text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
### Parameter Extraction for SLO-2 Ion Channel
```
"""
Example of using cwFitter to generate a HH model for SLO-2 ion channel
Based on experimental data from doi:10.1083/jcb.200203055
"""
import os.path
import sys
import time
import numpy as np
import matplotlib.pyplot as plt
sys.path.append('..')
sys.path.append('../..')
sys.path.append('../../..')
from channelworm.fitter import *
userData = dict()
cwd=os.getcwd()
csv_path_VC_1 = os.path.dirname(cwd)+'/examples/slo-2-data/slo-2-VClamp/1.csv'
csv_path_VC_2 = os.path.dirname(cwd)+'/examples/slo-2-data/slo-2-VClamp/2.csv'
csv_path_VC_3 = os.path.dirname(cwd)+'/examples/slo-2-data/slo-2-VClamp/3.csv'
csv_path_VC_4 = os.path.dirname(cwd)+'/examples/slo-2-data/slo-2-VClamp/4.csv'
x_var_VC = {'type':'Time','unit':'ms','toSI':1e-3}
y_var_VC = {'type':'Current','unit':'nA','toSI':1e-9,'adjust':-0.82}
traces_VC = [{'vol':110e-3,'csv_path':csv_path_VC_1,'x_var':x_var_VC,'y_var':y_var_VC},
{'vol':40e-3,'csv_path':csv_path_VC_2,'x_var':x_var_VC,'y_var':y_var_VC},
{'vol':-140e-3,'csv_path':csv_path_VC_3,'x_var':x_var_VC,'y_var':y_var_VC},
{'vol':-90e-3,'csv_path':csv_path_VC_4,'x_var':x_var_VC,'y_var':y_var_VC}]
ref_VC = {'fig':'6a','doi':'10.1038/77670'}
VClamp = {'ref':ref_VC,'traces':traces_VC}
# #
# csv_path_VC = os.path.dirname(cwd)+'/examples/slo-2-data/SLO-2-2000-VClamp.csv'
# x_var_VC = {'type':'Time','unit':'ms','toSI':1e-3}
# y_var_VC = {'type':'Current','unit':'nA','toSI':1e-9,'adjust':-0.82}
# traces_VC = [{'csv_path':csv_path_VC,'x_var':x_var_VC,'y_var':y_var_VC}]
# ref_VC = {'fig':'6a','doi':'10.1038/77670'}
# VClamp = {'ref':ref_VC,'traces':traces_VC}
#
csv_path = os.path.dirname(cwd)+'/examples/slo-2-data/SLO-2-2000-IV.csv'
ref = {'fig':'6a','doi':'10.1038/77670'}
x_var = {'type':'Voltage','unit':'mV','toSI':1e-3}
y_var = {'type':'Current','unit':'pA','toSI':1e-12}
IV = {'ref':ref,'csv_path':csv_path,'x_var':x_var,'y_var':y_var}
csv_path_POV = os.path.dirname(cwd)+'/examples/slo-2-data/SLO-2-2000-GV.csv'
ref_POV = {'fig':'6b','doi':'10.1038/77670'}
x_var_POV = {'type':'Voltage','unit':'mV','toSI':1e-3}
y_var_POV = {'type':'G/Gmax','unit':'','toSI':1}
POV = {'ref':ref_POV,'csv_path':csv_path_POV,'x_var':x_var_POV,'y_var':y_var_POV}
# userData['samples'] = {'IV':IV,'POV':POV}
userData['samples'] = {'VClamp':VClamp}
# userData['samples'] = {'IV':IV,'POV':POV,'VClamp':VClamp}
# args = {'weight':{'start':20,'peak':10,'tail':30,'end':30,4:50}}
myInitiator = initiators.Initiator(userData)
sampleData = myInitiator.get_sample_params()
bio_params = myInitiator.get_bio_params()
sim_params = myInitiator.get_sim_params()
myEvaluator = evaluators.Evaluator(sampleData,sim_params,bio_params)
# bio parameters for SLO-2
bio_params['cell_type'] = 'Xenopus oocytes'
bio_params['channel_type'] = 'SLO-2'
bio_params['ion_type'] = 'K'
bio_params['val_cell_params'][0] = 200e-9 # C_mem DOI: 10.1074/jbc.M605814200
bio_params['val_cell_params'][1] = 20e-6 # area DOI: 10.1101/pdb.top066308
bio_params['gate_params'] = {'vda': {'power': 1}}
bio_params['channel_params'] = ['g_dens','e_rev']
bio_params['unit_chan_params'] = ['S/m2','V']
bio_params['min_val_channel'] = [1e-4,-5e-3]
bio_params['max_val_channel'] = [10, 5e-3]
bio_params['channel_params'].extend(['v_half_a','k_a','T_a'])
bio_params['unit_chan_params'].extend(['V','V','s'])
bio_params['min_val_channel'].extend([-0.15, 0.001, 0.0001])
bio_params['max_val_channel'].extend([ 0.15, 0.1, 0.01])
# Simulation parameters for SLO-2 VClamp and I/V
sim_params['v_hold'] = -110e-3
sim_params['I_init'] = 0
sim_params['pc_type'] = 'VClamp'
sim_params['deltat'] = 1e-5
sim_params['duration'] = 0.059
sim_params['start_time'] = 0.0029
sim_params['end_time'] = 0.059
sim_params['protocol_start'] = -140e-3
sim_params['protocol_end'] = 110e-3
sim_params['protocol_steps'] = 10e-3
opt = '-pso'
# opt = '-ga'
# opt = None
if len(sys.argv) == 2:
opt = sys.argv[1]
if ('IV' or 'POV') in sampleData and opt is not None:
while True:
q = raw_input("\n\nTry fitting curves (y,n):")
if q == "n":
break # stops the loop
elif q == "y":
# Find initial guess for parameters using curve_fit, leastsq
popt = None
best_candidate = np.asarray(bio_params['min_val_channel']) + np.asarray(bio_params['max_val_channel']) / 2
best_candidate_params = dict(zip(bio_params['channel_params'],best_candidate))
cell_var = dict(zip(bio_params['cell_params'],bio_params['val_cell_params']))
mySimulator = simulators.Simulator(sim_params,best_candidate_params,cell_var,bio_params['gate_params'])
bestSim = mySimulator.patch_clamp()
if 'IV' in sampleData:
popt , p0 = mySimulator.optim_curve(params= bio_params['channel_params'],
best_candidate= best_candidate,
target= [sampleData['IV']['V'],sampleData['IV']['I']])
print 'Params after IV minimization:'
print p0
IV_fit_cost = myEvaluator.iv_cost(popt)
print 'IV cost:'
print IV_fit_cost
if 'POV' in sampleData:
POV_fit_cost = myEvaluator.pov_cost(popt)
print 'POV cost:'
print POV_fit_cost
if 'VClamp' in sampleData:
VClamp_fit_cost = myEvaluator.vclamp_cost(popt)
print 'VClamp cost:'
print VClamp_fit_cost
vData = np.arange(-0.140, 0.110, 0.001)
Iopt = mySimulator.iv_act(vData,*popt)
plt.plot([x*1e3 for x in bestSim['V_ss']],bestSim['I_ss'], label = 'Initial parameters', color='y')
plt.plot([x*1e3 for x in sampleData['IV']['V']],sampleData['IV']['I'], '--ko', label = 'sample data')
plt.plot([x*1e3 for x in vData],Iopt, color='r', label = 'Fitted to IV curve')
plt.legend()
plt.title("IV Curve Fit")
plt.xlabel('V (mV)')
plt.ylabel('I (A)')
plt.show()
elif 'POV' in sampleData:
popt , p0 = mySimulator.optim_curve(params= bio_params['channel_params'],
best_candidate= best_candidate,
target= [sampleData['POV']['V'],sampleData['POV']['PO']],curve_type='POV')
print 'Params after POV minimization:'
print p0
POV_fit_cost = myEvaluator.pov_cost(popt)
print 'POV cost:'
print POV_fit_cost
if 'VClamp' in sampleData:
VClamp_fit_cost = myEvaluator.vclamp_cost(popt)
print 'VClamp cost:'
print VClamp_fit_cost
vData = np.arange(-0.140, 0.110, 0.001)
POopt = mySimulator.pov_act(vData,*popt)
plt.plot([x*1e3 for x in bestSim['V_PO_ss']],bestSim['PO_ss'], label = 'Initial parameters', color='y')
plt.plot([x*1e3 for x in sampleData['POV']['V']],sampleData['POV']['PO'], '--ko', label = 'sample data')
plt.plot([x*1e3 for x in vData],POopt, color='r', label = 'Fitted to G/Gmax vs V curve')
plt.legend()
plt.title("G/Gmax vs V Curve Fit")
plt.xlabel('V (mV)')
plt.ylabel('G/Gmax')
plt.show()
if popt is not None:
if opt == '-pso':
bio_params['min_val_channel'][0:4] = popt[0:4] - abs(popt[0:4]/2)
bio_params['max_val_channel'][0:4] = popt[0:4] + abs(popt[0:4]/2)
else:
bio_params['min_val_channel'][0:4] = popt[0:4]
bio_params['max_val_channel'][0:4] = popt[0:4]
best_candidate_params = dict(zip(bio_params['channel_params'],popt))
cell_var = dict(zip(bio_params['cell_params'],bio_params['val_cell_params']))
mySimulator = simulators.Simulator(sim_params,best_candidate_params,cell_var,bio_params['gate_params'])
bestSim = mySimulator.patch_clamp()
myModelator = modelators.Modelator(bio_params,sim_params)
myModelator.compare_plots(sampleData,bestSim,show=True)
myModelator.ss_plots(bestSim,show=True)
start = time.time()
if opt == '-ga':
ga_args = myInitiator.get_opt_params()
best_candidate, score = myEvaluator.ga_evaluate(min=bio_params['min_val_channel'],
max=bio_params['max_val_channel'],
args=ga_args)
elif opt == '-pso':
pso_args = myInitiator.get_opt_params(type='PSO')
pso_args['minstep'] = 1e-24
pso_args['minfunc'] = 1e-24
pso_args['swarmsize'] = 100
pso_args['maxiter'] = 100
best_candidate, score = myEvaluator.pso_evaluate(lb=bio_params['min_val_channel'],
ub=bio_params['max_val_channel'],
args=pso_args)
else:
# best_candidate = [2.678373586024887e-08, -0.004343196320916513, -0.15148699378068883, 0.04457177073153084, 0.0006512657782666903]
best_candidate = [2.6713432536911465e-08, -0.0043477407996737093, -0.077423632426596764, 0.030752484500400822, 0.0007076266889846564]
best_candidate[0] = 2.6713432536911465e-08 / bio_params['val_cell_params'][1]
secs = time.time()-start
print("----------------------------------------------------\n\n"
+"Ran in %f seconds (%f mins)\n"%(secs, secs/60.0))
best_candidate_params = dict(zip(bio_params['channel_params'],best_candidate))
cell_var = dict(zip(bio_params['cell_params'],bio_params['val_cell_params']))
mySimulator = simulators.Simulator(sim_params,best_candidate_params,cell_var,bio_params['gate_params'])
bestSim = mySimulator.patch_clamp()
myModelator = modelators.Modelator(bio_params,sim_params)
myModelator.compare_plots(sampleData,bestSim,show=True)
myModelator.ss_plots(bestSim,show=True)
print 'best candidate after optimization:'
print best_candidate_params
# Only for tau_max
if 'VClamp' in sampleData:
for trace in sampleData['VClamp']['traces']:
if 'vol' in trace:
if trace['vol'] is None:
pass
elif trace['vol'] == 110e-3:
end = sim_params['protocol_end']
start = sim_params['protocol_start']
sim_params['protocol_end'] = trace['vol']
sim_params['protocol_start'] = trace['vol']
x = np.asarray(trace['t'])
on = sim_params['start_time']
off = sim_params['end_time']
onset = np.abs(x-on).argmin()
offset = np.abs(x-off).argmin()
t_sample_on = trace['t'][onset+1:offset]
I_sample_on = trace['I'][onset+1:offset]
vcSim = simulators.Simulator(sim_params,best_candidate_params,cell_var,bio_params['gate_params'])
pcSim = vcSim.patch_clamp()
popt , p0 = vcSim.optim_curve(params= bio_params['channel_params'],
best_candidate= best_candidate,
target= [t_sample_on,I_sample_on],curve_type='VClamp')
vcEval = evaluators.Evaluator(sampleData,sim_params,bio_params)
print 'Params after VClamp minimization:'
print p0
if 'IV' in sampleData:
IV_fit_cost = vcEval.iv_cost(popt)
print 'IV cost:'
print IV_fit_cost
if 'POV' in sampleData:
POV_fit_cost = vcEval.pov_cost(popt)
print 'POV cost:'
print POV_fit_cost
# VClamp_fit_cost = vcEval.vclamp_cost(popt)
# print 'VClamp cost:'
# print VClamp_fit_cost
tData = np.arange(on, off, sim_params['deltat'])
Iopt = vcSim.patch_clamp(tData,*popt)
# plt.plot(pcSim['t'],pcSim['I'][0], label = 'Initial parameters', color='y')
plt.plot(t_sample_on,I_sample_on, '--ko', label = 'sample data')
plt.plot(tData,Iopt, color='r', label = 'Fitted to VClamp trace')
plt.legend()
plt.title('VClamp Curve Fit for holding potential %i (mV)'%(trace['vol']*1e3))
plt.xlabel('T (s)')
plt.ylabel('I (A)')
plt.show()
sim_params['protocol_end'] = end
sim_params['protocol_start'] = start
# best_candidate_params = dict(zip(bio_params['channel_params'],popt))
best_candidate_params['T_a'] = popt[4]
cell_var = dict(zip(bio_params['cell_params'],bio_params['val_cell_params']))
mySimulator = simulators.Simulator(sim_params,best_candidate_params,cell_var,bio_params['gate_params'])
bestSim = mySimulator.patch_clamp()
myModelator = modelators.Modelator(bio_params,sim_params)
myModelator.compare_plots(sampleData,bestSim,show=True)
myModelator.ss_plots(bestSim,show=True)
# Generate NeuroML2 file
model_params = {}
model_params['channel_name'] = 'SLO2'
model_params['channel_id'] = '4'
model_params['model_id'] = '1'
model_params['contributors'] = [{'name': 'Vahid Ghayoomi','email': 'vahidghayoomi@gmail.com'}]
model_params['references'] = [{'doi': '10.1038/77670',
'PMID': '10903569',
'citation': 'SLO-2, a K+ channel with an unusual Cl- dependence. '
'(Yuan A; Dourado M; Butler A; Walton N; Wei A; Salkoff L. Nat. Neurosci., 3(8):771-9)'}]
model_params['file_name'] = cwd+'/slo-2-data/SLO-2.channel.nml'
nml2_file = myModelator.generate_channel_nml2(bio_params,best_candidate_params,model_params)
run_nml_out = myModelator.run_nml2(model_params['file_name'])
```
| github_jupyter |
# Procesado de votos
Este script modela la rama real-time (es decir, la única rama que hay) de la aplicación. Spark nos genera un dataframe continuo sobre el que podemos aplicar filtros y expresiones como en un dataframe normal, con la salvedad de que el resultado no se devuelve en el mismo instante de ejecutar un `df.show()` o `df.write()`, sino que debemos decidir en el modo de escritura con `.outputMode()`. Además, no todos los formatos de escritura soportan todos los modos de salida: https://spark.apache.org/docs/2.2.0/structured-streaming-programming-guide.html#basic-concepts y https://spark.apache.org/docs/2.2.0/structured-streaming-programming-guide.html#output-modes.
```
from pyspark.sql import SparkSession
from pyspark.sql.functions import explode
from pyspark.sql.functions import split
from pyspark.sql.functions import from_json
from pyspark.sql.functions import col
import pyspark.sql.functions as fn
from pyspark.sql.types import StructType, StructField, StringType, IntegerType
PACKAGES = "org.apache.spark:spark-sql-kafka-0-10_2.11:2.4.0"
spark = SparkSession \
.builder \
.appName("StructuredVotes") \
.config("spark.jars.packages", PACKAGES)\
.getOrCreate()
from ejercicios.votes import TOPIC_VOTES, TOPIC_VOTES_ENRICHED
```
Obtenemos el dataframe de manera similar a los dataframes de orígenes estáticos: indicamos el origen, Kafka, y varios parámetros de configuración: la dirección del broker, el topic y el offset inicial: https://spark.apache.org/docs/latest/structured-streaming-kafka-integration.html#creating-a-kafka-source-for-batch-queries. Debemos subscribirnos desde el offset más antiguo para poder recalcular el resultado sobre todos los votos recibidos.
```
df = spark \
.readStream \
.format("kafka") \
.option("kafka.bootstrap.servers", "localhost:9092") \
.option("startingOffsets", "earliest") \
.option("subscribe", TOPIC_VOTES_ENRICHED) \
.load()
```
Los mensajes llegan en formato JSON, pero al contrario que con `spark.read.csv`, debemos indicar el esquema completo.
```
schema = StructType([
StructField("CODIGO", IntegerType()),
StructField("COMUNIDAD", StringType()),
StructField("PROVINCIA", StringType()),
StructField("MUNICIPIO", StringType()),
StructField("PARTIDO", StringType())
])
```
Podemos usar el modo de salida por consola para hacer debugging. Si usamos el modo de salida `append`, en cada microbatch sólo se imprimen los mensajes procesados en ese batch. For formato por consola no soporta modo `complete`.
```
query = df \
.selectExpr("topic", "CAST(key AS STRING)", "CAST(value AS STRING) AS value") \
.withColumn("value_json", fn.from_json(col('value'), schema)) \
.select('topic', 'value_json') \
.writeStream \
.outputMode("append") \
.format("console") \
.start()
```
Al contrario que los clientes `Consumer` de otros ejercicios, `query.start()` arranca la query en el cluster de Spark sin bloquear el proceso python. Podemos pararla expresamente con `query.stop()` o bloquear el proceso hasta que termine la query con `query.awaitTermination()`.
```
query.stop()
query.awaitTermination()
```
También podemos volcar los datos en una vista SQL en memoria. Esta vista se puede leer como un dataframe por otros procesos de Spark.
```
query = df \
.selectExpr("topic", "CAST(key AS STRING)", "CAST(value AS STRING) AS value") \
.withColumn("value_json", fn.from_json(col('value'), schema)) \
.select('topic', 'value_json') \
.writeStream \
.outputMode("append") \
.format("memory") \
.queryName('preview') \
.start()
spark.sql('SELECT * FROM preview').show()
```
La tabla de destino y su dataframe asociada tiene un esquema, al igual que cualquier otro dataframe. La función `to_json` genera una columna de tipo complejo que podemos simplificar con una operación select.
```
spark.sql('SELECT * FROM preview').printSchema()
query = df \
.selectExpr("topic", "CAST(key AS STRING)", "CAST(value AS STRING) AS value") \
.withColumn("value_json", fn.from_json(col('value'), schema)) \
.select('value_json.COMUNIDAD', 'value_json.PROVINCIA', 'value_json.PARTIDO') \
.writeStream \
.outputMode("append") \
.format("memory") \
.queryName('preview2') \
.start()
spark.sql('SELECT * FROM preview2').show()
query.stop()
```
Tras leer el mensaje, una de las tareas es validar la _firma del mensaje_. El enunciado nos indica que asumamos que la firma digital está incluida en el mensaje y podemos hacer uso de una función que devuelva un booleano si la firma es satisfactoria. Como ya sabemos usar [UDFs](http://spark.apache.org/docs/latest/api/python/pyspark.sql.html#pyspark.sql.functions.udf), podemos escribir la función en python y convertirla a UDF:
```
def process_signature(comunidad, provincia, municipio):
return 'OK'
udf_process_signature = fn.udf(process_signature)
```
Ya estamos en disposición de agregar los resultados por comunidad autónoma y provincia. Al ejecutar una agregación debemos cambiar al modo de salida `complete` o `update`. El formato `memory` no soporta `update`, así que actualizaremos la tabla completa en cada iteración.
```
query = df \
.selectExpr("topic", "CAST(key AS STRING)", "CAST(value AS STRING) AS value") \
.withColumn("value_json", fn.from_json(col('value'), schema)) \
.select('value_json.COMUNIDAD', 'value_json.PROVINCIA', 'value_json.MUNICIPIO', 'value_json.PARTIDO') \
.withColumn('SIGNATURE', udf_process_signature(col('COMUNIDAD'), col('PROVINCIA'), col('MUNICIPIO'))) \
.where(~ fn.isnull(col('SIGNATURE'))) \
.groupBy('COMUNIDAD', 'PROVINCIA', 'PARTIDO') \
.agg(fn.count('*').alias('VOTOS')) \
.sort(col('COMUNIDAD').asc(), col('PROVINCIA').asc(), col('VOTOS').desc()) \
.writeStream \
.outputMode("complete") \
.format("memory") \
.queryName('dashboard') \
.start()
```
# Resultados
Sobre esta tabla podemos aplicar más filtros y podríamos usarla como origen de datos para escribir en un fichero externo (por ejemplo, al terminar las votaciones) o para mostrar gráficos con la composición de escaños.
```
spark.sql("""
SELECT COMUNIDAD, PARTIDO, sum(VOTOS) as VOTOS
FROM dashboard
WHERE VOTOS > 2 and COMUNIDAD LIKE 'And%'
GROUP BY COMUNIDAD, PARTIDO
ORDER BY VOTOS DESC
""").show(100, False)
query.stop()
```
# Arquitectura Kappa: cambio de core
El enunciado nos indica que a mitad de jornada se detecta que en una de las provincias hay problemas técnicos y la firma de los votos puede haber sido manipulada. Debemos recalcular los votos sin detener el sistema de recepción, ya que el resto de mesas electorales deben seguir funcionando.
En este caso podemos calcular una segunda tabla en la que descartamos todos los votos de la provincia afectada. Sólo es necesario cambiar la lógica de la función que valida las firmas. En un caso real reescribiríamos el código y arrancaríamos un segundo proceso (o contenedor, o llamada a pyspark-submit). En este caso podemos ejecutar la query desde el mismo notebook por simplificar.
Si la provincia afectada es, por ejemplo, Granada:
```
def process_signature_v2(comunidad, provincia, municipio):
if provincia == 'Granada':
return None
else:
return 'OK'
udf_process_signature_v2 = fn.udf(process_signature_v2)
query2 = df \
.selectExpr("topic", "CAST(key AS STRING)", "CAST(value AS STRING) AS value") \
.withColumn("value_json", fn.from_json(col('value'), schema)) \
.select('value_json.COMUNIDAD', 'value_json.PROVINCIA', 'value_json.MUNICIPIO', 'value_json.PARTIDO') \
.withColumn('SIGNATURE', udf_process_signature_v2(col('COMUNIDAD'), col('PROVINCIA'), col('MUNICIPIO'))) \
.where(~ fn.isnull(col('SIGNATURE'))) \
.groupBy('COMUNIDAD', 'PROVINCIA', 'PARTIDO') \
.agg(fn.count('*').alias('VOTOS')) \
.sort(col('COMUNIDAD').asc(), col('PROVINCIA').asc(), col('VOTOS').desc()) \
.writeStream \
.outputMode("complete") \
.format("memory") \
.queryName('dashboard_v2') \
.start()
spark.sql("""
SELECT COMUNIDAD, PARTIDO, sum(VOTOS) as VOTOS
FROM dashboard_v2
WHERE VOTOS > 2 and COMUNIDAD LIKE 'And%'
GROUP BY COMUNIDAD, PARTIDO
ORDER BY VOTOS DESC
""").show(100, False)
query2.stop()
```
| github_jupyter |
```
import numpy as np
import matplotlib.pyplot as plt
from EMAN2 import *
#### select one GPU when multiple GPUs are present
os.environ["CUDA_VISIBLE_DEVICES"]='0'
#### do not occupy the entire GPU memory at once
## seems necessary to avoid some errors...
os.environ["TF_FORCE_GPU_ALLOW_GROWTH"]='true'
#### finally initialize tensorflow
import tensorflow as tf
#### we will import some functions from e2gmm_refine later
emdir=e2getinstalldir()
sys.path.insert(0,os.path.join(emdir,'bin'))
#### need to unify the float type across tenforflow and numpy
## in theory float16 also works but it can be unsafe especially when the network is deeper...
floattype=np.float32
#### load particles and prepare some parameters
from e2gmm_refine import load_particles, set_indices_boxsz
fname="r3d_00/ptcls_01.lst"
## Fourier box size. will shrink particles accordingly
maxboxsz=48
## load metadata first
e=EMData(fname, 0, True)
raw_apix, raw_boxsz = e["apix_x"], e["ny"]
data_cpx, xfsnp = load_particles(fname, maxboxsz, shuffle=False)
print("Image size: ", data_cpx[0].shape)
## set up fourier indices for image generation/clipping later
## params is a dictionay that saves matrices for Fourier indexing
apix=raw_apix*raw_boxsz/maxboxsz
clipid=set_indices_boxsz(data_cpx[0].shape[1], apix, True)
params=set_indices_boxsz(maxboxsz)
## create an empty options object that is needed for some functions in e2gmm_refine
options=type('options', (object,), {})()
options.sym="c1"
#### load the Gaussian model produced by e2gmm_refine.py
from e2gmm_refine import get_clip
pts=np.loadtxt("gmm_00/model_gmm.txt").astype(floattype)
print("Gaussian model shape: ", pts.shape)
## turn model to tensorflow format
pts=tf.constant(pts[None,:,:])
#### now generate some images to make sure everything is working properly
from e2gmm_refine import pts2img, calc_frc
sz=params["sz"] ## size of image
bsz=8 ## batch size
## generate a batch of projection images from the GMM at neutral state
## pts2img produce complex images in (real, imag) format
## turn them into complex numbers and IFT to get the real images
ptsx=tf.Variable(tf.repeat(pts, bsz, axis=0))
imgs_real, imgs_imag=pts2img(ptsx, xfsnp[:bsz], params, lp=.05)
imgs_cpx=imgs_real.numpy().astype(floattype)+1j*imgs_imag.numpy().astype(floattype)
imgs_out=np.fft.irfft2(imgs_cpx)
## just show one projection-particle comparison
ii=1
plt.figure(figsize=(9,3))
plt.subplot(1,3,1)
plt.imshow(imgs_out[ii], cmap='gray')
## get the complex particle image and IFT to real space
dcpx=(data_cpx[0][:bsz], data_cpx[1][:bsz])
dcpx=get_clip(dcpx, sz, clipid)
dcpx_out=np.fft.irfft2(dcpx[0].numpy()+1j*dcpx[1].numpy())
plt.subplot(1,3,2)
plt.imshow(dcpx_out[ii], cmap='gray')
## calculate FRC between particle and projection
## the calc_frc function takes complex images in (real, imag) format
frc=calc_frc(dcpx, (imgs_real, imgs_imag), params["rings"], True)
plt.subplot(1,3,3)
plt.plot(frc[ii][1:])
fval=calc_frc(dcpx, (imgs_real, imgs_imag), params["rings"])
print(np.mean(fval))
#### calculate d(FRC)/d(GMM) for each particle
## this will be the input for the deep network in place of the particle images
from e2gmm_refine import calc_gradient
## shrink particles to the requested size
dcpx=get_clip(data_cpx, params["sz"], clipid)
## prepare training set to be fed into the function
trainset=tf.data.Dataset.from_tensor_slices((dcpx[0], dcpx[1], xfsnp))
trainset=trainset.batch(bsz)
allscr, allgrds=calc_gradient(trainset, pts, params, options)
## histogram FRC for each particle
plt.hist(allscr, 20);
plt.xlabel("FRC score");
plt.ylabel("Particle count");
#### now build the deep networks and do some test
from e2gmm_refine import build_encoder, build_decoder
nmid=4 ## size of latent space
conv=True ## convolution mode, should be more powerful
## build the actual network
encode_model=build_encoder(nout=nmid, conv=conv)
decode_model=build_decoder(pts[0].numpy(), ninp=nmid, conv=conv)
## test the network to make sure they are function properly
## before training, they should output GMMs that are very close to the neutral GMM
mid=encode_model(allgrds[:bsz])
print("Latent space shape: ", mid.shape)
out=decode_model(mid)
print("Output shape: ",out.shape)
print("Deviation from neutral model: ", np.mean(abs(out-pts)))
#### train the network from the particles
from e2gmm_refine import train_heterg
## parse parameters as options
options.niter=10 ## number of iterations
options.pas=[0,1,0] ## mask the (position, amplitude, sigma) of the GMM during training
options.learnrate=1e-4 ## learning rate
ptclidx=allscr>-1 ## this allows the selection of a subset of particles for training. now used now
## actual training
trainset=tf.data.Dataset.from_tensor_slices((allgrds[ptclidx], dcpx[0][ptclidx], dcpx[1][ptclidx], xfsnp[ptclidx]))
trainset=trainset.batch(bsz)
train_heterg(trainset, pts, encode_model, decode_model, params, options)
#### compute the conformation of each particle
from e2gmm_refine import calc_conf
## this is a 4D latent space and can be hard to visualize
mid=calc_conf(encode_model, allgrds[ptclidx])
print(mid.shape)
plt.scatter(mid[:,0], mid[:,1], mid[:,2]+10, mid[:,3]+1, alpha=.4, cmap="RdBu");
#### do pca on the latent space for better visualization
from sklearn.decomposition import PCA
pca=PCA(2)
p2=pca.fit_transform(mid)
plt.scatter(p2[:,0], p2[:,1],s=5, alpha=.2);
#### classify particles from PCA space
from sklearn import mixture
clust = mixture.GaussianMixture(3)
lbs=clust.fit_predict(p2[:,:2])
plt.scatter(p2[:,0], p2[:,1],s=10, c=lbs, alpha=.4, cmap="prism");
#### plot GMM for each class
cnt=clust.means_ ## start from class centers
cnt=pca.inverse_transform(cnt) ## inverse PCA to back to the neural network latent space
pcnt=decode_model(cnt.astype(floattype)) ## go through the decoder to get the Gaussian model
pcnt=pcnt.numpy()
plt.figure(figsize=(9,3))
for i,p in enumerate(pcnt):
plt.subplot(1,3,i+1)
plt.scatter(pts[0,:,0], pts[0,:,1], s=p[:,3]*10+5, c=p[:,3]*10, alpha=.4, cmap="Blues")
plt.axis("square")
#### generate particles subsets from classification
from EMAN2_utils import load_lst_params, save_lst_params
ptclinfo=load_lst_params(fname)
for l in np.unique(lbs):
cls=(lbs==l)
psave=[p for i,p in enumerate(ptclinfo) if cls[i]]
print("Class {} : {} particles".format(l, len(psave)))
save_lst_params(psave, "gmm_00/ptcl_cls_{:02d}.lst".format(l))
#### reconstruct particles for each class
for i in range(3):
launch_childprocess("e2make3dpar.py --input gmm_00/ptcl_cls_{:02d}.lst --output gmm_00/threed_cls_{:02d}.hdf --pad 168 --mode trilinear --no_wt --keep 1 --threads 12 --setsf strucfac.txt".format(i,i))
#### look at the 3d map from each class
## this should be similar to the Gaussian model of each class
plt.figure(figsize=(9,3))
for i in range(3):
e=EMData("gmm_00/threed_cls_{:02d}.hdf".format(i))
e=e.numpy().copy()
e=np.mean(e, axis=0)
plt.subplot(1,3,i+1)
plt.imshow(e, cmap='gray')
```
| github_jupyter |
### Aula 1
### Jupyter Notebook
"Jupyter Notebooks é uma aplicação web que pode ajudar a entender e visualizar dados e resultados de análises, juntamente com o código! Facilita a experimentação, colaboração e publicação online. Anteriormente conhecido como IPython notebooks, hoje permite o uso de outras linguagens, sendo Python o default."
Guia: https://www.datacamp.com/community/tutorials/tutorial-jupyter-notebook
#### Instalação:
Após o Python e o pip instalado execute:
***No Windows:***
```
python -m pip install -U pip setuptools
```
***No Linux***
```
pip install -U pip setuptools
```
E instale o jupyter:
***Python2***
```
pip install jupyter
```
***Python 3***
```
pip3 install jupyter
```
Para usar abra o terminal/bash na pasta do seu projeto e escreva jupyter-notebook.
O aplicativo rodará em seu broswer.
As caixas de texto são ***células***, cada uma delas pode ser executada separadamente, nelas podemos escrever códigos em python, markdown além de outras linguagens. Com o passar do tempo aprenderemos a trabalhar com elas.
Podemos escrever no formato markdown:
# Este é um título H1
## Este é um título H2
### Este é um título H3...
***Assim escrevemos em negrito***
* Agora
* Alguns
* Bullets
***Aprenda mais sobre markdown [clique aqui](https://guides.github.com/features/mastering-markdown/)***
#### Podemos colocar fórmulas matemáticas no formato latec usando $c
$$c = \sqrt{a^2 + b^2 } $$
## Plotando gráficos
#### Plotar gráficos com a matplotlib é muito fácil, tendo uma série de números basta chamar o método plot
Veja a documentação completa da matplotlib ***[AQUI](https://matplotlib.org/)***
No código abaixo importamos o pacote da biblioteca matplotlib e criamos um vetor com os pontos que desejamos plotar:
```
import matplotlib.pyplot as plt
%matplotlib inline
entradas = [1, -1, 3, 8, -3]
plt.plot(entradas)
```
Para saber mais use o comando help. Esse comando é seu maior aliado.
```
plt.plot?
```
#### Então podemos alterar os parâmetros para fazer criar um gráfico mais legível:
```
plt.plot(entradas, color="red", linestyle="dashed", marker = 'o', markerfacecolor="yellow")
```
### Podemos plotar funções matemáticas:
```
import numpy as np
x = np.linspace(0, 2, 100)
plt.plot(x, x, label='linear')
plt.plot(x, x**2, label='quadratico')
plt.plot(x, x**3, label='cubico')
plt.xlabel('Eixo X')
plt.ylabel('Eixo Y')
plt.title("Funções")
plt.legend()
plt.show()
x = np.arange(0, 10, 0.2)
y = np.sin(x)
fig, ax = plt.subplots()
ax.plot(x, y)
plt.show()
```
# Seaborn
Outra ferramenta muito boa para plotar gráficos é o Seaborn:
Conheça [aqui](https://seaborn.pydata.org)
________________________________________________________________________________________________________________________________
# Datasets
Dataset ou conjunto de dados são arquivos que contém milhares de informações sobre determinados assuntos, esses arquivos podem vir no formado XLS, XLSX, EXCEL, HTML, JSON, XML entre outros. Datasets são nada mais do que planilhas, com linhas e colunas. Por exemplo se quisermos guardar em um dataset o nome e idade de um milhão de pessoas, teremos um arquivo com um milhão de linhas e duas colunas.
#### Onde encontrar Datasets:
* [Kaggle](https://www.kaggle.com/datasets) O Kaggle é uma ótima ferramenta para encontrar desafios e se conectar com outros cientistas de dados do mundo todo, vai ser uma ótima ferramenta para aumentar seu portifolio, lá eles possuem vários datasets legais.
* [Dados.gov.br](http://dados.gov.br/) O portal brasileiro de dados abertos. Lá você vai encontrar datasets sobre diferentes áreas e setores do Brasil, de transportes a saúde ***Lembrando que com grandes poderes vem grandes responsabilidades***.
* [IBGE-sidra](https://sidra.ibge.gov.br/home/primpec/brasil) O site do IBGE fornece seus datasets pela plataforma sidra.
* [FiveThirtyEight](https://github.com/fivethirtyeight/data) Esse é um site que contém notícias e esportes, lá no git deles temos muitos datasets.
* ***Esse aqui nem vou falar nada só use: https://github.com/awesomedata/awesome-public-datasets***
Você também pode montar seu Dataset utilizando técnicas de Data Mining que serão abordadas a frente.
# Pandas
Para trabalhar com os datasets no python ultilizamos a biblioteca [pandas](https://pandas.pydata.org/).
Pandas é uma biblioteca open source BSD-licensed que garante alta performance e facilidade de uso com ferramentas de estrutura de dados para análise de dados.
Para abrir um dataset em csv no pandas ultilize o método 'read_csv' com o nome do arquivo, para visualizar chame escrevendo o nome da variável atribuida:
# Vamos aprender pandas com Pokemons
```
import pandas as pd
ds = pd.read_csv('Pokemon.csv')
ds
```
#### Como os datasets normalmente são grandes ultilizamos os metodos head e tail do pandas para ver o início e fim do arquivo
```
ds.head()
ds.tail()
```
### Para melhorar a visualização vamos passar os cabeçalhos para maiúsculo
```
ds.columns = ds.columns.str.upper().str.replace('_', '')
ds.head()
```
#### Para ver o número de linhas e colunas use o método shape.
```
ds.shape
```
#### O comando columns retorna o nome das colunas.
```
ds.columns
```
## Iniciando as análises
O pandas nos dá muitas ferramentas estatísticas prontas, para aprender vamos estudar um pouco sobre pokemons.
<img src="https://encrypted-tbn0.gstatic.com/images?q=tbn:ANd9GcTRo-QyGWQ9mTlBLZH_RfzyS1GLhb08b8rZmdi25oWb60dFFD36" width="100" height="100" />
No pandas cada coluna é uma variável e são acessadas com ponto. Vamos por exemplo listar o nome de todos os pokemons do dataset.
```
ds.NAME
```
#### O método describe nos mostra um resumo estatístico da coluna estatística. Por exemplo seleciona-se as estatísticas de ataque dos pokemons:
```
ds.ATTACK.describe()
```
#### Interpretando describe:
* count indica o número de linhas (pokemons)
* mean indica a média aritimética de ataque dos pokemons
* std Desvio padrão
* min o pokemon com o menor ataque
* 25% indica que 25% dos pokemons tem 55 de ataque
* max indica o pokemon com maior ataque
#### É muito ultil isolar essas informações por exemplo como capturar somente a média de ataque.
```
print("A média de ataque dos pokemons é " + str(ds.ATTACK.mean()))
```
#### Podemos ver as infos de um único pokemon, obviamente escolhemos o melhor de todos o Charmander
<img src="https://encrypted-tbn0.gstatic.com/images?q=tbn:ANd9GcRNHHfoobz73wfwm5NQFrrQ2RgMyese0BXYDazTVL-8aIkMNYglrw" width="100" height="100" />
```
ds[ds['NAME'] == 'Charmander']
```
#### Para dificultar eu gostaria de saber todos pokemons dragões ou de fogo
```
ds[((ds['TYPE 1']=='Fire') | (ds['TYPE 1']=='Dragon')) & ((ds['TYPE 2']=='Dragon') | (ds['TYPE 2']=='Fire'))]
```
#### Vamos mostrar os 10 pokemons com maior quantidade de indivíduos.
```
ds.sort_values('TOTAL',ascending=False).head(10)
```
### Contando e agrupando os pokemons de cada tipo:
```
print(ds['TYPE 1'].value_counts(), '\n' ,ds['TYPE 2'].value_counts())
ds.groupby(['TYPE 1']).size()
(ds['TYPE 1']=='Bug').sum()
```
#### Pode-se criar facilmente gráficos do matplotlib:
Plotando o valor de ataque de todos os pokemons marcando a média:
```
plt.hist(ds["ATTACK"])
plt.xlabel('Ataque')
plt.ylabel('Valor')
plt.plot()
plt.axvline(ds['ATTACK'].mean(),linestyle='dashed',color='purple')
plt.show()
```
#### Plotando pokemons pelo tipo
```
labels = 'Agua', 'Normal', 'Metal', 'Inseto', 'Psiquico', 'Fogo', 'Eletrico', 'Rocha', 'Outros'
sizes = [112, 98, 70, 69, 57, 52, 44, 44, 175]
colors = ['Y', 'B', '#00ff00', 'C', 'R', 'G', 'silver', 'white', 'M']
explode = (0, 0, 0, 0, 0, 0, 0, 0, 0)
plt.pie(sizes, explode=explode, labels=labels, colors=colors,
autopct='%1.1f%%', shadow=True, startangle=90)
plt.axis('equal')
plt.title("Dividindo os pokemons pelo tipo")
plt.plot()
fig=plt.gcf()
fig.set_size_inches(7,7)
plt.show()
```
#### Por ultimo vamos plotar as estatísticas de ataque dos pokemons em função do tipo, plotamos grafico do tipo bloxpot usando o Seaborn:
```
import seaborn as sns
plt.subplots(figsize = (15,5))
plt.title('Ataque pelo tipo')
sns.boxplot(x = "TYPE 1", y = "ATTACK",data = ds)
plt.ylim(0,200)
plt.show()
```
### Por ultimo usamos dispersão para plotar o ataque dos pokemons em função da geração.
```
plt.scatter(ds["ATTACK"], ds["GENERATION"]);
```
| github_jupyter |
# Spot detection with napari
### Overview
In this activity, we will perform spot detection on some in situ sequencing data ([Feldman and Singh et al., Cell, 2019](https://www.cell.com/cell/fulltext/S0092-8674(19)31067-0s)). In doing so, we will combine methods from [scipy](https://www.scipy.org/), [scikit-image](https://scikit-image.org/), and [cellpose](https://github.com/MouseLand/cellpose). The goal is to familiarize you with performing analysis that integrates the scientific python ecosystem and napari.
### Data source
The data were downloaded from the [OpticalPooledScreens github repository](https://github.com/feldman4/OpticalPooledScreens).
### Next steps
Following this activity, we will use the workflow generated in this activity to create a napari spot detection plugin.
### screenshots
For the solution notebook, we are including screenshots via the `nbscreenshot` utility. These are not required for your notebook.
An example usage: `nbscreenshot(viewer)`
```
from napari.utils import nbscreenshot
```
# Load the data
In the cells below load the data using the scikit-image `imread()` function. For more information about the `imread()` function, please see the [scikit-image docs](https://scikit-image.org/docs/dev/api/skimage.io.html#skimage.io.imread). We are loading two images:
- `nuclei`: an image of cell nuclei
- `spots`: an image of in situ sequencing spots
```
from skimage import io
nuclei_url = 'https://raw.githubusercontent.com/kevinyamauchi/napari-spot-detection-tutorial/main/data/nuclei_cropped.tif'
nuclei = io.imread(nuclei_url)
spots_url = 'https://raw.githubusercontent.com/kevinyamauchi/napari-spot-detection-tutorial/main/data/spots_cropped.tif'
spots = io.imread(spots_url)
```
# View the data
We will use napari to view our data. To do so, we first must create the viewer. Once the Viewer is created, we can add images to the viewer via the Viewer's `add_image()` method.
```
import napari
# create the napari viewer
viewer = napari.Viewer();
# add the nuclei image to the viewer
viewer.add_image(nuclei);
```
In the cell below, add the spots image to the viewer as was done above for the nuclei image. After loading the data, inspect it in the viewer and adjust the layer settings to your liking (e.g., contrast limits, colormap). You can pan/zoom around the image by click/dragging to pan and scrolling with your mousewheel or trackpad to zoom.
**Hint**: you can adjust a layer's opacity to see the change how much you see of the layers that are "under" it.
```
# add the spots image to the viewer
viewer.add_image(spots)
from napari.utils import nbscreenshot
nbscreenshot(viewer)
```
# Create an image filter
You may have noticed the the spots image contains background and autofluorescence from the cells. To improve spot detection, we will apply a high pass filter to improve the contrast of the spots.
```
import numpy as np
from scipy import ndimage as ndi
def gaussian_high_pass(image: np.ndarray, sigma: float = 2):
"""Apply a gaussian high pass filter to an image.
Parameters
----------
image : np.ndarray
The image to be filtered.
sigma : float
The sigma (width) of the gaussian filter to be applied.
The default value is 2.
Returns
-------
high_passed_im : np.ndarray
The image with the high pass filter applied
"""
low_pass = ndi.gaussian_filter(image, sigma)
high_passed_im = image - low_pass
return high_passed_im
```
In the cell below, apply the gaussian high pass filter to the `spots` image and add the image to the viewer.
```
# Use the gaussian_high_pass function to filter the spots image
filtered_spots = gaussian_high_pass(spots, 2)
# add the filtered image to the viewer
# hint: set the opacity < 1 in order to see the layers underneath
viewer.add_image(filtered_spots, opacity=0.6, colormap='viridis')
nbscreenshot(viewer)
```
# Detect spots
Next, we will create a function to detect the spots in the spot image. This function should take the raw image, apply the gaussian high pass filter from above and then use one of the blob detection algorithms from sci-kit image to perform the blob detection. The `detect_spots()` function should return a numpy array containing the coordinates of each spot and a numpy array containing the diameter of each spot.
Some hints:
- See the [blob detection tutorial from scikit-image](https://scikit-image.org/docs/dev/auto_examples/features_detection/plot_blob.html). - We recommend the [blob_log detector](https://scikit-image.org/docs/dev/api/skimage.feature.html#skimage.feature.blob_log), but feel free to experiment!
- See the "Note" from the blob_log docs: "The radius of each blob is approximately $\sqrt{2}\sigma$ for a 2-D image"
```
import numpy as np
from skimage.feature import blob_log
def detect_spots(
image: np.ndarray,
high_pass_sigma: float = 2,
spot_threshold: float = 0.01,
blob_sigma: float = 2
):
"""Apply a gaussian high pass filter to an image.
Parameters
----------
image : np.ndarray
The image in which to detect the spots.
high_pass_sigma : float
The sigma (width) of the gaussian filter to be applied.
The default value is 2.
spot_threshold : float
The threshold to be passed to the blob detector.
The default value is 0.01.
blob_sigma: float
The expected sigma (width) of the spots. This parameter
is passed to the "max_sigma" parameter of the blob
detector.
Returns
-------
points_coords : np.ndarray
An NxD array with the coordinate for each detected spot.
N is the number of spots and D is the number of dimensions.
sizes : np.ndarray
An array of size N, where N is the number of detected spots
with the diameter of each spot.
"""
# filter the image
filtered_spots = gaussian_high_pass(image, high_pass_sigma)
# detect the spots on the filtered image
blobs_log = blob_log(
filtered_spots,
max_sigma=blob_sigma,
num_sigma=1,
threshold=spot_threshold
)
# convert the output of the blob detector to the
# desired points_coords and sizes arrays
# (see the docstring for details)
points_coords = blobs_log[:, 0:2]
sizes = 2 * np.sqrt(2) * blobs_log[:, 2]
return points_coords, sizes
```
In the cell below, apply `detect_spots()` to our `spots` image. To visualize the results, add the spots to the viewer as a [Points layer](https://napari.org/tutorials/fundamentals/points.html). If you would like to see an example of using a points layer, see [this example](https://github.com/napari/napari/blob/master/examples/add_points.py). To test out your function, vary the detection parameters and see how they affect the results. Note that each time you run the cell, the new results are added as an additional Points layer, allowing you to compare results from different parameters. To make it easier to compare results, you can try modifying the layer opacity or toggling the visibility with the "eye" icon in the layer list.
```
# detect the spots
spot_coords, spot_sizes = detect_spots(
spots,
high_pass_sigma=2,
spot_threshold=0.01,
blob_sigma=2
)
# add the detected spots to the viewer as a Points layer
viewer.add_points(
spot_coords,
size=spot_sizes,
opacity=0.5
)
nbscreenshot(viewer)
```
## Conclusion
In this activity, we have interactively prototyped a spot detection function using a combination of jupyter notebook, scipy, scikit-image, and napari. In the next activity, we will take the spot detection function we created and turn it into a napari plugin.
| github_jupyter |
```
import gensim
word2vecmodel = gensim.models.KeyedVectors.load_word2vec_format(path.expanduser('~/work/jupyter_notebooks/notebooks/GoogleNews-vectors-negative300.bin'), binary=True)
word2vecmodel.save(path.expanduser('~/work/data/word2vec.bin'))
word2vecmodel = gensim.models.KeyedVectors.load(path.expanduser('~/work/data/word2vec.bin'), mmap='r')
import requests
url = 'https://www.ted.com/talks/craig_costello_cryptographers_quantum_computers_and_the_war_for_information'
title = url.split('/')[-1]
metaURL = 'https://api.ted.com/v1/talks/{}.json?api-key=uzdyad5pnc2mv2dd8r8vd65c'
r = requests.get(url = metaURL.format(title))
data = r.json()
id = data['talk']['id']
# [tag['tag'] for tag in data['talk']['tags']]
import numpy as np
import pickle
import pandas as pd
from os import path
import re
with open(path.expanduser('~/work/data/data.pkl'), "rb") as input_file:
data = pickle.load(input_file)
tags = [re.sub('\'','',taglist)[1:-1].split(', ') for taglist in data['tags'].values]
file = open(path.expanduser('~/work/data/tags.txt'), "w")
for talktags in tags:
file.write(' '.join(talktags) + '\n')
file.close()
import requests
import numpy as np
import nltk
import re
import gensim
nltk.download('stopwords')
nltk.download('wordnet')
def retrieve_prepare_subtitles(url):
metaDataURL = 'https://api.ted.com/v1/talks/{}.json?api-key=uzdyad5pnc2mv2dd8r8vd65c'
subURL = 'https://api.ted.com/v1/talks/{}/subtitles.json?api-key=uzdyad5pnc2mv2dd8r8vd65c'
r = requests.get(url = subURL.format(id))
data = r.json()
transcript = []
for dict in np.arange(len(data)-1):
text = data[str(dict)]['caption']['content']
text = re.sub(r'\((.*?)\)', ' ', text)
text = re.sub(r'\d+', '', text)
text = nltk.tokenize.RegexpTokenizer(r'\w+').tokenize(text.lower())
text = [w for w in text if w not in nltk.corpus.stopwords.words('english')]
text = [nltk.corpus.wordnet.morphy(w) if nltk.corpus.wordnet.morphy(w) else w for w in text]
transcript.extend(text)
return ' '.join(transcript)
import os
file = open(path.expanduser('~/work/test/lftm/doc.txt'), "w")
file.write(retrieve_prepare_subtitles(176))
file.close()
model = 'LFLDAinf'
paras = 'TEDLFLDA.paras'
corpus = 'doc.txt'
initer = '500'
niter = '50'
topn = '20'
name = 'TEDLFLDAinf'
sstep = '0'
os.chdir(os.path.expanduser('~/work/test/lftm/'))
os.system('java -jar jar/LFTM.jar -model {} -paras {} -corpus {} -initers {} -niters {} -twords {} -name {} -sstep {}'.format(
model,
paras,
corpus,
initer,
niter,
topn,
name,
sstep))
file = open(path.expanduser('~/work/test/lftm/TEDLFLDAinf.theta'), "r")
topics = file.readline()
file.close()
def train(datapath):
text = []
with open(datapath,"r+") as f:
while True:
vec = f.readline()
text.append(text)
import pickle
from os import path
import pandas as pd
with open(path.expanduser('~/work/data/data.pkl'), "rb") as input_file:
data = pickle.load(input_file)
import json
import requests
import pandas as pd
import numpy as np
metaDataURL = 'https://api.ted.com/v1/talks/{}.json?api-key=uzdyad5pnc2mv2dd8r8vd65c'
subURL = 'https://api.ted.com/v1/talks/{}/subtitles.json?api-key=uzdyad5pnc2mv2dd8r8vd65c'
_id=1
talk_meta = pd.DataFrame()
talk_sub = pd.DataFrame()
talk = {}
sub = {}
while True:
response_meta = requests.get(metaDataURL.format(_id))
response_meta = response_meta.json()
if 'error' in response_meta:
print("Id",_id, "not found.")
_id += 1
continue
response_sub = requests.get(subURL.format(_id))
response_sub = response_sub.json()
if 'error' in response_sub:
print("Sub for",_id, "not found.")
_id += 1
continue
talk['talk_id'] = _id
talk['name'] = response_meta['talk']['name']
talk['tags'] = ','.join([tag['tag'] for tag in response_meta['talk']['tags']])
talk_meta = talk_meta.append(talk, ignore_index=True)
chapter_id = 0
chapter = []
no_caption = len(response_sub)
chapter.append(response_sub['0']['caption']['content'])
for dict in np.arange(2,no_caption - 2):
if response_sub[str(dict)]['caption']['startOfParagraph']:
sub['talk_id'] = _id
sub['id'] = chapter_id
sub['transcript'] = ' '.join(chapter)
talk_sub = talk_sub.append(sub, ignore_index=True)
chapter_id += 1
chapter = []
chapter.append(response_sub[str(dict)]['caption']['content'])
else:
chapter.append(response_sub[str(dict)]['caption']['content'])
if response_sub[str(no_caption - 2)]['caption']['startOfParagraph']:
sub['talk_id'] = _id
sub['id'] = chapter_id
sub['transcript'] = ' '.join(chapter)
talk_sub = talk_sub.append(sub, ignore_index=True)
chapter_id += 1
chapter = []
chapter.append(response_sub[str(no_caption - 2)]['caption']['content'])
sub['talk_id'] = _id
sub['id'] = chapter_id
sub['transcript'] = ' '.join(chapter)
talk_sub = talk_sub.append(sub, ignore_index=True)
else:
sub['talk_id'] = _id
sub['id'] = chapter_id
chapter.append(response_sub[str(no_caption - 2)]['caption']['content'])
sub['transcript'] = ' '.join(chapter)
talk_sub = talk_sub.append(sub, ignore_index=True)
# text = re.sub(r'\((.*?)\)', ' ', text)
# text = re.sub(r'\d+', '', text)
# text = nltk.tokenize.RegexpTokenizer(r'\w+').tokenize(text.lower())
# text = [w for w in text if w not in nltk.corpus.stopwords.words('english')]
# text = [nltk.corpus.wordnet.morphy(w) if nltk.corpus.wordnet.morphy(w) else w for w in text]
# transcript.extend(text)
print("Loaded talk",_id)
_id += 1
talk_sub.groupby('talk_id').count()
talk_sub[(talk_sub.talk_id ==1) & (talk_sub.id == 0)].sort_values('id').values
talk_meta
talk_data = talk_meta.set_index('talk_id').join(talk_sub.set_index('talk_id'), 'talk_id')
talk_data = talk_meta.set_index('talk_id').join(talk_sub.set_index('talk_id'), on = 'talk_id')
talk_meta = talk_meta[~talk_meta['talk_id'].isin(talk_data[talk_data['transcript'].isna()].index)]
with open(path.expanduser('~/work/data/talk_meta.pkl'), 'wb') as output:
pickle.dump(talk_meta, output, pickle.HIGHEST_PROTOCOL)
with open(path.expanduser('~/work/data/talk_sub.pkl'), 'wb') as output:
pickle.dump(talk_sub, output, pickle.HIGHEST_PROTOCOL)
from os import path
import pickle
with open(path.expanduser('~/work/data/talk_meta.pkl'), "rb") as input_file:
talk_meta = pickle.load(input_file)
with open(path.expanduser('~/work/data/talk_sub.pkl'), "rb") as input_file:
talk_sub = pickle.load(input_file)
talk_sub.groupby('talk_id').count().sort_values('transcript')
talk_meta
!pip install nltk
import re
import nltk
nltk.download('stopwords')
nltk.download('wordnet')
def prepare_text(text):
text = re.sub(r'\((.*?)\)', ' ', text)
text = re.sub(r'\d+', '', text)
text = nltk.tokenize.RegexpTokenizer(r'\w+').tokenize(text.lower())
text = [w for w in text if not((w in ['thank', 'much']) or (w in nltk.corpus.stopwords.words('english')))]
text = [nltk.corpus.wordnet.morphy(w) if nltk.corpus.wordnet.morphy(w) else w for w in text]
return text
talk_sub[talk_sub.talk_id == 1]['transcript'].apply(lambda x: prepare_text(x)).values
tag2idx = {}
i=0
talktags = [taglist.split(',') for taglist in talk_meta['tags'].values]
for taglist in talktags:
for tag in taglist:
if tag not in tag2idx and not 'ted' in tag.lower():
tag2idx[tag] = i
i += 1
tag2idx
len(tag2idx)
from os import path
import pickle
with open(path.expanduser('~/work/data/talk_pre.pkl'), "rb") as input_file:
talk_pre = pickle.load(input_file)
# !sudo apt update -y
!pip install nltk
!pip install gensim
import re
import pandas as pd
from os import path
import nltk
import gensim
import time
from IPython.display import display, clear_output
nltk.download('stopwords')
nltk.download('wordnet')
data = pd.read_json('https://raw.githubusercontent.com/selva86/datasets/master/newsgroups.json')
data.head()
data.groupby('target_names').count()
lem = nltk.stem.WordNetLemmatizer()
def prepare_text(text):
text = text.lower()
text = re.sub('from:.+\n', '', text)
text = re.sub('article-i.d.:.+\n', '', text)
text = re.sub('nntp-posting-host:.+\n', '', text)
text = re.sub('organization:.+\n', '', text)
text = re.sub('x-newsreader:.+\n', '', text)
text = re.sub('distribution:.+\n', '', text)
text = re.sub('reply-to:.+\n', '', text)
text = re.sub('<.+>', '', text)
text = re.sub(r'\d+', '', text)
text = nltk.tokenize.RegexpTokenizer(r'\w+').tokenize(text)
text = [lem.lemmatize(w) for w in text]
text = [w for w in text if not(w in ['from', 'subject', 'edu', 'use', 'lines', 'ke'] or w in nltk.corpus.stopwords.words('english')) and len(w)>=3 and '__' not in w]
text = [w for w in text if len(w)>=3]
return text
data['content'] = data.content.apply(lambda x: prepare_text(x))
with open('20ng.txt', 'w') as f:
for d in data['content']:
f.write(' '.join(d))
f.write('\n')
data_words = data['content'].values
id2word = gensim.corpora.Dictionary(data_words)
id2word.filter_n_most_frequent(10)
# corpus = [id2word.doc2bow(text) for text in data_words]
def filter_text(text, tokens):
filtered_text = []
for word in text:
filtered_text.append(word)
return filtered_text
data['content'] = data.content.apply(lambda x: filter_text(x, id2word.token2id))
data['content'] = data.content.apply(lambda x: ' '.join(x))
f = open(path.expanduser('~/work/data/20NG.txt'),"w+")
for text in data['content']:
f.write(text+'\n')
f.close()
ref = pd.read_csv('~/work/data/wikipedia_utf8_filtered_20pageviews.csv',header=None)
ref.head()
lem = nltk.stem.WordNetLemmatizer()
def simple_prepare_text(text):
text = text.lower()
text = re.sub(r'\d+', '', text)
text = nltk.tokenize.RegexpTokenizer(r'\w+').tokenize(text)
text = [lem.lemmatize(w) for w in text]
text = [w for w in text if len(w)>=3]
text = ' '.join(text)
return text
# ref[1] = ref[1].apply(lambda x: simple_prepare_text(x))
len(ref)
f = open(path.expanduser('~/work/data/wiki.txt'),"w+")
i = 0
total = len(ref)
for text in ref[1]:
clear_output(wait=True)
start = time.time()
text = simple_prepare_text(text)
f.write(text+'\n')
dur = time.time() - start
display('Progress '+ str(100*i/total) + ' % - Duration per doc: '+ str(dur*1000) + ' ms')
i+=1
f.close()
print(str(5*463819/1000/60) + ' mins')
f = open(path.expanduser('~/work/data/sw.txt'),"w+")
for w in simple_prepare_text(' '.join(nltk.corpus.stopwords.words('english'))).split():
# s|\<the\>||g
f.write('s|\<'+w+'\>||g\n')
f.close()
import pandas as pd
import numpy as np
import pickle
from os import path
import re
import nltk
import gensim
import time
import matplotlib.pyplot as plt
from IPython.display import display, clear_output
nltk.download('stopwords')
nltk.download('wordnet')
lem = nltk.stem.WordNetLemmatizer()
def prepare_text(text):
text = text.lower()
text = re.sub(r'\d+', '', text)
text = nltk.tokenize.RegexpTokenizer(r'\w+').tokenize(text)
text = [lem.lemmatize(w) for w in text]
text = [w for w in text if w not in nltk.corpus.stopwords.words('english')]
text = [w for w in text if len(w)>=3]
return text
f1 = open(path.expanduser('~/work/data/asrael.txt'),"r")
f2 = open(path.expanduser('~/work/data/asrael_cleaned.txt'),"w")
i = 0
total = 125516
for line in f1:
clear_output(wait=True)
start = time.time()
text = ' '.join(prepare_text(line))
f2.write(text+'\n')
dur = time.time() - start
display('Progress '+ str(100*i/total) + ' % - Duration per doc: '+ str(dur*1000) + ' ms')
i+=1
f1.close()
f2.close()
```
| github_jupyter |
```
# default_exp core
```
# Core
> API details.
```
#hide
from nbdev.showdoc import *
#export
import os
import shutil
import requests
from pathlib import Path
from fastprogress import progress_bar
import zipfile, tarfile
#export
class Config:
config_path = Path('~/.aiadv').expanduser()
def __init__(self):
self.config_path.mkdir(parents=True, exist_ok=True)
#export
class URLs:
GDRIVE = "https://docs.google.com/uc?export=download&id="
# Datasets
YELP_REIVEWS = {'url': f'{GDRIVE}1Lmv4rsJiCWVs1nzs4ywA9YI-ADsTf6WB', 'fname': 'yelp_reveiw.csv'}
ENG_FRA_SAMPLE = {'url': f'{GDRIVE}1jLx6dZllBQ3LXZkCjZ4VciMQkZUInU10', 'fname': 'eng_fra_sample.csv'}
ENG_FRA = {'url': f'{GDRIVE}1o2ac0EliUod63sYUdpow_Dh-OqS3hF5Z', 'fname': 'eng_fra.txt'}
SURNAMES = {'url': f'{GDRIVE}1T1la2tYO1O7XkMRawG8VcFcvtjbxDqU-', 'fname': 'surnames.csv'}
CHD_49 = {'url': f'{GDRIVE}11yU-64VW4b9_tw-yWPUcnQ0A3-Uh-dd9', 'fname': 'chd_49.zip'}
ADV_SHERLOCK = {'url': 'https://norvig.com/big.txt', 'fname': 'adv_sherlock.txt'}
HUMAN_NUMBERS = {'url': 'http://files.fast.ai/data/examples/human_numbers.tgz', 'fname': 'human_numbers.tgz'}
MOVIELENS_SMALL = {'url': 'http://files.grouplens.org/datasets/movielens/ml-latest-small.zip', 'fname': 'ml-latest-small.zip'}
def path(ds=None):
fname = ds['fname']
path = Config().config_path/fname
return path
def stem(path):
if str(path).endswith('gz') or str(path).endswith('zip'):
parent = path.parent
return parent/path.stem
else: return path
#export
def download_data(ds, force_download=False):
"Download `url` to `fname`."
dest = URLs.path(ds)
if not dest.exists() or force_download:
download_url(ds['url'], dest, overwrite=force_download)
return dest
def file_extract(fname, dest=None):
"Extract `fname` using `tarfile` or `zipfile"
fname_str = str(fname)
if dest is None: dest = Path(fname).parent
if fname_str.endswith('gz' ): tarfile.open(fname, 'r:gz').extractall(dest)
elif fname_str.endswith('zip'): zipfile.ZipFile(fname ).extractall(dest)
else: raise Exception(f'Unrecognized archive: {fname}')
# cleaning up
os.remove(fname)
def download_url(url, dest, overwrite=False, pbar=None, show_progress=True, chunk_size=1024*1024,
timeout=4, retries=5):
"Download `url` to `dest` unless it exists and not `overwrite`"
if os.path.exists(dest) and not overwrite: return
s = requests.Session()
s.mount('http://',requests.adapters.HTTPAdapter(max_retries=retries))
# additional line to identify as a firefox browser, see fastai/#2438
s.headers.update({'User-Agent': 'Mozilla/5.0 (X11; Ubuntu; Linux x86_64; rv:71.0) Gecko/20100101 Firefox/71.0'})
u = s.get(url, stream=True, timeout=timeout)
try: file_size = int(u.headers["Content-Length"])
except: show_progress = False
with open(dest, 'wb') as f:
nbytes = 0
if show_progress: pbar = progress_bar(range(file_size), leave=False, parent=pbar)
try:
if show_progress: pbar.update(0)
for chunk in u.iter_content(chunk_size=chunk_size):
nbytes += len(chunk)
if show_progress: pbar.update(nbytes)
f.write(chunk)
except requests.exceptions.ConnectionError as e:
print('Connection Error, please check your internet')
#export
def untar_data(ds, force_download=False, extract_func=file_extract):
dest = URLs.path(ds)
stem = URLs.stem(dest)
fname = ds['fname']
if force_download:
if stem.exists():
try: os.remove(stem)
except: shutil.rmtree(stem)
if not stem.exists():
download_data(ds)
if str(fname).endswith('zip') or str(fname).endswith('gz'):
extract_func(dest)
return stem
path = untar_data(URLs.ENG_FRA, True)
path
!ls -lsh /home/ankur/.aiadv/
from nbdev.export import *
notebook2script()
```
| github_jupyter |
```
import os
import tensorflow as tf
import tensorflow_datasets as tfds
```
## One GPU strategy
```
# get available GPU
devices = tf.config.list_physical_devices('GPU')
tf.config.experimental.set_memory_growth(devices[0], True)
gpu_name = "GPU:0"
print(devices[0])
# Only one gpu available to set to OneDeviceStrategy
# Can be changed to MirroredStrategy if multiple GPU available
strategy = tf.distribute.OneDeviceStrategy(device=gpu_name)
```
## Get VOC 2012 dataset
```
# get voc 2012 dataset
splits = ['train[:80%]', 'train[80%:90%]', 'train[90%:]']
(train_examples, validation_examples, test_examples), info = tfds.load('voc/2012', batch_size=32, with_info=True, split=splits)
info.features
num_examples = info.splits['train'].num_examples
num_classes = info.features['labels'].num_classes
print(f"Number of train examples: {num_examples}, number of labels: {num_classes}")
```
## Preprocess VOC2012
```
# resize and normalize images
@tf.function
def format_image(tensor):
images = tf.image.resize(tensor['image'], IMAGE_SIZE) / 255.0
return images, tf.one_hot(tensor['objects']['label'], 20), tensor['objects']['bbox']
BUFFER_SIZE = num_examples
EPOCHS = 10
IMAGE_SIZE = (640, 640)
BATCH_SIZE_PER_REPLICA = 32
GLOBAL_BATCH_SIZE = BATCH_SIZE_PER_REPLICA * strategy.num_replicas_in_sync
# prepare batches
BATCH_SIZE = 32
train_batches = train_examples.shuffle(num_examples // 4).map(format_image).prefetch(1)
validation_batches = validation_examples.map(format_image)
test_batches = test_examples.map(format_image).batch(1)
for batch, labels, boxes in train_batches:
break
print(f"Batch shape: {batch.shape}")
print(f"Labels shape: {labels.shape} and boxes shape: {boxes.shape}")
```
## Distribute dataset over GPUs
```
# def distribute_datasets(strategy, train_batches, validation_batches, test_batches):
# train_dist_dataset = strategy.experimental_distribute_dataset(train_batches)
# val_dist_dataset = strategy.experimental_distribute_dataset(validation_batches)
# test_dist_dataset = strategy.experimental_distribute_dataset(test_batches)
# return train_dist_dataset, val_dist_dataset, test_dist_dataset
# train_dist_dataset, val_dist_dataset, test_dist_dataset = distribute_datasets(strategy, train_batches, validation_batches, test_batches)
# print(type(train_dist_dataset))
# for batch, labels, boxes in test_dist_dataset:
# break
# print(f"Batch shape: {batch.shape}")
# print(f"Labels shape: {labels.shape} and boxes shape: {boxes.shape}")
```
## Model
```
import os
import pathlib
# Clone the tensorflow models repository if it doesn't already exist
if "models" in pathlib.Path.cwd().parts:
while "models" in pathlib.Path.cwd().parts:
os.chdir('..')
elif not pathlib.Path('models').exists():
!git clone --depth 1 https://github.com/tensorflow/models
# Install the Object Detection API
%%bash
cd models/research/
protoc object_detection/protos/*.proto --python_out=.
cp object_detection/packages/tf2/setup.py .
python -m pip install .
import matplotlib
import matplotlib.pyplot as plt
import os
import random
import io
import imageio
import glob
import scipy.misc
import numpy as np
from six import BytesIO
from PIL import Image, ImageDraw, ImageFont
from IPython.display import display, Javascript
from IPython.display import Image as IPyImage
import tensorflow as tf
from object_detection.utils import label_map_util
from object_detection.utils import config_util
from object_detection.utils import visualization_utils as viz_utils
from object_detection.utils import colab_utils
from object_detection.builders import model_builder
%matplotlib inline
# Download the checkpoint and put it into models/research/object_detection/test_data/
!wget http://download.tensorflow.org/models/object_detection/tf2/20200711/ssd_resnet50_v1_fpn_640x640_coco17_tpu-8.tar.gz
!tar -xf ssd_resnet50_v1_fpn_640x640_coco17_tpu-8.tar.gz
!mv ssd_resnet50_v1_fpn_640x640_coco17_tpu-8/checkpoint models/research/object_detection/test_data/
tf.keras.backend.clear_session()
print('Building model and restoring weights for fine-tuning...', flush=True)
num_classes = 20
pipeline_config = 'models/research/object_detection/configs/tf2/ssd_resnet50_v1_fpn_640x640_coco17_tpu-8.config'
checkpoint_path = 'models/research/object_detection/test_data/checkpoint/ckpt-0'
# Load pipeline config and build a detection model.
#
# Since we are working off of a COCO architecture which predicts 90
# class slots by default, we override the `num_classes` field here to be just
# one (for our new rubber ducky class).
configs = config_util.get_configs_from_pipeline_file(pipeline_config)
model_config = configs['model']
model_config.ssd.num_classes = num_classes
model_config.ssd.freeze_batchnorm = True
detection_model = model_builder.build(
model_config=model_config, is_training=True)
# Set up object-based checkpoint restore --- RetinaNet has two prediction
# `heads` --- one for classification, the other for box regression. We will
# restore the box regression head but initialize the classification head
# from scratch (we show the omission below by commenting out the line that
# we would add if we wanted to restore both heads)
fake_box_predictor = tf.compat.v2.train.Checkpoint(
_base_tower_layers_for_heads=detection_model._box_predictor._base_tower_layers_for_heads,
# _prediction_heads=detection_model._box_predictor._prediction_heads,
# (i.e., the classification head that we *will not* restore)
_box_prediction_head=detection_model._box_predictor._box_prediction_head,
)
fake_model = tf.compat.v2.train.Checkpoint(
_feature_extractor=detection_model._feature_extractor,
_box_predictor=fake_box_predictor)
ckpt = tf.compat.v2.train.Checkpoint(model=fake_model)
ckpt.restore(checkpoint_path).expect_partial()
# Run model through a dummy image so that variables are created
image, shapes = detection_model.preprocess(tf.zeros([1, 640, 640, 3]))
prediction_dict = detection_model.predict(image, shapes)
_ = detection_model.postprocess(prediction_dict, shapes)
print('Weights restored!')
tf.get_logger().setLevel('ERROR')
from tqdm import tqdm
tf.keras.backend.set_learning_phase(True)
# These parameters can be tuned; since our training set has 5 images
# it doesn't make sense to have a much larger batch size, though we could
# fit more examples in memory if we wanted to.
batch_size = 32
learning_rate = 0.01
num_batches = len(train_batches)
# Select variables in top layers to fine-tune.
trainable_variables = detection_model.trainable_variables
to_fine_tune = []
prefixes_to_train = [
'WeightSharedConvolutionalBoxPredictor/WeightSharedConvolutionalBoxHead',
'WeightSharedConvolutionalBoxPredictor/WeightSharedConvolutionalClassHead']
for var in trainable_variables:
if any([var.name.startswith(prefix) for prefix in prefixes_to_train]):
to_fine_tune.append(var)
# Set up forward + backward pass for a single train step.
def get_model_train_step_function(model, optimizer, vars_to_fine_tune):
"""Get a tf.function for training step."""
# Use tf.function for a bit of speed.
# Comment out the tf.function decorator if you want the inside of the
# function to run eagerly.
@tf.function
def train_step_fn(image_tensors,
groundtruth_boxes_list,
groundtruth_classes_list):
"""A single training iteration.
Args:
image_tensors: A list of [1, height, width, 3] Tensor of type tf.float32.
Note that the height and width can vary across images, as they are
reshaped within this function to be 640x640.
groundtruth_boxes_list: A list of Tensors of shape [N_i, 4] with type
tf.float32 representing groundtruth boxes for each image in the batch.
groundtruth_classes_list: A list of Tensors of shape [N_i, num_classes]
with type tf.float32 representing groundtruth boxes for each image in
the batch.
Returns:
A scalar tensor representing the total loss for the input batch.
"""
shapes = tf.constant(batch_size * [[640, 640, 3]], dtype=tf.int32)
model.provide_groundtruth(
groundtruth_boxes_list=groundtruth_boxes_list,
groundtruth_classes_list=groundtruth_classes_list)
with tf.GradientTape() as tape:
preprocessed_images = tf.concat(
[detection_model.preprocess(image_tensor)[0]
for image_tensor in image_tensors], axis=0)
prediction_dict = model.predict(preprocessed_images, shapes)
losses_dict = model.loss(prediction_dict, shapes)
total_loss = losses_dict['Loss/localization_loss'] + losses_dict['Loss/classification_loss']
gradients = tape.gradient(total_loss, vars_to_fine_tune)
optimizer.apply_gradients(zip(gradients, vars_to_fine_tune))
return total_loss
return train_step_fn
optimizer = tf.keras.optimizers.SGD(learning_rate=learning_rate, momentum=0.9)
train_step_fn = get_model_train_step_function(
detection_model, optimizer, to_fine_tune)
print('Start fine-tuning!', flush=True)
#for idx in range(num_batches):
for epoch in range(20):
pbar = tqdm(total=num_batches, position=0, leave=True, bar_format='{l_bar}{bar}| {n_fmt}/{total_fmt} ')
for idx, (image_tensors, gt_classes_list, gt_boxes_list) in enumerate(train_batches):
image_tensors = tf.expand_dims(image_tensors, 1)
image_tensors, gt_boxes_list, gt_classes_list = list(image_tensors.numpy()), list(gt_boxes_list.numpy()), list(gt_classes_list.numpy())
# Training step (forward pass + backwards pass)
total_loss = train_step_fn(image_tensors, gt_boxes_list, gt_classes_list)
pbar.set_description("Training loss for step %s: %.4f" % (int(idx), float(total_loss)))
pbar.update()
print('Done fine-tuning!')
```
| github_jupyter |
```
import pandas as pd
import numpy as np
data=pd.read_csv('E:/PYTHON/WinterProject/england-premier-league-players-2018-to-2019-stats.csv')
#print(data)
#print(list(data.columns))
data_we_need=data[["full_name","position","Current Club","appearances_overall","goals_overall","assists_overall","penalty_goals","penalty_misses", 'clean_sheets_overall','yellow_cards_overall', 'red_cards_overall','Total_Price']]
print(data_we_need)
indexNames = pd.DataFrame()
#Running part of the code to be added at the start of the code
print("Welcome to the FPL Helper")
print("This will help you decide which player to keep in your team")
print("Please enter the Details of your team")
max_money_goalkeepers = 10
max_money_defenders = 26
max_mooney_midfielders = 42
max_money_forwards = 22
#This will leave a line
print("")
#this number should be between 3 to 5
total_team=[]
indexNames.drop(indexNames.index, inplace=True)
goalkeepers=data_we_need[data_we_need["position"]=="Goalkeeper"]
count_row = goalkeepers.shape[0]
def calculate_points(row):
return ((row["appearances_overall"]*2)+(row["goals_overall"]*6)+(row["assists_overall"]*3)+(row["penalty_goals"]*5)+(row["penalty_misses"]*-2)+(row["clean_sheets_overall"]*4)+(row['yellow_cards_overall']*-1)+(row['red_cards_overall']*-2))
goalkeepers["Points"] = goalkeepers.apply(calculate_points,axis=1)
#print(goalkeepers)
goalkeeper=[]
no_of_goalkeepers=int(input("Enter the number of goalkeepers you want in your team: "))
no_of_goalkeepers_selected=int(input("Enter the number of goalkeepers you have already added in your team: "))
no_of_gk_comp_suggest=int(no_of_goalkeepers-no_of_goalkeepers_selected)
print("No of suggestions in goalkeepers needed are: "+str(no_of_gk_comp_suggest))
for i in range(0,no_of_goalkeepers_selected):
temps_for_goalkeeper = str(input("Enter the player's name: "))
goalkeeper.append(temps_for_goalkeeper)
goalkeeper_temping = goalkeepers[goalkeepers['full_name'] == temps_for_goalkeeper ]
indexNames = indexNames.append(goalkeeper_temping)
temp_indexes = goalkeepers[goalkeepers['full_name'] == temps_for_goalkeeper ].index
# Delete these row indexes from dataFrame
goalkeepers.drop(temp_indexes , inplace=True)
#print(goalkeeper)
goalkeepers = goalkeepers.sort_values('Points',ascending=False)
selected_goalkeepers = goalkeepers.head(2)
#print(selected_goalkeepers)
#for i in range(no_of_gk_comp_suggest):
goalkeepers = goalkeepers.head(no_of_gk_comp_suggest)
#df[df['A'] == 2]['B']
indexNames = indexNames.append(goalkeepers)
print("Final Selected Goalkeepers")
print(indexNames)
defenders=data_we_need[data_we_need["position"]=="Defender"]
count_row = defenders.shape[0]
def calculate_points(row):
return ((row["appearances_overall"]*2)+(row["goals_overall"]*5)+(row["assists_overall"]*3)+(row["penalty_goals"]*5)+(row["penalty_misses"]*-2)+(row["clean_sheets_overall"]*4)+(row['yellow_cards_overall']*-1)+(row['red_cards_overall']*-2))
defenders["Points"] = defenders.apply(calculate_points,axis=1)
#print(defenders)
defender=[]
no_of_defenders=int(input("Enter the number of defenders you want in your team: "))
no_of_defenders_selected=int(input("Enter the number of defenders you have already added in your team: "))
no_of_gk_comp_suggest=int(no_of_defenders-no_of_defenders_selected)
print("No of suggestions in defenders needed are: "+str(no_of_gk_comp_suggest))
for i in range(0,no_of_defenders_selected):
temps_for_defenders = str(input("Enter the player's name: "))
defender.append(temps_for_defenders)
defenders_temping = defenders[defenders['full_name'] == temps_for_defenders ]
indexNames = indexNames.append(defenders_temping)
temp_indexes = defenders[defenders['full_name'] == temps_for_defenders ].index
# Delete these row indexes from dataFrame
defenders.drop(temp_indexes , inplace=True)
#print(defender)
defenders = defenders.sort_values('Points',ascending=False)
selected_defenders = defenders.head(2)
#print(selected_defenders)
#for i in range(no_of_gk_comp_suggest):
defenders = defenders.head(no_of_gk_comp_suggest)
#df[df['A'] == 2]['B']
indexNames = indexNames.append(defenders)
print("Final Selected Defenders")
print(indexNames)
midfielders=data_we_need[data_we_need["position"]=="Midfielder"]
count_row = midfielders.shape[0]
def calculate_points(row):
return ((row["appearances_overall"]*2)+(row["goals_overall"]*4)+(row["assists_overall"]*3)+(row["penalty_goals"]*5)+(row["penalty_misses"]*-2)+(row["clean_sheets_overall"]*4)+(row['yellow_cards_overall']*-1)+(row['red_cards_overall']*-2))
midfielders["Points"] = midfielders.apply(calculate_points,axis=1)
#print(midfielders)
midfielder=[]
no_of_midfielders=int(input("Enter the number of midfielders you want in your team: "))
no_of_midfielders_selected=int(input("Enter the number of midfielders you have already added in your team: "))
no_of_gk_comp_suggest=int(no_of_midfielders-no_of_midfielders_selected)
print("No of suggestions in midfielders needed are: "+str(no_of_gk_comp_suggest))
for i in range(0,no_of_midfielders_selected):
temps_for_midfielder = str(input("Enter the player's name: "))
midfielder.append(temps_for_midfielder)
midfielders_temping = midfielders[midfielders['full_name'] == temps_for_midfielder ]
indexNames = indexNames.append(midfielders_temping)
temp_indexes = midfielders[midfielders['full_name'] == temps_for_midfielder ].index
# Delete these row indexes from dataFrame
midfielders.drop(temp_indexes , inplace=True)
#print(midfielder)
midfielders = midfielders.sort_values('Points',ascending=False)
selected_midfielders = midfielders.head(2)
#print(selected_midfielders)
#for i in range(no_of_gk_comp_suggest):
midfielders = midfielders.head(no_of_gk_comp_suggest)
#df[df['A'] == 2]['B']
indexNames = indexNames.append(midfielders)
print("Final Selected Midfielders")
print(indexNames)
forwards=data_we_need[data_we_need["position"]=="Forward"]
count_row = forwards.shape[0]
def calculate_points(row):
return ((row["appearances_overall"]*2)+(row["goals_overall"]*4)+(row["assists_overall"]*3)+(row["penalty_goals"]*5)+(row["penalty_misses"]*-2)+(row["clean_sheets_overall"]*4)+(row['yellow_cards_overall']*-1)+(row['red_cards_overall']*-2))
forwards["Points"] = forwards.apply(calculate_points,axis=1)
#print(forwards)
forward=[]
no_of_forwards=int(input("Enter the number of forwards you want in your team: "))
no_of_forwards_selected=int(input("Enter the number of forwards you have already added in your team: "))
no_of_gk_comp_suggest=int(no_of_forwards-no_of_forwards_selected)
print("No of suggestions in forwards needed are: "+str(no_of_gk_comp_suggest))
for i in range(0,no_of_midfielders_selected):
temps_for_forward = str(input("Enter the player's name: "))
forward.append(temps_for_forward)
forwards_temping = forwards[forwards['full_name'] == temps_for_forward ]
indexNames = indexNames.append(forwards_temping)
temp_indexes = forwards[forwards['full_name'] == temps_for_forward ].index
# Delete these row indexes from dataFrame
forwards.drop(temp_indexes , inplace=True)
#print(forward)
forwards = forwards.sort_values('Points',ascending=False)
selected_forwards = forwards.head(3)
#print(selected_forwards)
#for i in range(no_of_gk_comp_suggest):
forwards = forwards.head(no_of_gk_comp_suggest)
#df[df['A'] == 2]['B']
indexNames = indexNames.append(forwards)
print("Final Selected Team will be")
Total_price_of_players = indexNames['Total_Price'].sum()
Total_Points = indexNames['Points'].sum()
print("The total price of the players :"+str(Total_price_of_players))
print("The total points of the players :"+str(Total_Points))
print(" ")
print(" ")
print("Team points per week :"+str(int(Total_Points/38)))
if(Total_price_of_players > 102):
print("You need more money to achieve this team which isn't possible so make some compromises")
print(indexNames)
```
| github_jupyter |
<table class="ee-notebook-buttons" align="left">
<td><a target="_blank" href="https://github.com/giswqs/earthengine-py-notebooks/tree/master/AssetManagement/export_ImageCollection.ipynb"><img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" /> View source on GitHub</a></td>
<td><a target="_blank" href="https://nbviewer.jupyter.org/github/giswqs/earthengine-py-notebooks/blob/master/AssetManagement/export_ImageCollection.ipynb"><img width=26px src="https://upload.wikimedia.org/wikipedia/commons/thumb/3/38/Jupyter_logo.svg/883px-Jupyter_logo.svg.png" />Notebook Viewer</a></td>
<td><a target="_blank" href="https://mybinder.org/v2/gh/giswqs/earthengine-py-notebooks/master?filepath=AssetManagement/export_ImageCollection.ipynb"><img width=58px src="https://mybinder.org/static/images/logo_social.png" />Run in binder</a></td>
<td><a target="_blank" href="https://colab.research.google.com/github/giswqs/earthengine-py-notebooks/blob/master/AssetManagement/export_ImageCollection.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" /> Run in Google Colab</a></td>
</table>
## Install Earth Engine API
Install the [Earth Engine Python API](https://developers.google.com/earth-engine/python_install) and [geehydro](https://github.com/giswqs/geehydro). The **geehydro** Python package builds on the [folium](https://github.com/python-visualization/folium) package and implements several methods for displaying Earth Engine data layers, such as `Map.addLayer()`, `Map.setCenter()`, `Map.centerObject()`, and `Map.setOptions()`.
The magic command `%%capture` can be used to hide output from a specific cell.
```
# %%capture
# !pip install earthengine-api
# !pip install geehydro
```
Import libraries
```
import ee
import folium
import geehydro
```
Authenticate and initialize Earth Engine API. You only need to authenticate the Earth Engine API once. Uncomment the line `ee.Authenticate()`
if you are running this notebook for this first time or if you are getting an authentication error.
```
# ee.Authenticate()
ee.Initialize()
```
## Create an interactive map
This step creates an interactive map using [folium](https://github.com/python-visualization/folium). The default basemap is the OpenStreetMap. Additional basemaps can be added using the `Map.setOptions()` function.
The optional basemaps can be `ROADMAP`, `SATELLITE`, `HYBRID`, `TERRAIN`, or `ESRI`.
```
Map = folium.Map(location=[40, -100], zoom_start=4)
Map.setOptions('HYBRID')
```
## Add Earth Engine Python script
```
# USDA NAIP ImageCollection
collection = ee.ImageCollection('USDA/NAIP/DOQQ')
# create an roi
polys = ee.Geometry.Polygon(
[[[-99.29615020751953, 46.725459351792374],
[-99.2116928100586, 46.72404725733022],
[-99.21443939208984, 46.772037733479884],
[-99.30267333984375, 46.77321343419932]]])
# create a FeatureCollection based on the roi and center the map
centroid = polys.centroid()
lng, lat = centroid.getInfo()['coordinates']
print("lng = {}, lat = {}".format(lng, lat))
Map.setCenter(lng, lat, 12)
fc = ee.FeatureCollection(polys)
# filter the ImageCollection using the roi
naip = collection.filterBounds(polys)
naip_2015 = naip.filterDate('2015-01-01', '2015-12-31')
mosaic = naip_2015.mosaic()
# print out the number of images in the ImageCollection
count = naip_2015.size().getInfo()
print("Count: ", count)
# add the ImageCollection and the roi to the map
vis = {'bands': ['N', 'R', 'G']}
Map.addLayer(mosaic,vis)
Map.addLayer(fc)
# export the ImageCollection to Google Drive
downConfig = {'scale': 30, "maxPixels": 1.0E13, 'driveFolder': 'image'} # scale means resolution.
img_lst = naip_2015.toList(100)
for i in range(0, count):
image = ee.Image(img_lst.get(i))
name = image.get('system:index').getInfo()
# print(name)
task = ee.batch.Export.image(image, name, downConfig)
task.start()
```
## Display Earth Engine data layers
```
Map.setControlVisibility(layerControl=True, fullscreenControl=True, latLngPopup=True)
Map
```
| github_jupyter |
```
import tensorflow as tf
# Get the GPU device name.
device_name = tf.test.gpu_device_name()
# The device name should look like the following:
if device_name == '/device:GPU:0':
print('Found GPU at: {}'.format(device_name))
else:
raise SystemError('GPU device not found')
import torch
# If there's a GPU available...
if torch.cuda.is_available():
# Tell PyTorch to use the GPU.
device = torch.device("cuda")
print('There are %d GPU(s) available.' % torch.cuda.device_count())
print('We will use the GPU:', torch.cuda.get_device_name(0))
# If not...
else:
print('No GPU available, using the CPU instead.')
device = torch.device("cpu")
#Parse
import pandas as pd
df = pd.read_csv('nsmc/ratings_train.txt', sep='\t')
test = pd.read_csv('nsmc/ratings_test.txt', sep='\t')
df.loc[df.label == 0].sample(5)[['document', 'label']]
# Get the lists of sentences and their labels.
sentences = df.document.values
labels = df.label.values
print(sentences) #리스트에 넣어줍니다.
import torch
from torch import nn, Tensor
from torch.optim import Optimizer
from torch.utils.data import DataLoader, RandomSampler, DistributedSampler, random_split
from torch.nn import CrossEntropyLoss
from pytorch_lightning.core.lightning import LightningModule
from pytorch_lightning.metrics.functional import accuracy, precision, recall
from transformers.modeling_bert import BertModel
from transformers import AdamW
""""""
#ImportError: cannot import name 'SAVE_STATE_WARNING'
#SAVE_STATE_WARNING from PyTorch, which only exists in 1.5.0
#downgrade pytorch version to 1.5.0
""""""
#토크나이저
from transformers import AutoTokenizer, AutoModelWithLMHead
tokenizer = AutoTokenizer.from_pretrained("beomi/kcbert-base")
#from transformers import AutoTokenizer
#tokenizer = AutoTokenizer.from_pretrained('bert-base-multilingual-cased')
# Print the original sentence.
print(' Original: ', sentences[0])
# Print the sentence split into tokens.
print('Tokenized: ', tokenizer.tokenize(sentences[0]))
# Print the sentence mapped to token ids.
print('Token IDs: ', tokenizer.convert_tokens_to_ids(tokenizer.tokenize(sentences[0])))
# BERT의 입력 형식에 맞게 토큰을 추가해줍니다
sentences = ["[CLS] " + str(sent) + " [SEP]" for sent in sentences]
sentences[:10]
tokenized_texts = [tokenizer.tokenize(sentence) for sentence in sentences]
print(tokenized_texts[:3])
from keras.preprocessing.sequence import pad_sequences
MAX_LEN = 128
input_ids = [tokenizer.convert_tokens_to_ids(x) for x in tokenized_texts]
input_ids = pad_sequences(input_ids, maxlen=MAX_LEN, dtype='long', truncating='post', padding='post')
input_ids[0]
""""""
#max_len = 128
# For every sentence...
#for sent in sentences:
# Tokenize the text and add `[CLS]` and `[SEP]` tokens.
# input_ids = tokenizer.encode(sent, add_special_tokens=True)
# Update the maximum sentence length.
# max_len = max(max_len, len(input_ids))
print('Max sentence length: ', MAX_LEN)
""""""
# Tokenize all of the sentences and map the tokens to thier word IDs.
input_ids = []
attention_masks = []
# For every sentence...
for sent in sentences:
# `encode_plus` will:
# (1) Tokenize the sentence.
# (2) Prepend the `[CLS]` token to the start.
# (3) Append the `[SEP]` token to the end.
# (4) Map tokens to their IDs.
# (5) Pad or truncate the sentence to `max_length`
# (6) Create attention masks for [PAD] tokens.
encoded_dict = tokenizer.encode_plus(
sent, # Sentence to encode.
add_special_tokens = True, # Add '[CLS]' and '[SEP]'
max_length = 64, # Pad & truncate all sentences.
pad_to_max_length = True,
truncation=True, #to eliminate the warning(truncation true로 설정하라는 에러를 받음)
return_attention_mask = True, # Construct attn. masks.
return_tensors = 'pt', # Return pytorch tensors.
)
# Add the encoded sentence to the list.
input_ids.append(encoded_dict['input_ids'])
# And its attention mask (simply differentiates padding from non-padding).
attention_masks.append(encoded_dict['attention_mask'])
# Convert the lists into tensors.
input_ids = torch.cat(input_ids, dim=0)
attention_masks = torch.cat(attention_masks, dim=0)
labels = torch.tensor(labels)
# Print sentence 0, now as a list of IDs.
print('Original: ', sentences[0])
print('Token IDs:', input_ids[0])
#Training & Validation Split
from torch.utils.data import TensorDataset, random_split
# Combine the training inputs into a TensorDataset.
dataset = TensorDataset(input_ids, attention_masks, labels)
# Create a 90-10 train-validation split.
# Calculate the number of samples to include in each set.
train_size = int(0.9 * len(dataset))
val_size = len(dataset) - train_size
# Divide the dataset by randomly selecting samples.
train_dataset, val_dataset = random_split(dataset, [train_size, val_size])
print('{:>5,} training samples'.format(train_size))
print('{:>5,} validation samples'.format(val_size))
from torch.utils.data import DataLoader, RandomSampler, SequentialSampler
# The DataLoader needs to know our batch size for training, so we specify it
# here. For fine-tuning BERT on a specific task, the authors recommend a batch
# size of 16 or 32.
batch_size = 32
# Create the DataLoaders for our training and validation sets.
# We'll take training samples in random order.
train_dataloader = DataLoader(
train_dataset, # The training samples.
sampler = RandomSampler(train_dataset), # Select batches randomly
batch_size = batch_size # Trains with this batch size.
)
# For validation the order doesn't matter, so we'll just read them sequentially.
validation_dataloader = DataLoader(
val_dataset, # The validation samples.
sampler = SequentialSampler(val_dataset), # Pull out batches sequentially.
batch_size = batch_size # Evaluate with this batch size.
)
#Train Our Classification Model
#BertForSequenceClassification
# normal BERT model with an added single linear layer on top for classification that we will use as a sentence classifier
from transformers import BertForSequenceClassification, AdamW, BertConfig
# Load BertForSequenceClassification, the pretrained BERT model with a single
# linear classification layer on top.
model = BertForSequenceClassification.from_pretrained(
"beomi/kcbert-base", # Use the 12-layer BERT model, with an uncased vocab.
num_labels = 2, # The number of output labels--2 for binary classification.
# You can increase this for multi-class tasks.
output_attentions = False, # Whether the model returns attentions weights.
output_hidden_states = False, # Whether the model returns all hidden-states.
)
# Tell pytorch to run this model on the GPU.
model.cuda()
#이 에러가 계속 나는데 이걸 해결하는 법을 모르겠어요
```
# 문제
Some weights of the model checkpoint at bert-base-uncased were not used when initializing
라는 에러를 계속 받고 있다. 찾아보니 fine-tuning이 제대로 되지 않았다는 뜻이라는데
```python
model = BertForSequenceClassification.from_pretrained(
"beomi/kcbert-base", # Use the 12-layer BERT model, with an uncased vocab.
num_labels = 2, # The number of output labels--2 for binary classification.
# You can increase this for multi-class tasks.
output_attentions = False, # Whether the model returns attentions weights.
output_hidden_states = False, # Whether the model returns all hidden-states.
)
```
라는 코드로 맨 위에 리니어 클래시피케이션을 수행하는 레이어를 쌓았기에 문제가 되지 않는다 생각해서 진행했다. 하지만 관련 이슈에 대해서는 더 찾아보기로 할게요.
# epoch
- 1epoch에서 90% accuracy를 달성했다...?
```
# Get all of the model's parameters as a list of tuples.
params = list(model.named_parameters())
print('The BERT model has {:} different named parameters.\n'.format(len(params)))
print('==== Embedding Layer ====\n')
for p in params[0:5]:
print("{:<55} {:>12}".format(p[0], str(tuple(p[1].size()))))
print('\n==== First Transformer ====\n')
for p in params[5:21]:
print("{:<55} {:>12}".format(p[0], str(tuple(p[1].size()))))
print('\n==== Output Layer ====\n')
for p in params[-4:]:
print("{:<55} {:>12}".format(p[0], str(tuple(p[1].size()))))
# Note: AdamW is a class from the huggingface library (as opposed to pytorch)
# I believe the 'W' stands for 'Weight Decay fix"
optimizer = AdamW(model.parameters(),
lr = 2e-5, # args.learning_rate - default is 5e-5, our notebook had 2e-5
eps = 1e-8 # args.adam_epsilon - default is 1e-8.
)
```
# Optimizer & Learning Rate Scheduler
For the purposes of fine-tuning, the authors recommend choosing from the following values (from Appendix A.3 of the BERT paper):
Batch size: 16, 32
Learning rate (Adam): 5e-5, 3e-5, 2e-5
Number of epochs: 2, 3, 4
파라미터 조절가능. batch size는 사이즈 때문에 작게 잡는게 좋을 것 같다. 64는 안됨.
```
from transformers import get_linear_schedule_with_warmup
# Number of training epochs. The BERT authors recommend between 2 and 4.
# We chose to run for 4, but we'll see later that this may be over-fitting the
# training data.
epochs = 4
# Total number of training steps is [number of batches] x [number of epochs].
# (Note that this is not the same as the number of training samples).
total_steps = len(train_dataloader) * epochs
# Create the learning rate scheduler.
scheduler = get_linear_schedule_with_warmup(optimizer,
num_warmup_steps = 0, # Default value in run_glue.py
num_training_steps = total_steps)
```
# Training Loop
Training:
Unpack our data inputs and labels
Load data onto the GPU for acceleration
Clear out the gradients calculated in the previous pass.
In pytorch the gradients accumulate by default (useful for things like RNNs) unless you explicitly clear them out.
Forward pass (feed input data through the network)
Backward pass (backpropagation)
Tell the network to update parameters with optimizer.step()
Track variables for monitoring progress
```
import numpy as np
# Function to calculate the accuracy of our predictions vs labels
def flat_accuracy(preds, labels):
pred_flat = np.argmax(preds, axis=1).flatten()
labels_flat = labels.flatten()
return np.sum(pred_flat == labels_flat) / len(labels_flat)
import time
import datetime
def format_time(elapsed):
'''
Takes a time in seconds and returns a string hh:mm:ss
'''
# Round to the nearest second.
elapsed_rounded = int(round((elapsed)))
# Format as hh:mm:ss
return str(datetime.timedelta(seconds=elapsed_rounded))
import random
import numpy as np
# This training code is based on the `run_glue.py` script here:
# https://github.com/huggingface/transformers/blob/5bfcd0485ece086ebcbed2d008813037968a9e58/examples/run_glue.py#L128
# Set the seed value all over the place to make this reproducible.
seed_val = 42
random.seed(seed_val)
np.random.seed(seed_val)
torch.manual_seed(seed_val)
torch.cuda.manual_seed_all(seed_val)
# We'll store a number of quantities such as training and validation loss,
# validation accuracy, and timings.
training_stats = []
# Measure the total training time for the whole run.
total_t0 = time.time()
# For each epoch...
for epoch_i in range(0, epochs):
# ========================================
# Training
# ========================================
# Perform one full pass over the training set.
print("")
print('======== Epoch {:} / {:} ========'.format(epoch_i + 1, epochs))
print('Training...')
# Measure how long the training epoch takes.
t0 = time.time()
# Reset the total loss for this epoch.
total_train_loss = 0
# Put the model into training mode. Don't be mislead--the call to
# `train` just changes the *mode*, it doesn't *perform* the training.
# `dropout` and `batchnorm` layers behave differently during training
# vs. test (source: https://stackoverflow.com/questions/51433378/what-does-model-train-do-in-pytorch)
model.train()
# For each batch of training data...
for step, batch in enumerate(train_dataloader):
# Progress update every 40 batches.
if step % 40 == 0 and not step == 0:
# Calculate elapsed time in minutes.
elapsed = format_time(time.time() - t0)
# Report progress.
print(' Batch {:>5,} of {:>5,}. Elapsed: {:}.'.format(step, len(train_dataloader), elapsed))
# Unpack this training batch from our dataloader.
#
# As we unpack the batch, we'll also copy each tensor to the GPU using the
# `to` method.
#
# `batch` contains three pytorch tensors:
# [0]: input ids
# [1]: attention masks
# [2]: labels
b_input_ids = batch[0].to(device)
b_input_mask = batch[1].to(device)
b_labels = batch[2].to(device)
# Always clear any previously calculated gradients before performing a
# backward pass. PyTorch doesn't do this automatically because
# accumulating the gradients is "convenient while training RNNs".
# (source: https://stackoverflow.com/questions/48001598/why-do-we-need-to-call-zero-grad-in-pytorch)
model.zero_grad()
# Perform a forward pass (evaluate the model on this training batch).
# The documentation for this `model` function is here:
# https://huggingface.co/transformers/v2.2.0/model_doc/bert.html#transformers.BertForSequenceClassification
# It returns different numbers of parameters depending on what arguments
# arge given and what flags are set. For our useage here, it returns
# the loss (because we provided labels) and the "logits"--the model
# outputs prior to activation.
loss, logits = model(b_input_ids,
token_type_ids=None,
attention_mask=b_input_mask,
labels=b_labels)
# Accumulate the training loss over all of the batches so that we can
# calculate the average loss at the end. `loss` is a Tensor containing a
# single value; the `.item()` function just returns the Python value
# from the tensor.
total_train_loss += loss.item()
# Perform a backward pass to calculate the gradients.
loss.backward()
# Clip the norm of the gradients to 1.0.
# This is to help prevent the "exploding gradients" problem.
torch.nn.utils.clip_grad_norm_(model.parameters(), 1.0)
# Update parameters and take a step using the computed gradient.
# The optimizer dictates the "update rule"--how the parameters are
# modified based on their gradients, the learning rate, etc.
optimizer.step()
# Update the learning rate.
scheduler.step()
# Calculate the average loss over all of the batches.
avg_train_loss = total_train_loss / len(train_dataloader)
# Measure how long this epoch took.
training_time = format_time(time.time() - t0)
print("")
print(" Average training loss: {0:.2f}".format(avg_train_loss))
print(" Training epcoh took: {:}".format(training_time))
# ========================================
# Validation
# ========================================
# After the completion of each training epoch, measure our performance on
# our validation set.
print("")
print("Running Validation...")
t0 = time.time()
# Put the model in evaluation mode--the dropout layers behave differently
# during evaluation.
model.eval()
# Tracking variables
total_eval_accuracy = 0
total_eval_loss = 0
nb_eval_steps = 0
# Evaluate data for one epoch
for batch in validation_dataloader:
# Unpack this training batch from our dataloader.
#
# As we unpack the batch, we'll also copy each tensor to the GPU using
# the `to` method.
#
# `batch` contains three pytorch tensors:
# [0]: input ids
# [1]: attention masks
# [2]: labels
b_input_ids = batch[0].to(device)
b_input_mask = batch[1].to(device)
b_labels = batch[2].to(device)
# Tell pytorch not to bother with constructing the compute graph during
# the forward pass, since this is only needed for backprop (training).
with torch.no_grad():
# Forward pass, calculate logit predictions.
# token_type_ids is the same as the "segment ids", which
# differentiates sentence 1 and 2 in 2-sentence tasks.
# The documentation for this `model` function is here:
# https://huggingface.co/transformers/v2.2.0/model_doc/bert.html#transformers.BertForSequenceClassification
# Get the "logits" output by the model. The "logits" are the output
# values prior to applying an activation function like the softmax.
(loss, logits) = model(b_input_ids,
token_type_ids=None,
attention_mask=b_input_mask,
labels=b_labels)
# Accumulate the validation loss.
total_eval_loss += loss.item()
# Move logits and labels to CPU
logits = logits.detach().cpu().numpy()
label_ids = b_labels.to('cpu').numpy()
# Calculate the accuracy for this batch of test sentences, and
# accumulate it over all batches.
total_eval_accuracy += flat_accuracy(logits, label_ids)
# Report the final accuracy for this validation run.
avg_val_accuracy = total_eval_accuracy / len(validation_dataloader)
print(" Accuracy: {0:.2f}".format(avg_val_accuracy))
# Calculate the average loss over all of the batches.
avg_val_loss = total_eval_loss / len(validation_dataloader)
# Measure how long the validation run took.
validation_time = format_time(time.time() - t0)
print(" Validation Loss: {0:.2f}".format(avg_val_loss))
print(" Validation took: {:}".format(validation_time))
# Record all statistics from this epoch.
training_stats.append(
{
'epoch': epoch_i + 1,
'Training Loss': avg_train_loss,
'Valid. Loss': avg_val_loss,
'Valid. Accur.': avg_val_accuracy,
'Training Time': training_time,
'Validation Time': validation_time
}
)
print("")
print("Training complete!")
print("Total training took {:} (h:mm:ss)".format(format_time(time.time()-total_t0)))
PATH = '/home/aiffel-dj57/project'
torch.save(model, PATH + 'kcbert.pt') # 전체 모델 저장
torch.save(model.state_dict(), PATH + 'model_state_dict.pt') # 모델 객체의 state_dict 저장
torch.save({
'kcbert': model.state_dict(),
'optimizer': optimizer.state_dict()
}, PATH + 'all.tar') # 여러 가지 값 저장, 학습 중 진행 상황 저장을 위해 epoch, loss 값 등 일반 scalar값 저장 가능
#이건 나중에 실행해보려고 했던거니 신경안쓰셔도 됩니다.
import pandas as pd
# Display floats with two decimal places.
pd.set_option('precision', 2)
# Create a DataFrame from our training statistics.
df_stats = pd.DataFrame(data=training_stats)
# Use the 'epoch' as the row index.
df_stats = df_stats.set_index('epoch')
# A hack to force the column headers to wrap.
#df = df.style.set_table_styles([dict(selector="th",props=[('max-width', '70px')])])
# Display the table.
df_stats
''''''
import matplotlib.pyplot as plt
% matplotlib inline
import seaborn as sns
# Use plot styling from seaborn.
sns.set(style='darkgrid')
# Increase the plot size and font size.
sns.set(font_scale=1.5)
plt.rcParams["figure.figsize"] = (12,6)
# Plot the learning curve.
plt.plot(df_stats['Training Loss'], 'b-o', label="Training")
plt.plot(df_stats['Valid. Loss'], 'g-o', label="Validation")
# Label the plot.
plt.title("Training & Validation Loss")
plt.xlabel("Epoch")
plt.ylabel("Loss")
plt.legend()
plt.xticks([1, 2, 3, 4])
plt.show()
''''''
#테스트셋에 평가해보기
import pandas as pd
# Load the dataset into a pandas dataframe.
test = pd.read_csv('nsmc/ratings_test.txt', sep='\t')
# Report the number of sentences.
print('Number of test sentences: {:,}\n'.format(df.shape[0]))
# Create sentence and label lists
sentences2 = test.document.values
labels2 = test.label.values
#왜 50,000이지 좀따 해결해볼게요.... #test sentence 수가 점점 늘어난다 뭔가 이상하다 ㅠㅠㅠㅠ(재부팅함)
# Print the original sentence.
print(' Original: ', sentences2[0])
# Print the sentence split into tokens.
print('Tokenized: ', tokenizer.tokenize(sentences2[0]))
# Print the sentence mapped to token ids.
print('Token IDs: ', tokenizer.convert_tokens_to_ids(tokenizer.tokenize(sentences2[0])))
# BERT의 입력 형식에 맞게 토큰을 추가해줍니다
sentences = ["[CLS] " + str(sent) + " [SEP]" for sent in sentences2]
sentences[:10]
tokenized_texts = [tokenizer.tokenize(sentence) for sentence in sentences]
print(tokenized_texts[:3])
from keras.preprocessing.sequence import pad_sequences
MAX_LEN = 128
input_ids = [tokenizer.convert_tokens_to_ids(x) for x in tokenized_texts]
input_ids = pad_sequences(input_ids, maxlen=MAX_LEN, dtype='long', truncating='post', padding='post')
input_ids[0]
#영어는 토큰화가 엉망으로 되네
# Tokenize all of the sentences and map the tokens to thier word IDs.
input_ids = []
attention_masks = []
# For every sentence...
for sent in sentences:
# `encode_plus` will:
# (1) Tokenize the sentence.
# (2) Prepend the `[CLS]` token to the start.
# (3) Append the `[SEP]` token to the end.
# (4) Map tokens to their IDs.
# (5) Pad or truncate the sentence to `max_length`
# (6) Create attention masks for [PAD] tokens.
encoded_dict = tokenizer.encode_plus(
sent, # Sentence to encode.
add_special_tokens = True, # Add '[CLS]' and '[SEP]'
max_length = 64, # Pad & truncate all sentences.
pad_to_max_length = True,
return_attention_mask = True, # Construct attn. masks.
return_tensors = 'pt', # Return pytorch tensors.
)
# Add the encoded sentence to the list.
input_ids.append(encoded_dict['input_ids'])
# And its attention mask (simply differentiates padding from non-padding).
attention_masks.append(encoded_dict['attention_mask'])
# Convert the lists into tensors.
input_ids = torch.cat(input_ids, dim=0)
attention_masks = torch.cat(attention_masks, dim=0)
labels = torch.tensor(labels)
# Set the batch size.
batch_size = 32
# Create the DataLoader.
prediction_data = TensorDataset(input_ids, attention_masks, labels)
prediction_sampler = SequentialSampler(prediction_data)
prediction_dataloader = DataLoader(prediction_data, sampler=prediction_sampler, batch_size=batch_size)
# Prediction on test set
print('Predicting labels for {:,} test sentences...'.format(len(input_ids)))
# Put model in evaluation mode
model.eval()
# Tracking variables
predictions , true_labels = [], []
# Predict
for batch in prediction_dataloader:
# Add batch to GPU
batch = tuple(t.to(device) for t in batch)
# Unpack the inputs from our dataloader
b_input_ids, b_input_mask, b_labels = batch
# Telling the model not to compute or store gradients, saving memory and
# speeding up prediction
with torch.no_grad():
# Forward pass, calculate logit predictions
outputs = model(b_input_ids, token_type_ids=None,
attention_mask=b_input_mask)
logits = outputs[0]
# Move logits and labels to CPU
logits = logits.detach().cpu().numpy()
label_ids = b_labels.to('cpu').numpy()
# Store predictions and true labels
predictions.append(logits)
true_labels.append(label_ids)
print(' DONE.')
print('Positive samples: %d of %d (%.2f%%)' % (df.label.sum(), len(df.label), (df.label.sum() / len(df.label) * 100.0)))
```
```python
from sklearn.metrics import matthews_corrcoef
matthews_set = []
# Evaluate each test batch using Matthew's correlation coefficient
print('Calculating Matthews Corr. Coef. for each batch...')
# For each input batch...
for i in range(len(true_labels)):
# The predictions for this batch are a 2-column ndarray (one column for "0"
# and one column for "1"). Pick the label with the highest value and turn this
# in to a list of 0s and 1s.
pred_labels_i = np.argmax(predictions[i], axis=1).flatten()
# Calculate and store the coef for this batch.
matthews = matthews_corrcoef(true_labels[i], pred_labels_i)
matthews_set.append(matthews)
```
# CHECK POINT
```python
import re
import emoji
from soynlp.normalizer import repeat_normalize
emojis = ''.join(emoji.UNICODE_EMOJI.keys())
pattern = re.compile(f'[^ .,?!/@$%~%·∼()\x00-\x7Fㄱ-ㅣ가-힣{emojis}]+')
url_pattern = re.compile(
r'https?:\/\/(www\.)?[-a-zA-Z0-9@:%._\+~#=]{1,256}\.[a-zA-Z0-9()]{1,6}\b([-a-zA-Z0-9()@:%_\+.~#?&//=]*)')
def clean(x):
x = pattern.sub(' ', x)
x = url_pattern.sub('', x)
x = x.strip()
x = repeat_normalize(x, num_repeats=2)
return x
```
kcBERT는 전처리 과정에서 이 과정을 실행하라고 함
| github_jupyter |
```
import seaborn as sns
import numpy as np
import json
from pprint import pprint
import matplotlib.pyplot as plt
```
## All cases, training time
```
warm_up_full_pretrain_perf = []
with open("../50_episodes_1_9/run_1/warm_up_full_pretrain_performances.json", "r") as full_pretrain_perf_file:
warm_up_full_pretrain_perf.extend(json.load(full_pretrain_perf_file))
no_warm_up_full_pretrain_perf = []
with open("../50_episodes_1_9/run_1/full_pretrain_performances.json", "r") as full_pretrain_perf_file:
no_warm_up_full_pretrain_perf.extend(json.load(full_pretrain_perf_file))
warm_up_full_scratch_perf = []
with open("../50_episodes_1_9/run_1/full_scratch_performances.json", "r") as full_scratch_perf_file:
warm_up_full_scratch_perf.extend(json.load(full_scratch_perf_file))
no_warm_full_scratch_perf = []
with open("../50_episodes_1_9/run_1/no_warm_up_full_scratch_performances.json", "r") as no_warm_full_scratch_perf_file:
no_warm_full_scratch_perf.extend(json.load(no_warm_full_scratch_perf_file))
print(len(warm_up_full_pretrain_perf))
print(len(no_warm_up_full_pretrain_perf))
print(len(warm_up_full_scratch_perf))
print(len(no_warm_full_scratch_perf))
warm_up_full_pretrain_mean = []
no_warm_up_full_pretrain_mean = []
warm_up_full_scratch_mean = []
no_warm_full_scratch_mean = []
for i in range(95):
warm_up_full_pretrain_mean.append([warm_up_full_pretrain_perf[i][str(j)] for j in range(50)])
no_warm_up_full_pretrain_mean.append([no_warm_up_full_pretrain_perf[i][str(j)] for j in range(50)])
warm_up_full_scratch_mean.append([warm_up_full_scratch_perf[i][str(j)] for j in range(50)])
no_warm_full_scratch_mean.append([no_warm_full_scratch_perf[i][str(j)] for j in range(50)])
plt.figure(figsize=(15, 9))
sns.set(font_scale=3)
sns.set_style("whitegrid")
data = np.asarray(warm_up_full_pretrain_mean)
ax = sns.tsplot(data=data, ci=[90], color='cyan', marker='*', markersize=15, markevery=5, linewidth=3.0, legend=True, condition="TF - WS")
data = np.asarray(no_warm_up_full_pretrain_mean)
ax = sns.tsplot(data=data, ci=[85], color='blue', marker='d', markersize=15, markevery=5, linewidth=3.0, legend=True, condition="TF - no WS")
data = np.asarray(warm_up_full_scratch_mean)
ax = sns.tsplot(data=data, ci=[85], color='red', marker='X', markersize=15, markevery=5, linewidth=3.0, legend=True, condition="no TF - WS")
data = np.asarray(no_warm_full_scratch_mean)
ax = sns.tsplot(data=data, ci=[90], color='orange', marker='^', markersize=15, markevery=5, linewidth=3.0, legend=True, condition="no TF - no WS")
ax.set_xlabel('Number of Epochs', weight='bold', size=35)
ax.set_ylabel('Success Rate', weight='bold', size=35)
sns.plt.title('Learning curve over training data set', weight='bold', size=35)
plt.xlim((0, 50))
plt.ylim((-0.02, 1))
plt.setp(ax.get_legend().get_texts(), fontsize="35")
plt.savefig('all_cases_learning_curve_training_data_set_200_reps_color.png', dpi=200, bbox_inches="tight", pad_inches=0)
plt.show()
```
## All cases, testing time
```
warm_up_full_pretrain_test_perf = []
with open("../50_episodes_1_9/run_1/warm_up_full_pretrain_test_performances.json", "r") as warm_up_full_pretrain_test_perf_file:
warm_up_full_pretrain_test_perf.extend(json.load(warm_up_full_pretrain_test_perf_file))
no_warm_up_full_pretrain_test_perf = []
with open("../50_episodes_1_9/run_1/full_pretrain_test_performances.json", "r") as full_pretrain_test_perf_file:
no_warm_up_full_pretrain_test_perf.extend(json.load(full_pretrain_test_perf_file))
warm_up_full_scratch_test_perf = []
with open("../50_episodes_1_9/run_1/full_scratch_test_performances.json", "r") as full_scratch_test_perf_file:
warm_up_full_scratch_test_perf.extend(json.load(full_scratch_test_perf_file))
no_warm_full_scratch_test_perf = []
with open("../50_episodes_1_9/run_1/no_warm_up_full_scratch_test_performances.json", "r") as no_warm_full_scratch_test_perf_file:
no_warm_full_scratch_test_perf.extend(json.load(no_warm_full_scratch_test_perf_file))
print(len(warm_up_full_pretrain_test_perf))
print(len(no_warm_up_full_pretrain_test_perf))
print(len(warm_up_full_scratch_test_perf))
print(len(no_warm_full_scratch_test_perf))
warm_up_full_pretrain_test_mean = []
no_warm_up_full_pretrain_test_mean = []
warm_up_full_scratch_test_mean = []
no_warm_full_scratch_test_mean = []
for i in range(95):
warm_up_full_pretrain_test_mean.append([warm_up_full_pretrain_test_perf[i][str(j)] for j in range(50)])
no_warm_up_full_pretrain_test_mean.append([no_warm_up_full_pretrain_test_perf[i][str(j)] for j in range(50)])
warm_up_full_scratch_test_mean.append([warm_up_full_scratch_test_perf[i][str(j)] for j in range(50)])
no_warm_full_scratch_test_mean.append([no_warm_full_scratch_test_perf[i][str(j)] for j in range(50)])
plt.figure(figsize=(15, 9))
sns.set(font_scale=3)
sns.set_style("whitegrid")
data = np.asarray(warm_up_full_pretrain_test_mean)
ax = sns.tsplot(data=data, ci=[85], color='cyan', marker='*', markersize=15, markevery=5, legend=True, condition="TF - WS")
data = np.asarray(no_warm_up_full_pretrain_test_mean)
ax = sns.tsplot(data=data, ci=[85], color='blue', marker='d', markersize=15, markevery=5, legend=True, condition="TF - no WS")
data = np.asarray(warm_up_full_scratch_test_mean)
ax = sns.tsplot(data=data, ci=[85], color='red', marker='X', markersize=15, markevery=5, linewidth=3.0, legend=True, condition="no TF - WS")
data = np.asarray(no_warm_full_scratch_test_mean)
ax = sns.tsplot(data=data, ci=[85], color='orange', marker='^', markersize=15, markevery=5, linewidth=3.0, legend=True, condition="no TF - no WS")
ax.set_xlabel('Number of Epochs', weight='bold', size=35)
ax.set_ylabel('Success Rate', weight='bold', size=35)
sns.plt.title('Learning curve over testing data set', weight='bold', size=35)
plt.xlim((0,50))
plt.ylim((-0.02, 1))
plt.setp(ax.get_legend().get_texts(), fontsize="35")
plt.savefig('all_cases_learning_curve_testing_data_set_200_reps_color.png', dpi=200, bbox_inches="tight", pad_inches=0)
plt.show()
```
| github_jupyter |
# GLM Example
Generalized linear models are a generalization of linear regression. In linear regression, responses are modeled as coming from gaussian distributions, each of equal variance, with each distribution's mean given by a linear combination of the predictor variables. GLMs allow other distributions where the variance might vary with the mean, and they allow the mean to be given by a function of the linear predictor instead of taking the linear predictor directly.
## Boston housing prices
We'll demonstrate by using GLMs to predict the median price of a home in Boston. The variables are crime rate, zoning information,
proportion of non-retail business, etc. This dataset has median prices in Boston for 1972. Even though the data is pretty old, the methodology for analytics is valid for more recent datasets.
The dataset is from Kaggle. https://www.kaggle.com/c/boston-housing. For tutorials use only.
## Housing Values in Suburbs of Boston in 1972
The <font color='red'>medv</font> variable is the target variable.
### Data description
The Boston data frame has 506 rows and 14 columns.
This data frame contains the following columns:
1. __crim__: per capita crime rate by town.
2. __zn__: proportion of residential land zoned for lots over 25,000 sq.ft.
3. __indus__: proportion of non-retail business acres per town.
4. __chas__: Charles River dummy variable (1 if tract bounds river; 0 otherwise).
5. __nox__: nitrogen oxides concentration (parts per 10 million).
6. __rm__: average number of rooms per dwelling.
7. __age__: proportion of owner-occupied units built prior to 1940.
8. __dis__: weighted mean of distances to five Boston employment centres.
9. __rad__: index of accessibility to radial highways.
10. __tax__: full-value property-tax rate per \$10000
11. __ptratio__: pupil-teacher ratio by town.
12. __black__: 1000(Bk - 0.63)^2 where Bk is the proportion of blacks by town.
13. __lstat__: lower status of the population (percent).
14. __medv__: median value of owner-occupied homes in $1000s.
</td></tr></table>
### Factoids
The prices in Boston across years are below. If we had a historical dataset, an analysis could be done to account for the macro trends as well.
The second graph shows the intuition we have with respect to prices in relation to crime rate. It is expected that house prices will be lower in areas where crime rates are higher.
The third figure is a chart showing how inflation may affect prices. So, for deeper analysis and prediction, we may want to consider inflation.
In this notebook, these factors are not considered. They are here to demonstrate the need for deep domain analysis.
<table><tr>
<td><img src="images/boston_prices_by_year.png" alt="Boston home prices" title="Boston housing prices" style="float:left;" /></td>
<td><img src="images/Crime-Rate-and-Median-House-Prices.png" alt="Boston home prices" title="Boston housing prices" /></td>
<td><img src="images/Inflation_Adjusted_Housing_Prices_1890_2006.jpg" alt="Inflation adjusted prices" title="Inflation adjusted prices" style="float:left;" />
</td></tr></table>
In this notebook, we will use the dataset for Boston housing prices and predict the price based on numerous factors.
```
from hana_ml import dataframe
from hana_ml.algorithms.pal import regression
import numpy as np
import matplotlib.pyplot as plt
import logging
```
## Load data
The data is loaded into 4 tables, for full, training, validation, and test sets:
<li>BOSTON_HOUSING_PRICES</li>
<li>BOSTON_HOUSING_PRICES_TRAINING</li>
<li>BOSTON_HOUSING_PRICES_VALIDATION</li>
<li>BOSTON_HOUSING_PRICES_TEST</li>
To do that, a connection is created and passed to the loader.
There is a config file, config/e2edata.ini that controls the connection parameters and whether or not to reload the data from scratch. In case the data is already loaded, there would be no need to load the data. A sample section is below. If the config parameter, reload_data is true then the tables for test, training, and validation are (re-)created and data inserted into them.
#########################<br>
[hana]<br>
url=host.sjc.sap.corp<br>
user=username<br>
passwd=userpassword<br>
port=3xx15<br>
<br>
[bostonhousingdataset]<br>
reload_data=true
#########################<br>
## Define Datasets - Training, validation, and test sets
Data frames represent data in HANA and HANA-side queries on that data, so computation on large data sets in HANA can happen in HANA. Trying to bring the entire data set into the client may be impractical or impossible for large data sets.
The original/full dataset is split into training, test and validation sets. In the example below, they reside in different tables.
```
from hana_ml.algorithms.pal.utility import DataSets, Settings
url, port, user, pwd = Settings.load_config("../../config/e2edata.ini")
connection_context = dataframe.ConnectionContext(url, port, user, pwd)
full_set, training_set, validation_set, test_set = DataSets.load_boston_housing_data(connection_context, force=True)
```
## Simple Exploration
Let us look at the number of rows in the data set.
```
print('Number of rows in full set: {}'.format(full_set.count()))
print('Number of rows in training set: {}'.format(training_set.count()))
print('Number of rows in validation set: {}'.format(validation_set.count()))
print('Number of rows in test set: {}'.format(test_set.count()))
```
### Let's look at the columns
```
print(full_set.columns)
```
### Let's look at the data types
```
full_set.dtypes()
```
### Set up the features and labels for the model
```
features=['CRIM', 'ZN', 'INDUS', 'CHAS', 'NOX', 'RM', 'AGE', 'DIS', 'RAD', 'TAX', 'PTRATIO', 'BLACK', 'LSTAT']
label='MEDV'
```
# Create model using training data
The first GLM we create will use the default settings of `family='gaussian'` and `link='identity'`. A GLM using these settings is equivalent to linear regression.
## Preprocessing
SAP HANA Predictive Analytics Library takes DOUBLE and INTEGER data types for most numeric types. Since we have DECIMALs and TINYINTs in our data set, we cast them to the types required by PAL.
```
# Cast to correct types so PAL can consume it.
dfts = training_set.cast(['CRIM', "ZN", "INDUS", "NOX", "RM", "AGE", "DIS", "PTRATIO", "BLACK", "LSTAT", "MEDV"], "DOUBLE")
dfts = dfts.cast(["CHAS", "RAD", "TAX"], "INTEGER")
dfts = dfts.to_head("ID")
dfts.head(5).collect()
```
## Create the model
```
linear_model = regression.GLM()
linear_model.fit(dfts, key='ID', features=features, label=label)
```
Let's see how well this model does. We'll compute the R^2 score of its predictions on the test set.
```
df_test = test_set.cast(['CRIM', "ZN", "INDUS", "NOX", "RM", "AGE", "DIS", "PTRATIO", "BLACK", "LSTAT", "MEDV"], "DOUBLE")
df_test = df_test.cast(["CHAS", "RAD", "TAX"], "INTEGER")
df_test = df_test.to_head("ID")
linear_model.score(df_test, key='ID', features=features, label=label)
```
Let's try a few others. We'll experiment with gamma distributions and inverse or logarithmic link functions.
```
gaussian_idlink = regression.GLM(family='gaussian', link='identity')
gamma_invlink = regression.GLM(family='gamma', link='inverse')
gaussian_invlink = regression.GLM(family='gaussian', link='inverse')
gamma_loglink = regression.GLM(family='gamma', link='log')
gaussian_loglink = regression.GLM(family='gaussian', link='log')
for model in [gaussian_idlink, gamma_invlink, gaussian_invlink, gamma_loglink, gaussian_loglink]:
model.fit(dfts, key='ID', features=features, label=label)
print(model.score(df_test, key='ID', features=features, label=label))
```
An inverse link seems to help model our data better. Our error terms seem to be modeled better by our original choice of `family='gaussian'` than `family='gamma'`.
| github_jupyter |
<a href="https://colab.research.google.com/github/NikolaZubic/AppliedGameTheoryHomeworkSolutions/blob/main/domaci4.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# ČETVRTI DOMAĆI ZADATAK iz predmeta "Primenjena teorija igara" (Applied Game Theory)
a) Poređenje rezultata kod pristupa Q-learning, SARSA i Monte Karlo metoda za igru Ajnc (BlackJack): https://github.com/NikolaZubic/AppliedGameTheoryHomeworkSolutions/blob/main/Homework%203/domaci3.ipynb
b) Razvoj bota za igranje igre Iks-Oks (Tic Tac Toe) koristeći "Q-learning" pristup.
# a)
Suštinska razlika između navedenih metoda jeste u načinu ažuriranja Q-vrijednosti. Uzeto je da Q-learning i SARSA koriste Temporal-Difference Learning, sa time da kod Q-learninga u narednom stanju uvijek biramo najbolju akciju, dok kod SARSA algoritma biramo akciju na osnovu politike odlučivanja.<br>
Dobijeni su sledeći rezultati:
Q LEARNING:<br>
Player wins: 3681<br>
Dealer wins: 5706<br>
Player wins percentage = 39.21%<br>
<br>
SARSA:<br>
Player wins: 3647<br>
Dealer wins: 5740<br>
Player wins percentage = 38.85%<br>
<br>
MONTE-CARLO METHODS:<br>
Player wins: 3314<br>
Dealer wins: 6186<br>
Player wins percentage = 34.88%<br>
# b)
# Potrebni import-i
```
import gym
import numpy as np
from gym import spaces
import pickle
import os
```
# Definisanje Iks-Oks okruženja koristeći "Open AI Gym" toolkit
```
class TicTacToeEnvironment(gym.Env):
# Because of human-friendly output
metadata = {'render.modes': ['human']}
def __init__(self, player_1, player_2):
"""
Board is predefined to a 3 x 3 grid.
We keep track of whether the game is over.
When initializing the TicTacToeEnvironment, we set game_over flag to 'False'.
:param player_1: First player
:param player_2: Second player
"""
self.observation_space = spaces.Discrete(3 * 3)
self.action_space = spaces.Discrete(9)
self.board = np.zeros((3, 3))
self.player_1 = player_1
self.player_2 = player_2
self.game_over = False
self.reset()
# Let player_1 play first
self.current_player = 1
# Board string representation
self.board_str = None
def reset(self):
# Resets the environment after one game.
self.board = np.zeros((3, 3)) # set board to zeros
self.board_str = None # set board string representation to null
self.game_over = False
self.current_player = 1
def get_board(self):
# getter for current board
return self.board
def get_board_str(self):
# synchronize board string representation with current board state
self.board_str = str(self.board.reshape(3 * 3))
return self.board_str
def get_free_positions(self):
# return positions on the board that are free / not occupied
positions = [(i, j) for i in range(3) for j in range(3) if self.board[i, j] == 0]
return positions
def update_state(self, new_position):
"""
Update the current state of the board. First player puts '1' on the board, second player puts '-1' on the board.
:param new_position: from set { (0, 0), (0, 1), (0, 2), (1, 0,), (1, 1), (1, 2), (2, 0), (2, 1), (2, 2) }
:return: None
"""
self.board[new_position] = self.current_player
# If current player puts 1 ("X") on the board, next move is player 2 who puts -1 "O" on the board.
if self.current_player == 1:
self.current_player = -1
else:
self.current_player = 1
def check_game_status(self):
"""
Check if game has finished.
:return: 1 if player 1 has won,
-1 if player 2 has won,
0 if draw,
None if game hasn't finished.
"""
for i in range(3):
if sum(self.board[i, :]) == 3:
self.game_over = True
return 1
if sum(self.board[i, :]) == -3:
self.game_over = True
return - 1
for i in range(3):
if sum(self.board[:, i]) == 3:
self.game_over = True
return 1
if sum(self.board[:, i]) == -3:
self.game_over = True
return -1
main_diagonal_sum = sum([self.board[i, i] for i in range(3)])
anti_diagonal_sum = sum([self.board[i, 3 - i - 1] for i in range(3)])
diagonal_sum = max(abs(main_diagonal_sum), abs(anti_diagonal_sum))
if diagonal_sum == 3:
self.game_over = True
if diagonal_sum == 3 or anti_diagonal_sum == 3:
return 1
else:
return - 1
# DRAW
if len(self.get_free_positions()) == 0:
self.game_over = True
return 0
self.game_over = False
return None
def step(self, action):
"""
Performs one action.
:param action: from set { (0, 0), (0, 1), (0, 2), (1, 0,), (1, 1), (1, 2), (2, 0), (2, 1), (2, 2) }
:return: current board state, reward, boolean indicating whether the game is over,
information about won the game if over
"""
if self.game_over:
return self.board, 0, True, None
self.update_state(action)
current_game_status = self.check_game_status()
if current_game_status is not None:
if current_game_status == 1:
reward = 1
info = {"Result": "Player 1 won the game."}
elif current_game_status == -1:
reward = -1
info = {"Result": "Player 2 won the game."}
else:
# DRAW
reward = 0
info = {"Result": "Draw."}
return self.board, reward, self.game_over, info
return self.board, None, self.game_over, None
def render(self):
for i in range(0, 3):
print('-------------')
out = '| '
for j in range(0, 3):
token = ''
if self.board[i, j] == 1:
token = 'X'
if self.board[i, j] == -1:
token = 'O'
if self.board[i, j] == 0:
token = ' '
out += token + ' | '
print(out)
print('-------------')
```
# Definisanje Q-learning agenta
```
class QLearningAgent(object):
def __init__(self, gamma=0.9, LAMBDA=0.1, epsilon=0.1):
self.gamma = gamma # Discounting factor
self.LAMBDA = LAMBDA # Used for filtering q-values, learning rate
self.epsilon = epsilon # Used in epsilon-greedy policy
self.states = []
self.states_values = {} # KEY = CURRENT_BOARD_STATE, VALUE = number
@staticmethod
def get_board_str(board):
return str(board.reshape(3 * 3))
def reset(self):
self.states = []
def add_state(self, state):
self.states.append(state)
def get_max_action(self, positions, current_board, symbol):
"""
Choose best action.
:param positions: set of free positions
:param current_board: current environment board
:param symbol: current player symbol (1 = "X" = PC and -1 = "O" = HUMAN)
:return: action, for example (1, 1) which is in the center of 3x3 grid
"""
if np.random.uniform(0, 1) <= self.epsilon:
idx = np.random.choice(len(positions)) # choose index from the interval [0, 9]
action = positions[idx] # choose certain action, for example positions[1] = (0, 2)
else:
action = None
value_max = - np.inf
for pos in positions:
next_board = current_board.copy()
next_board[pos] = symbol
next_board_str = self.get_board_str(next_board)
if self.states_values.get(next_board_str) is None:
action_value = 0
else:
action_value = self.states_values.get(next_board_str)
if action_value >= value_max:
value_max = action_value
action = pos
return action
def compute_q(self, reward):
"""
Compute q values at the end of the game, all states are saved in self.states.
:param reward: depends on whether the player has won/lost/draw the game
:return: None
"""
for current_board_state in reversed(self.states):
if self.states_values.get(current_board_state) is None:
self.states_values[current_board_state] = 0
q_current_s_a = self.states_values[current_board_state]
# compute new q
self.states_values[current_board_state] = q_current_s_a + self.LAMBDA * (self.gamma * reward -
q_current_s_a)
reward = self.states_values[current_board_state]
def save_policy(self):
file = open("saved_policy", "wb")
pickle.dump(self.states_values, file)
file.close()
def load_policy(self, file_name):
file = open(file_name, "rb")
self.states_values = pickle.load(file)
file.close()
```
# Definisanje igrača, unos sa tastature
```
class HumanPlayer(object):
@staticmethod
def act(positions):
while True:
user_input = input("['O' on move] x,y: ")
x, y = user_input.split(",")
x_int, y_int = int(x) - 1, int(y) - 1
pos = (x_int, y_int)
if pos in positions:
return pos
else:
print("Invalid move. Try again.")
```
# Treniranje agenta
```
def train(number_of_episodes):
player_1 = QLearningAgent()
player_2 = QLearningAgent()
agents = [player_1, player_2]
environment = TicTacToeEnvironment(player_1, player_2)
for i in range(number_of_episodes):
if i % 100000 == 0:
print("Episode {}".format(i))
environment.reset()
is_done = False
while not is_done:
for agent in agents:
if not is_done:
free_positions = environment.get_free_positions()
current_action = agent.get_max_action(free_positions, environment.get_board(),
environment.current_player)
current_board_state, reward, is_done, information = environment.step(current_action)
agent.add_state(agent.get_board_str(current_board_state))
if is_done:
if reward == 1:
# If player one wins, he gets the reward of 1
player_1.compute_q(1)
player_2.compute_q(0)
elif reward == -1:
# If player two wins, he gets the reward of 1
player_1.compute_q(0)
player_2.compute_q(1)
else:
# DRAW, computer gets smaller reward
player_1.compute_q(0.1)
player_2.compute_q(0.5)
# At the end, reset history of board states for both players
player_1.reset()
player_2.reset()
environment.reset()
player_1.save_policy()
```
# Pokretanje igre
```
def play_game():
player_1 = QLearningAgent()
player_1.load_policy("saved_policy")
player_2 = HumanPlayer()
agents = [player_1, player_2]
environment = TicTacToeEnvironment(player_1, player_2)
environment.reset()
is_done = False
environment.render()
while not is_done:
for agent in agents:
if agent == player_1:
action = agent.get_max_action(environment.get_free_positions(), environment.get_board(),
environment.current_player)
else:
action = agent.act(environment.get_free_positions())
state, reward, is_done, information = environment.step(action)
environment.render()
if is_done:
print(information['Result'])
break
```
# Glavni program
Preuzimanje istreniranog modela (treniran kroz **1 milion epizoda/partija**):
```
!wget "https://github.com/NikolaZubic/AppliedGameTheoryHomeworkSolutions/raw/main/Homework%204/saved_policy"
if __name__ == '__main__':
NUMBER_OF_EPISODES = 1000000
if not os.path.exists("saved_policy"):
train(number_of_episodes=NUMBER_OF_EPISODES)
play_game()
```
| github_jupyter |
# T81-558: Applications of Deep Neural Networks
**Module 7: Generative Adversarial Networks**
* Instructor: [Jeff Heaton](https://sites.wustl.edu/jeffheaton/), McKelvey School of Engineering, [Washington University in St. Louis](https://engineering.wustl.edu/Programs/Pages/default.aspx)
* For more information visit the [class website](https://sites.wustl.edu/jeffheaton/t81-558/).
# Module 7 Material
* **Part 7.1: Introduction to GANS for Image and Data Generation** [[Video]](https://www.youtube.com/watch?v=0QnCH6tlZgc&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_07_1_gan_intro.ipynb)
* Part 7.2: Implementing a GAN in Keras [[Video]](https://www.youtube.com/watch?v=T-MCludVNn4&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_07_2_Keras_gan.ipynb)
* Part 7.3: Face Generation with StyleGAN and Python [[Video]](https://www.youtube.com/watch?v=Wwwyr7cOBlU&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_07_3_style_gan.ipynb)
* Part 7.4: GANS for Semi-Supervised Learning in Keras [[Video]](https://www.youtube.com/watch?v=ZPewmEu7644&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_07_4_gan_semi_supervised.ipynb)
* Part 7.5: An Overview of GAN Research [[Video]](https://www.youtube.com/watch?v=cvCvZKvlvq4&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_07_5_gan_research.ipynb)
# Part 7.1: Introduction to GANS for Image and Data Generation
A generative adversarial network (GAN) is a class of machine learning systems invented by Ian Goodfellow in 2014. Two neural networks contest with each other in a game. Given a training set, this technique learns to generate new data with the same statistics as the training set. For example, a GAN trained on photographs can generate new photographs that look at least superficially authentic to human observers, having many realistic characteristics. Though originally proposed as a form of generative model for unsupervised learning, GANs have also proven useful for semi-supervised learning, fully supervised learning, and reinforcement learning. GANs were introduced in the following paper:
* Goodfellow, I., Pouget-Abadie, J., Mirza, M., Xu, B., Warde-Farley, D., Ozair, S., ... & Bengio, Y. (2014). [Generative adversarial nets](https://papers.nips.cc/paper/5423-generative-adversarial-nets.pdf). In *Advances in neural information processing systems* (pp. 2672-2680).
This paper used neural networks to automatically generate images for several datasets that we've seen previously: MINST and CIFAR. However, it also included the Toronto Face Dataset (a private dataset used by some researchers).

Only sub-figure D made use of convolutional neural networks. Figures A-C make use of fully connected neural networks. As we will see in this module, the role of convolutional neural networks with GANs was greatly increased.
A GAN is called a generative model because it generates new data. The overall process of a GAN is given by the following diagram.

| github_jupyter |
##### Copyright 2018 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Load text
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/tutorials/load_data/text"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/tutorials/load_data/text.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/tutorials/load_data/text.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/tutorials/load_data/text.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
</td>
</table>
This tutorial provides an example of how to use `tf.data.TextLineDataset` to load examples from text files. `TextLineDataset` is designed to create a dataset from a text file, in which each example is a line of text from the original file. This is potentially useful for any text data that is primarily line-based (for example, poetry or error logs).
In this tutorial, we'll use three different English translations of the same work, Homer's Illiad, and train a model to identify the translator given a single line of text.
## Setup
```
from __future__ import absolute_import, division, print_function, unicode_literals
try:
# %tensorflow_version only exists in Colab.
%tensorflow_version 2.x
except Exception:
pass
import tensorflow as tf
import tensorflow_datasets as tfds
import os
```
The texts of the three translations are by:
- [William Cowper](https://en.wikipedia.org/wiki/William_Cowper) — [text](https://storage.googleapis.com/download.tensorflow.org/data/illiad/cowper.txt)
- [Edward, Earl of Derby](https://en.wikipedia.org/wiki/Edward_Smith-Stanley,_14th_Earl_of_Derby) — [text](https://storage.googleapis.com/download.tensorflow.org/data/illiad/derby.txt)
- [Samuel Butler](https://en.wikipedia.org/wiki/Samuel_Butler_%28novelist%29) — [text](https://storage.googleapis.com/download.tensorflow.org/data/illiad/butler.txt)
The text files used in this tutorial have undergone some typical preprocessing tasks, mostly removing stuff — document header and footer, line numbers, chapter titles. Download these lightly munged files locally.
```
DIRECTORY_URL = 'https://storage.googleapis.com/download.tensorflow.org/data/illiad/'
FILE_NAMES = ['cowper.txt', 'derby.txt', 'butler.txt']
for name in FILE_NAMES:
text_dir = tf.keras.utils.get_file(name, origin=DIRECTORY_URL+name)
parent_dir = os.path.dirname(text_dir)
parent_dir
```
## Load text into datasets
Iterate through the files, loading each one into its own dataset.
Each example needs to be individually labeled, so use `tf.data.Dataset.map` to apply a labeler function to each one. This will iterate over every example in the dataset, returning (`example, label`) pairs.
```
def labeler(example, index):
return example, tf.cast(index, tf.int64)
labeled_data_sets = []
for i, file_name in enumerate(FILE_NAMES):
lines_dataset = tf.data.TextLineDataset(os.path.join(parent_dir, file_name))
labeled_dataset = lines_dataset.map(lambda ex: labeler(ex, i))
labeled_data_sets.append(labeled_dataset)
```
Combine these labeled datasets into a single dataset, and shuffle it.
```
BUFFER_SIZE = 50000
BATCH_SIZE = 64
TAKE_SIZE = 5000
all_labeled_data = labeled_data_sets[0]
for labeled_dataset in labeled_data_sets[1:]:
all_labeled_data = all_labeled_data.concatenate(labeled_dataset)
all_labeled_data = all_labeled_data.shuffle(
BUFFER_SIZE, reshuffle_each_iteration=False)
```
You can use `tf.data.Dataset.take` and `print` to see what the `(example, label)` pairs look like. The `numpy` property shows each Tensor's value.
```
for ex in all_labeled_data.take(5):
print(ex)
```
## Encode text lines as numbers
Machine learning models work on numbers, not words, so the string values need to be converted into lists of numbers. To do that, map each unique word to a unique integer.
### Build vocabulary
First, build a vocabulary by tokenizing the text into a collection of individual unique words. There are a few ways to do this in both TensorFlow and Python. For this tutorial:
1. Iterate over each example's `numpy` value.
2. Use `tfds.features.text.Tokenizer` to split it into tokens.
3. Collect these tokens into a Python set, to remove duplicates.
4. Get the size of the vocabulary for later use.
```
tokenizer = tfds.features.text.Tokenizer()
vocabulary_set = set()
for text_tensor, _ in all_labeled_data:
some_tokens = tokenizer.tokenize(text_tensor.numpy())
vocabulary_set.update(some_tokens)
vocab_size = len(vocabulary_set)
vocab_size
```
### Encode examples
Create an encoder by passing the `vocabulary_set` to `tfds.features.text.TokenTextEncoder`. The encoder's `encode` method takes in a string of text and returns a list of integers.
```
encoder = tfds.features.text.TokenTextEncoder(vocabulary_set)
```
You can try this on a single line to see what the output looks like.
```
example_text = next(iter(all_labeled_data))[0].numpy()
print(example_text)
encoded_example = encoder.encode(example_text)
print(encoded_example)
```
Now run the encoder on the dataset by wrapping it in `tf.py_function` and passing that to the dataset's `map` method.
```
def encode(text_tensor, label):
encoded_text = encoder.encode(text_tensor.numpy())
return encoded_text, label
def encode_map_fn(text, label):
return tf.py_function(encode, inp=[text, label], Tout=(tf.int64, tf.int64))
all_encoded_data = all_labeled_data.map(encode_map_fn)
```
## Split the dataset into test and train batches
Use `tf.data.Dataset.take` and `tf.data.Dataset.skip` to create a small test dataset and a larger training set.
Before being passed into the model, the datasets need to be batched. Typically, the examples inside of a batch need to be the same size and shape. But, the examples in these datasets are not all the same size — each line of text had a different number of words. So use `tf.data.Dataset.padded_batch` (instead of `batch`) to pad the examples to the same size.
```
train_data = all_encoded_data.skip(TAKE_SIZE).shuffle(BUFFER_SIZE)
train_data = train_data.padded_batch(BATCH_SIZE, padded_shapes=([-1],[]))
test_data = all_encoded_data.take(TAKE_SIZE)
test_data = test_data.padded_batch(BATCH_SIZE, padded_shapes=([-1],[]))
```
Now, `test_data` and `train_data` are not collections of (`example, label`) pairs, but collections of batches. Each batch is a pair of (*many examples*, *many labels*) represented as arrays.
To illustrate:
```
sample_text, sample_labels = next(iter(test_data))
sample_text[0], sample_labels[0]
```
Since we have introduced a new token encoding (the zero used for padding), the vocabulary size has increased by one.
```
vocab_size += 1
```
## Build the model
```
model = tf.keras.Sequential()
```
The first layer converts integer representations to dense vector embeddings. See the [Word Embeddings](../../tutorials/sequences/word_embeddings) tutorial for more details.
```
model.add(tf.keras.layers.Embedding(vocab_size, 64))
```
The next layer is a [Long Short-Term Memory](http://colah.github.io/posts/2015-08-Understanding-LSTMs/) layer, which lets the model understand words in their context with other words. A bidirectional wrapper on the LSTM helps it to learn about the datapoints in relationship to the datapoints that came before it and after it.
```
model.add(tf.keras.layers.Bidirectional(tf.keras.layers.LSTM(64)))
```
Finally we'll have a series of one or more densely connected layers, with the last one being the output layer. The output layer produces a probability for all the labels. The one with the highest probability is the models prediction of an example's label.
```
# One or more dense layers.
# Edit the list in the `for` line to experiment with layer sizes.
for units in [64, 64]:
model.add(tf.keras.layers.Dense(units, activation='relu'))
# Output layer. The first argument is the number of labels.
model.add(tf.keras.layers.Dense(3, activation='softmax'))
```
Finally, compile the model. For a softmax categorization model, use `sparse_categorical_crossentropy` as the loss function. You can try other optimizers, but `adam` is very common.
```
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
```
## Train the model
This model running on this data produces decent results (about 83%).
```
model.fit(train_data, epochs=3, validation_data=test_data)
eval_loss, eval_acc = model.evaluate(test_data)
print('\nEval loss: {:.3f}, Eval accuracy: {:.3f}'.format(eval_loss, eval_acc))
```
| github_jupyter |
```
import numpy as np
import mrcfile
import matplotlib.pyplot as plt
%matplotlib notebook
class SubgraphLoader():
def __init__(self,
inputShape,
sampleShape,
keys,
keyToPath,
labelFile):
self.inputShape = inputShape
self.sampleShape = sampleShape
self.keys = keys
self.keyToPath = keyToPath
self.particles = self._parseParticles(labelFile)
def getMicrograph(self, key):
""" Load micrograph. """
with mrcfile.open(self.keyToPath(key), permissive=True) as mrc:
data = mrc.data
return data
def boxContains(self, point, sampleId):
dimH, dimW = self.sampleShape
x, y = point
cx, cy = sampleId
return (cx*dimW < x < cx*dimW + dimW) and (cy*dimH < y < cy*dimH + dimH)
def _generateSubgraph(self, key):
""" Generate the subimages for a given micrograph. """
retDict = {}
h, w = self.inputShape
dimH, dimW = self.sampleShape
data = self.getMicrograph(key)
for idxh in range(int(h/dimH)):
for idxw in range(int(w/dimW)):
retDict[(idxh,idxw)] = data[idxh*dimH:idxh*dimH+dimH,
idxw*dimW:idxw*dimW+dimW]
return retDict
def _parseParticles(self, file):
"""
Read in the particles for all micrographs.
This will need to be edited if you change your key types.
"""
with open(file, "r") as f:
particles = f.readlines()
particleData = [particle.split()[0:3] for particle in particles[17:-1]]
particleDict = {}
for x in particleData:
key = int(x[0][18:22])
value = tuple(map(float, x[1:]))
particleDict.setdefault(key, []).append(value)
return particleDict
def getSubgraphAnnotation(self, shift = True):
"""
Searches through the particle list a dictionary which maps
(micrographKey, subgraphKey) -> [particles in subgraph]
subgraphKey - is the x,y position of a subgraph within the grid formed by the subgraphs over the micrograph.
shift specifies if you want the absolute or relative posiotions.
"""
subDict = {}
h, w = self.inputShape
dimH, dimW = self.sampleShape
for micrograph in self.keys:
for idxh in range(int(h/dimH)):
for idxw in range(int(w/dimW)):
subgraph_particles = np.array(
list(
filter(
lambda x : boxContains(x, (idxh, idxw)),
self.particles[micrograph]
)
)
)
try:
subDict[(micrograph, idxh, idxw)] = subgraph_particles - np.array([idxh*dimH, idxw*dimW])*shift
except:
continue
return subDict
def getSubgraphs(self):
subDict = {}
for micrograph in self.keys:
subgraphs = self._generateSubgraph(micrograph)
for k,v in subgraphs.items():
subDict[(micrograph, *k)] = v
return subDict
loader = SubgraphLoader (
(3710, 3838),
(512, 512),
[1, 2, 3],
lambda x: f'../data/full/stack_{str(x).zfill(4)}_2x_dfpt.mrc',
'../data/full/particles.star'
)
subgraphs = loader.getSubgraphs()
annotations = loader.getSubgraphAnnotation()
abs_annotations = loader.getSubgraphAnnotation(False)
key = (1,0,0)
plt.figure()
plt.imshow(subgraphs[key], cmap='gray')
plt.scatter(*zip(*annotations[key]), c='red', alpha=0.3, s=260)
particles = loader._parseParticles('../data/full/particles.star')
from matplotlib.patches import Rectangle
outdir = '../data/full/'
fname = 'stack_0001_2x_dfpt.mrc'
with mrcfile.open(f'{outdir}{fname}', permissive=True) as mrc:
data = mrc.data
key = (1,1,1)
dim = 512
plt.figure()
plt.imshow(data)
plt.scatter(*zip(*abs_annotations[key]), c='red', alpha=0.3, s=10)
plt.scatter(*zip(*particles[key[0]]), c='black', alpha=0.3, s=10)
rect = Rectangle((key[1]*dim,key[2]*dim), dim, dim,linewidth=1,edgecolor='r',facecolor='none')
# Add the patch to the Axes
plt.gca().add_patch(rect)
```
| github_jupyter |
<div style="width:1000 px">
<div style="float:right; width:98 px; height:98px;">
<img src="https://raw.githubusercontent.com/Unidata/MetPy/master/metpy/plots/_static/unidata_150x150.png" alt="Unidata Logo" style="height: 98px;">
</div>
<h1>Using Siphon to get NEXRAD Level 3 data from a TDS</h1>
<h3>Unidata Python Workshop</h3>
<div style="clear:both"></div>
</div>
<hr style="height:2px;">
<div style="float:right; width:250 px"><img src="https://upload.wikimedia.org/wikipedia/commons/4/4d/Siphoning.JPG" alt="Siphoning" style="height: 300px;"></div>
### Objectives
1. Learn more about Siphon
2. Use the RadarServer class to retrieve radar data from a TDS
3. Plot this data using numpy arrays and matplotlib
In this example, we'll focus on interacting with the Radar Query Service to retrieve radar data.
**But first!**
Bookmark these resources for when you want to use Siphon later!
+ [latest Siphon documentation](http://siphon.readthedocs.org/en/latest/)
+ [Siphon github repo](https://github.com/Unidata/siphon)
+ [TDS documentation](http://www.unidata.ucar.edu/software/thredds/v4.6/tds/TDS.html)
## Querying the server
First, we point at the top level of the Radar Query Service (the "Radar Server") to see what radar collections are available:
```
from siphon.catalog import TDSCatalog
cat = TDSCatalog('http://thredds.ucar.edu/thredds/radarServer/catalog.xml')
list(cat.catalog_refs)
```
Next we create an instance of the `RadarServer` object to point at one of these collections. This downloads some top level metadata and sets things up so we can easily query the server.
```
from siphon.radarserver import RadarServer
rs = RadarServer(cat.catalog_refs['NEXRAD Level III Radar from IDD'].href)
```
We can use rs.variables to see a list of radar products available to view from this access URL.
```
print(sorted(rs.variables))
```
If you're not a NEXRAD radar expert, there is more information available within the metadata downloaded from the server. (**NOTE:** Only the codes above are valid for queries.)
```
sorted(rs.metadata['variables'])
```
We can also see a list of the stations. Each station has associated location information.
```
print(sorted(rs.stations))
rs.stations['TLX']
```
Next, we'll create a new query object to help request the data. Using the chaining methods, let's ask for reflectivity data at the lowest tilt (NOQ) from radar TLX (Oklahoma City) for the current time. We see that when the query is represented as a string, it shows the encoded URL.
```
from datetime import datetime
query = rs.query()
query.stations('TLX').time(datetime.utcnow()).variables('N0Q')
```
The query also supports time range queries, queries for closest to a lon/lat point, or getting all radars within a lon/lat box.
We can use the RadarServer instance to check our query, to make sure we have required parameters and that we have chosen valid station(s) and variable(s).
```
rs.validate_query(query)
```
Make the request, which returns an instance of TDSCatalog. This handles parsing the catalog
```
catalog = rs.get_catalog(query)
```
We can look at the datasets on the catalog to see what data we found by the query. We find one NIDS file in the return
```
catalog.datasets
```
### Exercise: Querying the radar server
We'll work through doing some more queries on the radar server. Some useful links:
- RadarQuery [documentation](https://siphon.readthedocs.org/en/latest/api/radarserver.html#siphon.radarserver.RadarQuery)
- Documentation on Python's [datetime.timedelta](https://docs.python.org/3.5/library/datetime.html#timedelta-objects)
See if you can write Python code for the following queries:
Get ZDR (differential reflectivity) for 3 days ago from the radar nearest to Hays, KS (lon -99.324403, lat 38.874929). **No map necessary!**
Get base reflectivity for the last two hours from all of the radars in Wyoming (call it the bounding box with lower left corner 41.008717, -111.056360 and upper right corner 44.981008, -104.042719)
## Pulling out the data
We can pull that dataset out of the dictionary and look at the available access URLs. We see URLs for OPeNDAP, CDMRemote, and HTTPServer (direct download).
```
ds = list(catalog.datasets.values())[0]
ds.access_urls
```
We'll use the CDMRemote reader in Siphon and pass it the appropriate access URL. (This will all behave identically to using the 'OPENDAP' access, if we replace the `Dataset` from Siphon with that from `netCDF4`).
```
from siphon.cdmr import Dataset
data = Dataset(ds.access_urls['CdmRemote'])
```
The CDMRemote reader provides an interface that is almost identical to the usual python NetCDF interface.
```
list(data.variables)
```
We pull out the variables we need for azimuth and range, as well as the data itself.
```
rng = data.variables['gate'][:]
az = data.variables['azimuth'][:]
ref = data.variables['BaseReflectivityDR'][:]
```
Then convert the polar coordinates to Cartesian using numpy
```
import numpy as np
x = rng * np.sin(np.deg2rad(az))[:, None]
y = rng * np.cos(np.deg2rad(az))[:, None]
ref = np.ma.array(ref, mask=np.isnan(ref))
```
Finally, we plot them up using matplotlib and cartopy.
```
%matplotlib inline
import matplotlib.pyplot as plt
import cartopy
from metpy.plots import ctables # For NWS colortable
# Create projection centered on the radar. This allows us to use x
# and y relative to the radar.
proj = cartopy.crs.LambertConformal(central_longitude=data.RadarLongitude,
central_latitude=data.RadarLatitude)
# New figure with specified projection
fig = plt.figure(figsize=(10, 10))
ax = fig.add_subplot(1, 1, 1, projection=proj)
# Grab state borders
state_borders = cartopy.feature.NaturalEarthFeature(
category='cultural', name='admin_1_states_provinces_lakes',
scale='50m', facecolor='none')
ax.add_feature(state_borders, edgecolor='black', linewidth=2, zorder=2)
# Counties
counties = cartopy.io.shapereader.Reader('../../data/counties.shp')
ax.add_geometries(counties.geometries(), cartopy.crs.PlateCarree(),
facecolor='None', edgecolor='grey', zorder=1)
# Set limits in lat/lon space
ax.set_extent([data.RadarLongitude - 2.5, data.RadarLongitude + 2.5,
data.RadarLatitude - 2.5, data.RadarLatitude + 2.5])
# Get the NWS typical reflectivity color table, along with an appropriate norm that
# starts at 5 dBz and has steps in 5 dBz increments
norm, cmap = ctables.registry.get_with_steps('NWSReflectivity', 5, 5)
mesh = ax.pcolormesh(x, y, ref, cmap=cmap, norm=norm, zorder=0)
```
### Exercise: Your turn to plot!
Try making your own plot of radar data. Various options here, but this is pretty open-ended. Some options to inspire you:
- Try working with Level II Data (All variables are three-dimensional and in a single file)
- Try plotting Storm Total Precipitation or Digital Accumulation for an area that recently had a bunch of rain
- Maybe plot Doppler velocity or some dual-pol data (ZDR or correlation coefficient) for recent severe weather. Bonus points for making a multi-panel plot.
| github_jupyter |

👇👇👇点击可跳转!
1. [5大变量类型的联系和区别](#5大变量类型的联系和区别)
- 1.1 [binary](#binary)
- 1.2 [nominal](#nominal)
- 1.3 [ordinal](#ordinal)
- 1.4 [interval](#interval)
- 1.5 [ratio](#ratio)
2. [元数据框架](#元数据框架)
- 2.1 [metadata介绍](#metadata介绍)
- 2.2 [metadata构建方法](#metadata构建方法)
- 2.3 [元数据分析](#元数据分析)
3. [其他](#其他)
<div class="alert alert-block alert-warning">
<center><b>项目日志(2022年2月18日)</b></center>
第二阶段:【数据管理&元数据分析】
**第二节阶段核心目的:**
- 掌握5大变量类型的适用场景和应用方法
- 理解运用元数据方法论,掌握元数据框架
- 熟练运用元数据方法分析数据
- Matplotlib强化巩固
**难度(最高5星):**⭐⭐⭐
**第二阶段周期:** 2022年4月8日 至 2022年4月15日(北京时间)
Good Luck!
</div>
第二阶段,我们正式进入【数据分析】层面,不管大家是在校生、求职党、转行党、甚至纯粹为了自我提升的伙伴们,千万不要觉得画画柱形图、饼图,找找变量之间有啥关系就是数据分析了,远不止于此。
在我们这个项目中,有58个特征,我们需要用58个特征去预测司机明年发起索赔的概率是多少,那么上一期,大家已经讨论分析过这样做的意义。
包括但不限于:
- **在司机层面:** 可以帮助司机减少不必要的保险支出,提升司机的理性投保意愿。
- **在公司层面:** 可以帮助公司对保险产品更合理的定价,帮助公司对客户在投保意愿上更准确的分析其特征,提高行业竞争力。
- **在发展层面:** 可以帮助公司缩小对客户特征的调研范围,极大节省研究成本,扩大精英特征的研究投入,对新用户索赔情况快速响应等等。
那么这一阶段,我们主要来去研究一下,什么是元数据!
- <font color=red>元数据是什么?</font>
- <font color=red>为什么在做项目时非常重要?</font>
- <font color=red>元数据管理后有什么好处?</font>
- <font color=red>元数据分析又有哪些亮点?</font>
这一期帮大家一一解答。
## 5大变量类型的联系和区别
首先,本项目为了保护客户信息,没有公开数据的58个分析特征的具体含义,说白了,我们不能从【列名】进行直观上的分析,我们无法判断哪些特征对预测是切实有效的!
举个例子:我想预测一个人是男生还是女生,我们可能通过头发长度、身高、体重等直观维度去进行判别,我们会根据自然的意识和经验对特征的影响程度评估,这样不太回走弯路。
但是,这里所有维度的含义被延长,我们不能主观判断,只能通过不同的变量类型,拆解整合,期待找到其中的联系。
通过`df.info()`可以看出各个维度的数据类型,有 `int64` 和 `float64` 两种,说明什么?
说明数据已经在放出来之前,经过了一轮处理,文本数据已经被转换为整数(也是为了保护客户的一种手段)。
那么,总共58个特征,它们都属于那种变量类型呢?
**英文叫做:levels of measurement**
大概如下:
- binary ==> 0 or 1
- 这一列只含有两种值:不是0就是1
- nominal ==> categorical 不含顺序
- 最常用的类别型变量,**不含任何排序**
- 举例:**中国、美国、英国等国家就属于nominal变量,他们之间不存在顺序关系**
- ordinal ==> categorical 含顺序
- 顾名思义,跟nominal相比,含顺序
- 举例:成绩排名;小明第一,小红第二,小王第三等等,**虽然但看这三个任命不存在任何排序关系,但是在这里就代表着成绩高低**,所以是ordinal变量。
- interval ==> continuous 无绝对零点
- 连续型数据,可以参与统计计算,比较大小,量化差距
- 举例:温度;温度是衡量冷热而存在的物理量,40度比20度高20度,但是我们不能说40度是20度的两倍,**这是因为interval数据没有绝对零点!**
- ratio ==> continuous 有绝对零点
- 连续型数据,可以参与统计计算,比较大小,加减乘除运算都可以
- 举例:身高;200cm比100cm长100cm,也可以说200cm是100cm的两倍。**这是因为ratio数据有绝对零点!**
看下面👇👇👇
来自我的小红书作品:http://xhslink.com/AYivVf
【请移步到我的作品中查看细节哦!】
<div class="alert alert-block alert-success"><b>Step 1 </b>完成下面习题
将以下内容分类为为binary、nominal、ordinal、interval、ratio。
</div>
| id | Question | Your Answer |
| :--: | :------------------------------------------------------: | :---------: |
| 1 | Flavors of frozen yogurt | |
| 2 | Amount of money in savings accounts | |
| 3 | Letter grades on an English essay | |
| 4 | Religions | |
| 5 | Commuting times to work | |
| 6 | Ages (in years) of art students | |
| 7 | Ice cream flavor preference | |
| 8 | Years of important historical events | |
| 9 | Instructors classified as: Easy, Difficult or Impossible | |


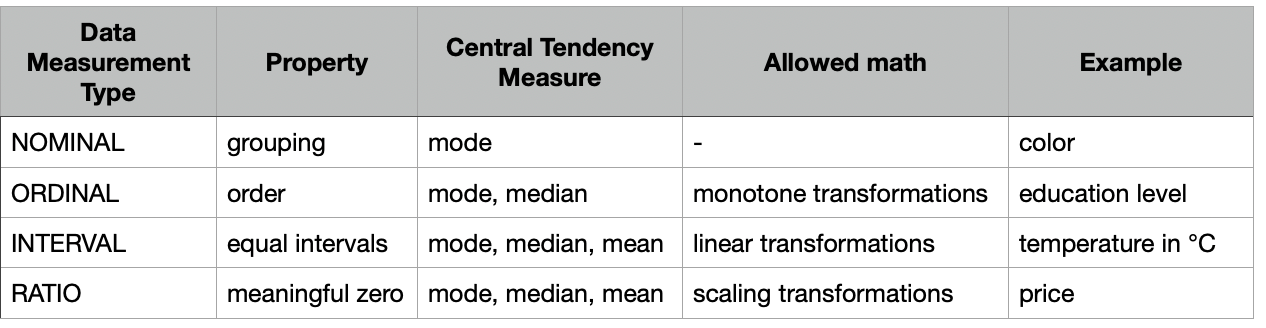


### 元数据
这个概念可能大家不是很熟悉,元宇宙最近这么火,大家都了解吧哈哈!
- 什么是元数据?
元数据(metadata),也叫做中介数据,或者叫数据的数据(data about data)。
- 为什么做项目时很重要?
因为做项目时,经常遇到维度很高,特征很多的数据,我们为了方便进一步的分析,必须对数据进行结构化的梳理,那么元数据是个很好的开始。
- 元数据有啥用?
主要来描述数据的属性(property)关于数据的组织、结构梳理,以及为以后的数据分析、可视化、建模都有很重要的意义。
<div class="alert alert-block alert-success"><b>Step 2</b>: 我们应该如何梳理高维度数据?别着急往下看,先思考一下!</div>
<div class="alert alert-block alert-info">
<p style="font-size:20px; display:inline">💡</p> 小提示:
高维数据换句话说就是数据的列数很多,所以我们应该围绕列来展开。
我们需要对所有列根据下列几个方面进行梳理:
- role ==> 序号|标签|特征
- category ==> 根据变量含义推断类别:车|人|地点|人工计算
- level ==> binary|nominal|ordinal|interval|ratio
- dtype ==> int|float|object...
- unique ==> 唯一值数量
- cardinality ==> high|low
- missing ==> 缺失值数量
- missing_percent ==> 缺失值百分比
- imputation ==> 填充缺失值手段
- keep ==> 是否保留
</div>
<div class="alert alert-block alert-success"><b>Step 2 (5分)</b>: 构建元数据结构表。
`from data_management import meta`
能复原我的output么?
</div>
```
import pandas as pd
import numpy as np
train = pd.read_csv('../../data/train.csv')
test = pd.read_csv('../../data/test.csv')
# train test 合并成一个 full set
fullset = "这里写你的答案!"
```
那么,变量类型的逻辑,帮大家梳理一下:
```diff
- role
+ 对每个列打上标签:序号|标签|特征
```
```diff
- category
+ 1. 列名含有ind表示individual的含义,返回"individual"
+ 2. 列名含有car表示car的含义,返回"car"
+ 3. 列名含有calc表示calculated的含义,返回"calculated"
```
```diff
- level
+ 1. 列名里有`bin`的或者这一列是target,我们归类为`binary` (if)
+ 2. 列名里有`cat`的或者这一列是id,归类为`nominal` (elif)
+ 3. 数据类型为`float`的是`interval` (elif)
+ 4. 数据类型是`int`的是`ordinal` (elif)
@ ordinal 解释一下:刨除开前面的情况,剩下的特征,如果他们的数据类型是`int`,我们认为他们是ordinal的,因为整数的排列是有序的,我们认为2>1,3>2等等,对应值的影响程度也满足这样的序列关系(虽然我们不知道值的具体含义)
```
**目标就是形成类似于下面的这样的表!**
【这个表有59行,相当于对数据的每一个维度进行梳理分析】
**注意:df的index设置为变量名称!**
```
from data_management import meta
metadata = meta(train,test)
```
<div class="alert alert-block alert-success"><b>Step 3</b>:
请基于meta,找出各个数据角色、变量类型各有多少个特征,目标形成下图的dataframe!
</div>
```
# 令起一行不要run
```
<div class="alert alert-block alert-success"><b>Step 4 (3分)</b>:
请基于meta,对连续型变量做统计描述型分析,使用describe函数!
- 请问连续型变量里面存在缺失值的有哪些?
- 请问连续型变量里面标准差最小的变量名称是什么?
- 请问连续型变量里面最大值大于4的变量名称是什么?
</div>
```
# 提示:用describe函数
stats = "在这里写你的答案"
stats
```
不要用眼睛看结果,要用代码逻辑输出哦!
```
# 请问连续型变量里面存在缺失值的有哪些?
"这里写你的代码"
# 请问连续型变量里面标准差最小的变量名称是什么?
"这里写你的代码"
# 请问连续型变量里面最大值大于4的变量名称是什么?
"这里写你的代码"
```
<div class="alert alert-block alert-success"><b>Step 5</b>:
根据下列代码和对应的结果,你有什么发现么?
</div>
你有什么发现么?
```
train.target.value_counts().plot(kind = 'bar')
```
<div class="alert alert-block alert-success"><b>Step 6</b>:
`from data_management import data_report`
看看这个函数都能得到什么信息!
</div>
```
from data_management import data_report
data_report(train,test,metadata,verbose=True)
```
补充:
如果你有余力,建议从下面链接中完成这100道Python Pandas puzzle!来为你下一阶段保驾护航!
链接:https://github.com/ajcr/100-pandas-puzzles/blob/master/100-pandas-puzzles.ipynb
[](https://imgtu.com/i/H65i3d)
<img src="https://s4.ax1x.com/2022/02/14/H65vxs.jpg" alt="image-20220214212808413" style="zoom:25%;" />
| github_jupyter |
# CaBi ML fitting - Random Forest
Trying out Random Forest here since it seems so good in terms of minimizing prediction error.
Using RandomizedSearchCV and GridSearchCV for hyperparameter tuning.
For now, not using PolynomialFeatures for transformation because (1) results are fairly good already and (2) computational speed is a bottleneck.
## 0. Data load, shaping, and split
* Read in data from AWS
* Encode time variable (day_of_year) as cyclical
* Split into Xtrain, Xtest, ytrain, ytest based on date
* Specify feature and target columns
```
# Read in data from AWS
from util_functions import *
import numpy as np
import pandas as pd
from pprint import pprint
import time
start_time = time.perf_counter()
set_env_path()
conn, cur = aws_connect()
# fullquery contains pretty much everything
fullquery = """
SELECT
EXTRACT(DOY FROM date) as day_of_year,
date,
year,
quarter,
month,
day_of_week,
daylight_hours,
apparenttemperaturehigh,
apparenttemperaturehightime,
apparenttemperaturelow,
apparenttemperaturelowtime,
precipintensitymaxtime,
sunrisetime,
sunsettime,
cloudcover,
dewpoint,
humidity,
precipaccumulation,
precipintensitymax,
precipprobability,
rain,
snow,
visibility,
windspeed,
us_holiday,
nats_single,
nats_double,
nats_attendance,
dc_bike_event,
dc_pop,
cabi_bikes_avail,
cabi_stations_alx,
cabi_stations_arl,
cabi_stations_ffx,
cabi_stations_mcn,
cabi_stations_mcs,
cabi_stations_wdc,
cabi_docks_alx,
cabi_docks_arl,
cabi_docks_ffx,
cabi_docks_mcn,
cabi_docks_mcs,
cabi_docks_wdc,
cabi_stations_tot,
cabi_docks_tot,
cabi_dur_empty_wdc,
cabi_dur_full_wdc,
cabi_dur_empty_arl,
cabi_dur_full_arl,
cabi_dur_full_alx,
cabi_dur_empty_alx,
cabi_dur_empty_mcs,
cabi_dur_full_mcs,
cabi_dur_full_mcn,
cabi_dur_empty_mcn,
cabi_dur_full_ffx,
cabi_dur_empty_ffx,
cabi_dur_empty_tot,
cabi_dur_full_tot,
cabi_active_members_day_key,
cabi_active_members_monthly,
cabi_active_members_annual,
cabi_trips_wdc_to_wdc,
cabi_trips_wdc_to_wdc_casual
from final_db"""
query = """
SELECT
EXTRACT(DOY FROM date) as day_of_year,
date,
year,
month,
day_of_week,
daylight_hours,
apparenttemperaturehigh,
apparenttemperaturehightime,
apparenttemperaturelow,
apparenttemperaturelowtime,
precipintensitymaxtime,
cloudcover,
dewpoint,
humidity,
precipaccumulation,
precipintensitymax,
precipprobability,
rain,
snow,
visibility,
windspeed,
us_holiday,
nats_single,
nats_double,
nats_attendance,
dc_bike_event,
dc_pop,
cabi_stations_tot,
cabi_docks_tot,
cabi_active_members_day_key,
cabi_active_members_monthly,
cabi_active_members_annual,
cabi_trips_wdc_to_wdc,
cabi_trips_wdc_to_wdc_casual
from final_db"""
pd.options.display.max_rows = None
pd.options.display.max_columns = None
df = pd.read_sql(query, con=conn)
# Setting date to index for easier splitting
df.set_index(df.date, drop=True, inplace=True)
df.index = pd.to_datetime(df.index)
print("We have {} instances and {} features".format(*df.shape))
```
#### Summary statistics
```
df.describe(percentiles=[.5]).round(3).transpose()
```
#### Which feature pairs are highly correlated?
```
def print_highly_correlated(df, features, threshold=0.75):
"""
Prints highly correlated feature pairs in df. Threshold set at 0.75 by default.
Selects pairs where abs(r) is above the threshold, puts them in a DataFrame,
making sure to avoid duplication, then sorts by abs(r) and prints.
"""
corr_df = df[features].corr()
correlated_features = np.where(np.abs(corr_df) > threshold)
correlated_features = [(corr_df.iloc[x,y], x, y) for x, y in zip(*correlated_features) if x != y and x < y]
s_corr_list = sorted(correlated_features, key=lambda x: -abs(x[0]))
print("There are {} feature pairs with pairwise correlation above {}".format(len(s_corr_list), threshold))
for v, i, j in s_corr_list:
cols = df[features].columns
print("{} and {} = {:0.3f}".format(corr_df.index[i], corr_df.columns[j], v))
print_highly_correlated(df, df.columns)
```
#### Encode day of year as cyclical
```
df['sin_day_of_year'] = np.sin(2*np.pi*df.day_of_year/365)
df['cos_day_of_year'] = np.cos(2*np.pi*df.day_of_year/365)
%matplotlib inline
df.sample(100).plot.scatter('sin_day_of_year','cos_day_of_year').set_aspect('equal')
```
#### Train/test split based on date
* Training dates = 2013-01-01 to 2016-12-31
* Test dates = 2017-01-01 to 2017-09-08
* New data (coincides with beginning of dockless pilot) = 2017-09-09 to present
```
# This can be tweaked, but we use ultimately use 5-fold cross-validation to pick the model
train = df.loc['2013-01-01':'2016-12-31']
test = df.loc['2017-01-01':'2017-09-08']
print(train.shape, test.shape)
tr = train.shape[0]
te = test.shape[0]
trpct = tr/(tr+te)
tepct = te/(tr+te)
print("{:0.3f} percent of the data is in the training set and {:0.3f} percent is in the test set".format(trpct, tepct))
# Specify columns to keep and drop for X and y
drop_cols = ['date']
y_cols = ['cabi_trips_wdc_to_wdc', 'cabi_trips_wdc_to_wdc_casual']
feature_cols = [col for col in df.columns if (col not in y_cols) & (col not in drop_cols)]
# X y split
Xtrain_raw = train[feature_cols]
# Our target variable here is all DC to DC trips
ytrain = train[y_cols[0]]
Xtest_raw = test[feature_cols]
ytest = test[y_cols[0]]
print(Xtrain_raw.shape, ytrain.shape, Xtest_raw.shape, ytest.shape)
```
## 1. Preprocessing
Trying it without any preprocessing first, but keeping PolynomialFeatures here in case we use it later.
```
from sklearn.preprocessing import PolynomialFeatures
'''This cell does nothing with PF(1), but will be useful for introducing PF'''
pf = PolynomialFeatures(1, include_bias=False)
Xtrain_pf_array = pf.fit_transform(Xtrain_raw)
Xtest_pf_array = pf.transform(Xtest_raw)
# Get feature names
Xtrain_cols = pf.get_feature_names(Xtrain_raw.columns)
# Convert arrays to dfs with the new pf column names
Xtrain = pd.DataFrame(Xtrain_pf_array, columns=Xtrain_cols)
Xtest = pd.DataFrame(Xtest_pf_array, columns=Xtrain_cols)
print(Xtrain.shape, Xtest.shape)
# Appending train and test to get full dataset for cross-validation
Xfull = Xtrain.append(Xtest)
yfull = ytrain.append(ytest)
print(Xfull.shape, yfull.shape)
print("Final vars=", Xfull.shape[1])
```
## 2. Model Hyperparameter Tuning
* Scoring functions
* RandomizedSearchCV
* GridSearchCV
#### Scoring functions
We need these functions to evaluate/score our models.
```
from sklearn.metrics import mean_squared_error as mse
from sklearn.metrics import r2_score
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import KFold
def score_model(model):
"""
Fits a model using the training set, predicts using the test set, and then calculates
and reports goodness of fit metrics.
"""
model.fit(Xtrain, ytrain)
yhat = model.predict(Xtest)
r2 = r2_score(ytest, yhat)
me = mse(ytest, yhat)
print("Results from {}: \nr2={:0.3f} \nMSE={:0.3f}".format(model, r2, me))
def cv_score(model, cv=5):
"""
Evaluates a model by 5-fold cross-validation and prints mean and 2*stdev of scores.
Shuffles before cross-validation but sets random_state=7 for reproducibility.
"""
kf = KFold(n_splits=cv, shuffle=True, random_state=7)
scores = cross_val_score(model, Xfull, yfull, cv=kf)
print(scores)
print("Accuracy: {:0.3f} (+/- {:0.3f})".format(scores.mean(), scores.std() * 2))
```
#### RandomizedSearchCV
We need to find appropriate values for our hyperparameters.
We can start by using RandomizedSearchCV to cast a wide net.
```
# Number of trees in random forest
n_estimators = [int(x) for x in np.linspace(200, 2000, 10)]
# Number of features to consider at every split
max_features = ['auto', 'sqrt']
# Maximum number of levels in tree
max_depth = [int(x) for x in np.linspace(10, 110, num = 11)]
max_depth.append(None)
# Minimum number of samples required to split a node
min_samples_split = [2, 5, 10]
# Minimum number of samples required at each leaf node
min_samples_leaf = [1, 2, 3, 4]
# Method of selecting samples for training each tree
bootstrap = [True, False]
# Create the parameter grid
param_distributions = {'n_estimators': n_estimators,
'max_features': max_features,
'max_depth': max_depth,
'min_samples_split': min_samples_split,
'min_samples_leaf': min_samples_leaf,
'bootstrap': bootstrap}
```
Altogether, there are 10 \* 2 \* 12 \* 3 \* 4 \* 2 = 5760 combinations.
We randomly sample 100 times per fold for a total of 300 fits.
```
from sklearn.model_selection import RandomizedSearchCV
from sklearn.ensemble import RandomForestRegressor
# Use the random grid to search for best hyperparameters
# First create the base model to tune
rf = RandomForestRegressor()
# Random search of parameters, using 3 fold cross validation,
# search across 100 different combinations, and use all available cores
cv = KFold(n_splits=3, shuffle=True, random_state=7)
ran_search = RandomizedSearchCV(estimator=rf,
param_distributions=param_distributions,
n_iter=100, cv=cv, verbose=3,
random_state=7, n_jobs=-1)
# Fit the random search model
ran_search.fit(Xtrain, ytrain)
```
We're interested in seeing if there's any improvement between the untuned default RF model and our new one
```
rf_random = ran_search.best_estimator_
print("Cross-validation score for base RF")
cv_score(rf)
print("\nCross-validation score for RF tuned by RandomizedSearchCV")
cv_score(rf_random)
print()
# What parameters are used?
pprint(ran_search.best_params_)
```
#### GridSearchCV
Slight increase in performance with the parameters suggested by RandomizedSearchCV.
Next, we use GridSearchCV which iterates over all of the possible combinations instead of randomly sampling.
*Note: User input required in the next section to create the GridSearch parameter grid based on RandomizedSearch results.*
```
from sklearn.model_selection import GridSearchCV
# Create the parameter grid based on the results of the random search
param_grid = {
'bootstrap': [False],
'max_depth': [50, 60, 70, 80, 90],
'max_features': ['sqrt'],
'min_samples_leaf': [1, 2, 3],
'min_samples_split': [2, 3, 4],
'n_estimators': [500, 1000, 1400, 1600, 2000]
}
# Create a base model
rf = RandomForestRegressor()
# Instantiate the grid search model
grid_search = GridSearchCV(estimator=rf, param_grid=param_grid,
cv=3, n_jobs=-1, verbose=3)
t = time.perf_counter()
# Fit the grid search to the data
grid_search.fit(Xtrain, ytrain)
elapsed_time = (time.perf_counter() - t)/60
print("This cell took {:0.2f} minutes to run".format(elapsed_time))
```
#### Cross-validation
How does this new model compare to the other model?
```
rf_best = grid_search.best_estimator_
print("Cross-validation score for untuned RF")
cv_score(rf)
print("\nCross-validation score for RF tuned by RandomizedSearchCV")
cv_score(rf_random)
print("\nCross-validation score for RF tuned by GridSearchCV")
cv_score(rf_best)
```
How do parameters differ between specifications?
```
print("RandomizedSearchCV params:")
pprint(ran_search.best_params_)
print("\nGridSearchCV params:")
pprint(grid_search.best_params_)
```
Which features are most important?
```
feature_importances = pd.DataFrame(rf_best.feature_importances_,
index = Xtrain.columns,
columns=['importance']).sort_values('importance', ascending=False)
feature_importances.head(20)
end_time = (time.perf_counter() - start_time)/60
print("This notebook took {:0.2f} minutes to run".format(end_time))
```
| github_jupyter |
```
!pip install --upgrade tables
!pip install eli5
!pip install xgboost
!pip install hyperopt
import pandas as pd
import numpy as np
import xgboost as xgb
from sklearn.metrics import mean_absolute_error as mea
from sklearn.model_selection import cross_val_score, KFold
from hyperopt import hp, fmin, tpe, STATUS_OK
import eli5
from eli5.sklearn import PermutationImportance
cd /content/drive/My Drive/Colab Notebooks/Matrix_repo/m_2/Car-Price-Prediction/data
df = pd.read_hdf('car.h5')
df.shape
```
Feature Engineering
```
SUFFIX_CAT = '__cat'
for feat in df.columns:
if isinstance(df[feat][0], list):continue
factorized_values = df[feat].factorize()[0]
if SUFFIX_CAT in feat:
df[feat] = factorized_values
else:
df[feat + SUFFIX_CAT] = factorized_values
df['param_moc'] = df['param_moc'].map(lambda x: -1 if str(x) == 'None' else int(str(x).split(' ')[0]) )
df['param_rok-produkcji']=df['param_rok-produkcji'].map(lambda x: -1 if str(x)== 'None' else int(x))
df['param_pojemność-skokowa']=df['param_pojemność-skokowa'].map(lambda x: -1 if str(x) == 'None' else int(str(x).split('cm')[0].replace(' ','')) )
def run_model(model,feats):
X= df[feats].values
y = df['price_value'].values
scores = cross_val_score(model, X,y, cv=3, scoring = 'neg_mean_absolute_error')
return np.mean(scores), np.std(scores)
```
XGBoost
```
feats = ["param_faktura-vat__cat",
"feature_kamera-cofania__cat",
"feature_łopatki-zmiany-biegów__cat",
"param_napęd__cat",
"param_skrzynia-biegów__cat",
"feature_asystent-pasa-ruchu__cat",
"param_stan__cat",
"feature_światła-led__cat",
"feature_bluetooth__cat",
"feature_regulowane-zawieszenie__cat",
"feature_wspomaganie-kierownicy__cat",
"feature_system-start-stop__cat",
"feature_światła-do-jazdy-dziennej__cat",
"feature_światła-xenonowe__cat",
"feature_czujniki-parkowania-przednie__cat",
"param_moc",
"param_rok-produkcji",
"param_pojemność-skokowa",
"feature_asystent-parkowania__cat",
"seller_name__cat"]
xgb_params={
'max_depth': 5,
'n_estimators': 50,
'learning_rate': 0.1,
'seed':0
}
run_model(xgb.XGBRegressor(**xgb_params),feats)
def obj_func(params):
print('training with params: ')
print(params)
mean_mea, score_std=run_model(xgb.XGBRegressor(**params), feats)
return {'loss': np.abs(mean_mea), 'status': STATUS_OK}
xgb_reg_params = {
'learning_rate': hp.choice('learning_rate', np.arange(0.05,0.31, 0.05)),
'max_depth': hp.choice('max_depth', np.arange(5, 16, 1, dtype=int)),
'subsample': hp.quniform('subsample', 0.5, 1, 0.05),
'colsample_bytree': hp.quniform('colsample_bytree', 0.5, 1, 0.05),
'objective': 'reg:squarederror',
'n_estimators': 100,
'seed': 0
}
best = fmin(obj_func, xgb_reg_params, algo=tpe.suggest, max_evals=25)
best
```
| github_jupyter |

# eodag-labextension: user manual
The `eodag-labextension` JupyterLab plugin brings satellite imagery search ability. With that extension you can search products, see results and check if results suits to your needs. And finally generate Python code allowing to access to those products in the notebook.
That tool is based on [eodag](https://github.com/CS-SI/eodag). See the [Python API User Guide](https://eodag.readthedocs.io/en/stable/api_user_guide.html) for a complete overview of eodag features.
## Configuration
For some providers we don’t need to set up credentials to search for products. But if you wish to download them, you should set the credentials beforehand, using these two environment variables for instance.
```
# Example for PEPS
os.environ["EODAG__PEPS__AUTH__CREDENTIALS__USERNAME"] = "PLEASE_CHANGE_ME"
os.environ["EODAG__PEPS__AUTH__CREDENTIALS__PASSWORD"] = "PLEASE_CHANGE_ME"
```
Credendials can be set dynamically using [environment variables](https://eodag.readthedocs.io/en/stable/getting_started_guide/configure.html#environment-variable-configuration) or also stored in the [configuration file](https://eodag.readthedocs.io/en/stable/getting_started_guide/configure.html#yaml-configuration-file) (recommanded). This file will be located at `$HOME/.config/eodag/eodag.yml`.
You can always choose to configure eodag using the configuration file or dynamically within your code, as shown in the [Configuration](https://eodag.readthedocs.io/en/stable/notebooks/api_user_guide/3_configuration.html) page of eodag's documentation.
Configuration also includes providers priority, download destination (`outputs_prefix`), extraction, ...
## Search
 Activate the plugin tab on the left of the JupuytuerLab interface by clicking on that icon.
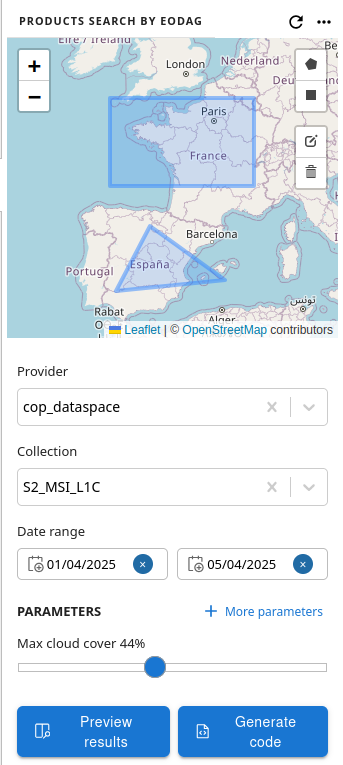
With displayed search form, you can enter search extent and following search criteria:
* **Product type**: the searched product type. For each entry of the drop-down list, a tooltip is displayed at hovering time with corresponding description.
* **Start date**: minimal date of the search temporal window.
* **End date**: maximal date of the search temporal window.
* **Max cloud cover**: maximum cloud cover allowed in search results in percent.
* **Additional parameters**: used to enter key-value pairs criteria for the request.
You can draw multiple extents, or use none. Each extent can be a rectangle or a free polygon.
Product type is mandatory. Other criteria are optional.
Once search criteria are filled out, click on the "Search" button to proceed to next step. At the end of the search, a popup opens and displays results.
## Results overview
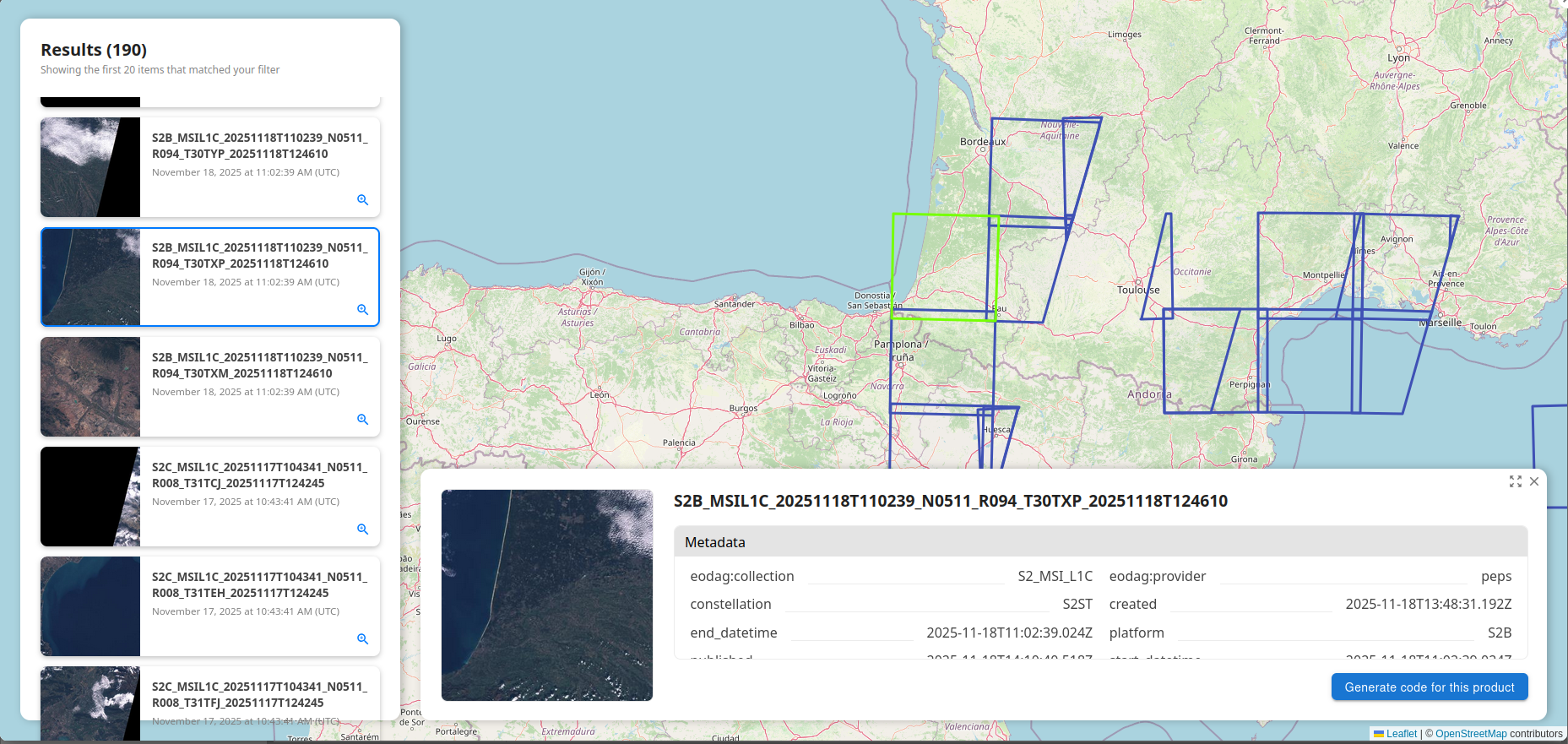
The results overview popup is compopsed of 3 parts:
* a map showing products extent,
* a table listing products,
* a pane containing metadata for currently selected product.
The results table allows you to access product metadata by clicking on the desired product line. The magnifying glass button allows you to zoom in on the product's in the map view. By scrolling down in the list of results, the search is automatically restarted to retrieve the following results.
In the metadata view, clicking on the thumbnail displays it in native resolution. Clicking it again reduces its size again.
## Apply to the Jupyter notebook
If the search result is correct, clicking on the "`Apply`" button will insert the Python eodag code in a new cell of the currently open notebook. The popup is automatically closed. From there, it is possible to work in the notebook on the search results by executing the eodag search.
Here is an example of generated code:
```
from eodag import EODataAccessGateway, setup_logging
setup_logging(1) # 0: nothing, 1: only progress bars, 2: INFO, 3: DEBUG
dag = EODataAccessGateway()
geometry = "POLYGON ((0.550136 43.005451, 0.550136 44.151469, 2.572104 44.151469, 2.572104 43.005451, 0.550136 43.005451))"
search_results, total_count = dag.search(
productType="S2_MSI_L1C",
geom=geometry,
start="2021-08-01",
end="2021-08-11",
cloudCover=17,
)
```
You may want to enforce usage of a particular provider. To do so, use [set_preferred_provider()](https://eodag.readthedocs.io/en/stablesearch/notebooks/api_user_guide/3_configuration.html?#Set-a-provider's-priority) in your search code after having instanciated [EODataAccessGateway](https://eodag.readthedocs.io/en/stable/api_reference/core.html#eodag.api.core.EODataAccessGateway):
```
dag = EODataAccessGateway()
dag.set_preferred_provider("theia")
```
## Using results
The obtained `SearchResult` will contain several `EOProduct` objects. See [SearchResult and EOProduct](https://eodag.readthedocs.io/en/stable/notebooks/api_user_guide/4_search.html#SearchResult-and-EOProduct) in the documentation for more information.
Here are some examples about how to use search results into a notebook:
```
from pprint import pprint
# Display results list
pprint(search_results)
# Display products access paths
pprint([p.location for p in search_results])
```
### Extract products extent
```
from shapely.geometry import GeometryCollection
features = GeometryCollection(
[product.geometry for product in search_results]
)
features
```
### Display products extent on a slippy map
```
from folium import Map, GeoJson, Figure
ext = features.bounds
bounds = [[ext[1], ext[0]], [ext[3], ext[2]]]
m = Map(tiles="Stamen Terrain", control_scale=True,)
GeoJson(search_results).add_to(m)
m.fit_bounds(bounds)
Figure(width=500, height=300).add_child(m)
```
## Downloading products
See [Download EO products](https://eodag.readthedocs.io/en/stable/notebooks/api_user_guide/7_download.html#Download-EO-products) in the documentation.
To download all products from the search request into a sub-directory called `downloaded`, run:
```
dag.download_all(search_results, outputs_prefix="downloaded")
```
| github_jupyter |
# Homework 7 - Berkeley STAT 157
**Your name: XX, SID YY, teammates A,B,C** (Please add your name, SID and teammates to ease Ryan and Rachel to grade.)
**Please submit your homework through [gradescope](http://gradescope.com/)**
Handout 4/2/2019, due 4/9/2019 by 4pm.
This homework deals with fine-tuning for computer vision. In this task, we attempt to identify 120 different breeds of dogs. The data set used in this competition is actually a subset of the ImageNet data set. Different from the images in the CIFAR-10 data set used in the previous homework, the images in the ImageNet data set are higher and wider and their dimensions are inconsistent. Again, you need to use GPU.
The dataset is available at [Kaggle](https://www.kaggle.com/c/dog-breed-identification). The rule is similar to homework 6:
- work as a team
- submit your results into Kaggle
- take a screen shot of your best score and insert it below
- the top 3 teams/individuals will be awarded with 500 dollar AWS credits
First, import the packages or modules required for the competition.
```
import collections
import d2l
import math
from mxnet import autograd, gluon, init, nd
from mxnet.gluon import model_zoo, nn
from mxnet.gluon import data as gdata, loss as gloss, utils as gutils
import os
import shutil
import time
import zipfile
```
## Obtain and Organize the Data Sets
The competition data is divided into a training set and testing set. The training set contains 10,222 images and the testing set contains 10,357 images. The images in both sets are in JPEG format. These images contain three RGB channels (color) and they have different heights and widths. There are 120 breeds of dogs in the training set, including Labradors, Poodles, Dachshunds, Samoyeds, Huskies, Chihuahuas, and Yorkshire Terriers.
### Download the Data Set
After logging in to Kaggle, we can click on the "Data" tab on the dog breed identification competition webpage shown in Figure 9.17 and download the training data set "train.zip", the testing data set "test.zip", and the training data set labels "label.csv.zip". After downloading the files, place them in the three paths below:
* kaggle_dog/train.zip
* kaggle_dog/test.zip
* kaggle_dog/labels.csv.zip
To make it easier to get started, we provide a small-scale sample of the data set mentioned above, "train_valid_test_tiny.zip". If you are going to use the full data set for the Kaggle competition, you will also need to change the `demo` variable below to `False`.
```
# If you use the full data set downloaded for the Kaggle competition,
# change the variable below to False.
demo = True
data_dir = './kaggle_dog'
if demo:
if not os.path.exists(data_dir):
os.mkdir(data_dir)
gutils.download('https://github.com/d2l-ai/d2l-en/raw/master/data/kaggle_dog/train_valid_test_tiny.zip',
data_dir)
zipfiles = ['train_valid_test_tiny.zip']
else:
zipfiles = ['train.zip', 'test.zip', 'labels.csv.zip']
for f in zipfiles:
with zipfile.ZipFile(data_dir + '/' + f, 'r') as z:
z.extractall(data_dir)
```
### Organize the Data Set
Next, we define the `reorg_train_valid` function to segment the validation set from the original Kaggle competition training set. The parameter `valid_ratio` in this function is the ratio of the number of examples of each dog breed in the validation set to the number of examples of the breed with the least examples (66) in the original training set. After organizing the data, images of the same breed will be placed in the same folder so that we can read them later.
```
def reorg_train_valid(data_dir, train_dir, input_dir, valid_ratio, idx_label):
# The number of examples of the least represented breed in the training set.
min_n_train_per_label = (
collections.Counter(idx_label.values()).most_common()[:-2:-1][0][1])
# The number of examples of each breed in the validation set.
n_valid_per_label = math.floor(min_n_train_per_label * valid_ratio)
label_count = {}
for train_file in os.listdir(os.path.join(data_dir, train_dir)):
idx = train_file.split('.')[0]
label = idx_label[idx]
d2l.mkdir_if_not_exist([data_dir, input_dir, 'train_valid', label])
shutil.copy(os.path.join(data_dir, train_dir, train_file),
os.path.join(data_dir, input_dir, 'train_valid', label))
if label not in label_count or label_count[label] < n_valid_per_label:
d2l.mkdir_if_not_exist([data_dir, input_dir, 'valid', label])
shutil.copy(os.path.join(data_dir, train_dir, train_file),
os.path.join(data_dir, input_dir, 'valid', label))
label_count[label] = label_count.get(label, 0) + 1
else:
d2l.mkdir_if_not_exist([data_dir, input_dir, 'train', label])
shutil.copy(os.path.join(data_dir, train_dir, train_file),
os.path.join(data_dir, input_dir, 'train', label))
```
The `reorg_dog_data` function below is used to read the training data labels, segment the validation set, and organize the training set.
```
def reorg_dog_data(data_dir, label_file, train_dir, test_dir, input_dir,
valid_ratio):
# Read the training data labels.
with open(os.path.join(data_dir, label_file), 'r') as f:
# Skip the file header line (column name).
lines = f.readlines()[1:]
tokens = [l.rstrip().split(',') for l in lines]
idx_label = dict(((idx, label) for idx, label in tokens))
reorg_train_valid(data_dir, train_dir, input_dir, valid_ratio, idx_label)
# Organize the training set.
d2l.mkdir_if_not_exist([data_dir, input_dir, 'test', 'unknown'])
for test_file in os.listdir(os.path.join(data_dir, test_dir)):
shutil.copy(os.path.join(data_dir, test_dir, test_file),
os.path.join(data_dir, input_dir, 'test', 'unknown'))
```
Because we are using a small data set, we set the batch size to 1. During actual training and testing, we would use the entire Kaggle Competition data set and call the `reorg_dog_data` function to organize the data set. Likewise, we would need to set the `batch_size` to a larger integer, such as 128.
```
if demo:
# Note: Here, we use a small data set and the batch size should be set
# smaller. When using the complete data set for the Kaggle competition,
# we can set the batch size to a larger integer.
input_dir, batch_size = 'train_valid_test_tiny', 1
else:
label_file, train_dir, test_dir = 'labels.csv', 'train', 'test'
input_dir, batch_size, valid_ratio = 'train_valid_test', 128, 0.1
reorg_dog_data(data_dir, label_file, train_dir, test_dir, input_dir,
valid_ratio)
```
## Image Augmentation
The size of the images in this section are larger than the images in the previous section. Here are some more image augmentation operations that might be useful.
```
transform_train = gdata.vision.transforms.Compose([
# Randomly crop the image to obtain an image with an area of 0.08 to 1
# of the original area and height to width ratio between 3/4 and 4/3.
# Then, scale the image to create a new image with a height and width
# of 224 pixels each.
gdata.vision.transforms.RandomResizedCrop(224, scale=(0.08, 1.0),
ratio=(3.0/4.0, 4.0/3.0)),
gdata.vision.transforms.RandomFlipLeftRight(),
# Randomly change the brightness, contrast, and saturation.
gdata.vision.transforms.RandomColorJitter(brightness=0.4, contrast=0.4,
saturation=0.4),
# Add random noise.
gdata.vision.transforms.RandomLighting(0.1),
gdata.vision.transforms.ToTensor(),
# Standardize each channel of the image.
gdata.vision.transforms.Normalize([0.485, 0.456, 0.406],
[0.229, 0.224, 0.225])])
```
During testing, we only use definite image preprocessing operations.
```
transform_test = gdata.vision.transforms.Compose([
gdata.vision.transforms.Resize(256),
# Crop a square of 224 by 224 from the center of the image.
gdata.vision.transforms.CenterCrop(224),
gdata.vision.transforms.ToTensor(),
gdata.vision.transforms.Normalize([0.485, 0.456, 0.406],
[0.229, 0.224, 0.225])])
```
## Read the Data Set
As in the previous section, we can create an `ImageFolderDataset` instance to read the data set containing the original image files.
```
train_ds = gdata.vision.ImageFolderDataset(
os.path.join(data_dir, input_dir, 'train'), flag=1)
valid_ds = gdata.vision.ImageFolderDataset(
os.path.join(data_dir, input_dir, 'valid'), flag=1)
train_valid_ds = gdata.vision.ImageFolderDataset(
os.path.join(data_dir, input_dir, 'train_valid'), flag=1)
test_ds = gdata.vision.ImageFolderDataset(
os.path.join(data_dir, input_dir, 'test'), flag=1)
```
Here, we create a `DataLoader` instance, just like in the previous section.
```
train_iter = gdata.DataLoader(train_ds.transform_first(transform_train),
batch_size, shuffle=True, last_batch='keep')
valid_iter = gdata.DataLoader(valid_ds.transform_first(transform_test),
batch_size, shuffle=True, last_batch='keep')
train_valid_iter = gdata.DataLoader(train_valid_ds.transform_first(
transform_train), batch_size, shuffle=True, last_batch='keep')
test_iter = gdata.DataLoader(test_ds.transform_first(transform_test),
batch_size, shuffle=False, last_batch='keep')
```
## Define the Model
The data set for this competition is a subset of the ImageNet data set. Therefore, we can use the approach discussed in the ["Fine Tuning"](fine-tuning.md) section to select a model pre-trained on the entire ImageNet data set and use it to extract image features to be input in the custom small-scale output network. Gluon provides a wide range of pre-trained models. Here, we will use the pre-trained ResNet-34 model. Because the competition data set is a subset of the pre-training data set, we simply reuse the input of the pre-trained model's output layer, i.e. the extracted features. Then, we can replace the original output layer with a small custom output network that can be trained, such as two fully connected layers in a series. Different from the experiment in the ["Fine Tuning"](fine-tuning.md) section, here, we do not retrain the pre-trained model used for feature extraction. This reduces the training time and the memory required to store model parameter gradients.
You must note that, during image augmentation, we use the mean values and standard deviations of the three RGB channels for the entire ImageNet data set for normalization. This is consistent with the normalization of the pre-trained model.
```
def get_net(ctx):
finetune_net = model_zoo.vision.resnet34_v2(pretrained=True)
# Define a new output network.
finetune_net.output_new = nn.HybridSequential(prefix='')
finetune_net.output_new.add(nn.Dense(256, activation='relu'))
# There are 120 output categories.
finetune_net.output_new.add(nn.Dense(120))
# Initialize the output network.
finetune_net.output_new.initialize(init.Xavier(), ctx=ctx)
# Distribute the model parameters to the CPUs or GPUs used for computation.
finetune_net.collect_params().reset_ctx(ctx)
return finetune_net
```
When calculating the loss, we first use the member variable `features` to obtain the input of the pre-trained model's output layer, i.e. the extracted feature. Then, we use this feature as the input for our small custom output network and compute the output.
```
loss = gloss.SoftmaxCrossEntropyLoss()
def evaluate_loss(data_iter, net, ctx):
l_sum, n = 0.0, 0
for X, y in data_iter:
y = y.as_in_context(ctx)
output_features = net.features(X.as_in_context(ctx))
outputs = net.output_new(output_features)
l_sum += loss(outputs, y).sum().asscalar()
n += y.size
return l_sum / n
```
## Define the Training Functions
We will select the model and tune hyper-parameters according to the model's performance on the validation set. The model training function `train` only trains the small custom output network.
```
def train(net, train_iter, valid_iter, num_epochs, lr, wd, ctx, lr_period,
lr_decay):
# Only train the small custom output network.
trainer = gluon.Trainer(net.output_new.collect_params(), 'sgd',
{'learning_rate': lr, 'momentum': 0.9, 'wd': wd})
for epoch in range(num_epochs):
train_l_sum, n, start = 0.0, 0, time.time()
if epoch > 0 and epoch % lr_period == 0:
trainer.set_learning_rate(trainer.learning_rate * lr_decay)
for X, y in train_iter:
y = y.as_in_context(ctx)
output_features = net.features(X.as_in_context(ctx))
with autograd.record():
outputs = net.output_new(output_features)
l = loss(outputs, y).sum()
l.backward()
trainer.step(batch_size)
train_l_sum += l.asscalar()
n += y.size
time_s = "time %.2f sec" % (time.time() - start)
if valid_iter is not None:
valid_loss = evaluate_loss(valid_iter, net, ctx)
epoch_s = ("epoch %d, train loss %f, valid loss %f, "
% (epoch + 1, train_l_sum / n, valid_loss))
else:
epoch_s = ("epoch %d, train loss %f, "
% (epoch + 1, train_l_sum / n))
print(epoch_s + time_s + ', lr ' + str(trainer.learning_rate))
```
## Train and Validate the Model
Now, we can train and validate the model. The following hyper-parameters can be tuned. For example, we can increase the number of epochs. Because `lr_period` and `lr_decay` are set to 10 and 0.1 respectively, the learning rate of the optimization algorithm will be multiplied by 0.1 after every 10 epochs.
```
ctx, num_epochs, lr, wd = d2l.try_gpu(), 1, 0.01, 1e-4
lr_period, lr_decay, net = 10, 0.1, get_net(ctx)
net.hybridize()
train(net, train_iter, valid_iter, num_epochs, lr, wd, ctx, lr_period,
lr_decay)
```
## Classify the Testing Set and Submit Results on Kaggle
After obtaining a satisfactory model design and hyper-parameters, we use all training data sets (including validation sets) to retrain the model and then classify the testing set. Note that predictions are made by the output network we just trained.
```
net = get_net(ctx)
net.hybridize()
train(net, train_valid_iter, None, num_epochs, lr, wd, ctx, lr_period,
lr_decay)
preds = []
for data, label in test_iter:
output_features = net.features(data.as_in_context(ctx))
output = nd.softmax(net.output_new(output_features))
preds.extend(output.asnumpy())
ids = sorted(os.listdir(os.path.join(data_dir, input_dir, 'test/unknown')))
with open('submission.csv', 'w') as f:
f.write('id,' + ','.join(train_valid_ds.synsets) + '\n')
for i, output in zip(ids, preds):
f.write(i.split('.')[0] + ',' + ','.join(
[str(num) for num in output]) + '\n')
```
After executing the above code, we will generate a "submission.csv" file. The format of this file is consistent with the Kaggle competition requirements.
## Hints to Improve Your Results
* You should download the whole data set from Kaggle and switch to `demo=False`.
* Try to increase the `batch_size` (batch size) and `num_epochs` (number of epochs).
* Try a deeper pre-trained model, you may find models from [gluoncv](https://gluon-cv.mxnet.io/model_zoo/classification.html).
| github_jupyter |
### Case Study 4 : Data Science Shark Tank: Pitch Your Ideas
** Due Date: April 27, 6pm**
<img src="https://cp.inkrefuge.com/images%5Cpressreleases/shark%20tank_large.jpg" width="400px">
After the previous 3 case studies, your team has now equipped with all the three powerful skills of data science: Hacking skill, Business skill and Math skill. In this project, your team is going to make use of these skills to come up with an idea of a new business/startup based upon data science technology. Your goal is to design a better service/solution on any data you like, develop a prototype/demo and prepare a pitch for your idea.
* Your team needs to decide which business problem is important for the market you are joining in (for example, social media market, housing market, search market, etc.).
* Then design a data science approach to improve one of the current services or design a new service on any data that you choose.
* The solution should include all the three components of data science: 1) the business part to analyze the potential impact of your new/improved service, why the idea can make money, how much are you evaluating the company; How are you planing to persuade the sharks to invest in your business; 2) the mathematical part to formulate the problem and develop math solution; 3) the programming part to collect the data, implement the math solution, and develop the prototype/demo.
# Background about Elevator Pitch (90 seconds) and Shark Tank
```
from IPython.display import YouTubeVideo
YouTubeVideo("mrSmaCo29U4")
YouTubeVideo("xIq8Sg59UdY")
```
### Two videos on storytelling
https://video.wpi.edu/Watch/g2T4NjBn
https://video.wpi.edu/Watch/q2A6Dbg3
**Optional Readings:**
* LinkedIn API: https://developer.linkedin.com/docs/rest-api
* Zillow API: https://pypi.python.org/pypi/pyzillow
* Google Map API: https://developers.google.com/api-client-library/python/apis/mapsengine/v1?hl=en
* More APIs: https://github.com/ptwobrussell/Mining-the-Social-Web-2nd-Edition
** Python libraries you may want to use:**
* Scikit-learn (http://scikit-learn.org): machine learning tools in Python.
** Data sources:**
* UCI Machine Learning Repository: http://archive.ics.uci.edu/ml/
* Statlib datasets: http://lib.stat.cmu.edu/
* Kaggel: www.kaggle.com
* Open Gov. Data: www.data.gov, www.data.gov.uk, www.data.gov.fr, http://opengovernmentdata.org/data/catalogues/
** NOTE **
* Please don't forget to save the notebook frequently when working in IPython Notebook, otherwise the changes you made can be lost.
*----------------------
### Problem 1: the Business Part (20 points)
As a group, learn about the data science related business and research about the current markets: such as search, social media, advertisement, recommendation and so on.
Pick one of the markets for further consideration, and design a new service which you believe to be important in the market.
Define precisely in the report and briefly in the cells below, what is the business problem that your team wants to solve.
Why the problem is important to solve?
Why you believe you could make a big difference with data science technology.
How are you planing to persuade the investors to buy in your idea.
**Please describe here *briefly* (please edit this cell)**
1) Your business problem to solve:
Understand when an article, published online, is likely to be "popular" in comparison to other articles released in similar formats. More precisely, the goal is to predict--before publication--whether an article is likely to be unpopular.
2) Why the problem is important to solve?
Text-based content (articles, news, blogs, etc.) is now largely consumed via online platforms. Without hard-copy sales numbers (as in the case of books and newspapers) to judge performance, those producing this content still need to understand the popularity of the articles they publish. A method of understanding likely popularity, before publication, can be extremely useful in deciding which content to release, as well as in attracting potential advertisers.
3) What is your idea to solve the problem?
Social media platforms have become the primary outlet for many to share their opinions. Thus while there are many measures to judge an articles "popularity" (such as page-visits, or external links) we choose to use the number of shares via social media as our metric of "popularity."
Moreover, while the specific words in an article influence the reader's enjoyment, the structure of the article itself also plays a role in its popularity. For example, perhaps longer articles are not read when published on a Monday. Technology articles with very short titles may not generate interest. Rather than considering the individual tokens within the text, we will used structure-based analysis.
Lastly, we will use ensemble-learning to make final predictions. While this may sacrifice some accuracy, it will help to decrease false negatives (the costly real-world mistake in this problem).
4) What differences you could make with your data science approach?
The use of structure-based article statistics offer a few advantages. First, this data can be computed and compared for all articles, regardless of the number available. This is especially useful if the number of training articles is small: there may be little (or no relevant) overlap in the individual text tokens. Second, choosing these predictors keeps our data low-dimensional. The number of token-based predictors for a collection of texts grows very quickly, resulting in very high-dimensional data. Fixing a number of structure-based predictors bounds the dimension of our data, giving us a better chance for accurate prediction. Lastly, these structure-based statistics offer interpretability: this could lead to suggestion features in our software (such as "increasing the title length will improve popularity" or "use shorter words in the first and last sentence to improve popularity").
5) Why do you believe the idea deserves the investment of the "sharks"?
Studies indicate that more and more people rely on online outlets to receive news and information. The amount of content generated online continues to grow as well. Thus the ability to understand, before release, an article's likelihood of success will become increasingly beneficial to online contributors. Utilizing structure-based analytics appeals to all contributors of online content. Large-scale publications may use our services with confidence that their pre-release content cannot be leaked. Large amounts of online content is generated by individuals. Professional bloggers, freelance writers, and "contributing authors," depend on article popularity, and social media buzz, to maintain their careers and attract potential advertisers. Most such individuals will lack the tools (and skills) to perform data analysis, and thus can benefit from our services.
Perhaps most important to potential investors, we are the first pre-release publication service offered in the market: news Media generated $63 Billion in US revenue last year.
---
### Problem 2: The Math Part (20 points)
Define the business problem as a math problem and design a math solution to the problem.
### 1) Problem formulation in Math
As described in the previous part, our goal is to predict the "popularity" of an online news article, based on structural features of the text. To begin, we must determine an appropriate metric of "popularity." Social media platforms are a primary outlet in which we express our opinions: we often "share" an online article that we believe others should read. Thus "shares" on social media can be used to estimate popularity. We will consider two metrics of popularity based on "shares":
#### Popularity
The raw number of shares an article receives. This metric is only relevant given a base-line number of shares for other articles posted in similar formats.
#### Buzz-Factor
While the total number of shares an article receives can be used to estimate popularity, the speed at which those shares are generated is also of interest. Suppose articles A and B each receive 2,000 shares. If article A generated those shares over 6 months, while article B gained the same shares in only 2 days, article B can be judged as more "popular." The number of shares received per day, an estimate of the share-rate, will be referred to as buzz-factor.
We focus our analysis even further. It would certainly be useful to be able to estimate the *extent* of an articles popularity, of the utmost interest of content producers is being able to anticipate when an article is *unpopular*. That is, while it would be convenient to predict whether an article receives 1,500 as opposed to 1,200 shares, it is more important to authors/publishers to predict when an article will receive *very low* numbers of shares. In this case, it is worthwhile to consider changing the format of the article, or perhaps not releasing the content at all. Therefore, our aim is to predict when an article will be *unpopular* or generate *no buzz*. Once again we must define these terms mathematically.
#### Unpopular
Our metric of popularity is number of shares. We consider an article to be *unpopular* if, among other articles of similar formats, it generates a number of shares which is in the bottom 25%. That is, given a collection of articles, the unpopular articles are those in percentiles 0-25 in number of shares.
#### No-Buzz
Our metric of popularity is number of shares per day. We consider an article to have *no buzz* if, among other articles of similar formats, it generates a buzz which is in the bottom 25%. That is, given a collection of articles, the articles with no-buzz are those in percentiles 0-25 in number of shares.
Therefore, our business problem can be phrased mathematically as follows:
#### *Given a collection of online news arctiles (of similar formats or origin) predict, which among those, will rank in the bottom 25% in numbers of shares or number of shares per day.*
### 2) Math Solution
We make a deliberate choice of how to transform articles into data for analysis. Specifically, we choose to compute attributes based on the structure of the article, as opposed to the textual content. This offers a few advantages:
* If the sample size of comparable articles is small, there may be little (or at least little relevant) overlap in the tokens contained within the text.
* Attributes based on the structure of an article may be computed for any text, and are comparable in a meaningful way regardless of the number of similar documents available.
* Software can be sent to customers which inputs an article, and outputs a set of structure-based attributes. This data can then be relayed to our services for analyis. This enables us to perform data analysis without needing access to the original text, thereby protecting the unpublished content of our customers.
#### Attributes
The following attributes are computed for training (comparable) articles:
* Number of "Shares" on social media ('shares')
* Number of days since publication ('timedelta')
The following attributes are computed for training and testing articles:
* Number of words in the Title ('n_tokens_title')
* Number of words in the text body ('n_tokens_content',)
* Number of unique words in the text body ('n_unique_tokens')
* Number of unique non-stop words in the text body ('n_non_stop_unique_tokens')
* Number of videos included in the article ('num_videos')
* Number of images included in the article ('num_imgs')
* Average word length ('average_token_length')
* Number of keywords associated with the article ('num_keywords')
* Genre of the article (As 6 dummy variables)
* 'is_lifestyle','is_entertainment','is_business','is_social_media','is_tech','is_world'
* Day of the week of publication (As 8 dummy variables)
* 'is_monday','is_tuesday','is_wednesday','is_thursday','is_friday','is_saturday','is_sunday'
* 'is_weekend'
* Text subjectivity ('global_subjectivity')
* Overal text polarity ('global_sentiment_polarity')
* Rate of positive words in the content ('global_rate_positive_words')
* Rate of negative words in the content ('global_rate_negative_words')
* Rate of positive words among non-neutral ('tokens rate_positive_words')
* Rate of negative words among non-neutral tokens ('rate_negative_words')
* Avg. polarity of positive words ('avg_positive_polarity')
* Min. polarity of positive words ('min_positive_polarity')
* Max. polarity of positive words ('max_positive_polarity')
* Avg. polarity of negative words ('avg_negative_polarity')
* Min. polarity of negative words ('min_negative_polarity ')
* Max. polarity of negative words ('max_negative_polarity')
* Title Subjectivity ('title_subjectivity')
* Title Polarity ('title_sentiment_polarity')
For each training article, we split the articles into four "popularity" bins based on percentile of 'shares'
* 0-25%-- "Unpopular"
* 25%-50%-- "Mildly Popular"
* 50%-75%--"Popular"
* 75%-100%-- "Very Popular"
We store the popularity classification as an additional attribute ('popularity')
For each training article, we compute a 'buzz_factor' attribute,
* buzz_factor = 'shares'/'timedelta'
For each training article we divide the articles into four "buzz" bins based on percentile of 'buzz factor'
* 0-25% "No Buzz"
* 25%-50% "Some Buzz"
* 50%-75% "Buzz"
* 75%-100% "Lots of Buzz"
We store the "buzz" classification as an additional attribute, ('buzz')
#### Target Variables
Our goal is to predict whether or not an article will be "unpopular" or generate "no buzz." Therefore we generate two boolean variables,
* 'unpopular' (Takes value 'True' if 'popularity' = "unpopular")
* 'no_buzz' (Takes vale 'True' if 'buzz' = "no_buzz")
#### Feature Selection
After all attribute constructions (including a number of dummy variables for classification), our data has approximately 50 predictors. In order to aid in interpretability of results, we reduce the number of features considered to 10. We perform this process twice, by using a feature-importance metric to determine those which are most relevant to the target variables ('unpopular', and 'no_buzz'). This is done using the ExtraTreesClassifier on a set of training documents. The ExtraTreesClassifier is a "class [that] implements a meta estimator that fits a number of randomized decision trees (a.k.a. extra-trees) on various sub-samples of the dataset and use averaging to improve the predictive accuracy and control over-fitting" [Per sklearn documentation]. This class includes a feature_importance attribute, from which we extract the top 10 features.
#### Machine Learning
We now train machine-learning algorithms on the set of (features computed for the ) training documents. For each target variable, these models only consider the top 10 features determined in the previous step. Our method considers two machine learning algorithms:
* Random Forest Classifier
* KNN (k chosen by cross-validation)
#### Prediction
We use ensemble learning to make predictions. Given a test article, the machine learning methods above are trained on a set of articles of a similar format or origin. In order to predict whether the test article will be "unpopular" or generate "no_buzz" each of the three algorithms make a prediction. We predict that an article is "unpopular" if EITHER of the trained models predict that it is unpopular. This method likely does not produce the highest accuracy, HOWEVER, it acts to minimize false negatives. That is, by predicting an article is "unpopular" if either of the models predicts "unpopular", it serves to DECREASE the number of arcticles which are unpopular that we (incorrectly) predict as "popular." Predicting an unpopular article as "popular" is the more expensive error within real world settings: this method acts to decrease such mistakes.
#### 3) Implementation of the Solution
```
# NOTE: THIS CODE IS NOT TO BE RUN, AS IT HAS NO DATA! IT IS THE SHELL FOR THE CODE, WHICH WILL BE RUN IN PROBLEM 3.
# GATHERING DATA FROM A WEB ARTICLE & EXTRACTING FEATURES
# import re
# import nltk
# import pprint
# from urllib import urlopen
# from bs4 import BeautifulSoup
# from collections import Counter
# # Input your own text
# raw = raw_input("Enter some text: ")
# # Online articles
# url = "http://www.foxnews.com/science/2016/04/22/nasa-marks-hubbles-birthday-with-this-captivating-image.html"
# html = urlopen(url).read()
# raw2 = BeautifulSoup(html).get_text()
# # Add to temporary local file
# f = open('text.txt', 'r')
# raw3 = f.read()
# f.close()
# # Filter stopwords
# stopwords = nltk.corpus.stopwords.words('english')
# text= [w for w in raw3 if w.lower() not in stopwords]
# # Number of words in title
# title=[t for t in raw3 if t.istitle()]
# wordcounttitle = Counter(title.split( ))
# n_tokens_title=len(wordcounttitle)
# # Number of words in the text body
# wordcount = Counter(raw3.split( ))
# n_tokens_content=len(wordcount)
# n_tokens_content
# # Number of unique words in text body
# n_unique_tokens=len(set(raw3))
# n_unique_tokens
# #number of unique non-stop words in text body
# n_non_stop_unique_tokens=len(set(text))
# n_non_stop_unique_tokens
# #average word length for original text
# average_token_length=len(text)/n_tokens_content
# average_token_length
# # Text subjectivity
# from textblob import TextBlob
# Global_subjectivity=TextBlob(text).sentiment.subjectivity
# # Overall text poluarity
# Global_sentiment_polarity=TextBlob(text).sentiment.polarity
#FEATURE IMPORTANCE
#INPUTS:
# df: data frame containing the (structure) features for the training data.
# feautures: list of (non-response) features
# target: target variable
#feature_selection_model = ExtraTreesClassifier().fit(df[features], df['unpopular'])
#feature_importance=feature_selection_model.feature_importances_
#importance_matrix=np.array([features,list(feature_importance)]).T
#def sortkey(s):
# return s[1]
#sort=zip(features,list(feature_importance))
#f = pd.DataFrame(sorted(sort,key=sortkey,reverse=True),columns=['variables','importance'])[:10]
# EXTRACT TOP FEATURES, DETERMINE TRAINING DOCUMENTS
#features2=f['variables']
#split data into two parts
#np.random.seed(0)
#x_train, x_test, y_train, y_test = train_test_split(df[features2], df.unpopular, test_size=0.4, random_state=None)
#x_train.shape
# ENSEMBLE LEARNING
# Random Forest
#print "RandomForest"
#rf = RandomForestClassifier(n_estimators=100,n_jobs=1)
#clf_rf = rf.fit(x_train,y_train)
#y_predicted_rf = clf_rf.predict(x_test)
# K-NN:
# Determine K by cross-validation.
#x_cv_train, x_cv_test, y_cv_train, y_cv_test = train_test_split(x_train, y_train, test_size=0.3, random_state=None)
# We use K-values ranging from 1-10
# k=[5,10,15,20,25,30,35,40,45,50]
# Train a model on the trainng set and use that model to predict on the testing set
# predicted_knn=[KNeighborsClassifier(n_neighbors=i).fit(x_cv_train,y_cv_train).predict(x_cv_test) for i in k]
#Compute accuracy on the testing set for each value of k
#score_knn=[metrics.accuracy_score(predicted_knn[i],y_cv_test) for i in range(10)]
# Plot accuracy on the test set vs. k
#fig=plt.figure(figsize=(8,6))
#plt.plot([5,10,15,20,25,30,35,40,45,50], score_knn, 'bo--',label='knn')
#plt.xlabel('K')
#plt.ylabel('score')
# Make predictions based on the best model above
# y_predicted_knn = KNeighborsClassifier(n_neighbors=6).fit(x_train,y_train).predict(x_test)
####FINAL PREDICTIONS: ENSEMBLE LEARNING
#y_predicted = y_predicted_knn + y_predicted_rf
#cm = metrics.confusion_matrix(y_test, y_predicted)
#print(cm)
#plt.matshow(cm)
#plt.title('Confusion matrix')
#plt.colorbar()
#plt.ylabel('True label')
#plt.xlabel('Predicted label')
#plt.show()
#print 'Prediction Accuracy'
#print (cm[0,0]+cm[1,1])/float(cm[0,0] + cm[0,1] + cm[1,0] + cm[1,1])
```
### Problem 3: The Hacking Part (20 points)
* Data Collection
* Implement a small Demo/Prototype/experiment result figures for the "product" of your data science company. You could use this demo during the Pitch
### Global Imports & Variables
```
%matplotlib inline
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
csv_path = 'data/OnlineNewsPopularity.csv'
hdf_path = 'data/online_news_popularity.h5'
blue = '#5898f1'
green = '#00b27f'
yellow = '#FEC04C'
red = '#fa5744'
```
### Download data, unzip, and save .csv
```
import requests, StringIO, csv, zipfile, sys
url = 'http://archive.ics.uci.edu/ml/machine-learning-databases/00332/OnlineNewsPopularity.zip'
request = requests.get(url)
print "Downloading data ...\nRequest status: {}".format(request.status_code)
archive = ZipFile(StringIO.StringIO(request.content))
print "Unzipping ..."
csv_data = archive.read('OnlineNewsPopularity/OnlineNewsPopularity.csv', 'r')
outfile = file(csv_path, 'w')
outfile.write(csv_data)
print "Saving into {}".format(csv_path)
outfile.close()
```
### Read .csv file into Pandas Data Frame & save into HDF5 (Hierarchical Data Format)
```
# Read .csv file into data frame
data_frame = pd.read_csv(csv_path, sep=', ', engine='python')
# Rename *channel* columns
data_frame.rename(columns={
'data_channel_is_lifestyle': 'is_lifestyle',
'data_channel_is_entertainment': 'is_entertainment',
'data_channel_is_bus': 'is_business',
'data_channel_is_socmed': 'is_social_media',
'data_channel_is_tech': 'is_tech',
'data_channel_is_world': 'is_world',
}, inplace=True)
# Rename *weekday* columns
data_frame.rename(columns={
'weekday_is_monday': 'is_monday',
'weekday_is_tuesday': 'is_tuesday',
'weekday_is_wednesday': 'is_wednesday',
'weekday_is_thursday': 'is_thursday',
'weekday_is_friday': 'is_friday',
'weekday_is_saturday': 'is_saturday',
'weekday_is_sunday': 'is_sunday',
}, inplace=True)
# Store data into HDF5 file
data_hdf = pd.HDFStore(hdf_path)
data_hdf['data_frame'] = data_frame
data_hdf.close()
```
### Read HDF5 file into Pandas Data Frame
```
# Read .h5 file into data frame
data_frame = pd.read_hdf(hdf_path)
# Drop columns included in the sample Dataset that are not considered in our methods.
data_frame.drop('LDA_00', axis=1, inplace=True)
data_frame.drop('LDA_01', axis=1, inplace=True)
data_frame.drop('LDA_02', axis=1, inplace=True)
data_frame.drop('LDA_03', axis=1, inplace=True)
data_frame.drop('LDA_04', axis=1, inplace=True)
data_frame.drop('kw_min_min', axis=1, inplace=True)
data_frame.drop('kw_max_min', axis=1, inplace=True)
data_frame.drop('kw_avg_min', axis=1, inplace=True)
data_frame.drop('kw_min_max', axis=1, inplace=True)
data_frame.drop('kw_max_max', axis=1, inplace=True)
data_frame.drop('kw_avg_max', axis=1, inplace=True)
data_frame.drop('kw_min_avg', axis=1, inplace=True)
data_frame.drop('kw_max_avg', axis=1, inplace=True)
data_frame.drop('kw_avg_avg', axis=1, inplace=True)
data_frame.drop('n_non_stop_words', axis=1, inplace=True)
data_frame.drop('url', axis=1, inplace=True)
# Data frame column headers
list(data_frame)
```
#### Remove outliers if applicable
```
# keep points that are within +2 to -2 standard deviations in column 'Data'.
df = data_frame[data_frame.shares-data_frame.shares.mean()<=(2*data_frame.shares.std())]
df.shape
```
Added 'buzz_factor' column: "Buzz-Factor" (using shares/day)
```
buzz_factor = df['shares'] / df['timedelta']
```
Added 'popularity' column: split # of shares into 4 "popularity" bins:
* 0-25%-- "Unpopular"
* 25%-50%-- "Mildly Popular"
* 50%-75%--"Popular"
* 75%-100%-- "Very Popular"
```
popularity = pd.qcut(df['shares'], 4, labels=[
"Unpopular",
"Midly Popular",
"Popular",
"Very Popular"
])
# Add these two statistics to the Data Frame
df.is_copy = False # turn off chain index warning
df['buzz_factor'] = buzz_factor.values
df['popularity'] = popularity.values
```
Similarly, split *buzz factor* into four percentile bins:
* 0-25% "No Buzz"
* 25%-50% "Some Buzz"
* 50%-75% "Buzz"
* 75%-100% "Lots of Buzz"
```
buzz = pd.qcut(df['buzz_factor'], 4, labels=["No Buzz","Some Buzz","Buzz","Lots of Buzz"])
df['buzz']=buzz.values
```
The real quantity of interest here is, in some sense, the LEAST successful articles. While it is interesting to predict the level of popularity/buzz factor, what we need (at the very least) to be able to do is predict whether an article will be "unpopular" or generate "no buzz." Thus we isolate these two bins.
```
# Compute Target variables and add to data frame
unpopular = df['popularity']== 'Unpopular'
df['unpopular'] = unpopular
no_buzz = df['buzz']=='No Buzz'
df['no_buzz'] = no_buzz
df.shape
# Brief Exploration of the "Popularity" classes: Consider mean number of shares per "popularity" bin
df_popularity = df.pivot_table('shares', index='popularity', aggfunc='mean')
df_popularity_count = df.pivot_table('shares', index='popularity', aggfunc='count')
print df_popularity_count
df_popularity.plot(kind='bar', color=green)
plt.title('Article Popularity')
plt.ylabel('mean shares')
# Brief Exploration of the "buzz" classes: Consider mean number of shares per day for each "buzz" bin
df_buzz = df.pivot_table('buzz_factor', index='buzz', aggfunc='mean')
df_buzz_count = df.pivot_table('buzz_factor', index='buzz', aggfunc='count')
print df_buzz_count
df_buzz.plot(kind='bar', color=green)
plt.title('Article Buzz')
plt.ylabel('buzz factor')
# Isolate non-response features
all_features = df.columns.values
excluded_features = [
'buzz',
'buzz_factor',
'no_buzz',
'popularity',
'shares',
'unpopular'
]
features = [f for f in all_features if f not in excluded_features]
```
#### Import Sklearn packages.
```
from sklearn import preprocessing
from sklearn.cross_validation import train_test_split
from sklearn.grid_search import GridSearchCV
from sklearn.metrics import confusion_matrix
from sklearn.decomposition import PCA
from sklearn.svm import SVC
import pandas as pd
from sklearn import metrics
from sklearn.metrics import fbeta_score, make_scorer
from sklearn.neighbors import KNeighborsClassifier
from sklearn.neighbors import RadiusNeighborsClassifier
from sklearn.ensemble import ExtraTreesClassifier
from sklearn.lda import LDA
from sklearn.qda import QDA
import numpy as np
from sklearn.ensemble import RandomForestClassifier
from sklearn.neighbors import NearestNeighbors
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import BaggingClassifier
from sklearn.svm import LinearSVC
from sklearn import cross_validation, metrics
from sklearn.naive_bayes import BernoulliNB
from time import time
```
# Unpopular Articles
At the very least, we want our software to predict whether an article will be unpopular. In this case, we define Unpopular as belonging in the bottom 25% in terms of shares amongst articles on a similar platform.
```
# Rank features by importance
feature_selection_model = ExtraTreesClassifier().fit(df[features], df['unpopular'])
feature_importance=feature_selection_model.feature_importances_
importance_matrix=np.array([features,list(feature_importance)]).T
def sortkey(s):
return s[1]
sort=zip(features,list(feature_importance))
# Extract top 10 important features
f = pd.DataFrame(sorted(sort,key=sortkey,reverse=True),columns=['variables','importance'])[:10]
f
features2=f['variables']
#split data into two parts
np.random.seed(0)
x_train, x_test, y_train, y_test = train_test_split(df[features2], df.unpopular, test_size=0.4, random_state=None)
x_train.shape
# Decision Tree accuracy and time elapsed caculation
#t0=time()
#print "DecisionTree"
#dt = DecisionTreeClassifier(min_samples_split=25,random_state=1)
#clf_dt=dt.fit(x_train,y_train)
#y_predicted_dt = clf_dt.predict(x_test)
#t1=time()
# Observe how decision tree performed on its own.
#print(metrics.classification_report(y_test, y_predicted_dt))
#print "time elapsed: ", t1-t0
# Random Forest
# Trainc classifier and time elapsed caculation
t2=time()
print "RandomForest"
rf = RandomForestClassifier(n_estimators=100,n_jobs=1)
clf_rf = rf.fit(x_train,y_train)
y_predicted_rf = clf_rf.predict(x_test)
t3=time()
# See how Random forest performed on its own.
print "Acurracy: ", clf_rf.score(x_test,y_test)
print "time elapsed: ", t3-t2
#KNN
from sklearn.neighbors import KNeighborsClassifier
# Determine K by cross-validation.
x_cv_train, x_cv_test, y_cv_train, y_cv_test = train_test_split(x_train, y_train, test_size=0.3, random_state=None)
# We use K-values ranging from 1-10
k=[5,10,15,20,25,30,35,40,45,50]
# Train a model on the training set and use that model to predict on the testing set
predicted_knn=[KNeighborsClassifier(n_neighbors=i).fit(x_cv_train,y_cv_train).predict(x_cv_test) for i in k]
#Compute accuracy on the testing set for each value of k
score_knn=[metrics.accuracy_score(predicted_knn[i],y_cv_test) for i in range(10)]
print score_knn
# Plot accuracy on the test set vs. k
fig=plt.figure(figsize=(8,6))
plt.plot([5,10,15,20,25,30,35,40,45,50], score_knn, 'bo--',label='knn', color=green)
plt.title('Unpopular K-NN')
plt.xlabel('K')
plt.ylabel('score')
# Make predictions based on best model above
y_predicted_knn = KNeighborsClassifier(n_neighbors=6).fit(x_train,y_train).predict(x_test)
# See how KNN did on its own.
# Print and plot a confusion matrix
cm = metrics.confusion_matrix(y_test, y_predicted_knn)
print(cm)
plt.matshow(cm)
plt.title('Confusion matrix')
plt.colorbar()
plt.ylabel('True label')
plt.xlabel('Predicted label')
plt.show()
```
### Final Predictions: Ensemble Learning
```
# Predict unpopular = True if either RF or KNN predict unpopular = True
y_predicted = y_predicted_knn + y_predicted_rf
#Print and plot confusion matrix
cm = metrics.confusion_matrix(y_test, y_predicted)
print(cm)
plt.matshow(cm)
plt.title('Confusion matrix')
plt.colorbar()
plt.ylabel('True label')
plt.xlabel('Predicted label')
plt.show()
# Print prediction Accuracy
print 'Prediction Accuracy'
print (cm[0,0]+cm[1,1])/float(cm[0,0] + cm[0,1] + cm[1,0] + cm[1,1])
```
# Articles with no Buzz
At the very least, we want our software to predict whether an article will generate no "buzz". In this case, we define "no buzz" as belonging in the bottom 25% in terms of shares per day amongst articles published on a similar platform.
```
# Select features
all_features = df.columns.values
excluded_features = [
'buzz',
'buzz_factor',
'no_buzz',
'popularity',
'shares',
'timedelta',
'unpopular'
]
features1 = [f for f in all_features if f not in excluded_features]
#Rank features by importance
feature_selection_model = ExtraTreesClassifier().fit(df[features1], df['no_buzz'])
feature_importance=feature_selection_model.feature_importances_
importance_matrix=np.array([features, list(feature_importance)]).T
def sortkey(s):
return s[1]
sort=zip(features,list(feature_importance))
# Extract top 10 important features
f_b=pd.DataFrame(sorted(sort,key=sortkey,reverse=True),columns=['variables','importance'])[:10]
f_b
features_b=f_b['variables']
#split data into two parts
np.random.seed(0)
x_train, x_test, y_train, y_test = train_test_split(df[features_b], df.no_buzz, test_size=0.4, random_state=None)
x_train.shape
# Decision Tree accuracy and time elapsed caculation
#t0=time()
#print "DecisionTree"
#dt = DecisionTreeClassifier(min_samples_split=25,random_state=1)
#clf_dt=dt.fit(x_train,y_train)
#y_predicted = clf_dt.predict(x_test)
#print(metrics.classification_report(y_test, y_predicted))
#t1=time()
#print "time elapsed: ", t1-t0
#Random Forest
# Train classifier and time elapsed caculation
t2=time()
print "RandomForest"
rf = RandomForestClassifier(n_estimators=100,n_jobs=1)
clf_rf = rf.fit(x_train,y_train)
y_predicted_rf = clf_rf.predict(x_test)
# See how random forest did on its own.
print "Acurracy: ", clf_rf.score(x_test,y_test)
t3=time()
print "time elapsed: ", t3-t2
# KNN
from sklearn.neighbors import KNeighborsClassifier
# Determine K by cross-validation.
x_cv_train, x_cv_test, y_cv_train, y_cv_test = train_test_split(x_train, y_train, test_size=0.3, random_state=None)
# We use K-values ranging from 1-10
k=[5,10,15,20,25,30,35,40,45,50]
# Train a model on the trainng set and use that model to predict on the testing set
predicted_knn=[KNeighborsClassifier(n_neighbors=i).fit(x_cv_train,y_cv_train).predict(x_cv_test) for i in k]
#Compute accuracy on the testing set for each value of k
score_knn=[metrics.accuracy_score(predicted_knn[i],y_cv_test) for i in range(10)]
print score_knn
# Plot accuracy on the test set vs. k
fig=plt.figure(figsize=(8,6))
plt.plot([5,10,15,20,25,30,35,40,45,50], score_knn, 'bo--',label='knn', color=green)
plt.title('No Buzz K-NN')
plt.xlabel('K')
plt.ylabel('score')
# Make predictions based on the best model above
y_predicted_knn = KNeighborsClassifier(n_neighbors=7).fit(x_train,y_train).predict(x_test)
# See how KNN did on its own.
# Print and plot a confusion matrix
cm = metrics.confusion_matrix(y_test, y_predicted_knn)
print(cm)
plt.matshow(cm)
plt.title('Confusion matrix')
plt.colorbar()
plt.ylabel('True label')
plt.xlabel('Predicted label')
plt.show()
```
### Final Predictions: Ensemble Learning
```
# Predict no_buzz = True if either RF or KNN predict no_buzz = True
y_predicted = y_predicted_knn + y_predicted_rf
# Print and plot confusion matrix
cm = metrics.confusion_matrix(y_test, y_predicted)
print(cm)
plt.matshow(cm)
plt.title('Confusion matrix')
plt.colorbar()
plt.ylabel('True label')
plt.xlabel('Predicted label')
plt.show()
# Print prediction accuracy
print 'Prediction Accuracy'
print (cm[0,0]+cm[1,1])/float(cm[0,0] + cm[0,1] + cm[1,0] + cm[1,1])
```
### Background analysis/exploration.
### IMPORTANT NOTE: The code cells above this one are the complete "solution" cells for problem 3. What follows was additional exploration/analysis. It was not used in our results, but we retained it in this notebook for completeness.
Here we try to use PCA to improve our results.
```
from sklearn.decomposition import PCA
pca = PCA(4)
plot_columns = pca.fit_transform(df[features1])
plt.scatter(x=plot_columns[:,0], y=plot_columns[:,1], c=df['unpopular'], color=[red, blue])
plt.show()
np.random.seed(0)
x_train, x_test, y_train, y_test = train_test_split(plot_columns, df.buzz_factor, test_size=0.4, random_state=None)
x_train.shape
#linear regression
from sklearn.linear_model import LinearRegression
model = LinearRegression(normalize=True)
model.fit(x_train, y_train)
from sklearn.metrics import mean_squared_error
predictions = model.predict(x_test)
mean_squared_error(predictions, y_test)
# Import the random forest model.
from sklearn.ensemble import RandomForestRegressor
model2 = RandomForestRegressor(n_estimators=60, min_samples_leaf=10, random_state=1)
model2.fit(x_train, y_train)
predictions = model2.predict(x_test)
mean_squared_error(predictions, y_test)
```
### A brief exploration
```
df[
['shares','n_tokens_title', 'n_tokens_content', 'n_unique_tokens']
].describe()
# Avg number of words in title and popular article content
popular_articles = df.ix[data_frame['shares'] >= 1400]
popular_articles[
['shares','n_tokens_title', 'n_tokens_content','n_unique_tokens']
].describe()
# Mean shares for each article type
type_articles = df.pivot_table('shares', index=[
'is_lifestyle', 'is_entertainment', 'is_business', 'is_social_media', 'is_tech', 'is_world'
], aggfunc=[np.mean])
print type_articles
type_articles.plot(kind='bar', color=red)
plt.legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0.)
plt.title('Mean shares by article type')
plt.xlabel('Article type')
plt.ylabel('Shares')
#### On avg, which day has more shares
# On avg, which day has more shares
day_articles = df.pivot_table('shares', index=[
'is_monday', 'is_tuesday', 'is_wednesday', 'is_thursday', 'is_friday', 'is_saturday', 'is_sunday'
], aggfunc=[np.mean])
print day_articles
day_articles.plot(kind='bar', color=green)
plt.legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0.)
plt.title('Mean shares by day')
plt.xlabel('Day')
plt.ylabel('Shares')
# Mean shares for tech and not tech channels during and not during weekends
df.pivot_table('shares', index=['is_weekend'], columns=['is_tech'], aggfunc=[np.mean], margins=True)
# Mean tech shares during work week (Monday to Friday)
tech_articles = df.ix[data_frame['is_tech'] == 1]
tech_articles = tech_articles.pivot_table('shares', index=[
'is_monday', 'is_tuesday', 'is_wednesday', 'is_thursday', 'is_friday'
], aggfunc=[np.mean])
print tech_articles
tech_articles.plot(kind='bar', color=blue)
plt.legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0.)
plt.title('Mean Tech Article Shares by Week Day')
plt.xlabel('Week Day')
plt.ylabel('Shares')
# Explore relationship with some features and number of shares
df.plot(kind='scatter', x='n_tokens_title', y='shares')
df.plot(kind='scatter', x='n_tokens_content', y='shares')
df.plot(kind='scatter', x='n_unique_tokens', y='shares')
json = df.to_json()
print json
```
---
*------------------------
### Problem 4: Prepare a 90 second Pitch and *present* it in the class (20 points)
* Prepare the slide(s) for the Pitch (10 points)
* Present it in the class (10 points).
*Advice: It should really only be one or two slides, but a really good one or two slides! Also, it is ok to select one person on the team to give the 90 second pitch (though a very organized multi-person 90 second pitch can be very impressive!) *
# Report: communicate the results (20 points)
(1) (5 points) What is your business proposition?
(2) (5 points) Why this topic is interesting or important to you? (Motivations)
(3) (5 points) How did you analyse the data?
(4) (5 points) How does your analysis support your business proposition?
(please include figures or tables in the report, but no source code)
# Slides (for 10 minutes of presentation) (20 points)
1. (5 points) Motivation about the data collection, why the topic is interesting to you.
2. (10 points) Communicating Results (figure/table)
3. (5 points) Story telling (How all the parts (data, analysis, result) fit together as a story?)
*-----------------
# Done
All set!
** What do you need to submit?**
* **Notebook File**: Save this IPython notebook, and find the notebook file in your folder (for example, "filename.ipynb"). This is the file you need to submit. Please make sure all the plotted tables and figures are in the notebook. If you used "ipython notebook --pylab=inline" to open the notebook, all the figures and tables should have shown up in the notebook.
* **PPT Slides**: **NOTE, for this Case Study you need to prepare two (2) PPT files!** One for the 90 second Pitch and one for a normal 10 minute presentation.
* ** Report**: please prepare a report (less than 10 pages) to report what you found in the data.
(please include figures or tables in the report, but no source code)
*Please compress all the files into a single zipped file.*
** How to submit: **
Send an email to rcpaffenroth@wpi.edu with the subject: "[DS501] Case study 4".
| github_jupyter |
```
from pytorch_tabnet.tab_model import TabNetRegressor
import torch
from sklearn.preprocessing import LabelEncoder
from sklearn.metrics import mean_squared_error
import pandas as pd
import numpy as np
np.random.seed(0)
import os
import wget
from pathlib import Path
```
# Download census-income dataset
```
url = "https://archive.ics.uci.edu/ml/machine-learning-databases/adult/adult.data"
dataset_name = 'census-income'
out = Path(os.getcwd()+'/data/'+dataset_name+'.csv')
out.parent.mkdir(parents=True, exist_ok=True)
if out.exists():
print("File already exists.")
else:
print("Downloading file...")
wget.download(url, out.as_posix())
```
# Load data and split
```
train = pd.read_csv(out)
target = ' <=50K'
if "Set" not in train.columns:
train["Set"] = np.random.choice(["train", "valid", "test"], p =[.8, .1, .1], size=(train.shape[0],))
train_indices = train[train.Set=="train"].index
valid_indices = train[train.Set=="valid"].index
test_indices = train[train.Set=="test"].index
```
# Simple preprocessing
Label encode categorical features and fill empty cells.
```
categorical_columns = []
categorical_dims = {}
for col in train.columns[train.dtypes == object]:
print(col, train[col].nunique())
l_enc = LabelEncoder()
train[col] = train[col].fillna("VV_likely")
train[col] = l_enc.fit_transform(train[col].values)
categorical_columns.append(col)
categorical_dims[col] = len(l_enc.classes_)
for col in train.columns[train.dtypes == 'float64']:
train.fillna(train.loc[train_indices, col].mean(), inplace=True)
```
# Define categorical features for categorical embeddings
```
unused_feat = ['Set']
features = [ col for col in train.columns if col not in unused_feat+[target]]
cat_idxs = [ i for i, f in enumerate(features) if f in categorical_columns]
cat_dims = [ categorical_dims[f] for i, f in enumerate(features) if f in categorical_columns]
# define your embedding sizes : here just a random choice
cat_emb_dim = [5, 4, 3, 6, 2, 2, 1, 10]
```
# Network parameters
```
clf = TabNetRegressor(cat_dims=cat_dims, cat_emb_dim=cat_emb_dim, cat_idxs=cat_idxs)
```
# Training
```
X_train = train[features].values[train_indices]
y_train = train[target].values[train_indices].reshape(-1, 1)
X_valid = train[features].values[valid_indices]
y_valid = train[target].values[valid_indices].reshape(-1, 1)
X_test = train[features].values[test_indices]
y_test = train[target].values[test_indices].reshape(-1, 1)
max_epochs = 1000 if not os.getenv("CI", False) else 2
clf.fit(
X_train=X_train, y_train=y_train,
X_valid=X_valid, y_valid=y_valid,
max_epochs=max_epochs,
patience=50,
batch_size=1024, virtual_batch_size=128,
num_workers=0,
drop_last=False
)
# Deprecated : best model is automatically loaded at end of fit
# clf.load_best_model()
preds = clf.predict(X_test)
y_true = y_test
test_score = mean_squared_error(y_pred=preds, y_true=y_true)
print(f"BEST VALID SCORE FOR {dataset_name} : {clf.best_cost}")
print(f"FINAL TEST SCORE FOR {dataset_name} : {test_score}")
```
# Global explainability : feat importance summing to 1
```
clf.feature_importances_
```
# Local explainability and masks
```
explain_matrix, masks = clf.explain(X_test)
from matplotlib import pyplot as plt
%matplotlib inline
fig, axs = plt.subplots(1, 3, figsize=(20,20))
for i in range(3):
axs[i].imshow(masks[i][:50])
axs[i].set_title(f"mask {i}")
```
# XGB
```
from xgboost import XGBRegressor
clf_xgb = XGBRegressor(max_depth=8,
learning_rate=0.1,
n_estimators=1000,
verbosity=0,
silent=None,
objective='reg:linear',
booster='gbtree',
n_jobs=-1,
nthread=None,
gamma=0,
min_child_weight=1,
max_delta_step=0,
subsample=0.7,
colsample_bytree=1,
colsample_bylevel=1,
colsample_bynode=1,
reg_alpha=0,
reg_lambda=1,
scale_pos_weight=1,
base_score=0.5,
random_state=0,
seed=None,)
clf_xgb.fit(X_train, y_train,
eval_set=[(X_valid, y_valid)],
early_stopping_rounds=40,
verbose=10)
preds = np.array(clf_xgb.predict(X_valid))
valid_auc = mean_squared_error(y_pred=preds, y_true=y_valid)
print(valid_auc)
preds = np.array(clf_xgb.predict(X_test))
test_auc = mean_squared_error(y_pred=preds, y_true=y_test)
print(test_auc)
```
| github_jupyter |
```
import numpy as np
import matplotlib.pyplot as plt
from scipy.interpolate import interp1d
import re
import ast
%matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
f, axarr = plt.subplots(2, 3)
x = range(1, 50)
#------------------------------------------------------------------------------------
file_scores = open('fusion_data/leaf_flower_top_50_scores.txt', 'r')
for line in file_scores:
if line.split(':')[0] == 'max rule':
y_max = ast.literal_eval(line.split(':')[1].strip())
elif line.split(':')[0] == 'sum rule':
y_sum = ast.literal_eval(line.split(':')[1].strip())
elif line.split(':')[0] == 'prod rule':
y_prod = ast.literal_eval(line.split(':')[1].strip())
elif line.split(':')[0] == 'leaf':
y_or1 = ast.literal_eval(line.split(':')[1].strip())
elif line.split(':')[0] == 'flower':
y_or2 = ast.literal_eval(line.split(':')[1].strip())
elif line.split(':')[0] == 'rh rule':
y_rh = ast.literal_eval(line.split(':')[1].strip())
else:
y_svm = ast.literal_eval(line.split(':')[1].strip())
file_scores.close()
f_svm = interp1d(x, y_svm, kind='cubic')
f_max = interp1d(x, y_max, kind='cubic')
f_sum = interp1d(x, y_sum, kind='cubic')
f_prod = interp1d(x, y_prod, kind='cubic')
f_rh = interp1d(x, y_rh, kind='cubic')
f_or1 = interp1d(x, y_or1, kind='cubic')
f_or2 = interp1d(x, y_or2, kind='cubic')
l1, l2, l3, l4, l5, l6, l7 = axarr[0, 0].plot(x, f_prod(x), '-', x, f_sum(x), '-', x, f_rh(x), '-',
x, f_svm(x), '-', x, f_max(x), '-', x, f_or1(x), '-', x, f_or2(x), '-')
axarr[0, 0].set_title('leaf vs flower')
axarr[0, 0].set_ylim([0.3,1])
axarr[0, 0].set_xlim([0,20])
#------------------------------------------------------------------------------------
file_scores = open('fusion_data/leaf_entire_top_50_scores.txt', 'r')
for line in file_scores:
if line.split(':')[0] == 'max rule':
y_max = ast.literal_eval(line.split(':')[1].strip())
elif line.split(':')[0] == 'sum rule':
y_sum = ast.literal_eval(line.split(':')[1].strip())
elif line.split(':')[0] == 'prod rule':
y_prod = ast.literal_eval(line.split(':')[1].strip())
elif line.split(':')[0] == 'leaf':
y_or1 = ast.literal_eval(line.split(':')[1].strip())
elif line.split(':')[0] == 'entire':
y_or2 = ast.literal_eval(line.split(':')[1].strip())
elif line.split(':')[0] == 'rh rule':
y_rh = ast.literal_eval(line.split(':')[1].strip())
else:
y_svm = ast.literal_eval(line.split(':')[1].strip())
file_scores.close()
f_svm = interp1d(x, y_svm, kind='cubic')
f_max = interp1d(x, y_max, kind='cubic')
f_sum = interp1d(x, y_sum, kind='cubic')
f_prod = interp1d(x, y_prod, kind='cubic')
f_rh = interp1d(x, y_rh, kind='cubic')
f_or1 = interp1d(x, y_or1, kind='cubic')
f_or2 = interp1d(x, y_or2, kind='cubic')
axarr[0, 1].plot(x, f_prod(x), '-', x, f_sum(x), '-', x, f_rh(x), '-',
x, f_svm(x), '-', x, f_max(x), '-', x, f_or1(x), '-', x, f_or2(x), '-')
axarr[0, 1].set_title('leaf vs entire')
axarr[0, 1].set_ylim([0.3,1])
axarr[0, 1].set_xlim([0,20])
#------------------------------------------------------------------------------------
file_scores = open('fusion_data/entire_flower_top_50_scores.txt', 'r')
for line in file_scores:
if line.split(':')[0] == 'max rule':
y_max = ast.literal_eval(line.split(':')[1].strip())
elif line.split(':')[0] == 'sum rule':
y_sum = ast.literal_eval(line.split(':')[1].strip())
elif line.split(':')[0] == 'prod rule':
y_prod = ast.literal_eval(line.split(':')[1].strip())
elif line.split(':')[0] == 'entire':
y_or1 = ast.literal_eval(line.split(':')[1].strip())
elif line.split(':')[0] == 'flower':
y_or2 = ast.literal_eval(line.split(':')[1].strip())
elif line.split(':')[0] == 'rh rule':
y_rh = ast.literal_eval(line.split(':')[1].strip())
else:
y_svm = ast.literal_eval(line.split(':')[1].strip())
file_scores.close()
f_svm = interp1d(x, y_svm, kind='cubic')
f_max = interp1d(x, y_max, kind='cubic')
f_sum = interp1d(x, y_sum, kind='cubic')
f_prod = interp1d(x, y_prod, kind='cubic')
f_rh = interp1d(x, y_rh, kind='cubic')
f_or1 = interp1d(x, y_or1, kind='cubic')
f_or2 = interp1d(x, y_or2, kind='cubic')
axarr[1, 0].plot(x, f_prod(x), '-', x, f_sum(x), '-', x, f_rh(x), '-',
x, f_svm(x), '-', x, f_max(x), '-', x, f_or1(x), '-', x, f_or2(x), '-')
axarr[1, 0].set_title('entire vs flower')
axarr[1, 0].set_ylim([0.3,1])
axarr[1, 0].set_xlim([0,20])
#------------------------------------------------------------------------------------
file_scores = open('fusion_data/branch_leaf_top_50_scores.txt', 'r')
for line in file_scores:
if line.split(':')[0] == 'max rule':
y_max = ast.literal_eval(line.split(':')[1].strip())
elif line.split(':')[0] == 'sum rule':
y_sum = ast.literal_eval(line.split(':')[1].strip())
elif line.split(':')[0] == 'prod rule':
y_prod = ast.literal_eval(line.split(':')[1].strip())
elif line.split(':')[0] == 'branch':
y_or1 = ast.literal_eval(line.split(':')[1].strip())
elif line.split(':')[0] == 'leaf':
y_or2 = ast.literal_eval(line.split(':')[1].strip())
elif line.split(':')[0] == 'rh rule':
y_rh = ast.literal_eval(line.split(':')[1].strip())
else:
y_svm = ast.literal_eval(line.split(':')[1].strip())
file_scores.close()
f_svm = interp1d(x, y_svm, kind='cubic')
f_max = interp1d(x, y_max, kind='cubic')
f_sum = interp1d(x, y_sum, kind='cubic')
f_prod = interp1d(x, y_prod, kind='cubic')
f_rh = interp1d(x, y_rh, kind='cubic')
f_or1 = interp1d(x, y_or1, kind='cubic')
f_or2 = interp1d(x, y_or2, kind='cubic')
axarr[1, 1].plot(x, f_prod(x), '-', x, f_sum(x), '-', x, f_rh(x), '-',
x, f_svm(x), '-', x, f_max(x), '-', x, f_or1(x), '-', x, f_or2(x), '-')
axarr[1, 1].set_title('branch vs leaf')
axarr[1, 1].set_ylim([0.3,1])
axarr[1, 1].set_xlim([0,20])
#------------------------------------------------------------------------------------
file_scores = open('fusion_data/branch_flower_top_50_scores.txt', 'r')
for line in file_scores:
if line.split(':')[0] == 'max rule':
y_max = ast.literal_eval(line.split(':')[1].strip())
elif line.split(':')[0] == 'sum rule':
y_sum = ast.literal_eval(line.split(':')[1].strip())
elif line.split(':')[0] == 'prod rule':
y_prod = ast.literal_eval(line.split(':')[1].strip())
elif line.split(':')[0] == 'branch':
y_or1 = ast.literal_eval(line.split(':')[1].strip())
elif line.split(':')[0] == 'flower':
y_or2 = ast.literal_eval(line.split(':')[1].strip())
elif line.split(':')[0] == 'rh rule':
y_rh = ast.literal_eval(line.split(':')[1].strip())
else:
y_svm = ast.literal_eval(line.split(':')[1].strip())
file_scores.close()
f_svm = interp1d(x, y_svm, kind='cubic')
f_max = interp1d(x, y_max, kind='cubic')
f_sum = interp1d(x, y_sum, kind='cubic')
f_prod = interp1d(x, y_prod, kind='cubic')
f_rh = interp1d(x, y_rh, kind='cubic')
f_or1 = interp1d(x, y_or1, kind='cubic')
f_or2 = interp1d(x, y_or2, kind='cubic')
axarr[0, 2].plot(x, f_prod(x), '-', x, f_sum(x), '-', x, f_rh(x), '-',
x, f_svm(x), '-', x, f_max(x), '-', x, f_or1(x), '-', x, f_or2(x), '-')
axarr[0, 2].set_title('branch vs flower')
axarr[0, 2].set_ylim([0.3,1])
axarr[0, 2].set_xlim([0,20])
#------------------------------------------------------------------------------------
file_scores = open('fusion_data/branch_entire_top_50_scores.txt', 'r')
for line in file_scores:
if line.split(':')[0] == 'max rule':
y_max = ast.literal_eval(line.split(':')[1].strip())
elif line.split(':')[0] == 'sum rule':
y_sum = ast.literal_eval(line.split(':')[1].strip())
elif line.split(':')[0] == 'prod rule':
y_prod = ast.literal_eval(line.split(':')[1].strip())
elif line.split(':')[0] == 'branch':
y_or1 = ast.literal_eval(line.split(':')[1].strip())
elif line.split(':')[0] == 'entire':
y_or2 = ast.literal_eval(line.split(':')[1].strip())
elif line.split(':')[0] == 'rh rule':
y_rh = ast.literal_eval(line.split(':')[1].strip())
else:
y_svm = ast.literal_eval(line.split(':')[1].strip())
file_scores.close()
f_svm = interp1d(x, y_svm, kind='cubic')
f_max = interp1d(x, y_max, kind='cubic')
f_sum = interp1d(x, y_sum, kind='cubic')
f_prod = interp1d(x, y_prod, kind='cubic')
f_rh = interp1d(x, y_rh, kind='cubic')
f_or1 = interp1d(x, y_or1, kind='cubic')
f_or2 = interp1d(x, y_or2, kind='cubic')
axarr[1, 2].plot(x, f_prod(x), '-', x, f_sum(x), '-', x, f_rh(x), '-',
x, f_svm(x), '-', x, f_max(x), '-', x, f_or1(x), '-', x, f_or2(x), '-')
axarr[1, 2].set_title('branch vs entire')
axarr[1, 2].set_ylim([0.2,1])
axarr[1, 2].set_xlim([0,20])
axarr[1, 2].set_xticks(np.arange(0, 21, 5))
# Fine-tune figure; hide x ticks for top plots and y ticks for right plots
plt.setp([a.get_xticklabels() for a in axarr[0, :]], visible=False)
plt.setp([a.get_yticklabels() for a in axarr[:, 1]], visible=False)
plt.setp([a.get_yticklabels() for a in axarr[:, 2]], visible=False)
f.legend((l1, l2, l3, l4, l5, l6, l7), ('Product rule', 'Sum rule','RHF',
'SVM', 'Max rule','Organ 1','Organ 2'), 'lower center',
ncol=7, frameon=False)
plt.show()
```
| github_jupyter |
# Linear regression baseline
In this notebook, we will create the linear regression baselines.
```
%load_ext autoreload
%autoreload 2
import numpy as np
import matplotlib.pyplot as plt
import xarray as xr
import seaborn as sns
import pickle
from src.score import *
from sklearn.linear_model import LinearRegression
from sklearn.metrics import mean_squared_error
from tqdm import tqdm_notebook as tqdm
sns.set_style('darkgrid')
sns.set_context('notebook')
def to_pickle(obj, fn):
with open(fn, 'wb') as f:
pickle.dump(obj, f)
def read_pickle(fn):
with open(fn, 'rb') as f:
return pickle.load(f)
```
## Load and prepare data for training
First up, we need to load and prepare the data so that we can feed it into our linear regression model.
```
DATADIR = '/data/WeatherBench/5.625deg/'
PREDDIR = '/data/WeatherBench/baselines/'
# DATADIR = '/data/stephan/WeatherBench/5.625deg/'
# PREDDIR = '/data/stephan/WeatherBench/baselines/'
# Load the entire dataset for the relevant variables
z500 = xr.open_mfdataset(f'{DATADIR}geopotential_500/*.nc', combine='by_coords').z
t850 = xr.open_mfdataset(f'{DATADIR}temperature_850/*.nc', combine='by_coords').t.drop('level')
tp = xr.open_mfdataset(f'{DATADIR}total_precipitation/*.nc', combine='by_coords').tp.rolling(time=6).sum()
tp.name = 'tp'
t2m = xr.open_mfdataset(f'{DATADIR}2m_temperature/*.nc', combine='by_coords').t2m
data = xr.merge([z500, t850, tp, t2m])
# Load the validation subset of the data: 2017 and 2018
z500_test = load_test_data(f'{DATADIR}geopotential_500', 'z')
t850_test = load_test_data(f'{DATADIR}temperature_850', 't')
tp_test = load_test_data(f'{DATADIR}total_precipitation', 'tp').rolling(time=6).sum()
tp_test.name = 'tp'
t2m_test = load_test_data(f'{DATADIR}2m_temperature', 't2m')
test_data = xr.merge([z500_test, t850_test, tp_test, t2m_test])
# The first 7+6 values of tp are missing, so let's just start after that
data.tp[:6+7+1].mean(('lat', 'lon')).compute()
data = data.isel(time=slice(7+6, None))
# Split into train and test data
# Yes, technically we should have a separate validation set but for LR this shouldn't matter.
data_train = data.sel(time=slice('1979', '2016'))
# data_train = data.sel(time=slice('2016', '2016'))
data_test = data.sel(time=slice('2017', '2018'))
# Compute normalization statistics
# Let's only take a sample of the time to speed this up.
data_mean = data_train.isel(time=slice(0, None, 10000)).mean().load()
data_std = data_train.isel(time=slice(0, None, 10000)).std().load()
# Normalize datasets
data_train = (data_train - data_mean) / data_std
data_test = (data_test - data_mean) / data_std
_, nlat, nlon = data_train.z.shape; nlat, nlon
data_train.z.isel(time=0).plot()
data_train.t.isel(time=0).plot()
data_train
def create_training_data(da, lead_time_h, return_valid_time=False):
"""Function to split input and output by lead time."""
X = da.isel(time=slice(0, -lead_time_h))
y = da.isel(time=slice(lead_time_h, None))
valid_time = y.time
if return_valid_time:
return X.values.reshape(-1, nlat*nlon), y.values.reshape(-1, nlat*nlon), valid_time
else:
return X.values.reshape(-1, nlat*nlon), y.values.reshape(-1, nlat*nlon)
```
## Train linear regression
Now let's train the model. We will use scikit-learn for this.
```
def train_lr(lead_time_h, input_vars, output_vars, data_subsample=1):
"""Create data, train a linear regression and return the predictions."""
X_train, y_train, X_test, y_test = [], [], [], []
for v in input_vars:
X, y = create_training_data(
data_train[v],
lead_time_h
)
X_train.append(X)
if v in output_vars: y_train.append(y)
X, y, valid_time = create_training_data(data_test[v], lead_time_h, return_valid_time=True)
X_test.append(X)
if v in output_vars: y_test.append(y)
X_train, y_train, X_test, y_test = [np.concatenate(d, 1) for d in [X_train, y_train, X_test, y_test]]
X_train = X_train[::data_subsample]
y_train = y_train[::data_subsample]
lr = LinearRegression(n_jobs=16)
lr.fit(X_train, y_train)
mse_train = mean_squared_error(y_train, lr.predict(X_train))
mse_test = mean_squared_error(y_test, lr.predict(X_test))
print(f'Train MSE = {mse_train}'); print(f'Test MSE = {mse_test}')
preds = lr.predict(X_test).reshape((-1, len(output_vars), nlat, nlon))
fcs = []
for i, v in enumerate(output_vars):
fc = xr.DataArray(
preds[:, i] * data_std[v].values + data_mean[v].values,
dims=['time', 'lat', 'lon'],
coords={
'time': valid_time,
'lat': data_train.lat,
'lon': data_train.lon
},
name=v
)
fcs.append(fc)
return xr.merge(fcs), lr
```
### 3 days
Here we train a model to directly predict the fields at 3 days lead time. Let's train a model that only predicts z or t and then a combined model. As we can see below, the model trained only on Z500 performs better than the combined model. But the same is not the case for T850. For the paper, we will use the combined model.
```
experiments = [
[['z'], ['z']],
[['t'], ['t']],
[['z', 't'], ['z', 't']],
[['tp'], ['tp']],
[['z', 't', 'tp'], ['tp']],
[['t2m'], ['t2m']],
[['z', 't', 't2m'], ['t2m']],
]
# Since training the LR on the full data takes up quite a lot of memory
# we only take every 5th time step which gives almost the same results (<0.5% difference)
data_subsample = 5
lead_time = 3*24
preds = []
models = []
for n, (i, o) in enumerate(experiments):
print(f'{n}: Input variables = {i}; output variables = {o}')
p, m = train_lr(lead_time, input_vars=i, output_vars=o, data_subsample=data_subsample)
preds.append(p); models.append(m)
r = compute_weighted_rmse(p, test_data).compute()
print('; '.join([f'{v} = {r[v].values}' for v in r]) + '\n')
p.to_netcdf(f'{PREDDIR}/lr_3d_{"_".join(i)}_{"_".join(o)}.nc');
to_pickle(m, f'{PREDDIR}/saved_models/lr_3d_{"_".join(i)}_{"_".join(o)}.pkl')
```
As we can see the models with just the output variable as input almost always perform better because of overfitting. We could try a regularized regression like ridge of lasso but the point of these models not to be good but rather to provide a solid baseline with as few hyperparameters as possible.
```
# Same for 5 days
data_subsample = 5
lead_time = 5*24
preds = []
models = []
for n, (i, o) in enumerate(experiments):
print(f'{n}: Input variables = {i}; output variables = {o}')
p, m = train_lr(lead_time, input_vars=i, output_vars=o, data_subsample=data_subsample)
preds.append(p); models.append(m)
r = compute_weighted_rmse(p, test_data).compute()
print('; '.join([f'{v} = {r[v].values}' for v in r]) + '\n')
p.to_netcdf(f'{PREDDIR}/lr_5d_{"_".join(i)}_{"_".join(o)}.nc');
to_pickle(m, f'{PREDDIR}/saved_models/lr_5d_{"_".join(i)}_{"_".join(o)}.pkl')
```
### Iterative forecast
Finally, an iterative forecast. First, we train a model for 6 hours lead time and then construct an iterative forecast up to 120 hours.
```
def create_iterative_fc(state, model, lead_time_h=6, max_lead_time_h=5*24):
max_fc_steps = max_lead_time_h // lead_time_h
fcs_z500, fcs_t850 = [], []
for fc_step in tqdm(range(max_fc_steps)):
state = model.predict(state)
fc_z500 = state[:, :nlat*nlon].copy() * data_std.z.values + data_mean.z.values
fc_t850 = state[:, nlat*nlon:].copy() * data_std.t.values + data_mean.t.values
fc_z500 = fc_z500.reshape((-1, nlat, nlon))
fc_t850 = fc_t850.reshape((-1, nlat, nlon))
fcs_z500.append(fc_z500); fcs_t850.append(fc_t850)
return [xr.DataArray(
np.array(fcs),
dims=['lead_time', 'time', 'lat', 'lon'],
coords={
'lead_time': np.arange(lead_time_h, max_lead_time_h + lead_time_h, lead_time_h),
'time': z500_test.time,
'lat': z500_test.lat,
'lon': z500_test.lon
}
) for fcs in [fcs_z500, fcs_t850]]
p, m = train_lr(6, input_vars=['z', 't'], output_vars=['z', 't'], data_subsample=5)
to_pickle(m, f'{PREDDIR}/saved_models/lr_6h_z_t_z_t.pkl')
m = read_pickle(f'{PREDDIR}/saved_models/lr_6h_z_t_z_t.pkl')
state = np.concatenate([data_test.z.values.reshape(-1, nlat*nlon),
data_test.t.values.reshape(-1, nlat*nlon)], 1)
fc_z500_6h_iter, fc_t850_6h_iter = create_iterative_fc(state, m)
fc_iter = xr.Dataset({'z': fc_z500_6h_iter, 't': fc_t850_6h_iter})
fc_iter.to_netcdf(f'{PREDDIR}/lr_6h_iter.nc');
```
# The End
| github_jupyter |
# Predicting sentiment from product reviews
# Fire up GraphLab Create
(See [Getting Started with SFrames](/notebooks/Week%201/Getting%20Started%20with%20SFrames.ipynb) for setup instructions)
```
import graphlab
# Limit number of worker processes. This preserves system memory, which prevents hosted notebooks from crashing.
graphlab.set_runtime_config('GRAPHLAB_DEFAULT_NUM_PYLAMBDA_WORKERS', 4)
```
# Read some product review data
Loading reviews for a set of baby products.
```
products = graphlab.SFrame('amazon_baby.gl/')
```
# Let's explore this data together
Data includes the product name, the review text and the rating of the review.
```
products.head()
```
# Build the word count vector for each review
```
products['word_count'] = graphlab.text_analytics.count_words(products['review'])
products.head()
graphlab.canvas.set_target('ipynb')
products['name'].show()
```
# Examining the reviews for most-sold product: 'Vulli Sophie the Giraffe Teether'
```
giraffe_reviews = products[products['name'] == 'Vulli Sophie the Giraffe Teether']
len(giraffe_reviews)
giraffe_reviews['rating'].show(view='Categorical')
```
# Build a sentiment classifier
```
products['rating'].show(view='Categorical')
```
## Define what's a positive and a negative sentiment
We will ignore all reviews with rating = 3, since they tend to have a neutral sentiment. Reviews with a rating of 4 or higher will be considered positive, while the ones with rating of 2 or lower will have a negative sentiment.
```
# ignore all 3* reviews
products = products[products['rating'] != 3]
# positive sentiment = 4* or 5* reviews
products['sentiment'] = products['rating'] >=4
products.head()
```
## Let's train the sentiment classifier
```
train_data,test_data = products.random_split(.8, seed=0)
sentiment_model = graphlab.logistic_classifier.create(train_data,
target='sentiment',
features=['word_count'],
validation_set=test_data)
```
# Evaluate the sentiment model
```
sentiment_model.evaluate(test_data, metric='roc_curve')
sentiment_model.show(view='Evaluation')
```
# Applying the learned model to understand sentiment for Giraffe
```
giraffe_reviews['predicted_sentiment'] = sentiment_model.predict(giraffe_reviews, output_type='probability')
giraffe_reviews.head()
```
## Sort the reviews based on the predicted sentiment and explore
```
giraffe_reviews = giraffe_reviews.sort('predicted_sentiment', ascending=False)
giraffe_reviews.head()
```
## Most positive reviews for the giraffe
```
giraffe_reviews[0]['review']
giraffe_reviews[1]['review']
```
## Show most negative reviews for giraffe
```
giraffe_reviews[-1]['review']
giraffe_reviews[-2]['review']
```
## Quizzes
```
selected_words = [
'awesome',
'great',
'fantastic',
'amazing',
'love',
'horrible',
'bad',
'terrible',
'awful',
'wow',
'hate']
```
### 1. Out of the 11 words in selected_words, which one is most used in the reviews in the dataset?
```
d = {}
for words in products['word_count']:
for word in words:
if word in selected_words:
if word not in d:
d[word] = words[word]
else:
d[word] += words[word]
d
results = sorted(d.items(), key=lambda x:x[1], reverse=True)
results
results[0]
```
### 2. Out of the 11 words in selected_words, which one is least used in the reviews in the dataset?
```
results[-1]
```
### 3. Out of the 11 words in selected_words, which one got the most positive weight in the selected_words_model?
```
# One hot encoding
for word in selected_words:
products[word] = products['word_count'].apply(
lambda word_count: word_count[word] if word in word_count else 0)
products.head()
train_data,test_data = products.random_split(.8, seed=0)
selected_words_model = graphlab.logistic_classifier.create(train_data,
target='sentiment',
features=selected_words,
validation_set=test_data)
selected_words_model['coefficients']
df = selected_words_model['coefficients'][1:]
max_positive = df['value'].max()
df[df['value'] == max_positive]
```
### 4.Out of the 11 words in selected_words, which one got the most negative weight in the selected_words_model?
```
df = selected_words_model['coefficients'][1:]
max_negative = df['value'].min()
df[df['value'] == max_negative]
```
### 5.Which of the following ranges contains the accuracy of the selected_words_model on the test_data?
```
selected_words_model.evaluate(test_data)
selected_words_model.evaluate(test_data)['accuracy']
```
### 6.Which of the following ranges contains the accuracy of the sentiment_model in the IPython Notebook from lecture on the test_data?
```
sentiment_model.evaluate(test_data)
sentiment_model.evaluate(test_data)['accuracy']
```
### 7.Which of the following ranges contains the accuracy of the majority class classifier, which simply predicts the majority class on the test_data?
```
total = test_data.num_rows()
total
positives = test_data[test_data['rating'] >= 4].num_rows()
positives
negatives = test_data[test_data['rating'] <= 2].num_rows()
negatives
majority_accuracy = float(positives) / total
majority_accuracy
```
### 8. How do you compare the different learned models with the baseline approach where we are just predicting the majority class?
- They all performed about the same.
- The model learned using all words performed much better than the other two. The other two approaches performed about the same.
- [x] The model learned using all words performed much better than the other two. The other two approaches performed about the same.
- Predicting the simply majority class performed much better than the other two models.
### 9.Which of the following ranges contains the ‘predicted_sentiment’ for the most positive review for ‘Baby Trend Diaper Champ’, according to the sentiment_model from the IPython Notebook from lecture?
```
diaper_champ_reviews = products[products['name']=='Baby Trend Diaper Champ']
diaper_champ_reviews.head()
sentiment_model.predict(diaper_champ_reviews[diaper_champ_reviews['rating'] == 5.0], output_type='probability').mean()
```
### 10.Consider the most positive review for ‘Baby Trend Diaper Champ’ according to the sentiment_model from the IPython Notebook from lecture. Which of the following ranges contains the predicted_sentiment for this review, if we use the selected_words_model to analyze it?
```
selected_words_model.predict(diaper_champ_reviews[diaper_champ_reviews['rating']][0:10], output_type='probability').mean()
```
### 11.Why is the value of the predicted_sentiment for the most positive review found using the sentiment_model much more positive than the value predicted using the selected_words_model?
- The sentiment_model is just too positive about everything.
- The selected_words_model is just too negative about everything.
- This review was positive, but used too many of the negative words in selected_words.
- [x]None of the selected_words appeared in the text of this review.
| github_jupyter |
# Developing an AI application
Going forward, AI algorithms will be incorporated into more and more everyday applications. For example, you might want to include an image classifier in a smart phone app. To do this, you'd use a deep learning model trained on hundreds of thousands of images as part of the overall application architecture. A large part of software development in the future will be using these types of models as common parts of applications.
In this project, you'll train an image classifier to recognize different species of flowers. You can imagine using something like this in a phone app that tells you the name of the flower your camera is looking at. In practice you'd train this classifier, then export it for use in your application. We'll be using [this dataset](http://www.robots.ox.ac.uk/~vgg/data/flowers/102/index.html) of 102 flower categories, you can see a few examples below.
<img src='assets/Flowers.png' width=500px>
The project is broken down into multiple steps:
* Load and preprocess the image dataset
* Train the image classifier on your dataset
* Use the trained classifier to predict image content
We'll lead you through each part which you'll implement in Python.
When you've completed this project, you'll have an application that can be trained on any set of labeled images. Here your network will be learning about flowers and end up as a command line application. But, what you do with your new skills depends on your imagination and effort in building a dataset. For example, imagine an app where you take a picture of a car, it tells you what the make and model is, then looks up information about it. Go build your own dataset and make something new.
First up is importing the packages you'll need. It's good practice to keep all the imports at the beginning of your code. As you work through this notebook and find you need to import a package, make sure to add the import up here.
Please make sure if you are running this notebook in the workspace that you have chosen GPU rather than CPU mode.
```
# Imports here
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
import matplotlib.pyplot as plt
import numpy as np
from collections import OrderedDict
from PIL import Image
import time
import json
import torchvision
from torchvision import datasets, transforms, models
import torch
from torch import nn, optim
```
## Load the data
Here you'll use `torchvision` to load the data ([documentation](http://pytorch.org/docs/0.3.0/torchvision/index.html)). The data should be included alongside this notebook, otherwise you can [download it here](https://s3.amazonaws.com/content.udacity-data.com/nd089/flower_data.tar.gz). The dataset is split into three parts, training, validation, and testing. For the training, you'll want to apply transformations such as random scaling, cropping, and flipping. This will help the network generalize leading to better performance. You'll also need to make sure the input data is resized to 224x224 pixels as required by the pre-trained networks.
The validation and testing sets are used to measure the model's performance on data it hasn't seen yet. For this you don't want any scaling or rotation transformations, but you'll need to resize then crop the images to the appropriate size.
The pre-trained networks you'll use were trained on the ImageNet dataset where each color channel was normalized separately. For all three sets you'll need to normalize the means and standard deviations of the images to what the network expects. For the means, it's `[0.485, 0.456, 0.406]` and for the standard deviations `[0.229, 0.224, 0.225]`, calculated from the ImageNet images. These values will shift each color channel to be centered at 0 and range from -1 to 1.
```
data_dir = 'flowers'
train_dir = data_dir + '/train'
valid_dir = data_dir + '/valid'
test_dir = data_dir + '/test'
# TODO: Define your transforms for the training, validation, and testing sets
train_transforms = transforms.Compose([transforms.RandomRotation(30),
transforms.RandomResizedCrop(224),
transforms.RandomHorizontalFlip(),
transforms.ToTensor(),
transforms.Normalize([0.485, 0.456, 0.406],
[0.229, 0.224, 0.225])])
test_and_valid_transforms = transforms.Compose([transforms.Resize(256),
transforms.CenterCrop(224),
transforms.ToTensor(),
transforms.Normalize([0.485, 0.456, 0.406],
[0.229, 0.224, 0.225])])
# TODO: Load the datasets with ImageFolder
train_data = datasets.ImageFolder(train_dir, transform=train_transforms)
test_data = datasets.ImageFolder(test_dir, transform=test_and_valid_transforms)
valid_data = datasets.ImageFolder(valid_dir, transform=test_and_valid_transforms)
# TODO: Using the image datasets and the trainforms, define the dataloaders
trainloader = torch.utils.data.DataLoader(train_data, batch_size=32, shuffle=True)
testloader = torch.utils.data.DataLoader(test_data, batch_size=32, shuffle=True)
validloader = torch.utils.data.DataLoader(valid_data, batch_size=32, shuffle=True)
```
### Label mapping
You'll also need to load in a mapping from category label to category name. You can find this in the file `cat_to_name.json`. It's a JSON object which you can read in with the [`json` module](https://docs.python.org/2/library/json.html). This will give you a dictionary mapping the integer encoded categories to the actual names of the flowers.
```
with open('cat_to_name.json', 'r') as f:
cat_to_name = json.load(f)
print(cat_to_name)
```
# Building and training the classifier
Now that the data is ready, it's time to build and train the classifier. As usual, you should use one of the pretrained models from `torchvision.models` to get the image features. Build and train a new feed-forward classifier using those features.
We're going to leave this part up to you. Refer to [the rubric](https://review.udacity.com/#!/rubrics/1663/view) for guidance on successfully completing this section. Things you'll need to do:
* Load a [pre-trained network](http://pytorch.org/docs/master/torchvision/models.html) (If you need a starting point, the VGG networks work great and are straightforward to use)
* Define a new, untrained feed-forward network as a classifier, using ReLU activations and dropout
* Train the classifier layers using backpropagation using the pre-trained network to get the features
* Track the loss and accuracy on the validation set to determine the best hyperparameters
We've left a cell open for you below, but use as many as you need. Our advice is to break the problem up into smaller parts you can run separately. Check that each part is doing what you expect, then move on to the next. You'll likely find that as you work through each part, you'll need to go back and modify your previous code. This is totally normal!
When training make sure you're updating only the weights of the feed-forward network. You should be able to get the validation accuracy above 70% if you build everything right. Make sure to try different hyperparameters (learning rate, units in the classifier, epochs, etc) to find the best model. Save those hyperparameters to use as default values in the next part of the project.
One last important tip if you're using the workspace to run your code: To avoid having your workspace disconnect during the long-running tasks in this notebook, please read in the earlier page in this lesson called Intro to GPU Workspaces about Keeping Your Session Active. You'll want to include code from the workspace_utils.py module.
```
# TODO: Build and train your network
# Set device variable to GPU or CPU
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
# Load pre-trained network
model = models.densenet201(pretrained=True)
# Freeze params of pre-trained model
for param in model.parameters():
param.requires_grad = False
# Define new classifier
classifier = nn.Sequential(OrderedDict([('fc1', nn.Linear(1920, 960)),
('relu1', nn.ReLU()),
('dropout1', nn.Dropout(p=0.5)),
('fc2', nn.Linear(960, 102)),
('output', nn.LogSoftmax(dim=1))]))
model.classifier = classifier
# Train classifier
model.to(device)
criterion = nn.NLLLoss()
optimizer = optim.Adam(model.classifier.parameters(), lr=0.001)
epochs = 4
print_every = 40
steps = 0
start = time.time()
for e in range(epochs):
running_loss = 0
for inputs, labels in trainloader:
steps += 1
inputs, labels = inputs.to(device), labels.to(device)
optimizer.zero_grad()
outputs = model.forward(inputs)
loss = criterion(outputs, labels)
loss.backward()
optimizer.step()
running_loss += loss.item()
# Track loss and accuracy on validation set
if steps % print_every == 0:
model.eval()
with torch.no_grad():
valid_loss = 0
accuracy = 0
for inputs, labels in validloader:
inputs, labels = inputs.to(device), labels.to(device)
outputs = model.forward(inputs)
valid_loss += criterion(outputs, labels).item()
ps = torch.exp(outputs)
equality = (labels.data == ps.max(dim=1)[1])
accuracy += equality.type(torch.FloatTensor).mean()
print("Epoch: {}/{}.. ".format(e+1, epochs),
"Training Loss: {:.3f}.. ".format(running_loss/print_every),
"Valid Loss: {:.3f}.. ".format(valid_loss/len(validloader)),
"Test Accuracy: {:.3f}".format(accuracy/len(validloader)))
running_loss = 0
model.train()
total_time = time.time() - start
print("\n** Total time to train model:", str(
int((total_time / 3600))) + ":" +
str(int((total_time % 3600) / 60)) + ":" + str(
int((total_time % 3600) % 60)))
```
## Testing your network
It's good practice to test your trained network on test data, images the network has never seen either in training or validation. This will give you a good estimate for the model's performance on completely new images. Run the test images through the network and measure the accuracy, the same way you did validation. You should be able to reach around 70% accuracy on the test set if the model has been trained well.
```
# TODO: Do validation on the test set
model.eval()
correct = 0
total = 0
with torch.no_grad():
for inputs, labels in testloader:
inputs, labels = inputs.to(device), labels.to(device)
outputs = model.forward(inputs)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the test images: %d %%' % (100 * correct / total))
```
## Save the checkpoint
Now that your network is trained, save the model so you can load it later for making predictions. You probably want to save other things such as the mapping of classes to indices which you get from one of the image datasets: `image_datasets['train'].class_to_idx`. You can attach this to the model as an attribute which makes inference easier later on.
```model.class_to_idx = image_datasets['train'].class_to_idx```
Remember that you'll want to completely rebuild the model later so you can use it for inference. Make sure to include any information you need in the checkpoint. If you want to load the model and keep training, you'll want to save the number of epochs as well as the optimizer state, `optimizer.state_dict`. You'll likely want to use this trained model in the next part of the project, so best to save it now.
```
# TODO: Save the checkpoint
model.class_to_idx = train_data.class_to_idx
checkpoint = {'class_to_idx': model.class_to_idx,
'classifier': model.classifier,
'arch': 'densenet201',
'state_dict': model.state_dict(),
'optimizer': optimizer}
torch.save(checkpoint, 'checkpoint.pth')
```
## Loading the checkpoint
At this point it's good to write a function that can load a checkpoint and rebuild the model. That way you can come back to this project and keep working on it without having to retrain the network.
```
# TODO: Write a function that loads a checkpoint and rebuilds the model
def load_checkpoint(filepath):
checkpoint = torch.load(filepath)
# Load pre-trained network
model = getattr(torchvision.models, checkpoint['arch'])(pretrained=True)
# Freeze params of pre-trained model
for param in model.parameters():
param.requires_grad = False
model.classifier = checkpoint['classifier']
model.load_state_dict(checkpoint['state_dict'])
model.optimizer = checkpoint['optimizer']
model.class_to_idx = checkpoint['class_to_idx']
return model
model = load_checkpoint('checkpoint.pth')
model
```
# Inference for classification
Now you'll write a function to use a trained network for inference. That is, you'll pass an image into the network and predict the class of the flower in the image. Write a function called `predict` that takes an image and a model, then returns the top $K$ most likely classes along with the probabilities. It should look like
```python
probs, classes = predict(image_path, model)
print(probs)
print(classes)
> [ 0.01558163 0.01541934 0.01452626 0.01443549 0.01407339]
> ['70', '3', '45', '62', '55']
```
First you'll need to handle processing the input image such that it can be used in your network.
## Image Preprocessing
You'll want to use `PIL` to load the image ([documentation](https://pillow.readthedocs.io/en/latest/reference/Image.html)). It's best to write a function that preprocesses the image so it can be used as input for the model. This function should process the images in the same manner used for training.
First, resize the images where the shortest side is 256 pixels, keeping the aspect ratio. This can be done with the [`thumbnail`](http://pillow.readthedocs.io/en/3.1.x/reference/Image.html#PIL.Image.Image.thumbnail) or [`resize`](http://pillow.readthedocs.io/en/3.1.x/reference/Image.html#PIL.Image.Image.thumbnail) methods. Then you'll need to crop out the center 224x224 portion of the image.
Color channels of images are typically encoded as integers 0-255, but the model expected floats 0-1. You'll need to convert the values. It's easiest with a Numpy array, which you can get from a PIL image like so `np_image = np.array(pil_image)`.
As before, the network expects the images to be normalized in a specific way. For the means, it's `[0.485, 0.456, 0.406]` and for the standard deviations `[0.229, 0.224, 0.225]`. You'll want to subtract the means from each color channel, then divide by the standard deviation.
And finally, PyTorch expects the color channel to be the first dimension but it's the third dimension in the PIL image and Numpy array. You can reorder dimensions using [`ndarray.transpose`](https://docs.scipy.org/doc/numpy-1.13.0/reference/generated/numpy.ndarray.transpose.html). The color channel needs to be first and retain the order of the other two dimensions.
```
def process_image(image):
''' Scales, crops, and normalizes a PIL image for a PyTorch model,
returns an Numpy array
'''
# TODO: Process a PIL image for use in a PyTorch model
width, height = image.size
size = 256
height = int(size) if height > width else int(max(height * size/width, 1))
width = int(size) if height < width else int(max(height * size/width, 1))
resized_image = image.resize((width, height))
crop_size = 224
crop_width, crop_height = resized_image.size
x1 = (crop_width - crop_size) / 2
x2 = (crop_height - crop_size) / 2
x3 = x1 + crop_size
x4 = x2 + crop_size
crop_image = resized_image.crop((x1, x2, x3, x4))
# Convert color channel values
np_image = np.array(crop_image) / 255
mean = np.array([0.485, 0.456, 0.406])
std = np.array([0.229, 0.224, 0.225])
np_image = (np_image - mean) / std
np_image = np_image.transpose((2, 0, 1))
return np_image
```
To check your work, the function below converts a PyTorch tensor and displays it in the notebook. If your `process_image` function works, running the output through this function should return the original image (except for the cropped out portions).
```
def imshow(image, ax=None, title=None):
if ax is None:
fig, ax = plt.subplots()
# PyTorch tensors assume the color channel is the first dimension
# but matplotlib assumes is the third dimension
image = image.transpose((1, 2, 0))
# Undo preprocessing
mean = np.array([0.485, 0.456, 0.406])
std = np.array([0.229, 0.224, 0.225])
image = std * image + mean
# Image needs to be clipped between 0 and 1 or it looks like noise when displayed
image = np.clip(image, 0, 1)
ax.imshow(image)
return ax
# Test function works correctly
path = 'flowers/test/15/image_06351.jpg'
test_image = Image.open(path)
imshow(process_image(test_image));
```
## Class Prediction
Once you can get images in the correct format, it's time to write a function for making predictions with your model. A common practice is to predict the top 5 or so (usually called top-$K$) most probable classes. You'll want to calculate the class probabilities then find the $K$ largest values.
To get the top $K$ largest values in a tensor use [`x.topk(k)`](http://pytorch.org/docs/master/torch.html#torch.topk). This method returns both the highest `k` probabilities and the indices of those probabilities corresponding to the classes. You need to convert from these indices to the actual class labels using `class_to_idx` which hopefully you added to the model or from an `ImageFolder` you used to load the data ([see here](#Save-the-checkpoint)). Make sure to invert the dictionary so you get a mapping from index to class as well.
Again, this method should take a path to an image and a model checkpoint, then return the probabilities and classes.
```python
probs, classes = predict(image_path, model)
print(probs)
print(classes)
> [ 0.01558163 0.01541934 0.01452626 0.01443549 0.01407339]
> ['70', '3', '45', '62', '55']
```
```
def predict(labels, image_path, model, topk=5):
''' Predict the class (or classes) of an image using a trained deep learning model.
'''
# TODO: Implement the code to predict the class from an image file
# Put model in eval mode
model.eval()
model.to(device)
# Process image
img = Image.open(image_path)
img = process_image(img)
# Convert 2D image to 1D vector
img = np.expand_dims(img, 0)
img = torch.from_numpy(img)
inputs = img.to(device)
with torch.no_grad():
output = model.forward(inputs.float())
ps = torch.exp(output)
probs, classes = torch.topk(ps, topk)
index_to_class = {model.class_to_idx[x]: x for x in model.class_to_idx}
top_classes = [index_to_class[each] for each in classes.cpu().numpy()[0]]
flower_names = [labels.get(str(each)) for each in top_classes]
return probs.cpu().detach().numpy()[0], top_classes, flower_names
probs, classes, flower_names = predict(cat_to_name, path, model)
print(probs)
print(classes)
print(flower_names)
```
## Sanity Checking
Now that you can use a trained model for predictions, check to make sure it makes sense. Even if the testing accuracy is high, it's always good to check that there aren't obvious bugs. Use `matplotlib` to plot the probabilities for the top 5 classes as a bar graph, along with the input image. It should look like this:
<img src='assets/inference_example.png' width=300px>
You can convert from the class integer encoding to actual flower names with the `cat_to_name.json` file (should have been loaded earlier in the notebook). To show a PyTorch tensor as an image, use the `imshow` function defined above.
```
# TODO: Display an image along with the top 5 classes
def sanity_check(image, ps, classes):
num_class = len(ps)
ps = np.array(ps)
image = process_image(image)
image = image.transpose((1, 2, 0))
# Undo preprocessing
mean = np.array([0.485, 0.456, 0.406])
std = np.array([0.229, 0.224, 0.225])
image = std * image + mean
# Image needs to be clipped between 0 and 1 or it looks like noise when displayed
image = np.clip(image, 0, 1)
topk_list = np.arange(num_class)
topk_list = topk_list[::-1]
fig, (ax1, ax2) = plt.subplots(figsize=(10,6), ncols=2)
ax1.imshow(image)
ax1.axis('off')
ax1.set_title(classes[0])
ax2.barh(topk_list, ps)
ax2.set_yticks(topk_list)
ax2.set_yticklabels(topk_list.astype(int), size='large')
ax2.set_title('Class Probability')
ax2.set_xlim(0, 1.1)
ax2.set_yticklabels(classes)
fig.subplots_adjust(wspace=.8);
sanity_check(test_image, probs, flower_names)
```
| github_jupyter |
# Analyzing the Pathway Commons 2 (PC2) database SIF file
## CS446/546 class session 1
### Goal: count the number of different types of biological interactions in PC2
### Approach: retrieve compressed tab-delimited "edge-list" file and tabulate "interaction" column
### Information you will need:
- The URL is: http://www.pathwaycommons.org/archives/PC2/v9/PathwayCommons9.All.hgnc.sif.gz
- You'll be using the Python modules `gzip`, `timeit`, `pandas`, `urllib.request`, `collections` and `operator`
### What is the ".sif" file format?
SIF stands for Simple Interaction File. The format is like this:
```
A1BG controls-expression-of A2M
A1BG interacts-with ABCC6
A1BG interacts-with ACE2
A1BG interacts-with ADAM10
A1BG interacts-with ADAM17
A1BG interacts-with ADAM9
...
```
### Other stuff you should do:
- Print the first six lines of the uncompressed data file
- Use a timer to time how long your program takes
- Count how many rows there are in the data file
- Estimate the number of proteins in the database; we'll define them operationally as strings in column 1 or column 3, for which the content of column 2 is one of these interactions: 'interacts-with', 'in-complex-with', 'neighbor-of'
- Count the total number of unique pairs of interacting molecules (ignoring interaction type)
- Count the number rows for each type of interaction in the database
- R aficionados: can you do it using cURL and gunzip, or download.file?
### Step-by-step instructions for R:
- Open a file object representing a stream of the remote, compressed data file, using `urlopen`
- Open a file object representing a stream of the uncompressed data file, using `gzip.GzipFile`
- Start the timer
- Read one line at a time, until the end of the file
- Split line on "\t" and pull out the tuple of species1, interaction_type, species2 from the line of text
```
# Class Session 2
# this is the URL of the SIF file at Pathway Commons
sif_file_url <- "http://www.pathwaycommons.org/archives/PC2/v9/PathwayCommons9.All.hgnc.sif.gz"
# for starters, we only want three possible interaction types
interaction_types_ppi <- c("interacts-with","in-complex-with","neighbor-of")
```
Read the remote compressed file into a data frame using `readr::read_delim_chunked`; time it using `system.time`; use `head` to print the first six lines.
```
library(readr)
# do the filtering while reading the data; reduce disk space and memory usage
system.time(interactions_df <- read_delim_chunked(sif_file_url,
callback=DataFrameCallback$new(function(df_chunk, pos){
subset(df_chunk, interaction_type %in% interaction_types_ppi)}),
chunk_size=10000,
delim="\t",
quote="",
comment="",
col_names=c("species1","interaction_type","species2"),
progress=FALSE))
# sanity check the resulting data frame
head(interactions_df)
```
Count the number of rows that correspond to protein-protein interactions
```
# how many rows are there in the protein-protein interaction edge-list?
nrow(interactions_df)
```
Count the number of proteins that participate in protein-protein interactions, using `unique`
```
# how many unique proteins are there in the interaction network?
length(unique(c(interactions_df$species1, interactions_df$species2)))
```
Count the number of unique interacting protein-protein pairs, regardless of interaction type
```
# how many unique interacting protein pairs are there in the interaction network?
length(unique(apply(interactions_df[,c(1,3)], 1, function(my_pair) {
paste(c(min(my_pair), "-", max(my_pair)),collapse="")
})))
```
Use `table` to count the number of each type of interaction in the database
```
# really quick-and-dirty approach using cURL and gunzip
system.time({
system(paste(c("curl -s --compressed ", sif_file_url, " | gunzip > pc.sif"), collapse=""),
intern=TRUE)
my_df <- read.table(file="pc.sif",
sep="\t",
comment.char="",
quote="",
col.names=c("species1",
"interaction_type",
"species2"),
stringsAsFactors=FALSE)
interactions_df5 <- subset(my_df,
interaction_type %in% interaction_types_ppi)})
all(interactions_df5 == interactions_df)
print(sort(table(as.factor(my_df$interaction_type)), decreasing=TRUE))
# alternative approach which is kind of slow and (transiently) memory-hungry; read the entire file into memory and
# then process the text contents of the file, line by line
system.time({
alltext <- readLines(gzcon(url(sif_file_url)))
nlines <- length(alltext)
interactions_df4 <- data.frame(do.call(rbind, lapply(1:length(alltext),
function(i) {
split_line <- strsplit(alltext[i], "\t")[[1]]
names(split_line) <- c("species1","interaction_type","species2")
split_line
})))
interactions_df4 <- subset(interactions_df4, interaction_type %in% interaction_types_ppi)
})
# sanity check that this slow approach at least gave us correct results
all(interactions_df4 == interactions_df)
rm(interactions_df4)
rm(alltext)
# quick and dirty approach which doesn't require looking up any
# newfangled commands, but transiently eats up disk space and memory
fname <- tempfile()
gzfname <- paste(c(fname, ".gz"),collapse="")
system.time({
download.file(sif_file_url, destfile=gzfname)
interactions_df3 <- subset(read.table(file=gzfile(gzfname),
sep="\t",
comment.char="",
quote="",
col.names=c("species1","interaction_type", "species2"),
stringsAsFactors=FALSE),
interaction_type %in% interaction_types_ppi)})
unlink(gzfname)
# sanity check the results
all(interactions_df3 == interactions_df)
# clean up
rm(interactions_df3)
```
| github_jupyter |
##### Copyright 2019 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Word embeddings
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/tutorials/text/word_embeddings">
<img src="https://www.tensorflow.org/images/tf_logo_32px.png" />
View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/tutorials/text/word_embeddings.ipynb">
<img src="https://www.tensorflow.org/images/colab_logo_32px.png" />
Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/tutorials/text/word_embeddings.ipynb">
<img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />
View source on GitHub</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/tutorials/text/word_embeddings.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
</td>
</table>
This tutorial introduces word embeddings. It contains complete code to train word embeddings from scratch on a small dataset, and to visualize these embeddings using the [Embedding Projector](http://projector.tensorflow.org) (shown in the image below).
<img src="https://github.com/tensorflow/docs/blob/master/site/en/tutorials/text/images/embedding.jpg?raw=1" alt="Screenshot of the embedding projector" width="400"/>
## Representing text as numbers
Machine learning models take vectors (arrays of numbers) as input. When working with text, the first thing we must do come up with a strategy to convert strings to numbers (or to "vectorize" the text) before feeding it to the model. In this section, we will look at three strategies for doing so.
### One-hot encodings
As a first idea, we might "one-hot" encode each word in our vocabulary. Consider the sentence "The cat sat on the mat". The vocabulary (or unique words) in this sentence is (cat, mat, on, sat, the). To represent each word, we will create a zero vector with length equal to the vocabulary, then place a one in the index that corresponds to the word. This approach is shown in the following diagram.
<img src="https://github.com/tensorflow/docs/blob/master/site/en/tutorials/text/images/one-hot.png?raw=1" alt="Diagram of one-hot encodings" width="400" />
To create a vector that contains the encoding of the sentence, we could then concatenate the one-hot vectors for each word.
Key point: This approach is inefficient. A one-hot encoded vector is sparse (meaning, most indicices are zero). Imagine we have 10,000 words in the vocabulary. To one-hot encode each word, we would create a vector where 99.99% of the elements are zero.
### Encode each word with a unique number
A second approach we might try is to encode each word using a unique number. Continuing the example above, we could assign 1 to "cat", 2 to "mat", and so on. We could then encode the sentence "The cat sat on the mat" as a dense vector like [5, 1, 4, 3, 5, 2]. This appoach is efficient. Instead of a sparse vector, we now have a dense one (where all elements are full).
There are two downsides to this approach, however:
* The integer-encoding is arbitrary (it does not capture any relationship between words).
* An integer-encoding can be challenging for a model to interpret. A linear classifier, for example, learns a single weight for each feature. Because there is no relationship between the similarity of any two words and the similarity of their encodings, this feature-weight combination is not meaningful.
### Word embeddings
Word embeddings give us a way to use an efficient, dense representation in which similar words have a similar encoding. Importantly, we do not have to specify this encoding by hand. An embedding is a dense vector of floating point values (the length of the vector is a parameter you specify). Instead of specifying the values for the embedding manually, they are trainable parameters (weights learned by the model during training, in the same way a model learns weights for a dense layer). It is common to see word embeddings that are 8-dimensional (for small datasets), up to 1024-dimensions when working with large datasets. A higher dimensional embedding can capture fine-grained relationships between words, but takes more data to learn.
<img src="https://github.com/tensorflow/docs/blob/master/site/en/tutorials/text/images/embedding2.png?raw=1" alt="Diagram of an embedding" width="400"/>
Above is a diagram for a word embedding. Each word is represented as a 4-dimensional vector of floating point values. Another way to think of an embedding is as "lookup table". After these weights have been learned, we can encode each word by looking up the dense vector it corresponds to in the table.
## Setup
```
from __future__ import absolute_import, division, print_function, unicode_literals
try:
# %tensorflow_version only exists in Colab.
%tensorflow_version 2.x
except Exception:
pass
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers
import tensorflow_datasets as tfds
tfds.disable_progress_bar()
```
## Using the Embedding layer
Keras makes it easy to use word embeddings. Let's take a look at the [Embedding](https://www.tensorflow.org/api_docs/python/tf/keras/layers/Embedding) layer.
The Embedding layer can be understood as a lookup table that maps from integer indices (which stand for specific words) to dense vectors (their embeddings). The dimensionality (or width) of the embedding is a parameter you can experiment with to see what works well for your problem, much in the same way you would experiment with the number of neurons in a Dense layer.
```
embedding_layer = layers.Embedding(1000, 5)
```
When you create an Embedding layer, the weights for the embedding are randomly initialized (just like any other layer). During training, they are gradually adjusted via backpropagation. Once trained, the learned word embeddings will roughly encode similarities between words (as they were learned for the specific problem your model is trained on).
If you pass an integer to an embedding layer, the result replaces each integer with the vector from the embedding table:
```
result = embedding_layer(tf.constant([1,2,3]))
result.numpy()
```
For text or sequence problems, the Embedding layer takes a 2D tensor of integers, of shape `(samples, sequence_length)`, where each entry is a sequence of integers. It can embed sequences of variable lengths. You could feed into the embedding layer above batches with shapes `(32, 10)` (batch of 32 sequences of length 10) or `(64, 15)` (batch of 64 sequences of length 15).
The returned tensor has one more axis than the input, the embedding vectors are aligned along the new last axis. Pass it a `(2, 3)` input batch and the output is `(2, 3, N)`
```
result = embedding_layer(tf.constant([[0,1,2],[3,4,5]]))
result.shape
```
When given a batch of sequences as input, an embedding layer returns a 3D floating point tensor, of shape `(samples, sequence_length, embedding_dimensionality)`. To convert from this sequence of variable length to a fixed representation there are a variety of standard approaches. You could use an RNN, Attention, or pooling layer before passing it to a Dense layer. This tutorial uses pooling because it's simplest. The [Text Classification with an RNN](text_classification_rnn.ipynb) tutorial is a good next step.
## Learning embeddings from scratch
In this tutorial you will train a sentiment classifier on IMDB movie reviews. In the process, the model will learn embeddings from scratch. We will use to a preprocessed dataset.
To load a text dataset from scratch see the [Loading text tutorial](../load_data/text.ipynb).
```
(train_data, test_data), info = tfds.load(
'imdb_reviews/subwords8k',
split = (tfds.Split.TRAIN, tfds.Split.TEST),
with_info=True, as_supervised=True)
```
Get the encoder (`tfds.features.text.SubwordTextEncoder`), and have a quick look at the vocabulary.
The "\_" in the vocabulary represent spaces. Note how the vocabulary includes whole words (ending with "\_") and partial words which it can use to build larger words:
```
encoder = info.features['text'].encoder
encoder.subwords[:20]
```
Movie reviews can be different lengths. We will use the `padded_batch` method to standardize the lengths of the reviews.
```
padded_shapes = ([None],())
train_batches = train_data.shuffle(1000).padded_batch(10, padded_shapes = padded_shapes)
test_batches = test_data.shuffle(1000).padded_batch(10, padded_shapes = padded_shapes)
```
As imported, the text of reviews is integer-encoded (each integer represents a specific word or word-part in the vocabulary).
Note the trailing zeros, because the batch is padded to the longest example.
```
train_batch, train_labels = next(iter(train_batches))
train_batch.numpy()
```
### Create a simple model
We will use the [Keras Sequential API](../../guide/keras) to define our model. In this case it is a "Continuous bag of words" style model.
* Next the Embedding layer takes the integer-encoded vocabulary and looks up the embedding vector for each word-index. These vectors are learned as the model trains. The vectors add a dimension to the output array. The resulting dimensions are: `(batch, sequence, embedding)`.
* Next, a GlobalAveragePooling1D layer returns a fixed-length output vector for each example by averaging over the sequence dimension. This allows the model to handle input of variable length, in the simplest way possible.
* This fixed-length output vector is piped through a fully-connected (Dense) layer with 16 hidden units.
* The last layer is densely connected with a single output node. Using the sigmoid activation function, this value is a float between 0 and 1, representing a probability (or confidence level) that the review is positive.
Caution: This model doesn't use masking, so the zero-padding is used as part of the input, so the padding length may affect the output. To fix this, see the [masking and padding guide](../../guide/keras/masking_and_padding).
```
embedding_dim=16
model = keras.Sequential([
layers.Embedding(encoder.vocab_size, embedding_dim),
layers.GlobalAveragePooling1D(),
layers.Dense(16, activation='relu'),
layers.Dense(1, activation='sigmoid')
])
model.summary()
```
### Compile and train the model
```
model.compile(optimizer='adam',
loss='binary_crossentropy',
metrics=['accuracy'])
history = model.fit(
train_batches,
epochs=10,
validation_data=test_batches, validation_steps=20)
```
With this approach our model reaches a validation accuracy of around 88% (note the model is overfitting, training accuracy is significantly higher).
```
import matplotlib.pyplot as plt
history_dict = history.history
acc = history_dict['accuracy']
val_acc = history_dict['val_accuracy']
loss = history_dict['loss']
val_loss = history_dict['val_loss']
epochs = range(1, len(acc) + 1)
plt.figure(figsize=(12,9))
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
plt.figure(figsize=(12,9))
plt.plot(epochs, acc, 'bo', label='Training acc')
plt.plot(epochs, val_acc, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.xlabel('Epochs')
plt.ylabel('Accuracy')
plt.legend(loc='lower right')
plt.ylim((0.5,1))
plt.show()
```
## Retrieve the learned embeddings
Next, let's retrieve the word embeddings learned during training. This will be a matrix of shape `(vocab_size, embedding-dimension)`.
```
e = model.layers[0]
weights = e.get_weights()[0]
print(weights.shape) # shape: (vocab_size, embedding_dim)
```
We will now write the weights to disk. To use the [Embedding Projector](http://projector.tensorflow.org), we will upload two files in tab separated format: a file of vectors (containing the embedding), and a file of meta data (containing the words).
```
import io
encoder = info.features['text'].encoder
out_v = io.open('vecs.tsv', 'w', encoding='utf-8')
out_m = io.open('meta.tsv', 'w', encoding='utf-8')
for num, word in enumerate(encoder.subwords):
vec = weights[num+1] # skip 0, it's padding.
out_m.write(word + "\n")
out_v.write('\t'.join([str(x) for x in vec]) + "\n")
out_v.close()
out_m.close()
```
If you are running this tutorial in [Colaboratory](https://colab.research.google.com), you can use the following snippet to download these files to your local machine (or use the file browser, *View -> Table of contents -> File browser*).
```
try:
from google.colab import files
except ImportError:
pass
else:
files.download('vecs.tsv')
files.download('meta.tsv')
```
## Visualize the embeddings
To visualize our embeddings we will upload them to the embedding projector.
Open the [Embedding Projector](http://projector.tensorflow.org/) (this can also run in a local TensorBoard instance).
* Click on "Load data".
* Upload the two files we created above: `vecs.tsv` and `meta.tsv`.
The embeddings you have trained will now be displayed. You can search for words to find their closest neighbors. For example, try searching for "beautiful". You may see neighbors like "wonderful".
Note: your results may be a bit different, depending on how weights were randomly initialized before training the embedding layer.
Note: experimentally, you may be able to produce more interpretable embeddings by using a simpler model. Try deleting the `Dense(16)` layer, retraining the model, and visualizing the embeddings again.
<img src="https://github.com/tensorflow/docs/blob/master/site/en/tutorials/text/images/embedding.jpg?raw=1" alt="Screenshot of the embedding projector" width="400"/>
## Next steps
This tutorial has shown you how to train and visualize word embeddings from scratch on a small dataset.
* To learn about recurrent networks see the [Keras RNN Guide](../../guide/keras/rnn.ipynb).
* To learn more about text classification (including the overall workflow, and if you're curious about when to use embeddings vs one-hot encodings) we recommend this practical text classification [guide](https://developers.google.com/machine-learning/guides/text-classification/step-2-5).
| github_jupyter |
<!-- dom:TITLE: Data Analysis and Machine Learning: Introduction and Representing data -->
# Data Analysis and Machine Learning: Introduction and Representing data
<!-- dom:AUTHOR: Morten Hjorth-Jensen at Department of Physics, University of Oslo & Department of Physics and Astronomy and National Superconducting Cyclotron Laboratory, Michigan State University -->
<!-- Author: -->
**Morten Hjorth-Jensen**, Department of Physics, University of Oslo and Department of Physics and Astronomy and National Superconducting Cyclotron Laboratory, Michigan State University
Date: **Dec 6, 2017**
Copyright 1999-2017, Morten Hjorth-Jensen. Released under CC Attribution-NonCommercial 4.0 license
## What is Machine Learning?
Machine learning is the science of giving computers the ability to
learn without being explicitly programmed. The idea is that there
exist generic algorithms which can be used to find patterns in a broad
class of data sets without having to write code specifically for each
problem. The algorithm will build its own logic based on the data.
Machine learning is a subfield of computer science, and is closely
related to computational statistics. It evolved from the study of
pattern recognition in artificial intelligence (AI) research, and has
made contributions to AI tasks like computer vision, natural language
processing and speech recognition. It has also, especially in later
years, found applications in a wide variety of other areas, including
bioinformatics, economy, physics, finance and marketing.
## Types of Machine Learning
The approaches to machine learning are many, but are often split into two main categories.
In *supervised learning* we know the answer to a problem,
and let the computer deduce the logic behind it. On the other hand, *unsupervised learning*
is a method for finding patterns and relationship in data sets without any prior knowledge of the system.
Some authours also operate with a third category, namely *reinforcement learning*. This is a paradigm
of learning inspired by behavioural psychology, where learning is achieved by trial-and-error,
solely from rewards and punishment.
Another way to categorize machine learning tasks is to consider the desired output of a system.
Some of the most common tasks are:
* Classification: Outputs are divided into two or more classes. The goal is to produce a model that assigns inputs into one of these classes. An example is to identify digits based on pictures of hand-written ones. Classification is typically supervised learning.
* Regression: Finding a functional relationship between an input data set and a reference data set. The goal is to construct a function that maps input data to continuous output values.
* Clustering: Data are divided into groups with certain common traits, without knowing the different groups beforehand. It is thus a form of unsupervised learning.
## Different algorithms
In this course we will build our machine learning approach on a statistical foundation, with elements
from data analysis, stochastic processes etc before we proceed with the following machine learning algorithms
1. Linear regression and its variants
2. Decision tree algorithms, from simpler to more complex ones
3. Nearest neighbors models
4. Bayesian statistics
5. Support vector machines and finally various variants of
6. Artifical neural networks
Before we proceed however, there are several practicalities with data analysis and software tools we would
like to present. These tools will help us in our understanding of various machine learning algorithms.
Our emphasis here is on understanding the mathematical aspects of different algorithms, however, where possible
we will emphasize the importance of using available software.
## Software and needed installations
We will make intensive use of python as programming language and the myriad of available libraries.
Furthermore, you will find IPython/Jupyter notebooks invaluable in your work.
You can run **R** codes in the Jupyter/IPython notebooks, with the immediate benefit of visualizing your data.
If you have Python installed (we recommend Python3) and you feel pretty familiar with installing different packages,
we recommend that you install the following Python packages via **pip** as
1. pip install numpy scipy matplotlib ipython scikit-learn mglearn sympy pandas pillow
For Python3, replace **pip** with **pip3**.
For OSX users we recommend also, after having installed Xcode, to install **brew**. Brew allows
for a seamless installation of additional software via for example
1. brew install python3
For Linux users, with its variety of distributions like for example the widely popular Ubuntu distribution
you can use **pip** as well and simply install Python as
1. sudo apt-get install python3 (or python for pyhton2.7)
etc etc.
## Python installers
If you don't want to perform these operations separately, we recommend two widely used distrubutions which set up
all relevant dependencies for Python, namely
1. [Anaconda](https://docs.anaconda.com/) Anaconda is an open source distribution of the Python and R programming languages for large-scale data processing, predictive analytics, and scientific computing, that aims to simplify package management and deployment. Package versions are managed by the package management system **conda**
2. [Enthought canopy](https://www.enthought.com/product/canopy/) is a Python distribution for scientific and analytic computing distribution and analysis environment, available for free and under a commercial license.
## Installing R, C++, cython or Julia
You will also find it convenient to utilize R.
Jupyter/Ipython notebook allows you run **R** code interactively in your browser. The software library **R** is
tuned to statistically analysis and allows for an easy usage of the tools we will discuss in these texts.
To install **R** with Jupyter notebook [following the link here](https://mpacer.org/maths/r-kernel-for-ipython-notebook)
## Installing R, C++, cython or Julia
For the C++ affecianodas, Jupyter/IPython notebook allows you also to install C++ and run codes written in this language
interactively in the browser. Since we will emphasize writing many of the algorithms yourself, you can thus opt for
either Python or C++ as programming languages.
To add more entropy, **cython** can also be used when running your notebooks. It means that Python with the Jupyter/IPython notebook
setup allows you to integrate widely popular softwares and tools for scientific computing. With its versatility,
including symbolic operations, Python offers a unique computational environment. Your Jupyter/IPython notebook
can easily be converted into a nicely rendered **PDF** file or a Latex file for further processing.
This never ends.
If you use the light mark-up language **doconce** you can convert a standard ascii text file into various HTML
formats, ipython notebooks, latex files, pdf files etc.
## Useful packages
If you already have a Python installation set up, you can use **pip** or **pip3** to install
1. pip3 install numpy scipy ipython
2. pip3 install pandas matplotlib scikit-learn pillow
Another useful package is **mglearn**.
## Introduction to Jupyter notebook and available tools
```
%matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
from scipy import sparse
import pandas as pd
from IPython.display import display
eye = np.eye(4)
print(eye)
sparse_mtx = sparse.csr_matrix(eye)
print(sparse_mtx)
x = np.linspace(-10,10,100)
y = np.sin(x)
plt.plot(x,y,marker='x')
plt.show()
data = {'Name': ["John", "Anna", "Peter", "Linda"], 'Location': ["Nairobi", "Napoli", "London", "Buenos Aires"], 'Age':[51, 21, 34, 45]}
data_pandas = pd.DataFrame(data)
display(data_pandas)
```
## Representing data, more examples
```
import numpy as np
import matplotlib.pyplot as plt
from scipy import sparse
import pandas as pd
from IPython.display import display
import mglearn
import sklearn
from sklearn.linear_model import LinearRegression
from sklearn.tree import DecisionTreeRegressor
x, y = mglearn.datasets.make_wave(n_samples=100)
line = np.linspace(-3,3,1000,endpoint=False).reshape(-1,1)
reg = DecisionTreeRegressor(min_samples_split=3).fit(x,y)
plt.plot(line, reg.predict(line), label="decision tree")
regline = LinearRegression().fit(x,y)
plt.plot(line, regline.predict(line), label= "Linear Rgression")
plt.show()
```
## Predator-Prey model from ecology
The population dynamics of a simple predator-prey system is a
classical example shown in many biology textbooks when ecological
systems are discussed. The system contains all elements of the
scientific method:
* The set up of a specific hypothesis combined with
* the experimental methods needed (one can study existing data or perform experiments)
* analyzing and interpreting the data and performing further experiments if needed
* trying to extract general behaviors and extract eventual laws or patterns
* develop mathematical relations for the uncovered regularities/laws and test these by per forming new experiments
## Case study from Hudson bay
Lots of data about populations of hares and lynx collected from furs in Hudson Bay, Canada, are available. It is known that the populations oscillate. Why?
Here we start by
1. plotting the data
2. derive a simple model for the population dynamics
3. (fitting parameters in the model to the data)
4. using the model predict the evolution other predator-pray systems
## Hudson bay data
% if FORMAT == 'ipynb':
Most mammalian predators rely on a variety of prey, which complicates mathematical modeling; however, a few predators have become highly specialized and seek almost exclusively a single prey species. An example of this simplified predator-prey interaction is seen in Canadian northern forests, where the populations of the lynx and the snowshoe hare are intertwined in a life and death struggle.
One reason that this particular system has been so extensively studied is that the Hudson Bay company kept careful records of all furs from the early 1800s into the 1900s. The records for the furs collected by the Hudson Bay company showed distinct oscillations (approximately 12 year periods), suggesting that these species caused almost periodic fluctuations of each other's populations. The table here shows data from 1900 to 1920.
% endif
<table border="1">
<thead>
<tr><th align="center">Year</th> <th align="center">Hares (x1000)</th> <th align="center">Lynx (x1000)</th> </tr>
</thead>
<tbody>
<tr><td align="left"> 1900 </td> <td align="right"> 30.0 </td> <td align="right"> 4.0 </td> </tr>
<tr><td align="left"> 1901 </td> <td align="right"> 47.2 </td> <td align="right"> 6.1 </td> </tr>
<tr><td align="left"> 1902 </td> <td align="right"> 70.2 </td> <td align="right"> 9.8 </td> </tr>
<tr><td align="left"> 1903 </td> <td align="right"> 77.4 </td> <td align="right"> 35.2 </td> </tr>
<tr><td align="left"> 1904 </td> <td align="right"> 36.3 </td> <td align="right"> 59.4 </td> </tr>
<tr><td align="left"> 1905 </td> <td align="right"> 20.6 </td> <td align="right"> 41.7 </td> </tr>
<tr><td align="left"> 1906 </td> <td align="right"> 18.1 </td> <td align="right"> 19.0 </td> </tr>
<tr><td align="left"> 1907 </td> <td align="right"> 21.4 </td> <td align="right"> 13.0 </td> </tr>
<tr><td align="left"> 1908 </td> <td align="right"> 22.0 </td> <td align="right"> 8.3 </td> </tr>
<tr><td align="left"> 1909 </td> <td align="right"> 25.4 </td> <td align="right"> 9.1 </td> </tr>
<tr><td align="left"> 1910 </td> <td align="right"> 27.1 </td> <td align="right"> 7.4 </td> </tr>
<tr><td align="left"> 1911 </td> <td align="right"> 40.3 </td> <td align="right"> 8.0 </td> </tr>
<tr><td align="left"> 1912 </td> <td align="right"> 57 </td> <td align="right"> 12.3 </td> </tr>
<tr><td align="left"> 1913 </td> <td align="right"> 76.6 </td> <td align="right"> 19.5 </td> </tr>
<tr><td align="left"> 1914 </td> <td align="right"> 52.3 </td> <td align="right"> 45.7 </td> </tr>
<tr><td align="left"> 1915 </td> <td align="right"> 19.5 </td> <td align="right"> 51.1 </td> </tr>
<tr><td align="left"> 1916 </td> <td align="right"> 11.2 </td> <td align="right"> 29.7 </td> </tr>
<tr><td align="left"> 1917 </td> <td align="right"> 7.6 </td> <td align="right"> 15.8 </td> </tr>
<tr><td align="left"> 1918 </td> <td align="right"> 14.6 </td> <td align="right"> 9.7 </td> </tr>
<tr><td align="left"> 1919 </td> <td align="right"> 16.2 </td> <td align="right"> 10.1 </td> </tr>
<tr><td align="left"> 1920 </td> <td align="right"> 24.7 </td> <td align="right"> 8.6 </td> </tr>
</tbody>
</table>
## Plotting the data
```
import numpy as np
from matplotlib import pyplot as plt
# Load in data file
data = np.loadtxt('Hudson_Bay.dat', delimiter=',', skiprows=1)
# Make arrays containing x-axis and hares and lynx populations
year = data[:,0]
hares = data[:,1]
lynx = data[:,2]
plt.plot(year, hares ,'b-+', year, lynx, 'r-o')
plt.axis([1900,1920,0, 100.0])
plt.xlabel(r'Year')
plt.ylabel(r'Numbers of hares and lynx ')
plt.legend(('Hares','Lynx'), loc='upper right')
plt.title(r'Population of hares and lynx from 1900-1920 (x1000)}')
plt.savefig('Hudson_Bay_data.pdf')
plt.savefig('Hudson_Bay_data.png')
plt.show()
```
% if FORMAT != 'ipynb':
## Hares and lynx in Hudson bay from 1900 to 1920
<!-- dom:FIGURE: [fig/Hudson_Bay_data.png, width=700 frac=0.9] -->
<!-- begin figure -->
<p></p>
<img src="fig/Hudson_Bay_data.png" width=700>
<!-- end figure -->
% endif
## Why now create a computer model for the hare and lynx populations?
% if FORMAT == 'ipynb':
We see from the plot that there are indeed fluctuations.
We would like to create a mathematical model that explains these
population fluctuations. Ecologists have predicted that in a simple
predator-prey system that a rise in prey population is followed (with
a lag) by a rise in the predator population. When the predator
population is sufficiently high, then the prey population begins
dropping. After the prey population falls, then the predator
population falls, which allows the prey population to recover and
complete one cycle of this interaction. Thus, we see that
qualitatively oscillations occur. Can a mathematical model predict
this? What causes cycles to slow or speed up? What affects the
amplitude of the oscillation or do you expect to see the oscillations
damp to a stable equilibrium? The models tend to ignore factors like
climate and other complicating factors. How significant are these?
% else:
* We see oscillations in the data
* What causes cycles to slow or speed up?
* What affects the amplitude of the oscillation or do you expect to see the oscillations damp to a stable equilibrium?
* With a model we can better *understand the data*
* More important: we can understand the ecology dynamics of
predator-pray populations
% endif
## The traditional (top-down) approach
The classical way (in all books) is to present the Lotka-Volterra equations:
$$
\begin{align*}
\frac{dH}{dt} &= H(a - b L)\\
\frac{dL}{dt} &= - L(d - c H)
\end{align*}
$$
Here,
* $H$ is the number of preys
* $L$ the number of predators
* $a$, $b$, $d$, $c$ are parameters
Most books quickly establish the model and then use considerable space on
discussing the qualitative properties of this *nonlinear system of
ODEs* (which cannot be solved)
## The "new" discrete bottom-up approach
**The bottom-up approach.**
* Start with experimental data and discuss the methods which have been used to collect the data, the assumptions, the electronic devices, the aims etc. That is, expose the students to the theory and assumptions behind the data that have been collected and motivate for the scientific method.
* Where appropriate the students should do the experiment(s) needed to collect the data.
* The first programming tasks are to read and visualize the data to see if there are patterns or regularities. This strengthens a research-driven intuition.
* Now we want to increase the understanding through modeling.
* Most of the biology lies in the *derivation* of the model. We shall
focus on an intuitive discrete approach that leads to difference
equations that can be programmed *and solved* directly.
## Basic (computer-friendly) mathematics notation
* Time points: $t_0,t_1,\ldots,t_m$
* Uniform distribution of time points: $t_n=n\Delta t$
* $H^n$: population of hares at time $t_n$
* $L^n$: population of lynx at time $t_n$
* We want to model the changes in populations, $\Delta H=H^{n+1}-H^n$
and $\Delta L=L^{n+1}-L^n$ during a general time interval $[t_{n+1},t_n]$
of length $\Delta t=t_{n+1}-t_n$
## Basic dynamics of the population of hares
The population of hares evolves due to births and deaths exactly as a bacteria population:
$$
\Delta H = a \Delta t H^n
$$
However, hares have an additional loss in the population because
they are eaten by lynx.
All the hares and lynx can form
$H\cdot L$ pairs in total. When such pairs meet during a time
interval $\Delta t$, there is some
small probablity that the lynx will eat the hare.
So in fraction $b\Delta t HL$, the lynx eat hares. This
loss of hares and must be accounted for:
subtracted in the equation for hares:
$$
\Delta H = a\Delta t H^n - b \Delta t H^nL^n
$$
## Basic dynamics of the population of lynx
We assume that the primary growth for the lynx population depends on sufficient food for raising lynx kittens, which implies an adequate source of nutrients from predation on hares. Thus, the growth of the lynx population does not only depend of how many lynx there are, but on how many hares they can eat.
In a time interval $\Delta t HL$ hares and lynx can meet, and in a
fraction $b\Delta t HL$ the lynx eats the hare. All of this does not
contribute to the growth of lynx, again just a fraction of
$b\Delta t HL$ that we write as
$d\Delta t HL$. In addition, lynx die just as in the population
dynamics with one isolated animal population, leading to a loss
$-c\Delta t L$.
The accounting of lynx then looks like
$$
\Delta L = d\Delta t H^nL^n - c\Delta t L^n
$$
## Evolution equations
By writing up the definition of $\Delta H$ and $\Delta L$, and putting
all assumed known terms $H^n$ and $L^n$ on the right-hand side, we have
$$
H^{n+1} = H^n + a\Delta t H^n - b\Delta t H^n L^n
$$
$$
L^{n+1} = L^n + d\Delta t H^nL^n - c\Delta t L^n
$$
Note:
* These equations are ready to be implemented!
* But to start, we need $H^0$ and $L^0$
(which we can get from the data)
* We also need values for $a$, $b$, $d$, $c$
## Adapt the model to the Hudson Bay case
* As always, models tend to be general - as here, applicable
to "all" predator-pray systems
* The critical issue is whether the *interaction* between hares and lynx
is sufficiently well modeled by $\hbox{const}HL$
* The parameters $a$, $b$, $d$, and $c$ must be
estimated from data
* Measure time in years
* $t_0=1900$, $t_m=1920$
## The program
```
import numpy as np
import matplotlib.pyplot as plt
def solver(m, H0, L0, dt, a, b, c, d, t0):
"""Solve the difference equations for H and L over m years
with time step dt (measured in years."""
num_intervals = int(m/float(dt))
t = np.linspace(t0, t0 + m, num_intervals+1)
H = np.zeros(t.size)
L = np.zeros(t.size)
print 'Init:', H0, L0, dt
H[0] = H0
L[0] = L0
for n in range(0, len(t)-1):
H[n+1] = H[n] + a*dt*H[n] - b*dt*H[n]*L[n]
L[n+1] = L[n] + d*dt*H[n]*L[n] - c*dt*L[n]
return H, L, t
# Load in data file
data = np.loadtxt('Hudson_Bay.csv', delimiter=',', skiprows=1)
# Make arrays containing x-axis and hares and lynx populations
t_e = data[:,0]
H_e = data[:,1]
L_e = data[:,2]
# Simulate using the model
H, L, t = solver(m=20, H0=34.91, L0=3.857, dt=0.1,
a=0.4807, b=0.02482, c=0.9272, d=0.02756,
t0=1900)
# Visualize simulations and data
plt.plot(t_e, H_e, 'b-+', t_e, L_e, 'r-o', t, H, 'm--', t, L, 'k--')
plt.xlabel('Year')
plt.ylabel('Numbers of hares and lynx')
plt.axis([1900, 1920, 0, 140])
plt.title(r'Population of hares and lynx 1900-1920 (x1000)')
plt.legend(('H_e', 'L_e', 'H', 'L'), loc='upper left')
plt.savefig('Hudson_Bay_sim.pdf')
plt.savefig('Hudson_Bay_sim.png')
plt.show()
```
% if FORMAT != 'ipynb':
## The plot
<!-- dom:FIGURE: [fig/Hudson_Bay_sim.png, width=700 frac=0.9] -->
<!-- begin figure -->
<p></p>
<img src="fig/Hudson_Bay_sim.png" width=700>
<!-- end figure -->
% else:
If we perform a least-square fitting, we can find optimal values for the parameters $a$, $b$, $d$, $c$. The optimal parameters are $a=0.4807$, $b=0.02482$, $d=0.9272$ and $c=0.02756$. These parameters result in a slightly modified initial conditions, namely $H(0) = 34.91$ and $L(0)=3.857$. With these parameters we are now ready to solve the equations and plot these data together with the experimental values.
% endif
## Linear Least squares in R
HudsonBay = read.csv("src/Hudson_Bay.csv",header=T)
fix(HudsonBay)
dim(HudsonBay)
names(HudsonBay)
plot(HudsonBay$Year, HudsonBay$Hares..x1000.)
attach(HudsonBay)
plot(Year, Hares..x1000.)
plot(Year, Hares..x1000., col="red", varwidth=T, xlab="Years", ylab="Haresx 1000")
summary(HudsonBay)
summary(Hares..x1000.)
library(MASS)
library(ISLR)
scatter.smooth(x=Year, y = Hares..x1000.)
linearMod = lm(Hares..x1000. ~ Year)
print(linearMod)
summary(linearMod)
plot(linearMod)
confint(linearMod)
predict(linearMod,data.frame(Year=c(1910,1914,1920)),interval="confidence")
## Linear Least squares in R
set.seed(1485)
len = 24
x = runif(len)
y = x^3+rnorm(len, 0,0.06)
ds = data.frame(x = x, y = y)
str(ds)
plot( y ~ x, main ="Known cubic with noise")
s = seq(0,1,length =100)
lines(s, s^3, lty =2, col ="green")
m = nls(y ~ I(x^power), data = ds, start = list(power=1), trace = T)
class(m)
summary(m)
power = round(summary(m)$coefficients[1], 3)
power.se = round(summary(m)$coefficients[2], 3)
plot(y ~ x, main = "Fitted power model", sub = "Blue: fit; green: known")
s = seq(0, 1, length = 100)
lines(s, s^3, lty = 2, col = "green")
lines(s, predict(m, list(x = s)), lty = 1, col = "blue")
text(0, 0.5, paste("y =x^ (", power, " +/- ", power.se, ")", sep = ""), pos = 4)
| github_jupyter |
```
#coding=utf-8
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import os
import time
import cPickle
def feature_rest(test_data, train_feature, ):
# train_feature = train_data.keys()
# test_feature = test_data.keys()
test_feature = test_data.keys()
result = test_data.copy()
for f2 in test_feature:
if f2 not in train_feature:
del result[f2]
for f1 in train_feature:
if f1 not in test_feature:
result[f1] = 0
result = result[train_feature]
return result
ageDic = {
'60后': 0,
'70后': 1,
'80后': 2,
'90后': 3,
'00后': 4,
}
genderDic = {
'男': 0,
'女': 1,
}
provinceDic1 = {
'北京': 0,
'浙江': 1,
'甘肃': 2,
'安徽': 3,
'贵州': 4,
'吉林': 5,
'四川': 6,
'江苏': 7,
'福建': 8,
'新疆': 9,
'上海': 10,
'海南': 11,
'天津': 12,
'重庆': 13,
'云南': 14,
'青海': 15,
'内蒙古': 16,
'湖北': 17,
'河北': 18,
'西藏': 19,
'黑龙江': 20,
'河南': 21,
'广西': 22,
'陕西': 23,
'山东': 24,
'广东': 25,
'湖南': 26,
'山西': 27,
'江西': 28,
'辽宁': 29,
'宁夏': 30,
}
provinceDic2 = {
'山东': 0,
'江苏': 0,
'安徽': 0,
'浙江': 0,
'福建': 0,
'上海': 0,
'广东': 1,
'广西': 1,
'海南': 1,
'湖北': 2,
'湖南': 2,
'河南': 2,
'江西': 2,
'北京': 3,
'天津': 3,
'河北': 3,
'山西': 3,
'内蒙古': 3,
'宁夏': 4,
'新疆': 4,
'青海': 4,
'陕西': 4,
'甘肃': 4,
'四川': 5,
'云南': 5,
'贵州': 5,
'西藏': 5,
'重庆': 5,
'辽宁': 6,
'吉林': 6,
'黑龙江': 6,
}
def time_transform(x):
time_local = time.localtime(x)
date = time.strftime("%Y%m%d%H%M%S%w", time_local)
return date
max_time = 1505138756
countryDic = {'日本':0,
'新加坡':1,
'中国香港':2,
'法国':3,
'意大利':4,
'加拿大':5}
countryDic1 = {'中国':0,
'中国台湾':1,
'中国澳门':2,
'中国香港':3,
'丹麦':4,
'俄罗斯':5,
'冰岛':6,
'加拿大':7,
'匈牙利':8,
'南非':9,
'卡塔尔':10,
'印度尼西亚':11,
'土耳其':12,
'埃及':13,
'墨西哥':14,
'奥地利':15,
'尼泊尔':16,
'巴西':17,
'希腊':18,
'德国':19,
'意大利':20,
'挪威':21,
'捷克':22,
'摩洛哥':23,
'斐济':24,
'新加坡':25,
'新西兰':26,
'日本':27,
'柬埔寨':28,
'比利时':29,
'毛里求斯':30,
'法国':31,
'波兰':32,
'泰国':33,
'澳大利亚':34,
'爱尔兰':35,
'瑞典':36,
'瑞士':37,
'缅甸':38,
'美国':39,
'老挝':40,
'芬兰':41,
'英国':42,
'荷兰':43,
'菲律宾':44,
'葡萄牙':45,
'西班牙':46,
'越南':47,
'阿联酋':48,
'韩国':49,
'马来西亚':50}
continentDic = {'亚洲':0,
'北美洲':1,
'南美洲':2,
'大洋洲':3,
'欧洲':4,
'非洲':5}
data_dir = "/home/x/hl/DC/datasets/data/"
test = False
action_train = pd.read_csv(data_dir+"trainingset/action_train.csv")
orderHistory_train = pd.read_csv(data_dir+"trainingset/orderHistory_train.csv")
userProfile_train = pd.read_csv(data_dir+"trainingset/userProfile_train.csv")
orderFuture_train = pd.read_csv(data_dir+"trainingset/orderFuture_train.csv")
userComment_train = pd.read_csv(data_dir+"trainingset/userComment_train.csv")
action_test=pd.read_csv(data_dir+'test/action_test.csv')
orderHistory_test=pd.read_csv(data_dir+'test/orderHistory_test.csv')
userComment_test=pd.read_csv(data_dir+'test/userComment_test.csv')
orderFuture_test=pd.read_csv(data_dir+'test/orderFuture_test.csv')
userProfile_test=pd.read_csv(data_dir+'test/userProfile_test.csv')
orderFuture_train.rename(columns={"orderType":"label"},inplace=True)
other_train = pd.read_csv("other_train_time_trans_42.csv")
other_test = pd.read_csv("other_test_time_trans_42.csv")
feature7_train = pd.read_csv("feature7_train.csv")
feature7_test = pd.read_csv("feature7_test.csv")
others_train = pd.merge(other_train,feature7_train,on="userid",how="left")
others_test = pd.merge(other_test,feature7_test,on="userid",how="left")
"""
返回用户的个人信息
1.性别
2.省份
3.年龄
<<dict见声明>>"""
def userprofile(userProfile):
df = userProfile.copy()
df["gender"] = df["gender"].map(genderDic)
df["province"] = df["province"].map(provinceDic1)
df["age"] = df["age"].map(ageDic)
return df
"""
用户评论相关特征
1.评分特征
2.用户评论内容特征(评论内容都是一些好的关键词,所以用数量来评判)
"""
def user_comment(userComment):
df = userComment.copy()
length = len(df)
key_words = [np.nan]*length
for index in range(length):
key_word = str(userComment["commentsKeyWords"][index])
if key_word != "nan":
key_words[index] = key_word.count(",")+1
df = df[["userid","rating"]]
df.rename(columns={"rating":"user_Comment_rating"},inplace=True)
df["user_Comment_key_words"] = key_words
return df.groupby("userid",as_index=False).sum()
"""统计去过每个大州的次数
"""
def ordered_continent(orderHistory):
df = orderHistory.copy()
df["continent"] = df["continent"].map(continentDic)
continent = pd.get_dummies(df["continent"], prefix='continent')
result = pd.concat([df["userid"],continent],axis =1)
result = result.groupby("userid",as_index=False).sum()
return result
"""统计去过每个城市的次数
"""
def ordered_country(orderHistory):
df = orderHistory.copy()
df["country"] = df["country"].map(countryDic)
country = pd.get_dummies(df["country"], prefix='country')
result = pd.concat([df["userid"],country],axis =1)
result = result.groupby("userid",as_index=False).sum()
return result
"""
1.订单总和
2.是否有过精品订单
3.精品订单总和
4.普通订单综合
5.精品订单比重
6.最后一次订单类型
7.最后一次订单时间
8.第一次订单时间
9.第一次订单类型
10.用户订单F1
"""
def orderhistory(orderHistory):
df = orderHistory.copy()
grouped = df[["userid", "orderType","orderTime"]].groupby("userid", as_index=False)
result = grouped.count()
result.rename(columns={"orderType":"ordered_count"},inplace=True)
result["ordered_sign"] = grouped.max()["orderType"]
result["ordered_supersum"] = grouped.sum()["orderType"]
result["ordered_end_type"] = grouped.last()["orderType"]
result["ordered_first_type"] = grouped.first()["orderType"]
result["ordered_end_Time"] = grouped.last()["orderTime"]
result["ordered_first_Time"] = grouped.first()["orderTime"]
result["ordered_simplesum"] = result["ordered_count"] - result["ordered_supersum"]
result["ordered_rate"] = result["ordered_supersum"]/result["ordered_count"]
result["ordered_historyF1"] = result["ordered_end_Time"] - result["ordered_first_Time"]
return result
"""
不同actionType占出现次数的比重
1.类型1数量
2.类型2数量
3.类型3数量
4.类型4数量
5.类型5数量
6.类型6数量
7.类型7数量
8.类型8数量
9.类型9数量
10.所有类型数量
11.类型1占比
12.类型2占比
13.类型3占比
14.类型4占比
15.类型5占比
16.类型6占比
17.类型7占比
18.类型8占比
19.类型9占比
"""
def action_actionType(action):
df = action.copy()
df["count"] = 1
df_count = df[["userid", "count"]].groupby("userid", as_index=False).count()
actionType = pd.get_dummies(action["actionType"], prefix='actionType')
result = pd.concat([df["userid"],actionType],axis=1)
result = result.groupby("userid", as_index=False).sum()
result["actionType_count"] = df_count["count"]
for column in range(1,10):
result["actionType{}_Per".format(column)] = result["actionType_{}".format(column)]/result["actionType_count"]
return result
"""
进行了聚类
234归为1类
789归为1类
每个类别:
最后一次出现时间
最后一次出现位置
第一次出现时间
第一次出现位置
"""
def actionType_sequence(action):
df = action.copy()
p = df[["userid", "actionType","actionTime"]].groupby("userid", as_index=False)
length = len(p.size())
type_total = 5
max_distance = [[np.nan] * length for _ in range(type_total)]
max_time = [[np.nan] * length for _ in range(type_total)]
min_distance = [[np.nan] * length for _ in range(type_total)]
min_time = [[np.nan] * length for _ in range(type_total)]
for index,(name, group) in enumerate(p):
actionType = np.array(group["actionType"])
actionTime = group["actionTime"]
actionType[actionType==3] = 2
actionType[actionType==4] = 2
actionType[actionType==5] = 3
actionType[actionType==6] = 4
actionType[actionType==7] = 5
actionType[actionType==8] = 5
actionType[actionType==9] = 5
actionType = list(actionType)
actionTime = list(actionTime)
firstTime = actionTime[0]
endTime = actionTime[-1]
action_set = set(actionType)
for number in range(type_total):
if (number + 1) in action_set:
loc = actionType.index(number + 1)
min_distance[number][index] = loc
min_time[number][index] = actionTime[loc]
actionType = actionType[::-1]
actionTime = actionTime[::-1]
for number in range(type_total):
if (number + 1) in action_set:
loc = actionType.index(number + 1)
max_distance[number][index] = loc
max_time[number][index] = actionTime[loc]
result = p.first()
del result["actionType"]
del result["actionTime"]
for column in range(type_total):
result["actionType_recent_position_{}".format(column + 1)] = max_distance[column]
for column in range(type_total):
result["actionType_recent_time_{}".format(column + 1)] = max_time[column]
for column in range(type_total):
result["actionType_first_position_{}".format(column + 1)] = min_distance[column]
for column in range(type_total):
result["actionType_first_time_{}".format(column + 1)] = min_time[column]
return result
"""
获取每个类别状态:
距离最后时间的时间差
距离最开始的时间差
begin-end持续时间差
"""
def actionType_recent(action):
p = action[["userid", "actionType", "actionTime"]].groupby("userid", as_index=False)
length = len(p.size())
type_total = 9
to_end = [[np.nan] * length for _ in range(type_total)]
to_begin = [[np.nan] * length for _ in range(type_total)]
begin_end = [[np.nan] * length for _ in range(type_total)]
for index,(name, group) in enumerate(p):
actionType = np.array(group["actionType"])
actionTime = np.array(group["actionTime"])
# actionType[actionType==3] = 2
# actionType[actionType==4] = 2
# actionType[actionType==5] = 3
# actionType[actionType==6] = 4
# actionType[actionType==7] = 5
# actionType[actionType==8] = 8
# actionType[actionType==9] = 8
begin_time = actionTime[0]
end_time = actionTime[-1]
action_set = set(actionType)
for number in range(type_total):
if (number + 1) in action_set:
begin,end = actionTime[actionType==number+1][0],actionTime[actionType==number+1][-1]
to_end[number][index] = end_time - end
to_begin[number][index] = begin - begin_time
begin_end[number][index] = end - begin
recentDic = p.first()
del recentDic["actionType"]
del recentDic["actionTime"]
for column in range(type_total):
recentDic["rangeTime_to_begin{}".format(column + 1)] = to_begin[column]
for column in range(type_total):
recentDic["rangeTime_to_end{}".format(column + 1)] = to_end[column]
for column in range(type_total):
recentDic["rangeTime_begin_end{}".format(column + 1)] = begin_end[column]
return recentDic
"""
1.第一个类型
2.第一个时间间隔
3.时间间隔最小值
4.时间间隔方差
5.时间间隔均值
6.最后一个类型
7.倒数第二个类型
8.倒数第三个类型
9.最后一个间隔
10.倒数第二个间隔
11.倒数第三个间隔
12.倒数第四个间隔
13.最后三个的均值
14.最后三个的方差
"""
def actionType_diff(action):
df = action.copy()
grouped = df[["userid", "actionTime", "actionType"]].groupby("userid", as_index=False)
length = len(grouped)
total_end,total_end_type = 4,3
end_stack = [[np.nan] * length for _ in range(total_end)]
end_type = [[np.nan] * length for _ in range(total_end_type)]
firstBrowse = [np.nan] * length
minBrowse = [np.nan] * length
varBrowse = [np.nan] * length
meanBrowse = [np.nan] * length
three_mean = [np.nan] * length
three_var = [np.nan] * length
for index, (name, group) in enumerate(grouped):
actionTime_diff = np.array(group["actionTime"].diff())[1:]
actionType = np.array(group["actionType"])[::-1]
if len(actionType)<1:continue
for number in range(min(len(actionTime_diff), total_end_type)):
end_type[number][index] = actionType[number]
if len(actionTime_diff) < 1: continue
firstBrowse[index] = actionTime_diff[0]
minBrowse[index] = np.min(actionTime_diff)
varBrowse[index] = np.var(actionTime_diff)
meanBrowse[index] = np.mean(actionTime_diff)
three_mean = np.mean(actionTime_diff[:3])
three_var = np.mean(actionTime_diff[:3])
actionTime_diff = actionTime_diff[::-1]
for number in range(min(len(actionTime_diff), total_end)):
end_stack[number][index] = actionTime_diff[number]
result = grouped.first()
result.rename(columns={"actionType": "action_firsttype"}, inplace=True)
del result["actionTime"]
result["action_firstBrowse"] = firstBrowse
result["action_minBrowse"] = minBrowse
result["action_varBrowse"] = varBrowse
result["action_meanBrowse"] = meanBrowse
result["action_endthreemean"] = three_mean
result["action_endthreevar"] = three_var
for number in range(total_end):
result["action_end{}Browse".format(number + 1)] = end_stack[number]
for number in range(total_end_type):
result["action_end{}type".format(number + 1)] = end_type[number]
return result
"""
针对类别x的时间间隔的mean,min,max,var等
"""
from scipy.fftpack import dct, fft
def actionType_X(action, *funcs):
grouped = action[["userid", "actionType", "actionTime"]].groupby("userid", as_index=False)
result = grouped.first()
total = 6
length = len(grouped.size())
interval_stack = [[[np.nan] * length for _ in range(total)] for _ in range(len(funcs))]
time_dct = [[np.nan] * length for _ in range(total)]
for index,(name, group) in enumerate(grouped):
actionTime = np.array(group["actionTime"])
actionType = np.array(group["actionType"])
actionType[actionType==3] = 2
actionType[actionType==4] = 2
actionType[actionType==5] = 3
actionType[actionType==6] = 4
actionType[actionType==7] = 5
actionType[actionType==8] = 6
actionType[actionType==9] = 6
for column in range(total):
test = actionTime[actionType==column+1]
if len(test)<1:continue
time_diff = test[1:] - test[:-1]
if len(time_diff)<1:continue
for number,func in enumerate(funcs):
interval_stack[number][column][index] = func(time_diff)
if len(time_diff)<3:continue
time_dct[column][index] = dct(time_diff)[2]
del result["actionType"]
del result["actionTime"]
for number,func in enumerate(funcs):
func_name = str(func).split(" ")[1]
for column in range(total):
result["actionType{}_interval{}".format(func_name,column+1)] = interval_stack[number][column]
# for column in range(total):
# result["dct_interval{}".format(column + 1)] = time_dct[column]
return result
"""
用户最后一次类别出现的时间差值
"""
def actionend_diff(action,pairs=None):
df = action.copy()
p = df[["userid", "actionType","actionTime"]].groupby("userid", as_index=False)
length = len(p.size())
type_total = 6
min_distance = np.array([[np.nan] * length for _ in range(type_total)])
min_time = np.array([[np.nan] * length for _ in range(type_total)])
for index,(name, group) in enumerate(p):
actionType = np.array(group["actionType"])
actionTime = group["actionTime"]
actionType[actionType==3] = 2
actionType[actionType==4] = 2
actionType[actionType==5] = 3
actionType[actionType==6] = 4
actionType[actionType==7] = 5
actionType[actionType==8] = 6
actionType[actionType==9] = 6
actionType = list(actionType)
actionTime = list(actionTime)
endTime = actionTime[-1]
actionType = actionType[::-1]
actionTime = actionTime[::-1]
action_set = set(actionType)
for number in range(type_total):
if (number + 1) in action_set:
loc = actionType.index(number + 1)
min_distance[number][index] = loc
min_time[number][index] = actionTime[loc]
result = p.first()
del result["actionType"]
del result["actionTime"]
for i in range(type_total-2):
for j in range(type_total-1):
if i == j:continue
result["typeend{}_{}diff".format(i+1,j+1)] = min_time[i]-min_time[j]
# for i in range(type_total):
# for j in range(type_total):
result["typeend{}_{}dist".format(i+1,j+1)] = min_distance[i] - min_distance[j]
return result
def actiontype_cloest(actions):
df = actions.copy()
total = 5
type_5 = {1: 1,
2: 2,
3: 2,
4: 2,
5: 3,
6: 4,
7: 5,
8: 5,
9: 5,
}
df["actionType"] = df["actionType"].map(type_5)
result = df[["userid"]].drop_duplicates().reset_index()
del result["index"]
for i in range(1,total+1):
for j in range(1,total+1):
type_ij = df[(df["actionType"] == i)|(df["actionType"] == j)]
next_type = list(type_ij["actionType"])[1:]
next_type.append(1)
next_time = list(type_ij["actionTime"])[1:]
next_time.append(max_time)
next_user = list(type_ij["userid"])[1:]
next_user.append(0)
type_ij["type_diff"] = next_type - type_ij["actionType"]
type_ij["time_diff"] = next_time - type_ij["actionTime"]
type_ij["user_diff"] = next_user - type_ij["userid"]
user_self = type_ij[(type_ij["type_diff"] == j - i) &(type_ij["user_diff"]==0)]
res = user_self.groupby("userid",as_index=False)["time_diff"].agg({"type{}_{}_closestmean".format(i,j):"mean",
}).reset_index()
# "type{}_{}_closestmin".format(i, j): "min",
del res["index"]
ij_first = user_self.sort_values(by='actionTime', ascending=True).drop_duplicates('userid')
ij_first = ij_first[['userid', 'time_diff']].rename(columns={'time_diff': 'firstclosest_{}_{}'.format(i, j)})
ij_last = user_self.sort_values(by='actionTime', ascending=False).drop_duplicates('userid')
ij_last = ij_last[['userid', 'time_diff']].rename(columns={'time_diff': 'endclosest_{}_{}'.format(i, j)})
result = pd.merge(result,res,on='userid',how="left")
# result = pd.merge(result,ij_first,on='userid',how="left")
result = pd.merge(result,ij_last,on='userid',how="left")
return result
"""
用户actiontype序列转移的时间间隔
原始的9类别进行了6类别划分(保证充足的数据)
"""
def type_typevalue(action,*funcs):
grouped = action[["userid", "actionType","actionTime"]].groupby("userid", as_index=False)
length = len(grouped.size())
total = 6
type2typeintervals = [[[[np.nan]*length for _ in range(total)] for _ in range(total)] for _ in range(len(funcs))]
for index,(name, group) in enumerate(grouped):
type2typeinterval = [[[] for _ in range(total)] for _ in range(total)]
actionType = np.array(group["actionType"])
actionType[actionType==3] = 2
actionType[actionType==4] = 2
actionType[actionType==5] = 3
actionType[actionType==6] = 4
actionType[actionType==7] = 5
actionType[actionType==8] = 6
actionType[actionType==9] = 6
actionTime = np.array(group["actionTime"])
interval = actionTime[1:] - actionTime[:-1]
for i in range(len(actionType)-1):
type2typeinterval[actionType[i]-1][actionType[i+1]-1].append(interval[i])
typeset = set(actionType)
for number,func in enumerate(funcs):
for type in set(typeset):
for typeto in set(typeset):
if type2typeinterval[type-1][typeto-1] == []:continue
type2typeintervals[number][type-1][typeto-1][index] = func(type2typeinterval[type-1][typeto-1])
result = grouped.first()
del result["actionType"]
del result["actionTime"]
for number, func in enumerate(funcs):
func_name = str(func).split(" ")[1]
for i in range(total-1):
for j in range(total-2):
result["type_{}to{}value{}".format(i + 1,j + 1,func_name)] = type2typeintervals[number][i][j]
return result
"""
用户actiontype序列转移的时间间隔
原始的9类别进行了6类别划分(保证充足的数据)
"""
def type_tototypevalue(action, *funcs):
grouped = action[["userid", "actionType", "actionTime"]].groupby("userid", as_index=False)
length = len(grouped.size())
total = 6
type2typeintervals = [[[[np.nan] * length for _ in range(total)] for _ in range(total)] for _ in range(len(funcs))]
for index, (name, group) in enumerate(grouped):
type2typeinterval = [[[] for _ in range(total)] for _ in range(total)]
actionType = np.array(group["actionType"])
actionType[actionType == 3] = 2
actionType[actionType == 4] = 2
actionType[actionType == 5] = 3
actionType[actionType == 6] = 4
actionType[actionType == 7] = 5
actionType[actionType == 8] = 6
actionType[actionType == 9] = 6
actionTime = np.array(group["actionTime"])
interval1 = actionTime[1:] - actionTime[:-1]
interval2 = actionTime[2:] - actionTime[:-2]
for i in range(len(actionType) - 2):
type2typeinterval[actionType[i] - 1][actionType[i + 1] - 1].append(interval1[i])
type2typeinterval[actionType[i] - 1][actionType[i + 2] - 1].append(interval2[i])
typeset = set(actionType)
for number, func in enumerate(funcs):
for type in set(typeset):
for typeto in set(typeset):
if type2typeinterval[type - 1][typeto - 1] == []: continue
type2typeintervals[number][type - 1][typeto - 1][index] = func(
type2typeinterval[type - 1][typeto - 1])
result = grouped.first()
del result["actionType"]
del result["actionTime"]
for number, func in enumerate(funcs):
func_name = str(func).split(" ")[1]
for i in range(total):
for j in range(total):
result["type_{}toto{}value{}".format(i + 1, j + 1, func_name)] = type2typeintervals[number][i][j]
return result
"""
获取最后window个动作的onehot编码
"""
def get_last_action_type(actions,*funcs):
"""
"""
grouped = actions[["userid","actionType"]].groupby('userid', as_index=False)
length = len(grouped)
total = 7
window = 3
type_end = [[np.nan]*length for _ in range(total)]
interval_stack = [[np.nan] * length for __ in range(len(funcs))]
for index,(name, group) in enumerate(grouped):
actionType = np.array(group['actionType'])
actionType[actionType==3] = 2
actionType[actionType==4] = 2
actionType[actionType==5] = 3
actionType[actionType==6] = 4
actionType[actionType==7] = 5
actionType[actionType==8] = 6
actionType[actionType==9] = 7
if len(actionType) < window: # 如果不够 n_action 个动作
actionType = np.concatenate([[np.nan] * (window - len(actionType)), actionType], axis=0)
else:
actionType = actionType[-window:]
for column in range(total):
type_end[column][index] = np.sum(actionType==column+1)
for number,func in enumerate(funcs):
interval_stack[number][index] = func(actionType)
result = grouped.first()
del result["actionType"]
for column in range(total):
result["type_end_{}".format(column+1)]= type_end[column]
for number, func in enumerate(funcs):
func_name = str(func).split(" ")[1]
result["type_end_{}".format(func_name)]= interval_stack[number]
return result
"""用户最后一单的年 月 日 小时 分特征
"""
def orderaction_date(action):
grouped = action[["userid", "actionTime"]].groupby("userid", as_index = False)
result = grouped.max()
# result["first_ordertime"] = result.actionTime.apply(lambda time: max_time - time)
# result["end_ordertime"] = grouped.max().actionTime.apply(lambda time: max_time - time)
# result["median_mediantime"] = grouped.median().actionTime.apply(lambda time:max_time - time)
date = result.actionTime.apply(time_transform)
result["year_last"] = date.apply(lambda x:int(x[:4]))
result["month_last"] = date.apply(lambda x:int(x[4:6]))
result["day_last"] = date.apply(lambda x:int(x[6:8]))
result["hour_last"] = date.apply(lambda x:int(x[8:10]))
result["minute_last"] = date.apply(lambda x:int(x[10:12]))
# result["weekday_last"] = date.apply(lambda x:int(x[12:13]))
del result["actionTime"]
return result
def action_weekday(action):
df = action.copy()
date = df.actionTime.apply(time_transform)
df["weekday"] = date.apply(lambda x: int(x[12:13]))
df["count"] = 1
df_count = df[["userid", "count"]].groupby("userid", as_index=False).count()
weekday = pd.get_dummies(df["weekday"], prefix='weekday')
tttt = pd.concat([df[["userid", "actionType"]], weekday], axis=1)
result = df[["userid"]].drop_duplicates().reset_index()
del result["index"]
for column in range(6):
tmp = tttt[tttt["weekday_{}".format(column)] == 1]
res = tmp[["userid", "actionType"]].groupby("userid", as_index=False)["actionType"].agg(
{"weekday{}actionTypemean".format(column): "mean",
"weekday{}actionTypevar".format(column): "var",
"weekday{}actionTypmin".format(column): "min",
# "weekday{}actionTypemax".format(column): "max",
})
result = pd.merge(result, res, on="userid", how="left")
return result
action_weekday(action_test)
"""
进行了聚类
234归为1类
789归为1类
每个类别:
最后一次出现时间得年月日表示
"""
def actionType_end_yearmd(action):
df = action.copy()
p = df[["userid", "actionType", "actionTime"]].groupby("userid", as_index=False)
length = len(p.size())
type_total = 5
max_time = [[0] * length for _ in range(type_total)]
for index, (name, group) in enumerate(p):
actionType = np.array(group["actionType"])
actionTime = group["actionTime"]
actionType[actionType == 3] = 2
actionType[actionType == 4] = 2
actionType[actionType == 5] = 3
actionType[actionType == 6] = 4
actionType[actionType == 7] = 5
actionType[actionType == 8] = 5
actionType[actionType == 9] = 5
actionType = list(actionType)
actionTime = list(actionTime)
action_set = set(actionType)
actionType = actionType[::-1]
actionTime = actionTime[::-1]
for number in range(type_total):
if (number + 1) in action_set:
loc = actionType.index(number + 1)
max_time[number][index] = actionTime[loc]
result = p.first()
del result["actionType"]
del result["actionTime"]
for column in range(type_total):
result["actionTime{}".format(column+1)] = max_time[column]
date = result["actionTime{}".format(column+1)].apply(time_transform)
# result["actionType{}year_last".format(column+1)] = date.apply(lambda x:int(x[:4]))
result["actionType{}month_last".format(column+1)] = date.apply(lambda x:int(x[4:6]))
result["actionType{}day_last".format(column+1)] = date.apply(lambda x:int(x[6:8]))
result["actionType{}hour_last".format(column+1)] = date.apply(lambda x:int(x[8:10]))
result["actionType{}minute_last".format(column+1)] = date.apply(lambda x:int(x[10:12]))
del result["actionTime{}".format(column+1)]
return result
def action_orderwindow(action):
df = action.copy()
grouped = df[["userid","actionTime","actionType"]].groupby("userid",as_index = False)
result = grouped.first()
length = len(grouped)
window_size = 10
total = 6
orderwindow = [[np.nan]*length for _ in range(total)]
for index,(name,group) in enumerate(grouped):
actionType = np.array(group["actionType"])
actionTime = np.array(group["actionTime"])
actionType[actionType == 3] = 2
actionType[actionType == 4] = 2
actionType[actionType == 5] = 3
actionType[actionType == 6] = 4
actionType[actionType == 7] = 5
actionType[actionType == 8] = 6
actionType[actionType == 9] = 6
interval = np.array(group["actionTime"].diff())
ordertype9 = actionType==total
window = np.array([False]*len(ordertype9))
for w in range(1,min(len(ordertype9),window_size)+1):
window[:-w] += ordertype9[w:]
for column in range(total):
orderwindow[column][index] = np.sum(actionType[window] == column+1)
for column in range(total):
result["ordered_inwindow_type{}".format(column+1)] = orderwindow[column]
del result["actionTime"]
del result["actionType"]
return result
"""
时间间隔相关特征
离最近一次的actiontype的时间间隔func
actiontype:1/2/3/4/5/6/7/8/9
func:np.mean / np.min / np.max / np.var
median后期可以考虑加入
"""
def actionType_sequence_recentdiff(action, *funcs):
df = action.copy()
p = df[["userid", "actionType", "actionTime"]].groupby("userid", as_index=False)
result = p.first()
type_total = 6
length = len(p.size())
interval_stack = [[[np.nan] * length for _ in range(type_total)] for __ in range(len(funcs))]
for index,(name, group) in enumerate(p):
#时间间隔取log,我也不知道为啥就提升了,不取就下降
interval = np.array(group["actionTime"].diff())[::-1]
actionType = np.array(group["actionType"])
actionType[actionType == 2] = 2
actionType[actionType == 3] = 2
actionType[actionType == 4] = 2
actionType[actionType == 5] = 3
actionType[actionType == 6] = 4
actionType[actionType == 7] = 5
actionType[actionType == 8] = 6
actionType[actionType == 9] = 6
actionType = list(actionType)[::-1]
action_set = set(actionType)
for number in range(type_total):
if number + 1 in action_set:
test = interval[:actionType.index(number + 1)]
if len(test) == 0:
continue
for i,func in enumerate(funcs):
interval_stack[i][number][index] = func(test)
del result["actionType"]
del result["actionTime"]
for i,func in enumerate(funcs):
func_name = str(func).split(" ")[1]
for column in range(type_total):
result["actionType_recent_diff_{}{}".format(func_name, column + 1)] = interval_stack[i][column]
return result
def end_day_action(actions,*funcs):
df = actions.copy()
df["yearmonthday"] = df["actionTime"].apply(time_transform).apply(lambda x: int(x[:8]))
grouped = df[["userid", "yearmonthday", "actionTime","actionType"]].groupby("userid", as_index=False)
result = grouped.last()
length = len(grouped)
total = 6
window = 3
count_stack = [np.nan] * length
max_stack = [np.nan] * length
end_datydiff = [[np.nan] * length for _ in range(len(funcs))]
end_daytype = [[np.nan] * length for _ in range(len(funcs))]
number_stack = [[np.nan] * length for _ in range(total)]
rate_stack = [[np.nan] * length for _ in range(total)]
for index, (name, group) in enumerate(grouped):
yearmonthday = np.array(group["yearmonthday"])
actionType = np.array(group["actionType"])
actionTime = np.array(group["actionTime"])
actionType[actionType == 3] = 2
actionType[actionType == 4] = 2
actionType[actionType == 5] = 3
actionType[actionType == 6] = 4
actionType[actionType == 7] = 5
actionType[actionType == 8] = 6
actionType[actionType == 9] = 6
end_day = np.max(yearmonthday)
windowday = np.array([False] * len(yearmonthday))
for w in range(window):
windowday += yearmonthday == end_day
end_day -= 1
count_stack[index] = np.sum(windowday)
if count_stack[index] == 0: continue
max_stack[index] = np.max(actionType[windowday])
for column in range(total):
number_stack[column][index] = 1.0 * np.sum(actionType[windowday] == column + 1) # / count_stack[index]
rate_stack[column][index] = 1.0 * np.sum(actionType[windowday] == column + 1)/ count_stack[index]
if count_stack[index] == 1:continue
time_diff = actionTime[windowday][1:] - actionTime[windowday][0:-1]
for column,func in enumerate(funcs):
end_datydiff[column][index] = func(time_diff)
end_daytype[column][index] = func(actionType[windowday])
del result["actionType"]
for column in range(total-1):
result["enddaytype{}number".format(column + 1)] = number_stack[column]
# result["enddaytype{}rate".format(column + 1)] = rate_stack[column]
for column,func in enumerate(funcs):
func_name = str(func).split(" ")[1]
result["endday_{}".format(func_name)] = end_datydiff[column]
# result["enddaytype_{}".format(func_name)] = end_daytype[column]
result["enddaymaxtype"] = max_stack
result["enddaycount"] = count_stack
return result
"""
类别转移概率矩阵
"""
def type_type(action):
grouped = action[["userid", "actionType"]].groupby("userid", as_index=False)
length = len(grouped.size())
total = 6
continue_type = [[[np.nan] * length for _ in range(total)] for _ in range(total)]
for index,(name, group) in enumerate(grouped):
actionType = np.array(group["actionType"])
actionType[actionType==3] = 2
actionType[actionType==4] = 2
actionType[actionType==5] = 3
actionType[actionType==6] = 4
actionType[actionType==7] = 5
actionType[actionType==8] = 6
actionType[actionType==9] = 6
if len(actionType) <= 2:
continue
t = actionType[:-2]
type_to = actionType[1:-1]
type_to_to = actionType[2:]
for i in range(total):
for j in range(total):
continue_type[i][j][index] = 1.0*np.sum((type_to==j+1)&(t==i+1)) + 1.0*np.sum((type_to_to==j+1)&(t==i+1))
result = grouped.first()
del result["actionType"]
for i in range(total - 1):
for j in range(total - 1):
result["type_{}_to_type_{}".format(i+1,j+1)] = continue_type[i][j]
return result
test = actiontype_cloest(action_train)
test
test.describe()
def gen_data(orderFuture_X, action_X, orderHistory_X,userProfile_X,userComment_X,others_X):
df = orderFuture_X.copy()
df = pd.merge(df,userprofile(userProfile_X),on="userid",how="left")
df = pd.merge(df,user_comment(userComment_X),on="userid",how="left")
df = pd.merge(df,ordered_country(orderHistory_X),on="userid",how="left")
df = pd.merge(df,ordered_continent(orderHistory_X),on="userid",how="left")
#f = pd.merge(df,history_f1(orderHistory_X),on="userid",how="left")
df = pd.merge(df,orderhistory(orderHistory_X),on="userid",how="left")
df = pd.merge(df,action_actionType(action_X),on="userid",how="left")
#df = pd.merge(df,actionType_sequence_begin(action_X),on="userid",how="left")
df = pd.merge(df,actionType_sequence(action_X),on="userid",how="left")
df = pd.merge(df,actionType_recent(action_X),on="userid",how="left")
df = pd.merge(df,actionType_diff(action_X),on="userid",how="left")
df = pd.merge(df,actionType_X(action_X,np.min),on="userid",how="left")
df = pd.merge(df,actionend_diff(action_X),on="userid",how="left")
df = pd.merge(df,type_typevalue(action_X,np.min,np.mean),on="userid",how="left")
df = pd.merge(df,orderaction_date(action_X),on="userid",how="left")
df = pd.merge(df,get_last_action_type(action_X,np.mean),on="userid",how="left")
df = pd.merge(df,actiontype_cloest(action_X),on="userid",how="left")
#96.896 707 97.17
df = pd.merge(df,others_X,on="userid",how="left")
# df = pd.merge(df,actionType_sequence_recentdiff(action_X,np.mean,np.min),on="userid",how="left")
# df = pd.merge(df,others_X,on="userid",how="left")
# df = pd.merge(df,type_type(action_X),on="userid",how="left")
# # df = pd.merge(df,continue_dic(action_X,total=7),on="userid",how="left")
return df
friend_train = pd.read_csv("friend_train.csv")
friend_test = pd.read_csv("friend_test.csv")
train_other_features1 = pd.read_csv("./liuqing/features/train_other_features1.csv")
test_other_features1 = pd.read_csv("./liuqing/features/test_other_features1.csv")
others_train = friend_train.copy()
others_test = friend_test.copy()
others_train = pd.merge(others_train,train_other_features1,on="userid",how="left")
others_test = pd.merge(others_test,test_other_features1,on="userid",how="left")
actions = gen_data(orderFuture_train, action_train, orderHistory_train,userProfile_train,userComment_train,others_train)
test_data = gen_data(orderFuture_test, action_test, orderHistory_test,userProfile_test,userComment_test,others_test)
others_test
"""
97.058 539 67 train_action_history_features
[616] cv_agg's auc: 0.970457 train_action_history_features10
[473] cv_agg's auc: 0.970434 train_action_history_features4 25
[510] cv_agg's auc: 0.970482 train_action_history_features6 52
[569] cv_agg's auc: 0.970475 train_action_order_features1 16
[673] cv_agg's auc: 0.97043 train_advance_action_features 42
[702] cv_agg's auc: 0.970963 train_other_features1 101
[529] cv_agg's auc: 0.97028 train_user_order_comment_features 7
"""
importance_feature = ["userid",
"goodorder_vs_actiontype_1_ratio",
'isOrder',
'total_good_order_ratio',
'has_good_order',
'goodorder_vs_actiontype_5_ratio',
'finalAction_4',
'action_type_511_time_delta_min',
'finalAction_8',
'action_type_511_time_delta_max',
'goodorder_vs_actiontype_6_ratio',
'type_1to4valuemean',
'histord_sum_cont4',
'age_lg90',
'three_gram_789_last_time',
'three_gram_789_time_mean',
'action_type_710_time_delta_min',
'three_gram_456_time_min',
'three_gram_action_456_ratio',
'pay_money_min_delta',
'three_gram_123_time_mean',
'two_gram_23_time_std',
'histord_sum_cont3',]
other_train = pd.read_csv("./liuqing/train_0.97329.csv")
other_test = pd.read_csv("./liuqing/test_0.97329.csv")
other_train = other_train[importance_feature]
other_test = other_test[importance_feature]
print len(other_train),len(other_train.keys())
train_data = actions.copy()
tttt = test_data.copy()
train_data = pd.merge(train_data,other_train,on="userid",how="left")
tttt = pd.merge(tttt,other_test,on="userid",how="left")
# train_data = pd.merge(train_data,type_tototypevalue(action_train,np.median,np.mean),on="userid",how="left")
# tttt = pd.merge(tttt,type_tototypevalue(action_test,np.median,np.mean),on="userid",how="left")
# train_data = pd.merge(train_data,action_features_train,on="userid",how="left")
# tttt = pd.merge(tttt,action_features_test,on="userid",how="left")
# train_data = pd.merge(train_data,action_orderwindow(action_train),on="userid",how="left")
# tttt = pd.merge(tttt,action_orderwindow(action_test),on="userid",how="left")
# train_data = pd.merge(train_data,end_day_action(action_train,np.mean,np.min,np.max,np.std),on="userid",how="left")
# tttt = pd.merge(tttt,end_day_action(action_test,np.mean,np.min,np.max,np.std),on="userid",how="left")
# train_data = pd.merge(train_data,actionType_end_yearmd(action_train),on="userid",how="left")
# tttt = pd.merge(tttt,actionType_end_yearmd(action_test),on="userid",how="left")
# train_data = pd.merge(train_data,action_weekday(action_train),on="userid",how="left")
# tttt = pd.merge(tttt,action_weekday(action_test),on="userid",how="left")
# train_data = pd.merge(train_data,actionType_sequence_recentdiff(action_train,np.mean,np.min),on="userid",how="left")
# tttt = pd.merge(tttt,actionType_sequence_recentdiff(action_test,np.mean,np.min),on="userid",how="left")
# train_data = pd.merge(train_data,type_type(action_train),on="userid",how="left")
# tttt = pd.merge(tttt,type_type(action_test),on="userid",how="left")
from sklearn.model_selection import train_test_split
import lightgbm as lgb
params = {
'objective': 'binary',
'metric': {'auc'},
'learning_rate': 0.05,
'num_leaves': 30,
'min_sum_hessian_in_leaf': 0.1,
'feature_fraction': 0.3,
'bagging_fraction': 0.5,
'lambda_l1': 0,
'lambda_l2': 5,
'num_thread':3
}
# params = {
# 'objective': 'binary',
# 'metric': {'auc'},
# 'learning_rate': 0.05,
# 'num_leaves': 30,
# 'min_sum_hessian_in_leaf': 0.1,
# 'feature_fraction': 0.3,
# 'bagging_fraction': 0.5,
# 'lambda_l1': 3,
# 'lambda_l2': 5,
# 'num_thread':3
# }
if "label" in train_data:
train_label = train_data["label"]
del train_data["label"]
X_train = train_data
dtrain = lgb.Dataset(X_train, label=train_label)
cv_result = lgb.cv(params, dtrain, num_boost_round = 3000,verbose_eval=True, nfold=5, seed=3,metrics='auc', early_stopping_rounds=30, show_stdv=False)
best_round = len(cv_result['auc-mean'])
best_auc = cv_result['auc-mean'][-1] # 最好的 auc 值
best_model = lgb.train(params, dtrain, best_round)
best_auc
for i,j in zip(best_model.feature_name(),best_model.feature_importance()):
print i,j
print train_data.keys()
print tttt.keys()
# ---将特征重要性写入文件--
feature_score = best_model.feature_importance()
feature_names = best_model.feature_name()
m = {}
for i in range(len(feature_names)):
m[feature_names[i]] = feature_score[i]
m = sorted(m.items(), key=lambda x: x[1], reverse=True)
fs = []
for (key, value) in m:
fs.append("{0},{1}\n".format(key, value))
with open('lgb_feature_confuseFeat.csv', 'w') as f:
f.writelines("feature,score\n")
f.writelines(fs)
# print(pd.DataFrame({"feature_names": feature_names, "feature_score": list(feature_score)}).sort_values(
# by="feature_score", ascending=False))
# ---得到预测值--
tttt = feature_rest(tttt, train_data.keys())
if "orderType" in orderFuture_test:
del orderFuture_test["orderType"]
if "orderType" in test_data:
del test_data["orderType"]
y = best_model.predict(tttt) # 输出的是概率结果
orderFuture_test['orderType']=y
orderFuture_test.to_csv('baseline_wxr.csv',index=False)
orderFuture_test.head()
plt.show()
m = {}
for i in range(len(feature_names)):
m[feature_names[i]] = feature_score[i]
m = sorted(m.items(), key=lambda x: x[1], reverse=True)
fs = []
number = 0
for (key, value) in m:
number += 1
fs.append("{0},{1}\n".format(key, value))
if number>=20:break
with open('lgb_featureimportance_top20HL.csv', 'w') as f:
f.writelines("feature,score\n")
f.writelines(fs)
best_model.
HL = pd.read_csv("9733aLQ.csv")
LQ = pd.read_csv("97369bHL.csv")
sum(HL.userid == LQ.userid)
HL.head()
LQ.head()
HL.orderType = (HL.orderType + LQ.orderType)/2
HL.to_csv('LQHL.csv',index=False)
tttt
# train = actiontype_cloest(action_train)
# test = actiontype_cloest(action_test)
# train_data.rename(columns={"label": "orderType"}, inplace=True)
train_data.to_csv('train_dataHL.csv',index=False)
tttt.to_csv('test_dataHL.csv',index=False)
tttt
```
| github_jupyter |
# 作業目標:
了解 Padding 的作法
了解 Pooling 的作法
# 作業重點:
(1) 調整Padding 的設定, 看 Pad 後的結果
(2) 調整Pooling 的設定, 看 Poolin 後的結果
```
import numpy as np
import h5py
import matplotlib.pyplot as plt
# 繪圖結果直接顯示在Jupyter cell 之內
%matplotlib inline
plt.rcParams['figure.figsize'] = (5.0, 4.0) # 設定繪圖板的大小
plt.rcParams['image.interpolation'] = 'nearest'
plt.rcParams['image.cmap'] = 'gray'
# autoreload。可以讓我們不退出IPython就動態修改代碼,在執行代碼前IPython會幫我們自動重載改動的模塊
%load_ext autoreload
%autoreload 2
np.random.seed(1)
# GRADED FUNCTION: zero_pad
def zero_pad(X, pad):
"""
對image X 做 zero-padding.
參數定義如下:
X -- python numpy array, 呈現維度 (m, n_H, n_W, n_C), 代表一批 m 個圖像
n_H: 圖高, n_W: 圖寬, n_C: color channels 數
pad -- 整數, 加幾圈的 zero padding.
Returns:
X_pad -- image of shape (m, n_H + 2*pad, n_W + 2*pad, n_C) 做完zero-padding 的結果
"""
X_pad = np.pad(X, ((0, 0), (pad, pad), (pad, pad), (0, 0)), 'constant', constant_values=(0, 0))
return X_pad
'''
作業修改
np.random.seed(1)
x =
x_pad =
'''
x = np.random.randn(4, 3, 3, 2) #產生gray image
x_pad = zero_pad(x, 1) # 加兩圈 Pad
print ("x.shape =", x.shape)
print ("x_pad.shape =", x_pad.shape)
print ("x[1,1] =", x[1,1])
print ("x_pad[1,1] =", x_pad[1,1])
fig, axarr = plt.subplots(1, 2)
axarr[0].set_title('x')
axarr[0].imshow(x[0,:,:,0])
axarr[1].set_title('x_pad')
axarr[1].imshow(x_pad[0,:,:,0])
# GRADED FUNCTION: pool_forward
def pool_forward(A_prev, hparameters, mode = "max"):
"""
設計一個前行網路的池化層
參數定義如下:
A_prev -- 輸入的numpy 陣列, 維度 (m, n_H_prev, n_W_prev, n_C_prev)
hparameter 超參數 -- "f" and "stride" 所形成的python 字典
mode -- 池化的模式: "max" or "average"
返回:
A -- 輸出的池化層, 維度為 (m, n_H, n_W, n_C) 的 numpy 陣列
cache -- 可以應用在 backward pass pooling layer 資料, 包含 input and hparameter
"""
# 檢索尺寸 from the input shape
(m, n_H_prev, n_W_prev, n_C_prev) = A_prev.shape
# 檢索超參數 from "hparameters"
f = hparameters["f"]
stride = hparameters["stride"]
# 定義輸出的dimensions
n_H = int(1 + (n_H_prev - f) / stride)
n_W = int(1 + (n_W_prev - f) / stride)
n_C = n_C_prev
# 初始化輸出的 matrix A
A = np.zeros((m, n_H, n_W, n_C))
### 程式起始位置 ###
for i in range(m): # 訓練樣本的for 迴圈
for h in range(n_H): # 輸出樣本的for 迴圈, 針對vertical axis
for w in range(n_W): # 輸出樣本的for 迴圈, 針對 horizontal axis
for c in range (n_C): # 輸出樣本的for 迴圈, 針對channels
# 找出特徵圖的寬度跟高度四個點
vert_start = h * stride
vert_end = h * stride+ f
horiz_start = w * stride
horiz_end = w * stride + f
# 定義第i個訓練示例
a_prev_slice = A_prev[i, vert_start:vert_end, horiz_start:horiz_end,c]
# 計算輸入data 的池化結果. 使用 if statment 去做分類
if mode == "max":
A[i, h, w, c] = np.max(a_prev_slice)
elif mode == "average":
A[i, h, w, c] = np.mean(a_prev_slice)
### 程式結束 ###
# 儲存輸入的特徵圖跟所設定的超參數, 可以用在 pool_backward()
cache = (A_prev, hparameters)
# 確認輸出的資料維度
assert(A.shape == (m, n_H, n_W, n_C))
return A, cache
'''
作業修改
np.random.seed(1)
A_prev =
hparameters =
'''
np.random.seed(1)
A_prev = np.random.randn(2, 4, 4, 3)
hparameters = {"stride" : 2, "f": 3}
A, cache = pool_forward(A_prev, hparameters)
A, cache = pool_forward(A_prev, hparameters)
print("mode = max")
print("A =", A)
print()
A, cache = pool_forward(A_prev, hparameters, mode = "average")
print("mode = average")
print("A =", A)
```
| github_jupyter |
# Display financial data.
This notebook assumes that you have downloaded daily data for each stock of the PHLX Oil Services Sector (^OLX) and the data is stored in CSV format with names: {ticker}\_data.csv. For example, for the CHX stocks the data is stored in CHX_data.csv.
All finantial data can be found in Yahoo finance [here](https://finance.yahoo.com/quote/%5EOSX/components?p=%5EOSX). The data can be manually downloaded from the web page and, while one can set up a downloading script in Python, I prefer not to publish it.
This notebook does the following:
1. Display the form of the correlation matrices: $\Sigma$ and $g$ for data between selected dates.
2. Plot the returns of the index and of each individual stock for a selected range of dates.
```
import sys
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import cm
from matplotlib.colors import ListedColormap, LinearSegmentedColormap
import pandas as pd
sys.path.append('../')
from index_tracking.get_data import make_correlation_matrices, read_stock_data
```
## Parameters
* Ticker name of the index
* Ticker name of the stocks
* Location of the stored market data
* Date range between which data is analysed
```
ticker_index = 'OSX'
# Tickers inside the index
tickers = ['CHX', 'CLB', 'DRQ', 'GLNG',
'HAL', 'HP', 'LBRT', 'NBR',
'NOV', 'OII', 'OIS', 'RIG',
'SLB', 'USAC', 'WHD']
# Location where all stock data is stored.
data_location = '../../../data/'
# Day range of data.
day_first = '2021-01-10'
day_last = '2021-01-30'
```
## Plot correlation matrices
```
def plot_correlation_matrices():
Σ, g, ϵ0 = make_correlation_matrices(
ticker_index, tickers, data_location, day_first, day_last
)
N = len(tickers)
# PLOTS.
viridis_big = cm.get_cmap('bwr', 512)
mins = np.min(Σ)
b = mins/2 + 0.5
newcmp = ListedColormap(viridis_big(np.linspace(b, 1.0, 256)))
fig, ax = plt.subplots(1, 1, figsize=(6, 5), constrained_layout=True)
# Reshape g to put it toghether with Σ.
g = np.reshape(g, (N, 1))
im = ax.imshow(np.concatenate((Σ, g), axis=1), cmap=newcmp)
ax.set_xticks(np.arange(N+1))
ax.set_yticks(np.arange(N))
ax.set_xticklabels([*tickers, ticker_index])
ax.set_yticklabels(tickers)
# Rotate the tick labels and set their alignment.
plt.setp(ax.get_xticklabels(), rotation=45, ha="right",
rotation_mode="anchor")
plt.colorbar(im, ax=ax)
plt.show()
return
plot_correlation_matrices()
```
## Plot daily return data
```
def plot_daily_returns():
# Get daily returns of the index and all stocks.
rI = read_stock_data(
ticker_index, data_location, day_first, day_last
)
N = len(tickers)
r = []
for ix, ticker in enumerate(tickers):
rstock = read_stock_data(
ticker, data_location, day_first, day_last
)
r.append(rstock)
# Get market dates between first and last days.
datafile = data_location + f'{ticker_index}_data.csv'
df = pd.read_csv(datafile, parse_dates=['Date'])
dates = df.loc[(df['Date'] >= day_first) & (df['Date'] <= day_last),'Date'].to_list()
dates = [date.strftime('%Y-%m-%d') for date in dates]
# PLOTS.
lfs = 20
tfs = 16
fig, ax = plt.subplots(1, 1, figsize=(6, 5), constrained_layout=True)
# Plot returns.
for i in range(N):
ax.plot(r[i], '0.7', lw=1)
ax.plot(rI, lw=3, label='Index')
# Indicate zero returns.
ax.axhline(0.0, c='k', ls='--', lw=1)
ax.legend(fontsize=lfs)
# Labels and ticks.
ax.set_xlabel('day', fontsize=lfs)
ax.set_ylabel('returns', fontsize=lfs)
ax.set_xticks(np.arange(len(dates)-1))
ax.set_xticklabels(dates[1:])
ax.tick_params(axis='x', labelsize=0.8*tfs)
ax.tick_params(axis='y', labelsize=tfs)
plt.setp(ax.get_xticklabels(), rotation=45, ha="right",
rotation_mode="anchor")
plt.show()
return
plot_daily_returns()
```
| github_jupyter |
# Use Your Own Inference Code with Amazon SageMaker XGBoost Algorithm
_**Customized inference for computing SHAP values with Amazon SageMaker XGBoost script mode**_
---
## Contents
1. [Introduction](#Introduction)
2. [Setup](#Setup)
3. [Training the XGBoost model](#Training-the-XGBoost-model)
4. [Deploying the XGBoost endpoint](#Deploying-the-XGBoost-endpoint)
---
## Introduction
This notebook shows how you can configure the SageMaker XGBoost model server by defining the following three functions in the Python source file you pass to the XGBoost constructor in the SageMaker Python SDK:
- `input_fn`: Takes request data and deserializes the data into an object for prediction,
- `predict_fn`: Takes the deserialized request object and performs inference against the loaded model, and
- `output_fn`: Takes the result of prediction and serializes this according to the response content type.
We will write a customized inference script that is designed to illustrate how [SHAP](https://github.com/slundberg/shap) values enable the interpretion of XGBoost models.
We use the [Abalone data](https://www.csie.ntu.edu.tw/~cjlin/libsvmtools/datasets/regression.html), originally from the [UCI data repository](https://archive.ics.uci.edu/ml/datasets/abalone). More details about the original dataset can be found [here](https://archive.ics.uci.edu/ml/machine-learning-databases/abalone/abalone.names). In this libsvm converted version, the nominal feature (Male/Female/Infant) has been converted into a real valued feature as required by XGBoost. Age of abalone is to be predicted from eight physical measurements.
This notebook uses the Abalone dataset to deploy a model server that returns SHAP values, which enable us to create model explanation such as the following plots that show each features contributing to push the model output from the base value.
<table><tr>
<td> <img src="images/shap_young_abalone.png" alt="Drawing"/> </td>
<td> <img src="images/shap_old_abalone.png" alt="Drawing"/> </td>
</tr></table>
---
## Setup
This notebook was created and tested on an ml.m5.2xlarge notebook instance.
Let's start by specifying:
1. The S3 bucket and prefix that you want to use for training and model data. This should be within the same region as the Notebook Instance, training, and hosting.
2. The IAM role arn used to give training and hosting access to your data. See the documentation for how to create these. Note, if more than one role is required for notebook instances, training, and/or hosting, please replace the boto regex with a the appropriate full IAM role arn string(s).
```
%%time
import io
import os
import boto3
import sagemaker
import time
import urllib
role = sagemaker.get_execution_role()
region = boto3.Session().region_name
# S3 bucket for saving code and model artifacts.
# Feel free to specify a different bucket here if you wish.
bucket = sagemaker.Session().default_bucket()
prefix = 'sagemaker/DEMO-xgboost-inference-script-mode'
```
### Fetching the dataset
The following methods download the Abalone dataset and upload files to S3.
```
%%time
# Load the dataset
FILE_DATA = 'abalone'
urllib.request.urlretrieve("https://www.csie.ntu.edu.tw/~cjlin/libsvmtools/datasets/regression/abalone", FILE_DATA)
sagemaker.Session().upload_data(FILE_DATA, bucket=bucket, key_prefix=prefix+'/train')
```
## Training the XGBoost model
SageMaker can now run an XGboost script using the XGBoost estimator. A typical training script loads data from the input channels, configures training with hyperparameters, trains a model, and saves a model to `model_dir` so that it can be hosted later. In this notebook, we use the same training script [abalone.py](https://github.com/awslabs/amazon-sagemaker-examples/blob/master/introduction_to_amazon_algorithms/xgboost_abalone/abalone.py) from [Regression with Amazon SageMaker XGBoost algorithm](https://github.com/awslabs/amazon-sagemaker-examples/blob/master/introduction_to_amazon_algorithms/xgboost_abalone/xgboost_abalone_dist_script_mode.ipynb). Refer to [Regression with Amazon SageMaker XGBoost algorithm](https://github.com/awslabs/amazon-sagemaker-examples/blob/master/introduction_to_amazon_algorithms/xgboost_abalone/xgboost_abalone_dist_script_mode.ipynb) for details on the training script.
After setting training parameters, we kick off training, and poll for status until training is completed, which in this example, takes between few minutes.
To run our training script on SageMaker, we construct a `sagemaker.xgboost.estimator.XGBoost` estimator, which accepts several constructor arguments:
* __entry_point__: The path to the Python script SageMaker runs for training and prediction.
* __role__: Role ARN
* __framework_version__: SageMaker XGBoost version you want to use for executing your model training code, e.g., `0.90-1`, `0.90-2`, or `1.0-1`.
* __train_instance_type__ *(optional)*: The type of SageMaker instances for training. __Note__: Because Scikit-learn does not natively support GPU training, Sagemaker Scikit-learn does not currently support training on GPU instance types.
* __sagemaker_session__ *(optional)*: The session used to train on Sagemaker.
* __hyperparameters__ *(optional)*: A dictionary passed to the train function as hyperparameters.
```
from sagemaker.session import s3_input
from sagemaker.xgboost.estimator import XGBoost
job_name = 'DEMO-xgboost-inference-script-mode-' + time.strftime("%Y-%m-%d-%H-%M-%S", time.gmtime())
print("Training job", job_name)
hyperparameters = {
"max_depth":"5",
"eta":"0.2",
"gamma":"4",
"min_child_weight":"6",
"subsample":"0.7",
"silent":"0",
"objective":"reg:squarederror",
"num_round":"50",
}
instance_type = "ml.c5.xlarge"
output_path = "s3://{}/{}/{}/output".format(bucket, prefix, "abalone-xgb")
xgb_script_mode_estimator = XGBoost(
entry_point="abalone.py",
hyperparameters=hyperparameters,
role=role,
train_instance_count=1,
train_instance_type=instance_type,
framework_version="1.0-1",
output_path="s3://{}/{}/{}/output".format(bucket, prefix, "xgboost-inference-script-mode"),
)
content_type = "text/libsvm"
train_input = s3_input("s3://{}/{}/{}/".format(bucket, prefix, "train"), content_type=content_type)
```
### Train XGBoost Estimator on Abalone Data
Training is as simple as calling `fit` on the Estimator. This will start a SageMaker Training job that will download the data, invoke the entry point code (in the provided script file), and save any model artifacts that the script creates. In this case, the script requires a `train` and a `validation` channel. Since we only created a `train` channel, we re-use it for validation.
```
xgb_script_mode_estimator.fit({'train': train_input, 'validation': train_input}, job_name=job_name)
```
## Deploying the XGBoost endpoint
After training, we can host the newly created model in SageMaker, and create an Amazon SageMaker endpoint – a hosted and managed prediction service that we can use to perform inference. If you call `deploy` after you call `fit` on an XGBoost estimator, it will create a SageMaker endpoint using the training script (i.e., `entry_point`). You can also optionally specify other functions to customize the behavior of deserialization of the input request (`input_fn()`), serialization of the predictions (`output_fn()`), and how predictions are made (`predict_fn()`). If any of these functions are not specified, the endpoint will use the default functions in the SageMaker XGBoost container. See the [SageMaker Python SDK documentation](https://sagemaker.readthedocs.io/en/stable/frameworks/xgboost/using_xgboost.html#sagemaker-xgboost-model-server) for details.
In this notebook, we will run a separate inference script and customize the endpoint to return [SHAP](https://github.com/slundberg/shap) values in addition to predictions. The inference script that we will run in this notebook is provided as the accompanying file (`inference.py`) and also shown below:
```python
import json
import os
import pickle as pkl
import numpy as np
import sagemaker_xgboost_container.encoder as xgb_encoders
def model_fn(model_dir):
"""
Deserialize and return fitted model.
"""
model_file = "xgboost-model"
booster = pkl.load(open(os.path.join(model_dir, model_file), "rb"))
return booster
def input_fn(request_body, request_content_type):
"""
The SageMaker XGBoost model server receives the request data body and the content type,
and invokes the `input_fn`.
Return a DMatrix (an object that can be passed to predict_fn).
"""
if request_content_type == "text/libsvm":
return xgb_encoders.libsvm_to_dmatrix(request_body)
else:
raise ValueError(
"Content type {} is not supported.".format(request_content_type)
)
def predict_fn(input_data, model):
"""
SageMaker XGBoost model server invokes `predict_fn` on the return value of `input_fn`.
Return a two-dimensional NumPy array where the first columns are predictions
and the remaining columns are the feature contributions (SHAP values) for that prediction.
"""
prediction = model.predict(input_data)
feature_contribs = model.predict(input_data, pred_contribs=True)
output = np.hstack((prediction[:, np.newaxis], feature_contribs))
return output
def output_fn(prediction, content_type):
"""
After invoking predict_fn, the model server invokes `output_fn`.
"""
if content_type == "application/json":
return json.dumps(prediction.tolist())
else:
raise ValueError("Content type {} is not supported.".format(content_type))
```
### transform_fn
If you would rather not structure your code around the three methods described above, you can instead define your own `transform_fn` to handle inference requests. An error is thrown if a `transform_fn` is present in conjunction with any `input_fn`, `predict_fn`, and/or `output_fn`. In our case, the `transform_fn` would look as follows:
```python
def transform_fn(model, request_body, content_type, accept_type):
dmatrix = xgb_encoders.libsvm_to_dmatrix(request_body)
prediction = model.predict(dmatrix)
feature_contribs = model.predict(dmatrix, pred_contribs=True)
output = np.hstack((prediction[:, np.newaxis], feature_contribs))
return json.dumps(output.tolist())
```
where `model` is the model object loaded by `model_fn`, `request_body` is the data from the inference request, `content_type` is the content type of the request, and `accept_type` is the request content type for the response.
### Deploy to an endpoint
Since the inference script is separate from the training script, here we use `XGBoostModel` to create a model from s3 artifacts and specify `inference.py` as the `entry_point`.
```
from sagemaker.xgboost.model import XGBoostModel
model_data = xgb_script_mode_estimator.model_data
xgb_inference_model = XGBoostModel(
model_data=model_data,
role=role,
entry_point="inference.py",
framework_version="1.0-1",
)
from sagemaker.predictor import json_deserializer
predictor = xgb_inference_model.deploy(
initial_instance_count=1,
instance_type="ml.c5.xlarge",
)
predictor.serializer = str
predictor.content_type = "text/libsvm"
predictor.deserializer = json_deserializer
predictor.accept = "application/json"
```
### Explain the model's predictions on each data point
```
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
def plot_feature_contributions(prediction):
attribute_names = [
"Sex", # nominal / -- / M, F, and I (infant)
"Length", # continuous / mm / Longest shell measurement
"Diameter", # continuous / mm / perpendicular to length
"Height", # continuous / mm / with meat in shell
"Whole weight", # continuous / grams / whole abalone
"Shucked weight", # continuous / grams / weight of meat
"Viscera weight", # continuous / grams / gut weight (after bleeding)
"Shell weight", # continuous / grams / after being dried
]
prediction, _, *shap_values, bias = prediction
if len(shap_values) != len(attribute_names):
raise ValueError("Length mismatch between shap values and attribute names.")
df = pd.DataFrame(data=[shap_values], index=["SHAP"], columns=attribute_names).T
df.sort_values(by="SHAP", inplace=True)
df["bar_start"] = bias + df.SHAP.cumsum().shift().fillna(0.0)
df["bar_end"] = df.bar_start + df.SHAP
df[["bar_start", "bar_end"]] = np.sort(df[["bar_start", "bar_end"]].values)
df["hue"] = df.SHAP.apply(lambda x: 0 if x > 0 else 1)
sns.set(style="white")
ax1 = sns.barplot(x=df.bar_end, y=df.index, data=df, orient="h", palette="vlag")
for idx, patch in enumerate(ax1.patches):
x_val = patch.get_x() + patch.get_width() + 0.8
y_val = patch.get_y() + patch.get_height() / 2
shap_value = df.SHAP.values[idx]
value = "{0}{1:.2f}".format("+" if shap_value > 0 else "-", shap_value)
ax1.annotate(value, (x_val, y_val), ha="right", va="center")
ax2 = sns.barplot(x=df.bar_start, y=df.index, data=df, orient="h", color="#FFFFFF")
ax2.set_xlim(
df[["bar_start", "bar_end"]].values.min() - 1,
df[["bar_start", "bar_end"]].values.max() + 1
)
ax2.axvline(x=bias, color="#000000", alpha=0.2, linestyle="--", linewidth=1)
ax2.set_title("base value: {0:.1f} → model output: {1:.1f}".format(bias, prediction))
ax2.set_xlabel("Abalone age")
sns.despine(left=True, bottom=True)
plt.tight_layout()
plt.show()
def predict_and_plot(predictor, libsvm_str):
label, *features = libsvm_str.strip().split()
prediction = predictor.predict(" ".join(["-99"] + features)) # use dummy label -99
plot_feature_contributions(prediction[0])
```
The below figure shows features each contributing to push the model output from the base value (9.9 rings) to the model output (6.9 rings). The primary indicator for a young abalone according to the model is low shell weight, which decreases the prediction by 3.0 rings from the base value of 9.9 rings. Whole weight and shucked weight are also powerful indicators. The whole weight pushes the prediction lower by 0.84 rings, while shucked weight pushes the prediction higher by 1.6 rings.
```
a_young_abalone = "6 1:3 2:0.37 3:0.29 4:0.095 5:0.249 6:0.1045 7:0.058 8:0.067"
predict_and_plot(predictor, a_young_abalone)
```
The second example shows feature contributions for another sample, an old abalone. We again see that the primary indicator for the age of abalone according to the model is shell weight, which increases the model prediction by 2.36 rings. Whole weight and shucked weight also contribute significantly, and they both push the model's prediction higher.
```
an_old_abalone = "15 1:1 2:0.655 3:0.53 4:0.175 5:1.2635 6:0.486 7:0.2635 8:0.415"
predict_and_plot(predictor, an_old_abalone)
```
### (Optional) Delete the Endpoint
If you're done with this exercise, please run the `delete_endpoint` line in the cell below. This will remove the hosted endpoint and avoid any charges from a stray instance being left on.
```
predictor.delete_endpoint()
```
| github_jupyter |
# Simple example
To get started with Logomaker, we begin by importing some useful python packages and Logomaker
```
# useful imports
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
%matplotlib inline
plt.ion()
import logomaker as lm
```
We now load an energy matrix for the transcription factor CRP into Logomaker using the built-in method [get_example_matrix](https://logomaker.readthedocs.io/en/latest/examples_src.html#get-example-matrix).
```
# logomaker method to load example matrices
crp_df = -lm.get_example_matrix('crp_energy_matrix')
crp_df.head()
```
These data are from [1](#sort_seq). To draw a logo, logomaker expects a pandas dataframe as input to its [Logo](https://logomaker.readthedocs.io/en/latest/Logo.html) class. The columns of this dataframe represent characters, rows represent positions, and values represent character heights conveying some type of information about their biological importance. Thus, we draw the logo as follows:
```
logo = lm.Logo(crp_df, font_name = 'Arial Rounded MT Bold')
```
# Basic styling
This section introduces basic styling options logomaker provides, applied to the CRP logo from the previous section. We still use the [Logo](https://logomaker.readthedocs.io/en/latest/Logo.html) class (specifically, its constructor) to draw the logo, however we now set a few addtional keyword arguments for styling: the *shade_below* and *fade_below* parameters set the amount of shading and fading to use for characters drawn below the x-axis.
```
# create and style logo
logo = lm.Logo(df=crp_df,
font_name='Arial Rounded MT Bold',
fade_below=0.5,
shade_below=0.5,
figsize=(10,3))
# set axes labels
logo.ax.set_xlabel('Position',fontsize=14)
logo.ax.set_ylabel("$-\Delta \Delta G$ (kcal/mol)", labelpad=-1,fontsize=14)
```
## Change logo font
The font of the characters drawn in the logo can easily be set by using the keyword argument *font_name*. A complete list of available fonts can by accessed by using the logomaker function *list_font_names()*, (e.g. call as *lm.list_font_names()*)
```
# create and style logo
logo = lm.Logo(df=crp_df,
font_name='Hobo Std',
fade_below=0.5,
shade_below=0.5,
figsize=(10,3))
# set axes labels
logo.ax.set_xlabel('Position',fontsize=14)
logo.ax.set_ylabel("$-\Delta \Delta G$ (kcal/mol)", labelpad=-1,fontsize=14)
```
# References
<a id='sort_seq'></a>
[1] Kinney, J. B. et al. (2010). `Using deep sequencing to characterize the biophysical mechanism of a transcriptional regulatory sequence.` Proc Natl Acad Sci USA, 107(20), 9158-9163.
| github_jupyter |
# AI Explanations: Explaining a tabular data model
## Overview
In this tutorial we will perform the following steps:
1. Build and train a Keras model.
1. Export the Keras model as a TF 1 SavedModel and deploy the model on Cloud AI Platform.
1. Compute explainations for our model's predictions using Explainable AI on Cloud AI Platform.
### Dataset
The dataset used for this tutorial was created from a BigQuery Public Dataset: [NYC 2018 Yellow Taxi data](https://console.cloud.google.com/bigquery?filter=solution-type:dataset&q=nyc%20taxi&id=e4902dee-0577-42a0-ac7c-436c04ea50b6&subtask=details&subtaskValue=city-of-new-york%2Fnyc-tlc-trips&project=michaelabel-gcp-training&authuser=1&subtaskIndex=3).
### Objective
The goal is to train a model using the Keras Sequential API that predicts how much a customer is compelled to pay (fares + tolls) for a taxi ride given the pickup location, dropoff location, the day of the week, and the hour of the day.
This tutorial focuses more on deploying the model to AI Explanations than on the design of the model itself. We will be using preprocessed data for this lab. If you wish to know more about the data and how it was preprocessed please see this [notebook](https://github.com/GoogleCloudPlatform/training-data-analyst/blob/master/courses/machine_learning/deepdive/01_bigquery/c_extract_and_benchmark.ipynb).
## Before you begin
This notebook was written with running in **Google Colabratory** in mind. The notebook will run on **Cloud AI Platform Notebooks** or your local environment if the proper packages are installed.
Make sure you're running this notebook in a **GPU runtime** if you have that option. In Colab, select **Runtime** --> **Change runtime type** and select **GPU** for **Hardward Accelerator**.
### Authenticate your GCP account
**If you are using AI Platform Notebooks**, your environment is already
authenticated. You should skip this step.
**Be sure to change the `PROJECT_ID` below to your project before running the cell!**
```
import os
PROJECT_ID = "michaelabel-gcp-training"
os.environ["PROJECT_ID"] = PROJECT_ID
```
**If you are using Colab**, run the cell below and follow the instructions
when prompted to authenticate your account via oAuth. Ignore the error message related to `tensorflow-serving-api`.
```
import sys
import warnings
warnings.filterwarnings('ignore')
os.environ['TF_CPP_MIN_LOG_LEVEL'] = '3'
# If you are running this notebook in Colab, follow the
# instructions to authenticate your GCP account. This provides access to your
# Cloud Storage bucket and lets you submit training jobs and prediction
# requests.
if 'google.colab' in sys.modules:
from google.colab import auth as google_auth
google_auth.authenticate_user()
!pip install witwidget --quiet
!pip install tensorflow==1.15.2 --quiet
!gcloud config set project $PROJECT_ID
elif "DL_PATH" in os.environ:
!sudo pip install tabulate --quiet
```
### Create a Cloud Storage bucket
**The following steps are required, regardless of your notebook environment.**
When you submit a training job using the Cloud SDK, you upload a Python package
containing your training code to a Cloud Storage bucket. AI Platform runs
the code from this package. In this tutorial, AI Platform also saves the
trained model that results from your job in the same bucket. You can then
create an AI Platform model version based on this output in order to serve
online predictions.
**Set the name of your Cloud Storage bucket below. It must be unique across all
Cloud Storage buckets.**
You may also change the `REGION` variable, which is used for operations
throughout the rest of this notebook. Make sure to [choose a region where Cloud
AI Platform services are
available](https://cloud.google.com/ml-engine/docs/tensorflow/regions). Note that you may
not use a Multi-Regional Storage bucket for training with AI Platform.
```
BUCKET_NAME = "michaelabel-gcp-training-ml"
REGION = "us-central1"
os.environ['BUCKET_NAME'] = BUCKET_NAME
os.environ['REGION'] = REGION
```
Run the following cell to create your Cloud Storage bucket if it does not already exist.
```
%%bash
exists=$(gsutil ls -d | grep -w gs://${BUCKET_NAME}/)
if [ -n "$exists" ]; then
echo -e "Bucket gs://${BUCKET_NAME} already exists."
else
echo "Creating a new GCS bucket."
gsutil mb -l ${REGION} gs://${BUCKET_NAME}
echo -e "\nHere are your current buckets:"
gsutil ls
fi
```
### Import libraries for creating model
Import the libraries we'll be using in this tutorial. **This tutorial has been tested with TensorFlow 1.15.2.**
```
%tensorflow_version 1.x
import tensorflow as tf
import tensorflow.feature_column as fc
import pandas as pd
import numpy as np
import json
import time
# Should be 1.15.2
print(tf.__version__)
```
## Downloading and preprocessing data
In this section you'll download the data to train and evaluate your model from a public GCS bucket. The original data has been preprocessed from the public BigQuery dataset linked above.
```
%%bash
# Copy the data to your notebook instance
mkdir taxi_preproc
gsutil cp -r gs://cloud-training/bootcamps/serverlessml/taxi_preproc/*_xai.csv ./taxi_preproc
ls -l taxi_preproc
```
### Read the data with Pandas
We'll use Pandas to read the training and validation data into a `DataFrame`. We will only use the first 7 columns of the csv files for our models.
```
CSV_COLUMNS = ['fare_amount', 'dayofweek', 'hourofday', 'pickuplon',
'pickuplat', 'dropofflon', 'dropofflat']
DAYS = ['Sun', 'Mon', 'Tue', 'Wed', 'Thu', 'Fri', 'Sat']
DTYPES = ['float32', 'str' , 'int32', 'float32' , 'float32' , 'float32' , 'float32' ]
def prepare_data(file_path):
df = pd.read_csv(file_path, usecols = range(7), names = CSV_COLUMNS,
dtype = dict(zip(CSV_COLUMNS, DTYPES)), skiprows=1)
labels = df['fare_amount']
df = df.drop(columns=['fare_amount'])
df['dayofweek'] = df['dayofweek'].map(dict(zip(DAYS, range(7)))).astype('float32')
return df, labels
train_data, train_labels = prepare_data('./taxi_preproc/train_xai.csv')
valid_data, valid_labels = prepare_data('./taxi_preproc/valid_xai.csv')
# Preview the first 5 rows of training data
train_data.head()
```
## Build, train, and evaluate our model with Keras
We'll use `tf.Keras` to build a our ML model that takes our features as input and predicts the fare amount.
But first, we will do some feature engineering. We will be utilizing `tf.feature_column` and `tf.keras.layers.Lambda` to implement our feature engineering in the model graph to simplify our `serving_input_fn` later.
```
# Create functions to compute engineered features in later Lambda layers
def euclidean(params):
lat1, lon1, lat2, lon2 = params
londiff = lon2 - lon1
latdiff = lat2 - lat1
return tf.sqrt(londiff*londiff + latdiff*latdiff)
NUMERIC_COLS = ['pickuplon', 'pickuplat', 'dropofflon', 'dropofflat', 'hourofday', 'dayofweek']
def transform(inputs):
transformed = inputs.copy()
transformed['euclidean'] = tf.keras.layers.Lambda(euclidean, name='euclidean')([
inputs['pickuplat'],
inputs['pickuplon'],
inputs['dropofflat'],
inputs['dropofflon']])
feat_cols = {colname: fc.numeric_column(colname)
for colname in NUMERIC_COLS}
feat_cols['euclidean'] = fc.numeric_column('euclidean')
print("BEFORE TRANSFORMATION")
print("INPUTS:", inputs.keys())
print("AFTER TRANSFORMATION")
print("TRANSFORMED:", transformed.keys())
print("FEATURES", feat_cols.keys())
return transformed, feat_cols
def build_model():
raw_inputs = {
colname : tf.keras.layers.Input(name=colname, shape=(), dtype='float32')
for colname in NUMERIC_COLS
}
transformed, feat_cols = transform(raw_inputs)
dense_inputs = tf.keras.layers.DenseFeatures(feat_cols.values(),
name = 'dense_input')(transformed)
h1 = tf.keras.layers.Dense(64, activation='relu', name='h1')(dense_inputs)
h2 = tf.keras.layers.Dense(32, activation='relu', name='h2')(h1)
output = tf.keras.layers.Dense(1, activation='linear', name = 'output')(h2)
model = tf.keras.models.Model(raw_inputs, output)
return model
model = build_model()
model.summary()
# Compile the model and see a summary
optimizer = tf.keras.optimizers.Adam(0.001)
model.compile(loss='mean_squared_error', optimizer=optimizer,
metrics = [tf.keras.metrics.RootMeanSquaredError()])
tf.keras.utils.plot_model(model, to_file='model_plot.png', show_shapes=True,
show_layer_names=True, rankdir="TB")
```
### Create an input data pipeline with tf.data
Per best practices, we will use `tf.Data` to create our input data pipeline. Our data is all in an in-memory dataframe, so we will use `tf.data.Dataset.from_tensor_slices` to create our pipeline.
```
def load_dataset(features, labels, mode):
dataset = tf.data.Dataset.from_tensor_slices(({"dayofweek" : features["dayofweek"],
"hourofday" : features["hourofday"],
"pickuplat" : features["pickuplat"],
"pickuplon" : features["pickuplon"],
"dropofflat" : features["dropofflat"],
"dropofflon" : features["dropofflon"]},
labels
))
if mode == tf.estimator.ModeKeys.TRAIN:
dataset = dataset.repeat().batch(256).shuffle(256*10)
else:
dataset = dataset.batch(256)
return dataset.prefetch(1)
train_dataset = load_dataset(train_data, train_labels, tf.estimator.ModeKeys.TRAIN)
valid_dataset = load_dataset(valid_data, valid_labels, tf.estimator.ModeKeys.EVAL)
```
### Train the model
Now we train the model. We will specify a number of epochs which to train the model and tell the model how many steps to expect per epoch.
```
tf.keras.backend.get_session().run(tf.tables_initializer(name='init_all_tables'))
steps_per_epoch = 426433 // 256
model.fit(train_dataset, steps_per_epoch=steps_per_epoch, validation_data=valid_dataset, epochs=10)
# Send test instances to model for prediction
predict = model.predict(valid_dataset, steps = 1)
predict[:5]
```
## Export the model as a TF 1 SavedModel
In order to deploy our model in a format compatible with AI Explanations, we'll follow the steps below to convert our Keras model to a TF Estimator, and then use the `export_saved_model` method to generate the SavedModel and save it in GCS.
```
## Convert our Keras model to an estimator
keras_estimator = tf.keras.estimator.model_to_estimator(keras_model=model, model_dir='export')
print(model.input)
# We need this serving input function to export our model in the next cell
serving_fn = tf.estimator.export.build_raw_serving_input_receiver_fn(
model.input
)
export_path = keras_estimator.export_saved_model(
'gs://' + BUCKET_NAME + '/explanations',
serving_input_receiver_fn=serving_fn
).decode('utf-8')
```
Use TensorFlow's `saved_model_cli` to inspect the model's SignatureDef. We'll use this information when we deploy our model to AI Explanations in the next section.
```
!saved_model_cli show --dir $export_path --all
```
## Deploy the model to AI Explanations
In order to deploy the model to Explanations, we need to generate an `explanations_metadata.json` file and upload this to the Cloud Storage bucket with our SavedModel. Then we'll deploy the model using `gcloud`.
### Prepare explanation metadata
We need to tell AI Explanations the names of the input and output tensors our model is expecting, which we print below.
The value for `input_baselines` tells the explanations service what the baseline input should be for our model. Here we're using the median for all of our input features. That means the baseline prediction for this model will be the fare our model predicts for the median of each feature in our dataset.
```
# Print the names of our tensors
print('Model input tensors: ', model.input)
print('Model output tensor: ', model.output.name)
baselines_med = train_data.median().values.tolist()
baselines_mode = train_data.mode().values.tolist()
print(baselines_med)
print(baselines_mode)
explanation_metadata = {
"inputs": {
"dayofweek": {
"input_tensor_name": "dayofweek:0",
"input_baselines": [baselines_mode[0][0]] # Thursday
},
"hourofday": {
"input_tensor_name": "hourofday:0",
"input_baselines": [baselines_mode[0][1]] # 8pm
},
"dropofflon": {
"input_tensor_name": "dropofflon:0",
"input_baselines": [baselines_med[4]]
},
"dropofflat": {
"input_tensor_name": "dropofflat:0",
"input_baselines": [baselines_med[5]]
},
"pickuplon": {
"input_tensor_name": "pickuplon:0",
"input_baselines": [baselines_med[2]]
},
"pickuplat": {
"input_tensor_name": "pickuplat:0",
"input_baselines": [baselines_med[3]]
},
},
"outputs": {
"dense": {
"output_tensor_name": "output/BiasAdd:0"
}
},
"framework": "tensorflow"
}
print(explanation_metadata)
```
Since this is a regression model (predicting a numerical value), the baseline prediction will be the same for every example we send to the model. If this were instead a classification model, each class would have a different baseline prediction.
```
# Write the json to a local file
with open('explanation_metadata.json', 'w') as output_file:
json.dump(explanation_metadata, output_file)
!gsutil cp explanation_metadata.json $export_path
```
### Create the model
Now we will create out model on Cloud AI Platform if it does not already exist.
```
MODEL = 'taxifare_explain'
os.environ["MODEL"] = MODEL
%%bash
exists=$(gcloud ai-platform models list | grep ${MODEL})
if [ -n "$exists" ]; then
echo -e "Model ${MODEL} already exists."
else
echo "Creating a new model."
gcloud ai-platform models create ${MODEL}
fi
```
### Create the model version
Creating the version will take ~5-10 minutes. Note that your first deploy may take longer.
```
# Each time you create a version the name should be unique
import datetime
now = datetime.datetime.now().strftime("%Y%m%d%H%M%S")
VERSION_IG = 'v_IG_{}'.format(now)
VERSION_SHAP = 'v_SHAP_{}'.format(now)
# Create the version with gcloud
!gcloud beta ai-platform versions create $VERSION_IG \
--model $MODEL \
--origin $export_path \
--runtime-version 1.15 \
--framework TENSORFLOW \
--python-version 3.7 \
--machine-type n1-standard-4 \
--explanation-method 'integrated-gradients' \
--num-integral-steps 25
!gcloud beta ai-platform versions create $VERSION_SHAP \
--model $MODEL \
--origin $export_path \
--runtime-version 1.15 \
--framework TENSORFLOW \
--python-version 3.7 \
--machine-type n1-standard-4 \
--explanation-method 'sampled-shapley' \
--num-paths 50
# Make sure the model deployed correctly. State should be `READY` in the following log
!gcloud ai-platform versions describe $VERSION_IG --model $MODEL
!echo "---"
!gcloud ai-platform versions describe $VERSION_SHAP --model $MODEL
```
## Getting predictions and explanations on deployed model
Now that your model is deployed, you can use the AI Platform Prediction API to get feature attributions. We'll pass it a single test example here and see which features were most important in the model's prediction. Here we'll use `gcloud` to call our deployed model.
### Format our request for gcloud
To use gcloud to make our AI Explanations request, we need to write the JSON to a file. Our example here is for a ride from the Google office in downtown Manhattan to LaGuardia Airport at 5pm on a Tuesday afternoon.
Note that we had to write our day of the week at "3" instead of "Tue" since we encoded the days of the week outside of our model and serving input function.
```
# Format data for prediction to our model
!rm taxi-data.txt
!touch taxi-data.txt
prediction_json = {"dayofweek": "3", "hourofday": "17", "pickuplon": "-74.0026", "pickuplat": "40.7410", "dropofflat": "40.7790", "dropofflon": "-73.8772"}
with open('taxi-data.txt', 'a') as outfile:
json.dump(prediction_json, outfile)
# Preview the contents of the data file
!cat taxi-data.txt
```
### Making the explain request
Now we make the explaination requests. We will go ahead and do this here for both integrated gradients and SHAP using the prediction JSON from above.
```
resp_obj = !gcloud beta ai-platform explain --model $MODEL --version $VERSION_IG --json-instances='taxi-data.txt'
response_IG = json.loads(resp_obj.s)
resp_obj
resp_obj = !gcloud beta ai-platform explain --model $MODEL --version $VERSION_SHAP --json-instances='taxi-data.txt'
response_SHAP = json.loads(resp_obj.s)
resp_obj
```
### Understanding the explanations response
First let's just look at the difference between our predictions using our baselines and our predicted taxi fare for the example.
```
explanations_IG = response_IG['explanations'][0]['attributions_by_label'][0]
explanations_SHAP = response_SHAP['explanations'][0]['attributions_by_label'][0]
predicted = round(explanations_SHAP['example_score'], 2)
baseline = round(explanations_SHAP['baseline_score'], 2 )
print('Baseline taxi fare: ' + str(baseline) + ' dollars')
print('Predicted taxi fare: ' + str(predicted) + ' dollars')
```
Next let's look at the feature attributions for this particular example. Positive attribution values mean a particular feature pushed our model prediction up by that amount, and vice versa for negative attribution values. Which features seem like they're the most important...well it seems like the location features are the most important!
```
from tabulate import tabulate
feature_names = valid_data.columns.tolist()
attributions_IG = explanations_IG['attributions']
attributions_SHAP = explanations_SHAP['attributions']
rows = []
for feat in feature_names:
rows.append([feat, prediction_json[feat], attributions_IG[feat], attributions_SHAP[feat]])
print(tabulate(rows,headers=['Feature name', 'Feature value', 'Attribution value (IG)', 'Attribution value (SHAP)']))
```
| github_jupyter |
```
!nvidia-smi
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
import seaborn as sns
import os, shutil
from glob import glob
from google.colab import drive
drive.mount('/content/grive')
import cv2
from keras.preprocessing.image import ImageDataGenerator
```
# Preprocessing images
```
image_gen = ImageDataGenerator(rescale=1./255)
train_dir = '/content/grive/MyDrive/Face Mask Dataset/Train'
val_dir = '/content/grive/MyDrive/Face Mask Dataset/Validation'
test_dir = '/content/grive/MyDrive/Face Mask Dataset/Test'
batch_size = 80
train_generator = image_gen.flow_from_directory(train_dir,
target_size=(150,150),
batch_size=batch_size,
seed=42,
shuffle=False,
class_mode='binary')
val_generator = image_gen.flow_from_directory(val_dir,
target_size=(150,150),
batch_size=batch_size,
seed=42,
shuffle=False,
class_mode='binary')
test_generator = image_gen.flow_from_directory(test_dir,
target_size=(150,150),
batch_size=batch_size,
seed=42,
shuffle=False,
class_mode='binary')
```
# Building CNN Model
```
from keras.models import Sequential
from keras.layers import Dense, Conv2D, MaxPool2D, Flatten, Dropout
from keras import optimizers
model = Sequential()
model.add(Conv2D(32, (3,3),input_shape=(150,150,3),activation='relu'))
model.add(MaxPool2D((2,2)))
model.add(Conv2D(64, (3,3),activation='relu'))
model.add(MaxPool2D((2,2)))
model.add(Conv2D(128, (3,3),activation='relu'))
model.add(MaxPool2D((2,2)))
model.add(Conv2D(128, (3,3),activation='relu'))
model.add(MaxPool2D((2,2)))
model.add(Flatten())
model.add(Dense(512, activation='relu'))
model.add(Dense(1, activation='sigmoid'))
model.compile(loss='binary_crossentropy',
optimizer='adam',
metrics=['accuracy'])
model.summary()
result = model.fit_generator(
train_generator,
steps_per_epoch=100,
epochs=30,
validation_data=val_generator,
validation_steps=100
)
model.save('model_large_data.h5')
```
# Evaluation on model
```
result.history['accuracy']
plt.plot(result.history['accuracy'])
model.metrics_names
plt.figure(figsize=(10,4))
plt.subplot(1,2,1)
plt.plot(result.history['accuracy'], label='train accuracy', color='red')
plt.plot(result.history['val_accuracy'], label='validation accuracy', color='blue')
plt.legend()
plt.subplot(1,2,2)
plt.plot(result.history['loss'], label='train loss', color='red')
plt.plot(result.history['val_loss'], label='validation loss', color='blue')
plt.legend()
plt.show()
from sklearn.metrics import confusion_matrix, classification_report
prediction = model.predict_classes(test_generator)
import math
num_of_examples = len(test_generator.filenames)
num_of_generator_calls = math.ceil(num_of_examples/(1.0*80))
test_labels=[]
for i in range(0,int(num_of_generator_calls)):
test_labels.extend(np.array(test_generator[i][1]))
train_generator.class_indices
confusion_matrix(test_labels, prediction)
plt.figure(figsize=(25,7))
plt.subplot(131)
ax = sns.heatmap(confusion_matrix(test_generator.labels, prediction.round()),
annot = True, annot_kws={"size":20}, fmt="d",cmap = "Blues")
cmlabels = ['True Negatives', "False Positives",
'Flase Negatives', "True Positives"]
for i,t in enumerate(ax.texts):
t.set_text(t.get_text() + "\n" + cmlabels[i])
plt.title('Confusion Matrix', size=25)
plt.xlabel('Predicted Outcome', size=20)
plt.ylabel('Actual Outcome', size=20)
labels = ['Mask', 'No Mask']
ax.set_xticklabels(labels, size=15)
ax.set_yticklabels(labels, size=15)
plt.tight_layout()
plt.show();
print(classification_report(test_labels, prediction))
```
| github_jupyter |
## 看下 0-255 模型在具有高斯分布的数据集中的精度
```
import matplotlib.pyplot as plt
import numpy as np
import sklearn
import torch
import torch.nn as nn
import torch.optim as optim
from torch.utils.data import DataLoader, Subset, RandomSampler, BatchSampler
from model.inceptionv4 import inceptionv4
from model.mobilenetv2 import mobilenetv2
from model.resnet import resnet18
from model.shufflenetv2 import shufflenetv2
from model.vgg import vgg9_bn
from s3_dataset import PlantDataSet, PlantDataSetB
import matplotlib.pyplot as plt
def get_pre(net, device, data_loader):
'''
得到整个测试集预测的结果,以及标签
'''
label_all = []
pre_all = []
with torch.no_grad():
net.eval()
for data in data_loader:
images, labels = data
images = images.float().to(device)
labels = labels.long().to(device)
outputs = net(images)
_, predicted = torch.max(outputs.data, 1)
label_all.extend(labels.data.cpu().numpy())
pre_all.extend(predicted.data.cpu().numpy())
return pre_all, label_all
from sklearn.metrics import accuracy_score # 精度
from sklearn.metrics import confusion_matrix # 混淆矩阵
device = torch.device('cuda:0' if torch.cuda.is_available() else 'cpu')
Func = [vgg9_bn, resnet18, shufflenetv2, mobilenetv2, inceptionv4]
Save_path = [
'../model_save/plant_disease2/vgg.pth',
'../model_save/plant_disease2/resnet18.pth',
'../model_save/plant_disease2/shufflenetv2.pth',
'../model_save/plant_disease2/mobilenetv2.pth',
'../model_save/plant_disease2/inceptionv4.pth'
]
data_test_a = DataLoader(PlantDataSet(flag='test'),
batch_size=8,
shuffle=False)
data_test_b = DataLoader(PlantDataSetB(flag='test'),
batch_size=8,
shuffle=False)
for Index in range(5):
# 导入模型和权重
net = Func[Index]()
path_saved_model = Save_path[Index]
net.load_state_dict(torch.load(path_saved_model))
net.to(device)
pre, label = get_pre(net, device, data_test_a)
pre, label = np.array(pre), np.array(label)
print('预测精度为:{:.9f}'.format(accuracy_score(label, pre)))
ans = confusion_matrix(label, pre, labels=list(range(38)))
from sklearn.metrics import classification_report
print(classification_report(label, pre, labels=list(range(38))))
from sklearn.metrics import accuracy_score # 精度
from sklearn.metrics import confusion_matrix # 混淆矩阵
device = torch.device('cuda:1' if torch.cuda.is_available() else 'cpu')
Func = [vgg9_bn, resnet18, shufflenetv2, mobilenetv2, inceptionv4]
Save_path = [
'../model_save/plant_disease_domain/vgg.pth',
'../model_save/plant_disease_domain/resnet18.pth',
'../model_save/plant_disease_domain/shufflenetv2.pth'
]
data_test_a = DataLoader(PlantDataSet(flag='test'),
batch_size=8,
shuffle=False)
data_test_b = DataLoader(PlantDataSetB(flag='test'),
batch_size=8,
shuffle=False)
for Index in range(3):
# 导入模型和权重
net = Func[Index]()
path_saved_model = Save_path[Index]
net.load_state_dict(torch.load(path_saved_model))
net.to(device)
pre, label = get_pre(net, device, data_test_b)
pre, label = np.array(pre), np.array(label)
print('预测精度为:{:.9f}'.format(accuracy_score(label, pre)))
```
| github_jupyter |
#RxId : CSV Data Cleaning
Data Source : https://pillbox.nlm.nih.gov/developers.html
https://dev.socrata.com/foundry/datadiscovery.nlm.nih.gov/crzr-uvwg
Issue : Two CSV files were downloaded from the above site.
Pillbox.NO.ID.csv has useful meds data but no image_id field to link it to a filename in the image library.
Pillbox.NO.IMAGE ID.csv lacks useful meds data but HAS image_id field to link it to a filename in the image library.
Solution: Clean CSV files individually and merge into single CSV that will be used to load an AWS RDS database
### Load CSVs into dataframes
```
import pandas as pd
pd.options.display.max_columns = None
url1="https://raw.githubusercontent.com/labs12-rxid/DS/master/CSV/Pillbox.NO.ID.csv"
df1=pd.read_csv(url1)
url2="https://raw.githubusercontent.com/labs12-rxid/DS/master/CSV/Pillbox.IMAGE%20ID.csv"
df2=pd.read_csv(url2)
print(df1.shape)
df2.shape
df1.columns
```
### Rename df2.id to df2.ID to match df1
```
df2.rename(columns={'id':'ID'}, inplace=True)
df2.columns
```
### Drop Useless/Duplicated Columns from df1 & df2
```
drop_col_1=['created at', 'updated at', 'ndc9', 'author',
'rxstring', 'has_image',
'Unnamed: 53', 'Unnamed: 54', 'Unnamed: 55',
'Unnamed: 56', 'Unnamed: 57', 'Unnamed: 58', 'Unnamed: 59',
'Unnamed: 60', 'Unnamed: 61', 'Unnamed: 62', 'Unnamed: 63',
'Unnamed: 64', 'Unnamed: 65', 'Unnamed: 66', 'Unnamed: 67',
'Unnamed: 68', 'Unnamed: 69', 'Unnamed: 70', 'Unnamed: 71']
df1.drop(columns=drop_col_1, inplace=True)
df1.columns
drop_col_2=['SETID', 'spp', 'INGREDIENTS','SPL_INACTIVE_ING','SPLSIZE', 'SPLSCORE',
'SPLIMPRINT', 'SPLCOLOR', 'SPLSHAPE', 'RXCUI', 'RXTTY', 'IMAGE_SOURCE']
df2.drop(columns=drop_col_2, inplace=True)
df2.columns
```
### compare same ID acrross Frames
```
df1.query('ID==3143')
df2.query('ID==3143')
```
### Combine dataframes
```
df_comb=pd.merge(df1,df2,how='left', on=['ID'])
df_comb.query('ID==3143')
```
### Write out combined CSV
```
# df_comb.to_csv('Pills.Final.csv', index=False ) #header=['id','status_group'])
```
# USING FINAL CSV FILE
```
df = pd.read_csv('Pills.Final.csv')
df.head()
# DATAFRAME with 'image_id' of ROUND observations
df_round = df['image_id'][df['splshape'] == 'C48348'][df['HAS_IMAGE'] == 1]
df_round.shape
# DATAFRAME with 'image_id' of OVAL observations
df_oval = df['image_id'][df['splshape'] == 'C48345'][df['HAS_IMAGE'] == 1]
df_oval.shape
# DATAFRAME with 'image_id' of CAPSULE observations
df_capsule = df['image_id'][df['splshape'] == 'C48336'][df['HAS_IMAGE'] == 1]
df_capsule.shape
# Quick test on CAPSULE files
df_captest = df[['image_id', 'splshape', 'splshape_text']][df['splshape'] == 'C48336'][df['HAS_IMAGE'] == 1]
df_captest.shape
df_captest['splshape_text'].value_counts()
# Quick test on OVAL files
df_ovaltest = df[['image_id', 'splshape', 'splshape_text']][df['splshape'] == 'C48345'][df['HAS_IMAGE'] == 1]
df_ovaltest.shape
df_ovaltest['splshape_text'].value_counts()
len(df_capsule)
```
### Copying ROUND image files to 'round_images' folder
```
# import os
# import shutil
# src = 'C:\\Users\\Gutierrez\\Documents\\DataScience\\lambda_school\\labs12-rxid\\DS\\pillbox_images'
# dest = 'C:\\Users\\Gutierrez\\Documents\\DataScience\\lambda_school\\labs12-rxid\\DS\\round_images'
# src_files = os.listdir(src)
# for file_name in df_round:
# full_file_name = os.path.join(src, file_name + '.jpg')
# if (os.path.isfile(full_file_name)):
# shutil.copy(full_file_name, dest)
pwd
```
### Copying OVAL image files to 'oval_images' folder
```
# import os
# import shutil
# src = 'C:\\Users\\Gutierrez\\Documents\\DataScience\\lambda_school\\labs12-rxid\\DS\\pillbox_images'
# dest = 'C:\\Users\\Gutierrez\\Documents\\DataScience\\lambda_school\\labs12-rxid\\DS\\oval_images'
# src_files = os.listdir(src)
# for file_name in df_oval:
# full_file_name = os.path.join(src, file_name + '.jpg')
# if (os.path.isfile(full_file_name)):
# shutil.copy(full_file_name, dest)
```
### Copying CAPSULE image files to 'capsule_images' folder
```
# import os
# import shutil
# src = 'C:\\Users\\Gutierrez\\Documents\\DataScience\\lambda_school\\labs12-rxid\\DS\\pillbox_images'
# dest = 'C:\\Users\\Gutierrez\\Documents\\DataScience\\lambda_school\\labs12-rxid\\DS\\capsule_images'
# src_files = os.listdir(src)
# for file_name in df_capsule:
# full_file_name = os.path.join(src, file_name + '.jpg')
# if (os.path.isfile(full_file_name)):
# shutil.copy(full_file_name, dest)
```
# GETTING IMAGES INTO `TRAIN` & `TEST` FOLDER
### Moving CAPSULE images to Training & Test folders
```
# import os
# import shutil
# src = 'C:\\Users\\Gutierrez\\Documents\\DataScience\\lambda_school\\labs12-rxid\\DS\\capsule_images'
# dest = 'C:\\Users\\Gutierrez\\Documents\\DataScience\\lambda_school\\labs12-rxid\\DS\\data\\train\\capsule'
# src_files = os.listdir(src)
# ctr = 1
# for file_name in df_capsule:
# if ctr <= 1025:
# full_file_name = os.path.join(src, file_name + '.jpg')
# if (os.path.isfile(full_file_name)):
# shutil.move(full_file_name, dest)
# ctr += 1
df_oval.head()
```
### Moving CAPSULE images to Test folder
```
def img_traintest(shape, df):
"""
Function to get images into Train and Test folders.
1025 for Train and 416 for Test.
shape: Takes in a string for the shape.
df: Takes in Pandas DataFrame/Series with images file names only.
"""
import os
import shutil
src = 'C:\\Users\\Gutierrez\\Documents\\DataScience\\lambda_school\\labs12-rxid\\DS\\' + shape + '_images'
dest1 = 'C:\\Users\\Gutierrez\\Documents\\DataScience\\lambda_school\\labs12-rxid\\DS\\data\\train\\' + shape
dest2 = 'C:\\Users\\Gutierrez\\Documents\\DataScience\\lambda_school\\labs12-rxid\\DS\\data\\test\\' + shape
src_files = os.listdir(src)
ctr1 = 1
ctr2 = 1
# print(src)
# print(dest1)
# print(dest2)
# print(f'df = {df.shape}')
for file_name in df:
# print(f'file_name:\n{file_name}')
if ctr1 <= 1025:
full_file_name = os.path.join(src, file_name + '.jpg')
# print(f'full_file_name:\n{full_file_name}')
if (os.path.isfile(full_file_name)):
shutil.move(full_file_name, dest1)
ctr1 += 1
if ctr2 <= 416:
full_file_name = os.path.join(src, file_name + '.jpg')
if (os.path.isfile(full_file_name)):
shutil.move(full_file_name, dest2)
ctr2 += 1
# img_traintest("capsule", df_capsule)
# img_traintest("round", df_round)
# img_traintest('oval', df_oval)
```
# Splitting images in half vertically
#### ROUND "Train" Folder
```
def image_splitter(shape, train_or_test):
'''
Function to split images in train or test folder according based on shape.
Function takes two strings, one for shape (i.e., round, capsule, oval) and another for either "train" or "test".
'''
import os
import imageio
# partial path for creating new directory and saving files
partial_path = "C:\\Users\\Gutierrez\\Documents\\DataScience\\lambda_school\\labs12-rxid\\DS\\data\\"
# making new folder for image halves
new_folder = partial_path + train_or_test + "\\" + shape + "_h\\"
os.makedirs(new_folder)
# path to loop through
folder_path = partial_path + train_or_test + "\\" + shape
# print(folder_path)
# image counter for image halves maded and saved
img_ctr = 0
# looping through all images in folder and processing
for file in os.listdir(folder_path):
file_name = os.path.join(folder_path, file)
# print(file_name)
img = imageio.imread(file_name)
height, width = img.shape[:2]
# Cut the image in half
width_cutoff = width // 2
s1 = img[:, :width_cutoff]
s2 = img[:, width_cutoff:]
# Save each half
filepath_h1 = new_folder + "h1_" + file
filepath_h2 = new_folder + "h2_" + file
imageio.imwrite(filepath_h1, s1)
imageio.imwrite(filepath_h2, s2)
img_ctr +=2
print('Total images (halves) saved:', img_ctr)
# image_splitter('round', 'train')
# image_splitter('round', 'test')
```
# Splitting images in half horizontally
```
def image_hsplitter(shape, train_or_test):
'''
Function to split images in train or test folder according based on shape.
Function takes two strings, one for shape (i.e., round, capsule, oval) and another for either "train" or "test".
'''
import os
import imageio
# partial path for creating new directory and saving files
partial_path = "C:\\Users\\Gutierrez\\Documents\\DataScience\\lambda_school\\labs12-rxid\\DS\\data\\"
# making new folder for image halves
new_folder = partial_path + train_or_test + "\\" + shape + "_h\\"
os.makedirs(new_folder)
# path to loop through
folder_path = partial_path + train_or_test + "\\" + shape
# print(folder_path)
# image counter for image halves maded and saved
img_ctr = 0
# looping through all images in folder and processing
for file in os.listdir(folder_path):
file_name = os.path.join(folder_path, file)
# print(file_name)
img = imageio.imread(file_name)
height, width = img.shape[:2]
# Cut the image in half horizontally
height_cutoff = height // 2
s1 = img[:height_cutoff, :]
s2 = img[height_cutoff:, :]
# Save each half
filepath_h1 = new_folder + "h1_" + file
filepath_h2 = new_folder + "h2_" + file
imageio.imwrite(filepath_h1, s1)
imageio.imwrite(filepath_h2, s2)
img_ctr +=2
print('Total images (halves) saved:', img_ctr)
```
#### CAPSULE "Train" Folder
```
image_hsplitter('capsule', 'train')
```
#### CAPSULE "Test" Folder
```
image_hsplitter('capsule', 'test')
```
# Renaming files for Neural Network
```
# Need images labeled like "cat.1" or "dog.3" and so on
# You could then use it in your example like this:
# rename(r'c:\temp\xx', r'*.doc', r'new(%s)')
# The above example will convert all *.doc files in c:\temp\xx dir to new(%s).doc,
# where %s is the previous base name of the file (without extension).
import glob, os
def rename_files(dir, pattern, titlePattern):
ctr = 1
for pathAndFilename in glob.iglob(os.path.join(dir, pattern)):
title, ext = os.path.splitext(os.path.basename(pathAndFilename))
os.rename(pathAndFilename,
os.path.join(dir, titlePattern % ctr + ext))
ctr += 1
```
#### Train images
```
# rename_files(r"C:\\Users\\Gutierrez\\Documents\\DataScience\\lambda_school\\labs12-rxid\\DS\\data\\train\\round_h",
# r"*.jpg",
# r"rnd.%s")
# rename_files(r"C:\\Users\\Gutierrez\\Documents\\DataScience\\lambda_school\\labs12-rxid\\DS\\data\\train\\capsule_h",
# r"*.jpg",
# r"cap.%s")
```
#### Test images
```
# rename_files(r"C:\\Users\\Gutierrez\\Documents\\DataScience\\lambda_school\\labs12-rxid\\DS\\data\\test\\round_h",
# r"*.jpg",
# r"rnd.%s")
# rename_files(r"C:\\Users\\Gutierrez\\Documents\\DataScience\\lambda_school\\labs12-rxid\\DS\\data\\test\\capsule_h",
# r"*.jpg",
# r"cap.%s")
```
# CHECKING FOR `COLOR` & `SHAPE`
```
df[df['splcolor_text'] == 'C48329']
df[df['product_code'] == '897714']
df[['spl_ingredients','splimprint', 'splcolor_text', 'splcolor', 'splshape_text', 'splshape', 'rxcui']][df['rxcui'].str.contains('Verapamil', na=False)]
df[['splcolor_text', 'splcolor']][df['splcolor']=='C48329']
df[df[('HAS_IMAGE' == 1)]]
df[(df['HAS_IMAGE'] == 1)].splshape_text.value_counts()
df[(df['HAS_IMAGE'] == 1)].splshape_text.value_counts().sum()
```
| github_jupyter |
# Regression Week 3: Assessing Fit (polynomial regression)
In this notebook you will compare different regression models in order to assess which model fits best. We will be using polynomial regression as a means to examine this topic. In particular you will:
* Write a function to take an SArray and a degree and return an SFrame where each column is the SArray to a polynomial value up to the total degree e.g. degree = 3 then column 1 is the SArray column 2 is the SArray squared and column 3 is the SArray cubed
* Use matplotlib to visualize polynomial regressions
* Use matplotlib to visualize the same polynomial degree on different subsets of the data
* Use a validation set to select a polynomial degree
* Assess the final fit using test data
We will continue to use the House data from previous notebooks.
# Fire up graphlab create
```
import graphlab
```
Next we're going to write a polynomial function that takes an SArray and a maximal degree and returns an SFrame with columns containing the SArray to all the powers up to the maximal degree.
The easiest way to apply a power to an SArray is to use the .apply() and lambda x: functions.
For example to take the example array and compute the third power we can do as follows: (note running this cell the first time may take longer than expected since it loads graphlab)
```
tmp = graphlab.SArray([1., 2., 3.])
tmp_cubed = tmp.apply(lambda x: x**3)
print tmp
print tmp_cubed
```
We can create an empty SFrame using graphlab.SFrame() and then add any columns to it with ex_sframe['column_name'] = value. For example we create an empty SFrame and make the column 'power_1' to be the first power of tmp (i.e. tmp itself).
```
ex_sframe = graphlab.SFrame()
ex_sframe['power_1'] = tmp
print ex_sframe
```
# Polynomial_sframe function
Using the hints above complete the following function to create an SFrame consisting of the powers of an SArray up to a specific degree:
```
def polynomial_sframe(feature, degree):
# assume that degree >= 1
# initialize the SFrame:
poly_sframe = graphlab.SFrame()
# and set poly_sframe['power_1'] equal to the passed feature
poly_sframe['power_1'] = feature
# first check if degree > 1
if degree > 1:
# then loop over the remaining degrees:
# range usually starts at 0 and stops at the endpoint-1. We want it to start at 2 and stop at degree
for power in range(2, degree+1):
# first we'll give the column a name:
name = 'power_' + str(power)
# then assign poly_sframe[name] to the appropriate power of feature
poly_sframe[name] = feature**power
return poly_sframe
```
To test your function consider the smaller tmp variable and what you would expect the outcome of the following call:
```
print polynomial_sframe(tmp, 3)
```
# Visualizing polynomial regression
Let's use matplotlib to visualize what a polynomial regression looks like on some real data.
```
sales = graphlab.SFrame('kc_house_data.gl/kc_house_data.gl')
```
As in Week 3, we will use the sqft_living variable. For plotting purposes (connecting the dots), you'll need to sort by the values of sqft_living. For houses with identical square footage, we break the tie by their prices.
```
sales = sales.sort(['sqft_living', 'price'])
```
Let's start with a degree 1 polynomial using 'sqft_living' (i.e. a line) to predict 'price' and plot what it looks like.
```
poly1_data = polynomial_sframe(sales['sqft_living'], 1)
poly1_data['price'] = sales['price'] # add price to the data since it's the target
```
NOTE: for all the models in this notebook use validation_set = None to ensure that all results are consistent across users.
```
model1 = graphlab.linear_regression.create(poly1_data, target = 'price', features = ['power_1'], validation_set = None)
#let's take a look at the weights before we plot
model1.get("coefficients")
import matplotlib.pyplot as plt
%matplotlib inline
plt.plot(poly1_data['power_1'],poly1_data['price'],'.',
poly1_data['power_1'], model1.predict(poly1_data),'-')
```
Let's unpack that plt.plot() command. The first pair of SArrays we passed are the 1st power of sqft and the actual price we then ask it to print these as dots '.'. The next pair we pass is the 1st power of sqft and the predicted values from the linear model. We ask these to be plotted as a line '-'.
We can see, not surprisingly, that the predicted values all fall on a line, specifically the one with slope 280 and intercept -43579. What if we wanted to plot a second degree polynomial?
```
poly2_data = polynomial_sframe(sales['sqft_living'], 2)
my_features = poly2_data.column_names() # get the name of the features
poly2_data['price'] = sales['price'] # add price to the data since it's the target
model2 = graphlab.linear_regression.create(poly2_data, target = 'price', features = my_features, validation_set = None)
model2.get("coefficients")
plt.plot(poly2_data['power_1'],poly2_data['price'],'.',
poly2_data['power_1'], model2.predict(poly2_data),'-')
```
The resulting model looks like half a parabola. Try on your own to see what the cubic looks like:
```
poly1_data = polynomial_sframe(sales['sqft_living'], 3)
poly1_data['price'] = sales['price'] # add price to the data since it's the target
model2 = graphlab.linear_regression.create(poly1_data, target = 'price', features = ['power_2'], validation_set = None)
plt.plot(poly1_data['power_2'],poly1_data['price'],'.',
poly1_data['power_2'], model2.predict(poly1_data),'-')
model3 = graphlab.linear_regression.create(poly1_data, target = 'price', features = ['power_3'], validation_set = None)
plt.plot(poly1_data['power_3'],poly1_data['price'],'.',
poly1_data['power_3'], model3.predict(poly1_data),'-')
```
Now try a 15th degree polynomial:
```
poly1_data = polynomial_sframe(sales['sqft_living'], 15)
poly1_data['price'] = sales['price'] # add price to the data since it's the target
model2 = graphlab.linear_regression.create(poly1_data, target = 'price', features = ['power_15'], validation_set = None)
plt.plot(poly1_data['power_15'],poly1_data['price'],'.',
poly1_data['power_15'], model2.predict(poly1_data),'-')
```
What do you think of the 15th degree polynomial? Do you think this is appropriate? If we were to change the data do you think you'd get pretty much the same curve? Let's take a look.
# Changing the data and re-learning
We're going to split the sales data into four subsets of roughly equal size. Then you will estimate a 15th degree polynomial model on all four subsets of the data. Print the coefficients (you should use .print_rows(num_rows = 16) to view all of them) and plot the resulting fit (as we did above). The quiz will ask you some questions about these results.
To split the sales data into four subsets, we perform the following steps:
* First split sales into 2 subsets with `.random_split(0.5, seed=0)`.
* Next split the resulting subsets into 2 more subsets each. Use `.random_split(0.5, seed=0)`.
We set `seed=0` in these steps so that different users get consistent results.
You should end up with 4 subsets (`set_1`, `set_2`, `set_3`, `set_4`) of approximately equal size.
```
temp_set, temp_set_plus = sales.random_split(.5,seed=0)
set_1, set_2 = temp_set.random_split(.5,seed=0)
set_3, set_4 = temp_set_plus.random_split(.5,seed=0)
```
Fit a 15th degree polynomial on set_1, set_2, set_3, and set_4 using sqft_living to predict prices. Print the coefficients and make a plot of the resulting model.
```
poly_data = polynomial_sframe(set_1['sqft_living'], 15)
poly_data['price'] = set_1['price'] # add price to the data since it's the target
model = graphlab.linear_regression.create(poly_data, target = 'price', features = ['power_15'], validation_set = None)
model.get("coefficients")
# plt.plot(poly_data['power_15'],poly_data['price'],'.',
# poly_data['power_15'], model.predict(poly_data),'-')
poly_data = polynomial_sframe(set_2['sqft_living'], 15)
poly_data['price'] = set_2['price'] # add price to the data since it's the target
model = graphlab.linear_regression.create(poly_data, target = 'price', features = ['power_15'], validation_set = None)
model.get("coefficients")
plt.plot(poly_data['power_15'],poly_data['price'],'.',
poly_data['power_15'], model.predict(poly_data),'-')
poly_data = polynomial_sframe(set_3['sqft_living'], 15)
poly_data['price'] = set_3['price'] # add price to the data since it's the target
model = graphlab.linear_regression.create(poly_data, target = 'price', features = ['power_15'], validation_set = None)
model.get("coefficients")
plt.plot(poly_data['power_15'],poly_data['price'],'.',
poly_data['power_15'], model.predict(poly_data),'-')
poly_data = polynomial_sframe(set_4['sqft_living'], 15)
poly_data['price'] = set_4['price'] # add price to the data since it's the target
model = graphlab.linear_regression.create(poly_data, target = 'price', features = ['power_15'], validation_set = None)
model.get("coefficients")
# plt.plot(poly_data['power_15'],poly_data['price'],'.',
# poly_data['power_15'], model.predict(poly_data),'-')
```
Some questions you will be asked on your quiz:
**Quiz Question: Is the sign (positive or negative) for power_15 the same in all four models?**
**Quiz Question: (True/False) the plotted fitted lines look the same in all four plots**
# Selecting a Polynomial Degree
Whenever we have a "magic" parameter like the degree of the polynomial there is one well-known way to select these parameters: validation set. (We will explore another approach in week 4).
We split the sales dataset 3-way into training set, test set, and validation set as follows:
* Split our sales data into 2 sets: `training_and_validation` and `testing`. Use `random_split(0.9, seed=1)`.
* Further split our training data into two sets: `training` and `validation`. Use `random_split(0.5, seed=1)`.
Again, we set `seed=1` to obtain consistent results for different users.
```
sales = sales.sort(['sqft_living', 'price'])
training_and_validation, testing = sales.random_split(.9,seed=1)
training, validation = training_and_validation.random_split(.5,seed=1)
print len(training)
print len(validation)
```
Next you should write a loop that does the following:
* For degree in [1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15] (to get this in python type range(1, 15+1))
* Build an SFrame of polynomial data of train_data['sqft_living'] at the current degree
* hint: my_features = poly_data.column_names() gives you a list e.g. ['power_1', 'power_2', 'power_3'] which you might find useful for graphlab.linear_regression.create( features = my_features)
* Add train_data['price'] to the polynomial SFrame
* Learn a polynomial regression model to sqft vs price with that degree on TRAIN data
* Compute the RSS on VALIDATION data (here you will want to use .predict()) for that degree and you will need to make a polynmial SFrame using validation data.
* Report which degree had the lowest RSS on validation data (remember python indexes from 0)
(Note you can turn off the print out of linear_regression.create() with verbose = False)
**Quiz Question: Which degree (1, 2, …, 15) had the lowest RSS on Validation data?**
Now that you have chosen the degree of your polynomial using validation data, compute the RSS of this model on TEST data. Report the RSS on your quiz.
```
for i in range(1,16):
poly_data = polynomial_sframe(training['sqft_living'], i)
my_features = poly_data.column_names()
poly_data['price'] = training['price'] # add price to the data since it's the target
model = graphlab.linear_regression.create(poly_data, target = 'price', features = my_features, validation_set = None, verbose = False)
validation_data = polynomial_sframe(validation['sqft_living'], i)
validation_data['price'] = validation['price']
predictions = model.predict(validation_data)
RSS = ((predictions - validation_data['price']) * (predictions - validation_data['price'])).sum()
print str(RSS)
```
**Quiz Question: what is the RSS on TEST data for the model with the degree selected from Validation data? (Make sure you got the correct degree from the previous question)**
```
poly_data = polynomial_sframe(testing['sqft_living'], 6)
my_features = poly_data.column_names()
poly_data['price'] = testing['price'] # add price to the data since it's the target
model = graphlab.linear_regression.create(poly_data, target = 'price', features = my_features, validation_set = None, verbose = False)
validation_data = polynomial_sframe(validation['sqft_living'], i)
validation_data['price'] = validation['price']
predictions = model.predict(validation_data)
RSS = ((predictions - validation_data['price']) * (predictions - validation_data['price'])).sum()
print str(RSS)
```
| github_jupyter |
```
%matplotlib inline
from IPython.display import display
import os
import subprocess
import requests
import datetime
import xarray as xr
import numpy as np
import pandas as pd
from scipy.interpolate import griddata
import matplotlib.pyplot as plt
import cartopy.crs as ccrs
def fetch_smn_data(from_date, to_date, output_dir):
"""
Fetch meteorological data from SMN
"""
if from_date < datetime.datetime(2017, 11, 26):
raise ValueError("No data before 2017-11-26")
files = _download_smn_data(from_date, to_date, output_dir)
dataframes = []
for file in files:
csv_file = _convert_to_csv(file)
dataframes.append(pd.read_csv(csv_file))
df = pd.concat(dataframes, ignore_index=True)
_convert_datetime(df)
# Convert Temperature to K
df.temp += 273.15
# Convert pressure to Pa
df.pressure *= 1e2
# Convert time to UTC
df.time += datetime.timedelta(hours=3)
# Drop nans
df.dropna(how="any", inplace=True)
return df
def _convert_to_csv(filename):
"""
"""
filename_csv = filename.replace(".txt", ".csv")
content = "date,hour,temp,humidity,pressure,wind_dir,wind_speed,station\n"
with open(filename, "rb") as f:
for i, line in enumerate(f):
if i < 2:
continue
line = line.decode('latin-1').strip()
content += line[:8].strip() + ","
content += line[8:14].strip() + ","
content += line[14:20].strip() + ","
content += line[20:25].strip() + ","
content += line[25:33].strip() + ","
content += line[33:38].strip() + ","
content += line[38:43].strip() + ","
content += line[43:].strip() + "\n"
with open(filename_csv, 'w') as f:
f.write(content)
return filename_csv
def _convert_datetime(df):
datetimes = []
for i in range(len(df)):
date = df.date[i]
hour = df.hour[i]
if np.isnan(hour):
datetimes.append(np.nan)
else:
hour = int(hour)
day, month, year = int(str(date)[:2]), int(str(date)[2:4]), int(str(date)[4:])
datetimes.append(datetime.datetime(year, month, day, hour))
df["time"] = datetimes
del df["hour"]
del df["date"]
def _download_smn_data(from_date, to_date, output_dir):
"""
Download meteorological data from SMN
"""
base_url = "https://ssl.smn.gob.ar/dpd/descarga_opendata.php?file=observaciones/"
day_interval = datetime.timedelta(days=1)
files = []
date = from_date
while True:
if date > to_date:
break
filename = "datohorario{:d}{:02d}{:02d}.txt".format(date.year, date.month, date.day)
file_path = os.path.join(output_dir, filename)
if not os.path.isfile(file_path):
response = requests.get(base_url + filename)
with open(file_path, "wb") as f:
f.write(response.content)
files.append(file_path)
date += day_interval
return files
output_dir = os.path.join("data", "smn")
from_date = datetime.datetime(2017, 11, 28)
to_date = datetime.datetime(2019, 11, 18)
df = fetch_smn_data(from_date, to_date, output_dir)
sj_aero = df.station == "SAN JUAN AERO"
sj = df[sj_aero]
sj = sj.set_index("time")
variables = [var for var in sj if var != "station"]
fix, axes = plt.subplots(
ncols=1, nrows=len(variables), sharex=True, figsize=(15, 6*len(variables))
)
for ax, variable in zip(axes, variables):
getattr(sj, variable).plot(ax=ax)
ax.set_title(variable)
plt.show()
sj.to_csv(os.path.join("data", "san_juan_airport_data.csv"))
```
| github_jupyter |
# UMAP Demo with Graphs
[UMAP](https://umap-learn.readthedocs.io/en/latest/) is a powerful dimensionality reduction tool which NVIDIA recently ported to GPUs with a python interface. In this notebook we will demostrate basic usage, plotting, and timing of the unsupervised CUDA (GPU) version of UMAP.
## Imports and Set Up
```
import os
import pandas as pd
import numpy as np
# libraries for scoring/clustering
from sklearn.manifold.t_sne import trustworthiness
# GPU UMAP
import cudf
from cuml.manifold.umap import UMAP as cumlUMAP
# plotting
import seaborn as sns
import matplotlib as mpl
import matplotlib.pyplot as plt
%matplotlib inline
sns.set(style='white', rc={'figure.figsize':(25, 12.5)})
# hide warnings
import warnings
warnings.filterwarnings("ignore", category=DeprecationWarning)
```
## Sanity Checks
We are going to work with the [fashion mnist](https://github.com/zalandoresearch/fashion-mnist) data set. This is a dataset consisting of 70,000 28x28 grayscale images of clothing. It should already be in the `data/fashion` folder, but let's do a sanity check!
```
if not os.path.exists('data/fashion'):
print("error, data is missing!")
```
Now let's make sure we have our RAPIDS compliant GPU. It must be Pascal or higher! You can also use this to define which GPU RAPIDS should use (advanced feature not covered here)
```
!nvidia-smi
```
## Helper Functions
```
# https://github.com/zalandoresearch/fashion-mnist/blob/master/utils/mnist_reader.py
def load_mnist(path, kind='train'):
import os
import gzip
import numpy as np
"""Load MNIST data from `path`"""
labels_path = os.path.join(path,
'%s-labels-idx1-ubyte.gz'
% kind)
images_path = os.path.join(path,
'%s-images-idx3-ubyte.gz'
% kind)
with gzip.open(labels_path, 'rb') as lbpath:
labels = np.frombuffer(lbpath.read(), dtype=np.uint8,
offset=8)
with gzip.open(images_path, 'rb') as imgpath:
images = np.frombuffer(imgpath.read(), dtype=np.uint8,
offset=16).reshape(len(labels), 784)
return images, labels
```
## Training
```
train, train_labels = load_mnist('data/fashion', kind='train')
test, test_labels = load_mnist('data/fashion', kind='t10k')
data = np.array(np.vstack([train, test]), dtype=np.float64) / 255.0
target = np.array(np.hstack([train_labels, test_labels]))
```
There are 60000 training images and 10000 test images
```
f"Train shape: {train.shape} and Test Shape: {test.shape}"
train[0].shape
```
As mentioned previously, each row in the train matrix is an image
```
# display a Nike? sneaker
pixels = train[0].reshape((28, 28))
plt.imshow(pixels, cmap='gray')
```
There is cost with moving data between host memory and device memory (GPU memory) and we will include that core when comparing speeds
```
%%time
record_data = (('fea%d'%i, data[:,i]) for i in range(data.shape[1]))
gdf = cudf.DataFrame(record_data)
```
`gdf` is a GPU backed dataframe -- all the data is stored in the device memory of the GPU. With the data converted, we can apply the `cumlUMAP` the same inputs as we do for the standard UMAP. Additionally, it should be noted that within cuml, [FAISS] https://github.com/facebookresearch/faiss) is used for extremely fast kNN and it's limited to single precision. `cumlUMAP` will automatically downcast to `float32` when needed.
```
%%timeit
g_embedding = cumlUMAP(n_neighbors=5, init="spectral").fit_transform(gdf)
```
## Visualization
OK, now let's plot the output of the embeddings so that we can see the seperation of the neighborhoods. Let's start by creating the classes.
```
classes = [
'T-shirt/top',
'Trouser',
'Pullover',
'Dress',
'Coat',
'Sandal',
'Shirt',
'Sneaker',
'Bag',
'Ankle boot']
#Needs to be redone because of timeit function sometimes loses our g_embedding variable
g_embedding = cumlUMAP(n_neighbors=5, init="spectral").fit_transform(gdf)
```
Just as the original author of UMAP, Leland McInnes, states in the [UMAP docs](https://umap-learn.readthedocs.io/en/latest/supervised.html), we can plot the results and show the separation between the various classes defined above.
```
g_embedding_numpy = g_embedding.to_pandas().values #it is necessary to convert to numpy array to do the visual mapping
fig, ax = plt.subplots(1, figsize=(14, 10))
plt.scatter(g_embedding_numpy[:,1], g_embedding_numpy[:,0], s=0.3, c=target, cmap='Spectral', alpha=1.0)
plt.setp(ax, xticks=[], yticks=[])
cbar = plt.colorbar(boundaries=np.arange(11)-0.5)
cbar.set_ticks(np.arange(10))
cbar.set_ticklabels(classes)
plt.title('Fashion MNIST Embedded via cumlUMAP');
```
Additionally, we can also quanititaviely compare the perfomance of `cumlUMAP` (GPU UMAP) to the reference/original implementation (CPU UMAP) using the [trustworthiness score](https://github.com/scikit-learn/scikit-learn/blob/master/sklearn/manifold/t_sne.py#L395). From the docstring:
> Trustworthiness expresses to what extent the local structure is retained. The trustworthiness is within [0, 1].
Like `t-SNE`, UMAP tries to capture both global and local structure and thus, we can apply the `trustworthiness` of the `g_embedding` data against the original input. With a higher score we are demonstrating that the algorithm does a better and better job of local structure retention. As [Corey Nolet](https://github.com/cjnolet) notes:
> Algorithms like UMAP aim to preserve local neighborhood structure and so measuring this property (trustworthiness) measures the algorithm's performance.
Scoring ~97% shows the GPU implementation is comparable to the original CPU implementation and the training time was ~9.5X faster
| github_jupyter |
# From Oliver Durr
## Variational Autoencoder (VAE)
A tutorial with code for a VAE as described in [Kingma and Welling, 2013](http://arxiv.org/abs/1312.6114). A talk with more details was given at the [DataLab Brown Bag Seminar](https://home.zhaw.ch/~dueo/bbs/files/vae.pdf).
Much of the code was taken, from https://jmetzen.github.io/2015-11-27/vae.html. However, I tried to focus more on the mathematical understanding, not so much on design of the algorithm.
### Some theoretical considerations
#### Outline
Situation: $x$ is from a high-dimensional space and $z$ is from a low-dimensional (latent) space, from which we like to reconstruct $p(x)$.
We consider a parameterized model $p_\theta(x|z)$ (with parameter $\theta$), to construct x for a given value of $z$. We build this model:
* $p_\theta(x | z)$ with a neural network determening the parameters $\mu, \Sigma$ of a Gaussian (or as done here with a Bernoulli-Density).
#### Inverting $p_\theta(x | z)$
The inversion is not possible, we therefore approximate $p(z|x)$ by $q_\phi (z|x)$ again a combination of a NN determening the parameters of a Gaussian
* $q_\phi(z | x)$ with a neural network + Gaussian
#### Training
We train the network treating it as an autoencoder.
#### Lower bound of the Log-Likelihood
The likelihood cannot be determined analytically. Therefore, in a first step we derive a lower (variational) bound $L^{v}$ of the log likelihood, for a given image. Technically we assume a discrete latent space. For a continous case simply replace the sum by the appropriate integral over the respective densities. We replace the inaccessible conditional propability $p(z|x)$ with an approximation $q(z|x)$ for which we later use a neural network topped by a Gaussian.
\begin{align}
L & = \log\left(p(x)\right) &\\
& = \sum_z q(z|x) \; \log\left(p(x)\right) &\text{multiplied with 1 }\\
& = \sum_z q(z|x) \; \log\left(\frac{p(z,x)}{p(z|x)}\right) &\\
& = \sum_z q(z|x) \; \log\left(\frac{p(z,x)}{q(z|x)} \frac{q(z|x)}{p(z|x)}\right) &\\
& = \sum_z q(z|x) \; \log\left(\frac{p(z,x)}{q(z|x)}\right) + \sum_z q(z|x) \; \log\left(\frac{q(z|x)}{p(z|x)}\right) &\\
& = L^{\tt{v}} + D_{\tt{KL}} \left( q(z|x) || p(z|x) \right) &\\
& \ge L^{\tt{v}} \\
\end{align}
The KL-Divergence $D_{\tt{KL}}$ is always positive, and the smaller the better $q(z|x)$ approximates $p(z|x)$
### Rewritting $L^\tt{v}$
We split $L^\tt{v}$ into two parts.
\begin{align}
L^{\tt{v}} & = \sum_z q(z|x) \; \log\left(\frac{p(z,x)}{q(z|x)}\right) & \text{with} \;\;p(z,x) = p(x|z) \,p(z)\\
& = \sum_z q(z|x) \; \log\left(\frac{p(x|z) p(z)}{q(z|x)}\right) &\\
& = \sum_z q(z|x) \; \log\left(\frac{p(z)}{q(z|x)}\right) + \sum_z q(z|x) \; \log\left(p(x|z)\right) &\\
& = -D_{\tt{KL}} \left( q(z|x) || p(z) \right) + \mathbb{E}_{q(z|x)}\left( \log\left(p(x|z)\right)\right) &\text{putting in } x^{(i)} \text{ for } x\\
& = -D_{\tt{KL}} \left( q(z|x^{(i)}) || p(z) \right) + \mathbb{E}_{q(z|x^{(i)})}\left( \log\left(p(x^{(i)}|z)\right)\right) &\\
\end{align}
Approximating $\mathbb{E}_{q(z|x^{(i)})}$ with sampling form the distribution $q(z|x^{(i)})$
#### Sampling
With $z^{(i,l)}$ $l = 1,2,\ldots L$ sampled from $z^{(i,l)} \thicksim q(z|x^{(i)})$
\begin{align}
L^{\tt{v}} & = -D_{\tt{KL}} \left( q(z|x^{(i)}) || p(z) \right)
+ \mathbb{E}_{q(z|x^{(i)})}\left( \log\left(p(x^{(i)}|z)\right)\right) &\\
L^{\tt{v}} & \approx -D_{\tt{KL}} \left( q(z|x^{(i)}) || p(z) \right)
+ \frac{1}{L} \sum_{i=1}^L \log\left(p(x^{(i)}|z^{(i,l)})\right) &\\
\end{align}
#### Calculation of $D_{\tt{KL}} \left( q(z|x^{(i)}) || p(z) \right)$
TODO
| github_jupyter |
# En<->Fr translation
```
%matplotlib inline
import importlib
import utils2; importlib.reload(utils2)
from utils2 import *
from gensim.models import word2vec
limit_mem()
# path='/data/jhoward/datasets/fr-en-109-corpus/'
# dpath = 'data/translate/'
path='data/translate/fr-en-109-corpus/'
dpath = 'data/translate/'
```
## Prepare corpus
```
fname=path+'giga-fren.release2.fixed'
en_fname = fname+'.en'
fr_fname = fname+'.fr'
re_eq = re.compile('^(Wh[^?.!]+\?)')
re_fq = re.compile('^([^?.!]+\?)')
lines = ((re_eq.search(eq), re_fq.search(fq))
for eq, fq in zip(open(en_fname), open(fr_fname)))
qs = [(e.group(), f.group()) for e,f in lines if e and f]
len(qs)
qs[:6]
dump(qs, dpath+'qs.pkl')
qs = load(dpath+'qs.pkl')
en_qs, fr_qs = zip(*qs)
re_mult_space = re.compile(r" *")
re_mw_punc = re.compile(r"(\w[’'])(\w)")
re_punc = re.compile("([\"().,;:/_?!—])")
re_apos = re.compile(r"(\w)'s\b")
def simple_toks(sent):
sent = re_apos.sub(r"\1 's", sent)
sent = re_mw_punc.sub(r"\1 \2", sent)
sent = re_punc.sub(r" \1 ", sent).replace('-', ' ')
sent = re_mult_space.sub(' ', sent)
return sent.lower().split()
fr_qtoks = list(map(simple_toks, fr_qs)); fr_qtoks[:4]
en_qtoks = list(map(simple_toks, en_qs)); en_qtoks[:4]
simple_toks("Rachel's baby is cuter than other's.")
def toks2ids(sents):
voc_cnt = collections.Counter(t for sent in sents for t in sent)
vocab = sorted(voc_cnt, key=voc_cnt.get, reverse=True)
vocab.insert(0, "<PAD>")
w2id = {w:i for i,w in enumerate(vocab)}
ids = [[w2id[t] for t in sent] for sent in sents]
return ids, vocab, w2id, voc_cnt
fr_ids, fr_vocab, fr_w2id, fr_counts = toks2ids(fr_qtoks)
en_ids, en_vocab, en_w2id, en_counts = toks2ids(en_qtoks)
len(en_vocab), len(fr_vocab)
```
## Word vectors
```
# en_vecs, en_wv_word, en_wv_idx = load_glove(
# '/data/jhoward/datasets/nlp/glove/results/6B.100d')
en_vecs, en_wv_word, en_wv_idx = load_glove('data/glove/results/6B.100d')
en_w2v = {w: en_vecs[en_wv_idx[w]] for w in en_wv_word}
n_en_vec, dim_en_vec = en_vecs.shape
dim_fr_vec = 200
# - not used
# fr_wik = pickle.load(open('/data/jhoward/datasets/nlp/polyglot-fr.pkl', 'rb'),
# encoding='latin1')
```
- Word vectors: http://fauconnier.github.io/index.html#wordembeddingmodels
- Corpus: https://www.sketchengine.co.uk/frwac-corpus/
```
# w2v_path='/data/jhoward/datasets/nlp/frWac_non_lem_no_postag_no_phrase_200_skip_cut100.bin'
w2v_path='data/frwac/frWac_non_lem_no_postag_no_phrase_200_skip_cut100.bin'
fr_model = word2vec.KeyedVectors.load_word2vec_format(w2v_path, binary=True)
fr_voc = fr_model.vocab
def create_emb(w2v, targ_vocab, dim_vec):
vocab_size = len(targ_vocab)
emb = np.zeros((vocab_size, dim_vec))
for i, word in enumerate(targ_vocab):
try:
emb[i] = w2v[word]
except KeyError:
# If we can't find the word, randomly initialize
emb[i] = normal(scale=0.6, size=(dim_vec,))
return emb
en_embs = create_emb(en_w2v, en_vocab, dim_en_vec); en_embs.shape
fr_embs = create_emb(fr_model, fr_vocab, dim_fr_vec); fr_embs.shape
```
## Prep data
```
en_lengths = collections.Counter(len(s) for s in en_ids)
maxlen = 30
len(list(filter(lambda x: len(x) > maxlen, en_ids))), len(
list(filter(lambda x: len(x) <= maxlen, en_ids)))
len(list(filter(lambda x: len(x) > maxlen, fr_ids))), len(
list(filter(lambda x: len(x) <= maxlen, fr_ids)))
en_padded = pad_sequences(en_ids, maxlen, padding="post", truncating="post")
fr_padded = pad_sequences(fr_ids, maxlen, padding="post", truncating="post")
en_padded.shape, fr_padded.shape, en_embs.shape
n = int(len(en_ids)*0.9)
idxs = np.random.permutation(len(en_ids))
fr_train, fr_test = fr_padded[idxs][:n], fr_padded[idxs][n:]
en_train, en_test = en_padded[idxs][:n], en_padded[idxs][n:]
```
## Model
```
en_train.shape
parms = {'verbose': 0, 'callbacks': [TQDMNotebookCallback()]}
fr_wgts = [fr_embs.T, np.zeros((len(fr_vocab,)))]
inp = Input((maxlen,))
x = Embedding(len(en_vocab), dim_en_vec, input_length=maxlen,
weights=[en_embs], trainable=False)(inp)
x = Bidirectional(LSTM(128, return_sequences=True))(x)
x = Bidirectional(LSTM(128, return_sequences=True))(x)
x = LSTM(128, return_sequences=True)(x)
x = TimeDistributed(Dense(dim_fr_vec))(x)
x = TimeDistributed(Dense(len(fr_vocab), weights=fr_wgts))(x)
x = Activation('softmax')(x)
model = Model(inp, x)
model.compile('adam', 'sparse_categorical_crossentropy')
K.set_value(model.optimizer.lr, 1e-3)
hist=model.fit(en_train, np.expand_dims(fr_train,-1), batch_size=64, epochs=20, **parms,
validation_data=[en_test, np.expand_dims(fr_test,-1)])
plot_train(hist)
model.save_weights(dpath+'trans.h5')
model.load_weights(dpath+'trans.h5')
```
## Testing
```
def sent2ids(sent):
sent = simple_toks(sent)
ids = [en_w2id[t] for t in sent]
return pad_sequences([ids], maxlen, padding="post", truncating="post")
def en2fr(sent):
ids = sent2ids(sent)
tr_ids = np.argmax(model.predict(ids), axis=-1)
return ' '.join(fr_vocab[i] for i in tr_ids[0] if i>0)
en2fr("what is the size of canada?")
```
| github_jupyter |
## Using scripts to perform Intel Advisor roofline profiling on Devito
This notebook uses the prewritten scripts `run_advisor.py`, `roofline.py` and `advisor_to_json.py` to show how you can easily profile a Devito application using Intel Advisor 2020 and plot memory-bound/compute-bound roofline models depicting the current state of the application's performance. These scripts can be found in the `benchmarks/user/advisor` folder of the full Devito repository. They are also available as part of the Devito package.
First, we are going to need a couple of imports to allow us to work with Devito and to run command line applications from inside this jupyter notebook. These will be needed for all three scripts.
```
import devito
from IPython.display import Image
import os
devito_path = os.path.dirname(''.join(devito.__path__))
os.environ['DEVITO_JUPYTER'] = devito_path
```
### Setting up the Advisor environment
Before running the following pieces of code, we must make sure that the Advisor environment, alongside the Intel C compiler `icc`, are correctly activated on the machine you wish to use the scripts on. To do so, run the following commands:
for Intel oneAPI:
```shell
source /opt/intel/oneapi/advisor/latest/advixe-vars.sh
```
```shell
source /opt/intel/oneapi/compiler/latest/env/vars.sh <architecture, e.g. intel64>
```
or
for Intel Parallel Studio:
```shell
source /opt/intel/advisor/advixe-vars.sh
```
```shell
source /opt/intel/compilers_and_libraries/linux/bin/compilervars.sh <architecture, e.g. intel64>
```
If your Advisor or icc have not been installed in the `/opt/intel/oneapi/advisor` (equivalently `/opt/intel/advisor`) or `/opt/intel/oneapi/compiler` (or `/opt/intel/compilers_and_libraries`) directory, replace them with your chosen path.
### Collecting performance data with `run_advisor.py`
Before generating graphical models or have data to be exported, we need to collect the performance data of our interested program. The command line that we will use is:
```shell
python3 <path-to-devito>/benchmarks/user/advisor/run_advisor.py --path <path-to-devito>/benchmarks/user/benchmark.py --exec-args "run -P acoustic -d 128 128 128 -so 4 --tn 50 --autotune off" --output <path-to-devito>/examples/performance/profilings --name JupyterProfiling
```
* `--path` specifies the path of the Devito/python executable,
* `--exec-args` specifies the command line arguments that we want to pass to our executable,
* `--output` specifies the directory where we want to permanently save our profiling reports,
* `--name` specifies the name of the single profiling that will be effected.
Let's run the command to do the profiling of our example application.
```
#NBVAL_SKIP
! python3 $DEVITO_JUPYTER/benchmarks/user/advisor/run_advisor.py --path $DEVITO_JUPYTER/benchmarks/user/benchmark.py --exec-args "run -P acoustic -d 128 128 128 -so 4 --tn 50 --autotune off" --output $DEVITO_JUPYTER/examples/performance/profilings --name JupyterProfiling
```
The above call might take a few minutes depending on what machine you are running the code on, please have patience. After it is done, we have Intel Advisor data from which we can generate rooflines and export data.
### Generating a roofline model to display profiling data
Now that we have collected the data inside a profiling directory, we can use the `roofline.py` script to produce a pdf of the roofline data that has been collected in the previous run. There are two visualisation modes for the generated roofline:
* overview: displays a single point with the overall GFLOPS/s and arithmetic intensity of the program
* top-loops: displays all points within runtime within one order of magnitude compared to the top time consuming loop
First, we will produce an 'overview' roofline. The command line that we will use is:
```shell
python3 <path-to-devito>/benchmarks/user/advisor/roofline.py --mode overview --name <path-to-devito>/examples/performance/resources/OverviewRoof --project <path-to-devito>/examples/performance/profilings/JupyterProfiling
```
* `--mode` specifies the mode as described (either `overview` or `top-loops`)
* `--name` specifies the name of the pdf file that will contain the roofline representation of the data
* `--project` specifies the directory where the profiling data is stored
Let's run the command.
```
#NBVAL_SKIP
! python3 $DEVITO_JUPYTER/benchmarks/user/advisor/roofline.py --mode overview --name $DEVITO_JUPYTER/examples/performance/resources/OverviewRoof --project $DEVITO_JUPYTER/examples/performance/profilings/JupyterProfiling
```
Once this command has completed, we can now observe the gathered profiling data through a roofline model.
```
Image(filename=os.path.join(devito_path, 'examples/performance/resources/OverviewRoof.png'))
```
Similarly, we can also produce a graph which displays the most time consuming loop alongside all other loops which have execution time within one order of magnitude from it. This is done by using `top-loops` mode.
```
#NBVAL_SKIP
! python3 $DEVITO_JUPYTER/benchmarks/user/advisor/roofline.py --mode top-loops --name $DEVITO_JUPYTER/examples/performance/resources/TopLoopsRoof --project $DEVITO_JUPYTER/examples/performance/profilings/JupyterProfiling
```
With the command having run, we can inspect the image that has been created and compare it to the overview mode roofline.
```
Image(filename=os.path.join(devito_path, 'examples/performance/resources/TopLoopsRoof.png'))
```
As you can see from this roofline graph, the main point is different from the single point of the previous graph. Moreover, each point is labelled with 'Time' and 'Incidence' indicators. These represent the total execution time of each loop's main body and their percentage incidence on the total execution time of the main time loop.
### Exporting Advisor roofline data as JSON
For easy and flexible exporting, the `advisor_to_json.py` script allows you to pack all the information represented within the previously seen graphs inside a JSON file. The command line which is needed to export our project's data in JSON format is:
```shell
python3 <path-to-devito>/benchmarks/user/advisor/advisor_to_json.py --name <path-to-devito>/examples/performance/resources/RoofsData --project <path-to-devito>/examples/performance/profilings/JupyterProfiling
```
* `--name` specifies the name of the JSON file which will be generated
* `--project` specifies the Advisor folder which contains all the collected data about the application
As always, let's run the command through bash.
```
#NBVAL_SKIP
! python3 $DEVITO_JUPYTER/benchmarks/user/advisor/advisor_to_json.py --name $DEVITO_JUPYTER/examples/performance/resources/RoofsData --project $DEVITO_JUPYTER/examples/performance/profilings/JupyterProfiling
```
We can now open the generated JSON file to inspect that we indeed have the data extracted from our roofline models.
```
json_file = open(os.path.join(devito_path, 'examples/performance/resources/RoofsData.json'), 'r')
json_file.read()
```
Let's take a look at the generated JSON. Inside we can see that the file is subdivided into three sections: one to hold the points used to draw the rooflines (`roofs`), one to contain the single overview point (`overview`) and another one that contains information about the top loops as specified earlier in this tutorial (`top-loops`).
By default, `advisor_to_json.py` collects both overview and top-loops data, but the information to collect can be specified using a `--mode` flag analogous to the one used in `roofline.py`.
### Further flags and functionality
The last two scripts contain more flags that you can use to adjust data collection, displaying and exporting.
`roofline.py`:
* `--scale` specifies how much rooflines should be scaled down due to using fewer cores than available
* `--precision` specifies the arithmetic precision of the integral operators
* `--th` specifies the threshold percentage over which to display loops in top-loops mode
`advisor_to_json.py`:
* `--scale`, as previously
* `--precision`, as previously
* `--th`, as previously
If you want to learn more about what they do, add a `--help` flag to the script that you are executing.
| github_jupyter |
```
import numpy as np
import librosa
import librosa.display
import matplotlib.pyplot as plt
from tqdm import tqdm_notebook as tqdm
import os
import multiprocessing
import threading
# musDB root dir
musdb_root_dir = '/home/scpark/hard/datasets/musdb18'
# directory to save the train data npz file
train_data_dir = '/home/scpark/hard/datasets/musdb18_wav'
# directory to save the test data npz file
test_data_dir = '/home/scpark/hard/datasets/musdb18_wav/test'
# chunk length (44100 sample rate, 10 seconds)
chunk_length = 10 * 44100
# the thread numbers to create data files
thread_nums = 4 # multiprocessing.cpu_count()
import musdb
# load the musDB
mus = musdb.DB(root_dir=musdb_root_dir)
# load the training tracks
train_tracks = mus.load_mus_tracks(subsets=['train'])
# load the test tracks
test_tracks = mus.load_mus_tracks(subsets=['test'])
def create_tracks_files(tracks, track_ids, save_dir):
for track_id, track in zip(track_ids, tracks):
length = len(track.audio)
for j, start_index in tqdm(enumerate(range(0, length, chunk_length))):
filename = save_dir + '/' + str(track_id) + '.' + str(j) + '.' + track.name + '.npz'
# check if file already exist
if os.path.exists(filename):
continue
# check if directory to save exist
if not os.path.exists(save_dir):
os.makedirs(save_dir)
audio = track.audio[start_index : start_index + chunk_length]
vocals = track.targets['vocals'].audio[start_index : start_index + chunk_length]
drums = track.targets['drums'].audio[start_index : start_index + chunk_length]
bass = track.targets['bass'].audio[start_index : start_index + chunk_length]
other = track.targets['other'].audio[start_index : start_index + chunk_length]
accompaniment = track.targets['accompaniment'].audio[start_index : start_index + chunk_length]
if len(audio) < chunk_length:
pad_length = chunk_length - len(audio)
audio = np.concatenate([audio, np.zeros([pad_length, 2])], axis=0)
vocals = np.concatenate([vocals, np.zeros([pad_length, 2])], axis=0)
drums = np.concatenate([drums, np.zeros([pad_length, 2])], axis=0)
bass = np.concatenate([bass, np.zeros([pad_length, 2])], axis=0)
other = np.concatenate([other, np.zeros([pad_length, 2])], axis=0)
accompaniment = np.concatenate([accompaniment, np.zeros([pad_length, 2])], axis=0)
elif len(audio) == 0:
continue
data = {
"audio": audio,
"vocals": vocals,
"drums": drums,
"bass": bass,
"other": other,
"accompaniment": accompaniment
}
np.savez(filename, **data, allow_pickle=False)
def create_files(tracks, save_dir, thread_nums):
tracks_list = [[] for _ in range(thread_nums)]
track_ids_list = [[] for _ in range(thread_nums)]
for track_id, track in enumerate(tracks):
tracks_list[track_id % thread_nums].append(track)
track_ids_list[track_id % thread_nums].append(track_id)
threads = []
for thread_id in range(thread_nums):
thread = threading.Thread(target=create_tracks_files, args=(tracks_list[thread_id], track_ids_list[thread_id], save_dir))
thread.start()
threads.append(thread)
for thread in threads:
thread.join()
create_files(train_tracks, train_data_dir, thread_nums)
create_files(test_tracks, test_data_dir, thread_nums)
```
| github_jupyter |
```
from muselsl import stream, list_muses, view, record
from multiprocessing import Process
from mne import Epochs, find_events
from time import time, strftime, gmtime
import os
from stimulus_presentation import n170
from utils import utils
from collections import OrderedDict
import warnings
warnings.filterwarnings('ignore')
```
# N170
<img style="height: 300px; float: right" src="https://www.researchgate.net/profile/Vincent_Walsh3/publication/49833438/figure/fig1/AS:394279194251295@1471014893499/The-P1-and-N170-ERP-components-The-graph-shows-the-grand-average-ERP-responses-from-ten.png"/>
The N170 is a large negative event-related potential (ERP) component that occurs after the detection of faces, but not objects, scrambled faces, or other body parts such as hands. The N170 occurs around 170ms after face perception and is most easily detected at lateral posterior electrodes such as T5 and T6 [1](http://citeseerx.ist.psu.edu/viewdoc/download?doi=10.1.1.601.6917&rep=rep1&type=pdf). Frontal or profile views of human (and animal [2](https://www.ncbi.nlm.nih.gov/pubmed/14995895)) faces elicit the strongest N170 and the strength of the N170 does not seem to be influenced by how familiar a face is. Thus, although there is no consensus on the specific source of the N170, researchers believe it is related to activity in the fusiform face area, an area of the brain that shows a similar response pattern and is involved in encoding the holistic representation of a face (i.e eyes, nose mouth all arranged in the appropriate way).
In this notebook, we will attempt to detect the N170 with the Muse headband using faces and houses as our stimuli. The Muse's temporal electrodes (TP9 and TP10) are well positioned to detect the N170 and we expect we'll be able to see an N170 emerge from just a few dozen trials. We will then run several different classification algorithms on our data in order to evaluate the performance of a potential brain-computer interface using the N170.
## Step 1: Connect to an EEG Device
*Note: if using Windows 10 and BlueMuse, skip this section and connect using the BlueMuse GUI*
Make sure your Muse 2016 is turned on and then run the following code. It should detect and connect to the device and begin 'Streaming...'
If the device is not found or the connection times out, try running this code again
```
# Search for available Muse devices
muses = list_muses()
# Start a background process that will stream data from the first available Muse
stream_process = Process(target=stream, args=(muses[0]['address'],))
stream_process.start()
```
## Step 2: Apply the EEG Device and Wait for Signal Quality to Stabilize
Once your Muse is connected and streaming data, put it on and run the following code to view the raw EEG data stream.
The numbers on the side of the graph indicate the variance of the signal. Wait until this decreases below 10 for all electrodes before proceeding.
```
%matplotlib
# On Windows, you may need to run the command %matplotlib tk
view()
```
## Step 3: Run the Experiment
Modify the variables in the following code chunk to define how long you want to run the experiment and the name of the subject and session you are collecting data from.
```
# Define these parameters
duration = 120 # in seconds. 120 is recommended
subject = 1 # unique id for each participant
session = 1 # represents a data collection session. Multiple trials can be performed for each session
```
Seat the subject in front of the computer and run the following cell to run a single trial of the experiment.
In order to maximise the possibility of success, participants should take the experiment in a quiet environment and do their best to minimize movement that might contaminate the signal. With their jaw and face relaxed, subjects should focus on the stimuli, mentally noting whether they see a "face" or a "house".
Data will be recorded into CSV files in the `eeg-notebooks/data` directory
```
recording_path = os.path.join(os.path.expanduser("~"), "eeg-notebooks", "data", "visual", "N170", "subject" + str(subject), "session" + str(session), ("recording_%s.csv" %
strftime("%Y-%m-%d-%H.%M.%S", gmtime())) + ".csv")
print('Recording data to: ', recording_path)
stimulus = Process(target=n170.present, args=(duration,))
recording = Process(target=record, args=(duration, recording_path))
stimulus.start()
recording.start()
```
### Repeat Data Collection 3-6 times
Visualizing ERPs requires averaging the EEG response over many different rounds of stimulus presentation. Depending on experimental conditions, this may require as little as one two minute trial or as many as 6. We recommend repeating the above experiment 3-6 times before proceeding.
Make sure to take breaks, though! Inattention, fatigue, and distraction will decrease the quality of event-related potentials such as the N170
## Step 4: Prepare the Data for Analysis
Once a suitable data set has been collected, it is now time to analyze the data and see if we can identify the N170
### Load data into MNE objects
[MNE](https://martinos.org/mne/stable/index.html) is a very powerful Python library for analyzing EEG data. It provides helpful functions for performing key tasks such as filtering EEG data, rejecting artifacts, and grouping EEG data into chunks (epochs).
The first step to using MNE is to read the data we've collected into an MNE `Raw` object
```
raw = utils.load_data('visual/N170', sfreq=256.,
subject_nb=subject, session_nb=session)
```
### Visualizing the Power Spectrum
Plotting the power spectral density (PSD) of our dataset will give us a glimpse at the different frequencies that are present. We won't be able to see the N170 in the PSD, but it will give us an impression of how noisy our data was. A very noisy or flat PSD may represent poor signal quality at certain electrodes
```
%matplotlib inline
raw.plot_psd();
```
This PSD looks good. There is a large peak at 60hz, representing background electrical activity.
### Filtering
Most ERP components are composed of lower frequency fluctuations in the EEG signal. Thus, we can filter out all frequencies between 1 and 30 hz in order to increase our ability to detect them.
```
raw.filter(1,30, method='iir')
raw.plot_psd(fmin=1, fmax=30);
```
This PSD of frequencies between 1 and 30 hz looks good. The difference between the temporal channels (red and black) and the frontal channels (blue and green) is clearly evident. The huge peak from 1 to 3hz is largely due to the presence of eye blinks, which produce large amplitude, low-frequency events in the EEG.
### Epoching
Next, we will chunk (epoch) the data into segments representing the data 100ms before to 800ms after each stimulus. No baseline correction is needed (signal is bandpass filtered) and we will reject every epoch where the amplitude of the signal exceeded 75 uV, which should most eye blinks.
```
# Create an array containing the timestamps and type of each stimulus (i.e. face or house)
events = find_events(raw)
event_id = {'House': 1, 'Face': 2}
# Create an MNE Epochs object representing all the epochs around stimulus presentation
epochs = Epochs(raw, events=events, event_id=event_id,
tmin=-0.1, tmax=0.8, baseline=None,
reject={'eeg': 75e-6}, preload=True,
verbose=False, picks=[0,1,2,3])
print('sample drop %: ', (1 - len(epochs.events)/len(events)) * 100)
epochs
```
Sample drop % is an important metric representing how noisy our data set was. If this is greater than 20%, consider ensuring that signal variances is very low in the raw EEG viewer and collecting more data
## Step 5: Analyze the Data
Finally, we can now analyze our results by averaging the epochs that occured during the different stimuli and looking for differences in the waveform
### Epoch average
With our `plot_conditions` utility function, we can plot the average ERP for all electrodes for both conditions:
```
%matplotlib inline
conditions = OrderedDict()
conditions['House'] = [1]
conditions['Face'] = [2]
fig, ax = utils.plot_conditions(epochs, conditions=conditions,
ci=97.5, n_boot=1000, title='',
diff_waveform=(1, 2))
```
Here we have a very nice deflection in the temporal channels around 200ms for face stimuli. This is likely the N170, although appearing slightly later due to delay in receiving the data over bluetooth.
There's not much to see in the frontal channels (AF7 and AF8), but that's to be expected based on the fact that the N170 is mostly a lateral posterior brain phenomenon
### Decoding the N170
Next, we will use 4 different machine learning pipelines to classify the N170 based on the data we collected. The
- **Vect + LR** : Vectorization of the trial + Logistic Regression. This can be considered the standard decoding pipeline for MEG / EEG.
- **Vect + RegLDA** : Vectorization of the trial + Regularized LDA. This one is very commonly used in P300 BCIs. It can outperform the previous one but become unusable if the number of dimension is too high.
- **ERPCov + TS**: ErpCovariance + Tangent space mapping. One of the most reliable Riemannian geometry-based pipeline.
- **ERPCov + MDM**: ErpCovariance + MDM. A very simple, yet effective (for low channel count), Riemannian geometry classifier.
Evaluation is done through cross-validation, with area-under-the-curve (AUC) as metric (AUC is probably the best metric for binary and unbalanced classification problem)
*Note: because we're doing machine learning here, the following cell may take a while to complete*
```
import pandas as pd
from sklearn.pipeline import make_pipeline
from mne.decoding import Vectorizer
from sklearn.linear_model import LogisticRegression
from sklearn.preprocessing import StandardScaler
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis as LDA
from sklearn.model_selection import cross_val_score, StratifiedShuffleSplit
from pyriemann.estimation import ERPCovariances, XdawnCovariances
from pyriemann.tangentspace import TangentSpace
from pyriemann.classification import MDM
from collections import OrderedDict
clfs = OrderedDict()
clfs['Vect + LR'] = make_pipeline(Vectorizer(), StandardScaler(), LogisticRegression())
clfs['Vect + RegLDA'] = make_pipeline(Vectorizer(), LDA(shrinkage='auto', solver='eigen'))
clfs['ERPCov + TS'] = make_pipeline(ERPCovariances(estimator='oas'), TangentSpace(), LogisticRegression())
clfs['ERPCov + MDM'] = make_pipeline(ERPCovariances(estimator='oas'), MDM())
clfs['XdawnCov + TS'] = make_pipeline(XdawnCovariances(estimator='oas'), TangentSpace(), LogisticRegression())
clfs['XdawnCov + MDM'] = make_pipeline(XdawnCovariances(estimator='oas'), MDM())
# format data
epochs.pick_types(eeg=True)
X = epochs.get_data() * 1e6
times = epochs.times
y = epochs.events[:, -1]
# define cross validation
cv = StratifiedShuffleSplit(n_splits=20, test_size=0.25,
random_state=42)
# run cross validation for each pipeline
auc = []
methods = []
for m in clfs:
print(m)
try:
res = cross_val_score(clfs[m], X, y==2, scoring='roc_auc',
cv=cv, n_jobs=-1)
auc.extend(res)
methods.extend([m]*len(res))
except:
pass
## Plot Decoding Results
import seaborn as sns
from matplotlib import pyplot as plt
results = pd.DataFrame(data=auc, columns=['AUC'])
results['Method'] = methods
fig = plt.figure(figsize=[8,4])
sns.barplot(data=results, x='AUC', y='Method')
plt.xlim(0.4, 0.9)
sns.despine()
```
The best classifiers for this data set appear to be the ERPCov and XdawnCov with tangent space projection pipelines. AUC is around .7, which is good, but on the low end for being able to run a brain-computer interface.
## Step 6: Share your Data!
How did your experiment go? If you're excited by your results we'd love to see your data!
Follow the instructions on our [Contributions](https://github.com/NeuroTechX/eeg-notebooks/blob/master/CONTRIBUTING.md) page to make a pull request with your data and we'll review it to be added to the EEG notebooks project.
| github_jupyter |
## Dispatching
In its simplest form, dispatching involves overloading methods based on their arguments. Implementations that do this based on argument types like [functools.singledispatch](https://docs.python.org/3/library/functools.html) and [multipledispatch](https://github.com/mrocklin/multipledispatch) make this straightforward.
This approach is problematic with duck array types used in functions that take multiple arguments though. A simple example here shows how \_\_array_ufunc__ and \_\_array_function__ make interoperability between backends possible to an extent, but the backend that ultimately handles an operation is arbitrary (it's the first one that doesn't throw NotImplemented) so the result type will often vary based on the argument order, as does the success of the operation:
```python
import dask.array as da
import sparse
a1 = da.array([1, 0])
a2 = sparse.COO.from_numpy(np.array([1, 0]))
# dask is better at handling arrays from other backends dispatched to ufunc
# while sparse only supports sparse or numpy arrays (the first case here
# works because dask is applying the function on numpy array chunks)
type(a1 + a2) # -> dask.array.core.Array
type(a2 + a1) # -> operand type(s) all returned NotImplemented
# similarly:
a3 = a1.map_blocks(sparse.COO.from_numpy)
type(np.stack([a1, a3])) # -> dask.array.core.Array
type(np.stack([a3, a1])) # -> All arrays must be instances of SparseArray.
type(np.stack([a1, a2])) # -> All arrays must be instances of SparseArray.
```
This means that code using multipledispatch would need to have overloads that match to each combination of backend types and perform coercion where necessary, likely targeting one of the backends present in the arguments. Some drawbacks to this are that defining overloads for more than 2 arguments or lists of arrays is hard and that the "target" backend is implicit in the implementation rather than controlled by the user.
A solution to this proposed in [unumpy](https://github.com/Quansight-Labs/unumpy) (via [uarray](https://github.com/Quansight-Labs/uarray)) is to make the target backend for any numpy functions explicit, as well as provide hooks for coercion of arguments to that backend. This is a good solution for much of the simpler functionality in a genetics toolkit but poses an issue for any more complex domain-specific algorithms in that it would be perfectly reasonable to expect that more than one array backend will be useful for doing things efficiently. From this perspective, the target implementations become something more like "algorithms" than "backends" and they should be free to make use of whatever array backends are most beneficial for a particular step (and coercion of arguments is simple when the target backend is clear). An example would be kinship estimation via CuPy followed by maximal independent set selection using a sparse array backend for relatedness pruning.
A further consideration is that dispatch may be something we want to eventually automate, rather than forcing our users to always think about it (e.g. an array with 1% sparsity should not go to an implementation that relies on a sparse backend, tiny arrays should not go to a chunked backend, big dask arrays should not be force into a numpy backend, etc.).
This prototype shows a small framework that is based to a degree on uarray and is centered around doing dispatching with all of the following in one place:
- **User preferences**: These always get highest priority in choosing an implementation
- **Configuration**: Inevitably we'll want a configuration framework (combined from disk, env vars, global vars, etc.) so this will be a good place to use it for parameterization of backends and dispatch automation
- **Arguments**: The arrays themselves for type, shape, and content analysis
Both uarray and multipledispatch are degenerate cases for this.
```
from lib import api
xr.set_options(display_style='html');
```
The abstraction consists of a "Fronted", a "Backend" and an API of stub functions.
An example Frontend would look like this where `Frontend` is an internal implementation used by the `MyFrontend` class. A class like `MyFrontend` would exist for every major piece of functionality in the library that benefits from moderately complex dispatching.
```
# This base class is defined once somewhere
class Frontend:
def __init__(self, config: Configuration = None):
self.config = config
self.backends = dict()
def register(self, backend: Backend) -> None:
# Frontends need to be aware of backends in order to choose intelligently between them
if backend.domain != self.domain:
raise ValueError('Backend with domain {backend.domain} not compatible with frontend domain {self.domain}')
self.backends[backend.id] = backend
def resolve(self, fn: Callable, *args, **kwargs) -> Backend:
# Choose a backend to dispatch to based on as much information as possible:
# First look for overrides in arguments passed to the function
backend_id = kwargs.get('backend')
# Next look for overrides in configuration
backend_id = backend_id or self.config.get(str(self.domain.append('backend')))
# Check to see what if any backends have the required packages installed
backend = [be for be in self.backends.values() if is_compatible(be)]
if backend is None:
raise ValueError(f'No suitable backend found for function {fn.__name__} (domain = {self.domain})')
# ** Analyze fn/args/kwargs here, in the future **
return backend
def dispatch(self, fn: Callable, *args, **kwargs):
self.resolve(fn, *args, **kwargs).dispatch(fn, *args, **kwargs)
def add(self, fn: Callable):
# Wrap a function to be dispatched and preserve docs
@functools.wraps(fn)
def wrapper(*args, **kwargs):
return self.dispatch(fn, *args, **kwargs)
return wrapper
# This is defined somewhere close to where the API to dispatch over lives (at least within the same package)
class MyFrontend(Frontend):
domain = 'genetics.method'
frontend = MyFrontend()
```
The actual API methods are simply stubs somewhere that have access to the frontend created above:
```
class API:
@frontend.add
def ld_prune(ds: Dataset) -> Dataset:
""" All documentation goes here """
pass
```
The backends should be separated from the frontend to make it as easy as possible to isolate imports for their optional dependencies. They do however need to know which frontend they are associated with and register themselves to it:
```
class AdvancedBackend(Backend):
domain = 'genetics.method'
id = 'advanced'
def ld_prune(ds: Dataset) -> Dataset:
# A potential mixed-backend workload:
# - Do pairwise calcs on GPU with CuPy
# - Do variant selection for those in high LD based on sparse arrays
# - Return selected indexes as Dask array with dense numpy chunks
...
def requirements() -> Sequence[Requirements]:
return [
# Packages are obvious requirments but this could also eventually include system resource constraints
Requirement('cupy', minimal_version='1.0'),
Requirement('sparse', minimal_version='0.5'),
Requirement('dask') # any version
]
class SimpleBackend(Backend):
domain = 'genetics.method'
id = 'simple'
def ld_prune(ds: Dataset) -> Dataset:
# Do everything assuming numpy
...
# As long as this step is isolated to the module the backend is defined in, it is
# easy to make sure that nothing is imported that is not installed:
frontend.register(AdvancedBackend())
frontend.register(SimpleBackend())
```
Usage then looks like this:
```
import api
ds: Dataset = ...
# Choose the best backend automatically based on environment, config, and arguments
# * To start, this is just an arbitrary choice based on which backends have installed dependencies
api.ld_prune(ds, backend='auto')
# Choose the backend explicitly
api.ld_prune(ds, backend='simple')
# OR explicitly within a block
with api.config.context('genetics.method.backend', 'simple'):
api.ld_prune(ds)
# OR explicitly globally
api.config.set('genetics.method.backend', 'simple')
api.ld_prune(ds)
```
| github_jupyter |
```
import pandas as pd
import numpy as np
from pathlib import Path
dPath = Path("../docs/dumps")
import pickle
with open(dPath / "train_data.pkl", 'rb') as filename:
train_data = pickle.load(filename)
with open(dPath / "valid_data.pkl", 'rb') as filename:
valid_data = pickle.load(filename)
X_train = train_data.drop("Detected", axis=1)
y_train = train_data.Detected
X_valid = valid_data.drop("Detected", axis=1)
y_valid = valid_data.Detected
with open(dPath / "rf_exp_03_names.pkl", 'rb') as filename:
names = pickle.load(filename)
X_train = X_train[names]
X_valid = X_valid[names]
X_train.head()
from imblearn.over_sampling import ADASYN
sm = ADASYN(random_state=42, n_jobs=-1)
%time X_res, y_res = sm.fit_resample(X_train, y_train)
X_train_not_running = not_running.drop("Detected", axis=1)
y_train_not_running = not_running.Detected
X_train_not_running = X_train_not_running[names]
#X_train_not_running['ExecutedRatio'] = -1
#X_res['ExecutedRatio'] = np.log(X_res.numExecuted)/X_res.ppn
X_train['ExecutedRatio'] = np.log(X_train.numExecuted)/X_train.ppnumberOfStatements
X_valid['ExecutedRatio'] = np.log(X_valid.numExecuted)/X_valid.ppnumberOfStatements
X_res = pd.concat([X_train, X_valid], ignore_index=True,axis=0)
y_res = pd.concat([y_train, y_valid], ignore_index=True,axis=0)
y_res.sum()/y_res.shape[0]
X_res.head()
from sklearn.ensemble import RandomForestClassifier
rf = RandomForestClassifier(n_estimators=100, max_features=0.7, oob_score=True, random_state=42, n_jobs=-1)
%time rf.fit(X_res, y_res)
from sklearn.metrics import roc_auc_score
from sklearn.metrics import classification_report
def print_score(m):
res = [m.score(X_train, y_train), m.score(X_valid, y_valid), roc_auc_score(y_valid, m.predict(X_valid))]
if hasattr(m, 'oob_score_'): res.append(m.oob_score_)
print(res)
def conf_matr(m):
y_pred = m.predict(X_valid)
print(classification_report(y_valid, y_pred))
print_score(rf)
conf_matr(rf)
with open(dPath / "rf_exp_10.pkl", 'wb') as filename:
pickle.dump(rf,filename)
names = X_train.columns
feat_importances = pd.Series(rf.feature_importances_, index=names)
feat_importances.sort_values(ascending=False,inplace=True)
feat_importances
feat_importances.to_excel(dPath / "proposed_approach_feat_importances.xlsx")
with open(dPath / "rf_exp_08_names.pkl", 'wb') as filename:
pickle.dump(names,filename)
with open(dPath / "test_data.pkl", 'rb') as filename:
test_data = pickle.load(filename)
X_test = test_data.drop("Detected", axis=1)
X_test['ExecutedRatio'] = np.log(X_test.numExecuted)/X_test.ppnumberOfStatements
y_test = test_data.Detected
X_test = X_test[names]
with open(dPath / "mutclslabels.pkl", 'rb') as filename:
mutclslabels = pickle.load(filename)
with open(dPath / "retypelabels.pkl", 'rb') as filename:
retypelabels = pickle.load(filename)
mutclscodes = dict(zip(mutclslabels,range(len(mutclslabels))))
retypecodes = dict(zip(retypelabels,range(len(retypelabels))))
X_test.replace(mutclscodes, inplace=True)
X_test.replace(retypecodes, inplace=True)
preds = rf.predict_proba(X_test)[:,1]
np.histogram(preds)
print(classification_report(y_test, preds >=0.50))
from sklearn.metrics import average_precision_score, precision_score, recall_score
print(average_precision_score(y_test, preds >= 0.5, average='micro'))
from sklearn import metrics
metrics.roc_auc_score(y_test, preds >= 0.5)
with open(dPath / "rf_exp_09_preds.pkl", 'wb') as filename:
pickle.dump(preds,filename)
```
| github_jupyter |
```
%matplotlib inline
%config InlineBackend.figure_format = "retina"
from matplotlib import rcParams
rcParams["savefig.dpi"] = 100
rcParams["figure.dpi"] = 100
rcParams["font.size"] = 20
import numpy as np
import matplotlib.pyplot as plt
np.random.seed(123)
# Choose the "true" parameters.
m_true = -0.9594
b_true = 4.294
# f_true = 0.534
from bioscrape.types import Model
from bioscrape.simulator import py_simulate_model
species = ['y']
parameters = {'m':m_true, 'b': b_true}
rule = ('assignment',{'equation':'y = _m*t + _b'})
x0 = {'y':0}
M = Model(species = species, parameters = parameters, rules = [rule], initial_condition_dict = x0)
#Simulate the Model deterministically
x0 = np.linspace(0, 10, 50)
results_det = py_simulate_model(x0, Model = M) #Returns a Pandas DataFrame
```
## Generate artificial data
```
# Generate some synthetic data from the model.
N = 50
x = np.sort(10 * np.random.rand(N))
yerr = 0.1 + 0.6 * np.random.rand(N)
y = m_true * x + b_true
# y += np.abs(f_true * y) * np.random.randn(N)
y += yerr * np.random.randn(N)
plt.errorbar(x, y, yerr=yerr, fmt=".k", capsize=0)
x0 = np.linspace(0, 10, 50)
plt.plot(x0, results_det['y'], "k", alpha=0.3, lw=3)
plt.xlim(0, 10)
plt.xlabel('x')
plt.ylabel('y');
```
## Create Pandas dataframes as required by bioscrape inference
```
import pandas as pd
exp_data = pd.DataFrame()
exp_data['x'] = x0
exp_data['y'] = y
exp_data
```
## Using bioscrape inference
```
# from bioscrape.pid_interfaces import
from bioscrape.inference import py_inference
# Import data from CSV
# Import a CSV file for each experiment run
prior = {'m' : ['gaussian', m_true, 500],'b' : ['gaussian', b_true, 1000]}
sampler, pid = py_inference(Model = M, exp_data = exp_data, measurements = ['y'],
time_column = ['x'], params_to_estimate = ['m','b'],
nwalkers = 32, nsteps = 2000, init_seed = 1e-4, prior = prior,
sim_type = 'deterministic', plot_show = False)
# Recommended to simply use sampler object/mcmc_results.csv and generate your own custom plots
truth_list, uncertainty_list = pid.plot_mcmc_results(sampler);
```
## Model simulations using parameter samples from the posterior distribution compared along side original model simulation
```
flat_samples = sampler.get_chain(discard=100, thin=15, flat=True)
inds = np.random.randint(len(flat_samples), size=100)
for ind in inds:
sample = flat_samples[ind]
plt.plot(x0, np.dot(np.vander(x0, 2), sample[:2]), "C1", alpha=0.1)
plt.errorbar(x, y, yerr=yerr, fmt=".k", capsize=0)
plt.plot(x0, m_true * x0 + b_true, "k", label="original model")
plt.legend(fontsize=14)
plt.xlim(0, 10)
plt.xlabel("x")
plt.ylabel("y");
```
# Other helpful utilities:
* Cost progress plots: You can look at the exploration of the log-likelihood by plotting cost_progress attribute of the inference object.
* The `plot_mcmc_results` returns the uncertainties around parameter values - this could indicate the accuracy of parameter inference predictions.
* You can use autocorrelation_time attribute of your inference object (pid.autocorrelation_time in the above example) to look at the autocorrelation time that it took for the chain to converge. A rule of thumb is that this value should be atleast 50 times lesser than the total number of steps for which you ran the MCMC chain. Higher autocorrelation time indicates that the chain was ended before it could converge to a distribution, whereas very low autocorrelation times could also be potentially something to look out for as it would indicate independence of all samples irrespective of the data.
* Check out Python emcee documentation on [**how to parallelize**](https://emcee.readthedocs.io/en/stable/tutorials/parallel/) the package using multiple CPU/GPU cores.
```
# rcParams['agg.path.chunksize'] = 10000
plt.close()
plt.figure()
plt.plot(pid.cost_progress)
plt.title('Log-likelihood progress')
plt.ylabel('Normed log-likelihood')
plt.xlabel('Number of steps in all chains')
plt.show()
pid.autocorrelation_time
```
| github_jupyter |
Monday, 21/06/2021
```
# ABS Function
import mysql.connector
import pandas as pd
conn = mysql.connector.connect(host ='localhost',
user ='root',
passwd ='***',
database ='dummy')
myquery = """
SELECT StudentID,FirstName,LastName,Semester1,Semester2, ABS(MarkGrowth) AS MarkGrowth FROM students;
"""
df = pd.read_sql(myquery,conn)
conn.close()
df
# CEILING Function
import mysql.connector
import pandas as pd
conn = mysql.connector.connect(host="localhost",
user = "root",
passwd = "***",
database = "dummy")
myquery = """
SELECT StudentID,FirstName,LastName,CEILING(Semester1) AS Semester1,CEILING(Semester2) AS Semester2, Markgrowth
FROM students;
"""
df = pd.read_sql(myquery,conn)
conn.close()
df
# FLOOR Function
import mysql.connector
import pandas as pd
conn = mysql.connector.connect(host="localhost",
user = "root",
passwd = "***",
database = "dummy")
myquery = """
SELECT StudentID,FirstName,LastName,FLOOR(Semester1) AS Semester1,FLOOR(Semester2) AS Semester2, Markgrowth
FROM students;
"""
df = pd.read_sql(myquery,conn)
conn.close()
df
# ROUND Function
import mysql.connector
import pandas as pd
conn = mysql.connector.connect(host="localhost",
user = "root",
passwd = "***",
database = "dummy")
myquery = """
SELECT StudentID,FirstName,LastName,ROUND(Semester1,1) AS Semester1,ROUND(Semester2,0) AS Semester2, Markgrowth
FROM students;
"""
df = pd.read_sql(myquery,conn)
conn.close()
df
# SQRT Function
import mysql.connector
import pandas as pd
conn = mysql.connector.connect(host="localhost",
user = "root",
passwd = "***",
database = "dummy")
myquery = """
SELECT StudentID,FirstName,LastName,SQRT(Semester1) AS Semester1,Semester2, Markgrowth
FROM students;
"""
df = pd.read_sql(myquery,conn)
conn.close()
df
# MOD and EXP Function
import mysql.connector
import pandas as pd
conn = mysql.connector.connect(host="localhost",
user = "root",
passwd = "***",
database = "dummy")
myquery = """
SELECT StudentID,FirstName,LastName,MOD(Semester1,2) AS Semester1,Semester2, EXP(Markgrowth)
FROM students;
"""
df = pd.read_sql(myquery,conn)
conn.close()
df
# CONCAT Function
import mysql.connector
import pandas as pd
conn = mysql.connector.connect(host="localhost",
user = "root",
passwd = "***",
database = "dummy")
myquery = """
SELECT StudentID,CONCAT(FirstName," ",LastName) AS Name,SQRT(Semester1) AS Semester1,Semester2,MarkGrowth
FROM students;
"""
df = pd.read_sql(myquery,conn)
conn.close()
df
# SUBSTRING_INDEX Function
import mysql.connector
import pandas as pd
conn = mysql.connector.connect(host="localhost",
user = "root",
passwd = "***",
database = "dummy")
myquery = """
SELECT StudentID,SUBSTRING_INDEX(Email,'@',1) AS Name
FROM students;
"""
df = pd.read_sql(myquery,conn)
conn.close()
df
# SUBSTR Function
import mysql.connector
import pandas as pd
conn = mysql.connector.connect(host="localhost",
user = "root",
passwd = "***",
database = "dummy")
myquery = """
SELECT StudentID,SUBSTR(FirstName,2,4) AS Initial_Name
FROM students;
"""
df = pd.read_sql(myquery,conn)
conn.close()
df
# LENGTH Function
import mysql.connector
import pandas as pd
conn = mysql.connector.connect(host="localhost",
user = "root",
passwd = "***",
database = "dummy")
myquery = """
SELECT StudentID,FirstName,LENGTH(FirstName) AS Total_Char
FROM students;
"""
df = pd.read_sql(myquery,conn)
conn.close()
df
# REPLACE Function
import mysql.connector
import pandas as pd
conn = mysql.connector.connect(host="localhost",
user = "root",
passwd = "***",
database = "dummy")
myquery = """
SELECT StudentID,Email,REPLACE(Email,'yahoo','gmail') AS New_Email
FROM students;
"""
df = pd.read_sql(myquery,conn)
conn.close()
df
# UPPER and LOWER Function
import mysql.connector
import pandas as pd
conn = mysql.connector.connect(host="localhost",
user = "root",
passwd = "***",
database = "dummy")
myquery = """
SELECT StudentID,UPPER(FirstName) AS FirstName, LOWER(LastName) AS LastName
FROM students;
"""
df = pd.read_sql(myquery,conn)
conn.close()
df
```
## For more text scalar function :
https://dev.mysql.com/doc/refman/8.0/en/string-functions.html
## For more mathematical scalar function :
https://dev.mysql.com/doc/refman/8.0/en/mathematical-functions.html
```
# SUM Function
import mysql.connector
import pandas as pd
conn = mysql.connector.connect(host="localhost",
user = "root",
passwd = "***",
database = "dummy")
myquery = """
SELECT SUM(Semester1) AS Total_1, SUM(Semester2) AS Total_2
FROM students;
"""
df = pd.read_sql(myquery,conn)
conn.close()
df
# COUNT Function
import mysql.connector
import pandas as pd
conn = mysql.connector.connect(host="localhost",
user = "root",
passwd = "***",
database = "dummy")
myquery = """
SELECT COUNT(FirstName) AS Total_Student
FROM students;
"""
df = pd.read_sql(myquery,conn)
conn.close()
df
# AVG Function
import mysql.connector
import pandas as pd
conn = mysql.connector.connect(host="localhost",
user = "root",
passwd = "***",
database = "dummy")
myquery = """
SELECT AVG(Semester1) AS AVG_1, AVG(Semester2) AS AVG_2, AVG(MarkGrowth) AS GROWTH_AVG
FROM students;
"""
df = pd.read_sql(myquery,conn)
conn.close()
df
# MIN and MAX Function
import mysql.connector
import pandas as pd
conn = mysql.connector.connect(host="localhost",
user = "root",
passwd = "***",
database = "dummy")
myquery = """
SELECT MIN(Semester1) AS Min1, MAX(Semester1) AS Max1, AVG(Semester1) AS AVG_1
FROM students;
"""
df = pd.read_sql(myquery,conn)
conn.close()
df
# GROUP BY Function
import mysql.connector
import pandas as pd
conn = mysql.connector.connect(host="localhost",
user = "root",
passwd = "***",
database = "sakila")
myquery = """
SELECT staff_id,
COUNT(payment_id) AS total_payment,
SUM(amount) AS total_amount
FROM payment
GROUP BY staff_id;
"""
df = pd.read_sql(myquery,conn)
conn.close()
df
# GROUP BY Function for Multiple Columns
import mysql.connector
import pandas as pd
conn = mysql.connector.connect(host="localhost",
user = "root",
passwd = "***",
database = "sakila")
myquery = """
SELECT country,
notes,
COUNT(DISTINCT id) AS total_customer
FROM customer_list
WHERE notes = 'active'
GROUP BY country,notes;
"""
df = pd.read_sql(myquery,conn)
conn.close()
df
# CASE ... END Statement
import mysql.connector
import pandas as pd
conn = mysql.connector.connect(host="localhost",
user = "root",
passwd = "***",
database = "sakila")
myquery = """
SELECT title,
rental_duration,
CASE
WHEN rental_duration > 5 THEN "Famous"
WHEN rental_duration < 5 THEN "Grow Up"
ELSE "Follow Up"
END as remark
FROM film;
"""
df = pd.read_sql(myquery,conn)
conn.close()
df
```
Learning Source : www.dqlab.id
| github_jupyter |
```
# data from https://github.com/cbaziotis/ekphrasis/blob/master/ekphrasis/utils/helpers.py
# reuploaded to husein's S3
# !wget https://malaya-dataset.s3-ap-southeast-1.amazonaws.com/counts_1grams.txt
import os
os.environ['CUDA_VISIBLE_DEVICES'] = ''
with open('counts_1grams.txt') as fopen:
f = fopen.read().split('\n')[:-1]
words = {}
for l in f:
w, c = l.split('\t')
c = int(c)
words[w] = c + words.get(w, 0)
# original from https://github.com/cbaziotis/ekphrasis/blob/master/ekphrasis/classes/spellcorrect.py
# improved it
import re
from collections import Counter
class SpellCorrector:
"""
The SpellCorrector extends the functionality of the Peter Norvig's
spell-corrector in http://norvig.com/spell-correct.html
"""
def __init__(self):
"""
:param corpus: the statistics from which corpus to use for the spell correction.
"""
super().__init__()
self.WORDS = words
self.N = sum(self.WORDS.values())
@staticmethod
def tokens(text):
return REGEX_TOKEN.findall(text.lower())
def P(self, word):
"""
Probability of `word`.
"""
return self.WORDS[word] / self.N
def most_probable(self, words):
_known = self.known(words)
if _known:
return max(_known, key=self.P)
else:
return []
@staticmethod
def edit_step(word):
"""
All edits that are one edit away from `word`.
"""
letters = 'abcdefghijklmnopqrstuvwxyz'
splits = [(word[:i], word[i:]) for i in range(len(word) + 1)]
deletes = [L + R[1:] for L, R in splits if R]
transposes = [L + R[1] + R[0] + R[2:] for L, R in splits if len(R) > 1]
replaces = [L + c + R[1:] for L, R in splits if R for c in letters]
inserts = [L + c + R for L, R in splits for c in letters]
return set(deletes + transposes + replaces + inserts)
def edits2(self, word):
"""
All edits that are two edits away from `word`.
"""
return (e2 for e1 in self.edit_step(word)
for e2 in self.edit_step(e1))
def known(self, words):
"""
The subset of `words` that appear in the dictionary of WORDS.
"""
return set(w for w in words if w in self.WORDS)
def edit_candidates(self, word, assume_wrong=False, fast=True):
"""
Generate possible spelling corrections for word.
"""
if fast:
ttt = self.known(self.edit_step(word)) or {word}
else:
ttt = self.known(self.edit_step(word)) or self.known(self.edits2(word)) or {word}
ttt = self.known([word]) | ttt
return list(ttt)
corrector = SpellCorrector()
possible_states = corrector.edit_candidates('eting')
possible_states
# !wget https://storage.googleapis.com/bert_models/2018_10_18/uncased_L-12_H-768_A-12.zip
# !unzip uncased_L-12_H-768_A-12.zip
BERT_VOCAB = 'uncased_L-12_H-768_A-12/vocab.txt'
BERT_INIT_CHKPNT = 'uncased_L-12_H-768_A-12/bert_model.ckpt'
BERT_CONFIG = 'uncased_L-12_H-768_A-12/bert_config.json'
import bert
from bert import run_classifier
from bert import optimization
from bert import tokenization
from bert import modeling
import tensorflow as tf
import numpy as np
tokenization.validate_case_matches_checkpoint(True,BERT_INIT_CHKPNT)
tokenizer = tokenization.FullTokenizer(
vocab_file=BERT_VOCAB, do_lower_case=True)
text = 'scientist suggests eting berger can lead to obesity'
text_mask = text.replace('eting', '**mask**')
text_mask
def tokens_to_masked_ids(tokens, mask_ind):
masked_tokens = tokens[:]
masked_tokens[mask_ind] = "[MASK]"
masked_tokens = ["[CLS]"] + masked_tokens + ["[SEP]"]
masked_ids = tokenizer.convert_tokens_to_ids(masked_tokens)
return masked_ids
bert_config = modeling.BertConfig.from_json_file(BERT_CONFIG)
class Model:
def __init__(
self,
):
self.X = tf.placeholder(tf.int32, [None, None])
model = modeling.BertModel(
config=bert_config,
is_training=False,
input_ids=self.X,
use_one_hot_embeddings=False)
output_layer = model.get_sequence_output()
embedding = model.get_embedding_table()
with tf.variable_scope('cls/predictions'):
with tf.variable_scope('transform'):
input_tensor = tf.layers.dense(
output_layer,
units = bert_config.hidden_size,
activation = modeling.get_activation(bert_config.hidden_act),
kernel_initializer = modeling.create_initializer(
bert_config.initializer_range
),
)
input_tensor = modeling.layer_norm(input_tensor)
output_bias = tf.get_variable(
'output_bias',
shape = [bert_config.vocab_size],
initializer = tf.zeros_initializer(),
)
logits = tf.matmul(input_tensor, embedding, transpose_b = True)
self.logits = tf.nn.bias_add(logits, output_bias)
tf.reset_default_graph()
sess = tf.InteractiveSession()
model = Model()
sess.run(tf.global_variables_initializer())
var_lists = tf.get_collection(tf.GraphKeys.TRAINABLE_VARIABLES, scope = 'bert')
cls = tf.get_collection(tf.GraphKeys.TRAINABLE_VARIABLES, scope = 'cls')
cls
saver = tf.train.Saver(var_list = var_lists + cls)
saver.restore(sess, BERT_INIT_CHKPNT)
replaced_masks = [text_mask.replace('**mask**', state) for state in possible_states]
replaced_masks
tokens = tokenizer.tokenize(replaced_masks[0])
input_ids = [tokens_to_masked_ids(tokens, i) for i in range(len(tokens))]
input_ids
tokens_ids = tokenizer.convert_tokens_to_ids(tokens)
tokens_ids
def generate_ids(mask):
tokens = tokenizer.tokenize(mask)
input_ids = [tokens_to_masked_ids(tokens, i) for i in range(len(tokens))]
tokens_ids = tokenizer.convert_tokens_to_ids(tokens)
return tokens, input_ids, tokens_ids
ids = [get_score(mask) for mask in replaced_masks]
tokens, input_ids, tokens_ids = list(zip(*ids))
indices, ids = [], []
for i in range(len(input_ids)):
indices.extend([i] * len(input_ids[i]))
ids.extend(input_ids[i])
ids[0]
masked_padded = tf.keras.preprocessing.sequence.pad_sequences(ids,padding='post')
masked_padded.shape
preds = sess.run(tf.nn.log_softmax(model.logits), feed_dict = {model.X: masked_padded})
preds.shape
indices = np.array(indices)
scores = []
for i in range(len(tokens)):
filter_preds = preds[indices == i]
total = np.sum([filter_preds[k, k + 1, x] for k, x in enumerate(tokens_ids[i])])
scores.append(total)
scores
prob_scores = np.array(scores) / np.sum(scores)
prob_scores
probs = list(zip(possible_states, prob_scores))
probs.sort(key = lambda x: x[1])
probs
```
| github_jupyter |
# Gluon CIFAR-10 Trained in Local Mode
_**ResNet model in Gluon trained locally in a notebook instance**_
---
---
_This notebook was created and tested on an ml.p3.8xlarge notebook instance._
## Setup
Import libraries and set IAM role ARN.
```
import sagemaker
from sagemaker.mxnet import MXNet
sagemaker_session = sagemaker.Session()
role = sagemaker.get_execution_role()
```
Install pre-requisites for local training.
```
!/bin/bash setup.sh
```
---
## Data
We use the helper scripts to download CIFAR-10 training data and sample images.
```
from cifar10_utils import download_training_data
download_training_data()
```
We use the `sagemaker.Session.upload_data` function to upload our datasets to an S3 location. The return value `inputs` identifies the location -- we will use this later when we start the training job.
Even though we are training within our notebook instance, we'll continue to use the S3 data location since it will allow us to easily transition to training in SageMaker's managed environment.
```
inputs = sagemaker_session.upload_data(path='data', key_prefix='data/DEMO-gluon-cifar10')
print('input spec (in this case, just an S3 path): {}'.format(inputs))
```
---
## Script
We need to provide a training script that can run on the SageMaker platform. When SageMaker calls your function, it will pass in arguments that describe the training environment. Check the script below to see how this works.
The network itself is a pre-built version contained in the [Gluon Model Zoo](https://mxnet.incubator.apache.org/versions/master/api/python/gluon/model_zoo.html).
```
!cat 'cifar10.py'
```
---
## Train (Local Mode)
The ```MXNet``` estimator will create our training job. To switch from training in SageMaker's managed environment to training within a notebook instance, just set `train_instance_type` to `local_gpu`.
```
m = MXNet('cifar10.py',
role=role,
train_instance_count=1,
train_instance_type='local_gpu',
framework_version='1.1.0',
hyperparameters={'batch_size': 1024,
'epochs': 50,
'learning_rate': 0.1,
'momentum': 0.9})
```
After we've constructed our `MXNet` object, we can fit it using the data we uploaded to S3. SageMaker makes sure our data is available in the local filesystem, so our training script can simply read the data from disk.
```
m.fit(inputs)
```
---
## Host
After training, we use the MXNet estimator object to deploy an endpoint. Because we trained locally, we'll also deploy the endpoint locally. The predictor object returned by `deploy` lets us call the endpoint and perform inference on our sample images.
```
predictor = m.deploy(initial_instance_count=1, instance_type='local_gpu')
```
### Evaluate
We'll use these CIFAR-10 sample images to test the service:
<img style="display: inline; height: 32px; margin: 0.25em" src="images/airplane1.png" />
<img style="display: inline; height: 32px; margin: 0.25em" src="images/automobile1.png" />
<img style="display: inline; height: 32px; margin: 0.25em" src="images/bird1.png" />
<img style="display: inline; height: 32px; margin: 0.25em" src="images/cat1.png" />
<img style="display: inline; height: 32px; margin: 0.25em" src="images/deer1.png" />
<img style="display: inline; height: 32px; margin: 0.25em" src="images/dog1.png" />
<img style="display: inline; height: 32px; margin: 0.25em" src="images/frog1.png" />
<img style="display: inline; height: 32px; margin: 0.25em" src="images/horse1.png" />
<img style="display: inline; height: 32px; margin: 0.25em" src="images/ship1.png" />
<img style="display: inline; height: 32px; margin: 0.25em" src="images/truck1.png" />
```
# load the CIFAR-10 samples, and convert them into format we can use with the prediction endpoint
from cifar10_utils import read_images
filenames = ['images/airplane1.png',
'images/automobile1.png',
'images/bird1.png',
'images/cat1.png',
'images/deer1.png',
'images/dog1.png',
'images/frog1.png',
'images/horse1.png',
'images/ship1.png',
'images/truck1.png']
image_data = read_images(filenames)
```
The predictor runs inference on our input data and returns the predicted class label (as a float value, so we convert to int for display).
```
for i, img in enumerate(image_data):
response = predictor.predict(img)
print('image {}: class: {}'.format(i, int(response)))
```
---
## Cleanup
After you have finished with this example, remember to delete the prediction endpoint. Only one local endpoint can be running at a time.
```
m.delete_endpoint()
```
| github_jupyter |
```
#Let's start by importing the most important libraries. We will primarily use Pandas and Numpy\
#, some modules from NLTK, and re for regular expressions
import pandas as pd;
import numpy as np;
import re;
import time;
from nltk.corpus import stopwords;
import json;
import sys;
import pickle;
#Fill in the data location, which has been obtained from https://www.kaggle.com/bittlingmayer/amazonreviews/data
data_location = 'input/amazon_review_full_csv/';
#Load in the train and test sets. No header.
data_train = pd.read_csv(data_location + 'train.csv', header=None);
data_test = pd.read_csv(data_location + 'test.csv', header=None);
#Update the headers with the right column names
data_train.columns = ['rating', 'subject', 'review'];
data_test.columns = ['rating', 'subject', 'review'];
print(len(data_train));
print(len(data_test));
#There are a lot of rows in both the train and test sets, so to make processing easier, we will
#use a small subset of the rows to verify that the pipeline works without errors.
#We can do the final training on an AWS server.
#First, we need to get a subset of indices. The sampling dataset
#will be divided into the following ratio: 75% TRAIN, 20% DEV, 5% TEST
np.random.seed(1024);
total_samples = int(0.06 * len(data_train))
rand_indices = np.random.choice(len(data_train), total_samples, replace=False);
train_split_index = int(0.75 * total_samples);
dev_split_index = int(0.95 * total_samples);
data_sample_train = data_train.iloc[rand_indices[:train_split_index]];
data_sample_dev = data_train.iloc[rand_indices[train_split_index:dev_split_index]];
data_sample_test = data_train.iloc[rand_indices[dev_split_index:]];
sample_ratio = len(data_train) / len(data_sample_train);
print("How much of the data we are using: " + str(100.0 / sample_ratio) + '%')
#Next step is cleaning. We will go through different steps, and in the end combine them into a single function. Because we
#are only writing the functions to build the pipeline, I will use a small subset of the data sample to verify it.
data_subsample = data_sample_train.iloc[0:100];
#First step: Remove all items that have a Neutral (3 star) rating
data_sub_filtered = data_subsample[data_subsample.rating != 3];
rows_removed = len(data_subsample) - len(data_sub_filtered);
print('Removing ' + str(100.0 * rows_removed / len(data_subsample)) + '% of rows');
#Second step: Binary ratings. 0 for {1, 2} and 1 for {4, 5}
data_sub_filtered.loc[data_sub_filtered.rating <= 2, 'rating'] = 0;
data_sub_filtered.loc[data_sub_filtered.rating >= 4, 'rating'] = 1;
#Third step: Remove all NaNs.
#We have enough rows in our dataset that we can safely remove all rows with NaNs and still have enough data.
rows_before = len(data_sub_filtered);
data_sub_filtered = data_sub_filtered.dropna();
print('Removed ' + str(100*(len(data_sub_filtered)-rows_before) / len(data_sub_filtered)) + '% of rows');
#I don't understand the warning above, but it looks like the data was properly processed
#Llet's look at a few rows of the data so far
data_sub_filtered.iloc[0:5]
#Fourth step: Remove all symbols such as ! % & * @ #
#To do this, I will use a regular expression that only keeps alphanumerics and punctuation symbols.
pattern_to_find = "[^a-zA-Z0-9' \.,!\?]";
pattern_to_repl = "";
start = time.time();
for row in data_sub_filtered.index:
data_sub_filtered.loc[row, 'subject'] = re.sub(pattern_to_find, pattern_to_repl, data_sub_filtered.loc[row, 'subject']).lower();
data_sub_filtered.loc[row, 'review'] = re.sub(pattern_to_find, pattern_to_repl, data_sub_filtered.loc[row, 'review']).lower();
print('Total time taken: ' + str(time.time() - start) + 's');
#Fifth step: Remove alll urls from the text
#ref: https://stackoverflow.com/a/6883094/1843486
start = time.time();
url_regex = 'http[s]?://(?:[a-zA-Z]|[0-9]|[$-_@.&+]|[!*\(\),]|(?:%[0-9a-fA-F][0-9a-fA-F]))+';
for row in data_sub_filtered.index:
data_sub_filtered.loc[row, 'subject'] = re.sub(url_regex, '', data_sub_filtered.loc[row, 'subject']);
data_sub_filtered.loc[row, 'review'] = re.sub(url_regex, '', data_sub_filtered.loc[row, 'review']);
print('Total time taken: ' + str(time.time() - start) + 's');
#Sixth step: Expand all contractions. This will reduce disambiguation between similar phrases such as I'll and I will.
#ref: https://stackoverflow.com/a/19794953/1843486
#I placed all contraction mappings in a text json file, which we can load in
#and look at a few values to see what we have
contractions = json.load(open('contractions.json', 'r'));
for pos, key in enumerate(contractions.keys()):
print(key, ' -> ', contractions[key]);
if pos > 5:
break;
#This is a little tricky, and not completely perfect. There are two types of contractions I will be looking at
#of the form < is'nt > and of the form < apple's >. The first can be mapped from the json dict,
#but the second form cannot because apple is a proper noun. For all instances of the second, I will just expand it to < apple is >
start = time.time();
apos_regex = "'[a-z]+|[a-z]+'[a-z]+|[a-z]+'";
#A function allows us to iterate over quickly rather than having a complicated lambda expression
def expand(word):
if "'" in word:
if word in contractions:
return contractions[word]
if word.endswith("'s"):
return word[:-2] + " is"
else:
return word;
else:
return word;
for row in data_sub_filtered.index:
data_sub_filtered.loc[row, 'subject'] = ' '.join(([expand(word) for word in data_sub_filtered.loc[row, 'subject'].split()]));
data_sub_filtered.loc[row, 'review'] = ' '.join(([expand(word) for word in data_sub_filtered.loc[row, 'review'].split()]));
print('Total time taken: ' + str(time.time() - start) + 's');
#Seventh step: Remove all stopwords such as and but if so then
eng_stopwords = set(stopwords.words("english"));
start = time.time();
for row in data_sub_filtered.index:
text_subj = data_sub_filtered.loc[row, 'subject'].split();
text_revi = data_sub_filtered.loc[row, 'review'].split();
data_sub_filtered.loc[row, 'subject'] = ' '.join([word for word in text_subj if word not in eng_stopwords]);
data_sub_filtered.loc[row, 'review'] = ' '.join([word for word in text_revi if word not in eng_stopwords]);
print('Total time taken: ' + str(time.time() - start) + 's');
#Now that all of the different processes have been completed, it is time to combine them all into a single function.
#The process_sentence method will process a single sentence \
#and return a version without URLs, symbols, contractions and stopwords.
def process_sentence(sentence):
#Remove special symbols and lowercase everything
alphanum = re.sub(pattern_to_find, pattern_to_repl, sentence).lower();
#Remove URLs
nourls = re.sub(url_regex, '', alphanum);
#Expand all contractions
noapos = ' '.join(([expand(word) for word in nourls.split()]));
#Remove stopwords
bigwords = ' '.join([word for word in noapos.split() if word not in eng_stopwords]);
return bigwords;
#The clean_data method will take in a dataframe, filter out neutral rows, binarize ratings, remove na's, and process rows.
def clean_data(dframe):
start = time.time();
dframe = dframe[dframe.rating != 3]
dframe.loc[dframe.rating <= 2, 'rating'] = 0;
dframe.loc[dframe.rating >= 4, 'rating'] = 1;
dframe = dframe.dropna();
for pos, row in enumerate(dframe.index):
dframe.loc[row, 'subject'] = process_sentence(dframe.loc[row, 'subject']);
dframe.loc[row, 'review'] = process_sentence(dframe.loc[row, 'review']);
if pos % 1000 == 0 and pos > 0:
time_so_far = (time.time() - start)/60;
time_eta = time_so_far * (len(dframe) / pos) - time_so_far;
sys.stdout.write("\rCompleted " + str(pos) + " / " + str(len(dframe)) + " in " + str(time_so_far) + "m eta: " + str(time_eta) + 'm');
print('\n')
print('Total time taken: ' + str(time.time() - start) + 's');
return dframe;
data_sub_processed = clean_data(data_subsample)
#sanity check
data_sub_processed.equals(data_sub_filtered)
data_train_processed = clean_data(data_sample_train);
data_dev_processed = clean_data(data_sample_dev);
data_test_processed = clean_data(data_sample_test);
pickle.dump(data_train_processed, open("picklefiles/data_train_processed.pkl", 'wb'));
pickle.dump(data_train_processed, open("picklefiles/data_dev_processed.pkl", 'wb'))
pickle.dump(data_train_processed, open("picklefiles/data_test_processed.pkl", 'wb'))
#Now that we have our data, the next step is to obtain features. I will be training a
#convolutional neural network, for which we need vectorized input. One way to do this
#is to define each word with a Word2Vec model.
#If you read my previous post on Word2Vec, the model is a shallow neural network trained
#to learn word embeddings. We will be learning features with the Word2Vec model first,
#and then using the features generated from it as inputs to the convolutional neural network.
```
| github_jupyter |
```
from tars import Tars, markets, portfolios, traders, strategies
%matplotlib inline
```
# Getting Started
Welcome on Tars, the coolest crypto trading bot for research purposes. Here's a great first guide to follow to get accustomed with the different module available in Tars.
We'll get familiar with :
1. Tars' concepts
2. the way to make the bot trade 24/7
3. the tutorials
Have a good read!
---
## Concept
The goal of Tars is to allow developers to create trading strategies with ease. The concept is composed of the following building blocks :
- market
- portfolio
- trader
- strategy
The idea behind this decomposition is to follow the nature of the trading activity. A **strategy** is used by a **trader** to manage a **portfolio** on a **market**.
When it makes sense, each object has a test version and a real version. So that it's possible to trade without financial risk.
Once you have those four components, you can give them to **Tars** so that he can run them.
### Market object
Allows to get the latest data from the market. Currently, only the cryptocurrency market is available. To get the latest OHLC data for ETHUSD :
```
markets.CryptoMarket().get_ohlc_data('XETHZUSD')[0].tail()
```
### Portfolio object
Represents the portfolio of currencies available for trading. Two objects are at your disposal :
- VirtualPortfolio -> for testing purposed
- CryptoPortfolio -> your real Kraken Exchange portfolio
```
# Create a virtual portfolio
portfolio = portfolios.VirtualPortfolio(
{'ZUSD': 1000,
'XETH': 30}
)
# Get the account balance
portfolio.get_account_balance()
# Get the total available amount for trading
portfolio.get_trade_balance()
```
### Trader object
Trader is the object allowing to make orders, cancel them, etc. To create a cryptocurrency trader, you also have two objects:
- VirtualCryptoTrader -> for paper trading
- CryptoTrader -> connects to Kraken Exchange for real transactions
```
# Create the virtual cryptocurrency trader
trader = traders.VirtualCryptoTrader(portfolio)
# Add an order. Here, buy some ETH
trader.add_order(
pair='XETHZUSD',
type='buy',
ordertype='market',
volume=0.2,
validate=True)
# You can trace back the actions of a trader to see what he did
trader.get_trades_history()[0]
```
### Strategy object
This is the most interesting, this package allows you to choose different ways to trade. Among them:
- BuyAndHold
- RandomInvestment
- SequentialInvestment
- TrendFollowingMACD
- and more...
For instance, to create the simplest `BuyAndHold` strategy, just do :
```
strategy = strategies.BuyAndHold(
trader,
'XETHZUSD',
0.2)
```
And don't forget to call for help when you're unsure about the signature needed for a strategy, they are sometimes different.
```
help(strategies.BuyAndHold)
```
#### Create your own strategy
It's super simple to create your own new strategy and I encourage you
to give it a shot!
To do it, you just need to inherit from
the `AbstractStrategy`and implement a `run` function without parameters. You can check the different existing strategies to see how to do it if you have a doubt.
---
## Run a strategy with Tars
```
tars = Tars()
tars.load(strategy)
```
### Start the bot
```
tars.start('00:01:00')
```
### Evaluate the strategy
```
tars.plot()
tars.evaluate()
```
### Stop the bot
```
tars.stop()
```
### Trade with real money
To do that, just change the portfolio and trader objects as follow:
- VirtualPortfolio should become CryptoPortfolio
- VirtualCryptoTrader should become CryptoTrader
Update the boolean `validating` trader arguments to `False`, run Tars and let him do his job!
---
## What else?
Some tutorials are available in the `notebooks/tutorials`folder. So far:
- One about historical data loading
- Trading for real
- Trading virtually
### Any Questions?
If you have any questions, feel free to contact me or put an issue on GitHub.
---
| github_jupyter |
## Sean Lahman’s Baseball Database
Our team of researchers has integrated playing statistics from the 2012 season. The updated version contains complete batting and pitching statistics back to 1871, plus fielding statistics, standings, team stats, managerial records, post-season data, and more. For more details on the latest release, please read the documentation.
The database can be used on any platform, but please be aware that this is not a standalone application. It is a database that requires Microsoft Access or some other relational database software to be useful.
Please help support the Baseball Archive. The database is free, but there are real costs associated with maintaining it and making it available for download. The more popular this site becomes, the more expensive it is to keep things going. Please consider making a donation as a show of your support. Like the PBS folks say, we need your support if we’re going to survive. Click here for more information.
### Limited Use License
This database is copyright 1996-2013 by Sean Lahman.
This work is licensed under a Creative Commons Attribution-ShareAlike 3.0 Unported License. For details see: http://creativecommons.org/licenses/by-sa/3.0/
### The 2012 Version
* [2012 Version - Microsoft Access](http://seanlahman.com/files/database/lahman2012-ms.zip)
* [2012 Version - comma-delimited version](http://seanlahman.com/files/database/lahman2012-csv.zip)
* [2012 Version - SQL version](http://seanlahman.com/files/database/lahman2012-sql.zip)
Files last updated January 9, 3:00 pm
## Documentation: The Lahman Baseball Database
2012 Version
Release Date: December 31, 2012
### CONTENTS
* 0.1 Copyright Notice
* 0.2 Contact Information
* 1.0 Release Contents
* 1.1 Introduction
* 1.2 What's New
* 1.3 Acknowledgements
* 1.4 Using this Database
* 1.5 Revision History
* 2.0 Data Tables
* 2.1 MASTER table
* 2.2 Batting Table
* 2.3 Pitching table
* 2.4 Fielding Table
* 2.5 All-Star table
* 2.6 Hall of Fame table
* 2.7 Managers table
* 2.8 Teams table
* 2.9 BattingPost table
* 2.10 PitchingPost table
* 2.11 TeamFranchises table
* 2.12 FieldingOF table
* 2.13 ManagersHalf table
* 2.14 TeamsHalf table
* 2.15 Salaries table
* 2.16 SeriesPost table
* 2.17 AwardsManagers table
* 2.18 AwardsPlayers table
* 2.19 AwardsShareManagers table
* 2.20 AwardsSharePlayers table
* 2.21 FieldingPost table
* 2.22 Appearances table
* 2.23 Schools table
* 2.24 SchoolsPlayers table
### 0.1 Copyright Notice & Limited Use License
This database is copyright 1996-2013 by Sean Lahman.
This work is licensed under a Creative Commons Attribution-ShareAlike 3.0 Unported License. For details see: http://creativecommons.org/licenses/by-sa/3.0/
For licensing information or further information, contact Sean Lahman
at: seanlahman@gmail.com
### 0.2 Contact Information
Web site: http://www.baseball1.com
E-Mail : seanlahman@gmail.com
If you're interested in contributing to the maintenance of this
database or making suggestions for improvement, please consider
joining our mailinglist at:
http://groups.yahoo.com/group/baseball-databank/
If you are interested in similar databases for other sports, please
vist the Open Source Sports website at http://OpenSourceSports.com
### 1.0 Release Contents
This release of the database can be downloaded in several formats. The
contents of each version are listed below.
**MS Access Versions**:
lahman2012.mdb
2012readme.txt
**SQL Version**:
lahman2012.sql
2012readme.txt
**Comma Delimited Version**:
2012readme.txt
AllStarFull.csv
Appearances.csv
AwardsManagers.csv
AwardsPlayers.csv
AwardsShareManagers.csv
AwardsSharePlayers.csv
Batting.csv
BattingPost.csv
Fielding.csv
FieldingOF.csv
FieldingPost.csv
HallOfFame.csv
Managers.csv
ManagersHalf.csv
Master.csv
Pitching.csv
PitchingPost.csv
Salaries.csv
Schools.csv
SchoolsPlayers.csv
SeriesPost.csv
Teams.csv
TeamsFranchises.csv
TeamsHalf.csv
### 1.1 Introduction
This database contains pitching, hitting, and fielding statistics for
Major League Baseball from 1871 through 2012. It includes data from
the two current leagues (American and National), the four other "major"
leagues (American Association, Union Association, Players League, and
Federal League), and the National Association of 1871-1875.
This database was created by Sean Lahman, who pioneered the effort to
make baseball statistics freely available to the general public. What
started as a one man effort in 1994 has grown tremendously, and now a
team of researchers have collected their efforts to make this the
largest and most accurate source for baseball statistics available
anywhere. (See Acknowledgements below for a list of the key
contributors to this project.)
None of what we have done would have been possible without the
pioneering work of Hy Turkin, S.C. Thompson, David Neft, and Pete
Palmer (among others). All baseball fans owe a debt of gratitude
to the people who have worked so hard to build the tremendous set
of data that we have today. Our thanks also to the many members of
the Society for American Baseball Research who have helped us over
the years. We strongly urge you to support and join their efforts.
Please vist their website (www.sabr.org).
This database can never take the place of a good reference book like
The Baseball Encyclopedia. But it will enable people do to the kind
of queries and analysis that those traditional sources don't allow.
If you have any problems or find any errors, please let us know. Any
feedback is appreciated
### 1.2 What's New in 2012
There has been significant cleanup in the master file
MLB's addition of wildcard games in 2012 adds two new types of records
to the post-season files. The abbreviations ALWC and NLWC are used to
denote each league's wild card game.
Added the MLB "Comeback Player of the Year" award to the awards table
Florida Marlins changed their name to the Miami Marlins, new team abbr is MIA
### 1.3 Acknowledgements
Much of the raw data contained in this database comes from the work of
Pete Palmer, the legendary statistician, who has had a hand in most
of the baseball encylopedias published since 1974. He is largely
responsible for bringing the batting, pitching, and fielding data out
of the dark ages and into the computer era. Without him, none of this
would be possible. For more on Pete's work, please read his own
account at: http://sabr.org/cmsfiles/PalmerDatabaseHistory.pdf
Two people have been key contributors to the work that followed, first
by taking the raw data and creating a relational database, and later
by extending the database to make it more accesible to researchers.
Sean Lahman launched the Baseball Archive's website back before
most people had heard of the world wide web. Frustrated by the
lack of sports data available, he led the effort to build a
baseball database that everyone could use. Baseball researchers
everywhere owe him a debt of gratitude. Lahman served as an associate
editor for three editions of Total Baseball and contributed to five
editions of The ESPN Baseball Encyclopedia. He has also been active in
developing databases for other sports.
The work of Sean Forman to create and maintain an online encyclopedia
at "baseball-reference.com" has been remarkable. Recognized as the
premier online reference source, Forman's site provides an oustanding
interface to the raw data. His efforts to help streamline the database
have been extremely helpful. Most importantly, Forman has spearheaded
the effort to provide standards that enable several different baseball
databases to be used together. He was also instrumental in launching
the Baseball Databank, a forum for researchers to gather and share
their work.
Since 2001, these two Seans have led a group of researchers
who volunteered to maintain and update the database.
A handful of researchers have made substantial contributions to
maintain this database in recent years. Listed alphabetically, they
are: Derek Adair, Mike Crain, Kevin Johnson, Rod Nelson, Tom Tango,
and Paul Wendt. These folks did much of the heavy lifting, and are
largely responsible for the improvements made in the last decade.
Others who made important contributions include: Dvd Avins,
Clifford Blau, Bill Burgess, Clem Comly, Jeff Burk, Randy Cox,
Mitch Dickerman, Paul DuBois, Mike Emeigh, F.X. Flinn, Bill Hickman,
Jerry Hoffman, Dan Holmes, Micke Hovmoller, Peter Kreutzer,
Danile Levine, Bruce Macleod, Ken Matinale, Michael Mavrogiannis,
Cliff Otto, Alberto Perdomo, Dave Quinn, John Rickert, Tom Ruane,
Theron Skyles, Hans Van Slootenm, Michael Westbay, and Rob Wood.
Many other people have made significant contributions to the database
over the years. The contribution of Tom Ruane's effort to the overall
quality of the underlying data has been tremendous. His work at
retrosheet.org integrates the yearly data with the day-by-day data,
creating a reference source of startling depth. It is unlikely than
any individual has contributed as much to the field of baseball
research in the past five years as Ruane has.
Sean Holtz helped with a major overhaul and redesign before the
2000 season. Keith Woolner was instrumental in helping turn
a huge collection of stats into a relational database in the mid-1990s.
Clifford Otto & Ted Nye also helped provide guidance to the early
versions. Lee Sinnis, John Northey & Erik Greenwood helped supply key
pieces of data. Many others have written in with corrections and
suggestions that made each subsequent version even better than what
preceded it.
The work of the SABR Baseball Records Committee, led by Lyle Spatz
has been invaluable. So has the work of Bill Carle and the SABR
Biographical Committee. David Vincent, keeper of the Home Run Log and
other bits of hard to find info, has always been helpful. The recent
addition of colleges to player bios is the result of much research by
members of SABR's Collegiate Baseball committee.
Salary data has been supplied by Doug Pappas, who passed away during
the summer of 2004. He was the leading authority on many subjects,
most significantly the financial history of Major League Baseball.
We are grateful that he allowed us to include some of the data he
compiled. His work has been continued by the SABR Business of
Baseball committee.
Thanks is also due to the staff at the National Baseball Library
in Cooperstown who have been so helpful -- Tim Wiles, Jim Gates,
Bruce Markusen, and the rest of the staff.
A special debt of gratitude is owed to Dave Smith and the folks at
Retrosheet. There is no other group working so hard to compile and
share baseball data. Their website (www.retrosheet.org) will give
you a taste of the wealth of information Dave and the gang have collected.
The 2012 database beneifited from the work of Ted Turocy and his
Chadwick baseball Bureau. For more details on his tools and services,
visit: http://chadwick.sourceforge.net/doc/index.html
Thanks to all contributors great and small. What you have created is
a wonderful thing.
### 1.4 Using this Database
This version of the database is available in Microsoft Access
format or in a generic, comma delimited format. Because this is a
relational database, you will not be able to use the data in a
flat-database application.
Please note that this is not a stand alone application. It requires
a database application or some other application designed specifically
to interact with the database.
If you are unable to import the data directly, you should download the
database in the delimted text format. Then use the documentation
in sections 2.1 through 2.22 of this document to import the data into
your database application.
### 1.5 Revision History
Version Date Comments
1.0 December 1992 Database ported from dBase
1.1 May 1993 Becomes fully relational
1.2 July 1993 Corrections made to full database
1.21 December 1993 1993 statistics added
1.3 July 1994 Pre-1900 data added
1.31 February 1995 1994 Statistics added
1.32 August 1995 Statistics added for other leagues
1.4 September 1995 Fielding Data added
1.41 November 1995 1995 statistics added
1.42 March 1996 HOF/All-Star tables added
1.5-MS October 1996 1st public release - MS Access format
1.5-GV October 1996 Released generic comma-delimted files
1.6-MS December 1996 Updated with 1996 stats, some corrections
1.61-MS December 1996 Corrected error in MASTER table
1.62 February 1997 Corrected 1914-1915 batters data and updated
2.0 February 1998 Major Revisions-added teams & managers
2.1 October 1998 Interim release w/1998 stats
2.2 January 1999 New release w/post-season stats & awards added
3.0 November 1999 Major release - fixed errors and 1999 statistics added
4.0 May 2001 Major release - proofed & redesigned tables
4.5 March 2002 Updated with 2001 stats and added new biographical data
5.0 December 2002 Major revision - new tables and data
5.1 January 2004 Updated with 2003 data, and new pitching categories
5.2 November 2004 Updated with 2004 season statistics.
5.3 December 2005 Updated with 2005 season statistics.
5.4 December 2006 Updated with 2006 season statistics.
5.5 December 2007 Updated with 2007 season statistics.
5.6 December 2008 Updated with 2008 season statistics.
5.7 December 2009 Updated for 2009 and added several tables.
5.8 December 2010 Updated with 2010 season statistics.
5.9 December 2011 Updated for 2011 and removed obsolete tables.
2012 December 2012 Updated with 2012 season statistics
------------------------------------------------------------------------------
2.0 Data Tables
The design follows these general principles. Each player is assigned a
unique number (playerID). All of the information relating to that player
is tagged with his playerID. The playerIDs are linked to names and
birthdates in the MASTER table.
The database is comprised of the following main tables:
MASTER - Player names, DOB, and biographical info
Batting - batting statistics
Pitching - pitching statistics
Fielding - fielding statistics
It is supplemented by these tables:
AllStarFull - All-Star appearances
Hall of Fame - Hall of Fame voting data
Managers - managerial statistics
Teams - yearly stats and standings
BattingPost - post-season batting statistics
PitchingPost - post-season pitching statistics
TeamFranchises - franchise information
FieldingOF - outfield position data
FieldingPost- post-season fieldinf data
ManagersHalf - split season data for managers
TeamsHalf - split season data for teams
Salaries - player salary data
SeriesPost - post-season series information
AwardsManagers - awards won by managers
AwardsPlayers - awards won by players
AwardsShareManagers - award voting for manager awards
AwardsSharePlayers - award voting for player awards
Appearances
Schools
SchoolsPlayers
Sections 2.1 through 2.27 of this document describe each of the tables in
detail and the fields that each contains.
### 2.1 MASTER table
lahmanID Unique number assigned to each player
playerID A unique code asssigned to each player. The playerID links
the data in this file with records in the other files.
managerID An ID for individuals who served as managers
hofID An ID for individuals who are in teh baseball Hall of Fame
birthYear Year player was born
birthMonth Month player was born
birthDay Day player was born
birthCountry Country where player was born
birthState State where player was born
birthCity City where player was born
deathYear Year player died
deathMonth Month player died
deathDay Day player died
deathCountry Country where player died
deathState State where player died
deathCity City where player died
nameFirst Player's first name
nameLast Player's last name
nameNote Note about player's name (usually signifying that they changed
their name or played under two differnt names)
nameGiven Player's given name (typically first and middle)
nameNick Player's nickname
weight Player's weight in pounds
height Player's height in inches
bats Player's batting hand (left, right, or both)
throws Player's throwing hand (left or right)
debut Date that player made first major league appearance
finalGame Date that player made first major league appearance (blank if still active)
college College attended
lahman40ID ID used in Lahman Database version 4.0
lahman45ID ID used in Lahman database version 4.5
retroID ID used by retrosheet
holtzID ID used by Sean Holtz's Baseball Almanac
bbrefID ID used by Baseball Reference website
### 2.2 Batting Table
playerID Player ID code
yearID Year
stint player's stint (order of appearances within a season)
teamID Team
lgID League
G Games
G_batting Game as batter
AB At Bats
R Runs
H Hits
2B Doubles
3B Triples
HR Homeruns
RBI Runs Batted In
SB Stolen Bases
CS Caught Stealing
BB Base on Balls
SO Strikeouts
IBB Intentional walks
HBP Hit by pitch
SH Sacrifice hits
SF Sacrifice flies
GIDP Grounded into double plays
G_Old Old version of games (deprecated)
### 2.3 Pitching table
playerID Player ID code
yearID Year
stint player's stint (order of appearances within a season)
teamID Team
lgID League
W Wins
L Losses
G Games
GS Games Started
CG Complete Games
SHO Shutouts
SV Saves
IPOuts Outs Pitched (innings pitched x 3)
H Hits
ER Earned Runs
HR Homeruns
BB Walks
SO Strikeouts
BAOpp Opponent's Batting Average
ERA Earned Run Average
IBB Intentional Walks
WP Wild Pitches
HBP Batters Hit By Pitch
BK Balks
BFP Batters faced by Pitcher
GF Games Finished
R Runs Allowed
SH Sacrifices by opposing batters
SF Sacrifice flies by opposing batters
GIDP Grounded into double plays by opposing batter
### 2.4 Fielding Table
playerID Player ID code
yearID Year
stint player's stint (order of appearances within a season)
teamID Team
lgID League
Pos Position
G Games
GS Games Started
InnOuts Time played in the field expressed as outs
PO Putouts
A Assists
E Errors
DP Double Plays
PB Passed Balls (by catchers)
WP Wild Pitches (by catchers)
SB Opponent Stolen Bases (by catchers)
CS Opponents Caught Stealing (by catchers)
ZR Zone Rating
### 2.5 AllstarFull table
playerID Player ID code
YearID Year
gameNum Game number (zero if only one All-Star game played that season)
gameID Retrosheet ID for the game idea
teamID Team
lgID League
GP 1 if Played in the game
startingPos If player was game starter, the position played
### 2.6 HallOfFame table
hofID Player ID code
yearID Year of ballot
votedBy Method by which player was voted upon
ballots Total ballots cast in that year
needed Number of votes needed for selection in that year
votes Total votes received
inducted Whether player was inducted by that vote or not (Y or N)
category Category in which candidate was honored
needed_note Explanation of qualifiers for special elections
### 2.7 Managers table
managerID Player ID Number
yearID Year
teamID Team
lgID League
inseason Managerial order. Zero if the individual managed the team
the entire year. Otherwise denotes where the manager appeared
in the managerial order (1 for first manager, 2 for second, etc.)
G Games managed
W Wins
L Losses
rank Team's final position in standings that year
plyrMgr Player Manager (denoted by 'Y')
### 2.8 Teams table
yearID Year
lgID League
teamID Team
franchID Franchise (links to TeamsFranchise table)
divID Team's division
Rank Position in final standings
G Games played
GHome Games played at home
W Wins
L Losses
DivWin Division Winner (Y or N)
WCWin Wild Card Winner (Y or N)
LgWin League Champion(Y or N)
WSWin World Series Winner (Y or N)
R Runs scored
AB At bats
H Hits by batters
2B Doubles
3B Triples
HR Homeruns by batters
BB Walks by batters
SO Strikeouts by batters
SB Stolen bases
CS Caught stealing
HBP Batters hit by pitch
SF Sacrifice flies
RA Opponents runs scored
ER Earned runs allowed
ERA Earned run average
CG Complete games
SHO Shutouts
SV Saves
IPOuts Outs Pitched (innings pitched x 3)
HA Hits allowed
HRA Homeruns allowed
BBA Walks allowed
SOA Strikeouts by pitchers
E Errors
DP Double Plays
FP Fielding percentage
name Team's full name
park Name of team's home ballpark
attendance Home attendance total
BPF Three-year park factor for batters
PPF Three-year park factor for pitchers
teamIDBR Team ID used by Baseball Reference website
teamIDlahman45 Team ID used in Lahman database version 4.5
teamIDretro Team ID used by Retrosheet
### 2.9 BattingPost table
yearID Year
round Level of playoffs
playerID Player ID code
teamID Team
lgID League
G Games
AB At Bats
R Runs
H Hits
2B Doubles
3B Triples
HR Homeruns
RBI Runs Batted In
SB Stolen Bases
CS Caught stealing
BB Base on Balls
SO Strikeouts
IBB Intentional walks
HBP Hit by pitch
SH Sacrifices
SF Sacrifice flies
GIDP Grounded into double plays
### 2.10 PitchingPost table
playerID Player ID code
yearID Year
round Level of playoffs
teamID Team
lgID League
W Wins
L Losses
G Games
GS Games Started
CG Complete Games
SHO Shutouts
SV Saves
IPOuts Outs Pitched (innings pitched x 3)
H Hits
ER Earned Runs
HR Homeruns
BB Walks
SO Strikeouts
BAOpp Opponents' batting average
ERA Earned Run Average
IBB Intentional Walks
WP Wild Pitches
HBP Batters Hit By Pitch
BK Balks
BFP Batters faced by Pitcher
GF Games Finished
R Runs Allowed
SH Sacrifice Hits allowed
SF Sacrifice Flies allowed
GIDP Grounded into Double Plays
### 2.11 TeamFranchises table
franchID Franchise ID
franchName Franchise name
active Whetehr team is currently active (Y or N)
NAassoc ID of National Association team franchise played as
### 2.12 FieldingOF table
playerID Player ID code
yearID Year
stint player's stint (order of appearances within a season)
Glf Games played in left field
Gcf Games played in center field
Grf Games played in right field
### 2.13 ManagersHalf table
managerID Manager ID code
yearID Year
teamID Team
lgID League
inseason Managerial order. One if the individual managed the team
the entire year. Otherwise denotes where the manager appeared
in the managerial order (1 for first manager, 2 for second, etc.)
half First or second half of season
G Games managed
W Wins
L Losses
rank Team's position in standings for the half
### 2.14 TeamsHalf table
yearID Year
lgID League
teamID Team
half First or second half of season
divID Division
DivWin Won Division (Y or N)
rank Team's position in standings for the half
G Games played
W Wins
L Losses
### 2.15 Salaries table
yearID Year
teamID Team
lgID League
playerID Player ID code
salary Salary
### 2.16 SeriesPost table
yearID Year
round Level of playoffs
teamIDwinner Team ID of the team that won the series
lgIDwinner League ID of the team that won the series
teamIDloser Team ID of the team that lost the series
lgIDloser League ID of the team that lost the series
wins Wins by team that won the series
losses Losses by team that won the series
ties Tie games
### 2.17 AwardsManagers table
managerID Manager ID code
awardID Name of award won
yearID Year
lgID League
tie Award was a tie (Y or N)
notes Notes about the award
### 2.18 AwardsPlayers table
playerID Player ID code
awardID Name of award won
yearID Year
lgID League
tie Award was a tie (Y or N)
notes Notes about the award
### 2.19 AwardsShareManagers table
awardID name of award votes were received for
yearID Year
lgID League
managerID Manager ID code
pointsWon Number of points received
pointsMax Maximum numner of points possible
votesFirst Number of first place votes
### 2.20 AwardsSharePlayers table
awardID name of award votes were received for
yearID Year
lgID League
playerID Player ID code
pointsWon Number of points received
pointsMax Maximum numner of points possible
votesFirst Number of first place votes
### 2.21 FieldingPost table
playerID Player ID code
yearID Year
teamID Team
lgID League
round Level of playoffs
Pos Position
G Games
GS Games Started
InnOuts Time played in the field expressed as outs
PO Putouts
A Assists
E Errors
DP Double Plays
TP Triple Plays
PB Passed Balls
SB Stolen Bases allowed (by catcher)
CS Caught Stealing (by catcher)
### 2.22 Appearances table
yearID Year
teamID Team
lgID League
playerID Player ID code
G_all Total games played
GS Games started
G_batting Games in which player batted
G_defense Games in which player appeared on defense
G_p Games as pitcher
G_c Games as catcher
G_1b Games as firstbaseman
G_2b Games as secondbaseman
G_3b Games as thirdbaseman
G_ss Games as shortstop
G_lf Games as leftfielder
G_cf Games as centerfielder
G_rf Games as right fielder
G_of Games as outfielder
G_dh Games as designated hitter
G_ph Games as pinch hitter
G_pr Games as pinch runner
### 2.23 Schools table
schoolID school ID code
schoolName school name
schoolCity city where school is located
schoolState state where school's city is located
schoolNick nickname for school's baseball team
### 2.24 SchoolsPlayers
playerid Player ID code
schoolID school ID code
yearMin year player's college career started
yearMax year player's college career started
### SQL Operators
#### SELECT
The SELECT statement is used to retrieve data from the database.
• The basic syntax is:
```sql
SELECT
<columns>
FROM
<table>
```
```sql
SELECT [DISTINCT | ALL]
{* | [columnExpression [AS newName]] [,...]
}
FROM TableName [alias] [, ...]
[WHERE condition]
[GROUP BY columnList] [HAVING condition]
[ORDER BY columnList]
```
`SELECT` is followed by the names of the columns in the output.
`SELECT` is always paired with `FROM`, which identifies the table from which we're retrieving the data.
```sql
SELECT
<columns>
FROM
<table>
```
`SELECT *` returns *all* of the columns.
Yelp example:
```sql
SELECT
*
FROM yelp_reviews;
```
`SELECT <columns>` returns *some* of the columns.
Yelp example:
```sql
SELECT
review_id, text, stars
FROM yelp_reviews;
```
#### WHERE
`WHERE`, which follows the `FROM` clause, is used to filter tables using specific criteria.
```sql
SELECT
<columns>
FROM
<table>
WHERE
<condition>
```
Yelp example:
```sql
SELECT
review_id, text, stars
FROM yelp_reviews
WHERE stars > 2 and useful != 0;
```
LIMIT
Limit caps the number of rows returned.
```sql
SELECT
review_id, text, stars
FROM yelp_reviews
WHERE stars > 2
LIMIT 10;
```
Calculations
```sql
SELECT
review_id, text, stars, stars+funny+useful+cool
FROM yelp_reviews
WHERE stars > 2;
LIMIT 10;
```
#### Order of Operations
The order of operation is the same as in algebra.
1. Whatever is in parentheses is executed first. If parentheses are nested, the innermost is executed first, then the next most inner, etc.
2. Then all division and multiplication left to right.
3. And finally all addition and subtraction left to right.
### Aggregations
Aggregations (or aggregate functions) are functions in which the values of multiple rows are grouped together as an input on certain criteria to form a single value of more significant meaning or measurement. Examples are sets, bags, or lists.
Aggregate funtions include:
- Average (i.e., arithmetic mean)
- Count
- Maximum
- Minimum
- Median
- Mode
- Sum
In SQL, they are performed in a `SELECT` statement like the following:
```sql
SELECT COUNT(useful)
FROM yelp_reviews;
```
```sql
SELECT
AVG(stars), MAX(funny), MIN(cool)
FROM yelp_reviews;
```
```sql
SELECT
AVG(stars), MAX(funny), MIN(cool)
FROM yelp_reviews;
WHERE stars > 2
```
#### Aliasing
• Sometimes it is useful to alias a column name to make a more
readable result set.
```sql
SELECT AVG(stars) AS Avg_Rating
FROM yelp_reviews
```
• The AS keyword is optional.
• Double quotes “ “ can be used instead of square brackets.
#### Like
• The LIKE keyword used in a WHERE operator with a wildcard (% or _)
allows you to search for patterns in character-based fields.
```sql
SELECT
review_id, text, stars, stars+funny+useful+cool
FROM yelp_reviews
WHERE text LIKE '%Boston%';
```
#### Between
• The BETWEEN keyword can be used in criteria to return values
between to other values.
• BETWEEN is inclusive of its ends.
```sql
SELECT
review_id, text, stars, stars+funny+useful+cool
FROM yelp_reviews
WHERE date BETWEEN ’11/1/2016’ AND
‘10/03/2017’;
```
#### NULL
• Nulls are special cases. They are not a value and so cannot be
compared to a value using = or < or >.
• To locate nulls you can use the IS keyword in a criteria:
```sql
SELECT
review_id, text, stars, stars+funny+useful+cool
FROM yelp_reviews
WHERE text IS NULL
```
```sql
SELECT
review_id, text, stars, stars+funny+useful+cool
FROM yelp_reviews
WHERE text IS NOT NULL
```
#### Subqueries
• Some SQL statements can have a SELECT embedded
within them.
• A subselect can be used in WHERE and HAVING
clauses of an outer SELECT, where it is called a
subquery or nested query.
• Subselects may also appear in INSERT, UPDATE, and
DELETE statements.
## Baseball SQL
Databases for sabermetricians [http://www.hardballtimes.com/main/article/databases-for-sabermetricians-part-one/](http://www.hardballtimes.com/main/article/databases-for-sabermetricians-part-one/)
SQL Views for Sabermetrics using Lahman's Baseball Database [https://gist.github.com/wesrice/f1cc48fa8d18d17bc2de](https://gist.github.com/wesrice/f1cc48fa8d18d17bc2de)
```sql
SELECT playerID,HR FROM `Batting`;
```
```sql
SELECT playerID,HR FROM `Batting`
ORDER BY HR DESC;
```
```sql
SELECT playerID,HR FROM `Batting`
ORDER BY HR DESC
LIMIT 10;
SELECT playerID,HR, yearID FROM `Batting`
WHERE yearID = 1999
ORDER BY HR DESC
LIMIT 10;
SELECT *
FROM Batting
WHERE yearID = "1990"
AND teamID = "KCA";
SELECT *, H/AB AS AVG
, (H+BB+HBP)/(AB+BB+HBP+SF) AS OBP
, (H+2B+2*3B+3*HR)/AB AS SLG
FROM Batting
WHERE yearID = "1990"AND teamID = "KCA"AND AB IS NOT NULL;
```
A simple one to show all of the players named "Sean:"
```sql
SELECT nameLast, nameFirst, debut
FROM Master
WHERE (nameFirst="Sean")
ORDER BY nameLast;
```
Here's one to show a list of players with 50 HRs in a season:
```sql
SELECT Master.nameLast, Master.nameFirst, Batting.HR, Batting.yearID
FROM Batting INNER JOIN Master ON Batting.playerID = Master.playerID
WHERE (((Batting.HR)>=50))
ORDER BY Batting.HR DESC;
```
Here's one to show the all-time leaders in strikeouts:
```sql
SELECT Master.nameLast, Master.nameFirst, Sum(Pitching.SO) AS SumOfSO
FROM Pitching INNER JOIN Master ON Pitching.playerID = Master.playerID
GROUP BY Pitching.playerID, Master.nameLast, Master.nameFirst
ORDER BY Master.nameLast;
```
```sql
SELECT playerID, AB, H, 2B, 3B, HR, BB
, SO, IBB, HBP, SH, SF
, H/AB AS AVG
, (H+BB+HBP)/(AB+BB+HBP+SF) AS OBP
, (H+2B+2*3B+3*HR)/AB AS SLG
FROM Batting
WHERE yearID = "1990"
AND teamID = "KCA"
AND AB IS NOT NULL;
SELECT playerID, yearID, teamID, HR
FROM Batting
ORDER BY HR DESC
LIMIT 50;
SELECT playerID, SUM(HR) AS HR
FROM Batting
GROUP BY playerID
ORDER BY HR DESC
LIMIT 50;
SELECT playerID
, SUM(H)/SUM(AB) AS AVG
, SUM(AB+BB+HBP+COALESCE(SF,0)) AS PA
FROM Batting
GROUP BY playerID
HAVING PA >= 3000
ORDER BY AVG DESC
LIMIT 50;
SELECT playerID
, SUM(ER)/SUM(IPOuts/3)*9 AS ERA
, SUM(IPOuts/3) AS IP
FROM Pitching
GROUP BY playerID
HAVING IP >= 1000
ORDER BY ERA ASC
LIMIT 50;
SELECT playerID
, yearID
, teamID
, R-RBI AS R_RBI
FROM Batting
ORDER BY R_RBI DESC
LIMIT 50;
SELECT b.playerID
, b.yearID
, b.teamID
, CAST(R-RBI AS SIGNED) AS R_RBI
FROM Batting b
ORDER BY R_RBI DESC
LIMIT 50;
CREATE VIEW view_name AS
CREATE TABLE table_name AS
```
Simply put either one of those at the front of your query.
So what’s the difference? A view:
1. Updates whenever the underlying data does.
2. Preserves the query used to generate it.
Creating a table:
1. Does not update to incorporate changes in data.
2. Does not run the query every time it’s called.
A view is more flexible; a table is quicker.
```sql
CREATE OR REPLACE VIEW sabermetrics_batting_simple AS
SELECT * FROM Batting;
CREATE OR REPLACE VIEW sabermetrics_batting AS
SELECT
Batting.*,
-- PA - Plate Appearances
(Batting.AB + Batting.BB + Batting.HBP + Batting.SF + Batting.SH) as PA,
-- BB% - Walk Percentage (http://www.fangraphs.com/library/offense/rate-stats/)
round((Batting.BB / (Batting.AB + Batting.BB + Batting.HBP + Batting.SF + Batting.SH)), 3) as BBpct,
-- K% - Strikeout Percentage (http://www.fangraphs.com/library/offense/rate-stats/)
round((Batting.SO / (Batting.AB + Batting.BB + Batting.HBP + Batting.SF + Batting.SH)), 3) as Kpct,
-- ISO - Isolated Power (http://www.fangraphs.com/library/offense/iso/)
round((((Batting.2B) + (2 * Batting.3B) + ( 3 * Batting.HR)) / Batting.AB), 3) as ISO,
-- BABIP - Batting Average on Balls in Play (http://www.fangraphs.com/library/offense/babip/)
round(((Batting.H - Batting.HR) / ((Batting.AB + Batting.BB + Batting.HBP + Batting.SF + Batting.SH) - Batting.SO - Batting.BB - Batting.HR)), 3) as BABIP,
-- AVG - Batting Average
round((Batting.H / Batting.AB), 3) as AVG,
-- OBP - On Base Percentage - (http://www.fangraphs.com/library/offense/obp/)
round(((Batting.H + Batting.BB + Batting.HBP) / (Batting.AB + Batting.BB + Batting.HBP + Batting.SF)), 3) as OBP,
-- SLG - Slugging Percentage
round(((Batting.H + Batting.2B + 2 * Batting.3B + 3 * Batting.HR) / Batting.AB), 3) as SLG,
-- OPS - On Base + Slugging (http://www.fangraphs.com/library/offense/ops/)
round(((Batting.H + Batting.BB + Batting.HBP) / (Batting.AB + Batting.BB + Batting.HBP + Batting.SF)) + (((Batting.H - Batting.2B - Batting.3B - Batting.HR) + (2 * Batting.2B) + (3 * Batting.3B) + (4 * Batting.HR)) / Batting.AB), 3) as OPS
FROM Batting
ORDER BY Batting.playerID ASC, Batting.yearID ASC;
CREATE OR REPLACE VIEW sabermetrics_battingpost AS
SELECT
Batting.*,
-- PA - Plate Appearances
(Batting.AB + Batting.BB + Batting.HBP + Batting.SF + Batting.SH) as PA,
-- BB% - Walk Percentage (http://www.fangraphs.com/library/offense/rate-stats/)
round((Batting.BB / (Batting.AB + Batting.BB + Batting.HBP + Batting.SF + Batting.SH)), 3) as BBpct,
-- K% - Strikeout Percentage (http://www.fangraphs.com/library/offense/rate-stats/)
round((Batting.SO / (Batting.AB + Batting.BB + Batting.HBP + Batting.SF + Batting.SH)), 3) as Kpct,
-- ISO - Isolated Power (http://www.fangraphs.com/library/offense/iso/)
round((((Batting.2B) + (2 * Batting.3B) + ( 3 * Batting.HR)) / Batting.AB), 3) as ISO,
-- BABIP - Batting Average on Balls in Play (http://www.fangraphs.com/library/offense/babip/)
round(((Batting.H - Batting.HR) / ((Batting.AB + Batting.BB + Batting.HBP + Batting.SF + Batting.SH) - Batting.SO - Batting.BB - Batting.HR)), 3) as BABIP,
-- AVG - Batting Average
round((Batting.H / Batting.AB), 3) as AVG,
-- OBP - On Base Percentage - (http://www.fangraphs.com/library/offense/obp/)
round(((Batting.H + Batting.BB + Batting.HBP) / (Batting.AB + Batting.BB + Batting.HBP + Batting.SF)), 3) as OBP,
-- SLG - Slugging Percentage
round(((Batting.H + Batting.2B + 2 * Batting.3B + 3 * Batting.HR) / Batting.AB), 3) as SLG,
-- OPS - On Base + Slugging (http://www.fangraphs.com/library/offense/ops/)
round(((Batting.H + Batting.BB + Batting.HBP) / (Batting.AB + Batting.BB + Batting.HBP + Batting.SF)) + (((Batting.H - Batting.2B - Batting.3B - Batting.HR) + (2 * Batting.2B) + (3 * Batting.3B) + (4 * Batting.HR)) / Batting.AB), 3) as OPS
FROM BattingPost AS Batting
ORDER BY Batting.playerID ASC, Batting.yearID ASC;
CREATE OR REPLACE VIEW sabermetrics_Fielding AS
SELECT
Fielding.*
-- PCT - Fielding Percentage
-- round(avg((Fielding.PO + Fielding.A) / (Fielding.PO + Fielding.A + Fielding.E)), 3) as PCT
FROM Fielding
ORDER BY Fielding.playerID ASC, Fielding.yearID ASC
CREATE OR REPLACE VIEW sabermetrics_Fieldingpost AS
SELECT
Fielding.*
-- PCT - Fielding Percentage
-- round(avg((Fielding.PO + Fielding.A) / (Fielding.PO + Fielding.A + Fielding.E)), 3) as PCT
FROM FieldingPost AS Fielding
ORDER BY Fielding.playerID ASC, Fielding.yearID ASC;
CREATE OR REPLACE VIEW sabermetrics_Pitching AS
SELECT
Pitching.*,
-- IP - Innings Pitched
round((Pitching.IPouts / 3), 3) as IP,
-- K/9 - Strikeouts per 9 innings (http://www.fangraphs.com/library/Pitching/rate-stats/)
round((Pitching.SO * 9) / (Pitching.IPouts / 3), 3) as k_9,
-- BB/9 - Walks per 9 innings (http://www.fangraphs.com/library/Pitching/rate-stats/)
round((Pitching.BB * 9) / (Pitching.IPouts / 3), 3) as BB_9,
-- K/BB - Strikeout to Walk Ratio
round((Pitching.SO / Pitching.BB), 3) as K_BB,
-- K% - Strikeout Percentage (http://www.fangraphs.com/library/Pitching/rate-stats/)
round((Pitching.SO / Pitching.BFP), 3) as Kpct,
-- BB% - Walk Percentage (http://www.fangraphs.com/library/Pitching/rate-stats/)
round((Pitching.BB / Pitching.BFP), 3) as BBpct,
-- HR/9 - Home Runs per 9 innings
round((Pitching.HR * 9) / (Pitching.IPouts / 3), 3) as HR_9,
-- AVG - Batting Average Against
round((Pitching.H / (Pitching.IPouts - Pitching.BB - Pitching.HBP - Pitching.SH - Pitching.SF)), 3) as AVG,
-- WHIP - Walks + Hits per Inning Pitch (http://www.fangraphs.com/library/Pitching/whip/)
round(((Pitching.BB + Pitching.H) / (Pitching.IPouts / 3)), 3) as WHIP,
-- BABIP - Batting Average on Balls in Play (http://www.fangraphs.com/library/Pitching/babip/)
round(((Pitching.H - Pitching.HR) / (Pitching.BFP - Pitching.SO - Pitching.BB - Pitching.HR)), 3) as BABIP
FROM Pitching
ORDER BY Pitching.playerID ASC, Pitching.yearID ASC;
CREATE OR REPLACE VIEW sabermetrics_Pitchingpost AS
SELECT
Pitching.*,
-- IP - Innings Pitched
round((Pitching.IPouts / 3), 3) as IP,
-- K/9 - Strikeouts per 9 innings (http://www.fangraphs.com/library/Pitching/rate-stats/)
round((Pitching.SO * 9) / (Pitching.IPouts / 3), 3) as k_9,
-- BB/9 - Walks per 9 innings (http://www.fangraphs.com/library/Pitching/rate-stats/)
round((Pitching.BB * 9) / (Pitching.IPouts / 3), 3) as BB_9,
-- K/BB - Strikeout to Walk Ratio
round((Pitching.SO / Pitching.BB), 3) as K_BB,
-- K% - Strikeout Percentage (http://www.fangraphs.com/library/Pitching/rate-stats/)
round((Pitching.SO / Pitching.BFP), 3) as Kpct,
-- BB% - Walk Percentage (http://www.fangraphs.com/library/Pitching/rate-stats/)
round((Pitching.BB / Pitching.BFP), 3) as BBpct,
-- HR/9 - Home Runs per 9 innings
round((Pitching.HR * 9) / (Pitching.IPouts / 3), 3) as HR_9,
-- AVG - Batting Average Against
round((Pitching.H / (Pitching.IPouts - Pitching.BB - Pitching.HBP - Pitching.SH - Pitching.SF)), 3) as AVG,
-- WHIP - Walks + Hits per Inning Pitch (http://www.fangraphs.com/library/Pitching/whip/)
round(((Pitching.BB + Pitching.H) / (Pitching.IPouts / 3)), 3) as WHIP,
-- BABIP - Batting Average on Balls in Play (http://www.fangraphs.com/library/Pitching/babip/)
round(((Pitching.H - Pitching.HR) / (Pitching.BFP - Pitching.SO - Pitching.BB - Pitching.HR)), 3) as BABIP
FROM PitchingPost AS Pitching
ORDER BY Pitching.playerID ASC, Pitching.yearID ASC;
```
*The below queries take a while *
```sql
SELECT p.playerID
, SUM(p.ER)/SUM(p.IPOuts/3)*9 AS ERA
, SUM(p.IPOuts/3) AS IP
FROM Pitching p, Master m
WHERE p.playerID = m.playerID
GROUP BY playerID
HAVING IP >= 1000
ORDER BY ERA ASC
LIMIT 50;
SELECT p.playerID, ERA, IP
FROM (SELECT playerID
, SUM(ER)/SUM(IPOuts/3)*9 AS ERA
, SUM(IPOuts/3) AS IP
FROM Pitching
GROUP BY playerID) p
HAVING IP >= 1000;
SELECT p.playerID, Name, ERA, IP
FROM (SELECT playerID
, SUM(ER)/SUM(IPOuts/3)*9 AS ERA
, SUM(IPOuts/3) AS IP
FROM Pitching
GROUP BY playerID) p
HAVING IP >= 1000
JOIN (SELECT CONCAT(nameFirst," ",nameLast) AS Name
, playerID
FROM Master) m
ON p.playerID = m.playerID
ORDER BY ERA ASC
LIMIT 50;
```
## Baseball Twitter Use Cases.
Use Case 1
Description: See which Players have the most followers
Actor: User
Precondition: Players must have Twitter accounts
Steps: Find all players with Twitter accounts and then find the most followed(Top 15)
Actor action: Request to see Players with Twitter accounts
System Responses: Return list of 15 players with full_names and Twitter handles
Post Condition: User will be given name and handle of most followed players
Alternate Path:
Error: User input is incorrect
```sql
SELECT a.full_name, a.user_id, a.followers
FROM (Account a LEFT JOIN Player p ON p.player_user_id = a.user_id)
ORDER BY a.followers DESC
LIMIT 15;
```
```sql
+-----------------------------------+-----------------+-----------+
| full_name | user_id | followers |
+-----------------------------------+-----------------+-----------+
| Nick Swisher | NickSwisher | 1736163 |
| ダルビッシュ有(Yu Darvish) | faridyu | 1252441 |
| Brandon Phillips | DatDudeBP | 1005157 |
| 田中将大/MASAHIRO TANAKA | t_masahiro18 | 844487 |
| David Ortiz | davidortiz | 804894 |
| Luis montes Jiménez | Chapomontes10 | 662539 |
| Jose Bautista | JoeyBats19 | 648481 |
| Mike Trout | Trouty20 | 629613 |
| Brian Wilson | BrianWilson38 | 605029 |
| Justin Verlander | JustinVerlander | 585342 |
| Miguel Cabrera | MiguelCabrera | 565891 |
| Robinson Cano | RobinsonCano | 490128 |
| Bryce Harper | Bharper3407 | 460107 |
| Matt Kemp | TheRealMattKemp | 434733 |
| CC Sabathia | CC_Sabathia | 416583 |
+-----------------------------------+-----------------+-----------+
```
Use Case 2
Description: Get a Players Twitter handle with their 2014 hits
Actor: User
Precondition: Player must have a Twitter account to be included
Steps: Find all players with Twitter accounts and then find each players hits
Actor action: Request to see Players with Twitter accounts
System Responses: Return list of all players on Twitter with their 2014 hits
Post Condition: User will be given name and handle as well as hits of players
Alternate Path:
Error: User input is incorrect
```sql
SELECT p.full_name, p.player_user_id, b.H
FROM ((Player p INNER JOIN Batting b ON b.full_name=p.full_name AND b.dob=p.dob)
LEFT JOIN Account a ON a.user_id=p.player_user_id)
WHERE b.h > 0 AND NOT p.player_user_id="NULL"
ORDER BY b.H DESC;
```
```sql
+-----------------------+------------------+-----+
| full_name | player_user_id | H |
+-----------------------+------------------+-----+
| Jose Altuve | @JoseAltuve27 | 225 |
| Miguel Cabrera | @MiguelCabrera | 191 |
| Ian Kinsler | @IKinsler3 | 188 |
| Robinson Cano | @RobinsonCano | 187 |
| Ben Revere | @BenRevere9 | 184 |
| Denard Span | @thisisdspan | 184 |
| Adam Jones | @SimplyAJ10 | 181 |
| Howie Kendrick | @HKendrick47 | 181 |
| Hunter Pence | @hunterpence | 180 |
| Jose Abreu | @79JoseAbreu | 176 |
| Dee Gordon | @FlashGJr | 176 |
| Jonathan Lucroy | @JLucroy20 | 176 |
| Freddie Freeman | @FreddieFreeman5 | 175 |
| Jose Reyes | @lamelaza_7 | 175 |
| James Loney | @theloney_s | 174 |
| Mike Trout | @Trouty20 | 173 |
| Andrew McCutchen | @TheCUTCH22 | 172 |
| Albert Pujols | @PujolsFive | 172 |
| Charlie Blackmon | @Chuck_Nazty | 171 |
| Buster Posey | @BusterPosey | 170 |
| Alexei Ramirez | @ImTheRealAlexei | 170 |
| Nelson Cruz | @ncboomstick23 | 166 |
| Alcides Escobar | @alcidesescobar2 | 165 |
| Yasiel Puig | @YasielPuig | 165 |
| Christian Yelich | @ChristianYelich | 165 |
| Erick Aybar | @aybarer01 | 164 |
```
Use Case 3
Description: Get all players who played for Mets in 2014 ordered by games played
Actor: User
Precondition: Only Includes players on the Mets
Steps: Find all players who played for the Mets in 2014 and then order them by games played
Actor action: Request to see amount of games played by each player on the Mets in 2014
System Responses: Return a list of all players who played for the Mets in 2014 ordered
from most games played to least games played
Post Condition: User will be given a list of all Mets players ordered by Games played
Alternate Path:
Error: User input is incorrect
```sql
SELECT p.full_name, b.G
FROM Player p
INNER JOIN Batting b
ON p.full_name=b.full_name and p.dob = b.dob
WHERE p.team_abbrev="NYM"
ORDER BY b.G DESC;
```
```sql
+-------------------+-----+
| full_name | G |
+-------------------+-----+
| Curtis Granderson | 155 |
| Lucas Duda | 153 |
| Daniel Murphy | 143 |
| David Wright | 134 |
| Ruben Tejada | 119 |
| Juan Lagares | 116 |
| Travis d'Arnaud | 108 |
| Eric Young | 100 |
| Chris Young | 88 |
| Eric Campbell | 85 |
| Bobby Abreu | 78 |
| Wilmer Flores | 78 |
| Jeurys Familia | 76 |
| Carlos Torres | 73 |
| Jenrry Mejia | 63 |
| Kirk Nieuwenhuis | 61 |
| Anthony Recker | 58 |
| Matt den Dekker | 53 |
| Josh Edgin | 47 |
| Vic Black | 41 |
| Daisuke Matsuzaka | 34 |
| Zach Wheeler | 34 |
| Scott Rice | 32 |
| Bartolo Colon | 31 |
| Jonathon Niese | 31 |
| Dana Eveland | 30 |
| Buddy Carlyle | 27 |
| Gonzalez Germen | 25 |
| Josh Satin | 25 |
| Jacob deGrom | 23 |
| Chris Young | 23 |
| Dillon Gee | 22 |
| Jose Valverde | 21 |
| Andrew Brown | 19 |
| Kyle Farnsworth | 19 |
| Dilson Herrera | 18 |
| Kyle Farnsworth | 16 |
| Omar Quintanilla | 15 |
| Ike Davis | 12 |
| Juan Centeno | 10 |
| Rafael Montero | 10 |
| Taylor Teagarden | 9 |
| Erik Goeddel | 6 |
| John Lannan | 5 |
| Dario Alvarez | 4 |
| Wilfredo Tovar | 2 |
| Bobby Parnell | 1 |
+-------------------+-----+
```
Use Case 4
Description: Get top 20 starting pitchers by ERA
Actor: User
Precondition: Only includes pitchers who started more than 10 games
Steps: Find all pitchers who have started more than 10 games and order them by ERA, lowest first
take the top 20 from the result
Actor action: Request to see best starting pitchers by ERA
System Responses: Return each pitchers full name, season ERA, and their team
Post Condition: User will be given a list of the top 20 pitchers, their ERA's and the team they play for.
Alternate Path:
Error: User input is incorrect
```sql
SELECT pi.full_name, pi.ERA, pl.team_abbrev as team
FROM Pitching pi
INNER JOIN (Player pl)
ON pi.full_name=pl.full_name and pi.dob=pl.dob
WHERE GS > 10
ORDER BY ERA
LIMIT 20;
```
```sql
+-------------------+------+------+
| full_name | ERA | team |
+-------------------+------+------+
| Clayton Kershaw | 1.77 | LAD |
| Michael Pineda | 1.89 | NYY |
| Felix Hernandez | 2.14 | SEA |
| Jake Peavy | 2.17 | BOS |
| Chris Sale | 2.17 | CHW |
| Johnny Cueto | 2.25 | CIN |
| Jon Lester | 2.35 | BOS |
| Adam Wainwright | 2.38 | STL |
| Doug Fister | 2.41 | WAS |
| Corey Kluber | 2.44 | CLE |
| Cole Hamels | 2.46 | PHI |
| Kyle Hendricks | 2.46 | CHC |
| Jon Lester | 2.52 | BOS |
| Jake Arrieta | 2.53 | CHC |
| Danny Duffy | 2.53 | KC |
| Carlos Carrasco | 2.55 | CLE |
| Andrew Cashner | 2.55 | SD |
| Garrett Richards | 2.61 | LAA |
| Henderson Alvarez | 2.65 | MIA |
| Jordan Zimmermann | 2.66 | WAS |
+-------------------+------+------+
```
Use Case 5
Description: List players, their twitter handles, and their HR's/RBI's
Actor: User
Precondition: Only includes players on Twitter
Steps: Find all players with a Twitter handle and then get their RBI's and HR's
Actor action: Request to see players with their twitter handles, HR's and RBI's
System Responses: Return each players full name, twitter handle, and their HR's/RBI's
Error: User input is incorrect
```sql
SELECT p.full_name, p.player_user_id, b.HR, b.RBI
FROM Batting b
INNER JOIN Player p
ON p.full_name=b.full_name AND b.dob=p.dob
WHERE b.RBI > 0 AND NOT p.player_user_id="NULL"
ORDER BY b.RBI DESC;
```
```sql
+-----------------------+------------------+----+-----+
| full_name | player_user_id | HR | RBI |
+-----------------------+------------------+----+-----+
| Adrian Gonzalez | @AdrianTitan23 | 27 | 116 |
| Mike Trout | @Trouty20 | 36 | 111 |
| Miguel Cabrera | @MiguelCabrera | 25 | 109 |
| Nelson Cruz | @ncboomstick23 | 40 | 108 |
| Jose Abreu | @79JoseAbreu | 36 | 107 |
| Albert Pujols | @PujolsFive | 28 | 105 |
| Giancarlo Stanton | @Giancarlo818 | 37 | 105 |
| David Ortiz | @davidortiz | 35 | 104 |
| Jose Bautista | @JoeyBats19 | 35 | 103 |
| Justin Upton | @JUST_JUP | 29 | 102 |
| Josh Donaldson | @BringerOfRain20 | 29 | 98 |
| Edwin Encarnacion | @Encadwin | 34 | 98 |
| Adam Jones | @SimplyAJ10 | 29 | 96 |
| Ryan Howard | @ryanhoward | 23 | 95 |
| Ian Kinsler | @IKinsler3 | 17 | 92 |
| Adam LaRoche | @e3laroche | 26 | 92 |
| Ian Desmond | @IanDesmond20 | 24 | 91 |
| Evan Longoria | @Evan3Longoria | 22 | 91 |
| Matt Holliday | @mattholliday7 | 20 | 90 |
| Matt Kemp | @TheRealMattKemp | 25 | 89 |
| Buster Posey | @BusterPosey | 22 | 89 |
| Marlon Byrd | @mjbsr6 | 25 | 85 |
```
| github_jupyter |
# Characterization of Systems in the Time Domain
*This Jupyter notebook is part of a [collection of notebooks](../index.ipynb) in the bachelors module Signals and Systems, Communications Engineering, Universität Rostock. Please direct questions and suggestions to [Sascha.Spors@uni-rostock.de](mailto:Sascha.Spors@uni-rostock.de).*
## Impulse Response
The response $y(t)$ of a linear time-invariant (LTI) system $\mathcal{H}$ to an arbitrary input signal $x(t)$ is derived in the following. The input signal can be represented as an integral when applying the [sifting-property of the Dirac impulse](../continuous_signals/standard_signals.ipynb#Dirac-Impulse)
\begin{equation}
x(t) = \int_{-\infty}^{\infty} x(\tau) \cdot \delta(t-\tau) \; d \tau
\end{equation}
Introducing above relation for the the input signal $x(t)$ into the output signal $y(t) = \mathcal{H} \{ x(t) \}$ of the system yields
\begin{equation}
y(t) = \mathcal{H} \left\{ \int_{-\infty}^{\infty} x(\tau) \cdot \delta(t-\tau) \; d \tau \right\}
\end{equation}
where $\mathcal{H} \{ \cdot \}$ denotes the system response operator. The integration and system response operator can be exchanged under the assumption that the system is linear
\begin{equation}
y(t) = \int_{-\infty}^{\infty} x(\tau) \cdot \mathcal{H} \left\{ \delta(t-\tau) \right\} \; d \tau
\end{equation}
where $\mathcal{H} \{\cdot\}$ was only applied to the Dirac impulse, since $x(\tau)$ can be regarded as constant factor with respect to the time $t$. It becomes evident that the response of a system to a Dirac impulse plays an important role in the calculation of the output signal for arbitrary input signals.
The response of a system to a Dirac impulse as input signal is denoted as [*impulse response*](https://en.wikipedia.org/wiki/Impulse_response). It is defined as
\begin{equation}
h(t) = \mathcal{H} \left\{ \delta(t) \right\}
\end{equation}
If the system is time-invariant, the response to a shifted Dirac impulse is $\mathcal{H} \left\{ \delta(t-\tau) \right\} = h(t-\tau)$. Hence, for an LTI system we finally get
\begin{equation}
y(t) = \int_{-\infty}^{\infty} x(\tau) \cdot h(t-\tau) \; d \tau
\end{equation}
Due to its relevance in the theory of LTI systems, this operation is explicitly termed as [*convolution*](https://en.wikipedia.org/wiki/Convolution). It is commonly abbreviated by $*$, hence for above integral we get $y(t) = x(t) * h(t)$. In some books the mathematically more precise nomenclature $y(t) = (x*h)(t)$ is used, since $*$ is the operator acting on the two signals $x$ and $h$ with regard to time $t$.
In can be concluded that the properties of an LTI system are entirely characterized by its impulse response. The response $y(t)$ of a system to an arbitrary input signal $x(t)$ is given by the convolution of the input signal $x(t)$ with its impulse response $h(t)$.
**Example**
The following example considers an LTI system whose relation between input $x(t)$ and output $y(t)$ is given by an ordinary differential equation (ODE) with constant coefficients
\begin{equation}
y(t) + \frac{d}{dt} y(t) = x(t)
\end{equation}
The system response is computed for the input signal $x(t) = e^{- 2 t} \cdot \epsilon(t)$ by
1. explicitly solving the ODE and by
2. computing the impulse response $h(t)$ and convolution with the input signal.
The solution should fulfill the initial conditions $y(t)\big\vert_{t = 0-} = 0$ and $\frac{d}{dt}y(t)\big\vert_{t = 0-} = 0$ due to causality.
First the ODE is defined in `SymPy`
```
import sympy as sym
sym.init_printing()
t = sym.symbols('t', real=True)
x = sym.Function('x')(t)
y = sym.Function('y')(t)
ode = sym.Eq(y + y.diff(t), x)
ode
```
The ODE is solved for the given input signal in order to calculate the output signal. The integration constant is calculated such that the solution fulfills the initial conditions
```
solution = sym.dsolve(ode.subs(x, sym.exp(-2*t)*sym.Heaviside(t)))
integration_constants = sym.solve(
(solution.rhs.limit(t, 0, '-'), solution.rhs.diff(t).limit(t, 0, '-')), 'C1')
y1 = solution.subs(integration_constants)
y1
```
Lets plot the output signal derived by explicit solution of the ODE
```
sym.plot(y1.rhs, (t, -1, 10), ylabel=r'$y(t)$');
```
The impulse response $h(t)$ is computed by solving the ODE for a Dirac impulse as input signal, $x(t) = \delta(t)$
```
h = sym.Function('h')(t)
solution2 = sym.dsolve(ode.subs(x, sym.DiracDelta(t)).subs(y, h))
integration_constants = sym.solve((solution2.rhs.limit(
t, 0, '-'), solution2.rhs.diff(t).limit(t, 0, '-')), 'C1')
h = solution2.subs(integration_constants)
h
```
Lets plot the impulse response $h(t)$ of the LTI system
```
sym.plot(h.rhs, (t, -1, 10), ylabel=r'$h(t)$');
```
As alternative to the explicit solution of the ODE, the system response is computed by evaluating the convolution $y(t) = x(t) * h(t)$. Since `SymPy` cannot handle the Heaviside function properly in integrands, the convolution integral is first simplified. Both the input signal $x(t)$ and the impulse response $h(t)$ are causal signals. Hence, the convolution integral degenerates to
\begin{equation}
y(t) = \int_{0}^{t} x(\tau) \cdot h(t - \tau) \; d\tau
\end{equation}
for $t \geq 0$. Note that $y(t) = 0$ for $t<0$.
```
tau = sym.symbols('tau', real=True)
y2 = sym.integrate(sym.exp(-2*tau) * h.rhs.subs(t, t-tau), (tau, 0, t))
y2
```
Lets plot the output signal derived by evaluation of the convolution
```
sym.plot(y2, (t, -1, 10), ylabel=r'$y(t)$');
```
**Exercise**
* Compare the output signal derived by explicit solution of the ODE with the signal derived by convolution. Are both equal?
* Check if the impulse response $h(t)$ is a solution of the ODE by manual calculation. Hint $\frac{d}{dt} \epsilon(t) = \delta(t)$.
* Check the solution of the convolution integral by manual calculation including the Heaviside functions.
**Copyright**
This notebook is provided as [Open Educational Resource](https://en.wikipedia.org/wiki/Open_educational_resources). Feel free to use the notebook for your own purposes. The text is licensed under [Creative Commons Attribution 4.0](https://creativecommons.org/licenses/by/4.0/), the code of the IPython examples under the [MIT license](https://opensource.org/licenses/MIT). Please attribute the work as follows: *Sascha Spors, Continuous- and Discrete-Time Signals and Systems - Theory and Computational Examples*.
| github_jupyter |
# Confined Aquifer Test
**This test is taken from AQTESOLV examples.**
```
%matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
from ttim import *
```
Set basic parameters:
```
Q = 6605.754 #constant discharge in m^3/d
b = -15.24 #aquifer thickness in m
rw = 0.1524 #well radius in m
```
Load data of three observation wells:
```
data1 = np.loadtxt('data/sioux100.txt')
t1 = data1[:, 0]
h1 = data1[:, 1]
r1 = 30.48 #distance between obs1 to pumping well
data2 = np.loadtxt('data/sioux200.txt')
t2 = data2[:, 0]
h2 = data2[:, 1]
r2 = 60.96 #distance between obs2 to pumping well
data3 = np.loadtxt('data/sioux400.txt')
t3 = data3[:, 0]
h3 = data3[:, 1]
r3 = 121.92 #distance between obs3 to pumping well
```
Create conceptual model:
```
ml_0 = ModelMaq(kaq=10, z=[0, b], Saq=0.001, tmin=0.001, tmax=10, topboundary='conf')
w_0 = Well(ml_0, xw=0, yw=0, rw=rw, tsandQ=[(0, Q)], layers = 0)
ml_0.solve()
```
Calibrate with three datasets simultaneously:
```
#unknown parameters: k, Saq
ca_0 = Calibrate(ml_0)
ca_0.set_parameter(name='kaq0', initial=10)
ca_0.set_parameter(name='Saq0', initial=1e-4)
ca_0.series(name='obs1', x=r1, y=0, t=t1, h=h1, layer=0)
ca_0.series(name='obs2', x=r2, y=0, t=t2, h=h2, layer=0)
ca_0.series(name='obs3', x=r3, y=0, t=t3, h=h3, layer=0)
ca_0.fit(report=True)
display(ca_0.parameters)
print('RMSE:', ca_0.rmse())
hm1_0 = ml_0.head(r1, 0, t1)
hm2_0 = ml_0.head(r2, 0, t2)
hm3_0 = ml_0.head(r3, 0, t3)
plt.figure(figsize = (8, 5))
plt.semilogx(t1, h1, '.', label='obs1')
plt.semilogx(t1, hm1_0[0], label='ttim1')
plt.semilogx(t2, h2, '.', label='obs2')
plt.semilogx(t2, hm2_0[0], label='ttim2')
plt.semilogx(t3, h3, '.', label='obs3')
plt.semilogx(t3, hm3_0[0], label='ttim3')
plt.xlabel('time(d)')
plt.ylabel('head(m)')
plt.legend()
plt.savefig('C:/Users/DELL/Python Notebook/MT BE/Fig/siouxsfit1.eps')
plt.show();
```
Try adding res and rc:
```
ml_1 = ModelMaq(kaq=10, z=[0, b], Saq=0.001, tmin=0.001, tmax=10, topboundary='conf')
w_1 = Well(ml_1, xw=0, yw=0, rw=rw, rc=0, res=0, tsandQ=[(0, Q)], layers=0)
ml_1.solve()
```
Calibrate with three datasets simultaneously:
When adding both res and rc into calibration, the optimized res value is bout 1.2e-12, which is close to the minimum limitation. Thus, adding res has nearly no effect on improving conceptual model's performance. res is removed from the calibration.
```
#unknown parameters: k, Saq, res, rc
ca_1 = Calibrate(ml_1)
ca_1.set_parameter(name='kaq0', initial=10)
ca_1.set_parameter(name='Saq0', initial=1e-4)
#ca_1.set_parameter_by_reference(name='res', parameter=w_1.res, initial=0, pmin = 0)
ca_1.set_parameter_by_reference(name='rc', parameter=w_1.rc, initial=0)
ca_1.series(name='obs1', x=r1, y=0, t=t1, h=h1, layer=0)
ca_1.series(name='obs2', x=r2, y=0, t=t2, h=h2, layer=0)
ca_1.series(name='obs3', x=r3, y=0, t=t3, h=h3, layer=0)
ca_1.fit(report=True)
display(ca_1.parameters)
print('RMSE:', ca_1.rmse())
hm1_1 = ml_1.head(r1, 0, t1)
hm2_1 = ml_1.head(r2, 0, t2)
hm3_1 = ml_1.head(r3, 0, t3)
plt.figure(figsize = (8, 5))
plt.semilogx(t1, h1, '.', label='obs1')
plt.semilogx(t1, hm1_1[0], label='ttim1')
plt.semilogx(t2, h2, '.', label='obs2')
plt.semilogx(t2, hm2_1[0], label='ttim2')
plt.semilogx(t3, h3, '.', label='obs3')
plt.semilogx(t3, hm3_1[0], label='ttim3')
plt.xlabel('time(d)')
plt.ylabel('head(m)')
plt.legend()
plt.savefig('C:/Users/DELL/Python Notebook/MT BE/Fig/siouxfit2.eps')
plt.show();
```
## Summary of values simulated by AQTESOLV
```
t = pd.DataFrame(columns=['k [m/d]', 'Ss [1/m]', 'rc'], \
index=['AQTESOLV', 'MLU', 'ttim', 'ttim-rc'])
t.loc['AQTESOLV'] = [282.659, 4.211E-03, '-']
t.loc['ttim'] = np.append(ca_0.parameters['optimal'].values, '-')
t.loc['ttim-rc'] = ca_1.parameters['optimal'].values
t.loc['MLU'] = [282.684, 4.209e-03, '-']
t['RMSE'] = [0.003925, 0.003897, ca_0.rmse(), ca_1.rmse()]
t
```
| github_jupyter |
# Introduction
Now we have an idea of three important components to analyzing neuroimaging data:
1. Data manipulation
2. Cleaning and confound regression
3. Parcellation and signal extraction
In this notebook the goal is to integrate these 3 basic components and perform a full analysis of group data using **Intranetwork Functional Connectivity (FC)**.
Intranetwork functional connectivity is essentially a result of performing correlational analysis on mean signals extracted from two ROIs. Using this method we can examine how well certain resting state networks, such as the **Default Mode Network (DMN)**, are synchronized across spatially distinct regions.
ROI-based correlational analysis forms the basis of many more sophisticated kinds of functional imaging analysis.
## Using Nilearn's High-level functionality to compute correlation matrices
Nilearn has a built in function for extracting timeseries from functional files and doing all the extra signal processing at the same time. Let's walk through how this is done
First we'll grab our imports as usual
```
from nilearn import image as nimg
from nilearn import plotting as nplot
import numpy as np
import pandas as pd
from bids import BIDSLayout
```
Let's grab the data that we want to perform our connectivity analysis on using PyBIDS:
```
#Use PyBIDS to parse BIDS data structure
fmriprep_dir = "../data/ds000030/derivatives/fmriprep/"
layout = BIDSLayout(fmriprep_dir,
config=['bids','derivatives'])
#Get resting state data (preprocessed, mask, and confounds file)
func_files = layout.get(datatype='func', task='rest',
desc='preproc',
space='MNI152NLin2009cAsym',
extension='nii.gz',
return_type='file')
mask_files = layout.get(datatype='func', task='rest',
desc='brain',
suffix="mask",
space='MNI152NLin2009cAsym',
extension='nii.gz',
return_type='file')
confound_files = layout.get(datatype='func',
task='rest',
desc='confounds',
extension='tsv',
return_type='file')
```
Now that we have a list of subjects to peform our analysis on, let's load up our parcellation template file
```
#Load separated parcellation
parcel_file = '../resources/rois/yeo_2011/Yeo_JNeurophysiol11_MNI152/relabeled_yeo_atlas.nii.gz'
yeo_7 = nimg.load_img(parcel_file)
```
Now we'll import a package from <code>nilearn</code>, called <code>input_data</code> which allows us to pull data using the parcellation file, and at the same time applying data cleaning!
We first create an object using the parcellation file <code>yeo_7</code> and our cleaning settings which are the following:
Settings to use:
- Confounds: trans_x, trans_y, trans_z, rot_x, rot_y, rot_z, white_matter, csf, global_signal
- Temporal Derivatives: Yes
- high_pass = 0.009
- low_pass = 0.08
- detrend = True
- standardize = True
The object <code>masker</code> is now able to be used on *any functional image of the same size*. The `input_data.NiftiLabelsMasker` object is a wrapper that applies parcellation, cleaning and averaging to an functional image. For example let's apply this to our first subject:
```
# Pull the first subject's data
func_file = func_files[0]
mask_file = mask_files[0]
confound_file = confound_files[0]
```
Before we go ahead and start using the <code>masker</code> that we've created, we have to do some preparatory steps. The following should be done prior to use the <code>masker</code> object:
1. Make your confounds matrix (as we've done in Episode 06)
2. Drop Dummy TRs that are to be excluded from our cleaning, parcellation, and averaging step
To help us with the first part, let's define a function to help extract our confound regressors from the .tsv file for us. Note that we've handled pulling the appropriate `{confounds}_derivative1` columns for you! You just need to supply the base regressors!
```
#Refer to part_06 for code + explanation
def extract_confounds(confound_tsv,confounds,dt=True):
'''
Arguments:
confound_tsv Full path to confounds.tsv
confounds A list of confounder variables to extract
dt Compute temporal derivatives [default = True]
Outputs:
confound_mat
'''
if dt:
dt_names = ['{}_derivative1'.format(c) for c in confounds]
confounds = confounds + dt_names
#Load in data using Pandas then extract relevant columns
confound_df = pd.read_csv(confound_tsv,delimiter='\t')
confound_df = confound_df[confounds]
#Convert into a matrix of values (timepoints)x(variable)
confound_mat = confound_df.values
#Return confound matrix
return confound_mat
```
Finally we'll set up our image file for confound regression (as we did in Episode 6). To do this we'll drop 4 TRs from *both our functional image and our confounds file*. Note that our <code>masker</code> object will not do this for us!
```
#Load functional image
#Remove the first 4 TRs
#Use the above function to pull out a confound matrix
#Drop the first 4 rows of the confounds matrix
```
### Using the masker
Finally with everything set up, we can now use the masker to perform our:
1. Confounds cleaning
2. Parcellation
3. Averaging within a parcel
All in one step!
```
#Apply cleaning, parcellation and extraction to functional data
```
Just to be clear, this data is *automatically parcellated for you, and, in addition, is cleaned using the confounds you've specified!*
The result of running <code>masker.fit_transform</code> is a matrix that has:
- Rows matching the number of timepoints (148)
- Columns, each for one of the ROIs that are extracted (43)
**But wait!**
We originally had **50 ROIs**, what happened to 7 of them? It turns out that <code>masker</code> drops ROIs that are empty (i.e contain no brain voxels inside of them), this means that 7 of our atlas' parcels did not correspond to any region with signal! To see which ROIs are kept after computing a parcellation you can look at the <code>labels_</code> property of <code>masker</code>:
```
print(masker.labels_)
print("Number of labels", len(masker.labels_))
```
This means that our ROIs of interest (44 and 46) *cannot be accessed using the 44th and 46th columns directly*!
There are many strategies to deal with this weirdness. What we're going to do is to create a new array that fills in the regions that were removed with <code>0</code> values. It might seem a bit weird now, but it'll simplify things when we start working with multiple subjects!
First we'll identify all ROIs from the original atlas. We're going to use the <code>numpy</code> package which will provide us with functions to work with our image arrays:
```
import numpy as np
# Get the label numbers from the atlas
# Get number of labels that we have
```
Now we're going to create an array that contains:
- A number of rows matching the number of timepoints
- A number of columns matching the total number of regions
```
# Remember fMRI images are of size (x,y,z,t)
# where t is the number of timepoints
# Create an array of zeros that has the correct size
# Get regions that are kept
# Fill columns matching labels with signal values
print(final_signal.shape)
```
It's a bit of work, but now we have an array where:
1. The column number matches the ROI label number
2. Any column that is lost during the <code>masker.fit_transform</code> is filled with <code>0</code> values!
To get the columns corresponding to the regions that we've kept, we can simply use the <code>regions_kept</code> variable to select columns corresponding to the regions that weren't removed:
This is identical to the original output of <code>masker.fit_transform</code>
This might seem unnecessary for now, but as you'll see in a bit, it'll come in handy when we deal with multiple subjects!
### Calculating Connectivity
In fMRI imaging, connectivity typically refers to the *correlation of the timeseries of 2 ROIs*. Therefore we can calculate a *full connectivity matrix* by computing the correlation between *all pairs of ROIs* in our parcellation scheme!
We'll use another nilearn tool called <code>ConnectivityMeasure</code> from <code>nilearn.connectome</code>. This tool will perform the full set of pairwise correlations for us
Like the masker, we need to make an object that will calculate connectivity for us.
Try using <code>SHIFT-TAB</code> to see what options you can put into the <code>kind</code> argument of <code>ConnectivityMeasure</code>
Then we use <code>correlation_measure.fit_transform()</code> in order to calculate the full correlation matrix for our parcellated data!
Note that we're using a list <code>[final_signal]</code>, this is because <code>correlation_measure</code> works on a *list of subjects*. We'll take advantage of this later!
The result is a matrix which has:
- A number of rows matching the number of ROIs in our parcellation atlas
- A number of columns, that also matches the number of ROIs in our parcellation atlas
You can read this correlation matrix as follows:
Suppose we wanted to know the correlation between ROI 30 and ROI 40
- Then Row 30, Column 40 gives us this correlation.
- Row 40, Column 40 can also give us this correlation
This is because the correlation of $A \to B = B \to A$
***
**NOTE**
Remember we were supposed to lose 7 regions from the <code>masker.fit_transform</code> step. The correlations for these regions will be 0!
***
Let's try pulling the correlation for ROI 44 and 46!
Note that it'll be the same if we swap the rows and columns!
## Exercise
Apply the data extract process shown above to all subjects in our subject list and collect the results. Your job is to fill in the blanks!
```
# First we're going to create some empty lists to store all our data in!
ctrl_subjects = []
schz_subjects = []
# We're going to keep track of each of our subjects labels here
# pulled from masker.labels_
labels_list = []
# Get the number of unique labels in our parcellation
# We'll use this to figure out how many columns to make (as we did earlier)
atlas_labels = np.unique(yeo_7.get_fdata().astype(int))
NUM_LABELS = len(atlas_labels)
# Set the list of confound variables we'll be using
confound_variables = ['trans_x','trans_y','trans_z',
'rot_x','rot_y','rot_z',
'global_signal',
'white_matter','csf']
# Number of TRs we should drop
TR_DROP=4
# Lets get all the subjects we have
subjects = layout.get_subjects()
for sub in subjects:
#Get the functional file for the subject (MNI space)
func_file = layout.get(subject=??,
datatype='??', task='rest',
desc='??',
space='??'
extension="nii.gz",
return_type='file')[0]
#Get the confounds file for the subject (MNI space)
confound_file=layout.get(subject=??, datatype='??',
task='rest',
desc='??',
extension='tsv',
return_type='file')[0]
#Load the functional file in
func_img = nimg.load_img(??)
#Drop the first 4 TRs
func_img = func_img.slicer[??,??,??,??]
#Extract the confound variables using the function
confounds = extract_confounds(confound_file,
confound_variables)
#Drop the first 4 rows from the confound matrix
confounds = confounds[??]
# Make our array of zeros to fill out
# Number of rows should match number of timepoints
# Number of columns should match the total number of regions
fill_array = np.zeros((func_img.shape[??], ??))
#Apply the parcellation + cleaning to our data
#What function of masker is used to clean and average data?
time_series = masker.fit_transform(??,??)
# Get the regions that were kept for this scan
regions_kept = np.array(masker.labels_)
# Fill the array, this is what we'll use
# to make sure that all our array are of the same size
fill_array[:, ??] = time_series
#If the subject ID starts with a "1" then they are control
if sub.startswith('1'):
ctrl_subjects.append(fill_array)
#If the subject ID starts with a "5" then they are case (case of schizophrenia)
if sub.startswith('5'):
schz_subjects.append(fill_array)
labels_list.append(masker.labels_)
```
The result of all of this code is that:
1. Subjects who start with a "1" in their ID, are controls, and are placed into the `ctrl_subjects` list
2. Subjects who start with a "2" in their ID, have schizophrenia, and are placed into the `schz_subjects` list
What's actually being placed into the list? The cleaned, parcellated time series data for each subject (the output of <code>masker.fit_transform</code>)!
A helpful trick is that we can re-use the <code>correlation_measure</code> object we made earlier and apply it to a *list of subject data*!
At this point, we have correlation matrices for each subject across two populations. The final step is to examine the differences between these groups in their correlation between ROI 44 and ROI 46.
### Visualizing Correlation Matrices and Group Differences
An important step in any analysis is visualizing the data that we have. We've cleaned data, averaged data and calculated correlations but we don't actually know what it looks like! Visualizing data is important to ensure that we don't throw pure nonsense into our final statistical analysis
To visualize data we'll be using a python package called <code>seaborn</code> which will allow us to create statistical visualizations with not much effort.
```
import seaborn as sns
import matplotlib.pyplot as plt
```
We can view a single subject's correlation matrix by using <code>seaborn</code>'s <code>heatmap</code> function:
Recall that cleaning and parcellating the data causes some ROIs to get dropped. We dealt with this by filling an array of zeros (<code>fill_array</code>) only for columns where the regions are kept (<code>regions_kept</code>). This means that we'll have some correlation values that are 0!
This is more apparent if we plot the data slightly differently. For demonstrative purposes we've:
- Taken the absolute value of our correlations so that the 0's are the darkest color
- Used a different color scheme
```
sns.heatmap(np.abs(ctrl_correlation_matrices[0]), cmap='viridis')
```
The dark lines in the correlation matrix correspond to regions that were dropped and therefore have 0 correlation!
We can now pull our ROI 44 and 46 by indexing our list of correlation matrices as if it were a 3D array (kind of like an MR volume). Take a look at the shape:
This is of form:
<code>ctrl_correlation_matrices[subject_index, row_index, column_index]</code>
Now we're going to pull out just the correlation values between ROI 44 and 46 *across all our subjects*. This can be done using standard array indexing:
Next we're going to arrange this data into a table. We'll create two tables (called dataframes in the python package we're using, `pandas`)
```
#Create control dataframe
# Create the schizophrenia dataframe
```
The result is:
- `ctrl_df` a table containing the correlation value for each control subject, with an additional column with the group label, which is 'control'
- `scz_df` a table containing the correlation value for each schizophrenia group subject, with an additional column with the group label, which is 'schizophrenia'
For visualization we're going to stack the two tables together...
```
#Stack the two dataframes together
# Show some random samples from dataframe
```
Finally we're going to visualize the results using the python package `seaborn`!
```
#Visualize results
# Create a figure canvas of equal width and height
# Create a box plot, with the x-axis as group
#the y-axis as the correlation value
# Create a "swarmplot" as well, you'll see what this is..
# Set the title and labels of the figure
```
Although the results here aren't significant they seem to indicate that there might be three subclasses in our schizophrenia group - of course we'd need *a lot* more data to confirm this! The interpretation of these results should ideally be based on some *a priori* hypothesis!
## Congratulations!
Hopefully now you understand that:
1. fMRI data needs to be pre-processed before analyzing
2. Manipulating images in python is easily done using `nilearn` and `nibabel`
3. You can also do post-processing like confound/nuisance regression using `nilearn`
4. Parcellating is a method of simplifying and "averaging" data. The type of parcellation reflect assumptions you make about the structure of your data
5. Functional Connectivity is really just time-series correlations between two signals!
| github_jupyter |
*This notebook contains an excerpt from the [Python Data Science Handbook](http://shop.oreilly.com/product/0636920034919.do) by Jake VanderPlas; the content is available [on GitHub](https://github.com/jakevdp/PythonDataScienceHandbook).*
# Aggregation and Grouping
An essential piece of analysis of large data is efficient summarization: computing aggregations like ``sum()``, ``mean()``, ``median()``, ``min()``, and ``max()``, in which a single number gives insight into the nature of a potentially large dataset.
In this section, we'll explore aggregations in Pandas, from simple operations akin to what we've seen on NumPy arrays, to more sophisticated operations based on the concept of a ``groupby``.
```
import numpy as np
import pandas as pd
class display(object):
"""Display HTML representation of multiple objects"""
template = """<div style="float: left; padding: 10px;">
<p style='font-family:"Courier New", Courier, monospace'>{0}</p>{1}
</div>"""
def __init__(self, *args):
self.args = args
def _repr_html_(self):
return '\n'.join(self.template.format(a, eval(a)._repr_html_())
for a in self.args)
def __repr__(self):
return '\n\n'.join(a + '\n' + repr(eval(a))
for a in self.args)
```
## Planets Data
Here we will use the Planets dataset, available via the [Seaborn package](http://seaborn.pydata.org/) (see [Visualization With Seaborn](04.14-Visualization-With-Seaborn.ipynb)).
It gives information on planets that astronomers have discovered around other stars (known as *extrasolar planets* or *exoplanets* for short). It can be downloaded with a simple Seaborn command:
```
#import seaborn as sns
#planets = sns.load_dataset('planets')
planets = pd.read_csv('data/planets.csv')
planets.shape
planets.head()
planets['mass'].mean()
planets.mean()
planets.describe()
```
The following table summarizes some other built-in Pandas aggregations:
| Aggregation | Description |
|--------------------------|---------------------------------|
| ``count()`` | Total number of items |
| ``first()``, ``last()`` | First and last item |
| ``mean()``, ``median()`` | Mean and median |
| ``min()``, ``max()`` | Minimum and maximum |
| ``std()``, ``var()`` | Standard deviation and variance |
| ``mad()`` | Mean absolute deviation |
| ``prod()`` | Product of all items |
| ``sum()`` | Sum of all items |
These are all methods of ``DataFrame`` and ``Series`` objects.
## GroupBy: Split, Apply, Combine
Simple aggregations can give you a flavor of your dataset, but often we would prefer to aggregate conditionally on some label or index: this is implemented in the so-called ``groupby`` operation.
The name "group by" comes from a command in the SQL database language, but it is perhaps more illuminative to think of it in the terms first coined by Hadley Wickham of Rstats fame: *split, apply, combine*.
### Split, apply, combine
A canonical example of this split-apply-combine operation, where the "apply" is a summation aggregation, is illustrated in this figure:

This makes clear what the ``groupby`` accomplishes:
- The *split* step involves breaking up and grouping a ``DataFrame`` depending on the value of the specified key.
- The *apply* step involves computing some function, usually an aggregate, transformation, or filtering, within the individual groups.
- The *combine* step merges the results of these operations into an output array.
While this could certainly be done manually using some combination of the masking, aggregation, and merging commands covered earlier, an important realization is that *the intermediate splits do not need to be explicitly instantiated*. Rather, the ``GroupBy`` can (often) do this in a single pass over the data, updating the sum, mean, count, min, or other aggregate for each group along the way.
The power of the ``GroupBy`` is that it abstracts away these steps: the user need not think about *how* the computation is done under the hood, but rather thinks about the *operation as a whole*.
As a concrete example, let's take a look at using Pandas for the computation shown in this diagram.
We'll start by creating the input ``DataFrame``:
```
df = pd.DataFrame({'key': ['A', 'B', 'C', 'A', 'B', 'C'],
'data': range(6)}, columns=['key', 'data'])
df
```
The most basic split-apply-combine operation can be computed with the ``groupby()`` method of ``DataFrame``s, passing the name of the desired key column:
```
grouping = df.groupby('key')
grouping
for group in grouping:
print(group)
grouping.get_group('A')
df.groupby('key').sum()
```
### The GroupBy object
The ``GroupBy`` object is a very flexible abstraction.
In many ways, you can simply treat it as if it's a collection of ``DataFrame``s, and it does the difficult things under the hood. Let's see some examples using the Planets data.
Perhaps the most important operations made available by a ``GroupBy`` are *aggregate*, *filter*, *transform*, and *apply*.
We'll discuss each of these more fully in ["Aggregate, Filter, Transform, Apply"](#Aggregate,-Filter,-Transform,-Apply), but before that let's introduce some of the other functionality that can be used with the basic ``GroupBy`` operation.
#### Column indexing
The ``GroupBy`` object supports column indexing in the same way as the ``DataFrame``, and returns a modified ``GroupBy`` object.
For example:
```
planets.groupby('method')
#%timeit planets.groupby('method').median()['orbital_period']
planets.groupby('method')['orbital_period']
#planets.groupby('method')[['orbital_period', 'mass']].mean()
#%timeit planets.groupby('method')['orbital_period'].median()
```
#### Iteration over groups
The ``GroupBy`` object supports direct iteration over the groups, returning each group as a ``Series`` or ``DataFrame``:
```
for (method, group) in planets.groupby('method'):
print("{0:30s} shape={1}".format(method, group.shape))
```
#### Dispatch methods
Through some Python class magic, any method not explicitly implemented by the ``GroupBy`` object will be passed through and called on the groups, whether they are ``DataFrame`` or ``Series`` objects.
For example, you can use the ``describe()`` method of ``DataFrame``s to perform a set of aggregations that describe each group in the data:
```
#planets.groupby('method')['year'].min()
#planets.groupby('method').min()
planets.groupby('method')['year'].describe()
```
### Aggregate, filter, transform, apply
The preceding discussion focused on aggregation for the combine operation, but there are more options available.
In particular, ``GroupBy`` objects have ``aggregate()``, ``filter()``, ``transform()``, and ``apply()`` methods that efficiently implement a variety of useful operations before combining the grouped data.
For the purpose of the following subsections, we'll use this ``DataFrame``:
```
rng = np.random.RandomState(0)
df = pd.DataFrame({'key': ['A', 'B', 'C', 'A', 'B', 'C'],
'data1': range(6),
'data2': rng.randint(0, 10, 6)},
columns = ['key', 'data1', 'data2'])
df
def my_agg(x):
return len([i for i in x if i%2 == 0])
#my_agg(df['data2'])
df.groupby('key').aggregate(my_agg)
```
#### Aggregation
We're now familiar with ``GroupBy`` aggregations with ``sum()``, ``median()``, and the like, but the ``aggregate()`` method allows for even more flexibility.
It can take a string, a function, or a list thereof, and compute all the aggregates at once.
Here is a quick example combining all these:
```
df.groupby('key').aggregate(['min', np.median, max, my_agg])
```
Another useful pattern is to pass a dictionary mapping column names to operations to be applied on that column:
```
df.groupby('key').aggregate({'data1': 'min',
'data2': 'max'})
```
#### Transformation
While aggregation must return a reduced version of the data, transformation can return some transformed version of the full data to recombine.
For such a transformation, the output is the same shape as the input.
A common example is to center the data by subtracting the group-wise mean:
```
df[['data1', 'data2']] - 1
df.mean()
df
df[['data1', 'data2']] - df.mean()
df
df.groupby('key').mean()
#df[['data1', 'data2']] - df[['data1', 'data2']].mean()
df.groupby('key').transform(lambda x: x - x.mean())
```
#### Filtering
A filtering operation allows you to drop data based on the group properties.
For example, we might want to keep all groups in which the standard deviation is larger than some critical value:
```
def filter_func(x):
return (x['data2'].std() > 4) # or (x['data1'].std() < 3)
display('df', "df.groupby('key').std()", "df.groupby('key').filter(filter_func)")
```
#### The apply() method
The ``apply()`` method lets you apply an arbitrary function to the group results.
The function should take a ``DataFrame``, and return either a Pandas object (e.g., ``DataFrame``, ``Series``) or a scalar; the combine operation will be tailored to the type of output returned.
For example, here is an ``apply()`` that normalizes the first column by the sum of the second:
```
def norm_by_data2(x):
# x is a DataFrame of group values
x['data1'] /= x['data2'].sum()
x['data2'] = x['data2']*2
return x
display('df', "df.groupby('key').apply(norm_by_data2)")
```
### Specifying the split key
In the simple examples presented before, we split the ``DataFrame`` on a single column name.
This is just one of many options by which the groups can be defined, and we'll go through some other options for group specification here.
#### A list, array, series, or index providing the grouping keys
The key can be any series or list with a length matching that of the ``DataFrame``. For example:
```
df
L = [0, 1, 0, 1, 2, 0]
display('df', 'df.groupby(L).sum()')
```
Of course, this means there's another, more verbose way of accomplishing the ``df.groupby('key')`` from before:
```
display('df', "df.groupby(df['key']).sum()")
```
#### A dictionary or series mapping index to group
Another method is to provide a dictionary that maps index values to the group keys:
```
df2 = df.set_index('key')
mapping = {'A': 'vowel', 'B': 'consonant', 'C': 'consonant'}
display('df2', 'df2.groupby(mapping).sum()')
df['type'] = df['key'].apply(lambda x: mapping[x])
df.groupby('type').sum()
```
#### Any Python function
Similar to mapping, you can pass any Python function that will input the index value and output the group:
```
display('df2', 'df2.groupby(str.lower).mean()')
```
### Grouping example
As an example of this, in a couple lines of Python code we can put all these together and count discovered planets by method and by decade:
```
decade = 10 * (planets['year'] // 10)
decade = decade.astype(str) + 's'
decade.name = 'decade'
planets.groupby(['method', decade])['number'].sum().unstack().fillna(0)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/EvenSol/NeqSim-Colab/blob/master/notebooks/thermodynamics/ThermodynamicsOfHydrogen.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
#@title Thermodynamics and physical properties of hydrogen
#@markdown This page will give an introduction to how to calculate thermodynamic and physical properties of hydrogen.
#@markdown <br><br>This document is part of the module ["Introduction to Gas Processing using NeqSim in Colab"](https://colab.research.google.com/github/EvenSol/NeqSim-Colab/blob/master/notebooks/examples_of_NeqSim_in_Colab.ipynb#scrollTo=_eRtkQnHpL70).
%%capture
!pip install neqsim
import neqsim
from neqsim.thermo.thermoTools import *
import matplotlib
import numpy as np
import matplotlib.pyplot as plt
import math
plt.style.use('classic')
%matplotlib inline
```
#Litterature
https://www.engineeringtoolbox.com/hydrogen-d_1419.html
Thermodynamic and transport properties of hydrogen containing streams:
https://www.nature.com/articles/s41597-020-0568-6
#Hydrogen properties
* Hydrogen is the lightest element and will explode at concentrations ranging from 4-75 percent by volume in the presence of sunlight, a flame, or a spark.
* Despite its stability, hydrogen forms many bonds and is present in many different compounds.
* Three naturally occurring isotopes of hydrogen exist: protium, deuterium, and tritium, each with different properties due to the difference in the number of neutrons in the nucleus.
Hydrogen is the smallest chemical element because it consists of only one proton in its nucleus. Its symbol is H, and its atomic number is 1. It has an average atomic weight of 1.0079 amu, making it the lightest element. Hydrogen is the most abundant chemical substance in the universe, especially in stars and gas giant planets. However, monoatomic hydrogen is rare on Earth is rare due to its propensity to form covalent bonds with most elements. At standard temperature and pressure, hydrogen is a nontoxic, nonmetallic, odorless, tasteless, colorless, and highly combustible diatomic gas with the molecular formula H2. Hydrogen is also prevalent on Earth in the form of chemical compounds such as hydrocarbons and water.
Hydrogen has one one proton and one electron; the most common isotope, protium (1H), has no neutrons. Hydrogen has a melting point of -259.14 °C and a boiling point of -252.87 °C. Hydrogen has a density of 0.08988 g/L, making it less dense than air. It has two distinct oxidation states, (+1, -1), which make it able to act as both an oxidizing and a reducing agent. Its covalent radius is 31.5 pm.
Hydrogen exists in two different spin isomers of hydrogen diatomic molecules that differ by the relative spin of their nuclei. The orthohydrogen form has parallel spins; the parahydrogen form has antiparallel spins. At standard temperature and pressure, hydrogen gas consists of 75 percent orthohydrogen and 25 percent parahydrogen. Hydrogen is available in different forms, such as compressed gaseous hydrogen, liquid hydrogen, and slush hydrogen (composed of liquid and solid), as well as solid and metallic forms.
```
#@title Introduction to properties of hydrogen
#@markdown This video gives an intriduction to properties of hydrogen
from IPython.display import YouTubeVideo
YouTubeVideo('U-MNKK20Z_g', width=600, height=400)
#@title Calculation of vapour pressure and density of hydrogen
#@markdown Hydrogen vapour pressure and phase densities can be calculated using a classic EoS such as the Peng Robinson.
#@markdown
from neqsim.thermo import fluid, TPflash
fluid1 = fluid('pr')
fluid1.addComponent('hydrogen', 1.0)
fluid1.setPressure(1.01325, 'bara')
fluid1.setTemperature(-250.0, 'C')
dewt(fluid1);
fluid1.initProperties()
print('buble point temperature', fluid1.getTemperature('C'), " at ", fluid1.getPressure(), " bara")
print('gas density ', fluid1.getPhase('gas').getDensity())
print('liquid density ', fluid1.getPhase('oil').getDensity())
```
#Accuracy of property calculations for hydrogen
In the following section we will evaluate models for calculation of preoprties of hydrogen dominated systems.
In the following example we compare density calculations using SRK/PR EoS to the GERG-2008 EoS.
##Calculation of density of hydrogen
In the following example we evaluate the accuracy density calculation of two common EoS (SRK/PR) by comparing the the GERG-2008 EoS reference EoS.
```
#@title Select component and equation of state. Set temperature [C] and pressure range [bara]. { run: "auto" }
temperature = 20.0 #@param {type:"number"}
minPressure = 1.0 #@param {type:"number"}
maxPressure = 1000.0 #@param {type:"number"}
eosname = "srk" #@param ["srk", "pr"]
R = 8.314 # J/mol/K
# Creating a fluid in neqsim
fluid1 = fluid(eosname) #create a fluid using the SRK-EoS
fluid1.addComponent('hydrogen', 1.0) #adding 1 mole to the fluid
fluid1.init(0);
def realgasdensity(pressure, temperature):
fluid1.setPressure(pressure, 'bara')
fluid1.setTemperature(temperature, "C")
TPflash(fluid1)
fluid1.initPhysicalProperties();
return fluid1.getDensity('kg/m3')
def GERGgasdensity(pressure, temperature):
fluid1.setPressure(pressure, 'bara')
fluid1.setTemperature(temperature, 'C')
TPflash(fluid1)
return fluid1.getPhase('gas').getDensity_GERG2008()
pressure = np.arange(minPressure, maxPressure, int((maxPressure-minPressure)/100)+1)
realdensity = [realgasdensity(P,temperature) for P in pressure]
GERG2008density = [GERGgasdensity(P,temperature) for P in pressure]
deviation = [((realgasdensity(P,temperature)-GERGgasdensity(P,temperature))/GERGgasdensity(P,temperature)*100.0) for P in pressure]
print('Pure component hydrogen parameters for SRK/PR EoS ')
print('Critical temperature ', fluid1.getComponent('hydrogen').getTC(), ' [K]')
print('Critical pressure ', fluid1.getComponent('hydrogen').getPC(), ' [bara]')
print('Accentric factor ', fluid1.getComponent('hydrogen').getAcentricFactor(), ' [-]')
fig = plt.figure()
plt.subplot(2, 1, 1)
plt.plot(pressure, realdensity, '-')
plt.plot(pressure, GERG2008density, '--')
plt.xlabel('Pressure [bara]')
plt.ylabel('Density [kg/m3]')
plt.legend([eosname, 'GERG-2008'],loc='best')
plt.subplot(2, 1, 2)
title = 'Density of hydrogen at '+ str(temperature)+ ' [C]'
fig.suptitle(title)
plt.plot(pressure, deviation)
plt.xlabel('Pressure [bara]')
plt.ylabel('Deviation [%]')
```
#Phase envelope of hydrogen dominated gas mixtures
In the follwing section we will evaluate the phase behaviour of gas mixtures dominated by hydrogen.
```
import pandas as pd
naturalgas = {'ComponentName': ["hydrogen", "CO2", "methane", "ethane", "propane", "i-butane", "n-butane","i-pentane", "n-pentane"],
'MolarComposition[-]': [90.0, 0.0, 10.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0]
}
naturalgasdf = pd.DataFrame(naturalgas)
print("Natural Gas Fluid:\n")
print(naturalgasdf.head(30).to_string())
naturalgasFluid = fluid_df(naturalgasdf)
gasPhaseEnvelope = phaseenvelope(naturalgasFluid,True)
cricobar = gasPhaseEnvelope.get("cricondenbar")
cricotherm = gasPhaseEnvelope.get("cricondentherm")
print("cricobarP ", cricobar[1], " [bara] ", " cricobarT ", cricobar[0], " °K")
print("cricothermP ", cricotherm[1], " [bara] ", " cricothermT ", cricotherm[0], " °K")
```
| github_jupyter |
```
import tensorflow as tf
import matplotlib as mpl
import matplotlib.pyplot as plt
%matplotlib inline
import numpy as np
import pandas as pd
import os
import sys
import time
from tensorflow import keras
for module in (tf, mpl, np, pd, keras):
print(module.__name__, module.__version__)
```
## Load play datasets
```
fashion_mnist = keras.datasets.fashion_mnist
(x_train_all, y_train_all), (x_test, y_test) = fashion_mnist.load_data()
x_valid, x_train = x_train_all[:5000], x_train_all[5000:]
y_valid, y_train = y_train_all[:5000], y_train_all[5000:]
print(x_valid.shape, y_valid.shape)
print(x_train.shape, y_train.shape)
print(x_test.shape, y_test.shape)
```
## Normalize inputdata by using Sklearn Standscalar
- made x = (x - u) / std
```
from sklearn.preprocessing import StandardScaler
# before normalization
print(np.max(x_train), np.min(x_train))
# perform normalization
scaler = StandardScaler()
# 1. data in x_train is int32, we need to convert them to float32 first
# 2. convert x_train data from
# [None, 28, 28] -> [None, 784]
# -> after all reshape back to [None, 28, 28]
x_train_scaled = scaler.fit_transform(x_train.astype(np.float32).reshape(-1, 1)).reshape(-1, 28, 28)
x_valid_scaled = scaler.transform(x_valid.astype(np.float32).reshape(-1, 1)).reshape(-1, 28, 28)
x_test_scaled = scaler.transform(x_test.astype(np.float32).reshape(-1, 1)).reshape(-1, 28, 28)
# after normalization
print(np.max(x_train_scaled), np.min(x_train_scaled))
```
## Build model by using tf.keras high level API
```
model = keras.models.Sequential()
model.add(keras.layers.Flatten(input_shape = [28, 28]))
model.add(keras.layers.Dense(300, activation = "relu"))
model.add(keras.layers.Dense(100, activation = "relu"))
model.add(keras.layers.Dense(10, activation = "softmax"))
# if y is one_hot vector then use categorical_crossentropy as loss function
model.compile(loss = "sparse_categorical_crossentropy",
optimizer = "sgd",
metrics = ["accuracy"])
```
## Train model with callback
```
# Tensorboard, EarlyStopping, ModelCheckPoint
# https://www.tensorflow.org/versions/r2.0/api_docs/python/tf/keras/callbacks
logdir = './callbacks' # logdir for tensorboard logdata
if not os.path.exists(logdir):
os.mkdir(logdir)
output_model_file = os.path.join(logdir, "fashion_mnist_model.h5")
callbacks = [
keras.callbacks.TensorBoard(log_dir=logdir),
keras.callbacks.ModelCheckpoint(output_model_file, save_best_only = True),
keras.callbacks.EarlyStopping(patience = 5, min_delta = 1e-3)
]
history = model.fit(x = x_train_scaled, y = y_train, epochs = 10,
validation_data = (x_valid_scaled, y_valid),
callbacks = callbacks)
# once training is finished, you can use tensorboard to visualize results
# tensorboard --logdir=callbacks
```
## Result visualization
```
from tensorflow.python.keras.callbacks import History
def plot_learning_curves(history: History):
pd.DataFrame(history.history).plot(figsize = (8, 5))
plt.grid(True)
plt.gca().set_ylim(0, 1)
plt.show()
plot_learning_curves(history)
```
## Evaluate model with test dataset
```
test_loss, test_acc = model.evaluate(x_test_scaled, y_test)
```
## Made a simple prediction on test dataset
```
# one_hot encoded results
predictions = model.predict(x_test)
print(predictions[0])
# compare ml-prediction to test label
print(np.argmax(predictions[0]), y_test[0])
```
| github_jupyter |
# प्रशिक्षण: खासगी डीप लर्निंगचे मूलभूत साधने
गोपनीयता जतन, विकेंद्रित गहन शिक्षणासाठी PySyft च्या प्रास्ताविक प्रशिक्षणात आपले स्वागत आहे. नोटबुकची ही मालिका एका अधिकाराखाली केंद्रीकृत न करता गुप्त / खाजगी डेटा / मॉडेल्सवर सखोल शिक्षण घेण्यासाठी आवश्यक असलेली नवीन साधने आणि तंत्रे जाणून घेण्यासाठी आपल्याला चरण-दर-चरण मार्गदर्शक आहेत.
**व्याप्ती:** लक्षात ठेवा आम्ही केवळ डेटाचे विकेंद्रीकरण(decentralized) /
एन्क्रिप्ट(encrypt) कसे करावे याबद्दल बोलत आहोत असे नाही तर डेटा संग्रहित आणि क्वेरी(query) केलेल्या डेटाबेस-समवेत डेटाच्या भोवतालच्या परिसंस्थेचे विकेंद्रीकरण करण्यात आणि डेटामधून माहिती काढण्यासाठी वापरल्या जाणार्या न्यूरल मॉडेल्सच्या(neural network)संधरबात PySyft चा कसा उपयोग करता येईल यावर आम्ही चर्चा करीत आहोत. जसजसे PySyft मध्ये नवीन extension तयार होत जातील, तसतसे या नोटबुकमध्ये नवीन कार्यक्षमतेचे स्पष्टीकरण देण्यासाठी नवीन ट्यूटोरियल पाठवले जाईल.
लेखक:
- Andrew Trask - Twitter: [@iamtrask](https://twitter.com/iamtrask)
अनुवादक:
- Krunal Kshirsagar - Twitter : [@krunal_wrote](https://twitter.com/krunal_wrote) - Github: [@Noob-can-Compile](https://github.com/Noob-can-Compile)
## सुरवातीचा आराखडा:
- भाग 1: खाजगी डीप लर्निंगचे मूळ साधने.
## हे ट्यूटोरियल का घ्यायचे?
**1) स्पर्धात्मक कारकीर्दीचा फायदा** - मागील 20 वर्षांपासून, डिजिटल क्रांतीमुळे डेटा जास्त प्रमाणात उपलब्ध झाला आहे कारण एनालॉग प्रक्रिया डिजिटल झाल्या आहेत. तथापि, [GDPR](https://eugdpr.org/) नवीन नियमनानुसार उद्योजकांवर कसे वापरावे - आणि महत्त्वाचे म्हणजे ते कसे विश्लेषित करतात - वैयक्तिक माहिती यावर कमी स्वातंत्र्य मिळवण्याचा दबाव असतो. **Bottom Line:** डेटा शास्त्रज्ञांना जुन्या(old school) साधनांसह जास्तीत जास्त डेटामध्ये प्रवेश मिळणार नाही, परंतु खाजगी डीप लर्निंगची साधने शिकून आपण या वक्रतेपेक्षा पुढे असू शकता आणि आपल्या कारकीर्दीत स्पर्धात्मक फायदा घेऊ शकता.
**2) उद्योजकीय संधी** - समाजात डिप्प लर्निंग सोडवू शकणार्या अनेक समस्या आहेत, परंतु बर्याच महत्त्वाच्या गोष्टींचा शोध लावला गेला नाही कारण त्यासाठी लोकांबद्दल आश्चर्यकारकपणे संवेदनशील माहिती मिळवणे आवश्यक आहे (तुम्ही डीप लर्निंगचा वापर मानसिक किंवा नातेसंबंध समस्या सोढवण्यासाठी लोकांना मदत करुशकतात!). अशाप्रकारे, खाजगी डीप लर्निंग शिकणे आपल्यासाठी नवीन स्टार्टअप संधींचे संपूर्ण विस्तार उघडते जे या टूलसेटशिवाय इतरांना पूर्वी उपलब्ध नव्हते.
**3) सोशल गुड** - डीप लर्निंगचा उपयोग खर्या जगातील विविध समस्या सोडविण्यासाठी केला जाऊ शकतो, परंतु वैयक्तिक माहितीवर डीप लर्निंग म्हणजे लोकांसंबंधी, लोकांसाठी डीप लर्निंग. आपल्या मालकीच्या नसलेल्या डेटावर डीप लर्निंग कसे करावे हे शिकणे हे करिअर किंवा उद्योजकीय संधीपेक्षा अधिक प्रतिनिधित्व करते, लोकांच्या जीवनातील काही सर्वात वैयक्तिक आणि महत्त्वपूर्ण समस्या सोडविण्यास मदत करण्याची संधी - आणि तेही प्रमाणात करणे.
## मला अतिरिक्त क्रेडिट कसे मिळेल?
- Pysyft ला Github वर Star करा! - [https://github.com/OpenMined/PySyft](https://github.com/OpenMined/PySyft)
- ह्या नोटबुकची माहिती देणारा किंवा शिकवणारा एक यूट्यूब व्हिडिओ बनवा!
... ठीक आहे ... चला सुरुवात करूया!
# भाग -1: पूर्वनिर्धारितता
- PyTorch जाणून घ्या - नसल्यास http://fast.ai कोर्स घ्या आणि परत या..
- PySyft फ्रेमवर्क पेपर https://arxiv.org/pdf/1811.04017.pdf वाचा! हे आपल्याला PySyft कसे तयार केले आहे यावर एक सखोल पार्श्वभूमी देईल जे गोष्टींना अधिक अर्थ प्राप्त करण्यास मदत करेल.
# भाग 0: सेटअप
सुरू करण्यासाठी, आपल्याला खात्री करुन घ्यावी लागेल की आपल्याकडे योग्य गोष्टी स्थापित केल्या आहेत. असे करण्यासाठी, PySyft च्या readme जा आणि सेटअपच्या सूचनांचे अनुसरण करा. बहुतेक लोकांसाठी TLDR आहे.
- Install Python 3.5 or higher
- Install PyTorch 1.1
- Clone PySyft (git clone https://github.com/OpenMined/PySyft.git)
- cd PySyft
- pip install -r pip-dep/requirements.txt
- pip install -r pip-dep/requirements_udacity.txt
- python setup.py install udacity
- python setup.py test
जर यापैकी कोणताही भाग आपल्यासाठी कार्य करत नसेल (किंवा कोणत्याही चाचण्या अयशस्वी झाल्या असतील तर) - प्रथम इन्स्टॉलेशन मदतीसाठी [README](https://github.com/OpenMined/PySyft.git) तपासा आणि नंतर एक GitHub Issue उघडा किंवा आमच्या स्लॅकमध्ये #beginner चॅनेलला पिंग करा! [slack.openmined.org](http://slack.openmined.org/)
```
# गोष्टी कार्यरत आहेत की नाही हे पाहण्यासाठी हा सेल चालवा
import sys
import torch
from torch.nn import Parameter
import torch.nn as nn
import torch.nn.functional as F
import syft as sy
hook = sy.TorchHook(torch)
torch.tensor([1,2,3,4,5])
```
जर या सेलची अंमलबजावणी झाली तर आपण शर्यतींसाठी निघाला आहात! चला हे करूया!
# भाग 1: खासगी (Private), विकेंद्रीकरण (Decentralized ) डेटा विज्ञान चे मूलभूत साधने
तर - आपण याच्यात पहिला प्रश्न विचारू शकता तो असा आहे - आपल्याकडे प्रवेश नसलेल्या डेटावर,आपण आपले मॉडेल कसे प्रशिक्षीत करू सकतो ?
पण, उत्तर आश्चर्यकारकपणे सोपे आहे. जर आपण PyTorch मध्ये काम करण्याची सवय लावत असाल, तर आपण torch.Tensor सारख्या वस्तूंच्या मदतीने काम करण्याची सवय लावली आहे!
```
x = torch.tensor([1,2,3,4,5])
y = x + x
print(y)
```
अर्थात हे सुपर फॅन्सी (आणि शक्तिशाली!) टेन्सर वापरणे महत्वाचे आहे, परंतु आपल्याकडे आपल्या स्थानिक मशीनवर डेटा असणे आवश्यक आहे. येथून आपला प्रवास सुरू होतो.
# विभाग 1.1 - बॉबच्या मशीनवर Tensor पाठवणे.
सामान्यत: आपण डेटा ठेवणाऱ्या मशीनवर डेटा सायन्स / डीप लर्निंगचे प्रदर्शन करत असे, आता आपल्याला दुसर्या मशीनवर या प्रकारचे गणित करायचे आहे. विशेषतः, आपण यापुढे डेटा आपल्या स्थानिक मशीनवर आहे असे समजू शकत नाही.
अशाप्रकारे, Torch Tensor वापरण्याऐवजी, आपण आता पॉईंटर्ससह टेन्सरवर काम करणार आहोत. मला काय म्हणायचे आहे ते मी तुम्हाला दाखवतो. प्रथम, "सोंग"(pretend) मशीन बनवू जे "सोंग"(pretend) व्यक्तीच्या मालकीचे आहे - आपण त्याला बॉब(Bob) म्हणू.
```
bob = sy.VirtualWorker(hook, id="bob")
```
समजा बॉबचे मशीन दुसर्या ग्रहावर आहे - कदाचित मंगळावर! पण, याक्षणी मशीन रिक्त आहे. चला काही डेटा तयार करू जेणेकरुन आपण ते बॉबला पाठवू आणि पॉईंटर्सबद्दल (Pointers) शिकू!
```
x = torch.tensor([1,2,3,4,5])
y = torch.tensor([1,1,1,1,1])
```
आणि आता - चला आपले टेन्सर बॉबला(Bob) पाठवू !!
```
x_ptr = x.send(bob)
y_ptr = y.send(bob)
x_ptr
```
BOOM! आता बॉबकडे(Bob) दोन Tensor आहेत! माझ्यावर विश्वास नाही? हे स्वत: पहा!
```
bob._objects
z = x_ptr + x_ptr
z
bob._objects
```
आता थोड लक्षात घ्या. जेव्हा आपण `x.send(bob)` म्हटले तेव्हा ते नवीन ऑब्जेक्ट परत केले ज्याला आपण `x_ptr` म्हटले. एका टेन्सरसाठी हा आमचा पहिला पॉईंटर आहे. Pointers to tensors प्रत्यक्षात स्वत: चा डेटा ठेवत नाहीत. त्याऐवजी, ते फक्त मशीनवर संग्रहित एका Tensor(डेटासह) बद्दल मेटाडेटा असतात. या टेन्सरचा हेतू आम्हाला अंतर्ज्ञानी एपीआय प्रदान करणे आहे जे या टेन्सरद्वारे इतर मशीनला कार्ये मोजण्यास सांगते. चला Pointer असलेल्या मेटाडेटावर एक नजर टाकू.
```
x_ptr
```
तो मेटाडेटा पहा!
Pointers शी संबंधित दोन मुख्य वैषिष्टे आहेत:
- `x_ptr.location : bob`, स्थान, (Pointer) ज्या ठिकाणी निर्देशित करीत आहे त्या स्थानाचा संदर्भ.
- `x_ptr.id_at_location : <random integer>`, ज्या ठिकाणी Tensor संग्रहित असेल तो आयडी.
ह्या स्वरूपात मुद्रित आहेत `<id_at_location>@<location>`
इतरही सामान्य गुणधर्म आहेत:
- `x_ptr.id : <random integer>`, आपल्या पॉईंटर Tensor चा आयडी, यादृच्छिकपणे(Randomly Allocated) त्याचा वाटप करण्यात आला.
- `x_ptr.owner : "me"`, पॉईंटर (Pointer) टेन्सरचा मालक असलेला वर्कर, येथे तो स्थानिक वर्कर आहे, ज्याचे नाव "me" आहे.
```
x_ptr.location
bob
bob == x_ptr.location
x_ptr.id_at_location
x_ptr.owner
```
आपण आश्चर्यचकित होऊ शकता की (Pointer) चा मालक असलेला स्थानिक कर्मचारीसुद्धा एक VirtualWorker आहे, जरी आम्ही तो तयार केलेला नाही. मजेदार तथ्य, जसे आपल्याकडे (Bob)साठी VirtualWorker ऑब्जेक्ट होता, तसे आपल्याकडे देखील (डीफॉल्टनुसार) नेहमीच असतो. जेव्हा आम्ही hook = sy.TorchHook() म्हणतो तेव्हा हा कार्यकर्ता आपोआप तयार होतो आणि म्हणूनच आपल्याला सहसा ते तयार करण्याची आवश्यकता नसते.
```
me = sy.local_worker
me
me == x_ptr.owner
```
आणि शेवटी, ज्याप्रमाणे आपण Tensor वर .send() कॉल करू शकतो, तसे Tensor परत मिळवण्यासाठी आपण पॉईंटर (Pointer) वर .get () कॉल करू शकतो !!!
```
x_ptr
x_ptr.get()
y_ptr
y_ptr.get()
z.get()
bob._objects
```
आणि जसे आपण पाहू शकता ... बॉब(Bob)कडे यापुढे टेन्सर नाही !!! ते परत आपल्या मशीनकडे गेले आहेत!
# विभाग 1.2 - Using Tensor Pointers
तर, बॉब(Bob)कडून टेन्सर(Tensor) पाठविणे आणि प्राप्त करणे चांगले आहे, परंतु हे डिप्प लर्निंगच नाही! आपल्याला रिमोट टेन्सर(Remote Tensor)वर Tensor _operations_ करण्यास सक्षम व्हायचे आहे. सुदैवाने, Tensor पॉईंटर्स(Pointers) हे बरेच सोपे करतात! आपण सामान्य Tensor सारखेच पॉईंटर्स(Pointers) वापरू शकता!
```
x = torch.tensor([1,2,3,4,5]).send(bob)
y = torch.tensor([1,1,1,1,1]).send(bob)
z = x + y
z
```
आणि पहा (And voilà!)
पडद्यामागे काहीतरी खूप शक्तिशाली घडले. स्थानिकरित्या x आणि y चे संगणन करण्याऐवजी, आज्ञा क्रमबद्ध केली गेली आणि बॉब(Bob)ला पाठविली गेली, त्याने गणना केली, एक टेन्सर(Tensor) z तयार केला, आणि नंतर पॉईंटरला z पाठवून परत आपल्याकडे परत केले!
आपण पॉईंटरवर(Pointer) .get () कॉल केल्यास आपल्याला आपल्या मशीनवर पुन्हा निकाल मिळेल!
```
z.get()
```
### Torch कार्य (Functions)
हा एपीआय(API) Torch चे सर्व ऑपरेशन्समध्ये विस्तारित केला गेला आहे !!!
```
x
y
z = torch.add(x,y)
z
z.get()
```
### व्हेरिएबल्स (Variables) (backpropagation सह!)
```
x = torch.tensor([1,2,3,4,5.], requires_grad=True).send(bob)
y = torch.tensor([1,1,1,1,1.], requires_grad=True).send(bob)
z = (x + y).sum()
z.backward()
x = x.get()
x
x.grad
```
म्हणूनच आपण पाहू शकता, एपीआय(API) खरोखरच लवचिक आहे आणि आपण साधारणपणे रिमोट डेटावरील(Remote Data) Torch मध्ये करता असे कोणतेही ऑपरेशन करण्यास सक्षम आहे. हे आमच्या अधिक प्रगत गोपनीयता संरक्षित प्रोटोकॉल(advanced privacy preserving protocols) जसे की फेडरेटेड लर्निंग(Federated Learning), सिक्युर मल्टी-पार्टी कंप्यूटेशन(Secure Multi-Party Computation) आणि डिफरेंशियल प्रायव्हसी(Differential Privacy) यासाठी आधारभूत कार्य करते!
# अभिनंदन !!! - समुदायात सामील होण्याची वेळ आली!
हे नोटबुक ट्यूटोरियल पूर्ण केल्याबद्दल अभिनंदन! आपण याचा आनंद घेत असल्यास आणि एआय(AI) आणि एआय सप्लाय चेन (डेटा) च्या विकेंद्रित(Decentralized) मालकीच्या गोपनीयतेच्या संरक्षणाच्या दिशेने चळवळीत सामील होऊ इच्छित असाल तर आपण हे खालील प्रकारे करू शकता!
### Pysyft ला Github वर Star करा!
आमच्या समुदायाला मदत करण्याचा सर्वात सोपा मार्ग म्हणजे फक्त गिटहब(GitHub) रेपो(Repo) तारांकित(Star) करणे! हे आम्ही तयार करीत असलेल्या छान साधनांविषयी जागरूकता वाढविण्यास मदत करते.
- [Star PySyft](https://github.com/OpenMined/PySyft)
### आमच्या Slack मध्ये सामील व्हा!
नवीनतम प्रगतीवर अद्ययावत राहण्याचा उत्तम मार्ग म्हणजे आमच्या समुदायामध्ये सामील होणे! आपण [http://slack.openmined.org](http://slack.openmined.org) येथे फॉर्म भरुन तसे करू शकता.
### एका कोड प्रोजेक्टमध्ये सामील व्हा!
आमच्या समुदायामध्ये योगदानाचा उत्तम मार्ग म्हणजे कोड योगदानकर्ता बनणे! कोणत्याही वेळी आपण (PySyft GitHub Issues Page) वर जाऊ शकता आणि "Project" साठी फिल्टर करू शकता. हे आपण कोणत्या प्रकल्पांमध्ये सामील होऊ शकता याबद्दल विहंगावलोकन देणारी सर्व उच्च स्तरीय तिकिटे दर्शवेल! आपण एखाद्या प्रकल्पात सामील होऊ इच्छित नसल्यास, परंतु आपण थोडं कोडिंग करू इच्छित असाल तर आपण "good first issues" म्हणून चिन्हांकित गिटहब(GitHub) अंक शोधून आणखी "one off" मिनी-प्रकल्प(mini project) शोधू शकता.
- [PySyft Projects](https://github.com/OpenMined/PySyft/issues?q=is%3Aopen+is%3Aissue+label%3AProject)
- [Good First Issue Tickets](https://github.com/OpenMined/PySyft/issues?q=is%3Aopen+is%3Aissue+label%3A%22good+first+issue%22)
### दान करा
आपल्याकडे आमच्या कोडेबेसमध्ये योगदान देण्यास वेळ नसल्यास, परंतु तरीही आपल्याला समर्थन द्यावयाचे असल्यास आपण आमच्या मुक्त संग्रहात बॅकर देखील होऊ शकता. सर्व देणगी आमच्या वेब होस्टिंग आणि हॅकॅथॉन आणि मेटअप्स सारख्या इतर सामुदायिक खर्चाकडे जातात!
[OpenMined's Open Collective Page](https://opencollective.com/openmined)
| github_jupyter |
```
%pylab inline
from numpy.lib.recfunctions import *
def filternan(a):
return a[isnan(a) != True]
segments = genfromtxt("segments.csv", delimiter=',', names=True)
names = genfromtxt("names.csv", delimiter=',', names=True)
callsigns = genfromtxt("callsigns.csv", delimiter=',', names=True)
imos = genfromtxt("imos.csv", delimiter=',', names=True)
fishing = genfromtxt("../Combinedfishing_2014.csv", delimiter=',', names=True)
fishing = append_fields(fishing, 'fishing', [], dtypes='<f4')
fishing['fishing'] = 1.0
tileset = genfromtxt("pipeline_205_08_07_13_33_31_mmsi_list.csv", delimiter=',', names=True)
tileset = append_fields(tileset, 'tileset', [], dtypes='<f4')
tileset['tileset'] = 1.0
d = join_by('mmsi', segments, names, jointype='outer', asrecarray=True)
d = join_by('mmsi', d, imos, jointype='outer', asrecarray=True)
d = join_by('mmsi', d, callsigns, jointype='outer', asrecarray=True)
d = join_by('mmsi', d, fishing, jointype='outer', asrecarray=True)
d = join_by('mmsi', d, tileset, jointype='outer', asrecarray=True)
d = d.filled(nan)
savez("mmsi_counts.npz", d)
d = load("mmsi_counts.npz")['arr_0']
f = d[d['fishing'] == 1]
histfig = figure(figsize=(20,5))
hist(filternan(f['seg_count']), bins=50, normed=False, color='b', alpha=0.5, log=True, label="segment count")
legend(); show()
histfig = figure(figsize=(20,5))
hist(filternan(f['radius']), bins=200, normed=False, color='r', alpha=0.5, log=True, label="radius")
legend(); show()
histfig = figure(figsize=(20,5))
hist(filternan(f['name_count']), bins=200, normed=False, color='b', alpha=0.5, log=True, label="name count")
legend(); show()
histfig = figure(figsize=(20,5))
hist(filternan(f['imo_count']), bins=200, normed=False, color='b', alpha=0.5, log=True, label="imo count")
legend(); show()
histfig = figure(figsize=(20,5))
hist(filternan(f['callsign_count']), bins=200, normed=False, color='b', alpha=0.5, log=True, label="callsign count")
legend(); show()
def compare(col1, col2):
col1_coord = unique(filternan(d[col1]))
col1_coord.sort()
col2_coord = unique(filternan(d[col2]))
col2_coord.sort()
all_counts = zeros((col1_coord.shape[0], col2_coord.shape[0]))
for i1, c1 in enumerate(col1_coord):
for i2, c2 in enumerate(col2_coord):
v = d[d[col1] > c1]
v = v[v[col2] > c2]
all_counts[i1,i2] = v.shape[0]
return col1_coord, col2_coord, all_counts
c1, c2, p = compare('seg_count', 'name_count')
x = log(p)
imshow(x); show()
imshow(x[:20,:20]); show()
imshow(x[:10,:10]); show()
f = d[d['tileset'] == 1]
print "#### 0.2"
print f[f['seg_count'] == 2.0][200]['mmsi']
print f[f['seg_count'] == 2.0][201]['mmsi']
print f[f['seg_count'] == 2.0][202]['mmsi']
print f[f['seg_count'] == 2.0][203]['mmsi']
print "#### 0.5"
print f[f['seg_count'] == 5.0][200]['mmsi']
print f[f['seg_count'] == 5.0][201]['mmsi']
print f[f['seg_count'] == 5.0][202]['mmsi']
print f[f['seg_count'] == 5.0][203]['mmsi']
```
* 0.2
* 205336000 no apparent spoofing: http://alpha-dev.globalfishingwatch.org/viz?workspace=/workspace/970bbf0e-1ddb-4d63-a46b-4d1b0a638201
* 205346000 three vessels: http://alpha-dev.globalfishingwatch.org/viz?workspace=/workspace/692d75af-91b5-4948-8bf4-4edb41263551
* 205482090 multiple vessels, NOT FISHING: http://alpha-dev.globalfishingwatch.org/viz?workspace=/workspace/f44fad67-549b-4c8f-95fb-6a1becdba350
* 205532000 multiple vessels: http://alpha-dev.globalfishingwatch.org/viz?workspace=/workspace/fd9a132c-65ff-49a7-821e-a152a0682a8c
* 0.5
* 440071000 two vessels: http://alpha-dev.globalfishingwatch.org/viz?workspace=/workspace/46d19ae5-e395-4c69-bea3-d4dc75113b9b
* 440107870 two vessels: http://alpha-dev.globalfishingwatch.org/viz?workspace=/workspace/5c0d7a81-335d-4c96-87a6-058e6c0e1f55
* 440111111 not properly despoofed, mulytiple vessels, broken points: http://alpha-dev.globalfishingwatch.org/viz?workspace=/workspace/483382d5-243d-49c9-8019-edef6d7c72ed
* 440120530 four vessels or bad despoofing: http://alpha-dev.globalfishingwatch.org/viz?workspace=/workspace/06ff655c-171a-4357-a3df-6f6ade288694
```
f = d[d['tileset'] == 1]
print "#### 0.2"
print f[f['name_count'] == 2.0][200]['mmsi']
print f[f['name_count'] == 2.0][201]['mmsi']
print f[f['name_count'] == 2.0][202]['mmsi']
print f[f['name_count'] == 2.0][203]['mmsi']
print "#### 0.5"
print f[f['name_count'] == 5.0][200]['mmsi']
print f[f['name_count'] == 5.0][201]['mmsi']
print f[f['name_count'] == 5.0][202]['mmsi']
print f[f['name_count'] == 5.0][203]['mmsi']
```
* 0.2
* 211495000 spoofing, no distinct vessels: http://alpha-dev.globalfishingwatch.org/viz?workspace=/workspace/b5128762-194c-4478-a808-c1bdb4bb4ea7
* 211501000 no spoofing: http://alpha-dev.globalfishingwatch.org/viz?workspace=/workspace/45516aa0-290f-4ae0-aff1-0f3825dd92a5
* 211502000 borken locations: http://alpha-dev.globalfishingwatch.org/viz?workspace=/workspace/ae46157c-a61e-47a8-974e-78bc2adf0d9e
* 211516530 no spoofing: http://alpha-dev.globalfishingwatch.org/viz?workspace=/workspace/362fd639-c19d-44e5-b979-5941aa8bdb58
* 0.5
* 261000660 bad despoofing?: http://alpha-dev.globalfishingwatch.org/viz?workspace=/workspace/76fe3ed2-8627-4d3c-bf7c-055a1c914e83
* 261003220 multiple vessels: http://alpha-dev.globalfishingwatch.org/viz?workspace=/workspace/664f36bd-bff2-42b5-9c1c-60104b02f558
* 261003640 multiple vessels, bad despoofing: http://alpha-dev.globalfishingwatch.org/viz?workspace=/workspace/efce1c0f-f902-4a08-90eb-db228688b246
* 261004780 bad positions, spoofing: http://alpha-dev.globalfishingwatch.org/viz?workspace=/workspace/b7604ea5-49b9-4a9d-9d67-11327aeaeef2
| github_jupyter |
# Text-mined dataset
```
!python ../../s4/scripts/001.thermonize_dmm.py \
--download-fn intermediate_data/Reactions_Solid_State_legacy_v14_2.pypickle \
--mp-output intermediate_data/DMM_Thermonized_MP.pypickle \
--freed-output intermediate_data/DMM_Thermonized_FREED.pypickle
import pickle
from collections import Counter
with open('intermediate_data/DMM_Thermonized_MP.pypickle', 'rb') as f:
data = pickle.load(f)
for k in data[1]:
v = data[1][k]
error_counting = Counter(x['exception_desc'] for x in v)
print('='*70)
print('Exception', k)
for err, cnt in error_counting.items():
print('%04d %s' % (cnt, err))
!python ../../s4/scripts/002.compute_cascades.py \
--reactions-file intermediate_data/DMM_Thermonized_MP.pypickle \
--output-fn intermediate_data/DMM_Cascades.pypickle \
--cascade-temperatures '[500,750,1000,1250,1500,1750]'
import pickle
from collections import defaultdict, Counter
with open('intermediate_data/DMM_Cascades.pypickle', 'rb') as f:
data = pickle.load(f)
cascade_error = defaultdict(Counter)
for key in data:
for k, reaction in data[key]:
for temp, cascade in reaction.items():
if cascade['error']:
cascade_error[cascade['exception']][cascade['exception_desc']] += 1
q = cascade
for exp_name in cascade_error:
print('='*70)
print('Exception', exp_name)
for err, cnt in cascade_error[exp_name].items():
print('%04d %s' % (cnt, err))
!python ../../s4/scripts/003.prepare_ml_data.py \
--reactions-file intermediate_data/DMM_Thermonized_MP.pypickle \
--text-mined-recipes-file intermediate_data/Reactions_Solid_State_legacy_v14_2.pypickle \
--output-fn intermediate_data/DMM_TrainingData.pypickle \
--cascade-file intermediate_data/DMM_Cascades.pypickle
```
# PCD dataset
```
with open('intermediate_data/PCD_Reactions.pypickle', 'wb') as f:
pickle.dump((data, {}), f)
!python ../../s4/scripts/002.compute_cascades.py \
--reactions-file intermediate_data/PCD_Reactions.pypickle \
--output-fn intermediate_data/PCD_Cascades.pypickle \
--cascade-temperatures '[1000]'
import pickle
from collections import defaultdict, Counter
with open('intermediate_data/PCD_Cascades.pypickle', 'rb') as f:
data = pickle.load(f)
cascade_error = defaultdict(Counter)
for key in data:
for k, reaction in data[key]:
for temp, cascade in reaction.items():
if cascade['error']:
cascade_error[cascade['exception']][cascade['exception_desc']] += 1
q = cascade
for exp_name in cascade_error:
print('='*70)
print('Exception', exp_name)
for err, cnt in cascade_error[exp_name].items():
print('%04d %s' % (cnt, err))
!python ../../s4/scripts/003.prepare_ml_data.py \
--reactions-file intermediate_data/PCD_Reactions.pypickle \
--output-fn intermediate_data/PCD_TrainingData.pypickle \
--cascade-file intermediate_data/PCD_Cascades.pypickle
```
| github_jupyter |
##### Copyright 2019 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# 循环神经网络(RNN)文本生成
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://tensorflow.google.cn/tutorials/text/text_generation"><img src="https://tensorflow.google.cn/images/tf_logo_32px.png" />在 tensorflow.google.cn 上查看</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/zh-cn/tutorials/text/text_generation.ipynb"><img src="https://tensorflow.google.cn/images/colab_logo_32px.png" />在 Google Colab 运行</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/zh-cn/tutorials/text/text_generation.ipynb"><img src="https://tensorflow.google.cn/images/GitHub-Mark-32px.png" />在 GitHub 上查看源代码</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/docs-l10n/site/zh-cn/tutorials/text/text_generation.ipynb"><img src="https://tensorflow.google.cn/images/download_logo_32px.png" />下载此 notebook</a>
</td>
</table>
本教程演示如何使用基于字符的 RNN 生成文本。我们将使用 Andrej Karpathy 在[《循环神经网络不合理的有效性》](http://karpathy.github.io/2015/05/21/rnn-effectiveness/)一文中提供的莎士比亚作品数据集。给定此数据中的一个字符序列 (“Shakespear”),训练一个模型以预测该序列的下一个字符(“e”)。通过重复调用该模型,可以生成更长的文本序列。
请注意:启用 GPU 加速可以更快地执行此笔记本。在 Colab 中依次选择:*运行时 > 更改运行时类型 > 硬件加速器 > GPU*。如果在本地运行,请确保 TensorFlow 的版本为 1.11 或更高。
本教程包含使用 [tf.keras](https://tensorflow.google.cn/programmers_guide/keras) 和 [eager execution](https://tensorflow.google.cn/programmers_guide/eager) 实现的可运行代码。以下是当本教程中的模型训练 30 个周期 (epoch),并以字符串 “Q” 开头时的示例输出:
<pre>
QUEENE:
I had thought thou hadst a Roman; for the oracle,
Thus by All bids the man against the word,
Which are so weak of care, by old care done;
Your children were in your holy love,
And the precipitation through the bleeding throne.
BISHOP OF ELY:
Marry, and will, my lord, to weep in such a one were prettiest;
Yet now I was adopted heir
Of the world's lamentable day,
To watch the next way with his father with his face?
ESCALUS:
The cause why then we are all resolved more sons.
VOLUMNIA:
O, no, no, no, no, no, no, no, no, no, no, no, no, no, no, no, no, no, no, no, no, it is no sin it should be dead,
And love and pale as any will to that word.
QUEEN ELIZABETH:
But how long have I heard the soul for this world,
And show his hands of life be proved to stand.
PETRUCHIO:
I say he look'd on, if I must be content
To stay him from the fatal of our country's bliss.
His lordship pluck'd from this sentence then for prey,
And then let us twain, being the moon,
were she such a case as fills m
</pre>
虽然有些句子符合语法规则,但是大多数句子没有意义。这个模型尚未学习到单词的含义,但请考虑以下几点:
* 此模型是基于字符的。训练开始时,模型不知道如何拼写一个英文单词,甚至不知道单词是文本的一个单位。
* 输出文本的结构类似于剧本 -- 文本块通常以讲话者的名字开始;而且与数据集类似,讲话者的名字采用全大写字母。
* 如下文所示,此模型由小批次 (batch) 文本训练而成(每批 100 个字符)。即便如此,此模型仍然能生成更长的文本序列,并且结构连贯。
## 设置
### 导入 TensorFlow 和其他库
```
import tensorflow as tf
import numpy as np
import os
import time
```
### 下载莎士比亚数据集
修改下面一行代码,在你自己的数据上运行此代码。
```
path_to_file = tf.keras.utils.get_file('shakespeare.txt', 'https://storage.googleapis.com/download.tensorflow.org/data/shakespeare.txt')
```
### 读取数据
首先,看一看文本:
```
# 读取并为 py2 compat 解码
text = open(path_to_file, 'rb').read().decode(encoding='utf-8')
# 文本长度是指文本中的字符个数
print ('Length of text: {} characters'.format(len(text)))
# 看一看文本中的前 250 个字符
print(text[:250])
# 文本中的非重复字符
vocab = sorted(set(text))
print ('{} unique characters'.format(len(vocab)))
```
## 处理文本
### 向量化文本
在训练之前,我们需要将字符串映射到数字表示值。创建两个查找表格:一个将字符映射到数字,另一个将数字映射到字符。
```
# 创建从非重复字符到索引的映射
char2idx = {u:i for i, u in enumerate(vocab)}
idx2char = np.array(vocab)
text_as_int = np.array([char2idx[c] for c in text])
```
现在,每个字符都有一个整数表示值。请注意,我们将字符映射至索引 0 至 `len(unique)`.
```
print('{')
for char,_ in zip(char2idx, range(20)):
print(' {:4s}: {:3d},'.format(repr(char), char2idx[char]))
print(' ...\n}')
# 显示文本首 13 个字符的整数映射
print ('{} ---- characters mapped to int ---- > {}'.format(repr(text[:13]), text_as_int[:13]))
```
### 预测任务
给定一个字符或者一个字符序列,下一个最可能出现的字符是什么?这就是我们训练模型要执行的任务。输入进模型的是一个字符序列,我们训练这个模型来预测输出 -- 每个时间步(time step)预测下一个字符是什么。
由于 RNN 是根据前面看到的元素维持内部状态,那么,给定此时计算出的所有字符,下一个字符是什么?
### 创建训练样本和目标
接下来,将文本划分为样本序列。每个输入序列包含文本中的 `seq_length` 个字符。
对于每个输入序列,其对应的目标包含相同长度的文本,但是向右顺移一个字符。
将文本拆分为长度为 `seq_length+1` 的文本块。例如,假设 `seq_length` 为 4 而且文本为 “Hello”, 那么输入序列将为 “Hell”,目标序列将为 “ello”。
为此,首先使用 `tf.data.Dataset.from_tensor_slices` 函数把文本向量转换为字符索引流。
```
# 设定每个输入句子长度的最大值
seq_length = 100
examples_per_epoch = len(text)//seq_length
# 创建训练样本 / 目标
char_dataset = tf.data.Dataset.from_tensor_slices(text_as_int)
for i in char_dataset.take(5):
print(idx2char[i.numpy()])
```
`batch` 方法使我们能轻松把单个字符转换为所需长度的序列。
```
sequences = char_dataset.batch(seq_length+1, drop_remainder=True)
for item in sequences.take(5):
print(repr(''.join(idx2char[item.numpy()])))
```
对于每个序列,使用 `map` 方法先复制再顺移,以创建输入文本和目标文本。`map` 方法可以将一个简单的函数应用到每一个批次 (batch)。
```
def split_input_target(chunk):
input_text = chunk[:-1]
target_text = chunk[1:]
return input_text, target_text
dataset = sequences.map(split_input_target)
```
打印第一批样本的输入与目标值:
```
for input_example, target_example in dataset.take(1):
print ('Input data: ', repr(''.join(idx2char[input_example.numpy()])))
print ('Target data:', repr(''.join(idx2char[target_example.numpy()])))
```
这些向量的每个索引均作为一个时间步来处理。作为时间步 0 的输入,模型接收到 “F” 的索引,并尝试预测 “i” 的索引为下一个字符。在下一个时间步,模型执行相同的操作,但是 `RNN` 不仅考虑当前的输入字符,还会考虑上一步的信息。
```
for i, (input_idx, target_idx) in enumerate(zip(input_example[:5], target_example[:5])):
print("Step {:4d}".format(i))
print(" input: {} ({:s})".format(input_idx, repr(idx2char[input_idx])))
print(" expected output: {} ({:s})".format(target_idx, repr(idx2char[target_idx])))
```
### 创建训练批次
前面我们使用 `tf.data` 将文本拆分为可管理的序列。但是在把这些数据输送至模型之前,我们需要将数据重新排列 (shuffle) 并打包为批次。
```
# 批大小
BATCH_SIZE = 64
# 设定缓冲区大小,以重新排列数据集
# (TF 数据被设计为可以处理可能是无限的序列,
# 所以它不会试图在内存中重新排列整个序列。相反,
# 它维持一个缓冲区,在缓冲区重新排列元素。)
BUFFER_SIZE = 10000
dataset = dataset.shuffle(BUFFER_SIZE).batch(BATCH_SIZE, drop_remainder=True)
dataset
```
## 创建模型
使用 `tf.keras.Sequential` 定义模型。在这个简单的例子中,我们使用了三个层来定义模型:
* `tf.keras.layers.Embedding`:输入层。一个可训练的对照表,它会将每个字符的数字映射到一个 `embedding_dim` 维度的向量。
* `tf.keras.layers.GRU`:一种 RNN 类型,其大小由 `units=rnn_units` 指定(这里你也可以使用一个 LSTM 层)。
* `tf.keras.layers.Dense`:输出层,带有 `vocab_size` 个输出。
```
# 词集的长度
vocab_size = len(vocab)
# 嵌入的维度
embedding_dim = 256
# RNN 的单元数量
rnn_units = 1024
def build_model(vocab_size, embedding_dim, rnn_units, batch_size):
model = tf.keras.Sequential([
tf.keras.layers.Embedding(vocab_size, embedding_dim,
batch_input_shape=[batch_size, None]),
tf.keras.layers.GRU(rnn_units,
return_sequences=True,
stateful=True,
recurrent_initializer='glorot_uniform'),
tf.keras.layers.Dense(vocab_size)
])
return model
model = build_model(
vocab_size = len(vocab),
embedding_dim=embedding_dim,
rnn_units=rnn_units,
batch_size=BATCH_SIZE)
```
对于每个字符,模型会查找嵌入,把嵌入当作输入运行 GRU 一个时间步,并用密集层生成逻辑回归 (logits),预测下一个字符的对数可能性。
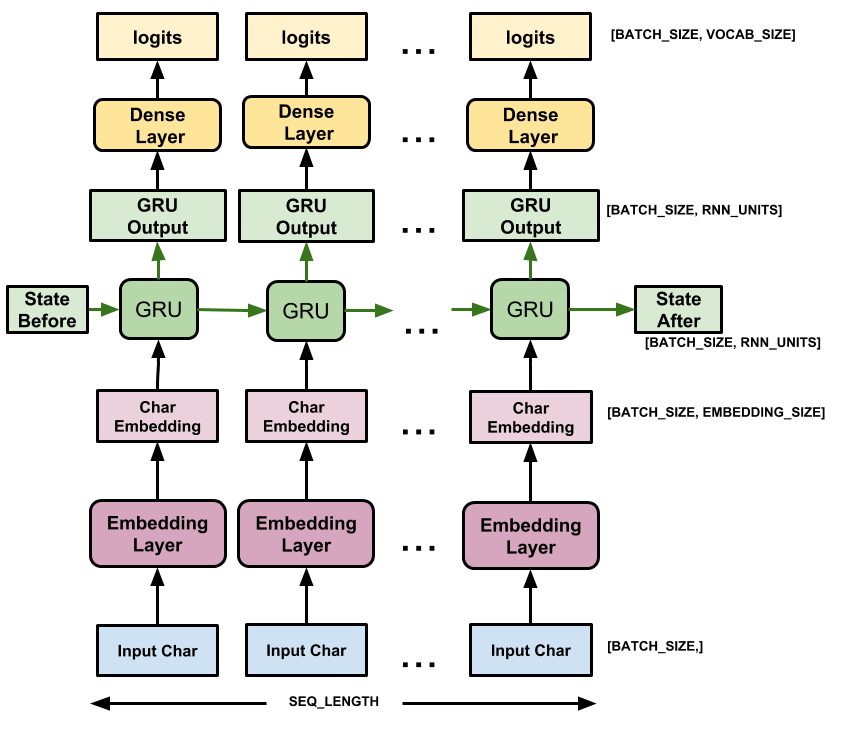
## 试试这个模型
现在运行这个模型,看看它是否按预期运行。
首先检查输出的形状:
```
for input_example_batch, target_example_batch in dataset.take(1):
example_batch_predictions = model(input_example_batch)
print(example_batch_predictions.shape, "# (batch_size, sequence_length, vocab_size)")
```
在上面的例子中,输入的序列长度为 `100`, 但是这个模型可以在任何长度的输入上运行:
```
model.summary()
```
为了获得模型的实际预测,我们需要从输出分布中抽样,以获得实际的字符索引。这个分布是根据对字符集的逻辑回归定义的。
请注意:从这个分布中 _抽样_ 很重要,因为取分布的 _最大值自变量点集(argmax)_ 很容易使模型卡在循环中。
试试这个批次中的第一个样本:
```
sampled_indices = tf.random.categorical(example_batch_predictions[0], num_samples=1)
sampled_indices = tf.squeeze(sampled_indices,axis=-1).numpy()
```
这使我们得到每个时间步预测的下一个字符的索引。
```
sampled_indices
```
解码它们,以查看此未经训练的模型预测的文本:
```
print("Input: \n", repr("".join(idx2char[input_example_batch[0]])))
print()
print("Next Char Predictions: \n", repr("".join(idx2char[sampled_indices ])))
```
## 训练模型
此时,这个问题可以被视为一个标准的分类问题:给定先前的 RNN 状态和这一时间步的输入,预测下一个字符的类别。
### 添加优化器和损失函数
标准的 `tf.keras.losses.sparse_categorical_crossentropy` 损失函数在这里适用,因为它被应用于预测的最后一个维度。
因为我们的模型返回逻辑回归,所以我们需要设定命令行参数 `from_logits`。
```
def loss(labels, logits):
return tf.keras.losses.sparse_categorical_crossentropy(labels, logits, from_logits=True)
example_batch_loss = loss(target_example_batch, example_batch_predictions)
print("Prediction shape: ", example_batch_predictions.shape, " # (batch_size, sequence_length, vocab_size)")
print("scalar_loss: ", example_batch_loss.numpy().mean())
```
使用 `tf.keras.Model.compile` 方法配置训练步骤。我们将使用 `tf.keras.optimizers.Adam` 并采用默认参数,以及损失函数。
```
model.compile(optimizer='adam', loss=loss)
```
### 配置检查点
使用 `tf.keras.callbacks.ModelCheckpoint` 来确保训练过程中保存检查点。
```
# 检查点保存至的目录
checkpoint_dir = './training_checkpoints'
# 检查点的文件名
checkpoint_prefix = os.path.join(checkpoint_dir, "ckpt_{epoch}")
checkpoint_callback=tf.keras.callbacks.ModelCheckpoint(
filepath=checkpoint_prefix,
save_weights_only=True)
```
### 执行训练
为保持训练时间合理,使用 10 个周期来训练模型。在 Colab 中,将运行时设置为 GPU 以加速训练。
```
EPOCHS=10
history = model.fit(dataset, epochs=EPOCHS, callbacks=[checkpoint_callback])
```
## 生成文本
### 恢复最新的检查点
为保持此次预测步骤简单,将批大小设定为 1。
由于 RNN 状态从时间步传递到时间步的方式,模型建立好之后只接受固定的批大小。
若要使用不同的 `batch_size` 来运行模型,我们需要重建模型并从检查点中恢复权重。
```
tf.train.latest_checkpoint(checkpoint_dir)
model = build_model(vocab_size, embedding_dim, rnn_units, batch_size=1)
model.load_weights(tf.train.latest_checkpoint(checkpoint_dir))
model.build(tf.TensorShape([1, None]))
model.summary()
```
### 预测循环
下面的代码块生成文本:
* 首先设置起始字符串,初始化 RNN 状态并设置要生成的字符个数。
* 用起始字符串和 RNN 状态,获取下一个字符的预测分布。
* 然后,用分类分布计算预测字符的索引。把这个预测字符当作模型的下一个输入。
* 模型返回的 RNN 状态被输送回模型。现在,模型有更多上下文可以学习,而非只有一个字符。在预测出下一个字符后,更改过的 RNN 状态被再次输送回模型。模型就是这样,通过不断从前面预测的字符获得更多上下文,进行学习。
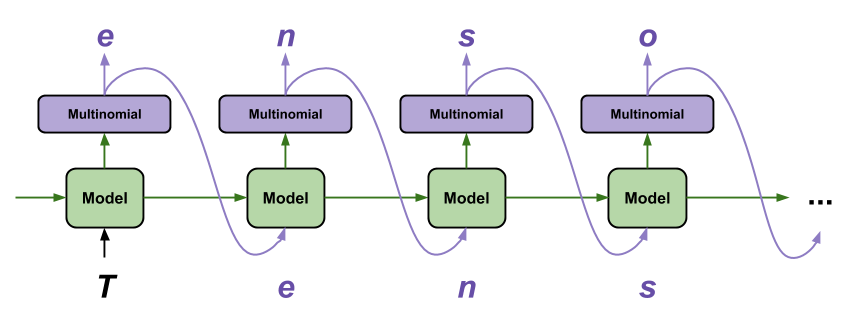
查看生成的文本,你会发现这个模型知道什么时候使用大写字母,什么时候分段,而且模仿出了莎士比亚式的词汇。由于训练的周期小,模型尚未学会生成连贯的句子。
```
def generate_text(model, start_string):
# 评估步骤(用学习过的模型生成文本)
# 要生成的字符个数
num_generate = 1000
# 将起始字符串转换为数字(向量化)
input_eval = [char2idx[s] for s in start_string]
input_eval = tf.expand_dims(input_eval, 0)
# 空字符串用于存储结果
text_generated = []
# 低温度会生成更可预测的文本
# 较高温度会生成更令人惊讶的文本
# 可以通过试验以找到最好的设定
temperature = 1.0
# 这里批大小为 1
model.reset_states()
for i in range(num_generate):
predictions = model(input_eval)
# 删除批次的维度
predictions = tf.squeeze(predictions, 0)
# 用分类分布预测模型返回的字符
predictions = predictions / temperature
predicted_id = tf.random.categorical(predictions, num_samples=1)[-1,0].numpy()
# 把预测字符和前面的隐藏状态一起传递给模型作为下一个输入
input_eval = tf.expand_dims([predicted_id], 0)
text_generated.append(idx2char[predicted_id])
return (start_string + ''.join(text_generated))
print(generate_text(model, start_string=u"ROMEO: "))
```
若想改进结果,最简单的方式是延长训练时间 (试试 `EPOCHS=30`)。
你还可以试验使用不同的起始字符串,或者尝试增加另一个 RNN 层以提高模型的准确率,亦或调整温度参数以生成更多或者更少的随机预测。
## 高级:自定义训练
上面的训练步骤简单,但是能控制的地方不多。
至此,你已经知道如何手动运行模型。现在,让我们打开训练循环,并自己实现它。这是一些任务的起点,例如实现 _课程学习_ 以帮助稳定模型的开环输出。
你将使用 `tf.GradientTape` 跟踪梯度。关于此方法的更多信息请参阅 [eager execution 指南](https://tensorflow.google.cn/guide/eager)。
步骤如下:
* 首先,初始化 RNN 状态,使用 `tf.keras.Model.reset_states` 方法。
* 然后,迭代数据集(逐批次)并计算每次迭代对应的 *预测*。
* 打开一个 `tf.GradientTape` 并计算该上下文时的预测和损失。
* 使用 `tf.GradientTape.grads` 方法,计算当前模型变量情况下的损失梯度。
* 最后,使用优化器的 `tf.train.Optimizer.apply_gradients` 方法向下迈出一步。
```
model = build_model(
vocab_size = len(vocab),
embedding_dim=embedding_dim,
rnn_units=rnn_units,
batch_size=BATCH_SIZE)
optimizer = tf.keras.optimizers.Adam()
@tf.function
def train_step(inp, target):
with tf.GradientTape() as tape:
predictions = model(inp)
loss = tf.reduce_mean(
tf.keras.losses.sparse_categorical_crossentropy(
target, predictions, from_logits=True))
grads = tape.gradient(loss, model.trainable_variables)
optimizer.apply_gradients(zip(grads, model.trainable_variables))
return loss
# 训练步骤
EPOCHS = 10
for epoch in range(EPOCHS):
start = time.time()
# 在每个训练周期开始时,初始化隐藏状态
# 隐藏状态最初为 None
hidden = model.reset_states()
for (batch_n, (inp, target)) in enumerate(dataset):
loss = train_step(inp, target)
if batch_n % 100 == 0:
template = 'Epoch {} Batch {} Loss {}'
print(template.format(epoch+1, batch_n, loss))
# 每 5 个训练周期,保存(检查点)1 次模型
if (epoch + 1) % 5 == 0:
model.save_weights(checkpoint_prefix.format(epoch=epoch))
print ('Epoch {} Loss {:.4f}'.format(epoch+1, loss))
print ('Time taken for 1 epoch {} sec\n'.format(time.time() - start))
model.save_weights(checkpoint_prefix.format(epoch=epoch))
```
| github_jupyter |
```
import json
import os
import sys
import fnmatch
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from collections import defaultdict
from sklearn import neural_network
from sklearn import linear_model
from sklearn import preprocessing
from sklearn import feature_selection, model_selection
from sklearn import multioutput
from sklearn import ensemble
from sklearn import svm
df = pd.read_csv('new_big_fbgm.csv')
#df = pd.read_csv('FBGM_League_1_all_seasons_Average_Stats(1).csv')
#df = df.sample(10000)
#['QB', 'OL', 'TE', 'DL', 'S', 'CB', 'WR', 'LB', 'RB', 'P', 'K']
clf2 = None
unique_pos = df.Pos.unique()
df.shape
#orig_df = df.copy()
#df = orig_df.sample(frac=0.1)
df.shape
stats_to_pred = (df.columns[105:])
stats_to_use = list(df.columns[11:103])
need_norm = ['Cmp', 'Att', 'Yds', 'TD', 'Int', 'Sk', 'Yds.1','Rush', 'Yds.2', 'TD.1','Tgt', 'Rec', 'Yds.3', 'TD.2', 'Tch','RRTD',
'Int.1',
'Yds.4',
'TD.3',
'PD',
'FF',
'FR',
'Yds.5',
'TD.4',
'Sk.1',
'Tck',
'Solo',
'Ast',
'TFL',
'Sfty',
'Fmb',
'FG10',
'FGA10',
'FG20',
'FGA20',
'FG30',
'FGA30',
'FG40',
'FGA40',
'FG50',
'FGA50',
'FGM',
'FGA',
'XPM',
'XPA',
'Pnt',
'Yds.7',
'PR',
'Yds.8',
'TD.5',
'KR',
'Yds.9',
'TD.6',
'FL',
'Pen',
'Yds.6',
'Blk',
'YScm',
'APY',
'AV']
need_norm = list(set(need_norm))
BAD_SET = set(['Blk','FL', 'Pen', 'Yds.6','Lng.4', 'APY'])
need_norm = [_ for _ in need_norm if _ not in BAD_SET]
for col in need_norm:
df[col + '_PerG'] = df[col]/np.maximum(df['G'],1)
stats_to_use.append(col + '_PerG')
for col in need_norm:
del stats_to_use[stats_to_use.index(col)]
stats_to_use = [_ for _ in stats_to_use if _ not in BAD_SET]
pos_t = 'QB'
stat_t = 'ThP'
stat_t_idx = list(y.columns).index(stat_t)
replacement_player_mean_bs = {}
replacement_player_std_bs = {}
replacement_player_cov_bs = {}
replacement_player_mean_r = {}
replacement_player_std_r = {}
replacement_player_cov_r = {}
clfs = {}
scalersX = {}
scalersY = {}
fexps = {}
for pos in unique_pos:
y = df[df.Pos == pos]#.iloc[:,105:]
X = df[df.Pos == pos]#.iloc[:,11:103]
y = y[stats_to_pred]
valid_col = stats_to_use#[_ for _ in X.columns if not _ in ['Blk','FL', 'Pen', 'Yds.6','Lng.4', 'APY']]
X = X[valid_col]
replacement_filter = (df.Pos == pos) & (df.Salary >= 0.10) & (df.Salary < 1.0)
replacement_player_mean_bs[pos] = X[replacement_filter].mean()
replacement_player_std_bs[pos] = X[replacement_filter].std()
replacement_player_cov_bs[pos] = X[replacement_filter].cov()
replacement_player_mean_r[pos] = y[replacement_filter].mean()
replacement_player_std_r[pos] = y[replacement_filter].std()
replacement_player_cov_r[pos] = y[replacement_filter].cov()
fexp = preprocessing.PolynomialFeatures(degree=1,interaction_only=True)
scalerX = preprocessing.StandardScaler()
scalery = preprocessing.StandardScaler()
prescale_X = scalerX.fit_transform(X)
prescale_y = scalery.fit_transform(y)
prescale_X = fexp.fit_transform(prescale_X)
X_train, X_test, y_train, y_test = model_selection.train_test_split(prescale_X, prescale_y, test_size=0.25, random_state=42)
clf = multioutput.MultiOutputRegressor(linear_model.ElasticNet(alpha=5e-2))
clf.fit(prescale_X,prescale_y)
yt = scalery.inverse_transform(clf.predict(prescale_X))
print(pos,clf.score(prescale_X,prescale_y))
clfs[pos] = clf
scalersX[pos] = scalersX
scalersY[pos] = scalery
fexps[pos] = fexp
if pos == pos_t:
fscores = sklearn.feature_selection.f_regression(prescale_X,prescale_y[:,stat_t_idx])[1]
for i in np.argsort(fscores):
if np.isnan(fscores[i]) or fscores[i] > 0.1:
continue
print(X.columns[i],fscores[i])
def get_norms(X):
means = np.zeros(X.shape[1])
stds = np.zeros(X.shape[1])
for col in range(X.shape[1]):
xv = np.array(X)[:,col]
#print(xv[np.where(xv !=0)].shape,X.columns[col])
means[col] = xv[np.where(xv !=0)].mean()
stds[col] = xv[np.where(xv !=0)].std()
rX = (np.array(X)-means)/stds
rX[np.where(X == 0)] = -10
return rX,means,stds
col_names = X.columns
col_names = fexp.get_feature_names(X.columns)
pos = 'QB'
stat_t = 'ThV'
stat_t_idx = list(y.columns).index(stat_t)
for i,c in enumerate(y.columns):
coeffs = clfs[pos].estimators_[i].coef_
v = np.argsort(abs(coeffs))[::-1]
print(c)
coeffs2 = [(coeffs[i2],col_names[i2]) for i2 in v[:10]]
#for v,n in sorted(coeffs2,reverse=True):
# print('{:.2f} * {} + '.format(v,n),end='')
print('| Variable | Coeff |')
print('|----------|-------|')
for v,n in sorted(coeffs2,reverse=True):
print('|{:25s}|{:.5f}|'.format(n,v))
#for v,n in sorted(coeffs2,reverse=True):
# print('\t{:25s}\t{:.2f}'.format(n,v))
GEN_YEAR = 2019
tyear = [GEN_YEAR]
#if tyear[0] < 1980:
# print("MY PARSING OF THE TABLES IS WRONG WITHOUT the 2PA/3PA TRACKS")
# raise
#CURRENT_YEAR = 2019
all_tables = {}
for ty in tyear:
all_tables[ty] = np.load('fb_tables_{}.pkl'.format(ty))
teams = all_tables[tyear[0]].keys()
print(all_tables[GEN_YEAR]['nwe'].keys())
import sklearn.feature_selection
player_stats = {k:{} for k in tyear}
table_columns = defaultdict(dict)
for ty in tyear:
tables = all_tables[ty]
for team in tables:
team_tables = tables[team]
for table_name in team_tables:
if table_name in ['draft-rights','team_and_opponent','conf','name','logo']:
continue
table = team_tables[table_name].fillna(0)
#print(table_name)
#print(table.index)
for row in table.itertuples():
name = row[0]
name = name.replace('\xa0\xa0',' ').replace('*','').replace('+','')
if name == 'Team Totals':
continue
nsplit = name.split(' ')
if nsplit[-1] in ['Jr.','Sr.','I','II','III',"IV",'(TW)']:
name = ' '.join(nsplit[:-1])
player_table = player_stats[ty].get(name,{})
player_row = player_table.get(table_name,[])
player_row = player_row + [row]
player_table[table_name] = player_row
player_table['team'] = team
player_stats[ty][name] = player_table
#if name == 'Dennis Smith Jr.' or name == 'Luka Doncic':
# print(player_stats[ty][name],team)
tcs = list(table.columns)
for i in range(len(tcs)):
while tcs.index(tcs[i]) != i:
if '.' in tcs[i]:
tsplit = tcs[i].split('.')
tsplit[-1] = str(int(tsplit[-1]) + 1)
tcs[i] = '.'.join(tsplit)
else:
tcs[i] += '.1'
table_columns[(ty,team)][table_name] = tcs
#player_stats[2007].keys()
table_mask = defaultdict(dict)
for key in table_columns:
for table in table_columns[key]:
table_mask[key][table] = [_.strip() !='' for _ in table_columns[key][table] ]
table_columns[key][table] = [_ for _ in table_columns[key][table] if _.strip() != '']
#for player in player_stats:
# for table_in in player_stats[player]:
# if 'on_off' in table_in or 'salaries' in table_in:
# continue
# if len(player_stats[player][table_in]) > 1:
# pass
#print(player,table_in,'MP' in player_stats[player][table_in][0]._fields)
#print(player_stats[player][table_in][0])
# add playoff data to normal data
if False:
for ty in tyear:
for player in player_stats[ty]:
for table_in in player_stats[ty][player]:
tableN = table_in.split('_')
tableS = '_'.join(tableN[1:])
if 'playoffs'==tableN[0] and not table_in in ['playoffs_pbp']:
#print(table_in)
if tableS in player_stats[ty][player]:
player_stats[ty][player][tableS] += player_stats[ty][player][table_in]
#player_stats[1968]['Bill Allen']
#player_stats[1968]['Jim Burns']['per_game'], table_columns[(1968,'DLC')]['per_game'].index('MP')
for ty in tyear:
for player in player_stats[ty]:
team = player_stats[ty][player]['team']
for tt in player_stats[ty][player]:
if tt == 'team':
continue
new_rows = []
for tablet in player_stats[ty][player][tt]:
vector = [_ if _ != '' else '0.0' for _ in tablet[1:]]
vector = [(float(_.replace('%',''))/100 if type(_) == str and'%' in _ else _) for _ in vector]
if 'on_off' in tt:
vector = vector[1:]
if 'contracts' in tt:
vector = vector[1:-2]
if tt in ['salaries2','contracts']:
vector = [_.replace(',','').replace('$','') for _ in vector]
try:
v2 = np.array(vector).astype(np.float)
except:
v2 = vector
new_rows.append(vector)
a = np.array(new_rows)
if tt in table_columns[(ty,team)]:
if 'MP' in table_columns[(ty,team)][tt] and not tt in ['pbp','on_off','on_off_p']:
try:
a = a.astype(np.float)
except:
a = list(a)
a[0] = np.array([float(_) for _ in a[0]])
a[1] = np.array([float(_) for _ in a[1]])
a = np.array(a)
try:
mins = a[:,table_columns[(ty,team)][tt].index('MP')].reshape((-1,1))
new_rows = ((a.T @ mins)/mins.sum()).T
a = new_rows
except:
#print(tt,a.shape,player,a,mins)
#print('.',end='')
print(ty,team,player,tt)
continue
player_stats[ty][player][tt] = a
player_stats
len(X.columns)
X.columns
#'returns', 'kicking'
#table_columns[(2018,'nwe')]['kicking']
lt = [('AV','games_played_team'),
('Cmp','passing'),('Att','passing'),('Pct','passing','Cmp%'),
('Yds','passing'),('TD','passing'),('TD%','passing'),('Int','passing'),
('Int%','passing'),('Lng','passing'),('Y/A','passing'),('AY/A','passing'),('QBRat','passing','Rate'),
('Y/C','passing'),('Y/G','passing'),('Sk','passing'),('Yds.1','passing'),('NY/A','passing'),
('ANY/A','passing'),('Sk%','passing'),('Rush','rushing_and_receiving','Att'),
('Yds.2','rushing_and_receiving','Yds'),('TD.1','rushing_and_receiving','TD'),('Lng.1','rushing_and_receiving','Lng'),
('Y/A.1','rushing_and_receiving','Y/A'),('Y/G.1','rushing_and_receiving','Y/G'),('A/G','rushing_and_receiving'),
('Tgt','rushing_and_receiving'),('Rec','rushing_and_receiving'),('Yds.3','rushing_and_receiving','Yds.1'),
('TD.2','rushing_and_receiving','TD.1'),('Lng.2','rushing_and_receiving','Lng.1'),('Y/R','rushing_and_receiving'),
('R/G','rushing_and_receiving'),('Y/G.2','rushing_and_receiving','Y/G.1'),('Ctch%','rushing_and_receiving'),
('Y/Tch','rushing_and_receiving'),('Y/Tgt','rushing_and_receiving'),('Tch','rushing_and_receiving','Touch'),
('YScm','rushing_and_receiving'),('RRTD','rushing_and_receiving'),('Int.1','defense','Int'),
('Yds.4','defense','Yds'),('TD.3','defense','TD'),('Lng.3','defense','Lng'),('PD','defense'),
('FF','defense'),('FR','defense'),('Yds.5','defense','Yds.1'),('TD.4','defense','TD.1'),('Sk.1','defense','Sk'),
('Tck','defense','Comb'),('Solo','defense'),('Ast','defense'),('TFL','defense'),('Sfty','defense'),('Fmb','defense'),
('FG10','kicking','FGM'),('FGA10','kicking','FGA'),('FG20','kicking','FGM.1'),('FGA20','kicking','FGA.1'),
('FG30','kicking','FGM.2'),('FGA30','kicking','FGA.2'),('FG40','kicking','FGM.3'),('FGA40','kicking','FGA.3'),
('FG50','kicking','FGM.4'),('FGA50','kicking','FGA.4'),('FGM','kicking','FGM.5'),('FGA','kicking','FGA.5'),
('Pct.1','kicking','FG%'),('XPM','kicking'),('XPA','kicking'),('Pct.2','kicking','XP%'),('Pnt','kicking'),
('Yds.7','kicking','Yds'),('Lng.5','kicking','Lng'),('Blk','kicking','Blck'),('Y/A.2','kicking','Y/P'),
('PR','returns','Ret'),('Yds.8','returns','Yds'),('TD.5','returns','TD'),('Lng.6','returns','Lng'),
('Y/A.3','returns','Y/R'),('KR','returns','Rt'),('Yds.9','returns','Yds.1'),('TD.6','returns','TD.1'),
('Lng.7','returns','Lng.1'),('Y/A.4','returns','Y/Rt')]
locations = []
for loc in lt:
if len(loc) == 2:
loc = list(loc)
loc.append(loc[0])
locations.append(loc)
locs = defaultdict(dict)
for ty in tyear:
for team in all_tables[ty]:
for loc in locations:
try:
locs[(ty,team)][loc[0]] = (table_columns[(ty,team)][loc[1]].index(loc[2]),loc[1])
except Exception as e:
#print(e)
#print(team,ty)
print(e,loc)
pass#print(loc,table_columns[loc[1]])
#if 'shooting' in table_columns:
# l = list(locs['LowPostFGP'])
# l[0] +=2
# locs['MidRangeFGP'] = l
len(locs[(GEN_YEAR,'nwe')]),locs[(GEN_YEAR,'nwe')]
player_vectors = []
player_names = []
player_years = []
player_scales = []
player_heights = []
player_weights = []
player_drafts = []
player_ages = []
player_teams = []
player_pos = []
for ty in tyear:
for name in player_stats[ty]:
try:
stats = player_stats[ty][name]
team = stats['team']
d = {}
for k,v in locs[(ty,team)].items():
SV = 0
try:
SV = float(stats[v[1]][0][v[0]])
except:
pass
d[k] = 0 if SV == '' else SV
d['Pos'] = []
for t in ['passing','defense','returns','kicking','rushing_and_receiving','games_played_team']:
try:
d['Age'] = int(float(stats[t][0][table_columns[(ty,team)][t].index('Age')]))
except:
pass
try:
d['Pos'].append(stats[t][0][table_columns[(ty,team)][t].index('Pos')])
except:
pass
try:
d['G'] = max(int(stats[t][0][table_columns[(ty,team)][t].index('G')]),d.get('G',0))
#print(d['G'])
except:
pass
if 'games_played_team' not in stats:
continue
wt = stats['games_played_team'][0][4]
ht = stats['games_played_team'][0][5]
exp = stats['games_played_team'][0][8]
for col in need_norm:
if col in d:
d[col + '_PerG'] = d[col]/np.maximum(d.get('G',0),1)
player_vectors.append([d[stat] for stat in X.columns])
player_names.append(name)
player_years.append(ty)
player_ages.append(d['Age'])
player_pos.append(d['Pos'])
player_heights.append(ht)
player_drafts.append(exp)
player_weights.append(wt)
player_teams.append(team)
except Exception as e:
print(name,team)
print(e)
print(d.keys())
#if name == 'Bill Bradley':
# raise
raise
X.columns
player_stats[ty][name]
first_n = len([yr for yr in player_years if yr == tyear[0]])
gen_FA = 0#len(teams)*5 if CURRENT_YEAR!=GEN_YEAR else 0
first_n,len(teams),gen_FA
#player_names
Xn = np.nan_to_num(np.array(player_vectors))
#Xn.shape,Xn_s.shape,prescale_X.shape,fexp.n_input_features_,X.shape
#clf2 = None
if clf2 is None:
clf2 = linear_model.LogisticRegression()
scalerX_pos_og = preprocessing.StandardScaler()
fexp_pos = preprocessing.PolynomialFeatures(degree=1,interaction_only=False)
#tmpX = df.iloc[:,11:103]
tmpX = df[valid_col]
clf2.fit(fexp_pos.fit_transform(scalerX_pos_og.fit_transform(tmpX)),df.Pos)
scalerX_pos = preprocessing.StandardScaler()
Xn_scale = fexp_pos.transform(scalerX_pos.fit_transform(Xn))
pred_pos = clf2.predict(Xn_scale)
#print(clf2.score(Xn_scale,df.Pos))
'QB', 'OL', 'TE', 'DL', 'S', 'CB', 'WR', 'LB', 'RB', 'P', 'K'
pos_mapping = {
'LG':'OL',
'RG':'OL',
'CB':'CB',
'LOLB':'LB',
'RB':'RB',
'FB':'RB',
'K':'K',
'C':'OL',
'LCB':'CB',
'RCB':'CB',
'FS':'S',
'RDT':'DL',
'QB':'QB',
'DL':'DL',
'MLB':'LB',
'WR':'WR',
'S':'S',
'SS':'S',
'TE':'TE',
'DT':'DL',
'RILB':'LB',
'T':'OL',
'LDT':'DL',
'P':'P',
'LILB':'LB',
'ROLB':'LB',
'LB':'LB',
'DB':'S',
'DE':'DL',
'NT':'DL',
'RT':'OL',
'LT': 'OL',
'SE': 'WR',
'FL': 'WR',
'OLB':'LB',
'DE ': 'DL',
'OT': 'OL',
'OL': 'OL',
'LS': 'OL',
'ILB':'LB',
'G':'OL',
'OG':'OL',
'EDGE':'DL',
'G,T': 'OL',
'LS,TE': 'TE',
'PR':'WR',
'KR':'WR',
'RDE':'DL',
'LDE':'DL',
'RLB':'LB',
'LLB':'LB'
}
import random
predicted_pos = []
for listed_p,model_p in zip(player_pos,pred_pos):
possible_pos = [_.upper() for _ in listed_p if _ != '0.0']
pposs = []
for pos in list(possible_pos):
if '/' in pos:
pp = pos.split('/')
for ppp in pp:
pposs.append(ppp)
else:
pposs.append(pos)
possible_pos = [pos_mapping[_.replace(' ','')] for _ in set(pposs)]
#print(possible_pos,model_p)
if model_p in possible_pos:
true_p = model_p
elif len(possible_pos) > 0:
random.shuffle(possible_pos)
true_p = possible_pos[0]
else:
true_p = model_p
predicted_pos.append(true_p)
predicted_pos = np.array(predicted_pos)
ratings = np.zeros((predicted_pos.shape[0],y.shape[1]))
for pos in unique_pos:
Xn = np.nan_to_num(np.array(player_vectors))[np.where(predicted_pos == pos)]
# tuned this to get roughly 8-12 players at 70 or above. Which seemed like normal for a league
#scalerX2 = preprocessing.StandardScaler()#(quantile_range=(30.0, 70.0))
scalerX2 = preprocessing.StandardScaler()
Xn_s =scalerX2.fit_transform(np.nan_to_num(Xn))
Xn_fs = fexps[pos].transform(np.nan_to_num(Xn_s))
predict = clfs[pos].predict(Xn_fs)
est_ratings = np.nan_to_num(scalersY[pos].inverse_transform(predict))
ratings[np.where(predicted_pos == pos)] = est_ratings
#all_tables[2018]['nwe'].keys()
#'logo', 'name', 'conf'
teams2 = []
for t in teams:
try:
aaaa = all_tables[GEN_YEAR][t]['conf'][:-9]
teams2.append(t)
except:
pass
teams = teams2
confs = ['AFC','NFC']
divs = list(set([all_tables[GEN_YEAR][t]['conf'][:-9] for t in teams]))
divs
import random
base = {}
base['startingSeason'] = GEN_YEAR
valid_tids = list(range(len(teams)))
new_teams = []
for i,t1 in enumerate(sorted(list(teams))):
t = {}
t['abbrev'] = t1.upper()
nsplit = all_tables[GEN_YEAR][t1]['name'].split(' ')
t['region'] = ' '.join(nsplit[:-1])
t['name'] = nsplit[-1]
t['imgURL'] = all_tables[GEN_YEAR][t1]['logo']
t['tid'] = i
div_name = all_tables[GEN_YEAR][t1]['conf'].rstrip().split(' ')
t['did'] = divs.index(' '.join(div_name[:-1]))
t['cid'] = [c in all_tables[GEN_YEAR][t1]['conf'] for c in confs].index(True)
new_teams.append(t)
print(new_teams)
team_abbbrevs = list(teams)
base['teams'] = new_teams
base['gameAttributes'] = []
base['version'] = 33
DIVS = []
for div in divs:
cid = [c in div for c in confs].index(True)
did = divs.index(div)
DIVS.append({'did':did, 'cid':cid, 'name':div})
print(DIVS)
base['gameAttributes'].append({'key':'divs', 'value': DIVS})
base['gameAttributes'].append({'key':'confs', 'value': [{'cid':i, 'name':n } for i,n in enumerate(confs)]})
confs
if gen_FA > 0:
player_names_f = player_names + ["Free Agent{}".format(write_roman(i)) for i in range(gen_FA)]
player_years_f = player_years + [GEN_YEAR for i in range(gen_FA)]
# scale them down, we don't want a bunch of amazing replacements
MEAN_S = 0.95
STD_S = 0.25
rp_ratings = np.random.multivariate_normal(MEAN_S*replacement_player_mean_r,STD_S*replacement_player_cov_r,size=(gen_FA))
ratings_f = np.vstack([ratings,rp_ratings])
else:
player_names_f = player_names
player_years_f = player_years
ratings_f = ratings
l2 = [_.lower() for _ in y.columns]
l2[1] = 'stre'
l2[3] = 'endu'
l2[-7] = 'tck'
import pprint
import copy
import random
gen_player = []
pp = pprint.PrettyPrinter()
for i,name in enumerate(player_names_f):
py = player_years_f[i]
name = player_names_f[i].replace('*','').replace('+','')
if name in ['Team Total','Opp Total']:
continue
sname = name.split(' ')
new_player = {}
new_player['firstName'] = sname[0]
new_player['lastName'] = ' '.join(sname[1:])
r_vec = {k: ratings_f[i][i2] for i2,k in enumerate(l2)}
r_vec = {k: int(np.clip(v,0,100)) for k,v in r_vec.items()}
#for t in ['hgt', 'stre', 'spd', 'jmp', 'endu', 'ins', 'dnk', 'ft', 'fg', 'tp', 'diq', 'oiq', 'drb', 'pss', 'reb']:
# if not t in r_vec:
# r_vec[t] = 50
true_pos = [predicted_pos[i]]
r_vec['pos'] = true_pos[0]
new_player['pos'] =true_pos[0]
#print(set(true_pos))
new_player['ratings'] = [r_vec]
age = player_ages[i]
new_player['born'] = {'year':GEN_YEAR-age,'loc':''}
new_player['tid'] = team_abbbrevs.index(player_teams[i])
exp = player_drafts[i]
try:
exp = int(exp)
except:
exp = 0
new_player['draft'] = {"originalTid": -1, "pick": 0, "round": 0, "tid": -1, "year": GEN_YEAR-exp}
new_player['weight'] = int(float(player_weights[i]))
try:
ht = [int(_) for _ in player_heights[i].split('-')]
hgt = ht[0]*12 + ht[1]
#print(ht,hgt)
new_player['hgt'] = int(hgt)#int(3.7*(hgt-66))
except:
pass
gen_player.append(new_player)
#gen_player
sum(['pos' in _['ratings'][0] for _ in gen_player]),len(gen_player)
base['players'] = gen_player
with open('fbgm_roster_{}.json'.format(tyear[0]),'wt') as fp:
json.dump(base,fp, sort_keys=True)
len(ratings),len(ratings_f),len(ratings)+gen_FA
gen_player
sorted(['S','DL'])
ppos = [_.upper() for _ in set(sum(player_pos,[])) if _ != '0.0']
pPos = []
for p in list(ppos):
if '/' in p:
for pp in p.split('/'):
pPos.append(pp)
else:
pPos.append(p)
pPos = list(set(pPos))
df.Pos.unique()
```
| github_jupyter |
```
# default_exp core
#hide
!pip install nbdev
```
# 00_Core
> This notebook will include the core functionalities needed to make our library operational within Google Colab
```
#hide
from nbdev.showdoc import *
```
As we are working out of our `Drive`, let's write a function to mount and refresh it each time (you will only need to sign in on the first)
```
#export
import os
from google.colab import drive
#export
def setup_drive():
"Connect Google Drive to use GitHub"
drive.mount('/content/drive', force_remount=True)
os._exit(00)
setup_drive()
```
Clone your repo into your Google Drive then re-open it (this only needs to be done once) so we are working in it's codebase
Now let's setup our instance to be utilized by Git and accepted
```
#export
from pathlib import Path
import os, subprocess
#export
def setup_git(path:Path, project_name:str, username:str, password:str, email:str):
"Link your mounted drive to GitHub. Remove sensitive information before pushing"
start = os.getcwd()
os.chdir(path)
commands = []
commands.append(f"git config --global user.email {email}")
commands.append(f"git config --global user.name {username}")
commands.append("git init")
commands.append("git remote rm origin")
commands.append(f"git remote add origin https://{username}:{password}@github.com/{username}/{project_name}.git")
commands.append("git pull origin master --allow-unrelated-histories")
for cmd in commands:
process = subprocess.Popen(cmd.split(), stdout=subprocess.PIPE)
output, err = process.communicate()
os.chdir(start)
```
We need to pass in the `Path` to point to our cloned repository along with the needed information. **REMEMBER to delete or replace the sensitive information BEFORE uploading to your library**
```
git_path = Path('drive/My Drive/nbdev_colab')
setup_git(git_path, 'nbdev_colab', 'muellerzr', 'myPass', 'myEmail@gmail.com')
```
Now let's make a command to push to our repository, similar to `setup_git`. This will also make our library for us
```
#export
from nbdev.export import *
#export
def git_push(path:Path, message:str):
"Convert the notebooks to scripts and then push to the library"
start = os.getcwd()
os.chdir(path)
commands = []
commands.append('nbdev_install_git_hooks')
commands.append('nbdev_build_lib')
commands.append('git add *')
commands.append(f'git commit -m "{message}"')
commands.append('git push origin master')
for cmd in commands:
process = subprocess.Popen(cmd, stdout=subprocess.PIPE)
output, err = process.communicate()
os.chdir(start)
```
Save your notebook, and now we can push to `GitHub`. When pushing, make sure not to include spaces, as this will post an error.
```
git_push(git_path, 'Final')
```
| github_jupyter |
# Noisy data
Suppose that we have a dataset in which we have some measured attributes. Now, these attributes might carry some random error or variance. Such errors in attribute values are called as noise in the data.
If such errors persist in our data, it will return inaccurate results.
## Data cleaning
Real-world data tend to be noisy. Noisy data is data with a large amount of additional meaningless
information in it called noise. Data cleaning (or data cleansing) routines attempt to smooth out noise
while identifying outliers in the data.
There are three data smoothing techniques as follows:
1. Binning : Binning methods smooth a sorted data value by consulting its “neighborhood”, that is, the values around it.
2. Regression : It conforms data values to a function. Linear regression involves finding the “best” line to fit two attributes (or variables) so that one attribute can be used to predict the other.
3. Outlier analysis : Outliers may be detected by clustering, for example, where similar values are organized into groups, or “clusters”. Intuitively, values that fall outside of the set of clusters may be considered as outliers.
## 1. Binning
Types of smoothing using binning:
Smoothing by bin means : In smoothing by bin means, each value in a bin is replaced by the mean value of the bin.
Smoothing by bin median : In this method each bin value is replaced by its bin median value.
Smoothing by bin boundary : In smoothing by bin boundaries, the minimum and maximum values in a given bin are identified as the bin boundaries. Each bin value is then replaced by the closest boundary value.
Bin = [ 2, 6, 7, 9, 13, 20, 21, 25, 30 ]
Partition using equal frequency approach:
Bin 1 : 2, 6, 7
Bin 2 : 9, 13, 20
Bin 3 : 21, 24, 30
Smoothing by bin mean :
Bin 1 : 5, 5, 5
Bin 2 : 14, 14, 14
Bin 3 : 25, 25, 25
Smoothing by bin median :
Bin 1 : 6,6,6
Bin 2 : 13,13,13
Bin 3 : 24,24,24
Smoothing by bin boundaries :
Boundary_bins = [0,7,14,21,30]
Bin = [ 2, 6, 7, 9, 13, 20, 21, 25, 30 ]
New_bin = [ (0,7] , (0,7] , (0,7] , (7,14], (7,14], (14,21], (14,21], (25,30] , (25,30] ]
```
import pandas as pd
import matplotlib.pyplot as plt
ages = [20, 22, 25, 27, 21, 23, 37, 31, 61, 45, 41, 32]
bound_bins = [18, 25, 35, 60, 100]
categories = ['18_25','25_35' , '35_60' , '60_100' ]
cats = pd.cut(ages, bound_bins)
cats
cats[6]
y = pd.value_counts(cats)
y
plt.bar(categories,y)
plt.show()
cut_labels = ['young', 'adult', 'old', 'very_old']
cut_bins = [18, 25, 35, 60, 100]
cats = pd.cut(ages, bins=cut_bins, labels=cut_labels)
cats
y = pd.value_counts(cats)
y
plt.bar(cut_labels,y,width= 0.7)
plt.show()
cats2 = pd.qcut(ages, q = 4)
cats2
y = pd.value_counts(cats2)
y
```
## 2. Regression
Used to predict for individuals on the basis of information gained from a previous sample of similar individuals
## 3. Clustering
Clustering is the task of dividing the data points into a number of groups (you choose it) such that data points in the same groups are more similar to other data points in the same group and dissimilar to the data points in other groups.
Reference:
https://en.wikipedia.org/wiki/Data_binning
https://www.geeksforgeeks.org/ml-binning-or-discretization/
| github_jupyter |
# Information maximiser
Using neural networks, sufficient statistics can be obtained from data by maximising the Fisher information.
The neural network takes some data ${\bf d}$ and maps it to a compressed summary $\mathscr{f}:{\bf d}\to{\bf x}$ where ${\bf x}$ can have the same size as the dimensionality of the parameter space, rather than the data space.
To train the neural network a batch of simulations ${\bf d}_{\sf sim}^{\sf fid}$ created at a fiducial parameter value $\boldsymbol{\theta}^{\rm fid}$ are compressed by the neural network to obtain ${\bf x}_{\sf sim}^{\sf fid}$. From this we can calculate the covariance ${\bf C_\mathscr{f}}$ of the compressed summaries. We also use simulations ${\bf d}_{\sf sim}^{\sf fid+}$ created above the fiducial parameter value $\boldsymbol{\theta}^{\sf fid+}$ and simulations ${\bf d}_{\sf sim}^{\sf fid-}$ created below the fiducial parameter value $\boldsymbol{\theta}^{\sf fid-}$. These are compressed using the network and used to find mean of the summaries $\partial\boldsymbol{\mu}_\mathscr{f}/\partial\theta_\alpha\equiv\boldsymbol{\mu}_\mathscr{f},_\alpha$ where the numerical derivative is $({\bf x}_{\sf sim}^{\sf fid+}-{\bf x}_{\sf sim}^{\sf fid-})/(\boldsymbol{\theta}^{\sf fid+}-\boldsymbol{\theta}^{\sf fid-})$. We then use ${\bf C}_\mathscr{f}$ and $\boldsymbol{\mu}_\mathscr{f},_\alpha$ to calculate the Fisher information
$${\bf F}_{\alpha\beta} = \boldsymbol{\mu}_\mathscr{f},^T_{\alpha}{\bf C}^{-1}_\mathscr{f}\boldsymbol{\mu}_\mathscr{f},_{\beta}.$$
We want to maximise the Fisher information so to train the network we use the loss function
$$\Lambda = -\frac{1}{2}|{\bf F}_{\alpha\beta}|.$$
When using this code please cite <a href="https://arxiv.org/abs/1802.03537">arXiv:1802.03537</a>.<br><br>
The code in the paper can be downloaded as v1 or v1.1 of the code kept on zenodo:<br><br>
[](https://doi.org/10.5281/zenodo.1175196)
<br>
The code presented below is version two (and is much more powerful) and is under constant development.
This code is run using<br>
>`python-3.6.6`
>`jupyter-4.4.0`
>`tensorflow-1.10.1`
>`numpy-1.14.5`
>`tqdm==4.25.0`
>`sys (native)`
Although these precise versions may not be necessary, I have put them here to avoid possible conflicts.
## Load modules
```
%matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
import IMNN
```
## Generate data
In this example we are going to use $n_{\bf d}=10$ data points of a 1D field of Gaussian noise with zero mean and unknown variance to see if the network can learn to summarise this variance.<br><br>
We start by defining a function to generate the data with the correct shape. The shape must be
```
data_shape = None + input shape
```
```
input_shape = [10]
```
It is useful to define the generating function so that it only takes in the value of the parameter as its input since the function can then be used for ABC later.<br><br>
The data needs to be generated at a fiducial parameter value and at perturbed values just below and above the fiducial parameter for the numerical derivative.
```
θ_fid = 1.
Δθpm = 0.1
```
The data at the perturbed values should have the shape
```
perturbed_data_shape = None + number of parameters + input shape
```
The generating function is defined so that the fiducial parameter is passed as a list so that many simulations can be made at once. This is very useful for the ABC function later.
```
def generate_data(θ, train = False):
if train:
return np.moveaxis(np.random.normal(0., np.sqrt(θ), [1] + input_shape + [len(θ)]), -1, 0)
else:
return np.moveaxis(np.random.normal(0., np.sqrt(θ), input_shape + [len(θ)]), -1, 0)
```
### Training data
Enough data needs to be made to approximate the covariance matrix of the output summaries. The number of simulations needed to approximate the covariance is `n_s`. If the data is particularly large then it might not be possible to pass all the data into active memory at once and so several the simulations can be split into batches.
For example if we wanted to make 2000 simulations, but estimate the covariance using 1000 simulations at a time
we would set
```python
n_s = 1000
n_train = 2
```
We're going to use 1000 simulations to approximate the covariance and use only 1 combination
```
n_s = 1000
n_train = 1
```
The training data can now be made
```
t = generate_data(θ = [θ_fid for i in range(n_train * n_s)], train = False)
```
By suppressing the sample variance between the simulations created at the lower and upper parameter values for the numerical derivative, far fewer simulations are needed. We choose to use 5% of the total number of simulations.
```
derivative_fraction = 0.05
n_p = int(n_s * derivative_fraction)
```
The sample variance can be supressed by choosing the same initial seed when creating the upper and lower simulations.
```
seed = np.random.randint(1e6)
np.random.seed(seed)
t_m = generate_data(θ = [θ_fid - Δθpm for i in range(n_train * n_p)], train = True)
np.random.seed(seed)
t_p = generate_data(θ = [θ_fid + Δθpm for i in range(n_train * n_p)], train = True)
np.random.seed()
```
We also need to get the denominator of the derivative which is given by the difference between the perturbed parameter values<br><br>
$$\frac{\partial}{\partial\theta} = \frac{1}{2\Delta\theta_{\pm}}.$$<br>
This needs to be done for every parameter and kept in a numpy array of shape `[number of parameters]`.
```
derivative_denominator = 1. / (2. * Δθpm)
der_den = np.array([derivative_denominator])
```
The fiducial simulations and simulations for the derivative must be collected in a dictionary to be stored on the GPU or passed to the training function.
```
data = {"x_central": t, "x_m": t_m, "x_p":t_p}
```
### Test data
We should also make some test data, but here we will use only one combination `n_train = 1`. This needs adding to the dictionary
```
tt = generate_data([θ_fid for i in range(n_s)])
seed = np.random.randint(1e6)
np.random.seed(seed)
tt_m = generate_data([θ_fid - Δθpm for i in range(n_p)], train = True)
np.random.seed(seed)
tt_p = generate_data([θ_fid + Δθpm for i in range(n_p)], train = True)
np.random.seed()
data["x_central_test"] = tt
data["x_m_test"] = tt_m
data["x_p_test"] = tt_p
```
### Data visualisation
We can plot the data to see what it looks like.
```
fig, ax = plt.subplots(1, 1, figsize = (10, 6))
ax.plot(data['x_central'][np.random.randint(n_train * n_s)], label = "training data")
ax.plot(data['x_central_test'][np.random.randint(n_s)], label = "test data")
ax.legend(frameon = False)
ax.set_xlim([0, 9])
ax.set_xticks([])
ax.set_ylabel("Data amplitude");
```
It is also very useful to plot the upper and lower derivatives to check that the sample variance is actually supressed since the network learns extremely slowly if this isn't done properly.
```
fig, ax = plt.subplots(2, 1, figsize = (10, 10))
plt.subplots_adjust(hspace = 0)
training_index = np.random.randint(n_train * n_p)
ax[0].plot(data['x_m'][training_index, 0], label = "lower training data", color = 'C0')
ax[0].plot(data['x_p'][training_index, 0], label = "upper training data", color = 'C0', linestyle = 'dashed')
test_index = np.random.randint(n_p)
ax[0].plot(data['x_m_test'][test_index, 0], label = "lower test data", color = 'C1')
ax[0].plot(data['x_p_test'][test_index, 0], label = "upper test data", color = 'C1')
ax[0].legend(frameon = False)
ax[0].set_xlim([0, 9])
ax[0].set_xticks([])
ax[0].set_ylabel("Data amplitude")
ax[1].axhline(xmin = 0., xmax = 1., y = 0., linestyle = 'dashed', color = 'black')
ax[1].plot(data['x_p'][training_index, 0] - data['x_m'][training_index, 0], color = 'C0')
ax[1].plot(data['x_p_test'][test_index, 0] - data['x_m_test'][test_index, 0], color = 'C1')
ax[1].set_xlim([0, 9])
ax[1].set_xticks([])
ax[1].set_ylabel("Difference between derivative data amplitudes");
```
## Initiliase the neural network
### Define network parameters
The network works with a base set of parameters which are<br>
> `'verbose'` - `bool` - whether to print out diagnostics
> `'number of simulations'` - `int` - the number of simulations to use in any one combination
> `'differentiation fraction'` - `float` - a fraction of the simulations to use for the numerical derivative
> `'fiducial θ'` - `array` - fiducial parameters in an array
> `'derivative denominator'` - `array` - denominator of the numerical derivative for each parameter
> `'number of summaries'` - `int` - number of summaries the network makes from the data
> `'input shape'` - `int` or `list` - the number of inputs or the shape of the input if image-like input
> `'preload data'` - `dict` or `None` - the training (and test) data to preload as a TensorFlow constant in a dictionary, no preloading is done if `None`
> `'calculate MLE'` - `bool` - whether to calculate the maximum likelihood estimate
> `'prebuild'` - `bool` - whether to get the network to build a network or to provided your own
> `'save file'` - `string` - a file name to save the graph (not saved if wrong type or not given)
```python
parameters = {
'verbose': True,
'number of simulations': n_s,
'differentiation fraction': derivative_fraction,
'fiducial θ': np.array([θ_fid]),
'derivative denominator': der_den,
'number of summaries': 1,
'input shape': input_shape,
'preload data': data,
'calculate MLE': True,
'prebuild': False,
}
```
The module can also build simple convolutional or dense networks (or a mixture of the two), which can be trigger by setting `'prebuild': True`. Several parameters are required to allow the network to build the network. These are<br>
> `'wv'` - `float` - the variance with which to initialise the weights. If this is 0 or less, the network will determine the weight variance according to He initialisation
> `'bb'` - `float` - the constant value with which to initialise the biases
> `'activation'` - `TensorFlow function` - a native tensorflow activation function
> `'α'` - `float` or `int` - an additional parameter, if needed, for the tensorflow activation function
> `'hidden layers'` - `list` - the architecture of the network. each element of the list is a hidden layer. A dense layer can be made using an integer where thet value indicates the number of neurons. A convolutional layer can be built by using a list where the first element is an integer where the number describes the number of filters, the second element is a list of the kernel size in the x and y directions, the third elemnet is a list of the strides in the x and y directions and the final element is string of 'SAME' or 'VALID' which describes the padding prescription.
Here is an example of the IMNN which uses 1000 simulations per combination and 50 upper and 50 lower simulations per derivative for a model with one parameter where we require one summary which are preloaded as a TensorFlow constant where we want to have access to the precomputed maximum likelihood estimate. The module will build the network which takes in an input array of shape `[10]` and allows the network to decide the weight initialisation, initialises the biases at `bb = 0.1` and uses `tf.nn.leaky_relu` activation with a negative gradient parameter of `α = 0.01`. The network architecture has two fully connected hidden layers with 128 neurons in each layer. The graph is saved into a file in the `data` folder called `model.meta`.
```
parameters = {
'verbose': True,
'number of simulations': n_s,
'fiducial θ': np.array([θ_fid]),
'derivative denominator': der_den,
'differentiation fraction': derivative_fraction,
'number of summaries': 1,
'calculate MLE': True,
'prebuild': True,
'input shape': input_shape,
'preload data': data,
'save file': "data/model",
'wv': 0.,
'bb': 0.1,
'activation': tf.nn.leaky_relu,
'α': 0.01,
'hidden layers': [128, 128]
}
```
### Self-defined network
A self defined network can be used instead of letting the module build the network for you. This function needs to take in two input tensors, the first is the shape of the input with `None` in the first axis and the second tensor is a tensorflow float (which will be the dropout). Since the weights need to be shared between several corresponding networks each set of trainable variables must be defined in its own scope. An example of the above network defined outside of the module is
```python
def network(input_tensor, dropout):
with tf.variable_scope('layer_1'):
weights = tf.get_variable("weights", [10, 128], initializer = tf.random_normal_initializer(0., 1.))
biases = tf.get_variable("biases", [128], initializer = tf.constant_initializer(0.1))
x = tf.matmul(input_tensor, weights)
x = tf.add(x, biases)
x = tf.nn.leaky_relu(x, 0.01)
x = tf.nn.dropout(x, dropout)
with tf.variable_scope('layer_2'):
weights = tf.get_variable("weights", [128, 128], initializer = tf.random_normal_initializer(0., 1.))
biases = tf.get_variable("biases", [128], initializer = tf.constant_initializer(0.1))
x = tf.matmul(x, weights)
x = tf.add(x, biases)
x = tf.nn.leaky_relu(x, 0.01)
x = tf.nn.dropout(x, dropout)
x = tf.reshape(x, (-1, 300))
with tf.variable_scope('layer_3'):
weights = tf.get_variable("weights", [128, 1], initializer = tf.random_normal_initializer(0., np.sqrt(2. / 300)))
biases = tf.get_variable("biases", [1], initializer = tf.constant_initializer(0.1))
x = tf.matmul(x, weights)
x = tf.add(x, biases)
x = tf.nn.leaky_relu(x, 0.01)
x = tf.nn.dropout(x, dropout)
return x
```
### Initialise the network
```
n = IMNN.IMNN(parameters = parameters)
```
### Build the network
To build the network a learning rate, η, must be defined.
```
η = 1e-3
```
The `setup(η)` function initialises the input tensors, builds the network and defines the optimisation scheme. If a self-defined network function (`network(a, b)`) has been constructed this can be passed to the setup function
```python
n.setup(η = η, network = network)
```
```
n.setup(η = η)
```
### Changing minimisation scheme
By default the optimation scheme is<br>
```python
n.backpropagate = tf.train.GradientDescentOptimizer(η).minimize(n.Λ)
```
To use any other training scheme, such as the `Adam` optimiser, it is sufficient to run
```python
n.backpropagate = tf.train.AdamOptimizer(η, β1, β2, ε).minimize(n.Λ)
```
after `setup(η)` to override the default minimisation routine. Note that testing with Adam optimiser has found it to be incredibly unstable. If you want to continue to use the default minimisation routine but want to change the learning rate without reinitialising you can run
```python
n.training_scheme(η = new_η)
```
## Train the network
With the data we can now train the network. The function simply takes the number of epochs, `num_epochs`, the fraction of neurons kept when using dropout `keep_rate`, and the denominator for the derivative calculated earlier, `der_den`.
```
num_epochs = 500
keep_rate = 0.8
```
If the data has not been preloaded as a TensorFlow constant then it can be passed to the train function
```python
train_F, test_F = n.train(num_epochs = num_epochs, n_train = n_train, keep_rate = keep_rate, data = data)
```
We can run
```
n.train(num_epochs = num_epochs, n_train = n_train, keep_rate = keep_rate, data = data, history = True)
```
The train function automatically collects a dictionary of history elements when `history = True`. When `history = False` the dictionary only contains
> `'F'` - the Fisher information at each epoch
> `'det(F)'` - the determinant of the Fisher information
When `history = True` then the dictionary also contains
> `'Λ'` - the loss function of the training data
> `'μ'` - the mean of the fiducial simulations
> `'C'` - the covariance of the fiducial simulations
> `'det(C)'` - the determinant of the fiducial simulations
> `'dμdθ'` - the mean of the numerical derivative of the simulations
Test version of each of these quantities is also calculated if there is provided test data
> `'test F'` - the Fisher information of the test data
> `'det(test F)'` - the determinant of the Fisher information from the test data
> `'Λ'` - the loss function of the test data
> `'μ'` - the mean of the fiducial test simulations
> `'C'` - the covariance of the fiducial test simulations
> `'det(C)'` - the determinant of the fiducial test simulations
> `'dμdθ'` - the mean of the numerical derivative of the test simulations
```
fig, ax = plt.subplots(5, 1, sharex = True, figsize = (8, 14))
plt.subplots_adjust(hspace = 0)
end = len(n.history["det(F)"])
epochs = np.arange(end)
a, = ax[0].plot(epochs, n.history["det(F)"], label = 'Training data')
b, = ax[0].plot(epochs, n.history["det(test F)"], label = 'Test data')
ax[0].legend(frameon = False)
ax[0].set_ylabel(r'$|{\bf F}_{\alpha\beta}|$')
ax[1].plot(epochs, n.history["Λ"])
ax[1].plot(epochs, n.history["test Λ"])
ax[1].set_xlabel('Number of epochs')
ax[1].set_ylabel(r'$\Lambda$')
ax[1].set_xlim([0, len(epochs)]);
ax[2].plot(epochs, n.history["det(C)"])
ax[2].plot(epochs, n.history["det(test C)"])
ax[2].set_xlabel('Number of epochs')
ax[2].set_ylabel(r'$|{\bf C}|$')
ax[2].set_xlim([0, len(epochs)]);
ax[3].plot(epochs, np.array(n.history["dμdθ"]).reshape((np.prod(np.array(n.history["dμdθ"]).shape))))
ax[3].plot(epochs, np.array(n.history["test dμdθ"]).reshape((np.prod(np.array(n.history["test dμdθ"]).shape))))
ax[3].set_ylabel(r'$\partial\mu/\partial\theta$')
ax[3].set_xlabel('Number of epochs')
ax[3].set_xlim([0, len(epochs)])
ax[4].plot(epochs, np.array(n.history["μ"]).reshape((np.prod(np.array(n.history["μ"]).shape))))
ax[4].plot(epochs, np.array(n.history["test μ"]).reshape((np.prod(np.array(n.history["test μ"]).shape))))
ax[4].set_ylabel('μ')
ax[4].set_xlabel('Number of epochs')
ax[4].set_xlim([0, len(epochs)])
print()
```
We can see that the test loss deviates from the training loss. This is to be expected because there are will be a lot of correlation within a small training set which isn't in the test set. As long as the test loss doesn't start increasing then it is likely that the network is still working, with the maximum Fisher available from the network is the value obtained from the test set.
## Resetting the network
If you need to reset the weights and biases for any reason then you can call
```python
n.reinitialise_session()
```
## Saving the network
If you don't initialise the network with a save name you can save the network as a `TensorFlow` `meta` graph. For example saving the model in the directory `/.data` called `saved_model.meta` can be done using the function
```python
n.save_network(file_name = "data/saved_model", first_time = True)
```
If `save file` is passed with a correct file name when initialising the module then the initialised network will be saved by
```python
n.begin_session()
```
and then saved at the end of training.
## Loading the network
You can load the network from a `TensorFlow` `meta` graph (from `/.data/saved_model.meta`) using the same parameter dictionary as used when originally training the network and then running
```python
n = IMNN.IMNN(parameters = parameters)
n.restore_network()
```
Training can be continued after restoring the model - although the Adam optimiser might need to reacquaint itself.
## Approximate Bayesian computation
We can now do ABC (or PMC-ABC) with our calculated summary. First we generate some simulated real data:
```
real_data = generate_data(θ = [1.], train = False)
```
We can plot this real data to see what it looks like.
```
fig, ax = plt.subplots(1, 1, figsize = (10, 6))
ax.plot(real_data[0], label = "real data")
ax.legend(frameon = False)
ax.set_xlim([0, 9])
ax.set_xticks([])
ax.set_ylabel("Data amplitude");
```
### ABC
We now perform ABC by drawing 100000 random samples from the prior. We define the upper and lower bounds of a uniform prior to be 0 and 10. Only a uniform prior is implemented at the moment. From the samples we create simulations at each parameter value and feed each simulation through the network to get summaries. The summaries are compared to the summary of the real data to find the distances which can be used to accept or reject points.
Because the simulations are created within the ABC function then the generation function must be passed. This is why the generator should be of the form defined above, which takes only a list of parameter values and returns a simulation at each parameter.
If the data is not preloaded as a TensorFlow constant then the data can be passed to the function as
```python
θ, summary, s, ρ, F = n.ABC(real_data = real_data, prior = [0, 10], draws = 100000, generate_simulation = generate_data, at_once = True, data = data)
```
Here we can use
```
θ, summary, s, ρ, F = n.ABC(real_data = real_data, prior = [0, 10], draws = 100000, generate_simulation = generate_data, at_once = True, data = data)
```
If the simulations are going to be too large to make all at once the `at_once` option can be set to false which will create one simulation at a time.
```python
θ, summary, s, ρ, F = n.ABC(real_data = real_data, der_den = der_den, prior = [0, 10], draws = 100000, generate_simulation = generate_data, at_once = False)
```
### Accept or reject
In ABC draws are accepted if the distance between the simulation summary and the simulation of the real data are "close", i.e. smaller than some ϵ value, which is chosen somewhat arbitrarily.
```
ϵ = 1
accept_indices = np.argwhere(ρ < ϵ)[:, 0]
reject_indices = np.argwhere(ρ >= ϵ)[:, 0]
```
### Plot samples
We can plot the output samples and the histogram of the accepted samples, which should peak around `θ = 1` (where we generated the real data). The monotonic function of all the output samples shows that the network has learned how to summarise the data.
```
fig, ax = plt.subplots(2, 1, sharex = True, figsize = (10, 10))
plt.subplots_adjust(hspace = 0)
ax[0].scatter(θ[accept_indices] , s[accept_indices, 0], s = 1)
ax[0].scatter(θ[reject_indices], s[reject_indices, 0], s = 1, alpha = 0.1)
ax[0].plot([0, 10], [summary[0], summary[0]], color = 'black', linestyle = 'dashed')
ax[0].set_ylabel('Network output', labelpad = 0)
ax[0].set_xlim([0, 10])
ax[1].hist(θ[accept_indices], np.linspace(0, 10, 100), histtype = u'step', density = True, linewidth = 1.5, color = '#9467bd');
ax[1].set_xlabel('$\\theta$')
ax[1].set_ylabel('$\\mathcal{P}(\\theta|{\\bf d})$')
ax[1].set_yticks([]);
```
There can be a lot of $\theta$ draws which are unconstrained by the network because no similar structures were seen in the data which is indicative of using too small of a small training set.
## PMC-ABC
Population Monte Carlo ABC is a way of reducing the number of draws by first sampling from a prior, accepting the closest 75% of the samples and weighting all the rest of the samples to create a new proposal distribution. The furthest 25% of the original samples are redrawn from the new proposal distribution. The furthest 25% of the simulation summaries are continually rejected and the proposal distribution updated until the number of draws needed accept all the 25% of the samples is much greater than this number of samples. This ratio is called the criterion. The inputs work in a very similar way to the `ABC` function above. If we want 1000 samples from the approximate distribution at the end of the PMC we need to set `num_keep = 1000`. The initial random draw (as in ABC above) initialises with `num_draws`, the larger this is the better proposal distribution will be on the first iteration.
If the data is not preloaded as a TensorFlow constant then the data can be passed to the function as
```python
θ_, summary_, ρ_, s_, W, total_draws, F = n.PMC(real_data = real_data, prior = [0, 10], num_draws = 1000, num_keep = 1000, generate_simulation = generate_data, criterion = 0.1, data = data, at_once = True, samples = None)```
Here we can use
```
θ_, summary_, ρ_, s_, W, total_draws, F = n.PMC(real_data = real_data, prior = [0, 10], num_draws = 1000, num_keep = 1000, generate_simulation = generate_data, criterion = 0.1, at_once = True, samples = None, data = data)
```
If we want the PMC to continue for longer we can provide the output of PMC as an input as
```python
θ_, summary_, ρ_, s_, W, total_draws, F = n.PMC(real_data = real_data, der_den = der_den, prior = [0, 10], num_draws = 1000, num_keep = 1000, generate_simulation = generate_data, criterion = 0.01, data = data, at_once = True, samples = [θ_, summary_, ρ_, s_, W, total_draws, F])```
Finally we can plot the accepted samples and plot their histogram.
```
fig, ax = plt.subplots(2, 1, sharex = True, figsize = (10, 10))
plt.subplots_adjust(hspace = 0)
ax[0].scatter(θ_ , s_, s = 1)
ax[0].plot([0, 10], [summary[0], summary[0]], color = 'black', linestyle = 'dashed')
ax[0].set_ylabel('Network output', labelpad = 0)
ax[0].set_xlim([0, 10])
ax[0].set_ylim([np.min(s_), np.max(s_)])
ax[1].hist(θ_, np.linspace(0, 10, 100), histtype = u'step', density = True, linewidth = 1.5, color = '#9467bd');
ax[1].set_xlabel('θ')
ax[1].set_ylabel('$\\mathcal{P}(\\theta|{\\bf d})$')
ax[1].set_yticks([]);
```
## Maximum likelihood estimate
We can also calculate the first-order Gaussian approximation of the posterior on the parameter and find a maximum likelihood estimate.
If the data is not preloaded as a TensorFlow constant then it can be passed using
```python
asymptotic_likelihood = n.asymptotic_likelihood(real_data = real_data, prior = np.linspace(0, 10, 1000).reshape((1, 1, 1000)), data = data)
MLE = n.θ_MLE(real_data = real_data, data = data)
```
Here we will use
```
asymptotic_likelihood = n.asymptotic_likelihood(real_data = real_data, prior = np.linspace(0, 10, 1000).reshape((1, 1, 1000)), data = data)
MLE = n.θ_MLE(real_data = real_data, data = data)
fig, ax = plt.subplots(1, 1, figsize = (10, 6))
ax.plot(np.linspace(0, 10, 1000), asymptotic_likelihood[0, 0], linewidth = 1.5)
ax.axvline(x = MLE[0, 0], ymin = 0., ymax = 1., linestyle = 'dashed', color = 'black')
ax.set_xlabel("θ")
ax.set_xlim([0, 10])
ax.set_ylabel('$\\mathcal{P}(\\theta|{\\bf d})$')
ax.set_yticks([]);
```
## Analytic posterior calculation
We know what the analytic posterior is for this model
$$\mathcal{P}(\boldsymbol{\theta}|{\bf d}) = \frac{\displaystyle{\rm exp}\left[-\frac{1}{2\boldsymbol{\theta}}\sum_{i = 1}^{n_{\bf d}}d_i\right]}{(2\pi\boldsymbol{\theta})^{n_{\bf d}/2}}.$$
We can there for plot this as a comparison.
```
θ_grid = np.linspace(0.001, 10, 1000)
analytic_posterior = np.exp(-0.5 * np.sum(real_data**2.) / θ_grid) / np.sqrt(2. * np.pi * θ_grid)**10.
analytic_posterior = analytic_posterior / np.sum(analytic_posterior * (θ_grid[1] - θ_grid[0]))
fig, ax = plt.subplots(1, 1, figsize = (10, 6))
ax.plot(θ_grid, analytic_posterior, linewidth = 1.5, color = 'C1', label = "Analytic posterior")
ax.hist(θ_, np.linspace(0, 10, 100), histtype = u'step', density = True, linewidth = 1.5, color = '#9467bd', label = "PMC posterior");
ax.hist(θ[accept_indices], np.linspace(0, 10, 100), histtype = u'step', density = True, linewidth = 1.5, color = 'C2', label = "ABC posterior")
ax.plot(np.linspace(0, 10, 1000), asymptotic_likelihood[0, 0], color = 'C0', linewidth = 1.5, label = "Asymptotic Gaussian likelihood")
ax.axvline(x = MLE[0, 0], ymin = 0., ymax = 1., linestyle = 'dashed', color = 'black', label = "Maximum likelihood estimate")
ax.legend(frameon = False)
ax.set_xlim([0, 10])
ax.set_xlabel('θ')
ax.set_ylabel('$\\mathcal{P}(\\theta|{\\bf d})$')
ax.set_yticks([]);
```
| github_jupyter |
# Apartado 5 - Funciones
- funciones con & sin `return`
- funciones con valores por defecto
- recursividad
--------------------------------------------------------------------------------------
## Funciones `print`
```
print("Yo soy 'así'")
print('Yo soy "así"')
_ = "así"
print("Yo soy " + _ + " .")
_ = 15
x = "Yo tengo " + str(_) + " años."
print(x)
print("Yo tengo", _ , " años.")
def prtn(x, r, y, t):
print(x + r + y + t)
prtn(x="Yo", r=" tengo", y=" 15", t=" años.")
def prtn(x, r, y, t):
print(x + r + y + t)
prtn("Yo", " tengo", " 15", " años.")
def p(*args):
print(args)
p(2, 4, 6, 8, 3, 1, 3)
def x():
a = 2
b = 4
suma = a + b
print(suma)
x()
```
--------------------------------------------------------------------------------------
## Funciones `return`
```
def x():
a = 2
b = 4
suma = a + b
#print("suma:", suma)
return a
z = x()
print("z:", z)
a = 3
def nombre_funcion():
a = 2
b = 4
suma3 = a + b
print("suma3:", suma3)
print("a:", a)
nombre_funcion()
a = 3
def nombre_funcion():
print("a2:", a)
b = 4
suma3 = a + b
print("suma3:", suma3)
print("a0:", a)
nombre_funcion()
def nombre_funcion():
a = 2
b = 4
suma3 = a + b
return suma3
print(nombre_funcion())
def nombre_funcion():
a = 2
b = 4
suma3 = a + b
print(nombre_funcion())
s = "HOLA"
s.lower()
def nombre_funcion():
a = 2
b = 4
suma3 = a + b
resta = a - b
print(suma3)
print(resta)
return resta
print(suma3)
return ["2", 2]
x = nombre_funcion()
print(x)
def suma():
a = 2
b = 3
return a + b
def multiplicacion():
a = 2
b = 3
return a * b
def f():
x = suma()
y = multiplicacion()
h = x / y
return h
x = f()
print(x)
suma = 2
x = suma()
# Esto es un comentario
""" Esto es otro"""
def suma_generica(arg1, arg2):
""" Suma arg1 y arg2
Esta es la segunda línea de la documentación de la función
"""
a = 2 # Esta variable vale 2
return arg1 + arg2
a = 3
b = 7
x = suma_generica(arg1=a, arg2=b)
print(x)
```
`help`
```
help(suma_generica)
def suma_generica(arg1, arg2):
""" Suma arg1 y arg2 """
if arg1.isnumeric() and arg2.isnumeric():
return int(arg1) + int(arg2)
else:
print("Solo se pueden sumar números")
a = input()
b = input()
x = suma_generica(arg1=a, arg2=b)
print(x)
def suma_generica(arg1, arg2):
""" Suma arg1 y arg2 """
if arg1.isnumeric() and arg2.isnumeric():
return int(arg1) + int(arg2)
else:
return "Solo se pueden sumar números"
a = input()
b = input()
x = suma_generica(arg1=a, arg2=b)
print(x)
```
--------------------------------------------------------------------------------------
## Funciones con valores por defecto
```
def mi_funcion_1(a, b=10):
return a**b
r = mi_funcion(a=2)
r
def mi_funcion_2(a, b=10):
return a**b
r = mi_funcion(a=2, b=5)
r
def mi_funcion_3(a, b):
return a**b
k = int(input("Edad:"))
h = int(input("Altura:"))
r = mi_funcion(a=k, b=h)
type(r)
b = None
str(b)
def mi_funcion_4(a=5, b=None, c=None, y=None, h=None):
if b == None:
b = len(str(None))
return a**b
r = mi_funcion()
r
def mi_funcion_5(a, b=4, c=None, y=None, h=None):
return a**b
r = mi_funcion(a=2)
r
def mi_funcion_6(a, b=4, c=676, y=None, h=None):
return a**b
r = mi_funcion(a=2)
r
def mi_funcion_7(b, a=7, c=676, y=None, h=None):
return a**b
r = mi_funcion(b=2)
r
```
**Ejemplos:**
- [Función de matplotlib](https://matplotlib.org/api/_as_gen/matplotlib.pyplot.pie.html#matplotlib.pyplot.pie)
- [Función de pandas](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.to_csv.html)
--------------------------------------------------------------------------------------
## Recursividad
```
def nombre_funcion():
print("X")
nombre_funcion()
def nombre_funcion(x):
print(x)
nombre_funcion(x="4")
def nombre_funcion(x):
return x
s = nombre_funcion(x=3)
print(s)
def f_recursiva():
print("X")
f_recursiva()
```
```python
def f():
print("Muestro esto")
f()
print("X")
f()
```
```
def f(acum):
if acum == 10:
return acum
print("acum:", acum)
f(acum=acum+1)
x = f(acum=0)
print("x:", x)
def f(acum):
if acum == 0:
return acum
print("acum:", acum)
p = f(acum=acum+1)
x = f(acum=0)
print(x)
print("Ha terminado esto")
def f(acum):
if acum == 10:
return acum
print("acum:", acum)
p = f(acum=acum+1)
return p
x = f(acum=0)
print(x)
print("Ha terminado esto")
```
--------------------------------------------------------------------------------------
| github_jupyter |
```
%%html
<link href="http://mathbook.pugetsound.edu/beta/mathbook-content.css" rel="stylesheet" type="text/css" />
<link href="https://aimath.org/mathbook/mathbook-add-on.css" rel="stylesheet" type="text/css" />
<style>.subtitle {font-size:medium; display:block}</style>
<link href="https://fonts.googleapis.com/css?family=Open+Sans:400,400italic,600,600italic" rel="stylesheet" type="text/css" />
<link href="https://fonts.googleapis.com/css?family=Inconsolata:400,700&subset=latin,latin-ext" rel="stylesheet" type="text/css" /><!-- Hide this cell. -->
<script>
var cell = $(".container .cell").eq(0), ia = cell.find(".input_area")
if (cell.find(".toggle-button").length == 0) {
ia.after(
$('<button class="toggle-button">Toggle hidden code</button>').click(
function (){ ia.toggle() }
)
)
ia.hide()
}
</script>
```
**Important:** to view this notebook properly you will need to execute the cell above, which assumes you have an Internet connection. It should already be selected, or place your cursor anywhere above to select. Then press the "Run" button in the menu bar above (the right-pointing arrowhead), or press Shift-Enter on your keyboard.
$\newcommand{\identity}{\mathrm{id}}
\newcommand{\notdivide}{\nmid}
\newcommand{\notsubset}{\not\subset}
\newcommand{\lcm}{\operatorname{lcm}}
\newcommand{\gf}{\operatorname{GF}}
\newcommand{\inn}{\operatorname{Inn}}
\newcommand{\aut}{\operatorname{Aut}}
\newcommand{\Hom}{\operatorname{Hom}}
\newcommand{\cis}{\operatorname{cis}}
\newcommand{\chr}{\operatorname{char}}
\newcommand{\Null}{\operatorname{Null}}
\newcommand{\lt}{<}
\newcommand{\gt}{>}
\newcommand{\amp}{&}
$
<div class="mathbook-content"><h2 class="heading hide-type" alt="Exercises 22.4 Additional Exercises: Error Correction for BCH Codes"><span class="type">Section</span><span class="codenumber">22.4</span><span class="title">Additional Exercises: Error Correction for <abbr class="acronym">BCH</abbr> Codes</span></h2><a href="finite-exercises-bch-codes.ipynb" class="permalink">¶</a></div>
<div class="mathbook-content"></div>
<div class="mathbook-content"><p id="p-3549"><abbr class="acronym">BCH</abbr> codes have very attractive error correction algorithms. Let $C$ be a <abbr class="acronym">BCH</abbr> code in $R_n\text{,}$ and suppose that a code polynomial $c(t) = c_0 + c_1 t + \cdots + c_{n-1} t^{n-1}$ is transmitted. Let $w(t) = w_0 + w_1 t + \cdots w_{n-1} t^{n-1}$ be the polynomial in $R_n$ that is received. If errors have occurred in bits $a_1, \ldots, a_k\text{,}$ then $w(t) = c(t) + e(t)\text{,}$ where $e(t) = t^{a_1} + t^{a_2} + \cdots + t^{a_k}$ is the <dfn class="terminology">error polynomial</dfn>. The decoder must determine the integers $a_i$ and then recover $c(t)$ from $w(t)$ by flipping the $a_i$th bit. From $w(t)$ we can compute $w( \omega^i ) = s_i$ for $i = 1, \ldots, 2r\text{,}$ where $\omega$ is a primitive $n$th root of unity over ${\mathbb Z}_2\text{.}$ We say the <dfn class="terminology">syndrome</dfn> of $w(t)$ is $s_1, \ldots, s_{2r}\text{.}$</p></div>
<div class="mathbook-content"><article class="exercise-like" id="exercise-759"><h6 class="heading"><span class="codenumber">1</span></h6><p id="p-3550">Show that $w(t)$ is a code polynomial if and only if $s_i = 0$ for all $i\text{.}$</p></article></div>
<div class="mathbook-content"><article class="exercise-like" id="exercise-760"><h6 class="heading"><span class="codenumber">2</span></h6><p id="p-3551">Show that</p><div class="displaymath">
\begin{equation*}
s_i = w( \omega^i) = e( \omega^i) = \omega^{i a_1} + \omega^{i a_2} + \cdots + \omega^{i a_k}
\end{equation*}
</div><p>for $i = 1, \ldots, 2r\text{.}$ The <dfn class="terminology">error-locator polynomial</dfn> is defined to be</p><div class="displaymath">
\begin{equation*}
s(x) = (x + \omega^{a_1})(x + \omega^{a_2}) \cdots (x + \omega^{a_k}).
\end{equation*}
</div></article></div>
<div class="mathbook-content"><article class="exercise-like" id="exercise-761"><h6 class="heading"><span class="codenumber">3</span></h6><p id="p-3552">Recall the $(15,7)$-block <abbr class="acronym">BCH</abbr> code in Example <a href="section-poly-codes.ipynb#example-finite-parity-check-x7-1" class="xref" alt="Example 22.19 " title="Example 22.19 ">22.19</a>. By Theorem <a href="section-error-detecting-correcting-codes.ipynb#theorem-min-distance" class="xref" alt="Theorem 8.13 " title="Theorem 8.13 ">8.13</a>, this code is capable of correcting two errors. Suppose that these errors occur in bits $a_1$ and $a_2\text{.}$ The error-locator polynomial is $s(x) = (x + \omega^{a_1})(x + \omega^{a_2})\text{.}$ Show that</p><div class="displaymath">
\begin{equation*}
s(x) = x^2 + s_1 x + \left( s_1^2 + \frac{s_3}{s_1} \right).
\end{equation*}
</div></article></div>
<div class="mathbook-content"><article class="exercise-like" id="exercise-762"><h6 class="heading"><span class="codenumber">4</span></h6><p id="p-3553">Let $w(t) = 1 + t^2 +t^4 + t^5 + t^7 + t^{12} + t^{13}\text{.}$ Determine what the originally transmitted code polynomial was.</p></article></div>
| github_jupyter |
```
import numpy as np
import pandas as pd
from bqplot import (
Axis, ColorAxis, LinearScale, DateScale, DateColorScale, OrdinalScale,
OrdinalColorScale, ColorScale, Scatter, Lines, Figure, Tooltip
)
from ipywidgets import Label
```
## Get Data
```
price_data = pd.DataFrame(np.cumsum(np.random.randn(150, 2).dot([[1.0, -0.8], [-0.8, 1.0]]), axis=0) + 100,
columns=['Security 1', 'Security 2'], index=pd.date_range(start='01-01-2007', periods=150))
size = 100
np.random.seed(0)
x_data = range(size)
y_data = np.cumsum(np.random.randn(size) * 100.0)
ord_keys = np.array(['A', 'B', 'C', 'D', 'E', 'F'])
ordinal_data = np.random.randint(5, size=size)
symbols = ['Security 1', 'Security 2']
dates_all = price_data.index.values
dates_all_t = dates_all[1:]
sec1_levels = np.array(price_data[symbols[0]].values.flatten())
log_sec1 = np.log(sec1_levels)
sec1_returns = log_sec1[1:] - log_sec1[:-1]
sec2_levels = np.array(price_data[symbols[1]].values.flatten())
```
# Basic Scatter
```
sc_x = DateScale()
sc_y = LinearScale()
scatt = Scatter(x=dates_all, y=sec2_levels, scales={'x': sc_x, 'y': sc_y})
ax_x = Axis(scale=sc_x, label='Date')
ax_y = Axis(scale=sc_y, orientation='vertical', tick_format='0.0f', label='Security 2')
Figure(marks=[scatt], axes=[ax_x, ax_y])
```
## Changing the marker and adding text to each point of the scatter
```
# Changing the marker as
sc_x = LinearScale()
sc_y = LinearScale()
scatt = Scatter(x=x_data[:10], y=y_data[:10], names=np.arange(10),
scales={'x': sc_x, 'y': sc_y}, colors=['red'], marker='cross')
ax_x = Axis(scale=sc_x)
ax_y = Axis(scale=sc_y, orientation='vertical', tick_format='0.2f')
Figure(marks=[scatt], axes=[ax_x, ax_y], padding_x=0.025)
```
## Changing the opacity of each marker
```
scatt.opacities = [0.3, 0.5, 1.]
```
# Representing additional dimensions of data
## Linear Scale for Color Data
```
sc_x = DateScale()
sc_y = LinearScale()
sc_c1 = ColorScale()
scatter = Scatter(x=dates_all, y=sec2_levels, color=sec1_returns,
scales={'x': sc_x, 'y': sc_y, 'color': sc_c1},
stroke='black')
ax_y = Axis(label='Security 2', scale=sc_y,
orientation='vertical', side='left')
ax_x = Axis(label='Date', scale=sc_x, num_ticks=10, label_location='end')
ax_c = ColorAxis(scale=sc_c1, tick_format='0.2%', label='Returns', orientation='vertical', side='right')
m_chart = dict(top=50, bottom=70, left=50, right=100)
Figure(axes=[ax_x, ax_c, ax_y], marks=[scatter], fig_margin=m_chart,
title='Scatter of Security 2 vs Dates')
## Changing the default color.
scatter.colors = ['blue'] # In this case, the dot with the highest X changes to blue.
## setting the fill to be empty
scatter.stroke = None
scatter.fill = False
## Setting the fill back
scatter.stroke = 'black'
scatter.fill = True
## Changing the color to a different variable
scatter.color = sec2_levels
ax_c.tick_format = '0.0f'
ax_c.label = 'Security 2'
## Changing the range of the color scale
sc_c1.colors = ['blue', 'green', 'orange']
```
## Date Scale for Color Data
```
sc_x = LinearScale()
sc_y = LinearScale()
sc_c1 = DateColorScale(scheme='Reds')
scatter = Scatter(x=sec2_levels, y=sec1_levels, color=dates_all,
scales={'x': sc_x, 'y': sc_y, 'color': sc_c1}, default_size=128,
stroke='black')
ax_y = Axis(label='Security 1 Level', scale=sc_y, orientation='vertical', side='left')
ax_x = Axis(label='Security 2', scale=sc_x)
ax_c = ColorAxis(scale=sc_c1, label='Date', num_ticks=5)
m_chart = dict(top=50, bottom=80, left=50, right=50)
Figure(axes=[ax_x, ax_c, ax_y], marks=[scatter], fig_margin=m_chart)
```
## Ordinal Scale for Color
```
factor = int(np.ceil(len(sec2_levels) * 1.0 / len(ordinal_data)))
ordinal_data = np.tile(ordinal_data, factor)
c_ord = OrdinalColorScale(colors=['DodgerBlue', 'SeaGreen', 'Yellow', 'HotPink', 'OrangeRed'])
sc_x = LinearScale()
sc_y = LinearScale()
scatter2 = Scatter(x=sec2_levels[1:],
y=sec1_returns,
color=ordinal_data,
scales={'x': sc_x, 'y': sc_y, 'color': c_ord},
legend='__no_legend__',
stroke='black')
ax_y = Axis(label='Security 1 Returns', scale=sc_y, orientation='vertical', tick_format='.0%')
ax_x = Axis(label='Security 2', scale=sc_x, label_location='end')
ax_c = ColorAxis(scale=c_ord, label='Class', side='right', orientation='vertical')
m_chart = dict(top=50, bottom=70, left=100, right=100)
Figure(axes=[ax_x, ax_y, ax_c], marks=[scatter2], fig_margin=m_chart)
ax_c.tick_format = '0.2f'
c_ord.colors = ['blue', 'red', 'green', 'yellow', 'orange']
```
## Setting size and opacity based on data
```
sc_x = LinearScale()
sc_y = LinearScale()
sc_y2 = LinearScale()
sc_size = LinearScale()
sc_opacity = LinearScale()
scatter2 = Scatter(x=sec2_levels[1:], y=sec1_levels, size=sec1_returns,
scales={'x': sc_x, 'y': sc_y, 'size': sc_size, 'opacity': sc_opacity},
default_size=128, colors=['orangered'], stroke='black')
ax_y = Axis(label='Security 1', scale=sc_y, orientation='vertical', side='left')
ax_x = Axis(label='Security 2', scale=sc_x)
Figure(axes=[ax_x, ax_y], marks=[scatter2])
## Changing the opacity of the scatter
scatter2.opacities = [0.5, 0.3, 0.1]
## Resetting the size for the scatter
scatter2.size=None
## Resetting the opacity and setting the opacity according to the date
scatter2.opacities = [1.0]
scatter2.opacity = dates_all
```
## Changing the skew of the marker
```
sc_x = LinearScale()
sc_y = LinearScale()
sc_e = LinearScale()
scatter = Scatter(scales={'x': sc_x, 'y': sc_y, 'skew': sc_e},
x=sec2_levels[1:], y=sec1_levels,
skew=sec1_returns, stroke="black",
colors=['gold'], default_size=200,
marker='rectangle', default_skew=0)
ax_y = Axis(label='Security 1', scale=sc_y, orientation='vertical', side='left')
ax_x = Axis(label='Security 2', scale=sc_x)
Figure(axes=[ax_x, ax_y], marks=[scatter], animation_duration=1000)
scatter.skew = None
scatter.skew = sec1_returns
```
## Rotation scale
```
sc_x = LinearScale()
sc_y = LinearScale()
sc_e = LinearScale()
sc_c = ColorScale(scheme='Reds')
x1 = np.linspace(-1, 1, 30)
y1 = np.linspace(-1, 1, 30)
x, y = np.meshgrid(x1,y1)
x, y = x.flatten(), y.flatten()
rot = x**2 + y**2
color=x-y
scatter = Scatter(scales={'x': sc_x, 'y': sc_y, 'color': sc_c, 'rotation': sc_e},
x=x, y=y, rotation=rot, color=color,
stroke="black", default_size=200,
marker='arrow', default_skew=0.5,)
Figure(marks=[scatter], animation_duration=1000)
scatter.rotation = 1.0 / (x ** 2 + y ** 2 + 1)
```
# Scatter Chart Interactions
## Moving points in Scatter
```
## Enabling moving of points in scatter. Try to click and drag any of the points in the scatter and
## notice the line representing the mean of the data update
sc_x = LinearScale()
sc_y = LinearScale()
scat = Scatter(x=x_data[:10], y=y_data[:10], scales={'x': sc_x, 'y': sc_y}, colors=['orange'],
enable_move=True)
lin = Lines(x=[], y=[], scales={'x': sc_x, 'y': sc_y}, line_style='dotted', colors=['orange'])
def update_line(change=None):
with lin.hold_sync():
lin.x = [np.min(scat.x), np.max(scat.x)]
lin.y = [np.mean(scat.y), np.mean(scat.y)]
update_line()
# update line on change of x or y of scatter
scat.observe(update_line, names=['x'])
scat.observe(update_line, names=['y'])
ax_x = Axis(scale=sc_x)
ax_y = Axis(scale=sc_y, tick_format='0.2f', orientation='vertical')
fig = Figure(marks=[scat, lin], axes=[ax_x, ax_y])
fig
```
### Updating X and Y while moving the point
```
## In this case on drag, the line updates as you move the points.
scat.update_on_move = True
latex_widget = Label(color='Green', font_size='16px')
def callback_help(name, value):
latex_widget.value = str(value)
latex_widget
scat.on_drag_start(callback_help)
scat.on_drag(callback_help)
scat.on_drag_end(callback_help)
## Restricting movement to only along the Y-axis
scat.restrict_y = True
```
## Adding/Deleting points
```
## Enabling adding the points to Scatter. Try clicking anywhere on the scatter to add points
with scat.hold_sync():
scat.enable_move = False
scat.interactions = {'click': 'add'}
```
## Switching between interactions
```
from ipywidgets import ToggleButtons, VBox
interact_control = ToggleButtons(options=['Add', 'Delete', 'Drag XY', 'Drag X', 'Drag Y'],
style={'button_width': '120px'})
def change_interact(change):
interact_parameters = {
'Add': {'interactions': {'click': 'add'},
'enable_move': False},
'Delete': {'interactions': {'click': 'delete'},
'enable_move': False},
'Drag XY': {'interactions': {'click': None},
'enable_move': True,
'restrict_x': False,
'restrict_y': False},
'Drag X': {'interactions': {'click': None},
'enable_move': True,
'restrict_x': True,
'restrict_y': False},
'Drag Y': {'interactions': {'click': None},
'enable_move': True,
'restrict_x': False,
'restrict_y': True}
}
for param, value in interact_parameters[interact_control.value].items():
setattr(scat, param, value)
interact_control.observe(change_interact, names='value')
fig.title = 'Adding/Deleting/Moving points'
VBox([fig, interact_control])
```
## Custom event on end of drag
```
## Whenever drag is ended, there is a custom event dispatched which can be listened to.
## try dragging a point and see the data associated with the event being printed
def test_func(self, content):
print("received drag end", content)
scat.on_drag_end(test_func)
```
## Adding tooltip and custom hover style
```
x_sc = LinearScale()
y_sc = LinearScale()
x_data = x_data[:50]
y_data = y_data[:50]
def_tt = Tooltip(fields=['x', 'y'], formats=['', '.2f'])
scatter_chart = Scatter(x=x_data, y=y_data, scales= {'x': x_sc, 'y': y_sc}, colors=['dodgerblue'],
tooltip=def_tt, unhovered_style={'opacity': 0.5})
ax_x = Axis(scale=x_sc)
ax_y = Axis(scale=y_sc, orientation='vertical', tick_format='0.2f')
Figure(marks=[scatter_chart], axes=[ax_x, ax_y])
## removing field names from the tooltip
def_tt.show_labels = False
## changing the fields displayed in the tooltip
def_tt.fields = ['y']
def_tt.fields = ['x']
```
| github_jupyter |
# Explorando Datasets - London Bike Sharing
Este documento tiene como propósito llevar a cabo los pasos durante un análisis de datos rutinario en un dataset como el London Bike Sharing.
Fuente: [Kaggle - Data Analysis](https://www.kaggle.com/hmavrodiev/london-bike-sharing-dataset)
## Metadata
El data set está compuesto por la siguientes columnas:
- "timestamp" - Campo de registro de fecha
- "cnt" - La cuenta de bicicletas compartidas
- "t1" - Temperatura real en grados Celsius
- "t2" - Temperatura percibida en Celsius
- "hum" - Porcentaje de humedad relativa
- "windspeed" - Velocidad del viento km/h
- "weathercode" - Categoría del clima
- "isholiday" - boolean - 1 Festivo / 0 no Festivo
- "isweekend" - boolean - 1 Si es en semana
- "season" - Categorías meteorologicas de cada temporada: 0-primavera ; 1-verano; 2-otoño; 3-invierno.
#### "weather_code" Descripción de las categorías del clima:
- 1 = Clear ; mostly clear but have some values with haze/fog/patches of fog/ fog in vicinity
- 2 = scattered clouds / few clouds
- 3 = Broken clouds
- 4 = Cloudy
- 7 = Rain/ light Rain shower/ Light rain
- 10 = rain with thunderstorm
- 26 = snowfall
- 94 = Freezing Fog
## Objetivo:
1. Cargar mi dataset
2. Identificar los tipos de variables
3. Si encuentro columnas que son tipo objeto y deberían ser formato fecha uso las funciones de pandas para dicha labor.
```
!pip install --upgrade pandas
import pandas as pd
import numpy as np
pd.__version__
```
Cuando trabajo con repositorio Local de información, cargo mis datos así:
```
df_lmerged = pd.read_csv('../../Data/london_merged.csv')
df_lmerged
```
Explorar los tipos de dato.
```
df_lmerged.dtypes
```
## Parseando Fechas.
Tranforma el tipo de dato de la columna timestamp de objeto a fecha. ¿Por qué es importante parsear la columna de tipo objeto a tipo fecha?.
```
df_lmerged['timestamp'] = pd.to_datetime(df_lmerged['timestamp'])
df_lmerged['timestamp']
df_lmerged.dtypes
```
Cuando la columna ya tiene formato tipo `datetime` puedo usar método de `.hour` para crear una columna de horas, porque puedo acceder a esta así como minutos, segundos.
```
df_lmerged['hour'] = df_lmerged['timestamp'].dt.hour
df_lmerged['hour']
```
## Creando Dataframe de trabajo.
Aqui lo que hice fue que eliminé la columna de tiempo con el propósito de posteriomente realizar operaciones matemáticas sobre un dataframe. Mediante `iloc` dice al dataframe que elimine `:` todas las filas y que me guarde desde la `1:` hasta el final.
```
df = df_lmerged.iloc[:, 1:]
df
# Si intento hacer una operación de un subset de un dataframe con una columna del dataframe.
# Tendré valores no numéricos.
df['t1'].iloc[::3]-df['t2']
# Para solucionar lo anterior puedo usar la función sub() que me resta y adicionalmente me permite usar el argumento fill_value, para rellenar ese NaN
df['t1'].iloc[::3].sub(df['t2'], fill_value = 1000)
```
```
+ add()
- sub(), subtract()
* mul(), multiply()
/ truediv(), div(), divide()
// floordiv()
% mod()
** pow()
```
```
# Puedo usar divisiones en columnas, al encontrarse con inf dice que se encontró sobre una cantidad infonita.
df['t1']/df['t2']
# La función dot()... me permite hacer ua multiplicación de matrices entre dataframes.
df['t1'].dot(df['t1'])
# Mas ejemplos sobre la función dot.
# df = pd.DataFrame([[0, 1, -2, -1], [1, 1, 1, 1]])
# s = pd.Series([1, 1, 2, 1])
# df.dot(s)
```
### Funciones más complejas y lambdas
```
def fun_1(x):
y = x**2 + 1
return y
fun_1(10)
np.arange(-5,6)
np.arange(-5,6).shape
fun_1(np.arange(-5,6))
df
```
Funciones apply() Se usan en filas o columnas
```
# Si quiero aplicar a mi dataframe, especificamente a mis series de horas del dataframe toda la función que recién escribí.x**2+1
df['hour'].apply(fun_1)
def fun_2(x, a=1, b=0):
y = x**2 + a*x + b
return y
fun_2(10, a = 20, b= -100)
# Si la función que quiero aplciar a mi dataframe posee argumentos me valgo de la función args para decirle cuales serán. Cabe aclarar que los argumentos que está usando
# son opcionales desde que sean llamados
df['hour'].apply(fun_2, args = (20, -100))
df['hour'].apply(fun_2, a =20, b= -100)
# Una forma mucho más rápida para no tener que definir la función es usar lambda.
df['hour'].apply(lambda x: x**2 + 1 )
# Una forma mucho más rápida para no tener que definir la función es usar lambda. Paso temperatura a Kelvin sumando mas 273
df['t1'].apply(lambda x: x+273)
# Puedo aplicar una función a todo el dataframe haciendo un promedio.
df.apply(lambda x: x.mean())
# Así le puedo escpecificar las columnas mediante ejes.
df.apply(lambda x: x.mean(), axis=1)
# Puedo usar tambien para desviaciones estaándara
df.apply(lambda x: x.std(), axis=1)
df.apply(lambda x: x['t1']-x['t2'], axis=1)
# Para aplciar un cambio a todo el mapa frame me permite operar a todos los valores individualmente.
df.applymap(lambda x: x/1000)
```
| github_jupyter |
```
%matplotlib inline
```
# Plot a confidence ellipse of a two-dimensional dataset
This example shows how to plot a confidence ellipse of a
two-dimensional dataset, using its pearson correlation coefficient.
The approach that is used to obtain the correct geometry is
explained and proved here:
https://carstenschelp.github.io/2018/09/14/Plot_Confidence_Ellipse_001.html
The method avoids the use of an iterative eigen decomposition algorithm
and makes use of the fact that a normalized covariance matrix (composed of
pearson correlation coefficients and ones) is particularly easy to handle.
```
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.patches import Ellipse
import matplotlib.transforms as transforms
```
The plotting function itself
""""""""""""""""""""""""""""
This function plots the confidence ellipse of the covariance of the given
array-like variables x and y. The ellipse is plotted into the given
axes-object ax.
The radiuses of the ellipse can be controlled by n_std which is the number
of standard deviations. The default value is 3 which makes the ellipse
enclose 99.7% of the points (given the data is normally distributed
like in these examples).
```
def confidence_ellipse(x, y, ax, n_std=3.0, facecolor='none', **kwargs):
"""
Create a plot of the covariance confidence ellipse of *x* and *y*.
Parameters
----------
x, y : array-like, shape (n, )
Input data.
ax : matplotlib.axes.Axes
The axes object to draw the ellipse into.
n_std : float
The number of standard deviations to determine the ellipse's radiuses.
Returns
-------
matplotlib.patches.Ellipse
Other parameters
----------------
kwargs : `~matplotlib.patches.Patch` properties
"""
if x.size != y.size:
raise ValueError("x and y must be the same size")
cov = np.cov(x, y)
pearson = cov[0, 1]/np.sqrt(cov[0, 0] * cov[1, 1])
# Using a special case to obtain the eigenvalues of this
# two-dimensionl dataset.
ell_radius_x = np.sqrt(1 + pearson)
ell_radius_y = np.sqrt(1 - pearson)
ellipse = Ellipse((0, 0),
width=ell_radius_x * 2,
height=ell_radius_y * 2,
facecolor=facecolor,
**kwargs)
# Calculating the stdandard deviation of x from
# the squareroot of the variance and multiplying
# with the given number of standard deviations.
scale_x = np.sqrt(cov[0, 0]) * n_std
mean_x = np.mean(x)
# calculating the stdandard deviation of y ...
scale_y = np.sqrt(cov[1, 1]) * n_std
mean_y = np.mean(y)
transf = transforms.Affine2D() \
.rotate_deg(45) \
.scale(scale_x, scale_y) \
.translate(mean_x, mean_y)
ellipse.set_transform(transf + ax.transData)
return ax.add_patch(ellipse)
```
A helper function to create a correlated dataset
""""""""""""""""""""""""""""""""""""""""""""""""
Creates a random two-dimesional dataset with the specified
two-dimensional mean (mu) and dimensions (scale).
The correlation can be controlled by the param 'dependency',
a 2x2 matrix.
```
def get_correlated_dataset(n, dependency, mu, scale):
latent = np.random.randn(n, 2)
dependent = latent.dot(dependency)
scaled = dependent * scale
scaled_with_offset = scaled + mu
# return x and y of the new, correlated dataset
return scaled_with_offset[:, 0], scaled_with_offset[:, 1]
```
Positive, negative and weak correlation
"""""""""""""""""""""""""""""""""""""""
Note that the shape for the weak correlation (right) is an ellipse,
not a circle because x and y are differently scaled.
However, the fact that x and y are uncorrelated is shown by
the axes of the ellipse being aligned with the x- and y-axis
of the coordinate system.
```
np.random.seed(0)
PARAMETERS = {
'Positive correlation': np.array([[0.85, 0.35],
[0.15, -0.65]]),
'Negative correlation': np.array([[0.9, -0.4],
[0.1, -0.6]]),
'Weak correlation': np.array([[1, 0],
[0, 1]]),
}
mu = 2, 4
scale = 3, 5
fig, axs = plt.subplots(1, 3, figsize=(9, 3))
for ax, (title, dependency) in zip(axs, PARAMETERS.items()):
x, y = get_correlated_dataset(800, dependency, mu, scale)
ax.scatter(x, y, s=0.5)
ax.axvline(c='grey', lw=1)
ax.axhline(c='grey', lw=1)
confidence_ellipse(x, y, ax, edgecolor='red')
ax.scatter(mu[0], mu[1], c='red', s=3)
ax.set_title(title)
plt.show()
```
Different number of standard deviations
"""""""""""""""""""""""""""""""""""""""
A plot with n_std = 3 (blue), 2 (purple) and 1 (red)
```
fig, ax_nstd = plt.subplots(figsize=(6, 6))
dependency_nstd = np.array([
[0.8, 0.75],
[-0.2, 0.35]
])
mu = 0, 0
scale = 8, 5
ax_nstd.axvline(c='grey', lw=1)
ax_nstd.axhline(c='grey', lw=1)
x, y = get_correlated_dataset(500, dependency_nstd, mu, scale)
ax_nstd.scatter(x, y, s=0.5)
confidence_ellipse(x, y, ax_nstd, n_std=1,
label=r'$1\sigma$', edgecolor='firebrick')
confidence_ellipse(x, y, ax_nstd, n_std=2,
label=r'$2\sigma$', edgecolor='fuchsia', linestyle='--')
confidence_ellipse(x, y, ax_nstd, n_std=3,
label=r'$3\sigma$', edgecolor='blue', linestyle=':')
ax_nstd.scatter(mu[0], mu[1], c='red', s=3)
ax_nstd.set_title('Different standard deviations')
ax_nstd.legend()
plt.show()
```
Using the keyword arguments
"""""""""""""""""""""""""""
Use the kwargs specified for matplotlib.patches.Patch in order
to have the ellipse rendered in different ways.
```
fig, ax_kwargs = plt.subplots(figsize=(6, 6))
dependency_kwargs = np.array([
[-0.8, 0.5],
[-0.2, 0.5]
])
mu = 2, -3
scale = 6, 5
ax_kwargs.axvline(c='grey', lw=1)
ax_kwargs.axhline(c='grey', lw=1)
x, y = get_correlated_dataset(500, dependency_kwargs, mu, scale)
# Plot the ellipse with zorder=0 in order to demonstrate
# its transparency (caused by the use of alpha).
confidence_ellipse(x, y, ax_kwargs,
alpha=0.5, facecolor='pink', edgecolor='purple', zorder=0)
ax_kwargs.scatter(x, y, s=0.5)
ax_kwargs.scatter(mu[0], mu[1], c='red', s=3)
ax_kwargs.set_title(f'Using kwargs')
fig.subplots_adjust(hspace=0.25)
plt.show()
```
| github_jupyter |
```
import matplotlib.pyplot as plt
import numpy as np
import copy
from qiskit import QuantumCircuit, QuantumRegister, ClassicalRegister, assemble
from collections.abc import Iterable
```
## A-gen-prep Circuit Verification
```
num_q = 3
qr_ = QuantumRegister(num_q+1)
cr_ = ClassicalRegister(num_q+1)
qc_ = QuantumCircuit(qr_, cr_)
a = 1/8
A = A_gen(num_q, a)
_, Q = Ctrl_Q(num_q, A)
qc_.append(A, qr_)
# qc_.append(Q, qr_)
qc_.measure(qr_, cr_)
from qiskit import execute, Aer
backend = Aer.get_backend("qasm_simulator")
job = execute(qc_, backend, shots=1000)
result = job.result()
counts = result.get_counts()
print(counts)
# Construct A operator that takes |0>_{n+1} to sqrt(1-a) |psi_0>|0> + sqrt(a) |psi_1>|1>
def A_gen(num_state_qubits, a, psi_zero=None, psi_one=None):
if psi_zero==None:
psi_zero = '0'*num_state_qubits
if psi_one==None:
psi_one = '1'*num_state_qubits
theta = 2 * np.arcsin(np.sqrt(a))
# Let the objective be qubit index n; state is on qubits 0 through n-1
qc_A = QuantumCircuit(num_state_qubits+1, name=f"A")
qc_A.ry(theta, num_state_qubits)
qc_A.x(num_state_qubits)
for i in range(num_state_qubits):
if psi_zero[i]=='1':
qc_A.cnot(num_state_qubits,i)
qc_A.x(num_state_qubits)
for i in range(num_state_qubits):
if psi_one[i]=='1':
qc_A.cnot(num_state_qubits,i)
return qc_A
```
## Ctrl_Q Validations
```
num_q = 1
qr_ = QuantumRegister(num_q+1)
cr_ = ClassicalRegister(num_q+1)
qc_ = QuantumCircuit(qr_, cr_)
a = 1/5
A = A_gen(num_q, a)
_, Q = Ctrl_Q(num_q, A)
qc_.append(Q, qr_)
display(qc_.draw())
qc_ = qc_.decompose().decompose()
usim = Aer.get_backend('unitary_simulator')
qobj = assemble(qc_)
unitary = usim.run(qobj).result().get_unitary()
print(np.matrix.round(unitary, 6))
# Construct the grover-like operator and a controlled version of it
def Ctrl_Q(num_state_qubits, A_circ):
# index n is the objective qubit, and indexes 0 through n-1 are state qubits
qc = QuantumCircuit(num_state_qubits+1, name=f"Q")
temp_A = copy.copy(A_circ)
A_gate = temp_A.to_gate()
A_gate_inv = temp_A.inverse().to_gate()
### Each cycle in Q applies in order: S_chi, A_circ_inverse, S_0, A_circ
# S_chi
qc.z(num_state_qubits)
# A_circ_inverse
qc.append(A_gate_inv, [i for i in range(num_state_qubits+1)])
# S_0
for i in range(num_state_qubits+1):
qc.x(i)
qc.h(num_state_qubits)
qc.mcx([x for x in range(num_state_qubits)], num_state_qubits)
qc.h(num_state_qubits)
for i in range(num_state_qubits+1):
qc.x(i)
# A_circ
qc.append(A_gate, [i for i in range(num_state_qubits+1)])
# add "global" phase
qc.x(num_state_qubits)
qc.z(num_state_qubits)
qc.x(num_state_qubits)
qc.z(num_state_qubits)
# Create a gate out of the Q operator
qc.to_gate(label='Q')
# and also a controlled version of it
Ctrl_Q_ = qc.control(1)
# and return both
return Ctrl_Q_, qc
```
| github_jupyter |
```
# Part of the LLVM Project, under the Apache License v2.0 with LLVM Exceptions.
# See https://llvm.org/LICENSE.txt for license information.
# SPDX-License-Identifier: Apache-2.0 WITH LLVM-exception
# Also available under a BSD-style license. See LICENSE.
```
## Setup
### Configuring Jupyter kernel.
We assume that you have followed the instructions for setting up Torch-MLIR. See [README.md](https://github.com/llvm/torch-mlir) if not.
To run this notebook, you need to configure Jupyter to access the Torch-MLIR Python modules that are built as part of your development setup. An easy way to do this is to run the following command with the same Python (and shell) that is correctly set up and able to run the Torch-MLIR end-to-end tests with RefBackend:
```shell
python -m ipykernel install --user --name=torch-mlir --env PYTHONPATH "$PYTHONPATH"
```
You should then have an option in Jupyter to select this kernel for running this notebook.
**TODO**: Make this notebook standalone and work based entirely on pip-installable packages.
### Additional dependencies for this notebook
```
!python -m pip install requests pillow
```
## Imports
### torch-mlir imports
```
import torch
import torchvision
import torch_mlir
from torch_mlir.dialects.torch.importer.jit_ir import ClassAnnotator, ModuleBuilder
from torch_mlir.dialects.torch.importer.jit_ir.torchscript_annotations import extract_annotations
from torch_mlir.passmanager import PassManager
from torch_mlir_e2e_test.linalg_on_tensors_backends.refbackend import RefBackendLinalgOnTensorsBackend
```
### General dependencies
```
import requests
from PIL import Image
```
### Utilities
```
def compile_and_load_on_refbackend(module):
"""Compile an MLIR Module to an executable module.
This uses the Torch-MLIR reference backend which accepts
linalg-on-tensors as the way to express tensor computations.
"""
backend = RefBackendLinalgOnTensorsBackend()
compiled = backend.compile(module)
return backend.load(compiled)
```
## Basic tanh module
A simple tiny module that is easier to understand and look at than a full ResNet.
```
class TanhModule(torch.nn.Module):
def forward(self, a):
return torch.tanh(a)
# Compile the model with an example input.
# We lower to the linalg-on-tensors form that the reference backend supports.
compiled = torch_mlir.compile(TanhModule(), torch.ones(3), output_type=torch_mlir.OutputType.LINALG_ON_TENSORS)
# Load it on the reference backend.
jit_module = compile_and_load_on_refbackend(compiled)
# Run it!
jit_module.forward(torch.tensor([-1.0, 1.0, 0.0]).numpy())
```
## ResNet Inference
Do some one-time preparation.
```
def _load_labels():
classes_text = requests.get(
"https://raw.githubusercontent.com/cathyzhyi/ml-data/main/imagenet-classes.txt",
stream=True,
).text
labels = [line.strip() for line in classes_text.splitlines()]
return labels
IMAGENET_LABELS = _load_labels()
def _get_preprocess_transforms():
# See preprocessing specification at: https://pytorch.org/vision/stable/models.html
T = torchvision.transforms
return T.Compose(
[
T.Resize(256),
T.CenterCrop(224),
T.ToTensor(),
T.Normalize(mean=[0.485, 0.456, 0.406], std=[0.229, 0.224, 0.225]),
]
)
PREPROCESS_TRANSFORMS = _get_preprocess_transforms()
```
Define some helper functions.
```
def fetch_image(url: str):
# Use some "realistic" User-Agent so that we aren't mistaken for being a scraper.
headers = {"User-Agent": "Mozilla/5.0 (X11; Linux x86_64) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/92.0.4515.131 Safari/537.36"}
return Image.open(requests.get(url, headers=headers, stream=True).raw).convert("RGB")
def preprocess_image(img: Image):
# Preprocess and add a batch dimension.
return torch.unsqueeze(PREPROCESS_TRANSFORMS(img), 0)
```
### Fetch our sample image.
```
img = fetch_image("https://upload.wikimedia.org/wikipedia/commons/thumb/3/31/Red_Smooth_Saluki.jpg/590px-Red_Smooth_Saluki.jpg")
img_preprocessed = preprocess_image(img)
img
```
### Define the module and compile it
```
resnet18 = torchvision.models.resnet18(weights=torchvision.models.ResNet18_Weights.DEFAULT)
resnet18.eval()
compiled = torch_mlir.compile(resnet18, torch.ones(1, 3, 224, 224), output_type=torch_mlir.OutputType.LINALG_ON_TENSORS)
jit_module = compile_and_load_on_refbackend(compiled)
```
### Execute the classification!
```
logits = torch.from_numpy(jit_module.forward(img_preprocessed.numpy()))
# Torch-MLIR doesn't currently support these final postprocessing operations, so perform them in Torch.
def top3_possibilities(logits):
_, indexes = torch.sort(logits, descending=True)
percentage = torch.nn.functional.softmax(logits, dim=1)[0] * 100
top3 = [(IMAGENET_LABELS[idx], percentage[idx].item()) for idx in indexes[0][:3]]
return top3
top3_possibilities(logits)
```
| github_jupyter |
# Week 2 Assignment: Zombie Detection
Welcome to this week's programming assignment! You will use the Object Detection API and retrain [RetinaNet](https://arxiv.org/abs/1708.02002) to spot Zombies using just 5 training images. You will setup the model to restore pretrained weights and fine tune the classification layers.
***Important:*** *This colab notebook has read-only access so you won't be able to save your changes. If you want to save your work periodically, please click `File -> Save a Copy in Drive` to create a copy in your account, then work from there.*
<img src='https://drive.google.com/uc?export=view&id=18Ck0qNSZy9F1KsUKWc4Jv7_x_1e_fXTN' alt='zombie'>
## Exercises
* [Exercise 1 - Import Object Detection API packages](#exercise-1)
* [Exercise 2 - Visualize the training images](#exercise-2)
* [Exercise 3 - Define the category index dictionary](#exercise-3)
* [Exercise 4 - Download checkpoints](#exercise-4)
* [Exercise 5.1 - Locate and read from the configuration file](#exercise-5-1)
* [Exercise 5.2 - Modify the model configuration](#exercise-5-2)
* [Exercise 5.3 - Modify model_config](#exercise-5-3)
* [Exercise 5.4 - Build the custom model](#exercise-5-4)
* [Exercise 6.1 - Define Checkpoints for the box predictor](#exercise-6-1)
* [Exercise 6.2 - Define the temporary model checkpoint](#exercise-6-2)
* [Exercise 6.3 - Restore the checkpoint](#exercise-6-2)
* [Exercise 7 - Run a dummy image to generate the model variables](#exercise-7)
* [Exercise 8 - Set training hyperparameters](#exercise-8)
* [Exercise 9 - Select the prediction layer variables](#exercise-9)
* [Exercise 10 - Define the training step](#exercise-10)
* [Exercise 11 - Preprocess, predict, and post process an image](#exercise-11)
## Installation
You'll start by installing the Tensorflow 2 [Object Detection API](https://github.com/tensorflow/models/tree/master/research/object_detection).
```
# uncomment the next line if you want to delete an existing models directory
!rm -rf ./models/
# clone the Tensorflow Model Garden
!git clone --depth 1 https://github.com/tensorflow/models/
# install the Object Detection API
!cd models/research/ && protoc object_detection/protos/*.proto --python_out=. && cp object_detection/packages/tf2/setup.py . && python -m pip install .
```
## Imports
Let's now import the packages you will use in this assignment.
```
import matplotlib
import matplotlib.pyplot as plt
import os
import random
import zipfile
import io
import scipy.misc
import numpy as np
import glob
import imageio
from six import BytesIO
from PIL import Image, ImageDraw, ImageFont
from IPython.display import display, Javascript
from IPython.display import Image as IPyImage
try:
# %tensorflow_version only exists in Colab.
%tensorflow_version 2.x
except Exception:
pass
import tensorflow as tf
tf.get_logger().setLevel('ERROR')
```
<a name='exercise-1'></a>
### **Exercise 1**: Import Object Detection API packages
Import the necessary modules from the `object_detection` package.
- From the [utils](https://github.com/tensorflow/models/tree/master/research/object_detection/utils) package:
- [label_map_util](https://github.com/tensorflow/models/blob/master/research/object_detection/utils/label_map_util.py)
- [config_util](https://github.com/tensorflow/models/blob/master/research/object_detection/utils/config_util.py): You'll use this to read model configurations from a .config file and then modify that configuration
- [visualization_utils](https://github.com/tensorflow/models/blob/master/research/object_detection/utils/visualization_utils.py): please give this the alias `viz_utils`, as this is what will be used in some visualization code that is given to you later.
- [colab_utils](https://github.com/tensorflow/models/blob/master/research/object_detection/utils/colab_utils.py)
- From the [builders](https://github.com/tensorflow/models/tree/master/research/object_detection/builders) package:
- [model_builder](https://github.com/tensorflow/models/blob/master/research/object_detection/builders/model_builder.py): This builds your model according to the model configuration that you'll specify.
```
### START CODE HERE (Replace Instances of `None` with your code) ###
# import the label map utility module
from object_detection.utils import label_map_util
# import module for reading and updating configuration files.
from object_detection.utils import config_util
# import module for visualization. use the alias `viz_utils`
from object_detection.utils import visualization_utils as viz_utils
# import module for building the detection model
from object_detection.builders import model_builder
### END CODE HERE ###
# import module for utilities in Colab
from object_detection.utils import colab_utils
```
## Utilities
You'll define a couple of utility functions for loading images and plotting detections. This code is provided for you.
```
def load_image_into_numpy_array(path):
"""Load an image from file into a numpy array.
Puts image into numpy array to feed into tensorflow graph.
Note that by convention we put it into a numpy array with shape
(height, width, channels), where channels=3 for RGB.
Args:
path: a file path.
Returns:
uint8 numpy array with shape (img_height, img_width, 3)
"""
img_data = tf.io.gfile.GFile(path, 'rb').read()
image = Image.open(BytesIO(img_data))
(im_width, im_height) = image.size
return np.array(image.getdata()).reshape(
(im_height, im_width, 3)).astype(np.uint8)
def plot_detections(image_np,
boxes,
classes,
scores,
category_index,
figsize=(12, 16),
image_name=None):
"""Wrapper function to visualize detections.
Args:
image_np: uint8 numpy array with shape (img_height, img_width, 3)
boxes: a numpy array of shape [N, 4]
classes: a numpy array of shape [N]. Note that class indices are 1-based,
and match the keys in the label map.
scores: a numpy array of shape [N] or None. If scores=None, then
this function assumes that the boxes to be plotted are groundtruth
boxes and plot all boxes as black with no classes or scores.
category_index: a dict containing category dictionaries (each holding
category index `id` and category name `name`) keyed by category indices.
figsize: size for the figure.
image_name: a name for the image file.
"""
image_np_with_annotations = image_np.copy()
viz_utils.visualize_boxes_and_labels_on_image_array(
image_np_with_annotations,
boxes,
classes,
scores,
category_index,
use_normalized_coordinates=True,
min_score_thresh=0.8)
if image_name:
plt.imsave(image_name, image_np_with_annotations)
else:
plt.imshow(image_np_with_annotations)
```
## Download the Zombie data
Now you will get 5 images of zombies that you will use for training.
- The zombies are hosted in a Google bucket.
- You can download and unzip the images into a local `training/` directory by running the cell below.
```
# uncomment the next 2 lines if you want to delete an existing zip and training directory
# !rm training-zombie.zip
# !rm -rf ./training
# download the images
!wget --no-check-certificate \
https://storage.googleapis.com/laurencemoroney-blog.appspot.com/training-zombie.zip \
-O ./training-zombie.zip
# unzip to a local directory
local_zip = './training-zombie.zip'
zip_ref = zipfile.ZipFile(local_zip, 'r')
zip_ref.extractall('./training')
zip_ref.close()
```
<a name='exercise-2'></a>
### **Exercise 2**: Visualize the training images
Next, you'll want to inspect the images that you just downloaded.
* Please replace instances of `None` below to load and visualize the 5 training images.
* You can inspect the *training* directory (using the `Files` button on the left side of this Colab) to see the filenames of the zombie images. The paths for the images will look like this:
```
./training/training-zombie1.jpg
./training/training-zombie2.jpg
./training/training-zombie3.jpg
./training/training-zombie4.jpg
./training/training-zombie5.jpg
```
- To set file paths, you'll use [os.path.join](https://www.geeksforgeeks.org/python-os-path-join-method/). As an example, if you wanted to create the path './parent_folder/file_name1.txt', you could write:
`os.path.join('parent_folder', 'file_name', str(1), '.txt')`
* You should see the 5 training images after running this cell. If not, please inspect your code, particularly the `image_path`.
```
%matplotlib inline
### START CODE HERE (Replace Instances of `None` with your code) ###
# assign the name (string) of the directory containing the training images
train_image_dir = './training'
# declare an empty list
train_images_np = []
# run a for loop for each image
for i in range(1, 6):
# define the path (string) for each image
image_path = os.path.join(train_image_dir, f'training-zombie{str(i)}.jpg')
print(image_path)
# load images into numpy arrays and append to a list
train_images_np.append(load_image_into_numpy_array(image_path))
### END CODE HERE ###
# configure plot settings via rcParams
plt.rcParams['axes.grid'] = False
plt.rcParams['xtick.labelsize'] = False
plt.rcParams['ytick.labelsize'] = False
plt.rcParams['xtick.top'] = False
plt.rcParams['xtick.bottom'] = False
plt.rcParams['ytick.left'] = False
plt.rcParams['ytick.right'] = False
plt.rcParams['figure.figsize'] = [14, 7]
# plot images
for idx, train_image_np in enumerate(train_images_np):
plt.subplot(1, 5, idx+1)
plt.imshow(train_image_np)
plt.show()
```
<a name='gt_boxes_definition'></a>
## Prepare data for training (Optional)
In this section, you will create your ground truth boxes. You can either draw your own boxes or use a prepopulated list of coordinates that we have provided below.
```
# Define the list of ground truth boxes
gt_boxes = []
```
#### Option 1: draw your own ground truth boxes
If you want to draw your own, please run the next cell and the following test code. If not, then skip these optional cells.
* Draw a box around the zombie in each image.
* Click the `next image` button to go to the next image
* Click `submit` when it says "All images completed!!".
- Make sure to not make the bounding box too big.
- If the box is too big, the model might learn the features of the background (e.g. door, road, etc) in determining if there is a zombie or not.
- Include the entire zombie inside the box.
- As an example, scroll to the beginning of this notebook to look at the bounding box around the zombie.
```
# Option 1: draw your own ground truth boxes
# annotate the training images
colab_utils.annotate(train_images_np, box_storage_pointer=gt_boxes)
# Option 1: draw your own ground truth boxes
# TEST CODE:
try:
assert(len(gt_boxes) == 5), "Warning: gt_boxes is empty. Did you click `submit`?"
except AssertionError as e:
print(e)
# checks if there are boxes for all 5 images
for gt_box in gt_boxes:
try:
assert(gt_box is not None), "There are less than 5 sets of box coordinates. " \
"Please re-run the cell above to draw the boxes again.\n" \
"Alternatively, you can run the next cell to load pre-determined " \
"ground truth boxes."
except AssertionError as e:
print(e)
break
ref_gt_boxes = [
np.array([[0.27333333, 0.41500586, 0.74333333, 0.57678781]]),
np.array([[0.29833333, 0.45955451, 0.75666667, 0.61078546]]),
np.array([[0.40833333, 0.18288394, 0.945, 0.34818288]]),
np.array([[0.16166667, 0.61899179, 0.8, 0.91910903]]),
np.array([[0.28833333, 0.12543962, 0.835, 0.35052755]]),
]
for gt_box, ref_gt_box in zip(gt_boxes, ref_gt_boxes):
try:
assert(np.allclose(gt_box, ref_gt_box, atol=0.04)), "One of the boxes is too big or too small. " \
"Please re-draw and make the box tighter around the zombie."
except AssertionError as e:
print(e)
break
```
<a name='gt-boxes'></a>
#### Option 2: use the given ground truth boxes
You can also use this list if you opt not to draw the boxes yourself.
```
# Option 2: use given ground truth boxes
# set this to `True` if you want to override the boxes you drew
override = False
# bounding boxes for each of the 5 zombies found in each image.
# you can use these instead of drawing the boxes yourself.
ref_gt_boxes = [
np.array([[0.27333333, 0.41500586, 0.74333333, 0.57678781]]),
np.array([[0.29833333, 0.45955451, 0.75666667, 0.61078546]]),
np.array([[0.40833333, 0.18288394, 0.945, 0.34818288]]),
np.array([[0.16166667, 0.61899179, 0.8, 0.91910903]]),
np.array([[0.28833333, 0.12543962, 0.835, 0.35052755]]),
]
# if gt_boxes is empty, use the reference
if not gt_boxes or override is True:
gt_boxes = ref_gt_boxes
# if gt_boxes does not contain 5 box coordinates, use the reference
for gt_box in gt_boxes:
try:
assert(gt_box is not None)
except:
gt_boxes = ref_gt_boxes
break
```
#### View your ground truth box coordinates
Whether you chose to draw your own or use the given boxes, please check your list of ground truth box coordinates.
```
# print the coordinates of your ground truth boxes
for gt_box in gt_boxes:
print(gt_box)
```
Below, we add the class annotations. For simplicity, we assume just a single class, though it should be straightforward to extend this to handle multiple classes. We will also convert everything to the format that the training loop expects (e.g., conversion to tensors, one-hot representations, etc.).
<a name='exercise-3'></a>
### **Exercise 3**: Define the category index dictionary
You'll need to tell the model which integer class ID to assign to the 'zombie' category, and what 'name' to associate with that integer id.
- zombie_class_id: By convention, class ID integers start numbering from 1,2,3, onward.
- If there is ever a 'background' class, it could be assigned the integer 0, but in this case, you're just predicting the one zombie class.
- Since you are just predicting one class (zombie), please assign `1` to the zombie class ID.
- category_index: Please define the `category_index` dictionary, which will have the same structure as this:
```
{human_class_id :
{'id' : human_class_id,
'name': 'human_so_far'}
}
```
- Define `category_index` similar to the example dictionary above, except for zombies.
- This will be used by the succeeding functions to know the class `id` and `name` of zombie images.
- num_classes: Since you are predicting one class, please assign `1` to the number of classes that the model will predict.
- This will be used during data preprocessing and again when you configure the model.
```
### START CODE HERE (Replace instances of `None` with your code ###
# Assign the zombie class ID
zombie_class_id = 1
# define a dictionary describing the zombie class
category_index = {
zombie_class_id: {
'id': zombie_class_id,
'name': 'zombie'
}
}
# Specify the number of classes that the model will predict
num_classes = 1
### END CODE HERE ###
# TEST CODE:
print(category_index[zombie_class_id])
```
**Expected Output:**
```txt
{'id': 1, 'name': 'zombie'}
```
### Data preprocessing
You will now do some data preprocessing so it is formatted properly before it is fed to the model:
- Convert the class labels to one-hot representations
- convert everything (i.e. train images, gt boxes and class labels) to tensors.
This code is provided for you.
```
# The `label_id_offset` here shifts all classes by a certain number of indices;
# we do this here so that the model receives one-hot labels where non-background
# classes start counting at the zeroth index. This is ordinarily just handled
# automatically in our training binaries, but we need to reproduce it here.
label_id_offset = 1
train_image_tensors = []
# lists containing the one-hot encoded classes and ground truth boxes
gt_classes_one_hot_tensors = []
gt_box_tensors = []
for (train_image_np, gt_box_np) in zip(train_images_np, gt_boxes):
# convert training image to tensor, add batch dimension, and add to list
train_image_tensors.append(tf.expand_dims(tf.convert_to_tensor(
train_image_np, dtype=tf.float32), axis=0))
# convert numpy array to tensor, then add to list
gt_box_tensors.append(tf.convert_to_tensor(gt_box_np, dtype=tf.float32))
# apply offset to to have zero-indexed ground truth classes
zero_indexed_groundtruth_classes = tf.convert_to_tensor(
np.ones(shape=[gt_box_np.shape[0]], dtype=np.int32) - label_id_offset)
# do one-hot encoding to ground truth classes
gt_classes_one_hot_tensors.append(tf.one_hot(
zero_indexed_groundtruth_classes, num_classes))
print('Done prepping data.')
```
## Visualize the zombies with their ground truth bounding boxes
You should see the 5 training images with the bounding boxes after running the cell below. If not, please re-run the [annotation tool](#gt_boxes_definition) again or use the prepopulated `gt_boxes` array given.
```
# give boxes a score of 100%
dummy_scores = np.array([1.0], dtype=np.float32)
# define the figure size
plt.figure(figsize=(30, 15))
# use the `plot_detections()` utility function to draw the ground truth boxes
for idx in range(5):
plt.subplot(2, 4, idx+1)
plot_detections(
train_images_np[idx],
gt_boxes[idx],
np.ones(shape=[gt_boxes[idx].shape[0]], dtype=np.int32),
dummy_scores, category_index)
plt.show()
```
## Download the checkpoint containing the pre-trained weights
Next, you will download [RetinaNet](https://arxiv.org/abs/1708.02002) and copy it inside the object detection directory.
When working with models that are at the frontiers of research, the models and checkpoints may not yet be organized in a central location like the TensorFlow Garden (https://github.com/tensorflow/models).
- You'll often read a blog post from the researchers, who will usually provide information on:
- how to use the model
- where to download the models and pre-trained checkpoints.
It's good practice to do some of this "detective work", so that you'll feel more comfortable when exploring new models yourself! So please try the following steps:
- Go to the [TensorFlow Blog](https://blog.tensorflow.org/), where researchers announce new findings.
- In the search box at the top of the page, search for "retinanet".
- In the search results, click on the blog post titled "TensorFlow 2 meets the Object Detection API" (it may be the first search result).
- Skim through this blog and look for links to either the checkpoints or to Colabs that will show you how to use the checkpoints.
- Try to fill out the following code cell below, which does the following:
- Download the compressed SSD Resnet 50 version 1, 640 x 640 checkpoint.
- Untar (decompress) the tar file
- Move the decompressed checkpoint to `models/research/object_detection/test_data/`
If you want some help getting started, please click on the "Initial Hints" cell to get some hints.
<details>
<summary>
<font size="3" color="darkgreen"><b>Initial Hints</b></font>
</summary>
<p>
General Hints to get started
<ul>
<li>The link to the blog is <a href="https://blog.tensorflow.org/2020/07/tensorflow-2-meets-object-detection-api.html">TensorFlow 2 meets the Object Detection API</a> </li>
<li>In the blog, you'll find the text "COCO pre-trained weights, which links to a list of checkpoints in GitHub titled
<a href="https://github.com/tensorflow/models/blob/master/research/object_detection/g3doc/tf2_detection_zoo.md">TensorFlow 2 Detection Model Zoo<a>.
</li>
<li>
If you read each checkpoint name, you'll find the one for SSD Resnet 50 version 1, 640 by 640. If you hover your mouse over
</li>
<li>
If you right-click on the desired checkpoint link, you can save the link address, and use it in the code cell below to get the checkpoint.
</li>
<li>For more hints, please click on the cell "More Hints"</li>
</ul>
</p>
<details>
<summary>
<font size="3" color="darkgreen"><b>More Hints</b></font>
</summary>
<p>
More Hints
<ul>
<li> To see how to download the checkpoint, look in the blog for links to Colab tutorials.
</li>
<li>
For example, the blog links to a Colab titled <a href="https://github.com/tensorflow/models/blob/master/research/object_detection/colab_tutorials/inference_tf2_colab.ipynb">Intro to Object Detection Colab</a>
</li>
<li>
In the Colab, you'll see the section titled "Build a detection model and load pre-trained model weights", which is followed by a code cell showing how to download, decompress, and relocate a checkpoint. Use similar syntax, except use the URL to the ssd resnet50 version 1 640x640 checkpoint instead.
</li>
<li> If you're feeling stuck, please click on the cell "Even More Hints".
</li>
</ul>
</p>
<details>
<summary>
<font size="3" color="darkgreen"><b>Even More Hints</b></font>
</summary>
<p>
Even More Hints
<ul>
<li> The blog post also links to a notebook titled
<a href="https://github.com/tensorflow/models/blob/master/research/object_detection/colab_tutorials/eager_few_shot_od_training_tf2_colab.ipynb">
Eager Few Shot Object Detection Colab</a>
</li>
<li> In this notebook, look for the section titled "Create model and restore weights for all but last layer".
The code cell below it shows how to download the exact checkpoint that you're interested in.
</li>
<li>You can also review the lecture videos for this week, which show the same code.</li>
</ul>
</p>
<a name='exercise-4'></a>
### Exercise 4: Download checkpoints
- Download the compressed SSD Resnet 50 version 1, 640 x 640 checkpoint.
- Untar (decompress) the tar file
- Move the decompressed checkpoint to `models/research/object_detection/test_data/`
```
### START CODE HERE ###
# Download the SSD Resnet 50 version 1, 640x640 checkpoint
!wget http://download.tensorflow.org/models/object_detection/tf2/20200711/ssd_resnet50_v1_fpn_640x640_coco17_tpu-8.tar.gz
# untar (decompress) the tar file
!tar -xf ssd_resnet50_v1_fpn_640x640_coco17_tpu-8.tar.gz
# copy the checkpoint to the test_data folder models/research/object_detection/test_data/
!mv ssd_resnet50_v1_fpn_640x640_coco17_tpu-8/checkpoint models/research/object_detection/test_data/
### END CODE HERE
```
## Configure the model
Here, you will configure the model for this use case.
<a name='exercise-5-1'></a>
### **Exercise 5.1**: Locate and read from the configuration file
#### pipeline_config
- In the Colab, on the left side table of contents, click on the folder icon to display the file browser for the current workspace.
- Navigate to `models/research/object_detection/configs/tf2`. The folder has multiple .config files.
- Look for the file corresponding to ssd resnet 50 version 1 640x640.
- You can double-click the config file to view its contents. This may help you as you complete the next few code cells to configure your model.
- Set the `pipeline_config` to a string that contains the full path to the resnet config file, in other words: `models/research/.../... .config`
#### configs
If you look at the module [config_util](https://github.com/tensorflow/models/blob/master/research/object_detection/utils/config_util.py) that you imported, it contains the following function:
```
def get_configs_from_pipeline_file(pipeline_config_path, config_override=None):
```
- Please use this function to load the configuration from your `pipeline_config`.
- `configs` will now contain a dictionary.
```
tf.keras.backend.clear_session()
### START CODE HERE ###
# define the path to the .config file for ssd resnet 50 v1 640x640
pipeline_config = '/content/models/research/object_detection/configs/tf2/ssd_resnet50_v1_fpn_640x640_coco17_tpu-8.config'
# Load the configuration file into a dictionary
configs = config_util.get_configs_from_pipeline_file(pipeline_config)
### END CODE HERE ###
# See what configs looks like
configs
```
<a name='exercise-5-2'></a>
### **Exercise 5.2**: Get the model configuration
#### model_config
- From the `configs` dictionary, access the object associated with the key 'model'.
- `model_config` now contains an object of type `object_detection.protos.model_pb2.DetectionModel`.
- If you print `model_config`, you'll see something like this:
```
ssd {
num_classes: 90
image_resizer {
fixed_shape_resizer {
height: 640
width: 640
}
}
feature_extractor {
...
...
freeze_batchnorm: false
```
```
### START CODE HERE ###
# Read in the object stored at the key 'model' of the configs dictionary
model_config = configs.get('model')
### END CODE HERE
# see what model_config looks like
model_config
```
<a name='exercise-5-3'></a>
### **Exercise 5.3**: Modify model_config
- Modify num_classes from the default `90` to the `num_classes` that you set earlier in this notebook.
- num_classes is nested under ssd. You'll need to use dot notation 'obj.x' and NOT bracket notation obj['x']` to access num_classes.
- Freeze batch normalization
- Batch normalization is not frozen in the default configuration.
- If you inspect the `model_config` object, you'll see that `freeze_batchnorm` is nested under `ssd` just like `num_classes`.
- Freeze batch normalization by setting the relevant field to `True`.
```
### START CODE HERE ###
# Modify the number of classes from its default of 90
model_config.ssd.num_classes = num_classes
# Freeze batch normalization
model_config.ssd.freeze_batchnorm = True
### END CODE HERE
# See what model_config now looks like after you've customized it!
model_config
```
## Build the model
Recall that you imported [model_builder](https://github.com/tensorflow/models/blob/master/research/object_detection/builders/model_builder.py).
- You'll use `model_builder` to build the model according to the configurations that you have just downloaded and customized.
<a name='exercise-5.4'></a>
### **Exercise 5.4**: Build the custom model
#### model_builder
model_builder has a function `build`:
```
def build(model_config, is_training, add_summaries=True):
```
- model_config: Set this to the model configuration that you just customized.
- is_training: Set this to True.
- You can keep the default value for the remaining parameter.
- Note that it will take some time to build the model.
```
### START CODE HERE (Replace instances of `None` with your code) ###
detection_model = model_builder.build(model_config=model_config, is_training=True)
### END CODE HERE ###
print(type(detection_model))
```
**Expected Output**:
```txt
<class 'object_detection.meta_architectures.ssd_meta_arch.SSDMetaArch'>
```
## Restore weights from your checkpoint
Now, you will selectively restore weights from your checkpoint.
- Your end goal is to create a custom model which reuses parts of, but not all of the layers of RetinaNet (currently stored in the variable `detection_model`.)
- The parts of RetinaNet that you want to reuse are:
- Feature extraction layers
- Bounding box regression prediction layer
- The part of RetinaNet that you will not want to reuse is the classification prediction layer (since you will define and train your own classification layer specific to zombies).
- For the parts of RetinaNet that you want to reuse, you will also restore the weights from the checkpoint that you selected.
#### Inspect the detection_model
First, take a look at the type of the detection_model and its Python class.
```
# Run this to check the type of detection_model
detection_model
```
#### Find the source code for detection_model
You'll see that the type of the model is `object_detection.meta_architectures.ssd_meta_arch.SSDMetaArch`.
Please practice some detective work and open up the source code for this class in GitHub repository. Recall that at the start of this assignment, you cloned from this repository: [TensorFlow Models](https://github.com/tensorflow/models).
- Navigate through these subfolders: models -> research -> object_detection.
- If you get stuck, go to this link: [object_detection](https://github.com/tensorflow/models/tree/master/research/object_detection)
- Take a look at this 'object_detection' folder and look for the remaining folders to navigate based on the class type of detection_model: object_detection.meta_architectures.ssd_meta_arch.SSDMetaArch
- Hopefully you'll find the meta_architectures folder, and within it you'll notice a file named `ssd_meta_arch.py`.
- Please open and view this [ssd_meta_arch.py](https://github.com/tensorflow/models/blob/master/research/object_detection/meta_architectures/ssd_meta_arch.py) file.
#### View the variables in detection_model
Now, check the class variables that are in `detection_model`.
```
vars(detection_model)
```
You'll see that detection_model contains several variables:
Two of these will be relevant to you:
```
...
_box_predictor': <object_detection.predictors.convolutional_keras_box_predictor.WeightSharedConvolutionalBoxPredictor at 0x7f5205eeb1d0>,
...
_feature_extractor': <object_detection.models.ssd_resnet_v1_fpn_keras_feature_extractor.SSDResNet50V1FpnKerasFeatureExtractor at 0x7f52040f1ef0>,
```
#### Inspect `_feature_extractor`
Take a look at the [ssd_meta_arch.py](https://github.com/tensorflow/models/blob/master/research/object_detection/meta_architectures/ssd_meta_arch.py) code.
```
# Line 302
feature_extractor: a SSDFeatureExtractor object.
```
Also
```
# Line 380
self._feature_extractor = feature_extractor
```
So `detection_model._feature_extractor` is a feature extractor, which you will want to reuse for your zombie detector model.
#### Inspect `_box_predictor`
- View the [ssd_meta_arch.py](https://github.com/tensorflow/models/blob/master/research/object_detection/meta_architectures/ssd_meta_arch.py) file (which is the source code for detection_model)
- Notice that in the __init__ constructor for class SSDMetaArch(model.DetectionModel),
```
...
box_predictor: a box_predictor.BoxPredictor object
...
self._box_predictor = box_predictor
```
#### Inspect _box_predictor
Please take a look at the class type of `detection_model._box_predictor`
```
# view the type of _box_predictor
detection_model._box_predictor
```
You'll see that the class type of _box_predictor is
```
object_detection.predictors.convolutional_keras_box_predictor.WeightSharedConvolutionalBoxPredictor
```
You can navigate through the GitHub repository to this path:
- [objection_detection/predictors](https://github.com/tensorflow/models/tree/master/research/object_detection/predictors)
- Notice that there is a file named convolutional_keras_box_predictor.py. Please open that file.
#### View variables in `_box_predictor`
Also view the variables contained in _box_predictor:
```
vars(detection_model._box_predictor)
```
Among the variables listed, a few will be relevant to you:
```
...
_base_tower_layers_for_heads
...
_box_prediction_head
...
_prediction_heads
```
In the source code for [convolutional_keras_box_predictor.py](https://github.com/tensorflow/models/blob/master/research/object_detection/predictors/convolutional_keras_box_predictor.py) that you just opened, look at the source code to get a sense for what these three variables represent.
#### Inspect `base_tower_layers_for_heads`
If you look at the [convolutional_keras_box_predictor.py](https://github.com/tensorflow/models/blob/master/research/object_detection/predictors/convolutional_keras_box_predictor.py) file, you'll notice this:
```
# line 302
self._base_tower_layers_for_heads = {
BOX_ENCODINGS: [],
CLASS_PREDICTIONS_WITH_BACKGROUND: [],
}
```
- `base_tower_layers_for_heads` is a dictionary with two key-value pairs.
- `BOX_ENCODINGS`: points to a list of layers
- `CLASS_PREDICTIONS_WITH_BACKGROUND`: points to a list of layers
- If you scan the code, you'll see that for both of these, the lists are filled with all layers that appear BEFORE the prediction layer.
```
# Line 377
# Stack the base_tower_layers in the order of conv_layer, batch_norm_layer
# and activation_layer
base_tower_layers = []
for i in range(self._num_layers_before_predictor):
```
So `detection_model.box_predictor._base_tower_layers_for_heads` contains:
- The layers for the prediction before the final bounding box prediction
- The layers for the prediction before the final class prediction.
You will want to use these in your model.
#### Inspect `_box_prediction_head`
If you again look at [convolutional_keras_box_predictor.py](https://github.com/tensorflow/models/blob/master/research/object_detection/predictors/convolutional_keras_box_predictor.py) file, you'll see this
```
# Line 248
box_prediction_head: The head that predicts the boxes.
```
So `detection_model.box_predictor._box_prediction_head` points to the bounding box prediction layer, which you'll want to use for your model.
#### Inspect `_prediction_heads`
If you again look at [convolutional_keras_box_predictor.py](https://github.com/tensorflow/models/blob/master/research/object_detection/predictors/convolutional_keras_box_predictor.py) file, you'll see this
```
# Line 121
self._prediction_heads = {
BOX_ENCODINGS: box_prediction_heads,
CLASS_PREDICTIONS_WITH_BACKGROUND: class_prediction_heads,
}
```
You'll also see this docstring
```
# Line 83
class_prediction_heads: A list of heads that predict the classes.
```
So `detection_model.box_predictor._prediction_heads` is a dictionary that points to both prediction layers:
- The layer that predicts the bounding boxes
- The layer that predicts the class (category).
#### Which layers will you reuse?
Remember that you are reusing the model for its feature extraction and bounding box detection.
- You will create your own classification layer and train it on zombie images.
- So you won't need to reuse the class prediction layer of `detection_model`.
## Define checkpoints for desired layers
You will now isolate the layers of `detection_model` that you wish to reuse so that you can restore the weights to just those layers.
- First, define checkpoints for the box predictor
- Next, define checkpoints for the model, which will point to this box predictor checkpoint as well as the feature extraction layers.
Please use [tf.train.Checkpoint](https://www.tensorflow.org/api_docs/python/tf/train/Checkpoint).
As a reminder of how to use tf.train.Checkpoint:
```
tf.train.Checkpoint(
**kwargs
)
```
Pretend that `detection_model` contains these variables for which you want to restore weights:
- `detection_model._ice_cream_sundae`
- 'detection_model._pies._apple_pie`
- 'detection_model._pies._pecan_pie`
Notice that the pies are nested within `._pies`.
If you just want the ice cream sundae and apple pie variables (and not the pecan pie) then you can do the following:
```
tmp_pies_checkpoint = tf.train.Checkpoint(
_apple_pie = detection_model._pies._apple_pie
)
```
Next, in order to connect these together in a node graph, do this:
```
tmp_model_checkpoint = tf.train.Checkpoint(
_pies = tmp_pies_checkpoint,
_ice_cream_sundae = detection_model._ice_cream_sundae
)
```
Finally, define a checkpoint that uses the key `model` and takes in the tmp_model_checkpoint.
```
checkpoint = tf.train.Checkpoint(
model = tmp_model_checkpoint
)
```
You'll then be ready to restore the weights from the checkpoint that you downloaded.
Try this out step by step!
<a name='exercise-6-1'></a>
### Exercise 6.1: Define Checkpoints for the box predictor
- Please define `box_predictor_checkpoint` to be checkpoint for these two layers of the `detection_model`'s box predictor:
- The base tower layer (the layers the precede both the class prediction and bounding box prediction layers).
- The box prediction head (the prediction layer for bounding boxes).
- Note, you won't include the class prediction layer.
```
### START CODE HERE ###
tmp_box_predictor_checkpoint = tf.train.Checkpoint(
_base_tower_layers_for_heads = detection_model._box_predictor._base_tower_layers_for_heads,
_box_prediction_head = detection_model._box_predictor._box_prediction_head
)
### END CODE HERE
# Check the datatype of this checkpoint
type(tmp_box_predictor_checkpoint)
# Expected output:
# tensorflow.python.training.tracking.util.Checkpoint
# Check the variables of this checkpoint
vars(tmp_box_predictor_checkpoint)
```
#### Expected output
You should expect to see a list of variables that include the following:
```
'_base_tower_layers_for_heads': DictWrapper({'box_encodings': ListWrapper([]), 'class_predictions_with_background': ListWrapper([])}),
'_box_prediction_head': <object_detection.predictors.heads.keras_box_head.WeightSharedConvolutionalBoxHead at 0x7fefac014710>,
...
```
<a name='exercise-6-2'></a>
### Exercise 6.2: Define the temporary model checkpoint**
Now define `tmp_model_checkpoint` so that it points to these two layers:
- The feature extractor of the detection model.
- The temporary box predictor checkpoint that you just defined.
```
### START CODE HERE ###
tmp_model_checkpoint = tf.train.Checkpoint(
_box_predictor = tmp_box_predictor_checkpoint,
_feature_extractor = detection_model._feature_extractor
)
### END CODE HERE ###
# Check the datatype of this checkpoint
type(tmp_model_checkpoint)
# Expected output
# tensorflow.python.training.tracking.util.Checkpoint
# Check the vars of this checkpoint
vars(tmp_model_checkpoint)
```
#### Expected output
Among the variables of this checkpoint, you should see:
```
'_box_predictor': <tensorflow.python.training.tracking.util.Checkpoint at 0x7fefac044a20>,
'_feature_extractor': <object_detection.models.ssd_resnet_v1_fpn_keras_feature_extractor.SSDResNet50V1FpnKerasFeatureExtractor at 0x7fefac0240b8>,
```
<a name='exercise-6-3'></a>
### Exercise 6.3: Restore the checkpoint
You can now restore the checkpoint.
First, find and set the `checkpoint_path`
- checkpoint_path:
- Using the "files" browser in the left side of Colab, navigate to `models -> research -> object_detection -> test_data`.
- If you completed the previous code cell that downloads and moves the checkpoint, you'll see a subfolder named "checkpoint".
- The 'checkpoint' folder contains three files:
- checkpoint
- ckpt-0.data-00000-of-00001
- ckpt-0.index
- Please set checkpoint_path to the path to the full path `models/.../ckpt-0`
- Notice that you don't want to include a file extension after `ckpt-0`.
- **IMPORTANT**: Please don't set the path to include the `.index` extension in the checkpoint file name.
- If you do set it to `ckpt-0.index`, there won't be any immediate error message, but later during training, you'll notice that your model's loss doesn't improve, which means that the pre-trained weights were not restored properly.
Next, define one last checkpoint using `tf.train.Checkpoint()`.
- For the single keyword argument,
- Set the key as `model=`
- Set the value to your temporary model checkpoint that you just defined.
- **IMPORTANT**: You'll need to set the keyword argument as `model=` and not something else like `detection_model=`.
- If you set this keyword argument to anything else, it won't show an immmediate error, but when you train your model on the zombie images, your model loss will not decrease (your model will not learn).
Finally, call this checkpoint's `.restore()` function, passing in the path to the checkpoint.
```
### START CODE HERE ###
checkpoint_path = 'models/research/object_detection/test_data/checkpoint/ckpt-0'
# Define a checkpoint that sets `model= None
checkpoint = tf.train.Checkpoint(
model=tmp_model_checkpoint
)
# Restore the checkpoint to the checkpoint path
checkpoint.restore(checkpoint_path)
### END CODE HERE ###
```
<a name='exercise-7'></a>
### **Exercise 7**: Run a dummy image to generate the model variables
Run a dummy image through the model so that variables are created. We need to select the trainable variables later in Exercise 9 and right now, it is still empty. Try running `len(detection_model.trainable_variables)` in a code cell and you will get `0`. We will pass in a dummy image through the forward pass to create these variables.
Recall that `detection_model` is an object of type [object_detection.meta_architectures.ssd_meta_arch.SSDMetaArch]()
Important methods that are available in the `detection_model` object are:
- [preprocess()](https://github.com/tensorflow/models/blob/dc4d11216b738920ddb136729e3ae71bddb75c7e/research/object_detection/meta_architectures/ssd_meta_arch.py#L459):
- takes in a tensor representing an image and returns
- returns `image, shapes`
- For the dummy image, you can declare a [tensor of zeros](https://www.tensorflow.org/api_docs/python/tf/zeros) that has a shape that the `preprocess()` method can accept (i.e. [batch, height, width, channels]).
- Remember that your images have dimensions 640 x 640 x 3.
- You can pass in a batch of 1 when making the dummy image.
- [predict()](https://github.com/tensorflow/models/blob/dc4d11216b738920ddb136729e3ae71bddb75c7e/research/object_detection/meta_architectures/ssd_meta_arch.py#L525)
- takes in `image, shapes` which are created by the `preprocess()` function call.
- returns a prediction in a Python dictionary
- this will pass the dummy image through the forward pass of the network and create the model variables
- [postprocess()](https://github.com/tensorflow/models/blob/dc4d11216b738920ddb136729e3ae71bddb75c7e/research/object_detection/meta_architectures/ssd_meta_arch.py#L655)
- Takes in the prediction_dict and shapes
- returns a dictionary of post-processed predictions of detected objects ("detections").
**Note**: Please use the recommended variable names, which include the prefix `tmp_`, since these variables won't be used later, but you'll define similarly-named variables later for predicting on actual zombie images.
```
### START CODE HERE (Replace instances of `None` with your code)###
# use the detection model's `preprocess()` method and pass a dummy image
tmp_image, tmp_shapes = detection_model.preprocess(tf.zeros([1, 640, 640, 3]))
# run a prediction with the preprocessed image and shapes
tmp_prediction_dict = detection_model.predict(tmp_image, tmp_shapes)
# postprocess the predictions into final detections
tmp_detections = detection_model.postprocess(tmp_prediction_dict, tmp_shapes)
### END CODE HERE ###
print('Weights restored!')
# Test Code:
assert len(detection_model.trainable_variables) > 0, "Please pass in a dummy image to create the trainable variables."
print(detection_model.weights[0].shape)
print(detection_model.weights[231].shape)
print(detection_model.weights[462].shape)
```
**Expected Output**:
```txt
(3, 3, 256, 24)
(512,)
(256,)
```
## Eager mode custom training loop
With the data and model now setup, you can now proceed to configure the training.
<a name='exercise-8'></a>
### **Exercise 8**: Set training hyperparameters
Set an appropriate learning rate and optimizer for the training.
- batch_size: you can use 4
- You can increase the batch size up to 5, since you have just 5 images for training.
- num_batches: You can use 100
- You can increase the number of batches but the training will take longer to complete.
- learning_rate: You can use 0.01
- When you run the training loop later, notice how the initial loss INCREASES` before decreasing.
- You can try a lower learning rate to see if you can avoid this increased loss.
- optimizer: you can use [tf.keras.optimizers.SGD](https://www.tensorflow.org/api_docs/python/tf/keras/optimizers/SGD)
- Set the learning rate
- Set the momentum to 0.9
Training will be fairly quick, so we do encourage you to experiment a bit with these hyperparameters!
```
tf.keras.backend.set_learning_phase(True)
### START CODE HERE (Replace instances of `None` with your code)###
# set the batch_size
batch_size = 5
# set the number of batches
num_batches = 100
# Set the learning rate
learning_rate = 0.01
# set the optimizer and pass in the learning_rate
optimizer = tf.keras.optimizers.SGD(learning_rate=learning_rate, momentum=0.9)
### END CODE HERE ###
```
## Choose the layers to fine-tune
To make use of transfer learning and pre-trained weights, you will train just certain parts of the detection model, namely, the last prediction layers.
- Please take a minute to inspect the layers of `detection_model`.
```
# Inspect the layers of detection_model
for i,v in enumerate(detection_model.trainable_variables):
print(f"i: {i} \t name: {v.name} \t shape:{v.shape} \t dtype={v.dtype}")
```
Notice that there are some layers whose names are prefixed with the following:
```
WeightSharedConvolutionalBoxPredictor/WeightSharedConvolutionalBoxHead
...
WeightSharedConvolutionalBoxPredictor/WeightSharedConvolutionalClassHead
...
WeightSharedConvolutionalBoxPredictor/BoxPredictionTower
...
WeightSharedConvolutionalBoxPredictor/ClassPredictionTower
...
```
Among these, which do you think are the prediction layers at the "end" of the model?
- Recall that when inspecting the source code to restore the checkpoints ([convolutional_keras_box_predictor.py](https://github.com/tensorflow/models/blob/master/research/object_detection/predictors/convolutional_keras_box_predictor.py)) you noticed that:
- `_base_tower_layers_for_heads`: refers to the layers that are placed right before the prediction layer
- `_box_prediction_head` refers to the prediction layer for the bounding boxes
- `_prediction_heads`: refers to the set of prediction layers (both for classification and for bounding boxes)
So you can see that in the source code for this model, "tower" refers to layers that are before the prediction layer, and "head" refers to the prediction layers.
<a name='exercise-9'></a>
### **Exercise 9**: Select the prediction layer variables
Based on inspecting the `detection_model.trainable_variables`, please select the prediction layer variables that you will fine tune:
- The bounding box head variables (which predict bounding box coordinates)
- The class head variables (which predict the class/category)
You have a few options for doing this:
- You can access them by their list index:
```
detection_model.trainable_variables[92]
```
- Alternatively, you can use string matching to select the variables:
```
tmp_list = []
for v in detection_model.trainable_variables:
if v.name.startswith('ResNet50V1_FPN/bottom_up_block5'):
tmp_list.append(v)
```
**Hint**: There are a total of four variables that you want to fine tune.
```
### START CODE HERE (Replace instances of `None` with your code) ###
# define a list that contains the layers that you wish to fine tune
to_fine_tune = []
for v in detection_model.trainable_variables:
if v.name.startswith('WeightSharedConvolutionalBoxPredictor/WeightSharedConvolutional'):
to_fine_tune.append(v)
### END CODE HERE
# Test Code:
print(to_fine_tune[0].name)
print(to_fine_tune[2].name)
```
**Expected Output**:
```txt
WeightSharedConvolutionalBoxPredictor/WeightSharedConvolutionalBoxHead/BoxPredictor/kernel:0
WeightSharedConvolutionalBoxPredictor/WeightSharedConvolutionalClassHead/ClassPredictor/kernel:0
```
## Train your model
You'll define a function that handles training for one batch, which you'll later use in your training loop.
First, walk through these code cells to learn how you'll perform training using this model.
```
# Get a batch of your training images
g_images_list = train_image_tensors[0:2]
```
The `detection_model` is of class [SSDMetaArch](https://github.com/tensorflow/models/blob/dc4d11216b738920ddb136729e3ae71bddb75c7e/research/object_detection/meta_architectures/ssd_meta_arch.py#L655), and its source code shows that is has this function [preprocess](https://github.com/tensorflow/models/blob/dc4d11216b738920ddb136729e3ae71bddb75c7e/research/object_detection/meta_architectures/ssd_meta_arch.py#L459).
- This preprocesses the images so that they can be passed into the model (for training or prediction):
```
def preprocess(self, inputs):
"""Feature-extractor specific preprocessing.
...
Args:
inputs: a [batch, height_in, width_in, channels] float tensor representing
a batch of images with values between 0 and 255.0.
Returns:
preprocessed_inputs: a [batch, height_out, width_out, channels] float
tensor representing a batch of images.
true_image_shapes: int32 tensor of shape [batch, 3] where each row is
of the form [height, width, channels] indicating the shapes
of true images in the resized images, as resized images can be padded
with zeros.
```
```
# Use .preprocess to preprocess an image
g_preprocessed_image = detection_model.preprocess(g_images_list[0])
print(f"g_preprocessed_image type: {type(g_preprocessed_image)}")
print(f"g_preprocessed_image length: {len(g_preprocessed_image)}")
print(f"index 0 has the preprocessed image of shape {g_preprocessed_image[0].shape}")
print(f"index 1 has information about the image's true shape excluding padding: {g_preprocessed_image[1]}")
```
You can pre-process each image and save their outputs into two separate lists
- One list of the preprocessed images
- One list of the true shape for each preprocessed image
```
preprocessed_image_list = []
true_shape_list = []
for img in g_images_list:
processed_img, true_shape = detection_model.preprocess(img)
preprocessed_image_list.append(processed_img)
true_shape_list.append(true_shape)
print(f"preprocessed_image_list is of type {type(preprocessed_image_list)}")
print(f"preprocessed_image_list has length {len(preprocessed_image_list)}")
print()
print(f"true_shape_list is of type {type(true_shape_list)}")
print(f"true_shape_list has length {len(true_shape_list)}")
```
## Make a prediction
The `detection_model` also has a `.predict` function. According to the source code for [predict](https://github.com/tensorflow/models/blob/dc4d11216b738920ddb136729e3ae71bddb75c7e/research/object_detection/meta_architectures/ssd_meta_arch.py#L525)
```
def predict(self, preprocessed_inputs, true_image_shapes):
"""Predicts unpostprocessed tensors from input tensor.
This function takes an input batch of images and runs it through the forward
pass of the network to yield unpostprocessesed predictions.
...
Args:
preprocessed_inputs: a [batch, height, width, channels] image tensor.
true_image_shapes: int32 tensor of shape [batch, 3] where each row is
of the form [height, width, channels] indicating the shapes
of true images in the resized images, as resized images can be padded
with zeros.
Returns:
prediction_dict: a dictionary holding "raw" prediction tensors:
1) preprocessed_inputs: the [batch, height, width, channels] image
tensor.
2) box_encodings: 4-D float tensor of shape [batch_size, num_anchors,
box_code_dimension] containing predicted boxes.
3) class_predictions_with_background: 3-D float tensor of shape
[batch_size, num_anchors, num_classes+1] containing class predictions
(logits) for each of the anchors. Note that this tensor *includes*
background class predictions (at class index 0).
4) feature_maps: a list of tensors where the ith tensor has shape
[batch, height_i, width_i, depth_i].
5) anchors: 2-D float tensor of shape [num_anchors, 4] containing
the generated anchors in normalized coordinates.
6) final_anchors: 3-D float tensor of shape [batch_size, num_anchors, 4]
containing the generated anchors in normalized coordinates.
If self._return_raw_detections_during_predict is True, the dictionary
will also contain:
7) raw_detection_boxes: a 4-D float32 tensor with shape
[batch_size, self.max_num_proposals, 4] in normalized coordinates.
8) raw_detection_feature_map_indices: a 3-D int32 tensor with shape
[batch_size, self.max_num_proposals].
"""
```
Notice that `.predict` takes its inputs as tensors. If you tried to pass in the preprocessed images and true shapes, you'll get an error.
```
# Try to call `predict` and pass in lists; look at the error message
try:
detection_model.predict(preprocessed_image_list, true_shape_list)
except AttributeError as e:
print("Error message:", e)
```
But don't worry! You can check how to properly use `predict`:
- Notice that the source code documentation says that `preprocessed_inputs` and `true_image_shapes` are expected to be tensors and not lists of tensors.
- One way to turn a list of tensors into a tensor is to use [tf.concat](https://www.tensorflow.org/api_docs/python/tf/concat)
```
tf.concat(
values, axis, name='concat'
)
```
```
# Turn a list of tensors into a tensor
preprocessed_image_tensor = tf.concat(preprocessed_image_list, axis=0)
true_shape_tensor = tf.concat(true_shape_list, axis=0)
print(f"preprocessed_image_tensor shape: {preprocessed_image_tensor.shape}")
print(f"true_shape_tensor shape: {true_shape_tensor.shape}")
```
Now you can make predictions for the images.
According to the source code, `predict` returns a dictionary containing the prediction information, including:
- The bounding box predictions
- The class predictions
```
# Make predictions on the images
prediction_dict = detection_model.predict(preprocessed_image_tensor, true_shape_tensor)
print("keys in prediction_dict:")
for key in prediction_dict.keys():
print(key)
```
#### Calculate loss
Now that your model has made its prediction, you want to compare it to the ground truth in order to calculate a loss.
- The `detection_model` has a [loss](https://github.com/tensorflow/models/blob/dc4d11216b738920ddb136729e3ae71bddb75c7e/research/object_detection/meta_architectures/ssd_meta_arch.py#L807) function.
```Python
def loss(self, prediction_dict, true_image_shapes, scope=None):
"""Compute scalar loss tensors with respect to provided groundtruth.
Calling this function requires that groundtruth tensors have been
provided via the provide_groundtruth function.
Args:
prediction_dict: a dictionary holding prediction tensors with
1) box_encodings: 3-D float tensor of shape [batch_size, num_anchors,
box_code_dimension] containing predicted boxes.
2) class_predictions_with_background: 3-D float tensor of shape
[batch_size, num_anchors, num_classes+1] containing class predictions
(logits) for each of the anchors. Note that this tensor *includes*
background class predictions.
true_image_shapes: int32 tensor of shape [batch, 3] where each row is
of the form [height, width, channels] indicating the shapes
of true images in the resized images, as resized images can be padded
with zeros.
scope: Optional scope name.
Returns:
a dictionary mapping loss keys (`localization_loss` and
`classification_loss`) to scalar tensors representing corresponding loss
values.
"""
```
It takes in:
- The prediction dictionary that comes from your call to `.predict()`.
- the true images shape that comes from your call to `.preprocess()` followed by the conversion from a list to a tensor.
Try calling `.loss`. You'll see an error message that you'll addres in order to run the `.loss` function.
```
try:
losses_dict = detection_model.loss(prediction_dict, true_shape_tensor)
except RuntimeError as e:
print(e)
```
This is giving an error about groundtruth_classes_list:
```
The graph tensor has name: groundtruth_classes_list:0
```
Notice in the docstring for `loss` (shown above), it says:
```
Calling this function requires that groundtruth tensors have been
provided via the provide_groundtruth function.
```
So you'll first want to set the ground truth (true labels and true bounding boxes) before you calculate the loss.
- This makes sense, since the loss is comparing the prediction to the ground truth, and so the loss function needs to know the ground truth.
#### Provide the ground truth
The source code for providing the ground truth is located in the parent class of `SSDMetaArch`, `model.DetectionModel`.
- Here is the link to the code for [provide_ground_truth](https://github.com/tensorflow/models/blob/fd6b24c19c68af026bb0a9efc9f7b1719c231d3d/research/object_detection/core/model.py#L297)
```Python
def provide_groundtruth(
self,
groundtruth_boxes_list,
groundtruth_classes_list,
... # more parameters not show here
"""
Args:
groundtruth_boxes_list: a list of 2-D tf.float32 tensors of shape
[num_boxes, 4] containing coordinates of the groundtruth boxes.
Groundtruth boxes are provided in [y_min, x_min, y_max, x_max]
format and assumed to be normalized and clipped
relative to the image window with y_min <= y_max and x_min <= x_max.
groundtruth_classes_list: a list of 2-D tf.float32 one-hot (or k-hot)
tensors of shape [num_boxes, num_classes] containing the class targets
with the 0th index assumed to map to the first non-background class.
"""
```
You'll set two parameters in `provide_ground_truth`:
- The true bounding boxes
- The true classes
```
# Get the ground truth bounding boxes
gt_boxes_list = gt_box_tensors[0:2]
# Get the ground truth class labels
gt_classes_list = gt_classes_one_hot_tensors[0:2]
# Provide the ground truth to the model
detection_model.provide_groundtruth(
groundtruth_boxes_list=gt_boxes_list,
groundtruth_classes_list=gt_classes_list)
```
Now you can calculate the loss
```
# Calculate the loss after you've provided the ground truth
losses_dict = detection_model.loss(prediction_dict, true_shape_tensor)
# View the loss dictionary
losses_dict = detection_model.loss(prediction_dict, true_shape_tensor)
print(f"loss dictionary keys: {losses_dict.keys()}")
print(f"localization loss {losses_dict['Loss/localization_loss']:.8f}")
print(f"classification loss {losses_dict['Loss/classification_loss']:.8f}")
```
You can now calculate the gradient and optimize the variables that you selected to fine tune.
- Use tf.GradientTape
```Python
with tf.GradientTape() as tape:
# Make the prediction
# calculate the loss
# calculate the gradient of each model variable with respect to each loss
gradients = tape.gradient([some loss], variables to fine tune)
# apply the gradients to update these model variables
optimizer.apply_gradients(zip(gradients, variables to fine tune))
```
```
# Let's just reset the model so that you can practice setting it up yourself!
detection_model.provide_groundtruth(groundtruth_boxes_list=[], groundtruth_classes_list=[])
```
<a name='exercise-10'></a>
### **Exercise 10**: Define the training step
Please complete the function below to set up one training step.
- Preprocess the images
- Make a prediction
- Calculate the loss (and make sure the loss function has the ground truth to compare with the prediction)
- Calculate the total loss:
- `total_loss` = `localization_loss + classification_loss`
- Note: this is different than the example code that you saw above
- Calculate gradients with respect to the variables you selected to train.
- Optimize the model's variables
```
# decorate with @tf.function for faster training (remember, graph mode!)
@tf.function
def train_step_fn(image_list,
groundtruth_boxes_list,
groundtruth_classes_list,
model,
optimizer,
vars_to_fine_tune):
"""A single training iteration.
Args:
image_list: A list of [1, height, width, 3] Tensor of type tf.float32.
Note that the height and width can vary across images, as they are
reshaped within this function to be 640x640.
groundtruth_boxes_list: A list of Tensors of shape [N_i, 4] with type
tf.float32 representing groundtruth boxes for each image in the batch.
groundtruth_classes_list: A list of Tensors of shape [N_i, num_classes]
with type tf.float32 representing groundtruth boxes for each image in
the batch.
Returns:
A scalar tensor representing the total loss for the input batch.
"""
# Provide the ground truth to the model
model.provide_groundtruth(
groundtruth_boxes_list=groundtruth_boxes_list,
groundtruth_classes_list=groundtruth_classes_list
)
with tf.GradientTape() as tape:
### START CODE HERE (Replace instances of `None` with your code) ###
# Preprocess the images
preprocessed_image_list = []
true_shape_list = []
for img in image_list:
processed_img, true_shape = model.preprocess(img)
preprocessed_image_list.append(processed_img)
true_shape_list.append(true_shape)
preprocessed_image_tensor = tf.concat(preprocessed_image_list, axis=0)
true_shape_tensor = tf.concat(true_shape_list, axis=0)
# Make a prediction
prediction_dict = model.predict(preprocessed_image_tensor, true_shape_tensor)
# Calculate the total loss (sum of both losses)
losses_dict = model.loss(prediction_dict, true_shape_tensor)
total_loss = losses_dict['Loss/localization_loss'] + losses_dict['Loss/classification_loss']
# Calculate the gradients
gradients = tape.gradient([total_loss], vars_to_fine_tune)
# Optimize the model's selected variables
optimizer.apply_gradients(zip(gradients, vars_to_fine_tune))
### END CODE HERE ###
return total_loss
```
## Run the training loop
Run the training loop using the training step function that you just defined.
```
print('Start fine-tuning!', flush=True)
for idx in range(num_batches):
# Grab keys for a random subset of examples
all_keys = list(range(len(train_images_np)))
random.shuffle(all_keys)
example_keys = all_keys[:batch_size]
# Get the ground truth
gt_boxes_list = [gt_box_tensors[key] for key in example_keys]
gt_classes_list = [gt_classes_one_hot_tensors[key] for key in example_keys]
# get the images
image_tensors = [train_image_tensors[key] for key in example_keys]
# Training step (forward pass + backwards pass)
total_loss = train_step_fn(image_tensors,
gt_boxes_list,
gt_classes_list,
detection_model,
optimizer,
to_fine_tune
)
if idx % 10 == 0:
print('batch ' + str(idx) + ' of ' + str(num_batches)
+ ', loss=' + str(total_loss.numpy()), flush=True)
print('Done fine-tuning!')
```
**Expected Output:**
Total loss should be decreasing and should be less than 1 after fine tuning. For example:
```txt
Start fine-tuning!
batch 0 of 100, loss=1.2559178
batch 10 of 100, loss=16.067217
batch 20 of 100, loss=8.094654
batch 30 of 100, loss=0.34514275
batch 40 of 100, loss=0.033170983
batch 50 of 100, loss=0.0024622646
batch 60 of 100, loss=0.00074224477
batch 70 of 100, loss=0.0006149876
batch 80 of 100, loss=0.00046916265
batch 90 of 100, loss=0.0004159231
Done fine-tuning!
```
## Load test images and run inference with new model!
You can now test your model on a new set of images. The cell below downloads 237 images of a walking zombie and stores them in a `results/` directory.
```
# uncomment if you want to delete existing files
!rm zombie-walk-frames.zip
!rm -rf ./zombie-walk
!rm -rf ./results
# download test images
!wget --no-check-certificate \
https://storage.googleapis.com/laurencemoroney-blog.appspot.com/zombie-walk-frames.zip \
-O zombie-walk-frames.zip
# unzip test images
local_zip = './zombie-walk-frames.zip'
zip_ref = zipfile.ZipFile(local_zip, 'r')
zip_ref.extractall('./results')
zip_ref.close()
```
You will load these images into numpy arrays to prepare it for inference.
```
test_image_dir = './results/'
test_images_np = []
# load images into a numpy array. this will take a few minutes to complete.
for i in range(0, 237):
image_path = os.path.join(test_image_dir, 'zombie-walk' + "{0:04}".format(i) + '.jpg')
print(image_path)
test_images_np.append(np.expand_dims(
load_image_into_numpy_array(image_path), axis=0))
```
<a name='exercise-11'></a>
### **Exercise 11**: Preprocess, predict, and post process an image
Define a function that returns the detection boxes, classes, and scores.
```
# Again, uncomment this decorator if you want to run inference eagerly
@tf.function
def detect(input_tensor):
"""Run detection on an input image.
Args:
input_tensor: A [1, height, width, 3] Tensor of type tf.float32.
Note that height and width can be anything since the image will be
immediately resized according to the needs of the model within this
function.
Returns:
A dict containing 3 Tensors (`detection_boxes`, `detection_classes`,
and `detection_scores`).
"""
preprocessed_image, shapes = detection_model.preprocess(input_tensor)
prediction_dict = detection_model.predict(preprocessed_image, shapes)
### START CODE HERE (Replace instances of `None` with your code) ###
# use the detection model's postprocess() method to get the the final detections
detections = detection_model.postprocess(prediction_dict, shapes)
### END CODE HERE ###
return detections
```
You can now loop through the test images and get the detection scores and bounding boxes to overlay in the original image. We will save each result in a `results` dictionary and the autograder will use this to evaluate your results.
```
# Note that the first frame will trigger tracing of the tf.function, which will
# take some time, after which inference should be fast.
label_id_offset = 1
results = {'boxes': [], 'scores': []}
for i in range(len(test_images_np)):
input_tensor = tf.convert_to_tensor(test_images_np[i], dtype=tf.float32)
detections = detect(input_tensor)
plot_detections(
test_images_np[i][0],
detections['detection_boxes'][0].numpy(),
detections['detection_classes'][0].numpy().astype(np.uint32)
+ label_id_offset,
detections['detection_scores'][0].numpy(),
category_index, figsize=(15, 20), image_name="./results/gif_frame_" + ('%03d' % i) + ".jpg")
results['boxes'].append(detections['detection_boxes'][0][0].numpy())
results['scores'].append(detections['detection_scores'][0][0].numpy())
# TEST CODE
print(len(results['boxes']))
print(results['boxes'][0].shape)
print()
# compare with expected bounding boxes
print(np.allclose(results['boxes'][0], [0.28838485, 0.06830047, 0.7213766 , 0.19833465], rtol=0.18))
print(np.allclose(results['boxes'][5], [0.29168868, 0.07529271, 0.72504973, 0.20099735], rtol=0.18))
print(np.allclose(results['boxes'][10], [0.29548776, 0.07994056, 0.7238164 , 0.20778716], rtol=0.18))
```
**Expected Output:** Ideally the three boolean values at the bottom should be `True`. But if you only get two, you can still try submitting. This compares your resulting bounding boxes for each zombie image to some preloaded coordinates (i.e. the hardcoded values in the test cell above). Depending on how you annotated the training images,it's possible that some of your results differ for these three frames but still get good results overall when all images are examined by the grader. If two or all are False, please try annotating the images again with a tighter bounding box or use the [predefined `gt_boxes` list](#gt-boxes).
```txt
237
(4,)
True
True
True
```
You can also check if the model detects a zombie class in the images by examining the `scores` key of the `results` dictionary. You should get higher than 88.0 here.
```
x = np.array(results['scores'])
# percent of frames where a zombie is detected
zombie_detected = (np.where(x > 0.9, 1, 0).sum())/237*100
print(zombie_detected)
```
You can also display some still frames and inspect visually. If you don't see a bounding box around the zombie, please consider re-annotating the ground truth or use the predefined `gt_boxes` [here](#gt-boxes)
```
print('Frame 0')
display(IPyImage('./results/gif_frame_000.jpg'))
print()
print('Frame 5')
display(IPyImage('./results/gif_frame_005.jpg'))
print()
print('Frame 10')
display(IPyImage('./results/gif_frame_010.jpg'))
```
## Create a zip of the zombie-walk images.
You can download this if you like to create your own animations
```
zipf = zipfile.ZipFile('./zombie.zip', 'w', zipfile.ZIP_DEFLATED)
filenames = glob.glob('./results/gif_frame_*.jpg')
filenames = sorted(filenames)
for filename in filenames:
zipf.write(filename)
zipf.close()
```
## Create Zombie animation
```
imageio.plugins.freeimage.download()
!rm -rf ./results/zombie-anim.gif
anim_file = './zombie-anim.gif'
filenames = glob.glob('./results/gif_frame_*.jpg')
filenames = sorted(filenames)
last = -1
images = []
for filename in filenames:
image = imageio.imread(filename)
images.append(image)
imageio.mimsave(anim_file, images, 'GIF-FI', fps=10)
```
Unfortunately, using `IPyImage` in the notebook (as you've done in the rubber ducky detection tutorial) for the large `gif` generated will disconnect the runtime. To view the animation, you can instead use the `Files` pane on the left and double-click on `zombie-anim.gif`. That will open a preview page on the right. It will take 2 to 3 minutes to load and see the walking zombie.
## Save results file for grading
Run the cell below to save your results. Download the `results.data` file and upload it to the grader in the classroom.
```
import pickle
# remove file if it exists
!rm results.data
# write results to binary file. upload for grading.
with open('results.data', 'wb') as filehandle:
pickle.dump(results['boxes'], filehandle)
print('Done saving! Please download `results.data` from the Files tab\n' \
'on the left and submit for grading.\nYou can also use the next cell as a shortcut for downloading.')
from google.colab import files
files.download('results.data')
```
**Congratulations on completing this assignment! Please go back to the Coursera classroom and upload `results.data` to the Graded Lab item for Week 2.**
| github_jupyter |
# [deplacy](https://koichiyasuoka.github.io/deplacy/)'de sözdizimsel analiz
## [Stanza](https://stanfordnlp.github.io/stanza) ile analiz
```
!pip install deplacy stanza
import stanza
stanza.download("tr")
nlp=stanza.Pipeline("tr")
doc=nlp("İyi insan sözünün üstüne gelir.")
import deplacy
deplacy.render(doc)
deplacy.serve(doc,port=None)
# import graphviz
# graphviz.Source(deplacy.dot(doc))
```
## [NLP-Cube](https://github.com/Adobe/NLP-Cube) ile analiz
```
!pip install deplacy nlpcube
from cube.api import Cube
nlp=Cube()
nlp.load("tr")
doc=nlp("İyi insan sözünün üstüne gelir.")
import deplacy
deplacy.render(doc)
deplacy.serve(doc,port=None)
# import graphviz
# graphviz.Source(deplacy.dot(doc))
```
## [spaCy-COMBO](https://github.com/KoichiYasuoka/spaCy-COMBO) ile analiz
```
!pip install deplacy spacy_combo
import spacy_combo
nlp=spacy_combo.load("tr_imst")
doc=nlp("İyi insan sözünün üstüne gelir.")
import deplacy
deplacy.render(doc)
deplacy.serve(doc,port=None)
# import graphviz
# graphviz.Source(deplacy.dot(doc))
```
## [spaCy-jPTDP](https://github.com/KoichiYasuoka/spaCy-jPTDP) ile analiz
```
!pip install deplacy spacy_jptdp
import spacy_jptdp
nlp=spacy_jptdp.load("tr_imst")
doc=nlp("İyi insan sözünün üstüne gelir.")
import deplacy
deplacy.render(doc)
deplacy.serve(doc,port=None)
# import graphviz
# graphviz.Source(deplacy.dot(doc))
```
## [Camphr-Udify](https://camphr.readthedocs.io/en/latest/notes/udify.html) ile analiz
```
!pip install deplacy camphr 'unofficial-udify>=0.3.0' en-udify@https://github.com/PKSHATechnology-Research/camphr_models/releases/download/0.7.0/en_udify-0.7.tar.gz
import pkg_resources,imp
imp.reload(pkg_resources)
import spacy
nlp=spacy.load("en_udify")
doc=nlp("İyi insan sözünün üstüne gelir.")
import deplacy
deplacy.render(doc)
deplacy.serve(doc,port=None)
# import graphviz
# graphviz.Source(deplacy.dot(doc))
```
## [Turku-neural-parser-pipeline](https://turkunlp.org/Turku-neural-parser-pipeline/) ile analiz
```
!pip install deplacy ufal.udpipe configargparse 'tensorflow<2' torch==0.4.1 torchtext==0.3.1 torchvision==0.2.1
!test -d Turku-neural-parser-pipeline || git clone --depth=1 https://github.com/TurkuNLP/Turku-neural-parser-pipeline
!cd Turku-neural-parser-pipeline && git submodule update --init --recursive && test -d models_tr_imst || python fetch_models.py tr_imst
import sys,subprocess
nlp=lambda t:subprocess.run([sys.executable,"full_pipeline_stream.py","--gpu","-1","--conf","models_tr_imst/pipelines.yaml"],cwd="Turku-neural-parser-pipeline",input=t,encoding="utf-8",stdout=subprocess.PIPE).stdout
doc=nlp("İyi insan sözünün üstüne gelir.")
import deplacy
deplacy.render(doc)
deplacy.serve(doc,port=None)
# import graphviz
# graphviz.Source(deplacy.dot(doc))
```
## [UDPipe 2](http://ufal.mff.cuni.cz/udpipe/2) ile analiz
```
!pip install deplacy
def nlp(t):
import urllib.request,urllib.parse,json
with urllib.request.urlopen("https://lindat.mff.cuni.cz/services/udpipe/api/process?model=tr&tokenizer&tagger&parser&data="+urllib.parse.quote(t)) as r:
return json.loads(r.read())["result"]
doc=nlp("İyi insan sözünün üstüne gelir.")
import deplacy
deplacy.render(doc)
deplacy.serve(doc,port=None)
# import graphviz
# graphviz.Source(deplacy.dot(doc))
```
## [spacy-udpipe](https://github.com/TakeLab/spacy-udpipe) ile analiz
```
!pip install deplacy spacy-udpipe
import spacy_udpipe
spacy_udpipe.download("tr")
nlp=spacy_udpipe.load("tr")
doc=nlp("İyi insan sözünün üstüne gelir.")
import deplacy
deplacy.render(doc)
deplacy.serve(doc,port=None)
# import graphviz
# graphviz.Source(deplacy.dot(doc))
```
## [COMBO-pytorch](https://gitlab.clarin-pl.eu/syntactic-tools/combo) ile analiz
```
!pip install --index-url https://pypi.clarin-pl.eu/simple deplacy combo
import combo.predict
nlp=combo.predict.COMBO.from_pretrained("turkish-ud27")
doc=nlp("İyi insan sözünün üstüne gelir.")
import deplacy
deplacy.render(doc)
deplacy.serve(doc,port=None)
# import graphviz
# graphviz.Source(deplacy.dot(doc))
```
## [Trankit](https://github.com/nlp-uoregon/trankit) ile analiz
```
!pip install deplacy trankit transformers
import trankit
nlp=trankit.Pipeline("turkish")
doc=nlp("İyi insan sözünün üstüne gelir.")
import deplacy
deplacy.render(doc)
deplacy.serve(doc,port=None)
# import graphviz
# graphviz.Source(deplacy.dot(doc))
```
| github_jupyter |
```
import pandas as pd
from tensorflow import keras
from tensorflow.keras.datasets import mnist
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Dropout, Conv2D, MaxPooling2D, Flatten
from tensorflow.keras.optimizers import RMSprop
import matplotlib.pyplot as plt
from sklearn.metrics import classification_report, confusion_matrix
train = pd.read_csv("train.csv")
Y = train["label"]
X = train.drop(labels = ["label"],axis = 1)
x = X.values.reshape(42000, 28, 28, 1)
x = x.astype('float32')
x /= 255
num_classes = 10
y = keras.utils.to_categorical(Y, num_classes)
print(y[0])
from sklearn.model_selection import train_test_split
x_train, x_val, y_train, y_val = train_test_split(x, y, test_size = 0.1, random_state=5)
print('Qtde de treino: {}'.format(len(x_train)))
print('Qtde de validação: {}'.format(len(x_val)))
model = Sequential()
model.add(Conv2D(20, kernel_size=(3, 3),
activation='relu',
input_shape=(28,28,1)))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Conv2D(40, kernel_size=(3,3), activation='relu'))
model.add(Flatten())
model.add(Dense(50, activation='relu'))
model.add(Dropout(0.2))
model.add(Dense(num_classes, activation='softmax'))
model.summary()
model.compile(loss='categorical_crossentropy',
optimizer=RMSprop(),
metrics=['accuracy'])
batch_size = 32
epochs = 20
callbacks_list = [
keras.callbacks.ModelCheckpoint(
filepath='model.h5',
monitor='val_loss', save_best_only=True, verbose=1),
keras.callbacks.EarlyStopping(monitor='val_loss', patience=10,verbose=1)
]
history = model.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
callbacks = callbacks_list,
verbose=1,
validation_data=(x_val, y_val))
fig, ax = plt.subplots(1,2, figsize=(16,8))
ax[0].plot(history.history['loss'], color='b', label="Training loss")
ax[0].plot(history.history['val_loss'], color='r', label="validation loss",axes =ax[0])
legend = ax[0].legend(loc='best', shadow=True)
ax[1].plot(history.history['accuracy'], color='b', label="Training accuracy")
ax[1].plot(history.history['val_accuracy'], color='r',label="Validation accuracy")
legend = ax[1].legend(loc='best', shadow=True)
from tensorflow.keras.models import load_model
model = load_model('model.h5')
score = model.evaluate(x_val, y_val, verbose=0)
print('Test loss:', score[0])
print('Test accuracy:', score[1])
print(y_train[10])
print(model.predict(x_train[10].reshape((1,28,28,1))))
print(model.predict_classes(x_train[10].reshape((1,28,28,1))))
import itertools
#Plot the confusion matrix. Set Normalize = True/False
def plot_confusion_matrix(cm, classes, normalize=True, title='Confusion matrix', cmap=plt.cm.Blues):
"""
This function prints and plots the confusion matrix.
Normalization can be applied by setting `normalize=True`.
"""
plt.figure(figsize=(10,10))
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
cm = np.around(cm, decimals=2)
cm[np.isnan(cm)] = 0.0
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, cm[i, j],
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
import numpy as np
# Classificando toda base de teste
y_pred = model.predict_classes(x_val)
# voltando pro formato de classes
y_test_c = np.argmax(y_val, axis=1)
target_names = ['0', '1', '2', '3', '4', '5', '6', '7', '8', '9']
#Confution Matrix
cm = confusion_matrix(y_test_c, y_pred)
plot_confusion_matrix(cm, target_names, normalize=False, title='Confusion Matrix')
print('Classification Report')
print(classification_report(y_test_c, y_pred, target_names=target_names))
```
| github_jupyter |
# Prerequisites
1. Workspace & Compute exists, and is created outside of this notebook
2. A Tabular Dataset exists, and is created and registered outside of this notebook
3. Pip requirements:
- dpv2-sdk (for creating and accessing AzureML resources)
- azureml-mlflow (for loading the tracking URI)
# Setup
These environment variables enable private preview features, such as AutoML
```
%env AZURE_EXTENSION_DIR=/home/schrodinger/automl/sdk-cli-v2/src/cli/src
%env AZURE_ML_CLI_PRIVATE_FEATURES_ENABLED=true
```
## Imports
```
from azure.ml import MLClient
import mlflow
```
## Setting necessary context
```
subscription_id = '381b38e9-9840-4719-a5a0-61d9585e1e91'
resource_group_name = 'gasi_rg_centraleuap'
# The workspace under which to log experiments and trials
workspace_name = "gasi_ws_centraleuap"
# The experiment under which AutoML will track its trials and artifacts
experiment_name = "automl-classification-bmarketing-all"
# The compute target where AutoML will execute its trials
compute_name = "cpu-cluster"
# The datasets along with their versions
training_dataset = "bankmarketing_train:1"
test_dataset = "bankmarketing_test:1"
validation_dataset = "bankmarketing_valid:1"
```
### Question:
Append 'azureml:...' to the dataset names above?
## Initialize MLClient
Create an MLClient object - which is used to manage all Azure ML resources, such as workspaces, jobs, models, etc.
```
client = MLClient(subscription_id, resource_group_name, workspace_name=workspace_name)
assert client is not None
```
## Initialize MLFlow Client
The models and artifacts that are produced by AutoML can be accessed by the MLFlow interface. Initialize the MLFlow client here, and set the backend as Azure ML, via. the MLFlow Client.
```
tracking_uri = "TODO --> Get this from MLClient"
################################################################################
# TODO: The API to get tracking URI is not yet available on Worksapce object.
from azureml.core import Workspace as WorkspaceV1
ws = WorkspaceV1(workspace_name=workspace_name, resource_group=resource_group_name, subscription_id=subscription_id)
tracking_uri = ws.get_mlflow_tracking_uri()
del ws
################################################################################
mlflow.set_tracking_uri(tracking_uri)
mlflow.set_experiment(experiment_name)
print("\nCurrent tracking uri: {}".format(mlflow.get_tracking_uri()))
```
### Questions
Q: Can we set this (the tracking URI) inside AutoML, given things won't work at all w/o setting MLFlow context above?
Q: Do we need MLFlow client for job submissions?
# AutoML Job
## Job Configuration - A minimal example
Using default primary metric (accuracy for classification)
```
from azure.ml.entities import AutoMLJob
automl_job = AutoMLJob(
compute=compute_name,
task="classification",
target="y",
dataset={"train": training_dataset, "test": test_dataset, "valid": validation_dataset},
configuration={
"blocked_models": ["KNN", "LinearSVM"],
"exit_criterion": {"timeout_hours": 1},
"max_concurrent_trials": 4,
"validation": {"n_cross_validations": 5},
},
properties={"save_mlflow": True} # This should be enabled by default
)
created_job = client.jobs.create_or_update(automl_job)
created_job
print("Studio URL: ", created_job.interaction_endpoints["Studio"].endpoint)
```
## Alternate Job Configurations
### Enable ONNX compatible Models
Shows a mixed use of promoted properties along with the original entity (TrainingSettings)
```
from azure.ml.entities import AutoMLJob
from azure.ml.entities._job.automl.training_settings import TrainingSettings
training_settings = TrainingSettings(enable_onnx_compatible_models=True)
automl_job = AutoMLJob(
compute=compute_name,
task="classification",
target="y",
dataset={"train": training_dataset, "test": test_dataset, "valid": validation_dataset},
training_settings = training_settings,
configuration={
"blocked_models": ["KNN", "LinearSVM"],
"exit_criterion": {"timeout_hours": 1},
"max_concurrent_trials": 4,
"validation": {"n_cross_validations": 5},
},
properties={"save_mlflow": True}
)
automl_job.training_settings.__dict__
```
### Use a non-default primary metric
```
from azure.ml.entities import AutoMLJob
from azure.ml._restclient.v2020_09_01_preview.models import GeneralSettings
general_settings = GeneralSettings(primary_metric= "auc_weighted")
automl_job = AutoMLJob(
compute=compute_name,
task="classification",
target="y",
dataset={"train": training_dataset, "test": test_dataset, "valid": validation_dataset},
general_settings = general_settings,
configuration={
"blocked_models": ["KNN", "LinearSVM"],
"exit_criterion": {"timeout_hours": 1},
"max_concurrent_trials": 4,
"validation": {"n_cross_validations": 5},
},
properties={"save_mlflow": True}
)
automl_job.general_settings.__dict__
```
### Enable Deep Neural Nets + train-valid percentage split
```
from azure.ml.entities import AutoMLJob
automl_job = AutoMLJob(
compute=compute_name,
task="classification",
target="y",
dataset={"train": training_dataset, "test": test_dataset, "valid": validation_dataset},
configuration={
"blocked_models": ["KNN", "LinearSVM"],
"exit_criterion": {"timeout_hours": 1},
"max_concurrent_trials": 4,
"enable_dnn": True,
"validation": {"valid_percent": 0.2},
},
properties={"save_mlflow": True} # This should be enabled by default
)
automl_job.training_settings.__dict__, "-----", automl_job.data_settings.validation_data.__dict__
```
### Disable Ensembling
```
from azure.ml.entities import AutoMLJob
from azure.ml.entities._job.automl.training_settings import TrainingSettings
training_settings = TrainingSettings(
enable_vote_ensemble=False, enable_stack_ensemble=False
)
automl_job = AutoMLJob(
compute=compute_name,
task="classification",
target="y",
dataset={"train": training_dataset, "test": test_dataset, "valid": validation_dataset},
training_settings = training_settings,
configuration={
"blocked_models": ["KNN", "LinearSVM"],
"exit_criterion": {"timeout_hours": 1},
"max_concurrent_trials": 4,
"validation": {"n_cross_validations": 5},
"enable_dnn": True,
},
properties={"save_mlflow": True}
)
automl_job.training_settings.__dict__
```
### Forecasting
```
from azure.ml.entities import AutoMLJob
from azure.ml.entities._job.automl.forecasting import ForecastingSettings
forecast_settings = ForecastingSettings(
time_column_name="DATE", forecast_horizon=12, frequency='MS'
)
automl_job = AutoMLJob(
compute=compute_name,
task="forecasting",
target="BeerProduction",
dataset={"train": training_dataset, "test": test_dataset, "valid": validation_dataset},
forecasting_settings = forecast_settings,
configuration={
"blocked_models": ["KNN", "LinearSVM"],
"exit_criterion": {"timeout_hours": 1},
"max_concurrent_trials": 4,
"validation": {"n_cross_validations": 5},
"enable_dnn": True,
},
properties={"save_mlflow": True}
)
automl_job.forecasting_settings.__dict__
```
### Custom Featurization Settings
```
from azure.ml.entities._job.automl.featurization import ColumnTransformer, FeaturizationConfig
featurization_config = FeaturizationConfig()
featurization_config.blocked_transformers = ['LabelEncoder']
featurization_config.drop_columns = ['MMIN']
featurization_config.column_purposes = {
'MYCT': 'Numeric',
'VendorName': 'CategoricalHash'
}
#default strategy mean, add transformer param for for 3 columns
transformer_params_dict = {
"Imputer": [
ColumnTransformer(fields=["CACH"], parameters={'strategy': 'median'}),
ColumnTransformer(fields=["CHMIN"], parameters={'strategy': 'median'}),
ColumnTransformer(fields=["PRP"], parameters={'strategy': 'most_frequent'}),
],
"HashOneHotEncoder": [
ColumnTransformer(fields=[], parameters={'number_of_bits': 3.0})
]
}
featurization_config.transformer_params = transformer_params_dict
from azure.ml.entities import AutoMLJob
from azure.ml.entities._job.automl.featurization import FeaturizationSettings
featurization_settings = FeaturizationSettings(featurization_config=featurization_config)
automl_job = AutoMLJob(
compute=compute_name,
task="classification",
target="y",
dataset={"train": training_dataset, "test": test_dataset, "valid": validation_dataset},
featurization_settings=featurization_settings,
configuration={
"blocked_models": ["KNN", "LinearSVM"],
"exit_criterion": {"timeout_hours": 1},
"max_concurrent_trials": 4,
"validation": {"valid_percent": 0.2},
},
properties={"save_mlflow": True} # This should be enabled by default
)
automl_job.featurization_settings.featurization_config.__dict__
```
| github_jupyter |
```
from pydrive.auth import GoogleAuth
from pydrive.drive import GoogleDrive
from google.colab import auth
from oauth2client.client import GoogleCredentials
# Authenticate and create the PyDrive client.
auth.authenticate_user()
gauth = GoogleAuth()
gauth.credentials = GoogleCredentials.get_application_default()
drive = GoogleDrive(gauth)
#This part linkes the dataset in the drive to the google colab file
link = 'https://drive.google.com/file/d/1yzZTYQXxJ2KmUfBddh4N63DoFholzsuV/view?usp=sharing' #Moving Average
link2 = 'https://drive.google.com/file/d/1e1LUjGWbSo3FAVJkl-YrK-mIwRwTZG9I/view?usp=sharing' #Actual Prices
link3 = 'https://drive.google.com/file/d/16wCdkMXOxIh75QjYuN9vxlNo2RuF2MSr/view?usp=sharing' #PCA Predicted
link4 = 'https://drive.google.com/file/d/1-PzbGALW502qtO4VOUOahtqWR3IUJSNC/view?usp=sharing' #LR Predicted
link5 = 'https://drive.google.com/file/d/1-1pSSwP8xzLh2MDx29nU3bPCNcSmHL8X/view?usp=sharing' #PCA Actuals
link6 = 'https://drive.google.com/file/d/1jrbnWW45MeEPq3JEox9sP2tOyGEdBzAo/view?usp=sharing' #LR Actuals
id = link.split('/')[-2]
id2 = link2.split('/')[-2]
id3 = link3.split('/')[-2]
id4 = link4.split('/')[-2]
id5 = link5.split('/')[-2]
id6 = link6.split('/')[-2]
# id7 = link7.split('/')[-2]
downloaded = drive.CreateFile({'id' : id})
downloaded2 = drive.CreateFile({'id' : id2})
downloaded3 = drive.CreateFile({'id' : id3})
downloaded4 = drive.CreateFile({'id' : id4})
downloaded5 = drive.CreateFile({'id' : id5})
downloaded6 = drive.CreateFile({'id' : id6})
downloaded.GetContentFile('MovingAverage.csv')
downloaded.GetContentFile('y_dataset.csv')
downloaded3.GetContentFile('PCA_Predicted_Prices.csv')
downloaded4.GetContentFile('LR_Predicted_Prices.csv')
downloaded5.GetContentFile('PCA_Actual_Prices.csv')
downloaded6.GetContentFile('LR_Actual_prices.csv')
#S&P 500 Data
link = 'https://drive.google.com/file/d/1MMKb7NW7wfggpfpAm8xS2UFwzomjZJju/view?usp=sharing'
id7 = link.split('/')[-2]
downloaded7 = drive.CreateFile({'id' : id7})
downloaded7.GetContentFile('SP500.csv')
SP500 = pd.read_csv("SP500.csv")
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
#Model 1: Moving Average
Moving_Average = pd.read_csv('MovingAverage.csv')
Moving_Average['Date'] = pd.to_datetime(Moving_Average['Date'])
Moving_Average=Moving_Average.set_index('Date')
Closing_Prices = pd.read_csv('y_dataset.csv')
Closing_Prices['Date'] = pd.to_datetime(Closing_Prices['Date'])
Closing_Prices=Closing_Prices.set_index('Date')
#Model 2: Linear Regression
LR_Predicted_Prices = pd.read_csv('LR_Predicted_Prices.csv')
LR_Predicted_Prices['Date'] = pd.to_datetime(LR_Predicted_Prices['Date'])
LR_Predicted_Prices=LR_Predicted_Prices.set_index('Date')
LR_Actual_Prices = pd.read_csv('LR_Actual_prices.csv')
LR_Actual_Prices['Date'] = pd.to_datetime(LR_Actual_Prices['Date'])
LR_Actual_Prices=LR_Actual_Prices.set_index('Date')
#Model 3: LSTM
PCA_Predicted_Prices = pd.read_csv('PCA_Predicted_Prices.csv')
PCA_Predicted_Prices['Date'] = pd.to_datetime(PCA_Predicted_Prices['Date'])
PCA_Predicted_Prices=PCA_Predicted_Prices.set_index('Date')
PCA_Actual_Prices = pd.read_csv('PCA_Actual_Prices.csv')
PCA_Actual_Prices['Date'] = pd.to_datetime(PCA_Actual_Prices['Date'])
PCA_Actual_Prices=PCA_Actual_Prices.set_index('Date')
#Computing Log Returns
PCA_Predicted_Returns = PCA_Predicted_Prices.apply(lambda x: np.log(x) - np.log(x.shift(1))).iloc[1:]
PCA_Actual_Returns = PCA_Actual_Prices.apply(lambda x: np.log(x) - np.log(x.shift(1))).iloc[1:]
LR_Predicted_Returns = LR_Predicted_Prices.apply(lambda x: np.log(x) - np.log(x.shift(1))).iloc[1:]
LR_Actual_Returns = LR_Actual_Prices.apply(lambda x: np.log(x) - np.log(x.shift(1))).iloc[1:]
Closing_Prices_Returns = Closing_Prices.apply(lambda x: np.log(x) - np.log(x.shift(1))).iloc[1:]
from sklearn.metrics import mean_squared_error
mean_squared_error(PCA_Actual_Prices, PCA_Predicted_Prices)
```
# Creation of Different Lookback and Forward Windows
```
#Mean Returns function
def mean_returns(df, length):
mu = df.sum(axis = 0)/length
return mu
from datetime import timedelta
from dateutil.parser import parse
def monthdelta(date, delta):
m, y = (date.month+delta) % 12, date.year + ((date.month)+delta-1) // 12
if not m: m = 12
d = min(date.day, [31,
29 if y%4==0 and not y%400==0 else 28,31,30,31,30,31,31,30,31,30,31][m-1])
new_date = (date.replace(day=d,month=m, year=y))
return parse(new_date.strftime('%Y-%m-%d'))
#This part of the code takes in a dataset and splits it into datasets w/ lookback months and forward looking months
def windowGenerator (dataframe, lookback, horizon,
step, cummulative = False):
#takes pandas dataframe with DatetimeIndex
if cummulative:
c = lookback
step = horizon
initial = min(dataframe.index)
windows = []
horizons = []
while initial <= monthdelta(max(dataframe.index), -lookback):
windowStart = initial
windowEnd = monthdelta(windowStart, lookback)
if cummulative:
windowStart = min(dataframe.index)
windowEnd = monthdelta(windowStart, c) + timedelta(days=1)
c += horizon
horizonStart = windowEnd + timedelta(days=1)
horizonEnd = monthdelta(horizonStart, horizon)
lookbackWindow = dataframe[windowStart:windowEnd]
horizonWindow = dataframe[horizonStart:horizonEnd]
windows.append(lookbackWindow)
horizons.append(horizonWindow)
initial = monthdelta(initial, step)
return windows, horizons
```
# Optimize Function
```
from scipy.optimize import minimize, Bounds, LinearConstraint
from numpy.linalg import norm
def actual_return(actual_returns, w):
actual_returns = actual_returns
mean_return = mean_returns(actual_returns, actual_returns.shape[0])
actual_covariance = actual_returns.cov()
portfolio_returns = mean_return.T.dot(w)
portfolio_variance = w.T.dot(actual_covariance).dot(w)
return portfolio_returns, portfolio_variance
#Input entire predicted returns df, actual returns df, starting date
def scipy_opt(predicted_returns, actual_returns, lam1, lam2):
mean_return = mean_returns(predicted_returns, predicted_returns.shape[0])
predicted_covariance = predicted_returns.cov()
#Cost Function
def f(w):
return -(mean_return.T.dot(w) - lam1*(w.T.dot(predicted_covariance).dot(w)) + lam2*norm(w, ord=1))
#out custom maximises
#Bounds of Weights
opt_bounds = Bounds(0, 1)
#Equality Constraints
def h(w):
return sum(w) - 1
#Constraints Dictionary
cons = ({
'type' : 'eq',
'fun' : lambda w: h(w)
})
#Solver
sol = minimize(f,
x0 = np.ones(mean_return.shape[0]),
constraints = cons,
bounds = opt_bounds,
options = {'disp': False},
tol=10e-10)
#Predicted Results
w = sol.x
predicted_portfolio_returns = w.dot(mean_return)
portfolio_STD = w.T.dot(predicted_covariance).dot(w)
#Actual Results
portfolio_actual_returns, portfolio_actual_variance = actual_return(actual_returns, w)
sharpe_ratio = portfolio_actual_returns/np.std(portfolio_actual_variance)
ret_dict = {'weights' : w,
'predicted_returns' : predicted_portfolio_returns,
'predicted_variance' : portfolio_STD,
'actual_returns' : portfolio_actual_returns,
'actual_variance' : portfolio_actual_variance,
'sharpe_ratio': sharpe_ratio}
return ret_dict
```
# Method 1: Moving Average
```
#Moving average only needs to look at historical mean returns to make a prediction for the future. It
#essentially follows the trend of our lookback window
MA_act_windows, MA_act_horizons = windowGenerator(Closing_Prices_Returns, 12, 1, 1)
MA_scipy_returns= []
MA_scipy_variance = []
MA_scipy_SR = []
# Testing on 5 years of data
for i in range(len(LR_act_horizons)-72,len(LR_act_horizons)-12):
#Scipy optimize results
scipy = scipy_opt(MA_act_windows[i], MA_act_horizons[i], .5, 2)
MA_scipy_returns.append(scipy['actual_returns'])
MA_scipy_variance.append(scipy['actual_variance'])
MA_scipy_SR.append(scipy['sharpe_ratio'])
# print("Month " + str(i) + " complete")
timestamps = []
for i in range(len(MA_act_horizons)-72,len(MA_act_horizons)-12):
time = MA_act_horizons[i].index[-1]
timestamps.append(time)
MA_Portfolio_Returns = pd.DataFrame(data = np.array([MA_scipy_returns,MA_scipy_variance,MA_scipy_SR]).T, columns = ['Returns', 'Variance', 'Sharpe Ratio'], index=timestamps)
MA_Portfolio_Returns.index.rename('Date')
MA_Portfolio_Returns.to_csv('MA_Portfolio_Returns.csv')
!cp MA_Portfolio_Returns.csv "drive/My Drive/Machine Learning Project/ML Section Exports"
MA_Portfolio_Returns.head()
```
# Method 2: PCA + Linear Regression
```
LR_pred_windows, LR_pred_horizons = windowGenerator(LR_Predicted_Returns, 12, 1, 1)
LR_act_windows, LR_act_horizons = windowGenerator(LR_Actual_Returns, 12, 1, 1)
LR_scipy_returns= []
LR_scipy_variance = []
LR_scipy_SR = []
#Testing on 5 years of data
for i in range(len(LR_act_horizons)-72,len(LR_act_horizons)-12):
#Scipy optimize results
scipy = scipy_opt(LR_pred_horizons[i], LR_act_horizons[i], .5, 2)
LR_scipy_returns.append(scipy['actual_returns'])
LR_scipy_variance.append(scipy['actual_variance'])
LR_scipy_SR.append(scipy['sharpe_ratio'])
# print("Month " + str(i) + " complete")
timestamps = []
for i in range(len(LR_act_horizons)-72,len(LR_act_horizons)-12):
time = LR_act_horizons[i].index[-1]
timestamps.append(time)
LR_Portfolio_Returns = pd.DataFrame(data = np.array([LR_scipy_returns,LR_scipy_variance,LR_scipy_SR]).T, columns = ['Returns', 'Variance', 'Sharpe Ratio'], index=timestamps)
## Exporting the Dataset
from google.colab import drive
drive.mount('drive')
LR_Portfolio_Returns.to_csv('LR_Portfolio_Returns.csv')
!cp LR_Portfolio_Returns.csv "drive/My Drive/Machine Learning Project/ML Section Exports"
```
# Method 3: LSTM + PCA
```
LSTM_pred_windows, LSTM_pred_horizons = windowGenerator(PCA_Predicted_Returns,12,1,1)
LSTM_act_windows, LSTM_act_horizons = windowGenerator(PCA_Actual_Returns,12,1,1)
len(LSTM_pred_horizons)
scipy_returns= []
scipy_variance = []
scipy_SR = []
for i in range(len(LSTM_act_horizons)-60,len(LSTM_act_horizons)):
#Scipy optimize results
scipy = scipy_opt(LSTM_pred_horizons[i], LSTM_act_horizons[i], .5, 2)
scipy_returns.append(scipy['actual_returns'])
scipy_variance.append(scipy['actual_variance'])
scipy_SR.append(scipy['sharpe_ratio'])
print("Month " + str(i) + " complete")
timestamps = []
for i in range(len(LSTM_act_horizons)-60,len(LSTM_act_horizons)):
time = LSTM_act_horizons[i].index[0]
timestamps.append(time)
LSTM_Portfolio_Returns = pd.DataFrame(data = np.array([scipy_returns,scipy_variance,scipy_SR]).T, columns = ['Returns', 'Variance', 'Sharpe Ratio'], index=timestamps)
LSTM_Portfolio_Returns.to_csv('LSTM_Portfolio_Returns.csv')
!cp LSTM_Portfolio_Returns.csv "drive/My Drive/Machine Learning Project/ML Section Exports"
timestamps
LSTM_Portfolio_Returns
```
# Diagnostics
The calculation for seeing how much the portfolio grows in dollar terms:
$Price_t * e^{LogReturns} = Price_{t+1}$
- Given $100, our equity graph shows how much our portfolio value increase (or decreases)
```
import math
MA_equity = [100]
LR_equity = [100]
LSTM_equity = [100]
#This is the calculation for the for seeing how much the portfolio grows
for i in range(1,60):
MA_equity.append(MA_equity[i-1]* math.exp(MA_scipy_returns[i]))
LR_equity.append(LR_equity[i-1]* math.exp(LR_scipy_returns[i]))
LSTM_equity.append(LSTM_equity[i-1]* math.exp(scipy_returns[i]))
plt.plot(timestamps, MA_equity, label = "Moving Average")
plt.plot(timestamps, LR_equity, label = "Linear Regression")
plt.plot(timestamps, LSTM_equity, label = "LSTM")
plt.title("Equity Graph")
plt.legend()
plt.show();
plt.hist(LR_scipy_returns, bins = 60, label = 'returns', alpha = 1, color = 'green')
plt.title("Moving Average Portfolio Distribution")
plt.legend()
plt.show();
plt.hist(MA_scipy_returns, bins = 60, label = 'returns', alpha = 1, color = 'red')
plt.title("Linear Regression Portfolio Distribution")
plt.legend()
plt.show();
plt.hist(scipy_returns, bins = 60, label = 'returns', alpha = 1, color = 'blue')
plt.title("LSTM Portfolio Returns Distribution")
plt.legend()
plt.show();
print("Moving Average Ending Equity: " , MA_equity[-1])
print("Linear Regression Ending Equity: " , LR_equity[-1])
print("LSTM Ending Equity" , LSTM_equity[-1])
import math
def metrics(returns):
sharpe = returns.mean() / returns.std()
annualized_sharpe = sharpe.item() / math.sqrt(252)
stdev = returns.std()
annualized_vol = stdev.item() / math.sqrt(252)
return {"Annualized Sharpe Ratio": annualized_sharpe,
"Annualized Volatility": annualized_vol}
#Annualized info for Moving Average
metrics(np.array(MA_scipy_returns))
#Annualized info for Linear Regression
metrics(np.array(LR_scipy_returns))
#Annualized info for LSTM
metrics(np.array(scipy_returns))
```
$$
| github_jupyter |
<!-- dom:TITLE: Statistical physics -->
# Statistical physics
<!-- dom:AUTHOR: Morten Hjorth-Jensen Email morten.hjorth-jensen@fys.uio.no at Department of Physics and Center of Mathematics for Applications, University of Oslo & National Superconducting Cyclotron Laboratory, Michigan State University -->
<!-- Author: --> **Morten Hjorth-Jensen Email morten.hjorth-jensen@fys.uio.no**, Department of Physics and Center of Mathematics for Applications, University of Oslo and National Superconducting Cyclotron Laboratory, Michigan State University
Date: **Fall 2015**
## Ensembles
In statistical physics the concept of an ensemble is one of the
cornerstones in the definition of thermodynamical quantities. An
ensemble is a collection of microphysics systems from which we derive
expectations values and thermodynamical properties related to
experiment. As an example, the specific heat (which is a measurable
quantity in the laboratory) of a system of infinitely many particles,
can be derived from the basic interactions between the microscopic
constituents. The latter can span from electrons to atoms and
molecules or a system of classical spins. All these microscopic
constituents interact via a well-defined interaction. We say
therefore that statistical physics bridges the gap between the
microscopic world and the macroscopic world. Thermodynamical
quantities such as the specific heat or net magnetization of a system
can all be derived from a microscopic theory.
## Famous Ensembles
The table lists the most used ensembles in statistical physics
together with frequently arising extensive (depend on the size of the
systems such as the number of particles) and intensive variables
(apply to all components of a system), in addition to associated
potentials.
<table border="1">
<tr></tr>
<tbody>
<tr><td align="left"> </td> <td align="center"> Microcanonical </td> <td align="center"> Canonical </td> <td align="center"> </td> </tr>
<tr><td align="left"> Grand canonical </td> <td align="center"> Pressure canonical </td> </tr>
<tr><td align="left"> </td> <td align="center"> </td> <td align="center"> </td> <td align="center"> </td> <td align="center"> </td> </tr>
<tr><td align="left"> </td> <td align="center"> </td> <td align="center"> </td> <td align="center"> </td> <td align="center"> </td> </tr>
<tr><td align="left"> Exchange of heat </td> <td align="center"> no </td> <td align="center"> yes </td> <td align="center"> yes </td> <td align="center"> yes </td> </tr>
<tr><td align="left"> with the environment </td> <td align="center"> </td> <td align="center"> </td> <td align="center"> </td> <td align="center"> </td> </tr>
<tr><td align="left"> </td> <td align="center"> </td> <td align="center"> </td> <td align="center"> </td> <td align="center"> </td> </tr>
<tr><td align="left"> Exchange of particles </td> <td align="center"> no </td> <td align="center"> no </td> <td align="center"> yes </td> <td align="center"> no </td> </tr>
<tr><td align="left"> with the environemt </td> <td align="center"> </td> <td align="center"> </td> <td align="center"> </td> <td align="center"> </td> </tr>
<tr><td align="left"> </td> <td align="center"> </td> <td align="center"> </td> <td align="center"> </td> <td align="center"> </td> </tr>
<tr><td align="left"> Thermodynamical </td> <td align="center"> $V, \cal M, \cal D$ </td> <td align="center"> $V, \cal M, \cal D$ </td> <td align="center"> $V, \cal M, \cal D$ </td> </tr>
<tr><td align="left"> </td> <td align="center"> $P, \cal H, \cal E$ </td> </tr>
<tr><td align="left"> parameters </td> <td align="center"> $E$ </td> <td align="center"> $T$ </td> <td align="center"> $T$ </td> <td align="center"> $T$ </td> </tr>
<tr><td align="left"> </td> <td align="center"> $N$ </td> <td align="center"> $N$ </td> <td align="center"> $\mu$ </td> <td align="center"> $N$ </td> </tr>
<tr><td align="left"> </td> <td align="center"> </td> <td align="center"> </td> <td align="center"> </td> <td align="center"> </td> </tr>
<tr><td align="left"> Potential </td> <td align="center"> Entropy </td> <td align="center"> Helmholtz </td> <td align="center"> $PV$ </td> <td align="center"> Gibbs </td> </tr>
<tr><td align="left"> </td> <td align="center"> </td> <td align="center"> </td> <td align="center"> </td> <td align="center"> </td> </tr>
<tr><td align="left"> Energy </td> <td align="center"> Internal </td> <td align="center"> Internal </td> <td align="center"> Internal </td> <td align="center"> Enthalpy </td> </tr>
<tr><td align="left"> </td> <td align="center"> </td> <td align="center"> </td> <td align="center"> </td> <td align="center"> </td> </tr>
<tr><td align="left"> </td> <td align="center"> </td> <td align="center"> </td> <td align="center"> </td> <td align="center"> </td> </tr>
<tr><td align="left"> </td> <td align="center"> </td> <td align="center"> </td> <td align="center"> </td> <td align="center"> </td> </tr>
</tbody>
</table>
## Canonical Ensemble
One of the most used ensembles is the canonical one, which is related to the microcanonical ensemble
via a Legendre transformation. The temperature is an intensive variable in this ensemble whereas the energy
follows as an expectation value.
In order to calculate expectation values such as the mean energy $\langle E \rangle $
at a given temperature, we need a probability distribution.
It is given by the Boltzmann distribution
$$
P_i(\beta) = \frac{e^{-\beta E_i}}{Z}
$$
with $\beta=1/k_BT$ being the inverse temperature, $k_B$ is the
Boltzmann constant, $E_i$ is the energy of a microstate $i$ while
$Z$ is the partition function for the canonical ensemble
defined as
## The partition function is a normalization constant
In the canonical ensemble the partition function is
$$
Z=\sum_{i=1}^{M}e^{-\beta E_i},
$$
where the sum extends over all microstates $M$.
## Helmoltz free energy, what does it mean?
The potential of interest in this case is Helmholtz' free energy. It
relates the expectation value of the energy at a given temperatur $T$
to the entropy at the same temperature via
$$
F=-k_{B}TlnZ=\langle E \rangle-TS.
$$
Helmholtz' free energy expresses the
struggle between two important principles in physics, namely the
strive towards an energy minimum and the drive towards higher entropy
as the temperature increases. A higher entropy may be interpreted as a
larger degree of disorder. When equilibrium is reached at a given
temperature, we have a balance between these two principles. The
numerical expression is Helmholtz' free energy.
## Thermodynamical quantities
In the canonical ensemble the entropy is given by
$$
S =k_{B}lnZ
+k_{B}T\left(\frac{\partial lnZ}{\partial T}\right)_{N, V},
$$
and the pressure by
$$
p=k_{B}T\left(\frac{\partial lnZ}{\partial V}\right)_{N, T}.
$$
Similarly we can compute the chemical potential as
$$
\mu =-k_{B}T\left(\frac{\partial lnZ}{\partial N}\right)_{V, T}.
$$
## Thermodynamical quantities, the energy in the canonical ensemble
For a system described by the canonical ensemble, the energy is an
expectation value since we allow energy to be exchanged with the surroundings
(a heat bath with temperature $T$).
This expectation value, the mean energy,
can be calculated using
$$
\langle E\rangle =k_{B}T^{2}\left(\frac{\partial lnZ}{\partial T}\right)_{V, N}
$$
or
using the probability distribution
$P_i$ as
$$
\langle E \rangle = \sum_{i=1}^M E_i P_i(\beta)=
\frac{1}{Z}\sum_{i=1}^M E_ie^{-\beta E_i}.
$$
## Energy and specific heat in the canonical ensemble
The energy is proportional to the first derivative of the potential,
Helmholtz' free energy. The corresponding variance is defined as
$$
\sigma_E^2=\langle E^2 \rangle-\langle E \rangle^2=
\frac{1}{Z}\sum_{i=1}^M E_i^2e^{-\beta E_i}-
\left(\frac{1}{Z}\sum_{i=1}^M E_ie^{-\beta E_i}\right)^2.
$$
If we divide the latter quantity with
$kT^2$ we obtain the specific heat at constant volume
$$
C_V= \frac{1}{k_BT^2}\left(\langle E^2 \rangle-\langle E \rangle^2\right),
$$
which again can be related to the second derivative of Helmholtz' free energy.
## Magnetic moments and susceptibility in the canonical ensemble
Using the same prescription, we can also evaluate the mean magnetization
through
$$
\langle {\cal M} \rangle = \sum_i^M {\cal M}_i P_i(\beta)=
\frac{1}{Z}\sum_i^M {\cal M}_ie^{-\beta E_i},
$$
and the corresponding variance
$$
\sigma_{{\cal M}}^2=\langle {\cal M}^2 \rangle-\langle {\cal M} \rangle^2=
\frac{1}{Z}\sum_{i=1}^M {\cal M}_i^2e^{-\beta E_i}-
\left(\frac{1}{Z}\sum_{i=1}^M {\cal M}_ie^{-\beta E_i}\right)^2.
$$
This quantity defines also the susceptibility
$\chi$
$$
\chi=\frac{1}{k_BT}\left(\langle {\cal M}^2 \rangle-\langle {\cal M} \rangle^2\right).
$$
## Our model, the Ising model in one and two dimensions
The model we will employ in our studies of phase transitions at finite temperature for
magnetic systems is the so-called Ising model. In its simplest form
the energy is expressed as
$$
E=-J\sum_{<kl>}^{N}s_ks_l-{\cal B}\sum_k^Ns_k,
$$
with $s_k=\pm 1$, $N$ is the total number of spins,
$J$ is a coupling constant expressing the strength of the interaction
between neighboring spins and
${\cal B}$ is an external magnetic field interacting with the magnetic
moment set up by the spins.
The symbol $<kl>$ indicates that we sum over nearest
neighbors only.
Notice that for $J>0$ it is energetically favorable for neighboring spins
to be aligned. This feature leads to, at low enough temperatures,
a cooperative phenomenon called spontaneous magnetization. That is,
through interactions between nearest neighbors, a given magnetic
moment can influence the alignment of spins that are separated
from the given spin by a macroscopic distance. These long range correlations
between spins are associated with a long-range order in which
the lattice has a net magnetization in the absence of a magnetic field.
## Our model, the Ising model in one and two dimensions
In order to calculate expectation values such as the mean energy
$\langle E \rangle $ or
magnetization $\langle {\cal M} \rangle $
in statistical physics
at a given temperature, we need a probability distribution
$$
P_i(\beta) = \frac{e^{-\beta E_i}}{Z}
$$
with $\beta=1/kT$ being the inverse temperature, $k$ the
Boltzmann constant, $E_i$ is the energy of a state $i$ while
$Z$ is the partition function for the canonical ensemble
defined as
$$
Z=\sum_{i=1}^{M}e^{-\beta E_i},
$$
where the sum extends over all microstates
$M$.
$P_i$ expresses the probability of finding the system in a given
configuration $i$.
## Our model, the Ising model in one and two dimensions
The energy for a specific configuration $i$
is given by
$$
E_i =-J\sum_{<kl>}^{N}s_ks_l.
$$
## Our model, the Ising model in one and two dimensions
To better understand what is meant with a configuration,
consider first the case of the one-dimensional Ising model
with ${\cal B}=0$.
In general, a given configuration of $N$ spins in one
dimension may look like
$$
\begin{array}{cccccccccc}
\uparrow&\uparrow&\uparrow&\dots&\uparrow&\downarrow&\uparrow&\dots&\uparrow&\downarrow\\
1&2&3&\dots& i-1&i&i+1&\dots&N-1&N\end{array}
$$
In order to illustrate these features
let us further specialize to
just two spins.
With two spins, since each spin takes two values only,
we have $2^2=4$ possible arrangements of the two spins.
These four possibilities are
$$
1= \uparrow\uparrow\hspace{1cm}
2= \uparrow\downarrow\hspace{1cm}
3= \downarrow\uparrow\hspace{1cm}
4=\downarrow\downarrow
$$
## Our model, the Ising model in one and two dimensions
What is the energy of each of these configurations?
For small systems, the way we treat the ends matters. Two cases are
often used.
In the first case we employ what is called free ends. This means that there is no contribution from points to the right or left of the endpoints. For the one-dimensional case, the energy is then written as a sum over a single index
$$
E_i =-J\sum_{j=1}^{N-1}s_js_{j+1},
$$
## Our model, the Ising model in one and two dimensions
If we label the first spin as $s_1$ and the second as $s_2$
we obtain the following
expression for the energy
$$
E=-Js_1s_2.
$$
The calculation of the energy for the one-dimensional lattice
with free ends for one specific spin-configuration
can easily be implemented in the following lines
for ( j=1; j < N; j++) {
energy += spin[j]*spin[j+1];
}
where the vector $spin[]$ contains the spin value $s_k=\pm 1$.
## Our model, the Ising model in one and two dimensions
For the specific state $E_1$, we have chosen all spins up. The energy of
this configuration becomes then
$$
E_1=E_{\uparrow\uparrow}=-J.
$$
The other configurations give
$$
E_2=E_{\uparrow\downarrow}=+J,
$$
$$
E_3=E_{\downarrow\uparrow}=+J,
$$
and
$$
E_4=E_{\downarrow\downarrow}=-J.
$$
## Our model, the Ising model in one and two dimensions
We can also choose so-called periodic boundary conditions. This means
that the neighbour to the right of $s_N$ is assumed to take the value
of $s_1$. Similarly, the neighbour to the left of $s_1$ takes the
value $s_N$. In this case the energy for the one-dimensional lattice
reads
$$
E_i =-J\sum_{j=1}^{N}s_js_{j+1},
$$
and we obtain the following expression for the
two-spin case
$$
E=-J(s_1s_2+s_2s_1).
$$
## Our model, the Ising model in one and two dimensions
In this case the energy for $E_1$ is different, we obtain namely
$$
E_1=E_{\uparrow\uparrow}=-2J.
$$
The other cases do also differ and we have
$$
E_2=E_{\uparrow\downarrow}=+2J,
$$
$$
E_3=E_{\downarrow\uparrow}=+2J,
$$
and
$$
E_4=E_{\downarrow\downarrow}=-2J.
$$
## Our model, the Ising model in one and two dimensions
If we choose to use periodic boundary conditions we can code the above
expression as
jm=N;
for ( j=1; j <=N ; j++) {
energy += spin[j]*spin[jm];
jm = j ;
}
The magnetization is however the same, defined as
$$
{\cal M}_i=\sum_{j=1}^N s_j,
$$
where we sum over all spins for a given configuration $i$.
## Our model, the Ising model in one and two dimensions
The table lists the energy and magnetization for both free ends
and periodic boundary conditions.
<table border="1">
<thead>
<tr><th align="center"> State </th> <th align="center">Energy (FE)</th> <th align="center">Energy (PBC)</th> <th align="center">Magnetization</th> </tr>
</thead>
<tbody>
<tr><td align="center"> $1= \uparrow\uparrow$ </td> <td align="right"> $-J$ </td> <td align="right"> $-2J$ </td> <td align="right"> 2 </td> </tr>
<tr><td align="center"> $2=\uparrow\downarrow$ </td> <td align="right"> $J$ </td> <td align="right"> $2J$ </td> <td align="right"> 0 </td> </tr>
<tr><td align="center"> $ 3=\downarrow\uparrow$ </td> <td align="right"> $J$ </td> <td align="right"> $2J$ </td> <td align="right"> 0 </td> </tr>
<tr><td align="center"> $ 4=\downarrow\downarrow$ </td> <td align="right"> $-J$ </td> <td align="right"> $-2J$ </td> <td align="right"> -2 </td> </tr>
</tbody>
</table>
## Our model, the Ising model in one and two dimensions
We can reorganize according to the number of spins pointing up, as shown in the table here
<table border="1">
<thead>
<tr><th align="center">Number spins up</th> <th align="center">Degeneracy</th> <th align="center">Energy (FE)</th> <th align="center">Energy (PBC)</th> <th align="center">Magnetization</th> </tr>
</thead>
<tbody>
<tr><td align="left"> 2 </td> <td align="left"> 1 </td> <td align="right"> $-J$ </td> <td align="right"> $-2J$ </td> <td align="right"> 2 </td> </tr>
<tr><td align="left"> 1 </td> <td align="left"> 2 </td> <td align="right"> $J$ </td> <td align="right"> $2J$ </td> <td align="right"> 0 </td> </tr>
<tr><td align="left"> 0 </td> <td align="left"> 1 </td> <td align="right"> $-J$ </td> <td align="right"> $-2J$ </td> <td align="right"> -2 </td> </tr>
</tbody>
</table>
## Our model, the Ising model in one and two dimensions
It is worth noting that for small dimensions of the lattice,
the energy differs depending on whether we use
periodic boundary conditions or free ends. This means also
that the partition functions will be different, as discussed
below. In the thermodynamic limit we have $N\rightarrow \infty$,
and the final results do not depend on the kind of boundary conditions
we choose.
For a one-dimensional lattice with periodic boundary conditions,
each spin sees two neighbors. For a
two-dimensional lattice each spin sees four neighboring spins.
How many neighbors does a spin see in three dimensions?
## Our model, the Ising model in one and two dimensions
In a similar way, we could enumerate the number of states for
a two-dimensional system consisting of two spins, i.e.,
a $2\times 2$ Ising model on a square lattice with {\em periodic
boundary conditions}. In this case we have a total of
$2^4=16$ states.
Some
examples of configurations with their respective energies are
listed here
$$
E=-8J\hspace{1cm}\begin{array}{cc}\uparrow & \uparrow \\
\uparrow & \uparrow\end{array}
\hspace{0.5cm}
E=0\hspace{1cm}\begin{array}{cc}\uparrow & \uparrow \\
\uparrow & \downarrow\end{array}
\hspace{0.5cm}
E=0\hspace{1cm}\begin{array}{cc}\downarrow & \downarrow \\
\uparrow & \downarrow\end{array}
\hspace{0.5cm}
E=-8J\hspace{1cm}\begin{array}{cc}\downarrow & \downarrow \\
\downarrow & \downarrow\end{array}
$$
## Our model, the Ising model in one and two dimensions
In the table here we group these configurations
according to their total energy and magnetization.
<table border="1">
<thead>
<tr><th align="center">Number spins up</th> <th align="center">Degeneracy</th> <th align="center">Energy</th> <th align="center">Magnetization</th> </tr>
</thead>
<tbody>
<tr><td align="left"> 4 </td> <td align="left"> 1 </td> <td align="right"> $-8J$ </td> <td align="right"> 4 </td> </tr>
<tr><td align="left"> 3 </td> <td align="left"> 4 </td> <td align="right"> $0$ </td> <td align="right"> 2 </td> </tr>
<tr><td align="left"> 2 </td> <td align="left"> 4 </td> <td align="right"> $0$ </td> <td align="right"> 0 </td> </tr>
<tr><td align="left"> 2 </td> <td align="left"> 2 </td> <td align="right"> $8J$ </td> <td align="right"> 0 </td> </tr>
<tr><td align="left"> 1 </td> <td align="left"> 4 </td> <td align="right"> $0$ </td> <td align="right"> -2 </td> </tr>
<tr><td align="left"> 0 </td> <td align="left"> 1 </td> <td align="right"> $-8J$ </td> <td align="right"> -4 </td> </tr>
</tbody>
</table>
## Phase Transitions and Critical Phenomena
A phase transition is marked by abrupt macroscopic changes as external
parameters are changed, such as an increase of temperature. The point
where a phase transition takes place is called a critical point.
We distinguish normally between two types of phase transitions;
first-order transitions and second-order transitions. An important
quantity in studies of phase transitions is the so-called correlation
length $\xi$ and various correlations functions like spin-spin
correlations. For the Ising model we shall show below that the
correlation length is related to the spin-correlation function, which
again defines the magnetic susceptibility. The spin-correlation
function is nothing but the covariance and expresses the degree of
correlation between spins.
## Phase Transitions and Critical Phenomena
The correlation length defines the length scale at which the overall
properties of a material start to differ from its bulk properties. It
is the distance over which the fluctuations of the microscopic degrees
of freedom (for example the position of atoms) are significantly
correlated with each other. Usually it is of the order of few
interatomic spacings for a solid. The correlation length $\xi$
depends however on external conditions such as pressure and
temperature.
<!-- !split -->
## Phase Transitions and Critical Phenomena
First order/discontinuous phase transitions are characterized by two or more
states on either side of the critical point that can coexist at the
critical point. As we pass through the critical point we observe a
discontinuous behavior of thermodynamical functions. The correlation
length is normally finite at the critical point. Phenomena such as
hysteris occur, viz. there is a continuation of state below the
critical point into one above the critical point. This continuation is
metastable so that the system may take a macroscopically long time to
readjust. A classical example is the melting of ice. It takes a
specific amount of time before all the ice has melted. The temperature
remains constant and water and ice can coexist for a macroscopic
time. The energy shows a discontinuity at the critical point,
reflecting the fact that a certain amount of heat is needed in order
to melt all the ice
## Phase Transitions and Critical Phenomena
Second order or continuous transitions are different and in general
much difficult to understand and model. The correlation length
diverges at the critical point, fluctuations are correlated over all
distance scales, which forces the system to be in a unique critical
phase. The two phases on either side of the critical point become
identical. The disappearance of a spontaneous magnetization is a
classical example of a second-order phase transitions. Structural
transitions in solids are other types of second-order phase
transitions. Strong correlations make a perturbative treatment
impossible. From a theoretical point of view, the way out is
renormalization group theory. The table lists some typical system
with their pertinent order parameters.
## Phase Transitions and Critical Phenomena
<table border="1">
<tr></tr>
<tbody>
<tr><td align="center"> System </td> <td align="center"> Transition </td> <td align="center"> Order Parameter </td> </tr>
<tr><td align="center"> </td> <td align="center"> </td> <td align="center"> </td> </tr>
<tr><td align="center"> </td> <td align="center"> </td> <td align="center"> </td> </tr>
<tr><td align="center"> Liquid-gas </td> <td align="center"> Condensation/evaporation </td> <td align="center"> Density difference $\Delta\rho=\rho_{liquid}-\rho_{gas}$ </td> </tr>
<tr><td align="center"> Binary liquid </td> <td align="center"> mixture/Unmixing </td> <td align="center"> Composition difference </td> </tr>
<tr><td align="center"> Quantum liquid </td> <td align="center"> Normal fluid/superfluid </td> <td align="center"> $<\phi>$, $\psi$ = wavefunction </td> </tr>
<tr><td align="center"> Liquid-solid </td> <td align="center"> Melting/crystallisation </td> <td align="center"> Reciprocal lattice vector </td> </tr>
<tr><td align="center"> Magnetic solid </td> <td align="center"> Ferromagnetic </td> <td align="center"> Spontaneous magnetisation $M$ </td> </tr>
<tr><td align="center"> </td> <td align="center"> Antiferromagnetic </td> <td align="center"> Sublattice magnetisation $M$ </td> </tr>
<tr><td align="center"> Dielectric solid </td> <td align="center"> Ferroelectric </td> <td align="center"> Polarization $P$ </td> </tr>
<tr><td align="center"> </td> <td align="center"> Antiferroelectric </td> <td align="center"> Sublattice polarisation $P$ </td> </tr>
<tr><td align="center"> </td> <td align="center"> </td> <td align="center"> </td> </tr>
<tr><td align="center"> </td> <td align="center"> </td> <td align="center"> </td> </tr>
</tbody>
</table>
## Phase Transitions and Critical Phenomena
Using Ehrenfest's definition of the order of a phase transition we can
relate the behavior around the critical point to various derivatives
of the thermodynamical potential. In the canonical ensemble we are
using, the thermodynamical potential is Helmholtz' free energy
$$
F= \langle E\rangle -TS = -kTln Z
$$
meaning $ lnZ = -F/kT = -F\beta$. The energy is given as the first derivative of $F$
$$
\langle E \rangle=-\frac{\partial lnZ}{\partial \beta} =\frac{\partial (\beta F)}{\partial \beta}.
$$
and the specific heat is defined via the second derivative of $F$
$$
C_V=-\frac{1}{kT^2}\frac{\partial^2 (\beta F)}{\partial\beta^2}.
$$
## Phase Transitions and Critical Phenomena
We can relate observables to various derivatives of the partition
function and the free energy. When a given derivative of the free
energy or the partition function is discontinuous or diverges
(logarithmic divergence for the heat capacity from the Ising model) we
talk of a phase transition of order of the derivative. A first-order
phase transition is recognized in a discontinuity of the energy, or
the first derivative of $F$. The Ising model exhibits a second-order
phase transition since the heat capacity diverges. The susceptibility
is given by the second derivative of $F$ with respect to external
magnetic field. Both these quantities diverge.
## The Ising Model and Phase Transitions
The Ising model in two dimensions with ${\cal B} = 0$ undergoes a
phase transition of second order. What it actually means is that below
a given critical temperature $T_C$, the Ising model exhibits a
spontaneous magnetization with $\langle {\cal M} \rangle\ne 0$. Above
$T_C$ the average magnetization is zero. The mean magnetization
approaches zero at $T_C$ with an infinite slope. Such a behavior is
an example of what are called critical phenomena. A critical
phenomenon is normally marked by one or more thermodynamical variables
which vanish above a critical point. In our case this is the mean
magnetization $\langle {\cal M} \rangle\ne 0$. Such a parameter is
normally called the order parameter.
## The Ising Model and Phase Transitions
Critical phenomena have been extensively studied in physics. One major
reason is that we still do not have a satisfactory understanding of
the properties of a system close to a critical point. Even for the
simplest three-dimensional systems we cannot predict exactly the
values of various thermodynamical variables. Simplified theoretical
approaches like mean-field models discussed below, can even predict
the wrong physics. Mean-field theory results in a second-order phase
transition for the one-dimensional Ising model, whereas we saw in the
previous section that the one-dimensional Ising model does not predict
any spontaneous magnetization at any finite temperature. The physical
reason for this can be understood from the following simple
consideration. Assume that the ground state for an $N$-spin system in
one dimension is characterized by the following configuration
## The Ising Model and Phase Transitions
It is possibleto show that the mean magnetization is given by
(for temperature below $T_C$)
$$
\langle {\cal M}(T) \rangle \sim \left(T-T_C\right)^{\beta},
$$
where $\beta=1/8$ is a so-called critical exponent. A similar relation
applies to the heat capacity
$$
C_V(T) \sim \left|T_C-T\right|^{-\alpha},
$$
and the susceptibility
$$
\chi(T) \sim \left|T_C-T\right|^{-\gamma},
$$
with $\alpha = 0$ and $\gamma = -7/4$.
## The Ising Model and Phase Transitions
Another important quantity is the correlation length, which is expected
to be of the order of the lattice spacing for $T$ is close to $T_C$. Because the spins
become more and more correlated as $T$ approaches $T_C$, the correlation
length increases as we get closer to the critical temperature. The discontinuous
behavior of the correlation $\xi$ near $T_C$ is
<!-- Equation labels as ordinary links -->
<div id="eq:xi"></div>
$$
\begin{equation}
\xi(T) \sim \left|T_C-T\right|^{-\nu}.
\label{eq:xi} \tag{1}
\end{equation}
$$
## The Ising Model and Phase Transitions
A second-order phase transition is characterized by a correlation
length which spans the whole system. The correlation length is
typically of the order of some few interatomic distances. The fact
that a system like the Ising model, whose energy is described by the
interaction between neighboring spins only, can yield correlation
lengths of macroscopic size at a critical point is still a feature
which is not properly understood. Stated differently, how can the
spins propagate their correlations so extensively when we approach the
critical point, in particular since the interaction acts only between
nearest spins? Below we will compute the correlation length via the
spin-sin correlation function for the one-dimensional Ising model.
## The Ising Model and Phase Transitions
In our actual calculations of the two-dimensional Ising model, we are however
always limited to a finite lattice and $\xi$ will
be proportional with the size of the lattice at the critical point.
Through finite size scaling relations
it is possible to relate the behavior at finite lattices with the
results for an infinitely large lattice.
The critical temperature scales then as
<!-- Equation labels as ordinary links -->
<div id="eq:tc"></div>
$$
\begin{equation}
T_C(L)-T_C(L=\infty) \propto aL^{-1/\nu},
\label{eq:tc} \tag{2}
\end{equation}
$$
with $a$ a constant and $\nu$ defined in Eq. [(1)](#eq:xi).
## The Ising Model and Phase Transitions
The correlation length for a finite lattice size can then be shown to be proportional to
$$
\xi(T) \propto L\sim \left|T_C-T\right|^{-\nu}.
$$
and if we set $T=T_C$ one can obtain the following relations for the
magnetization, energy and susceptibility for $T \le T_C$
$$
\langle {\cal M}(T) \rangle \sim \left(T-T_C\right)^{\beta}
\propto L^{-\beta/\nu},
$$
$$
C_V(T) \sim \left|T_C-T\right|^{-\gamma} \propto L^{\alpha/\nu},
$$
and
$$
\chi(T) \sim \left|T_C-T\right|^{-\alpha} \propto L^{\gamma/\nu}.
$$
## The Metropolis Algorithm and the Two-dimensional Ising Model
In our case we have as the Monte Carlo sampling function the probability
for finding the system in a state $s$ given by
$$
P_s=\frac{e^{-(\beta E_s)}}{Z},
$$
with energy $E_s$, $\beta=1/kT$ and $Z$ is a normalization constant which
defines the partition function in the canonical ensemble. As discussed
above
$$
Z(\beta)=\sum_se^{-(\beta E_s)}
$$
is difficult to compute since we need all states.
## The Metropolis Algorithm and the Two-dimensional Ising Model
In a calculation of the Ising model in two dimensions, the number of
configurations is given by $2^N$ with $N=L\times L$ the number of
spins for a lattice of length $L$. Fortunately, the Metropolis
algorithm considers only ratios between probabilities and we do not
need to compute the partition function at all. The algorithm goes as
follows
* Establish an initial state with energy $E_b$ by positioning yourself at a random configuration in the lattice
* Change the initial configuration by flipping e.g., one spin only. Compute the energy of this trial state $E_t$.
* Calculate $\Delta E=E_t-E_b$. The number of values $\Delta E$ is limited to five for the Ising model in two dimensions, see the discussion below.
* If $\Delta E \le 0$ we accept the new configuration, meaning that the energy is lowered and we are hopefully moving towards the energy minimum at a given temperature. Go to step 7.
* If $\Delta E > 0$, calculate $w=e^{-(\beta \Delta E)}$.
* Compare $w$ with a random number $r$. If
$$
r \le w,
$$
then accept the new configuration, else we keep the old configuration.
* The next step is to update various expectations values.
* The steps (2)-(7) are then repeated in order to obtain a sufficently good representation of states.
* Each time you sweep through the lattice, i.e., when you have summed over all spins, constitutes what is called a Monte Carlo cycle. You could think of one such cycle as a measurement. At the end, you should divide the various expectation values with the total number of cycles. You can choose whether you wish to divide by the number of spins or not. If you divide with the number of spins as well, your result for e.g., the energy is now the energy per spin.
## The Metropolis Algorithm and the Two-dimensional Ising Model
The crucial step is the calculation of the energy difference and the
change in magnetization. This part needs to be coded in an as
efficient as possible way since the change in energy is computed many
times. In the calculation of the energy difference from one spin
configuration to the other, we will limit the change to the flipping
of one spin only. For the Ising model in two dimensions it means that
there will only be a limited set of values for $\Delta E$. Actually,
there are only five possible values.
## The Metropolis Algorithm and the Two-dimensional Ising Model
To see this, select first a
random spin position $x,y$ and assume that this spin and its nearest
neighbors are all pointing up. The energy for this configuration is
$E=-4J$. Now we flip this spin as shown below. The energy of the new
configuration is $E=4J$, yielding $\Delta E=8J$.
$$
E=-4J\hspace{1cm}\begin{array}{ccc} & \uparrow & \\
\uparrow & \uparrow & \uparrow\\
& \uparrow & \end{array}
\hspace{1cm}\Longrightarrow\hspace{1cm}
E=4J\hspace{1cm}\begin{array}{ccc} & \uparrow & \\
\uparrow & \downarrow & \uparrow\\
& \uparrow & \end{array}
$$
The four other possibilities are as follows
$$
E=-2J\hspace{1cm}\begin{array}{ccc} & \uparrow & \\
\downarrow & \uparrow & \uparrow\\
& \uparrow & \end{array}
\hspace{1cm}\Longrightarrow\hspace{1cm}
E=2J\hspace{1cm}\begin{array}{ccc} & \uparrow & \\
\downarrow & \downarrow & \uparrow\\
& \uparrow & \end{array}
$$
with $\Delta E=4J$,
$$
E=0\hspace{1cm}\begin{array}{ccc} & \uparrow & \\
\downarrow & \uparrow & \uparrow\\
& \downarrow & \end{array}
\hspace{1cm}\Longrightarrow\hspace{1cm}
E=0\hspace{1cm}\begin{array}{ccc} & \uparrow & \\
\downarrow & \downarrow & \uparrow\\
& \downarrow & \end{array}
$$
with $\Delta E=0$,
$$
E=2J\hspace{1cm}\begin{array}{ccc} & \downarrow & \\
\downarrow & \uparrow & \uparrow\\
& \downarrow & \end{array}
\hspace{1cm}\Longrightarrow\hspace{1cm}
E=-2J\hspace{1cm}\begin{array}{ccc} & \downarrow & \\
\downarrow & \downarrow & \uparrow\\
& \downarrow & \end{array}
$$
with $\Delta E=-4J$ and finally
$$
E=4J\hspace{1cm}\begin{array}{ccc} & \downarrow & \\
\downarrow & \uparrow & \downarrow\\
& \downarrow & \end{array}
\hspace{1cm}\Longrightarrow\hspace{1cm}
E=-4J\hspace{1cm}\begin{array}{ccc} & \downarrow & \\
\downarrow & \downarrow & \downarrow\\
& \downarrow & \end{array}
$$
with $\Delta E=-8J$.
## The Metropolis Algorithm and the Two-dimensional Ising Model
This means in turn that we could construct an array which contains all values
of $e^{\beta \Delta E}$ before doing the Metropolis sampling. Else, we
would have to evaluate the exponential at each Monte Carlo sampling.
For the two-dimensional Ising model there are only five possible values. It is rather easy
to convice oneself that for the one-dimensional Ising model we have only three possible values.
The main part of the Ising model program is shown here
/*
Program to solve the two-dimensional Ising model
The coupling constant J = 1
Boltzmann's constant = 1, temperature has thus dimension energy
Metropolis sampling is used. Periodic boundary conditions.
*/
#include <iostream>
#include <fstream>
#include <iomanip>
#include "lib.h"
using namespace std;
ofstream ofile;
// inline function for periodic boundary conditions
inline int periodic(int i, int limit, int add) {
return (i+limit+add) % (limit);
}
// Function to read in data from screen
void read_input(int&, int&, double&, double&, double&);
// Function to initialise energy and magnetization
void initialize(int, double, int **, double&, double&);
// The metropolis algorithm
void Metropolis(int, long&, int **, double&, double&, double *);
// prints to file the results of the calculations
void output(int, int, double, double *);
// main program
int main(int argc, char* argv[])
{
char *outfilename;
long idum;
int **spin_matrix, n_spins, mcs;
double w[17], average[5], initial_temp, final_temp, E, M, temp_step;
// Read in output file, abort if there are too few command-line arguments
if( argc <= 1 ){
cout << "Bad Usage: " << argv[0] <<
" read also output file on same line" << endl;
exit(1);
}
else{
outfilename=argv[1];
}
ofile.open(outfilename);
// Read in initial values such as size of lattice, temp and cycles
read_input(n_spins, mcs, initial_temp, final_temp, temp_step);
spin_matrix = (int**) matrix(n_spins, n_spins, sizeof(int));
idum = -1; // random starting point
for ( double temp = initial_temp; temp <= final_temp; temp+=temp_step){
// initialise energy and magnetization
E = M = 0.;
// setup array for possible energy changes
for( int de =-8; de <= 8; de++) w[de+8] = 0;
for( int de =-8; de <= 8; de+=4) w[de+8] = exp(-de/temp);
// initialise array for expectation values
for( int i = 0; i < 5; i++) average[i] = 0.;
initialize(n_spins, double temp, spin_matrix, E, M);
// start Monte Carlo computation
for (int cycles = 1; cycles <= mcs; cycles++){
Metropolis(n_spins, idum, spin_matrix, E, M, w);
// update expectation values
average[0] += E; average[1] += E*E;
average[2] += M; average[3] += M*M; average[4] += fabs(M);
}
// print results
output(n_spins, mcs, temp, average);
}
free_matrix((void **) spin_matrix); // free memory
ofile.close(); // close output file
return 0;
}
## The Metropolis Algorithm and the Two-dimensional Ising Model
The array $w[17]$ contains values of $\Delta E$ spanning from $-8J$ to
$8J$ and it is precalculated in the main part for every new
temperature. The program takes as input the initial temperature, final
temperature, a temperature step, the number of spins in one direction
(we force the lattice to be a square lattice, meaning that we have the
same number of spins in the $x$ and the $y$ directions) and the number
of Monte Carlo cycles.
## The Metropolis Algorithm and the Two-dimensional Ising Model
For every Monte Carlo cycle we run through all
spins in the lattice in the function metropolis and flip one spin at
the time and perform the Metropolis test. However, every time we flip
a spin we need to compute the actual energy difference $\Delta E$ in
order to access the right element of the array which stores $e^{\beta
\Delta E}$. This is easily done in the Ising model since we can
exploit the fact that only one spin is flipped, meaning in turn that
all the remaining spins keep their values fixed. The energy
difference between a state $E_1$ and a state $E_2$ with zero external
magnetic field is
$$
\Delta E = E_2-E_1 =J\sum_{<kl>}^{N}s_k^1s_{l}^1-J\sum_{<kl>}^{N}s_k^2s_{l}^2,
$$
which we can rewrite as
$$
\Delta E = -J \sum_{<kl>}^{N}s_k^2(s_l^2-s_{l}^1),
$$
where the sum now runs only over the nearest neighbors $k$.
## The Metropolis Algorithm and the Two-dimensional Ising Model
Since the spin to be flipped takes only two values, $s_l^1=\pm 1$ and $s_l^2=\pm 1$, it means that if
$s_l^1= 1$, then $s_l^2=-1$ and if $s_l^1= -1$, then $s_l^2=1$.
The other spins keep their values, meaning that
$s_k^1=s_k^2$.
If $s_l^1= 1$ we must have $s_l^1-s_{l}^2=2$, and
if $s_l^1= -1$ we must have $s_l^1-s_{l}^2=-2$. From these results we see that the energy difference
can be coded efficiently as
<!-- Equation labels as ordinary links -->
<div id="eq:deltaeising"></div>
$$
\begin{equation}
\Delta E = 2Js_l^1\sum_{<k>}^{N}s_k,
\label{eq:deltaeising} \tag{3}
\end{equation}
$$
where the sum runs only over the nearest neighbors $k$ of spin $l$.
We can compute the change in magnetisation by flipping one spin as well.
Since only spin $l$ is flipped, all the surrounding spins remain unchanged.
## The Metropolis Algorithm and the Two-dimensional Ising Model
The difference in magnetisation is therefore only given by the difference
$s_l^1-s_{l}^2=\pm 2$, or in a more compact way as
<!-- Equation labels as ordinary links -->
<div id="eq:deltamising"></div>
$$
\begin{equation}
M_2 = M_1+2s_l^2,
\label{eq:deltamising} \tag{4}
\end{equation}
$$
where $M_1$ and $M_2$ are the magnetizations before and after the spin flip, respectively.
Eqs. [(3)](#eq:deltaeising) and [(4)](#eq:deltamising) are implemented in the function **metropolis** shown here
void Metropolis(int n_spins, long& idum, int **spin_matrix, double& E, double&M, double *w)
{
// loop over all spins
for(int y =0; y < n_spins; y++) {
for (int x= 0; x < n_spins; x++){
// Find random position
int ix = (int) (ran1(&idum)*(double)n_spins);
int iy = (int) (ran1(&idum)*(double)n_spins);
int deltaE = 2*spin_matrix[iy][ix]*
(spin_matrix[iy][periodic(ix,n_spins,-1)]+
spin_matrix[periodic(iy,n_spins,-1)][ix] +
spin_matrix[iy][periodic(ix,n_spins,1)] +
spin_matrix[periodic(iy,n_spins,1)][ix]);
// Here we perform the Metropolis test
if ( ran1(&idum) <= w[deltaE+8] ) {
spin_matrix[iy][ix] *= -1; // flip one spin and accept new spin config
// update energy and magnetization
M += (double) 2*spin_matrix[iy][ix];
E += (double) deltaE;
}
}
}
} // end of Metropolis sampling over spins
## The Metropolis Algorithm and the Two-dimensional Ising Model
Note that we loop over all spins but that we choose the lattice positions $x$ and $y$ randomly.
If the move is accepted after performing the Metropolis test, we update the energy and the magnetisation.
The new values are used to update the averages computed in the main function.
## The Metropolis Algorithm and the Two-dimensional Ising Model
We need also to initialize various variables.
This is done in the function here.
// function to initialise energy, spin matrix and magnetization
void initialize(int n_spins, double temp, int **spin_matrix,
double& E, double& M)
{
// setup spin matrix and intial magnetization
for(int y =0; y < n_spins; y++) {
for (int x= 0; x < n_spins; x++){
if (temp < 1.5) spin_matrix[y][x] = 1; // spin orientation for the ground state
M += (double) spin_matrix[y][x];
}
}
// setup initial energy
for(int y =0; y < n_spins; y++) {
for (int x= 0; x < n_spins; x++){
E -= (double) spin_matrix[y][x]*
(spin_matrix[periodic(y,n_spins,-1)][x] +
spin_matrix[y][periodic(x,n_spins,-1)]);
}
}
}// end function initialise
## Two-dimensional Ising Model and analysis of spin values
The following python code displays the values of the spins as function of temperature.
```
# coding=utf-8
#2-dimensional ising model with visualization
import numpy, sys, math
import pygame
#Needed for visualize when using SDL
screen = None;
font = None;
BLOCKSIZE = 10
def periodic (i, limit, add):
"""
Choose correct matrix index with periodic
boundary conditions
Input:
- i: Base index
- limit: Highest \"legal\" index
- add: Number to add or subtract from i
"""
return (i+limit+add) % limit
def visualize(spin_matrix, temp, E, M, method):
"""
Visualize the spin matrix
Methods:
method = -1:No visualization (testing)
method = 0: Just print it to the terminal
method = 1: Pretty-print to terminal
method = 2: SDL/pygame single-pixel
method = 3: SDL/pygame rectangle
"""
#Simple terminal dump
if method == 0:
print "temp:", temp, "E:", E, "M:", M
print spin_matrix
#Pretty-print to terminal
elif method == 1:
out = ""
size = len(spin_matrix)
for y in xrange(size):
for x in xrange(size):
if spin_matrix.item(x,y) == 1:
out += "X"
else:
out += " "
out += "\n"
print "temp:", temp, "E:", E, "M:", M
print out + "\n"
#SDL single-pixel (useful for large arrays)
elif method == 2:
size = len(spin_matrix)
screen.lock()
for y in xrange(size):
for x in xrange(size):
if spin_matrix.item(x,y) == 1:
screen.set_at((x,y),(0,0,255))
else:
screen.set_at((x,y),(255,0,0))
screen.unlock()
pygame.display.flip()
#SDL block (usefull for smaller arrays)
elif method == 3:
size = len(spin_matrix)
screen.lock()
for y in xrange(size):
for x in xrange(size):
if spin_matrix.item(x,y) == 1:
rect = pygame.Rect(x*BLOCKSIZE,y*BLOCKSIZE,BLOCKSIZE,BLOCKSIZE)
pygame.draw.rect(screen,(0,0,255),rect)
else:
rect = pygame.Rect(x*BLOCKSIZE,y*BLOCKSIZE,BLOCKSIZE,BLOCKSIZE)
pygame.draw.rect(screen,(255,0,0),rect)
screen.unlock()
pygame.display.flip()
#SDL block w/ data-display
elif method == 4:
size = len(spin_matrix)
screen.lock()
for y in xrange(size):
for x in xrange(size):
if spin_matrix.item(x,y) == 1:
rect = pygame.Rect(x*BLOCKSIZE,y*BLOCKSIZE,BLOCKSIZE,BLOCKSIZE)
pygame.draw.rect(screen,(255,255,255),rect)
else:
rect = pygame.Rect(x*BLOCKSIZE,y*BLOCKSIZE,BLOCKSIZE,BLOCKSIZE)
pygame.draw.rect(screen,(0,0,0),rect)
s = font.render("<E> = %5.3E; <M> = %5.3E" % E,M,False,(255,0,0))
screen.blit(s,(0,0))
screen.unlock()
pygame.display.flip()
def monteCarlo(temp, size, trials, visual_method):
"""
Calculate the energy and magnetization
(\"straight\" and squared) for a given temperature
Input:
- temp: Temperature to calculate for
- size: dimension of square matrix
- trials: Monte-carlo trials (how many times do we
flip the matrix?)
- visual_method: What method should we use to visualize?
Output:
- E_av: Energy of matrix averaged over trials, normalized to spins**2
- E_variance: Variance of energy, same normalization * temp**2
- M_av: Magnetic field of matrix, averaged over trials, normalized to spins**2
- M_variance: Variance of magnetic field, same normalization * temp
- Mabs: Absolute value of magnetic field, averaged over trials
"""
#Setup spin matrix, initialize to ground state
spin_matrix = numpy.zeros( (size,size), numpy.int8) + 1
#Create and initialize variables
E = M = 0
E_av = E2_av = M_av = M2_av = Mabs_av = 0
#Setup array for possible energy changes
w = numpy.zeros(17,numpy.float64)
for de in xrange(-8,9,4): #include +8
w[de+8] = math.exp(-de/temp)
#Calculate initial magnetization:
M = spin_matrix.sum()
#Calculate initial energy
for j in xrange(size):
for i in xrange(size):
E -= spin_matrix.item(i,j)*\
(spin_matrix.item(periodic(i,size,-1),j) + spin_matrix.item(i,periodic(j,size,1)))
#Start metropolis MonteCarlo computation
for i in xrange(trials):
#Metropolis
#Loop over all spins, pick a random spin each time
for s in xrange(size**2):
x = int(numpy.random.random()*size)
y = int(numpy.random.random()*size)
deltaE = 2*spin_matrix.item(x,y)*\
(spin_matrix.item(periodic(x,size,-1), y) +\
spin_matrix.item(periodic(x,size,1), y) +\
spin_matrix.item(x, periodic(y,size,-1)) +\
spin_matrix.item(x, periodic(y,size,1)))
if numpy.random.random() <= w[deltaE+8]:
#Accept!
spin_matrix[x,y] *= -1
M += 2*spin_matrix[x,y]
E += deltaE
#Update expectation values
E_av += E
E2_av += E**2
M_av += M
M2_av += M**2
Mabs_av += int(math.fabs(M))
visualize(spin_matrix, temp,E/float(size**2),M/float(size**2), method);
#Normalize average values
E_av /= float(trials);
E2_av /= float(trials);
M_av /= float(trials);
M2_av /= float(trials);
Mabs_av /= float(trials);
#Calculate variance and normalize to per-point and temp
E_variance = (E2_av-E_av*E_av)/float(size*size*temp*temp);
M_variance = (M2_av-M_av*M_av)/float(size*size*temp);
#Normalize returned averages to per-point
E_av /= float(size*size);
M_av /= float(size*size);
Mabs_av /= float(size*size);
return (E_av, E_variance, M_av, M_variance, Mabs_av)
# Main program
size = 100
trials = 100000
temp = 2.5
method = 3
#Initialize pygame
if method == 2 or method == 3 or method == 4:
pygame.init()
if method == 2:
screen = pygame.display.set_mode((size,size))
elif method == 3:
screen = pygame.display.set_mode((size*10,size*10))
elif method == 4:
screen = pygame.display.set_mode((size*10,size*10))
font = pygame.font.Font(None,12)
(E_av, E_variance, M_av, M_variance, Mabs_av) = monteCarlo(temp,size,trials, method)
print "%15.8E %15.8E %15.8E %15.8E %15.8E %15.8E\n" % (temp, E_av, E_variance, M_av, M_variance, Mabs_av)
pygame.quit();
```
| github_jupyter |
```
import open3d as o3d
import numpy as np
import sys
# monkey patches visualization and provides helpers to load geometries
sys.path.append('..')
import open3d_tutorial as o3dtut
# change to True if you want to interact with the visualization windows
o3dtut.interactive = False
```
# KDTree
Open3D uses [FLANN](https://www.cs.ubc.ca/research/flann/) to build KDTrees for fast retrieval of nearest neighbors.
## Build KDTree from point cloud
The code below reads a point cloud and builds a KDTree. This is a preprocessing step for the following nearest neighbor queries.
```
print("Testing kdtree in open3d ...")
print("Load a point cloud and paint it gray.")
pcd = o3d.io.read_point_cloud("../../TestData/Feature/cloud_bin_0.pcd")
pcd.paint_uniform_color([0.5, 0.5, 0.5])
pcd_tree = o3d.geometry.KDTreeFlann(pcd)
```
## Find neighboring points
We pick the 1500-th point as the anchor point and paint it red.
```
print("Paint the 1500th point red.")
pcd.colors[1500] = [1, 0, 0]
```
### Using search_knn_vector_3d
The function `search_knn_vector_3d` returns a list of indices of the k nearest neighbors of the anchor point. These neighboring points are painted with blue color. Note that we convert `pcd.colors` to a numpy array to make batch access to the point colors, and broadcast a blue color [0, 0, 1] to all the selected points. We skip the first index since it is the anchor point itself.
```
print("Find its 200 nearest neighbors, paint blue.")
[k, idx, _] = pcd_tree.search_knn_vector_3d(pcd.points[1500], 200)
np.asarray(pcd.colors)[idx[1:], :] = [0, 0, 1]
```
### Using search_radius_vector_3d
Similarly, we can use `search_radius_vector_3d` to query all points with distances to the anchor point less than a given radius. We paint these points with green color.
```
print("Find its neighbors with distance less than 0.2, paint green.")
[k, idx, _] = pcd_tree.search_radius_vector_3d(pcd.points[1500], 0.2)
np.asarray(pcd.colors)[idx[1:], :] = [0, 1, 0]
print("Visualize the point cloud.")
o3d.visualization.draw_geometries([pcd], zoom=0.5599,
front=[-0.4958, 0.8229, 0.2773],
lookat=[2.1126, 1.0163, -1.8543],
up=[0.1007, -0.2626, 0.9596])
```
<div class="alert alert-info">
**Note:**
Besides the KNN search `search_knn_vector_3d` and the RNN search `search_radius_vector_3d`, Open3D provides a hybrid search function `search_hybrid_vector_3d`. It returns at most k nearest neighbors that have distances to the anchor point less than a given radius. This function combines the criteria of KNN search and RNN search. It is known as RKNN search in some literatures. It has performance benefits in many practical cases, and is heavily used in a number of Open3D functions.
</div>
| github_jupyter |
```
from scipy.stats import norm
import numpy as np
from math import cos, sin
import matplotlib.pyplot as plt
%matplotlib inline
import seaborn
from copy import copy
from sklearn.decomposition import PCA
```
# 7.1 The Curse of Dimensionality
Consider a D-dimensional uniform distribution with coordinates between [r,-r] on each axis.
What is the probability of randomly drawing a point that lies within a distance $ r=\sqrt{x_1^2+\dots+x_{D}^2} $ of the origin?
The volume of a hypersphere with radius r is $$ V_D(r)=\frac{2 r^D \pi^{D/2}}{D~\Gamma(D/2)} $$ so this probability is given by $$ f_D=\frac{V_D(r)}{(2r)^D} = \frac{\pi^{D/2}}{D~2^{D-1}~\Gamma(D/2)} $$ The limit of which goes to zero as D approaches infinity: $$ \lim_ {D\to \infty} f_d = 0 $$
The larger the dimensionality of your data set, the more difficult it is to evenly sample its volume.
# 7.3 Principal Component Analysis
### Simple explanation:
#### PCA is a method for reducing the dimensionality of a data set while preserving as much of the valuable information in that set as possible.
#### Some reasons why PCA is useful:
- By reducing the dimensionality of our data, we can more easily visualize it (e.g., 3D->2D transformation)
- It reduces the amount of space needed to store the data
- It can speed up computation time in working with the data (less things to calculate)
- It can give us useful insights into the way our data's features relate to one another
#### The PCA algorithm essentially performs the following steps:
1. Given a set of data $\vec{x}$ in $k$-dimensions aligned with axes $\hat{e}_{i}$, find a new set of $k$ axes $\hat{e}_{i}'$ such that the covariances of the data set with respect to the new axes is zero: $\sigma_{\hat{e}_{i}'}\sigma_{\hat{e}_{j}'} = 0 \text{ for all } i\neq j$.
2. Drop the axes with the least variances
#### A very informative link:
http://setosa.io/ev/principal-component-analysis/
```
# Author: Jake VanderPlas
# License: BSD
# The figure produced by this code is published in the textbook
# "Statistics, Data Mining, and Machine Learning in Astronomy" (2013)
# For more information, see http://astroML.github.com
# To report a bug or issue, use the following forum:
# https://groups.google.com/forum/#!forum/astroml-general
import numpy as np
from matplotlib import pyplot as plt
from matplotlib.patches import Ellipse
#----------------------------------------------------------------------
# This function adjusts matplotlib settings for a uniform feel in the textbook.
# Note that with usetex=True, fonts are rendered with LaTeX. This may
# result in an error if LaTeX is not installed on your system. In that case,
# you can set usetex to False.
from astroML.plotting import setup_text_plots
setup_text_plots(fontsize=8, usetex=True)
#------------------------------------------------------------
# Set parameters and draw the random sample
np.random.seed(42)
r = 0.9
sigma1 = 0.25
sigma2 = 0.08
rotation = np.pi / 6
s = np.sin(rotation)
c = np.cos(rotation)
X = np.random.normal(0, [sigma1, sigma2], size=(100, 2)).T
R = np.array([[c, -s],
[s, c]])
X = np.dot(R, X)
#------------------------------------------------------------
# Plot the diagram
fig = plt.figure(figsize=(10, 10), facecolor='w')
ax = plt.axes((0, 0, 1, 1), xticks=[], yticks=[], frameon=False)
# draw axes
ax.annotate(r'$x$', (-r, 0), (r, 0),
ha='center', va='center',
arrowprops=dict(arrowstyle='<->', color='k', lw=1))
ax.annotate(r'$y$', (0, -r), (0, r),
ha='center', va='center',
arrowprops=dict(arrowstyle='<->', color='k', lw=1))
# draw rotated axes
ax.annotate(r'$x^\prime$', (-r * c, -r * s), (r * c, r * s),
ha='center', va='center',
arrowprops=dict(color='k', arrowstyle='<->', lw=1))
ax.annotate(r'$y^\prime$', (r * s, -r * c), (-r * s, r * c),
ha='center', va='center',
arrowprops=dict(color='k', arrowstyle='<->', lw=1))
# scatter points
ax.scatter(X[0], X[1], s=25, lw=0, c='k', zorder=2)
# draw lines
vnorm = np.array([s, -c])
for v in (X.T):
d = np.dot(v, vnorm)
v1 = v - d * vnorm
ax.plot([v[0], v1[0]], [v[1], v1[1]], '-k')
ax.scatter(v1[0], v1[1], color = (1,0,0), s = 50)
# draw ellipses
for sigma in (1, 2, 3):
ax.add_patch(Ellipse((0, 0), 2 * sigma * sigma1, 2 * sigma * sigma2,
rotation * 180. / np.pi,
ec='k', fc='gray', alpha=0.2, zorder=1))
ax.set_xlim(-1, 1)
ax.set_ylim(-1, 1)
plt.show()
```
## 7.3.1 Derivation of Principal Component Analysis (page 293)
- Start with a set of data $\boldsymbol{X}$ comprising a series of $N$ observations (e.g., stars), each observation made up of $K$ features (e.g. magnitude, color, etc.). $X^{\left(j\right)}_{i}$ is feature $j$ of the $i^{\text{th}}$ observation.
- Subtract the mean of each feature from every data point: $X^{\left(j\right)}_{i}\rightarrow X^{\left(j\right)}_{i}-\bar{X}^{\left(j\right)}$
- Optionally, one can scale all features by dividing data by the feature standard deviation
- Calculate the variance of the data along some arbitrary axis $\hat{r}_{1}$:
$$\begin{align}
\sigma_{\hat{r}_{1}}^{2} &= \sum_{i}^{N}\left(\vec{X_{i}}\cdot\hat{r}_{1}\right)^{2}\\
&=\sum_{i}^{N}\left(\hat{r}_{1}^{T}\cdot\vec{X_{i}}^{T}\right)\left(\vec{X_{i}}\cdot\hat{r}_{1}\right)\\
\end{align}
$$
- Choose $\hat{r}_{1}$ to be the axis of maximum variance. To find $\hat{r}_{1}$, maximize the variance subject to the constraint $\hat{r}_{1}^{T}\hat{r}_{1} = 1$ (the axis is a unit vector); this can be done by using Lagrange multipliers:
$$\phi\left(\hat{r}_{1},\lambda_{1}\right)=\
\sum_{i}^{N}\left(\hat{r}_{1}^{T}\cdot\vec{X_{i}}^{T}\right)\left(\vec{X_{i}}\cdot\hat{r}_{1}\right)\
-\lambda_{1}\left(\hat{r}_{1}^{T}\hat{r}_{1}-1\right)$$
- Setting $\frac{\text{d}\phi}{\text{d}\hat{r}_{1}} = 0$ yields
$$\left(\sum_{i}^{N}\vec{X}_{i}^{T}\vec{X}_{i}\right)\hat{r}_{1}-\lambda_{1}\hat{r}_{1}=0$$
- This is an eigenvalue equation for the maximum variance $\lambda_{1}$ and the corresponding axis of maximal variance $\hat{r}_{1}$.
- Notice that $\sum_{i}^{N}\vec{X}_{i}^{T}\vec{X}_{i} = \boldsymbol{X}^{T}\boldsymbol{X}=\boldsymbol{C}_{\boldsymbol{X}}$--the axis of maximum variance is an eigenvector of the covariance matrix
- To get the next axis $\hat{r}_{2}$, we solve the same Lagrange multiplier equation with the additional constraint that the covariance $\sigma_{\hat{r}_{1}\hat{r}_{2}} = 0$, and repeat
- Finally, we transform the data by taking $\boldsymbol{Y}=\boldsymbol{X}\boldsymbol{R}$ and dropping low-variance features (the right-most rows in the transformed data
### Singular value decomposition (SVD)
- The PCA can be computed through the data matrix $\boldsymbol{X}$ itself rather than through the eigenvalue decomposition of the covariance matrix; this method is known as **singular value decomposition**
- SVD:
$$\boldsymbol{U\Sigma V}^{T}=\frac{1}{\sqrt{N-1}}\boldsymbol{X}$$
- $\boldsymbol{U}$: *left-singular matrix*
- $\boldsymbol{\Sigma}$: matrix of singular values
- $\boldsymbol{V}$: *right-singular matrix*
$$\boldsymbol{C}_{\boldsymbol{X}}=\left[\frac{1}{\sqrt{N-1}}\right]$$
- Can show that
- $\boldsymbol{V}=\boldsymbol{R}$
- $\boldsymbol{\Sigma}^{2}=\boldsymbol{C}_{\boldsymbol{Y}}$
- SVD of the data directly yields the projection $\boldsymbol{R}$ and variances $\boldsymbol{C}_{\boldsymbol{Y}}$
### Three ways to calculate the principal components $\boldsymbol{R}$ and the eigenvalues $\boldsymbol{C}_{\boldsymbol{Y}}$:
1. Eigenvalue decomposition of covariance matrix $\boldsymbol{C}_{\boldsymbol{X}}$
- Use if $N\gg K$
2. SVD of $\boldsymbol{X}$
- Use if $N\sim K$
3. Eigenvalue decomposition of correlation matrix $\boldsymbol{M}_{\boldsymbol{X}}$
- Use if $K\gg N$
## 7.3.2 The Application of PCA
```
#We have ~100 carbon enhanced metal poor stars with 23 chemical abundances measured [Fe/H] and [X/Fe]
#importing and floatify data
infile=open('CEMPabundances.dat','r')
per_row = []
for line in infile:
per_row.append(line.split('\t'))
elem_abundance_arr_str = zip(*per_row)
truncated_elem_abundance_arr=[]
tot_elem_abundance_arr=[]
for arr0 in elem_abundance_arr_str:
tot_elem_abundance_arr.append([float(i) for i in arr0])
#remove unmeasured elements
for n in range(len(tot_elem_abundance_arr)):
arr0=tot_elem_abundance_arr[n]
truncated_elem_abundance_arr.append([x for x in arr0 if x!=-10000.])
#read labels for abundances
infile.close()
#for arr in elem_abundance_arr:
# for i in range(len(arr)):
# arr[i]=float(arr[i])
infile=open('elem_list.dat','r')
per_row = []
for line in infile:
per_row.append(line.split('\t'))
elems=per_row[0]
infile.close()
#histogram distributions in abundances
fig = plt.figure(figsize=(20, 20))
for n in range(0,23):
ax=fig.add_subplot(5,5,(n+1))
ax.hist(truncated_elem_abundance_arr[n], 10, normed=1, facecolor='green', alpha=0.75)
#ax.set_xlabel(elems[n])
ax.annotate(elems[n] ,xy=(0.05,0.9), xycoords='axes fraction', size=14)
plt.show()
```
# searching for structure in a 23-dimensional data set
Without reducing the dimensionality, we can play around and try to search for some structure in the abundances. Looking for correlations between two dimensions is doable. If we were looking for structure in a plane defined by just two of these axes, we could make a triangle plot (no room here though) and visually inspect for any pairs of related abundances. We could do something similar to search for structure in the data that depended on 3 dimensions, but that would be hard to visualize and we'd have $\mathcal{O}(D^3)$ combinations to test.
```
Dim1=10 #0-22 cheat: try combinations of 1,17,18,20,21,22
Dim2=17 #0-22
Nobjs=len(tot_elem_abundance_arr[0])
#reform this array since next cell is changing it for whatever reason
tot_elem_abundance_arr=[]
for arr0 in elem_abundance_arr_str:
tot_elem_abundance_arr.append([float(i) for i in arr0])
#only use objects with measurements for both elements
arr1,arr2=[],[]
for obj in range(Nobjs):
if tot_elem_abundance_arr[Dim1][obj]!=-10000. and tot_elem_abundance_arr[Dim2][obj]!=-10000.:
arr1.append(tot_elem_abundance_arr[Dim1][obj])
arr2.append(tot_elem_abundance_arr[Dim2][obj])
fig=plt.figure(figsize=(5, 5))
ax=fig.add_subplot(111)
ax.plot(arr1,arr2, 'o')
ax.annotate('N$_{obj}$='+repr(len(arr1)) ,xy=(0.05,0.9), xycoords='axes fraction', size=14)
ax.set_xlabel(elems[Dim1],fontsize=22)
ax.set_ylabel(elems[Dim2],fontsize=22)
```
# use PCA to find vectors in 23-space that account for most variance
```
#unmeasured elements replaced with median of measurements
processed_arr=[]
copy_arr=list(copy(tot_elem_abundance_arr))
for n in range(len(copy_arr)):
arr=copy_arr[n]
processed_arr.append(arr)
temp_arr=[]
for i in range(Nobjs):
if arr[i]!=-10000:
temp_arr.append(arr[i])
for i in range(Nobjs):
if arr[i]==-10000:
processed_arr[n][i]=np.median(temp_arr)
processed_arr=np.transpose(processed_arr)
pca = PCA(n_components=3)
pca.fit(processed_arr)
print 'Percent of variance in each PC'
print(pca.explained_variance_ratio_)
pca_score = pca.explained_variance_ratio_
V = pca.components_
PC1, PC2, PC3=V[0],V[1],V[2]
# plot orientations of principal components
fig=plt.figure(figsize=(20, 12))
ax=fig.add_subplot(311)
x=np.linspace(0,22,23)
zeroline=np.zeros(23)
ax.set_xlim([-1,24])
ax.set_ylim([-0.3,1.2])
ax.set_ylabel('PC 1',fontsize=20)
ax.plot(x,zeroline,'--',color='grey')
for n in range(len(elems)):
ax.annotate(elems[n] ,xy=(n,1.0+((-1)**n)*0.05), xycoords='data',ha="center", size=14)
ax.plot(x,PC1,'o')
ax2=fig.add_subplot(312)
ax2.set_xlim([-1,24])
ax2.set_ylim([-0.6,1.0])
ax2.set_ylabel('PC 2',fontsize=20)
ax2.plot(x,zeroline,'--',color='grey')
for n in range(len(elems)):
ax2.annotate(elems[n] ,xy=(n,0.7+((-1)**n)*0.05), xycoords='data',ha="center", size=14)
ax2.plot(x,PC2,'o')
ax3=fig.add_subplot(313)
ax3.set_xlim([-1,24])
ax3.set_ylim([-0.6,1.0])
ax3.set_ylabel('PC 3',fontsize=20)
ax3.plot(x,zeroline,'--',color='grey')
for n in range(len(elems)):
ax3.annotate(elems[n] ,xy=(n,0.7+((-1)**n)*0.05), xycoords='data',ha="center", size=14)
ax3.plot(x,PC3,'o')
# Author: Jake VanderPlas
# License: BSD
# The figure produced by this code is published in the textbook
# "Statistics, Data Mining, and Machine Learning in Astronomy" (2013)
# For more information, see http://astroML.github.com
# To report a bug or issue, use the following forum:
# https://groups.google.com/forum/#!forum/astroml-general
import numpy as np
from matplotlib import pyplot as plt
from astroML import datasets
#----------------------------------------------------------------------
# This function adjusts matplotlib settings for a uniform feel in the textbook.
# Note that with usetex=True, fonts are rendered with LaTeX. This may
# result in an error if LaTeX is not installed on your system. In that case,
# you can set usetex to False.
from astroML.plotting import setup_text_plots
setup_text_plots(fontsize=8, usetex=True)
#------------------------------------------------------------
# load data:
data = datasets.fetch_sdss_corrected_spectra()
spectra = datasets.sdss_corrected_spectra.reconstruct_spectra(data)
# Eigenvalues can be computed using PCA as in the commented code below:
#from sklearn.decomposition import PCA
#pca = PCA()
#pca.fit(spectra)
#evals = pca.explained_variance_ratio_
#evals_cs = evals.cumsum()
# because the spectra have been reconstructed from masked values, this
# is not exactly correct in this case: we'll use the values computed
# in the file compute_sdss_pca.py
evals = data['evals'] ** 2
evals_cs = evals.cumsum()
evals_cs /= evals_cs[-1]
#------------------------------------------------------------
# plot the eigenvalues
fig = plt.figure(figsize=(5, 3.75))
fig.subplots_adjust(hspace=0.05, bottom=0.12)
ax = fig.add_subplot(211, xscale='log', yscale='log')
ax.grid()
ax.plot(evals, c='k')
ax.set_ylabel('Normalized Eigenvalues')
ax.xaxis.set_major_formatter(plt.NullFormatter())
ax.set_ylim(5E-4, 100)
ax = fig.add_subplot(212, xscale='log')
ax.grid()
ax.semilogx(evals_cs, color='k')
ax.set_xlabel('Eigenvalue Number')
ax.set_ylabel('Cumulative Eigenvalues')
ax.set_ylim(0.65, 1.00)
plt.show()
```

```
# Author: Jake VanderPlas
# License: BSD
# The figure produced by this code is published in the textbook
# "Statistics, Data Mining, and Machine Learning in Astronomy" (2013)
# For more information, see http://astroML.github.com
# To report a bug or issue, use the following forum:
# https://groups.google.com/forum/#!forum/astroml-general
import numpy as np
from matplotlib import pyplot as plt
from sklearn.decomposition import PCA
from astroML.datasets import sdss_corrected_spectra
from astroML.decorators import pickle_results
#----------------------------------------------------------------------
# This function adjusts matplotlib settings for a uniform feel in the textbook.
# Note that with usetex=True, fonts are rendered with LaTeX. This may
# result in an error if LaTeX is not installed on your system. In that case,
# you can set usetex to False.
from astroML.plotting import setup_text_plots
setup_text_plots(fontsize=8, usetex=True)
#------------------------------------------------------------
# Download data
data = sdss_corrected_spectra.fetch_sdss_corrected_spectra()
spectra = sdss_corrected_spectra.reconstruct_spectra(data)
wavelengths = sdss_corrected_spectra.compute_wavelengths(data)
#------------------------------------------------------------
# Compute PCA components
# Eigenvalues can be computed using PCA as in the commented code below:
#from sklearn.decomposition import PCA
#pca = PCA()
#pca.fit(spectra)
#evals = pca.explained_variance_ratio_
#evals_cs = evals.cumsum()
# because the spectra have been reconstructed from masked values, this
# is not exactly correct in this case: we'll use the values computed
# in the file compute_sdss_pca.py
evals = data['evals'] ** 2
evals_cs = evals.cumsum()
evals_cs /= evals_cs[-1]
evecs = data['evecs']
spec_mean = spectra.mean(0)
#------------------------------------------------------------
# Find the coefficients of a particular spectrum
spec = spectra[1]
coeff = np.dot(evecs, spec - spec_mean)
#------------------------------------------------------------
# Plot the sequence of reconstructions
fig = plt.figure(figsize=(5, 5))
fig.subplots_adjust(hspace=0, top=0.95, bottom=0.1, left=0.12, right=0.93)
for i, n in enumerate([0, 4, 8, 20]):
ax = fig.add_subplot(411 + i)
ax.plot(wavelengths, spec, '-', c='gray')
ax.plot(wavelengths, spec_mean + np.dot(coeff[:n], evecs[:n]), '-k')
if i < 3:
ax.xaxis.set_major_formatter(plt.NullFormatter())
ax.set_ylim(-2, 21)
ax.set_ylabel('flux')
if n == 0:
text = "mean"
elif n == 1:
text = "mean + 1 component\n"
text += r"$(\sigma^2_{tot} = %.2f)$" % evals_cs[n - 1]
else:
text = "mean + %i components\n" % n
text += r"$(\sigma^2_{tot} = %.2f)$" % evals_cs[n - 1]
ax.text(0.02, 0.93, text, ha='left', va='top', transform=ax.transAxes)
fig.axes[-1].set_xlabel(r'${\rm wavelength\ (\AA)}$')
plt.show()
```

### Reconstruction of data from the eigenbasis:
$$\vec{x}_{i}\left(k\right)=\vec{\mu}\left(k\right)+\sum_{j}^{R}\theta_{ij}\vec{e}_{j}\left(k\right)$$
$$\theta_{ij}=\sum_{k}\vec{e}_{j}\left(k\right)\left(\vec{x}_{i}\left(k\right)-\vec{\mu}\left(k\right)\right)$$
- Truncate the expansion to compress the data:
$\vec{x}_{i}\left(k\right)=\sum_{i}^{r<R}\theta_{i}\vec{e}_{i}\left(k\right)$
- How many terms to include in the expansion? One way is to define a bound $\alpha$ on the fraction of the variance we want to capture:
$$\frac{\sum_{i}^{i=r}\sigma_{i}}{\sum_{i}^{i=R}\sigma_{i}}<\alpha$$
- $\alpha$ typically taken from 0.70 to 0.95;
- Manually inspect the cumulative-eigenvalues plot, truncate at the 'knee'
## PCA with missing data
```
# Author: Jake VanderPlas
# License: BSD
# The figure produced by this code is published in the textbook
# "Statistics, Data Mining, and Machine Learning in Astronomy" (2013)
# For more information, see http://astroML.github.com
# To report a bug or issue, use the following forum:
# https://groups.google.com/forum/#!forum/astroml-general
import numpy as np
from matplotlib import pyplot as plt
from matplotlib import ticker
from astroML.datasets import fetch_sdss_corrected_spectra
from astroML.datasets import sdss_corrected_spectra
#----------------------------------------------------------------------
# This function adjusts matplotlib settings for a uniform feel in the textbook.
# Note that with usetex=True, fonts are rendered with LaTeX. This may
# result in an error if LaTeX is not installed on your system. In that case,
# you can set usetex to False.
from astroML.plotting import setup_text_plots
setup_text_plots(fontsize=8, usetex=True)
#------------------------------------------------------------
# Get spectra and eigenvectors used to reconstruct them
data = fetch_sdss_corrected_spectra()
spec = sdss_corrected_spectra.reconstruct_spectra(data)
lam = sdss_corrected_spectra.compute_wavelengths(data)
evecs = data['evecs']
mu = data['mu']
norms = data['norms']
mask = data['mask']
#------------------------------------------------------------
# plot the results
i_plot = ((lam > 5750) & (lam < 6350))
lam = lam[i_plot]
specnums = [20, 8, 9]
subplots = [311, 312, 313]
fig = plt.figure(figsize=(5, 6.25))
fig.subplots_adjust(left=0.09, bottom=0.08, hspace=0, right=0.92, top=0.95)
for subplot, i in zip(subplots, specnums):
ax = fig.add_subplot(subplot)
# compute eigen-coefficients
spec_i_centered = spec[i] / norms[i] - mu
coeffs = np.dot(spec_i_centered, evecs.T)
# blank out masked regions
spec_i = spec[i]
mask_i = mask[i]
spec_i[mask_i] = np.nan
# plot the raw masked spectrum
ax.plot(lam, spec_i[i_plot], '-', color='k',
label='True spectrum', lw=1.5)
# plot two levels of reconstruction
for nev in [10]:
if nev == 0:
label = 'mean'
else:
label = 'reconstruction\n(nev=%i)' % nev
spec_i_recons = norms[i] * (mu + np.dot(coeffs[:nev], evecs[:nev]))
ax.plot(lam, spec_i_recons[i_plot], label=label, color='grey')
# plot shaded background in masked region
ylim = ax.get_ylim()
mask_shade = ylim[0] + mask[i][i_plot].astype(float) * ylim[1]
plt.fill(np.concatenate([lam[:1], lam, lam[-1:]]),
np.concatenate([[ylim[0]], mask_shade, [ylim[0]]]),
lw=0, fc='k', alpha=0.2)
ax.set_xlim(lam[0], lam[-1])
ax.set_ylim(ylim)
ax.yaxis.set_major_formatter(ticker.NullFormatter())
if subplot == 311:
ax.legend(loc=1)
ax.set_xlabel('$\lambda\ (\AA)$')
ax.set_ylabel('normalized flux')
plt.show()
```

| github_jupyter |
<h1>Table of Contents<span class="tocSkip"></span></h1>
<div class="toc" style="margin-top: 1em;"><ul class="toc-item"><li><span><a href="#Name" data-toc-modified-id="Name-1"><span class="toc-item-num">1 </span>Name</a></span></li><li><span><a href="#Search" data-toc-modified-id="Search-2"><span class="toc-item-num">2 </span>Search</a></span><ul class="toc-item"><li><span><a href="#Load-Cached-Results" data-toc-modified-id="Load-Cached-Results-2.1"><span class="toc-item-num">2.1 </span>Load Cached Results</a></span></li><li><span><a href="#Run-From-Scratch" data-toc-modified-id="Run-From-Scratch-2.2"><span class="toc-item-num">2.2 </span>Run From Scratch</a></span></li></ul></li><li><span><a href="#Analysis" data-toc-modified-id="Analysis-3"><span class="toc-item-num">3 </span>Analysis</a></span><ul class="toc-item"><li><span><a href="#Gender-Breakdown" data-toc-modified-id="Gender-Breakdown-3.1"><span class="toc-item-num">3.1 </span>Gender Breakdown</a></span></li><li><span><a href="#Face-Sizes" data-toc-modified-id="Face-Sizes-3.2"><span class="toc-item-num">3.2 </span>Face Sizes</a></span></li><li><span><a href="#Appearances-on-a-Single-Show" data-toc-modified-id="Appearances-on-a-Single-Show-3.3"><span class="toc-item-num">3.3 </span>Appearances on a Single Show</a></span></li><li><span><a href="#Screen-Time-Across-All-Shows" data-toc-modified-id="Screen-Time-Across-All-Shows-3.4"><span class="toc-item-num">3.4 </span>Screen Time Across All Shows</a></span></li></ul></li><li><span><a href="#Persist-to-Cloud" data-toc-modified-id="Persist-to-Cloud-4"><span class="toc-item-num">4 </span>Persist to Cloud</a></span><ul class="toc-item"><li><span><a href="#Save-Model-to-GCS" data-toc-modified-id="Save-Model-to-GCS-4.1"><span class="toc-item-num">4.1 </span>Save Model to GCS</a></span><ul class="toc-item"><li><span><a href="#Make-sure-the-GCS-file-is-valid" data-toc-modified-id="Make-sure-the-GCS-file-is-valid-4.1.1"><span class="toc-item-num">4.1.1 </span>Make sure the GCS file is valid</a></span></li></ul></li><li><span><a href="#Save-Labels-to-DB" data-toc-modified-id="Save-Labels-to-DB-4.2"><span class="toc-item-num">4.2 </span>Save Labels to DB</a></span><ul class="toc-item"><li><span><a href="#Commit-the-person-and-labeler" data-toc-modified-id="Commit-the-person-and-labeler-4.2.1"><span class="toc-item-num">4.2.1 </span>Commit the person and labeler</a></span></li><li><span><a href="#Commit-the-FaceIdentity-labels" data-toc-modified-id="Commit-the-FaceIdentity-labels-4.2.2"><span class="toc-item-num">4.2.2 </span>Commit the FaceIdentity labels</a></span></li></ul></li></ul></li></ul></div>
```
from esper.prelude import *
from esper.identity import *
from esper import embed_google_images
```
# Name
```
name = 'Sean Hannity'
```
# Search
## Load Cached Results
```
assert name != ''
results = FaceIdentityModel.load(name=name)
imshow(np.hstack([cv2.resize(x[1][0], (200, 200)) for x in results.model_params['images']]))
plt.show()
plot_precision_and_cdf(results)
```
## Run From Scratch
Run this section if you do not have a cached model and precision curve estimates.
```
assert name != ''
img_dir = embed_google_images.fetch_images(name)
face_imgs = load_and_select_faces_from_images(img_dir)
face_embs = embed_google_images.embed_images(face_imgs)
assert(len(face_embs) == len(face_imgs))
imshow(np.hstack([cv2.resize(x[0], (200, 200)) for x in face_imgs if x]))
plt.show()
face_ids_by_bucket, face_ids_by_score = face_search_by_embeddings(face_embs)
precision_model = PrecisionModel(face_ids_by_bucket)
print('Select all MISTAKES. Ordered by DESCENDING score. Expecting {} frames'.format(precision_model.get_lower_count()))
lower_widget = precision_model.get_lower_widget()
lower_widget
print('Select all NON-MISTAKES. Ordered by ASCENDING distance. Expecting {} frames'.format(precision_model.get_upper_count()))
upper_widget = precision_model.get_upper_widget()
upper_widget
```
Run the following cell after labelling.
```
lower_precision = precision_model.compute_precision_for_lower_buckets(lower_widget.selected)
upper_precision = precision_model.compute_precision_for_upper_buckets(upper_widget.selected)
precision_by_bucket = {**lower_precision, **upper_precision}
results = FaceIdentityModel(
name=name,
face_ids_by_bucket=face_ids_by_bucket,
face_ids_to_score=face_ids_to_score,
precision_by_bucket=precision_by_bucket,
model_params={
'images': list(zip(face_embs, face_imgs))
}
)
plot_precision_and_cdf(results)
# Save the model
results.save()
```
# Analysis
## Gender Breakdown
```
gender_breakdown = compute_gender_breakdown(results)
print('Raw counts:')
for k, v in gender_breakdown.items():
print(' ', k, ':', v)
print()
print('Proportions:')
denominator = sum(v for v in gender_breakdown.values())
for k, v in gender_breakdown.items():
print(' ', k, ':', v / denominator)
print()
print('Showing examples:')
show_gender_examples(results)
```
## Face Sizes
```
plot_histogram_of_face_sizes(results)
```
## Appearances on a Single Show
```
show_name = 'Hannity'
screen_time_by_video_id = compute_screen_time_by_video(results, show_name)
plot_histogram_of_screen_times_by_video(name, show_name, screen_time_by_video_id)
plot_screentime_over_time(name, show_name, screen_time_by_video_id)
plot_distribution_of_appearance_times_by_video(results, show_name)
```
## Screen Time Across All Shows
```
screen_time_by_show = get_screen_time_by_show(results)
plot_screen_time_by_show(name, screen_time_by_show)
```
# Persist to Cloud
## Save Model to GCS
```
gcs_model_path = results.save_to_gcs()
```
### Make sure the GCS file is valid
```
gcs_results = FaceIdentityModel.load_from_gcs(name=name)
plot_precision_and_cdf(gcs_results)
```
## Save Labels to DB
```
from django.core.exceptions import ObjectDoesNotExist
def standardize_name(name):
return name.lower()
person_type = ThingType.objects.get(name='person')
try:
person = Thing.objects.get(name=standardize_name(name), type=person_type)
print('Found person:', person.name)
except ObjectDoesNotExist:
person = Thing(name=standardize_name(name), type=person_type)
print('Creating person:', person.name)
labeler = Labeler(name='face-identity-{}'.format(person.name), data_path=gcs_model_path)
```
### Commit the person and labeler
```
person.save()
labeler.save()
```
### Commit the FaceIdentity labels
```
commit_face_identities_to_db(results, person, labeler)
print('Committed {} labels to the db'.format(FaceIdentity.objects.filter(labeler=labeler).count()))
```
| github_jupyter |
```
import pandas as pd
import datetime
import numpy as np
```
## Loading Production Data from 1987 to 2008
The production data from these years follows the same file format.
We can therefore import using the same format and put the dataframes into a dictionary.
In 1990 we manually fix well API No: 21451, DUCKETT "A" and set it's well number to 1 as unspecified.
Same in 1991.
```
dates_cols_oil = ["OIL."+str(i) for i in range(0, 12, 1)]
dates_cols_gas = ["GAS."+str(i) for i in range(0, 12, 1)]
dates_cols = dates_cols_oil + dates_cols_gas
headers_old_2003 = ['API_COUNTY', 'API_NUMBER', 'SUFFIX', 'WELL_NAME','WELL_NO', ' OPER_NO', 'OPER_SUFFIX',
'OPERATOR', 'ME', 'SECTION', 'TWP','RAN', 'Q4', 'Q3', 'Q2', 'Q1', 'LATITUDE', 'LONGITUDE', 'OTC_COUNTY',
'OTC_LEASE_NO', 'OTC_SUB_NO', 'OTC_MERGE', 'POOL_NO', 'CODE','FORMATION', 'OFB', 'ALLOWABLE_CLASS',
'ALLOWABLE_TYPE', ' PURCH_NO',
'PURCHASER', 'PURCH_SUFFIX', 'OFB.1',
'YEAR', 'JAN', 'OIL.0', 'GAS.0', 'FEB',
'OIL.1', 'GAS.1', 'MAR', 'OIL.2', 'GAS.2',
'APR', 'OIL.3', 'GAS.3', 'MAY', 'OIL.4',
'GAS.4', 'JUN', 'OIL.5', 'GAS.5', 'JUL',
'OIL.6', 'GAS.6', 'AUG', 'OIL.7', 'GAS.7',
'SEP', 'OIL.8', 'GAS.8', 'OCT', 'OIL.9',
'GAS.9', 'NOV', 'OIL.10', 'GAS.10', 'DEC',
'OIL.11', 'GAS.11']
headers_new_2004 = ['API_COUNTY', 'API_NUMBER', 'S', 'WELL_NAME','WELL_NO', ' OPER_NO',
'OPERATOR', 'ME', 'SECTION', 'TWP','RAN', 'Q4', 'Q3', 'Q2', 'Q1', 'LATITUDE', 'LONGITUDE', 'OTC_COUNTY',
'OTC_LEASE_NO', 'OTC_SUB_NO', 'OTC_MERGE', 'POOL_NO', 'CODE','FORMATION','ALLOWABLE_CLASS',
'ALLOWABLE_TYPE', ' PURCH_NO',
'PURCHASER', 'OFB.1',
'YEAR', 'JAN', 'OIL.0', 'GAS.0', 'FEB',
'OIL.1', 'GAS.1', 'MAR', 'OIL.2', 'GAS.2',
'APR', 'OIL.3', 'GAS.3', 'MAY', 'OIL.4',
'GAS.4', 'JUN', 'OIL.5', 'GAS.5', 'JUL',
'OIL.6', 'GAS.6', 'AUG', 'OIL.7', 'GAS.7',
'SEP', 'OIL.8', 'GAS.8', 'OCT', 'OIL.9',
'GAS.9', 'NOV', 'OIL.10', 'GAS.10', 'DEC',
'OIL.11', 'GAS.11']
df_in = None
production_data = {}
for i in range(1987, 2016, 1):
dates_oil = [ "OIL_"+str(datetime.date(i, j+1, 1)) for j in range(0, 12, 1)]
dates_gas = [ "GAS_"+str(datetime.date(i, j+1, 1)) for j in range(0, 12, 1)]
renamed_oil = {old: new for old, new in zip(dates_cols_oil, dates_oil)}
renamed_gas = {old: new for old, new in zip(dates_cols_gas, dates_gas)}
renamed_cols = {**renamed_oil, **renamed_gas}
#print(renamed_cols)
if i != 1994: #No Data from 1994
print(i)
if i <= 2008:
df = None
if i < 2004:
df = pd.read_csv("../raw/"+str(i)+"prodn.txt", delimiter="|", skiprows=[0, 2], names=headers_old_2003)
else:
df = pd.read_csv("../raw/"+str(i)+"prodn.txt", delimiter="|", skiprows=[0, 2], names=headers_new_2004)
df_in = df.copy()
print(df.columns)
print(renamed_cols)
df.rename(index=str, columns=renamed_cols, inplace=True)
df = df.drop(['YEAR','JAN', 'FEB', 'MAR', 'APR', 'MAY', 'JUN', 'JUL','AUG', 'SEP', 'OCT', 'NOV', 'DEC'], axis=1)
production_data[i] = df
else:
df = pd.read_csv("../raw/"+str(i)+"prodn.txt", delimiter="|")
df[["API_COUNTY", "API_NUMBER"]].apply(lambda x: pd.to_numeric(x, errors='coerce',downcast='integer'))
df_in = df.copy()
df.rename(renamed_cols)
production_data[i] = df
df_in.head()
def filter_data(row):
buffer = []
for val in row:
val_parsed = None
try:
val_parsed = int(val)
except ValueError:
val_parsed = 0
buffer.append(val_parsed)
return np.array(buffer, dtype=np.int32)
meta_dataframe = None
meta_prod_dfs = []
meta_data = {}
columns = ['API_NUMBER','API_COUNTY','LATITUDE', 'LONGITUDE', 'FORMATION']
for year in range(1987, 2016):
print(year)
if year != 1994:
filter_col = columns
yearly_meta_data = production_data[year]#.dropna()
for i in range(1, len(yearly_meta_data.index)):
row = yearly_meta_data.iloc[[i]]
api_num = row["API_NUMBER"].values.astype(np.int32)[0]
mdata = row[filter_col].values[0]
if api_num in meta_data.keys():
pass
else:
if not np.isnan(api_num):
meta_data[api_num] = {}
try:
meta_data[api_num]["API_COUNTY"] = int(mdata[1])
meta_data[api_num]["LATITUDE"] = float(mdata[2])
meta_data[api_num]["LONGITUDE"] = float(mdata[3])
form_str = str(mdata[4]).strip(" ")
meta_data[api_num]["FORMATION"] = form_str
except ValueError:
print("Found invalid value: ", api_num, year, mdata)
del meta_data[-2147483648 ]
meta_out = {}
for key in meta_data.keys():
meta_out[str(key)] = meta_data[key]
with open('../processed/immutable/immutable.json', 'w') as fp:
json.dump(meta_out, fp, sort_keys=True)
```
| github_jupyter |
```
import statistical_analysis_toolkit as stat_tools
import pandas as pd
import scipy.stats as stats
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
#que up some data
data = pd.DataFrame.from_dict({"fired":[34, 37, 37, 38, 41, 42, 43, 44, 44, 45, 45, 45, 46, 48, 49, 53, 53, 54, 54, 55, 56],
'not_fired':[27, 33, 36, 37, 38, 38, 39, 42, 42, 43, 43, 44, 44, 44, 45, 45, 45, 45, 46, 46, 47, 47, 48, 48, 49, 49, 51, 51, 52, 54]}
, orient='index').T
#expected to have two columns (one for each sample, col_names = feature_names)
#null hyp (equal means)
h0 = 0.0
#95% CL
alpha = 0.05
analysis = stat_tools.analyze_distribution(data, alpha=alpha, h0=h0)
p_val = analysis.loc['POOLED'].p_value_t_test
evidence = stat_tools.strength_of_evidence(p_val)
print(f'Base on a p-value of {p_val} there is {evidence} evidence against H0={h0}')
analysis
#n = stat_tools.critical_norm(.01)
df = 8
a = 0.05
t = stat_tools.critical_t(a, df) #two sided
t1 = stats.t.ppf(a, df) #one sided, lower
[t, t1]
p = stat_tools.p_value_from_t(-2.54, 8, sides=1) #normal distribution - add t-distribution as well?
data = pd.read_csv('C:\SMU\Stats\Homework\HW4\EducationData.csv')
data = pd.DataFrame.from_dict({'12 Years':data[data.Educ == 12].Income2005,
'16 Years':data[data.Educ == 16].Income2005}, orient='index').T
#expected to have two columns (one for each sample, col_names = feature_names)
#null hyp (equal means)
h0 = 0.0
#95% CL
alpha = 0.05
analysis = stat_tools.analyze_distribution(data, alpha=alpha, h0=h0)
p_val = analysis.loc['POOLED'].p_value_t_test
evidence = stat_tools.strength_of_evidence(p_val)
print(f'Base on a p-value of {p_val} there is {evidence} evidence against H0={h0}')
analysis
data = pd.read_csv('~/Documents/SMU/Stats/Homework/HW4/Autism.csv')[['After', 'Before']]
#expected to have two columns (one for each sample, col_names = feature_names)
#null hyp (equal means)
h0 = 0.0
#95% CL
alpha = 0.05
analysis = stat_tools.analyze_distribution(data, alpha=alpha, h0=h0, is_log_data=False)
p_val = analysis.loc['POOLED'].p_value_t_test
evidence = stat_tools.strength_of_evidence(p_val)
print(f'Base on a p-value of {p_val} there is {evidence} evidence against H0={h0}')
analysis
education_data = pd.read_csv('/Users/skennedy/Documents/SMU/Stats/Homework/HW5/ex0525.csv')[['Educ', 'Income2005']]
#education_data['LogIncome2005'] = np.log(education_data.Income2005)
education_data.Educ.replace('<12', '12', inplace=True)
education_data.Educ.replace('>16', '16', inplace=True)
education_data.to_csv('education_data.csv')
anova(education_data, 'Educ', 'Income2005')
#{ for education_level, data in education_data.groupby(['Educ'])}
#prep data for an anova test (one data frame with multiple columns as samples)
def anova(data, partition_column, observable_column):
#test assumptions of anova graphically and numerically
'''1) Normality: Similar to t-tools hypothesis testing, ANOVA is robust to this assumption.
Extremely long-tailed distributions (outliers) or skewed distributions,
coupled with different sample sizes (especially when the sample sizes are small)
present the only serious distributional problems.'''
'''2) Equal Standard Deviations: This assumption is crucial, paramount, and VERY important.'''
'''3) The assumptions of independence within and across groups are critical.
If lacking, different analysis should be attempted.'''
#Create logical groupings
groups = data.groupby([partition_column])
hist_fig, hist_ax = plt.subplots(len(groups), 1, sharex=True)
hist_fig.suptitle('Histograms Raw Data')
hist_fig_log, hist_ax_log = plt.subplots(len(groups), 1, sharex=True)
hist_fig_log.suptitle('Histograms Log Data')
i = 0
group_stats = []
for group_key, group in groups:
clean_data = group[observable_column].dropna()
group_std = clean_data.std()
group_mean = clean_data.mean()
#plot histograms
hist_ax[i].hist(clean_data)
hist_ax[i].set_title(group_key)
log_data = np.log(clean_data[clean_data != 0])
hist_ax_log[i].hist(log_data)
i = i + 1
```
| github_jupyter |
# Agglomerative Hierarchical Clustering
In this technique, initially each data point is considered as an individual cluster. At each iteration, the similar clusters merge with other clusters until one cluster or K clusters are formed. There are multiple ways to calculate similarity between clusters. In this project we used 2 ways-
#### 1) Single Linkage (MIN)
#### 2) Complete Linkage (MAX)
```
import numpy as np
import pandas as pd
from math import sqrt
dataset = pd.read_csv('india-basemap/data_clubbed.csv', header=None)
dataset.head()
states = dataset[0]
dataset.drop(0, inplace=True, axis=1)
dataset.head()
```
### Year-wise Clustering Analysis
We perform year-wise clustering analysis for all states. To find the clusters for a particular based on their crime records, specify the year in the 'year' variable
```
year = 2014
year_list = []
start_range = 0
for i in range(len(dataset)):
if dataset[1].loc[i] == year:
start_range = i
break
end_range = 0
for i in range(len(dataset)):
if dataset[1].loc[i] == year:
end_range += 1
end_range = end_range + start_range
dataset.drop(1, inplace=True, axis=1)
for i in range(start_range, end_range+1):
year_list.append(dataset.loc[i])
```
### Step 1 - Distance Matrix
Euclidean Distance is used to calculate the similarity between 2 rows of the dataset. This is used to create the distance matrix.
```
def euclidean_distance(row1, row2):
distance = 0.0
for i in range(0,11):
distance += (row1[i] - row2[i])**2
return sqrt(distance)
```
### Step 2 - Smallest Value
The smallest value in the upper triangle of the distance matrix is used to find the smallest distance between 2 cluster.
```
def smallest_val(arr, clusters, n, linkage):
smallest = 10**9
small_i = 0
small_j = 0
for i in range(0, n):
for j in range(i+1, n):
if arr[i][j] < smallest:
smallest = arr[i][j]
small_i = i
small_j = j
update_matrix(arr, clusters, small_i, small_j, n, linkage)
#print(arr)
return smallest
```
### Step 3 - Update the distance matrix
The distance matrix is updated according to the linkage specified in the argument. The 2 different linkages are -
- Single Linkage - Sim(C1,C2) = Min Sim(Pi,Pj) such that Pi ∈ C1 & Pj ∈ C2
- Complete Linkage - im(C1,C2) = Max Sim(Pi,Pj) such that Pi ∈ C1 & Pj ∈ C2
```
def update_matrix(arr, clusters, i, j, n, linkage):
temp_list = []
temp_list.append(i)
temp_list.append(j)
temp_list.append(arr[i][j])
clusters.append(temp_list)
print('Merged' , i, j, 'with distance', arr[i][j])
for a in range(0, n):
for b in range(a+1, n):
if ((a == i) and (b != j)) or ((a == j) and (b != i)):
if linkage == 'single':
arr[a][b] = min(arr[i][b], arr[j][b])
arr[b][a] = min(arr[i][b], arr[j][b])
elif linkage == 'complete':
arr[a][b] = max(arr[i][b], arr[j][b])
arr[b][a] = max(arr[i][b], arr[j][b])
elif ((a != i) and (b == j)) or ((a != j) and (b == i)):
if linkage == 'single':
arr[a][b] = min(arr[a][i], arr[a][j])
arr[b][a] = min(arr[a][i], arr[a][j])
elif linkage == 'complete':
arr[a][b] = max(arr[a][i], arr[a][j])
arr[b][a] = max(arr[a][i], arr[a][j])
for a in range(0, n):
arr[i][a] = 10**9
arr[a][i] = 10**9
data = dataset.values.tolist()
n = end_range - start_range
arr = [[0 for i in range(n)] for j in range(n)]
for i in range(start_range, end_range):
for j in range(i+1, end_range):
arr[i-start_range][j-end_range] = euclidean_distance(data[i],data[j])
arr[j-end_range][i-start_range] = euclidean_distance(data[i],data[j])
```
### Step 4 - Find the clusters using DFS
The connected components i.e the clusters are found by creating a graph and finding the connect components using DFS.
```
class Graph:
def __init__(self,V):
self.V = V
self.adj = [[] for i in range(V)]
def DFSUtil(self, temp, v, visited):
visited[v] = True
temp.append(v)
for i in self.adj[v]:
if visited[i] == False:
temp = self.DFSUtil(temp, i, visited)
return temp
def addEdge(self, v, w):
self.adj[v].append(w)
self.adj[w].append(v)
def connectedComponents(self):
visited = []
cc = []
for i in range(self.V):
visited.append(False)
for v in range(self.V):
if visited[v] == False:
temp = []
cc.append(self.DFSUtil(temp, v, visited))
return cc
```
### Results -
- Using elbow method we found that 10 clusters yield the best results for this dataset
- After finding the first 10 clusters, we perform a demographic survey of the data
- The states that fall into the same cluster have similar crime patterns i.e the number and type of crimes
- It is noticed that all the coastal states like Lakshadweep, Andaman and Nicobar Islands, Daman & Diu and D&N haveli fall into the same cluster very often
- Another observation is that most of the Northeastern states fall into the same cluster
```
g = Graph(n);
linkage = 'complete'
clusters = []
labelList = []
for cnt in range(0,n-1):
val = smallest_val(arr, clusters, n, linkage)
for i in range(0,len(clusters)):
g.addEdge(clusters[i][0], clusters[i][1])
cc = g.connectedComponents()
if len(cc) == 10:
print('\n')
for i in range(0, len(cc)):
temp_states = []
for j in range(0, len(cc[i])):
temp_states.append(states[cc[i][j]])
print("Cluster "+ str(i)+" :"+ str(temp_states))
labelList.append(" ".join(temp_states))
break
labelList
```
### Cross-checking the results
The scipy library is used to cross-check the results and to get a dendogram for the data
```
from scipy.cluster.hierarchy import dendrogram, linkage
from matplotlib import pyplot as plt
X = np.array(year_list)
Z = linkage(X, 'complete')
fig = plt.figure(figsize=(25, 15))
plt.xlabel('States')
plt.ylabel('Cluster Distance')
dn = dendrogram(Z, orientation='right')
```
| github_jupyter |
# Compute the density of railspace patches
For each railspace patch, this notebook computes the percentage of neighboring railspace patches, i.e.:
```python
railspace_neighboring_patches / total_neighboring_patches
```
The neighboring patches are all patches in a user-defined radius (in meters).
```
# solve issue with autocomplete
%config Completer.use_jedi = False
%load_ext autoreload
%autoreload 2
%matplotlib notebook
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:80% !important; }</style>"))
import numpy as np
import pandas as pd
import pyproj
from scipy import spatial
```
## Inputs
```
path2all_patches = "XXX"
path2rail_patches = "XXX"
path2save_output = "XXX"
# user-defined radius (in meters) for neighbors
distance_in_meters = 500
# --- example
# path2all_patches = "./resources/all_patches_latlonpred.csv"
# path2rail_patches = "./results_v003/pred_0103_keep_1_250.csv"
# path2save_output = "./df_pred_0103_rail_density.csv"
```
## Read all patches
We use this to later compute `total_neighboring_patches` in this equation:
```python
railspace_neighboring_patches / total_neighboring_patches
```
```
all_patches_pd = pd.read_csv(path2all_patches)
```
## KD-tree for all patches
```
ecef = pyproj.Proj(proj='geocent', ellps='WGS84', datum='WGS84')
lla = pyproj.Proj(proj='latlong', ellps='WGS84', datum='WGS84')
x, y, z = pyproj.transform(lla, ecef,
all_patches_pd["center_lon"].to_numpy(),
all_patches_pd["center_lat"].to_numpy(),
np.zeros(len(all_patches_pd["center_lat"])),
radians=False)
# add x, y, z to df
all_patches_pd["x"] = x
all_patches_pd["y"] = y
all_patches_pd["z"] = z
kdtree_patches = spatial.cKDTree(all_patches_pd[["x", "y", "z"]].to_numpy())
```
## Read railspace patches
```
df_pred_0103 = pd.read_csv(path2rail_patches, index_col=0)
df_pred_0103
```
## KD-tree for railspace patches
```
ecef = pyproj.Proj(proj='geocent', ellps='WGS84', datum='WGS84')
lla = pyproj.Proj(proj='latlong', ellps='WGS84', datum='WGS84')
x, y, z = pyproj.transform(lla, ecef,
df_pred_0103["center_lon"].to_numpy(),
df_pred_0103["center_lat"].to_numpy(),
np.zeros(len(df_pred_0103["center_lat"])),
radians=False)
# add x, y, z to df
df_pred_0103["x"] = x
df_pred_0103["y"] = y
df_pred_0103["z"] = z
kdtree_0103 = spatial.cKDTree(df_pred_0103[["x", "y", "z"]].to_numpy())
kdtree_pred_0103 = spatial.cKDTree(df_pred_0103[["x", "y", "z"]].to_numpy())
```
## Calculate density
Now we can compute the terms in:
```python
railspace_neighboring_patches / total_neighboring_patches
```
```
print("[INFO] rails.....")
railspace_neighboring_patches = kdtree_pred_0103.query_ball_tree(kdtree_pred_0103, distance_in_meters)
print("[INFO] patches...")
total_neighboring_patches = kdtree_pred_0103.query_ball_tree(kdtree_patches, distance_in_meters)
num_railspace_neighboring_patches = [len(x) for x in railspace_neighboring_patches]
num_total_neighboring_patches = [len(x) for x in total_neighboring_patches]
df_pred_0103["perc_neigh_rails"] = num_railspace_neighboring_patches / (np.array(num_total_neighboring_patches) + 1e-32) * 100.
df_pred_0103.iloc[0]
df_pred_0103.to_csv(path2save_output)
```
| github_jupyter |
<h1>Overview<span class="tocSkip"></span></h1>
<div class="toc"><ul class="toc-item"><li><span><a href="#WMA-Tokenization" data-toc-modified-id="WMA-Tokenization-1"><span class="toc-item-num">1 </span>WMA Tokenization</a></span><ul class="toc-item"><li><span><a href="#WMA-en-de-train/val" data-toc-modified-id="WMA-en-de-train/val-1.1"><span class="toc-item-num">1.1 </span>WMA en-de train/val</a></span></li></ul></li></ul></div>
```
import os
import re
import sys
import numpy as np
import pandas as pd
from time import sleep
from keras import backend as K
from keras.models import Model
from keras.models import Sequential as SequentialModel
from keras.layers import Dense, Conv1D, LSTM, Dropout, Embedding, Layer, Input, Flatten, concatenate as Concatenate, Lambda
from keras.callbacks import Callback
from keras.utils import to_categorical
from keras.preprocessing.text import Tokenizer as KerasTokenizer
%load_ext autoreload
%autoreload 2
%reload_ext autoreload
sys.path.insert(0, '../ct')
import load
from preprocess import preprocess
from preprocess import Tokenizer
from preprocess.preprocess import separator_samples
from model.layers import LayerNormalization
from model.layers import ContentBasedAttention_CT
from model.layers import ScaledDotProductAttention
from model.layers import MultiHeadAttention
from model import CompressiveTransformer
from load.wma import load as load_wma
def file_utf_to_ascii(input_path, output_path=None):
if output_path is None:
s = input_path.split('.')
output_path = '.'.join(s[:-1]) + '-ascii.' + s[-1]
with open(input_path, 'r', encoding='utf8') as file:
content = file.read()
content = content.encode('ascii', 'xmlcharrefreplace')
content = content.decode('ascii')
with open(output_path, 'w', encoding='ascii') as file:
file.write(content)
print(f'converted utf->ascii for {input_path}')
# for p in input_paths:
# file_utf_to_ascii(p)
```
# WMA Tokenization
```
vocab_size=1024
lowercase=False
input_paths = {'en': '../data/wma-en-de/input/train-en.txt',
'de': '../data/wma-en-de/input/train-de.txt'}
tokenizer_output_path = f'../data/wma-en-de/tokenizer/en-de-v0-t{vocab_size}' \
f'{"-lowercase" if lowercase else ""}.tok'
tokenizer = Tokenizer(input_paths=list(input_paths.values()),
tokenizer_output_path=tokenizer_output_path,
vocab_size=vocab_size,
lowercase=lowercase)
hello = tokenizer.encode_batch(['hello'])[0]
hello
print(hello.ids)
print(hello.tokens)
```
## WMA en-de train/val
```
from load.wma import load as load_wma
wma = load_wma(input_paths['en'],
input_paths['de'])
wma
english_encodings = tokenizer.encode_batch(wma.english.tolist())
wma['english_ids'] = [encoding.ids for encoding in english_encodings]
del english_encodings
# german_encodings = tokenizer.encode_batch(wma.german.tolist())
# wma['german_ids'] = [encoding.ids for encoding in german_encodings]
# del german_encodings
wma[['english_ids']]
val_index = int(len(wma)*0.8)
x_train = wma[['english_ids']][:val_index]
x_val = wma[['english_ids']][-val_index:]
x_train.to_pickle('../data/wma-en-de/processed/train-en.pkl.zip')
x_val.to_pickle('../data/wma-en-de/processed/val-en.pkl.zip')
```
| github_jupyter |
# Recap
Büyük veri nedir?
Büyük veri bir problemdir. Geleneksel yöntemlerle işlenemeyen verilere büyük veri denir.
Apache Hadoop: Büyük veri ekosisteminin temeli.
MapReduce: Bir bilgisayar kümesinin tek bir bilgisayar gibi hareket etmesini sağlayacak paralel programlama modelidir.
Apache Spark: Cluster üzerinde hızlı bilgi işleme kütühanesidir.
RDDs: Veri bellek içi işlenerek hız kazanılmıştır.
# Kurulum
```
import warnings
warnings.filterwarnings("ignore", category=DeprecationWarning)
warnings.filterwarnings("ignore", category=FutureWarning)
#!pip install findspark
import findspark
findspark.init("/Users/mvahit/spark/spark-3.0.0-preview-bin-hadoop2.7")
import pyspark
from pyspark import SparkContext
from pyspark.sql import SparkSession
from pyspark.conf import SparkConf
spark = SparkSession.builder \
.master("local") \
.appName("pyspark_giris") \
.getOrCreate()
sc = spark.sparkContext
sc
#sc.stop()
```
# Temel DataFrame İşlemleri
```
spark_df = spark.read.csv("./churn.csv", header = True, inferSchema = True)
spark_df
type(spark_df)
spark_df.printSchema()
import seaborn as sns
df = sns.load_dataset("diamonds")
type(df)
df.head()
spark_df.head()
df.dtypes
spark_df.dtypes
df.ndim
spark_df.ndim
spark_df.show(3, truncate = True)
spark_df.count()
spark_df.columns
spark_df.describe().show()
spark_df.describe("Age").show()
spark_df.select("Age","Names").show()
spark_df.filter(spark_df.Age > 40).count()
spark_df.filter(spark_df.Age > 40).show()
spark_df.groupby("Churn").count().show()
spark_df.groupby("Churn").agg({"Age":"mean"}).show()
```
# SQL İşlemleri
```
spark_df.createOrReplaceTempView("tbl_df")
spark.sql("show databases").show()
spark.sql("show tables").show()
spark.sql("select Age from tbl_df").show(5)
spark.sql("select Churn, mean(Age) from tbl_df group by Churn").show()
```
1. Senaryo: Server'da python notebook araçlar kurulu olabilir.
- Basit bağlantı ayarlarıyla server'daki final tablo ile python arasında bağlantı sağlanıp veri server ortamındaki python'da işlenebilir hale gelir.
2. Senaryo: Final tablolar oluşturumuş olur. SQL ile yapılır. Final tabloları locale cekilmek durumundadır.
3. Senaryo: Big data
# Büyük Veride Görselleştirme
```
import matplotlib.pyplot as plt
import seaborn as sns
sns.barplot(x = "Churn", y = spark_df.Churn.index, data = spark_df)
spark_df.groupby("Churn").agg({"Age":"mean"}).show()
spark_df.groupby("Churn").agg({"Age":"mean"}).toPandas()
sdf = spark_df.toPandas()
sns.barplot(x = "Churn", y = sdf.Churn.index, data = sdf);
```
# GBM ile Müşteri Terk Modellemesi
```
spark_df = spark.read.csv("./churn.csv", header = True,inferSchema = True)
spark_df.show(5)
spark_df = spark_df.toDF(*[c.lower() for c in spark_df.columns])
spark_df.show(5)
spark_df = spark_df.withColumnRenamed("_c0", "index")
spark_df.show(5)
spark_df.count()
len(spark_df.columns)
spark_df.describe().show()
spark_df.select("age","total_purchase","account_manager","years","num_sites","churn").describe().toPandas().transpose()
spark_df = spark_df.dropna()
spark_df = spark_df.withColumn("age_kare", spark_df.age**2)
spark_df.show()
#bagimli degiskeni belirtmek
from pyspark.ml.feature import StringIndexer
stringIndexer = StringIndexer(inputCol = "churn", outputCol = "label")
mod = stringIndexer.fit(spark_df)
indexed = mod.transform(spark_df)
spark_df = indexed.withColumn("label", indexed["label"].cast("integer"))
spark_df.show()
#bagimsiz degiskenler
from pyspark.ml.feature import VectorAssembler
spark_df.columns
cols = ["age","total_purchase","account_manager","years","num_sites"]
va = VectorAssembler(inputCols = cols, outputCol = "features")
va_df = va.transform(spark_df)
va_df.show()
final_df = va_df.select(["features","label"])
final_df.show()
splits = final_df.randomSplit([0.70, 0.30])
train_df = splits[0]
test_df = splits[1]
test_df.show()
#base model
from pyspark.ml.classification import GBTClassifier
gbm = GBTClassifier(maxIter = 10, featuresCol = "features", labelCol = "label")
gbm_model = gbm.fit(train_df)
sc
y_pred = gbm_model.transform(test_df)
y_pred.show(20)
ac = y_pred.select("label","prediction")
ac.show(3)
ac.filter(ac.label == ac.prediction).count()/ac.count()
#model tuning
from pyspark.ml.evaluation import BinaryClassificationEvaluator
from pyspark.ml.tuning import ParamGridBuilder, CrossValidator
evaluator = BinaryClassificationEvaluator()
paramGrid = (ParamGridBuilder()
.addGrid(gbm.maxDepth, [2, 4, 6])
.addGrid(gbm.maxBins, [20, 30])
.addGrid(gbm.maxIter, [10, 20])
.build())
cv = CrossValidator(estimator = gbm, estimatorParamMaps = paramGrid, evaluator = evaluator, numFolds = 10)
cv_model = cv.fit(train_df)
y_pred = cv_model.transform(test_df)
ac = y_pred.select("label","prediction")
ac.filter(ac.label == ac.prediction).count() / ac.count()
# yeni ya da var olan bir musterimiz var. Bu musteri bizi terk eder mi?
import pandas as pd
names = pd.Series(["Ali Ahmetoğlu", "Taner Gün", "Berkay","Polat Konak", "Kamil Atasoy"])
age = pd.Series([38, 43, 34, 50, 40])
total_purchase = pd.Series([30000, 10000, 6000, 30000, 100000])
account_manager = pd.Series([1,0,0,1,1])
years = pd.Series([20, 10, 3, 8, 30])
num_sites = pd.Series([30,8,8,6,50])
yeni_musteriler = pd.DataFrame({
'names':names,
'age': age,
'total_purchase': total_purchase,
'account_manager': account_manager ,
'years': years,
'num_sites': num_sites})
yeni_musteriler.columns
yeni_musteriler.head()
yeni_sdf = spark.createDataFrame(yeni_musteriler)
yeni_sdf.show(3)
new_customers = va.transform(yeni_sdf)
new_customers.show(3)
results = cv_model.transform(new_customers)
results.select("names","prediction").show()
sc.stop()
```
| github_jupyter |
# 3. LightGBM_GridSearchCV
**Start from the most basic features, and try to improve step by step.**
## Run name
```
import time
project_name = 'TalkingdataAFD2018'
step_name = 'LightGBM_GridSearchCV'
time_str = time.strftime("%Y%m%d_%H%M%S", time.localtime())
run_name = '%s_%s_%s' % (project_name, step_name, time_str)
print('run_name: %s' % run_name)
t0 = time.time()
```
## Important params
```
date = 6
print('date: ', date)
test_n_rows = None
# test_n_rows = 18790469
# test_n_rows = 10*10000
day_rows = {
0: {
'n_skiprows': 1,
'n_rows': 10 * 10000
},
6: {
'n_skiprows': 1,
'n_rows': 9308568
},
7: {
'n_skiprows': 1 + 9308568,
'n_rows': 59633310
},
8: {
'n_skiprows': 1 + 9308568 + 59633310,
'n_rows': 62945075
},
9: {
'n_skiprows': 1 + 9308568 + 59633310 + 62945075,
'n_rows': 53016937
}
}
n_skiprows = day_rows[date]['n_skiprows']
n_rows = day_rows[date]['n_rows']
```
## Import PKGs
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
from IPython.display import display
import os
import gc
import time
import random
import zipfile
import h5py
import pickle
import math
from PIL import Image
import shutil
from tqdm import tqdm
import multiprocessing
from multiprocessing import cpu_count
from sklearn.model_selection import train_test_split, GridSearchCV
from sklearn.metrics import accuracy_score
random_num = np.random.randint(10000)
print('random_num: %s' % random_num)
```
## Project folders
```
cwd = os.getcwd()
input_folder = os.path.join(cwd, 'input')
output_folder = os.path.join(cwd, 'output')
model_folder = os.path.join(cwd, 'model')
log_folder = os.path.join(cwd, 'log')
print('input_folder: \t\t\t%s' % input_folder)
print('output_folder: \t\t\t%s' % output_folder)
print('model_folder: \t\t\t%s' % model_folder)
print('log_folder: \t\t\t%s' % log_folder)
train_csv_file = os.path.join(input_folder, 'train.csv')
train_sample_csv_file = os.path.join(input_folder, 'train_sample.csv')
test_csv_file = os.path.join(input_folder, 'test.csv')
sample_submission_csv_file = os.path.join(input_folder, 'sample_submission.csv')
print('\ntrain_csv_file: \t\t%s' % train_csv_file)
print('train_sample_csv_file: \t\t%s' % train_sample_csv_file)
print('test_csv_file: \t\t\t%s' % test_csv_file)
print('sample_submission_csv_file: \t%s' % sample_submission_csv_file)
```
## Load data
```
%%time
train_csv = pd.read_csv(train_csv_file, skiprows=range(1, n_skiprows), nrows=n_rows, parse_dates=['click_time'])
test_csv = pd.read_csv(test_csv_file, nrows=test_n_rows, parse_dates=['click_time'])
sample_submission_csv = pd.read_csv(sample_submission_csv_file)
print('train_csv.shape: \t\t', train_csv.shape)
print('test_csv.shape: \t\t', test_csv.shape)
print('sample_submission_csv.shape: \t', sample_submission_csv.shape)
print('train_csv.dtypes: \n', train_csv.dtypes)
display(train_csv.head(2))
display(test_csv.head(2))
display(sample_submission_csv.head(2))
y_data = train_csv['is_attributed']
train_csv.drop(['is_attributed'], axis=1, inplace=True)
display(y_data.head())
```
## Features
```
train_csv['day'] = train_csv['click_time'].dt.day.astype('uint8')
train_csv['hour'] = train_csv['click_time'].dt.hour.astype('uint8')
train_csv['minute'] = train_csv['click_time'].dt.minute.astype('uint8')
train_csv['second'] = train_csv['click_time'].dt.second.astype('uint8')
print('train_csv.shape: \t', train_csv.shape)
display(train_csv.head(2))
test_csv['day'] = test_csv['click_time'].dt.day.astype('uint8')
test_csv['hour'] = test_csv['click_time'].dt.hour.astype('uint8')
test_csv['minute'] = test_csv['click_time'].dt.minute.astype('uint8')
test_csv['second'] = test_csv['click_time'].dt.second.astype('uint8')
print('test_csv.shape: \t', test_csv.shape)
display(test_csv.head(2))
arr = np.array([[3,6,6],[4,5,1]])
print(arr)
np.ravel_multi_index(arr, (7,6))
print(arr)
print(np.ravel_multi_index(arr, (7,6), order='F'))
def df_add_counts(df, cols, tag="_count"):
arr_slice = df[cols].values
unq, unqtags, counts = np.unique(np.ravel_multi_index(arr_slice.T, arr_slice.max(0) + 1), return_inverse=True, return_counts=True)
df["_".join(cols) + tag] = counts[unqtags]
return df
def df_add_uniques(df, cols, tag="_unique"):
gp = df[cols] \
.groupby(by=cols[0:len(cols) - 1])[cols[len(cols) - 1]] \
.nunique() \
.reset_index() \
.rename(index=str, columns={cols[len(cols) - 1]: "_".join(cols)+tag})
df = df.merge(gp, on=cols[0:len(cols) - 1], how='left')
return df
train_csv = df_add_counts(train_csv, ['ip', 'day', 'hour'])
train_csv = df_add_counts(train_csv, ['ip', 'app'])
train_csv = df_add_counts(train_csv, ['ip', 'app', 'os'])
train_csv = df_add_counts(train_csv, ['ip', 'device'])
train_csv = df_add_counts(train_csv, ['app', 'channel'])
train_csv = df_add_uniques(train_csv, ['ip', 'channel'])
display(train_csv.head())
test_csv = df_add_counts(test_csv, ['ip', 'day', 'hour'])
test_csv = df_add_counts(test_csv, ['ip', 'app'])
test_csv = df_add_counts(test_csv, ['ip', 'app', 'os'])
test_csv = df_add_counts(test_csv, ['ip', 'device'])
test_csv = df_add_counts(test_csv, ['app', 'channel'])
test_csv = df_add_uniques(test_csv, ['ip', 'channel'])
display(test_csv.head())
```
## Prepare data
```
train_useless_features = ['click_time', 'attributed_time']
train_csv.drop(train_useless_features, axis=1, inplace=True)
test_useless_features = ['click_time', 'click_id']
test_csv.drop(test_useless_features, axis=1, inplace=True)
display(train_csv.head())
display(test_csv.head())
x_data = train_csv
x_train, x_val, y_train, y_val = train_test_split(x_data, y_data, test_size=0.01, random_state=2017)
x_test = test_csv
print(x_data.shape)
print(y_data.shape)
print(x_train.shape)
print(y_train.shape)
print(x_val.shape)
print(y_val.shape)
print(x_test.shape)
```
## Train
```
%%time
import lightgbm as lgb
from sklearn.metrics import roc_auc_score
from sklearn.model_selection import GridSearchCV
# lgb_train = lgb.Dataset(x_train, label=y_train)
# lgb_val = lgb.Dataset(x_val, label=y_val, reference=lgb_train)
# LightGBM parameters
param_grid = {
# 'task': 'train',
# 'num_boost_round': [200],
# 'early_stopping_rounds': [10],
# 'boosting_type': ['gbdt'], # (default="gbdt")
# 'num_leaves': [300], # (default=31)
# 'max_depth': [-1], # (default=-1)
# 'learning_rate': [0.1], # (default=0.1)
# 'n_estimators': [1000, 500], # (default=10)
# 'max_bin': [1000, 255], # (default=255)
# 'subsample_for_bin': [100*10000], # (default=50000)
# 'objective': ['binary'], # (default=None)
# 'min_split_gain': [0.], # (default=0.)
# 'min_child_weight': [1e-3], # (default=1e-3)
# 'min_child_samples': [10], # (default=20)
# 'subsample': [0.7], # (default=1.)
# 'subsample_freq': [1], # (default=1)
# 'colsample_bytree': [0.9], # (default=1.)
# 'reg_alpha': [0.], # (default=0.)
# 'reg_lambda': [0.], # (default=0.)
# 'random_state': [random_num], # (default=None)
# 'n_jobs': [-1], # (default=-1)
# 'silent': [False], # (default=True)
# 'metric': ['auc', 'binary_logloss'],
}
# print('params: ', params)
# train
clf = lgb.LGBMClassifier(
# 'num_boost_round'=200,
# 'early_stopping_rounds'=10,
boosting_type='gbdt', # (default="gbdt")
num_leaves=30000, # (default=31)
max_depth=-1, # (default=-1)
learning_rate=0.1, # (default=0.1)
n_estimators=500, # (default=10)
# 'max_bin'=255, # (default=255)
subsample_for_bin=100*10000, # (default=50000)
objective='binary', # (default=None)
class_weight=None,
min_split_gain=0., # (default=0.)
min_child_weight=1e-3, # (default=1e-3)
min_child_samples=10, # (default=20)
subsample=0.7, # (default=1.)
# 'subsample_freq'=1, # (default=1)
colsample_bytree=0.9, # (default=1.)
reg_alpha=0., # (default=0.)
reg_lambda=0., # (default=0.)
random_state=random_num, # (default=None)
n_jobs=-1, # (default=-1)
silent=False, # (default=True)
# 'metric'=['auc', 'binary_logloss'],
)
# gbm = lgb.train(
# params,
# train_set=lgb_train,
# valid_sets=lgb_val
# )
grid_search = GridSearchCV(estimator=clf, param_grid=param_grid, cv=3, n_jobs=1)
grid_search.fit(x_data, y_data)
y_data_proba = grid_search.predict_proba(x_data)
y_data_pred = (y_data_proba>=0.5).astype(int)
acc_data = accuracy_score(y_data, y_data_pred)
roc_data = roc_auc_score(y_data, y_data_proba)
print('acc_data: %.4f \t roc_data: %.4f' % (acc_data, roc_data))
print(grid_search.best_estimator_)
print(grid_search.grid_scores_)
print('*' * 80)
y_train_proba = gbm.predict(x_train, num_iteration=gbm.best_iteration)
y_train_pred = (y_train_proba>=0.5).astype(int)
acc_train = accuracy_score(y_train, y_train_pred)
roc_train = roc_auc_score(y_train, y_train_proba)
print('acc_train: %.4f \t roc_train: %.4f' % (acc_train, roc_train))
y_val_proba = gbm.predict(x_val, num_iteration=gbm.best_iteration)
y_val_pred = (y_val_proba>=0.5).astype(int)
acc_val = accuracy_score(y_val, y_val_pred)
roc_val = roc_auc_score(y_val, y_val_proba)
print('acc_val: %.4f \t roc_val: %.4f' % (acc_val, roc_val))
```
## Predict
```
run_name_acc = run_name + '_' + str(int(roc_val*10000)).zfill(4)
print(run_name_acc)
y_test_proba = gbm.predict(x_test, num_iteration=gbm.best_iteration)
print(y_test_proba.shape)
print(y_test_proba[:20])
def save_proba(y_train_proba, y_train, y_val_proba, y_val, y_test_proba, click_ids, file_name):
print(click_ids[:5])
if os.path.exists(file_name):
os.remove(file_name)
print('File removed: \t%s' % file_name)
with h5py.File(file_name) as h:
h.create_dataset('y_train_proba', data=y_train_proba)
h.create_dataset('y_train', data=y_train)
h.create_dataset('y_val_proba', data=y_val_proba)
h.create_dataset('y_val', data=y_val)
h.create_dataset('y_test_proba', data=y_test_proba)
h.create_dataset('click_ids', data=click_ids)
print('File saved: \t%s' % file_name)
def load_proba(file_name):
with h5py.File(file_name, 'r') as h:
y_train_proba = np.array(h['y_train_proba'])
y_train = np.array(h['y_train'])
y_val_proba = np.array(h['y_val_proba'])
y_val = np.array(h['y_val'])
y_test_proba = np.array(h['y_test_proba'])
click_ids = np.array(h['click_ids'])
print('File loaded: \t%s' % file_name)
print(click_ids[:5])
return y_train_proba, y_train, y_val_proba, y_val, y_test_proba, click_ids
y_proba_file = os.path.join(model_folder, 'proba_%s.p' % run_name_acc)
save_proba(y_train_proba, y_train, y_val_proba, y_val, y_test_proba, np.array(sample_submission_csv['click_id']), y_proba_file)
y_train_proba, y_train, y_val_proba, y_val, y_test_proba, click_ids = load_proba(y_proba_file)
print(y_train_proba.shape)
print(y_train.shape)
print(y_val_proba.shape)
print(y_val.shape)
print(y_test_proba.shape)
print(len(click_ids))
%%time
submission_csv_file = os.path.join(output_folder, 'pred_%s.csv' % run_name_acc)
print(submission_csv_file)
submission_csv = pd.DataFrame({ 'click_id': click_ids , 'is_attributed': y_test_proba })
submission_csv.to_csv(submission_csv_file, index = False)
print('Time cost: %.2f s' % (time.time() - t0))
print('random_num: ', random_num)
print('date: ', date)
print(run_name_acc)
print('Done!')
```
| github_jupyter |
```
import nltk, re, pickle, os
import pandas as pd
import numpy as np
from textblob import TextBlob
from nltk.tokenize import sent_tokenize, word_tokenize, wordpunct_tokenize, MWETokenizer
from nltk.stem import porter, WordNetLemmatizer
from nltk.corpus import stopwords
from nltk.util import ngrams
from sklearn.feature_extraction.text import CountVectorizer, TfidfVectorizer
from sklearn.decomposition import LatentDirichletAllocation, TruncatedSVD, NMF
from sklearn.neighbors import NearestNeighbors
#import pyLDAvis, pyLDAvis.sklearn
#from IPython.display import display
from sklearn.preprocessing import StandardScaler
import seaborn as sns
%matplotlib inline
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
path = '/Volumes/ext200/Dropbox/metis/p4_fletcher/pick/'
#open metadata and cleaned talks
with open(path + 'ted_all.pkl', 'rb') as picklefile:
ted_all = pickle.load(picklefile)
with open(path + 'cleaned_talks.pkl', 'rb') as picklefile:
cleaned_talks = pickle.load(picklefile)
```
# Vectorize + Topic Modeling
look at tfidf in addition to count vectorizer for topic modeling methods
Count vect: Convert a collection of text documents to a matrix of token counts This implementation produces a sparse representation.
# LDA
```
def topic_mod_lda(data,topics=5,iters=10,ngram_min=1, ngram_max=3, max_df=0.6, max_feats=5000):
""" vectorizer - turn words into numbers for each document(rows)
then use Latent Dirichlet Allocation to get topics"""
vectorizer = CountVectorizer(ngram_range=(ngram_min,ngram_max),
stop_words='english',
max_df = max_df,
max_features=max_feats)
# `fit (train), then transform` to convert text to a bag of words
vect_data = vectorizer.fit_transform(data)
lda = LatentDirichletAllocation(n_components=topics,
max_iter=iters,
random_state=42,
learning_method='online',
n_jobs=-1)
lda_dat = lda.fit_transform(vect_data)
# to display a list of topic words and their scores
def display_topics(model, feature_names, no_top_words):
for ix, topic in enumerate(model.components_):
print("Topic ", ix)
print(" ".join([feature_names[i]
for i in topic.argsort()[:-no_top_words - 1:-1]]))
display_topics(lda, vectorizer.get_feature_names(),20)
return vectorizer, vect_data, lda, lda_dat
```
best yet
cleaned_talks,topics=23,
iters=100,
ngram_min=1,
ngram_max=2,
max_df=0.4,
max_feats=2000)
```
vect_mod, vect_data, lda_mod, lda_data = topic_mod_lda(cleaned_talks,
topics=20,
iters=100,
ngram_min=1,
ngram_max=2,
max_df=0.5,
max_feats=2000)
```
# for each document, assign the topic (column) with the highest score from the LDA
```
topic_ind = np.argmax(lda_data, axis=1)
topic_ind.shape
y=topic_ind
# create text labels for plotting
tsne_labels = pd.DataFrame(y)
# save to csv
tsne_labels.to_csv(path + 'tsne_labels.csv')
topic_names = tsne_labels
topic_names[topic_names==0] = "family"
topic_names[topic_names==1] = "agriculture"
topic_names[topic_names==2] = "space"
topic_names[topic_names==3] = "environment"
topic_names[topic_names==4] = "global economy"
topic_names[topic_names==5] = "writing"
topic_names[topic_names==6] = "sounds"
topic_names[topic_names==7] = "belief, mortality"
topic_names[topic_names==8] = "transportation"
topic_names[topic_names==9] = "gaming"
topic_names[topic_names==10] = "architecture"
topic_names[topic_names==11] = "education"
topic_names[topic_names==12] = "neuroscience"
topic_names[topic_names==13] = "climate, energy"
topic_names[topic_names==14] = "politics"
topic_names[topic_names==15] = "robotics"
topic_names[topic_names==16] = "disease biology"
topic_names[topic_names==17] = "medicine"
topic_names[topic_names==18] = "technology, privacy"
topic_names[topic_names==19] = "war"
topic_names
#save text labels to csv and pkl for plotting
topic_names.to_csv(path + 'topic_names.csv')
with open(path + 'topic_names.pkl', 'wb') as picklefile:
pickle.dump(topic_names, picklefile)
```
Visualize the first 2 components from the topic modeling (LDA).
Not really the best way to look at clusters, but a good place to start.
```
import pyLDAvis, pyLDAvis.sklearn
from IPython.display import display
# Setup to run in Jupyter notebook
pyLDAvis.enable_notebook()
# Create the visualization
vis = pyLDAvis.sklearn.prepare(lda_mod, vect_data, vect_mod)
# Export as a standalone HTML web page
pyLDAvis.save_html(vis, 'lda.html')
# # Let's view it!
display(vis)
```
# tSNE
a way to reduce dimensionality to three components so that we can visualize where/if there are clusters in document-topic space.
```
def plot_tsne(X,y,v1=0,v2=0):
""" pass in the X from pca.transform ,and the corresponding y values and a string to add to the title
plots the first three PCA directions/eigenvectors with target values as the color
___________________________________________________________________"""
fig = plt.figure(1, figsize=(13, 10))
ax = Axes3D(fig, elev=-150, azim=110)
# plot transformed values (the three features that we have decomposed to) , colors correspond to target values
ax.scatter(X[:, 0], X[:, 1], X[:, 2], c=y,
cmap=plt.cm.hot, edgecolor='k', s=50)
ax.set_title("tSNE Ted Topics ", fontsize=16)
ax.set_xlabel("1st ",fontsize=16)
ax.w_xaxis.set_ticklabels([])
ax.set_ylabel("2nd",fontsize=16)
ax.w_yaxis.set_ticklabels([])
ax.set_zlabel("3rd",fontsize=16)
ax.w_zaxis.set_ticklabels([])
ax.view_init(v1,v2)
```
Want to tweak the hyperparameters so that we get the lowest KL divergence
```
from sklearn.manifold import TSNE
# a t-SNE model
# angle value close to 1 means sacrificing accuracy for speed
# pca initializtion usually leads to better results
tsne_model = TSNE(n_components=3, verbose=1, random_state=44, angle=.50,
perplexity=18,early_exaggeration=1,learning_rate=50.0)#, init='pca'
# 20-D -> 3-D
tsne_lda = tsne_model.fit_transform(lda_data)
tsne_data = pd.DataFrame(tsne_lda)
tsne_data.to_csv(path + 'tsne_lda.csv')
plot_tsne(tsne_lda,y, 0, 60)
```
# Below are models that gave sub-par results compared to the LDA
# NMF - must do count vectorizer with this one
```
def topic_mod_nmf(data, topics=5,iters=10,ngram_min=1, ngram_max=3, max_df=0.6, max_feats=5000):
""" vectorizer - turn words into numbers for each document(rows)
then use Latent Dirichlet Allocation to get topics"""
vectorizer = CountVectorizer(ngram_range=(ngram_min,ngram_max),
stop_words='english',
max_df = max_df,
max_features=max_feats)
# call `fit` to build the vocabulary
vectorizer.fit(data)
# finally, call `transform` to convert text to a bag of words
x = vectorizer.transform(data)
mod = NMF(n_components=topics,
max_iter=iters,
random_state=42)
topiced_dat = mod.fit_transform(x)
# to display a list of topic words and their scores
def display_topics(model_, feature_names, no_top_words):
for ix, topic in enumerate(model_.components_):
print("Topic ", ix)
print(" ".join([feature_names[i]
for i in topic.argsort()[:-no_top_words - 1:-1]]))
display_topics(mod, vectorizer.get_feature_names(),20)
return mod, vectorizer, x, topiced_dat
nmf_,vect_mod, vect_data, topic_data = topic_mod_nmf(cleaned_talks,
topics=23,
iters=50,
ngram_min=1,
ngram_max=2,
max_df=0.6,
max_feats=2000)
```
# LSA count vec
```
def topic_mod_lsa(data, topics=5,ngram_min=1, ngram_max=3, max_df=0.6, max_feats=5000):
""" vectorizer - turn words into numbers for each document(rows)
then use Latent Dirichlet Allocation to get topics"""
vectorizer = CountVectorizer(ngram_range=(ngram_min,ngram_max),
stop_words='english',
max_df = max_df,
max_features=max_feats)
# call `fit` to build the vocabulary
vect_data = vectorizer.fit_transform(data)
#stdScale = Normalizer()
#vect_scale = stdScale.fit_transform(vect_data)
mod = TruncatedSVD(n_components=topics,random_state=42)
topiced_dat = mod.fit_transform(vect_data)
# to display a list of topic words and their scores
def display_topics(model_, feature_names, no_top_words):
for ix, topic in enumerate(model_.components_):
print("Topic ", ix)
print(" ".join([feature_names[i]
for i in topic.argsort()[:-no_top_words - 1:-1]]))
display_topics(mod, vectorizer.get_feature_names(),20)
return mod, vectorizer, vect_data
lsa_,vect_mod,lsa_topic_data = topic_mod_lsa(cleaned_talks,
topics=23,
ngram_min=1,
ngram_max=2,
max_df=0.6,
max_feats=2000)
```
# LSA Tfidf
```
def topic_mod_lsa_t(data, topics=5,ngram_min=1, ngram_max=3, max_df=0.6, max_feats=5000):
""" vectorizer - turn words into numbers for each document(rows)
then use Latent Dirichlet Allocation to get topics"""
vectorizer = TfidfVectorizer(ngram_range=(ngram_min,ngram_max),
stop_words='english',
max_df = max_df,
max_features=max_feats)
vect_data = vectorizer.fit_transform(data)
stdScale = Normalizer()
vect_scale = stdScale.fit_transform(vect_data)
mod = TruncatedSVD(n_components=topics,random_state=42)
topiced_dat = mod.fit_transform(vect_scale)
# to display a list of topic words and their scores
def display_topics(model_, feature_names, no_top_words):
for ix, topic in enumerate(model_.components_):
print("Topic ", ix)
print(" ".join([feature_names[i]
for i in topic.argsort()[:-no_top_words - 1:-1]]))
display_topics(mod, vectorizer.get_feature_names(),20)
return mod, vectorizer, vect_scale
lsa_,vect_mod,lsa_topic_data = topic_mod_lsa(cleaned_talks,
topics=23,
ngram_min=1,
ngram_max=2,
max_df=0.6,
max_feats=2000)
```
| github_jupyter |
```
import numpy as np
import cvxpy as cp
import matplotlib.pyplot as plt
%matplotlib inline
```

# Formulation
minimize $\sum{[u_i^T u_i -2u_i^T x_c + x_c^T x_c - r^2]^2}$
using: $z = x_c^T x_c - r^2$
minimize $\sum{(u_i^T u_i -2u_i^T x_c + z)^2} = \lVert A[x_c,z] - b\rVert^2$
It's least squares!
But we need : $r^2 \geq 0 \implies x_c^T x_c - z \geq 0$
Can we assure this always happens at optimal point?
$x_* = (A^T A)^{-1} A^T b$
# Problem data
```
U = [
[-3.8355737e+00 , 5.9061250e+00],
[-3.2269177e+00 , 7.5112709e+00],
[-1.6572955e+00 , 7.4704730e+00],
[-2.8202585e+00 , 7.7378120e+00],
[-1.7831869e+00 , 5.4818448e+00],
[-2.1605783e+00 , 7.7231450e+00],
[-2.0960803e+00 , 7.7072529e+00],
[-1.3866295e+00 , 6.1452654e+00],
[-3.2077849e+00 , 7.6023307e+00],
[-2.0095986e+00 , 7.6382459e+00],
[-2.0965432e+00 , 5.2421510e+00],
[-2.8128775e+00 , 5.1622157e+00],
[-3.6501826e+00 , 7.2585500e+00],
[-2.1638414e+00 , 7.6899057e+00],
[-1.7274710e+00 , 5.4564872e+00],
[-1.5743230e+00 , 7.3510769e+00],
[-1.3761806e+00 , 6.9730981e+00],
[-1.3602495e+00 , 6.9056362e+00],
[-1.5257654e+00 , 5.7518622e+00],
[-1.9231176e+00 , 7.6775030e+00],
[-2.9296195e+00 , 7.7561481e+00],
[-3.2828270e+00 , 5.4188036e+00],
[-2.9078414e+00 , 5.1741322e+00],
[-3.5423007e+00 , 5.5660735e+00],
[-3.1388035e+00 , 7.7008514e+00],
[-1.7957226e+00 , 5.4273243e+00],
[-2.6267585e+00 , 7.7336173e+00],
[-3.6652627e+00 , 7.2686635e+00],
[-3.7394118e+00 , 6.0293335e+00],
[-3.7898021e+00 , 5.9057623e+00],
[-3.6200108e+00 , 5.7754097e+00],
[-3.0386294e+00 , 5.3028798e+00],
[-2.0320023e+00 , 5.2594588e+00],
[-2.9577808e+00 , 5.3040353e+00],
[-2.9146706e+00 , 7.7731243e+00],
[-3.2243786e+00 , 5.4402982e+00],
[-2.1781976e+00 , 7.7681141e+00],
[-2.2545150e+00 , 5.2233652e+00],
[-1.2559218e+00 , 6.2741755e+00],
[-1.8875105e+00 , 5.4133273e+00],
[-3.6122685e+00 , 7.2743342e+00],
[-2.6552417e+00 , 7.7564498e+00],
[-1.4127560e+00 , 6.0732284e+00],
[-3.7475311e+00 , 7.2351834e+00],
[-2.1367633e+00 , 7.6955709e+00],
[-3.9263527e+00 , 6.2241593e+00],
[-2.3118969e+00 , 7.7636052e+00],
[-1.4249518e+00 , 7.1457752e+00],
[-2.0196394e+00 , 5.3154475e+00],
[-1.4021445e+00 , 5.9675466e+00],
]
U = np.array(U)
A = np.concatenate((-2*U , np.ones((U.shape[0],1))), axis = 1)
b = -np.sum(U * U , axis = 1, keepdims = True)
print("A shape:", A.shape)
print("b shape:", b.shape)
```
# Solving
```
x = cp.Variable((A.shape[1], 1))
obj = cp.Minimize(cp.sum_squares(A @ x - b))
prob = cp.Problem(obj)
prob.solve()
assert prob.status == cp.OPTIMAL
x_c = x.value[:-1]
z = x.value[-1]
r2 = x_c.T @ x_c - z
assert r2 >= 0
r = r2**0.5
print('x_c:\n', x_c)
print('r:\n', r)
from matplotlib.patches import Circle
fig, ax = plt.subplots(figsize=(8,8))
ax.scatter(U[:,0], U[:,1], marker = 'x')
ax.scatter(x_c[0],x_c[1], c = 'r')
circle = Circle(x_c, r , facecolor='none', edgecolor='gray', linewidth=1, ls = '--', alpha=0.5)
ax.add_patch(circle)
plt.show()
```
| github_jupyter |
```
import torch
import torch.nn as nn
import torch.nn.functional as F
tod = torch.distributions
import copy
import numpy as np
```
### Entire structure of BNN with Bayes Backprop
```
def isotropic_gauss_loglike(x, mu, sigma, do_sum=True):
cte_term = -(0.5) * np.log(2 * np.pi)
det_sig_term = -torch.log(sigma)
inner = (x - mu) / sigma
dist_term = -(0.5) * (inner ** 2)
if do_sum:
out = (cte_term + det_sig_term + dist_term).sum() # sum over all weights
else:
out = (cte_term + det_sig_term + dist_term)
return out
class laplace_prior(object):
def __init__(self, mu, b):
self.mu = mu
self.b = b
def loglike(self, x, do_sum=True):
if do_sum:
return (-np.log(2 * self.b) - torch.abs(x - self.mu) / self.b).sum()
else:
return (-np.log(2 * self.b) - torch.abs(x - self.mu) / self.b)
"Sample according to parameterized posterior distribution of weights"
"posterior is assumed to be element-wise normal distribuiton for the weights"
def sample_weights(W_mu, b_mu, W_p, b_p):
"Quick method for sampling weights and exporting weights"
".data.new is just to create same typed tensor of certain shape"
"Q1. what is the type of W_mu, and why is tensor type failed for the command .normal_()?"
"""
A1. W_mu is a nn.Paramter which accepts .normal_(), this essentially create N(0,1) sampled values on
each of its entries
A2. W is (entry-wise) normal distributed sample with mean W_mu, (entry-wise) standard deviation std_W.
A3. W_p is used to model std_W, through a softplus function added with 1e-6.
A4. Similarly for b, too. Except that we can use no bias, which is the case when b_mu is None.
A5. A softplus function always keeps output positive, making std reasonable, at the same time, its input
which is the W_p & b_p is not constrained at all, which is why we use it.
"""
eps_W = W_mu.new(W_mu.shape).normal_()
std_W = 1e-6 + F.softplus(W_p, beta=1, threshold=20)
W = W_mu + 1 * std_W * eps_W
if b_mu is not None:
std_b = 1e-6 + F.softplus(b_p, beta=1, threshold=20)
eps_b = b_mu.new(b_mu.shape).normal_()
b = b_mu + 1 * std_b * eps_b
else:
b = None
return W, b
class BayesLinear_Normalq(torch.nn.Module):
"""
Linear Layer where weights are sampled from a fully
factorised Normal with learnable parameters.
The likelihood of the weight samples under the prior
and the approximate posterior are returned with each
forward pass in order to estimate the KL term in
the ELBO.
"""
def __init__(self, n_in, n_out, prior_class):
super(BayesLinear_Normalq, self).__init__()
self.n_in = n_in
self.n_out= n_out
self.prior = prior_class
# Learnable parameters
self.W_mu = nn.Parameter(torch.Tensor(self.n_in, self.n_out).uniform_(-0.1,0.1))
self.W_p = nn.Parameter(torch.Tensor(self.n_in, self.n_out).uniform_(-3,-2))
self.b_mu = nn.Parameter(torch.Tensor(self.n_out).uniform_(-0.1, 0.1))
self.b_p = nn.Parameter(torch.Tensor(self.n_out).uniform_(-3, -2))
self.lpw = 0
self.lqw = 0
"""
X shape (batch_size, n_in)
"""
def forward(self, X, sample=False):
"the self.training is True by default so it doesn't really matter what sample is here"
if not self.training and not sample:
"Expand simply copies and broadcast along first axis to shape (batch_size, n_out)"
output = X @ self.W_mu + self.b_mu.unsqueeze(0)
return output, 0, 0
else:
# the same random sample is used for every element in the minibatch
"Source of randomness, in fact a Gaussian Noise Here"
"Note that the way we generate W & b is in align with the choice of the approximate posterior"
"And it has nothing to do with the prior"
eps_W = self.W_mu.new(self.W_mu.shape).normal_()
eps_b = self.b_mu.new(self.b_mu.shape).normal_()
# sample parameters
std_w = 1e-6 + F.softplus(self.W_p, beta=1, threshold=20)
std_b = 1e-6 + F.softplus(self.b_p, beta=1, threshold=20)
W = self.W_mu + 1 * std_w * eps_W
b = self.b_mu + 1 * std_b * eps_b
output = X @ W + b.unsqueeze(0) # (batch_size, n_output)
"approximate posterior: isotropic_gauss_loglike is in the 'prior.py' file "
lqw = isotropic_gauss_loglike(W, self.W_mu, std_w) + isotropic_gauss_loglike(b, self.b_mu, std_b)
"log-likelihood of the parameters (weights and biases) under the prior"
lpw = self.prior.loglike(W) + self.prior.loglike(b)
return output, lqw, lpw
class bayes_linear_2L(nn.Module):
"""2 hidden layer Bayes By Backprop (VI) Network"""
def __init__(self, input_dim, output_dim, n_hid, prior_instance):
super(bayes_linear_2L, self).__init__()
# prior_instance = isotropic_gauss_prior(mu=0, sigma=0.1)
# prior_instance = spike_slab_2GMM(mu1=0, mu2=0, sigma1=0.135, sigma2=0.001, pi=0.5)
# prior_instance = isotropic_gauss_prior(mu=0, sigma=0.1)
self.prior_instance = prior_instance
self.input_dim = input_dim
self.output_dim = output_dim
"n_hid: input dim and output dim for the hidden layers (2nd layer)"
self.bfc1 = BayesLinear_Normalq(input_dim, n_hid, self.prior_instance)
self.bfc2 = BayesLinear_Normalq(n_hid, n_hid, self.prior_instance)
self.bfc3 = BayesLinear_Normalq(n_hid, output_dim, self.prior_instance)
# choose your non linearity
# self.act = nn.Tanh()
# self.act = nn.Sigmoid()
self.act = nn.ReLU(inplace=True)
# self.act = nn.ELU(inplace=True)
# self.act = nn.SELU(inplace=True)
def forward(self, x, sample=False):
"note that sample is just a bool type with True or False value"
tlqw = 0
tlpw = 0
"Q2. what does view do? The answer is with the type of input x"
"A2: this is the key which reshaped input into (batch_size, input_dim)"
x = x.view(-1, self.input_dim) # view(batch_size, input_dim)
# -----------------
"Calling the Forward Pass of first layer, essentially"
x, lqw, lpw = self.bfc1(x, sample)
tlqw = tlqw + lqw
tlpw = tlpw + lpw
# -----------------
x = self.act(x)
# -----------------
x, lqw, lpw = self.bfc2(x, sample)
tlqw = tlqw + lqw
tlpw = tlpw + lpw
# -----------------
x = self.act(x)
# -----------------
y, lqw, lpw = self.bfc3(x, sample)
tlqw = tlqw + lqw
tlpw = tlpw + lpw
return y, tlqw, tlpw
def sample_predict(self, x, Nsamples):
"""
Not sure what it means: Used for estimating the data's likelihood by approximately marginalising the weights with MC
Take a number (Nsamples) of samples of weight sets according to the posterior distribution assumption, compute
the output or predictions of the network based on those values of weights and biases.
"""
# Just copies type from x, initializes new vector
predictions = x.new(Nsamples, x.shape[0], self.output_dim)
tlqw_vec = np.zeros(Nsamples)
tlpw_vec = np.zeros(Nsamples)
for i in range(Nsamples):
y, tlqw, tlpw = self.forward(x, sample=True)
predictions[i] = y
tlqw_vec[i] = tlqw
tlpw_vec[i] = tlpw
return predictions, tlqw_vec, tlpw_vec
class BaseNet(object):
def __init__(self):
cprint('c', '\nNet:')
def get_nb_parameters(self):
return sum(p.numel() for p in self.model.parameters())
def set_mode_train(self, train=True):
if train:
"essentially call the nn.Module.train() which sets the training mode for the model"
self.model.train()
else:
"set the nn.Module to eval mode"
self.model.eval()
def update_lr(self, epoch, gamma=0.99):
self.epoch += 1
if self.schedule is not None:
if len(self.schedule) == 0 or epoch in self.schedule:
self.lr *= gamma
print('learning rate: %f (%d)\n' % self.lr, epoch)
for param_group in self.optimizer.param_groups:
param_group['lr'] = self.lr
def save(self, filename):
cprint('c', 'Writting %s\n' % filename)
torch.save({
'epoch': self.epoch,
'lr': self.lr,
'model': self.model,
'optimizer': self.optimizer}, filename)
def load(self, filename):
cprint('c', 'Reading %s\n' % filename)
state_dict = torch.load(filename)
self.epoch = state_dict['epoch']
self.lr = state_dict['lr']
self.model = state_dict['model']
self.optimizer = state_dict['optimizer']
print(' restoring epoch: %d, lr: %f' % (self.epoch, self.lr))
return self.epoch
class BBP_Bayes_Net(BaseNet):
"""
Full network wrapper for Bayes By Backprop nets with methods for training,
prediction and weight prunning
"""
eps = 1e-6
def __init__(self, lr=1e-3, channels_in=5, side_in=1, cuda=True, classes=5, batch_size=5, Nbatches=1,
nhid=10, prior_instance=laplace_prior(mu=0, b=0.1)):
super(BBP_Bayes_Net, self).__init__()
cprint('y', ' Creating Net!! ')
self.lr = lr
self.schedule = None # [] #[50,200,400,600]
self.cuda = cuda
self.channels_in = channels_in
self.classes = classes
"entire number of data is the product of batch_size and Nbatches"
"more like size of each minibatch"
self.batch_size = batch_size
"more like no. of minibatches"
self.Nbatches = Nbatches
self.prior_instance = prior_instance
self.nhid = nhid
self.side_in = side_in
self.create_net()
self.create_opt()
self.epoch = 0
self.test = False
def create_net(self):
torch.manual_seed(42)
if self.cuda:
torch.cuda.manual_seed(42)
"Q3: why is the input dim equals the product and not just channels_in ?"
"A3: We input image data, which has side_length * side_length * no. layers(RGB) number of data points"
self.model = bayes_linear_2L(input_dim=self.channels_in * self.side_in * self.side_in,
output_dim=self.classes, n_hid=self.nhid, prior_instance=self.prior_instance)
if self.cuda:
self.model.cuda()
# cudnn.benchmark = True
print(' Total params: %.2fM' % (self.get_nb_parameters() / 1000000.0))
def create_opt(self):
# self.optimizer = torch.optim.Adam(self.model.parameters(), lr=self.lr, betas=(0.9, 0.999), eps=1e-08,
# weight_decay=0)
self.optimizer = torch.optim.SGD(self.model.parameters(), lr=self.lr, momentum=0)
# self.optimizer = torch.optim.SGD(self.model.parameters(), lr=self.lr, momentum=0.9)
# self.sched = torch.optim.lr_scheduler.StepLR(self.optimizer, step_size=1, gamma=10, last_epoch=-1)
def fit(self, x, y, samples=1):
x, y = to_variable(var=(x, y.long()), cuda=self.cuda)
self.optimizer.zero_grad()
if samples == 1:
"model used here is STOCHASTIC but with only 1 samples used"
out, tlqw, tlpw = self.model(x)
"""
1. mean log probability of data given weights: mlpdw
2. out shape: (batch_size, out_dim), y shape: (batch_size)
3. This is a classification task, y takes value {1,2,...,out_dim}, out_dim is actually
the number of classes also. For the F.cross_entropy function, it first convert 'out',
whose value can be any real number, into a valid discrete distribution mass function,
then compute - log(p_{out}(y)), then summed over all the batches, which we can use
to construct MC estimate of Cross Entropy (Likelihood Cost) by taking average over batch
size.
"""
mlpdw = F.cross_entropy(out, y, reduction='sum')
"expected KL divergence"
"Note that this term get scaled furthur by Nbatches"
Edkl = (tlqw - tlpw) / self.Nbatches
elif samples > 1:
mlpdw_cum = 0
Edkl_cum = 0
for i in range(samples):
out, tlqw, tlpw = self.model(x, sample=True)
mlpdw_i = F.cross_entropy(out, y, reduction='sum')
Edkl_i = (tlqw - tlpw) / self.Nbatches
mlpdw_cum = mlpdw_cum + mlpdw_i
Edkl_cum = Edkl_cum + Edkl_i
mlpdw = mlpdw_cum / samples
Edkl = Edkl_cum / samples
"loss function we wish to minimize, negative ELBO"
loss = Edkl + mlpdw
loss.backward()
self.optimizer.step()
# out: (batch_size, out_channels, out_caps_dims)
pred = out.data.max(dim=1, keepdim=False)[1] # get the index of the max log-probability
err = pred.ne(y.data).sum()
return Edkl.data, mlpdw.data, err
def eval(self, x, y, train=False):
x, y = to_variable(var=(x, y.long()), cuda=self.cuda)
out, _, _ = self.model(x)
loss = F.cross_entropy(out, y, reduction='sum')
probs = F.softmax(out, dim=1).data.cpu()
pred = out.data.max(dim=1, keepdim=False)[1] # get the index of the max log-probability
err = pred.ne(y.data).sum()
return loss.data, err, probs
def sample_eval(self, x, y, Nsamples, logits=True, train=False):
"""Prediction, only returining result with weights marginalised"""
x, y = to_variable(var=(x, y.long()), cuda=self.cuda)
out, _, _ = self.model.sample_predict(x, Nsamples)
"""
1. Cross-entropy in torch is basically a softmax (to get estimate prob) +
negative log-likelihood loss
2. NLLLoss in torch assumes the input to be already log-scale
3. Considering above two points, the main difference for the if/else below is that
we take mean over samples and softmanx to get prob or we take probs for each sample
and then average to get mean probs
"""
if logits:
mean_out = out.mean(dim=0, keepdim=False)
loss = F.cross_entropy(mean_out, y, reduction='sum')
probs = F.softmax(mean_out, dim=1).data.cpu()
else:
mean_out = F.softmax(out, dim=2).mean(dim=0, keepdim=False)
probs = mean_out.data.cpu()
log_mean_probs_out = torch.log(mean_out)
loss = F.nll_loss(log_mean_probs_out, y, reduction='sum')
pred = mean_out.data.max(dim=1, keepdim=False)[1] # get the index of the max log-probability
err = pred.ne(y.data).sum()
return loss.data, err, probs
def all_sample_eval(self, x, y, Nsamples):
"""Returns predictions for each MC sample"""
x, y = to_variable(var=(x, y.long()), cuda=self.cuda)
out, _, _ = self.model.sample_predict(x, Nsamples)
prob_out = F.softmax(out, dim=2)
prob_out = prob_out.data
return prob_out
"samples weights, flatten and record it but not the bias"
def get_weight_samples(self, Nsamples=10):
state_dict = self.model.state_dict()
weight_vec = []
Nsamples=10
for i in range(Nsamples):
for key in state_dict.keys():
"each key for loop is a 'str' class object"
"the split('.') split up the str object according to position of '.' "
"and divide into list, weight_name is 'W_mu', 'W_p', 'b_mu' and 'b_p' "
weight_dict = {}
weight_name = key.split('.')[1]
weight_dict[weight_name] = state_dict[key].cpu().data
if weight_name == 'b_p':
W, b = sample_weights(W_mu=W_mu, b_mu=b_mu, W_p=W_p, b_p=b_p)
for weight in W.cpu().view(-1):
weight_vec.append(weight)
return np.array(weight_vec)
"""
Record here the value of absolute value of mean divided by std for weights (elementwise)
posterior distribuiton, probably useful for reparameterization
1. With thresh, then present element-wise whether threshold is exceeded
2. Without thresh, report the element-wise _SNR value
"""
def get_weight_SNR(self, thresh=None):
state_dict = self.model.state_dict()
weight_SNR_vec = []
if thresh is not None:
mask_dict = {}
weight_dict = {}
for key in state_dict.keys():
weight_name = key.split('.')[1]
layer_name = key.split('.')[0]
weight_dict[weight_name] = state_dict[key].data
if weight_name == 'b_p':
W_mu, W_p, b_mu, b_p = weight_dict.values()
"compute elementwise posterior std"
sig_W = 1e-6 + F.softplus(W_p, beta=1, threshold=20)
sig_b = 1e-6 + F.softplus(b_p, beta=1, threshold=20)
"element-wise posterior absolute mean divided by std"
W_snr = (torch.abs(W_mu) / sig_W)
b_snr = (torch.abs(b_mu) / sig_b)
"if thresh exist, return element-wise True/False: whether _snr > thresh"
if thresh is not None:
mask_dict[layer_name + '.W'] = (W_snr > thresh)
mask_dict[layer_name + '.b'] = (b_snr > thresh)
"if no thresh, record the _snr value"
if thresh is None:
for weight_SNR in W_snr.cpu().view(-1):
weight_SNR_vec.append(weight_SNR)
for weight_SNR in b_snr.cpu().view(-1):
weight_SNR_vec.append(weight_SNR)
if thresh is not None:
return mask_dict
else:
return np.array(weight_SNR_vec)
"""
Sample independetly Nsamples of weights and compute element-wise KL divergence
between approximate posterior and the prior distributions
1. With thresh, then present element-wise whether threshold is exceeded
2. Without thresh, report the element-wise KL divergence value
"""
def get_weight_KLD(self, Nsamples=20, thresh=None):
state_dict = self.model.state_dict()
weight_KLD_vec = []
if thresh is not None:
mask_dict = {}
weight_dict = {}
for key in state_dict.keys():
weight_name = key.split('.')[1]
layer_name = key.split('.')[0]
weight_dict[weight_name] = state_dict[key].data
if weight_name == 'b_p':
W_mu, W_p, b_mu, b_p = weight_dict.values()
"compute elementwise posterior std"
std_W = 1e-6 + F.softplus(W_p, beta=1, threshold=20)
std_b = 1e-6 + F.softplus(b_p, beta=1, threshold=20)
KL_W = W_mu.new(W_mu.shape).zero_()
KL_b = b_mu.new(b_mu.shape).zero_()
for i in range(Nsamples):
W, b = sample_weights(W_mu=W_mu, b_mu=b_mu, W_p=W_p, b_p=b_p)
# Note that this will currently not work with slab and spike prior
"posterior element-wise log like minus prior element-wise log like"
KL_W += isotropic_gauss_loglike(W, W_mu, std_W,
do_sum=False) - self.model.prior_instance.loglike(W,
do_sum=False)
"posterior element-wise log like minus prior element-wise log like"
KL_b += isotropic_gauss_loglike(b, b_mu, std_b,
do_sum=False) - self.model.prior_instance.loglike(b,
do_sum=False)
"average over number of samples"
KL_W /= Nsamples
KL_b /= Nsamples
"thresh here is for the KL divergence value specifically"
if thresh is not None:
mask_dict[layer_name + '.W'] = KL_W > thresh
mask_dict[layer_name + '.b'] = KL_b > thresh
else:
for weight_KLD in KL_W.cpu().view(-1):
weight_KLD_vec.append(weight_KLD)
for weight_KLD in KL_b.cpu().view(-1):
weight_KLD_vec.append(weight_KLD)
if thresh is not None:
return mask_dict
else:
return np.array(weight_KLD_vec)
"""
Masking the model's parameter, if threshold is not exceeded, reset the mean/std for
posterior distribuiton for the weights to zero/~0.01.
"""
def mask_model(self, Nsamples=0, thresh=0):
'''
Nsamples is used to select SNR (0) or KLD (>0) based masking
'''
original_state_dict = copy.deepcopy(self.model.state_dict())
"Note that = means that changing value of state_dict will also change values for RHS"
state_dict = self.model.state_dict()
if Nsamples == 0:
mask_dict = self.get_weight_SNR(thresh=thresh)
else:
mask_dict = self.get_weight_KLD(Nsamples=Nsamples, thresh=thresh)
n_unmasked = 0
previous_layer_name = ''
for key in state_dict.keys():
layer_name = key.split('.')[0]
if layer_name != previous_layer_name:
previous_layer_name = layer_name
"if element value below threshold, reset to mean zero and small std (~0.01)"
"this procedure is called masking, put a maks on these values"
state_dict[layer_name + '.W_mu'][~mask_dict[layer_name + '.W']] = 0
state_dict[layer_name + '.W_p'][~mask_dict[layer_name + '.W']] = -1000
state_dict[layer_name + '.b_mu'][~mask_dict[layer_name + '.b']] = 0
state_dict[layer_name + '.b_p'][~mask_dict[layer_name + '.b']] = -1000
"number of un-masked weight values"
n_unmasked += mask_dict[layer_name + '.W'].sum()
n_unmasked += mask_dict[layer_name + '.b'].sum()
return original_state_dict, n_unmasked
```
### Training on MNIST dataset
```
import time
import torch.utils.data
from torchvision import transforms, datasets
import matplotlib
models_dir = 'test/models_weight_uncertainty_MC_MNIST_gaussian'
results_dir = 'test/results_weight_uncertainty_MC_MNIST_gaussian'
"""
create folder(s) in current location named with element in paths,
after converting to list object
"""
def mkdir(paths):
"if not already a list, make it a list"
if not isinstance(paths, (list, tuple)):
paths = [paths]
""
for path in paths:
"if not a directory, make it a directory"
if not os.path.isdir(path):
"creates a folder named path in the current location"
os.makedirs(path)
mkdir(models_dir)
mkdir(results_dir)
# train config
NTrainPointsMNIST = 60000
batch_size = 100
nb_epochs = 160
log_interval = 1
savemodel_its = [20, 50, 80, 120]
save_dicts = []
```
The MNIST dataset is comprised of 70,000 handwritten numeric digit images and their respective labels.
There are 60,000 training images and 10,000 test images, all of which are 28 pixels by 28 pixels.
```
cprint('c', '\nData:')
# load data
"FIX for data loading issue: 503"
"seems to be a server issue which is fixed by using another amazon server link"
new_mirror = 'https://ossci-datasets.s3.amazonaws.com/mnist'
datasets.MNIST.resources = [
('/'.join([new_mirror, url.split('/')[-1]]), md5)
for url, md5 in datasets.MNIST.resources
]
# data augmentation
transform_train = transforms.Compose([
transforms.ToTensor(),
transforms.Normalize(mean=(0.1307,), std=(0.3081,))
])
transform_test = transforms.Compose([
transforms.ToTensor(),
transforms.Normalize(mean=(0.1307,), std=(0.3081,))
])
trainset = datasets.MNIST("./data", train=True, download=True, transform=transform_train)
valset = datasets.MNIST(root='./data', train=False, download=True, transform=transform_test)
use_cuda = torch.cuda.is_available()
if use_cuda:
trainloader = torch.utils.data.DataLoader(trainset, batch_size=batch_size, shuffle=True, pin_memory=True, num_workers=3)
valloader = torch.utils.data.DataLoader(valset, batch_size=batch_size, shuffle=False, pin_memory=True, num_workers=3)
else:
trainloader = torch.utils.data.DataLoader(trainset, batch_size=batch_size, shuffle=True, pin_memory=False,
num_workers=3)
valloader = torch.utils.data.DataLoader(valset, batch_size=batch_size, shuffle=False, pin_memory=False,
num_workers=3)
```
### Create BNN Network
```
# net dims
cprint('c', '\nNetwork:')
lr = 1e-3
nsamples = 3
########################################################################################
net = BBP_Bayes_Net(lr=lr, channels_in=1, side_in=28, cuda=use_cuda, classes=10, batch_size=batch_size,
Nbatches=(NTrainPointsMNIST/batch_size))
epoch = 0
```
## Training
```
# train
cprint('c', '\nTrain:')
print(' init cost variables:')
kl_cost_train = np.zeros(nb_epochs)
pred_cost_train = np.zeros(nb_epochs)
err_train = np.zeros(nb_epochs)
cost_dev = np.zeros(nb_epochs)
err_dev = np.zeros(nb_epochs)
# best_cost = np.inf
best_err = np.inf
nb_its_dev = 1
tic0 = time.time()
for i in range(epoch, nb_epochs):
"the object class BaseNet grants the sub-object class BBP_Bayes_Net a .set_mode_train function"
net.set_mode_train(True)
tic = time.time()
nb_samples = 0
for x, y in trainloader:
".fit trains the model with one step of optimization"
"x shape: (batch_size, channels_in, side_in, side_in)"
"Q4: why is shape of x not (batch_size, input_dim) and still work?"
"A4: See A2, bayes_linear_2L explicitly reshapes the input"
"y shape: (batch_size)"
"nsamples is MC samples for weight used to evaluate loss/err, here set to 3"
cost_dkl, cost_pred, err = net.fit(x, y, samples=nsamples)
err_train[i] += err
kl_cost_train[i] += cost_dkl
pred_cost_train[i] += cost_pred
"batch_size: len(x) / number of training points accumulate: nb_samples"
nb_samples += len(x)
kl_cost_train[i] /= nb_samples
pred_cost_train[i] /= nb_samples
err_train[i] /= nb_samples
toc = time.time()
net.epoch = i
# ---- print
print("it %d/%d, Jtr_KL = %f, Jtr_pred = %f, err = %f, " % (i, nb_epochs, kl_cost_train[i], pred_cost_train[i], err_train[i]), end="")
cprint('r', ' time: %f seconds\n' % (toc - tic))
# Save state dict
if i in savemodel_its:
save_dicts.append(copy.deepcopy(net.model.state_dict()))
# ---- dev
"compute the test error among validation data set, save the best model / final model"
if i % nb_its_dev == 0:
net.set_mode_train(False)
nb_samples = 0
for j, (x, y) in enumerate(valloader):
cost, err, probs = net.eval(x, y)
cost_dev[i] += cost
err_dev[i] += err
nb_samples += len(x)
cost_dev[i] /= nb_samples
err_dev[i] /= nb_samples
cprint('g', ' Jdev = %f, err = %f\n' % (cost_dev[i], err_dev[i]))
if err_dev[i] < best_err:
best_err = err_dev[i]
cprint('b', 'best test error')
net.save(models_dir+'/theta_best.dat')
toc0 = time.time()
runtime_per_it = (toc0 - tic0) / float(nb_epochs)
cprint('r', ' average time: %f seconds\n' % runtime_per_it)
net.save(models_dir+'/theta_last.dat')
```
## Result
```
## ---------------------------------------------------------------------------------------------------------------------
# results
cprint('c', '\nRESULTS:')
nb_parameters = net.get_nb_parameters()
best_cost_dev = np.min(cost_dev)
best_cost_train = np.min(pred_cost_train)
err_dev_min = err_dev[::nb_its_dev].min()
print(' cost_dev: %f (cost_train %f)' % (best_cost_dev, best_cost_train))
print(' err_dev: %f' % (err_dev_min))
print(' nb_parameters: %d (%s)' % (nb_parameters, humansize(nb_parameters)))
print(' time_per_it: %fs\n' % (runtime_per_it))
## Save results for plots
# np.save('results/test_predictions.npy', test_predictions)
np.save(results_dir + '/cost_train.npy', kl_cost_train)
np.save(results_dir + '/cost_train.npy', pred_cost_train)
np.save(results_dir + '/cost_dev.npy', cost_dev)
np.save(results_dir + '/err_train.npy', err_train)
np.save(results_dir + '/err_dev.npy', err_dev)
## ---------------------------------------------------------------------------------------------------------------------
# fig cost vs its
textsize = 15
marker=5
plt.figure(dpi=100)
fig, ax1 = plt.subplots()
ax1.plot(pred_cost_train, 'r--')
ax1.plot(range(0, nb_epochs, nb_its_dev), cost_dev[::nb_its_dev], 'b-')
ax1.set_ylabel('Cross Entropy')
plt.xlabel('epoch')
plt.grid(b=True, which='major', color='k', linestyle='-')
plt.grid(b=True, which='minor', color='k', linestyle='--')
lgd = plt.legend(['test error', 'train error'], markerscale=marker, prop={'size': textsize, 'weight': 'normal'})
ax = plt.gca()
plt.title('classification costs')
for item in ([ax.title, ax.xaxis.label, ax.yaxis.label] +
ax.get_xticklabels() + ax.get_yticklabels()):
item.set_fontsize(textsize)
item.set_weight('normal')
plt.savefig(results_dir + '/cost.png', bbox_extra_artists=(lgd,), bbox_inches='tight')
plt.figure()
fig, ax1 = plt.subplots()
ax1.plot(kl_cost_train, 'r')
ax1.set_ylabel('nats?')
plt.xlabel('epoch')
plt.grid(b=True, which='major', color='k', linestyle='-')
plt.grid(b=True, which='minor', color='k', linestyle='--')
ax = plt.gca()
plt.title('DKL (per sample)')
for item in ([ax.title, ax.xaxis.label, ax.yaxis.label] +
ax.get_xticklabels() + ax.get_yticklabels()):
item.set_fontsize(textsize)
item.set_weight('normal')
plt.figure(dpi=100)
fig2, ax2 = plt.subplots()
ax2.set_ylabel('% error')
ax2.semilogy(range(0, nb_epochs, nb_its_dev), 100 * err_dev[::nb_its_dev], 'b-')
ax2.semilogy(100 * err_train, 'r--')
plt.xlabel('epoch')
plt.grid(b=True, which='major', color='k', linestyle='-')
plt.grid(b=True, which='minor', color='k', linestyle='--')
ax2.get_yaxis().set_minor_formatter(matplotlib.ticker.ScalarFormatter())
ax2.get_yaxis().set_major_formatter(matplotlib.ticker.ScalarFormatter())
lgd = plt.legend(['test error', 'train error'], markerscale=marker, prop={'size': textsize, 'weight': 'normal'})
ax = plt.gca()
for item in ([ax.title, ax.xaxis.label, ax.yaxis.label] +
ax.get_xticklabels() + ax.get_yticklabels()):
item.set_fontsize(textsize)
item.set_weight('normal')
plt.savefig(results_dir + '/err.png', bbox_extra_artists=(lgd,), box_inches='tight')
```
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
import warnings
warnings.filterwarnings('ignore')
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.linear_model import LinearRegression, Lasso, LogisticRegression, ElasticNet
from sklearn.ensemble import GradientBoostingRegressor, GradientBoostingClassifier
from sklearn.preprocessing import PolynomialFeatures, StandardScaler, RobustScaler, QuantileTransformer
from sklearn.pipeline import Pipeline
import scipy.special
from dml_iv import DMLIV
from dr_iv import DRIV, ProjectedDRIV
from dml_ate_iv import DMLATEIV
```
# Synthetic Data Generation - TripAdvisor
- Used for generating data that looks similar in structure (value distributions of private features are not representative of true data) to the data used for the intent-to-treat A/B test at TripAdvisor.
```
import numpy as np
import pandas as pd
import locale
n = 10000 # Generated dataset size
np.random.seed(123)
X_colnames = {
'days_visited_exp_pre': 'day_count_pre', # How many days did they visit TripAdvisor attractions pages in the pre-period
'days_visited_free_pre': 'day_count_pre', # How many days did they visit TripAdvisor through free channels (e.g. domain direct) in the pre-period
'days_visited_fs_pre': 'day_count_pre', # How many days did they visit TripAdvisor fs pages in the pre-period
'days_visited_hs_pre': 'day_count_pre', # How many days did they visit TripAdvisor hotels pages in the pre-period
'days_visited_rs_pre': 'day_count_pre', # How many days did they visit TripAdvisor restaurant pages in the pre-period
'days_visited_vrs_pre': 'day_count_pre', # How many days did they visit TripAdvisor vrs pages in the pre-period
'is_existing_member': 'binary', #Binary indicator of whether they are existing member
'locale_en_US': 'binary', # User's locale
'os_type': 'os', # User's operating system
'revenue_pre': 'revenue', # Revenue in the pre-period
}
treat_colnames = {
'treatment': 'binary', # Did they receive the easier sign-up process in the experiment? [This is the instrument]
'is_member': 'is_member' # Did they become a member during the experiment period (through any means)? [This is the treatment of interest]
}
outcome_colnames = {
'days_visited': 'days_visited', # How many days did they visit TripAdvisor in the experimental period
}
def gen_data(data_type, n):
gen_func = {'day_count_pre': lambda: np.random.randint(0, 29 , n), # Pre-experiment period was 28 days
'day_count_post': lambda: np.random.randint(0, 15, n), # Experiment ran for 14 days
'os': lambda: np.random.choice(['osx', 'windows', 'linux'], n),
'locale': lambda: np.random.choice(list(locale.locale_alias.keys()), n),
'count': lambda: np.random.lognormal(1, 1, n).astype('int'),
'binary': lambda: np.random.binomial(1, .5, size=(n,)),
##'days_visited': lambda:
'revenue': lambda: np.round(np.random.lognormal(0, 3, n), 2)
}
return gen_func[data_type]() if data_type else None
X_data = {colname: gen_data(datatype, n) for colname, datatype in X_colnames.items()}
##treat_data = {colname: gen_data(datatype, N) for colname, datatype in treat_colnames.items()}
##outcome_data = {colname: gen_data(datatype, N) for colname, datatype in outcome_colnames.items()}
X_data=pd.DataFrame({**X_data})
# Turn strings into categories for numeric mapping
X_data['os_type'] = X_data.os_type.astype('category').cat.codes
print(X_data.columns.values)
X_pre=X_data.values.astype('float')
def dgp_binary(X,n,true_fn):
##X = np.random.uniform(-1, 1, size=(n, d))
Z = np.random.binomial(1, .5, size=(n,))
nu = np.random.uniform(-5, 5, size=(n,))
coef_Z = 0.8
plt.title("Pr[T=1 | Z=1, X] vs Pr[T=1 | Z=0, X]")
plt.scatter(X[:, 0], coef_Z*scipy.special.expit(0.4*X[:, 0] + nu))
plt.scatter(X[:, 0], .1*np.ones(X.shape[0]))
plt.show()
C = np.random.binomial(1, coef_Z*scipy.special.expit(0.4*X[:, 0] + nu)) # Compliers when recomended
C0 = np.random.binomial(1, .006*np.ones(X.shape[0])) # Non-compliers when not recommended
T = C * Z + C0 * (1 - Z)
y = true_fn(X) * T + 2*nu + 5*(X[:, 0]>0) + 0.1*np.random.uniform(0, 1, size=(n,))
return y, T, Z
true_fn = lambda X: (.8+.5*(5*(X[:,0]>5) + 10*(X[:,0]>15) + 5*(X[:, 0]>20)) - 3*X[:, 6])
y, T, Z = dgp_binary(X_pre, n, true_fn)
X = QuantileTransformer(subsample=100000).fit_transform(X_pre)
#### y menas number of days visit TripAdvisor in the experimental period, should be in range [0,14],
##should be strong right skewed
plt.hist(y)
plt.show()
print("True ATE: {:.3f}".format(np.mean(true_fn(X_pre))))
plt.figure(figsize=(10, 2))
plt.subplot(1, 2, 1)
plt.hist(T[Z==0])
plt.title("T[Z=0]")
plt.subplot(1, 2, 2)
plt.hist(T[Z==1])
plt.title("T[Z=1]")
plt.show()
print("New members: in treatment = {:f}, in control = {:f}".format(T[Z == 1].sum()/Z.sum(), T[Z == 0].sum()/(1-Z).sum()))
print("Z treatment proportion: {:.5f}".format(np.mean(Z)))
```
### Defining some generic regressors and classifiers
```
from dml_ate_iv import DMLATEIV
from utilities import RegWrapper, SeparateModel
from sklearn.model_selection import GridSearchCV, StratifiedKFold, train_test_split
from sklearn.ensemble import RandomForestClassifier, RandomForestRegressor
from sklearn.dummy import DummyClassifier
from sklearn.linear_model import LinearRegression, LogisticRegression, LogisticRegressionCV, LassoCV
from sklearn import metrics
from xgboost import XGBClassifier, XGBRegressor
from xgb_utilities import XGBWrapper
# Define a generic non-parametric regressor
#model = lambda: GradientBoostingRegressor(n_estimators=20, max_depth=3, min_samples_leaf=20,
# n_iter_no_change=5, min_impurity_decrease=.001, tol=0.001)
model = lambda: XGBWrapper(XGBRegressor(gamma=0.001, n_estimators=100, min_child_weight=50, n_jobs=10),
early_stopping_rounds=5, eval_metric='rmse', binary=False)
# model = lambda: RandomForestRegressor(n_estimators=100)
# model = lambda: Lasso(alpha=0.0001) #CV(cv=5)
# model = lambda: GradientBoostingRegressor(n_estimators=60)
# model = lambda: LinearRegression(n_jobs=-1)
# model = lambda: LassoCV(cv=5, n_jobs=-1)
# Define a generic non-parametric classifier. We have to wrap it with the RegWrapper, because
# we want to use predict_proba and not predict. The RegWrapper calls predict_proba of the
# underlying model whenever predict is called.
#model_clf = lambda: RegWrapper(GradientBoostingClassifier(n_estimators=20, max_depth=3, min_samples_leaf=20,
# n_iter_no_change=5, min_impurity_decrease=.001, tol=0.001))
model_clf = lambda: RegWrapper(XGBWrapper(XGBClassifier(gamma=0.001, n_estimators=100, min_child_weight=50, n_jobs=10),
early_stopping_rounds=5, eval_metric='logloss', binary=True))
# model_clf = lambda: RandomForestClassifier(n_estimators=100)
# model_clf = lambda: RegWrapper(GradientBoostingClassifier(n_estimators=60))
# model_clf = lambda: RegWrapper(LogisticRegression(C=10, penalty='l1', solver='liblinear'))
# model_clf = lambda: RegWrapper(LogisticRegressionCV(n_jobs=-1, cv=3, scoring='neg_log_loss'))
model_clf_dummy = lambda: RegWrapper(DummyClassifier(strategy='prior'))
# We need to specify models to be used for each of these residualizations
model_Y_X = lambda: model() # model for E[Y | X]
model_T_X = lambda: model_clf() # model for E[T | X]. We use a classifier since T is binary
model_Z_X = lambda: model_clf_dummy() # model for E[Z | X]. We use a classifier since Z is binary
# For DMLIV we also need a model for E[T | X, Z]. We use a classifier since T is binary
# Because Z is also binary, we could have also done a more complex model_T_XZ, where we split
# the data based on Z=1 and Z=0 and fit a separate sub-model for each case.
model_T_XZ = lambda: SeparateModel(model_clf(), model_clf())
def plot_separate(X, X_pre, cate):
ones = X[:, 6]>.5
lower = []
upper = []
unique_X = np.unique(X[ones, 0])
for t in unique_X:
upper.append(np.percentile(cate[ones & (X[:, 0]==t)], 99))
lower.append(np.percentile(cate[ones & (X[:, 0]==t)], 1))
p = plt.fill_between(unique_X, lower, upper, label='est1', alpha=.5)
plt.plot(unique_X, lower, color=p.get_facecolor()[0], alpha=.5)
plt.plot(unique_X, upper, color=p.get_facecolor()[0], alpha=.5)
plt.scatter(X[ones, 0], true_fn(X_pre[ones]), label='true1', alpha=.8)
ones = X[:, 6]<.5
lower = []
upper = []
unique_X = np.unique(X[ones, 0])
for t in unique_X:
upper.append(np.percentile(cate[ones & (X[:, 0]==t)], 99))
lower.append(np.percentile(cate[ones & (X[:, 0]==t)], 1))
p = plt.fill_between(unique_X, lower, upper, label='est0', alpha=.5)
plt.plot(unique_X, lower, color=p.get_facecolor()[0], alpha=.5)
plt.plot(unique_X, upper, color=p.get_facecolor()[0], alpha=.5)
plt.scatter(X[ones, 0], true_fn(X_pre[ones]), label='true0', alpha=.8)
plt.legend()
```
# DMLATEIV
```
dmlateiv_obj = DMLATEIV(model_Y_X(), model_T_X(), model_Z_X(),
n_splits=10, # n_splits determines the number of splits to be used for cross-fitting.
binary_instrument=True, # a flag whether to stratify cross-fitting by instrument
binary_treatment=True # a flag whether to stratify cross-fitting by treatment
)
dmlateiv_obj.fit(y, T, X, Z)
ta_effect = dmlateiv_obj.effect()
ta_effect_conf = dmlateiv_obj.normal_effect_interval(lower=2.5, upper=97.5)
print("True ATE: {:.3f}".format(np.mean(true_fn(X_pre))))
print("Estimate: {:.3f}".format(ta_effect))
print("ATE Estimate Interval: ({:.3f}, {:.3f})".format(ta_effect_conf[0], ta_effect_conf[1]))
print("ATE Estimate Std: {:.3f}".format(dmlateiv_obj.std))
```
# Exogenous Treatment Approaches
```
# Simply regressing y on T
from utilities import StatsModelLinearRegression
StatsModelLinearRegression().fit(T.reshape(-1, 1), y).summary()
# Regressing y on X, X*T, T and setting CATE(X) = Pred(X, X*1, 1) - Pred(X, X*0, 0))
from econml.utilities import hstack
est = StatsModelLinearRegression().fit(hstack([X, X*T.reshape(-1, 1), T.reshape(-1, 1)]), y)
T_one = np.ones(T.shape[0])
T_zero = np.zeros(T.shape[0])
driv_cate = est.predict(hstack([X, X*T_one.reshape(-1, 1), T_one.reshape(-1, 1)]))-\
est.predict(hstack([X, X*T_zero.reshape(-1, 1), T_zero.reshape(-1, 1)]))
print(np.mean(driv_cate))
# We can also see how it compares to the true CATE at each target point and calculate MSE
plt.title("Exog-T CATE: MSE {:.2}".format(np.mean((true_fn(X_pre) - driv_cate)**2)))
plot_separate(X, X_pre, driv_cate)
plt.show()
# Doing a full fledged DML for conditionally exogenous treatment
from econml.dml import DML
dml = DML(model_Y_X(), model_T_X(), LassoCV(cv=3))
dml.fit(y, T, X)
dml_cate = dml.effect(X)
print(np.mean(dml_cate))
# We can also see how it compares to the true CATE at each target point and calculate MSE
plt.title("Exog-T CATE: MSE {:.2}".format(np.mean((true_fn(X_pre) - dml_cate)**2)))
plot_separate(X, X_pre, dml_cate)
plt.show()
```
# DMLIV CATE
```
from dml_iv import DMLIV, GenericDMLIV
from utilities import SelectiveLasso, SeparateModel
from sklearn.linear_model import LassoCV
from econml.utilities import hstack
np.random.seed(123)
# We now specify the features to be used for heterogeneity. We will fit a CATE model of the form
# theta(X) = <theta, phi(X)>
# for some set of features phi(X). The featurizer needs to support fit_transform, that takes
# X and returns phi(X). We need to include a bias if we also want a constant term.
dmliv_featurizer = lambda: PolynomialFeatures(degree=1, include_bias=True)
# Then we need to specify a model to be used for fitting the parameters theta in the linear form.
# This model will minimize the square loss:
# (Y - E[Y|X] - <theta, phi(X)> * (E[T|X,Z] - E[T|X]))**2
# potentially with some regularization on theta. Here we use an ell_1 penalty on theta
# dmliv_model_effect = lambda: LinearRegression()
# We could also use LassoCV to select the regularization weight in the final stage with
# cross validation.
# dmliv_model_effect = lambda: LassoCV(fit_intercept=False, cv=3)
# If we also have a prior that there is no effect heterogeneity we can use a selective lasso
# that does not penalize the constant term in the CATE model
feature_inds = np.arange(1, X.shape[1]+1)
dmliv_model_effect = lambda: SelectiveLasso(feature_inds, LassoCV(cv=5, n_jobs=-1))
cate = DMLIV(model_Y_X(), model_T_X(), model_T_XZ(),
dmliv_model_effect(), dmliv_featurizer(),
n_splits=10, # number of splits to use for cross-fitting
binary_instrument=True, # a flag whether to stratify cross-fitting by instrument
binary_treatment=True # a flag whether to stratify cross-fitting by treatment
)
"""
dmliv_model_effect = lambda: model()
cate = GenericDMLIV(model_Y_X(), model_T_X(), model_T_XZ(),
dmliv_model_effect(),
n_splits=10, # number of splits to use for cross-fitting
binary_instrument=True, # a flag whether to stratify cross-fitting by instrument
binary_treatment=True # a flag whether to stratify cross-fitting by treatment
)"""
%%time
cate.fit(y, T, X, Z, store_final=True)
dmliv_effect = cate.effect(X)
# We can average the CATE to get an ATE
print("ATE Estimate: {:.3f}".format(np.mean(dmliv_effect)))
print("True ATE: {:.3f}".format(np.mean(true_fn(X_pre))))
# We can also see how it compares to the true CATE at each target point and calculate MSE
plt.title("DMLIV CATE: MSE {:.2}".format(np.mean((true_fn(X_pre) - dmliv_effect)**2)))
plot_separate(X, X_pre, dmliv_effect)
plt.show()
##histogram of estimated treatment effect
import seaborn as sns
sns.distplot(dmliv_effect, label='est')
sns.distplot(true_fn(X_pre), label='true')
plt.legend()
plt.xlabel("Treatment Effect")
plt.ylabel("Frequency")
plt.show()
```
# Algorithm 3 - DRIV ATE
```
from utilities import SubsetWrapper, StatsModelLinearRegression, ConstantModel
from dr_iv import IntentToTreatDRIV
from utilities import WeightWrapper
np.random.seed(123)
# For intent to treat DRIV we need a flexible model of the CATE to be used in the preliminary estimation.
# This flexible model needs to accept sample weights at fit time. Here we use a weightWrapper to wrap
# a lasso estimator. WeightWrapper requires a linear model with no intercept, hence the Pipeline
# that adds a bias to the features.
driv_flexible_model_effect = lambda: WeightWrapper(Pipeline([('bias', PolynomialFeatures(degree=1, include_bias=True)),
('lasso', SelectiveLasso(np.arange(1, X.shape[1]+1), LassoCV(cv=5, n_jobs=-1)))]))
# Then we can also define any final model to project to. Here we project to a constant model to get an ATE
driv_final_model_effect = lambda: ConstantModel()
dr_cate = IntentToTreatDRIV(model_Y_X(), model_T_XZ(),
driv_flexible_model_effect(),
final_model_effect=driv_final_model_effect(),
cov_clip=0.0001,
n_splits=10)
%%time
dr_cate.fit(y, T, X, Z, store_final=True)
dr_cate.effect_model.summary()
lr = LinearRegression().fit(X, true_fn(X_pre))
print(lr.coef_)
print(lr.intercept_)
```
## Projecting to subset
```
from utilities import WeightWrapper
subset_names = X_data.columns.values
#subset_names = set(['days_visited_as_pre', 'is_existing_member'])
# list of indices of features X to use in the final model
feature_inds = np.argwhere([(x in subset_names) for x in X_data.columns.values]).flatten()
print(feature_inds)
# Because we are projecting to a low dimensional model space, we can
# do valid inference and we can use statsmodel linear regression to get all
# the hypothesis testing capability
lr_driv_model_effect = lambda: SubsetWrapper(StatsModelLinearRegression(),
feature_inds # list of indices of features X to use in the final model
)
dr_cate.refit_final(lr_driv_model_effect())
driv_cate = dr_cate.effect(X[:, feature_inds])
##histogram of estimated treatment effect
import seaborn as sns
sns.distplot(driv_cate, label='est')
sns.distplot(true_fn(X_pre), label='true')
plt.legend()
plt.xlabel("Treatment Effect")
plt.ylabel("Frequency")
plt.show()
# We can also see how it compares to the true CATE at each target point and calculate MSE
plt.title("DMLIV CATE: MSE {:.2}".format(np.mean((true_fn(X_pre) - driv_cate)**2)))
plot_separate(X, X_pre, driv_cate)
plt.show()
# To get the ATE we look at the params of the fitted statsmodel
dr_effect = np.mean(driv_cate)
dr_effect
# To get the statsmodel summary we look at the effect_model, which is the pipeline, we then look
# at the reg step of the pipeline which is the statsmodel wrapper and then we look
# at the model attribute of the statsmodel wrapper and print the summary()
dr_cate.effect_model.summary()
print("ATE Estimate: {:.2f}".format(dr_effect))
print("True ATE: {:.2f}".format(np.mean(true_fn(X_pre))))
lr = LinearRegression().fit(X[:, feature_inds], true_fn(X_pre).reshape(-1, 1))
print(lr.coef_)
print(lr.intercept_)
# We can also evaluate coverage and create prediction intervals using statsmodels attributes
from statsmodels.sandbox.regression.predstd import wls_prediction_std
res = dr_cate.effect_model.model
predictions = res.get_prediction(PolynomialFeatures(degree=1, include_bias=True).fit_transform(X[:, feature_inds]))
frame = predictions.summary_frame(alpha=0.05)
pred = frame['mean']
iv_l = frame['mean_ci_lower']
iv_u = frame['mean_ci_upper']
# This is the true CATE functions
theta_true = true_fn(X_pre)
# This is the true projection of the CATE function on the subspace of linear functions of the
# subset of the features used in the projection
true_proj = LinearRegression().fit(X[:, feature_inds], theta_true).predict(X[:, feature_inds])
# Are we covering the true projection
covered = (true_proj <= iv_u) & (true_proj >= iv_l)
print("Coverage of True Projection: {:.2f}".format(np.mean(covered)))
fig, ax = plt.subplots(figsize=(8,6))
order = np.argsort(X[:, feature_inds[0]])
ax.plot(X[order, feature_inds[0]], iv_u[order], 'r--')
ax.plot(X[order, feature_inds[0]], iv_l[order], 'r--')
ax.plot(X[order, feature_inds[0]], pred[order], 'g--.', label="pred")
#ax.plot(X[order, feature_inds[0]], theta_true[order], 'b-', label="True", alpha=.3)
ax.plot(X[order, feature_inds[0]], true_proj[order], 'y-', label="TrueProj", alpha=.3)
ax.legend(loc='best')
plt.show()
```
# Lasso Final CATE
```
lasso_driv_model_effect = lambda: WeightWrapper(Pipeline([('bias', PolynomialFeatures(degree=1, include_bias=True)),
('lasso', SelectiveLasso(np.arange(1, X.shape[1]+1),
LassoCV(cv=5, n_jobs=-1, fit_intercept=False)))]))
dr_cate.refit_final(lasso_driv_model_effect())
# We can also see how it compares to the true CATE at each target point and calculate MSE
dr_effect = dr_cate.effect(X)
plt.title("DMLIV CATE: MSE {:.2}".format(np.mean((true_fn(X_pre) - dr_effect)**2)))
plot_separate(X, X_pre, dr_effect)
plt.show()
```
# Random Forest Based CATE and Tree Explainer
```
from dml_iv import DMLIV
from dr_iv import DRIV, ProjectedDRIV
from utilities import SubsetWrapper
from sklearn.ensemble import RandomForestRegressor, GradientBoostingRegressor
np.random.seed(123)
# We need a model for the final regression that will fit the function theta(X)
# Now we use a linear model and a lasso.
rf_driv_model_effect = lambda: RandomForestRegressor(n_estimators=100, max_depth=3, min_impurity_decrease=0.1,
min_samples_leaf=500, bootstrap=True)
rf_dr_cate = dr_cate.refit_final(rf_driv_model_effect())
rf_dr_effect = rf_dr_cate.effect(X)
##histogram of estimated treatment effect
import seaborn as sns
sns.distplot(rf_dr_effect, label='est')
sns.distplot(true_fn(X_pre), label='true')
plt.legend()
plt.xlabel("Treatment Effect")
plt.ylabel("Frequency")
plt.show()
print("ATE Estimate: {:.2f}".format(np.mean(rf_dr_effect)))
print("True ATE: {:.2f}".format(np.mean(true_fn(X_pre))))
plt.title("DRIV CATE: MSE {:.2}".format(np.mean((true_fn(X_pre) - rf_dr_effect)**2)))
plot_separate(X, X_pre, rf_dr_effect)
plt.show()
import shap
import pandas as pd
Xdf = pd.DataFrame(X, columns=X_data.columns)
# explain the model's predictions using SHAP values
explainer = shap.TreeExplainer(rf_dr_cate.effect_model)
shap_values = explainer.shap_values(Xdf)
# visualize the first prediction's explanation (use matplotlib=True to avoid Javascript)
shap.force_plot(explainer.expected_value, shap_values[0,:], Xdf.iloc[0,:], matplotlib=True)
shap.summary_plot(shap_values, Xdf)
shap.summary_plot(shap_values, Xdf, plot_type='bar')
```
# CATE with DRIV-RW
```
from utilities import SubsetWrapper, StatsModelLinearRegression, ConstantModel
from dr_iv import IntentToTreatDRIV
from utilities import WeightWrapper
np.random.seed(123)
# For intent to treat DRIV we need a flexible model of the CATE to be used in the preliminary estimation.
# This flexible model needs to accept sample weights at fit time. Here we use a weightWrapper to wrap
# a lasso estimator. WeightWrapper requires a linear model with no intercept, hence the Pipeline
# that adds a bias to the features.
rf_driv_model_effect = lambda: GradientBoostingRegressor(n_estimators=30, max_depth=2, min_impurity_decrease=0.001,
min_samples_leaf=200)
rf_dr_cate = IntentToTreatDRIV(model_Y_X(), model_T_XZ(),
rf_driv_model_effect(),
opt_reweighted=True, # re-weighting the final loss for variance reduction
cov_clip=1e-7,
n_splits=10)
%%time
rf_dr_cate.fit(y, T, X, Z, store_final=True)
rf_dr_effect = rf_dr_cate.effect(X)
plt.title("DRIV CATE: MSE {:.2}".format(np.mean((true_fn(X_pre) - rf_dr_effect)**2)))
plot_separate(X, X_pre, rf_dr_effect)
plt.show()
import shap
import pandas as pd
Xdf = pd.DataFrame(X, columns=X_data.columns)
# explain the model's predictions using SHAP values
explainer = shap.TreeExplainer(rf_dr_cate.effect_model)
shap_values = explainer.shap_values(Xdf)
# visualize the first prediction's explanation (use matplotlib=True to avoid Javascript)
shap.force_plot(explainer.expected_value, shap_values[0,:], Xdf.iloc[0,:], matplotlib=True)
shap.summary_plot(shap_values, Xdf)
shap.summary_plot(shap_values, Xdf, plot_type='bar')
```
### Lasso CATE
```
lasso_driv_model_effect = lambda: WeightWrapper(Pipeline([('bias', PolynomialFeatures(degree=1, include_bias=True)),
('lasso', SelectiveLasso(np.arange(1, X.shape[1]+1),
LassoCV(cv=5, n_jobs=-1, fit_intercept=False)))]))
rf_dr_cate.refit_final(lasso_driv_model_effect())
# We can also see how it compares to the true CATE at each target point and calculate MSE
dr_effect = rf_dr_cate.effect(X)
plt.title("DMLIV CATE: MSE {:.2}".format(np.mean((true_fn(X_pre) - dr_effect)**2)))
plot_separate(X, X_pre, dr_effect)
plt.show()
print("ATE Estimate: {:.3f}".format(np.mean(dr_effect)))
from utilities import ConstantModel
rf_dr_cate.refit_final(ConstantModel(), opt_reweighted=False)
rf_dr_cate.effect_model.summary()
```
# Using Generic DRIV
```
from dml_iv import DMLIV
from dr_iv import DRIV, ProjectedDRIV
from utilities import SubsetWrapper, StatsModelLinearRegression, ConstantModel
np.random.seed(123)
# For DRIV we need a model for predicting E[T*Z | X]. We use a classifier
model_TZ_X = lambda: model_clf()
# For generic DRIV we also need to provide a preliminary effect model. Here we use DMLIV
dmliv_featurizer = lambda: PolynomialFeatures(degree=1, include_bias=True)
dmliv_model_effect = lambda: SelectiveLasso(np.arange(1, X.shape[1]+1), LassoCV(cv=5, n_jobs=-1))
prel_model_effect = DMLIV(model_Y_X(), model_T_X(), model_T_XZ(),
dmliv_model_effect(), dmliv_featurizer(), n_splits=1)
dr_cate = DRIV(model_Y_X(), model_T_X(), model_Z_X(), # same as in DMLATEIV
prel_model_effect, # preliminary model for CATE, must support fit(y, T, X, Z) and effect(X)
model_TZ_X(), # model for E[T * Z | X]
ConstantModel(), # model for final stage of fitting theta(X)
cov_clip=.0001, # covariance clipping to avoid large values in final regression from weak instruments
n_splits=10, # number of splits to use for cross-fitting
binary_instrument=True, # a flag whether to stratify cross-fitting by instrument
binary_treatment=True, # a flag whether to stratify cross-fitting by treatment
opt_reweighted=False
)
%%time
dr_cate.fit(y, T, X, Z, store_final=True)
dr_cate.effect_model.summary()
```
# Deep CATE Models
```
from deep_dr_iv import DeepIntentToTreatDRIV
import keras
treatment_model = lambda: keras.Sequential([keras.layers.Dense(100, activation='relu'),
keras.layers.Dropout(0.17),
keras.layers.Dense(100, activation='relu'),
keras.layers.Dropout(0.17),
keras.layers.Dense(100, activation='relu', name='final'),
keras.layers.Dropout(0.17),
keras.layers.Dense(1)])
keras_fit_options = { "epochs": 100,
"batch_size": 32,
"validation_split": 0.1,
"callbacks": [keras.callbacks.EarlyStopping(patience=10, restore_best_weights=True)],
"verbose": 0}
dr_cate = DeepIntentToTreatDRIV(model_Y_X(), model_T_XZ(),
treatment_model(),
training_options=keras_fit_options,
cov_clip=0.0001,
opt_reweighted=False,
n_splits=3)
%%time
dr_cate.fit(y, T, X, Z, store_final=True)
dr_effect = dr_cate.effect(X)
plt.title("DRIV CATE: MSE {:.2}".format(np.mean((true_fn(X_pre) - dr_effect)**2)))
plot_separate(X, X_pre, dr_effect)
plt.show()
from utilities import ConstantModel
dr_cate.refit_final(ConstantModel(), opt_reweighted=False)
dr_cate.effect_model.summary()
```
| github_jupyter |
<a href="https://colab.research.google.com/github/DanielaManate/SentimentAnalysis-TopicModeling/blob/master/SA2_Lexical_Complexity.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Importing Libraries
```
import pandas as pd
import numpy as np
# Convert text to vectors of numbers => document-term-matrix
from sklearn.feature_extraction.text import CountVectorizer
# for tokenizing
import nltk
nltk.download('punkt')
from nltk.tokenize import word_tokenize
# plots
import matplotlib.pyplot as plt
# for wordlcloud graphs
from wordcloud import WordCloud
from google.colab import drive
drive.mount('/content/gdrive')
path = 'gdrive/My Drive/GH_NLP_Course/Data/'
```
# Reading Dataset
```
reviews = pd.read_csv(path + 'Input/2.input_data_prepped.csv')
reviews.head()
reviews.dtypes
# Convert text_prep to string to counteract issues later
reviews['text_prep'] = reviews['text_prep'].astype(str)
```
# Lexical Complexity
- Word Frequency: number of words per corpus. Top Words
- Word Length: average length of individual words in a text
- Lexical Diversity: number of unique words used in a text
- Lexical Density: the number of unique words divided by the total number of words (word repetition)
# Bag of Words - Top 6000
```
len(reviews)
```
Matrix VS Array
- matrices are strictly 2-dimensional
- numpy arrays (ndarrays) are N-dimensional.
DF VS Array
- DF = 2-dimensional array with labeled axes (matrix with column names for columns, and index labels for rows)
- A single column or row in a Pandas DataFrame is a Pandas series — a one-dimensional array with axis labels
Sparse Matrix
= Matrices that contain mostly zero values are called sparse, distinct from matrices where most of the values are non-zero, called dense.
Compressed Sparse Row Matrix: a compressed matrix, to save space (contains one vector for index pointers, one vector for indexes, one vector with data, does not save 0s)
Converting compressed sparse matrix to
- dense matrix: counts_per_review.todense()
- array: counts_per_review.toarray()
### Create a document-term-matrix
```
# max_features = If not None, build a vocabulary that only consider the top
# max_features ordered by term frequency across the corpus.
# construct the vocab on most frequent 6000 words
vec = CountVectorizer(max_features = 6000)
# creates a matrix with rows = documents, cols = words
counts_per_review = vec.fit_transform(reviews.text_prep)
# the 6000 words => will become the name of the cols
review_words = vec.get_feature_names()
dtm = pd.DataFrame(counts_per_review.toarray(),
columns=review_words)
dtm.head()
dtm['pron']
# Delete pron column
del dtm['pron']
dtm.shape
reviews.text_prep.iloc[250]
dtm['third'].iloc[250]
# Count number of words in text prep
reviews['text_prep_tokens'] = reviews['text_prep'].apply(word_tokenize)
reviews['word_len_prep'] = reviews['text_prep_tokens'].apply(len)
reviews.head()
```
### Create a new column, text_prep_lim, that contains only the words in BoW
Lambda Function: anonymous function (a function that is defined without a name => we don't use def)
```
reviews['text_prep_lim_tokens'] = reviews['text_prep_tokens'].apply(lambda x:
[word for word in x if word in review_words])
reviews.head()
reviews['word_len_prep_lim'] = reviews['text_prep_lim_tokens'].apply(len)
reviews.head()
# # Change output from list of strings to sentence
reviews['text_prep_lim'] = reviews['text_prep_lim_tokens'].apply(lambda x: ' '.join(x))
reviews.head()
reviews['text_prep_lim'].iloc[250]
print(reviews.text.iloc[0])
print(reviews.text_prep.iloc[0])
print(reviews.text_prep_lim.iloc[0])
# the word 'croed' & pronoun '-PRON-' are missing from text_prep_lim
reviews.columns
reviews = reviews[['rest_id', 'text', 'rating', 'char_count', 'positive',
'text_prep', 'text_prep_tokens', 'word_len_prep',
'text_prep_lim', 'text_prep_lim_tokens','word_len_prep_lim']]
reviews.head(2)
```
### Save BOW DTM to csv
```
dtm.to_csv(path + 'Output/1.dtm_1_bow.csv', index=False)
reviews.to_csv(path + 'Input/3.input_data_prepped_bow.csv',
index = False)
```
# Word Cloud
```
dtm.head()
```
- Axis 0 will act on all the ROWS in each COLUMN
- Axis 1 will act on all the COLUMNS in each ROW
```
word_cloud_freq = dtm.sum(axis=0)
word_cloud_freq
wordcloud = WordCloud(max_words=100,
background_color="white",
font_path = path + 'Input/SignPainter.ttc',
colormap="Blues")
wordcloud.generate_from_frequencies(word_cloud_freq)
plt.imshow(wordcloud)
# Turn off axis lines and labels
plt.axis("off")
plt.show()
```
# Word Cloud Positive - Top 20
```
dtm_pos = dtm[reviews['positive']==1].copy()
dtm_pos.shape
word_cloud_freq_pos = dtm_pos.sum(axis=0)
wordcloud = WordCloud(max_words=20,
background_color="white",
font_path = path + 'Input/SignPainter.ttc',
colormap="Greens")
wordcloud.generate_from_frequencies(word_cloud_freq_pos)
plt.imshow(wordcloud)
plt.axis("off")
plt.show()
wordcloud.words_
# 8 of the 20 words are positive words
# always, amazing, delicious, friendly, good, great, like, love
```
# Word Cloud Negative - Top 20
```
dtm_neg = dtm[reviews['positive']==0].copy()
dtm_neg.shape
word_cloud_freq_neg = dtm_neg.sum(axis=0)
word_cloud_freq_neg
wordcloud = WordCloud(max_words=20,
background_color="white",
font_path = path + 'Input/SignPainter.ttc',
colormap="Reds")
wordcloud.generate_from_frequencies(word_cloud_freq_neg)
plt.imshow(wordcloud)
plt.axis("off")
plt.show()
wordcloud.words_
# 1 of the 20 words is negative, 2 are positive
# wait
# good, like
```
# Word Frequency
```
# Most Frequent Words
word_freq = pd.DataFrame(dtm.sum(axis=0)).reset_index()
word_freq.head()
word_freq.columns = ['Word', 'Count']
word_freq.head()
word_freq = word_freq.sort_values(by = 'Count',
ascending = False)
word_freq.head()
# Plot Top 20 Words
top20 = word_freq[0:20].copy().reset_index(drop=True)
top20.head()
plt.barh(y = top20['Word'],
width = top20['Count'],
height = 0.5,
align='center')
plt.title('Top 20 Tokens')
plt.xlabel('Word')
plt.ylabel('Frequency')
plt.show();
```
# Word Length
```
# Word Length
word_freq['word_len'] = word_freq['Word'].apply(len)
word_freq.head()
word_freq.describe()
# Average word length = 6 chars
```
# Lexical Diversity: How many unique words in each review?
```
dtm.head()
# For each doc, count number of unique words
# <=> count how many columns have non-zero values
lexical = pd.DataFrame(data=np.count_nonzero(dtm, axis=1),
columns = ['Unique Words'])
lexical.head()
lexical['Total Words'] = dtm.sum(axis=1)
lexical.head()
reviews['text_prep_lim'].iloc[0]
# dinner & pm appear twice;
lexical.describe()
# On avg, 34 unique words per review, 40 total words per review
lexical['positive']=reviews['positive'].copy()
lexical.head()
lexical.groupby('positive').mean()
# On avg, 42 unique words per negative review, 30 unique words per positive review
# On avg, 53 total words per negative review, 35 unique words per positive review
```
# Lexical Density: Unique Words / Total Words
```
# As lexical density increases, repetition decreases.
lexical.head()
lexical['Density'] = lexical['Unique Words']/lexical['Total Words']
lexical.head()
lexical['Density'].hist()
lexical.groupby('positive').mean()
# Positive Reviews have larger density, which suggests lower repetition.
```
# Insights
Conclusions
- Negative reviews have a larger average number of unique and total words, which suggests that users leave a longer review when they have negative feedback
- Negative reviews have a lower density, which suggests that users tend to repeat words when they have negative feedback
- The most common 20 words in negative reviews are not negative words (except one word - 'wait'), while almost half of the most common 20 words in positive reviews are positive words. Negations (e.g. 'not good') greatly affect Sentiment Analysis, and we extracted words like 'not' using stopwords from reviews.
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.