pixel values within 0 and 1
train_images = train_images / 255
test_images = test_images / 255
# Add extra dimension for convolution channels
train_images = np.array(train_images[..., tf.newaxis], requires_grad=False)
test_images = np.array(test_images[..., tf.newaxis], requires_grad=False)
# !pip3 install qiskit.ibmq
token1 = '2b740fada862fb6d9683487f84455ff8fccb2caaa3988e4f406d321ef67ce0644eb679211a1b6a377ec5ee9b3839d34434255304ab535a9ed11ce60703f07446'
token2 = '08952555d263c29c9b015855d49bcbf0e7a3cb1354b5520743285bb48d19f0db420522be05c80dd1fb19179c42d798b238ae7b944568bb6e7e7284d9bfeb0209'
from qiskit import IBMQ
# IBMQ.save_account(token2)
# provider = IBMQ.enable_account(token1)
# dev = qml.device("default.qubit", wires=9)
# dev = qml.device('qiskit.ibmq', wires=9, backend='ibmq_qasm_simulator',provider=provider)
# IBMQ.get_provider(hub='ibm-q', group='open', project='main')
dev = qml.device('qiskit.ibmq', wires=9, backend='ibmq_qasm_simulator',
ibmqx_token=token1, hub='ibm-q', group='open', project='main')
print(dev.capabilities()['backend'])
# import pennylane as qml
# from pennylane_ionq import ops
# dev = qml.device("ionq.qpu", wires=9)
# Random circuit parameters
@qml.qnode(dev)
def circuit(phi):
# Encoding of 4 classical input values
for j in range(9):
qml.RY(np.pi * phi[j], wires=j)
# rand_params = np.random.uniform(high=2 * np.pi, size=(n_layers, 9))
# print("rand", rand_params)
# Random quantum circuit
RandomLayers(np.random.uniform(high=2 * np.pi, size=(n_layers, 9)), wires=list(range(9)), seed=None)
# Measurement producing 9 classical output values
return [qml.expval(qml.PauliZ(j)) for j in range(9)]
def quanv(image,filters=9):
"""Convolves the input image with many applications of the same quantum circuit."""
out = np.zeros((26, 26, filters))
# Loop over the coordinates of the top-left pixel of 2X2 squares
for j in range(1, 27, 1):
for k in range(1, 27, 1):
# Process a squared 2x2 region of the image with a quantum circuit
q_results = []
for i in range(filters):
q_result = circuit(
[
image[j - 1, k - 1, 0],
image[j - 1, k, 0],
image[j - 1, k + 1, 0],
image[j, k - 1, 0],
image[j, k, 0],
image[j, k + 1, 0],
image[j + 1, k - 1, 0],
image[j + 1, k, 0],
image[j + 1, k + 1, 0]
]
)
# print(q_result)
q_results.append(sum(q_result))
# print(q_results,"hi")
# Assign expectation values to different channels of the output pixel (j/2, k/2)
for c in range(filters):
out[j - 1, k - 1, c] = q_results[c]
return out
if PREPROCESS == True:
q_train_images = []
print("Quantum pre-processing of train images:")
for idx, img in enumerate(train_images):
print("{}/{} ".format(idx + 1, n_train), end="\r")
q_train_images.append(quanv(img))
q_train_images = np.asarray(q_train_images)
q_test_images = []
print("\nQuantum pre-processing of test images:")
for idx, img in enumerate(test_images):
print("{}/{} ".format(idx + 1, n_test), end="\r")
q_test_images.append(quanv(img))
q_test_images = np.asarray(q_test_images)
# Save pre-processed images
np.save(SAVE_PATH + "q_train_images_f1_c9.npy", q_train_images)
np.save(SAVE_PATH + "q_test_images_f1_c9.npy", q_test_images)
# Load pre-processed images
q_train_images = np.load(SAVE_PATH + "q_train_images_f1_c9.npy")
q_test_images = np.load(SAVE_PATH + "q_test_images_f1_c9.npy")
n_samples = 4
n_channels = 9
fig, axes = plt.subplots(1 + n_channels, n_samples, figsize=(10, 10))
for k in range(n_samples):
axes[0, 0].set_ylabel("Input")
if k != 0:
axes[0, k].yaxis.set_visible(False)
axes[0, k].imshow(train_images[k, :, :, 0], cmap="gray")
# Plot all output channels
for c in range(n_channels):
axes[c + 1, 0].set_ylabel("Output [ch. {}]".format(c))
if k != 0:
axes[c, k].yaxis.set_visible(False)
axes[c + 1, k].imshow(q_train_images[k, :, :, c], cmap="gray")
plt.tight_layout()
plt.show()
def MyModel():
"""Initializes and returns a custom Keras model
which is ready to be trained."""
model = keras.models.Sequential([
keras.layers.Conv2D(filters=50,kernel_size=(5,5),activation='relu'),
tf.keras.layers.MaxPooling2D(),
keras.layers.Conv2D(filters=64,kernel_size=(5,5),activation='relu'),
tf.keras.layers.MaxPooling2D(),
keras.layers.Flatten(),
keras.layers.Dense(1024, activation="relu"),
keras.layers.Dropout(0.4),
keras.layers.Dense(10, activation="softmax")
])
model.compile(
optimizer='adam',
loss="sparse_categorical_crossentropy",
metrics=["accuracy"],
)
return model
q_model = MyModel()
q_history = q_model.fit(
q_train_images,
train_labels,
validation_data=(q_test_images, test_labels),
batch_size=4,
epochs=n_epochs,
verbose=2,
)
c_model = MyModel()
c_history = c_model.fit(
train_images,
train_labels,
validation_data=(test_images, test_labels),
batch_size=4,
epochs=n_epochs,
verbose=2,
)
import matplotlib.pyplot as plt
plt.style.use("seaborn")
fig, (ax1, ax2) = plt.subplots(2, 1, figsize=(6, 9))
ax1.plot(q_history.history["val_accuracy"], "-ob", label="With quantum layer")
ax1.plot(c_history.history["val_accuracy"], "-og", label="Without quantum layer")
ax1.set_ylabel("Accuracy")
ax1.set_ylim([0, 1])
ax1.set_xlabel("Epoch")
ax1.legend()
ax2.plot(q_history.history["val_loss"], "-ob", label="With quantum layer")
ax2.plot(c_history.history["val_loss"], "-og", label="Without quantum layer")
ax2.set_ylabel("Loss")
ax2.set_ylim(top=2.5)
ax2.set_xlabel("Epoch")
ax2.legend()
plt.tight_layout()
plt.show()
```
| github_jupyter |
```
# imports
from keras.models import Sequential
from keras.layers import Dense
from keras.layers import Dropout
from sklearn.metrics import r2_score
import matplotlib.pyplot as plt
import numpy
from keras.optimizers import Adam
import keras
from matplotlib import pyplot
from keras.callbacks import EarlyStopping
import pandas as pd
from sklearn.preprocessing import LabelEncoder
# Read data from csv file for training and validation data
TrainingSet = numpy.genfromtxt("./training.csv", delimiter=",", skip_header=True)
ValidationSet = numpy.genfromtxt("./validation.csv", delimiter=",", skip_header=True)
# Split into input (X) and output (Y) variables
X1 = TrainingSet[:,0:6]
Y1 = TrainingSet[:,6]
X2 = ValidationSet[:,0:6]
Y2 = ValidationSet[:,6]
# Create model
model = Sequential()
model.add(Dense(128, activation="relu", input_dim=6))
model.add(Dense(32, activation="relu"))
model.add(Dense(8, activation="relu"))
# Since the regression is performed, a Dense layer containing a single neuron with a linear activation function.
# Typically ReLu-based activation are used but since it is performed regression, it is needed a linear activation.
model.add(Dense(1, activation="linear"))
# Compile model: The model is initialized with the Adam optimizer and then it is compiled.
model.compile(loss='mean_squared_error', optimizer=Adam(lr=1e-3, decay=1e-3 / 200))
# Patient early stopping
es = EarlyStopping(monitor='val_loss', mode='min', verbose=1, patience=200)
# Fit the model
history = model.fit(X1, Y1, validation_data=(X2, Y2), epochs=10000000, batch_size=100, verbose=2, callbacks=[es])
# Calculate predictions
PredTestSet = model.predict(X1)
PredValSet = model.predict(X2)
# Save predictions
numpy.savetxt("trainresults.csv", PredTestSet, delimiter=",")
numpy.savetxt("valresults.csv", PredValSet, delimiter=",")
# Plot training history
pyplot.plot(history.history['loss'], label='train')
pyplot.plot(history.history['val_loss'], label='test')
pyplot.legend()
plt.title('Training History'),
plt.xlabel('Epoch'),
plt.ylabel('Validation Loss')
pyplot.show()
# Plot actual vs prediction for training set
TestResults = numpy.genfromtxt("trainresults.csv", delimiter=",")
plt.plot(Y1,TestResults,'ro')
plt.title('Training Set')
plt.xlabel('Actual')
plt.ylabel('Predicted')
# Compute R-Square value for training set
TestR2Value = r2_score(Y1,TestResults)
print("Training Set R-Square=", TestR2Value)
# Plot actual vs prediction for validation set
ValResults = numpy.genfromtxt("valresults.csv", delimiter=",")
plt.plot(Y2,ValResults,'ro')
plt.title('Validation Set')
plt.xlabel('Actual')
plt.ylabel('Predicted')
# Compute R-Square value for validation set
ValR2Value = r2_score(Y2,ValResults)
print("Validation Set R-Square=",ValR2Value)
```
| github_jupyter |
# PyGw Showcase
This notebook demonstrates the some of the utility provided by the `pygw` python package.
In this guide, we will show how you can use `pygw` to easily:
- **Define** a data schema for Geotools SimpleFeature/Vector data (aka create a new data type)
- **Create** instances for the new type
- **Create** a RocksDB GeoWave Data Store
- **Register** a DataType Adapter & Index to the data store for your new data type
- **Write** user-created data into the GeoWave Data Store
- **Query** data out of the data store
```
%pip install ../../../../python/src/main/python
```
### Loading state capitals test data set
Load state capitals from CSV
```
import csv
with open("../../../java-api/src/main/resources/stateCapitals.csv", encoding="utf-8-sig") as f:
reader = csv.reader(f)
raw_data = [row for row in reader]
# Let's take a look at what the data looks like
raw_data[0]
```
For the purposes of this exercise, we will use the state name (`[0]`), capital name (`[1]`), longitude (`[2]`), latitude (`[3]`), and the year that the capital was established (`[4]`).
### Creating a new SimpleFeatureType for the state capitals data set
We can define a data schema for our data by using a `SimpleFeatureTypeBuilder` to build a `SimpleFeatureType`.
We can use the convenience methods defined in `AttributeDescriptor` to define each field of the feature type.
```
from pygw.geotools import SimpleFeatureTypeBuilder
from pygw.geotools import AttributeDescriptor
# Create the feature type builder
type_builder = SimpleFeatureTypeBuilder()
# Set the name of the feature type
type_builder.set_name("StateCapitals")
# Add the attributes
type_builder.add(AttributeDescriptor.point("location"))
type_builder.add(AttributeDescriptor.string("state_name"))
type_builder.add(AttributeDescriptor.string("capital_name"))
type_builder.add(AttributeDescriptor.date("established"))
# Build the feature type
state_capitals_type = type_builder.build_feature_type()
```
### Creating features for each data point using our new SimpleFeatureType
`pygw` allows you to create `SimpleFeature` instances for `SimpleFeatureType` using a `SimpleFeatureBuilder`.
The `SimpleFeatureBuilder` allows us to specify all of the attributes of a feature, and then build it by providing a feature ID. For this exercise, we will use the index of the data as the unique feature id. We will use `shapely` to create the geometries for each feature.
```
from pygw.geotools import SimpleFeatureBuilder
from shapely.geometry import Point
from datetime import datetime
feature_builder = SimpleFeatureBuilder(state_capitals_type)
features = []
for idx, capital in enumerate(raw_data):
state_name = capital[0]
capital_name = capital[1]
longitude = float(capital[2])
latitude = float(capital[3])
established = datetime(int(capital[4]), 1, 1)
feature_builder.set_attr("location", Point(longitude, latitude))
feature_builder.set_attr("state_name", state_name)
feature_builder.set_attr("capital_name", capital_name)
feature_builder.set_attr("established", established)
feature = feature_builder.build(str(idx))
features.append(feature)
```
### Creating a data store
Now that we have a set of `SimpleFeatures`, let's create a data store to write the features into. `pygw` supports all of the data store types that GeoWave supports. All that is needed is to first construct the appropriate `DataStoreOptions` variant that defines the parameters of the data store, then to pass those options to a `DataStoreFactory` to construct the `DataStore`. In this example we will create a new RocksDB data store.
```
from pygw.store import DataStoreFactory
from pygw.store.rocksdb import RocksDBOptions
# Specify the options for the data store
options = RocksDBOptions()
options.set_geowave_namespace("geowave.example")
# NOTE: Directory is relative to the JVM working directory.
options.set_directory("./datastore")
# Create the data store
datastore = DataStoreFactory.create_data_store(options)
```
#### An aside: `help()`
Much of `pygw` is well-documented, and the `help` method in python can be useful for figuring out what a `pygw` instance can do. Let's try it out on our data store.
```
help(datastore)
```
### Adding our data to the data store
To store data into our data store, we first have to register a `DataTypeAdapter` for our simple feature data and create an index that defines how the data is queried. GeoWave supports simple feature data through the use of a `FeatureDataAdapter`. All that is needed for a `FeatureDataAdapter` is a `SimpleFeatureType`. We will also add both spatial and spatial/temporal indices.
```
from pygw.geotools import FeatureDataAdapter
# Create an adapter for feature type
state_capitals_adapter = FeatureDataAdapter(state_capitals_type)
from pygw.index import SpatialIndexBuilder
from pygw.index import SpatialTemporalIndexBuilder
# Add a spatial index
spatial_idx = SpatialIndexBuilder().set_name("spatial_idx").create_index()
# Add a spatial/temporal index
spatial_temporal_idx = SpatialTemporalIndexBuilder().set_name("spatial_temporal_idx").create_index()
# Now we can add our type to the data store with our spatial index
datastore.add_type(state_capitals_adapter, spatial_idx, spatial_temporal_idx)
# Check that we've successfully registered an index and type
registered_types = datastore.get_types()
for t in registered_types:
print(t.get_type_name())
registered_indices = datastore.get_indices(state_capitals_adapter.get_type_name())
for i in registered_indices:
print(i.get_name())
```
### Writing data to our store
Now our data store is ready to receive our feature data. To do this, we must create a `Writer` for our data type.
```
# Create a writer for our data
writer = datastore.create_writer(state_capitals_adapter.get_type_name())
# Writing data to the data store
for ft in features:
writer.write(ft)
# Close the writer when we are done with it
writer.close()
```
### Querying our store to make sure the data was ingested properly
`pygw` supports querying data in the same fashion as the Java API. You can use a `VectorQueryBuilder` to create queries on simple feature data sets. We will use one now to query all of the state capitals in the data store.
```
from pygw.query import VectorQueryBuilder
# Create the query builder
query_builder = VectorQueryBuilder()
# When you don't supply any constraints to the query builder, everything will be queried
query = query_builder.build()
# Execute the query
results = datastore.query(query)
```
The results returned above is a closeable iterator of `SimpleFeature` objects. Let's define a function that we can use to print out some information about these feature and then close the iterator when we are finished with it.
```
def print_results(results):
for result in results:
capital_name = result.get_attribute("capital_name")
state_name = result.get_attribute("state_name")
established = result.get_attribute("established")
print("{}, {} was established in {}".format(capital_name, state_name, established.year))
# Close the iterator
results.close()
# Print the results
print_results(results)
```
### Constraining the results
Querying all of the data can be useful occasionally, but most of the time we will want to filter the data to only return results that we are interested in. `pygw` supports several types of constraints to make querying data as flexible as possible.
#### CQL Constraints
One way you might want to query the data is using a simple CQL query.
```
# A CQL expression for capitals that are in the northeastern part of the US
cql_expression = "BBOX(location, -87.83,36.64,-66.74,48.44)"
# Create the query builder
query_builder = VectorQueryBuilder()
query_builder.add_type_name(state_capitals_adapter.get_type_name())
# If we want, we can tell the query builder to use the spatial index, since we aren't using time
query_builder.index_name(spatial_idx.get_name())
# Get the constraints factory
constraints_factory = query_builder.constraints_factory()
# Create the cql constraints
constraints = constraints_factory.cql_constraints(cql_expression)
# Set the constraints and build the query
query = query_builder.constraints(constraints).build()
# Execute the query
results = datastore.query(query)
# Display the results
print_results(results)
```
#### Spatial/Temporal Constraints
You may also want to contrain the data by both spatial and temporal constraints using the `SpatialTemporalConstraintsBuilder`. For this example, we will query all capitals that were established after 1800 within 10 degrees of Washington DC.
```
# Create the query builder
query_builder = VectorQueryBuilder()
query_builder.add_type_name(state_capitals_adapter.get_type_name())
# We can tell the builder to use the spatial/temporal index
query_builder.index_name(spatial_temporal_idx.get_name())
# Get the constraints factory
constraints_factory = query_builder.constraints_factory()
# Create the spatial/temporal constraints builder
constraints_builder = constraints_factory.spatial_temporal_constraints()
# Create the spatial constraint geometry.
washington_dc_buffer = Point(-77.035, 38.894).buffer(10.0)
# Set the spatial constraint
constraints_builder.spatial_constraints(washington_dc_buffer)
# Set the temporal constraint
constraints_builder.add_time_range(datetime(1800,1,1), datetime.now())
# Build the constraints
constraints = constraints_builder.build()
# Set the constraints and build the query
query = query_builder.constraints(constraints).build()
# Execute the query
results = datastore.query(query)
# Display the results
print_results(results)
```
#### Filter Factory Constraints
We can also use the `FilterFactory` to create more complicated filters. For example, if we wanted to find all of the capitals within 500 miles of Washington DC that contain the letter L that were established after 1830.
```
from pygw.query import FilterFactory
# Create the filter factory
filter_factory = FilterFactory()
# Create a filter that passes when the capital location is within 500 miles of the
# literal location of Washington DC
location_prop = filter_factory.property("location")
washington_dc_lit = filter_factory.literal(Point(-77.035, 38.894))
distance_km = 500 * 1.609344 # Convert miles to kilometers
distance_filter = filter_factory.dwithin(location_prop, washington_dc_lit, distance_km, "kilometers")
# Create a filter that passes when the capital name contains the letter L.
capital_name_prop = filter_factory.property("capital_name")
name_filter = filter_factory.like(capital_name_prop, "*l*")
# Create a filter that passes when the established date is after 1830
established_prop = filter_factory.property("established")
date_lit = filter_factory.literal(datetime(1830, 1, 1))
date_filter = filter_factory.after(established_prop, date_lit)
# Combine the name, distance, and date filters
combined_filter = filter_factory.and_([distance_filter, name_filter, date_filter])
# Create the query builder
query_builder = VectorQueryBuilder()
query_builder.add_type_name(state_capitals_adapter.get_type_name())
# Get the constraints factory
constraints_factory = query_builder.constraints_factory()
# Create the filter constraints
constraints = constraints_factory.filter_constraints(combined_filter)
# Set the constraints and build the query
query = query_builder.constraints(constraints).build()
# Execute the query
results = datastore.query(query)
# Display the results
print_results(results)
```
### Using Pandas with GeoWave query results
It's fairly easy to load vector features from GeoWave queries into a Pandas DataFrame. To do this, make sure pandas is installed.
```
%pip install pandas
```
Next we will import pandas and issue a query to the datastore to load into a dataframe.
```
from pandas import DataFrame
# Query everything
query = VectorQueryBuilder().build()
results = datastore.query(query)
# Load the results into a pandas dataframe
dataframe = DataFrame.from_records([feature.to_dict() for feature in results])
# Display the dataframe
dataframe
```
| github_jupyter |
```
data = """
Chisăliţă-Creţu, M.-C.: Refactoring in Object-Oriented Modeling, Todesco Publisher House, Cluj-Napoca, 2011, ISBN 978-606-595-014-6, P.234.2.Book Chapters 2.1.Chisăliţă-Creţu, M.-C.: The Multi-Objective Refactoring Set Selection Problem - A Solution Representation Analysis, book title: Advances in Computer Science and Engineering, Edited by: Matthias Schmidt, InTech Publisher House, March 2011 2011, ISBN 978-953-307-173-2, P. 441-462. 3.International Journals 3.1.Chisăliţă-Creţu M.-C.: Conceptual Modeling Evolution. A Formal Approach, MathSciNet, http://www.ams.org/mathscinet, Studia Universitatis Babeş-Bolyai, Series Informatica, Categ CNCSIS B+, XVI(1), 2011, P.62 – 83.3.2.Chisăliţă-Creţu M.-C.: An evolutionary approach for the entity refactoring setselection problem, Scopus, http://www.scopus.com/home.url, Journal ofInformation Technology Review, 2010, P.107-118.3.3.Chisăliţă-Creţu, M.-C., Vescan (Fanea), A.:The multi-objective refactoring selection problem, MathSciNet, http://www.ams.org/mathscinet, Studia Universitatis Babeş-Bolyai, Series Informatica, 2009, P.249-253.4.ISI Conferences 4.1.Chisăliţă-Creţu, M.-C., Vescan (Fanea), A.,The multiobjective refactoring selection problem, International Conference ”Knowledge Engineering: Principles and Techniques”, Presa Universitară Clujeană, www.cs.ubbcluj.ro/kept2009/, 2009, P. 291-298.4.2.Chisăliţă-Creţu, M.-C.: The Entity Refactoring Set Selection Problem - Practical Experiments for an Evolutionary Approach, The World Congress on Engineering and Computer Science (WCECS2009), October 20-22, 2009, San Francisco, USA, Newswood Limited, Editor: S. I. Ao, Craig Douglas, W. S. Grundfest, Jon Burgstone, 978-988-17012-6-8, http://www.iaeng.org/WCECS2009/ ICCSA2009.html, 2009, P. 285-290.4.3.Chisăliţă-Creţu, M.-C. A Multi-Objective Approach for Entity Refactoring Set Selection Problem, the 2nd International Conference on the Applications of Digital
Maria-Camelia Chisăliţă-Creţu 2/4 Information and Web Technologies (ICADIWT 2009), August 4-6, 2009, London, UK, Scopus, 2009, P. 100-105.5.International Conferences 5.1.Chisăliţă-Creţu, M.-C.: Refactoring Impact Formal Representation on the Internal Program Structure, The 6th International Conference on virtual Learning, October 28-29, 2011, Cluj-Napoca, România, Editura Universităţii Bucureşti, ISSN 1844 - 8933, http://c3.icvl.eu/2011, 2011, P. 500-510.5.2.Chisăliţă-Creţu, M.-C.: The refactoring plan configuration. A formal model, The 5thInternational Conference on virtual Learning, October 29-31, 2010, Târgu Mureş, România, Bucharest University Publisher House, ISSN 1844 - 8933, http://c3.icvl.eu/2010, 2010, P. 418-424.5.3.Chisăliţă-Creţu, M.-C.: The optimal refactoring selection problem - a multi-objective evolutionary approach, The 5th International Conference on virtual Learning, October 29-31, 2010, Târgu Mureş, România, Editura Universităţii Bucureşti, ISSN 1844 - 8933, http://c3.icvl.eu/2010, 2010, P. 410-417.5.4.Chisăliţă-Creţu, M.-C., Mihiş, A.-D.:A model for conceptual modeling evolution, International Conference on Applied Mathematics (ICAM 7), September 1-4, 2010, Baia-Mare, Romania, 2010, P. 100-107.5.5.Chisăliţă-Creţu, M.-C.: A refactoring impact based approach for the internal quality assessment, International Conference on Applied Mathematics (ICAM 7), September 1-4, 2010, Baia-Mare, Romania, 2010, P. 108-115.5.6.Chisăliţă-Creţu, M.-C.: Solution Representation Analysis For The EvolutionaryApproach of the Entity Refactoring Set Selection Problem, the 12th International Multiconference "Information Society" (IS 2009), October 12–16, 2009, Ljubljana, Slovenia, Informacijska družba, Editor: Marko Bohanec, Matjaž Gams, Vladislav Rajkovič, 978-961-264-010-1, Inspec, Scopus, 2009, P. 269-272.5.7.Chisăliţă-Creţu, M.-C.: First Results of an Evolutionary Approach for the Entity Refactoring Set Selection Problem, the 4th International Conference "Interdisciplinarity in Engineering" (INTER-ENG 2009), November 12-13, 2009, Târgu Mureş, România, 2009, P. 200-205.5.8.Chisăliţă-Creţu M.-C.: Search-Based Software Entity Refactoring – A New Solution Representation For The Multi-Objective Evolutionary Approach Of The Entity Set Selection Refactoring Problem, the 12th International Scientific and Professional Conference (DidMatTech 2009), September 10-11, 2009, Trnava, Slovakia, Editor: Veronika Stoffová, 2009, P. 100-103.5.9.Chisăliţă-Creţu, M.-C.: Identifying Patterns to Solve Low-Level Problems, International Conference on Competitiveness and European Integration (ICCEI 2007), October 26-27, 2007, Risoprint, 2007, P. 89-96.5.10.Chisăliţă-Creţu, M.-C.: Hidden Relations Between Code Duplication and Change Couplings, International Conference on Competitiveness and European Integration (ICCEI 2007), October 26-27, 2007, Risoprint, 2007, P. 80-88.
Maria-Camelia Chisăliţă-Creţu 3/4 5.11.Chisăliţă-Creţu, M.-C.: Applying Graph Transformation Rules for Refactoring, International Conference on Competitiveness and European Integration (ICCEI 2007), October 26-27, 2007, Risoprint, 2007, P. 73-79.5.12.Mihiş, A.-D., Chisăliţă-Creţu, M.-C., Mihăilă, C., Şerban, C.-A.:A Tool That Supports Simplifying Conditional Expressions Using Boolean Functions, International Conference of Mathematics and Informatics (ICMI45), Studii şi cercetări ştiinţifice, Editor: Mocanu Marcelina & Nimineţ Valer, 2006, P. 493-502.6.National Journals 6.1.Chisăliţă-Creţu, M.-C.: Describing low level problems as patterns and solving them via refactorings, Studii şi Cercetări Ştiinţifice, Seria Matematică, Bacău, Categ CNCSIS B+, 17, 2007, P.29 – 48.6.2.Chisăliţă-Creţu, M.-C.: Refactorizarea automată a codului sursă prin aplicarea regulilor limbajului Constraint Java , Universitatea Creştină „Dimitrie Cantemir” Bucureşti, Facultatea de Ştiinţe Economice Cluj-Napoca, „Analele Facultăţii”, Seria Ştiinţe Economice, 2007, P.192-201.6.3.Chisăliţă-Creţu, M.-C.: Modele eficiente de descriere pentru anti-şabloanele de refactorizare a codului sursă , Universitatea Creştină „Dimitrie Cantemir” Bucureşti, Facultatea de Ştiinţe Economice Cluj-Napoca, „Analele Facultăţii”, Seria Ştiinţe Economice, 2007, P.181-191.6.4.Chisăliţă-Creţu, M.-C.: Problema redundanţei în codul sursă. Definiţie, cauze,consecinţe şi soluţii , Universitatea Creştină „Dimitrie Cantemir” Bucureşti,Facultatea de Ştiinţe Economice Cluj-Napoca, „Analele Facultăţii”, Seria Ştiinţe Economice, 2006, P.183-189.6.5.Chisăliţă-Creţu, M.-C.: Efecte ale refactorizării asupra structurii interne a codului , Universitatea Creştină „Dimitrie Cantemir” Bucureşti, Facultatea de Ştiinţe Economice Cluj-Napoca, „Analele Facultăţii”, Seria Ştiinţe Economice, 2005, P.214-230.7.National Conferences 7.1.Chisăliţă-Creţu, M.-C.: Introducing Composition Strategies for the Refactoring Plan Building Problem, Symposium ”Zilele Academice Clujene”, Presa Universitară Clujeană, 2012, P. 56-63.7.2.Chisăliţă-Creţu, M.-C.: Formalizing the refactoring impact on internal program quality, Symposium ”Zilele Academice Clujene”, Presa Universitară Clujeană, 2010, P. 86-91.7.3.Chisăliţă-Creţu, M.-C.: Introducing Open-Closed Principle In Object Oriented Design Via Refactorings, Zilele Informaticii Economice Clujene, 2008, P. 104-115.7.4.Chisăliţă-Creţu, M.-C.: Describing Low-Level Problems as Patterns and Solving Them via Refactorings, Proceedings of the Symposium „Zilele Academice Clujene”, 2008, P. 75-86.7.5.Chisăliţă-Creţu, M.-C.: Describing Low Level Problems As Patterns And Solving Them Via Refactorings, Conferinţa Naţională de Matematică şi Informatică (CNMI 2007), 16-17 Noiembrie, 2007, P. 10-23.
Maria-Camelia Chisăliţă-Creţu 4/4 7.6.Chisăliţă-Creţu, M.-C.: Consecinţe asupra proiectării orientate obiect prin aplicarea refactorizărilor, folosind metrici soft, Sesiunea Ştiinţifică a Universităţii Creştine „Dimitrie Cantemir” Bucureşti, Facultatea de Ştiinţe Economice Cluj-Napoca „Probleme actuale ale gândirii, ştiinţei şi practicii economico-sociale”, Risoprint, 2007, P. 212-224.7.7.Chisăliţă-Creţu, M.-C.: Utilizarea analizei conceptelor formale în refactorizare, Sesiunea Ştiinţifică a Universităţii Creştine „Dimitrie Cantemir” Bucureşti, Facultatea de Ştiinţe Economice Cluj-Napoca „Probleme actuale ale gândirii, ştiinţei şi practicii economico-sociale”, Risoprint, 2007, P. 205-211.7.8.Chisăliţă-Creţu, M.-C., Şerban, C.-A.:Impact on Design Quality of Refactorings on Code via Metrics, Proceedings of the Symposium „Zilele Academice Clujene”, 2006, P. 39-44.7.9.Chisăliţă-Creţu, M.-C., Cheia publică şi aplicaţiile ei, Sesiunea Ştiinţifică a Universităţii Creştine „Dimitrie Cantemir” Bucureşti, Facultatea de Ştiinţe Economice Cluj-Napoca „Probleme actuale ale gândirii, ştiinţei şi practicii economico-sociale”, Risoprint, 2006, P. 258-268.7.10.Chisăliţă-Creţu, M.-C.: Program Internal Structure View with Formal Concept Analysis, Proceedings of the Symposium „Zilele Academice Clujene”, 2006, P. 33-38.7.11.Chisăliţă-Creţu, M.-C.: Direcţii de cercetare în aplicarea refactorizării, Sesiunea Ştiinţifică a Universităţii Creştine „Dimitrie Cantemir” Bucureşti, Facultatea de Ştiinţe Economice Cluj-Napoca „Probleme actuale ale gândirii, ştiinţei şi practicii economico-sociale”, Risoprint, 2005, P. 287-292.7.12.Chisăliţă-Creţu, M.-C.: General Aspects of Refactoring Applicability to Conceptual Models, Proceedings of the Symposium „Colocviul Academic Clujean de INFORMATICĂ”, 2005, P. 99-104.7.13.Chisăliţă-Creţu, M.-C.: Current Problems In Refactoring Activities, Proceedings of the Symposium „Zilele Academice Clujene”, 2004, P. 27-32.7.14.Chisăliţă-Creţu, M.-C.: Rolul refactorizării în procesul de dezvoltare a produselor soft, Sesiunea Ştiinţifică a Universităţii Creştine „Dimitrie Cantemir” Bucureşti, Facultatea de Ştiinţe Economice Cluj-Napoca „Probleme actuale ale gândirii, ştiinţei şi practicii economico-sociale”, Ecoexpert, 2004, P. 429-432.7.15.Chisăliţă-Creţu, M.-C., Mihiş, A.-D., Guran, A.-M.:3D Modeling vs. 2D Modeling, Proceedings of the Symposium „Colocviul Academic Clujean de INFORMATICĂ, 2003, P. 59-64.7.16.Barticel, S., Bretan, H., Costea, C., Chisăliţă-Creţu, M.-C., Mara, A., Măgeruşan, C., Mărcuş, G., Mihiş, A.-D., Mureşan, R., Petraşcu, D., Pitiş, A., Tompa, A.:An Application for Higher Education Admission, Proceedings of the Symposium „Zilele Academice Clujene, 2002, P. 163-170.7.17.Barticel, S., Chisăliţă-Creţu, M.-C., Mara, A., Măgeruşan, C., Marcuş, G., Mihiş, A.-D., Petraşcu, D., Tompa, A.:A Documentation Server, Research Seminars, Seminar on Computer Science, 2001, P. 65-74
"""
import re
class HelperMethods:
@staticmethod
def IsDate(text):
# print("text")
# print(text)
for c in text.lstrip():
if c not in "1234567890 ":
return False
return True
def ProcessLine(line):
authors = ""
title = ""
if len(line.split()) < 5:
return False, authors, title
#re.search("", line)
authors = line.split(':')[0]
title = line.split(':')[1].split(',')[0]
date = [date for date in line.split(',') if HelperMethods.IsDate(date.lstrip())][0]
return True, authors, title, date
def ProcessLineB(line):
return False, "", ""
pubs = []
for i in re.split("[0-9]{1}\.", data):
print(i)
try:
rv, authors, title, date = ProcessLine(i)
except:
rv, authors, title = ProcessLineB(i)
date = ""
if rv:
print("authors: ", authors.lstrip())
print("title: ", title)
print("date: ", date)
pubs.append((authors.lstrip(), title, date))
for pub in pubs:
print(pub)
import mariadb
import json
with open('../credentials.json', 'r') as crd_json_fd:
json_text = crd_json_fd.read()
json_obj = json.loads(json_text)
credentials = json_obj["Credentials"]
username = credentials["username"]
password = credentials["password"]
table_name = "publications_cache"
db_name = "ubbcluj"
mariadb_connection = mariadb.connect(user=username, password=password, database=db_name)
mariadb_cursor = mariadb_connection.cursor()
for paper in pubs:
title = ""
pub_date = ""
affiliations = ""
authors = ""
try:
pub_date = paper[2].lstrip()
pub_date = str(pub_date) + "-01-01"
if len(pub_date) != 10:
pub_date = ""
except:
pass
try:
affiliations = paper[2].lstrip().split('\'')[0]
except:
pass
try:
title = paper[1].lstrip().split('\'')[0]
except:
pass
try:
authors = paper[0].lstrip().split('\'')[0]
except AttributeError:
pass
table_name = "publications_cache"
insert_string = "INSERT INTO {0} SET ".format(table_name)
insert_string += "Title=\'{0}\', ".format(title)
insert_string += "ProfessorId=\'{0}\', ".format(21)
if pub_date != "":
insert_string += "PublicationDate=\'{0}\', ".format(str(pub_date))
insert_string += "Authors=\'{0}\', ".format(authors)
insert_string += "Affiliations=\'{0}\' ".format("")
print(insert_string)
#print(pub_date)
try:
mariadb_cursor.execute(insert_string)
except mariadb.ProgrammingError as pe:
print("Error")
raise pe
except mariadb.IntegrityError:
continue
```
| github_jupyter |
```
# Copyright 2020 The Google Research Authors.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
'''
# Introduction
This is a demo of the SOFT top-k operator (https://arxiv.org/pdf/2002.06504.pdf).
We demostrate the usage of the provided `Topk_custom` module in the forward and the backward pass.
'''
```
'''
# Set up
'''
import torch
from torch.nn.parameter import Parameter
import numpy as np
import soft_ot as soft
import matplotlib.pyplot as plt
import matplotlib
matplotlib.rc('xtick', labelsize=20)
matplotlib.rc('ytick', labelsize=20)
torch.manual_seed(1)
num_iter = int(1e2)
k = 3
epsilon=5e-2 # larger epsilon lead to smoother relaxation, and requires less num_iter
soft_topk = soft.TopK_custom(k, epsilon=epsilon, max_iter=num_iter)
'''
# Input the scores
'''
scores = [5,2,3,4,1,6] #input the scores here
scores_tensor = Parameter(torch.FloatTensor([scores]))
print('======scores======')
print(scores)
'''
# Forward pass.
The goal of the forward pass is to identify the scores that belongs to top-k.
The `soft_topk` object returns a smoothed indicator function: The entries are close to 1 for top-k scores, and close to 0 for non-top-k scores.
The smoothness is controled by hyper-parameter `epsilon`.
'''
A = soft_topk(scores_tensor)
indicator_vector = A.data.numpy()
print('======topk scores======')
print(indicator_vector[0,:])
plt.imshow(indicator_vector, cmap='Greys')
# plt.axis('off')
plt.yticks([])
plt.xticks(range(len(scores)), scores)
plt.colorbar()
plt.show()
'''
# Backward Pass
The goal of training is to push the scores that should have been top-k to really be top-k.
For example, in neural kNN, we want to push the scores with the same labels as the query sample to be top-k.
In this demo, we mimick the loss function of neural kNN.
`picked` is the scores ids with the same label as the query sample. Our training goal is to push them to be top-k.
'''
picked = [1,2,3]
loss = 0
for item in picked:
loss += A[0,item]
loss.backward()
A_grad = scores_tensor.grad.clone()
print('======w.r.t score grad======')
print(A_grad.data.numpy())
'''
# Visualization of the Grad
'''
x = scores
grad = A_grad.numpy()[0,:]
grad = grad/np.linalg.norm(grad)
plt.figure(figsize=(len(scores),5))
plt.scatter(range(len(x)), x)
picked_scores = [x[item] for item in picked]
plt.scatter(picked, picked_scores, label='scores we want to push to smallest top-k')
for i, item in enumerate(x):
plt.arrow(i, item, 0, grad[i], head_width=abs(grad[i])/4, fc='k')
plt.xticks(range(len(x)), x)
plt.yticks([])
plt.xlim([-0.5, len(scores)-0.5])
plt.ylim([min(scores)-1, max(scores)+1])
plt.legend()
plt.show()
# clear the grad before rerun the forward-backward code
scores_tensor.grad.data.zero_()
```
| github_jupyter |
# Missing Position Prediction - GRU Context Result
```
%load_ext autoreload
%autoreload 2
%matplotlib inline
from sentencebert_missingpositionprediction import *
import numpy as np
import seaborn as sns
import matplotlib as mpl
import matplotlib.pyplot as plt
import pandas as pd
%matplotlib inline
device = torch.device('cuda')
```
## Dataset
```
# train
train_dataset = ROCStoriesDataset_random_missing(data_path = "../data/rocstories_completion_train.csv")
# dev
val_dataset = ROCStoriesDataset_with_missing(data_path = "../data/rocstories_completion_dev.csv")
# test
test_dataset = ROCStoriesDataset_with_missing(data_path = "../data/rocstories_completion_test.csv")
```
## Model
```
%%capture
batch_size = 32
block_size = 32
sentbertmodel = SentenceTransformer('bert-base-nli-mean-tokens')
tokenizer = BertTokenizer.from_pretrained('bert-base-cased')
# --- model ---
model = MissingPisitionPredictionModel(sentbertmodel, device).to(device)
# --- DataLoader ---|
collate_fn = lambda data: collate(data, tokenizer, block_size=block_size, device=device)
train_iterator = DataLoader(
train_dataset, sampler=RandomSampler(train_dataset), batch_size=batch_size, collate_fn=collate_fn,
)
valid_iterator = DataLoader(
val_dataset, sampler=SequentialSampler(val_dataset), batch_size=batch_size, collate_fn=collate_fn,
)
test_iterator = DataLoader(
test_dataset, sampler=SequentialSampler(test_dataset), batch_size=batch_size, collate_fn=collate_fn,
)
TRG_PAD_IDX = tokenizer.pad_token_id
START_ID = tokenizer.cls_token_id
mpe_criterion = nn.CrossEntropyLoss()
```
### load model
```
model_dir = "<PATH TO model_dir>"
model_name = "sentbert-positionestimation_model.pt"
model_path = os.path.join(model_dir, model_name)
model.load_state_dict(torch.load(model_path))
```
## Show Result
```
batch = iter(valid_iterator).next()
story = batch.src
cls = batch.missing_ids
original_sentence = batch.tgt_str
story[0]
cls[0].to("cpu").numpy()
" ".join(np.insert(story[0], cls[0].to("cpu").numpy(), "____________________."))
out = show_result(model, test_iterator)
out.to_csv("result/mpp_gru_predict_result_20200421.csv", index=False)
```
## Heat map
```
acc_heatmap, cls_count = for_heatmap(model, test_iterator)
acc_heatmap_df = pd.DataFrame(acc_heatmap,
index=['1', '2', '3', '4', '5'], columns=['1', '2', '3', '4', '5'])
sns.set()
fig = plt.figure(dpi=1200)
ax = fig.add_subplot(1, 1, 1)
sns.heatmap(acc_heatmap_df, cmap="Blues",
annot=True, fmt=".3f",
linewidths=.5,
square=True,
vmin=0.0, vmax=1.0,
ax=ax)
ax.set(xlabel ='Ground Truth Missing Position',ylabel='Predicted Missing Position')
#plt.title('Training Epochs : 30')
#plt.title('Ours')
plt.title('GRU Context')
#plt.savefig('images/mpp_heatmap.png')
#plt.savefig('images/mpp_heatmap_with_title.png')
#plt.savefig('images/mpp_heatmap_ours.png')
plt.savefig('images/mpp_heatmap_gru.eps')
plt.show()
acc_heatmap_df.to_csv("./result/mpp_gru_acc_heatmap.csv")
```
| github_jupyter |
# 1. CNN with Pattern images based on Lengyel-Epstein model
## 1) Import Packages
```
import warnings
warnings.simplefilter(action='ignore', category=FutureWarning)
import sys
import tensorflow as tf
import keras
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten
from keras.layers.convolutional import Conv2D, MaxPooling2D
from PIL import Image
import numpy as np
import matplotlib.pyplot as plt
import random
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import OneHotEncoder
from tensorflow.python.framework import ops
```
## 2) Make Dataset
```
# Make dataset (144)
x_orig = []
y_orig = np.zeros((1,48))
for i in range(1,145):
if i <= 48 :
folder = 0
elif i <=96 :
folder = 1
else:
folder = 2
img = Image.open('144/{0}/pattern_{1}.jpg'.format(folder,i))
data = np.array(img)
x_orig.append(data)
for i in range(1,3):
y_orig = np.append(y_orig, np.full((1, 48),i), axis = 1)
# Make dataset (360)
x_orig = []
y_orig = np.zeros((1,120))
for i in range(1,361):
if i <= 120 :
folder = 0
elif i <=240 :
folder = 1
else:
folder = 2
img = Image.open('360/{0}/pattern_{1}.jpg'.format(folder,i))
data = np.array(img)
x_orig.append(data)
for i in range(1,3):
y_orig = np.append(y_orig, np.full((1, 120),i), axis = 1)
# Make dataset (720)
x_orig = []
y_orig = np.zeros((1,240))
for i in range(1,721):
if i <= 240 :
folder = 0
elif i <=480 :
folder = 1
else:
folder = 2
img = Image.open('720/{0}/pattern_{1}.jpg'.format(folder,i))
data = np.array(img)
x_orig.append(data)
for i in range(1,3):
y_orig = np.append(y_orig, np.full((1, 240),i), axis = 1)
x_orig = np.array(x_orig)
print(x_orig.shape)
print(y_orig.shape)
# Random shuffle
s = np.arange(x_orig.shape[0])
np.random.shuffle(s)
x_shuffle = x_orig[s,:]
y_shuffle = y_orig[:,s]
print(x_shuffle.shape)
print(y_shuffle.shape)
# y_shuffle
# Split train and test datasets
x_train_orig, x_test_orig, y_train_orig, y_test_orig = train_test_split(x_shuffle,y_shuffle.T, test_size=0.3, shuffle=True, random_state=1004)
# Normalize image vectors
x_train = x_train_orig/255.
x_test = x_test_orig/255.
# Convert train and test labels to one hot matrices
enc = OneHotEncoder()
y1 = y_train_orig.reshape(-1,1)
enc.fit(y1)
y_train = enc.transform(y1).toarray()
y2 = y_test_orig.reshape(-1,1)
enc.fit(y2)
y_test = enc.transform(y2).toarray()
# Explore your dataset
print ("number of training examples = " + str(x_train.shape[1]))
print ("number of test examples = " + str(x_test.shape[1]))
print ("x_train shape: " + str(x_train.shape))
print ("y_train shape: " + str(y_train.shape))
print ("x_test shape: " + str(x_test.shape))
print ("y_test shape: " + str(y_test.shape))
```
## 3) CNN(Convolutional Neural Network)
```
x_train = x_train.reshape(x_train.shape[0], 64, 64, 1)
x_test = x_test.reshape(x_test.shape[0], 64, 64, 1)
batch_size = 128
num_classes = 3
epochs = 100
model = Sequential()
model.add(Conv2D(32, kernel_size=(5, 5), strides=(1, 1), padding='same',
activation='relu',
input_shape=(64,64,1)))
model.add(MaxPooling2D(pool_size=(2, 2), strides=(2, 2)))
model.add(Conv2D(64, (2, 2), activation='relu', padding='same'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.25))
model.add(Flatten())
model.add(Dense(1000, activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(num_classes, activation='softmax'))
model.summary()
model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
hist = model.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
verbose=1,
validation_data=(x_test, y_test))
```
## 4) Accuracy analysis
```
score = model.evaluate(x_test, y_test, verbose=0)
print('Test loss:', score[0])
print('Test accuracy:', score[1])
predictions = model.predict(x_test)
predictions[10]
print(np.argmax(predictions[10])) # highest confidence
# This model is convinced that this image is "2"
plt.imshow(x_test[10].reshape(64,64))
```
| github_jupyter |
# Introduction to CNN Keras - Acc 0.997 (top 8%)
### **Yassine Ghouzam, PhD**
#### 18/07/2017
* **1. Introduction**
* **2. Data preparation**
* 2.1 Load data
* 2.2 Check for null and missing values
* 2.3 Normalization
* 2.4 Reshape
* 2.5 Label encoding
* 2.6 Split training and valdiation set
* **3. CNN**
* 3.1 Define the model
* 3.2 Set the optimizer and annealer
* 3.3 Data augmentation
* **4. Evaluate the model**
* 4.1 Training and validation curves
* 4.2 Confusion matrix
* **5. Prediction and submition**
* 5.1 Predict and Submit results
# 1. Introduction
This is a 5 layers Sequential Convolutional Neural Network for digits recognition trained on MNIST dataset. I choosed to build it with keras API (Tensorflow backend) which is very intuitive. Firstly, I will prepare the data (handwritten digits images) then i will focus on the CNN modeling and evaluation.
I achieved 99.671% of accuracy with this CNN trained in 2h30 on a single CPU (i5 2500k). For those who have a >= 3.0 GPU capabilites (from GTX 650 - to recent GPUs), you can use tensorflow-gpu with keras. Computation will be much much faster !!!
**For computational reasons, i set the number of steps (epochs) to 2, if you want to achieve 99+% of accuracy set it to 30.**
This Notebook follows three main parts:
* The data preparation
* The CNN modeling and evaluation
* The results prediction and submission
<img src="http://img1.imagilive.com/0717/mnist-sample.png" ></img>
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import seaborn as sns
%matplotlib inline
np.random.seed(2)
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix
import itertools
from keras.utils.np_utils import to_categorical # convert to one-hot-encoding
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten, Conv2D, MaxPool2D
from keras.optimizers import RMSprop
from keras.preprocessing.image import ImageDataGenerator
from keras.callbacks import ReduceLROnPlateau
sns.set(style='white', context='notebook', palette='deep')
```
# 2. Data preparation
## 2.1 Load data
```
# Load the data
train = pd.read_csv("../input/train.csv")
test = pd.read_csv("../input/test.csv")
Y_train = train["label"]
# Drop 'label' column
X_train = train.drop(labels = ["label"],axis = 1)
# free some space
del train
g = sns.countplot(Y_train)
Y_train.value_counts()
```
We have similar counts for the 10 digits.
## 2.2 Check for null and missing values
```
# Check the data
X_train.isnull().any().describe()
test.isnull().any().describe()
```
I check for corrupted images (missing values inside).
There is no missing values in the train and test dataset. So we can safely go ahead.
## 2.3 Normalization
We perform a grayscale normalization to reduce the effect of illumination's differences.
Moreover the CNN converg faster on [0..1] data than on [0..255].
```
# Normalize the data
X_train = X_train / 255.0
test = test / 255.0
```
## 2.3 Reshape
```
# Reshape image in 3 dimensions (height = 28px, width = 28px , canal = 1)
X_train = X_train.values.reshape(-1,28,28,1)
test = test.values.reshape(-1,28,28,1)
```
Train and test images (28px x 28px) has been stock into pandas.Dataframe as 1D vectors of 784 values. We reshape all data to 28x28x1 3D matrices.
Keras requires an extra dimension in the end which correspond to channels. MNIST images are gray scaled so it use only one channel. For RGB images, there is 3 channels, we would have reshaped 784px vectors to 28x28x3 3D matrices.
## 2.5 Label encoding
```
# Encode labels to one hot vectors (ex : 2 -> [0,0,1,0,0,0,0,0,0,0])
Y_train = to_categorical(Y_train, num_classes = 10)
```
Labels are 10 digits numbers from 0 to 9. We need to encode these lables to one hot vectors (ex : 2 -> [0,0,1,0,0,0,0,0,0,0]).
## 2.6 Split training and valdiation set
```
# Set the random seed
random_seed = 2
# Split the train and the validation set for the fitting
X_train, X_val, Y_train, Y_val = train_test_split(X_train, Y_train, test_size = 0.1, random_state=random_seed)
```
I choosed to split the train set in two parts : a small fraction (10%) became the validation set which the model is evaluated and the rest (90%) is used to train the model.
Since we have 42 000 training images of balanced labels (see 2.1 Load data), a random split of the train set doesn't cause some labels to be over represented in the validation set. Be carefull with some unbalanced dataset a simple random split could cause inaccurate evaluation during the validation.
To avoid that, you could use stratify = True option in train_test_split function (**Only for >=0.17 sklearn versions**).
We can get a better sense for one of these examples by visualising the image and looking at the label.
```
# Some examples
g = plt.imshow(X_train[0][:,:,0])
```
# 3. CNN
## 3.1 Define the model
I used the Keras Sequential API, where you have just to add one layer at a time, starting from the input.
The first is the convolutional (Conv2D) layer. It is like a set of learnable filters. I choosed to set 32 filters for the two firsts conv2D layers and 64 filters for the two last ones. Each filter transforms a part of the image (defined by the kernel size) using the kernel filter. The kernel filter matrix is applied on the whole image. Filters can be seen as a transformation of the image.
The CNN can isolate features that are useful everywhere from these transformed images (feature maps).
The second important layer in CNN is the pooling (MaxPool2D) layer. This layer simply acts as a downsampling filter. It looks at the 2 neighboring pixels and picks the maximal value. These are used to reduce computational cost, and to some extent also reduce overfitting. We have to choose the pooling size (i.e the area size pooled each time) more the pooling dimension is high, more the downsampling is important.
Combining convolutional and pooling layers, CNN are able to combine local features and learn more global features of the image.
Dropout is a regularization method, where a proportion of nodes in the layer are randomly ignored (setting their wieghts to zero) for each training sample. This drops randomly a propotion of the network and forces the network to learn features in a distributed way. This technique also improves generalization and reduces the overfitting.
'relu' is the rectifier (activation function max(0,x). The rectifier activation function is used to add non linearity to the network.
The Flatten layer is use to convert the final feature maps into a one single 1D vector. This flattening step is needed so that you can make use of fully connected layers after some convolutional/maxpool layers. It combines all the found local features of the previous convolutional layers.
In the end i used the features in two fully-connected (Dense) layers which is just artificial an neural networks (ANN) classifier. In the last layer(Dense(10,activation="softmax")) the net outputs distribution of probability of each class.
```
# Set the CNN model
# my CNN architechture is In -> [[Conv2D->relu]*2 -> MaxPool2D -> Dropout]*2 -> Flatten -> Dense -> Dropout -> Out
model = Sequential()
model.add(Conv2D(filters = 32, kernel_size = (5,5),padding = 'Same',
activation ='relu', input_shape = (28,28,1)))
model.add(Conv2D(filters = 32, kernel_size = (5,5),padding = 'Same',
activation ='relu'))
model.add(MaxPool2D(pool_size=(2,2)))
model.add(Dropout(0.25))
model.add(Conv2D(filters = 64, kernel_size = (3,3),padding = 'Same',
activation ='relu'))
model.add(Conv2D(filters = 64, kernel_size = (3,3),padding = 'Same',
activation ='relu'))
model.add(MaxPool2D(pool_size=(2,2), strides=(2,2)))
model.add(Dropout(0.25))
model.add(Flatten())
model.add(Dense(256, activation = "relu"))
model.add(Dropout(0.5))
model.add(Dense(10, activation = "softmax"))
```
## 3.2 Set the optimizer and annealer
Once our layers are added to the model, we need to set up a score function, a loss function and an optimisation algorithm.
We define the loss function to measure how poorly our model performs on images with known labels. It is the error rate between the oberved labels and the predicted ones. We use a specific form for categorical classifications (>2 classes) called the "categorical_crossentropy".
The most important function is the optimizer. This function will iteratively improve parameters (filters kernel values, weights and bias of neurons ...) in order to minimise the loss.
I choosed RMSprop (with default values), it is a very effective optimizer. The RMSProp update adjusts the Adagrad method in a very simple way in an attempt to reduce its aggressive, monotonically decreasing learning rate.
We could also have used Stochastic Gradient Descent ('sgd') optimizer, but it is slower than RMSprop.
The metric function "accuracy" is used is to evaluate the performance our model.
This metric function is similar to the loss function, except that the results from the metric evaluation are not used when training the model (only for evaluation).
```
# Define the optimizer
optimizer = RMSprop(lr=0.001, rho=0.9, epsilon=1e-08, decay=0.0)
# Compile the model
model.compile(optimizer = optimizer , loss = "categorical_crossentropy", metrics=["accuracy"])
```
<img src="http://img1.imagilive.com/0717/learningrates.jpg"> </img>
In order to make the optimizer converge faster and closest to the global minimum of the loss function, i used an annealing method of the learning rate (LR).
The LR is the step by which the optimizer walks through the 'loss landscape'. The higher LR, the bigger are the steps and the quicker is the convergence. However the sampling is very poor with an high LR and the optimizer could probably fall into a local minima.
Its better to have a decreasing learning rate during the training to reach efficiently the global minimum of the loss function.
To keep the advantage of the fast computation time with a high LR, i decreased the LR dynamically every X steps (epochs) depending if it is necessary (when accuracy is not improved).
With the ReduceLROnPlateau function from Keras.callbacks, i choose to reduce the LR by half if the accuracy is not improved after 3 epochs.
```
# Set a learning rate annealer
learning_rate_reduction = ReduceLROnPlateau(monitor='val_acc',
patience=3,
verbose=1,
factor=0.5,
min_lr=0.00001)
epochs = 1 # Turn epochs to 30 to get 0.9967 accuracy
batch_size = 86
```
## 3.3 Data augmentation
In order to avoid overfitting problem, we need to expand artificially our handwritten digit dataset. We can make your existing dataset even larger. The idea is to alter the training data with small transformations to reproduce the variations occuring when someone is writing a digit.
For example, the number is not centered
The scale is not the same (some who write with big/small numbers)
The image is rotated...
Approaches that alter the training data in ways that change the array representation while keeping the label the same are known as data augmentation techniques. Some popular augmentations people use are grayscales, horizontal flips, vertical flips, random crops, color jitters, translations, rotations, and much more.
By applying just a couple of these transformations to our training data, we can easily double or triple the number of training examples and create a very robust model.
The improvement is important :
- Without data augmentation i obtained an accuracy of 98.114%
- With data augmentation i achieved 99.67% of accuracy
```
# Without data augmentation i obtained an accuracy of 0.98114
#history = model.fit(X_train, Y_train, batch_size = batch_size, epochs = epochs,
# validation_data = (X_val, Y_val), verbose = 2)
# With data augmentation to prevent overfitting (accuracy 0.99286)
datagen = ImageDataGenerator(
featurewise_center=False, # set input mean to 0 over the dataset
samplewise_center=False, # set each sample mean to 0
featurewise_std_normalization=False, # divide inputs by std of the dataset
samplewise_std_normalization=False, # divide each input by its std
zca_whitening=False, # apply ZCA whitening
rotation_range=10, # randomly rotate images in the range (degrees, 0 to 180)
zoom_range = 0.1, # Randomly zoom image
width_shift_range=0.1, # randomly shift images horizontally (fraction of total width)
height_shift_range=0.1, # randomly shift images vertically (fraction of total height)
horizontal_flip=False, # randomly flip images
vertical_flip=False) # randomly flip images
datagen.fit(X_train)
```
For the data augmentation, i choosed to :
- Randomly rotate some training images by 10 degrees
- Randomly Zoom by 10% some training images
- Randomly shift images horizontally by 10% of the width
- Randomly shift images vertically by 10% of the height
I did not apply a vertical_flip nor horizontal_flip since it could have lead to misclassify symetrical numbers such as 6 and 9.
Once our model is ready, we fit the training dataset .
```
# Fit the model
history = model.fit_generator(datagen.flow(X_train,Y_train, batch_size=batch_size),
epochs = epochs, validation_data = (X_val,Y_val),
verbose = 2, steps_per_epoch=X_train.shape[0] // batch_size
, callbacks=[learning_rate_reduction])
```
# 4. Evaluate the model
## 4.1 Training and validation curves
```
# Plot the loss and accuracy curves for training and validation
fig, ax = plt.subplots(2,1)
ax[0].plot(history.history['loss'], color='b', label="Training loss")
ax[0].plot(history.history['val_loss'], color='r', label="validation loss",axes =ax[0])
legend = ax[0].legend(loc='best', shadow=True)
ax[1].plot(history.history['acc'], color='b', label="Training accuracy")
ax[1].plot(history.history['val_acc'], color='r',label="Validation accuracy")
legend = ax[1].legend(loc='best', shadow=True)
```
The code below is for plotting loss and accuracy curves for training and validation. Since, i set epochs = 2 on this notebook .
I'll show you the training and validation curves i obtained from the model i build with 30 epochs (2h30)
<img src="http://img1.imagilive.com/0717/mnist_099671_train_val_loss_acc.png"></img>
The model reaches almost 99% (98.7+%) accuracy on the validation dataset after 2 epochs. The validation accuracy is greater than the training accuracy almost evry time during the training. That means that our model dosen't not overfit the training set.
Our model is very well trained !!!
<img src="http://img1.imagilive.com/0717/accuracies1de.jpg"/>
## 4.2 Confusion matrix
Confusion matrix can be very helpfull to see your model drawbacks.
I plot the confusion matrix of the validation results.
```
# Look at confusion matrix
def plot_confusion_matrix(cm, classes,
normalize=False,
title='Confusion matrix',
cmap=plt.cm.Blues):
"""
This function prints and plots the confusion matrix.
Normalization can be applied by setting `normalize=True`.
"""
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, cm[i, j],
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
# Predict the values from the validation dataset
Y_pred = model.predict(X_val)
# Convert predictions classes to one hot vectors
Y_pred_classes = np.argmax(Y_pred,axis = 1)
# Convert validation observations to one hot vectors
Y_true = np.argmax(Y_val,axis = 1)
# compute the confusion matrix
confusion_mtx = confusion_matrix(Y_true, Y_pred_classes)
# plot the confusion matrix
plot_confusion_matrix(confusion_mtx, classes = range(10))
```
Here we can see that our CNN performs very well on all digits with few errors considering the size of the validation set (4 200 images).
However, it seems that our CNN has some little troubles with the 4 digits, hey are misclassified as 9. Sometime it is very difficult to catch the difference between 4 and 9 when curves are smooth.
Let's investigate for errors.
I want to see the most important errors . For that purpose i need to get the difference between the probabilities of real value and the predicted ones in the results.
```
# Display some error results
# Errors are difference between predicted labels and true labels
errors = (Y_pred_classes - Y_true != 0)
Y_pred_classes_errors = Y_pred_classes[errors]
Y_pred_errors = Y_pred[errors]
Y_true_errors = Y_true[errors]
X_val_errors = X_val[errors]
def display_errors(errors_index,img_errors,pred_errors, obs_errors):
""" This function shows 6 images with their predicted and real labels"""
n = 0
nrows = 2
ncols = 3
fig, ax = plt.subplots(nrows,ncols,sharex=True,sharey=True)
for row in range(nrows):
for col in range(ncols):
error = errors_index[n]
ax[row,col].imshow((img_errors[error]).reshape((28,28)))
ax[row,col].set_title("Predicted label :{}\nTrue label :{}".format(pred_errors[error],obs_errors[error]))
n += 1
# Probabilities of the wrong predicted numbers
Y_pred_errors_prob = np.max(Y_pred_errors,axis = 1)
# Predicted probabilities of the true values in the error set
true_prob_errors = np.diagonal(np.take(Y_pred_errors, Y_true_errors, axis=1))
# Difference between the probability of the predicted label and the true label
delta_pred_true_errors = Y_pred_errors_prob - true_prob_errors
# Sorted list of the delta prob errors
sorted_dela_errors = np.argsort(delta_pred_true_errors)
# Top 6 errors
most_important_errors = sorted_dela_errors[-6:]
# Show the top 6 errors
display_errors(most_important_errors, X_val_errors, Y_pred_classes_errors, Y_true_errors)
```
The most important errors are also the most intrigous.
For those six case, the model is not ridiculous. Some of these errors can also be made by humans, especially for one the 9 that is very close to a 4. The last 9 is also very misleading, it seems for me that is a 0.
```
# predict results
results = model.predict(test)
# select the indix with the maximum probability
results = np.argmax(results,axis = 1)
results = pd.Series(results,name="Label")
submission = pd.concat([pd.Series(range(1,28001),name = "ImageId"),results],axis = 1)
submission.to_csv("cnn_mnist_datagen.csv",index=False)
```
** you found this notebook helpful or you just liked it , some upvotes would be very much appreciated - That will keep me motivated :)**
| github_jupyter |
```
%%time
import malaya
string1 = 'xjdi ke, y u xsuke makan HUSEIN kt situ tmpt, i hate it. pelikle, pada'
string2 = 'i mmg xske mknn HUSEIN kampng tmpat, i love them. pelikle saye'
string3 = 'perdana menteri ke11 sgt suka makn ayam, harganya cuma rm15.50'
string4 = 'pada 10/4, kementerian mengumumkan, 1/100'
string5 = 'Husein Zolkepli dapat tempat ke-12 lumba lari hari ni'
string6 = 'Husein Zolkepli (2011 - 2019) adalah ketua kampng di kedah sekolah King Edward ke-IV'
```
## Load spell normalizer
```
corrector = malaya.spell.probability()
normalizer = malaya.normalize.spell(corrector)
normalizer.normalize('boleh dtg 8pagi esok tak atau minggu depan? 2 oktober 2019 2pm, tlong bayar rm 3.2k sekali tau')
```
Here you can see, Malaya normalizer will normalize `minggu depan` to datetime object, also `3.2k ringgit` to `RM3200`
```
print(normalizer.normalize(string1))
print(normalizer.normalize(string2))
print(normalizer.normalize(string3))
print(normalizer.normalize(string4))
print(normalizer.normalize(string5))
print(normalizer.normalize(string6))
```
## Normalizing rules
#### 1. Normalize title,
```python
{
'dr': 'Doktor',
'yb': 'Yang Berhormat',
'hj': 'Haji',
'ybm': 'Yang Berhormat Mulia',
'tyt': 'Tuan Yang Terutama',
'yab': 'Yang Berhormat',
'ybm': 'Yang Berhormat Mulia',
'yabhg': 'Yang Amat Berbahagia',
'ybhg': 'Yang Berbahagia',
'miss': 'Cik',
}
```
```
normalizer.normalize('Dr yahaya')
```
#### 2. expand `x`
```
normalizer.normalize('xtahu')
```
#### 3. normalize `ke -`
```
normalizer.normalize('ke-12')
normalizer.normalize('ke - 12')
```
#### 4. normalize `ke - roman`
```
normalizer.normalize('ke-XXI')
normalizer.normalize('ke - XXI')
```
#### 5. normalize `NUM - NUM`
```
normalizer.normalize('2011 - 2019')
normalizer.normalize('2011.01-2019')
```
#### 6. normalize `pada NUM (/ | -) NUM`
```
normalizer.normalize('pada 10/4')
normalizer.normalize('PADA 10 -4')
```
#### 7. normalize `NUM / NUM`
```
normalizer.normalize('10 /4')
```
#### 8. normalize `rm NUM`
```
normalizer.normalize('RM 10.5')
```
#### 9. normalize `rm NUM sen`
```
normalizer.normalize('rm 10.5 sen')
```
#### 10. normalize `NUM sen`
```
normalizer.normalize('10.5 sen')
```
#### 11. normalize money
```
normalizer.normalize('rm10.4m')
normalizer.normalize('$10.4K')
```
#### 12. normalize cardinal
```
normalizer.normalize('123')
```
#### 13. normalize ordinal
```
normalizer.normalize('ke123')
```
#### 14. normalize date / time / datetime string to datetime.datetime
```
normalizer.normalize('2 hari lepas')
normalizer.normalize('esok')
normalizer.normalize('okt 2019')
normalizer.normalize('2pgi')
normalizer.normalize('pukul 8 malam')
normalizer.normalize('jan 2 2019 12:01pm')
normalizer.normalize('2 ptg jan 2 2019')
```
#### 15. normalize money string to string number representation
```
normalizer.normalize('50 sen')
normalizer.normalize('20.5 ringgit')
normalizer.normalize('20m ringgit')
normalizer.normalize('22.5123334k ringgit')
```
#### 16. normalize date string to %d/%m/%y
```
normalizer.normalize('1 nov 2019')
normalizer.normalize('januari 1 1996')
normalizer.normalize('januari 2019')
```
#### 17. normalize time string to %H:%M:%S
```
normalizer.normalize('2pm')
normalizer.normalize('2:01pm')
normalizer.normalize('2AM')
```
| github_jupyter |
# Converting the Parquet data format to recordIO-wrapped protobuf
---
---
## Contents
1. [Introduction](#Introduction)
1. [Optional data ingestion](#Optional-data-ingestion)
1. [Download the data](#Download-the-data)
1. [Convert into Parquet format](#Convert-into-Parquet-format)
1. [Data conversion](#Data-conversion)
1. [Convert to recordIO protobuf format](#Convert-to-recordIO-protobuf-format)
1. [Upload to S3](#Upload-to-S3)
1. [Training the linear model](#Training-the-linear-model)
## Introduction
In this notebook we illustrate how to convert a Parquet data format into the recordIO-protobuf format that many SageMaker algorithms consume. For the demonstration, first we'll convert the publicly available MNIST dataset into the Parquet format. Subsequently, it is converted into the recordIO-protobuf format and uploaded to S3 for consumption by the linear learner algorithm.
```
import os
import io
import re
import boto3
import pandas as pd
import numpy as np
import time
import sagemaker
from sagemaker import get_execution_role
role = get_execution_role()
sagemaker_session = sagemaker.Session()
bucket = sagemaker_session.default_bucket()
prefix = "sagemaker/DEMO-parquet"
!conda install -y -c conda-forge fastparquet scikit-learn
```
## Optional data ingestion
### Download the data
```
%%time
import pickle, gzip, numpy, urllib.request, json
# Load the dataset
urllib.request.urlretrieve("http://deeplearning.net/data/mnist/mnist.pkl.gz", "mnist.pkl.gz")
with gzip.open("mnist.pkl.gz", "rb") as f:
train_set, valid_set, test_set = pickle.load(f, encoding="latin1")
from fastparquet import write
from fastparquet import ParquetFile
def save_as_parquet_file(dataset, filename, label_col):
X = dataset[0]
y = dataset[1]
data = pd.DataFrame(X)
data[label_col] = y
data.columns = data.columns.astype(str) # Parquet expexts the column names to be strings
write(filename, data)
def read_parquet_file(filename):
pf = ParquetFile(filename)
return pf.to_pandas()
def features_and_target(df, label_col):
X = df.loc[:, df.columns != label_col].values
y = df[label_col].values
return [X, y]
```
### Convert into Parquet format
```
trainFile = "train.parquet"
validFile = "valid.parquet"
testFile = "test.parquet"
label_col = "target"
save_as_parquet_file(train_set, trainFile, label_col)
save_as_parquet_file(valid_set, validFile, label_col)
save_as_parquet_file(test_set, testFile, label_col)
```
## Data conversion
Since algorithms have particular input and output requirements, converting the dataset is also part of the process that a data scientist goes through prior to initiating training. E.g., the Amazon SageMaker implementation of Linear Learner takes recordIO-wrapped protobuf. Most of the conversion effort is handled by the Amazon SageMaker Python SDK, imported as `sagemaker` below.
```
dfTrain = read_parquet_file(trainFile)
dfValid = read_parquet_file(validFile)
dfTest = read_parquet_file(testFile)
train_X, train_y = features_and_target(dfTrain, label_col)
valid_X, valid_y = features_and_target(dfValid, label_col)
test_X, test_y = features_and_target(dfTest, label_col)
```
### Convert to recordIO protobuf format
```
import io
import numpy as np
import sagemaker.amazon.common as smac
trainVectors = np.array([t.tolist() for t in train_X]).astype("float32")
trainLabels = np.where(np.array([t.tolist() for t in train_y]) == 0, 1, 0).astype("float32")
bufTrain = io.BytesIO()
smac.write_numpy_to_dense_tensor(bufTrain, trainVectors, trainLabels)
bufTrain.seek(0)
validVectors = np.array([t.tolist() for t in valid_X]).astype("float32")
validLabels = np.where(np.array([t.tolist() for t in valid_y]) == 0, 1, 0).astype("float32")
bufValid = io.BytesIO()
smac.write_numpy_to_dense_tensor(bufValid, validVectors, validLabels)
bufValid.seek(0)
```
### Upload to S3
```
import boto3
import os
key = "recordio-pb-data"
boto3.resource("s3").Bucket(bucket).Object(os.path.join(prefix, "train", key)).upload_fileobj(
bufTrain
)
s3_train_data = "s3://{}/{}/train/{}".format(bucket, prefix, key)
print("uploaded training data location: {}".format(s3_train_data))
boto3.resource("s3").Bucket(bucket).Object(os.path.join(prefix, "validation", key)).upload_fileobj(
bufValid
)
s3_validation_data = "s3://{}/{}/validation/{}".format(bucket, prefix, key)
print("uploaded validation data location: {}".format(s3_validation_data))
```
## Training the linear model
Once we have the data preprocessed and available in the correct format for training, the next step is to actually train the model using the data. Since this data is relatively small, it isn't meant to show off the performance of the Linear Learner training algorithm, although we have tested it on multi-terabyte datasets.
This example takes four to six minutes to complete. Majority of the time is spent provisioning hardware and loading the algorithm container since the dataset is small.
First, let's specify our containers. Since we want this notebook to run in all 4 of Amazon SageMaker's regions, we'll create a small lookup. More details on algorithm containers can be found in [AWS documentation](https://docs-aws.amazon.com/sagemaker/latest/dg/sagemaker-algo-docker-registry-paths.html).
```
from sagemaker.amazon.amazon_estimator import get_image_uri
container = get_image_uri(boto3.Session().region_name, "linear-learner")
linear_job = "DEMO-linear-" + time.strftime("%Y-%m-%d-%H-%M-%S", time.gmtime())
print("Job name is:", linear_job)
linear_training_params = {
"RoleArn": role,
"TrainingJobName": linear_job,
"AlgorithmSpecification": {"TrainingImage": container, "TrainingInputMode": "File"},
"ResourceConfig": {"InstanceCount": 1, "InstanceType": "ml.c4.2xlarge", "VolumeSizeInGB": 10},
"InputDataConfig": [
{
"ChannelName": "train",
"DataSource": {
"S3DataSource": {
"S3DataType": "S3Prefix",
"S3Uri": "s3://{}/{}/train/".format(bucket, prefix),
"S3DataDistributionType": "FullyReplicated",
}
},
"CompressionType": "None",
"RecordWrapperType": "None",
},
{
"ChannelName": "validation",
"DataSource": {
"S3DataSource": {
"S3DataType": "S3Prefix",
"S3Uri": "s3://{}/{}/validation/".format(bucket, prefix),
"S3DataDistributionType": "FullyReplicated",
}
},
"CompressionType": "None",
"RecordWrapperType": "None",
},
],
"OutputDataConfig": {"S3OutputPath": "s3://{}/{}/".format(bucket, prefix)},
"HyperParameters": {
"feature_dim": "784",
"mini_batch_size": "200",
"predictor_type": "binary_classifier",
"epochs": "10",
"num_models": "32",
"loss": "absolute_loss",
},
"StoppingCondition": {"MaxRuntimeInSeconds": 60 * 60},
}
```
Now let's kick off our training job in SageMaker's distributed, managed training, using the parameters we just created. Because training is managed (AWS handles spinning up and spinning down hardware), we don't have to wait for our job to finish to continue, but for this case, let's setup a while loop so we can monitor the status of our training.
```
%%time
sm = boto3.Session().client("sagemaker")
sm.create_training_job(**linear_training_params)
status = sm.describe_training_job(TrainingJobName=linear_job)["TrainingJobStatus"]
print(status)
sm.get_waiter("training_job_completed_or_stopped").wait(TrainingJobName=linear_job)
if status == "Failed":
message = sm.describe_training_job(TrainingJobName=linear_job)["FailureReason"]
print("Training failed with the following error: {}".format(message))
raise Exception("Training job failed")
sm.describe_training_job(TrainingJobName=linear_job)["TrainingJobStatus"]
```
| github_jupyter |
# The Semi-Supervised VAE
## Introduction
Most of the models we've covered in the tutorials are unsupervised:
- [Variational Autoencoder (VAE)](vae.ipynb)
- [DMM](dmm.ipynb)
- [Attend-Infer-Repeat](air.ipynb)
We've also covered a simple supervised model:
- [Bayesian Regression](bayesian_regression.ipynb)
The semi-supervised setting represents an interesting intermediate case where some of the data is labeled and some is not. It is also of great practical importance, since we often have very little labeled data and much more unlabeled data. We'd clearly like to leverage labeled data to improve our models of the unlabeled data.
The semi-supervised setting is also well suited to generative models, where missing data can be accounted for quite naturally—at least conceptually.
As we will see, in restricting our attention to semi-supervised generative models, there will be no shortage of different model variants and possible inference strategies.
Although we'll only be able to explore a few of these variants in detail, hopefully you will come away from the tutorial with a greater appreciation for the abstractions and modularity offered by probabilistic programming.
So let's go about building a generative model. We have a dataset
$\mathcal{D}$ with $N$ datapoints,
$$ \mathcal{D} = \{ ({\bf x}_i, {\bf y}_i) \} $$
where the $\{ {\bf x}_i \}$ are always observed and the labels $\{ {\bf y}_i \}$ are only observed for some subset of the data. Since we want to be able to model complex variations in the data, we're going to make this a latent variable model with a local latent variable ${\bf z}_i$ private to each pair $({\bf x}_i, {\bf y}_i)$. Even with this set of choices, a number of model variants are possible: we're going to focus on the model variant depicted in Figure 1 (this is model M2 in reference [1]).
For convenience—and since we're going to model MNIST in our experiments below—let's suppose the $\{ {\bf x}_i \}$ are images and the $\{ {\bf y}_i \}$ are digit labels. In this model setup, the latent random variable ${\bf z}_i$ and the (partially observed) digit label _jointly_ generate the observed image.
The ${\bf z}_i$ represents _everything but_ the digit label, possibly handwriting style or position.
Let's sidestep asking when we expect this particular factorization of $({\bf x}_i, {\bf y}_i, {\bf z}_i)$ to be appropriate, since the answer to that question will depend in large part on the dataset in question (among other things). Let's instead highlight some of the ways that inference in this model will be challenging as well as some of the solutions that we'll be exploring in the rest of the tutorial.
## The Challenges of Inference
For concreteness we're going to continue to assume that the partially-observed $\{ {\bf y}_i \}$ are discrete labels; we will also assume that the $\{ {\bf z}_i \}$ are continuous.
- If we apply the general recipe for stochastic variational inference to our model (see [SVI Part I](svi_part_i.ipynb)) we would be sampling the discrete (and thus non-reparameterizable) variable ${\bf y}_i$ whenever it's unobserved. As discussed in [SVI Part III](svi_part_iii.ipynb) this will generally lead to high-variance gradient estimates.
- A common way to ameliorate this problem—and one that we'll explore below—is to forego sampling and instead sum out all ten values of the class label ${\bf y}_i$ when we calculate the ELBO for an unlabeled datapoint ${\bf x}_i$. This is more expensive per step, but can help us reduce the variance of our gradient estimator and thereby take fewer steps.
- Recall that the role of the guide is to 'fill in' _latent_ random variables. Concretely, one component of our guide will be a digit classifier $q_\phi({\bf y} | {\bf x})$ that will randomly 'fill in' labels $\{ {\bf y}_i \}$ given an image $\{ {\bf x}_i \}$. Crucially, this means that the only term in the ELBO that will depend on $q_\phi(\cdot | {\bf x})$ is the term that involves a sum over _unlabeled_ datapoints. This means that our classifier $q_\phi(\cdot | {\bf x})$—which in many cases will be the primary object of interest—will not be learning from the labeled datapoints (at least not directly).
- This seems like a potential problem. Luckily, various fixes are possible. Below we'll follow the approach in reference [1], which involves introducing an additional objective function for the classifier to ensure that the classifier learns directly from the labeled data.
We have our work cut out for us so let's get started!
## First Variant: Standard objective function, naive estimator
As discussed in the introduction, we're considering the model depicted in Figure 1. In more detail, the model has the following structure:
- $p({\bf y}) = Cat({\bf y}~|~{\bf \pi})$: multinomial (or categorical) prior for the class label
- $p({\bf z}) = \mathcal{N}({\bf z}~|~{\bf 0,I})$: unit normal prior for the latent code $\bf z$
- $p_{\theta}({\bf x}~|~{\bf z,y}) = Bernoulli\left({\bf x}~|~\mu\left({\bf z,y}\right)\right)$: parameterized Bernoulli likelihood function; $\mu\left({\bf z,y}\right)$ corresponds to `decoder` in the code
We structure the components of our guide $q_{\phi}(.)$ as follows:
- $q_{\phi}({\bf y}~|~{\bf x}) = Cat({\bf y}~|~{\bf \alpha}_{\phi}\left({\bf x}\right))$: parameterized multinomial (or categorical) distribution; ${\bf \alpha}_{\phi}\left({\bf x}\right)$ corresponds to `encoder_y` in the code
- $q_{\phi}({\bf z}~|~{\bf x, y}) = \mathcal{N}({\bf z}~|~{\bf \mu}_{\phi}\left({\bf x, y}\right), {\bf \sigma^2_{\phi}\left(x, y\right)})$: parameterized normal distribution; ${\bf \mu}_{\phi}\left({\bf x, y}\right)$ and ${\bf \sigma^2_{\phi}\left(x, y\right)}$ correspond to the neural digit classifier `encoder_z` in the code
These choices reproduce the structure of model M2 and its corresponding inference network in reference [1].
We translate this model and guide pair into Pyro code below. Note that:
- The labels `ys`, which are represented with a one-hot encoding, are only partially observed (`None` denotes unobserved values).
- `model()` handles both the observed and unobserved case.
- The code assumes that `xs` and `ys` are mini-batches of images and labels, respectively, with the size of each batch denoted by `batch_size`.
```
def model(self, xs, ys=None):
# register this pytorch module and all of its sub-modules with pyro
pyro.module("ss_vae", self)
batch_size = xs.size(0)
# inform Pyro that the variables in the batch of xs, ys are conditionally independent
with pyro.plate("data"):
# sample the handwriting style from the constant prior distribution
prior_loc = xs.new_zeros([batch_size, self.z_dim])
prior_scale = xs.new_ones([batch_size, self.z_dim])
zs = pyro.sample("z", dist.Normal(prior_loc, prior_scale).to_event(1))
# if the label y (which digit to write) is supervised, sample from the
# constant prior, otherwise, observe the value (i.e. score it against the constant prior)
alpha_prior = xs.new_ones([batch_size, self.output_size]) / (1.0 * self.output_size)
ys = pyro.sample("y", dist.OneHotCategorical(alpha_prior), obs=ys)
# finally, score the image (x) using the handwriting style (z) and
# the class label y (which digit to write) against the
# parametrized distribution p(x|y,z) = bernoulli(decoder(y,z))
# where `decoder` is a neural network
loc = self.decoder([zs, ys])
pyro.sample("x", dist.Bernoulli(loc).to_event(1), obs=xs)
def guide(self, xs, ys=None):
with pyro.plate("data"):
# if the class label (the digit) is not supervised, sample
# (and score) the digit with the variational distribution
# q(y|x) = categorical(alpha(x))
if ys is None:
alpha = self.encoder_y(xs)
ys = pyro.sample("y", dist.OneHotCategorical(alpha))
# sample (and score) the latent handwriting-style with the variational
# distribution q(z|x,y) = normal(loc(x,y),scale(x,y))
loc, scale = self.encoder_z([xs, ys])
pyro.sample("z", dist.Normal(loc, scale).to_event(1))
```
### Network Definitions
In our experiments we use the same network configurations as used in reference [1]. The encoder and decoder networks have one hidden layer with $500$ hidden units and softplus activation functions. We use softmax as the activation function for the output of `encoder_y`, sigmoid as the output activation function for `decoder` and exponentiation for the scale part of the output of `encoder_z`. The latent dimension is 50.
### MNIST Pre-Processing
We normalize the pixel values to the range $[0.0, 1.0]$. We use the [MNIST data loader](http://pytorch.org/docs/master/torchvision/datasets.html#torchvision.datasets.MNIST) from the `torchvision` library. The testing set consists of $10000$ examples. The default training set consists of $60000$ examples. We use the first $50000$ examples for training (divided into supervised and un-supervised parts) and the remaining $10000$ images for validation. For our experiments, we use $4$ configurations of supervision in the training set, i.e. we consider $3000$, $1000$, $600$ and $100$ supervised examples selected randomly (while ensuring that each class is balanced).
### The Objective Function
The objective function for this model has the two terms (c.f. Eqn. 8 in reference [1]):
$$\mathcal{J} = \!\!\sum_{({\bf x,y}) \in \mathcal{D}_{supervised} } \!\!\!\!\!\!\!\!\mathcal{L}\big({\bf x,y}\big) +\!\!\! \sum_{{\bf x} \in \mathcal{D}_{unsupervised}} \!\!\!\!\!\!\!\mathcal{U}\left({\bf x}\right)
$$
To implement this in Pyro, we setup a single instance of the `SVI` class. The two different terms in the objective functions will emerge automatically depending on whether we pass the `step` method labeled or unlabeled data. We will alternate taking steps with labeled and unlabeled mini-batches, with the number of steps taken for each type of mini-batch depending on the total fraction of data that is labeled. For example, if we have 1,000 labeled images and 49,000 unlabeled ones, then we'll take 49 steps with unlabeled mini-batches for each labeled mini-batch. (Note that there are different ways we could do this, but for simplicity we only consider this variant.) The code for this setup is given below:
```
from pyro.infer import SVI, Trace_ELBO, TraceEnum_ELBO, config_enumerate
from pyro.optim import Adam
# setup the optimizer
adam_params = {"lr": 0.0003}
optimizer = Adam(adam_params)
# setup the inference algorithm
svi = SVI(model, guide, optimizer, loss=Trace_ELBO())
```
When we run this inference in Pyro, the performance seen during test time is degraded by the noise inherent in the sampling of the categorical variables (see Figure 2 and Table 1 at the end of this tutorial). To deal with this we're going to need a better ELBO gradient estimator.
## Interlude: Summing Out Discrete Latents
As highlighted in the introduction, when the discrete latent labels ${\bf y}$ are not observed, the ELBO gradient estimates rely on sampling from $q_\phi({\bf y}|{\bf x})$. These gradient estimates can be very high-variance, especially early in the learning process when the guessed labels are often incorrect. A common approach to reduce variance in this case is to sum out discrete latent variables, replacing the Monte Carlo expectation
$$\mathbb E_{{\bf y}\sim q_\phi(\cdot|{\bf x})}\nabla\operatorname{ELBO}$$
with an explicit sum
$$\sum_{\bf y} q_\phi({\bf y}|{\bf x})\nabla\operatorname{ELBO}$$
This sum is usually implemented by hand, as in [1], but Pyro can automate this in many cases. To automatically sum out all discrete latent variables (here only ${\bf y}$), we simply wrap the guide in `config_enumerate()`:
```python
svi = SVI(model, config_enumerate(guide), optimizer, loss=TraceEnum_ELBO(max_plate_nesting=1))
```
In this mode of operation, each `svi.step(...)` computes a gradient term for each of the ten latent states of $y$. Although each step is thus $10\times$ more expensive, we'll see that the lower-variance gradient estimate outweighs the additional cost.
Going beyond the particular model in this tutorial, Pyro supports summing over arbitrarily many discrete latent variables. Beware that the cost of summing is exponential in the number of discrete variables, but is cheap(er) if multiple independent discrete variables are packed into a single tensor (as in this tutorial, where the discrete labels for the entire mini-batch are packed into the single tensor ${\bf y}$). To use this parallel form of `config_enumerate()`, we must inform Pyro that the items in a minibatch are indeed independent by wrapping our vectorized code in a `with pyro.plate("name")` block.
## Second Variant: Standard Objective Function, Better Estimator
Now that we have the tools to sum out discrete latents, we can see if doing so helps our performance. First, as we can see from Figure 3, the test and validation accuracies now evolve much more smoothly over the course of training. More importantly, this single modification improved test accuracy from around `20%` to about `90%` for the case of $3000$ labeled examples. See Table 1 for the full results. This is great, but can we do better?
## Third Variant: Adding a Term to the Objective
For the two variants we've explored so far, the classifier $q_{\phi}({\bf y}~|~ {\bf x})$ doesn't learn directly from labeled data. As we discussed in the introduction, this seems like a potential problem. One approach to addressing this problem is to add an extra term to the objective so that the classifier learns directly from labeled data. Note that this is exactly the approach adopted in reference [1] (see their Eqn. 9). The modified objective function is given by:
\begin{align}
\mathcal{J}^{\alpha} &= \mathcal{J} + \alpha \mathop{\mathbb{E}}_{\tilde{p_l}({\bf x,y})} \big[-\log\big(q_{\phi}({\bf y}~|~ {\bf x})\big)\big] \\
&= \mathcal{J} + \alpha' \sum_{({\bf x,y}) \in \mathcal{D}_{\text{supervised}}} \big[-\log\big(q_{\phi}({\bf y}~|~ {\bf x})\big)\big]
\end{align}
where $\tilde{p_l}({\bf x,y})$ is the empirical distribution over the labeled (or supervised) data and $\alpha' \equiv \frac{\alpha}{|\mathcal{D}_{\text{supervised}}|}$. Note that we've introduced an arbitrary hyperparameter $\alpha$ that modulates the importance of the new term.
To learn using this modified objective in Pyro we do the following:
- We use a new model and guide pair (see the code snippet below) that corresponds to scoring the observed label ${\bf y}$ for a given image ${\bf x}$ against the predictive distribution $q_{\phi}({\bf y}~|~ {\bf x})$
- We specify the scaling factor $\alpha'$ (`aux_loss_multiplier` in the code) in the `pyro.sample` call by making use of `poutine.scale`. Note that `poutine.scale` was used to similar effect in the [Deep Markov Model](dmm.ipynb) to implement KL annealing.
- We create a new `SVI` object and use it to take gradient steps on the new objective term
```
def model_classify(self, xs, ys=None):
pyro.module("ss_vae", self)
with pyro.plate("data"):
# this here is the extra term to yield an auxiliary loss
# that we do gradient descent on
if ys is not None:
alpha = self.encoder_y(xs)
with pyro.poutine.scale(scale=self.aux_loss_multiplier):
pyro.sample("y_aux", dist.OneHotCategorical(alpha), obs=ys)
def guide_classify(xs, ys):
# the guide is trivial, since there are no
# latent random variables
pass
svi_aux = SVI(model_classify, guide_classify, optimizer, loss=Trace_ELBO())
```
When we run inference in Pyro with the additional term in the objective, we outperform both previous inference setups. For example, the test accuracy for the case with $3000$ labeled examples improves from `90%` to `96%` (see Figure 4 below and Table 1 in the next section). Note that we used validation accuracy to select the hyperparameter $\alpha'$.
## Results
| Supervised data | First variant | Second variant | Third variant | Baseline classifier |
|------------------|----------------|----------------|----------------|---------------------|
| 100 | 0.2007(0.0353) | 0.2254(0.0346) | 0.9319(0.0060) | 0.7712(0.0159) |
| 600 | 0.1791(0.0244) | 0.6939(0.0345) | 0.9437(0.0070) | 0.8716(0.0064) |
| 1000 | 0.2006(0.0295) | 0.7562(0.0235) | 0.9487(0.0038) | 0.8863(0.0025) |
| 3000 | 0.1982(0.0522) | 0.8932(0.0159) | 0.9582(0.0012) | 0.9108(0.0015) |
<center> <b>Table 1:</b> Result accuracies (with 95% confidence bounds) for different inference methods</center>
Table 1 collects our results from the three variants explored in the tutorial. For comparison, we also show results from a simple classifier baseline, which only makes use of the supervised data (and no latent random variables). Reported are mean accuracies (with 95% confidence bounds in parentheses) across five random selections of supervised data.
We first note that the results for the third variant—where we summed out the discrete latent random variable $\bf y$ and made use of the additional term in the objective function—reproduce the results reported in reference [1]. This is encouraging, since it means that the abstractions in Pyro proved flexible enough to accomodate the required modeling and inference setup. Significantly, this flexibility was evidently necessary to outperform the baseline. It's also worth emphasizing that the gap between the baseline and third variant of our generative model setup increases as the number of labeled datapoints decreases (maxing out at about 15% for the case with only 100 labeled datapoints). This is a tantalizing result because it's precisely in the regime where we have few labeled data points that semi-supervised learning is particularly attractive.
### Latent Space Visualization
We use <a href="https://lvdmaaten.github.io/tsne/"> T-SNE</a> to reduce the dimensionality of the latent $\bf z$ from $50$ to $2$ and visualize the 10 digit classes in Figure 5. Note that the structure of the embedding is quite different than that in the [VAE](vae.ipynb) case, where the digits are clearly separated from one another in the embedding. This make sense, since for the semi-supervised case the latent $\bf z$ is free to use its representational capacity to model, e.g., handwriting style, since the variation between digits is provided by the (partially observed) labels.
### Conditional image generation
We sampled $100$ images for each class label ($0$ to $9$) by sampling different values of the latent variable ${\bf z}$. The diversity of handwriting styles exhibited by each digit is consistent with what we saw in the T-SNE visualization, suggesting that the representation learned by $\bf z$ is disentangled from the class labels.
## Final thoughts
We've seen that generative models offer a natural approach to semi-supervised machine learning. One of the most attractive features of generative models is that we can explore a large variety of models in a single unified setting. In this tutorial we've only been able to explore a small fraction of the possible model and inference setups that are possible. There is no reason to expect that one variant is best; depending on the dataset and application, there will be reason to prefer one over another. And there are a lot of variants (see Figure 7)!
Some of these variants clearly make more sense than others, but a priori it's difficult to know which ones are worth trying out. This is especially true once we open the door to more complicated setups, like the two models at the bottom of the figure, which include an always latent random variable $\tilde{\bf y}$ in addition to the partially observed label ${\bf y}$. (Incidentally, this class of models—see reference [2] for similar variants—offers another potential solution to the 'no training' problem that we identified above.)
The reader probably doesn't need any convincing that a systematic exploration of even a fraction of these options would be incredibly time-consuming and error-prone if each model and each inference procedure were coded up by scratch. It's only with the modularity and abstraction made possible by a probabilistic programming system that we can hope to explore the landscape of generative models with any kind of nimbleness—and reap any awaiting rewards.
See the full code on [Github](https://github.com/pyro-ppl/pyro/blob/dev/examples/vae/ss_vae_M2.py).
## References
[1] `Semi-supervised Learning with Deep Generative Models`,<br/>
Diederik P. Kingma, Danilo J. Rezende, Shakir Mohamed, Max Welling
[2] `Learning Disentangled Representations with Semi-Supervised Deep Generative Models`,
<br/>
N. Siddharth, Brooks Paige, Jan-Willem Van de Meent, Alban Desmaison, Frank Wood, <br/>
Noah D. Goodman, Pushmeet Kohli, Philip H.S. Torr
| github_jupyter |
### Note
* Instructions have been included for each segment. You do not have to follow them exactly, but they are included to help you think through the steps.
```
# Dependencies and Setup
import pandas as pd
# File to Load (Remember to Change These)
file_to_load = "Resources/purchase_data.csv"
# Read Purchasing File and store into Pandas data frame
purchase_data = pd.read_csv(file_to_load)
purchase_data
```
## Player Count
* Display the total number of players
```
total=purchase_data["SN"].nunique()
total_players= {"total Players":[total]}
df=pd.DataFrame(total_players,index=[0])
df
```
## Purchasing Analysis (Total)
* Run basic calculations to obtain number of unique items, average price, etc.
* Create a summary data frame to hold the results
* Optional: give the displayed data cleaner formatting
* Display the summary data frame
```
#number of unique items
purchase_data_groubby= purchase_data.groupby("Item ID")
unique_item=len(purchase_data_groubby["Item ID"].nunique())
unique_item
#average price(I could not round a single value)
average_price=purchase_data["Price"].mean()
#average_price="${:,.2f}".format(purchase_data["Price"].mean())
##purchase_data["Price"]=purchase_data["Price"].map("${:,.2f}".format)
average_price
#number of perchases
number_purchases=purchase_data["Item ID"].count()
number_purchases
#Total Revenue
Total_Revenue=purchase_data["Price"].sum()
Total_Revenue
Purchasing_Analysis=pd.DataFrame({"number of Unique Items":unique_item,
"Average Price":[average_price],
"Number of Purchases":number_purchases,
"Total Revenue":[Total_Revenue]},index=[0])
Purchasing_Analysis
Purchasing_Analysis["Average Price"]=Purchasing_Analysis["Average Price"].map("${:,.2f}".format)
Purchasing_Analysis["Total Revenue"]=Purchasing_Analysis["Total Revenue"].map("${:,.2f}".format)
Purchasing_Analysis
```
## Gender Demographics
* Percentage and Count of Male Players
* Percentage and Count of Female Players
* Percentage and Count of Other / Non-Disclosed
```
#purchase_data.groupby("Gender")
#purchase_data.groupby("Gender")
total_count=purchase_data.groupby(["Gender"]).nunique()
total_count_percentage=total_count["SN"]/total*100
total_count_percentage=total_count_percentage.map("{:,.2f}%".format)
Gender_Demographics=pd.DataFrame({"Total Count":total_count["SN"],
"Percentage of Players":total_count_percentage},
index=["Male", "Female", "Other / Non-Disclosed"])
Gender_Demographics
total_count
#purchase_data=purchase_data.drop_duplicates(inplace=True)
#male_players=len(purchase_data["Gender"].loc[purchase_data["Gender"]=="Male"])
#female_players=len(purchase_data["Gender"].loc[purchase_data["Gender"]=="Female"])
#tother_players=len(purchase_data["Gender"])-male_players-female_players
#total_players=len(purchase_data["Gender"])
#Gender_Demographics=pd.DataFrame({"Total Count":[male_players,female_players,other_players],
# "Percentage of Players":[male_players/total_players*100,female_players/total_players*100,other_players/total_players*100]
# },index=["Male", "Female", "Other / Non-Disclosed"])
#Gender_Demographics
```
## Purchasing Analysis (Gender)
* Run basic calculations to obtain purchase count, avg. purchase price, avg. purchase total per person etc. by gender
* Create a summary data frame to hold the results
* Optional: give the displayed data cleaner formatting
* Display the summary data frame
```
total_count=purchase_data.groupby(["Gender"]).nunique()
total_count_percentage=total_count["SN"]/total*100
total_count_percentage=total_count_percentage.map("{:,.2f}%".format)
Gender_Demographics=pd.DataFrame({"Total Count":total_count["SN"],
"Percentage of Players":total_count_percentage},
index=["Male", "Female", "Other / Non-Disclosed"])
Gender_Demographics
#purchase count
pd_group_gender=purchase_data.groupby(["Gender"])
total_count_2=pd_group_gender["SN"].count()
#average price
ave_count=pd_group_gender["Price"].mean()
ave_count=ave_count.map("{:,.2f}%".format)
#Total Purchase Value(T_P_V)
T_P_V=pd_group_gender["Price"].sum()
#Avg Total Purchase per Person(A_T_P_P_P)
A_T_P_P_P=T_P_V/Gender_Demographics["Total Count"]
#clear formating
T_P_V=T_P_V.map("${:,.2f}".format)
A_T_P_P_P=A_T_P_P_P.map("${:,.2f}%".format)
#Create a summary data frame to hold the results
summery_gender=pd.DataFrame({"Total Count":total_count_2,
"Average Price":ave_count,
"Total Purchase Value":T_P_V,
"Avg Total Purchase Per Person": A_T_P_P_P
})
summery_gender
```
## Age Demographics
* Establish bins for ages
* Categorize the existing players using the age bins. Hint: use pd.cut()
* Calculate the numbers and percentages by age group
* Create a summary data frame to hold the results
* Optional: round the percentage column to two decimal points
* Display Age Demographics Table
```
#Age Demographics
bins=[0,9,14,19,24,29,34,39,2000]
labels_group=["<10","10-14","15-19","20-24","25-29","30-34","35-39","40+"]
purchase_data["score summery"]=pd.cut(purchase_data["Age"],bins, labels=labels_group)
age_group=purchase_data.groupby(["score summery"])
#Calculate the numbers and percentages by age group(n_age_groups) and (p_age_groups)
n_age_groups=age_group["SN"].nunique()
p_age_groups=n_age_groups/total*100
# round the percentage column to two decimal points
p_age_groups=p_age_groups.map("{:,.2f}%".format)
#Create a summary data frame to hold the results
Age_Demographics=pd.DataFrame({"Total Count":n_age_groups,
"Percentage of Players":p_age_groups,
})
Age_Demographics
```
## Purchasing Analysis (Age)
* Bin the purchase_data data frame by age
* Run basic calculations to obtain purchase count, avg. purchase price, avg. purchase total per person etc. in the table below
* Create a summary data frame to hold the results
* Optional: give the displayed data cleaner formatting
* Display the summary data frame
```
purchase_data["score summery"]=pd.cut(purchase_data["Age"],bins, labels=labels_group)
age_group=purchase_data.groupby(["score summery"])
#purchase count
total_count_age=age_group["Purchase ID"].nunique()
#Average Purchase Price(A_P_P_A)
A_P_P_A=age_group["Price"].mean()
#Total purchase value (abreviation:tpv)
tpv=total_count_age*A_P_P_A
#Avg Total Purchase per Person(abreviation:atpp),total count including repetetive customer(abreviation:tcirc)
tcirc=age_group["Purchase ID"].count()
atpp =tpv/Age_Demographics["Total Count"]
#give the displayed data cleaner formatting
A_P_P_A=A_P_P_A.map("${:,.2f}".format)
tpv=tpv.map("${:,.2f}".format)
atpp=atpp.map("${:,.2f}%".format)
#Create a summary data frame to hold the results
Purchasing_Analysis_Age=pd.DataFrame({"purchase count":total_count_age,
"Average Purchase Price":A_P_P_A,
"Total purchase value":tpv,
"Avg Total Purchase per Person":atpp
})
Purchasing_Analysis_Age
```
## Top Spenders
* Run basic calculations to obtain the results in the table below
* Create a summary data frame to hold the results
* Sort the total purchase value column in descending order
* Optional: give the displayed data cleaner formatting
* Display a preview of the summary data frame
```
#number of unique items(abreviation:nui)
#Group by Spendors ( "SN" )
nui=purchase_data.groupby(["SN"])
nui_count=nui["Purchase ID"].count()
#Average Purchase Price
nui_ave=nui["Price"].mean()
#Total Purchase Value
nui_sum=nui["Price"].sum()
#give the displayed data cleaner formatting
nui_ave=nui_ave.map("${:,.2f}".format)
#Create a summary data frame to hold the results
Top_Spenders=pd.DataFrame({"purchase count":nui_count,
"Average Purchase Price":nui_ave,
"Total purchase value":nui_sum,
})
#Sort the total purchase value column in descending order
Top_Spenders=Top_Spenders.sort_values("Total purchase value",ascending=False)
#give the displayed data cleaner formatting
Top_Spenders["Total purchase value"]=nui_sum.map("${:,.2f}".format)
Top_Spenders
```
## Most Popular Items
* Retrieve the Item ID, Item Name, and Item Price columns
* Group by Item ID and Item Name. Perform calculations to obtain purchase count, item price, and total purchase value
* Create a summary data frame to hold the results
* Sort the purchase count column in descending order
* Optional: give the displayed data cleaner formatting
* Display a preview of the summary data frame
```
#Group by Item ID and Item Name(abbreviation:giiin)
giiin=purchase_data.groupby(["Item ID","Item Name"])
#Purchase Count(abbreviation:pc)
pc=giiin["Purchase ID"].nunique()
#Item Price (abbreviasion:ip)
ip=giiin["Price"].mean()
#total purchase value by item(tpvi)
tpvi=giiin["Price"].sum()
#item price formating
ip=ip.map("${:,.2f}".format)
tpvi=tpvi.map("${:,.2f}".format)
#Create a summary data frame to hold the results
Most_Popular_Items=pd.DataFrame({"purchase count":pc,
"Item Price":ip,
"Total purchase value":tpvi,
})
#Sort the purchase count in descending order
Most_Popular_Items=Most_Popular_Items.sort_values("purchase count",ascending=False)
Most_Popular_Items
```
## Most Profitable Items
* Sort the above table by total purchase value in descending order
* Optional: give the displayed data cleaner formatting
* Display a preview of the data frame
```
#total purchase value by item(tpvi)
tpvi=giiin["Price"].sum()
#Create a summary data frame to hold the results
Most_Profitable_Items=pd.DataFrame({"purchase count":pc,
"Item Price":ip,
"Total purchase value":tpvi,
})
#Sort the purchase count in descending order
Most_Profitable_Items=Most_Profitable_Items.sort_values("Total purchase value",ascending=False)
Most_Profitable_Items["Total purchase value"]=Most_Profitable_Items["Total purchase value"].map("${:,.2f}".format)
Most_Profitable_Items
```
# Conclusions
Of the 780 active players, the vast majority are male (84%). There also exists, a smaller, but notable proportion of female layers (14%).
Our peak age demographic falls between 20-24 (46.8%) with secondary groups falling between 15-19 (17.4%) and 25-29 (13%).
The majority of purchases are also done by the age group 20-24 (46.8%) with secondary groups falling between 15-19 (17.4%) and 25-29 (13%).
Out of 183 items offered, the most popular and profitable ones are "Oathbreaker, Last Hope of the Breaking Storm" (12 buys), brought $51 and "Nirvana" and "Fiery Glass Crusader" having (9 buys) each and brought $44 and $41 respectively. Generally, all players (780) prefer different items, there are no significantly more popular item(s) than others.
Average purchase is about $3 per person with the top spenders paying up to $19 for their purchases. Still, 97% are paying way under $10. The total profit from the sold items is about $2400 for 780 players.
| github_jupyter |
<script async src="https://www.googletagmanager.com/gtag/js?id=UA-59152712-8"></script>
<script>
window.dataLayer = window.dataLayer || [];
function gtag(){dataLayer.push(arguments);}
gtag('js', new Date());
gtag('config', 'UA-59152712-8');
</script>
# The BSSN Formulation of General Relativity in Generic Curvilinear Coordinates: An Overview
## Author: Zach Etienne
## This tutorial notebook demonstrates how Einstein's equations of general relativity in this formulation are constructed and output within NRPy+.
### As Einstein's equations in this formalism take the form of highly nonlinear, coupled *wave equations*, the [tutorial notebook on the scalar wave equation in curvilinear coordinates](Tutorial-ScalarWaveCurvilinear.ipynb) is *required* reading before beginning this module. That module, as well as its own prerequisite [module on reference metrics within NRPy+](Tutorial-Reference_Metric.ipynb) provides the needed overview of how NRPy+ handles reference metrics.
## Introduction:
NRPy+'s original purpose was to be an easy-to-use code capable of generating Einstein's equations in a broad class of [singular](https://en.wikipedia.org/wiki/Coordinate_singularity), curvilinear coordinate systems, where the user need only input the scale factors of the underlying reference metric. Upon generating these equations, NRPy+ would then leverage SymPy's [common-expression-elimination (CSE)](https://en.wikipedia.org/wiki/Common_subexpression_elimination) and C code generation routines, coupled to its own [single-instruction, multiple-data (SIMD)](https://en.wikipedia.org/wiki/SIMD) functions, to generate highly-optimized C code.
### Background Reading/Lectures:
* Mathematical foundations of BSSN and 3+1 initial value problem decompositions of Einstein's equations:
* [Thomas Baumgarte's lectures on mathematical formulation of numerical relativity](https://www.youtube.com/watch?v=t3uo2R-yu4o&list=PLRVOWML3TL_djTd_nsTlq5aJjJET42Qke)
* [Yuichiro Sekiguchi's introduction to BSSN](http://www2.yukawa.kyoto-u.ac.jp/~yuichiro.sekiguchi/3+1.pdf)
* Extensions to the standard BSSN approach used in NRPy+
* [Brown's covariant "Lagrangian" formalism of BSSN](https://arxiv.org/abs/0902.3652)
* [BSSN in spherical coordinates, using the reference-metric approach of Baumgarte, Montero, Cordero-Carrión, and Müller (2012)](https://arxiv.org/abs/1211.6632)
* [BSSN in generic curvilinear coordinates, using the extended reference-metric approach of Ruchlin, Etienne, and Baumgarte (2018)](https://arxiv.org/abs/1712.07658)
### A Note on Notation:
As is standard in NRPy+,
* Greek indices refer to four-dimensional quantities where the zeroth component indicates temporal (time) component.
* Latin indices refer to three-dimensional quantities. This is somewhat counterintuitive since Python always indexes its lists starting from 0. As a result, the zeroth component of three-dimensional quantities will necessarily indicate the first *spatial* direction.
As a corollary, any expressions involving mixed Greek and Latin indices will need to offset one set of indices by one: A Latin index in a four-vector will be incremented and a Greek index in a three-vector will be decremented (however, the latter case does not occur in this tutorial notebook).
<a id='toc'></a>
# Table of Contents
$$\label{toc}$$
This module lays out the mathematical foundation for the BSSN formulation of Einstein's equations, as detailed in the references in the above Background Reading/Lectures section. It is meant to provide an overview of the basic equations and point of reference for **full tutorial notebooks** linked below:
1. [Step 1](#brownslagrangebssn): [Brown](https://arxiv.org/abs/0902.3652)'s covariant formulation of the BSSN time-evolution equations (see next section for gauge conditions)
1. [Step 1.a](#fullequations): Numerical implementation of BSSN time-evolution equations
1. [Step 1.a.i](#liederivs) ([**BSSN quantities module [start here]**](Tutorial-BSSN_quantities.ipynb); [**BSSN time-evolution module**](Tutorial-BSSN_time_evolution-BSSN_RHSs.ipynb)): Expanding the Lie derivatives; the BSSN time-evolution equations in their final form
1. [Step 2](#gaugeconditions) ([**full tutorial notebook**](Tutorial-BSSN_time_evolution-BSSN_gauge_RHSs.ipynb)): Time-evolution equations for the BSSN |gauge quantities $\alpha$ and $\beta^i$
1. [Step 3](#constraintequations) ([**full tutorial notebook**](Tutorial-BSSN_constraints.ipynb)): The BSSN constraint equations
1. [Step 3.a](#hamiltonianconstraint): The Hamiltonian constraint
1. [Step 3.b](#momentumconstraint): The momentum constraint
1. [Step 4](#gammaconstraint) ([**full tutorial notebook**](Tutorial-BSSN_enforcing_determinant_gammabar_equals_gammahat_constraint.ipynb)): The BSSN algebraic constraint $\hat{\gamma}=\bar{\gamma}$
1. [Step 5](#latex_pdf_output) Output this notebook to $\LaTeX$-formatted PDF file
<a id='brownslagrangebssn'></a>
# Step 1: [Brown](https://arxiv.org/abs/0902.3652)'s covariant formulation of BSSN \[Back to [top](#toc)\]
$$\label{brownslagrangebssn}$$
The covariant "Lagrangian" BSSN formulation of [Brown (2009)](https://arxiv.org/abs/0902.3652), which requires
$$
\partial_t \bar{\gamma} = 0,
$$
results in the BSSN equations taking the following form (Eqs. 11 and 12 in [Ruchlin, Etienne, and Baumgarte (2018)](https://arxiv.org/abs/1712.07658)):
\begin{align}
\partial_{\perp} \bar{\gamma}_{i j} {} = {} & \frac{2}{3} \bar{\gamma}_{i j} \left (\alpha \bar{A}_{k}^{k} - \bar{D}_{k} \beta^{k}\right ) - 2 \alpha \bar{A}_{i j} \; , \\
\partial_{\perp} \bar{A}_{i j} {} = {} & -\frac{2}{3} \bar{A}_{i j} \bar{D}_{k} \beta^{k} - 2 \alpha \bar{A}_{i k} {\bar{A}^{k}}_{j} + \alpha \bar{A}_{i j} K \nonumber \\
& + e^{-4 \phi} \left \{-2 \alpha \bar{D}_{i} \bar{D}_{j} \phi + 4 \alpha \bar{D}_{i} \phi \bar{D}_{j} \phi \right . \nonumber \\
& \left . + 4 \bar{D}_{(i} \alpha \bar{D}_{j)} \phi - \bar{D}_{i} \bar{D}_{j} \alpha + \alpha \bar{R}_{i j} \right \}^{\text{TF}} \; , \\
\partial_{\perp} \phi {} = {} & \frac{1}{6} \left (\bar{D}_{k} \beta^{k} - \alpha K \right ) \; , \\
\partial_{\perp} K {} = {} & \frac{1}{3} \alpha K^{2} + \alpha \bar{A}_{i j} \bar{A}^{i j} \nonumber \\
& - e^{-4 \phi} \left (\bar{D}_{i} \bar{D}^{i} \alpha + 2 \bar{D}^{i} \alpha \bar{D}_{i} \phi \right ) \; , \\
\partial_{\perp} \bar{\Lambda}^{i} {} = {} & \bar{\gamma}^{j k} \hat{D}_{j} \hat{D}_{k} \beta^{i} + \frac{2}{3} \Delta^{i} \bar{D}_{j} \beta^{j} + \frac{1}{3} \bar{D}^{i} \bar{D}_{j} \beta^{j} \nonumber \\
& - 2 \bar{A}^{i j} \left (\partial_{j} \alpha - 6 \partial_{j} \phi \right ) + 2 \bar{A}^{j k} \Delta_{j k}^{i} \nonumber \\
& -\frac{4}{3} \alpha \bar{\gamma}^{i j} \partial_{j} K \\
\end{align}
where
* the $\text{TF}$ superscript denotes the trace-free part.
* $\bar{\gamma}_{ij} = \varepsilon_{i j} + \hat{\gamma}_{ij}$, where $\bar{\gamma}_{ij} = e^{-4\phi} \gamma_{ij}$ is the conformal metric, $\gamma_{ij}$ is the physical metric (see below), and $\varepsilon_{i j}$ encodes information about the non-hatted metric.
* $\gamma_{ij}$, $\beta^i$, and $\alpha$ are the physical (as opposed to conformal) spatial 3-metric, shift vector, and lapse, respectively, which may be defined via the 3+1 decomposition line element (in [$G=c=1$ units](https://en.wikipedia.org/wiki/Planck_units)):
$$ds^2 = -\alpha^2 dt^2 + \gamma_{ij}\left(dx^i + \beta^i dt\right)\left(dx^j + \beta^j dt\right).$$
* $\bar{R}_{ij}$ is the conformal Ricci tensor, computed via
\begin{align}
\bar{R}_{i j} {} = {} & - \frac{1}{2} \bar{\gamma}^{k l} \hat{D}_{k} \hat{D}_{l} \bar{\gamma}_{i j} + \bar{\gamma}_{k(i} \hat{D}_{j)} \bar{\Lambda}^{k} + \Delta^{k} \Delta_{(i j) k} \nonumber \\
& + \bar{\gamma}^{k l} \left (2 \Delta_{k(i}^{m} \Delta_{j) m l} + \Delta_{i k}^{m} \Delta_{m j l} \right ) \; .
\end{align}
* $\partial_{\perp} = \partial_t - \mathcal{L}_\beta$; $\mathcal{L}_\beta$ is the [Lie derivative](https://en.wikipedia.org/wiki/Lie_derivative) along the shift vector $\beta^i$.
* $\partial_0 = \partial_t - \beta^i \partial_i$ is an advective time derivative.
* $\hat{D}_j$ is the [covariant derivative](https://en.wikipedia.org/wiki/Covariant_derivative) with respect to the reference metric $\hat{\gamma}_{ij}$.
* $\bar{D}_j$ is the [covariant derivative](https://en.wikipedia.org/wiki/Covariant_derivative) with respect to the barred spatial 3-metric $\bar{\gamma}_{ij}$
* $\Delta^i_{jk}$ is the tensor constructed from the difference of barred and hatted Christoffel symbols:
$$\Delta^i_{jk} = \bar{\Gamma}^i_{jk} - \hat{\Gamma}^i_{jk}$$
* The related quantity $\Delta^i$ is defined $\Delta^i \equiv \bar{\gamma}^{jk} \Delta^i_{jk}$.
* $\bar{A}_{ij}$ is the conformal, trace-free extrinsic curvature:
$$\bar{A}_{ij} = e^{-4\phi} \left(K_{ij} - \frac{1}{3}\gamma_{ij} K\right),$$
where $K$ is the trace of the extrinsic curvature $K_{ij}$.
<a id='fullequations'></a>
## Step 1.a: Numerical implementation of BSSN time-evolution equations \[Back to [top](#toc)\]
$$\label{fullequations}$$
Regarding the numerical implementation of the above equations, first notice the left-hand sides of the equations include the time derivatives. Numerically, these equations are solved using as an [initial value problem](https://en.wikipedia.org/wiki/Initial_value_problem), where data are specified at a given time $t$, so that the solution at any later time can be obtained using the [Method of Lines (MoL)](https://en.wikipedia.org/wiki/Method_of_lines). MoL requires that the equations be written in the form:
$$\partial_t \vec{U} = \vec{f}\left(\vec{U},\partial_i \vec{U}, \partial_i \partial_j \vec{U},...\right),$$
for the vector of "evolved quantities" $\vec{U}$, where the right-hand side vector $\vec{f}$ *does not* contain *explicit* time derivatives of $\vec{U}$.
Thus we must first rewrite the above equations so that *only* partial derivatives of time appear on the left-hand sides of the equations, meaning that the Lie derivative terms must be moved to the right-hand sides of the equations.
<a id='liederivs'></a>
### Step 1.a.i: Expanding the Lie derivatives; BSSN equations in their final form \[Back to [top](#toc)\]
$$\label{liederivs}$$
In this Step, we provide explicit expressions for the [Lie derivatives](https://en.wikipedia.org/wiki/Lie_derivative) $\mathcal{L}_\beta$ appearing inside the $\partial_\perp = \partial_t - \mathcal{L}_\beta$ operators for $\left\{\bar{\gamma}_{i j},\bar{A}_{i j},W, K, \bar{\Lambda}^{i}\right\}$.
In short, the Lie derivative of tensor weight $w$ is given by (from [the wikipedia article on Lie derivatives](https://en.wikipedia.org/wiki/Lie_derivative))
\begin{align}
(\mathcal {L}_X T) ^{a_1 \ldots a_r}{}_{b_1 \ldots b_s} &= X^c(\partial_c T^{a_1 \ldots a_r}{}_{b_1 \ldots b_s}) \\
&\quad - (\partial_c X ^{a_1}) T ^{c a_2 \ldots a_r}{}_{b_1 \ldots b_s} - \ldots - (\partial_c X^{a_r}) T ^{a_1 \ldots a_{r-1}c}{}_{b_1 \ldots b_s} \\
&\quad + (\partial_{b_1} X^c) T ^{a_1 \ldots a_r}{}_{c b_2 \ldots b_s} + \ldots + (\partial_{b_s} X^c) T ^{a_1 \ldots a_r}{}_{b_1 \ldots b_{s-1} c} + w (\partial_{c} X^c) T ^{a_1 \ldots a_r}{}_{b_1 \ldots b_{s}}
\end{align}
Thus to evaluate the Lie derivative, one must first know the tensor density weight $w$ for each tensor. In this formulation of Einstein's equations, **all evolved quantities have density weight $w=0$**, so according to the definition of Lie derivative above,
\begin{align}
\mathcal{L}_\beta \bar{\gamma}_{ij} &= \beta^k \partial_k \bar{\gamma}_{ij} + \partial_i \beta^k \bar{\gamma}_{kj} + \partial_j \beta^k \bar{\gamma}_{ik}, \\
\mathcal{L}_\beta \bar{A}_{ij} &= \beta^k \partial_k \bar{A}_{ij} + \partial_i \beta^k \bar{A}_{kj} + \partial_j \beta^k \bar{A}_{ik}, \\
\mathcal{L}_\beta \phi &= \beta^k \partial_k \phi, \\
\mathcal{L}_\beta K &= \beta^k \partial_k K, \\
\mathcal{L}_\beta \bar{\Lambda}^i &= \beta^k \partial_k \bar{\Lambda}^i - \partial_k \beta^i \bar{\Lambda}^k
\end{align}
With these definitions, the BSSN equations for the un-rescaled evolved variables in the form $\partial_t \vec{U} = f\left(\vec{U},\partial_i \vec{U}, \partial_i \partial_j \vec{U},...\right)$ become
\begin{align}
\partial_t \bar{\gamma}_{i j} {} = {} & \left[\beta^k \partial_k \bar{\gamma}_{ij} + \partial_i \beta^k \bar{\gamma}_{kj} + \partial_j \beta^k \bar{\gamma}_{ik} \right] + \frac{2}{3} \bar{\gamma}_{i j} \left (\alpha \bar{A}_{k}^{k} - \bar{D}_{k} \beta^{k}\right ) - 2 \alpha \bar{A}_{i j} \; , \\
\partial_t \bar{A}_{i j} {} = {} & \left[\beta^k \partial_k \bar{A}_{ij} + \partial_i \beta^k \bar{A}_{kj} + \partial_j \beta^k \bar{A}_{ik} \right] - \frac{2}{3} \bar{A}_{i j} \bar{D}_{k} \beta^{k} - 2 \alpha \bar{A}_{i k} {\bar{A}^{k}}_{j} + \alpha \bar{A}_{i j} K \nonumber \\
& + e^{-4 \phi} \left \{-2 \alpha \bar{D}_{i} \bar{D}_{j} \phi + 4 \alpha \bar{D}_{i} \phi \bar{D}_{j} \phi + 4 \bar{D}_{(i} \alpha \bar{D}_{j)} \phi - \bar{D}_{i} \bar{D}_{j} \alpha + \alpha \bar{R}_{i j} \right \}^{\text{TF}} \; , \\
\partial_t \phi {} = {} & \left[\beta^k \partial_k \phi \right] + \frac{1}{6} \left (\bar{D}_{k} \beta^{k} - \alpha K \right ) \; , \\
\partial_{t} K {} = {} & \left[\beta^k \partial_k K \right] + \frac{1}{3} \alpha K^{2} + \alpha \bar{A}_{i j} \bar{A}^{i j} - e^{-4 \phi} \left (\bar{D}_{i} \bar{D}^{i} \alpha + 2 \bar{D}^{i} \alpha \bar{D}_{i} \phi \right ) \; , \\
\partial_t \bar{\Lambda}^{i} {} = {} & \left[\beta^k \partial_k \bar{\Lambda}^i - \partial_k \beta^i \bar{\Lambda}^k \right] + \bar{\gamma}^{j k} \hat{D}_{j} \hat{D}_{k} \beta^{i} + \frac{2}{3} \Delta^{i} \bar{D}_{j} \beta^{j} + \frac{1}{3} \bar{D}^{i} \bar{D}_{j} \beta^{j} \nonumber \\
& - 2 \bar{A}^{i j} \left (\partial_{j} \alpha - 6 \partial_{j} \phi \right ) + 2 \alpha \bar{A}^{j k} \Delta_{j k}^{i} -\frac{4}{3} \alpha \bar{\gamma}^{i j} \partial_{j} K
\end{align}
where the terms moved from the right-hand sides to the left-hand sides are enclosed in square braces.
Notice that the shift advection operator $\beta^k \partial_k \left\{\bar{\gamma}_{i j},\bar{A}_{i j},\phi, K, \bar{\Lambda}^{i}, \alpha, \beta^i, B^i\right\}$ appears on the right-hand side of *every* expression. As the shift determines how the spatial coordinates $x^i$ move on the next 3D slice of our 4D manifold, we find that representing $\partial_k$ in these shift advection terms via an *upwinded* finite difference stencil results in far lower numerical errors. This trick is implemented below in all shift advection terms.
As discussed in the [NRPy+ tutorial notebook on BSSN quantities](Tutorial-BSSN_quantities.ipynb), tensorial expressions can diverge at coordinate singularities, so each tensor in the set of BSSN variables
$$\left\{\bar{\gamma}_{i j},\bar{A}_{i j},\phi, K, \bar{\Lambda}^{i}, \alpha, \beta^i, B^i\right\},$$
is written in terms of the corresponding rescaled quantity in the set
$$\left\{h_{i j},a_{i j},\text{cf}, K, \lambda^{i}, \alpha, \mathcal{V}^i, \mathcal{B}^i\right\},$$
respectively, as defined in the [BSSN quantities tutorial](Tutorial-BSSN_quantities.ipynb).
<a id='gaugeconditions'></a>
# Step 2: Time-evolution equations for the BSSN gauge quantities $\alpha$ and $\beta^i$ \[Back to [top](#toc)\]
$$\label{gaugeconditions}$$
As described in the **Background Reading/Lectures** linked to above, the gauge quantities $\alpha$ and $\beta^i$ specify how coordinate time and spatial points adjust from one spatial hypersurface to the next, in our 3+1 decomposition of Einstein's equations.
As choosing $\alpha$ and $\beta^i$ is equivalent to choosing coordinates for where we sample our solution to Einstein's equations, we are completely free to choose $\alpha$ and $\beta^i$ on any given spatial hypersuface. It has been found that fixing $\alpha$ and $\beta^i$ to constant values in the context of dynamical spacetimes results in instabilities, so we generally need to define expressions for $\partial_t \alpha$ and $\partial_t \beta^i$ and couple these equations to the rest of the BSSN time-evolution equations.
Though we are free to choose the form of the right-hand sides of the gauge time evolution equations, very few have been found robust in the presence of (puncture) black holes.
The most commonly adopted gauge conditions for BSSN (i.e., time-evolution equations for the BSSN gauge quantities $\alpha$ and $\beta^i$) are the
* $1+\log$ lapse condition:
$$
\partial_0 \alpha = -2 \alpha K
$$
* Second-order Gamma-driving shift condition:
\begin{align}
\partial_0 \beta^i &= B^{i} \\
\partial_0 B^i &= \frac{3}{4} \partial_{0} \bar{\Lambda}^{i} - \eta B^{i},
\end{align}
where $\partial_0$ is the advection operator; i.e., $\partial_0 A^i = \partial_t A^i - \beta^j \partial_j A^i$. Note that $\partial_{0} \bar{\Lambda}^{i}$ in the right-hand side of the $\partial_{0} B^{i}$ equation is computed by adding $\beta^j \partial_j \bar{\Lambda}^i$ to the right-hand side expression given for $\partial_t \bar{\Lambda}^i$, so no explicit time dependence occurs in the right-hand sides of the BSSN evolution equations and the Method of Lines can be applied directly.
While it is incredibly robust in Cartesian coordinates, [Brown](https://arxiv.org/abs/0902.3652) pointed out that the above time-evolution equation for the shift is not covariant. In fact, we have found this non-covariant version to result in very poor results when solving Einstein's equations in spherical coordinates for a spinning black hole with spin axis pointed in the $\hat{x}$ direction. Therefore we adopt Brown's covariant version as described in the [**full time-evolution equations for the BSSN gauge quantities $\alpha$ and $\beta^i$ tutorial notebook**](Tutorial-BSSN_time_evolution-BSSN_gauge_RHSs.ipynb).
<a id='constraintequations'></a>
# Step 3: The BSSN constraint equations \[Back to [top](#toc)\]
$$\label{constraintequations}$$
In a way analogous to Maxwell's equations, the BSSN decomposition of Einstein's equations are written as a set of time-evolution equations and a set of constraint equations. In this step we present the BSSN constraints
\begin{align}
\mathcal{H} &= 0 \\
\mathcal{M^i} &= 0,
\end{align}
where $\mathcal{H}=0$ is the **Hamiltonian constraint**, and $\mathcal{M^i} = 0$ is the **momentum constraint**. When constructing our spacetime from the initial data, one spatial hypersurface at a time, to confirm that at a given time, the Hamiltonian and momentum constraint violations converge to zero as expected with increased numerical resolution.
<a id='hamiltonianconstraint'></a>
## Step 3.a: The Hamiltonian constraint $\mathcal{H}$ \[Back to [top](#toc)\]
$$\label{hamiltonianconstraint}$$
The Hamiltonian constraint is written (Eq. 13 of [Baumgarte *et al.*](https://arxiv.org/pdf/1211.6632.pdf)):
$$
\mathcal{H} = \frac{2}{3} K^2 - \bar{A}_{ij} \bar{A}^{ij} + e^{-4\phi} \left(\bar{R} - 8 \bar{D}^i \phi \bar{D}_i \phi - 8 \bar{D}^2 \phi\right)
$$
<a id='momentumconstraint'></a>
## Step 3.b: The momentum constraint $\mathcal{M}^i$ \[Back to [top](#toc)\]
$$\label{momentumconstraint}$$
The momentum constraint is written (Eq. 47 of [Ruchlin, Etienne, & Baumgarte](https://arxiv.org/pdf/1712.07658.pdf)):
$$ \mathcal{M}^i = e^{-4\phi} \left(
\frac{1}{\sqrt{\bar{\gamma}}} \hat{D}_j\left(\sqrt{\bar{\gamma}}\bar{A}^{ij}\right) +
6 \bar{A}^{ij}\partial_j \phi -
\frac{2}{3} \bar{\gamma}^{ij}\partial_j K +
\bar{A}^{jk} \Delta\Gamma^i_{jk} + \bar{A}^{ik} \Delta\Gamma^j_{jk}\right)
$$
Notice the momentum constraint as written in [Baumgarte *et al.*](https://arxiv.org/pdf/1211.6632.pdf) is missing a term, as described in [Ruchlin, Etienne, & Baumgarte](https://arxiv.org/pdf/1712.07658.pdf).
<a id='gammaconstraint'></a>
# Step 4: The BSSN algebraic constraint: $\hat{\gamma}=\bar{\gamma}$ \[Back to [top](#toc)\]
$$\label{gammaconstraint}$$
[Brown](https://arxiv.org/abs/0902.3652)'s covariant Lagrangian formulation of BSSN, which we adopt, requires that $\partial_t \bar{\gamma} = 0$, where $\bar{\gamma}=\det \bar{\gamma}_{ij}$. We generally choose to set $\bar{\gamma}=\hat{\gamma}$ in our initial data.
Numerical errors will cause $\bar{\gamma}$ to deviate from a constant in time. This actually disrupts the hyperbolicity of the PDEs (causing crashes), so to cure this, we adjust $\bar{\gamma}_{ij}$ at the end of each Runge-Kutta timestep, so that its determinant satisfies $\bar{\gamma}=\hat{\gamma}$ at all times. We adopt the following, rather standard prescription (Eq. 53 of [Ruchlin, Etienne, and Baumgarte (2018)](https://arxiv.org/abs/1712.07658)):
$$
\bar{\gamma}_{ij} \to \left(\frac{\hat{\gamma}}{\bar{\gamma}}\right)^{1/3} \bar{\gamma}_{ij}.
$$
Notice the expression on the right is guaranteed to have determinant equal to $\hat{\gamma}$.
<a id='latex_pdf_output'></a>
# Step 5: Output this notebook to $\LaTeX$-formatted PDF file \[Back to [top](#toc)\]
$$\label{latex_pdf_output}$$
The following code cell converts this Jupyter notebook into a proper, clickable $\LaTeX$-formatted PDF file. After the cell is successfully run, the generated PDF may be found in the root NRPy+ tutorial directory, with filename
[Tutorial-BSSN_formulation.pdf](Tutorial-BSSN_formulation.pdf) (Note that clicking on this link may not work; you may need to open the PDF file through another means.)
```
import cmdline_helper as cmd # NRPy+: Multi-platform Python command-line interface
cmd.output_Jupyter_notebook_to_LaTeXed_PDF("Tutorial-BSSN_formulation")
```
| github_jupyter |
##### Copyright 2019 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Save and serialize models with Keras
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/guide/keras/save_and_serialize"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/guide/keras/save_and_serialize.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/guide/keras/save_and_serialize.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/guide/keras/save_and_serialize.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
</td>
</table>
The first part of this guide covers saving and serialization for Sequential models and models built using the Functional API and for Sequential models. The saving and serialization APIs are the exact same for both of these types of models.
Saving for custom subclasses of `Model` is covered in the section "Saving Subclassed Models". The APIs in this case are slightly different than for Sequential or Functional models.
## Setup
```
from __future__ import absolute_import, division, print_function, unicode_literals
try:
# %tensorflow_version only exists in Colab.
%tensorflow_version 2.x
except Exception:
pass
import tensorflow as tf
tf.keras.backend.clear_session() # For easy reset of notebook state.
```
## Part I: Saving Sequential models or Functional models
Let's consider the following model:
```
from tensorflow import keras
from tensorflow.keras import layers
inputs = keras.Input(shape=(784,), name='digits')
x = layers.Dense(64, activation='relu', name='dense_1')(inputs)
x = layers.Dense(64, activation='relu', name='dense_2')(x)
outputs = layers.Dense(10, name='predictions')(x)
model = keras.Model(inputs=inputs, outputs=outputs, name='3_layer_mlp')
model.summary()
```
Optionally, let's train this model, just so it has weight values to save, as well as an optimizer state.
Of course, you can save models you've never trained, too, but obviously that's less interesting.
```
(x_train, y_train), (x_test, y_test) = keras.datasets.mnist.load_data()
x_train = x_train.reshape(60000, 784).astype('float32') / 255
x_test = x_test.reshape(10000, 784).astype('float32') / 255
model.compile(loss=keras.losses.SparseCategoricalCrossentropy(from_logits=True),
optimizer=keras.optimizers.RMSprop())
history = model.fit(x_train, y_train,
batch_size=64,
epochs=1)
# Save predictions for future checks
predictions = model.predict(x_test)
```
### Whole-model saving
You can save a model built with the Functional API into a single file. You can later recreate the same model from this file, even if you no longer have access to the code that created the model.
This file includes:
- The model's architecture
- The model's weight values (which were learned during training)
- The model's training config (what you passed to `compile`), if any
- The optimizer and its state, if any (this enables you to restart training where you left off)
```
# Save the model
model.save('path_to_my_model.h5')
# Recreate the exact same model purely from the file
new_model = keras.models.load_model('path_to_my_model.h5')
import numpy as np
# Check that the state is preserved
new_predictions = new_model.predict(x_test)
np.testing.assert_allclose(predictions, new_predictions, rtol=1e-6, atol=1e-6)
# Note that the optimizer state is preserved as well:
# you can resume training where you left off.
```
### Export to SavedModel
You can also export a whole model to the TensorFlow `SavedModel` format. `SavedModel` is a standalone serialization format for TensorFlow objects, supported by TensorFlow serving as well as TensorFlow implementations other than Python.
```
# Export the model to a SavedModel
model.save('path_to_saved_model', save_format='tf')
# Recreate the exact same model
new_model = keras.models.load_model('path_to_saved_model')
# Check that the state is preserved
new_predictions = new_model.predict(x_test)
np.testing.assert_allclose(predictions, new_predictions, rtol=1e-6, atol=1e-6)
# Note that the optimizer state is preserved as well:
# you can resume training where you left off.
```
The `SavedModel` files that were created contain:
- A TensorFlow checkpoint containing the model weights.
- A `SavedModel` proto containing the underlying TensorFlow graph.
### Architecture-only saving
Sometimes, you are only interested in the architecture of the model, and you don't need to save the weight values or the optimizer. In this case, you can retrieve the "config" of the model via the `get_config()` method. The config is a Python dict that enables you to recreate the same model -- initialized from scratch, without any of the information learned previously during training.
```
config = model.get_config()
reinitialized_model = keras.Model.from_config(config)
# Note that the model state is not preserved! We only saved the architecture.
new_predictions = reinitialized_model.predict(x_test)
assert abs(np.sum(predictions - new_predictions)) > 0.
```
You can alternatively use `to_json()` from `from_json()`, which uses a JSON string to store the config instead of a Python dict. This is useful to save the config to disk.
```
json_config = model.to_json()
reinitialized_model = keras.models.model_from_json(json_config)
```
### Weights-only saving
Sometimes, you are only interested in the state of the model -- its weights values -- and not in the architecture. In this case, you can retrieve the weights values as a list of Numpy arrays via `get_weights()`, and set the state of the model via `set_weights`:
```
weights = model.get_weights() # Retrieves the state of the model.
model.set_weights(weights) # Sets the state of the model.
```
You can combine `get_config()`/`from_config()` and `get_weights()`/`set_weights()` to recreate your model in the same state. However, unlike `model.save()`, this will not include the training config and the optimizer. You would have to call `compile()` again before using the model for training.
```
config = model.get_config()
weights = model.get_weights()
new_model = keras.Model.from_config(config)
new_model.set_weights(weights)
# Check that the state is preserved
new_predictions = new_model.predict(x_test)
np.testing.assert_allclose(predictions, new_predictions, rtol=1e-6, atol=1e-6)
# Note that the optimizer was not preserved,
# so the model should be compiled anew before training
# (and the optimizer will start from a blank state).
```
The save-to-disk alternative to `get_weights()` and `set_weights(weights)`
is `save_weights(fpath)` and `load_weights(fpath)`.
Here's an example that saves to disk:
```
# Save JSON config to disk
json_config = model.to_json()
with open('model_config.json', 'w') as json_file:
json_file.write(json_config)
# Save weights to disk
model.save_weights('path_to_my_weights.h5')
# Reload the model from the 2 files we saved
with open('model_config.json') as json_file:
json_config = json_file.read()
new_model = keras.models.model_from_json(json_config)
new_model.load_weights('path_to_my_weights.h5')
# Check that the state is preserved
new_predictions = new_model.predict(x_test)
np.testing.assert_allclose(predictions, new_predictions, rtol=1e-6, atol=1e-6)
# Note that the optimizer was not preserved.
```
But remember that the simplest, recommended way is just this:
```
model.save('path_to_my_model.h5')
del model
model = keras.models.load_model('path_to_my_model.h5')
```
### Weights-only saving using TensorFlow checkpoints
Note that `save_weights` can create files either in the Keras HDF5 format,
or in the [TensorFlow Checkpoint format](https://www.tensorflow.org/api_docs/python/tf/train/Checkpoint). The format is inferred from the file extension you provide: if it is ".h5" or ".keras", the framework uses the Keras HDF5 format. Anything else defaults to Checkpoint.
```
model.save_weights('path_to_my_tf_checkpoint')
```
For total explicitness, the format can be explicitly passed via the `save_format` argument, which can take the value "tf" or "h5":
```
model.save_weights('path_to_my_tf_checkpoint', save_format='tf')
```
## Saving Subclassed Models
Sequential models and Functional models are datastructures that represent a DAG of layers. As such,
they can be safely serialized and deserialized.
A subclassed model differs in that it's not a datastructure, it's a piece of code. The architecture of the model
is defined via the body of the `call` method. This means that the architecture of the model cannot be safely serialized. To load a model, you'll need to have access to the code that created it (the code of the model subclass). Alternatively, you could be serializing this code as bytecode (e.g. via pickling), but that's unsafe and generally not portable.
For more information about these differences, see the article ["What are Symbolic and Imperative APIs in TensorFlow 2.0?"](https://medium.com/tensorflow/what-are-symbolic-and-imperative-apis-in-tensorflow-2-0-dfccecb01021).
Let's consider the following subclassed model, which follows the same structure as the model from the first section:
```
class ThreeLayerMLP(keras.Model):
def __init__(self, name=None):
super(ThreeLayerMLP, self).__init__(name=name)
self.dense_1 = layers.Dense(64, activation='relu', name='dense_1')
self.dense_2 = layers.Dense(64, activation='relu', name='dense_2')
self.pred_layer = layers.Dense(10, name='predictions')
def call(self, inputs):
x = self.dense_1(inputs)
x = self.dense_2(x)
return self.pred_layer(x)
def get_model():
return ThreeLayerMLP(name='3_layer_mlp')
model = get_model()
```
First of all, *a subclassed model that has never been used cannot be saved*.
That's because a subclassed model needs to be called on some data in order to create its weights.
Until the model has been called, it does not know the shape and dtype of the input data it should be
expecting, and thus cannot create its weight variables. You may remember that in the Functional model from the first section, the shape and dtype of the inputs was specified in advance (via `keras.Input(...)`) -- that's why Functional models have a state as soon as they're instantiated.
Let's train the model, so as to give it a state:
```
(x_train, y_train), (x_test, y_test) = keras.datasets.mnist.load_data()
x_train = x_train.reshape(60000, 784).astype('float32') / 255
x_test = x_test.reshape(10000, 784).astype('float32') / 255
model.compile(loss=keras.losses.SparseCategoricalCrossentropy(from_logits=True),
optimizer=keras.optimizers.RMSprop())
history = model.fit(x_train, y_train,
batch_size=64,
epochs=1)
```
The recommended way to save a subclassed model is to use `save_weights` to create a TensorFlow SavedModel checkpoint, which will contain the value of all variables associated with the model:
- The layers' weights
- The optimizer's state
- Any variables associated with stateful model metrics (if any)
```
model.save_weights('path_to_my_weights', save_format='tf')
# Save predictions for future checks
predictions = model.predict(x_test)
# Also save the loss on the first batch
# to later assert that the optimizer state was preserved
first_batch_loss = model.train_on_batch(x_train[:64], y_train[:64])
```
To restore your model, you will need access to the code that created the model object.
Note that in order to restore the optimizer state and the state of any stateful metric, you should
compile the model (with the exact same arguments as before) and call it on some data before calling `load_weights`:
```
# Recreate the model
new_model = get_model()
new_model.compile(loss=keras.losses.SparseCategoricalCrossentropy(from_logits=True),
optimizer=keras.optimizers.RMSprop())
# This initializes the variables used by the optimizers,
# as well as any stateful metric variables
new_model.train_on_batch(x_train[:1], y_train[:1])
# Load the state of the old model
new_model.load_weights('path_to_my_weights')
# Check that the model state has been preserved
new_predictions = new_model.predict(x_test)
np.testing.assert_allclose(predictions, new_predictions, rtol=1e-6, atol=1e-6)
# The optimizer state is preserved as well,
# so you can resume training where you left off
new_first_batch_loss=new_model.train_on_batch(x_train[:64], y_train[:64])
assert first_batch_loss == new_first_batch_loss
```
You've reached the end of this guide! This covers everything you need to know about saving and serializing models with tf.keras in TensorFlow 2.0.
| github_jupyter |
# Pooling Layer
---
In this notebook, we add and visualize the output of a maxpooling layer in a CNN.
### Import the image
```
import cv2
import matplotlib.pyplot as plt
%matplotlib inline
# TODO: Feel free to try out your own images here by changing img_path
# to a file path to another image on your computer!
img_path = 'images/udacity_sdc.png'
# load color image
bgr_img = cv2.imread(img_path)
# convert to grayscale
gray_img = cv2.cvtColor(bgr_img, cv2.COLOR_BGR2GRAY)
# normalize, rescale entries to lie in [0,1]
gray_img = gray_img.astype("float32")/255
# plot image
plt.imshow(gray_img, cmap='gray')
plt.show()
```
### Define and visualize the filters
```
import numpy as np
## TODO: Feel free to modify the numbers here, to try out another filter!
filter_vals = np.array([[-1, -1, 1, 1], [-1, -1, 1, 1], [-1, -1, 1, 1], [-1, -1, 1, 1]])
print('Filter shape: ', filter_vals.shape)
# Defining four different filters,
# all of which are linear combinations of the `filter_vals` defined above
# define four filters
filter_1 = filter_vals
filter_2 = -filter_1
filter_3 = filter_1.T
filter_4 = -filter_3
filters = np.array([filter_1, filter_2, filter_3, filter_4])
# For an example, print out the values of filter 1
print('Filter 1: \n', filter_1)
```
### Define convolutional and pooling layers
Initialize a convolutional layer so that it contains all your created filters. Then add a maxpooling layer, [documented here](https://pytorch.org/docs/stable/nn.html#maxpool2d), with a kernel size of (4x4) so you can really see that the image resolution has been reduced after this step!
```
import torch
import torch.nn as nn
import torch.nn.functional as F
# define a neural network with a convolutional layer with four filters
# AND a pooling layer of size (4, 4)
class Net(nn.Module):
def __init__(self, weight):
super(Net, self).__init__()
# initializes the weights of the convolutional layer to be the weights of the 4 defined filters
k_height, k_width = weight.shape[2:]
# assumes there are 4 grayscale filters
self.conv = nn.Conv2d(1, 4, kernel_size=(k_height, k_width), bias=False)
self.conv.weight = torch.nn.Parameter(weight)
# define a pooling layer
self.pool = nn.MaxPool2d(4, 4)
def forward(self, x):
# calculates the output of a convolutional layer
# pre- and post-activation
conv_x = self.conv(x)
activated_x = F.relu(conv_x)
# applies pooling layer
pooled_x = self.pool(activated_x)
# returns all layers
return conv_x, activated_x, pooled_x
# instantiate the model and set the weights
weight = torch.from_numpy(filters).unsqueeze(1).type(torch.FloatTensor)
model = Net(weight)
# print out the layer in the network
print(model)
```
### Visualize the output of each filter
First, we'll define a helper function, `viz_layer` that takes in a specific layer and number of filters (optional argument), and displays the output of that layer once an image has been passed through.
```
# helper function for visualizing the output of a given layer
# default number of filters is 4
def viz_layer(layer, n_filters= 4):
fig = plt.figure(figsize=(20, 20))
for i in range(n_filters):
ax = fig.add_subplot(1, n_filters, i+1, xticks=[], yticks=[])
# grab layer outputs
ax.imshow(np.squeeze(layer[0,i].data.numpy()), cmap='gray')
ax.set_title('Output %s' % str(i+1))
```
Let's look at the output of a convolutional layer after a ReLu activation function is applied.
```
# plot original image
plt.imshow(gray_img, cmap='gray')
# visualize all filters
fig = plt.figure(figsize=(12, 6))
fig.subplots_adjust(left=0, right=1.5, bottom=0.8, top=1, hspace=0.05, wspace=0.05)
for i in range(4):
ax = fig.add_subplot(1, 4, i+1, xticks=[], yticks=[])
ax.imshow(filters[i], cmap='gray')
ax.set_title('Filter %s' % str(i+1))
# convert the image into an input Tensor
gray_img_tensor = torch.from_numpy(gray_img).unsqueeze(0).unsqueeze(1)
# get all the layers
conv_layer, activated_layer, pooled_layer = model(gray_img_tensor)
# visualize the output of the activated conv layer
viz_layer(activated_layer)
```
### Visualize the output of the pooling layer
Then, take a look at the output of a pooling layer. The pooling layer takes as input the feature maps pictured above and reduces the dimensionality of those maps, by some pooling factor, by constructing a new, smaller image of only the maximum (brightest) values in a given kernel area.
```
# visualize the output of the pooling layer
viz_layer(pooled_layer)
```
| github_jupyter |
# MP2
## Some useful resources:
- [original paper](https://journals.aps.org/pr/abstract/10.1103/PhysRev.46.618)
- Levine Chapter 16
- [psi4numpy tutorial](https://github.com/psi4/psi4numpy/blob/master/Tutorials/05_Moller-Plesset/5a_conventional-mp2.ipynb)
- [Crawdad programming notes](http://sirius.chem.vt.edu/wiki/doku.php?id=crawdad:programming:project4)
# MP2 algorithm
1. The starting point will be the Hartree-Fock wavefunction
## Imports
```
import numpy as np
import scipy.linalg as spla
import psi4
import matplotlib.pyplot as plt
import time
%matplotlib notebook
```
## Specify the molecule
```
# start timer
start_time = time.time()
# define molecule
mol = psi4.geometry("""
O 0.0000000 0.0000000 0.0000000
H 0.7569685 0.0000000 -0.5858752
H -0.7569685 0.0000000 -0.5858752
symmetry c1
""")
psi4.set_options({'basis': 'sto-3g'})
wfn = psi4.core.Wavefunction.build(mol, psi4.core.get_global_option('BASIS'))
mints = psi4.core.MintsHelper(wfn.basisset())
# get number of electrons
num_elec_alpha = wfn.nalpha()
num_elec_beta = wfn.nbeta()
num_elec = num_elec_alpha + num_elec_beta
# get nuclear repulsion energy
E_nuc = mol.nuclear_repulsion_energy()
```
# STEP 2 : Calculate molecular integrals
Overlap
$$ S_{\mu\nu} = (\mu|\nu) = \int dr \phi^*_{\mu}(r) \phi_{\nu}(r) $$
Kinetic
$$ T_{\mu\nu} = (\mu\left|-\frac{\nabla}{2}\right|\nu) = \int dr \phi^*_{\mu}(r) \left(-\frac{\nabla}{2}\right) \phi_{\nu}(r) $$
Nuclear Attraction
$$ V_{\mu\nu} = (\mu|r^{-1}|\nu) = \int dr \phi^*_{\mu}(r) r^{-1} \phi_{\nu}(r) $$
Form Core Hamiltonian
$$ H = T + V $$
Two electron integrals
$$ (\mu\nu|\lambda\sigma) = \int dr_1 dr_2 \phi^*_{\mu}(r_1) \phi_{\nu}(r_1) r_{12}^{-1} \phi_{\lambda}(r_2) \phi_{\sigma}(r_2) $$
```
# calculate overlap integrals
S = np.asarray(mints.ao_overlap())
# calculate kinetic energy integrals
T = np.asarray(mints.ao_kinetic())
# calculate nuclear attraction integrals
V = np.asarray(mints.ao_potential())
# form core Hamiltonian
H = T + V
# calculate two electron integrals
eri = np.asarray(mints.ao_eri())
# get number of atomic orbitals
num_ao = np.shape(S)[0]
print(np.shape(eri))
# set inital density matrix to zero
D = np.zeros((num_ao,num_ao))
# 2 helper functions for printing during SCF
def print_start_iterations():
print("{:^79}".format("{:>4} {:>11} {:>11} {:>11} {:>11}".format("Iter", "Time(s)", "RMSC DM", "delta E", "E_scf_elec")))
print("{:^79}".format("{:>4} {:>11} {:>11} {:>11} {:>11}".format("****", "*******", "*******", "*******", "******")))
def print_iteration(iteration_num, iteration_start_time, iteration_end_time, iteration_rmsc_dm, iteration_E_diff, E_scf_elec):
print("{:^79}".format("{:>4d} {:>11f} {:>.5E} {:>.5E} {:>11f}".format(iteration_num, iteration_end_time - iteration_start_time, iteration_rmsc_dm, iteration_E_diff, E_scf_elec)))
# set stopping criteria
iteration_max = 100
convergence_E = 1e-9
convergence_DM = 1e-5
# loop variables
iteration_num = 0
E_scf_total = 0
E_scf_elec = 0.0
iteration_E_diff = 0.0
iteration_rmsc_dm = 0.0
converged = False
exceeded_iterations = False
print_start_iterations()
while (not converged and not exceeded_iterations):
# store last iteration and increment counters
iteration_start_time = time.time()
iteration_num += 1
E_elec_last = E_scf_elec
D_last = np.copy(D)
# form G matrix
G = np.zeros((num_ao,num_ao))
for i in range(num_ao):
for j in range(num_ao):
for k in range(num_ao):
for l in range(num_ao):
G[i,j] += D[k,l] * ((2.0*(eri[i,j,k,l])) - (eri[i,k,j,l]))
# build fock matrix
F = H + G
# solve the generalized eigenvalue problem
E_orbitals, C = spla.eigh(F,S)
# compute new density matrix
D = np.zeros((num_ao,num_ao))
for i in range(num_ao):
for j in range(num_ao):
for k in range(num_elec_alpha):
D[i,j] += C[i,k] * C[j,k]
# calculate electronic energy
E_scf_elec = np.sum(np.multiply(D , (H + F)))
# calculate energy change of iteration
iteration_E_diff = np.abs(E_scf_elec - E_elec_last)
# rms change of density matrix
iteration_rmsc_dm = np.sqrt(np.sum((D - D_last)**2))
iteration_end_time = time.time()
print_iteration(iteration_num, iteration_start_time, iteration_end_time, iteration_rmsc_dm, iteration_E_diff, E_scf_elec)
if(np.abs(iteration_E_diff) < convergence_E and iteration_rmsc_dm < convergence_DM):
converged = True
if(iteration_num == iteration_max):
exceeded_iterations = True
# calculate total energy
E_scf_total = E_scf_elec + E_nuc
print("{:^79}".format("Total HF energy : {:>11f}".format(E_scf_total)))
```
# Perform MP2 calculation
## Convert the two-electron integrals from AO basis to the MO basis
$$(pq|rs) = \sum_\mu \sum_\nu \sum_\lambda \sum_\sigma C_\mu^p C_\nu^q
(\mu \nu|\lambda \sigma) C_\lambda^r C_\sigma^s.$$
Attempt to code this conversion below, remember that the electron repulsion integrals above are stored as vector `eri` that is of the shape (num_ao,num_ao,num_ao,num_ao). Here the num_ao's for sto-3g water is 7. The resulting tensor will have the same shape as `eri`.
```
## place code for two-electron integral conversion here.
```
### Compute the MP2 Energy
Now we can calculate the MP2 estimation of the correlation energy.
$$E_{\mathrm{corr(MP2)}}\ =\ \frac{( ia \mid jb ) [ 2 (ia \mid jb ) - ( ib \mid ja )]}{\epsilon_i + \epsilon_j + \epsilon_a - \epsilon_b}$$
Here $i$ and $j$ represent all occupied orbitals, where as $a$ and $b$ will be unoccupied orbitals.
Remember during this coding step that we are basing our MP2 correction on an RHF calculation and thus there are the same amount of $\alpha$ and $\beta$ electrons.
```
#initialize the variable forthe mp2 correlation energy
E_corr_mp2 = 0
# code the equation above and adjust the value of E_corr_mp2
#this will print your E_corr mp2
print("{:^79}".format("Total MP2 correlation energy : {:>11f}".format(E_corr_mp2)))
```
The correlation energy should be very small compared to the total energy (-0.035493 Ha), which is generally the case. However, this correlation energy can be very important to describing properties such as dispersion.
## A comparison with Psi4
```
# Get the SCF wavefunction & energies# Get t
scf_e, scf_wfn = psi4.energy('scf', return_wfn=True)
mp2_e = psi4.energy('mp2')
print(mp2_e)
E_diff = (mp2_e - (E_total + E_corr_mp2))
print(E_diff)
```
| github_jupyter |
```
from analysis import *
from docking_analysis import *
from IPython.display import Markdown
task_name = 'Keck_Pria_FP'
file_paths = {0: 'single_classification_22/45710870/',
1: 'single_classification_42/45710871/',
2: 'single_regression_2/45710874/',
3: 'single_regression_11/45710875/',
4: 'vanilla_lstm_8/46547695/',
5: 'vanilla_lstm_19/46547696/',
6: 'multi_classification_15/eval_keck/',
7: 'multi_classification_18/eval_keck/',
8: 'sklearn_rf_390014_96/',
9: 'sklearn_rf_390014_97/'}
file_paths = {0: 'single_classification_22/45710870/',
1: 'single_regression_2/45710874/',
2: 'vanilla_lstm_19/46547696/',
3: 'multi_classification_15/eval_keck/',
4: 'random_forest_97/',
5: 'irv_80/'}
number = 20
evaluations = {0: 'train prec', 1: 'train roc', 2: 'train bedroc',
3: 'val prec', 4: 'val roc', 5: 'val bedroc',
6: 'test prec', 7: 'test roc', 8: 'test bedroc',
9: 'EF_2', 10: 'EF_1', 11: 'EF_015', 12: 'EF_01',
13: 'NEF_2', 14: 'NEF_1', 15: 'NEF_015', 16:'NEF_01'}
paths = ['../../output/stage_1/cross_validation_Keck_FP/{}'.format(p) for p in file_paths.values()]
model_list = ['single_classification',
'single_regression',
'vanilla_lstm',
'multi_classification',
'random_forest',
'irv']
%matplotlib inline
import matplotlib
matplotlib.rcParams['figure.figsize'] = (20.0, 10.0)
%load_ext autoreload
%autoreload 2
for k in range(len(file_paths)):
print 'Check ', file_paths[k]
check_result_completeness(file_path=paths[k], number=number)
print
plot_cross_validation(paths,
evaluation_list=[evaluations[0], evaluations[3], evaluations[6]],
model_list=model_list,
title='AUC[PR]',
task_name=task_name)
mean_content, median_content, std_content = table_cross_validation(paths,
evaluation_list=[evaluations[0], evaluations[3], evaluations[6]],
model_list=model_list,
evaluation_mode='AUC[PR]')
content = '{}\n{}\n{}'.format(mean_content, median_content, std_content)
Markdown(content)
plot_cross_validation(paths,
evaluation_list=[evaluations[1], evaluations[4], evaluations[7]],
model_list=model_list,
title='AUC[ROC]',
task_name=task_name)
mean_content, median_content, std_content = table_cross_validation(paths,
evaluation_list=[evaluations[1], evaluations[4], evaluations[7]],
model_list=model_list,
evaluation_mode='AUC[ROC]')
content = '{}\n{}\n{}'.format(mean_content, median_content, std_content)
Markdown(content)
plot_cross_validation(paths,
evaluation_list=[evaluations[2], evaluations[5], evaluations[8]],
model_list=model_list,
title='AUC[BED ROC]',
task_name=task_name)
mean_content, median_content, std_content = table_cross_validation(paths,
evaluation_list=[evaluations[2], evaluations[5], evaluations[8]],
model_list=model_list,
evaluation_mode='AUC[BED ROC]')
content = '{}\n{}\n{}'.format(mean_content, median_content, std_content)
Markdown(content)
plot_cross_validation(paths,
evaluation_list=[evaluations[9], evaluations[10], evaluations[11], evaluations[12]],
model_list=model_list,
title='Enrichment Factor on Test Set',
task_name=task_name)
mean_content, median_content, std_content = table_cross_validation(paths,
evaluation_list=[evaluations[9], evaluations[10], evaluations[11], evaluations[12]],
model_list=model_list,
evaluation_mode='Enrichment Factor (Test-set)')
content = '{}\n{}\n{}'.format(mean_content, median_content, std_content)
Markdown(content)
plot_cross_validation(paths,
evaluation_list=[evaluations[13], evaluations[14], evaluations[15], evaluations[16]],
model_list=model_list,
title='Normalized Enrichment Factor on Test Set',
task_name=task_name)
mean_content, median_content, std_content = table_cross_validation(paths,
evaluation_list=[evaluations[13], evaluations[14], evaluations[15], evaluations[16]],
model_list=model_list,
evaluation_mode='Normalized Enrichment Factor (Test-set)')
content = '{}\n{}\n{}'.format(mean_content, median_content, std_content)
Markdown(content)
```
# Docking Results
```
content = get_auc_table(file_path='../../output/docking/stage_1/lc123_all_docking_scores.csv',
target_name='Keck_Pria_FP_data',
auc_list=['precision_auc_single', 'roc_auc_single', 'bedroc_auc_single'],
auc_header=['AUC[PR]', 'AUC[ROC]', 'AUC[BED ROC]'],
title='AUC for Docking Methods')
Markdown(content)
EF_ratio_list = [0.02, 0.01, 0.0015, 0.001]
content = get_ef_table(file_path='../../output/docking/stage_1/lc123_all_docking_scores.csv',
target_name='Keck_Pria_FP_data',
efr_list=EF_ratio_list,
ef_header=['EF_2', 'EF_1', 'EF_015', 'EF_01'],
title='Enrichment Factor for Docking Methods')
Markdown(content)
EF_ratio_list = [0.02, 0.01, 0.0015, 0.001]
content = get_nef_table(file_path='../../output/docking/stage_1/lc123_all_docking_scores.csv',
target_name='Keck_Pria_FP_data',
nefr_list=EF_ratio_list,
nef_header=['NEF_2', 'NEF_1', 'NEF_015', 'NEF_01'],
title='Normalized Enrichment Factor for Docking Methods')
Markdown(content)
```
# Plot EF curve
```
from EF_curve_support_loader import *
%matplotlib inline
import matplotlib
matplotlib.rcParams['figure.figsize'] = (20.0, 30.0)
predictions_path = '../../output/stage_1_predictions/cross_validation_Keck_FP'
model_names_list = ['single_classification_22',
'single_regression_2',
'multi_classification_15',
'sklearn_rf_390014_97',
'deepchem_irv_80']
EF_ratio_list = np.linspace(0.001, 0.15, 100)
N = len(model_names_list)
data_set_name = 'Keck_Pria_FP_data'
data_pd_list = []
for i in range(N):
temp_pd = get_EF_curve_in_pd(EF_ratio_list=EF_ratio_list,
data_set_name=data_set_name,
model_name=model_names_list[i],
predictions_path=predictions_path,
regenerate=True)
data_pd_list.append(temp_pd)
whole_pd = data_pd_list[0]
for i in range(1, N):
whole_pd = whole_pd.append(data_pd_list[i])
%matplotlib inline
import matplotlib
matplotlib.rcParams['figure.figsize'] = (20.0, 30.0)
sns.tsplot(data=whole_pd, time="EFR", unit="running process", condition="model", value="EF")
figure_dir = 'plottings/{}'.format(task_name)
if not os.path.isdir(figure_dir):
os.makedirs(figure_dir)
plt.savefig('{}/EF_curve.png'.format(figure_dir), bbox_inches = 'tight')
```
| github_jupyter |
<img src="http://hilpisch.com/tpq_logo.png" alt="The Python Quants" width="35%" align="right" border="0"><br>
# Derivatives Analytics with Python
**_Chapters 12 & 13_**
**Wiley Finance (2015)**
<img src="http://hilpisch.com/images/derivatives_analytics_front.jpg" alt="Derivatives Analytics with Python" width="30%" align="left" border="0">
Yves Hilpisch
The Python Quants GmbH
<a href='mailto:analytics@pythonquants.com'>analytics@pythonquants.com</a>
<a href='http://pythonquants.com'>www.pythonquants.com</a>
```
import matplotlib.pyplot as plt
%matplotlib inline
```
## Chapter 12: Valuation
```
%run 12_val/BCC97_simulation.py
np.array((S0, r0, kappa_r, theta_r, sigma_r))
opt
```
### Simulation
```
plot_rate_paths(r)
plt.savefig('../images/12_val/rate_paths.pdf')
plot_volatility_paths(v)
plt.savefig('../images/12_val/volatility_paths.pdf')
plot_index_paths(S)
plt.savefig('../images/12_val/ES50_paths.pdf')
plot_index_histogram(S)
plt.savefig('../images/12_val/ES50_histogram.pdf')
```
### European Options Valuation
```
%run 12_val/BCC97_valuation_comparison.py
%time compare_values(M0=50, I=50000)
%time compare_values(M0=50, I=200000)
%time compare_values(M0=200, I=50000)
```
### American Options Valuation
```
%run 12_val/BCC97_american_valuation.py
%time lsm_compare_values(M0=150, I=50000)
```
## Chapter 13: Dynamic Hedging
### BSM Model
```
%run 13_dyh/BSM_lsm_hedging_algorithm.py
S, po, vt, errs, t = BSM_hedge_run(p=25)
plot_hedge_path(S, po, vt, errs, t)
# plt.savefig('../images/13_dyh/BSM_hedge_run_1.pdf')
S, po, vt, errs, t = BSM_hedge_run(p=5)
plot_hedge_path(S, po, vt, errs, t)
# plt.savefig('../images/13_dyh/BSM_hedge_run_2.pdf')
%run 13_dyh/BSM_lsm_hedging_histogram.py
%time pl_list = BSM_dynamic_hedge_mcs()
plot_hedge_histogram(pl_list)
plt.savefig('../images/13_dyh/BSM_hedge_histo.pdf')
%time pl_list = BSM_dynamic_hedge_mcs(M=200, I=150000)
plot_hedge_histogram(pl_list)
# plt.savefig('../images/13_dyh/BSM_hedge_histo_more.pdf')
```
### BCC97 Model
```
%run 13_dyh/BCC97_lsm_hedging_algorithm.py
S, po, vt, errs, t = BCC97_hedge_run(2)
plot_hedge_path(S, po, vt, errs, t)
plt.savefig('../images/13_dyh/BCC_hedge_run_1.pdf')
S, po, vt, errs, t = BCC97_hedge_run(8)
plot_hedge_path(S, po, vt, errs, t + 1)
plt.savefig('../images/13_dyh/BCC_hedge_run_2.pdf')
S, po, vt, errs, t = BCC97_hedge_run(4)
plot_hedge_path(S, po, vt, errs, t + 1)
plt.savefig('../images/13_dyh/BCC_hedge_run_3.pdf')
%run 13_dyh/BCC97_lsm_hedging_histogram.py
%time pl_list = BCC97_hedge_simulation(M=150, I=150000)
plot_hedge_histogram(pl_list)
plt.savefig('../images/13_dyh/BCC_hedge_histo.pdf')
```
<img src="http://hilpisch.com/tpq_logo.png" alt="The Python Quants" width="35%" align="right" border="0"><br>
<a href="http://www.pythonquants.com" target="_blank">www.pythonquants.com</a> | <a href="http://twitter.com/dyjh" target="_blank">@dyjh</a>
<a href="mailto:analytics@pythonquants.com">analytics@pythonquants.com</a>
**Python Quant Platform** |
<a href="http://quant-platform.com">http://quant-platform.com</a>
**Derivatives Analytics with Python** |
<a href="http://www.derivatives-analytics-with-python.com" target="_blank">Derivatives Analytics @ Wiley Finance</a>
**Python for Finance** |
<a href="http://python-for-finance.com" target="_blank">Python for Finance @ O'Reilly</a>
| github_jupyter |
# A simple parameter exploration
This notebook demonstrates a very simple parameter exploration of a custom function that we have defined. It is a simple function that returns the distance to a unit circle, so we expect our parameter exploration to resemble a circle.
```
# change to the root directory of the project
import os
if os.getcwd().split("/")[-1] == "examples":
os.chdir('..')
# This will reload all imports as soon as the code changes
%load_ext autoreload
%autoreload 2
try:
import matplotlib.pyplot as plt
except ImportError:
import sys
!{sys.executable} -m pip install matplotlib
import matplotlib.pyplot as plt
import numpy as np
from neurolib.utils.parameterSpace import ParameterSpace
from neurolib.optimize.exploration import BoxSearch
```
## Define the evaluation function
Here we define a very simple evaluation function. The function needs to take in `traj` as an argument, which is the pypet trajectory. This is how the function knows what parameters were assigned to it. Using the builtin function `search.getParametersFromTraj(traj)` we can then retrieve the parameters for this run. They are returned as a dictionary and can be accessed in the function.
In the last step, we use `search.saveToPypet(result_dict, traj)` to save the results to the pypet trajectory and to an HDF. In between, the computational magic happens!
```
def explore_me(traj):
pars = search.getParametersFromTraj(traj)
# let's calculate the distance to a circle
computation_result = abs((pars['x']**2 + pars['y']**2) - 1)
result_dict = {"distance" : computation_result}
search.saveToPypet(result_dict, traj)
```
## Define the parameter space and exploration
Here we define which space we want to cover. For this, we use the builtin class `ParameterSpace` which provides a very easy interface to the exploration. To initialize the exploration, we simply pass the evaluation function and the parameter space to the `BoxSearch` class.
```
parameters = ParameterSpace({"x": np.linspace(-2, 2, 2), "y": np.linspace(-2, 2, 2)})
# info: chose np.linspace(-2, 2, 40) or more, values here are low for testing
search = BoxSearch(evalFunction = explore_me, parameterSpace = parameters, filename="example-1.1.hdf")
```
## Run
And off we go!
```
search.run()
```
## Get results
We can easily obtain the results from pypet. First we call `search.loadResults()` to make sure that the results are loaded from the hdf file to our instance.
```
search.loadResults()
print("Number of results: {}".format(len(search.results)))
```
The runs are also ordered in a simple pandas dataframe called `search.dfResults`. We cycle through all results by calling `search.results[i]` and loading the desired result (here the distance to the circle) into the dataframe
```
for i in search.dfResults.index:
search.dfResults.loc[i, 'distance'] = search.results[i]['distance']
search.dfResults
```
And of course a plot can visualize the results very easily.
```
plt.imshow(search.dfResults.pivot_table(values='distance', index = 'x', columns='y'), \
extent = [min(search.dfResults.x), max(search.dfResults.x),
min(search.dfResults.y), max(search.dfResults.y)], origin='lower')
plt.colorbar(label='Distance to the unit circle')
```
| github_jupyter |
# AutoEncoders
### In this notebook you will learn the definition of an autoencoder, how it works, and see an implementation in TensorFlow.
---
### Table of Contents
1. Introduction
2. Feature Extraction and Dimensionality Reduction
3. Autoencoder Structure
4. Performance
5. Training: Loss Function
6. Code
---
### By the end of this notebook, you should be able to create simple autoencoders apply them to problems in the field of unsupervised learning.
---
## Introduction
---
An autoencoder, also known as autoassociator or Diabolo networks, is an artificial neural network employed to recreate the given input.
It takes a set of <b>unlabeled</b> inputs, encodes them and then tries to extract the most valuable information from them.
They are used for feature extraction, learning generative models of data, dimensionality reduction and can be used for compression.
A 2006 paper named *[Reducing the Dimensionality of Data with Neural Networks](https://www.cs.toronto.edu/~hinton/science.pdf)*, done by G. E. Hinton and R. R. Salakhutdinov</b>, showed better results than years of refining other types of network, and was a breakthrough in the field of Neural Networks, a field that was "stagnant" for 10 years.
Now, autoencoders, based on Restricted Boltzmann Machines, are employed in some of the largest deep learning applications. They are the building blocks of Deep Belief Networks (DBN).
<p align="center">
<img src="../images/DBNpic.png">
</p>
---
## Feature Extraction and Dimensionality Reduction
---
An example given by Nikhil Buduma in KdNuggets *[(link)]("http://www.kdnuggets.com/2015/03/deep-learning-curse-dimensionality-autoencoders.html")* gives an excellent explanation of the utility of this type of Neural Network.
Say that you want to extract the emotion that a person in a photograph is feeling. Take the following 256x256 pixel grayscale picture as an example:
<p align="center">
<img src="../images/examplePFP.png">
</p>
If we just use the raw image, we have too many dimensions to analyze. This image is 256x256 pixels, which corresponds to an input vector of 65536 dimensions! Conventional cell phones can produce images in the 4000 x 3000 pixels range, which gives us 12 million dimensions to analyze.
This is particularly problematic, since the difficulty of a machine learning problem is vastly increased as more dimensions are involved. According to a 1982 study by C.J. Stone *[(link)]("http://www-personal.umich.edu/~jizhu/jizhu/wuke/Stone-AoS82.pdf")*, the time to fit a model, is optimal if:
$ m^{-p/(2p+d)} $
Where:
* m: Number of data points
* d: Dimensionality of the data
* p: Number of Parameters in the model
As you can see, it increases exponentially!
Returning to our example, we don't need to use all of the 65,536 dimensions to classify an emotion.
A human identifies emotions according to specific facial expressions, and some <b>key features</b>, like the shape of the mouth and eyebrows.
<p align="center">
<img src="../images/examplePFPMarked.png" height="256" width="256">
</p>
---
## Autoencoder Structure
---
<p align="center">
<img src="../images/encoderDecoder.png" style="width: 400px;">
</p>
An autoencoder can be divided in two parts, the **encoder** and the **decoder**.
The *encoder* needs to compress the representation of an input. In this case, we are going to reduce the dimensions of the image of the example face from 2000 dimensions to only 30 dimensions. We will acomplish this by running the data through the layers of our encoder.
The *decoder* works like encoder network in reverse. It works to recreate the input as closely as possible. The training procedure produces at the center of the network a compressed, low dimensional representation that can be decoded to obtain the higher dimensional representation with minimal loss of information between the input and the output.
---
## Performance
---
After training has been completed, you can use the encoded data as a reliable low dimensional representation of the data. This can be applied to many problems where dimensionality reduction seems appropriate.
<img src="../images/dimensionalityPic.png">
This image was extracted from the G. E. Hinton and R. R. Salakhutdinovcomparing's <a href="https://www.cs.toronto.edu/~hinton/science.pdf">paper</a>, on the two-dimensional reduction for 500 digits of the MNIST, with PCA (Principal Component Analysis) on the left and autoencoder on the right. We can see that the autoencoder provided us with a better separation of data.
---
## Training: Loss function
---
An autoencoder uses the <b>Loss</b> function to properly train the network. The Loss function will calculate the differences between our output and the expected results. After that, we can minimize this error with gradient descent. There are many types of Loss functions, and it is important to consider the type of problem (classification, regression, etc.) when choosing this funtion.
### Binary Values:
$$L(W) = -\sum_{k} (x_k log(\hat{x}_k) + (1 - x_k) \log (1 - \hat{x}_k)) $$
For binary values, we can use an equation based on the sum of Bernoulli's cross-entropy. This loss function is best for binary classification problems.
$x_k$ is one of our inputs and $\hat{x}\_k $ is the respective output. Note that:
$$\hat{x} = f(x,W) $$
Where $W$ is the full parameter set of the neural network.
We use this function so that when $x_k=1$, we want the calculated value of $\hat{x}_k $ to be very close to one, and likewise if $x_k=0$.
If the value is one, we just need to calculate the first part of the formula, that is, $-x_klog(\hat{x}_k)$. Which, turns out to just calculate $-log(\hat{x}_k)$.
We explicitly exclude the second term to avoid numerical difficulties when computing the logarithm of very small numbers.
Likewise, if the value is zero, we need to calculate just the second part, $(1 - x_k)log(1 - \hat{x}_k)$, which turns out to be $log(1 - \hat{x}_k)$.
### Real values:
$$L(W) = - \frac{1}{2}\sum_{k} (\hat{x}_k- x_k \ )^2$$
For data where the value (not category) is important to reproduce, we can use the sum of squared errors (SSE) for our *Loss* function. This function is usually used in regressions.
As it was with the above example, $x_k$ is one of our inputs and $\hat{x}_k$ is the respective output, and we want to make our output as similar as possible to our input.
### Computing Gradient
The gradient of the loss function is an important and complex function. It is defined as:
$$\nabla_{W} L(W)_j = \frac{\partial f(x,W)}{\partial{W_j}}$$
Fortunately for us, TensorFlow computes these complex functions automatically when we define our functions that are used to compute loss! They automatically manage the backpropagation algorithm, which is an efficient way of computing the gradients in complex neural networks.
---
## Code
---
We are going to use the MNIST dataset for our example.
The following code was created by Aymeric Damien. You can find some of his code in <a href="https://github.com/aymericdamien">here</a>. We made some modifications which allow us to import the datasets to Jupyter Notebooks.
Let's call our imports and make the MNIST data available to use.
```
!pip install tensorflow==2.2.0rc0
```
Once tensorflow, numpy and matplotlib have been succesfully imported, the data set must be imported to the environment.
```
#from __future__ import division, print_function, absolute_import
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
if not tf.__version__ == '2.2.0-rc0':
print(tf.__version__)
raise ValueError('please upgrade to TensorFlow 2.2.0-rc0, or restart your Kernel (Kernel->Restart & Clear Output)')
```
Load the imported data using tensor flow (tf).
```
# Import MINST data
mnist = tf.keras.datasets.mnist
(x_train, y_train), (x_test, y_test) = mnist.load_data()
```
When training a neural network, it is popular practice to use 32 bit precision. The newly configured type is then divided by 255 because that is the maximum value of a byte. By doing so, we will get an output that is between 0.0 and 1.0, which normalizes the data and allows for it to be scaled.
```
x_train = x_train.astype('float32') / 255.
x_test = x_test.astype('float32') / 255.
y_train = y_train.astype('float32') / 255.
y_test = y_test.astype('float32') / 255.
```
Using -1 for the first dimension means that it is inferred using Tensor, based on the number of elements.
```
x_image_train = tf.reshape(x_train, [-1,28,28,1])
x_image_train = tf.cast(x_image_train, 'float32')
x_image_test = tf.reshape(x_test, [-1,28,28,1])
x_image_test = tf.cast(x_image_test, 'float32')
```
Print the newly trained shape.
```
print(x_train.shape)
```
We use the tf.keras.layers.Flatten() function to prepare the training data to be compatible with the encoding and decoding layer
```
flatten_layer = tf.keras.layers.Flatten()
x_train = flatten_layer(x_train)
```
Notice how the <code>x_train.shape</code> changes from (60000,28,28) to (60000, 784)
```
print(x_train.shape)
```
Now, let's give the parameters that are going to be used by our NN.
```
learning_rate = 0.01
training_epochs = 20
batch_size = 256
display_step = 1
examples_to_show = 10
global_step = tf.Variable(0)
total_batch = int(len(x_train) / batch_size)
# Network Parameters
n_hidden_1 = 256 # 1st layer num features
n_hidden_2 = 128 # 2nd layer num features
encoding_layer = 32 # final encoding bottleneck features
n_input = 784 # MNIST data input (img shape: 28*28)
```
### Encoder
---
Now we need to create our encoder. For this, we are going to use tf.keras.layers.Dense with sigmoidal activation functions. Sigmoidal functions delivers great results with this type of network. This is due to having a good derivative that is well-suited to backpropagation. We can create our encoder using the sigmoidal function like this:
```
enocoding_1 = tf.keras.layers.Dense(n_hidden_1, activation=tf.nn.sigmoid)
encoding_2 = tf.keras.layers.Dense(n_hidden_2, activation=tf.nn.sigmoid)
encoding_final = tf.keras.layers.Dense(encoding_layer, activation=tf.nn.relu)
# Building the encoder
def encoder(x):
x_reshaped = flatten_layer(x)
# Encoder first layer with sigmoid activation #1
layer_1 = enocoding_1(x_reshaped)
# Encoder second layer with sigmoid activation #2
layer_2 = encoding_2(layer_1)
code = encoding_final(layer_2)
return code
```
### Decoder
---
You can see that the layer_1 in the encoder is the layer_2 in the decoder and vice-versa.
```
decoding_1 = tf.keras.layers.Dense(n_hidden_2, activation=tf.nn.sigmoid)
decoding_2 = tf.keras.layers.Dense(n_hidden_1, activation=tf.nn.sigmoid)
decoding_final = tf.keras.layers.Dense(n_input)
# Building the decoder
def decoder(x):
# Decoder first layer with sigmoid activation #1
layer_1 = decoding_1(x)
# Decoder second layer with sigmoid activation #2
layer_2 = decoding_2(layer_1)
decode = self.decoding_final(layer_2)
return decode
```
### Model Building
---
Let's construct our model.
We define a <code>cost</code> function to calculate the loss and a <code>grad</code> function to calculate gradients that will be used in backpropagation.
```
class AutoEncoder(tf.keras.Model):
def __init__(self):
super(AutoEncoder, self).__init__()
self.n_hidden_1 = n_hidden_1 # 1st layer num features
self.n_hidden_2 = n_hidden_2 # 2nd layer num features
self.encoding_layer = encoding_layer
self.n_input = n_input # MNIST data input (img shape: 28*28)
self.flatten_layer = tf.keras.layers.Flatten()
self.enocoding_1 = tf.keras.layers.Dense(self.n_hidden_1, activation=tf.nn.sigmoid)
self.encoding_2 = tf.keras.layers.Dense(self.n_hidden_2, activation=tf.nn.sigmoid)
self.encoding_final = tf.keras.layers.Dense(self.encoding_layer, activation=tf.nn.relu)
self.decoding_1 = tf.keras.layers.Dense(self.n_hidden_2, activation=tf.nn.sigmoid)
self.decoding_2 = tf.keras.layers.Dense(self.n_hidden_1, activation=tf.nn.sigmoid)
self.decoding_final = tf.keras.layers.Dense(self.n_input)
# Building the encoder
def encoder(self,x):
#x = self.flatten_layer(x)
layer_1 = self.enocoding_1(x)
layer_2 = self.encoding_2(layer_1)
code = self.encoding_final(layer_2)
return code
# Building the decoder
def decoder(self, x):
layer_1 = self.decoding_1(x)
layer_2 = self.decoding_2(layer_1)
decode = self.decoding_final(layer_2)
return decode
def call(self, x):
encoder_op = self.encoder(x)
# Reconstructed Images
y_pred = self.decoder(encoder_op)
return y_pred
def cost(y_true, y_pred):
loss = tf.losses.mean_squared_error(y_true, y_pred)
cost = tf.reduce_mean(loss)
return cost
def grad(model, inputs, targets):
#print('shape of inputs : ',inputs.shape)
#targets = flatten_layer(targets)
with tf.GradientTape() as tape:
reconstruction = model(inputs)
loss_value = cost(targets, reconstruction)
return loss_value, tape.gradient(loss_value, model.trainable_variables),reconstruction
```
### Model Training
---
For training we will run for 20 epochs.
```
model = AutoEncoder()
optimizer = tf.keras.optimizers.RMSprop(learning_rate)
for epoch in range(training_epochs):
for i in range(total_batch):
x_inp = x_train[i : i + batch_size]
loss_value, grads, reconstruction = grad(model, x_inp, x_inp)
optimizer.apply_gradients(zip(grads, model.trainable_variables))
# Display logs per epoch step
if epoch % display_step == 0:
print("Epoch:", '%04d' % (epoch+1),
"cost=", "{:.9f}".format(loss_value))
print("Optimization Finished!")
```
### Model Testing
---
Now, let's apply encoder and decoder for our tests.
```
# Applying encode and decode over test set
encode_decode = model(flatten_layer(x_image_test[:examples_to_show]))
```
Let's simply visualize our graphs!
```
# Compare original images with their reconstructions
f, a = plt.subplots(2, 10, figsize=(10, 2))
for i in range(examples_to_show):
a[0][i].imshow(np.reshape(x_image_test[i], (28, 28)))
a[1][i].imshow(np.reshape(encode_decode[i], (28, 28)))
```
As you can see, the reconstructions were successful. It can be seen that some noise were added to the image.
<hr>
## Want to learn more?
Running deep learning programs usually needs a high performance platform. **PowerAI** speeds up deep learning and AI. Built on IBM’s Power Systems, **PowerAI** is a scalable software platform that accelerates deep learning and AI with blazing performance for individual users or enterprises. The **PowerAI** platform supports popular machine learning libraries and dependencies including TensorFlow, Caffe, Torch, and Theano. You can use [PowerAI on IMB Cloud](https://cocl.us/ML0120EN_PAI).
Also, you can use **Watson Studio** to run these notebooks faster with bigger datasets. **Watson Studio** is IBM’s leading cloud solution for data scientists, built by data scientists. With Jupyter notebooks, RStudio, Apache Spark and popular libraries pre-packaged in the cloud, **Watson Studio** enables data scientists to collaborate on their projects without having to install anything. Join the fast-growing community of **Watson Studio** users today with a free account at [Watson Studio](https://cocl.us/ML0120EN_DSX). This is the end of this lesson. Thank you for reading this notebook, and good luck!
### Thanks for completing this lesson!
Created by <a href="https://www.linkedin.com/in/franciscomagioli">Francisco Magioli</a>, <a href="https://ca.linkedin.com/in/erich-natsubori-sato">Erich Natsubori Sato</a>, <a href="https://ca.linkedin.com/in/saeedaghabozorgi">Saeed Aghabozorgi</a>
Updated to TF 2.X by <a href="https://www.linkedin.com/in/samaya-madhavan"> Samaya Madhavan </a>
Added to IBM Developer by [Dhivya Lakshminarayanan](https://www.linkedin.com/in/dhivya-lak/)
### References:
- [https://en.wikipedia.org/wiki/Autoencoder](https://en.wikipedia.org/wiki/Autoencoder?cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DL0120EN-SkillsNetwork-20629446&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ)
- [http://ufldl.stanford.edu/tutorial/unsupervised/Autoencoders/](http://ufldl.stanford.edu/tutorial/unsupervised/Autoencoders?cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DL0120EN-SkillsNetwork-20629446&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ)
- [http://www.slideshare.net/billlangjun/simple-introduction-to-autoencoder](http://www.slideshare.net/billlangjun/simple-introduction-to-autoencoder?cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DL0120EN-SkillsNetwork-20629446&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ)
- [http://www.slideshare.net/danieljohnlewis/piotr-mirowski-review-autoencoders-deep-learning-ciuuk14](http://www.slideshare.net/danieljohnlewis/piotr-mirowski-review-autoencoders-deep-learning-ciuuk14?cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DL0120EN-SkillsNetwork-20629446&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ)
- [https://cs.stanford.edu/~quocle/tutorial2.pdf](https://cs.stanford.edu/~quocle/tutorial2.pdf?cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DL0120EN-SkillsNetwork-20629446&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ)
- <https://gist.github.com/hussius/1534135a419bb0b957b9>
- [http://www.deeplearningbook.org/contents/autoencoders.html](http://www.deeplearningbook.org/contents/autoencoders.html?cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DL0120EN-SkillsNetwork-20629446&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ)
- [http://www.kdnuggets.com/2015/03/deep-learning-curse-dimensionality-autoencoders.html/](http://www.kdnuggets.com/2015/03/deep-learning-curse-dimensionality-autoencoders.html?cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DL0120EN-SkillsNetwork-20629446&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ)
- [https://www.youtube.com/watch?v=xTU79Zs4XKY](https://www.youtube.com/watch?v=xTU79Zs4XKY&cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DL0120EN-SkillsNetwork-20629446&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ)
- [http://www-personal.umich.edu/~jizhu/jizhu/wuke/Stone-AoS82.pdf](http://www-personal.umich.edu/~jizhu/jizhu/wuke/Stone-AoS82.pdf?cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DL0120EN-SkillsNetwork-20629446&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ)
<hr>
Copyright © 2018 [Cognitive Class](https://cocl.us/DX0108EN_CC). This notebook and its source code are released under the terms of the [MIT License](https://bigdatauniversity.com/mit-license?cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DL0120EN-SkillsNetwork-20629446&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ&cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DL0120EN-SkillsNetwork-20629446&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ).
| github_jupyter |
# 含并行连结的网络(GoogLeNet)
在2014年的ImageNet图像识别挑战赛中,一个名叫GoogLeNet的网络结构大放异彩 [1]。它虽然在名字上向LeNet致敬,但在网络结构上已经很难看到LeNet的影子。GoogLeNet吸收了NiN中网络串联网络的思想,并在此基础上做了很大改进。在随后的几年里,研究人员对GoogLeNet进行了数次改进,本节将介绍这个模型系列的第一个版本。
## Inception 块
GoogLeNet中的基础卷积块叫作Inception块,得名于同名电影《盗梦空间》(Inception)。与上一节介绍的NiN块相比,这个基础块在结构上更加复杂,如图5.8所示。

由图5.8可以看出,Inception块里有4条并行的线路。前3条线路使用窗口大小分别是$1\times 1$、$3\times 3$和$5\times 5$的卷积层来抽取不同空间尺寸下的信息,其中中间2个线路会对输入先做$1\times 1$卷积来减少输入通道数,以降低模型复杂度。第四条线路则使用$3\times 3$最大池化层,后接$1\times 1$卷积层来改变通道数。4条线路都使用了合适的填充来使输入与输出的高和宽一致。最后我们将每条线路的输出在通道维上连结,并输入接下来的层中去。
Inception块中可以自定义的超参数是每个层的输出通道数,我们以此来控制模型复杂度。
```
import tensorflow as tf
print(tf.__version__)
for gpu in tf.config.experimental.list_physical_devices('GPU'):
tf.config.experimental.set_memory_growth(gpu, True)
class Inception(tf.keras.layers.Layer):
def __init__(self,c1, c2, c3, c4):
super().__init__()
# 线路1,单1 x 1卷积层
self.p1_1 = tf.keras.layers.Conv2D(c1, kernel_size=1, activation='relu', padding='same')
# 线路2,1 x 1卷积层后接3 x 3卷积层
self.p2_1 = tf.keras.layers.Conv2D(c2[0], kernel_size=1, padding='same', activation='relu')
self.p2_2 = tf.keras.layers.Conv2D(c2[1], kernel_size=3, padding='same',
activation='relu')
# 线路3,1 x 1卷积层后接5 x 5卷积层
self.p3_1 = tf.keras.layers.Conv2D(c3[0], kernel_size=1, padding='same', activation='relu')
self.p3_2 = tf.keras.layers.Conv2D(c3[1], kernel_size=5, padding='same',
activation='relu')
# 线路4,3 x 3最大池化层后接1 x 1卷积层
self.p4_1 = tf.keras.layers.MaxPool2D(pool_size=3, padding='same', strides=1)
self.p4_2 = tf.keras.layers.Conv2D(c4, kernel_size=1, padding='same', activation='relu')
def call(self, x):
p1 = self.p1_1(x)
p2 = self.p2_2(self.p2_1(x))
p3 = self.p3_2(self.p3_1(x))
p4 = self.p4_2(self.p4_1(x))
return tf.concat([p1, p2, p3, p4], axis=-1) # 在通道维上连结输出
Inception(64, (96, 128), (16, 32), 32)
```
## GoogLeNet模型
GoogLeNet跟VGG一样,在主体卷积部分中使用5个模块(block),每个模块之间使用步幅为2的$3\times 3$最大池化层来减小输出高宽。第一模块使用一个64通道的$7\times 7$卷积层。
```
b1 = tf.keras.models.Sequential()
b1.add(tf.keras.layers.Conv2D(64, kernel_size=7, strides=2, padding='same', activation='relu'))
b1.add(tf.keras.layers.MaxPool2D(pool_size=3, strides=2, padding='same'))
```
第二模块使用2个卷积层:首先是64通道的$1\times 1$卷积层,然后是将通道增大3倍的$3\times 3$卷积层。它对应Inception块中的第二条线路。
```
b2 = tf.keras.models.Sequential()
b2.add(tf.keras.layers.Conv2D(64, kernel_size=1, padding='same', activation='relu'))
b2.add(tf.keras.layers.Conv2D(192, kernel_size=3, padding='same', activation='relu'))
b2.add(tf.keras.layers.MaxPool2D(pool_size=3, strides=2, padding='same'))
```
第三模块串联2个完整的Inception块。第一个Inception块的输出通道数为$64+128+32+32=256$,其中4条线路的输出通道数比例为$64:128:32:32=2:4:1:1$。其中第二、第三条线路先分别将输入通道数减小至$96/192=1/2$和$16/192=1/12$后,再接上第二层卷积层。第二个Inception块输出通道数增至$128+192+96+64=480$,每条线路的输出通道数之比为$128:192:96:64 = 4:6:3:2$。其中第二、第三条线路先分别将输入通道数减小至$128/256=1/2$和$32/256=1/8$。
```
b3 = tf.keras.models.Sequential()
b3.add(Inception(64, (96, 128), (16, 32), 32))
b3.add(Inception(128, (128, 192), (32, 96), 64))
b3.add(tf.keras.layers.MaxPool2D(pool_size=3, strides=2, padding='same'))
```
第四模块更加复杂。它串联了5个Inception块,其输出通道数分别是$192+208+48+64=512$、$160+224+64+64=512$、$128+256+64+64=512$、$112+288+64+64=528$和$256+320+128+128=832$。这些线路的通道数分配和第三模块中的类似,首先含$3\times 3$卷积层的第二条线路输出最多通道,其次是仅含$1\times 1$卷积层的第一条线路,之后是含$5\times 5$卷积层的第三条线路和含$3\times 3$最大池化层的第四条线路。其中第二、第三条线路都会先按比例减小通道数。这些比例在各个Inception块中都略有不同。
```
b4 = tf.keras.models.Sequential()
b4.add(Inception(192, (96, 208), (16, 48), 64))
b4.add(Inception(160, (112, 224), (24, 64), 64))
b4.add(Inception(128, (128, 256), (24, 64), 64))
b4.add(Inception(112, (144, 288), (32, 64), 64))
b4.add(Inception(256, (160, 320), (32, 128), 128))
b4.add(tf.keras.layers.MaxPool2D(pool_size=3, strides=2, padding='same'))
```
第五模块有输出通道数为$256+320+128+128=832$和$384+384+128+128=1024$的两个Inception块。其中每条线路的通道数的分配思路和第三、第四模块中的一致,只是在具体数值上有所不同。需要注意的是,第五模块的后面紧跟输出层,该模块同NiN一样使用全局平均池化层来将每个通道的高和宽变成1。最后我们将输出变成二维数组后接上一个输出个数为标签类别数的全连接层。
```
b5 = tf.keras.models.Sequential()
b5.add(Inception(256, (160, 320), (32, 128), 128))
b5.add(Inception(384, (192, 384), (48, 128), 128))
b5.add(tf.keras.layers.GlobalAvgPool2D())
net = tf.keras.models.Sequential([b1, b2, b3, b4, b5, tf.keras.layers.Dense(10)])
```
GoogLeNet模型的计算复杂,而且不如VGG那样便于修改通道数。本节里我们将输入的高和宽从224降到96来简化计算。下面演示各个模块之间的输出的形状变化。
```
X = tf.random.uniform(shape=(1, 96, 96, 1))
for layer in net.layers:
X = layer(X)
print(layer.name, 'output shape:\t', X.shape)
```
## 获取数据和训练模型
我们使用高和宽均为96像素的图像来训练GoogLeNet模型。训练使用的图像依然来自Fashion-MNIST数据集。
```
import numpy as np
class DataLoader():
def __init__(self):
fashion_mnist = tf.keras.datasets.fashion_mnist
(self.train_images, self.train_labels), (self.test_images, self.test_labels) = fashion_mnist.load_data()
self.train_images = np.expand_dims(self.train_images.astype(np.float32)/255.0,axis=-1)
self.test_images = np.expand_dims(self.test_images.astype(np.float32)/255.0,axis=-1)
self.train_labels = self.train_labels.astype(np.int32)
self.test_labels = self.test_labels.astype(np.int32)
self.num_train, self.num_test = self.train_images.shape[0], self.test_images.shape[0]
def get_batch_train(self, batch_size):
index = np.random.randint(0, np.shape(self.train_images)[0], batch_size)
#need to resize images to (224,224)
resized_images = tf.image.resize_with_pad(self.train_images[index],224,224,)
return resized_images.numpy(), self.train_labels[index]
def get_batch_test(self, batch_size):
index = np.random.randint(0, np.shape(self.test_images)[0], batch_size)
#need to resize images to (224,224)
resized_images = tf.image.resize_with_pad(self.test_images[index],224,224,)
return resized_images.numpy(), self.test_labels[index]
batch_size = 128
dataLoader = DataLoader()
x_batch, y_batch = dataLoader.get_batch_train(batch_size)
print("x_batch shape:",x_batch.shape,"y_batch shape:", y_batch.shape)
def train_googlenet():
net.load_weights("5.9_googlenet_weights.h5")
epoch = 5
num_iter = dataLoader.num_train//batch_size
for e in range(epoch):
for n in range(num_iter):
x_batch, y_batch = dataLoader.get_batch_train(batch_size)
net.fit(x_batch, y_batch)
if n%20 == 0:
net.save_weights("5.9_googlenet_weights.h5")
# optimizer = tf.keras.optimizers.SGD(learning_rate=0.05, momentum=0.0, nesterov=False)
optimizer = tf.keras.optimizers.Adam(lr=1e-7)
net.compile(optimizer=optimizer,
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
x_batch, y_batch = dataLoader.get_batch_train(batch_size)
net.fit(x_batch, y_batch)
train_googlenet()
net.load_weights("5.9_googlenet_weights.h5")
x_test, y_test = dataLoader.get_batch_test(2000)
net.evaluate(x_test, y_test, verbose=2)
```
## 小结
* Inception块相当于一个有4条线路的子网络。它通过不同窗口形状的卷积层和最大池化层来并行抽取信息,并使用$1\times 1$卷积层减少通道数从而降低模型复杂度。
* GoogLeNet将多个设计精细的Inception块和其他层串联起来。其中Inception块的通道数分配之比是在ImageNet数据集上通过大量的实验得来的。
* GoogLeNet和它的后继者们一度是ImageNet上最高效的模型之一:在类似的测试精度下,它们的计算复杂度往往更低。
| github_jupyter |
## Callin Switzer
10 Dec 2018
### Modified TLD script for running simulation
```
import matplotlib.pyplot as plt
%matplotlib inline
import numpy as np
import os
import math
import seaborn as sns
from scipy.integrate import odeint
import random
import time
from datetime import datetime
from matplotlib.patches import Ellipse
from collections import OrderedDict
import matplotlib.gridspec as gridspec
import sys
import pandas as pd
import importlib
print(sys.version)
now = datetime.now()
print("last run on " + str(now))
import multiProcTraj
import multiProcTraj as mpt
# define directories
baseDir = os.getcwd()
dataDir = r'D:\MothSimulations\11c-AggressiveManeuver\Qstore\hws_am_con'
figDir = r'D:\Dropbox\AcademiaDropbox\mothMachineLearning_dataAndFigs\Figs'
dataOutput = r'D:\Dropbox\AcademiaDropbox\mothMachineLearning_dataAndFigs\DataOutput'
savedModels = r'D:\Dropbox\AcademiaDropbox\mothMachineLearning_dataAndFigs\savedModels'
randomRawData = r'D:/Dropbox/AcademiaDropbox/mothMachineLearning_dataAndFigs/PythonGeneratedData'
# specify ranges
# x,xd,y,yd,
# theta,thetad,phi,phid,
# F, alpha, tau0
np.random.seed(seed=123)
ranges = np.array([[0, 0], [-1500, 1500], [0, 0], [-1500, 1500],
[0, 2*np.pi], [-25, 25], [0, 2*np.pi], [-25, 25],
[0, 44300], [0, 2*np.pi], [-100000, 100000]])
tic = time.time()
mpt.nstep = 10000
mpt.t = np.linspace(0, 0.02, num = mpt.nstep, endpoint = False) # time cut into 100 timesteps
mpt.nrun = 100 #number of trajectories.
# initialize the matrix of 0's
zeroMatrix = np.zeros([mpt.nrun, mpt.nstep])
x, xd, y, yd, \
theta, thetad, phi, phid = [zeroMatrix.copy() for ii in
range(len([ "x", "xd", "y", "yd",
"theta", "thetad", "phi", "phid"]))]
# generate random initial conditions for state 0
state0 = np.random.uniform(ranges[:, 0], ranges[:, 1],
size=(mpt.nrun, ranges.shape[0]), )
# loop through all the runs
for i in range(0,mpt.nrun):
# run ODE
state = odeint(mpt.FlyTheBug, state0[i, :], mpt.t)
x[i,:], xd[i,:] = state[:,0], state[:,1]
y[i,:], yd[i, :] = state[:,2], state[:,3]
theta[i,:], thetad[i, :] = state[:,4],state[:,5]
phi[i,:], phid[i, :] = state[:, 6], state[:,7]
print('elapsed time = ',time.time()-tic)
plt.figure(figsize = [10,10])
plt.axes().set_aspect('equal', 'datalim')
for i in range(0,mpt.nrun):
plt.plot(x[i, :],y[i, :], label = 'trajectory x vs y')
plt.scatter(x[:,-1:][:,0], y[:,-1:][:,0])
dfa = pd.DataFrame(state0, columns=["x","xd","y","yd","theta","thetad","phi","phid", "F", "alpha", "tau0"])
df_final = pd.DataFrame(OrderedDict({"xf" : x[:,-1:][:,0],
"xdf" : xd[:,-1:][:,0],
"yf": y[:,-1:][:,0],
"ydf" : yd[:,-1:][:,0],
"thetaf" : theta[:,-1:][:,0],
"thetadf" : thetad[:,-1:][:,0],
"phif" : phi[:,-1:][:,0],
"phidf" : phid[:,-1:][:,0],
} ))
df_final.head()
df_c = pd.concat([dfa.reset_index(drop=True), df_final], axis=1)
df_c.head()
dataOutput
#df_c.to_csv(os.path.join(dataOutput, "UpdatedCodeCheck.csv"), index= False)
# read in Jorge's comparison
checkData = pd.read_csv(os.path.join(dataOutput, "UpdatedCodeCheck_output_all.csv"), )
checkData.head()
plt.figure(figsize = [10,10])
plt.axes().set_aspect('equal', 'datalim')
for i in range(0,mpt.nrun):
plt.plot(x[i, :],y[i, :], alpha = 0.2)
plt.scatter(x[:, -1:], y[:, -1:], c= 'orange', label = "Python")
plt.scatter(checkData.x_a, checkData.y_a, label = "Matlab")
plt.legend()
plt.savefig(os.path.join(figDir, "PythonVsMatlab.png") )
plt.show()
x.shape # nrun, nstep
# plotnum of timesteps vs. xf (don't get too high or too low)
# check parameters
# check duration
plt.axes().set_aspect('equal', 'datalim')
plt.scatter(x[:, -1:].reshape(-1) - checkData.x_a, y[:, -1:].reshape(-1) - checkData.y_a, s = 15, alpha = 0.4, c= "black")
plt.title("Error from matlab vs. python")
plt.savefig(os.path.join(figDir, "PythonVsMatlab_error.png") )
plt.show()
```
## Visualize moth at different timesteps
* See if moth is spinning
* See if abdomen is rotating too far
```
def cart2pol(x, y):
rho = np.sqrt(x**2 + y**2)
phi = np.arctan2(y, x)
return(rho, phi)
def pol2cart(rho, phi):
'''
rho: radius
phi: angle (in radians)
'''
x = rho * np.cos(phi)
y = rho * np.sin(phi)
return(x, y)
def midpoint(p1, p2):
return ((p1[0]+p2[0])/2, (p1[1]+p2[1])/2)
plt.figure(figsize = [10,10])
plt.axes().set_aspect('equal', 'datalim')
i =54
plt.plot(x[i, :],y[i, :], label = 'trajectory x vs y')
nstep = mpt.nstep
# plot moth and force
timestep = 0
# theta = head angle
# phi = abdomen angle
thoraxLen = 0.908 * 2# cm
abLen = 1.747 *2 #cm
bodyWidth = 1.1
# plot trajectory
fig, ax = plt.subplots( figsize = [10,10])
ax.set_aspect('equal', 'datalim')
i += 1
ax.plot(x[i, :],y[i, :], label = 'trajectory x vs y')
# add body positions:
for timestep in (np.linspace(0, nstep - 1, num = 8 )).astype(int):
center = np.array([x[i, timestep], y[i, timestep]])
head = center + np.array(pol2cart(thoraxLen, theta[i, timestep]))
abTip = center + np.array(pol2cart(abLen, phi[i, timestep]))
xx, yy = zip(*[center, head])
xab,yab = zip(*[center, abTip])
el = Ellipse(midpoint(center, head), width = thoraxLen, height = bodyWidth, facecolor='#907760', alpha=0.8, angle = math.degrees(theta[i, timestep]))
el2 = Ellipse(midpoint(center, abTip), width = abLen, height = bodyWidth, facecolor='#DEC9B0', alpha=0.8, angle = math.degrees(phi[i, timestep]))
ax.add_artist(el)
ax.add_artist(el2)
ax.plot(xx, yy, 'k', alpha = 0.2)
#ax.scatter(xx, yy, s= 10, c = 'k', alpha = 0.2)
ax.plot(xab,yab, 'k', alpha = 0.2)
#ax.scatter(xab,yab, s = 10, c = 'k', alpha = 0.2)
# plot force
forceAlpha = state0[i, 9]
forceCenter = midpoint(center, head)
forceMagnitude = state0[i, 8] / 15000 # scale
#ax.scatter(forceCenter[0], forceCenter[1], s= 30, c= 'r', alpha = 1)
forceAngle = theta[i, timestep] + forceAlpha
forceTip = np.add(pol2cart(forceMagnitude, forceAngle), forceCenter)
#ax.scatter(forceTip[0], forceTip[1], s= 30, c= 'r', alpha = 1)
ax.arrow(x = forceCenter[0], y = forceCenter[1],
dx = forceTip[0] - forceCenter[0], dy = forceTip[1] - forceCenter[1],
head_width = 0.2, color = "#B61212")
nrun = mpt.nrun
# plot moth with ellipses
# refref: what force would it take for the moth to counter-act gravity?
timestep = 0
# theta = head angle
# phi = abdomen angle
thoraxLen = 0.908 * 2# cm
abLen = 1.7475 *2 #cm
bodyWidth = 1.1
# plot trajectory
fig, ax = plt.subplots( figsize = [10,10])
ax.set_aspect('equal', 'datalim')
for i in np.random.randint(0, high = nrun, size = 3):
# plot trajectory
ax.plot(x[i, :],y[i, :], label = 'trajectory x vs y')
# add body positions:
for timestep in (np.linspace(0, nstep -1, num = 8 )).astype(int):
center = np.array([x[i, timestep], y[i, timestep]])
head = center + np.array(pol2cart(thoraxLen, theta[i, timestep]))
abTip = center + np.array(pol2cart(abLen, phi[i, timestep]))
xx, yy = zip(*[center, head])
xab,yab = zip(*[center, abTip])
el = Ellipse(midpoint(center, head), width = thoraxLen, height = bodyWidth, facecolor='#907760', alpha=0.8, angle = math.degrees(theta[i, timestep]))
el2 = Ellipse(midpoint(center, abTip), width = abLen, height = bodyWidth, facecolor='#DEC9B0', alpha=0.8, angle = math.degrees(phi[i, timestep]))
ax.add_artist(el)
ax.add_artist(el2)
ax.plot(xx, yy, 'k', alpha = 0.2)
#ax.scatter(xx, yy, s= 10, c = 'k', alpha = 0.2)
ax.plot(xab,yab, 'k', alpha = 0.2)
#ax.scatter(xab,yab, s = 10, c = 'k', alpha = 0.2)
# plot force
forceAlpha = state0[i, 9]
forceCenter = midpoint(center, head)
forceMagnitude = state0[i, 8] / 15000 # scale
#ax.scatter(forceCenter[0], forceCenter[1], s= 30, c= 'r', alpha = 1)
forceAngle = theta[i, timestep] + forceAlpha
forceTip = np.add(pol2cart(forceMagnitude, forceAngle), forceCenter)
#ax.scatter(forceTip[0], forceTip[1], s= 30, c= 'r', alpha = 1)
ax.arrow(x = forceCenter[0], y = forceCenter[1],
dx = forceTip[0] - forceCenter[0], dy = forceTip[1] - forceCenter[1],
head_width = 0.2, color = "#B61212")
# from mpl_toolkits.axes_grid1.anchored_artists import AnchoredSizeBar
# scalebar = AnchoredSizeBar(ax.transData,
# 980/1500, 'Force of gravity', 'lower right',
# pad=0.1,
# color="#B61212",
# frameon=False,
# size_vertical=0.07, sep = 8)
# plt.gca().add_artist(scalebar)
# ax.add_artist(scalebar)
```
## Calculate Angle from torque
```
# reset k to original
# 7 sec, tao = 100, g = 0
# drop back to linear -- see what happens with fixed toque
# do again with damping (set c to 0) set to 0 (should see resonance) -- we should know the resonance for a linear spring
# do c = 0 with no torque (can set nstep = 1000 / sec)
# plot beta (angle b/w thorax and abdomen) vs. time
# plot derivative of beta
# redo with nonlinear spring
# set body angle to 90 degrees, and see what happens.
# redo with c == 0
# set drag to 0 c_d = 0
# reload multiProcTrag
importlib.reload(multiProcTraj)
#print(mpt.g, mpt.K, mpt.springExponent)
mpt.c = 0.1
mpt.g = 0.0
mpt.K = 29.3*100
print(mpt.g, mpt.K, mpt.springExponent)
np.random.seed(seed=12035)
# x,xd,y,yd,
# theta,thetad,phi,phid,
# F, alpha, tau0
# ranges = np.array([[0, 0], [0.0001, 0.0001], [0, 0], [0.0001, 0.0001],
# [np.pi/2, np.pi/2], [0, 0], [3*np.pi/2, 3*np.pi/2], [0, 0],
# [0, 0], [0, 0], [0, -100000]])
############# manually enter values #################
x,xd,y,yd = 0, 0.0001, 0, 0.0001
theta,thetad,phi,phid = np.pi/2, 0.0001, np.pi + np.pi/2, 0.0001
F, alpha, tau0 = 0, 0, 1000
#####################################################
ranges = np.array([[x, x], [xd, xd], [y, y], [yd, yd],
[theta, theta], [thetad, thetad], [phi,phi], [phid, phid],
[F, F], [alpha, alpha], [tau0, tau0]])
tic = time.time()
simLength = 8 # seconds
samplesPerSec= 100
mpt.nstep = int(simLength * samplesPerSec) # steps per second
mpt.t = np.linspace(0, simLength, num = mpt.nstep, endpoint = False) # time cut into 100 timesteps
nrun = 1 #number of trajectories.
# initialize the matrix of 0's
zeroMatrix = np.zeros([nrun, len(mpt.t)])
x, xd, y, yd, \
theta, thetad, phi, phid = [zeroMatrix.copy() for ii in
range(len([ "x", "xd", "y", "yd",
"theta", "thetad", "phi", "phid"]))]
print(x.shape)
# generate random initial conditions for state 0
state0 = np.random.uniform(ranges[:, 0], ranges[:, 1],
size=(nrun, ranges.shape[0]), )
# loop through all the runs
for i in range(0,nrun):
# run ODE
state = odeint(mpt.FlyTheBug, state0[i, :], mpt.t)
x[i,:], xd[i,:] = state[:,0], state[:,1]
y[i,:], yd[i, :] = state[:,2], state[:,3]
theta[i,:], thetad[i, :] = state[:,4],state[:,5]
phi[i,:], phid[i, :] = state[:, 6], state[:,7]
print(i)
print('elapsed time = ',time.time()-tic)
# plot moth and force
timestep = 0
# theta = head angle
# phi = abdomen angle
thoraxLen = 0.908 * 2# cm
abLen = 1.747 *2 #cm
bodyWidth = 1.1
# plot trajectory
fig, ax = plt.subplots( figsize = [10,10])
ax.set_aspect('equal', 'datalim')
i = 0
ax.plot(x[i, :],y[i, :], label = 'trajectory x vs y')
# add body positions:
for timestep in (np.linspace(0, mpt.nstep -1 , num = 8 )).astype(int):
center = np.array([x[i, timestep], y[i, timestep]])
head = center + np.array(pol2cart(thoraxLen, theta[i, timestep]))
abTip = center + np.array(pol2cart(abLen, phi[i, timestep]))
xx, yy = zip(*[center, head])
xab,yab = zip(*[center, abTip])
el = Ellipse(midpoint(center, head), width = thoraxLen, height = bodyWidth, facecolor='#907760', alpha=0.8, angle = math.degrees(theta[i, timestep]))
el2 = Ellipse(midpoint(center, abTip), width = abLen, height = bodyWidth, facecolor='#DEC9B0', alpha=0.8, angle = math.degrees(phi[i, timestep]))
ax.add_artist(el)
ax.add_artist(el2)
ax.plot(xx, yy, 'k', alpha = 0.2)
#ax.scatter(xx, yy, s= 10, c = 'k', alpha = 0.2)
ax.plot(xab,yab, 'k', alpha = 0.2)
#ax.scatter(xab,yab, s = 10, c = 'k', alpha = 0.2)
# plot force
forceAlpha = state0[i, 9]
forceCenter = midpoint(center, head)
forceMagnitude = state0[i, 8] / 15000 # scale
#ax.scatter(forceCenter[0], forceCenter[1], s= 30, c= 'r', alpha = 1)
forceAngle = theta[i, timestep] + forceAlpha
forceTip = np.add(pol2cart(forceMagnitude, forceAngle), forceCenter)
#ax.scatter(forceTip[0], forceTip[1], s= 30, c= 'r', alpha = 1)
ax.arrow(x = forceCenter[0], y = forceCenter[1],
dx = forceTip[0] - forceCenter[0], dy = forceTip[1] - forceCenter[1],
head_width = 0.2, color = "#B61212")
# plot moth and force
timestep = 0
# theta = head angle
# phi = abdomen angle
thoraxLen = 0.908 * 2# cm
abLen = 1.747 *2 #cm
bodyWidth = 1.1
# plot trajectory
if i >= (nrun - 1):
i = 0
i += 1
# ax.plot(x[i, :],y[i, :], label = 'trajectory x vs y')
fig = plt.subplots(figsize = np.array([30,8])/ 1.4)
gs = gridspec.GridSpec(3, 1,height_ratios=[1,1,1])
ax = plt.subplot(gs[0])
ax1 = plt.subplot(gs[1])
ax2 = plt.subplot(gs[2])
ax.set_aspect('equal')
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
ax.spines['left'].set_visible(False)
ctr1 = 0
for i in range(0, nrun):
# add body positions:
ctr = 0
for timestep in (np.linspace(0, mpt.nstep - 1, num = 40)).astype(int):
#center = np.array([x[i, timestep], y[i, timestep]])
center = np.array([ctr,ctr1])
ctr += 3
head = center + np.array(pol2cart(thoraxLen, theta[i, timestep]))
abTip = center + np.array(pol2cart(abLen, phi[i, timestep]))
xx, yy = zip(*[center, head])
xab,yab = zip(*[center, abTip])
el = Ellipse(midpoint(center, head), width = thoraxLen, height = bodyWidth, facecolor='#907760', alpha=0.8, angle = math.degrees(theta[i, timestep]))
el2 = Ellipse(midpoint(center, abTip), width = abLen, height = bodyWidth, facecolor='#DEC9B0', alpha=0.8, angle = math.degrees(phi[i, timestep]))
if np.mod(ctr, 1) == 0:
ax.add_artist(el)
ax.add_artist(el2)
ax.plot(xx, yy, 'k', alpha = 0.2)
ax.plot(xab,yab, 'k', alpha = 0.2)
#plt.annotate("tau = " + str(np.round(state0[i,10])), center + (5, 1))
ax.set_title(r'$\tau$ =' + str(np.round(state0[i,10], 5))+ ", g = " + str(mpt.g) + ", spring Exp. = " + str(mpt.springExponent) + ", c = " + str(mpt.c))
ax.set_xlabel("")
ax.set_xticks([0, 120, 126])
ax.set_xticklabels([0, np.round(np.max(mpt.t)), ""])
ax.set_ylim([-4,3])
ax.set_yticks([])
ctr1 -= 8
ax1.plot(mpt.t, [math.degrees(phi[i, jj] - theta[i, jj] - np.pi) for jj in range(len(phi[i,:]))])
#ax1.hlines(-0, xmin = 0, xmax = np.max(mpt.t))
ax1.set_ylabel("degrees b/w\nthorax and abdomen")
ax1.set_xlabel("")
ax1.spines['right'].set_visible(False)
ax1.spines['top'].set_visible(False)
degs = np.array([math.degrees(phi[i, jj] - theta[i, jj]) for jj in range(len(phi[i,:]))])
ax2.plot(mpt.t[2:], np.diff(degs[1:]) * samplesPerSec)
ax2.set_ylabel("change in degrees")
ax2.set_xlabel("Time (s)")
ax2.spines['top'].set_visible(False)
ax2.spines['right'].set_visible(False)
plt.savefig(os.path.join(mpt.figDir, "TauAndAngle3_noDamping.png"), dpi = 150)
plt.show()
# plot moth and force
timestep = 0
# theta = head angle
# phi = abdomen angle
thoraxLen = 0.908 * 2# cm
abLen = 1.747 *2 #cm
bodyWidth = 1.1
# plot trajectory
if i >= (nrun - 1):
i = 0
i += 1
# ax.plot(x[i, :],y[i, :], label = 'trajectory x vs y')
fig = plt.subplots(figsize = np.array([30,8])/ 1.4)
gs = gridspec.GridSpec(3, 1,height_ratios=[1,1,1])
ax = plt.subplot(gs[0])
ax1 = plt.subplot(gs[1])
ax2 = plt.subplot(gs[2])
ax.set_aspect('equal')
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
ax.spines['left'].set_visible(False)
ctr1 = 0
for i in range(0, nrun):
# add body positions:
ctr = 0
for timestep in (np.linspace(0, mpt.nstep - 1, num = 40)).astype(int):
#center = np.array([x[i, timestep], y[i, timestep]])
center = np.array([ctr,ctr1])
ctr += 3
head = center + np.array(pol2cart(thoraxLen, theta[i, timestep]))
abTip = center + np.array(pol2cart(abLen, phi[i, timestep]))
xx, yy = zip(*[center, head])
xab,yab = zip(*[center, abTip])
el = Ellipse(midpoint(center, head), width = thoraxLen, height = bodyWidth, facecolor='#907760', alpha=0.8, angle = math.degrees(theta[i, timestep]))
el2 = Ellipse(midpoint(center, abTip), width = abLen, height = bodyWidth, facecolor='#DEC9B0', alpha=0.8, angle = math.degrees(phi[i, timestep]))
if np.mod(ctr, 1) == 0:
ax.add_artist(el)
ax.add_artist(el2)
ax.plot(xx, yy, 'k', alpha = 0.2)
ax.plot(xab,yab, 'k', alpha = 0.2)
#plt.annotate("tau = " + str(np.round(state0[i,10])), center + (5, 1))
ax.set_title(r'$\tau$ =' + str(np.round(state0[i,10], 5))+ ", g = " + str(mpt.g) + ", spring Exp. = " + str(mpt.springExponent) + ", c = " + str(mpt.c))
ax.set_xlabel("")
ax.set_xticks([0, 120, 126])
ax.set_xticklabels([0, np.round(np.max(mpt.t)), ""])
ax.set_ylim([-4,3])
ax.set_yticks([])
ctr1 -= 8
ax1.plot(mpt.t, [math.degrees(phi[i, jj]) for jj in range(len(phi[i,:]))])
#ax1.hlines(-0, xmin = 0, xmax = np.max(mpt.t))
ax1.set_ylabel("phi")
ax1.set_xlabel("")
ax1.spines['right'].set_visible(False)
ax1.spines['top'].set_visible(False)
degs = np.array([math.degrees(phi[i, jj] - theta[i, jj]) for jj in range(len(phi[i,:]))])
ax2.plot(mpt.t, [math.degrees(theta[i, jj]) for jj in range(len(phi[i,:]))])
ax2.set_ylabel("theta")
ax2.set_xlabel("Time (s)")
ax2.spines['top'].set_visible(False)
ax2.spines['right'].set_visible(False)
plt.show()
fig, ax = plt.subplots( figsize = [10,3])
plt.plot(mpt.t, [math.degrees(phi[i, jj] - theta[i, jj]) for jj in range(len(phi[i,:]))])
plt.hlines(-180, xmin = 0, xmax = np.max(mpt.t))
plt.show()
plt.plot(mpt.t)
nrun
for i in range(0, nrun):
fig, ax = plt.subplots( figsize = [10,3])
plt.plot([math.degrees(theta[i, jj]) for jj in range(len(phi[i,:]))])
plt.show()
for i in range(0, nrun):
fig, ax = plt.subplots( figsize = [10,3])
plt.plot([math.degrees(theta[i, jj]) for jj in range(len(phi[i,:]))])
plt.show()
# plot moth and force
timestep = 0
# theta = head angle
# phi = abdomen angle
thoraxLen = 0.908 * 2# cm
abLen = 1.747 *2 #cm
bodyWidth = 1.1
# plot trajectory
if i >= (nrun - 1):
i = 0
i += 1
# ax.plot(x[i, :],y[i, :], label = 'trajectory x vs y')
#ax.set_aspect('equal')
ctr1 = 0
for i in range(0, nrun):
fig, ax = plt.subplots( figsize = [10,3])
plt.plot((phi[i, :] - theta[i, :]) - np.pi)
plt.ylim(0, 10)
plt.hlines(y = np.pi, xmin = 0, xmax = 10000)
plt.show()
print("tau = ", state0[i,10])
plt.show()
plt.plot((phi[i, :] - theta[i, :]) - np.pi)
plt.show()
center
```
# See if abdomen twangs
```
np.random.seed(seed=12035)
# x,xd,y,yd,
# theta,thetad,phi,phid,
# F, alpha, tau0
# ranges = np.array([[0, 0], [0.0001, 0.0001], [0, 0], [0.0001, 0.0001],
# [np.pi/2, np.pi/2], [0, 0], [3*np.pi/2, 3*np.pi/2], [0, 0],
# [0, 0], [0, 0], [0, -100000]])
ranges = np.array([[0, 0], [0.0001, 0.0001], [0, 0], [0.0001, 0.0001],
[-np.pi/4, -np.pi/4], [0, 0], [3*np.pi/2, 3*np.pi/2], [0, 0],
[0, 0], [0, 0], [0, 0]])
tic = time.time()
nstep = 5000000
t = np.linspace(0, 7, num = nstep, endpoint = False) # time cut into 100 timesteps
nrun = 1 #number of trajectories.
# initialize the matrix of 0's
zeroMatrix = np.zeros([nrun, nstep])
x, xd, y, yd, \
theta, thetad, phi, phid = [zeroMatrix.copy() for ii in
range(len([ "x", "xd", "y", "yd",
"theta", "thetad", "phi", "phid"]))]
# generate random initial conditions for state 0
state0 = np.random.uniform(ranges[:, 0], ranges[:, 1],
size=(nrun, ranges.shape[0]), )
# loop through all the runs
for i in range(0,nrun):
# run ODE
state = odeint(FlyTheBug, state0[i, :], t)
x[i,:], xd[i,:] = state[:,0], state[:,1]
y[i,:], yd[i, :] = state[:,2], state[:,3]
theta[i,:], thetad[i, :] = state[:,4],state[:,5]
phi[i,:], phid[i, :] = state[:, 6], state[:,7]
print(i)
print('elapsed time = ',time.time()-tic)
# plot moth and force
timestep = 0
# theta = head angle
# phi = abdomen angle
thoraxLen = 0.908 * 2# cm
abLen = 1.747 *2 #cm
bodyWidth = 1.1
# plot trajectory
if i >= (nrun - 1):
i = 0
i += 1
# ax.plot(x[i, :],y[i, :], label = 'trajectory x vs y')
fig, ax = plt.subplots( figsize = [30,10])
ax.set_aspect('equal')
ctr1 = 0
for i in range(0, nrun):
# add body positions:
ctr = 0
for timestep in (np.linspace(0, nstep - 1, num = 40)).astype(int):
#center = np.array([x[i, timestep], y[i, timestep]])
center = np.array([ctr,ctr1])
ctr += 3
head = center + np.array(pol2cart(thoraxLen, theta[i, timestep]))
abTip = center + np.array(pol2cart(abLen, phi[i, timestep]))
xx, yy = zip(*[center, head])
xab,yab = zip(*[center, abTip])
el = Ellipse(midpoint(center, head), width = thoraxLen, height = bodyWidth, facecolor='#907760', alpha=0.8, angle = math.degrees(theta[i, timestep]))
el2 = Ellipse(midpoint(center, abTip), width = abLen, height = bodyWidth, facecolor='#DEC9B0', alpha=0.8, angle = math.degrees(phi[i, timestep]))
ax.add_artist(el)
ax.add_artist(el2)
ax.plot(xx, yy, 'k', alpha = 0.2)
ax.plot(xab,yab, 'k', alpha = 0.2)
#plt.annotate("tau = " + str(np.round(state0[i,10])), center + (5, 1))
plt.annotate(r'$\tau$ =' + str(np.round(state0[i,10], 5)), center + (5, 1))
plt.xlabel("Time (sec)")
plt.xticks([0, 120, 130], [0, np.round(np.max(t)), ""])
plt.yticks([-4,3])
ctr1 -= 8
fig.savefig(os.path.join(figDir, "ZeroTorqueAngle.png"), dpi = 150)
plt.show()
for i in range(0, nrun):
fig, ax = plt.subplots( figsize = [10,3])
plt.plot([math.degrees(phi[i, jj] - theta[i, jj] ) for jj in range(len(phi[i,:]))])
#plt.hlines(360, xmin = 0, xmax = nstep)
plt.ylabel("Degrees")
plt.xlabel("nstep")
plt.show()
```
| github_jupyter |
## Title :
Classification using Decision Tree
## Description :
The goal of this exercise is to get comfortable using Decision Trees for classification in sklearn. Eventually, you will produce a plot similar to the one given below:
<img src="../fig/fig1.png" style="width: 500px;">
## Instructions:
- Read the train and test datafile as Pandas data frame.
- Use `minority` and `bachelor` as the predictor variables and `won` as the response.
- Fit a decision tree of depth 2 and another of depth 10 on the training data.
- Call the function `plot_boundary` to visualise the decision boundary of these 2 classifiers.
- Increase the number of predictor variables as mentioned in scaffold.
- Initialize a decision tree classifier of depth 2, 10 and 15.
- Fit the model on the train data.
- Compute the train and test accuracy scores for each classifier.
- Use the helper code to look at the feature importance of the predictors from the decision tree of depth 15.
## Hints:
<a href="https://scikit-learn.org/stable/modules/generated/sklearn.tree.DecisionTreeClassifier.html" target="_blank">sklearn.DecisionTreeClassifier()</a>
Generates a Logistic Regression classifier
<a href="https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.LogisticRegression.html#sklearn.linear_model.LogisticRegression.score" target="_blank">sklearn.score()</a>
Accuracy classification score.
<a href="https://scikit-learn.org/stable/modules/generated/sklearn.tree.DecisionTreeClassifier.html#sklearn.tree.DecisionTreeClassifier.fit" target="_blank">classifier.fit()</a>
Build a decision tree classifier from the training set (X, y).
**Note: This exercise is auto-graded and you can try multiple attempts.**
```
# Import necessary libraries
import numpy as np
import pandas as pd
import sklearn as sk
import seaborn as sns
from sklearn import tree
import matplotlib.pyplot as plt
from helper import plot_boundary
from prettytable import PrettyTable
from sklearn.tree import DecisionTreeClassifier
from sklearn.model_selection import cross_val_score
pd.set_option('display.width', 100)
pd.set_option('display.max_columns', 20)
plt.rcParams["figure.figsize"] = (12,8)
# Read the data file "election_train.csv" as a Pandas dataframe
elect_train = pd.read_csv("election_train.csv")
# Read the data file "election_test.csv" as a Pandas dataframe
elect_test = pd.read_csv("election_test.csv")
# Take a quick look at the train data
elect_train.head()
# Set the columns minority and bachelor as train data predictors
X_train = ___
# Set the columns minority and bachelor as test data predictors
X_test = ___
# Set the column "won" as the train response variable
y_train = ___
# Set the column "won" as the test response variable
y_test = ___
# Initialize a Decision Tree classifier with a depth of 2
dt1 = ___
# Fit the classifier on the train data
___
# Initialize a Decision Tree classifier with a depth of 10
dt2 = ___
# Fit the classifier on the train data
___
# Call the function plot_boundary from the helper file to get
# the decision boundaries of both the classifiers
plot_boundary(elect_train, dt1, dt2)
# Set of predictor columns
pred_cols = ['minority', 'density','hispanic','obesity','female','income','bachelor','inactivity']
# Use the columns above as the predictor data from the train data
X_train = elect_train[pred_cols]
# Use the columns above as the predictor data from the test data
X_test = elect_test[pred_cols]
# Initialize a Decision Tree classifier with a depth of 2
dt1 = ___
# Initialize a Decision Tree classifier with a depth of 10
dt2 = ___
# Initialize a Decision Tree classifier with a depth of 15
dt3 = ___
# Fit all the classifier on the train data
___
___
___
### edTest(test_accuracy) ###
# Compute the train and test accuracy for the first decision tree classifier of depth 2
dt1_train_acc = ___
dt1_test_acc = ___
# Compute the train and test accuracy for the second decision tree classifier of depth 10
dt2_train_acc = ___
dt2_test_acc = ___
# Compute the train and test accuracy for the third decision tree classifier of depth 15
dt3_train_acc = ___
dt3_test_acc = ___
# Helper code to plot the scores of each classifier as a table
pt = PrettyTable()
pt.field_names = ['Max Depth', 'Number of Features', 'Train Accuracy', 'Test Accuracy']
pt.add_row([2, 2, round(dt1_train_acc, 4), round(dt1_test_acc,4)])
pt.add_row([10, 2, round(dt2_train_acc,4), round(dt2_test_acc,4)])
pt.add_row([15, len(pred_cols), round(dt3_train_acc,4), round(dt3_test_acc,4)])
print(pt)
```
| github_jupyter |
```
import sys
import time
import os.path
from glob import glob
from datetime import datetime, timedelta
import h5py
import numpy as np
sys.path.insert(0, '/glade/u/home/ksha/WORKSPACE/utils/')
sys.path.insert(0, '/glade/u/home/ksha/WORKSPACE/Analog_BC/')
sys.path.insert(0, '/glade/u/home/ksha/WORKSPACE/Analog_BC/utils/')
sys.path.insert(0, '/glade/u/home/ksha/PUBLISH/fcstpp/')
import data_utils as du
import graph_utils as gu
from fcstpp import utils as fu
from namelist import *
# graph tools
import cmaps
import cartopy.crs as ccrs
import cartopy.mpl.geoaxes
import cartopy.feature as cfeature
from cartopy.io.shapereader import Reader
from cartopy.feature import ShapelyFeature
from cartopy.mpl.gridliner import LONGITUDE_FORMATTER, LATITUDE_FORMATTER
import matplotlib.pyplot as plt
import matplotlib.lines as mlines
import matplotlib.colors as colors
import matplotlib.patches as patches
from matplotlib.collections import PatchCollection
from matplotlib import ticker
import matplotlib.ticker as mticker
import matplotlib.gridspec as gridspec
from mpl_toolkits.axes_grid1.inset_locator import inset_axes
%matplotlib inline
import warnings
warnings.filterwarnings("ignore")
need_publish = False
# True: publication quality figures
# False: low resolution figures in the notebook
if need_publish:
dpi_ = fig_keys['dpi']
else:
dpi_ = 75
```
# Data
```
# importing domain information
with h5py.File(save_dir+'BC_domain_info.hdf', 'r') as h5io:
base_lon = h5io['base_lon'][...]
base_lat = h5io['base_lat'][...]
bc_lon = h5io['bc_lon'][...]
bc_lat = h5io['bc_lat'][...]
etopo_bc = h5io['etopo_bc'][...]
land_mask = h5io['land_mask_base'][...]
land_mask_bc = h5io['land_mask_bc'][...]
with h5py.File(save_dir+'BCH_wshed_groups.hdf', 'r') as h5io:
flag_sw = h5io['flag_sw'][...]
flag_si = h5io['flag_si'][...]
flag_n = h5io['flag_n'][...]
```
## Datetime info
```
mon_rain = np.array([9, 10, 11, 0, 1, 2])
mon_dry = np.array([3, 4, 5, 6, 7, 8])
base = datetime(2017, 1, 1)
date_list = [base + timedelta(days=x) for x in range(365+365+365)]
rain_inds = np.zeros((len(date_list),), dtype=bool)
dry_inds = np.zeros((len(date_list),), dtype=bool)
mon_inds = []
for d, date in enumerate(date_list):
mon_inds.append(date.month-1)
if date.month-1 in mon_dry:
dry_inds[d] = True
else:
rain_inds[d] = True
mon_inds = np.array(mon_inds)
fcst_leads = np.arange(3, 72*3+3, 3, dtype=np.float)
```
## CRPS results
```
from scipy.stats import wilcoxon
def wilcoxon_by_leads(FCST1, FCST2):
N, L = FCST1.shape
w_stat = np.empty((L,))
p_vals = np.empty((L,))
series1 = FCST1
series2 = FCST2
for i in range(L):
w_, p_ = wilcoxon(series1[:, i], series2[:, i],)
w_stat[i] = w_
p_vals[i] = p_
return w_stat, p_vals
def boost_mean_std(data):
temp = fu.score_bootstrap_1d(data, bootstrap_n=100)
mean_ = np.mean(temp, axis=-1)
p95_ = np.quantile(temp-mean_[..., None], 0.95, axis=-1)
return mean_, p95_
# with h5py.File(save_dir+'CLIM_CRPS_BCH_2017.hdf', 'r') as h5io:
# CLIM_CRPS_17 = h5io['CRPS'][...]
# with h5py.File(save_dir+'CLIM_CRPS_BCH_2018.hdf', 'r') as h5io:
# CLIM_CRPS_18 = h5io['CRPS'][...]
# with h5py.File(save_dir+'CLIM_CRPS_BCH_2019.hdf', 'r') as h5io:
# CLIM_CRPS_19 = h5io['CRPS'][...]
# with h5py.File(save_dir+'BASE_final_CRPS_BCH_2017.hdf', 'r') as h5io:
# BASE_CRPS_17 = h5io['CRPS'][...]
# with h5py.File(save_dir+'BASE_final_CRPS_BCH_2018.hdf', 'r') as h5io:
# BASE_CRPS_18 = h5io['CRPS'][...]
# with h5py.File(save_dir+'BASE_final_CRPS_BCH_2019.hdf', 'r') as h5io:
# BASE_CRPS_19 = h5io['CRPS'][...]
# with h5py.File(save_dir+'SL_final_CRPS_BCH_2017.hdf', 'r') as h5io:
# SL_CRPS_17 = h5io['CRPS'][...]
# with h5py.File(save_dir+'SL_final_CRPS_BCH_2018.hdf', 'r') as h5io:
# SL_CRPS_18 = h5io['CRPS'][...]
# with h5py.File(save_dir+'SL_final_CRPS_BCH_2019.hdf', 'r') as h5io:
# SL_CRPS_19 = h5io['CRPS'][...]
# with h5py.File(save_dir+'BASE_CNN_CRPS_BCH_2017.hdf', 'r') as h5io:
# BCNN_CRPS_17 = h5io['CRPS'][...]
# with h5py.File(save_dir+'BASE_CNN_CRPS_BCH_2018.hdf', 'r') as h5io:
# BCNN_CRPS_18 = h5io['CRPS'][...]
# with h5py.File(save_dir+'BASE_CNN_CRPS_BCH_2019.hdf', 'r') as h5io:
# BCNN_CRPS_19 = h5io['CRPS'][...]
# with h5py.File(save_dir+'SL_CNN_CRPS_BCH_2017.hdf', 'r') as h5io:
# SCNN_CRPS_17 = h5io['CRPS'][...]
# with h5py.File(save_dir+'SL_CNN_CRPS_BCH_2018.hdf', 'r') as h5io:
# SCNN_CRPS_18 = h5io['CRPS'][...]
# with h5py.File(save_dir+'SL_CNN_CRPS_BCH_2019.hdf', 'r') as h5io:
# SCNN_CRPS_19 = h5io['CRPS'][...]
# with h5py.File(save_dir+'GEFS_CRPS_BCH_2017.hdf', 'r') as h5io:
# GEFS_CRPS_17 = h5io['CRPS'][...]
# with h5py.File(save_dir+'GEFS_CRPS_BCH_2018.hdf', 'r') as h5io:
# GEFS_CRPS_18 = h5io['CRPS'][...]
# with h5py.File(save_dir+'GEFS_CRPS_BCH_2019.hdf', 'r') as h5io:
# GEFS_CRPS_19 = h5io['CRPS'][...]
# CLIM_CRPS = np.concatenate((CLIM_CRPS_17, CLIM_CRPS_18, CLIM_CRPS_19), axis=0)
# BASE_CRPS = np.concatenate((BASE_CRPS_17, BASE_CRPS_18, BASE_CRPS_19), axis=0)
# SL_CRPS = np.concatenate((SL_CRPS_17, SL_CRPS_18, SL_CRPS_19), axis=0)
# BCNN_CRPS = np.concatenate((BCNN_CRPS_17, BCNN_CRPS_18, BCNN_CRPS_19), axis=0)
# SCNN_CRPS = np.concatenate((SCNN_CRPS_17, SCNN_CRPS_18, SCNN_CRPS_19), axis=0)
# GEFS_CRPS = np.concatenate((GEFS_CRPS_17, GEFS_CRPS_18, GEFS_CRPS_19), axis=0)
# BASE_CRPS_rain = np.transpose(BASE_CRPS[rain_inds, :], (0, 2, 1))
# SL_CRPS_rain = np.transpose(SL_CRPS[rain_inds, :], (0, 2, 1))
# BCNN_CRPS_rain = np.transpose(BCNN_CRPS[rain_inds, :], (0, 2, 1))
# SCNN_CRPS_rain = np.transpose(SCNN_CRPS[rain_inds, :], (0, 2, 1))
# GEFS_CRPS_rain = np.transpose(GEFS_CRPS[rain_inds, :], (0, 2, 1))
# CLIM_CRPS_rain = np.transpose(CLIM_CRPS[rain_inds, :], (0, 2, 1))
# BASE_CRPS_dry = np.transpose(BASE_CRPS[dry_inds, :], (0, 2, 1))
# SL_CRPS_dry = np.transpose(SL_CRPS[dry_inds, :], (0, 2, 1))
# BCNN_CRPS_dry = np.transpose(BCNN_CRPS[dry_inds, :], (0, 2, 1))
# SCNN_CRPS_dry = np.transpose(SCNN_CRPS[dry_inds, :], (0, 2, 1))
# GEFS_CRPS_dry = np.transpose(GEFS_CRPS[dry_inds, :], (0, 2, 1))
# CLIM_CRPS_dry = np.transpose(CLIM_CRPS[dry_inds, :], (0, 2, 1))
# BASE_CRPS_rain_mean, BASE_CRPS_rain_std = boost_mean_std(BASE_CRPS_rain)
# SL_CRPS_rain_mean, SL_CRPS_rain_std = boost_mean_std(SL_CRPS_rain)
# BCNN_CRPS_rain_mean, BCNN_CRPS_rain_std = boost_mean_std(BCNN_CRPS_rain)
# SCNN_CRPS_rain_mean, SCNN_CRPS_rain_std = boost_mean_std(SCNN_CRPS_rain)
# GEFS_CRPS_rain_mean, GEFS_CRPS_rain_std = boost_mean_std(GEFS_CRPS_rain)
# CLIM_CRPS_rain_mean = np.nanmean(CLIM_CRPS_rain, axis=(0, 1))
# BASE_CRPS_dry_mean, BASE_CRPS_dry_std = boost_mean_std(BASE_CRPS_dry)
# SL_CRPS_dry_mean, SL_CRPS_dry_std = boost_mean_std(SL_CRPS_dry)
# BCNN_CRPS_dry_mean, BCNN_CRPS_dry_std = boost_mean_std(BCNN_CRPS_dry)
# SCNN_CRPS_dry_mean, SCNN_CRPS_dry_std = boost_mean_std(SCNN_CRPS_dry)
# GEFS_CRPS_dry_mean, GEFS_CRPS_dry_std = boost_mean_std(GEFS_CRPS_dry)
# CLIM_CRPS_dry_mean = np.nanmean(CLIM_CRPS_dry, axis=(0, 1))
# pad_nan = np.array([np.nan, np.nan])
# CRPS_MEAN = {}
# CRPS_MEAN['gfs_rain'] = np.concatenate((pad_nan, 1-GEFS_CRPS_rain_mean/CLIM_CRPS_rain_mean), axis=0)
# CRPS_MEAN['base_rain'] = np.concatenate((pad_nan, 1-BASE_CRPS_rain_mean/CLIM_CRPS_rain_mean), axis=0)
# CRPS_MEAN['sl_rain'] = np.concatenate((pad_nan, 1-SL_CRPS_rain_mean/CLIM_CRPS_rain_mean), axis=0)
# CRPS_MEAN['bcnn_rain'] = np.concatenate((pad_nan, 1-BCNN_CRPS_rain_mean/CLIM_CRPS_rain_mean), axis=0)
# CRPS_MEAN['scnn_rain'] = np.concatenate((pad_nan, 1-SCNN_CRPS_rain_mean/CLIM_CRPS_rain_mean), axis=0)
# CRPS_MEAN['gfs_dry'] = np.concatenate((pad_nan, 1-GEFS_CRPS_dry_mean/CLIM_CRPS_dry_mean), axis=0)
# CRPS_MEAN['base_dry'] = np.concatenate((pad_nan, 1-BASE_CRPS_dry_mean/CLIM_CRPS_dry_mean), axis=0)
# CRPS_MEAN['sl_dry'] = np.concatenate((pad_nan, 1-SL_CRPS_dry_mean/CLIM_CRPS_dry_mean), axis=0)
# CRPS_MEAN['bcnn_dry'] = np.concatenate((pad_nan, 1-BCNN_CRPS_dry_mean/CLIM_CRPS_dry_mean), axis=0)
# CRPS_MEAN['scnn_dry'] = np.concatenate((pad_nan, 1-SCNN_CRPS_dry_mean/CLIM_CRPS_dry_mean), axis=0)
# CRPS_STD = {}
# CRPS_STD['gfs_rain'] = np.concatenate((pad_nan, GEFS_CRPS_rain_std), axis=0)
# CRPS_STD['base_rain'] = np.concatenate((pad_nan, BASE_CRPS_rain_std), axis=0)
# CRPS_STD['sl_rain'] = np.concatenate((pad_nan, SL_CRPS_rain_std), axis=0)
# CRPS_STD['bcnn_rain'] = np.concatenate((pad_nan, BCNN_CRPS_rain_std), axis=0)
# CRPS_STD['scnn_rain'] = np.concatenate((pad_nan, SCNN_CRPS_rain_std), axis=0)
# CRPS_STD['gfs_dry'] = np.concatenate((pad_nan, GEFS_CRPS_dry_std), axis=0)
# CRPS_STD['base_dry'] = np.concatenate((pad_nan, BASE_CRPS_dry_std), axis=0)
# CRPS_STD['sl_dry'] = np.concatenate((pad_nan, SL_CRPS_dry_std), axis=0)
# CRPS_STD['bcnn_dry'] = np.concatenate((pad_nan, BCNN_CRPS_dry_std), axis=0)
# CRPS_STD['scnn_dry'] = np.concatenate((pad_nan, SCNN_CRPS_dry_std), axis=0)
# BASE_CRPS_rain_ave = np.nanmean(BASE_CRPS[rain_inds, ...], axis=(2,))
# SL_CRPS_rain_ave = np.nanmean(SL_CRPS[rain_inds, ...], axis=(2,))
# BCNN_CRPS_rain_ave = np.nanmean(BCNN_CRPS[rain_inds, ...], axis=(2,))
# SCNN_CRPS_rain_ave = np.nanmean(SCNN_CRPS[rain_inds, ...], axis=(2,))
# BASE_CRPS_dry_ave = np.nanmean(BASE_CRPS[dry_inds, ...], axis=(2,))
# SL_CRPS_dry_ave = np.nanmean(SL_CRPS[dry_inds, ...], axis=(2,))
# BCNN_CRPS_dry_ave = np.nanmean(BCNN_CRPS[dry_inds, ...], axis=(2,))
# SCNN_CRPS_dry_ave = np.nanmean(SCNN_CRPS[dry_inds, ...], axis=(2,))
# CRPS_wilcox = {}
# w_stat, p_vals = wilcoxon_by_leads(BASE_CRPS_rain_ave, SL_CRPS_rain_ave)
# CRPS_wilcox['BASE_SL_rain'] = np.concatenate((pad_nan, p_vals), axis=0)
# w_stat, p_vals = wilcoxon_by_leads(BCNN_CRPS_rain_ave, SCNN_CRPS_rain_ave)
# CRPS_wilcox['BCNN_SCNN_rain'] = np.concatenate((pad_nan, p_vals), axis=0)
# w_stat, p_vals = wilcoxon_by_leads(BASE_CRPS_rain_ave, BCNN_CRPS_rain_ave)
# CRPS_wilcox['BASE_BCNN_rain'] = np.concatenate((pad_nan, p_vals), axis=0)
# w_stat, p_vals = wilcoxon_by_leads(SL_CRPS_rain_ave, SCNN_CRPS_rain_ave)
# CRPS_wilcox['SL_SCNN_rain'] = np.concatenate((pad_nan, p_vals), axis=0)
# w_stat, p_vals = wilcoxon_by_leads(BASE_CRPS_dry_ave, SL_CRPS_dry_ave)
# CRPS_wilcox['BASE_SL_dry'] = np.concatenate((pad_nan, p_vals), axis=0)
# w_stat, p_vals = wilcoxon_by_leads(BCNN_CRPS_dry_ave, SCNN_CRPS_dry_ave)
# CRPS_wilcox['BCNN_SCNN_dry'] = np.concatenate((pad_nan, p_vals), axis=0)
# w_stat, p_vals = wilcoxon_by_leads(BASE_CRPS_dry_ave, BCNN_CRPS_dry_ave)
# CRPS_wilcox['BASE_BCNN_dry'] = np.concatenate((pad_nan, p_vals), axis=0)
# w_stat, p_vals = wilcoxon_by_leads(SL_CRPS_dry_ave, SCNN_CRPS_dry_ave)
# CRPS_wilcox['SL_SCNN_dry'] = np.concatenate((pad_nan, p_vals), axis=0)
# np.save(save_dir+'CRPS_BCH_rain_dry.npy', CRPS_MEAN)
# np.save(save_dir+'CRPS_BCH_rain_dry_std.npy', CRPS_STD)
# np.save(save_dir+'CRPS_BCH_wilcox.npy', CRPS_wilcox)
CRPS_MEAN = np.load(save_dir+'CRPS_BCH_rain_dry.npy', allow_pickle=True)[()]
CRPS_STD = np.load(save_dir+'CRPS_BCH_rain_dry_std.npy', allow_pickle=True)[()]
CRPS_wilcox = np.load(save_dir+'CRPS_BCH_wilcox.npy', allow_pickle=True)[()]
```
# Figure
```
edge_bc = [-141, -113.25, 48.25, 60]
def aspc_cal(edge):
return (edge[3]-edge[2])/(edge[1]-edge[0])
r_bc = aspc_cal(edge_bc)
cmap_pct, A = gu.precip_cmap()
gray = [0.5, 0.5, 0.5]
KW = {}
KW['gfs'] = {'linestyle': '-', 'color': gray, 'ecolor':'0.25', 'linewidth':2.5, 'elinewidth':1.5, 'barsabove':False}
KW['era'] = {'linestyle': '-', 'color': 'k', 'ecolor':'0.25', 'linewidth':2.5, 'elinewidth':1.5, 'barsabove':False}
KW['base'] = {'linestyle': '-', 'color': orange, 'ecolor':'0.25', 'linewidth':2.5, 'elinewidth':1.5, 'barsabove':False}
KW['bcnn'] = {'linestyle': '-', 'color': red, 'ecolor':'0.25', 'linewidth':2.5, 'elinewidth':1.5, 'barsabove':False}
KW['sl'] = {'linestyle': '-', 'color': cyan, 'ecolor':'0.25', 'linewidth':2.5, 'elinewidth':1.5, 'barsabove':False}
KW['scnn'] = {'linestyle': '-', 'color': blue, 'ecolor':'0.25', 'linewidth':2.5, 'elinewidth':1.5, 'barsabove':False}
kw_lines = {}
kw_lines['gfs'] = {'linestyle': '-', 'color': gray, 'linewidth':2.5}
kw_lines['era'] = {'linestyle': '-', 'color': 'k', 'linewidth':2.5}
kw_lines['base'] = {'linestyle': '-', 'color': orange, 'linewidth':2.5}
kw_lines['bcnn'] = {'linestyle': '-', 'color': red, 'linewidth':2.5}
kw_lines['sl'] = {'linestyle': '-', 'color': cyan, 'linewidth':2.5}
kw_lines['scnn'] = {'linestyle': '-', 'color': blue, 'linewidth':2.5}
kw_bar = {'bottom':0.0, 'width': 3.0, 'color': '0.75', 'edgecolor': 'k', 'linestyle': '-', 'linewidth': 0}
kw_step = {'color': 'k', 'linestyle': '-', 'linewidth': 1.5, 'where':'mid'}
fontsize = 13.5
```
## Line graph
```
cates = ['rain', 'dry']
method_crps = ['base', 'bcnn', 'sl', 'scnn', 'gfs']
fig = plt.figure(figsize=(13, 8), dpi=dpi_)
gs = gridspec.GridSpec(3, 5, height_ratios=[1, 0.6, 0.6], width_ratios=[1, 1, 0.1, 1, 1])
ax11 = plt.subplot(gs[0, 0])
ax12 = plt.subplot(gs[0, 1])
ax13 = plt.subplot(gs[1, 0])
ax14 = plt.subplot(gs[1, 1])
ax15 = plt.subplot(gs[2, 0])
ax16 = plt.subplot(gs[2, 1])
ax21 = plt.subplot(gs[0, 3])
ax22 = plt.subplot(gs[0, 4])
ax23 = plt.subplot(gs[1, 3])
ax24 = plt.subplot(gs[1, 4])
ax25 = plt.subplot(gs[2, 3])
ax26 = plt.subplot(gs[2, 4])
plt.subplots_adjust(0, 0, 1, 1, hspace=0, wspace=0)
AX_all = [ax11, ax12, ax13, ax14, ax15, ax16, ax21, ax22, ax23, ax24, ax25, ax26]
AX_crps = [ax11, ax12, ax21, ax22]
AX_diff_all = [ax13, ax14, ax15, ax16, ax23, ax24, ax25, ax26]
AX_diff_sub1 = [ax13, ax23]
AX_diff_sub2 = [ax14, ax24]
AX_diff_sub3 = [ax15, ax25]
AX_diff_sub4 = [ax16, ax26]
x_start1, y_start1 = 0.025, 0.975
x_start2, y_start2 = 0.025, 0.95
x_start3, y_start3 = 0.025, 0.95
handles = []
ax_t1 = fig.add_axes([0, 1.0, (2/4.1), 0.031875])
ax_t1.set_axis_off()
handles.append(ax_t1.text(0.5, 1, 'Station-wise-mean CRPSS in Oct-Mar, 2017-2019', ha='center', va='top',
fontsize=fontsize, transform=ax_t1.transAxes))
handles.append(ax_t1.text(0.368+0.006, 0.999, '[*]', ha='left', va='top', fontsize=10, transform=ax_t1.transAxes))
handles.append(ax_t1.text(0.510, 0.999, '[**]', ha='left', va='top', fontsize=10, transform=ax_t1.transAxes))
ax_t2 = fig.add_axes([2.1/4.1, 1.0, (2/4.1), 0.031875])
ax_t2.set_axis_off()
handles.append(ax_t2.text(0.5, 1, 'Station-wise-mean CRPSS in Apr-Sept, 2017-2019', ha='center', va='top',
fontsize=fontsize, transform=ax_t2.transAxes))
handles += gu.string_partial_format(fig, ax11, x_start1, y_start1, 'left', 'top',
['(a) CRPSS of ', 'noSL-H15', ','],
['k', orange, 'k'], [fontsize,]*3, ['normal', 'bold', 'normal'])
handles += gu.string_partial_format(fig, ax11, x_start1+0.1, y_start1-0.055, 'left', 'top',
['noSL-CNN', ', and quantile'], [red, 'k'], [fontsize,]*2, ['bold', 'normal'])
handles += gu.string_partial_format(fig, ax11, x_start1+0.1, y_start1-0.11, 'left', 'top',
['mapped ', 'GEFS',], ['k', gray, 'k'], [fontsize,]*3, ['normal', 'bold', 'normal'])
handles += gu.string_partial_format(fig, ax12, x_start1, y_start1, 'left', 'top',
['(b) CRPSS of ', 'SL-H15', ', ', 'SL-CNN', ','],
['k', cyan, 'k', blue, 'k'], [fontsize,]*5, ['normal', 'bold', 'normal', 'bold', 'normal'])
handles += gu.string_partial_format(fig, ax12, x_start1+0.1, y_start1-0.055, 'left', 'top',
['and quantile mapped ', 'GEFS'], ['k', gray], [fontsize,]*2, ['normal', 'bold'])
handles.append(ax21.text(x_start1, y_start1, '(c) Same as (a), but in Apr-Sept.', ha='left', va='top',
fontsize=fontsize, transform=ax21.transAxes, zorder=6))
handles.append(ax22.text(x_start1, y_start1, '(d) Same as (b), but in Apr-Sept.', ha='left', va='top',
fontsize=fontsize, transform=ax22.transAxes, zorder=6))
fontsize_list = [fontsize,]*5
fontweight_list = ['normal', 'bold', 'normal', 'bold', 'normal']
handles += gu.string_partial_format(fig, ax13, x_start2, y_start2, 'left', 'top',
['(e) ', 'noSL-CNN', ' minus ', 'noSL-H15'],
['k', red, 'k', orange, 'k'], fontsize_list, fontweight_list)
handles += gu.string_partial_format(fig, ax14, x_start2, y_start2, 'left', 'top',
['(f) ', 'SL-CNN', ' minus ', 'SL-H15'],
['k', blue, 'k', cyan, 'k'], fontsize_list, fontweight_list)
handles += gu.string_partial_format(fig, ax23, x_start2, y_start2, 'left', 'top',
['(g) ', 'noSL-CNN', ' minus ', 'noSL-H15'],
['k', red, 'k', orange, 'k'], fontsize_list, fontweight_list)
handles += gu.string_partial_format(fig, ax24, x_start2, y_start2, 'left', 'top',
['(h) ', 'SL-CNN', ' minus ', 'SL-H15'],
['k', blue, 'k', cyan, 'k'], fontsize_list, fontweight_list)
handles += gu.string_partial_format(fig, ax15, x_start3, y_start3, 'left', 'top',
['(i) ', 'SL-H15', ' minus ', 'noSL-H15'],
['k', cyan, 'k', orange, 'k'], fontsize_list, fontweight_list)
handles += gu.string_partial_format(fig, ax16, x_start3, y_start3, 'left', 'top',
['(j) ', 'SL-CNN', ' minus ', 'noSL-CNN'],
['k', blue, 'k', red, 'k'], fontsize_list, fontweight_list)
handles += gu.string_partial_format(fig, ax25, x_start3, y_start3, 'left', 'top',
['(k) ', 'SL-H15', ' minus ', 'noSL-H15'],
['k', cyan, 'k', orange, 'k'], fontsize_list, fontweight_list)
handles += gu.string_partial_format(fig, ax26, x_start3, y_start3, 'left', 'top',
['(l) ', 'SL-CNN', ' minus ', 'noSL-CNN'],
['k', blue, 'k', red, 'k'], fontsize_list, fontweight_list)
for ax in AX_all:
ax = gu.ax_decorate_box(ax)
ax.xaxis.set_tick_params(labelsize=fontsize)
ax.yaxis.set_tick_params(labelsize=fontsize)
ax.set_xlim([0, 168])
ax.set_xticks(np.arange(12, 168, 24))
ax.axhline(0, xmin=0, xmax=1.0, linewidth=1.5, linestyle='-', color='k', zorder=3)
for d in range(1, 8):
ax.text(d/7.0-1/14, 0.02, 'd-{}'.format(int(d-1)), ha='center', va='bottom',
fontsize=fontsize, transform=ax.transAxes, zorder=6)
for i, ax in enumerate(AX_crps):
ax.set_ylim([0.1, 0.6])
ax.set_yticks([0.2, 0.3, 0.4, 0.5,])
if i > 1:
for y in np.arange(0.1, 0.7, 0.1):
if np.abs(y-0)>0.01:
ax.axhline(y, xmin=0, xmax=1.0, linewidth=1.5, linestyle=':', color='0.5', zorder=2)
for day in np.arange(24, 168+24, 24):
ax.axvline(day, ymin=0, ymax=0.9, linewidth=1.5, linestyle=':', color='0.5', zorder=2)
else:
for y in np.arange(0.1, 0.6, 0.1):
if np.abs(y-0)>0.01:
ax.axhline(y, xmin=0, xmax=1.0, linewidth=1.5, linestyle=':', color='0.5', zorder=2)
for day in np.arange(24, 168+24, 24):
ax.axvline(day, ymin=0, ymax=0.85, linewidth=1.5, linestyle=':', color='0.5', zorder=2)
for ax in AX_diff_sub1 + AX_diff_sub2:
ax.set_ylim([-0.02, 0.1])
ax.set_yticks([0.0, 0.02, 0.04, 0.06, 0.08,])
for y in np.arange(0.02, 0.1, 0.02):
ax.axhline(y, xmin=0, xmax=1.0, linewidth=1.5, linestyle=':', color='0.5')
for day in np.arange(24, 168+24, 24):
ax.axvline(day, ymin=0, ymax=5/6, linewidth=1.5, linestyle=':', color='0.5', zorder=5)
for ax in AX_diff_sub3 + AX_diff_sub4:
ax.set_ylim([-0.04, 0.08])
ax.set_yticks([0.0, 0.02, 0.04, 0.06])
for y in np.arange(-0.02, 0.08, 0.02):
ax.axhline(y, xmin=0, xmax=1.0, linewidth=1.5, linestyle=':', color='0.5')
for day in np.arange(24, 168+24, 24):
ax.axvline(day, ymin=0, ymax=5/6, linewidth=1.5, linestyle=':', color='0.5', zorder=5)
ax11.tick_params(labelleft=True)
ax13.tick_params(labelleft=True)
ax15.tick_params(labelleft=True)
for i, key in enumerate(['base', 'bcnn', 'gfs']):
AX_crps[0].errorbar(fcst_leads[:56], CRPS_MEAN['{}_rain'.format(key)],
yerr=CRPS_STD['{}_rain'.format(key)], **KW[key], zorder=4)
AX_crps[2].errorbar(fcst_leads[:56], CRPS_MEAN['{}_dry'.format(key)],
yerr=CRPS_STD['{}_dry'.format(key)], **KW[key], zorder=4)
for i, key in enumerate(['sl', 'scnn', 'gfs']):
AX_crps[1].errorbar(fcst_leads[:56], CRPS_MEAN['{}_rain'.format(key)],
yerr=CRPS_STD['{}_rain'.format(key)], **KW[key], zorder=4)
AX_crps[3].errorbar(fcst_leads[:56], CRPS_MEAN['{}_dry'.format(key)],
yerr=CRPS_STD['{}_dry'.format(key)], **KW[key], zorder=4)
for i, key in enumerate(['rain', 'dry']):
diff_ = CRPS_MEAN['bcnn_{}'.format(key)]-CRPS_MEAN['base_{}'.format(key)]
AX_diff_sub1[i].bar(fcst_leads[:56], diff_, **kw_bar, zorder=2)
AX_diff_sub1[i].step(fcst_leads[:56], diff_, **kw_step, zorder=4)
AX_diff_sub1[i].vlines(fcst_leads[2]-1.5, ymin=0.0, ymax=diff_[2], color='k', linestyle='-', linewidth=1.5, zorder=5)
diff_ = CRPS_MEAN['scnn_{}'.format(key)]-CRPS_MEAN['sl_{}'.format(key)]
AX_diff_sub2[i].bar(fcst_leads[:56], diff_, **kw_bar, zorder=2)
AX_diff_sub2[i].step(fcst_leads[:56], diff_, **kw_step, zorder=4)
AX_diff_sub2[i].vlines(fcst_leads[2]-1.5, ymin=0.0, ymax=diff_[2], color='k', linestyle='-', linewidth=1.5, zorder=5)
diff_ = np.copy(CRPS_MEAN['sl_{}'.format(key)]-CRPS_MEAN['base_{}'.format(key)])
AX_diff_sub3[i].step(fcst_leads[:56], diff_, **kw_step, zorder=4)
AX_diff_sub3[i].vlines(fcst_leads[2]-1.5, ymin=0.0, ymax=diff_[2], color='k', linestyle='-', linewidth=1.5, zorder=5)
diff_[CRPS_wilcox['BASE_SL_{}'.format(key)] > 0.01] = np.nan
AX_diff_sub3[i].bar(fcst_leads[:56], diff_, **kw_bar, zorder=2)
diff_ = np.copy(CRPS_MEAN['scnn_{}'.format(key)]-CRPS_MEAN['bcnn_{}'.format(key)])
AX_diff_sub4[i].step(fcst_leads[:56], diff_, **kw_step, zorder=4)
AX_diff_sub4[i].vlines(fcst_leads[2]-1.5, ymin=0.0, ymax=diff_[2], color='k', linestyle='-', linewidth=1.5, zorder=5)
diff_[CRPS_wilcox['BCNN_SCNN_{}'.format(key)] > 0.01] = np.nan
AX_diff_sub4[i].bar(fcst_leads[:56], diff_, **kw_bar, zorder=2)
#ax11.axvline(fcst_leads[2]-0.75, ymin=0.4, ymax=0.8, color='k', linestyle='-', linewidth=1.5, zorder=5)
handles.append(ax11.text(0.025, 0.375, '+9 hr', ha='left', va='top', fontsize=fontsize, transform=ax11.transAxes))
ax_w1 = fig.add_axes([-0.035, -0.1/1.2/8*10.2, 0.275, 0.075/1.2/8*10.2])
ax_w1.set_axis_off()
ax_w1.text(0, 1, '* The station-wise-mean curves (a-d) are\n averaged over 100 bootstrap replicates.\n Error bars represent the 95% CI.',
ha='left', va='top', fontsize=fontsize, transform=ax_w1.transAxes);
ax_w2 = fig.add_axes([-0.035, -0.145/1.2/8*10.2, 0.5, 0.025/1.2/8*10.2])
ax_w2.set_axis_off()
ax_w2.text(0, 1, '** CRP Skill Score (CRPSS) is calculated relative to the CRPS of 2000-2014 ERA5 monthly CDFs.',
ha='left', va='top', fontsize=fontsize, transform=ax_w2.transAxes);
label_ = [' ',
' ',
' ',
' ',
' ',]
handle_lines = []
handle_lines.append(mlines.Line2D([], [], label=label_[0], **kw_lines['bcnn']))
handle_lines.append(mlines.Line2D([], [], label=label_[1], **kw_lines['base']))
handle_lines.append(mlines.Line2D([], [], label=label_[2], **kw_lines['scnn']))
handle_lines.append(mlines.Line2D([], [], label=label_[3], **kw_lines['sl']))
handle_lines.append(mlines.Line2D([], [], label=label_[4], **kw_lines['gfs']))
ax_lg1 = fig.add_axes([0, -0.075/1.2/8*10.2, 1.0, 0.05/1.2/8*10.2])
ax_lg1.set_axis_off()
LG1 = ax_lg1.legend(handles=handle_lines, bbox_to_anchor=(1, 0.75), ncol=6, loc=7, prop={'size':fontsize}, fancybox=False);
LG1.get_frame().set_facecolor('none')
LG1.get_frame().set_linewidth(0)
LG1.get_frame().set_alpha(1.0)
ax_lg1.text(0.3+0.085, 1, 'noSL-CNN', ha='left', va='top', fontsize=fontsize, fontweight='bold',
color=KW['bcnn']['color'], transform=ax_lg1.transAxes)
ax_lg1.text(0.3+0.205, 1, 'noSL-H15', ha='left', va='top', fontsize=fontsize, fontweight='bold',
color=KW['base']['color'], transform=ax_lg1.transAxes)
ax_lg1.text(0.3+0.325, 1, 'SL-CNN', ha='left', va='top', fontsize=fontsize, fontweight='bold',
color=KW['scnn']['color'], transform=ax_lg1.transAxes)
ax_lg1.text(0.3+0.425, 1, 'SL-H15', ha='left', va='top', fontsize=fontsize, fontweight='bold',
color=KW['sl']['color'], transform=ax_lg1.transAxes)
gu.string_partial_format(fig, ax_lg1, 0.325+0.505, 1, 'left', 'top', ['quantile mapped ', 'GEFS'],
['k', gray, 'k'], [fontsize,]*3, ['normal', 'bold', 'normal']);
handle_legneds = []
handle_legneds.append(patches.Patch(facecolor='none', edgecolor='k', linewidth=1.5,
label='CRPSS difference'))
handle_legneds.append(patches.Patch(facecolor=kw_bar['color'], edgecolor='none',
label='Wilcoxon signed-rank test with p < 0.01'))
ax_lg2 = fig.add_axes([1.9/4.1, -0.0975/1.2/8*10.2, 2.2/4.1, 0.03/1.2/8*10.2])
ax_lg2.set_axis_off()
LG2 = ax_lg2.legend(handles=handle_legneds, bbox_to_anchor=(0.0, 0.5), ncol=2, loc=6,
prop={'size':fontsize}, fancybox=False);
LG2.get_frame().set_facecolor('none')
LG2.get_frame().set_linewidth(0)
LG2.get_frame().set_alpha(1.0)
for handle in handles:
handle.set_bbox(dict(facecolor='w', pad=0, edgecolor='none', zorder=6))
if need_publish:
# Save figure
fig.savefig(fig_dir+'AnEn_BCH_CRPS.png', format='png', **fig_keys)
```
| github_jupyter |
```
import numpy as np
import pandas as pd
%matplotlib inline
df = pd.read_csv("data/weather_m4.csv")
df
df.info()
```
# Detecting and Inspecting Missing Values
```
df[["MIN_TEMP_GROUND", "VIEW_RANGE", "MIST", "WEATHER_CODE"]].head(15)
df.isnull()
df.isnull().any()
df[df.isnull().any(axis=1)]
# rows with all null values
df[df.isnull().all(axis=1)]
df.notnull().all()
every_6th_row = pd.Series(range(5, len(df), 6))
every_6th_row
# getting values from every 6th row
df["MIN_TEMP_GROUND"][every_6th_row]
# double check whether every 6th row has data by ensuring not null
# 1)
df["MIN_TEMP_GROUND"][every_6th_row].notnull().all()
# double check other remaining rows (excpet for every 6th row)
# 2)
df["MIN_TEMP_GROUND"].drop(every_6th_row).isnull().all()
# Rewriting the same logic as above two lines using loc
# 1)
df.loc[every_6th_row, "MIN_TEMP_GROUND"].notnull().all()
# 2)
df.drop(df.index[every_6th_row]).index
df.loc[df.drop(df.index[every_6th_row]).index, "MIN_TEMP_GROUND"]
```
# Handling missing data
```
df.drop(columns="WEATHER_CODE")
# remove unecessary columns
df.drop(columns="WEATHER_CODE", inplace=True)
df.head()
df["MIN_TEMP_GROUND"].fillna(0)
# if we want to fill empty values with forward values or backward values
# forwards method will fill based on the previous values
df['MIN_TEMP_GROUND'].fillna(method='ffill', inplace=True)
df.head(15)
# backward methods use values from backwards so it won't leave first 5 rows with NaN
df['MIN_TEMP_GROUND'].fillna(method='bfill', inplace=True)
df.head(15)
# if we check again , MIN_TEMP_GROUND will have no Null Values anymore
df.isnull().any()
# if we want to see null values rows
df[df.isnull().any(axis=1)]
# if we want to see misisng by which months and year
df.loc[df.isnull().any(axis=1), "YYYYMMDD"].value_counts()
# collecting all null values into seperate df
nulls_dropped = df.dropna()
nulls_dropped.info()
# if we look at the data, we can see indexes are out of align
nulls_dropped[5300:5310]
```
## using thresh
- thresh allow to set what is the minium thresh level
```
# here we set thresh level of 7, keeping rows with at least 7 non NA values
# however we see only 5 non NA values are thre
drop_thresh = df.dropna(thresh=7)
drop_thresh[drop_thresh.isnull().any(axis=1)]
```
# replacing with values
```
rows_to_fill = df.isnull().any(axis=1)
rows_to_fill
df[rows_to_fill]
# we want to fill mean values of data set to those rows with null values
# first, fill the null values with mean
# second, get the values of the previously NA rows
nulls_filled = df.fillna(df.mean())
nulls_filled[rows_to_fill]
# fill null values with Mode
df.fillna(df.mode().iloc[0], inplace=True)
df.head()
```
# Removing Outliers
```
athletes = pd.read_csv("data/athletes.csv")
athletes.info()
%matplotlib inline
athletes.plot.scatter(x="height", y="weight");
heights = athletes["height"]
heights.plot.box();
q1 = heights.quantile(.25)
q3 = heights.quantile(.75)
iqr = q3 - q1
pmin = q1 - 1.5 * iqr
pmax = q3 + 1.5 * iqr
heights.between(pmin, pmax)
nwh = heights[heights.between(pmin, pmax)]
nwh.head()
#new variable without outliners
new_heights = heights.where(heights.between(pmin, pmax))
# for comparions
compare = pd.DataFrame({"before": heights, "after": new_heights})
compare.describe()
compare.plot.box();
# changes to original heights
heights.where(heights.between(pmin, pmax), inplace=True)
athletes.plot.scatter(x="height", y="weight")
```
# Removing Duplicates
```
athletes.duplicated().any()
athletes[athletes.duplicated()]
athletes.drop_duplicates(inplace=True)
athletes.duplicated().any()
# checking how many unique nationality are there
athletes["nationality"].drop_duplicates()
athletes["nationality"].drop_duplicates().sort_values()
# there is bultin function for this
athletes["nationality"].value_counts()
# ratio of men vs women
athletes["sex"].value_counts()
```
# Conversion Types
```
athletes.info()
athletes[["gold", 'silver', 'bronze']].head()
athletes["bronze"].sum()
# above result is not what we want, we want medals columns as integer not as string
athletes['bronze'].astype(int)
# we can see that someone type letter O instead of zero 0 in one of the values
# so we need to fix it first.
athletes[athletes["bronze"] == 'O']
athletes.loc[7521, ['gold', 'silver', 'bronze']] = 0
# after fixing, converting to int
athletes[['gold', 'silver', 'bronze']] = athletes[['gold', 'silver', 'bronze']].astype(int)
# now getting sum of medals
athletes[['gold', 'silver', 'bronze']].sum()
athletes.info()
```
# Fixing Indexes
```
athletes.head()
# as original data have id columns, we can use it as index
athletes.set_index("id", drop=True, inplace=True)
athletes.head()
# changing column name
athletes.rename(
columns={"nationality": "country", "sport": "discipline"},
inplace=True
)
athletes.head()
```
----------
```
df = pd.read_csv("data/weather_m4.csv")
df.dropna(inplace=True)
df.info()
df.head()
# reset index
df.reset_index(drop=True)
```
| github_jupyter |
<small><small><i>
All the IPython Notebooks in **Python Functions** lecture series by Dr. Milaan Parmar are available @ **[GitHub](https://github.com/milaan9/04_Python_Functions)**
</i></small></small>
# Python Global, Local and Nonlocal variables
In this class, you’ll learn about Python Global variables, Local variables, Nonlocal variables and where to use them.
When we define a function with variables, then those variables scope is limited to that function. In Python, the scope of a variable is the portion of a program where the variable is declared. Parameters and variables defined inside a function are not visible from outside the function. Hence, it is called the variable’s local scope.
>**Note:** The inner function does have access to the outer function’s local scope.
When we are executing a function, the life of the variables is up to running time. Once we return from the function, those variables get destroyed. So function does no need to remember the value of a variable from its previous call.
## Global Variables
In Python, a variable declared outside of the function or in global scope is known as a global variable. This means that a global variable can be accessed inside or outside of the function.
For example:
```
# Example 1: Create a Global Variable
global_var = 999
def fun1():
print("Value in 1st function:", global_var)
def fun2():
print("Value in 2nd function:", global_var)
fun1()
fun2()
# Example 2:
x = "global"
def fun():
print("x inside:", x)
fun()
print("x outside:", x)
```
In the above code, we created **`x`** as a global variable and defined a **`fun()`** to print the global variable **`x`**. Finally, we call the **`fun()`** which will print the value of **`x`**.
What if you want to change the value of **`x`** inside a function?
```
# Example 3:
x = "global"
def fun():
x = x * 2
print(x)
fun()
```
**Explanation:**
The output shows an error because Python treats **`x`** as a local variable and **`x`** is also not defined inside **`fun()`**.
To make this work, we use the **`global`** keyword. Visit **[Python Global Keyword](https://github.com/milaan9/04_Python_Functions/blob/main/003_Python_Function_global_Keywords.ipynb)** to learn more.
```
# Example 4:
global_lang = 'DataScience'
def var_scope_test():
local_lang = 'Python'
print(local_lang)
var_scope_test() # Output 'Python'
# outside of function
print(global_lang) # Output 'DataScience'
print(local_lang) # NameError: name 'local_lang' is not defined
# Example 5:
a=90 # 'a' is a variable defined outside of function, i.e., Global variable
def print_data():
a=6 # 'a' is a variable defined inside of function, i.e., local variable
b=30
print("(a,b):(",a,",",b,")")
print_data() #(a,b):( 5 , 10 )
print("Global a :",a) #Global x : 50
print("Local b : ",b) #b is local veriable - throw NameError
```
## Local Variables
A variable declared inside the function's body or in the local scope is known as a local variable.
If we try to access the local variable from the outside of the function, we will get the error as **`NameError`**.
```
# Example 1: Accessing local variable outside the scope
def fun():
y = "local"
fun()
print(y)
```
The output shows an error because we are trying to access a local variable **`y`** in a global scope whereas the local variable only works inside **`fun()`** or local scope.
```
# Example 2: Create a Local Variable
# Normally, we declare a variable inside the function to create a local variable.
def fun():
y = "local"
print(y)
fun()
```
Let's take a look at the cell **In [2]: # Example 3:** where **`x`** was a global variable and we wanted to modify **`x`** inside **`fun()`**.
```
# Exercise 3:
def fun1():
loc_var = 999 # local variable
print("Value is :", loc_var)
def fun2():
print("Value is :", loc_var)
fun1()
fun2()
```
## Global and local variables
Here, we will show how to use global variables and local variables in the same code.
```
# Example 1: Using Global and Local variables in the same code
x = "global"
def fun():
global x
y = "local"
x = x * 2
print(x)
print(y)
fun()
```
**Explanation**:
In the above code, we declare **`x`** as a global and **`y`** as a local variable in the **`fun()`**. Then, we use multiplication operator **`*`** to modify the global variable **`x`** and we print both **`x`** and **`y`**.
After calling the **`fun()`**, the value of **`x`** becomes **`global global`** because we used the **`x * 2`** to print two times **`global`**. After that, we print the value of local variable **`y`** i.e **`local`**.
```
# Example 2: Global variable and Local variable with same name
x = 9
def fun():
x = 19
print("local x:", x)
fun()
print("global x:", x)
```
**Explanation**:
In the above code, we used the same name **`x`** for both global variable and local variable. We get a different result when we print the same variable because the variable is declared in both scopes, i.e. the local scope inside **`fun()`** and global scope outside **`fun()`**.
When we print the variable inside **`fun()`** it outputs **`local x: 19`**. This is called the local scope of the variable.
Similarly, when we print the variable outside the **`fun()`**, it outputs **`global x: 9`**. This is called the global scope of the variable.
```
# Exercise 3:
def my_func(): # for this Function I am not writing any argument in parenthesis '()'
x = 10
print("Value inside the body of function:",x)
x = 20 # first, this line to execute
my_func() # second, the body of function will execute
print("Value outside of function:",x) # finally, this line will execute
```
**Explanation:**
Here, we can see that the value of **`x`** is 20 initially. Even though the function **`my_func()`** changed the value of **`x`** to 10, it did not affect the value outside the function.
This is because the variable **`x`** inside the function is different (local to the function) from the one outside. Although they have the same names, they are two different variables with different scopes.
On the other hand, variables outside of the function are visible from inside. They have a global scope.
We can read these values from inside the function but cannot change (write) them. In order to modify the value of variables outside the function, they must be declared as global variables using the keyword global.
## Nonlocal Variables
Nonlocal variables are used in nested functions whose local scope is not defined. This means that the variable can be neither in the local nor the global scope.
Let's see an example of how a global variable is created in Python.
We use **`nonlocal`** keywords to create nonlocal variables.
```
# Example 1: Create a nonlocal variable
x1 = "global" # Global variable
def outer_fun(): # main function
x1 = "local" # 'x' is local variable for main function and it is nested variable for nested function
print("variable type for Outer function:", x1)
def inner_fun(): # nested fucntion
nonlocal x1 # using local variable 'x' in nested function as nonloval variable
x1 = "nonlocal" # changing the value of my 'x'
print("variable type for Inner function:", x1) # print 'nonlocal'
inner_fun() #print("outer:", x1) # print 'nonlocal'
outer_fun()
print("Variable type of x1:", x1)
```
In the above code, there is a nested **`inner()`** function. We use nonlocal keywords to create a **`nonlocal`** variable. The **`inner()`** function is defined in the scope of another function **`outer()`**.
> **Note**: If we change the value of a nonlocal variable, the changes appear in the local variable.
```
# Exercise 2:
def outer_fun():
x = 999
def inner_fun():
# local variable now acts as global variable
nonlocal x
x = 900
print("value of x inside inner function is:", x)
inner_fun()
print("value of x inside outer function is:", x)
outer_fun()
```
| github_jupyter |
# Experiment on New York Times Dataset
```
%load_ext cython
%%cython -a
import cython
import numpy as np
cimport numpy as np
from libc.math cimport pow, exp
from scipy.special.cython_special cimport psi
@cython.boundscheck(False)
@cython.wraparound(False)
@cython.cdivision(True)
def LDASVI_cython(double[:,:] doc_cnt, int K, double alpha0, double gamma0,
int MB, double eps, double kappa, double tau0, int epoch, int seed, int printcycle):
cdef int D, vocab_size, epoch_len
vocab_size = doc_cnt.shape[0] # number of words in vocabulary
D = doc_cnt.shape[1] # number of documents
epoch_len = D//MB
np.random.seed(seed)
cdef double[:,:] gamma = np.random.rand(vocab_size,K) # initialization of topics
cdef long[:] sample_id
cdef double[:,:] alpha
cdef double diff
cdef double[:] word_phi = np.empty(K)
cdef double word_phi_sum
cdef double[:] tmp_alpha
cdef double[:,:] tmp_gamma
cdef int i,j,k
cdef double[:] psi_sum_gam
cdef double[:] psi_sum_alpha
for ep in range(epoch):
order = np.random.permutation(D)
for t in range(epoch_len):
itr = ep * epoch_len + t
if printcycle!=0 and itr%printcycle==0:
print('starting iteration %i'%itr)
sample_id = order[t*MB:(t+1)*MB]
alpha = np.ones((K,MB)) # initialization of topic proportions
diff = eps + 1
# compute psi(sum(gamma, axis=0))
psi_sum_gam = np.zeros(K)
for k in range(K):
for i in range(vocab_size):
psi_sum_gam[k] += gamma[i,k]
psi_sum_gam[k] = psi(psi_sum_gam[k])
'''E-Step: update local variational parameters(phi,alpha) till convergent'''
while(diff>eps):
diff = 0
tmp_gamma = np.zeros((vocab_size,K))
# compute psi(sum(alpha, axis=0))
psi_sum_alpha = np.zeros(MB)
for i in range(MB):
for k in range(K):
psi_sum_alpha[i] += alpha[k,i]
psi_sum_alpha[i] = psi(psi_sum_alpha[i])
for i in range(MB):
tmp_alpha = np.ones(K) * alpha0
for j in range(vocab_size):
if doc_cnt[j,sample_id[i]] != 0:
word_phi_sum = 0
for k in range(K):
word_phi[k] = exp(psi(gamma[j,k]) - psi_sum_gam[k] + psi(alpha[k,i]) - psi_sum_alpha[i])
word_phi_sum += word_phi[k]
# normalize phi, update tmp_alpha, tmp_gamma
for k in range(K):
word_phi[k] /= word_phi_sum
tmp_alpha[k] += word_phi[k] * doc_cnt[j,sample_id[i]]
tmp_gamma[j,k] += word_phi[k] * doc_cnt[j,sample_id[i]]
# accumulate diff to decide local convergence
for k in range(K):
diff += abs(tmp_alpha[k] - alpha[k,i])
alpha[k,i] = tmp_alpha[k]
diff = diff / K / MB
'''M-Step: update global variational parameters(gamma)'''
rho_t = pow(tau0+itr, -kappa)
for i in range(vocab_size):
for j in range(K):
gamma[i,j] = (1-rho_t)*gamma[i,j] + rho_t*(gamma0+tmp_gamma[i,j]*D/MB)
return gamma # no need to return alpha, since alpha only includes topic proportion of a mini-batch of documents
from sta663_project_lda.preprocessing.gen_nytdata import nytdata_generator
nytdata_generator()
nytdata_mat = np.load('./data/nytdata_mat.npy')
nytdata_voc = np.load('./data/nytdata_voc.npy')
training_params = {
'K':25, # topic_num
'alpha0':0.01, # alpha_prior
'gamma0':0.01, # gamma_prior
'MB':256, # minibatch_size
'kappa':0.5, # learning_rate=(tau0+itr)^(-kappa)
'tau0':256,
'eps':1e-3, # convergence criteria for local updates
}
gamma = LDASVI_cython(nytdata_mat, **training_params, epoch=10, seed=0, printcycle=10)
gamma = np.array(gamma)
from sta663_project_lda.visualization.demo_topics import topic_viz
topic_viz(gamma, nytdata_voc, topk=10)
```
| github_jupyter |
# Self-Driving Car Engineer Nanodegree
## Project: **Finding Lane Lines on the Road**
***
#### I have removed some instructions and intermediate code for students' reference from this page, to make a submission.
#### However, this jupyter notebook is from my branched workspce, created especially for project submission
[`project_submission branch`](https://github.com/mohan-barathi/SD_Car_ND_LaneLines_P1/tree/project_submission)
#### The actual workspce with all my temporary code and student instructions can be found at
[`master branch`](https://github.com/mohan-barathi/SD_Car_ND_LaneLines_P1/tree/master)
**The writeup for this project can be found at :** `../writeup_P1_Lane_Marking.md`
In this project, we will use the tools we learned about in the lesson to identify lane lines on the road. I have developed a pipeline on a series of individual images, and later applied the result to a video stream.
Please refer the video clip "raw-lines-example.mp4" (also contained in this repository) to see what the output should look like, and the output generated as a result of the below work.
**The tools I have are color selection, region of interest selection, grayscaling, Gaussian smoothing, Canny Edge Detection and Hough Tranform line detection. The goal is piece together a pipeline to detect the line segments in the image, then average/extrapolate them and draw them onto the image for display (as below). The pipeline should work on Sample images, and the given video file**
---
<figure>
<img src="examples/line-segments-example.jpg" width="380" alt="Combined Image" />
<figcaption>
<p></p>
<p style="text-align: center;"> The output should look something like this (above) after detecting line segments using the helper functions below </p>
</figcaption>
</figure>
<p></p>
<figure>
<img src="examples/laneLines_thirdPass.jpg" width="380" alt="Combined Image" />
<figcaption>
<p></p>
<p style="text-align: center;"> The goal is to connect/average/extrapolate line segments to get output like this</p>
</figcaption>
</figure>
## Import Packages
```
#importing some useful packages
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import numpy as np
import cv2
%matplotlib inline
```
## Read in an Image
```
#reading in an image
image = mpimg.imread('test_images/solidWhiteRight.jpg')
#printing out some stats and plotting
print('This image is:', type(image), 'with dimensions:', image.shape)
plt.imshow(image) # if you wanted to show a single color channel image called 'gray', for example, call as plt.imshow(gray, cmap='gray')
```
## Helper Functions
Below are some helper functions to help get you started. They should look familiar from the lesson!
### Here, as directed in later part, changes are made in draw_lines() function
#### The changes are too much to place inside a single function, but as directed in the instructions, all the below operations are made within this function
##### The changes include:
* Segregating the lanes as left and right
* Finding the average Gradient for left and right lane lines
* Finding the top most co-ordinate for left and right lanes
* Using the top-most co-ordinate and average slope, the bottom-most co-ordinate for left and right lane are derived
* Respective error handlings
* And finally, drawing these 2 lane lines on given image
```
def draw_lines(img, lines, color=[255, 0, 0], thickness=2):
"""
NOTE: this is the function you might want to use as a starting point once you want to
average/extrapolate the line segments you detect to map out the full
extent of the lane (going from the result shown in raw-lines-example.mp4
to that shown in P1_example.mp4).
Think about things like separating line segments by their
slope ((y2-y1)/(x2-x1)) to decide which segments are part of the left
line vs. the right line. Then, you can average the position of each of
the lines and extrapolate to the top and bottom of the lane.
This function draws `lines` with `color` and `thickness`.
Lines are drawn on the image inplace (mutates the image).
If you want to make the lines semi-transparent, think about combining
this function with the weighted_img() function below
"""
# Declare Left and Right Lanes
left_lanes = []
right_lanes = []
left_slope = 0
right_slope = 0
# setting the Right Bottom of image as initial values
left_x_top = img.shape[1]
left_y_top = img.shape[0]
right_x_top = img.shape[1]
right_y_top = img.shape[0]
# use the slope to separate the lanes accordingly
# and find the average slope and top most point of lane lines
for line in lines:
for x1,y1,x2,y2 in line:
slope = ((y2-y1)/(x2-x1))
# Segregate the line as left and right lane, based on the slope
if (slope<0):
left_lanes.append(line)
left_slope = left_slope + slope
# Handle if the up and down co-ordinates are interchanged for a line
if (y2 < y1):
if (y2 < left_y_top):
left_y_top = y2
left_x_top = x2
else:
if (y1 < left_y_top):
left_y_top = y1
left_x_top = x1
else:
right_lanes.append(line)
right_slope = right_slope + slope
# Handle if the up and down co-ordinates are interchanged for a line
if (y2 < y1):
if (y2 < right_y_top):
right_y_top = y2
right_x_top = x2
else:
if (y1 < right_y_top):
right_y_top = y1
right_x_top = x1
#print ("The 2 top co-ordinates are :", left_x_top, left_y_top, right_x_top, right_y_top)
#cv2.line(img, (left_x_top, left_y_top), (right_x_top, right_y_top), [0,255,0], 30)
# find the average slope of both the lanes
try :
left_average_slope = left_slope/len(left_lanes)
#print ("Average Left lane slope :", left_average_slope)
except(ZeroDivisionError):
print ("No Left lane found") #to-do : Handle the exception upon further calculation
try:
right_average_slope = right_slope/len(right_lanes)
#print ( "Average Right lane slope :", right_average_slope)
except(ZeroDivisionError):
print ("No Right Lane found") #to-do : Handle the exception upon further calculation
# Now we have the top most co-ordinate of the lane, and the slop of the lane
# It's time to find out the bottom co-ordinate, to draw single lane line
# Left-Bottom Co-ordinate, using the function y=mx+b
# left_y_top = left_average_slope*left_x_top + b
try:
b = left_y_top - (left_average_slope*left_x_top)
left_y_bottom = img.shape[0]
left_x_bottom = int((left_y_bottom - b) /left_average_slope) # re-arrange (y=mx+b)
cv2.line(img, (left_x_top, left_y_top), (left_x_bottom, left_y_bottom), [255,0,0], 15)
except(UnboundLocalError):
print ("No Left Lane found in this frame..!")
# In a Similar Way, finding the Right Bottom Co-ordinate
try :
b = right_y_top - (right_average_slope*right_x_top)
right_y_bottom = img.shape[0]
right_x_bottom = int((right_y_bottom - b) /right_average_slope) # re-arrange (y=mx+b)
cv2.line(img, (right_x_top, right_y_top), (right_x_bottom, right_y_bottom), [255,0,0], 15)
except(UnboundLocalError):
print ("No Right Lane found in this Frame..!")
#print ("Left lanes :",len(left_lanes)," Right lanes : ",len(right_lanes))
import math
def grayscale(img):
"""Applies the Grayscale transform
This will return an image with only one color channel
but NOTE: to see the returned image as grayscale
(assuming your grayscaled image is called 'gray')
you should call plt.imshow(gray, cmap='gray')"""
return cv2.cvtColor(img, cv2.COLOR_RGB2GRAY)
# Or use BGR2GRAY if you read an image with cv2.imread()
# return cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
def canny(img, low_threshold, high_threshold):
"""Applies the Canny transform"""
return cv2.Canny(img, low_threshold, high_threshold)
def gaussian_blur(img, kernel_size):
"""Applies a Gaussian Noise kernel"""
return cv2.GaussianBlur(img, (kernel_size, kernel_size), 0)
def region_of_interest(img, vertices):
"""
Applies an image mask.
Only keeps the region of the image defined by the polygon
formed from `vertices`. The rest of the image is set to black.
`vertices` should be a numpy array of integer points.
"""
#defining a blank mask to start with
mask = np.zeros_like(img)
#defining a 3 channel or 1 channel color to fill the mask with depending on the input image
if len(img.shape) > 2:
channel_count = img.shape[2] # i.e. 3 or 4 depending on your image
ignore_mask_color = (255,) * channel_count
else:
ignore_mask_color = 255
#filling pixels inside the polygon defined by "vertices" with the fill color
cv2.fillPoly(mask, vertices, ignore_mask_color)
#returning the image only where mask pixels are nonzero
masked_image = cv2.bitwise_and(img, mask)
return masked_image
def hough_lines(img, rho, theta, threshold, min_line_len, max_line_gap):
"""
`img` should be the output of a Canny transform.
Returns an image with hough lines drawn.
"""
lines = cv2.HoughLinesP(img, rho, theta, threshold, np.array([]), minLineLength=min_line_len, maxLineGap=max_line_gap)
line_img = np.zeros((img.shape[0], img.shape[1], 3), dtype=np.uint8)
draw_lines(line_img, lines)
return line_img
# Python 3 has support for cool math symbols.
def weighted_img(img, initial_img, α=0.8, β=1., γ=0.):
"""
`img` is the output of the hough_lines(), An image with lines drawn on it.
Should be a blank image (all black) with lines drawn on it.
`initial_img` should be the image before any processing.
The result image is computed as follows:
initial_img * α + img * β + γ
NOTE: initial_img and img must be the same shape!
"""
return cv2.addWeighted(initial_img, α, img, β, γ)
```
# Build a Lane Finding Pipeline
The Below pipeline takes raw image as input, uses the above helper functions to draw the lane lines over it, and return the annotated images with lane lines marked.
The output copies of `test_images` are placed into the `test_images_output` directory.
```
# TODO: Build your pipeline that will draw lane lines on the test_images
# then save them to the test_images_output directory.
#plt.imshow(image)
def draw_lane_lines(input_image):
"""
This Function takes in the coloured image as input,
finds the lane in the image using all the helper functions, and draws the lane over the coloured image,
and gives back the resultant image.
"""
#fetch the image shape, and take a copy of the image to work on
ysize = input_image.shape[0]
xsize = input_image.shape[1]
copy_image = np.copy(input_image)
# convert the image to grayscale, and use gaussian blur to smoothen it
gray_image = grayscale(copy_image)
arbitrary_kernal_size = 5
blurred_gray_image = gaussian_blur(gray_image, arbitrary_kernal_size)
# use canny edge detection method to find the edges of the grayscled image
arbitrary_low_threshold = 50
arbitrary_high_threshold = 150
edge_detected_image = canny(blurred_gray_image, arbitrary_low_threshold, arbitrary_high_threshold)
# Mask the parts of the image other than lane, using RoI helper function
mask = np.zeros_like(edge_detected_image)
ignore_mask_color = 255
imshape = copy_image.shape
vertices = np.array([[(0,imshape[0]),(435, 330), (525,330), (imshape[1],imshape[0])]], dtype=np.int32)
cv2.fillPoly(mask, vertices, ignore_mask_color)
masked_edges = cv2.bitwise_and(edge_detected_image, mask)
# applying hough lines
# Define the Hough transform parameters
rho = 2
theta = np.pi/180
threshold = 15
min_line_length = 40
max_line_gap = 20
image_with_hough_line = hough_lines(masked_edges,rho, theta, threshold, min_line_length, max_line_gap)
# The Final image is made by overlaying the hough lines over original image
final_image = weighted_img(image_with_hough_line, copy_image)
return final_image
```
## Test Images
As mentioned in the below cell, there are 6 sample images for testing.
**The next cell iterates through this list, uses the pipeline to annotate the lane lines, and places the output image in test_images_output folder**
```
import os
os.listdir("test_images/")
def iterate_test_image():
tes_images_list = os.listdir("test_images/")
for each_image in tes_images_list:
print (each_image)
test_output = draw_lane_lines(mpimg.imread('test_images/'+each_image))
plt.imshow(test_output)
mpimg.imsave("test_images_output/"+"output_Solid_Lanes_"+each_image, test_output)
iterate_test_image()
```
## Test on Videos
We can test our solution on two provided videos:
`solidWhiteRight.mp4`
`solidYellowLeft.mp4`
**Note: if you get an import error when you run the next cell, try changing your kernel (select the Kernel menu above --> Change Kernel). Still have problems? Try relaunching Jupyter Notebook from the terminal prompt. Also, consult the forums for more troubleshooting tips.**
**If you get an error that looks like this:**
```
NeedDownloadError: Need ffmpeg exe.
You can download it by calling:
imageio.plugins.ffmpeg.download()
```
**Follow the instructions in the error message and check out [this forum post](https://discussions.udacity.com/t/project-error-of-test-on-videos/274082) for more troubleshooting tips across operating systems.**
```
# Import everything needed to edit/save/watch video clips
from moviepy.editor import VideoFileClip
from IPython.display import HTML
def process_image(image):
# NOTE: The output you return should be a color image (3 channel) for processing video below
# TODO: put your pipeline here,
result = draw_lane_lines(image)
# you should return the final output (image where lines are drawn on lanes)
return result
```
Let's try the one with the solid white lane on the right first ...
```
white_output = 'test_videos_output/solidWhiteRight.mp4'
## To speed up the testing process you may want to try your pipeline on a shorter subclip of the video
## To do so add .subclip(start_second,end_second) to the end of the line below
## Where start_second and end_second are integer values representing the start and end of the subclip
## You may also uncomment the following line for a subclip of the first 5 seconds
##clip1 = VideoFileClip("test_videos/solidWhiteRight.mp4").subclip(0,5)
clip1 = VideoFileClip("test_videos/solidWhiteRight.mp4")
white_clip = clip1.fl_image(process_image) #NOTE: this function expects color images!!
%time white_clip.write_videofile(white_output, audio=False)
```
Play the video inline, or if you prefer find the video in your filesystem (should be in the same directory) and play it in your video player of choice.
```
HTML("""
<video width="960" height="540" controls>
<source src="{0}">
</video>
""".format(white_output))
```
Now for the one with the solid yellow lane on the left. This one's more tricky!
```
yellow_output = 'test_videos_output/solidYellowLeft.mp4'
## To speed up the testing process you may want to try your pipeline on a shorter subclip of the video
## To do so add .subclip(start_second,end_second) to the end of the line below
## Where start_second and end_second are integer values representing the start and end of the subclip
## You may also uncomment the following line for a subclip of the first 5 seconds
##clip2 = VideoFileClip('test_videos/solidYellowLeft.mp4').subclip(0,5)
clip2 = VideoFileClip('test_videos/solidYellowLeft.mp4')
yellow_clip = clip2.fl_image(process_image)
%time yellow_clip.write_videofile(yellow_output, audio=False)
HTML("""
<video width="960" height="540" controls>
<source src="{0}">
</video>
""".format(yellow_output))
```
## Optional Challenge
The above approach did not work well with this optional chanllenge.
I'll try to make the above solution fit to this video clip, in near future.
| github_jupyter |
### Real-time human hearing preference acquisition
```
#=================================================
# User's hearing preference data collection (main)
# Author: Nasim Alamdari
# Date: Dec. 2020
#=================================================
import math
import argparse
import sys
from PyQt5.QtGui import *
from PyQt5.QtWidgets import *
from PyQt5.QtCore import *
from IPython import get_ipython
import threading
from multiprocessing import Queue
import time
from time import sleep
import sounddevice as sd
import soundfile as sf
from copy import copy
from fittingEnv import *
from askHuman_realtime import *
from interface import *
# ### Main: Initialization
env_name = "fittingEnv"
env = fittingEnv()
obs_agent, obs_RePrd, audio = env.reset()
action_is_box = type(env.action_space) == gym.spaces.box.Box
if action_is_box:
action_space_n = np.sum(env.action_space.shape)
else:
action_space_n = env.action_space.n
datetime_str = str(datetime.now())
min_Datasize = 2 # minimum data collection
askH_model = askHuman_realtime(datetime_str, min_Datasize)
### Reward predictor settings
hc_ask_human_freq_episodes1 = 1 #every two episode, ask human preferences
hc_trijectory_interval = 1
hc_max = 2800 #10 segment is used for traning reward predictor
trijectory = None # create empty array
run_id = 0 # number of episodes (actually all is one eposodes, there is not reset condition in this project)
nIter = 0 # number of steps
sess_maxIter = 1 # it was 3 before
# ## Run the Main
# window allocation for human interface :
if QApplication.instance() is None:
app = QApplication(sys.argv)
%gui qt5 ## enable PyQt5 event loop integration for .pynb files
#get_ipython().run_line_magic('gui', 'qt5 ## enable PyQt5 event loop integration')
cnt = 0;
cnt_max = 30
while True:
done=False
run_start = nIter
## Ask human
if run_id > 40 and run_id % hc_ask_human_freq_episodes1 == 0:
cnt+=1
print("Counting human feedback: ", cnt)
askH_model.ask_human()
## Trijectory
trijectory = []
while done == False:
# just for testing reward predictor, action is random (instead of using DRL agent):
action = np.random.randint(action_space_n)
#obs_agent, obs_RePrd, audio, adjustCR, _, done, info = env.step(action)
env.take_action(action)
adjustCR = np.reshape(env.adjustVal, (5,))
if trijectory != None:
trijectory.append(copy(adjustCR)) # add compression ratio adjustment for Pref Platform
nIter += 1
if done or nIter - run_start > sess_maxIter:
print("Session ends")
if trijectory != None:
print("Add trij")
askH_model.add_trijactory(trijectory)
trijectory = None #empty session's trijactory
break
run_id+=1
if cnt > cnt_max: # end of asking preferences
break
print("nIter = ",nIter)
print("run_id = ", run_id)
print("===================================")
## Collect and Save all the data and preferences
askH_model.collect_preferences()
print("***** Data collection from human is completed!******")
A=1
B = 1
```
| github_jupyter |
# Classification of b-quark jets in the Aleph simulated data
Python macro for selecting b-jets in Aleph Z->qqbar MC using (kernel) Principle Component Analysis, (k)PCA.
These are among the simplest **unsupervised learning** techniques, and can be considered a quick way to look at new high dimensional data, without knowing much about it. However, notice that cleaning and pre-processing might be needed, as NaN/9999 values will ruin the approach.
### Data:
The input variables (X) are:
* energy: Measured energy of the jet in GeV. Should be 45 GeV, but fluctuates.
* cTheta: cos(theta), i.e. the polar angle of the jet with respect to the beam axis.
The detector works best in the central region (|cTheta| small) and less well in the forward regions.
* phi: The azimuth angle of the jet. As the detector is uniform in phi, this should not matter (much).
* prob_b: Probability of being a b-jet from the pointing of the tracks to the vertex.
* spheri: Sphericity of the event, i.e. how spherical it is.
* pt2rel: The transverse momentum squared of the tracks relative to the jet axis, i.e. width of the jet.
* multip: Multiplicity of the jet (in a relative measure).
* bqvjet: b-quark vertex of the jet, i.e. the probability of a detached vertex.
* ptlrel: Transverse momentum (in GeV) of possible lepton with respect to jet axis (about 0 if no leptons).
The target variable (Y) - **which is not to be used here for anything other than final evaluation** is:
* isb: 1 if it is from a b-quark and 0, if it is not.
### Task:
Thus, in the following we apply PCA and kPCA to the Aleph b-jet data.
* Authors: Troels C. Petersen, Carl Johnsen (NBI)
* Email: petersen@nbi.dk, cjjohnsen@nbi.ku.dk
* Date: 3rd of May 2021
```
from __future__ import print_function, division # Ensures Python3 printing & division standard
from matplotlib import pyplot as plt
from matplotlib import colors
from matplotlib.colors import LogNorm
import numpy as np
import csv
from sklearn.preprocessing import quantile_transform
from sklearn.decomposition import PCA
from sklearn.decomposition import KernelPCA
r = np.random
r.seed(42)
SavePlots = False
plt.close('all')
```
***
# PCA:
```
Btag1 = np.loadtxt('../Week1/AlephBtag_MC_small_v2.csv',skiprows=1)
X = Btag1[:,:-1]
y = Btag1[:,-1]
print ('First X entry: ', X[0])
X_scaled = quantile_transform(X, copy=True)
print ('First scaled X entry: ', X_scaled[0])
pca = PCA(n_components=2)
X_pca = pca.fit_transform(X_scaled)
print ('First PCA X entry:', X_pca[0])
print (pca.explained_variance_ratio_)
fig, ax = plt.subplots(figsize=(12, 8))
truths = np.array([X_pca[i] for i in range(len(y)) if y[i] == 1])
falses = np.array([X_pca[i] for i in range(len(y)) if y[i] != 1])
plt.scatter(falses[:,0], falses[:,1], color='blue', alpha=.1)
plt.scatter(truths[:,0], truths[:,1], color='red', alpha=.1)
plt.show()
kpca = KernelPCA(n_components=2)
kpca.fit(X)
kpca_x = kpca.transform(X)
truths = np.array([kpca_x[i] for i in range(len(y)) if y[i] == 1])
falses = np.array([kpca_x[i] for i in range(len(y)) if y[i] != 1])
plt.scatter(truths[:,0], truths[:,1], color='red', alpha=.1)
plt.scatter(falses[:,0], falses[:,1], color='blue', alpha=.1)
plt.show()
```
| github_jupyter |
####################################
# Bounding boxes visualization logic - Hawaii region
####################################
```
from matplotlib import pyplot as plt
import matplotlib.dates as mdates
from matplotlib.patches import Rectangle
from datetime import datetime, timedelta
import numpy as np
import pandas as pd
import xarray as xr
import zarr
import fsspec
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
plt.rcParams['figure.figsize'] = 12,8
################################
# Read numpy arrays of heat events data
################################
# This notebook uses the output of daymet_hawaii_early_extremes,
# and implements the next func. These are the size arrays that hold
# the heat events detected by 1-0.
# here, now I read this persisted data to write the second part of the algorithm:
# how to fuse these pixel-wise detected heat events into single event and its extend in
# x,y coordinated (not time, yet).
# Pick one file, for instance 1998.
year = 1998
arr3d = np.load(f'./arr_heat3d/arr_heat3d-{year}.npy')
np.where(arr3d==1) # pre-labeled heat events
################################
# progress of an example heat event
################################
t_indexes = range(121,128)
dr = pd.date_range(start=f'1/1/{year}', periods=365, freq='D').date
fig, axs = plt.subplots(1, len(t_indexes), figsize=(24,12))
for i, idx in enumerate(t_indexes):
H = arr3d[idx,:,:]
ax = axs.ravel()[i]
ax.imshow(H, interpolation='none')
ax.set_title(dr[idx])
################################
# find boundary boxes
################################
# uses parent-child hierarchy to merge small areas into fewer heat events
# runs for a single time slice (day)
# later, we will find a way to fuse these boundary bozes in time axis (3rd dimension)
# to find our objective: 'heat events' in 3d,
# defined by ((x1,y1),(x2,y2),(t1,t2)) 3d slices
# OpenCV Contours Hierarchy documentation:
# https://vovkos.github.io/doxyrest-showcase/opencv/sphinxdoc/page_tutorial_py_contours_hierarchy.html#doxid-d9-d8b-tutorial-py-contours-hierarchy
# !pip install opencv-python-headless
import cv2
from matplotlib.patches import Rectangle
from typing import List, Tuple
def bounding_boxes(arr2d: np.array) -> List[tuple]:
H = arr2d.astype(np.uint8)
ret, thresh = cv2.threshold(H, 0, 1, 0, cv2.THRESH_BINARY)
contours, hier = cv2.findContours(thresh, cv2.RETR_EXTERNAL, cv2.CHAIN_APPROX_SIMPLE)
boxes = [cv2.boundingRect(c) for c in contours]
return boxes
fig, axs = plt.subplots(1, len(t_indexes), figsize=(24,12))
for i, idx in enumerate(t_indexes):
arr2d = arr3d[idx,:,:]
ax = axs.ravel()[i]
ax.imshow(arr2d, interpolation='none')
ax.set_title(dr[idx])
boxes = bounding_boxes(arr2d)
for b in boxes:
x,y,w,h = b
rect = Rectangle((x, y), w, h, color='red', fill=False, linewidth=2)
ax.add_patch(rect)
################################
# loop to filter tiny windows and fuse overlapping ones
################################
import itertools
def isoverlap(box1:tuple, box2:tuple) -> bool:
"""Return True if two windows overlap"""
x1,y1,w1,h1 = box1
x2,y2,w2,h2 = box2
return not (x2>x1+w1 or x2+w2<x1 or y2>y1+h1 or y2+h2<y1)
assert(isoverlap((10,10,10,10), (9,9,10,10)) == True)
assert(isoverlap((10,10,10,10), (100,100,2,2)) == False)
def outer(box1:tuple, box2:tuple) -> tuple:
"""Fuse two windows into one, parent window."""
x1,y1,w1,h1 = box1
x2,y2,w2,h2 = box2
x = min(x1,x2)
y = min(y1,y2)
w = max(x1+w1,x2+w2)-x
h = max(y1+h1,y2+h2)-y
return (x, y, w, h)
def istiny(box:tuple, min_area:int) -> bool:
x,y,w,h = box
return w*h <= min_area
def filter_tiny_ones(boxes:List[tuple]) -> List[tuple]:
return [c for c in boxes if not istiny(c, 10)]
def collapse(boxes:List[tuple]) -> List[tuple]:
for box1, box2 in itertools.combinations(boxes, 2):
if isoverlap(box1,box2):
boxes.remove(box1)
boxes.remove(box2)
boxes.append(outer(box1,box2))
return collapse(boxes) # recursion
boxes.sort(key=lambda _:_[0])
return boxes
# test
boxes = [(186, 496, 77, 51), (204, 491, 4, 6), (244, 376, 1, 1), (10,12,3,4), (9,9,10,10)]
boxes = filter_tiny_ones(boxes)
boxes = collapse(boxes)
boxes
################################
# First: apply the functions above to the row-windows openCV found:
################################
fig, axs = plt.subplots(1, len(t_indexes), figsize=(24,12))
for i, idx in enumerate(t_indexes):
arr2d = arr3d[idx,:,:]
ax = axs.ravel()[i]
ax.imshow(arr2d, interpolation='none')
ax.set_title(dr[idx])
boxes = bounding_boxes(arr2d)
for b in boxes:
x,y,w,h = b
rect = Rectangle((x, y), w, h, color='red', fill=False, linewidth=1)
ax.add_patch(rect)
boxes2 = collapse(filter_tiny_ones(boxes))
for c in boxes2:
x,y,w,h = c
rect = Rectangle((x, y), w, h, color='green', fill=False, linewidth=2)
ax.add_patch(rect)
################################
# Next: merge these 2d-merged windows, this time in'time' axis, i.e in 3D
################################
"""Walk on time axis and collapse boxes of consequtive days that
hold boxes, i.e. heat event(s). The output is metadata only, with
the start and end time of each heat event detected, and the largest
box that frames the geolocation of its extend.
I.e., the func `collapse` acts on detected windows,
this time along the 3rd, 'time' axis.
For example, an event is detected from 1998-01-01 to 1998-01-05, started in
LA and moved along to San Diago during that 5 days. The rectangle will be
large enough to encapsulate both cities.
pseudo-code be like:
ds.groupBy(heat-event, axis='time').agg(min-date, max-date, encapsulating-box(x,y,w,h))..
In the output, t1 and t2 are inclusive, like [t1,t2]
"""
def array2boxes(arr2d:np.array) -> List[tuple]:
"""Pipeline. Takes a time-slice (2D array) and returns the boxes."""
boxes = bounding_boxes(arr2d)
boxes = filter_tiny_ones(boxes)
boxes = collapse(boxes)
return boxes
# (Later we replaced the below for loop with equal pandas ops, as follows.)
def groupby_heat_events(arr3d:np.array) -> List[dict]:
bucket = []
events = []
num_days = arr3d.shape[0]
for i in range(num_days):
arr2d = arr3d[i,:,:]
boxes = array2boxes(arr2d)
if not boxes and not bucket:
# no boxes that day
pass
elif boxes and not bucket:
# 1st day of an event
t1 = i
bucket += boxes
elif boxes and bucket:
# 2nd, 3rd,.. day of an event
bucket += boxes
if i == num_days-1:
# if the last day of the data was still a heat event
t2 = i
events += [dict(t1=t1, t2=t2, boxes=bucket)]
bucket = []
elif not boxes and bucket:
# last day of an event
t2 = i-1
events += [dict(t1=t1, t2=t2, boxes=bucket)]
bucket = []
else:
raise KeyError("!?")
for e in events:
e['boxes'] = collapse(e['boxes'])
return pd.DataFrame(events)
groupby_heat_events(arr3d)
################################
# Group bounding boxes by their start and end times
################################
# This func does the same, just shorter.
def groupby_heat_events(arr3d:np.array) -> List[dict]:
rows = []
num_days = arr3d.shape[0]
for i in range(num_days):
arr2d = arr3d[i,:,:]
boxes = array2boxes(arr2d)
rows += [dict(time=i, boxes=boxes)]
df = pd.DataFrame(rows)
df['hasEvent'] = df['boxes'].apply(lambda x: len(x)) > 0
df['label'] = df['hasEvent'].diff().ne(False).cumsum()
dff = df[df['hasEvent']]
dfg = dff.groupby('label').agg({
'time':[np.min,np.max],
'boxes':lambda _: collapse(np.sum(_))
}).reset_index()
dfg.columns = ['label', 'i1', 'i2', 'boxes']
dfg = dfg.assign(d1=dr[dfg['i1']], d2=dr[dfg['i2']])
dfg = dfg.drop('label', axis=1)
return dfg
# Lets test on another year!
year = 1993
dr = pd.date_range(start=f'1/1/{year}', periods=365, freq='D').date
arr3d = np.load(f'./arr_heat3d/arr_heat3d-{year}.npy')
df_events = groupby_heat_events(arr3d)
df_events
################################
# Visualize Hawaii data and bounding boxes, year=1993
################################
for ev, (i1, i2) in df_events[['i1','i2']].iterrows():
t_indexes = range(i1,i2+1)
fig, axs = plt.subplots(1, len(t_indexes), figsize=(24,12))
for i, idx in enumerate(t_indexes):
arr2d = arr3d[idx,:,:]
ax = axs.ravel()[i]
ax.imshow(arr2d, interpolation='none')
ax.set_title(dr[idx])
boxes = df_events['boxes'].iloc[ev]
for b in boxes:
x,y,w,h = b
rect = Rectangle((x, y), w, h, color='red', fill=False, linewidth=2)
ax.add_patch(rect)
fig.suptitle(f"Year {year}, Heat Event {ev}", fontsize=14)
fig.tight_layout()
```
| github_jupyter |
# Preprocessing Functions
@author: Régis Gbenou <br>
@email: regis.gbenou@outlook.fr
The aim here is to build a dictionary that collects all of the exchange rates based on US dollars ($) in a given time periode.
## LIBRARIES
```
import numpy as np # library for matrix manipulation.
import re # library for regular expressions.
import requests # library for requesting web pages.
from bs4 import BeautifulSoup # library for getting information from HTML pages.
from ast import literal_eval # library for interpreting python objects.
```
## FUNCTIONS
```
## Recognization of python objects.
def literal_ev(row):
'''
<lit_ev> abréviation for litteral_eval, is a function which interprets
python object, here sets and lists.
Parameters
----------
row : str
Any character string.
Returns
-------
Python object
DESCRIPTION.
'''
if row == "set()": # Instead of '{}' to signal the presence of an empty set there is 'set()'
row = "{}"
else:
pass
return literal_eval(row)
## Getting Currency.
pattern_currency = "\D+" # sequence of non digit characters whose length >= 0
prog_currency = re.compile(pattern_currency) # transforms <pattern_currency> in search criterion.
def get_currency(row, name):
'''
Getting the first non-numerical character sequence of the <row> dictionary
at <name> key.
Parameters
< ----------
row : dict
Dictionary.
name : str
One of the keys of <row>.
Returns
-------
str
Frst non numerical chararcter sequence.
'''
x = name
res = 'non-existent'
if x in list(row):
res= prog_currency.match(row[x]).group(0) # First sequence that matches with <pattern_currency>.
return res
## Getting money amount.
pattern_amount = "(\d+[\d,]*)*\d" # Either sequence of number that can contains comas.
prog_amount = re.compile(pattern_amount) # transforms <pattern_amount> in search criterion.
def get_amount(row, name):
'''
Getting the number the firtst numerical sequence of the <row> dictionnary
at <name> key.
Parameters
----------
row : dict
Dictionary.
name : str
One of the keys of <row>.
Returns
-------
str
First numerical sequence.
'''
x = name
res = np.nan
if x in list(row):
y = prog_amount.search(row[x]).group(0) # First sequence that matches with <pattern_amount>.
res = int(re.sub(",", "", y))
return res
## Standadization
def build_dict_currency(year_start, year_end):
'''
<build_dict_currency> builds a dictionary collecting all of available exchange
rates in the https://www.xe.com/ website at each year between <year_start>
and <year_end>. Precisely at the date year-01-25.
Parameters
----------
year_start : int
The firtst year from which we want to collect exchange rates.
year_end : int
The last year from which we want to collect exchange rates.
Returns
-------
set_set : list
Dictionary indexed by year and containing at each entry a dictionary
that contains all of available exchange rates at the given year
between <year_start> and <year_end>.
'''
set_set = dict()
for year in np.arange(year_start, year_end+1):
page = requests.get("https://www.xe.com/currencytables/?from=USD&"+\
f"date={year}-12-31")
soup = BeautifulSoup(page.content, "html.parser") # accesses to the html page structure.
""" Use of tags to found the intersting data. """
body = soup.body
content = body.find(id="content")
frame_table = content.find(attrs=({"class": "historicalRateTable-wrap"}))
table = frame_table.find(id="historicalRateTbl")
body_table = table.find_all("tr")
table_list = [k.find_all("td") for k in body_table]
set_set[str(year)] = {k[0].get_text(): float(k[3].get_text()) for\
k in table_list[1:]}
set_set[str(year)]["$"] = 1.0
''' Special processing for the currencies that have different name in
our data and the website.'''
currency_list = [k[0].get_text() for k in table_list[1:]]
currency_ue = ['ATS','BYR', 'BGL','DEM', 'EEK', 'ESP', 'FIM', 'FRF', # Some coutries of UE have abandonned their currency for UE currency.
'IEP', 'ITL', 'LTL', 'LVL', 'PTE', 'ROL', 'SKK', 'YUM']
for k in currency_ue:
if k in currency_list: # If they are present in currency list
pass # there is nothing to do.
else: # Otherwise,
set_set[str(year)][k] = set_set[str(year)]['EUR'] # we will associate them to the UE currency.
corr_dict_int = {'NLG':'ANG', 'RUR':'RUB', 'TRL':'TRY', 'VEB':'VEF'} # Some of the currencies in our data have a name different from those present in <currency_list>
for k in corr_dict_int: # besides these currencies appear only from a certain year.
if corr_dict_int[k] in currency_list: # if the equivalent is present
set_set[str(year)][k] = set_set[str(year)][corr_dict_int[k]] # we affect the equivalent exchange rate.
else:
pass
print(f'{year}: downloaded exchange rates.')
return set_set
```
| github_jupyter |
```
%matplotlib inline
import matplotlib
matplotlib.rcParams['image.interpolation'] = 'nearest'
import numpy as np
import matplotlib.pyplot as plt
```
# Image processing and machine learning
Some image processing numerical techniques are very specific to image processing, such as mathematical morphology or anisotropic diffusion segmentation. However, it is also possible to adapt generic machine learning techniques for image processing.
## A short introduction to machine learning
This section is adapted from the [quick start tutorial](http://scikit-learn.org/stable/tutorial/basic/tutorial.html) from the scikit-learn documentation.
In general, a learning problem considers a set of N samples of data and then tries to predict properties of unknown data. If each sample is more than a single number and, for instance, a multi-dimensional entry (aka multivariate data), it is said to have several attributes or features.
Typical machine learning tasks are :
- **classification**: samples belong to two or more classes and we want to learn from already labeled data how to predict the class of unlabeled data. For example, given examples of pixels belonging to an object of interest and background, we want the algorithm to label all the other pixels of the image. Or given images of cats and dogs, we want to label automatically images whether they show cats or dogs.
- **clustering**: grouping together similar samples. For example, given a set of pictures, can we group them automatically by suject (e.g. people, monuments, animals...)?
In image processing, a sample can either be
- a whole image, its features being pixel values, or sub-regions of an image (e.g. for face detection)
- a pixel, its features being intensity values in colorspace, or statistical information about a neighbourhood centered on the pixel,
- a labeled region, e.g. for classifying particles in an image of labels
The only requirement is to create a dataset composed of N samples, of m features each, which can be passed to the **estimators** of scikit-learn.
Let us start with an example, using the **digits dataset** from scikit-learn.
```
from sklearn import datasets
digits = datasets.load_digits()
print(digits)
```
The dataset is a dictionary-like object that holds all the data and some metadata about the data. This data is stored in the ``.data`` member, which is a ``n_samples, n_features`` array. Response variables (if available, as here) are stored in the ``.target member.``
```
print(digits.data.shape)
print(digits.target.shape)
```
From the shape of the ``data`` array, we see that there are 1797 samples, each having 64 features. In fact, these 64 pixels are the raveled values of an 8x8 image. For convenience, the 2D images are also provided as in the ``.images`` member. In a machine learning problem, a sample always consists of a **flat array** of features, which sometimes require reshaping data.
```
print(digits.images.shape)
np.all(digits.data[0].reshape((8, 8)) == digits.images[0])
plt.imshow(digits.images[0], cmap='gray')
print("target: ", digits.target[0])
```
We now use one of scikit-learn's estimators classes in order to predict the digit from an image.
Here we use an SVC (support vector machine classification) classifier, which uses a part of the dataset (the **training set**) to find the best way to separate the different classes. Even without knowing the details of the SVC, we can use it as a black box thanks to the common estimator API of scikit-learn. An estimator is created by initializing an estimator object:
```
from sklearn import svm
clf = svm.SVC(gamma=0.001, C=100.)
```
The estimator is trained from the learning set using its ``.fit`` method.
```
clf.fit(digits.data[:-10], digits.target[:-10])
```
Then the target value of new data is predicted using the ``.predict`` method of the estimator.
```
print(clf.predict(digits.data[-2:]))
fig, axes = plt.subplots(1, 2)
axes[0].imshow(digits.images[-2], cmap='gray')
axes[1].imshow(digits.images[-1], cmap='gray')
```
So far, so good? We completed our first machine learning example!
In the following, we will see how to use machine learning for image processing. We will use different kinds of samples and features, starting from low-level pixel-based features (e.g. RGB color), to mid-level features (e.g. corner, patches of high contrast), and finally to properties of segmented regions.
**Outline**
- Image segmentation using pixel-based features (color and texture)
- Panorama stitching / image registration based on mid-level features
- Classifying labeled objects using their properties
**What we will not cover**
- computer vision: automatic detection / recognition of objects (faces, ...)
**A follow-up by Stéfan after this part** : image classification using deep learning with Keras.
## Thresholding and vector quantization
Image binarization is a common operation. For grayscale images, finding the best threshold for binarization can be a manual operation. Alternatively, algorithms can select a threshold value automatically; which is convenient for computer vision, or for batch-processing a series of images.
Otsu algorithm is the most famous thresholding algorithm. It maximizes the variance between the two segmented groups of pixels. Therefore, it is can be interpreted as a **clustering** algorithm. Samples are pixels and have a single feature, which is their grayscale value.
```
from skimage import data, exposure, filters
camera = data.camera()
hi = exposure.histogram(camera)
val = filters.threshold_otsu(camera)
fig, axes = plt.subplots(1, 2)
axes[0].imshow(camera, cmap='gray')
axes[0].contour(camera, [val], colors='y')
axes[1].plot(hi[1], hi[0])
axes[1].axvline(val, ls='--')
```
How can we transpose the idea of Otsu thresholding to RGB or multichannel images? We can use the k-means algorithm, which aims to partition samples in k clusters, where each sample belongs to the cluster of nearest mean.
Below we show a simple example of k-means clustering, based on the Iris dataset of ``scikit-learn``. Note that the ``KMeans`` estimator
uses a similar API as the SVC we used for digits classification, with the .fit method.
```
# From http://scikit-learn.org/stable/auto_examples/cluster/plot_cluster_iris.html
from mpl_toolkits.mplot3d import Axes3D
from sklearn.cluster import KMeans
np.random.seed(5)
iris = datasets.load_iris()
X = iris.data
y = iris.target
clf = KMeans(n_clusters=3)
fig = plt.figure(figsize=(4, 3))
ax = Axes3D(fig, rect=[0, 0, .95, 1], elev=48, azim=134)
clf.fit(X)
labels = clf.labels_
ax.scatter(X[:, 3], X[:, 0], X[:, 2], c=labels.astype(np.float), cmap='jet')
ax.w_xaxis.set_ticklabels([])
ax.w_yaxis.set_ticklabels([])
ax.w_zaxis.set_ticklabels([])
ax.set_xlabel('Petal width')
ax.set_ylabel('Sepal length')
ax.set_zlabel('Petal length')
```
k-means clustering uses the Euclidean distance in feature space to cluster samples. If we want to cluster together pixels of similar color, the RGB space is not well suited since it mixes together information about color and light intensity. Therefore, we first transform the RGB image into [Lab colorspace](https://en.wikipedia.org/wiki/Lab_color_space), and only use the color channels (a and b) for clustering.
```
from skimage import io, color
im = io.imread('../images/round_pill.jpg')
im_lab = color.rgb2lab(im)
data = np.array([im_lab[..., 1].ravel(), im_lab[..., 2].ravel()])
```
Then we create a ``KMeans`` estimator for two clusters.
```
from sklearn.cluster import KMeans
kmeans = KMeans(n_clusters=2, random_state=0).fit(data.T)
segmentation = kmeans.labels_.reshape(im.shape[:-1])
plt.imshow(im)
plt.contour(segmentation, colors='y')
```
Of course we can generalize this method to more than two clusters.
```
im = io.imread('../images/chapel_floor.png')
im_lab = color.rgb2lab(im)
data = np.array([im_lab[..., 0].ravel(),
im_lab[..., 1].ravel(),
im_lab[..., 2].ravel()])
kmeans = KMeans(n_clusters=4, random_state=0).fit(data.T)
segmentation = kmeans.labels_.reshape(im.shape[:-1])
color_mean = color.label2rgb(segmentation, im, kind='mean')
fig, axes = plt.subplots(1, 2)
axes[0].imshow(im)
axes[0].axis('off')
axes[1].imshow(color_mean)
axes[1].axis('off')
```
### Exercise:
For the chapel floor image, cluster the image in 3 clusters, using only the color channels (not the lightness one). What happens?
## SLIC algorithm: clustering using color and spatial features
In the thresholding / vector quantization approach presented above, pixels are characterized only by their color features. However, in most images neighboring pixels correspond to the same object. Hence, information on spatial proximity between pixels can be used in addition to color information.
SLIC (Simple Linear Iterative Clustering) is a segmentation algorithm which clusters pixels in both space and color. Therefore, regions of space that are similar in color will end up in the same segment.
```
spices = io.imread('../images/spices.jpg')
plt.imshow(spices)
```
Let us try to segment the different spices using the previous k-means approach. One problem is that there is a lot of texture coming from the relief and shades.
```
im_lab = color.rgb2lab(spices)
data = np.array([im_lab[..., 1].ravel(),
im_lab[..., 2].ravel()])
kmeans = KMeans(n_clusters=10, random_state=0).fit(data.T)
labels = kmeans.labels_.reshape(spices.shape[:-1])
color_mean = color.label2rgb(labels, spices, kind='mean')
plt.imshow(color_mean)
from skimage import segmentation
plt.imshow(segmentation.mark_boundaries(spices, labels))
```
SLIC is a superpixel algorithm, which segments an image into patches (superpixels) of neighboring pixels with a similar color. SLIC also works in the Lab colorspace. The ``compactness`` parameter controls the relative importance of the distance in image- and color-space.
```
from skimage import segmentation
segments = segmentation.slic(spices, n_segments=200, compactness=20)
plt.imshow(segmentation.mark_boundaries(spices, segments))
result = color.label2rgb(segments, spices, kind='mean')
plt.imshow(result)
```
After the super-pixel segmentation (which is also called oversegmentation, because we end up with more segments that we want to), we can add a second clustering step to join superpixels belonging to the same spice heap.
```
im_lab = color.rgb2lab(result)
data = np.array([im_lab[..., 1].ravel(),
im_lab[..., 2].ravel()])
kmeans = KMeans(n_clusters=5, random_state=0).fit(data.T)
labels = kmeans.labels_.reshape(spices.shape[:-1])
color_mean = color.label2rgb(labels, spices, kind='mean')
plt.imshow(segmentation.mark_boundaries(spices, labels))
```
Note that other superpixel algorithms are available, such as **Felzenswalb** segmentation.
```
result = segmentation.felzenszwalb(spices, scale=100)
plt.imshow(color.label2rgb(result, spices, kind='mean'))
plt.imshow(segmentation.mark_boundaries(spices, result))
```
### Exercise
Repeat the same operations (SLIC superpixel segmentation, followed by K-Means clustering on the average color of superpixels) on the astronaut image. Vary the following parameters
- slic: n_segments and compactness
- KMeans: n_clusters (start with 8 for example)
```
from skimage import data
astro = data.astronaut()
# solution goes here
```
## Increasing the number of low-level features: trained segmentation using Gabor filters and random forests
In the examples above, a small number of features per pixel was used: either a color triplet only, or a color triplet and its (x, y) position. However, it is possible to use other features, such as the local texture. Texture features can be obtained using Gabor filters, which are Gaussian kernels modulated by a sinusoidal wave.
```
# From http://scikit-image.org/docs/dev/auto_examples/features_detection/plot_gabor.html
from skimage import data, img_as_float
from skimage.filters import gabor_kernel
import scipy.ndimage as ndi
shrink = (slice(0, None, 3), slice(0, None, 3))
brick = img_as_float(data.load('brick.png'))[shrink]
grass = img_as_float(data.load('grass.png'))[shrink]
wall = img_as_float(data.load('rough-wall.png'))[shrink]
image_names = ('brick', 'grass', 'wall')
images = (brick, grass, wall)
def power(image, kernel):
# Normalize images for better comparison.
image = (image - image.mean()) / image.std()
return np.sqrt(ndi.convolve(image, np.real(kernel), mode='wrap')**2 +
ndi.convolve(image, np.imag(kernel), mode='wrap')**2)
# Plot a selection of the filter bank kernels and their responses.
results = []
kernel_params = []
for frequency in (0.1, 0.4):
kernel = gabor_kernel(frequency, theta=0)
params = 'frequency=%.2f' % (frequency)
kernel_params.append(params)
# Save kernel and the power image for each image
results.append((kernel, [power(img, kernel) for img in images]))
fig, axes = plt.subplots(nrows=3, ncols=4, figsize=(5, 4))
plt.gray()
fig.suptitle('Image responses for Gabor filter kernels', fontsize=12)
axes[0][0].axis('off')
# Plot original images
for label, img, ax in zip(image_names, images, axes[0][1:]):
ax.imshow(img)
ax.set_title(label, fontsize=9)
ax.axis('off')
for label, (kernel, powers), ax_row in zip(kernel_params, results, axes[1:]):
# Plot Gabor kernel
ax = ax_row[0]
ax.imshow(np.real(kernel), interpolation='nearest')
ax.set_ylabel(label, fontsize=7)
ax.set_xticks([])
ax.set_yticks([])
# Plot Gabor responses with the contrast normalized for each filter
vmin = np.min(powers)
vmax = np.max(powers)
for patch, ax in zip(powers, ax_row[1:]):
ax.imshow(patch, vmin=vmin, vmax=vmax)
ax.axis('off')
```
We define a segmentation algorithms which:
- computes different features for Gabor filters of different scale and angle, for every pixel
- trains a **RandomForest** classifier from user-labeled data, which are given as a mask of labels
- and predicts the label of the remaining non-labeled pixels
The RandomForest algorithm chooses automatically thresholds along the different feature directions, and also decides which features are the most significant to discriminate between the different classes. This is very useful when we don't know if all features are relevant.
```
from sklearn.ensemble import RandomForestClassifier
from skimage import filters
from skimage import img_as_float
def _compute_features(im):
gabor_frequencies = np.logspace(-3, 1, num=5, base=2)
thetas = [0, np.pi/2]
nb_fq = len(gabor_frequencies) * len(thetas)
im = np.atleast_3d(im)
im_gabor = np.empty((im.shape[-1], nb_fq) + im.shape[:2])
for ch in range(im.shape[-1]):
img = img_as_float(im[..., ch])
for i_fq, fq in enumerate(gabor_frequencies):
for i_th, theta in enumerate(thetas):
tmp = filters.gabor(img, fq, theta=theta)
im_gabor[ch, len(thetas) * i_fq + i_th] = \
np.abs(tmp[0] + 1j * tmp[1])
return im_gabor
def trainable_segmentation(im, mask):
"""
Parameters
----------
im : ndarray
2-D image (grayscale or RGB) to be segmented
mask : ndarray of ints
Array of labels. Non-zero labels are known regions that are used
to train the classification algorithm.
"""
# Define features
im_gabor = _compute_features(im)
nb_ch, nb_fq, sh_1, sh2 = im_gabor.shape
# Training data correspond to pixels labeled in mask
training_data = im_gabor[:, :, mask>0]
training_data = training_data.reshape((nb_ch * nb_fq,
(mask>0).sum())).T
training_labels = mask[mask>0].ravel()
# Data are from the remaining pixels
data = im_gabor[:, :, mask == 0].reshape((nb_ch * nb_fq,
(mask == 0).sum())).T
# classification
clf = RandomForestClassifier()
clf.fit(training_data, training_labels)
labels = clf.predict(data)
result = np.copy(mask)
result[mask == 0] = labels
return result
# Image from https://fr.wikipedia.org/wiki/Fichier:Bells-Beach-View.jpg
beach = io.imread('../images/Bells-Beach.jpg')
# Define mask of user-labeled pixels, which will be used for training
mask = np.zeros(beach.shape[:-1], dtype=np.uint8)
mask[700:] = 1
mask[:550, :650] = 2
mask[400:450, 1000:1100] = 3
plt.imshow(beach)
plt.contour(mask, colors='y')
result = trainable_segmentation(beach, mask)
plt.imshow(color.label2rgb(result, beach, kind='mean'))
```
## Using mid-level features
```
from skimage import data
camera = data.camera()
from skimage import feature
corner_camera = feature.corner_harris(camera)
coords = feature.corner_peaks(corner_camera)
plt.imshow(camera, cmap='gray')
plt.plot(coords[:, 1], coords[:, 0], 'o')
plt.xlim(0, 512)
plt.ylim(512, 0)
```
[Panorama stitching](example_pano.ipynb)
[A longer example](adv3_panorama-stitching.ipynb)
### Exercise
Represent the ORB keypoint of the camera-man
```
# solution goes here
```
## Clustering or classifying labeled objects
We have already seen how to use ``skimage.measure.regionprops`` to extract the properties (area, perimeter, ...) of labeled objects. These properties can be used as features in order to cluster the objects in different groups, or to classify them if given a training set.
In the example below, we use ``skimage.data.binary_blobs`` to generate a binary image. We use several properties to generate features: the area, the ratio between squared perimeter and area, and the solidity (which is the area fraction of the object as compared to its convex hull). We would like to separate the big convoluted particles from the smaller round ones. Here I did not want to bother with a training set, so we will juste use clustering instead of classifying.
```
from skimage import measure
from skimage import data
im = data.binary_blobs(length=1024, blob_size_fraction=0.05,
volume_fraction=0.2)
labels = measure.label(im)
props = measure.regionprops(labels)
data = np.array([(prop.area,
prop.perimeter**2/prop.area,
prop.solidity) for prop in props])
plt.imshow(labels, cmap='spectral')
```
Once again we use the KMeans algorithm to cluster the objects. We visualize the result as an array of labels.
```
clf = KMeans(n_clusters=2)
clf.fit(data)
def reshape_cluster_labels(cluster_labels, image_labels):
"""
Some NumPy magic
"""
cluster_labels = np.concatenate(([0], cluster_labels + 1))
return cluster_labels[image_labels]
object_clusters = reshape_cluster_labels(clf.labels_, labels)
plt.imshow(object_clusters, cmap='spectral')
```
However, our features were not carefully designed. Since the ``area`` property can take much larger values than the other properties, it dominates the other ones. To correct this effect, we can normalize the area to its maximal value.
```
data[:, 0] /= data[:, 0].max()
clf.fit(data)
object_clusters = reshape_cluster_labels(clf.labels_, labels)
plt.imshow(object_clusters, cmap='spectral')
```
A better way to do the rescaling is to use of the scaling methods provided by ``sklearn.preprocessing``. The ``StandardScaler`` makes sure that every feature has a zero mean and a unit standard deviation.
```
from sklearn import preprocessing
min_max_scaler = preprocessing.StandardScaler()
data_scaled = min_max_scaler.fit_transform(data)
clf = KMeans(n_clusters=2)
clf.fit(data_scaled)
object_clusters = reshape_cluster_labels(clf.labels_, labels)
plt.imshow(object_clusters, cmap='spectral')
```
###Exercise
Replace the area property by the eccentricity, so that clustering separates compact and convoluted particles, regardless of their size.
| github_jupyter |
# Self-Driving Car Engineer Nanodegree
## Deep Learning
## Project: Build a Traffic Sign Recognition Classifier
In this notebook, a template is provided for you to implement your functionality in stages, which is required to successfully complete this project. If additional code is required that cannot be included in the notebook, be sure that the Python code is successfully imported and included in your submission if necessary.
> **Note**: Once you have completed all of the code implementations, you need to finalize your work by exporting the iPython Notebook as an HTML document. Before exporting the notebook to html, all of the code cells need to have been run so that reviewers can see the final implementation and output. You can then export the notebook by using the menu above and navigating to \n",
"**File -> Download as -> HTML (.html)**. Include the finished document along with this notebook as your submission.
In addition to implementing code, there is a writeup to complete. The writeup should be completed in a separate file, which can be either a markdown file or a pdf document. There is a [write up template](https://github.com/udacity/CarND-Traffic-Sign-Classifier-Project/blob/master/writeup_template.md) that can be used to guide the writing process. Completing the code template and writeup template will cover all of the [rubric points](https://review.udacity.com/#!/rubrics/481/view) for this project.
The [rubric](https://review.udacity.com/#!/rubrics/481/view) contains "Stand Out Suggestions" for enhancing the project beyond the minimum requirements. The stand out suggestions are optional. If you decide to pursue the "stand out suggestions", you can include the code in this Ipython notebook and also discuss the results in the writeup file.
>**Note:** Code and Markdown cells can be executed using the **Shift + Enter** keyboard shortcut. In addition, Markdown cells can be edited by typically double-clicking the cell to enter edit mode.
---
## Import all necessary packages
```
import pickle
from sklearn.utils import shuffle
import numpy as np
import random
import csv
import matplotlib.pyplot as plt
%matplotlib inline
from textwrap import wrap
import cv2
import tensorflow as tf
from tensorflow.contrib.layers import flatten
```
---
## Step 0: Load The Data
```
# Load pickled data
# TODO: Fill this in based on where you saved the training and testing data
# define input files
training_file = '../../GD_GitHubData/traffic-signs-data/train.p'
validation_file = '../../GD_GitHubData/traffic-signs-data/valid.p'
testing_file = '../../GD_GitHubData/traffic-signs-data/test.p'
# load inputs from input files
with open(training_file, mode='rb') as f:
train = pickle.load(f)
with open(validation_file, mode='rb') as f:
valid = pickle.load(f)
with open(testing_file, mode='rb') as f:
test = pickle.load(f)
# get data from inputs
X_train, y_train = train['features'], train['labels']
X_valid, y_valid = valid['features'], valid['labels']
X_test, y_test = test['features'], test['labels']
# check data consistency
assert(len(X_train) == len(y_train))
assert(len(X_valid) == len(y_valid))
assert(len(X_test) == len(y_test))
```
---
## Step 1: Dataset Summary & Exploration
The pickled data is a dictionary with 4 key/value pairs:
- `'features'` is a 4D array containing raw pixel data of the traffic sign images, (num examples, width, height, channels).
- `'labels'` is a 1D array containing the label/class id of the traffic sign. The file `signnames.csv` contains id -> name mappings for each id.
- `'sizes'` is a list containing tuples, (width, height) representing the original width and height the image.
- `'coords'` is a list containing tuples, (x1, y1, x2, y2) representing coordinates of a bounding box around the sign in the image. **THESE COORDINATES ASSUME THE ORIGINAL IMAGE. THE PICKLED DATA CONTAINS RESIZED VERSIONS (32 by 32) OF THESE IMAGES**
Complete the basic data summary below. Use python, numpy and/or pandas methods to calculate the data summary rather than hard coding the results. For example, the [pandas shape method](http://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.shape.html) might be useful for calculating some of the summary results.
### Provide a Basic Summary of the Data Set Using Python, Numpy and/or Pandas
```
### Replace each question mark with the appropriate value.
### Use python, pandas or numpy methods rather than hard coding the results
# TODO: Number of training examples
n_train = X_train[:,0,0,0].shape[0]
# TODO: Number of validation examples
n_validation = X_valid[:,0,0,0].shape[0]
# TODO: Number of testing examples.
n_test = X_test[:,0,0,0].shape[0]
# TODO: What's the shape of an traffic sign image?
image_shape = X_train[0,:,:,:].shape if (X_train[0,:,:,:].shape == X_valid[0,:,:,:].shape == X_test[0,:,:,:].shape) else []
# TODO: How many unique classes/labels there are in the dataset.
l_classes = []
n_classes = len([l_classes.append(label) for label in np.concatenate((y_train, y_valid, y_test)) if label not in l_classes])
print("Number of training examples =", n_train)
print("Number of validation examples =", n_validation)
print("Number of testing examples =", n_test)
print("Image data shape =", image_shape)
print("Number of classes =", n_classes)
```
### Include an exploratory visualization of the dataset
Visualize the German Traffic Signs Dataset using the pickled file(s). This is open ended, suggestions include: plotting traffic sign images, plotting the count of each sign, etc.
The [Matplotlib](http://matplotlib.org/) [examples](http://matplotlib.org/examples/index.html) and [gallery](http://matplotlib.org/gallery.html) pages are a great resource for doing visualizations in Python.
**NOTE:** It's recommended you start with something simple first. If you wish to do more, come back to it after you've completed the rest of the sections. It can be interesting to look at the distribution of classes in the training, validation and test set. Is the distribution the same? Are there more examples of some classes than others?
```
### Data exploration visualization code goes here.
### Feel free to use as many code cells as needed.
# Visualizations will be shown in the notebook.
# define constants
displayimages = 30
def plot_traffic_signs(images, labels, totalimages):
# ...
# This function plots a set of traffic sign images
# ...
# Inputs
# ...
# images : list of images
# labels : list of image labels
# totalimages : total number of images to plot
# define constants
maxhorizontalimages = 10
dpi = 80
horizontalimagesize = 15
titlebasefontsize = 100
titlechars_per_line = 18
# initialize variables
horizontalimages = min(totalimages, maxhorizontalimages)
# create figure with subplot
verticalimages = np.int(np.ceil(totalimages / horizontalimages))
verticalimagesize = (horizontalimagesize * (verticalimages / horizontalimages))
figure, axes = plt.subplots(verticalimages, horizontalimages, figsize=(horizontalimagesize, verticalimagesize), dpi = dpi)
figure.tight_layout()
# figure.subplots_adjust(left=None, bottom=None, right=None, top=None, wspace=None, hspace=None)
axes = axes.reshape(-1)
# plot all images
titlefontsize = (titlebasefontsize / horizontalimages)
for idx, axis in enumerate(axes):
# configure axis
axis.set_axis_off()
axis.get_xaxis().set_visible(False)
axis.get_yaxis().set_visible(False)
# print label and plot image
if (len(labels) > idx):
axis.set_title("\n".join(wrap(labels[idx], titlechars_per_line)), fontsize = titlefontsize)
axis.imshow(images[idx, :, :])
# make sure plot is shown
plt.show()
# read description of labels
with open('signnames.csv') as csvfile:
reader = csv.DictReader(csvfile)
labeldict = dict([(np.int(row['ClassId']), row['SignName']) for row in reader])
# select and plot random images
print('Random images:')
indices = np.random.randint(0, len(X_train), displayimages)
images = X_train[indices].squeeze()
labels = [labeldict[idx] for idx in y_train[indices]]
plot_traffic_signs(images, labels, displayimages)
# select and plot random images of the same random label
label = np.random.randint(min(y_train), max(y_train), 1)
print('Random images of the same random label:', labeldict[label[0]])
images = np.asarray([image for idx, image in enumerate(X_train) if (y_train[idx] == label)])
indices = np.random.randint(0, images.shape[0], displayimages)
images = images[indices, :, :]
labels = [labeldict[label[0]] for image in images]
plot_traffic_signs(images, labels, displayimages)
# select and plot a high contrast image for each label
labelnums = np.arange(min(y_train), (max(y_train) + 1))
contrastimages = []
averageimages = []
for labelnum in labelnums:
labelimages = np.asarray([image for idx, image in enumerate(X_train) if (y_train[idx] == labelnum)])
grays = np.asarray([cv2.cvtColor(labelimage, cv2.COLOR_RGB2GRAY) for labelimage in labelimages])
smalls = [gray[15:18, 15:18] for gray in grays]
deviations = np.asarray([np.sum(np.absolute(small - np.average(small))) for small in smalls])
contrastimages.append(labelimages[np.argmax(deviations)])
averageimages.append(np.array(np.average(labelimages, axis = 0), dtype = np.int32))
contrastimages = np.asarray(contrastimages)
averageimages = np.asarray(averageimages)
labels = [labeldict[labelnums[idx]] for idx, labelnum in enumerate(labelnums)]
print('High contrast image for each label:')
plot_traffic_signs(contrastimages, labels, len(labelnums))
print('Average image for each label:')
plot_traffic_signs(averageimages, labels, len(labelnums))
```
### Modify existing data to extend the database and visualize it
ToDo
----
## Step 2: Design and Test a Model Architecture
Design and implement a deep learning model that learns to recognize traffic signs. Train and test your model on the [German Traffic Sign Dataset](http://benchmark.ini.rub.de/?section=gtsrb&subsection=dataset).
The LeNet-5 implementation shown in the [classroom](https://classroom.udacity.com/nanodegrees/nd013/parts/fbf77062-5703-404e-b60c-95b78b2f3f9e/modules/6df7ae49-c61c-4bb2-a23e-6527e69209ec/lessons/601ae704-1035-4287-8b11-e2c2716217ad/concepts/d4aca031-508f-4e0b-b493-e7b706120f81) at the end of the CNN lesson is a solid starting point. You'll have to change the number of classes and possibly the preprocessing, but aside from that it's plug and play!
With the LeNet-5 solution from the lecture, you should expect a validation set accuracy of about 0.89. To meet specifications, the validation set accuracy will need to be at least 0.93. It is possible to get an even higher accuracy, but 0.93 is the minimum for a successful project submission.
There are various aspects to consider when thinking about this problem:
- Neural network architecture (is the network over or underfitting?)
- Play around preprocessing techniques (normalization, rgb to grayscale, etc)
- Number of examples per label (some have more than others).
- Generate fake data.
Here is an example of a [published baseline model on this problem](http://yann.lecun.com/exdb/publis/pdf/sermanet-ijcnn-11.pdf). It's not required to be familiar with the approach used in the paper but, it's good practice to try to read papers like these.
### Pre-process the Data Set (normalization, grayscale, etc.)
Minimally, the image data should be normalized so that the data has mean zero and equal variance. For image data, `(pixel - 128)/ 128` is a quick way to approximately normalize the data and can be used in this project.
Other pre-processing steps are optional. You can try different techniques to see if it improves performance.
Use the code cell (or multiple code cells, if necessary) to implement the first step of your project.
```
### Preprocess the data here. It is required to normalize the data. Other preprocessing steps could include
### converting to grayscale, etc.
### Feel free to use as many code cells as needed.
# normalize input data
X_train_norm = np.asarray(((X_train / 127.5) - 1), np.float32)
X_valid_norm = np.asarray(((X_valid / 127.5) - 1), np.float32)
X_test_norm = np.asarray(((X_test / 127.5) - 1), np.float32)
# ensure correct label data type
y_train_conv = np.asarray(y_train, np.int32)
y_valid_conv = np.asarray(y_valid, np.int32)
y_test_conv = np.asarray(y_test, np.int32)
```
### Model Architecture
```
### Define your architecture here.
### Feel free to use as many code cells as needed.
def LeNet(x, bdisplay):
# ...
# This function calculates the LeNet logits for input x
# ...
# Inputs
# ...
# x : input image
# bdisplay : boolean for 'display information'
# ...
# Outputs
# ...
# logits : probabilities of individual classes in input image
# Arguments used for tf.truncated_normal, randomly defines variables for the weights and biases for each layer
mu = 0
sigma = 0.1
# SOLUTION: Layer 1: Convolutional. Input = 32x32x3. Output = 28x28x6.
conv1_W = tf.Variable(tf.truncated_normal(shape = (5, 5, 3, 6), mean = mu, stddev = sigma))
conv1_b = tf.Variable(tf.zeros(6))
conv1 = tf.nn.conv2d(x, conv1_W, strides = [1, 1, 1, 1], padding = 'VALID') + conv1_b
# display information
if bdisplay:
print('Conv: shape', (5, 5, 3, 6), 'zeros', 6, 'mean', mu, 'stddev', sigma)
print('Conv: strides', [1, 1, 1, 1], 'padding', 'VALID')
print('Conv: inp', x)
print('Conv: W', conv1_W)
print('Conv: b', conv1_b)
print('Conv: out', conv1)
# SOLUTION: Activation.
conv1 = tf.nn.relu(conv1)
print('Convolutional layer 1 :', [32, 32], 'input dimension with depth', 3, 'and', [28, 28], 'output dimensions with depth', 6)
# SOLUTION: Pooling. Input = 28x28x6. Output = 14x14x6.
conv1 = tf.nn.max_pool(conv1, ksize = [1, 2, 2, 1], strides = [1, 2, 2, 1], padding = 'VALID')
# display information
if bdisplay:
print('Pool: filter', [1, 2, 2, 1], 'strides', [1, 2, 2, 1], 'padding', 'VALID')
print('Pool: inp', conv1)
print('Pool: out', conv1)
print('Pooling layer 1 :', [28, 28], 'input dimension with depth', 6, 'and', [14, 14], 'output dimensions with depth', 6)
# SOLUTION: Layer 2: Convolutional. Output = 10x10x16.
conv2_W = tf.Variable(tf.truncated_normal(shape = (5, 5, 6, 16), mean = mu, stddev = sigma))
conv2_b = tf.Variable(tf.zeros(16))
conv2 = tf.nn.conv2d(conv1, conv2_W, strides = [1, 1, 1, 1], padding = 'VALID') + conv2_b
# display information
if bdisplay:
print('Conv: shape', [5, 5, 6, 16], 'zeros', 16, 'mean', mu, 'stddev', sigma)
print('Conv: strides', [1, 1, 1, 1], 'padding', 'VALID')
print('Conv: inp', conv1)
print('Conv: W', conv2_W)
print('Conv: b', conv2_b)
print('Conv: out', conv2)
# SOLUTION: Activation.
conv2 = tf.nn.relu(conv2)
print('Convolutional layer 2 :', [14, 14], 'input dimension with depth', 6, 'and', [10, 10], 'output dimensions with depth', 16)
# SOLUTION: Pooling. Input = 10x10x16. Output = 5x5x16.
conv2 = tf.nn.max_pool(conv2, ksize = [1, 2, 2, 1], strides = [1, 2, 2, 1], padding = 'VALID')
# display information
if bdisplay:
print('Pool: filter', [1, 2, 2, 1], 'strides', [1, 2, 2, 1], 'padding', 'VALID')
print('Pool: inp', conv2)
print('Pool: out', conv2)
print('Pooling layer 2 :', [10, 10], 'input dimension with depth', 16, 'and', [5, 5], 'output dimensions with depth', 16)
# SOLUTION: Flatten. Input = 5x5x16. Output = 400.
fc0 = flatten(conv2)
# SOLUTION: Layer 3: Fully Connected. Input = 400. Output = 120.
fc1_W = tf.Variable(tf.truncated_normal(shape=(400, 120), mean = mu, stddev = sigma))
fc1_b = tf.Variable(tf.zeros(120))
fc1 = tf.matmul(fc0, fc1_W) + fc1_b
# display information
if bdisplay:
print('Full: shape', [400, 120], 'zeros', 120, 'mean', mu, 'stddev', sigma)
print('Full: inp', fc0)
print('Full: W', fc1_W)
print('Full: b', fc1_b)
print('Full: fc', fc1)
# SOLUTION: Activation.
fc1 = tf.nn.relu(fc1)
print('Fully connected layer 1 :', 400, 'input dimensions and', 120, 'output dimensions')
# SOLUTION: Layer 4: Fully Connected. Input = 120. Output = 84.
fc2_W = tf.Variable(tf.truncated_normal(shape = (120, 84), mean = mu, stddev = sigma))
fc2_b = tf.Variable(tf.zeros(84))
fc2 = tf.matmul(fc1, fc2_W) + fc2_b
# display information
if bdisplay:
print('Full: shape', [120, 84], 'zeros', 84, 'mean', mu, 'stddev', sigma)
print('Full: inp', fc1)
print('Full: W', fc2_W)
print('Full: b', fc2_b)
print('Full: fc', fc2)
# SOLUTION: Activation.
fc2 = tf.nn.relu(fc2)
print('Fully connected layer 2 :', 120, 'input dimensions and', 84, 'output dimensions')
# SOLUTION: Layer 5: Fully Connected. Input = 84. Output = 43.
fc3_W = tf.Variable(tf.truncated_normal(shape = (84, 43), mean = mu, stddev = sigma))
fc3_b = tf.Variable(tf.zeros(43))
logits = tf.matmul(fc2, fc3_W) + fc3_b
# display information
if bdisplay:
print('Full: shape', [84, 43], 'zeros', 43, 'mean', mu, 'stddev', sigma)
print('Full: inp', fc2)
print('Full: W', fc3_W)
print('Full: b', fc3_b)
print('Full: fc', logits)
print('Fully connected layer 3 :', 84, 'input dimensions and', 43, 'output dimensions')
return logits
### Define your architecture here.
### Feel free to use as many code cells as needed.
def LeNet_adjusted(x, tf_keep_prob, bdisplay):
# ...
# This function calculates the LeNet logits for input x
# ...
# Inputs
# ...
# x : input image
# bdisplay : boolean for 'display information'
# ...
# Outputs
# ...
# logits : probabilities of individual classes in input image
# Arguments used for tf.truncated_normal, randomly defines variables for the weights and biases for each layer
mu = 0
sigma = 0.1
# Layer 1: Convolutional. Input = 32x32x3. Output = 28x28x18.
conv1_W = tf.Variable(tf.truncated_normal(shape = (5, 5, 3, 18), mean = mu, stddev = sigma))
conv1_b = tf.Variable(tf.zeros(18))
conv1 = tf.nn.conv2d(x, conv1_W, strides = [1, 1, 1, 1], padding = 'VALID') + conv1_b
# display information
if bdisplay:
print('Conv: shape', (5, 5, 3, 18), 'zeros', 18, 'mean', mu, 'stddev', sigma)
print('Conv: strides', [1, 1, 1, 1], 'padding', 'VALID')
print('Conv: inp', x)
print('Conv: W', conv1_W)
print('Conv: b', conv1_b)
print('Conv: out', conv1)
# Activation and drop out
conv1 = tf.nn.relu(conv1)
conv1 = tf.nn.dropout(conv1, tf_keep_prob)
print('Convolutional layer 1 :', [32, 32], 'input dimension with depth', 3, 'and', [28, 28], 'output dimensions with depth', 18)
# Pooling. Input = 28x28x18. Output = 14x14x18.
#conv1 = tf.nn.max_pool(conv1, ksize = [1, 2, 2, 1], strides = [1, 2, 2, 1], padding = 'VALID')
# display information
#if bdisplay:
# print('Pool: filter', [1, 2, 2, 1], 'strides', [1, 2, 2, 1], 'padding', 'VALID')
# print('Pool: inp', conv1)
# print('Pool: out', conv1)
#print('Pooling layer 1 :', [28, 28], 'input dimension with depth', 18, 'and', [14, 14], 'output dimensions with depth', 18)
# Layer 2: Convolutional. Output = 24x24x54.
conv2_W = tf.Variable(tf.truncated_normal(shape = (5, 5, 18, 54), mean = mu, stddev = sigma))
conv2_b = tf.Variable(tf.zeros(54))
conv2 = tf.nn.conv2d(conv1, conv2_W, strides = [1, 1, 1, 1], padding = 'VALID') + conv2_b
# display information
if bdisplay:
print('Conv: shape', [5, 5, 18, 54], 'zeros', 54, 'mean', mu, 'stddev', sigma)
print('Conv: strides', [1, 1, 1, 1], 'padding', 'VALID')
print('Conv: inp', conv1)
print('Conv: W', conv2_W)
print('Conv: b', conv2_b)
print('Conv: out', conv2)
# Activation and drop out
conv2 = tf.nn.relu(conv2)
conv2 = tf.nn.dropout(conv2, tf_keep_prob)
print('Convolutional layer 2 :', [28, 28], 'input dimension with depth', 18, 'and', [24, 24], 'output dimensions with depth', 54)
# Pooling. Input = 24x24x54. Output = 12x12x54.
conv2 = tf.nn.max_pool(conv2, ksize = [1, 2, 2, 1], strides = [1, 2, 2, 1], padding = 'VALID')
# display information
if bdisplay:
print('Pool: filter', [1, 2, 2, 1], 'strides', [1, 2, 2, 1], 'padding', 'VALID')
print('Pool: inp', conv2)
print('Pool: out', conv2)
print('Pooling layer 2 :', [24, 24], 'input dimension with depth', 54, 'and', [12, 12], 'output dimensions with depth', 54)
# Flatten. Input = 12x12x54. Output = 7776.
fc0 = flatten(conv2)
# Layer 3: Fully Connected. Input = 7776. Output = 800.
fc1_W = tf.Variable(tf.truncated_normal(shape=(7776, 800), mean = mu, stddev = sigma))
fc1_b = tf.Variable(tf.zeros(800))
fc1 = tf.matmul(fc0, fc1_W) + fc1_b
# display information
if bdisplay:
print('Full: shape', [7776, 800], 'zeros', 800, 'mean', mu, 'stddev', sigma)
print('Full: inp', fc0)
print('Full: W', fc1_W)
print('Full: b', fc1_b)
print('Full: fc', fc1)
# Activation and drop out
fc1 = tf.nn.relu(fc1)
fc1 = tf.nn.dropout(fc1, tf_keep_prob)
print('Fully connected layer 1 :', 7776, 'input dimensions and', 800, 'output dimensions')
# Layer 4: Fully Connected. Input = 800. Output = 84.
fc2_W = tf.Variable(tf.truncated_normal(shape = (800, 84), mean = mu, stddev = sigma))
fc2_b = tf.Variable(tf.zeros(84))
fc2 = tf.matmul(fc1, fc2_W) + fc2_b
# display information
if bdisplay:
print('Full: shape', [800, 84], 'zeros', 84, 'mean', mu, 'stddev', sigma)
print('Full: inp', fc1)
print('Full: W', fc2_W)
print('Full: b', fc2_b)
print('Full: fc', fc2)
# Activation and drop out
fc2 = tf.nn.relu(fc2)
fc2 = tf.nn.dropout(fc2, tf_keep_prob)
print('Fully connected layer 2 :', 800, 'input dimensions and', 84, 'output dimensions')
# Layer 5: Fully Connected. Input = 84. Output = 43.
fc3_W = tf.Variable(tf.truncated_normal(shape = (84, 43), mean = mu, stddev = sigma))
fc3_b = tf.Variable(tf.zeros(43))
logits = tf.matmul(fc2, fc3_W) + fc3_b
# display information
if bdisplay:
print('Full: shape', [84, 43], 'zeros', 43, 'mean', mu, 'stddev', sigma)
print('Full: inp', fc2)
print('Full: W', fc3_W)
print('Full: b', fc3_b)
print('Full: fc', logits)
print('Fully connected layer 3 :', 84, 'input dimensions and', 43, 'output dimensions')
return logits
def determine_outputs(input_dims, filter_size = [5, 5], strides = [1, 1], padding = 'VALID'):
# ...
# This function calculates the output dimensions of a convolutional operation
# ...
# Inputs
# ...
# input_dims : dimensions of input (height, width)
# filter_size : size of filter [height, width]
# strides : value of strides [height_step, width_step]
# padding : type of padding operation {'VALID', 'SAME'}
# ...
# Outputs
# ...
# output_dims : dimensions of output (height, width)
# initialize output
output_dims = None
# valid padding
if padding == 'VALID':
output_dims = np.array([np.ceil(float(input_dims[0] - filter_size[0] + 1) / float(strides[0])), \
np.ceil(float(input_dims[1] - filter_size[1] + 1) / float(strides[1]))], dtype = np.int32)
# same padding
elif padding == 'SAME':
output_dims = np.array([np.ceil(float(input_dims[0]) / float(strides[0])), \
np.ceil(float(input_dims[1]) / float(strides[1]))], dtype = np.int32)
# unknown padding
else:
output_dims = None
return output_dims
```
### Train, Validate and Test the Model
A validation set can be used to assess how well the model is performing. A low accuracy on the training and validation
sets imply underfitting. A high accuracy on the training set but low accuracy on the validation set implies overfitting.
```
### Train your model here.
### Calculate and report the accuracy on the training and validation set.
### Once a final model architecture is selected,
### the accuracy on the test set should be calculated and reported as well.
### Feel free to use as many code cells as needed.
# Use DROPOUT etc. to not overfit !!!
# define constants
bdisplay = False
epochs = 50
#epochs = 10
#batch_size = 256
batch_size = 128
rate = 0.0002
#rate = 0.001
keep_prob = 0.6
#keep_prob = 1
# define tensorflow parameters
x = tf.placeholder(tf.float32, (None, 32, 32, 3), name = 'x')
y = tf.placeholder(tf.int32, (None), name = 'y')
tf_keep_prob = tf.placeholder(tf.float32, (None), name = 'tf_keep_prob')
### define pipelines
# forward propagation
one_hot_y = tf.one_hot(y, n_classes)
logits_orig = LeNet(x, bdisplay)
logits_adjusted = LeNet_adjusted(x, tf_keep_prob, bdisplay)
logits = logits_adjusted
# training (backpropagation)
cross_entropy = tf.nn.softmax_cross_entropy_with_logits(labels = one_hot_y, logits = logits)
loss_operation = tf.reduce_mean(cross_entropy)
optimizer = tf.train.AdamOptimizer(learning_rate = rate)
training_operation = optimizer.minimize(loss_operation)
# evaluation
correct_prediction = tf.equal(tf.argmax(logits, 1), tf.argmax(one_hot_y, 1))
accuracy_operation = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
# initialize saver object
saver = tf.train.Saver()
def evaluate(session, X_data, y_data, batch_size):
# ...
# This function plots a set of traffic sign images
# ...
# Inputs
# ...
# session : tensorflow session
# X_data : feature list
# y_data : label list
# batch_size : batch size
# ...
# Outputs
# ...
# relative_accuracy : relative accuracy
# create session object
session = tf.get_default_session()
# initialize output
total_accuracy = 0
# initial calculations
num_examples = len(X_data)
# for all batches do
for offset in range(0, num_examples, batch_size):
# define current batch
batch_x, batch_y = X_data[offset:(offset + batch_size)], y_data[offset:(offset + batch_size)]
# calculate accuracy
accuracy = session.run(accuracy_operation, feed_dict={x: batch_x, y: batch_y, tf_keep_prob: keep_prob})
# increment total accuracy
total_accuracy += (accuracy * len(batch_x))
# calculate output
relative_accuracy = total_accuracy / num_examples
return relative_accuracy
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
# display message
print("Training...")
# iterate over all epochs
for idx in range(epochs):
# shuffle training data in each epoch
X_train_norm, y_train_conv = shuffle(X_train_norm, y_train_conv)
# iterate over all batches
for offset in range(0, n_train, batch_size):
# get current batch training set
end = offset + batch_size
batch_x, batch_y = X_train_norm[offset:end], y_train_conv[offset:end]
# execute training operation
sess.run(training_operation, feed_dict = {x: batch_x, y: batch_y, tf_keep_prob: keep_prob})
# validate accuracy current of epoch
validation_accuracy = evaluate(sess, X_valid_norm, y_valid_conv, batch_size)
# display validation accuracy of current epoch
print("EPOCH {:4d}:".format(idx + 1), "Validation Accuracy = {:.3f}".format(validation_accuracy))
# save training result
saver.save(sess, '../../GD_GitHubData/temp_data/tsclass')
# display message
print("Model saved")
def outputFeatureMap(image_input, tf_activation, activation_min=-1, activation_max=-1 ,plt_num=1):
# Here make sure to preprocess your image_input in a way your network expects
# with size, normalization, ect if needed
# image_input =
# Note: x should be the same name as your network's tensorflow data placeholder variable
# If you get an error tf_activation is not defined it maybe having trouble accessing the variable from inside a function
activation = tf_activation.eval(session=sess,feed_dict={x : image_input})
featuremaps = activation.shape[3]
plt.figure(plt_num, figsize=(15,15))
for featuremap in range(featuremaps):
plt.subplot(6,8, featuremap+1) # sets the number of feature maps to show on each row and column
plt.title('FeatureMap ' + str(featuremap)) # displays the feature map number
if activation_min != -1 & activation_max != -1:
plt.imshow(activation[0,:,:, featuremap], interpolation="nearest", vmin =activation_min, vmax=activation_max, cmap="gray")
elif activation_max != -1:
plt.imshow(activation[0,:,:, featuremap], interpolation="nearest", vmax=activation_max, cmap="gray")
elif activation_min !=-1:
plt.imshow(activation[0,:,:, featuremap], interpolation="nearest", vmin=activation_min, cmap="gray")
else:
plt.imshow(activation[0,:,:, featuremap], interpolation="nearest", cmap="gray")
with tf.Session() as sess:
saver.restore(sess, tf.train.latest_checkpoint('../../GD_GitHubData/temp_data'))
test_accuracy = evaluate(sess, X_test_norm, y_test_conv, batch_size)
print("Test Accuracy = {:.3f}".format(test_accuracy))
```
---
## Step 3: Test a Model on New Images
To give yourself more insight into how your model is working, download at least five pictures of German traffic signs from the web and use your model to predict the traffic sign type.
You may find `signnames.csv` useful as it contains mappings from the class id (integer) to the actual sign name.
### Load and Output the Images
```
### Load the images and plot them here.
### Feel free to use as many code cells as needed.
```
### Predict the Sign Type for Each Image
```
### Run the predictions here and use the model to output the prediction for each image.
### Make sure to pre-process the images with the same pre-processing pipeline used earlier.
### Feel free to use as many code cells as needed.
```
### Analyze Performance
```
### Calculate the accuracy for these 5 new images.
### For example, if the model predicted 1 out of 5 signs correctly, it's 20% accurate on these new images.
```
### Output Top 5 Softmax Probabilities For Each Image Found on the Web
For each of the new images, print out the model's softmax probabilities to show the **certainty** of the model's predictions (limit the output to the top 5 probabilities for each image). [`tf.nn.top_k`](https://www.tensorflow.org/versions/r0.12/api_docs/python/nn.html#top_k) could prove helpful here.
The example below demonstrates how tf.nn.top_k can be used to find the top k predictions for each image.
`tf.nn.top_k` will return the values and indices (class ids) of the top k predictions. So if k=3, for each sign, it'll return the 3 largest probabilities (out of a possible 43) and the correspoding class ids.
Take this numpy array as an example. The values in the array represent predictions. The array contains softmax probabilities for five candidate images with six possible classes. `tf.nn.top_k` is used to choose the three classes with the highest probability:
```
# (5, 6) array
a = np.array([[ 0.24879643, 0.07032244, 0.12641572, 0.34763842, 0.07893497,
0.12789202],
[ 0.28086119, 0.27569815, 0.08594638, 0.0178669 , 0.18063401,
0.15899337],
[ 0.26076848, 0.23664738, 0.08020603, 0.07001922, 0.1134371 ,
0.23892179],
[ 0.11943333, 0.29198961, 0.02605103, 0.26234032, 0.1351348 ,
0.16505091],
[ 0.09561176, 0.34396535, 0.0643941 , 0.16240774, 0.24206137,
0.09155967]])
```
Running it through `sess.run(tf.nn.top_k(tf.constant(a), k=3))` produces:
```
TopKV2(values=array([[ 0.34763842, 0.24879643, 0.12789202],
[ 0.28086119, 0.27569815, 0.18063401],
[ 0.26076848, 0.23892179, 0.23664738],
[ 0.29198961, 0.26234032, 0.16505091],
[ 0.34396535, 0.24206137, 0.16240774]]), indices=array([[3, 0, 5],
[0, 1, 4],
[0, 5, 1],
[1, 3, 5],
[1, 4, 3]], dtype=int32))
```
Looking just at the first row we get `[ 0.34763842, 0.24879643, 0.12789202]`, you can confirm these are the 3 largest probabilities in `a`. You'll also notice `[3, 0, 5]` are the corresponding indices.
```
### Print out the top five softmax probabilities for the predictions on the German traffic sign images found on the web.
### Feel free to use as many code cells as needed.
```
### Project Writeup
Once you have completed the code implementation, document your results in a project writeup using this [template](https://github.com/udacity/CarND-Traffic-Sign-Classifier-Project/blob/master/writeup_template.md) as a guide. The writeup can be in a markdown or pdf file.
> **Note**: Once you have completed all of the code implementations and successfully answered each question above, you may finalize your work by exporting the iPython Notebook as an HTML document. You can do this by using the menu above and navigating to \n",
"**File -> Download as -> HTML (.html)**. Include the finished document along with this notebook as your submission.
---
## Step 4 (Optional): Visualize the Neural Network's State with Test Images
This Section is not required to complete but acts as an additional excersise for understaning the output of a neural network's weights. While neural networks can be a great learning device they are often referred to as a black box. We can understand what the weights of a neural network look like better by plotting their feature maps. After successfully training your neural network you can see what it's feature maps look like by plotting the output of the network's weight layers in response to a test stimuli image. From these plotted feature maps, it's possible to see what characteristics of an image the network finds interesting. For a sign, maybe the inner network feature maps react with high activation to the sign's boundary outline or to the contrast in the sign's painted symbol.
Provided for you below is the function code that allows you to get the visualization output of any tensorflow weight layer you want. The inputs to the function should be a stimuli image, one used during training or a new one you provided, and then the tensorflow variable name that represents the layer's state during the training process, for instance if you wanted to see what the [LeNet lab's](https://classroom.udacity.com/nanodegrees/nd013/parts/fbf77062-5703-404e-b60c-95b78b2f3f9e/modules/6df7ae49-c61c-4bb2-a23e-6527e69209ec/lessons/601ae704-1035-4287-8b11-e2c2716217ad/concepts/d4aca031-508f-4e0b-b493-e7b706120f81) feature maps looked like for it's second convolutional layer you could enter conv2 as the tf_activation variable.
For an example of what feature map outputs look like, check out NVIDIA's results in their paper [End-to-End Deep Learning for Self-Driving Cars](https://devblogs.nvidia.com/parallelforall/deep-learning-self-driving-cars/) in the section Visualization of internal CNN State. NVIDIA was able to show that their network's inner weights had high activations to road boundary lines by comparing feature maps from an image with a clear path to one without. Try experimenting with a similar test to show that your trained network's weights are looking for interesting features, whether it's looking at differences in feature maps from images with or without a sign, or even what feature maps look like in a trained network vs a completely untrained one on the same sign image.
<figure>
<img src="visualize_cnn.png" width="380" alt="Combined Image" />
<figcaption>
<p></p>
<p style="text-align: center;"> Your output should look something like this (above)</p>
</figcaption>
</figure>
<p></p>
```
### Visualize your network's feature maps here.
### Feel free to use as many code cells as needed.
# image_input: the test image being fed into the network to produce the feature maps
# tf_activation: should be a tf variable name used during your training procedure that represents the calculated state of a specific weight layer
# activation_min/max: can be used to view the activation contrast in more detail, by default matplot sets min and max to the actual min and max values of the output
# plt_num: used to plot out multiple different weight feature map sets on the same block, just extend the plt number for each new feature map entry
def outputFeatureMap(image_input, tf_activation, activation_min=-1, activation_max=-1 ,plt_num=1):
# Here make sure to preprocess your image_input in a way your network expects
# with size, normalization, ect if needed
# image_input =
# Note: x should be the same name as your network's tensorflow data placeholder variable
# If you get an error tf_activation is not defined it may be having trouble accessing the variable from inside a function
activation = tf_activation.eval(session=sess,feed_dict={x : image_input})
featuremaps = activation.shape[3]
plt.figure(plt_num, figsize=(15,15))
for featuremap in range(featuremaps):
plt.subplot(6,8, featuremap+1) # sets the number of feature maps to show on each row and column
plt.title('FeatureMap ' + str(featuremap)) # displays the feature map number
if activation_min != -1 & activation_max != -1:
plt.imshow(activation[0,:,:, featuremap], interpolation="nearest", vmin =activation_min, vmax=activation_max, cmap="gray")
elif activation_max != -1:
plt.imshow(activation[0,:,:, featuremap], interpolation="nearest", vmax=activation_max, cmap="gray")
elif activation_min !=-1:
plt.imshow(activation[0,:,:, featuremap], interpolation="nearest", vmin=activation_min, cmap="gray")
else:
plt.imshow(activation[0,:,:, featuremap], interpolation="nearest", cmap="gray")
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import os
import torch
from PIL import Image
from torch.utils.data import Dataset, DataLoader
from torchvision import io, datasets, transforms
from tqdm.notebook import tqdm
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
#enable cuda
use_cuda = torch.cuda.is_available()
if use_cuda:
device = torch.device('cuda')
loader_kwargs = {'num_workers': 1, 'pin_memory': True}
else:
device = torch.device('cpu')
loader_kwargs = {}
print(device)
class ExpressionDataset(Dataset):
"""Binomial expression dataset"""
def __init__(self, csv_file, root_dir):
self.labels = pd.read_csv(csv_file)
self.root_dir = root_dir
def __len__(self):
return len(self.labels)
def __getitem__(self, idx):
if torch.is_tensor(idx):
idx = idx.tolist()
img_name = os.path.join(self.root_dir, self.labels.iloc[idx,0])
img = io.read_image(img_name).float()
exprtype, ans = self.labels.iloc[idx,1:]
exprtype_int = 0
if exprtype == "infix":
exprtype_int = 1
elif exprtype == "postfix":
exprtype_int = 2
return [img, exprtype_int, ans+9]
dataset = ExpressionDataset(csv_file='../input/soml-hackathon/SoML/SoML-50/annotations.csv', root_dir='../input/soml-hackathon/SoML/SoML-50/data')
plt.imshow(dataset[10][0].squeeze())
train_set, test_set, validation_set = torch.utils.data.random_split(dataset, [40000,5000,5000])
train_loader = DataLoader(train_set, batch_size=50, shuffle=True)
test_loader = DataLoader(test_set, batch_size=50, shuffle=True)
validation_loader = DataLoader(validation_set, batch_size=50, shuffle=True)
class ExpressionTypeNetwork(torch.nn.Module):
def __init__(self):
super().__init__()
self.conv1 = nn.Conv2d(1, 10, kernel_size = 8)
self.conv2 = nn.Conv2d(10, 20, kernel_size = 4)
self.conv2_drop = nn.Dropout2d()
self.fc1 = nn.Linear(1900, 100)
self.fc2 = nn.Linear(100, 3)
# self.input_layer = torch.nn.Linear(128*384, 192)
#self.hl1 = torch.nn.Linear(384, 14*3)
# self.hl2 = torch.nn.Linear(192, 3)
# self.relu = torch.nn.ReLU()
def forward(self, x):
x = F.relu(F.max_pool2d(self.conv1(x), 6))
x = F.relu(F.max_pool2d(self.conv2_drop(self.conv2(x)), 3))
# print(x.shape)
x = torch.flatten(x,1)
x = F.relu(self.fc1(x))
x = F.dropout(x, training = self.training)
x = self.fc2(x)
return x
# x = self.input_layer(x)
# x = self.relu(x)
# x = self.hl2(x)
# x = self.relu(x)
# x = self.hl2(x)
# return x
def acc(self, loader):
total = 0
correct = 0
with torch.no_grad():
for images, types, answers in loader:
images, types = images.to(device), types.to(device)
batch_size = images.shape[0]
# images = images.reshape(batch_size, (128//4)*(384//4))
output = self(images)
predicted = torch.argmax(output, dim=1)
correct += torch.sum(types == predicted)
total += batch_size
return correct/total
type_net = ExpressionTypeNetwork().to(device)
lossfn = torch.nn.CrossEntropyLoss()
optimizer = torch.optim.Adam(type_net.parameters(), lr=0.001)
type_net.eval()
print(f"Accuracy: {type_net.acc(validation_loader)}")
for epoch in range(10):
avg_loss = 0
num_iters = 0
type_net.train()
for images, types, answers in tqdm(train_loader):
images, types = images.to(device), types.to(device)
optimizer.zero_grad()
batch_size = images.shape[0]
# images = images.reshape(batch_size, 128*384)
output = type_net(images)
loss = lossfn(output, types)
loss.backward()
optimizer.step()
avg_loss += loss.item()
num_iters += 1
type_net.eval()
print(f"Loss: {avg_loss/num_iters}")
print(f"Accuracy: {type_net.acc(validation_loader)}")
torch.save(type_net.state_dict(), '/kaggle/working/type_net_dict')
```
| github_jupyter |
```
# Copyright 2020 Google LLC
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Vertex client library: AutoML tabular classification model for batch prediction with explanation
<table align="left">
<td>
<a href="https://colab.research.google.com/github/GoogleCloudPlatform/vertex-ai-samples/blob/master/notebooks/community/gapic/automl/showcase_automl_tabular_classification_batch_explain.ipynb">
<img src="https://cloud.google.com/ml-engine/images/colab-logo-32px.png" alt="Colab logo"> Run in Colab
</a>
</td>
<td>
<a href="https://github.com/GoogleCloudPlatform/vertex-ai-samples/blob/master/notebooks/community/gapic/automl/showcase_automl_tabular_classification_batch_explain.ipynb">
<img src="https://cloud.google.com/ml-engine/images/github-logo-32px.png" alt="GitHub logo">
View on GitHub
</a>
</td>
</table>
<br/><br/><br/>
## Overview
This tutorial demonstrates how to use the Vertex client library for Python to create tabular classification models and do batch prediction with explanation using Google Cloud's [AutoML](https://cloud.google.com/vertex-ai/docs/start/automl-users).
### Dataset
The dataset used for this tutorial is the [Iris dataset](https://www.tensorflow.org/datasets/catalog/iris) from [TensorFlow Datasets](https://www.tensorflow.org/datasets/catalog/overview). This dataset does not require any feature engineering. The version of the dataset you will use in this tutorial is stored in a public Cloud Storage bucket. The trained model predicts the type of Iris flower species from a class of three species: setosa, virginica, or versicolor.
### Objective
In this tutorial, you create an AutoML tabular classification model from a Python script, and then do a batch prediction with explainability using the Vertex client library. You can alternatively create and deploy models using the `gcloud` command-line tool or online using the Google Cloud Console.
The steps performed include:
- Create a Vertex `Dataset` resource.
- Train the model.
- View the model evaluation.
- Make a batch prediction with explainability.
There is one key difference between using batch prediction and using online prediction:
* Prediction Service: Does an on-demand prediction for the entire set of instances (i.e., one or more data items) and returns the results in real-time.
* Batch Prediction Service: Does a queued (batch) prediction for the entire set of instances in the background and stores the results in a Cloud Storage bucket when ready.
### Costs
This tutorial uses billable components of Google Cloud (GCP):
* Vertex AI
* Cloud Storage
Learn about [Vertex AI
pricing](https://cloud.google.com/vertex-ai/pricing) and [Cloud Storage
pricing](https://cloud.google.com/storage/pricing), and use the [Pricing
Calculator](https://cloud.google.com/products/calculator/)
to generate a cost estimate based on your projected usage.
## Installation
Install the latest version of Vertex client library.
```
import os
import sys
# Google Cloud Notebook
if os.path.exists("/opt/deeplearning/metadata/env_version"):
USER_FLAG = "--user"
else:
USER_FLAG = ""
! pip3 install -U google-cloud-aiplatform $USER_FLAG
```
Install the latest GA version of *google-cloud-storage* library as well.
```
! pip3 install -U google-cloud-storage $USER_FLAG
```
### Restart the kernel
Once you've installed the Vertex client library and Google *cloud-storage*, you need to restart the notebook kernel so it can find the packages.
```
if not os.getenv("IS_TESTING"):
# Automatically restart kernel after installs
import IPython
app = IPython.Application.instance()
app.kernel.do_shutdown(True)
```
## Before you begin
### GPU runtime
*Make sure you're running this notebook in a GPU runtime if you have that option. In Colab, select* **Runtime > Change Runtime Type > GPU**
### Set up your Google Cloud project
**The following steps are required, regardless of your notebook environment.**
1. [Select or create a Google Cloud project](https://console.cloud.google.com/cloud-resource-manager). When you first create an account, you get a $300 free credit towards your compute/storage costs.
2. [Make sure that billing is enabled for your project.](https://cloud.google.com/billing/docs/how-to/modify-project)
3. [Enable the Vertex APIs and Compute Engine APIs.](https://console.cloud.google.com/flows/enableapi?apiid=ml.googleapis.com,compute_component)
4. [The Google Cloud SDK](https://cloud.google.com/sdk) is already installed in Google Cloud Notebook.
5. Enter your project ID in the cell below. Then run the cell to make sure the
Cloud SDK uses the right project for all the commands in this notebook.
**Note**: Jupyter runs lines prefixed with `!` as shell commands, and it interpolates Python variables prefixed with `$` into these commands.
```
PROJECT_ID = "[your-project-id]" # @param {type:"string"}
if PROJECT_ID == "" or PROJECT_ID is None or PROJECT_ID == "[your-project-id]":
# Get your GCP project id from gcloud
shell_output = !gcloud config list --format 'value(core.project)' 2>/dev/null
PROJECT_ID = shell_output[0]
print("Project ID:", PROJECT_ID)
! gcloud config set project $PROJECT_ID
```
#### Region
You can also change the `REGION` variable, which is used for operations
throughout the rest of this notebook. Below are regions supported for Vertex. We recommend that you choose the region closest to you.
- Americas: `us-central1`
- Europe: `europe-west4`
- Asia Pacific: `asia-east1`
You may not use a multi-regional bucket for training with Vertex. Not all regions provide support for all Vertex services. For the latest support per region, see the [Vertex locations documentation](https://cloud.google.com/vertex-ai/docs/general/locations)
```
REGION = "us-central1" # @param {type: "string"}
```
#### Timestamp
If you are in a live tutorial session, you might be using a shared test account or project. To avoid name collisions between users on resources created, you create a timestamp for each instance session, and append onto the name of resources which will be created in this tutorial.
```
from datetime import datetime
TIMESTAMP = datetime.now().strftime("%Y%m%d%H%M%S")
```
### Authenticate your Google Cloud account
**If you are using Google Cloud Notebook**, your environment is already authenticated. Skip this step.
**If you are using Colab**, run the cell below and follow the instructions when prompted to authenticate your account via oAuth.
**Otherwise**, follow these steps:
In the Cloud Console, go to the [Create service account key](https://console.cloud.google.com/apis/credentials/serviceaccountkey) page.
**Click Create service account**.
In the **Service account name** field, enter a name, and click **Create**.
In the **Grant this service account access to project** section, click the Role drop-down list. Type "Vertex" into the filter box, and select **Vertex Administrator**. Type "Storage Object Admin" into the filter box, and select **Storage Object Admin**.
Click Create. A JSON file that contains your key downloads to your local environment.
Enter the path to your service account key as the GOOGLE_APPLICATION_CREDENTIALS variable in the cell below and run the cell.
```
# If you are running this notebook in Colab, run this cell and follow the
# instructions to authenticate your GCP account. This provides access to your
# Cloud Storage bucket and lets you submit training jobs and prediction
# requests.
# If on Google Cloud Notebook, then don't execute this code
if not os.path.exists("/opt/deeplearning/metadata/env_version"):
if "google.colab" in sys.modules:
from google.colab import auth as google_auth
google_auth.authenticate_user()
# If you are running this notebook locally, replace the string below with the
# path to your service account key and run this cell to authenticate your GCP
# account.
elif not os.getenv("IS_TESTING"):
%env GOOGLE_APPLICATION_CREDENTIALS ''
```
### Create a Cloud Storage bucket
**The following steps are required, regardless of your notebook environment.**
This tutorial is designed to use training data that is in a public Cloud Storage bucket and a local Cloud Storage bucket for your batch predictions. You may alternatively use your own training data that you have stored in a local Cloud Storage bucket.
Set the name of your Cloud Storage bucket below. Bucket names must be globally unique across all Google Cloud projects, including those outside of your organization.
```
BUCKET_NAME = "gs://[your-bucket-name]" # @param {type:"string"}
if BUCKET_NAME == "" or BUCKET_NAME is None or BUCKET_NAME == "gs://[your-bucket-name]":
BUCKET_NAME = "gs://" + PROJECT_ID + "aip-" + TIMESTAMP
```
**Only if your bucket doesn't already exist**: Run the following cell to create your Cloud Storage bucket.
```
! gsutil mb -l $REGION $BUCKET_NAME
```
Finally, validate access to your Cloud Storage bucket by examining its contents:
```
! gsutil ls -al $BUCKET_NAME
```
### Set up variables
Next, set up some variables used throughout the tutorial.
### Import libraries and define constants
#### Import Vertex client library
Import the Vertex client library into our Python environment.
```
import time
import google.cloud.aiplatform_v1beta1 as aip
from google.protobuf import json_format
from google.protobuf.json_format import MessageToJson, ParseDict
from google.protobuf.struct_pb2 import Struct, Value
```
#### Vertex constants
Setup up the following constants for Vertex:
- `API_ENDPOINT`: The Vertex API service endpoint for dataset, model, job, pipeline and endpoint services.
- `PARENT`: The Vertex location root path for dataset, model, job, pipeline and endpoint resources.
```
# API service endpoint
API_ENDPOINT = "{}-aiplatform.googleapis.com".format(REGION)
# Vertex location root path for your dataset, model and endpoint resources
PARENT = "projects/" + PROJECT_ID + "/locations/" + REGION
```
#### AutoML constants
Set constants unique to AutoML datasets and training:
- Dataset Schemas: Tells the `Dataset` resource service which type of dataset it is.
- Data Labeling (Annotations) Schemas: Tells the `Dataset` resource service how the data is labeled (annotated).
- Dataset Training Schemas: Tells the `Pipeline` resource service the task (e.g., classification) to train the model for.
```
# Tabular Dataset type
DATA_SCHEMA = "gs://google-cloud-aiplatform/schema/dataset/metadata/tables_1.0.0.yaml"
# Tabular Labeling type
LABEL_SCHEMA = (
"gs://google-cloud-aiplatform/schema/dataset/ioformat/table_io_format_1.0.0.yaml"
)
# Tabular Training task
TRAINING_SCHEMA = "gs://google-cloud-aiplatform/schema/trainingjob/definition/automl_tables_1.0.0.yaml"
```
#### Hardware Accelerators
Set the hardware accelerators (e.g., GPU), if any, for prediction.
Set the variable `DEPLOY_GPU/DEPLOY_NGPU` to use a container image supporting a GPU and the number of GPUs allocated to the virtual machine (VM) instance. For example, to use a GPU container image with 4 Nvidia Telsa K80 GPUs allocated to each VM, you would specify:
(aip.AcceleratorType.NVIDIA_TESLA_K80, 4)
For GPU, available accelerators include:
- aip.AcceleratorType.NVIDIA_TESLA_K80
- aip.AcceleratorType.NVIDIA_TESLA_P100
- aip.AcceleratorType.NVIDIA_TESLA_P4
- aip.AcceleratorType.NVIDIA_TESLA_T4
- aip.AcceleratorType.NVIDIA_TESLA_V100
Otherwise specify `(None, None)` to use a container image to run on a CPU.
```
if os.getenv("IS_TESTING_DEPOLY_GPU"):
DEPLOY_GPU, DEPLOY_NGPU = (
aip.AcceleratorType.NVIDIA_TESLA_K80,
int(os.getenv("IS_TESTING_DEPOLY_GPU")),
)
else:
DEPLOY_GPU, DEPLOY_NGPU = (aip.AcceleratorType.NVIDIA_TESLA_K80, 1)
```
#### Container (Docker) image
For AutoML batch prediction, the container image for the serving binary is pre-determined by the Vertex prediction service. More specifically, the service will pick the appropriate container for the model depending on the hardware accelerator you selected.
#### Machine Type
Next, set the machine type to use for prediction.
- Set the variable `DEPLOY_COMPUTE` to configure the compute resources for the VM you will use for prediction.
- `machine type`
- `n1-standard`: 3.75GB of memory per vCPU.
- `n1-highmem`: 6.5GB of memory per vCPU
- `n1-highcpu`: 0.9 GB of memory per vCPU
- `vCPUs`: number of \[2, 4, 8, 16, 32, 64, 96 \]
*Note: You may also use n2 and e2 machine types for training and deployment, but they do not support GPUs*
```
if os.getenv("IS_TESTING_DEPLOY_MACHINE"):
MACHINE_TYPE = os.getenv("IS_TESTING_DEPLOY_MACHINE")
else:
MACHINE_TYPE = "n1-standard"
VCPU = "4"
DEPLOY_COMPUTE = MACHINE_TYPE + "-" + VCPU
print("Deploy machine type", DEPLOY_COMPUTE)
```
# Tutorial
Now you are ready to start creating your own AutoML tabular classification model.
## Set up clients
The Vertex client library works as a client/server model. On your side (the Python script) you will create a client that sends requests and receives responses from the Vertex server.
You will use different clients in this tutorial for different steps in the workflow. So set them all up upfront.
- Dataset Service for `Dataset` resources.
- Model Service for `Model` resources.
- Pipeline Service for training.
- Job Service for batch prediction and custom training.
```
# client options same for all services
client_options = {"api_endpoint": API_ENDPOINT}
def create_dataset_client():
client = aip.DatasetServiceClient(client_options=client_options)
return client
def create_model_client():
client = aip.ModelServiceClient(client_options=client_options)
return client
def create_pipeline_client():
client = aip.PipelineServiceClient(client_options=client_options)
return client
def create_job_client():
client = aip.JobServiceClient(client_options=client_options)
return client
clients = {}
clients["dataset"] = create_dataset_client()
clients["model"] = create_model_client()
clients["pipeline"] = create_pipeline_client()
clients["job"] = create_job_client()
for client in clients.items():
print(client)
```
## Dataset
Now that your clients are ready, your first step is to create a `Dataset` resource instance. This step differs from Vision, Video and Language. For those products, after the `Dataset` resource is created, one then separately imports the data, using the `import_data` method.
For tabular, importing of the data is deferred until the training pipeline starts training the model. What do we do different? Well, first you won't be calling the `import_data` method. Instead, when you create the dataset instance you specify the Cloud Storage location of the CSV file or BigQuery location of the data table, which contains your tabular data as part of the `Dataset` resource's metadata.
#### Cloud Storage
`metadata = {"input_config": {"gcs_source": {"uri": [gcs_uri]}}}`
The format for a Cloud Storage path is:
gs://[bucket_name]/[folder(s)/[file]
#### BigQuery
`metadata = {"input_config": {"bigquery_source": {"uri": [gcs_uri]}}}`
The format for a BigQuery path is:
bq://[collection].[dataset].[table]
Note that the `uri` field is a list, whereby you can input multiple CSV files or BigQuery tables when your data is split across files.
### Data preparation
The Vertex `Dataset` resource for tabular has a couple of requirements for your tabular data.
- Must be in a CSV file or a BigQuery query.
#### CSV
For tabular classification, the CSV file has a few requirements:
- The first row must be the heading -- note how this is different from Vision, Video and Language where the requirement is no heading.
- All but one column are features.
- One column is the label, which you will specify when you subsequently create the training pipeline.
#### Location of Cloud Storage training data.
Now set the variable `IMPORT_FILE` to the location of the CSV index file in Cloud Storage.
```
IMPORT_FILE = "gs://cloud-samples-data/tables/iris_1000.csv"
```
#### Quick peek at your data
You will use a version of the Iris dataset that is stored in a public Cloud Storage bucket, using a CSV index file.
Start by doing a quick peek at the data. You count the number of examples by counting the number of rows in the CSV index file (`wc -l`) and then peek at the first few rows.
You also need for training to know the heading name of the label column, which is save as `label_column`. For this dataset, it is the last column in the CSV file.
```
count = ! gsutil cat $IMPORT_FILE | wc -l
print("Number of Examples", int(count[0]))
print("First 10 rows")
! gsutil cat $IMPORT_FILE | head
heading = ! gsutil cat $IMPORT_FILE | head -n1
label_column = str(heading).split(",")[-1].split("'")[0]
print("Label Column Name", label_column)
if label_column is None:
raise Exception("label column missing")
```
## Dataset
Now that your clients are ready, your first step in training a model is to create a managed dataset instance, and then upload your labeled data to it.
### Create `Dataset` resource instance
Use the helper function `create_dataset` to create the instance of a `Dataset` resource. This function does the following:
1. Uses the dataset client service.
2. Creates an Vertex `Dataset` resource (`aip.Dataset`), with the following parameters:
- `display_name`: The human-readable name you choose to give it.
- `metadata_schema_uri`: The schema for the dataset type.
- `metadata`: The Cloud Storage or BigQuery location of the tabular data.
3. Calls the client dataset service method `create_dataset`, with the following parameters:
- `parent`: The Vertex location root path for your `Database`, `Model` and `Endpoint` resources.
- `dataset`: The Vertex dataset object instance you created.
4. The method returns an `operation` object.
An `operation` object is how Vertex handles asynchronous calls for long running operations. While this step usually goes fast, when you first use it in your project, there is a longer delay due to provisioning.
You can use the `operation` object to get status on the operation (e.g., create `Dataset` resource) or to cancel the operation, by invoking an operation method:
| Method | Description |
| ----------- | ----------- |
| result() | Waits for the operation to complete and returns a result object in JSON format. |
| running() | Returns True/False on whether the operation is still running. |
| done() | Returns True/False on whether the operation is completed. |
| canceled() | Returns True/False on whether the operation was canceled. |
| cancel() | Cancels the operation (this may take up to 30 seconds). |
```
TIMEOUT = 90
def create_dataset(name, schema, src_uri=None, labels=None, timeout=TIMEOUT):
start_time = time.time()
try:
if src_uri.startswith("gs://"):
metadata = {"input_config": {"gcs_source": {"uri": [src_uri]}}}
elif src_uri.startswith("bq://"):
metadata = {"input_config": {"bigquery_source": {"uri": [src_uri]}}}
dataset = aip.Dataset(
display_name=name,
metadata_schema_uri=schema,
labels=labels,
metadata=json_format.ParseDict(metadata, Value()),
)
operation = clients["dataset"].create_dataset(parent=PARENT, dataset=dataset)
print("Long running operation:", operation.operation.name)
result = operation.result(timeout=TIMEOUT)
print("time:", time.time() - start_time)
print("response")
print(" name:", result.name)
print(" display_name:", result.display_name)
print(" metadata_schema_uri:", result.metadata_schema_uri)
print(" metadata:", dict(result.metadata))
print(" create_time:", result.create_time)
print(" update_time:", result.update_time)
print(" etag:", result.etag)
print(" labels:", dict(result.labels))
return result
except Exception as e:
print("exception:", e)
return None
result = create_dataset("iris-" + TIMESTAMP, DATA_SCHEMA, src_uri=IMPORT_FILE)
```
Now save the unique dataset identifier for the `Dataset` resource instance you created.
```
# The full unique ID for the dataset
dataset_id = result.name
# The short numeric ID for the dataset
dataset_short_id = dataset_id.split("/")[-1]
print(dataset_id)
```
## Train the model
Now train an AutoML tabular classification model using your Vertex `Dataset` resource. To train the model, do the following steps:
1. Create an Vertex training pipeline for the `Dataset` resource.
2. Execute the pipeline to start the training.
### Create a training pipeline
You may ask, what do we use a pipeline for? You typically use pipelines when the job (such as training) has multiple steps, generally in sequential order: do step A, do step B, etc. By putting the steps into a pipeline, we gain the benefits of:
1. Being reusable for subsequent training jobs.
2. Can be containerized and ran as a batch job.
3. Can be distributed.
4. All the steps are associated with the same pipeline job for tracking progress.
Use this helper function `create_pipeline`, which takes the following parameters:
- `pipeline_name`: A human readable name for the pipeline job.
- `model_name`: A human readable name for the model.
- `dataset`: The Vertex fully qualified dataset identifier.
- `schema`: The dataset labeling (annotation) training schema.
- `task`: A dictionary describing the requirements for the training job.
The helper function calls the `Pipeline` client service'smethod `create_pipeline`, which takes the following parameters:
- `parent`: The Vertex location root path for your `Dataset`, `Model` and `Endpoint` resources.
- `training_pipeline`: the full specification for the pipeline training job.
Let's look now deeper into the *minimal* requirements for constructing a `training_pipeline` specification:
- `display_name`: A human readable name for the pipeline job.
- `training_task_definition`: The dataset labeling (annotation) training schema.
- `training_task_inputs`: A dictionary describing the requirements for the training job.
- `model_to_upload`: A human readable name for the model.
- `input_data_config`: The dataset specification.
- `dataset_id`: The Vertex dataset identifier only (non-fully qualified) -- this is the last part of the fully-qualified identifier.
- `fraction_split`: If specified, the percentages of the dataset to use for training, test and validation. Otherwise, the percentages are automatically selected by AutoML.
```
def create_pipeline(pipeline_name, model_name, dataset, schema, task):
dataset_id = dataset.split("/")[-1]
input_config = {
"dataset_id": dataset_id,
"fraction_split": {
"training_fraction": 0.8,
"validation_fraction": 0.1,
"test_fraction": 0.1,
},
}
training_pipeline = {
"display_name": pipeline_name,
"training_task_definition": schema,
"training_task_inputs": task,
"input_data_config": input_config,
"model_to_upload": {"display_name": model_name},
}
try:
pipeline = clients["pipeline"].create_training_pipeline(
parent=PARENT, training_pipeline=training_pipeline
)
print(pipeline)
except Exception as e:
print("exception:", e)
return None
return pipeline
```
### Construct the task requirements
Next, construct the task requirements. Unlike other parameters which take a Python (JSON-like) dictionary, the `task` field takes a Google protobuf Struct, which is very similar to a Python dictionary. Use the `json_format.ParseDict` method for the conversion.
The minimal fields you need to specify are:
- `prediction_type`: Whether we are doing "classification" or "regression".
- `target_column`: The CSV heading column name for the column we want to predict (i.e., the label).
- `train_budget_milli_node_hours`: The maximum time to budget (billed) for training the model, where 1000 = 1 hour.
- `disable_early_stopping`: Whether True/False to let AutoML use its judgement to stop training early or train for the entire budget.
- `transformations`: Specifies the feature engineering for each feature column.
For `transformations`, the list must have an entry for each column. The outer key field indicates the type of feature engineering for the corresponding column. In this tutorial, you set it to `"auto"` to tell AutoML to automatically determine it.
Finally, create the pipeline by calling the helper function `create_pipeline`, which returns an instance of a training pipeline object.
```
TRANSFORMATIONS = [
{"auto": {"column_name": "sepal_width"}},
{"auto": {"column_name": "sepal_length"}},
{"auto": {"column_name": "petal_length"}},
{"auto": {"column_name": "petal_width"}},
]
PIPE_NAME = "iris_pipe-" + TIMESTAMP
MODEL_NAME = "iris_model-" + TIMESTAMP
task = Value(
struct_value=Struct(
fields={
"target_column": Value(string_value=label_column),
"prediction_type": Value(string_value="classification"),
"train_budget_milli_node_hours": Value(number_value=1000),
"disable_early_stopping": Value(bool_value=False),
"transformations": json_format.ParseDict(TRANSFORMATIONS, Value()),
}
)
)
response = create_pipeline(PIPE_NAME, MODEL_NAME, dataset_id, TRAINING_SCHEMA, task)
```
Now save the unique identifier of the training pipeline you created.
```
# The full unique ID for the pipeline
pipeline_id = response.name
# The short numeric ID for the pipeline
pipeline_short_id = pipeline_id.split("/")[-1]
print(pipeline_id)
```
### Get information on a training pipeline
Now get pipeline information for just this training pipeline instance. The helper function gets the job information for just this job by calling the the job client service's `get_training_pipeline` method, with the following parameter:
- `name`: The Vertex fully qualified pipeline identifier.
When the model is done training, the pipeline state will be `PIPELINE_STATE_SUCCEEDED`.
```
def get_training_pipeline(name, silent=False):
response = clients["pipeline"].get_training_pipeline(name=name)
if silent:
return response
print("pipeline")
print(" name:", response.name)
print(" display_name:", response.display_name)
print(" state:", response.state)
print(" training_task_definition:", response.training_task_definition)
print(" training_task_inputs:", dict(response.training_task_inputs))
print(" create_time:", response.create_time)
print(" start_time:", response.start_time)
print(" end_time:", response.end_time)
print(" update_time:", response.update_time)
print(" labels:", dict(response.labels))
return response
response = get_training_pipeline(pipeline_id)
```
# Deployment
Training the above model may take upwards of 30 minutes time.
Once your model is done training, you can calculate the actual time it took to train the model by subtracting `end_time` from `start_time`. For your model, you will need to know the fully qualified Vertex Model resource identifier, which the pipeline service assigned to it. You can get this from the returned pipeline instance as the field `model_to_deploy.name`.
```
while True:
response = get_training_pipeline(pipeline_id, True)
if response.state != aip.PipelineState.PIPELINE_STATE_SUCCEEDED:
print("Training job has not completed:", response.state)
model_to_deploy_id = None
if response.state == aip.PipelineState.PIPELINE_STATE_FAILED:
raise Exception("Training Job Failed")
else:
model_to_deploy = response.model_to_upload
model_to_deploy_id = model_to_deploy.name
print("Training Time:", response.end_time - response.start_time)
break
time.sleep(60)
print("model to deploy:", model_to_deploy_id)
```
## Model information
Now that your model is trained, you can get some information on your model.
## Evaluate the Model resource
Now find out how good the model service believes your model is. As part of training, some portion of the dataset was set aside as the test (holdout) data, which is used by the pipeline service to evaluate the model.
### List evaluations for all slices
Use this helper function `list_model_evaluations`, which takes the following parameter:
- `name`: The Vertex fully qualified model identifier for the `Model` resource.
This helper function uses the model client service's `list_model_evaluations` method, which takes the same parameter. The response object from the call is a list, where each element is an evaluation metric.
For each evaluation (you probably only have one) we then print all the key names for each metric in the evaluation, and for a small set (`logLoss` and `auPrc`) you will print the result.
```
def list_model_evaluations(name):
response = clients["model"].list_model_evaluations(parent=name)
for evaluation in response:
print("model_evaluation")
print(" name:", evaluation.name)
print(" metrics_schema_uri:", evaluation.metrics_schema_uri)
metrics = json_format.MessageToDict(evaluation._pb.metrics)
for metric in metrics.keys():
print(metric)
print("logloss", metrics["logLoss"])
print("auPrc", metrics["auPrc"])
return evaluation.name
last_evaluation = list_model_evaluations(model_to_deploy_id)
```
## Model deployment for batch prediction
Now deploy the trained Vertex `Model` resource you created for batch prediction. This differs from deploying a `Model` resource for on-demand prediction.
For online prediction, you:
1. Create an `Endpoint` resource for deploying the `Model` resource to.
2. Deploy the `Model` resource to the `Endpoint` resource.
3. Make online prediction requests to the `Endpoint` resource.
For batch-prediction, you:
1. Create a batch prediction job.
2. The job service will provision resources for the batch prediction request.
3. The results of the batch prediction request are returned to the caller.
4. The job service will unprovision the resoures for the batch prediction request.
## Make a batch prediction request
Now do a batch prediction to your deployed model.
### Make test items
You will use synthetic data as a test data items. Don't be concerned that we are using synthetic data -- we just want to demonstrate how to make a prediction.
```
HEADING = "petal_length,petal_width,sepal_length,sepal_width"
INSTANCE_1 = "1.4,1.3,5.1,2.8"
INSTANCE_2 = "1.5,1.2,4.7,2.4"
```
### Make the batch input file
Now make a batch input file, which you will store in your local Cloud Storage bucket. Unlike image, video and text, the batch input file for tabular is only supported for CSV. For CSV file, you make:
- The first line is the heading with the feature (fields) heading names.
- Each remaining line is a separate prediction request with the corresponding feature values.
For example:
"feature_1", "feature_2". ...
value_1, value_2, ...
```
import tensorflow as tf
gcs_input_uri = BUCKET_NAME + "/test.csv"
with tf.io.gfile.GFile(gcs_input_uri, "w") as f:
f.write(HEADING + "\n")
f.write(str(INSTANCE_1) + "\n")
f.write(str(INSTANCE_2) + "\n")
print(gcs_input_uri)
! gsutil cat $gcs_input_uri
```
### Compute instance scaling
You have several choices on scaling the compute instances for handling your batch prediction requests:
- Single Instance: The batch prediction requests are processed on a single compute instance.
- Set the minimum (`MIN_NODES`) and maximum (`MAX_NODES`) number of compute instances to one.
- Manual Scaling: The batch prediction requests are split across a fixed number of compute instances that you manually specified.
- Set the minimum (`MIN_NODES`) and maximum (`MAX_NODES`) number of compute instances to the same number of nodes. When a model is first deployed to the instance, the fixed number of compute instances are provisioned and batch prediction requests are evenly distributed across them.
- Auto Scaling: The batch prediction requests are split across a scaleable number of compute instances.
- Set the minimum (`MIN_NODES`) number of compute instances to provision when a model is first deployed and to de-provision, and set the maximum (`MAX_NODES) number of compute instances to provision, depending on load conditions.
The minimum number of compute instances corresponds to the field `min_replica_count` and the maximum number of compute instances corresponds to the field `max_replica_count`, in your subsequent deployment request.
```
MIN_NODES = 1
MAX_NODES = 1
```
### Make batch prediction request
Now that your batch of two test items is ready, let's do the batch request. Use this helper function `create_batch_prediction_job`, with the following parameters:
- `display_name`: The human readable name for the prediction job.
- `model_name`: The Vertex fully qualified identifier for the `Model` resource.
- `gcs_source_uri`: The Cloud Storage path to the input file -- which you created above.
- `gcs_destination_output_uri_prefix`: The Cloud Storage path that the service will write the predictions to.
- `parameters`: Additional filtering parameters for serving prediction results.
The helper function calls the job client service's `create_batch_prediction_job` metho, with the following parameters:
- `parent`: The Vertex location root path for Dataset, Model and Pipeline resources.
- `batch_prediction_job`: The specification for the batch prediction job.
Let's now dive into the specification for the `batch_prediction_job`:
- `display_name`: The human readable name for the prediction batch job.
- `model`: The Vertex fully qualified identifier for the `Model` resource.
- `dedicated_resources`: The compute resources to provision for the batch prediction job.
- `machine_spec`: The compute instance to provision. Use the variable you set earlier `DEPLOY_GPU != None` to use a GPU; otherwise only a CPU is allocated.
- `starting_replica_count`: The number of compute instances to initially provision, which you set earlier as the variable `MIN_NODES`.
- `max_replica_count`: The maximum number of compute instances to scale to, which you set earlier as the variable `MAX_NODES`.
- `model_parameters`: Additional filtering parameters for serving prediction results. *Note*, image segmentation models do not support additional parameters.
- `input_config`: The input source and format type for the instances to predict.
- `instances_format`: The format of the batch prediction request file: `csv` only supported.
- `gcs_source`: A list of one or more Cloud Storage paths to your batch prediction requests.
- `output_config`: The output destination and format for the predictions.
- `prediction_format`: The format of the batch prediction response file: `csv` only supported.
- `gcs_destination`: The output destination for the predictions.
This call is an asychronous operation. You will print from the response object a few select fields, including:
- `name`: The Vertex fully qualified identifier assigned to the batch prediction job.
- `display_name`: The human readable name for the prediction batch job.
- `model`: The Vertex fully qualified identifier for the Model resource.
- `generate_explanations`: Whether True/False explanations were provided with the predictions (explainability).
- `state`: The state of the prediction job (pending, running, etc).
Since this call will take a few moments to execute, you will likely get `JobState.JOB_STATE_PENDING` for `state`.
```
BATCH_MODEL = "iris_batch-" + TIMESTAMP
def create_batch_prediction_job(
display_name,
model_name,
gcs_source_uri,
gcs_destination_output_uri_prefix,
parameters=None,
):
if DEPLOY_GPU:
machine_spec = {
"machine_type": DEPLOY_COMPUTE,
"accelerator_type": DEPLOY_GPU,
"accelerator_count": DEPLOY_NGPU,
}
else:
machine_spec = {
"machine_type": DEPLOY_COMPUTE,
"accelerator_count": 0,
}
batch_prediction_job = {
"display_name": display_name,
# Format: 'projects/{project}/locations/{location}/models/{model_id}'
"model": model_name,
"model_parameters": json_format.ParseDict(parameters, Value()),
"input_config": {
"instances_format": IN_FORMAT,
"gcs_source": {"uris": [gcs_source_uri]},
},
"output_config": {
"predictions_format": OUT_FORMAT,
"gcs_destination": {"output_uri_prefix": gcs_destination_output_uri_prefix},
},
"dedicated_resources": {
"machine_spec": machine_spec,
"starting_replica_count": MIN_NODES,
"max_replica_count": MAX_NODES,
},
"generate_explanation": True,
}
response = clients["job"].create_batch_prediction_job(
parent=PARENT, batch_prediction_job=batch_prediction_job
)
print("response")
print(" name:", response.name)
print(" display_name:", response.display_name)
print(" model:", response.model)
try:
print(" generate_explanation:", response.generate_explanation)
except:
pass
print(" state:", response.state)
print(" create_time:", response.create_time)
print(" start_time:", response.start_time)
print(" end_time:", response.end_time)
print(" update_time:", response.update_time)
print(" labels:", response.labels)
return response
IN_FORMAT = "csv"
OUT_FORMAT = "csv" # [csv]
response = create_batch_prediction_job(
BATCH_MODEL, model_to_deploy_id, gcs_input_uri, BUCKET_NAME, None
)
```
Now get the unique identifier for the batch prediction job you created.
```
# The full unique ID for the batch job
batch_job_id = response.name
# The short numeric ID for the batch job
batch_job_short_id = batch_job_id.split("/")[-1]
print(batch_job_id)
```
### Get information on a batch prediction job
Use this helper function `get_batch_prediction_job`, with the following paramter:
- `job_name`: The Vertex fully qualified identifier for the batch prediction job.
The helper function calls the job client service's `get_batch_prediction_job` method, with the following paramter:
- `name`: The Vertex fully qualified identifier for the batch prediction job. In this tutorial, you will pass it the Vertex fully qualified identifier for your batch prediction job -- `batch_job_id`
The helper function will return the Cloud Storage path to where the predictions are stored -- `gcs_destination`.
```
def get_batch_prediction_job(job_name, silent=False):
response = clients["job"].get_batch_prediction_job(name=job_name)
if silent:
return response.output_config.gcs_destination.output_uri_prefix, response.state
print("response")
print(" name:", response.name)
print(" display_name:", response.display_name)
print(" model:", response.model)
try: # not all data types support explanations
print(" generate_explanation:", response.generate_explanation)
except:
pass
print(" state:", response.state)
print(" error:", response.error)
gcs_destination = response.output_config.gcs_destination
print(" gcs_destination")
print(" output_uri_prefix:", gcs_destination.output_uri_prefix)
return gcs_destination.output_uri_prefix, response.state
predictions, state = get_batch_prediction_job(batch_job_id)
```
### Get the predictions with explanations
When the batch prediction is done processing, the job state will be `JOB_STATE_SUCCEEDED`.
Finally you view the predictions and corresponding explanations stored at the Cloud Storage path you set as output. The explanations will be in a CSV format, which you indicated at the time we made the batch explanation job, under a subfolder starting with the name `prediction`, and under that folder will be a file called `explanations*.csv`.
Now display (cat) the contents. You will see one line for each explanation.
- The first four fields are the values (features) you did the prediction on.
- The remaining fields are the confidence values, between 0 and 1, for each prediction.
```
def get_latest_predictions(gcs_out_dir):
""" Get the latest prediction subfolder using the timestamp in the subfolder name"""
folders = !gsutil ls $gcs_out_dir
latest = ""
for folder in folders:
subfolder = folder.split("/")[-2]
if subfolder.startswith("prediction-"):
if subfolder > latest:
latest = folder[:-1]
return latest
while True:
predictions, state = get_batch_prediction_job(batch_job_id, True)
if state != aip.JobState.JOB_STATE_SUCCEEDED:
print("The job has not completed:", state)
if state == aip.JobState.JOB_STATE_FAILED:
raise Exception("Batch Job Failed")
else:
folder = get_latest_predictions(predictions)
! gsutil ls $folder/explanation*.csv
! gsutil cat $folder/explanation*.csv
break
time.sleep(60)
```
# Cleaning up
To clean up all GCP resources used in this project, you can [delete the GCP
project](https://cloud.google.com/resource-manager/docs/creating-managing-projects#shutting_down_projects) you used for the tutorial.
Otherwise, you can delete the individual resources you created in this tutorial:
- Dataset
- Pipeline
- Model
- Endpoint
- Batch Job
- Custom Job
- Hyperparameter Tuning Job
- Cloud Storage Bucket
```
delete_dataset = True
delete_pipeline = True
delete_model = True
delete_endpoint = True
delete_batchjob = True
delete_customjob = True
delete_hptjob = True
delete_bucket = True
# Delete the dataset using the Vertex fully qualified identifier for the dataset
try:
if delete_dataset and "dataset_id" in globals():
clients["dataset"].delete_dataset(name=dataset_id)
except Exception as e:
print(e)
# Delete the training pipeline using the Vertex fully qualified identifier for the pipeline
try:
if delete_pipeline and "pipeline_id" in globals():
clients["pipeline"].delete_training_pipeline(name=pipeline_id)
except Exception as e:
print(e)
# Delete the model using the Vertex fully qualified identifier for the model
try:
if delete_model and "model_to_deploy_id" in globals():
clients["model"].delete_model(name=model_to_deploy_id)
except Exception as e:
print(e)
# Delete the endpoint using the Vertex fully qualified identifier for the endpoint
try:
if delete_endpoint and "endpoint_id" in globals():
clients["endpoint"].delete_endpoint(name=endpoint_id)
except Exception as e:
print(e)
# Delete the batch job using the Vertex fully qualified identifier for the batch job
try:
if delete_batchjob and "batch_job_id" in globals():
clients["job"].delete_batch_prediction_job(name=batch_job_id)
except Exception as e:
print(e)
# Delete the custom job using the Vertex fully qualified identifier for the custom job
try:
if delete_customjob and "job_id" in globals():
clients["job"].delete_custom_job(name=job_id)
except Exception as e:
print(e)
# Delete the hyperparameter tuning job using the Vertex fully qualified identifier for the hyperparameter tuning job
try:
if delete_hptjob and "hpt_job_id" in globals():
clients["job"].delete_hyperparameter_tuning_job(name=hpt_job_id)
except Exception as e:
print(e)
if delete_bucket and "BUCKET_NAME" in globals():
! gsutil rm -r $BUCKET_NAME
```
| github_jupyter |
# Software Development 1
Topics for today will include:
- Javadoc
- Reference Data Types
- Linear Data Structures
- Arrays
- ArrayLists
- Queues
- LinkedList
- Stacks
- Revisiting Objects
## JavaDoc
Coming off of a lab dealing with Markdown, discussing documentation seems like a good place to go next. Documentation is something that is super simple in concept but rather difficult to implement. At it's core documentation is simple. It's a set of instructions on how to use a product. When you really start to think about it though, it can quickly balloon into something rather complex.
When thinking about documentation, is your audience:
- Tech Savvy
- Tech Illiterate
- Younger
- Older
- Familiary With The Technology
- A Developer Themselves
- A Malicious User
Thinking about all of these different scenarios you'd potentially write to them all in a different way. For someone that's Tech Savvy you may not need to give them as much detail because they may get the underlying theme of how your technology works and just need you to tell them how to implement it.
For some one younger you may be able to use parallels to compare it to something they understand so that they get it, but that same example doesn't make any sense to someone older.
We also need to alter our documentation for the medium that it's going to be consumed. As tech has gotten better we have the ability to make out lives easier by adding things like IntelliJ and VS Code into our arsenals. These both come with something called IntelliSense that can tell us what a function does before we commit to using it.
Now the way that it does this is through the proper documentation of code. In Java that's done by using Java Doc. Java Doc in the wholistic sense will write documentation and make a static webpage with that documentation based on the way that you comment your code. On the one hand people may think that documenting code is kind of tedious. On the other having a tool like Java Doc makes it very rewarding.
For the purpose of this class we won't go to the full lengths there but we'll use the commenting style to add usage tips to our functions. As we talk through our lesson today.
## What Are Reference Data Types Again?
## Linear Data Structures
What are linear data structures and why do we need them?
First off linear data structure are comprised of a type of data and can be accessed sequentially. A good example of this that we already understand is an array.
What are our problems with arrays in Java though? Why can't we just use those?
### Arrays
Arrays are our initial linear data stucture and as we all know we put object in and they're ordered and accessible through providing an index when referencing the array.
```
int[] numbers = {0, 1, 2, 3, 4, 5};
System.out.println("Printing out list with for loop:");
for (int i = 0; i < numbers.length; i++){
System.out.println(numbers[i]);
}
System.out.println("\nPrinting out list with for each loop:");
for (int number: numbers) {
System.out.println(number);
}
```
Going through both lists we get the same thing no matter the method. This is because we're iterating through it sequentially and there's an order to our array. We have 3 other examples of this that we need to understand for certain scenarios that we could come across
First let's look at an ArrayList
### Array List
Array Lists are great because it gives us the ability to have an mutable array when it comes to size. Callback to before. The issue that we run into mainly with primitave arrays is that they're immutable in size. Meaning that after we make it the first time we can't make it bigger. We have what we have.
Sucks...
But we can actually just use an ArrayList to get around that. ArrayList and Arrays are basically the same functionallity wise besides that one detail. The syntax for some of these things do change. The underlying concepts are the same though!
```
import java.util.ArrayList;
ArrayList<Integer> listExample1 = new ArrayList();
// Preferred with out data in the liust to start.
ArrayList<Integer> listExample2 = new ArrayList<Integer>();
ArrayList<Integer> numbers = new ArrayList(Arrays.asList(0,1,2,3,4,5));
ArrayList<Integer> numbers2 = new ArrayList<Integer>(Arrays.asList(0,1,2,3,4,5));
int demo = 6;
System.out.println("Printing out list with for loop:");
for (int i = 0; i < numbers.size(); i++){
System.out.println(numbers.get(i));
}
System.out.println("\nPrinting out list with for each loop:");
for (int number: numbers) {
System.out.println(number);
}
```
So as we can see here we're able to do the same things that we did above with the ArrayList. Some of the keywords and methods changed a little bit but things are still the same.
Now let's try and add something to out array! Then we can reprint these things and see if they actually change!
```
numbers.add(6);
numbers.add(7);
numbers.add(8);
numbers.add(9);
numbers.add(10);
System.out.println("Printing out list with for loop:");
for (int i = 0; i < numbers.size(); i++){
System.out.println(numbers.get(i));
}
System.out.println("\nPrinting out list with for each loop:");
for (int number: numbers) {
System.out.println(number);
}
numbers.add(1, 10);
System.out.println(numbers)
```
Just like that we're able to add elements to our list! Pretty painless!
Now let's take it up a little bit and talk about Linked Lists!
## LinkedLists
Now LinkedList are again similar to ArrayLists we we should actually go and learn how both of these things are being created under the covers!
ArrayList have spots assigned to them and then we assign items to those spots. So we grab a contiguous block of storage and put all of the stuff we're storing there. Think when you go to the movies with friends. If there are four of you, you try and find 4 available seats all next to each other so that you can sit together. Same idea for Arrays and Array Lists.
Now when it comes to LinkedList lets morph that. Let's say that you want to give something to a friends sibling. You'd probably give the thing to your friend and then your friend would give it to the sibling. This works because you know where to find your friend and your friend knows where to find their sibling.
That knowledge is what we'd call a pointer. Pointers are used all over the place in LinkedLists.
```
import java.util.LinkedList;
LinkedList<String> recipe = new LinkedList<String>();
recipe.add("Bottom Bun");
recipe.add("Mayo");
recipe.add("Lettuce");
recipe.add("Tomato");
recipe.add("Burger Patty");
recipe.add("Cheese");
recipe.add("Bacon");
recipe.add("Mayo");
recipe.add("Ketchup");
recipe.add("Top Bun");
System.out.println(recipe);
```
This is done differently under the covers than an array list. When an arraylist runs out of space that becomes very pricey in memory to move everything.
We can circumnavigate that by using LinkedLists and pointers. Now in terms of code you'll rarely see a visual difference unless you're paying attention to preformance but when it comes to knowing data type and how they work this knowledge is invaluable.
Let's look at manipulating our created list a little bit where we move some things around.
```
recipe.add(2, "Pickles");
recipe.add(6, "Burger Patty");
recipe.add(6, "Cheese");
recipe.remove("Tomato");
System.out.println(recipe);
```
Here, same as before it's not much effort to move things around. What's important to note here is that we can put the index that we want the place of the item to be at along with the data and the LinkedList will figure out the rest. Fixing the pointers and what not under the covers.
Next up we'll talk about Queues
### Queues
So similar to real life we have lines to process things.
To do this in code we can use a Queue!
Queues are generally FIFO(First In First Out) Queue's will be our first exposure to interfaces which we'll talk about later. They're initialized by using other reference types. The two common ones are LinkedLists which we just learned about and PriorityQueues.
```
import java.util.Queue;
Queue<String> line = new LinkedList<String>();
line.add("Chris");
line.add("Sharon");
line.add("Ava");
line.add("Tim");
System.out.println(line);
```
Unlike some of our previous examples when we grab something here we're actually taking it completely out of the list.
```
String headOfLine = line.remove();
System.out.println(line);
System.out.println(headOfLine);
System.out.println(line.element());
System.out.println(line);
```
This again is just like a line and isn't that complex.
Getting through that allows us to move onto the last type that we'll go through(There are many more of these linear data types) We've talked about Stacks so lets see one in action!
### Stacks
Stack we talked about last week and out example was a stack of plates. It's LIFO (Last In First Out) the best example that I have for this is when we're dealing with recursion.
I have an example that we can look at in the QuickDemos folder under Fibonacci.
```
import java.util.Stack;
Stack<String> stack = new Stack<String>();
stack.push("One");
stack.push("Two");
stack.push("Three");
stack.push("Four");
stack.push("Five");
System.out.println(stack);
```
Now when we remove the items it's going to come from the end of the list. Not the top.
```
stack.pop();
stack.pop();
System.out.println(stack);
```
Next we're going to move on to how all of these are relate and how they came to be!
For that we're going to talk about Objects!
| github_jupyter |
# Richter's Nepal Earthquake
```
#Importing the Libraries
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
sns.set(style="white", context="notebook", palette="deep", color_codes=True)
from collections import Counter
from sklearn.preprocessing import StandardScaler
from sklearn.decomposition import PCA
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score, confusion_matrix, f1_score
#Importing the Data
train = pd.read_csv(r'...\train_values.csv')
trainy = pd.read_csv(r'...\train_labels.csv')
train = train.join(trainy['damage_grade'])
train.head()
train = train.fillna(np.nan)
train.isnull().sum()
# Outlier detection
def detect_outliers(df,n,features):
"""
Takes a dataframe df of features and returns a list of the indices
corresponding to the observations containing more than n outliers according
to the Tukey method.
"""
outlier_indices = []
# iterate over features(columns)
for col in features:
# 1st quartile (25%)
Q1 = np.percentile(df[col], 25)
# 3rd quartile (75%)
Q3 = np.percentile(df[col],75)
# Interquartile range (IQR)
IQR = Q3 - Q1
# outlier step
outlier_step = 1.5 * IQR
# Determine a list of indices of outliers for feature col
outlier_list_col = df[(df[col] < Q1 - outlier_step) | (df[col] > Q3 + outlier_step )].index
# append the found outlier indices for col to the list of outlier indices
outlier_indices.extend(outlier_list_col)
# select observations containing more than 2 outliers
outlier_indices = Counter(outlier_indices)
multiple_outliers = list( k for k, v in outlier_indices.items() if v > n )
return multiple_outliers
# detect outliers
Outliers_to_drop = detect_outliers(train,2,["count_floors_pre_eq", "age", "area_percentage", "height_percentage", "count_families"])
train.loc[Outliers_to_drop]
# Drop outliers
train = train.drop(Outliers_to_drop, axis = 0).reset_index(drop=True)
plt.figure(figsize=(15,10))
c = sns.heatmap(train[["geo_level_1_id","geo_level_2_id", "geo_level_3_id", "count_floors_pre_eq", "age", "area_percentage", "height_percentage",
"count_families", "damage_grade"]].corr(), annot=True, fmt="0.2f")
plt.figure(figsize=(20,15))
sns.barplot(train['geo_level_1_id'], train['damage_grade'])
sns.barplot(train['count_floors_pre_eq'], train['damage_grade'])
plt.figure(figsize=(15,10))
sns.barplot(train['age'], train['damage_grade'])
plt.figure(figsize=(15,10))
sns.barplot(train['area_percentage'], train['damage_grade'])
plt.figure(figsize=(15,10))
sns.barplot(train['count_families'], train['damage_grade'])
plt.figure(figsize=(15,10))
sns.barplot(train['height_percentage'], train['damage_grade'])
trainy = train['damage_grade']
trainy.shape
# train = train.drop("damage_grade", axis=1)
dataset = train.append(test, ignore_index = True)
dataset = dataset.drop(["geo_level_2_id", "geo_level_3_id"], axis=1)
```
## Feature Engineering
```
dataset['lowHeight'] = dataset['height_percentage'].map(lambda s: 1 if s <= 11 else 0)
dataset['medHeight'] = dataset['height_percentage'].map(lambda s: 1 if 12 <= s <= 23 else 0)
dataset['highHeight'] = dataset['height_percentage'].map(lambda s: 1 if 24 <= s else 0)
sns.barplot("highHeight", "damage_grade", data=dataset)
dataset['lessFloors'] = dataset['count_floors_pre_eq'].map(lambda s: 1 if s <= 5 else 0)
dataset['lotFloors'] = dataset['count_floors_pre_eq'].map(lambda s: 1 if s >= 6 else 0)
train.shape
dataset = pd.get_dummies(dataset, columns = ["land_surface_condition"], prefix="LSC")
dataset = pd.get_dummies(dataset, columns = ["foundation_type"], prefix="FT")
dataset = pd.get_dummies(dataset, columns = ["roof_type"], prefix="RT")
dataset = pd.get_dummies(dataset, columns = ["ground_floor_type"], prefix="GFT")
dataset = pd.get_dummies(dataset, columns = ["other_floor_type"], prefix="OFT")
dataset = pd.get_dummies(dataset, columns = ["position"], prefix="P")
dataset = pd.get_dummies(dataset, columns = ["plan_configuration"], prefix="PC")
dataset = pd.get_dummies(dataset, columns = ["legal_ownership_status"], prefix="LOS")
X_train = dataset[:254104]
X_test = dataset[254104:]
X_train = X_train.drop(["building_id", "damage_grade"], axis=1)
X_test = X_test.drop(["building_id", "damage_grade"], axis=1)
scaler = StandardScaler()
X_train = scaler.fit_transform(X_train)
X_test = scaler.transform(X_test)
pca = PCA(n_components=71)
pca.fit(X_train)
print(pca.explained_variance_ratio_.cumsum())
pca_fin = PCA(n_components=55)
X_train = pca_fin.fit_transform(X_train)
X_test = pca_fin.transform(X_test)
y_train = trainy.iloc[:].values
y_train
X_train_split , X_test_split, y_train_split, y_test_split = train_test_split(X_train, y_train, test_size=0.2, random_state=42)
classifier = RandomForestClassifier(n_estimators=300, n_jobs=4)
classifier.fit(X_train_split,y_train_split)
y_pred = classifier.predict(X_test_split)
print(accuracy_score(y_test_split, y_pred))
print(confusion_matrix(y_test_split, y_pred))
print(f1_score(y_test_split, y_pred, average='micro'))
```
| github_jupyter |
## Dependencies
```
import os
import sys
import cv2
import shutil
import random
import warnings
import numpy as np
import pandas as pd
import seaborn as sns
import multiprocessing as mp
import matplotlib.pyplot as plt
from tensorflow import set_random_seed
from sklearn.utils import class_weight
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix, cohen_kappa_score
from keras import backend as K
from keras.models import Model
from keras.utils import to_categorical
from keras import optimizers, applications
from keras.preprocessing.image import ImageDataGenerator
from keras.layers import Dense, Dropout, GlobalAveragePooling2D, Input
from keras.callbacks import EarlyStopping, ReduceLROnPlateau, Callback, LearningRateScheduler, ModelCheckpoint
def seed_everything(seed=0):
random.seed(seed)
os.environ['PYTHONHASHSEED'] = str(seed)
np.random.seed(seed)
set_random_seed(0)
seed = 0
seed_everything(seed)
%matplotlib inline
sns.set(style="whitegrid")
warnings.filterwarnings("ignore")
sys.path.append(os.path.abspath('../input/efficientnet/efficientnet-master/efficientnet-master/'))
from efficientnet import *
```
## Load data
```
fold_set = pd.read_csv('../input/aptos-data-split/5-fold.csv')
X_train = fold_set[fold_set['fold_3'] == 'train']
X_val = fold_set[fold_set['fold_3'] == 'validation']
test = pd.read_csv('../input/aptos2019-blindness-detection/test.csv')
# Preprocecss data
X_train["id_code"] = X_train["id_code"].apply(lambda x: x + ".png")
X_val["id_code"] = X_val["id_code"].apply(lambda x: x + ".png")
test["id_code"] = test["id_code"].apply(lambda x: x + ".png")
print('Number of train samples: ', X_train.shape[0])
print('Number of validation samples: ', X_val.shape[0])
print('Number of test samples: ', test.shape[0])
display(X_train.head())
```
# Model parameters
```
# Model parameters
model_path = '../working/effNetB4_img256_noBen_fold4.h5'
FACTOR = 4
BATCH_SIZE = 8 * FACTOR
EPOCHS = 10
WARMUP_EPOCHS = 5
LEARNING_RATE = 1e-4 * FACTOR
WARMUP_LEARNING_RATE = 1e-3 * FACTOR
HEIGHT = 256
WIDTH = 256
CHANNELS = 3
TTA_STEPS = 1
ES_PATIENCE = 5
RLROP_PATIENCE = 3
LR_WARMUP_EPOCHS = 3
STEP_SIZE = len(X_train) // BATCH_SIZE
TOTAL_STEPS = EPOCHS * STEP_SIZE
WARMUP_STEPS = LR_WARMUP_EPOCHS * STEP_SIZE
```
# Pre-procecess images
```
new_data_base_path = '../input/aptos2019-blindness-detection/train_images/'
test_base_path = '../input/aptos2019-blindness-detection/test_images/'
train_dest_path = 'base_dir/train_images/'
validation_dest_path = 'base_dir/validation_images/'
test_dest_path = 'base_dir/test_images/'
# Making sure directories don't exist
if os.path.exists(train_dest_path):
shutil.rmtree(train_dest_path)
if os.path.exists(validation_dest_path):
shutil.rmtree(validation_dest_path)
if os.path.exists(test_dest_path):
shutil.rmtree(test_dest_path)
# Creating train, validation and test directories
os.makedirs(train_dest_path)
os.makedirs(validation_dest_path)
os.makedirs(test_dest_path)
def crop_image(img, tol=7):
if img.ndim ==2:
mask = img>tol
return img[np.ix_(mask.any(1),mask.any(0))]
elif img.ndim==3:
gray_img = cv2.cvtColor(img, cv2.COLOR_RGB2GRAY)
mask = gray_img>tol
check_shape = img[:,:,0][np.ix_(mask.any(1),mask.any(0))].shape[0]
if (check_shape == 0): # image is too dark so that we crop out everything,
return img # return original image
else:
img1=img[:,:,0][np.ix_(mask.any(1),mask.any(0))]
img2=img[:,:,1][np.ix_(mask.any(1),mask.any(0))]
img3=img[:,:,2][np.ix_(mask.any(1),mask.any(0))]
img = np.stack([img1,img2,img3],axis=-1)
return img
def circle_crop(img):
img = crop_image(img)
height, width, depth = img.shape
largest_side = np.max((height, width))
img = cv2.resize(img, (largest_side, largest_side))
height, width, depth = img.shape
x = width//2
y = height//2
r = np.amin((x, y))
circle_img = np.zeros((height, width), np.uint8)
cv2.circle(circle_img, (x, y), int(r), 1, thickness=-1)
img = cv2.bitwise_and(img, img, mask=circle_img)
img = crop_image(img)
return img
def preprocess_image(image_id, base_path, save_path, HEIGHT=HEIGHT, WIDTH=WIDTH, sigmaX=10):
image = cv2.imread(base_path + image_id)
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
image = circle_crop(image)
image = cv2.resize(image, (HEIGHT, WIDTH))
# image = cv2.addWeighted(image, 4, cv2.GaussianBlur(image, (0,0), sigmaX), -4 , 128)
cv2.imwrite(save_path + image_id, image)
def preprocess_data(df, HEIGHT=HEIGHT, WIDTH=WIDTH, sigmaX=10):
df = df.reset_index()
for i in range(df.shape[0]):
item = df.iloc[i]
image_id = item['id_code']
item_set = item['fold_3']
if item_set == 'train':
preprocess_image(image_id, new_data_base_path, train_dest_path)
if item_set == 'validation':
preprocess_image(image_id, new_data_base_path, validation_dest_path)
def preprocess_test(df, base_path=test_base_path, save_path=test_dest_path, HEIGHT=HEIGHT, WIDTH=WIDTH, sigmaX=10):
df = df.reset_index()
for i in range(df.shape[0]):
image_id = df.iloc[i]['id_code']
preprocess_image(image_id, base_path, save_path)
n_cpu = mp.cpu_count()
train_n_cnt = X_train.shape[0] // n_cpu
val_n_cnt = X_val.shape[0] // n_cpu
test_n_cnt = test.shape[0] // n_cpu
# Pre-procecss old data train set
pool = mp.Pool(n_cpu)
dfs = [X_train.iloc[train_n_cnt*i:train_n_cnt*(i+1)] for i in range(n_cpu)]
dfs[-1] = X_train.iloc[train_n_cnt*(n_cpu-1):]
res = pool.map(preprocess_data, [x_df for x_df in dfs])
pool.close()
# Pre-procecss validation set
pool = mp.Pool(n_cpu)
dfs = [X_val.iloc[val_n_cnt*i:val_n_cnt*(i+1)] for i in range(n_cpu)]
dfs[-1] = X_val.iloc[val_n_cnt*(n_cpu-1):]
res = pool.map(preprocess_data, [x_df for x_df in dfs])
pool.close()
# Pre-procecss test set
pool = mp.Pool(n_cpu)
dfs = [test.iloc[test_n_cnt*i:test_n_cnt*(i+1)] for i in range(n_cpu)]
dfs[-1] = test.iloc[test_n_cnt*(n_cpu-1):]
res = pool.map(preprocess_test, [x_df for x_df in dfs])
pool.close()
```
# Data generator
```
datagen=ImageDataGenerator(rescale=1./255,
rotation_range=360,
horizontal_flip=True,
vertical_flip=True)
train_generator=datagen.flow_from_dataframe(
dataframe=X_train,
directory=train_dest_path,
x_col="id_code",
y_col="diagnosis",
class_mode="raw",
batch_size=BATCH_SIZE,
target_size=(HEIGHT, WIDTH),
seed=seed)
valid_generator=datagen.flow_from_dataframe(
dataframe=X_val,
directory=validation_dest_path,
x_col="id_code",
y_col="diagnosis",
class_mode="raw",
batch_size=BATCH_SIZE,
target_size=(HEIGHT, WIDTH),
seed=seed)
test_generator=datagen.flow_from_dataframe(
dataframe=test,
directory=test_dest_path,
x_col="id_code",
batch_size=1,
class_mode=None,
shuffle=False,
target_size=(HEIGHT, WIDTH),
seed=seed)
def classify(x):
if x < 0.5:
return 0
elif x < 1.5:
return 1
elif x < 2.5:
return 2
elif x < 3.5:
return 3
return 4
labels = ['0 - No DR', '1 - Mild', '2 - Moderate', '3 - Severe', '4 - Proliferative DR']
def plot_confusion_matrix(train, validation, labels=labels):
train_labels, train_preds = train
validation_labels, validation_preds = validation
fig, (ax1, ax2) = plt.subplots(1, 2, sharex='col', figsize=(24, 7))
train_cnf_matrix = confusion_matrix(train_labels, train_preds)
validation_cnf_matrix = confusion_matrix(validation_labels, validation_preds)
train_cnf_matrix_norm = train_cnf_matrix.astype('float') / train_cnf_matrix.sum(axis=1)[:, np.newaxis]
validation_cnf_matrix_norm = validation_cnf_matrix.astype('float') / validation_cnf_matrix.sum(axis=1)[:, np.newaxis]
train_df_cm = pd.DataFrame(train_cnf_matrix_norm, index=labels, columns=labels)
validation_df_cm = pd.DataFrame(validation_cnf_matrix_norm, index=labels, columns=labels)
sns.heatmap(train_df_cm, annot=True, fmt='.2f', cmap="Blues",ax=ax1).set_title('Train')
sns.heatmap(validation_df_cm, annot=True, fmt='.2f', cmap=sns.cubehelix_palette(8),ax=ax2).set_title('Validation')
plt.show()
def plot_metrics(history, figsize=(20, 14)):
fig, (ax1, ax2) = plt.subplots(2, 1, sharex='col', figsize=figsize)
ax1.plot(history['loss'], label='Train loss')
ax1.plot(history['val_loss'], label='Validation loss')
ax1.legend(loc='best')
ax1.set_title('Loss')
ax2.plot(history['acc'], label='Train accuracy')
ax2.plot(history['val_acc'], label='Validation accuracy')
ax2.legend(loc='best')
ax2.set_title('Accuracy')
plt.xlabel('Epochs')
sns.despine()
plt.show()
def apply_tta(model, generator, steps=10):
step_size = generator.n//generator.batch_size
preds_tta = []
for i in range(steps):
generator.reset()
preds = model.predict_generator(generator, steps=step_size)
preds_tta.append(preds)
return np.mean(preds_tta, axis=0)
def evaluate_model(train, validation):
train_labels, train_preds = train
validation_labels, validation_preds = validation
print("Train Cohen Kappa score: %.3f" % cohen_kappa_score(train_preds, train_labels, weights='quadratic'))
print("Validation Cohen Kappa score: %.3f" % cohen_kappa_score(validation_preds, validation_labels, weights='quadratic'))
print("Complete set Cohen Kappa score: %.3f" % cohen_kappa_score(np.append(train_preds, validation_preds), np.append(train_labels, validation_labels), weights='quadratic'))
def cosine_decay_with_warmup(global_step,
learning_rate_base,
total_steps,
warmup_learning_rate=0.0,
warmup_steps=0,
hold_base_rate_steps=0):
"""
Cosine decay schedule with warm up period.
In this schedule, the learning rate grows linearly from warmup_learning_rate
to learning_rate_base for warmup_steps, then transitions to a cosine decay
schedule.
:param global_step {int}: global step.
:param learning_rate_base {float}: base learning rate.
:param total_steps {int}: total number of training steps.
:param warmup_learning_rate {float}: initial learning rate for warm up. (default: {0.0}).
:param warmup_steps {int}: number of warmup steps. (default: {0}).
:param hold_base_rate_steps {int}: Optional number of steps to hold base learning rate before decaying. (default: {0}).
:param global_step {int}: global step.
:Returns : a float representing learning rate.
:Raises ValueError: if warmup_learning_rate is larger than learning_rate_base, or if warmup_steps is larger than total_steps.
"""
if total_steps < warmup_steps:
raise ValueError('total_steps must be larger or equal to warmup_steps.')
learning_rate = 0.5 * learning_rate_base * (1 + np.cos(
np.pi *
(global_step - warmup_steps - hold_base_rate_steps
) / float(total_steps - warmup_steps - hold_base_rate_steps)))
if hold_base_rate_steps > 0:
learning_rate = np.where(global_step > warmup_steps + hold_base_rate_steps,
learning_rate, learning_rate_base)
if warmup_steps > 0:
if learning_rate_base < warmup_learning_rate:
raise ValueError('learning_rate_base must be larger or equal to warmup_learning_rate.')
slope = (learning_rate_base - warmup_learning_rate) / warmup_steps
warmup_rate = slope * global_step + warmup_learning_rate
learning_rate = np.where(global_step < warmup_steps, warmup_rate,
learning_rate)
return np.where(global_step > total_steps, 0.0, learning_rate)
class WarmUpCosineDecayScheduler(Callback):
"""Cosine decay with warmup learning rate scheduler"""
def __init__(self,
learning_rate_base,
total_steps,
global_step_init=0,
warmup_learning_rate=0.0,
warmup_steps=0,
hold_base_rate_steps=0,
verbose=0):
"""
Constructor for cosine decay with warmup learning rate scheduler.
:param learning_rate_base {float}: base learning rate.
:param total_steps {int}: total number of training steps.
:param global_step_init {int}: initial global step, e.g. from previous checkpoint.
:param warmup_learning_rate {float}: initial learning rate for warm up. (default: {0.0}).
:param warmup_steps {int}: number of warmup steps. (default: {0}).
:param hold_base_rate_steps {int}: Optional number of steps to hold base learning rate before decaying. (default: {0}).
:param verbose {int}: quiet, 1: update messages. (default: {0}).
"""
super(WarmUpCosineDecayScheduler, self).__init__()
self.learning_rate_base = learning_rate_base
self.total_steps = total_steps
self.global_step = global_step_init
self.warmup_learning_rate = warmup_learning_rate
self.warmup_steps = warmup_steps
self.hold_base_rate_steps = hold_base_rate_steps
self.verbose = verbose
self.learning_rates = []
def on_batch_end(self, batch, logs=None):
self.global_step = self.global_step + 1
lr = K.get_value(self.model.optimizer.lr)
self.learning_rates.append(lr)
def on_batch_begin(self, batch, logs=None):
lr = cosine_decay_with_warmup(global_step=self.global_step,
learning_rate_base=self.learning_rate_base,
total_steps=self.total_steps,
warmup_learning_rate=self.warmup_learning_rate,
warmup_steps=self.warmup_steps,
hold_base_rate_steps=self.hold_base_rate_steps)
K.set_value(self.model.optimizer.lr, lr)
if self.verbose > 0:
print('\nBatch %02d: setting learning rate to %s.' % (self.global_step + 1, lr))
class RAdam(optimizers.Optimizer):
"""RAdam optimizer.
# Arguments
lr: float >= 0. Learning rate.
beta_1: float, 0 < beta < 1. Generally close to 1.
beta_2: float, 0 < beta < 1. Generally close to 1.
epsilon: float >= 0. Fuzz factor. If `None`, defaults to `K.epsilon()`.
decay: float >= 0. Learning rate decay over each update.
weight_decay: float >= 0. Weight decay for each param.
amsgrad: boolean. Whether to apply the AMSGrad variant of this
algorithm from the paper "On the Convergence of Adam and
Beyond".
# References
- [Adam - A Method for Stochastic Optimization](https://arxiv.org/abs/1412.6980v8)
- [On the Convergence of Adam and Beyond](https://openreview.net/forum?id=ryQu7f-RZ)
- [On The Variance Of The Adaptive Learning Rate And Beyond](https://arxiv.org/pdf/1908.03265v1.pdf)
"""
def __init__(self, lr=0.001, beta_1=0.9, beta_2=0.999,
epsilon=None, decay=0., weight_decay=0., amsgrad=False, **kwargs):
super(RAdam, self).__init__(**kwargs)
with K.name_scope(self.__class__.__name__):
self.iterations = K.variable(0, dtype='int64', name='iterations')
self.lr = K.variable(lr, name='lr')
self.beta_1 = K.variable(beta_1, name='beta_1')
self.beta_2 = K.variable(beta_2, name='beta_2')
self.decay = K.variable(decay, name='decay')
self.weight_decay = K.variable(weight_decay, name='weight_decay')
if epsilon is None:
epsilon = K.epsilon()
self.epsilon = epsilon
self.initial_decay = decay
self.initial_weight_decay = weight_decay
self.amsgrad = amsgrad
def get_updates(self, loss, params):
grads = self.get_gradients(loss, params)
self.updates = [K.update_add(self.iterations, 1)]
lr = self.lr
if self.initial_decay > 0:
lr = lr * (1. / (1. + self.decay * K.cast(self.iterations, K.dtype(self.decay))))
t = K.cast(self.iterations, K.floatx()) + 1
ms = [K.zeros(K.int_shape(p), dtype=K.dtype(p), name='m_' + str(i)) for (i, p) in enumerate(params)]
vs = [K.zeros(K.int_shape(p), dtype=K.dtype(p), name='v_' + str(i)) for (i, p) in enumerate(params)]
if self.amsgrad:
vhats = [K.zeros(K.int_shape(p), dtype=K.dtype(p), name='vhat_' + str(i)) for (i, p) in enumerate(params)]
else:
vhats = [K.zeros(1, name='vhat_' + str(i)) for i in range(len(params))]
self.weights = [self.iterations] + ms + vs + vhats
beta_1_t = K.pow(self.beta_1, t)
beta_2_t = K.pow(self.beta_2, t)
sma_inf = 2.0 / (1.0 - self.beta_2) - 1.0
sma_t = sma_inf - 2.0 * t * beta_2_t / (1.0 - beta_2_t)
for p, g, m, v, vhat in zip(params, grads, ms, vs, vhats):
m_t = (self.beta_1 * m) + (1. - self.beta_1) * g
v_t = (self.beta_2 * v) + (1. - self.beta_2) * K.square(g)
m_corr_t = m_t / (1.0 - beta_1_t)
if self.amsgrad:
vhat_t = K.maximum(vhat, v_t)
v_corr_t = K.sqrt(vhat_t / (1.0 - beta_2_t) + self.epsilon)
self.updates.append(K.update(vhat, vhat_t))
else:
v_corr_t = K.sqrt(v_t / (1.0 - beta_2_t) + self.epsilon)
r_t = K.sqrt((sma_t - 4.0) / (sma_inf - 4.0) *
(sma_t - 2.0) / (sma_inf - 2.0) *
sma_inf / sma_t)
p_t = K.switch(sma_t > 5, r_t * m_corr_t / v_corr_t, m_corr_t)
if self.initial_weight_decay > 0:
p_t += self.weight_decay * p
p_t = p - lr * p_t
self.updates.append(K.update(m, m_t))
self.updates.append(K.update(v, v_t))
new_p = p_t
# Apply constraints.
if getattr(p, 'constraint', None) is not None:
new_p = p.constraint(new_p)
self.updates.append(K.update(p, new_p))
return self.updates
def get_config(self):
config = {
'lr': float(K.get_value(self.lr)),
'beta_1': float(K.get_value(self.beta_1)),
'beta_2': float(K.get_value(self.beta_2)),
'decay': float(K.get_value(self.decay)),
'weight_decay': float(K.get_value(self.weight_decay)),
'epsilon': self.epsilon,
'amsgrad': self.amsgrad,
}
base_config = super(RAdam, self).get_config()
return dict(list(base_config.items()) + list(config.items()))
```
# Model
```
def create_model(input_shape):
input_tensor = Input(shape=input_shape)
base_model = EfficientNetB4(weights=None,
include_top=False,
input_tensor=input_tensor)
# base_model.load_weights('../input/efficientnet-keras-weights-b0b5/efficientnet-b5_imagenet_1000_notop.h5')
x = GlobalAveragePooling2D()(base_model.output)
final_output = Dense(1, activation='linear', name='final_output')(x)
model = Model(input_tensor, final_output)
model.load_weights('../input/aptos-pretrain-olddata-effnetb4/effNetB4_img224_noBen_oldData.h5')
return model
```
# Train top layers
```
model = create_model(input_shape=(HEIGHT, WIDTH, CHANNELS))
for layer in model.layers:
layer.trainable = False
for i in range(-2, 0):
model.layers[i].trainable = True
metric_list = ["accuracy"]
optimizer = RAdam(lr=WARMUP_LEARNING_RATE)
model.compile(optimizer=optimizer, loss='mean_squared_error', metrics=metric_list)
model.summary()
STEP_SIZE_TRAIN = train_generator.n//train_generator.batch_size
STEP_SIZE_VALID = valid_generator.n//valid_generator.batch_size
history_warmup = model.fit_generator(generator=train_generator,
steps_per_epoch=STEP_SIZE_TRAIN,
validation_data=valid_generator,
validation_steps=STEP_SIZE_VALID,
epochs=WARMUP_EPOCHS,
verbose=2).history
```
# Fine-tune the model
```
for layer in model.layers:
layer.trainable = True
checkpoint = ModelCheckpoint(model_path, monitor='val_loss', mode='min', save_best_only=True, save_weights_only=True)
es = EarlyStopping(monitor='val_loss', mode='min', patience=ES_PATIENCE, restore_best_weights=True, verbose=1)
cosine_lr = WarmUpCosineDecayScheduler(learning_rate_base=LEARNING_RATE,
total_steps=TOTAL_STEPS,
warmup_learning_rate=0.0,
warmup_steps=WARMUP_STEPS,
hold_base_rate_steps=(2 * STEP_SIZE))
callback_list = [checkpoint, es, cosine_lr]
optimizer = RAdam(lr=LEARNING_RATE)
model.compile(optimizer=optimizer, loss='mean_squared_error', metrics=metric_list)
model.summary()
history = model.fit_generator(generator=train_generator,
steps_per_epoch=STEP_SIZE_TRAIN,
validation_data=valid_generator,
validation_steps=STEP_SIZE_VALID,
epochs=EPOCHS,
callbacks=callback_list,
verbose=2).history
fig, ax = plt.subplots(1, 1, sharex='col', figsize=(20, 4))
ax.plot(cosine_lr.learning_rates)
ax.set_title('Fine-tune learning rates')
plt.xlabel('Steps')
plt.ylabel('Learning rate')
sns.despine()
plt.show()
```
# Model loss graph
```
plot_metrics(history)
# Create empty arays to keep the predictions and labels
df_preds = pd.DataFrame(columns=['label', 'pred', 'set'])
train_generator.reset()
valid_generator.reset()
# Add train predictions and labels
for i in range(STEP_SIZE_TRAIN + 1):
im, lbl = next(train_generator)
preds = model.predict(im, batch_size=train_generator.batch_size)
for index in range(len(preds)):
df_preds.loc[len(df_preds)] = [lbl[index], preds[index][0], 'train']
# Add validation predictions and labels
for i in range(STEP_SIZE_VALID + 1):
im, lbl = next(valid_generator)
preds = model.predict(im, batch_size=valid_generator.batch_size)
for index in range(len(preds)):
df_preds.loc[len(df_preds)] = [lbl[index], preds[index][0], 'validation']
df_preds['label'] = df_preds['label'].astype('int')
# Classify predictions
df_preds['predictions'] = df_preds['pred'].apply(lambda x: classify(x))
train_preds = df_preds[df_preds['set'] == 'train']
validation_preds = df_preds[df_preds['set'] == 'validation']
```
# Model Evaluation
## Confusion Matrix
### Original thresholds
```
plot_confusion_matrix((train_preds['label'], train_preds['predictions']), (validation_preds['label'], validation_preds['predictions']))
```
## Quadratic Weighted Kappa
```
evaluate_model((train_preds['label'], train_preds['predictions']), (validation_preds['label'], validation_preds['predictions']))
```
## Apply model to test set and output predictions
```
preds = apply_tta(model, test_generator, TTA_STEPS)
predictions = [classify(x) for x in preds]
results = pd.DataFrame({'id_code':test['id_code'], 'diagnosis':predictions})
results['id_code'] = results['id_code'].map(lambda x: str(x)[:-4])
# Cleaning created directories
if os.path.exists(train_dest_path):
shutil.rmtree(train_dest_path)
if os.path.exists(validation_dest_path):
shutil.rmtree(validation_dest_path)
if os.path.exists(test_dest_path):
shutil.rmtree(test_dest_path)
```
# Predictions class distribution
```
fig = plt.subplots(sharex='col', figsize=(24, 8.7))
sns.countplot(x="diagnosis", data=results, palette="GnBu_d").set_title('Test')
sns.despine()
plt.show()
results.to_csv('submission.csv', index=False)
display(results.head())
```
| github_jupyter |
## benchmarking with the speedglm implementation
revised 03 August 2017, by [Nima Hejazi](http://nimahejazi.org)
The purpose of this notebook is to benchmark the performance of the `survtmle` package using the standard `glm` implementation, on simulated data sets of varying sample sizes ($n = 100, 1000, 5000$). This is one of two notebooks meant to compare the performance of `glm` against that of `speedglm`.
### preliminaries
```
# preliminaries
library(microbenchmark)
# set seed and constants
set.seed(341796)
t_0 <- 15
## get correct version of `survtmle`
if ("survtmle" %in% installed.packages()) {
remove.packages("survtmle")
}
suppressMessages(devtools::install_github("benkeser/survtmle", ref = "speedglm"))
library(survtmle)
```
## <u>Example 1: simple simulated data set</u>
This is a rather trivial example wherein the simulated data set contains few covariates.
### case 1: _n = 100 (trivial example)_
```
# simulate data
n <- 100
W <- data.frame(W1 = runif(n), W2 = rbinom(n, 1, 0.5))
A <- rbinom(n, 1, 0.5)
T <- rgeom(n,plogis(-4 + W$W1 * W$W2 - A)) + 1
C <- rgeom(n, plogis(-6 + W$W1)) + 1
ftime <- pmin(T, C)
ftype <- as.numeric(ftime == T)
system.time(
fit <- survtmle(ftime = ftime, ftype = ftype, trt = A, adjustVars = W,
glm.trt = "1", glm.ftime = "I(W1*W2) + trt + t",
glm.ctime = "W1 + t", method = "hazard", t0 = t_0)
)
suppressMessages(
m1 <- microbenchmark(unit = "s",
fit <- survtmle(ftime = ftime, ftype = ftype, trt = A, adjustVars = W,
glm.trt = "1", glm.ftime = "I(W1*W2) + trt + t",
glm.ctime = "W1 + t", method = "hazard", t0 = t_0)
)
)
summary(m1)
```
This trivial example is merely provided for comparison against the following cases with larger sample sizes.
### case 2: _n = 1000_
```
# simulate data
n <- 1000
W <- data.frame(W1 = runif(n), W2 = rbinom(n, 1, 0.5))
A <- rbinom(n, 1, 0.5)
T <- rgeom(n,plogis(-4 + W$W1 * W$W2 - A)) + 1
C <- rgeom(n, plogis(-6 + W$W1)) + 1
ftime <- pmin(T, C)
ftype <- as.numeric(ftime == T)
system.time(
fit <- survtmle(ftime = ftime, ftype = ftype, trt = A, adjustVars = W,
glm.trt = "1", glm.ftime = "I(W1*W2) + trt + t",
glm.ctime = "W1 + t", method = "hazard", t0 = t_0)
)
suppressMessages(
m2 <- microbenchmark(unit = "s",
fit <- survtmle(ftime = ftime, ftype = ftype, trt = A, adjustVars = W,
glm.trt = "1", glm.ftime = "I(W1*W2) + trt + t",
glm.ctime = "W1 + t", method = "hazard", t0 = t_0)
)
)
summary(m2)
```
...
### case 3: _n = 5000_
```
# simulate data
n <- 5000
W <- data.frame(W1 = runif(n), W2 = rbinom(n, 1, 0.5))
A <- rbinom(n, 1, 0.5)
T <- rgeom(n,plogis(-4 + W$W1 * W$W2 - A)) + 1
C <- rgeom(n, plogis(-6 + W$W1)) + 1
ftime <- pmin(T, C)
ftype <- as.numeric(ftime == T)
system.time(
fit <- survtmle(ftime = ftime, ftype = ftype, trt = A, adjustVars = W,
glm.trt = "1", glm.ftime = "I(W1*W2) + trt + t",
glm.ctime = "W1 + t", method = "hazard", t0 = t_0)
)
#m3 <- microbenchmark(unit = "s",
# fit <- survtmle(ftime = ftime, ftype = ftype, trt = A, adjustVars = W,
# glm.trt = "1", glm.ftime = "I(W1*W2) + trt + t",
# glm.ctime = "W1 + t", method = "hazard", t0 = t_0)
#)
#summary(m3)
```
This case takes too long to benchmark properly.
## <u>Example 2: a "more real" simulated data set</u>
This is a more interesting example wherein the simulated data set contains a larger number of covariates, which might be interesting in real-world / practical applications.
```
# functions for this simulation
get.ftimeForm <- function(trt, site){
form <- "-1"
for(i in unique(trt)){
for(s in unique(site)){
form <- c(form,
paste0("I(trt==",i,"& site == ",s," & t==",
unique(ftime[ftype>0 & trt==i & site == s]),")",
collapse="+"))
}
}
return(paste(form,collapse="+"))
}
get.ctimeForm <- function(trt, site){
form <- "-1"
for(i in unique(trt)){
for(s in unique(site)){
form <- c(form,
paste0("I(trt==",i,"& site == ",s," & t==",
unique(ftime[ftype==0 & trt==i & site == s]),")",
collapse="+"))
}
}
return(paste(form,collapse="+"))
}
```
### case 1: _n = 100 (trivial example)_
```
# simulate data
n <- 100
trt <- rbinom(n, 1, 0.5)
# e.g., study site
adjustVars <- data.frame(site = (rbinom(n,1,0.5) + 1))
ftime <- round(1 + runif(n, 1, 350) - trt + adjustVars$site)
ftype <- round(runif(n, 0, 1))
glm.ftime <- get.ftimeForm(trt = trt, site = adjustVars$site)
glm.ctime <- get.ctimeForm(trt = trt, site = adjustVars$site)
system.time(
fit <- survtmle(ftime = ftime, ftype = ftype, trt = trt, adjustVars = adjustVars,
glm.trt = "1", glm.ftime = glm.ftime, glm.ctime = glm.ctime,
method = "hazard", t0 = t_0)
)
suppressMessages(
m4 <- microbenchmark(unit = "s", times = 10,
fit <- survtmle(ftime = ftime, ftype = ftype, trt = trt, adjustVars = adjustVars,
glm.trt = "1", glm.ftime = glm.ftime, glm.ctime = glm.ctime,
method = "hazard", t0 = t_0)
)
)
summary(m4)
```
This trivial example is merely provided for comparison against the following cases with larger sample sizes.
### case 2: _n = 1000_
```
# simulate data
n <- 1000
trt <- rbinom(n, 1, 0.5)
# e.g., study site
adjustVars <- data.frame(site = (rbinom(n,1,0.5) + 1))
ftime <- round(1 + runif(n, 1, 350) - trt + adjustVars$site)
ftype <- round(runif(n, 0, 1))
glm.ftime <- get.ftimeForm(trt = trt, site = adjustVars$site)
glm.ctime <- get.ctimeForm(trt = trt, site = adjustVars$site)
system.time(
fit <- survtmle(ftime = ftime, ftype = ftype, trt = trt, adjustVars = adjustVars,
glm.trt = "1", glm.ftime = glm.ftime, glm.ctime = glm.ctime,
method = "hazard", t0 = t_0)
)
suppressMessages(
m5 <- microbenchmark(unit = "s", times = 10,
fit <- survtmle(ftime = ftime, ftype = ftype, trt = trt, adjustVars = adjustVars,
glm.trt = "1", glm.ftime = glm.ftime, glm.ctime = glm.ctime,
method = "hazard", t0 = t_0)
)
)
summary(m5)
```
commentary here...
### case 3: _n = 5000_
```
# simulate data
#n <- 5000
#trt <- rbinom(n, 1, 0.5)
# e.g., study site
#adjustVars <- data.frame(site = (rbinom(n,1,0.5) + 1))
#ftime <- round(1 + runif(n, 1, 350) - trt + adjustVars$site)
#ftype <- round(runif(n, 0, 1))
#glm.ftime <- get.ftimeForm(trt = trt, site = adjustVars$site)
#glm.ctime <- get.ctimeForm(trt = trt, site = adjustVars$site)
#system.time(
# fit <- survtmle(ftime = ftime, ftype = ftype, trt = trt, adjustVars = adjustVars,
# glm.trt = "1", glm.ftime = glm.ftime, glm.ctime = glm.ctime,
# method = "hazard", t0 = t_0)
#)
#m6 <- microbenchmark(unit = "s",
# fit <- survtmle(ftime = ftime, ftype = ftype, trt = trt, adjustVars = adjustVars,
# glm.trt = "1", glm.ftime = glm.ftime, glm.ctime = glm.ctime,
# method = "hazard", t0 = t_0)
#)
#summary(m6)
```
As in the previous example, this case takes too long to benchmark properly.
```
fit <- survtmle(ftime = ftime, ftype = ftype, trt = trt, adjustVars = adjustVars,
glm.trt = "1", glm.ftime = glm.ftime, glm.ctime = glm.ctime,
method = "hazard", t0 = t_0)
traceback()
```
| github_jupyter |
# Plagiarism Detection Model
Now that you've created training and test data, you are ready to define and train a model. Your goal in this notebook, will be to train a binary classification model that learns to label an answer file as either plagiarized or not, based on the features you provide the model.
This task will be broken down into a few discrete steps:
* Upload your data to S3.
* Define a binary classification model and a training script.
* Train your model and deploy it.
* Evaluate your deployed classifier and answer some questions about your approach.
To complete this notebook, you'll have to complete all given exercises and answer all the questions in this notebook.
> All your tasks will be clearly labeled **EXERCISE** and questions as **QUESTION**.
It will be up to you to explore different classification models and decide on a model that gives you the best performance for this dataset.
---
## Load Data to S3
In the last notebook, you should have created two files: a `training.csv` and `test.csv` file with the features and class labels for the given corpus of plagiarized/non-plagiarized text data.
>The below cells load in some AWS SageMaker libraries and creates a default bucket. After creating this bucket, you can upload your locally stored data to S3.
Save your train and test `.csv` feature files, locally. To do this you can run the second notebook "2_Plagiarism_Feature_Engineering" in SageMaker or you can manually upload your files to this notebook using the upload icon in Jupyter Lab. Then you can upload local files to S3 by using `sagemaker_session.upload_data` and pointing directly to where the training data is saved.
```
import pandas as pd
import boto3
import sagemaker
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
# session and role
sagemaker_session = sagemaker.Session()
role = sagemaker.get_execution_role()
# create an S3 bucket
bucket = sagemaker_session.default_bucket()
```
## EXERCISE: Upload your training data to S3
Specify the `data_dir` where you've saved your `train.csv` file. Decide on a descriptive `prefix` that defines where your data will be uploaded in the default S3 bucket. Finally, create a pointer to your training data by calling `sagemaker_session.upload_data` and passing in the required parameters. It may help to look at the [Session documentation](https://sagemaker.readthedocs.io/en/stable/session.html#sagemaker.session.Session.upload_data) or previous SageMaker code examples.
You are expected to upload your entire directory. Later, the training script will only access the `train.csv` file.
```
# should be the name of directory you created to save your features data
data_dir = 'plagiarism_data'
# set prefix, a descriptive name for a directory
prefix = 'data_plagiarism'
# upload all data to S3
input_data = sagemaker_session.upload_data(path=data_dir, bucket=bucket, key_prefix=prefix)
print(input_data)
```
### Test cell
Test that your data has been successfully uploaded. The below cell prints out the items in your S3 bucket and will throw an error if it is empty. You should see the contents of your `data_dir` and perhaps some checkpoints. If you see any other files listed, then you may have some old model files that you can delete via the S3 console (though, additional files shouldn't affect the performance of model developed in this notebook).
```
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
# confirm that data is in S3 bucket
empty_check = []
for obj in boto3.resource('s3').Bucket(bucket).objects.all():
empty_check.append(obj.key)
print(obj.key)
assert len(empty_check) !=0, 'S3 bucket is empty.'
print('Test passed!')
```
---
# Modeling
Now that you've uploaded your training data, it's time to define and train a model!
The type of model you create is up to you. For a binary classification task, you can choose to go one of three routes:
* Use a built-in classification algorithm, like LinearLearner.
* Define a custom Scikit-learn classifier, a comparison of models can be found [here](https://scikit-learn.org/stable/auto_examples/classification/plot_classifier_comparison.html).
* Define a custom PyTorch neural network classifier.
It will be up to you to test out a variety of models and choose the best one. Your project will be graded on the accuracy of your final model.
---
## EXERCISE: Complete a training script
To implement a custom classifier, you'll need to complete a `train.py` script. You've been given the folders `source_sklearn` and `source_pytorch` which hold starting code for a custom Scikit-learn model and a PyTorch model, respectively. Each directory has a `train.py` training script. To complete this project **you only need to complete one of these scripts**; the script that is responsible for training your final model.
A typical training script:
* Loads training data from a specified directory
* Parses any training & model hyperparameters (ex. nodes in a neural network, training epochs, etc.)
* Instantiates a model of your design, with any specified hyperparams
* Trains that model
* Finally, saves the model so that it can be hosted/deployed, later
### Defining and training a model
Much of the training script code is provided for you. Almost all of your work will be done in the `if __name__ == '__main__':` section. To complete a `train.py` file, you will:
1. Import any extra libraries you need
2. Define any additional model training hyperparameters using `parser.add_argument`
2. Define a model in the `if __name__ == '__main__':` section
3. Train the model in that same section
Below, you can use `!pygmentize` to display an existing `train.py` file. Read through the code; all of your tasks are marked with `TODO` comments.
**Note: If you choose to create a custom PyTorch model, you will be responsible for defining the model in the `model.py` file,** and a `predict.py` file is provided. If you choose to use Scikit-learn, you only need a `train.py` file; you may import a classifier from the `sklearn` library.
```
# directory can be changed to: source_sklearn or source_pytorch
!pygmentize source_sklearn/train.py
```
### Provided code
If you read the code above, you can see that the starter code includes a few things:
* Model loading (`model_fn`) and saving code
* Getting SageMaker's default hyperparameters
* Loading the training data by name, `train.csv` and extracting the features and labels, `train_x`, and `train_y`
If you'd like to read more about model saving with [joblib for sklearn](https://scikit-learn.org/stable/modules/model_persistence.html) or with [torch.save](https://pytorch.org/tutorials/beginner/saving_loading_models.html), click on the provided links.
---
# Create an Estimator
When a custom model is constructed in SageMaker, an entry point must be specified. This is the Python file which will be executed when the model is trained; the `train.py` function you specified above. To run a custom training script in SageMaker, construct an estimator, and fill in the appropriate constructor arguments:
* **entry_point**: The path to the Python script SageMaker runs for training and prediction.
* **source_dir**: The path to the training script directory `source_sklearn` OR `source_pytorch`.
* **entry_point**: The path to the Python script SageMaker runs for training and prediction.
* **source_dir**: The path to the training script directory `train_sklearn` OR `train_pytorch`.
* **entry_point**: The path to the Python script SageMaker runs for training.
* **source_dir**: The path to the training script directory `train_sklearn` OR `train_pytorch`.
* **role**: Role ARN, which was specified, above.
* **train_instance_count**: The number of training instances (should be left at 1).
* **train_instance_type**: The type of SageMaker instance for training. Note: Because Scikit-learn does not natively support GPU training, Sagemaker Scikit-learn does not currently support training on GPU instance types.
* **sagemaker_session**: The session used to train on Sagemaker.
* **hyperparameters** (optional): A dictionary `{'name':value, ..}` passed to the train function as hyperparameters.
Note: For a PyTorch model, there is another optional argument **framework_version**, which you can set to the latest version of PyTorch, `1.0`.
## EXERCISE: Define a Scikit-learn or PyTorch estimator
To import your desired estimator, use one of the following lines:
```
from sagemaker.sklearn.estimator import SKLearn
```
```
from sagemaker.pytorch import PyTorch
```
```
# your import and estimator code, here
from sagemaker.sklearn.estimator import SKLearn
# specify an output path
# prefix is specified above
output_path = 's3://{}/{}'.format(bucket, prefix)
# instantiate a pytorch estimator
estimator = SKLearn(entry_point='train.py',
source_dir='source_sklearn',
role=role,
train_instance_count=1,
train_instance_type='ml.c4.xlarge',
output_path=output_path,
sagemaker_session=sagemaker_session,
hyperparameters={
'epochs': 200
})
```
## EXERCISE: Train the estimator
Train your estimator on the training data stored in S3. This should create a training job that you can monitor in your SageMaker console.
```
%%time
# Train your estimator on S3 training data
estimator.fit({'train': input_data})
```
## EXERCISE: Deploy the trained model
After training, deploy your model to create a `predictor`. If you're using a PyTorch model, you'll need to create a trained `PyTorchModel` that accepts the trained `<model>.model_data` as an input parameter and points to the provided `source_pytorch/predict.py` file as an entry point.
To deploy a trained model, you'll use `<model>.deploy`, which takes in two arguments:
* **initial_instance_count**: The number of deployed instances (1).
* **instance_type**: The type of SageMaker instance for deployment.
Note: If you run into an instance error, it may be because you chose the wrong training or deployment instance_type. It may help to refer to your previous exercise code to see which types of instances we used.
```
%%time
# uncomment, if needed
# from sagemaker.pytorch import PyTorchModel
# deploy your model to create a predictor
predictor = estimator.deploy(initial_instance_count=1, instance_type='ml.t2.medium')
```
---
# Evaluating Your Model
Once your model is deployed, you can see how it performs when applied to our test data.
The provided cell below, reads in the test data, assuming it is stored locally in `data_dir` and named `test.csv`. The labels and features are extracted from the `.csv` file.
```
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
import os
# read in test data, assuming it is stored locally
test_data = pd.read_csv(os.path.join(data_dir, "test.csv"), header=None, names=None)
# labels are in the first column
test_y = test_data.iloc[:,0]
test_x = test_data.iloc[:,1:]
```
## EXERCISE: Determine the accuracy of your model
Use your deployed `predictor` to generate predicted, class labels for the test data. Compare those to the *true* labels, `test_y`, and calculate the accuracy as a value between 0 and 1.0 that indicates the fraction of test data that your model classified correctly. You may use [sklearn.metrics](https://scikit-learn.org/stable/modules/classes.html#module-sklearn.metrics) for this calculation.
**To pass this project, your model should get at least 90% test accuracy.**
```
# First: generate predicted, class labels
test_y_preds = predictor.predict(test_x)
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
# test that your model generates the correct number of labels
assert len(test_y_preds)==len(test_y), 'Unexpected number of predictions.'
print('Test passed!')
import sklearn.metrics as metrics
# Second: calculate the test accuracy
accuracy = metrics.accuracy_score(test_y, test_y_preds)
print(accuracy)
## print out the array of predicted and true labels, if you want
print('\nPredicted class labels: ')
print(test_y_preds)
print('\nTrue class labels: ')
print(test_y.values)
```
### Question 1: How many false positives and false negatives did your model produce, if any? And why do you think this is?
** Answer**: The model predicted 0 false positives and 0 false negatives which gives us a very accurate model (100%).
### Question 2: How did you decide on the type of model to use?
** Answer**: I decided to use a LinearSVC model based on https://scikit-learn.org/stable/tutorial/machine_learning_map/index.html
----
## EXERCISE: Clean up Resources
After you're done evaluating your model, **delete your model endpoint**. You can do this with a call to `.delete_endpoint()`. You need to show, in this notebook, that the endpoint was deleted. Any other resources, you may delete from the AWS console, and you will find more instructions on cleaning up all your resources, below.
```
# uncomment and fill in the line below!
#<name_of_deployed_predictor>.delete_endpoint()
# Deletes a precictor.endpoint
def delete_endpoint(predictor):
try:
boto3.client('sagemaker').delete_endpoint(EndpointName=predictor.endpoint)
print('Deleted {}'.format(predictor.endpoint))
except:
print('Already deleted: {}'.format(predictor.endpoint))
delete_endpoint(predictor)
```
### Deleting S3 bucket
When you are *completely* done with training and testing models, you can also delete your entire S3 bucket. If you do this before you are done training your model, you'll have to recreate your S3 bucket and upload your training data again.
```
# deleting bucket, uncomment lines below
bucket_to_delete = boto3.resource('s3').Bucket(bucket)
bucket_to_delete.objects.all().delete()
```
### Deleting all your models and instances
When you are _completely_ done with this project and do **not** ever want to revisit this notebook, you can choose to delete all of your SageMaker notebook instances and models by following [these instructions](https://docs.aws.amazon.com/sagemaker/latest/dg/ex1-cleanup.html). Before you delete this notebook instance, I recommend at least downloading a copy and saving it, locally.
---
## Further Directions
There are many ways to improve or add on to this project to expand your learning or make this more of a unique project for you. A few ideas are listed below:
* Train a classifier to predict the *category* (1-3) of plagiarism and not just plagiarized (1) or not (0).
* Utilize a different and larger dataset to see if this model can be extended to other types of plagiarism.
* Use language or character-level analysis to find different (and more) similarity features.
* Write a complete pipeline function that accepts a source text and submitted text file, and classifies the submitted text as plagiarized or not.
* Use API Gateway and a lambda function to deploy your model to a web application.
These are all just options for extending your work. If you've completed all the exercises in this notebook, you've completed a real-world application, and can proceed to submit your project. Great job!
| github_jupyter |
# XYZ, a Now Python-Friendly <br/>Geospatial Data Management Service
<p style="color:black;font-size:18px;">
<strong>Dinu Gherman, <a href="https://here.com/">HERE Technologies</a><br/>
<a href="http://2020.geopython.net">Online Python Machine Learning Conference & GeoPython 2020</a><br />Sept. 21, 2020</strong></p>
## Introduction
**HERE.com:** the location company, growing beyond automotive/logistics, 30+ years experience
**XYZ:** a new, unified, accessible, scalable, managed, open geo-database
**Components:** Hub, REST API, CLI, Studio (Web-Viewer)
**Features:** Open Source, Geospatial database/service, large volume, cloud, real-time, now Python.
**This talk:** about using XYZ programmatically from Python as intended for data scientists/analysts.
## Key Concepts
**XYZ:** Hub, Spaces with IDs, Features (geometry data) with IDs, Tags, Properties
**Spaces:** Create, read, list, update, share, delete. Plus info and stats
**Features:** Add, read, update, iterate, search, cluster (hex/quad bins), delete
**Search:** Features by ID, tag, property, bbox, tile, radius or geometry
**[XYZ-Spaces-Python](https://github.com/heremaps/xyz-spaces-python)**, short `xyzspaces`:
- wraps the XYZ Hub RESTful API
- provides a higher-level `Space` class
- imports GeoJSON, plus CSV, GeoBuff, GPX, KML, WKT, Shapefiles, and GeoPandas dataframes
**Pro plan (supported):** Virtual spaces, feature clustering, activity log (plus more to come)
## Mini-Tutorial
This section gives a short overview of basic interaction patterns with the HERE Data Hub (based on XYZ Hub, with some pro features added).
```
import json
import os
from IPython.display import JSON
import geojson
import pandas as pd
import requests
import turfpy.measurement
from branca.colormap import linear
from ipyleaflet import basemaps, GeoJSON, LayersControl, Map, Polygon
from ipywidgets import interact
import xyzspaces
import xyzspaces.datasets
from xyzspaces.apis import HubApi
from xyzspaces import XYZ
xyz_token = os.getenv("XYZ_TOKEN")
xyz = XYZ(credentials=xyz_token)
```
### List spaces
```
sp_list = xyz.spaces.list()
len(sp_list)
sp = sp_list[0]
JSON(sp)
```
### Create a space
`XYZ.spaces.new()`
```
data1 = xyzspaces.datasets.get_countries_data()
JSON(data1)
m = Map(center=[0, 0], zoom=1)
m.layout.height = "300px"
m += GeoJSON(data=data1)
m
world_space = xyz.spaces.new(title="PyML & GeoPy 2020 Demo", description="Demo")
world_space.info
sp_id = world_space.info["id"]
sp_id
world_space.get_statistics()
```
### Add features
`space.add_features()`
```
features_info = world_space.add_features(features=data1)
world_space.get_statistics()
```
### Access features
`space.iter_feature()`, `space.get_feature()`, `space.get_features()`
```
m = Map(center=[0, 0], zoom=1)
m.layout.height = "300px"
data2 = geojson.FeatureCollection(list(world_space.iter_feature()))
m += GeoJSON(data=data2)
m
JSON(data1["features"][0])
JSON(data2["features"][0])
```
`api.get_space_feature()`, `space.get_feature()`...
```
api = HubApi(credentials=xyz_token)
JSON(api.get_space_feature(sp_id, "AFG"))
```
Using more convenient abstractions:
```
afg = world_space.get_feature("AFG")
JSON(afg)
m = Map(center=[30, 70], zoom=3)
m.layout.height = "300px"
m += GeoJSON(data=afg)
m
```
Multiple features:
```
JSON(world_space.get_features(["AFG", "IND"]))
```
### Delete space
```
# world_space.delete()
```
### Update features, add tags
`space.update_feature`
```
for feat in world_space.iter_feature():
name = feat["properties"]["name"].lower()
if name[0] == name[-1]:
world_space.update_feature(feature_id=feat["id"],
data=feat,
add_tags=["palindromish"])
```
### Search features with tags
`space.search()`
```
for feat in world_space.search(tags=["palindromish"]):
print(feat["properties"]["name"])
```
### Search features in bounding box
`space.features_in_bbox()`
```
# Features in the Southern hemishpere:
m = Map(center=[-20, 0], zoom=1)
m.layout.height = "300px"
feats = list(world_space.features_in_bbox([-180, -90, 180, 0], clip=False))
m += GeoJSON(data=geojson.FeatureCollection(feats))
m
```
### Search features by geometry
`space.spatial_search_geometry()`
```
deu = world_space.get_feature("DEU")
del deu["geometry"]["bbox"] # hack
m = Map(center=[50, 9], zoom=4)
feats = list(world_space.spatial_search_geometry(data=deu["geometry"]))
m += GeoJSON(data=geojson.FeatureCollection(feats))
m += GeoJSON(data=deu, style={"color": "red", "fillOpacity": 0})
m
eu_feats = list(world_space.features_in_bbox([-20, 38, 35, 70]))
list(sorted([feat["properties"]["name"] for feat in eu_feats]))
df = pd.DataFrame(data=[{
"Name": eu_feat["properties"]["name"],
"Neighbors":
len(list(world_space.spatial_search_geometry(data=eu_feat["geometry"]))) - 1}
for eu_feat in eu_feats
]).sort_values(by=["Neighbors", "Name"], ascending=False, axis=0)
df.head()
df.index = df.Name
df.plot.bar(title="#Neighboring Countries in Europe", figsize=(10, 5), xlabel='')
```
### Search features by tags/parameters
`space.search()`
Now using the Microsoft US Building Footprints [dataset](https://github.com/Microsoft/USBuildingFootprints) (ca. 125 M buildings), also available in a HERE Data Hub space (ID: R4QDHvd1), used in this simple [ZIP code example](https://studio.here.com/viewer/?project_id=58604429-3919-437d-8ae4-9ee9693104d1) on [HERE Studio](https://studio.here.com).
```
from xyzspaces.datasets import get_microsoft_buildings_space as msbs
ms_space = msbs()
JSON(ms_space.get_statistics())
lat, lon = 38.89759, -77.03665 # White House, USA
m = Map(center=[lat, lon], zoom=18)
feats = list(ms_space.search(tags=["postalcode@20500"]))
# feats = list(space.spatial_search(lat=lat, lon=lon, radius=100)) # same
m += GeoJSON(data=geojson.FeatureCollection(feats))
m
JSON(feats[0])
```
Compared with original feature from https://github.com/Microsoft/USBuildingFootprints:
```
JSON({"type":"Feature","geometry":{"type":"Polygon","coordinates":[[[-77.037293,38.897578],[-77.036459,38.897577],[-77.036459,38.897592],[-77.035855,38.897591],[-77.035856,38.897546],[-77.035778,38.897545],[-77.035778,38.897399],[-77.03549,38.897399],[-77.035489,38.897735],[-77.03556,38.897735],[-77.03556,38.897771],[-77.035783,38.897771],[-77.035783,38.897706],[-77.036228,38.897707],[-77.036228,38.897817],[-77.036439,38.897818],[-77.036438,38.897928],[-77.036619,38.897929],[-77.036619,38.89781],[-77.036853,38.89781],[-77.036853,38.897708],[-77.03695,38.897708],[-77.03695,38.897708],[-77.037851,38.897709],[-77.037851,38.89747],[-77.037929,38.89747],[-77.037929,38.897427],[-77.037815,38.897427],[-77.037816,38.897339],[-77.037294,38.897338],[-77.037293,38.897578]]]},"properties":{}})
gen = ms_space.search(params={"p.city": "Washington"})
wash_dc_1000 = [next(gen) for i in range(1000)]
lat, lon = 38.89759, -77.03665 # White House, USA
m = Map(center=[lat, lon], zoom=11, basemap=basemaps.CartoDB.Positron)
m += GeoJSON(data=geojson.FeatureCollection(wash_dc_1000))
m
```
### Search features by radius
`space.spatial_search()`
```
lat, lon = 38.89759, -77.03665 # White House, USA
m = Map(center=[lat, lon], zoom=14, basemap=basemaps.CartoDB.Positron)
features = list(ms_space.spatial_search(lat=lat, lon=lon, radius=2000))
m += GeoJSON(data=geojson.FeatureCollection(features))
m
area = turfpy.measurement.area
sizes = [area(f) for f in features]
cm = linear.Oranges_04.scale(min(sizes), max(sizes))
cm
lat, lon = 38.89759, -77.03665 # White House, USA
m = Map(center=[lat, lon], zoom=14, basemap=basemaps.CartoDB.Positron)
features = list(ms_space.spatial_search(lat=lat, lon=lon, radius=2000))
m += GeoJSON(
data=geojson.FeatureCollection(features),
hover_style={"fillOpacity": 1},
style_callback=lambda feat: {"weight": 2, "color": cm(area(feat))}
)
m
```
See also a more expanded version using building house numbers in this notebook on GitHub: [docs/notebooks/building_numbers.ipynb](https://github.com/heremaps/xyz-spaces-python/blob/master/docs/notebooks/building_numbers.ipynb).
```
# ms_space.delete()
```
### Cluster all space features
`space.cluster()`
```
xyz_pro_token = os.getenv("XYZ_PRO_TOKEN")
xyz = XYZ(credentials=xyz_pro_token)
cluster_space = xyz.spaces.new(title="Cluster Demo GeoPy", description="...")
info = cluster_space.add_feature(afg)
lat, lon = list(reversed(turfpy.measurement.center(afg)["geometry"]["coordinates"]))
m = Map(center=[lat, lon], zoom=4, basemap=basemaps.CartoDB.Positron)
m += GeoJSON(data=afg, name="AFG", style={"color": "red"})
m
@interact(abs_res=(0, 4))
def overlay(abs_res=0):
global m
fc = cluster_space.cluster(
"hexbin",
clustering_params = {"absoluteResolution": abs_res}
)
lay = GeoJSON(data=fc, name=f"Hexbin {abs_res}")
try:
prev = [l for l in m.layers if l.name.startswith("Hexbin")][0]
m.substitute_layer(prev, lay)
except IndexError:
m += lay
cluster_space.delete_feature("AFG")
```
`space.add_features_csv()`
```
# https://ourairports.com/data/
# https://ourairports.com/data/airports.csv
url = "https://ourairports.com/data/airports.csv"
fn = os.path.basename(url)
try:
df = pd.read_csv(fn)
except:
df = pd.read_csv(url)
df.to_csv(fn, index=False)
df1 = df[df.continent=="EU"]
df1 = df1[df1.iso_country!="RU"]
fn = "airports_eu.csv"
df1.to_csv(fn, index=False)
m = Map(center=[50, 13], zoom=3, basemap=basemaps.CartoDB.Positron)
m += LayersControl(position="topright")
m
# add many single point XYZ feature from CSV file
info = cluster_space.add_features_csv(fn, "longitude_deg", "latitude_deg", id_col="id")
# calculate clustered cells
fcc = cluster_space.cluster("hexbin", clustering_params={"absoluteResolution": 2})
# add hex cluster cells to map
values = [f["properties"]["aggregation"]["qty"]
for f in fcc["features"]]
cm = linear.Oranges_04.scale(min(values), max(values))
m += GeoJSON(data=fcc,
name="Hex Clusters",
hover_style={"fillOpacity": 0.75},
style_callback=lambda feat: {
"color": cm(feat["properties"]["aggregation"]["qty"])
})
# build one multi-point GeoJSON object
coords = [[tup.longitude_deg, tup.latitude_deg] for tup in df1.itertuples()]
mp = geojson.MultiPoint([[lon, lat] for [lon, lat] in coords])
f = geojson.Feature(geometry=mp)
f["id"] = "airports"
fc = geojson.FeatureCollection([f])
m += GeoJSON(data=fc,
name="Airports",
point_style={'radius': 1, "weight": 1, "fillOpacity": 1})
```
### Cleanup
```
cluster_space.delete()
world_space.delete()
```
More examples are available in the [docs/notebooks](github.com/heremaps/xyz-spaces-python) folder on GitHub.
## Not shown...
- Search by tiles
- Schema validation (pro)
- Virtual spaces (pro)
- Activity log (pro)
- Rule-based tags (pro)
## Conclusions
### Main take-aways
XYZ:
- is an Open Source geospatial cloud database/service
- is the foundation of the HERE Data Hub (commercial, free plan w/o CC)
- stores geodata features as GeoJSON, organized in *spaces*
- allows to manage, scale and filter geodata easily
- loves Python: `pip install xyzspaces`, also on conda-forge
- wants you to engage and give feedback!
### Links
- https://here.xyz
- https://github.com/heremaps/xyz-spaces-python
- https://developer.here.com/products/data-hub
### Questions?
| github_jupyter |
```
import numpy as np
import pandas as pd
from sklearn import preprocessing
def step_gradient(points,learning_rate,m):
M=len(points)
m_slope=list([0 for i in range(len(points[0])-1)])
for i in range(M):
y=points[i,len(points[0])-2]
x_total=0;
q=0
for k in range(len(points[0])):
if(k==len(points[0])-2):
continue
x_total+=m[q]*points[i,k]
q=q+1
l=0
for j in range(len(points[0])):
if (j==len(points[0])-2):
continue
m_slope[l]+=(-2/M)*((y-x_total)*points[i,j])
l=l+1
new_m=list([0 for j in range(len(points[0])-1)])
a=0
for i in range(len(m)):
new_m[i]=m[i]-learning_rate*m_slope[i]
return new_m
def gd(points,learning_rate,num_iter):
m=list([0 for j in range(len(points[0])-1)])
for i in range(num_iter):
m = step_gradient(points,learning_rate,m)
print(i," cost: ",cost(points,m))
return m
def run():
data = np.loadtxt('training_boston_x_y_train.csv',delimiter=',')
x = data[0:,0:13]
y = data[0:,13:]
#min_max_scaler_object = preprocessing.MinMaxScaler()
#x = min_max_scaler_object.fit_transform(x)
c = np.hstack((x,y))
o = np.ones((len(data),1))
c = np.hstack((c,o))
df = pd.DataFrame(c)
x2 = df.values
learning_rate = 0.1
num_iter =800
m = gd(x2,learning_rate,num_iter)
return m
def cost(points,m):
#total_cost=0
M=len(points)
total_cost=0
for i in range(M):
y=points[i,len(points[0])-2]
x_total=0
q=0
for k in range(len(points[0])):
if(k==len(points[0])-2):
continue
x_total+=m[q]*points[i,k]
q=q+1
total_cost+=(1/M)*((y-x_total)**2)
return total_cost
def predict(p,m):
y_pred=list([0 for j in range(len(p))])
k=0
for i in range(len(p)):
for j in range(len(m)):
y_pred[k]+=m[j]*p[i,j]
k=k+1
return y_pred
m = run()
m
p=np.genfromtxt("test_boston_x_test.csv",delimiter=",")
b=np.ones((len(p),1))
c=np.hstack((p,b))
y_pred=predict(c,m)
x=np.array(y_pred)
x=np.row_stack(x)
np.savetxt("y_test.csv",x,fmt="%.5f")
data = np.genfromtxt('training_boston_x_y_train.csv',delimiter=',')
df = pd.DataFrame(data)
data = np.loadtxt('training_boston_x_y_train.csv',delimiter=',')
x = data[0:,0:13]
y = data[0:,13:]
b=np.ones((len(data),1))
c=np.hstack((x,b))
y_pred=predict(c,m)
y_pred = pd.DataFrame(y_pred)
y_pred.shape
y.shape
out = np.hstack((y_pred,y))
out = pd.DataFrame(out)
out
```
| github_jupyter |
# Tutorial: Learning a digit classifier with the MNIST dataset
## Introduction
The goal of this tutorial is to learn basic machine learning skills. The goal is to make the best digit classifier you can.
The dataset we will work on is old but it is a reference benchmark to evaluate new algorithms (and early concepts).
How well our classification algorithms performs ?
The MNIST handwritten digit database is a collection of 70,000 handwritten digits and their corresponding labels (from 0 to 9).
The dataset is split into a Training set (60,000 images) and a Validation set (10,000 images). You will train your model on the Training set and test it on the Test set.
__[Who is the best at MNIST?](https://rodrigob.github.io/are_we_there_yet/build/classification_datasets_results.html#4d4e495354)__
## Requirements
- We will need [Scikit-Learn](https://scikit-learn.org/stable/) and [Keras](https://keras.io/) + [TensorFlow](https://www.tensorflow.org/). The MNIST dataset is downloaded by Tensorflow/Keras.
- __DO__ read the [Scikit-Learn documentation](https://scikit-learn.org/stable/tutorial/basic/tutorial.html) which is exhaustive and completely awesome.
- Scikit_Learn is from [INRIA](https://scikit-learn.fondation-inria.fr/en/home/), Keras/TF from Google.
```
from keras.datasets import mnist
(x_train, y_train), (x_test, y_test) = mnist.load_data()
import matplotlib
import matplotlib.pyplot as plt
%matplotlib inline
# %matplotlib nbagg
# %matplotlib ipympl
# %matplotlib notebook
images_and_labels = list(zip(x_train, y_train))
plt.subplots_adjust(bottom=0, left=.01, right=.99, top=1.6, hspace=.35)
for i, (image, label) in enumerate(images_and_labels[:25]):
plt.subplot(5, 5, i + 1)
plt.axis('off')
plt.imshow(image, cmap=plt.cm.gray, interpolation='nearest')
plt.title('N°%i Label: %i' % (i, label))
%%timeit -n 1 -r 1
# Explore the first 5 digits in the training dataset
for i, image in enumerate(x_train[0:5]):
print('Image n°', i, 'Label:', y_train[i])
plt.imshow(image, cmap='gray')
plt.show()
```
### Let's evaluate three classical Machine Learning methods
- Support Vector Machine (SVM) -> [1_ML_Tutorial_SVM.ipynb](1_ML_Tutorial_SVM.ipynb)
- Neural Networks -> [2_ML_Tutorial_NN.ipynb](2_ML_Tutorial_NN.ipynb)
- Convolutionnal Neural Networks (CNN) -> [3_ML_Tutorial_CNN.ipynb](3_ML_Tutorial_CNN.ipynb)
# Some Machine Learning ressources
- [Scikit-Learn Quick Start Tutorial](https://scikit-learn.org/stable/tutorial/basic/tutorial.html)
- [Jake VanderPlas' _Python Data Science Handbook_](https://jakevdp.github.io/PythonDataScienceHandbook/index.html)
- [Fast.ai ML MOOC](http://course18.fast.ai/ml)
| github_jupyter |
[Reference](https://tensorflow.google.cn/tutorials/wide)
These tutorial introdue how to implment feature column out of pandas dataframe
```
import tensorflow as tf
import pandas as pd
from pandas import DataFrame
import tempfile
```
## Explore Data
```
# import tempfile
# import urllib
# train_file = tempfile.NamedTemporaryFile()
# test_file = tempfile.NamedTemporaryFile()
# urllib.request.urlretrieve("https://archive.ics.uci.edu/ml/machine-learning-databases/adult/adult.data", train_file.name)
# urllib.request.urlretrieve("https://archive.ics.uci.edu/ml/machine-learning-databases/adult/adult.test", test_file.name)
import pandas as pd
train_file_name = "adult.data.txt"
test_file_name = "adult.test.txt"
CSV_COLUMNS = [
"age", "workclass", "fnlwgt", "education", "education_num",
"marital_status", "occupation", "relationship", "race", "gender",
"capital_gain", "capital_loss", "hours_per_week", "native_country",
"income_bracket"]
df_train = pd.read_csv("adult.data.txt", names=CSV_COLUMNS, skipinitialspace=True)
df_test = pd.read_csv("adult.test.txt", names=CSV_COLUMNS, skipinitialspace=True, skiprows=1)
df_test.head()
df_test.head()
```
Transform the label
```
train_label = df_train["income_bracket"].apply(lambda x: ">50K" in x).astype(int)
test_label = df_test["income_bracket"].apply(lambda x: ">50K" in x).astype(int)
def input_fn(data_file, num_epochs, shuffle):
"""Input builder function."""
df_data = pd.read_csv(
tf.gfile.Open(data_file),
names=CSV_COLUMNS,
skipinitialspace=True,
engine="python",
skiprows=1)
# remove NaN elements
df_data = df_data.dropna(how="any", axis=0)
labels = df_data["income_bracket"].apply(lambda x: ">50K" in x).astype(int)
return tf.estimator.inputs.pandas_input_fn(
x=df_data,
y=labels,
batch_size=100,
num_epochs=num_epochs,
shuffle=shuffle,
num_threads=5)
```
# Selecting and Engineering feature for the model
A feature coloum can be one of these
* base feature column: raw column in original dataframe
* derived feature column: new column created base on some transformation over one or many raw column in original dataframe.
#### Base categorical column
```
gender = tf.feature_column.categorical_column_with_vocabulary_list(
"gender", ["Female", "Male"])
# if we don't know possible value in advance
occupation = tf.feature_column.categorical_column_with_hash_bucket(
"occupation", hash_bucket_size=1000)
education = tf.feature_column.categorical_column_with_vocabulary_list(
"education", [
"Bachelors", "HS-grad", "11th", "Masters", "9th",
"Some-college", "Assoc-acdm", "Assoc-voc", "7th-8th",
"Doctorate", "Prof-school", "5th-6th", "10th", "1st-4th",
"Preschool", "12th"
])
marital_status = tf.feature_column.categorical_column_with_vocabulary_list(
"marital_status", [
"Married-civ-spouse", "Divorced", "Married-spouse-absent",
"Never-married", "Separated", "Married-AF-spouse", "Widowed"
])
relationship = tf.feature_column.categorical_column_with_vocabulary_list(
"relationship", [
"Husband", "Not-in-family", "Wife", "Own-child", "Unmarried",
"Other-relative"
])
workclass = tf.feature_column.categorical_column_with_vocabulary_list(
"workclass", [
"Self-emp-not-inc", "Private", "State-gov", "Federal-gov",
"Local-gov", "?", "Self-emp-inc", "Without-pay", "Never-worked"
])
native_country = tf.feature_column.categorical_column_with_hash_bucket(
"native_country", hash_bucket_size=1000)
```
#### Base continuous column
```
education_num = tf.feature_column.numeric_column("education_num")
age = tf.feature_column.numeric_column("age")
education_num = tf.feature_column.numeric_column("education_num")
capital_gain = tf.feature_column.numeric_column("capital_gain")
capital_loss = tf.feature_column.numeric_column("capital_loss")
hours_per_week = tf.feature_column.numeric_column("hours_per_week")
```
#### Making continuous feature categorical by bucketization
```
age = tf.feature_column.numeric_column("age")
age_buckets = tf.feature_column.bucketized_column(
age, boundaries=[18, 25, 30, 35, 40, 45, 50, 55, 60, 65])
```
### Intersecting multiple column with CrossedColumn
```
education_x_occupation = tf.feature_column.crossed_column(
["education", "occupation"], hash_bucket_size=1000)
```
# Define Model
```
# first list all the feature column we defined
base_columns = [
gender, native_country, education, occupation, workclass, relationship,
age_buckets,
]
crossed_columns = [
tf.feature_column.crossed_column(
["education", "occupation"], hash_bucket_size=1000),
tf.feature_column.crossed_column(
[age_buckets, "education", "occupation"], hash_bucket_size=1000),
tf.feature_column.crossed_column(
["native_country", "occupation"], hash_bucket_size=1000)
]
model_dir = tempfile.mkdtemp()
m = tf.estimator.LinearClassifier(
model_dir=model_dir, feature_columns=base_columns + crossed_columns)
model_dir = tempfile.mkdtemp()
m = tf.estimator.LinearClassifier(
model_dir=model_dir, feature_columns=base_columns + crossed_columns)
```
# Train Model
```
# set num_epochs to None to get infinite stream of data.
train_steps = 100
m.train(
input_fn=input_fn(train_file_name, num_epochs=None, shuffle=True),
steps=train_steps)
results = m.evaluate(input_fn=input_fn(test_file_name,num_epochs=1, shuffle=False))
print("model directory = %s" % model_dir)
for key in sorted(results):
print("%s: %s" % (key, results[key]))
```
| github_jupyter |
<img src="NotebookAddons/blackboard-banner.png" width="100%" />
<font face="Calibri">
<br>
<font size="5"> <b>Exploring SAR Time Series Data over Ecosystems and Deforestation Sites</b></font>
<br>
<font size="4"> <b> Franz J Meyer; University of Alaska Fairbanks & Josef Kellndorfer, <a href="http://earthbigdata.com/" target="_blank">Earth Big Data, LLC</a> </b> <br>
<img style="padding:7px;" src="NotebookAddons/UAFLogo_A_647.png" width="170" align="right" /></font>
<font size="3">This notebook introduces you to the time series signatures over forested sites and sites affected by deforestation. The data analysis is done in the framework of *Jupyter Notebooks*. The Jupyter Notebook environment is easy to launch in any web browser for interactive data exploration with provided or new training data. Notebooks are comprised of text written in a combination of executable python code and markdown formatting including latex style mathematical equations. Another advantage of Jupyter Notebooks is that they can easily be expanded, changed, and shared with new data sets or newly available time series steps. Therefore, they provide an excellent basis for collaborative and repeatable data analysis. <br>
<b>This notebook covers the following data analysis concepts:</b>
- How to load time series stacks into Jupyter Notebooks and how to explore image content using basic functions such as mean value calculation and histogram analysis.
- How to extract time series information for individual pixels of an image.
- Typical time series signatures over forests and deforestation sites.
</font>
</font>
<hr>
<font face="Calibri" size="5" color='rgba(200,0,0,0.2)'> <b>Important Notes about JupyterHub</b> </font>
<br><br>
<font face="Calibri" size="3"> <b>Your JupyterHub server will automatically shutdown when left idle for more than 1 hour. Your notebooks will not be lost but you will have to restart their kernels and re-run them from the beginning. You will not be able to seamlessly continue running a partially run notebook.</b> </font>
<br><br>
</font>
```
%%javascript
var kernel = Jupyter.notebook.kernel;
var command = ["notebookUrl = ",
"'", window.location, "'" ].join('')
kernel.execute(command)
from IPython.display import Markdown
from IPython.display import display
user = !echo $JUPYTERHUB_USER
env = !echo $CONDA_PREFIX
if env[0] == '':
env[0] = 'Python 3 (base)'
if env[0] != '/home/jovyan/.local/envs/rtc_analysis':
display(Markdown(f'<text style=color:red><strong>WARNING:</strong></text>'))
display(Markdown(f'<text style=color:red>This notebook should be run using the "rtc_analysis" conda environment.</text>'))
display(Markdown(f'<text style=color:red>It is currently using the "{env[0].split("/")[-1]}" environment.</text>'))
display(Markdown(f'<text style=color:red>Select the "rtc_analysis" from the "Change Kernel" submenu of the "Kernel" menu.</text>'))
display(Markdown(f'<text style=color:red>If the "rtc_analysis" environment is not present, use <a href="{notebookUrl.split("/user")[0]}/user/{user[0]}/notebooks/conda_environments/Create_OSL_Conda_Environments.ipynb"> Create_OSL_Conda_Environments.ipynb </a> to create it.</text>'))
display(Markdown(f'<text style=color:red>Note that you must restart your server after creating a new environment before it is usable by notebooks.</text>'))
```
<hr>
<font face="Calibri">
<font size="5"> <b> 0. Importing Relevant Python Packages </b> </font>
<font size="3">In this notebook we will use the following scientific libraries:
<ol type="1">
<li> <b><a href="https://pandas.pydata.org/" target="_blank">Pandas</a></b> is a Python library that provides high-level data structures and a vast variety of tools for analysis. The great feature of this package is the ability to translate rather complex operations with data into one or two commands. Pandas contains many built-in methods for filtering and combining data, as well as the time-series functionality. </li>
<li> <b><a href="https://www.gdal.org/" target="_blank">GDAL</a></b> is a software library for reading and writing raster and vector geospatial data formats. It includes a collection of programs tailored for geospatial data processing. Most modern GIS systems (such as ArcGIS or QGIS) use GDAL in the background.</li>
<li> <b><a href="http://www.numpy.org/" target="_blank">NumPy</a></b> is one of the principal packages for scientific applications of Python. It is intended for processing large multidimensional arrays and matrices, and an extensive collection of high-level mathematical functions and implemented methods makes it possible to perform various operations with these objects. </li>
<li> <b><a href="https://matplotlib.org/index.html" target="_blank">Matplotlib</a></b> is a low-level library for creating two-dimensional diagrams and graphs. With its help, you can build diverse charts, from histograms and scatterplots to non-Cartesian coordinates graphs. Moreover, many popular plotting libraries are designed to work in conjunction with matplotlib. </li>
</font>
```
%%capture
import os # for chdir, getcwd, path.basename, path.exists
from math import ceil
import pandas as pd # for DatetimeIndex
import numpy as np #for log10, mean, percentile, power
from osgeo import gdal # for GetRasterBand, Open, ReadAsArray
%matplotlib notebook
import matplotlib.patches as patches # for Rectangle
import matplotlib.pyplot as plt # for add_subplot, axis, figure, imshow, legend, plot, set_axis_off, set_data,
# set_title, set_xlabel, set_ylabel, set_ylim, subplots, title, twinx
plt.rcParams.update({'font.size': 12})
import asf_notebook as asfn
asfn.jupytertheme_matplotlib_format()
```
<hr>
<font face="Calibri">
<font size="5"> <b> 1. Load Data Stack</b> </font> <img src="NotebookAddons/Deforest-MadreDeDios.jpg" width="350" style="padding:5px;" align="right" />
<font size="3"> This notebook will be using a 78-image deep dual-polarization C-band SAR data stack over Madre de Dios in Peru to analyze time series signatures of vegetation covers, water bodies, and areas affected by deforestation. The C-band data were acquired by ESA's Sentinel-1 SAR sensor constellation and are available to you through the services of the <a href="https://www.asf.alaska.edu/" target="_blank">Alaska Satellite Facility</a>.
The site in question is interesting as it has experienced extensive logging over the last 10 years (see image to the right; <a href="https://blog.globalforestwatch.org/" target="_blank">Monitoring of the Andean Amazon Project</a>). Since the 1980s, people have been clearing forests in this area for farming, cattle ranching, logging, and (recently) gold mining. Creating RGB color composites is an easy way to visualize ongoing changes in the landscape.
</font></font>
<br><br>
<font face="Calibri" size="3">Before we get started, let's first <b>create a working directory for this analysis and change into it:</b> </font>
```
path = "/home/jovyan/notebooks/SAR_Training/English/Hazards/data_Ex2-4_S1-MadreDeDios"
asfn.new_directory(path)
os.chdir(path)
print(f"Current working directory: {os.getcwd()}")
```
<font face="Calibri" size="3">We will <b>retrieve the relevant data</b> from an <a href="https://aws.amazon.com/" target="_blank">Amazon Web Service (AWS)</a> cloud storage bucket <b>using the following command</b>:</font></font>
```
time_series_path = 's3://asf-jupyter-data/MadreDeDios.zip'
time_series = os.path.basename(time_series_path)
!aws --region=us-east-1 --no-sign-request s3 cp $time_series_path $time_series
```
<font face="Calibri" size="3"> Now, let's <b>unzip the file (overwriting previous extractions) and clean up after ourselves:</b> </font>
```
if asfn.path_exists(time_series):
asfn.asf_unzip(os.getcwd(), time_series)
os.remove(time_series)
```
<br>
<font face="Calibri" size="5"> <b> 2. Define Data Directory and Path to VRT </b> </font>
<br><br>
<font face="Calibri" size="3"><b>Create a variable containing the VRT filename and the image acquisition dates:</b></font>
```
!gdalbuildvrt -separate raster_stack.vrt tiffs/*_VV.tiff
image_file_VV = "raster_stack.vrt"
!gdalbuildvrt -separate raster_stack_VH.vrt tiffs/*_VH.tiff
image_file_VH = "raster_stack_VH.vrt"
```
<font face="Calibri" size="3"><b>Create an index of timedelta64 data with Pandas:</b></font>
```
!ls tiffs/*_VV.tiff | sort | cut -c 7-21 > raster_stack_VV.dates
datefile_VV = 'raster_stack_VV.dates'
dates_VV = open(datefile_VV).readlines()
tindex_VV = pd.DatetimeIndex(dates_VV)
!ls tiffs/*_VH.tiff | sort | cut -c 7-21 > raster_stack_VH.dates
datefile_VH = 'raster_stack_VH.dates'
dates_VH = open(datefile_VH).readlines()
tindex_VH = pd.DatetimeIndex(dates_VH)
```
<br>
<hr>
<font face="Calibri" size="5"> <b> 3. Assess Image Acquisition Dates </b> </font>
<font face="Calibri" size="3"> Before we start analyzing the available image data, we want to examine the content of our data stack. From the date index, we <b>make and print a lookup table for band numbers and dates:</b></font>
```
stindex=[]
for i in [datefile_VV,datefile_VH]:
sdates=open(i).readlines()
stindex.append(pd.DatetimeIndex(sdates))
j=1
print('\nBands and dates for',i.strip('.dates'))
for k in stindex[-1]:
print("{:4d} {}".format(j, k.date()),end=' ')
j+=1
if j%5==1: print()
```
<hr>
<br>
<font face="Calibri" size="5"> <b> 4. Create Minimum Image to Identify Likely Areas of Deforestation </b> </font>
<font face="Calibri" size="4"> <b> 4.1 Load Time Series Stack </b> </font>
<b>First, we load the raster stack into memory and calculate the minimum backscatter in the time series:</b>
</font>
```
img = gdal.Open(image_file_VV)
band = img.GetRasterBand(1)
raster0 = band.ReadAsArray()
band_number = 0 # Needed for updates
rasterstack_VV = img.ReadAsArray()
```
<font face="Calibri" size="3"> To <b>explore the image (number of bands, pixels, lines),</b> you can use several functions associated with the image object (img) created in the last code cell: </font>
```
print(img.RasterCount) # Number of Bands
print(img.RasterXSize) # Number of Pixels
print(img.RasterYSize) # Number of Lines
```
<font face="Calibri" size="3"> The following line <b>calculates the minimum backscatter per pixel</b> across the time series: </font>
```
db_mean = np.min(rasterstack_VV, axis=0)
```
<br>
<font face="Calibri" size="4"> <b> 4.2 Visualize the Minimum Image and Select a Coordinate for a Time Series</b> </font>
<font face="Calibri" size="3"> <b>Write a class to create an interactive plot from which we can select interesting image locations for a time series.</b></font>
```
class pixelPicker:
def __init__(self, image, width, height):
self.x = None
self.y = None
self.fig = plt.figure(figsize=(width, height))
self.ax = self.fig.add_subplot(111, visible=False)
self.rect = patches.Rectangle(
(0.0, 0.0), width, height,
fill=False, clip_on=False, visible=False)
self.rect_patch = self.ax.add_patch(self.rect)
self.cid = self.rect_patch.figure.canvas.mpl_connect('button_press_event',
self)
self.image = image
self.plot = self.gray_plot(self.image, fig=self.fig, return_ax=True)
self.plot.set_title('Select a Point of Interest')
def gray_plot(self, image, vmin=None, vmax=None, fig=None, return_ax=False):
'''
Plots an image in grayscale.
Parameters:
- image: 2D array of raster values
- vmin: Minimum value for colormap
- vmax: Maximum value for colormap
- return_ax: Option to return plot axis
'''
if vmin is None:
vmin = np.nanpercentile(self.image, 1)
if vmax is None:
vmax = np.nanpercentile(self.image, 99)
#if fig is None:
# my_fig = plt.figure()
ax = fig.add_axes([0.1,0.1,0.8,0.8])
ax.imshow(image, cmap=plt.cm.gist_gray, vmin=vmin, vmax=vmax)
if return_ax:
return(ax)
def __call__(self, event):
print('click', event)
self.x = event.xdata
self.y = event.ydata
for pnt in self.plot.get_lines():
pnt.remove()
plt.plot(self.x, self.y, 'ro')
```
<font face="Calibri" size="3"> Now we are ready to plot the minimum image. <b>Click a point interest for which you want to analyze radar brightness over time</b>: </font>
```
# Large plot of multi-temporal average of VV values to inspect pixel values
fig_xsize = 7.5
fig_ysize = 7.5
my_plot = pixelPicker(db_mean, fig_xsize, fig_ysize)
```
<font face="Calibri" size="3"><b>Save the selected coordinates</b>: </font>
```
sarloc = (ceil(my_plot.x), ceil(my_plot.y))
print(sarloc)
```
<br>
<font face="Calibri" size="5"> <b> 5. Plot SAR Brightness Time Series at Point Locations </b> </font>
<font face="Calibri" size="4"> <b> 5.1 SAR Brightness Time Series at Point Locations </b> </font>
<font face="Calibri" size="3"> We will pick a pixel location identified in the SAR image above and plot the time series for this identified point. By focusing on image locations undergoing deforestation, we should see the changes in the radar cross section related to the deforestation event.
First, for processing of the imagery in this notebook we <b>generate a list of image handles and retrieve projection and georeferencing information.</b></font>
```
imagelist=[image_file_VV, image_file_VH]
geotrans=[]
proj=[]
img_handle=[]
xsize=[]
ysize=[]
bands=[]
for i in imagelist:
img_handle.append(gdal.Open(i))
geotrans.append(img_handle[-1].GetGeoTransform())
proj.append(img_handle[-1].GetProjection())
xsize.append(img_handle[-1].RasterXSize)
ysize.append(img_handle[-1].RasterYSize)
bands.append(img_handle[-1].RasterCount)
# for i in proj:
# print(i)
# for i in geotrans:
# print(i)
# for i in zip(['C-VV','C-VH','NDVI','B3','B4','B5'],bands,ysize,xsize):
# print(i)
```
<font face="Calibri" size="3"> Now, let's <b>pick a 5x5 image area around a center pixel defined in variable <i>sarloc</i></b>...</font>
```
ref_x=geotrans[0][0]+sarloc[0]*geotrans[0][1]
ref_y=geotrans[0][3]+sarloc[1]*geotrans[0][5]
print('UTM Coordinates ',ref_x, ref_y)
print('SAR pixel/line ',sarloc[0], sarloc[1])
subset_sentinel=(sarloc[0], sarloc[1], 5, 5)
```
<font face="Calibri" size="3">... and <b>extract the time series</b> for this small area around the selected center pixel:</font>
```
s_ts=[]
for idx in (0, 1):
means=[]
for i in range(bands[idx]):
rs=img_handle[idx].GetRasterBand(i+1).ReadAsArray(*subset_sentinel)
rs_means_pwr = np.mean(rs)
rs_means_dB = 10.*np.log10(rs_means_pwr)
means.append(rs_means_dB)
s_ts.append(pd.Series(means,index=stindex[idx]))
means = []
```
<font face="Calibri" size="3"><b>Plot the extracted time series</b> for VV and VH polarizations:</font>
```
%matplotlib inline
fig, ax = plt.subplots(1, 1, figsize=(8, 4))
s_ts[0].plot(ax=ax, color='red', label='C-VV')#,xlim=(min(min(stindex),min(stindex[0])),
# max(max(stindex),max(stindex[0]))))
s_ts[1].plot(ax=ax, color='blue', label='C-VH')
ax.set_xlabel('Date')
ax.set_ylabel('Sentinel-1 $\gamma^o$ [dB]')
ax.set_title('Sentinel-1 Backscatter')
plt.grid()
_ = ax.legend(loc='best')
_ = fig.suptitle('Time Series Profiles of Sentinel-1 SAR Backscatter')
figname = f"RCSTimeSeries-{ref_x:.0f}_{ref_y:.0f}.png"
plt.savefig(figname, dpi=300, transparent='true')
```
<br>
<div class="alert alert-success">
<font face="Calibri" size="5"> <b> <font color='rgba(200,0,0,0.2)'> <u>EXERCISE</u>: </font> Explore Time Series at Different Point Locations </b> </font>
<font face="Calibri" size="3"> Explore this data set some more by picking different point coordinates to explore. Use the time series animation together with the minimum plot to identify interesting areas and explore the radar brightness history. Discuss with your colleagues what you find.
</font>
</div>
<br>
<hr>
<font face="Calibri" size="2"> <i>Exercise3B-ExploreSARTimeSeriesDeforestation.ipynb - Version 1.5.0 - April 2021
<br>
<b>Version Changes</b>
<ul>
<li>from osgeo import gdal</li>
<li>namespace asf_notebook</li>
</ul>
</i>
</font>
| github_jupyter |
# Prepare tweets and news data for IBM topic
Last modifed: 2017-10-24
# Roadmap
1. Prepare multiprocessing and MongoDB scripts available in ibm_tweets_analysis project
2. Filter out tweets with keyword 'ibm' in tweet_text field from MongoDB db
3. Check basic statistics of embedded URL link in tweet_text to external news article
4. Manually collect external news articles
5. Check ibm_news basic statistics
6. Updated Objective: focus on "social_capital_ceo_palihapitiya_watson_joke" news and tweets
# Steps
```
"""
Initialization
"""
'''
Standard modules
'''
import os
import pickle
import csv
import time
from pprint import pprint
import json
import pymongo
import multiprocessing
import logging
import collections
'''
Analysis modules
'''
%matplotlib inline
%config InlineBackend.figure_format = 'retina' # render double resolution plot output for Retina screens
import matplotlib.pyplot as plt
import pandas as pd
'''
Custom modules
'''
import config
import utilities
import mongodb
import multiprocessing_workers
'''
R magic and packages
'''
# hide all RRuntimeWarnings
import warnings
warnings.filterwarnings('ignore')
# add home for R in anaconda on PATH sys env
os.environ['PATH'] += ':/opt/anaconda3/bin'
# load R magic
%load_ext rpy2.ipython
# load R packages
%R require(ggplot2)
'''
Misc
'''
nb_name = '20171024-daheng-prepare_ibm_tweets_news_data'
# all tweets with keywork 'ibm' in tweet_text field from ND IBM dataset
ibm_tweets_file = os.path.join(config.IBM_TWEETS_NEWS_DIR, 'ibm_tweets.json')
# based on ibm_tweets_file. Duplicate tweets with the same or similar tweet_text are removed
ibm_unique_tweets_file = os.path.join(config.IBM_TWEETS_NEWS_DIR, 'ibm_unique_tweets.json')
# manually selected news sources list by examing most common news sources of valid urls embedded in ibm unique tweets
# selected_news_sources_lst = ['www.forbes.com', 'finance.yahoo.com', 'venturebeat.com',
# 'medium.com', 'www.engadget.com', 'alltheinternetofthings.com',
# 'www.zdnet.com', 'www.wsj.com', 'www.cnbc.com']
selected_news_sources_lst = ['venturebeat', 'engadget', 'wsj', 'cnbc']
# manually collected ibm news data
ibm_news_file = os.path.join(config.HR_DIR, 'selected_ibm_news.csv')
# all tweets related to the 'social_capital_ceo_palihapitiya_watson_joke' news by cnbc
palihapitiya_watson_joke_tweets_file = os.path.join(config.HR_DIR, 'palihapitiya_watson_joke_tweets.csv')
# manually tag information of all tweets related to the 'social_capital_ceo_palihapitiya_watson_joke' news by cnbc
palihapitiya_watson_joke_tweets_tag_file = os.path.join(config.HR_DIR, 'palihapitiya_watson_joke_tweets_tag.csv')
```
## Prepare multiprocessing and MongoDB scripts available in ibm_tweets_analysis project
Copy `mongodb.py` and `multiprocessing_workers.py` files to the project root dir.
- `mongodb.py` can be used to get connection to local MongoDB database.
- `multiprocessing_workers.py` can be used to query MongoDB database in multiple processes to save time (need modifications)
Native tweets are stored in `tweets_ek-2` db and `tw_nt` table.
## Filter out tweets with keyword 'ibm' in tweet_text field from MongoDB db
### Query tweets from MongoDB db
```
%%time
"""
Register
IBM_TWEETS_NEWS_DIR = os.path.join(DATA_DIR, 'ibm_tweets_news')
in config
"""
DB_NAME = 'tweets_ek-2'
COL_NAME = 'tw_nt'
if 0 == 1:
multiprocessing.log_to_stderr(logging.DEBUG)
'''
Use multiprocessing to parse tweet_text field for "ibm" keyword
'''
procedure_name = 'tag_native_tweets_text_ibm'
# set processes number to CPU numbers minus 1
process_num = multiprocessing.cpu_count() - 1
process_file_names_lst = ['{}-{}.json'.format(process_ind, procedure_name)
for process_ind in range(process_num)]
process_files_lst = [os.path.join(config.IBM_TWEETS_NEWS_DIR, process_file_name)
for process_file_name in process_file_names_lst]
jobs = []
for process_ind in range(process_num):
p = multiprocessing.Process(target=multiprocessing_workers.find_keywords_in_tweet_text,
args=(DB_NAME, COL_NAME, process_ind, process_num, process_files_lst[process_ind], ['ibm']),
name='Process-{}/{}'.format(process_ind, process_num))
jobs.append(p)
for job in jobs:
job.start()
for job in jobs:
job.join()
```
### Merge process files
```
%%time
"""
Merger all process files into a single file
Register
ibm_tweets_file = os.path.join(config.IBM_TWEETS_NEWS_DIR, 'ibm_tweets.json')
in Initialization section.
"""
if 0 == 1:
'''
Re-generate process file names
'''
procedure_name = 'tag_native_tweets_text_ibm'
process_num = multiprocessing.cpu_count() - 1
process_file_names_lst = ['{}-{}.json'.format(process_ind, procedure_name)
for process_ind in range(process_num)]
process_files_lst = [os.path.join(config.IBM_TWEETS_NEWS_DIR, process_file_name)
for process_file_name in process_file_names_lst]
with open(ibm_tweets_file, 'w') as output_f:
for process_file in process_files_lst:
with open(process_file, 'r') as input_f:
for line in input_f:
output_f.write(line)
```
### Remove duplicate tweets
```
%%time
"""
Remove tweets with the same or silimar tweet_text field
Register
ibm_unique_tweets_file = os.path.join(config.IBM_TWEETS_NEWS_DIR, 'ibm_unique_tweets.json')
in Initialization section.
"""
if 0 == 1:
with open(ibm_unique_tweets_file, 'w') as output_f:
with open(ibm_tweets_file, 'r') as input_f:
uniqe_tweet_text_field = set()
for line in input_f:
tweet_json = json.loads(line)
tweet_text = tweet_json['text']
cleaned_tweet_text = utilities.clean_tweet_text(tweet_text)
if cleaned_tweet_text not in uniqe_tweet_text_field:
uniqe_tweet_text_field.add(cleaned_tweet_text)
output_f.write(line)
```
## Check basic statistics of embedded URL link in tweet_text to external news article
```
"""
Check number of ibm tweets and number of ibm unique tweets
"""
if 1 == 1:
with open(ibm_tweets_file, 'r') as f:
ibm_tweets_num = sum([1 for line in f])
print('Number of ibm tweets: {}'.format(ibm_tweets_num))
with open(ibm_unique_tweets_file, 'r') as f:
ibm_unique_tweets_num = sum([1 for line in f])
print('Number of unique ibm tweets: {}'.format(ibm_unique_tweets_num))
"""
Check number of ibm unique tweets with URL
"""
if 1 == 1:
with open(ibm_unique_tweets_file, 'r') as f:
# if entities.urls field is not empty
ibm_unique_tweets_url_num = sum([1 for line in f
if json.loads(line)['entities']['urls']])
print('Number of unique ibm tweets with URL: {}'.format(ibm_unique_tweets_url_num))
%%time
"""
Check most popular domain names in URLs embedded in ibm unique tweets
"""
if 1 == 1:
url_domain_names_counter = collections.Counter()
with open(ibm_unique_tweets_file, 'r') as f:
for line in f:
tweet_json = json.loads(line)
# if tweet contains at least one url, entities.urls is not empty
entities_urls = tweet_json['entities']['urls']
if entities_urls:
for entities_url in entities_urls:
# expanded_url field may contain full unshortened url
expanded_url = entities_url['expanded_url']
url_domain_name = expanded_url.split('/')[2]
url_domain_names_counter.update([url_domain_name])
pprint(url_domain_names_counter.most_common(50))
%%time
"""
Re-compute most popular domain names in URLs embedded in ibm unique tweets
- ignore misc irrelevant website domain names
- ignore all shortened urls
Register
selected_news_sources_lst
in Initialization section.
"""
misc_irrelevant_websites_lst = ['twitter', 'youtube', 'youtu.be', 'amazon', 'paper.li', 'linkedin', 'lnkd.in', 'instagram']
shortened_url_identifiers_lst = ['bit.ly', 'ift.tt', 'dlvr.it', 'ow.ly', 'buff.ly', 'oal.lu', 'goo.gl', 'ln.is', 'gag.gl', 'fb.me', 'trap.it', 'ibm.co',
'ibm.biz', 'shar.es', 'crwd.fr', 'klou.tt', 'tek.io', 'owler.us', 'upflow.co', 'hubs.ly', 'zd.net', 'spr.ly', 'flip.it']
if 0 == 1:
valid_url_domain_names_counter = collections.Counter()
ignore_lst = misc_irrelevant_websites_lst + shortened_url_identifiers_lst
with open(ibm_unique_tweets_file, 'r') as f:
for line in f:
tweet_json = json.loads(line)
# if tweet contains at least one url, entities.urls is not empty
entities_urls = tweet_json['entities']['urls']
if entities_urls:
for entities_url in entities_urls:
# expanded_url field may contain full unshortened url
expanded_url = entities_url['expanded_url']
# ignore all urls with manually selected tokens
if not any(token in expanded_url for token in ignore_lst):
# ignore all shortned urls by HEURISTIC
if len(expanded_url.split('/')) > 4:
valid_url_domain_name = expanded_url.split('/')[2]
valid_url_domain_names_counter.update([valid_url_domain_name])
pprint(valid_url_domain_names_counter.most_common(50))
%%time
"""
Check most common valid links
"""
misc_irrelevant_websites_lst = ['twitter', 'youtube', 'youtu.be', 'amazon', 'paper.li', 'linkedin', 'lnkd.in', 'instagram']
shortened_url_identifiers_lst = ['bit.ly', 'ift.tt', 'dlvr.it', 'ow.ly', 'buff.ly', 'oal.lu', 'goo.gl', 'ln.is', 'gag.gl', 'fb.me', 'trap.it', 'ibm.co',
'ibm.biz', 'shar.es', 'crwd.fr', 'klou.tt', 'tek.io', 'owler.us', 'upflow.co', 'hubs.ly', 'zd.net', 'spr.ly', 'flip.it']
if 0 == 1:
urls_counter = collections.Counter()
ignore_lst = misc_irrelevant_websites_lst + shortened_url_identifiers_lst
with open(ibm_unique_tweets_file, 'r') as f:
for line in f:
tweet_json = json.loads(line)
# if tweet contains at least one url, entities.urls is not empty
entities_urls = tweet_json['entities']['urls']
if entities_urls:
for entities_url in entities_urls:
# expanded_url field may contain full unshortened url
expanded_url = entities_url['expanded_url']
# ignore all urls with manually selected tokens
if not any(token in expanded_url for token in ignore_lst):
# ignore all shortned urls by HEURISTIC
if len(expanded_url.split('/')) > 4:
urls_counter.update([expanded_url])
pprint(urls_counter.most_common(50))
%%time
"""
Check most common links to selected news sources
"""
if 0 == 1:
selected_news_sources_urls_counter = collections.Counter()
with open(ibm_tweets_file, 'r') as f:
for line in f:
tweet_json = json.loads(line)
# if tweet contains at least one url, entities.urls is not empty
entities_urls = tweet_json['entities']['urls']
if entities_urls:
for entities_url in entities_urls:
# expanded_url field may contain full unshortened url
expanded_url = entities_url['expanded_url']
# filter out only url links to selected news sources
if any(selected_news_source in expanded_url for selected_news_source in selected_news_sources_lst):
selected_news_sources_urls_counter.update([expanded_url])
pprint(selected_news_sources_urls_counter.most_common(50))
```
## Manually collect external news articles
After examining
- most common valid links
- most common links to selected news sources
manually collect external news articles.
__Note__:
- single news article may have multiple links (shortened by different services; picture/video materials; trivial parameters)
```
"""
Register
ibm_news_file
in Initialization section.
"""
```
## Check ibm_news basic statistics
```
"""
Load in csv file
"""
if 1 == 1:
ibm_news_df = pd.read_csv(filepath_or_buffer=ibm_news_file, sep='\t')
with pd.option_context('display.max_colwidth', 100, 'expand_frame_repr', False):
display(ibm_news_df[['NEWS_DATE', 'NEWS_NAME', 'NEWS_DOC']])
"""
Print any news_doc by paragraphs
"""
test_lst = ibm_news_df.iloc[10]['NEWS_DOC'].split('::::::::')
for ind, item in enumerate(test_lst):
print('({})'.format(ind+1))
print(item)
```
## Updated Objective: focus on "social_capital_ceo_palihapitiya_watson_joke" news and tweets
__New Objective__:
- Only focus on the "social_capital_ceo_palihapitiya_watson_joke" news and tweets.
- generate a illustrative figure, which should be placed in the Introduction section of the paper, to demonstrate the interaction/cycle between news and tweets.
### Find out all related tweets
```
%%time
"""
Find out all tweets related to the 'social_capital_ceo_palihapitiya_watson_joke' news
News URL 1: https://www.cnbc.com/2017/05/08/ibms-watson-is-a-joke-says-social-capital-ceo-palihapitiya.html
News URL 2: https://www.cnbc.com/2017/05/09/no-joke-id-like-to-see-my-firm-go-head-to-head-with-ibm-on-a-i-palihapitiya.html
Register
palihapitiya_watson_joke_tweets_file
in Initialization section
"""
if 0 == 1:
target_news_keywords_lst = ['social capital', 'chamath', 'palihapitiya']
target_tweets_dict_lst = []
with open(ibm_unique_tweets_file, 'r') as f:
for line in f:
tweet_json = json.loads(line)
tweet_text = tweet_json['text'].replace('\n', ' ').replace('\r', ' ')
tweet_user_screen_name = tweet_json['user']['screen_name']
tweet_created_at = utilities.parse_tweet_post_time(tweet_json['created_at'])
if any(kw.lower() in tweet_text.lower() for kw in target_news_keywords_lst):
target_tweet_dict = {'tweet_created_at': tweet_created_at,
'tweet_user_screen_name': tweet_user_screen_name,
'tweet_text': tweet_text}
target_tweets_dict_lst.append(target_tweet_dict)
target_tweets_df = pd.DataFrame(target_tweets_dict_lst)
target_tweets_df.to_csv(path_or_buf=palihapitiya_watson_joke_tweets_file, sep='\t', index=True, quoting=csv.QUOTE_MINIMAL)
"""
Read in data
"""
if 1 == 1:
target_tweets_df = pd.read_csv(filepath_or_buffer=palihapitiya_watson_joke_tweets_file,
sep='\t',
index_col=0,
parse_dates=['tweet_created_at'],
quoting=csv.QUOTE_MINIMAL)
with pd.option_context('display.max_rows', 260, 'display.max_colwidth', 150, 'expand_frame_repr', False):
display(target_tweets_df)
```
### Manually tag each tweet
Manually tag each tweet related to "social_capital_ceo_palihapitiya_watson_joke" news for:
- tweet_sentiment: being neutral (1), mild (2), or stimulant/sarcastic (3)
- tweet_news: correspond to first news (1), or second news (2)
```
"""
Register
palihapitiya_watson_joke_tweets_tag_file
in Initialization section
"""
```
### Check data and quick plot
```
"""
Load data
"""
if 1 == 1:
'''
Read in all tweets related to the 'social_capital_ceo_palihapitiya_watson_joke' news
'''
target_tweets_df = pd.read_csv(filepath_or_buffer=palihapitiya_watson_joke_tweets_file,
sep='\t',
index_col=0,
parse_dates=['tweet_created_at'],
quoting=csv.QUOTE_MINIMAL)
'''
Read in manually tagged information for all tweets just loaded
'''
target_tweets_tag_df = pd.read_csv(filepath_or_buffer=palihapitiya_watson_joke_tweets_tag_file,
sep='\t',
index_col=0)
'''
Combine dfs and set index
'''
test_tweets_df = target_tweets_df.join(target_tweets_tag_df)
test_tweets_df['tweet_index'] = test_tweets_df.index
test_tweets_df = test_tweets_df.set_index('tweet_created_at')
"""
Check tweets related to second news
"""
if 1 == 1:
test_df = test_tweets_df[test_tweets_df['tweet_news'] == 2]
display(test_df)
"""
For tweets related to first news
Build tmp dfs for tweets in mild sentiment and harsh sentiment separately
"""
if 1 == 1:
mild_cond = (test_tweets_df['tweet_news'] == 1) & (test_tweets_df['tweet_sentiment'] == 2)
harsh_cond = (test_tweets_df['tweet_news'] == 1) & (test_tweets_df['tweet_sentiment'] == 3)
mild_tweets_df = test_tweets_df[mild_cond]
harsh_tweets_df = test_tweets_df[harsh_cond]
"""
Check tweets in mild sentiment
"""
print(mild_tweets_df['tweet_index'].count())
with pd.option_context('display.max_rows', 100, 'display.max_colwidth', 150, 'expand_frame_repr', False):
display(mild_tweets_df)
"""
Check tweets in harsh sentiment
"""
print(harsh_tweets_df['tweet_index'].count())
with pd.option_context('display.max_rows', 100, 'display.max_colwidth', 150, 'expand_frame_repr', False):
display(harsh_tweets_df)
"""
Bin mild/harsh tweets by 4H period and count numbers
"""
if 1 == 1:
mild_tweets_bin_count = mild_tweets_df['tweet_index'].resample('4H', convention='start').count().rename('mild_tweets_count')
harsh_tweets_bin_count = harsh_tweets_df['tweet_index'].resample('4H', convention='start').count().rename('harsh_tweets_count')
tweets_count = pd.concat([mild_tweets_bin_count, harsh_tweets_bin_count], axis=1)[:24]
with pd.option_context('display.max_rows', 100, 'display.max_colwidth', 150, 'expand_frame_repr', False):
display(tweets_count)
if 1 == 1:
tweets_count.plot(kind="bar", figsize=(12,6), title='# of mild/harsh tweets', stacked=True)
```
### Plot use R ggplot2
```
"""
Prepare df data
"""
if 1 == 1:
'''
Read in all tweets related to the 'social_capital_ceo_palihapitiya_watson_joke' news
'''
target_tweets_df = pd.read_csv(filepath_or_buffer=palihapitiya_watson_joke_tweets_file,
sep='\t',
index_col=0,
parse_dates=['tweet_created_at'],
quoting=csv.QUOTE_MINIMAL)
'''
Read in manually tagged information for all tweets just loaded
'''
target_tweets_tag_df = pd.read_csv(filepath_or_buffer=palihapitiya_watson_joke_tweets_tag_file,
sep='\t',
index_col=0)
'''
Join dfs and set index
'''
test_tweets_df = target_tweets_df.join(target_tweets_tag_df)
test_tweets_df['tweet_index'] = test_tweets_df.index
test_tweets_df = test_tweets_df.set_index('tweet_created_at')
'''
Bin mild/harsh tweets by 4H period and count numbers
'''
mild_tweets_df = test_tweets_df[(test_tweets_df['tweet_news'] == 1) & (test_tweets_df['tweet_sentiment'] == 2)]
harsh_tweets_df = test_tweets_df[(test_tweets_df['tweet_news'] == 1) & (test_tweets_df['tweet_sentiment'] == 3)]
second_news_mild_tweets_df = test_tweets_df[(test_tweets_df['tweet_news'] == 2) & (test_tweets_df['tweet_sentiment'] == 2)]
mild_tweets_bin_count = mild_tweets_df['tweet_index'].resample('4H', label='start', loffset='2H 1S').count().rename('mild_tweets_count')
harsh_tweets_bin_count = harsh_tweets_df['tweet_index'].resample('4H', label='start', loffset='2H 1S').count().rename('harsh_tweets_count')
second_news_mild_tweets_bin_count = second_news_mild_tweets_df['tweet_index'].resample('4H', label='start', loffset='2H 1S').count().rename('second_news_mild_tweets_count')
tweets_count = pd.concat([mild_tweets_bin_count, harsh_tweets_bin_count, second_news_mild_tweets_bin_count], axis=1)
'''
Misc operations
'''
tweets_count = tweets_count.fillna(0)
tweets_count['mild_tweets_count'] = tweets_count['mild_tweets_count'].astype(int)
tweets_count['harsh_mild_diff'] = tweets_count['harsh_tweets_count'] - tweets_count['mild_tweets_count']
tweets_count['mild_tweets_count_neg'] = - tweets_count['mild_tweets_count']
tweets_count['second_news_mild_tweets_count'] = tweets_count['second_news_mild_tweets_count'].astype(int)
tweets_count['second_news_mild_tweets_count_neg'] = - tweets_count['second_news_mild_tweets_count']
tweets_count.reset_index(drop=False, inplace=True)
tweets_r_df = tweets_count
tweets_r_df
%%R -i tweets_r_df
#
# Prepare data
#
# cast data types
tweets_r_df$tweet_created_at <- as.POSIXct(strptime(tweets_r_df$tweet_created_at, format="%Y-%m-%d %H:%M:%S"))
#
# Plot and tweak histogram
#
# initialize new plot
# cols <- c('Harsh'='red', 'Mild'='blue', 'diff_line'='black')
plt <- ggplot(data=tweets_r_df, aes(x=tweet_created_at)) +
# layers of ref lines for publishing times of first and second news
geom_vline(xintercept=as.POSIXct(strptime('2017-05-08 16:45:00', format="%Y-%m-%d %H:%M:%S")), linetype='dashed', color='grey80') +
geom_vline(xintercept=as.POSIXct(strptime('2017-05-09 09:55:00', format="%Y-%m-%d %H:%M:%S")), linetype='dashed', color='grey80') +
# layer of geom_bar for harsh tweets
geom_bar(aes(y=harsh_tweets_count, fill='Harsh'), stat='identity', alpha=0.65) +
# layer of geom_rect for highlighting largest bar
geom_rect(aes(xmin=as.POSIXct(strptime('2017-05-09 12:15:00', format="%Y-%m-%d %H:%M:%S")),
xmax=as.POSIXct(strptime('2017-05-09 15:45:00', format="%Y-%m-%d %H:%M:%S")),
ymin=0, ymax=27), fill=NA, color="red", size=0.7, alpha=1) +
# layer of geom_bar for mild tweets
geom_bar(aes(y=mild_tweets_count_neg, fill='Mild'), stat='identity', alpha=0.65) +
# layer of geom_line for diff between harsh tweets and mild tweets
geom_line(aes(x=(tweet_created_at), y=harsh_mild_diff), stat='identity', linetype='solid') +
# layer of geom_bar for a few tweets related to second news in mild sentiment
geom_bar(aes(y=second_news_mild_tweets_count_neg), stat='identity', alpha=0.65, fill='green') +
# x-axis and y-axis
scale_x_datetime(name = 'Time',
date_labels = "%b %d %I%p",
date_breaks = "4 hour",
expand = c(0, 0),
limits = c(as.POSIXct(strptime('2017-05-08 12:00:00', format="%Y-%m-%d %H:%M:%S")),
as.POSIXct(strptime('2017-05-10 19:00:00', format="%Y-%m-%d %H:%M:%S")))) +
scale_y_continuous(name = 'Number of users',
breaks = c(-10, -5, 0, 5, 10, 15, 20, 25),
labels = c('10', '5', '0', '5', '10', '15', '20', '25'),
limits = c(-15, 30)) +
# legend
scale_fill_manual(name = "Sentiment Intensity",
values = c('Harsh'='red', 'Mild'='blue')) +
# theme
theme(panel.background = element_blank(),
axis.line = element_line(color='black'),
panel.grid.major.y = element_line(color='grey80'),
panel.grid.major.x = element_blank(),
panel.grid.minor = element_blank(),
axis.text.x = element_text(angle=90),
legend.position = 'top')
#
# Output figure
#
ggsave('./fig/ibm_joke_or_not.png', plt, height=5, width=5, dpi=200)
```
| github_jupyter |
## Matching catalogues to the VAST Pilot Survey
This notebook gives an example of how to use vast-tools in a notebook environment to perform a crossmatch between a catalogue and the VAST Pilot Survey.
**Note** The settings and filters applied in this notebook, while sensible, are somewhat generic - always consider your science goals on what filters you want to make!
It is **highly recommended** that results from the VAST Pipeline are used and this is what will be primarily covered in this example. It is possible to run a search just using vast-tools but the results are nowhere near as rich as the pipeline - this is covered at the of this document.
### The VAST Pipeline
The pipeline hosted on the Nimbus server will contain the pipeline run for the full pilot survey. For a complete demo of what can be done with the vast-tools `Pipeline` class see the `vast-pipeline-example.ipynb` example notebook.
### The Imports
Below are the imports required for this example. The main imports required from vast-tools are the Pipeline and VASTMOCS objects. The Query object is for the vast-tools query option that is shown at the end of this notebook. Astropy objects are also imported as they are critical to perfoming the crossmatch.
```
from vasttools.moc import VASTMOCS
from vasttools.pipeline import Pipeline
from vasttools.query import Query
from mocpy import World2ScreenMPL
import matplotlib.pyplot as plt
from astropy import units as u
from astropy.coordinates import Angle, SkyCoord
```
### Catalogue selection
For this example we will be using the `Quasars and Active Galactic Nuclei (13th Ed.) (Veron+ 2010)` catalogue, which has the Vizier ID of `VII/258`.
_**Note:** Of course your catalogue doesn't have to come from Vizier. If you have a `csv` or `FITS` file then simply load this data into a DataFrame, create a SkyCoord object and you'll be good to go._
To start our search, the first question we want to answer is:
*What sources from the catalogue are in the VAST Pilot Survey footprint?*
This can be efficiently answered by using the `query_vizier_vast_pilot()` method in VASTMOCS.
First we initialise the VASTMOCS object:
```
mocs = VASTMOCS()
```
We then use the query vizier method to obtain all the sources from the Veron catalogue which are contained within the footprint. It will likely take a bit of time to complete.
```
veron_vast_sources = mocs.query_vizier_vast_pilot('VII/258', max_rows=200000)
veron_vast_sources
```
We see that 44,704 sources are within the VAST Pilot Survey footprint.
_**Tip:** The table returned above is an astropy.table. This can be converted to pandas by using `veron_vast_sources = veron_vast_sources.to_pandas()`._
These can be plotted along with the VAST Pilot Survey footprint using the MOC. See the vast-mocs-example notebook for more on using MOCS and the `Wordl2ScreenMPL` method.
```
from astropy.visualization.wcsaxes.frame import EllipticalFrame
fig = plt.figure(figsize=(16,8))
# Load the Epoch 1 MOC file to use
epoch1_moc = mocs.load_pilot_epoch_moc('1')
#
with World2ScreenMPL(
fig,
fov=320 * u.deg,
center=SkyCoord(0, 0, unit='deg', frame='icrs'),
coordsys="icrs",
rotation=Angle(0, u.degree),
) as wcs:
ax = fig.add_subplot(111, projection=wcs, frame_class=EllipticalFrame)
ax.set_title("Veron Catalogue Sources in the VAST Pilot Survey")
ax.grid(color="black", linestyle="dotted")
epoch1_moc.fill(ax=ax, wcs=wcs, alpha=0.5, fill=True, linewidth=0, color="#00bb00")
epoch1_moc.border(ax=ax, wcs=wcs, alpha=0.5, color="black")
ax.scatter(
veron_vast_sources['_RAJ2000'],
veron_vast_sources['_DEJ2000'],
transform=ax.get_transform('world'),
zorder=10,
s=3
)
fig
```
### Loading the VAST Pipeline Data
Now the results of the VAST Pipeline need to be loaded. This example will not give full details of the Pipeline class, but please refer to the `vast-pipeline-example.ipynb` example notebook for a full example and description.
We'll be using the full VAST Pilot Survey pipeline containing epochs 0–13 (a test version called `tiles_corrected`).
```
# below I suppress DeprecationWarnings due to ipykernel bug and an astropy warning due to FITS header warnings.
import warnings
from astropy.utils.exceptions import AstropyWarning
warnings.simplefilter('ignore', category=AstropyWarning)
warnings.filterwarnings("ignore", category=DeprecationWarning)
# define pipeline object
pipe = Pipeline()
# load the run
pipe_run = pipe.load_run('tiles_corrected')
```
We now have access to the unique sources found by the pipeline:
```
pipe_run.sources.head()
```
### Performing the Crossmatch
The crossmatch can be performed using the astropy `match_to_catalog_sky` function. The first step is to create the sky coord objects for each catalogue. First the Veron catalog which was already obtained above:
```
# Unfortunately we cannot use guess_from_table for the Vizier results, so we construct manually
veron_skycoord = SkyCoord(veron_vast_sources['_RAJ2000'], veron_vast_sources['_DEJ2000'], unit=(u.deg, u.deg))
veron_names = veron_vast_sources['Name'].tolist()
```
and then by default the pipeline run object has the default sources saved as a sky coord object as `pipe_run.sources_skycoord`:
```
pipe_run.sources_skycoord
```
Now the crossmatching can be performed. See https://docs.astropy.org/en/stable/coordinates/matchsep.html#astropy-coordinates-matching for details on the astropy functions and outputs.
```
idx, d2d, d3d = veron_skycoord.match_to_catalog_sky(pipe_run.sources_skycoord)
radius_limit = 15 * u.arcsec
(d2d <= radius_limit).sum()
```
From above we can see that 5048 Veron objects have a match to the pipeline sources. If you wish you could merge the results together:
```
# Convert Veron to pandas first
veron_vast_sources_pd = veron_vast_sources.to_pandas()
# Create a d2d mask
d2d_mask = d2d <= radius_limit
# Select the crossmatches less than 15 arcsec
veron_crossmatch_result_15asec = veron_vast_sources_pd.loc[d2d_mask].copy()
# Append the id and distance of the VAST crossmatch to the veron sources
veron_crossmatch_result_15asec['vast_xmatch_id'] = pipe_run.sources.iloc[idx[d2d_mask]].index.values
veron_crossmatch_result_15asec['vast_xmatch_d2d_asec'] = d2d[d2d_mask].arcsec
# Join the result
veron_crossmatch_result_15asec = veron_crossmatch_result_15asec.merge(pipe_run.sources, how='left', left_on='vast_xmatch_id', right_index=True, suffixes=("_veron", "_vast"))
veron_crossmatch_result_15asec
```
With the crossmatches in hand you can now start to do any kind of analysis you wish to perform. For example we can perform a quick check to see if the pipeline has picked out any of these sources as having significant two-epoch variability:
```
veron_crossmatch_result_15asec[veron_crossmatch_result_15asec['m_abs_significant_max_peak'] > 0.00]
```
And remember you can use the vast-toools source tools to view any source as in the other example notebooks:
```
# Get the first VAST source above from the table above
first_source_id = veron_crossmatch_result_15asec[veron_crossmatch_result_15asec['m_abs_significant_max_peak'] > 0.00].iloc[0].vast_xmatch_id
first_source = pipe_run.get_source(first_source_id)
first_source.plot_lightcurve(min_points=1)
first_source.show_all_png_cutouts(columns=5, figsize=(12,7), size=Angle(2. * u.arcmin))
```
### Filtering the Pipeline Sources (Optional)
The example above has used all the sources from the pipeline results, but these may need to be filtered further to improve results. For example Below is an example of how to filter the sources.
```
my_query_string = (
"n_measurements >= 3 "
"& n_selavy >= 2 "
"& n_neighbour_dist > 1./60. "
"& 0.8 < avg_compactness < 1.4 "
"& n_relations == 0 "
"& max_snr > 7.0"
)
pipe_run_filtered_sources = pipe_run.sources.query(my_query_string)
pipe_run_filtered_sources
```
You can either:
* apply this to the crossmatch results above, or
* substitute `pipe_run_filtered_sources` into the complete crossmatch process above in the place of `my_run.sources` (you need to create a new SkyCoord object first).
```
pipe_run_filtered_sources_skycoord = pipe_run.get_sources_skycoord(user_sources=pipe_run_filtered_sources)
pipe_run_filtered_sources_skycoord
```
### Finding All Crossmatches Between Sources
The crossmatch above only finds the nearest neighbour to the sources in your catalogue. Astropy also offers the functionality to find all matches between objects within a defined radius. See https://docs.astropy.org/en/stable/coordinates/matchsep.html#searching-around-coordinates for full details. This is done by performing the below, using the 15 arcsec radius:
```
idx_vast, idx_veron, d2d, d3d = veron_skycoord.search_around_sky(pipe_run.sources_skycoord, 15 * u.arcsec)
```
A merged dataframe of this crossmatch can be made like that below. Note there are multiple matches to sources so this will generate duplicate sources within the dataframe.
```
# Create a subset dataframe of the Veron sources with a match
veron_search_around_results_15asec = veron_vast_sources_pd.iloc[idx_veron].copy()
# Add the VAST d2d and match id columns
veron_search_around_results_15asec['vast_xmatch_d2d_asec'] = d2d.arcsec
veron_search_around_results_15asec['vast_xmatch_id'] = pipe_run.sources.iloc[idx_vast].index.values
# Perform the merge
veron_search_around_results_15asec = veron_search_around_results_15asec.merge(pipe_run.sources, how='left', left_on='vast_xmatch_id', right_index=True, suffixes=("_veron", "_vast"))
veron_search_around_results_15asec
```
This is the end of the example of performing a catalogue crossmatch using the VAST Pipeline. The information below this point is about using the vast-tools query method to find sources from the pilot survey if a pipeline run is not available. A pipeline run should be used whenever possible due to the superior quality of data it generates.
## Find VAST Matches Using VAST Tools
If a pipeline run isn't available you can use VAST Tools to match to the **VAST Pilot Survey only**.
Here the same Veron dataframe that was created in the pipeline section above is used.
The first step is to construct a Query to see how many sources have matches to selavy componentst in the VAST Pilot Survey. In the Query definition below we use the `matches_only` argument. This means that only those sources that have an actual match are returned. I also explicitly do not select RACS to search here, I'm only interested in the VAST Pilot data, so I select `all-vast`. Note you must pre-create the output directory for the query if you intend to use it.
```
veron_query = Query(
coords=veron_skycoord,
source_names=veron_names,
epochs='all-vast',
max_sep=1.5,
crossmatch_radius=10.0,
base_folder='/data/vast-survey/pilot/',
matches_only=True,
no_rms=True,
output_dir='veron-vast-crossmatching',
)
```
And run `find_sources` - again a warning that this will take a little while to process.
```
veron_query.find_sources()
```
We can check the results attribute to see how many sources return a match.
```
veron_query.results.shape[0]
```
### Using the results
4664 sources have returned a match in the VAST Pilot Survey in any epoch.
We can create new skycoord and name objects ready for a new query:
```
matches_mask = [i in (veron_query.results) for i in veron_vast_sources['Name']]
matched_names = veron_vast_sources['Name'][matches_mask].tolist()
matched_skycoords = veron_skycoord[matches_mask]
```
Or loop through and save all the measurements for each source.
```
# for i in veron_query.results:
# i.write_measurements()
```
While you can explore the sources as normal, for example
```
my_source = veron_query.results['1AXG J134412+0016']
lc = my_source.plot_lightcurve()
lc
cutout = my_source.show_png_cutout('1')
cutout
```
it's not recommended to produce cut outs for all sources in the notebook as this will start to take a lot of memory and be quite slow. If you'd like to do this then please use the `find_sources.py` script.
### VAST Tools Variability
Unlike the Pipeline, the sources returned using this method do not contain any of the caluclated metrics. However, you can also perform some rudimentary variablility analysis on the results if you wish.
I would recommened using the VAST Pipeline if possible for this kind of analysis as the associations will be much better and the you'll get a lot more information, but nevertheless this is an example of what you **can** do with the data from vast-tools.
In the code below I create a dataframe from the query results (which is a pandas series) and assign it to `variables_df` and define a function that returns the eta and V metrics for each source when passed through `.apply()`. These are then assigned to new `eta` and `v` columns in the `variables_df` dataframe.
```
import pandas as pd
def get_variable_metrics(row):
"""
Function to return the eta and v metrics using apply.
"""
return row['object'].calc_eta_and_v_metrics()
# create the variables_df dataframe, rename the column holding the objects as 'object'
variables_df = pd.DataFrame(veron_query.results).rename(columns={'name':'object'})
# obtain the metrics
variables_df[['eta', 'v']] = variables_df.apply(get_variable_metrics, result_type='expand', axis=1)
```
We can then, for example, plot the log eta distribution, making sure we choose sources that have more than 2 detections.
```
%matplotlib inline
mask = [i.detections > 2 for i in variables_df['object']]
import numpy as np
np.log10(variables_df.eta[mask]).hist(bins=100)
plt.show()
```
You could then do the same for `v` and start to fit Gaussians to the distributions and select candidates.
**Note** for large queries it is recommened to use the script version of `find_sources.py` to get cutouts for **all** results.
| github_jupyter |
# 📃 Solution for Exercise M6.04
The aim of this exercise is to:
* verify if a GBDT tends to overfit if the number of estimators is not
appropriate as previously seen for AdaBoost;
* use the early-stopping strategy to avoid adding unnecessary trees, to
get the best generalization performances.
We will use the California housing dataset to conduct our experiments.
```
from sklearn.datasets import fetch_california_housing
from sklearn.model_selection import train_test_split
data, target = fetch_california_housing(return_X_y=True, as_frame=True)
target *= 100 # rescale the target in k$
data_train, data_test, target_train, target_test = train_test_split(
data, target, random_state=0, test_size=0.5)
```
<div class="admonition note alert alert-info">
<p class="first admonition-title" style="font-weight: bold;">Note</p>
<p class="last">If you want a deeper overview regarding this dataset, you can refer to the
Appendix - Datasets description section at the end of this MOOC.</p>
</div>
Create a gradient boosting decision tree with `max_depth=5` and
`learning_rate=0.5`.
```
# solution
from sklearn.ensemble import GradientBoostingRegressor
gbdt = GradientBoostingRegressor(max_depth=5, learning_rate=0.5)
```
Create a validation curve to assess the impact of the number of trees
on the generalization performance of the model. Evaluate the list of parameters
`param_range = [1, 2, 5, 10, 20, 50, 100]` and use the mean absolute error
to assess the generalization performance of the model.
```
# solution
from sklearn.model_selection import validation_curve
param_range = [1, 2, 5, 10, 20, 50, 100]
gbdt_train_scores, gbdt_test_scores = validation_curve(
gbdt,
data_train,
target_train,
param_name="n_estimators",
param_range=param_range,
scoring="neg_mean_absolute_error",
n_jobs=2,
)
gbdt_train_errors, gbdt_test_errors = -gbdt_train_scores, -gbdt_test_scores
import matplotlib.pyplot as plt
plt.errorbar(
param_range,
gbdt_train_errors.mean(axis=1),
yerr=gbdt_train_errors.std(axis=1),
label="Training",
)
plt.errorbar(
param_range,
gbdt_test_errors.mean(axis=1),
yerr=gbdt_test_errors.std(axis=1),
label="Cross-validation",
)
plt.legend()
plt.ylabel("Mean absolute error in k$\n(smaller is better)")
plt.xlabel("# estimators")
_ = plt.title("Validation curve for GBDT regressor")
```
Unlike AdaBoost, the gradient boosting model will always improve when
increasing the number of trees in the ensemble. However, it will reach a
plateau where adding new trees will just make fitting and scoring slower.
To avoid adding new unnecessary tree, gradient boosting offers an
early-stopping option. Internally, the algorithm will use an out-of-sample
set to compute the generalization performance of the model at each addition of a
tree. Thus, if the generalization performance is not improving for several
iterations, it will stop adding trees.
Now, create a gradient-boosting model with `n_estimators=1000`. This number
of trees will be too large. Change the parameter `n_iter_no_change` such
that the gradient boosting fitting will stop after adding 5 trees that do not
improve the overall generalization performance.
```
# solution
gbdt = GradientBoostingRegressor(n_estimators=1000, n_iter_no_change=5)
gbdt.fit(data_train, target_train)
gbdt.n_estimators_
```
We see that the number of trees used is far below 1000 with the current
dataset. Training the GBDT with the entire 1000 trees would have been
useless.
| github_jupyter |
Source: https://github.com/yandexdataschool/nlp_course/tree/2021/week04_seq2seq
## Seminar and homework (12 points total)
Today we shall compose encoder-decoder neural networks and apply them to the task of machine translation.
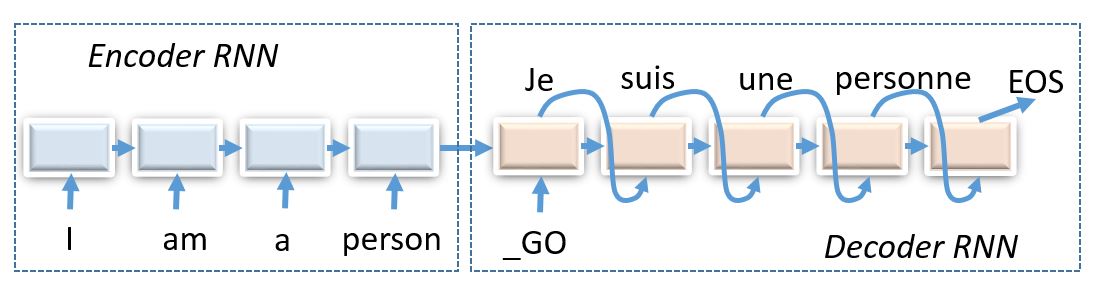
_(img: esciencegroup.files.wordpress.com)_
Encoder-decoder architectures are about converting anything to anything, including
* Machine translation and spoken dialogue systems
* [Image captioning](http://mscoco.org/dataset/#captions-challenge2015) and [image2latex](https://openai.com/requests-for-research/#im2latex) (convolutional encoder, recurrent decoder)
* Generating [images by captions](https://arxiv.org/abs/1511.02793) (recurrent encoder, convolutional decoder)
* Grapheme2phoneme - convert words to transcripts
## Our task: machine translation
We gonna try our encoder-decoder models on russian to english machine translation problem. More specifically, we'll translate hotel and hostel descriptions. This task shows the scale of machine translation while not requiring you to train your model for weeks if you don't use GPU.
Before we get to the architecture, there's some preprocessing to be done. ~~Go tokenize~~ Alright, this time we've done preprocessing for you. As usual, the data will be tokenized with WordPunctTokenizer.
However, there's one more thing to do. Our data lines contain unique rare words. If we operate on a word level, we will have to deal with large vocabulary size. If instead we use character-level models, it would take lots of iterations to process a sequence. This time we're gonna pick something inbetween.
One popular approach is called [Byte Pair Encoding](https://github.com/rsennrich/subword-nmt) aka __BPE__. The algorithm starts with a character-level tokenization and then iteratively merges most frequent pairs for N iterations. This results in frequent words being merged into a single token and rare words split into syllables or even characters.
```
!pip3 install torch>=1.3.0
!pip3 install subword-nmt &> log
!wget https://www.dropbox.com/s/yy2zqh34dyhv07i/data.txt?dl=1 -O data.txt
!wget https://raw.githubusercontent.com/yandexdataschool/nlp_course/2020/week04_seq2seq/vocab.py -O vocab.py
# thanks to tilda and deephack teams for the data, Dmitry Emelyanenko for the code :)
from nltk.tokenize import WordPunctTokenizer
from subword_nmt.learn_bpe import learn_bpe
from subword_nmt.apply_bpe import BPE
tokenizer = WordPunctTokenizer()
def tokenize(x):
return ' '.join(tokenizer.tokenize(x.lower()))
# split and tokenize the data
with open('train.en', 'w') as f_src, open('train.ru', 'w') as f_dst:
for line in open('data.txt'):
src_line, dst_line = line.strip().split('\t')
f_src.write(tokenize(src_line) + '\n')
f_dst.write(tokenize(dst_line) + '\n')
# build and apply bpe vocs
bpe = {}
for lang in ['en', 'ru']:
learn_bpe(open('./train.' + lang), open('bpe_rules.' + lang, 'w'), num_symbols=8000)
bpe[lang] = BPE(open('./bpe_rules.' + lang))
with open('train.bpe.' + lang, 'w') as f_out:
for line in open('train.' + lang):
f_out.write(bpe[lang].process_line(line.strip()) + '\n')
```
### Building vocabularies
We now need to build vocabularies that map strings to token ids and vice versa. We're gonna need these fellas when we feed training data into model or convert output matrices into words.
```
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
data_inp = np.array(open('./train.bpe.ru').read().split('\n'))
data_out = np.array(open('./train.bpe.en').read().split('\n'))
from sklearn.model_selection import train_test_split
train_inp, dev_inp, train_out, dev_out = train_test_split(data_inp, data_out, test_size=3000,
random_state=42)
for i in range(3):
print('inp:', train_inp[i])
print('out:', train_out[i], end='\n\n')
from vocab import Vocab
inp_voc = Vocab.from_lines(train_inp)
out_voc = Vocab.from_lines(train_out)
# Here's how you cast lines into ids and backwards.
batch_lines = sorted(train_inp, key=len)[5:10]
batch_ids = inp_voc.to_matrix(batch_lines)
batch_lines_restored = inp_voc.to_lines(batch_ids)
print("lines")
print(batch_lines)
print("\nwords to ids (0 = bos, 1 = eos):")
print(batch_ids)
print("\nback to words")
print(batch_lines_restored)
```
Draw source and translation length distributions to estimate the scope of the task.
```
plt.figure(figsize=[8, 4])
plt.subplot(1, 2, 1)
plt.title("source length")
plt.hist(list(map(len, map(str.split, train_inp))), bins=20);
plt.subplot(1, 2, 2)
plt.title("translation length")
plt.hist(list(map(len, map(str.split, train_out))), bins=20);
```
### Encoder-decoder model (2 points)
The code below contains a template for a simple encoder-decoder model: single GRU encoder/decoder, no attention or anything. This model is implemented for you as a reference and a baseline for your homework assignment.
```
import torch
import torch.nn as nn
import torch.nn.functional as F
device = 'cuda' if torch.cuda.is_available() else 'cpu'
class BasicModel(nn.Module):
def __init__(self, inp_voc, out_voc, emb_size=64, hid_size=128):
"""
A simple encoder-decoder seq2seq model
"""
super().__init__() # initialize base class to track sub-layers, parameters, etc.
self.inp_voc, self.out_voc = inp_voc, out_voc
self.hid_size = hid_size
self.emb_inp = nn.Embedding(len(inp_voc), emb_size)
self.emb_out = nn.Embedding(len(out_voc), emb_size)
self.enc0 = nn.GRU(emb_size, hid_size, batch_first=True)
self.dec_start = nn.Linear(hid_size, hid_size)
self.dec0 = nn.GRUCell(emb_size, hid_size)
self.logits = nn.Linear(hid_size, len(out_voc))
def forward(self, inp, out):
""" Apply model in training mode """
initial_state = self.encode(inp)
return self.decode(initial_state, out)
def encode(self, inp, **flags):
"""
Takes symbolic input sequence, computes initial state
:param inp: matrix of input tokens [batch, time]
:returns: initial decoder state tensors, one or many
"""
inp_emb = self.emb_inp(inp)
batch_size = inp.shape[0]
enc_seq, [last_state_but_not_really] = self.enc0(inp_emb)
# enc_seq: [batch, time, hid_size], last_state: [batch, hid_size]
# note: last_state is not _actually_ last because of padding, let's find the real last_state
lengths = (inp != self.inp_voc.eos_ix).to(torch.int64).sum(dim=1).clamp_max(inp.shape[1] - 1)
last_state = enc_seq[torch.arange(len(enc_seq)), lengths]
# ^-- shape: [batch_size, hid_size]
dec_start = self.dec_start(last_state)
return [dec_start]
def decode_step(self, prev_state, prev_tokens, **flags):
"""
Takes previous decoder state and tokens, returns new state and logits for next tokens
:param prev_state: a list of previous decoder state tensors, same as returned by encode(...)
:param prev_tokens: previous output tokens, an int vector of [batch_size]
:return: a list of next decoder state tensors, a tensor of logits [batch, len(out_voc)]
"""
prev_gru0_state = prev_state[0]
# <YOUR CODE HERE>
return new_dec_state, output_logits
def decode(self, initial_state, out_tokens, **flags):
""" Iterate over reference tokens (out_tokens) with decode_step """
batch_size = out_tokens.shape[0]
state = initial_state
# initial logits: always predict BOS
onehot_bos = F.one_hot(torch.full([batch_size], self.out_voc.bos_ix, dtype=torch.int64),
num_classes=len(self.out_voc)).to(device=out_tokens.device)
first_logits = torch.log(onehot_bos.to(torch.float32) + 1e-9)
logits_sequence = [first_logits]
for i in range(out_tokens.shape[1] - 1):
state, logits = self.decode_step(state, out_tokens[:, i])
logits_sequence.append(logits)
return torch.stack(logits_sequence, dim=1)
def decode_inference(self, initial_state, max_len=100, **flags):
""" Generate translations from model (greedy version) """
batch_size, device = len(initial_state[0]), initial_state[0].device
state = initial_state
outputs = [torch.full([batch_size], self.out_voc.bos_ix, dtype=torch.int64,
device=device)]
all_states = [initial_state]
for i in range(max_len):
state, logits = self.decode_step(state, outputs[-1])
outputs.append(logits.argmax(dim=-1))
all_states.append(state)
return torch.stack(outputs, dim=1), all_states
def translate_lines(self, inp_lines, **kwargs):
inp = self.inp_voc.to_matrix(inp_lines).to(device)
initial_state = self.encode(inp)
out_ids, states = self.decode_inference(initial_state, **kwargs)
return self.out_voc.to_lines(out_ids.cpu().numpy()), states
# debugging area
model = BasicModel(inp_voc, out_voc).to(device)
dummy_inp_tokens = inp_voc.to_matrix(sorted(train_inp, key=len)[5:10]).to(device)
dummy_out_tokens = out_voc.to_matrix(sorted(train_out, key=len)[5:10]).to(device)
h0 = model.encode(dummy_inp_tokens)
h1, logits1 = model.decode_step(h0, torch.arange(len(dummy_inp_tokens), device=device))
assert isinstance(h1, list) and len(h1) == len(h0)
assert h1[0].shape == h0[0].shape and not torch.allclose(h1[0], h0[0])
assert logits1.shape == (len(dummy_inp_tokens), len(out_voc))
logits_seq = model.decode(h0, dummy_out_tokens)
assert logits_seq.shape == (dummy_out_tokens.shape[0], dummy_out_tokens.shape[1], len(out_voc))
# full forward
logits_seq2 = model(dummy_inp_tokens, dummy_out_tokens)
assert logits_seq2.shape == logits_seq.shape
dummy_translations, dummy_states = model.translate_lines(train_inp[:3], max_len=25)
print("Translations without training:")
print('\n'.join([line for line in dummy_translations]))
```
### Training loss (2 points)
Our training objective is almost the same as it was for neural language models:
$$ L = {\frac1{|D|}} \sum_{X, Y \in D} \sum_{y_t \in Y} - \log p(y_t \mid y_1, \dots, y_{t-1}, X, \theta) $$
where $|D|$ is the __total length of all sequences__, including BOS and first EOS, but excluding PAD.
```
def compute_loss(model, inp, out, **flags):
"""
Compute loss (float32 scalar) as in the formula above
:param inp: input tokens matrix, int32[batch, time]
:param out: reference tokens matrix, int32[batch, time]
In order to pass the tests, your function should
* include loss at first EOS but not the subsequent ones
* divide sum of losses by a sum of input lengths (use voc.compute_mask)
"""
mask = model.out_voc.compute_mask(out) # [batch_size, out_len]
targets_1hot = F.one_hot(out, len(model.out_voc)).to(torch.float32)
# outputs of the model, [batch_size, out_len, num_tokens]
logits_seq = <YOUR CODE HERE>
# log-probabilities of all tokens at all steps, [batch_size, out_len, num_tokens]
logprobs_seq = <YOUR CODE HERE>
# log-probabilities of correct outputs, [batch_size, out_len]
logp_out = (logprobs_seq * targets_1hot).sum(dim=-1)
# ^-- this will select the probability of the actual next token.
# Note: you can compute loss more efficiently using using F.cross_entropy
# average cross-entropy over tokens where mask == True
return <YOUR CODE HERE> # average loss, scalar
dummy_loss = compute_loss(model, dummy_inp_tokens, dummy_out_tokens)
print("Loss:", dummy_loss)
assert np.allclose(dummy_loss.item(), 7.5, rtol=0.1, atol=0.1), "We're sorry for your loss"
# test autograd
dummy_loss.backward()
for name, param in model.named_parameters():
assert param.grad is not None and abs(param.grad.max()) != 0, f"Param {name} received no gradients"
```
### Evaluation: BLEU
Machine translation is commonly evaluated with [BLEU](https://en.wikipedia.org/wiki/BLEU) score. This metric simply computes which fraction of predicted n-grams is actually present in the reference translation. It does so for n=1,2,3 and 4 and computes the geometric average with penalty if translation is shorter than reference.
While BLEU [has many drawbacks](http://www.cs.jhu.edu/~ccb/publications/re-evaluating-the-role-of-bleu-in-mt-research.pdf), it still remains the most commonly used metric and one of the simplest to compute.
```
from nltk.translate.bleu_score import corpus_bleu
def compute_bleu(model, inp_lines, out_lines, bpe_sep='@@ ', **flags):
"""
Estimates corpora-level BLEU score of model's translations given inp and reference out
Note: if you're serious about reporting your results, use https://pypi.org/project/sacrebleu
"""
with torch.no_grad():
translations, _ = model.translate_lines(inp_lines, **flags)
translations = [line.replace(bpe_sep, '') for line in translations]
actual = [line.replace(bpe_sep, '') for line in out_lines]
return corpus_bleu(
[[ref.split()] for ref in actual],
[trans.split() for trans in translations],
smoothing_function=lambda precisions, **kw: [p + 1.0 / p.denominator for p in precisions]
) * 100
compute_bleu(model, dev_inp, dev_out)
```
### Training loop
Training encoder-decoder models isn't that different from any other models: sample batches, compute loss, backprop and update
```
from IPython.display import clear_output
from tqdm import tqdm, trange
metrics = {'train_loss': [], 'dev_bleu': [] }
model = BasicModel(inp_voc, out_voc).to(device)
opt = torch.optim.Adam(model.parameters(), lr=1e-3)
batch_size = 32
for _ in trange(25000):
step = len(metrics['train_loss']) + 1
batch_ix = np.random.randint(len(train_inp), size=batch_size)
batch_inp = inp_voc.to_matrix(train_inp[batch_ix]).to(device)
batch_out = out_voc.to_matrix(train_out[batch_ix]).to(device)
<YOUR CODE: training step using batch_inp and batch_out>
metrics['train_loss'].append((step, loss_t.item()))
if step % 100 == 0:
metrics['dev_bleu'].append((step, compute_bleu(model, dev_inp, dev_out)))
clear_output(True)
plt.figure(figsize=(12,4))
for i, (name, history) in enumerate(sorted(metrics.items())):
plt.subplot(1, len(metrics), i + 1)
plt.title(name)
plt.plot(*zip(*history))
plt.grid()
plt.show()
print("Mean loss=%.3f" % np.mean(metrics['train_loss'][-10:], axis=0)[1], flush=True)
# Note: it's okay if bleu oscillates up and down as long as it gets better on average over long term (e.g. 5k batches)
assert np.mean(metrics['dev_bleu'][-10:], axis=0)[1] > 15, "We kind of need a higher bleu BLEU from you. Kind of right now."
for inp_line, trans_line in zip(dev_inp[::500], model.translate_lines(dev_inp[::500])[0]):
print(inp_line)
print(trans_line)
print()
```
### Your Attention Required
In this section we want you to improve over the basic model by implementing a simple attention mechanism.
This is gonna be a two-parter: building the __attention layer__ and using it for an __attentive seq2seq model__.
### Attention layer (3 points)
Here you will have to implement a layer that computes a simple additive attention:
Given encoder sequence $ h^e_0, h^e_1, h^e_2, ..., h^e_T$ and a single decoder state $h^d$,
* Compute logits with a 2-layer neural network
$$a_t = linear_{out}(tanh(linear_{e}(h^e_t) + linear_{d}(h_d)))$$
* Get probabilities from logits,
$$ p_t = {{e ^ {a_t}} \over { \sum_\tau e^{a_\tau} }} $$
* Add up encoder states with probabilities to get __attention response__
$$ attn = \sum_t p_t \cdot h^e_t $$
You can learn more about attention layers in the lecture slides or [from this post](https://distill.pub/2016/augmented-rnns/).
```
class AttentionLayer(nn.Module):
def __init__(self, name, enc_size, dec_size, hid_size, activ=torch.tanh):
""" A layer that computes additive attention response and weights """
super().__init__()
self.name = name
self.enc_size = enc_size # num units in encoder state
self.dec_size = dec_size # num units in decoder state
self.hid_size = hid_size # attention layer hidden units
self.activ = activ # attention layer hidden nonlinearity
# create trainable paramteres like this:
self.<PARAMETER_NAME> = nn.Parameter(<INITIAL_VALUES>, requires_grad=True)
<...> # you will need a couple of these
def forward(self, enc, dec, inp_mask):
"""
Computes attention response and weights
:param enc: encoder activation sequence, float32[batch_size, ninp, enc_size]
:param dec: single decoder state used as "query", float32[batch_size, dec_size]
:param inp_mask: mask on enc activatons (0 after first eos), float32 [batch_size, ninp]
:returns: attn[batch_size, enc_size], probs[batch_size, ninp]
- attn - attention response vector (weighted sum of enc)
- probs - attention weights after softmax
"""
# Compute logits
<...>
# Apply mask - if mask is 0, logits should be -inf or -1e9
# You may need torch.where
<...>
# Compute attention probabilities (softmax)
probs = <...>
# Compute attention response using enc and probs
attn = <...>
return attn, probs
```
### Seq2seq model with attention (3 points)
You can now use the attention layer to build a network. The simplest way to implement attention is to use it in decoder phase:

_image from distill.pub [article](https://distill.pub/2016/augmented-rnns/)_
On every step, use __previous__ decoder state to obtain attention response. Then feed concat this response to the inputs of next attention layer.
The key implementation detail here is __model state__. Put simply, you can add any tensor into the list of `encode` outputs. You will then have access to them at each `decode` step. This may include:
* Last RNN hidden states (as in basic model)
* The whole sequence of encoder outputs (to attend to) and mask
* Attention probabilities (to visualize)
_There are, of course, alternative ways to wire attention into your network and different kinds of attention. Take a look at [this](https://arxiv.org/abs/1609.08144), [this](https://arxiv.org/abs/1706.03762) and [this](https://arxiv.org/abs/1808.03867) for ideas. And for image captioning/im2latex there's [visual attention](https://arxiv.org/abs/1502.03044)_
```
class AttentiveModel(BasicModel):
def __init__(self, name, inp_voc, out_voc,
emb_size=64, hid_size=128, attn_size=128):
""" Translation model that uses attention. See instructions above. """
nn.Module.__init__(self) # initialize base class to track sub-layers, trainable variables, etc.
self.inp_voc, self.out_voc = inp_voc, out_voc
self.hid_size = hid_size
# <YOUR CODE: initialize layers>
def encode(self, inp, **flags):
"""
Takes symbolic input sequence, computes initial state
:param inp: matrix of input tokens [batch, time]
:return: a list of initial decoder state tensors
"""
# encode input sequence, create initial decoder states
# <YOUR CODE>
# apply attention layer from initial decoder hidden state
first_attn_probas = <...>
# Build first state: include
# * initial states for decoder recurrent layers
# * encoder sequence and encoder attn mask (for attention)
# * make sure that last state item is attention probabilities tensor
first_state = [<...>, first_attn_probas]
return first_state
def decode_step(self, prev_state, prev_tokens, **flags):
"""
Takes previous decoder state and tokens, returns new state and logits for next tokens
:param prev_state: a list of previous decoder state tensors
:param prev_tokens: previous output tokens, an int vector of [batch_size]
:return: a list of next decoder state tensors, a tensor of logits [batch, n_tokens]
"""
# <YOUR CODE HERE>
return [new_dec_state, output_logits]
```
### Training attentive model
Please reuse the infrastructure you've built for the regular model. I hope you didn't hard-code anything :)
```
<YOUR CODE: create AttentiveModel and training utilities>
<YOUR CODE: training loop>
<YOUR CODE: measure final BLEU>
```
### Visualizing model attention (2 points)
After training the attentive translation model, you can check it's sanity by visualizing its attention weights.
We provided you with a function that draws attention maps using [`Bokeh`](https://bokeh.pydata.org/en/latest/index.html). Once you managed to produce something better than random noise, please save at least 3 attention maps and __submit them to anytask__ alongside this notebook to get the max grade. Saving bokeh figures as __cell outputs is not enough!__ (TAs can't see saved bokeh figures in anytask). You can save bokeh images as screenshots or using this button:

__Note:__ you're not locked into using bokeh. If you prefer a different visualization method, feel free to use that instead of bokeh.
```
import bokeh.plotting as pl
import bokeh.models as bm
from bokeh.io import output_notebook, show
output_notebook()
def draw_attention(inp_line, translation, probs):
""" An intentionally ambiguous function to visualize attention weights """
inp_tokens = inp_voc.tokenize(inp_line)
trans_tokens = out_voc.tokenize(translation)
probs = probs[:len(trans_tokens), :len(inp_tokens)]
fig = pl.figure(x_range=(0, len(inp_tokens)), y_range=(0, len(trans_tokens)),
x_axis_type=None, y_axis_type=None, tools=[])
fig.image([probs[::-1]], 0, 0, len(inp_tokens), len(trans_tokens))
fig.add_layout(bm.LinearAxis(axis_label='source tokens'), 'above')
fig.xaxis.ticker = np.arange(len(inp_tokens)) + 0.5
fig.xaxis.major_label_overrides = dict(zip(np.arange(len(inp_tokens)) + 0.5, inp_tokens))
fig.xaxis.major_label_orientation = 45
fig.add_layout(bm.LinearAxis(axis_label='translation tokens'), 'left')
fig.yaxis.ticker = np.arange(len(trans_tokens)) + 0.5
fig.yaxis.major_label_overrides = dict(zip(np.arange(len(trans_tokens)) + 0.5, trans_tokens[::-1]))
show(fig)
inp = dev_inp[::500]
trans, states = model.translate_lines(inp)
# select attention probs from model state (you may need to change this for your custom model)
# attention_probs below must have shape [batch_size, translation_length, input_length], extracted from states
# e.g. if attention probs are at the end of each state, use np.stack([state[-1] for state in states], axis=1)
attention_probs = <YOUR CODE>
for i in range(5):
draw_attention(inp[i], trans[i], attention_probs[i])
# Does it look fine already? don't forget to save images for anytask!
```
__Note 1:__ If the attention maps are not iterpretable, try starting encoder from zeros (instead of dec_start), forcing model to use attention.
__Note 2:__ If you're studying this course as a YSDA student, please submit __attention screenshots__ alongside your notebook.
## Bonus: goind deeper (2++ points each)
We want you to find the best model for the task. Use everything you know.
* different recurrent units: rnn/gru/lstm; deeper architectures
* bidirectional encoder, different attention methods for decoder (additive, dot-product, multi-head)
* word dropout, training schedules, anything you can imagine
* replace greedy inference with beam search
For a better grasp of seq2seq We recommend you to conduct at least one experiment from one of the bullet-points or your alternative ideas. As usual, describe what you tried and what results you obtained in a short report.
`[your report/log here or anywhere you please]`
| github_jupyter |
# CLX DGA Detection
This is an introduction to CLX DGA Detection.
## What is DGA Detection?
[Domain Generation Algorithms](https://en.wikipedia.org/wiki/Domain_generation_algorithm) (DGAs) are used to generate domain names that can be used by the malware to communicate with the command and control servers. IP addresses and static domain names can be easily blocked, and a DGA provides an easy method to generate a large number of domain names and rotate through them to circumvent traditional block lists.
## When to use CLX DGA Detection?
Use CLX DGA Detection to build your own DGA Detection model that can then be used to predict whether a given domain is malicious or not. We will use a type of recurrent neural network called the Gated Recurrent Unit (GRU) for this example. The [CLX](https://github.com/rapidsai/clx) and [RAPIDS](https://rapids.ai/) libraries enable users train their models with up-to-date domain names representative of both benign and DGA generated strings. Using a [CLX workflow](./intro-clx-workflow.ipynb), this capability could also be used in production environments.
**For a more advanced, in-depth example of CLX DGA Detection view this Jupyter** [notebook](https://github.com/rapidsai/clx/blob/main/notebooks/dga_detection/DGA_Detection.ipynb).
## How to train a CLX DGA Detection model
To train a CLX DGA Detection model you simply need a training data set which contains a column of domains and their associated `type` which can be either `1` (benign) or `0` (malicious).
First initialize your new model
```
LR = 0.001
N_LAYERS = 3
CHAR_VOCAB = 128
HIDDEN_SIZE = 100
N_DOMAIN_TYPE = 2 # Will be 2 since there are a total of 2 different types
from clx.analytics.dga_detector import DGADetector
dd = DGADetector(lr=LR)
dd.init_model(
n_layers=N_LAYERS,
char_vocab=CHAR_VOCAB,
hidden_size=HIDDEN_SIZE,
n_domain_type=N_DOMAIN_TYPE,
)
```
Next, train your DGA detector. The below example uses a small dataset for demonstration only. Ideally you will want a larger training set.
To develop a more expansive training set, these resources are available:
* DGA : http://osint.bambenekconsulting.com/feeds/dga-feed.txt
* Benign : http://s3.amazonaws.com/alexa-static/top-1m.csv.zip
```
import cudf
train_df = cudf.DataFrame()
train_df["domain"] = [
"google.com",
"youtube.com",
"tmall.com",
"duiwlqeejymdb.com",
"kofsmyaiufarb.net",
"xskphhmrlcihr.biz",
"yahoo.com",
"linkedin.com",
"twitter.com",
"wejaecjhycwss.co.uk",
"xtorhktvpblmr.info",
"xvljisbfalkts.com",
]
train_df["type"] = [1, 1, 1, 0, 0, 0, 1, 1, 1, 0, 0, 0]
```
When we train a model, the total loss is returned
```
dd.train_model(train_df['domain'], train_df['type'])
```
Ideally, you will want to train your model over a number of `epochs` as detailed in our example DGA Detection [notebook](https://github.com/rapidsai/clx/blob/main/notebooks/dga_detection/DGA_Detection.ipynb).
## Save a trained model checkpoint
```
dd.save_checkpoint("clx_dga_classifier.pth")
```
## Load a model checkpoint
Let's create a new dga detector and load the saved model from above.
```
dga_detector = DGADetector(lr=0.001)
dga_detector.load_checkpoint("clx_dga_classifier.pth")
```
## DGA Inferencing
Use your new model to predict malicious domains
```
test_df = cudf.DataFrame()
test_df['domain'] = ['facebook.com','ylqblbltqkynb.net']
dga_detector.predict(test_df['domain'])
```
## Conclusion
DGA detector in CLX enables users to train their models for detection and also use existing models. This capability could also be used in conjunction with log parsing efforts if the logs contain domain names. DGA detection done with CLX and RAPIDS keeps data in GPU memory, removing unnecessary copy/converts and providing a 4X speed advantage over CPU only implementations. This is esepcially true with large batch sizes.
| github_jupyter |
## Tweet Emotion Recognition: Natural Language Processing with TensorFlow
---
Dataset: [Tweet Emotion Dataset](https://github.com/dair-ai/emotion_dataset)
---
## Task 1: Introduction
## Task 2: Setup and Imports
1. Installing Hugging Face's nlp package
2. Importing libraries
```
!pip install nlp
%matplotlib inline
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
import nlp
import random
def show_history(h):
epochs_trained = len(h.history['loss'])
plt.figure(figsize=(16, 6))
plt.subplot(1, 2, 1)
plt.plot(range(0, epochs_trained), h.history.get('accuracy'), label='Training')
plt.plot(range(0, epochs_trained), h.history.get('val_accuracy'), label='Validation')
plt.ylim([0., 1.])
plt.xlabel('Epochs')
plt.ylabel('Accuracy')
plt.legend()
plt.subplot(1, 2, 2)
plt.plot(range(0, epochs_trained), h.history.get('loss'), label='Training')
plt.plot(range(0, epochs_trained), h.history.get('val_loss'), label='Validation')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
def show_confusion_matrix(y_true, y_pred, classes):
from sklearn.metrics import confusion_matrix
cm = confusion_matrix(y_true, y_pred, normalize='true')
plt.figure(figsize=(8, 8))
sp = plt.subplot(1, 1, 1)
ctx = sp.matshow(cm)
plt.xticks(list(range(0, 6)), labels=classes)
plt.yticks(list(range(0, 6)), labels=classes)
plt.colorbar(ctx)
plt.show()
print('Using TensorFlow version', tf.__version__)
```
## Task 3: Importing Data
1. Importing the Tweet Emotion dataset
2. Creating train, validation and test sets
3. Extracting tweets and labels from the examples
```
dataset=nlp.load_dataset('emotion')
dataset
train=dataset['train']
val=dataset['validation']
test=dataset['test']
def get_tweet(data):
tweets=[x['text'] for x in data]
labels=[x['label'] for x in data]
return tweets,labels
tweets,labels=get_tweet(train)
tweets[0],labels[0]
```
## Task 4: Tokenizer
1. Tokenizing the tweets
```
from tensorflow.keras.preprocessing.text import Tokenizer
tokenizer=Tokenizer(num_words=1000,oov_token='<UNK>')
tokenizer.fit_on_texts(tweets)
tokenizer.texts_to_sequences([tweets[0]])
tweets[0]
```
## Task 5: Padding and Truncating Sequences
1. Checking length of the tweets
2. Creating padded sequences
```
lengths=[len(t.split(' ')) for t in tweets]
plt.hist(lengths,bins=len(set(lengths)))
plt.show()
maxlen=50
from tensorflow.keras.preprocessing.sequence import pad_sequences
def get_sequences(tokenizer,tweets):
sequences=tokenizer.texts_to_sequences(tweets)
padded=pad_sequences(sequences,truncating='post',padding='post',maxlen=maxlen)
return padded
padded_train_seq=get_sequences(tokenizer,tweets)
padded_train_seq[0]
```
## Task 6: Preparing the Labels
1. Creating classes to index and index to classes dictionaries
2. Converting text labels to numeric labels
```
classes=set(labels)
print(classes)
plt.hist(labels,bins=11)
plt.show()
class_to_index=dict((c,i) for i,c in enumerate(classes))
index_to_class=dict((v,k)for k,v in class_to_index.items())
class_to_index
index_to_class
names_to_ids=lambda labels:np.array([class_to_index.get(x) for x in labels])
train_labels=names_to_ids(labels)
print(train_labels[0])
```
## Task 7: Creating the Model
1. Creating the model
2. Compiling the model
```
model=tf.keras.models.Sequential([
tf.keras.layers.Embedding(1000,16,input_length=maxlen),
tf.keras.layers.Bidirectional(tf.keras.layers.LSTM(20,return_sequences=True)),
tf.keras.layers.Bidirectional(tf.keras.layers.LSTM(20)),
tf.keras.layers.Dense(6,activation='softmax')
])
model.compile(
loss='sparse_categorical_crossentropy',
optimizer='adam',
metrics=['accuracy']
)
model.summary()
```
## Task 8: Training the Model
1. Preparing a validation set
2. Training the model
```
val_tweets,val_labels=get_tweet(val)
val_seq=get_sequences(tokenizer,val_tweets)
val_labels=names_to_ids(val_labels)
val_tweets[0],val_labels[0]
h=model.fit(
padded_train_seq,train_labels,
validation_data=(val_seq,val_labels),
epochs=20,
callbacks=[
tf.keras.callbacks.EarlyStopping(monitor='val_accuracy',patience=2)
]
)
```
## Task 9: Evaluating the Model
1. Visualizing training history
2. Prepraring a test set
3. A look at individual predictions on the test set
4. A look at all predictions on the test set
```
show_history(h)
test_tweets,test_labels=get_tweet(test)
test_seq=get_sequences(tokenizer,test_tweets)
test_labels=names_to_ids(test_labels)
_=model.evaluate(test_seq,test_labels)
i=random.randint(0,len(test_labels)-1)
print('Sentence:',test_tweets[i])
print('Emotion:',index_to_class[test_labels[i]])
p=model.predict(np.expand_dims(test_seq[i],axis=0))[0]
pred_class=index_to_class[np.argmax(p).astype('uint8')]
print('Predicted Emotion:',pred_class)
preds=model.predict_classes(test_seq)
show_confusion_matrix(test_labels,preds,list(classes))
```
| github_jupyter |
# Molecule Objects in <span style="font-variant: small-caps"> Psi4 </span>
This tutorial provides an overview on creating and manipulating molecule objects in <span style='font-variant: small-caps'> Psi4</span>, illustrated with an example parameterization of the Lennard-Jones potential for Helium dimer.
In order to use <span style="font-variant: small-caps"> Psi4 </span> within a Julia environment, we may install PyCall
executing `using Pkg; Pkg.add("PyCall")`. We point PyCall to the Python installation that has <span style="font-variant: small-caps"> Psi4 </span> installed, that is `ENV["PYTHON"] = ~/miniconda3/envs/p4env/bin/python`, rebuild PyCall, `using Pkg; Pkg.build("PyCall")` and restart Julia. The next time we start Julia we may import <span style="font-variant: small-caps"> Psi4 </span> with pyimport:
(check [here](https://github.com/dgasmith/psi4numpy/blob/master/README.md) for more details)
```
using PyCall: pyimport
psi4 = pyimport("psi4")
```
Unlike in <span style="font-variant: small-caps"> Psi4 </span> input files, defining a molecule in Julia is done by passing the molecular coordinates as a triple-quoted string to the [`psi4.geometry()`](http://psicode.org/psi4manual/master/api/psi4.driver.geometry.html#psi4.driver.geometry "API Details") function:
```
he = psi4.geometry("""
He
""")
```
Here, not only does the variable `he` refer to the helium molecule, but also an instance of the [`psi4.core.Molecule`](http://psicode.org/psi4manual/master/psi4api.html#psi4.core.Molecule "Go to API")
class in <span style='font-variant: small-caps'> Psi4</span>; this will be discussed in more detail later. For a more
complicated system than an isolated atom, the coordinates can be given in Cartesian or Z-Matrix formats:
```
h2o = psi4.geometry("""
O
H 1 0.96
H 1 0.96 2 104.5
""")
```
Information like the molecular charge, multiplicity, and units are assumed to be 0, 1, and Angstroms, respectively, if not specified within the molecule definition. This is done by adding one or more [molecule keywords](http://psicode.org/psi4manual/master/psithonmol.html#molecule-keywords "Go to Documentation") to the geometry string used to define the molecule. Additionally, <span style="font-variant: small-caps"> Psi4 </span> can detect molecular symmetry, or it can be specified manually. For example, to define a doublet water cation in $C_{2V}$ symmetry using lengths in Bohr,
```
doublet_h2o_cation = psi4.geometry("""
1 2
O
H 1 1.814
H 1 1.814 2 104.5
units bohr
symmetry c2v
""")
```
where the line `1 2` defines the charge and multiplicity, respectively. For systems of non-bonded fragments, the coordinates of each fragment are separated by a double-hyphen `"--"`; this allows for one fragment to be defined with Cartesian and another to be defined with Z-Matrix. For example, the hydronium-benzene complex can be defined with:
```
hydronium_benzene = psi4.geometry("""
0 1
C 0.710500000000 -0.794637665924 -1.230622098778
C 1.421000000000 -0.794637665924 0.000000000000
C 0.710500000000 -0.794637665924 1.230622098778
C -0.710500000000 -0.794637665924 1.230622098778
H 1.254500000000 -0.794637665924 -2.172857738095
H -1.254500000000 -0.794637665924 2.172857738095
C -0.710500000000 -0.794637665924 -1.230622098778
C -1.421000000000 -0.794637665924 0.000000000000
H 2.509000000000 -0.794637665924 0.000000000000
H 1.254500000000 -0.794637665924 2.172857738095
H -1.254500000000 -0.794637665924 -2.172857738095
H -2.509000000000 -0.794637665924 0.000000000000
--
1 1
X 1 CC 3 30 2 A2
O 13 R 1 90 2 90
H 14 OH 13 TDA 1 0
H 14 OH 15 TDA 13 A1
H 14 OH 15 TDA 13 -A1
CC = 1.421
CH = 1.088
A1 = 120.0
A2 = 180.0
OH = 1.05
R = 4.0
units angstrom
""")
```
For non-bonded fragments, the charge and multiplicity should be given explicitly for each fragment. If not, the
charge and multiplicity given (or inferred) for the first fragment is assumed to be the same for all fragments. In
addition to defining the coordinates outright, we have used variables within the geometry specification strings to
define bond lengths, angles, and dihedrals in the molecule. Similarly, we could define the X, Y, or Z Cartesian
coordinate for any atom in our molecule.
In order to define these variables after the molecule is built, as opposed to within the geometry specification
itself, there are several ways to do so; one of which will be illustrated in the Lennard-Jones potential example
below.
When a Psi4 molecule is first built using ``psi4.geometry()``, it is in an unfinished state, as a user may wish to
tweak the molecule. This can be solved by calling [``psi4.Molecule.update_geometry()``](http://psicode.org/psi4manual/master/psi4api.html#psi4.core.Molecule.update_geometry "Go to API"). This will update the molecule and restore sanity
to chemistry.
```
h2cch2 = psi4.geometry("""
H
C 1 HC
H 2 HC 1 A1
C 2 CC 3 A1 1 D1
H 4 HC 2 A1 1 D1
H 4 HC 2 A1 1 D2
HC = 1.08
CC = 1.4
A1 = 120.0
D1 = 180.0
D2 = 0.0
""")
println("Ethene has $(h2cch2.natom()) atoms")
h2cch2.update_geometry()
println("Ethene has $(h2cch2.natom()) atoms")
```
Finally, one can obtain useful information from a molecule by invoking one of several [`psi4.core.Molecule`](http://psicode.org/psi4manual/master/psi4api.html#psi4.core.Molecule "Go to Documentation") class methods on the molecule of interest. For example, if we were interested in verifying that our doublet water cation from above is, in fact, a doublet, we could invoke
~~~python
doublet_h2o_cation.multiplicity()
~~~
Below, some useful Molecule class methods are tabulated; please refer to the documentation for more details.
| Method | Description |
|--------|-------------|
| [center_of_mass()](http://psicode.org/psi4manual/master/psi4api.html#psi4.core.Molecule.center_of_mass "Go to Documentation") | Computes center of mass of molecule |
| [molecular_charge()](http://psicode.org/psi4manual/master/psi4api.html#psi4.core.Molecule.molecular_charge "Go to Documentation") | Gets the molecular charge |
| [multiplicity()](http://psicode.org/psi4manual/master/psi4api.html#psi4.core.Molecule.multiplicity "Go to Documentation") | Gets the total multiplicity |
| [nuclear_repulsion_energy()](http://psicode.org/psi4manual/master/psi4api.html#psi4.core.Molecule.nuclear_repulsion_energy "Go to Documentation") | Computes the nuclear repulsion energy of the molecule |
## Example: Fitting Lennard-Jones Parameters from Potential Energy Scan
In this example, we will compute and fit a potential energy curve for the Helium dimer. To begin with, let's create a string representation for our He dimer in Z-Matrix format, with the variable `**R**` representing the distance between the He atoms. The stars surrounding the ``R`` aren't any special syntax, just a convenient marker for future substitution.
```
# Define He Dimer
he_dimer = """
He
--
He 1 **R**
"""
```
Now we can build a series of dimers with the He atoms at different separations, and compute the energy at each point:
```
distances = [2.875, 3.0, 3.125, 3.25, 3.375, 3.5, 3.75, 4.0, 4.5, 5.0, 6.0, 7.0]
energies = []
for d in distances
# Build a new molecule at each separation
mol = psi4.geometry(replace(he_dimer,"**R**" => string(d)))
# Compute the Counterpoise-Corrected interaction energy
en = psi4.energy("MP2/aug-cc-pVDZ", molecule=mol, bsse_type="cp")
# Place in a reasonable unit, Wavenumbers in this case
en *= 219474.6
# Append the value to our list
push!(energies,en)
end
println("Finished computing the potential!")
```
Next, we can use the [NumPy](http://www.numpy.org/) library to fit a curve to these points along the potential scan. In this case, we will fit a Lennard-Jones potential.
```
np = pyimport("numpy")
# Fit data in least-squares way to a -12, -6 polynomial
powers = [-12, -6]
x = np.power(np.reshape(np.array(distances),(-1, 1)), powers)
coeffs = np.linalg.lstsq(x, energies)[1]
# Build list of points
fpoints = np.reshape(np.linspace(2, 7, 50),(-1, 1))
fdata = np.power(fpoints, powers)
fit_energies = np.dot(fdata, coeffs)
```
To visualize our results, we can use the [Matplotlib](http://matplotlib.org/) library. Since we're working in a Jupyter notebook, we can also use the `%matplotlib inline` "magic" command so that the graphs will show up in the notebook itself (check [here](https://ipython.org/ipython-doc/3/interactive/magics.html) for a comprehensive list of magic commands).
```
%matplotlib inline
import PyPlot
plt = PyPlot
plt.xlim((2, 7)) # X limits
plt.ylim((-7, 2)) # Y limits
plt.scatter(distances, energies) # Scatter plot of the distances/energies
plt.plot(fpoints, fit_energies) # Fit data
plt.plot([0,10], [0,0], "k-") # Make a line at 0
```
| github_jupyter |
<a href="https://colab.research.google.com/github/CanopySimulations/canopy-python-examples/blob/master/encrypting_config_components.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Upgrade Runtime
This cell ensures the runtime supports `asyncio` async/await, and is needed on Google Colab. If the runtime is upgraded, you will be prompted to restart it, which you should do before continuing execution.
```
!pip install "ipython>=7"
```
# Set Up Environment
### Import required libraries
```
!pip install -q canopy
import canopy
import logging
import nest_asyncio
logging.basicConfig(level=logging.INFO)
nest_asyncio.apply()
```
### Authenticate
```
authentication_data = canopy.prompt_for_authentication()
session = canopy.Session(authentication_data)
```
# Example: Encrypting config components
## Encrypting arbitrary data
The API exposes endpoints for encrypting and decrypting any JSON structure.
We can access these endpoints using the OpenAPI generated wrappers in the Canopy Python library.
In the example below we encrypt some arbitrary data:
```
# Created the OpenAPI generated wrapper for the config APIs.
config_api = canopy.openapi.ConfigApi(session.async_client)
# Create some arbitrary data.
initial_data = { 'hello': 'world' }
# Encrypt the data using the API.
encrypted_data = await config_api.config_encrypt(initial_data)
print(f'Encrypted: {encrypted_data}')
```
We can then decrypt it:
```
decrypted_data = await config_api.config_decrypt(encrypted_data)
print(f'Decrypted: {decrypted_data}')
```
**Encrypted data can only be decrypted by users within the same Canopy tenant (users who enter the same "company" name when logging in to the Canopy platform).**
Users in other tenants can use the encrypted data in their configs, but cannot decrypt it themselves.
This makes it a secure way to share car components with other Canopy tenants for simulation, without them being able to see the parameters. We can also hide sensitive output channels when certain areas of the car are encrypted.
## Encrypting part of a config
Only certain parts of a config can be encrypted, as the encrypted version must still conform to the schema. You can contact Canopy to discuss your encryption requirements.
In this example we will encrypt the electric motors of a car, which is an area that has already been enabled for encryption.
First we will fetch a default car and output its `electricMotors` section.
```
default_car = await canopy.load_default_config(session, 'car', 'Canopy F1 Car 2019')
print(default_car.data.powertrain.electric.electricMotors)
```
Next we will encrypt the `electricMotors` section.
```
encryption_result = await config_api.config_encrypt(default_car.data.powertrain.electric.electricMotors)
print(encryption_result)
```
Next we will replace the `electricMotors` section of the car with the encrypted version and save the car to the platform.
```
default_car.data.powertrain.electric.electricMotors = encryption_result
user_car_id = await canopy.create_config(
session,
'car',
'Python Example - Car with encrypted electric motors',
default_car.raw_data)
print(user_car_id)
```
And finally we will load the car again and output the `electricMotors` section, to show that it has been saved encrypted.
```
user_car = await canopy.load_config(session, user_car_id)
print(user_car.data.powertrain.electric.electricMotors)
```
| github_jupyter |
tgb - 8/18/2019 - The goal is to calculate smooth linear reponse functions about the base state used for Noah's paper and for 2 different neural networks:
1) One that exhibit stable behavior once coupled to CAM (Stable NN)
2) One that does not (Unstable NN)
We will take the following steps:
1) Perturb the base state with normally-distributed perturbations at each level
2) Calculate the linear response of each perturbed profile
3) Average the linear response to get the final linear response
4) Save the perturbed profiles and the perturbed response, as well as their average for Noah
5) Use eigenvalue analysis to see how the averaging affect the stability of the response. We will use 3 types of stability analysis:
5.1) Stability of the coupled (T,q) system, all normalized to energy units
5.2) Stability of the (q) system on its own
5.3) Stability of the (T,q) system reduced to a single q system via the WTG approximation (see Beucler and Cronin, 2018)
# 1) Initialization
Define imports
```
from cbrain.imports import *
from cbrain.data_generator import *
from cbrain.utils import *
from cbrain.normalization import *
from cbrain.models import fc_model
import matplotlib.pyplot as plt
from numpy import linalg as LA
import pickle
import scipy.integrate as sin
from tensorflow.python.ops.parallel_for.gradients import batch_jacobian
%matplotlib inline
```
Define neural networks
```
in_vars = ['QBP', 'TBP', 'VBP', 'PS', 'SOLIN', 'SHFLX', 'LHFLX']
out_vars = ['PHQ', 'TPHYSTND', 'FSNT', 'FSNS', 'FLNT', 'FLNS', 'PRECT']
scale_dict = load_pickle('../../nn_config/scale_dicts/002_pnas_scaling.pkl')
```
Define input and output transform
```
CAMDIR = '/scratch/05488/tg847872/revision_debug/'
input_transform = InputNormalizer(
xr.open_dataset(f'{CAMDIR}/001_norm.nc'),
in_vars,
'mean', 'maxrs')
output_transform = DictNormalizer(xr.open_dataset(CAMDIR+'001_norm.nc'),
out_vars, scale_dict)
```
tgb - 2/26/2020 - Use right norm file for unstable NN only
```
path_UNSTAB = '/project/meteo/w2w/A6/S.Rasp/SP-CAM/saved_models/007_32col_pnas_exact/'
InputNormalizer
pathdiv = path_UNSTAB+'inp_div.txt'
pathsub = path_UNSTAB+'inp_sub.txt'
File_div = open(pathdiv,"r")
input_div = File_div.read().split(',')
input_div[-1] = input_div[-1][:-1]
input_div = np.array(input_div).astype(float)
File_sub = open(pathsub,"r")
input_sub = File_sub.read().split(',')
input_sub[-1] = input_sub[-1][:-1]
input_sub = np.array(input_sub).astype(float)
```
tgb - 2/26/2020 - Load unstable NN only
```
from tensorflow.keras.models import Sequential, Model
from tensorflow.keras.layers import *
def fc_model(input_shape, output_shape, hidden_layers, conservation_layer=False,
inp_sub=None, inp_div=None, norm_q=None):
inp = Input(shape=(input_shape,))
# First hidden layer
x = Dense(hidden_layers[0])(inp)
x = LeakyReLU()(x)
# Remaining hidden layers
for h in hidden_layers[1:]:
x = Dense(h)(x)
x = LeakyReLU()(x)
# if conservation_layer:
# x = SurRadLayer(inp_sub, inp_div, norm_q)([inp, x])
# x = MassConsLayer(inp_sub, inp_div, norm_q)([inp, x])
# out = EntConsLayer(inp_sub, inp_div, norm_q)([inp, x])
else:
out = Dense(output_shape)(x)
return tf.keras.models.Model(inp, out)
unstabNN = fc_model(94, 65, [256]*9, 'LeakyReLU')
unstabNN.load_weights(
f'{path_UNSTAB}weights.h5'
)
```
On Stampede - Load stable and unstable NNs
```
model = fc_model(66, 65, [256]*9, 'LeakyReLU')
model.load_weights(
f'{CAMDIR}weights.h5'
)
STABLENN_path = '/home1/05823/tg851228/006_8col_pnas_exact/'
UNSTABNN_path = '/home1/05823/tg851228/007_32col_pnas_exact/'
stabNN = fc_model(94, 65, [256]*9, 'LeakyReLU')
stabNN.load_weights(
f'{STABLENN_path}weights.h5'
)
unstabNN = fc_model(94, 65, [256]*9, 'LeakyReLU')
unstabNN.load_weights(
f'{UNSTABNN_path}weights.h5'
)
```
# 2) Load and perturb Noah's tropical profiles
Load Noah's dataset for base tropical profile
```
path = '/home1/05823/tg851228/NC_DATA/2019-05-01-tropical_mean.nc'
noah_ds = xr.open_mfdataset(path, decode_times=False, concat_dim='time')
```
Load CAM dataset for coordinates used in CAM
```
cam_ds = xr.open_mfdataset(CAMDIR+ 'debug.cam2.h1.*.nc', decode_times=False, concat_dim='time')
```
Calculate rho and z from ideal gas law and hydrostasy
```
# Height [m]
Z_INTERP = np.interp(cam_ds.lev.values,np.fliplr(noah_ds.p.values)[0,:],
np.flip(noah_ds.z.values))
# Interface height [m]
ZI_INTERP = np.interp(cam_ds.ilev.values,np.fliplr(noah_ds.p.values)[0,:],
np.flip(noah_ds.z.values))
# Density [kg/m3]
RHO_INTERP = np.interp(cam_ds.lev.values,np.fliplr(noah_ds.p.values)[0,:],
np.flip(noah_ds.rho.values)[0,:])
# Specific humidity [kg/kg]
NNQBP = 1e-3*np.interp(cam_ds.lev.values,np.fliplr(noah_ds.p.values)[0,:],
np.fliplr(noah_ds.QV.values)[0,:])
# Temperature [K]
NNTBP = np.interp(cam_ds.lev.values,np.fliplr(noah_ds.p.values)[0,:],
np.fliplr(noah_ds.TABS.values)[0,:])
# Meridional velocity [m/s]
NNVBP = np.interp(cam_ds.lev.values,np.fliplr(noah_ds.p.values)[0,:],
np.fliplr(noah_ds.V.values)[0,:])
# Surface pressure [Pa]
NNPS = 1e2*noah_ds.p.values[:,0]
# Solar insolation [W/m2]
NNSOLIN = noah_ds.SOLIN.values
# Sensible heat flux [W/m2]
NNSHF = noah_ds.SHF.values
# Latent heat flux [W/m2]
NNLHF = noah_ds.LHF.values
```
Calculate rho and z from ideal gas law and hydrostasy
```
# Ideal gas law -> rho=p(R_d*T_v)
eps = 0.622 # Ratio of molecular weight(H2O)/molecular weight(dry air)
R_D = 287 # Specific gas constant of dry air in J/K/kg
r = NNQBP/(NNQBP**0-NNQBP)
Tv = NNTBP*(r**0+r/eps)/(r**0+r)
RHO = 1e2*cam_ds.lev.values/(R_D*Tv)
Z = -sin.cumtrapz(x=1e2*cam_ds.lev.values,y=1/(G*RHO))
Z = np.concatenate((np.zeros(1,),Z))
Z = Z-Z[-1]+Z_INTERP[-1]
```
For interface levels, first interpolate the temperature and specific humidity before infering the interface geopotential heights
```
qvI = 1e-3*np.interp(cam_ds.ilev.values,np.fliplr(noah_ds.p.values)[0,:],
np.fliplr(noah_ds.QV.values)[0,:])
TI = np.interp(cam_ds.ilev.values,np.fliplr(noah_ds.p.values)[0,:],
np.fliplr(noah_ds.TABS.values)[0,:])
rI = qvI/(qvI**0-qvI)
TvI = TI*(rI**0+rI/eps)/(rI**0+rI)
RHOI = 1e2*cam_ds.ilev.values/(R_D*TvI)
ZI = -sin.cumtrapz(x=1e2*cam_ds.ilev.values,y=1/(G*RHOI))
ZI = np.concatenate((np.zeros(1,),ZI))
ZI = ZI-ZI[-1]+ZI_INTERP[-1]
```
Save them in a vector called base state
```
base_state = {}
base_state['qv'] = NNQBP
base_state['T'] = NNTBP
base_state['v'] = NNVBP
base_state['ps'] = NNPS
base_state['S0'] = NNSOLIN
base_state['SHF'] = NNSHF
base_state['LHF'] = NNLHF
base_state['p'] = cam_ds.lev.values
base_state['p_interface'] = cam_ds.ilev.values
base_state['z'] = Z
base_state['z_interface'] = ZI
base_state['rho'] = RHO
base_state['rho_interface'] = RHOI
# Variables that we will perturb
profiles = {'qv','T','v'}
scalars = {'ps','S0','SHF','LHF'}
```
Perturb profiles and scalars
```
# We perturb each variable using a normal distribution of mean 0 and standard deviation 10%
Np = np.size(base_state['p']) # Number of vertical levels
Npert = 1000 # Number of perturbations
pert_state = {};
for i,profile in enumerate(profiles):
print('i=',i,' profile=',profile,' ',end='\r')
pert_state[profile] = np.zeros((Np,Npert))
for j,lev in enumerate(base_state['p']):
#print('j=',j,' level=',lev)
pert_state[profile][j,:] = base_state[profile][j]+\
np.random.normal(loc=0,scale=0.1,size=(Npert,))*np.tile(base_state[profile][j],(Npert,))
for i,scalar in enumerate(scalars):
print('i=',i,' scalar=',scalar,' ',end='\r')
pert_state[scalar] = base_state[scalar]+\
np.random.normal(loc=0,scale=0.1,size=(Npert,))*np.tile(base_state[scalar],(Npert,))
```
Check that the perturbation follows a normal distribution
```
plt.hist(pert_state['LHF'],bins=100)
plt.axvline(x=base_state['LHF'],color='k')
plt.hist(1e3*pert_state['qv'][28],bins=100)
plt.axvline(x=1e3*base_state['qv'][28],color='k')
```
# 3) Calculate the linear response of each perturbed profile, and compare the mean linear response function to the linear response function of the mean
Define the functions to get single jacobian and jacobian in a batch
```
def get_jacobian(x, model):
sess = tf.keras.backend.get_session()
jac = jacobian(model.output, model.input)
J = sess.run(jac, feed_dict={model.input: x.astype(np.float32)[None]})
return J.squeeze()
def get_batch_jacobian(x, model):
sess = tf.keras.backend.get_session()
jac = batch_jacobian(model.output, model.input)
J = sess.run(jac, feed_dict={model.input: x.astype(np.float32)})
return J.squeeze()
```
Create input vectors to feed to the neural network
```
in_vec = np.concatenate([base_state['qv'],
base_state['T'],
base_state['v'],
base_state['ps'],
base_state['S0'],
base_state['SHF'],
base_state['LHF'],
])[None, :].astype('float32')
in_vec_pert = np.concatenate([pert_state['qv'],
pert_state['T'],
pert_state['v'],
np.tile(pert_state['ps'],(1,1)),
np.tile(pert_state['S0'],(1,1)),
np.tile(pert_state['SHF'],(1,1)),
np.tile(pert_state['LHF'],(1,1)),
])[None, :].astype('float32')
in_vec_pert = np.transpose(in_vec_pert[0,:,:]) # Shape = [#batches,input size]
```
Define normalization for inputs and outputs
```
cf_inp = np.zeros((1,94))
for index in range (94):
if index<30: cf_inp[0,index]=L_V;
elif index<60: cf_inp[0,index]=C_P;
else: cf_inp[0,index]=1;
cf_oup = np.zeros((1,65))
for index in range (65):
if index<30: cf_oup[0,index]=L_V;
elif index<60: cf_oup[0,index]=C_P;
else: cf_oup[0,index]=1;
```
Use the get_batch_jacobian function to calculate linear responses about all perturbed states
```
Jstab = get_batch_jacobian(input_transform.transform(in_vec_pert),stabNN)*\
np.transpose(cf_oup/output_transform.scale)/\
(cf_inp*input_transform.div)
Junstab = get_batch_jacobian(input_transform.transform(in_vec_pert),unstabNN)*\
np.transpose(cf_oup/output_transform.scale)/\
(cf_inp*input_transform.div)
LRFstab = Jstab[:,:(2*Np),:(2*Np)] # Only keep the d(dq/dt,dT/dt)/d(q,T) Jacobian
LRFunstab = Junstab[:,:(2*Np),:(2*Np)] # Only keep the d(dq/dt,dT/dt)/d(q,T) Jacobian
LRFstab_mean = np.mean(LRFstab,axis=0)
LRFunstab_mean = np.mean(LRFunstab,axis=0)
```
Plot the mean linear response for both the stable and unstable cases
```
plt.rc('font', family='serif')
for ifig in range(2):
plt.figure(num=None,dpi=80, facecolor='w', edgecolor='k')
fig, ax = plt.subplots(figsize=(7.5,7.5))
if ifig==0: toplot = 24*3600*LRFstab_mean; case = 'Stable'
elif ifig==1: toplot = 24*3600*LRFunstab_mean; case = 'Unstable'
print('Case #',ifig+1,case)
cax = ax.matshow(toplot,cmap='bwr', vmin=-5, vmax=5)
x = np.linspace(0.,60.,100);
plt.plot(x,Np*x**0, color='k')
plt.plot(Np*x**0,x, color='k')
plt.xlim((0,2*Np)); plt.ylim((2*Np,0))
cbar = fig.colorbar(cax, pad = 0.1)
cbar.ax.tick_params(labelsize=20)
cbar.set_label(r'$\mathrm{Growth\ rate\ \left(1/day\right)}$',\
rotation=90, fontsize = 20)
plt.xticks(fontsize=20); plt.yticks(fontsize=20)
ax.xaxis.set_label_position('top')
X = plt.xlabel(r'Input level (QV left, T right)', fontsize = 20)
ax.xaxis.labelpad = 20
Y = plt.ylabel(r'Output level (QV_t top, T_t bottom)', fontsize = 20)
ax.yaxis.labelpad = 20
```
Compare to linear response of the mean
```
Jstab = get_batch_jacobian(input_transform.transform(in_vec),stabNN)*\
np.transpose(cf_oup/output_transform.scale)/\
(cf_inp*input_transform.div)
Junstab = get_batch_jacobian(input_transform.transform(in_vec),unstabNN)*\
np.transpose(cf_oup/output_transform.scale)/\
(cf_inp*input_transform.div)
LRFmean_stab = Jstab[:(2*Np),:(2*Np)] # Only keep the d(dq/dt,dT/dt)/d(q,T) Jacobian
LRFmean_unstab = Junstab[:(2*Np),:(2*Np)] # Only keep the d(dq/dt,dT/dt)/d(q,T) Jacobian
plt.rc('font', family='serif')
for ifig in range(2):
plt.figure(num=None,dpi=80, facecolor='w', edgecolor='k')
fig, ax = plt.subplots(figsize=(7.5,7.5))
if ifig==0: toplot = 24*3600*LRFmean_stab; case = 'Stable'
elif ifig==1: toplot = 24*3600*LRFmean_unstab; case = 'Unstable'
print('Case #',ifig+1,case)
cax = ax.matshow(toplot,cmap='bwr', vmin=-5, vmax=5)
x = np.linspace(0.,60.,100);
plt.plot(x,Np*x**0, color='k')
plt.plot(Np*x**0,x, color='k')
plt.xlim((0,2*Np)); plt.ylim((2*Np,0))
cbar = fig.colorbar(cax, pad = 0.1)
cbar.ax.tick_params(labelsize=20)
cbar.set_label(r'$\mathrm{Growth\ rate\ \left(1/day\right)}$',\
rotation=90, fontsize = 20)
plt.xticks(fontsize=20); plt.yticks(fontsize=20)
ax.xaxis.set_label_position('top')
X = plt.xlabel(r'Input level (QV left, T right)', fontsize = 20)
ax.xaxis.labelpad = 20
Y = plt.ylabel(r'Output level (QV_t top, T_t bottom)', fontsize = 20)
ax.yaxis.labelpad = 20
```
# 4) Save profiles and linear reponse functions for Noah
Save four linear response functions:
1) Mean LRF for stable NN
2) Mean LRF for unstable NN
3) LRF for stable NN about mean base state
4) LRF for unstable NN about mean base state
```
Name = ['MeanLRF_stable','MeanLRF_unstable','LRFMean_stable','LRFMean_unstable']
jac = {}
for ijac,name in enumerate(Name):
if ijac==0: tmp = LRFstab_mean
elif ijac==1: tmp = LRFunstab_mean
elif ijac==2: tmp = LRFmean_stab
elif ijac==3: tmp = LRFmean_unstab
jac[name] = {}
jac[name]['q'] = {}
jac[name]['T'] = {}
jac[name]['q']['q'] = tmp[:30,:30]
jac[name]['q']['T'] = tmp[:30,30:]
jac[name]['T']['q'] = tmp[30:,:30]
jac[name]['T']['T'] = tmp[30:,30:]
```
Take mean of pert_state (which should be close to base_state) for reference
```
pert_state_mean = {};
for i,profile in enumerate(profiles):
print('i=',i,' profile=',profile,' ',end='\r')
pert_state_mean[profile] = np.mean(pert_state[profile],axis=1)
for i,scalar in enumerate(scalars):
print('i=',i,' scalar=',scalar,' ',end='\r')
pert_state_mean[scalar] = np.mean(pert_state[scalar])
```
Save all LRFs, base state, and mean of perturbed states in an pkl file for Noah
```
path = '/home1/05823/tg851228/SPCAM/CBRAIN-CAM/notebooks/tbeucler_devlog/PKL_DATA/'
hf = open(path+'8_19_LRF.pkl','wb')
ForNoah = {"base_state" : base_state,
"mean_pert_state" : pert_state_mean,
"linear_response_functions" : jac}
pickle.dump(ForNoah,hf)
hf.close()
```
# 5) Eigenvalue analysis of LRFs
## 5.1) Coupled (T,q) system
Spectrum of all 4 LRFs
Of course, note that for the mean LRFs, one could also take the mean of the spectrum
```
TQ_sp = {}
plt.figure(figsize=(10,5))
plt.axhline(y=0,color='k')
for ijac,name in enumerate(Name):
if ijac==0: tmp = LRFstab_mean; ls = '-'
elif ijac==1: tmp = LRFunstab_mean; ls = '-'
elif ijac==2: tmp = LRFmean_stab; ls = '--'
elif ijac==3: tmp = LRFmean_unstab; ls = '--'
Eval, Evec = LA.eig(tmp)
lam = 24*3600*np.real(Eval)
TQ_sp[name] = [np.min(lam),np.max(lam)]
plt.plot(np.sort(lam),label=name,linestyle=ls)
plt.legend()
plt.xlabel('Index of eigenvalue')
plt.ylabel('Eigenvalue [1/day]')
```
From this graph, and given that we may trust the spectrum of the smooth mean LRFs more than the spectrum of the noisy LRF about the mean base state, we can infer that:
The instability may be caused by a lack of damping of multiple modes rather than an amplification of a single mode
## 5.2) (T,q) system coupled via WTG following Beucler and Cronin (2018)
Calculate ham coefficient for the base state
```
dq_dp = np.gradient(base_state['qv'],base_state['p'])
dT_dp = np.gradient(base_state['T'],base_state['p'])
dz_dp = np.gradient(base_state['z'],base_state['p'])
ham = -L_V*dq_dp/(C_P*dT_dp+G*dz_dp)
plt.plot(ham,base_state['p'])
plt.axvline(x=1,color='k')
plt.gca().invert_yaxis()
plt.xlabel('HAM [1]')
plt.ylabel('Pressure [hPa]')
def nearest_index(array, value):
idx = (np.abs(array-value)).argmin()
return idx
LS = ['-','-','--','--']
LRF_WTG = {}
WTG_sp = {}
i150 = nearest_index(base_state['p'],150)
i900 = nearest_index(base_state['p'],925)
for ijac,name in enumerate(Name):
LRF_WTG[name] = jac[name]['q']['q']+(ham*C_P/L_V)*jac[name]['q']['q']
Eval, Evec = LA.eig(LRF_WTG[name][i150:i900,i150:i900])
lam = 24*3600*np.real(Eval)
WTG_sp[name] = [np.min(lam),np.max(lam)]
plt.plot(np.sort(lam),label=name,linestyle=LS[ijac])
plt.legend()
plt.xlabel('Index of eigenvalue')
plt.ylabel('Eigenvalue [1/day]')
```
This eigenvalue analysis is consistent with the one for the full (T,q) system: It is the lack of damping of the "unstable NN" that may be causing the instability rather than the positive eigenvalues at the end of the spectrum. The positive eigenvalues are not present in this analysis, suggesting that the positive modes may have been either BL or upper-atmospheric modes that we have filtered out here.
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.