text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
# Analyze game tree with position evaluation
## Import
```
# Game graph library
import igraph
import chess
import math
class Game():
def __init__(self, game):
self.game = game
@property
def moves_uci(self):
res = []
node = self.game
while not node.is_end():
node = node.next()
res.append(node.uci())
return res
@property
def moves_nodes(self):
res = []
node = self.game
while not node.is_end():
node = node.next()
res.append(node)
return res
class GamesGraph():
def __init__(self):
self.graph = igraph.Graph(directed=True)
def add_move(self, start_fen, end_fen, uci):
vs = self._ensure_vertex(start_fen)
vt = self._ensure_vertex(end_fen)
try:
e = self.graph.es.find(_source=vs.index, _target=vt.index)
e["count"] += 1
except:
e = self.graph.add_edge(vs, vt)
e["uci"] = uci
e["count"] = 1
@property
def start_node(self):
return self.graph.vs.find(chess.STARTING_FEN)
def _ensure_vertex(self, fen):
try:
return self.graph.vs.find(fen)
except:
v = self.graph.add_vertex(name=fen)
v["fen"] = fen
v["turn"] = chess.Board(fen).turn
return v
def games_graph(games, max_moves):
gr = GamesGraph()
for game in games:
start_fen = game.game.board().fen()
for move in game.moves_nodes[:max_moves]:
fen = move.board().fen()
uci = move.uci()
gr.add_move(start_fen, fen, uci)
start_fen = fen
return gr
def compute_edge_weights_uniform(vertex):
all_count = vertex.degree(mode="out")
for edge in vertex.out_edges():
edge["prob"] = 1.0
edge["weight"] = 0.0
def compute_edge_weights_counting(vertex):
all_count = sum(map(lambda x: x["count"], vertex.out_edges()))
for edge in vertex.out_edges():
# Certainty doesn't exist... Let's put a 90% ceiling.
prob = min(edge["count"] / all_count, 0.9)
edge["prob"] = prob
edge["weight"] = -math.log(prob)
def compute_graph_weights(graph, black_uniform=False, white_uniform=False):
# Compute the graph weights such that:
# * The distance is the inverse of the probability to go from source to destination.
# * Summation of two weights is the same as multiplying the probability.
#
# count: ranges from 1 to max; max = sum(out_edges["count"]).
# prob: count / max_count; [0; 1]
# weigth: -log(prob); [0; +inf] ~ [very_likely; unlikely]
for vertex in graph.graph.vs:
if vertex["turn"] == chess.WHITE and white_uniform:
compute_edge_weights_uniform(vertex)
elif vertex["turn"] == chess.BLACK and black_uniform:
compute_edge_weights_uniform(vertex)
else:
compute_edge_weights_counting(vertex)
import chess.pgn
import io
import json
with open('playerx-games.json') as f:
data = json.load(f)
games = []
for game in data:
pgn = io.StringIO(game)
games.append(Game(chess.pgn.read_game(pgn)))
white_games = [g for g in games if g.game.headers["White"] == "playerx"]
black_games = [g for g in games if g.game.headers["Black"] == "playerx"]
len(games), len(white_games), len(black_games)
# Load the evaluations
import chess
def load_evals_json(path):
with open(path) as f:
evaljs = json.load(f)
evals = {}
for pos in evaljs:
evals[pos["fen"]] = pos["eval"]
# add the initial position
evals[chess.STARTING_FEN] = {
"type": "cp",
"value": 0,
}
return evals
# Returns [-1;1]
def rating(ev, fen):
val = ev["value"]
if ev["type"] == "cp":
# Clamp to -300, +300. Winning a piece is enough.
val = max(-300, min(300, val))
return val / 300.0
if val > 0: return 1.0
if val < 0: return -1.0
# This is mate, but is it white or black?
b = chess.Board(fen)
return 1.0 if b.turn == chess.WHITE else -1.0
# Returns [0;1], where 0 is min advantage, 1 is max for black.
def rating_black(ev, fen):
return -rating(ev, fen) * 0.5 + 0.5
# Returns [0;1], where 0 is min advantage, 1 is max for black.
def rating_white(ev, fen):
return rating(ev, fen) * 0.5 + 0.5
def compute_rating(evals, rating_fn):
for fen in evals.keys():
ev = evals[fen]
evals[fen]["rating"] = rating_fn(ev, fen)
def update_graph_rating(g, evals):
for v in g.graph.vs:
v["rating"] = evals[v["fen"]]["rating"]
import chess
from functools import reduce
class Line():
def __init__(self, end_node, moves, cost):
self.end_node = end_node
self.moves = moves
self.cost = cost
@property
def moves_uci(self):
return [e["uci"] for e in self.moves]
@property
def rating(self):
return self.end_node["rating"]
@property
def probability(self):
return reduce(lambda x, y: x*y, [e["prob"] for e in self.moves])
@property
def end_board(self):
return chess.Board(self.end_node["fen"])
@property
def end_fen(self):
return self.end_node["fen"]
def __str__(self):
return "{}: {} (prob={} rating={})".format(
self.end_node["fen"],
self.moves_uci,
self.probability,
self.rating,
)
def __repr__(self):
return self.__str__()
def compute_line(graph, path):
# Skip empty paths.
if len(path) < 2:
return None
end_node = graph.graph.vs.find(path[-1])
cost = 0
moves = []
for i in range(len(path) - 1):
edge = graph.graph.es.find(_source=path[i], _target=path[i+1])
cost += edge["weight"]
moves.append(edge)
return Line(end_node, moves, cost)
def best_lines(lines, min_rating=0.5):
lines = filter(lambda x: x is not None, lines)
return [
l for l in sorted(lines, key=lambda x: x.cost)
if l.rating > min_rating
]
```
## Graph test
```
import igraph
g = igraph.Graph(directed=True)
g.add_vertex(name="a")
g.add_vertex(name="b")
a = g.vs.find("a")
b = g.vs.find("b")
print("a: ", a, "b: ", b)
print(a.index)
g.add_edge(a, b, name="foo")
print("edge 0, 1: ", g.es.find(_source=a.index, _target=b.index) != None)
igraph.plot(g, bbox=(200, 200))
import igraph
import chess
g = games_graph(black_games[:8], 4)
fen_evals = load_evals_json('eval-black.json')
compute_rating(fen_evals, rating_white)
update_graph_rating(g, fen_evals)
style = {
"edge_label": g.graph.es["count"],
"edge_label_color": "blue",
"vertex_label": ["{:.2}".format(x) for x in g.graph.vs["rating"]],
"vertex_label_size": 8,
"vertex_label_color": "red",
"vertex_color": [("white" if x == chess.WHITE else "black") for x in g.graph.vs["turn"]],
"vertex_shape": ["rectangle"] + ["circle" for _ in g.graph.vs][1:]
}
igraph.plot(g.graph, **style)
# Test weights without correcting for my moves equal probability.
compute_graph_weights(g)
style = {
"edge_label": ["{:.2}".format(x) for x in g.graph.es["weight"]],
"edge_label_color": "blue",
"vertex_color": [("white" if x == chess.WHITE else "black") for x in g.graph.vs["turn"]],
"vertex_shape": ["rectangle"] + ["circle" for _ in g.graph.vs][1:]
}
igraph.plot(g.graph, **style)
# Test the weights.
compute_graph_weights(g, white_uniform=True)
style = {
"edge_label": ["{:.2}".format(x) for x in g.graph.es["weight"]],
"edge_label_color": "blue",
"vertex_color": [("white" if x == chess.WHITE else "black") for x in g.graph.vs["turn"]],
"vertex_shape": ["rectangle"] + ["circle" for _ in g.graph.vs][1:]
}
igraph.plot(g.graph, **style)
# Shortest paths from initial to every other position.
sp = g.graph.get_shortest_paths(g.start_node, weights=g.graph.es["weight"])
sp
lines = [compute_line(g, p) for p in sp]
lines = best_lines(lines)
lines
print(lines[8])
lines[8].end_board
```
## Black games
```
# Load the games
graph = games_graph(black_games, 10)
len(graph.graph.vs["name"])
# Load the evaluations
fen_evals = load_evals_json('eval-black.json')
compute_rating(fen_evals, rating_white)
len(fen_evals)
update_graph_rating(graph, fen_evals)
compute_graph_weights(graph, white_uniform=False)
# Shortest paths from initial to every other position.
sp = graph.graph.get_shortest_paths(graph.start_node, weights=graph.graph.es["weight"])
all_lines = [compute_line(graph, p) for p in sp]
lines = best_lines(all_lines, 0.6)
lines[:10]
import pandas
data = pandas.Series([
x.probability
for x in all_lines
if x is not None and x.probability < 0.02
])
df = pandas.DataFrame({"prob": data}, columns=["prob"])
df.plot.hist(bins=40)
high_adv = [x for x in lines if x.rating > 0.7][:10]
print(high_adv[0])
high_adv[0].end_board
```
Not good. Moves the Q out early, hoping for a black mistake.
```
print(high_adv[1])
high_adv[1].end_board
```
+1.4 but played the wrong move only 3/16 times.
```
print(high_adv[2])
high_adv[2].end_board
not_e4 = [l for l in lines if l.moves_uci[0] != "e2e4" and l.rating > 0.7]
print(not_e4[1])
not_e4[1].end_board
```
Interesting position. He seems to often get it wrong. The main last moves are:
* Nc6 (2) +0.8
* e6 (2) +0.1
* c5 +0.3
* a6 +0.2
* e5 +1.5
| github_jupyter |
# Research Problem
Last year, I read a paper titled, "Feature Selection Methods for Identifying Genetic Determinants of Host Species in RNA Viruses". This year, I read another paper titled, "Predicting host tropism of influenza A virus proteins using random forest". The essence of these papers were to predict influenza virus host tropism from sequence features. The particular feature engineering steps were somewhat distinct, in which the former used amino acid sequences encoded as binary 1/0s, while the latter used physiochemical characteristics of the amino acid sequences instead. However, the core problem was essentially identical - predict a host classification from influenza protein sequence features. Random forest classifiers were used in both papers, and is a powerful method for identifying non-linear mappings from features to class labels. My question here was to see if I could get comparable performance using a simple neural network.
# Data
I downloaded influenza HA sequences from the Influenza Research Database. Sequences dated from 1980 to 2015. Lab strains were excluded, duplicates allowed (captures host tropism of certain sequences). All viral subtypes were included.
Below, let's take a deep dive into what it takes to construct an artificial neural network!
The imports necessary for running this notebook.
```
! echo $PATH
! echo $CUDA_ROOT
import pandas as pd
import numpy as np
from Bio import SeqIO
from Bio import AlignIO
from Bio.Align import MultipleSeqAlignment
from collections import Counter
from sklearn.preprocessing import LabelBinarizer
from sklearn.cross_validation import train_test_split
from sklearn.ensemble import RandomForestClassifier, ExtraTreesClassifier, GradientBoostingClassifier
from sklearn.metrics import mutual_info_score as mi
from lasagne import layers
from lasagne.updates import nesterov_momentum
from nolearn.lasagne import NeuralNet
import theano
```
Read in the viral sequences.
```
sequences = SeqIO.to_dict(SeqIO.parse('20150902_nnet_ha.fasta', 'fasta'))
# sequences
```
The sequences are going to be of variable length. To avoid the problem of doing multiple sequence alignments, filter to just the most common length (i.e. 566 amino acids).
```
lengths = Counter()
for accession, seqrecord in sequences.items():
lengths[len(seqrecord.seq)] += 1
lengths.most_common(1)[0][0]
```
There are sequences that are ambiguously labeled. For example, "Environment" and "Avian" samples. We would like to give a more detailed prediction as to which hosts it likely came from. Therefore, take out the "Environment" and "Avian" samples.
```
# For convenience, we will only work with amino acid sequencees of length 566.
final_sequences = dict()
for accession, seqrecord in sequences.items():
host = seqrecord.id.split('|')[1]
if len(seqrecord.seq) == lengths.most_common(1)[0][0]:
final_sequences[accession] = seqrecord
```
Create a `numpy` array to store the alignment.
```
alignment = MultipleSeqAlignment(final_sequences.values())
alignment_array = np.array([list(rec) for rec in alignment])
```
The first piece of meat in the code begins here. In the cell below, we convert the sequence matrix into a series of binary `1`s and `0`s, to encode the features as numbers. This is important - AFAIK, almost all machine learning algorithms require numerical inputs.
```
# Create an empty dataframe.
# df = pd.DataFrame()
# # Create a dictionary of position + label binarizer objects.
# pos_lb = dict()
# for pos in range(lengths.most_common(1)[0][0]):
# # Convert position 0 by binarization.
# lb = LabelBinarizer()
# # Fit to the alignment at that position.
# lb.fit(alignment_array[:,pos])
# # Add the label binarizer to the dictionary.
# pos_lb[pos] = lb
# # Create a dataframe.
# pos = pd.DataFrame(lb.transform(alignment_array[:,pos]))
# # Append the columns to the dataframe.
# for col in pos.columns:
# maxcol = len(df.columns)
# df[maxcol + 1] = pos[col]
from isoelectric_point import isoelectric_points
df = pd.DataFrame(alignment_array).replace(isoelectric_points)
# Add in host data
df['host'] = [s.id.split('|')[1] for s in final_sequences.values()]
df = df.replace({'X':np.nan, 'J':np.nan, 'B':np.nan, 'Z':np.nan})
df.dropna(inplace=True)
df.to_csv('isoelectric_point_data.csv')
# Normalize data to between 0 and 1.
from sklearn.preprocessing import StandardScaler
df_std = pd.DataFrame(StandardScaler().fit_transform(df.ix[:,:-1]))
df_std['host'] = df['host']
ambiguous_hosts = ['Environment', 'Avian', 'Unknown', 'NA', 'Bird', 'Sea_Mammal', 'Aquatic_Bird']
unknowns = df_std[df_std['host'].isin(ambiguous_hosts)]
train_test_df = df_std[df_std['host'].isin(ambiguous_hosts) == False]
train_test_df.dropna(inplace=True)
```
With the cell above, we now have a sequence feature matrix, in which the 566 amino acids positions have been expanded to 6750 columns of binary sequence features.
The next step is to grab out the host species labels, and encode them as 1s and 0s as well.
```
set([i for i in train_test_df['host'].values])
# Grab out the labels.
output_lb = LabelBinarizer()
output_lb.fit(train_test_df['host'])
Y = output_lb.fit_transform(train_test_df['host'])
Y = Y.astype(np.float32) # Necessary for passing the data into nolearn.
Y.shape
X = train_test_df.ix[:,:-1].values
X = X.astype(np.float32) # Necessary for passing the data into nolearn.
X.shape
```
Next up, we do the train/test split.
```
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size=0.25, random_state=42)
```
For comparison, let's train a random forest classifier, and see what the concordance is between the predicted labels and the actual labels.
```
rf = RandomForestClassifier()
rf.fit(X_train, Y_train)
predictions = rf.predict(X_test)
predicted_labels = output_lb.inverse_transform(predictions)
# Compute the mutual information between the predicted labels and the actual labels.
mi(predicted_labels, output_lb.inverse_transform(Y_test))
```
By the majority-consensus rule, and using mutual information as the metric for scoring, things look not so bad! As mentioned above, the `RandomForestClassifier` is a pretty powerful method for finding non-linear patterns between features and class labels.
Uncomment the cell below if you want to try the `scikit-learn`'s `ExtraTreesClassifier`.
```
# et = ExtraTreesClassifier()
# et.fit(X_train, Y_train)
# predictions = et.predict(X_test)
# predicted_labels = output_lb.inverse_transform(predictions)
# mi(predicted_labels, output_lb.inverse_transform(Y_test))
```
As a demonstration of how this model can be used, let's look at the ambiguously labeled sequences, i.e. those from "Environment" and "Avian", to see whether we can make a prediction as to what host it likely came frome.
```
# unknown_hosts = unknowns.ix[:,:-1].values
# preds = rf.predict(unknown_hosts)
# output_lb.inverse_transform(preds)
```
Alrighty - we're now ready to try out a neural network! For this try, we will use `lasagne` and `nolearn`, two packages which have made things pretty easy for building neural networks. In this segment, I'm going to not show experiments with multiple architectures, activations and the like. The goal is to illustrate how easy the specification of a neural network is.
The network architecture that we'll try is as such:
- 1 input layer, of shape 6750 (i.e. taking in the columns as data).
- 1 hidden layer, with 300 units.
- 1 output layer, of shape 140 (i.e. each of the class labels).
```
from lasagne import nonlinearities as nl
net1 = NeuralNet(layers=[
('input', layers.InputLayer),
('hidden1', layers.DenseLayer),
#('dropout', layers.DropoutLayer),
#('hidden2', layers.DenseLayer),
#('dropout2', layers.DropoutLayer),
('output', layers.DenseLayer),
],
# Layer parameters:
input_shape=(None, X.shape[1]),
hidden1_num_units=300,
#dropout_p=0.3,
#hidden2_num_units=500,
#dropout2_p=0.3,
output_nonlinearity=nl.softmax,
output_num_units=Y.shape[1],
#allow_input_downcast=True,
# Optimization Method:
update=nesterov_momentum,
update_learning_rate=0.01,
update_momentum=0.9,
regression=True,
max_epochs=100,
verbose=1
)
```
Training a simple neural network on my MacBook Air takes quite a bit of time :). But the function call for fitting it is a simple `nnet.fit(X, Y)`.
```
net1.fit(X_train, Y_train)
```
Let's grab out the predictions!
```
preds = net1.predict(X_test)
preds.shape
```
We're going to see how good the classifier did by examining the class labels. The way to visualize this is to have, say, the class labels on the X-axis, and the probability of prediction on the Y-axis. We can do this sample by sample. Here's a simple example with no frills in the matplotlib interface.
```
import matplotlib.pyplot as plt
%matplotlib inline
plt.bar(np.arange(len(preds[0])), preds[0])
```
Alrighty, let's add some frills - the class labels, the probability of each class label, and the original class label.
```
### NOTE: Change the value of i to anything above!
i = 111
plt.figure(figsize=(20,5))
plt.bar(np.arange(len(output_lb.classes_)), preds[i])
plt.xticks(np.arange(len(output_lb.classes_)) + 0.5, output_lb.classes_, rotation='vertical')
plt.title('Original Label: ' + output_lb.inverse_transform(Y_test)[i])
plt.show()
# print(output_lb.inverse_transform(Y_test)[i])
```
Let's do a majority-consensus rule applied to the labels, and then compute the mutual information score again.
```
preds_labels = []
for i in range(preds.shape[0]):
maxval = max(preds[i])
pos = list(preds[i]).index(maxval)
preds_labels.append(output_lb.classes_[pos])
mi(preds_labels, output_lb.inverse_transform(Y_test))
```
With a score of 0.73, that's not bad either! It certainly didn't outperform the `RandomForestClassifier`, but the default parameters on the RFC were probably pretty good to begin with. Notice how little tweaking on the neural network we had to do as well.
For good measure, these were the class labels. Notice how successful influenza has been in replicating across the many different species!
```
output_lb.classes_
```
# The biology behind this dataset.
A bit more about the biology of influenza.
If you made it this far, thank you for hanging on! How does this mini project relate to the biology of flu?
As the flu evolves and moves between viral hosts, it gradually adapts to that host. This allows it to successfully establish an infection in the host population.
We can observe the viral host as we sample viruses from it. Sometimes, we don't catch it in its adapted state, but it's un-adapted state, as if it had freshly joined in from its other population. That is likely why some of the class labels are mis-identified.
Also, there are environmentally sampled isolates. They obviously aren't simply replicating in the environment (i.e. bodies of water), but in some host, and were shed into the water. For these guys, the host labels won't necessarily match up, as there'll be a stronger signal with particular hosts - whether it be from ducks, pigs or even humans.
# Next steps?
There's a few obvious things that can be done.
1. Latin hypercube sampling for Random Forest parameters.
2. Experimenting with adding more layers, tweaking the layer types etc.
What else might be done? Ping me at ericmajinglong@gmail.com with the subject "neural nets and HA". :)
| github_jupyter |
```
import matplotlib.pyplot as plt
import numpy as np
# if using a jupyter notebook
%matplotlib inline
theta = np.arange(0,np.pi,0.001) # start,stop,step
scale = 1
cos_theta = np.cos(theta)
phi_theta = 3*np.cos(0.5*theta-3/2*np.pi)+1
phi_linear = -(1+2 * np.cos(0.5))/np.pi * theta + np.cos(0.5)
phi_large = -0.876996 * theta + 0.5
phi_zero = -0.876996 * theta
phi_log = -np.log(theta+1)+1
sphereFace = np.cos(1.35 * theta)
arcFace = np.cos(theta+0.5)
cosFace = np.cos(theta)-0.35
cm1 = np.cos(theta+0.3)-0.2
cm2 = np.cos(0.9*theta +0.4) - 0.15
y = 340/(19*np.pi**2) * theta**2 - 397/(19*np.pi) * theta+1
linear_arcface = (np.pi- 2 * theta)/np.pi
li_margin_arcface = (np.pi- 2 * (theta+0.45))/np.pi
plt.plot(theta, cos_theta, label='Softmax $(1.0, 1.0, 0.0, 0.00)$')
#plt.plot(theta, phi_theta, label='Phi_theta $(3.0, 0.5, -4.71, -1)$')
plt.plot(theta, phi_linear, label=r'Phi_linear $\frac{-(1+2*cos(0.5))}{\pi}\theta+cos(0.5)$')
#plt.plot(theta, phi_large, label=r'large_linear')
#plt.plot(theta, phi_zero, label=r'Phi_zero')
#plt.plot(theta, sphereFace, label='SphereFace $(1.0, 1.35, 0.0, 0.00)$')
plt.plot(theta, arcFace, label='ArcFace $(1.0, 1.0, 0.5, 0.00)$')
#plt.plot(theta, cosFace, label='CosFace $(1.0, 1.0, 0.0, 0.35)$')
plt.plot(theta, linear_arcface, label='li-arcface')
plt.plot(theta, li_margin_arcface, label='li_margin_arcface 0.45')
#plt.plot(theta, cm1, label='CM1 (1.0, 1.0, 0.3, 0.20)')
#plt.plot(theta, cm2, label='CM1 (1.0, 0.9, 0.4, 0.15)' )
#plt.plot(theta, y, label='y theta 10')
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.grid(linestyle='--')
plt.show()
start = -4
stop = 4
step = 0.01
def compute_smooth_x(sigma):
sigma_y = 1.0/(sigma**2)
x_neg_1 = np.arange(start, -1*sigma_y, step) # start,stop,step
x_neg_2 = np.arange(-1*sigma_y, 0, step) # start,stop,step
x_pos_1 = np.arange(0, sigma_y, step) # start,stop,step
x_pos_2 = np.arange(sigma_y, stop, step) # start,stop,step
x_center = np.concatenate((x_neg_2, x_pos_1))
smooth_x_center = (sigma * x_center)**2 / 2
smooth_x_left = np.abs(x_neg_1) - 0.5/ (sigma**2)
smooth_x_right = np.abs(x_pos_2) - 0.5/ (sigma**2)
smooth_x = np.concatenate((smooth_x_left, smooth_x_center, smooth_x_right))
x = np.concatenate((x_neg_1, x_neg_2, x_pos_1, x_pos_2))
return x, smooth_x
x = np.arange(start, stop, step) # start,stop,step
square_x = x**2
abs_x = np.abs(x)
x_sigma_1, smooth_x_sigma_1 = compute_smooth_x(1.0)
x_sigma_3, smooth_x_sigma_3 = compute_smooth_x(3.0)
plt.plot(x, square_x, label='$x^2$')
plt.plot(x, abs_x, label='$abs(x)$')
plt.plot(x_sigma_1, smooth_x_sigma_1, label='smooth(x)')
plt.plot(x_sigma_3, smooth_x_sigma_3, label='smooth(x) sigma 3')
plt.plot(x_sigma_3, 4.0*smooth_x_sigma_3, label='smooth(x) sigma 3 scale 4')
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.grid(linestyle='--')
plt.show()
x = np.arange(np.exp(-1), np.exp(1), 0.01) # start,stop,step
y_2 = x**2
y_4 = x**4
y_8 = x**8
y_16 = x**16
y_32 = x**32
y_64 = x**64
plt.plot(x, y_2, label='$x^{2}$')
plt.plot(x, y_4, label='$x^{4}$')
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.grid(linestyle='--')
plt.show()
plt.plot(x, y_8, label='$x^{8}$')
plt.plot(x, y_16, label='$x^{16}$')
plt.plot(x, y_32, label='$x^{32}$')
plt.plot(x, y_64, label='$x^{64}$')
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.grid(linestyle='--')
plt.show()
```
| github_jupyter |
This is a companion notebook for the book [Deep Learning with Python, Second Edition](https://www.manning.com/books/deep-learning-with-python-second-edition?a_aid=keras&a_bid=76564dff). For readability, it only contains runnable code blocks and section titles, and omits everything else in the book: text paragraphs, figures, and pseudocode.
**If you want to be able to follow what's going on, I recommend reading the notebook side by side with your copy of the book.**
This notebook was generated for TensorFlow 2.6.
# Deep learning for text
## Natural Language Processing: the bird's eye view
## Preparing text data
### Text standardization
### Text splitting (tokenization)
### Vocabulary indexing
### Using the `TextVectorization` layer
```
import string
class Vectorizer:
def standardize(self, text):
text = text.lower()
return "".join(char for char in text if char not in string.punctuation)
def tokenize(self, text):
text = self.standardize(text)
return text.split()
def make_vocabulary(self, dataset):
self.vocabulary = {"": 0, "[UNK]": 1}
for text in dataset:
text = self.standardize(text)
tokens = self.tokenize(text)
for token in tokens:
if token not in self.vocabulary:
self.vocabulary[token] = len(self.vocabulary)
self.inverse_vocabulary = dict(
(v, k) for k, v in self.vocabulary.items())
def encode(self, text):
text = self.standardize(text)
tokens = self.tokenize(text)
return [self.vocabulary.get(token, 1) for token in tokens]
def decode(self, int_sequence):
return " ".join(
self.inverse_vocabulary.get(i, "[UNK]") for i in int_sequence)
vectorizer = Vectorizer()
dataset = [
"I write, erase, rewrite",
"Erase again, and then",
"A poppy blooms.",
]
vectorizer.make_vocabulary(dataset)
test_sentence = "I write, rewrite, and still rewrite again"
encoded_sentence = vectorizer.encode(test_sentence)
print(encoded_sentence)
decoded_sentence = vectorizer.decode(encoded_sentence)
print(decoded_sentence)
from tensorflow.keras.layers import TextVectorization
text_vectorization = TextVectorization(
output_mode="int",
)
import re
import string
import tensorflow as tf
def custom_standardization_fn(string_tensor):
lowercase_string = tf.strings.lower(string_tensor)
return tf.strings.regex_replace(
lowercase_string, f"[{re.escape(string.punctuation)}]", "")
def custom_split_fn(string_tensor):
return tf.strings.split(string_tensor)
text_vectorization = TextVectorization(
output_mode="int",
standardize=custom_standardization_fn,
split=custom_split_fn,
)
dataset = [
"I write, erase, rewrite",
"Erase again, and then",
"A poppy blooms.",
]
text_vectorization.adapt(dataset)
```
**Displaying the vocabulary**
```
text_vectorization.get_vocabulary()
vocabulary = text_vectorization.get_vocabulary()
test_sentence = "I write, rewrite, and still rewrite again"
encoded_sentence = text_vectorization(test_sentence)
print(encoded_sentence)
inverse_vocab = dict(enumerate(vocabulary))
decoded_sentence = " ".join(inverse_vocab[int(i)] for i in encoded_sentence)
print(decoded_sentence)
```
## Two approaches for representing groups of words: sets and sequences
### Preparing the IMDB movie reviews data
```
!curl -O https://ai.stanford.edu/~amaas/data/sentiment/aclImdb_v1.tar.gz
!tar -xf aclImdb_v1.tar.gz
!rm -r aclImdb/train/unsup
!cat aclImdb/train/pos/4077_10.txt
import os, pathlib, shutil, random
base_dir = pathlib.Path("aclImdb")
val_dir = base_dir / "val"
train_dir = base_dir / "train"
for category in ("neg", "pos"):
os.makedirs(val_dir / category)
files = os.listdir(train_dir / category)
random.Random(1337).shuffle(files)
num_val_samples = int(0.2 * len(files))
val_files = files[-num_val_samples:]
for fname in val_files:
shutil.move(train_dir / category / fname,
val_dir / category / fname)
from tensorflow import keras
batch_size = 32
train_ds = keras.utils.text_dataset_from_directory(
"aclImdb/train", batch_size=batch_size
)
val_ds = keras.utils.text_dataset_from_directory(
"aclImdb/val", batch_size=batch_size
)
test_ds = keras.utils.text_dataset_from_directory(
"aclImdb/test", batch_size=batch_size
)
```
**Displaying the shapes and dtypes of the first batch**
```
for inputs, targets in train_ds:
print("inputs.shape:", inputs.shape)
print("inputs.dtype:", inputs.dtype)
print("targets.shape:", targets.shape)
print("targets.dtype:", targets.dtype)
print("inputs[0]:", inputs[0])
print("targets[0]:", targets[0])
break
```
### Processing words as a set: the bag-of-words approach
#### Single words (unigrams) with binary encoding
**Preprocessing our datasets with a `TextVectorization` layer**
```
text_vectorization = TextVectorization(
max_tokens=20000,
output_mode="binary",
)
text_only_train_ds = train_ds.map(lambda x, y: x)
text_vectorization.adapt(text_only_train_ds)
binary_1gram_train_ds = train_ds.map(lambda x, y: (text_vectorization(x), y))
binary_1gram_val_ds = val_ds.map(lambda x, y: (text_vectorization(x), y))
binary_1gram_test_ds = test_ds.map(lambda x, y: (text_vectorization(x), y))
```
**Inspecting the output of our binary unigram dataset**
```
for inputs, targets in binary_1gram_train_ds:
print("inputs.shape:", inputs.shape)
print("inputs.dtype:", inputs.dtype)
print("targets.shape:", targets.shape)
print("targets.dtype:", targets.dtype)
print("inputs[0]:", inputs[0])
print("targets[0]:", targets[0])
break
```
**Our model-building utility**
```
from tensorflow import keras
from tensorflow.keras import layers
def get_model(max_tokens=20000, hidden_dim=16):
inputs = keras.Input(shape=(max_tokens,))
x = layers.Dense(hidden_dim, activation="relu")(inputs)
x = layers.Dropout(0.5)(x)
outputs = layers.Dense(1, activation="sigmoid")(x)
model = keras.Model(inputs, outputs)
model.compile(optimizer="rmsprop",
loss="binary_crossentropy",
metrics=["accuracy"])
return model
```
**Training and testing the binary unigram model**
```
model = get_model()
model.summary()
callbacks = [
keras.callbacks.ModelCheckpoint("binary_1gram.keras",
save_best_only=True)
]
model.fit(binary_1gram_train_ds.cache(),
validation_data=binary_1gram_val_ds.cache(),
epochs=10,
callbacks=callbacks)
model = keras.models.load_model("binary_1gram.keras")
print(f"Test acc: {model.evaluate(binary_1gram_test_ds)[1]:.3f}")
```
#### Bigrams with binary encoding
**Configuring the `TextVectorization` layer to return bigrams**
```
text_vectorization = TextVectorization(
ngrams=2,
max_tokens=20000,
output_mode="binary",
)
```
**Training and testing the binary bigram model**
```
text_vectorization.adapt(text_only_train_ds)
binary_2gram_train_ds = train_ds.map(lambda x, y: (text_vectorization(x), y))
binary_2gram_val_ds = val_ds.map(lambda x, y: (text_vectorization(x), y))
binary_2gram_test_ds = test_ds.map(lambda x, y: (text_vectorization(x), y))
model = get_model()
model.summary()
callbacks = [
keras.callbacks.ModelCheckpoint("binary_2gram.keras",
save_best_only=True)
]
model.fit(binary_2gram_train_ds.cache(),
validation_data=binary_2gram_val_ds.cache(),
epochs=10,
callbacks=callbacks)
model = keras.models.load_model("binary_2gram.keras")
print(f"Test acc: {model.evaluate(binary_2gram_test_ds)[1]:.3f}")
```
#### Bigrams with TF-IDF encoding
**Configuring the `TextVectorization` layer to return token counts**
```
text_vectorization = TextVectorization(
ngrams=2,
max_tokens=20000,
output_mode="count"
)
```
**Configuring the `TextVectorization` layer to return TF-IDF-weighted outputs**
```
text_vectorization = TextVectorization(
ngrams=2,
max_tokens=20000,
output_mode="tf_idf",
)
```
**Training and testing the TF-IDF bigram model**
```
text_vectorization.adapt(text_only_train_ds)
tfidf_2gram_train_ds = train_ds.map(lambda x, y: (text_vectorization(x), y))
tfidf_2gram_val_ds = val_ds.map(lambda x, y: (text_vectorization(x), y))
tfidf_2gram_test_ds = test_ds.map(lambda x, y: (text_vectorization(x), y))
model = get_model()
model.summary()
callbacks = [
keras.callbacks.ModelCheckpoint("tfidf_2gram.keras",
save_best_only=True)
]
model.fit(tfidf_2gram_train_ds.cache(),
validation_data=tfidf_2gram_val_ds.cache(),
epochs=10,
callbacks=callbacks)
model = keras.models.load_model("tfidf_2gram.keras")
print(f"Test acc: {model.evaluate(tfidf_2gram_test_ds)[1]:.3f}")
inputs = keras.Input(shape=(1,), dtype="string")
processed_inputs = text_vectorization(inputs)
outputs = model(processed_inputs)
inference_model = keras.Model(inputs, outputs)
import tensorflow as tf
raw_text_data = tf.convert_to_tensor([
["That was an excellent movie, I loved it."],
])
predictions = inference_model(raw_text_data)
print(f"{float(predictions[0] * 100):.2f} percent positive")
```
| github_jupyter |
# 2.1.6 APIによる入手
Yahoo APIを利用してショッピングのレビューコメントを取得
```
# リスト 2.1.20
# Yahoo ショッピングのカテゴリID一覧を取得する
import requests
import json
import time
import csv
# エンドポイント
url_cat = 'https://shopping.yahooapis.jp/ShoppingWebService/V1/json/categorySearch'
# アプリケーションid (p.34の方法で取得した値を設定して下さい)
appid = 'xxxx'
# 全カテゴリファイル
all_categories_file = './all_categories.csv'
# APIリクエスト呼び出し用関数
def r_get(url, dct):
time.sleep(1) # 1回で1秒あける
return requests.get(url, params=dct)
# カテゴリ取得用関数
def get_cats(cat_id):
try:
result = r_get(url_cat, {'appid': appid, 'category_id': cat_id})
cats = result.json()['ResultSet']['0']['Result']['Categories']['Children']
for i, cat in cats.items():
if i != '_container':
yield cat['Id'], {'short': cat['Title']['Short'], 'medium': cat['Title']['Medium'], 'long': cat['Title']['Long']}
except:
pass
# リスト 2.1.21
# カテゴリ一覧CSVファイルの生成
# ヘッダ
output_buffer = [['カテゴリコードlv1', 'カテゴリコードlv2', 'カテゴリコードlv3',
'カテゴリ名lv1', 'カテゴリ名lv2', 'カテゴリ名lv3', 'カテゴリ名lv3_long']]
with open(all_categories_file, 'w') as f:
writer = csv.writer(f, lineterminator='\n')
writer.writerows(output_buffer)
output_buffer = []
# カテゴリレベル1
for id1, title1 in get_cats(1):
print('カテゴリレベル1 :', title1['short'])
try:
# カテゴリレベル2
for id2, title2 in get_cats(id1):
# カテゴリレベル3
for id3, title3 in get_cats(id2):
wk = [id1, id2, id3, title1['short'], title2['short'], title3['short'], title3['long']]
output_buffer.append(wk)
# ファイル書き込み
with open(all_categories_file, 'a') as f:
writer = csv.writer(f, lineterminator='\n')
writer.writerows(output_buffer)
output_buffer = []
except KeyError:
continue
# リスト 2.1.22
# CSVファイルの内容確認
import pandas as pd
from IPython.display import display
df = pd.read_csv(all_categories_file)
display(df.head())
# リスト 2.1.23
# スマホのコード確認
df1 = df.query("カテゴリコードlv3 == '49331'")
display(df1)
# リスト 2.1.24
# レビューコメントの取得
import requests
import time
url_review = 'https://shopping.yahooapis.jp/ShoppingWebService/V1/json/reviewSearch'
# アプリケーションid
# 書籍ではappidは伏せ字にしてください
appid = 'dj0zaiZpPUZCZFh2WjRYM1V1WCZzPWNvbnN1bWVyc2VjcmV0Jng9ZmE-'
# レビュー取得件数。最大50。APIの仕様。
num_results = 50
num_reviews_per_cat = 99999999
# テキストの最大・最小文字数。レビュー本文がこれより長い・短いものは読み飛ばす。
max_len = 10000
min_len = 50
def r_get(url, dct):
time.sleep(1) # 1回で1秒あける
return requests.get(url, params=dct)
# 指定したカテゴリidのレビューを返す
def get_reviews(cat_id, max_items):
# 実際に返した件数
items = 0
# 結果配列
results = []
# 開始位置
start = 1
while (items < max_items):
result = r_get(url_review, {'appid': appid, 'category_id': cat_id, 'results': num_results, 'start': start})
if result.ok:
rs = result.json()['ResultSet']
else:
print('エラーが返されました : [cat id] {} [reason] {}-{}'.format(cat_id, result.status_code, result.reason))
if result.status_code == 400:
print('ステータスコード400(badrequestは中止せず読み飛ばします')
break
else:
exit(True)
avl = int(rs['totalResultsAvailable'])
pos = int(rs['firstResultPosition'])
ret = int(rs['totalResultsReturned'])
#print('総ヒット数: %d 開始位置: %d 取得数: %d' % (avl, pos, ret))
reviews = result.json()['ResultSet']['Result']
for rev in reviews:
desc_len = len(rev['Description'])
if min_len > desc_len or max_len < desc_len:
continue
items += 1
buff = {}
buff['id'] = items
buff['title'] = rev['ReviewTitle'].replace('\n', '').replace(',', '、')
buff['rate'] = int(float(rev['Ratings']['Rate']))
buff['comment'] = rev['Description'].replace('\n', '').replace(',', '、')
buff['name'] = rev['Target']['Name']
buff['code'] = rev['Target']['Code']
results.append(buff)
if items >= max_items:
break
start += ret
#print('有効件数: %d' % items)
return results
# リスト2.1.25
# コメント一覧の取得と保存
import json
import pickle
# get_reviews(code, count) レビューコメントの取得
# code: カテゴリコード (all_categories.csv)に記載のもの
# count: 何件取得するか
result = get_reviews(49331,5)
print(json.dumps(result, indent=2,ensure_ascii=False))
```
| github_jupyter |
<a href="https://colab.research.google.com/github/Blackman9t/Advanced-Data-Science/blob/master/pyspark_fundamentals_2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# IBM intro to spark lab, part 3 and DataCamp intro to Pyspark, lessons 3 and 4.
**This is a very comprehensive notebook on pyspark. It's a sequel to the notebook pyspark_fundamentals_1. This notebook contains tutorials from IBM and DataCamp combined.<br>
This notebook holds the IBM intro to spark tutorials part 3 and the DataCamp intro to pyspark course lessons 3 and 4.<br> A continuation link is provided at the end of this notebook**
First let's load spark dependencies to run in colab
```
!apt-get install openjdk-8-jdk-headless -qq > /dev/null
!wget -q http://apache.osuosl.org/spark/spark-2.4.5/spark-2.4.5-bin-hadoop2.7.tgz
!tar xf spark-2.4.5-bin-hadoop2.7.tgz
!pip install -q findspark
!pip install pyspark
# Set up required environment variables
import os
os.environ["JAVA_HOME"] = "/usr/lib/jvm/java-8-openjdk-amd64"
os.environ["SPARK_HOME"] = "/content/spark-2.4.5-bin-hadoop2.7"
```
Next let's instantiate a SparkContext object to connect to the spark cluster if none exists
```
from pyspark import SparkConf, SparkContext
try:
conf = SparkConf().setMaster('local').setAppName('myApp')
sc = SparkContext(conf=conf)
print('SparkContext Initialised successfully!')
except Exception as e:
print(e)
# Let's see the SparkContext Object
sc
```
Next let's create a SparkSession as our interface to the SparkContect we created above
```
from pyspark.sql import SparkSession
spark = SparkSession.builder.appName('myApp').getOrCreate()
# Let's see the SparkSession
spark
# Let's import other possible libraries we may use
import pandas as pd
import numpy as np
```
## DataCamp, Course 1 Contd..:
**Lesson 3: Getting started with machine learning pipelines**
________________________________________________________________________________
At the core of the pyspark.ml module are the Transformer and Estimator classes. Almost every other class in the module behaves similarly to these two basic classes.
Transformer classes have a .transform() method that takes a DataFrame and returns a new DataFrame; usually the original one with a new column appended. For example, you might use the class Bucketizer to create discrete bins from a continuous feature or the class PCA to reduce the dimensionality of your dataset using principal component analysis.
Estimator classes all implement a .fit() method. These methods also take a DataFrame, but instead of returning another DataFrame they return a model object. This can be something like a StringIndexerModel for including categorical data saved as strings in your models, or a RandomForestModel that uses the random forest algorithm for classification or regression.
First let's grab our Flights, Airports and Planes data sets using Wget and then read these into Spark DataFrames
```
! wget 'https://assets.datacamp.com/production/repositories/1237/datasets/fa47bb54e83abd422831cbd4f441bd30fd18bd15/flights_small.csv'
! wget 'https://assets.datacamp.com/production/repositories/1237/datasets/6e5c4ac2a4799338ba7e13d54ce1fa918da644ba/airports.csv'
! wget 'https://assets.datacamp.com/production/repositories/1237/datasets/231480a2696c55fde829ce76d936596123f12c0c/planes.csv'
flights = spark.read.csv('flights_small.csv', header=True)
flights.show(5)
airports = spark.read.csv('airports.csv',header=True)
airports.show(5)
planes = spark.read.csv('planes.csv',header=True)
planes.show(5)
```
**Join the DataFrames**
In the next two chapters you'll be working to build a model that predicts whether or not a flight will be delayed based on the flights data. This model will also include information about the plane that flew the route, so the first step is to join the two tables: flights and planes!
```
# First, rename the year column of planes to plane_year to avoid duplicate column names.
planes = planes.withColumnRenamed('year','plane_year')
planes.show(5)
# Create a new DataFrame called model_data by joining the flights table with planes using the tailnum column as the key.
model_data = flights.join(planes, on='tailnum', how='left_outer')
model_data.show(5)
# Let's see how many rows exist
model_data.count()
# let's print the schema and see how it's like
model_data.printSchema()
```
**Data types**
Good work! Before you get started modeling, it's important to know that Spark only handles numeric data. That means all of the columns in your DataFrame must be either integers or decimals (called 'doubles' in Spark).
When we imported our data, we let Spark guess what kind of information each column held. Unfortunately, Spark doesn't always guess right and you can see that some of the columns in our DataFrame are strings containing numbers as opposed to actual numeric values.
To remedy this, you can use the .cast() method in combination with the .withColumn() method. It's important to note that .cast() works on columns, while .withColumn() works on DataFrames.
The only argument you need to pass to .cast() is the kind of value you want to create, in string form. For example, to create integers, you'll pass the argument "integer" and for decimal numbers you'll use "double".
You can put this call to .cast() inside a call to .withColumn() to overwrite the already existing column, just like you did in the previous chapter!
**String to integer**
Now you'll use the .cast() method you learned in the previous exercise to convert all the appropriate columns from your DataFrame model_data to integers!
To convert the type of a column using the .cast() method, you can write code like this:
```
dataframe = dataframe.withColumn("col", dataframe.col.cast("new_type"))
```
```
# Use the method .withColumn() to .cast() the following columns to type "integer". Access the columns using the df.col notation:
# model_data.arr_delay
# model_data.air_time
# model_data.month
# model_data.plane_year
model_data = model_data.withColumn("arr_delay", model_data.arr_delay.cast('integer'))
model_data = model_data.withColumn("air_time", model_data.air_time.cast('integer'))
model_data = model_data.withColumn("month", model_data.month.cast('integer'))
model_data = model_data.withColumn("plane_year", model_data.plane_year.cast('integer'))
# Let'see the schema data types again
model_data.printSchema()
```
**Create a new column**
In the last exercise, you converted the column plane_year to an integer. This column holds the year each plane was manufactured. However, your model will use the planes' age, which is slightly different from the year it was made!
Create the column plane_age using the .withColumn() method and subtracting the year of manufacture (column plane_year) from the year (column year) of the flight.
```
model_data = model_data.withColumn('plane_age', model_data.year - model_data.plane_year)
model_data.show(5)
```
**Making a Boolean**
Consider that you're modeling a yes or no question: is the flight late? However, your data contains the arrival delay in minutes for each flight. Thus, you'll need to create a boolean column which indicates whether the flight was late or not!
```
#Use the .withColumn() method to create the column is_late. This column is equal to model_data.arr_delay > 0.
model_data = model_data.withColumn('is_arrival_late', model_data.arr_delay > 0)
model_data.show(3)
# Convert this column to an integer column so that you can use it in your model and name it label
# (this is the default name for the response variable in Spark's machine learning routines).
model_data = model_data.withColumn('label', model_data.is_arrival_late.cast('integer'))
model_data.show(3)
```
**Remove missing values**
```
cols = model_data.columns
for col_name in cols:
model_data = model_data.filter(col_name + ' is not Null')
# Let's see how many rows are left from the filter exercise above
model_data.count()
```
**Strings and factors**
As you know, Spark requires numeric data for modeling. So far this hasn't been an issue; even boolean columns can easily be converted to integers without any trouble. But you'll also be using the airline and the plane's destination as features in your model. These are coded as strings and there isn't any obvious way to convert them to a numeric data type.
Fortunately, PySpark has functions for handling this built into the pyspark.ml.features submodule. You can create what are called 'one-hot vectors' to represent the carrier and the destination of each flight. A one-hot vector is a way of representing a categorical feature where every observation has a vector in which all elements are zero except for at most one element, which has a value of one (1).
Each element in the vector corresponds to a level of the feature, so it's possible to tell what the right level is by seeing which element of the vector is equal to one (1).
The first step to encoding your categorical feature is to create a StringIndexer. Members of this class are Estimators that take a DataFrame with a column of strings and map each unique string to a number. Then, the Estimator returns a Transformer that takes a DataFrame, attaches the mapping to it as metadata, and returns a new DataFrame with a numeric column corresponding to the string column.
The second step is to encode this numeric column as a one-hot vector using a OneHotEncoder. This works exactly the same way as the StringIndexer by creating an Estimator and then a Transformer. The end result is a column that encodes your categorical feature as a vector that's suitable for machine learning routines!
This may seem complicated, but don't worry! All you have to remember is that you need to create a StringIndexer and a OneHotEncoder, and the Pipeline will take care of the rest.
**Carrier column**
In this exercise you'll create a StringIndexer and a OneHotEncoder to code the carrier column. To do this, you'll call the class constructors with the arguments inputCol and outputCol.
The inputCol is the name of the column you want to index or encode, and the outputCol is the name of the new column that the Transformer should create.
```
# Create a StringIndexer called carr_indexer by calling StringIndexer() with inputCol="carrier" and outputCol="carrier_index".
from pyspark.ml.feature import StringIndexer
carr_indexer = StringIndexer(inputCol='carrier', outputCol='carrier_index')
# to immediately see the effect of the StringIndexer, lets say...
indexed = carr_indexer.fit(model_data).transform(model_data) # This creates a new data frame
indexed.show(3)
# Create a OneHotEncoder called carr_encoder by calling OneHotEncoder() with inputCol="carrier_index" and outputCol="carrier_fact".
from pyspark.ml.feature import OneHotEncoder
carr_encoder = OneHotEncoder(inputCol='carrier_index', outputCol='carrier_fact')
# to immediately see the effect of the OneHotEncoder, lets say...
encoded = carr_encoder.transform(indexed) # This creates a new data frame. Note that encoder is a transformer and has no fit method
encoded.show(3)
```
**Destination Column**
Now you'll encode the dest column just like we did the carrier column.
```
# Create a StringIndexer called dest_indexer by calling StringIndexer() with inputCol="dest" and outputCol="dest_index".
dest_indexer = StringIndexer(inputCol='dest', outputCol='dest_index')
# Create a OneHotEncoder called dest_encoder by calling OneHotEncoder() with inputCol="dest_index" and outputCol="dest_fact".
dest_encoder = OneHotEncoder(inputCol='dest_index', outputCol='dest_fact')
```
**Assemble a vector**
The last step in the Pipeline is to combine all of the columns containing our features into a single column. This has to be done before modeling can take place because every Spark modeling routine expects the data to be in this form. You can do this by storing each of the values from a column as an entry in a vector. Then, from the model's point of view, every observation is a vector that contains all of the information about it and a label that tells the modeler what value that observation corresponds to.
Because of this, the pyspark.ml.feature submodule contains a class called VectorAssembler. This Transformer takes all of the columns you specify and combines them into a new vector column.
```
# Create a VectorAssembler by calling VectorAssembler() with the inputCols names as a list and the outputCol name "features".
# The list of columns should be ["month", "air_time", "carrier_fact", "dest_fact", "plane_age"].
from pyspark.ml.feature import VectorAssembler
vec_assembler = VectorAssembler(inputCols=["month", "air_time", "carrier_fact", "dest_fact", "plane_age"], outputCol='features')
```
**Create the pipeline**
You're finally ready to create a Pipeline!
Pipeline is a class in the pyspark.ml module that combines all the Estimators and Transformers that you've already created. This lets you reuse the same modeling process over and over again by wrapping it up in one simple object. Neat, right?
_Import Pipeline from pyspark.ml._
Call the Pipeline() constructor with the keyword argument stages to create a Pipeline called flights_pipe.<br>
stages should be a list holding all the stages you want your data to go through in the pipeline. Here this is just:
```
[dest_indexer, dest_encoder, carr_indexer, carr_encoder, vec_assembler]
```
```
from pyspark.ml import Pipeline
flights_pipe = Pipeline(stages=[dest_indexer, dest_encoder, carr_indexer, carr_encoder, vec_assembler])
```
**Test vs Train**
After you've cleaned your data and gotten it ready for modeling, one of the most important steps is to split the data into a test set and a train set. After that, don't touch your test data until you think you have a good model! As you're building models and forming hypotheses, you can test them on your training data to get an idea of their performance.
Once you've got your favorite model, you can see how well it predicts the new data in your test set. This never-before-seen data will give you a much more realistic idea of your model's performance in the real world when you're trying to predict or classify new data.
In Spark it's important to make sure you split the data after all the transformations. This is because operations like StringIndexer don't always produce the same index even when given the same list of strings.
**Transform the data:**
Hooray, now you're finally ready to pass your data through the Pipeline you created!
```
# Create the DataFrame piped_data by calling the Pipeline methods .fit() and .transform() in a chain.
# Both of these methods take model_data as their only argument.
piped_data = flights_pipe.fit(model_data).transform(model_data)
piped_data.show(3)
```
**Split the data**
Now that you've done all your manipulations, the last step before modeling is to split the data!
Use the DataFrame method .randomSplit() to split piped_data into two pieces, training with 75% of the data, and test with 25% of the data by passing the list [.75, .25] to the .randomSplit() method.
```
training, testing = piped_data.randomSplit([0.75, 0.25])
```
Let's see howmany rows in both training and testing sets
```
print('Training set has {}, while testing set has {} observations. Total is {} observations.'.format(training.count(), testing.count(), (training.count() + testing.count())))
```
## DataCamp, Course 1 Contd...:
**Lesson 4: Model tuning and selection**
________________________________________________________________________________
**What is logistic regression?**
The model you'll be fitting in this chapter is called a logistic regression. This model is very similar to a linear regression, but instead of predicting a numeric variable, it predicts the probability (between 0 and 1) of an event.
To use this as a classification algorithm, all you have to do is assign a cutoff point to these probabilities. If the predicted probability is above the cutoff point, you classify that observation as a 'yes' (in this case, the flight being late), if it's below, you classify it as a 'no'!
You'll tune this model by testing different values for several hyperparameters. A hyperparameter is just a value in the model that's not estimated from the data, but rather is supplied by the user to maximize performance. For this course it's not necessary to understand the mathematics behind all of these values - what's important is that you'll try out a few different choices and pick the best one.
**Create the modeler**
The Estimator you'll be using is a LogisticRegression from the pyspark.ml.classification submodule.
```
# Import the LogisticRegression class from pyspark.ml.classification.
# Create a LogisticRegression called lr by calling LogisticRegression() with no arguments.
from pyspark.ml.classification import LogisticRegression
lr = LogisticRegression()
print(lr)
```
**Cross validation**
In the next few exercises you'll be tuning your logistic regression model using a procedure called k-fold cross validation. This is a method of estimating the model's performance on unseen data (like your test DataFrame).
It works by splitting the training data into a few different partitions. The exact number is up to you, but in this course you'll be using PySpark's default value of three. Once the data is split up, one of the partitions is set aside, and the model is fit to the others. Then the error is measured against the held out partition. This is repeated for each of the partitions, so that every block of data is held out and used as a test set exactly once. Then the error on each of the partitions is averaged. This is called the cross validation error of the model, and is a good estimate of the actual error on the held out data.
You'll be using cross validation to choose the hyperparameters by creating a grid of the possible pairs of values for the two hyperparameters, elasticNetParam and regParam, and using the cross validation error to compare all the different models so you can choose the best one!
Cross validation helps us to estimate the model error on the held-out testing data set
**Create the evaluator**
The first thing you need when doing cross validation for model selection is a way to compare different models. Luckily, the pyspark.ml.evaluation submodule has classes for evaluating different kinds of models. Your model is a binary classification model, so you'll be using the BinaryClassificationEvaluator from the pyspark.ml.evaluation module.
This evaluator calculates the area under the ROC. This is a metric that combines the two kinds of errors a binary classifier can make (false positives and false negatives) into a simple number. You'll learn more about this towards the end of the chapter!
```
# Import the submodule pyspark.ml.evaluation as evals.
# Create evaluator by calling evals.BinaryClassificationEvaluator() with the argument metricName="areaUnderROC".
import pyspark.ml.evaluation as evals
evaluator = evals.BinaryClassificationEvaluator(metricName='areaUnderROC')
```
**Make a grid**
Next, you need to create a grid of values to search over when looking for the optimal hyperparameters. The submodule pyspark.ml.tuning includes a class called ParamGridBuilder that does just that (maybe you're starting to notice a pattern here; PySpark has a submodule for just about everything!).
You'll need to use the .addGrid() and .build() methods to create a grid that you can use for cross validation. The .addGrid() method takes a model parameter (an attribute of the model Estimator, lr, that you created a few exercises ago) and a list of values that you want to try. The .build() method takes no arguments, it just returns the grid that you'll use later.
```
# Import the submodule pyspark.ml.tuning under the alias tune.
import pyspark.ml.tuning as tune
# Call the class constructor ParamGridBuilder() with no arguments. Save this as grid.
grid = tune.ParamGridBuilder()
# Call the .addGrid() method on grid with lr.regParam as the first argument and np.arange(0, .1, .01) as the second argument.
# This second call is a function from the numpy module (imported as np) that creates a list of numbers from 0 to .1, incrementing by .01. Overwrite grid with the result.
# Add the hyperparameter
grid = grid.addGrid(lr.regParam, np.arange(0, .1, .01))
# Update grid again by calling the .addGrid() method a second time create a grid for lr.elasticNetParam that includes only the values [0, 1].
grid = grid.addGrid(lr.elasticNetParam, [0,1])
# Build the grid
grid = grid.build()
```
**Make the validator**
The submodule pyspark.ml.tuning also has a class called CrossValidator for performing cross validation. This Estimator takes the modeler you want to fit, the grid of hyperparameters you created, and the evaluator you want to use to compare your models.
The submodule pyspark.ml.tune has already been imported as tune. You'll create the CrossValidator by passing it the logistic regression Estimator lr, the parameter grid, and the evaluator you created in the previous exercises.
<br>Name this object cv.
```
# Create the CrossValidator
cv = tune.CrossValidator(estimator=lr,
estimatorParamMaps=grid,
evaluator=evaluator)
```
**Fit the model(s)**
You're finally ready to fit the models and select the best one!
```
# Fit cross validation models
models = cv.fit(training)
# Extract the best model
best_lr = models.bestModel
print(best_lr)
# We can also print the coefficient and intercept of the Logistic Regression model by using the following command:
#coefficient of the regression model
coeff = best_lr.coefficients
#X and Y intercept
intrcpt = best_lr.intercept
print ("The coefficient of the model is : %a" %coeff)
print ("The Intercept of the model is : %f" %intrcpt)
```
**Evaluating binary classifiers**
For this course we'll be using a common metric for binary classification algorithms call the AUC, or area under the curve. In this case, the curve is the ROC, or receiver operating curve. The details of what these things actually measure isn't important for this course. All you need to know is that for our purposes, the closer the AUC is to one (1), the better the model is!
**Evaluate the model**
Remember the test data that you set aside waaaaaay back in chapter 3? It's finally time to test your model on it! You can use the same evaluator you made to fit the model.
```
# Use your model to generate predictions by applying best_lr.transform() to the test data. Save this as test_results.
test_results = best_lr.transform(testing)
# Call evaluator.evaluate() on test_results to compute the AUC. Print the output.
print(evaluator.evaluate(test_results))
```
# DataCamp Course 2: Big Data Fundamentals with PySpark.
### Lesson 1: Introduction to Big Data analysis with Spark
**Inspecting The SparkContext**
SparkContext(sc) is an entry point into the world of spark functionality. An entry point is where control is transferred from operating system(os) to the provided program. in simpler terms, it's like a key to your house, without which there is no entry.
<br>Let's inspect some of the attributes of teh SparkContect.
```
# Let's print the SparkContext Version
sc.version
# Let's print the Python Version that sc runs on.
sc.pythonVer
# Let's print the Master. Master is the URL of the Cluster, or 'local' string to run in local mode.
# sc.master returns 'local', meaning the SparkContext acts as a master on a local node, using all available threads on the computer where it is running.
sc.master
```
**Loading Data in Pyspark:**
We can load raw data in pyspark using SparkContext by two distinct means:
<br>1. SparkContext parallelize() method on a list:
```
rdd = sc.parallelize(range(10))
```
<br>2. SparkContext textFile() method on a file:
```
rdd2 = sc.textFile('text.txt')
```
**Loading data in PySpark shell**
In PySpark, we express our computation through operations on distributed collections that are automatically parallelized across the cluster. In the previous exercise, you have seen an example of loading a list as parallelized collections and in this exercise, you'll load the data from a local file in PySpark shell.
```
# Load a local file into PySpark shell
lines = sc.textFile('movie_quotes.txt')
```
**Use of lambda() with map()**
The map() function in Python returns a list of the results after applying the given function to each item of a given iterable (list, tuple etc.). The general syntax of map() function is map(fun, iter). We can also use lambda functions with map(). The general syntax of map() function with lambda() is map(lambda <agument>:<expression>, iter). Refer to slide 5 of video 1.7 for general help of map() function with lambda().
```
my_list = list(range(1,11))
print(my_list)
# Square each item in my_list using map() and lambda().
squrd_list = list(map(lambda x: x**2, my_list))
print(squrd_list)
```
**Use of lambda() with filter():**
Another function that is used extensively in Python is the filter() function. The filter() function in Python takes in a function and a list as arguments. The general syntax of the filter() function is filter(function, list_of_input). Similar to the map(), filter() can be used with lambda() function. The general syntax of the filter() function with lambda() is filter(lambda <argument>:<expression>, list). Refer to slide 6 of video 1.7 for general help of the filter() function with lambda().
```
my_list2 = [10, 21, 31, 40, 51, 60, 72, 80, 93, 101]
# Filter the numbers divisible by 10 from my_list2 using filter() and lambda().
filtered_list2 = list(filter(lambda x: x % 10 == 0, my_list2))
print(filtered_list2)
```
### Lesson 2: Programming in PySpark RDD’s
**RDDs** stand for _Resilient Distributed Dataset_, they are the first class citizen of Apache Spark.<br>It is simply a collection of data distributed across the cluster. RDD is the fundamental and backbone data type in Pyspark.
**Decomposing RDDs:**
Let's look at the different features of RDDs:
1. **_Resilient :_** Means the ability to withstand failures and recompute missing or damaged partitions.
2. **_Distributed :_** Means spanning the jobs across multiple nodes in the cluster for efficient computation.
3. **_Datasets :_** These are a collection of partitioned data e.g Arrays, Tuples, Tables, E.t.c.
**Creating RDDs:**
RDDs are created in 3 different ways:
1. Using the Spark Context .parallelize() method on an iterable like a list or range
```
rdd = sc.parallelize(range(100))
rdd2 = sc.parallelize('hello world')
```
2. creating RDDs from an external files like a text file or CSV file. This is by far the most common way to create RDDs in Pyspark. This method uses the Spark Context's textFile() method. Here we can read a README.md file stored locally on our computer.
```
rdd2 = sc.textFile('README.md')
```
3. Creating RDDs from other RDDs
**Understanding Partitioning in Pyspark**
Understanding how Spark deals with partitions allows one to control parallelism
. A partition in Spark is the division of the large dataset, with each part being stored in multiple locations across the cluster. By default Spark partitions the Data at the time of creating RDDs, based on several factors such as available resources, external datasets e.t.c
However this behavior can be controlled when we create an RDD via textFile() by passing a second argument called minPartitions, which specifies the minimum number of partitions to be created for an RDD.
In the parallelize() method, no need to use the keyword instead just pass the number of partitions you want as the second argument
```
rdd1 = sc.parallelize(range(100), 6)
# Let's see the type of rdd1
type(rdd1)
# let's see the number of partitions in rdd1
rdd1.getNumPartitions()
```
**RDDs from Parallelized collections**
Resilient Distributed Dataset (RDD) is the basic abstraction in Spark. It is an immutable distributed collection of objects. Since RDD is a fundamental and backbone data type in Spark, it is important that you understand how to create it. In this exercise, you'll create your first RDD in PySpark from a collection of words.
```
RDD = sc.parallelize(["Spark", "is", "a", "framework", "for", "Big Data processing"])
RDD.collect()
type(RDD)
```
**RDDs from External Datasets**
PySpark can easily create RDDs from files that are stored in external storage devices such as HDFS (Hadoop Distributed File System), Amazon S3 buckets, etc. However, the most common method of creating RDD's is from files stored in your local file system. This method takes a file path and reads it as a collection of lines. In this exercise, you'll create an RDD from the file path
```
from google.colab import files
uploaded = files.upload()
file_rdd = sc.textFile('movie_quotes.txt')
# Let's see the first 6 items
file_rdd.take(6)
type(file_rdd)
file_rdd.getNumPartitions()
```
**Partitions in your data**
SparkContext's textFile() method takes an optional second argument called minPartitions for specifying the minimum number of partitions. In this exercise, you'll create an RDD named fileRDD_part with 5 partitions and then compare that with fileRDD that you created in the previous
```
fileRDD_part = sc.textFile('movie_quotes.txt', minPartitions=5)
type(fileRDD_part)
fileRDD_part.getNumPartitions()
```
**Overview of Pyspark Operations**
RDDs in Pyspark support two different types of operations:
1. **Transformations :** Transformations are operations on an RDD that return a new RDD
2. **Actions :** Are operations that perform some computation on an RDD.
**RDD Transformations:**
Transformations follow lazy evaluation
<br>Lazy evaluations denote that Spark creates a graph from all the operations we perform on an RDD. And execution of the graph only starts when an action is performed on an RDD. This is called **_lazy evaluation_** in Spark.
**The RDD Transformations we will look at are :**
1. map()
2. filter()
3. flatmap()
4. union()
_map() transformation :_ Takes in a function and applies it to each element in an RDD, with the result of the func being the new value of each element in the resulting RDD.
_filter() transformation :_ Takes in a function and returns an RDD that only has elements that pass the filter condition.
_flatMap() transformation :_ This is like map() but with the exception that it returns multiple values for each element in the source RDD.
_union() transformation :_ returns the union of one RDD with another RDD
```
# Example of flatMap() transformation
rdd = sc.parallelize(['Hello world','How are you?'])
flat_map_rdd = rdd.flatMap(lambda x: x.split(' '))
flat_map_rdd.collect()
# Example of union() transformation...
# Let's define an input RDD to filter other RDDs from
inputRDD = sc.textFile('movie_quotes.txt')
# Next let's filter out 2 other RDDs
fuckRDD = inputRDD.filter(lambda x: 'fuck' in x)
youRDD = inputRDD.filter(lambda x: 'you' in x)
fuckRDD.collect()
youRDD.collect()
# next lets union or join the fuckRDD to the youRDD
combinedRDD = fuckRDD.union(youRDD)
# Let's see it
combinedRDD.collect()
```
**RDD Actions:**
After transformations, we usually need to do some action in the RDD. Remember RDDs are inherently lazy and must be actioned to derive any outputs.
<br>The Actions we would use here are:
1. **collect() :_** returns all elemnets in the RDD
2. **take(N) :_** returns the N number of elements from the RDD
3. **first() :_** returns the first element and is similar to take(1)
4. **count() :_** returns the total number of elements in the RDD
**Map and Collect :**
The main method by which you can manipulate data in PySpark is using map(). The map() transformation takes in a function and applies it to each element in the RDD. It can be used to do any number of things, from fetching the website associated with each URL in our collection to just squaring the numbers. In this simple exercise, you'll use map() transformation to cube each element in an RDD
```
rdd = sc.parallelize(range(0,101,10))
# use map to cube each element in rdd
cubed = rdd.map(lambda x: x**3)
cubed.collect()
```
**Filter and Count:**
The RDD transformation filter() returns a new RDD containing only the elements that satisfy a particular function. It is useful for filtering large datasets based on a keyword. For this exercise, you'll filter out lines containing keyword Spark from fileRDD RDD which consists of lines of text from the README.md file. Next, you'll count the total number of lines containing the keyword Spark and finally print the first 4 lines of the filtered RDD.
```
# Let's grab the spark README.md file using wget
!rm README.md* -f
!wget https://raw.githubusercontent.com/carloapp2/SparkPOT/master/README.md
file_rdd = sc.textFile('README.md')
file_rdd.take(5)
# Create filter() transformation to select only the lines containing the keyword Spark.
fileRDD_filter = file_rdd.filter(lambda x: 'Spark' in x)
fileRDD_filter.collect()
print("The total number of lines with the keyword Spark is", fileRDD_filter.count())
```
**<h3>Pair-RDDs in PySpark:</h3>**
Working with RDDs of key/Value pairs, which are a common data type for many operations in Spark.
<br>Real life datasets are usually Key/Value pairs. Each row is a key that maps to one or more values.
<br>In order to deal with this kind of data set, Pyspark provides a special kind of data structure called _Pair-RDDs._ In Pair-RDDs, the key refers to the identifier, while the value refers to the data.
**Creating Pair-RDDs:-**
Two common ways to create Pair-RDDs
1. From a list of Key/Value tuples
2. From a regular RDD
Irrespective of the method, the first step to create Pair-RDD is getting the data into Key/Value form.
Example 1: Creating a Pair-RDD from a list of tuples
```
# A list of tuples
my_tuple = [('Sam',23),('Tim',46),('May',33),('John', 54)]
# Creating a Pair RDD
pairRDD_tuple = sc.parallelize(my_tuple)
# Let's see the first element
pairRDD_tuple.first()
type(pairRDD_tuple)
```
Example 2: Creating a Pair-RDD from a regular RDD
```
my_list = ['Sam 23','Tim 46','May 33','John 54']
regular_rdd = sc.parallelize(my_list)
regular_rdd.take(1)
# Now let's create a pair RDD drom the regualr_rdd
pairRDD_regular = regular_rdd.map(lambda x: (x.split(' ')[0], int(x.split(' ')[1])))
pairRDD_regular.collect()
```
**Transformations on Pair-RDDs:**
Pair-RDDs are RDDS, so all transformations applicable to regular RDDs are applicable to them.
<Br>Since Pair-RDDs contain tuples, we need to pass functions that operate on Key/Value pairs.
<br>A few special operations are available for this kind, such as:
1. reduceByKey(func): Combines values with the same key.
2. groupByKey(): Group values with the same key.
3. sortByKey(): Return an RDD sorted by the key.
4. join(): Join two Pair-RDDs based on their key.
**reduceByKey() Transformation:**
This is the most popular Pair-RDD transformation, which combines values of the same key using a function.
<br>reduceByKey() runs several parallel operations, one for each key in the dataset. reduceByKey() returns a new RDD consisting of each key and the reduced value for that key.
For example let's combine or add the goals scored for each player in the list of tuples below
```
tup_list = [('Messi',23), ('Ronaldo',34), ('Neymar',22), ('Messi',24)]
# Next we create a regular RDD and chain it to the reduceByKey() transformation
# In one line of code, creating an RDD with total goals scored per player.
tup_list_reduceByKey = sc.parallelize(tup_list).reduceByKey(lambda x,y: x + y)
# Finally display all key-value pairs of the new RDD
tup_list_reduceByKey.collect()
```
**sortByKey() Transformation:**
Sorting of data is important for many applications. We can sort Pair-RDDs as long as there's an order defined in the keys.
<br>This transformation returns an RDD, sorted by keys in ascending or descending order.
Using the previous example, let's sort the last RDD by descending number of goals scored per player.
```
# First let's reverse the RDD making goals scored the key, player name the value
tup_list_reduceByKey_reversed = tup_list_reduceByKey.map(lambda x: (x[1], x[0]))
# Now let's sort it by descending order
tup_list_reduceByKey_reversed.sortByKey(ascending=False).collect()
```
**groupByKey() Transformation:**
This allows us to group the data by key. It groups all the values with the same key in the Pair-RDD. For example grouping all of the airports for a particular country together.
```
# Example of groupByKey() transformation
airports_tups = [('US','JFK'), ('UK','LHR'), ('FR','CDG'), ('US','SFO')]
airports_tups_groupd = sc.parallelize(airports_tups).groupByKey().collect()
for cont, airport in airports_tups_groupd:
print(cont,list(airport))
```
**join() Transformation:**
This joins two Pair-RDDs based on their key
```
# First let's create 3 lists of players, goals and countries
players = ['Messi','Ronaldo','Neymar']
goals_scored = [34, 32, 24]
origin_country = ['Argentina','Portugal','Brazil']
# Next using zip to zip 2 lists per time from above together to form 2 Pair-RDDs
players_goal_rdd = sc.parallelize(list(zip(players,goals_scored)))
players_country_rdd = sc.parallelize(list(zip(players, origin_country)))
# Next let's join the 2 Pair-RDDs based on their key to get an RDD that shows,
# The name as key and goals scored and country as values for each player above.
players_goal_rdd.join(players_country_rdd).collect()
```
| github_jupyter |
```
import warnings
warnings.simplefilter('ignore')
from sklearn import svm
import pickle
import numpy as np
import pandas as pd
from pandas.io.json import json_normalize
import math
from scipy import sparse
from nltk.metrics.agreement import AnnotationTask
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.svm import SVC
from sklearn.naive_bayes import MultinomialNB
from sklearn.dummy import DummyClassifier
from sklearn.preprocessing import LabelEncoder
from sklearn.feature_extraction import DictVectorizer
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.feature_extraction.text import TfidfTransformer
from sklearn.metrics import precision_recall_fscore_support
from sklearn.linear_model import LogisticRegressionCV
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
from sklearn.base import BaseEstimator
from sklearn.base import TransformerMixin
from sklearn.model_selection import GridSearchCV
from sklearn.pipeline import Pipeline
from sklearn.pipeline import FeatureUnion
from sklearn.pipeline import make_pipeline
from sklearn.metrics import classification_report
from sklearn.metrics import accuracy_score
from sklearn.metrics import confusion_matrix
from sklearn.metrics import cohen_kappa_score
from sklearn.externals import joblib
from sklearn.preprocessing import FunctionTransformer
from stop_words import get_stop_words
pd.options.mode.chained_assignment = None
import string
import time
import re
import os
import json
import subprocess
import csv
import gzip
from datetime import datetime
import itertools
import matplotlib.pyplot as plt
# Tokenizers
from nltk.tokenize.casual import TweetTokenizer
# Imbalanced classes
from imblearn.under_sampling import RandomUnderSampler
from imblearn.pipeline import make_pipeline as make_pipeline_imb
from imblearn.metrics import classification_report_imbalanced
from imblearn.over_sampling import SMOTE, ADASYN
import sys
!{sys.executable} -m pip install --upgrade eli5
import eli5
sys.path.append('../../../../external_repositories/word2vec-twitter/')
from word2vecReader import Word2Vec
import krippendorff
```
# Set Notebook Parameters & Define Key Functions
Here we set variables necessary for pre-processing.
Krippendorff's alpha (inter-rater reliability) data fromat looks as follows (from module documentation - https://github.com/grrrr/krippendorff-alpha):
[
{unit1:value, unit2:value, ...}, # coder 1
{unit1:value, unit3:value, ...}, # coder 2
... # more coders
]
or
it is a sequence of (masked) sequences (list, numpy.array, numpy.ma.array, e.g.)
with rows corresponding to coders and columns to items
metric: function calculating the pairwise distance
force_vecmath: force vector math for custom metrics (numpy required)
convert_items: function for the type conversion of items (default: float)
missing_items: indicator for missing items (default: None)
```
## from http://scikit-learn.org/stable/auto_examples/model_selection/plot_confusion_matrix.html#sphx-glr-auto-examples-model-selection-plot-confusion-matrix-py
def plot_confusion_matrix(cm, classes,
normalize=False,
title='Confusion matrix',
cmap=plt.cm.Blues):
## This function prints and plots the confusion matrix.
## Normalization can be applied by setting `normalize=True`.
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
print("Normalized confusion matrix")
else:
print('Confusion matrix, without normalization')
print(cm)
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
fmt = '.2f' if normalize else 'd'
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, format(cm[i, j], fmt),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
## from http://scikit-learn.org/stable/auto_examples/model_selection/plot_confusion_matrix.html#sphx-glr-auto-examples-model-selection-plot-confusion-matrix-py
def show_conf_matrices(y_test, y_predictions):
## compute confusion matrix
cnf_matrix = confusion_matrix(y_test, y_predictions)
np.set_printoptions(precision=2)
## make class labels
y_labels = []
for item in y_test:
if item not in y_labels:
y_labels.append(item)
else:
continue
## plot non-normalized confusion matrix
plt.figure(figsize=(8, 6))
plot_confusion_matrix(cnf_matrix,classes=y_labels,
title='Confusion matrix, without normalization')
## plot normalized confusion matrix
plt.figure(figsize=(8, 6))
plot_confusion_matrix(cnf_matrix, classes=y_labels, normalize=True,
title='Normalized confusion matrix')
plt.show()
def eval_classifier(pipeline, X_test, y_test):
y_pred = pipeline.predict(X_test)
krippendorff_df = pd.DataFrame({'pred': y_pred, 'test': y_test})
krippendorff_df['pred'] = krippendorff_df['pred'].astype('int')
krippendorff_df['test'] = krippendorff_df['test'].astype('int')
print(classification_report(y_test, y_pred) +
"\nCohen's kappa: " + str(cohen_kappa_score(y_test,y_pred)) +
"\nKrippendorff's alpha: " + str(krippendorff.alpha(krippendorff_df)))
show_conf_matrices(y_test, y_pred)
# tokenizer = RegexpTokenizer(r'\w+')
tokenizer = TweetTokenizer(preserve_case=False, strip_handles=True, reduce_len=True) # which tokenizer to use
stop_words_en = get_stop_words('en') # English stoplist
stop_words_sp = get_stop_words('spanish') # Spanish stoplist
stop_words_2 = []
for word in stop_words_en:
stop_words_2.append(word)
for word in stop_words_sp:
stop_words_2.append(word)
def tokenize_func(doc):
## remove URLs
doc = re.sub(r'^(https|http)?:\/\/.*(\r|\n|\b)', '', doc, flags=re.MULTILINE)
## tokenize
tokens = tokenizer.tokenize(doc)
## remove punctuation from each token
table = str.maketrans('', '', string.punctuation)
nopunct_tokens = [w.translate(table) for w in tokens]
for tok in nopunct_tokens:
if 'httptco' in tok:
ind = nopunct_tokens.index(tok)
del(nopunct_tokens[ind])
elif 'httpstco' in tok:
ind = nopunct_tokens.index(tok)
del(nopunct_tokens[ind])
## keep words length 3 or more
long_tokens = [token for token in nopunct_tokens if len(token) > 2]
## remove remaining tokens that are not alphabetic
alpha_tokens = [word for word in long_tokens if word.isalpha()]
## remove stop words from tokens
stopped_tokens = [i for i in alpha_tokens if not i in stop_words_2]
return stopped_tokens
def replace_urls(text):
text_clean = re.sub(r'^(https|http)?:\/\/.*(\r|\n|\b)', '', text, flags=re.MULTILINE)
return text_clean
```
# Get Data
Here we have the call for our datasets. Datasets necesssary for this process include:
'../../../../data/russell/processed_hand_labeled_w2v_both.csv.gz'
produced by notebooks/models/create/feature_extraction/3.0-ams-w2v-featureextraction-pretrained, and
'../../../../data/russell/processed_hand_labeled_w2v_both_liwc.csv.gz'
produced by running the preceding dataset through LIWC2015 software's csv analysis function. We do not include these intermediary datasets in our repository. As such, we comment out code that requires these datasets. However, in the event that you would like to run this commented-out code, these are reproducible using our notebooks/models/create/feature_extraction/ code and LIWC2015's software.
The final dataset resulting from the preprocessing steps which follow can be found at:
'../../../../data/russell/russell_liwc_w2v_both.csv.gz'
```
## this function is necessary to reformat complete dataset (with both w2v and liwc features) with appropriate
## column names because liwc software renames non-liwc columns alphabetically.
def get_data(file_w2v, file_w2v_liwc):
## get dataset with no liwc features
tweets = pd.read_csv(file_w2v,
dtype=str, compression='gzip')
tweets = tweets.drop(['Unnamed: 0'], axis=1)
## get dataset with liwc features
liwc = pd.read_csv(file_w2v_liwc,
compression='gzip', dtype=str)
liwc = liwc.drop(['A'], axis=1)
## rename columns on dataset with liwc features
tweets_header = list(tweets.columns.values)
liwc_header = list(liwc.columns.values)[5:]
new_header = tweets_header+liwc_header
liwc.columns = new_header
## recast dataset with liwc features as full "tweets" df
tweets = liwc
## preprocess text_no_urls field to make sure no na values remain
tweets['text_no_urls'] = tweets['text_no_urls'].fillna(' ')
return tweets
def preprocess_w2v_features(df, original_w2v_column, pretrained_w2v_column):
## turn original w2v features column (now strings) into list
df[original_w2v_column] = [df[:-1]+[df[-1][:-1]]
for df in [df.split()[1:]
for df in list(df[original_w2v_column].values)]]
## turn original w2v features' vectors (now lists of strings) into lists of floats
df[original_w2v_column] = [pd.to_numeric(vect) for vect in list(df[original_w2v_column].values)]
## turn pretrained w2v features column (now strings) into list
df[pretrained_w2v_column] = [df[:-1]+[df[-1][:-1]]
for df in [df.split()[1:]
for df in list(df[pretrained_w2v_column].values)]]
## turn pretrained w2v features' vectors (now lists of strings) into lists of floats
df[pretrained_w2v_column] = [pd.to_numeric(vect) for vect in list(df[pretrained_w2v_column].values)]
return df
def preprocess_liwc_features(df):
## define list of column names to be turned into numeric values
# change back to 55 !!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!
cols_to_numeric = list(tweets.columns.values)[5:]
## convert list of columns to numeric values
for n in cols_to_numeric:
df[n] = df[n].astype(float)
return df
```
### Call Data Functions
```
# tweets = get_data('../../../../data/russell/processed_hand_labeled_w2v_both.csv.gz',
# '../../../../data/russell/processed_hand_labeled_w2v_both_liwc.csv.gz')
# tweets = preprocess_w2v_features(tweets, 'w2v_features_original', 'w2v_features_pretrained')
# tweets = preprocess_liwc_features(tweets)
```
# Separate into Training and Test Sets
```
## if not re-running the above data preprocessing code, load preprocessed dataset here
tweets = pd.read_csv( '../../../../data/russell/russell_liwc_w2v_both.csv.gz',
compression='gzip', dtype=str)
def make_combined_training_docs_all(df):
col_names = list(tweets.columns.values)
features = [col_names[1]]+col_names[3:]
x_train, x_test, y_train, y_test = train_test_split(df[features],
df['new_topic_1'], test_size=0.10)
return x_train, x_test, y_train, y_test
x_train_all, x_test_all, y_train_all, y_test_all = make_combined_training_docs_all(tweets)
```
# Vectorizers
Each of these classes is used to select and vectorize the appropriate features during model training
```
class MeanEmbeddingVectorizer(object):
def __init__(self, word2vec):
self.word2vec = word2vec
# if a text is empty we should return a vector of zeros
# with the same dimensionality as all the other vectors
self.dim = len(word2vec)
def fit(self, X, y):
return self
def transform(self, X):
means = np.array([sum(vector_vals)/len(vector_vals) if pd.isnull(vector_vals).all() == False
else 0
for vector_vals in X])
return means
class ArrayCaster(BaseEstimator, TransformerMixin):
def fit(self, x, y=None):
return self
def transform(self, data):
## get mean of all values in vector to put in place of nan values
col_mean = np.nanmean(data)
col_means = np.full(data.shape, col_mean)
## find indicies that you need to replace
inds = np.where(pd.isnull(data))
## place column means in the indices
data[inds] = np.take(col_means, inds)
return np.transpose(np.matrix(data))
class TextSelector(BaseEstimator, TransformerMixin):
def __init__(self,key):
self.key = key
def fit(self,X,y=None):
return self
def transform(self,X):
X[self.key] = X[self.key].fillna(' ')
return X[self.key]
```
# Build Classifiers
## Logistic Regression Classifier
When attempting to run a logistic regression classifier on the child class (original) topic labeling system Dr. Russell used, receive the following error: "The least populated class in y has only 1 members, which is too few. The minimum number of members in any class cannot be less than n_splits=3." This could also be what is affecting the Naive Bayes performance and SVM. As such, the below logistic regression attempt attempts to predict only the PARENT class in Dr. Russell's data. This parent class was added as an additional column to Dr. Russell's data.
### Logistic Regression with LIWC Features
```
get_liwc_cols = FunctionTransformer(lambda x: x[list(tweets.columns.values)[5:]], validate=False)
%%time
feature_pipeline = Pipeline([
('feature-extraction', FeatureUnion([
('text-vector', Pipeline([
('select', TextSelector(key='text_no_urls')),
('countvec', CountVectorizer(tokenizer=tokenize_func)),
])),
('liwc-features', Pipeline([
('selector', get_liwc_cols),
('tfidf', TfidfTransformer(use_idf = False))
]))
]
)),
('lr', LogisticRegressionCV(multi_class='multinomial', solver='saga', max_iter=100))
])
feature_pipeline.fit(x_train_all, y_train_all)
## uncomment to save model
# joblib.dump(feature_pipeline, '../../../../models/best/lr/lr_multinomial_saga_liwc.pkl')
eval_classifier(feature_pipeline, x_test_all, y_test_all)
```
### Logistic Regression with Original OR Pretrained w2v Features
```
%%time
feature_pipeline = Pipeline([
('feature-extraction', FeatureUnion([
('text-vector', Pipeline([
('select', TextSelector(key='text_no_urls')),
('countvec', CountVectorizer(tokenizer=tokenize_func)),
])),
('w2v-features', Pipeline([
('select', TextSelector(key='w2v_features_original')),
('meanw2v', MeanEmbeddingVectorizer('w2v_features_original')),
('caster', ArrayCaster())
]))
]
)),
('lr', LogisticRegressionCV(multi_class='multinomial', solver='saga', max_iter=100))
])
feature_pipeline.fit(x_train_all, y_train_all)
## uncomment to save model
# joblib.dump(feat_pipe, '../../../../models/best/lr/lr_multinomial_saga_originalw2v.pkl')
eval_classifier(feature_pipeline, x_test_all, y_test_all)
%%time
feature_pipeline = Pipeline([
('feature-extraction', FeatureUnion([
('text-vector', Pipeline([
('select', TextSelector(key='text_no_urls')),
('countvec', CountVectorizer(tokenizer=tokenize_func)),
])),
('w2v-features', Pipeline([
('select', TextSelector(key='w2v_features_pretrained')),
('meanw2v', MeanEmbeddingVectorizer('w2v_features_pretrained')),
('caster', ArrayCaster())
]))
]
)),
('lr', LogisticRegressionCV(multi_class='multinomial', solver='saga', max_iter=100))
])
feature_pipeline.fit(x_train_all, y_train_all)
## uncomment to save model
# joblib.dump(feat_pipe, '../../../../models/best/lr/lr_multinomial_saga_pretrainedw2v.pkl')
eval_classifier(feature_pipeline, x_test_all, y_test_all)
eli5.show_weights(lr, top=20, vec=vec)
```
## SVM Classifier
### Identify Optimal SVM C and Gamma Parameters (GridSearch)
Note that this process can take several hours.
```
# %%time
# param_grid = {'svc__C': [1, 5, 10, 50],
# 'svc__gamma': [0.0001, 0.0005, 0.001, 0.005]}
# grid = GridSearchCV(pipeline, param_grid)
# grid.fit(x_train_all, y_train_all)
# print(grid.best_params_)
```
### SVM with w2v or LIWC Features
The prepopulated kernel (best performing) is 'rbf' here. We also tried 'linear' and 'poly' kernels.
```
%%time
## uncomment features from featureunion as needed to evaluate each feature individually
## or in combination with others
feature_pipeline = Pipeline([
('feature-extraction', FeatureUnion([
('text-vector', Pipeline([
('select', TextSelector(key='text_no_urls')),
('countvec', CountVectorizer(tokenizer=tokenize_func)),
])),
('liwc-features', Pipeline([
('select', get_liwc_cols),
('tfidf', TfidfTransformer(use_idf = False)),
]))
# ('w2v-features-original', Pipeline([
# ('select', TextSelector(key='w2v_features_original')),
# ('meanw2v', MeanEmbeddingVectorizer('w2v_features_original')),
# ('caster', ArrayCaster())
# ])),
# ('w2v-features-pretrained', Pipeline([
# ('select', TextSelector(key='w2v_features_pretrained')),
# ('meanw2v', MeanEmbeddingVectorizer('w2v_features_pretrained')),
# ('caster', ArrayCaster())
# ]))
]
)),
('svm', SVC(kernel='rbf', gamma=0.005, C=50))
])
feature_pipeline.fit(x_train_all, y_train_all)
## uncomment to save model(s)
# joblib.dump(feature_pipeline, '../../../../models/best/svm/svm_rbf_gridoptimized.pkl')
# joblib.dump(feature_pipeline, '../../../../models/best/svm/svm_rbf_gridoptimized_liwc.pkl')
# joblib.dump(feature_pipeline, '../../../../models/best/svm/svm_rbf_gridoptimized_originalw2v.pkl')
# joblib.dump(feature_pipeline, '../../../../models/best/svm/svm_rbf_gridoptimized_pretrainedw2v.pkl')
# joblib.dump(feature_pipeline, '../../../../models/best/svm/svm_rbf_gridoptimized_liwc_pretrainedw2v.pkl')
eval_classifier(feature_pipeline, x_test_all, y_test_all)
```
| github_jupyter |
# Tutorial Goal
This tutorial aims to show how to **configure and run** a **synthetic workload** using the **wlgen module** provided by LISA.
# Configure logging
```
import logging
from conf import LisaLogging
LisaLogging.setup()
# Execute this cell to enabled devlib debugging statements
logging.getLogger('ssh').setLevel(logging.DEBUG)
# Other python modules required by this notebook
import json
import os
```
# Test environment setup
```
# Setup a target configuration
conf = {
# Target is localhost
"platform" : 'linux',
"board" : "juno",
# Login credentials
"host" : "192.168.0.1",
"username" : "root",
"password" : "",
# Binary tools required to run this experiment
# These tools must be present in the tools/ folder for the architecture
"tools" : ['rt-app', 'taskset', 'trace-cmd'],
# Comment the following line to force rt-app calibration on your target
# "rtapp-calib" : {
# "0": 355, "1": 138, "2": 138, "3": 355, "4": 354, "5": 354
# },
# FTrace events end buffer configuration
"ftrace" : {
"events" : [
"sched_switch",
"cpu_frequency"
],
"buffsize" : 10240
},
# Where results are collected
"results_dir" : "WlgenExample",
# Devlib modules we'll need
"modules": ["cpufreq"]
}
# Support to access the remote target
from env import TestEnv
# Initialize a test environment using:
# the provided target configuration (my_target_conf)
# the provided test configuration (my_test_conf)
te = TestEnv(conf)
target = te.target
```
## Workload execution utility
```
def execute(te, wload, res_dir):
logging.info('# Create results folder for this execution')
!mkdir {res_dir}
logging.info('# Setup FTrace')
te.ftrace.start()
logging.info('## Start energy sampling')
te.emeter.reset()
logging.info('### Start RTApp execution')
wload.run(out_dir=res_dir)
logging.info('## Read energy consumption: %s/energy.json', res_dir)
nrg_report = te.emeter.report(out_dir=res_dir)
logging.info('# Stop FTrace')
te.ftrace.stop()
trace_file = os.path.join(res_dir, 'trace.dat')
logging.info('# Save FTrace: %s', trace_file)
te.ftrace.get_trace(trace_file)
logging.info('# Save platform description: %s/platform.json', res_dir)
plt, plt_file = te.platform_dump(res_dir)
logging.info('# Report collected data:')
logging.info(' %s', res_dir)
!ls -la {res_dir}
return nrg_report, plt, plt_file, trace_file
```
# Single task RTApp workload
## 1) creation
```
# Support to configure and run RTApp based workloads
from wlgen import RTA
# Create a new RTApp workload generator using the calibration values
# reported by the TestEnv module
rtapp_name = 'example1'
rtapp = RTA(target, rtapp_name, calibration=te.calibration())
```
## 2) configuration
```
# RTApp configurator for generation of PERIODIC tasks
from wlgen import Periodic
# Configure this RTApp instance to:
rtapp.conf(
# 1. generate a "profile based" set of tasks
kind = 'profile',
# 2. define the "profile" of each task
params = {
# 3. PERIODIC task with
'task_p20': Periodic (
period_ms = 100, # period [ms]
duty_cycle_pct = 20, # duty cycle [%]
duration_s = 5, # duration [s]
delay_s = 0, # start after that delay [s]
sched = { # run as a low-priority SCHED_OTHER task
'policy' : 'OTHER',
'priotity' : 130,
},
cpus = # pinned on first online CPU
str(target.list_online_cpus()[0])
# ADD OTHER PARAMETERS
).get(),
},
);
# Inspect the JSON file used to run the application
with open('./{}_00.json'.format(rtapp_name), 'r') as fh:
rtapp_json = json.load(fh)
logging.info('Generated RTApp JSON file:')
print json.dumps(rtapp_json, indent=4, sort_keys=True)
```
## 3) execution
```
res_dir = os.path.join(te.res_dir, rtapp_name)
nrg_report, plt, plt_file, trace_file = execute(te, rtapp, res_dir)
```
## 4) Check collected data
```
# Dump the energy measured for the LITTLE and big clusters
logging.info('Energy: %s', nrg_report.report_file)
print json.dumps(nrg_report.channels, indent=4, sort_keys=True)
# Dump the platform descriptor, which could be useful for further analysis
# of the generated results
logging.info('Platform description: %s', plt_file)
print json.dumps(plt, indent=4, sort_keys=True)
```
## 5) trace inspection
```
!kernelshark {trace_file} 2>/dev/null
```
# Workload composition using RTApp
## 1) creation
```
# Support to configure and run RTApp based workloads
from wlgen import RTA
# Create a new RTApp workload generator using the calibration values
# reported by the TestEnv module
rtapp_name = 'example2'
rtapp = RTA(target, rtapp_name, calibration=te.calibration())
```
## 2) configuration
```
# RTApp configurator for generation of PERIODIC tasks
from wlgen import Periodic, Ramp
cpus = str(target.bl.bigs_online[0])
# Light workload
light = Periodic(duty_cycle_pct=10, duration_s=1.0, period_ms= 10,
cpus=cpus)
# Ramp workload
ramp = Ramp(start_pct=10, end_pct=90, delta_pct=20, time_s=1, period_ms=50,
cpus=cpus)
# Heavy workload
heavy = Periodic(duty_cycle_pct=90, duration_s=0.1, period_ms=100,
cpus=cpus)
# Composed workload
lrh_task = light + ramp + heavy
# Configure this RTApp instance to:
rtapp.conf(
# 1. generate a "profile based" set of tasks
kind = 'profile',
# 2. define the "profile" of each task
params = {
# 3. PERIODIC task with
'task_ramp': lrh_task.get()
},
);
# Inspect the JSON file used to run the application
with open('./{}_00.json'.format(rtapp_name), 'r') as fh:
rtapp_json = json.load(fh)
logging.info('Generated RTApp JSON file:')
print json.dumps(rtapp_json, indent=4, sort_keys=True)
```
## 3) execution
```
res_dir = os.path.join(te.res_dir, rtapp_name)
nrg_report, plt, plt_file, trace_file = execute(te, rtapp, res_dir)
```
## 4) trace inspection
```
!kernelshark {trace_file} 2>/dev/null
```
# Custom RTApp connfiguration
```
# Support to configure and run RTApp based workloads
from wlgen import RTA
# Create a new RTApp workload generator using the calibration values
# reported by the TestEnv module
rtapp_name = 'example3'
rtapp = RTA(target, rtapp_name, calibration=te.calibration())
# Configure this RTApp to use a custom JSON
rtapp.conf(
# 1. generate a "custom" set of tasks
kind = 'custom',
# 2. define the "profile" of each task
params = "../../assets/mp3-short.json",
# In this case only few values of the orignal JSON can be tuned:
# DURATION : maximum duration of the workload [s]
# PVALUE : calibration value
# LOGDIR : folder used for generated logs
# WORKDIR : working directory on target
# 3. defined a maximum duration for that workload
duration = 5,
);
res_dir = os.path.join(te.res_dir, rtapp_name)
nrg_report, plt, plt_file, trace_file = execute(te, rtapp, res_dir)
# Inspect the JSON file used to run the application
with open('./{}_00.json'.format(rtapp_name), 'r') as fh:
rtapp_json = json.load(fh)
logging.info('Generated RTApp JSON file:')
print json.dumps(rtapp_json, indent=4, sort_keys=True)
```
# Running Hackbench
## 1) creation and configuration
```
# Support to configure and run RTApp based workloads
from wlgen import PerfMessaging
# Create a "perf bench sched messages" (i.e. hackbench) workload
perf_name = 'hackbench'
perf = PerfMessaging(target, perf_name)
perf.conf(group=1, loop=100, pipe=True, thread=True)
```
## 2) execution
```
res_dir = os.path.join(te.res_dir, perf_name)
nrg_report, plt, plt_file, trace_file = execute(te, perf, res_dir)
```
## 3) explore the performance report
```
# Inspect the generated performance report
perf_file = os.path.join(te.res_dir, perf_name, 'performance.json')
with open(perf_file, 'r') as fh:
perf_json = json.load(fh)
logging.info('Generated performance JSON file:')
print json.dumps(perf_json, indent=4, sort_keys=True)
```
## 4) trace inspection
```
!kernelshark {trace_file} 2>/dev/null
```
| github_jupyter |
```
# Load Libraries - Make sure to run this cell!
import pandas as pd
import numpy as np
import re
from collections import Counter
from sklearn import feature_extraction, tree, model_selection, metrics
from yellowbrick.features import Rank2D
from yellowbrick.features import RadViz
from yellowbrick.features import ParallelCoordinates
import seaborn as sns
import matplotlib.pyplot as plt
import matplotlib
%matplotlib inline
```
## Worksheet - Answer - DGA Detection using Machine Learning
This worksheet is a step-by-step guide on how to detect domains that were generated using "Domain Generation Algorithm" (DGA). We will walk you through the process of transforming raw domain strings to Machine Learning features and creating a decision tree classifer which you will use to determine whether a given domain is legit or not. Once you have implemented the classifier, the worksheet will walk you through evaluating your model.
Overview 2 main steps:
1. **Feature Engineering** - from raw domain strings to numeric Machine Learning features using DataFrame manipulations
2. **Machine Learning Classification** - predict whether a domain is legit or not using a Decision Tree Classifier
**DGA - Background**
"Various families of malware use domain generation
algorithms (DGAs) to generate a large number of pseudo-random
domain names to connect to a command and control (C2) server.
In order to block DGA C2 traffic, security organizations must
first discover the algorithm by reverse engineering malware
samples, then generate a list of domains for a given seed. The
domains are then either preregistered, sink-holed or published
in a DNS blacklist. This process is not only tedious, but can
be readily circumvented by malware authors. An alternative
approach to stop malware from using DGAs is to intercept DNS
queries on a network and predict whether domains are DGA
generated. Much of the previous work in DGA detection is based
on finding groupings of like domains and using their statistical
properties to determine if they are DGA generated. However,
these techniques are run over large time windows and cannot be
used for real-time detection and prevention. In addition, many of
these techniques also use contextual information such as passive
DNS and aggregations of all NXDomains throughout a network.
Such requirements are not only costly to integrate, they may not
be possible due to real-world constraints of many systems (such
as endpoint detection). An alternative to these systems is a much
harder problem: detect DGA generation on a per domain basis
with no information except for the domain name. Previous work
to solve this harder problem exhibits poor performance and many
of these systems rely heavily on manual creation of features;
a time consuming process that can easily be circumvented by
malware authors..."
[Citation: Woodbridge et. al 2016: "Predicting Domain Generation Algorithms with Long Short-Term Memory Networks"]
A better alternative for real-world deployment would be to use "featureless deep learning" - We have a separate notebook where you can see how this can be implemented!
**However, let's learn the basics first!!!**
## Worksheet for Part 1 - Feature Engineering
```
## Load data
df = pd.read_csv('../../data/dga_data_small.csv')
df.drop(['host', 'subclass'], axis=1, inplace=True)
print(df.shape)
df.sample(n=5).head() # print a random sample of the DataFrame
df[df.isDGA == 'legit'].head()
# Google's 10000 most common english words will be needed to derive a feature called ngrams...
# therefore we already load them here.
top_en_words = pd.read_csv('../../data/google-10000-english.txt', header=None, names=['words'])
top_en_words.sample(n=5).head()
# Source: https://github.com/first20hours/google-10000-english
```
## Part 1 - Feature Engineering
Option 1 to derive Machine Learning features is to manually hand-craft useful contextual information of the domain string. An alternative approach (not covered in this notebook) is "Featureless Deep Learning", where an embedding layer takes care of deriving features - a huge step towards more "AI".
Previous academic research has focused on the following features that are based on contextual information:
**List of features**:
1. Length ["length"]
2. Number of digits ["digits"]
3. Entropy ["entropy"] - use ```H_entropy``` function provided
4. Vowel to consonant ratio ["vowel-cons"] - use ```vowel_consonant_ratio``` function provided
5. N-grams ["n-grams"] - use ```ngram``` functions provided
**Tasks**:
Split into A and B parts, see below...
Please run the following function cell and then continue reading the next markdown cell with more details on how to derive those features. Have fun!
```
def H_entropy (x):
# Calculate Shannon Entropy
prob = [ float(x.count(c)) / len(x) for c in dict.fromkeys(list(x)) ]
H = - sum([ p * np.log2(p) for p in prob ])
return H
def vowel_consonant_ratio (x):
# Calculate vowel to consonant ratio
x = x.lower()
vowels_pattern = re.compile('([aeiou])')
consonants_pattern = re.compile('([b-df-hj-np-tv-z])')
vowels = re.findall(vowels_pattern, x)
consonants = re.findall(consonants_pattern, x)
try:
ratio = len(vowels) / len(consonants)
except: # catch zero devision exception
ratio = 0
return ratio
```
### Tasks - A - Feature Engineering
Please try to derive a new pandas 2D DataFrame with a new column for each of feature. Focus on ```length```, ```digits```, ```entropy``` and ```vowel-cons``` here. Also make sure to encode the ```isDGA``` column as integers. [pandas.Series.str](http://pandas.pydata.org/pandas-docs/stable/generated/pandas.Series.str.html), [pandas.Series.replace](http://pandas.pydata.org/pandas-docs/stable/generated/pandas.Series.replace.html) and [pandas.Series,apply](http://pandas.pydata.org/pandas-docs/stable/generated/pandas.Series.apply.html) can be very helpful to quickly derive those features. Functions you need to apply here are provided in above cell.
The ```ngram``` is a bit more complicated, see next instruction cell to add this feature...
```
# derive features
df['length'] = df.domain.str.len()
df['digits'] = df.domain.str.count('[0-9]')
df['entropy'] = df.domain.apply(H_entropy)
df['vowel-cons'] = df.domain.apply(vowel_consonant_ratio)
# encode strings of target variable as integers
df.isDGA = df.isDGA.replace(to_replace = 'dga', value=1)
df.isDGA = df.isDGA.replace(to_replace = 'legit', value=0)
print(df.isDGA.value_counts())
# check intermediate 2D pandas DataFrame
df.sample(n=5).head()
```
### Tasks - B - Feature Engineering
Finally, let's tackle the **ngram** feature. There are multiple steps involved to derive this feature. Here in this notebook, we use an implementation outlined in the this academic paper [Schiavoni 2014: "Phoenix: DGA-based Botnet Tracking and Intelligence" - see section: Linguistic Features](http://s2lab.isg.rhul.ac.uk/papers/files/dimva2014.pdf).
- **What are ngrams???** Imagine a string like 'facebook', if I were to derive all n-grams for n=2 (aka bi-grams) I would get '['fa', 'ac', 'ce', 'eb', 'bo', 'oo', 'ok']', so you see that you slide with one step from the left and just group 2 characters together each time, a tri-gram for 'facebook' would yielfd '['fac', 'ace', 'ceb', 'ebo', 'boo', 'ook']'. Ngrams have a long history in natural language processing, but are also used a lot for example in detecting malicious executable (raw byte ngrams in this case).
Steps involved:
1. We have the 10000 most common english words (see data file we loaded, we call this DataFrame ```top_en_words``` in this notebook). Now we run the ```ngrams``` functions on a list of all these words. The output here is a list that contains ALL 1-grams, bi-grams and tri-grams of these 10000 most common english words.
2. We use the ```Counter``` function from collections to derive a dictionary ```d``` that contains the counts of all unique 1-grams, bi-grams and tri-grams.
3. Our ```ngram_feature``` function will do the core magic. It takes your domain as input, splits it into ngrams (n is a function parameter) and then looks up these ngrams in the english dictionary ```d``` we derived in step 2. Function returns the normalized sum of all ngrams that were contained in the english dictionary. For example, running ```ngram_feature('facebook', d, 2)``` will return 171.28 (this value is just like the one published in the Schiavoni paper).
4. Finally ```average_ngram_feature``` wraps around ```ngram_feature```. You will use this function as your task is to derive a feature that gives the average of the ngram_feature for n=1,2 and 3. Input to this function should be a simple list with entries calling ```ngram_feature``` with n=1,2 and 3, hence a list of 3 ngram_feature results.
5. **YOUR TURN: Apply ```average_ngram_feature``` to you domain column in the DataFrame thereby adding ```ngram``` to the df.**
6. **YOUR TURN: Finally drop the ```domain``` column from your DataFrame**.
Please run the following function cell and then write your code in the following cell.
```
# ngrams: Implementation according to Schiavoni 2014: "Phoenix: DGA-based Botnet Tracking and Intelligence"
# http://s2lab.isg.rhul.ac.uk/papers/files/dimva2014.pdf
def ngrams(word, n):
# Extract all ngrams and return a regular Python list
# Input word: can be a simple string or a list of strings
# Input n: Can be one integer or a list of integers
# if you want to extract multipe ngrams and have them all in one list
l_ngrams = []
if isinstance(word, list):
for w in word:
if isinstance(n, list):
for curr_n in n:
ngrams = [w[i:i+curr_n] for i in range(0,len(w)-curr_n+1)]
l_ngrams.extend(ngrams)
else:
ngrams = [w[i:i+n] for i in range(0,len(w)-n+1)]
l_ngrams.extend(ngrams)
else:
if isinstance(n, list):
for curr_n in n:
ngrams = [word[i:i+curr_n] for i in range(0,len(word)-curr_n+1)]
l_ngrams.extend(ngrams)
else:
ngrams = [word[i:i+n] for i in range(0,len(word)-n+1)]
l_ngrams.extend(ngrams)
# print(l_ngrams)
return l_ngrams
def ngram_feature(domain, d, n):
# Input is your domain string or list of domain strings
# a dictionary object d that contains the count for most common english words
# finally you n either as int list or simple int defining the ngram length
# Core magic: Looks up domain ngrams in english dictionary ngrams and sums up the
# respective english dictionary counts for the respective domain ngram
# sum is normalized
l_ngrams = ngrams(domain, n)
# print(l_ngrams)
count_sum=0
for ngram in l_ngrams:
if d[ngram]:
count_sum+=d[ngram]
try:
feature = count_sum/(len(domain)-n+1)
except:
feature = 0
return feature
def average_ngram_feature(l_ngram_feature):
# input is a list of calls to ngram_feature(domain, d, n)
# usually you would use various n values, like 1,2,3...
return sum(l_ngram_feature)/len(l_ngram_feature)
l_en_ngrams = ngrams(list(top_en_words['words']), [1,2,3])
d = Counter(l_en_ngrams)
from six.moves import cPickle as pickle
with open('../../data/d_common_en_words' + '.pickle', 'wb') as f:
pickle.dump(d, f, pickle.HIGHEST_PROTOCOL)
df['ngrams'] = df.domain.apply(lambda x: average_ngram_feature([ngram_feature(x, d, 1),
ngram_feature(x, d, 2),
ngram_feature(x, d, 3)]))
# check final 2D pandas DataFrame containing all final features and the target vector isDGA
df.sample(n=5).head()
df_final = df
df_final = df_final.drop(['domain'], axis=1)
df_final.to_csv('../../data/dga_features_final_df.csv', index=False)
df_final.head()
```
#### Breakpoint: Load Features and Labels
If you got stuck in Part 1, please simply load the feature matrix we prepared for you, so you can move on to Part 2 and train a Decision Tree Classifier.
```
df_final = pd.read_csv('../../data/dga_features_final_df.csv')
print(df_final.isDGA.value_counts())
df_final.head()
```
### Visualizing the Results
At this point, we've created a dataset which has many features that can be used for classification. Using YellowBrick, your final step is to visualize the features to see which will be of value and which will not.
First, let's create a Rank2D visualizer to compute the correlations between all the features. Detailed documentation available here: http://www.scikit-yb.org/en/latest/examples/methods.html#feature-analysis
```
feature_names = ['length','digits','entropy','vowel-cons','ngrams']
features = df_final[feature_names]
target = df_final.isDGA
visualizer = Rank2D(algorithm='pearson',features=feature_names)
visualizer.fit_transform( features )
visualizer.poof()
```
Now let's use a Seaborn pairplot as well. This will really show you which features have clear dividing lines between the classes. Docs are available here: http://seaborn.pydata.org/generated/seaborn.pairplot.html
```
sns.pairplot(df_final, hue='isDGA', vars=feature_names)
```
Finally, let's try making a RadViz of the features. This visualization will help us see whether there is too much noise to make accurate classifications.
```
X = df_final[feature_names].as_matrix()
y = df_final.isDGA.as_matrix()
radvizualizer = RadViz(classes=['Benign','isDga'], features=feature_names)
radvizualizer.fit_transform( X, y)
radvizualizer.poof()
```
| github_jupyter |
```
# default_exp sklearn_lda
```
# sklearn.LatentDirichletAllocation
> Run LDA Model using 'sklearn'
Run LDA Model using 'sklearn'.
## Preprocessing
```
#export
import pandas as pd
#export
import jieba
#export
def make_df(csv_name, column = 'Content', output_column = 'text'):
'''Use jieba, create data frame.'''
df = pd.read_csv(csv_name)
df = df.dropna(subset=[column])
df[output_column] = df[column].apply(lambda x: " ".join(jieba.cut(x)))
return df
affirmative = make_df("data/affirmative.csv")
affirmative.head()
negative = make_df("data/negative.csv")
negative.head()
```
## LDA
```
#export
import pyLDAvis
import pyLDAvis.sklearn
#export
pyLDAvis.enable_notebook()
#export
import pandas as pd
from sklearn.feature_extraction.text import CountVectorizer
import jieba
from sklearn.decomposition import LatentDirichletAllocation
def chinese_word_cut(mytext):
return " ".join(jieba.cut(mytext))
def print_top_words(model, feature_names, n_top_words):
for topic_idx, topic in enumerate(model.components_):
print("Topic #%d:" % topic_idx)
print(" ".join([feature_names[i]
for i in topic.argsort()[:-n_top_words - 1:-1]]))
#export
def get_custom_stopwords(stop_words_file, encoding = 'utf-8'):
with open(stop_words_file, encoding = encoding) as f:
stopwords = f.read()
stopwords_list = stopwords.split('\n')
custom_stopwords_list = [i for i in stopwords_list]
return custom_stopwords_list
stopwords = get_custom_stopwords("data/stopwords.txt", encoding='utf-8') # HIT停用词词典
max_df = 0.9 # 在超过这一比例的文档中出现的关键词(过于平凡),去除掉。
min_df = 5 # 在低于这一数量的文档中出现的关键词(过于独特),去除掉。
n_features = 1000 # 最大提取特征数量
n_top_words = 20 # 显示主题下关键词的时候,显示多少个
col_content = "text" # 说明其中的文本信息所在列名称
#export
def lda_on_chinese_articles_with_param(df, n_topics,
col_content,
stopwords,
n_features,
max_df,
min_df,
n_top_words):
articles_cutted = df[col_content].apply(chinese_word_cut)
vect = CountVectorizer(max_df = max_df,
min_df = min_df,
token_pattern=u'(?u)\\b[^\\d\\W]\\w+\\b',
stop_words=frozenset(stopwords))
tf = vect.fit_transform(articles_cutted)
lda = LatentDirichletAllocation(n_components=n_topics, max_iter=50,
learning_method='online',
learning_offset=50,
random_state=0)
lda.fit(tf)
print_top_words(lda, vect.get_feature_names(), n_top_words)
return lda, tf, vect
#export
def lda_on_chinese_articles(df, n_topics):
return lda_on_chinese_articles_with_param(df, n_topics,
col_content = col_content,
stopwords = stopwords,
n_features = n_features,
max_df = max_df,
min_df = min_df,
n_top_words = n_top_words)
lda, tf, vect = lda_on_chinese_articles(df = affirmative, n_topics = 3)
pyLDAvis.sklearn.prepare(lda, tf, vect)
```
```python
TypeError: __init__() got an unexpected keyword argument 'n_topics'
```
一般出现这种问题都是程序中字母写错、漏写之类的问题
————————————————
版权声明:本文为CSDN博主「zhuimengshaonian66」的原创文章,遵循 CC 4.0 BY-SA 版权协议,转载请附上原文出处链接及本声明。
原文链接:https://blog.csdn.net/zhuimengshaonian66/article/details/81700959
`n_components` 参数名称修改了。
```
lda, tf, vect = lda_on_chinese_articles(df = negative, n_topics = 3)
pyLDAvis.sklearn.prepare(lda, tf, vect)
pyLDAvis.sklearn.prepare(lda, tf, vect)
```
参考 https://github.com/bmabey/pyLDAvis/issues/132
```python
D:\install\miniconda\lib\site-packages\pyLDAvis\_prepare.py:257: FutureWarning: Sorting because non-concatenation axis is not aligned. A future version
of pandas will change to not sort by default.
To accept the future behavior, pass 'sort=False'.
To retain the current behavior and silence the warning, pass 'sort=True'.
return pd.concat([default_term_info] + list(topic_dfs))
```
重新安装后依然没有解决。
```
pyLDAvis.__version__
pd.__version__
# !pip install pyldavis
import pickle as pkl
with open("model/sklearn-lda.pkl", 'wb') as fp:
pkl.dump(lda, fp)
with open("model/sklearn-lda.pkl", 'rb') as fp:
model0 = pkl.load(fp)
print(model0.__class__)
```
| github_jupyter |
# Learning Objectives
- [ ] 2.2.1 Understand the different types: integer `int`, real `float`, char `chr`, string `str` and Boolean `Boolean` and initialise arrays `list`, `tuple` (1-dimensional and 2-dimensional).
# 3 Basic Data Structures
In computer science, a data structure is a data organization, management, and storage format that enables efficient access and modification. More precisely, a data structure is a collection of data values, the relationships among them, and the functions or operations that can be applied to the data.
## 3.1 Array
An array, is a data structure consisting of a collection of elements (values or variables), each identified by at least one array index or key. We will focus on two commonly used ones in Python.
### 3.1.1 List
**Lists** are the most commonly used data structure in Python.
* It is a **mutable** collection, i.e. its items can be added and removed.
* Each of these data can be accessed by calling its index value.
#### 3.1.1.1 Creating a list
Lists are declared/created by just equating a variable to `[ ]` or list.
>```
>empty_list = []
>print(type(empty_list))
>```
Items in the list are seperated by comma `,`.
>```
>nums = [1, 2, 3, 4]
>nums
>```
```
# YOUR CODE HERE
```
List element can be of any data type.
>```
>fruits = ['apple', 'banana', 'cherry', 'durian']
>fruits
>```
```
# YOUR CODE HERE
```
In fact, it is able to hold elements of *mixed data types*, although this is not commonly used.
>```
>mixed = ['apple', 3, 'banana', 5.0, True, None, (1,), [1,23]]
>mixed
>```
```
# YOUR CODE HERE
```
List can also have **lists as its element**, which creates a `nested list`.
>```
>nested = [ [10, 11, 12, 13],
> [20, 21, 22, 23] ]
>nested
>```
```
# YOUR CODE HERE
```
#### 3.1.1.2 Accessing Elements in a list via Indexing
Items in collection can be accessed by their indexes. Python uses zero-based indexing, i.e. index starts from 0.
>```
>print(fruits)
>print(fruits[0])
>print(fruits[1])
>```
```
# YOUR CODE HERE
```
Indexing can also be done in reverse order by using a negative value as the index. That is the last element of the list has an index of -1, and second last element has index of -2 etc. This is called **negative indexing**.
<center><img src="./images/list-indexing.png" alt="Set Venn Diagram" style="width: 400px;"/></center>
>```
>fruits[-1]
>fruits[-2]
>```
```
# YOUR CODE HERE
```
For nested list, we can access items by **multi-level indexing**. Each level of the index always starts from 0.
For example, access 1st element in 1st list, and 2nd element in 2nd list
>```python
>print(nested)
>print(nested[0][0])
>print(nested[1][1])
>```
#### Exercise 3.1
How do you access element `Blackcurrant` in following list?
>```
>nested_fruits = [
> ['Apple', 'Apricots', 'Avocado'],
> ['Banana', 'Blackcurrant', 'Blueberries'],
> ['Cherries', 'Cranberries', 'Custard-Apple']]
>
>```
```
# YOUR CODE HERE
```
#### 3.1.1.2 Accessing Subsets Elements in a list via Slicing
**Indexing** was only limited to accessing a single element.
**Slicing** on the other hand is accessing a sequence of data inside the list.
**Slicing** is done by defining the index values of the `first element` and the `last element` from the parent list that is required in the sliced list.
>```
>sub = num[a : b]
>sub = num[a : ]
>sub = num[: b]
>sub = num[:]
>```
* if both `a` and `b` are specified, `a` is the first index, `b` is the **last index + 1**.
* if `b` is omitted, it will slice till last element.
* if `a` is omitted, it will starts from first element.
* if neither `a` or `b` is specified, it is effectively copy the whole list
**Note: the upper bound index is NOT inclusive!**
#### Exercise 3.2
* Create a list contain number 0-9
* Print 3rd to 5th items
* Print all items after 6th position
>```
>num = [0,1,2,3,4,5,6,7,8,9]
>```
```
# YOUR CODE HERE
```
#### Exercise 3.3
The `num` is a list of integers from 0 to 9, split the list into 2 equal size sub list, `sub1` and `sub2`.
>```
>num = [0,1,2,3,4,5,6,7,8,9]
>
>```
```
# YOUR CODE HERE
print(sub1)
print(sub2)
```
Remember list items can be accessed using `negative index`. Same technique can be applied for slicing too, i.e. we can **slice with negative index**.
* Last item has index of -1
#### Exercise 3.4
For a list with integer 0-9,
* How to get last 3 items from a list?
* How to ignore last 3 items from a list?
* How to strip first and last items from a list?
>```
>num = [0,1,2,3,4,5,6,7,8,9]
>
>
>
>
>
>```
```
# YOUR CODE HERE
```
#### 3.1.1.3 Working with Lists
##### 3.1.1.3.1 Length of lists `len()`
To find the length of the list or the number of elements in a list, the function `len( )` is used. Syntax is
>```
>len(your_list)
>```
#### Exercise 3.5
Find the length of list `num = [0,1,2,3,4,5,6,7,8,9]`
```
# YOUR CODE HERE
```
##### 3.1.1.3.2 Maximum `max()`, minimum `min()` and sum `sum()` of all the values in the list
If the list consists of all integer elements, the functions `min( )`, `max( )` and `sum()` gives the minimum item, maximum item and total sum value of the list. Syntax is
>```
>max(your_list)
>min(your_list)
>sum(your_list)
>```
#### Exercise 3.5
Find the minimum value, max value and sum of all the values of list `num = [0,1,2,3,4,5,6,7,8,9]`
```
# YOUR CODE HERE
```
#### Exercise 3.6
Find the maximum, minimum and sum of all the values of list `num = [0,1,2,3,4,5,6,7,8,9]` and use `.format()` string method to print out the following message.
>```
>min = 0, max = 9, sum = 45
>```
```
# YOUR CODE HERE
```
For a list with elements as string, the `max( )` and `min( )` is still applicable.
* `max( )` would return a string element whose ASCII value is the highest
* `min( )` is used to return the lowest
Note that only the first index of each element is considered each time and if they value is the same then second index considered so on and so forth.
#### Exercise 3.7
What is the output of `min()` and `max()` on following list? What would happen if `sum()` is applied?
```
jc = ['tmjc','vjc','tjc','yijc','acjc','njc','hci','asrjc','jpjc','rjc','cjc','mi','dhs','ejc','sajc','nyjc']
```
```
# YOUR CODE HERE
```
##### 3.1.1.3.3 Reversing the values in a list with `reversed()` function
The entire elements present in the list can be reversed by using the `reversed()` function.
#### Exercise 3.8
Modify following list by arranging items in reverse order.
```
jc = ['tmjc','vjc','tjc','yijc''acjc','njc','hci','asrjc','jpjc','rjc','cjc','mi','dhs','ejc','sajc','nyjc']
```
```
# YOUR CODE HERE
```
#### Exercise 3.9
Can you print out the items in reverse order without modifying the list?
* Hint: use indexing with step `-1`
```
jc = ['tmjc','vjc','tjc','yijc''acjc','njc','hci','asrjc','jpjc','rjc','cjc','mi','dhs','ejc','sajc','nyjc']
```
```
# YOUR CODE HERE
```
##### 3.1.1.3.3 Sorting elements in a list with `sorted()` function
Python offers built in operation `sorted( )` to arrange the elements in **ascending** order. Syntax is
>```
>sorted(your_list)
>```
For **descending** order, specify the named argument `reverse = True`. Syntax is
>```
>sorted(your_list, reverse = True)
>```
By default the reverse condition will be `False` for reverse. Hence changing it to `True` would arrange the elements in descending order.
#### Exercise 3.10
* Check out the documentation of `list.sort()` method
* Sort following list in <u>ascending</u> order, and then in <u>descending</u> order
```
jc = ['tmjc','vjc','tjc','yijc''acjc','njc','hci','asrjc','jpjc','rjc','cjc','mi','dhs','ejc','sajc','nyjc']
```
```
# YOUR CODE HERE
```
#### Exercise 3.11
* What's the difference between `list.sort()` method and `sorted()` function?
* Write code to illustrate the difference.
```
# YOUR CODE HERE
```
The `sorted()` function has another named argument `key`, which allows us to specify a callable function to adjust our sorting criteria. Syntax is
>```
>sorted(your_list, key = your_function)
>```
The sorting will be done based on returned value from this callable function.
#### Exercise 3.12
For following list of items, sort them by number of characters in each item.
* *Hint:* The `len()` function returns length of a string. To sort based on string length, `key = len` can be specified as shown.
```
names = ['duck', 'chicken', 'goose']
```
```
# YOUR CODE HERE
```
#### Exercise 3.13
For a list `[-1, 5, -30, -10, 2, 20, -3]`, sort the list in descending order by their absolute value.
* *Hint:* The `abs()` function returns absolute value of a number.
```
# YOUR CODE HERE
```
Two lists can also be join together simply using `+` operator. This is called the **concatenation of lists**.
#### Exercise 3.14
Concatenate the lists `[1,2,3,4,5]` and `[6,7,8,9]`
```
# YOUR CODE HERE
```
Similar to String, we can repeat a list multiple times with `*` operator.
#### Exercise 3.15
Create a list `[1,2,3,1,2,3,1,2,3,1,2,3]` from list `[1,2,3]`.
```
# YOUR CODE HERE
```
Recap: String has many similar behaviors as list.
#### Exercise 3.16
* How to concatenate 2 strings?
* How to repeat a string 2 times?
```
# YOUR CODE HERE
```
#### 3.1.1.4 Membership and Searching elements in a list
You might need to check if a particular item is in a list.
Instead of using `for` loop to iterate over the list and use the if condition, Python provides a simple **`in`** statement to check membership of an item.
#### Exercise 3.17
Write code to find out whether `'duck'` and `'dog'` are in the list `['duck', 'chicken', 'goose']` respectively.
```
# YOUR CODE HERE
```
To count the occurence of a particular item in a list, we can use the `.count()` method. Syntax is
>```
>your_list.count(item)
>```
#### Exercise 3.18
* Create a list `['duck', 'chicken', 'goose', 'duck', 'chicken', 'goose', 'duck', 'chicken', 'goose']` from `['duck', 'chicken', 'goose']`
* Count number of occurence of `'duck'`
```
# YOUR CODE HERE
```
To find the index value of a particular item, we can use `.index()` method. Syntax is
>```
>your_list.index(item)
>```
* If there are multiple items of the same value, only the first index value of that item is returned.
* You can add 2nd argument `x` to start searching from index `x` onwards.
Note: the string functions `find()` and `rfind()` are <u>not available</u> for list.
#### Exercise 3.19
* Create a list `['duck', 'chicken', 'goose', 'duck', 'chicken', 'goose', 'duck', 'chicken', 'goose']` from `['duck', 'chicken', 'goose']`
* Find the index of the first occurence of `'duck'` in the list.
* Find ALL the indices of occurences of `'duck'` in the list.
```
# YOUR CODE HERE
l=['duck', 'chicken', 'goose', 'duck', 'chicken', 'goose', 'duck', 'chicken', 'goose']
l_copy=['duck', 'chicken', 'goose','goose','duck', 'chicken', 'goose', 'duck', 'chicken', 'goose']
elem='goose'
index=l_copy.index(elem)
print(index)
try:
while l_copy.index(elem)+1:
l_copy=l_copy[l_copy.index(elem)+1:]
index=index+l_copy.index(elem)+1
print(index)
except:
print(f"That's all the indices of {elem} in the list.")
```
#### 3.1.1.5 Iterating through List
To iterate through a collection, e.g. list or tuple, you can use **for-loop**.
>```
>for item in my_list:
> # Process each item
>```
#### Exercise 3.20
Print out each item in `names` list.
```
names = ['duck', 'chicken', 'goose']
```
```
# YOUR CODE HERE
```
To find the index value of each item in the list, we can use `enumerate()` function. Syntax is
>```
>enumerate(your_list)
>```
#### Exercise 3.21
* Print items in list `['duck', 'chicken', 'goose']` as following output.
>```
>0 duck
>1 chicken
>2 goose
>```
Python provides a very handy way to perform same operate on all items in a list, and return a new list, called *list comprehension*.
#### Exercise 3.21
Create a list which contains len() value of each item in `['duck', 'chicken', 'goose']`
```
# YOUR CODE HERE
```
#### Exercise 3.22
Create a list which contain the first 100 terms of an arithmetic sequence with first term 2 and common difference 3.
```
# YOUR CODE HERE
```
#### Exercise 3.23
How to prefix all items with a string 'big', which resulted in `['big duck', 'big chicken', 'big goose']`?
```
names = ['duck', 'chicken', 'goose']
```
```
# YOUR CODE HERE
```
#### 3.1.1.4 Modifying a list
List is a **mutable** collection, i.e. list can be modified and item value can be updated.
It is easy to update an item in the list by its index value.
#### Exercise 3.24
For a list `s = [0,1,2,3,4]`, update its 3rd item to `9`.
```
# YOUR CODE HERE
```
The `list.append( )` method is used to append (add) a element at the end of the list. Syntax is
>```
>your_list.append(element)
>```
#### Exercise 3.24
For a list `s = [0,1,2,3,4,5]`, append a value `6` to it.
```
# YOUR CODE HERE
```
#### Exercise 3.24
What happens if you append a list `[7,8,9]` to a list `[1,2,3,4,5]`?
```
# YOUR CODE HERE
```
A list can also be **extended** with items from another list using `list.extend()` method. It will modify the first list. The resultant list will contain all the elements of the lists that were added, i.e. the resultant list is NOT a nested list. Syntax is
>```
>your_list.extend(another_list)
>```
**Note** the difference between `append()` and `extend()`.
#### Exercise 3.25
Extend a list `[1,2,3,4,5,6]` with all items in another list `[7,8,9]`.
```
# YOUR CODE HERE
```
**Question:**
* Can you re-write above code using `+` operator?
* Can you insert `[7,8,9]` in the middle of `[1,2,3,4,5,6]` using `+` operator?
```
# YOUR CODE HERE
```
The method `list.insert(position,new_value)` is used to insert a element `new_value` at a specified index value `position`. Syntax is
>```
>your_list.insert(position,new_value)
>```
* `list.insert()` method does not replace element at the index.
* `list.append( )` method can only insert item at the end.
#### Exercise 3.26
Use `list.insert()` method to modify a string `'What a day'` to `'What a sunny day'`.
* *Hint:* Use `str.split()` and `str.join()` methods.
```
# YOUR CODE HERE
```
`list.pop( )` method can be used to remove the last element in the list. This is similar to the operation of a stack (which we will cover later in the course.) Syntax is
>```
>your_list.pop()
>```
#### Exercise 3.27
Use `list.pop()` function to remove items in list `[0,1,2,3,4]` in reverse order.
```
# YOUR CODE HERE
```
Index value can be specified to `list.pop()` method to remove a certain element corresponding to that index value. Syntax is
>```
>your_list.pop(index)
>```
#### Exercise 3.28
Use `list.pop()` method to remove `'c'` from list `['a','b','c','d','e']`.
`list.remove( )` method is used to remove an item based on its value. Syntax is
>```
>your_list.remove(value)
>```
* If there are multiple items of same value, it will only remove 1st item.
* It will throw an **exception** if the value is not found in the list. You may need to enclose it with `try-except` block. (We will come back to this)
#### Exercise 3.29
Use `list.remove()` function to remove value `3` three times.
```
lst = [0,1,2,3,4] * 2
```
```
# YOUR CODE HERE
```
#### Exercise 3.30
How to remove all values `3` in the list `[1,2,3,4,1,2,3,4,1,2,3,4]`?
* *Hint:* Use `while`, `try-except`, and `break`
```
# YOUR CODE HERE
```
To clear all elements in a list, use its `list.clear()` method.
#### Exercise 3.31
Clear all items in `s = [1,2,3]`.
```
# YOUR CODE HERE
```
## Recap
* How to create a new list?
* Can a list hold elements of different data type?
* Why do you need multiple level indexing?
* Name 3 functions or operators which works with list.
* What is the keyword used to check membership in a list?
* How to add an item to a list?
* How to remove an item from a list?
* How to merge 2 lists?
### 3.1.2 Tuples
A Tuple is a collection of Python objects which is **immutable**, i.e. not modifiable after creation.
#### 3.1.2.1 Creating a Tuple
Tuple is created with listed of items surrounded by parentheses **"( )"**, and seperated by comma **","**.
* To create an empty tuple, simple use `()`
* To create a single-item tuple, need to add **common `,`** behind the element. E.g. `tup = (3,)`
#### Exercise 3.32
* Create a tuple `t` with values `1, 2, 3, 4`
* Print it and its type
```
# YOUR CODE HERE
t=(1,2,3,4)
print(type(t))
```
In fact, parentheses is optional unless it is to create an empty tuple.
```
# YOUR CODE HERE
```
Tuple can also be created using by typecasting a list using `tuple()` constructor function.
* when string is passed in as argument, it turns string into collection of characters.
#### Exercise 3.33
* Create a tuple from list `[1,2,3]`.
* What happen if you apply `tuple()` constructor function on a string `Good day!`?
```
# YOUR CODE HERE
a=tuple([1,2,3])
type(a)
tuple('Good day!')
```
Python collections allows mix data types in the same collection. We can also create tuple with items of different data type, although this is not commonly used.
#### Exercise 3.34
* Create a tuple with items `'apple', 3.0, 'banana', 4`.
```
# YOUR CODE HERE
```
Tuple can contain other tuples as its elements. Such tuples are called **nested tuples**.
#### Exercise 3.35
* Create a tuple `nested` with items `0, 1, (2, 3, 4), (5, 6)`
* What's the length of above tuple?
```
# YOUR CODE HERE
b=(0, 1, (2, 3, 4), (5, 6))
len(b)
```
#### 3.1.2.2 Accessing an Element in tuple via indexing.
Items in collection can be accessed by their indexes. Python uses zero-based indexing, i.e. index starts from 0.
#### Exercise 3.36
* Create a tuple `('apple', 'banana', 'cherry', 'durian')` and assign it to the variable `fruits`.
* Print out 2nd and 4th item in the tuple.
```
# YOUR CODE HERE
fruits=('apple', 'banana', 'cherry', 'durian')
print(fruits[1])
print(fruits[3])
```
Indexing can also be done in reverse order. That is the last element has an index of -1, and second last element has index of -2.
<center>
<img src="./images/list-indexing.png" alt="Set Venn Diagram" style="width: 400px;"/>
</center>
#### Exercise 3.37
* Use <u>negative indexing</u> to print out <u>last</u> and <u>2nd last</u> item in `fruits`.
```
# YOUR CODE HERE
```
For nested tuples, we can access items by multi-level indexing as well. Each level of the index always starts from 0.
#### Exercise 3.38
* Create a tuple `nested` with items `0, 1, (2, 3, 4), (5, 6)`.
* How do you access value `4` and `5`?
```
# YOUR CODE HERE
```
#### 3.1.2.3 Accessing a Subset of Tuple via Slicing.
**Indexing** was only limited to accessing a single element.
**Slicing** on the other hand is accessing a sequence of data inside the tuple.
**Slicing** is done by defining the index values of the `first element` and the `last element` from the parent list that is required in the sliced tuple.
>```
>sub = num[a : b]
>sub = num[a : ]
>sub = num[: b]
>sub = num[:]
>```
* if both `a` and `b` are specified, `a` is the first index, `b` is the **last index + 1**.
* if `b` is omitted, it will slice till last element.
* if `a` is omitted, it will starts from first element.
* if neither `a` or `b` is specified, it is effectively copy the whole tuple.
**Note: the upper bound index is NOT inclusive!**
Try out following code.
>```
>num = tuple(range(10))
>
># Get item with index 2 to 4
>print(num[2:5])
>
># Get first 5 items
>print(num[:5])
>
># Get from item with index = 5 onwards
>print(num[5:])
>```
```
# YOUR CODE HERE
```
Remember tuple items can be accessed using `negative index`. Same technique can be applied for slicing too.
* Last item has index of -1
#### Exercise 3.39
Consider the tuple `num = (0,1,2,3,4,5,6,7,8,9)`.
* How to get last 3 items from a tuple?
* How to ignore last 3 items from a tuple?
* How to strip first and last items from a tuple?
```
# YOUR CODE HERE
```
#### 3.1.2.4 Working with Tuple
##### 3.1.2.4.1 Length of tuples `len()`
To find the length of the tuple or the number of elements in a tuple, the function `len( )` is used. Syntax is
>```
>len(tuples)
>```
##### 3.1.2.4.2 Maximum `max()`, minimum `min()` and sum `sum()` of all the values in the tuple
If the tuple consists of all integer elements, the functions `min( )`, `max( )` and `sum()` gives the minimum item, maximum item and total sum value of the tuple. Syntax is
>```
>max(your_tuple)
>min(your_tuple)
>sum(your_tuple)
>```
#### Exercise 3.40
Print out min value, max value and sum of tuple with items 0-9.
```
# YOUR CODE HERE
```
#### Exercise 3.41
How do you create a tuple by reversing another tuple?
```
# YOUR CODE HERE
```
If elements are string type, `max()` and `min()` is still applicable. `max()` would return a string element whose ASCII value is the highest and the lowest when `min()` is used. Note that only the first index of each element is considered each time and if they value is the same then second index considered so on and so forth.
#### Exercise 3.42
What's the minimum and the maximum value of tuple `poly = ('np','sp','tp','rp','nyp')`?
```
# YOUR CODE HERE
```
##### 3.1.2.4.3 Checking Values in the tuple with `any()` and `all()` function
`any()` function returns `True` if any item in tuple (collection) is evaluated `True`.
`all()` function returns `True` if all items in tuple (collection) is evaluated `True`.
Python evaluates following values as `False`
* `False`, `None`, numeric zero of all types
* Empty strings and containers (including strings, tuples, lists, dictionaries, sets and frozensets)
#### Exercise 3.43
For each list of `[2, 4, 0, 9]`, `['', 'hello', 'world']`.
* use `any()` to check if the list contains any non-zero item.
* use `all()` to check if all items in the list are non-zero .
```
# YOUR CODE HERE
```
##### 3.1.2.4.4 Reversing the values in a tuple with `reversed()` function
Unlike list, tuple is immutable. Thus `.reverse()` method is **NOT** applicable to tuple.
The `reversed()` function returns a reversed object which can be converted to be a tuple or list. Syntax is
>```
>reversed(my_tuple)
>```
```
poly = ('np','sp','tp','rp','nyp')
r = reversed(poly)
print(type(r), tuple(r))
```
```
# YOUR CODE HERE
# poly=('np','sp','tp','rp','nyp')
# poly.reverse()
# print(poly)
poly = ('np','sp','tp','rp','nyp')
r = reversed(poly)
print(tuple(r))
print(type(r), tuple(r))
```
##### 3.1.2.4.5 Arranging the values in a tuple with `reversed()` function
Similarly, `.sort()` method cannot be applied directly to tuple itself.
`sorted()` function to arrange the elements in **ascending** order. Syntax is
>```
>sorted(my_tuple)
>```
```
poly = ('np','sp','tp','rp','nyp')
s = sorted(poly)
print(poly)
print(s)
```
```
# YOUR CODE HERE
```
For **descending** order, specify the named argument `reverse = True`.
By default the reverse condition will be `False` for `reverse`. Hence changing it to `True` would arrange the elements in descending order.
The `sort()` function has another named argument `key`, which allows us to specify a callable function. The sorting will be done based on returned value from this callable function.
For example, `len()` function returns length of a string.
To sort based on string length, `key = len` can be specified as shown.
```
names = ('duck', 'chicken', 'goose')
s1 = sorted(names, key=len)
print(s1)
# YOUR CODE HERE where s2 is in reverse order
print(s2)
```
Two tuples can also be join together simply using `+` operator.
Besides tuples, `str` objects are also immutable. New string will be created when a string is modified.
```
(1,2,3)+(4,5)
s = "hello world."
t = s
print(t == s, t is s)
s = s + "abc"
print(t == s, t is s)
```
Similarly, tuple is immutable. Modification to tuple will return a new tuple object.
Similar to list, we can repeat a tuple multiple times with `*` operator.
```
t1 = (1,2,3,4,5,6)
t2 = t1 * 2
print(t1, t2)
```
#### 3.1.2.5 Membership and Searching elements in a tuple
You might need to check if a particular item is in a tuple.
Instead of using `for` loop to iterate over the tuple and use the `if` condition, Python provides a simple `in` statement to check membership of an item.
>```
>names = ('duck', 'chicken', 'goose')
>found1 = 'duck' in names
>found2 = 'dog' in names
>print(found1, found2)
>```
`count( )` is used to count the occurence of a particular item in a list.
```
names = ('duck', 'chicken', 'goose') * 2
print(names)
names.count('duck')
```
`index( )` is used to find the index value of a particular item.
* Note that if there are multiple items of the same value, the first index value of that item is returned.
* You can add 2nd argument `x` to start searching from index `x` onwards.
```
names2 = ('duck', 'chicken', 'goose') * 2
idx = names2.index('goose')
idx2 = names2.index('goose', 4)
print('Gooses at index {} and {}'.format(idx, idx2))
```
#### 3.1.2.6 Membership and Searching elements in a tuple
To iterate through a tuple (collection), you can use `for` loop.
```
names = ('duck', 'chicken', 'goose')
for name in names:
print(name)
```
If you need the index value, you can use `enumerate()` function.
```
for idx, name in enumerate(names):
print(idx, name)
```
#### 3.1.2.7 Function with Multiple Returning Values
In most programming languages, function/method can only return a single value. It is the same practice in Python.
But in Python, you can return a tuple which can easily pack multiple values together.
## Exercise 3.44
Define a function `minmax()` which fulfils following conditions. Test the function with list `\[1,2,3,4,5\]'.
- accept a list as input
- return both min and max values of the list
Tuple can be easily unpacked into multiple values.
During unpacking, number of variable needs to match number of items in tuple
```
x, y, z = 1, 2, 3
print(x, y, z)
```
It is common to use underscore `_` for items to be ignored.
```
times = '9am to 5pm to 6pm to 8pm'.split('to')
print(times)
start, _, end, _ = tuple(times)
print(start, end)
```
#### Exercise 3.45
How to swap two values `x` and `y` in a single statement?
```
x = 10
y = 20
# YOUR CODE HERE
print('x = {}, y = {}'.format(x, y))
x,y =y,x
print('x = {}, y = {}'.format(x, y))
```
You can use `*` to hold any number of unpacked values.
For example, from a tuple, you would like to get its last item, and put all other items in a list.
```
t = (1,2,3,4,5)
#a,b =t
*a, b = t
print(a, b)
```
#### Exercise 3.46
How to extract only first and last items from a tuple `(1,2,3,4,5)`.
```
t = (1,2,3,4,5)
# YOUR CODE HERE
a,*b,c=t
print(a, b, c)
```
## Recap
### Difference between Tuple and List
A tuple is **immutable** whereas a list is **mutable**.
* You can't add elements to a tuple. Tuples have no append or extend method.
* You can't remove elements from a tuple. Tuples have no remove or pop method.
### When to use Tuple?
* Tuples are used in function to return multiple values together.
* Tuples are lighter-weight and are more memory efficient and often faster if used in appropriate places.
* When using a tuple you protect against accidental modification when passing it between functions.
* Tuples, being immutable, can be used as a key in a dictionary, which we’re about to learn about.
#
## 3.2 Hash Table
A **hash table** (hash map) is a data structure that stores data in an associative manner. Roughly speaking, it is an unordered collection of key-value pairs, which are just a pair of values where by knowing the `key` value, you can retrieve `value` value.
Hash table uses a fixed size array as a storage medium and uses **hash function** to generate an index where an element is to be inserted or is to be located from. Hash table allows us to do a key-value lookup.
#### Example
#### Linear Probing
Reference: https://www.youtube.com/watch?v=sfWyugl4JWA
Hashing
### 3.2.1 Dictionary
Dictionaries are a common feature of modern languages (often known as maps, associative arrays, or hashmaps) which let you associate pairs of values together.
In Python, dictionaries are defined in **dict** data type.
* It stores keys and their corresponding values.
* Keys must be **unique** and **immutable**.
* It is **mutable**, i.e. you can add and remove items from a dictionary.
* It is **unordered**, i.e. items in a dictionary are not ordered.
* Elements in a dictionary is of the form `key_1:value_1, key_2:value_2,....`
#### 3.2.1.1 Creating a dictionary
Dictionary is created with listed of items surrounded by curly brackets `{}`, and seperated by comma `,`.
* To create an empty dictionary, simple use `{}`
* Key and value are separated by colon `:`
* Key needs to be **immutable** type, e.g. data type like scalar, string or tuple
#### Example
```
# empty dictionary
d0 = {}
# dictionary with mixed data type
d1 = {'name': 'John', 1: [2, 4, 3]}
print(d1)
```
#### Example
Create a dictionary `fruits` which has following keys and values.
| key | value |
|-----|----------|
| a | Apple |
| b | Banana |
| c | Cherries |
| d | Durian |
```
# YOUR CODE HERE
```
New dictionary can be created from a list of tuples too using the `dict()` constructor function, where each tuple contains a key and a value. Syntax is
>`dict(my_list_of_tuples)`
**Example**
Construct a dictionary `f3` using list `[('a','Apple'), ('b','Banana'), ('c','Cherries'), ('d','Durian')]`.
```
# YOUR CODE HERE
```
#### 3.2.1.2 Accessing an Element in dictionary by its respective key.
Items in dictionary can be accessed by their respective keys.
* Key can be used either inside square brackets or with the `get()` method.
* The difference while using `get()` is that it returns `None` instead of `KeyError` Exception, if the key is not found.
* `get()` method can take in a default value argument, which will be returned if the key is not found. Syntax is
>`your_dict.get(key,message_if_unavailable)`
#### Example
What happens when you try to use a non-existing key?
```
print(fruits)
print(fruits['a'])
print(fruits['b'])
print(fruits['z'])
```
#### Example
What happens when you use `.get()` and try to use a non-existing key?
```
print(fruits)
print(fruits.get('a', 'Not available'))
print(fruits.get('z', 'Not available'))
```
3.2.1.3 Finding number of elements in a dictionary with `len()` function
To find the number of elements in a dictionary, `len()` function is used. Syntax is
>```
>len(my_dictionary)
>```
#### Example:
Find the length of `fruits` dictionary.
```
# YOUR CODE HERE
```
#### 3.2.1.4 `dict.keys()`, `dict.values()`, `dict.items()` methods
* `keys()` method return the dictionary's keys as `dict_keys` object.
* `values()` method return the dictionary's values as `dict_values` object.
* `items()` return the dictionary's key-value pairs as `dict_items` object.
If you want the various collections as a list typecast the objects using the `list` constructor function.
#### Example
Print out the keys, values and key-value pairs of the dictionary `fruits`.
```
print(fruits.keys())
print(fruits.values())
print(fruits.items())
```
#### 3.2.1.5 Modifying and Updating a dictionary
Similar to list, dictionary is a **mutable** collection, i.e. dictionary can be modified and the values of existing items can be updated. Syntax is
>```
>your_dictionary[your_key]=your_value
>```
* If the key exists in the dictionary, existing value will be updated.
* If the key doesn't exists in the dictionary, new key:value pair is added to dictionary.
#### Example
Using the `fruits` dictionary defined earlier.
- Update its key `a` value to `['Apple', 'Apricots', 'Avocado']`
- Add another key-value pair `{'f':'Fig'}` to `fruits` dictionary.
```
# YOUR CODE HERE
```
#### 3.2.1.6 Merging Dictionaries with `.update()`
`.update()` method is used to merge items from another dictionary.
* Adds element(s) to the dictionary if the key is not in the dictionary.
* If the key is in the dictionary, it updates the key with the new value.
#### Example
* Create another dictionary `fruits_too` with items `{'d':'Dates', 'e':'Eldercherry', 'f':'Fig', 'g':'Grape'}`.
* Add/update items from `fruits_too` to `fruits`.
```
# YOUR CODE HERE
```
#### 3.2.1.7 Removing Items with `.pop()`, `.popitem()`, `.clear()`
`.pop()` method is used to remove an item by key and returns the value. Syntax is
>```
>my_dictionary.pop(my_item)
>```
It throws exception if key is not found.
#### Example
```
fruits = {'a': 'captain', 'b': 'Banana', 'c': 'Cherry', 'd': 'Durian', 'f': 'Fig'}
print(fruits)
p = fruits.pop('b')
print(fruits)
print(p)
b = fruits.popitem()
print(fruits)
print(b)
fruits.clear()
print(fruits)
```
`.popitem()` removes any arbitrary item.
`.clear()` clears all items in a dictionary.
```
mixed = dict(fruits)
print(mixed.popitem())
mixed.clear()
print(mixed)
```
#### 3.2.1.8 Iterating Through Dictionary
To iterate through a dictionary, you can use for-loop. By default, the iteration is done ONLY on **keys** of the dictionary.
#### Example
```
fruits = {'a': 'captain', 'b': 'Banana', 'c': 'Cherry', 'd': 'Durian', 'f': 'Fig'}
for key in fruits:
print(key)
```
#### Exercise
Write a code to:
1. iterate through the values in the `fruits` dictionary, and
2. iterate through the keys and values in the `fruits` dictionary at the same time.
#### 3.2.1.9 Dictionary Comprehension
Similiar to list, we can also use dictionary comprehension to easily generate a dictionary.
#### Example
```
s = [x*2 for x in range(10)]
print(s)
d = {x: x*x for x in range(1,10)}
print(d)
```
#### 3.2.1.10 Membership Test
We can use `in` operator to check membership of a key in a dictionary.
#### Example
Check whether key `a` and `z` are in the `fruits` dictionary. By default, membership testing is again done on keys.
```
print(fruits)
# by default, membership testing is done on keys
print('a' in fruits)
print('z' in fruits)
```
#### Exercise
* How to test if a value `Apple` is in a dictionary?
* How to test if a key-value pair `{'a':'Apple'}` is in the dictionary?
* Let `d1 = {'a':'Apple', 'c':'Cherries'}`. How to check if all key-value pairs in one dictionary `d1` are in the dictionary `fruits`?
```
#YOUR CODE HERE
```
In a dictionary, to find key by matching its value, we can either use:
* Option 1: for-loop
* Option 2: `.index()` method
#### Example
```
x = 'Cherry'
s = list(fruits.values())
print(s)
i = s.index(x)
print(i)
k = list(fruits.keys())
print(k[i])
```
## Recap
* How to create a dictionary?
* How to copy a dictionary?
* How to retrieve an item by key? by `[]` & by `.get()`
* How to update an item?
* How to add an item?
* How to remove an item?
* How to merge an dictionary to another?
* What's the differences among `dict.keys()`, `dict.values()` and `dict.items()`
## 3.3 Set
A set is an **unordered** collection of **unique** values. In Python, `set` is the data type for Sets.
* Every element in set is unique (no duplicates) and must be **immutable** (which cannot be changed).
* `set` itself is mutable. We can add or remove items from it.
* `set` is often used to eliminate repeated numbers in a sequence/list.
### 3.3.1. How to create a set?
A set is created by either of following methods:
* Placing all the items (elements) inside curly braces `{}`, separated by comma `,`
* Using the built-in constructor function `set()`, which takes in a collection (list or tuple)
**Note:**
* Set can contain mixed data type.
* All duplicate value will be discarded.
#### Example
```
s = [1,2,3,4]*3
print(s)
set3 = set(s)
print(set3)
```
**Question**
* Why we can't use `{}` to create an empty set?
#### Example
```
s = {}
print(type(s))
# empty set
set0 = set()
print(type(set0))
```
#### 3.3.2 How to modify a set?
Set is mutable. Thus you can perform following actions on a set.
* Add a single item
* Updae/add multiple items
* remove items
##### 3.3.2.1 Add a Single Item with `add()`
`add()` function is used to add a single item to the list.
If the item is already exists in the set, it will be ignored.
#### Example
```
set1 = {1,2,3,4}
set1.add(5)
set1.add(5)
print(set1)
```
##### 3.3.2.2 Add Multiple Items with `update()`
`update()` function is used to add multiple items to a set. It can take in one or more collections, e.g. list, tuple or another set.
* If the item already exists, it will be discarded.
* If the item does not exist, it will be added.
#### Example
```
set1 = {1,2,3,4}
print(set1)
set1.update([2,3,4,5])
print(set1)
set1.update([(5,6)], {7,8}, (9,10))
print(set1)
```
##### 3.3.2.3 Remove Item by Value
An item can be removed from set using `discard()` and `remove()` methods.
* `discard()` method does not throw exception if the item is not found in the set.
* `remove()` method will raise an `Exception` in such condition.
```
set1 = {1,2,3,4}
set1.discard(4)
set1.discard(44)
print(set1)
set1.remove(33)
print(set1)
```
##### 3.3.2.4 Remove an Arbitrary Item
`pop()` function can be used to remove an **arbitrary** item from set. This is because set is unordered.
* The popped value may seem following a sequence. But that is due to the internal hashmap implementation. It is not reliable and depends on values.
set1 = {1,2,3,4}
print(set1.pop())
print(set1.pop())
print(set1)
##### 3.3.2.5 Remove an Arbitrary Item
Sets can be used to carry out mathematical set operations
* intersection
* union
* symmetric difference
* difference (subtracting)
<img src=".images/set-venn.png" alt="Set Venn Diagram" style="width: 500px;"/>
##### Example
Create 2 sets using `set()` constructor.
```
set1 = set(range(0,8))
set2 = set(range(5,13))
print(set1)
print(set2)
```
##### 3.3.2.6 Intersection
`intersection()` function outputs a set which contains all the elements that are in both sets.
* Operator `&` can be used for Intersection operation.
```
print(set1.intersection(set2))
print(set1 & set2)
```
##### 3.3.2.7 Union
`union( )` function returns a set which contains all the elements of both the sets without repition.
* Operator `|` can be used for Union operation.
```
print(set1.union(set2))
print(set1 | set2)
```
##### 3.3.2.8 Difference (Subtracting)
`difference()` function ouptuts a set which contains elements that are in set1 and not in set2.
* Operator `-` can be use for Subtracting operation.
```
print(set1.difference(set2))
print(set1 - set2)
```
#### 3.3.2.9 Symetric Difference
`symmetric_difference()` function ouputs a function which contains elements that are in one of the sets.
```
s7 = set1.symmetric_difference(set2)
print(s7)
```
#### 3.3.2.10 Subset, Superset
`issubset()`, `issuperset()` is used to check if the set1/set2 is a subset, superset of set2/set1 respectively.
```
set3 = set(range(0,15))
set4 = set(range(5,10))
print(set3)
print(set4)
r1 = set3.issubset(set4)
r2 = set3.issuperset(set4)
print(r1, r2)
set3 = set(range(0,15))
set4 = set(range(5,10))
r1 = set4.issubset(set3)
r2 = set3.issuperset(set4)
print(r1, r2)
```
#### 3.3.2.11 Disjoint
`isdisjoint()` is used to check if the set1/set2 is disjoint.
```
r3 = set3.isdisjoint(set4)
print(r3)
set5 = set(range(5))
set6 = set(range(6,10))
r4 = set5.isdisjoint(set6)
print(r4)
```
| github_jupyter |
# Use Spark to predict product line with `ibm-watson-machine-learning`
This notebook contains steps and code to get data from the IBM Data Science Experience Community, create a predictive model, and start scoring new data. It introduces commands for getting data and for basic data cleaning and exploration, pipeline creation, model training, model persistance to Watson Machine Learning repository, model deployment, and scoring.
Some familiarity with Python is helpful. This notebook uses Python 3.8 and Apache® Spark 2.4.
You will use a publicly available data set, **GoSales Transactions**, which details anonymous outdoor equipment purchases. Use the details of this data set to predict clients' interests in terms of product line, such as golf accessories, camping equipment, and others.
## Learning goals
The learning goals of this notebook are:
- Load a CSV file into an Apache® Spark DataFrame.
- Explore data.
- Prepare data for training and evaluation.
- Create an Apache® Spark machine learning pipeline.
- Train and evaluate a model.
- Persist a pipeline and model in Watson Machine Learning repository.
- Deploy a model for online scoring using Wastson Machine Learning API.
- Score sample scoring data using the Watson Machine Learning API.
- Explore and visualize prediction result using the plotly package.
## Contents
This notebook contains the following parts:
1. [Setup](#setup)
2. [Load and explore data](#load)
3. [Create spark ml model](#model)
4. [Persist model](#persistence)
5. [Predict locally](#visualization)
6. [Deploy and score](#scoring)
7. [Clean up](#cleanup)
8. [Summary and next steps](#summary)
<a id="setup"></a>
## 1. Set up the environment
Before you use the sample code in this notebook, you must perform the following setup tasks:
- Contact with your Cloud Pack for Data administrator and ask him for your account credentials
### Connection to WML
Authenticate the Watson Machine Learning service on IBM Cloud Pack for Data. You need to provide platform `url`, your `username` and `api_key`.
```
username = 'PASTE YOUR USERNAME HERE'
api_key = 'PASTE YOUR API_KEY HERE'
url = 'PASTE THE PLATFORM URL HERE'
wml_credentials = {
"username": username,
"apikey": api_key,
"url": url,
"instance_id": 'openshift',
"version": '4.0'
}
```
Alternatively you can use `username` and `password` to authenticate WML services.
```
wml_credentials = {
"username": ***,
"password": ***,
"url": ***,
"instance_id": 'openshift',
"version": '4.0'
}
```
### Install and import the `ibm-watson-machine-learning` package
**Note:** `ibm-watson-machine-learning` documentation can be found <a href="http://ibm-wml-api-pyclient.mybluemix.net/" target="_blank" rel="noopener no referrer">here</a>.
```
!pip install -U ibm-watson-machine-learning
from ibm_watson_machine_learning import APIClient
client = APIClient(wml_credentials)
```
### Working with spaces
First of all, you need to create a space that will be used for your work. If you do not have space already created, you can use `{PLATFORM_URL}/ml-runtime/spaces?context=icp4data` to create one.
- Click New Deployment Space
- Create an empty space
- Go to space `Settings` tab
- Copy `space_id` and paste it below
**Tip**: You can also use SDK to prepare the space for your work. More information can be found [here](https://github.com/IBM/watson-machine-learning-samples/blob/master/cpd4.0/notebooks/python_sdk/instance-management/Space%20management.ipynb).
**Action**: Assign space ID below
```
space_id = 'PASTE YOUR SPACE ID HERE'
```
You can use `list` method to print all existing spaces.
```
client.spaces.list(limit=10)
```
To be able to interact with all resources available in Watson Machine Learning, you need to set **space** which you will be using.
```
client.set.default_space(space_id)
```
<a id="load"></a>
## 2. Load and explore data
In this section you will load the data as an Apache® Spark DataFrame and perform a basic exploration.
Load the data to the Spark DataFrame by using *wget* to upload the data to gpfs and then *read* method.
### Test Spark
```
try:
from pyspark.sql import SparkSession
except:
print('Error: Spark runtime is missing. If you are using Watson Studio change the notebook runtime to Spark.')
raise
```
The csv file GoSales_Tx.csv is availble on the same repository where this notebook is located. Load the file to Apache® Spark DataFrame using code below.
```
import os
from wget import download
sample_dir = 'spark_sample_model'
if not os.path.isdir(sample_dir):
os.mkdir(sample_dir)
filename = os.path.join(sample_dir, 'GoSales_Tx.csv')
if not os.path.isfile(filename):
filename = download('https://github.com/IBM/watson-machine-learning-samples/raw/master/cpd4.0/data/product-line-prediction/GoSales_Tx.csv', out=sample_dir)
spark = SparkSession.builder.getOrCreate()
df_data = spark.read\
.format('org.apache.spark.sql.execution.datasources.csv.CSVFileFormat')\
.option('header', 'true')\
.option('inferSchema', 'true')\
.load(filename)
df_data.take(3)
```
Explore the loaded data by using the following Apache® Spark DataFrame methods:
- print schema
- print top ten records
- count all records
```
df_data.printSchema()
```
As you can see, the data contains five fields. PRODUCT_LINE field is the one we would like to predict (label).
```
df_data.show()
df_data.count()
```
As you can see, the data set contains 60252 records.
<a id="model"></a>
## 3. Create an Apache® Spark machine learning model
In this section you will learn how to prepare data, create an Apache® Spark machine learning pipeline, and train a model.
### 3.1: Prepare data
In this subsection you will split your data into: train, test and predict datasets.
```
splitted_data = df_data.randomSplit([0.8, 0.18, 0.02], 24)
train_data = splitted_data[0]
test_data = splitted_data[1]
predict_data = splitted_data[2]
print("Number of training records: " + str(train_data.count()))
print("Number of testing records : " + str(test_data.count()))
print("Number of prediction records : " + str(predict_data.count()))
```
As you can see our data has been successfully split into three datasets:
- The train data set, which is the largest group, is used for training.
- The test data set will be used for model evaluation and is used to test the assumptions of the model.
- The predict data set will be used for prediction.
### 3.2: Create pipeline and train a model
In this section you will create an Apache® Spark machine learning pipeline and then train the model.
In the first step you need to import the Apache® Spark machine learning packages that will be needed in the subsequent steps.
```
from pyspark.ml.feature import OneHotEncoder, StringIndexer, IndexToString, VectorAssembler
from pyspark.ml.classification import RandomForestClassifier
from pyspark.ml.evaluation import MulticlassClassificationEvaluator
from pyspark.ml import Pipeline, Model
```
In the following step, convert all the string fields to numeric ones by using the StringIndexer transformer.
```
stringIndexer_label = StringIndexer(inputCol="PRODUCT_LINE", outputCol="label").fit(df_data)
stringIndexer_prof = StringIndexer(inputCol="PROFESSION", outputCol="PROFESSION_IX")
stringIndexer_gend = StringIndexer(inputCol="GENDER", outputCol="GENDER_IX")
stringIndexer_mar = StringIndexer(inputCol="MARITAL_STATUS", outputCol="MARITAL_STATUS_IX")
```
In the following step, create a feature vector by combining all features together.
```
vectorAssembler_features = VectorAssembler(inputCols=["GENDER_IX", "AGE", "MARITAL_STATUS_IX", "PROFESSION_IX"], outputCol="features")
```
Next, define estimators you want to use for classification. Random Forest is used in the following example.
```
rf = RandomForestClassifier(labelCol="label", featuresCol="features")
```
Finally, indexed labels back to original labels.
```
labelConverter = IndexToString(inputCol="prediction", outputCol="predictedLabel", labels=stringIndexer_label.labels)
```
Let's build the pipeline now. A pipeline consists of transformers and an estimator.
```
pipeline_rf = Pipeline(stages=[stringIndexer_label, stringIndexer_prof, stringIndexer_gend, stringIndexer_mar, vectorAssembler_features, rf, labelConverter])
```
Now, you can train your Random Forest model by using the previously defined **pipeline** and **train data**.
```
train_data.printSchema()
model_rf = pipeline_rf.fit(train_data)
```
You can check your **model accuracy** now. To evaluate the model, use **test data**.
```
predictions = model_rf.transform(test_data)
evaluatorRF = MulticlassClassificationEvaluator(labelCol="label", predictionCol="prediction", metricName="accuracy")
accuracy = evaluatorRF.evaluate(predictions)
print("Accuracy = %g" % accuracy)
print("Test Error = %g" % (1.0 - accuracy))
```
You can tune your model now to achieve better accuracy. For simplicity of this example tuning section is omitted.
<a id="persistence"></a>
## 4. Persist model
In this section you will learn how to store your pipeline and model in Watson Machine Learning repository by using python client libraries.
**Note**: Apache® Spark 2.4 is required.
### 4.1: Save pipeline and model
In this subsection you will learn how to save pipeline and model artifacts to your Watson Machine Learning instance.
```
saved_model = client.repository.store_model(
model=model_rf,
meta_props={
client.repository.ModelMetaNames.NAME:'Product Line model',
client.repository.ModelMetaNames.TYPE: "mllib_2.4",
client.repository.ModelMetaNames.SOFTWARE_SPEC_UID: client.software_specifications.get_id_by_name('spark-mllib_2.4'),
client.repository.ModelMetaNames.LABEL_FIELD: "PRODUCT_LINE",
},
training_data=train_data,
pipeline=pipeline_rf)
```
Get saved model metadata from Watson Machine Learning.
```
published_model_id = client.repository.get_model_uid(saved_model)
print("Model Id: " + str(published_model_id))
```
**Model Id** can be used to retrive latest model version from Watson Machine Learning instance.
Below you can see stored model details.
```
client.repository.get_model_details(published_model_id)
```
### 4.2: Load model
In this subsection you will learn how to load back saved model from specified instance of Watson Machine Learning.
```
loaded_model = client.repository.load(published_model_id)
print(type(loaded_model))
```
As you can see the name is correct. You have already learned how save and load the model from Watson Machine Learning repository.
<a id="visualization"></a>
## 5. Predict locally
In this section you will learn how to score test data using loaded model and visualize the prediction results with plotly package.
### 5.1: Make local prediction using previously loaded model and test data
In this subsection you will score *predict_data* data set.
```
predictions = loaded_model.transform(predict_data)
```
Preview the results by calling the *show()* method on the predictions DataFrame.
```
predictions.show(5)
```
By tabulating a count, you can see which product line is the most popular.
```
predictions.select("predictedLabel").groupBy("predictedLabel").count().show()
```
<a id="scoring"></a>
## 6. Deploy and score
In this section you will learn how to create online scoring and to score a new data record using `ibm-watson-machine-learning`.
**Note:** You can also use REST API to deploy and score.
For more information about REST APIs, see the [Swagger Documentation](https://watson-ml-v4-api.mybluemix.net/wml-restapi-cloud.html#/Deployments/deployments_create).
### 6.1: Create online scoring endpoint
Now you can create an online scoring endpoint.
#### Create online deployment for published model
```
deployment_details = client.deployments.create(
published_model_id,
meta_props={
client.deployments.ConfigurationMetaNames.NAME: "Product Line model deployment",
client.deployments.ConfigurationMetaNames.ONLINE: {}
}
)
deployment_details
```
Now, you can use above scoring url to make requests from your external application.
<a id="cleanup"></a>
## 7. Clean up
If you want to clean up all created assets:
- experiments
- trainings
- pipelines
- model definitions
- models
- functions
- deployments
please follow up this sample [notebook](https://github.com/IBM/watson-machine-learning-samples/blob/master/cpd4.0/notebooks/python_sdk/instance-management/Machine%20Learning%20artifacts%20management.ipynb).
<a id="summary"></a>
## 8. Summary and next steps
You successfully completed this notebook! You learned how to use Apache Spark machine learning as well as Watson Machine Learning for model creation and deployment.
Check out our [Online Documentation](https://dataplatform.cloudibm.com/docs/content/analyze-data/wml-setup.html) for more samples, tutorials, documentation, how-tos, and blog posts.
### Authors
**Amadeusz Masny**, Python Software Developer in Watson Machine Learning at IBM
Copyright © 2020, 2021 IBM. This notebook and its source code are released under the terms of the MIT License.
| github_jupyter |
```
"""
Update Parameters Here
"""
FILE = "Quaks"
ADDRESS = "0xd9d1c2623fbb4377d9bf29075e610a9b8b4805b4" # set to "" if you want to print graphs for all addresses that minted more than MIN_TOKENS_MINTED
TOKEN_COL = "TOKEN_ID"
MIN_TOKENS_MINTED = 25
"""
Optional parameters
Customise these values and set ZOOM_IN to True if you want to zoom in on a specific range
"""
ZOOM_IN = False
LOWER_BOUND = 0
UPPER_BOUND = 6000
TOP_N = 6000
"""
@author: mdigi14
"""
import pandas as pd
import matplotlib.pyplot as plt
import math
from utils import config
"""
Plot params
"""
plt.rcParams.update({"figure.facecolor": "white", "savefig.facecolor": "white"})
"""
Helper Functions
"""
def print_graph(grifter_address, MINTING_DB, RARITY_DB, zoom_in=False):
if zoom_in:
lower_bound = LOWER_BOUND
upper_bound = UPPER_BOUND
top_n = TOP_N
else:
# Count number of tokens in Rarity DB and round up to nearest 100
TOKEN_COUNT = int(math.ceil(max(RARITY_DB[TOKEN_COL] / 100.0)) * 100)
upper_bound = top_n = TOKEN_COUNT
lower_bound = 0
GRIFTER_DB = MINTING_DB[MINTING_DB["to_account"] == grifter_address]
tokens_minted = GRIFTER_DB["TOKEN_ID"]
rarity_ranks = GRIFTER_DB["rank"]
if len(tokens_minted) >= MIN_TOKENS_MINTED or ADDRESS != "":
if TOKEN_COL == "TOKEN_NAME":
RARITY_DB["TOKEN_ID"] = (
RARITY_DB["TOKEN_NAME"].str.split("#").str[1].astype(int)
)
RARITY_DB = RARITY_DB.sort_values("TOKEN_ID")
ax = RARITY_DB.plot.scatter(
x="TOKEN_ID",
y="Rank",
grid=True,
alpha=0.25,
title="{} - {}".format(FILE, grifter_address),
figsize=(14, 7),
)
plt.scatter(x=tokens_minted, y=rarity_ranks, color="black", s=55)
ax.set_xlabel("Token ID")
ax.set_ylabel("Rarity Rank")
plt.xlim(lower_bound, upper_bound)
plt.ylim(0, top_n)
plt.show()
"""
Generate Plot
"""
PATH = f"{config.RARITY_FOLDER}/{FILE}_raritytools.csv"
MINT_PATH = f"{config.MINTING_FOLDER}/{FILE}_minting.csv"
RARITY_DB = pd.read_csv(PATH)
RARITY_DB = RARITY_DB[RARITY_DB["TOKEN_ID"].duplicated() == False]
MINTING_DB = pd.read_csv(MINT_PATH)
addresses = set(MINTING_DB["to_account"].unique())
if ADDRESS != "":
print_graph(ADDRESS, MINTING_DB, RARITY_DB, zoom_in=ZOOM_IN)
else:
for address in addresses:
print_graph(address, MINTING_DB, RARITY_DB, zoom_in=ZOOM_IN)
```
| github_jupyter |
# Create miRNASNP-v3 index for BIGD
Guide to for index file [NGDC](https://bigd.big.ac.cn/standards/dis) Bigsearch system (index.bs)
**index.bs** file structure as following:
```
DB JSON_ENCODED_STRING
ENTRY JSON_ENCODED_STRING
```
```
import json
import uuid
from pymongo import MongoClient
db_dict = {
"id": "mirnasnp",
"title": "miRNASNP-v3",
"url": "http://bioinfo.life.hust.edu.cn/miRNASNP/",
"description": "miRNASNP-v3 is a comprehensive database for SNPs and disease-related variations in miRNAs and miRNA targets",
"basicInfo": "In miRNASNP-v3, 46,826 SNPs in human 1,897 pre-miRNAs (2,624 mature miRNAs) and 7,115,796 SNPs in 3'UTRs of 18,151 genes were characterized. Besides, 505,417 disease-related variations (DRVs) from GWAS, ClinVar and COSMIC were identified in miRNA and gene 3'UTR. Gene enrichment of target gain/loss by variations in miRNA seed region was provided.",
"categories": ["miRNA", "mutation", "disease"],
"species": ["Homo Sapiens"],
"updatedAt": "2015-08-30 11:11:11"
}
class MongoMir:
__mongo = MongoClient("mongodb://username:passwd@ip:port/dbname")
def __init__(self, col_name = 'mirinfo'):
self.__col_name = col_name
def get_data(self, output={}, condition={}):
output['_id'] = 0
mcur = self.__mongo.mirnasnp[self.__col_name].find(
condition, output, no_cursor_timeout=True
)
return mcur.count()
def get_mirnas(self):
mcur = self.__mongo.mirnasnp.pri_mir_summary.find(
{}, {'_id': 0, 'mir_id': 1, 'mir_chr': 1, 'mir_start': 1, 'mir_end': 1}
)
# res = [{'mir_id': item['mir_id'], 'loci': f"{item['mir_chr']}:{item['mir_start']}-{item['mir_end']}"} for item in mcur]
res = [item['mir_id'] for item in mcur]
return res
def get_genes(self):
mcur = self.__mongo.mirnasnp.mutation_summary_genelist.find(
{}, {'_id': 0, 'gene_symbol': 1}
)
m_symbol = set([item['gene_symbol'] for item in mcur])
mcur = self.__mongo.mirnasnp.snp_summary_genelist.find(
{}, {'_id': 0, 'gene_symbol': 1}
)
s_symbol = set([item['gene_symbol'] for item in mcur])
return list(m_symbol.union(s_symbol))
class ENTRY(object):
def __init__(self, type, title, url):
self.id = str(uuid.uuid4())
self.type = type
self.title = title
self.url = url
self.dbId = "mirnasnp"
self.updatedAt = "2015-08-30 11:11:11"
self.description = ""
self.basicInfo = ""
self.species = ["Homo Sapiens"]
self.attrs = {
"symbol": title,
}
def __getattr__(self, attr):
return self[attr]
def get_entry(it, type = 'miRNA ID'):
if type.startswith('miRNA'):
url = f'http://bioinfo.life.hust.edu.cn/miRNASNP/#!/mirna?mirna_id={it}'
else:
url = f'http://bioinfo.life.hust.edu.cn/miRNASNP/#!/gene?query_gene={it}&has_snp=1&has_phenotype=1'
e = ENTRY(type, it, url)
return json.dumps(e.__dict__)
mongo_mirnasnp = MongoMir()
mirna_ids = mongo_mirnasnp.get_mirnas()
gene_ids = mongo_mirnasnp.get_genes()
with open('/home/liucj/tmp/index.bs', 'w') as fh:
header = 'DB' + '\t' + json.dumps(db_dict) + '\n'
fh.write(header)
for it in mirna_ids:
line = 'ENTRY' + '\t' + get_entry(it = it, type = 'miRNA ID') + '\n'
fh.write(line)
for it in gene_ids:
line = 'ENTRY' + '\t' + get_entry(it = it, type = 'Official gene symbol') + '\n'
fh.write(line)
```
Check the index.bs file
```
/home/miaoyr/software/BSChecker/bschecker-1.1.4-bin/bin/bschecker /home/liucj/tmp/index.bs
```
| github_jupyter |
# MODULE 1 - Basic Python programming: IF statements
### Simple example using several concepts
Use for loop and nested if to find member of list (if any)
```
# Define list of automobiles
cars = ['honda', 'mini', 'subaru', 'toyota']
# Loop through all items in list
for car in cars: # car is loop variable containing current item in list
if car == 'mini':
print(car.upper()) # use upper() method function
else:
print(car.title()) # use title() method function to capitalize 1st letter
```
### Evaluating logical expressions
```
car="mini"
car=="mini" # Logical expression test for equality with ==
car="honda"
car=="mini" # Logical expression evaluates to False when not equal
car="mini"
car=="Mini" # Not equal since different case
car="Mini"
car.lower()=="mini" # Convert car variable to lower case before comparison
#print(car) # Use of lower() method does not alter value of car variable
# Mini
```
### Use if statement to alter flow-of-control
```
additional_item = "pickles"
if additional_item != "mushrooms":
print("Hold the mushrooms please!")
number_bedrooms = 3 # Logical expression with numeric values
number_bedrooms == 3
quiz_answer = 3
# logical expression determines flow
if quiz_answer != 5:
print("That is not the correct answer. Please try again!")
age = 25
age < 30 # Less than operator <
age <= 26 # Less than or equal to operator <=
age > 42 # Greater than operator >
age >= 33 # Greater than or equal to operator >=
```
### Evaluating more complex conditions
```
number_bedrooms = 2
square_ft = 1800
number_bedrooms >= 2 and square_ft >= 2400 # Using the "and" logical operator
square_ft = 2650
number_bedrooms >= 2 and square_ft >= 2400 # Using the "and" logical operator
number_bedrooms = 2
square_ft = 1800
number_bedrooms >= 2 or square_ft >= 2400 # Using the "or" logical operator
number_bedrooms = 1
number_bedrooms >= 2 or square_ft >= 2000 # Using the "or" logical operator
```
### Checking if a value is in a list
```
PAC_12 = ['UCLA', 'Arizona', 'Oregon', "Stanford", "UC Berkeley", "USC", "Washington"]
print('UCLA' in PAC_12)
print('MIT' in PAC_12)
# Now check if not in a list
banned_users = ['ellen', 'catherine', 'stephen']
user='marie'
if user not in banned_users:
print(f"{user.title()}, you can post a response if you wish.")
```
### Boolean values: True, False
```
face_image_recognized = True
if face_image_recognized:
print("image recognition complete!")
face_image_recognized = False
if not face_image_recognized: # Use "not" logical operator
print("sorry, face not recognized")
```
### Using if-else and if-elif-else statements
```
age = 19
if age >= 18:
print("You are welcome to vote!")
age = 19
if age >= 18: # You can have multple lines in the block after the if statement
print("You are welcome to vote!")
print("Please be sure to register")
```
Can also have an if ... else statement
```
age = 17
if age >= 18: # Block of code if logical expression evaluates to True
print("You are old enough to vote!")
print("Have you registered to vote yet?")
else: # Block of code if logical expression evaluates to False
print("Sorry, you are too young to vote.")
print("Please register to vote as soon as you turn 18!")
```
Can also use an if-elif-else chain
```
age = 12
if age < 4: # NOTE: indenting matters! Syntax errors otherwise
print("Your admission is free of charge")
elif age < 18:
print("Your admission is $10")
else:
print("Your admisstion is 25")
# Assign value to variable used with if below to control flow-of-control
age = 12
if age < 4:
price = 0
elif age < 18:
price = 25
else:
price = 40
# Use print functions in code for debugging purposes
print(f"Your admission cost is ${price}.")
age = 32
if age < 4:
price = 0
elif age < 18:
price = 25
elif age < 65:
price = 40
else:
price = 20
# Use print functions in code for debugging purposes
print(f"Your admission cost is ${price}.")
```
Use if to implement "case" statement
```
# Assign value to variable used with if below to control flow-of-control
age = 12
if age < 4:
price = 0
elif age < 18:
price = 25
elif age < 65:
price = 40
elif age >= 65:
price = 20
# Use print functions in code for debugging purposes
print(f"Your admission cost is ${price}.")
```
### Testing multiple conditions
```
# if-elif-else chain is useful, but only if you need just one test to pass.
# Sometimes it's important to check all of the conditions as below:
# More than one condition could be True
# Below would NOT work if we used if-elif chain!
requested_toppings = ["mushrooms", "extra cheese"]
if "mushrooms" in requested_toppings:
print("Adding mushrooms.")
if "pepperoni" in requested_toppings:
print("Adding pepperoni.")
if "extra cheese" in requested_toppings:
print("Adding extra cheese")
print("\nFinished making your pizza")
```
### Checking for special items
```
requested_toppings = ['mushrooms', 'green peppers', 'extra cheese']
# Loop through all members of requested_toppings list
for requested_topping in requested_toppings:
print(f"Adding {requested_topping}.")
print("\nFinished making your pizza!")
requested_toppings = ['mushrooms', 'green peppers', 'extra cheese']
# Loop through all members of requested_toppings list
for requested_topping in requested_toppings:
if requested_topping == "green peppers":
print("Sorry, we are out of green peppers right now.")
else:
print(f"Adding {requested_topping}.")
print("\nFinished making your pizza!")
requested_toppings = []
if requested_toppings:
for requested_topping in requested_toppings:
print(f"Adding {requested_topping}.")
print("\nFinished making your pizza!")
else:
print("Are you sure you want a plain pizza?")
available_toppings = ['mushrooms', 'olives', 'green peppers',
'pepperoni', 'pineapple', 'extra cheese']
requested_toppings = ['mushrooms', 'french fries', 'extra cheese']
# Loop through all members of requested_toppings list
for requested_topping in requested_toppings:
if requested_topping in available_toppings:
print(f"Adding {requested_topping}.")
else:
print(f"Sorry, we don't have {requested_topping}.")
print("\nFinished making your pizza!")
```
| github_jupyter |
```
import pandas as pd
from collections import Counter
import seaborn as sns
import umap
import matplotlib.pyplot as plt
import texthero as hero
import polarice
import polarice.preprocessing
from gensim.models import KeyedVectors
import glob
import numpy as np
np.random.seed(42)
data_dir = "./data/framing/elites_tweets/"
limit_top_w2v = 500_000
all_files = glob.glob(data_dir + "*.jsonl")
li = []
for filename in all_files:
df = pd.read_json(filename, lines=True)
li.append(df)
frame = pd.concat(li, axis=0, ignore_index=True)
elites_df = pd.read_csv(data_dir + "../input/elites-data.csv")
elites_df.info()
frame["user_id"] = frame["user"].map(lambda x: x["id"])
frame.info()
frame["user_id"].nunique()
labelled_tweets = frame.merge(elites_df, left_on="user_id", right_on="twitter_id")
german_tweet_pipeline = polarice.preprocessing.ENGLISH_TWEET_PIPELINE
labelled_tweets["cleaned"] = labelled_tweets["full_text"].pipe(hero.clean, pipeline=german_tweet_pipeline)
labelled_tweets # output clear for privacy
model = KeyedVectors.load_word2vec_format("glove.840B.300d_w2v.txt", binary=False, limit=limit_top_w2v)
%run frame_axis.py
fs = FrameSystem.load("moral.pkl")
fs.attach_model(model)
fs.compute()
fs.frame_axes
%%time
fs.axes_ordered_by_effect_sizes(labelled_tweets["cleaned"], num_bootstrap_samples=1)
%%time
trans_df = fs.transform_df(labelled_tweets, "cleaned", model)
trans_df # output clear for privacy
bias_cols = [col for col in labelled_tweets if col.endswith("_bias")]
intensity_cols = [col for col in labelled_tweets if col.endswith("_inte")]
moral_cols = bias_cols + intensity_cols
cm = sns.light_palette("green", as_cmap=True)
groups = labelled_tweets.groupby("party")[moral_cols]
group_moral = groups.mean()
latex_df = group_moral.transpose()
latex_df.index = pd.MultiIndex.from_tuples(latex_df.index.str.split('_').tolist())
latex_df = latex_df.swaplevel(0, 1, 0)
latex_df = latex_df.rename(index={
"care": "Care",
"fair": "Fairness",
"loya": "Loyalty",
"auth": "Authority",
"sanc": "Sanctity",
"bias": "Bias",
"intensity": "Intensity"
})
latex_df = latex_df * 10 # for easier readability
print(latex_df.to_latex(multirow=True, float_format="%.3f"))
latex_df.style.background_gradient(cmap=cm, axis=1)
```
# Classification
```
y_label = "party"
bias_cols = [col for col in labelled_tweets if col.endswith("_bias")]
intensity_cols = [col for col in labelled_tweets if col.endswith("_inte")]
moral_cols = bias_cols + intensity_cols
Y = labelled_tweets[y_label].values
X = labelled_tweets[moral_cols].values
print(X.shape)
print(np.expand_dims(Y, axis=1).shape)
np.concatenate([X, np.expand_dims(Y, axis=1)], axis=1)
from sklearn.linear_model import LogisticRegression
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
from sklearn.metrics import classification_report
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size=0.25, shuffle=True)
classifier = LogisticRegression(max_iter=1000).fit(X_train, Y_train)
Y_pred = classifier.predict(X_test)
classification_report(Y_test, Y_pred, output_dict=True)
print(classification_report(Y_test, Y_pred))
classifier.coef_
coef_df = pd.DataFrame.from_records(classifier.coef_, index=classifier.classes_, columns=moral_cols)
print(coef_df.to_latex())
coef_df
coef_df = coef_df.stack().to_frame().reset_index()
coef_df[["moral", "bias/inte"]] = coef_df["level_1"].str.split("_", expand=True)
coef_df = coef_df.drop(columns=["level_1"])
coef_df = coef_df.rename(columns={"level_0": "handle", 0: "coef"})
coef_df
bias_df = coef_df[coef_df["bias/inte"] == "bias"]
sns.pointplot(data=bias_df, x="moral", y="coef", hue="handle")
inte_df = coef_df[coef_df["bias/inte"] == "inte"]
sns.pointplot(data=inte_df, x="moral", y="coef", hue="handle")
# pairplot on traindata: https://scikit-learn.org/stable/auto_examples/inspection/plot_linear_model_coefficient_interpretation.html
dataset = labelled_tweets[[y_label] + moral_cols].copy()
dataset # output clear for privacy
ds = dataset.sample(10_000)
%%time
g = sns.pairplot(ds, kind="reg", diag_kind="kde", hue="party")
g.map_lower(sns.kdeplot, levels=4, color=".2")
plt.savefig("pairplot_us.pdf", dpi=300)
plt.show()
```
| github_jupyter |
# Tensor Manipulation: Psi4 and NumPy manipulation routines
Contracting tensors together forms the core of the Psi4Julia project. First let us consider the popluar [Einstein Summation Notation](https://en.wikipedia.org/wiki/Einstein_notation) which allows for very succinct descriptions of a given tensor contraction.
For example, let us consider a [inner (dot) product](https://en.wikipedia.org/wiki/Dot_product):
$$c = \sum_{ij} A_{ij} * B_{ij}$$
With the Einstein convention, all indices that are repeated are considered summed over, and the explicit summation symbol is dropped:
$$c = A_{ij} * B_{ij}$$
This can be extended to [matrix multiplication](https://en.wikipedia.org/wiki/Matrix_multiplication):
\begin{align}
\rm{Conventional}\;\;\; C_{ik} &= \sum_{j} A_{ij} * B_{jk} \\
\rm{Einstein}\;\;\; C &= A_{ij} * B_{jk} \\
\end{align}
Where the $C$ matrix has *implied* indices of $C_{ik}$ as the only repeated index is $j$.
However, there are many cases where this notation fails. Thus we often use the generalized Einstein convention. To demonstrate let us examine a [Hadamard product](https://en.wikipedia.org/wiki/Hadamard_product_(matrices)):
$$C_{ij} = \sum_{ij} A_{ij} * B_{ij}$$
This operation is nearly identical to the dot product above, and is not able to be written in pure Einstein convention. The generalized convention allows for the use of indices on the left hand side of the equation:
$$C_{ij} = A_{ij} * B_{ij}$$
Usually it should be apparent within the context the exact meaning of a given expression.
Finally we also make use of Matrix notation:
\begin{align}
{\rm Matrix}\;\;\; \bf{D} &= \bf{A B C} \\
{\rm Einstein}\;\;\; D_{il} &= A_{ij} B_{jk} C_{kl}
\end{align}
Note that this notation is signified by the use of bold characters to denote matrices and consecutive matrices next to each other imply a chain of matrix multiplications!
## Tensor Operations
To perform most operations we turn to tensor packages (here we use [TensorOperations.jl](https://github.com/Jutho/TensorOperations.jl)). Those allow Einstein convention as an input. In addition to being much easier to read, manipulate, and change, it has (usually) optimal performance.
First let us import our normal suite of modules:
```
using PyCall: pyimport
psi4 = pyimport("psi4")
np = pyimport("numpy")
using TensorOperations: @tensor
```
We can then use conventional Julia loops or `@tensor` to perform the same task.
```
using BenchmarkTools: @btime, @belapsed
```
With `@btime`/`@belapsed` we average time over several executions to have more reliable timings than `@time`/`@elapsed` (single execution).
<font color="red">**WARNING: We are using Julia's global variables, and those are known to be less efficient than local variables. It is better to wrap code inside function.**</font>
To begin let us consider the construction of the following tensor (which you may recognize):
$$G_{pq} = 2.0 * I_{pqrs} D_{rs} - 1.0 * I_{prqs} D_{rs}$$
Keep size relatively small as these 4-index tensors grow very quickly in size.
```
dims = 20
@assert dims <= 30 "Size must be smaller than 30."
D = rand(dims, dims)
I = rand(dims, dims, dims, dims)
# Build the Fock matrix using loops, while keeping track of time
println("Time for loop G build:")
Gloop = @btime begin
Gloop = np.zeros((dims, dims))
@inbounds for ind in CartesianIndices(I)
p, q, r, s = Tuple(ind)
Gloop[p, q] += 2I[p, q, r, s] * D[r, s]
Gloop[p, q] -= I[p, r, q, s] * D[r, s]
end
Gloop
end
# Build the Fock matrix using einsum, while keeping track of time
println("Time for @tensor G build:")
G = @btime @tensor G[p,q] := 2I[p,q,r,s] * D[r,s] - I[p,r,q,s] * D[r,s]
# Make sure the correct answer is obtained
println("Loop and einsum builds of the Fock matrix match? ", np.allclose(G, Gloop))
println()
# Print out relative times for explicit loop vs einsum Fock builds
#println("G builds with einsum are $(g_loop_time/einsum_time) times faster than Julia loops!")
```
As you can see, the `@tensor` macro can be considerably faster than plain Julia loops.
## Matrix multiplication chain/train
Now let us turn our attention to a more canonical matrix multiplication example such as:
$$D_{il} = A_{ij} B_{jk} C_{kl}$$
Matrix multiplication is an extremely common operation in all branches of linear algebra. Thus, these functions have been optimized to be extremely efficient. `@tensor` uses it. The matrix product will explicitly compute the following operation:
$$C_{ij} = A_{ij} * B_{ij}$$
This is Julia's matrix multiplication method `*` for matrices.
```
dims = 200
A = rand(dims, dims)
B = rand(dims, dims)
C = rand(dims, dims)
# First compute the pair product
tmp_dot = A * B
@tensor tmp_tensor[i,k] := A[i,j] * B[j,k]
println("Pair product allclose? ", np.allclose(tmp_dot, tmp_tensor))
```
Now that we have proved exactly what `*` product does, let us consider the full chain and do a timing comparison:
```
D_dot = A * B * C
@tensor D_tensor[i,l] := A[i,j] * B[j,k] * C[k,l]
println("Chain multiplication allclose? ", np.allclose(D_dot, D_tensor))
println()
println("* time:")
@btime A * B * C
println()
println("@tensor time:")
@btime @tensor D_tensor[i,l] := A[i,j] * B[j,k] * C[k,l];
```
Both have similar timings, and both call [Basic Linear Algebra Subprograms (BLAS)](https://en.wikipedia.org/wiki/Basic_Linear_Algebra_Subprograms). The BLAS routines are highly optimized and threaded versions of the code.
- The `@tensor` code will factorize the operation by default; Thus, the overall cost is not ${\cal O}(N^4)$ (as there are four indices) rather it is the factored $(\bf{A B}) \bf{C}$ which runs ${\cal O}(N^3)$.
Therefore you do not need to factorize the expression yourself (sometimes you might need):
```
println("@tensor factorized time:")
@btime @tensor begin
tmp[i,k] := A[i,j] * B[j,k]
tmp2[i,l] := tmp[i,k] * C[k,l]
end
nothing
```
On most machines the three have similar timings. The BLAS usage is usually recommended. Thankfully, in Julia its syntax is very clear. The Psi4Julia project tends to lean toward usage of tensor packages but if Julia's built-in matrix multiplication is significantly cleaner/faster we would use it. The real value of tensor packages will become tangible for more complicated expressions.
## Complicated tensor manipulations
Let us consider a popular index transformation example:
$$M_{pqrs} = C_{pi} C_{qj} I_{ijkl} C_{rk} C_{sl}$$
Here, a naive loop implementation would scale like $\mathcal{O}(N^8)$ which translates to an extremely costly computation for all but the smallest $N$. A smarter implementation (factorizing the whole expression) would scale
like $\mathcal{O}(N^5)$.
<font color="red">**WARNING: First execution is slow because of compilation time. Successive are more honest to the running time.**</font>
```
# Grab orbitals
dims = 15
@assert dims <= 15 || "Size must be smaller than 15."
C = rand(dims, dims)
I = rand(dims, dims, dims, dims)
# @tensor full transformation.
print("\nStarting @tensor full transformation...")
n8_time = @elapsed @tensor MO_n8[I,J,K,L] := C[p,I] * C[q,J] * I[p,q,r,s] * C[r,K] * C[s,L]
print("complete in $n8_time s\n")
# @tensor factorized N^5 transformation.
print("\nStarting @tensor factorized N^5 transformation with einsum ... ")
n5_time = @elapsed @tensor begin
MO_n5[A,q,r,s] := C[p,A] * I[p,q,r,s]
MO_n5[A,B,r,s] := C[q,B] * MO_n5[A,q,r,s]
MO_n5[A,B,C,s] := C[r,C] * MO_n5[A,B,r,s]
MO_n5[A,B,C,D] := C[s,D] * MO_n5[A,B,C,s]
end
print("complete in $n5_time s \n")
println(" @tensor factorized is $(n8_time/n5_time) faster than full @tensor algorithm!")
println(" Allclose? ", np.allclose(MO_n8, MO_n5))
# Julia's GEMM N^5 transformation.
# Try to figure this one out!
print("\nStarting Julia's factorized transformation with * ... ")
dgemm_time = @elapsed begin
MO = C' * reshape(I, dims, :)
MO = reshape(MO, :, dims) * C
MO = permutedims(reshape(MO, dims, dims, dims, dims), (2, 1, 4, 3))
MO = C' * reshape(MO, dims, :)
MO = reshape(MO, :, dims) * C
MO = permutedims(reshape(MO, dims, dims, dims, dims),(2, 1, 4, 3))
end
print("complete in $dgemm_time s \n")
println(" * factorized is $(n8_time/dgemm_time) faster than full @tensor algorithm!")
println(" Allclose? ", np.allclose(MO_n8, MO))
# There are still several possibilities to explore:
# @inbounds, @simd, LinearAlgebra.LAPACK calls, Einsum.jl, Tullio.jl, ...
```
None of the above algorithms is $\mathcal{O}(N^8)$. `@tensor` factorizes the expression to achieve better performance. There is a small edge in doing the factorization manually. Factorized algorithms have similar timings, although it is clear that with `@tensor` is easier than with Julia's built-in `*`. To use the usual matrix multiplication with tensors we have to reshape and permute their dimensions, subtracting appeal to the simple `*` syntax.
| github_jupyter |
# Route optimization of a pub crawl with ORS and `ortools`
It's this of the year again (or will be in 6 months): the freshmen pour into the institute and as the diligent student council you are, you want to welcome them for their geo adventure with a stately pub crawl to prepare them for the challenges lying ahead.
We want to give you the opportunity to route the pack of rookies in a fairly optimal way:
```
mkdir ors-pubcrawl
conda create -n ors-pubcrawl python=3.6 folium shapely
cd ors-pubcrawl
pip install openrouteservice ortools
import folium
from shapely import wkt, geometry
import json
from pprint import pprint
```
Now we're ready to start our most optimally planned pub crawl ever through hipster Kreuzberg! It will also be the most un-hipster pub crawl ever, as we'll cover ground with a taxi. At least it's safer than biking half-delirious.
First the basic parameters: API key and the district polygon to limit our pub search. The Well Known Text was prepared in QGIS from Berlin authority's [WFS](http://fbinter.stadt-berlin.de/fb/wfs/geometry/senstadt/re_ortsteil/) (QGIS field calculator has a `geom_to_wkt` method). BTW, Berlin, hope you don't wonder why your feature services are so slow... Simplify is the magic word, simplify.
```
api_key = 'your_key'
wkt_str = 'Polygon ((13.43926404 52.48961046, 13.42040115 52.49586382, 13.42541101 52.48808523, 13.42368155 52.48635829, 13.40788599 52.48886084, 13.40852944 52.487142, 13.40745989 52.48614988, 13.40439187 52.48499746, 13.40154731 52.48500125, 13.40038591 52.48373202, 13.39423818 52.4838664, 13.39425346 52.48577149, 13.38629096 52.48582648, 13.38626853 52.48486362, 13.3715694 52.48495055, 13.37402099 52.4851697, 13.37416365 52.48771105, 13.37353615 52.48798191, 13.37539925 52.489432, 13.37643416 52.49167597, 13.36821531 52.49333093, 13.36952826 52.49886974, 13.37360623 52.50416333, 13.37497726 52.50337776, 13.37764916 52.5079675, 13.37893813 52.50693045, 13.39923153 52.50807711, 13.40022883 52.50938108, 13.40443425 52.50777471, 13.4052848 52.50821063, 13.40802944 52.50618019, 13.40997081 52.50692569, 13.41152096 52.50489127, 13.41407284 52.50403794, 13.41490921 52.50491634, 13.41760145 52.50417013, 13.41943091 52.50564912, 13.4230412 52.50498109, 13.42720031 52.50566607, 13.42940229 52.50857222, 13.45335235 52.49752496, 13.45090795 52.49710803, 13.44765912 52.49472124, 13.44497623 52.49442276, 13.43926404 52.48961046))'
aoi_geom = wkt.loads(wkt_str) # load geometry from WKT string
aoi_coords = list(aoi_geom.exterior.coords) # get coords from exterior ring
aoi_coords = [(y,x) for x,y in aoi_coords] # swap (x,y) to (y,x). Really leaflet?!
aoi_centroid = aoi_geom.centroid # Kreuzberg center for map center
```
Next, add the Kreuzberg polygon as marker to the map, so we get a bit of orientation.
```
m = folium.Map(tiles='Stamen Toner',location=(aoi_centroid.y, aoi_centroid.x), zoom_start=14)
folium.vector_layers.Polygon(aoi_coords,
color='#ffd699',
fill_color='#ffd699',
fill_opacity=0.2,
weight=3).add_to(m)
m
```
Now it's time to see which are the lucky bars to host a bunch of increasingly drunk geos. We use the [**Places API**](https://openrouteservice.org/documentation/#/reference/places/places/location-service-(get), where we can pass a GeoJSON as object right into. As we want to crawl only bars and not churches, we have to limit the query to category ID's which represent pubs. We can get the mapping easily when passing `category_list`:
```
from openrouteservice import client, places
clnt = client.Client(key=api_key)
```
[**Here**](https://github.com/GIScience/openrouteservice-docs#places-response) is a nicer list. If you look for pub, you'll find it under `sustenance : 560` with ID 569. Chucking that into a query, yields:
```
aoi_json = geometry.mapping(geometry.shape(aoi_geom))
query = {'request': 'pois',
'geojson': aoi_json,
'filter_category_ids': [569],
'sortby': 'distance'}
pubs = clnt.places(**query)['features'] # Perform the actual request and get inner json
# Amount of pubs in Kreuzberg
print("\nAmount of pubs: {}".format(len(pubs)))
```
107 bars in one night might be a stretch, even for such a resilient species. Coincidentally, the rate of smokers is unproportionally high within the undergrad geo community. So, we really would like to hang out in smoker bars:
```
query['filters_custom'] = {'smoking':['yes']} # Filter out smoker bars
pubs_smoker = clnt.places(**query)['features']
print("\nAmount of smoker pubs: {}".format(len(pubs_smoker)))
```
A bit better. Let's see where they are.
**Optionally**, use the [**Geocoding API**](https://openrouteservice.org/documentation/#/reference/places/places/location-service-(get) to get representable names. Note, it'll be 25 API calls. Means, you can only run one per minute.
```
from openrouteservice import geocode
pubs_addresses = []
for feat in pubs_smoker:
lon, lat = feat['geometry']['coordinates']
name = clnt.pelias_reverse(point=(lon, lat))['features'][0]['properties']['name']
popup = "<strong>{0}</strong><br>Lat: {1:.3f}<br>Long: {2:.3f}".format(name, lat, lon)
icon = folium.map.Icon(color='lightgray',
icon_color='#b5231a',
icon='beer', # fetches font-awesome.io symbols
prefix='fa')
folium.map.Marker([lat, lon], icon=icon, popup=popup).add_to(m)
pubs_addresses.append(name)
# folium.map.LayerControl().add_to(m)
m
```
Ok, we have an idea where we go. But not in which order. To determine the optimal route, we first have to know the distance between all pubs. We can conveniently solve this with the [**Matrix API**](https://openrouteservice.org/documentation/#/reference/places/places/location-service-(get).
>I'd have like to do this example for biking/walking, but I realized too late that we restricted matrix calls to 5x5 locations for those profiles...
```
from openrouteservice import distance_matrix
pubs_coords = [feat['geometry']['coordinates'] for feat in pubs_smoker]
request = {'locations': pubs_coords,
'profile': 'driving-car',
'metrics': ['duration']}
pubs_matrix = clnt.distance_matrix(**request)
print("Calculated {}x{} routes.".format(len(pubs_matrix['durations']),len(pubs_matrix['durations'][0])))
```
Check, 26x26. So, we got the durations now in `pubs_matrix['durations']`. Then there's finally the great entrance of [**ortools**](https://github.com/google/or-tools).
Note, this is a local search.
```
from ortools.constraint_solver import pywrapcp
from ortools.constraint_solver import routing_enums_pb2
def getDistance(from_id, to_id):
return int(pubs_matrix['durations'][from_id][to_id])
tsp_size = len(pubs_addresses)
num_routes = 1
start = 0 # arbitrary start location
coords_aoi = [(y,x) for x,y in aoi_coords] # swap (x,y) to (y,x)
optimal_coords = []
if tsp_size > 0:
routing = pywrapcp.RoutingModel(tsp_size, num_routes, start)
search_parameters = pywrapcp.RoutingModel.DefaultSearchParameters()
# Create the distance callback, which takes two arguments (the from and to node indices)
# and returns the distance between these nodes.
dist_callback = getDistance
routing.SetArcCostEvaluatorOfAllVehicles(dist_callback)
# Solve, returns a solution if any.
assignment = routing.SolveWithParameters(search_parameters)
if assignment:
# Total cost of the 'optimal' solution.
print("Total duration: " + str(round(assignment.ObjectiveValue(), 3) / 60) + " minutes\n")
index = routing.Start(start) # Index of the variable for the starting node.
route = ''
# while not routing.IsEnd(index):
for node in range(routing.nodes()):
optimal_coords.append(pubs_coords[routing.IndexToNode(index)])
route += str(pubs_addresses[routing.IndexToNode(index)]) + ' -> '
index = assignment.Value(routing.NextVar(index))
route += str(pubs_addresses[routing.IndexToNode(index)])
optimal_coords.append(pubs_coords[routing.IndexToNode(index)])
print("Route:\n" + route)
```
Visualizing both, the optimal route, and the more or less random waypoint order of the intial GeoJSON, look like this:
```
from openrouteservice import directions
import os.path
def style_function(color):
return lambda feature: dict(color=color,
weight=3,
opacity=1)
# See what a 'random' tour would have been
pubs_coords.append(pubs_coords[0])
request = {'coordinates': pubs_coords,
'profile': 'driving-car',
'geometry': 'true',
'format_out': 'geojson',
# 'instructions': 'false'
}
random_route = clnt.directions(**request)
folium.features.GeoJson(data=random_route,
name='Random Bar Crawl',
style_function=style_function('#84e184'),
overlay=True).add_to(m)
# And now the optimal route
request['coordinates'] = optimal_coords
optimal_route = clnt.directions(**request)
folium.features.GeoJson(data=optimal_route,
name='Optimal Bar Crawl',
style_function=style_function('#6666ff'),
overlay=True).add_to(m)
m.add_child(folium.map.LayerControl())
m
```
The purple route looks a bit less painful. Let's see what the actual numbers say:
```
optimal_duration = 0
random_duration = 0
optimal_duration = optimal_route['features'][0]['properties']['summary']['duration'] / 60
random_duration = random_route['features'][0]['properties']['summary']['duration'] / 60
print("Duration optimal route: {0:.3f} mins\nDuration random route: {1:.3f} mins".format(optimal_duration,
random_duration))
```
Optimizing that route saved us a good 120€ worth of taxi costs.
| github_jupyter |
# Loops
## for Loops
A [for loop](https://docs.python.org/3/reference/compound_stmts.html#for) allows us to execute a block of code multiple times with some parameters updated each time through the loop. A `for` loop begins with the `for` statement:
```
iterable = [1,2,3]
for item in iterable:
# code block indented 4 spaces
print(item)
```
The main points to observe are:
* `for` and `in` keywords
* `iterable` is a sequence object such as a list, tuple or range
* `item` is a variable which takes each value in `iterable`
* end `for` statement with a colon `:`
* code block indented 4 spaces which executes once for each value in `iterable`
For example, let's print $n^2$ for $n$ from 0 to 5:
```
for n in [0,1,2,3,4,5]:
square = n**2
print(n,'squared is',square)
print('The for loop is complete!')
```
Copy and paste this code and any of the examples below into the [Python visualizer](http://www.pythontutor.com/visualize.html#mode=edit) to see each step in a `for` loop!
## while Loops
What if we want to execute a block of code multiple times but we don't know exactly how many times? We can't write a `for` loop because this requires us to set the length of the loop in advance. This is a situation when a [while loop](https://en.wikipedia.org/wiki/While_loop#Python) is useful.
The following example illustrates a [while loop](https://docs.python.org/3/tutorial/introduction.html#first-steps-towards-programming):
```
n = 5
while n > 0:
print(n)
n = n - 1
```
The main points to observe are:
* `while` keyword
* a logical expression followed by a colon `:`
* loop executes its code block if the logical expression evaluates to `True`
* update the variable in the logical expression each time through the loop
* **BEWARE!** If the logical expression *always* evaluates to `True`, then you get an [infinite loop](https://en.wikipedia.org/wiki/While_loop#Python)!
We prefer `for` loops over `while` loops because of the last point. A `for` loop will never result in an infinite loop. If a loop can be constructed with `for` or `while`, we'll always choose `for`.
## Constructing Sequences
There are several ways to construct a sequence of values and to save them as a Python list. We have already seen Python's list comprehension syntax. There is also the `append` list method described below.
### Sequences by a Formula
If a sequence is given by a formula then we can use a list comprehension to construct it. For example, the sequence of squares from 1 to 100 can be constructed using a list comprehension:
```
squares = [d**2 for d in range(1,11)]
print(squares)
```
However, we can achieve the same result with a `for` loop and the `append` method for lists:
```
# Intialize an empty list
squares = []
for d in range(1,11):
# Append the next square to the list
squares.append(d**2)
print(squares)
```
In fact, the two examples above are equivalent. The purpose of list comprehensions is to simplify and compress the syntax into a one-line construction.
### Recursive Sequences
We can only use a list comprehension to construct a sequence when the sequence values are defined by a formula. But what if we want to construct a sequence where the next value depends on previous values? This is called a [recursive sequence](https://en.wikipedia.org/wiki/Recursion).
For example, consider the [Fibonacci sequence](https://en.wikipedia.org/wiki/Fibonacci_number):
$$
x_1 = 1, x_2 = 1, x_3 = 2, x_4 = 3, x_5 = 5, ...
$$
where
$$
x_{n} = x_{n-1} + x_{n-2}
$$
We can't use a list comprehension to build the list of Fibonacci numbers, and so we must use a `for` loop with the `append` method instead. For example, the first 15 Fibonacci numbers are:
```
fibonacci_numbers = [1,1]
for n in range(2,15):
fibonacci_n = fibonacci_numbers[n-1] + fibonacci_numbers[n-2]
fibonacci_numbers.append(fibonacci_n)
print(fibonacci_numbers)
```
## Computing Sums
Suppose we want to compute the sum of a sequence of numbers $x_0$, $x_1$, $x_2$, $x_3$, $\dots$, $x_n$. There are at least two approaches:
1. Compute the entire sequence, store it as a list $[x_0,x_1,x_2,\dots,x_n]$ and then use the built-in function `sum`.
2. Initialize a variable with value 0 (and name it `result` for example), create and add each element in the sequence to `result` one at a time.
The advantage of the second approach is that we don't need to store all the values at once. For example, here are two ways to write a function which computes the sum of squares.
For the first approach, use a list comprehension:
```
def sum_of_squares_1(N):
"Compute the sum of squares 1**2 + 2**2 + ... + N**2."
return sum([n**2 for n in range(1,N + 1)])
sum_of_squares_1(4)
```
For the second approach, use a `for` loop with the initialize-and-update construction:
```
def sum_of_squares_2(N):
"Compute the sum of squares 1**2 + 2**2 + ... + N**2."
# Initialize the output value to 0
result = 0
for n in range(1,N + 1):
# Update the result by adding the next term
result = result + n**2
return result
sum_of_squares_2(4)
```
Again, both methods yield the same result however the second uses less memory!
## Computing Products
There is no built-in function to compute products of sequences therefore we'll use an initialize-and-update construction similar to the example above for computing sums.
Write a function called `factorial` which takes a positive integer $N$ and return the factorial $N!$.
```
def factorial(N):
"Compute N! = N(N-1) ... (2)(1) for N >= 1."
# Initialize the output variable to 1
product = 1
for n in range(2,N + 1):
# Update the output variable
product = product * n
return product
```
Let's test our function for input values for which we know the result:
```
factorial(2)
factorial(5)
```
We can use our function to approximate $e$ using the Taylor series for $e^x$:
$$
e^x = \sum_{k=0}^{\infty} \frac{x^k}{k!}
$$
For example, let's compute the 100th partial sum of the series with $x=1$:
```
sum([1/factorial(k) for k in range(0,101)])
```
## Searching for Solutions
We can use `for` loops to search for integer solutions of equations. For example, suppose we would like to find all representations of a positive integer $N$ as a [sum of two squares](https://en.wikipedia.org/wiki/Sum_of_two_squares_theorem). In other words, we want to find all integer solutions $(x,y)$ of the equation:
$$
x^2 + y^2 = N
$$
Write a function called `reps_sum_squares` which takes an integer $N$ and finds all representations of $N$ as a sum of squares $x^2 + y^2 = N$ for $0 \leq x \leq y$. The function returns the representations as a list of tuples. For example, if $N = 50$ then $1^2 + 7^2 = 50$ and $5^2 + 5^2 = 50$ and the function returns the list `[(1, 7),(5, 5)]`.
Let's outline our approach before we write any code:
1. Given $x \leq y$, the largest possible value for $x$ is $\sqrt{\frac{N}{2}}$
2. For $x \leq \sqrt{\frac{N}{2}}$, the pair $(x,y)$ is a solution if $N - x^2$ is a square
3. Define a helper function called `is_square` to test if an integer is square
```
def is_square(n):
"Determine if the integer n is a square."
if round(n**0.5)**2 == n:
return True
else:
return False
def reps_sum_squares(N):
'''Find all representations of N as a sum of squares x**2 + y**2 = N.
Parameters
----------
N : integer
Returns
-------
reps : list of tuples of integers
List of tuples (x,y) of positive integers such that x**2 + y**2 = N.
Examples
--------
>>> reps_sum_squares(1105)
[(4, 33), (9, 32), (12, 31), (23, 24)]
'''
reps = []
if is_square(N/2):
# If N/2 is a square, search up to x = (N/2)**0.5
max_x = round((N/2)**0.5)
else:
# If N/2 is not a square, search up to x = floor((N/2)**0.5)
max_x = int((N/2)**0.5)
for x in range(0,max_x + 1):
y_squared = N - x**2
if is_square(y_squared):
y = round(y_squared**0.5)
# Append solution (x,y) to list of solutions
reps.append((x,y))
return reps
reps_sum_squares(1105)
```
What is the smallest integer which can be expressed as the sum of squares in 5 different ways?
```
N = 1105
num_reps = 4
while num_reps < 5:
N = N + 1
reps = reps_sum_squares(N)
num_reps = len(reps)
print(N,':',reps_sum_squares(N))
```
## Examples
### Prime Numbers
A positive integer is [prime](https://en.wikipedia.org/wiki/Prime_number) if it is divisible only by 1 and itself. Write a function called `is_prime` which takes an input parameter `n` and returns `True` or `False` depending on whether `n` is prime or not.
Let's outline our approach before we write any code:
1. An integer $d$ divides $n$ if there is no remainder of $n$ divided by $d$.
2. Use the modulus operator `%` to compute the remainder.
3. If $d$ divides $n$ then $n = d q$ for some integer $q$ and either $d \leq \sqrt{n}$ or $q \leq \sqrt{n}$ (and not both), therefore we need only test if $d$ divides $n$ for integers $d \leq \sqrt{n}$
```
def is_prime(n):
"Determine whether or not n is a prime number."
if n <= 1:
return False
# Test if d divides n for d <= n**0.5
for d in range(2,round(n**0.5) + 1):
if n % d == 0:
# n is divisible by d and so n is not prime
return False
# If we exit the for loop, then n is not divisible by any d
# and therefore n is prime
return True
```
Let's test our function on the first 30 numbers:
```
for n in range(0,31):
if is_prime(n):
print(n,'is prime!')
```
Our function works! Let's find all the primes between 20,000 and 20,100.
```
for n in range(20000,20100):
if is_prime(n):
print(n,'is prime!')
```
### Divisors
Let's write a function called `divisors` which takes a positive integer $N$ and returns the list of positive integers which divide $N$.
```
def divisors(N):
"Return the list of divisors of N."
# Initialize the list of divisors (which always includes 1)
divisor_list = [1]
# Check division by d for d <= N/2
for d in range(2,N // 2 + 1):
if N % d == 0:
divisor_list.append(d)
# N divides itself and so we append N to the list of divisors
divisor_list.append(N)
return divisor_list
```
Let's test our function:
```
divisors(10)
divisors(100)
divisors(59)
```
### Collatz Conjecture
Let $a$ be a positive integer and consider the recursive sequence where $x_0 = a$ and
$$
x_{n+1} = \left\\{ \begin{array}{cl} x_n/2 & \text{if } x_n \text{ is even} \\\\ 3x_n+1 & \text{if } x_n \text{ is odd} \end{array} \\right.
$$
The [Collatz conjecture](https://en.wikipedia.org/wiki/Collatz_conjecture) states that this sequence will *always* reach 1. For example, if $a = 10$ then $x_0 = 10$, $x_1 = 5$, $x_2 = 16$, $x_3 = 8$, $x_4 = 4$, $x_5 = 2$ and $x_6 = 1$.
Write a function called `collatz` which takes one input parameter `a` and returns the sequence of integers defined above and ending with the first occurrence $x_n=1$.
```
def collatz(a):
"Compute the Collatz sequence starting at a and ending at 1."
# Initialize list with first value a
sequence = [a]
# Compute values until we reach 1
while sequence[-1] > 1:
# Check if the last element in the list is even
if sequence[-1] % 2 == 0:
# Compute and append the new value
sequence.append(sequence[-1] // 2)
else:
# Compute and append the new value
sequence.append(3*sequence[-1] + 1)
return sequence
```
Let's test our function:
```
print(collatz(10))
collatz(22)
```
The Collatz conjecture is quite amazing. No matter where we start, the sequence always terminates at 1!
```
a = 123456789
seq = collatz(a)
print("Collatz sequence for a =",a)
print("begins with",seq[:5])
print("ends with",seq[-5:])
print("and has",len(seq),"terms.")
```
Which $a < 1000$ produces the longest sequence?
```
max_length = 1
a_max = 1
for a in range(1,1001):
seq_length = len(collatz(a))
if seq_length > max_length:
max_length = seq_length
a_max = a
print('Longest sequence begins with a =',a_max,'and has length',max_length)
```
## Exercises
1. [Fermat's theorem on the sum of two squares](https://en.wikipedia.org/wiki/Fermat%27s_theorem_on_sums_of_two_squares) states that every prime number $p$ of the form $4k+1$ can be expressed as the sum of two squares. For example, $5 = 2^2 + 1^2$ and $13 = 3^2 + 2^2$. Find the smallest prime greater than $2019$ of the form $4k+1$ and write it as a sum of squares. (Hint: Use the functions `is_prime` and `reps_sum_squares` from this section.)
2. What is the smallest prime number which can be represented as a sum of squares in 2 different ways?
3. What is the smallest integer which can be represented as a sum of squares in 3 different ways?
4. Write a function called `primes_between` which takes two integer inputs $a$ and $b$ and returns the list of primes in the closed interval $[a,b]$.
5. Write a function called `primes_d_mod_N` which takes four integer inputs $a$, $b$, $d$ and $N$ and returns the list of primes in the closed interval $[a,b]$ which are congruent to $d$ mod $N$ (this means that the prime has remainder $d$ after division by $N$). This kind of list is called [primes in an arithmetic progression](https://en.wikipedia.org/wiki/Dirichlet%27s_theorem_on_arithmetic_progressions).
6. Write a function called `reciprocal_recursion` which takes three positive integers $x_0$, $x_1$ and $N$ and returns the sequence $[x_0,x_1,x_2,\dots,x_N]$ where
$$
x_n = \frac{1}{x_{n-1}} + \frac{1}{x_{n-2}}
$$
7. Write a function called `root_sequence` which takes input parameters $a$ and $N$, both positive integers, and returns the $N$th term $x_N$ in the sequence:
$$
\begin{align}
x_0 &= a \\\
x_n &= 1 + \sqrt{x_{n-1}}
\end{align}
$$
Does the sequence converge to different values for different starting values $a$?
8. Write a function called `fib_less_than` which takes one input $N$ and returns the list of Fibonacci numbers less than $N$.
9. Write a function called `fibonacci_primes` which takes an input parameter $N$ and returns the list of Fibonacci numbers less than $N$ which are also prime numbers.
10. Let $w(N)$ be the number of ways $N$ can be expressed as a sum of two squares $x^2 + y^2 = N$ with $1 \leq x \leq y$. Then
$$
\lim_{N \to \infty} \frac{1}{N} \sum_{n=1}^{N} w(n) = \frac{\pi}{8}
$$
Compute the left side of the formula for $N=100$ and compare the result to $\pi / 8$.
11. A list of positive integers $[a,b,c]$ (with $1 \leq a < b$) are a [Pythagorean triple](https://en.wikipedia.org/wiki/Pythagorean_triple) if $a^2 + b^2 = c^2$. Write a function called `py_triples` which takes an input parameter $N$ and returns the list of Pythagorean triples `[a,b,c]` with $c \leq N$.
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
%matplotlib inline
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from scipy.signal import convolve2d
from render_galaxy import *
from matplotlib.patches import Ellipse
import cProfile
from shapely.geometry import LineString, Point
from pprint import pprint
psf_size = 11
psf_sigma = 2
psf_x = np.linspace(-10, 10, psf_size)
psf_y = np.linspace(-10, 10, psf_size)
psf_x, psf_y = np.meshgrid(psf_x, psf_y)
psf = (1/(2*np.pi*psf_sigma**2) * np.exp(-(psf_x**2/(2*psf_sigma**2)
+ psf_y**2/(2*psf_sigma**2))))
psf /= np.sum(psf)
plt.imshow(psf, cmap='bone')
plt.axis('off')
plt.colorbar()
None
disk_comp = {
'mu': np.zeros(2) + 256,
'roll': np.pi / 3,
'rEff': 250,
'axRatio': 0.5,
'c': 2,
'i0': 1,
'n': 1
}
bulge_comp = {
'mu': np.zeros(2) + 256,
'roll': 0,
'rEff': 40,
'axRatio': 1,
'c': 2,
'i0': 1,
'n': 1
}
image_size = 512
oversample_n = 5
import time
t0 = time.time()
disk_arr = sersic_comp(disk_comp)
bulge_arr = sersic_comp(bulge_comp)
post_processed_image = asinh_stretch(
convolve2d(disk_arr + bulge_arr, psf)
)
print(time.time() - t0)
plt.imshow(post_processed_image, cmap='gray_r', origin='lower')
plt.colorbar()
disk_isophote = Ellipse(
disk_comp['mu'],
disk_comp['rEff'],
disk_comp['rEff'] * disk_comp['axRatio'],
-np.rad2deg(disk_comp['roll']),
facecolor='none',
edgecolor='r'
)
plt.imshow(post_processed_image, cmap='gray_r', origin='lower')
plt.gca().add_artist(disk_isophote)
print(disk_isophote)
arm_points = np.stack((np.arange(100, 250)+30, np.arange(100, 250)+30)).T
spiral_params = {
'i0': 1, 'spread': 10, 'falloff': 1,
}
spiral_arr = spiral_arm(arm_points, spiral_params, disk_comp)
post_processed_image = asinh_stretch(
convolve2d(disk_arr + bulge_arr + spiral_arr, psf)
)
plt.imshow(post_processed_image, cmap='gray_r', origin='lower')
disk_isophote = Ellipse(
disk_comp['mu'],
disk_comp['rEff'],
disk_comp['rEff'] * disk_comp['axRatio'],
-np.rad2deg(disk_comp['roll']),
facecolor='none',
edgecolor='r'
)
plt.gca().add_artist(disk_isophote)
plt.plot(*arm_points.T)
import json
with open('../../component-clustering/tmp_cls_dump.json') as f:
classifications = json.load(f)
a = classifications[6720]
pprint(a)
import parse_annotation as pa
parsed_annotation = pa.parse_annotation(a)
parsed_annotation
print('Rendering...')
print('\tDisk')
disk_arr = sersic_comp(parsed_annotation['disk'])
print('\tBulge')
bulge_arr = sersic_comp(parsed_annotation['bulge'])
print('\tBar')
bar_arr = sersic_comp(parsed_annotation['bar'])
print('\tSpiral arms')
spiral_arr = np.zeros(disk_arr.shape)
for arm_points, spiral_params in parsed_annotation['spiral']:
spiral_arr += spiral_arm(arm_points, spiral_params, parsed_annotation['disk'])
print('Post-processing')
post_processed_image = asinh_stretch(
convolve2d(disk_arr + bulge_arr + bar_arr + spiral_arr, psf)
)
plt.figure(figsize=(10, 10))
plt.imshow(post_processed_image, cmap='gray_r', origin='lower')
disk_isophote = Ellipse(
parsed_annotation['disk']['mu'],
parsed_annotation['disk']['rEff'],
parsed_annotation['disk']['rEff'] * parsed_annotation['disk']['axRatio'],
-np.rad2deg(parsed_annotation['disk']['roll']),
facecolor='none',
edgecolor='r'
)
bulge_isophote = Ellipse(
parsed_annotation['bulge']['mu'],
parsed_annotation['bulge']['rEff'],
parsed_annotation['bulge']['rEff'] * parsed_annotation['bulge']['axRatio'],
-np.rad2deg(parsed_annotation['bulge']['roll']),
facecolor='none',
edgecolor='C0'
)
plt.gca().add_artist(disk_isophote)
plt.gca().add_artist(bulge_isophote)
plt.colorbar()
```
| github_jupyter |
# Identify a CPU bottleneck caused by a callback process with Amazon SageMaker Debugger
In this notebook we demonstrate how to identify a training bottleneck that is caused by a TensorFlow Keras callback.
To simulate this type of bottleneck, we will program the callback associated with the tensor monitoring feature of Amazon SageMaker Debugger, to collect an excessive number of tensors, and at a high frequency.
### Install sagemaker
To use the new Debugger profiling features, ensure that you have the latest version of SageMaker SDK installed. The following cell updates the library and restarts the Jupyter kernel to apply the updates.
```
import sys
import IPython
install_needed = True # should only be True once
if install_needed:
print("installing deps and restarting kernel")
!{sys.executable} -m pip install -U sagemaker
IPython.Application.instance().kernel.do_shutdown(True)
```
## 1. Prepare training dataset
### Tensorflow Datasets package
First of all, set the notebook kernel to Tensorflow 2.x.
We will use CIFAR-10 dataset for this experiment. To download CIFAR-10 datasets and convert it into TFRecord format, install `tensorflow-datasets` package, run `demo/generate_cifar10_tfrecords`, and upload tfrecord files to your S3 bucket.
```
!python demo/generate_cifar10_tfrecords.py --data-dir=./data
import sagemaker
s3_bucket = sagemaker.Session().default_bucket()
dataset_prefix='data/cifar10-tfrecords'
desired_s3_uri = f's3://{s3_bucket}/{dataset_prefix}'
dataset_location = sagemaker.s3.S3Uploader.upload(local_path='data', desired_s3_uri=desired_s3_uri)
print(f'Dataset uploaded to {dataset_location}')
```
# 2. Create a Training Job with Profiling Enabled<a class="anchor" id="option-1"></a>
We will use the standard [SageMaker Estimator API for Tensorflow](https://sagemaker.readthedocs.io/en/stable/frameworks/tensorflow/sagemaker.tensorflow.html#tensorflow-estimator) to create a training job. To enable profiling, we create a `ProfilerConfig` object and pass it to the `profiler_config` parameter of the `TensorFlow` estimator. For this demo, we set the the profiler to probe the system once every 500 miliseconds.
### Set a profiler configuration
```
from sagemaker.debugger import ProfilerConfig, FrameworkProfile
profiler_config = ProfilerConfig(
system_monitor_interval_millis=500,
framework_profile_params=FrameworkProfile(local_path="/opt/ml/output/profiler/", start_step=5, num_steps=2)
)
```
### Configure Debugger hook
We configure the debugger hook to collect an excessive number of tensors, every 50 steps.
```
import os
from sagemaker.debugger import DebuggerHookConfig, CollectionConfig
debugger_hook_config = DebuggerHookConfig(
hook_parameters={
'save_interval': '50'
},
collection_configs=[
CollectionConfig(name="outputs"),
CollectionConfig(name="gradients"),
CollectionConfig(name="weights"),
CollectionConfig(name="layers")
]
)
```
### Define hyperparameters
The start-up script is set to [train_tf_bottleneck.py](./demo/train_tf_bottleneck.py). Define hyperparameters such as number of epochs, and batch size.
```
hyperparameters = {'epoch': 2,
'batch_size': 128
}
```
### Get the image URI
The image that we will is dependent on the region that you are running this notebook in.
```
import boto3
session = boto3.session.Session()
region = session.region_name
image_uri = f"763104351884.dkr.ecr.{region}.amazonaws.com/tensorflow-training:2.3.1-gpu-py37-cu110-ubuntu18.04"
```
### Define SageMaker Tensorflow Estimator
To enable profiling, you need to pass the Debugger profiling configuration (`profiler_config`), a list of Debugger rules (`rules`), and the image URI (`image_uri`) to the estimator. Debugger enables monitoring and profiling while the SageMaker estimator requests a training job.
```
import sagemaker
from sagemaker.tensorflow import TensorFlow
job_name = 'network-bottleneck'
instance_count = 1
instance_type = 'ml.p2.xlarge'
entry_script = 'train_tf_bottleneck.py'
estimator = TensorFlow(
role=sagemaker.get_execution_role(),
image_uri=image_uri,
base_job_name=job_name,
instance_type=instance_type,
instance_count=instance_count,
entry_point=entry_script,
source_dir='demo',
profiler_config=profiler_config,
debugger_hook_config=debugger_hook_config,
script_mode=True,
hyperparameters=hyperparameters,
input_mode='Pipe'
)
```
> If you see an error, `TypeError: __init__() got an unexpected keyword argument 'instance_type'`, that means SageMaker Python SDK is out-dated. Please update your SageMaker Python SDK to 2.x by executing the below command and restart this notebook.
```bash
pip install --upgrade sagemaker
```
### Start training job
The following `estimator.fit()` with `wait=False` argument initiates the training job in the background. You can proceed to run the dashboard or analysis notebooks.
```
remote_inputs = {'train' : dataset_location+'/train'}
estimator.fit(remote_inputs, wait=False)
```
# 3: Monitor the system resource utilization using SageMaker Studio
SageMaker Studio provides the visualization tool for Sagemaker Debugger where you can find the analysis report and the system and framework resource utilization history.
To access this information in SageMaker Studio, click on the last icon on the left to open `SageMaker Components and registries` and choose `Experiments and trials`. You will see the list of training jobs. Right click on the job you want to investigate shows a pop-up menu, then click on `Open Debugger for insights` which opens a new tab for SageMaker Debugger.
There are two tabs, `Overview` and `Nodes`. `Overview` gives profiling summaries for quick review, and `Nodes` gives a detailed utilization information on all nodes.
# 4: SageMaker Debugger profiling analysis utilities
We can use the profiling analysis utilities to gain deeper insights into what the source of the issue is.
For this step, we will rely on the bokeh and smdebug packages
```
! pip install bokeh==2.1.1
! pip install smdebug
```
Use smdebug to extract gpu and framework metrics
```
from smdebug.profiler.analysis.notebook_utils.training_job import TrainingJob
from smdebug.profiler.analysis.utils.profiler_data_to_pandas import PandasFrame
training_job_name = estimator.latest_training_job.name
region = 'us-east-1'
tj = TrainingJob(training_job_name, region)
pf = PandasFrame(tj.profiler_s3_output_path)
# extract gpu metrics
system_metrics_df = pf.get_all_system_metrics()
gpus = system_metrics_df[system_metrics_df['dimension'] == 'GPUUtilization']
timestamps = gpus['timestamp_us'].to_numpy()
values = gpus['value'].to_numpy()
# exctract framework metrics
framework_metrics_df = pf.get_all_framework_metrics(selected_framework_metrics=['Step:ModeKeys.TRAIN','Step:ModeKeys.GLOBAL'])
train_steps = framework_metrics_df[framework_metrics_df['framework_metric'].isin(['Step:ModeKeys.TRAIN', 'Step:ModeKeys.GLOBAL'])]
start_step = train_steps['start_time_us'].to_numpy()
end_step = train_steps['end_time_us'].to_numpy()
step_num = train_steps['step'].to_numpy()
```
Use bokeh to plot the gpu metrics and the training progression on the same graph. This enables us to correlate between the two. We can see that the drops in gpu utilization coincide with every 50th step, which are marked in yellow. These are precisely the steps in which we have chosen to capture all of the graph tensors.

```
import numpy as np
from bokeh.models import ColumnDataSource, CustomJS, Div, HoverTool, HBar
from bokeh.models.glyphs import Circle, Line
from bokeh.plotting import figure, show
plot = figure(
plot_height=400,
plot_width=1400,
x_range=(timestamps[0], timestamps[-1]),
y_range=(-1, 110),
tools="crosshair,xbox_select,pan,reset,save,xwheel_zoom",
)
x_range = plot.x_range
plot.xgrid.visible = False
plot.ygrid.visible = False
colors = np.where(step_num % 50 == 0, "yellow", "purple")
# pad framework metrics to match length of system metrics
pad = values.size - step_num.size
source = ColumnDataSource(data=dict(x=timestamps, y=values,
left=np.pad(start_step,(0,pad)),
right=np.pad(end_step,(0,pad)),
color=np.pad(colors,(0,pad))))
callback = CustomJS(
args=dict(s1=source, div=Div(width=250, height=100, height_policy="fixed")),
code="""
console.log('Running CustomJS callback now.');
var inds = s1.selected.indices;
console.log(inds);
var line = "<span style=float:left;clear:left;font_size=13px><b> Selected index range: [" + Math.min.apply(Math,inds) + "," + Math.max.apply(Math,inds) + "]</b></span>\\n";
console.log(line)
var text = div.text.concat(line);
var lines = text.split("\\n")
if (lines.length > 35)
lines.shift();
div.text = lines.join("\\n");""",
)
plot.js_on_event("selectiongeometry", callback)
line = Line(x="x", y="y", line_color="white")
circle = Circle(x="x", y="y", fill_alpha=0, line_width=0)
hbar = HBar(y=105, height=5, right="right", left="left", fill_color="color", line_cap='round', line_width=0)
p = plot.add_glyph(source, line)
p = plot.add_glyph(source, circle)
p = plot.add_glyph(source, hbar)
# create tooltip for hover tool
hover = HoverTool(
renderers=[p], tooltips=[("index", "$index"), ("(x,y)", "($x, $y)")]
)
plot.xaxis.axis_label = "Time in ms"
plot.yaxis.axis_label = "GPU Utilization"
plot.add_tools(hover)
show(plot, notebook_handle=True)
```
| github_jupyter |
## Problem
- You are maintaining a sorted list of reviews for a movie and would like to know if a particular score has been given to that movie and it the case what is the rank of that review (eg: 0 if the worse review)
- However you want to search for that information in the most efficient way possible
```
sorted_reviews = [1.0, 1.5, 2.1, 2.5, 2.5, 2.8, 3.0, 3.4, 3.9, 4.1, 4.2, 4.7, 5.0]
print(f'sorted_reviews = {sorted_reviews}')
reviews_to_search = [1.5, 3.1, 6.0, 5.0]
print(f'reviews_to_search = {reviews_to_search}')
```
- Time-Complexity Comparaison
<p align="center"><img width="35%" src="images/time-complexity.png"/></p>
## Answer
- use the bisect.bisect_left function which implements binary-search and enables us to search for a sorted list
```
import bisect #<0>
```
#### Let's search if if a given score has been given to the movie
```
# BAD WAY: linear-search ==> O(n) time-complexity
for new_review in reviews_to_search:
# we could use in operator instead but just to make the point explicit let's use a for loop
for position, review in enumerate(sorted_reviews): #<1>
if review == new_review:
print(f'[HIT]review={new_review} found in position={position}')
break
else:
print(f'[MISS]review={new_review} not found in sorted_reviews')
# GOOD WAY: binary-search ==> 0(log n) time-complexity
for new_review in reviews_to_search:
position = bisect.bisect_left(sorted_reviews, new_review) #<2>
if position < len(sorted_reviews) and sorted_reviews[position] == new_review: #<3>
print(f'[HIT]review={new_review} found in position={position}')
else:
print(f'[MISS]review={new_review} not found in sorted_reviews')
```
## Discussion
- <0> importing the bisect module which provides 0(log n) efficient implementation of search and insertion based on binary-search.
- <1> by doing a linear search we are not taking advantage of the fact that the list of reviews is already sorted.
- <2> bisect.bisect_left takes advantage of the fact that the list is sorted to reduce the search complexity from 0(n) to 0(log n)
- <3> bisect.bisect_left returns the position where the search value IS or SHOULD BE if not yet in the list.
## Problem
- How to insert new reviews in a list of sorted reviews efficiently
## Answer
- use the bisect.insort_left function which search for the right position then insert by shifting all the other elements to the right.
```
# VERY BAD WAY: append to the end then sort ==> O(n * log n)
sorted_reviews = [1.0, 2.1, 2.5, 2.8, 3.0, 3.9, 4.1, 4.7]
reviews_to_insert = [6.0, 3.4, 5.0, 1.5]
sorted_reviews += reviews_to_insert
sorted_reviews #<0>
sorted_reviews.sort() #<1>
sorted_reviews
# GOOD WAY: use bisect.insort_left ==> O(n) + 0(log n) ~ O(n)
sorted_reviews = [1.0, 2.1, 2.5, 2.8, 3.0, 3.9, 4.1, 4.7]
reviews_to_insert = [6.0, 3.4, 5.0, 1.5]
print(f'Initial sorted_reviews = {sorted_reviews}')
for new_review in reviews_to_insert:
bisect.insort_left(sorted_reviews, new_review) #<2>
print(f'sorted_reviews after inserting {new_review} = {sorted_reviews}')
```
## Discussion
- <0> we append all the new reviews to insert and the end of the sorted_reviews list
- <1> best sorting algorithms take O(n * log n) time to sort
- <2> takes 0(log n) to search right insert position and O(n) to shift.
| github_jupyter |
<h1>Table of Contents<span class="tocSkip"></span></h1>
<div class="toc"><ul class="toc-item"><li><span><a href="#Nearest-Neighbors" data-toc-modified-id="Nearest-Neighbors-1"><span class="toc-item-num">1 </span>Nearest Neighbors</a></span><ul class="toc-item"><li><span><a href="#TF-IDF" data-toc-modified-id="TF-IDF-1.1"><span class="toc-item-num">1.1 </span>TF-IDF</a></span></li></ul></li><li><span><a href="#Reference" data-toc-modified-id="Reference-2"><span class="toc-item-num">2 </span>Reference</a></span></li></ul></div>
```
# code for loading the format for the notebook
import os
# path : store the current path to convert back to it later
path = os.getcwd()
os.chdir(os.path.join('..', '..', 'notebook_format'))
from formats import load_style
load_style(plot_style = False)
os.chdir(path)
# 1. magic for inline plot
# 2. magic to print version
# 3. magic so that the notebook will reload external python modules
# 4. magic to enable retina (high resolution) plots
# https://gist.github.com/minrk/3301035
%matplotlib inline
%load_ext watermark
%load_ext autoreload
%autoreload 2
%config InlineBackend.figure_format = 'retina'
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from collections import Counter
from sklearn.neighbors import NearestNeighbors
from sklearn.metrics.pairwise import paired_distances
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.feature_extraction.text import TfidfTransformer, TfidfVectorizer
%watermark -a 'Ethen' -d -t -v -p numpy,pandas,matplotlib,sklearn
```
# Nearest Neighbors
When exploring a large set of text documents -- such as Wikipedia, news articles, StackOverflow, etc. -- it can be useful to get a list of related material. To find relevant documents you typically need to convert the text document into bag of words or TF-IDF format and choose the distance metric to measure the notion of similarity. In this documentation we'll explore the tradeoffs with representing documents using bag of words and TF-IDF.
We will be using the Wikipedia pages dataset. Each row of the dataset consists of a link to the Wikipedia article, the name of the person, and the text of the article (in lowercase). To follow along, please download the file from this [dropbox link](https://www.dropbox.com/s/mriz0nq35ore8cr/people_wiki.csv?dl=0).
```
# the file is placed one level above this notebook,
# since it's also used by other notebooks
# change this if you liked
filepath = os.path.join('..', 'people_wiki.csv')
wiki = pd.read_csv(filepath)
wiki.head(5)
```
We start by converting the document into bag of words format and utilize the euclidean distance to find the nearest neighbors of the Barack Obama.
```
# bag of words
vect = CountVectorizer()
word_weight = vect.fit_transform(wiki['text'])
# query Barack Obama's top 10 nearest neighbors
nn = NearestNeighbors(metric = 'euclidean')
nn.fit(word_weight)
obama_index = wiki[wiki['name'] == 'Barack Obama'].index[0]
distances, indices = nn.kneighbors(word_weight[obama_index], n_neighbors = 10)
# 1. flatten the 2d-array distance and indices to 1d
# 2. merge the distance information with the original wiki dataset,
# to obtain the name matching the id.
neighbors = pd.DataFrame({'distance': distances.flatten(), 'id': indices.flatten()})
nearest_info = (wiki.
merge(neighbors, right_on = 'id', left_index = True).
sort_values('distance')[['id', 'name', 'distance']])
nearest_info
```
Looking at the result, it seems nice that all of the 10 people are politicians. Let's dig a bit deeper and find out why Joe Biden was considered a close neighbor of Obama by looking at the most frequently used words in both pages. To do this, we'll take the sparse row matrix that we had and extract the word count dictionary for each document.
```
def unpack_word_weight(vect, word_weight):
"""
Given the CountVector and the fit_transformed
word count sparse array obtain each documents'
word count dictionary
In the Compressed Sparse Row format,
`indices` stands for indexes inside the row vectors of the matrix
and `indptr` (index pointer) tells where the row starts in the data and in the indices
attributes. nonzero values of the i-th row are data[indptr[i]:indptr[i+1]]
with column indices indices[indptr[i]:indptr[i+1]]
References
----------
http://www.scipy-lectures.org/advanced/scipy_sparse/csr_matrix.html
"""
feature_names = np.array(vect.get_feature_names())
data = word_weight.data
indptr = word_weight.indptr
indices = word_weight.indices
n_docs = word_weight.shape[0]
word_weight_list = []
for i in range(n_docs):
doc = slice(indptr[i], indptr[i + 1])
count, idx = data[doc], indices[doc]
feature = feature_names[idx]
word_weight_dict = Counter({k: v for k, v in zip(feature, count)})
word_weight_list.append(word_weight_dict)
return word_weight_list
wiki['word_weight'] = unpack_word_weight(vect, word_weight)
wiki.head(3)
def get_top_words(wiki, name, column_name, top_n = None):
row = wiki.loc[wiki['name'] == name, column_name]
# when converting Series to dictionary, the row index
# will be the key to the content
word_weight_dict = row.to_dict()[row.index[0]]
if top_n is None:
top_n = len(word_weight_dict)
word_weight_table = pd.DataFrame(word_weight_dict.most_common(top_n),
columns = ['word', 'weight'])
return word_weight_table
words_obama = get_top_words(wiki, name = 'Barack Obama', column_name = 'word_weight')
words_biden = get_top_words(wiki, name = 'Joe Biden', column_name = 'word_weight')
# merge the two DataFrame, since both tables contained the same column name weight,
# it will automatically renamed one of them by adding suffix _x and _y to prevent confusion.
# hence we'll rename the columns to tell which one is for which
words_combined = (words_obama.
merge(words_biden, on = 'word').
rename(columns = {'weight_x': 'Obama', 'weight_y': 'Biden'}))
words_combined.head(6)
```
The next question we want to ask is, among these common words that appear in both documents, how many other documents in the Wikipedia dataset also contain them?
```
def has_top_words(word_weight_vector, common_words):
"""
return True if common_words is a subset of unique_words
return False otherwise
References
----------
http://stackoverflow.com/questions/12182744/python-pandas-apply-a-function-with-arguments-to-a-series
"""
# extract the keys of word_weight_vector and convert it to a set
unique_words = set(word_weight_vector.keys())
boolean = common_words.issubset(unique_words)
return boolean
# we'll extract the 5 most frequent
common_words = set(words_combined['word'].head(5))
print('top 5 common words: ', common_words)
wiki['has_top_words'] = wiki['word_weight'].apply(has_top_words, args = (common_words,))
print('number of articles that also contain the common words: ', wiki['has_top_words'].sum())
```
Given this result, we saw that much of the perceived commonalities between the two articles were due to occurrences of extremely frequent words, such as "the", "and", and "his". All of these words appear very often in all of the documents. To retrieve articles that are more relevant, maybe we should be focusing more on rare words that don't happen in every article. And this is where TF-IDF (term frequency–inverse document frequency) comes in. Note that although we can remove stop words to prevent some of this behavior, sometimes the frequently appeared words might not be a stop word.
## TF-IDF
TF-IDF, short for term frequency–inverse document frequency, is a numeric measure that is use to score the importance of a word in a document based on how often did it appear in that document and a given collection of documents. The intuition for this measure is : If a word appears frequently in a document, then it should be important and we should give that word a high score. But if a word appears in too many other documents, it's probably not a unique identifier, therefore we should assign a lower score to that word. In short, it is a feature representation that penalizes words that are too common.
Let us consider the following toy dataset that consists of 3 documents.
```
docs = np.array([
'The sun is shining',
'The weather is sweet',
'The sun is shining and the weather is sweet'
])
```
Most commonly, TF-IDF is calculated as follows:
$$\text{tf-idf}(t, d, D) = \text{tf}(t, d) \times \text{idf}(t, d, D)$$
Where `t` denotes the terms; `d` denotes each document; `D` denotes the collection of documents.
The first part of the formula $tf(t, d)$ stands for term frequency, which is defined by the number of times a term $t$ occurs in a document $d$.
```
vect = CountVectorizer()
tf = vect.fit_transform(docs).toarray()
tf
vect.vocabulary_
```
Based on the vocabulary, the word "and" would be the first column in each document vector in `tf` and it appears once in the third document.
In order to understand the second part of the formlua $\text{idf}(t, d, D)$, inverse document frequency, let's first write down the complete math formula for IDF.
$$ idf(t, d, D) = log \frac{ \mid \text{ } D \text{ } \mid }{ 1 + \mid \{ d : t \in d \} \mid } $$
- The numerator : `D` is infering to our document space. It can also be seen as D = ${ d_{1}, d_{2}, \dots, d_{n} }$ where n is the number of documents in your collection. Thus for our example $\mid \text{ } D \text{ } \mid$, the size of our document space is 3, since we have 3 documents.
- The denominator : $\mid \{ d: t \in d \} \mid$ is the document freqency. To be explicit, it is the number of documents $d$ that contain the term $t$. Note that this implies it doesn't matter if a term appeared 1 time or 100 times in a document, it will still be counted as 1, since the term simply did appear in the document.
- The constant 1 is added to the denominator to avoid a zero-division error if a term is not contained in any document in the test dataset.
Note that there are very different variation of the formula. For example, The tf-idfs in scikit-learn are calculated by:
$$ idf(t, d, D) = log \frac{ \mid \text{ } D \text{ } \mid }{ \mid \{ d : t \in d \} \mid } + 1 $$
Here, the `+1` count is added directly to the idf, instead of the denominator. The effect of this is that terms with zero idf, i.e. that occur in all documents of a training set, will not be entirely ignored. We can demonstrate this by calculating the idfs manually using the equation above and do the calculation ourselves and compare the results to the TfidfTransformer output using the settings `use_idf=True, smooth_idf=False, norm=None` (more on `smooth_idf` and `norm` later).
```
# compute manually the tfidf score for the first document
n_docs = len(docs)
df = np.sum(tf != 0, axis = 0)
idf = np.log(n_docs / df) + 1
tf_idf = tf[0] * idf
print(tf_idf)
print()
# use the library to do the computation
tfidf = TfidfTransformer(use_idf = True, smooth_idf = False, norm = None)
doc_tfidf = tfidf.fit_transform(tf).toarray()
print(doc_tfidf[0])
assert np.allclose(tf_idf, doc_tfidf[0])
```
Next, recall that in the tf (term frequency) section, we’re representing each term as the number of times they appeared in the document. The main issue for this representation is that it will create a bias towards long documents, as a given term has more chance to appear in longer document, making them look more important than actually they are. Thus the approach to resolve this issue is the good old L2 normalization. i.e., dividing the raw term frequency vector $v$ by its length $||v||_2$ (L2- or Euclidean norm).
$$v_{norm} = \frac{v}{ \parallel v \parallel_2} = \frac{v}{\sqrt{v{_1}^2 + v{_2}^2 + \dots + v{_n}^2}} = \frac{v}{\big(\sum_{i=1}^n v_i^2 \big)^{\frac{1}{2}}}$$
```
# manual
tf_norm = tf_idf / np.sqrt(np.sum(tf_idf ** 2))
print(tf_norm)
print()
# library
tfidf = TfidfTransformer(use_idf = True, smooth_idf = False, norm = 'l2')
doc_tfidf = tfidf.fit_transform(tf).toarray()
print(doc_tfidf[0])
assert np.allclose(tf_norm, doc_tfidf[0])
```
Another parameter in the `TfidfTransformer` is the `smooth_idf`, which is described as
> smooth_idf : boolean, default=True
Smooth idf weights by adding one to document frequencies, as if an extra document was seen containing every term in the collection exactly once. Prevents zero divisions.
So, our idf would then be defined as follows:
$$idf(t, d, D) = log \frac{ 1 + \mid D \text{ } \mid }{ 1 + \mid \{ d : t \in d \} \mid } + 1 $$
Again let's confirm this by computing it manually and using the library.
```
# manual
n_docs = len(docs)
df = np.sum(tf != 0, axis = 0)
idf = np.log((1 + n_docs) / (1 + df)) + 1
tf_idf = tf[0] * idf
tf_norm = tf_idf / np.sqrt(np.sum(tf_idf ** 2))
print(tf_norm)
print()
# library
tfidf = TfidfTransformer(use_idf = True, smooth_idf = True, norm = 'l2')
doc_tfidf = tfidf.fit_transform(tf).toarray()
print(doc_tfidf[0])
assert np.allclose(tf_norm, doc_tfidf[0])
```
To sum it up, the default settings in the `TfidfTransformer` is:
- `use_idf=True`
- `smooth_idf=True`
- `norm='l2'`
And we can use the `TfidfVectorizer` to compute the TF-IDF score from raw text in one step without having to do use `CountVectorizer` to convert it to bag of words representation and then transform it to TF-IDF using `TfidfTransformer`. For those interested, this [link](https://github.com/ethen8181/machine-learning/blob/master/clustering/tfidf/feature_extraction.py) contains the full TF-IDF implemented from scratch.
Now that we've understood the inner details on TF-IDF let's return back to our initial task and use this weighting scheme instead of bag of words. We start by converting the document into TF-IDF format and use this along with cosine distance to find the nearest neighbors of the Barack Obama (if we normalized our articles in the TF-IDF transformation, then the euclidean distance and the cosine distance is proportional to each other, hence they're doing the same thing). Then we extract the commonly used words (now weighted by the TF-IDF score instead of word count) in both the Joe Biden and Obama article. Finally we compute how many other documents in the Wikipedia dataset also contain these words. In short, we're doing the exact same thing as the beginning except we now use TF-IDF to represent the documents and use cosine distance to measure similarity between documents.
```
# tf-idf instead of bag of words
tfidf_vect = TfidfVectorizer()
tfidf_weight = tfidf_vect.fit_transform(wiki['text'])
# this time, query Barack Obama's top 100 nearest neighbors using
# cosine distance, we have to specify the algorithm to `brute` since
# the default 'auto' does not work with cosine distance
# http://stackoverflow.com/questions/32745541/dbscan-error-with-cosine-metric-in-python
nn_cosine = NearestNeighbors(metric = 'cosine', algorithm = 'brute')
nn_cosine.fit(tfidf_weight)
obama_index = wiki[wiki['name'] == 'Barack Obama'].index[0]
cosine, indices = nn_cosine.kneighbors(tfidf_weight[obama_index], n_neighbors = 100)
# 1. flatten the 2d-array distance and indices to 1d
# 2. merge the distance information with the original wiki dataset,
# to obtain the name matching the id.
neighbors_cosine = pd.DataFrame({'cosine': cosine.flatten(), 'id': indices.flatten()})
nearest_info = (wiki.
merge(neighbors_cosine, right_on = 'id', left_index = True).
sort_values('cosine')[['id', 'name', 'cosine']])
nearest_info.head()
wiki['tfidf_weight'] = unpack_word_weight(tfidf_vect, tfidf_weight)
wiki.head(3)
words_obama = get_top_words(wiki, name = 'Barack Obama', column_name = 'tfidf_weight')
words_biden = get_top_words(wiki, name = 'Joe Biden', column_name = 'tfidf_weight')
words_combined = (words_obama.
merge(words_biden, on = 'word').
rename(columns = {'weight_x': 'Obama', 'weight_y': 'Biden'}))
words_combined.head(6)
# we'll extract the 5 most frequent
common_words = set(words_combined['word'].head(5))
print('top 5 common words: ', common_words)
wiki['has_top_words'] = wiki['tfidf_weight'].apply(has_top_words, args = (common_words,))
print('number of articles that also contain the common words: ', wiki['has_top_words'].sum())
```
Notice the huge difference in this calculation using TF-IDF scores instead of raw word counts. We've eliminated noise arising from extremely common words. Happily ever after? Not so fast. To illustrate another possible issue, let's first compute the length of each Wikipedia document, and examine the document lengths for the 100 nearest neighbors to Obama's page.
```
def compute_length(row):
return len(row.split(' '))
wiki['length'] = wiki['text'].apply(compute_length)
wiki.head(3)
nearest_cosine = (wiki.
merge(neighbors_cosine, right_on = 'id', left_index = True).
sort_values('cosine')[['id', 'name', 'cosine', 'length']])
nearest_cosine.head()
plt.figure(figsize = (10.5, 4.5))
plt.hist(wiki['length'], 50, histtype = 'stepfilled',
normed = True, label = 'Entire Wikipedia', alpha = 0.5)
plt.hist(nearest_cosine['length'], 50, histtype = 'stepfilled',
normed = True, label = '100 NNs of Obama (cosine)', alpha = 0.8)
plt.axvline(nearest_cosine.loc[nearest_cosine['name'] == 'Barack Obama', 'length'].values,
color = 'g', linestyle = '--', linewidth = 4,
label = 'Length of Barack Obama')
plt.axis([0, 1500, 0, 0.04])
plt.legend(loc = 'best', prop = {'size': 15})
plt.title('Distribution of document length')
plt.xlabel('# of words')
plt.ylabel('Percentage')
plt.tight_layout()
```
The visualization is basically telling us that 100 nearest neighbors using TF-IDF weighting and cosine distance provide a sampling across articles with different length. The thing is that whether we're choosing euclidean or cosine distance to measure our articles' similarity, both of them still ignore the document's length, which may be great in certain situations but not in others. For instance, consider the following (admittedly contrived) tweet.
```
+--------------------------------------------------------+
| +--------+ |
| One that shall not be named | Follow | |
| @username +--------+ |
| |
| Democratic governments control law in response to |
| popular act. |
| |
| 8:05 AM - 16 May 2016 |
| |
| Reply Retweet (1,332) Like (300) |
| |
+--------------------------------------------------------+
```
How similar is this tweet to Barack Obama's Wikipedia article? Let's transform the tweet into TF-IDF features; compute the cosine distance between the Barack Obama article and this tweet and compare this distance to the distance between the Barack Obama article and all of its Wikipedia 10 nearest neighbors.
```
tweet = tfidf_vect.transform(['democratic governments control law in response to popular act'])
tweet_dist = paired_distances(tfidf_weight[obama_index], tweet, metric = 'cosine')
# compare to the 100 articles that were nearest to Obama's,
# the distance of this tweet is shorter than how many articles
np.sum(tweet_dist < nearest_cosine['cosine'].values)
```
With cosine distances, the tweet is "nearer" to Barack Obama than most of the articles. If someone is reading the Barack Obama Wikipedia page, would we want to recommend they read this tweet? Ignoring article lengths completely resulted in nonsensical results. In practice, it is common to enforce maximum or minimum document lengths. After all, when someone is reading a long article from *The Atlantic*, we wouldn't recommend him/her a tweet.
# Reference
- [Coursera: Washington Clustering & Retrieval](https://www.coursera.org/learn/ml-clustering-and-retrieval)
- [Notebooks: Tf-idf Walkthrough for scikit-learn](http://nbviewer.jupyter.org/github/rasbt/pattern_classification/blob/master/machine_learning/scikit-learn/tfidf_scikit-learn.ipynb)
- [Blog: Machine Learning :: Text feature extraction (tf-idf)](http://blog.christianperone.com/2011/09/machine-learning-text-feature-extraction-tf-idf-part-i/)
| github_jupyter |
# Multigroup (Delayed) Cross Section Generation Part II: Advanced Features
This IPython Notebook illustrates the use of the **`openmc.mgxs.Library`** class. The `Library` class is designed to automate the calculation of multi-group cross sections for use cases with one or more domains, cross section types, and/or nuclides. In particular, this Notebook illustrates the following features:
* Calculation of multi-energy-group and multi-delayed-group cross sections for a **fuel assembly**
* Automated creation, manipulation and storage of `MGXS` with **`openmc.mgxs.Library`**
* Steady-state pin-by-pin **delayed neutron fractions (beta)** for each delayed group.
* Generation of surface currents on the interfaces and surfaces of a Mesh.
## Generate Input Files
```
%matplotlib inline
import math
import matplotlib.pyplot as plt
import numpy as np
import openmc
import openmc.mgxs
```
First we need to define materials that will be used in the problem: fuel, water, and cladding.
```
# 1.6 enriched fuel
fuel = openmc.Material(name='1.6% Fuel')
fuel.set_density('g/cm3', 10.31341)
fuel.add_nuclide('U235', 3.7503e-4)
fuel.add_nuclide('U238', 2.2625e-2)
fuel.add_nuclide('O16', 4.6007e-2)
# borated water
water = openmc.Material(name='Borated Water')
water.set_density('g/cm3', 0.740582)
water.add_nuclide('H1', 4.9457e-2)
water.add_nuclide('O16', 2.4732e-2)
water.add_nuclide('B10', 8.0042e-6)
# zircaloy
zircaloy = openmc.Material(name='Zircaloy')
zircaloy.set_density('g/cm3', 6.55)
zircaloy.add_nuclide('Zr90', 7.2758e-3)
```
With our three materials, we can now create a `Materials` object that can be exported to an actual XML file.
```
# Create a materials collection and export to XML
materials = openmc.Materials((fuel, water, zircaloy))
materials.export_to_xml()
```
Now let's move on to the geometry. This problem will be a square array of fuel pins and control rod guide tubes for which we can use OpenMC's lattice/universe feature. The basic universe will have three regions for the fuel, the clad, and the surrounding coolant. The first step is to create the bounding surfaces for fuel and clad, as well as the outer bounding surfaces of the problem.
```
# Create cylinders for the fuel and clad
fuel_outer_radius = openmc.ZCylinder(r=0.39218)
clad_outer_radius = openmc.ZCylinder(r=0.45720)
# Create boundary planes to surround the geometry
min_x = openmc.XPlane(x0=-10.71, boundary_type='reflective')
max_x = openmc.XPlane(x0=+10.71, boundary_type='reflective')
min_y = openmc.YPlane(y0=-10.71, boundary_type='reflective')
max_y = openmc.YPlane(y0=+10.71, boundary_type='reflective')
min_z = openmc.ZPlane(z0=-10., boundary_type='reflective')
max_z = openmc.ZPlane(z0=+10., boundary_type='reflective')
```
With the surfaces defined, we can now construct a fuel pin cell from cells that are defined by intersections of half-spaces created by the surfaces.
```
# Create a Universe to encapsulate a fuel pin
fuel_pin_universe = openmc.Universe(name='1.6% Fuel Pin')
# Create fuel Cell
fuel_cell = openmc.Cell(name='1.6% Fuel')
fuel_cell.fill = fuel
fuel_cell.region = -fuel_outer_radius
fuel_pin_universe.add_cell(fuel_cell)
# Create a clad Cell
clad_cell = openmc.Cell(name='1.6% Clad')
clad_cell.fill = zircaloy
clad_cell.region = +fuel_outer_radius & -clad_outer_radius
fuel_pin_universe.add_cell(clad_cell)
# Create a moderator Cell
moderator_cell = openmc.Cell(name='1.6% Moderator')
moderator_cell.fill = water
moderator_cell.region = +clad_outer_radius
fuel_pin_universe.add_cell(moderator_cell)
```
Likewise, we can construct a control rod guide tube with the same surfaces.
```
# Create a Universe to encapsulate a control rod guide tube
guide_tube_universe = openmc.Universe(name='Guide Tube')
# Create guide tube Cell
guide_tube_cell = openmc.Cell(name='Guide Tube Water')
guide_tube_cell.fill = water
guide_tube_cell.region = -fuel_outer_radius
guide_tube_universe.add_cell(guide_tube_cell)
# Create a clad Cell
clad_cell = openmc.Cell(name='Guide Clad')
clad_cell.fill = zircaloy
clad_cell.region = +fuel_outer_radius & -clad_outer_radius
guide_tube_universe.add_cell(clad_cell)
# Create a moderator Cell
moderator_cell = openmc.Cell(name='Guide Tube Moderator')
moderator_cell.fill = water
moderator_cell.region = +clad_outer_radius
guide_tube_universe.add_cell(moderator_cell)
```
Using the pin cell universe, we can construct a 17x17 rectangular lattice with a 1.26 cm pitch.
```
# Create fuel assembly Lattice
assembly = openmc.RectLattice(name='1.6% Fuel Assembly')
assembly.pitch = (1.26, 1.26)
assembly.lower_left = [-1.26 * 17. / 2.0] * 2
```
Next, we create a NumPy array of fuel pin and guide tube universes for the lattice.
```
# Create array indices for guide tube locations in lattice
template_x = np.array([5, 8, 11, 3, 13, 2, 5, 8, 11, 14, 2, 5, 8,
11, 14, 2, 5, 8, 11, 14, 3, 13, 5, 8, 11])
template_y = np.array([2, 2, 2, 3, 3, 5, 5, 5, 5, 5, 8, 8, 8, 8,
8, 11, 11, 11, 11, 11, 13, 13, 14, 14, 14])
# Create universes array with the fuel pin and guide tube universes
universes = np.tile(fuel_pin_universe, (17,17))
universes[template_x, template_y] = guide_tube_universe
# Store the array of universes in the lattice
assembly.universes = universes
```
OpenMC requires that there is a "root" universe. Let us create a root cell that is filled by the pin cell universe and then assign it to the root universe.
```
# Create root Cell
root_cell = openmc.Cell(name='root cell', fill=assembly)
# Add boundary planes
root_cell.region = +min_x & -max_x & +min_y & -max_y & +min_z & -max_z
# Create root Universe
root_universe = openmc.Universe(universe_id=0, name='root universe')
root_universe.add_cell(root_cell)
```
We now must create a geometry that is assigned a root universe and export it to XML.
```
# Create Geometry and export to XML
geometry = openmc.Geometry(root_universe)
geometry.export_to_xml()
```
With the geometry and materials finished, we now just need to define simulation parameters. In this case, we will use 10 inactive batches and 40 active batches each with 2500 particles.
```
# OpenMC simulation parameters
batches = 50
inactive = 10
particles = 2500
# Instantiate a Settings object
settings = openmc.Settings()
settings.batches = batches
settings.inactive = inactive
settings.particles = particles
settings.output = {'tallies': False}
# Create an initial uniform spatial source distribution over fissionable zones
bounds = [-10.71, -10.71, -10, 10.71, 10.71, 10.]
uniform_dist = openmc.stats.Box(bounds[:3], bounds[3:], only_fissionable=True)
settings.source = openmc.Source(space=uniform_dist)
# Export to "settings.xml"
settings.export_to_xml()
```
Let us also create a plot to verify that our fuel assembly geometry was created successfully.
```
# Plot our geometry
plot = openmc.Plot.from_geometry(geometry)
plot.pixels = (250, 250)
plot.color_by = 'material'
openmc.plot_inline(plot)
```
As we can see from the plot, we have a nice array of fuel and guide tube pin cells with fuel, cladding, and water!
## Create an MGXS Library
Now we are ready to generate multi-group cross sections! First, let's define a 20-energy-group and 1-energy-group.
```
# Instantiate a 20-group EnergyGroups object
energy_groups = openmc.mgxs.EnergyGroups()
energy_groups.group_edges = np.logspace(-3, 7.3, 21)
# Instantiate a 1-group EnergyGroups object
one_group = openmc.mgxs.EnergyGroups()
one_group.group_edges = np.array([energy_groups.group_edges[0], energy_groups.group_edges[-1]])
```
Next, we will instantiate an `openmc.mgxs.Library` for the energy and delayed groups with our the fuel assembly geometry.
```
# Instantiate a tally mesh
mesh = openmc.RegularMesh(mesh_id=1)
mesh.dimension = [17, 17, 1]
mesh.lower_left = [-10.71, -10.71, -10000.]
mesh.width = [1.26, 1.26, 20000.]
# Initialize an 20-energy-group and 6-delayed-group MGXS Library
mgxs_lib = openmc.mgxs.Library(geometry)
mgxs_lib.energy_groups = energy_groups
mgxs_lib.num_delayed_groups = 6
# Specify multi-group cross section types to compute
mgxs_lib.mgxs_types = ['total', 'transport', 'nu-scatter matrix', 'kappa-fission', 'inverse-velocity', 'chi-prompt',
'prompt-nu-fission', 'chi-delayed', 'delayed-nu-fission', 'beta']
# Specify a "mesh" domain type for the cross section tally filters
mgxs_lib.domain_type = 'mesh'
# Specify the mesh domain over which to compute multi-group cross sections
mgxs_lib.domains = [mesh]
# Construct all tallies needed for the multi-group cross section library
mgxs_lib.build_library()
# Create a "tallies.xml" file for the MGXS Library
tallies_file = openmc.Tallies()
mgxs_lib.add_to_tallies_file(tallies_file, merge=True)
# Instantiate a current tally
mesh_filter = openmc.MeshSurfaceFilter(mesh)
current_tally = openmc.Tally(name='current tally')
current_tally.scores = ['current']
current_tally.filters = [mesh_filter]
# Add current tally to the tallies file
tallies_file.append(current_tally)
# Export to "tallies.xml"
tallies_file.export_to_xml()
```
Now, we can run OpenMC to generate the cross sections.
```
# Run OpenMC
openmc.run()
```
## Tally Data Processing
Our simulation ran successfully and created statepoint and summary output files. We begin our analysis by instantiating a `StatePoint` object.
```
# Load the last statepoint file
sp = openmc.StatePoint('statepoint.50.h5')
```
The statepoint is now ready to be analyzed by the `Library`. We simply have to load the tallies from the statepoint into the `Library` and our `MGXS` objects will compute the cross sections for us under-the-hood.
```
# Initialize MGXS Library with OpenMC statepoint data
mgxs_lib.load_from_statepoint(sp)
# Extrack the current tally separately
current_tally = sp.get_tally(name='current tally')
```
## Using Tally Arithmetic to Compute the Delayed Neutron Precursor Concentrations
Finally, we illustrate how one can leverage OpenMC's [tally arithmetic](tally-arithmetic.ipynb) data processing feature with `MGXS` objects. The `openmc.mgxs` module uses tally arithmetic to compute multi-group cross sections with automated uncertainty propagation. Each `MGXS` object includes an `xs_tally` attribute which is a "derived" `Tally` based on the tallies needed to compute the cross section type of interest. These derived tallies can be used in subsequent tally arithmetic operations. For example, we can use tally artithmetic to compute the delayed neutron precursor concentrations using the `Beta` and `DelayedNuFissionXS` objects. The delayed neutron precursor concentrations are modeled using the following equations:
$$\frac{\partial}{\partial t} C_{k,d} (t) = \int_{0}^{\infty}\mathrm{d}E'\int_{\mathbf{r} \in V_{k}}\mathrm{d}\mathbf{r} \beta_{k,d} (t) \nu_d \sigma_{f,x}(\mathbf{r},E',t)\Phi(\mathbf{r},E',t) - \lambda_{d} C_{k,d} (t) $$
$$C_{k,d} (t=0) = \frac{1}{\lambda_{d}} \int_{0}^{\infty}\mathrm{d}E'\int_{\mathbf{r} \in V_{k}}\mathrm{d}\mathbf{r} \beta_{k,d} (t=0) \nu_d \sigma_{f,x}(\mathbf{r},E',t=0)\Phi(\mathbf{r},E',t=0) $$
```
# Set the time constants for the delayed precursors (in seconds^-1)
precursor_halflife = np.array([55.6, 24.5, 16.3, 2.37, 0.424, 0.195])
precursor_lambda = math.log(2.0) / precursor_halflife
beta = mgxs_lib.get_mgxs(mesh, 'beta')
# Create a tally object with only the delayed group filter for the time constants
beta_filters = [f for f in beta.xs_tally.filters if type(f) is not openmc.DelayedGroupFilter]
lambda_tally = beta.xs_tally.summation(nuclides=beta.xs_tally.nuclides)
for f in beta_filters:
lambda_tally = lambda_tally.summation(filter_type=type(f), remove_filter=True) * 0. + 1.
# Set the mean of the lambda tally and reshape to account for nuclides and scores
lambda_tally._mean = precursor_lambda
lambda_tally._mean.shape = lambda_tally.std_dev.shape
# Set a total nuclide and lambda score
lambda_tally.nuclides = [openmc.Nuclide(name='total')]
lambda_tally.scores = ['lambda']
delayed_nu_fission = mgxs_lib.get_mgxs(mesh, 'delayed-nu-fission')
# Use tally arithmetic to compute the precursor concentrations
precursor_conc = beta.xs_tally.summation(filter_type=openmc.EnergyFilter, remove_filter=True) * \
delayed_nu_fission.xs_tally.summation(filter_type=openmc.EnergyFilter, remove_filter=True) / lambda_tally
# The difference is a derived tally which can generate Pandas DataFrames for inspection
precursor_conc.get_pandas_dataframe().head(10)
```
Another useful feature of the Python API is the ability to extract the surface currents for the interfaces and surfaces of a mesh. We can inspect the currents for the mesh by getting the pandas dataframe.
```
current_tally.get_pandas_dataframe().head(10)
```
## Cross Section Visualizations
In addition to inspecting the data in the tallies by getting the pandas dataframe, we can also plot the tally data on the domain mesh. Below is the delayed neutron fraction tallied in each mesh cell for each delayed group.
```
# Extract the energy-condensed delayed neutron fraction tally
beta_by_group = beta.get_condensed_xs(one_group).xs_tally.summation(filter_type='energy', remove_filter=True)
beta_by_group.mean.shape = (17, 17, 6)
beta_by_group.mean[beta_by_group.mean == 0] = np.nan
# Plot the betas
plt.figure(figsize=(18,9))
fig = plt.subplot(231)
plt.imshow(beta_by_group.mean[:,:,0], interpolation='none', cmap='jet')
plt.colorbar()
plt.title('Beta - delayed group 1')
fig = plt.subplot(232)
plt.imshow(beta_by_group.mean[:,:,1], interpolation='none', cmap='jet')
plt.colorbar()
plt.title('Beta - delayed group 2')
fig = plt.subplot(233)
plt.imshow(beta_by_group.mean[:,:,2], interpolation='none', cmap='jet')
plt.colorbar()
plt.title('Beta - delayed group 3')
fig = plt.subplot(234)
plt.imshow(beta_by_group.mean[:,:,3], interpolation='none', cmap='jet')
plt.colorbar()
plt.title('Beta - delayed group 4')
fig = plt.subplot(235)
plt.imshow(beta_by_group.mean[:,:,4], interpolation='none', cmap='jet')
plt.colorbar()
plt.title('Beta - delayed group 5')
fig = plt.subplot(236)
plt.imshow(beta_by_group.mean[:,:,5], interpolation='none', cmap='jet')
plt.colorbar()
plt.title('Beta - delayed group 6')
```
| github_jupyter |
```
from glob import glob
import os
import pickle
import json
from configparser import ConfigParser
import operator
import itertools
import tensorflow as tf
from sklearn import metrics
import joblib
import numpy as np
from scipy.sparse import coo_matrix
from scipy import optimize
import matplotlib.pyplot as plt
import matplotlib.colors as colors
%matplotlib inline
```
## Results
### Framewise error rate
```
root = '/media/ildefonso/HD-LCU3/tf_syl_seg/koumura_repo_results/'
these_dirs_branch = [
('results_181014_194418', 'bird 0'),
('results_181015_075005', 'bird 1'),
('results_181014_194508', 'bird 2'),
#('results_180306_145149', 'bird 3'),
#('results_180308_134732', 'bird 4'),
('results_181017_105732', 'bird 5'),
('results_181016_010102', 'bird 6'),
('results_181016_074937', 'bird 7'),
('results_181015_103121', 'bird 9')
]
these_dirs = [(root + this_dir[0], this_dir[1]) for this_dir in these_dirs_branch]
config = ConfigParser()
all_results_list = []
for this_dir, bird_ID in these_dirs:
results_dict = {}
results_dict['dir'] = this_dir
os.chdir(this_dir)
config.read(glob('config*')[0])
results_dict['data_dir'] = config['DATA']['data_dir']
results_dict['time_steps'] = config['NETWORK']['time_steps']
results_dict['num_hidden'] = config['NETWORK']['num_hidden']
results_dict['train_set_durs'] = [int(element)
for element in
config['TRAIN']['train_set_durs'].split(',')]
with open(glob('summary*/train_err')[0], 'rb') as f:
results_dict['train_err'] = pickle.load(f)
with open(glob('summary*/test_err')[0], 'rb') as f:
results_dict['test_err'] = pickle.load(f)
pe = joblib.load(glob('summary*/y_preds_and_err_for_train_and_test')[0])
results_dict['train_syl_err_rate'] = pe['train_syl_err_rate']
results_dict['test_syl_err_rate'] = pe['test_syl_err_rate']
results_dict['bird_ID'] = bird_ID
all_results_list.append(results_dict)
os.chdir('..')
for el in all_results_list:
el['mn_test_err'] = np.mean(el['test_err'], axis=1)
el['mn_train_err'] = np.mean(el['train_err'], axis=1)
el['mn_train_syl_err'] = np.mean(el['train_syl_err_rate'], axis=1)
el['mn_test_syl_err'] = np.mean(el['test_syl_err_rate'], axis=1)
el['train_set_durs'] = np.asarray(el['train_set_durs'])
```
plot framewise error
```
all_mn_test_err = []
for el in all_results_list:
all_mn_test_err.append(el['mn_test_err'])
all_mn_test_err = np.asarray(all_mn_test_err)
plt.yticks?
plt.style.use('ggplot')
fig, ax = plt.subplots()
for el in all_results_list:
lbl = (el['bird_ID'])
ax.plot(el['train_set_durs'],
el['mn_test_err'],
label=lbl,
linestyle='--',
marker='o')
ax.plot(el['train_set_durs'],np.median(all_mn_test_err,axis=0),
linestyle='--',marker='o',linewidth=3,color='k',label='median across birds')
fig.set_size_inches(16,8)
plt.legend(fontsize=20)
plt.xticks(el['train_set_durs'])
plt.tick_params(axis='both', which='major', labelsize=20, rotation=45)
plt.title('Frame error rate as a function of training set size', fontsize=40)
plt.ylabel('Frame error rate\nas measured on test set', fontsize=32)
plt.xlabel('Training set size: duration in s', fontsize=32);
plt.tight_layout()
plt.savefig('frame-err-rate-v-train-set-size.png')
```
### syllable error rate
```
plt.style.use('ggplot')
fig, ax = plt.subplots()
fig.set_size_inches(16,8)
for el in all_results_list:
lbl = (el['bird_ID'])
ax.plot(el['train_set_durs'],
el['mn_test_syl_err'],
label=lbl,
linestyle=':',
marker='o')
plt.scatter(120,0.84,s=75)
plt.text(75,0.7,'Koumura & Okanoya 2016,\n0.84 note error rate\nwith 120s training data', fontsize=20)
plt.scatter(480,0.5,s=75)
plt.text(355,0.35,'Koumura & Okanoya 2016,\n0.46 note error rate\nwith 480s training data', fontsize=20)
plt.legend(fontsize=20, loc='upper right');
plt.title('Syllable error rate as a function of training set size', fontsize=40)
plt.xticks(el['train_set_durs'])
plt.tick_params(axis='both', which='major', labelsize=20, rotation=45)
plt.ylabel('Syllable error rate\nas measured on test set', fontsize=32)
plt.xlabel('Training set size: duration in s', fontsize=32);
plt.tight_layout()
plt.savefig('syl-error-rate-v-train-set-size.png')
```
| github_jupyter |
```
import cv2
import matplotlib.pyplot as plt
import numpy as np
import random
import os
from ssd_model import SSD300, SSD512
from ssd_utils import PriorUtil
from ssd_data import InputGenerator
from ssd_data import preprocess
from ssd_training import SSDLoss
from utils.model import load_weights
from data_voc import GTUtility
gt_util = GTUtility('data/VOC2007/')
gt_util_train, gt_util_val = gt_util.split(0.8)
experiment = 'ssd_voc'
num_classes = gt_util.num_classes
#model = SSD300(num_classes=num_classes)
model = SSD512(num_classes=num_classes)
image_size = model.image_size
_, inputs, images, data = gt_util_train.sample_random_batch(batch_size=16, input_size=image_size)
test_idx = 5
test_img = images[test_idx]
test_gt = data[test_idx]
plt.imshow(test_img)
gt_util.plot_gt(test_gt, show_labels=True)
plt.show()
prior_util = PriorUtil(model)
egt = prior_util.encode(test_gt)
idxs = np.where(np.logical_not(egt[:,4]))[0]
egt[idxs][:,:5]
#x = prior_util.encode(test_gt)
#y = prior_util.decode(x)
self = prior_util
from utils.vis import to_rec
def plot_assignment(self, map_idx):
ax = plt.gca()
im = plt.gci()
image_height, image_width = image_size = im.get_size()
# ground truth
boxes = self.gt_boxes
boxes_x = (boxes[:,0] + boxes[:,2]) / 2. * image_height
boxes_y = (boxes[:,1] + boxes[:,3]) / 2. * image_width
for box in boxes:
xy_rec = to_rec(box[:4], image_size)
ax.add_patch(plt.Polygon(xy_rec, fill=False, edgecolor='b', linewidth=2))
plt.plot(boxes_x, boxes_y, 'bo', markersize=8)
# prior boxes
for idx, box_idx in self.match_indices.items():
if idx >= self.map_offsets[map_idx] and idx < self.map_offsets[map_idx+1]:
x, y = self.priors_xy[idx]
w, h = self.priors_wh[idx]
plt.plot(x, y, 'ro', markersize=4)
plt.plot([x, boxes_x[box_idx]], [y, boxes_y[box_idx]], '-r', linewidth=1)
ax.add_patch(plt.Rectangle((x-w/2, y-h/2), w+1, h+1,
fill=False, edgecolor='y', linewidth=2))
from ssd_utils import iou
def encode(self, gt_data, overlap_threshold=0.5, debug=False):
# calculation is done with normalized sizes
gt_boxes = self.gt_boxes = np.copy(gt_data[:,:4]) # normalized xmin, ymin, xmax, ymax
gt_labels = self.gt_labels = np.copy(gt_data[:,4:]) # one_hot classes including background
num_priors = self.priors.shape[0]
num_classes = gt_labels.shape[1]
# TODO: empty ground truth
if gt_data.shape[0] == 0:
print('gt_data', type(gt_data), gt_data.shape)
gt_iou = np.array([iou(b, self.priors_norm) for b in gt_boxes]).T
# assigne gt to priors
max_idxs = np.argmax(gt_iou, axis=1)
max_val = gt_iou[np.arange(num_priors), max_idxs]
prior_mask = max_val > overlap_threshold
match_indices = max_idxs[prior_mask]
self.match_indices = dict(zip(list(np.argwhere(prior_mask)[:,0]), list(match_indices)))
# prior labels
confidence = np.zeros((num_priors, num_classes))
confidence[:,0] = 1
confidence[prior_mask] = gt_labels[match_indices]
# compute local offsets from ground truth boxes
gt_xy = (gt_boxes[:,2:4] + gt_boxes[:,0:2]) / 2.
gt_wh = gt_boxes[:,2:4] - gt_boxes[:,0:2]
gt_xy = gt_xy[match_indices]
gt_wh = gt_wh[match_indices]
priors_xy = self.priors_xy[prior_mask] / self.image_size
priors_wh = self.priors_wh[prior_mask] / self.image_size
offsets = np.zeros((num_priors, 4))
offsets[prior_mask, 0:2] = (gt_xy - priors_xy) / priors_wh
offsets[prior_mask, 2:4] = np.log(gt_wh / priors_wh)
offsets[prior_mask, :] /= self.priors[prior_mask,-4:] # variances
return np.concatenate([offsets, confidence], axis=1)
def decode(self, model_output, confidence_threshold=0.01, keep_top_k=200):
# calculation is done with normalized sizes
offsets = model_output[:,:4]
confidence = model_output[:,4:]
num_priors = offsets.shape[0]
num_classes = confidence.shape[1]
priors_xy = self.priors_xy / self.image_size
priors_wh = self.priors_wh / self.image_size
# compute bounding boxes from local offsets
boxes = np.empty((num_priors, 4))
offsets *= self.priors[:,-4:] # variances
boxes_xy = priors_xy + offsets[:,0:2] * priors_wh
boxes_wh = priors_wh * np.exp(offsets[:,2:4])
boxes[:,0:2] = boxes_xy - boxes_wh / 2. # xmin, ymin
boxes[:,2:4] = boxes_xy + boxes_wh / 2. # xmax, ymax
boxes = np.clip(boxes, 0.0, 1.0)
prior_mask = confidence > confidence_threshold
# TODO: number of confident boxes, compute bounding boxes only for those?
#print(np.sum(np.any(prior_mask[:,1:], axis=1)))
# do non maximum suppression
results = []
for c in range(1, num_classes):
mask = prior_mask[:,c]
boxes_to_process = boxes[mask]
if len(boxes_to_process) > 0:
confs_to_process = confidence[mask, c]
feed_dict = {
self.boxes: boxes_to_process,
self.scores: confs_to_process
}
idx = self.sess.run(self.nms, feed_dict=feed_dict)
good_boxes = boxes_to_process[idx]
good_confs = confs_to_process[idx][:, None]
labels = np.ones((len(idx),1)) * c
c_pred = np.concatenate((good_boxes, good_confs, labels), axis=1)
results.extend(c_pred)
results = np.array(results)
if len(results) > 0:
order = np.argsort(results[:, 1])[::-1]
results = results[order]
results = results[:keep_top_k]
self.results = results
return results
x = encode(self, test_gt)
y = decode(self, x)
#for idx in range(len(prior_util.prior_maps)):
for idx in [1,2,3,4]:
m = prior_util.prior_maps[idx]
plt.figure(figsize=[10]*2)
plt.imshow(test_img)
m.plot_locations()
#m.plot_boxes([0, 10, 100])
#gt_util.plot_gt(test_gt)
plot_assignment(self, idx)
prior_util.plot_results(y, show_labels=False)
plt.show()
enc = prior_util.encode(test_gt)
prior_util.decode(enc)[:,:4]
test_gt[:,:4]
# plot ground truth
for i in range(4):
plt.figure(figsize=[8]*2)
plt.imshow(images[i])
gt_util.plot_gt(data[i])
plt.show()
# plot prior boxes
for i, m in enumerate(prior_util.prior_maps):
plt.figure(figsize=[8]*2)
#plt.imshow(images[7])
plt.imshow(images[8])
m.plot_locations()
#m.plot_boxes([0, 10, 100])
m.plot_boxes([0])
plt.axis('off')
#plt.savefig('plots/ssd_priorboxes_%i.pgf' % (i), bbox_inches='tight')
#print(m.map_size)
plt.show()
for l in model.layers:
try:
ks = l.weights[0].shape
except:
ks = ""
print("%30s %16s %24s %20s" % (l.name, l.__class__.__name__, l.output_shape, ks))
model.summary()
batch_size = 24
gen = InputGenerator(gt_util_train, prior_util, batch_size, model.image_size,
augmentation=False,
vflip_prob=0.0, do_crop=False)
g = gen.generate()
%%time
for j in range(3000):
i, o = next(g)
print(j, len(i))
if len(i) != batch_size:
print('FOOOOOOOOOOO')
import cProfile
g = gen.generate()
p = cProfile.Profile()
p.enable()
i, o = next(g)
p.disable()
p.print_stats(sort='cumulative')
```
| github_jupyter |
# <img style="float: left; padding-right: 10px; width: 45px" src="https://raw.githubusercontent.com/Harvard-IACS/2018-CS109A/master/content/styles/iacs.png"> CS-109B Introduction to Data Science
## Lab 5: Convolutional Neural Networks
**Harvard University**<br>
**Spring 2019**<br>
**Lab instructor:** Eleni Kaxiras<br>
**Instructors:** Pavlos Protopapas and Mark Glickman<br>
**Authors:** Eleni Kaxiras, Pavlos Protopapas, Patrick Ohiomoba, and Davis Sontag
```
# RUN THIS CELL TO PROPERLY HIGHLIGHT THE EXERCISES
import requests
from IPython.core.display import HTML
styles = requests.get("https://raw.githubusercontent.com/Harvard-IACS/2019-CS109B/master/content/styles/cs109.css").text
HTML(styles)
```
## Learning Goals
In this lab we will look at Convolutional Neural Networks (CNNs), and their building blocks.
By the end of this lab, you should:
- know how to put together the building blocks used in CNNs - such as convolutional layers and pooling layers - in `keras` with an example.
- have a good undertanding on how images, a common type of data for a CNN, are represented in the computer and how to think of them as arrays of numbers.
- be familiar with preprocessing images with `keras` and `sckit-learn`.
- use `keras-viz` to produce Saliency maps.
- learn best practices for configuring the hyperparameters of a CNN.
- run your first CNN and see the error rate.
```
import matplotlib.pyplot as plt
plt.rcParams["figure.figsize"] = (5,5)
import numpy as np
from scipy.optimize import minimize
import tensorflow as tf
import keras
from keras import layers
from keras import models
from keras import utils
from keras.layers import Dense
from keras.models import Sequential
from keras.layers import Flatten
from keras.layers import Dropout
from keras.layers import Activation
from keras.regularizers import l2
from keras.optimizers import SGD
from keras.optimizers import RMSprop
from keras import datasets
from keras.preprocessing.image import ImageDataGenerator
from keras.callbacks import LearningRateScheduler
from keras.callbacks import History
from keras import losses
from keras.datasets import mnist
from keras.utils import to_categorical
from sklearn.utils import shuffle
print(tf.VERSION)
print(tf.keras.__version__)
%matplotlib inline
```
## Prologue: `keras-viz` Visualization Toolkit
`keras-vis` is a high-level toolkit for visualizing and debugging your trained keras neural net models. Currently supported visualizations include:
- Activation maximization
- **Saliency maps**
- Class activation maps
All visualizations by default support N-dimensional image inputs. i.e., it generalizes to N-dim image inputs to your model. Compatible with both theano and tensorflow backends with 'channels_first', 'channels_last' data format.
Read the documentation at https://raghakot.github.io/keras-vis.https://github.com/raghakot/keras-vis
To install use `pip install git+https://github.com/raghakot/keras-vis.git --upgrade`
## SEAS JupyterHub
[Instructions for Using SEAS JupyterHub](https://canvas.harvard.edu/courses/48088/pages/instructions-for-using-seas-jupyterhub)
SEAS and FAS are providing you with a platform in AWS to use for the class (accessible from the 'Jupyter' menu link in Canvas). These are AWS p2 instances with a GPU, 10GB of disk space, and 61 GB of RAM, for faster training for your networks. Most of the libraries such as keras, tensorflow, pandas, etc. are pre-installed. If a library is missing you may install it via the Terminal.
**NOTE : The AWS platform is funded by SEAS and FAS for the purposes of the class. It is not running against your individual credit. You are not allowed to use it for purposes not related to this course.**
**Help us keep this service: Make sure you stop your instance as soon as you do not need it.**

## Part 1: Parts of a Convolutional Neural Net
There are three types of layers in a Convolutional Neural Network:
- Convolutional Layers
- Pooling Layers.
- Dropout Layers.
- Fully Connected Layers.
### a. Convolutional Layers.
Convolutional layers are comprised of **filters** and **feature maps**. The filters are essentially the **neurons** of the layer. They have the weights and produce the input for the next layer. The feature map is the output of one filter applied to the previous layer.
The fundamental difference between a densely connected layer and a convolution layer is that dense layers learn global patterns in their input feature space (for example, for an MNIST digit, patterns involving all pixels), whereas convolution layers learn local patterns: in the case of images, patterns found in small 2D windows of the inputs called *receptive fields*.
This key characteristic gives convnets two interesting properties:
- The patterns they learn are **translation invariant**. After learning a certain pattern in the lower-right corner of a picture, a convnet can recognize it anywhere: for example, in the upper-left corner. A densely connected network would have to learn the pattern anew if it appeared at a new location. This makes convnets data efficient when processing images (because the visual world is fundamentally translation invariant): they need fewer training samples to learn representations that have generalization power.
- They can learn **spatial hierarchies of patterns**. A first convolution layer will learn small local patterns such as edges, a second convolution layer will learn larger patterns made of the features of the first layers, and so on. This allows convnets to efficiently learn increasingly complex and abstract visual concepts (because the visual world is fundamentally spatially hierarchical).
Convolutions operate over 3D tensors, called feature maps, with two spatial axes (height and width) as well as a depth axis (also called the channels axis). For an RGB image, the dimension of the depth axis is 3, because the image has three color channels: red, green, and blue. For a black-and-white picture, like the MNIST digits, the depth is 1 (levels of gray). The convolution operation extracts patches from its input feature map and applies the same transformation to all of these patches, producing an output feature map. This output feature map is still a 3D tensor: it has a width and a height. Its depth can be arbitrary, because the output depth is a parameter of the layer, and the different channels in that depth axis no longer stand for specific colors as in RGB input; rather, they stand for filters. Filters encode specific aspects of the input data: at a high level, a single filter could encode the concept “presence of a face in the input,” for instance.
In the MNIST example that we will see, the first convolution layer takes a feature map of size (28, 28, 1) and outputs a feature map of size (26, 26, 32): it computes 32 filters over its input. Each of these 32 output channels contains a 26×26 grid of values, which is a response map of the filter over the input, indicating the response of that filter pattern at different locations in the input.
Convolutions are defined by two key parameters:
- Size of the patches extracted from the inputs. These are typically 3×3 or 5×5
- The number of filters computed by the convolution.
**Padding**: One of "valid", "causal" or "same" (case-insensitive). "valid" means "no padding". "same" results in padding the input such that the output has the same length as the original input. "causal" results in causal (dilated) convolutions,
In `keras` see [convolutional layers](https://keras.io/layers/convolutional/)
**keras.layers.Conv2D**(filters, kernel_size, strides=(1, 1), padding='valid', activation=None, use_bias=True,
kernel_initializer='glorot_uniform', data_format='channels_last',
bias_initializer='zeros')
#### How are the values in feature maps calculated?

### Exercise 1:
- Compute the operations by hand (assuming zero padding and same arrays for all channels) to produce the first element of the 4x4 feature map. How did we get the 4x4 output size?
- Write this Conv layer in keras
-- your answer here
### b. Pooling Layers.
Pooling layers are also comprised of filters and feature maps. Let's say the pooling layer has a 2x2 receptive field and a stride of 2. This stride results in feature maps that are one half the size of the input feature maps. We can use a max() operation for each receptive field.
In `keras` see [pooling layers](https://keras.io/layers/pooling/)
**keras.layers.MaxPooling2D**(pool_size=(2, 2), strides=None, padding='valid', data_format=None)

### c. Dropout Layers.
Dropout consists in randomly setting a fraction rate of input units to 0 at each update during training time, which helps prevent overfitting.
In `keras` see [Dropout layers](https://keras.io/layers/core/)
keras.layers.Dropout(rate, seed=None)
rate: float between 0 and 1. Fraction of the input units to drop.<br>
seed: A Python integer to use as random seed.
References
[Dropout: A Simple Way to Prevent Neural Networks from Overfitting](http://www.jmlr.org/papers/volume15/srivastava14a/srivastava14a.pdf)
### d. Fully Connected Layers.
A fully connected layer flattens the square feature map into a vector. Then we can use a sigmoid or softmax activation function to output probabilities of classes.
In `keras` see [FC layers](https://keras.io/layers/core/)
**keras.layers.Dense**(units, activation=None, use_bias=True,
kernel_initializer='glorot_uniform', bias_initializer='zeros')
#### IT'S ALL ABOUT THE HYPERPARAMETERS!
- stride
- size of filter
- number of filters
- poolsize
## Part 2: Preprocessing the data
### Taking a look at how images are represented in a computer using a photo of a Picasso sculpture
```
img = plt.imread('data/picasso.png')
img.shape
img[1,:,1]
print(type(img[50][0][0]))
# let's see the image
imgplot = plt.imshow(img)
```
#### Visualizing the channels
```
R_img = img[:,:,0]
G_img = img[:,:,1]
B_img = img[:,:,2]
plt.subplot(221)
plt.imshow(R_img, cmap=plt.cm.Reds)
plt.subplot(222)
plt.imshow(G_img, cmap=plt.cm.Greens)
plt.subplot(223)
plt.imshow(B_img, cmap=plt.cm.Blues)
plt.subplot(224)
plt.imshow(img)
plt.show()
```
More on preprocessing data below!
If you want to learn more: [Image Processing with Python and Scipy](http://prancer.physics.louisville.edu/astrowiki/index.php/Image_processing_with_Python_and_SciPy)
## Part 3: Putting the Parts together to make a small ConvNet Model
Let's put all the parts together to make a convnet for classifying our good old MNIST digits.
```
# Load data and preprocess
(train_images, train_labels), (test_images, test_labels) = mnist.load_data() # load MNIST data
train_images.shape
train_images.max(), train_images.min()
train_images = train_images.reshape((60000, 28, 28, 1)) # Reshape to get third dimension
train_images = train_images.astype('float32') / 255 # Normalize between 0 and 1
test_images = test_images.reshape((10000, 28, 28, 1)) # Reshape to get third dimension
test_images = test_images.astype('float32') / 255 # Normalize between 0 and 1
# Convert labels to categorical data
train_labels = to_categorical(train_labels)
test_labels = to_categorical(test_labels)
mnist_cnn_model = models.Sequential() # Create sequential model
# Add network layers
mnist_cnn_model.add(layers.Conv2D(32, (3, 3), activation='relu', input_shape=(28, 28, 1)))
mnist_cnn_model.add(layers.MaxPooling2D((2, 2)))
mnist_cnn_model.add(layers.Conv2D(64, (3, 3), activation='relu'))
mnist_cnn_model.add(layers.MaxPooling2D((2, 2)))
mnist_cnn_model.add(layers.Conv2D(64, (3, 3), activation='relu'))
```
The next step is to feed the last output tensor (of shape (3, 3, 64)) into a densely connected classifier network like those you’re already familiar with: a stack of Dense layers. These classifiers process vectors, which are 1D, whereas the current output is a 3D tensor. First we have to flatten the 3D outputs to 1D, and then add a few Dense layers on top.
```
mnist_cnn_model.add(layers.Flatten())
mnist_cnn_model.add(layers.Dense(64, activation='relu'))
mnist_cnn_model.add(layers.Dense(10, activation='softmax'))
mnist_cnn_model.summary()
# Compile model
mnist_cnn_model.compile(optimizer='rmsprop',
loss='categorical_crossentropy',
metrics=['accuracy'])
# Fit the model
mnist_cnn_model.fit(train_images, train_labels, epochs=5, batch_size=64)
# Evaluate the model on the test data:
test_loss, test_acc = mnist_cnn_model.evaluate(test_images, test_labels)
test_acc
```
A densely connected network (MLP) running MNIST usually has a test accuracy of 97.8%, whereas our basic convnet has a test accuracy of 99.03%: we decreased the error rate by 68% (relative) with only 5 epochs. Not bad! But why does this simple convnet work so well, compared to a densely connected model? The answer is above on how convolutional layers work!
### Data Preprocessing : Meet the `ImageDataGenerator` class in `keras` [(docs)](https://keras.io/preprocessing/image/)
The MNIST and other pre-loaded dataset are formatted in a way that is almost ready for feeding into the model. What about plain images? They should be formatted into appropriately preprocessed floating-point tensors before being fed into the network.
The Dogs vs. Cats dataset that you’ll use isn’t packaged with Keras. It was made available by Kaggle as part of a computer-vision competition in late 2013, back when convnets weren’t mainstream. The data has been downloaded for you from https://www.kaggle.com/c/dogs-vs-cats/data The pictures are medium-resolution color JPEGs.
```
# TODO: set your base dir to your correct local location
base_dir = 'data/cats_and_dogs_small'
import os, shutil
# Set up directory information
train_dir = os.path.join(base_dir, 'train')
validation_dir = os.path.join(base_dir, 'validation')
test_dir = os.path.join(base_dir, 'test')
train_cats_dir = os.path.join(train_dir, 'cats')
train_dogs_dir = os.path.join(train_dir, 'dogs')
validation_cats_dir = os.path.join(validation_dir, 'cats')
validation_dogs_dir = os.path.join(validation_dir, 'dogs')
test_cats_dir = os.path.join(test_dir, 'cats')
test_dogs_dir = os.path.join(test_dir, 'dogs')
print('total training cat images:', len(os.listdir(train_cats_dir)))
print('total training dog images:', len(os.listdir(train_dogs_dir)))
print('total validation cat images:', len(os.listdir(validation_cats_dir)))
print('total validation dog images:', len(os.listdir(validation_dogs_dir)))
print('total test cat images:', len(os.listdir(test_cats_dir)))
print('total test dog images:', len(os.listdir(test_dogs_dir)))
```
So you do indeed have 2,000 training images, 1,000 validation images, and 1,000 test images. Each split contains the same number of samples from each class: this is a balanced binary-classification problem, which means classification accuracy will be an appropriate measure of success.
#### Building the network
```
from keras import layers
from keras import models
model = models.Sequential()
model.add(layers.Conv2D(32, (3, 3), activation='relu',
input_shape=(150, 150, 3)))
model.add(layers.MaxPooling2D((2, 2)))
model.add(layers.Conv2D(64, (3, 3), activation='relu'))
model.add(layers.MaxPooling2D((2, 2)))
model.add(layers.Conv2D(128, (3, 3), activation='relu'))
model.add(layers.MaxPooling2D((2, 2)))
model.add(layers.Conv2D(128, (3, 3), activation='relu'))
model.add(layers.MaxPooling2D((2, 2)))
model.add(layers.Flatten())
model.add(layers.Dense(512, activation='relu'))
model.add(layers.Dense(1, activation='sigmoid'))
model.summary()
```
For the compilation step, you’ll go with the RMSprop optimizer. Because you ended the network with a single sigmoid unit, you’ll use binary crossentropy as the loss.
```
from keras import optimizers
model.compile(loss='binary_crossentropy',
optimizer=optimizers.RMSprop(lr=1e-4),
metrics=['acc'])
```
The steps for getting it into the network are roughly as follows:
1. Read the picture files.
2. Decode the JPEG content to RGB grids of pixels.
3. Convert these into floating-point tensors.
4. Rescale the pixel values (between 0 and 255) to the [0, 1] interval (as you know, neural networks prefer to deal with small input values).
It may seem a bit daunting, but fortunately Keras has utilities to take care of these steps automatically with the class `ImageDataGenerator`, which lets you quickly set up Python generators that can automatically turn image files on disk into batches of preprocessed tensors. This is what you’ll use here.
```
from keras.preprocessing.image import ImageDataGenerator
train_datagen = ImageDataGenerator(rescale=1./255)
test_datagen = ImageDataGenerator(rescale=1./255)
train_generator = train_datagen.flow_from_directory(
train_dir,
target_size=(150, 150),
batch_size=20,
class_mode='binary')
validation_generator = test_datagen.flow_from_directory(
validation_dir,
target_size=(150, 150),
batch_size=20,
class_mode='binary')
```
Let’s look at the output of one of these generators: it yields batches of 150×150 RGB images (shape (20, 150, 150, 3)) and binary labels (shape (20,)). There are 20 samples in each batch (the batch size). Note that the generator yields these batches indefinitely: it loops endlessly over the images in the target folder. For this reason, you need to break the iteration loop at some point:
```
for data_batch, labels_batch in train_generator:
print('data batch shape:', data_batch.shape)
print('labels batch shape:', labels_batch.shape)
break
```
Let’s fit the model to the data using the generator. You do so using the `.fit_generator` method, the equivalent of `.fit` for data generators like this one. It expects as its first argument a Python generator that will yield batches of inputs and targets indefinitely, like this one does.
Because the data is being generated endlessly, the Keras model needs to know how many samples to draw from the generator before declaring an epoch over. This is the role of the `steps_per_epoch` argument: after having drawn steps_per_epoch batches from the generator—that is, after having run for steps_per_epoch gradient descent steps - the fitting process will go to the next epoch. In this case, batches are 20 samples, so it will take 100 batches until you see your target of 2,000 samples.
When using fit_generator, you can pass a validation_data argument, much as with the fit method. It’s important to note that this argument is allowed to be a data generator, but it could also be a tuple of Numpy arrays. If you pass a generator as validation_data, then this generator is expected to yield batches of validation data endlessly; thus you should also specify the validation_steps argument, which tells the process how many batches to draw from the validation generator for evaluation
```
history = model.fit_generator(
train_generator,
steps_per_epoch=100,
epochs=5, # TODO: should be 30
validation_data=validation_generator,
validation_steps=50)
# It’s good practice to always save your models after training.
model.save('cats_and_dogs_small_1.h5')
```
Let’s plot the accuracy of the model over the training and validation data during training:
```
fig, ax = plt.subplots(1, 1, figsize=(10,6))
ax.plot((history.history['acc']), 'r', label='train')
ax.plot((history.history['val_acc']), 'b' ,label='val')
ax.set_xlabel(r'Epoch', fontsize=20)
ax.set_ylabel(r'Accuracy', fontsize=20)
ax.legend()
ax.tick_params(labelsize=20)
```
Let's try data augmentation
```
datagen = ImageDataGenerator(
rotation_range=40,
width_shift_range=0.2,
height_shift_range=0.2,
shear_range=0.2,
zoom_range=0.2,
horizontal_flip=True,
fill_mode='nearest')
```
These are just a few of the options available (for more, see the Keras documentation).
Let’s quickly go over this code:
- rotation_range is a value in degrees (0–180), a range within which to randomly rotate pictures.
- width_shift and height_shift are ranges (as a fraction of total width or height) within which to randomly translate pictures vertically or horizontally.
- shear_range is for randomly applying shearing transformations.
- zoom_range is for randomly zooming inside pictures.
- horizontal_flip is for randomly flipping half the images horizontally—relevant when there are no assumptions of - horizontal asymmetry (for example, real-world pictures).
- fill_mode is the strategy used for filling in newly created pixels, which can appear after a rotation or a width/height shift.
Let’s look at the augmented images
```
from keras.preprocessing import image
fnames = [os.path.join(train_dogs_dir, fname) for
fname in os.listdir(train_dogs_dir)]
img_path = fnames[3] # Chooses one image to augment
img = image.load_img(img_path, target_size=(150, 150))
# Reads the image and resizes it
x = image.img_to_array(img) # Converts it to a Numpy array with shape (150, 150, 3)
x = x.reshape((1,) + x.shape) # Reshapes it to (1, 150, 150, 3)
i=0
for batch in datagen.flow(x, batch_size=1):
plt.figure(i)
imgplot = plt.imshow(image.array_to_img(batch[0]))
i += 1
if i % 4 == 0:
break
plt.show()
```
If you train a new network using this data-augmentation configuration, the network will never see the same input twice. But the inputs it sees are still heavily intercorrelated, because they come from a small number of original images—you can’t produce new information, you can only remix existing information. As such, this may not be enough to completely get rid of overfitting. To further fight overfitting, you’ll also add a **Dropout** layer to your model right before the densely connected classifier.
```
model = models.Sequential()
model.add(layers.Conv2D(32, (3, 3), activation='relu',
input_shape=(150, 150, 3)))
model.add(layers.MaxPooling2D((2, 2)))
model.add(layers.Conv2D(64, (3, 3), activation='relu'))
model.add(layers.MaxPooling2D((2, 2)))
model.add(layers.Conv2D(128, (3, 3), activation='relu'))
model.add(layers.MaxPooling2D((2, 2)))
model.add(layers.Conv2D(128, (3, 3), activation='relu'))
model.add(layers.MaxPooling2D((2, 2)))
model.add(layers.Flatten())
model.add(layers.Dropout(0.5))
model.add(layers.Dense(512, activation='relu'))
model.add(layers.Dense(1, activation='sigmoid'))
model.compile(loss='binary_crossentropy',
optimizer=optimizers.RMSprop(lr=1e-4),
metrics=['acc'])
# Let’s train the network using data augmentation and dropout.
train_datagen = ImageDataGenerator(
rescale=1./255,
rotation_range=40,
width_shift_range=0.2,
height_shift_range=0.2,
shear_range=0.2,
zoom_range=0.2,
horizontal_flip=True,)
test_datagen = ImageDataGenerator(rescale=1./255)
# Note that the validation data shouldn’t be augmented!
train_generator = train_datagen.flow_from_directory(
train_dir,
target_size=(150, 150),
batch_size=32,
class_mode='binary')
validation_generator = test_datagen.flow_from_directory(
validation_dir,
target_size=(150, 150),
batch_size=32,
class_mode='binary')
history = model.fit_generator(
train_generator,
steps_per_epoch=100,
epochs=5, # TODO: should be 100
validation_data=validation_generator,
validation_steps=50)
model.save('cats_and_dogs_small_2.h5')
```
And let’s plot the results again. Thanks to data augmentation and dropout, you’re no longer overfitting: the training curves are closely tracking the validation curves. You now reach an accuracy of 82%, a 15% relative improvement over the non-regularized model. (Note: these numbers are for 100 epochs..)
```
fig, ax = plt.subplots(1, 1, figsize=(10,6))
ax.plot((history.history['acc']), 'r', label='train')
ax.plot((history.history['val_acc']), 'b' ,label='val')
ax.set_xlabel(r'Epoch', fontsize=20)
ax.set_ylabel(r'Accuracy', fontsize=20)
ax.legend()
ax.tick_params(labelsize=20)
```
By using regularization techniques even further, and by tuning the network’s parameters (such as the number of filters per convolution layer, or the number of layers in the network), you may be able to get an even better accuracy, likely up to 86% or 87%. But it would prove difficult to go any higher just by training your own convnet from scratch, because you have so little data to work with. As a next step to improve your accuracy on this problem, you’ll have to use a pretrained model.
## Part 4: keras viz toolkit
https://github.com/raghakot/keras-vis/blob/master/examples/mnist/attention.ipynb
```
class_idx = 0
indices = np.where(test_labels[:, class_idx] == 1.)[0]
# pick some random input from here.
idx = indices[0]
# Lets sanity check the picked image.
from matplotlib import pyplot as plt
%matplotlib inline
plt.rcParams['figure.figsize'] = (18, 6)
plt.imshow(test_images[idx][..., 0])
input_shape=(28, 28, 1)
num_classes = 10
batch_size = 128
epochs = 5
model = Sequential()
model.add(layers.Conv2D(32, kernel_size=(3, 3),
activation='relu',
input_shape=input_shape))
model.add(layers.Conv2D(64, (3, 3), activation='relu'))
model.add(layers.MaxPooling2D(pool_size=(2, 2)))
model.add(layers.Dropout(0.25))
model.add(layers.Flatten())
model.add(layers.Dense(128, activation='relu'))
model.add(layers.Dropout(0.5))
model.add(layers.Dense(num_classes, activation='softmax', name='preds'))
model.compile(loss=keras.losses.categorical_crossentropy,
optimizer=keras.optimizers.Adam(),
metrics=['accuracy'])
model.fit(train_images, train_labels,
batch_size=batch_size,
epochs=epochs,
verbose=1,
validation_data=(test_images, test_labels))
score = model.evaluate(test_images, test_labels, verbose=0)
print('Test loss:', score[0])
print('Test accuracy:', score[1])
from vis.visualization import visualize_saliency
from vis.utils import utils
from keras import activations
# Utility to search for layer index by name.
# Alternatively we can specify this as -1 since it corresponds to the last layer.
layer_idx = utils.find_layer_idx(model, 'preds')
plt.rcParams["figure.figsize"] = (5,5)
from vis.visualization import visualize_cam
import warnings
warnings.filterwarnings('ignore')
# This corresponds to the Dense linear layer.
for class_idx in np.arange(10):
indices = np.where(test_labels[:, class_idx] == 1.)[0]
idx = indices[0]
f, ax = plt.subplots(1, 4)
ax[0].imshow(test_images[idx][..., 0])
for i, modifier in enumerate([None, 'guided', 'relu']):
grads = visualize_cam(model, layer_idx, filter_indices=class_idx,
seed_input=test_images[idx], backprop_modifier=modifier)
if modifier is None:
modifier = 'vanilla'
ax[i+1].set_title(modifier)
ax[i+1].imshow(grads, cmap='jet')
```
#### References and Acknowledgements
The cats and dogs part of this lab is based on the book Deep Learning with Python, Chapter 5 written by the Francois Chollet, the author of Keras. It is a very practical introduction to Deep Learning. It is appropriate for those with some Python knowledge who want to start with machine learning.
The saliency maps are from https://github.com/raghakot/keras-vis/blob/master/examples/mnist/attention.ipynb
| github_jupyter |
<table class="ee-notebook-buttons" align="left">
<td><a target="_blank" href="https://github.com/giswqs/earthengine-py-notebooks/tree/master/Image/composite_bands.ipynb"><img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" /> View source on GitHub</a></td>
<td><a target="_blank" href="https://nbviewer.jupyter.org/github/giswqs/earthengine-py-notebooks/blob/master/Image/composite_bands.ipynb"><img width=26px src="https://upload.wikimedia.org/wikipedia/commons/thumb/3/38/Jupyter_logo.svg/883px-Jupyter_logo.svg.png" />Notebook Viewer</a></td>
<td><a target="_blank" href="https://colab.research.google.com/github/giswqs/earthengine-py-notebooks/blob/master/Image/composite_bands.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" /> Run in Google Colab</a></td>
</table>
## Install Earth Engine API and geemap
Install the [Earth Engine Python API](https://developers.google.com/earth-engine/python_install) and [geemap](https://geemap.org). The **geemap** Python package is built upon the [ipyleaflet](https://github.com/jupyter-widgets/ipyleaflet) and [folium](https://github.com/python-visualization/folium) packages and implements several methods for interacting with Earth Engine data layers, such as `Map.addLayer()`, `Map.setCenter()`, and `Map.centerObject()`.
The following script checks if the geemap package has been installed. If not, it will install geemap, which automatically installs its [dependencies](https://github.com/giswqs/geemap#dependencies), including earthengine-api, folium, and ipyleaflet.
```
# Installs geemap package
import subprocess
try:
import geemap
except ImportError:
print('Installing geemap ...')
subprocess.check_call(["python", '-m', 'pip', 'install', 'geemap'])
import ee
import geemap
```
## Create an interactive map
The default basemap is `Google Maps`. [Additional basemaps](https://github.com/giswqs/geemap/blob/master/geemap/basemaps.py) can be added using the `Map.add_basemap()` function.
```
Map = geemap.Map(center=[40,-100], zoom=4)
Map
```
## Add Earth Engine Python script
```
# Add Earth Engine dataset
# There are many fine places to look here is one. Comment
# this out if you want to twiddle knobs while panning around.
Map.setCenter(-61.61625, -11.64273, 14)
# Grab a sample L7 image and pull out the RGB and pan bands
# in the range (0, 1). (The range of the pan band values was
# chosen to roughly match the other bands.)
image1 = ee.Image('LANDSAT/LE7/LE72300681999227EDC00')
rgb = image1.select('B3', 'B2', 'B1').unitScale(0, 255)
gray = image1.select('B8').unitScale(0, 155)
# Convert to HSV, swap in the pan band, and convert back to RGB.
huesat = rgb.rgbToHsv().select('hue', 'saturation')
upres = ee.Image.cat(huesat, gray).hsvToRgb()
# Display before and after layers using the same vis parameters.
visparams = {'min': [.15, .15, .25], 'max': [1, .9, .9], 'gamma': 1.6}
Map.addLayer(rgb, visparams, 'Orignal')
Map.addLayer(upres, visparams, 'Pansharpened')
```
## Display Earth Engine data layers
```
Map.addLayerControl() # This line is not needed for ipyleaflet-based Map.
Map
```
| github_jupyter |
```
import pandas as pd
import numpy as np
%matplotlib inline
import matplotlib.pyplot as plt
```
=================================================================================================================
# Lecture Notes: Linear Support Vector Machines
##### D.Vidotto, Data Mining: JBI030 2019/2020
=================================================================================================================
Linear Support Vector Machines (SVM) is a classifier that, similarly to logistic regression, tries to find a *linear boundary* to separate the classes. Unlike Logistic Regression, however, SVM finds such a boundary under a geometric perspective, rather than a probabilistic one.
For this notebook, students are assumed to have become familiar with the following concepts during the class:
* (hard/soft) margins
* support vectors
* maximum margin hyperplane
* linearly separable/non-separable data
In this notebook, we will review such concepts and will see how to implement SVM in Python:
1. SVM for Linearly Separable Data
* Geometric Intuition
* The optimization problem
* Decision function, predictions, and comparison with Logistic Regression
1. SVM for Non-Separable Data
* The slack variables
* The New Optimization Problem
* Regularization
1. SVM: Other remarks
1. Examples in Python
* Two-Class case
* Multi-Class case
1. Application to the Heart dataset
## 1. SVM for Linearly Separable Data
### 1.1 Geometric Intuition
Consider the following dataset (2 features; and 2 classes called -1 (negative class) and 1, the positive class):
<img src="./img/linear_svm/lin_sep_svm.png" width="350" height="50"/>
In this case the classes are said to be *linearly separable*: there is at least one straight line (a hyperplane in multiple dimensions) that can perfectly separate examples of the negative class from examples of the positive class. Actually, the number of linear boundaries that we can find to divide the two classes is potentially infinite:
<img src="./img/linear_svm/lin_sep_svm_separated.png" width="350" height="50"/>
The [Perceptron](https://en.wikipedia.org/wiki/Perceptron) (not covered in this course) is an algorithm that finds a separating hyperplane in case of linearly separable data, as the ones shown in this example. However, the Perceptron will rarely find a unique solution: it can find potentially an infinite number of hyperplanes as we slightly perturb the training sample, or as we modify the starting values of its algorithm.
**Support Vector Machines** is a Machine Learning algorithm that allows finding a unique and stable solution by finding a well-defined hyperplane: the one that is equally distant from both classes. Thus, if we define the *margin* as the shortest distance between the hyperplane and the closest data point, SVM seeks to find the **maximum margin hyperplane**: that is, the hyperplane that maximizes the distances (the margins) from the data points of both classes. Consider the following two hyperplanes for the dataset introduced above:
<img src="./img/linear_svm/svm_two_hyperplanes.png" width="800" height="50"/>
It seems clear from the figure that the hyperplane on the right is better than the one on the left, as it seems equally far away from the closes examples of both classes (the hyperplane on the left seems much closer to the positive class), and therefore it looks reasonable to assume that it can generalize better to new examples. Indeed, the hyperplane of the right plot is the *maximum margin hyperplane* found by fitting SVM on the dataset. By definition, it leaves exactly the same distance from the closest point of the two classes:
<img src="./img/linear_svm/svm_support_vectors.png" width="350" height="50"/>
The circled data points are the closest to the optimal hyperplane; such points are called *support vectors* (hence the algorithm name). The dashed lines represent the two margins of this maximum margin hyperplane; by definition, the distances of the support vectors from the hyperplane are exactly the same for both the positive and the negative class. Notice that, once trained, SVM only needs the support vectors to define the hyperplane (discarding the remaining data points), and therefore to perform predictions. Similar to logistic regression, predictions occur by considering in which part of the hyperplane the new data points fall, and classifying it accordinly. Unlike logistic regression, however, SVM doesn't use a probabilistic model, and therefore it cannot be interpreted probabilistically. Notice, also, that if we find a hyperplane that has a lower margin from one class w.r.t. the other class, we can always move it in such a way to obtain maximum margin from both classes (Why is that? Try to give your own intuition.)
This type of margin found by SVM is also known as **hard margin**, as it seeks to perfectly separate the two classes, not allowing for miss-classifications of the data points. We are now going to review the model more formally.
## 1.2 The optimization problem
Here we are going to use the following notation:
* $y_i \in \{-1,1\}$, where -1 denoted the negative class and 1 the positive class (As seen for the logistic regression model, the labels are encoded internally by `scikit-learn` during the optimization stage)
* The *linear decision function* for observation $i$ is defined as $f(\mathbf{x})=w_0+\mathbf{w}^T\mathbf{x}_i=w_0+w_1x_{i1}+...+w_px_{ip}$, where the weights (or coefficients) vector $\mathbf{w}=[w_1,....,w_p]$ and the $i$-th unit vector $\mathbf{x}_i=[x_{i1},...,x_{ip}]$ are defined analogously to what done in the logistic regression notebook, and the same for the *bias* (or intercept) term $w_0$
* the optimal hyperplane (the decision boundary) is defined by all those points $x_1,...,x_p$ such that $w_0+\mathbf{w}^T\mathbf{x}=0$; this will be denoted with $\mathcal{H}_{w_0,\mathbf{w}}=\{x: w_0+\mathbf{w}^T\mathbf{x}=0\}$
* we use the Euclidean norm to define the length of a vector; such norm is denoted with $||\mathbf{w}||_2=\sqrt{\sum_{j=1}^{p}w_j^2}$ (to simplify notation, the subscript $_2$ will be dropped, and the norm will be simply denoted with $||\mathbf{w}||$)
With the help of a graph we are going to add some other definitions before delving into the optimization problem:
<img src="./img/linear_svm/svm_definitions.png" width="900" height="50"/>
From the figures we can retrieve the following information:
* the weights vector $\mathbf{w}$ is orthogonal to the hyperplane (by definition)
* the (normalized) distance between each margin (and therefore the support vectors) and the optimum hyperplane is $\frac{1}{||\mathbf{w}||}$ (more on this shortly); the total margin size is therefore $\frac{2}{||\mathbf{w}||}$
* the equations for the hard margins are $w_0+\mathbf{w}^T\mathbf{x}=1$ (margin of the positive class) and $w_0+\mathbf{w}^T\mathbf{x}=-1$ (margin of the negative class)
* the *projection* of a generic point $\mathbf{x}$ onto the hyperplane is denoted with $\mathbf{x}_p$, and the vector $\mathbf{d}$ contains the distances between the coordinates of $\mathbf{x}$ and the coordinates of its projection ($\mathbf{d} = \mathbf{x} - \mathbf{x}_p$)
Now, how do we formalize the SVM model?
* Consider the projection of a generic point $\mathbf{x}$, namely $\mathbf{x}_p = \mathbf{x}-\mathbf{d}$: because it lies in the hyperplane, we know that $w_0+\mathbf{w}^T\mathbf{x}_p = 0$ and as a consequence $$w_0+\mathbf{w}^T(\mathbf{x}-\mathbf{d}) = 0$$
<br>
* because $\mathbf{w}$ and $\mathbf{d}$ point to the same direction (for any $\mathbf{x})$, we can see $\mathbf{d}$ as a rescaled version of the weights vector, $\mathbf{d}=c \cdot\mathbf{w}$ with $c$ a real scalar; therefore $$w_0+\mathbf{w}^T(\mathbf{x}-c\cdot\mathbf{w}) = 0$$
<br>
* solving the above equation for $c$, we obtain $c=\frac{w_0+\mathbf{w}^T\mathbf{x}}{\mathbf{w}^T\mathbf{w}}$ and in turn $\mathbf{d}=\frac{w_0+\mathbf{w}^T\mathbf{x}}{\mathbf{w}^T\mathbf{w}}\mathbf{w}$
<br>
* the total length of the distance $\mathbf{d}$ between the point and the plane is the Euclidean norm $||\mathbf{d}||$, which is equal to
<br>
$$||\mathbf{d}|| = \sqrt{\mathbf{d}^T\mathbf{d}} = |c|\cdot\sqrt{\mathbf{w}^T\mathbf{w}} =
\frac{|w_0+\mathbf{w}^T\mathbf{x}|}{\mathbf{w}^T\mathbf{w}}\sqrt{\mathbf{w}^T\mathbf{w}} =
\frac{|w_0+\mathbf{w}^T\mathbf{x}|}{||\mathbf{w}||} $$
* across all the training data points $\mathbf{x}_1,...,\mathbf{x}_n$, we are interested in finding those points that minimize such distance for a given value of $w_0$ and $\mathbf{w}$, and therefore we are interested in the following minimization problem: <br> $$ \delta(w_0,\mathbf{w}) = \min_{\mathbf{x}_{1},...,\mathbf{x}_n} \frac{|w_0+\mathbf{w}^T\mathbf{x}_i|}{||\mathbf{w}||} $$ <br> where $\delta(w_0,\mathbf{w})$ is the *margin* of the hyperplane defined by $w_0$ and $\mathbf{w}$
* now, across all $w_0$ in $\mathbb{R}$ and $\mathbf{w}$ in $\mathbb{R}^p$, the final goal is to find the optimum values $w_0^{opt}$ and $\mathbf{w}^{opt}$ that lead to a hyperplane which *maximizes* the margins (i.e., which maximizes the distances from the training data points): <br> $$w_0^{opt},\mathbf{w}^{opt} = \max_{w_0,\mathbf{w}} \delta(w_0,\mathbf{w}) = \max_{w_0,\mathbf{w}}\left[\min_{\mathbf{x}_{1},...,\mathbf{x}_n} \frac{|w_0+\mathbf{w}^T\mathbf{x}_i|}{||\mathbf{w}||}\right] $$
<br>
* the problem is incomplete: we are not specifying that the optimal hyperplane must lie in between the two classes (and therefore a solution such as $-\infty$ or $\infty$ is the optimal one at the moment); to add this requirement, we must *constrain* the optimization problem. The constrain must state that the observations all lie in the right side of the hyperplane, which is equivalent to say that $y_i(w_0+\mathbf{w}^T\mathbf{x}_i) \geq 0\ \forall i$ (check this result by yourself, with the help of the plot above). The new optimization problem becomes <br><br>$$\max_{w_0,\mathbf{w}}\left[\min_{\mathbf{x}_{1},...,\mathbf{x}_n} \frac{|w_0+\mathbf{w}^T\mathbf{x}_i|}{||\mathbf{w}||}\right]$$<br>$$\ subject\ to\ y_i(w_0+\mathbf{w}^T\mathbf{x}_i) \geq 0\ \forall i$$
* it is possible to simplify the problem:
1. in the solution for the margins, $||\mathbf{w}||$ is constant for the $\mathbf{x}_i$'s and can be pulled outside the minimization problem:<br>$$\max_{w_0,\mathbf{w}}\frac{1}{||\mathbf{w}||}\left[\min_{\mathbf{x}_{1},...,\mathbf{x}_n} |w_0+\mathbf{w}^T\mathbf{x}_i|\right]$$<br>$$\ s.t.\ y_i(w_0+\mathbf{w}^T\mathbf{x}_i) \geq 0\ \forall i$$
<br>
1. the hyperplane and the margins are "scale-invariant": $\mathcal{H}_{w_0,\mathbf{w}} = \mathcal{H}_{k\cdot w_0,k\cdot\mathbf{w}}$ and $\delta(w_0,\mathbf{w}) = \delta(k\cdot w_0,k\cdot\mathbf{w})$ for any $k \neq 0$: we can obtain different solutions for $\mathbf{w}$ and $w_0$, but the hyperplane is the same; we can use this fact to choose a scale for the bias and the coefficients such that <br> $$\min_{\mathbf{x}_{1},...,\mathbf{x}_n} |w_0+\mathbf{w}^T\mathbf{x}_i| = 1$$ <br> and the new optimization problem can be reformulated as <br>$$\max_{w_0,\mathbf{w}}\frac{1}{||\mathbf{w}||}$$<br>$$\ s.t.\
\left\{
\begin{array}{l}
y_i(w_0+\mathbf{w}^T\mathbf{x}_i) \geq 0\ \forall i \\
\min_{\mathbf{x}_{1},...,\mathbf{x}_n} |w_0+\mathbf{w}^T\mathbf{x}_i| = 1\ \forall i
\end{array}
\right.
$$<br> Notice that this is what we observed on the left plot above: the margin is now "fixed" to have a length of $\frac{1}{||\mathbf{w}||}$, and our goal is to find the values of $w_0$ and $\mathbf{w}$ that maximize such length
1. the maximization problem can be easily reformulated as a minimization problem:<br>$$\min_{w_0,\mathbf{w}}\mathbf{w}^T\mathbf{w}$$<br>$$\ s.t.\
\left\{
\begin{array}{l}
y_i(w_0+\mathbf{w}^T\mathbf{x}_i) \geq 0\ \forall i \\
\min_{\mathbf{x}_{1},...,\mathbf{x}_n} |w_0+\mathbf{w}^T\mathbf{x}_i| = 1\ \forall i
\end{array}
\right.
$$<br>
1. last, the constraints can be also simplified; (a) $y_i(w_0+\mathbf{w}^T\mathbf{x}_i) \geq 0$ can be joined to (b) $\min_{\mathbf{x}_{1},...,\mathbf{x}_n} |w_0+\mathbf{w}^T\mathbf{x}_i| = 1$, implying that <br> (c) $\forall\ i,\ y_i(w_0+\mathbf{w}^T\mathbf{x}_i) \geq 1$ (this is true, as all points are on the correct side of the hyperplane, the absolute values can be replaced by the multiplication by $y_i$, and their minimum is equal to 1); but the opposite is also true, that is, knowing (c) implies (a) and (b): if $y_i(w_0+\mathbf{w}^T\mathbf{x}_i)$ was larger than one, we can still minimize (rescale) $\mathbf{w}$ and $w_0$ s.t. the minimum becomes one, making (a) and (b) automatically true
Thus, the final optimization problem is:
$$\min_{w_0,\mathbf{w}}\mathbf{w}^T\mathbf{w}$$<br>$$\ s.t.\ y_i(w_0+\mathbf{w}^T\mathbf{x}_i) \geq 1\ \forall i$$
<br>
Notice that the *support vectors* are those data points for which the constraint $y_i(w_0+\mathbf{w}^T\mathbf{x}_i)$ is equal to 1. This problem is a Quadratic Programming (QP) otpimization problem, where the objective function to be minimized is quadratic ($\mathbf{w}^T\mathbf{w}$), and the constraint is linear: this implies that a unique solution exists. There is a number of libraries and softwares that can help to find solutions to QP problems; scikit-learn, for example, uses [liblinear](https://www.csie.ntu.edu.tw/~cjlin/liblinear/). How such libraries work is beyond the scope of this course; it is important, though, to be aware of the kind of optimization problem SVM is solving.
### 1.3 Decision function, predictions, and comparison with Logistic Regression
SVM shares some similarities, while having some profound differences, with the logistic regression (LR) model:
* similar to LR, the decision function of linear SVM $f(\mathbf{x})$ is the linear combination of the features (with the weights), $w_0+\mathbf{w}^T\mathbf{x}_i$; as a matter of fact, the two models often lead to similar results
* similar to LR, a new instance $\mathbf{x}^*$ is predicted to the positive class if $f(\mathbf{x}^*) \geq 0$ and to the negative class otherwise
* unlike LR, SVM only needs the support vectors to define the optimal separating hyperplane
* unlike LR, SVM doesn't have a probabilistic interpreation, but only a geometrical one; therefore, it doesn't return probabilities, but only the values of the decision function (quantities such as ROC curve and P/R curves can be computed using the decision function, and setting $0$ as the 'default threshold')
In the plot below, you can see the decision boundaries found by (unregularized) LR and (hard-margins) SVM for the dataset introduced above. As you can notice, the two algorithms lead to very similar boundaries.
<img src="./img/linear_svm/svm_vs_lr.png" width="350" height="50"/>
## 2. SVM for Non-Linearly Separable Data
What if a perfect linear separation was not possible? Consider the following dataset.
<img src="./img/linear_svm/non_lin_sep_svm.png" width="350" height="50"/>
In this case, because of that one positive point in the region of the negative class, there is no straight line that can perfectly separate the classes. In this case, Logistic Regression can still find a solution, while SVM with hard margins won't find any solution: its optimization problem becomes *infeasible*.
In order to find a solution to such problem, the hard-margins requirement must be relaxed with the introduction of *soft margins*. These margins can be found with the introduction of *slack variables*.
### 2.1 The slack variables
In the hard-margins problem, the constraint
$$y_i(w_0+\mathbf{w}^T\mathbf{x}_i) \geq 1\ \forall i$$
doesn't allow for miss-classification of the training points: each units needs to stay in the correct side of the hyperplane. By introducing *slack variables* $\xi_i \geq 0\ \forall\ i$, we can allow for violations of such constraint; this can be done by subracting the value of the slack variables from the right-hand side of the constraint:
$$y_i(w_0+\mathbf{w}^T\mathbf{x}_i) \geq 1 - \xi_i\ \forall i$$
In particular, the new constraint can be interpreted as follows:
* points that lie exactly on the margins are all those points $i$ such that $\xi_i=0$ and the equality holds: that is, as in the hard-margin case, they are the units for which $y_i(w_0+\mathbf{w}^T\mathbf{x}_i) = 1$
* all the points that are on the right side of the hyperplane (i.e., the points that are correctly classified) and *outside the margin* have $\xi_i=0$ and the inequality holds: as in the hard-margin case, for these units ) $y_i(w_0+\mathbf{w}^T\mathbf{x}_i) > 1$
* all the points on the right side of the hyperplane, but *inside the margin*, have $0<\xi_i\leq1$, and therefore $1> y_i(w_0+\mathbf{w}^T\mathbf{x}_i) \geq 0$
* all the points in the wrong side of the hyperplane (miss-classified observations) have $\xi_i\geq1$, which implies $y_i(w_0+\mathbf{w}^T\mathbf{x}_i) \leq 0$
The absolute value of the $\xi_i$'s depends on how much the unit violates the constraint, or it is inside the margin. Under this new formulation, the *support vectors* are all those units that define the margins, are inside of them, or that are in the wrong side of the hyperplane.
The following plots give a graphical intuition of how the slack variables work:
<img src="./img/linear_svm/svm_slack_variables.png" width="900" height="50"/>
### 2.2 The new optimization problem
It is clear that it is always possible to find values for the $\xi_i$'s such that the constraints $\{y_i(w_0+\mathbf{w}^T\mathbf{x}_i) \geq 1 - \xi_i,\ i=1,...,n\}$ are satisfied. However, as we want to relax as few violations as possible, we want the values of the $\xi_i$'s we subtract to be as little as possible. This means that we can add the sum of the $\xi_i$'s to the minimization problem, and associate a cost $C\geq0$ to this sum. The "soft-constraints" optimization problems thus becomes:
$$\min_{w_0,\mathbf{w}}\mathbf{w}^T\mathbf{w} + C\sum_{i=1}^n\xi_i$$
$$\ s.t.\
\left\{
\begin{array}{l}
y_i(w_0+\mathbf{w}^T\mathbf{x}_i) \geq 1-\xi_i\ \forall i \\
\xi_i \geq 0\ \forall i
\end{array}
\right.
$$
As we will see in the next section, $C$ is a hyperparameter that controls for model complexity. Here, we finish by recasting the optimization problem into one that can make the objective function differentiable, and therefore suitable for optimization techniques such as gradient descent.
In order to achieve this, we can easily get rid of the constraints by noticing that the following equality holds:
<br>
$$
\xi_i =
\left\{
\begin{array}{cl}
1 - y_i(w_0+\mathbf{w}^T\mathbf{x}_i) & \ if\ y_i(w_0+\mathbf{w}^T\mathbf{x}_i) < 1 \\
0 & \ if \ y_i(w_0+\mathbf{w}^T\mathbf{x}_i) \geq 1
\end{array}
\right.
$$
<br>
In practice, the slack variables are equal to 0 for all the training examples in the correct side of the hyperplane *and* outside the margins. To the extent that the training observations are inside the margins, or even in the wrong side of the hyperplane, the value of the corresponding $\xi_i$'s decreases accordingly. This means that, for each $i$ in the training set, we have that $$\xi_i = \max(1 - y_i(w_0+\mathbf{w}^T\mathbf{x}_i), 0)$$
The function $\max(1 - t, 0)$ is known as *hinge loss* in the Machine Learning and optimization communities. This new insight allows us to rewrite the whole minimization problem as follows:
$$\min_{w_0,\mathbf{w}}\left[\mathbf{w}^T\mathbf{w} + C\sum_{i=1}^n \max\left(1 - y_i(w_0+\mathbf{w}^T\mathbf{x}_i), 0\right)\right]$$
where the first term ($\mathbf{w}^T\mathbf{w}$) acts as regularizer, and the *hinge loss* seeks to find the best fitting solution for a given cost $C$ (Once again, notice the form Regularizer + Loss already encountered with logistic regression). The following plot shows the *hinge loss function* (and compares it with the log-loss used by logistic regression, as well the 0-1 missclassification loss):
<img src="./img/linear_svm/hinge_vs_log.png" width="450" height="50"/>
We can notice three facts:
* the log loss looks like a *smoother* version of the hinge-loss; for large negative values (that is: for large mistakes made by the classifiers), the two functions becomes approximatively parallel
* both the hinge loss and the log-loss try to approximate the misclassification loss, which is non-differentiable and non-convex
* the hinge loss is linear for values smaller than 1 (observations inside or on the wrong side of the margins), and constant (equal to 0) for larger values (observations correctly classified); this means that it can be easily differentiated *almost everywhere*, except at $x=1$ (in which case the gradient can be substituted with a *sub-gradient* value)
### 2.3 Regularization
As mentioned above, the hyperparameter $C$ can be used to regularize the model. In particular, $C$ can be interpreted as the *cost* we are willing to pay for the miss-classifications:
* if $C$ is small, the penalty is cheap and we allow more training units to be on the wrong side of the margin (or within the margins), obtaining larger margins
* if $C$ is large, the penalty is expensive and we prefer to have as more points as possible on the correct side of the hyperplane (and possibly outside the marings); in this way, margins become *smaller*
* as $C \rightarrow 0$, the penalty becomes "free" and it doesn't matter anymore
* as $C \rightarrow +\infty$, we seek to have 0 slacks and we move towards the *hard margins* solution
This can be easily seen in the following plots (notice the number of data points that are inside the margins as we increase $C$):
<img src="./img/linear_svm/svm_regularization.png" width="750" height="50"/>
Can you recognize for what values of $C$ the model has larger bias, and for what values of $C$ the model has larger variance? Therefore, when is it more likely underfitting, and when is it more likely overfitting?
## 3. SVM: Other Remarks
* Exactly like other linear models, Linear SVM works better in higher dimensions (it is also faster to train than the non-linear counterpart, which we will explore in the next lecture)
* As a downside, of course, it doesn't work well when the classes are divided by non-linear decision boundaries
* SVM doesn't have a probabilistic interpretation; it just seeks a hyperplane in the feature space that can separate the classes
* the coefficient weights $\mathbf{w}$ can be interpreted and used as *variable importance* indicators (therefore, it can also be used for feature selection)
* besides $l_2$ regularization, Linear SVM can also be trained with a $l_1$ penalty, in which case the weights vector becomes sparse (some of the elements of $\mathbf{w}$ are set to 0)
* unlike logistic regression, multiclass SVM cannot be trained with several classes simultaneously, therefore we must resort to approaches such as One-Versus-Rest for this type of classification
* it is very important to **rescale the dataset** before training a SVM classifier (either with standardization and/or normalization), otherwise features taking on larger values will dominate over other features when determining the distance from the hyperplane and performing predictions
* there exists a Regression version of the support vector classifier, which is called *Support Vector Regression*; it sometimes leads to results similar to linear regression, but it is not as intuitive, and therefore not used that often. We won't cover such algorithm in this course, but feel free to try the [scikit-learn implementation](https://scikit-learn.org/stable/modules/generated/sklearn.svm.SVR.html) of such algorithm with continuous output variables
## 4. Examples in Python
We are now going to explore the output of the `LinearSVC` function from the `sklearn.svm` module. In the first part we will see an example in the binary case; in the second, we will see an example with the multiclass case (OvR approach).
### 4.1 Two-Class Case
We will generate a toy dataset with the `make_classification` function of scikit-learn (we will use the same dataset generated in the notebook of Logistic Regression). Furthermore, we scale the training and test datasets by standardizing them.
```
from sklearn.datasets import make_classification
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
X, y = make_classification(n_samples=200, n_features=2, n_redundant=0,
n_informative=2, n_clusters_per_class=1,
flip_y=0.05, class_sep=0.8, random_state=60)
X_train, X_test, y_train, y_test = train_test_split(X, y, stratify=y, test_size=0.2,
random_state=1)
scaler = StandardScaler()
X_train_scaled = scaler.fit_transform(X_train)
X_test_scaled = scaler.transform(X_test)
plt.figure(figsize=(5,5))
plt.plot(X_train_scaled[y_train==0,0],X_train_scaled[y_train==0,1], "bo", label="Class 0" )
plt.plot(X_train_scaled[y_train==1,0],X_train_scaled[y_train==1,1], "o", color="orange", label="Class 1" )
plt.xlabel(r"$x_1$", fontsize=12)
plt.ylabel(r"$x_2$", fontsize=12)
plt.legend()
plt.show()
```
We will show an application of `LinearSVC` with `C=0.5`; notice that this is not necessarily the best value, which should be found by Cross-Validation or with the Holdout Method (By default, LinearSVC has the options `penalty='l2'` and `C=1.0`. More information in the [documentation](https://scikit-learn.org/stable/modules/generated/sklearn.svm.LinearSVC.html)).
```
from sklearn.svm import LinearSVC
svm_mod = LinearSVC(C=0.5, random_state=1)
svm_mod.fit(X_train_scaled, y_train)
```
As done with the logistic regression model, we can explore the coefficients vector $\mathbf{w}$:
```
svm_mod.coef_
```
and the bias term $w_0$:
```
svm_mod.intercept_
```
We can also calculate the decision function $w_0+\mathbf{w}^T\mathbf{x}$; let's do it for the first five units of the test set:
```
svm_mod.decision_function(X_test_scaled[:5])
```
which suggests us that the first, second, and fifth unit of the test set are predicted to belong to the positive class, while the third and fourth unit to the negative class. Let's confirm our intuition with the `predict` method:
```
svm_mod.predict(X_test_scaled[:5])
```
Last, let's plot the data for the found decision boundary and its margins (remember: the *decision boundary* are all those points in the feature space for which the decision function is equal to 0, and the margins all those values for which the decision function is equal to 1 or -1).
```
x1_range = np.linspace(X_train_scaled[:,0].min()-0.1, X_train_scaled[:,0].max()+0.1, 100)
x2_range = np.linspace(X_train_scaled[:,1].min()-0.1, X_train_scaled[:,1].max()+0.1, 100)
xx1, xx2 = np.meshgrid(x1_range, x2_range)
df_grid = svm_mod.decision_function(np.c_[xx1.ravel(), xx2.ravel()])
df_grid = df_grid.reshape(xx1.shape)
f, ax = plt.subplots(figsize=(5, 5))
contour = ax.contour(xx1, xx2, df_grid, [-1,0,1],
alpha=1., vmin=0, vmax=0)
plt.clabel(contour, inline=1, fontsize=12)
plt.plot(X_train_scaled[y_train==0,0], X_train_scaled[y_train==0,1],
"bo", label="Class 0")
plt.plot(X_train_scaled[y_train==1,0], X_train_scaled[y_train==1,1],
"o", color="orange", label="Class 1")
plt.title("SVM: Boundary and Margins", fontsize=16)
plt.xlabel(r"$x_1$", fontsize=12)
plt.ylabel(r"$x_2$", fontsize=12)
plt.show()
```
### 4.2 Multiclass case
As done for the two-class case, we use `make_classification` to generate the same four-class case used with the Logistic Regression model. When more than two classes are chosen, `LinearSVC` selects the option `multi_class='ovr'` approach by default, and so there is no need for us to specify it manually.
```
X, y = make_classification(n_samples=240, n_features=2, n_redundant=0,
n_informative=2, n_clusters_per_class=1, n_classes=4,
flip_y=0.05, class_sep=1.2, random_state=2)
X_train, X_test, y_train, y_test = train_test_split(X, y, stratify=y, test_size=0.2,
random_state=1)
scaler = StandardScaler()
X_train_scaled = scaler.fit_transform(X_train)
X_test_scaled = scaler.transform(X_test)
plt.figure(figsize=(5,5))
plt.plot(X_train_scaled[y_train==0,0],X_train_scaled[y_train==0,1], "bo", label="Class 0" )
plt.plot(X_train_scaled[y_train==1,0],X_train_scaled[y_train==1,1], "o", color="orange", label="Class 1" )
plt.plot(X_train_scaled[y_train==2,0],X_train_scaled[y_train==2,1], "mo", label="Class 2" )
plt.plot(X_train_scaled[y_train==3,0],X_train_scaled[y_train==3,1], "go", label="Class 3" )
plt.xlabel(r"$x_1$", fontsize=12)
plt.ylabel(r"$x_2$", fontsize=12)
plt.legend()
plt.show()
```
As done above, let's train a Linear SVM algorithm with `C=0.5` and explore the estimated coefficients:
```
svm_mod = LinearSVC(C=0.5, random_state=1)
svm_mod.fit(X_train_scaled, y_train)
svm_mod.coef_
```
Exactly as seen for the logistic regression model, a set of coefficients is returned for each class (which corresponds to the coefficients estimated for each separate OvR model). The same can be said for the bias terms...
```
svm_mod.intercept_
```
...and for the decision functions (here calculated for the first five units of the test set):
```
svm_mod.decision_function(X_test_scaled[:5])
```
Thus, test units 1, 2, 3, 4, and 5 are predicted to belong to Classes 3, 0, 1, 1, and 2 respectively:
```
svm_mod.predict(X_test_scaled[:5])
```
Last, we are going to plot the class-specific estimated decision boundaries:
```
from matplotlib.colors import ListedColormap
custom_cmap = ListedColormap(['blue','orange','magenta', 'green'])
x0, x1 = np.meshgrid(
np.linspace(X_train_scaled[:,0].min()-0.1, X_train_scaled[:,0].max()+0.1, 500).reshape(-1, 1),
np.linspace(X_train_scaled[:,1].min()-0.1, X_train_scaled[:,1].max()+0.1, 500).reshape(-1, 1),
)
X_new = np.c_[x0.ravel(), x1.ravel()]
y_predict = svm_mod.predict(X_new)
zz = y_predict.reshape(x0.shape)
plt.figure(figsize=(5, 5))
plt.plot(X_train_scaled[y_train==3, 0], X_train_scaled[y_train==3, 1], "go", label="Class 3")
plt.plot(X_train_scaled[y_train==2, 0], X_train_scaled[y_train==2, 1], "mo", label="Class 2")
plt.plot(X_train_scaled[y_train==1, 0], X_train_scaled[y_train==1, 1], "o", color="orange", label="Class 1")
plt.plot(X_train_scaled[y_train==0, 0], X_train_scaled[y_train==0, 1], "bo", label="Class 0")
plt.contour(x0, x1, zz, cmap=custom_cmap, alpha=0.25)
plt.xlabel("x1")
plt.ylabel("x2")
plt.title("SVM: Decision Boundaries")
plt.legend()
plt.show()
```
Once again, you can see that the decision boundaries found by LinearSVM are not too different from the ones found by Logistic Regression! (Compare it with the graphs plotted in the Logistic Regression notebook.)
### 4.3 LinearSVM in Python, other remarks and exercises
* the `sklear.linear_model` module provides the [SGDclassifier](https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.SGDClassifier.html) function, which can estimate linear SVM with stochastic gradient descent when `loss='hinge'` is set
* the option `pnealty='l_1;` is also possible, in which case $l_1$ regularization is performed
* for both the binary and multiclass case, try to plot the decision boundaries (and margins in the binary case) for different values of C, and see how they change
### 5. Application to the Heart Dataset
As done for the logistic regression model, we apply also LinearSVM to the heart dataset. We will work with the $l_2$ penalty, and evaluate different values of `C` with 10-fold-cross-validation. As an exercise, try to see whether the $l_1$ penalty performs better than the $l_2$ penalty, and what features are selected with this penalty!
```
# 1.Load datasets
data_train = pd.read_csv("./data/heart_data/heart_train_processed.csv")
data_test = pd.read_csv("./data/heart_data/heart_test_processed.csv")
X_train = data_train.drop("y", axis=1)
X_test = data_test.drop("y", axis=1)
y_train = data_train["y"]
y_test = data_test["y"]
# 2a. Prepare parameter Grid for elastic-net
from sklearn.model_selection import GridSearchCV
# For C, we will use a grid of 120 equi-distant values in the log space between 1e-4 and 70
param_grid = {'C': np.exp( np.linspace(np.log(1e-4), np.log(70), 120)) }
# 2b. Run GridSearchCV with 10-fold-cv
clf = GridSearchCV(LinearSVC(max_iter=1e+07, random_state=1),
param_grid, cv=10, n_jobs=-1)
clf.fit(X_train, y_train)
print("CV Grid Search: Done")
```
Let's visualize the plot of C values vs. CV accuracies:
```
cv_res = pd.DataFrame(clf.cv_results_).sort_values(by="param_C")
plt.figure(figsize=(10,5))
x_ax = np.linspace(0, cv_res.shape[0], cv_res.shape[0])
plt.plot(x_ax, cv_res["mean_test_score"], "b-")
plt.plot(cv_res["mean_test_score"].idxmax()+1,clf.best_score_, "go", label="Best CV Estimator")
plt.xticks(x_ax, (cv_res["param_C"]).astype("float64").round(20), rotation=90, fontsize=6)
plt.xlabel("C")
plt.ylabel("CV Accuracy")
plt.title("SVM: l2 Regularization")
plt.legend(loc=2)
plt.show()
```
The best CV score is obtained for:
```
clf.best_params_
```
which obtains a CV accuracy of
```
clf.best_score_
```
which is in line with the CV accuracy observed with the elastic-net penalty for logistic regression. The final accuracy on the test set is:
```
clf.best_estimator_.score(X_test,y_test)
```
Which is only slightly worse than the test-set accuracy calculated with Logistic Regression. Last, let's see how each features contribute to the final classification of the linear SVM model. How would you interpret this results?
```
colors = ['r' if coef < 0 else 'b' for coef in clf.best_estimator_.coef_[0]]
plt.figure(figsize=(7,5))
plt.barh(range(X_train.shape[1]), clf.best_estimator_.coef_[0], color=colors)
plt.yticks(range(X_train.shape[1]), X_train.columns)
plt.xticks(np.arange(clf.best_estimator_.coef_[0].min(), clf.best_estimator_.coef_[0].max(),0.07))
plt.xlabel("$w_j$")
plt.title("SVM: Features Contribution")
plt.show()
```
| github_jupyter |
```
import meanderpy as mp
import matplotlib.pyplot as plt
import numpy as np
%matplotlib qt
cd /Users/zoltan/Dropbox/Python
```
## Input parameters
```
W = 200.0 # channel width (m)
D = 12.0 # channel depth (m)
pad = 100 # padding (number of nodepoints along centerline)
deltas = 50.0 # sampling distance along centerline
nit = 2000 # number of iterations
Cf = 0.022 # dimensionless Chezy friction factor
crdist = 1.5*W # threshold distance at which cutoffs occur
kl = 60.0/(365*24*60*60.0) # migration rate constant (m/s)
kv = 1.0E-11 # vertical slope-dependent erosion rate constant (m/s)
dt = 2*0.05*365*24*60*60.0 # time step (s)
dens = 1000 # density of water (kg/m3)
saved_ts = 20 # which time steps will be saved
n_bends = 30 # approximate number of bends you want to model
Sl = 0.0 # initial slope (matters more for submarine channels than rivers)
t1 = 500 # time step when incision starts
t2 = 700 # time step when lateral migration starts
t3 = 1400 # time step when aggradation starts
aggr_factor = 4e-9 # aggradation factor (m/s, about 0.18 m/year, it kicks in after t3)
```
## Initialize model
```
from imp import reload
reload(mp)
ch = mp.generate_initial_channel(W,D,Sl,deltas,pad,n_bends) # initialize channel
chb = mp.ChannelBelt(channels=[ch],cutoffs=[],cl_times=[0.0],cutoff_times=[]) # create channel belt object
```
## Run simulation
```
chb.migrate(nit,saved_ts,deltas,pad,crdist,Cf,kl,kv,dt,dens,t1,t2,t3,aggr_factor) # channel migration
fig = chb.plot('strat',20,60) # plotting
```
## Build 3D fluvial model
```
h_mud = 0.4 # thickness of overbank deposit for each time step
dx = 10.0 # gridcell size in meters
chb_3d, xmin, xmax, ymin, ymax = chb.build_3d_model('fluvial',h_mud=h_mud,levee_width=4000.0,h=12.0,w=W,bth=0.0,
dcr=10.0,dx=dx,delta_s=deltas,starttime=chb.cl_times[20],endtime=chb.cl_times[-1],
xmin=0,xmax=0,ymin=0,ymax=0)
# create plots
fig1,fig2,fig3 = chb_3d.plot_xsection(200, [[0.5,0.25,0],[0.9,0.9,0],[0.5,0.25,0]], 4)
```
## Build 3D submarine channel model
```
W = 200.0 # channel width (m)
D = 12.0 # channel depth (m)
pad = 50 # padding (number of nodepoints along centerline)
deltas = 100.0 # sampling distance along centerline
nit = 1500 # number of iterations
Cf = 0.02 # dimensionless Chezy friction factor
crdist = 1.5*W # threshold distance at which cutoffs occur
kl = 60.0/(365*24*60*60.0) # migration rate constant (m/s)
kv = 1.0E-11 # vertical slope-dependent erosion rate constant (m/s)
dt = 2*0.05*365*24*60*60.0 # time step (s)
dens = 1000 # density of water (kg/m3)
saved_ts = 20 # which time steps will be saved
n_bends = 50 # approximate number of bends you want to model
Sl = 0.01 # initial slope (matters more for submarine channels than rivers)
t1 = 500 # time step when incision starts
t2 = 700 # time step when lateral migration starts
t3 = 1000 # time step when aggradation starts
aggr_factor = 4.0 # aggradation factor (it kicks in after t3)
reload(mp)
ch = mp.generate_initial_channel(W,D,Sl,deltas,pad,n_bends) # initialize channel
chb = mp.ChannelBelt(channels=[ch],cutoffs=[],cl_times=[0.0],cutoff_times=[]) # create channel belt object
chb.migrate(nit,saved_ts,deltas,pad,crdist,Cf,kl,kv,dt,dens,t1,t2,t3,aggr_factor) # channel migration
fig = chb.plot('strat',20,60) # plotting
h_mud = 3.0*np.ones((len(chb.cl_times[20:]),))
dx = 15.0
chb_3d, xmin, xmax, ymin, ymax = chb.build_3d_model('submarine',h_mud=h_mud,levee_width=5000.0,h=12.0,w=W,bth=6.0,
dcr=7.0,dx=dx,delta_s=deltas,starttime=chb.cl_times[20],endtime=chb.cl_times[-1],
xmin=0,xmax=0,ymin=0,ymax=0)
fig1,fig2,fig3 = chb_3d.plot_xsection(400, [[0.5,0.25,0],[0.9,0.9,0],[0.5,0.25,0]], 10)
```
| github_jupyter |
# Spectral Clustering
In this notebook, we will use cuGraph to identify the cluster in a test graph using Spectral Clustering with both the (A) Balance Cut metric, and (B) the Modularity Maximization metric
Notebook Credits
* Original Authors: Bradley Rees and James Wyles
* Last Edit: 05/03/2019
RAPIDS Versions: 0.7.0
Test Hardware
* GP100 32G, CUDA 9,2
## Introduction
Spectral clustering uses the eigenvectors of a Laplacian of the input graph to find a given number of clusters which satisfy a given quality metric. Balanced Cut and Modularity Maximization are such quality metrics.
@See: https://en.wikipedia.org/wiki/Spectral_clustering
To perform spectral clustering using the balanced cut metric in cugraph use:
__cugraph.spectralBalancedCutClustering(G, num_clusters, num_eigen_vects)__
<br>or<br>
__cugraph.spectralModularityMaximizationClustering(G, num_clusters, num_eigen_vects)__
Input
* __G__: A cugraph.Graph object
* __num_clusters__: The number of clusters to find
* __num_eig__: (optional) The number of eigenvectors to use
Returns
* __df__: cudf.DataFrame with two names columns:
* df["vertex"]: The vertex id.
* df["cluster"]: The assigned partition.
## cuGraph 0.7 Notice
cuGraph version 0.7 has some limitations:
* Only Int32 Vertex ID are supported
* Only float (FP32) edge data is supported
* Vertex numbering is assumed to start at zero
These limitations are being addressed and will be fixed future versions.
These example notebooks will illustrate how to manipulate the data so that it comforms to the current limitations
----
### Test Data
We will be using the Zachary Karate club dataset
*W. W. Zachary, An information flow model for conflict and fission in small groups, Journal of
Anthropological Research 33, 452-473 (1977).*

Zachary used a min-cut flow model to partition the graph into two clusters, shown by the circles and squares. Zarchary wanted just two cluster based on a conflict that caused the Karate club to break into two separate clubs. Many social network clustering methods identify more that two social groups in the data.
```
# Import needed libraries
import cugraph
import cudf
import numpy as np
from collections import OrderedDict
```
### Read the CSV datafile using cuDF
```
# Test file
datafile='./data/karate-data.csv'
# Read the data file
cols = ["src", "dst"]
dtypes = OrderedDict([
("src", "int32"),
("dst", "int32")
])
gdf = cudf.read_csv(datafile, names=cols, delimiter='\t', dtype=list(dtypes.values()) )
```
### Adjusting the vertex ID
Let's adjust all the vertex IDs to be zero based. We are going to do this by adding two new columns with the adjusted IDs
```
gdf["src"] = gdf["src"] - 1
gdf["dst"] = gdf["dst"] - 1
# The algorithm requires that there are edge weights. In this case all the weights are being ste to 1
gdf["data"] = cudf.Series(np.ones(len(gdf), dtype=np.float32))
# Look at the first few data records - the output should be two colums src and dst
gdf.head().to_pandas()
# verify data type
gdf.dtypes
```
Everything looks good, we can now create a graph
```
# create a CuGraph
G = cugraph.Graph()
G.add_edge_list(gdf["src"], gdf["dst"], gdf["data"])
```
----
#### Define and print function, but adjust vertex ID so that they match the illustration
```
def print_cluster(_df, id):
_f = _df.query('cluster == @id')
part = []
for i in range(len(_f)):
part.append(_f['vertex'][i] + 1)
print(part)
```
----
#### Using Balanced Cut
```
# Call spectralBalancedCutClustering on the graph for 3 clusters
# using 3 eigenvectors:
bc_gdf = cugraph.spectralBalancedCutClustering(G, 3, num_eigen_vects=3)
# Check the edge cut score for the produced clustering
score = cugraph.analyzeClustering_edge_cut(G, 3, bc_gdf['cluster'])
score
# See which nodes are in cluster 0:
print_cluster(bc_gdf, 0)
# See which nodes are in cluster 1:
print_cluster(bc_gdf, 1)
# See which nodes are in cluster 2:
print_cluster(bc_gdf, 2)
```
----
#### Modularity Maximization
Let's now look at the clustering using the modularity maximization metric
```
# Call spectralModularityMaximizationClustering on the graph for 3 clusters
# using 3 eigenvectors:
mm_gdf = cugraph.spectralModularityMaximizationClustering(G, 3, num_eigen_vects=3)
# Check the modularity score for the produced clustering
score = cugraph.analyzeClustering_modularity(G, 3, mm_gdf['cluster'])
score
# See which nodes are in cluster 0:
print_cluster(mm_gdf, 0)
print_cluster(mm_gdf, 1)
print_cluster(mm_gdf, 2)
```
Notice that the two metrics produce different results
___
Copyright (c) 2019, NVIDIA CORPORATION.
Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0
Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
___
| github_jupyter |
# Build machine learning workflow to predict new data with Amazon SageMaker and AWS Step Functions
This script creates a Step Function state machine to preprocess the inference data and predict with the images in ECR.
## Import modules
```
import uuid
import boto3
import sagemaker
from sagemaker.amazon.amazon_estimator import get_image_uri
from sagemaker.s3 import S3Uploader
from sagemaker import get_execution_role
from sagemaker.sklearn.processing import SKLearnProcessor
from sagemaker.processing import Processor, ProcessingInput, ProcessingOutput
import stepfunctions
from stepfunctions.steps import (
Chain,
ProcessingStep,
TransformStep
)
from stepfunctions.inputs import ExecutionInput
from stepfunctions.workflow import Workflow
```
## Setup
Modify according to your configurations.
```
# Bucket name in S3
bucket = "hermione-sagemaker"
# Set session
region_name="us-east-1"
boto3.setup_default_session(region_name=region_name)
# Get user role
role = get_execution_role()
# Role to create and execute step functions
# paste the AmazonSageMaker-StepFunctionsWorkflowExecutionRole ARN
workflow_execution_role = ""
# SageMaker expects unique names for each job, model and endpoint.
# Otherwise, the execution will fail. The ExecutionInput creates
# dynamically names for each execution.
execution_input = ExecutionInput(
schema={
"PreprocessingJobName": str,
"TransformJobName": str
}
)
# Get AWS Account ID
account_number = boto3.client("sts").get_caller_identity()["Account"]
# Processor image name previous uploaded in ECR
image_name_processor = "hermione-processor"
# Inference image name previous uploaded in ECR
image_name_inference = "hermione-inference"
# Input and output paths to execute train and inference
paths = {
'expectations': f"s3://{bucket}/PREPROCESSING/EXPECTATIONS",
'preprocessing': f"s3://{bucket}/PREPROCESSING/PREPROCESSING",
'test_raw': f"s3://{bucket}/TEST_RAW",
'inference_processed': f"s3://{bucket}/PREPROCESSING/INFERENCE_PROCESSED",
'validations': f"s3://{bucket}/PREPROCESSING/VALIDATIONS",
'model': f"s3://{bucket}/PREPROCESSING/MODEL/Hermione-train-2021-05-26-12-41-29-505/output/model.tar.gz",
'output_path': f"s3://{bucket}/PREPROCESSING/OUTPUT"
}
# instance to run the code
instance_type_preprocessing="ml.t3.medium"
instance_type_inference="ml.m5.large"
```
## Preprocessing Step
```
# Processor image previous uploaded in ECR
image_uri_processor = f"{account_number}.dkr.ecr.{region_name}.amazonaws.com/{image_name_processor}"
# Creates the processor to access the ECR image
processor = Processor(image_uri=image_uri_processor,
role=role,
instance_count=1,
instance_type=instance_type_preprocessing)
# Creates input and output objects for ProcessingStep
inputs=[
ProcessingInput(source=paths['test_raw'],
destination='/opt/ml/processing/input/raw_data',
input_name='raw_data'),
ProcessingInput(source=paths['preprocessing'],
destination='/opt/ml/processing/input/preprocessing',
input_name='preprocessing'),
ProcessingInput(source=paths['expectations'],
destination='/opt/ml/processing/input/expectations',
input_name='expectations')
]
outputs = [
ProcessingOutput(
source="/opt/ml/processing/output/processed/inference",
destination=paths['inference_processed'],
output_name="inference_data",
),
ProcessingOutput(
source="/opt/ml/processing/output/validations",
destination=paths['validations'],
output_name="validations",
)
]
# Creates the ProcessingStep
processing_step = ProcessingStep(
"SageMaker Preprocessing step",
processor=processor,
job_name=execution_input["PreprocessingJobName"],
inputs=inputs,
outputs=outputs,
container_arguments=["--step", "test"]
)
```
## Inference Step
```
# Inference image previous uploaded in ECR
image_uri_inference = f"{account_number}.dkr.ecr.{region_name}.amazonaws.com/{image_name_inference}"
# Creates input and output objects for TransformStep
input_path = paths['inference_processed']
model_path = paths['model']
output_path = paths['output_path']
# Creates the model to access the ECR image
model = sagemaker.model.Model(
image_uri = image_uri_inference,
model_data=model_path,
role=role)
# Creates a transformer object from the trained model
transformer = model.transformer(
instance_count=1,
instance_type=instance_type_inference,
output_path=output_path,
accept = 'text/csv')
# Creates the TransformStep
transform_step = TransformStep(
"Inference Step",
transformer=transformer,
job_name=execution_input["TransformJobName"],
data=input_path,
content_type='text/csv',
wait_for_completion=True,
model_name=model.name
)
```
## Create Workflow and Execute
```
# Creates Fail state to mark the workflow failed in case any of the steps fail.
failed_state_sagemaker_processing_failure = stepfunctions.steps.states.Fail(
"ML Workflow failed", cause="SageMakerProcessingJobFailed"
)
# Adds the Error handling in the workflow
catch_state_processing = stepfunctions.steps.states.Catch(
error_equals=["States.TaskFailed"],
next_step=failed_state_sagemaker_processing_failure,
)
processing_step.add_catch(catch_state_processing)
transform_step.add_catch(catch_state_processing)
# Creates workflow with Pre-Processing Job and Transform Job
workflow_graph = Chain([processing_step, transform_step])
branching_workflow = Workflow(
name="SFN_Hermione_Inference",
definition=workflow_graph,
role=workflow_execution_role,
)
branching_workflow.create()
# Generates unique names for Pre-Processing Job and Training Job
# Each job requires a unique name
preprocessing_job_name = "Hermione-Preprocessing-{}".format(
uuid.uuid1().hex
)
inference_job_name = "Hermione-Inference-{}".format(
uuid.uuid1().hex
)
# Executes the workflow
execution = branching_workflow.execute(
inputs={
"PreprocessingJobName": preprocessing_job_name,
"TransformJobName": inference_job_name
}
)
execution_output = execution.get_output(wait=False)
execution.render_progress()
```
## Results
```
import pandas as pd
pd.read_csv('s3://hermione-sagemaker/PREPROCESSING/OUTPUT/inference.csv.out')
```
| github_jupyter |
**Portrait Segmentation Using Prisma-Unet**
Set up the GPU runtime
```
# Check GPU
!nvidia-smi
# Mount G-drive
from google.colab import drive
drive.mount('/content/drive')
```
**Imports**
```
# Import libraries
import os
import tensorflow as tf
import keras
from keras.models import Model
from keras.layers import Dense, Input,Flatten, concatenate,Reshape, Conv2D, MaxPooling2D, Lambda,Activation,Conv2DTranspose, SeparableConv2D
from keras.layers import UpSampling2D, Conv2DTranspose, BatchNormalization, Dropout, DepthwiseConv2D, Add
from keras.callbacks import TensorBoard, ModelCheckpoint, Callback, ReduceLROnPlateau
from keras.regularizers import l1
from keras.optimizers import SGD, Adam
import keras.backend as K
from keras.utils import plot_model
from keras.callbacks import TensorBoard, ModelCheckpoint, Callback
import numpy as np
import matplotlib.pyplot as plt
from scipy.ndimage.filters import gaussian_filter
from random import randint
from keras.models import load_model
from keras.preprocessing.image import ImageDataGenerator
from PIL import Image
import matplotlib.pyplot as plt
from random import randint
%matplotlib inline
```
**Load dataset**
Load the datset for training the model from directory.
Ensure the images are in **RGB** format and masks (**ALPHA**) have pixel values **0 or 255**.
```
IMGDS="/content/portrait256/images";
MSKDS="/content/portrait256/masks";
# Total number of images
num_images=len(os.listdir(IMGDS+"/img"))
```
Copy pretrained model to local runtime disk. Save the checkpoints to your google drive (safe).
```
# Configure save paths and batch size
CHECKPOINT="/content/drive/My Drive/portrait256/prisma-net-{epoch:02d}-{val_loss:.2f}.hdf5"
LOGS='./logs'
BATCH_SIZE=64
```
**Data Generator**
Create a data generator to load images and masks together at runtime.
Use same seed for performing run-time augmentation for images and masks. Here we use 80/20 tran-val split.
**Note:** The keras 'flow_from_directory' expects a specific directory structure for loading datasets. Your parent data-set directory should contain two sub-directories 'images' and 'masks'. Now, each of these directories should have a sub-directory(say 'img' and 'msk) for storing images or masks.
```
# Data generator for training and validation
data_gen_args = dict(rescale=1./255,
width_shift_range=0.2,
height_shift_range=0.2,
zoom_range=0.2,
horizontal_flip=True,
validation_split=0.2
)
image_datagen = ImageDataGenerator(**data_gen_args)
mask_datagen = ImageDataGenerator(**data_gen_args)
# Provide the same seed and keyword arguments to the fit and flow methods
seed = 1
batch_sz=BATCH_SIZE
# Train-val split (80-20)
num_train=int(num_images*0.8)
num_val=int(num_images*0.2)
train_image_generator = image_datagen.flow_from_directory(
IMGDS,
batch_size=batch_sz,
shuffle=True,
subset='training',
color_mode="rgb",
class_mode=None,
seed=seed)
train_mask_generator = mask_datagen.flow_from_directory(
MSKDS,
batch_size=batch_sz,
shuffle=True,
subset='training',
color_mode="grayscale",
class_mode=None,
seed=seed)
val_image_generator = image_datagen.flow_from_directory(
IMGDS,
batch_size = batch_sz,
shuffle=True,
subset='validation',
color_mode="rgb",
class_mode=None,
seed=seed)
val_mask_generator = mask_datagen.flow_from_directory(
MSKDS,
batch_size = batch_sz,
shuffle=True,
subset='validation',
color_mode="grayscale",
class_mode=None,
seed=seed)
# combine generators into one which yields image and masks
train_generator = zip(train_image_generator, train_mask_generator)
val_generator = zip(val_image_generator, val_mask_generator)
```
**Model Architecture**
The prisma-net basically uses a **U-Net** encoder-decoder structure. However, the architecture incorporates a few significant **changes**. Firstly, we replace the concatenation of features after upsampling with **element-wise addition**. Further, instead of normal Conv+ReLu block, we use a **residual block with depth-wise separable convolutions**. Finally, to improve the accuracy the **decoder** part contains **more blocks** than encoder.
```
def residual_block(x, nfilters):
y = Activation("relu")(x)
y= SeparableConv2D(filters=nfilters, kernel_size=3, padding="same")(y)
y = Activation("relu")(y)
y= SeparableConv2D(filters=nfilters, kernel_size=3, padding="same")(y)
z = Add()([x, y])
return z
def prisma_unet(finetuene=False, alpha=1):
input = Input(shape=(256,256,3))
# Encoder part
x = Conv2D(filters=8, kernel_size=3,padding = 'same' )(input)
res1= residual_block(x, nfilters=8)
x = Conv2D(filters=32, kernel_size=3, strides=2, padding = 'same' )(res1)
res2= residual_block(x, nfilters=32)
x = Conv2D(filters=64, kernel_size=3, strides=2, padding = 'same' )(res2)
res3= residual_block(x, nfilters=64)
x = Conv2D(filters=128, kernel_size=3, strides=2, padding = 'same' )(res3)
x= residual_block(x, nfilters=128)
res4= residual_block(x, nfilters=128)
x = Conv2D(filters=128, kernel_size=3, strides=2, padding = 'same' )(res4)
x= residual_block(x, nfilters=128)
x= residual_block(x, nfilters=128)
x= residual_block(x, nfilters=128)
x= residual_block(x, nfilters=128)
x= residual_block(x, nfilters=128)
x= residual_block(x, nfilters=128)
# Decoder part
x=Conv2DTranspose(filters=128, kernel_size=3, strides=2, padding = "same")(x)
x = Add()([x, res4 ])
x= residual_block(x, nfilters=128)
x= residual_block(x, nfilters=128)
x= residual_block(x, nfilters=128)
x = Conv2DTranspose(filters=64, kernel_size=3, strides=2, padding = 'same' )(x)
x = Add()([x, res3 ])
x= residual_block(x, nfilters=64)
x= residual_block(x, nfilters=64)
x= residual_block(x, nfilters=64)
x = Conv2DTranspose(filters=32, kernel_size=3, strides=2, padding = 'same' )(x)
x = Add()([x, res2 ])
x= residual_block(x, nfilters=32)
x= residual_block(x, nfilters=32)
x= residual_block(x, nfilters=32)
x = Conv2DTranspose(filters=8, kernel_size=3, strides=2, padding = 'same' )(x)
x = Add()([x, res1 ])
x= residual_block(x, nfilters=8)
x= residual_block(x, nfilters=8)
x= residual_block(x, nfilters=8)
x = Conv2DTranspose(1, (1,1), padding='same')(x)
x = Activation('sigmoid', name="op")(x)
model = Model(inputs=input, outputs=x)
model.compile(loss='binary_crossentropy', optimizer=Adam(lr=1e-3),metrics=['accuracy'])
return model
# Get prisma network
model = prisma_unet()
# Model summary
model.summary()
# Layer specifications
for i, layer in enumerate(model.layers):
print(i, layer.output.name, layer.output.shape)
# Plot model architecture
plot_model(model, to_file='prisma-net.png')
# Save checkpoints
checkpoint = ModelCheckpoint(CHECKPOINT, monitor='val_loss', verbose=1, save_weights_only=False , save_best_only=True, mode='min')
# Callbacks
reduce_lr = ReduceLROnPlateau(factor=0.5, patience=3, min_lr=0.000001, verbose=1)
tensorboard = TensorBoard(log_dir=LOGS, histogram_freq=0,
write_graph=True, write_images=True)
callbacks_list = [checkpoint, tensorboard, reduce_lr]
```
**Train**
Initially train the model for **300 epochs** using **supervisely person** dataset. Finally, train the model on **portrait datasets**, using the result of the previous step as initial values for **weights**.
Use keras callbacks for **tensorboard** visulaization and **learning rate decay** as shown below. You can resume your training from a previous session by loading the entire **pretrained model** (weights & optimzer state) as a hdf5 file.
```
# Load pretrained model (if any)
model=load_model('/content/drive/My Drive/portrait256/prisma-net-07-0.09.hdf5')
# Train the model
model.fit_generator(
train_generator,
epochs=300,
steps_per_epoch=num_train/batch_sz,
validation_data=val_generator,
validation_steps=num_val/batch_sz,
use_multiprocessing=True,
workers=4,
callbacks=callbacks_list)
```
**Test**
Test the model on a new portrait image and plot the results.
```
# Load a test image
im=Image.open('/content/baby.jpg')
# Load the model
model=load_model('/content/drive/My Drive/portrait256/prisma-net-15-0.08.hdf5')
# Inference
im=im.resize((256,256),Image.ANTIALIAS)
img=np.float32(np.array(im)/255.0)
plt.imshow(img[:,:,0:3])
img=img[:,:,0:3]
# Reshape input and threshold output
out=model.predict(img.reshape(1,256,256,3))
out=np.float32((out>0.5))
# Output mask
plt.imshow(np.squeeze(out.reshape((256,256))))
```
**Export Model**
Export the model to **tflite** format for **real-time** inference on a **smart-phone**.
```
# Flatten output and save model
output = model.output
newout=Reshape((65536,))(output)
new_model=Model(model.input,newout)
new_model.save('prisma-net.h5')
# For Float32 Model
converter = tf.lite.TFLiteConverter.from_keras_model_file('/content/prisma-net.h5')
tflite_model = converter.convert()
open("prisma-net.tflite", "wb").write(tflite_model)
```
**Post-training Quantization**
We can **reduce the model size and latency** by performing post training quantization. Fixed precison conversion (**UINT8**) allows us to reduce the model size significantly by quantizing the model weights.We can run this model on the mobile **CPU**. The **FP16** (experimental) conversion allows us to reduce the model size by half and the corresponding model can be run directly on mobile **GPU**.
```
# For UINT8 Quantization
converter = tf.lite.TFLiteConverter.from_keras_model_file('/content/prisma-net.h5')
converter.optimizations = [tf.lite.Optimize.OPTIMIZE_FOR_SIZE]
tflite_model = converter.convert()
open("prisma-net_uint8.tflite", "wb").write(tflite_model)
# For Float16 Quantization (Experimental)
import tensorflow as tf
converter = tf.lite.TFLiteConverter.from_keras_model_file('/content/prisma-net.h5')
converter.optimizations = [tf.lite.Optimize.DEFAULT]
converter.target_spec.supported_types = [tf.lite.constants.FLOAT16]
tflite_model = converter.convert()
open("prisma-net_fp16.tflite", "wb").write(tflite_model)
```
**Plot sample output**
Load the test data as a batch using a numpy array.
Crop the image using the output mask and plot the result.
```
# Load test images and model
model=load_model('/content/prisma-net.h5',compile=False)
test_imgs=np.load('/content/kids.npy')
test_imgs= np.float32(np.array(test_imgs)/255.0)
# Perform batch prediction
out=model.predict(test_imgs)
out=np.float32((out>0.5))
out=out.reshape((4,256,256,1))
# Plot the output using matplotlib
fig=plt.figure(figsize=(16, 16))
columns = 4
rows = 2
for i in range(1, columns+1):
img = test_imgs[i-1].squeeze()
fig.add_subplot(rows, columns, i)
plt.imshow(img)
plt.show()
fig=plt.figure(figsize=(16, 16))
columns = 4
rows = 2
for i in range(1, columns+1):
img = out[i-1].squeeze()/255.0
fig.add_subplot(rows, columns, 4+i)
plt.imshow(out[i-1]*test_imgs[i-1])
plt.show()
```
| github_jupyter |
<center>
<img src="https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-PY0101EN-SkillsNetwork/IDSNlogo.png" width="300" alt="cognitiveclass.ai logo" />
</center>
# Watson Speech to Text Translator
Estimated time needed: **25** minutes
## Objectives
After completing this lab you will be able to:
* Operate a Speech to Text Translator through an API
### Introduction
<p>In this notebook, you will learn to convert an audio file of an English speaker to text using a Speech to Text API. Then, you will translate the English version to a Spanish version using a Language Translator API. <b>Note:</b> You must obtain the API keys and endpoints to complete the lab.</p>
<div class="alert alert-block alert-info" style="margin-top: 20px">
<h2>Table of Contents</h2>
<ul>
<li><a href="#ref0">Speech To Text</a></li>
<li><a href="#ref1">Language Translator</a></li>
<li><a href="#ref2">Exercise</a></li>
</ul>
</div>
```
#you will need the following library
!pip install ibm_watson wget
```
<h2 id="ref0">Speech to Text</h2>
<p>First we import <code>SpeechToTextV1</code> from <code>ibm_watson</code>. For more information on the API, please click on this <a href="https://cloud.ibm.com/apidocs/speech-to-text?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDeveloperSkillsNetworkPY0101ENSkillsNetwork19487395-2021-01-01&code=python">link</a>.</p>
```
from ibm_watson import SpeechToTextV1
import json
from ibm_cloud_sdk_core.authenticators import IAMAuthenticator
```
<p>The service endpoint is based on the location of the service instance, we store the information in the variable URL. To find out which URL to use, view the service credentials and paste the url here.</p>
```
url_s2t = "https://api.eu-gb.speech-to-text.watson.cloud.ibm.com/instances/9893cab4-b9d3-4ff1-88cd-bacee5369596"
```
<p>You require an API key, and you can obtain the key on the <a href="https://cloud.ibm.com/resources?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDeveloperSkillsNetworkPY0101ENSkillsNetwork19487395-2021-01-01">Dashboard </a>.</p>
```
iam_apikey_s2t = "wyUxOK2-zTQNX2DBiRWeQuL0HaPh2uw-GJdRi5iVvy4T"
```
<p>You create a <a href="http://watson-developer-cloud.github.io/python-sdk/v0.25.0/apis/watson_developer_cloud.speech_to_text_v1.html?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDeveloperSkillsNetworkPY0101ENSkillsNetwork19487395-2021-01-01">Speech To Text Adapter object</a> the parameters are the endpoint and API key.</p>
```
authenticator = IAMAuthenticator(iam_apikey_s2t)
s2t = SpeechToTextV1(authenticator=authenticator)
s2t.set_service_url(url_s2t)
s2t
```
<p>Lets download the audio file that we will use to convert into text.</p>
```
!wget -O PolynomialRegressionandPipelines.mp3 https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-PY0101EN-SkillsNetwork/labs/Module%205/data/PolynomialRegressionandPipelines.mp3
```
<p>We have the path of the .wav file we would like to convert to text</p>
```
filename='PolynomialRegressionandPipelines.mp3'
```
<p>We create the file object <code>wav</code> with the wav file using <code>open</code>. We set the <code>mode</code> to "rb" , this is similar to read mode, but it ensures the file is in binary mode. We use the method <code>recognize</code> to return the recognized text. The parameter <code>audio</code> is the file object <code>wav</code>, the parameter <code>content_type</code> is the format of the audio file.</p>
```
with open(filename, mode="rb") as wav:
response = s2t.recognize(audio=wav, content_type='audio/mp3')
```
<p>The attribute result contains a dictionary that includes the translation:</p>
```
response.result
from pandas import json_normalize
json_normalize(response.result['results'],"alternatives")
response
```
<p>We can obtain the recognized text and assign it to the variable <code>recognized_text</code>:</p>
```
recognized_text=response.result['results'][0]["alternatives"][0]["transcript"]
type(recognized_text)
```
<h2 id="ref1">Language Translator</h2>
<p>First we import <code>LanguageTranslatorV3</code> from ibm_watson. For more information on the API click <a href="https://cloud.ibm.com/apidocs/speech-to-text?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDeveloperSkillsNetworkPY0101ENSkillsNetwork19487395-2021-01-01&code=python"> here</a></p>
```
from ibm_watson import LanguageTranslatorV3
```
<p>The service endpoint is based on the location of the service instance, we store the information in the variable URL. To find out which URL to use, view the service credentials.</p>
```
url_lt='https://api.us-south.language-translator.watson.cloud.ibm.com/instances/0a37d337-5c9d-4e0a-973d-015362c77bdf'
```
<p>You require an API key, and you can obtain the key on the <a href="https://cloud.ibm.com/resources?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDeveloperSkillsNetworkPY0101ENSkillsNetwork19487395-2021-01-01">Dashboard</a>.</p>
```
apikey_lt='0MOycL06bk_eKKZ6h4XEq8brm1p5_mZ-IYGVATXeN4Pu'
```
<p>API requests require a version parameter that takes a date in the format version=YYYY-MM-DD. This lab describes the current version of Language Translator, 2018-05-01</p>
```
version_lt='2018-05-01'
```
<p>we create a Language Translator object <code>language_translator</code>:</p>
```
authenticator = IAMAuthenticator(apikey_lt)
language_translator = LanguageTranslatorV3(version=version_lt,authenticator=authenticator)
language_translator.set_service_url(url_lt)
language_translator
```
<p>We can get a Lists the languages that the service can identify.
The method Returns the language code. For example English (en) to Spanis (es) and name of each language.</p>
```
from pandas import json_normalize
json_normalize(language_translator.list_identifiable_languages().get_result(), "languages")
```
<p>We can use the method <code>translate</code>. This will translate the text. The parameter text is the text, Model_id is the type of model we would like to use use we use list the language. In this case, we set it to 'en-es' or English to Spanish. We get a Detailed Response object translation_response</p>
```
translation_response = language_translator.translate(\
text=recognized_text, model_id='en-es')
translation_response
```
<p>The result is a dictionary.</p>
```
translation=translation_response.get_result()
translation
```
<p>We can obtain the actual translation as a string as follows:</p>
```
spanish_translation =translation['translations'][0]['translation']
spanish_translation
```
<p>We can translate back to English</p>
```
translation_new = language_translator.translate(text=spanish_translation ,model_id='es-en').get_result()
```
<p>We can obtain the actual translation as a string as follows:</p>
```
translation_eng=translation_new['translations'][0]['translation']
translation_eng
```
<br>
<h2>Quiz</h2>
Translate to French.
```
French_translation=language_translator.translate(
text=translation_eng , model_id='en-fr').get_result()
```
<details><summary>Click here for the solution</summary>
```python
French_translation=language_translator.translate(
text=translation_eng , model_id='en-fr').get_result()
French_translation['translations'][0]['translation']
```
</details>
<h3>Language Translator</h3>
<b>References</b>
https://cloud.ibm.com/apidocs/speech-to-text?code=python
https://cloud.ibm.com/apidocs/language-translator?code=python
<hr>
## Authors:
[Joseph Santarcangelo](https://www.linkedin.com/in/joseph-s-50398b136/?utm_medium=Exinfluencer\&utm_source=Exinfluencer\&utm_content=000026UJ\&utm_term=10006555\&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDeveloperSkillsNetworkPY0101ENSkillsNetwork19487395-2021-01-01)
Joseph Santarcangelo has a PhD in Electrical Engineering, his research focused on using machine learning, signal processing, and computer vision to determine how videos impact human cognition. Joseph has been working for IBM since he completed his PhD.
## Other Contributor(s)
<a href="https://www.linkedin.com/in/fanjiang0619/?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDeveloperSkillsNetworkPY0101ENSkillsNetwork19487395-2021-01-01">Fan Jiang</a>
## Change Log
| Date (YYYY-MM-DD) | Version | Changed By | Change Description |
|---|---|---|---|
| 2021-04-07 | 2.2 | Malika | Updated the libraries |
| 2021-01-05 | 2.1 | Malika | Added a library |
| 2020-08-26 | 2.0 | Lavanya | Moved lab to course repo in GitLab |
| | | | |
| | | | |
<hr/>
## <h3 align="center"> © IBM Corporation 2020. All rights reserved. <h3/>
| github_jupyter |
# Multi-Layer Perceptron, MNIST
---
In this notebook, we will train an MLP to classify images from the [MNIST database](http://yann.lecun.com/exdb/mnist/) hand-written digit database.
The process will be broken down into the following steps:
>1. Load and visualize the data
2. Define a neural network
3. Train the model
4. Evaluate the performance of our trained model on a test dataset!
Before we begin, we have to import the necessary libraries for working with data and PyTorch.
```
# import libraries
import torch
import numpy as np
```
---
## Load and Visualize the [Data](http://pytorch.org/docs/stable/torchvision/datasets.html)
Downloading may take a few moments, and you should see your progress as the data is loading. You may also choose to change the `batch_size` if you want to load more data at a time.
This cell will create DataLoaders for each of our datasets.
```
from torchvision import datasets
import torchvision.transforms as transforms
# number of subprocesses to use for data loading
num_workers = 0
# how many samples per batch to load
batch_size = 20
# convert data to torch.FloatTensor
transform = transforms.ToTensor()
# choose the training and test datasets
train_data = datasets.MNIST(root='data', train=True,
download=True, transform=transform)
test_data = datasets.MNIST(root='data', train=False,
download=True, transform=transform)
# prepare data loaders
train_loader = torch.utils.data.DataLoader(train_data, batch_size=batch_size,
num_workers=num_workers)
test_loader = torch.utils.data.DataLoader(test_data, batch_size=batch_size,
num_workers=num_workers)
```
### Visualize a Batch of Training Data
The first step in a classification task is to take a look at the data, make sure it is loaded in correctly, then make any initial observations about patterns in that data.
```
import matplotlib.pyplot as plt
%matplotlib inline
# obtain one batch of training images
dataiter = iter(train_loader)
images, labels = dataiter.next()
images = images.numpy()
# plot the images in the batch, along with the corresponding labels
fig = plt.figure(figsize=(25, 4))
for idx in np.arange(20):
ax = fig.add_subplot(2, 20/2, idx+1, xticks=[], yticks=[])
ax.imshow(np.squeeze(images[idx]), cmap='gray')
# print out the correct label for each image
# .item() gets the value contained in a Tensor
ax.set_title(str(labels[idx].item()))
```
### View an Image in More Detail
```
img = np.squeeze(images[1])
fig = plt.figure(figsize = (12,12))
ax = fig.add_subplot(111)
ax.imshow(img, cmap='gray')
width, height = img.shape
thresh = img.max()/2.5
for x in range(width):
for y in range(height):
val = round(img[x][y],2) if img[x][y] !=0 else 0
ax.annotate(str(val), xy=(y,x),
horizontalalignment='center',
verticalalignment='center',
color='white' if img[x][y]<thresh else 'black')
```
---
## Define the Network [Architecture](http://pytorch.org/docs/stable/nn.html)
The architecture will be responsible for seeing as input a 784-dim Tensor of pixel values for each image, and producing a Tensor of length 10 (our number of classes) that indicates the class scores for an input image. This particular example uses two hidden layers and dropout to avoid overfitting.
```
import torch.nn as nn
import torch.nn.functional as F
## TODO: Define the NN architecture
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
# linear layer (784 -> 1 hidden node)
self.fc1 = nn.Linear(28 * 28, 256)
self.fc2 = nn.Linear(256, 64)
self.fc3 = nn.Linear(64, 10)
self.dropout = nn.Dropout(p = 0.2)
def forward(self, x):
# flatten image input
x = x.view(x.shape[0], -1)
# add hidden layer, with relu activation function
x = F.relu(self.fc1(x))
x = self.dropout(x)
x = F.relu(self.fc2(x))
x = self.dropout(x)
x = self.fc3(x)
return x
# initialize the NN
model = Net()
print(model)
```
### Specify [Loss Function](http://pytorch.org/docs/stable/nn.html#loss-functions) and [Optimizer](http://pytorch.org/docs/stable/optim.html)
It's recommended that you use cross-entropy loss for classification. If you look at the documentation (linked above), you can see that PyTorch's cross entropy function applies a softmax funtion to the output layer *and* then calculates the log loss.
```
## TODO: Specify loss and optimization functions
from torch import optim
# specify loss function
criterion = nn.CrossEntropyLoss()
# specify optimizer
optimizer = optim.Adam(model.parameters(), lr = 0.005)
```
---
## Train the Network
The steps for training/learning from a batch of data are described in the comments below:
1. Clear the gradients of all optimized variables
2. Forward pass: compute predicted outputs by passing inputs to the model
3. Calculate the loss
4. Backward pass: compute gradient of the loss with respect to model parameters
5. Perform a single optimization step (parameter update)
6. Update average training loss
The following loop trains for 30 epochs; feel free to change this number. For now, we suggest somewhere between 20-50 epochs. As you train, take a look at how the values for the training loss decrease over time. We want it to decrease while also avoiding overfitting the training data.
```
# number of epochs to train the model
n_epochs = 30 # suggest training between 20-50 epochs
model.train() # prep model for training
for epoch in range(n_epochs):
# monitor training loss
train_loss = 0.0
###################
# train the model #
###################
for data, target in train_loader:
# clear the gradients of all optimized variables
optimizer.zero_grad()
# forward pass: compute predicted outputs by passing inputs to the model
output = model(data)
# calculate the loss
loss = criterion(output, target)
# backward pass: compute gradient of the loss with respect to model parameters
loss.backward()
# perform a single optimization step (parameter update)
optimizer.step()
# update running training loss
train_loss += loss.item()*data.size(0)
# print training statistics
# calculate average loss over an epoch
train_loss = train_loss/len(train_loader.sampler)
print('Epoch: {} \tTraining Loss: {:.6f}'.format(
epoch+1,
train_loss
))
```
---
## Test the Trained Network
Finally, we test our best model on previously unseen **test data** and evaluate it's performance. Testing on unseen data is a good way to check that our model generalizes well. It may also be useful to be granular in this analysis and take a look at how this model performs on each class as well as looking at its overall loss and accuracy.
#### `model.eval()`
`model.eval(`) will set all the layers in your model to evaluation mode. This affects layers like dropout layers that turn "off" nodes during training with some probability, but should allow every node to be "on" for evaluation!
```
# initialize lists to monitor test loss and accuracy
test_loss = 0.0
class_correct = list(0. for i in range(10))
class_total = list(0. for i in range(10))
model.eval() # prep model for *evaluation*
for data, target in test_loader:
# forward pass: compute predicted outputs by passing inputs to the model
output = model(data)
# calculate the loss
loss = criterion(output, target)
# update test loss
test_loss += loss.item()*data.size(0)
# convert output probabilities to predicted class
_, pred = torch.max(output, 1)
# compare predictions to true label
correct = np.squeeze(pred.eq(target.data.view_as(pred)))
# calculate test accuracy for each object class
for i in range(len(target)):
label = target.data[i]
class_correct[label] += correct[i].item()
class_total[label] += 1
# calculate and print avg test loss
test_loss = test_loss/len(test_loader.sampler)
print('Test Loss: {:.6f}\n'.format(test_loss))
for i in range(10):
if class_total[i] > 0:
print('Test Accuracy of %5s: %2d%% (%2d/%2d)' % (
str(i), 100 * class_correct[i] / class_total[i],
np.sum(class_correct[i]), np.sum(class_total[i])))
else:
print('Test Accuracy of %5s: N/A (no training examples)' % (classes[i]))
print('\nTest Accuracy (Overall): %2d%% (%2d/%2d)' % (
100. * np.sum(class_correct) / np.sum(class_total),
np.sum(class_correct), np.sum(class_total)))
```
### Visualize Sample Test Results
This cell displays test images and their labels in this format: `predicted (ground-truth)`. The text will be green for accurately classified examples and red for incorrect predictions.
```
# obtain one batch of test images
dataiter = iter(test_loader)
images, labels = dataiter.next()
# get sample outputs
output = model(images)
# convert output probabilities to predicted class
_, preds = torch.max(output, 1)
# prep images for display
images = images.numpy()
# plot the images in the batch, along with predicted and true labels
fig = plt.figure(figsize=(25, 4))
for idx in np.arange(20):
ax = fig.add_subplot(2, 20/2, idx+1, xticks=[], yticks=[])
ax.imshow(np.squeeze(images[idx]), cmap='gray')
ax.set_title("{} ({})".format(str(preds[idx].item()), str(labels[idx].item())),
color=("green" if preds[idx]==labels[idx] else "red"))
```
| github_jupyter |
```
import matplotlib.pyplot as plt
import seaborn as sns
import pandas as pd
import json
import numpy as np
from acquire import remove_stopwords, basic_clean, tokenize , prep_and_split_data
from prepare_jag import prep_train, basic_clean3
import re
import os
from re import search
import tensorflow as tf
from sklearn.model_selection import train_test_split
from tensorflow.keras import layers
from tensorflow import keras
from nltk.corpus import stopwords
import nltk
import random
random.seed(333)
train_df, val_df, test_df = prep_and_split_data()
train_df
```
## Multi-label binarization
Let's preprocess our labels using the [StringLookup](https://keras.io/api/layers/preprocessing_layers/categorical/string_lookup) layer.
```
features = tf.ragged.constant(train_df['targets'].values)
lookup = tf.keras.layers.StringLookup(output_mode='multi_hot')
lookup.adapt(features)
vocab = lookup.get_vocabulary()
def invert_multi_hot(encoded_labels):
'''Reverse a single multi-hot encoded label to a tuple of vocab terms.'''
hot_indices = np.argwhere(encoded_labels == 1.0)[...,0]
return np.take(vocab, hot_indices)
print('Vocabulary:\n')
print(vocab)
```
### Separate the individual targets from the label pool and then use it to represent a given label set with 0's and 1's
```
sample_label = train_df['targets'].iloc[0]
print(f'Original label: {sample_label}')
label_binarized = lookup([sample_label])
print(f'Label-binarized representation: {label_binarized}')
```
## Data preprocessing and [tf.data.Dataset](https://www.tensorflow.org/api_docs/python/tf/data/Dataset) objects
```
train_df['clean'].apply(lambda x: len(x.split(" "))).describe()
max_seqlen = 98
batch_size = 128
padding_token = '<pad>'
auto = tf.data.AUTOTUNE
def make_dataset(dataframe, is_train=True):
labels = tf.ragged.constant(dataframe['targets'].values)
label_binarized = lookup(labels).numpy()
dataset = tf.data.Dataset.from_tensor_slices(
(dataframe['clean'].values, label_binarized)
)
dataset = dataset.shuffle(batch_size * 10) if is_train else dataset
return dataset.batch(batch_size)
```
## Prepare the [tf.data.Dataset](https://www.tensorflow.org/api_docs/python/tf/data/Dataset) objects.
```
train_dataset = make_dataset(train_df, is_train=True)
validation_dataset = make_dataset(val_df, is_train=False)
test_dataset = make_dataset(test_df, is_train=False)
```
## Preview the dataset
```
text_batch, label_batch = next(iter(train_dataset))
for i, text in enumerate(text_batch[:5]):
label = label_batch[i].numpy()[None, ...]
print(f'Student note: {text}')
print(f'Targets: {invert_multi_hot(label[0])}')
print(' ')
```
## Vectorization
Vectorize the text to represent it as a quantitative value. We will use [TextVectorization layer](https://keras.io/api/layers/preprocessing_layers/text/text_vectorization)
```
# Get unique words in student notes.
vocabulary = set()
train_df['clean'].str.split().apply(vocabulary.update)
vocabulary_size = len(vocabulary)
print(vocabulary_size)
```
## Now we create our vectorization layer and map() to the [tf.data.Datasets](https://www.tensorflow.org/api_docs/python/tf/data/Dataset) created earlier.
```
text_vectorizer = layers.TextVectorization(
max_tokens=vocabulary_size, ngrams=2, output_mode='tf_idf'
)
with tf.device('/CPU:0'):
text_vectorizer.adapt(train_dataset.map(lambda text, label: text))
train_dataset = train_dataset.map(
lambda text, label: (text_vectorizer(text), label), num_parallel_calls=auto).prefetch(auto)
validation_dataset = validation_dataset.map(
lambda text, label: (text_vectorizer(text), label), num_parallel_calls=auto).prefetch(auto)
test_dataset = test_dataset.map(
lambda text, label: (text_vectorizer(text), label), num_parallel_calls=auto).prefetch(auto)
text_batch, label_batch = next(iter(train_dataset))
len(text_batch[0]), len(label_batch[0])
```
## Create a text classification model
```
def make_model():
shallow_mlp_model = keras.Sequential(
[
layers.Dense(512, activation="relu"),
layers.Dense(256, activation="relu"),
layers.Dense(lookup.vocabulary_size(), activation="softmax"),
]
)
return shallow_mlp_model
```
## Train our model
```
epochs = 2
shallow_mlp_model = make_model()
shallow_mlp_model.compile(
loss="binary_crossentropy", optimizer="adam", metrics=["categorical_accuracy"]
)
history = shallow_mlp_model.fit(
train_dataset, validation_data=validation_dataset, epochs=epochs
)
def plot_result(item):
plt.plot(history.history[item], label=item)
plt.plot(history.history["val_" + item], label="val_" + item)
plt.xlabel("Epochs")
plt.ylabel(item)
plt.title("Train and Validation {} Over Epochs".format(item), fontsize=14)
plt.legend()
plt.grid()
plt.show()
plot_result("loss")
plot_result("categorical_accuracy")
```
## Evaluate the model
```
train_dataset
validation_dataset
test_dataset
_, categorical_acc = shallow_mlp_model.evaluate(test_dataset)
print(f"Categorical accuracy on the test set: {round(categorical_acc * 100, 2)}%.")
# Create a model for inference.
model_for_inference = keras.Sequential([text_vectorizer, shallow_mlp_model])
# Create a small dataset just for demoing inference.
inference_dataset = make_dataset(test_df.sample(100), is_train=False)
text_batch, label_batch = next(iter(inference_dataset))
predicted_probabilities = model_for_inference.predict(text_batch)
# Perform inference.
for i, text in enumerate(text_batch[:10]):
label = label_batch[i].numpy()[None, ...]
print(f"Student notes: {text}")
print(f"Targets(s): {invert_multi_hot(label[0])}")
predicted_proba = [proba for proba in predicted_probabilities[i]]
all_labels = [
x
for _, x in sorted(
zip(predicted_probabilities[i], lookup.get_vocabulary()),
key=lambda pair: pair[0],
reverse=True,
)
][:len(invert_multi_hot(label[0]))]
print(f"Predicted Targets(s): ({', '.join([label for label in all_labels])})")
print(" ")
```
| github_jupyter |
# Optimizing a mesh using a Differentiable Renderer
Differentiable rendering can be used to optimize the underlying 3D properties, like geometry and lighting, by backpropagating gradients from the loss in the image space. In this tutorial, we optimize geometry and texture of a single object based on a dataset of rendered ground truth views. This tutorial demonstrates functionality in `kaolin.render.mesh`, including the key `dibr_rasterization` function. See detailed [API documentation](https://kaolin.readthedocs.io/en/latest/modules/kaolin.render.mesh.html).
In addition, we demonstrate the use of [Kaolin's 3D checkpoints and training visualization](https://kaolin.readthedocs.io/en/latest/modules/kaolin.visualize.html) with the [Omniverse Kaolin App](https://docs.omniverse.nvidia.com/app_kaolin/app_kaolin/user_manual.html).
```
!pip install -q matplotlib
import json
import os
import glob
import time
from PIL import Image
import torch
import numpy as np
from matplotlib import pyplot as plt
import kaolin as kal
# path to the rendered image (using the data synthesizer)
rendered_path = "../samples/rendered_clock/"
# path to the output logs (readable with the training visualizer in the omniverse app)
logs_path = './logs/'
# We initialize the timelapse that will store USD for the visualization apps
timelapse = kal.visualize.Timelapse(logs_path)
# Hyperparameters
num_epoch = 40
batch_size = 2
laplacian_weight = 0.1
flat_weight = 0.001
image_weight = 0.1
mask_weight = 1.
lr = 5e-2
scheduler_step_size = 15
scheduler_gamma = 0.5
texture_res = 400
# select camera angle for best visualization
test_batch_ids = [2, 5, 10]
test_batch_size = len(test_batch_ids)
```
# Generating Training Data
To optimize a mesh, typical training data includes RGB images and segmentation mask. One way to generate this data is to use the Data Generator in the [Omniverse Kaolin App](https://docs.omniverse.nvidia.com/app_kaolin/app_kaolin/user_manual.html#data-generator). We provide sample output of the app in `examples/samples/`.
## Parse synthetic data
We first need to parse the synthetic data generated by the omniverse app.
The omniverse app generate 1 file per type of data (which can be depth map, rgb image, segmentation map), and an additional metadata json file.
The json file contains two main fields:
- camera_properties: Contains all the data related to camera setting such as "clipping_range", "horizontal_aperture", "focal_length", "tf_mat"
- asset_transforms: Those are transformations that are applied by the [Omniverse Kaolin App](https://docs.omniverse.nvidia.com/app_kaolin/app_kaolin/user_manual.html#data-generator), such as rotation / translation between objects or normalization.
```
num_views = len(glob.glob(os.path.join(rendered_path,'*_rgb.png')))
train_data = []
for i in range(num_views):
data = kal.io.render.import_synthetic_view(
rendered_path, i, rgb=True, semantic=True)
train_data.append(data)
dataloader = torch.utils.data.DataLoader(train_data, batch_size=batch_size,
shuffle=True, pin_memory=True)
```
# Loading the Sphere Template
The optimization starts from deforming an input template mesh according to the input image. We will use a sphere template that provides better performance on objects without topological holes. We use "/kaolin/examples/samples/sphere.obj" for convenience.
```
mesh = kal.io.obj.import_mesh('../samples/sphere.obj', with_materials=True)
# the sphere is usually too small (this is fine-tuned for the clock)
vertices = mesh.vertices.cuda().unsqueeze(0) * 75
vertices.requires_grad = True
faces = mesh.faces.cuda()
uvs = mesh.uvs.cuda().unsqueeze(0)
face_uvs_idx = mesh.face_uvs_idx.cuda()
face_uvs = kal.ops.mesh.index_vertices_by_faces(uvs, face_uvs_idx).detach()
face_uvs.requires_grad = False
texture_map = torch.ones((1, 3, texture_res, texture_res), dtype=torch.float, device='cuda',
requires_grad=True)
# The topology of the mesh and the uvs are constant
# so we can initialize them on the first iteration only
timelapse.add_mesh_batch(
iteration=0,
category='optimized_mesh',
faces_list=[mesh.faces.cpu()],
uvs_list=[mesh.uvs.cpu()],
face_uvs_idx_list=[mesh.face_uvs_idx.cpu()],
)
```
# Preparing the losses and regularizer
During training we will use different losses:
- an image loss: an L1 loss based on RGB image.
- a mask loss: an Intersection over Union (IoU) of the segmentation mask with the soft_mask output by DIB-R rasterizer.
- a laplacian loss: to avoid deformation that are too strong.
- a flat loss: to keep a smooth surface and avoid faces intersecting.
For that we need to compute the laplacian matrix and some adjacency information
(the face idx of faces connected to each edge)
```
## Separate vertices center as a learnable parameter
vertices_init = vertices.detach()
vertices_init.requires_grad = False
# This is the center of the optimized mesh, separating it as a learnable parameter helps the optimization.
vertice_shift = torch.zeros((3,), dtype=torch.float, device='cuda',
requires_grad=True)
def recenter_vertices(vertices, vertice_shift):
"""Recenter vertices on vertice_shift for better optimization"""
vertices_min = vertices.min(dim=1, keepdim=True)[0]
vertices_max = vertices.max(dim=1, keepdim=True)[0]
vertices_mid = (vertices_min + vertices_max) / 2
vertices = vertices - vertices_mid + vertice_shift
return vertices
nb_faces = faces.shape[0]
nb_vertices = vertices_init.shape[1]
face_size = 3
## Set up auxiliary connectivity matrix of edges to faces indexes for the flat loss
edges = torch.cat([faces[:,i:i+2] for i in range(face_size - 1)] +
[faces[:,[-1,0]]], dim=0)
edges = torch.sort(edges, dim=1)[0]
face_ids = torch.arange(nb_faces, device='cuda', dtype=torch.long).repeat(face_size)
edges, edges_ids = torch.unique(edges, sorted=True, return_inverse=True, dim=0)
nb_edges = edges.shape[0]
# edge to faces
sorted_edges_ids, order_edges_ids = torch.sort(edges_ids)
sorted_faces_ids = face_ids[order_edges_ids]
# indices of first occurences of each key
idx_first = torch.where(
torch.nn.functional.pad(sorted_edges_ids[1:] != sorted_edges_ids[:-1],
(1,0), value=1))[0]
nb_faces_per_edge = idx_first[1:] - idx_first[:-1]
# compute sub_idx (2nd axis indices to store the faces)
offsets = torch.zeros(sorted_edges_ids.shape[0], device='cuda', dtype=torch.long)
offsets[idx_first[1:]] = nb_faces_per_edge
sub_idx = (torch.arange(sorted_edges_ids.shape[0], device='cuda', dtype=torch.long) -
torch.cumsum(offsets, dim=0))
nb_faces_per_edge = torch.cat([nb_faces_per_edge,
sorted_edges_ids.shape[0] - idx_first[-1:]],
dim=0)
max_sub_idx = 2
edge2faces = torch.zeros((nb_edges, max_sub_idx), device='cuda', dtype=torch.long)
edge2faces[sorted_edges_ids, sub_idx] = sorted_faces_ids
## Set up auxiliary laplacian matrix for the laplacian loss
vertices_laplacian_matrix = kal.ops.mesh.uniform_laplacian(
nb_vertices, faces)
```
# Setting up optimizer
```
optim = torch.optim.Adam(params=[vertices, texture_map, vertice_shift],
lr=lr)
scheduler = torch.optim.lr_scheduler.StepLR(optim, step_size=scheduler_step_size,
gamma=scheduler_gamma)
```
# Training
This toy tutorial optimizes geometry and texture of the mesh directly to demonstrate losses, rasterization and 3D checkpoints available in Kaolin.
These components can be combined with a neural architecture of your choice to learn tasks like image to 3D mesh.
```
for epoch in range(num_epoch):
for idx, data in enumerate(dataloader):
optim.zero_grad()
gt_image = data['rgb'].cuda()
gt_mask = data['semantic'].cuda()
cam_transform = data['metadata']['cam_transform'].cuda()
cam_proj = data['metadata']['cam_proj'].cuda()
### Prepare mesh data with projection regarding to camera ###
vertices_batch = recenter_vertices(vertices, vertice_shift)
face_vertices_camera, face_vertices_image, face_normals = \
kal.render.mesh.prepare_vertices(
vertices_batch.repeat(batch_size, 1, 1),
faces, cam_proj, camera_transform=cam_transform
)
### Perform Rasterization ###
# Construct attributes that DIB-R rasterizer will interpolate.
# the first is the UVS associated to each face
# the second will make a hard segmentation mask
face_attributes = [
face_uvs.repeat(batch_size, 1, 1, 1),
torch.ones((batch_size, nb_faces, 3, 1), device='cuda')
]
image_features, soft_mask, face_idx = kal.render.mesh.dibr_rasterization(
gt_image.shape[1], gt_image.shape[2], face_vertices_camera[:, :, :, -1],
face_vertices_image, face_attributes, face_normals[:, :, -1])
# image_features is a tuple in composed of the interpolated attributes of face_attributes
texture_coords, mask = image_features
image = kal.render.mesh.texture_mapping(texture_coords,
texture_map.repeat(batch_size, 1, 1, 1),
mode='bilinear')
image = torch.clamp(image * mask, 0., 1.)
### Compute Losses ###
image_loss = torch.mean(torch.abs(image - gt_image))
mask_loss = kal.metrics.render.mask_iou(soft_mask,
gt_mask.squeeze(-1))
# laplacian loss
vertices_mov = vertices - vertices_init
vertices_mov_laplacian = torch.matmul(vertices_laplacian_matrix, vertices_mov)
laplacian_loss = torch.mean(vertices_mov_laplacian ** 2) * nb_vertices * 3
# flat loss
mesh_normals_e1 = face_normals[:, edge2faces[:, 0]]
mesh_normals_e2 = face_normals[:, edge2faces[:, 1]]
faces_cos = torch.sum(mesh_normals_e1 * mesh_normals_e2, dim=2)
flat_loss = torch.mean((faces_cos - 1) ** 2) * edge2faces.shape[0]
loss = (
image_loss * image_weight +
mask_loss * mask_weight +
laplacian_loss * laplacian_weight +
flat_loss * flat_weight
)
### Update the mesh ###
loss.backward()
optim.step()
scheduler.step()
print(f"Epoch {epoch} - loss: {float(loss)}")
### Write 3D Checkpoints ###
pbr_material = [
{'rgb': kal.io.materials.PBRMaterial(diffuse_texture=torch.clamp(texture_map[0], 0., 1.))}
]
vertices_batch = recenter_vertices(vertices, vertice_shift)
# We are now adding a new state of the mesh to the timelapse
# we only modify the texture and the vertices position
timelapse.add_mesh_batch(
iteration=epoch,
category='optimized_mesh',
vertices_list=[vertices_batch[0]],
materials_list=pbr_material
)
```
# Visualize training
You can now use [the Omniverse app](https://docs.omniverse.nvidia.com/app_kaolin/app_kaolin) to visualize the mesh optimization over training by using the training visualizer on "./logs/", where we stored the checkpoints.
You can also show the rendered image generated by DIB-R and the learned texture map with your 2d images libraries.
```
with torch.no_grad():
# This is similar to a training iteration (without the loss part)
data_batch = [train_data[idx] for idx in test_batch_ids]
cam_transform = torch.stack([data['metadata']['cam_transform'] for data in data_batch], dim=0).cuda()
cam_proj = torch.stack([data['metadata']['cam_proj'] for data in data_batch], dim=0).cuda()
vertices_batch = recenter_vertices(vertices, vertice_shift)
face_vertices_camera, face_vertices_image, face_normals = \
kal.render.mesh.prepare_vertices(
vertices_batch.repeat(test_batch_size, 1, 1),
faces, cam_proj, camera_transform=cam_transform
)
face_attributes = [
face_uvs.repeat(test_batch_size, 1, 1, 1),
torch.ones((test_batch_size, nb_faces, 3, 1), device='cuda')
]
image_features, soft_mask, face_idx = kal.render.mesh.dibr_rasterization(
256, 256, face_vertices_camera[:, :, :, -1],
face_vertices_image, face_attributes, face_normals[:, :, -1])
texture_coords, mask = image_features
image = kal.render.mesh.texture_mapping(texture_coords,
texture_map.repeat(test_batch_size, 1, 1, 1),
mode='bilinear')
image = torch.clamp(image * mask, 0., 1.)
## Display the rendered images
f, axarr = plt.subplots(1, test_batch_size, figsize=(7, 22))
f.subplots_adjust(top=0.99, bottom=0.79, left=0., right=1.4)
f.suptitle('DIB-R rendering', fontsize=30)
for i in range(test_batch_size):
axarr[i].imshow(image[i].cpu().detach())
## Display the texture
plt.figure(figsize=(10, 10))
plt.title('2D Texture Map', fontsize=30)
plt.imshow(torch.clamp(texture_map[0], 0., 1.).cpu().detach().permute(1, 2, 0))
```
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
%matplotlib inline
import warnings; warnings.simplefilter('ignore')
```
# Random Graphs
Draw random Delaunay triangulations
```
from graphs import draw_graph, random_delaunay, random_realization
from matplotlib import pyplot as plt
import numpy as np
def draw_random_graph( g, s, t, pos, hidden_state, realization, cut, pruned, ax=None, with_removed=False):
if ax is None:
ax = plt.subplot(111)
hidden_edges = {l[0]: k for l, k in zip(hidden_state, realization)}
if with_removed:
removed_edges = list(cut.edges()) + list(pruned.edges())
else:
removed_edges = []
return draw_graph(g, s, t, pos, hidden_edges=hidden_edges,
removed_edges=removed_edges, ax=ax)
def draw_lot_of_graphs(rows, columns, width=5, with_removed=False):
f, axs = plt.subplots(rows, columns, sharey=False, figsize=(columns * width, rows* width))
if columns == 1:
axs = np.array([axs]).T
if rows == 1:
axs = np.array([axs])
for i in range(rows):
for j in range(columns):
g, hidden_state, s, t, cut, pruned = random_delaunay(30, 0.5, 7, iters=1)
realization = random_realization(g, hidden_state, s, t)
draw_random_graph(g, s, t, g.pos, hidden_state=hidden_state, realization=realization,
cut=cut, pruned=pruned, ax=axs[i,j], with_removed=with_removed)
plt.tight_layout()
draw_lot_of_graphs(2,3)
```
# Experiments
One Experiment is defined by a type of map, a set of classifiers and a set of policies
```
from experiment import RandomDelaunayExperiment, all_classifier_samples
exp_config = {'description': 'Random triangulations test',
'map': {
'type': 'delaunay',
'number': 10,
'p_not_traversable': 0.5,
'n_hidden':7,
'size': 30,
'iters': 1
},
'classifier': {
'sigma': [0.5],
'samples': 1
},
'policy': {
'thresholds': [0, 0.5, 1]
}
}
exp = RandomDelaunayExperiment('test', exp_config, save=False, pool=1)
```
# Sample a graph
An experiment repeatedly samples a planning instance made by
- a graph
- source and target nodes
- a list of hidden states
- a realization
```
realization, planner, sources = exp.sample(0)
draw_random_graph(planner._graph, sources[0], planner.target, planner._graph.pos,
planner.hidden_state, realization, None, None)
```
# Compute all policy cost using classifiers
For a planning instance, we compute the competitive ration of all policies when using all classifiers
```
classifiers = exp_config['classifier']
policies = exp_config['policy']
all_classifier_samples(realization, planner, sources=sources,
classifier_config=classifiers,
policy_config=policies)
```
which is the same as the experiment method
```
exp.compute_sample(2)
```
that is applied to all maps to compute the final result.
```
exp.compute()
```
| github_jupyter |
# Project: Image Manipulation
[](https://colab.research.google.com/github/ShawnHymel/computer-vision-with-embedded-machine-learning/blob/master/1.1.5%20-%20Project%20-%20Loading%20and%20Manipulating%20Images/project_image_manipulation.ipynb)
Welcome to your first project in the Computer Vision with Embedded Machine Learning course! Follow along with the project prompts to complete this project. Press 'shift + enter' to run a cell.
Author: EdgeImpulse, Inc.<br>
Date: July 30, 2021<br>
License: [Apache-2.0](apache.org/licenses/LICENSE-2.0)<br>
```
### Example: Try running this cell. Press 'shift + enter'.
var = 1 + 2
print("Answer:", var)
```
Google Colab runs Jupyter Notebook, which is a web-based client application that allows you to run code (Python in this case) on a server. Similar to an interactive Python console, you can run pieces of code (stored in "cells") at a time rather than running (and debugging) an entire program.
Variables are saved between cells, unless you select *Runtime > Restart Runtime* to delete all your stored variables. You'll need to run all of your cells again if you do that.
If you need help with Jupyter Notebook, you can read the documentation here: https://jupyter-notebook.readthedocs.io/en/stable/
If this is your first time using Colab, I recommend reading through this [getting start guide](https://www.tutorialspoint.com/google_colab/google_colab_quick_guide.htm).
## Part 1: Load an Image
On the left side of Colab, click the folder icon. Click the upload icon and select one of your captured dataset images.
```
### You will likely need these libraries
import os
import PIL
```
You might need to use the `os.path.join()` function to create a string that points to the location of your image. You can read more about it here: https://docs.python.org/3/library/os.path.html#os.path.join
There are a few ways to open an image in Python. I recommend using the Image module in the PIL package to do that. You can read about the Image module here: https://pillow.readthedocs.io/en/stable/reference/Image.html. You will also need to use the `Image.convert()` function to convert the image to grayscale (documentation found on that same page).
```
### Open image and convert to grayscale.
# Placeholder--you need to save the grayscale image in the 'img' variable
img = None
# >>> ENTER YOUR CODE HERE <<<
```
We can preview the image using the `imshow()` function as part of the matplotlib library. You can read about it here: https://matplotlib.org/stable/api/_as_gen/matplotlib.pyplot.imshow.html
```
### Load the pyplot module (from matplotlib)
import matplotlib.pyplot as plt
### View the image
# At this point, your image should be loaded and saved in the 'img' variable. Let's view it.
plt.imshow(img, cmap='gray', vmin=0, vmax=255)
```
## Part 2: Convert Image to Numpy Array
PIL Image objects are useful for some object manipulation, but to really get our hands dirty, we want to convert everything to numerical arrays. We'll use Numpy to work with such arrays.
Numpy is a powerful Python library that allows us to efficiently work with multi-dimensional arrays. If this is your first time using Numpy, I recommend reading through this tutorial (try running some of the commands!): https://towardsdatascience.com/getting-started-with-numpy-59b22df56729
```
### We'll obviously need the Numpy package
import numpy as np
```
I recommend using the `asarray()` function in Numpy to conver the PIL Image object. You can read about it here: https://numpy.org/doc/stable/reference/generated/numpy.asarray.html
```
### Conver the Image object to a Numpy array
# Save your Numpy array in the 'img_array' variable
img_array = None
# >>> ENTER YOUR CODE HERE <<<
### View the array as an image
# At this point, your image should be saved as a Numpy array in the 'img_array' variable
plt.imshow(img_array, cmap='gray', vmin=0, vmax=255)
### View details about the array: print the shape, entire array, first row, and first column
# Note that Python may leave out some values when printing the entire array.
# Make sure that the first row and first column look correct!
# >>> ENTER YOUR CODE HERE <<<
```
## Part 3: Resize Image
We can use a variety of functions to change the image, which will help us curate our dataset as well as work with image data on our embedded devices.
To start, we'll resize the image to something smaller. Resizing, scaling, and zooming images is often not trivial, so we'll rely on the transform module in the scikit-image package to help us. You can read more about the resize function here: https://scikit-image.org/docs/dev/api/skimage.transform.html#skimage.transform.resize
```
### We'll use the Scikit-Image Transform module to help us resize the image
from skimage.transform import resize
### Use the "resize" function to resize your image array to 28x28 pixels.
# I recommend setting the "anti_aliasing" parameter to True.
# Print the array's shape when you are done (it should be (28, 28))
# Save your resized image in the 'img_resized' variable
img_resized = None
# >>> ENTER YOUR CODE HERE <<<
### View the new array as an image (this should look all black--that's OK!)
# At this point, your resized image should be saved in the 'img_resized' variable
plt.imshow(img_resized, cmap='gray', vmin=0, vmax=255)
```
What happened?!? Why is the image all black?
The resize function in skimage automatically *normalizes* all of the values in the image to be between 0.0 and 1.0 (they were integers between 0 and 255 previously). Feel free to print out some values from the array to verify this.
Let's go back to those 8-bit grayscale values!
You can use the Numpy [rint()](https://numpy.org/doc/stable/reference/generated/numpy.rint.html#numpy.rint) and [astype()](https://numpy.org/doc/stable/reference/generated/numpy.ndarray.astype.html) functions to round and convert to integers, respectively. You can use the [clip()](https://numpy.org/doc/stable/reference/generated/numpy.clip.html) function to clamp elements in an array to be between 2 values.
```
### Convert the normalized 0.0 to 1.0 values in your image to 0 to 255 integers
# Save this 8-bit array back into the 'img_resized' variable
# Round or truncate all elements to integer values (no floating point decimals)
# Make sure all elements are between 0 and 255 (known as "clamping")
# >>> ENTER YOUR CODE HERE <<<
### Let's try drawing the image again
# At this point, your resized image should be saved in the 'img_resized' variable
plt.imshow(img_resized, cmap='gray', vmin=0, vmax=255)
```
## Part 4: Draw on the Image
Here's your final challenge for this project: draw a white 10x10 square (unfilled, outline thickness of 1 pixel) in the middle of your image.
You must do this by manipulating the raw values in the Numpy array. You may not use other libraries to help you (such as by drawing a [Rectangle](https://matplotlib.org/stable/api/_as_gen/matplotlib.patches.Rectangle.html) with matplotlib).
I'd like you to try this exercise for two reasons:
1. You get practice slicing arrays in Numpy
2. You won't have matplotlib on some embedded systems, and you'll have to draw things like bounding boxes manually
If you have not used Numpy (or other linear algebra systems, like R and MATLAB) before, this can be tricky. I highly recommend avoiding using for loops, as Numpy is highly optimized for array operations without them. You may want to read through this tutorial to get an understanding of this style of array programming: https://realpython.com/numpy-array-programming/
```
### Change the required elements in the Numpy array so that a 10x10 white square is drawn in the
# middle. Try to avoid using for loops if possible.
# Start with your resized image
img_with_box = np.copy(img_resized)
# >>> ENTER YOUR CODE HERE <<<
### Draw the image--you should see a white squre in the middle!
# Your image should be saved in the 'img_with_square
plt.imshow(img_with_box, cmap='gray', vmin=0, vmax=255)
```
| github_jupyter |
<h1>Table of Contents<span class="tocSkip"></span></h1>
<div class="toc"><ul class="toc-item"><li><span><a href="#Training-Notebook" data-toc-modified-id="Training-Notebook-1"><span class="toc-item-num">1 </span>Training Notebook</a></span><ul class="toc-item"><li><span><a href="#Imports-&-Constants" data-toc-modified-id="Imports-&-Constants-1.1"><span class="toc-item-num">1.1 </span>Imports & Constants</a></span></li><li><span><a href="#Loading-data-&-processing-pipeline" data-toc-modified-id="Loading-data-&-processing-pipeline-1.2"><span class="toc-item-num">1.2 </span>Loading data & processing pipeline</a></span></li><li><span><a href="#Part-I:-Training-the-distance-model" data-toc-modified-id="Part-I:-Training-the-distance-model-1.3"><span class="toc-item-num">1.3 </span>Part I: Training the distance model</a></span><ul class="toc-item"><li><span><a href="#Instantiating-model,-loss-and-optimizer" data-toc-modified-id="Instantiating-model,-loss-and-optimizer-1.3.1"><span class="toc-item-num">1.3.1 </span>Instantiating model, loss and optimizer</a></span></li><li><span><a href="#Training-&-Validation-loops" data-toc-modified-id="Training-&-Validation-loops-1.3.2"><span class="toc-item-num">1.3.2 </span>Training & Validation loops</a></span></li></ul></li><li><span><a href="#Part-II:-Training-the-distance-classifier" data-toc-modified-id="Part-II:-Training-the-distance-classifier-1.4"><span class="toc-item-num">1.4 </span>Part II: Training the distance classifier</a></span><ul class="toc-item"><li><span><a href="#Dataloaders-creation" data-toc-modified-id="Dataloaders-creation-1.4.1"><span class="toc-item-num">1.4.1 </span>Dataloaders creation</a></span></li></ul></li></ul></li></ul></div>
# Training Notebook
---
In this notebook, we will train our model to perform data similarity detection.
## Imports & Constants
```
import os
import sys
# Ignore warnings
import warnings
warnings.filterwarnings("ignore")
# Used to import libraries from an absolute path starting with the project's root
module_path = os.path.abspath(os.path.join('../'))
if module_path not in sys.path:
sys.path.append(module_path)
from PIL import Image
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import time
from datetime import datetime
import importlib
import torch
import torch.nn as nn
import torch.optim as optim
from torch.utils.data.sampler import SubsetRandomSampler
from sklearn.linear_model import LogisticRegression
# Local imports
from src.dataset.similarityVectorizedDataset import SimilarityVectorizedDataset
from src.model.losses import ContrastiveLoss
BATCH_SIZE = 128
EMBEDDING_DIM = 40
EPOCHS = 25
TRAIN = 0.8
TEST = 0.1
VAL = 0.1
SHUFFLE = True
SEED = 42
LR = 1e-3
TO_SAVE = True
torch.manual_seed(SEED)
device = torch.device('cuda:0' if torch.cuda.is_available() else 'cpu')
print(f"Device used: {device}")
```
## Loading data & processing pipeline
We are loading the PyTorch compatible Dataset class and create 3 dataloaders, one per set of data (train, val, test)
```
def padding_collate(batch):
"""
Used as a PyTorch collate_fn function in PyTorch dataloaders.
Given a batch of vectors of shape (word_count, word_size),
make the word_count of each sequence uniform by doing right-side 0 padding.
/!\ The sequences size between batches may vary /!\
"""
max_shape_val = max(
[
#b[0] to get X, b[0][0] to get the first sentence of every X, b[0][1] to get the second sentence of every X
max(b[0][0].shape[0], b[0][1].shape[0]) for b in batch
]
)
X1 = []
X2 = []
y = []
for i in range(len(batch)):
#batch[i][0] is X, batch[i][1] is y
x1 = batch[i][0][0]
if x1.shape[0] < max_shape_val:
to_be_padded_shape = (max_shape_val - x1.shape[0], x1.shape[1])
padding = torch.zeros(to_be_padded_shape)
x1 = torch.cat((x1, padding), dim=0)
x2 = batch[i][0][1]
if x2.shape[0] < max_shape_val:
to_be_padded_shape = (max_shape_val - x2.shape[0], x2.shape[1])
padding = torch.zeros(to_be_padded_shape)
x2 = torch.cat((x2, padding), dim=0)
X1.append(x1)
X2.append(x2)
y.append([batch[i][1]])
X1 = torch.stack(X1)
X2 = torch.stack(X2)
return (X1, X2), torch.FloatTensor(y)
dataset = SimilarityVectorizedDataset()
# Data preparation
dataset_size = len(dataset)
indices = list(range(dataset_size))
val_split = int(np.floor(VAL * dataset_size))
test_split = int(np.floor(TEST * dataset_size))
if SHUFFLE:
np.random.seed(SEED)
np.random.shuffle(indices)
train_indices, val_indices, test_indices = indices[val_split+test_split:], indices[:val_split], indices[val_split:val_split+test_split]
# Creating PT data samplers and loaders:
train_sampler = SubsetRandomSampler(train_indices)
val_sampler = SubsetRandomSampler(val_indices)
test_sampler = SubsetRandomSampler(test_indices)
num_train_batch = int(np.ceil(TRAIN * dataset_size / BATCH_SIZE))
num_val_batch = int(np.ceil(VAL * dataset_size / BATCH_SIZE))
num_test_batch = int(np.ceil(TEST * dataset_size / BATCH_SIZE))
print("Creating dataloaders..")
dataloader_train = torch.utils.data.dataloader.DataLoader(
dataset = dataset,
batch_size = BATCH_SIZE,
collate_fn = padding_collate,
sampler = train_sampler
)
dataloader_val = torch.utils.data.dataloader.DataLoader(
dataset = dataset,
batch_size = BATCH_SIZE,
collate_fn = padding_collate,
sampler = val_sampler
)
dataloader_test = torch.utils.data.dataloader.DataLoader(
dataset = dataset,
collate_fn = padding_collate,
sampler = test_sampler
)
```
## Part I: Training the distance model
### Instantiating model, loss and optimizer
```
import src.model.contrastiveModel as net
importlib.reload(net)
model = net.SiameseLSTM(embedding_dim = EMBEDDING_DIM)
model.cuda()
model.train()
contrastive_loss = ContrastiveLoss()
optimizer = optim.Adam(model.parameters(), lr=LR)
scheduler = optim.lr_scheduler.ReduceLROnPlateau(optimizer, mode='max', factor=0.1, patience=0, verbose=True)
```
### Training & Validation loops
```
min_val_loss = np.inf
for i in range(EPOCHS):
print(f"Epochs {i}")
n = 1
total_loss = 0
total_duration = 0
t0 = time.time()
total_timesteps = len(dataloader_train)
for local_batch, local_labels in dataloader_train:
model.zero_grad()
# Transfer to GPU
local_batch_X1, local_batch_X2, local_labels = local_batch[0].to(device), local_batch[1].to(device), local_labels.to(device)
preds1, preds2 = model(local_batch_X1, local_batch_X2)
preds = torch.dist(preds1, preds2, 2)
# Compute the loss, gradients, and update the parameters by
#loss = loss_function(preds, local_labels)
loss = contrastive_loss(preds, local_labels)
loss.backward()
optimizer.step()
# Statistics to follow progress
total_loss += loss.item()
duration = time.time() - t0
total_duration += duration
total_duration = round(total_duration, 2)
estimated_duration_left = round((total_duration / n) * (total_timesteps), 2)
print(f"\r Epochs {i} - Loss: {total_loss/n} - Acc: {0} - Batch: {n}/{num_train_batch} - Dur: {total_duration}s/{estimated_duration_left}s", end="")
n+=1
t0 = time.time()
print("\n")
# End of epochs validation
n = 1
total_loss = 0
with torch.no_grad():
model.eval()
for local_batch, local_labels in dataloader_val:
# Transfer to GPU
local_batch_X1, local_batch_X2, local_labels = local_batch[0].to(device), local_batch[1].to(device), local_labels.to(device)
preds1, preds2 = model(local_batch_X1, local_batch_X2)
preds = torch.dist(preds1, preds2, 2)
loss = contrastive_loss(preds, local_labels)
total_loss += loss.item()
acc=0
print(f"\r Epochs {i} - Val_loss: {total_loss/n} - Batch: {n}/{num_val_batch}", end="")
n+=1
if TO_SAVE and total_loss < min_val_loss:
print("Saving weights..")
date = datetime.now().strftime("%m_%d_%H_%M_%S" )
torch.save(model.state_dict(), f"siamese_lstm_sequence_{date}_epoch{i}.pt")
min_val_loss = total_loss
model.train()
print("\n---")
date = datetime.now().strftime("%m_%d_%H_%M_%S" )
torch.save(model.state_dict(), f"siamese_smaller_lstm_sequence_{date}_epoch{i}.pt")
```
## Part II: Training the distance classifier
### Dataloaders creation
```
def model_and_titles_to_distance_dataset(model, dataloader):
"""
Given a similarity learning model and a dataloader, transforms the data of the dataloader in the shape of a distance dataset.
The distances are computed using the model. We then use the target variables to train a linear model to classify the distances
to 2 different classes: similar or dissimilar.
"""
X = []
y = []
n = 1
total_duration = 0
total_steps = len(dataloader)
model.eval()
for local_batch, local_labels in dataloader:
t0 = time.time()
# Transfer to GPU
local_batch_X1, local_batch_X2, local_labels = local_batch[0].to(device), local_batch[1].to(device), local_labels.to(device)
preds1, preds2 = model(local_batch_X1, local_batch_X2)
preds = torch.dist(preds1, preds2, 2)
with torch.no_grad():
# Transfering distances and labels to cpu
distances_cpu = preds.cpu().numpy().reshape(-1, 1)
labels = torch.flatten(local_labels).cpu().numpy().reshape(-1, 1)
# Fitting logreg
X.append(distances_cpu)
y.append(labels)
duration = time.time() - t0
total_duration += duration
per_step_mean_duration = total_duration / n
rest_of_time = per_step_mean_duration * (total_steps)
n+=1
print(f"\r{n}-{total_steps} (ETA: {total_duration}/{rest_of_time}s)", end="")
model.train()
X = np.array(X).flatten().reshape(-1, 1)
y = np.array(y).flatten()
return X, y
dataloader_train = torch.utils.data.dataloader.DataLoader(
dataset = dataset,
collate_fn = padding_collate,
sampler = train_sampler
)
dataloader_val = torch.utils.data.dataloader.DataLoader(
dataset = dataset,
collate_fn = padding_collate,
sampler = val_sampler
)
dataloader_test = torch.utils.data.dataloader.DataLoader(
dataset = dataset,
collate_fn = padding_collate,
sampler = test_sampler
)
print("Generating training dataset...")
X_train, y_train = model_and_titles_to_distance_dataset(model, dataloader_train)
print("Generating validation dataset...")
X_test, y_test = model_and_titles_to_distance_dataset(model, dataloader_test)
print("Generating test dataset...")
X_val, y_val = model_and_titles_to_distance_dataset(model, dataloader_val)
logreg = LogisticRegression()
logreg.fit(X_train, y_train)
train_acc = logreg.score(X_train, y_train)
val_acc = logreg.score(X_val, y_val)
test_acc = logreg.score(X_test, y_test)
print(f"\nFinal perfs: {train_acc} - {val_acc} - {test_acc}")
from sklearn.metrics import plot_roc_curve
plot_roc_curve(logreg, X_test, y_test)
```
| github_jupyter |
# 10장. 회귀 분석으로 연속적 타깃 변수 예측하기
**아래 링크를 통해 이 노트북을 주피터 노트북 뷰어(nbviewer.jupyter.org)로 보거나 구글 코랩(colab.research.google.com)에서 실행할 수 있습니다.**
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://nbviewer.jupyter.org/github/rickiepark/python-machine-learning-book-2nd-edition/blob/master/code/ch10/ch10.ipynb"><img src="https://jupyter.org/assets/main-logo.svg" width="28" />주피터 노트북 뷰어로 보기</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/rickiepark/python-machine-learning-book-2nd-edition/blob/master/code/ch10/ch10.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />구글 코랩(Colab)에서 실행하기</a>
</td>
</table>
`watermark`는 주피터 노트북에 사용하는 파이썬 패키지를 출력하기 위한 유틸리티입니다. `watermark` 패키지를 설치하려면 다음 셀의 주석을 제거한 뒤 실행하세요.
```
#!pip install watermark
%load_ext watermark
%watermark -u -d -v -p numpy,pandas,matplotlib,sklearn,seaborn
```
맷플롯립을 기반의 그래픽 라이브러리인 seaborn 패키지는 다음 명령으로 설치할 수 있습니다.
conda install seaborn
또는
pip install seaborn
# 주택 데이터셋 탐색
## 데이터프레임으로 주택 데이터셋 읽기
이 설명은 [https://archive.ics.uci.edu/ml/datasets/Housing](https://archive.ics.uci.edu/ml/datasets/Housing)을 참고했습니다:
속성:
<pre>
1. CRIM: 도시의 인당 범죄율
2. ZN: 25,000 평방 피트가 넘는 주택 비율
3. INDUS: 도시에서 소매 업종이 아닌 지역 비율
4. CHAS: 찰스강 인접 여부(강 주변=1, 그 외=0)
5. NOX: 일산화질소 농도(10ppm 당)
6. RM: 주택의 평균 방 개수
7. AGE: 1940년 이전에 지어진 자가 주택 비율
8. DIS: 다섯 개의 보스턴 고용 센터까지 가중치가 적용된 거리
9. RAD: 방사형으로 뻗은 고속도로까지 접근성 지수
10. TAX: $10,000당 재산세율
11. PTRATIO: 도시의 학생-교사 비율
12. B: 1000(Bk - 0.63)^2, 여기에서 Bk는 도시의 아프리카계 미국인 비율
13. LSTAT: 저소득 계층의 비율
14. MEDV: 자가 주택의 중간 가격($1,000 단위)
</pre>
```
import pandas as pd
df = pd.read_csv('https://raw.githubusercontent.com/rasbt/'
'python-machine-learning-book-2nd-edition'
'/master/code/ch10/housing.data.txt',
header=None,
sep='\s+')
df.columns = ['CRIM', 'ZN', 'INDUS', 'CHAS',
'NOX', 'RM', 'AGE', 'DIS', 'RAD',
'TAX', 'PTRATIO', 'B', 'LSTAT', 'MEDV']
df.head()
```
## 데이터셋의 중요 특징을 시각화하기
```
import matplotlib.pyplot as plt
import seaborn as sns
cols = ['LSTAT', 'INDUS', 'NOX', 'RM', 'MEDV']
sns.pairplot(df[cols], height=2.5)
plt.tight_layout()
plt.show()
import numpy as np
cm = np.corrcoef(df[cols].values.T)
#sns.set(font_scale=1.5)
hm = sns.heatmap(cm,
cbar=True,
annot=True,
square=True,
fmt='.2f',
annot_kws={'size': 15},
yticklabels=cols,
xticklabels=cols)
plt.tight_layout()
plt.show()
```
# 최소 제곱 선형 회귀 모델 구현하기
## 경사 하강법으로 회귀 모델의 파라미터 구하기
```
class LinearRegressionGD(object):
def __init__(self, eta=0.001, n_iter=20):
self.eta = eta
self.n_iter = n_iter
def fit(self, X, y):
self.w_ = np.zeros(1 + X.shape[1])
self.cost_ = []
for i in range(self.n_iter):
output = self.net_input(X)
errors = (y - output)
self.w_[1:] += self.eta * X.T.dot(errors)
self.w_[0] += self.eta * errors.sum()
cost = (errors**2).sum() / 2.0
self.cost_.append(cost)
return self
def net_input(self, X):
return np.dot(X, self.w_[1:]) + self.w_[0]
def predict(self, X):
return self.net_input(X)
X = df[['RM']].values
y = df['MEDV'].values
from sklearn.preprocessing import StandardScaler
sc_x = StandardScaler()
sc_y = StandardScaler()
X_std = sc_x.fit_transform(X)
y_std = sc_y.fit_transform(y[:, np.newaxis]).flatten()
lr = LinearRegressionGD()
lr.fit(X_std, y_std)
plt.plot(range(1, lr.n_iter+1), lr.cost_)
plt.ylabel('SSE')
plt.xlabel('Epoch')
plt.show()
def lin_regplot(X, y, model):
plt.scatter(X, y, c='steelblue', edgecolor='white', s=70)
plt.plot(X, model.predict(X), color='black', lw=2)
return
lin_regplot(X_std, y_std, lr)
plt.xlabel('Average number of rooms [RM] (standardized)')
plt.ylabel('Price in $1000s [MEDV] (standardized)')
plt.show()
print('기울기: %.3f' % lr.w_[1])
print('절편: %.3f' % lr.w_[0])
num_rooms_std = sc_x.transform(np.array([[5.0]]))
price_std = lr.predict(num_rooms_std)
print("$1,000 단위 가격: %.3f" % sc_y.inverse_transform(price_std))
```
## 사이킷런으로 회귀 모델의 가중치 추정하기
```
from sklearn.linear_model import LinearRegression
slr = LinearRegression()
slr.fit(X, y)
y_pred = slr.predict(X)
print('기울기: %.3f' % slr.coef_[0])
print('절편: %.3f' % slr.intercept_)
lin_regplot(X, y, slr)
plt.xlabel('Average number of rooms [RM]')
plt.ylabel('Price in $1000s [MEDV]')
plt.show()
```
**정규 방정식**을 사용한 방법:
```
# 1로 채워진 열 벡터 추가
Xb = np.hstack((np.ones((X.shape[0], 1)), X))
w = np.zeros(X.shape[1])
z = np.linalg.inv(np.dot(Xb.T, Xb))
w = np.dot(z, np.dot(Xb.T, y))
print('기울기: %.3f' % w[1])
print('절편: %.3f' % w[0])
```
# RANSAC을 사용하여 안정된 회귀 모델 훈련하기
```
from sklearn.linear_model import RANSACRegressor
ransac = RANSACRegressor(LinearRegression(),
max_trials=100,
min_samples=50,
loss='absolute_loss',
residual_threshold=5.0,
random_state=0)
ransac.fit(X, y)
inlier_mask = ransac.inlier_mask_
outlier_mask = np.logical_not(inlier_mask)
line_X = np.arange(3, 10, 1)
line_y_ransac = ransac.predict(line_X[:, np.newaxis])
plt.scatter(X[inlier_mask], y[inlier_mask],
c='steelblue', edgecolor='white',
marker='o', label='Inliers')
plt.scatter(X[outlier_mask], y[outlier_mask],
c='limegreen', edgecolor='white',
marker='s', label='Outliers')
plt.plot(line_X, line_y_ransac, color='black', lw=2)
plt.xlabel('Average number of rooms [RM]')
plt.ylabel('Price in $1000s [MEDV]')
plt.legend(loc='upper left')
plt.show()
print('기울기: %.3f' % ransac.estimator_.coef_[0])
print('절편: %.3f' % ransac.estimator_.intercept_)
```
# 선형 회귀 모델의 성능 평가
```
from sklearn.model_selection import train_test_split
X = df.iloc[:, :-1].values
y = df['MEDV'].values
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.3, random_state=0)
slr = LinearRegression()
slr.fit(X_train, y_train)
y_train_pred = slr.predict(X_train)
y_test_pred = slr.predict(X_test)
import numpy as np
import scipy as sp
ary = np.array(range(100000))
%timeit np.linalg.norm(ary)
%timeit sp.linalg.norm(ary)
%timeit np.sqrt(np.sum(ary**2))
plt.scatter(y_train_pred, y_train_pred - y_train,
c='steelblue', marker='o', edgecolor='white',
label='Training data')
plt.scatter(y_test_pred, y_test_pred - y_test,
c='limegreen', marker='s', edgecolor='white',
label='Test data')
plt.xlabel('Predicted values')
plt.ylabel('Residuals')
plt.legend(loc='upper left')
plt.hlines(y=0, xmin=-10, xmax=50, color='black', lw=2)
plt.xlim([-10, 50])
plt.tight_layout()
plt.show()
from sklearn.metrics import r2_score
from sklearn.metrics import mean_squared_error
print('훈련 MSE: %.3f, 테스트 MSE: %.3f' % (
mean_squared_error(y_train, y_train_pred),
mean_squared_error(y_test, y_test_pred)))
print('훈련 R^2: %.3f, 테스트 R^2: %.3f' % (
r2_score(y_train, y_train_pred),
r2_score(y_test, y_test_pred)))
```
# 회귀에 규제 적용하기
```
from sklearn.linear_model import Lasso
lasso = Lasso(alpha=0.1)
lasso.fit(X_train, y_train)
y_train_pred = lasso.predict(X_train)
y_test_pred = lasso.predict(X_test)
print(lasso.coef_)
print('훈련 MSE: %.3f, 테스트 MSE: %.3f' % (
mean_squared_error(y_train, y_train_pred),
mean_squared_error(y_test, y_test_pred)))
print('훈련 R^2: %.3f, 테스트 R^2: %.3f' % (
r2_score(y_train, y_train_pred),
r2_score(y_test, y_test_pred)))
```
릿지 회귀:
```
from sklearn.linear_model import Ridge
ridge = Ridge(alpha=1.0)
```
라쏘 회귀:
```
from sklearn.linear_model import Lasso
lasso = Lasso(alpha=1.0)
```
엘라스틱 넷 회귀:
```
from sklearn.linear_model import ElasticNet
elanet = ElasticNet(alpha=1.0, l1_ratio=0.5)
```
# 선형 회귀 모델을 다항 회귀로 변환하기
```
X = np.array([258.0, 270.0, 294.0,
320.0, 342.0, 368.0,
396.0, 446.0, 480.0, 586.0])\
[:, np.newaxis]
y = np.array([236.4, 234.4, 252.8,
298.6, 314.2, 342.2,
360.8, 368.0, 391.2,
390.8])
from sklearn.preprocessing import PolynomialFeatures
lr = LinearRegression()
pr = LinearRegression()
quadratic = PolynomialFeatures(degree=2)
X_quad = quadratic.fit_transform(X)
# 선형 특성 학습
lr.fit(X, y)
X_fit = np.arange(250, 600, 10)[:, np.newaxis]
y_lin_fit = lr.predict(X_fit)
# 이차항 특성 학습
pr.fit(X_quad, y)
y_quad_fit = pr.predict(quadratic.fit_transform(X_fit))
# 결과 그래프
plt.scatter(X, y, label='training points')
plt.plot(X_fit, y_lin_fit, label='linear fit', linestyle='--')
plt.plot(X_fit, y_quad_fit, label='quadratic fit')
plt.legend(loc='upper left')
plt.tight_layout()
plt.show()
y_lin_pred = lr.predict(X)
y_quad_pred = pr.predict(X_quad)
print('훈련 MSE 비교 - 선형 모델: %.3f, 다항 모델: %.3f' % (
mean_squared_error(y, y_lin_pred),
mean_squared_error(y, y_quad_pred)))
print('훈련 R^2 비교 - 선형 모델: %.3f, 다항 모델: %.3f' % (
r2_score(y, y_lin_pred),
r2_score(y, y_quad_pred)))
```
## 주택 데이터셋을 사용한 비선형 관계 모델링
```
X = df[['LSTAT']].values
y = df['MEDV'].values
regr = LinearRegression()
# 이차, 삼차 다항식 특성을 만듭니다
quadratic = PolynomialFeatures(degree=2)
cubic = PolynomialFeatures(degree=3)
X_quad = quadratic.fit_transform(X)
X_cubic = cubic.fit_transform(X)
# 학습된 모델을 그리기 위해 특성 범위를 만듭니다
X_fit = np.arange(X.min(), X.max(), 1)[:, np.newaxis]
regr = regr.fit(X, y)
y_lin_fit = regr.predict(X_fit)
linear_r2 = r2_score(y, regr.predict(X))
regr = regr.fit(X_quad, y)
y_quad_fit = regr.predict(quadratic.fit_transform(X_fit))
quadratic_r2 = r2_score(y, regr.predict(X_quad))
regr = regr.fit(X_cubic, y)
y_cubic_fit = regr.predict(cubic.fit_transform(X_fit))
cubic_r2 = r2_score(y, regr.predict(X_cubic))
# 결과 그래프를 그립니다
plt.scatter(X, y, label='training points', color='lightgray')
plt.plot(X_fit, y_lin_fit,
label='linear (d=1), $R^2=%.2f$' % linear_r2,
color='blue',
lw=2,
linestyle=':')
plt.plot(X_fit, y_quad_fit,
label='quadratic (d=2), $R^2=%.2f$' % quadratic_r2,
color='red',
lw=2,
linestyle='-')
plt.plot(X_fit, y_cubic_fit,
label='cubic (d=3), $R^2=%.2f$' % cubic_r2,
color='green',
lw=2,
linestyle='--')
plt.xlabel('% lower status of the population [LSTAT]')
plt.ylabel('Price in $1000s [MEDV]')
plt.legend(loc='upper right')
plt.show()
```
데이터셋을 변환합니다:
```
X = df[['LSTAT']].values
y = df['MEDV'].values
# 특성을 변환합니다
X_log = np.log(X)
y_sqrt = np.sqrt(y)
# 학습된 모델을 그리기 위해 특성 범위를 만듭니다
X_fit = np.arange(X_log.min()-1, X_log.max()+1, 1)[:, np.newaxis]
regr = regr.fit(X_log, y_sqrt)
y_lin_fit = regr.predict(X_fit)
linear_r2 = r2_score(y_sqrt, regr.predict(X_log))
# 결과 그래프를 그립니다
plt.scatter(X_log, y_sqrt, label='training points', color='lightgray')
plt.plot(X_fit, y_lin_fit,
label='linear (d=1), $R^2=%.2f$' % linear_r2,
color='blue',
lw=2)
plt.xlabel('log(% lower status of the population [LSTAT])')
plt.ylabel('$\sqrt{Price \; in \; \$1000s \; [MEDV]}$')
plt.legend(loc='lower left')
plt.tight_layout()
plt.show()
```
# 랜덤 포레스트를 사용하여 비선형 관계 다루기
## 결정 트리 회귀
```
from sklearn.tree import DecisionTreeRegressor
X = df[['LSTAT']].values
y = df['MEDV'].values
tree = DecisionTreeRegressor(max_depth=3)
tree.fit(X, y)
sort_idx = X.flatten().argsort()
lin_regplot(X[sort_idx], y[sort_idx], tree)
plt.xlabel('% lower status of the population [LSTAT]')
plt.ylabel('Price in $1000s [MEDV]')
plt.show()
```
## 랜덤 포레스트 회귀
```
X = df.iloc[:, :-1].values
y = df['MEDV'].values
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.4, random_state=1)
from sklearn.ensemble import RandomForestRegressor
forest = RandomForestRegressor(n_estimators=1000,
criterion='mse',
random_state=1,
n_jobs=-1)
forest.fit(X_train, y_train)
y_train_pred = forest.predict(X_train)
y_test_pred = forest.predict(X_test)
print('훈련 MSE: %.3f, 테스트 MSE: %.3f' % (
mean_squared_error(y_train, y_train_pred),
mean_squared_error(y_test, y_test_pred)))
print('훈련 R^2: %.3f, 테스트 R^2: %.3f' % (
r2_score(y_train, y_train_pred),
r2_score(y_test, y_test_pred)))
plt.scatter(y_train_pred,
y_train_pred - y_train,
c='steelblue',
edgecolor='white',
marker='o',
s=35,
alpha=0.9,
label='training data')
plt.scatter(y_test_pred,
y_test_pred - y_test,
c='limegreen',
edgecolor='white',
marker='s',
s=35,
alpha=0.9,
label='test data')
plt.xlabel('Predicted values')
plt.ylabel('Residuals')
plt.legend(loc='upper left')
plt.hlines(y=0, xmin=-10, xmax=50, lw=2, color='black')
plt.xlim([-10, 50])
plt.tight_layout()
plt.show()
```
| github_jupyter |
#GINN
This notebook is an exemple of how to deploy GINN in the wild.
GINN is a missing data imputation algorithm, you can find more info [HERE](https://arxiv.org/pdf/1905.01907.pdf).
To install this framework use pip.
```
!pip install git+https://github.com/spindro/GINN.git
```
In this example we use the heart dataset and remove features completely at random. This dataset contains both categorical and numerical features and will show the ability of handling them at the same time. You just need to specify wich columns contains categorical variables and viceversa.
```
import csv
import numpy as np
from sklearn import model_selection, preprocessing
from ginn import GINN
from ginn.utils import degrade_dataset, data2onehot
datafile_w = 'heart.csv'
X = np.zeros((303,13),dtype='float')
y = np.zeros((303,1),dtype='int')
with open(datafile_w,'r') as f:
reader=csv.reader(f)
for i, row in enumerate(reader):
data=[float(datum) for datum in row[:-1]]
X[i]=data
y[i]=row[-1]
cat_cols = [1,2,5,6,8,10,11,12,13]
num_cols = [0,3,4,7,9]
y = np.reshape(y,-1)
num_classes = len(np.unique(y))
```
We divide the dataset in train and test set to show what our framework can do when new data arrives. We induce missingness with a completely at random mechanism and remove 20% of elements from the data matrix of both sets. We store also the matrices indicating wether an element is missing or not.
```
missingness= 0.2
seed = 42
x_train, x_test, y_train, y_test = model_selection.train_test_split(
X, y, test_size=0.3, stratify=y
)
cx_train, cx_train_mask = degrade_dataset(x_train, missingness,seed, np.nan)
cx_test, cx_test_mask = degrade_dataset(x_test, missingness,seed, np.nan)
cx_tr = np.c_[cx_train, y_train]
cx_te = np.c_[cx_test, y_test]
mask_tr = np.c_[cx_train_mask, np.ones(y_train.shape)]
mask_te = np.c_[cx_test_mask, np.ones(y_test.shape)]
```
Here we proprecess the data applying a one-hot encoding for the categorical variables.
We get the encoded dataset three different masks that indicates the missing features and if these features are categorical or numerical, plus the new columns for the categorical variables with their one-hot range.
```
[oh_x, oh_mask, oh_num_mask, oh_cat_mask, oh_cat_cols] = data2onehot(
np.r_[cx_tr,cx_te], np.r_[mask_tr,mask_te], num_cols, cat_cols
)
```
We scale the features with a min max scaler that will preserve the one-hot encoding
```
oh_x_tr = oh_x[:x_train.shape[0],:]
oh_x_te = oh_x[x_train.shape[0]:,:]
oh_mask_tr = oh_mask[:x_train.shape[0],:]
oh_num_mask_tr = oh_mask[:x_train.shape[0],:]
oh_cat_mask_tr = oh_mask[:x_train.shape[0],:]
oh_mask_te = oh_mask[x_train.shape[0]:,:]
oh_num_mask_te = oh_mask[x_train.shape[0]:,:]
oh_cat_mask_te = oh_mask[x_train.shape[0]:,:]
scaler_tr = preprocessing.MinMaxScaler()
oh_x_tr = scaler_tr.fit_transform(oh_x_tr)
scaler_te = preprocessing.MinMaxScaler()
oh_x_te = scaler_te.fit_transform(oh_x_te)
```
Now we are ready to impute the missing values on the training set!
```
imputer = GINN(oh_x_tr,
oh_mask_tr,
oh_num_mask_tr,
oh_cat_mask_tr,
oh_cat_cols,
num_cols,
cat_cols
)
imputer.fit()
imputed_tr = scaler_tr.inverse_transform(imputer.transform())
### OR ###
# imputed_ginn = scaler_tr.inverse_transform(imputer.fit_transorm())
# for the one-liners
```
In case arrives new data, you can just reuse the model...
* Add the new data
* Impute!
```
imputer.add_data(oh_x_te,oh_mask_te,oh_num_mask_te,oh_cat_mask_te)
imputed_te = imputer.transform()
imputed_te = scaler_te.inverse_transform(imputed_te[x_train.shape[0]:])
### OR ###
# imputed_te = scaler_te.inverse_transform(imputer.fit_transorm()[x_train.shape[0]:])
# for the one-liners
```
... or fine tune the model on this new evidence.
```
imputer.fit(fine_tune=True)
imputed_te_ft = imputer.transform()
imputed_te_ft = scaler_te.inverse_transform(imputed_te_ft[x_train.shape[0]:])
### OR ###
# imputed_te_ft = scaler_te.inverse_transform(imputer.fit_transorm()[x_train.shape[0]:])
# for the one-liners
```
Now use your imputed dataset as you wish!
| github_jupyter |
```
import os
use_gpu = False
if use_gpu:
os.environ['CUDA_VISIBLE_DEVICES'] = '1'
import random
import pandas as pd
import cv2
import numpy as np
import Mask_RCNN.model as modellib
from Mask_RCNN.inference_config import inference_config
from Mask_RCNN.bowl_dataset import BowlDataset
import Mask_RCNN.visualize as visualize
import Mask_RCNN.utils as utils
import matplotlib.pyplot as plt
%matplotlib inline
def get_ax(rows=1, cols=1, size=8):
"""Return a Matplotlib Axes array to be used in
all visualizations in the notebook. Provide a
central point to control graph sizes.
Change the default size attribute to control the size
of rendered images
"""
_, ax = plt.subplots(rows, cols, figsize=(size*cols, size*rows))
return ax
ROOT_DIR = os.getcwd()
MODEL_DIR = os.path.join(ROOT_DIR, "model/checkpoint/")
print (MODEL_DIR)
# Recreate the model in inference mode
model = modellib.MaskRCNN(mode="inference",
config=inference_config,
model_dir=MODEL_DIR)
# Get path to saved weights
# Either set a specific path or find last trained weights
model_path = os.path.join(MODEL_DIR, "RCNN_checkpoint.h5")
# model_path = model.find_last()[1]
print (model_path)
# Load trained weights (fill in path to trained weights here)
assert model_path != "", "Provide path to trained weights"
print("Loading weights from ", model_path)
model.load_weights(model_path, by_name=True)
TEST_DIR = './test_2/'
dataset_test = BowlDataset()
dataset_test.load_bowl(TEST_DIR)
dataset_test.prepare()
sample_submission = pd.read_csv('./stage2_sample_submission_final.csv')
output = []
visualize_all = False
ix = random.randint(0, len(sample_submission.ImageId)-1)
if visualize_all != True:
visualize_image_ids = sample_submission.ImageId[ix:ix+1]
else:
visualize_image_ids = sample_submission.ImageId
for image_id in visualize_image_ids:
image_path = os.path.join(TEST_DIR, image_id, 'images', image_id + '.png')
print (image_path)
original_image = cv2.imread(image_path)
results = model.detect([original_image], verbose=1)
r = results[0]
visualize.display_instances(original_image,
r['rois'],
r['masks'],
r['class_ids'],
['', 'nuclei'],
r['scores'],
ax=get_ax())
image_path = os.path.join("./test.jpg")
print ("loading images from: " + str(image_path))
original_image = cv2.imread(image_path)
results = model.detect([original_image],
verbose=1)
r = results[0]
visualize.display_instances(original_image,
r['rois'],
r['masks'],
r['class_ids'],
['', 'nuclei'],
r['scores'],
ax=get_ax())
```
| github_jupyter |
```
from google.colab import drive
drive.mount('/content/gdrive')
import os
os.chdir('/content/gdrive/My Drive/finch/tensorflow1/multi_turn_rewrite/chinese/main')
%tensorflow_version 1.x
!pip install texar
import tensorflow as tf
import texar.tf as tx
import numpy as np
import pprint
import logging
from pathlib import Path
from modified_beam_search_decoder import BeamSearchDecoder
print("TensorFlow Version", tf.__version__)
print('GPU Enabled:', tf.test.is_gpu_available())
def get_vocab(f_path):
word2idx = {}
with open(f_path) as f:
for i, line in enumerate(f):
line = line.rstrip('\n')
word2idx[line] = i
return word2idx
def align_pad(li):
max_len = max([len(sent) for sent in li])
for sent in li:
if len(sent) < max_len:
sent += [0] * (max_len - len(sent))
def data_generator(f_paths, params):
for f_path in f_paths:
with open(f_path) as f:
print('Reading', f_path)
for line in f:
line = line.rstrip()
h1, h2, q, a = line.split('\t')
char2idx_fn = lambda x: [params['char2idx'].get(c, len(params['char2idx'])) for c in list(x)]
h1, h2, q, a = char2idx_fn(h1), char2idx_fn(h2), char2idx_fn(q), char2idx_fn(a)
a = [c for c in a if c in (h1 + h2 + q)]
align_pad([h1, h2])
a_in = [1] + a
a_out = a + [2]
q = q + [2]
nested = ({'history': [h1, h2], 'query': q}, (a_in, a_out))
yield nested
def dataset(is_training, params):
_shapes = ({'history':[None, None], 'query':[None]}, ([None], [None]))
_types = ({'history':tf.int32, 'query': tf.int32}, (tf.int32, tf.int32))
_pads = ({'history':0, 'query':0}, (0, 0))
if is_training:
ds = tf.data.Dataset.from_generator(
lambda: data_generator(params['train_path'], params),
output_shapes = _shapes,
output_types = _types,)
ds = ds.shuffle(params['buffer_size'])
ds = ds.padded_batch(params['batch_size'], _shapes, _pads)
ds = ds.prefetch(tf.data.experimental.AUTOTUNE)
else:
ds = tf.data.Dataset.from_generator(
lambda: data_generator(params['test_path'], params),
output_shapes = _shapes,
output_types = _types,)
ds = ds.padded_batch(params['batch_size'], _shapes, _pads)
ds = ds.prefetch(tf.data.experimental.AUTOTUNE)
return ds
def clip_grads(loss):
variables = tf.trainable_variables()
pprint.pprint(variables)
grads = tf.gradients(loss, variables)
clipped_grads, _ = tf.clip_by_global_norm(grads, params['clip_norm'])
return zip(clipped_grads, variables)
def rnn_cell():
def cell_fn():
cell = tf.nn.rnn_cell.GRUCell(params['hidden_units'],
kernel_initializer=tf.orthogonal_initializer())
return cell
if params['dec_layers'] > 1:
cells = []
for i in range(params['dec_layers']):
if i == params['dec_layers'] - 1:
cells.append(cell_fn())
else:
cells.append(tf.nn.rnn_cell.ResidualWrapper(cell_fn(), residual_fn=lambda i,o: tf.concat((i,o), -1)))
return tf.nn.rnn_cell.MultiRNNCell(cells)
else:
return cell_fn()
def dec_cell(enc_out, q_enc_len):
h_enc_out, q_enc_out = enc_out
attn_h = tf.contrib.seq2seq.BahdanauAttention(
num_units = params['hidden_units'],
memory = h_enc_out,
memory_sequence_length = None)
attn_q = tf.contrib.seq2seq.BahdanauAttention(
num_units = params['hidden_units'],
memory = q_enc_out,
memory_sequence_length = q_enc_len)
return tf.contrib.seq2seq.AttentionWrapper(
cell = rnn_cell(),
attention_mechanism = [attn_h, attn_q],
attention_layer_size = [params['hidden_units']//2, params['hidden_units']//2])
class Pointer(tf.layers.Layer):
def __init__(self, encoder_ids, encoder_out, vocab_size, is_beam_search):
super().__init__()
self.encoder_ids = tf.cast(encoder_ids, tf.int32)
self.encoder_out = encoder_out
self.vocab_size = vocab_size
self.is_beam_search = is_beam_search
def call(self, inputs):
_max_len = tf.shape(self.encoder_ids)[1]
_batch_size_ori = tf.shape(inputs)[0]
if self.is_beam_search:
_batch_size= _batch_size_ori * params['beam_width']
else:
_batch_size = _batch_size_ori
inputs = tf.reshape(inputs, (_batch_size, params['hidden_units']))
attn_weights = tf.matmul(self.encoder_out, tf.expand_dims(inputs, -1))
attn_weights = tf.squeeze(attn_weights, -1)
updates = tf.nn.softmax(attn_weights)
batch_nums = tf.range(0, _batch_size)
batch_nums = tf.expand_dims(batch_nums, axis=1)
batch_nums = tf.tile(batch_nums, [1, _max_len])
indices = tf.stack([batch_nums, self.encoder_ids], axis=2)
if self.is_beam_search:
x = tf.scatter_nd(indices, updates, (_batch_size, self.vocab_size))
return tf.reshape(x, (_batch_size_ori, params['beam_width'], self.vocab_size))
else:
x = tf.scatter_nd(indices, updates, (_batch_size, self.vocab_size))
return x
def compute_output_shape(self, input_shape):
return input_shape[:-1].concatenate(self.vocab_size)
class OutputProj(tf.layers.Layer):
def __init__(self, h_encoder_ids, q_encoder_ids, h_enc_out, q_enc_out, vocab_size, is_beam_search):
super().__init__()
self.h_pointer = Pointer(h_encoder_ids, h_enc_out, vocab_size, is_beam_search)
self.q_pointer = Pointer(q_encoder_ids, q_enc_out, vocab_size, is_beam_search)
self.vocab_size = vocab_size
def build(self, input_shape):
self.gate_fc = tf.layers.Dense(1, tf.sigmoid, use_bias=False)
super().build(input_shape)
def call(self, inputs):
h_dist = self.h_pointer(inputs)
q_dist = self.q_pointer(inputs)
gate = self.gate_fc(inputs)
return gate * h_dist + (1 - gate) * q_dist
def compute_output_shape(self, input_shape):
return input_shape[:-1].concatenate(self.vocab_size)
def bigru_encode(encoder, x, mask):
enc_out, state_fw, state_bw = encoder(x, mask=mask)
enc_state = tf.concat((state_fw, state_bw), axis=-1)
return enc_out, enc_state
def teach_forcing(labels, embedding, enc_out, enc_len, enc_state, batch_sz, params, is_training, encoder_ids):
h_enc_out, q_enc_out = enc_out
h_ids, q_ids = encoder_ids
_, q_enc_len = enc_len
output_proj = OutputProj(h_ids, q_ids, h_enc_out, q_enc_out, len(params['char2idx'])+1, is_beam_search=False)
dec_inputs, dec_outputs = labels
dec_seq_len = tf.count_nonzero(dec_inputs, 1, dtype=tf.int32)
dec_inputs = tf.nn.embedding_lookup(embedding, dec_inputs)
dec_inputs = tf.layers.dropout(dec_inputs, params['dropout_rate'], training=is_training)
cell = dec_cell((h_enc_out, q_enc_out), q_enc_len)
init_state = cell.zero_state(batch_sz, tf.float32).clone(
cell_state=enc_state)
helper = tf.contrib.seq2seq.TrainingHelper(
inputs = dec_inputs,
sequence_length = dec_seq_len,)
decoder = tf.contrib.seq2seq.BasicDecoder(
cell = cell,
helper = helper,
initial_state = init_state,
output_layer = output_proj)
decoder_output, _, _ = tf.contrib.seq2seq.dynamic_decode(
decoder = decoder,
maximum_iterations = tf.reduce_max(dec_seq_len))
return decoder_output.rnn_output
def beam_search(embedding, enc_out, enc_len, enc_state, batch_sz, params, encoder_ids):
h_enc_out, q_enc_out = enc_out
h_enc_out_t = tf.contrib.seq2seq.tile_batch(h_enc_out, params['beam_width'])
q_enc_out_t = tf.contrib.seq2seq.tile_batch(q_enc_out, params['beam_width'])
enc_state_t = tf.contrib.seq2seq.tile_batch(enc_state, params['beam_width'])
h_ids, q_ids = encoder_ids
h_ids_t = tf.contrib.seq2seq.tile_batch(h_ids, params['beam_width'])
q_ids_t = tf.contrib.seq2seq.tile_batch(q_ids, params['beam_width'])
_, q_enc_len = enc_len
q_enc_len_t = tf.contrib.seq2seq.tile_batch(q_enc_len, params['beam_width'])
output_proj = OutputProj(h_ids_t, q_ids_t, h_enc_out_t, q_enc_out_t, len(params['char2idx'])+1, is_beam_search=True)
cell = dec_cell((h_enc_out_t, q_enc_out_t), q_enc_len_t)
init_state = cell.zero_state(batch_sz*params['beam_width'], tf.float32).clone(
cell_state=enc_state_t)
decoder = BeamSearchDecoder(
cell = cell,
embedding = embedding,
start_tokens = tf.tile(tf.constant([1], tf.int32), [batch_sz]),
end_token = 2,
initial_state = init_state,
beam_width = params['beam_width'],
output_layer = output_proj,)
decoder_output, _, _ = tf.contrib.seq2seq.dynamic_decode(
decoder = decoder,
maximum_iterations = params['max_len'],)
return decoder_output.predicted_ids[:, :, 0]
def forward(features, labels, mode):
history = features['history']
query = features['query']
is_training = (mode == tf.estimator.ModeKeys.TRAIN)
batch_sz = tf.shape(query)[0]
query_valid_len = tf.count_nonzero(query, 1, dtype=tf.int32)
query_mask = tf.sign(query)
num_history = tf.shape(history)[1]
history_len = tf.shape(history)[2]
history = tf.reshape(history, (num_history*batch_sz, history_len))
history_mask = tf.sign(history)
history_ = tf.reshape(history, (batch_sz, num_history*history_len))
encoder_ids = (history_, query)
with tf.variable_scope('Embedding'):
embedding = tf.Variable(np.load('../vocab/char.npy'),
dtype=tf.float32,
name='fasttext_vectors')
def embed_fn(x):
x = tf.nn.embedding_lookup(embedding, x)
x = tf.layers.dropout(x, params['dropout_rate'], training=is_training)
return x
query = embed_fn(query)
history = embed_fn(history)
with tf.variable_scope('Encoder'):
encoder = tf.keras.layers.Bidirectional(tf.keras.layers.GRU(
params['hidden_units'], return_state=True, return_sequences=True, zero_output_for_mask=True))
query_out, query_state = bigru_encode(encoder, query, query_mask)
history_out, history_state = bigru_encode(encoder, history, history_mask)
history_out = tf.reshape(history_out, (batch_sz, num_history*history_len, 2*params['hidden_units']))
fc_h = tf.keras.layers.Dense(params['hidden_units'], params['activation'], name='fc_encode_history')
fc_q = tf.keras.layers.Dense(params['hidden_units'], params['activation'], name='fc_encode_query')
encoder_out = (fc_h(history_out), fc_q(query_out))
encoder_state = tf.layers.dense(query_state, params['hidden_units'], params['activation'], name='fc_encode_state')
enc_len = (None, query_valid_len)
with tf.variable_scope('Decoder'):
if is_training or (mode == tf.estimator.ModeKeys.EVAL):
return teach_forcing(labels, embedding, encoder_out, enc_len, encoder_state, batch_sz, params, is_training, encoder_ids)
else:
return beam_search(embedding, encoder_out, enc_len, encoder_state, batch_sz, params, encoder_ids)
def clr(step,
initial_learning_rate,
maximal_learning_rate,
step_size,
scale_fn,
scale_mode,):
step = tf.cast(step, tf.float32)
initial_learning_rate = tf.convert_to_tensor(
initial_learning_rate, name='initial_learning_rate')
dtype = initial_learning_rate.dtype
maximal_learning_rate = tf.cast(maximal_learning_rate, dtype)
step_size = tf.cast(step_size, dtype)
cycle = tf.floor(1 + step / (2 * step_size))
x = tf.abs(step / step_size - 2 * cycle + 1)
mode_step = cycle if scale_mode == 'cycle' else step
return initial_learning_rate + (
maximal_learning_rate - initial_learning_rate) * tf.maximum(
tf.cast(0, dtype), (1 - x)) * scale_fn(mode_step)
def cross_entropy_loss(logits, labels, vocab_size, smoothing):
soft_targets = tf.one_hot(tf.cast(labels, tf.int32), depth=vocab_size)
soft_targets = ((1-smoothing) * soft_targets) + (smoothing / vocab_size)
logits = tf.minimum(1., logits + 1e-6)
log_probs = tf.log(logits)
xentropy = - tf.reduce_sum(soft_targets * log_probs, axis=-1)
weights = tf.to_float(tf.not_equal(labels, 0))
xentropy *= weights
return tf.reduce_sum(xentropy) / tf.reduce_sum(weights)
def model_fn(features, labels, mode, params):
logits_or_ids = forward(features, labels, mode)
if mode == tf.estimator.ModeKeys.PREDICT:
return tf.estimator.EstimatorSpec(mode, predictions=logits_or_ids)
dec_inputs, dec_outputs = labels
loss_op = cross_entropy_loss(logits_or_ids, dec_outputs, len(params['char2idx'])+1, params['label_smoothing'])
if mode == tf.estimator.ModeKeys.TRAIN:
global_step=tf.train.get_or_create_global_step()
decay_lr = clr(
step = global_step,
initial_learning_rate = 1e-4,
maximal_learning_rate = 8e-4,
step_size = 8 * params['buffer_size'] // params['batch_size'],
scale_fn=lambda x: 1 / (2.0 ** (x - 1)),
scale_mode = 'cycle',)
train_op = tf.train.AdamOptimizer(decay_lr).apply_gradients(
clip_grads(loss_op), global_step=global_step)
hook = tf.train.LoggingTensorHook({'lr': decay_lr}, every_n_iter=100)
return tf.estimator.EstimatorSpec(
mode=mode, loss=loss_op, train_op=train_op, training_hooks=[hook],)
if mode == tf.estimator.ModeKeys.EVAL:
return tf.estimator.EstimatorSpec(mode=mode, loss=loss_op)
def minimal_test(estimator):
test_str = '成都房价是多少|不买就后悔了成都房价还有上涨空间|买不起'
h1, h2, q = test_str.split('|')
char2idx_fn = lambda x: [params['char2idx'].get(c, len(params['char2idx'])) for c in list(x)]
h1, h2, q = char2idx_fn(h1), char2idx_fn(h2), char2idx_fn(q)
q = q + [2]
align_pad([h1, h2])
predicted = list(estimator.predict(tf.estimator.inputs.numpy_input_fn(
x = {'history':np.reshape([h1, h2], (1, 2, len(h1))),
'query':np.reshape(q, (1, len(q)))},
shuffle = False)))[0]
predicted = ''.join([params['idx2char'].get(idx, '<unk>') for idx in predicted if (idx != 0 and idx != 2)])
print('-'*12)
print('minimal test')
print('Q:', test_str)
print('A:', predicted)
print('-'*12)
params = {
'model_dir': '../model/pointer_gru_clr',
'log_path': '../log/pointer_gru_clr.txt',
'train_path': ['../data/train_pos.txt', '../data/train_neg.txt'],
'test_path': ['../data/test_pos.txt'],
'vocab_path': '../vocab/char.txt',
'max_len': 30,
'activation': tf.nn.elu,
'dropout_rate': .2,
'hidden_units': 300,
'dec_layers': 1,
'num_hops': 3,
'gating_fn': tf.sigmoid,
'beam_width': 10,
'clip_norm': .1,
'buffer_size': 18986 * 2,
'batch_size': 32,
'num_patience': 10,
'label_smoothing': .1,
}
params['char2idx'] = get_vocab(params['vocab_path'])
params['idx2char'] = {idx: char for char, idx in params['char2idx'].items()}
# Create directory if not exist
Path(os.path.dirname(params['log_path'])).mkdir(exist_ok=True)
Path(params['model_dir']).mkdir(exist_ok=True, parents=True)
# Logging
logger = logging.getLogger('tensorflow')
logger.setLevel(logging.INFO)
fh = logging.FileHandler(params['log_path'])
logger.addHandler(fh)
# Create an estimator
estimator = tf.estimator.Estimator(
model_fn=model_fn,
model_dir=params['model_dir'],
config=tf.estimator.RunConfig(
save_checkpoints_steps=params['buffer_size']//params['batch_size']+1,
keep_checkpoint_max=3),
params=params)
best_em = 0.
count = 0
tf.enable_eager_execution()
while True:
estimator.train(input_fn=lambda: dataset(is_training=True, params=params))
minimal_test(estimator)
# BLEU
labels = [label for _, (_, label) in dataset(is_training=False, params=params)]
labels = [j for i in labels for j in i.numpy()]
labels = [[params['idx2char'].get(idx, '<unk>') for idx in arr if (idx!=0 and idx!=2)] for arr in labels]
preds = list(estimator.predict(input_fn=lambda: dataset(is_training=False, params=params)))
assert len(labels) == len(preds)
preds = [[params['idx2char'].get(idx, '<unk>') for idx in arr if (idx!=0 and idx!=2)] for arr in preds]
em = [np.array_equal(p, l) for p, l in zip(preds, labels)]
em = np.asarray(em).mean()
bleu, bleu_1, bleu_2, bleu_3, bleu_4 = tx.evals.corpus_bleu_moses(list_of_references=[[l] for l in labels], hypotheses=preds, return_all=True)
logger.info("BLEU: {:.3f}, BELU-1: {:.3f}, BLEU-2: {:.3f}, BLEU-4: {:.3f}, EM: {:.3f}".format(bleu, bleu_1, bleu_2, bleu_4, em))
if em > best_em:
best_em = em
count = 0
else:
count += 1
logger.info("Best EM: {:.3f}".format(best_em))
if count == params['num_patience']:
print(params['num_patience'], "times not improve the best result, therefore stop training")
break
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import allel
import matplotlib.pyplot as plt
from sklearn.decomposition import PCA
```
# Import variants VCF file & Data Cleaning
```
vcf = allel.read_vcf("../data/raw/1349 sample and all 253k unfiltered SNPs.vcf", )
variants = np.char.array(vcf["variants/CHROM"].astype(str)) + ":" + np.char.array(vcf["variants/POS"].astype(str))
vcf_arr = vcf["calldata/GT"].astype("float")
vcf_arr[vcf_arr == -1] = np.nan
mutations = vcf_arr
# mutations = np.abs(mutations)
mutations = mutations.sum(axis=2)
mutations = mutations.T
mutations.shape
mutations_df = pd.DataFrame(data=mutations, index=vcf["samples"], columns=variants)
mutations_df.shape
mutations_df.dropna(axis=1, how="any", thresh=800, inplace=True)
mutations_df.shape
mutations_df.dropna(axis=0, how="any", thresh=200000, inplace=True)
mutations_df.fillna(value=0, inplace=True)
mutations_df.isna().sum().sum()
```
# Subset patients
```
samples_phenotypes = pd.read_table("../data/raw/Sample metadata.csv", sep=",")
samples_phenotypes.set_index("ID", inplace=True)
good_samples = pd.read_table("../data/interim/samples_metadata.csv", sep=",")
good_samples.set_index("ID", inplace=True)
good_samples = good_samples[good_samples["SRC"] != "LGS"]
good_samples = good_samples[good_samples["SRC"] != "D2"]
good_samples = good_samples[good_samples["SRC"] != "U2"]
SLE_samples = good_samples[good_samples["SLE"] == 1]
SLE_samples.shape
hla_protein_samples = pd.Index(['55062', '56104', '34903', '16820', '41060', '54687', '44119', '48523',
'33287', '14947', '21560', '87483', '42335', '30146', '28289', '40007'])
highdsdna_samples = pd.Index(["32588", "55062"]) # High dsDNA
lowdsdna_samples = pd.Index(["54687", "16820"]) # low dsDNA
validation_samples = highdsdna_samples.append(lowdsdna_samples)
validation_samples.shape
```
# Filtered data
```
# filtered_df = mutations_df
filtered_df = mutations_df.filter(items=SLE_samples.index, axis=0)
filtered_df.shape
```
# Subset variants by gene
```
variants_top56_genes = pd.read_table("../data/interim/variants_top56_genes.csv", sep=",")
variants_top56_genes.set_index("Variant ID", inplace=True)
variants_top56_genes = variants_top56_genes.filter(items=["Gene(s)", "Variant ID"])
genes_matched_variants = filtered_df.columns & variants_top56_genes.index
mutations_gene_matched = filtered_df.T.join(variants_top56_genes, how="right")
mutations_by_gene = {}
mutations_gene_gb = mutations_gene_matched.groupby("Gene(s)")
for x in mutations_gene_gb.groups:
mutations_gene_df = mutations_gene_gb.get_group(x)
gene_name = mutations_gene_df["Gene(s)"].iloc[0]
print(gene_name)
mutations_gene_df = mutations_gene_df.drop(columns=["Gene(s)"]).dropna(axis=0).T
# mutations_gene_df.to_csv("../data/processed/SNPs_by_gene/"+gene_name+".csv")
mutations_by_gene[gene_name] = mutations_gene_df
```
# Gene - mutations
```
variant_genes = pd.read_table("../data/raw/216k Variants with gene name.csv", sep=",")
variant_genes = variant_genes[variant_genes["Gene(s)"] != "?"]
# variant_genes = variant_genes.filter(items=["Gene(s)", "Variant ID"])
variant_genes["Position"] = variant_genes["Position"].astype("object")
genes_56 = variant_genes["Gene(s)"].value_counts()[:56]
```
# Run PCA
```
def get_top_k_components(snp_data, var_threshold=0.80, return_fit_transform=False):
pca = PCA()
pca.fit(snp_data)
top_k = np.argmax(np.cumsum(np.power(pca.singular_values_, 2)) /
np.sum(np.power(pca.singular_values_, 2)) > var_threshold)
if return_fit_transform:
pca.n_components = top_k
pca.n_components_ = top_k
return pca.transform(snp_data)[:, :top_k]
else:
return pca.components_[:top_k]
def get_top_variant_by_coef(pca_components, coef_percentile=70):
gene_coefs_sum = np.abs(pca_components).sum(axis=0)
coefs_percentile = np.percentile(gene_coefs_sum, coef_percentile)
return np.where(gene_coefs_sum > coefs_percentile)
def select_top_variants(snp_data, var_threshold=0.80, coef_percentile=70):
variants = snp_data.columns
top_k_components = get_top_k_components(snp_data, var_threshold)
top_variants_idx = get_top_variant_by_coef(top_k_components, coef_percentile)
return variants[top_variants_idx]
plt.figure(figsize=(10, 7))
plt.title("Variant coefficients from PCA top-k components feature selection for "+gene)
plt.ylabel("coefficient")
plt.xlabel("genes")
plt.scatter(x=range(pca.components_.shape[1]), y=gene_coefs_sum)
```
# Compute PCA projections of all SNP data
```
pca_projs_by_gene = {}
for gene in mutations_by_gene.keys():
pca_projs_by_gene[gene] = get_top_k_components(mutations_by_gene[gene],
var_threshold=0.90,
return_fit_transform=True)
print(gene, mutations_by_gene[gene].shape[1], pca_projs_by_gene[gene].shape)
genes = list(pca_projs_by_gene.keys())
num_top_components = [pca_projs_by_gene[gene].shape[1] for gene in genes]
plt.figure(figsize=(12, 7), dpi=150)
plt.title("Top-k PCA components selected with 90% variance threshold, split by genes, only SLE")
plt.xlabel("Genes")
plt.xticks(rotation=90)
plt.ylabel("# of PCA components selected")
plt.bar(x=genes, height=num_top_components)
pca_projs_concat = np.concatenate([pca_projs_by_gene[gene] for gene in pca_projs_by_gene.keys()],
axis=1)
pca_projs_concat.shape
pca_proj_names = []
for i, gene in enumerate(mutations_by_gene.keys()):
for n in range(num_top_components[i]):
pca_proj_names.append(gene + "_" + str(n))
len(pca_proj_names)
pca_projs_concat_df = pd.DataFrame(pca_projs_concat, index=mutations_gene_df.index, columns=pca_proj_names)
pca_projs_concat_df.index.name = "Sample ID"
patient_gene_proj_scores = pd.DataFrame(index=pca_projs_concat_df.index, columns=genes)
for gene in genes:
projs_by_gene = pca_projs_concat_df.iloc[:, pca_projs_concat_df.columns.str.contains(gene)]
patient_gene_proj_scores[gene] = np.power(projs_by_gene, 2).sum(axis=1)/projs_by_gene.columns.shape[0]
patient_gene_proj_scores
patient_gene_proj_scores.to_csv("../data/interim/patient_gene_proj_scores.csv")
pca_projs_concat_df
pca_projs_concat_df.to_csv("../data/interim/PCA_projections_80_variance_SLE_samples.csv",
index=True)
```
# Select variants by percentile
```
top_variants_by_gene = {}
for gene in mutations_by_gene.keys():
top_variants_by_gene[gene] = select_top_variants(mutations_by_gene[gene], var_threshold=0.80, coef_percentile=70)
print(gene, mutations_by_gene[gene].shape[1], top_variants_by_gene[gene].shape[0])
all_top_variants = []
for gene in top_variants_by_gene.keys():
all_top_variants.extend(top_variants_by_gene[gene])
len(all_top_variants)
```
# Concatenate all selected variant into one DataFrame
```
mutations_df = mutations_df.loc[:,~mutations_df.columns.duplicated()]
mutations_top_variants_df = mutations_df.filter(items=all_top_variants)
mutations_top_variants_df
mutations_top_variants_df.to_csv("../data/interim/mutations_top_variants_80_variance.csv")
```
# Graph the eigenvalues
```
import matplotlib.pyplot as plt
plt.figure(figsize=(10, 7))
plt.bar(x=range(pca.singular_values_.shape[0]),
height=pca.singular_values_, )
plt.ylabel('log of singular values')
plt.xlabel('Principal components')
plt.title('Elbow plot from PCA of mutations data')
plt.xlim(-1, 45)
# plt.ylim(0, 10)
plt.show()
```
| github_jupyter |
```
from IPython.core.display import display, HTML, Markdown
def is_subseq(s, subs):
"""
bool: verifica se subs è sottosequenza di s.
"""
found = 0
pos_r = 0
while pos_r < len(s):
if s[pos_r] == subs[found]:
found+=1
if found >= len(subs):
return True
pos_r += 1
return False
def is_monotone(s, mono_type):
"""
(bool, NO_cert_string): verifica se s presenta la monotonicità di tipo mono_type (vedi tabella):
"SC" : strettamente crescente
"ND" : non-decrescente
"SD" : strettamente decrescente
"NC" : non-crescente
In caso affermativo il bool è True.
Altrimenti il bool è False e viene restituita una stringa che riporta una violazione puntuale alla monotonicità richiesta.
"""
for i in range(1,len(s)):
if s[i] < s[i-1] and mono_type in {"SC","ND"}:
return (0,f"L'elemento {s[i]} in posizione {i+1} è minore dell'elemento {s[i-1]} in posizione {i}.")
if s[i] > s[i-1] and mono_type in {"SD","NC"}:
return (0,f"L'elemento {s[i]} in posizione {i+1} è maggiore dell'elemento {s[i-1]} in posizione {i}.")
if s[i] == s[i-1] and mono_type in {"SC","SD"}:
return (0,f"L'elemento {s[i]} in posizione {i+1} non è maggiore dell'elemento {s[i-1]} in posizione {i}.")
return (1,None)
def is_monotone_subseq(s, subs, mono_type):
"""
(evaluation_string): verifica se s presenta la monotonicità di tipo mono_type (vedi tabella):
"SC" : strettamente crescente
"ND" : non-decrescente
"SD" : strettamente decrescente
"NC" : non-crescente
Restituisce una stringa contenete la valutazione del certificato immesso dallo studente.
"""
submission_string = f"Hai inserito il certificato $subs= {subs}$."
submission_string += f"<br>L'istanza era data da $s= {s}$.<br>"
NO_eval = "No. Totalizzeresti <span style='color:green'>[0 safe pt]</span>, \
<span style='color:blue'>[0 possible pt]</span>, \
<span style='color:red'>[10 out of reach pt]</span>.<br>"
SI_eval = "Si. Totalizzeresti <span style='color:green'>[ 1 safe pt]</span>, \
<span style='color:blue'>[9 possible pt]</span>, \
<span style='color:red'>[0 out of reach pt]</span>.<br>"
ans,NO_cert_string = is_monotone(subs, mono_type)
if not ans:
return submission_string + NO_eval + NO_cert_string
if not is_subseq(s, subs):
return submission_string + NO_eval + "La sequenza $subs$ proposta non è sottosequenza di $s$."
return submission_string + SI_eval
def is_monotone_subseq_without_interval(s, subs, mono_type, start, end):
"""
(evaluation_string): verifica se s presenta la monotonicità di tipo mono_type (vedi tabella):
"SC" : strettamente crescente
"ND" : non-decrescente
"SD" : strettamente decrescente
"NC" : non-crescente
Restituisce una stringa contenete la valutazione del certificato immesso dallo studente sulla stringa s
privata degli elementi tra start e end
"""
aux = s[:]
del aux[start:end]
return is_monotone_subseq(aux,subs, mono_type)
```
## Esercizio \[60 pts\]
(poldo) Ricerca di sottosequenze strettamente crescenti di massima lughezza.
Si consideri la seguente sequenza di numeri naturali:
```
s = [1, 14, 8, 2, 4, 21, 28, 48, 5, 26, 49, 9, 32, 19, 12, 46, 10, 7, 3, 25, 11, 6, 29, 39, 44, 13]
print(s)
```
__Richieste__:
1. __[10 pts]__ Trovare una sottosequenza $s1$ strettamente crescente di $s$ che sia la più lunga possibile.
```
subs1 = [4]
display(Markdown(is_monotone(s, subs1, "SC")))
```
2. __[10 pts]__ Trovare una sottosequenza $s2$ strettamente decrescente di $s$ che sia la più lunga possibile.
```
subs2 = [4]
display(Markdown(is_monotone_subseq(s, subs2, "SD")))
```
3. __[10 pts]__ Trovare la più lunga sottosequenza crescente che includa l'elemento di valore 7
4. __[10 pts]__ Una sequenza è detta una _V-sequenza_ se cala fino ad un certo punto, e da lì in poi cresce sempre. Trovare la più lunga V-sequenza che sia una sottosequenza della sequenza data
5. __[20 pts]__ Qual è il minor numero possibile di colori _C_ per colorare gli elementi della sequenza in input in modo che, per ogni colore, la sottosequenza degli elementi di quel colore sia monotona non crescente? Specificare per ogni elemento il colore (come colori, usare i numeri da 1 a _C_)
| github_jupyter |
# Explaining IoU
- toc: true
- badges: true
- comments: true
- categories: [jupyter]
- image: images/chart-preview.png
# An IOU explanation and implementaion walk through
In this blogpost i will explain what is IOU, where is it used , how is it implemented
## What is IOU
IOU is pretty much clear by the name intersection over union.
The formula is
- **IOU = Area of Intersection / Area of union**
- **Area of union = First Box Area + Second Box Area -Intersection Area**

## How is it implemented(basic)
Here i will show a simple implementation in pytorch.If you look at the below picture we
will get a basic idea of how to get the intersection between two boxes, the rest are simple

For the basic implementation of this can be found in this nice [blogpost](http://ronny.rest/tutorials/module/localization_001/iou/) and from that is basic implemenation is like this
```
#collapse-hide
def batch_iou(a, b, epsilon=1e-5):
""" Given two arrays `a` and `b` where each row contains a bounding
box defined as a list of four numbers:
[x1,y1,x2,y2]
where:
x1,y1 represent the upper left corner
x2,y2 represent the lower right corner
It returns the Intersect of Union scores for each corresponding
pair of boxes.
Args:
a: (numpy array) each row containing [x1,y1,x2,y2] coordinates
b: (numpy array) each row containing [x1,y1,x2,y2] coordinates
epsilon: (float) Small value to prevent division by zero
Returns:
(numpy array) The Intersect of Union scores for each pair of bounding
boxes.
"""
# COORDINATES OF THE INTERSECTION BOXES
x1 = np.array([a[:, 0], b[:, 0]]).max(axis=0)
y1 = np.array([a[:, 1], b[:, 1]]).max(axis=0)
x2 = np.array([a[:, 2], b[:, 2]]).min(axis=0)
y2 = np.array([a[:, 3], b[:, 3]]).min(axis=0)
# AREAS OF OVERLAP - Area where the boxes intersect
width = (x2 - x1)
height = (y2 - y1)
# handle case where there is NO overlap
width[width < 0] = 0
height[height < 0] = 0
area_overlap = width * height
# COMBINED AREAS
area_a = (a[:, 2] - a[:, 0]) * (a[:, 3] - a[:, 1])
area_b = (b[:, 2] - b[:, 0]) * (b[:, 3] - b[:, 1])
area_combined = area_a + area_b - area_overlap
# RATIO OF AREA OF OVERLAP OVER COMBINED AREA
iou = area_overlap / (area_combined + epsilon)
return iou
```
## Where is it used and how to implement for that use case
But the above implementation assumes that both the bounding boxes have the same set of batches,which is rarely the case. IOU is mainly used in **object detection tasks**.
1. We will have a set of anchors for each position in the feature map,for eg say if we have a feature map of shape 5x5 and there are 3 anchors per position then there will be 5x5x3=75 total anchors
2. The Ground trouth boxes for that feature map may be much less the number of anchors
3. We need to find the matching anchors to the bounding boxes, so we can select that portion of the feature map for the downstream predictions.
### Implementing for the above use case
Basically when we get two boxes say
`a- B,M,4` -- the anchor boxes after reshaping(B,A,H,W,4) where A is number of anchors
`b- B,N,4` --the real bboxes. N is the max number of boxes in certain image and the other images will be padded with -1.
we need to compute iou between `a` and `b` so each box in `a` is compare with each box
in `b`. So we should make N copies of copies of each box in `a` to be compare with
` N bboxes`. Also if we want to vectorise this operation then we need to make `M` copies of `b`. So the final dimensions will be
`a - B,M,N,4`
`b - B,M,N,4`
Now we can say like each slice of the both `a` and `b` can be compared
```
import torch
#say the given anchors and bboxes are in shape x_top,y_top,x_btm,y_btm
sample_anchors = torch.tensor([[[[[5.,5,15,15], [25,25,35,35],[1,1,9,9]]]]]) #only 1 batch
bboxes = torch.tensor([[[1.,1,11,11], [20,20,30,30]]])
B = bboxes.shape[0]
no_of_bboxes = bboxes.shape[1]
print('sample anchors \n', sample_anchors,'\n')
print('sample bboxes \n', bboxes,'\n')
print('sample number of anchors shape ',sample_anchors.shape)
print('sample bboxes shape ',bboxes.shape,'\n')
```
Here we need to compare the 3 anchor boxes with the two bboxes, first we reshape the anchors to be of shape `batch,total_anchors,4`,
we need to compute iou between `sample_anchors` and `bboxes` so each of the `3` anchors are
compared with the bboxes which is `2` here. So for vectorized implementation we should make 3 copies of copies of each anchor in `sample_anchors` to be compare with
` 2 bboxes`. Also if we should make `3` copies of `b` to aid in vectorized implementation. So the final dimensions will be
- `sample_anchors - B,3,2,4`
- `b=boxes - B,3,2,4`
```
sample_anchors = sample_anchors.reshape(B,-1,4)
no_of_anchors = sample_anchors.shape[1]
sample_anchors = sample_anchors.unsqueeze(2).expand(-1,-1,no_of_bboxes,-1)
print(sample_anchors)
print(sample_anchors.shape)
bboxes = bboxes.unsqueeze(1).expand(-1,no_of_anchors,-1,-1)
print(bboxes)
print(bboxes.shape)
#first we need to find the intersection for that width and height of the intersection area
#this inturn can be obtained by finding the lefttop and bottom corner cordinates and subtracting them
left_top = torch.max(sample_anchors[:,:,:,:2],bboxes[:,:,:,:2])
right_bottom = torch.min(sample_anchors[:,:,:,2:],bboxes[:,:,:,2:])
delta = right_bottom - left_top
print(delta)
#The first element of delta is width and the next element is height, we can remove negative values
#since this will be boxes that are not intersecting
#(remember the the image top left if (0,0) and bottom y is positive downwards)
delta[delta<0]=0
#now find the intersection area
interesection_area = delta[:,:,:,0]*delta[:,:,:,1]
print(interesection_area)
print(interesection_area.shape)
```
A small picture represntation is tried below,we can see that first and 3rd anchors intersect with first bounding box while the 2nd anchor intersect with the next one

From the intersection area above we can see that the where there are no itersection the area is zero
and thus in this case the first and last anchor mathces with the first bbox while the second
anchor mathces with the second one
```
#now we need to find the Area of union which is
#Area of union = First Box Area + Second Box Area -Intersection Area
sample_anchors_area = (sample_anchors[:,:,:,2]-sample_anchors[:,:,:,0])*(sample_anchors[:,:,:,3] -
sample_anchors[:,:,:,1])
bbox_area = (bboxes[:,:,:,2] - bboxes[:,:,:,0]) * (bboxes[:,:,:,3] - bboxes[:,:,:,1])
iou = interesection_area/(sample_anchors_area+bbox_area - interesection_area)
print(iou)
print(iou.shape)
```
so the final iou matrix will have shape **(Batch,no_of_anchors,no_of_bboxes)**
### Downstream usage of this iou
This iou matrix will be used for calculation the regression offsets, negative anchors,ground truth class . The other place where iou is used is for mean Average Precision at the end which if possible i will explain in another post
## Complete code
Below i will provide a small code for implementing this in a batch
```
def IOU(anchors,bboxes):
#anchors B,A,H,W,4
#bboxes B,N,4
B = anchors.shape[0]
anchors = anchors.reshape(B,-1,4)
M,N = anchors.shape[1],bboxes.shape[1]
#expanding
anchors = anchors.unsqueeze(2).expand(-1,-1,N,-1)
bboxes = bboxes.unsqueeze(1).expand(-1,M,-1,-1)
left_top = torch.max(anchors[:,:,:,:2],bboxes[:,:,:,:2])
right_bottom = torch.min(anchors[:,:,:,2:],bboxes[:,:,:,2:])
delta = right_bottom - left_top
delta[delta<0] = 0
intersection_area = delta[:,:,:,0]*delta[:,:,:,1]
anchors_area = (anchors[:,:,:,2]-anchors[:,:,:,0])*(anchors[:,:,:,3] -anchors[:,:,:,1])
bbox_area = (bboxes[:,:,:,2] - bboxes[:,:,:,0])* (bboxes[:,:,:,3] - bboxes[:,:,:,1])
iou = interesection_area/(anchors_area+bbox_area - interesection_area)
return iou
```
| github_jupyter |
# TEST ANGULAR POWER SPECTRA
```
%matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
from scipy.interpolate import interp1d
from astropy.cosmology import FlatLambdaCDM
plt.rcParams.update({
'text.usetex': False,
'font.family': 'serif',
'legend.frameon': False,
'legend.handlelength': 1.5,
})
```
## 1. Load data
* Cosmology
```
cosmo = {}
with open('../../data/des-y1-test/cosmological_parameters/values.txt') as cosmo_values:
for line in cosmo_values:
if line:
key, val = line.partition('=')[::2]
cosmo[key.strip()] = float(val)
cosmo_astropy = FlatLambdaCDM(H0=cosmo['hubble'], Ob0=cosmo['omega_b'], Om0= cosmo['omega_m'], Tcmb0=2.7)
```
* Distance functions
```
zdM = np.loadtxt('../../data/des-y1-test/distances/z.txt')
dM = np.loadtxt('../../data/des-y1-test/distances/d_m.txt')
```
* Matter power spectrum
```
zp = np.loadtxt('../../data/des-y1-test/matter_power_nl/z.txt')
k_h = np.loadtxt('../../data/des-y1-test/matter_power_nl/k_h.txt')
p_h = np.loadtxt('../../data/des-y1-test/matter_power_nl/p_k.txt')
xp = np.interp(zp, zdM, dM)
```
#### 1.a. LSST Y10 quantities
Lens sample
```
nbinlsst = 10
binlsst_a, binlsst_b = np.tril_indices(nbinlsst)
binlsst_a += 1
binlsst_b += 1
import math
import scipy.integrate as integrate
zph = [0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 0.9, 1.0, 1.10, 1.20]
def sigma_lens(z):
return 0.03 * (1 + z)
def dndz(z, z0, alpha):
return np.square(z) * np.exp( - np.power(z / z0, alpha))
def nz_lens(z, z0, alpha):
def dndz_lens(z):
return dndz(z, 0.28, 0.90)
norm = integrate.quad(lambda z: dndz_lens(z), 0.0, 1100.0 )[0]
return np.square(z) * np.exp( - np.power(z / z0, alpha)) / norm
def nzi(z, z0, alpha):
ni = [0.0]*(len(zph) + 1)
xi = [0.0]*(len(zph) + 1)
for j,elem in enumerate(zph):
xi[j] = (zph[j] - z)/(sigma_lens(z) * np.sqrt(2))
for k,char in enumerate(zph):
ni[k] = 0.5*nz_lens(z,z0, alpha)*( math.erf(xi[k+1]) - math.erf(xi[k]))
return ni
```
#### 1.b. Matter power spectrum
```
zp = np.loadtxt('../../data/des-y1-test/matter_power_nl/z.txt')
k_h = np.loadtxt('../../data/des-y1-test/matter_power_nl/k_h.txt')
p_h = np.loadtxt('../../data/des-y1-test/matter_power_nl/p_k.txt')
xp = np.interp(zp, zdM, dM)
k0, kf = k_h[0]*(cosmo['hubble']/100), k_h[-1]*(cosmo['hubble']/100)
k_h2 = np.logspace(np.log10(k0), np.log10(kf), 1024)
k = k_h2*cosmo['h0']
```
* Unequal-time power spectra
```
import sys
sys.path.append("../../unequalpy")
from skypy.power_spectrum import growth_function
from approximation import growth_midpoint
from matter import matter_power_spectrum_1loop as P1loop
from matter import matter_unequal_time_power_spectrum as Puetc
from approximation import geometric_approx as Pgeom
from approximation import midpoint_approx as Pmid
d = np.loadtxt('../../data/Pfastpt.txt',unpack=True)
ks, pk, p22, p13 = d[:, 0], d[:, 1], d[:, 2], d[:, 3]
p11_int = interp1d( ks, pk, fill_value="extrapolate")
p22_int = interp1d( ks, p22, fill_value="extrapolate")
p13_int = interp1d( ks, p13, fill_value="extrapolate")
powerk = (p11_int, p22_int, p13_int)
g = growth_function(np.asarray(zp), cosmo_astropy)/growth_function(0, cosmo_astropy)
gm = growth_midpoint(np.asarray(zp), np.asarray(zp), growth_function, cosmo_astropy)
pet = P1loop(k, g, powerk)
puet = Puetc(k, g, g, powerk)
pgeom = Pgeom(pet)
pmid = Pmid(k, gm, powerk)
```
## 2. The correlation function
```
import corfu
r_uet, xi_uet = corfu.ptoxi(k, puet, q=0.2)
r_limb, xi_limb = corfu.ptoxi(k, pet, q=0, limber=True)
r_geom, xi_geom = corfu.ptoxi(k, pgeom, q=0)
r_mid, xi_mid = corfu.ptoxi(k, pmid, q=0)
plt.figure(figsize=(6,4))
plt.loglog(r_uet, +xi_uet[0,0], 'k', label='Unequal-time', lw=1)
plt.loglog(r_uet, -xi_uet[0,0], '--k', lw=1)
plt.loglog(r_limb, +xi_limb[0], '--b', label='Limber', lw=1)
plt.loglog(r_limb, -xi_limb[0], ':b', lw=1)
plt.loglog(r_geom, +xi_geom[0,0], '--r', label='Geometric', lw=1)
plt.loglog(r_geom, -xi_geom[0,0], ':r', lw=1)
plt.loglog(r_mid, +xi_mid[0,0], '--g', label='Midpoint', lw=2)
plt.loglog(r_mid, -xi_mid[0,0], ':g', lw=2)
plt.legend()
plt.xlabel('r')
plt.ylabel(r'$\xi(r)$')
plt.show()
```
## 3. Lensing filters
```
from lens_filter import filter_galaxy_clustering
```
* Redshift distribution of galaxies
```
zn = np.loadtxt('../../data/des-y1-test/nz_lens/z.txt')
nz = [np.loadtxt('../../data/des-y1-test/nz_lens/bin_%d.txt' % i) for i in range(1, 5+1)]
xf = np.interp(zn, zdM, dM)
nzlsst = np.zeros((len(zn),(len(zph) + 1)))
for i in range(0,len(zn)):
nzlsst[i] = nzi(zn[i], 0.28, 0.90)
```
* Galaxy clustering
```
g0 = growth_function(0, cosmo_astropy)
Dz = growth_function(zph, cosmo_astropy) / g0
bias_lsst = 0.95 / Dz
fglsst = [filter_galaxy_clustering(xf, zn, nzlsst[:,i], bias_lsst[i], cosmo_astropy) for i in range(10)]
```
## 4. Angular correlation function
```
theta = np.logspace(-3, np.log10(np.pi), 64)
theta_arcmin = np.degrees(theta)*60
w_limb = [corfu.eqt(theta, (xf, fglsst[a-1]*fglsst[b-1]), (xp, r_limb, xi_limb)) for a, b in zip(binlsst_a, binlsst_b)]
w_geom = [corfu.uneqt(theta, (xf, fglsst[a-1]), (xf, fglsst[b-1]), (xp, xp, r_geom, xi_geom), 1) for a, b in zip(binlsst_a, binlsst_b)]
w_uet = [corfu.uneqt(theta, (xf, fglsst[a-1]), (xf, fglsst[b-1]), (xp, xp, r_uet, xi_uet), True) for a, b in zip(binlsst_a, binlsst_b)]
w_mid = [corfu.uneqt(theta, (xf, fglsst[a-1]), (xf, fglsst[b-1]), (xp, xp, r_mid, xi_mid), True) for a, b in zip(binlsst_a, binlsst_b)]
fig, axes = plt.subplots(1,10, figsize=(17, 4), sharey=True)
for ax in axes.ravel():
ax.axis('off')
for i, (a, b) in enumerate(zip(binlsst_a, binlsst_b)):
if a == b:
ax = axes[b-1]
ax.axis('on')
ax.loglog(theta_arcmin, +w_limb[i], '--b', label='Limber', lw=1)
ax.loglog(theta_arcmin, -w_limb[i], ':b', lw=1)
ax.loglog(theta_arcmin, +w_geom[i], '--r', label='Geometric', lw=1)
ax.loglog(theta_arcmin, -w_geom[i], ':r', lw=1)
ax.loglog(theta_arcmin, +w_mid[i], '--g', label='Midpoint', lw=2)
ax.loglog(theta_arcmin, -w_mid[i], ':g', lw=2)
ax.loglog(theta_arcmin, +w_uet[i], 'k', label='Unequal-time', lw=1)
ax.loglog(theta_arcmin, -w_uet[i], '--k', lw=1)
# ax.set_xlim(5e0, 1e4)
# ax.set_ylim(1e-5, 2e-1)
ax.set_xticks([1e1, 1e2, 1e3, 1e4])
ax.tick_params(axis='y', which='minor', labelcolor='none')
string = '({0},{1})'.format(a,b)
ax.text(2e3,5e-2,string)
axes[0].legend(markerfirst=False, loc='lower left', frameon=False)
ax = fig.add_subplot(111, frameon=False)
ax.set_xlabel(r'Angular separation, $\theta$ [arcmin]', size=12)
ax.set_ylabel(r'Angular correlation, $w(\theta)$', size=12)
ax.tick_params(labelcolor='none', top=False, bottom=False, left=False, right=False)
ax.tick_params(axis='y', pad=15)
fig.tight_layout(pad=0.5)
# fig.savefig('plots/w_galaxy_clustering.pdf', bbox_inches='tight')
plt.show()
```
## 5. Angular power spectrum analysis
### 5.1. Angular power spectra
```
l_limb, cl_limb = np.transpose([corfu.wtocl(theta, w, lmax=2000) for w in w_limb], (1, 0, 2))
l_geom, cl_geom = np.transpose([corfu.wtocl(theta, w, lmax=2000) for w in w_geom], (1, 0, 2))
l_uet, cl_uet = np.transpose([corfu.wtocl(theta, w, lmax=2000) for w in w_uet], (1, 0, 2))
l_mid, cl_mid = np.transpose([corfu.wtocl(theta, w, lmax=2000)for w in w_mid], (1, 0, 2))
fig, axes = plt.subplots(1,10, figsize=(15,4), sharey=True)
for ax in axes.ravel():
ax.axis('off')
for i, (a, b) in enumerate(zip(binlsst_a, binlsst_b)):
if a == b:
ax = axes[b-1]
ax.axis('on')
ax.loglog(l_limb[i], cl_limb[i], ':b', label='Limber', lw=1)
ax.loglog(l_geom[i], cl_geom[i], ':r', label='Geometric', lw=1)
ax.loglog(l_mid[i], cl_mid[i], '--g', label='Midpoint', lw=2)
ax.loglog(l_uet[i], cl_uet[i], 'k', label='Unequal-time', lw=1)
# ax.set_xlim(5e0, 2e3)
# ax.set_ylim(2e-7, 3e-4)
ax.set_xticks([1e1, 1e2, 1e3])
string = '({0},{1})'.format(a,b)
ax.text(0.9e2,1.2e-8,string)
axes[0].legend(markerfirst=False, bbox_to_anchor=(0.9, 1.1), bbox_transform=plt.gcf().transFigure, loc='upper left')
ax = fig.add_subplot(111, frameon=False)
ax.set_xlabel(r'Angular mode, $\ell$', size=12)
ax.set_ylabel(r'Angular power, $C_{\ell}$', size=12)
ax.tick_params(labelcolor='none', top=False, bottom=False, left=False, right=False)
ax.tick_params(axis='y', pad=12)
# fig.tight_layout(pad=0.5)
# fig.savefig('plots/cl_galaxy_clustering.pdf', bbox_inches='tight')
plt.show()
```
### 5.2. Relative error
```
frac_limb = cl_limb/cl_uet
frac_geom = cl_geom/cl_uet
frac_mid = cl_mid/cl_uet
frac_uet = cl_uet/cl_uet
fig, axes = plt.subplots(1,10, figsize=(15,4), sharey=True)
for ax in axes.ravel():
ax.axis('off')
for i, (a, b) in enumerate(zip(binlsst_a, binlsst_b)):
if a == b:
ax = axes[b-1]
ax.axis('on')
ax.semilogx(l_limb[i], frac_limb[i], 'b', label='Limber', lw=1)
ax.semilogx(l_geom[i], frac_geom[i], ':r', label='Geometric', lw=1)
ax.semilogx(l_mid[i], frac_mid[i], '--g', label='Midpoint', lw=2)
ax.semilogx(l_uet[i], frac_uet[i], ':k', label='Unequal-time', lw=0.5)
ax.set_xlim(1, 2e3)
ax.set_ylim(0.95, 1.06)
ax.set_yticks([0.96, 0.98, 1, 1.02, 1.04])
string = '({0},{1})'.format(a,b)
ax.text(1e2,1.05,string)
ax.fill_between(l_limb[0], 0.99, 1.01, alpha=0.05)
ax.fill_between(l_limb[0], 0.98, 1.02, alpha=0.05)
axes[0].legend(markerfirst=False, bbox_to_anchor=(0.9, 1.1), bbox_transform=plt.gcf().transFigure, loc='upper left')
ax = fig.add_subplot(111, frameon=False)
ax.set_xlabel(r'Angular mode, $\ell$', size=12)
ax.set_ylabel(r'$C_{\ell}^{approx} / C_{\ell}^{uetc}$', size=12)
ax.tick_params(labelcolor='none', top=False, bottom=False, left=False, right=False)
ax.tick_params(axis='y', pad=12)
# fig.tight_layout(pad=0.5)
# fig.savefig('plots/fraction_cl_clustering.pdf', bbox_inches='tight')
plt.show()
fig, axes = plt.subplots(1,1, figsize=(6,4), sharex=True, sharey=True)
ax = axes
ax.axis('on')
ax.semilogx(l_limb[0], frac_limb[0], 'b', label='Limber', lw=1)
ax.semilogx(l_geom[0], frac_geom[0], ':r', label='Geometric', lw=1)
ax.semilogx(l_mid[0], frac_mid[0], '--g', label='Midpoint', lw=2)
ax.semilogx(l_uet[0], frac_uet[0], ':k', label='Unequal-time', lw=0.5)
ax.set_xlim(1, 2e3)
ax.set_ylim(0.95, 1.06)
ax.set_yticks([0.96, 0.98, 1, 1.02, 1.04])
string = '({0},{1})'.format(1,1)
ax.text(5e2,1.05,string)
ax.fill_between(l_limb[0], 0.99, 1.01, alpha=0.05)
ax.fill_between(l_limb[0], 0.98, 1.02, alpha=0.05)
axes.legend(markerfirst=False, loc='upper left')
ax = fig.add_subplot(111, frameon=False)
ax.set_xlabel(r'Angular mode, $\ell$', size=12)
ax.set_ylabel(r'$C_{\ell}^{approx} / C_{\ell}^{uetc}$', size=12)
ax.tick_params(labelcolor='none', top=False, bottom=False, left=False, right=False)
ax.tick_params(axis='y', pad=12)
ax.set_title('Galaxy clustering')
fig.tight_layout(pad=0.5)
# fig.savefig('plots/bin11_cl_clustering.pdf', bbox_inches='tight')
plt.show()
```
| github_jupyter |
# CAISO OASIS Data downloader
Data taken from [CAISO Oasis portal](http://oasis.caiso.com/mrioasis)
API documentation located [here](http://www.caiso.com/Documents/OASIS-InterfaceSpecification_v5_1_3Clean_Fall2017Release.pdf)
| SubLAP | Node Name |
|-----------------------------|-------------------------------------------------------------------------------------------------------------------------------------------------------|
| Pacific Gas & Electric | 'PGCC', 'PGEB', 'PGF1', 'PGFG', 'PGHB', 'PGKN', 'PGLP', 'PGNB', 'PGNC', 'PGNP','PGNV', 'PGP2', 'PGSA', 'PGSB', 'PGSF', 'PGSI', 'PGSN', 'PGST', 'PGZP' |
| Southern California Edison | 'SCEC', 'SCEN', 'SCEW', 'SCHD', 'SCLD', 'SCNW' |
| San Diego Gas & Electric | 'SDG1' |
| Valley Electric Association | 'VEA' |
```
## Import required libraries
import requests
from selenium import webdriver
from functools import reduce
import pandas as pd
import zipfile as zp
import time
import os
import glob
import altair as alt
from altair_saver import save
## Define required variables
month_dict = {'Jan': ['0101', '0131'],
'Feb': ['0201', '0228'],
'Mar': ['0301', '0331'],
'Apr': ['0401', '0430'],
'May': ['0501', '0531'],
'Jun': ['0601', '0630'],
'Jul': ['0701', '0731'],
'Aug': ['0801', '0831'],
'Sep': ['0901', '0930'],
'Oct': ['1001', '1031'],
'Nov': ['1101', '1130'],
'Dec': ['1201', '1231']
}
node_list = ['PGCC', 'PGEB', 'PGF1', 'PGFG', 'PGHB', 'PGKN', 'PGLP', 'PGNB', 'PGNC', 'PGNP',
'PGNV', 'PGP2', 'PGSA', 'PGSB', 'PGSF', 'PGSI', 'PGSN', 'PGST', 'PGZP',
'SCEC', 'SCEN', 'SCEW', 'SCHD', 'SCLD', 'SCNW', 'SDG1', 'VEA']
node_dict = {}
for node in node_list:
node_dict[node] = 'SLAP_' + node + '-APND'
def extract_hourly_data(year, nodes):
"""
Downloads Southern California monthly energy prices from CAISO's Oasis Portal for a year of choice
and aggregates separate files into a single csv file
Parameters:
year: str
Outputs:
None
Example:
extract_monthly_data(2017)
"""
node_entry = reduce(lambda x, y: x +','+ y, list(map(lambda x: 'SLAP_' + x + '-APND', nodes)))
name_entry = reduce(lambda x, y: x +','+ y, nodes)
chrome_options = webdriver.ChromeOptions()
prefs = {'download.default_directory' : \
'/Users/jaromeleslie/Documents/MDS/Personal_projects/CAISOprices/data/'+str(year)}
chrome_options.add_experimental_option('prefs', prefs)
driver = webdriver.Chrome(chrome_options=chrome_options)
for month in month_dict.keys():
api_call = "http://oasis.caiso.com/oasisapi/SingleZip?queryname=PRC_LMP&resultformat=6&startdatetime=" +\
str(year)+month_dict[month][0] + "T07:00-0000&enddatetime=" + str(year)+month_dict[month][1] + \
f"T07:00-0000&version=1&market_run_id=DAM&node={node_entry}"
print(api_call)
driver.get(api_call)
time.sleep(15)
zip_files = glob.glob('../data/' +str(year)+'/*.zip')
for zip_filename in zip_files:
dir_name = os.path.splitext(zip_filename)[0]
if not os.path.isdir(dir_name):
os.mkdir(dir_name)
zip_handler = zp.ZipFile(zip_filename, "r")
zip_handler.extractall(dir_name)
# path = dir_name
csv_files = glob.glob('../data/' + str(year) +'/*/*.csv')
entries =[]
for csv in csv_files:
entries.append(pd.read_csv(csv))
combined_csvs = pd.concat(entries)
combined_csvs.to_csv(f"../data/{str(year)}/{str(year)}_{name_entry}.csv")
driver.close()
print("Download and aggregation complete!")
return combined_csvs
test_2_2019 = extract_hourly_data(2019, ['SCEC', 'SCEN'])
test_2_2019.XML_DATA_ITEM.unique()
sample_day = test_2_2019.query('OPR_DT == "2019-06-01"').query('NODE == "SLAP_SCEC-APND"')
sample_day
read_test = pd.read_csv('../data/2019/2019_SCEC,SCEN.csv')
sample_day = read_test.query('OPR_DT == "2019-06-01"').query('NODE == "SLAP_SCEC-APND" and XML_DATA_ITEM == "LMP_PRC"')
sample_day
sample_chart = alt.Chart(sample_day).mark_line().encode(
x=alt.X('OPR_HR:O', axis=alt.Axis(title='Hour',format=',.0f', labelAngle=0)),
y=alt.Y('MW:Q', axis = alt.Axis( title='Price')),
color=alt.Color('XML_DATA_ITEM', legend=alt.Legend(title = 'Category',orient="right"))
).properties(title="", width=500).interactive()
sample_chart
save(sample_chart, "../img/sample_day.png")
combined_csvs['NODE_ID'].unique()
def extract_5min_data(year, node):
chrome_options = webdriver.ChromeOptions()
prefs = {'download.default_directory' : '/Users/jaromeleslie/Documents/MDS/Personal_projects/CAISO_oasis_extractor/data/5min/'+str(year)}
chrome_options.add_experimental_option('prefs', prefs)
driver = webdriver.Chrome(chrome_options=chrome_options)
for month in month_dict.keys():
api_call = "http://oasis.caiso.com/oasisapi/SingleZip?queryname=PRC_INTVL_LMP&resultformat=6&startdatetime=" + str(year)+month_dict[month][0] + "T07:00-0000&enddatetime=" + str(year)+month_dict[month][1] + "T07:00-0000&version=1&market_run_id=HASP&node=" + node
print(api_call)
driver.get(api_call)
time.sleep(15)
zip_files = glob.glob('../data/5min/' +str(year)+'/*.zip')
for zip_filename in zip_files:
dir_name = os.path.splitext(zip_filename)[0]
os.mkdir(dir_name)
zip_handler = zp.ZipFile(zip_filename, "r")
zip_handler.extractall(dir_name)
# path = dir_name
csv_files = glob.glob('../data/5min/' + str(year) +'/*/*.csv')
entries =[]
for csv in csv_files:
entries.append(pd.read_csv(csv))
combined_csvs = pd.concat(entries)
combined_csvs.to_csv('../data/5min/' + str(year) +'/'+ str(year) + node +'.csv')
extract_5min_data(2018, "CLEARKE_6_N012")
extract_monthly_15min_data(2019, "SLAP_SCEC-APND")
```
| github_jupyter |
# Introduction to Python and Natural Language Technologies
## Lecture 06, NLP Introduction
March 16, 2020
Ádám Kovács
This lecture aims to give an introduction to the main concepts of NLP and word representations.
## Preparation
[Download GLOVE](http://sandbox.hlt.bme.hu/~adaamko/glove.6B.100d.txt)
```
!pip install spacy
!pip install textacy
!pip install flair
!pip install gensim
!pip install -U scikit-learn
!python -m spacy download en
```
## NLP Tasks
### Why do we need NLP?
- Make the computer understand text
- Extract useful information from it
- A collection that helps us processing huge amount of texts
- We have two directions:
- Analysis: Convert text to a structural representation
- Generation: Generate text from formal representation
### Tasks most people would think of
- Spellchecking
- Machine translation
- Chatbots
these are not an exhaustive list
### "Real" tasks?
- Basic NLP tasks (but very important):
- tokenization
- lemmatization
- POS tagging
- syntactic parsing
- More semantic tasks:
- summarization
- question answering
- information extraction (e.g. NER tagging)
- relation extraction
- chatbots
- machine translation
- ...
## Spacy
- For demonstrating NLP tasks, we are going to use the library [spacy](https://spacy.io/) a lot.
- It is an open-source NLP library for Python
- It features a lot of out-of-the-box models for NLP
- NER, POS tagging, dependency parsing, vectorization...
```
import spacy
from spacy import displacy
#loading the english model
nlp = spacy.load('en')
```
## Basic preprocessing tasks, text normalization
<h3 id="Tokenization">Tokenization</h3>
<ul>
<li>Splitting text into words, sentences, documents, etc..</li>
<li>One of the goals of tokenizing text into words is to create a <strong>vocabulary</strong></li>
</ul>
<p><em>Muffins cost <strong>$3.88</strong> in New York. Please buy me two as I <strong>can't</strong> go. <strong>They'll</strong> taste good. I'm going to <strong>Finland's</strong> capital to hear about <strong>state-of-the-art</strong> solutions in NLP.</em></p>
- $3.88 - split on the period?
- can't - can not?
- They'll - they will?
- Finland's - Finland?
- state-of-the-art?
```
sens = "Muffins cost $3.88 in New York. Please buy me two as I can't go." \
" They'll taste good. I'm going to Finland's capital to hear about state-of-the-art solutions in NLP."
print(sens.split())
print(len(sens.split()))
sens = "Muffins cost $3.88 in New York. Please buy me two as I can't go." \
" They'll taste good. I'm going to Finland's capital to hear about state-of-the-art solutions in NLP."
doc = nlp(sens)
tokens = [token.text for token in doc]
print(tokens)
for sen in doc.sents:
print(sen)
for token in doc:
print(token.text, token.is_alpha, token.is_stop)
```
### Lemmatization, stemming
- The goal of lemmatization is to find the dictionary form of the words
- Called the "lemma" of a word
- _dogs_ -> _dog_ , _went_ -> _go_
- Ambiguity plays a role: _saw_ -> _see_?
- Needs POS tag to disambiguate
```
doc = nlp("I saw two dogs yesterday.")
lemmata = [token.lemma_ for token in doc]
print(lemmata)
```
### POS tagging
- Words can be groupped into grammatical categories.
- These are called the Part Of Speech tags of the words.
- Words belonging to the same group are interchangable
- Ambiguity: _guard_ ?
```
doc = nlp("The white dog went to play football yesterday.")
[token.pos_ for token in doc]
```
<h3 id="Morphological-analysis">Morphological analysis</h3>
<ul>
<li>Splitting words into morphemes</li>
<li>Morphemes are the smallest meaningful units in a language (part of the words)</li>
<li>friend<span style="color: #e03e2d;">s</span>, wait<span style="color: #e03e2d;">ing</span>, friend<span style="color: #e03e2d;">li</span><span style="color: #3598db;">er</span></li>
<li>Tagging them with morphological tags</li>
<li>Ambiguity: <em>várnak</em></li>
</ul>
```
doc = nlp("Yesterday I went to buy two dogs")
nlp.vocab.morphology.tag_map[doc[-1].tag_]
```
## Advanced tasks
### Syntactic parsing
- *Colorless green ideas sleep furiously.*
- *Furiously sleep ideas green colorless.*
Chomsky (1956)
Two types.
- Phrase structure grammar
- __Dependency grammar__
### Universal Dependency Parsing
- Started and standardized in the [UD](http://universaldependencies.org/) project.
- The types are Language-independent
- The annotations are trying to be consistent accross 70+ languages
```
doc = nlp("Colorless green ideas sleep furiously")
displacy.render(doc, style='dep', jupyter=True, options={'distance': 100})
```
### Named entity recognition
- Identify the present entities in the text
```
sens = "Muffins cost $3.88 in New York. Please buy me two as I can't go." \
" They'll taste good. I'm going to Finland's capital to hear about state-of-the-art solutions in NLP."
doc = nlp(sens)
for ent in doc.ents:
print(ent)
displacy.render(doc, style='ent', jupyter=True)
```
### Language modelling
- One of the most important task in NLP
- The goal is to compute the "probability" of a sentence
- Can be used in:
- Machine Translation
- Text generation
- Correcting spelling
- Word vectors?
- P(the quick brown __fox__) > P(the quick brown __stick__)
```
#!pip install transformers
from transformers import pipeline
text_generator = pipeline("text-generation")
print(text_generator("The quick brown ", max_length=10, do_sample=False))
```
## Semantic tasks
### Summarization
```
summarizer = pipeline("summarization")
summarizer("Deep learning is used almost exclusively in a Linux environment.\
You need to be comfortable using the command line if you are serious about deep learning and NLP.\
Most NLP and deep learning libraries have better support for Linux and MacOS than Windows. \
Most papers nowadays release the source code for their experiments with Linux support only.",
min_length=5)
```
### Sentiment Analysis
- In the simplest case, decide whether a text is negative or positive.
```
sentiment = pipeline("sentiment-analysis")
sentiment(['This class is really cool! I would recommend this to anyone!'])
```
### Question Answering
- Given a context and a question choose the right answer
- Can be extractive or abstractive
```
question_answerer = pipeline('question-answering')
question_answerer({
'question': 'Who went to the store ?',
'context': 'Adam went to the store yesterday.'})
```
### <center>Lexical Inference, Natural Language Inference</center>
<div class="frame">
| **entailment** | | |
|:--------------------------------------------------------------|:----|:----|
| A young family enjoys feeling ocean waves lap at their feet. | | |
| A family is at the beach | | |
| **contradiction** | | |
| There is no man wearing a black helmet and pushing a bicycle | | |
| One man is wearing a black helmet and pushing a bicycle | | |
| **neutral** | | |
| An old man with a package poses in front of an advertisement. | | |
| A man poses in front of an ad for beer. | | |
</div>
### Machine Comprehension
- https://demo.allennlp.org/reading-comprehension
### Machine translation
```
translation = pipeline("translation_en_to_de")
text = "I like to study Data Science and Machine Learning"
translated_text = translation(text, max_length=40)[0]['translation_text']
print(translated_text)
```
### Chatbots
```
#!pip install chatbotAI
from chatbot import demo
demo()
```
## Demos
- http://e-magyar.hu/hu/parser
- https://demo.allennlp.org/
- https://talktotransformer.com/
- [GPT-3](https://github.com/elyase/awesome-gpt3) (*has 175B parameters*)
## Representations
To be able to run machine learning algorithms the computer needs numerical representations. For natural text input this means we need a mapping that converts strings to a numerical represenatation. **one-hot encoding** is the easiest approach where we map each word to an integer id.
```
sentence = "yesterday the lazy dog went to the store to buy food"
mapping = dict()
max_id = 0
for word in sentence.split():
if word not in mapping:
mapping[word] = max_id
max_id = max_id + 1
print(mapping)
```
### Load matplotlib and pandas
```
import os
import pandas as pd
import re
```
# Data analysis
- we use nlp frameworks for the basic tasks
- for the preprocessing tasks (lemmatization, tokenization) we use [spaCy](https://spacy.io/)
- for keyword extraction and various text analyzation tasks we use [textacy](https://github.com/chartbeat-labs/textacy)
- textacy builds on spaCy output
- both are open source python libraries
<p><strong>AG_NEWS</strong> classes:</p>
<ul>
<li>
<p>World - <em>Venezuela Prepares for Chavez Recall Vote</em></p>
</li>
<li>
<p>Sports - <em>Johnson Back to His Best as D-Backs End Streak</em></p>
</li>
<li>
<p>Business - <em>Intel to delay product aimed for high-definition TVs</em></p>
</li>
<li>
<p>Sci/Tech - <em>China's Red Flag Linux to focus on enterprise</em></p>
</li>
</ul>
```
NGRAMS = 2
from torchtext import data
from torchtext.datasets import text_classification
import os
if not os.path.isdir('./data'):
os.mkdir('./data')
text_classification.DATASETS['AG_NEWS'](
root='./data', ngrams=NGRAMS, vocab=None)
train_data = pd.read_csv("./data/ag_news_csv/train.csv",quotechar='"', names=['label', 'title', 'description'])
test_data = pd.read_csv("./data/ag_news_csv/test.csv",quotechar='"', names=['label', 'title', 'description'])
train_data.head()
train_data.label.value_counts()
text_sports = train_data[train_data.label == 2]
text = " ".join(text_sports.title.tolist())
doc_text = nlp(text[:200000])
import textacy
from textacy.extract import ngrams
from collections import Counter
Counter([ng.text.lower() for n in [2,4] for ng in ngrams(doc_text, n)]).most_common(10)
from textacy.ke import textrank
textrank(
doc_text,
normalize = "lemma",
window_size=2, edge_weighting="binary", position_bias=False
)
textrank(
doc_text,
window_size=10, edge_weighting="count", position_bias=False
)
import math
from collections import Counter
words = [tok for tok in doc_text if tok.is_alpha and not tok.is_stop]
word_probs = {tok.text.lower(): tok.prob for tok in words}
freqs = Counter(tok.text for tok in words)
#!pip install wordcloud
from wordcloud import WordCloud
print(len(freqs))
wordcloud = WordCloud(background_color="white", max_words=30, scale=1.5).generate_from_frequencies(freqs)
image = wordcloud.to_image()
image.save("./wordcloud.png")
from IPython.display import Image
Image(filename='./wordcloud.png')
```
## Building a classficiation pipeline
The AG's news topic classification dataset is constructed by Xiang Zhang (xiang.zhang@nyu.edu) from the dataset above. It is used as a text classification benchmark in the following paper: Xiang Zhang, Junbo Zhao, Yann LeCun. Character-level Convolutional Networks for Text Classification. Advances in Neural Information Processing Systems 28 (NIPS 2015).
The AG's news topic classification dataset is constructed by choosing 4 largest classes from the original corpus. Each class contains 30,000 training samples and 1,900 testing samples. The total number of training samples is 120,000 and testing 7,600.
```
!pip install torchtext==0.4
!pip install torch
!pip install gensim
!pip install scikit-learn
NGRAMS = 2
from torchtext import data
from torchtext.datasets import text_classification
import os
if not os.path.isdir('./data'):
os.mkdir('./data')
text_classification.DATASETS['AG_NEWS'](
root='./data', ngrams=NGRAMS, vocab=None)
#Import the needed libraries
from tqdm import tqdm
from sklearn.model_selection import train_test_split as split
import numpy as np
```
Now we use pandas to read in the dataset into a DataFrame. We are also going to just take a fraction of the dataset to be more efficient.
```
#1-World, 2-Sports, 3-Business, 4-Sci/Tech
train_data = pd.read_csv("./data/ag_news_csv/train.csv",quotechar='"', names=['label', 'title', 'description'])
test_data = pd.read_csv("./data/ag_news_csv/test.csv",quotechar='"', names=['label', 'title', 'description'])
train_data = train_data.groupby('label').apply(lambda x: x.sample(frac=0.2, random_state=1234)).sample(frac=1.0)
train_data
```
We need a way of converting raw data to features!

_(image from [link](https://developers.google.com/machine-learning))_
The easiest way of converting raw data to features is called the [Bag of Words](https://en.wikipedia.org/wiki/Bag-of-words_model) model.
```
from collections import defaultdict
word_to_ix = defaultdict(int)
for sent in train_data.title:
for word in sent.split():
if word not in word_to_ix:
word_to_ix[word] = len(word_to_ix)
len(word_to_ix)
```
We are going to use Python's machine learning library, called scikit-learn to build a classical ML pipeline
```
from sklearn.feature_extraction.text import CountVectorizer
corpora = ['hello my name is adam','i am the instructor for this class']
# instantiate the vectorizer object
vectorizer = CountVectorizer()
# convert th documents into a matrix
wm = vectorizer.fit_transform(corpora)
#retrieve the terms found in the corpora
tokens = vectorizer.get_feature_names()
df_vect = pd.DataFrame(data = wm.toarray(),index = ['Doc1','Doc2'],columns = tokens)
df_vect
vectorizer = CountVectorizer(max_features=10000, stop_words="english")
X = vectorizer.fit(train_data.title)
X.vocabulary_
c = X.transform(["Hello my name is adam"]).toarray()
print(c)
```
## Displaying the most frequent terms from CountVectorizer
```
import matplotlib.pyplot as plt
%matplotlib inline
import matplotlib
matplotlib.style.use('ggplot')
matplotlib.pyplot.rcParams['figure.figsize'] = (16, 10)
matplotlib.pyplot.rcParams['font.family'] = 'sans-serif'
matplotlib.pyplot.rcParams['font.size'] = 20
vectorize = CountVectorizer(stop_words="english")
c = vectorize.fit(train_data.title)
C = c.transform(train_data.title)
summ = np.sum(C,axis=0)
total = np.squeeze(np.asarray(summ))
term_freq_df = pd.DataFrame([total],columns=c.get_feature_names()).transpose()
term_freq_df.columns = ["frequency"]
term_freq_df.sort_values(by="frequency", ascending=False)
y_pos = np.arange(500)
plt.figure(figsize=(10,8))
s = 1
expected_zipf = [term_freq_df.sort_values(by='frequency', ascending=False)['frequency'][0]/(i+1)**s for i in y_pos]
plt.bar(y_pos, term_freq_df.sort_values(by='frequency', ascending=False)['frequency'][:500], align='center', alpha=0.5)
plt.plot(y_pos, expected_zipf, color='r', linestyle='--',linewidth=2,alpha=0.5)
plt.ylabel('Frequency')
plt.title('Top 500 tokens')
```
We first build a feature extraction method that takes raw texts as input and runs builds features on the whole dataset
```
import gensim
from tqdm import tqdm
from sklearn.model_selection import train_test_split as split
import numpy as np
def vectorize_to_bow(tr_data, tst_data):
tr_vectors = X.transform(tr_data)
tst_vectors = X.transform(tst_data)
return tr_vectors, tst_vectors
def get_features_and_labels(data, labels):
tr_data,tst_data,tr_labels,tst_labels = split(data,labels, test_size=0.3, random_state=1234)
tst_vecs = []
tr_vecs = []
tr_vecs, tst_vecs = vectorize_to_bow(tr_data, tst_data)
return tr_vecs, tr_labels, tst_vecs, tst_labels
tr_vecs, tr_labels, tst_vecs, tst_labels = get_features_and_labels(train_data.title, train_data.label)
tr_vecs.shape
```
<h2 id="Machine-Learning">Machine Learning</h2>
<ul>
<li>We have a datasets with labels</li>
<li>We can train a machine learning algorithm using the labels as "gold" data - <span style="color: #e03e2d;">Supervised learning</span></li>
<li>The algorithm will predict unseen data points using the trained model</li>
<li>We will use <a href="https://scikit-learn.org/stable/" target="_blank" rel="noopener">sklearn</a> for the models</li>
</ul>
### Logistic Regression
- One of the simplest method for classification tasks

```
#Import a bunch of stuff from sklearn
from sklearn.ensemble import RandomForestClassifier, AdaBoostClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import cross_val_score
lr = LogisticRegression(n_jobs=-1)
lr.fit(tr_vecs, tr_labels)
#!pip install eli5
import eli5
eli5.show_weights(lr, feature_names=X.get_feature_names())
from sklearn.metrics import accuracy_score
print(type(tst_vecs))
lr_pred = lr.predict(tst_vecs)
print("Logistic Regression Test accuracy : {}".format(accuracy_score(tst_labels, lr_pred)))
```
Bag of words are the simplest method for featurizing your data. If we want a more sophisticated method, we could use [TF-IDf](https://scikit-learn.org/stable/modules/generated/sklearn.feature_extraction.text.TfidfVectorizer.html).
- __TF__: The term frequency of a word in a document.
- __IDF__: The inverse document frequency of the word across a set of documents. This means, how common or rare a word is in the entire document set. The closer it is to 0, the more common a word is.
- The higher the score, the more relevant that word is in that particular document
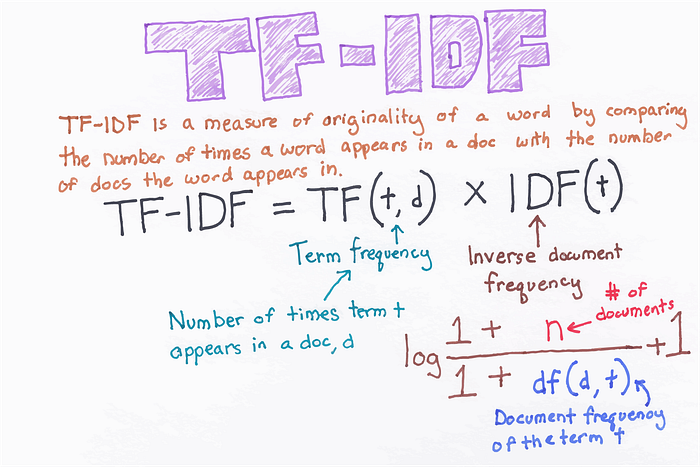

_(image from [link](https://miro.medium.com))_
```
from sklearn.feature_extraction.text import TfidfVectorizer
vectorizer = TfidfVectorizer(max_features=1000, use_idf=True)
vectors = vectorizer.fit(train_data.title)
tfidf_vectorizer_vectors = vectors.transform(train_data.title)
first_vector_tfidfvectorizer=tfidf_vectorizer_vectors[6]
# place tf-idf values in a pandas data frame
df = pd.DataFrame(first_vector_tfidfvectorizer.T.todense(), index=vectors.get_feature_names(), columns=["tfidf"])
df = df.sort_values(by=["tfidf"],ascending=False)
df
```
Sklearn allows us to build [pipelines](https://scikit-learn.org/stable/modules/generated/sklearn.pipeline.Pipeline.html) with defining each step of the pipeline, like:
- Vectorizers
- Classifiers
- Voting strategies
- Optionally merge feature extraction from multiple sources
## Problems
- When representing words with id's we assign them to the words in the order of the encounter.
- This means that we may assign different vectors to the words each time we run the algorithm.
- Doesn't include any concept of similarity, e.g: `similarity(embedding(cat, dog)) > similarity(embedding(cat, computer))`
- The representation is very sparse and could have very high dimension, which would also slow the computations. The size is given by the vocabulary of our corpus, that can be over 100000 dimension.
## Word embeddings
- map each word to a small dimensional (around 100-300) continuous vectors.
- this means that similar words should have similar vectors.
- what do we mean by word similarity ?
### Cosine similarity
- Now that we have word vectors, we need a way to quantify the similarity between individual words, according to these vectors. One such metric is cosine-similarity. We will be using this to find words that are "close" and "far" from one another.
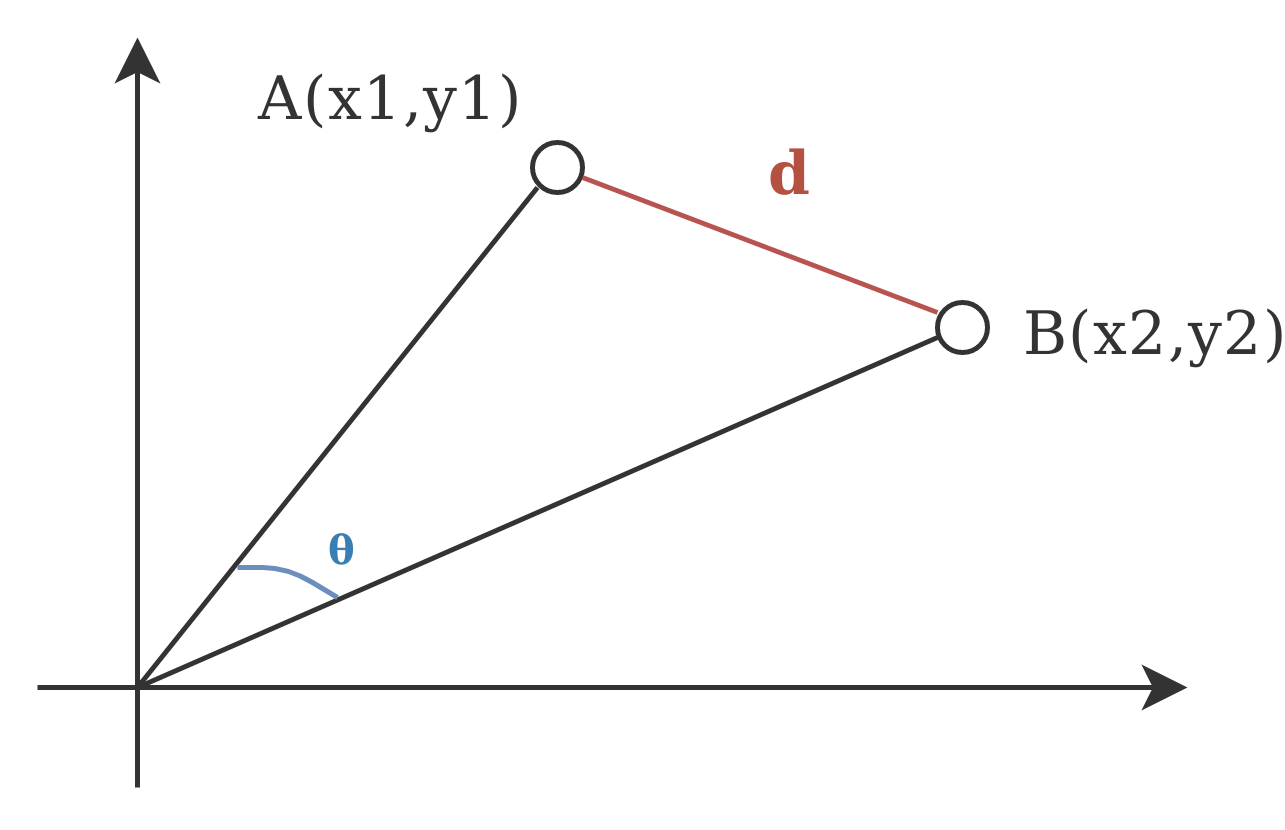
<h2 id="Creating-word-embeddings">Creating word embeddings</h2>
<p>"a word is characterized by the company it keeps" -- popularized by <em>John Rupert Firth</em></p>
<ul>
<li>A popular theory is that words are as similar as their context is</li>
<li>Word embeddings are also created with neural networks that predicts the word's context from the word itself</li>
</ul>
<p>To create word embeddings, a neural network is trained to perform the tasks. But then it is not used actually for the task it was trained it on. The goal is actually to learn the weights of the hidden layer. Then, these weights will be our vectors called "word embeddings".</p>
<p><strong>Neural Network?</strong></p>
<p>Instead of computing the actual angle, we can leave the similarity in terms of <span class="MathJax_Preview" style="color: inherit;"><span id="MJXp-Span-73" class="MJXp-math"><span id="MJXp-Span-74" class="MJXp-mi MJXp-italic">s</span><span id="MJXp-Span-75" class="MJXp-mi MJXp-italic">i</span><span id="MJXp-Span-76" class="MJXp-mi MJXp-italic">m</span><span id="MJXp-Span-77" class="MJXp-mi MJXp-italic">i</span><span id="MJXp-Span-78" class="MJXp-mi MJXp-italic">l</span><span id="MJXp-Span-79" class="MJXp-mi MJXp-italic">a</span><span id="MJXp-Span-80" class="MJXp-mi MJXp-italic">r</span><span id="MJXp-Span-81" class="MJXp-mi MJXp-italic">i</span><span id="MJXp-Span-82" class="MJXp-mi MJXp-italic">t</span><span id="MJXp-Span-83" class="MJXp-mi MJXp-italic">y</span><span id="MJXp-Span-84" class="MJXp-mo" style="margin-left: 0.333em; margin-right: 0.333em;">=</span><span id="MJXp-Span-85" class="MJXp-mi MJXp-italic">c</span><span id="MJXp-Span-86" class="MJXp-mi MJXp-italic">o</span><span id="MJXp-Span-87" class="MJXp-mi MJXp-italic">s</span><span id="MJXp-Span-88" class="MJXp-mo" style="margin-left: 0em; margin-right: 0em;">(</span><span id="MJXp-Span-89" class="MJXp-mi">Θ</span><span id="MJXp-Span-90" class="MJXp-mo" style="margin-left: 0em; margin-right: 0em;">)</span></span></span>. Formally the <a href="https://en.wikipedia.org/wiki/Cosine_similarity" target="_blank" rel="noopener">Cosine Similarity</a> <span class="MathJax_Preview" style="color: inherit;"><span id="MJXp-Span-91" class="MJXp-math"><span id="MJXp-Span-92" class="MJXp-mi MJXp-italic">s</span></span></span> between two vectors <span class="MathJax_Preview" style="color: inherit;"><span id="MJXp-Span-93" class="MJXp-math"><span id="MJXp-Span-94" class="MJXp-mi MJXp-italic">p</span></span></span> and <span class="MathJax_Preview" style="color: inherit;"><span id="MJXp-Span-95" class="MJXp-math"><span id="MJXp-Span-96" class="MJXp-mi MJXp-italic">q</span></span></span> is defined as:</p>
<p><span class="MathJax_Preview" style="color: inherit;"><span id="MJXp-Span-97" class="MJXp-math MJXp-display"><span id="MJXp-Span-98" class="MJXp-mi MJXp-italic">s</span><span id="MJXp-Span-99" class="MJXp-mo" style="margin-left: 0.333em; margin-right: 0.333em;">=</span><span id="MJXp-Span-100" class="MJXp-mfrac" style="vertical-align: 0.25em;"><span class="MJXp-box"><span id="MJXp-Span-101" class="MJXp-mi MJXp-italic">p</span><span id="MJXp-Span-102" class="MJXp-mo" style="margin-left: 0.267em; margin-right: 0.267em;">⋅</span><span id="MJXp-Span-103" class="MJXp-mi MJXp-italic">q</span></span><span class="MJXp-box" style="margin-top: -0.9em;"><span class="MJXp-denom"><span class="MJXp-box"><span id="MJXp-Span-104" class="MJXp-mrow"><span id="MJXp-Span-105" class="MJXp-mo" style="margin-left: 0.167em; margin-right: 0.167em;">|</span></span><span id="MJXp-Span-106" class="MJXp-mrow"><span id="MJXp-Span-107" class="MJXp-mo" style="margin-left: 0.167em; margin-right: 0.167em;">|</span></span><span id="MJXp-Span-108" class="MJXp-mi MJXp-italic">p</span><span id="MJXp-Span-109" class="MJXp-mrow"><span id="MJXp-Span-110" class="MJXp-mo" style="margin-left: 0.167em; margin-right: 0.167em;">|</span></span><span id="MJXp-Span-111" class="MJXp-mrow"><span id="MJXp-Span-112" class="MJXp-mo" style="margin-left: 0.167em; margin-right: 0.167em;">|</span></span><span id="MJXp-Span-113" class="MJXp-mrow"><span id="MJXp-Span-114" class="MJXp-mo" style="margin-left: 0.167em; margin-right: 0.167em;">|</span></span><span id="MJXp-Span-115" class="MJXp-mrow"><span id="MJXp-Span-116" class="MJXp-mo" style="margin-left: 0.167em; margin-right: 0.167em;">|</span></span><span id="MJXp-Span-117" class="MJXp-mi MJXp-italic">q</span><span id="MJXp-Span-118" class="MJXp-mrow"><span id="MJXp-Span-119" class="MJXp-mo" style="margin-left: 0.167em; margin-right: 0.167em;">|</span></span><span id="MJXp-Span-120" class="MJXp-mrow"><span id="MJXp-Span-121" class="MJXp-mo" style="margin-left: 0.167em; margin-right: 0.167em;">|</span></span></span></span></span></span><span id="MJXp-Span-122" class="MJXp-mo" style="margin-left: 0em; margin-right: 0.222em;">,</span><span id="MJXp-Span-123" class="MJXp-mrow"><span id="MJXp-Span-124" class="MJXp-mtext"> where </span></span><span id="MJXp-Span-125" class="MJXp-mi MJXp-italic">s</span><span id="MJXp-Span-126" class="MJXp-mo" style="margin-left: 0.333em; margin-right: 0.333em;">∈</span><span id="MJXp-Span-127" class="MJXp-mo" style="margin-left: 0em; margin-right: 0em;">[</span><span id="MJXp-Span-128" class="MJXp-mo" style="margin-left: 0.267em; margin-right: 0.267em;">−</span><span id="MJXp-Span-129" class="MJXp-mn">1</span><span id="MJXp-Span-130" class="MJXp-mo" style="margin-left: 0em; margin-right: 0.222em;">,</span><span id="MJXp-Span-131" class="MJXp-mn">1</span><span id="MJXp-Span-132" class="MJXp-mo" style="margin-left: 0em; margin-right: 0em;">]</span></span></span></p>
## Creating word embeddings 2
Word embeddings are learned with neural networks. The target can be:
- Tries to predict the word given the context - The Continous Bag Of Words model (CBOW)
- Tries to predict the context given a words - The SkipGram model
The training examples are generated from big text corpora. For example from the sentence “The quick brown fox jumps over the lazy dog.” we can generate the following inputs:
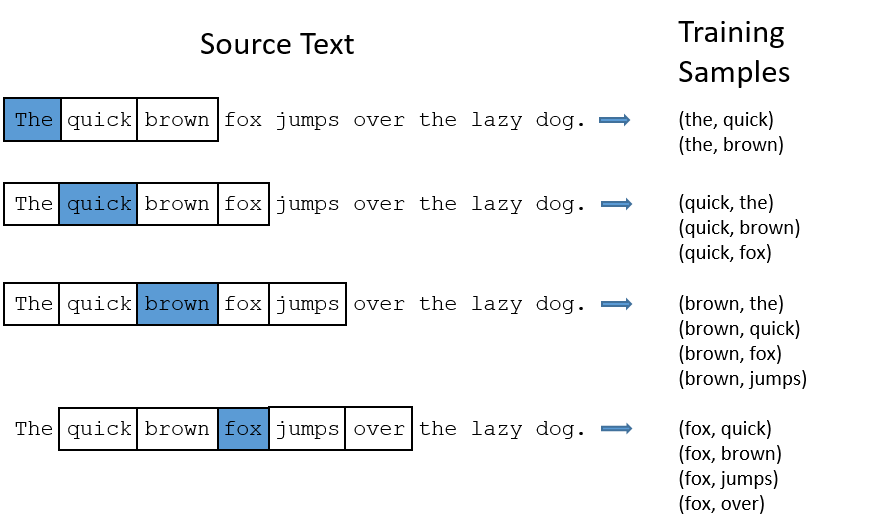
To do this, we first build a vocabulary of words from our training documents–let’s say we have a vocabulary of 10,000 unique words. The vocabulary of big corporas can be much more then 10,000 unique words, to handle them we usually substitute rare words with a special token (this is usually the _UNK_ token).
First we build the vocabulary of our documents, then for representing words, we will use one-hot vectors. The output of the network will be a single vector that contains the probabilities for the "nearby" words.
### Famous static word embeddings for English
- [Word2vec](https://arxiv.org/pdf/1301.3781.pdf)
- [GLOVE](https://nlp.stanford.edu/projects/glove/)
### Contextual embeddings?
- [Elmo](https://allennlp.org/elmo)
- [BERT](https://arxiv.org/abs/1810.04805)
- [Flair](https://www.aclweb.org/anthology/N19-4010/)
For static embeddings, we will use a GLOVE embedding of 100 dimensional vectors trained on 6B tokens.
[Download GLOVE](http://sandbox.hlt.bme.hu/~adaamko/glove.6B.100d.txt)
# Using word embeddings
As we discussed, more recently prediction-based word vectors have demonstrated better performance, such as word2vec and GloVe (which also utilizes the benefit of counts). Here, we shall explore the embeddings produced by GloVe. If you want to know more about embeddings, try reading [GloVe's original paper](https://nlp.stanford.edu/pubs/glove.pdf).
```
embedding_file = "glove.6B.100d.txt"
embedding = gensim.models.KeyedVectors.load_word2vec_format(embedding_file, binary=False)
dog_vector = embedding["dog"]
dog_vector.shape
embedding.most_similar("president")
embedding.most_similar(positive=['woman', 'king'], negative=['man'])
embedding.similarity("woman", "computer")
from sklearn.manifold import TSNE
def tsne_plot(model, size=500):
"Creates and TSNE model and plots it"
labels = []
tokens = []
for i, word in enumerate(model.wv.vocab):
if len(tokens) > size:
break
tokens.append(model[word])
labels.append(word)
tsne_model = TSNE(perplexity=40, n_components=2, init='pca', n_iter=2500, random_state=23)
new_values = tsne_model.fit_transform(tokens)
x = []
y = []
for value in new_values:
x.append(value[0])
y.append(value[1])
for i in range(len(x)):
plt.scatter(x[i],y[i])
plt.annotate(labels[i],
xy=(x[i], y[i]),
xytext=(5, 2),
textcoords='offset points',
ha='right',
va='bottom')
plt.show()
tsne_plot(embedding, 100)
```
## Analogies
```
def analogy(word1, word2, word3, n=5):
#get vectors for each word
word1_vector = embedding[word1]
word2_vector = embedding[word2]
word3_vector = embedding[word3]
#calculate analogy vector
analogy_vector = embedding.most_similar(positive=[word3, word2], negative=[word1])
print(word1 + " is to " + word2 + " as " + word3 + " is to...")
return analogy_vector
analogy('man', 'king', 'woman')
```
## Spacy also has pretrained embeddings!
```
from sklearn.metrics.pairwise import cosine_similarity
doc = nlp("man woman")
cosine_similarity(doc[0].vector.reshape(1, -1), doc[1].vector.reshape(1, -1))
```
__Or with using the built in similarity function:__
```
nlp("My name is adam").similarity(nlp("My name is andrea"))
```
## Contextual embeddings
In GloVe and Word2vec representations, words have a static representation. But words can have different meaning in different contexts, e.g. the word "stick":
1. Find some dry sticks and we'll make a campfire.
2. Let's stick with glove embeddings.

_(Peters et. al., 2018 in the ELMo paper)_
```
# The sentence objects holds a sentence that we may want to embed or tag
from flair.data import Sentence
from flair.embeddings import FlairEmbeddings
# init embedding
flair_embedding_forward = FlairEmbeddings('news-forward')
# create a sentence
sentence1 = Sentence("Find some dry sticks and we'll make a campfire.")
sentence2 = Sentence("Let's stick with glove embeddings.")
# embed words in sentence
flair_embedding_forward.embed(sentence2)
for token in sentence2:
print(token)
print(token.embedding)
```
In Flair, a pretrained NER tagger is also available for use
| github_jupyter |
## Imports and Data Loading
```
# Imports
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib
import matplotlib.pyplot as plt
from scipy.stats import skew
from scipy.stats.stats import pearsonr
%config InlineBackend.figure_format = 'retina' #set 'png' here when working on notebook
%matplotlib inline
# Sklearn imports
from sklearn.model_selection import cross_val_score
from sklearn.linear_model import Ridge, RidgeCV, ElasticNet, LassoCV, LassoLarsCV
# Read the train and the test data
train = pd.read_csv("./data/train.csv")
test = pd.read_csv("./data/test.csv")
# Shape of the data we are dealing with
print(train.shape)
print(test.shape)
all_data = pd.concat((train.loc[:,'MSSubClass':'SaleCondition'],
test.loc[:,'MSSubClass':'SaleCondition']))
all_data.shape
```
## Data Preprocessing
- First I'll transform the skewed numeric features by taking log(feature + 1) - this will make the features more normal
- Create Dummy variables for the categorical features
- Replace the numeric missing values (NaN's) with the mean of their respective columns
```
# Experimental purpose
prices = pd.DataFrame({"price":train["SalePrice"], "log(price + 1)":np.log1p(train["SalePrice"])})
# Prior being normalized
sns.distplot(prices['price'])
# After being normalized, a more symmetric skew
sns.distplot(prices['log(price + 1)'])
```
### Step 1: Log transform the numeric features
```
# Step 1: Log transform the numeric features
train["SalePrice"] = np.log1p(train["SalePrice"])
# Log transform skewed numeric features:
numeric_feats = all_data.dtypes[all_data.dtypes != "object"].index
skewed_feats = train[numeric_feats].apply(lambda x: skew(x.dropna())) # Compute skewness
skewed_feats = skewed_feats[skewed_feats > 0.75] # Filter out index's where skew is high
skewed_feats = skewed_feats.index
all_data[skewed_feats] = np.log1p(all_data[skewed_feats]) # Perform log1p on all the remaining features
print(skewed_feats)
all_data.head()
```
### Step 2: One Hot Encoding categorical values
```
# Step 2:One Hot Encoding categorical values
all_data = pd.get_dummies(all_data)
all_data.head()
```
### Step 3: filling NaN's with the mean of the column
```
all_data = all_data.fillna(all_data.mean())
```
## Models
### Ridge model
```
# Creating matrices for sklearn, bringing back data to it's original shape
X_train = all_data[:train.shape[0]]
X_test = all_data[train.shape[0]:]
y = train.SalePrice
```
Now running regularized linear regression models from the scikit learn module. We'll be trying both l_1(Lasso) and l_2(Ridge) regularization.
Defining a function that returns the cross-validation rmse error so we can evaluate our models and pick the best tuning parameters
```
# General fun to return rmse score of the model
def rmse_cv(model):
"""
:param: model = model for evaluatio
:return: rmse = list of all cros-eval scores
"""
rmse= np.sqrt(-cross_val_score(model, X_train, y, scoring="neg_mean_squared_error", cv = 5))
return(rmse)
```
The main tuning parameter for the Ridge model is alpha - a regularization parameter that measures how flexible our model is. The higher the regularization the less prone our model will be to overfit. However it will also lose flexibility and might not capture all of the signal in the data.
```
model_ridge = Ridge()
alphas = [0.05, 0.1, 0.3, 1, 3, 5, 10, 15, 30, 50, 75]
cv_ridge = [rmse_cv(Ridge(alpha = alpha)).mean()
for alpha in alphas]
# RMSE scores
print(cv_ridge)
# Plotting the RMSE Score
cv_res = pd.DataFrame({"cv_ridge": cv_ridge, "alphas": alphas})
sns.lineplot(data=cv_res, x="alphas", y="cv_ridge")
```
Note the U-ish shaped curve above. When alpha is too large the regularization is too strong and the model cannot capture all the complexities in the data. If however we let the model be too flexible (alpha small) the model begins to overfit.
alpha = 10 is about right based on the plot above
```
min(cv_ridge)
```
### Lasso model
```
model_lasso = LassoCV(alphas = [1, 0.1, 0.001, 0.0005]).fit(X_train, y)
rmse_cv(model_lasso).mean()
```
The **lasso performs** even better so we'll just use this one to predict on the test set. Another neat thing about the Lasso is that it does feature selection for you - setting coefficients of features it deems unimportant to zero. Let's take a look at the coefficients:
```
coef = pd.Series(model_lasso.coef_, index = X_train.columns)
print("Lasso picked " + str(sum(coef != 0)) + " variables and eliminated the other " + str(sum(coef == 0)) + " variables")
# Having a look at the most imporant features
imp_coef = pd.concat([coef.sort_values().head(10),
coef.sort_values().tail(10)])
print(imp_coef)
matplotlib.rcParams['figure.figsize'] = (8.0, 10.0)
imp_coef.plot(kind = "barh")
plt.title("Coefficients in the Lasso Model")
```
The most important positive feature is GrLivArea - the above ground area by area square feet. This definitely sense. Then a few other location and quality features contributed positively. Some of the negative features make less sense and would be worth looking into more - it seems like they might come from unbalanced categorical variables.
Also note that unlike the feature importance you'd get from a random forest these are actual coefficients in your model - so you can say precisely why the predicted price is what it is. The only issue here is that we log_transformed both the target and the numeric features so the actual magnitudes are a bit hard to interpret.
```
#let's look at the residuals as well:
matplotlib.rcParams['figure.figsize'] = (6.0, 6.0)
preds = pd.DataFrame({"preds":model_lasso.predict(X_train), "true":y})
preds["residuals"] = preds["true"] - preds["preds"]
preds.plot(x = "preds", y = "residuals",kind = "scatter")
```
### XGboost Model
For XGBoost Please look at this [kaggle article](https://www.kaggle.com/c/house-prices-advanced-regression-techniques/notebooks?competitionId=5407&sortBy=voteCount)
## Keras
```
from keras.layers import Dense
from keras.models import Sequential
from keras.regularizers import l1
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import train_test_split
X_train = StandardScaler().fit_transform(X_train)
X_tr, X_val, y_tr, y_val = train_test_split(X_train, y, random_state = 3)
X_tr.shape
X_tr
model = Sequential()
#model.add(Dense(256, activation="relu", input_dim = X_train.shape[1]))
model.add(Dense(1, input_dim = X_train.shape[1], W_regularizer=l1(0.001)))
model.compile(loss = "mse", optimizer = "adam")
```
## Resources
[Stacked Regressions to predict House Prices](https://www.kaggle.com/serigne/stacked-regressions-top-4-on-leaderboard)
| github_jupyter |
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.preprocessing import MinMaxScaler
from sklearn.preprocessing import OneHotEncoder
import keras
from keras import backend as K
from keras.models import Sequential
from keras.layers import Activation
from keras.layers.core import Dense
from keras.optimizers import Adam
from keras.metrics import categorical_crossentropy
from keras.utils import to_categorical
from keras import metrics
# global
filtered_cols = ['MEAN_RR', 'HR', 'SDRR', 'pNN50', 'LF', 'HF', 'condition']
input_shape = len(filtered_cols) - 1
# SWELL-Knowledge Worker Dataset
train_file = pd.read_csv('stress_train.csv').drop(columns="datasetId")
# relevant features
train_file = train_file[filtered_cols]
train_file
train_file.describe()
plt.figure()
cor = train_file.corr()
sns.heatmap(cor, annot=True, cmap=plt.cm.Reds)
plt.show()
cor_target = abs(cor["HF"])
relevant_features = cor_target[cor_target>0.5]
relevant_features
test_file = pd.read_csv('stress_test.csv').drop(columns="datasetId")
test_file = test_file[filtered_cols]
test_file.describe()
# train
train_samples = train_file.drop(columns='condition').to_numpy()
train_labels = train_file['condition'].to_numpy()
# test
test_samples = test_file.drop(columns='condition').to_numpy()
test_labels = test_file['condition'].to_numpy()
# normalize features
scaler = MinMaxScaler(feature_range=(0,1))
train_samples = scaler.fit_transform(train_samples)
test_samples = scaler.fit_transform(test_samples)
# one-hot-encode labels
one_hot_encoder = OneHotEncoder(categories='auto')
train_labels = one_hot_encoder.fit_transform(train_labels.reshape(-1, 1)).toarray()
test_labels = one_hot_encoder.fit_transform(test_labels.reshape(-1, 1)).toarray()
# build the model
model = Sequential([
Dense(input_shape, input_shape=[input_shape], activation='relu'),
Dense(100, activation='relu'),
Dense(100, activation='relu'),
Dense(100, activation='relu'),
Dense(100, activation='relu'),
Dense(100, activation='relu'),
Dense(3, activation='softmax')
])
model.summary()
model.compile(Adam(lr=.0001),
loss='categorical_crossentropy',
metrics=['accuracy'])
history = model.fit(train_samples, train_labels, validation_split=0.1, batch_size=100, epochs=150, shuffle=True, verbose=2)
model.save('model_stress.h5')
plt.plot(history.history['accuracy'])
plt.plot(history.history['val_accuracy'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'val'], loc='upper left')
plt.show()
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'val'], loc='upper left')
plt.show()
```
| github_jupyter |
# Monte Carlo Example
[](https://github.com/eabarnes1010/course_objective_analysis/tree/main/code)
[](https://colab.research.google.com/github/eabarnes1010/course_objective_analysis/blob/main/code/monte_carlo_example.ipynb)
An illustration of using Monte Carlo techniques.
#### The problem
This example is based on the "Monte Carlo simulation" example in Chapter 1 of the lecture notes. The idea is as follows...
In January (31 days), the maximum daily temperature was 2.2 standard deviations from the climatological mean temperature. If we assume that daily temperatures are normally distributed, how rare is it to have a maximum of 2.2$\sigma$ or greater in 31 daily samples?
#### The approach
While we have lots of nice statistical tests for the mean, and even the variance, we've ventured outside of this into the realm of maxima! So what do we do? Since the maximum is a pretty common quantity, there are in fact tests we could use. However, we could also take a Monte Carlo approach (which is what we will do given the title of this example) to solve our problem.
The idea is that since we _know_ the underlying population is normal, we can just make some "fake" data and play around with it to find out how rare a 2.2$\sigma$ maximum really is.
The first step, of course, is to import our packages.
```
#.............................................
# IMPORT STATEMENTS
#.............................................
import numpy as np
import matplotlib.pyplot as plt
import scipy.stats as stats
#.............................................
# PLOTTING COMMANDS
#.............................................
import matplotlib as mpl
# set figure defaults
mpl.rcParams['figure.dpi'] = 150
plt.rcParams['figure.figsize'] = (12.0/2, 8.0/2)
```
Setup our variables. The "drawn_max" variable is where we set the maximum we actually got in our 31 samples, and "sample_length" sets the number of values in our sample (i.e. N).
```
drawn_max = 2.2
sample_length = 31
```
Next, we are going to make a long list of values drawn from a random normal distribution (call this data set "Z"). You can think of this as our _popoulation bucket_. We will also initialize an empty array "M" where we will ultimately store the maximum values from many many different samples of length N=31.
```
Z = np.random.randn(1_000_000,1)
M = np.empty([100_000, 1])
```
Next we do the heavy lifting. The idea is that since we know our underlying data is normal, we are going to randomly grab N=31 values from this huge list (Z), calculate the maximum in the sample, save it, then rinse and repeat. We will do this 10,000,000 times.
```
for iloop in range(np.size(M)):
ip = np.random.randint(low=0,high=Z.shape[0],size=sample_length)
M[iloop] = np.max(Z[ip])
```
Ok - so what do we have now? Well, we now have 100,000 maximum values from samples of N=31. So, we can plot the distribution of these maxima (M), and see what they look like! We can then see how rare our actual maximum that we actually got (i.e. drawn_max = 2.2) is. Check it out.
```
plt.figure()
bin_width = .1
bin_list = np.arange(-1,10,bin_width)
n, bins = np.histogram(M, bins=bin_list, density=False)
plt.plot([drawn_max, drawn_max],[0, .12],color='red',linewidth=3,linestyle='--', label='actual Jan. maximum')
plt.bar(bins[0:-1],n/float(len(M)),bin_width, facecolor='blue', alpha=0.4)
plt.plot(bins[0:-1]+bin_width/2,n/float(len(M)),color='blue', label='random samples')
plt.xlabel('maximum value')
plt.ylabel('frequency')
titlename = 'Maximum value for sample of length N = ' + str(sample_length)
plt.title(titlename)
plt.legend(fontsize=8)
plt.show()
```
The blue shading and line denotes the frequency of the maxima we got from the 100,000 samples of length N=31 (drawn from a normal don't forget). The red line denotes what we _actually_ got from our real data. How rare is it? Well, just looking at the plot it is pretty clear that a maximum of 2.2 is not rare at all. Of course, we can calculate the number of values in M above 2.2 to see the probability.
```
100.*float(sum(M>drawn_max)/np.size(M))
```
So, 35% of the 100,000 samples actually had maxima of 2.2 or bigger! So truly, not rare at all.
| github_jupyter |
<a href="https://colab.research.google.com/github/jeffheaton/t81_558_deep_learning/blob/master/manual_setup.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# T81-558: Applications of Deep Neural Networks
**Manual Python Setup**
* Instructor: [Jeff Heaton](https://sites.wustl.edu/jeffheaton/), McKelvey School of Engineering, [Washington University in St. Louis](https://engineering.wustl.edu/Programs/Pages/default.aspx)
* For more information visit the [class website](https://sites.wustl.edu/jeffheaton/t81-558/).
# Software Installation
This class is technically oriented. A successful student needs to be able to compile and execute Python code that makes use of TensorFlow for deep learning. There are two options for you to accomplish this:
* Install Python, TensorFlow and some IDE (Jupyter, TensorFlow, and others)
* Use Google CoLab in the cloud
## Installing Python and TensorFlow
It is possible to install and run Python/TensorFlow entirely from your computer. Google provides TensorFlow for Windows, Mac, and Linux. Previously, TensorFlow did not support Windows. However, as of December 2016, TensorFlow supports Windows for both CPU and GPU operation.
The first step is to install Python 3.7. As of August 2019, this is the latest version of Python 3. I recommend using the Miniconda (Anaconda) release of Python, as it already includes many of the data science related packages that are needed by this class. Anaconda directly supports Windows, Mac, and Linux. Miniconda is the minimal set of features from the extensive Anaconda Python distribution. Download Miniconda from the following URL:
* [Miniconda](https://docs.conda.io/en/latest/miniconda.html)
# Dealing with TensorFlow incompatibility with Python 3.7
*Note: I will remove this section once all needed libraries add support for Python 3.7.
**VERY IMPORTANT** Once Miniconda has been downloaded you must create a Python 3.6 environment. Not all TensorFlow 2.0 packages currently (as of August 2019) support Python 3.7. This is not unusual, usually you will need to stay one version back from the latest Python to maximize compatibility with common machine learning packages. So you must execute the following commands:
```
conda create -y --name tensorflow python=3.6
```
To enter this environment, you must use the following command (**for Windows**), this command must be done every time you open a new Anaconda/Miniconda terminal window:
```
activate tensorflow
```
For **Mac**, do this:
```
source activate tensorflow
```
# Installing Jupyter
it is easy to install Jupyter notebooks with the following command:
```
conda install -y jupyter
```
Once Jupyter is installed, it is started with the following command:
```
jupyter notebook
```
The following packages are needed for this course:
```
conda install -y scipy
pip install --exists-action i --upgrade sklearn
pip install --exists-action i --upgrade pandas
pip install --exists-action i --upgrade pandas-datareader
pip install --exists-action i --upgrade matplotlib
pip install --exists-action i --upgrade pillow
pip install --exists-action i --upgrade tqdm
pip install --exists-action i --upgrade requests
pip install --exists-action i --upgrade h5py
pip install --exists-action i --upgrade pyyaml
pip install --exists-action i --upgrade tensorflow_hub
pip install --exists-action i --upgrade bayesian-optimization
pip install --exists-action i --upgrade spacy
pip install --exists-action i --upgrade gensim
pip install --exists-action i --upgrade flask
pip install --exists-action i --upgrade boto3
pip install --exists-action i --upgrade gym
pip install --exists-action i --upgrade tensorflow==2.0.0-beta1
pip install --exists-action i --upgrade keras-rl2 --user
conda update -y --all
```
Notice that I am installing as specific version of TensorFlow. As of the current semester, this is the latest version of TensorFlow. It is very likely that Google will upgrade this during this semester. The newer version may have some incompatibilities, so it is important that we start with this version and end with the same.
You should also link your new **tensorflow** environment to Jupyter so that you can choose it as a Kernal. Always make sure to run your Jupyter notebooks from your 3.6 kernel. This is demonstrated in the video.
```
python -m ipykernel install --user --name tensorflow --display-name "Python 3.6 (tensorflow)"
```
```
# What version of Python do you have?
import sys
import tensorflow.keras
import pandas as pd
import sklearn as sk
import tensorflow as tf
print(f"Tensor Flow Version: {tf.__version__}")
print(f"Keras Version: {tensorflow.keras.__version__}")
print()
print(f"Python {sys.version}")
print(f"Pandas {pd.__version__}")
print(f"Scikit-Learn {sk.__version__}")
print("GPU is", "available" if tf.test.is_gpu_available() else "NOT AVAILABLE")
```
| github_jupyter |
### Imports and Version Check
```
import numpy as np
import tensorflow as tf
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Conv2D, MaxPooling2D
from tensorflow.keras.layers import Activation, Dropout, Flatten, Dense
from tensorflow.keras.models import Model
from tensorflow.keras import backend as K
from tensorflow.keras.optimizers import SGD
from tensorflow.keras.layers import Dense, GlobalAveragePooling2D, Input
from tensorflow.keras.applications.inception_v3 import InceptionV3
from tensorflow.keras.applications.resnet50 import ResNet50, preprocess_input
from tensorflow.keras.applications import MobileNet
from tensorflow.keras.preprocessing.image import (ImageDataGenerator, Iterator,
array_to_img, img_to_array, load_img)
from tensorflow.keras.callbacks import ModelCheckpoint, TensorBoard, EarlyStopping
from tensorflow.keras.optimizers import SGD, Adam
from tensorflow.keras import regularizers
from tensorflow.keras import layers
import sys, os
import matplotlib.pyplot as plt
from sklearn import metrics
import tensorflow_model_optimization as tfmot
import shap
import keras
import matplotlib.cm as cm
from IPython.display import Image
import pandas as pd
import seaborn as sns
# os.environ["CUDA_DEVICE_ORDER"] = "PCI_BUS_ID" # see issue #152
# os.environ["CUDA_VISIBLE_DEVICES"] = "0"
# config = tf.compat.v1.ConfigProto()
# config.gpu_options.allow_growth = True
# sess = tf.compat.v1.Session(config=config)
# sess.as_default()
# physical_devices = tf.config.experimental.list_physical_devices('GPU')
# for physical_device in physical_devices:
# tf.config.experimental.set_memory_growth(physical_device, True)
tf.__version__
%matplotlib inline
# os.environ['TF_FORCE_GPU_ALLOW_GROWTH'] = 'true'
```
### Data Generator
```
train_data_dir = '../data/CollisionData/'
img_width, img_height = 96, 96 # 224, 224
nb_train_samples = 730
nb_validation_samples = 181
epochs = 25
batch_size = 16
if K.image_data_format() == 'channels_first':
input_shape = (3, img_width, img_height)
else:
input_shape = (img_width, img_height, 3)
train_datagen = ImageDataGenerator(rescale=1./255,
shear_range=0.2,
zoom_range=0.2,
horizontal_flip=True,
) # set validation split
#### train with 100% of data. See other notebooks for validation of the models
train_generator = train_datagen.flow_from_directory(
train_data_dir,
target_size=(img_height, img_width),
batch_size=batch_size,
class_mode='binary') # set as training data
```
## Model Architectures
```
models = [] # can be used for multiple models, for this we are using a single model
```
### Custom CNN
Tiny Compatible
```
finetune_model_CNN = tf.keras.Sequential([
layers.InputLayer(input_shape=input_shape),
layers.BatchNormalization(),
layers.Conv2D(filters=16, kernel_size=5, padding='same', activation='relu'),
layers.MaxPool2D(),
layers.BatchNormalization(),
layers.Conv2D(32, 3, padding='same', activation='relu'),
layers.Conv2D(32, 3, padding='same', activation='relu'),
layers.MaxPool2D(),
layers.BatchNormalization(),
layers.Conv2D(64, 3, padding='same', activation='relu'),
layers.Conv2D(64, 3, padding='same', activation='relu'),
layers.MaxPool2D(),
layers.Flatten(),
layers.Dense(128, activation='relu'),
layers.Dropout(0.5),
layers.Dense(64, activation='relu'),
layers.Dropout(0.5),
layers.Dense(1, activation='sigmoid')
])
finetune_model_CNN._name="Custom_CNN"
finetune_model_CNN.compile(
optimizer=tf.keras.optimizers.Adam(lr = 1e-5, decay = 1e-5),
loss='binary_crossentropy',
metrics=['accuracy']
)
models.append(finetune_model_CNN)
finetune_model_CNN.summary()
```
## Training and Testing
```
callbacks = [
tf.keras.callbacks.EarlyStopping(
monitor='val_loss', patience = 15,
min_delta=0.001, restore_best_weights=True
)
]
history = []
for each in models:
print("="*40)
print("Training and Testing Model: %s" % str(each.name))
temp_history = each.fit(train_generator,
steps_per_epoch = nb_train_samples // batch_size,
epochs = epochs,
validation_steps = nb_validation_samples // batch_size, shuffle=True, callbacks = callbacks)
print("="*40)
history.append(temp_history)
```
### Model Summary
#### Number of Parameters
```
for each in models:
print("Number of parameters: %s" % str(each.name))
print(each.count_params())
```
#### Number of Ops
```
def get_flops(model_h5_path):
session = tf.compat.v1.Session()
graph = tf.compat.v1.get_default_graph()
with graph.as_default():
with session.as_default():
model = tf.keras.models.load_model(model_h5_path)
run_meta = tf.compat.v1.RunMetadata()
opts = tf.compat.v1.profiler.ProfileOptionBuilder.float_operation()
flops = tf.compat.v1.profiler.profile(graph=graph,
run_meta=run_meta, cmd='op', options=opts)
tf.compat.v1.reset_default_graph()
return flops.total_float_ops
for each in models:
model_file = '../my-log-dir/saved_model/' + str(each.name) + '.h5'
print("Number of OPS: %s" % str(each.name))
print(get_flops(model_file))
```
#### Model Size
```
import os
for each in models:
model_file = '../my-log-dir/saved_model/' + str(each.name) + '.h5'
print("Model: %s" % str(each.name))
b = os.path.getsize(model_file)
print ("Size(mb): %d" % (b/1000000))
```
# Optimization
## Pruning
```
model_file = '../my-log-dir/saved_model/Custom_CNN.h5'
model = tf.keras.models.load_model(model_file)
end_step = np.ceil(1.0 * nb_train_samples / batch_size).astype(np.int32) * epochs
pruning_schedule = tfmot.sparsity.keras.PolynomialDecay(initial_sparsity=0.50,
final_sparsity=0.90,
begin_step=0,
end_step=end_step,
frequency=100)
model_for_pruning = tfmot.sparsity.keras.prune_low_magnitude(model, pruning_schedule=pruning_schedule)
model_for_pruning.summary()
logdir = '../my-log-dir/'
callbacks = [
tfmot.sparsity.keras.UpdatePruningStep(),
tfmot.sparsity.keras.PruningSummaries(log_dir=logdir),
]
model_for_pruning.compile(
optimizer='adam',
loss='binary_crossentropy',
metrics=['accuracy']
)
model_for_pruning.fit(train_generator,
steps_per_epoch = nb_train_samples // batch_size,
epochs = epochs,
validation_steps = nb_validation_samples // batch_size, shuffle=True, callbacks = callbacks)
```
#### Export Pruned Model
```
from tensorflow_model_optimization.sparsity import keras as sparsity
final_model = sparsity.strip_pruning(model_for_pruning)
final_model.summary()
from tensorflow.keras.models import load_model
import numpy as np
# model = tf.keras.models.load_model(final_model)
for i, w in enumerate(final_model.get_weights()):
print(
"{} -- Total:{}, Zeros: {:.2f}%".format(
final_model.weights[i].name, w.size, np.sum(w == 0) / w.size * 100
)
)
```
##### Pruned Model Size and Store
```
import tempfile
import zipfile
# _, new_pruned_keras_file = tempfile.mkstemp(".h5")
new_pruned_keras_file = "../my-log-dir/saved_model/pruned_model.pb"
print("Saving pruned model to: ", new_pruned_keras_file)
tf.keras.models.save_model(final_model, new_pruned_keras_file, include_optimizer=False)
print(
"Size of the pruned model: %.2f Mb"
% (os.path.getsize(new_pruned_keras_file) / float(2 ** 20))
)
```
## Quantize
#### Post training quantization
##### Full integer quantization.
To 8-bits
```
keras_model = tf.keras.models.load_model(new_pruned_keras_file)
tflite_fullint_model_file = "../my-log-dir/saved_model/post_fullint_quantized.tflite"
# converter = tf.lite.TFLiteConverter.from_saved_model('../log/saved_model/pruned_model.pb')
converter = tf.lite.TFLiteConverter.from_keras_model(final_model)
converter.optimizations = [tf.lite.Optimize.DEFAULT]
num_calibration_steps = 1
def representative_dataset_gen():
for _ in range(num_calibration_steps):
# Get sample input data as a numpy array in a method of your choosing.
## Ideally we should do a validation calibration but we are using all of the training data for max acc
# x =np.concatenate([validation_generator.next()[0] for i in range(validation_generator.__len__())])
x =np.concatenate([train_generator.next()[0] for i in range(train_generator.__len__())])
print(x.shape)
yield [x]
converter.representative_dataset = representative_dataset_gen
converter.target_spec.supported_ops = [tf.lite.OpsSet.TFLITE_BUILTINS_INT8]
converter.inference_input_type = tf.int8 # or tf.uint8
converter.inference_output_type = tf.int8 # or tf.uint8
tflite_model = converter.convert()
with open(tflite_fullint_model_file, "wb") as f:
f.write(tflite_model)
```
### Output Files
`pruned_model.pb` - Pruned model file (file size is not going to reduce but the sparsity is incorporated)
`post_fullint_quantized.tflite` - Integer 8-bit quantized model
## Quantized Model to OpenMV
Here we show the MicroPython code that uses the `post_fullint_quantized.tflite` file to perform inferencing on the device (OpenMV Cam H7+).
```
# %load CollisionCam.py
# Dune Collision Camera
# written by EB Goldstein and SD Mohanty
# started 10/2020
# last revision 4/2021
# import what we need
import pyb, sensor, image, time, os, tf, random
#setup LEDs and set into known off state
red_led = pyb.LED(1)
green_led = pyb.LED(2)
red_led.off()
green_led.off()
#red light during setup
red_led.on()
# get sensor set up
sensor.reset() # Reset & initialize sensor
sensor.set_pixformat(sensor.RGB565) # Set pixel format to RGB
sensor.set_framesize((sensor.QVGA)) # Set frame size to QVGA (320x240)
sensor.set_windowing((240,240)) # Set window to 240x240
sensor.skip_frames(time=2000) # Let the camera adjust.
#Load the TFlite model and the labels
net = tf.load('/post_quantized_full_int.tflite', load_to_fb=True)
labels = ['collision', 'no collision']
#turn red off when model is loaded
red_led.off()
#MAIN LOOP
# loop needs to do a few things:
# x-take a picture
# x-record the picture on sd card
# x-do the inference
# x-record the inference in a db w/ rand as name
# x-blink LED for field debugging
# x-delay
while(True):
#toggle LED for visual indication that script is running
green_led.toggle()
#get the image/take the picture
img = sensor.snapshot()
#Do the classification and get the object returned by the inference.
TF_objs = net.classify(img)
#The object has a output, which is a list of classifcation scores
#for each of the output channels. this model only has 1 (Collision).
#So now we extract that float value and print to the serial terminal.
collision_score = TF_objs[0].output()[0]
print("Collision = %f" % collision_score)
#we don;t have an RTC attached now, so we save the images and the
#collision scores with names according to a random bit stream.
#generate random bits for stream of names
rand_label = str(random.getrandbits(30))
#save image on camera (bit as name)
img.save("./imgs/" + rand_label + ".jpg")
#save inference (bit as name, and score) to a file
with open("./inference.txt", 'a') as file:
file.write(rand_label + "," + str(collision_score) + "\n")
#wait some number of milliseconds
pyb.delay(1000)
```
| github_jupyter |
## Exploring the impact of double-scoring on PRMSE
In previous notebooks, we have seen that in order to compute PRMSE, we need the responses to have scores from two human raters. However, it may not be practical to have every single repsonses double-scored. In this notebook, we examine how PRMSE depends on the percentage/number of double-scored responses that may be available in the dataset.
To do this, we randomly choose a fixed number of rater pairs in each rater category, then simulate scenarios with a different percentage of double-scored responses for each of those rater pairs, and compute PRMSE for a pre-determined in each scenario.
```
import itertools
import json
import pandas as pd
import numpy as np
import seaborn as sns
from matplotlib import pyplot as plt
from pathlib import Path
from rsmtool.utils.prmse import prmse_true
from simulation.dataset import Dataset
from simulation.utils import (compute_agreement_one_system_one_rater_pair,
get_rater_pairs,
simulate_percent_double_scored)
from sklearn.metrics import r2_score
```
### Step 1: Setup
To set up the experiment, we first load the dataset we have already created and saved in the `making_a_dataset.ipynb` notebook and use that for this experiment.
For convenience and replicability, we have pre-computed many of the parameters that are used in our notebooks and saved them in the file `settings.json`. We load this file below.
```
# load the dataset file
dataset = Dataset.from_file('../data/default.dataset')
# let's remind ourselves what the dataset looks like
print(dataset)
# load the experimental settings file
experiment_settings = json.load(open('settings.json', 'r'))
# now get the data frames for our loaded dataset
df_scores, df_rater_metadata, df_system_metadata = dataset.to_frames()
# get the ID of the simulated system that we have chosen
# as the source of our single automated score
chosen_system = experiment_settings['sample_system']
# get the number of fixed human-rater pairs per category that we want to use
rater_pairs_per_category = experiment_settings['rater_pairs_per_category']
# get the various percentages of double-scored responses we want to simulate
double_scored_percentages = experiment_settings['double_scored_percentages']
# now let's get the pre-determined number of randomly-sampled pairs of raters in each rater category
rater_category_pairs = df_rater_metadata.groupby('rater_category').apply(lambda row: get_rater_pairs(row['rater_id'], rater_pairs_per_category))
```
### Step 2: Simulate various percentages of double-scored responses
Now, for each chosen pair of simulated raters, we randomly mask out a fixed percentage of the second rater's scores, and then compute the PRMSE (and other conventional agreement metrics) for the chosen system against this modified set of rater scores. Another thing worth checking is whether it's the _percentage_ of double-scored responses that matters or simply the _number_ of double-scored responses in the dataset. To do this, we also compute a second PRMSE value over only the double-scored repsonses available in each case. For example, if we are simulating the scenario where we only have 10% of the responses double-scored, then we compute two PRMSE values: (a) over the full dataset with 10% double-scores and 90% single-scored responses and (b) over a smaller dataset that has only the 10% double-scored responses.
Note that since we are computing a number of metrics for a dataset of 10,000 responses multiple times in the cell below, it will take a while.
```
# create a chained iterator over all of the chosen pairs
# so that we can iterate over all 200 of them at once
rater_pairs = list(itertools.chain.from_iterable(rater_category_pairs.values))
# initialize a list that will hold the information for each rater pair x percentavge
metric_values = []
# iterate over the cross-product of the 200 rater pairs and the percentages
for (rater_id1, rater_id2), percent_double_scored in itertools.product(rater_pairs, double_scored_percentages):
# get a data frame that only has the scores from the two given raters
# with a given percentage of the second rater's scores masked out;
# this function also returns the actual number of double scored
# responses in the returned data frame
(df_percent_double_scored,
num_double_scored) = simulate_percent_double_scored(df_scores,
rater_id1,
rater_id2,
percent_double_scored)
# add the system scores into the same data frame for convenience
df_percent_double_scored[chosen_system] = df_scores[chosen_system]
# also compute another version of this data frame where the double-scored responses#
# are the only responses; we need this to check whether what matters is the % of
# double-scored responses or just the sheer *number* of double-scored responses
df_only_double_scores = df_percent_double_scored.dropna()
# compute the PRMSE using this dataset with only ``percentage``-percent double scored responses
prmse_percent = prmse_true(df_percent_double_scored[chosen_system],
df_percent_double_scored[[rater_id1, rater_id2]])
# also compute the PRMSE using the double-scores only dataset
prmse_number = prmse_true(df_only_double_scores[chosen_system],
df_only_double_scores[[rater_id1, rater_id2]])
# compute the other conventional agreement metrics for the same dataset
agreement_metrics = compute_agreement_one_system_one_rater_pair(df_percent_double_scored,
chosen_system,
rater_id1,
rater_id2,
include_mean=True)[0]
# save the PRMSE value along with other metadata
metric_values.append({'rater_id1': rater_id1,
'rater_id2': rater_id2,
'percent_double_scored': percent_double_scored,
'num_double_scored': num_double_scored,
'r': agreement_metrics['r'],
'QWK': agreement_metrics['QWK'],
'R2': agreement_metrics['R2'],
'prmse_percent': prmse_percent,
'prmse_number': prmse_number})
```
### Step 3: Examine impact of double-scoring
Now we use some plots to examine the actual impact of double-scoring on the PRMSE metric.
```
# now create a data frame from the PRMSE list
df_metrics_double_scored = pd.DataFrame(metric_values)
# also add in the rater category from the rater metadata since we need that for our plots
# note that since both the raters in any pair are from the same category, we can just
# merge on the first rater ID
df_metrics_double_scored_with_categories = df_metrics_double_scored.merge(df_rater_metadata,
left_on=['rater_id1'],
right_on=['rater_id'])
```
So, the first plot we look at is that of the two PRMSE values: (a) one computed over datasets with varying percentage of double-scored responses and (b) the other computed over datasets of varying sizes that only-contain double-scored responses. We also facet both plots by the rater category -- i.e., the average inter-rater agreement between the two simulated raters.
```
# create a figure with 2 subplots
fig, axes = plt.subplots(1, 2);
fig.set_size_inches(20, 5)
with sns.plotting_context('notebook', font_scale=1.3):
for prmse_type, ax in zip(['percent', 'number'], axes):
# choose what goes on the x and y axes
x_column = 'percent_double_scored' if prmse_type == 'percent' else 'num_double_scored'
y_column = 'prmse_percent' if prmse_type == 'percent' else 'prmse_number'
# set the x-axis labels and tick labels appropriately
x_label = "% (n) double-scored responses" if prmse_type == 'percent' else 'Number of responses (all double-scored)'
percent_ticklabels = [f"{pct}%\n({num})" for pct, num in zip(double_scored_percentages,
df_metrics_double_scored_with_categories['num_double_scored'])]
number_ticklabels = df_metrics_double_scored_with_categories['num_double_scored'].tolist()
x_ticklabels = percent_ticklabels if prmse_type == 'percent' else number_ticklabels
# now add the the boxplot of the PRMSE values
sns.boxplot(x=x_column,
y=y_column,
hue='rater_category',
hue_order=dataset.rater_categories,
data=df_metrics_double_scored_with_categories,
ax=ax)
# and set the axis labels
ax.set_xlabel(x_label, fontsize=14)
ax.set_ylabel("PRMSE", fontsize=14)
ax.set_xticklabels(x_ticklabels)
ax.tick_params('x', labelsize=14)
# There are several outlier values of PRMSE at 1.6. We remove them to reduce the white space on the plots
# and make a note in the caption.
# We'll add them back before release.
ax.set_ylim(0.4, 1.2)
# we also want to add in a line showing the R2 score between the simulated true scores
# and the scores from our chosen simulated system
r2_true_system_score = r2_score(df_scores['true'], df_scores[chosen_system])
ax.axhline(r2_true_system_score, color='black', ls=':')
plt.show()
```
These plots shows that PRMSE scores are much more stable with a larger percentage of double-scored responses and what matters is the number of double scored responses, not the percentage. A good rule of thumb is to have at least 1000 double-scored responses.
We can also look at the same data in tabular format. For simplicity, we will use the range of values as the measure of spread.
```
# group the metrics by the double-scoring percentage and then by rater category
grouper = df_metrics_double_scored_with_categories.groupby(['percent_double_scored',
'rater_category'])
# compute the descriptives over the groups, for the PRMSE metric
df_prmse_descriptives = grouper['prmse_percent'].describe()
# compute the range as the difference between the maximum PRMSE value and the minimum
df_prmse_descriptives['range'] = df_prmse_descriptives['max'] - df_prmse_descriptives['min']
# get the range as a separate dataframe and print out in rater category order
df_prmse_range = df_prmse_descriptives['range'].unstack()[dataset.rater_categories]
df_prmse_range
```
Next, we look at the other conventional agrement metrics to see how they vary with the number of double-scored responses available.
```
# extract out only the agreement metrics columns along with the percentage and the rater category
df_agreement_double_scored = df_metrics_double_scored_with_categories[['rater_category', 'r', 'QWK', 'R2', 'percent_double_scored']]
# now create a longer version of this data frame that's more amenable to plotting
df_agreement_double_scored_long = df_agreement_double_scored.melt(id_vars=['percent_double_scored', 'rater_category'],
var_name='metric')
# make the plot
g = sns.catplot(x='percent_double_scored',
y='value',
col='metric',
hue='rater_category',
hue_order=dataset.rater_categories,
kind='box',
sharey=False,
data=df_agreement_double_scored_long)
g.set_axis_labels("% double-scored responses", "value")
g._legend.set_title('rater category')
plt.show()
```
These plots show that the agreement metrics are not as stable as PRMSE even with more double-scored responses and still depend significantly on how well the two raters agreed with each other.
| github_jupyter |
### Introduction and Motivation
Machine learning courses often fixate on discriminative models (such as decision trees and linear models) to the point where many machine learning engineers ignore generative models (such as Gaussian mixture models and generative adversarial networks) altogether. In my experience, I think the same is often done for time series modeling and anomaly detection. It is my goal to introduce my readers to the world of simple generative models. As an example, I'll be discussing DGAs (we'll talk about DGAs in a bit).
It is my goal in this post to give my readers a taste of what generative models can do and why they are important tools in any data scientist's toolkit.
I'll start with a motivating example in cyber security: detecting DGAs. I'll explain what DGAs are, why we care about them, and why discriminative models can sometimes fail miserably in finding them. I'll then discuss a simple generative model (the markov chain) and then finish with a few parting words on how to learn more.
Many readers might (and probably will) ask why I am writing a post about domain generating algorithms (DGAs). DGAs have been done to death in the cyber security space! That is mostly true. There has been a lot of attention on DGAs and they have been covered quite thoroughly from a variety of different angles. However, data on DGAs is very convenient to find and easy to work with, so they present a nice data set to work on.
A big thanks to my friend Kate Highnam who introduced me to the world of DGAs and often patiently listens to my rants on statistics.
### What is a domain?
Before we define a domain generating algorithm, we first need to define a domain! Domains (or domain names) are single character strings that we use to communicate with a DNS server to tell the server what website we want to go to. The DNS server takes this string and connects us to the IP address or addresses of the servers that the domain belongs to.
For example: I might want to go to the domain "google.com". I would put Google into my search engine that would look up the servers Google runs to process requests for google.com and then return whatever those servers want to send back to me.
Why do we like this? There are actually a ton of reasons that I won't get into, but the simplest reason is that it's hard to remember an IP address like 151.101.129.121 but easy to remember a domain name like lifewire.com.
Often in cyber security you will see domains printed as such: malware[.]com. The reason for this is that the domain may link to actual malware and inserting brackets avoids accident hyper link clicking.
### What is a domain generating algorithm?
DGAs are just pieces of code that produce domain names. They often have some set of rules such that the domains they generate look random but follow some set pattern. There are many uses for DGAs. Content distribution networks (CDN) and ad networks use DGAs to quickly create domains for transient content and to get around ad blocker tools. Bad actors (hackers and other miscreants) uses DGAs to disguise communications between a command and control server (C&C) and malware installed on a target computer.
We want to detect DGAs so that we can find instances where malware is communicating with its C&C, which acts like a sort of mother ship, and silence those communications. There is a lot more than can be said on this topic on the cyber security front, but I'm easily distracted and I want to at least try to stay on point.
The basic procedure of using a DGA is to have your malware generate many (often hundreds r thousands) of domain names a day (cadence can vary depending on beaconing behavior). The bad actor will know what domains will be generated on that day because he created the DGA and so he can buy a few of those domains in advance and point them to his C&C server. Rinse and repeat to ensure that cyber security analysts never catch on to where your C&C.
What is hard about detecting malware DGAs is that they are intentionally trying NOT to be found! Hackers are always coming up with new DGAs to fool our DGA detection models.
Below is a super simple example of a DGA from: https://en.wikipedia.org/wiki/Domain_generation_algorithm
You can also look at these resources to see actual DGAs that ahve been reverse engineered from malware: [Endgame Github](https://github.com/endgameinc/dga_predict/tree/master/dga_classifier/dga_generators), [Paul Chaignon Repo](https://github.com/pchaigno/dga-collection)
```
def generate_domain(year, month, day):
"""Generates a domain name for the given date."""
domain = ""
for i in range(16):
year = ((year ^ 8 * year) >> 11) ^ ((year & 0xFFFFFFF0) << 17)
month = ((month ^ 4 * month) >> 25) ^ 16 * (month & 0xFFFFFFF8)
day = ((day ^ (day << 13)) >> 19) ^ ((day & 0xFFFFFFFE) << 12)
domain += chr(((year ^ month ^ day) % 25) + 97)
return domain
print "Sample Domain from DGA:", generate_domain(2018,5,1) + '.com'
```
### Discriminators vs Generators?
Let's discuss many of the more popular ways of identifying DGAs. First we can simply look up if someone else has identified a domain as a DGA using services like [virus total](https://www.virustotal.com/), [whois](https://www.whois.net/), or [DGArchive](https://dgarchive.caad.fkie.fraunhofer.de/).
A very popular next step is to train a discriminator to identify dgas from non-dgas. A discriminator is simply a model that 'learns' a hard or soft boundary between classes. There are numerous papers on how to use GBM, neural nets, and a host of other model types to detect DGAs. The work is quite interesting, but it is hard to produce novel research is such a saturated space. If you want to play around in this space, I recommend this fun [github project](https://github.com/andrewaeva/DGA).
The problem with discriminators is that they establish their class boundaries based on what they've seen. For the benign case this is usually fine. The nature of benign domain names rarely changes that much. DGAs, however, can change a lot as bad actors try to get around our detection models. Therefore, the decision boundary that worked for our train data might not work for our test data a few months after we deploy it (or may not work as well).
We can try to combat this with [generative models](https://stats.stackexchange.com/questions/12421/generative-vs-discriminative). These models try to find the distribution over each individual class. Rather than asking the question "is this domain class A or B?", we ask "is this class A?". Now because we are trying to model an entire distribution instead of just finding a decision boundary, generative models can be much less accurate than discriminative models. This is why discriminative models are favored in cyber security. Cyber teams need to feed potential DGAs through human agents and so need to lower false positive rates to keep queue sizes reasonable for their agents. [Here](http://www.raid-symposium.org/raid99/PAPERS/Axelsson.pdf) is a good discussion of this topic.
However, running both a generative model and discriminative model might allow a small cyber security team to get the best of both worlds:
* accurate decision boundaries that produce low FP rates
* a detection system that isn't 'over fit' to the train classes
If you felt like this went over your head just wait, things will become clearer with the examples.
### What is a markov chain?
Ok, so let's get started. There are many kinds of generative models. I've built generators from neural nets (like GANs and Autoencoders), kernel density estimates over hand crafted feature spaces, Gaussian processes, etc. But these are hard models to build and can be very finicky. I often feel like I am practicing black magic when trying to tune a GAN model.
So we're starting small. Markov models are simple models to understand but are SUPER powerful. I always try to use markov models if I can because you can implement them easily, often get great results, and they are easily interpreted.
Let me preface next part of this section with a couple important notes. Firstly, I will be covering Markov chains in more detail in a later blog post on Markov Decision Processes (MDP) in the context of reinforcement learning, so forgive me if I move quickly over them. Second, my code for building a markov chain will be very crude and inefficient. This is because I wrote the code to be easy to understand and straight forward. In the MDP I will use a much more efficient methodology.
In the early 1900s Andrey Markov produced some interesting work concerning discrete processes he called 'chains'. These chains were a set of successive states and a process that describes the movement from one state to the next. Each move is a single step and is based on a transition model. A Markov chain is based on the Markov Property. The Markov property states that given the present, the future is conditionally independent of the past. That’s it, the state in which the process is now it is dependent only from the state it was at
t-1.
If we can empirically arrive at the transition matrix that describes the probability of transitioning from one state to another, then we can know the probability that a certain sequence of states was observed (given the assumption that the process has markovian properties).
### Building our DGA generator
So how do we use the markov chain? Well, we can think of each character in a domain as a state. We can then determine the probability to seeing a certain character given the character before it. Now, any linguist worth their salt will tell you that language is more complex than this (especially English) and that words do not perfectly adhere to the markov property. Well, it's an approximation!
Let's put this into practice.
First, get the all_legit.txt and all_dga.txt files from this handy dandy source: https://github.com/andrewaeva/DGA
The all_legit.txt contains the alexa one million dataset of popular benign domains. The all_dga.txt contains actual examples of dgas from popular malware families.
```
import pandas as pd
import numpy as np
df_alexa1m = pd.read_csv("all_legit.txt",header=None)
df_alexa1m.columns = ['domains']
df_alexa1m.domains = df_alexa1m.domains.apply( lambda x: x.split(".")[0] )
df_alexa1m.domains = df_alexa1m.domains.apply( lambda x: x.lower() )
```
We then encode each possible character in the domain as an index value and store those values in a dictionary (we also create a look up dictionary for convenience). This is a popular trick to transform strings into a vector format that a model can ingest. Here I do it because it makes my life easier filling in the transition matrix later on. I'm lazy.
We include '!' and '@' as start and stop characters to represent the probability of starting or ending with a certain character. This is important or our transition matrix won't take into account the probability of having a particular start or stop character!
```
character_dict = {}
i = 0
for domain in df_alexa1m.domains.values:
for char in domain:
if char not in character_dict.keys():
character_dict[char] = i
i += 1
else:
pass
# add padding chars
character_dict['!'] = i
i += 1
character_dict['@'] = i
character_lookup_dict = {v: k for k, v in character_dict.iteritems()} # python 2 version
#character_dict_lookup = {v: k for k, v in character_dict.items()} # python 3 version
```
We then initialize a matrix that is size nxn where n is the number of possible states (characters). The way we arrive at our transition probabilities is by counting the instances of those transitions in our alexa one million set. Since some transitions might not exist at all (our data might not be 100% representative of the true population) we initialize every transition count to 1 so that when we normalize the transition counts to get probability scores no probability will be exactly zero (just really really close). We are therefore making the assumption that no transition is exactly impossible, only possibly highly unlikely.
```
num_chars = np.max( character_dict.values() ) + 1
transition_matrix = np.ones((num_chars,num_chars))
```
Then we simply count the number fo transitions we see in our alexa one million domains!
```
for domain in df_alexa1m.domains.values:
domain = '!' + domain + '@'
domain = [ character_dict[x] for x in domain]
for i, char in enumerate(domain):
if i+1 <= len(domain)-1:
char_1 = domain[i]
char_2 = domain[i+1]
transition_matrix[char_1][char_2] += 1
```
We normalize our counts to get probabilities.
```
row_sums = transition_matrix.sum(axis=1)
transition_matrix = transition_matrix / row_sums[:, np.newaxis]
```
So how do we interpret this transition matrix? Well, each value in the matrix represents the probability of transitioning from one state (character) to another. For instance, what is the probability of transitioning to 'a' given that we are in state 'b'?
```
char_a = character_dict['a']
char_b = character_dict['b']
print "Prob of going from b -> a: {}%".format( 100.00 * transition_matrix[char_b,char_a] )
```
Now we could try to find the probability of seeing a particular domain as the power series of the probability of each of it's character transitions...but we will often run into underflow issues (take a peak at the transition matrix to see why). Instead we will borrow from our handy stats textbooks and just take the summed log likelihoods.
```
log_liklihood_list = []
for domain in df_alexa1m.domains.values:
domain = '!' + domain + '@'
domain = [ character_dict[x] for x in domain]
log_liklihood = 0
for i, char in enumerate(domain):
if i+1 <= len(domain)-1:
char_1 = domain[i]
char_2 = domain[i+1]
transition_probability = transition_matrix[char_1,char_2]
log_liklihood += np.log(transition_probability)
# normalize by length
log_liklihood = log_liklihood / len(domain)
log_liklihood_list.append(log_liklihood)
df_alexa1m['LL'] = log_liklihood_list
import matplotlib.pyplot as plt
import seaborn as sns
plt.figure(figsize=(20,5))
sns.distplot( df_alexa1m.LL.values )
plt.show()
```
Above we see the distribution of log likelihoods for the alexa one million domains given our markov chain model. This is cool to look at, but tells us very little. Let's compare this to the distribution over the zeus dga domains (zeus is a famous family of malware which sometimes makes use of particular domain generators).
```
df_dgas = pd.read_csv("all_dga.txt",header=None)
df_dgas.columns = ['domains']
df_dgas['dga_family'] = df_dgas.domains.apply(lambda x: x[-1])
df_dgas.domains = df_dgas.domains.apply( lambda x: x.split(".")[0] )
df_dgas.domains = df_dgas.domains.apply( lambda x: x.lower() )
def dga_family(x):
if x == '0':
return "legit"
elif x == '1':
return "cryptolocker"
elif x == '2':
return "zeus"
elif x == '3':
return "pushdo"
elif x == '4':
return "rovnix"
elif x == '5':
return "tinba"
elif x == '6':
return "conflicker"
elif x == '7':
return "matsnu"
elif x == '8':
return "rambo"
else:
return None
df_dgas.dga_family = df_dgas.dga_family.apply(lambda x: dga_family(x))
log_liklihood_list = []
for domain in df_dgas.domains.values:
domain = '!' + domain + '@'
domain = [ character_dict[x] for x in domain]
log_liklihood = 0
for i, char in enumerate(domain):
if i+1 <= len(domain)-1:
char_1 = domain[i]
char_2 = domain[i+1]
transition_probability = transition_matrix[char_1,char_2]
log_liklihood += np.log(transition_probability)
# normalize by length
log_liklihood = log_liklihood / len(domain)
log_liklihood_list.append(log_liklihood)
df_dgas['LL'] = log_liklihood_list
plt.figure(figsize=(20,5))
sns.distplot( df_alexa1m.LL.values ,label = 'alexa1m')
sns.distplot( df_dgas[df_dgas.dga_family == 'zeus'].LL.values ,label='zeus')
plt.legend()
plt.show()
```
As we can see, the zeus domains all have MUCH lower log likelihoods than the alexa one million domains. What does this mean? It means that given the transition probabilities in the alexa one million set, the zeus set is very unlikely to have been generated using the same process.
Ok...so what? Well, we just built a model that can tell that the zeus domains are not benign WITHOUT training on a single zeus domain. We didn't need any samples of the zeus dgas. Theoretically, we could detect DGAs we've never seen before and never trained on. Now there are a variety of nuances, practical considerations, and qualifications to that statement...but I think this is pretty neato.
Now some of you out there are thinking...this is super cool, look how easy it would be to draw a line between those humps to separate zeus domains from alexa domains. With generators we look at the probability or likelihood that a data point came from the process our generator describes. We're typically not in the business of drawing lines (although we might for practical purposes). Let's look at why drawing lines can be bad.
Below I'm going to plot the alexa domains and zeus domains against each other with domain length on the y axis and log likelihood (from our markov model) on the x axis. These distributions look pretty different right? We could easily draw a line between then to define the zeus and alexa classes!
```
df_alexa1m['length'] = df_alexa1m.domains.apply(lambda x: len(x))
df_dgas['length'] = df_dgas.domains.apply(lambda x: len(x))
plt.figure(figsize=(20,5))
plt.scatter(df_alexa1m.LL.values,df_alexa1m.length.values,alpha=.05,label='alexa1m')
plt.scatter(df_dgas[df_dgas.dga_family == 'zeus'].LL.values,df_dgas[df_dgas.dga_family == 'zeus'].length.values,alpha=.05,label='zeus')
plt.legend()
plt.show()
```
Below I've drawn our line. Everything looks fine, right? This is a model that we can be proud of and deploy into production!
```
plt.figure(figsize=(20,5))
plt.scatter(df_alexa1m.LL.values,df_alexa1m.length.values,alpha=.05,label='alexa1m')
plt.scatter(df_dgas[df_dgas.dga_family == 'zeus'].LL.values,df_dgas[df_dgas.dga_family == 'zeus'].length.values,alpha=.05,label='zeus')
plt.legend()
plt.plot([0, -5.0], [95, -4.0], 'r-', lw=2)
plt.show()
```
But what if hackers figure out (or outright guess) that domain length is a feature in our model? It’s pretty plausible that a hacker might do this and I may address that in a future blog post. Let's see what happens when the "hacker" shortens the length of the zeus domains to match the alexa domains more closely.
```
plt.figure(figsize=(20,5))
plt.scatter(df_alexa1m.LL.values,df_alexa1m.length.values,alpha=.05,label='alexa1m')
plt.scatter(df_dgas[df_dgas.dga_family == 'zeus'].LL.values,df_dgas[df_dgas.dga_family == 'zeus'].length.values - 20.0,alpha=.05,label='zeus')
plt.legend()
plt.plot([0, -5.0], [95, -4.0], 'r-', lw=2)
plt.show()
```
Suddenly our line doesn't seem like such a great decision boundary! However, you'll notice that the zeus domains are still at the very tail end of our generative model (If this isn't clear, try graphing each axis separately as a histogram. Scatter plots can make interpreting densities difficult). This is a toy example of how a generative model might tell us that something is wrong even if the discriminative model doesn't detect anything bad.
### Practical Tips for Using Your Model
Before I turn you loose on the world with your new found knowledge, I wanted to go through a few practical concerns. This isn't a thorough treatment of how to use generative models, so please don't take this code and try to productionize a model!
So how do we sue our model? Well, the easiest way is to use our log likelihood scores as anomaly scores. The bigger the LL the less likely we are to have seen that data point given the train data. We could then send x% of the anomalies to our cyber security team for a thorough analysis.
That's not an ideal solution for two reasons. Firstly, it assumes that lower likelihood scores are 'better'. However, we know that hackers try to make DGAs look like non-DGAs (One interesting example: [DeepDGA: Adversarially-Tuned Domain Generation and Detection](https://arxiv.org/abs/1610.01969)). What if a hacker uses adversarial training techniques to produce DGAs that perfectly fit our detection model? We might start to see unusually low log likelihoods (if the hacker is sloppy). More importantly, it's hard to tune log likelihood. We would ideally like to use the model in a way such that we can intelligently tune it's sensitivity.
To this end, we'll discuss (at a very high level) two methods: Kernel Density Estimation and Gaussian Mixture Models. I'm going to gloss over quite a bit of stats and assumptions. I don't feel too guilty given that this is just a fun blog post...if you are a statistical purist; please don't hate me too much!
First we'll cover a very simple trick: [kernel density estimation](https://en.wikipedia.org/wiki/Kernel_density_estimation) (kde). KDEs are a smoothing technique used to approximate a continuous probability density function from finite data. KDEs are useful for a lot of problems in signal processing and econometrics. Typically we sue caution when fitting KDEs given that our choice of kernel and kernel parameters can lead to over fitting...but our distributions here are simple and so we don't need to be too worried.
Below I've fit a one dimensional kde to the histogram of our markov model likelihoods. Do you think the kde accurately represents the distribution of likelihoods?
```
from scipy.stats import gaussian_kde
class Kernel_Density_Estimate(object):
"""Kenerl Density Estimation with Scipy"""
# Note that scipy weights its bandwidth by the covariance
# of theinput data. To make the results comparable to the other methods,
# we devide the bandwidth by the sample standard deviation here.
def __init__(self,bandwidth=0.2):
self.bandwidth = bandwidth
self.kde = None
def train(self, x, **kwargs):
self.kde = gaussian_kde(x, bw_method = self.bandwidth / x.std(), **kwargs)
def predict(self, x):
return self.kde.evaluate(x)
kde = Kernel_Density_Estimate(bandwidth=.05)
kde.train( df_alexa1m.LL.values )
x = np.linspace(-6,0,1000)
pdf = kde.predict(x)
fig = plt.figure(figsize=(20,5))
plt.subplot(1, 2, 1)
plt.plot(x,pdf)
plt.xlim(-6,0)
plt.ylim(0,2)
plt.title("Kernel Density Estimate of Alexa 1M LL")
plt.subplot(1, 2, 2)
sns.distplot(df_alexa1m.LL.values, kde=False)
plt.xlim(-6,0)
plt.title("Actual Data of Alexa 1M LL")
plt.show()
```
Now that we have an estimate of the probability density function (pdf) of our data given our markov model we can artificially discretize it and then approximate the cumulative density function (cdf) of our data. With the cdf we can find the 99% credibility interval (yes, I mean credibility and not confidence!) of the data. Assuming our alexa1m set is representative of our test data and our assumptions around our model and cdf approximation are correct, then we will identify 1% of our benign domains as DGAs.
Below I print the % of DGAs caught, the % of alexa1m domains accidently caught, and the 99% credible interval. You'll notice that this is a two-tailed credible interval. Why do I consider anomalies with very low LLs? Well, a hacker might be producing domains using an Oracle he created based on the alexa1m data (that he knows is a popular set to build DGA detectors on). His DGAs might look TOO much like the alexa1m set and so be perfectly modeled by the markov chain. It turns out that this isn't the case here, but it's interesting to think about.
```
cdf = 100.00 * pdf / pdf.sum()
cdf = cdf.cumsum()
lower_bound = x[ cdf[cdf <= .5].shape[0] ].round(2)
upper_bound = x[ cdf[cdf <= 99.5].shape[0] ].round(2)
print "99% Credible Interval for LL: {} : {}".format(lower_bound, upper_bound)
DGA_Anomalies = 0
for LL in df_dgas[df_dgas.dga_family == 'zeus']['LL']:
if LL < lower_bound or LL > upper_bound:
DGA_Anomalies += 1
Alexa_Anomalies = 0
for LL in df_alexa1m.LL:
if LL < lower_bound or LL > upper_bound:
Alexa_Anomalies += 1
dgas_caught = np.round(100.0 * DGA_Anomalies / df_dgas[df_dgas.dga_family == 'zeus'].shape[0],1)
alexa1m_caught = np.round(100.0 * Alexa_Anomalies / df_alexa1m.shape[0],1)
print "Percent of Zeus Domains Caught: {}% \nPercent of Alexa 1M Domains Caught: {}%".format(dgas_caught,alexa1m_caught)
```
Not bad! This is a pretty good catch rate! However, we need to take into account the base rate fallacy. If you are not familiar with the base rate fallacy, I **HIGHLY** recommend you read through this short paper: [The Base-Rate Fallacy and its Implications for the
Difficulty of Intrusion Detection](http://www.raid-symposium.org/raid99/PAPERS/Axelsson.pdf).
Basically, the number of benign domains we see in the real world is always FAR bigger than the number of DGAs. We can use this information to see how many benign domains we will flag for our human cyber security agents to detect vs the number of actual DGAs.
Let's say we see 10,000,000 unique domains a day and that .1% of these are actual DGAs. That means that 10,000 DGAs and 9,990,000 benign domains exist in our proxy logs.
$.999 * 10,000 = 9,990$ <br>
$.001 * 10,000 = 10$ <br>
We catch 9,990 DGAs and miss 10 at the 99% credible interval.
$.009 * 9,990,000 = 89,910$ <br>
$.991 * 9,990,000 = 9,900,090$ <br>
Unfortunately we falsely identify almost 90,000 benign domains as DGAs! That means about 9 / 10 domains sent to our human agents will NOT be malicious (and this is often optimistic compared to reality).
Just for fun, let's use look at the one tailed 99.5% credibility interval.
```
cdf = 100.00 * pdf / pdf.sum()
cdf = cdf.cumsum()
lower_bound = x[ cdf[cdf <= .5].shape[0] ].round(2)
print "99.5% Credible Interval for LL: <{}".format(lower_bound)
DGA_Anomalies = 0
for LL in df_dgas[df_dgas.dga_family == 'zeus']['LL']:
if LL < lower_bound:
DGA_Anomalies += 1
Alexa_Anomalies = 0
for LL in df_alexa1m.LL:
if LL < lower_bound:
Alexa_Anomalies += 1
dgas_caught = np.round(100.0 * DGA_Anomalies / df_dgas[df_dgas.dga_family == 'zeus'].shape[0],1)
alexa1m_caught = np.round(100.0 * Alexa_Anomalies / df_alexa1m.shape[0],1)
print "Percent of Zeus Domains Caught: {}% \nPercent of Alexa 1M Domains Caught: {}%".format(dgas_caught,alexa1m_caught)
```
We'll use the same assumptions as before, but with the one tailed 99.5% credibility interval.
$.999 * 10,000 = 9,990$ <br>
$.001 * 10,000 = 10$ <br>
We catch 9,990 DGAs and miss 10 at the 99% credible interval.
$.005 * 9,990,000 = 49,950$ <br>
$.995 * 9,990,000 = 9,940,050$ <br>
Now only about 8 / 10 domains sent to our human agents will NOT be malicious.
Perhaps we can do a little better than that! Rather than just considering the LL from our markov model, we'll also consider the length of the domain as a feature of interest. Now that we have two dimensions to consider, we could use a 2D kernel...but as the number of dimensions gets higher, that gets tricky. Instead, we'll use a mixture model to approximate the distribution of our data. A mixture model combines a set of simple distributions (like a normal/Gaussian, Poisson , or beta distribution) together to create something more complex (like the basis functions in Fourier transforms that I cover in my time series course). There are really complicated mixture models that we could sue, but we're going to make some naive assumptions. We'll assume that we can model our data as a combination of Gaussian distributions (we call this a gmm or [Gaussian mixture model](https://en.wikipedia.org/wiki/Mixture_model)).
I'll skip over the details of GMMs, but I highly encourage you to check them out since they are super cool and highly useful. I've made use of GMMs in a wide variety of use cases from voice recognition to reinforcement learning. I'll start with a single component GMM (one Gaussian). I plot the Gaussian in 2D space with ellipses to show x standard deviations away from the mean.
```
from sklearn.mixture import GMM
from matplotlib.patches import Ellipse
def draw_ellipse(position, covariance, ax=None, **kwargs):
"""Draw an ellipse with a given position and covariance"""
ax = ax or plt.gca()
# Convert covariance to principal axes
if covariance.shape == (2, 2):
U, s, Vt = np.linalg.svd(covariance)
angle = np.degrees(np.arctan2(U[1, 0], U[0, 0]))
width, height = 2 * np.sqrt(s)
else:
angle = 0
width, height = 1.96 * np.sqrt(covariance)
# Draw the Ellipse
for nsig in range(1, 4):
ax.add_patch(Ellipse(position, nsig * width, nsig * height,
angle, **kwargs))
def plot_gmm(gmm, X, label=True, ax=None):
plt.figure(figsize=(20,10))
sns.set_style("white")
ax = ax or plt.gca()
X = X[np.random.randint(X.shape[0], size= int(X.shape[0]*.005) ), :]
labels = gmm.fit(X).predict(X)
w_factor = 0.2 / gmm.weights_.max()
for pos, covar, w in zip(gmm.means_, gmm.covars_, gmm.weights_):
draw_ellipse(pos, covar, alpha=.1)#alpha=w * w_factor)
alpha = 0.08
if label:
ax.scatter(X[:, 0], X[:, 1], c=labels, s=40, cmap='viridis', zorder=2, alpha=alpha)
else:
ax.scatter(X[:, 0], X[:, 1], s=40, zorder=2, alpha=alpha)
plt.show()
gmm = GMM(n_components=1, random_state=42)
plot_gmm(gmm, df_alexa1m[['LL','length']].as_matrix() )
```
Next, we'll plot the log liklihoods that each zeus and alexa1m domain belongs to our gaussian.
```
x1 = df_dgas[df_dgas.dga_family == 'zeus'][['LL','length']]
dga_scores = gmm.score( x1 )
x1 = df_alexa1m[['LL','length']]
alexa1m_scores = gmm.score( x1 )
plt.figure(figsize=(20,5))
sns.distplot(dga_scores,label='zeus',bins=1000)
sns.distplot(alexa1m_scores,label='alexa1m',bins=1000)
plt.xlim(-60,0)
plt.legend()
plt.show()
```
This is great, we see that zeus domains have MUCH lower log liklihoods. If we so wanted, we could use our KDE trick on the LL of our GMM. Let's try it.
```
kde = Kernel_Density_Estimate(bandwidth=.1)
kde.train( alexa1m_scores )
x = np.linspace(-60,0,1000)
pdf = kde.predict(x)
fig = plt.figure(figsize=(20,5))
plt.subplot(1, 2, 1)
plt.plot(x,pdf)
plt.xlim(-60,0)
plt.ylim(0,1.20)
plt.title("Kernel Density Estimate of Alexa 1M LL")
plt.subplot(1, 2, 2)
sns.distplot(alexa1m_scores, kde=False, bins=500)
plt.xlim(-60,0)
plt.title("Actual Data of Alexa 1M LL")
plt.show()
cdf = 100.00 * pdf / pdf.sum()
cdf = cdf.cumsum()
lower_bound = x[ cdf[cdf <= .5].shape[0] ].round(2)
upper_bound = x[ cdf[cdf <= 99.5].shape[0] ].round(2)
print "99% Credible Interval for LL: {} : {}".format(lower_bound, upper_bound)
DGA_Anomalies = 0
for LL in dga_scores:
if LL < lower_bound or LL > upper_bound:
DGA_Anomalies += 1
Alexa_Anomalies = 0
for LL in alexa1m_scores:
if LL < lower_bound or LL > upper_bound:
Alexa_Anomalies += 1
dgas_caught = np.round(100.0 * DGA_Anomalies / dga_scores.shape[0],1)
alexa1m_caught = np.round(100.0 * Alexa_Anomalies / alexa1m_scores.shape[0],1)
print "Percent of Zeus Domains Caught: {}% \nPercent of Alexa 1M Domains Caught: {}%".format(dgas_caught,alexa1m_caught)
```
Wow, now we've caught all of the zeus domains without increasing the number of alexa1m domains caught!
However, let's consider our calculation from earlier.
We'll use the same assumptions as before, but with the one tailed 99.5% credibility interval.
$1.0 * 10,000 = 10,000$ <br>
$0.0 * 10,000 = 0$ <br>
We catch 9,990 DGAs and miss 10 at the 99% credible interval.
$.005 * 9,990,000 = 49,950$ <br>
$.995 * 9,990,000 = 9,940,050$ <br>
We still have about 8 / 10 domains sent to our human agents that will NOT be malicious! Let's try using a one tailed 99.9% credible interval and see what happens.
```
cdf = 100.00 * pdf / pdf.sum()
cdf = cdf.cumsum()
lower_bound = x[ cdf[cdf <= .1].shape[0] ].round(2)
print "99.9% Credible Interval for LL: <{}".format(lower_bound)
DGA_Anomalies = 0
for LL in dga_scores:
if LL < lower_bound:
DGA_Anomalies += 1
Alexa_Anomalies = 0
for LL in alexa1m_scores:
if LL < lower_bound:
Alexa_Anomalies += 1
dgas_caught = np.round(100.0 * DGA_Anomalies / dga_scores.shape[0],1)
alexa1m_caught = np.round(100.0 * Alexa_Anomalies / alexa1m_scores.shape[0],1)
print "Percent of Zeus Domains Caught: {}% \nPercent of Alexa 1M Domains Caught: {}%".format(dgas_caught,alexa1m_caught)
```
We'll use the same assumptions as before, but with the one tailed 99.5% credibility interval.
$.987 * 10,000 = 9870$ <br>
$.015 * 10,000 = 130$ <br>
We catch 9,990 DGAs and miss 10 at the 99% credible interval.
$.001 * 9,990,000 = 9,990$ <br>
$.999 * 9,990,000 = 9,980,010$ <br>
Now only about 5 / 10 domains sent to our human agents that will NOT be malicious! Pretty neat right? We're catching about the same volume of DGAs, but we're using our human agents much more efficiently. It should be noted that you can tune discriminators in a very similar way as we've tuned the output of our generative model, but I won't cover that here.
Just for fun let's try using more than one Gaussian component. I won't get into how I arrived at 2 components (honestly, there are many evaluation metrics you can use like AIC, BIC, discriminator attributes, etc.). I leave it to the reader to delve into the wacky world of mixed models and their components.
```
gmm = GMM(n_components=2, random_state=42)
plot_gmm(gmm, df_alexa1m[['LL','length']].as_matrix() )
x1 = df_dgas[df_dgas.dga_family == 'zeus'][['LL','length']]
dga_scores = gmm.score( x1 )
x1 = df_alexa1m[['LL','length']]
alexa1m_scores = gmm.score( x1 )
kde = Kernel_Density_Estimate(bandwidth=.1)
kde.train( alexa1m_scores )
x = np.linspace(-60,0,1000)
pdf = kde.predict(x)
fig = plt.figure(figsize=(20,5))
plt.subplot(1, 2, 1)
plt.plot(x,pdf)
plt.xlim(-60,0)
plt.ylim(0,1.20)
plt.title("Kernel Density Estimate of Alexa 1M LL")
plt.subplot(1, 2, 2)
sns.distplot(alexa1m_scores, kde=False, bins=500)
plt.xlim(-60,0)
plt.title("Actual Data of Alexa 1M LL")
plt.show()
cdf = 100.00 * pdf / pdf.sum()
cdf = cdf.cumsum()
lower_bound = x[ cdf[cdf <= .1].shape[0] ].round(2)
print "99.9% Credible Interval for LL: <{}".format(lower_bound)
DGA_Anomalies = 0
for LL in dga_scores:
if LL < lower_bound:
DGA_Anomalies += 1
Alexa_Anomalies = 0
for LL in alexa1m_scores:
if LL < lower_bound:
Alexa_Anomalies += 1
dgas_caught = np.round(100.0 * DGA_Anomalies / dga_scores.shape[0],1)
alexa1m_caught = np.round(100.0 * Alexa_Anomalies / alexa1m_scores.shape[0],1)
print "Percent of Zeus Domains Caught: {}% \nPercent of Alexa 1M Domains Caught: {}%".format(dgas_caught,alexa1m_caught)
```
Well slap my knee and call me Greg, those are some great results! Now I will note that I chose the zeus DGA family because they are actually very easy to identify using either a generator or discriminator. I leave it to the reader to see how the generator works on the other families of DGAs in our dataset.
One last note, based on the math from before 50% of the anomalies our model detects will be actual malicious DGAs. However, we are detecting ~20,000 anomalies a day! Let's say we have 100 human agents that can each go through 60 DGAs in a full work day. We're only vetting 6000 anomalies a day (and that's assuming that it's a good day for the human agents and that no one is sick or tired or just having an off day).
Naively we might assume that 50% of the 6000 vetted are DGAs and so we only actually catch about 30% of the zeus DGAs in our proxy logs due to the bandwidth limitations of our agents. The real picture is a little more rosy. It turns out that we could rank order which anomalies our agents look at first by their log likelihood scores. Less likely domains go to the top of the agent's queue. For the zeus case we would then detect 6000 DGAs a day (I leave it to the reader to confirm this if they don't trust me)...but 6000 is still only 60% of the zeus DGAs in our proxy logs!
Yep, cyber security sucks, but we've been able to make it suck a lot less by intelligently using our generative model to make the most efficient use of our human agents. Discriminators are much better at identifying different classes, but are terrible at making predictions outside of their training sets. I often try to (although I have not seen this in production elsewhere) create a generator to determine the probability that a new test sample was generated from the same process as my training data. If I get a decent likelihood score, I then run the sample through a discriminator. I then get the benefits of the discriminator but have an early warning system in case I suddenly start seeing domains that are unlike any domains (malignant or benign) that I have ever seen before (which may be a sign that there is a new DGA in the wild unlike any of the old ones).
I have very strong opinions about how uncertainties in modeling could be improved using statistics, but too often in practice data scientists rush to production and don't have time to create robust models that really take into account likelihood uncertainty, model uncertainty, and out-of-sample uncertainty. I'd like to one day post a small tutorial on fully accounting for uncertainty in a model.
### Let's create our own DGA!
Before we close out this fun adventure with generative models, let's look at one more cool feature of them. Since our model is generative in nature, we can actually generate fake examples of alexa1m domains. Since these are fake domains, we could actually use them as a DGA to fool DGA detectors into thinking they are Alexa1M domains!
**Don't try using this DGA in a real attack!** Firstly, there is more to creating malware than creating a decent DGA. Secondly, you will get caught and the repercussions can be HUGE. Lastly, malicious hacking is immoral, get a real job. While I love thinking about how to break through various cyber security defenses, I would never put on a black hat.
So if we aren't going to use this DGA in a cyber security attack, how might we use it? Well, we could use it in adversarial training to build a discriminator that can tell real alexa1m domains from fake alexa1m domains created using markov chains. We would need to be careful, however, because using a generator in this way can introduce major bias into this model.
Anyhoo, let's look at our sample domains...
```
def markov_dga_generator():
domain = [38]
while domain[-1] <> 39 and len(domain) <= 12:
a = domain[-1]
d = np.random.choice( [i for i in range(transition_matrix.shape[0])] , 1, p=transition_matrix[a,:] )
domain.append(d[0])
domain = ''.join( [character_lookup_dict[char] for char in domain if char not in [38,39]] )
return domain
markov_dgas = [markov_dga_generator() for i in range(20)]
markov_dgas
```
### Final Words
There's a lot more I could say about generative models, but I'll contain myself. The key to a good generative model is to describe the data as best you can, but in trying to do so you will run into all of the same problems you would with classic inference problems. Therefore, I recommend taking a decent Bayesian statistics course where you can gain familiarity with the proper way to solve these kinds of problems estimating probability distributions.
If you are looking to get some practice in with generative models, try building a generative model of alexa one million that can identify matsnu (a famous dictionary dga included in our dataset) domains as anomalies. If you want an extra hard challenge, try detecting pushdo domains as anomalies!
If you want to move past markov models as generators. I recommend looking at building mixture models or embeddings. However, Markov models are pretty great and I'll have future posts on Markov Decision Processes and Hidden markov Models (I also cover Markov switching models in my time series course).
For example, try build a domain2vec model to identify important latent variables describing a domain and then model clusters within that embedding space with a Gaussian mixture model. Alternatively, you could create an auto-encoder trained on the alexa one million data and use the reconstruction errors to identify domains that are dissimilar to alexa one million. I won't go into detail on how to implement these ideas (at least not in this blog post).
Finally, try thinking about how you would use discriminators and generators together in a practical setting. Would it be wise to feed the output of the generator into the discriminator? (Think about the case where the generator is an auto-encoder and the discriminator is a simple neural net/multi layer perceptron)
Thanks for reading; I hope you found this post interesting and helpful!
### Post Scriptum (P.S.)
So after writing this tutorial I decided to go back and play with a GMM model that was 'trained' using full bayesian inference. I won't go into detail about that this means (feel free to read my tutorial on bayesian statistics if you are curious), but I do want to show my results. The advantage of this approach (although I don't explore it here) is that we can pull model uncertainty estimates. Anyway, I base my example off of this [edward code](http://nbviewer.jupyter.org/github/blei-lab/edward/blob/master/notebooks/unsupervised.ipynb) for simplicity. Feel free to read through that notebook to build intuition around what I am doing.
One neat benefit of the approach below is that I won't have to bother with kde at all and can jsut use actual probabilities from my gaussian componenents.
Also note that edward is almsot fully integrated into tensorflow contrib, so you can just rebuild this work in tf if you so choose.
Warning! Because the GMM below is 'trained' using full bayesian inference and I make predictions using very small samples approximating the posterior, the code below yields variable results and the numbers don't always line up with my comments. If you have the time and want more reliability from the model, increase the epochs during the inference step and take more samples of the posterior when making predictions.
```
from edward.models import (
Categorical, Dirichlet, Empirical, InverseGamma,
MultivariateNormalDiag, Normal, ParamMixture)
import tensorflow as tf
import edward as ed
x_train = df_alexa1m[['LL','length']]
x_train = np.asmatrix(x_train)
N, D = x_train.shape # number of data points, dimensionality of data
K = 2 # number of components
pi = Dirichlet(tf.ones(K))
mu = Normal(tf.zeros(D), tf.ones(D), sample_shape=K)
sigmasq = InverseGamma(tf.ones(D), tf.ones(D), sample_shape=K)
x = ParamMixture(pi, {'loc': mu, 'scale_diag': tf.sqrt(sigmasq)},
MultivariateNormalDiag,
sample_shape=N)
z = x.cat
T = 500 # number of MCMC samples
qpi = Empirical(tf.get_variable(
"qpi/params", [T, K],
initializer=tf.constant_initializer(1.0 / K)))
qmu = Empirical(tf.get_variable(
"qmu/params", [T, K, D],
initializer=tf.zeros_initializer()))
qsigmasq = Empirical(tf.get_variable(
"qsigmasq/params", [T, K, D],
initializer=tf.ones_initializer()))
qz = Empirical(tf.get_variable(
"qz/params", [T, N],
initializer=tf.zeros_initializer(),
dtype=tf.int32))
inference = ed.Gibbs({pi: qpi, mu: qmu, sigmasq: qsigmasq, z: qz},
data={x: x_train})
inference.initialize()
sess = ed.get_session()
tf.global_variables_initializer().run()
t_ph = tf.placeholder(tf.int32, [])
running_cluster_means = tf.reduce_mean(qmu.params[:t_ph], 0)
for _ in range(inference.n_iter):
info_dict = inference.update()
inference.print_progress(info_dict)
t = info_dict['t']
if t % inference.n_print == 0:
print("\nInferred cluster means:")
print(sess.run(running_cluster_means, {t_ph: t - 1}))
# Make predictions for alexa domains
x_train = df_alexa1m[['LL','length']]
x_train = np.asmatrix(x_train)
N, D = x_train.shape
mu_sample = qmu.sample(100)
sigmasq_sample = qsigmasq.sample(100)
x_post = Normal(loc=tf.ones([N, 1, 1, 1]) * mu_sample,
scale=tf.ones([N, 1, 1, 1]) * tf.sqrt(sigmasq_sample))
x_broadcasted = tf.tile(tf.reshape(x_train, [N, 1, 1, D]), [1, 100, K, 1])
x_broadcasted = tf.to_float(x_broadcasted)
d = x_post.prob(x_broadcasted).eval()
# multiply out over latent variables
d = d.prod(axis=3)
# take the expectation across samples
e = d.mean(axis=1)
# Make predictions for zeus domains
x = df_dgas[df_dgas.dga_family == 'zeus'][['LL','length']]
x = np.asmatrix(x)
N, D = x.shape
x_post = Normal(loc=tf.ones([N, 1, 1, 1]) * mu_sample,
scale=tf.ones([N, 1, 1, 1]) * tf.sqrt(sigmasq_sample))
x_broadcasted = tf.tile(tf.reshape(x, [N, 1, 1, D]), [1, 100, K, 1])
x_broadcasted = tf.to_float(x_broadcasted)
d = x_post.prob(x_broadcasted).eval()
# multiply out over latent variables
d = d.prod(axis=3)
# take the expectation across samples
f = d.mean(axis=1)
# take the mean probability across mixture components
e = e.mean(axis=1)
f = f.mean(axis=1)
alexa1m_caught = 100.0 * e[e < .001].shape[0] / e.shape[0]
dgas_caught = 100.0 * f[f < .001].shape[0] / f.shape[0]
print "Percent of Zeus Domains Caught: {}% \nPercent of Alexa 1M Domains Caught: {}%".format(dgas_caught,alexa1m_caught)
```
We'll use the same assumptions as in the original post to assess efficacy.
$.999 * 10,000 = 9990$ <br>
$.001 * 10,000 = 10$ <br>
We catch 9,990 DGAs and miss 10 at the 99% credible interval.
$.003 * 9,990,000 = 29,970$ <br>
$.997 * 9,990,000 = 9,960,030$ <br>
These results are pretty neat. We basicaly get similar resutls as before (3/10 anaomalies are DGAs). Let's just try this with a different probability threshold.
```
alexa1m_caught = 100.0 * e[e < .0001].shape[0] / e.shape[0]
dgas_caught = 100.0 * f[f < .0001].shape[0] / f.shape[0]
print "Percent of Zeus Domains Caught: {}% \nPercent of Alexa 1M Domains Caught: {}%".format(dgas_caught,alexa1m_caught)
```
$.73 * 10,000 = 7,300$ <br>
$.27 * 10,000 = 2,700$ <br>
We catch 9,990 DGAs and miss 10 at the 99% credible interval.
$.00654 * 9,990,000 = 6,533$ <br>
$.99999346 * 9,990,000 = 9,983,467$ <br>
We're now getting a ratio of about 1:1 in our detected anomalies of DGAs vs Benign Domains! We're also doing this in a much simpler way than before (no need for kde). While we're not catching 100% of the zeus domains in our model, this is a much more practical framework if we are to utilize human agents to comb through our anomalies.
| github_jupyter |
<a href="https://www.bigdatauniversity.com"><img src="https://ibm.box.com/shared/static/cw2c7r3o20w9zn8gkecaeyjhgw3xdgbj.png" width="400" align="center"></a>
<h1 align="center"><font size="5">COLLABORATIVE FILTERING</font></h1>
Recommendation systems are a collection of algorithms used to recommend items to users based on information taken from the user. These systems have become ubiquitous can be commonly seen in online stores, movies databases and job finders. In this notebook, we will explore recommendation systems based on Collaborative Filtering and implement simple version of one using Python and the Pandas library.
<h1>Table of contents</h1>
<div class="alert alert-block alert-info" style="margin-top: 20px">
<ol>
<li><a href="#ref1">Acquiring the Data</a></li>
<li><a href="#ref2">Preprocessing</a></li>
<li><a href="#ref3">Collaborative Filtering</a></li>
</ol>
</div>
<br>
<hr>
<a id="ref1"></a>
# Acquiring the Data
To acquire and extract the data, simply run the following Bash scripts:
Dataset acquired from [GroupLens](http://grouplens.org/datasets/movielens/). Lets download the dataset. To download the data, we will use **`!wget`** to download it from IBM Object Storage.
__Did you know?__ When it comes to Machine Learning, you will likely be working with large datasets. As a business, where can you host your data? IBM is offering a unique opportunity for businesses, with 10 Tb of IBM Cloud Object Storage: [Sign up now for free](http://cocl.us/ML0101EN-IBM-Offer-CC)
```
!wget -O moviedataset.zip https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/ML0101ENv3/labs/moviedataset.zip
print('unziping ...')
!unzip -o -j moviedataset.zip
```
Now you're ready to start working with the data!
<hr>
<a id="ref2"></a>
# Preprocessing
First, let's get all of the imports out of the way:
```
#Dataframe manipulation library
import pandas as pd
#Math functions, we'll only need the sqrt function so let's import only that
from math import sqrt
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
```
Now let's read each file into their Dataframes:
```
#Storing the movie information into a pandas dataframe
movies_df = pd.read_csv('movies.csv')
#Storing the user information into a pandas dataframe
ratings_df = pd.read_csv('ratings.csv')
```
Let's also take a peek at how each of them are organized:
```
#Head is a function that gets the first N rows of a dataframe. N's default is 5.
movies_df.head()
```
So each movie has a unique ID, a title with its release year along with it (Which may contain unicode characters) and several different genres in the same field. Let's remove the year from the title column and place it into its own one by using the handy [extract](http://pandas.pydata.org/pandas-docs/stable/generated/pandas.Series.str.extract.html#pandas.Series.str.extract) function that Pandas has.
Let's remove the year from the __title__ column by using pandas' replace function and store in a new __year__ column.
```
#Using regular expressions to find a year stored between parentheses
#We specify the parantheses so we don't conflict with movies that have years in their titles
movies_df['year'] = movies_df.title.str.extract('(\(\d\d\d\d\))',expand=False)
#Removing the parentheses
movies_df['year'] = movies_df.year.str.extract('(\d\d\d\d)',expand=False)
#Removing the years from the 'title' column
movies_df['title'] = movies_df.title.str.replace('(\(\d\d\d\d\))', '')
#Applying the strip function to get rid of any ending whitespace characters that may have appeared
movies_df['title'] = movies_df['title'].apply(lambda x: x.strip())
```
Let's look at the result!
```
movies_df.head()
```
With that, let's also drop the genres column since we won't need it for this particular recommendation system.
```
#Dropping the genres column
movies_df = movies_df.drop('genres', 1)
```
Here's the final movies dataframe:
```
movies_df.head()
```
<br>
Next, let's look at the ratings dataframe.
```
ratings_df.head()
```
Every row in the ratings dataframe has a user id associated with at least one movie, a rating and a timestamp showing when they reviewed it. We won't be needing the timestamp column, so let's drop it to save on memory.
```
#Drop removes a specified row or column from a dataframe
ratings_df = ratings_df.drop('timestamp', 1)
```
Here's how the final ratings Dataframe looks like:
```
ratings_df.head()
```
<hr>
<a id="ref3"></a>
# Collaborative Filtering
Now, time to start our work on recommendation systems.
The first technique we're going to take a look at is called __Collaborative Filtering__, which is also known as __User-User Filtering__. As hinted by its alternate name, this technique uses other users to recommend items to the input user. It attempts to find users that have similar preferences and opinions as the input and then recommends items that they have liked to the input. There are several methods of finding similar users (Even some making use of Machine Learning), and the one we will be using here is going to be based on the __Pearson Correlation Function__.
<img src="https://ibm.box.com/shared/static/1ql8cbwhtkmbr6nge5e706ikzm5mua5w.png" width=800px>
The process for creating a User Based recommendation system is as follows:
- Select a user with the movies the user has watched
- Based on his rating to movies, find the top X neighbours
- Get the watched movie record of the user for each neighbour.
- Calculate a similarity score using some formula
- Recommend the items with the highest score
Let's begin by creating an input user to recommend movies to:
Notice: To add more movies, simply increase the amount of elements in the userInput. Feel free to add more in! Just be sure to write it in with capital letters and if a movie starts with a "The", like "The Matrix" then write it in like this: 'Matrix, The' .
```
userInput = [
{'title':'Breakfast Club, The', 'rating':5},
{'title':'Toy Story', 'rating':3.5},
{'title':'Jumanji', 'rating':2},
{'title':"Pulp Fiction", 'rating':5},
{'title':'Akira', 'rating':4.5}
]
inputMovies = pd.DataFrame(userInput)
inputMovies
```
#### Add movieId to input user
With the input complete, let's extract the input movies's ID's from the movies dataframe and add them into it.
We can achieve this by first filtering out the rows that contain the input movies' title and then merging this subset with the input dataframe. We also drop unnecessary columns for the input to save memory space.
```
#Filtering out the movies by title
inputId = movies_df[movies_df['title'].isin(inputMovies['title'].tolist())]
#Then merging it so we can get the movieId. It's implicitly merging it by title.
inputMovies = pd.merge(inputId, inputMovies)
#Dropping information we won't use from the input dataframe
inputMovies = inputMovies.drop('year', 1)
#Final input dataframe
#If a movie you added in above isn't here, then it might not be in the original
#dataframe or it might spelled differently, please check capitalisation.
inputMovies
```
#### The users who has seen the same movies
Now with the movie ID's in our input, we can now get the subset of users that have watched and reviewed the movies in our input.
```
#Filtering out users that have watched movies that the input has watched and storing it
userSubset = ratings_df[ratings_df['movieId'].isin(inputMovies['movieId'].tolist())]
userSubset.head()
```
We now group up the rows by user ID.
```
#Groupby creates several sub dataframes where they all have the same value in the column specified as the parameter
userSubsetGroup = userSubset.groupby(['userId'])
```
lets look at one of the users, e.g. the one with userID=1130
```
userSubsetGroup.get_group(1130)
```
Let's also sort these groups so the users that share the most movies in common with the input have higher priority. This provides a richer recommendation since we won't go through every single user.
```
#Sorting it so users with movie most in common with the input will have priority
userSubsetGroup = sorted(userSubsetGroup, key=lambda x: len(x[1]), reverse=True)
```
Now lets look at the first user
```
userSubsetGroup[0:3]
```
#### Similarity of users to input user
Next, we are going to compare all users (not really all !!!) to our specified user and find the one that is most similar.
we're going to find out how similar each user is to the input through the __Pearson Correlation Coefficient__. It is used to measure the strength of a linear association between two variables. The formula for finding this coefficient between sets X and Y with N values can be seen in the image below.
Why Pearson Correlation?
Pearson correlation is invariant to scaling, i.e. multiplying all elements by a nonzero constant or adding any constant to all elements. For example, if you have two vectors X and Y,then, pearson(X, Y) == pearson(X, 2 * Y + 3). This is a pretty important property in recommendation systems because for example two users might rate two series of items totally different in terms of absolute rates, but they would be similar users (i.e. with similar ideas) with similar rates in various scales .

The values given by the formula vary from r = -1 to r = 1, where 1 forms a direct correlation between the two entities (it means a perfect positive correlation) and -1 forms a perfect negative correlation.
In our case, a 1 means that the two users have similar tastes while a -1 means the opposite.
We will select a subset of users to iterate through. This limit is imposed because we don't want to waste too much time going through every single user.
```
userSubsetGroup = userSubsetGroup[0:100]
```
Now, we calculate the Pearson Correlation between input user and subset group, and store it in a dictionary, where the key is the user Id and the value is the coefficient
```
#Store the Pearson Correlation in a dictionary, where the key is the user Id and the value is the coefficient
pearsonCorrelationDict = {}
#For every user group in our subset
for name, group in userSubsetGroup:
#Let's start by sorting the input and current user group so the values aren't mixed up later on
group = group.sort_values(by='movieId')
inputMovies = inputMovies.sort_values(by='movieId')
#Get the N for the formula
nRatings = len(group)
#Get the review scores for the movies that they both have in common
temp_df = inputMovies[inputMovies['movieId'].isin(group['movieId'].tolist())]
#And then store them in a temporary buffer variable in a list format to facilitate future calculations
tempRatingList = temp_df['rating'].tolist()
#Let's also put the current user group reviews in a list format
tempGroupList = group['rating'].tolist()
#Now let's calculate the pearson correlation between two users, so called, x and y
Sxx = sum([i**2 for i in tempRatingList]) - pow(sum(tempRatingList),2)/float(nRatings)
Syy = sum([i**2 for i in tempGroupList]) - pow(sum(tempGroupList),2)/float(nRatings)
Sxy = sum( i*j for i, j in zip(tempRatingList, tempGroupList)) - sum(tempRatingList)*sum(tempGroupList)/float(nRatings)
#If the denominator is different than zero, then divide, else, 0 correlation.
if Sxx != 0 and Syy != 0:
pearsonCorrelationDict[name] = Sxy/sqrt(Sxx*Syy)
else:
pearsonCorrelationDict[name] = 0
pearsonCorrelationDict.items()
pearsonDF = pd.DataFrame.from_dict(pearsonCorrelationDict, orient='index')
pearsonDF.columns = ['similarityIndex']
pearsonDF['userId'] = pearsonDF.index
pearsonDF.index = range(len(pearsonDF))
pearsonDF.head()
```
#### The top x similar users to input user
Now let's get the top 50 users that are most similar to the input.
```
topUsers=pearsonDF.sort_values(by='similarityIndex', ascending=False)[0:50]
topUsers.head()
```
Now, let's start recommending movies to the input user.
#### Rating of selected users to all movies
We're going to do this by taking the weighted average of the ratings of the movies using the Pearson Correlation as the weight. But to do this, we first need to get the movies watched by the users in our __pearsonDF__ from the ratings dataframe and then store their correlation in a new column called _similarityIndex". This is achieved below by merging of these two tables.
```
topUsersRating=topUsers.merge(ratings_df, left_on='userId', right_on='userId', how='inner')
topUsersRating.head()
```
Now all we need to do is simply multiply the movie rating by its weight (The similarity index), then sum up the new ratings and divide it by the sum of the weights.
We can easily do this by simply multiplying two columns, then grouping up the dataframe by movieId and then dividing two columns:
It shows the idea of all similar users to candidate movies for the input user:
```
#Multiplies the similarity by the user's ratings
topUsersRating['weightedRating'] = topUsersRating['similarityIndex']*topUsersRating['rating']
topUsersRating.head()
#Applies a sum to the topUsers after grouping it up by userId
tempTopUsersRating = topUsersRating.groupby('movieId').sum()[['similarityIndex','weightedRating']]
tempTopUsersRating.columns = ['sum_similarityIndex','sum_weightedRating']
tempTopUsersRating.head()
#Creates an empty dataframe
recommendation_df = pd.DataFrame()
#Now we take the weighted average
recommendation_df['weighted average recommendation score'] = tempTopUsersRating['sum_weightedRating']/tempTopUsersRating['sum_similarityIndex']
recommendation_df['movieId'] = tempTopUsersRating.index
recommendation_df.head()
```
Now let's sort it and see the top 20 movies that the algorithm recommended!
```
recommendation_df = recommendation_df.sort_values(by='weighted average recommendation score', ascending=False)
recommendation_df.head(10)
movies_df.loc[movies_df['movieId'].isin(recommendation_df.head(10)['movieId'].tolist())]
```
### Advantages and Disadvantages of Collaborative Filtering
##### Advantages
* Takes other user's ratings into consideration
* Doesn't need to study or extract information from the recommended item
* Adapts to the user's interests which might change over time
##### Disadvantages
* Approximation function can be slow
* There might be a low of amount of users to approximate
* Privacy issues when trying to learn the user's preferences
<h2>Want to learn more?</h2>
IBM SPSS Modeler is a comprehensive analytics platform that has many machine learning algorithms. It has been designed to bring predictive intelligence to decisions made by individuals, by groups, by systems – by your enterprise as a whole. A free trial is available through this course, available here: <a href="http://cocl.us/ML0101EN-SPSSModeler">SPSS Modeler</a>
Also, you can use Watson Studio to run these notebooks faster with bigger datasets. Watson Studio is IBM's leading cloud solution for data scientists, built by data scientists. With Jupyter notebooks, RStudio, Apache Spark and popular libraries pre-packaged in the cloud, Watson Studio enables data scientists to collaborate on their projects without having to install anything. Join the fast-growing community of Watson Studio users today with a free account at <a href="https://cocl.us/ML0101EN_DSX">Watson Studio</a>
<h3>Thanks for completing this lesson!</h3>
<h4>Author: <a href="https://ca.linkedin.com/in/saeedaghabozorgi">Saeed Aghabozorgi</a></h4>
<p><a href="https://ca.linkedin.com/in/saeedaghabozorgi">Saeed Aghabozorgi</a>, PhD is a Data Scientist in IBM with a track record of developing enterprise level applications that substantially increases clients’ ability to turn data into actionable knowledge. He is a researcher in data mining field and expert in developing advanced analytic methods like machine learning and statistical modelling on large datasets.</p>
<hr>
<p>Copyright © 2018 <a href="https://cocl.us/DX0108EN_CC">Cognitive Class</a>. This notebook and its source code are released under the terms of the <a href="https://bigdatauniversity.com/mit-license/">MIT License</a>.</p>
| github_jupyter |
```
import sys
sys.path.append('../../../')
sys.path.append('../../../examples/')
sys.path.append('../../performance_tools/')
import os
import pickle
import logging
import numpy as np
import pandas as pd
from dumb_containers import evaluate_performance
import torch
import torch.nn as nn
from torch.nn import NLLLoss, CrossEntropyLoss
from argparse import Namespace
from tqdm import tqdm
from pytorch_pretrained_bert.modeling_fine_tune import BertForPointWiseClassification
from run_classifier_dataset_utils_fine_tune import LCQMCProcessor, compute_metrics, output_modes
from run_classifier_dataset_utils_fine_tune import convert_examples_to_features_fine_tune as convert_examples_to_features
from torch.utils.data import (DataLoader, RandomSampler, SequentialSampler,
TensorDataset)
from pytorch_pretrained_bert.tokenization import BertTokenizer
FINE_TUNED_PATH = '/efs/fine_tune/lcqmc/pointwise/lcqmc_fine_tune_40_1_1e-5/'
task_name = 'lcqmc'
output_mode = output_modes[task_name]
args = Namespace(data_dir = '/efs/projects/bert_fine_tune/fine_tune/data/train_dev_test/LCQMC/processed',
bert_model = '/efs/downloads/bert/pytorch/bert_base_chinese',
max_seq_length = 40,
local_rank = -1,
eval_batch_size = 8,
do_train = False
)
logger = logging.getLogger("LCQMC_pointwise_eval")
logging.basicConfig(format = '%(asctime)s - %(levelname)s - %(name)s - %(message)s',
datefmt = '%m/%d/%Y %H:%M:%S',
level = logging.DEBUG)
device = torch.device('cuda')
processor = LCQMCProcessor()
tokenizer = BertTokenizer.from_pretrained(FINE_TUNED_PATH)
model = BertForPointWiseClassification.from_pretrained(FINE_TUNED_PATH)
model.to(device)
label_list = processor.get_labels()
num_labels = len(label_list)
# eval_examples = processor.get_dev_examples(args.data_dir)
eval_examples = processor.get_test_examples(args.data_dir)
# cached_eval_features_file = os.path.join(args.data_dir, 'test_{0}_{1}_{2}'.format(
# list(filter(None, args.bert_model.split('/'))).pop(),
# str(args.max_seq_length),
# str(task_name)))
# try:
# with open(cached_eval_features_file, "rb") as reader:
# eval_features = pickle.load(reader)
# except:
# eval_features = convert_examples_to_features(
# eval_examples, label_list, args.max_seq_length, tokenizer, output_mode)
# if args.local_rank == -1 or torch.distributed.get_rank() == 0:
# logger.info(" Saving eval features into cached file %s", cached_eval_features_file)
# with open(cached_eval_features_file, "wb") as writer:
# pickle.dump(eval_features, writer)
eval_features = convert_examples_to_features(
eval_examples, label_list, args.max_seq_length, tokenizer, output_mode)
all_input_ids_a = torch.tensor([f.input_ids_a for f in eval_features], dtype=torch.long)
all_input_mask_a = torch.tensor([f.input_mask_a for f in eval_features], dtype=torch.long)
all_segment_ids_a = torch.tensor([f.segment_ids_a for f in eval_features], dtype=torch.long)
all_input_ids_b = torch.tensor([f.input_ids_b for f in eval_features], dtype=torch.long)
all_input_mask_b = torch.tensor([f.input_mask_b for f in eval_features], dtype=torch.long)
all_segment_ids_b = torch.tensor([f.segment_ids_b for f in eval_features], dtype=torch.long)
all_label_ids = torch.tensor([f.label_id for f in eval_features], dtype=torch.long)
eval_data = TensorDataset(all_input_ids_a,
all_input_ids_b,
all_input_mask_a,
all_input_mask_b,
all_segment_ids_a,
all_segment_ids_b,
all_label_ids)
eval_sampler = SequentialSampler(eval_data)
eval_dataloader = DataLoader(eval_data, sampler=eval_sampler, batch_size=args.eval_batch_size)
model.eval()
eval_loss = 0
nb_eval_steps = 0
preds = []
out_label_ids = None
for (input_ids_a, input_ids_b,
input_mask_a, input_mask_b,
segment_ids_a, segment_ids_b,
label_ids
) in tqdm(eval_dataloader, desc="Evaluating"):
input_ids_a = input_ids_a.to(device)
input_mask_a = input_mask_a.to(device)
segment_ids_a = segment_ids_a.to(device)
input_ids_b = input_ids_b.to(device)
input_mask_b = input_mask_b.to(device)
segment_ids_b = segment_ids_b.to(device)
label_ids = label_ids.to(device)
with torch.no_grad():
logits, _, _ = model(input_ids_1 = input_ids_a,
input_ids_2 = input_ids_b,
token_type_ids_1=segment_ids_a,
token_type_ids_2=segment_ids_b,
attention_mask_1=input_mask_a,
attention_mask_2=input_mask_b,)
probs = torch.softmax(logits, dim = 1)
loss_fct = CrossEntropyLoss()
tmp_eval_loss = loss_fct(logits.view(-1, num_labels), label_ids.view(-1))
if tmp_eval_loss.mean().item() == np.inf:
logger.debug("invalid loss")
print(cos_sim, probs)
print(input_ids_a)
print(input_ids_b)
break
eval_loss += tmp_eval_loss.mean().item()
nb_eval_steps += 1
if len(preds) == 0:
preds.append(probs.detach().cpu().numpy())
out_label_ids = label_ids.detach().cpu().numpy()
else:
preds[0] = np.append(
preds[0], probs.detach().cpu().numpy(), axis=0)
out_label_ids = np.append(
out_label_ids, label_ids.detach().cpu().numpy(), axis=0)
eval_loss = eval_loss / nb_eval_steps
preds = preds[0]
probs = preds[:,1]
pred_outs = np.argmax(preds, axis=1)
result = compute_metrics(task_name, pred_outs, out_label_ids)
loss = tr_loss/global_step if args.do_train else None
result['eval_loss'] = eval_loss
# result['global_step'] = global_step
result['loss'] = loss
# output_eval_file = os.path.join(args.output_dir, "eval_results.txt")
logger.info("***** Eval results *****")
for key in sorted(result.keys()):
logger.info(" %s = %s", key, str(result[key]))
# writer.write("%s = %s\n" % (key, str(result[key])))
gt = all_label_ids.numpy()
evaluate_performance(gt, probs)
```
| github_jupyter |
```
import os
import sys
sys.path.append(os.path.dirname(os.path.abspath(os.path.dirname('../..'))))
import torch
from PIL import Image
import matplotlib.pyplot as plt
import numpy as np
from src.models.cifar10.resnet import ResNet18
from torch.nn import functional as F
if torch.cuda.is_available():
device = 'cuda'
torch.set_default_tensor_type('torch.cuda.FloatTensor')
else:
device = 'cpu'
torch.set_default_tensor_type('torch.FloatTensor')
class_name = ['airplane', 'automobile', 'bird', 'cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck']
#class_name = ['airplane', 'bird', 'car', 'cat', 'deer', 'dog', 'horse', 'monkey', 'ship', 'truck']
model = ResNet18(alpha=1).to(device)
model.load_state_dict(torch.load('./pretrained/resnet18_cifar10_gvp_model_10.pth'))
model.eval()
img_path = './test_img/cifar10/test1.png'
img = Image.open(img_path)
plt.imshow(img)
target_class = 8
import torchvision.transforms as transforms
from torch.autograd import Variable
cvt_tensor = transforms.Compose([transforms.Resize((32,32)),
transforms.ToTensor(),
transforms.Normalize((0.4914, 0.4822, 0.4465), (0.2023, 0.1994, 0.2010))])
tensor_img = cvt_tensor(img).to(device)
tensor_img = tensor_img.view(1, 3, 32,32)
tensor_img = Variable(tensor_img, requires_grad=True)
gradients = None
def hook_function(module, grad_input, grad_output):
global gradients
gradients = grad_input[0]
first_layer = list(model._modules.items())[0][1]
first_layer.register_backward_hook(hook_function)
output = model(tensor_img).to(device)
model.zero_grad()
one_hot_output = torch.cuda.FloatTensor(1, output.size()[-1]).zero_()
one_hot_output[0][target_class] = 1
output.backward(gradient=one_hot_output)
grad_img = gradients.cpu().data.numpy()[0]
def scaling(img):
img = img - np.min(img)
img = img / np.max(img)
return img
grad_img = scaling(grad_img)
grad_img = grad_img.transpose(1,2,0)
if np.max(grad_img) <= 1:
grad_img = (grad_img*255).astype(np.uint8)
plt.imshow(grad_img)
grad_times_img = gradients.cpu().data.numpy()[0] + tensor_img.detach().cpu().numpy()[0]
grad_times_img.shape
def convert_to_grayscale(im_as_arr):
grayscale_im = np.sum(np.abs(im_as_arr), axis=0)
im_max = np.percentile(grayscale_im, 99)
im_min = np.min(grayscale_im)
grayscale_im = (np.clip((grayscale_im - im_min) / (im_max - im_min), 0, 1))
grayscale_im = np.expand_dims(grayscale_im, axis=0)
return grayscale_im
grad_times_img = convert_to_grayscale(grad_times_img)
grad_times_img = grad_times_img - grad_times_img.min()
grad_times_img /= grad_times_img.max()
grad_times_img.shape
grad_times_img = np.repeat(grad_times_img, 3, axis=0)
grad_times_img = grad_times_img.transpose(1,2,0)
plt.imshow(grad_times_img)
```
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
#%matplotlib inline
from classic_envs.random_integrator import DiscRandomIntegratorEnv
from lyapunov_reachability.speculation_tabular import DefaultQAgent, ExplorerQAgent, LyapunovQAgent
from lyapunov_reachability.shortest_path.dp import SimplePIAgent
from gridworld.utils import test, play, visualize
from cplex.exceptions import CplexSolverError
import matplotlib
matplotlib.rcParams['figure.dpi'] = 300
matplotlib.rcParams['pdf.fonttype'] = 42
matplotlib.rcParams['ps.fonttype'] = 42
import matplotlib.pyplot as plt
import torch.nn as nn
import numpy as np
import pickle
import os
import seaborn as sns
sns.set()
env_name = 'integrator'
# DO NOT CHANGE THIS ----
n = 10
grid_points = 40
# You can change them ----
episode_length = 200
confidence = 0.8
name = '{}-{}-{}'.format(n, grid_points, env_name)
ans_data = np.load(os.path.join(name, 'answer', 'answer.npz'))
ans = ans_data['safety_v']
del ans_data
max_safe_set = np.sum(ans >= confidence)
```
### Load baseline log files
```
baseline_dir = os.path.join(name, 'tabular-initial')
baseline_step = int(5e6)
a = np.load(os.path.join(baseline_dir, '{}.npz'.format(int(baseline_step))))['reachability_q']
a = np.min(a, -1)
init_found = np.sum((a <= 1. - confidence) * (ans >= confidence))
init_notsafe = np.sum((a <= 1. - confidence) * (ans < confidence))
init_error = np.mean((a - ans) ** 2)
del a
```
### Set necessary parameters to load log files
```
steps = int(1e8)
improve_interval = int(1e6)
log_interval = int(5e6)
save_interval= int(5e6)
fig_kwargs = {'format': 'eps',
'dpi': 300,
'rasterized': True,
'bbox_inches': 'tight',
'pad_inches': 0,
'frameon': False,
}
# Figsize default: (6., 4.); do not change this
```
### Get statistics
```
ckpts = int(steps // save_interval)
xaxis = save_interval * np.array(range(0, ckpts+1))#(np.array(range(1, ckpts+1))-0.5)
bl_seeds = list(range(1001, 1021))
seeds = bl_seeds
bl_error = []
bl_found = []
bl_notsafe = []
bl_cover = []
for seed in seeds:
file_prev = np.load(os.path.join(name, 'tabular-initial',
'{}.npz'.format(int(baseline_step))))
map_prev = np.sum(file_prev['reachability_q'] * file_prev['policy'], -1)
del file_prev
for i in range(1, ckpts+1):
file_now = np.load(os.path.join(name, 'spec-tb-default-{}'.format(seed),
'{}.npz'.format(int(save_interval * i))))
map_now = np.sum(file_now['reachability_q'] * file_now['policy'], -1)
bl_found.append(np.sum((map_now <= 1. - confidence) * (ans >= confidence)))
bl_notsafe.append( np.sum((map_now <= 1. - confidence) * (ans < confidence)))
bl_error.append(np.mean((map_now - ans) ** 2))
bl_cover.append( np.sum((map_now <= 1. - confidence) * (map_prev <= 1. - confidence)) / np.sum(map_prev <= 1. - confidence) )
map_prev[:] = map_now[:]
del map_now, file_now
del map_prev
bl_error = np.array(bl_error).reshape((len(seeds), ckpts))
bl_found = np.array(bl_found).reshape((len(seeds), ckpts))
bl_notsafe = np.array(bl_notsafe).reshape((len(seeds), ckpts))
bl_cover = np.array(bl_cover).reshape((len(seeds), ckpts))
lyap_seeds = list(range(1001, 1021))
seeds = lyap_seeds
lyap_error = []
lyap_found = []
lyap_notsafe = []
lyap_cover = []
for seed in seeds:
file_prev = np.load(os.path.join(name, 'tabular-initial',
'{}.npz'.format(int(baseline_step))))
map_prev = np.sum(file_prev['reachability_q'] * file_prev['policy'], -1)
del file_prev
for i in range(1, ckpts+1):
file_now = np.load(os.path.join(name, 'spec-tb-lyapunov-{}'.format(seed),
'{}.npz'.format(int(save_interval * i))))
map_now = np.sum(file_now['reachability_q'] * file_now['policy'], -1)
lyap_found.append(np.sum((map_now <= 1. - confidence) * (ans >= confidence)))
lyap_notsafe.append(np.sum((map_now <= 1. - confidence) * (ans < confidence)))
lyap_error.append(np.mean((map_now - ans) ** 2))
lyap_cover.append( np.sum((map_now <= 1. - confidence) * (map_prev <= 1. - confidence)) / np.sum(map_prev <= 1. - confidence) )
map_prev[:] = map_now[:]
del map_now, file_now
del map_prev
lyap_error = np.array(lyap_error).reshape((len(seeds), ckpts))
lyap_found = np.array(lyap_found).reshape((len(seeds), ckpts))
lyap_notsafe = np.array(lyap_notsafe).reshape((len(seeds), ckpts))
lyap_cover = np.array(lyap_cover).reshape((len(seeds), ckpts))
exp_seeds = list(range(1001, 1021))
seeds = exp_seeds
exp_error = []
exp_found = []
exp_notsafe = []
exp_cover = []
for seed in seeds:
file_prev = np.load(os.path.join(name, 'tabular-initial',
'{}.npz'.format(int(baseline_step))))
map_prev = np.sum(file_prev['reachability_q'] * file_prev['policy'], -1)
del file_prev
for i in range(1, ckpts+1):
file_now = np.load(os.path.join(name, 'spec-tb-explorer-{}'.format(seed),
'{}.npz'.format(int(save_interval * i))))
map_now = np.sum(file_now['reachability_q'] * file_now['policy'], -1)
exp_found.append(np.sum((map_now <= 1. - confidence) * (ans >= confidence)))
exp_notsafe.append(np.sum((map_now <= 1. - confidence) * (ans < confidence)))
exp_error.append(np.mean((map_now - ans) ** 2))
exp_cover.append( np.sum((map_now <= 1. - confidence) * (map_prev <= 1. - confidence)) / np.sum(map_prev <= 1. - confidence) )
map_prev[:] = map_now[:]
del map_now, file_now
del map_prev
exp_error = np.array(exp_error).reshape((len(seeds), ckpts))
exp_found = np.array(exp_found).reshape((len(seeds), ckpts))
exp_notsafe = np.array(exp_notsafe).reshape((len(seeds), ckpts))
exp_cover = np.array(exp_cover).reshape((len(seeds), ckpts))
del seeds
fig, ax = plt.subplots(1, 1, sharex=True)
bl_mu = np.concatenate(([init_found], np.mean(bl_found, axis=0)), axis=0) / max_safe_set
bl_std = np.concatenate(([0], np.std(bl_found, axis=0)), axis=0) / max_safe_set
lyap_mu = np.concatenate(([init_found], np.mean(lyap_found, axis=0)), axis=0) / max_safe_set
lyap_std = np.concatenate(([0], np.std(lyap_found, axis=0)), axis=0) / max_safe_set
exp_mu = np.concatenate(([init_found], np.mean(exp_found, axis=0)), axis=0) / max_safe_set
exp_std = np.concatenate(([0], np.std(exp_found, axis=0)), axis=0) / max_safe_set
ax.fill_between(xaxis, bl_mu - bl_std, bl_mu + bl_std, alpha=0.25, color='teal')
ax.fill_between(xaxis, lyap_mu - lyap_std, lyap_mu + lyap_std, alpha=0.25, color='coral')
ax.fill_between(xaxis, exp_mu - exp_std, exp_mu + exp_std, alpha=0.25, color='mediumblue')
ax.plot(xaxis, bl_mu, label='No Lyapunov', color='teal')
ax.plot(xaxis, lyap_mu, label='LSS', color='coral')
ax.plot(xaxis, exp_mu, label='ESS', color='mediumblue')
ax.legend(ncol=3, loc='upper right')
# plt.xlabel('Steps (1 step=128 samples)')
# plt.ylabel('Ratio of safe states found')
plt.xlabel('Steps')
ax.ticklabel_format(style='sci', scilimits=(-3,4), axis='both')
plt.xlim(0, ckpts*save_interval)
plt.ylim(.1, .65)
# plt.ylim(-0.05, 1.05)
# ax.set_rasterized(True)
ax.set_rasterization_zorder(0)
fig.set_dpi(300)
fig.patch.set_alpha(0)
fig.tight_layout()
plt.savefig(os.path.join(name, '{}-spec-tb-[safe_set]over[max_safe_set].pdf'.format(env_name)), format='pdf')
fig, ax = plt.subplots(1, 1, sharex=True)
bl_mu = np.concatenate(([init_notsafe], np.mean(bl_notsafe, axis=0)), axis=0) / np.prod(ans.shape)
bl_std = np.concatenate(([0], np.std(bl_notsafe, axis=0)), axis=0) / np.prod(ans.shape)
lyap_mu = np.concatenate(([init_notsafe], np.mean(lyap_notsafe, axis=0)), axis=0) / np.prod(ans.shape)
lyap_std = np.concatenate(([0], np.std(lyap_notsafe, axis=0)), axis=0) / np.prod(ans.shape)
exp_mu = np.concatenate(([init_notsafe], np.mean(exp_notsafe, axis=0)), axis=0) / np.prod(ans.shape)
exp_std = np.concatenate(([0], np.std(exp_notsafe, axis=0)), axis=0) / np.prod(ans.shape)
ax.fill_between(xaxis, bl_mu - bl_std, bl_mu + bl_std, alpha=0.25, color='teal')
ax.fill_between(xaxis, lyap_mu - lyap_std, lyap_mu + lyap_std, alpha=0.25, color='coral')
ax.fill_between(xaxis, exp_mu - exp_std, exp_mu + exp_std, alpha=0.25, color='mediumblue')
ax.plot(xaxis, bl_mu, label='No Lyapunov', color='teal')
ax.plot(xaxis, lyap_mu, label='LSS', color='coral')
ax.plot(xaxis, exp_mu, label='ESS', color='mediumblue')
ax.legend(ncol=3, loc='upper right')
# plt.xlabel('Steps (1 step=128 samples)')
# plt.ylabel('Ratio of false-positive safe states')
plt.xlabel('Steps')
ax.ticklabel_format(style='sci', scilimits=(-3,4), axis='both')
plt.xlim(save_interval, ckpts*save_interval)
plt.ylim(-0.05, 0.20)
# ax.set_rasterized(True)
ax.set_rasterization_zorder(0)
fig.set_dpi(300)
fig.patch.set_alpha(0)
fig.tight_layout()
plt.savefig(os.path.join(name, '{}-spec-tb-[false_positive_safe_set]over[state_space].pdf'.format(env_name)), format='pdf')
fig, ax = plt.subplots(1, 1, sharex=True)
bl_mu = np.concatenate(([1], np.mean(bl_cover, axis=0)), axis=0)
bl_std = np.concatenate(([0], np.std(bl_cover, axis=0)), axis=0)
lyap_mu = np.concatenate(([1], np.mean(lyap_cover, axis=0)), axis=0)
lyap_std = np.concatenate(([0], np.std(lyap_cover, axis=0)), axis=0)
exp_mu = np.concatenate(([1], np.mean(exp_cover, axis=0)), axis=0)
exp_std = np.concatenate(([0], np.std(exp_cover, axis=0)), axis=0)
ax.fill_between(xaxis, bl_mu - bl_std, bl_mu + bl_std, alpha=0.25, color='teal')
ax.fill_between(xaxis, lyap_mu - lyap_std, lyap_mu + lyap_std, alpha=0.25, color='coral')
ax.fill_between(xaxis, exp_mu - exp_std, exp_mu + exp_std, alpha=0.25, color='mediumblue')
ax.plot(xaxis, bl_mu, label='No Lyapunov', color='teal')
ax.plot(xaxis, lyap_mu, label='LSS', color='coral')
ax.plot(xaxis, exp_mu, label='ESS', color='mediumblue')
ax.legend(ncol=2, loc='lower right')
# plt.xlabel('Steps (1 step=128 samples)')
# plt.ylabel('Ratio of safe states found')
plt.xlabel('Steps')
ax.ticklabel_format(style='sci', scilimits=(-3,4), axis='both')
plt.xlim(int(0. * save_interval), int(ckpts * save_interval))
plt.ylim(0.95, 1.05)
# ax.set_rasterized(True)
ax.set_rasterization_zorder(0)
fig.set_dpi(300)
fig.patch.set_alpha(0)
fig.tight_layout()plt.savefig(os.path.join(name, '{}-spec-tb-[cover_ratio].pdf'.format(env_name)), format='pdf')
fig, ax = plt.subplots(1, 1, sharex=True)
bl_mu = np.concatenate(([init_error], np.mean(bl_error, axis=0)), axis=0)
bl_std = np.concatenate(([0], np.std(bl_error, axis=0)), axis=0)
lyap_mu = np.concatenate(([init_error], np.mean(lyap_error, axis=0)), axis=0)
lyap_std = np.concatenate(([0], np.std(lyap_error, axis=0)), axis=0)
exp_mu = np.concatenate(([init_error], np.mean(exp_error, axis=0)), axis=0)
exp_std = np.concatenate(([0], np.std(exp_error, axis=0)), axis=0)
ax.fill_between(xaxis, bl_mu - bl_std, bl_mu + bl_std, alpha=0.25, color='teal')
ax.fill_between(xaxis, lyap_mu - lyap_std, lyap_mu + lyap_std, alpha=0.25, color='coral')
ax.fill_between(xaxis, exp_mu - exp_std, exp_mu + exp_std, alpha=0.25, color='mediumblue')
ax.plot(xaxis, bl_mu, label='No Lyapunov', color='teal')
ax.plot(xaxis, lyap_mu, label='LSS', color='coral')
ax.plot(xaxis, exp_mu, label='ESS', color='mediumblue')
ax.legend(ncol=2, loc='lower right')
# plt.xlabel('Steps (1 step=128 samples)')
# plt.ylabel('Ratio of false-positive safe states')
plt.xlabel('Steps')
ax.ticklabel_format(style='sci', scilimits=(-3,4), axis='both')
plt.xlim(save_interval, ckpts*save_interval)
#plt.ylim(-0.05, 1.05)
# ax.set_rasterized(True)
ax.set_rasterization_zorder(0)
fig.set_dpi(300)
fig.patch.set_alpha(0)
fig.tight_layout()
plt.savefig(os.path.join(name, '{}-spec-tb-[mean_square_error].pdf'.format(env_name)), format='pdf')
```
### Learning curve
```
init_safety = 0
with open(os.path.join(baseline_dir, 'log.pkl'), 'rb') as f:
data = pickle.load(f)
init_safety = data['average_safety'][-1]
del data
bl_episode_safety = []
lyap_episode_safety = []
exp_episode_safety = []
ckpts = int(steps // log_interval)
xaxis = log_interval * np.array(range(0, ckpts+1))#(np.array(range(1, ckpts+1))-0.5)
seeds = list(range(1001, 1021))
for seed in seeds:
with open(os.path.join(name, 'spec-tb-default-{}'.format(seed), 'log.pkl'), 'rb') as f:
data = pickle.load(f)
bl_episode_safety += data['average_safety']
del data
bl_episode_safety = np.array(bl_episode_safety).reshape((len(seeds), ckpts))
seeds = list(range(1001, 1021))
for seed in seeds:
with open(os.path.join(name, 'spec-tb-lyapunov-{}'.format(seed), 'log.pkl'), 'rb') as f:
data = pickle.load(f)
lyap_episode_safety += data['average_safety']
del data
lyap_episode_safety = np.array(lyap_episode_safety).reshape((len(seeds), ckpts))
seeds = list(range(1001, 1021))
for seed in seeds:
with open(os.path.join(name, 'spec-tb-explorer-{}'.format(seed), 'log.pkl'), 'rb') as f:
data = pickle.load(f)
exp_episode_safety += data['average_safety']
del data
exp_episode_safety = np.array(exp_episode_safety).reshape((len(seeds), ckpts))
fig, ax = plt.subplots(1, 1, sharex=True)
bl_mu = np.concatenate(([init_safety], np.mean(bl_episode_safety, axis=0)), axis=0)
bl_std = np.concatenate(([0], np.std(bl_episode_safety, axis=0)), axis=0)
lyap_mu = np.concatenate(([init_safety], np.mean(lyap_episode_safety, axis=0)), axis=0)
lyap_std = np.concatenate(([0], np.std(lyap_episode_safety, axis=0)), axis=0)
exp_mu = np.concatenate(([init_safety], np.mean(exp_episode_safety, axis=0)), axis=0)
exp_std = np.concatenate(([0], np.std(exp_episode_safety, axis=0)), axis=0)
ax.fill_between(xaxis, bl_mu - bl_std, bl_mu + bl_std, alpha=0.25, color='teal')
ax.fill_between(xaxis, lyap_mu - lyap_std, lyap_mu + lyap_std, alpha=0.25, color='coral')
ax.fill_between(xaxis, exp_mu - exp_std, exp_mu + exp_std, alpha=0.25, color='mediumblue')
ax.plot(xaxis, bl_mu, label='No Lyapunov', color='teal')
ax.plot(xaxis, lyap_mu, label='LSS', color='coral')
ax.plot(xaxis, exp_mu, label='ESS', color='mediumblue')
ax.plot(xaxis, confidence * np.ones((ckpts+1,)), 'r--')
ax.legend(ncol=2, loc='lower right')
# plt.xlabel('Steps (1 step=128 samples)')
# plt.ylabel('Average episode safety')
plt.xlabel('Steps')
ax.ticklabel_format(style='sci', scilimits=(-3,4), axis='both')
plt.xlim(0, ckpts*log_interval)
plt.ylim(.70, 1.05)
# plt.ylim(-0.05, 1.05)
# ax.set_rasterized(True)
ax.set_rasterization_zorder(0)
fig.set_dpi(300)
fig.patch.set_alpha(0)
fig.tight_layout()
plt.savefig(os.path.join(name, '{}-spec-tb-[train_episode_safety].pdf'.format(env_name)), format='pdf')
```
### Visualization
```
def get_reachability(name, logdir, seeds, ckpts, reshape=True, reference=None):
reachability_list = []
for seed in seeds:
tmp = []
for i in range(1, ckpts+1):
a = np.load(os.path.join(name, '{}-{}'.format(logdir, seed),
'{}.npz'.format(int(save_interval * i))))['reachability_q']
a = np.min(a, -1)
tmp.append(a)
del a
tmp = np.array(tmp)
reachability_list.append(tmp)
if reference is None:
reachability_list = np.array(reachability_list).mean(0)
else:
idx = np.argmax(reference[:, -1])
reachability_list = np.array(reachability_list)[idx, ...]
if reshape:
try:
reachability_list = reachability_list.reshape((ckpts, grid_points, grid_points))
except ValueError:
print("Reshape unavailable.")
return reachability_list
ckpts = int(steps // save_interval)
xaxis = save_interval * np.array(range(1, ckpts+1))#(np.array(range(1, ckpts+1))-0.5)
bl_list = get_reachability(name, 'spec-tb-default', bl_seeds, ckpts, reshape=True, reference=bl_found)
lyap_list = get_reachability(name, 'spec-tb-lyapunov', lyap_seeds, ckpts, reshape=True, reference=lyap_found)
exp_list = get_reachability(name, 'spec-tb-explorer', exp_seeds, ckpts, reshape=True, reference=exp_found)
idx = ckpts
fig, ax = plt.subplots(1,1)
img = plt.imshow(ans.reshape((grid_points, grid_points)) >= confidence,
cmap='inferno', extent=[.5, -.5, -1., 1.,], aspect=.5)
#img = plt.imshow(ans.reshape((grid_points, grid_points)) >= confidence, cmap='plasma', extent=[.5, -.5, -1., 1.,], aspect=.5)
ax.set_xlabel('Velocity')
ax.set_xticks(np.arange(-.5, .5+1e-3, .2))
ax.set_ylabel('Position')
ax.set_yticks(np.arange(-1., 1.+2e-3, .4))
# ax.get_yaxis().set_visible(False)
plt.clim(0., 1.)
# fig.colorbar(img)
plt.grid(False)
fig.set_dpi(300)
fig.patch.set_facecolor('none')
fig.patch.set_alpha(0)
fig.tight_layout()
ax.patch.set_facecolor('none')
ax.patch.set_alpha(0)
plt.savefig(os.path.join(name, 'integrator-spec-tb-visualize-answer.pdf'), format='pdf',
facecolor=fig.get_facecolor(), edgecolor='none', bbox_inches='tight')
fig, ax = plt.subplots(1,1)
# Show False-positive and True-positive altogether.
img = plt.imshow((1.-bl_list[idx-1] >= confidence) * (ans.reshape((grid_points, grid_points)) >= confidence)
+ (1.-bl_list[idx-1] >= confidence) * (ans.reshape((grid_points, grid_points)) < confidence) * 0.5,
cmap='inferno', extent=[.5, -.5, -1., 1.,], aspect=.5)
#img = plt.imshow((1.-bl_list[idx-1] >= confidence), cmap='plasma', extent=[.5, -.5, -1., 1.,], aspect=.5)
ax.set_xlabel('Velocity')
ax.set_xticks(np.arange(-.5, .5+1e-3, .2))
ax.set_ylabel('Position')
ax.set_yticks(np.arange(-1., 1.+2e-3, .4))
ax.get_yaxis().set_visible(False)
plt.clim(0., 1.)
# fig.colorbar(img)
plt.grid(False)
fig.set_dpi(300)
fig.patch.set_facecolor('none')
fig.patch.set_alpha(0)
fig.tight_layout()
ax.patch.set_facecolor('none')
ax.patch.set_alpha(0)
fig.savefig(os.path.join(name, 'integrator-spec-tb-visualize-baseline-{}.pdf'.format(save_interval * idx)), format='pdf',
facecolor=fig.get_facecolor(), edgecolor='none', bbox_inches='tight')
fig, ax = plt.subplots(1,1)
# Show False-positive and True-positive altogether.
img = plt.imshow((1.-lyap_list[idx-1] >= confidence) * (ans.reshape((grid_points, grid_points)) >= confidence)
+ (1.-lyap_list[idx-1] >= confidence) * (ans.reshape((grid_points, grid_points)) < confidence) * 0.5,
cmap='inferno', extent=[.5, -.5, -1., 1.,], aspect=.5)
#img = plt.imshow((1.-lyap_list[idx-1] >= confidence), cmap='plasma', extent=[.5, -.5, -1., 1.,], aspect=.5)
ax.set_xlabel('Velocity')
ax.set_xticks(np.arange(-.5, .5+1e-3, .2))
ax.set_ylabel('Position')
ax.set_yticks(np.arange(-1., 1.+2e-3, .4))
ax.get_yaxis().set_visible(False)
plt.clim(0., 1.)
# fig.colorbar(img)
plt.grid(False)
fig.set_dpi(300)
fig.patch.set_facecolor('none')
fig.patch.set_alpha(0)
fig.tight_layout()
ax.patch.set_facecolor('none')
ax.patch.set_alpha(0)
plt.savefig(os.path.join(name, 'integrator-spec-tb-visualize-lyapunov-{}.pdf'.format(save_interval * idx)),
format='pdf', facecolor=fig.get_facecolor(), edgecolor='none', bbox_inches='tight')
fig, ax = plt.subplots(1,1)
# Show False-positive and True-positive altogether.
img = plt.imshow((1.-exp_list[idx-1] >= confidence) * (ans.reshape((grid_points, grid_points)) >= confidence)
+ (1.-exp_list[idx-1] >= confidence) * (ans.reshape((grid_points, grid_points)) < confidence) * 0.5,
cmap='inferno', extent=[.5, -.5, -1., 1.,], aspect=.5)
#img = plt.imshow((1.-exp_list[idx-1] >= confidence), cmap='plasma', extent=[.5, -.5, -1., 1.,], aspect=.5)
ax.set_xlabel('Velocity')
ax.set_xticks(np.arange(-.5, .5+1e-3, .2))
ax.set_ylabel('Position')
ax.set_yticks(np.arange(-1., 1.+2e-3, .4))
ax.get_yaxis().set_visible(False)
plt.clim(0., 1.)
# fig.colorbar(img)
plt.grid(False)
fig.set_dpi(300)
fig.patch.set_facecolor('none')
fig.patch.set_alpha(0)
fig.tight_layout()
ax.patch.set_facecolor('none')
ax.patch.set_alpha(0)
plt.savefig(os.path.join(name, 'integrator-spec-tb-visualize-explorer-{}.pdf'.format(save_interval * idx)),
format='pdf', facecolor=fig.get_facecolor(), edgecolor='none', bbox_inches='tight')
```
### Trial
```
env = DiscRandomIntegratorEnv(n=n, grid_points=grid_points, seed=None)
model = ExplorerQAgent.load(os.path.join(name, 'spec-tb-lyapunov-123'), int(1e8), env=env)
def tester(env, act, trials=100, initial_state=None, gamma=1.):
count = 0
safe_run = 0.
done = False
while count < trials:
_ = env.reset()
if initial_state is not None:
env.set_state(initial_state)
st = initial_state
else:
st = env.quantize(env.state)
episode_rew = 0.
episode_safety = 1.
t = 0
while not done: #t <= episode_length and not done:
_, rew, done, info = env.step(act.step(st))
st = info['state']
episode_safety *= info['safety']
episode_rew = gamma * episode_rew + rew
t += 1
count += 1
if episode_safety > 0.:
safe_run += 1.
env.close()
return 1. * safe_run / trials
tester(env, model, trials=10, initial_state=763)
```
| github_jupyter |
# <font color='blue'>Data Science Academy - Python Fundamentos - Capítulo 3</font>
## Download: http://github.com/dsacademybr
```
# Versão da Linguagem Python
from platform import python_version
print('Versão da Linguagem Python Usada Neste Jupyter Notebook:', python_version())
```
## Exercícios - Loops e Condiconais
```
# Exercício 1 - Crie uma estrutura que pergunte ao usuário qual o dia da semana. Se o dia for igual a Domingo ou
# igual a sábado, imprima na tela "Hoje é dia de descanso", caso contrário imprima na tela "Você precisa trabalhar!"
dia = input('Qual o dia da semana?') #Função input permite ao usuário inserir informações.
if (dia == 'Domingo'): #Se o dia inserido pelo usuário for Domingo será printado na tela 'Hoje é dia de descanso'
print('Hoje é dia de descanso.')
elif (dia == 'Sabado'): #Se o dia for Sabado irá aparecer na tela 'Hoje é dia de descanso'
print('Hoje é dia de descanso.')
else: #Senão irá aparecer 'Você precisa trabalhar'
print('Você precisa trabalhar.')
# Exercício 2 - Crie uma lista de 5 frutas e verifique se a fruta 'Morango' faz parte da lista
lista = ['Morango','Abacaxi','Melão','Melancia','Pera']
for fruta in lista: #Para cada fruta na lista será analisado
if (fruta == 'Morango'): #se aparece morango
print('Morango faz parte da lista de frutas.') #Caso apareça, será printado na tela
# Exercício 3 - Crie uma tupla de 4 elementos, multiplique cada elemento da tupla por 2 e guarde os resultados em uma
# lista
tupla = (1,2,3,4)
lista = []
for elemento in tupla:
x = elemento * 2 #Cada elemento da tupla será multiplicado por 2
lista.append(x) #Cada elemento será adicionado a uma lista por meio da função append
print(lista) #Lista final
# Exercício 4 - Crie uma sequência de números pares entre 100 e 150 e imprima na tela
for sequencia in range(100,151,2): #Realiza uma sequência a partir do número 100, de dois em dois, até o número 151(número exclusivo)
print(sequencia)
# Exercício 5 - Crie uma variável chamada temperatura e atribua o valor 40. Enquanto temperatura for maior que 35,
# imprima as temperaturas na tela
temperatura = 40
while temperatura > 35:
print(temperatura)
temperatura = temperatura - 1
# Exercício 6 - Crie uma variável chamada contador = 0. Enquanto counter for menor que 100, imprima os valores na tela,
# mas quando for encontrado o valor 23, interrompa a execução do programa
contador = 0
while (contador < 100) and (contador != 23):
print(contador)
contador = contador +1
else:
print('Concluído')
# Exercício 7 - Crie uma lista vazia e uma variável com valor 4. Enquanto o valor da variável for menor ou igual a 20,
# adicione à lista, apenas os valores pares e imprima a lista
lista = list() #List é a representação tipo lista
a = 4
while (a <= 20):
lista.append(a)
a = a + 2
print(lista)
# Exercício 8 - Transforme o resultado desta função range em uma lista: range(5, 45, 2)
list(range(5,45,2)) #A função list organiza os elementos da função range em uma lista
# Exercício 9 - Faça a correção dos erros no código abaixo e execute o programa. Dica: são 3 erros.
temperatura = float(input('Qual a temperatura? '))
if temperatura > 30:
print('Vista roupas leves.')
else:
print('Busque seus casacos.')
# Exercício 10 - Faça um programa que conte quantas vezes a letra "r" aparece na frase abaixo. Use um placeholder na
# sua instrução de impressão
# “É melhor, muito melhor, contentar-se com a realidade; se ela não é tão brilhante como os sonhos, tem pelo menos a
# vantagem de existir.” (Machado de Assis)
frase = ("É melhor, muito melhor, contentar-se com a realidade; se ela não é tão brilhante como os sonhos, tem pelo menos a vantagem de existir.")
a = frase.count('r') #A função .count conta quantas vezes a letra 'r' apareceu na frase
letraR = a
print('A letra r aparece {0} vezes na frase.'.format(letraR)) #O placeholder .format indica qual elemento deve aparecer na frase por meio do índice
```
# Fim
### Obrigado - Data Science Academy - <a href="http://facebook.com/dsacademybr">facebook.com/dsacademybr</a>
| github_jupyter |
<center>
<img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/Logos/organization_logo/organization_logo.png" width="300" alt="cognitiveclass.ai logo" />
</center>
# Multiple Linear Regression
Estimated time needed: **15** minutes
## Objectives
After completing this lab you will be able to:
- Use scikit-learn to implement Multiple Linear Regression
- Create a model, train,test and use the model
<h1>Table of contents</h1>
<div class="alert alert-block alert-info" style="margin-top: 20px">
<ol>
<li><a href="#understanding-data">Understanding the Data</a></li>
<li><a href="#reading_data">Reading the Data in</a></li>
<li><a href="#multiple_regression_model">Multiple Regression Model</a></li>
<li><a href="#prediction">Prediction</a></li>
<li><a href="#practice">Practice</a></li>
</ol>
</div>
<br>
<hr>
### Importing Needed packages
```
import matplotlib.pyplot as plt
import pandas as pd
import pylab as pl
import numpy as np
%matplotlib inline
```
### Downloading Data
To download the data, we will use !wget to download it from IBM Object Storage.
```
#!wget -O FuelConsumption.csv https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/ML0101ENv3/labs/FuelConsumptionCo2.csv
```
**Did you know?** When it comes to Machine Learning, you will likely be working with large datasets. As a business, where can you host your data? IBM is offering a unique opportunity for businesses, with 10 Tb of IBM Cloud Object Storage: [Sign up now for free](http://cocl.us/ML0101EN-IBM-Offer-CC)
<h2 id="understanding_data">Understanding the Data</h2>
### `FuelConsumption.csv`:
We have downloaded a fuel consumption dataset, **`FuelConsumption.csv`**, which contains model-specific fuel consumption ratings and estimated carbon dioxide emissions for new light-duty vehicles for retail sale in Canada. [Dataset source](http://open.canada.ca/data/en/dataset/98f1a129-f628-4ce4-b24d-6f16bf24dd64?cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-ML0101EN-SkillsNetwork-20718538&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ&cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-ML0101EN-SkillsNetwork-20718538&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ&cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-ML0101EN-SkillsNetwork-20718538&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ&cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-ML0101EN-SkillsNetwork-20718538&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ)
- **MODELYEAR** e.g. 2014
- **MAKE** e.g. Acura
- **MODEL** e.g. ILX
- **VEHICLE CLASS** e.g. SUV
- **ENGINE SIZE** e.g. 4.7
- **CYLINDERS** e.g 6
- **TRANSMISSION** e.g. A6
- **FUELTYPE** e.g. z
- **FUEL CONSUMPTION in CITY(L/100 km)** e.g. 9.9
- **FUEL CONSUMPTION in HWY (L/100 km)** e.g. 8.9
- **FUEL CONSUMPTION COMB (L/100 km)** e.g. 9.2
- **CO2 EMISSIONS (g/km)** e.g. 182 --> low --> 0
<h2 id="reading_data">Reading the data in</h2>
```
df = pd.read_csv("FuelConsumption.csv")
# take a look at the dataset
df.head()
```
Lets select some features that we want to use for regression.
```
cdf = df[['ENGINESIZE','CYLINDERS','FUELCONSUMPTION_CITY','FUELCONSUMPTION_HWY','FUELCONSUMPTION_COMB','CO2EMISSIONS']]
cdf.head(9)
```
Lets plot Emission values with respect to Engine size:
```
plt.scatter(cdf.ENGINESIZE, cdf.CO2EMISSIONS, color='blue')
plt.xlabel("Engine size")
plt.ylabel("Emission")
plt.show()
```
#### Creating train and test dataset
Train/Test Split involves splitting the dataset into training and testing sets respectively, which are mutually exclusive. After which, you train with the training set and test with the testing set.
This will provide a more accurate evaluation on out-of-sample accuracy because the testing dataset is not part of the dataset that have been used to train the data. It is more realistic for real world problems.
This means that we know the outcome of each data point in this dataset, making it great to test with! And since this data has not been used to train the model, the model has no knowledge of the outcome of these data points. So, in essence, it’s truly an out-of-sample testing.
```
msk = np.random.rand(len(df)) < 0.8
train = cdf[msk]
test = cdf[~msk]
```
#### Train data distribution
```
plt.scatter(train.ENGINESIZE, train.CO2EMISSIONS, color='blue')
plt.xlabel("Engine size")
plt.ylabel("Emission")
plt.show()
```
<h2 id="multiple_regression_model">Multiple Regression Model</h2>
In reality, there are multiple variables that predict the Co2emission. When more than one independent variable is present, the process is called multiple linear regression. For example, predicting co2emission using FUELCONSUMPTION_COMB, EngineSize and Cylinders of cars. The good thing here is that Multiple linear regression is the extension of simple linear regression model.
```
from sklearn import linear_model
regr = linear_model.LinearRegression()
x = np.asanyarray(train[['ENGINESIZE','CYLINDERS','FUELCONSUMPTION_COMB']])
y = np.asanyarray(train[['CO2EMISSIONS']])
regr.fit (x, y)
# The coefficients
print ('Coefficients: ', regr.coef_)
```
As mentioned before, **Coefficient** and **Intercept** , are the parameters of the fit line.
Given that it is a multiple linear regression, with 3 parameters, and knowing that the parameters are the intercept and coefficients of hyperplane, sklearn can estimate them from our data. Scikit-learn uses plain Ordinary Least Squares method to solve this problem.
#### Ordinary Least Squares (OLS)
OLS is a method for estimating the unknown parameters in a linear regression model. OLS chooses the parameters of a linear function of a set of explanatory variables by minimizing the sum of the squares of the differences between the target dependent variable and those predicted by the linear function. In other words, it tries to minimizes the sum of squared errors (SSE) or mean squared error (MSE) between the target variable (y) and our predicted output ($\\hat{y}$) over all samples in the dataset.
OLS can find the best parameters using of the following methods:
```
- Solving the model parameters analytically using closed-form equations
- Using an optimization algorithm (Gradient Descent, Stochastic Gradient Descent, Newton’s Method, etc.)
```
<h2 id="prediction">Prediction</h2>
```
y_hat= regr.predict(test[['ENGINESIZE','CYLINDERS','FUELCONSUMPTION_COMB']])
x = np.asanyarray(test[['ENGINESIZE','CYLINDERS','FUELCONSUMPTION_COMB']])
y = np.asanyarray(test[['CO2EMISSIONS']])
print("Residual sum of squares: %.2f"
% np.mean((y_hat - y) ** 2))
# Explained variance score: 1 is perfect prediction
print('Variance score: %.2f' % regr.score(x, y))
```
**explained variance regression score:**
If $\\hat{y}$ is the estimated target output, y the corresponding (correct) target output, and Var is Variance, the square of the standard deviation, then the explained variance is estimated as follow:
$\\texttt{explainedVariance}(y, \\hat{y}) = 1 - \\frac{Var{ y - \\hat{y}}}{Var{y}}$
The best possible score is 1.0, lower values are worse.
<h2 id="practice">Practice</h2>
Try to use a multiple linear regression with the same dataset but this time use __FUEL CONSUMPTION in CITY__ and
__FUEL CONSUMPTION in HWY__ instead of FUELCONSUMPTION_COMB. Does it result in better accuracy?
```
# write your code here
regr = linear_model.LinearRegression()
x = train[['ENGINESIZE','CYLINDERS','FUELCONSUMPTION_COMB']]
y = train[['CO2EMISSIONS']]
regr.fit (x, y)
# The coefficients
print ('Coefficients: ', regr.coef_)
```
Double-click **here** for the solution.
<!-- Your answer is below:
regr = linear_model.LinearRegression()
x = np.asanyarray(train[['ENGINESIZE','CYLINDERS','FUELCONSUMPTION_CITY','FUELCONSUMPTION_HWY']])
y = np.asanyarray(train[['CO2EMISSIONS']])
regr.fit (x, y)
print ('Coefficients: ', regr.coef_)
y_= regr.predict(test[['ENGINESIZE','CYLINDERS','FUELCONSUMPTION_CITY','FUELCONSUMPTION_HWY']])
x = np.asanyarray(test[['ENGINESIZE','CYLINDERS','FUELCONSUMPTION_CITY','FUELCONSUMPTION_HWY']])
y = np.asanyarray(test[['CO2EMISSIONS']])
print("Residual sum of squares: %.2f"% np.mean((y_ - y) ** 2))
print('Variance score: %.2f' % regr.score(x, y))
-->
<h2>Want to learn more?</h2>
IBM SPSS Modeler is a comprehensive analytics platform that has many machine learning algorithms. It has been designed to bring predictive intelligence to decisions made by individuals, by groups, by systems – by your enterprise as a whole. A free trial is available through this course, available here: <a href="https://www.ibm.com/analytics/spss-statistics-software">SPSS Modeler</a>
Also, you can use Watson Studio to run these notebooks faster with bigger datasets. Watson Studio is IBM's leading cloud solution for data scientists, built by data scientists. With Jupyter notebooks, RStudio, Apache Spark and popular libraries pre-packaged in the cloud, Watson Studio enables data scientists to collaborate on their projects without having to install anything. Join the fast-growing community of Watson Studio users today with a free account at <a href="https://www.ibm.com/cloud/watson-studio">Watson Studio</a>
### Thank you for completing this lab!
## Author
Saeed Aghabozorgi
### Other Contributors
<a href="https://www.linkedin.com/in/joseph-s-50398b136/" target="_blank">Joseph Santarcangelo</a>
## Change Log
| Date (YYYY-MM-DD) | Version | Changed By | Change Description |
| ----------------- | ------- | ---------- | ---------------------------------- |
| 2020-08-27 | 2.0 | Lavanya | Moved lab to course repo in GitLab |
| | | | |
| | | | |
## <h3 align="center"> © IBM Corporation 2020. All rights reserved. <h3/>
| github_jupyter |
# Sentiment analysis using Amazon Comprehend
This notebook contains the script to perform a sentiment analysis using Amazon Comprehend.
We will run the sentiment analysis for all the clean tweets (tweets with only keywords of one category) to extract the sentiment towards that BC category.
When running this notebook, we saved the output in "AWScomprenhend_output"
```
import pandas as pd
from collections import OrderedDict
import requests
import boto3
comprehend = boto3.client('comprehend', region_name='us-east-1')
import os
path=os.getcwd()
import timeit
```
**Note that the average time to process one tweet is 0.0591 seconds**
```
###########
# LNG IUD #
###########
df = pd.read_csv('/home/ec2-user/SageMaker/CleanAndAggregateTweets/LNG-IUD_CleanTweets.txt',
sep = '\t' )
len(df.text)
start = timeit.default_timer()
dfTweet = pd.DataFrame(columns=["tweets" ,"sentiments" ,"positive" ,"negative" ,"neutral", "mixed" ])
for i in range(len(df.text)):
#print(i)
if pd.notna(df.text[i]):
res = comprehend.detect_sentiment(Text=df.text[i] , LanguageCode='en')
s = res.get('Sentiment')
p = res.get('SentimentScore')['Positive']
neg = res.get('SentimentScore')['Negative']
neu = res.get('SentimentScore')['Neutral']
mix = res.get('SentimentScore')['Mixed']
dfTweet = dfTweet.append({"tweets": df.text[i],"sentiments": s, 'positive': p, 'negative': neg,
'neutral': neu, 'mixed': mix},ignore_index=True)
stop = timeit.default_timer()
print('Time: ', stop - start)
dfTweet.to_csv(path_or_buf=path+'/'+"AWScomprenhend_output/LNG-IUD_AWScomprehend_complete.csv")
##############
# Copper IUD #
##############
df = pd.read_csv('/home/ec2-user/SageMaker/CleanAndAggregateTweets/copperIUD_CleanTweets.txt',
sep = '\t' )
len(df.text)
start = timeit.default_timer()
dfTweet = pd.DataFrame(columns=["tweets" ,"sentiments" ,"positive" ,"negative" ,"neutral", "mixed" ])
for i in range(len(df.text)):
#print(i)
if pd.notna(df.text[i]):
res = comprehend.detect_sentiment(Text=df.text[i] , LanguageCode='en')
s = res.get('Sentiment')
p = res.get('SentimentScore')['Positive']
neg = res.get('SentimentScore')['Negative']
neu = res.get('SentimentScore')['Neutral']
mix = res.get('SentimentScore')['Mixed']
dfTweet = dfTweet.append({"tweets": df.text[i],"sentiments": s, 'positive': p, 'negative': neg,
'neutral': neu, 'mixed': mix},ignore_index=True)
stop = timeit.default_timer()
print('Time: ', stop - start)
dfTweet.to_csv(path_or_buf=path+'/'+"AWScomprenhend_output/copperIUD_AWScomprehend_complete.csv")
#############
# The Patch #
############
df = pd.read_csv('/home/ec2-user/SageMaker/CleanAndAggregateTweets/Patch_CleanTweets.txt',
sep = '\t' )
len(df.text)
start = timeit.default_timer()
dfTweet = pd.DataFrame(columns=["tweets" ,"sentiments" ,"positive" ,"negative" ,"neutral", "mixed" ])
for i in range(len(df.text)):
#print(i)
if pd.notna(df.text[i]):
res = comprehend.detect_sentiment(Text=df.text[i] , LanguageCode='en')
s = res.get('Sentiment')
p = res.get('SentimentScore')['Positive']
neg = res.get('SentimentScore')['Negative']
neu = res.get('SentimentScore')['Neutral']
mix = res.get('SentimentScore')['Mixed']
dfTweet = dfTweet.append({"tweets": df.text[i],"sentiments": s, 'positive': p, 'negative': neg,
'neutral': neu, 'mixed': mix},ignore_index=True)
stop = timeit.default_timer()
print('Time: ', stop - start)
dfTweet.to_csv(path_or_buf=path+'/'+"AWScomprenhend_output/Patch_AWScomprehend_complete.csv")
############
# The Ring #
############
df = pd.read_csv('/home/ec2-user/SageMaker/CleanAndAggregateTweets/Ring_CleanTweets.txt',
sep = '\t' )
len(df.text)
start = timeit.default_timer()
dfTweet = pd.DataFrame(columns=["tweets" ,"sentiments" ,"positive" ,"negative" ,"neutral", "mixed" ])
for i in range(len(df.text)):
#print(i)
if pd.notna(df.text[i]):
res = comprehend.detect_sentiment(Text=df.text[i] , LanguageCode='en')
s = res.get('Sentiment')
p = res.get('SentimentScore')['Positive']
neg = res.get('SentimentScore')['Negative']
neu = res.get('SentimentScore')['Neutral']
mix = res.get('SentimentScore')['Mixed']
dfTweet = dfTweet.append({"tweets": df.text[i],"sentiments": s, 'positive': p, 'negative': neg,
'neutral': neu, 'mixed': mix},ignore_index=True)
stop = timeit.default_timer()
print('Time: ', stop - start)
dfTweet.to_csv(path_or_buf=path+'/'+"AWScomprenhend_output/Ring_AWScomprehend_complete.csv")
###########
# Implant #
###########
df = pd.read_csv('/home/ec2-user/SageMaker/CleanAndAggregateTweets/Implant_CleanTweets.txt',
sep = '\t' )
len(df.text)
start = timeit.default_timer()
dfTweet = pd.DataFrame(columns=["tweets" ,"sentiments" ,"positive" ,"negative" ,"neutral", "mixed" ])
for i in range(len(df.text)):
#print(i)
if pd.notna(df.text[i]):
res = comprehend.detect_sentiment(Text=df.text[i] , LanguageCode='en')
s = res.get('Sentiment')
p = res.get('SentimentScore')['Positive']
neg = res.get('SentimentScore')['Negative']
neu = res.get('SentimentScore')['Neutral']
mix = res.get('SentimentScore')['Mixed']
dfTweet = dfTweet.append({"tweets": df.text[i],"sentiments": s, 'positive': p, 'negative': neg,
'neutral': neu, 'mixed': mix},ignore_index=True)
stop = timeit.default_timer()
print('Time: ', stop - start)
dfTweet.to_csv(path_or_buf=path+'/'+"AWScomprenhend_output/Implant_AWScomprehend_complete.csv")
############
# The pill #
############
df = pd.read_csv('/home/ec2-user/SageMaker/CleanAndAggregateTweets/Pill_CleanTweets.txt',
sep = '\t' )
len(df.text)
start = timeit.default_timer()
dfTweet = pd.DataFrame(columns=["tweets" ,"sentiments" ,"positive" ,"negative" ,"neutral", "mixed" ])
for i in range(len(df.text)):
#print(i)
if pd.notna(df.text[i]):
res = comprehend.detect_sentiment(Text=df.text[i] , LanguageCode='en')
s = res.get('Sentiment')
p = res.get('SentimentScore')['Positive']
neg = res.get('SentimentScore')['Negative']
neu = res.get('SentimentScore')['Neutral']
mix = res.get('SentimentScore')['Mixed']
dfTweet = dfTweet.append({"tweets": df.text[i],"sentiments": s, 'positive': p, 'negative': neg,
'neutral': neu, 'mixed': mix},ignore_index=True)
stop = timeit.default_timer()
print('Time: ', stop - start)
dfTweet.to_csv(path_or_buf=path+'/'+"AWScomprenhend_output/Pill_AWScomprehend_complete.csv")
###########
# TheShot #
###########
df = pd.read_csv('/home/ec2-user/SageMaker/CleanAndAggregateTweets/Shot_CleanTweets.txt',
sep = '\t' )
len(df.text)
start = timeit.default_timer()
dfTweet = pd.DataFrame(columns=["tweets" ,"sentiments" ,"positive" ,"negative" ,"neutral", "mixed" ])
for i in range(len(df.text)):
#print(i)
if pd.notna(df.text[i]):
res = comprehend.detect_sentiment(Text=df.text[i] , LanguageCode='en')
s = res.get('Sentiment')
p = res.get('SentimentScore')['Positive']
neg = res.get('SentimentScore')['Negative']
neu = res.get('SentimentScore')['Neutral']
mix = res.get('SentimentScore')['Mixed']
dfTweet = dfTweet.append({"tweets": df.text[i],"sentiments": s, 'positive': p, 'negative': neg,
'neutral': neu, 'mixed': mix},ignore_index=True)
stop = timeit.default_timer()
print('Time: ', stop - start)
dfTweet.to_csv(path_or_buf=path+'/'+"AWScomprenhend_output/Shot_AWScomprehend_complete.csv")
#######
# IUD #
#######
df = pd.read_csv('/home/ec2-user/SageMaker/CleanAndAggregateTweets/IUD_CleanTweets.txt',
sep = '\t' )
len(df.text)
start = timeit.default_timer()
dfTweet = pd.DataFrame(columns=["tweets" ,"sentiments" ,"positive" ,"negative" ,"neutral", "mixed" ])
for i in range(len(df.text)):
#print(i)
if pd.notna(df.text[i]):
res = comprehend.detect_sentiment(Text=df.text[i] , LanguageCode='en')
s = res.get('Sentiment')
p = res.get('SentimentScore')['Positive']
neg = res.get('SentimentScore')['Negative']
neu = res.get('SentimentScore')['Neutral']
mix = res.get('SentimentScore')['Mixed']
dfTweet = dfTweet.append({"tweets": df.text[i],"sentiments": s, 'positive': p, 'negative': neg,
'neutral': neu, 'mixed': mix},ignore_index=True)
stop = timeit.default_timer()
print('Time: ', stop - start)
dfTweet.to_csv(path_or_buf=path+'/'+"AWScomprenhend_output/IUD_AWScomprehend_complete.csv")
```
| github_jupyter |
### Analyzing the Stroop Effect
Perform the analysis in the space below. Remember to follow [the instructions](https://docs.google.com/document/d/1-OkpZLjG_kX9J6LIQ5IltsqMzVWjh36QpnP2RYpVdPU/pub?embedded=True) and review the [project rubric](https://review.udacity.com/#!/rubrics/71/view) before submitting. Once you've completed the analysis and write up, download this file as a PDF or HTML file and submit in the next section.
(1) What is the independent variable? What is the dependent variable?
**independent variable**: congruent or incongruent condition.
**dependent variable**: Time to complete test.
(2) What is an appropriate set of hypotheses for this task? What kind of statistical test do you expect to perform? Justify your choices.
Null Hypothsis, H0 - No change in time between two reading tasks (congruent or incongruent)
Alternate Hypothesis, H1 - incongruent task take more time than congruent.
H0: μi ≤ μc (μi - population mean of incongruent values, μc - population mean of congruent values)
H1: μi > μc (μi - population mean of incongruent values, μc - population mean of congruent values)
**statistical test** dependent t-test (two tailed)
- We need to compare the means of two related groups to determine the statistically significant difference between two means.
- We are assuming distributions are nearly normal and we are comparing 2 dependent samples of data
- our sample size less than 30 and we don't know the population standard deviations
(3) Report some descriptive statistics regarding this dataset. Include at least one measure of central tendency and at least one measure of variability. The name of the data file is 'stroopdata.csv'.
```
import math
import pandas as pd
import numpy as np
import seaborn as sns
from scipy.stats import t
%matplotlib inline
df = pd.read_csv('./stroopdata.csv')
print(df.mean(axis=0))
print(df.std(axis=0))
print("standard deviation for congruent {0:.3f}".format(np.std(df['Congruent'].values)))
print("standard deviation for Incongruent {0:.3f}".format(np.std(df['Incongruent'].values)))
```
(4) Provide one or two visualizations that show the distribution of the sample data. Write one or two sentences noting what you observe about the plot or plots.
```
sns.distplot(df['Congruent'])
```
The data is more or less normally distributed and the middle of the data is a little bit less than 15
```
sns.distplot(df['Incongruent'])
```
There are some interesting data points on the upper end of this distribution that skew it right
The histogram plots, although both graphs visually appear somewhat positively skewed,
the mean is pretty close to the peak in both graphs which would indicate a normal distribution
(5) Now, perform the statistical test and report the results. What is the confidence level and your critical statistic value? Do you reject the null hypothesis or fail to reject it? Come to a conclusion in terms of the experiment task. Did the results match up with your expectations?
```
#Sample size
print(df['Congruent'].size)
print(df['Incongruent'].size)
#t-critical value for a 95% confidence level and 23 d.f.
t.ppf(.95, 23)
```
For a confidence level of 95% and 23 degrees of freedom, our t-critical value ends up being *1.7139*
Our point estimate for the difference of the means is: 22.02 - 14.05 = 7.97
Our standard deviation of the differences is calculated below.
```
df['Difference'] = df['Congruent'] - df['Incongruent']
print("standard deviation for congruent {0:.4f}".format(df['Difference'].std(axis=0)))
```
### t-statistic:
```
7.97/(4.8648 / math.sqrt(24))
```
Our t-statistic (**8.02**) is greater than our critical value (**1.7139**),So we can **reject the null hypothesis**.
Which matches up with what we expected, That it takes much less time to do the congruent task than it does to do the incongruent task.
## References
https://en.wikipedia.org/wiki/Stroop_effect
http://www.statisticshowto.com/when-to-use-a-t-score-vs-z-score/
| github_jupyter |
# Задание 1.1 - Метод К-ближайших соседей (K-neariest neighbor classifier)
В первом задании вы реализуете один из простейших алгоритмов машинного обучения - классификатор на основе метода K-ближайших соседей.
Мы применим его к задачам
- бинарной классификации (то есть, только двум классам)
- многоклассовой классификации (то есть, нескольким классам)
Так как методу необходим гиперпараметр (hyperparameter) - количество соседей, мы выберем его на основе кросс-валидации (cross-validation).
Наша основная задача - научиться пользоваться numpy и представлять вычисления в векторном виде, а также ознакомиться с основными метриками, важными для задачи классификации.
Перед выполнением задания:
- запустите файл `download_data.sh`, чтобы скачать данные, которые мы будем использовать для тренировки
- установите все необходимые библиотеки, запустив `pip install -r requirements.txt` (если раньше не работали с `pip`, вам сюда - https://pip.pypa.io/en/stable/quickstart/)
Если вы раньше не работали с numpy, вам может помочь tutorial. Например этот:
http://cs231n.github.io/python-numpy-tutorial/
```
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
%load_ext autoreload
%autoreload 2
from dataset import load_svhn
from knn import KNN
from metrics import binary_classification_metrics, multiclass_accuracy
```
# Загрузим и визуализируем данные
В задании уже дана функция `load_svhn`, загружающая данные с диска. Она возвращает данные для тренировки и для тестирования как numpy arrays.
Мы будем использовать цифры из датасета Street View House Numbers (SVHN, http://ufldl.stanford.edu/housenumbers/), чтобы решать задачу хоть сколько-нибудь сложнее MNIST.
```
train_X, train_y, test_X, test_y = load_svhn("data", max_train=1000, max_test=100)
samples_per_class = 5 # Number of samples per class to visualize
plot_index = 1
for example_index in range(samples_per_class):
for class_index in range(10):
plt.subplot(5, 10, plot_index)
image = train_X[train_y == class_index][example_index]
plt.imshow(image.astype(np.uint8))
plt.axis('off')
plot_index += 1
```
# Сначала реализуем KNN для бинарной классификации
В качестве задачи бинарной классификации мы натренируем модель, которая будет отличать цифру 0 от цифры 9.
```
# First, let's prepare the labels and the source data
# Only select 0s and 9s
binary_train_mask = (train_y == 0) | (train_y == 9)
binary_train_X = train_X[binary_train_mask]
binary_train_y = train_y[binary_train_mask] == 0
binary_test_mask = (test_y == 0) | (test_y == 9)
binary_test_X = test_X[binary_test_mask]
binary_test_y = test_y[binary_test_mask] == 0
# Reshape to 1-dimensional array [num_samples, 3*3*32]
binary_train_X = binary_train_X.reshape(binary_train_X.shape[0], -1)
binary_test_X = binary_test_X.reshape(binary_test_X.shape[0], -1)
# Create the classifier and call fit to train the model
# KNN just remembers all the data
knn_classifier = KNN(k=1)
knn_classifier.fit(binary_train_X, binary_train_y)
```
## Пришло время написать код!
Последовательно реализуйте функции `compute_distances_two_loops`, `compute_distances_one_loop` и `compute_distances_no_loops`
в файле `knn.py`
```
# TODO: implement compute_distances_two_loops in knn.py
dists = knn_classifier.compute_distances_two_loops(binary_test_X)
assert np.isclose(dists[0, 10], np.sum(np.abs(binary_test_X[0] - binary_train_X[10])))
# TODO: implement compute_distances_one_loop in knn.py
dists = knn_classifier.compute_distances_one_loop(binary_test_X)
assert np.isclose(dists[0, 10], np.sum(np.abs(binary_test_X[0] - binary_train_X[10])))
# TODO: implement compute_distances_no_loops in knn.py
dists = knn_classifier.compute_distances_no_loops(binary_test_X)
assert np.isclose(dists[0, 10], np.sum(np.abs(binary_test_X[0] - binary_train_X[10])))
# Lets look at the performance difference
%timeit knn_classifier.compute_distances_two_loops(binary_test_X)
%timeit knn_classifier.compute_distances_one_loop(binary_test_X)
%timeit knn_classifier.compute_distances_no_loops(binary_test_X)
# TODO: implement predict_labels_binary in knn.py
prediction = knn_classifier.predict(binary_test_X)
# TODO: implement binary_classification_metrics in metrics.py
precision, recall, f1, accuracy = binary_classification_metrics(prediction, binary_test_y)
print("KNN with k = %s" % knn_classifier.k)
print("Accuracy: %4.2f, Precision: %4.2f, Recall: %4.2f, F1: %4.2f" % (accuracy, precision, recall, f1))
# Let's put everything together and run KNN with k=3 and see how we do
knn_classifier_3 = KNN(k=3)
knn_classifier_3.fit(binary_train_X, binary_train_y)
prediction = knn_classifier_3.predict(binary_test_X)
precision, recall, f1, accuracy = binary_classification_metrics(prediction, binary_test_y)
print("KNN with k = %s" % knn_classifier_3.k)
print("Accuracy: %4.2f, Precision: %4.2f, Recall: %4.2f, F1: %4.2f" % (accuracy, precision, recall, f1))
```
# Кросс-валидация (cross-validation)
Попробуем найти лучшее значение k!
Для этого мы воспользуемся k-fold cross-validation (https://en.wikipedia.org/wiki/Cross-validation_(statistics)#k-fold_cross-validation). Мы разделим тренировочные данные на 5 фолдов (folds), и по очереди будем использовать каждый из них в качестве проверочных данных (validation data), а остальные -- в качестве тренировочных (training data).
В качестве финальной оценки эффективности k мы усредним значения F1 score на всех фолдах.
После этого мы просто выберем значение k с лучшим значением метрики.
*Бонус*: есть ли другие варианты агрегировать F1 score по всем фолдам? Напишите плюсы и минусы в клетке ниже.
```
# Find the best k using cross-validation based on F1 score
num_folds = 5
train_folds_X = []
train_folds_y = []
# TODO: split the training data in 5 folds and store them in train_folds_X/train_folds_y
k_choices = [1, 2, 3, 5, 8, 10, 15, 20, 25, 50]
k_to_f1 = {} # dict mapping k values to mean F1 scores (int -> float)
for k in k_choices:
# TODO: perform cross-validation
# Go through every fold and use it for testing and all other folds for validation
# Perform training and produce F1 score metric on the validation dataset
# Average F1 from all the folds and write it into k_to_f1
pass
for k in sorted(k_to_f1):
print('k = %d, f1 = %f' % (k, k_to_f1[k]))
```
### Проверим, как хорошо работает лучшее значение k на тестовых данных (test data)
```
# TODO Set the best k to the best value found by cross-validation
best_k = 1
best_knn_classifier = KNN(k=best_k)
best_knn_classifier.fit(binary_train_X, binary_train_y)
prediction = best_knn_classifier.predict(binary_test_X)
precision, recall, f1, accuracy = binary_classification_metrics(prediction, binary_test_y)
print("Best KNN with k = %s" % best_k)
print("Accuracy: %4.2f, Precision: %4.2f, Recall: %4.2f, F1: %4.2f" % (accuracy, precision, recall, f1))
```
# Многоклассовая классификация (multi-class classification)
Переходим к следующему этапу - классификации на каждую цифру.
```
# Now let's use all 10 classes
train_X = train_X.reshape(train_X.shape[0], -1)
test_X = test_X.reshape(test_X.shape[0], -1)
knn_classifier = KNN(k=1)
knn_classifier.fit(train_X, train_y)
# TODO: Implement predict_labels_multiclass
predict = knn_classifier.predict(test_X)
# TODO: Implement multiclass_accuracy
accuracy = multiclass_accuracy(predict, test_y)
print("Accuracy: %4.2f" % accuracy)
```
Снова кросс-валидация. Теперь нашей основной метрикой стала точность (accuracy), и ее мы тоже будем усреднять по всем фолдам.
```
# Find the best k using cross-validation based on accuracy
num_folds = 5
train_folds_X = []
train_folds_y = []
# TODO: split the training data in 5 folds and store them in train_folds_X/train_folds_y
k_choices = [1, 2, 3, 5, 8, 10, 15, 20, 25, 50]
k_to_accuracy = {}
for k in k_choices:
# TODO: perform cross-validation
# Go through every fold and use it for testing and all other folds for validation
# Perform training and produce accuracy metric on the validation dataset
# Average accuracy from all the folds and write it into k_to_accuracy
pass
for k in sorted(k_to_accuracy):
print('k = %d, accuracy = %f' % (k, k_to_accuracy[k]))
```
### Финальный тест - классификация на 10 классов на тестовой выборке (test data)
Если все реализовано правильно, вы должны увидеть точность не менее **0.2**.
```
# TODO Set the best k as a best from computed
best_k = 1
best_knn_classifier = KNN(k=best_k)
best_knn_classifier.fit(train_X, train_y)
prediction = best_knn_classifier.predict(test_X)
# Accuracy should be around 20%!
accuracy = multiclass_accuracy(prediction, test_y)
print("Accuracy: %4.2f" % accuracy)
```
| github_jupyter |
### Manipulating Volumes
If we manipulate the image data, for example a crop or flip, we need to update the affine matrix as well. If not, the image geometry will be wrong and this could be dangerous for future use of the data.
Authors: David Atkinson
First version: 20 June 2021
CCP SyneRBI Synergistic Image Reconstruction Framework (SIRF).
Copyright 2021 University College London.
This is software developed for the Collaborative Computational Project in Synergistic Reconstruction for Biomedical Imaging (http://www.ccpsynerbi.ac.uk/).
Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0 Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
```
# Setup the working directory for the notebook
import notebook_setup
from sirf_exercises import cd_to_working_dir
cd_to_working_dir('Geometry')
# Import required packages
import nibabel
import matplotlib.pyplot as plt
import numpy as np
import os
import sirf.Reg as Reg
# Data for geometry notebooks when run is ./nifti/*.nii
data_path = os.getcwd()
# Set numpy print options to print small numbers as zero etc.
np.set_printoptions(precision=3,suppress=True)
%matplotlib notebook
def vdisp(ax, vol, A, falpha, frame, ocmap='gray'):
# 3D Display of volume
# sdisp(ax, s_ido, falpha, frame, ocmap='gray')
# ax axes predefined
# vol array of volume data
# A 4x4 affine matrix
# falpha face alpha (0-1)
# frame frame number (0-based)
# ocmap colormap defaults to gray
#
# Calculates the vertices of pixels and uses to create a surface with transparency
# falpha and intensities corresponding to pixwl values
img = vol[:,:,frame]
nrow = img.shape[0]
ncol = img.shape[1]
L = np.zeros((nrow+1, ncol+1)) # allocate memory
P = np.zeros((nrow+1, ncol+1)) # +1 because this is for vertices
H = np.zeros((nrow+1, ncol+1))
for ir in range(0,nrow+1):
for ic in range(0,ncol+1):
# VLPH are LPH patient coordinates corresponding to
# pixel vertices, which are at image coords -0.5, 0.5, 1.5, ...
VLPH = np.matmul(A, np.array([ [ir-0.5], [ic-0.5], [frame], [1] ]))
L[ir,ic] = VLPH[0] # separate the components for surf plot
P[ir,ic] = VLPH[1]
H[ir,ic] = VLPH[2]
scamap = plt.cm.ScalarMappable(cmap=ocmap)
fcolors = scamap.to_rgba(img, alpha=falpha)
ax.plot_surface(L, P, H, facecolors=fcolors, cmap=ocmap, linewidth=0, rcount=100, ccount=100)
ax.set_xlabel('Left')
ax.set_ylabel('Posterior')
ax.set_zlabel('Head')
# Read in data
# Get the affine matrix from NIfTI and convert to LPH
fpath = os.path.join(data_path , 'nifti')
fn_cor = "OBJECT_phantom_T2W_TSE_Cor_14_1.nii" # Coronal volume, 30 slices
ffn = os.path.join(fpath, fn_cor) # full file name
s_imd = Reg.ImageData(ffn) # SIRF ImageData object
vol = s_imd.as_array() # SIRF array (the volume)
s_geom_info = s_imd.get_geometrical_info()
A_LPH = s_geom_info.get_index_to_physical_point_matrix() # 4x4 affine matrix
print(A_LPH)
print(vol.shape)
# Find the 3D coordinate of the offset point
Q = np.matmul(A_LPH,[0,0,0,1])
print(Q)
```
We are going to create a new volume from the central 200x200x20 region.
Will the spacing change?
Will the orientations change?
Will the offset change?
```
# Calculate the image coordinates of the region we are going to extract.
# There might be more elegant ways of doing this, but Python has made a mess of
# rounding and division in its various versions, so this is supposed to be clear
fov = np.array([200, 200, 20]) # new field of view in pixels
center0b = np.floor(np.array(vol.shape) / 2.0) # 0-based coordinate of centre
hw = np.floor(fov/2.0) # half width of new fov
lim = (center0b - hw).astype(int) # Python ....
# Extract new volume from old
volnew = vol[lim[0]:lim[0]+fov[0], lim[1]:lim[1]+fov[1], lim[2]:lim[2]+fov[2]]
print(volnew.shape)
# Calculate the new offset in 3D LPH
# lim is in 0-based units
Qnew = np.matmul(A_LPH, [lim[0],lim[1],lim[2],1])
# The new A is the same as the old, excet for the updated offset
Anew = np.array([[A_LPH[0,0], A_LPH[0,1], A_LPH[0,2], Qnew[0]],
[A_LPH[1,0], A_LPH[1,1], A_LPH[1,2], Qnew[1]],
[A_LPH[2,0], A_LPH[2,1], A_LPH[2,2], Qnew[2]],
[A_LPH[3,0], A_LPH[3,1], A_LPH[3,2], Qnew[3]] ])
print(A_LPH)
print(Anew)
fig = plt.figure() # Open figure and get 3D axes (can rotate with mouse)
ax = plt.axes(projection='3d')
vdisp(ax, volnew, Anew, 0.6, 10, 'gray')
vdisp(ax, vol, A_LPH, 0.2, 15, 'gray')
```
The figure above shows that the cropped region comes correctly from the original.
Now lets look at an example of flipping the second dimension.
Will the spacing change?
Will the orientations change?
Will the offset change?
```
# The new offset will be at the position of the last voxel in the 2nd dimension in the original colume
Qnew = np.matmul(A_LPH, [0, vol.shape[1]-1, 0, 1])
print(Qnew)
# The new A will use the updated offset and swap the sign of the 2nd column as this
# corresponds to the 2nd array index.
Anew = np.array([[A_LPH[0,0], -A_LPH[0,1], A_LPH[0,2], Qnew[0]],
[A_LPH[1,0], -A_LPH[1,1], A_LPH[1,2], Qnew[1]],
[A_LPH[2,0], -A_LPH[2,1], A_LPH[2,2], Qnew[2]],
[A_LPH[3,0], A_LPH[3,1], A_LPH[3,2], Qnew[3]] ])
# Flip the volume in the 2nd dimension (1 in 0-based units)
volnew = np.flip(vol, axis=1)
ax = plt.figure()
slc = 15
plt.subplot(1,2,1, title='original')
plt.imshow(vol[:,:,slc])
plt.xticks([]), plt.yticks([])
plt.subplot(1,2,2, title='flipped in second dimension')
plt.imshow(volnew[:,:,slc])
plt.xticks([]), plt.yticks([])
# Although flipped, we have correctly updated the geometry:
fig = plt.figure() # Open figure and get 3D axes (can rotate with mouse)
ax = plt.axes(projection='3d')
vdisp(ax, volnew, Anew, 0.6, slc, 'gray')
vdisp(ax, vol, A_LPH, 0.2, slc, 'hot')
```
Despite the flipped orientation in the array, the images coincide because they are the same slice and are correctly positioned in 3D space.
Possible Exercises:
Flip in the first dimension.
Apply a 90 degree rotation about the 3rd dimension axis (a simple rotation)
| github_jupyter |
```
# IPYTHON_TEST_SKIP_REMAINDER
#
# This flag causes execution to skip the remainder of the notebook in the Jenkins test suite.
# This is because the functionality for plotting the spectra etc. below seems to be broken and
# needs to be revisited.
```
# Computing normal modes using the ringdown and eigenvalue method
This notebook illustrates how to use the class `NormalModeSimulation` to perform normal mode computations. This class contains a number of convenience methods which make it easy to compute eigenmodes of a system using either the ringdown method (where the system is excited using a short field pulse, for example) or the "analytical" eigenvalue method.
```
from finmag import normal_mode_simulation, sim_with
from finmag.util.meshes import nanodisk
```
## Creating and relaxing a simulation
First we create a simulation and bring it into a relaxed state. We start by creating a mesh of a nanodisk with diameter 60 nm and height 10 nm. For illustration purposes (so that the calculations won't take too long) we choose a relatively coarse mesh discretisation of `maxh` = 5 nm. In a real simulation this should probably be chosen smaller (e.g. `maxh` = 3 nm).
```
d = 60
h = 10
maxh = 5.0
mesh = nanodisk(d, h, maxh)
print mesh
```
Next we define the material parameters for Permalloy, and choose an initial magnetisation from which to start the simulation (which in our case points along the x-axis). We also choose a large value `alpha=1.0` for the damping in order to speed up the relaxation. Finally, we apply an external field during the relaxation phase which points just slightly off the x-axis. In the ringdown phase below this field will be re-set to point exactly along the x-axis. This causes the magnetisation to precess, and this precession will be recorded and analysed to compute the normal modes.
```
# Material parameters for Permalloy
Ms = 8e5
A=13e-12
m_init = [1, 0, 0]
alpha_relax = 1.0
H_ext_relax = [1e5, 1e3, 0]
```
Now we create the simulation with the parameters defined above.
```
sim = normal_mode_simulation(mesh, Ms, m_init, alpha=alpha_relax, unit_length=1e-9, A=A, H_ext=H_ext_relax)
```
... and relax it (in the presence of the external field).
```
sim.relax()
```
This is what the relaxed state looks like (with the magnetisation essentially pointing along the field axis).
```
sim.render_scene()
#sim.render_scene(use_display=0) # use_display=0 results in pop-up window, but seems more robust (problem on osiris.sesnet)
```
Now we redefine some parameters and define a few new ones which define the behaviour during the ringdown phase.
- We choose a small value of `alpha` so that the precession is nearly undamped (which is important to obtain well-defined peaks in the Fourier spectrum).
- We set the external field so that it points exactly along the x-axis. This introduces a small excitation into the system which we can use to measure the eigenmodes.
- We also save the averaged magnetisation every 10 ps, as well as snapshots of the spatially resolved magnetisation every 10 ps. These will be used to compute the spectrum and export animations of the eigenmodes to VTK.
```
alpha_ringdown = 0.01
H_ext_ringdown = [1e5, 0, 0]
t_end = 1e-9
save_ndt_every = 1e-11
save_m_every = 1e-11
m_snapshots_filename = 'snapshots/m_ringdown.npy'
```
Now we run the actual ringdown. This is as simple as calling the function `sim.run_ringdown` with the parameters defined above.
```
sim.run_ringdown(t_end=t_end, alpha=alpha_ringdown, H_ext=H_ext_ringdown, save_ndt_every=save_ndt_every,
save_m_every=save_m_every, m_snapshots_filename=m_snapshots_filename, overwrite=True)
```
Let's have a look at the dynamics of the averaged magnetisation during the ringdown. We only plot the components `m_y` and `m_z` because `m_x` remains virtually unchanged. We can achieve this by calling the helper method `sim.plot_dynamics`, which automatically reads the data from the `.ndt` file that is associated with the simulation (this only works because we told it to save averaged magnetisation values to this file during the ringdown phase above).
```
_ = sim.plot_dynamics('yz')
```
We can also plot the FFT spectrum by calling `sim.plot_spectrum`. Again, this automatically reads the data from the `.ndt` file which was saved during the ringdown phase above. The resolution and maximum frequency depend on the length of the simulation (defined by the parameter `t_end` above) and the interval at which the magnetisation was saved (defined by `save_ndt_every`). Since we only ran the simulation for 1 ns, the frequency resolution is relatively coarse here.
```
_ = sim.plot_spectrum(figsize=(16, 4), components='y')
```
We notice a single peak above, and would like to visualise the mode by exporting a VTK file which we can animate in Paraview. In order to do this, we first need to find the exact location of the peak. It seems to be somewhere near 10 GHz, so we use this as a first approximation. The helper function `sim.find_peak_near_frequency` can be used to determine the exact location, as well as the index of the peak in the array of frequencies. This function also expects an argument specifying the magnetisation component ('x', 'y', or 'z') in whose spectrum to search for a peak. We use 'y' here because that ones seems to have the most prominent peak.
```
peak_freq, peak_idx = sim.find_peak_near_frequency(10e9, 'y')
print "Found peak at {:.3f} GHz (index in FFT array: {})".format(peak_freq / 1e9, peak_idx)
```
In order to have a visual confirmation whether the peak was detected correctly, let's use the helper function `sim.plot_peak_near_frequency` (which accepts the same parameters as `sim.find_peak_near_frequency`).
```
_ = sim.plot_peak_near_frequency(10e9, 'y', figsize=(16, 4))
```
We can now export an animation of the eigenmode corresponding to that peak. The helper function `sim.export_normal_mode_animation_from_ringdown` does this for us. We again give it the approximate frequency and component (these have the same meaning as in the functions discussed previously, as well as an output directory (it is also possible to specify a filename directly). We also tell it to animate one cycle of the oscillation using 10 frames.
```
sim.export_normal_mode_animation_from_ringdown(\
'snapshots/m_ringdown_*.npy', f_approx=10e-9, component='y', directory='animations/ringdown',
num_cycles=1, num_frames_per_cycle=10)
```
To confirm, the output directory now contains a bunch of VTK files which can be imported into Paraview to inspect the mode in detail.
```
ls -1 animations/ringdown/
```
## Computing normal modes using the eigenvalue method
An alternative to the ringdown method is to compute the normal modes using an analytical eigenvalue approach as described in reference [1]. All we need to do is to tell the system how many eigenvalues we want to compute. Optionally, it is possible specify the tolerance, and whether to save the computed matrices for later use (neither of which we are doing here).
[1] D'Aquino et al., "A novel formulation for the numerical computation of magnetization modes in complex micromagnetic systems", J Comp Phys 228, 17, 6130–6149 (2009).
```
omega, w = sim.compute_normal_modes(n_values=20)
```
The computed normal mode frequencies (in GHz) are returned in the vector `omega` (sorted from smallest to largest magnitude). Note that for each mode there is also a corresponding mode at the negative frequency. However, these are discarded by default because they do not contain any additional information.
```
print omega
```
The corresponding eigenvectors (which encode the "shapes" of the normal mode) are returned in the vector `w`. But rather than looking at it directly, we directly export an animation of the first eigenmode, which we can then animate in Paraview.
```
sim.export_normal_mode_animation(0, directory='animations/eigenvalue', num_cycles=1, num_snapshots_per_cycle=10)
```
Again, to confirm that it worked let's list the output files.
```
ls -1 animations/eigenvalue/
```
| github_jupyter |
# Advanced topic: Climate sensitivity and feedback
This notebook is part of [The Climate Laboratory](https://brian-rose.github.io/ClimateLaboratoryBook) by [Brian E. J. Rose](http://www.atmos.albany.edu/facstaff/brose/index.html), University at Albany.
____________
<a id='section1'></a>
## 1. Radiative forcing
____________
Let’s say we instantaneously double atmospheric CO$_2$. What happens?
- The atmosphere is less efficient at radiating energy away to space.
- OLR will decrease
- The climate system will begin gaining energy.
We will call this abrupt decrease in OLR the **radiative forcing**, a positive number in W m$^{-2}$
$$ \Delta R = -\Delta OLR$$
$\Delta R$ is a measure of the rate at which energy begins to accumulate in the climate system after an abrupt increase in greenhouse gases, but *before any change in climate* (i.e. temperature).
What happens next?
____________
<a id='section2'></a>
## 2. Climate sensitivity (without feedback)
____________
Let’s use our simple zero-dimensional EBM to calculate the resulting change in **equilibrium temperature**.
How much warming will we get once the climate system has adjusted to the radiative forcing?
First note that at equilibrium we must have
$$ASR = OLR$$
and in our very simple model, there is no change in ASR, so
$$ ASR_f = ASR_f $$
(with standing for final.)
From this we infer that
$$ OLR_f = OLR_i $$
The new equilibrium will have **exactly the same OLR** as the old equilibrium, but a **different surface temperature**.
The climate system must warm up by a certain amount to get the OLR back up to its original value! The question is, **how much warming is necessary**? In other words, **what is the new equilibrium temperature**?
### Equilibrium Climate Sensitivity (ECS)
We now define the Equilibrium Climate Sensitivity (denoted ECS or $\Delta T_{2xCO2}$):
*The global mean surface warming necessary to balance the planetary energy budget after a doubling of atmospheric CO$_2$.*
The temperature must increase so that the increase in OLR is exactly equal to the radiative forcing:
$$ OLR_f - OLR_{2xCO2} = \Delta R $$
From the [lecture on analytical solutions of the global EBM](https://brian-rose.github.io/ClimateLaboratoryBook/courseware/analytical-efolding.html), we have linearized our model for OLR with a slope $\lambda_0 = 3.3$ W m$^{-2}$ K$^{-1}$. This means that a global warming of 1 degree causes a 3.3 W m$^{-2}$ increase in the OLR. So we can write:
$$OLR_f \approx OLR_{2xCO2} + \lambda_0 \Delta T_0 $$
where we are writing the change in temperature as
$$ \Delta T_0 = T_f - T_i $$
(and the subscript zero will remind us that this is the response in the simplest model, in the absence of any feedbacks)
To achieve energy balance, the planet must warm up by
$$ \Delta T_0 = \frac{\Delta R}{\lambda_0} $$
As we will see later, the actual radiative forcing due CO$_2$ doubling is about 4 W m$^{-2}$.
So our model without feedback gives a prediction for climate sensitivity:
```
# Repeating code from Lecture 2
sigma = 5.67E-8 # Stefan-Boltzmann constant in W/m2/K4
Q = 341.3 # global mean insolation in W/m2
alpha = 101.9 / Q # observed planetary albedo
Te = ((1-alpha)*Q/sigma)**0.25 # Emission temperature (definition)
Tsbar = 288. # global mean surface temperature in K
beta = Te / Tsbar # Calculate value of beta from observations
lambda_0 = 4 * sigma * beta**4 * Tsbar**3
DeltaR = 4. # Radiative forcing in W/m2
DeltaT0 = DeltaR / lambda_0
print( 'The Equilibrium Climate Sensitivity in the absence of feedback is {:.1f} K.'.format(DeltaT0))
```
Question: what are the current best estimates for the actual warming (including all feedbacks) in response to a doubling of CO$_2$?
We’ll now look at the feedback concept. Climate feedbacks tend to amplify the response to increased CO$_2$. But $\Delta T_0$ is a meaningful climate sensitivity in the absence of feedback.
$\Delta T_0 = 1.2$ K is the **warming that we would have if the Earth radiated the excess energy away to space as a blackbody**, and with no change in the planetary albedo.
____________
<a id='section3'></a>
## 3. The feedback concept
____________
A concept borrowed from electrical engineering. You have all heard or used the term before, but we’ll try take a more precise approach today.
A feedback occurs when a portion of the output from the action of a system is added to the input and subsequently alters the output:

The result of a loop system can either be **amplification** or **dampening** of the process, depending on the sign of the gain in the loop, which we will denote $f$.
We will call amplifying feedbacks **positive** ($f>0$) and damping feedbacks **negative** ($f<0$).
We can think of the “process” here as the entire climate system, which contains many examples of both positive and negative feedback.
### Example: the water vapor feedback
The capacity of the atmosphere to hold water vapor (saturation specific humidity) increases exponentially with temperature. Warming is thus accompanied by moistening (more water vapor), which leads to more warming due to the enhanced water vapor greenhouse effect.
**Positive or negative feedback?**
### Example: the ice-albedo feedback
Colder temperatures lead to expansion of the areas covered by ice and snow, which tend to be more reflective than water and vegetation. This causes a reduction in the absorbed solar radiation, which leads to more cooling.
**Positive or negative feedback?**
*Make sure it’s clear that the sign of the feedback is the same whether we are talking about warming or cooling.*
_____________
<a id='section4'></a>
## 4. Climate feedback: some definitions
____________
We start with an initial radiative forcing , and get a response
$$ \Delta T_0 = \frac{\Delta R}{\lambda_0} $$
Now consider what happens in the presence of a feedback process. For a concrete example, let’s take the **water vapor feedback**. For every degree of warming, there is an additional increase in the greenhouse effect, and thus additional energy added to the system.
Let’s denote this extra energy as
$$ f \lambda_0 \Delta T_0 $$
where $f$ is the **feedback amount**, a number that represents what fraction of the output gets added back to the input. $f$ must be between $-\infty$ and +1.
For the example of the water vapor feedback, $f$ is positive (between 0 and +1) – the process adds extra energy to the original radiative forcing.
The amount of energy in the full "input" is now
$$ \Delta R + f \lambda_0 \Delta T_0 $$
or
$$ (1+f) \lambda_0 \Delta T_0 $$
But now we need to consider the next loop. A fraction $f$ of the additional energy is also added to the input, giving us
$$ (1+f+f^2) \lambda_0 \Delta T_0 $$
and we can go round and round, leading to the infinite series
$$ (1+f+f^2+f^3+ ...) \lambda_0 \Delta T_0 = \lambda_0 \Delta T_0 \sum_{n=0}^{\infty} f^n $$
Question: what happens if $f=1$?
It so happens that this infinite series has an exact solution
$$ \sum_{n=0}^{\infty} f^n = \frac{1}{1-f} $$
So the full response including all the effects of the feedback is actually
$$ \Delta T = \frac{1}{1-f} \Delta T_0 $$
This is also sometimes written as
$$ \Delta T = g \Delta T_0 $$
where
$$ g = \frac{1}{1-f} = \frac{\Delta T}{\Delta T_0} $$
is called the **system gain** -- the ratio of the actual warming (including all feedbacks) to the warming we would have in the absence of feedbacks.
So if the overall feedback is positive, then $f>0$ and $g>1$.
And if the overall feedback is negative?
_____________
<a id='section5'></a>
## 5. Climate sensitivity with feedback
____________
ECS is an important number. A major goal of climate modeling is to provide better estimates of ECS and its uncertainty.
Latest IPCC report AR5 gives a likely range of 1.5 to 4.5 K. (There is lots of uncertainty in these numbers – we will definitely come back to this question)
So our simple estimate of the no-feedback change $\Delta T_0$ is apparently underestimating climate sensitivity.
Saying the same thing another way: the overall net climate feedback is positive, amplifying the response, and the system gain $g>1$.
Let’s assume that the true value is $\Delta T_{2xCO2} = 3$ K (middle of the range). This implies that the gain is
$$ g = \frac{\Delta T_{2xCO2}}{\Delta T_0} = \frac{3}{1.2} = 2.5 $$
The actual warming is substantially amplified!
There are lots of reasons for this, but the water vapor feedback is probably the most important.
Question: if $g=2.5$, what is the feedback amount $f$?
$$ g = \frac{1}{1-f} $$
or rearranging,
$$ f = 1 - 1/g = 0.6 $$
The overall feedback (due to water vapor, clouds, etc.) is **positive**.
_____________
<a id='section6'></a>
## 6. Contribution of individual feedback processes to Equilibrium Climate Sensitivity
____________
Now what if we have several individual feedback processes occurring simultaneously?
We can think of individual feedback amounts $f_1, f_2, f_3, ...$, with each representing a physically distinct mechanism, e.g. water vapor, surface snow and ice, cloud changes, etc.
Each individual process takes a fraction $f_i$ of the output and adds to the input. So the feedback amounts are additive,
$$ f = f_1 + f_2 + f_3 + ... = \sum_{i=0}^N f_i $$
This gives us a way to compare the importance of individual feedback processes!
The climate sensitivity is now
$$ \Delta T_{2xCO2} = \frac{1}{1- \sum_{i=0}^N f_i } \Delta T_0 $$
The climate sensitivity is thus **increased by positive feedback processes**, and **decreased by negative feedback processes**.
### Climate feedback parameters
We can also write this in terms of the original radiative forcing as
$$ \Delta T_{2xCO2} = \frac{\Delta R}{\lambda_0 - \sum_{i=1}^{N} \lambda_i} $$
where
$$ \lambda_i = \lambda_0 f_i $$
known as **climate feedback parameters**, in units of W m$^{-2}$ K$^{-1}$.
With this choice of sign conventions, $\lambda_i > 0$ for a positive feedback process.
Individual feedback parameters $\lambda_i$ are then additive, and can be compared to the no-feedback parameter $\lambda_0$.
Based on our earlier numbers, the net feedback necessary to get a climate sensitivity of 3 K is
$$ \sum_{i=1}^N \lambda_i = \lambda_0 \sum_{i=1}^N f_i = (3.3 \text{ W m}^{-2} \text{ K}^{-1}) (0.6) = 2 \text{ W m}^{-2} \text{ K}^{-1} $$
We might decompose this net climate feedback into, for example
- longwave and shortwave processes
- cloud and non-cloud processes
These individual feedback processes may be positive or negative. This is very powerful, because we can **measure the relative importance of different feedback processes** simply by comparing their $\lambda_i$ values.
### Every climate model has a Planck feedback
The "Planck feedback" represented by the negative of our reference parameter $-\lambda_0$ is not really a feedback at all.
It is the most basic and universal climate process, and is present in every climate model. It is simply an expression of the fact that a warm planet radiates more to space than a cold planet.
As we will see, our estimate of $-\lambda_0 = -3.3 ~\text{W} ~\text{m}^{-2} ~\text{K}^{-1} $ is essentially the same as the Planck feedback diagnosed from complex GCMs. Unlike our simple zero-dimensional EBM, however, most other climate models (and the real climate system) have other radiative feedback processes, such that
$$\lambda = \lambda_0 - \sum_{i=1}^{N} \lambda_i \ne \lambda_0 $$
____________
<a id='section7'></a>
## 7. Feedbacks diagnosed from complex climate models
____________
### Data from the IPCC AR5
This figure is reproduced from the recent IPCC AR5 report. It shows the feedbacks diagnosed from the various models that contributed to the assessment.
(Later in the term we will discuss how the feedback diagnosis is actually done)
See below for complete citation information.
<img src='http://www.climatechange2013.org/images/figures/WGI_AR5_Fig9-43.jpg' width=800>
**Figure 9.43** | (a) Strengths of individual feedbacks for CMIP3 and CMIP5 models (left and right columns of symbols) for Planck (P), water vapour (WV), clouds (C), albedo (A), lapse rate (LR), combination of water vapour and lapse rate (WV+LR) and sum of all feedbacks except Planck (ALL), from Soden and Held (2006) and Vial et al. (2013), following Soden et al. (2008). CMIP5 feedbacks are derived from CMIP5 simulations for abrupt fourfold increases in CO2 concentrations (4 × CO2). (b) ECS obtained using regression techniques by Andrews et al. (2012) against ECS estimated from the ratio of CO2 ERF to the sum of all feedbacks. The CO2 ERF is one-half the 4 × CO2 forcings from Andrews et al. (2012), and the total feedback (ALL + Planck) is from Vial et al. (2013).
*Figure caption reproduced from the AR5 WG1 report*
Legend:
- P: Planck feedback
- WV: Water vapor feedback
- LR: Lapse rate feedback
- WV+LR: combined water vapor plus lapse rate feedback
- C: cloud feedback
- A: surface albedo feedback
- ALL: sum of all feedback except Plank, i.e. ALL = WV+LR+C+A
Things to note:
- The models all agree strongly on the Planck feedback.
- The Planck feedback is about $-\lambda_0 = -3.3 ~\text{W} ~\text{m}^{-2} ~\text{K}^{-1} $ just like our above estimate (but with opposite sign convention -- watch carefully for that in the literature)
- The water vapor feedback is strongly positive in every model.
- The lapse rate feedback is something we will study later. It is slightly negative.
- For reasons we will discuss later, the best way to measure the water vapor feedback is to combine it with lapse rate feedback.
- Models agree strongly on the combined water vapor plus lapse rate feedback.
- The albedo feedback is slightly positive but rather small globally.
- By far the largest spread across the models occurs in the cloud feedback.
- Global cloud feedback ranges from slighly negative to strongly positive across the models.
- Most of the spread in the total feedback is due to the spread in the cloud feedback.
- Therefore, most of the spread in the ECS across the models is due to the spread in the cloud feedback.
- Our estimate of $+2.0 ~\text{W} ~\text{m}^{-2} ~\text{K}^{-1}$ for all the missing processes is consistent with the GCM ensemble.
### Citation
This is Figure 9.43 from Chapter 9 of the IPCC AR5 Working Group 1 report.
The report and images can be found online at
<http://www.climatechange2013.org/report/full-report/>
The full citation is:
Flato, G., J. Marotzke, B. Abiodun, P. Braconnot, S.C. Chou, W. Collins, P. Cox, F. Driouech, S. Emori, V. Eyring, C. Forest, P. Gleckler, E. Guilyardi, C. Jakob, V. Kattsov, C. Reason and M. Rummukainen, 2013: Evaluation of Climate Models. In: Climate Change 2013: The Physical Science Basis. Contribution of Working Group I to the Fifth Assessment Report of the Intergovernmental Panel on Climate Change [Stocker, T.F., D. Qin, G.-K. Plattner, M. Tignor, S.K. Allen, J. Boschung, A. Nauels, Y. Xia, V. Bex and P.M. Midgley (eds.)]. Cambridge University Press, Cambridge, United Kingdom and New York, NY, USA, pp. 741–866, doi:10.1017/CBO9781107415324.020
____________
<a id='section8'></a>
## 8. The zero-dimensional model with variable albedo
____________
### The model
In homework you will be asked to include a new process in the zero-dimensional EBM: a temperature-dependent albedo.
We use the following formula:
$$ \alpha(T) = \left\{ \begin{array}{ccc}
\alpha_i & & T \le T_i \\
\alpha_o + (\alpha_i-\alpha_o) \frac{(T-T_o)^2}{(T_i-T_o)^2} & & T_i < T < T_o \\
\alpha_o & & T \ge T_o \end{array} \right\}$$
with parameter values:
- $\alpha_o = 0.289$ is the albedo of a warm, ice-free planet
- $\alpha_i = 0.7$ is the albedo of a very cold, completely ice-covered planet
- $T_o = 293$ K is the threshold temperature above which our model assumes the planet is ice-free
- $T_i = 260$ K is the threshold temperature below which our model assumes the planet is completely ice covered.
For intermediate temperature, this formula gives a smooth variation in albedo with global mean temperature. It is tuned to reproduce the observed albedo $\alpha = 0.299$ for $T = 288$ K.
### Coding up the model in Python
This largely repeats what I asked you to do in your homework.
```
%matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
def albedo(T, alpha_o = 0.289, alpha_i = 0.7, To = 293., Ti = 260.):
alb1 = alpha_o + (alpha_i-alpha_o)*(T-To)**2 / (Ti - To)**2
alb2 = np.where(T>Ti, alb1, alpha_i)
alb3 = np.where(T<To, alb2, alpha_o)
return alb3
def ASR(T, Q=341.3):
alpha = albedo(T)
return Q * (1-alpha)
def OLR(T, sigma=5.67E-8, tau=0.57):
return tau * sigma * T**4
def Ftoa(T):
return ASR(T) - OLR(T)
T = np.linspace(220., 300., 100)
plt.plot(T, albedo(T))
plt.xlabel('Temperature (K)')
plt.ylabel('albedo')
plt.ylim(0,1)
plt.title('Albedo as a function of global mean temperature')
```
### Graphical solution: TOA fluxes as functions of temperature
```
plt.plot(T, OLR(T), label='OLR')
plt.plot(T, ASR(T), label='ASR')
plt.plot(T, Ftoa(T), label='Ftoa')
plt.xlabel('Surface temperature (K)')
plt.ylabel('TOA flux (W m$^{-2}$)')
plt.grid()
plt.legend(loc='upper left')
```
The graphs meet at three different points! That means there are actually three different possible equilibrium temperatures in this model.
### Numerical solution to get the three equilibrium temperatures
```
# Use numerical root-finding to get the equilibria
from scipy.optimize import brentq
# brentq is a root-finding function
# Need to give it a function and two end-points
# It will look for a zero of the function between those end-points
Teq1 = brentq(Ftoa, 280., 300.)
Teq2 = brentq(Ftoa, 260., 280.)
Teq3 = brentq(Ftoa, 200., 260.)
print( Teq1, Teq2, Teq3)
```
### Bonus exercise
Using numerical timestepping and different initial temperatures, can you get the model to converge on all three equilibria, or only some of them?
What do you think this means?
____________
## Credits
This notebook is part of [The Climate Laboratory](https://brian-rose.github.io/ClimateLaboratoryBook), an open-source textbook developed and maintained by [Brian E. J. Rose](http://www.atmos.albany.edu/facstaff/brose/index.html), University at Albany.
It is licensed for free and open consumption under the
[Creative Commons Attribution 4.0 International (CC BY 4.0)](https://creativecommons.org/licenses/by/4.0/) license.
Development of these notes and the [climlab software](https://github.com/brian-rose/climlab) is partially supported by the National Science Foundation under award AGS-1455071 to Brian Rose. Any opinions, findings, conclusions or recommendations expressed here are mine and do not necessarily reflect the views of the National Science Foundation.
____________
| github_jupyter |
```
import torch
import torch.nn as nn
import time
import argparse
import os
import datetime
import torch.nn.functional as F
import random
from torch.distributions.categorical import Categorical
import math
import numpy as np
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from torch.autograd import Variable
from torch.optim import lr_scheduler
import matplotlib.pyplot as plt
import time
from tqdm import tqdm_notebook
from tqdm import tqdm_notebook
import math
import numpy as np
import torch
import tqdm
import torch.nn as nn
import torch.nn.functional as F
from torch.utils.data import Dataset
import matplotlib.pyplot as plt
from scipy.spatial import distance
# visualization
%matplotlib inline
from IPython.display import set_matplotlib_formats, clear_output
import matplotlib_inline
matplotlib_inline.backend_inline.set_matplotlib_formats('png2x','pdf')
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import warnings
warnings.filterwarnings("ignore", category=UserWarning)
device = torch.device("cpu"); gpu_id = -1 # select CPU
gpu_id = '0' # select a single GPU
#gpu_id = '2,3' # select multiple GPUs
os.environ["CUDA_VISIBLE_DEVICES"] = str(gpu_id)
if torch.cuda.is_available():
device = torch.device("cuda")
print('GPU name: {:s}, gpu_id: {:s}'.format(torch.cuda.get_device_name(0),gpu_id))
print(device)
```
# HPN Large
```
def compute_tour_length(x, tour):
"""
Compute the length of a batch of tours
Inputs : x of size (bsz, nb_nodes, 2) batch of tsp tour instances
tour of size (bsz, nb_nodes) batch of sequences (node indices) of tsp tours
Output : L of size (bsz,) batch of lengths of each tsp tour
"""
bsz = x.shape[0]
nb_nodes = x.shape[1]
arange_vec = torch.arange(bsz, device=x.device)
first_cities = x[arange_vec, tour[:,0], :] # size(first_cities)=(bsz,2)
previous_cities = first_cities
L = torch.zeros(bsz, device=x.device)
with torch.no_grad():
for i in range(1,nb_nodes):
current_cities = x[arange_vec, tour[:,i], :]
L += torch.sum( (current_cities - previous_cities)**2 , dim=1 )**0.5 # dist(current, previous node)
previous_cities = current_cities
L += torch.sum((current_cities - first_cities)**2 , dim=1)**0.5 # dist(last, first node)
return L
class TransEncoderNet(nn.Module):
"""
Encoder network based on self-attention transformer
Inputs :
h of size (bsz, nb_nodes+1, dim_emb) batch of input cities
Outputs :
h of size (bsz, nb_nodes+1, dim_emb) batch of encoded cities
score of size (bsz, nb_nodes+1, nb_nodes+1) batch of attention scores
"""
def __init__(self, nb_layers, dim_emb, nb_heads, dim_ff, batchnorm):
super(TransEncoderNet, self).__init__()
assert dim_emb == nb_heads* (dim_emb//nb_heads) # check if dim_emb is divisible by nb_heads
self.MHA_layers = nn.ModuleList( [nn.MultiheadAttention(dim_emb, nb_heads) for _ in range(nb_layers)] )
self.linear1_layers = nn.ModuleList( [nn.Linear(dim_emb, dim_ff) for _ in range(nb_layers)] )
self.linear2_layers = nn.ModuleList( [nn.Linear(dim_ff, dim_emb) for _ in range(nb_layers)] )
if batchnorm:
self.norm1_layers = nn.ModuleList( [nn.BatchNorm1d(dim_emb) for _ in range(nb_layers)] )
self.norm2_layers = nn.ModuleList( [nn.BatchNorm1d(dim_emb) for _ in range(nb_layers)] )
else:
self.norm1_layers = nn.ModuleList( [nn.LayerNorm(dim_emb) for _ in range(nb_layers)] )
self.norm2_layers = nn.ModuleList( [nn.LayerNorm(dim_emb) for _ in range(nb_layers)] )
self.nb_layers = nb_layers
self.nb_heads = nb_heads
self.batchnorm = batchnorm
def forward(self, h):
# PyTorch nn.MultiheadAttention requires input size (seq_len, bsz, dim_emb)
h = h.transpose(0,1) # size(h)=(nb_nodes, bsz, dim_emb)
# L layers
for i in range(self.nb_layers):
h_rc = h # residual connection, size(h_rc)=(nb_nodes, bsz, dim_emb)
h, score = self.MHA_layers[i](h, h, h) # size(h)=(nb_nodes, bsz, dim_emb), size(score)=(bsz, nb_nodes, nb_nodes)
# add residual connection
h = h_rc + h # size(h)=(nb_nodes, bsz, dim_emb)
if self.batchnorm:
# Pytorch nn.BatchNorm1d requires input size (bsz, dim, seq_len)
h = h.permute(1,2,0).contiguous() # size(h)=(bsz, dim_emb, nb_nodes)
h = self.norm1_layers[i](h) # size(h)=(bsz, dim_emb, nb_nodes)
h = h.permute(2,0,1).contiguous() # size(h)=(nb_nodes, bsz, dim_emb)
else:
h = self.norm1_layers[i](h) # size(h)=(nb_nodes, bsz, dim_emb)
# feedforward
h_rc = h # residual connection
h = self.linear2_layers[i](torch.relu(self.linear1_layers[i](h)))
h = h_rc + h # size(h)=(nb_nodes, bsz, dim_emb)
if self.batchnorm:
h = h.permute(1,2,0).contiguous() # size(h)=(bsz, dim_emb, nb_nodes)
h = self.norm2_layers[i](h) # size(h)=(bsz, dim_emb, nb_nodes)
h = h.permute(2,0,1).contiguous() # size(h)=(nb_nodes, bsz, dim_emb)
else:
h = self.norm2_layers[i](h) # size(h)=(nb_nodes, bsz, dim_emb)
# Transpose h
h = h.transpose(0,1) # size(h)=(bsz, nb_nodes, dim_emb)
return h, score
class Attention(nn.Module):
def __init__(self, n_hidden):
super(Attention, self).__init__()
self.size = 0
self.batch_size = 0
self.dim = n_hidden
v = torch.FloatTensor(n_hidden).cuda()
self.v = nn.Parameter(v)
self.v.data.uniform_(-1/math.sqrt(n_hidden), 1/math.sqrt(n_hidden))
# parameters for pointer attention
self.Wref = nn.Linear(n_hidden, n_hidden)
self.Wq = nn.Linear(n_hidden, n_hidden)
def forward(self, q, ref): # query and reference
self.batch_size = q.size(0)
self.size = int(ref.size(0) / self.batch_size)
q = self.Wq(q) # (B, dim)
ref = self.Wref(ref)
ref = ref.view(self.batch_size, self.size, self.dim) # (B, size, dim)
q_ex = q.unsqueeze(1).repeat(1, self.size, 1) # (B, size, dim)
# v_view: (B, dim, 1)
v_view = self.v.unsqueeze(0).expand(self.batch_size, self.dim).unsqueeze(2)
# (B, size, dim) * (B, dim, 1)
u = torch.bmm(torch.tanh(q_ex + ref), v_view).squeeze(2)
return u, ref
class LSTM(nn.Module):
def __init__(self, n_hidden):
super(LSTM, self).__init__()
# parameters for input gate
self.Wxi = nn.Linear(n_hidden, n_hidden) # W(xt)
self.Whi = nn.Linear(n_hidden, n_hidden) # W(ht)
self.wci = nn.Linear(n_hidden, n_hidden) # w(ct)
# parameters for forget gate
self.Wxf = nn.Linear(n_hidden, n_hidden) # W(xt)
self.Whf = nn.Linear(n_hidden, n_hidden) # W(ht)
self.wcf = nn.Linear(n_hidden, n_hidden) # w(ct)
# parameters for cell gate
self.Wxc = nn.Linear(n_hidden, n_hidden) # W(xt)
self.Whc = nn.Linear(n_hidden, n_hidden) # W(ht)
# parameters for forget gate
self.Wxo = nn.Linear(n_hidden, n_hidden) # W(xt)
self.Who = nn.Linear(n_hidden, n_hidden) # W(ht)
self.wco = nn.Linear(n_hidden, n_hidden) # w(ct)
def forward(self, x, h, c): # query and reference
# input gate
i = torch.sigmoid(self.Wxi(x) + self.Whi(h) + self.wci(c))
# forget gate
f = torch.sigmoid(self.Wxf(x) + self.Whf(h) + self.wcf(c))
# cell gate
c = f * c + i * torch.tanh(self.Wxc(x) + self.Whc(h))
# output gate
o = torch.sigmoid(self.Wxo(x) + self.Who(h) + self.wco(c))
h = o * torch.tanh(c)
return h, c
class HPN(nn.Module):
def __init__(self, n_feature, n_hidden):
super(HPN, self).__init__()
self.city_size = 0
self.batch_size = 0
self.dim = n_hidden
# pointer layer
self.pointer = Attention(n_hidden)
self.TransPointer = Attention(n_hidden)
# lstm encoder
self.encoder = LSTM(n_hidden)
# trainable first hidden input
h0 = torch.FloatTensor(n_hidden)
c0 = torch.FloatTensor(n_hidden)
self.h0 = nn.Parameter(h0)
self.c0 = nn.Parameter(c0)
self.h0.data.uniform_(-1/math.sqrt(n_hidden), 1/math.sqrt(n_hidden))
self.c0.data.uniform_(-1/math.sqrt(n_hidden), 1/math.sqrt(n_hidden))
r1 = torch.ones(1)
r2 = torch.ones(1)
r3 = torch.ones(1)
self.r1 = nn.Parameter(r1)
self.r2 = nn.Parameter(r2)
self.r3 = nn.Parameter(r3)
# embedding
self.embedding_x = nn.Linear(n_feature, n_hidden)
self.embedding_all1 = nn.Linear(n_feature, n_hidden)
self.embedding_all2 = nn.Linear(n_feature + 1, n_hidden)
self.Transembedding_all = TransEncoderNet(6, 128, 8, 512, batchnorm=True)
# vector to start decoding
self.start_placeholder = nn.Parameter(torch.randn(n_hidden))
# weights for GNN
self.W1 = nn.Linear(n_hidden, n_hidden)
self.W2 = nn.Linear(n_hidden, n_hidden)
self.W3 = nn.Linear(n_hidden, n_hidden)
# aggregation function for GNN
self.agg_1 = nn.Linear(n_hidden, n_hidden)
self.agg_2 = nn.Linear(n_hidden, n_hidden)
self.agg_3 = nn.Linear(n_hidden, n_hidden)
def forward(self, Transcontext, x, X_all, mask, h=None, c=None, latent=None):
'''
Inputs (B: batch size, size: city size, dim: hidden dimension)
x: current city coordinate (B, 2)
X_all: all cities' cooridnates (B, size, 2)
mask: mask visited cities
h: hidden variable (B, dim)
c: cell gate (B, dim)
latent: latent pointer vector from previous layer (B, size, dim)
Outputs
softmax: probability distribution of next city (B, size)
h: hidden variable (B, dim)
c: cell gate (B, dim)
latent_u: latent pointer vector for next layer
'''
self.batch_size = X_all.size(0)
self.city_size = X_all.size(1)
# Check if this iteration is the first one
if h is None or c is None:
# Letting the placeholder be the first input
x = self.start_placeholder
# init-embedding for All Cities
context = self.embedding_all1(X_all)
# Transormer context
Transcontext,_ = self.Transembedding_all(context)
Transcontext = Transcontext.reshape(-1, self.dim) # (B, size, dim)
# =============================
# handling the cell and the hidden state for the first iteration
# =============================
h0 = self.h0.unsqueeze(0).expand(self.batch_size, self.dim)
c0 = self.c0.unsqueeze(0).expand(self.batch_size, self.dim)
h0 = h0.unsqueeze(0).contiguous()
c0 = c0.unsqueeze(0).contiguous()
# let h0, c0 be the hidden variable of first turn
h = h0.squeeze(0)
c = c0.squeeze(0)
else:
# =============================
# Feature context
# =============================
X_all = torch.cat((torch.cdist(X_all,x.view(self.batch_size,1,2),p=2), X_all - x.unsqueeze(1).repeat(1, self.city_size, 1)), 2)
# sequential input Embedding
x = self.embedding_x(x)
# init-embedding for All Cities
context = self.embedding_all2(X_all)
# =============================
# graph neural network encoder
# =============================
# Handling contextes's size
context = context.reshape(-1, self.dim) # (B, size, dim)
context = self.r1 * self.W1(context) + (1-self.r1) * F.relu(self.agg_1(context/(self.city_size-1)))
context = self.r2 * self.W2(context) + (1-self.r2) * F.relu(self.agg_2(context/(self.city_size-1)))
context = self.r3 * self.W3(context) + (1-self.r3) * F.relu(self.agg_3(context/(self.city_size-1)))
# LSTM encoder
h, c = self.encoder(x, h, c)
# =============================
# Decoding Phase
# =============================
u1, _ = self.pointer(h, context)
u2 ,_ = self.TransPointer(h,Transcontext)
u = u1 + u2
latent_u = u.clone()
u = 100 * torch.tanh(u) + mask
return Transcontext,F.softmax(u, dim=1), h, c, latent_u
size = 50
TOL = 1e-3
TINY = 1e-15
learn_rate = 1e-3 # learning rate
B = 128 # batch_size
B_val = 64 # validation size
B_valLoop = 20
size_val = 500
steps = 2500 # training steps
n_epoch = 100 # epochs
print('=========================')
print('prepare to train')
print('=========================')
print('Hyperparameters:')
print('size', size)
print('size_val', size_val)
print('learning rate', learn_rate)
print('batch size', B)
print('validation size', B_val)
print('steps', steps)
print('epoch', n_epoch)
print('=========================')
###################
# Instantiate a training network and a baseline network
###################
try:
del Actor # remove existing model
del Critic # remove existing model
except:
pass
Actor = HPN(n_feature=2, n_hidden=128)
Critic = HPN(n_feature=2, n_hidden=128)
optimizer = optim.Adam(Actor.parameters(), lr=learn_rate)
lr_decay_step = 2500
lr_decay_rate = 0.96
opt_scheduler = lr_scheduler.MultiStepLR(optimizer, range(lr_decay_step, lr_decay_step*1000,lr_decay_step), gamma=lr_decay_rate)
# Putting Critic model on the eval mode
Actor = Actor.to(device)
Critic = Critic.to(device)
Critic.eval()
########################
# Remember to first initialize the model and optimizer, then load the dictionary locally.
#######################
epoch_ckpt = 0
tot_time_ckpt = 0
plot_performance_train = []
plot_performance_baseline = []
#********************************************# Uncomment these lines to re-start training with saved checkpoint #********************************************#
#checkpoint_file = "../input/hpnlarge487e/checkpoint_21-08-08--09-15-31-n50-gpu0.pkl"
#checkpoint = torch.load(checkpoint_file, map_location=device)
#epoch_ckpt = checkpoint['epoch'] + 1
#tot_time_ckpt = checkpoint['tot_time']
#plot_performance_train = checkpoint['plot_performance_train']
#plot_performance_baseline = checkpoint['plot_performance_baseline']
#Critic.load_state_dict(checkpoint['model_baseline'])
#Actor.load_state_dict(checkpoint['model_train'])
#optimizer.load_state_dict(checkpoint['optimizer'])
#print('Re-start training with saved checkpoint file={:s}\n Checkpoint at epoch= {:d} and time={:.3f}min\n'.format(checkpoint_file,epoch_ckpt-1,tot_time_ckpt/60))
#del checkpoint
#*********************************************# Uncomment these lines to re-start training with saved checkpoint #********************************************#
###################
# Main training loop
###################
start_training_time = time.time()
time_stamp = datetime.datetime.now().strftime("%y-%m-%d--%H-%M-%S")
C = 0 # baseline
R = 0 # reward
zero_to_bsz = torch.arange(B, device=device) # [0,1,...,bsz-1]
for epoch in range(0,n_epoch):
# re-start training with saved checkpoint
epoch += epoch_ckpt
###################
# Train model for one epoch
###################
start = time.time()
Actor.train()
for i in range(1,steps+1):
X = torch.rand(B, size, 2).cuda()
mask = torch.zeros(B,size).cuda()
R = 0
logprobs = 0
reward = 0
Y = X.view(B,size,2)
x = Y[:,0,:]
h = None
c = None
Transcontext = None
#Actor Sampling phase
for k in range(size):
Transcontext,output, h, c, _ = Actor(Transcontext,x=x, X_all=X, h=h, c=c, mask=mask)
sampler = torch.distributions.Categorical(output)
idx = sampler.sample()
Y1 = Y[zero_to_bsz, idx.data].clone()
if k == 0:
Y_ini = Y1.clone()
if k > 0:
reward = torch.sum((Y1 - Y0)**2 , dim=1 )**0.5
Y0 = Y1.clone() # --> insert current node into prev node for the next iteration
x = Y[zero_to_bsz, idx.data].clone()
R += reward
logprobs += torch.log(output[zero_to_bsz, idx.data] + TINY)
mask[zero_to_bsz, idx.data] += -np.inf
R += torch.sum((Y1 - Y_ini)**2 , dim=1 )**0.5
# Critic Baseline phase
mask = torch.zeros(B,size).cuda()
C = 0
baseline = 0
Y = X.view(B,size,2)
x = Y[:,0,:]
h = None
c = None
Transcontext = None
# compute tours for baseline without grad
with torch.no_grad():
for k in range(size):
Transcontext,output, h, c, _ = Critic(Transcontext,x=x, X_all=X, h=h, c=c, mask=mask)
idx = torch.argmax(output, dim=1) # ----> greedy baseline critic
Y1 = Y[zero_to_bsz, idx.data].clone()
if k == 0:
Y_ini = Y1.clone()
if k > 0:
baseline = torch.sum((Y1 - Y0)**2 , dim=1 )**0.5
Y0 = Y1.clone()
x = Y[zero_to_bsz, idx.data].clone()
C += baseline
mask[zero_to_bsz, idx.data] += -np.inf
C += torch.sum((Y1 - Y_ini)**2 , dim=1 )**0.5
###################
# Loss and backprop handling
###################
loss = torch.mean((R - C) * logprobs)
optimizer.zero_grad()
loss.backward()
torch.nn.utils.clip_grad_norm_(Actor.parameters(),1.0, norm_type=2)
optimizer.step()
opt_scheduler.step()
if i % 50 == 0:
print("epoch:{}, batch:{}/{}, reward:{}".format(epoch, i, steps, R.mean().item()))
# R_mean.append(R.mean().item())
# R_std.append(R.std().item())
# greedy validation
tour_len = 0
X_val = np.random.rand(B_val, size_val, 2)
X = X_val
X = torch.Tensor(X).cuda()
mask = torch.zeros(B_val,size_val).cuda()
R = 0
logprobs = 0
Idx = []
reward = 0
Y = X.view(B_val, size_val, 2) # to the same batch size
x = Y[:,0,:]
h = None
c = None
Transcontext = None
for k in range(size_val):
Transcontext,output, h, c, _ = Actor(Transcontext,x=x, X_all=X, h=h, c=c, mask=mask)
sampler = torch.distributions.Categorical(output)
# idx = sampler.sample()
idx = torch.argmax(output, dim=1)
Idx.append(idx.data)
Y1 = Y[[i for i in range(B_val)], idx.data]
if k == 0:
Y_ini = Y1.clone()
if k > 0:
reward = torch.norm(Y1-Y0, dim=1)
Y0 = Y1.clone()
x = Y[[i for i in range(B_val)], idx.data]
R += reward
mask[[i for i in range(B_val)], idx.data] += -np.inf
R += torch.norm(Y1-Y_ini, dim=1)
tour_len += R.mean().item()
print('validation tour length:', tour_len)
#print('validation tour length:', R.std().item())
time_one_epoch = time.time() - start
time_tot = time.time() - start_training_time + tot_time_ckpt
###################
# Evaluate train model and baseline on 1k random TSP instances
###################
Actor.eval()
mean_tour_length_actor = 0
mean_tour_length_critic = 0
for step in range(0,B_valLoop):
# compute tour for model and baseline
X = np.random.rand(B, size, 2)
X = torch.Tensor(X).cuda()
mask = torch.zeros(B,size).cuda()
R = 0
reward = 0
Y = X.view(B,size,2)
x = Y[:,0,:]
h = None
c = None
Transcontext = None
with torch.no_grad():
for k in range(size):
Transcontext,output, h, c, _ = Actor(Transcontext,x=x, X_all=X, h=h, c=c, mask=mask)
idx = torch.argmax(output, dim=1)
Y1 = Y[zero_to_bsz, idx.data].clone()
if k == 0:
Y_ini = Y1.clone()
if k > 0:
#reward = torch.linalg.norm(Y1 - Y0, dim=1) # --> Calculation of the distance between two node
reward = torch.sum((Y1 - Y0)**2 , dim=1 )**0.5
Y0 = Y1.clone() # --> insert current node into prev node for the next iteration
x = Y[zero_to_bsz, idx.data].clone()
R += reward
mask[zero_to_bsz, idx.data] += -np.inf
#R += torch.linalg.norm(Y1 - Y_ini, dim=1)
R += torch.sum((Y1 - Y_ini)**2 , dim=1 )**0.5
# critic baseline
mask = torch.zeros(B,size).cuda()
C = 0
baseline = 0
Y = X.view(B,size,2)
x = Y[:,0,:]
h = None
c = None
Transcontext = None
with torch.no_grad():
for k in range(size):
Transcontext,output, h, c, _ = Critic(Transcontext,x=x, X_all=X, h=h, c=c, mask=mask)
idx = torch.argmax(output, dim=1)
Y1 = Y[zero_to_bsz, idx.data].clone()
if k == 0:
Y_ini = Y1.clone()
if k > 0:
#baseline = torch.linalg.norm(Y1-Y0, dim=1)
baseline = torch.sum((Y1 - Y0)**2 , dim=1 )**0.5
Y0 = Y1.clone()
x = Y[zero_to_bsz, idx.data].clone()
C += baseline
mask[zero_to_bsz, idx.data] += -np.inf
#C += torch.linalg.norm(Y1-Y_ini, dim=1) # ---> Last point to intial point
C += torch.sum((Y1 - Y_ini)**2 , dim=1 )**0.5
mean_tour_length_actor += R.mean().item()
mean_tour_length_critic += C.mean().item()
mean_tour_length_actor = mean_tour_length_actor / B_valLoop
mean_tour_length_critic = mean_tour_length_critic / B_valLoop
# evaluate train model and baseline and update if train model is better
update_baseline = mean_tour_length_actor + TOL < mean_tour_length_critic
print('Avg Actor {} --- Avg Critic {}'.format(mean_tour_length_actor,mean_tour_length_critic))
if update_baseline:
Critic.load_state_dict(Actor.state_dict())
print('My actor is going on the right road Hallelujah :) Updated')
# For checkpoint
plot_performance_train.append([(epoch+1), mean_tour_length_actor])
plot_performance_baseline.append([(epoch+1), mean_tour_length_critic])
# Compute optimality gap
if size==50: gap_train = mean_tour_length_actor/5.692- 1.0
elif size==100: gap_train = mean_tour_length_actor/7.765- 1.0
else: gap_train = -1.0
# Print and save in txt file
mystring_min = 'Epoch: {:d}, epoch time: {:.3f}min, tot time: {:.3f}day, L_actor: {:.3f}, L_critic: {:.3f}, gap_train(%): {:.3f}, update: {}'.format(
epoch, time_one_epoch/60, time_tot/86400, mean_tour_length_actor, mean_tour_length_critic, 100 * gap_train, update_baseline)
print(mystring_min)
print('Save Checkpoints')
# Saving checkpoint
checkpoint_dir = os.path.join("checkpoint")
if not os.path.exists(checkpoint_dir):
os.makedirs(checkpoint_dir)
torch.save({
'epoch': epoch,
'time': time_one_epoch,
'tot_time': time_tot,
'loss': loss.item(),
'plot_performance_train': plot_performance_train,
'plot_performance_baseline': plot_performance_baseline,
'mean_tour_length_val': tour_len,
'model_baseline': Critic.state_dict(),
'model_train': Actor.state_dict(),
'optimizer': optimizer.state_dict(),
}, '{}.pkl'.format(checkpoint_dir + "/checkpoint_" + time_stamp + "-n{}".format(50) + "-gpu{}".format(gpu_id)))
```
| github_jupyter |
<a href="https://colab.research.google.com/github/Serbeld/RX-COVID-19/blob/master/Resize.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
import matplotlib.pyplot as plt
import numpy as np
import cv2
from google.colab import drive
drive.mount('/content/drive')
methods = ['Original','INTER_NEAREST','Original','INTER_LINEAR','Original', 'INTER_AREA','Original','INTER_CUBIC']
grid = cv2.imread("/content/drive/My Drive/2.jpeg")
gridshape = np.array(grid)
#print(gridshape.shape)
fig, axes = plt.subplots(2, 4, figsize=(16, 8),
subplot_kw={'xticks': [], 'yticks': []})
#1k to 500x500px
num_size_max = int((500/1080)*(gridshape.shape[0]))
print(num_size_max)
size = (num_size_max,num_size_max)
for ax, interp_method in zip(axes.flat, methods):
if interp_method == 'Original':
grid2 = grid
if interp_method == 'INTER_NEAREST':
grid2 = cv2.resize(grid, size,interpolation = cv2.INTER_NEAREST)
if interp_method == 'INTER_LINEAR':
grid2 = cv2.resize(grid, size,interpolation = cv2.INTER_LINEAR)
if interp_method == 'INTER_AREA':
grid2 = cv2.resize(grid, size,interpolation = cv2.INTER_AREA)
if interp_method == 'INTER_CUBIC':
grid2 = cv2.resize(grid, size,interpolation = cv2.INTER_CUBIC)
grid2 = np.array(grid2)
ax.imshow(grid2)
ax.set_title(interp_method)
plt.show()
methods = ['Original','INTER_NEAREST','Original','INTER_LINEAR','Original', 'INTER_AREA','Original','INTER_CUBIC']
fig, axes = plt.subplots(2, 4, figsize=(16, 8),
subplot_kw={'xticks': [], 'yticks': []})
#1k to 500x500px
num_size_max = int((500/1080)*(gridshape.shape[0]))
print(num_size_max)
size = (num_size_max,num_size_max)
aporcent = int(num_size_max*0.4)
bporcent = int(num_size_max*1)
cporcent = int(num_size_max*0.1)
dporcent = int(num_size_max*0.7)
for ax, interp_method in zip(axes.flat, methods):
if interp_method == 'Original':
grid2 = grid[int((aporcent/num_size_max)*426):int((bporcent/num_size_max)*426),
int((cporcent/num_size_max)*426):int((dporcent/num_size_max)*426)]
if interp_method == 'INTER_NEAREST':
grid2 = cv2.resize(grid, size,interpolation = cv2.INTER_NEAREST)
grid2 = grid2[aporcent:bporcent,cporcent:dporcent]
if interp_method == 'INTER_LINEAR':
grid2 = cv2.resize(grid, size,interpolation = cv2.INTER_LINEAR)
grid2 = grid2[aporcent:bporcent,cporcent:dporcent]
if interp_method == 'INTER_AREA':
grid2 = cv2.resize(grid, size,interpolation = cv2.INTER_AREA)
grid2 = grid2[aporcent:bporcent,cporcent:dporcent]
if interp_method == 'INTER_CUBIC':
grid2 = cv2.resize(grid, size,interpolation = cv2.INTER_CUBIC)
grid2 = grid2[aporcent:bporcent,cporcent:dporcent]
grid2 = np.array(grid2)
ax.imshow(grid2)
ax.set_title(interp_method)
plt.show()
methods = ['Original','INTER_NEAREST','Original','INTER_LINEAR','Original', 'INTER_AREA','Original','INTER_CUBIC']
fig, axes = plt.subplots(2, 4, figsize=(16, 8),
subplot_kw={'xticks': [], 'yticks': []})
#1k to 500x500px
num_size_max = int((500/1080)*(gridshape.shape[0]))
print(num_size_max)
size = (num_size_max,num_size_max)
aporcent = int(num_size_max*0.7)
bporcent = int(num_size_max*0.9)
cporcent = int(num_size_max*0.3)
dporcent = int(num_size_max*0.5)
for ax, interp_method in zip(axes.flat, methods):
if interp_method == 'Original':
grid2 = grid[int((aporcent/num_size_max)*426):int((bporcent/num_size_max)*426),
int((cporcent/num_size_max)*426):int((dporcent/num_size_max)*426)]
if interp_method == 'INTER_NEAREST':
grid2 = cv2.resize(grid, size,interpolation = cv2.INTER_NEAREST)
grid2 = grid2[aporcent:bporcent,cporcent:dporcent]
if interp_method == 'INTER_LINEAR':
grid2 = cv2.resize(grid, size,interpolation = cv2.INTER_LINEAR)
grid2 = grid2[aporcent:bporcent,cporcent:dporcent]
if interp_method == 'INTER_AREA':
grid2 = cv2.resize(grid, size,interpolation = cv2.INTER_AREA)
grid2 = grid2[aporcent:bporcent,cporcent:dporcent]
if interp_method == 'INTER_CUBIC':
grid2 = cv2.resize(grid, size,interpolation = cv2.INTER_CUBIC)
grid2 = grid2[aporcent:bporcent,cporcent:dporcent]
grid2 = np.array(grid2)
ax.imshow(grid2)
ax.set_title(interp_method)
plt.show()
methods = ['Original','INTER_NEAREST','Original','INTER_LINEAR','Original', 'INTER_AREA','Original','INTER_CUBIC']
fig, axes = plt.subplots(2, 4, figsize=(16, 8),
subplot_kw={'xticks': [], 'yticks': []})
grid = cv2.imread("/content/drive/My Drive/2.jpeg")
gridshape = np.array(grid)
#print(gridshape.shape)
#2k to 500x500px
num_size_max = int((500/2048)*(gridshape.shape[0]))
print(num_size_max)
size = (num_size_max,num_size_max)
for ax, interp_method in zip(axes.flat, methods):
if interp_method == 'Original':
grid2 = grid
if interp_method == 'INTER_NEAREST':
grid2 = cv2.resize(grid, size,interpolation = cv2.INTER_NEAREST)
if interp_method == 'INTER_LINEAR':
grid2 = cv2.resize(grid, size,interpolation = cv2.INTER_LINEAR)
if interp_method == 'INTER_AREA':
grid2 = cv2.resize(grid, size,interpolation = cv2.INTER_AREA)
if interp_method == 'INTER_CUBIC':
grid2 = cv2.resize(grid, size,interpolation = cv2.INTER_CUBIC)
grid2 = np.array(grid2)
ax.imshow(grid2)
ax.set_title(interp_method)
plt.show()
methods = ['Original','INTER_NEAREST','Original','INTER_LINEAR','Original', 'INTER_AREA','Original','INTER_CUBIC']
fig, axes = plt.subplots(2, 4, figsize=(16, 8),
subplot_kw={'xticks': [], 'yticks': []})
#2k to 500x500px
num_size_max = int((500/2048)*(gridshape.shape[0]))
print(num_size_max)
size = (num_size_max,num_size_max)
aporcent = int(num_size_max*0.6)
bporcent = int(num_size_max*1)
cporcent = int(num_size_max*0.2)
dporcent = int(num_size_max*0.6)
for ax, interp_method in zip(axes.flat, methods):
if interp_method == 'Original':
grid2 = grid[int((aporcent/num_size_max)*426):int((bporcent/num_size_max)*426),
int((cporcent/num_size_max)*426):int((dporcent/num_size_max)*426)]
if interp_method == 'INTER_NEAREST':
grid2 = cv2.resize(grid, size,interpolation = cv2.INTER_NEAREST)
grid2 = grid2[aporcent:bporcent,cporcent:dporcent]
if interp_method == 'INTER_LINEAR':
grid2 = cv2.resize(grid, size,interpolation = cv2.INTER_LINEAR)
grid2 = grid2[aporcent:bporcent,cporcent:dporcent]
if interp_method == 'INTER_AREA':
grid2 = cv2.resize(grid, size,interpolation = cv2.INTER_AREA)
grid2 = grid2[aporcent:bporcent,cporcent:dporcent]
if interp_method == 'INTER_CUBIC':
grid2 = cv2.resize(grid, size,interpolation = cv2.INTER_CUBIC)
grid2 = grid2[aporcent:bporcent,cporcent:dporcent]
grid2 = np.array(grid2)
ax.imshow(grid2)
ax.set_title(interp_method)
plt.show()
methods = ['Original','INTER_NEAREST','Original','INTER_LINEAR','Original', 'INTER_AREA','Original','INTER_CUBIC']
fig, axes = plt.subplots(2, 4, figsize=(16, 8),
subplot_kw={'xticks': [], 'yticks': []})
#2k to 500x500px
num_size_max = int((500/2048)*(gridshape.shape[0]))
print(num_size_max)
size = (num_size_max,num_size_max)
aporcent = int(num_size_max*0.75)
bporcent = int(num_size_max*0.9)
cporcent = int(num_size_max*0.35)
dporcent = int(num_size_max*0.5)
for ax, interp_method in zip(axes.flat, methods):
if interp_method == 'Original':
grid2 = grid[int((aporcent/num_size_max)*426):int((bporcent/num_size_max)*426),
int((cporcent/num_size_max)*426):int((dporcent/num_size_max)*426)]
if interp_method == 'INTER_NEAREST':
grid2 = cv2.resize(grid, size,interpolation = cv2.INTER_NEAREST)
grid2 = grid2[aporcent:bporcent,cporcent:dporcent]
if interp_method == 'INTER_LINEAR':
grid2 = cv2.resize(grid, size,interpolation = cv2.INTER_LINEAR)
grid2 = grid2[aporcent:bporcent,cporcent:dporcent]
if interp_method == 'INTER_AREA':
grid2 = cv2.resize(grid, size,interpolation = cv2.INTER_AREA)
grid2 = grid2[aporcent:bporcent,cporcent:dporcent]
if interp_method == 'INTER_CUBIC':
grid2 = cv2.resize(grid, size,interpolation = cv2.INTER_CUBIC)
grid2 = grid2[aporcent:bporcent,cporcent:dporcent]
grid2 = np.array(grid2)
ax.imshow(grid2)
ax.set_title(interp_method)
plt.show()
methods = ['Original','INTER_NEAREST','Original','INTER_LINEAR','Original', 'INTER_AREA','Original','INTER_CUBIC']
fig, axes = plt.subplots(2, 4, figsize=(16, 8),
subplot_kw={'xticks': [], 'yticks': []})
grid = cv2.imread("/content/drive/My Drive/Noise.png")
ret,grid = cv2.threshold(grid,127,255,cv2.THRESH_BINARY)
grid = cv2.resize(grid,(10,10),interpolation = cv2.INTER_NEAREST)
#grid = cv2.cvtColor(grid, cv2.COLOR_BGR2GRAY)
grid = np.array(grid)
gridshape = grid
#print(gridshape.shape)
#1k to 500x500px
num_size_max = int((500/1080)*(gridshape.shape[0]))
print(num_size_max)
size = (num_size_max,num_size_max)
aporcent = int(num_size_max*0)
bporcent = int(num_size_max*1)
cporcent = int(num_size_max*0)
dporcent = int(num_size_max*1)
for ax, interp_method in zip(axes.flat, methods):
if interp_method == 'Original':
grid2 = grid[int((aporcent/num_size_max)*gridshape.shape[0]):int((bporcent/num_size_max)*gridshape.shape[0]),
int((cporcent/num_size_max)*gridshape.shape[0]):int((dporcent/num_size_max)*gridshape.shape[0])]
if interp_method == 'INTER_NEAREST':
grid2 = cv2.resize(grid, size,interpolation = cv2.INTER_NEAREST)
grid2 = grid2[aporcent:bporcent,cporcent:dporcent]
if interp_method == 'INTER_LINEAR':
grid2 = cv2.resize(grid, size,interpolation = cv2.INTER_LINEAR)
grid2 = grid2[aporcent:bporcent,cporcent:dporcent]
if interp_method == 'INTER_AREA':
grid2 = cv2.resize(grid, size,interpolation = cv2.INTER_AREA)
grid2 = grid2[aporcent:bporcent,cporcent:dporcent]
if interp_method == 'INTER_CUBIC':
grid2 = cv2.resize(grid, size,interpolation = cv2.INTER_CUBIC)
grid2 = grid2[aporcent:bporcent,cporcent:dporcent]
grid2 = np.array(grid2)
ax.imshow(grid2)
ax.set_title(interp_method)
plt.show()
methods = ['Original','INTER_NEAREST','Original','INTER_LINEAR','Original', 'INTER_AREA','Original','INTER_CUBIC']
fig, axes = plt.subplots(2, 4, figsize=(16, 8),
subplot_kw={'xticks': [], 'yticks': []})
grid = cv2.imread("/content/drive/My Drive/Noise.png")
ret,grid = cv2.threshold(grid,127,255,cv2.THRESH_BINARY)
grid = cv2.resize(grid,(8,8),interpolation = cv2.INTER_NEAREST)
grid[0:3,0:1] = 0
grid[0:1,0:3] = 0
grid[3:6,3:6] = 255
grid[7:8,6:8] = 0
grid[6:8,7:8] = 0
#grid = cv2.cvtColor(grid, cv2.COLOR_BGR2GRAY)
grid = np.array(grid)
gridshape = grid
#print(gridshape.shape)
#1k to 500x500px
num_size_max = int((500/1080)*(gridshape.shape[0]))
print(num_size_max)
size = (num_size_max,num_size_max)
aporcent = int(num_size_max*0)
bporcent = int(num_size_max*1)
cporcent = int(num_size_max*0)
dporcent = int(num_size_max*1)
for ax, interp_method in zip(axes.flat, methods):
if interp_method == 'Original':
grid2 = grid[int((aporcent/num_size_max)*gridshape.shape[0]):int((bporcent/num_size_max)*gridshape.shape[0]),
int((cporcent/num_size_max)*gridshape.shape[0]):int((dporcent/num_size_max)*gridshape.shape[0])]
if interp_method == 'INTER_NEAREST':
grid2 = cv2.resize(grid, size,interpolation = cv2.INTER_NEAREST)
grid2 = grid2[aporcent:bporcent,cporcent:dporcent]
if interp_method == 'INTER_LINEAR':
grid2 = cv2.resize(grid, size,interpolation = cv2.INTER_LINEAR)
grid2 = grid2[aporcent:bporcent,cporcent:dporcent]
if interp_method == 'INTER_AREA':
grid2 = cv2.resize(grid, size,interpolation = cv2.INTER_AREA)
grid2 = grid2[aporcent:bporcent,cporcent:dporcent]
if interp_method == 'INTER_CUBIC':
grid2 = cv2.resize(grid, size,interpolation = cv2.INTER_CUBIC)
grid2 = grid2[aporcent:bporcent,cporcent:dporcent]
grid2 = np.array(grid2)
ax.imshow(grid2)
ax.set_title(interp_method)
plt.show()
```
| github_jupyter |
```
%matplotlib inline
import pandas as pd
compras_df = pd.read_csv('data/compras_df_enriched.csv')
```
# Probar un modelo supervisado
```
data = compras_df.copy()
for i, column in enumerate(list([str(d) for d in data.dtypes])):
if column == "object":
data[data.columns[i]] = data[data.columns[i]].fillna(data[data.columns[i]].mode())
data[data.columns[i]] = data[data.columns[i]].astype("category").cat.codes
else:
data[data.columns[i]] = data[data.columns[i]].fillna(data[data.columns[i]].median())
data.head(5)
from sklearn.ensemble import RandomForestRegressor
from sklearn.model_selection import train_test_split
y = data.pop('IMPORTE')
X = data.copy()
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.33, random_state=42)
clf = RandomForestRegressor(n_estimators=1000, max_depth = 4)
clf.fit(X_train, y_train)
from sklearn.metrics import r2_score
r2_score(y_test, clf.predict(X_test))
```
# Probar un modelo no supervisado o de clustering para texto
```
import gensim
from nltk.corpus import stopwords
import numpy as np
np.random.seed(2021)
def preprocess(text):
result = []
for token in text.split():
if token not in stopwords.words('spanish') and len(token)>4:
result.append(token)
return result
process_business = [preprocess(x) for x in list(compras_df['BENEFICIARIO'].unique())]
dictionary = gensim.corpora.Dictionary(process_business)
dictionary.filter_extremes(no_below=1, no_above=0.5)
bow_corpus = [dictionary.doc2bow(doc) for doc in process_business]
lda_model = gensim.models.LdaMulticore(bow_corpus, num_topics=6, id2word=dictionary, passes=10, workers=2)
for idx, type_business in lda_model.print_topics(-1):
print('Topic:{} Words:{}'.format(idx, type_business))
def get_max_topic(index):
results = lda_model[bow_corpus[index]]
return max(lda_model[results], key=lambda x: x[1])[0]
business_cluster = {x: get_max_topic(i) for i, x in enumerate(list(compras_df['BENEFICIARIO'].unique()))}
compras_df['CLUSTER_NAME'] = compras_df['BENEFICIARIO'].apply(lambda x: business_cluster[x])
compras_df.groupby('CLUSTER_NAME')['IMPORTE'].mean().plot.bar()
```
# Probar un modelo no supervisado de anomalías en cuanto a importes
```
import numpy as np
from scipy.stats import norm
params = norm.fit(compras_df['IMPORTE'])
params
compras_df['PROB_TOTAL'] = [norm(params[0], params[1]).cdf(x) for x in compras_df['IMPORTE']]
compras_df[compras_df['PROB_TOTAL']>0.95]
models_sheet={}
for sheet in list(compras_df['SHEET'].unique()):
sheet_compras = compras_df[compras_df['SHEET'] == sheet]
params = norm.fit(sheet_compras['IMPORTE'])
models_sheet[sheet] = norm(params[0], params[1])
sheet_prob = []
for i, row in compras_df.iterrows():
sheet_prob.append(models_sheet[row['SHEET']].cdf(row['IMPORTE']))
compras_df['PROB_SHEET'] = sheet_prob
compras_df[compras_df['PROB_SHEET']>0.95]
```
# Formulando conclusiones
* Plantear la tendencia central
* Hay historias en los outeliers
* Esfuerzate en contestar la pregunta original
* Si tu enfoque cambio, documentalo.
| github_jupyter |
# Regression Week 5: LASSO (coordinate descent)
In this notebook, you will implement your very own LASSO solver via coordinate descent. You will:
* Write a function to normalize features
* Implement coordinate descent for LASSO
* Explore effects of L1 penalty
# Fire up graphlab create
Make sure you have the latest version of graphlab (>= 1.7)
```
import graphlab
graphlab.product_key.set_product_key("C0C2-04B4-D94B-70F6-8771-86F9-C6E1-E122")
```
# Load in house sales data
Dataset is from house sales in King County, the region where the city of Seattle, WA is located.
```
sales = graphlab.SFrame('kc_house_data.gl/kc_house_data.gl')
# In the dataset, 'floors' was defined with type string,
# so we'll convert them to int, before using it below
sales['floors'] = sales['floors'].astype(int)
```
If we want to do any "feature engineering" like creating new features or adjusting existing ones we should do this directly using the SFrames as seen in the first notebook of Week 2. For this notebook, however, we will work with the existing features.
# Import useful functions from previous notebook
As in Week 2, we convert the SFrame into a 2D Numpy array. Copy and paste `get_num_data()` from the second notebook of Week 2.
```
import numpy as np # note this allows us to refer to numpy as np instead
def get_numpy_data(data_sframe, features, output):
data_sframe['constant'] = 1 # this is how you add a constant column to an SFrame
# add the column 'constant' to the front of the features list so that we can extract it along with the others:
features = ['constant'] + features # this is how you combine two lists
# select the columns of data_SFrame given by the features list into the SFrame features_sframe (now including constant):
features_sframe = data_sframe[features]
# the following line will convert the features_SFrame into a numpy matrix:
feature_matrix = features_sframe.to_numpy()
# assign the column of data_sframe associated with the output to the SArray output_sarray
output_sarray = data_sframe['price']
# the following will convert the SArray into a numpy array by first converting it to a list
output_array = output_sarray.to_numpy()
return(feature_matrix, output_array)
```
Also, copy and paste the `predict_output()` function to compute the predictions for an entire matrix of features given the matrix and the weights:
```
def predict_output(feature_matrix, weights):
# assume feature_matrix is a numpy matrix containing the features as columns and weights is a corresponding numpy array
# create the predictions vector by using np.dot()
predictions = np.dot(feature_matrix, weights)
return(predictions)
```
# Normalize features
In the house dataset, features vary wildly in their relative magnitude: `sqft_living` is very large overall compared to `bedrooms`, for instance. As a result, weight for `sqft_living` would be much smaller than weight for `bedrooms`. This is problematic because "small" weights are dropped first as `l1_penalty` goes up.
To give equal considerations for all features, we need to **normalize features** as discussed in the lectures: we divide each feature by its 2-norm so that the transformed feature has norm 1.
Let's see how we can do this normalization easily with Numpy: let us first consider a small matrix.
```
X = np.array([[3.,5.,8.],[4.,12.,15.]])
print X
```
Numpy provides a shorthand for computing 2-norms of each column:
```
norms = np.linalg.norm(X, axis=0) # gives [norm(X[:,0]), norm(X[:,1]), norm(X[:,2])]
print norms
```
To normalize, apply element-wise division:
```
print X / norms # gives [X[:,0]/norm(X[:,0]), X[:,1]/norm(X[:,1]), X[:,2]/norm(X[:,2])]
```
Using the shorthand we just covered, write a short function called `normalize_features(feature_matrix)`, which normalizes columns of a given feature matrix. The function should return a pair `(normalized_features, norms)`, where the second item contains the norms of original features. As discussed in the lectures, we will use these norms to normalize the test data in the same way as we normalized the training data.
```
import numpy as np
def normalize_features(feature_matrix):
norms = np.linalg.norm(feature_matrix, axis=0)
features = feature_matrix / norms
return features, norms
```
To test the function, run the following:
```
features, norms = normalize_features(np.array([[3.,6.,9.],[4.,8.,12.]]))
print features
# should print
# [[ 0.6 0.6 0.6]
# [ 0.8 0.8 0.8]]
print norms
# should print
# [5. 10. 15.]
```
# Implementing Coordinate Descent with normalized features
We seek to obtain a sparse set of weights by minimizing the LASSO cost function
```
SUM[ (prediction - output)^2 ] + lambda*( |w[1]| + ... + |w[k]|).
```
(By convention, we do not include `w[0]` in the L1 penalty term. We never want to push the intercept to zero.)
The absolute value sign makes the cost function non-differentiable, so simple gradient descent is not viable (you would need to implement a method called subgradient descent). Instead, we will use **coordinate descent**: at each iteration, we will fix all weights but weight `i` and find the value of weight `i` that minimizes the objective. That is, we look for
```
argmin_{w[i]} [ SUM[ (prediction - output)^2 ] + lambda*( |w[1]| + ... + |w[k]|) ]
```
where all weights other than `w[i]` are held to be constant. We will optimize one `w[i]` at a time, circling through the weights multiple times.
1. Pick a coordinate `i`
2. Compute `w[i]` that minimizes the cost function `SUM[ (prediction - output)^2 ] + lambda*( |w[1]| + ... + |w[k]|)`
3. Repeat Steps 1 and 2 for all coordinates, multiple times
For this notebook, we use **cyclical coordinate descent with normalized features**, where we cycle through coordinates 0 to (d-1) in order, and assume the features were normalized as discussed above. The formula for optimizing each coordinate is as follows:
```
┌ (ro[i] + lambda/2) if ro[i] < -lambda/2
w[i] = ├ 0 if -lambda/2 <= ro[i] <= lambda/2
└ (ro[i] - lambda/2) if ro[i] > lambda/2
```
where
```
ro[i] = SUM[ [feature_i]*(output - prediction + w[i]*[feature_i]) ].
```
Note that we do not regularize the weight of the constant feature (intercept) `w[0]`, so, for this weight, the update is simply:
```
w[0] = ro[i]
```
## Effect of L1 penalty
Let us consider a simple model with 2 features:
```
simple_features = ['sqft_living', 'bedrooms']
my_output = 'price'
(simple_feature_matrix, output) = get_numpy_data(sales, simple_features, my_output)
```
Don't forget to normalize features:
```
simple_feature_matrix, norms = normalize_features(simple_feature_matrix)
```
We assign some random set of initial weights and inspect the values of `ro[i]`:
```
weights = np.array([1., 4., 1.])
```
Use `predict_output()` to make predictions on this data.
```
prediction = predict_output(simple_feature_matrix, weights)
```
Compute the values of `ro[i]` for each feature in this simple model, using the formula given above, using the formula:
```
ro[i] = SUM[ [feature_i]*(output - prediction + w[i]*[feature_i]) ]
```
*Hint: You can get a Numpy vector for feature_i using:*
```
simple_feature_matrix[:,i]
```
```
ro = [0 for i in range((simple_feature_matrix.shape)[1])]
for j in range((simple_feature_matrix.shape)[1]):
ro[j] = (simple_feature_matrix[:,j] * (output - prediction + (weights[j] * simple_feature_matrix[:,j]))).sum()
print ro
```
***QUIZ QUESTION***
Recall that, whenever `ro[i]` falls between `-l1_penalty/2` and `l1_penalty/2`, the corresponding weight `w[i]` is sent to zero. Now suppose we were to take one step of coordinate descent on either feature 1 or feature 2. What range of values of `l1_penalty` **would not** set `w[1]` zero, but **would** set `w[2]` to zero, if we were to take a step in that coordinate?
```
diff = abs((ro[1]*2) - (ro[2]*2))
print('λ = (%e, %e)' %((ro[2]-diff/2+1)*2, (ro[2]+diff/2-1)*2))
```
***QUIZ QUESTION***
What range of values of `l1_penalty` would set **both** `w[1]` and `w[2]` to zero, if we were to take a step in that coordinate?
```
print ro[1]*2
print ro[2]*2
```
So we can say that `ro[i]` quantifies the significance of the i-th feature: the larger `ro[i]` is, the more likely it is for the i-th feature to be retained.
## Single Coordinate Descent Step
Using the formula above, implement coordinate descent that minimizes the cost function over a single feature i. Note that the intercept (weight 0) is not regularized. The function should accept feature matrix, output, current weights, l1 penalty, and index of feature to optimize over. The function should return new weight for feature i.
```
def lasso_coordinate_descent_step(i, feature_matrix, output, weights, l1_penalty):
# compute prediction
prediction = predict_output(feature_matrix, weights)
# compute ro[i] = SUM[ [feature_i]*(output - prediction + weight[i]*[feature_i]) ]
ro_i = np.sum(feature_matrix[:,i]*(output - prediction + weights[i]*feature_matrix[:,i]))
if i == 0: # intercept -- do not regularize
new_weight_i = ro_i
elif ro_i < -l1_penalty/2.:
new_weight_i = ro_i + (l1_penalty/2)
elif ro_i > l1_penalty/2.:
new_weight_i = ro_i - (l1_penalty/2)
else:
new_weight_i = 0.
return new_weight_i
```
To test the function, run the following cell:
```
# should print 0.425558846691
import math
print lasso_coordinate_descent_step(1, np.array([[3./math.sqrt(13),1./math.sqrt(10)],[2./math.sqrt(13),3./math.sqrt(10)]]),
np.array([1., 1.]), np.array([1., 4.]), 0.1)
```
## Cyclical coordinate descent
Now that we have a function that optimizes the cost function over a single coordinate, let us implement cyclical coordinate descent where we optimize coordinates 0, 1, ..., (d-1) in order and repeat.
When do we know to stop? Each time we scan all the coordinates (features) once, we measure the change in weight for each coordinate. If no coordinate changes by more than a specified threshold, we stop.
For each iteration:
1. As you loop over features in order and perform coordinate descent, measure how much each coordinate changes.
2. After the loop, if the maximum change across all coordinates is falls below the tolerance, stop. Otherwise, go back to step 1.
Return weights
**IMPORTANT: when computing a new weight for coordinate i, make sure to incorporate the new weights for coordinates 0, 1, ..., i-1. One good way is to update your weights variable in-place. See following pseudocode for illustration.**
```
for i in range(len(weights)):
old_weights_i = weights[i] # remember old value of weight[i], as it will be overwritten
# the following line uses new values for weight[0], weight[1], ..., weight[i-1]
# and old values for weight[i], ..., weight[d-1]
weights[i] = lasso_coordinate_descent_step(i, feature_matrix, output, weights, l1_penalty)
# use old_weights_i to compute change in coordinate
...
```
```
def lasso_cyclical_coordinate_descent(feature_matrix, output, initial_weights, l1_penalty, tolerance):
weights = initial_weights.copy()
# converged condition variable
converged = False
while not converged:
max_change = 0
for i in range(len(weights)):
old_weights_i = weights[i]
weights[i] = lasso_coordinate_descent_step(i, feature_matrix, output, weights, l1_penalty)
change_i = np.abs(old_weights_i - weights[i])
if change_i > max_change:
max_change = change_i
if max_change < tolerance:
converged = True
return weights
```
Using the following parameters, learn the weights on the sales dataset.
```
simple_features = ['sqft_living', 'bedrooms']
my_output = 'price'
initial_weights = np.zeros(3)
l1_penalty = 1e7
tolerance = 1.0
```
First create a normalized version of the feature matrix, `normalized_simple_feature_matrix`
```
(simple_feature_matrix, output) = get_numpy_data(sales, simple_features, my_output)
(normalized_simple_feature_matrix, simple_norms) = normalize_features(simple_feature_matrix) # normalize features
```
Then, run your implementation of LASSO coordinate descent:
```
weights = lasso_cyclical_coordinate_descent(normalized_simple_feature_matrix, output,
initial_weights, l1_penalty, tolerance)
print weights
# predictions = predict_output(normalized_simple_feature_matrix, weights)
# rss = 0
# for i in range(0, len(predictions)):
# error = predictions[i] - sales['price'][i]
# rss += error * error
# print rss
```
***QUIZ QUESTIONS***
1. What is the RSS of the learned model on the normalized dataset?
2. Which features had weight zero at convergence?
# Evaluating LASSO fit with more features
Let us split the sales dataset into training and test sets.
```
train_data,test_data = sales.random_split(.8,seed=0)
```
Let us consider the following set of features.
```
all_features = ['bedrooms',
'bathrooms',
'sqft_living',
'sqft_lot',
'floors',
'waterfront',
'view',
'condition',
'grade',
'sqft_above',
'sqft_basement',
'yr_built',
'yr_renovated']
```
First, create a normalized feature matrix from the TRAINING data with these features. (Make you store the norms for the normalization, since we'll use them later)
```
(all_feature_matrix, output) = get_numpy_data(train_data, all_features, my_output)
(normalized_all_feature_matrix, simple_norms) = normalize_features(all_feature_matrix) # normalize features
my_output = 'price'
initial_weights = np.zeros(14)
l1_penalty = 1e7
tolerance = 1.0
```
First, learn the weights with `l1_penalty=1e7`, on the training data. Initialize weights to all zeros, and set the `tolerance=1`. Call resulting weights `weights1e7`, you will need them later.
```
weights1e7 = lasso_cyclical_coordinate_descent(normalized_all_feature_matrix, output,
initial_weights, l1_penalty=1e7, tolerance=1)
print weights1e7
```
***QUIZ QUESTION***
What features had non-zero weight in this case?
Next, learn the weights with `l1_penalty=1e8`, on the training data. Initialize weights to all zeros, and set the `tolerance=1`. Call resulting weights `weights1e8`, you will need them later.
```
weights1e8 = lasso_cyclical_coordinate_descent(normalized_all_feature_matrix, output,
initial_weights, l1_penalty=1e8, tolerance=1)
print weights1e8
```
***QUIZ QUESTION***
What features had non-zero weight in this case?
Finally, learn the weights with `l1_penalty=1e4`, on the training data. Initialize weights to all zeros, and set the `tolerance=5e5`. Call resulting weights `weights1e4`, you will need them later. (This case will take quite a bit longer to converge than the others above.)
```
weights1e4 = lasso_cyclical_coordinate_descent(normalized_all_feature_matrix, output,
initial_weights, l1_penalty=1e4, tolerance=5e5)
print weights1e4
```
***QUIZ QUESTION***
What features had non-zero weight in this case?
## Rescaling learned weights
Recall that we normalized our feature matrix, before learning the weights. To use these weights on a test set, we must normalize the test data in the same way.
Alternatively, we can rescale the learned weights to include the normalization, so we never have to worry about normalizing the test data:
In this case, we must scale the resulting weights so that we can make predictions with *original* features:
1. Store the norms of the original features to a vector called `norms`:
```
features, norms = normalize_features(features)
```
2. Run Lasso on the normalized features and obtain a `weights` vector
3. Compute the weights for the original features by performing element-wise division, i.e.
```
weights_normalized = weights / norms
```
Now, we can apply `weights_normalized` to the test data, without normalizing it!
Create a normalized version of each of the weights learned above. (`weights1e4`, `weights1e7`, `weights1e8`).
```
# (normalized_simple_feature_matrix, simple_norms) = normalize_features(all_features) # normalize features
normalized_weights1e7 = weights1e7 / simple_norms
print normalized_weights1e7[3]
normalized_weights1e4 = weights1e4 / simple_norms
normalized_weights1e8 = weights1e8 / simple_norms
```
To check your results, if you call `normalized_weights1e7` the normalized version of `weights1e7`, then:
```
print normalized_weights1e7[3]
```
should return 161.31745624837794.
## Evaluating each of the learned models on the test data
Let's now evaluate the three models on the test data:
```
(test_feature_matrix, test_output) = get_numpy_data(test_data, all_features, 'price')
```
Compute the RSS of each of the three normalized weights on the (unnormalized) `test_feature_matrix`:
```
prediction = predict_output(test_feature_matrix, normalized_weights1e4)
rss = 0
for i in range(0, len(prediction)):
error = prediction[i] - test_data['price'][i]
rss += error * error
print rss
prediction = predict_output(test_feature_matrix, normalized_weights1e7)
rss = 0
for i in range(0, len(prediction)):
error = prediction[i] - test_data['price'][i]
rss += error * error
print rss
prediction = predict_output(test_feature_matrix, normalized_weights1e8)
rss = 0
for i in range(0, len(prediction)):
error = prediction[i] - test_data['price'][i]
rss += error * error
print rss
```
***QUIZ QUESTION***
Which model performed best on the test data?
| github_jupyter |
Copyright (c) Microsoft Corporation. All rights reserved.
Licensed under the MIT License.

# Reinforcement Learning in Azure Machine Learning - Cartpole Problem on Compute Instance
Reinforcement Learning in Azure Machine Learning is a managed service for running reinforcement learning training and simulation. With Reinforcement Learning in Azure Machine Learning, data scientists can start developing reinforcement learning systems on one machine, and scale to compute targets with 100’s of nodes if needed.
This example shows how to use Reinforcement Learning in Azure Machine Learning to train a Cartpole playing agent on a compute instance.
### Cartpole problem
Cartpole, also known as [Inverted Pendulum](https://en.wikipedia.org/wiki/Inverted_pendulum), is a pendulum with a center of mass above its pivot point. This formation is essentially unstable and will easily fall over but can be kept balanced by applying appropriate horizontal forces to the pivot point.
<table style="width:50%">
<tr>
<th>
<img src="./images/cartpole.png" alt="Cartpole image" />
</th>
</tr>
<tr>
<th><p>Fig 1. Cartpole problem schematic description (from <a href="https://towardsdatascience.com/cartpole-introduction-to-reinforcement-learning-ed0eb5b58288">towardsdatascience.com</a>).</p></th>
</tr>
</table>
The goal here is to train an agent to keep the cartpole balanced by applying appropriate forces to the pivot point.
See [this video](https://www.youtube.com/watch?v=XiigTGKZfks) for a real-world demonstration of cartpole problem.
### Prerequisite
The user should have completed the Azure Machine Learning Tutorial: [Get started creating your first ML experiment with the Python SDK](https://docs.microsoft.com/en-us/azure/machine-learning/tutorial-1st-experiment-sdk-setup). You will need to make sure that you have a valid subscription ID, a resource group, and an Azure Machine Learning workspace. All datastores and datasets you use should be associated with your workspace.
## Set up Development Environment
The following subsections show typical steps to setup your development environment. Setup includes:
* Connecting to a workspace to enable communication between your local machine and remote resources
* Creating an experiment to track all your runs
* Using a Compute Instance as compute target
### Azure Machine Learning SDK
Display the Azure Machine Learning SDK version.
```
import azureml.core
print("Azure Machine Learning SDK Version:", azureml.core.VERSION)
```
### Get Azure Machine Learning workspace
Get a reference to an existing Azure Machine Learning workspace.
```
from azureml.core import Workspace
ws = Workspace.from_config()
print(ws.name, ws.location, ws.resource_group, sep = ' | ')
```
### Use Compute Instance as compute target
A compute target is a designated compute resource where you run your training and simulation scripts. This location may be your local machine or a cloud-based compute resource. For more information see [What are compute targets in Azure Machine Learning?](https://docs.microsoft.com/en-us/azure/machine-learning/concept-compute-target)
The code below shows how to use current compute instance as a compute target. First some helper functions:
```
import os.path
# Get information about the currently running compute instance (notebook VM), like its name and prefix.
def load_nbvm():
if not os.path.isfile("/mnt/azmnt/.nbvm"):
return None
with open("/mnt/azmnt/.nbvm", 'r') as nbvm_file:
return {key:value for (key, value) in line.strip().split('=') for line in nbvm_file}
```
Then we use these helper functions to get a handle to current compute instance.
```
from azureml.core.compute import ComputeInstance
from azureml.core.compute_target import ComputeTargetException
# Load current compute instance info
current_compute_instance = load_nbvm()
# For this demo, let's use the current compute instance as the compute target, if available
if current_compute_instance:
print("Current compute instance:", current_compute_instance)
instance_name = current_compute_instance['instance']
else:
instance_name = "cartpole-ci-stdd2v2"
try:
instance = ComputeInstance(workspace=ws, name=instance_name)
print('Found existing instance, use it.')
except ComputeTargetException:
print("Creating new compute instance...")
compute_config = ComputeInstance.provisioning_configuration(
vm_size='STANDARD_D2_V2'
)
instance = ComputeInstance.create(ws, instance_name, compute_config)
instance.wait_for_completion(show_output=True)
print("Instance name:", instance_name)
compute_target = ws.compute_targets[instance_name]
print("Compute target status:")
print(compute_target.get_status().serialize())
```
### Create Azure Machine Learning experiment
Create an experiment to track the runs in your workspace.
```
from azureml.core.experiment import Experiment
experiment_name = 'CartPole-v0-CI'
exp = Experiment(workspace=ws, name=experiment_name)
```
## Train Cartpole Agent
To facilitate reinforcement learning, Azure Machine Learning Python SDK provides a high level abstraction, the _ReinforcementLearningEstimator_ class, which allows users to easily construct reinforcement learning run configurations for the underlying reinforcement learning framework. Reinforcement Learning in Azure Machine Learning supports the open source [Ray framework](https://ray.io/) and its highly customizable [RLlib](https://ray.readthedocs.io/en/latest/rllib.html#rllib-scalable-reinforcement-learning). In this section we show how to use _ReinforcementLearningEstimator_ and Ray/RLlib framework to train a cartpole playing agent.
### Create reinforcement learning estimator
The code below creates an instance of *ReinforcementLearningEstimator*, `training_estimator`, which then will be used to submit a job to Azure Machine Learning to start the Ray experiment run.
Note that this example is purposely simplified to the minimum. Here is a short description of the parameters we are passing into the constructor:
- `source_directory`, local directory containing your training script(s) and helper modules,
- `entry_script`, path to your entry script relative to the source directory,
- `script_params`, constant parameters to be passed to each run of training script,
- `compute_target`, reference to the compute target in which the trainer and worker(s) jobs will be executed,
- `rl_framework`, the reinforcement learning framework to be used (currently must be Ray).
We use the `script_params` parameter to pass in general and algorithm-specific parameters to the training script.
```
from azureml.contrib.train.rl import ReinforcementLearningEstimator, Ray
training_algorithm = "PPO"
rl_environment = "CartPole-v0"
script_params = {
# Training algorithm
"--run": training_algorithm,
# Training environment
"--env": rl_environment,
# Algorithm-specific parameters
"--config": '\'{"num_gpus": 0, "num_workers": 1}\'',
# Stop conditions
"--stop": '\'{"episode_reward_mean": 200, "time_total_s": 300}\'',
# Frequency of taking checkpoints
"--checkpoint-freq": 2,
# If a checkpoint should be taken at the end - optional argument with no value
"--checkpoint-at-end": "",
# Log directory
"--local-dir": './logs'
}
training_estimator = ReinforcementLearningEstimator(
# Location of source files
source_directory='files',
# Python script file
entry_script='cartpole_training.py',
# A dictionary of arguments to pass to the training script specified in ``entry_script``
script_params=script_params,
# The Azure Machine Learning compute target set up for Ray head nodes
compute_target=compute_target,
# Reinforcement learning framework. Currently must be Ray.
rl_framework=Ray()
)
```
### Training script
As recommended in RLlib documentations, we use Ray Tune API to run the training algorithm. All the RLlib built-in trainers are compatible with the Tune API. Here we use `tune.run()` to execute a built-in training algorithm. For convenience, down below you can see part of the entry script where we make this call.
This is the list of parameters we are passing into `tune.run()` via the `script_params` parameter:
- `run_or_experiment`: name of the [built-in algorithm](https://ray.readthedocs.io/en/latest/rllib-algorithms.html#rllib-algorithms), 'PPO' in our example,
- `config`: Algorithm-specific configuration. This includes specifying the environment, `env`, which in our example is the gym **[CartPole-v0](https://gym.openai.com/envs/CartPole-v0/)** environment,
- `stop`: stopping conditions, which could be any of the metrics returned by the trainer. Here we use "mean of episode reward", and "total training time in seconds" as stop conditions, and
- `checkpoint_freq` and `checkpoint_at_end`: Frequency of taking checkpoints (number of training iterations between checkpoints), and if a checkpoint should be taken at the end.
We also specify the `local_dir`, the directory in which the training logs, checkpoints and other training artificats will be recorded.
See [RLlib Training APIs](https://ray.readthedocs.io/en/latest/rllib-training.html#rllib-training-apis) for more details, and also [Training (tune.run, tune.Experiment)](https://ray.readthedocs.io/en/latest/tune/api_docs/execution.html#training-tune-run-tune-experiment) for the complete list of parameters.
```python
import ray
import ray.tune as tune
if __name__ == "__main__":
# parse arguments ...
# Intitialize ray
ay.init(address=args.ray_address)
# Run training task using tune.run
tune.run(
run_or_experiment=args.run,
config=dict(args.config, env=args.env),
stop=args.stop,
checkpoint_freq=args.checkpoint_freq,
checkpoint_at_end=args.checkpoint_at_end,
local_dir=args.local_dir
)
```
### Submit the estimator to start experiment
Now we use the *training_estimator* to submit a run.
```
training_run = exp.submit(training_estimator)
```
### Monitor experiment
Azure Machine Learning provides a Jupyter widget to show the status of an experiment run. You could use this widget to monitor the status of the runs.
Note that _ReinforcementLearningEstimator_ creates at least two runs: (a) A parent run, i.e. the run returned above, and (b) a collection of child runs. The number of the child runs depends on the configuration of the reinforcement learning estimator. In our simple scenario, configured above, only one child run will be created.
The widget will show a list of the child runs as well. You can click on the link under **Status** to see the details of a child run. It will also show the metrics being logged.
```
from azureml.widgets import RunDetails
RunDetails(training_run).show()
```
### Stop the run
To stop the run, call `training_run.cancel()`.
```
# Uncomment line below to cancel the run
# training_run.cancel()
```
### Wait for completion
Wait for the run to complete before proceeding.
**Note: The run may take a few minutes to complete.**
```
training_run.wait_for_completion()
```
### Get a handle to the child run
You can obtain a handle to the child run as follows. In our scenario, there is only one child run, we have it called `child_run_0`.
```
import time
child_run_0 = None
timeout = 30
while timeout > 0 and not child_run_0:
child_runs = list(training_run.get_children())
print('Number of child runs:', len(child_runs))
if len(child_runs) > 0:
child_run_0 = child_runs[0]
break
time.sleep(2) # Wait for 2 seconds
timeout -= 2
print('Child run info:')
print(child_run_0)
```
## Evaluate Trained Agent and See Results
We can evaluate a previously trained policy using the `rollout.py` helper script provided by RLlib (see [Evaluating Trained Policies](https://ray.readthedocs.io/en/latest/rllib-training.html#evaluating-trained-policies) for more details). Here we use an adaptation of this script to reconstruct a policy from a checkpoint taken and saved during training. We took these checkpoints by setting `checkpoint-freq` and `checkpoint-at-end` parameters above.
In this section we show how to get access to these checkpoints data, and then how to use them to evaluate the trained policy.
### Create a dataset of training artifacts
To evaluate a trained policy (a checkpoint) we need to make the checkpoint accessible to the rollout script. All the training artifacts are stored in workspace default datastore under **azureml/<run_id>** directory.
Here we create a file dataset from the stored artifacts, and then use this dataset to feed these data to rollout estimator.
```
from azureml.core import Dataset
run_id = child_run_0.id # Or set to run id of a completed run (e.g. 'rl-cartpole-v0_1587572312_06e04ace_head')
run_artifacts_path = os.path.join('azureml', run_id)
print("Run artifacts path:", run_artifacts_path)
# Create a file dataset object from the files stored on default datastore
datastore = ws.get_default_datastore()
training_artifacts_ds = Dataset.File.from_files(datastore.path(os.path.join(run_artifacts_path, '**')))
```
To verify, we can print out the number (and paths) of all the files in the dataset, as follows.
```
artifacts_paths = training_artifacts_ds.to_path()
print("Number of files in dataset:", len(artifacts_paths))
# Uncomment line below to print all file paths
#print("Artifacts dataset file paths: ", artifacts_paths)
```
### Evaluate a trained policy
We need to configure another reinforcement learning estimator, `rollout_estimator`, and then use it to submit another run. Note that the entry script for this estimator now points to `cartpole-rollout.py` script.
Also note how we pass the checkpoints dataset to this script using `inputs` parameter of the _ReinforcementLearningEstimator_.
We are using script parameters to pass in the same algorithm and the same environment used during training. We also specify the checkpoint number of the checkpoint we wish to evaluate, `checkpoint-number`, and number of the steps we shall run the rollout, `steps`.
The checkpoints dataset will be accessible to the rollout script as a mounted folder. The mounted folder and the checkpoint number, passed in via `checkpoint-number`, will be used to create a path to the checkpoint we are going to evaluate. The created checkpoint path then will be passed into RLlib rollout script for evaluation.
Let's find the checkpoints and the last checkpoint number first.
```
# Find checkpoints and last checkpoint number
checkpoint_files = [
os.path.basename(file) for file in training_artifacts_ds.to_path() \
if os.path.basename(file).startswith('checkpoint-') and \
not os.path.basename(file).endswith('tune_metadata')
]
checkpoint_numbers = []
for file in checkpoint_files:
checkpoint_numbers.append(int(file.split('-')[1]))
print("Checkpoints:", checkpoint_numbers)
last_checkpoint_number = max(checkpoint_numbers)
print("Last checkpoint number:", last_checkpoint_number)
```
Now let's configure rollout estimator. Note that we use the last checkpoint for evaluation. The assumption is that the last checkpoint points to our best trained agent. You may change this to any of the checkpoint numbers printed above and observe the effect.
```
script_params = {
# Checkpoint number of the checkpoint from which to roll out
"--checkpoint-number": last_checkpoint_number,
# Training algorithm
"--run": training_algorithm,
# Training environment
"--env": rl_environment,
# Algorithm-specific parameters
"--config": '{}',
# Number of rollout steps
"--steps": 2000,
# If should repress rendering of the environment
"--no-render": ""
}
rollout_estimator = ReinforcementLearningEstimator(
# Location of source files
source_directory='files',
# Python script file
entry_script='cartpole_rollout.py',
# A dictionary of arguments to pass to the rollout script specified in ``entry_script``
script_params = script_params,
# Data inputs
inputs=[
training_artifacts_ds.as_named_input('artifacts_dataset'),
training_artifacts_ds.as_named_input('artifacts_path').as_mount()],
# The Azure Machine Learning compute target
compute_target=compute_target,
# Reinforcement learning framework. Currently must be Ray.
rl_framework=Ray(),
# Additional pip packages to install
pip_packages = ['azureml-dataprep[fuse,pandas]'])
```
Same as before, we use the *rollout_estimator* to submit a run.
```
rollout_run = exp.submit(rollout_estimator)
```
And then, similar to the training section, we can monitor the real-time progress of the rollout run and its chid as follows. If you browse logs of the child run you can see the evaluation results recorded in driver_log.txt file. Note that you may need to wait several minutes before these results become available.
```
RunDetails(rollout_run).show()
```
Wait for completion of the rollout run, or you may cancel the run.
```
# Uncomment line below to cancel the run
#rollout_run.cancel()
rollout_run.wait_for_completion()
```
## Cleaning up
For your convenience, below you can find code snippets to clean up any resources created as part of this tutorial that you don't wish to retain.
```
# To archive the created experiment:
#exp.archive()
# To delete created compute instance
if not current_compute_instance:
compute_target.delete()
```
## Next
This example was about running Reinforcement Learning in Azure Machine Learning (Ray/RLlib Framework) on a compute instance. Please see [Cartpole Problem on Single Compute](../cartpole-on-single-compute/cartpole_sc.ipynb)
example which uses Ray RLlib to train a Cartpole playing agent on a single node remote compute.
| github_jupyter |
<a href="https://colab.research.google.com/github/NeuromatchAcademy/course-content-dl/blob/main/tutorials/W2D5_GenerativeModels/W2D5_Tutorial3.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Neuromatch Academy: Week 2, Day 5, Tutorial 3
# VAEs and GANs : Conditional GANs and Implications of GAN Technology
__Content creators:__ Kai Xu, Seungwook Han, Akash Srivastava
__Content reviewers:__ Polina Turishcheva, Melvin Selim Atay, Hadi Vafaei, Deepak Raya
__Content editors:__ Spiros Chavlis
__Production editors:__ Arush Tagade, Spiros Chavlis
---
## Tutorial Objectives
The goal of this notebook is to understand conditional GANs. Then you will have the opportunity to experience first-hand how effective GANs are at modeling the data distribution and to question what the consequences of this technology may be.
By the end of this tutorial you will be able to:
- Understand the differences in conditional GANs.
- Generate high-dimensional natural images from a BigGAN.
- Understand the efficacy of GANs in modeling the data distribution (e.g., faces).
- Understand the energy inefficiency / environmental impact of training these large generative models.
- Understand the implications of this technology (ethics, environment, *etc*.).
```
#@markdown Tutorial slides (pt. 1)
# you should link the slides for all tutorial videos here (we will store pdfs on osf)
from IPython.display import HTML
HTML('<iframe src="https://docs.google.com/presentation/d/1eP79mRMzD2Q7Utol3kZ5hooYIXRTQMMb" frameborder="0" width="960" height="569" allowfullscreen="true" mozallowfullscreen="true" webkitallowfullscreen="true"></iframe>')
```
---
# Setup
```
# Imports
# Install Huggingface BigGAN library
!pip install pytorch-pretrained-biggan --quiet
!pip install Pillow libsixel-python --quiet
# Import libraries
import nltk
import torch
import random
import torchvision
import numpy as np
import matplotlib.pyplot as plt
from pytorch_pretrained_biggan import BigGAN
from pytorch_pretrained_biggan import one_hot_from_names
from pytorch_pretrained_biggan import truncated_noise_sample
nltk.download('wordnet')
device = 'cuda' if torch.cuda.is_available() else 'cpu'
#@title Figure settings
from IPython.display import display
import ipywidgets
import ipywidgets as widgets # interactive display
%config InlineBackend.figure_format = 'retina'
plt.style.use("https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/nma.mplstyle")
# @title Set seed for reproducibility in Pytorch
# https://pytorch.org/docs/stable/notes/randomness.html
def set_seed(seed):
"""
Set random seed for reproducibility
Args:
seed: integer
A positive integer to ensure reproducibility
"""
random.seed(seed)
torch.manual_seed(seed)
torch.cuda.manual_seed_all(seed)
torch.cuda.manual_seed(seed)
np.random.seed(seed)
torch.backends.cudnn.deterministic = True
torch.backends.cudnn.benchmark = False
print(f'Seed {seed} has been set.')
def seed_worker(worker_id):
worker_seed = torch.initial_seed() % 2**32
np.random.seed(worker_seed)
random.seed(worker_seed)
set_seed(522)
```
---
# Section 1: Generating with a conditional GAN (BigGAN)
```
#@title Video 1: Conditional Generative Models
# Insert the ID of the corresponding youtube video
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="lV6zH2xDZck", width=854, height=480, fs=1)
print("Video available at https://youtu.be/" + video.id)
video
```
In this section, we will load a pre-trained conditional GAN, BigGAN, which is the state-of-the-art model in conditional high-dimensional natural image generation, and generate samples from it. Since it is a class conditional model, we will be able to use the class label to generate images from the respective classes of objects.
Read here for more details on BigGAN: https://arxiv.org/pdf/1809.11096.pdf
```
# Load respective BigGAN model for the specified resolution (biggan-deep-128, biggan-deep-256, biggan-deep-512)
def load_biggan(model_res):
return BigGAN.from_pretrained('biggan-deep-{}'.format(model_res))
# Create class and noise vectors for sampling from BigGAN
def create_class_noise_vectors(class_str, trunc, num_samples):
class_vector = one_hot_from_names([class_str]*num_samples, batch_size=num_samples)
noise_vector = truncated_noise_sample(truncation=trunc, batch_size=num_samples)
return class_vector, noise_vector
# Generate samples from BigGAN
def generate_biggan_samples(model, class_vector, noise_vector, truncation=0.4,
device=device):
# Convert to tensor
noise_vector = torch.from_numpy(noise_vector)
class_vector = torch.from_numpy(class_vector)
# Move to GPU
noise_vector = noise_vector.to(device)
class_vector = class_vector.to(device)
model.to(device)
# Generate an image
with torch.no_grad():
output = model(noise_vector, class_vector, truncation)
# Back to CPU
output = output.to('cpu')
# The output layer of BigGAN has a tanh layer, resulting the range of [-1, 1] for the output image
# Therefore, we normalize the images properly to [0, 1] range.
# Clipping is only in case of numerical instability problems
output = torch.clip(((output.detach().clone() + 1) / 2.0), 0, 1)
output = output
# Make grid and show generated samples
output_grid = torchvision.utils.make_grid(output, nrow=min(4, output.shape[0]), padding=5)
plt.imshow(output_grid.permute(1,2,0))
return output_grid
def generate(b):
# Create BigGAN model
model = load_biggan(MODEL_RESOLUTION)
# Use specified parameters (resolution, class, number of samples, etc) to generate from BigGAN
class_vector, noise_vector = create_class_noise_vectors(CLASS, TRUNCATION, NUM_SAMPLES)
samples_grid = generate_biggan_samples(model, class_vector, noise_vector, TRUNCATION, device)
torchvision.utils.save_image(samples_grid, 'samples.png')
### If CUDA out of memory issue, lower NUM_SAMPLES (number of samples)
```
## Section 1.1: Define configurations
We will now define the configurations (resolution of model, number of samples, class to sample from, truncation level) under which we will sample from BigGAN.
***Question***: What is the truncation trick employed by BigGAN? How does sample variety and fidelity change by varying the truncation level? (Hint: play with the truncation slider and try sampling at different levels)
```
#@title { run: "auto" }
### RUN THIS BLOCK EVERY TIME YOU CHANGE THE PARAMETERS FOR GENERATION
# Resolution at which to generate
MODEL_RESOLUTION = "256" #@param [128, 256, 512]
# Number of images to generate
NUM_SAMPLES = 1 #@param {type:"slider", min:1, max:100, step:1}
# Class of images to generate
CLASS = 'German shepherd' #@param ['tench', 'magpie', 'jellyfish', 'German shepherd', 'bee', 'acoustic guitar', 'coffee mug', 'minibus', 'monitor']
# Truncation level of the normal distribution we sample z from
TRUNCATION = 0.4 #@param {type:"slider", min:0.1, max:1, step:0.1}
# Create generate button, given parameters specified above
button = widgets.Button(description="GENERATE!",
layout=widgets.Layout(width='30%', height='80px'),
button_style='danger')
output = widgets.Output()
display(button, output)
button.on_click(generate)
```
# Food for thought
1. How does BigGAN differ from previous state-of-the-art generative models for high-dimensional natural images? In other words, how does BigGAN solve high-dimensional image generation? (Hint: look into model architecture and training configurations) (BigGAN paper: https://arxiv.org/pdf/1809.11096.pdf)
2. Continuing from Question 1, what are the drawbacks of introducing such techniques into training large models for high-dimensional, diverse datasets?
3. Play with other pre-trained generative models like StyleGAN here -- where code for sampling and interpolation in the latent space is available: https://github.com/NVlabs/stylegan
---
# Section 2: Ethical issues
```
#@markdown Tutorial slides (pt. 2)
# you should link the slides for all tutorial videos here (we will store pdfs on osf)
from IPython.display import HTML
HTML('<iframe src="https://docs.google.com/presentation/d/1-omVjYriCQumx1q_dtAGw-gWmnx7uPz5" frameborder="0" width="960" height="569" allowfullscreen="true" mozallowfullscreen="true" webkitallowfullscreen="true"></iframe>')
#@title Video 2: Ethical Issues
# Insert the ID of the corresponding youtube video
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="ZtWFeUZgfVk", width=854, height=480, fs=1)
print("Video available at https://youtu.be/" + video.id)
video
```
## Section 2.1: Faces Quiz
Now is your turn to test your abilities on recognizing a real vs. a fake image!
```
#@markdown Real or Fake?
from IPython import display
from IPython.display import IFrame
IFrame(src='https://docs.google.com/forms/d/e/1FAIpQLSeGjn2S2bn6Q1qWjVgDS5LG7G1GsQQh2Q0T9dEUO1z5_W0yYg/viewform?usp=sf_link', width=900, height=600)
```
## Section 2.2: Energy Efficiency Quiz
```
#@markdown Make a guess
IFrame(src='https://docs.google.com/forms/d/e/1FAIpQLSe8suNt4ZmadSr_6IWq6s_nUYxC1VCpjR2cBBmQ7cR_5znCZw/viewform?usp=sf_link', width=900, height=600)
```
---
# Summary
```
#@markdown Tutorial slides (pt. 3)
# you should link the slides for all tutorial videos here (we will store pdfs on osf)
from IPython.display import HTML
HTML('<iframe src="https://docs.google.com/presentation/d/1s8jHcZUudPp0h1yZLHRI9Rk8hz3zuoUS" frameborder="0" width="960" height="569" allowfullscreen="true" mozallowfullscreen="true" webkitallowfullscreen="true"></iframe>')
#@title Video 3: Recap and advanced topics
# Insert the ID of the corresponding youtube video
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="7nUjFG3N04I", width=854, height=480, fs=1)
print("Video available at https://youtu.be/" + video.id)
video
```
Hooray! You have finished the second week of NMA-DL course!!!
In the first section of this tutorial, we have learned:
- How conditional GANs differ from unconditional models
- How to use a pre-trained BigGAN model to generate high-dimensional photo-realistic images and its tricks to modulate diversity and image fidelity
In the second section, we learned about the broader ethical implications of GAN technology on society through deepfakes and their tremendous energy inefficiency.
On the brighter side, as we learned throughout the week, GANs are very effective in modeling the data distribution and have many practical applications.
For example, as personalized healthcare and applications of AI in healthcare rise, the need to remove any Personally Identifiable Information (PII) becomes more important. As shown in this paper (https://link.springer.com/chapter/10.1007/978-3-030-45385-5_36), GANs can be leveraged to anonymize healthcare data.
As a food for thought, what are some other practical applications of GANs that you can think of? Discuss with your pod your ideas.
| github_jupyter |
```
import keras
from keras.models import Sequential, Model, load_model
from keras.layers import Dense, Dropout, Activation, Flatten, Input, Lambda
from keras.layers import Conv2D, MaxPooling2D, AveragePooling2D, Conv1D, MaxPooling1D, LSTM, ConvLSTM2D, GRU, CuDNNLSTM, CuDNNGRU, BatchNormalization, LocallyConnected2D, Permute, TimeDistributed, Bidirectional
from keras.layers import Concatenate, Reshape, Softmax, Conv2DTranspose, Embedding, Multiply
from keras.callbacks import ModelCheckpoint, EarlyStopping, Callback
from keras import regularizers
from keras import backend as K
from keras.utils.generic_utils import Progbar
from keras.layers.merge import _Merge
import keras.losses
from keras.datasets import mnist
from functools import partial
from collections import defaultdict
import tensorflow as tf
from tensorflow.python.framework import ops
import isolearn.keras as iso
import numpy as np
import tensorflow as tf
import logging
logging.getLogger('tensorflow').setLevel(logging.ERROR)
import os
import pickle
import numpy as np
import isolearn.io as isoio
import isolearn.keras as isol
import pandas as pd
import scipy.sparse as sp
import scipy.io as spio
import matplotlib.pyplot as plt
from sequence_logo_helper import dna_letter_at, plot_dna_logo
from keras.backend.tensorflow_backend import set_session
def contain_tf_gpu_mem_usage() :
config = tf.ConfigProto()
config.gpu_options.allow_growth = True
sess = tf.Session(config=config)
set_session(sess)
contain_tf_gpu_mem_usage()
class EpochVariableCallback(Callback) :
def __init__(self, my_variable, my_func) :
self.my_variable = my_variable
self.my_func = my_func
def on_epoch_begin(self, epoch, logs={}) :
K.set_value(self.my_variable, self.my_func(K.get_value(self.my_variable), epoch))
#Define dataset/experiment name
dataset_name = "apa_doubledope"
#Load cached dataframe
cached_dict = pickle.load(open('apa_doubledope_cached_set.pickle', 'rb'))
data_df = cached_dict['data_df']
print("len(data_df) = " + str(len(data_df)) + " (loaded)")
#Make generators
valid_set_size = 0.05
test_set_size = 0.05
batch_size = 32
#Generate training and test set indexes
data_index = np.arange(len(data_df), dtype=np.int)
train_index = data_index[:-int(len(data_df) * (valid_set_size + test_set_size))]
valid_index = data_index[train_index.shape[0]:-int(len(data_df) * test_set_size)]
test_index = data_index[train_index.shape[0] + valid_index.shape[0]:]
print('Training set size = ' + str(train_index.shape[0]))
print('Validation set size = ' + str(valid_index.shape[0]))
print('Test set size = ' + str(test_index.shape[0]))
data_gens = {
gen_id : iso.DataGenerator(
idx,
{'df' : data_df},
batch_size=batch_size,
inputs = [
{
'id' : 'seq',
'source_type' : 'dataframe',
'source' : 'df',
'extractor' : iso.SequenceExtractor('padded_seq', start_pos=180, end_pos=180 + 205),
'encoder' : iso.OneHotEncoder(seq_length=205),
'dim' : (1, 205, 4),
'sparsify' : False
}
],
outputs = [
{
'id' : 'hairpin',
'source_type' : 'dataframe',
'source' : 'df',
'extractor' : lambda row, index: row['proximal_usage'],
'transformer' : lambda t: t,
'dim' : (1,),
'sparsify' : False
}
],
randomizers = [],
shuffle = True if gen_id == 'train' else False
) for gen_id, idx in [('all', data_index), ('train', train_index), ('valid', valid_index), ('test', test_index)]
}
#Load data matrices
x_train = np.concatenate([data_gens['train'][i][0][0] for i in range(len(data_gens['train']))], axis=0)
x_test = np.concatenate([data_gens['test'][i][0][0] for i in range(len(data_gens['test']))], axis=0)
y_train = np.concatenate([data_gens['train'][i][1][0] for i in range(len(data_gens['train']))], axis=0)
y_test = np.concatenate([data_gens['test'][i][1][0] for i in range(len(data_gens['test']))], axis=0)
print("x_train.shape = " + str(x_train.shape))
print("x_test.shape = " + str(x_test.shape))
print("y_train.shape = " + str(y_train.shape))
print("y_test.shape = " + str(y_test.shape))
#Define sequence template (APA Doubledope sublibrary)
#sequence_template = 'CTTCCGATCTNNNNNNNNNNNNNNNNNNNNCATTACTCGCATCCANNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNCAGCCAATTAAGCCNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNCTAC'
sequence_template = 'CTTCCGATCTNNNNNNNNNNNNNNNNNNNNCATTACTCGCATCCANNNNNNNNNNNNNNNNNNNNNNNNNANTAAANNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNCAGCCAATTAAGCCNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNCTAC'
sequence_mask = np.array([1 if sequence_template[j] == 'N' else 0 for j in range(len(sequence_template))])
#Visualize background sequence distribution
save_figs = True
pseudo_count = 1.0
x_mean = (np.sum(x_train, axis=(0, 1)) + pseudo_count) / (x_train.shape[0] + 4. * pseudo_count)
x_mean_logits = np.log(x_mean / (1. - x_mean))
plot_dna_logo(np.copy(x_mean), sequence_template=sequence_template, figsize=(14, 0.65), logo_height=1.0, plot_start=0, plot_end=205, plot_sequence_template=True, save_figs=save_figs, fig_name="benchmark_inclusion_" + dataset_name + "_background")
#Calculate mean training set conservation
entropy = np.sum(x_mean * -np.log(x_mean), axis=-1) / np.log(2.0)
conservation = 2.0 - entropy
x_mean_conservation = np.sum(conservation) / np.sum(sequence_mask)
print("Mean conservation (bits) = " + str(x_mean_conservation))
#Calculate mean training set kl-divergence against background
x_train_clipped = np.clip(np.copy(x_train[:, 0, :, :]), 1e-8, 1. - 1e-8)
kl_divs = np.sum(x_train_clipped * np.log(x_train_clipped / np.tile(np.expand_dims(x_mean, axis=0), (x_train_clipped.shape[0], 1, 1))), axis=-1) / np.log(2.0)
x_mean_kl_divs = np.sum(kl_divs * sequence_mask, axis=-1) / np.sum(sequence_mask)
x_mean_kl_div = np.mean(x_mean_kl_divs)
print("Mean KL Div against background (bits) = " + str(x_mean_kl_div))
from tensorflow.python.framework import ops
#Stochastic Binarized Neuron helper functions (Tensorflow)
#ST Estimator code adopted from https://r2rt.com/beyond-binary-ternary-and-one-hot-neurons.html
#See Github https://github.com/spitis/
def st_sampled_softmax(logits):
with ops.name_scope("STSampledSoftmax") as namescope :
nt_probs = tf.nn.softmax(logits)
onehot_dim = logits.get_shape().as_list()[1]
sampled_onehot = tf.one_hot(tf.squeeze(tf.multinomial(logits, 1), 1), onehot_dim, 1.0, 0.0)
with tf.get_default_graph().gradient_override_map({'Ceil': 'Identity', 'Mul': 'STMul'}):
return tf.ceil(sampled_onehot * nt_probs)
def st_hardmax_softmax(logits):
with ops.name_scope("STHardmaxSoftmax") as namescope :
nt_probs = tf.nn.softmax(logits)
onehot_dim = logits.get_shape().as_list()[1]
sampled_onehot = tf.one_hot(tf.argmax(nt_probs, 1), onehot_dim, 1.0, 0.0)
with tf.get_default_graph().gradient_override_map({'Ceil': 'Identity', 'Mul': 'STMul'}):
return tf.ceil(sampled_onehot * nt_probs)
@ops.RegisterGradient("STMul")
def st_mul(op, grad):
return [grad, grad]
#Gumbel Distribution Sampler
def gumbel_softmax(logits, temperature=0.5) :
gumbel_dist = tf.contrib.distributions.RelaxedOneHotCategorical(temperature, logits=logits)
batch_dim = logits.get_shape().as_list()[0]
onehot_dim = logits.get_shape().as_list()[1]
return gumbel_dist.sample()
#PWM Masking and Sampling helper functions
def mask_pwm(inputs) :
pwm, onehot_template, onehot_mask = inputs
return pwm * onehot_mask + onehot_template
def sample_pwm_st(pwm_logits) :
n_sequences = K.shape(pwm_logits)[0]
seq_length = K.shape(pwm_logits)[2]
flat_pwm = K.reshape(pwm_logits, (n_sequences * seq_length, 4))
sampled_pwm = st_sampled_softmax(flat_pwm)
return K.reshape(sampled_pwm, (n_sequences, 1, seq_length, 4))
def sample_pwm_gumbel(pwm_logits) :
n_sequences = K.shape(pwm_logits)[0]
seq_length = K.shape(pwm_logits)[2]
flat_pwm = K.reshape(pwm_logits, (n_sequences * seq_length, 4))
sampled_pwm = gumbel_softmax(flat_pwm, temperature=0.5)
return K.reshape(sampled_pwm, (n_sequences, 1, seq_length, 4))
#Generator helper functions
def initialize_sequence_templates(generator, sequence_templates, background_matrices) :
embedding_templates = []
embedding_masks = []
embedding_backgrounds = []
for k in range(len(sequence_templates)) :
sequence_template = sequence_templates[k]
onehot_template = iso.OneHotEncoder(seq_length=len(sequence_template))(sequence_template).reshape((1, len(sequence_template), 4))
for j in range(len(sequence_template)) :
if sequence_template[j] not in ['N', 'X'] :
nt_ix = np.argmax(onehot_template[0, j, :])
onehot_template[:, j, :] = -4.0
onehot_template[:, j, nt_ix] = 10.0
elif sequence_template[j] == 'X' :
onehot_template[:, j, :] = -1.0
onehot_mask = np.zeros((1, len(sequence_template), 4))
for j in range(len(sequence_template)) :
if sequence_template[j] == 'N' :
onehot_mask[:, j, :] = 1.0
embedding_templates.append(onehot_template.reshape(1, -1))
embedding_masks.append(onehot_mask.reshape(1, -1))
embedding_backgrounds.append(background_matrices[k].reshape(1, -1))
embedding_templates = np.concatenate(embedding_templates, axis=0)
embedding_masks = np.concatenate(embedding_masks, axis=0)
embedding_backgrounds = np.concatenate(embedding_backgrounds, axis=0)
generator.get_layer('template_dense').set_weights([embedding_templates])
generator.get_layer('template_dense').trainable = False
generator.get_layer('mask_dense').set_weights([embedding_masks])
generator.get_layer('mask_dense').trainable = False
generator.get_layer('background_dense').set_weights([embedding_backgrounds])
generator.get_layer('background_dense').trainable = False
#Generator construction function
def build_sampler(batch_size, seq_length, n_classes=1, n_samples=1, sample_mode='st') :
#Initialize Reshape layer
reshape_layer = Reshape((1, seq_length, 4))
#Initialize background matrix
onehot_background_dense = Embedding(n_classes, seq_length * 4, embeddings_initializer='zeros', name='background_dense')
#Initialize template and mask matrices
onehot_template_dense = Embedding(n_classes, seq_length * 4, embeddings_initializer='zeros', name='template_dense')
onehot_mask_dense = Embedding(n_classes, seq_length * 4, embeddings_initializer='ones', name='mask_dense')
#Initialize Templating and Masking Lambda layer
masking_layer = Lambda(mask_pwm, output_shape = (1, seq_length, 4), name='masking_layer')
background_layer = Lambda(lambda x: x[0] + x[1], name='background_layer')
#Initialize PWM normalization layer
pwm_layer = Softmax(axis=-1, name='pwm')
#Initialize sampling layers
sample_func = None
if sample_mode == 'st' :
sample_func = sample_pwm_st
elif sample_mode == 'gumbel' :
sample_func = sample_pwm_gumbel
upsampling_layer = Lambda(lambda x: K.tile(x, [n_samples, 1, 1, 1]), name='upsampling_layer')
sampling_layer = Lambda(sample_func, name='pwm_sampler')
permute_layer = Lambda(lambda x: K.permute_dimensions(K.reshape(x, (n_samples, batch_size, 1, seq_length, 4)), (1, 0, 2, 3, 4)), name='permute_layer')
def _sampler_func(class_input, raw_logits) :
#Get Template and Mask
onehot_background = reshape_layer(onehot_background_dense(class_input))
onehot_template = reshape_layer(onehot_template_dense(class_input))
onehot_mask = reshape_layer(onehot_mask_dense(class_input))
#Add Template and Multiply Mask
pwm_logits = masking_layer([background_layer([raw_logits, onehot_background]), onehot_template, onehot_mask])
#Compute PWM (Nucleotide-wise Softmax)
pwm = pwm_layer(pwm_logits)
#Tile each PWM to sample from and create sample axis
pwm_logits_upsampled = upsampling_layer(pwm_logits)
sampled_pwm = sampling_layer(pwm_logits_upsampled)
sampled_pwm = permute_layer(sampled_pwm)
sampled_mask = permute_layer(upsampling_layer(onehot_mask))
return pwm_logits, pwm, sampled_pwm, onehot_mask, sampled_mask
return _sampler_func
#Initialize Encoder and Decoder networks
batch_size = 32
seq_length = 205
n_samples = 128
sample_mode = 'st'
#sample_mode = 'gumbel'
#Load sampler
sampler = build_sampler(batch_size, seq_length, n_classes=1, n_samples=n_samples, sample_mode=sample_mode)
#Load Predictor
predictor_path = '../../../aparent/saved_models/aparent_plasmid_iso_cut_distalpas_all_libs_no_sampleweights_sgd.h5'
predictor = load_model(predictor_path)
predictor.trainable = False
predictor.compile(optimizer=keras.optimizers.SGD(lr=0.1), loss='mean_squared_error')
#Build scrambler model
dummy_class = Input(shape=(1,), name='dummy_class')
input_logits = Input(shape=(1, seq_length, 4), name='input_logits')
pwm_logits, pwm, sampled_pwm, pwm_mask, sampled_mask = sampler(dummy_class, input_logits)
scrambler_model = Model([input_logits, dummy_class], [pwm_logits, pwm, sampled_pwm, pwm_mask, sampled_mask])
#Initialize Sequence Templates and Masks
initialize_sequence_templates(scrambler_model, [sequence_template], [x_mean_logits])
scrambler_model.trainable = False
scrambler_model.compile(
optimizer=keras.optimizers.Adam(lr=0.001, beta_1=0.9, beta_2=0.999),
loss='mean_squared_error'
)
scrambler_model.summary()
file_names = [
"autoscrambler_dataset_apa_doubledope_sample_mode_st_n_samples_32_resnet_1_4_32_8_025_n_epochs_50_target_bits_025_smooth_importance_scores_test.npy",
"perturbation_apa_doubledope_importance_scores_test.npy",
"deepexplain_apa_doubledope_method_gradient_importance_scores_test.npy",
"deeplift_apa_doubledope_method_guided_backprop_importance_scores_test.npy",
"deepexplain_apa_doubledope_method_rescale_importance_scores_test.npy",
#"deeplift_apa_doubledope_method_revealcancel_importance_scores_test.npy",
"deepshap_apa_doubledope_importance_scores_test.npy",
"deepexplain_apa_doubledope_method_integrated_gradients_importance_scores_test.npy",
"extremal_apa_doubledope_mode_preserve_perturbation_blur_area_01_importance_scores_test.npy",
"extremal_apa_doubledope_mode_preserve_perturbation_fade_area_01_importance_scores_test.npy",
"pytorch_saliency_apa_doubledope_smaller_blur_importance_scores_test.npy",
"sufficient_input_subsets_apa_doubledope_thresh_07_mean_importance_scores_test.npy",
"autoscrambler_dataset_apa_doubledope_resnet_1_4_32_8_025_n_epochs_50_target_lum_005_gumbel_no_bg_lum_importance_scores_test.npy",
"l2x_apa_doubledope_importance_scores_test.npy",
"invase_apa_doubledope_conv_importance_scores_test.npy",
]
model_names =[
"scrambler",
"perturbation",
"gradient",
"guided_backprop",
"deeplift",
"deepshap",
"integrated_gradients",
"torchray_blur",
"torchray_fade",
"saliency_model",
"sis_mean",
"zero_scrambler",
"l2x",
"invase",
]
model_importance_scores_test = [np.load(file_name) for file_name in file_names]
for model_i in range(len(model_names)) :
if model_importance_scores_test[model_i].shape[-1] > 1 :
model_importance_scores_test[model_i] = np.sum(model_importance_scores_test[model_i], axis=-1, keepdims=True)
feature_quantiles = [0.80, 0.90, 0.95, 0.98]
on_state_logit_val = 50.
dummy_test = np.zeros((x_test.shape[0], 1))
x_test_logits = 2. * x_test - 1.
aparent_l_test = np.zeros((x_test.shape[0], 13))
aparent_l_test[:, 4] = 1.
aparent_d_test = np.ones((x_test.shape[0], 1))
y_pred_ref = predictor.predict([np.transpose(x_test, (0, 2, 3, 1)), aparent_l_test, aparent_d_test], batch_size=32, verbose=True)[0]
acc_ref = np.sum(np.sign(y_pred_ref[:, 0] - 0.5) == np.sign(y_test[:, 0] - 0.5)) / float(x_test.shape[0])
_, _, _, pwm_mask, sampled_mask = scrambler_model.predict([x_test_logits, dummy_test], batch_size=batch_size)
model_kl_divergences = []
model_accs = []
for model_i in range(len(model_names)) :
print("Benchmarking model '" + str(model_names[model_i]) + "'...")
feature_quantile_kl_divergences = []
feature_quantile_accs = []
for feature_quantile_i, feature_quantile in enumerate(feature_quantiles) :
print("Feature quantile = " + str(feature_quantile))
importance_scores_test = None
if model_names[model_i] not in ['zero_scrambler', 'l2x'] :
importance_scores_test = np.abs(model_importance_scores_test[model_i])
else :
importance_scores_test = model_importance_scores_test[model_i]
n_to_test = importance_scores_test.shape[0] // batch_size * batch_size
importance_scores_test = importance_scores_test[:n_to_test]
importance_scores_test *= np.expand_dims(np.max(pwm_mask[:n_to_test], axis=-1), axis=-1)
quantile_vals = np.quantile(importance_scores_test, axis=(1, 2, 3), q=feature_quantile, keepdims=True)
quantile_vals = np.tile(quantile_vals, (1, importance_scores_test.shape[1], importance_scores_test.shape[2], importance_scores_test.shape[3]))
top_logits_test = np.zeros(importance_scores_test.shape)
top_logits_test[importance_scores_test > quantile_vals] = on_state_logit_val
top_logits_test = np.tile(top_logits_test, (1, 1, 1, 4)) * x_test_logits[:n_to_test]
_, _, samples_test, _, _ = scrambler_model.predict([top_logits_test, dummy_test[:n_to_test]], batch_size=batch_size)
mean_kl_divs = []
mean_accs = []
for data_ix in range(samples_test.shape[0]) :
if data_ix % 100 == 0 :
print("Processing example " + str(data_ix) + "...")
y_pred_var_samples = predictor.predict([np.transpose(samples_test[data_ix, ...], (0, 2, 3, 1)), aparent_l_test[:n_samples], aparent_d_test[:n_samples]], batch_size=n_samples)[0][:, 0]
y_pred_ref_samples = np.tile(y_pred_ref[data_ix, :], (n_samples,))
y_pred_var_samples = np.clip(y_pred_var_samples, 1e-6, 1. - 1e-6)
y_pred_ref_samples = np.clip(y_pred_ref_samples, 1e-6, 1. - 1e-6)
left_kl_divs = y_pred_ref_samples * np.log(y_pred_ref_samples / y_pred_var_samples) + (1. - y_pred_ref_samples) * np.log((1. - y_pred_ref_samples) / (1. - y_pred_var_samples))
right_kl_divs = y_pred_var_samples * np.log(y_pred_var_samples / y_pred_ref_samples) + (1. - y_pred_var_samples) * np.log((1. - y_pred_var_samples) / (1. - y_pred_ref_samples))
mean_kl_div = np.mean(left_kl_divs + right_kl_divs)
mean_acc = np.sum(np.sign(y_pred_var_samples - 0.5) == np.sign(y_test[data_ix, 0] - 0.5)) / float(y_pred_var_samples.shape[0])
mean_kl_divs.append(mean_kl_div)
mean_accs.append(1. if mean_acc > 0.9 else 0.)
mean_kl_divs = np.array(mean_kl_divs)
mean_accs = np.array(mean_accs)
feature_quantile_kl_divergences.append(mean_kl_divs)
feature_quantile_accs.append(mean_accs)
model_kl_divergences.append(feature_quantile_kl_divergences)
model_accs.append(feature_quantile_accs)
model_names =[
"scrambler",
"perturbation",
"gradient",
"guided\nbackprop",
"deeplift\nrescale",
"deepshap",
"integrated\ngradients",
"torchray\nblur",
"torchray\nfade",
"saliency\nmodel",
"sis\nmean",
"zero\nscrambler",
"l2x",
"invase",
]
#Store benchmark results as tables
benchmark_name = "benchmark_inclusion_apa_doubledope_new_rescale_smooth"
save_figs = True
median_kl_table = np.zeros((len(model_kl_divergences), len(model_kl_divergences[0])))
mean_acc_table = np.zeros((len(model_accs), len(model_accs[0])))
for i, model_name in enumerate(model_names) :
for j, feature_quantile in enumerate(feature_quantiles) :
median_kl_table[i, j] = np.median(model_kl_divergences[i][j])
mean_acc_table[i, j] = np.mean(model_accs[i][j])
#Plot and store median kl table
f = plt.figure(figsize = (4, 6))
cells = np.round(median_kl_table, 3).tolist()
print("--- Median KL Divergences ---")
max_len = np.max([len(model_name.upper().replace("\n", " ")) for model_name in model_names])
print(("-" * max_len) + " " + " ".join([(str(feature_quantile) + "0")[:4] for feature_quantile in feature_quantiles]))
for i in range(len(cells)) :
curr_len = len([model_name.upper().replace("\n", " ") for model_name in model_names][i])
row_str = [model_name.upper().replace("\n", " ") for model_name in model_names][i] + (" " * (max_len - curr_len))
for j in range(len(cells[i])) :
cells[i][j] = (str(cells[i][j]) + "00000")[:4]
row_str += " " + cells[i][j]
print(row_str)
print("")
table = plt.table(cellText=cells, rowLabels=[model_name.upper().replace("\n", " ") for model_name in model_names], colLabels=feature_quantiles, loc='center')
ax = plt.gca()
#f.patch.set_visible(False)
ax.axis('off')
ax.axis('tight')
plt.tight_layout()
if save_figs :
plt.savefig(benchmark_name + "_kl_table.png", dpi=300, transparent=True)
plt.savefig(benchmark_name + "_kl_table.eps")
plt.show()
#Plot and store accuracy table
f = plt.figure(figsize = (4, 6))
cells = np.concatenate([np.array([acc_ref] * mean_acc_table.shape[0]).reshape(-1, 1), mean_acc_table], axis=1)
cells = np.round(cells, 3).tolist()
print("--- Mean Accuracies ---")
max_len = np.max([len(model_name.upper().replace("\n", " ")) for model_name in model_names])
print(("-" * max_len) + " " + " ".join([(str(feature_quantile) + "0")[:4] for feature_quantile in (["REF "] + feature_quantiles)]))
for i in range(len(cells)) :
curr_len = len([model_name.upper().replace("\n", " ") for model_name in model_names][i])
row_str = [model_name.upper().replace("\n", " ") for model_name in model_names][i] + (" " * (max_len - curr_len))
for j in range(len(cells[i])) :
cells[i][j] = (str(cells[i][j]) + "00000")[:4]
row_str += " " + cells[i][j]
print(row_str)
table = plt.table(cellText=cells, rowLabels=[model_name.upper().replace("\n", " ") for model_name in model_names], colLabels=["REF"] + feature_quantiles, loc='center')
ax = plt.gca()
#f.patch.set_visible(False)
ax.axis('off')
ax.axis('tight')
plt.tight_layout()
if save_figs :
plt.savefig(benchmark_name + "_acc_table.png", dpi=300, transparent=True)
plt.savefig(benchmark_name + "_acc_table.eps")
plt.show()
def lighten_color(color, amount=0.5):
import matplotlib.colors as mc
import colorsys
try:
c = mc.cnames[color]
except:
c = color
c = colorsys.rgb_to_hls(*mc.to_rgb(c))
return colorsys.hls_to_rgb(c[0], 1 - amount * (1 - c[1]), c[2])
fig = plt.figure(figsize=(13, 6))
benchmark_name = "benchmark_inclusion_apa_doubledope_new_rescale_smooth"
save_figs = True
width = 0.2
max_y_val = None
cm = plt.get_cmap('viridis_r')
shades = [0.4, 0.6, 0.8, 1]
quantiles = [0.5, 0.8, 0.9, 0.95]
all_colors = plt.rcParams['axes.prop_cycle'].by_key()['color'] + plt.rcParams['axes.prop_cycle'].by_key()['color']
model_colors = {model_names[i]: all_colors[i] for i in range(len(model_names))}
results = np.zeros((len(quantiles), len(model_names), 1))
for i in range(1, len(feature_quantiles) + 1) :
for j in range(len(model_names)):
kl_div_samples = model_kl_divergences[j][i-1]
for l in range(len(quantiles)):
quantile = quantiles[l]
results[l, j, 0] = np.quantile(kl_div_samples, q=quantile)
xs = range(len(model_names))
xs = [xi + i*width for xi in xs]
for j in range(len(model_names)) :
for l in range(len(quantiles)) :
model_name = model_names[j]
c = model_colors[model_name]
val = results[l, j, 0]
if i == 1 and j == 0 :
lbl = "$%i^{th}$ Perc." % int(100*quantiles[l])
else :
lbl=None
if l == 0 :
plt.bar(xs[j], val, width=width, color=lighten_color(c, shades[l]), edgecolor='k', linewidth=1, label=lbl, zorder=l+1)
else :
prev_val = results[l-1, j].mean(axis=-1)
plt.bar(xs[j],val-prev_val, width=width, bottom = prev_val, color=lighten_color(c, shades[l]), edgecolor='k', linewidth=1, label=lbl, zorder=l+1)
#if l == len(quantiles) - 1 and (max_y_val is None or val < 0.95 * max_y_val) :
# plt.text(xs[j], val, "Top\n" + str(int(100 - 100 * feature_quantiles[i-1])) + "%", horizontalalignment='center', verticalalignment='bottom', fontdict={ 'family': 'serif', 'color': 'black', 'weight': 'bold', 'size': 10 })
prev_results = results
plt.xticks([i + 2.5*width for i in range(len(model_names))])
all_lbls = [model_names[j].upper() for j in range(len(model_names))]
plt.gca().set_xticklabels(all_lbls, rotation=60)
plt.ylabel("Test Set KL-Divergence")
#max_y_val = np.max(results) * 1.1
if max_y_val is not None :
plt.ylim([0, max_y_val])
plt.grid(True)
plt.gca().set_axisbelow(True)
plt.gca().grid(color='gray', alpha=0.2)
plt.gca().spines['right'].set_visible(False)
plt.gca().spines['top'].set_visible(False)
plt.gca().yaxis.set_ticks_position('left')
plt.gca().xaxis.set_ticks_position('bottom')
plt.legend(fontsize=12, frameon=True, loc='upper left')
leg = plt.gca().get_legend()
for l in range(len(quantiles)):
leg.legendHandles[l].set_color(lighten_color(all_colors[7], shades[l]))
leg.legendHandles[l].set_edgecolor('k')
plt.tight_layout()
if save_figs :
plt.savefig(benchmark_name + ".png", dpi=300, transparent=True)
plt.savefig(benchmark_name + ".eps")
plt.show()
#Gradient saliency/backprop visualization
import matplotlib.collections as collections
import operator
import matplotlib.pyplot as plt
import matplotlib.cm as cm
import matplotlib.colors as colors
import matplotlib as mpl
from matplotlib.text import TextPath
from matplotlib.patches import PathPatch, Rectangle
from matplotlib.font_manager import FontProperties
from matplotlib import gridspec
from matplotlib.ticker import FormatStrFormatter
def plot_importance_scores(importance_scores, ref_seq, figsize=(12, 2), score_clip=None, sequence_template='', plot_start=0, plot_end=96, plot_sequence_template=False, save_figs=False, fig_name=None) :
end_pos = ref_seq.find("#")
fig = plt.figure(figsize=figsize)
ax = plt.gca()
if score_clip is not None :
importance_scores = np.clip(np.copy(importance_scores), -score_clip, score_clip)
max_score = np.max(np.sum(importance_scores[:, :], axis=0)) + 0.01
for i in range(0, len(ref_seq)) :
if sequence_template[i] == 'N' :
mutability_score = np.sum(importance_scores[:, i])
dna_letter_at(ref_seq[i], i + 0.5, 0, mutability_score, ax)
elif plot_sequence_template :
dna_letter_at(ref_seq[i], i + 0.5, 0, max_score, ax, color='black')
plt.sca(ax)
plt.xlim((0, len(ref_seq)))
plt.ylim((0, max_score))
plt.axis('off')
#plt.yticks([0.0, max_score], [0.0, max_score], fontsize=16)
for axis in fig.axes :
axis.get_xaxis().set_visible(False)
axis.get_yaxis().set_visible(False)
plt.tight_layout()
if save_figs :
plt.savefig(fig_name + ".png", transparent=True, dpi=300)
plt.savefig(fig_name + ".eps")
plt.show()
#Visualize a few example patterns
save_figs = True
plot_examples = [3, 4, 6, 7]
feature_quantiles = [0.80, 0.90, 0.95, 0.98]
score_clips = [
None,
0.5,
0.5,
0.5,
0.5,
0.5,
0.5,
None,
None,
None,
None,
None,
None,
None,
]
on_state_logit_val = 50.
encoder = isol.OneHotEncoder(205)
dummy_test = np.zeros((x_test.shape[0], 1))
x_test_logits = 2. * x_test - 1.
aparent_l_test = np.zeros((x_test.shape[0], 13))
aparent_l_test[:, 4] = 1.
aparent_d_test = np.ones((x_test.shape[0], 1))
y_pred_ref = predictor.predict([np.transpose(x_test, (0, 2, 3, 1)), aparent_l_test, aparent_d_test], batch_size=32, verbose=True)[0]
_, _, _, pwm_mask, sampled_mask = scrambler_model.predict([x_test_logits, dummy_test], batch_size=batch_size)
for data_ix in plot_examples :
print("Test pattern = " + str(data_ix) + ":")
y_test_hat_ref = predictor.predict(x=[np.expand_dims(np.expand_dims(x_test[data_ix, 0, :, :], axis=0), axis=-1), aparent_l_test[:1], aparent_d_test[:1]], batch_size=1)[0][0, 0]
print(" - Prediction (original) = " + str(round(y_test_hat_ref, 2))[:4])
plot_dna_logo(x_test[data_ix, 0, :, :], sequence_template=sequence_template, figsize=(14, 0.65), plot_start=0, plot_end=205, plot_sequence_template=True, save_figs=save_figs, fig_name="benchmark_inclusion_" + dataset_name + "_test_ix_" + str(data_ix))
for model_i in range(len(model_names)) :
print("Model = '" + str(model_names[model_i]) + "'...")
if model_names[model_i] not in ['zero\nscrambler', 'l2x'] :
if len(model_importance_scores_test[model_i].shape) >= 5 :
importance_scores_test = np.abs(model_importance_scores_test[model_i][1, ...])
else :
importance_scores_test = np.abs(model_importance_scores_test[model_i])
else :
importance_scores_test = model_importance_scores_test[model_i]
importance_scores_test = importance_scores_test[:32]
importance_scores_test *= np.expand_dims(np.max(pwm_mask[:32], axis=-1), axis=-1)
y_test_hat_mean_qts = []
for feature_quantile_i, feature_quantile in enumerate(feature_quantiles) :
quantile_vals = np.quantile(importance_scores_test, axis=(1, 2, 3), q=feature_quantile, keepdims=True)
quantile_vals = np.tile(quantile_vals, (1, importance_scores_test.shape[1], importance_scores_test.shape[2], importance_scores_test.shape[3]))
top_logits_test = np.zeros(importance_scores_test.shape)
top_logits_test[importance_scores_test > quantile_vals] = on_state_logit_val
top_logits_test = np.tile(top_logits_test, (1, 1, 1, 4)) * x_test_logits[:32]
_, _, samples_test, _, _ = scrambler_model.predict([top_logits_test, dummy_test[:32]], batch_size=batch_size)
y_test_hat = predictor.predict(x=[np.expand_dims(samples_test[data_ix, :, 0, :, :], axis=-1), aparent_l_test[:n_samples], aparent_d_test[:n_samples]], batch_size=32)[0][:, 0]
y_test_hat_mean = np.mean(y_test_hat)
y_test_hat_mean_qts.append(str(round(y_test_hat_mean, 2))[:4])
print(" - Prediction (scrambled qts) = " + str(y_test_hat_mean_qts))
if model_names[model_i] in ['scrambler'] :
scrambled_logits_test = np.tile(importance_scores_test[:32], (1, 1, 1, 4)) * x_test_logits[:32]
_, pwm_test, samples_test, _, _ = scrambler_model.predict_on_batch([scrambled_logits_test, dummy_test[:32]])
y_test_hat = predictor.predict(x=[np.expand_dims(samples_test[data_ix, :, 0, :, :], axis=-1), aparent_l_test[:n_samples], aparent_d_test[:n_samples]], batch_size=32)[0][:, 0]
y_test_hat_mean = np.mean(y_test_hat)
print(" - Prediction (scrambled natural) = " + str(round(y_test_hat_mean, 2))[:4])
subtracted_pwm_test = np.exp(scrambled_logits_test) / np.sum(np.exp(scrambled_logits_test), axis=-1, keepdims=True)
plot_dna_logo(subtracted_pwm_test[data_ix, 0, :, :], sequence_template=sequence_template, figsize=(14, 0.65), plot_start=0, plot_end=205, plot_sequence_template=True, save_figs=save_figs, fig_name="benchmark_inclusion_" + dataset_name + "_test_ix_" + str(data_ix) + "_" + model_names[model_i])
else :
plot_importance_scores(importance_scores_test[data_ix, 0, :, :].T, encoder.decode(x_test[data_ix, 0, :, :]), figsize=(14, 0.65), score_clip=score_clips[model_i], sequence_template=sequence_template, plot_start=0, plot_end=205, plot_sequence_template=True, save_figs=save_figs, fig_name="benchmark_inclusion_" + dataset_name + "_test_ix_" + str(data_ix) + "_" + model_names[model_i])
```
| github_jupyter |
```
import numpy as np
import networkx as nx
import matplotlib.pyplot as plt
from IPython.display import Image
from collections import defaultdict
from sklearn.metrics.pairwise import euclidean_distances
```
# 1 Node Classification [25 points]
### 1.1 Relational Classification [10 points]
As we discussed in class, we can use relational classification to predict node labels. Consider the
graph G as shown in Figure 1. We would like to classify nodes into 2 classes ”+” and ”-”. Labels
for node 3, 5, 8 and 10 are given (red for ”+”, blue for ”-”). Recall that using a probabilistic
relational classifier to predict label Y i for node i is defined as:
$ P(Y_{i} = c) = \frac{1}{|N_{i}|} \sum_{(i,j) \in E} W(i,j) P(Y_{j} = c)$
where $|N_{i}|$ is the number of neighbors of node i. Assume all the edges have edge weight $W (i, j) = 1$ in this graph. For labeled nodes, initialize with the ground-truth Y labels, i.e., $P (Y_{3} = +) = P(Y_{5} = +) = 1.0, P(Y_{8} = +) = P(Y_{10} = +) = 0 $. For unlabeled nodes, use unbiased initialization $P(Y_{i}= +) = 0.5$. Update nodes by node ID in ascending order (i.e., update node 1 first, then node 2, etc.)
```
edges = [(1,2), (1,3), (2,3),
(2,4), (3,6), (4,7),
(4,8), (5,8), (5,9),
(5,6), (6,9), (6, 10),
(7,8), (8,9), (9,10)]
nodes = list(range(10))
pos_nodes = [3,5]
neg_nodes = [8, 10]
G = defaultdict(list)
for i, j in edges:
G[i-1].append(j-1)
G[j-1].append(i-1)
G = dict(G)
```
#### (i) After the second iteration, give $P(Y_{i} = +)$ for i = 2, 4, 6. [6 points]
```
def Q1_1(nb_it=20, ep=1e-6, idxs=[2,4,6]):
P = [ 0.5 for i in nodes]
for i in pos_nodes:
P[i-1] = 1
for i in neg_nodes:
P[i-1] = 0
print(P)
it = 1
has_change = True
while (it <= nb_it) and has_change:
print(f'It: {it}')
has_change = False
for node in nodes:
if (node+1 in pos_nodes) or (node+1 in neg_nodes):
continue
nids = G[node]
p = sum([P[j] for j in nids])
p_new = p / len(nids)
if abs(P[node] - p_new) > ep:
has_change = True
P[node] = p_new
if it == 2:
print([(i, P[i-1]) for i in idxs])
it += 1
return P
P = Q1_1()
```
#### If we use 0.5 as the probability threshold, i.e., consider a node i belonging to class ”+” if $P(Y_{i} = +) > 0.5$, which node will belong to class ”+” in the end? Which will belong to class”-”? [4 points]
```
P
for n, p in zip(nodes, P):
if p > 0.5:
print(f'Node: {n+1} ;P={p:.2f}--> +')
else:
print(f'Node: {n+1} ;P={p:.2f}--> -')
```
### 1.2 Belief Propagation [15 points]
* reading: http://helper.ipam.ucla.edu/publications/gss2013/gss2013_11344.pdf
In this problem, we will be using Belief Propagation(BP) on Conditional Random Field to solve inference problems. Conditional Random Fields (CRF) are an important special case of Markov Random Fields to model conditional probability distribution. They define a probability distribution p over variables
* Message $ m_{ij}(x_{j})$: can be intuitively understood as a message from hidden node $x_{i}$ to hidden node $x_{j}$ about what state node i thinks node j should be in.
$ m_{ij}(x_{j}) = \sum_{x_{i}} φ_{i}(x_{i}) ψ_{ij}(x_{i}, x_{j}) \prod_{k\in N_{i} / j} m_{ki}(x_{i})$
* Belief $ b_{i}(x_{i})$ : $ b_{i}(x_{i}) = \frac{1}{Z} φ_{i}(x_{i}) \prod_{j \in N_{i}} m_{ji}(x_{i})$, where Z denotes the normalizing constant that ensures elements in $b_{i}(x_{i})$ sum to 1
##### (i) Consider the network with four hidden nodes shown in Figure 3. Compute the belief at node 1, b 1 (x 1 ), using the belief propagation rules, and write the result in terms of φ’s and ψ’s. [5 points]
* $ b_{1}(x_{1}) = \frac{1}{Z} φ_{1}(x_{1}) m_{21}(x_{1})$
* $\quad\quad\quad = \frac{1}{Z} φ_{1}(x_{1}) m_{21}(x_{1})$
* $ m_{21}(x_{1}) = \sum_{x_{2}} φ_{2}(x_{2}) ψ_{21}(x_{2}, x_{1}) m_{32}(x_{2}) m_{42}(x_{2})$
* $ \quad\quad\quad = \sum_{x_{2}} φ_{2}(x_{2}) ψ_{21}(x_{2}, x_{1}) \sum_{x_{3}} φ_{3}(x_{3}) ψ_{32}(x_{3}, x_{2}) \sum_{x_{4}} φ_{4}(x_{4}) ψ_{42}(x_{4}, x_{2})$
* $ \quad\quad\quad = \sum_{x_{2}} \sum_{x_{3}} \sum_{x_{4}} φ_{2}(x_{2}) φ_{3}(x_{3}) φ_{4}(x_{4}) ψ_{21}(x_{2}, x_{1}) ψ_{32}(x_{3}, x_{2}) ψ_{42}(x_{4}, x_{2})$
* $ b_{1}(x_{1}) = \frac{1}{Z} φ_{1}(x_{1}) \sum_{x_{2}} \sum_{x_{3}} \sum_{x_{4}} φ_{2}(x_{2}) φ_{3}(x_{3}) φ_{4}(x_{4}) ψ_{21}(x_{2}, x_{1}) ψ_{32}(x_{3}, x_{2}) ψ_{42}(x_{4}, x_{2})$
#### (ii) Prove that the belief at node 1 calculated above, $b_{1}(x_{1})$, is the same as the marginal probability of x 1 conditioned on the observations, $p(x_{1} |y_{1}, y_{2}, y_{3}, y_{4})$. [2 points],
ref: http://helper.ipam.ucla.edu/publications/gss2013/gss2013_11344.pdf
* $ b_{1}(x_{1}) = \frac{1}{Z} φ_{1}(x_{1}) \sum_{x_{2}} \sum_{x_{3}} \sum_{x_{4}} φ_{2}(x_{2}) φ_{3}(x_{3}) φ_{4}(x_{4}) ψ_{21}(x_{2}, x_{1}) ψ_{32}(x_{3}, x_{2}) ψ_{42}(x_{4}, x_{2})$
* $ b_{1}(x_{1}) = \frac{1}{Z} \sum_{x_{2}} \sum_{x_{3}} \sum_{x_{4}} φ_{1}(x_{1}) φ_{2}(x_{2}) φ_{3}(x_{3}) φ_{4}(x_{4}) ψ_{21}(x_{2}, x_{1}) ψ_{32}(x_{3}, x_{2}) ψ_{42}(x_{4}, x_{2})$
* $ b_{1}(x_{1}) = \frac{1}{Z} \sum_{x_{2}} \sum_{x_{3}} \sum_{x_{4}} p(x_{1}, x_{2}, x_{3}, x_{4} |y_{1}, y_{2}, y_{3}, y_{4})$
* $ b_{1}(x_{1}) = \frac{1}{Z} p(x_{1} |y_{1}, y_{2}, y_{3}, y_{4})$
#### (iii) Let’s work with a graph without cycles as shown in Figure 4. Assume x and y only have two states (0 and 1) and the graphical model has 5 hidden variables, and two variables observed with y 2 = 0, y 4 = 1. The compatibility matrices are given in the arrays below
```
# States --> {0, 1}
y2 = 0
y4 = 1
psi_1_2 = np.array([[1, 0.9],
[0.9, 1]])
psi_3_4 = psi_1_2
psi_2_3 = np.array([[0.1, 1],
[1, 0.1]])
psi_3_5 = psi_2_3
phi_2 = np.array([[1, 0.1],
[0.1, 1]])
phi_4 = phi_2
```
$ m_{53}(x_{3}) = \sum_{x_{5}} φ_{5}(x_{5}) ψ_{53}(x_{5}, x_{3}) $
$ m_{43}(x_{3}) = \sum_{x_{4}} φ_{4}(x_{4}) ψ_{43}(x_{4}, x_{3}) $
$ m_{32}(x_{2}) = \sum_{x_{3}} φ_{3}(x_{3}) ψ_{32}(x_{3}, x_{4}) m_{43}(x_{3}) m_{53}(x_{3})$
$ m_{21}(x_{1}) = \sum_{x_{2}} φ_{2}(x_{2}) ψ_{21}(x_{2}, x_{1}) m_{32}(x_{2})$
$ m_{12}(x_{2}) = \sum_{x_{1}} φ_{1}(x_{1}) ψ_{12}(x_{1}, x_{2})$
$ m_{23}(x_{3}) = \sum_{x_{2}} φ_{2}(x_{2}) ψ_{23}(x_{2}, x_{3}) m_{12}(x_{2})$
$ m_{34}(x_{4}) = \sum_{x_{3}} φ_{3}(x_{3}) ψ_{34}(x_{3}, x_{4}) m_{23}(x_{3}) m_{53}(x_{3})$
$ m_{35}(x_{5}) = \sum_{x_{3}} φ_{3}(x_{3}) ψ_{35}(x_{3}, x_{5}) m_{23}(x_{3}) m_{43}(x_{3})$
```
m53_x3 = np.sum(psi_3_5, axis=0)
m43_x3 = np.dot(psi_3_4, phi_4[:, y4])
m32_x2 = np.dot(psi_2_3, (m53_x3 * m43_x3).reshape(-1, 1))
m21_x1 = np.dot(psi_1_2, phi_2[:,y2].reshape(-1,1) * m32_x2)
m12_x2 = np.sum(psi_1_2, axis=0)
m23_x3 = np.dot(psi_2_3, phi_2[:,y2] * m12_x2)
m34_x4 = np.dot(psi_3_4, m23_x3 * m53_x3)
m35_x5 = np.dot(psi_3_5, m23_x3 * m43_x3)
```
$ b_{1}(x_{1}) = \frac{1}{Z} φ_{1}(x_{1}) m_{21}(x_{1})$
$ b_{2}(x_{2}) = \frac{1}{Z} φ_{2}(x_{2}) m_{12}(x_{2}) m_{32}(x_{2}) $
$ b_{3}(x_{3}) = \frac{1}{Z} φ_{3}(x_{3}) m_{23}(x_{3}) m_{43}(x_{3}) m_{53}(x_{3}) $
$ b_{4}(x_{4}) = \frac{1}{Z} φ_{4}(x_{4}) m_{34}(x_{4})$
$ b_{5}(x_{5}) = \frac{1}{Z} φ_{5}(x_{5}) m_{35}(x_{5})$
```
Z_1 = np.sum(m21_x1)
b1_x1 = (1/Z_1) * m21_x1
aux = phi_2[:,y2].reshape(-1, 1) * m12_x2.reshape(-1, 1) * m32_x2
Z_2 = np.sum(aux)
b2_x2 = (1/Z_2) * aux
aux = (m23_x3 * m43_x3 * m53_x3).reshape(-1, 1)
Z_3 = np.sum(aux)
b3_x3 = (1/Z_3) * aux
aux = (phi_4[:, y4] * m34_x4).reshape(-1, 1)
Z_4 = np.sum(aux)
b4_x4 = (1/Z_4) * aux
Z_5 = np.sum(m35_x5)
b5_x5 = (1/Z_5) * m35_x5.reshape(-1, 1)
for i, p in enumerate([b1_x1, b2_x2, b3_x3, b4_x4, b5_x5], 1):
print(f'b{i}(x{i}) = ')
print(p, "\n")
```
#### Is the prediction what you expected? [8 points]
* x2, x4 are strong biases for being in state 0, 1 (y2, y4)
* x1 shows a slight preference for state 0 (influences of x2)
* x3 strong biases for being in state 1 (compatibility matrix)(x3 tended to be opposite to x2 and slighty similiar to x4)
* x5 biases for being in state 1 (compatibility matrix)(x5 tended to be opposite to x3)
# 2 Node Embeddings with TransE [25 points]
### 2.1 Warmup: Why the Comparative Loss? [3 points]
* $ L_{simple} = \sum_{(h,l,t) \in S} d(\boldsymbol h + \boldsymbol l, \boldsymbol t)$
```
g = nx.DiGraph()
for node in [1,2,3,4]:
g.add_node(node)
for edge in [(1,2), (3,4)]:
g.add_edge(*edge)
nx.draw(g, with_labels=True, font_weight='bold')
labels = nx.get_edge_attributes(g, 'weight')
pos = nx.spring_layout(g)
nx.draw_networkx_edge_labels(g, pos, edge_labels=labels)
```
Let have only one relation, and the embeddings the followings:
* $ \boldsymbol l = [1, -1]$
* $ \boldsymbol e_{1} = \boldsymbol e_{3} = [0, 1]$
* $ \boldsymbol e_{2} = \boldsymbol e_{4} = [1, 0]$
```
l = np.array([[1, -1]])
e1 = np.array([[0, 1]])
e3 = e1
e2 = np.array([[1, 0]])
e4 = e2
```
* $ L_{simple} = d(\boldsymbol e_1 + \boldsymbol l, \boldsymbol e_2) + d(\boldsymbol e_1 + \boldsymbol l, \boldsymbol e_2) = 0$
```
l_simple = euclidean_distances(e1+l, e2)[0][0] + euclidean_distances(e3+l, e4)[0][0]
l_simple
samples = np.concatenate([e1, e2, e3, e4])
x = samples[:, 0]
y = samples[:, 1]
n = ["e1", "e2", "e3", "e4"]
fig, ax = plt.subplots()
ax.scatter(x=x, y=y)
for i, txt in enumerate(n):
ax.annotate(txt, (x[i], y[i]))
```
* So we had a loss value of 0 and useless embeddings because $ e_{1}$ is equal to $e_{3} $ and $e_{2} $ to $e_{4}$ but in the graph it make non sense to be close enough.
### 2.2 The Purpose of the Margin [5 points]
* Same example as above.
```
neg_sampling_1 = {
euclidean_distances(e1+l, e3)[0][0],
euclidean_distances(e1+l, e4)[0][0],
euclidean_distances(e3+l, e2)[0][0],
euclidean_distances(e4+l, e2)[0][0]
}
neg_sampling_2 = {
euclidean_distances(e3+l, e1)[0][0],
euclidean_distances(e3+l, e2)[0][0],
euclidean_distances(e1+l, e4)[0][0],
euclidean_distances(e2+l, e4)[0][0]
}
l_margin1 = np.sum([
max(euclidean_distances(e1+l, e2)[0][0] - neg_sampling, 0)
for neg_sampling in neg_sampling_1
])
l_margin2 = np.sum([
max(euclidean_distances(e3+l, e4)[0][0] - neg_sampling, 0)
for neg_sampling in neg_sampling_2
])
l_margin = l_margin1 + l_margin2
l_margin
```
# 3 GNN Expressiveness [25 points]
## 3.1 Effect of Depth on Expressiveness [7 Points]
(i) Consider the following 2 graphs, where all nodes have 1-dimensional feature [1]. We use a simplified version of GNN, with no nonlinearity and linear layers, and sum aggregation. We run the GNN to compute node embeddings for the 2 red nodes respectively. Note that the 2 red nodes have different 4-hop neighborhood structure. How many layers of message passing are needed so that these 2 nodes can be distinguished (i.e., have different GNN embeddings)?
* 3
(ii) Consider training a GNN on a node classification task, where nodes are classified to be positive
if they are part of an induced cyclic subgraph of length 10, and negative otherwise. Give an
example of a graph and the node which should be classified as True. Show that no GNN with
fewer than 5 layers can perfectly perform this classification task.
*
### 3.2 Relation to Random Walk [7 Points]
* (i) $ P = D^{-1} A$
* (ii) $ P = \frac{1}{2} D^{-1} A + \frac{1}{2} I$
### 3.3 Over-Smoothing Effect [5 Points]
*
### 3.4 Learning BFS with GNN [6 Points]
* (i)
* (ii) $ h_{i}^{l+1} = MAX([h_{j}^{l} | j \in N_{i}])$
# 4 GNN training [25 Points]
### 4.1 Examine Dataset [1 Point]
(i) Warm-up. First let’s examine the data. To do this, you will need use train.py. How many
nodes are there in the test set of CORA? How many graphs are there in the test set of
ENZYMES?
```
from torch_geometric.datasets import TUDataset
from torch_geometric.datasets import Planetoid
from torch_geometric.data import DataLoader
enzymes_dataset = TUDataset(root='/tmp/ENZYMES', name='ENZYMES')
cora_dataset = Planetoid(root='/tmp/Cora', name='Cora')
len(enzymes_dataset), len(cora_dataset)
nb_nodes_test_set = cora_dataset.data.test_mask.sum().item()
print(f'Core Dataset:\n >Number of Nodes in test set: {nb_nodes_test_set}')
data_size = len(enzymes_dataset)
nb_g_test_set = len(enzymes_dataset[int(data_size * 0.8):])
print(f'ENZYMES Dataset:\n >Number of Graphs in test set: {nb_g_test_set}')
```
### 4.3 Training [9 Points]
#### Cora:
```
Image(filename='q4_starter_code/img/loss_cora.png')
Image(filename='q4_starter_code/img/acc_cora.png')
Image(filename='q4_starter_code/img/loss_enz.png')
Image(filename='q4_starter_code/img/acc_enz.png')
```
Conclusion:
* Without any hyper paramater tunning GCN seem to work better for both cases.
* GraphSage and GAT models for enzymes dataset are not learning nothing at all. We should work with the hyper parameters
| github_jupyter |
# Non-Parametric, Unsupervised UMAP and HDBSCAN
```
#Imports
### Unfortunately, UMAP takes a while to import. One of its dependencies (pynndescent) uses numba,
### which is the cause of the performance bottleneck here.
import cv2
import hdbscan
import json
import matplotlib.pyplot as plt
import numpy as np
import os
import pandas as pd
import random
import re
import seaborn as sns
import skimage
import umap
import umap.plot
from bokeh.embed import json_item
from bokeh.models import HoverTool
from bokeh.models.tools import LassoSelectTool
from bokeh.plotting import show as bokeh_show, output_notebook
from glob import glob
from hdbscan import HDBSCAN
from IPython.display import display, HTML, Image, Javascript
from pathlib import Path
from sklearn.preprocessing import StandardScaler
#Constants
IMG_SIZE = 256
IMAGES_DIR = '../images/final_to_match'
MAX_FILES = 300
```
We first load the images...
```
files = Path(IMAGES_DIR)
image_files = []
for file in files.rglob('*'):
ext = file.suffix.lower()
if ext == ".jpg": image_files.append(file)
if len(image_files) >= MAX_FILES: break
image_files[0:4]
```
Then apply additional preprocessing...
```
images = []
names = []
files = []
for i, file in enumerate(image_files):
file_str = str(file)
image = cv2.imread(file_str)
names.append(file.stem)
files.append(file_str)
image = cv2.cvtColor(image, cv2.COLOR_BGR2GRAY)
image = cv2.resize(image, (IMG_SIZE, IMG_SIZE))
image = image.astype('float32')
#image = (image - image.mean(axis=(0, 1), keepdims = True)) / image.std(axis=(0, 1), keepdims = True)
#image = cv2.normalize(image, None, 0, 1, cv2.NORM_MINMAX)
images.append(image)
progress = i/MAX_FILES * 100
if progress % 5 == 0: print(f'{progress}% done')
print("Complete!")
```
We then convert it to a numpy array.
```
images = np.asarray(images)
images.shape
data = images.reshape((images.shape[0], -1))
#data = StandardScaler().fit_transform(data)
data.shape
```
We then fit UMAP to the data.
```
mapper = umap.UMAP(
n_neighbors = 200,
min_dist = 0.5,
n_components = 2,
metric = 'euclidean',
random_state = 100,
densmap = False
).fit(data)
umap_res = mapper.transform(data)
```
We apply HDBSCAN to the UMAP results.
```
cluster = HDBSCAN(
algorithm ='best',
approx_min_span_tree = True,
gen_min_span_tree = False,
leaf_size = 40,
metric='euclidean',
min_cluster_size = 15,
min_samples = 15,
p = None
).fit(umap_res)
```
The results are shown below.
```
hover_data = pd.DataFrame(
index = np.arange(
data.shape[0]
),
data = {
"file": files,
'index' : np.arange(data.shape[0]),
'name': names,
'class': cluster.labels_,
'probability': cluster.probabilities_,
'outlier': [ 1 if item == -1 else 0 for item in cluster.labels_ ]
}
)
hover_data.head(10).drop("index", axis = 1)
hover = HoverTool(
tooltips="""
<div>
<div>
<img
src="@file" height="128" alt="@file" width="128"
style="float: left; margin: 0px 15px 15px 0px;"
border="2"
></img>
</div>
<div>
<span style="font-size: 10px; font-weight: bold;">@name</span>
<span style="font-size: 10px; color: #966;">[$index]</span>
</div>
<div>
<span style="font-size: 9px; color: #966;">@file</span>
</div>
<div>
<span style="font-size: 9px; color: #966;">Class: @class</span>
</div>
<div>
<span style="font-size: 9px; color: #966;">Probability: @probability</span>
</div>
<div>
<span style="font-size: 9px; color: #966;">Outlier: @outlier</span>
</div>
</div>
"""
)
p = umap.plot.interactive(mapper, labels = cluster.labels_,hover_data = hover_data, point_size = 5, interactive_text_search = False)
del p.tools[len(p.tools)-1]
p.add_tools(hover)
p.add_tools(LassoSelectTool())
output_notebook()
bokeh_show(p)
```
The front-end has been built using Vue.js. Therefore, the data must be consumable using a JS-friendly format. We can convert the Bokeh plot into a JSON blob, and visualise it using Bokeh.js.
```
p_json = json.dumps(json_item(p))
p_json[0:150]
display(HTML('<div id="umap"></div>'))
Javascript(f'''Bokeh.embed.embed_item({p_json}, "umap")''')
```
| github_jupyter |
```
import os
os.environ['CUDA_VISIBLE_DEVICES'] = ''
import malaya_speech.train.model.mini_jasper as jasper
import malaya_speech
import tensorflow as tf
import numpy as np
import json
with open('malaya-speech-sst-vocab.json') as fopen:
unique_vocab = json.load(fopen) + ['{', '}', '[']
featurizer = malaya_speech.tf_featurization.STTFeaturizer(
normalize_per_feature = True
)
X = tf.placeholder(tf.float32, [None, None])
X_len = tf.placeholder(tf.int32, [None])
batch_size = tf.shape(X)[0]
features = tf.TensorArray(dtype = tf.float32, size = batch_size, dynamic_size = True, infer_shape = False)
features_len = tf.TensorArray(dtype = tf.int32, size = batch_size)
init_state = (0, features, features_len)
def condition(i, features, features_len):
return i < batch_size
def body(i, features, features_len):
f = featurizer(X[i, :X_len[i]])
f_len = tf.shape(f)[0]
return i + 1, features.write(i, f), features_len.write(i, f_len)
_, features, features_len = tf.while_loop(condition, body, init_state)
features_len = features_len.stack()
padded_features = tf.TensorArray(dtype = tf.float32, size = batch_size)
maxlen = tf.reduce_max(features_len)
init_state = (0, padded_features)
def condition(i, padded_features):
return i < batch_size
def body(i, padded_features):
f = features.read(i)
f = tf.pad(f, [[0, maxlen - tf.shape(f)[0]], [0,0]])
return i + 1, padded_features.write(i, f)
_, padded_features = tf.while_loop(condition, body, init_state)
padded_features = padded_features.stack()
padded_features.set_shape((None, None, 80))
model = jasper.Model(padded_features, features_len, training = False)
logits = tf.layers.dense(model.logits['outputs'], len(unique_vocab) + 1)
seq_lens = model.logits['src_length']
logits = tf.transpose(logits, [1, 0, 2])
logits = tf.identity(logits, name = 'logits')
seq_lens = tf.identity(seq_lens, name = 'seq_lens')
decoded = tf.nn.ctc_beam_search_decoder(logits, seq_lens, beam_width=100, top_paths=1, merge_repeated=True)
preds = tf.sparse.to_dense(tf.to_int32(decoded[0][0]))
preds = tf.identity(preds, 'preds')
sess = tf.InteractiveSession()
sess.run(tf.global_variables_initializer())
var_lists = tf.get_collection(tf.GraphKeys.GLOBAL_VARIABLES)
saver = tf.train.Saver(var_list = var_lists)
saver.restore(sess, 'asr-mini-jasper-ctc/model.ckpt-365000')
files = [
'savewav_2020-11-26_22-36-06_294832.wav',
'savewav_2020-11-26_22-40-56_929661.wav',
'download.wav',
'husein-zolkepli.wav',
'mas-aisyah.wav',
'khalil-nooh.wav',
'wadi-annuar.wav',
'675.wav',
'664.wav',
]
ys = [malaya_speech.load(f)[0] for f in files]
padded, lens = malaya_speech.padding.sequence_1d(ys, return_len = True)
decoded = sess.run(preds, feed_dict = {X: padded, X_len: lens})
results = []
for i in range(len(decoded)):
results.append(malaya_speech.char.decode(decoded[i], lookup = unique_vocab).replace('<PAD>', ''))
results
results = []
for i in range(len(decoded)):
results.append(malaya_speech.char.decode(decoded[i], lookup = unique_vocab).replace('<PAD>', ''))
results
saver = tf.train.Saver()
saver.save(sess, 'asr-mini-jasper-ctc-output/model.ckpt')
strings = ','.join(
[
n.name
for n in tf.get_default_graph().as_graph_def().node
if ('Variable' in n.op
or 'Placeholder' in n.name
or 'logits' in n.name
or 't_logits' in n.name
or 'seq_lens' in n.name
or 'alphas' in n.name
or 'self/Softmax' in n.name)
and 'adam' not in n.name
and 'beta' not in n.name
and 'global_step' not in n.name
and 'Assign' not in n.name
]
)
strings.split(',')
def freeze_graph(model_dir, output_node_names):
if not tf.gfile.Exists(model_dir):
raise AssertionError(
"Export directory doesn't exists. Please specify an export "
'directory: %s' % model_dir
)
checkpoint = tf.train.get_checkpoint_state(model_dir)
input_checkpoint = checkpoint.model_checkpoint_path
absolute_model_dir = '/'.join(input_checkpoint.split('/')[:-1])
output_graph = absolute_model_dir + '/frozen_model.pb'
clear_devices = True
with tf.Session(graph = tf.Graph()) as sess:
saver = tf.train.import_meta_graph(
input_checkpoint + '.meta', clear_devices = clear_devices
)
saver.restore(sess, input_checkpoint)
output_graph_def = tf.graph_util.convert_variables_to_constants(
sess,
tf.get_default_graph().as_graph_def(),
output_node_names.split(','),
)
with tf.gfile.GFile(output_graph, 'wb') as f:
f.write(output_graph_def.SerializeToString())
print('%d ops in the final graph.' % len(output_graph_def.node))
freeze_graph('asr-mini-jasper-ctc-output', strings)
def load_graph(frozen_graph_filename):
with tf.gfile.GFile(frozen_graph_filename, 'rb') as f:
graph_def = tf.GraphDef()
graph_def.ParseFromString(f.read())
with tf.Graph().as_default() as graph:
tf.import_graph_def(graph_def)
return graph
g = load_graph('asr-mini-jasper-ctc-output/frozen_model.pb')
x = g.get_tensor_by_name('import/Placeholder:0')
x_lens = g.get_tensor_by_name('import/Placeholder_1:0')
logits = g.get_tensor_by_name('import/logits:0')
seq_lens = g.get_tensor_by_name('import/seq_lens:0')
test_sess = tf.InteractiveSession(graph = g)
result = test_sess.run([logits, seq_lens], feed_dict = {x: padded, x_lens: lens})
result
from tensorflow.tools.graph_transforms import TransformGraph
transforms = ['add_default_attributes',
'remove_nodes(op=Identity, op=CheckNumerics, op=Dropout)',
'fold_batch_norms',
'fold_old_batch_norms',
'quantize_weights(fallback_min=-10, fallback_max=10)',
'strip_unused_nodes',
'sort_by_execution_order']
pb = 'asr-mini-jasper-ctc-output/frozen_model.pb'
input_graph_def = tf.GraphDef()
with tf.gfile.FastGFile(pb, 'rb') as f:
input_graph_def.ParseFromString(f.read())
transformed_graph_def = TransformGraph(input_graph_def,
['Placeholder', 'Placeholder_1'],
['logits', 'seq_lens'], transforms)
with tf.gfile.GFile(f'{pb}.quantized', 'wb') as f:
f.write(transformed_graph_def.SerializeToString())
g = load_graph(f'{pb}.quantized')
x = g.get_tensor_by_name('import/Placeholder:0')
x_lens = g.get_tensor_by_name('import/Placeholder_1:0')
logits = g.get_tensor_by_name('import/logits:0')
seq_lens = g.get_tensor_by_name('import/seq_lens:0')
test_sess = tf.InteractiveSession(graph = g)
result = test_sess.run([logits, seq_lens], feed_dict = {x: padded, x_lens: lens})
result
```
| github_jupyter |
# Introduction
Selecting specific values of a pandas DataFrame or Series to work on is an implicit step in almost any data operation you'll run, so one of the first things you need to learn in working with data in Python is how to go about selecting the data points relevant to you quickly and effectively.
```
import pandas as pd
reviews = pd.read_csv("../input/wine-reviews/winemag-data-130k-v2.csv", index_col=0)
pd.set_option('max_rows', 5)
```
**To start the exercise for this topic, please click [here](https://www.kaggle.com/kernels/fork/587910).**
# Native accessors
Native Python objects provide good ways of indexing data. Pandas carries all of these over, which helps make it easy to start with.
Consider this DataFrame:
```
reviews
```
In Python, we can access the property of an object by accessing it as an attribute. A `book` object, for example, might have a `title` property, which we can access by calling `book.title`. Columns in a pandas DataFrame work in much the same way.
Hence to access the `country` property of `reviews` we can use:
```
reviews.country
```
If we have a Python dictionary, we can access its values using the indexing (`[]`) operator. We can do the same with columns in a DataFrame:
```
reviews['country']
```
These are the two ways of selecting a specific Series out of a DataFrame. Neither of them is more or less syntactically valid than the other, but the indexing operator `[]` does have the advantage that it can handle column names with reserved characters in them (e.g. if we had a `country providence` column, `reviews.country providence` wouldn't work).
Doesn't a pandas Series look kind of like a fancy dictionary? It pretty much is, so it's no surprise that, to drill down to a single specific value, we need only use the indexing operator `[]` once more:
```
reviews['country'][0]
```
# Indexing in pandas
The indexing operator and attribute selection are nice because they work just like they do in the rest of the Python ecosystem. As a novice, this makes them easy to pick up and use. However, pandas has its own accessor operators, `loc` and `iloc`. For more advanced operations, these are the ones you're supposed to be using.
### Index-based selection
Pandas indexing works in one of two paradigms. The first is **index-based selection**: selecting data based on its numerical position in the data. `iloc` follows this paradigm.
To select the first row of data in a DataFrame, we may use the following:
```
reviews.iloc[0]
```
Both `loc` and `iloc` are row-first, column-second. This is the opposite of what we do in native Python, which is column-first, row-second.
This means that it's marginally easier to retrieve rows, and marginally harder to get retrieve columns. To get a column with `iloc`, we can do the following:
```
reviews.iloc[:, 0]
```
On its own, the `:` operator, which also comes from native Python, means "everything". When combined with other selectors, however, it can be used to indicate a range of values. For example, to select the `country` column from just the first, second, and third row, we would do:
```
reviews.iloc[:3, 0]
```
Or, to select just the second and third entries, we would do:
```
reviews.iloc[1:3, 0]
```
It's also possible to pass a list:
```
reviews.iloc[[0, 1, 2], 0]
```
Finally, it's worth knowing that negative numbers can be used in selection. This will start counting forwards from the _end_ of the values. So for example here are the last five elements of the dataset.
```
reviews.iloc[-5:]
```
### Label-based selection
The second paradigm for attribute selection is the one followed by the `loc` operator: **label-based selection**. In this paradigm, it's the data index value, not its position, which matters.
For example, to get the first entry in `reviews`, we would now do the following:
```
reviews.loc[0, 'country']
```
`iloc` is conceptually simpler than `loc` because it ignores the dataset's indices. When we use `iloc` we treat the dataset like a big matrix (a list of lists), one that we have to index into by position. `loc`, by contrast, uses the information in the indices to do its work. Since your dataset usually has meaningful indices, it's usually easier to do things using `loc` instead. For example, here's one operation that's much easier using `loc`:
```
reviews.loc[:, ['taster_name', 'taster_twitter_handle', 'points']]
```
### Choosing between `loc` and `iloc`
When choosing or transitioning between `loc` and `iloc`, there is one "gotcha" worth keeping in mind, which is that the two methods use slightly different indexing schemes.
`iloc` uses the Python stdlib indexing scheme, where the first element of the range is included and the last one excluded. So `0:10` will select entries `0,...,9`. `loc`, meanwhile, indexes inclusively. So `0:10` will select entries `0,...,10`.
Why the change? Remember that loc can index any stdlib type: strings, for example. If we have a DataFrame with index values `Apples, ..., Potatoes, ...`, and we want to select "all the alphabetical fruit choices between Apples and Potatoes", then it's a lot more convenient to index `df.loc['Apples':'Potatoes']` than it is to index something like `df.loc['Apples', 'Potatoet']` (`t` coming after `s` in the alphabet).
This is particularly confusing when the DataFrame index is a simple numerical list, e.g. `0,...,1000`. In this case `df.iloc[0:1000]` will return 1000 entries, while `df.loc[0:1000]` return 1001 of them! To get 1000 elements using `loc`, you will need to go one lower and ask for `df.loc[0:999]`.
Otherwise, the semantics of using `loc` are the same as those for `iloc`.
# Manipulating the index
Label-based selection derives its power from the labels in the index. Critically, the index we use is not immutable. We can manipulate the index in any way we see fit.
The `set_index()` method can be used to do the job. Here is what happens when we `set_index` to the `title` field:
```
reviews.set_index("title")
```
This is useful if you can come up with an index for the dataset which is better than the current one.
# Conditional selection
So far we've been indexing various strides of data, using structural properties of the DataFrame itself. To do *interesting* things with the data, however, we often need to ask questions based on conditions.
For example, suppose that we're interested specifically in better-than-average wines produced in Italy.
We can start by checking if each wine is Italian or not:
```
reviews.country == 'Italy'
```
This operation produced a Series of `True`/`False` booleans based on the `country` of each record. This result can then be used inside of `loc` to select the relevant data:
```
reviews.loc[reviews.country == 'Italy']
```
This DataFrame has ~20,000 rows. The original had ~130,000. That means that around 15% of wines originate from Italy.
We also wanted to know which ones are better than average. Wines are reviewed on a 80-to-100 point scale, so this could mean wines that accrued at least 90 points.
We can use the ampersand (`&`) to bring the two questions together:
```
reviews.loc[(reviews.country == 'Italy') & (reviews.points >= 90)]
```
Suppose we'll buy any wine that's made in Italy _or_ which is rated above average. For this we use a pipe (`|`):
```
reviews.loc[(reviews.country == 'Italy') | (reviews.points >= 90)]
```
Pandas comes with a few built-in conditional selectors, two of which we will highlight here.
The first is `isin`. `isin` is lets you select data whose value "is in" a list of values. For example, here's how we can use it to select wines only from Italy or France:
```
reviews.loc[reviews.country.isin(['Italy', 'France'])]
```
The second is `isnull` (and its companion `notnull`). These methods let you highlight values which are (or are not) empty (`NaN`). For example, to filter out wines lacking a price tag in the dataset, here's what we would do:
```
reviews.loc[reviews.price.notnull()]
```
# Assigning data
Going the other way, assigning data to a DataFrame is easy. You can assign either a constant value:
```
reviews['critic'] = 'everyone'
reviews['critic']
```
Or with an iterable of values:
```
reviews['index_backwards'] = range(len(reviews), 0, -1)
reviews['index_backwards']
```
# Your turn
If you haven't started the exercise, you can **[get started here](https://www.kaggle.com/kernels/fork/587910)**.
---
*Have questions or comments? Visit the [course discussion forum](https://www.kaggle.com/learn/pandas/discussion) to chat with other learners.*
| github_jupyter |
```
import os
from datetime import datetime
import pandas as pd
from zeep import Client
import loadenv
import db
from db import get_connection
# execute last cell for definitions first
client = Client("https://tools.rki.de/SurvStat/SurvStatWebService.svc?singleWsdl")
factory = client.type_factory("ns2")
state_ids = {
"Baden-Württemberg": "08",
"Bayern": "09",
"Berlin": "11",
"Brandenburg": "12",
"Bremen": "04",
"Hamburg": "02",
"Hessen": "06",
"Mecklenburg-Vorpommern": "13",
"Niedersachsen": "03",
"Nordrhein-Westfalen": "05",
"Rheinland-Pfalz": "07",
"Saarland": "10",
"Sachsen": "14",
"Sachsen-Anhalt": "15",
"Schleswig-Holstein": "01",
"Thüringen": "16",
}
counties = {
"LK Ahrweiler": {"State": "07", "Region": "DEB1", "County": "07131"},
"LK Aichach-Friedberg": {"State": "09", "Region": "DE27", "County": "09771"},
"LK Alb-Donau-Kreis": {"State": "08", "Region": "DE14", "County": "08425"},
"LK Altenburger Land": {"State": "16", "Region": "DEG0", "County": "16077"},
"LK Altenkirchen": {"State": "07", "Region": "DEB1", "County": "07132"},
"LK Altmarkkreis Salzwedel": {"State": "15", "Region": "DEE0", "County": "15081"},
"LK Altötting": {"State": "09", "Region": "DE21", "County": "09171"},
"LK Alzey-Worms": {"State": "07", "Region": "DEB3", "County": "07331"},
"LK Amberg-Sulzbach": {"State": "09", "Region": "DE23", "County": "09371"},
"LK Ammerland": {"State": "03", "Region": "DE94", "County": "03451"},
"LK Anhalt-Bitterfeld": {"State": "15", "Region": "DEE0", "County": "15082"},
"LK Ansbach": {"State": "09", "Region": "DE25", "County": "09571"},
"LK Aschaffenburg": {"State": "09", "Region": "DE26", "County": "09671"},
"LK Augsburg": {"State": "09", "Region": "DE27", "County": "09772"},
"LK Aurich": {"State": "03", "Region": "DE94", "County": "03452"},
"LK Bad Dürkheim": {"State": "07", "Region": "DEB3", "County": "07332"},
"LK Bad Kissingen": {"State": "09", "Region": "DE26", "County": "09672"},
"LK Bad Kreuznach": {"State": "07", "Region": "DEB1", "County": "07133"},
"LK Bad Tölz-Wolfratshausen": {"State": "09", "Region": "DE21", "County": "09173"},
"LK Bamberg": {"State": "09", "Region": "DE24", "County": "09471"},
"LK Barnim": {"State": "12", "Region": "DE40", "County": "12060"},
"LK Bautzen": {"State": "14", "Region": "DED2", "County": "14625"},
"LK Bayreuth": {"State": "09", "Region": "DE24", "County": "09472"},
"LK Berchtesgadener Land": {"State": "09", "Region": "DE21", "County": "09172"},
"LK Bergstraße": {"State": "06", "Region": "DE71", "County": "06431"},
"LK Bernkastel-Wittlich": {"State": "07", "Region": "DEB2", "County": "07231"},
"LK Biberach": {"State": "08", "Region": "DE14", "County": "08426"},
"LK Birkenfeld": {"State": "07", "Region": "DEB1", "County": "07134"},
"LK Bitburg-Prüm": {"State": "07", "Region": "DEB2", "County": "07232"},
"LK Bodenseekreis": {"State": "08", "Region": "DE14", "County": "08435"},
"LK Borken": {"State": "05", "Region": "DEA3", "County": "05554"},
"LK Breisgau-Hochschwarzwald": {"State": "08", "Region": "DE13", "County": "08315"},
"LK Burgenlandkreis": {"State": "15", "Region": "DEE0", "County": "15084"},
"LK Böblingen": {"State": "08", "Region": "DE11", "County": "08115"},
"LK Börde": {"State": "15", "Region": "DEE0", "County": "15083"},
"LK Calw": {"State": "08", "Region": "DE12", "County": "08235"},
"LK Celle": {"State": "03", "Region": "DE93", "County": "03351"},
"LK Cham": {"State": "09", "Region": "DE23", "County": "09372"},
"LK Cloppenburg": {"State": "03", "Region": "DE94", "County": "03453"},
"LK Coburg": {"State": "09", "Region": "DE24", "County": "09473"},
"LK Cochem-Zell": {"State": "07", "Region": "DEB1", "County": "07135"},
"LK Coesfeld": {"State": "05", "Region": "DEA3", "County": "05558"},
"LK Cuxhaven": {"State": "03", "Region": "DE93", "County": "03352"},
"LK Dachau": {"State": "09", "Region": "DE21", "County": "09174"},
"LK Dahme-Spreewald": {"State": "12", "Region": "DE40", "County": "12061"},
"LK Darmstadt-Dieburg": {"State": "06", "Region": "DE71", "County": "06432"},
"LK Deggendorf": {"State": "09", "Region": "DE22", "County": "09271"},
"LK Diepholz": {"State": "03", "Region": "DE92", "County": "03251"},
"LK Dillingen a.d.Donau": {"State": "09", "Region": "DE27", "County": "09773"},
"LK Dingolfing-Landau": {"State": "09", "Region": "DE22", "County": "09279"},
"LK Dithmarschen": {"State": "01", "Region": "DEF0", "County": "01051"},
"LK Donau-Ries": {"State": "09", "Region": "DE27", "County": "09779"},
"LK Donnersbergkreis": {"State": "07", "Region": "DEB3", "County": "07333"},
"LK Düren": {"State": "05", "Region": "DEA2", "County": "05358"},
"LK Ebersberg": {"State": "09", "Region": "DE21", "County": "09175"},
"LK Eichsfeld": {"State": "16", "Region": "DEG0", "County": "16061"},
"LK Eichstätt": {"State": "09", "Region": "DE21", "County": "09176"},
"LK Elbe-Elster": {"State": "12", "Region": "DE40", "County": "12062"},
"LK Emmendingen": {"State": "08", "Region": "DE13", "County": "08316"},
"LK Emsland": {"State": "03", "Region": "DE94", "County": "03454"},
"LK Ennepe-Ruhr-Kreis": {"State": "05", "Region": "DEA5", "County": "05954"},
"LK Enzkreis": {"State": "08", "Region": "DE12", "County": "08236"},
"LK Erding": {"State": "09", "Region": "DE21", "County": "09177"},
"LK Erlangen-Höchstadt": {"State": "09", "Region": "DE25", "County": "09572"},
"LK Erzgebirgskreis": {"State": "14", "Region": "DED4", "County": "14521"},
"LK Esslingen": {"State": "08", "Region": "DE11", "County": "08116"},
"LK Euskirchen": {"State": "05", "Region": "DEA2", "County": "05366"},
"LK Forchheim": {"State": "09", "Region": "DE24", "County": "09474"},
"LK Freising": {"State": "09", "Region": "DE21", "County": "09178"},
"LK Freudenstadt": {"State": "08", "Region": "DE12", "County": "08237"},
"LK Freyung-Grafenau": {"State": "09", "Region": "DE22", "County": "09272"},
"LK Friesland": {"State": "03", "Region": "DE94", "County": "03455"},
"LK Fulda": {"State": "06", "Region": "DE73", "County": "06631"},
"LK Fürstenfeldbruck": {"State": "09", "Region": "DE21", "County": "09179"},
"LK Fürth": {"State": "09", "Region": "DE25", "County": "09573"},
"LK Garmisch-Partenkirchen": {"State": "09", "Region": "DE21", "County": "09180"},
"LK Germersheim": {"State": "07", "Region": "DEB3", "County": "07334"},
"LK Gießen": {"State": "06", "Region": "DE72", "County": "06531"},
"LK Gifhorn": {"State": "03", "Region": "DE91", "County": "03151"},
"LK Goslar": {"State": "03", "Region": "DE91", "County": "03153"},
"LK Gotha": {"State": "16", "Region": "DEG0", "County": "16067"},
"LK Grafschaft Bentheim": {"State": "03", "Region": "DE94", "County": "03456"},
"LK Greiz": {"State": "16", "Region": "DEG0", "County": "16076"},
"LK Groß-Gerau": {"State": "06", "Region": "DE71", "County": "06433"},
"LK Göppingen": {"State": "08", "Region": "DE11", "County": "08117"},
"LK Görlitz": {"State": "14", "Region": "DED2", "County": "14626"},
"LK Göttingen": {"State": "03", "Region": "DE91", "County": "03159"},
"LK Günzburg": {"State": "09", "Region": "DE27", "County": "09774"},
"LK Gütersloh": {"State": "05", "Region": "DEA4", "County": "05754"},
"LK Hameln-Pyrmont": {"State": "03", "Region": "DE92", "County": "03252"},
"LK Harburg": {"State": "03", "Region": "DE93", "County": "03353"},
"LK Harz": {"State": "15", "Region": "DEE0", "County": "15085"},
"LK Havelland": {"State": "12", "Region": "DE40", "County": "12063"},
"LK Haßberge": {"State": "09", "Region": "DE26", "County": "09674"},
"LK Heidekreis": {"State": "03", "Region": "DE93", "County": "03358"},
"LK Heidenheim": {"State": "08", "Region": "DE11", "County": "08135"},
"LK Heilbronn": {"State": "08", "Region": "DE11", "County": "08125"},
"LK Heinsberg": {"State": "05", "Region": "DEA2", "County": "05370"},
"LK Helmstedt": {"State": "03", "Region": "DE91", "County": "03154"},
"LK Herford": {"State": "05", "Region": "DEA4", "County": "05758"},
"LK Hersfeld-Rotenburg": {"State": "06", "Region": "DE73", "County": "06632"},
"LK Herzogtum Lauenburg": {"State": "01", "Region": "DEF0", "County": "01053"},
"LK Hildburghausen": {"State": "16", "Region": "DEG0", "County": "16069"},
"LK Hildesheim": {"State": "03", "Region": "DE92", "County": "03254"},
"LK Hochsauerlandkreis": {"State": "05", "Region": "DEA5", "County": "05958"},
"LK Hochtaunuskreis": {"State": "06", "Region": "DE71", "County": "06434"},
"LK Hof": {"State": "09", "Region": "DE24", "County": "09475"},
"LK Hohenlohekreis": {"State": "08", "Region": "DE11", "County": "08126"},
"LK Holzminden": {"State": "03", "Region": "DE92", "County": "03255"},
"LK Höxter": {"State": "05", "Region": "DEA4", "County": "05762"},
"LK Ilm-Kreis": {"State": "16", "Region": "DEG0", "County": "16070"},
"LK Jerichower Land": {"State": "15", "Region": "DEE0", "County": "15086"},
"LK Kaiserslautern": {"State": "07", "Region": "DEB3", "County": "07335"},
"LK Karlsruhe": {"State": "08", "Region": "DE12", "County": "08215"},
"LK Kassel": {"State": "06", "Region": "DE73", "County": "06633"},
"LK Kelheim": {"State": "09", "Region": "DE22", "County": "09273"},
"LK Kitzingen": {"State": "09", "Region": "DE26", "County": "09675"},
"LK Kleve": {"State": "05", "Region": "DEA1", "County": "05154"},
"LK Konstanz": {"State": "08", "Region": "DE13", "County": "08335"},
"LK Kronach": {"State": "09", "Region": "DE24", "County": "09476"},
"LK Kulmbach": {"State": "09", "Region": "DE24", "County": "09477"},
"LK Kusel": {"State": "07", "Region": "DEB3", "County": "07336"},
"LK Kyffhäuserkreis": {"State": "16", "Region": "DEG0", "County": "16065"},
"LK Lahn-Dill-Kreis": {"State": "06", "Region": "DE72", "County": "06532"},
"LK Landsberg a.Lech": {"State": "09", "Region": "DE21", "County": "09181"},
"LK Landshut": {"State": "09", "Region": "DE22", "County": "09274"},
"LK Leer": {"State": "03", "Region": "DE94", "County": "03457"},
"LK Leipzig": {"State": "14", "Region": "DED5", "County": "14729"},
"LK Lichtenfels": {"State": "09", "Region": "DE24", "County": "09478"},
"LK Limburg-Weilburg": {"State": "06", "Region": "DE72", "County": "06533"},
"LK Lindau": {"State": "09", "Region": "DE27", "County": "09776"},
"LK Lippe": {"State": "05", "Region": "DEA4", "County": "05766"},
"LK Ludwigsburg": {"State": "08", "Region": "DE11", "County": "08118"},
"LK Ludwigslust-Parchim": {"State": "13", "Region": "DE80", "County": "13076"},
"LK Lörrach": {"State": "08", "Region": "DE13", "County": "08336"},
"LK Lüchow-Dannenberg": {"State": "03", "Region": "DE93", "County": "03354"},
"LK Lüneburg": {"State": "03", "Region": "DE93", "County": "03355"},
"LK Main-Kinzig-Kreis": {"State": "06", "Region": "DE71", "County": "06435"},
"LK Main-Spessart": {"State": "09", "Region": "DE26", "County": "09677"},
"LK Main-Tauber-Kreis": {"State": "08", "Region": "DE11", "County": "08128"},
"LK Main-Taunus-Kreis": {"State": "06", "Region": "DE71", "County": "06436"},
"LK Mainz-Bingen": {"State": "07", "Region": "DEB3", "County": "07339"},
"LK Mansfeld-Südharz": {"State": "15", "Region": "DEE0", "County": "15087"},
"LK Marburg-Biedenkopf": {"State": "06", "Region": "DE72", "County": "06534"},
"LK Mayen-Koblenz": {"State": "07", "Region": "DEB1", "County": "07137"},
"LK Mecklenburgische Seenplatte": {
"State": "13",
"Region": "DE80",
"County": "13071",
},
"LK Meißen": {"State": "14", "Region": "DED2", "County": "14627"},
"LK Merzig-Wadern": {"State": "10", "Region": "DEC0", "County": "10042"},
"LK Mettmann": {"State": "05", "Region": "DEA1", "County": "05158"},
"LK Miesbach": {"State": "09", "Region": "DE21", "County": "09182"},
"LK Miltenberg": {"State": "09", "Region": "DE26", "County": "09676"},
"LK Minden-Lübbecke": {"State": "05", "Region": "DEA4", "County": "05770"},
"LK Mittelsachsen": {"State": "14", "Region": "DED4", "County": "14522"},
"LK Märkisch-Oderland": {"State": "12", "Region": "DE40", "County": "12064"},
"LK Märkischer Kreis": {"State": "05", "Region": "DEA5", "County": "05962"},
"LK Mühldorf a.Inn": {"State": "09", "Region": "DE21", "County": "09183"},
"LK München": {"State": "09", "Region": "DE21", "County": "09184"},
"LK Neckar-Odenwald-Kreis": {"State": "08", "Region": "DE12", "County": "08225"},
"LK Neu-Ulm": {"State": "09", "Region": "DE27", "County": "09775"},
"LK Neuburg-Schrobenhausen": {"State": "09", "Region": "DE21", "County": "09185"},
"LK Neumarkt i.d.OPf.": {"State": "09", "Region": "DE23", "County": "09373"},
"LK Neunkirchen": {"State": "10", "Region": "DEC0", "County": "10043"},
"LK Neustadt a.d.Aisch-Bad Windsheim": {
"State": "09",
"Region": "DE25",
"County": "09575",
},
"LK Neustadt a.d.Waldnaab": {"State": "09", "Region": "DE23", "County": "09374"},
"LK Neuwied": {"State": "07", "Region": "DEB1", "County": "07138"},
"LK Nienburg (Weser)": {"State": "03", "Region": "DE92", "County": "03256"},
"LK Nordfriesland": {"State": "01", "Region": "DEF0", "County": "01054"},
"LK Nordhausen": {"State": "16", "Region": "DEG0", "County": "16062"},
"LK Nordsachsen": {"State": "14", "Region": "DED5", "County": "14730"},
"LK Nordwestmecklenburg": {"State": "13", "Region": "DE80", "County": "13074"},
"LK Northeim": {"State": "03", "Region": "DE91", "County": "03155"},
"LK Nürnberger Land": {"State": "09", "Region": "DE25", "County": "09574"},
"LK Oberallgäu": {"State": "09", "Region": "DE27", "County": "09780"},
"LK Oberbergischer Kreis": {"State": "05", "Region": "DEA2", "County": "05374"},
"LK Oberhavel": {"State": "12", "Region": "DE40", "County": "12065"},
"LK Oberspreewald-Lausitz": {"State": "12", "Region": "DE40", "County": "12066"},
"LK Odenwaldkreis": {"State": "06", "Region": "DE71", "County": "06437"},
"LK Oder-Spree": {"State": "12", "Region": "DE40", "County": "12067"},
"LK Offenbach": {"State": "06", "Region": "DE71", "County": "06438"},
"LK Oldenburg": {"State": "03", "Region": "DE94", "County": "03458"},
"LK Olpe": {"State": "05", "Region": "DEA5", "County": "05966"},
"LK Ortenaukreis": {"State": "08", "Region": "DE13", "County": "08317"},
"LK Osnabrück": {"State": "03", "Region": "DE94", "County": "03459"},
"LK Ostalbkreis": {"State": "08", "Region": "DE11", "County": "08136"},
"LK Ostallgäu": {"State": "09", "Region": "DE27", "County": "09777"},
"LK Osterholz": {"State": "03", "Region": "DE93", "County": "03356"},
"LK Ostholstein": {"State": "01", "Region": "DEF0", "County": "01055"},
"LK Ostprignitz-Ruppin": {"State": "12", "Region": "DE40", "County": "12068"},
"LK Paderborn": {"State": "05", "Region": "DEA4", "County": "05774"},
"LK Passau": {"State": "09", "Region": "DE22", "County": "09275"},
"LK Peine": {"State": "03", "Region": "DE91", "County": "03157"},
"LK Pfaffenhofen a.d.Ilm": {"State": "09", "Region": "DE21", "County": "09186"},
"LK Pinneberg": {"State": "01", "Region": "DEF0", "County": "01056"},
"LK Plön": {"State": "01", "Region": "DEF0", "County": "01057"},
"LK Potsdam-Mittelmark": {"State": "12", "Region": "DE40", "County": "12069"},
"LK Prignitz": {"State": "12", "Region": "DE40", "County": "12070"},
"LK Rastatt": {"State": "08", "Region": "DE12", "County": "08216"},
"LK Ravensburg": {"State": "08", "Region": "DE14", "County": "08436"},
"LK Recklinghausen": {"State": "05", "Region": "DEA3", "County": "05562"},
"LK Regen": {"State": "09", "Region": "DE22", "County": "09276"},
"LK Regensburg": {"State": "09", "Region": "DE23", "County": "09375"},
"LK Rems-Murr-Kreis": {"State": "08", "Region": "DE11", "County": "08119"},
"LK Rendsburg-Eckernförde": {"State": "01", "Region": "DEF0", "County": "01058"},
"LK Reutlingen": {"State": "08", "Region": "DE14", "County": "08415"},
"LK Rhein-Erft-Kreis": {"State": "05", "Region": "DEA2", "County": "05362"},
"LK Rhein-Hunsrück-Kreis": {"State": "07", "Region": "DEB1", "County": "07140"},
"LK Rhein-Kreis Neuss": {"State": "05", "Region": "DEA1", "County": "05162"},
"LK Rhein-Lahn-Kreis": {"State": "07", "Region": "DEB1", "County": "07141"},
"LK Rhein-Neckar-Kreis": {"State": "08", "Region": "DE12", "County": "08226"},
"LK Rhein-Pfalz-Kreis": {"State": "07", "Region": "DEB3", "County": "07338"},
"LK Rhein-Sieg-Kreis": {"State": "05", "Region": "DEA2", "County": "05382"},
"LK Rheingau-Taunus-Kreis": {"State": "06", "Region": "DE71", "County": "06439"},
"LK Rheinisch-Bergischer Kreis": {
"State": "05",
"Region": "DEA2",
"County": "05378",
},
"LK Rhön-Grabfeld": {"State": "09", "Region": "DE26", "County": "09673"},
"LK Rosenheim": {"State": "09", "Region": "DE21", "County": "09187"},
"LK Rostock": {"State": "13", "Region": "DE80", "County": "13072"},
"LK Rotenburg (Wümme)": {"State": "03", "Region": "DE93", "County": "03357"},
"LK Roth": {"State": "09", "Region": "DE25", "County": "09576"},
"LK Rottal-Inn": {"State": "09", "Region": "DE22", "County": "09277"},
"LK Rottweil": {"State": "08", "Region": "DE13", "County": "08325"},
"LK Saale-Holzland-Kreis": {"State": "16", "Region": "DEG0", "County": "16074"},
"LK Saale-Orla-Kreis": {"State": "16", "Region": "DEG0", "County": "16075"},
"LK Saalekreis": {"State": "15", "Region": "DEE0", "County": "15088"},
"LK Saalfeld-Rudolstadt": {"State": "16", "Region": "DEG0", "County": "16073"},
"LK Saar-Pfalz-Kreis": {"State": "10", "Region": "DEC0", "County": "10045"},
"LK Saarlouis": {"State": "10", "Region": "DEC0", "County": "10044"},
"LK Salzlandkreis": {"State": "15", "Region": "DEE0", "County": "15089"},
"LK Sankt Wendel": {"State": "10", "Region": "DEC0", "County": "10046"},
"LK Schaumburg": {"State": "03", "Region": "DE92", "County": "03257"},
"LK Schleswig-Flensburg": {"State": "01", "Region": "DEF0", "County": "01059"},
"LK Schmalkalden-Meiningen": {"State": "16", "Region": "DEG0", "County": "16066"},
"LK Schwalm-Eder-Kreis": {"State": "06", "Region": "DE73", "County": "06634"},
"LK Schwandorf": {"State": "09", "Region": "DE23", "County": "09376"},
"LK Schwarzwald-Baar-Kreis": {"State": "08", "Region": "DE13", "County": "08326"},
"LK Schweinfurt": {"State": "09", "Region": "DE26", "County": "09678"},
"LK Schwäbisch Hall": {"State": "08", "Region": "DE11", "County": "08127"},
"LK Segeberg": {"State": "01", "Region": "DEF0", "County": "01060"},
"LK Siegen-Wittgenstein": {"State": "05", "Region": "DEA5", "County": "05970"},
"LK Sigmaringen": {"State": "08", "Region": "DE14", "County": "08437"},
"LK Soest": {"State": "05", "Region": "DEA5", "County": "05974"},
"LK Sonneberg": {"State": "16", "Region": "DEG0", "County": "16072"},
"LK Spree-Neiße": {"State": "12", "Region": "DE40", "County": "12071"},
"LK Stade": {"State": "03", "Region": "DE93", "County": "03359"},
"LK Stadtverband Saarbrücken": {"State": "10", "Region": "DEC0", "County": "10041"},
"LK Starnberg": {"State": "09", "Region": "DE21", "County": "09188"},
"LK Steinburg": {"State": "01", "Region": "DEF0", "County": "01061"},
"LK Steinfurt": {"State": "05", "Region": "DEA3", "County": "05566"},
"LK Stendal": {"State": "15", "Region": "DEE0", "County": "15090"},
"LK Stormarn": {"State": "01", "Region": "DEF0", "County": "01062"},
"LK Straubing-Bogen": {"State": "09", "Region": "DE22", "County": "09278"},
"LK Sächsische Schweiz-Osterzgebirge": {
"State": "14",
"Region": "DED2",
"County": "14628",
},
"LK Sömmerda": {"State": "16", "Region": "DEG0", "County": "16068"},
"LK Südliche Weinstraße": {"State": "07", "Region": "DEB3", "County": "07337"},
"LK Südwestpfalz": {"State": "07", "Region": "DEB3", "County": "07340"},
"LK Teltow-Fläming": {"State": "12", "Region": "DE40", "County": "12072"},
"LK Tirschenreuth": {"State": "09", "Region": "DE23", "County": "09377"},
"LK Traunstein": {"State": "09", "Region": "DE21", "County": "09189"},
"LK Trier-Saarburg": {"State": "07", "Region": "DEB2", "County": "07235"},
"LK Tuttlingen": {"State": "08", "Region": "DE13", "County": "08327"},
"LK Tübingen": {"State": "08", "Region": "DE14", "County": "08416"},
"LK Uckermark": {"State": "12", "Region": "DE40", "County": "12073"},
"LK Uelzen": {"State": "03", "Region": "DE93", "County": "03360"},
"LK Unna": {"State": "05", "Region": "DEA5", "County": "05978"},
"LK Unstrut-Hainich-Kreis": {"State": "16", "Region": "DEG0", "County": "16064"},
"LK Unterallgäu": {"State": "09", "Region": "DE27", "County": "09778"},
"LK Vechta": {"State": "03", "Region": "DE94", "County": "03460"},
"LK Verden": {"State": "03", "Region": "DE93", "County": "03361"},
"LK Viersen": {"State": "05", "Region": "DEA1", "County": "05166"},
"LK Vogelsbergkreis": {"State": "06", "Region": "DE72", "County": "06535"},
"LK Vogtlandkreis": {"State": "14", "Region": "DED4", "County": "14523"},
"LK Vorpommern-Greifswald": {"State": "13", "Region": "DE80", "County": "13075"},
"LK Vorpommern-Rügen": {"State": "13", "Region": "DE80", "County": "13073"},
"LK Vulkaneifel": {"State": "07", "Region": "DEB2", "County": "07233"},
"LK Waldeck-Frankenberg": {"State": "06", "Region": "DE73", "County": "06635"},
"LK Waldshut": {"State": "08", "Region": "DE13", "County": "08337"},
"LK Warendorf": {"State": "05", "Region": "DEA3", "County": "05570"},
"LK Wartburgkreis": {"State": "16", "Region": "DEG0", "County": "16063"},
"LK Weilheim-Schongau": {"State": "09", "Region": "DE21", "County": "09190"},
"LK Weimarer Land": {"State": "16", "Region": "DEG0", "County": "16071"},
"LK Weißenburg-Gunzenhausen": {"State": "09", "Region": "DE25", "County": "09577"},
"LK Werra-Meißner-Kreis": {"State": "06", "Region": "DE73", "County": "06636"},
"LK Wesel": {"State": "05", "Region": "DEA1", "County": "05170"},
"LK Wesermarsch": {"State": "03", "Region": "DE94", "County": "03461"},
"LK Westerwaldkreis": {"State": "07", "Region": "DEB1", "County": "07143"},
"LK Wetteraukreis": {"State": "06", "Region": "DE71", "County": "06440"},
"LK Wittenberg": {"State": "15", "Region": "DEE0", "County": "15091"},
"LK Wittmund": {"State": "03", "Region": "DE94", "County": "03462"},
"LK Wolfenbüttel": {"State": "03", "Region": "DE91", "County": "03158"},
"LK Wunsiedel i.Fichtelgebirge": {
"State": "09",
"Region": "DE24",
"County": "09479",
},
"LK Würzburg": {"State": "09", "Region": "DE26", "County": "09679"},
"LK Zollernalbkreis": {"State": "08", "Region": "DE14", "County": "08417"},
"LK Zwickau": {"State": "14", "Region": "DED4", "County": "14524"},
"Region Hannover": {"State": "03", "Region": "DE92", "County": "03241"},
"SK Amberg": {"State": "09", "Region": "DE23", "County": "09361"},
"SK Ansbach": {"State": "09", "Region": "DE25", "County": "09561"},
"SK Aschaffenburg": {"State": "09", "Region": "DE26", "County": "09661"},
"SK Augsburg": {"State": "09", "Region": "DE27", "County": "09761"},
"SK Baden-Baden": {"State": "08", "Region": "DE12", "County": "08211"},
"SK Bamberg": {"State": "09", "Region": "DE24", "County": "09461"},
"SK Bayreuth": {"State": "09", "Region": "DE24", "County": "09462"},
"SK Berlin Charlottenburg-Wilmersdorf": {
"State": "11",
"Region": "DE30",
"County": "11004",
},
"SK Berlin Friedrichshain-Kreuzberg": {
"State": "11",
"Region": "DE30",
"County": "11002",
},
"SK Berlin Lichtenberg": {"State": "11", "Region": "DE30", "County": "11011"},
"SK Berlin Marzahn-Hellersdorf": {
"State": "11",
"Region": "DE30",
"County": "11010",
},
"SK Berlin Mitte": {"State": "11", "Region": "DE30", "County": "11001"},
"SK Berlin Neukölln": {"State": "11", "Region": "DE30", "County": "11008"},
"SK Berlin Pankow": {"State": "11", "Region": "DE30", "County": "11003"},
"SK Berlin Reinickendorf": {"State": "11", "Region": "DE30", "County": "11012"},
"SK Berlin Spandau": {"State": "11", "Region": "DE30", "County": "11005"},
"SK Berlin Steglitz-Zehlendorf": {
"State": "11",
"Region": "DE30",
"County": "11006",
},
"SK Berlin Tempelhof-Schöneberg": {
"State": "11",
"Region": "DE30",
"County": "11007",
},
"SK Berlin Treptow-Köpenick": {"State": "11", "Region": "DE30", "County": "11009"},
"SK Bielefeld": {"State": "05", "Region": "DEA4", "County": "05711"},
"SK Bochum": {"State": "05", "Region": "DEA5", "County": "05911"},
"SK Bonn": {"State": "05", "Region": "DEA2", "County": "05314"},
"SK Bottrop": {"State": "05", "Region": "DEA3", "County": "05512"},
"SK Brandenburg a.d.Havel": {"State": "12", "Region": "DE40", "County": "12051"},
"SK Braunschweig": {"State": "03", "Region": "DE91", "County": "03101"},
"SK Bremen": {"State": "04", "Region": "DE50", "County": "04011"},
"SK Bremerhaven": {"State": "04", "Region": "DE50", "County": "04012"},
"SK Chemnitz": {"State": "14", "Region": "DED4", "County": "14511"},
"SK Coburg": {"State": "09", "Region": "DE24", "County": "09463"},
"SK Cottbus": {"State": "12", "Region": "DE40", "County": "12052"},
"SK Darmstadt": {"State": "06", "Region": "DE71", "County": "06411"},
"SK Delmenhorst": {"State": "03", "Region": "DE94", "County": "03401"},
"SK Dessau-Roßlau": {"State": "15", "Region": "DEE0", "County": "15001"},
"SK Dortmund": {"State": "05", "Region": "DEA5", "County": "05913"},
"SK Dresden": {"State": "14", "Region": "DED2", "County": "14612"},
"SK Duisburg": {"State": "05", "Region": "DEA1", "County": "05112"},
"SK Düsseldorf": {"State": "05", "Region": "DEA1", "County": "05111"},
"SK Eisenach": {"State": "16", "Region": "DEG0", "County": "16056"},
"SK Emden": {"State": "03", "Region": "DE94", "County": "03402"},
"SK Erfurt": {"State": "16", "Region": "DEG0", "County": "16051"},
"SK Erlangen": {"State": "09", "Region": "DE25", "County": "09562"},
"SK Essen": {"State": "05", "Region": "DEA1", "County": "05113"},
"SK Flensburg": {"State": "01", "Region": "DEF0", "County": "01001"},
"SK Frankenthal": {"State": "07", "Region": "DEB3", "County": "07311"},
"SK Frankfurt (Oder)": {"State": "12", "Region": "DE40", "County": "12053"},
"SK Frankfurt am Main": {"State": "06", "Region": "DE71", "County": "06412"},
"SK Freiburg i.Breisgau": {"State": "08", "Region": "DE13", "County": "08311"},
"SK Fürth": {"State": "09", "Region": "DE25", "County": "09563"},
"SK Gelsenkirchen": {"State": "05", "Region": "DEA3", "County": "05513"},
"SK Gera": {"State": "16", "Region": "DEG0", "County": "16052"},
"SK Hagen": {"State": "05", "Region": "DEA5", "County": "05914"},
"SK Halle": {"State": "15", "Region": "DEE0", "County": "15002"},
"SK Hamburg": {"State": "02", "Region": "DE60", "County": "02000"},
"SK Hamm": {"State": "05", "Region": "DEA5", "County": "05915"},
"SK Heidelberg": {"State": "08", "Region": "DE12", "County": "08221"},
"SK Heilbronn": {"State": "08", "Region": "DE11", "County": "08121"},
"SK Herne": {"State": "05", "Region": "DEA5", "County": "05916"},
"SK Hof": {"State": "09", "Region": "DE24", "County": "09464"},
"SK Ingolstadt": {"State": "09", "Region": "DE21", "County": "09161"},
"SK Jena": {"State": "16", "Region": "DEG0", "County": "16053"},
"SK Kaiserslautern": {"State": "07", "Region": "DEB3", "County": "07312"},
"SK Karlsruhe": {"State": "08", "Region": "DE12", "County": "08212"},
"SK Kassel": {"State": "06", "Region": "DE73", "County": "06611"},
"SK Kaufbeuren": {"State": "09", "Region": "DE27", "County": "09762"},
"SK Kempten": {"State": "09", "Region": "DE27", "County": "09763"},
"SK Kiel": {"State": "01", "Region": "DEF0", "County": "01002"},
"SK Koblenz": {"State": "07", "Region": "DEB1", "County": "07111"},
"SK Krefeld": {"State": "05", "Region": "DEA1", "County": "05114"},
"SK Köln": {"State": "05", "Region": "DEA2", "County": "05315"},
"SK Landau i.d.Pfalz": {"State": "07", "Region": "DEB3", "County": "07313"},
"SK Landshut": {"State": "09", "Region": "DE22", "County": "09261"},
"SK Leipzig": {"State": "14", "Region": "DED5", "County": "14713"},
"SK Leverkusen": {"State": "05", "Region": "DEA2", "County": "05316"},
"SK Ludwigshafen": {"State": "07", "Region": "DEB3", "County": "07314"},
"SK Lübeck": {"State": "01", "Region": "DEF0", "County": "01003"},
"SK Magdeburg": {"State": "15", "Region": "DEE0", "County": "15003"},
"SK Mainz": {"State": "07", "Region": "DEB3", "County": "07315"},
"SK Mannheim": {"State": "08", "Region": "DE12", "County": "08222"},
"SK Memmingen": {"State": "09", "Region": "DE27", "County": "09764"},
"SK Mönchengladbach": {"State": "05", "Region": "DEA1", "County": "05116"},
"SK Mülheim a.d.Ruhr": {"State": "05", "Region": "DEA1", "County": "05117"},
"SK München": {"State": "09", "Region": "DE21", "County": "09162"},
"SK Münster": {"State": "05", "Region": "DEA3", "County": "05515"},
"SK Neumünster": {"State": "01", "Region": "DEF0", "County": "01004"},
"SK Neustadt a.d.Weinstraße": {"State": "07", "Region": "DEB3", "County": "07316"},
"SK Nürnberg": {"State": "09", "Region": "DE25", "County": "09564"},
"SK Oberhausen": {"State": "05", "Region": "DEA1", "County": "05119"},
"SK Offenbach": {"State": "06", "Region": "DE71", "County": "06413"},
"SK Oldenburg": {"State": "03", "Region": "DE94", "County": "03403"},
"SK Osnabrück": {"State": "03", "Region": "DE94", "County": "03404"},
"SK Passau": {"State": "09", "Region": "DE22", "County": "09262"},
"SK Pforzheim": {"State": "08", "Region": "DE12", "County": "08231"},
"SK Pirmasens": {"State": "07", "Region": "DEB3", "County": "07317"},
"SK Potsdam": {"State": "12", "Region": "DE40", "County": "12054"},
"SK Regensburg": {"State": "09", "Region": "DE23", "County": "09362"},
"SK Remscheid": {"State": "05", "Region": "DEA1", "County": "05120"},
"SK Rosenheim": {"State": "09", "Region": "DE21", "County": "09163"},
"SK Rostock": {"State": "13", "Region": "DE80", "County": "13003"},
"SK Salzgitter": {"State": "03", "Region": "DE91", "County": "03102"},
"SK Schwabach": {"State": "09", "Region": "DE25", "County": "09565"},
"SK Schweinfurt": {"State": "09", "Region": "DE26", "County": "09662"},
"SK Schwerin": {"State": "13", "Region": "DE80", "County": "13004"},
"SK Solingen": {"State": "05", "Region": "DEA1", "County": "05122"},
"SK Speyer": {"State": "07", "Region": "DEB3", "County": "07318"},
"SK Straubing": {"State": "09", "Region": "DE22", "County": "09263"},
"SK Stuttgart": {"State": "08", "Region": "DE11", "County": "08111"},
"SK Suhl": {"State": "16", "Region": "DEG0", "County": "16054"},
"SK Trier": {"State": "07", "Region": "DEB2", "County": "07211"},
"SK Ulm": {"State": "08", "Region": "DE14", "County": "08421"},
"SK Weiden i.d.OPf.": {"State": "09", "Region": "DE23", "County": "09363"},
"SK Weimar": {"State": "16", "Region": "DEG0", "County": "16055"},
"SK Wiesbaden": {"State": "06", "Region": "DE71", "County": "06414"},
"SK Wilhelmshaven": {"State": "03", "Region": "DE94", "County": "03405"},
"SK Wolfsburg": {"State": "03", "Region": "DE91", "County": "03103"},
"SK Worms": {"State": "07", "Region": "DEB3", "County": "07319"},
"SK Wuppertal": {"State": "05", "Region": "DEA1", "County": "05124"},
"SK Würzburg": {"State": "09", "Region": "DE26", "County": "09663"},
"SK Zweibrücken": {"State": "07", "Region": "DEB3", "County": "07320"},
"StadtRegion Aachen": {"State": "05", "Region": "DEA2", "County": "05334"},
"Unbekannt": {"State": "-1", "Region": "????", "County": "?????"},
}
# os.makedirs('data-rki')
def fetch_county(county, incidence=False):
measures = {"Count": 0}
if incidence is True:
measures = {"Incidence": 1}
res = client.service.GetOlapData(
{
"Language": "German",
"Measures": measures,
"Cube": "SurvStat",
# Totals still included, setting `true` yields duplicates
"IncludeTotalColumn": False,
"IncludeTotalRow": False,
"IncludeNullRows": False,
"IncludeNullColumns": True,
"HierarchyFilters": factory.FilterCollection(
[
{
"Key": {
"DimensionId": "[PathogenOut].[KategorieNz]",
"HierarchyId": "[PathogenOut].[KategorieNz].[Krankheit DE]",
},
"Value": factory.FilterMemberCollection(
["[PathogenOut].[KategorieNz].[Krankheit DE].&[COVID-19]"]
),
},
{
"Key": {
"DimensionId": "[ReferenzDefinition]",
"HierarchyId": "[ReferenzDefinition].[ID]",
},
"Value": factory.FilterMemberCollection(
["[ReferenzDefinition].[ID].&[1]"]
),
},
{
"Key": {
"DimensionId": "[DeutschlandNodes].[Kreise71Web]",
"HierarchyId": "[DeutschlandNodes].[Kreise71Web].[FedStateKey71]",
},
"Value": factory.FilterMemberCollection(
[
f"[DeutschlandNodes].[Kreise71Web].[FedStateKey71].&[{ counties[county]['State'] }].&[{ counties[county]['Region'] }].&[{ counties[county]['County'] }]"
]
),
},
]
),
"RowHierarchy": "[ReportingDate].[YearWeek]",
"ColumnHierarchy": "[AlterPerson80].[AgeGroupName8]"
}
)
# TODO Wolfgang: Set the correct parameters above
# likely
# [ReportingDate].[WeekYear]
# [ReportingDate].[Season27 Year Week]
columns = [i["Caption"] for i in res.Columns.QueryResultColumn]
df = pd.DataFrame(
[
[i["Caption"]]
+ [float(c.replace(".", "").replace(",", ".")) if c is not None else None for c in i["Values"]["string"]]
for i in res.QueryResults.QueryResultRow
],
columns=["KW"] + columns,
)
df = df.set_index("KW")
df = df.astype("Float64") # Allow for None values
df = df.drop("Gesamt")
df = df.drop("Gesamt", axis=1)
return df
```
Test one LK
```
counties["LK Konstanz"]
df = fetch_county("LK Konstanz")
df2 = fetch_county("LK Konstanz", True)
```
All LKs
```
df_count = df['A80+'].nlargest(3)
df_count
df_incidence = df2['A80+'].nlargest(3)
df_incidence
```
Definitions
```
def estimate_pop(v_count, v_incidence):
res = []
for i in range(0, min(len(v_count), len(v_incidence))):
if v_count[i] is not None and v_incidence[i] is not None:
res.append((v_count[i] * 100000) / v_incidence[i])
return round(sum(res) / float(len(res)))
estimate_pop(df_count.to_list(), df_incidence.to_list())
conn, cur = get_connection()
def process_county(county):
df = fetch_county("LK Konstanz")
df2 = fetch_county("LK Konstanz", True)
for colName, col in df.iteritems():
if colName == "Unbekannt":
continue
v_count = df[colName].nlargest(3).to_list()
v_incidence = df2[colName].nlargest(3).to_list()
pop = estimate_pop(v_count, v_incidence)
age = int(colName[1:3])
ags = counties[county]['County']
if age == 0:
cur.execute(f"INSERT INTO population_survstat_agegroup2 (ags, a00) VALUES ('{'{:05d}'.format(int(ags))}', {pop})")
elif age > 0 and age < 80:
cur.execute(f"UPDATE population_survstat_agegroup2 SET {'a{:02d}'.format(int(age))} = {pop} WHERE ags = '{'{:05d}'.format(int(ags))}'")
else:
cur.execute(f"UPDATE population_survstat_agegroup2 SET \"A80+\" = {pop} WHERE ags = '{'{:05d}'.format(int(ags))}'")
conn.commit()
return pop
import logging
logging.getLogger("zeep").setLevel(logging.INFO)
logging.getLogger("urllib3").setLevel(logging.INFO)
i = 0
total = len(counties.keys())
for c in counties.keys():
process_county(c)
i += 1
print(f"{i}/{total}")
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import scipy
from datetime import datetime, timedelta
import sys
sys.path.append('/Users/tarabaris/GitHub/odp-sdk-python/Examples')
## For SDK
from getpass import getpass
from odp_sdk import ODPClient
from UtilityFunctions import *
## For plotting
import seaborn as sns
import matplotlib.pyplot as plt
import mpl_toolkits
import cartopy.crs as ccrs
from cartopy.mpl.ticker import LongitudeFormatter, LatitudeFormatter
import cartopy.feature as cfeature
import cmocean
from matplotlib.colors import BoundaryNorm
from matplotlib.ticker import MaxNLocator
from mpl_toolkits.axes_grid1 import make_axes_locatable
from cartopy.mpl.gridliner import LONGITUDE_FORMATTER, LATITUDE_FORMATTER
from matplotlib.lines import Line2D
## For geopandas
import geopandas as gpd
from shapely.geometry import Polygon
from shapely.geometry import MultiPolygon
## Extra functions
from tqdm import notebook
import warnings
warnings.filterwarnings("ignore")
pd.set_option('display.max_columns', None)
pd.set_option('display.max_rows', None)
sns.set_palette(sns.color_palette("hls", 47))
```
# Connect to API
```
client = ODPClient(api_key=getpass(prompt='Insert your personal ODP API key:'),
project="odp", client_name="odp")
```
Run function you will need to fetch polygons of different regions
```
def get_poly(df):
polys=[]
for i in notebook.tqdm(range(len(df))):
if df['poly_count'][i]>1:
m_polys=[]
for i in range(df['poly_count'][i]):
df_p = client.sequences.data.retrieve(id=df['id'][i], start=1, end=None).to_pandas()
m_polys.append(Polygon(zip(df_p[df_p.polygonId==i]['lat'], df_p[df_p.polygonId==i]['lon'])))
polys.append(MultiPolygon(m_polys))
else:
polys.append(Polygon(zip(client.sequences.data.retrieve(id=df['id'][i], start=1, end=None).to_pandas()['lat'], client.sequences.data.retrieve(id=df['id'][i], start=1, end=None).to_pandas()['lon'])))
df['geometry']=polys
```
# Get casts and data from Antarctic EEZ
First get all sequences of Marine Regions and EEZs that belong to Antarctica
```
seqs = client.sequences.list(external_id_prefix = 'marine-regions-Intersect_EEZ_IHO', metadata={'SOVEREIGN1': 'ANTARCTICA'}, limit=-1).to_pandas()
seqs.head()
```
Create a dataframe with marine regions, eezs, and associated polygons
```
df_ant = pd.DataFrame({'marine_regions':[seqs.iloc[i]['metadata']['MARREGION'] for i in range(len(seqs))],
'MRGID':[seqs.iloc[i]['metadata']['MRGID'] for i in range(len(seqs))],
'IHO_Sea':[seqs.iloc[i]['metadata']['IHO_SEA'] for i in range(len(seqs))],
'EEZ':[seqs.iloc[i]['metadata']['EEZ'] for i in range(len(seqs))],
'SOVEREIGN1':[seqs.iloc[i]['metadata']['SOVEREIGN1'] for i in range(len(seqs))],
'id': seqs['id'],
'poly_count':[int(seqs.iloc[i]['metadata']['polygonCount']) for i in range(len(seqs))]})
get_poly(df_ant)
df_ant = gpd.GeoDataFrame(df_ant)
df_ant.head()
```
Pull all casts from 2018 and find the ones that intersect with the Antarctic EEZ Polygons
```
## Download casts from 2018 and turn into a geopandas dataframe
casts2018 = client.get_available_casts([-180, 180], [-90, 90], ['2018-01-01', '2018-12-31'], n_threads=35)
casts2018 = gpd.GeoDataFrame(
casts2018, geometry=gpd.points_from_xy(casts2018.lon, casts2018.lat))
casts2018.head()
## Find intersect of casts and Antarctic EEZ
casts2018_ant = gpd.sjoin(casts2018, df_ant, how="inner", op='intersects')
casts2018_ant.head(2)
```
You can then fetch the measurement data from each of these casts
```
ant_data = client.download_data_from_casts(casts2018_ant.extId.unique(), n_threads=40)
ant_data.head()
```
And you can plot the measurements for Temperature in the Antarctic EEZ
```
plot_casts('Temperature', ant_data, cmap=cmocean.cm.thermal)
```
# Assign World Seas Marine Regions to casts
```
seqs = client.sequences.list(external_id_prefix = 'marine-regions-Intersect_EEZ_IHO', limit=-1).to_pandas()
seqs.head()
```
Create dataframe with name of marine region and polygons
```
df_mr = pd.DataFrame({'marine_regions':[seqs.iloc[i]['metadata']['MARREGION'] for i in range(len(seqs))],
'iho_sea':[seqs.iloc[i]['metadata']['IHO_SEA'] for i in range(len(seqs))],
'MRGID':[seqs.iloc[i]['metadata']['MRGID'] for i in range(len(seqs))],
'poly_count':[int(seqs.iloc[i]['metadata']['polygonCount']) for i in range(len(seqs))],
'id':seqs['id']})
get_poly(df_mr)
df_mr = gpd.GeoDataFrame(df_mr)
df_mr.head()
```
# Join oceanographic data to Marine Regions
Here we find the intersect of points from the casts with the polygons from the marine regions.
The resulting dataframe will have the associated marine regions for each cast location.
```
casts2018_mr = gpd.sjoin(casts2018, df_mr, how="inner", op='intersects')
casts2018_mr.head(2)
def plot_marine_regions_data(df_casts, df_marine_regions, lat=[-90, 90], lon=[-180,180]):
df_casts = df_casts[(df_casts.lat.between(lat[0], lat[1])) & (df_casts.lon.between(lon[0], lon[1]))]
fig = plt.figure(figsize=(14, 14))
colors = sns.color_palette('hls', n_colors=len(df_casts.marine_regions.unique()))
ax = fig.add_subplot(1, 1, 1, projection=ccrs.PlateCarree())
mr = df_casts.marine_regions.unique()
legend_elements = []
for i, j in enumerate(mr):
df_marine_regions[df_marine_regions.marine_regions == j]['geometry'].plot(ax=ax, markersize=5, color=colors[i], zorder=1);
legend_elements.append(Line2D([0], [0], color = colors[i], lw=4, label=mr[i]))
sns.scatterplot(x="lon", y="lat", data=df_casts, color = 'navy', s=20, marker='o', edgecolor='white', linewidths=0.05)
ax.set_extent([lon[0], lon[1], lat[0], lat[1]],crs=ccrs.PlateCarree())
gl = ax.gridlines(crs=ccrs.PlateCarree(), draw_labels=True, linewidth=1, color='gray', alpha=0.7, linestyle=':')
gl.xformatter = LONGITUDE_FORMATTER
gl.yformatter = LATITUDE_FORMATTER
ax.legend(handles=legend_elements, loc='lower center',
ncol=3, borderaxespad=-12)
geo_map(ax)
plot_marine_regions_data(casts2018_mr, df_mr, lat=[-5,15], lon=[-100,-75])
```
| github_jupyter |
# Lindstedt Poincare Example - Rayleigh's Equation
For the Lindstedt-Poincare method we introduce a dimensionless time $\tau$
$$
\begin{align*}
\tau &= \omega t \\
\dfrac{d}{dt} &= \omega \dfrac{d}{d\tau}\\
\dfrac{d^2}{dt^2} &= \omega^2 \dfrac{^2}{d\tau^2}
\end{align*}
$$
We will use Rayleigh's Equation as an example
$$
\begin{gather*}
\ddot{x} + \epsilon \left( \dfrac{1}{3}\dot{x}^3 - \dot{x} \right) + x = 0
\end{gather*}
$$
with initial conditions of
$$
\begin{gather*}
x(0) = a\\
\dot{x}(0) = 0
\end{gather*}
$$
and the system will be investigated up to only the second order.
```
import sympy as sp
from sympy.simplify.fu import TR0, TR5, TR7, TR8, TR11
from math import factorial
N = 3 # first order
# Define the symbolic parameters
epsilon = sp.symbols('epsilon')
omega = sp.symbols('omega')
omega_i = sp.symbols('omega_(0:' + str(N) + ')')
tau = sp.symbols('tau')
x = sp.Function('x')(tau)
xdot = sp.Derivative(x, tau)
xddot = sp.Derivative(xdot, tau)
x0 = sp.Function('x_0')(tau)
x1 = sp.Function('x_1')(tau)
x2 = sp.Function('x_2')(tau)
x3 = sp.Function('x_3')(tau)
# EOM
EOM = omega**2 * xddot + x + epsilon * (omega**3 * xdot**3/3 - omega * xdot)
EOM
x_i = (x0, x1, x2, x3)
x_e = sum([epsilon**i * x_i[i] for i in range(N+1)])
x_e
_omega = sum([epsilon**i * omega_i[i] for i in range(N)])
_omega
# Substitute these into the EOM
EOM = EOM.subs([
(x, x_e), (omega, _omega)
])
EOM
EOM = sp.expand(EOM).doit()
EOM = sp.expand(EOM)
EOM
# Collect the coefficients for the epsilons
epsilon_Eq = sp.collect(EOM, epsilon, evaluate=False)
epsilon_0_Eq = sp.Eq(epsilon_Eq[epsilon**0], 0)
epsilon_0_Eq
epsilon_1_Eq = sp.Eq(epsilon_Eq[epsilon], 0)
epsilon_1_Eq
epsilon_2_Eq = sp.Eq(epsilon_Eq[epsilon**2], 0)
epsilon_2_Eq
epsilon_3_Eq = sp.Eq(epsilon_Eq[epsilon**3], 0)
epsilon_3_Eq
sp.dsolve(epsilon_0_Eq, x0)
```
Not as convenient as working with the polar form
```
a = sp.symbols('a')
beta = sp.symbols('beta')
phi = tau + beta
x0_polar = a * sp.cos(tau + beta)
x0_polar
```
Update $\epsilon^1$ equation
```
epsilon_1_Eq = sp.expand(epsilon_1_Eq.subs(x0, x0_polar)).doit()
epsilon_1_Eq = sp.expand(TR8(sp.expand(TR7(TR5((epsilon_1_Eq))))))
epsilon_1_Eq
```
In the equation above we can see that the $\cos\phi$ terms are secular which causes the approximation fail due to resonance. Therefore, we must set those terms to go to zero.
$\omega_1$ = 0
$a = \pm \dfrac{2}{\omega_0}$
and let us pick $a = 2/\omega_0$
```
omega_1_new = 0
epsilon_1_Eq = epsilon_1_Eq.subs([(omega_i[1], omega_1_new), (a, 2/omega_i[0])])
epsilon_1_Eq = sp.simplify(epsilon_1_Eq)
epsilon_1_Eq
temp = sp.dsolve(epsilon_1_Eq)
temp
```
We only care about the particular solution
```
x1_p = 2 * sp.sin(3 * beta + 3 * tau) / 3 / (9 * omega_i[0]**2 - 1)
x1_p
epsilon_2_Eq = epsilon_2_Eq.subs([
(omega_i[1], omega_1_new), (x0, x0_polar), (x1, x1_p)
]).doit()
epsilon_2_Eq = sp.expand(TR8(sp.expand(TR7(TR5((epsilon_2_Eq))))))
epsilon_2_Eq
```
Remove the secular term by setting
$$
\omega_2 = -\dfrac{a\omega_0^2}{36\omega_0^2 - 4}
$$
```
omega_2_new = -a*omega_i[0]**2 / (36*omega_i[0]**2 - 4)
epsilon_2_Eq = epsilon_2_Eq.subs([(omega_i[2], omega_2_new), (a, 2/omega_i[0])])
epsilon_2_Eq = sp.simplify(epsilon_2_Eq)
epsilon_2_Eq
temp = sp.dsolve(epsilon_2_Eq)
temp
x2_p = (-2 * omega_i[0] * sp.cos(5*beta + 5*tau) / (225 * omega_i[0]**4 - 34*omega_i[0]**2 + 1)
+ 2 *omega_i[0]*sp.cos(3*beta + 3*tau) / (81*omega_i[0]**4 - 18*omega_i[0]**2 + 1))
x2_p
omega = omega_i[0] + epsilon * omega_1_new + epsilon**2 * omega_2_new
omega
t = sp.symbols('t')
x_new = epsilon * a * sp.cos(omega * t + beta)
x_new
```
We can find $x_3(t)$ after this, but we do not have to. This is because we will "reconstitute" the solution to $O(\epsilon^3)$
$$
\begin{align*}
\begin{cases}
x &= x_0 + \epsilon x_1 + \epsilon^2 x_2 + O(\epsilon^3)\\
\omega &= \omega_0 + \epsilon \omega_1 + \epsilon^2 \omega_2 + O(\epsilon^3)\\
\tau &= \omega t
\end{cases}
\end{align*}
$$
Establish the expansion for $x$ (found during the Lindstedt-Poincare analysis)
```
x0_new = a * sp.cos(omega * t + beta)
x1_new = x1_p.subs(tau, omega * t )
x2_new = x2_p.subs(tau, omega * t)
x_final = x0_new + epsilon * x1_new + epsilon ** 2 * x2_new
x_final
# From the initial conditions we know that omega0 = 1, a = 2, and beta = 0
x_final = x_final.subs([
(omega_i[0], 1), (a, 2), (beta, 0)])
x_final
omega.subs([(a,2), (omega_i[0],1)])
import numpy as np
import matplotlib.pyplot as plt
from scipy.integrate import solve_ivp, DOP853
from typing import List
# Generate phase plane of Rayleigh's equation to confirm limit cycle
def rayleigh(t, x, e):
return [x[1], -x[0] - e*(x[1]**3 / 3 - x[1])]
def solve_diffeq(func, t, tspan, ic, parameters={}, algorithm='DOP853', stepsize=np.inf):
return solve_ivp(fun=func, t_span=tspan, t_eval=t, y0=ic, method=algorithm,
args=tuple(parameters.values()), atol=1e-8, rtol=1e-5, max_step=stepsize)
def phasePlane(x1, x2, func, params):
X1, X2 = np.meshgrid(x1, x2) # create grid
u, v = np.zeros(X1.shape), np.zeros(X2.shape)
NI, NJ = X1.shape
for i in range(NI):
for j in range(NJ):
x = X1[i, j]
y = X2[i, j]
dx = func(0, (x, y), *params.values()) # compute values on grid
u[i, j] = dx[0]
v[i, j] = dx[1]
M = np.hypot(u, v)
u /= M
v /= M
return X1, X2, u, v, M
def DEplot(sys: object, tspan: tuple, x0: List[List[float]],
x: np.ndarray, y: np.ndarray, params: dict):
if len(tspan) != 3:
raise Exception('tspan should be tuple of size 3: (min, max, number of points).')
# Set up the figure the way we want it to look
plt.figure(figsize=(12, 9))
X1, X2, dx1, dx2, M = phasePlane(
x, y, sys, params
)
# Quiver plot
plt.quiver(X1, X2, dx1, dx2, M, scale=None, pivot='mid')
plt.grid()
if tspan[0] < 0:
t1 = np.linspace(0, tspan[0], tspan[2])
t2 = np.linspace(0, tspan[1], tspan[2])
if min(tspan) < 0:
t_span1 = (np.max(t1), np.min(t1))
else:
t_span1 = (np.min(t1), np.max(t1))
t_span2 = (np.min(t2), np.max(t2))
for x0i in x0:
sol1 = solve_diffeq(sys, t1, t_span1, x0i, params)
plt.plot(sol1.y[0, :], sol1.y[1, :], '-r')
sol2 = solve_diffeq(sys, t2, t_span2, x0i, params)
plt.plot(sol2.y[0, :], sol2.y[1, :], '-r')
else:
t = np.linspace(tspan[0], tspan[1], tspan[2])
t_span = (np.min(t), np.max(t))
for x0i in x0:
sol = solve_diffeq(sys, t, t_span, x0i, params)
plt.plot(sol.y[0, :], sol.y[1, :], '-r')
plt.xlim([np.min(x), np.max(x)])
plt.ylim([np.min(y), np.max(y)])
plt.show()
x10 = np.arange(0, 10, 1)
x20 = np.arange(0, 10, 1)
x0 = np.stack((x10, x20), axis=-1)
p = {'e': 0.01}
x1 = np.linspace(-5, 5, 20)
x2 = np.linspace(-5, 5, 20)
DEplot(rayleigh, (-8, 8, 1000), x0, x1, x2, p)
# Compare the approximation to the actual solution
# let a = 2
tmax = 2
tmin = 30
tspan = np.linspace(tmin, tmax, 1000)
# ODE solver solution
sol = solve_diffeq(rayleigh, tspan, (tmin, tmax), [2, 0], p)
# Approximation
def rayleigh_LP2(t, e):
omega = 1 - e**2/16
A = 2 * np.cos(t * omega)
B = e * np.sin(3 * t * omega)/12
C = e**2 * np.cos(3 * t * omega)/32
D = -e**2 * np.cos(5 * t * omega) / 96
return A + B + C + D
approx = rayleigh_LP2(tspan, 0.01)
plt.figure(figsize=(12, 9))
plt.plot(tspan, sol.y[0, :])
plt.plot(tspan, approx)
plt.grid(True)
plt.xlabel('$t$')
plt.ylabel('$x$')
plt.show()
```
### Better than first order Multiple Scale method
| github_jupyter |
```
from google.colab import drive
drive.mount('/content/gdrive')
import os
os.chdir('/content/gdrive/My Drive/finch/tensorflow2/text_matching/snli/main')
%tensorflow_version 2.x
!pip install tensorflow-addons
!pip install transformers
from transformers import BertTokenizer, TFBertModel
import tensorflow as tf
import tensorflow_addons as tfa
import numpy as np
import pprint
import logging
import time
print("TensorFlow Version", tf.__version__)
print('GPU Enabled:', tf.test.is_gpu_available())
params = {
'train_path': '../data/train.txt',
'test_path': '../data/test.txt',
'pretrain_path': 'bert-base-uncased',
'num_samples': 550152,
'buffer_size': 200000,
'batch_size': 32,
'max_len': 128 + 3,
'num_patience': 5,
'init_lr': 1e-5,
'max_lr': 3e-5,
}
tokenizer = BertTokenizer.from_pretrained(params['pretrain_path'],
lowercase = True,
add_special_tokens = True)
# stream data from text files
def data_generator(f_path, params):
label2idx = {'neutral': 0, 'entailment': 1, 'contradiction': 2,}
with open(f_path) as f:
print('Reading', f_path)
for line in f:
line = line.rstrip()
label, text1, text2 = line.split('\t')
if label == '-':
continue
text1 = tokenizer.tokenize(text1)
text2 = tokenizer.tokenize(text2)
if len(text1) + len(text2) + 3 > params['max_len']:
_max_len = (params['max_len'] - 3) // 2
text1 = text1[:_max_len]
text2 = text2[:_max_len]
text = ['[CLS]'] + text1 + ['[SEP]'] + text2 + ['[SEP]']
text = tokenizer.convert_tokens_to_ids(text)
seg = [0] + [0] * len(text1) + [0] + [1] * len(text2) + [1]
yield text, seg, label2idx[label]
def dataset(is_training, params):
_shapes = ([None], [None], ())
_types = (tf.int32, tf.int32, tf.int32)
_pads = (0, 0, -1)
if is_training:
ds = tf.data.Dataset.from_generator(
lambda: data_generator(params['train_path'], params),
output_shapes = _shapes,
output_types = _types,)
ds = ds.shuffle(params['buffer_size'])
ds = ds.padded_batch(params['batch_size'], _shapes, _pads)
ds = ds.prefetch(tf.data.experimental.AUTOTUNE)
else:
ds = tf.data.Dataset.from_generator(
lambda: data_generator(params['test_path'], params),
output_shapes = _shapes,
output_types = _types,)
ds = ds.padded_batch(params['batch_size'], _shapes, _pads)
ds = ds.prefetch(tf.data.experimental.AUTOTUNE)
return ds
# input stream ids check
text, seg, _ = next(data_generator(params['train_path'], params))
print(text)
print(seg)
class BertFinetune(tf.keras.Model):
def __init__(self, params):
super(BertFinetune, self).__init__()
self.bert = TFBertModel.from_pretrained(params['pretrain_path'],
trainable = True)
self.drop_1 = tf.keras.layers.Dropout(.1)
self.fc = tf.keras.layers.Dense(300, tf.nn.swish, name='down_stream/fc')
self.drop_2 = tf.keras.layers.Dropout(.1)
self.out = tf.keras.layers.Dense(3, name='down_stream/out')
def call(self, bert_inputs, training):
bert_inputs = [tf.cast(inp, tf.int32) for inp in bert_inputs]
x = self.bert(bert_inputs, training=training)[1]
x = self.drop_1(x, training=training)
x = self.fc(x)
x = self.drop_2(x, training=training)
x = self.out(x)
return x
model = BertFinetune(params)
model.build([[None, None], [None, None], [None, None]])
pprint.pprint([(v.name, v.shape) for v in model.trainable_variables])
step_size = 2 * params['num_samples'] // params['batch_size']
decay_lr = tfa.optimizers.Triangular2CyclicalLearningRate(
initial_learning_rate = params['init_lr'],
maximal_learning_rate = params['max_lr'],
step_size = step_size,)
optim = tf.optimizers.Adam(params['init_lr'])
global_step = 0
best_acc = .0
count = 0
t0 = time.time()
logger = logging.getLogger('tensorflow')
logger.setLevel(logging.INFO)
while True:
# TRAINING
for (text, seg, labels) in dataset(is_training=True, params=params):
with tf.GradientTape() as tape:
logits = model([text, tf.sign(text), seg], training=True)
loss = tf.compat.v1.losses.softmax_cross_entropy(
tf.one_hot(labels, 3, dtype=tf.float32),
logits = logits,
label_smoothing = .2,)
optim.lr.assign(decay_lr(global_step))
grads = tape.gradient(loss, model.trainable_variables)
grads, _ = tf.clip_by_global_norm(grads, 5.)
optim.apply_gradients(zip(grads, model.trainable_variables))
if global_step % 100 == 0:
logger.info("Step {} | Loss: {:.4f} | Spent: {:.1f} secs | LR: {:.6f}".format(
global_step, loss.numpy().item(), time.time()-t0, optim.lr.numpy().item()))
t0 = time.time()
global_step += 1
# EVALUATION
m = tf.keras.metrics.Accuracy()
for (text, seg, labels) in dataset(is_training=False, params=params):
logits = model([text, tf.sign(text), seg], training=False)
m.update_state(y_true=labels, y_pred=tf.argmax(logits, -1))
acc = m.result().numpy()
logger.info("Evaluation: Testing Accuracy: {:.3f}".format(acc))
if acc > best_acc:
best_acc = acc
# you can save model here
count = 0
else:
count += 1
logger.info("Best Accuracy: {:.3f}".format(best_acc))
if count == params['num_patience']:
print(params['num_patience'], "times not improve the best result, therefore stop training")
break
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
#read csv file
data=pd.read_csv("spam.csv",encoding="latin_1")
data.head()
#find no.of rows and columns
data.shape
#find if any NaN numbers is there in data
data.isnull().sum()
#Drop columns
data = data.drop(["Unnamed: 2", "Unnamed: 3", "Unnamed: 4"], axis=1)
#column names rechange
data = data.rename(columns={"v1":"label", "v2":"text"})
#Count observations in each label
data.label.value_counts()
# convert label to a numerical variable
data['label_num'] = data.label.map({'ham':1, 'spam':0})
data.head()
data['length'] = data['text'].apply(len)
data.head()
x=np.array(data['text'])
x
y = np.array(data['label_num'])
y
import seaborn as sns
sns.countplot(data["label"])
plt.show()
data["label"].value_counts().plot(kind="pie",autopct="%1.1f%%")
plt.axis("equal")
plt.show()
spam1=data.loc[data['label']=='spam']
spam1["text"].head()
ham1=data.loc[data['label']=='ham']
ham1["text"].head()
x_train=np.array(data.iloc[0:500,1])
x_train.shape
y_train=np.array(data.iloc[0:500,0])
y_train[0:5]
from sklearn.model_selection import train_test_split
x_train,x_test,y_train,y_test=train_test_split(x,y,test_size=0.3,random_state=1)
from sklearn.feature_extraction.text import CountVectorizer
count_vector = CountVectorizer()
train_data = count_vector.fit_transform(x_train)
test_data = count_vector.transform(x_test)
from sklearn.naive_bayes import MultinomialNB
model=MultinomialNB()
model.fit(train_data,y_train)
test_data
pred=model.predict(test_data)
pred
model.score(test_data,y_test)
#creating testing data
x_test=[ "hi how are you","when will you go to home", "i will call you back", "are you busy now",
"WINNER!! As a valued network customer you have..."]
x_test.append("goodmoring")
x_test
x_test1=np.array(x_test)
x_test1
X_train=data.iloc[0:200,1]
X_train[0:6]
Y_train=data.iloc[0:200,0]
Y_train[0:5]
from sklearn.feature_extraction.text import CountVectorizer
count_vector = CountVectorizer()
print(count_vector)
train_data = count_vector.fit_transform(X_train)
test_data = count_vector.transform(x_test1)
train_data.shape
test_data.shape
Y_train.shape
from sklearn.naive_bayes import MultinomialNB
model=MultinomialNB()
model.fit(train_data,Y_train)
pred=model.predict(test_data)
pred
df = pd.DataFrame(dict(x=x_test1, y=pred))
df
df.iloc[3:]
data.head()
ham=data[data['label']=='ham'][:747]
spam=data[data['label']=='spam']
ham.shape,spam.shape
newdata=pd.concat([ham,spam])
newdata=newdata.sample(frac=1,random_state=40)
newdata.head()
sns.countplot(x='label',data=newdata)
newdata['label'] = newdata.label.map({'ham':1, 'spam':0})
newdata.head()
X=np.array(newdata['text'])
X
Y=np.array(newdata['label'])
Y
from sklearn.model_selection import train_test_split
X_train,X_test,Y_train,Y_test=train_test_split(X,Y,test_size=0.2,random_state=42)
from sklearn.feature_extraction.text import CountVectorizer
count_vector = CountVectorizer()
train_data = count_vector.fit_transform(X_train)
test_data = count_vector.transform(X_test)
train_data.toarray()
from sklearn.naive_bayes import MultinomialNB
model=MultinomialNB()
model.fit(train_data,Y_train)
pred=model.predict(test_data)
pred
model.score(test_data,Y_test)
```
| github_jupyter |
# Exploratory Data Analysis
In statistics, **exploratory data analysis (EDA)** is an approach to analyzing data sets to summarize their main characteristics, often with visual methods ([Wikipedia](https://en.wikipedia.org/wiki/Exploratory_data_analysis)).
## The origins of EDA
[John Wilder Tukey](https://en.wikipedia.org/wiki/John_Tukey) (1915 – 2000)
<img src="./_img/EDA_Tukey_77.png" style="height: 350px;">
Tukey, John W. (1977) _Exploratory data analysis_.
## Minard's Map
[Charles Joseph Minard](https://en.wikipedia.org/wiki/Charles_Joseph_Minard) (1781 – 1870)
<img src="./_img/minards_map.png" style="height: 400px;">
## Anscombe's quartet
[Francis John Anscombe](https://en.wikipedia.org/wiki/Frank_Anscombe) (1918 – 2001)
#### *Based on descriptive statistics such as mean, variance, and regression coefficients, among others others, these 4 data sets are the same!!*
<p><p>
<font size=25>
<table style="text-align: center; margin-left:auto;
margin-right:auto; font-size: 60%; font-family : courier" border="1", width="800px", >
<tr>
<th colspan="2">I</th>
<th colspan="2">II</th>
<th colspan="2">III</th>
<th colspan="2">IV</th>
</tr>
<tr>
<td>x</td>
<td>y</td>
<td>x</td>
<td>y</td>
<td>x</td>
<td>y</td>
<td>x</td>
<td>y</td>
</tr>
<tr>
<td>10.0</td>
<td>8.04</td>
<td>10.0</td>
<td>9.14</td>
<td>10.0</td>
<td>7.46</td>
<td>8.0</td>
<td>6.58</td>
</tr>
<tr>
<td>8.0</td>
<td>6.95</td>
<td>8.0</td>
<td>8.14</td>
<td>8.0</td>
<td>6.77</td>
<td>8.0</td>
<td>5.76</td>
</tr>
<tr>
<td>13.0</td>
<td>7.58</td>
<td>13.0</td>
<td>8.74</td>
<td>13.0</td>
<td>12.74</td>
<td>8.0</td>
<td>7.71</td>
</tr>
<tr>
<td>9.0</td>
<td>8.81</td>
<td>9.0</td>
<td>8.77</td>
<td>9.0</td>
<td>7.11</td>
<td>8.0</td>
<td>8.84</td>
</tr>
<tr>
<td>11.0</td>
<td>8.33</td>
<td>11.0</td>
<td>9.26</td>
<td>11.0</td>
<td>7.81</td>
<td>8.0</td>
<td>8.47</td>
</tr>
<tr>
<td>14.0</td>
<td>9.96</td>
<td>14.0</td>
<td>8.10</td>
<td>14.0</td>
<td>8.84</td>
<td>8.0</td>
<td>7.04</td>
</tr>
<tr>
<td>6.0</td>
<td>7.24</td>
<td>6.0</td>
<td>6.13</td>
<td>6.0</td>
<td>6.08</td>
<td>8.0</td>
<td>5.25</td>
</tr>
<tr>
<td>4.0</td>
<td>4.26</td>
<td>4.0</td>
<td>3.10</td>
<td>4.0</td>
<td>5.39</td>
<td>19.0</td>
<td>12.50</td>
</tr>
<tr>
<td>12.0</td>
<td>10.84</td>
<td>12.0</td>
<td>9.13</td>
<td>12.0</td>
<td>8.15</td>
<td>8.0</td>
<td>5.56</td>
</tr>
<tr>
<td>7.0</td>
<td>4.82</td>
<td>7.0</td>
<td>7.26</td>
<td>7.0</td>
<td>6.42</td>
<td>8.0</td>
<td>7.91</td>
</tr>
<tr>
<td>5.0</td>
<td>5.68</td>
<td>5.0</td>
<td>4.74</td>
<td>5.0</td>
<td>5.73</td>
<td>8.0</td>
<td>6.89</td>
</tr>
</table>
</font>
### Plotting of the data reveals the structure of the data set
<img src="./_img/anscombes_quartet.png" style="height: 700px;">
## Let's code!
***
| github_jupyter |
# RadarCOVID-Report
## Data Extraction
```
import datetime
import json
import logging
import os
import shutil
import tempfile
import textwrap
import uuid
import matplotlib.pyplot as plt
import matplotlib.ticker
import numpy as np
import pandas as pd
import retry
import seaborn as sns
%matplotlib inline
current_working_directory = os.environ.get("PWD")
if current_working_directory:
os.chdir(current_working_directory)
sns.set()
matplotlib.rcParams["figure.figsize"] = (15, 6)
extraction_datetime = datetime.datetime.utcnow()
extraction_date = extraction_datetime.strftime("%Y-%m-%d")
extraction_previous_datetime = extraction_datetime - datetime.timedelta(days=1)
extraction_previous_date = extraction_previous_datetime.strftime("%Y-%m-%d")
extraction_date_with_hour = datetime.datetime.utcnow().strftime("%Y-%m-%d@%H")
current_hour = datetime.datetime.utcnow().hour
are_today_results_partial = current_hour != 23
```
### Constants
```
from Modules.ExposureNotification import exposure_notification_io
spain_region_country_code = "ES"
germany_region_country_code = "DE"
default_backend_identifier = spain_region_country_code
backend_generation_days = 7 * 2
daily_summary_days = 7 * 4 * 3
daily_plot_days = 7 * 4
tek_dumps_load_limit = daily_summary_days + 1
```
### Parameters
```
environment_backend_identifier = os.environ.get("RADARCOVID_REPORT__BACKEND_IDENTIFIER")
if environment_backend_identifier:
report_backend_identifier = environment_backend_identifier
else:
report_backend_identifier = default_backend_identifier
report_backend_identifier
environment_enable_multi_backend_download = \
os.environ.get("RADARCOVID_REPORT__ENABLE_MULTI_BACKEND_DOWNLOAD")
if environment_enable_multi_backend_download:
report_backend_identifiers = None
else:
report_backend_identifiers = [report_backend_identifier]
report_backend_identifiers
environment_invalid_shared_diagnoses_dates = \
os.environ.get("RADARCOVID_REPORT__INVALID_SHARED_DIAGNOSES_DATES")
if environment_invalid_shared_diagnoses_dates:
invalid_shared_diagnoses_dates = environment_invalid_shared_diagnoses_dates.split(",")
else:
invalid_shared_diagnoses_dates = []
invalid_shared_diagnoses_dates
```
### COVID-19 Cases
```
report_backend_client = \
exposure_notification_io.get_backend_client_with_identifier(
backend_identifier=report_backend_identifier)
@retry.retry(tries=10, delay=10, backoff=1.1, jitter=(0, 10))
def download_cases_dataframe_from_ecdc():
return pd.read_csv(
"https://opendata.ecdc.europa.eu/covid19/casedistribution/csv/data.csv")
confirmed_df_ = download_cases_dataframe_from_ecdc()
confirmed_df = confirmed_df_.copy()
confirmed_df = confirmed_df[["dateRep", "cases", "geoId"]]
confirmed_df.rename(
columns={
"dateRep":"sample_date",
"cases": "new_cases",
"geoId": "country_code",
},
inplace=True)
confirmed_df["sample_date"] = pd.to_datetime(confirmed_df.sample_date, dayfirst=True)
confirmed_df.sort_values("sample_date", inplace=True)
confirmed_df.tail()
confirmed_days = pd.date_range(
start=confirmed_df.iloc[0].sample_date,
end=extraction_datetime)
source_regions_at_date_df = pd.DataFrame(data=confirmed_days, columns=["sample_date"])
source_regions_at_date_df["source_regions_at_date"] = \
source_regions_at_date_df.sample_date.apply(
lambda x: report_backend_client.source_regions_for_date(date=x))
source_regions_at_date_df.sort_values("sample_date", inplace=True)
source_regions_at_date_df.tail()
confirmed_df = source_regions_at_date_df.merge(confirmed_df, how="left")
confirmed_df = confirmed_df[confirmed_df.apply(
lambda x: x.country_code in x.source_regions_at_date, axis=1)]
confirmed_df.drop(columns=["source_regions_at_date"], inplace=True)
confirmed_df = source_regions_at_date_df.merge(confirmed_df, how="left")
confirmed_df["sample_date"] = confirmed_df.sample_date.dt.strftime("%Y-%m-%d")
confirmed_df.sort_values("sample_date", inplace=True)
confirmed_df.tail()
def sort_source_regions_for_display(source_regions: list) -> list:
if report_backend_identifier in source_regions:
source_regions = [report_backend_identifier] + \
list(sorted(set(source_regions).difference([report_backend_identifier])))
else:
source_regions = list(sorted(source_regions))
return source_regions
report_source_regions = confirmed_df.country_code.dropna().unique().tolist()
report_source_regions = sort_source_regions_for_display(
source_regions=report_source_regions)
confirmed_df["_source_regions_group"] = confirmed_df.source_regions_at_date.apply(
lambda x: ",".join(sort_source_regions_for_display(x)))
confirmed_df = confirmed_df.groupby(["sample_date", "_source_regions_group"]).new_cases.sum(min_count=1).reset_index()
confirmed_df.sort_values("sample_date", inplace=True)
confirmed_df.tail()
confirmed_output_columns = ["sample_date", "new_cases", "covid_cases"]
confirmed_output_df = pd.DataFrame(columns=confirmed_output_columns)
for source_regions_group in confirmed_df._source_regions_group.unique().tolist():
confirmed_source_regions_group_df = \
confirmed_df[confirmed_df._source_regions_group == source_regions_group].copy()
confirmed_df.sort_values("sample_date", inplace=True)
confirmed_source_regions_group_df["covid_cases"] = \
confirmed_source_regions_group_df.new_cases.rolling(7, min_periods=0).mean().round()
confirmed_source_regions_group_df = \
confirmed_source_regions_group_df[confirmed_output_columns]
confirmed_source_regions_group_df.fillna(method="ffill", inplace=True)
confirmed_output_df = confirmed_output_df.append(confirmed_source_regions_group_df)
confirmed_df = confirmed_output_df.copy()
confirmed_df.tail()
confirmed_df.rename(columns={"sample_date": "sample_date_string"}, inplace=True)
confirmed_df.sort_values("sample_date_string", inplace=True)
confirmed_df.tail()
confirmed_df[["new_cases", "covid_cases"]].plot()
```
### Extract API TEKs
```
raw_zip_path_prefix = "Data/TEKs/Raw/"
fail_on_error_backend_identifiers = [report_backend_identifier]
multi_backend_exposure_keys_df = \
exposure_notification_io.download_exposure_keys_from_backends(
backend_identifiers=report_backend_identifiers,
generation_days=backend_generation_days,
fail_on_error_backend_identifiers=fail_on_error_backend_identifiers,
save_raw_zip_path_prefix=raw_zip_path_prefix)
multi_backend_exposure_keys_df["region"] = multi_backend_exposure_keys_df["backend_identifier"]
multi_backend_exposure_keys_df.rename(
columns={
"generation_datetime": "sample_datetime",
"generation_date_string": "sample_date_string",
},
inplace=True)
multi_backend_exposure_keys_df.head()
early_teks_df = multi_backend_exposure_keys_df[
multi_backend_exposure_keys_df.rolling_period < 144].copy()
early_teks_df["rolling_period_in_hours"] = early_teks_df.rolling_period / 6
early_teks_df[early_teks_df.sample_date_string != extraction_date] \
.rolling_period_in_hours.hist(bins=list(range(24)))
early_teks_df[early_teks_df.sample_date_string == extraction_date] \
.rolling_period_in_hours.hist(bins=list(range(24)))
multi_backend_exposure_keys_df = multi_backend_exposure_keys_df[[
"sample_date_string", "region", "key_data"]]
multi_backend_exposure_keys_df.head()
active_regions = \
multi_backend_exposure_keys_df.groupby("region").key_data.nunique().sort_values().index.unique().tolist()
active_regions
multi_backend_summary_df = multi_backend_exposure_keys_df.groupby(
["sample_date_string", "region"]).key_data.nunique().reset_index() \
.pivot(index="sample_date_string", columns="region") \
.sort_index(ascending=False)
multi_backend_summary_df.rename(
columns={"key_data": "shared_teks_by_generation_date"},
inplace=True)
multi_backend_summary_df.rename_axis("sample_date", inplace=True)
multi_backend_summary_df = multi_backend_summary_df.fillna(0).astype(int)
multi_backend_summary_df = multi_backend_summary_df.head(backend_generation_days)
multi_backend_summary_df.head()
def compute_keys_cross_sharing(x):
teks_x = x.key_data_x.item()
common_teks = set(teks_x).intersection(x.key_data_y.item())
common_teks_fraction = len(common_teks) / len(teks_x)
return pd.Series(dict(
common_teks=common_teks,
common_teks_fraction=common_teks_fraction,
))
multi_backend_exposure_keys_by_region_df = \
multi_backend_exposure_keys_df.groupby("region").key_data.unique().reset_index()
multi_backend_exposure_keys_by_region_df["_merge"] = True
multi_backend_exposure_keys_by_region_combination_df = \
multi_backend_exposure_keys_by_region_df.merge(
multi_backend_exposure_keys_by_region_df, on="_merge")
multi_backend_exposure_keys_by_region_combination_df.drop(
columns=["_merge"], inplace=True)
if multi_backend_exposure_keys_by_region_combination_df.region_x.nunique() > 1:
multi_backend_exposure_keys_by_region_combination_df = \
multi_backend_exposure_keys_by_region_combination_df[
multi_backend_exposure_keys_by_region_combination_df.region_x !=
multi_backend_exposure_keys_by_region_combination_df.region_y]
multi_backend_exposure_keys_cross_sharing_df = \
multi_backend_exposure_keys_by_region_combination_df \
.groupby(["region_x", "region_y"]) \
.apply(compute_keys_cross_sharing) \
.reset_index()
multi_backend_cross_sharing_summary_df = \
multi_backend_exposure_keys_cross_sharing_df.pivot_table(
values=["common_teks_fraction"],
columns="region_x",
index="region_y",
aggfunc=lambda x: x.item())
multi_backend_cross_sharing_summary_df
multi_backend_without_active_region_exposure_keys_df = \
multi_backend_exposure_keys_df[multi_backend_exposure_keys_df.region != report_backend_identifier]
multi_backend_without_active_region = \
multi_backend_without_active_region_exposure_keys_df.groupby("region").key_data.nunique().sort_values().index.unique().tolist()
multi_backend_without_active_region
exposure_keys_summary_df = multi_backend_exposure_keys_df[
multi_backend_exposure_keys_df.region == report_backend_identifier]
exposure_keys_summary_df.drop(columns=["region"], inplace=True)
exposure_keys_summary_df = \
exposure_keys_summary_df.groupby(["sample_date_string"]).key_data.nunique().to_frame()
exposure_keys_summary_df = \
exposure_keys_summary_df.reset_index().set_index("sample_date_string")
exposure_keys_summary_df.sort_index(ascending=False, inplace=True)
exposure_keys_summary_df.rename(columns={"key_data": "shared_teks_by_generation_date"}, inplace=True)
exposure_keys_summary_df.head()
```
### Dump API TEKs
```
tek_list_df = multi_backend_exposure_keys_df[
["sample_date_string", "region", "key_data"]].copy()
tek_list_df["key_data"] = tek_list_df["key_data"].apply(str)
tek_list_df.rename(columns={
"sample_date_string": "sample_date",
"key_data": "tek_list"}, inplace=True)
tek_list_df = tek_list_df.groupby(
["sample_date", "region"]).tek_list.unique().reset_index()
tek_list_df["extraction_date"] = extraction_date
tek_list_df["extraction_date_with_hour"] = extraction_date_with_hour
tek_list_path_prefix = "Data/TEKs/"
tek_list_current_path = tek_list_path_prefix + f"/Current/RadarCOVID-TEKs.json"
tek_list_daily_path = tek_list_path_prefix + f"Daily/RadarCOVID-TEKs-{extraction_date}.json"
tek_list_hourly_path = tek_list_path_prefix + f"Hourly/RadarCOVID-TEKs-{extraction_date_with_hour}.json"
for path in [tek_list_current_path, tek_list_daily_path, tek_list_hourly_path]:
os.makedirs(os.path.dirname(path), exist_ok=True)
tek_list_df.drop(columns=["extraction_date", "extraction_date_with_hour"]).to_json(
tek_list_current_path,
lines=True, orient="records")
tek_list_df.drop(columns=["extraction_date_with_hour"]).to_json(
tek_list_daily_path,
lines=True, orient="records")
tek_list_df.to_json(
tek_list_hourly_path,
lines=True, orient="records")
tek_list_df.head()
```
### Load TEK Dumps
```
import glob
def load_extracted_teks(mode, region=None, limit=None) -> pd.DataFrame:
extracted_teks_df = pd.DataFrame(columns=["region"])
file_paths = list(reversed(sorted(glob.glob(tek_list_path_prefix + mode + "/RadarCOVID-TEKs-*.json"))))
if limit:
file_paths = file_paths[:limit]
for file_path in file_paths:
logging.info(f"Loading TEKs from '{file_path}'...")
iteration_extracted_teks_df = pd.read_json(file_path, lines=True)
extracted_teks_df = extracted_teks_df.append(
iteration_extracted_teks_df, sort=False)
extracted_teks_df["region"] = \
extracted_teks_df.region.fillna(spain_region_country_code).copy()
if region:
extracted_teks_df = \
extracted_teks_df[extracted_teks_df.region == region]
return extracted_teks_df
daily_extracted_teks_df = load_extracted_teks(
mode="Daily",
region=report_backend_identifier,
limit=tek_dumps_load_limit)
daily_extracted_teks_df.head()
exposure_keys_summary_df_ = daily_extracted_teks_df \
.sort_values("extraction_date", ascending=False) \
.groupby("sample_date").tek_list.first() \
.to_frame()
exposure_keys_summary_df_.index.name = "sample_date_string"
exposure_keys_summary_df_["tek_list"] = \
exposure_keys_summary_df_.tek_list.apply(len)
exposure_keys_summary_df_ = exposure_keys_summary_df_ \
.rename(columns={"tek_list": "shared_teks_by_generation_date"}) \
.sort_index(ascending=False)
exposure_keys_summary_df = exposure_keys_summary_df_
exposure_keys_summary_df.head()
```
### Daily New TEKs
```
tek_list_df = daily_extracted_teks_df.groupby("extraction_date").tek_list.apply(
lambda x: set(sum(x, []))).reset_index()
tek_list_df = tek_list_df.set_index("extraction_date").sort_index(ascending=True)
tek_list_df.head()
def compute_teks_by_generation_and_upload_date(date):
day_new_teks_set_df = tek_list_df.copy().diff()
try:
day_new_teks_set = day_new_teks_set_df[
day_new_teks_set_df.index == date].tek_list.item()
except ValueError:
day_new_teks_set = None
if pd.isna(day_new_teks_set):
day_new_teks_set = set()
day_new_teks_df = daily_extracted_teks_df[
daily_extracted_teks_df.extraction_date == date].copy()
day_new_teks_df["shared_teks"] = \
day_new_teks_df.tek_list.apply(lambda x: set(x).intersection(day_new_teks_set))
day_new_teks_df["shared_teks"] = \
day_new_teks_df.shared_teks.apply(len)
day_new_teks_df["upload_date"] = date
day_new_teks_df.rename(columns={"sample_date": "generation_date"}, inplace=True)
day_new_teks_df = day_new_teks_df[
["upload_date", "generation_date", "shared_teks"]]
day_new_teks_df["generation_to_upload_days"] = \
(pd.to_datetime(day_new_teks_df.upload_date) -
pd.to_datetime(day_new_teks_df.generation_date)).dt.days
day_new_teks_df = day_new_teks_df[day_new_teks_df.shared_teks > 0]
return day_new_teks_df
shared_teks_generation_to_upload_df = pd.DataFrame()
for upload_date in daily_extracted_teks_df.extraction_date.unique():
shared_teks_generation_to_upload_df = \
shared_teks_generation_to_upload_df.append(
compute_teks_by_generation_and_upload_date(date=upload_date))
shared_teks_generation_to_upload_df \
.sort_values(["upload_date", "generation_date"], ascending=False, inplace=True)
shared_teks_generation_to_upload_df.tail()
today_new_teks_df = \
shared_teks_generation_to_upload_df[
shared_teks_generation_to_upload_df.upload_date == extraction_date].copy()
today_new_teks_df.tail()
if not today_new_teks_df.empty:
today_new_teks_df.set_index("generation_to_upload_days") \
.sort_index().shared_teks.plot.bar()
generation_to_upload_period_pivot_df = \
shared_teks_generation_to_upload_df[
["upload_date", "generation_to_upload_days", "shared_teks"]] \
.pivot(index="upload_date", columns="generation_to_upload_days") \
.sort_index(ascending=False).fillna(0).astype(int) \
.droplevel(level=0, axis=1)
generation_to_upload_period_pivot_df.head()
new_tek_df = tek_list_df.diff().tek_list.apply(
lambda x: len(x) if not pd.isna(x) else None).to_frame().reset_index()
new_tek_df.rename(columns={
"tek_list": "shared_teks_by_upload_date",
"extraction_date": "sample_date_string",}, inplace=True)
new_tek_df.tail()
shared_teks_uploaded_on_generation_date_df = shared_teks_generation_to_upload_df[
shared_teks_generation_to_upload_df.generation_to_upload_days == 0] \
[["upload_date", "shared_teks"]].rename(
columns={
"upload_date": "sample_date_string",
"shared_teks": "shared_teks_uploaded_on_generation_date",
})
shared_teks_uploaded_on_generation_date_df.head()
estimated_shared_diagnoses_df = shared_teks_generation_to_upload_df \
.groupby(["upload_date"]).shared_teks.max().reset_index() \
.sort_values(["upload_date"], ascending=False) \
.rename(columns={
"upload_date": "sample_date_string",
"shared_teks": "shared_diagnoses",
})
invalid_shared_diagnoses_dates_mask = \
estimated_shared_diagnoses_df.sample_date_string.isin(invalid_shared_diagnoses_dates)
estimated_shared_diagnoses_df[invalid_shared_diagnoses_dates_mask] = 0
estimated_shared_diagnoses_df.head()
```
### Hourly New TEKs
```
hourly_extracted_teks_df = load_extracted_teks(
mode="Hourly", region=report_backend_identifier, limit=25)
hourly_extracted_teks_df.head()
hourly_new_tek_count_df = hourly_extracted_teks_df \
.groupby("extraction_date_with_hour").tek_list. \
apply(lambda x: set(sum(x, []))).reset_index().copy()
hourly_new_tek_count_df = hourly_new_tek_count_df.set_index("extraction_date_with_hour") \
.sort_index(ascending=True)
hourly_new_tek_count_df["new_tek_list"] = hourly_new_tek_count_df.tek_list.diff()
hourly_new_tek_count_df["new_tek_count"] = hourly_new_tek_count_df.new_tek_list.apply(
lambda x: len(x) if not pd.isna(x) else 0)
hourly_new_tek_count_df.rename(columns={
"new_tek_count": "shared_teks_by_upload_date"}, inplace=True)
hourly_new_tek_count_df = hourly_new_tek_count_df.reset_index()[[
"extraction_date_with_hour", "shared_teks_by_upload_date"]]
hourly_new_tek_count_df.head()
hourly_summary_df = hourly_new_tek_count_df.copy()
hourly_summary_df.set_index("extraction_date_with_hour", inplace=True)
hourly_summary_df = hourly_summary_df.fillna(0).astype(int).reset_index()
hourly_summary_df["datetime_utc"] = pd.to_datetime(
hourly_summary_df.extraction_date_with_hour, format="%Y-%m-%d@%H")
hourly_summary_df.set_index("datetime_utc", inplace=True)
hourly_summary_df = hourly_summary_df.tail(-1)
hourly_summary_df.head()
```
### Data Merge
```
result_summary_df = exposure_keys_summary_df.merge(
new_tek_df, on=["sample_date_string"], how="outer")
result_summary_df.head()
result_summary_df = result_summary_df.merge(
shared_teks_uploaded_on_generation_date_df, on=["sample_date_string"], how="outer")
result_summary_df.head()
result_summary_df = result_summary_df.merge(
estimated_shared_diagnoses_df, on=["sample_date_string"], how="outer")
result_summary_df.head()
result_summary_df = confirmed_df.tail(daily_summary_days).merge(
result_summary_df, on=["sample_date_string"], how="left")
result_summary_df.head()
result_summary_df["sample_date"] = pd.to_datetime(result_summary_df.sample_date_string)
source_regions_for_summary_df = \
source_regions_at_date_df.rename(columns={"source_regions_at_date": "source_regions"})
source_regions_for_summary_df["source_regions"] = \
source_regions_for_summary_df.source_regions.apply(lambda x: ",".join(sort_source_regions_for_display(x)))
source_regions_for_summary_df.head()
result_summary_df = result_summary_df.merge(source_regions_for_summary_df, how="left")
result_summary_df.set_index(["sample_date", "source_regions"], inplace=True)
result_summary_df.drop(columns=["sample_date_string"], inplace=True)
result_summary_df.sort_index(ascending=False, inplace=True)
result_summary_df.head()
with pd.option_context("mode.use_inf_as_na", True):
result_summary_df = result_summary_df.fillna(0).astype(int)
result_summary_df["teks_per_shared_diagnosis"] = \
(result_summary_df.shared_teks_by_upload_date / result_summary_df.shared_diagnoses).fillna(0)
result_summary_df["shared_diagnoses_per_covid_case"] = \
(result_summary_df.shared_diagnoses / result_summary_df.covid_cases).fillna(0)
result_summary_df.head(daily_plot_days)
weekly_result_summary_df = result_summary_df \
.sort_index(ascending=True).fillna(0).rolling(7).agg({
"covid_cases": "sum",
"shared_teks_by_generation_date": "sum",
"shared_teks_by_upload_date": "sum",
"shared_diagnoses": "sum"
}).sort_index(ascending=False)
with pd.option_context("mode.use_inf_as_na", True):
weekly_result_summary_df = weekly_result_summary_df.fillna(0).astype(int)
weekly_result_summary_df["teks_per_shared_diagnosis"] = \
(weekly_result_summary_df.shared_teks_by_upload_date / weekly_result_summary_df.shared_diagnoses).fillna(0)
weekly_result_summary_df["shared_diagnoses_per_covid_case"] = \
(weekly_result_summary_df.shared_diagnoses / weekly_result_summary_df.covid_cases).fillna(0)
weekly_result_summary_df.head()
last_7_days_summary = weekly_result_summary_df.to_dict(orient="records")[1]
last_7_days_summary
```
## Report Results
```
display_column_name_mapping = {
"sample_date": "Sample\u00A0Date\u00A0(UTC)",
"source_regions": "Source Countries",
"datetime_utc": "Timestamp (UTC)",
"upload_date": "Upload Date (UTC)",
"generation_to_upload_days": "Generation to Upload Period in Days",
"region": "Backend",
"region_x": "Backend\u00A0(A)",
"region_y": "Backend\u00A0(B)",
"common_teks": "Common TEKs Shared Between Backends",
"common_teks_fraction": "Fraction of TEKs in Backend (A) Available in Backend (B)",
"covid_cases": "COVID-19 Cases in Source Countries (7-day Rolling Average)",
"shared_teks_by_generation_date": "Shared TEKs by Generation Date",
"shared_teks_by_upload_date": "Shared TEKs by Upload Date",
"shared_diagnoses": "Shared Diagnoses (Estimation)",
"teks_per_shared_diagnosis": "TEKs Uploaded per Shared Diagnosis",
"shared_diagnoses_per_covid_case": "Usage Ratio (Fraction of Cases in Source Countries Which Shared Diagnosis)",
"shared_teks_uploaded_on_generation_date": "Shared TEKs Uploaded on Generation Date",
}
summary_columns = [
"covid_cases",
"shared_teks_by_generation_date",
"shared_teks_by_upload_date",
"shared_teks_uploaded_on_generation_date",
"shared_diagnoses",
"teks_per_shared_diagnosis",
"shared_diagnoses_per_covid_case",
]
```
### Daily Summary Table
```
result_summary_df_ = result_summary_df.copy()
result_summary_df = result_summary_df[summary_columns]
result_summary_with_display_names_df = result_summary_df \
.rename_axis(index=display_column_name_mapping) \
.rename(columns=display_column_name_mapping)
result_summary_with_display_names_df
```
### Daily Summary Plots
```
result_plot_summary_df = result_summary_df.head(daily_plot_days)[summary_columns] \
.droplevel(level=["source_regions"]) \
.rename_axis(index=display_column_name_mapping) \
.rename(columns=display_column_name_mapping)
summary_ax_list = result_plot_summary_df.sort_index(ascending=True).plot.bar(
title=f"Daily Summary",
rot=45, subplots=True, figsize=(15, 22), legend=False)
ax_ = summary_ax_list[-1]
ax_.get_figure().tight_layout()
ax_.get_figure().subplots_adjust(top=0.95)
ax_.yaxis.set_major_formatter(matplotlib.ticker.PercentFormatter(1.0))
_ = ax_.set_xticklabels(sorted(result_plot_summary_df.index.strftime("%Y-%m-%d").tolist()))
```
### Daily Generation to Upload Period Table
```
display_generation_to_upload_period_pivot_df = \
generation_to_upload_period_pivot_df \
.head(backend_generation_days)
display_generation_to_upload_period_pivot_df \
.head(backend_generation_days) \
.rename_axis(columns=display_column_name_mapping) \
.rename_axis(index=display_column_name_mapping)
fig, generation_to_upload_period_pivot_table_ax = plt.subplots(
figsize=(12, 1 + 0.6 * len(display_generation_to_upload_period_pivot_df)))
generation_to_upload_period_pivot_table_ax.set_title(
"Shared TEKs Generation to Upload Period Table")
sns.heatmap(
data=display_generation_to_upload_period_pivot_df
.rename_axis(columns=display_column_name_mapping)
.rename_axis(index=display_column_name_mapping),
fmt=".0f",
annot=True,
ax=generation_to_upload_period_pivot_table_ax)
generation_to_upload_period_pivot_table_ax.get_figure().tight_layout()
```
### Hourly Summary Plots
```
hourly_summary_ax_list = hourly_summary_df \
.rename_axis(index=display_column_name_mapping) \
.rename(columns=display_column_name_mapping) \
.plot.bar(
title=f"Last 24h Summary",
rot=45, subplots=True, legend=False)
ax_ = hourly_summary_ax_list[-1]
ax_.get_figure().tight_layout()
ax_.get_figure().subplots_adjust(top=0.9)
_ = ax_.set_xticklabels(sorted(hourly_summary_df.index.strftime("%Y-%m-%d@%H").tolist()))
```
### Publish Results
```
def get_temporary_image_path() -> str:
return os.path.join(tempfile.gettempdir(), str(uuid.uuid4()) + ".png")
def save_temporary_plot_image(ax):
if isinstance(ax, np.ndarray):
ax = ax[0]
media_path = get_temporary_image_path()
ax.get_figure().savefig(media_path)
return media_path
def save_temporary_dataframe_image(df):
import dataframe_image as dfi
media_path = get_temporary_image_path()
dfi.export(df, media_path)
return media_path
github_repository = os.environ.get("GITHUB_REPOSITORY")
if github_repository is None:
github_repository = "pvieito/Radar-STATS"
github_project_base_url = "https://github.com/" + github_repository
display_formatters = {
display_column_name_mapping["teks_per_shared_diagnosis"]: lambda x: f"{x:.2f}",
display_column_name_mapping["shared_diagnoses_per_covid_case"]: lambda x: f"{x:.2%}",
}
daily_summary_table_html = result_summary_with_display_names_df \
.head(daily_plot_days) \
.rename_axis(index=display_column_name_mapping) \
.rename(columns=display_column_name_mapping) \
.to_html(formatters=display_formatters)
multi_backend_summary_table_html = multi_backend_summary_df \
.head(daily_plot_days) \
.rename_axis(columns=display_column_name_mapping) \
.rename(columns=display_column_name_mapping) \
.rename_axis(index=display_column_name_mapping) \
.to_html(formatters=display_formatters)
def format_multi_backend_cross_sharing_fraction(x):
if pd.isna(x):
return "-"
elif round(x * 100, 1) == 0:
return ""
else:
return f"{x:.1%}"
multi_backend_cross_sharing_summary_table_html = multi_backend_cross_sharing_summary_df \
.rename_axis(columns=display_column_name_mapping) \
.rename(columns=display_column_name_mapping) \
.rename_axis(index=display_column_name_mapping) \
.to_html(
classes="table-center",
formatters=display_formatters,
float_format=format_multi_backend_cross_sharing_fraction)
multi_backend_cross_sharing_summary_table_html = \
multi_backend_cross_sharing_summary_table_html \
.replace("<tr>","<tr style=\"text-align: center;\">")
extraction_date_result_summary_df = \
result_summary_df[result_summary_df.index.get_level_values("sample_date") == extraction_date]
extraction_date_result_hourly_summary_df = \
hourly_summary_df[hourly_summary_df.extraction_date_with_hour == extraction_date_with_hour]
covid_cases = \
extraction_date_result_summary_df.covid_cases.sum()
shared_teks_by_generation_date = \
extraction_date_result_summary_df.shared_teks_by_generation_date.sum()
shared_teks_by_upload_date = \
extraction_date_result_summary_df.shared_teks_by_upload_date.sum()
shared_diagnoses = \
extraction_date_result_summary_df.shared_diagnoses.sum()
teks_per_shared_diagnosis = \
extraction_date_result_summary_df.teks_per_shared_diagnosis.sum()
shared_diagnoses_per_covid_case = \
extraction_date_result_summary_df.shared_diagnoses_per_covid_case.sum()
shared_teks_by_upload_date_last_hour = \
extraction_date_result_hourly_summary_df.shared_teks_by_upload_date.sum().astype(int)
display_source_regions = ", ".join(report_source_regions)
if len(report_source_regions) == 1:
display_brief_source_regions = report_source_regions[0]
else:
display_brief_source_regions = f"{len(report_source_regions)} 🇪🇺"
summary_plots_image_path = save_temporary_plot_image(
ax=summary_ax_list)
summary_table_image_path = save_temporary_dataframe_image(
df=result_summary_with_display_names_df)
hourly_summary_plots_image_path = save_temporary_plot_image(
ax=hourly_summary_ax_list)
multi_backend_summary_table_image_path = save_temporary_dataframe_image(
df=multi_backend_summary_df)
generation_to_upload_period_pivot_table_image_path = save_temporary_plot_image(
ax=generation_to_upload_period_pivot_table_ax)
```
### Save Results
```
report_resources_path_prefix = "Data/Resources/Current/RadarCOVID-Report-"
result_summary_df.to_csv(
report_resources_path_prefix + "Summary-Table.csv")
result_summary_df.to_html(
report_resources_path_prefix + "Summary-Table.html")
hourly_summary_df.to_csv(
report_resources_path_prefix + "Hourly-Summary-Table.csv")
multi_backend_summary_df.to_csv(
report_resources_path_prefix + "Multi-Backend-Summary-Table.csv")
multi_backend_cross_sharing_summary_df.to_csv(
report_resources_path_prefix + "Multi-Backend-Cross-Sharing-Summary-Table.csv")
generation_to_upload_period_pivot_df.to_csv(
report_resources_path_prefix + "Generation-Upload-Period-Table.csv")
_ = shutil.copyfile(
summary_plots_image_path,
report_resources_path_prefix + "Summary-Plots.png")
_ = shutil.copyfile(
summary_table_image_path,
report_resources_path_prefix + "Summary-Table.png")
_ = shutil.copyfile(
hourly_summary_plots_image_path,
report_resources_path_prefix + "Hourly-Summary-Plots.png")
_ = shutil.copyfile(
multi_backend_summary_table_image_path,
report_resources_path_prefix + "Multi-Backend-Summary-Table.png")
_ = shutil.copyfile(
generation_to_upload_period_pivot_table_image_path,
report_resources_path_prefix + "Generation-Upload-Period-Table.png")
```
### Publish Results as JSON
```
summary_results_api_df = result_summary_df.reset_index()
summary_results_api_df["sample_date_string"] = \
summary_results_api_df["sample_date"].dt.strftime("%Y-%m-%d")
summary_results_api_df["source_regions"] = \
summary_results_api_df["source_regions"].apply(lambda x: x.split(","))
today_summary_results_api_df = \
summary_results_api_df.to_dict(orient="records")[0]
summary_results = dict(
backend_identifier=report_backend_identifier,
source_regions=report_source_regions,
extraction_datetime=extraction_datetime,
extraction_date=extraction_date,
extraction_date_with_hour=extraction_date_with_hour,
last_hour=dict(
shared_teks_by_upload_date=shared_teks_by_upload_date_last_hour,
shared_diagnoses=0,
),
today=today_summary_results_api_df,
last_7_days=last_7_days_summary,
daily_results=summary_results_api_df.to_dict(orient="records"))
summary_results = \
json.loads(pd.Series([summary_results]).to_json(orient="records"))[0]
with open(report_resources_path_prefix + "Summary-Results.json", "w") as f:
json.dump(summary_results, f, indent=4)
```
### Publish on README
```
with open("Data/Templates/README.md", "r") as f:
readme_contents = f.read()
readme_contents = readme_contents.format(
extraction_date_with_hour=extraction_date_with_hour,
github_project_base_url=github_project_base_url,
daily_summary_table_html=daily_summary_table_html,
multi_backend_summary_table_html=multi_backend_summary_table_html,
multi_backend_cross_sharing_summary_table_html=multi_backend_cross_sharing_summary_table_html,
display_source_regions=display_source_regions)
with open("README.md", "w") as f:
f.write(readme_contents)
```
### Publish on Twitter
```
enable_share_to_twitter = os.environ.get("RADARCOVID_REPORT__ENABLE_PUBLISH_ON_TWITTER")
github_event_name = os.environ.get("GITHUB_EVENT_NAME")
if enable_share_to_twitter and github_event_name == "schedule" and \
(shared_teks_by_upload_date_last_hour or not are_today_results_partial):
import tweepy
twitter_api_auth_keys = os.environ["RADARCOVID_REPORT__TWITTER_API_AUTH_KEYS"]
twitter_api_auth_keys = twitter_api_auth_keys.split(":")
auth = tweepy.OAuthHandler(twitter_api_auth_keys[0], twitter_api_auth_keys[1])
auth.set_access_token(twitter_api_auth_keys[2], twitter_api_auth_keys[3])
api = tweepy.API(auth)
summary_plots_media = api.media_upload(summary_plots_image_path)
summary_table_media = api.media_upload(summary_table_image_path)
generation_to_upload_period_pivot_table_image_media = api.media_upload(generation_to_upload_period_pivot_table_image_path)
media_ids = [
summary_plots_media.media_id,
summary_table_media.media_id,
generation_to_upload_period_pivot_table_image_media.media_id,
]
if are_today_results_partial:
today_addendum = " (Partial)"
else:
today_addendum = ""
status = textwrap.dedent(f"""
#RadarCOVID – {extraction_date_with_hour}
Source Countries: {display_brief_source_regions}
Today{today_addendum}:
- Uploaded TEKs: {shared_teks_by_upload_date:.0f} ({shared_teks_by_upload_date_last_hour:+d} last hour)
- Shared Diagnoses: ≤{shared_diagnoses:.0f}
- Usage Ratio: ≤{shared_diagnoses_per_covid_case:.2%}
Last 7 Days:
- Shared Diagnoses: ≤{last_7_days_summary["shared_diagnoses"]:.0f}
- Usage Ratio: ≤{last_7_days_summary["shared_diagnoses_per_covid_case"]:.2%}
Info: {github_project_base_url}#documentation
""")
status = status.encode(encoding="utf-8")
api.update_status(status=status, media_ids=media_ids)
```
| github_jupyter |
```
######################################## Prediction ###############################################
#generating patches from predictions image
def prediction_patch(X,patch_size=320):
patch_paths=[]
patch_count=0
print(X.shape)
n_vertical_patches=math.ceil(X.shape[0]/patch_size)
n_horizontal_patches=math.ceil(X.shape[1]/patch_size)
print("no. of horizontal patches:",n_horizontal_patches," no. of vertical patches:",n_vertical_patches)
print(X.shape)
for k in range(0,n_vertical_patches):
for l in range(0,n_horizontal_patches):
temp_x=X[k*patch_size:(k+1)*patch_size,l*patch_size:(l+1)*patch_size,:]
print("loading entry "+str(patch_count)+"......")
#print(k*patch_size,(k+1)*patch_size,l*patch_size,(l+1)*patch_size)
np.save("Image_patch_test/img_patch"+str(patch_count)+".npy",temp_x)
patch_paths.append("Image_patch_test/img_patch"+str(patch_count)+".npy")
patch_count=patch_count+1
return patch_count,patch_paths #returns the no. of patches generated and the corresponding patch paths
############################### Prediction ############################################
#returns the prediction mask for a given image (path)
def get_prediction(image_path):
#image_path is the path of the image over which prediction has to be generated
if os.path.exists(image_path):
#loading the image
img=np.array(imageio.imread(image_path,format="tiff"))
shape=img.shape
print(img.shape,end=" ")
#preprocessing the image
img = normalize(img)
img=resize(img)
print("shape of image:",img.shape)
img=np.array(img)
n_patches,patch_paths=prediction_patch(img)
count=0
patches=[]
predict_paths=[]
for path in patch_paths:
patch=np.load(path)
patches.append(patch)
patches=np.array(patches)
predictions=new_model.predict(patches,batch_size=4)
for prediction in predictions:
prediction=np.argmax(prediction,axis=2)
image_class=get_image_from_classes(prediction)
np.save("Prediction_patches/patch"+str(count)+".npy",image_class)
predict_paths.append("Prediction_patches/patch"+str(count)+".npy")
count=count+1
image=restore_image_from_patches(predict_paths,shape)
return image
return "error"
## function to generate rgb image from previosly generated patches
def restore_image_from_patches(patch_paths,image_shape,patch_size=320):
count=0
img_height,img_width,n_channels=image_shape
n_channels=3
#calculating the extended height and width based on true shape
n_vertical_patches=math.ceil(img_height/patch_size)
n_horizontal_patches=math.ceil(img_width/patch_size)
extended_height = patch_size * n_vertical_patches
extended_width = patch_size * n_horizontal_patches
ext_image = np.zeros(shape=(extended_height, extended_width, n_channels), dtype=np.float32)
print(n_vertical_patches,n_horizontal_patches)
print(extended_height,extended_width)
#joining patches to form the actual image
for k in range(0,n_vertical_patches):
for l in range(0,n_horizontal_patches):
print(count)
patch=np.load(patch_paths[count])
#print(patch.shape)
print(k*patch_size,(k+1)*patch_size,l*patch_size,(l+1)*patch_size)
print("loading entry "+str(count)+"......")
ext_image[k*patch_size:(k+1)*patch_size,l*patch_size:(l+1)*patch_size,:]=patch[:,:,:]
count=count+1
image=ext_image[:img_height,:img_width,:n_channels]
return image
!mkdir Image_patch_test
!mkdir Prediction_patches
path="./sat/4.tif" #give path of the test image for which prediction is to be generated
im=get_prediction(path)
print(im.shape)
################################Calculating metric scores ######################################################3
from mlxtend.evaluate import confusion_matrix
from mlxtend.plotting import plot_confusion_matrix
from sklearn.metrics import cohen_kappa_score
def slice(img):
mask=[]
for i in range(9):
mask.append(img[:,:,i])
return mask
#function to calculate the kappa coefficient per class on a training images and their corresponding predictions
def get_output():
path='./'
i=1
scores=[]
while True:
sat_path=path+'sat/'+str(i)+'.tif'
if os.path.exists(sat_path):
K_score=[]
img=np.array(imageio.imread(sat_path,format="tiff"))
print(img.shape,end=" ")
print(img.shape)
gt_path = path+'gt/'+str(i)+'.tif'
mask = cv2.imread(gt_path)
mask = cv2.cvtColor(mask, cv2.COLOR_BGR2RGB)
mask = get_mask(img,mask)
mask = slice(mask)
pred_mask=get_prediction(sat_path)
pred_mask=get_mask(img,pred_mask)
pred_mask=slice(pred_mask)
for j in range(9):
K_score.append(cohen_kappa_score(mask[j].argmax(axis=0),pred_mask[j].argmax(axis=0),labels=None))
print(K_score[j])
scores.append(K_score)
else:
print("end of loop")
break
i=i+1
return K_score
### kappa coefficient calculated on training images
### the kappa scores generated per class can be found in the output below for each of the 14 training images respectively
### the scores are printed in a series of 9 values, after loading the patches for a particular images
### the scores are for classes 0,1,2,3,4,5,6,7,8 as printes respectively
K_score=get_output()
#function to calculate the kappa coefficient per class on a training images and their corresponding predictions
def confusion_matrix1():
path="./"
i=1
stack=[]
while True:
sat_path=path+'sat/'+str(i)+'.tif'
if os.path.exists(sat_path):
img=np.array(imageio.imread(sat_path,format="tiff"))
print(img.shape)
gt_path = path+'gt/'+str(i)+'.tif'
mask = cv2.imread(gt_path)
#print("mask before getting classes",mask.shape)
mask = cv2.cvtColor(mask, cv2.COLOR_BGR2RGB)
mask=get_classes_from_image(mask)
#print("mask after getting classes",mask.shape)
#mask = resize(mask)
pred_img=get_prediction(sat_path)
#print("pred before getting classes",pred_img.shape)
pred_img=get_classes_from_image(pred_img)
#print("pred after getting classes",pred_img.shape)
mask=mask.flatten()
#print("mask after flatten",mask.shape)
pred_img=pred_img.flatten()
#print("pred after flatten",pred_img.shape)
y=confusion_matrix(mask, pred_img,binary=False)
print(y,y.shape)
stack.append(y)
else:
break
i=i+1
return stack
### the confusion matrix for each training image is printed after loading the data for each image
### the horizontal rows denote the actual classes while the columns show the predictions for each class(classes can be inferred from the corresponding indices i.e. index 0 represents class 0)
cm=confusion_matrix1()
```
| github_jupyter |
# Developing a Pretrained Alexnet model using ManufacturingNet
###### To know more about the manufacturingnet please visit: http://manufacturingnet.io/
```
import ManufacturingNet
import numpy as np
```
First we import manufacturingnet. Using manufacturingnet we can create deep learning models with greater ease.
It is important to note that all the dependencies of the package must also be installed in your environment
##### Now we first need to download the data. You can use our dataset class where we have curated different types of datasets and you just need to run two lines of code to download the data :)
```
from ManufacturingNet import datasets
datasets.CastingData()
```
##### Alright! Now please check your working directory. The data should be present inside it. That was super easy !!
The Casting dataset is an image dataset with 2 classes. The task that we need to perform using Pretrained Alexnet is classification. ManufacturingNet has also provided different datasets in the package which the user can choose depending on type of application
Pretrained models use Imagefolder dataset from pytorch and image size is (224,224,channels). The pretrained model needs the root folder path of train and test images(Imagefolder format). Manufacturing pretrained models have image resizing feature.
```
#paths of root folder
train_data_address='casting_data/train/'
val_data_address='casting_data/test/'
```
#### Now all we got to do is import the pretrained model class and answer a few simple questions and we will be all set. The manufacturingnet has been designed in a way to make things easy for user and provide them the tools to implement complex used
```
from ManufacturingNet.models import AlexNet
# from ManufacturingNet.models import ResNet
# from ManufacturingNet.models import DenseNet
# from ManufacturingNet.models import MobileNet
# from ManufacturingNet.models import GoogleNet
# from ManufacturingNet.models import VGG
```
###### We import the pretrained Alexnet model (AlexNet) from package and answer a few simple questions
```
model=ResNet(train_data_address,val_data_address)
model=ResNet(train_data_address,val_data_address)
model=ResNet(train_data_address,val_data_address)
model=AlexNet(train_data_address,val_data_address)
```
Alright! Its done you have built your pretrained AlexNet using the manufacturingnet package. Just by answering a few simple questions. It is really easy
A few pointers about developing the pretrained models. These models require image size to be (224,224,channels) as the input size of the image. The number of classes for classification can be varied and is handled by the package. User can use only the architecture without using the pretrained weights.
The loss functions, optimizer, epochs, scheduler should be chosen by the user. The model summary, training accuracy, validation accuracy, confusion matrix, Loss vs epoch are also provided by the package.
ManufacturingNet provides many pretrained models with similar scripts. ManufacturingNet offer ResNet(different variants), AlexNet, GoogleNet, VGG(different variants) and DenseNet(different variants).
Users can follow a similar tutorial on pretrained ResNet(different variants).
| github_jupyter |
This notebook produces a comparison of DES 4 photo-z estimate methods, an astroML Nadaraya Watson photo-z estimation trained on DES photometry, and the spectroscopic redshift. The tables used were produced by matching a catalog of known quasars [Drexel local file: GTR-ADM-QSO-ir-testhighz_findbw_lup_2016_starclean.fits] to the DES SVA1 "gold" catalog and then cross-matching to the 4 DES photo-z catalogs.
GTR quasars: 4193,
GTR quasars matched to DES Sva1: 35
```
import numpy as np
from astropy.table import Table
from sklearn.model_selection import train_test_split, cross_val_predict
from sklearn.metrics import classification_report
from astroML.linear_model import NadarayaWatson
import matplotlib as mpl
import matplotlib.pyplot as plt
import matplotlib.patches as mpatches
import palettable
import richardsplot as rplot
%matplotlib inline
```
These cells load in our data tables, then separate out the relevant data for training the model. The model is then given our test set and produces a photo-z prediction. The resulting predictions are then sorted by object classification. This process happens once for each of the four DES photo-z methods before plotting. If not interested in the minute details, skip to cell 14 for the plots.
```
#read in table
destable_annz = Table.read('gtr_qso_+dessva1_annz.fits')
#prevent a bug involving MaskedColumns
destable_annz = destable_annz.filled()
destable_bpz = Table.read('gtr_qso_+dessva1_bpz.fits')
destable_bpz = destable_bpz.filled()
destable_skynet = Table.read('gtr_qso_+dessva1_skynet.fits')
destable_skynet = destable_skynet.filled()
destable_tpz = Table.read('gtr_qso_+dessva1_tpz.fits')
destable_tpz = destable_tpz.filled()
#stack photometry bands, des photo-z, and des object class
X_annz = np.vstack([destable_annz['MAG_AUTO_G'], destable_annz['MAG_AUTO_R'], destable_annz['MAG_AUTO_I'], destable_annz['MAG_AUTO_Z'], destable_annz['Z_MEAN'], destable_annz['MODEST_CLASS'] ]).T
y_annz = np.array(destable_annz['zspec'])
#split into training and test sets of 1/2 and 1/2 respectively
X_train_annz, X_test_annz, y_train_annz, y_test_annz = train_test_split(X_annz, y_annz, test_size=0.5, random_state=76)
#empty arrays for photometry bands
X_traintrue_annz = np.empty((X_train_annz.shape[0], X_train_annz.shape[1]-2), dtype=float)
X_testtrue_annz = np.empty((X_test_annz.shape[0], X_test_annz.shape[1]-2), dtype=float)
#empty array for des photo-z
DesZs_annz = np.empty((X_test_annz.shape[0], 1), dtype=float)
#empty array for object class
ModestClass_annz = np.empty((X_test_annz.shape[0], 1), dtype=int)
#loop through and separate photometry bands, des photo-z and object class
for i in range(len(X_train_annz)):
X_traintrue_annz[i] = X_train_annz[i][:4] #first four entries i.e. the four photometry bands
for i in range(len(X_test_annz)):
X_testtrue_annz[i] = X_test_annz[i][:4]
DesZs_annz[i] = X_test_annz[i][4] #fifth entry, the des photo-z
ModestClass_annz[i] = X_test_annz[i][5] #sixth entry, the des object classification
#initialize model with gaussian kernel of width 0.05
model_annz = NadarayaWatson('gaussian', 0.05)
#fit model to training set
model_annz.fit(X_traintrue_annz, y_train_annz)
#produce a photo-z estimate
pred_annz = model_annz.predict(X_testtrue_annz)
X_bpz = np.vstack([destable_bpz['MAG_AUTO_G'], destable_bpz['MAG_AUTO_R'], destable_bpz['MAG_AUTO_I'], destable_bpz['MAG_AUTO_Z'], destable_bpz['Z_MEAN'], destable_bpz['MODEST_CLASS'] ]).T
y_bpz = np.array(destable_bpz['zspec'])
X_train_bpz, X_test_bpz, y_train_bpz, y_test_bpz = train_test_split(X_bpz, y_bpz, test_size=0.5, random_state=76)
X_traintrue_bpz = np.empty((X_train_bpz.shape[0], X_train_bpz.shape[1]-2), dtype=float)
X_testtrue_bpz = np.empty((X_test_bpz.shape[0], X_test_bpz.shape[1]-2), dtype=float)
DesZs_bpz = np.empty((X_test_bpz.shape[0], 1), dtype=float)
ModestClass_bpz = np.empty((X_test_bpz.shape[0], 1), dtype=int)
for i in range(len(X_train_bpz)):
X_traintrue_bpz[i] = X_train_bpz[i][:4]
for i in range(len(X_test_bpz)):
X_testtrue_bpz[i] = X_test_bpz[i][:4]
DesZs_bpz[i] = X_test_bpz[i][4]
ModestClass_bpz[i] = X_test_bpz[i][5]
model_bpz = NadarayaWatson('gaussian', 0.05)
model_bpz.fit(X_traintrue_bpz, y_train_bpz)
pred_bpz = model_bpz.predict(X_testtrue_bpz)
X_skynet = np.vstack([destable_skynet['MAG_AUTO_G'], destable_skynet['MAG_AUTO_R'], destable_skynet['MAG_AUTO_I'], destable_skynet['MAG_AUTO_Z'], destable_skynet['Z_MEAN'], destable_skynet['MODEST_CLASS'] ]).T
y_skynet = np.array(destable_skynet['zspec'])
X_train_skynet, X_test_skynet, y_train_skynet, y_test_skynet = train_test_split(X_skynet, y_skynet, test_size=0.5, random_state=76)
X_traintrue_skynet = np.empty((X_train_skynet.shape[0], X_train_skynet.shape[1]-2), dtype=float)
X_testtrue_skynet = np.empty((X_test_skynet.shape[0], X_test_skynet.shape[1]-2), dtype=float)
DesZs_skynet = np.empty((X_test_skynet.shape[0], 1), dtype=float)
ModestClass_skynet = np.empty((X_test_skynet.shape[0], 1), dtype=int)
for i in range(len(X_train_skynet)):
X_traintrue_skynet[i] = X_train_skynet[i][:4]
for i in range(len(X_test_skynet)):
X_testtrue_skynet[i] = X_test_skynet[i][:4]
DesZs_skynet[i] = X_test_skynet[i][4]
ModestClass_skynet[i] = X_test_skynet[i][5]
model_skynet = NadarayaWatson('gaussian', 0.05)
model_skynet.fit(X_traintrue_skynet, y_train_skynet)
pred_skynet = model_skynet.predict(X_testtrue_skynet)
X_tpz = np.vstack([destable_tpz['MAG_AUTO_G'], destable_tpz['MAG_AUTO_R'], destable_tpz['MAG_AUTO_I'], destable_tpz['MAG_AUTO_Z'], destable_tpz['Z_MEAN'], destable_tpz['MODEST_CLASS'] ]).T
y_tpz = np.array(destable_tpz['zspec'])
X_train_tpz, X_test_tpz, y_train_tpz, y_test_tpz = train_test_split(X_tpz, y_tpz, test_size=0.5, random_state=76)
X_traintrue_tpz = np.empty((X_train_tpz.shape[0], X_train_tpz.shape[1]-2), dtype=float)
X_testtrue_tpz = np.empty((X_test_tpz.shape[0], X_test_tpz.shape[1]-2), dtype=float)
DesZs_tpz = np.empty((X_test_tpz.shape[0], 1), dtype=float)
ModestClass_tpz = np.empty((X_test_tpz.shape[0], 1), dtype=int)
for i in range(len(X_train_tpz)):
X_traintrue_tpz[i] = X_train_tpz[i][:4]
for i in range(len(X_test_tpz)):
X_testtrue_tpz[i] = X_test_tpz[i][:4]
DesZs_tpz[i] = X_test_tpz[i][4]
ModestClass_tpz[i] = X_test_tpz[i][5]
model_tpz = NadarayaWatson('gaussian', 0.05)
model_tpz.fit(X_traintrue_tpz, y_train_tpz)
pred_tpz = model_tpz.predict(X_testtrue_tpz)
print len(pred_annz)
#empty arrays for object sorting
stars_annz = np.empty(shape=(0,3))
gals_annz = np.empty(shape=(0,3))
uns_annz = np.empty(shape=(0,3))
#loop through and sort objects based on des object class
for i in range(len(ModestClass_annz)):
if ModestClass_annz[i] == 2:
stars_annz = np.append(stars_annz, [[pred_annz[i], DesZs_annz[i], y_test_annz[i]]], axis = 0)
elif ModestClass_annz[i] == 1:
gals_annz = np.append(gals_annz, [[pred_annz[i], DesZs_annz[i], y_test_annz[i]]], axis = 0)
else:
uns_annz = np.append(uns_annz, [[pred_annz[i], DesZs_annz[i], y_test_annz[i]]], axis = 0)
print len(stars_annz)
print len(gals_annz)
print len(uns_annz)
stars_bpz = np.empty(shape=(0,3))
gals_bpz = np.empty(shape=(0,3))
uns_bpz = np.empty(shape=(0,3))
for i in range(len(ModestClass_bpz)):
if ModestClass_bpz[i] == 2:
stars_bpz = np.append(stars_bpz, [[pred_bpz[i], DesZs_bpz[i], y_test_bpz[i]]], axis = 0)
elif ModestClass_bpz[i] == 1:
gals_bpz = np.append(gals_bpz, [[pred_bpz[i], DesZs_bpz[i], y_test_bpz[i]]], axis = 0)
else:
uns_bpz = np.append(uns_bpz, [[pred_bpz[i], DesZs_bpz[i], y_test_bpz[i]]], axis = 0)
stars_skynet = np.empty(shape=(0,3))
gals_skynet = np.empty(shape=(0,3))
uns_skynet = np.empty(shape=(0,3))
for i in range(len(ModestClass_skynet)):
if ModestClass_skynet[i] == 2:
stars_skynet = np.append(stars_skynet, [[pred_skynet[i], DesZs_skynet[i], y_test_skynet[i]]], axis = 0)
elif ModestClass_skynet[i] == 1:
gals_skynet = np.append(gals_skynet, [[pred_skynet[i], DesZs_skynet[i], y_test_skynet[i]]], axis = 0)
else:
uns_skynet = np.append(uns_skynet, [[pred_skynet[i], DesZs_skynet[i], y_test_skynet[i]]], axis = 0)
stars_tpz = np.empty(shape=(0,3))
gals_tpz = np.empty(shape=(0,3))
uns_tpz = np.empty(shape=(0,3))
for i in range(len(ModestClass_tpz)):
if ModestClass_tpz[i] == 2:
stars_tpz = np.append(stars_tpz, [[pred_tpz[i], DesZs_tpz[i], y_test_tpz[i]]], axis = 0)
elif ModestClass_tpz[i] == 1:
gals_tpz = np.append(gals_tpz, [[pred_tpz[i], DesZs_tpz[i], y_test_tpz[i]]], axis = 0)
else:
uns_tpz = np.append(uns_tpz, [[pred_tpz[i], DesZs_tpz[i], y_test_tpz[i]]], axis = 0)
plt.figure(figsize=(16,16))
plt.subplot(221)
plt.scatter(stars_annz.T[0], stars_annz.T[2], s=25, facecolor='none', edgecolor='blue')
plt.scatter(stars_annz.T[1], stars_annz.T[2], s=10, c='blue')
plt.scatter(gals_annz.T[0], gals_annz.T[2], s=25, facecolor='none', edgecolor='orange')
plt.scatter(gals_annz.T[1], gals_annz.T[2], s=10, c='orange')
legendhelp1_annz = plt.scatter(uns_annz.T[0], uns_annz.T[2], s=25, facecolor='none', edgecolor='k', label = 'NW photo-z')
legendhelp2_annz = plt.scatter(uns_annz.T[1], uns_annz.T[2], s=10, c='k', label = 'DES ANNZ photo-z')
plt.plot([0,1,2,3,4,5], 'r')
#plt.xlim(0,5)
#plt.ylim(0,5)
plt.xlabel('Photo-z Estimation')
plt.ylabel('z-spec')
plt.title('ANNZ')
orange_patch = mpatches.Patch(color='orange', label='DES Galaxy')
blue_patch = mpatches.Patch(color='blue', label='DES Star')
black_patch = mpatches.Patch(color='k', label='DES Undetermined')
plt.legend(handles=[blue_patch, orange_patch, black_patch, legendhelp1_annz, legendhelp2_annz])
plt.subplot(222)
plt.scatter(stars_bpz.T[0], stars_bpz.T[2], s=25, facecolor='none', edgecolor='blue')
plt.scatter(stars_bpz.T[1], stars_bpz.T[2], s=10, c='blue')
plt.scatter(gals_bpz.T[0], gals_bpz.T[2], s=25, facecolor='none', edgecolor='orange')
plt.scatter(gals_bpz.T[1], gals_bpz.T[2], s=10, c='orange')
legendhelp1_bpz = plt.scatter(uns_bpz.T[0], uns_bpz.T[2], s=25, facecolor='none', edgecolor='k', label = 'NW photo-z')
legendhelp2_bpz = plt.scatter(uns_bpz.T[1], uns_bpz.T[2], s=10, c='k', label = 'DES BPZ photo-z')
plt.plot([0,1,2,3,4,5], 'r')
plt.xlim(0,5)
plt.ylim(0,5)
plt.xlabel('Photo-z Estimation')
plt.ylabel('z-spec')
plt.title('BPZ')
plt.legend(handles=[blue_patch, orange_patch, black_patch, legendhelp1_bpz, legendhelp2_bpz])
plt.subplot(223)
plt.scatter(stars_skynet.T[0], stars_skynet.T[2], s=25, facecolor='none', edgecolor='blue')
plt.scatter(stars_skynet.T[1], stars_skynet.T[2], s=10, c='blue')
plt.scatter(gals_skynet.T[0], gals_skynet.T[2], s=25, facecolor='none', edgecolor='orange')
plt.scatter(gals_skynet.T[1], gals_skynet.T[2], s=10, c='orange')
legendhelp1_skynet = plt.scatter(uns_skynet.T[0], uns_skynet.T[2], s=25, facecolor='none', edgecolor='k', label = 'NW photo-z')
legendhelp2_skynet = plt.scatter(uns_skynet.T[1], uns_skynet.T[2], s=10, c='k', label = 'DES Skynet photo-z')
plt.plot([0,1,2,3,4,5], 'r')
plt.xlim(0,5)
plt.ylim(0,5)
plt.xlabel('Photo-z Estimation')
plt.ylabel('z-spec')
plt.title('Skynet')
plt.legend(handles=[blue_patch, orange_patch, black_patch, legendhelp1_skynet, legendhelp2_skynet])
plt.subplot(224)
plt.scatter(stars_tpz.T[0], stars_tpz.T[2], s=25, facecolor='none', edgecolor='blue')
plt.scatter(stars_tpz.T[1], stars_tpz.T[2], s=10, c='blue')
plt.scatter(gals_tpz.T[0], gals_tpz.T[2], s=25, facecolor='none', edgecolor='orange')
plt.scatter(gals_tpz.T[1], gals_tpz.T[2], s=10, c='orange')
legendhelp1_tpz = plt.scatter(uns_tpz.T[0], uns_tpz.T[2], s=25, facecolor='none', edgecolor='k', label = 'NW photo-z')
legendhelp2_tpz = plt.scatter(uns_tpz.T[1], uns_tpz.T[2], s=10, c='k', label = 'DES TPZ photo-z')
plt.plot([0,1,2,3,4,5], 'r')
plt.xlim(0,5)
plt.ylim(0,5)
plt.xlabel('Photo-z Estimation')
plt.ylabel('z-spec')
plt.title('TPZ')
plt.legend(handles=[blue_patch, orange_patch, black_patch, legendhelp1_tpz, legendhelp2_tpz])
```
Above plots show comparisons between the 4 DES photo-z methods and our Nadaraya-Watson method, both as compared to the spectroscopic redshift of the object. The open circles are the NW predictions and the closed points are DES's photometric redshift estimates. The red line represents the line along which photometric redshift equals spectroscopic redshift i.e. a "correct" photo-z estimate. This set of four plots is colored according to DES's object classification, though our test set here contains mostly objects classified by DES as stars i.e. point sources.
| github_jupyter |
```
import sys
sys.path.insert(0,"/home/nico/Documents/TEAR/Codes_TEAR/PythonCodes/LibFolder")
from Lib_GeneralFunctions import *
from Lib_GeneralSignalProcNAnalysis import *
from Lib_SigmoidProcessing import *
import pandas as pd
from matplotlib.gridspec import GridSpec
# Save into a class the
class SSCreference:
def __init__(self, filename, coordinates, RefSource="SEM2DPACK"):
line = pd.read_csv(filename.format("slip"), header=None)
self.Time = line[0]
self.Slip = line[1]
line = pd.read_csv(filename.format("sr"), header=None)
self.SlipRate = line[1]
self.Coord = coordinates #Only used for labels and printing
self.RefSource = RefSource
#end __init__
# Default object printing information
def __repr__(self):
return "The TPV3reference object was generated from: {} and the receiver is located at {}".format(self.RefSource, self.Coord)
#end __repr__
def __str__(self):
return "The TPV3reference object was generated from: {} and the receiver is located at {}".format(self.RefSource, self.Coord)
#end __str__
def PlotReference(self, ax, SlipSlipRate, filtering=True, **kwargs):
if SlipSlipRate=="Slip":
if(filtering):
ax.plot(self.Time, Butterworth(self.Slip, **kwargs), label = "", c = "k", ls = "--", zorder=1)
else:
ax.plot(self.Time, self.Slip, label = "", c = "k", ls = "--", zorder=1)
elif SlipSlipRate=="SlipRate":
if(filtering):
ax.plot(self.Time, Butterworth(self.SlipRate, **kwargs), label = "", c = "k", ls = "--", zorder=1)
else:
ax.plot(self.Time, self.SlipRate, label = "", c = "k", ls = "--", zorder=1)
return ax
def GenericFigAxis():
fig = plt.figure(figsize=[15,5])
gs = GridSpec(1, 2)
ax1 = fig.add_subplot(gs[0, 0])
ax2 = fig.add_subplot(gs[0, 1])
return fig, [ax1, ax2]
def format_axes(fig):
"""
Format a figure and 4 equidistant reveivers' lines from a single file. Receiver distance defines the color.
"""
for i, ax in enumerate(fig.axes):
ax.set_xlim(-0.5,4)
ax.set_ylim(-0.5,8)
ax.set_xlabel("time(s)")
Lines = fig.axes[-1].get_lines()
legend2 = fig.axes[-1].legend(Lines, ['2km','4km', '6km', '8km'], loc=1)
fig.axes[-1].add_artist(legend2)
fig.axes[-1].set_ylabel("Slip Rate (m/s)")
fig.axes[0].set_ylabel("Slip (m)")
def Multi_format_axes(fig,cmap, LabelsPerColor):
"""
Format a figure that contains different files with
information from several receivers for simulations under sets of blending parameters.
"""
ColorDict = dict(enumerate(LabelsPerColor))
for i, ax in enumerate(fig.axes):
ax.set_xlim(-0.5,4)
ax.set_ylim(-0.5,8)
ax.set_xlabel("time(s)")
Lines = []
for idx,colcol in enumerate(cmap.colors):
Lines.append(mlines.Line2D([], [], color = colcol,
linewidth = 3, label = ColorDict.get(idx)))
legend2 = fig.axes[-1].legend(Lines, LabelsPerColor, loc = 2)
fig.axes[-1].add_artist(legend2)
fig.axes[-1].set_ylabel("Slip Rate (m/s)")
fig.axes[0].set_ylabel("Slip (m)")
path = "/home/nico/Documents/TEAR/Codes_TEAR/ProfilePicking/Output/"
# Reference saved into a list of objects
RefList = [SSCreference(path + "Reference/sem2dpack/sem2d-{}-1.txt", "2km"),
SSCreference(path + "Reference/sem2dpack/sem2d-{}-2.txt", "4km"),
SSCreference(path + "Reference/sem2dpack/sem2d-{}-3.txt", "6km"),
SSCreference(path + "Reference/sem2dpack/sem2d-{}-4.txt", "8km"),
]
from matplotlib.colors import ListedColormap
import matplotlib.lines as mlines
from palettable.scientific.sequential import Oslo_4
cmap = ListedColormap(Oslo_4.mpl_colors[:])
FolderSigmoidPath = "/home/nico/Documents/TEAR/Codes_TEAR/PythonCodes/[SSC]Sigmoid/ProcessedData/"
ListOfFileNames = ["20210204-T4-50x50-P1-100.05",
"20210204-T5-25x25-P1-50.025",
"20210215-T6-12.5x12.5-P1-25.0125",
"20210204-T7-50x50-P1-50.05",
"20210204-T8-25x25-P1-25.025",
"20210215-T9-12.5x12.5-P1-12.5125",]
Delta2 = ListOfFileNames[:3]
Delta1 = ListOfFileNames[3:6]
SigmoidFiles2 = [LoadPickleFile(FolderSigmoidPath, fname) for fname in Delta2]
SigmoidFiles1 = [LoadPickleFile(FolderSigmoidPath, fname) for fname in Delta1]
fig, axis = GenericFigAxis()
# Sigmoid case plotting
for iidx,SFile in enumerate(SigmoidFiles2):
print(SFile)
for Test1 in SFile:
axis[0].plot(Test1.Time, Test1.Slip, color= cmap.colors[iidx],linewidth=2,zorder=iidx)
axis[1].plot(Test1.Time, Test1.SlipRate, color= cmap.colors[iidx],linewidth=2,zorder=iidx)
LabelsPerColor= ["50x50-P1-$\delta$:100.05", "25x25-P1-$\delta$:50.025","12.5x12.5-P1-$\delta$:25.012"]
Multi_format_axes(fig, cmap, LabelsPerColor)
fig.suptitle("P1 - No blending - $\delta_f: 2.001$")
[item.PlotReference(axis[0], "Slip", filtering=False) for item in RefList]
[item.PlotReference(axis[1], "SlipRate", filtering=False) for item in RefList]
##############################################
fig, axis = GenericFigAxis()
# Sigmoid case plotting
for iidx,SFile in enumerate(SigmoidFiles1):
for Test1 in SFile:
axis[0].plot(Test1.Time, Test1.Slip, color= cmap.colors[iidx],linewidth=2,zorder=iidx)
axis[1].plot(Test1.Time, Test1.SlipRate, color= cmap.colors[iidx],linewidth=2,zorder=iidx)
LabelsPerColor= ["50x50-P1-$\delta$:50.05", "25x25-P1-$\delta$:25.025","12.5x12.5-P1-$\delta$:12.512"]
Multi_format_axes(fig, cmap, LabelsPerColor)
fig.suptitle("P1 - No blending - $\delta_f: 1.001$")
[item.PlotReference(axis[0], "Slip", filtering=False) for item in RefList]
[item.PlotReference(axis[1], "SlipRate", filtering=False) for item in RefList]
from palettable.scientific.sequential import Oslo_3
cmap = ListedColormap(Oslo_3.mpl_colors[:])
FolderSigmoidPath = "/home/nico/Documents/TEAR/Codes_TEAR/PythonCodes/[SSC]Sigmoid/ProcessedData/"
FolderTiltedPath = "/home/nico/Documents/TEAR/Codes_TEAR/PythonCodes/[SSC]Sigmoid/ProcessedData/20210203-Tilting/"
SigmoidFile = LoadPickleFile(FolderSigmoidPath, "20210215-T3-25x25-P3-12000FaultPoints50.025")
TiltedFile = LoadPickleFile(Filename = "TPList_t8180_d50.025.pickle",FolderPath = FolderTiltedPath)
fig, axis = GenericFigAxis()
# Sigmoid case plotting
iidx = 0
for Test1 in SigmoidFile:
axis[0].plot(Test1.Time, Test1.Slip, color= cmap.colors[iidx],linewidth=2,zorder=iidx)
axis[1].plot(Test1.Time, Test1.SlipRate, color= cmap.colors[iidx],linewidth=2,zorder=iidx)
# Tilted case plotting
iidx = 1
for Test1 in TiltedFile[:-1]:
axis[0].plot(Test1.Time, Test1.DispX, color= cmap.colors[iidx],linewidth=2,zorder=iidx)
axis[1].plot(Test1.Time, Test1.VelX, color= cmap.colors[iidx],linewidth=2,zorder=iidx)
LabelsPerColor= ["Sigmoid","Tilted"]
Multi_format_axes(fig, cmap, LabelsPerColor)
fig.suptitle("Cell Dims:25x25 - P3 - $\delta$50.025 \nA:4.0p/$\delta$ , $\phi_o$:0.65$\delta$")
[item.PlotReference(axis[0], "Slip", filtering=False) for item in RefList]
[item.PlotReference(axis[1], "SlipRate", filtering=False) for item in RefList]
```
# Update Sigmoid vs tilting
# Constant fault zone
```
FolderSigmoidPath = "/home/nico/Documents/TEAR/Codes_TEAR/PythonCodes/[SSC]Sigmoid/ProcessedData/"
ListOfFileNames = ["20210204-T4-50x50-P1-100.05",
"20210216-T4-25x25-P3-100.03"]
Delta2 = ListOfFileNames
SigmoidFiles2 = [LoadPickleFile(FolderSigmoidPath, fname) for fname in Delta2]
fig, axis = GenericFigAxis()
# Sigmoid case plotting
for iidx,SFile in enumerate(SigmoidFiles2):
print(SFile)
for Test1 in SFile:
axis[0].plot(Test1.Time, Test1.Slip, color= cmap.colors[iidx],linewidth=2,zorder=iidx)
axis[1].plot(Test1.Time, Test1.SlipRate, color= cmap.colors[iidx],linewidth=2,zorder=iidx)
LabelsPerColor= ["50x50-P1-$\delta_f$:2.001", "25x25-P1-$\delta_f$:4.001"]
Multi_format_axes(fig, cmap, LabelsPerColor)
fig.suptitle("P1 - No blending - $\delta: 100.$")
[item.PlotReference(axis[0], "Slip", filtering=False) for item in RefList]
[item.PlotReference(axis[1], "SlipRate", filtering=False) for item in RefList]
from palettable.scientific.sequential import Oslo_5
cmap = ListedColormap(Oslo_5.mpl_colors[:])
FolderSigmoidPath = "/home/nico/Documents/TEAR/Codes_TEAR/PythonCodes/[SSC]Sigmoid/ProcessedData/"
ListOfFileNames = ["20210218-T1-25x25-P3-37.525",
"20210218-T2-25x25-P3-50.025",
"20210218-T4-25x25-P3-62.525",
"20210218-T6-25x25-P3-75.02499999999999"]
Delta2 = ListOfFileNames
SigmoidFiles2 = [LoadPickleFile(FolderSigmoidPath, fname) for fname in Delta2]
fig, axis = GenericFigAxis()
# Sigmoid case plotting
for iidx,SFile in enumerate(SigmoidFiles2):
print(SFile)
for Test1 in SFile:
axis[0].plot(Test1.Time, Test1.Slip, color= cmap.colors[iidx],linewidth=2,zorder=iidx)
axis[1].plot(Test1.Time, Test1.SlipRate, color= cmap.colors[iidx],linewidth=2,zorder=iidx)
LabelsPerColor= ["$\delta_f$:1.501","$\delta_f$:2.001","$\delta_f$:2.501","$\delta_f$:3.001"]
Multi_format_axes(fig, cmap, LabelsPerColor)
fig.suptitle("P1 - No blending - 25x25, ")
[item.PlotReference(axis[0], "Slip", filtering=False) for item in RefList]
[item.PlotReference(axis[1], "SlipRate", filtering=False) for item in RefList]
FolderSigmoidPath = "/home/nico/Documents/TEAR/Codes_TEAR/PythonCodes/[SSC]Sigmoid/ProcessedData/"
ListOfFileNames = ["20210218-T1-25x25-P3-37.525",
"20210218-T2-25x25-P3-50.025",
"20210218-T4-25x25-P3-62.525"]
Delta2 = ListOfFileNames
SigmoidFiles2 = [LoadPickleFile(FolderSigmoidPath, fname) for fname in Delta2]
fig, axis = GenericFigAxis()
# Sigmoid case plotting
for iidx,SFile in enumerate(SigmoidFiles2):
print(SFile)
for Test1 in SFile:
axis[0].plot(Test1.Time, Test1.Slip, color= cmap.colors[iidx],linewidth=2,zorder=iidx)
axis[1].plot(Test1.Time, Test1.SlipRate, color= cmap.colors[iidx],linewidth=2,zorder=iidx)
LabelsPerColor= ["$\delta_f$:1.501","$\delta_f$:2.001","$\delta_f$:2.501"]
Multi_format_axes(fig, cmap, LabelsPerColor)
fig.suptitle("P1 - No blending - 25x25, ")
[item.PlotReference(axis[0], "Slip", filtering=False) for item in RefList]
[item.PlotReference(axis[1], "SlipRate", filtering=False) for item in RefList]
```
| github_jupyter |
```
%matplotlib inline
```
# TP1: The Interaction-driven Metal-Insulator Transition
## GOAL:
1) To understand the difference between the electronic spectral functions of a metal and of a Mott insulator.
2) To understand the interaction-driven metal-insulator transition in the Hubbard Model (the reference model for this problem), known as the **Mott Metal-Insulator Transition** (MIT).
To achieve these goals you will have to understand how to read Green's function and connect them to physical properties. We will use the (exact!) Dynamical-Mean-Field-Theory (DMFT) solution of the Hubbard Model on the infinite-dimensional Bethe lattice. The DMFT is numerically implemented using the Iterative Perturbation Theory (IPT), which provides (approximate but good) Green's functions. To this purpose you will have to run a PYTHON code, manipulating the input/output in order to simulate different physical situations and interpret the outputs.
It would be interesting to study also the details of the DMFT implementation and the IPT impurity solver, but it will not be possible on this occasion. Use it then as a black box to perform virtual theoretical experiments.
You will have to provide us at the end of the Tutorial a complete report, answering to the questions of the section EXERCISES. Include graphs and commentaries whenever asked (or possible) at your best convenience.
Real frequency IPT solver single band Hubbard model
===================================================
Here it is the IPT code.
```
# Author: Óscar Nájera
# License: 3-clause BSD
from __future__ import division, absolute_import, print_function
import scipy.signal as signal
import numpy as np
import matplotlib.pyplot as plt
plt.matplotlib.rcParams.update({'axes.labelsize': 22,
'axes.titlesize': 22, })
def fermi_dist(energy, beta):
""" Fermi Dirac distribution"""
exponent = np.asarray(beta * energy).clip(-600, 600)
return 1. / (np.exp(exponent) + 1)
def semi_circle_hiltrans(zeta, D=1):
"""Calculate the Hilbert transform with a semicircular DOS """
sqr = np.sqrt(zeta**2 - D**2)
sqr = np.sign(sqr.imag) * sqr
return 2 * (zeta - sqr) / D**2
def pertth(Aw, nf, U):
"""Imaginary part of the second order diagram"""
# because of ph and half-fill in the Single band one can work with
# A^+ only
Ap = Aw * nf
# convolution A^+ * A^+
App = signal.fftconvolve(Ap, Ap, mode='same')
# convolution A^-(-w) * App
Appp = signal.fftconvolve(Ap, App, mode='same')
return -np.pi * U**2 * (Appp + Appp[::-1])
def dmft_loop(gloc, w, U, beta, loops):
"""DMFT Loop for the single band Hubbard Model at Half-Filling
Parameters
----------
gloc : complex 1D ndarray
local Green's function to use as seed
w : real 1D ndarray
real frequency points
U : float
On site interaction, Hubbard U
beta : float
Inverse temperature
loops : int
Amount of DMFT loops to perform
Returns
-------
gloc : complex 1D ndarray
DMFT iterated local Green's function
sigma : complex 1D ndarray
DMFT iterated self-energy
"""
dw = w[1] - w[0]
eta = 2j * dw
nf = fermi_dist(w, beta)
for i in range(loops):
# Self-consistency
g0 = 1 / (w + eta - .25 * gloc)
# Spectral-function of Weiss field
A0 = -g0.imag / np.pi
# Clean for PH and Half-fill
A0 = 0.5 * (A0 + A0[::-1])
# Second order diagram
isi = pertth(A0, nf, U) * dw * dw
# Kramers-Kronig relation, uses Fourier Transform to speed convolution
hsi = -signal.hilbert(isi, len(isi) * 4)[:len(isi)].imag
sigma = hsi + 1j * isi
# Semi-circle Hilbert Transform
gloc = semi_circle_hiltrans(w - sigma)
return gloc, sigma
```
## Example Simulation
```
w = np.linspace(-6, 6, 2**12)
gloc = semi_circle_hiltrans(w + 1e-3j)
#gloc= 0.0
gloc, sigma_loc = dmft_loop(gloc, w, 0.3, 100, 100)
#plt.plot(w, -gloc.imag, lw=3, label=r'$\pi A(\omega)$')
plt.plot(w, sigma_loc.real, '-', label=r'$Re \Sigma(\omega)$')
plt.plot(w, sigma_loc.imag, '-', label=r'$Im \Sigma(\omega)$')
plt.xlabel(r'$\omega$')
plt.legend(loc=0)
#plt.ylim([0, 0.1])
#plt.xlim([-1, 1])
plt.show()
```
## The effect of the self-consistency condition
1. The DMFT implementation is an iterative process which requires a certain number of loops before convergence is reached. But one needs a starting point, i.e. one needs to input an initial guess of the spectral function. Can you say which is the starting guess in the DMFT implementation considered above?
2. For $U < Uc_2$, run the code with *just one dmft iteration*, using the same values of $U$ that you have employed above. This one-loop run corresponds to the solution of the Single Impurity Anderson Model SIAM (or **Kondo Model**, see Marc's lecture notes) for a semi-circular conduction band. Compare the spectral function $A(\omega)$ of the one-iteration loop with the fully converged one. Do you see the MIT?
3. Repeat the same excercise of point 2., this time inputing no bath (i.e. the intial guess is zero), and always one iteration loop. This means that you are essentially solving the isolated atom problem with on-site interaction $U$. Draw a conclusions about the effect of the self-conistency in solving the MIT problem.
# Let's start now
## Excercises
We shall first work at low temperature, choose $\beta=100$. The imaginary part of the Green's function $A(\omega)= -\Im G(\omega)/\pi$ can be directly connected to physical observables (e.g. spectral functions) or transport (e.g. specific heat).
## The Metal-to-Insulator Transition
1. Plot first $A(\omega)$ in the case $U=0$. This is the semicircular density of states used in the DMFT code, i.e. $-\Im G(\omega)/\pi= D(\omega)$. The system is in this case "halffilled", i.e. there is one elctron per atomic orbital. Can you indicate then where the occupied states are on the plot of the density of states $A(\omega)$? Why the system is a metal according to the band theory of solids?
2. Run the code for several values of $U$ (the half-bandwidth is set $D=1$) to check out the metal-insulator transition. Display $A(\omega)$ for some rapresentative values of $U$, in the metallic and insulating sides.
3. Approach the MIT at $Uc_2$ (metal to insulator) from below ($U< Uc_2$) and describe how the different contributions to the $A(\omega)$ (quasiparticle peak and Hubbard bands) evolve as you get close to the critical value. Why can we state that for $U> Uc_2$ the system is finally insulating? Give your extimation of $Uc_2$.
4. At the MIT point, the quaiparticle peak has completely disappered. But where is the spectral weight of the peak gone? Explain why we expect that the total spectral weight [i.e. the area under the curve $A(\omega)$] is conserved.
## Selfenergy
1. Plot now the real part and the imaginary part of the self-energy $\Sigma(\omega)$, first for $U< Uc_2$, then for $U \ge Uc_2$. Use the same values of $U$ that you have used above.
2. We are going now to concentrate on the low energy part $\omega\to 0$, where a quasiparticle peak is observable in the metallic phase, even for values of $U$ very close to the MIT. The Fermi liquid theory of metals states that the selfenergy is a regular function of $\omega$, which can be Taylor expanded: $$
\Re\Sigma \simeq \mu_0 + \alpha \omega+ O(\omega^2) \quad \quad
\Im\Sigma \sim \omega^2+ O(\omega^3)
$$
For which values of $U$ can we state that the system is in a Fermi liquid state?
3. We shall now see how the Fermi liquid theory may be useful to understand the beahvior of spectral functions. The local Green's function (entering the DMFT self-consistency condition) is the Hilbert transform of the lattice Green's function $G(\omega,\varepsilon)$
$$
G_{loc}(\omega)= \,\int D(\varepsilon) \, G(\omega, \varepsilon)\, d\varepsilon=
\,\int D(\varepsilon)/\left[ \omega-\varepsilon- \Sigma(\omega) \right] \, d\varepsilon $$
where $D(\varepsilon)= -(1/\pi) \Im G_{loc}(\varepsilon,U=0)$ and in our case (infinite dimension) $\Sigma(\omega)$ is independent of $\varepsilon$. Show that the lattice Green's function can be written within the Fermi liquid theory ($\omega\to0$) as a almost free-particle Green's function $$
G(\omega, \xi)\sim \frac{Z}{\omega-\xi} $$
where the main difference is the factor $Z$, known as *quasi-particle renormalization factor* .
* Evaluate $Z$ for different values of $U$ (use the real part of the self-energy) and plot it as a function of $U$. Explain why $Z$ is useful to describe the MIT. *Hint: in order to extract a part of the Green's function array (let's call it e.g. $gx$) between frequencies, e.g. $[-2,2]$, define a logical array of frequencies $w_r= (w< 2 )*(w> -2)$, then the desired reduced Green's function array is simply given by $gx[w_r]$.*
* Calculate (for a couple of $U$ values used above) the area under the quasiparticle peaks and plot them as a function of $U$. Relate this area with the quasiparticle residue $Z$ (you should plot them as a function of $U$).
* Determine the exact relation between the quasiparticle peak and $Z$ also analytically. Explain finally why it is said that *a Mott transition is a breaking of the Fermi Liquid Theory*.
# First order transition
The MIT is a first order transiton, i.e. there is a region of coexistence between the insulator and the metallic phase.
1. Show that there is an interval of $U$ where DMFT self-consistency provides two solutions. Determine the $Uc_1$, i.e. the critical value of $U$ where the insulating state changes into a metal by reducing $U$. *(Hint: you have to use an appropriate starting guess to obtain the insulating solution, start your investigations with values of $U$ in the insulating side).*
2. Is $Z$ the right parameter to describe the insulator-to-metal transition too? Argument your answer.
# $T-U \,$ phase diagram and the $T$-driven transition
1. Run now the code for higher temperatures $T$, i.e. lower inverse temperatures $\beta= 75, 50, 25$ and determine for each temperature $T$ the $Uc_1$ and the $Uc_2$. Sketch then the coexistence region in the $T-U$ plane.
2. Observe the shape of the coexistence region in the $T-U$ space. Fix the interaction $U$ inside this co-existence region (e.g. $U=2.9$) and systematically increase the temperature. Compare the different density of states. Can you now explain the phase-diagram of VO$_2$, in particular the unusual not-metallic behavior at high temperature?
3. Can you give an interpretation of this physical behaviour with temperature in terms of the associated SIAM model, i.e. a spin impurity fluctuating with temperature in a bath of free electrons?
| github_jupyter |
# VGP 245 Assignment 2
Question 1: Implement a function that sums a list of tuple paired numbers i.e coordinates (x,y) i.e. vector2 and sums it together into a single tuple. i.e. vector2Add([(1,1), (2,3), (3,3), (5,10)]) is (11,17)
```
# a.
def vector2Sum(list):
a = 0
b = 0
for e in list:
a = a + int(e[0])
b = b + int(e[1])
return (a, b)
list1 = [(1,1), (2,3), (3,3), (5,10)]
finalVector = vector2Sum(list1)
print (finalVector)
```
b. What can go wrong with the function? How do you check that the value in the parameter is correct?
A: It will try to sum the value regardless of it's type, since the tuple can have elements of every type. To fix this, we can do a try-catch to make sure the element is from the proper type, otherwise it will ignore the value.
Question 2: Implement a function that checks if the age specified is 18 or above, and print a message whether or not you can vote, and if the age is above 65, return a special friendly message.
```
# i. can_vote(18) prints the message "Don't forget to vote on Oct. 20th."
# ii. can_vote(10) prints you have "8 more years before you can vote."
# iii. can_vote(65) prints "Don't forget to vote on Oct. 20th. You're strong and awesome!
def check_age(value):
if(value > 18 and value < 65):
print("Don't forget to vote on Oct. 20th.")
elif (value > 65):
print("Don't forget to vote on Oct. 20th. You're strong and awesome!")
else:
print("You have a few more years before you can vote.")
age = int(input("Please enter your age: "))
check_age(age)
```
Question 3. Implement a increment function that takes a number and increments it by 1 using a default argument, so that user can replace that increment unit to with a different number. Use that increment function in a for loop to show at least 10 increments for each case.
```
# i. increment(3) returns 3
# ii. increment(2, 2) returns 4
def increment_function(value, other_value = 1):
value = value + other_value
print(value)
return value
increment_function(2)
increment_function(2, 3)
print('-'*10)
print("\n")
count = 0
value = 3
print ("Initial value: ", value)
while count != 10:
print("Count = ", count)
value = increment_function(value, 4)
count += 1
```
Question 4: Create sets using the of the following information:
SSJ: Goku, Vegeta, Gohan, Gohan, Trunks, Broly
Sayian: Goku, Vegeta, Pan, Nappa, Gohan, Trunks, Broly
Human: Krillian, Yamcha, Tien, Chichi, Bulma, Gohan, Trunks, Pan
Finding the differences and intersects between the sets using the set methods:
```
# Creating the Sets:
SSJ = set(["Goku", "Vegeta", "Gohan", "Goten", "Trunks", "Broly"])
Sayian = set([ "Goku", "Vegeta", "Pan", "Nappa", "Gohan", "Trunks", "Broly"])
Human = set(["Krillin", "Yamcha", "Tien", "Chichi", "Bulma", "Gohan", "Trunks", "Pan"])
# a. Find the elements that are only in "SSJ" set and not in "Sayian" set
print (SSJ.difference(Sayian))
# b. Find the elements that are in the "Human" set and "Sayian" set
print(Human.intersection(Sayian))
# c. Find the elements that are in the "SSJ" set and "Human" set
print(SSJ.intersection(Human))
# d. Find the elements that are in both "Sayian" set and the "Human" set.
print(Sayian.intersection(Human))
```
Question 5: What is the type return from the execution of the following functions:
```
def sum(left, right):
return left + right
print(sum(3,4))
print(sum(1.1, 2.1))
print(sum('kame', 'hameha'))
def print_greetings(name):
print(f'Welcome {name} to Namek, have a nice day!')
print_greetings('Master Roshi')
def capitalize(message):
return message.upper()
capitalize("It's over 9000!")
# a. sum(3,4)
# b. sum(1.1, 2.1)
# c. sum('kame', 'hameha')
# d. print_greetings('Master Roshi')
# e. capitalize("It's over 9000!")
```
Question 6: What is tuple unpacking and show an example of it.
```
# Tuple unpacking is also called multiple assignment. It's the association of values to each element of the tuple.
x, y = 5, 10
# or
(x, y) = (5, 10)
```
Question 7: Make the following function throw an exception when the count is 0
```
senzu_bean_count = 2
# a. implement throw / raise exception
def eat_senzu_bean():
senzu_bean_count -= 1
print('consuming senzu bean...')
raise Exception()
try:
eat_senzu_bean()
except Exception:
print("No more beans!!")
finally:
print("You finished the meal.")
# b. Handle the exception, so that it shows a useful error and continues gracefully
```
Question 8: What is finally used for?
The keyword "finally" makes sure the following section will always be called. It's used to terminate a process that hang-up in an error
Question 9: How do you create an empty set?
s = set()
Question 10: What is the difference between a set and a dictionary? and what are their similarities?
A set is like a dict with keys but no values, and they're both implemented using a hash table. Both cannot accept repeated keys, but the set can work without associated values.
| github_jupyter |
```
from pandas_datareader import data
from matplotlib import pyplot as plt
import pandas as pd
import datetime
import numpy as np
import pandas_datareader
print pandas_datareader.__version__
# Define the instruments to download. We would like to see Apple, Microsoft and others.
companies_dict = {
'Amazon': 'AMZN',
'Apple': 'AAPL',
'Walgreen': 'WBA',
'Northrop Grumman': 'NOC',
'Boeing': 'BA',
'Lockheed Martin': 'LMT',
'McDonalds': 'MCD',
'Intel': 'INTC',
'Navistar': 'NAV',
'IBM': 'IBM',
'Texas Instruments': 'TXN',
'MasterCard': 'MA',
'Microsoft': 'MSFT',
'General Electrics': 'GE',
'Symantec': 'SYMC',
'American Express': 'AXP',
'Pepsi': 'PEP',
'Coca Cola': 'KO',
'Johnson & Johnson': 'JNJ',
'Toyota': 'TM',
'Honda': 'HMC',
'Mitsubishi': 'MSBHY',
'Sony': 'SNE',
'Exxon': 'XOM',
'Chevron': 'CVX',
'Valero Energy': 'VLO',
'Ford': 'F',
'Bank of America': 'BAC'}
companies = sorted(companies_dict.items(), key=lambda x: x[1])
#print(companies)
# Define which online source one should use
data_source = 'morningstar'
# Define the start and end dates that we want to see
start_date = '2015-01-01'
end_date = '2017-12-31'
# User pandas_reader.data.DataReader to load the desired data. As simple as that.
panel_data = data.DataReader(companies_dict.values(), data_source, start_date, end_date).unstack(level=0)
# Print Axes Labels
print(panel_data.axes)
# Find Stock Open and Close Values
stock_close = panel_data['Close']
stock_open = panel_data['Open']
print(stock_close.iloc[0])
#print(stock_open.iloc[0])
# Calculate daily stock movement
stock_close = np.array(stock_close).T
stock_open = np.array(stock_open).T
row, col = stock_close.shape
movements = np.zeros([row, col])
for i in range(0, row):
movements[i,:] = np.subtract(stock_close[i,:], stock_open[i,:])
for i in range(0, len(companies)):
print('Company: {}, Change: {}'.format(companies[i][0], sum(movements[i][:])))
# print(movements.shape)
# Import Normalizer
from sklearn.preprocessing import Normalizer
from sklearn.pipeline import make_pipeline
from sklearn.cluster import KMeans
# Create a normalizer: normalizer
normalizer = Normalizer()
new = normalizer.fit_transform(movements)
print(new.max())
print(new.min())
print(new.mean())
# Normalizer for use in pipeline
normalizer = Normalizer()
# Create a KMeans model with 10 clusters: kmeans
kmeans = KMeans(n_clusters=10, max_iter=1000)
# Make a pipeline chaining normalizer and kmeans: pipeline
pipeline = make_pipeline(normalizer, kmeans)
# Fit pipeline to the daily price movements
pipeline.fit(movements)
print(kmeans.inertia_)
# Import pandas
import pandas as pd
# Predict the cluster labels: labels
labels = pipeline.predict(movements)
# Create a DataFrame aligning labels and companies: df
df = pd.DataFrame({'labels': labels, 'companies': companies})
# Display df sorted by cluster label
print(df.sort_values('labels'))
# Visualization - Plot Stock Movements
plt.clf
plt.figure(figsize=(18, 16))
ax1 = plt.subplot(221)
plt.plot(new[19][:])
plt.title(companies[19])
plt.subplot(222, sharey=ax1)
plt.plot(new[13][:])
plt.title(companies[13])
plt.show()
from sklearn.decomposition import PCA
# Visualize the results on PCA-reduced data
# Principal component analysis (PCA)
# Linear dimensionality reduction using Singular Value Decomposition of the data to project it to a lower dimensional space
reduced_data = PCA(n_components=2).fit_transform(new)
kmeans = KMeans(init='k-means++', n_clusters=10, n_init=10)
kmeans.fit(reduced_data)
labels = kmeans.predict(reduced_data)
# Create a DataFrame aligning labels and companies: df
df = pd.DataFrame({'labels': labels, 'companies': companies})
# Display df sorted by cluster label
print(df.sort_values('labels'))
# Step size of the mesh. Decrease to increase the quality of the VQ.
h = .01 # point in the mesh [x_min, x_max]x[y_min, y_max].
# Plot the decision boundary. For that, we will assign a color to each
x_min, x_max = reduced_data[:, 0].min() - 1, reduced_data[:, 0].max() + 1
y_min, y_max = reduced_data[:, 1].min() - 1, reduced_data[:, 1].max() + 1
xx, yy = np.meshgrid(np.arange(x_min, x_max, h), np.arange(y_min, y_max, h))
# Obtain labels for each point in mesh. Use last trained model.
Z = kmeans.predict(np.c_[xx.ravel(), yy.ravel()])
# Put the result into a color plot
Z = Z.reshape(xx.shape)
# Define Colormap
cmap = plt.cm.Paired
plt.figure(figsize=(10, 10))
plt.clf()
plt.imshow(Z, interpolation='nearest',
extent=(xx.min(), xx.max(), yy.min(), yy.max()),
cmap=cmap,
aspect='auto', origin='lower')
plt.plot(reduced_data[:, 0], reduced_data[:, 1], 'k.', markersize=5)
# Plot the centroids as a white X
centroids = kmeans.cluster_centers_
plt.scatter(centroids[:, 0], centroids[:, 1],
marker='x', s=169, linewidths=3,
color='w', zorder=10)
plt.title('K-means clustering on Stock Market Movements (PCA-reduced data)\n'
'Centroids are marked with white cross')
plt.xlim(x_min, x_max)
plt.ylim(y_min, y_max)
plt.show()
```
| github_jupyter |
# Strain stratification
Summer provides a special disease-strain stratification class ([StrainStratification](http://summerepi.com/api/stratification.html#summer.stratification.StrainStratification)) that treats each stratum as a separate strain of the infection. This allows you to model multiple strains of an infection (or multiple infectious diseases with similar compartmental structures) which are able to infect people separately.
They key difference between StrainStratification vs. a normal Stratification is that the strain stratification adjusts the count of infectious people per strain. For example, in a normal stratification with an age-based split into 'young' and 'old' you will have frequency-dependent infection flows calculated as follows:
```python
# Find a common force of infection for both young and old
num_infected = num_old_infected + num_young_infected
force_of_infection = contact_rate * num_infected / num_pop
# Use that common force of infection to get flow rates for young/old infection
infect_rate_young = force_of_infection * num_young_susceptible
infect_rate_old = force_of_infection * num_old_susceptible
```
Consider now how this will be calculated for two strains ("mild" and "wild") when applied to an unstratified susceptible compartment:
```python
# Find a different force of infection for mild and wild
force_of_infection_mild = contact_rate * num_infected_mild / num_pop
force_of_infection_wild = contact_rate * num_infected_wild / num_pop
# Use the different force of infection values to get flow rates for mild/wild infection
infect_rate_mild = force_of_infection_mild * num_susceptible
infect_rate_wild = force_of_infection_wild * num_susceptible
```
Let's work through a code example. For starters, let's create an SIR model:
```
import numpy as np
import matplotlib.pyplot as plt
from summer import CompartmentalModel
def build_model():
"""Returns a model for the stratification examples"""
model = CompartmentalModel(
times=[1990, 2010],
compartments=["S", "I", "R"],
infectious_compartments=["I"],
timestep=0.1,
)
# Add people to the model
model.set_initial_population(distribution={"S": 990, "I": 10})
# Susceptible people can get infected.
model.add_infection_frequency_flow(name="infection", contact_rate=2, source="S", dest="I")
# Infectious people take 3 years, on average, to recover.
model.add_transition_flow(name="recovery", fractional_rate=1/3, source="I", dest="R")
# Add an infection-specific death flow to the I compartment.
model.add_death_flow(name="infection_death", death_rate=0.05, source="I")
return model
def plot_compartments(model):
"""Plot model compartment sizes over time"""
fig, ax = plt.subplots(1, 1, figsize=(12, 6), dpi=120)
for i in range(model.outputs.shape[1]):
ax.plot(model.times, model.outputs.T[i])
ax.set_title("SIR Model Outputs")
ax.set_xlabel("Year")
ax.set_ylabel("Compartment size")
start, end = ax.get_xlim()
ax.xaxis.set_ticks(np.arange(start + 1, end, 5))
ax.legend([str(c) for c in model.compartments], loc='upper right')
plt.show()
print('Compartment names:', model.compartments)
print('Initial values:', model.outputs[0])
print('Final values:', model.outputs[-1])
# Force NumPy to format output arrays nicely.
np.set_printoptions(formatter={'all': lambda f: f"{f:0.2f}"})
```
Lets see what this model looks like without any stratifications:
```
model = build_model()
model.run()
plot_compartments(model)
```
Now we can add a strain stratification to the infected (I) and recovered (R) compartments. We will assume immunity to one strain gives you immunity to the other.
```
from summer import StrainStratification
strata = ['mild', 'wild']
strat = StrainStratification(name="strain", strata=strata, compartments=['I', 'R'])
# At the start of the simulation, 20% of infected people have wild strain.
strat.set_population_split({'mild': 0.8, 'wild': 0.2})
model = build_model()
model.stratify_with(strat)
model.run()
plot_compartments(model)
```
Note that despite the stratification, the model results are the same in aggregate, because we have not applied any adjustments to the flows or strata infectiousness yet. Let's do that:
```
from summer import StrainStratification, Multiply
strata = ['mild', 'wild']
strat = StrainStratification(name="strain", strata=strata, compartments=['I', 'R'])
# Again, 20% of infected people have wild strain at the start.
strat.set_population_split({'mild': 0.8, 'wild': 0.2})
# The wild strain kills at 1.2x the rate as the mild strain does.
strat.set_flow_adjustments("infection_death", {
"mild": None,
"wild": Multiply(1.2),
})
# Wild strain is twice as infectious than the mild strain (or equivalently, people are twice as susceptible to it).
strat.set_flow_adjustments("infection", {
"mild": None,
"wild": Multiply(2),
})
model = build_model()
model.stratify_with(strat)
model.run()
plot_compartments(model)
```
| github_jupyter |
# Example
This page was rendered from a Jupyter notebook found [here](https://github.com/benlindsay/pbcluster/docs/example.ipynb).
In this example, we will analyze particle clustering by loading a `.gro` trajectory file (see [here](http://manual.gromacs.org/archive/5.0.3/online/gro.html) for details about this format), then passing that trajectory data to PBCluster. Your trajectories can come from wherever you want, but it's your responsibility to get the particle coordinate data out of them to pass into PBCluster. That data needs to either be in the form of a 3D numpy array with shape `(n_timesteps, n_particles, n_dimensions)`, or a pandas dataframe with columns for `timestep`, `particle_id`, `x0`, `x1`, and `x2` (assuming a 3D simulation box).
Here's a visualization made with [Ovito](https://ovito.org/) of the particle trajectory we'll be analyzing in this example:

As the simulation progresses from timestep 0 to 39, the particles start dispersed and aggregate into a single cluster.
At the end of this trajectory, it looks like there is a large cluster on the bottom and a separate, small cluster on top, but those are both part of the same cluster.
Since this box has periodic boundaries (i.e. particles wrap around to the other side of the box if they pass through a face), the cluster wraps across the top and bottom faces of the box.
## Import Libraries
```
import mdtraj # for loading .gro data
import numpy as np
import pandas as pd
import plotnine as pn # for ggplot2-style plotting
from pbcluster import Trajectory
```
## Load `.gro` data
We need the particle coordinates and the simulation box size data.
```
traj_data = mdtraj.load("traj.gro")
# The factor of 10 is because .gro files store coordinates in Angstroms instead of nanometers
particle_coords = traj_data.xyz * 10
n_iterations, n_particles, n_dimensions = particle_coords.shape
print(f"n_iterations: {n_iterations}")
print(f"n_particles: {n_particles}")
print(f"n_dimensions: {n_dimensions}")
```
So this trajectory file represents a 3D box of 34 particles moving around for 40 iterations.
Now let's get the box lengths data:
```
box_lengths = traj_data.unitcell_lengths * 10
assert box_lengths.shape == (n_iterations, n_dimensions)
box_lengths[:5, :]
```
The PBCluster package requires the box size to stay the same for every iteration, so let's double-check that that's true and store just the first row in this array as the box lengths data.
```
assert np.allclose(box_lengths, box_lengths[0, 0])
box_lengths = box_lengths[0, :]
box_lengths
```
## Compute Cluster and Particle Properties
This package clusters particles based on a choice of a cutoff distance.
Any 2 particles that are within that cutoff distance of each other will be part of the same cluster.
The [radial distribution function](https://en.wikipedia.org/wiki/Radial_distribution_function) (calculated elsewhere) of this trajectory shows a first peak at $r \approx 4.4$, so I chose $r = 4.95$ to capture particle pairs in that peak.
```
cutoff_distance = 4.95
trajectory = Trajectory(particle_coords, box_lengths, cutoff_distance)
```
Let's first calculate properties associated with particle clusters. In the dataframe below, each row represents a single cluster with an ID of `cluster_id` at timestep `timestep`.
```
cluster_properties_df = trajectory.compute_cluster_properties(properties="all")
cluster_properties_df.head()
```
Now let's calculate properties associated with individual particles. In the dataframe below, each row represents a single particle with an ID of `particle_id` at timestep `timestep`.
```
particle_properties_df = trajectory.compute_particle_properties(properties="all")
particle_properties_df.head()
```
## Plot Cluster Properties
The following plot shows how the particle size distribution changes over time. Just like the rows in `cluster_properties_df`, each point represents a single cluster at a given iteration. I added some transparency to each point so overlapping points are seen as a little darker.
```
(
pn.ggplot(cluster_properties_df)
+ pn.geom_point(pn.aes(x='timestep', y='n_particles'), alpha=0.3)
)
```
## Plot Particle Properties
The following plot shows how the coordination number distribution changes over time.
Coordination number means the number of neighbors any given particle has.
Here, just like the rows of `particle_properties_df`, each point represents a single particle at a given point in time.
Since there are more total points in this chart, I added a little vertical jitter to each point to reduce the amount of overlap we see.
```
(
pn.ggplot(particle_properties_df)
+ pn.geom_jitter(pn.aes(x='timestep', y='coordination_number'), width=0, height=0.2, alpha=0.3)
+ pn.scale_y_continuous(breaks=[0, 2, 4, 6, 8])
)
```
Overall, both of these plots corroborate what can be seen in the visualization at the top: that particles aggregate over time, forming larger and larger and more densely connected clusters until all particles in the box are part of a single cluster.
| github_jupyter |
```
# Configure ipython to hide long tracebacks.
import sys
ipython = get_ipython()
def minimal_traceback(*args, **kwargs):
etype, value, tb = sys.exc_info()
value.__cause__ = None # suppress chained exceptions
stb = ipython.InteractiveTB.structured_traceback(etype, value, tb)
del stb[3:-1]
return ipython._showtraceback(etype, value, stb)
ipython.showtraceback = minimal_traceback
```
# How to Think in JAX
[](https://colab.sandbox.google.com/github/google/jax/blob/master/docs/notebooks/thinking_in_jax.ipynb)
JAX provides a simple and powerful API for writing accelerated numerical code, but working effectively in JAX sometimes requires extra consideration. This document is meant to help build a ground-up understanding of how JAX operates, so that you can use it more effectively.
## JAX vs. NumPy
**Key Concepts:**
- JAX provides a NumPy-inspired interface for convenience.
- Through duck-typing, JAX arrays can often be used as drop-in replacements of NumPy arrays.
- Unlike NumPy arrays, JAX arrays are always immutable.
NumPy provides a well-known, powerful API for working with numerical data. For convenience, JAX provides `jax.numpy` which closely mirrors the numpy API and provides easy entry into JAX. Almost anything that can be done with `numpy` can be done with `jax.numpy`:
```
import matplotlib.pyplot as plt
import numpy as np
x_np = np.linspace(0, 10, 1000)
y_np = 2 * np.sin(x_np) * np.cos(x_np)
plt.plot(x_np, y_np);
import jax.numpy as jnp
x_jnp = jnp.linspace(0, 10, 1000)
y_jnp = 2 * jnp.sin(x_jnp) * jnp.cos(x_jnp)
plt.plot(x_jnp, y_jnp);
```
The code blocks are identical aside from replacing `np` with `jnp`, and the results are the same. As we can see, JAX arrays can often be used directly in place of NumPy arrays for things like plotting.
The arrays themselves are implemented as different Python types:
```
type(x_np)
type(x_jnp)
```
Python's [duck-typing](https://en.wikipedia.org/wiki/Duck_typing) allows JAX arrays and NumPy arrays to be used interchangeably in many places.
However, there is one important difference between JAX and NumPy arrays: JAX arrays are immutable, meaning that once created their contents cannot be changed.
Here is an example of mutating an array in NumPy:
```
# NumPy: mutable arrays
x = np.arange(10)
x[0] = 10
print(x)
```
The equivalent in JAX results in an error, as JAX arrays are immutable:
```
# JAX: immutable arrays
x = jnp.arange(10)
x[0] = 10
```
For updating individual elements, JAX provides an [indexed update syntax](https://jax.readthedocs.io/en/latest/jax.ops.html#syntactic-sugar-for-indexed-update-operators) that returns an updated copy:
```
y = x.at[0].set(10)
print(x)
print(y)
```
## NumPy, lax & XLA: JAX API layering
**Key Concepts:**
- `jax.numpy` is a high-level wrapper that provides a familiar interface.
- `jax.lax` is a lower-level API that is stricter and often more powerful.
- All JAX operations are implemented in terms of operations in [XLA](https://www.tensorflow.org/xla/) – the Accelerated Linear Algebra compiler.
If you look at the source of `jax.numpy`, you'll see that all the operations are eventually expressed in terms of functions defined in `jax.lax`. You can think of `jax.lax` as a stricter, but often more powerful, API for working with multi-dimensional arrays.
For example, while `jax.numpy` will implicitly promote arguments to allow operations between mixed data types, `jax.lax` will not:
```
import jax.numpy as jnp
jnp.add(1, 1.0) # jax.numpy API implicitly promotes mixed types.
from jax import lax
lax.add(1, 1.0) # jax.lax API requires explicit type promotion.
```
If using `jax.lax` directly, you'll have to do type promotion explicitly in such cases:
```
lax.add(jnp.float32(1), 1.0)
```
Along with this strictness, `jax.lax` also provides efficient APIs for some more general operations than are supported by NumPy.
For example, consider a 1D convolution, which can be expressed in NumPy this way:
```
x = jnp.array([1, 2, 1])
y = jnp.ones(10)
jnp.convolve(x, y)
```
Under the hood, this NumPy operation is translated to a much more general convolution implemented by [`lax.conv_general_dilated`](https://jax.readthedocs.io/en/latest/_autosummary/jax.lax.conv_general_dilated.html):
```
from jax import lax
result = lax.conv_general_dilated(
x.reshape(1, 1, 3).astype(float), # note: explicit promotion
y.reshape(1, 1, 10),
window_strides=(1,),
padding=[(len(y) - 1, len(y) - 1)]) # equivalent of padding='full' in NumPy
result[0, 0]
```
This is a batched convolution operation designed to be efficient for the types of convolutions often used in deep neural nets. It requires much more boilerplate, but is far more flexible and scalable than the convolution provided by NumPy (See [JAX Sharp Bits: Convolutions](https://jax.readthedocs.io/en/latest/notebooks/Common_Gotchas_in_JAX.html#%F0%9F%94%AA-Convolutions) for more detail on JAX convolutions).
At their heart, all `jax.lax` operations are Python wrappers for operations in XLA; here, for example, the convolution implementation is provided by [XLA:ConvWithGeneralPadding](https://www.tensorflow.org/xla/operation_semantics#convwithgeneralpadding_convolution).
Every JAX operation is eventually expressed in terms of these fundamental XLA operations, which is what enables just-in-time (JIT) compilation.
## To JIT or not to JIT
**Key Concepts:**
- By default JAX executes operations one at a time, in sequence.
- Using a just-in-time (JIT) compilation decorator, sequences of operations can be optimized together and run at once.
- Not all JAX code can be JIT compiled, as it requires array shapes to be static & known at compile time.
The fact that all JAX operations are expressed in terms of XLA allows JAX to use the XLA compiler to execute blocks of code very efficiently.
For example, consider this function that normalizes the rows of a 2D matrix, expressed in terms of `jax.numpy` operations:
```
import jax.numpy as jnp
def norm(X):
X = X - X.mean(0)
return X / X.std(0)
```
A just-in-time compiled version of the function can be created using the `jax.jit` transform:
```
from jax import jit
norm_compiled = jit(norm)
```
This function returns the same results as the original, up to standard floating-point accuracy:
```
np.random.seed(1701)
X = jnp.array(np.random.rand(10000, 10))
np.allclose(norm(X), norm_compiled(X), atol=1E-6)
```
But due to the compilation (which includes fusing of operations, avoidance of allocating temporary arrays, and a host of other tricks), execution times can be orders of magnitude faster in the JIT-compiled case (note the use of `block_until_ready()` to account for JAX's [asynchronous dispatch](https://jax.readthedocs.io/en/latest/async_dispatch.html)):
```
%timeit norm(X).block_until_ready()
%timeit norm_compiled(X).block_until_ready()
```
That said, `jax.jit` does have limitations: in particular, it requires all arrays to have static shapes. That means that some JAX operations are incompatible with JIT compilation.
For example, this operation can be executed in op-by-op mode:
```
def get_negatives(x):
return x[x < 0]
x = jnp.array(np.random.randn(10))
get_negatives(x)
```
But it returns an error if you attempt to execute it in jit mode:
```
jit(get_negatives)(x)
```
This is because the function generates an array whose shape is not known at compile time: the size of the output depends on the values of the input array, and so it is not compatible with JIT.
## JIT mechanics: tracing and static variables
**Key Concepts:**
- JIT and other JAX transforms work by *tracing* a function to determine its effect on inputs of a specific shape and type.
- Variables that you don't want to be traced can be marked as *static*
To use `jax.jit` effectively, it is useful to understand how it works. Let's put a few `print()` statements within a JIT-compiled function and then call the function:
```
@jit
def f(x, y):
print("Running f():")
print(f" x = {x}")
print(f" y = {y}")
result = jnp.dot(x + 1, y + 1)
print(f" result = {result}")
return result
x = np.random.randn(3, 4)
y = np.random.randn(4)
f(x, y)
```
Notice that the print statements execute, but rather than printing the data we passed to the function, though, it prints *tracer* objects that stand-in for them.
These tracer objects are what `jax.jit` uses to extract the sequence of operations specified by the function. Basic tracers are stand-ins that encode the **shape** and **dtype** of the arrays, but are agnostic to the values. This recorded sequence of computations can then be efficiently applied within XLA to new inputs with the same shape and dtype, without having to re-execute the Python code.
When we call the compiled fuction again on matching inputs, no re-compilation is required and nothing is printed because the result is computed in compiled XLA rather than in Python:
```
x2 = np.random.randn(3, 4)
y2 = np.random.randn(4)
f(x2, y2)
```
The extracted sequence of operations is encoded in a JAX expression, or *jaxpr* for short. You can view the jaxpr using the `jax.make_jaxpr` transformation:
```
from jax import make_jaxpr
def f(x, y):
return jnp.dot(x + 1, y + 1)
make_jaxpr(f)(x, y)
```
Note one consequence of this: because JIT compilation is done *without* information on the content of the array, control flow statements in the function cannot depend on traced values. For example, this fails:
```
@jit
def f(x, neg):
return -x if neg else x
f(1, True)
```
If there are variables that you would not like to be traced, they can be marked as static for the purposes of JIT compilation:
```
from functools import partial
@partial(jit, static_argnums=(1,))
def f(x, neg):
return -x if neg else x
f(1, True)
```
Note that calling a JIT-compiled function with a different static argument results in re-compilation, so the function still works as expected:
```
f(1, False)
```
Understanding which values and operations will be static and which will be traced is a key part of using `jax.jit` effectively.
## Static vs Traced Operations
**Key Concepts:**
- Just as values can be either static or traced, operations can be static or traced.
- Static operations are evaluated at compile-time in Python; traced operations are compiled & evaluated at run-time in XLA.
- Use `numpy` for operations that you want to be static; use `jax.numpy` for operations that you want to be traced.
This distinction between static and traced values makes it important to think about how to keep a static value static. Consider this function:
```
import jax.numpy as jnp
from jax import jit
@jit
def f(x):
return x.reshape(jnp.array(x.shape).prod())
x = jnp.ones((2, 3))
f(x)
```
This fails with an error specifying that a tracer was found in `jax.numpy.reshape`. Let's add some print statements to the function to understand why this is happening:
```
@jit
def f(x):
print(f"x = {x}")
print(f"x.shape = {x.shape}")
print(f"jnp.array(x.shape).prod() = {jnp.array(x.shape).prod()}")
# comment this out to avoid the error:
# return x.reshape(jnp.array(x.shape).prot())
f(x)
```
Notice that although `x` is traced, `x.shape` is a static value. However, when we use `jnp.array` and `jnp.prod` on this static value, it becomes a traced value, at which point it cannot be used in a function like `reshape()` that requires a static input (recall: array shapes must be static).
A useful pattern is to use `numpy` for operations that should be static (i.e. done at compile-time), and use `jax.numpy` for operations that should be traced (i.e. compiled and executed at run-time). For this function, it might look like this:
```
from jax import jit
import jax.numpy as jnp
import numpy as np
@jit
def f(x):
return x.reshape((np.prod(x.shape),))
f(x)
```
For this reason, a standard convention in JAX programs is to `import numpy as np` and `import jax.numpy as jnp` so that both interfaces are available for finer control over whether operations are performed in a static matter (with `numpy`, once at compile-time) or a traced manner (with `jax.numpy`, optimized at run-time).
| github_jupyter |
# Test on Fashion Dataset
This notebook shows how to load a pretrained model and perform test on Fashion Dataset. The model is trained on 40K images. It achieves >70% mAP when test on 100 images.
```
import os
import sys
import random
import math
import re
import time
import numpy as np
import cv2
import matplotlib
import matplotlib.pyplot as plt
# Root directory of the project
ROOT_DIR = os.path.abspath("../../")
# Import Mask RCNN
sys.path.append(ROOT_DIR) # To find local version of the library
from mrcnn.config import Config
from mrcnn import utils
import mrcnn.model as modellib
from mrcnn import visualize
from mrcnn.model import log
from pycocotools.coco import COCO
%matplotlib inline
# Directory to save logs and trained model
MODEL_DIR = os.path.join(ROOT_DIR, "logs")
```
## Configurations
```
class TestConfig(Config):
"""Configuration for training on the toy shapes dataset.
Derives from the base Config class and overrides values specific
to the toy shapes dataset.
"""
# Give the configuration a recognizable name
NAME = "fashion"
# Train on 1 GPU and 8 images per GPU. We can put multiple images on each
# GPU because the images are small. Batch size is 8 (GPUs * images/GPU).
GPU_COUNT = 1
IMAGES_PER_GPU = 1
# Number of classes (including background)
NUM_CLASSES = 1 + 13 # background + 13 shapes
# Use small images for faster training. Set the limits of the small side
# the large side, and that determines the image shape.
IMAGE_MIN_DIM = 256
IMAGE_MAX_DIM = 256
# Use smaller anchors because our image and objects are small
RPN_ANCHOR_SCALES = (8, 16, 32, 64, 128) # anchor side in pixels
# Reduce training ROIs per image because the images are small and have
# few objects. Aim to allow ROI sampling to pick 33% positive ROIs.
TRAIN_ROIS_PER_IMAGE = 32
# Use a small epoch since the data is simple
STEPS_PER_EPOCH = 10
# use small validation steps since the epoch is small
VALIDATION_STEPS = 5
```
## Notebook Preferences
```
def get_ax(rows=1, cols=1, size=8):
"""Return a Matplotlib Axes array to be used in
all visualizations in the notebook. Provide a
central point to control graph sizes.
Change the default size attribute to control the size
of rendered images
"""
_, ax = plt.subplots(rows, cols, figsize=(size*cols, size*rows))
return ax
```
## Dataset
Raw dataset related.
```
import io
import lmdb
import sqlite3
import pandas as pd
import json
from PIL import Image
from IPython.display import display
class PhotoData(object):
def __init__(self, path):
self.env = lmdb.open(
path, map_size=2**36, readonly=True, lock=False
)
def __iter__(self):
with self.env.begin() as t:
with t.cursor() as c:
for key, value in c:
yield key, value
def __getitem__(self, index):
key = str(index).encode('ascii')
with self.env.begin() as t:
data = t.get(key)
if not data:
return None
with io.BytesIO(data) as f:
image = Image.open(f)
image.load()
return image
def __len__(self):
return self.env.stat()['entries']
photo_data = PhotoData(r'..'+os.path.sep+'..'+os.path.sep+'..'+os.path.sep+'photos.lmdb')
json_file = r'..' + os.path.sep + '..' + os.path.sep + '..' + os.path.sep + 'modanet2018_instances_train.json'
d = json.load(open(json_file))
coco=COCO(json_file)
```
# Fashion Dataset Class
Extend the Dataset class and add a method to load the shapes dataset, `load_shapes()`, and override the following methods:
* load_image()
* load_mask()
* image_reference()
```
from pycocotools import mask as maskUtils
class FashionDataset(utils.Dataset):
def load_fashion(self, count=5, start=0, class_ids=None):
json_file = r'..' + os.path.sep + '..' + os.path.sep + '..' + os.path.sep + 'modanet2018_instances_train.json'
d = json.load(open(json_file))
coco=COCO(json_file)
if not class_ids:
class_ids = sorted(coco.getCatIds())
if class_ids:
all_ids = []
for id in class_ids:
all_ids.extend(list(coco.getImgIds(catIds=[id])))
# Remove duplicates
all_ids = list(set(all_ids))
else:
# All images
all_ids = list(coco.imgs.keys())
random.seed(3)
random.shuffle(all_ids)
all_class_ids = sorted(coco.getCatIds())
for i in all_class_ids:
print('{}:{}'.format(i, coco.loadCats(i)[0]['name']), end='|')
self.add_class("fashion", i, coco.loadCats(i)[0]['name'])
image_ids = []
for c in range(count):
image_ids.append(all_ids[c+start])
# Add images
for i in image_ids:
self.add_image(
"fashion", image_id=i,
path=None,
width=coco.imgs[i]["width"],
height=coco.imgs[i]["height"],
annotations=coco.loadAnns(coco.getAnnIds(
imgIds=[i], catIds=class_ids, iscrowd=None)))
return image_ids
def load_image(self, image_id):
imgId = self.image_info[image_id]['id']
image = photo_data[imgId]
out = np.array(image.getdata()).astype(np.int32).reshape((image.size[1], image.size[0], 3))
return out
def image_reference(self, image_id):
"""Return the shapes data of the image."""
pass
def load_mask(self, image_id):
"""Load instance masks for the given image.
Different datasets use different ways to store masks. This
function converts the different mask format to one format
in the form of a bitmap [height, width, instances].
Returns:
masks: A bool array of shape [height, width, instance count] with
one mask per instance.
class_ids: a 1D array of class IDs of the instance masks.
"""
# If not a COCO image, delegate to parent class.
image_info = self.image_info[image_id]
instance_masks = []
class_ids = []
annotations = self.image_info[image_id]["annotations"]
# Build mask of shape [height, width, instance_count] and list
# of class IDs that correspond to each channel of the mask.
for annotation in annotations:
class_id = annotation['category_id']
if class_id:
m = self.annToMask(annotation, image_info["height"],
image_info["width"])
# Some objects are so small that they're less than 1 pixel area
# and end up rounded out. Skip those objects.
if m.max() < 1:
continue
# Is it a crowd? If so, use a negative class ID.
if annotation['iscrowd']:
# Use negative class ID for crowds
class_id *= -1
# For crowd masks, annToMask() sometimes returns a mask
# smaller than the given dimensions. If so, resize it.
if m.shape[0] != image_info["height"] or m.shape[1] != image_info["width"]:
m = np.ones([image_info["height"], image_info["width"]], dtype=bool)
instance_masks.append(m)
class_ids.append(class_id)
# Pack instance masks into an array
if class_ids:
mask = np.stack(instance_masks, axis=2).astype(np.bool)
class_ids = np.array(class_ids, dtype=np.int32)
return mask, class_ids
else:
# Call super class to return an empty mask
return super(FashionDataset, self).load_mask(image_id)
def annToRLE(self, ann, height, width):
"""
Convert annotation which can be polygons, uncompressed RLE to RLE.
:return: binary mask (numpy 2D array)
"""
segm = ann['segmentation']
if isinstance(segm, list):
# polygon -- a single object might consist of multiple parts
# we merge all parts into one mask rle code
rles = maskUtils.frPyObjects(segm, height, width)
rle = maskUtils.merge(rles)
elif isinstance(segm['counts'], list):
# uncompressed RLE
rle = maskUtils.frPyObjects(segm, height, width)
else:
# rle
rle = ann['segmentation']
return rle
def annToMask(self, ann, height, width):
"""
Convert annotation which can be polygons, uncompressed RLE, or RLE to binary mask.
:return: binary mask (numpy 2D array)
"""
rle = self.annToRLE(ann, height, width)
m = maskUtils.decode(rle)
return m
```
## Load pre-trained weights
```
subset = sorted(coco.getCatIds(catNms=['bag', 'belt', 'outer', 'dress', 'pants', 'top', 'shorts', 'skirt', 'scarf/tie']))
test_config = TestConfig()
# Recreate the model in inference mode
model = modellib.MaskRCNN(mode="inference",
config=test_config,
model_dir=MODEL_DIR)
# Get path to saved weights
# Either set a specific path or find last trained weights
# model_path = os.path.join(ROOT_DIR, ".h5 file name here")
model_path = model.find_last()
# Load trained weights
print("Loading weights from ", model_path)
model.load_weights(model_path, by_name=True)
```
## Test 100 Images
We trained our model with previous 40K, so we perform test use images after 40K.
## Test Type
We test following types:
'bag', 'belt', 'outer', 'dress', 'pants', 'top', 'shorts', 'skirt', 'scarf/tie'
```
test_count = 100
dataset_test = FashionDataset()
ids = dataset_test.load_fashion(test_count, start=40001, class_ids=subset)
dataset_test.prepare()
%%javascript
IPython.OutputArea.prototype._should_scroll = function(lines) {
return false;
}
```
## Evaluation
```
# Compute VOC-Style mAP @ IoU=0.5
# Running on 100 images. Increase for better accuracy.
image_ids = range(100)
APs = []
for image_id in image_ids:
# Load image and ground truth data
image, image_meta, gt_class_id, gt_bbox, gt_mask =\
modellib.load_image_gt(dataset_test, test_config,
image_id, use_mini_mask=False)
molded_images = np.expand_dims(modellib.mold_image(image, test_config), 0)
# Run object detection
results = model.detect([image], verbose=0)
r = results[0]
# Compute AP
AP, precisions, recalls, overlaps =\
utils.compute_ap(gt_bbox, gt_class_id, gt_mask,
r["rois"], r["class_ids"], r["scores"], r['masks'])
print('image '+str(image_id)+": id:"+str(ids[image_id])+ ', AP:' + str(AP) )
APs.append(AP)
print("mAP: ", np.mean(APs))
```
| github_jupyter |
# Romanian Transformer Cased Fine-Tuning with PyTorch
# 1. Setup
```
!rm -r sample_data
from google.colab import drive
drive.mount('/content/drive')
```
## 1.1. Using Colab GPU for Training
```
import tensorflow as tf
# Get the GPU device name.
device_name = tf.test.gpu_device_name()
# The device name should look like the following:
if device_name == '/device:GPU:0':
print('Found GPU at: {}'.format(device_name))
else:
raise SystemError('GPU device not found')
import torch
# If there's a GPU available...
if torch.cuda.is_available():
# Tell PyTorch to use the GPU.
device = torch.device("cuda")
print('There are %d GPU(s) available.' % torch.cuda.device_count())
print('We will use the GPU:', torch.cuda.get_device_name(0))
# If not...
else:
print('No GPU available, using the CPU instead.')
device = torch.device("cpu")
```
## 1.2. Installing the Hugging Face Library
```
!pip install transformers
```
The code in this notebook is actually a simplified version of the [run_glue.py](https://github.com/huggingface/transformers/blob/master/examples/run_glue.py) example script from huggingface.
`run_glue.py` is a helpful utility which allows you to pick which GLUE benchmark task you want to run on, and which pre-trained model you want to use (you can see the list of possible models [here](https://github.com/huggingface/transformers/blob/e6cff60b4cbc1158fbd6e4a1c3afda8dc224f566/examples/run_glue.py#L69)). It also supports using either the CPU, a single GPU, or multiple GPUs. It even supports using 16-bit precision if you want further speed up.
Unfortunately, all of this configurability comes at the cost of *readability*. In this Notebook, we've simplified the code greatly and added plenty of comments to make it clear what's going on.
# 2. Loading the Dataset
## 2.1. Parse
```
import numpy as np
train_file = "/content/drive/My Drive/Colab Notebooks/NLP/Projects/VarDial 2020 MOROCO/corpus/RDI-Train+Dev-VARDIAL2020/processed/train.tsv"
sentences = np.array([line.strip().split('\t')[0] for line in open(train_file)])
labels = np.array([line.strip().split('\t')[1] for line in open(train_file)])
labels = np.array(labels, dtype=int)
# Report the number of sentences.
print('Number of training sentences: {:,}\n'.format(len(sentences)))
```
# 3. Tokenization & Input Formatting
In this section, we'll transform our dataset into the format that BERT can be trained on.
## 3.1. Tokenizer
To feed our text to BERT, it must be split into tokens, and then these tokens must be mapped to their index in the tokenizer vocabulary.
The tokenization must be performed by the tokenizer included with BERT--the below cell will download this for us. We'll be using the "cased" version of Multilingual BERT here, as it is recommended over the previous "uncased" one.
```
from transformers import AutoTokenizer
# Load the Romanian Transformer tokenizer.
print('Loading Romanian Transformer tokenizer...')
tokenizer = AutoTokenizer.from_pretrained("dumitrescustefan/bert-base-romanian-cased-v1", do_lower_case=False)
```
Let's apply the tokenizer to one sentence just to see the output.
```
# Print the original sentence.
print(' Original: ', sentences[0])
# Print the sentence split into tokens.
print('Tokenized: ', tokenizer.tokenize(sentences[0]))
# Print the sentence mapped to token ids.
print('Token IDs: ', tokenizer.convert_tokens_to_ids(tokenizer.tokenize(sentences[0])))
```
When we actually convert all of our sentences, we'll use the `tokenize.encode` function to handle both steps, rather than calling `tokenize` and `convert_tokens_to_ids` separately.
Before we can do that, though, we need to talk about some of BERT's formatting requirements.
## 3.2. Required Formatting
The above code left out a few required formatting steps that we'll look at here.
*Side Note: The input format to BERT seems "over-specified" to me... We are required to give it a number of pieces of information which seem redundant, or like they could easily be inferred from the data without us explicity providing it. But it is what it is, and I suspect it will make more sense once I have a deeper understanding of the BERT internals.*
We are required to:
1. Add special tokens to the start and end of each sentence.
2. Pad & truncate all sentences to a single constant length.
3. Explicitly differentiate real tokens from padding tokens with the "attention mask".
## 3.3. Tokenize Dataset
Now we're ready to perform the real tokenization.
The `tokenizer.encode_plus` function combines multiple steps for us:
1. Split the sentence into tokens.
2. Add the special `[CLS]` and `[SEP]` tokens.
3. Map the tokens to their IDs.
4. Pad or truncate all sentences to the same length.
5. Create the attention masks which explicitly differentiate real tokens from `[PAD]` tokens.
The first four features are in `tokenizer.encode`, but I'm using `tokenizer.encode_plus` to get the fifth item (attention masks). Documentation is [here](https://huggingface.co/transformers/main_classes/tokenizer.html?highlight=encode_plus#transformers.PreTrainedTokenizer.encode_plus).
```
# Tokenize all of the sentences and map the tokens to thier word IDs.
input_ids = []
attention_masks = []
# For every sentence...
for sent in sentences:
# `encode_plus` will:
# (1) Tokenize the sentence.
# (2) Prepend the `[CLS]` token to the start.
# (3) Append the `[SEP]` token to the end.
# (4) Map tokens to their IDs.
# (5) Pad or truncate the sentence to `max_length`
# (6) Create attention masks for [PAD] tokens.
encoded_dict = tokenizer.encode_plus(
sent, # Sentence to encode.
add_special_tokens = True, # Add '[CLS]' and '[SEP]'
max_length = 128, # Pad & truncate all sentences.
pad_to_max_length = True,
return_attention_mask = True, # Construct attn. masks.
return_tensors = 'pt', # Return pytorch tensors.
)
# Add the encoded sentence to the list.
input_ids.append(encoded_dict['input_ids'])
# And its attention mask (simply differentiates padding from non-padding).
attention_masks.append(encoded_dict['attention_mask'])
# Convert the lists into tensors.
import torch
input_ids = torch.cat(input_ids, dim=0)
attention_masks = torch.cat(attention_masks, dim=0)
labels = torch.tensor(labels)
# Print sentence 0, now as a list of IDs.
print('Original: ', sentences[0])
print('Token IDs:', input_ids[0])
```
## 3.4. Training & Validation Split
Load the validation dataset.
```
import pandas as pd
# Load the dataset into a pandas dataframe.
val_df = pd.read_csv("/content/drive/My Drive/Colab Notebooks/NLP/Projects/VarDial 2020 MOROCO/corpus/RDI-Train+Dev-VARDIAL2020/processed/dev-target.tsv", delimiter='\t', header=None, names=['sentence', 'label'])
# Report the number of sentences.
print('Number of validation sentences: {:,}\n'.format(val_df.shape[0]))
# Get the lists of sentences and their labels.
val_sentences = val_df.sentence.values
val_labels = val_df.label.values.astype(int)
# Tokenize all of the sentences and map the tokens to thier word IDs.
val_input_ids = []
val_attention_masks = []
# For every sentence...
for sent in val_sentences:
# `encode_plus` will:
# (1) Tokenize the sentence.
# (2) Prepend the `[CLS]` token to the start.
# (3) Append the `[SEP]` token to the end.
# (4) Map tokens to their IDs.
# (5) Pad or truncate the sentence to `max_length`
# (6) Create attention masks for [PAD] tokens.
encoded_dict = tokenizer.encode_plus(
sent, # Sentence to encode.
add_special_tokens = True, # Add '[CLS]' and '[SEP]'
max_length = 128, # Pad & truncate all sentences.
pad_to_max_length = True,
return_attention_mask = True, # Construct attn. masks.
return_tensors = 'pt', # Return pytorch tensors.
)
# Add the encoded sentence to the list.
val_input_ids.append(encoded_dict['input_ids'])
# And its attention mask (simply differentiates padding from non-padding).
val_attention_masks.append(encoded_dict['attention_mask'])
# Convert the lists into tensors.
import torch
val_input_ids = torch.cat(val_input_ids, dim=0)
val_attention_masks = torch.cat(val_attention_masks, dim=0)
val_labels = torch.tensor(val_labels)
# Print sentence 0, now as a list of IDs.
print('Original: ', val_sentences[0])
print('Token IDs:', val_input_ids[0])
```
Divide up our training set to use 90% for training and 10% for validation.
```
from torch.utils.data import TensorDataset, random_split
# Combine the training inputs into a TensorDataset.
dataset = TensorDataset(input_ids, attention_masks, labels)
train_dataset = dataset
val_dataset = TensorDataset(val_input_ids, val_attention_masks, val_labels)
print('{:>5,} training samples'.format(len(train_dataset)))
print('{:>5,} validation samples'.format(len(val_dataset)))
```
We'll also create an iterator for our dataset using the torch DataLoader class. This helps save on memory during training because, unlike a for loop, with an iterator the entire dataset does not need to be loaded into memory.
```
from torch.utils.data import DataLoader, RandomSampler, SequentialSampler
# The DataLoader needs to know our batch size for training, so we specify it
# here. For fine-tuning BERT on a specific task, the authors recommend a batch
# size of 16 or 32.
batch_size = 32
# Create the DataLoaders for our training and validation sets.
# We'll take training samples in random order.
train_dataloader = DataLoader(
train_dataset, # The training samples.
sampler = RandomSampler(train_dataset), # Select batches randomly
batch_size = batch_size # Trains with this batch size.
)
# For validation the order doesn't matter, so we'll just read them sequentially.
validation_dataloader = DataLoader(
val_dataset, # The validation samples.
sampler = SequentialSampler(val_dataset), # Pull out batches sequentially.
batch_size = batch_size # Evaluate with this batch size.
)
```
# 4. Train Our Classification Model
Now that our input data is properly formatted, it's time to fine tune the BERT model.
## 4.1. BertForSequenceClassification
We'll be using [BertForSequenceClassification](https://huggingface.co/transformers/v2.2.0/model_doc/bert.html#bertforsequenceclassification). This is the normal BERT model with an added single linear layer on top for classification that we will use as a sentence classifier. As we feed input data, the entire pre-trained BERT model and the additional untrained classification layer is trained on our specific task.
```
# del df
del sentences
del labels
del dataset
del train_dataset
del val_dataset
del input_ids
del attention_masks
torch.cuda.empty_cache()
from transformers import BertForSequenceClassification, AdamW, BertConfig
# Load XLMRobertaForSequenceClassification model
model = BertForSequenceClassification.from_pretrained(
"dumitrescustefan/bert-base-romanian-cased-v1", # Use the recommended Multilingual BERT model, with a cased vocab
num_labels = 2, # The number of output labels -- 2 for binary classification.
# You can increase this for multi-class tasks.
output_attentions = False, # Whether the model returns attentions weights.
output_hidden_states = False, # Whether the model returns all hidden-states.
)
# Tell pytorch to run this model on the GPU.
# model.cuda()
print(model)
```
Just for curiosity's sake, we can browse all of the model's parameters by name here.
In the below cell, I've printed out the names and dimensions of the weights for:
1. The embedding layer.
2. The first of the twelve transformers.
3. The output layer.
```
# Get all of the model's parameters as a list of tuples.
params = list(model.named_parameters())
print('The Romanian Transformer model has {:} different named parameters.\n'.format(len(params)))
print('==== Embedding Layer ====\n')
for p in params[0:5]:
print("{:<55} {:>12}".format(p[0], str(tuple(p[1].size()))))
print('\n==== First Transformer ====\n')
for p in params[5:21]:
print("{:<55} {:>12}".format(p[0], str(tuple(p[1].size()))))
print('\n==== Output Layer ====\n')
for p in params[-4:]:
print("{:<55} {:>12}".format(p[0], str(tuple(p[1].size()))))
```
## 4.2. Optimizer & Learning Rate Scheduler
Now that we have our model loaded we need to grab the training hyperparameters from within the stored model.
For the purposes of fine-tuning, the authors recommend choosing from the following values (from Appendix A.3 of the [BERT paper](https://arxiv.org/pdf/1810.04805.pdf)):
>- **Batch size:** 16, 32
- **Learning rate (Adam):** 5e-5, 3e-5, 2e-5
- **Number of epochs:** 2, 3, 4
We chose:
* Batch size: 32 (set when creating our DataLoaders)
* Learning rate: 2e-5
* Epochs: 3
The epsilon parameter `eps = 1e-8` is "a very small number to prevent any division by zero in the implementation" (from [here](https://machinelearningmastery.com/adam-optimization-algorithm-for-deep-learning/)).
You can find the creation of the AdamW optimizer in `run_glue.py` [here](https://github.com/huggingface/transformers/blob/5bfcd0485ece086ebcbed2d008813037968a9e58/examples/run_glue.py#L109).
```
# Note: AdamW is a class from the huggingface library (as opposed to pytorch)
# I believe the 'W' stands for 'Weight Decay fix"
optimizer = AdamW(model.parameters(),
lr = 2e-5, # args.learning_rate - default is 5e-5, our notebook had 2e-5
eps = 1e-8 # args.adam_epsilon - default is 1e-8.
)
from transformers import get_linear_schedule_with_warmup
# Number of training epochs. The BERT authors recommend between 2 and 4.
# We chose to run for 4, but we'll see later that this may be over-fitting the
# training data.
epochs = 2
# Total number of training steps is [number of batches] x [number of epochs].
# (Note that this is not the same as the number of training samples).
total_steps = len(train_dataloader) * epochs
# Create the learning rate scheduler.
scheduler = get_linear_schedule_with_warmup(optimizer,
num_warmup_steps = int(0.06 * total_steps), # Default value in run_glue.py
num_training_steps = total_steps)
```
## 4.3. Training Loop
Below is our training loop. There's a lot going on, but fundamentally for each pass in our loop we have a training phase and a validation phase.
> *Thank you to [Stas Bekman](https://ca.linkedin.com/in/stasbekman) for contributing the insights and code for using validation loss to detect over-fitting!*
**Training:**
- Unpack our data inputs and labels
- Load data onto the GPU for acceleration
- Clear out the gradients calculated in the previous pass.
- In pytorch the gradients accumulate by default (useful for things like RNNs) unless you explicitly clear them out.
- Forward pass (feed input data through the network)
- Backward pass (backpropagation)
- Tell the network to update parameters with optimizer.step()
- Track variables for monitoring progress
**Evalution:**
- Unpack our data inputs and labels
- Load data onto the GPU for acceleration
- Forward pass (feed input data through the network)
- Compute loss on our validation data and track variables for monitoring progress
Pytorch hides all of the detailed calculations from us, but we've commented the code to point out which of the above steps are happening on each line.
> *PyTorch also has some [beginner tutorials](https://pytorch.org/tutorials/beginner/blitz/cifar10_tutorial.html#sphx-glr-beginner-blitz-cifar10-tutorial-py) which you may also find helpful.*
Define a helper function for calculating accuracy.
```
import numpy as np
# Function to calculate the accuracy of our predictions vs labels
def flat_accuracy(preds, labels):
pred_flat = np.argmax(preds, axis=1).flatten()
labels_flat = labels.flatten()
return np.sum(pred_flat == labels_flat) / len(labels_flat)
```
Helper function for formatting elapsed times as `hh:mm:ss`
```
import time
import datetime
def format_time(elapsed):
'''
Takes a time in seconds and returns a string hh:mm:ss
'''
# Round to the nearest second.
elapsed_rounded = int(round((elapsed)))
# Format as hh:mm:ss
return str(datetime.timedelta(seconds=elapsed_rounded))
```
We're ready to kick off the training!
```
import random
import numpy as np
# This training code is based on the `run_glue.py` script here:
# https://github.com/huggingface/transformers/blob/5bfcd0485ece086ebcbed2d008813037968a9e58/examples/run_glue.py#L128
# Use GPU
import torch
# If there's a GPU available...
if torch.cuda.is_available():
# Tell PyTorch to use the GPU.
device = torch.device("cuda")
print('There are %d GPU(s) available.' % torch.cuda.device_count())
print('We will use the GPU:', torch.cuda.get_device_name(0))
# If not...
else:
print('No GPU available, using the CPU instead.')
device = torch.device("cpu")
# device = torch.device("cpu")
model.cuda()
# Set the seed value all over the place to make this reproducible.
seed_val = 42
random.seed(seed_val)
np.random.seed(seed_val)
torch.manual_seed(seed_val)
# torch.cuda.manual_seed_all(seed_val)
# We'll store a number of quantities such as training and validation loss,
# validation accuracy, and timings.
training_stats = []
# Measure the total training time for the whole run.
total_t0 = time.time()
# For each epoch...
for epoch_i in range(0, epochs):
# ========================================
# Training
# ========================================
# Perform one full pass over the training set.
print("")
print('======== Epoch {:} / {:} ========'.format(epoch_i + 1, epochs))
print('Training...')
# Measure how long the training epoch takes.
t0 = time.time()
# Reset the total loss for this epoch.
total_train_loss = 0
# Put the model into training mode. Don't be mislead--the call to
# `train` just changes the *mode*, it doesn't *perform* the training.
# `dropout` and `batchnorm` layers behave differently during training
# vs. test (source: https://stackoverflow.com/questions/51433378/what-does-model-train-do-in-pytorch)
model.train()
# For each batch of training data...
for step, batch in enumerate(train_dataloader):
# Progress update every 40 batches.
if step % 10 == 0 and not step == 0:
# Calculate elapsed time in minutes.
elapsed = format_time(time.time() - t0)
# Report progress.
print(' Batch {:>5,} of {:>5,}. Elapsed: {:}.'.format(step, len(train_dataloader), elapsed))
# Unpack this training batch from our dataloader.
#
# As we unpack the batch, we'll also copy each tensor to the GPU using the
# `to` method.
#
# `batch` contains three pytorch tensors:
# [0]: input ids
# [1]: attention masks
# [2]: labels
b_input_ids = batch[0].to(device)
b_input_mask = batch[1].to(device)
b_labels = batch[2].to(device)
# Always clear any previously calculated gradients before performing a
# backward pass. PyTorch doesn't do this automatically because
# accumulating the gradients is "convenient while training RNNs".
# (source: https://stackoverflow.com/questions/48001598/why-do-we-need-to-call-zero-grad-in-pytorch)
model.zero_grad()
# Perform a forward pass (evaluate the model on this training batch).
# The documentation for this `model` function is here:
# https://huggingface.co/transformers/v2.2.0/model_doc/bert.html#transformers.BertForSequenceClassification
# It returns different numbers of parameters depending on what arguments
# arge given and what flags are set. For our useage here, it returns
# the loss (because we provided labels) and the "logits"--the model
# outputs prior to activation.
loss, logits = model(b_input_ids,
token_type_ids=None,
attention_mask=b_input_mask,
labels=b_labels)
# Accumulate the training loss over all of the batches so that we can
# calculate the average loss at the end. `loss` is a Tensor containing a
# single value; the `.item()` function just returns the Python value
# from the tensor.
total_train_loss += loss.item()
# Perform a backward pass to calculate the gradients.
loss.backward()
# Clip the norm of the gradients to 1.0.
# This is to help prevent the "exploding gradients" problem.
torch.nn.utils.clip_grad_norm_(model.parameters(), 1.0)
# Update parameters and take a step using the computed gradient.
# The optimizer dictates the "update rule"--how the parameters are
# modified based on their gradients, the learning rate, etc.
optimizer.step()
# Update the learning rate.
scheduler.step()
# Calculate the average loss over all of the batches.
avg_train_loss = total_train_loss / len(train_dataloader)
# Measure how long this epoch took.
training_time = format_time(time.time() - t0)
print("")
print(" Average training loss: {0:.2f}".format(avg_train_loss))
print(" Training epoch took: {:}".format(training_time))
# ========================================
# Validation
# ========================================
# After the completion of each training epoch, measure our performance on
# our validation set.
print("")
print("Running Validation...")
t0 = time.time()
# Put the model in evaluation mode--the dropout layers behave differently
# during evaluation.
model.eval()
# Tracking variables
total_eval_accuracy = 0
total_eval_loss = 0
nb_eval_steps = 0
# Evaluate data for one epoch
for batch in validation_dataloader:
# Unpack this training batch from our dataloader.
#
# As we unpack the batch, we'll also copy each tensor to the GPU using
# the `to` method.
#
# `batch` contains three pytorch tensors:
# [0]: input ids
# [1]: attention masks
# [2]: labels
b_input_ids = batch[0].to(device)
b_input_mask = batch[1].to(device)
b_labels = batch[2].to(device)
# Tell pytorch not to bother with constructing the compute graph during
# the forward pass, since this is only needed for backprop (training).
with torch.no_grad():
# Forward pass, calculate logit predictions.
# token_type_ids is the same as the "segment ids", which
# differentiates sentence 1 and 2 in 2-sentence tasks.
# The documentation for this `model` function is here:
# https://huggingface.co/transformers/v2.2.0/model_doc/bert.html#transformers.BertForSequenceClassification
# Get the "logits" output by the model. The "logits" are the output
# values prior to applying an activation function like the softmax.
(loss, logits) = model(b_input_ids,
token_type_ids=None,
attention_mask=b_input_mask,
labels=b_labels)
# Accumulate the validation loss.
total_eval_loss += loss.item()
# Move logits and labels to CPU
logits = logits.detach().cpu().numpy()
label_ids = b_labels.to('cpu').numpy()
# Calculate the accuracy for this batch of test sentences, and
# accumulate it over all batches.
total_eval_accuracy += flat_accuracy(logits, label_ids)
# Report the final accuracy for this validation run.
avg_val_accuracy = total_eval_accuracy / len(validation_dataloader)
print(" Accuracy: {0:.2f}".format(avg_val_accuracy))
# Calculate the average loss over all of the batches.
avg_val_loss = total_eval_loss / len(validation_dataloader)
# Measure how long the validation run took.
validation_time = format_time(time.time() - t0)
print(" Validation Loss: {0:.2f}".format(avg_val_loss))
print(" Validation took: {:}".format(validation_time))
# Record all statistics from this epoch.
training_stats.append(
{
'epoch': epoch_i + 1,
'Training Loss': avg_train_loss,
'Valid. Loss': avg_val_loss,
'Valid. Accur.': avg_val_accuracy,
'Training Time': training_time,
'Validation Time': validation_time
}
)
print("")
print("Training complete!")
print("Total training took {:} (h:mm:ss)".format(format_time(time.time()-total_t0)))
```
Save the trained model and the arguments it's been trained with.
```
import os
# Saving best-practices: if you use defaults names for the model, you can reload it using from_pretrained()
output_dir = '/content/drive/My Drive/Colab Notebooks/NLP/Projects/VarDial 2020 MOROCO/ro-trans-cased_v2/'
# Create output directory if needed
if not os.path.exists(output_dir):
os.makedirs(output_dir)
print("Saving model to %s" % output_dir)
# /content/model_save/pytorch_model.bin
# Save a trained model, configuration and tokenizer using `save_pretrained()`.
# They can then be reloaded using `from_pretrained()`
model_to_save = model.module if hasattr(model, 'module') else model # Take care of distributed/parallel training
model_to_save.save_pretrained(output_dir)
tokenizer.save_pretrained(output_dir)
# Good practice: save your training arguments together with the trained model
# torch.save(args, os.path.join(output_dir, 'training_args.bin'))
# Programatically terminate session
!kill -9 -1
```
Let's view the summary of the training process.
```
import pandas as pd
# Display floats with two decimal places.
pd.set_option('precision', 2)
# Create a DataFrame from our training statistics.
df_stats = pd.DataFrame(data=training_stats)
# Use the 'epoch' as the row index.
df_stats = df_stats.set_index('epoch')
# A hack to force the column headers to wrap.
#df = df.style.set_table_styles([dict(selector="th",props=[('max-width', '70px')])])
# Display the table.
df_stats
```
Notice that, while the the training loss is going down with each epoch, the validation loss is increasing! This suggests that we are training our model too long, and it's over-fitting on the training data.
(For reference, we are using 7,695 training samples and 856 validation samples).
Validation Loss is a more precise measure than accuracy, because with accuracy we don't care about the exact output value, but just which side of a threshold it falls on.
If we are predicting the correct answer, but with less confidence, then validation loss will catch this, while accuracy will not.
```
import matplotlib.pyplot as plt
% matplotlib inline
import seaborn as sns
# Use plot styling from seaborn.
sns.set(style='darkgrid')
# Increase the plot size and font size.
sns.set(font_scale=1.5)
plt.rcParams["figure.figsize"] = (12,6)
# Plot the learning curve.
plt.plot(df_stats['Training Loss'], 'b-o', label="Training")
plt.plot(df_stats['Valid. Loss'], 'g-o', label="Validation")
# Label the plot.
plt.title("Training & Validation Loss")
plt.xlabel("Epoch")
plt.ylabel("Loss")
plt.legend()
plt.xticks([1, 2, 3, 4])
plt.show()
```
# 5. Performance On Test Set
Now we'll load the holdout dataset and prepare inputs just as we did with the training set. Then we'll evaluate predictions using [Matthew's correlation coefficient](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.matthews_corrcoef.html) because this is the metric used by the wider NLP community to evaluate performance on CoLA. With this metric, +1 is the best score, and -1 is the worst score. This way, we can see how well we perform against the state of the art models for this specific task.
### 5.1. Data Preparation
We'll need to apply all of the same steps that we did for the training data to prepare our test data set.
```
!rm -r sample_data
from google.colab import drive
drive.mount('/content/drive')
!pip install transformers
from transformers import BertForSequenceClassification, AutoTokenizer
# Load a trained model and vocabulary that you have fine-tuned
model = BertForSequenceClassification.from_pretrained('/content/drive/My Drive/Colab Notebooks/NLP/Projects/VarDial 2020 MOROCO/ro-trans-cased_v2')
tokenizer = AutoTokenizer.from_pretrained('/content/drive/My Drive/Colab Notebooks/NLP/Projects/VarDial 2020 MOROCO/ro-trans-cased_v2', do_lower_case=False)
# Use GPU
import torch
# If there's a GPU available...
if torch.cuda.is_available():
# Tell PyTorch to use the GPU.
device = torch.device("cuda")
print('There are %d GPU(s) available.' % torch.cuda.device_count())
print('We will use the GPU:', torch.cuda.get_device_name(0))
# If not...
else:
print('No GPU available, using the CPU instead.')
device = torch.device("cpu")
# Tell pytorch to run this model on the GPU.
# model.cuda()
# Copy the model to the GPU.
model.to(device)
import pandas as pd
# Load the dataset into a pandas dataframe.
df = pd.read_csv("/content/drive/My Drive/Colab Notebooks/NLP/Projects/VarDial 2020 MOROCO/corpus/RDI-Train+Dev-VARDIAL2020/processed/dev-target.tsv", delimiter='\t', header=None, names=['sentence', 'label'])
# Report the number of sentences.
print('Number of test sentences: {:,}\n'.format(df.shape[0]))
# Create sentence and label lists
sentences = df.sentence.values
labels = df.label.values
# Tokenize all of the sentences and map the tokens to thier word IDs.
input_ids = []
attention_masks = []
# For every sentence...
for sent in sentences:
# `encode_plus` will:
# (1) Tokenize the sentence.
# (2) Prepend the `[CLS]` token to the start.
# (3) Append the `[SEP]` token to the end.
# (4) Map tokens to their IDs.
# (5) Pad or truncate the sentence to `max_length`
# (6) Create attention masks for [PAD] tokens.
encoded_dict = tokenizer.encode_plus(
sent, # Sentence to encode.
add_special_tokens = True, # Add '[CLS]' and '[SEP]'
max_length = 512, # Pad & truncate all sentences.
pad_to_max_length = True,
return_attention_mask = True, # Construct attn. masks.
return_tensors = 'pt', # Return pytorch tensors.
)
# Add the encoded sentence to the list.
input_ids.append(encoded_dict['input_ids'])
# And its attention mask (simply differentiates padding from non-padding).
attention_masks.append(encoded_dict['attention_mask'])
# Convert the lists into tensors.
import torch
from torch.utils.data import TensorDataset, DataLoader, RandomSampler, SequentialSampler
input_ids = torch.cat(input_ids, dim=0)
attention_masks = torch.cat(attention_masks, dim=0)
labels = torch.tensor(labels)
# Set the batch size.
batch_size = 8
# Create the DataLoader.
prediction_data = TensorDataset(input_ids, attention_masks, labels)
prediction_sampler = SequentialSampler(prediction_data)
prediction_dataloader = DataLoader(prediction_data, sampler=prediction_sampler, batch_size=batch_size)
```
## 5.2. Evaluate on Test Set
With the test set prepared, we can apply our fine-tuned model to generate predictions on the test set.
```
import numpy as np
# Prediction on test set
print('Predicting labels for {:,} test sentences...'.format(len(input_ids)))
# Put model in evaluation mode
model.eval()
# Tracking variables
predictions , true_labels = [], []
# Predict
for batch in prediction_dataloader:
# Add batch to GPU
batch = tuple(t.to(device) for t in batch)
# Unpack the inputs from our dataloader
b_input_ids, b_input_mask, b_labels = batch
# Telling the model not to compute or store gradients, saving memory and
# speeding up prediction
with torch.no_grad():
# Forward pass, calculate logit predictions
outputs = model(b_input_ids, token_type_ids=None,
attention_mask=b_input_mask)
logits = outputs[0]
# Move logits and labels to CPU
logits = logits.detach().cpu().numpy()
label_ids = b_labels.to('cpu').numpy()
# Store predictions and true labels
predictions.extend(logits)
true_labels.extend(label_ids)
# predictions = np.argmax(predictions, axis=1).flatten()
from scipy.special import expit
predictions = expit(np.array(predictions)[:, 1])
print(' DONE.')
```
Compute the AUC and plot the ROC.
```
from sklearn.metrics import roc_curve, auc
fpr, tpr, threshold = roc_curve(true_labels, predictions)
roc_auc = auc(fpr, tpr)
print(roc_auc)
import matplotlib.pyplot as plt
plt.title('Receiver Operating Characteristic')
plt.plot(fpr, tpr, 'b', label = 'AUC = %0.6f' % roc_auc)
plt.legend(loc = 'lower right')
plt.plot([0, 1], [0, 1],'r--')
plt.xlim([0, 1])
plt.ylim([0, 1])
plt.ylabel('True Positive Rate')
plt.xlabel('False Positive Rate')
plt.show()
```
Compute the F1 score.
```
from sklearn.metrics import f1_score
print(f1_score(true_labels, [1 if prediction > 0.5 else 0 for prediction in predictions]))
print(predictions)
```
# Appendix
## A1. Saving & Loading Fine-Tuned Model
This first cell (taken from `run_glue.py` [here](https://github.com/huggingface/transformers/blob/35ff345fc9df9e777b27903f11fa213e4052595b/examples/run_glue.py#L495)) writes the model and tokenizer out to disk.
```
import os
# Saving best-practices: if you use defaults names for the model, you can reload it using from_pretrained()
output_dir = './model_save/'
# Create output directory if needed
if not os.path.exists(output_dir):
os.makedirs(output_dir)
print("Saving model to %s" % output_dir)
# Save a trained model, configuration and tokenizer using `save_pretrained()`.
# They can then be reloaded using `from_pretrained()`
model_to_save = model.module if hasattr(model, 'module') else model # Take care of distributed/parallel training
model_to_save.save_pretrained(output_dir)
tokenizer.save_pretrained(output_dir)
# Good practice: save your training arguments together with the trained model
# torch.save(args, os.path.join(output_dir, 'training_args.bin'))
```
Let's check out the file sizes, out of curiosity.
```
!ls -l --block-size=K ./model_save/
```
The largest file is the model weights, at around 418 megabytes.
```
!ls -l --block-size=M ./model_save/pytorch_model.bin
```
To save your model across Colab Notebook sessions, download it to your local machine, or ideally copy it to your Google Drive.
```
# Mount Google Drive to this Notebook instance.
from google.colab import drive
drive.mount('/content/drive')
# Copy the model files to a directory in your Google Drive.
!cp -r ./model_save/ "./drive/Shared drives/ChrisMcCormick.AI/Blog Posts/BERT Fine-Tuning/"
```
The following functions will load the model back from disk.
```
# Load a trained model and vocabulary that you have fine-tuned
model = model_class.from_pretrained(output_dir)
tokenizer = tokenizer_class.from_pretrained(output_dir)
# Copy the model to the GPU.
model.to(device)
```
## A.2. Weight Decay
The huggingface example includes the following code block for enabling weight decay, but the default decay rate is "0.0", so I moved this to the appendix.
This block essentially tells the optimizer to not apply weight decay to the bias terms (e.g., $ b $ in the equation $ y = Wx + b $ ). Weight decay is a form of regularization--after calculating the gradients, we multiply them by, e.g., 0.99.
```
# This code is taken from:
# https://github.com/huggingface/transformers/blob/5bfcd0485ece086ebcbed2d008813037968a9e58/examples/run_glue.py#L102
# Don't apply weight decay to any parameters whose names include these tokens.
# (Here, the BERT doesn't have `gamma` or `beta` parameters, only `bias` terms)
no_decay = ['bias', 'LayerNorm.weight']
# Separate the `weight` parameters from the `bias` parameters.
# - For the `weight` parameters, this specifies a 'weight_decay_rate' of 0.01.
# - For the `bias` parameters, the 'weight_decay_rate' is 0.0.
optimizer_grouped_parameters = [
# Filter for all parameters which *don't* include 'bias', 'gamma', 'beta'.
{'params': [p for n, p in param_optimizer if not any(nd in n for nd in no_decay)],
'weight_decay_rate': 0.1},
# Filter for parameters which *do* include those.
{'params': [p for n, p in param_optimizer if any(nd in n for nd in no_decay)],
'weight_decay_rate': 0.0}
]
# Note - `optimizer_grouped_parameters` only includes the parameter values, not
# the names.
```
# Revision History
**Version 3** - *Mar 18th, 2020* - (current)
* Simplified the tokenization and input formatting (for both training and test) by leveraging the `tokenizer.encode_plus` function.
`encode_plus` handles padding *and* creates the attention masks for us.
* Improved explanation of attention masks.
* Switched to using `torch.utils.data.random_split` for creating the training-validation split.
* Added a summary table of the training statistics (validation loss, time per epoch, etc.).
* Added validation loss to the learning curve plot, so we can see if we're overfitting.
* Thank you to [Stas Bekman](https://ca.linkedin.com/in/stasbekman) for contributing this!
* Displayed the per-batch MCC as a bar plot.
**Version 2** - *Dec 20th, 2019* - [link](https://colab.research.google.com/drive/1Y4o3jh3ZH70tl6mCd76vz_IxX23biCPP)
* huggingface renamed their library to `transformers`.
* Updated the notebook to use the `transformers` library.
**Version 1** - *July 22nd, 2019*
* Initial version.
## Further Work
* It might make more sense to use the MCC score for “validation accuracy”, but I’ve left it out so as not to have to explain it earlier in the Notebook.
* Seeding -- I’m not convinced that setting the seed values at the beginning of the training loop is actually creating reproducible results…
* The MCC score seems to vary substantially across different runs. It would be interesting to run this example a number of times and show the variance.
| github_jupyter |
# CMPS 392 - Machine Learning
## *Assignment 3*
##### **Mohamed Nassar, Spring 2020**
---
## Ex 1
```
# Explain
import numpy as np
a = np.array([0., np.finfo(np.float32).eps/2 ]).astype('float32')
print (a.argmax())
print ( (a+1).argmax() )
```
## Ex 2
```
# Explain and propose a better solution to compute the variance of the numpy array x
import numpy.random as rand
x = np.array([10000 + rand.random() for i in range(10)]).astype('float32')
variance = np.mean(np.power(x,2,dtype='float32'),dtype='float32') - np.power(np.mean(x, dtype='float32'),2,dtype='float32')
print (variance)
stddev = np.sqrt(variance)
np.std(x)
```
## Ex 3
```
# Take learning rate = 0.18, 0.01, 0.1, 0.2 and explain what's happening when we perform gradient descent
# Why learning rate = 0.1 performs so nicely at the begining of the descent. Justify.
import matplotlib.pyplot as plt
A = np.array ([[0.1,0],[0,10]])
theta = np.pi/3
R = np.array ( [[ np.cos(theta), np.sin(theta)] , [-np.sin(theta), np.cos(theta)]] )
H = np.matmul ( np.matmul (np.transpose(R), A ) , R )
print (H)
x1_vals = np.arange(-200, 200, 1)
x2_vals = np.arange(-200, 200 , 1)
x1, x2 = np.meshgrid(x1_vals , x2_vals)
z = 7.525/2 * x1**2 + 2.575/2 * x2**2 + -4.32 * x1 * x2 + -9 * x2 + 15
fig = plt.figure(figsize=(10,10))
ax = plt.axes()
cp = ax.contour(x1, x2, z, [0, 1000, 10000, 100000])
ax.clabel(cp, inline=True, fontsize=10)
ax.set_title('Contour Plot')
ax.set_xlabel('x1 ')
ax.set_ylabel('x2 ')
# ax.set_xlim([-100,-70])
# ax.set_ylim([-200,-150])
# plt.show()
# gradient descent
x1, x2 = -190, -150
eps = 0.18
pts_x1 = [x1]
pts_x2 = [x2]
for i in range (100 ):
g= np.array ( [(7.525 * x1 -4.32 * x2 ) , (2.575*x2 -4.32 * x1 -9) ])
gt_h_g = np.dot ( np.dot ( g , H ) , g)
gt_g = np.dot ( g , g )
# print (gt_g/gt_h_g)
(x1, x2) = (x1 - eps * g[0] , x2 - eps * g[1] )
pts_x1.append(x1)
pts_x2.append(x2)
plt.plot(pts_x1, pts_x2, 'r-x')
plt.show()
```
## Ex 4
```
# explain what is going wrong, propose a fix
# n.b. you cannot change the hard coded numbers
def softmax (x):
return np.exp(x)/np.sum(np.exp(x))
def logloss ( probs, y ):
return -np.log (np.sum( probs * y))
logits = np.array([89, 50, 60]).astype('float32')
probs = softmax(logits)
y = np.array([1, 0, 0])
loss = logloss ( probs, y )
print (loss)
```
## Ex 5
```
# explain what is going wrong, propose a fix
def sigmoid(x):
return (1/(1+ np.exp(-x)))
def logloss ( prob, y ):
return -np.log (prob * y)
logit = np.float32(-89)
prob = sigmoid(logit)
y = 1
loss = logloss ( prob, y )
print (loss)
```
## Ex 6
Propose an example of your choice to show why it is worth keeping an eye on numerical computations issues when implementing machine learning algorithms
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.