text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
# Batch Normalization
One way to make deep networks easier to train is to use more sophisticated optimization procedures such as SGD+momentum, RMSProp, or Adam. Another strategy is to change the architecture of the network to make it easier to train.
One idea along these lines is batch normalization which was proposed by [1] in 2015.
The idea is relatively straightforward. Machine learning methods tend to work better when their input data consists of uncorrelated features with zero mean and unit variance. When training a neural network, we can preprocess the data before feeding it to the network to explicitly decorrelate its features; this will ensure that the first layer of the network sees data that follows a nice distribution. However, even if we preprocess the input data, the activations at deeper layers of the network will likely no longer be decorrelated and will no longer have zero mean or unit variance since they are output from earlier layers in the network. Even worse, during the training process the distribution of features at each layer of the network will shift as the weights of each layer are updated.
The authors of [1] hypothesize that the shifting distribution of features inside deep neural networks may make training deep networks more difficult. To overcome this problem, [1] proposes to insert batch normalization layers into the network. At training time, a batch normalization layer uses a minibatch of data to estimate the mean and standard deviation of each feature. These estimated means and standard deviations are then used to center and normalize the features of the minibatch. A running average of these means and standard deviations is kept during training, and at test time these running averages are used to center and normalize features.
It is possible that this normalization strategy could reduce the representational power of the network, since it may sometimes be optimal for certain layers to have features that are not zero-mean or unit variance. To this end, the batch normalization layer includes learnable shift and scale parameters for each feature dimension.
[1] [Sergey Ioffe and Christian Szegedy, "Batch Normalization: Accelerating Deep Network Training by Reducing
Internal Covariate Shift", ICML 2015.](https://arxiv.org/abs/1502.03167)
```
# As usual, a bit of setup
import time
import numpy as np
import matplotlib.pyplot as plt
from cs231n.classifiers.fc_net import *
from cs231n.data_utils import get_CIFAR10_data
from cs231n.gradient_check import eval_numerical_gradient, eval_numerical_gradient_array
from cs231n.solver import Solver
%matplotlib inline
plt.rcParams['figure.figsize'] = (10.0, 8.0) # set default size of plots
plt.rcParams['image.interpolation'] = 'nearest'
plt.rcParams['image.cmap'] = 'gray'
# for auto-reloading external modules
# see http://stackoverflow.com/questions/1907993/autoreload-of-modules-in-ipython
%load_ext autoreload
%autoreload 2
def rel_error(x, y):
""" returns relative error """
return np.max(np.abs(x - y) / (np.maximum(1e-8, np.abs(x) + np.abs(y))))
def print_mean_std(x,axis=0):
print(' means: ', x.mean(axis=axis))
print(' stds: ', x.std(axis=axis))
print()
# Load the (preprocessed) CIFAR10 data.
data = get_CIFAR10_data()
for k, v in data.items():
print('%s: ' % k, v.shape)
```
## Batch normalization: forward
In the file `cs231n/layers.py`, implement the batch normalization forward pass in the function `batchnorm_forward`. Once you have done so, run the following to test your implementation.
Referencing the paper linked to above in [1] may be helpful!
```
# Check the training-time forward pass by checking means and variances
# of features both before and after batch normalization
# Simulate the forward pass for a two-layer network
np.random.seed(231)
N, D1, D2, D3 = 200, 50, 60, 3
X = np.random.randn(N, D1)
W1 = np.random.randn(D1, D2)
W2 = np.random.randn(D2, D3)
a = np.maximum(0, X.dot(W1)).dot(W2)
print('Before batch normalization:')
print_mean_std(a,axis=0)
gamma = np.ones((D3,))
beta = np.zeros((D3,))
# Means should be close to zero and stds close to one
print('After batch normalization (gamma=1, beta=0)')
a_norm, _ = batchnorm_forward(a, gamma, beta, {'mode': 'train'})
print_mean_std(a_norm,axis=0)
gamma = np.asarray([1.0, 2.0, 3.0])
beta = np.asarray([11.0, 12.0, 13.0])
# Now means should be close to beta and stds close to gamma
print('After batch normalization (gamma=', gamma, ', beta=', beta, ')')
a_norm, _ = batchnorm_forward(a, gamma, beta, {'mode': 'train'})
print_mean_std(a_norm,axis=0)
# Check the test-time forward pass by running the training-time
# forward pass many times to warm up the running averages, and then
# checking the means and variances of activations after a test-time
# forward pass.
np.random.seed(231)
N, D1, D2, D3 = 200, 50, 60, 3
W1 = np.random.randn(D1, D2)
W2 = np.random.randn(D2, D3)
bn_param = {'mode': 'train'}
gamma = np.ones(D3)
beta = np.zeros(D3)
for t in range(50):
X = np.random.randn(N, D1)
a = np.maximum(0, X.dot(W1)).dot(W2)
batchnorm_forward(a, gamma, beta, bn_param)
bn_param['mode'] = 'test'
X = np.random.randn(N, D1)
a = np.maximum(0, X.dot(W1)).dot(W2)
a_norm, _ = batchnorm_forward(a, gamma, beta, bn_param)
# Means should be close to zero and stds close to one, but will be
# noisier than training-time forward passes.
print('After batch normalization (test-time):')
print_mean_std(a_norm,axis=0)
```
## Batch normalization: backward
Now implement the backward pass for batch normalization in the function `batchnorm_backward`.
To derive the backward pass you should write out the computation graph for batch normalization and backprop through each of the intermediate nodes. Some intermediates may have multiple outgoing branches; make sure to sum gradients across these branches in the backward pass.
Once you have finished, run the following to numerically check your backward pass.
```
# Gradient check batchnorm backward pass
np.random.seed(231)
N, D = 4, 5
x = 5 * np.random.randn(N, D) + 12
gamma = np.random.randn(D)
beta = np.random.randn(D)
dout = np.random.randn(N, D)
bn_param = {'mode': 'train'}
fx = lambda x: batchnorm_forward(x, gamma, beta, bn_param)[0]
fg = lambda a: batchnorm_forward(x, a, beta, bn_param)[0]
fb = lambda b: batchnorm_forward(x, gamma, b, bn_param)[0]
dx_num = eval_numerical_gradient_array(fx, x, dout)
da_num = eval_numerical_gradient_array(fg, gamma.copy(), dout)
db_num = eval_numerical_gradient_array(fb, beta.copy(), dout)
_, cache = batchnorm_forward(x, gamma, beta, bn_param)
dx, dgamma, dbeta = batchnorm_backward(dout, cache)
#You should expect to see relative errors between 1e-13 and 1e-8
print('dx error: ', rel_error(dx_num, dx))
print('dgamma error: ', rel_error(da_num, dgamma))
print('dbeta error: ', rel_error(db_num, dbeta))
```
## Batch normalization: alternative backward
In class we talked about two different implementations for the sigmoid backward pass. One strategy is to write out a computation graph composed of simple operations and backprop through all intermediate values. Another strategy is to work out the derivatives on paper. For example, you can derive a very simple formula for the sigmoid function's backward pass by simplifying gradients on paper.
Surprisingly, it turns out that you can do a similar simplification for the batch normalization backward pass too!
In the forward pass, given a set of inputs $X=\begin{bmatrix}x_1\\x_2\\...\\x_N\end{bmatrix}$,
we first calculate the mean $\mu$ and variance $v$.
With $\mu$ and $v$ calculated, we can calculate the standard deviation $\sigma$ and normalized data $Y$.
The equations and graph illustration below describe the computation ($y_i$ is the i-th element of the vector $Y$).
\begin{align}
& \mu=\frac{1}{N}\sum_{k=1}^N x_k & v=\frac{1}{N}\sum_{k=1}^N (x_k-\mu)^2 \\
& \sigma=\sqrt{v+\epsilon} & y_i=\frac{x_i-\mu}{\sigma}
\end{align}
<img src="notebook_images/batchnorm_graph.png" width=691 height=202>
The meat of our problem during backpropagation is to compute $\frac{\partial L}{\partial X}$, given the upstream gradient we receive, $\frac{\partial L}{\partial Y}.$ To do this, recall the chain rule in calculus gives us $\frac{\partial L}{\partial X} = \frac{\partial L}{\partial Y} \cdot \frac{\partial Y}{\partial X}$.
The unknown/hart part is $\frac{\partial Y}{\partial X}$. We can find this by first deriving step-by-step our local gradients at
$\frac{\partial v}{\partial X}$, $\frac{\partial \mu}{\partial X}$,
$\frac{\partial \sigma}{\partial v}$,
$\frac{\partial Y}{\partial \sigma}$, and $\frac{\partial Y}{\partial \mu}$,
and then use the chain rule to compose these gradients (which appear in the form of vectors!) appropriately to compute $\frac{\partial Y}{\partial X}$.
If it's challenging to directly reason about the gradients over $X$ and $Y$ which require matrix multiplication, try reasoning about the gradients in terms of individual elements $x_i$ and $y_i$ first: in that case, you will need to come up with the derivations for $\frac{\partial L}{\partial x_i}$, by relying on the Chain Rule to first calculate the intermediate $\frac{\partial \mu}{\partial x_i}, \frac{\partial v}{\partial x_i}, \frac{\partial \sigma}{\partial x_i},$ then assemble these pieces to calculate $\frac{\partial y_i}{\partial x_i}$.
You should make sure each of the intermediary gradient derivations are all as simplified as possible, for ease of implementation.
After doing so, implement the simplified batch normalization backward pass in the function `batchnorm_backward_alt` and compare the two implementations by running the following. Your two implementations should compute nearly identical results, but the alternative implementation should be a bit faster.
```
np.random.seed(231)
N, D = 100, 500
x = 5 * np.random.randn(N, D) + 12
gamma = np.random.randn(D)
beta = np.random.randn(D)
dout = np.random.randn(N, D)
bn_param = {'mode': 'train'}
out, cache = batchnorm_forward(x, gamma, beta, bn_param)
t1 = time.time()
dx1, dgamma1, dbeta1 = batchnorm_backward(dout, cache)
t2 = time.time()
dx2, dgamma2, dbeta2 = batchnorm_backward_alt(dout, cache)
t3 = time.time()
print('dx difference: ', rel_error(dx1, dx2))
print('dgamma difference: ', rel_error(dgamma1, dgamma2))
print('dbeta difference: ', rel_error(dbeta1, dbeta2))
print('speedup: %.2fx' % ((t2 - t1) / (t3 - t2)))
```
## Fully Connected Nets with Batch Normalization
Now that you have a working implementation for batch normalization, go back to your `FullyConnectedNet` in the file `cs231n/classifiers/fc_net.py`. Modify your implementation to add batch normalization.
Concretely, when the `normalization` flag is set to `"batchnorm"` in the constructor, you should insert a batch normalization layer before each ReLU nonlinearity. The outputs from the last layer of the network should not be normalized. Once you are done, run the following to gradient-check your implementation.
HINT: You might find it useful to define an additional helper layer similar to those in the file `cs231n/layer_utils.py`. If you decide to do so, do it in the file `cs231n/classifiers/fc_net.py`.
```
np.random.seed(231)
N, D, H1, H2, C = 2, 15, 20, 30, 10
X = np.random.randn(N, D)
y = np.random.randint(C, size=(N,))
# You should expect losses between 1e-4~1e-10 for W,
# losses between 1e-08~1e-10 for b,
# and losses between 1e-08~1e-09 for beta and gammas.
for reg in [0, 3.14]:
print('Running check with reg = ', reg)
model = FullyConnectedNet([H1, H2], input_dim=D, num_classes=C,
reg=reg, weight_scale=5e-2, dtype=np.float64,
normalization='batchnorm')
loss, grads = model.loss(X, y)
print('Initial loss: ', loss)
for name in sorted(grads):
f = lambda _: model.loss(X, y)[0]
grad_num = eval_numerical_gradient(f, model.params[name], verbose=False, h=1e-5)
print('%s relative error: %.2e' % (name, rel_error(grad_num, grads[name])))
if reg == 0: print()
```
# Batchnorm for deep networks
Run the following to train a six-layer network on a subset of 1000 training examples both with and without batch normalization.
```
np.random.seed(231)
# Try training a very deep net with batchnorm
hidden_dims = [100, 100, 100, 100, 100]
num_train = 1000
small_data = {
'X_train': data['X_train'][:num_train],
'y_train': data['y_train'][:num_train],
'X_val': data['X_val'],
'y_val': data['y_val'],
}
weight_scale = 2e-2
bn_model = FullyConnectedNet(hidden_dims, weight_scale=weight_scale, normalization='batchnorm')
model = FullyConnectedNet(hidden_dims, weight_scale=weight_scale, normalization=None)
print('Solver with batch norm:')
bn_solver = Solver(bn_model, small_data,
num_epochs=10, batch_size=50,
update_rule='adam',
optim_config={
'learning_rate': 1e-3,
},
verbose=True,print_every=20)
bn_solver.train()
print('\nSolver without batch norm:')
solver = Solver(model, small_data,
num_epochs=10, batch_size=50,
update_rule='adam',
optim_config={
'learning_rate': 1e-3,
},
verbose=True, print_every=20)
solver.train()
```
Run the following to visualize the results from two networks trained above. You should find that using batch normalization helps the network to converge much faster.
```
def plot_training_history(title, label, baseline, bn_solvers, plot_fn, bl_marker='.', bn_marker='.', labels=None):
"""utility function for plotting training history"""
plt.title(title)
plt.xlabel(label)
bn_plots = [plot_fn(bn_solver) for bn_solver in bn_solvers]
bl_plot = plot_fn(baseline)
num_bn = len(bn_plots)
for i in range(num_bn):
label='with_norm'
if labels is not None:
label += str(labels[i])
plt.plot(bn_plots[i], bn_marker, label=label)
label='baseline'
if labels is not None:
label += str(labels[0])
plt.plot(bl_plot, bl_marker, label=label)
plt.legend(loc='lower center', ncol=num_bn+1)
plt.subplot(3, 1, 1)
plot_training_history('Training loss','Iteration', solver, [bn_solver], \
lambda x: x.loss_history, bl_marker='o', bn_marker='o')
plt.subplot(3, 1, 2)
plot_training_history('Training accuracy','Epoch', solver, [bn_solver], \
lambda x: x.train_acc_history, bl_marker='-o', bn_marker='-o')
plt.subplot(3, 1, 3)
plot_training_history('Validation accuracy','Epoch', solver, [bn_solver], \
lambda x: x.val_acc_history, bl_marker='-o', bn_marker='-o')
plt.gcf().set_size_inches(15, 15)
plt.show()
```
# Batch normalization and initialization
We will now run a small experiment to study the interaction of batch normalization and weight initialization.
The first cell will train 8-layer networks both with and without batch normalization using different scales for weight initialization. The second layer will plot training accuracy, validation set accuracy, and training loss as a function of the weight initialization scale.
```
np.random.seed(231)
# Try training a very deep net with batchnorm
hidden_dims = [50, 50, 50, 50, 50, 50, 50]
num_train = 1000
small_data = {
'X_train': data['X_train'][:num_train],
'y_train': data['y_train'][:num_train],
'X_val': data['X_val'],
'y_val': data['y_val'],
}
bn_solvers_ws = {}
solvers_ws = {}
weight_scales = np.logspace(-4, 0, num=20)
for i, weight_scale in enumerate(weight_scales):
print('Running weight scale %d / %d' % (i + 1, len(weight_scales)))
bn_model = FullyConnectedNet(hidden_dims, weight_scale=weight_scale, normalization='batchnorm')
model = FullyConnectedNet(hidden_dims, weight_scale=weight_scale, normalization=None)
bn_solver = Solver(bn_model, small_data,
num_epochs=10, batch_size=50,
update_rule='adam',
optim_config={
'learning_rate': 1e-3,
},
verbose=False, print_every=200)
bn_solver.train()
bn_solvers_ws[weight_scale] = bn_solver
solver = Solver(model, small_data,
num_epochs=10, batch_size=50,
update_rule='adam',
optim_config={
'learning_rate': 1e-3,
},
verbose=False, print_every=200)
solver.train()
solvers_ws[weight_scale] = solver
# Plot results of weight scale experiment
best_train_accs, bn_best_train_accs = [], []
best_val_accs, bn_best_val_accs = [], []
final_train_loss, bn_final_train_loss = [], []
for ws in weight_scales:
best_train_accs.append(max(solvers_ws[ws].train_acc_history))
bn_best_train_accs.append(max(bn_solvers_ws[ws].train_acc_history))
best_val_accs.append(max(solvers_ws[ws].val_acc_history))
bn_best_val_accs.append(max(bn_solvers_ws[ws].val_acc_history))
final_train_loss.append(np.mean(solvers_ws[ws].loss_history[-100:]))
bn_final_train_loss.append(np.mean(bn_solvers_ws[ws].loss_history[-100:]))
plt.subplot(3, 1, 1)
plt.title('Best val accuracy vs weight initialization scale')
plt.xlabel('Weight initialization scale')
plt.ylabel('Best val accuracy')
plt.semilogx(weight_scales, best_val_accs, '-o', label='baseline')
plt.semilogx(weight_scales, bn_best_val_accs, '-o', label='batchnorm')
plt.legend(ncol=2, loc='lower right')
plt.subplot(3, 1, 2)
plt.title('Best train accuracy vs weight initialization scale')
plt.xlabel('Weight initialization scale')
plt.ylabel('Best training accuracy')
plt.semilogx(weight_scales, best_train_accs, '-o', label='baseline')
plt.semilogx(weight_scales, bn_best_train_accs, '-o', label='batchnorm')
plt.legend()
plt.subplot(3, 1, 3)
plt.title('Final training loss vs weight initialization scale')
plt.xlabel('Weight initialization scale')
plt.ylabel('Final training loss')
plt.semilogx(weight_scales, final_train_loss, '-o', label='baseline')
plt.semilogx(weight_scales, bn_final_train_loss, '-o', label='batchnorm')
plt.legend()
plt.gca().set_ylim(1.0, 3.5)
plt.gcf().set_size_inches(15, 15)
plt.show()
```
## Inline Question 1:
Describe the results of this experiment. How does the scale of weight initialization affect models with/without batch normalization differently, and why?
## Answer:
[A network without batch normalization is highly sensitive to the weight initialization scale to avoid for example vanishing gradients. With batch normalization the network can perform well with a greater range weight initialization scales which is seen by the final training loss as well as the accuracy on the training and validation set. ]
# Batch normalization and batch size
We will now run a small experiment to study the interaction of batch normalization and batch size.
The first cell will train 6-layer networks both with and without batch normalization using different batch sizes. The second layer will plot training accuracy and validation set accuracy over time.
```
def run_batchsize_experiments(normalization_mode):
np.random.seed(231)
# Try training a very deep net with batchnorm
hidden_dims = [100, 100, 100, 100, 100]
num_train = 1000
small_data = {
'X_train': data['X_train'][:num_train],
'y_train': data['y_train'][:num_train],
'X_val': data['X_val'],
'y_val': data['y_val'],
}
n_epochs=10
weight_scale = 2e-2
batch_sizes = [5,10,50]
lr = 10**(-3.5)
solver_bsize = batch_sizes[0]
print('No normalization: batch size = ',solver_bsize)
model = FullyConnectedNet(hidden_dims, weight_scale=weight_scale, normalization=None)
solver = Solver(model, small_data,
num_epochs=n_epochs, batch_size=solver_bsize,
update_rule='adam',
optim_config={
'learning_rate': lr,
},
verbose=False)
solver.train()
bn_solvers = []
for i in range(len(batch_sizes)):
b_size=batch_sizes[i]
print('Normalization: batch size = ',b_size)
bn_model = FullyConnectedNet(hidden_dims, weight_scale=weight_scale, normalization=normalization_mode)
bn_solver = Solver(bn_model, small_data,
num_epochs=n_epochs, batch_size=b_size,
update_rule='adam',
optim_config={
'learning_rate': lr,
},
verbose=False)
bn_solver.train()
bn_solvers.append(bn_solver)
return bn_solvers, solver, batch_sizes
batch_sizes = [5,10,50]
bn_solvers_bsize, solver_bsize, batch_sizes = run_batchsize_experiments('batchnorm')
plt.subplot(2, 1, 1)
plot_training_history('Training accuracy (Batch Normalization)','Epoch', solver_bsize, bn_solvers_bsize, \
lambda x: x.train_acc_history, bl_marker='-^', bn_marker='-o', labels=batch_sizes)
plt.subplot(2, 1, 2)
plot_training_history('Validation accuracy (Batch Normalization)','Epoch', solver_bsize, bn_solvers_bsize, \
lambda x: x.val_acc_history, bl_marker='-^', bn_marker='-o', labels=batch_sizes)
plt.gcf().set_size_inches(15, 10)
plt.show()
```
## Inline Question 2:
Describe the results of this experiment. What does this imply about the relationship between batch normalization and batch size? Why is this relationship observed?
## Answer:
[FILL THIS IN]
# Layer Normalization
Batch normalization has proved to be effective in making networks easier to train, but the dependency on batch size makes it less useful in complex networks which have a cap on the input batch size due to hardware limitations.
Several alternatives to batch normalization have been proposed to mitigate this problem; one such technique is Layer Normalization [2]. Instead of normalizing over the batch, we normalize over the features. In other words, when using Layer Normalization, each feature vector corresponding to a single datapoint is normalized based on the sum of all terms within that feature vector.
[2] [Ba, Jimmy Lei, Jamie Ryan Kiros, and Geoffrey E. Hinton. "Layer Normalization." stat 1050 (2016): 21.](https://arxiv.org/pdf/1607.06450.pdf)
## Inline Question 3:
Which of these data preprocessing steps is analogous to batch normalization, and which is analogous to layer normalization?
1. Scaling each image in the dataset, so that the RGB channels for each row of pixels within an image sums up to 1.
2. Scaling each image in the dataset, so that the RGB channels for all pixels within an image sums up to 1.
3. Subtracting the mean image of the dataset from each image in the dataset.
4. Setting all RGB values to either 0 or 1 depending on a given threshold.
## Answer:
[FILL THIS IN]
# Layer Normalization: Implementation
Now you'll implement layer normalization. This step should be relatively straightforward, as conceptually the implementation is almost identical to that of batch normalization. One significant difference though is that for layer normalization, we do not keep track of the moving moments, and the testing phase is identical to the training phase, where the mean and variance are directly calculated per datapoint.
Here's what you need to do:
* In `cs231n/layers.py`, implement the forward pass for layer normalization in the function `layernorm_backward`.
Run the cell below to check your results.
* In `cs231n/layers.py`, implement the backward pass for layer normalization in the function `layernorm_backward`.
Run the second cell below to check your results.
* Modify `cs231n/classifiers/fc_net.py` to add layer normalization to the `FullyConnectedNet`. When the `normalization` flag is set to `"layernorm"` in the constructor, you should insert a layer normalization layer before each ReLU nonlinearity.
Run the third cell below to run the batch size experiment on layer normalization.
```
# Check the training-time forward pass by checking means and variances
# of features both before and after layer normalization
# Simulate the forward pass for a two-layer network
np.random.seed(231)
N, D1, D2, D3 =4, 50, 60, 3
X = np.random.randn(N, D1)
W1 = np.random.randn(D1, D2)
W2 = np.random.randn(D2, D3)
a = np.maximum(0, X.dot(W1)).dot(W2)
print('Before layer normalization:')
print_mean_std(a,axis=1)
gamma = np.ones(D3)
beta = np.zeros(D3)
# Means should be close to zero and stds close to one
print('After layer normalization (gamma=1, beta=0)')
a_norm, _ = layernorm_forward(a, gamma, beta, {'mode': 'train'})
print_mean_std(a_norm,axis=1)
gamma = np.asarray([3.0,3.0,3.0])
beta = np.asarray([5.0,5.0,5.0])
# Now means should be close to beta and stds close to gamma
print('After layer normalization (gamma=', gamma, ', beta=', beta, ')')
a_norm, _ = layernorm_forward(a, gamma, beta, {'mode': 'train'})
print_mean_std(a_norm,axis=1)
# Gradient check batchnorm backward pass
np.random.seed(231)
N, D = 4, 5
x = 5 * np.random.randn(N, D) + 12
gamma = np.random.randn(D)
beta = np.random.randn(D)
dout = np.random.randn(N, D)
ln_param = {}
fx = lambda x: layernorm_forward(x, gamma, beta, ln_param)[0]
fg = lambda a: layernorm_forward(x, a, beta, ln_param)[0]
fb = lambda b: layernorm_forward(x, gamma, b, ln_param)[0]
dx_num = eval_numerical_gradient_array(fx, x, dout)
da_num = eval_numerical_gradient_array(fg, gamma.copy(), dout)
db_num = eval_numerical_gradient_array(fb, beta.copy(), dout)
_, cache = layernorm_forward(x, gamma, beta, ln_param)
dx, dgamma, dbeta = layernorm_backward(dout, cache)
#You should expect to see relative errors between 1e-12 and 1e-8
print('dx error: ', rel_error(dx_num, dx))
print('dgamma error: ', rel_error(da_num, dgamma))
print('dbeta error: ', rel_error(db_num, dbeta))
```
# Layer Normalization and batch size
We will now run the previous batch size experiment with layer normalization instead of batch normalization. Compared to the previous experiment, you should see a markedly smaller influence of batch size on the training history!
```
ln_solvers_bsize, solver_bsize, batch_sizes = run_batchsize_experiments('layernorm')
plt.subplot(2, 1, 1)
plot_training_history('Training accuracy (Layer Normalization)','Epoch', solver_bsize, ln_solvers_bsize, \
lambda x: x.train_acc_history, bl_marker='-^', bn_marker='-o', labels=batch_sizes)
plt.subplot(2, 1, 2)
plot_training_history('Validation accuracy (Layer Normalization)','Epoch', solver_bsize, ln_solvers_bsize, \
lambda x: x.val_acc_history, bl_marker='-^', bn_marker='-o', labels=batch_sizes)
plt.gcf().set_size_inches(15, 10)
plt.show()
```
## Inline Question 4:
When is layer normalization likely to not work well, and why?
1. Using it in a very deep network
2. Having a very small dimension of features
3. Having a high regularization term
## Answer:
[2] Since the sample mean and variance are computed over the features of the input, a small dimension of features will lead to features with a large scale of values influencing the computation unproportionally to its importance.
| github_jupyter |
## SLU05 - Functions Intermediate: Exercise notebook
```
# Students don't need to worry about this cell
# Just make sure that the first thing you do is
# to run it or the rest of the grading won't work!
import math
import hashlib
import inspect
def _hash(s):
return hashlib.blake2b(
bytes(str(s), encoding='utf8'),
digest_size=5
).hexdigest()
```
## Exercise 1
Open up your favorite editor (word, google docs, notes, whatever) and write answers to the following questions in your own words:
1. Explain what local scope is
1. Explain what global scope is
What you want to be able to do is get to the point where you can explain it (even if you don't understand it yet) using the right words and concepts. Having a good idea yourself about what all of these are is essential.
## What is local scope
Write your answer here
## What is global scope
Write your answer here
## Exercise 2
Consider the following code:
```
def hello_world():
a_var = 10
return a_var
another_var = 100
```
What please identify which variable is in a local scope and which variale is in global scope and what variable is in
```
# assign the string values of either 'local' or 'global'
# Assign a_var_scope the value of 'local' or 'global'
a_var_scope = ''
# Assign another_var_scope the value of 'local' or 'global'
another_var_scope = ''
# YOUR CODE HERE
raise NotImplementedError()
assert _hash(a_var_scope) == '702153dc06', 'incorrect scope, try again!'
assert _hash(another_var_scope) == 'ec9f60ee7b', 'incorrect scope, try again!'
```
## Exercise 3
In this exercise, you'll learn the difference between scopes regardless of the name of a variable. The name of the variable is not the important part, it's all about being inside or outside of a function when the variable is assigned it's value.
```
outside_var = 100
def hello_world_complicated():
print(outside_var)
a_var = 10
outside_var = 200
print(outside_var)
# The print(outside_var) which is the first line in the function is printing
# the value of a global or locally scoped variable
# Assign outside_var_first_print_scope the value of 'local' or 'global'
outside_var_first_print_scope = ''
# The print(outside_var) which is the last line in the function is printing
# the value of a global or locally scoped variable
# Assign outside_var_second_print_scope the value of 'local' or 'global'
outside_var_second_print_scope = ''
# YOUR CODE HERE
raise NotImplementedError()
assert _hash('a' + outside_var_first_print_scope) == 'fc6003ae12', 'incorrect scope, try again!'
assert _hash('b' + outside_var_second_print_scope) == '650bfd57ec', 'incorrect scope, try again!'
```
## Exercise 4
Now you'll learn about how local and global scope can clash (and potentially create problems) when a local function accesses things in the global scope.
### DO NOT RUN THIS CODE
The point of this exercise is to read and understand what is happening between the variables and the scopes. You can easily run this code and come up with the answers but if you do without trying to reason about it first, you'll be robbing yourself of a valueable learning activity!
```py
a_list = [1, 2, 3]
def hello():
another_list = a_list
another_list.append(4)
my_value_1 = a_list + [5]
hello()
my_value_2 = a_list
```
```
# Fill in the values by ONLY READING the code.
# DO NOT RUN THE CELL ABOVE
my_value_1 = [] # fill in the values in the list for my_value_1
my_value_2 = [] # fill in the values in the list for my_value_2
# YOUR CODE HERE
raise NotImplementedError()
assert _hash(my_value_1) == '90cd62da52', 'not quite, follow the code and the concepts!'
assert _hash(my_value_2) == '76d2a49034', 'not quite, follow the code and the concepts!'
```
## Exercise 5
In this exercise, you will create a variable in the global scope and then use it inside the local scope
of another function. Your instructions are:
1. Create a global variable called `a_number` and assign it the value of `2`
1. Create a function called `use_global_scope` that takes one argument
1. The argument is called `multiply_by` and should be an integer
1. If `multiply_by` is not an integer, the function returns `None`
1. This function should return the value of the global variable `a_number` multiplied by `multiply_by`
```
# first define a global variable called a_number
# next define your function use_global_scope that multiplies a_number by multiply_by
# YOUR CODE HERE
raise NotImplementedError()
sig = inspect.signature(use_global_scope)
source = inspect.getsource(use_global_scope)
params = sig.parameters
assert len(sig.parameters) == 1, 'your function should take one argument'
assert 'multiply_by' in params, 'you must have an argument called my_list'
assert 'a_number' in globals(), 'be sure to define a_number in the global scope'
assert use_global_scope(10) == 20, 'check the spec and your implementation'
assert use_global_scope('hello') == None, 'check the spec and your implementation'
assert 'a_number' in source, 'be sure to use a_number in side of the function'
assert '=' not in source, 'you do not need to assign any values to any variables inside of this function'
assert '10' not in source, 'are you cheating by hard-coding?'
assert '20' not in source, 'no cheating by hard-coding stuff!'
```
## Exercise 6
In this exercise, you will create a function that multiplies all the elements of a list
by a given number
1. Create a function named `mult_by`
1. It takes one positional argument called `my_list`
1. It take one keyword argument called is called `multiply_by` and is a number
1. The default value is `2`
1. The function must multiply all elements of the list by `multiply_by` and return the result
```
# YOUR CODE HERE
raise NotImplementedError()
assert 'mult_by' in globals(), 'did you name the function correctly?'
sig = inspect.signature(mult_by)
source = inspect.getsource(mult_by)
params = sig.parameters
assert len(sig.parameters) == 2, 'your function should define two arguments'
assert 'my_list' in params, 'you must have an argument called my_list'
assert 'multiply_by' in params, 'you must have an argument called multiply_by'
assert sig.parameters['my_list'].default == inspect._empty, 'my_list should be a positional arg'
assert sig.parameters['multiply_by'].default == 2, 'multiply_by should be position and default to 2'
assert (
'8' not in source
and
'10' not in source
), 'are you cheeeeeting?'
assert mult_by([4, 5]) == [8, 10], 'wrong return value, check spec and implementation!'
assert mult_by([4, 5], multiply_by=1) == [4, 5], 'wrong return value, check spec and implementation!'
```
| github_jupyter |
# Chapter 2: Working With Lists
Much of the remainder of this book is dedicated to using data structures to produce analysis that is elegant and efficient. To use the words of economics, you are making a long-term investment in your human capital by working through these exercises. Once you have invested in these fixed-costs, you can work with data at low marginal cost.
If you are familiar with other programming languages, you may be accustomed to working with arrays. An array is must be cast to house particular data types (_float_, _int_, _string_, etc…). By default, Python works with dynamic lists instead of arrays. Dynamic lists are not cast as a particular type.
## Working with Lists
|New Concepts | Description|
| --- | --- |
| Dynamic List | A dynamic list is encapsulated by brackets _([])_. A list is mutable. Elements can be added to or deleted from a list on the fly.|
| List Concatenation | Two lists can be joined together in the same manner that strings are concatenated. |
| List Indexing | Lists are indexed with the first element being indexed as zero and the last element as the length of (number of elements in) the list less one. Indexes are called using brackets – i.e., _lst[0]_ calls the 0th element in the list. |
In later chapters, we will combine lists with dictionaries to essential data structures. We will also work with more efficient and convenient data structures using the numpy and pandas libraries.
Below we make our first lists. One will be empty. Another will contain integers. Another will have floats. Another strings. Another will mix these:
```
#lists.py
empty_list = []
int_list = [1,2,3,4,5]
float_list = [1.0,2.0,3.0,4.0,5.0]
string_list = ["Many words", "impoverished meaning"]
mixed_list = [1,2.0, "Mix it up"]
print(empty_list)
print(int_list)
print(float_list)
print(string_list)
print(mixed_list)
```
Often we will want to transform lists. In the following example, we will concatenate two lists, which means we will join the lists together:
```
#concatenateLists
list1 = [5, 4, 9, 10, 3, 5]
list2 = [6, 3, 2, 1, 5, 3]
join_lists = list1 + list2
print("list1:", list1)
print("list2:", list2)
print(join_lists)
```
We have joined the lists together to make one long list. We can already observe one way in which Python will be useful for helping us to organize data. If we were doing this in a spread sheet, we would have to identify the row and column values of the elements or copy and paste the desired values into new rows or enter formulas into cells. Python accomplishes this for us with much less work.
For a list of numbers, we will usually perform some arithmetic operation or categorize these values in order to identify meaningful subsets within the data. This requires access the elements, which Python allows us to do efficiently.
In the next exercise we will call elements by index number from the same lists we have already made. We will use the list’s append method to make a copy of a list. The append method adds an element to the end of a list.
```
#copyListElementsForLoop.py
list1 = [5, 4, 9, 10, 3, 5]
list2 = [6, 3, 2, 1, 5, 3]
print("list1 elements:", list1[0], list1[1], list1[2], list1[3], list1[4])
print("list2 elements:", list2[0], list2[1], list2[2], list2[3], list2[4])
list3 = []
j = len(list1)
for i in range(j):
list3.append(list1[i])
k = len(list2)
for i in range(k):
list3.append(list2[i])
print("list3 elements:", list3)
```
# For Loops and _range()_
| New Concepts | Description |
| --- | --- |
| _list(obj)_ | List transforms an iterable object, such as a tuple or set, into a dynamic list. |
| _range(j, k , l)_ | Identifies a range of integers from _j _ to _k–1_ separated by some interval _l_. |
|_len(obj)_ | Measure the length of an iterable object. |
We can use a for loop to more efficiently execute this task. As we saw in the last chapter, the for loop will execute a series of elements: for element in list. Often, this list is a range of numbers that represent the index of a dynamic list. For this purpose we call:
```
for i in range(j, k, l):
<execute script>
```
The for loop cycles through all integers of interval _l _ between _j _ and _k - 1_, executing a script for each value. This script may explicitly integrate the value _i_.
If you do not specify a starting value, _j _, the range function assumes that you are calling an array of elements from _0 _ to _j _. Likewise, if you do not specify an interval, _l _, range assumes that this interval is _1 _. Thus, _for i in range(k)_ is interpreted as _for i in range(0, k, 1)_. We will again use the loop in its simplest form, cycling through number from _0 _ to _(k – 1)_, where the length of the list is the value _k _. These cases are illustrated below in _range.py_.
```
#range.py
list1 = list(range(9))
list2 = list(range(-9,9))
list3 = list(range(-9,9,3))
print(list1)
print(list2)
print(list3)
```
The for loop will automatically identify the elements contained in _range()_ without requiring you to call _list()_. This is illustrated below in _forLoopAndRange.py_.
```
#forLoopAndRange.py
for i in range(10):
print(i)
```
Having printed _i for all i in range(0, 10, 1)_, we produce a set of integers from 0 to 9.
If we were only printing index numbers from a range, for loops would not be very useful. For loops can be used to produce a wide variety of outputs. Often, you will call a for loop to cycle through the index of a particular array. Since arrays are indexed starting with 0 and for loops also assume 0 as an initial value, cycling through a list with a for loop is straight-forward. For a list named _A _, just use the command:
```
for i in range(len(A)):
<execute script>
```
This command will call all integers between 0 and 1 less than the length of _A _. In other words, it will call all indexers associated with _A _.
```
#copyListElementsForLoop.py
list1 = [5, 4, 9, 10, 3, 5]
list2 = [6, 3, 2, 1, 5, 3]
print("list1 elements:", list1[0], list1[1], list1[2], list1[3], list1[4])
print("list2 elements:", list2[0], list2[1], list2[2], list2[3], list2[4])
list3 = []
j = len(list1)
for i in range(j):
list3.append(list1[i])
k = len(list2)
for i in range(k):
list3.append(list2[i])
print("list3 elements:", list3)
```
## Creating a New List with Values from Other Lists
| New Concepts | Description |
| --- | --- |
| List Methods i.e., _.append()_, _.insert()_ | List methods append and insert increse the length of a list by adding in element to the list. |
| If Statements | An if statement executes the block of code contained in it if conditions stipulated by the if statement are met (they return True). |
| Else Statement | In the case that the conditions stipulated by an if statement are not met, and else statement executes an alternate block of code |
| Operator i.e., _==_, _!=_, _< _,_> _, _<=_, _>=_ | The operator indicates the condition relating two variables that is to be tested. |
We can extend the exercise by summing the ith elements in each list. In the exercise below, _list3_ is the sum of the ith elements from _list1_ and _list2_.
```
#addListElements.py
list1 = [5, 4, 9, 10, 3, 5]
list2 = [6, 3, 2, 1, 5, 3]
print("list1 elements:", list1[0], list1[1], list1[2], list1[3], list1[4])
print("list2 elements:", list2[0], list2[1], list2[2], list2[3], list2[4])
list3 = []
j = len(list1)
for i in range(j):
list3.append(list1[i] + list2[i])
print("list3:", list3)
```
In the last exercise, we created an empty list, _list3_. We could not fill the list by calling element in it directly, as no elements yet exist in the list. Instead, we use the append method that is owned by the list-object. Alternately, we can use the insert method. It takes the form, _list.insert(index, object)_. This is shown in a later example. We appended the summed values of the first two lists in the order that the elements are ranked. We could have summed them in opposite order by summing element 5, then 4, ..., then 0.
```
#addListElements.py
list1 = [5, 4, 9, 10, 3, 5]
list2 = [6, 3, 2, 1, 5, 3]
print("list1 elements:", list1[0], list1[1], list1[2], list1[3], list1[4])
print("list2 elements:", list2[0], list2[1], list2[2], list2[3], list2[4])
list3 = []
j = len(list1)
for i in range(j):
list3.insert(0,list1[i] + list2[i])
print("list3:", list3)
```
In the next exercise we will us a function that we have not used before. We will check the length of each list whose elements are summed. We want to make sure that if we call an index from one list, it exists in the other. We do not want to call a list index if it does not exist. That would produce an error.
We can check if a statement is true using an if statement. As with the for loop, the if statement is followed by a colon. This tells the program that the execution below or in front of the if statement depends upon the truth of the condition specified. The code that follows below an if statement must be indented, as this identifies what block of code is subject to the statement.
```
if True:
print("execute script")
```
If the statement returns _True_, then the commands that follow the if-statement will be executed. Though not stated explicitly, we can think of the program as passing over the if statement to the remainder of the script:
```
if True:
print("execute script")
else:
pass
```
If the statement returns _False_, then the program will continue reading the script.
```
if False:
print("execute script")
else:
pass
```
Nothing is printed in the console since there is no further script to execute.
We will want to check if the lengths of two different lists are the same. To check that a variable has a stipulated value, we use two equals signs. Using _==_ allows the program to compare two values rather setting the value of the variable on the left, as would occur with only one equals sign.
Following the if statement is a for loop. If the length of _list1_ and _list2_ are equal, the program will set the ith element of _list3_ equal to the sum of the ith elements from _list1_ and _list2_. In this example, the for loop will cycle through index values 0, 1, 2, 3, 4, and 5.
We can take advantage of the for loop to use _.insert()_ in a manner that replicates the effect of our use of _append()_. We will insert the sum of the ith elements of _list1_ and _list2_ at the ith element of _list3_.
```
#addListElements3.py
list1 = [5, 4, 9, 10, 3, 5]
list2 = [6, 3, 2, 1, 5, 3]
print("list1 elements:", list1[0], list1[1], list1[2], list1[3], list1[4])
print("list2 elements:", list2[0], list2[1], list2[2], list2[3], list2[4])
list3 = []
j = len(list1)
if j == len(list2):
for i in range(0, len(list2)):
list3.insert(i,list1[i] + list2[i])
print("list3:", list3)
```
The if condition may be followed by an else statement. This tells the program to run a different command if the condition of the if statement is not met. In this case, we want the program to tell us why the condition was not met. In other cases, you may want to create other if statements to create a tree of possible outcomes. Below we use an if-else statement to identify when list’s are not the same length. We remove the last element from _list2_ to create lists of different lengths:
```
#addListElements4.py
list1 = [5, 4, 9, 10, 3, 5]
list2 = [6, 3, 2, 1, 5]
print("list1 elements:", list1[0], list1[1], list1[2], list1[3], list1[4])
print("list2 elements:", list2[0], list2[1], list2[2], list2[3], list2[4])
list3 = []
j = len(list1)
if j == len(list2):
for i in range(0, len(list2)):
list3.insert(i,list1[i] + list2[i])
else:
print("Lists are not the same length, cannot perform element-wise operations.")
print("list3:", list3)
```
Since the condition passed to the if statement was false, no values were appended to *list3*.
## Removing List Elements
| New Concepts | Description |
| --- | --- |
| _del_ | The command del is used to delete an element from a list |
|List Methods i.e., _.pop()_, _.remove()_, _.append()_ | Lists contains methods that can be used to modify the list. These include _.pop()_ which removes the last element of a list, allowing it to be saved as a separate object. Another method, _.remove()_ deletes an explicitly identified element. _.append(x)_ adds an additional element at the end of the list. |
Perhaps you want to remove an element from a list. There are a few means of accomplishing this. Which one you choose depends on the ends desired.
```
#deleteListElements.py
list1 = ["red", "blue", "orange", "black", "white", "golden"]
list2 = ["nose", "ice", "fire", "cat", "mouse", "dog"]
print("lists before deletion: ")
for i in range(len(list1)):
print(list1[i],"\t", list2[i])
del list1[0]
del list2[5]
print()
print("lists after deletion: ")
for i in range(len(list1)):
print(list1[i], "\t",list2[i])
```
We have deleted _"red"_ from _list1_ and _"dog"_ from _list2_. By printing the elements of each list once before and once after one element is deleted from each, we can note the difference in the lists over time.
What if we knew that we wanted to remove the elements but did not want to check what index each element is associated with? We can use the remove function owned by each list. We will tell _list1_ to remove _"red"_ and _list2_ to remove _"dog"_.
```
#removeListElements.py
list1 = ["red", "blue", "orange", "black", "white", "golden"]
list2 = ["nose", "ice", "fire", "cat", "mouse", "dog"]
print("lists before deletion: ")
for i in range(len(list1)):
print(list1[i],"\t", list2[i])
list1.remove("red")
list2.remove("dog")
print()
print("lists after deletion: ")
for i in range(len(list1)):
print(list1[i], "\t",list2[i])
```
We have achieved the same result using a different means. What if we wanted to keep track of the element that we removed? Before deleting or removing the element, we could assign the value to a different object. Let's do this before using the remove function:
```
#removeAndSaveListElementsPop.py
#define list1 and list2
list1 = ["red", "blue", "orange", "black", "white", "golden"]
list2 = ["nose", "ice", "fire", "cat", "mouse", "dog"]
#identify what is printed in for loop
print("lists before deletion: ")
if len(list1) == len(list2):
# use for loop to print lists in parallel
for i in range(len(list1)):
print(list1[i],"\t", list2[i])
# remove list elements and save them as variables '_res"
list1_res = "red"
list2_res = "dog"
list1.remove(list1_res)
list2.remove(list2_res)
print()
# print lists again as in lines 8-11
print("lists after deletion: ")
if len(list1) == len(list2):
for i in range(len(list1)):
print(list1[i], "\t",list2[i])
print()
print("Res1", "\tRes2")
print(list1_res, "\t" + (list2_res))
```
An easier way to accomplish this is to use _.pop_, another method owned by each list.
```
#removeListElementsPop.py
#define list1 and list2
list1 = ["red", "blue", "orange", "black", "white", "golden"]
list2 = ["nose", "ice", "fire", "cat", "mouse", "dog"]
#identify what is printed in for loop
print("lists before deletion: ")
# use for loop to print lists in parallel
for i in range(len(list1)):
print(list1[i],"\t", list2[i])
# remove list elements and save them as variables '_res"
list1_res = list1.pop(0)
list2_res = list2.pop(5)
print()
# print lists again as in lines 8-11
print("lists after deletion: ")
for i in range(len(list1)):
print(list1[i], "\t",list2[i])
print()
print("Res1", "\tRes2")
print(list1_res, "\t" + (list2_res))
```
## More with For Loops
When you loop through element values, it is not necessary that these are consecutive. You may skip values at some interval. The next example returns to the earlier _addListElements#.py_ examples. This time, we pass the number 2 as the third element in _range()_. Now range will count by twos from _0 _ to _j – 1_. This will make _list3_ shorter than before.
```
#addListElements5.py
list1 = [5, 4, 9, 10, 3, 5]
list2 = [6, 3, 2, 1, 5, 3]
print("list1 elements:", list1[0], list1[1], list1[2], list1[3], list1[4])
print("list2 elements:", list2[0], list2[1], list2[2], list2[3], list2[4])
list3 = []
j = len(list1)
if j == len(list2):
for i in range(0, j, 2):
list3.append(list1[i] + list2[i])
else:
print("Lists are not the same length, cannot perform element-wise operations.")
print("list3:", list3)
```
We entered the sum of elements 0, 2, and 4 from _list1_ and _list2_ into _list3_. Since these were appended to _list3_, they are indexed in _list3[0]_, _list3[1]_, and _list3[2]_.
For loops in python can call in sequence element of objects that are iterable. These include lists, strings, keys and values from dictionaries, as well as the range function we have already used. You may use a for loop that calls each element in the list without identifying the index of each element.
```
obj = ["A", "few", "words", "to", "print"]
for x in obj:
print(x)
```
Each _x_A _ called is an element from _obj_. Where before we passed _len(list1)_ to the for loop, we now pass _list1_ itself to the for loop and append each element _x _ to _list2_.
```
#forLoopWithoutIndexer.py
list1 = ["red", "blue", "orange", "black", "white", "golden"]
list2 = []
for x in list1:
list2.append(x)
print("list1\t", "list2")
k = len(list1)
j = len(list2)
if len(list1) == len(list2):
for i in range(0, len(list1)):
print(list1[i], "\t", list2[i])
```
## Sorting Lists, Errors, and Exceptions
| New Concepts | Description |
| --- | --- |
| _sorted()_ | The function sorted() sorts a list in order of numerical or alphabetical value. |
| passing errors i.e., _try_ and _except_ | A try statement will pass over an error if one is generated by the code in the try block. In the case that an error is passed, code from the except block well be called. This should typically identify the type of error that was passed. |
We can sort lists using the sorted list function that orders the list either by number or alphabetically. We reuse lists from the last examples to show this.
```
#sorting.py
list1 = [5, 4, 9, 10, 3, 5]
list2 = ["red", "blue", "orange", "black", "white", "golden"]
print("list1:", list1)
print("list2:", list2)
sorted_list1 = sorted(list1)
sorted_list2 = sorted(list2)
print("sortedList1:", sorted_list1)
print("sortedList2:", sorted_list2)
```
What happens if we try to sort a that has both strings and integers? You might expect that Python would sort integers and then strings or vice versa. If you try this, you will raise an error:
```
#sortingError.py
list1 = [5, 4, 9, 10, 3, 5]
list2 = ["red", "blue", "orange", "black", "white", "golden"]
list3 = list1 + list2
print("list1:", list1)
print("list2:", list2)
print("list3:", list3)
sorted_list1 = sorted(list1)
sorted_list2 = sorted(list2)
print("sortedList1:", sorted_list1)
print("sortedList2:", sorted_list2)
sorted_list3 = sorted(list3)
print("sortedList3:", sorted_list3)
print("Execution complete!")
```
The script returns an error. If this error is raised during execution, it will interrupt the program. One way to deal with this is to ask Python to try to execute some script and to execute some other command if an error would normally be raised:
```
#sortingError.py
list1 = [5, 4, 9, 10, 3, 5]
list2 = ["red", "blue", "orange", "black", "white", "golden"]
list3 = list1 + list2
print("list1:", list1)
print("list2:", list2)
print("list3:", list3)
sorted_list1 = sorted(list1)
sorted_list2 = sorted(list2)
print("sortedList1:", sorted_list1)
print("sortedList2:", sorted_list2)
try:
sorted_list3 = sorted(list3)
print("sortedList3:", sorted_list3)
except:
print("TypeError: unorderable types: str() < int() "
"ignoring error")
print("Execution complete!")
```
We successfully avoided the error and instead called an alternate operation defined under except. The use for this will become more obvious as we move along. We will except use them from time to time and note the reason when we do.
## Slicing a List
| New Concepts | Description |
| --- | --- |
| slice i.e., _list\[a:b\]_|A slice of a list is a copy of a portion (or all) of a list from index a to b – 1.|
Sometimes, we may want to access several elements instantly. Python allows us to do this with a slice. Technically, when you call a list in its entirety, you take a slice that includes the entire list. We can do this explicitly like this:
```
#fullSlice.py
some_list = [3, 1, 5, 6, 1]
print(some_list[:])
```
Using *some_list\[:\]* is equivalent of creating a slice using *some_list\[min_index: list_length\]* where *min_index = 0* and *list_length= len(some_list)*:
```
#fullSlice2.py
some_list = [3, 1, 5, 6, 1]
min_index = 0
max_index = len(some_list)
print("minimum:", min_index)
print("maximum:", max_index)
print("Full list using slice", some_list[min_index:max_index])
print("Full list without slice", some_list)
```
This is not very useful if we do not use this to take a smaller subsection of a list. Below, we create a new array that is a subset of the original array. As you might expect by now, *full_list\[7\]* calls the 8th element. Since indexing begins with the 0th element, this element is actually counted as the 7th element. Also, similar to the command *for i in range(3, 7)*, the slice calls elements 3, 4, 5, and 6:
```
#partialSlice.py
min_index = 3
max_index = 7
full_list = [1, 2, 3, 4, 5, 6, 7, 8, 9]
partial_list = full_list[min_index:max_index]
print("Full List:", full_list)
print("Partial List:", partial_list)
print("full_list[7]:", full_list[7])
```
## Nested For Loops
| New Concepts | Description |
| --- | --- |
| Nested For Loops | A for loop may contain other for loops. They are useful for multidimensional data structures. |
Creative use of for loops can save the programmer a lot of work. While you should be careful not to create so many layers of for loops and if statements that code is difficult to interpret (“Flat is better than nested”), you should be comfortable with the structure of nested for loops and, eventually, their use in structures like dictionaries and generators.
A useful way to become acquainted with the power of multiple for loops is to identify what each the result of each iteration of nested for loops. In the code below, the first for loop will count from 0 to 4. For each value of *i*, the second for loop will cycle through values 0 to 4 for _j _.
```
#nestedForLoop.py
print("i", "j")
for i in range(5):
for j in range(5):
print(i, j)
```
Often, we will want to employ values generated by for loops in a manner other than printing the values generated directly by the for loops. We may, for example, want to create a new value constructed from _i _ and _j _. Below, this value is constructed as the sum of _i _ and _j _.
```
#nestedForLoop.py
print("i", "j", "i+j")
for i in range(5):
for j in range(5):
val = i + j
print(i, j, val)
```
If we interpret the results as a table, we can better understand the intuition of for loops. Lighter shading indicates lower values of _i _ with shading growing darker as the value of _i _ increases.
| | | | |__j__| | |
| --- | --- | --- | --- | --- | --- | --- |
| | | __0__ | __1__ |__2__|__3__ | __4__ |
| | __0__ | 0 | 1 | 2 | 3 | 4 |
| | __1__ | 1 | 2 | 3 | 4 | 5 |
| __i__ | __2__ | 2 | 3 | 4 | 5 | 6 |
| | __3__ | 3 | 4 | 5 | 6 | 7 |
| | __4__ | 4 | 5 | 6 | 7 | 8 |
## Lists, Lists, and More Lists
| New Concepts | Description |
| --- | --- |
| _min(lst)_ | The function _min()_ returns the lowest value from a list of values passed to it. |
| _max(lst)_ | The function _max()_ returns that highest value from a list of values passed to it. |
| generators i.e., _[val for val in lst]_ |Generators use a nested for loop to create an iterated data structure. |
Lists have some convenient features. You can find the maximum and minimum values in a list with the _min()_ and _max()_ functions:
```
# minMaxFunctions.py
list1 = [20, 30, 40, 50]
max_list_value = max(list1)
min_list_value = min(list1)
print("maximum:", max_list_value, "minimum:", min_list_value)
```
We could have used a for loop to find these values. The program below performs the same task:
```
#minMaxFuntionsByHand.py
list1 = [20, 30, 40, 50]
# initial smallest value is infinite
# will be replaced if a value from the list is lower
min_list_val = float("inf")
# initial largest values is negative infinite
# will be replaced if a value from the list is higher
max_list_val = float("-inf")
for x in list1:
if x < min_list_val:
min_list_val = x
if x > max_list_val:
max_list_val = x
print("maximum:", max_list_val, "minimum:", min_list_val)
```
We chose to make the starting value of min_list_value large and positive and the starting value of *max_list_value* large and negative. The for loop cycles through these values and assigns the value, _x _, from the list to *min_list_value* if the value is less than the current value assigned to *min_list_value* and to *max_list_value* if the value is greater than the current value assigned to *max_list_value*.
Earlier in the chapter, we constructed lists using list comprehension (i.e., the _list()_ function) and by generating lists and setting values with _.append()_ and _.insert()_. We may also use a generator to create a list. Generators are convenient as they provide a compact means of creating a list that is easier to interpret. They follow the same format as the _list()_ function.
```
#listFromGenerator.py
generator = (i for i in range(20))
print(generator)
list1 = list(generator)
print(list1)
list2 = [2 * i for i in range(20)]
print(list2)
```
### Exercises
1. Create a list of numbers 100 elements in length that counts by 3s - i.e., [3,6,9,12,...]
2. Using the list from question 1, create a second list whose elements are the same values converted to strings. hint: use a for loop and the function str().
3. Using the list from question 2, create a variable that concatenates each of the elements in order of index (Hint: result should be like "36912...").
4. Using .pop() and .append(), create a list whose values are the same as the list from question 1 but in reverse order. (Hint: .pop() removes the last element from a list. The value can be save, i.e., x = lst.pop().)
5. Using len(), calculate the midpoint of the list from question 1. Pass this midpoint to slice the list so that the resultant copy includes only the second half of the list from question 1.
6. Create a string that includes only every other element, starting from the 0th element, from the string in question 3 while maintaining the order of these elements (Hint: this can be done be using a for loop whose values are descending).
7. Explain the difference between a dynamic list in Python (usually referred to as a list) and a tuple.
### Exploration
1. Use a generator to create a list of the first 100 prime numbers. Include a paragraph explaining how a generator works.
2. Using a for loop and the pop function, create a list of the values from the list of prime numbers whose values are descending from highest to lowest.
3. Using either of prime numbers, create another list that includes the same numbers but is randomly ordered. Do this without shuffling the initial list (Hint: you will need to import random and import copy). Explain in a paragraph the process that you followed to accomplish the task.
| github_jupyter |
# RadarCOVID-Report
## Data Extraction
```
import datetime
import json
import logging
import os
import shutil
import tempfile
import textwrap
import uuid
import matplotlib.pyplot as plt
import matplotlib.ticker
import numpy as np
import pandas as pd
import retry
import seaborn as sns
%matplotlib inline
current_working_directory = os.environ.get("PWD")
if current_working_directory:
os.chdir(current_working_directory)
sns.set()
matplotlib.rcParams["figure.figsize"] = (15, 6)
extraction_datetime = datetime.datetime.utcnow()
extraction_date = extraction_datetime.strftime("%Y-%m-%d")
extraction_previous_datetime = extraction_datetime - datetime.timedelta(days=1)
extraction_previous_date = extraction_previous_datetime.strftime("%Y-%m-%d")
extraction_date_with_hour = datetime.datetime.utcnow().strftime("%Y-%m-%d@%H")
current_hour = datetime.datetime.utcnow().hour
are_today_results_partial = current_hour != 23
```
### Constants
```
from Modules.ExposureNotification import exposure_notification_io
spain_region_country_code = "ES"
germany_region_country_code = "DE"
default_backend_identifier = spain_region_country_code
backend_generation_days = 7 * 2
daily_summary_days = 7 * 4 * 3
daily_plot_days = 7 * 4
tek_dumps_load_limit = daily_summary_days + 1
```
### Parameters
```
environment_backend_identifier = os.environ.get("RADARCOVID_REPORT__BACKEND_IDENTIFIER")
if environment_backend_identifier:
report_backend_identifier = environment_backend_identifier
else:
report_backend_identifier = default_backend_identifier
report_backend_identifier
environment_enable_multi_backend_download = \
os.environ.get("RADARCOVID_REPORT__ENABLE_MULTI_BACKEND_DOWNLOAD")
if environment_enable_multi_backend_download:
report_backend_identifiers = None
else:
report_backend_identifiers = [report_backend_identifier]
report_backend_identifiers
environment_invalid_shared_diagnoses_dates = \
os.environ.get("RADARCOVID_REPORT__INVALID_SHARED_DIAGNOSES_DATES")
if environment_invalid_shared_diagnoses_dates:
invalid_shared_diagnoses_dates = environment_invalid_shared_diagnoses_dates.split(",")
else:
invalid_shared_diagnoses_dates = []
invalid_shared_diagnoses_dates
```
### COVID-19 Cases
```
report_backend_client = \
exposure_notification_io.get_backend_client_with_identifier(
backend_identifier=report_backend_identifier)
@retry.retry(tries=10, delay=10, backoff=1.1, jitter=(0, 10))
def download_cases_dataframe_from_ecdc():
return pd.read_csv(
"https://opendata.ecdc.europa.eu/covid19/casedistribution/csv/data.csv")
confirmed_df_ = download_cases_dataframe_from_ecdc()
confirmed_df = confirmed_df_.copy()
confirmed_df = confirmed_df[["dateRep", "cases", "geoId"]]
confirmed_df.rename(
columns={
"dateRep":"sample_date",
"cases": "new_cases",
"geoId": "country_code",
},
inplace=True)
confirmed_df["sample_date"] = pd.to_datetime(confirmed_df.sample_date, dayfirst=True)
confirmed_df["sample_date"] = confirmed_df.sample_date.dt.strftime("%Y-%m-%d")
confirmed_df.sort_values("sample_date", inplace=True)
confirmed_df.tail()
def sort_source_regions_for_display(source_regions: list) -> list:
if report_backend_identifier in source_regions:
source_regions = [report_backend_identifier] + \
list(sorted(set(source_regions).difference([report_backend_identifier])))
else:
source_regions = list(sorted(source_regions))
return source_regions
report_source_regions = report_backend_client.source_regions_for_date(
date=extraction_datetime.date())
report_source_regions = sort_source_regions_for_display(
source_regions=report_source_regions)
report_source_regions
confirmed_days = pd.date_range(
start=confirmed_df.iloc[0].sample_date,
end=extraction_datetime)
confirmed_days_df = pd.DataFrame(data=confirmed_days, columns=["sample_date"])
confirmed_days_df["sample_date_string"] = \
confirmed_days_df.sample_date.dt.strftime("%Y-%m-%d")
confirmed_days_df.tail()
source_regions_at_date_df = confirmed_days_df.copy()
source_regions_at_date_df["source_regions_at_date"] = \
source_regions_at_date_df.sample_date.apply(
lambda x: report_backend_client.source_regions_for_date(date=x))
source_regions_at_date_df.sort_values("sample_date", inplace=True)
source_regions_at_date_df["_source_regions_group"] = source_regions_at_date_df. \
source_regions_at_date.apply(lambda x: ",".join(sort_source_regions_for_display(x)))
source_regions_at_date_df.tail()
source_regions_for_summary_df = \
source_regions_at_date_df[["sample_date", "_source_regions_group"]].copy()
source_regions_for_summary_df.rename(columns={"_source_regions_group": "source_regions"}, inplace=True)
source_regions_for_summary_df.tail()
confirmed_output_columns = ["sample_date", "new_cases", "covid_cases"]
confirmed_output_df = pd.DataFrame(columns=confirmed_output_columns)
for source_regions_group, source_regions_group_series in \
source_regions_at_date_df.groupby("_source_regions_group"):
source_regions_set = set(source_regions_group.split(","))
confirmed_source_regions_set_df = \
confirmed_df[confirmed_df.country_code.isin(source_regions_set)].copy()
confirmed_source_regions_group_df = \
confirmed_source_regions_set_df.groupby("sample_date").new_cases.sum() \
.reset_index().sort_values("sample_date")
confirmed_source_regions_group_df["covid_cases"] = \
confirmed_source_regions_group_df.new_cases.rolling(7, min_periods=0).mean().round()
confirmed_source_regions_group_df = \
confirmed_source_regions_group_df[confirmed_output_columns]
confirmed_source_regions_group_df.fillna(method="ffill", inplace=True)
confirmed_source_regions_group_df = \
confirmed_source_regions_group_df[
confirmed_source_regions_group_df.sample_date.isin(
source_regions_group_series.sample_date_string)]
confirmed_output_df = confirmed_output_df.append(confirmed_source_regions_group_df)
confirmed_df = confirmed_output_df.copy()
confirmed_df.tail()
confirmed_df.rename(columns={"sample_date": "sample_date_string"}, inplace=True)
confirmed_df = confirmed_days_df[["sample_date_string"]].merge(confirmed_df, how="left")
confirmed_df.sort_values("sample_date_string", inplace=True)
confirmed_df.fillna(method="ffill", inplace=True)
confirmed_df.tail()
confirmed_df[["new_cases", "covid_cases"]].plot()
```
### Extract API TEKs
```
raw_zip_path_prefix = "Data/TEKs/Raw/"
fail_on_error_backend_identifiers = [report_backend_identifier]
multi_backend_exposure_keys_df = \
exposure_notification_io.download_exposure_keys_from_backends(
backend_identifiers=report_backend_identifiers,
generation_days=backend_generation_days,
fail_on_error_backend_identifiers=fail_on_error_backend_identifiers,
save_raw_zip_path_prefix=raw_zip_path_prefix)
multi_backend_exposure_keys_df["region"] = multi_backend_exposure_keys_df["backend_identifier"]
multi_backend_exposure_keys_df.rename(
columns={
"generation_datetime": "sample_datetime",
"generation_date_string": "sample_date_string",
},
inplace=True)
multi_backend_exposure_keys_df.head()
early_teks_df = multi_backend_exposure_keys_df[
multi_backend_exposure_keys_df.rolling_period < 144].copy()
early_teks_df["rolling_period_in_hours"] = early_teks_df.rolling_period / 6
early_teks_df[early_teks_df.sample_date_string != extraction_date] \
.rolling_period_in_hours.hist(bins=list(range(24)))
early_teks_df[early_teks_df.sample_date_string == extraction_date] \
.rolling_period_in_hours.hist(bins=list(range(24)))
multi_backend_exposure_keys_df = multi_backend_exposure_keys_df[[
"sample_date_string", "region", "key_data"]]
multi_backend_exposure_keys_df.head()
active_regions = \
multi_backend_exposure_keys_df.groupby("region").key_data.nunique().sort_values().index.unique().tolist()
active_regions
multi_backend_summary_df = multi_backend_exposure_keys_df.groupby(
["sample_date_string", "region"]).key_data.nunique().reset_index() \
.pivot(index="sample_date_string", columns="region") \
.sort_index(ascending=False)
multi_backend_summary_df.rename(
columns={"key_data": "shared_teks_by_generation_date"},
inplace=True)
multi_backend_summary_df.rename_axis("sample_date", inplace=True)
multi_backend_summary_df = multi_backend_summary_df.fillna(0).astype(int)
multi_backend_summary_df = multi_backend_summary_df.head(backend_generation_days)
multi_backend_summary_df.head()
def compute_keys_cross_sharing(x):
teks_x = x.key_data_x.item()
common_teks = set(teks_x).intersection(x.key_data_y.item())
common_teks_fraction = len(common_teks) / len(teks_x)
return pd.Series(dict(
common_teks=common_teks,
common_teks_fraction=common_teks_fraction,
))
multi_backend_exposure_keys_by_region_df = \
multi_backend_exposure_keys_df.groupby("region").key_data.unique().reset_index()
multi_backend_exposure_keys_by_region_df["_merge"] = True
multi_backend_exposure_keys_by_region_combination_df = \
multi_backend_exposure_keys_by_region_df.merge(
multi_backend_exposure_keys_by_region_df, on="_merge")
multi_backend_exposure_keys_by_region_combination_df.drop(
columns=["_merge"], inplace=True)
if multi_backend_exposure_keys_by_region_combination_df.region_x.nunique() > 1:
multi_backend_exposure_keys_by_region_combination_df = \
multi_backend_exposure_keys_by_region_combination_df[
multi_backend_exposure_keys_by_region_combination_df.region_x !=
multi_backend_exposure_keys_by_region_combination_df.region_y]
multi_backend_exposure_keys_cross_sharing_df = \
multi_backend_exposure_keys_by_region_combination_df \
.groupby(["region_x", "region_y"]) \
.apply(compute_keys_cross_sharing) \
.reset_index()
multi_backend_cross_sharing_summary_df = \
multi_backend_exposure_keys_cross_sharing_df.pivot_table(
values=["common_teks_fraction"],
columns="region_x",
index="region_y",
aggfunc=lambda x: x.item())
multi_backend_cross_sharing_summary_df
multi_backend_without_active_region_exposure_keys_df = \
multi_backend_exposure_keys_df[multi_backend_exposure_keys_df.region != report_backend_identifier]
multi_backend_without_active_region = \
multi_backend_without_active_region_exposure_keys_df.groupby("region").key_data.nunique().sort_values().index.unique().tolist()
multi_backend_without_active_region
exposure_keys_summary_df = multi_backend_exposure_keys_df[
multi_backend_exposure_keys_df.region == report_backend_identifier]
exposure_keys_summary_df.drop(columns=["region"], inplace=True)
exposure_keys_summary_df = \
exposure_keys_summary_df.groupby(["sample_date_string"]).key_data.nunique().to_frame()
exposure_keys_summary_df = \
exposure_keys_summary_df.reset_index().set_index("sample_date_string")
exposure_keys_summary_df.sort_index(ascending=False, inplace=True)
exposure_keys_summary_df.rename(columns={"key_data": "shared_teks_by_generation_date"}, inplace=True)
exposure_keys_summary_df.head()
```
### Dump API TEKs
```
tek_list_df = multi_backend_exposure_keys_df[
["sample_date_string", "region", "key_data"]].copy()
tek_list_df["key_data"] = tek_list_df["key_data"].apply(str)
tek_list_df.rename(columns={
"sample_date_string": "sample_date",
"key_data": "tek_list"}, inplace=True)
tek_list_df = tek_list_df.groupby(
["sample_date", "region"]).tek_list.unique().reset_index()
tek_list_df["extraction_date"] = extraction_date
tek_list_df["extraction_date_with_hour"] = extraction_date_with_hour
tek_list_path_prefix = "Data/TEKs/"
tek_list_current_path = tek_list_path_prefix + f"/Current/RadarCOVID-TEKs.json"
tek_list_daily_path = tek_list_path_prefix + f"Daily/RadarCOVID-TEKs-{extraction_date}.json"
tek_list_hourly_path = tek_list_path_prefix + f"Hourly/RadarCOVID-TEKs-{extraction_date_with_hour}.json"
for path in [tek_list_current_path, tek_list_daily_path, tek_list_hourly_path]:
os.makedirs(os.path.dirname(path), exist_ok=True)
tek_list_df.drop(columns=["extraction_date", "extraction_date_with_hour"]).to_json(
tek_list_current_path,
lines=True, orient="records")
tek_list_df.drop(columns=["extraction_date_with_hour"]).to_json(
tek_list_daily_path,
lines=True, orient="records")
tek_list_df.to_json(
tek_list_hourly_path,
lines=True, orient="records")
tek_list_df.head()
```
### Load TEK Dumps
```
import glob
def load_extracted_teks(mode, region=None, limit=None) -> pd.DataFrame:
extracted_teks_df = pd.DataFrame(columns=["region"])
file_paths = list(reversed(sorted(glob.glob(tek_list_path_prefix + mode + "/RadarCOVID-TEKs-*.json"))))
if limit:
file_paths = file_paths[:limit]
for file_path in file_paths:
logging.info(f"Loading TEKs from '{file_path}'...")
iteration_extracted_teks_df = pd.read_json(file_path, lines=True)
extracted_teks_df = extracted_teks_df.append(
iteration_extracted_teks_df, sort=False)
extracted_teks_df["region"] = \
extracted_teks_df.region.fillna(spain_region_country_code).copy()
if region:
extracted_teks_df = \
extracted_teks_df[extracted_teks_df.region == region]
return extracted_teks_df
daily_extracted_teks_df = load_extracted_teks(
mode="Daily",
region=report_backend_identifier,
limit=tek_dumps_load_limit)
daily_extracted_teks_df.head()
exposure_keys_summary_df_ = daily_extracted_teks_df \
.sort_values("extraction_date", ascending=False) \
.groupby("sample_date").tek_list.first() \
.to_frame()
exposure_keys_summary_df_.index.name = "sample_date_string"
exposure_keys_summary_df_["tek_list"] = \
exposure_keys_summary_df_.tek_list.apply(len)
exposure_keys_summary_df_ = exposure_keys_summary_df_ \
.rename(columns={"tek_list": "shared_teks_by_generation_date"}) \
.sort_index(ascending=False)
exposure_keys_summary_df = exposure_keys_summary_df_
exposure_keys_summary_df.head()
```
### Daily New TEKs
```
tek_list_df = daily_extracted_teks_df.groupby("extraction_date").tek_list.apply(
lambda x: set(sum(x, []))).reset_index()
tek_list_df = tek_list_df.set_index("extraction_date").sort_index(ascending=True)
tek_list_df.head()
def compute_teks_by_generation_and_upload_date(date):
day_new_teks_set_df = tek_list_df.copy().diff()
try:
day_new_teks_set = day_new_teks_set_df[
day_new_teks_set_df.index == date].tek_list.item()
except ValueError:
day_new_teks_set = None
if pd.isna(day_new_teks_set):
day_new_teks_set = set()
day_new_teks_df = daily_extracted_teks_df[
daily_extracted_teks_df.extraction_date == date].copy()
day_new_teks_df["shared_teks"] = \
day_new_teks_df.tek_list.apply(lambda x: set(x).intersection(day_new_teks_set))
day_new_teks_df["shared_teks"] = \
day_new_teks_df.shared_teks.apply(len)
day_new_teks_df["upload_date"] = date
day_new_teks_df.rename(columns={"sample_date": "generation_date"}, inplace=True)
day_new_teks_df = day_new_teks_df[
["upload_date", "generation_date", "shared_teks"]]
day_new_teks_df["generation_to_upload_days"] = \
(pd.to_datetime(day_new_teks_df.upload_date) -
pd.to_datetime(day_new_teks_df.generation_date)).dt.days
day_new_teks_df = day_new_teks_df[day_new_teks_df.shared_teks > 0]
return day_new_teks_df
shared_teks_generation_to_upload_df = pd.DataFrame()
for upload_date in daily_extracted_teks_df.extraction_date.unique():
shared_teks_generation_to_upload_df = \
shared_teks_generation_to_upload_df.append(
compute_teks_by_generation_and_upload_date(date=upload_date))
shared_teks_generation_to_upload_df \
.sort_values(["upload_date", "generation_date"], ascending=False, inplace=True)
shared_teks_generation_to_upload_df.tail()
today_new_teks_df = \
shared_teks_generation_to_upload_df[
shared_teks_generation_to_upload_df.upload_date == extraction_date].copy()
today_new_teks_df.tail()
if not today_new_teks_df.empty:
today_new_teks_df.set_index("generation_to_upload_days") \
.sort_index().shared_teks.plot.bar()
generation_to_upload_period_pivot_df = \
shared_teks_generation_to_upload_df[
["upload_date", "generation_to_upload_days", "shared_teks"]] \
.pivot(index="upload_date", columns="generation_to_upload_days") \
.sort_index(ascending=False).fillna(0).astype(int) \
.droplevel(level=0, axis=1)
generation_to_upload_period_pivot_df.head()
new_tek_df = tek_list_df.diff().tek_list.apply(
lambda x: len(x) if not pd.isna(x) else None).to_frame().reset_index()
new_tek_df.rename(columns={
"tek_list": "shared_teks_by_upload_date",
"extraction_date": "sample_date_string",}, inplace=True)
new_tek_df.tail()
shared_teks_uploaded_on_generation_date_df = shared_teks_generation_to_upload_df[
shared_teks_generation_to_upload_df.generation_to_upload_days == 0] \
[["upload_date", "shared_teks"]].rename(
columns={
"upload_date": "sample_date_string",
"shared_teks": "shared_teks_uploaded_on_generation_date",
})
shared_teks_uploaded_on_generation_date_df.head()
estimated_shared_diagnoses_df = shared_teks_generation_to_upload_df \
.groupby(["upload_date"]).shared_teks.max().reset_index() \
.sort_values(["upload_date"], ascending=False) \
.rename(columns={
"upload_date": "sample_date_string",
"shared_teks": "shared_diagnoses",
})
invalid_shared_diagnoses_dates_mask = \
estimated_shared_diagnoses_df.sample_date_string.isin(invalid_shared_diagnoses_dates)
estimated_shared_diagnoses_df[invalid_shared_diagnoses_dates_mask] = 0
estimated_shared_diagnoses_df.head()
```
### Hourly New TEKs
```
hourly_extracted_teks_df = load_extracted_teks(
mode="Hourly", region=report_backend_identifier, limit=25)
hourly_extracted_teks_df.head()
hourly_new_tek_count_df = hourly_extracted_teks_df \
.groupby("extraction_date_with_hour").tek_list. \
apply(lambda x: set(sum(x, []))).reset_index().copy()
hourly_new_tek_count_df = hourly_new_tek_count_df.set_index("extraction_date_with_hour") \
.sort_index(ascending=True)
hourly_new_tek_count_df["new_tek_list"] = hourly_new_tek_count_df.tek_list.diff()
hourly_new_tek_count_df["new_tek_count"] = hourly_new_tek_count_df.new_tek_list.apply(
lambda x: len(x) if not pd.isna(x) else 0)
hourly_new_tek_count_df.rename(columns={
"new_tek_count": "shared_teks_by_upload_date"}, inplace=True)
hourly_new_tek_count_df = hourly_new_tek_count_df.reset_index()[[
"extraction_date_with_hour", "shared_teks_by_upload_date"]]
hourly_new_tek_count_df.head()
hourly_summary_df = hourly_new_tek_count_df.copy()
hourly_summary_df.set_index("extraction_date_with_hour", inplace=True)
hourly_summary_df = hourly_summary_df.fillna(0).astype(int).reset_index()
hourly_summary_df["datetime_utc"] = pd.to_datetime(
hourly_summary_df.extraction_date_with_hour, format="%Y-%m-%d@%H")
hourly_summary_df.set_index("datetime_utc", inplace=True)
hourly_summary_df = hourly_summary_df.tail(-1)
hourly_summary_df.head()
```
### Data Merge
```
result_summary_df = exposure_keys_summary_df.merge(
new_tek_df, on=["sample_date_string"], how="outer")
result_summary_df.head()
result_summary_df = result_summary_df.merge(
shared_teks_uploaded_on_generation_date_df, on=["sample_date_string"], how="outer")
result_summary_df.head()
result_summary_df = result_summary_df.merge(
estimated_shared_diagnoses_df, on=["sample_date_string"], how="outer")
result_summary_df.head()
result_summary_df = confirmed_df.tail(daily_summary_days).merge(
result_summary_df, on=["sample_date_string"], how="left")
result_summary_df.head()
result_summary_df["sample_date"] = pd.to_datetime(result_summary_df.sample_date_string)
result_summary_df = result_summary_df.merge(source_regions_for_summary_df, how="left")
result_summary_df.set_index(["sample_date", "source_regions"], inplace=True)
result_summary_df.drop(columns=["sample_date_string"], inplace=True)
result_summary_df.sort_index(ascending=False, inplace=True)
result_summary_df.head()
with pd.option_context("mode.use_inf_as_na", True):
result_summary_df = result_summary_df.fillna(0).astype(int)
result_summary_df["teks_per_shared_diagnosis"] = \
(result_summary_df.shared_teks_by_upload_date / result_summary_df.shared_diagnoses).fillna(0)
result_summary_df["shared_diagnoses_per_covid_case"] = \
(result_summary_df.shared_diagnoses / result_summary_df.covid_cases).fillna(0)
result_summary_df.head(daily_plot_days)
weekly_result_summary_df = result_summary_df \
.sort_index(ascending=True).fillna(0).rolling(7).agg({
"covid_cases": "sum",
"shared_teks_by_generation_date": "sum",
"shared_teks_by_upload_date": "sum",
"shared_diagnoses": "sum"
}).sort_index(ascending=False)
with pd.option_context("mode.use_inf_as_na", True):
weekly_result_summary_df = weekly_result_summary_df.fillna(0).astype(int)
weekly_result_summary_df["teks_per_shared_diagnosis"] = \
(weekly_result_summary_df.shared_teks_by_upload_date / weekly_result_summary_df.shared_diagnoses).fillna(0)
weekly_result_summary_df["shared_diagnoses_per_covid_case"] = \
(weekly_result_summary_df.shared_diagnoses / weekly_result_summary_df.covid_cases).fillna(0)
weekly_result_summary_df.head()
last_7_days_summary = weekly_result_summary_df.to_dict(orient="records")[1]
last_7_days_summary
```
## Report Results
```
display_column_name_mapping = {
"sample_date": "Sample\u00A0Date\u00A0(UTC)",
"source_regions": "Source Countries",
"datetime_utc": "Timestamp (UTC)",
"upload_date": "Upload Date (UTC)",
"generation_to_upload_days": "Generation to Upload Period in Days",
"region": "Backend",
"region_x": "Backend\u00A0(A)",
"region_y": "Backend\u00A0(B)",
"common_teks": "Common TEKs Shared Between Backends",
"common_teks_fraction": "Fraction of TEKs in Backend (A) Available in Backend (B)",
"covid_cases": "COVID-19 Cases in Source Countries (7-day Rolling Average)",
"shared_teks_by_generation_date": "Shared TEKs by Generation Date",
"shared_teks_by_upload_date": "Shared TEKs by Upload Date",
"shared_diagnoses": "Shared Diagnoses (Estimation)",
"teks_per_shared_diagnosis": "TEKs Uploaded per Shared Diagnosis",
"shared_diagnoses_per_covid_case": "Usage Ratio (Fraction of Cases in Source Countries Which Shared Diagnosis)",
"shared_teks_uploaded_on_generation_date": "Shared TEKs Uploaded on Generation Date",
}
summary_columns = [
"covid_cases",
"shared_teks_by_generation_date",
"shared_teks_by_upload_date",
"shared_teks_uploaded_on_generation_date",
"shared_diagnoses",
"teks_per_shared_diagnosis",
"shared_diagnoses_per_covid_case",
]
```
### Daily Summary Table
```
result_summary_df_ = result_summary_df.copy()
result_summary_df = result_summary_df[summary_columns]
result_summary_with_display_names_df = result_summary_df \
.rename_axis(index=display_column_name_mapping) \
.rename(columns=display_column_name_mapping)
result_summary_with_display_names_df
```
### Daily Summary Plots
```
result_plot_summary_df = result_summary_df.head(daily_plot_days)[summary_columns] \
.droplevel(level=["source_regions"]) \
.rename_axis(index=display_column_name_mapping) \
.rename(columns=display_column_name_mapping)
summary_ax_list = result_plot_summary_df.sort_index(ascending=True).plot.bar(
title=f"Daily Summary",
rot=45, subplots=True, figsize=(15, 22), legend=False)
ax_ = summary_ax_list[-1]
ax_.get_figure().tight_layout()
ax_.get_figure().subplots_adjust(top=0.95)
ax_.yaxis.set_major_formatter(matplotlib.ticker.PercentFormatter(1.0))
_ = ax_.set_xticklabels(sorted(result_plot_summary_df.index.strftime("%Y-%m-%d").tolist()))
```
### Daily Generation to Upload Period Table
```
display_generation_to_upload_period_pivot_df = \
generation_to_upload_period_pivot_df \
.head(backend_generation_days)
display_generation_to_upload_period_pivot_df \
.head(backend_generation_days) \
.rename_axis(columns=display_column_name_mapping) \
.rename_axis(index=display_column_name_mapping)
fig, generation_to_upload_period_pivot_table_ax = plt.subplots(
figsize=(12, 1 + 0.6 * len(display_generation_to_upload_period_pivot_df)))
generation_to_upload_period_pivot_table_ax.set_title(
"Shared TEKs Generation to Upload Period Table")
sns.heatmap(
data=display_generation_to_upload_period_pivot_df
.rename_axis(columns=display_column_name_mapping)
.rename_axis(index=display_column_name_mapping),
fmt=".0f",
annot=True,
ax=generation_to_upload_period_pivot_table_ax)
generation_to_upload_period_pivot_table_ax.get_figure().tight_layout()
```
### Hourly Summary Plots
```
hourly_summary_ax_list = hourly_summary_df \
.rename_axis(index=display_column_name_mapping) \
.rename(columns=display_column_name_mapping) \
.plot.bar(
title=f"Last 24h Summary",
rot=45, subplots=True, legend=False)
ax_ = hourly_summary_ax_list[-1]
ax_.get_figure().tight_layout()
ax_.get_figure().subplots_adjust(top=0.9)
_ = ax_.set_xticklabels(sorted(hourly_summary_df.index.strftime("%Y-%m-%d@%H").tolist()))
```
### Publish Results
```
def get_temporary_image_path() -> str:
return os.path.join(tempfile.gettempdir(), str(uuid.uuid4()) + ".png")
def save_temporary_plot_image(ax):
if isinstance(ax, np.ndarray):
ax = ax[0]
media_path = get_temporary_image_path()
ax.get_figure().savefig(media_path)
return media_path
def save_temporary_dataframe_image(df):
import dataframe_image as dfi
media_path = get_temporary_image_path()
dfi.export(df, media_path)
return media_path
github_repository = os.environ.get("GITHUB_REPOSITORY")
if github_repository is None:
github_repository = "pvieito/Radar-STATS"
github_project_base_url = "https://github.com/" + github_repository
display_formatters = {
display_column_name_mapping["teks_per_shared_diagnosis"]: lambda x: f"{x:.2f}",
display_column_name_mapping["shared_diagnoses_per_covid_case"]: lambda x: f"{x:.2%}",
}
daily_summary_table_html = result_summary_with_display_names_df \
.head(daily_plot_days) \
.rename_axis(index=display_column_name_mapping) \
.rename(columns=display_column_name_mapping) \
.to_html(formatters=display_formatters)
multi_backend_summary_table_html = multi_backend_summary_df \
.head(daily_plot_days) \
.rename_axis(columns=display_column_name_mapping) \
.rename(columns=display_column_name_mapping) \
.rename_axis(index=display_column_name_mapping) \
.to_html(formatters=display_formatters)
def format_multi_backend_cross_sharing_fraction(x):
if pd.isna(x):
return "-"
elif round(x * 100, 1) == 0:
return ""
else:
return f"{x:.1%}"
multi_backend_cross_sharing_summary_table_html = multi_backend_cross_sharing_summary_df \
.rename_axis(columns=display_column_name_mapping) \
.rename(columns=display_column_name_mapping) \
.rename_axis(index=display_column_name_mapping) \
.to_html(
classes="table-center",
formatters=display_formatters,
float_format=format_multi_backend_cross_sharing_fraction)
multi_backend_cross_sharing_summary_table_html = \
multi_backend_cross_sharing_summary_table_html \
.replace("<tr>","<tr style=\"text-align: center;\">")
extraction_date_result_summary_df = \
result_summary_df[result_summary_df.index.get_level_values("sample_date") == extraction_date]
extraction_date_result_hourly_summary_df = \
hourly_summary_df[hourly_summary_df.extraction_date_with_hour == extraction_date_with_hour]
covid_cases = \
extraction_date_result_summary_df.covid_cases.item()
shared_teks_by_generation_date = \
extraction_date_result_summary_df.shared_teks_by_generation_date.item()
shared_teks_by_upload_date = \
extraction_date_result_summary_df.shared_teks_by_upload_date.item()
shared_diagnoses = \
extraction_date_result_summary_df.shared_diagnoses.item()
teks_per_shared_diagnosis = \
extraction_date_result_summary_df.teks_per_shared_diagnosis.item()
shared_diagnoses_per_covid_case = \
extraction_date_result_summary_df.shared_diagnoses_per_covid_case.item()
shared_teks_by_upload_date_last_hour = \
extraction_date_result_hourly_summary_df.shared_teks_by_upload_date.sum().astype(int)
display_source_regions = ", ".join(report_source_regions)
if len(report_source_regions) == 1:
display_brief_source_regions = report_source_regions[0]
else:
display_brief_source_regions = f"{len(report_source_regions)} 🇪🇺"
summary_plots_image_path = save_temporary_plot_image(
ax=summary_ax_list)
summary_table_image_path = save_temporary_dataframe_image(
df=result_summary_with_display_names_df)
hourly_summary_plots_image_path = save_temporary_plot_image(
ax=hourly_summary_ax_list)
multi_backend_summary_table_image_path = save_temporary_dataframe_image(
df=multi_backend_summary_df)
generation_to_upload_period_pivot_table_image_path = save_temporary_plot_image(
ax=generation_to_upload_period_pivot_table_ax)
```
### Save Results
```
report_resources_path_prefix = "Data/Resources/Current/RadarCOVID-Report-"
result_summary_df.to_csv(
report_resources_path_prefix + "Summary-Table.csv")
result_summary_df.to_html(
report_resources_path_prefix + "Summary-Table.html")
hourly_summary_df.to_csv(
report_resources_path_prefix + "Hourly-Summary-Table.csv")
multi_backend_summary_df.to_csv(
report_resources_path_prefix + "Multi-Backend-Summary-Table.csv")
multi_backend_cross_sharing_summary_df.to_csv(
report_resources_path_prefix + "Multi-Backend-Cross-Sharing-Summary-Table.csv")
generation_to_upload_period_pivot_df.to_csv(
report_resources_path_prefix + "Generation-Upload-Period-Table.csv")
_ = shutil.copyfile(
summary_plots_image_path,
report_resources_path_prefix + "Summary-Plots.png")
_ = shutil.copyfile(
summary_table_image_path,
report_resources_path_prefix + "Summary-Table.png")
_ = shutil.copyfile(
hourly_summary_plots_image_path,
report_resources_path_prefix + "Hourly-Summary-Plots.png")
_ = shutil.copyfile(
multi_backend_summary_table_image_path,
report_resources_path_prefix + "Multi-Backend-Summary-Table.png")
_ = shutil.copyfile(
generation_to_upload_period_pivot_table_image_path,
report_resources_path_prefix + "Generation-Upload-Period-Table.png")
```
### Publish Results as JSON
```
def generate_summary_api_results(df: pd.DataFrame) -> list:
api_df = df.reset_index().copy()
api_df["sample_date_string"] = \
api_df["sample_date"].dt.strftime("%Y-%m-%d")
api_df["source_regions"] = \
api_df["source_regions"].apply(lambda x: x.split(","))
return api_df.to_dict(orient="records")
summary_api_results = \
generate_summary_api_results(df=result_summary_df)
today_summary_api_results = \
generate_summary_api_results(df=extraction_date_result_summary_df)[0]
summary_results = dict(
backend_identifier=report_backend_identifier,
source_regions=report_source_regions,
extraction_datetime=extraction_datetime,
extraction_date=extraction_date,
extraction_date_with_hour=extraction_date_with_hour,
last_hour=dict(
shared_teks_by_upload_date=shared_teks_by_upload_date_last_hour,
shared_diagnoses=0,
),
today=today_summary_api_results,
last_7_days=last_7_days_summary,
daily_results=summary_api_results)
summary_results = \
json.loads(pd.Series([summary_results]).to_json(orient="records"))[0]
with open(report_resources_path_prefix + "Summary-Results.json", "w") as f:
json.dump(summary_results, f, indent=4)
```
### Publish on README
```
with open("Data/Templates/README.md", "r") as f:
readme_contents = f.read()
readme_contents = readme_contents.format(
extraction_date_with_hour=extraction_date_with_hour,
github_project_base_url=github_project_base_url,
daily_summary_table_html=daily_summary_table_html,
multi_backend_summary_table_html=multi_backend_summary_table_html,
multi_backend_cross_sharing_summary_table_html=multi_backend_cross_sharing_summary_table_html,
display_source_regions=display_source_regions)
with open("README.md", "w") as f:
f.write(readme_contents)
```
### Publish on Twitter
```
enable_share_to_twitter = os.environ.get("RADARCOVID_REPORT__ENABLE_PUBLISH_ON_TWITTER")
github_event_name = os.environ.get("GITHUB_EVENT_NAME")
if enable_share_to_twitter and github_event_name == "schedule" and \
(shared_teks_by_upload_date_last_hour or not are_today_results_partial):
import tweepy
twitter_api_auth_keys = os.environ["RADARCOVID_REPORT__TWITTER_API_AUTH_KEYS"]
twitter_api_auth_keys = twitter_api_auth_keys.split(":")
auth = tweepy.OAuthHandler(twitter_api_auth_keys[0], twitter_api_auth_keys[1])
auth.set_access_token(twitter_api_auth_keys[2], twitter_api_auth_keys[3])
api = tweepy.API(auth)
summary_plots_media = api.media_upload(summary_plots_image_path)
summary_table_media = api.media_upload(summary_table_image_path)
generation_to_upload_period_pivot_table_image_media = api.media_upload(generation_to_upload_period_pivot_table_image_path)
media_ids = [
summary_plots_media.media_id,
summary_table_media.media_id,
generation_to_upload_period_pivot_table_image_media.media_id,
]
if are_today_results_partial:
today_addendum = " (Partial)"
else:
today_addendum = ""
status = textwrap.dedent(f"""
#RadarCOVID – {extraction_date_with_hour}
Source Countries: {display_brief_source_regions}
Today{today_addendum}:
- Uploaded TEKs: {shared_teks_by_upload_date:.0f} ({shared_teks_by_upload_date_last_hour:+d} last hour)
- Shared Diagnoses: ≤{shared_diagnoses:.0f}
- Usage Ratio: ≤{shared_diagnoses_per_covid_case:.2%}
Last 7 Days:
- Shared Diagnoses: ≤{last_7_days_summary["shared_diagnoses"]:.0f}
- Usage Ratio: ≤{last_7_days_summary["shared_diagnoses_per_covid_case"]:.2%}
Info: {github_project_base_url}#documentation
""")
status = status.encode(encoding="utf-8")
api.update_status(status=status, media_ids=media_ids)
```
| github_jupyter |
# 周期驱动有阻尼非线性单摆的混沌特性 #
以前的作业中,我们已经定义了在给定时间区间内求解一般微分方程的函数dSolve,
[dSolve in Chap1](https://github.com/loading99pct/computationalphysics_N2013301020062/tree/master/chap-1)
借助之前的求解方法函数,定义函数dSolveUntil, 此函数不断迭代,直到状态(t,X)满足某个条件stopQ,
返回解列表,此函数结构由nestUntilList给出
已经定义了dSolveUntil,它和dSolve一样可以调用四种求解方法,默认采用 RK-4 求解
[dSolveUntil in Chap2](https://github.com/loading99pct/computationalphysics_N2013301020062/tree/master/Chap_2.1)
本次作业中:
1, 对初值的敏感依赖性
2, 庞加莱截面
## 对初值的敏感依赖性 ##
dSolve可以调用四种求解方法,默认采用 RK-4 求解
定义周期驱动有阻尼非线性单摆的运动微分方程(被映射为某个函数)
```
def forcedDampedNonlinearPendulum(omgd, fd = 1.2, q = 0.5 , gOverL = 1.):
def foo(t,X):
[omega, theta] = X
return array([- gOverL*math.sin(theta) - q*omega + fd*math.sin(omgd*t), omega])
return foo
```
考虑在初始条件有微小差别下的情况,这里角度有细微差别
[omega, theta] = [0., 0.20]
[omega, theta] = [0., 0.21]
分别用两个初值求解运动方程
```
ans1 = dSolve(forcedDampedNonlinearPendulum(2./3.), array([0., 0.20]), tMax, stepSize, method = "RK4")
ans1Omega = reduceStateList(ans1, [0])
ans1Theta = reduceStateList(ans1, [1])
ans2 = dSolve(forcedDampedNonlinearPendulum(2./3.), array([0., 0.21]), tMax, stepSize, method = "RK4")
ans2Omega = reduceStateList(ans2, [0])
ans2Theta = reduceStateList(ans2, [1])
```
分别画出角速度随时间变化,角度随时间变化,以及相空间轨迹
```
# Omega
ddp(ans2Omega,["ForwardEuler"], xLabel = "t/s", yLabel = "Omega/ 1/s")
ddp(ans1Omega,["RK4"], xLabel = "t/s", yLabel = "Omega/ 1/s")
done()
# Theta
ddp(ans2Theta,["ForwardEuler"], xLabel = "t/s", yLabel = "Theta/ 1")
ddp(ans1Theta,["RK4"], xLabel = "t/s", yLabel = "Theta/ 1")
done()
# phase diag
easyPlotXYHoldON(getXiList(ans2,2),getXiList(ans2,1), "ForwardEuler","Theta/ 1","Omega/ 1/s", "Phase Diagram")
easyPlotXYHoldON(getXiList(ans1,2),getXiList(ans1,1), "RK4","Theta/ 1","Omega/ 1/s", "Phase Diagram")
done()
```
角度随时间变化(未取模)
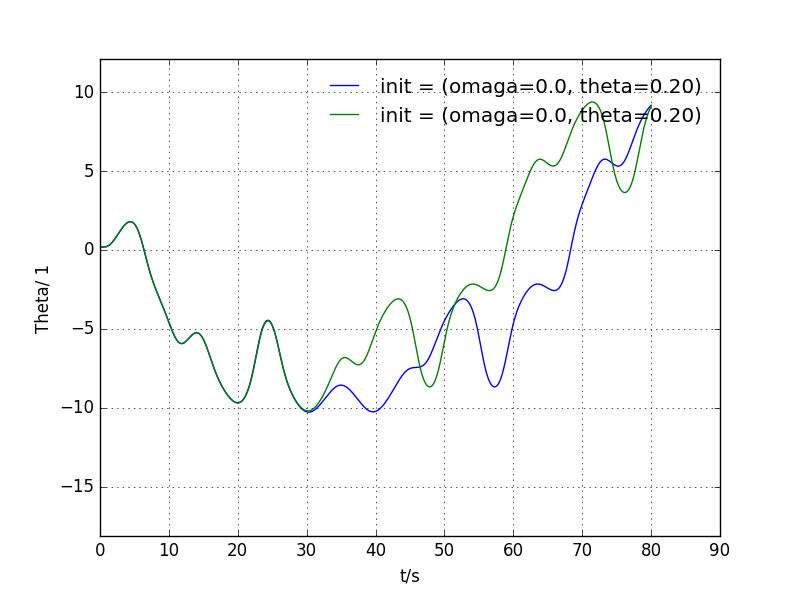
角度随时间变化(取模)
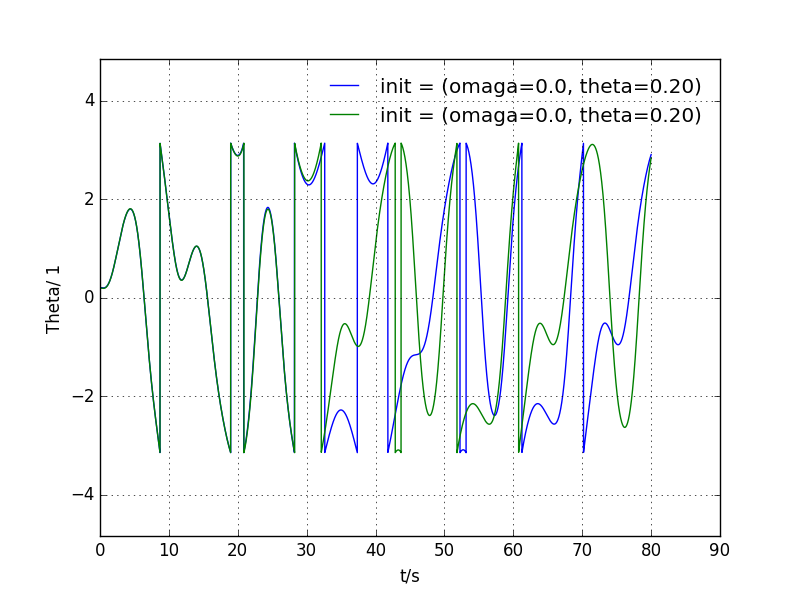
角速度随时间变化
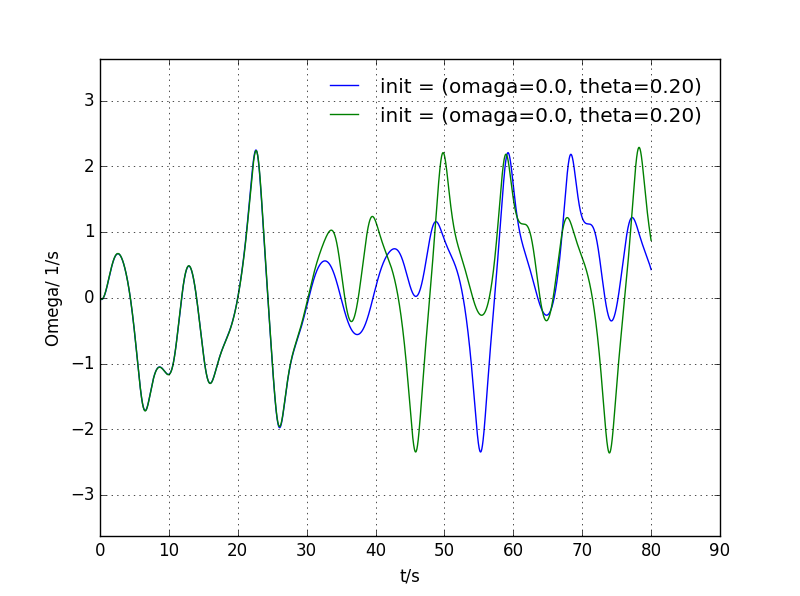
相图
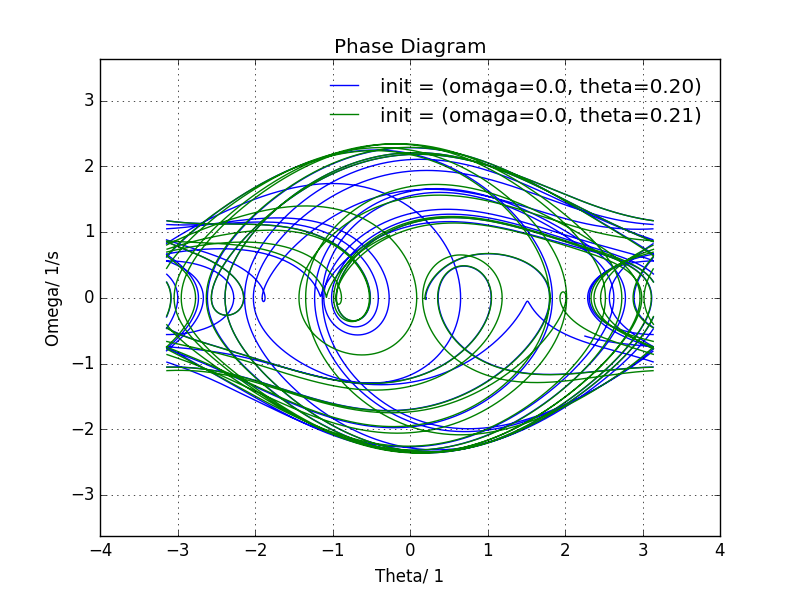
可以看出,初始结果的微小差别会随着时间被放大,并占据主导
## 庞加莱截面 ##
将庞加莱截面定义为对解的频闪提取
```
def poincareSection(solution, omgd, phase = 0., tmin = 50):
def mulOfTQ(t, omgd):
n = (t - phase/omgd)/ (2*math.pi/omgd)
return fEqQ(round(n), n)
n = len(solution)
sol = solution#[int(round(0.2*n)):-1]
sectionPoints = filter(lambda state: mulOfTQ(getT(state), omgd) and getT(state) > tmin, sol)
return sectionPoints
```
求解运动方程,并提取出不同初始相位下的庞加莱截面,并画出如下
以及,画出不同相位的动态图如下(gif,加载需要时间)
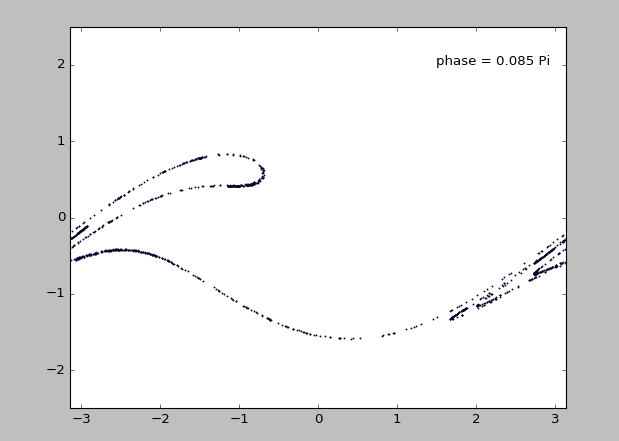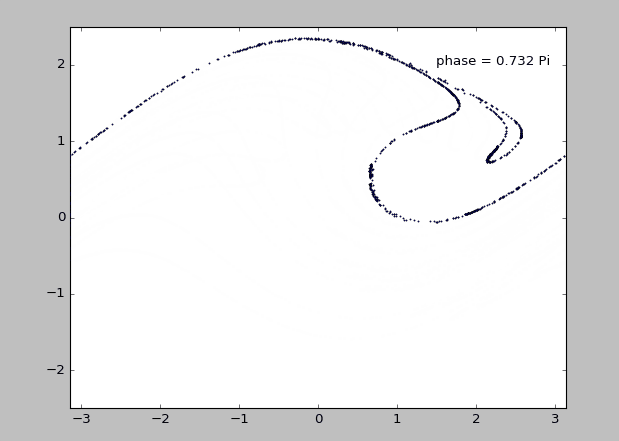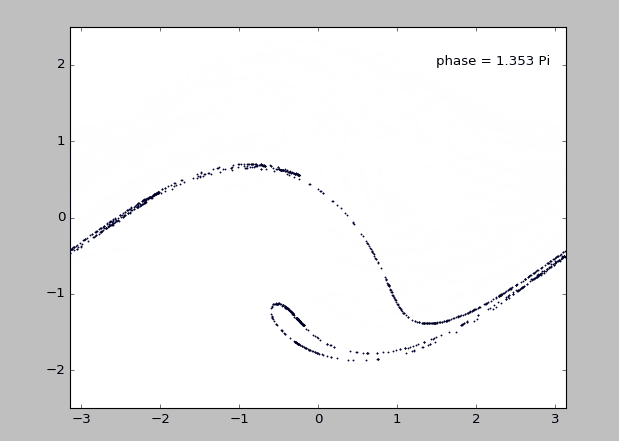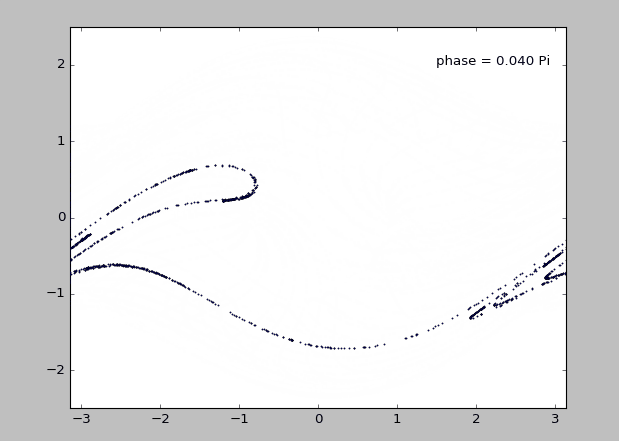
从庞加莱截面的有序性可以看出,即使混沌之中,也存在规律
| github_jupyter |
# Load and process molecules with `rdkit`
This notebook does the following:
- Molecules downloaded in the [previous notebook](./ 1_Get_Molecular_libraries.ipynb) are processed using `rdkit`.
The output of this notebook is a the file `rdKit_db_molecules.obj`, which is a pandas data frame containing the rdkit object of each molecule.
```
import pandas as pd
import numpy as np
from glob import glob
from pathlib import Path
from rdkit import Chem
from rdkit.Chem import AllChem
from rdkit import RDLogger
RDLogger.DisableLog('rdApp.*')
import sys
sys.path.append('../..')
from helper_modules.analyze_molecules_with_rdkit import *
```
## DEKOIS Molecules
```
sdf_input_path = './datasets/DEKOIS2/sdf/'
list_sdf_files = get_files_list(sdf_input_path, actives_name='ligand')
list_sdf_files
df_DEKOIS = get_mol_dataframe(load_molecules_from_dir(list_sdf_files))
df_DEKOIS.head()
df_DEKOIS.loc['ligand_4', 'mol_rdk']
```
## DUD 2006
```
sdf_input_path = './datasets/DUD/sdf/'
list_sdf_files = get_files_list(sdf_input_path, actives_name='ligand')
df_DUD = get_mol_dataframe(load_molecules_from_dir(list_sdf_files))
df_DUD.head()
df_DUD.loc['ligand_4', 'mol_rdk']
```
## CSAR 2012 molecules
```
sdf_input_path = './datasets/CSAR/sdf/'
# Function to sort the molecules alphanumerically
sort_function = lambda x: int(x.split('CS')[-1].split('.sdf')[0])
# Get the list of files
list_sdf_files = get_files_list(sdf_input_path,
actives_name = '',
inactives_name = '',
sort_func = sort_function
)
# Get the dataframe with the rdkit objects
df_CSAR = get_mol_dataframe(load_molecules_from_dir(list_sdf_files))
# The `Activity` column is not correct, we will update it
df_CSAR = df_CSAR.drop('Activity', axis = 1)
df_CSAR.index = df_CSAR.index.set_names(['Compound_ID'])
df_CSAR = df_CSAR.reset_index()
# Include information about activity of the molecule
csar_dat_file = (f'./datasets/CSAR/csar_dataset.csv') # Created in the previous notebook
csar_data = pd.read_csv(csar_dat_file)
# homogenize the df format
df_CSAR = df_CSAR.merge(csar_data[['Compound_ID', 'activity']], on = 'Compound_ID')
df_CSAR = df_CSAR.rename(columns = {'Compound_ID': 'Lig', 'activity': 'Activity'})
df_CSAR = df_CSAR.set_index('Lig')
df_CSAR = df_CSAR[['Activity', 'mol_rdk', 'sanitized']]
df_CSAR.head()
df_CSAR.loc['CS262', 'mol_rdk']
```
## Cocrystalized molecules
```
# Path to SDF Files
sdf_input_path = './datasets/COCRYS/sdf/'
# Function to sort the molecules by their PDB id
sort_function = lambda x: x.split('/')[-1].split('_')[0]
# Get the list of files
list_sdf_files = get_files_list(sdf_input_path,
actives_name='LIG',
sufix='',
sort_func = sort_function
)
# Compute a dataframe with the molecules as rdkit objects
df_pdi_lig = load_cocrys_molecules_from_dir(list_sdf_files)
# Update the dataframe
df_COCRYS = df_pdi_lig[['Lig', 'mol_rdk']]
df_COCRYS['Activity'] = 'active'
df_COCRYS['sanitized'] = [True if i != 'v3' else False for i in df_pdi_lig.validation]
df_COCRYS = df_COCRYS[['Lig', 'Activity', 'mol_rdk', 'sanitized']]
df_COCRYS = df_COCRYS.drop_duplicates('Lig').set_index('Lig')
df_COCRYS.sanitized.value_counts()
print('Shape', df_COCRYS.shape)
df_COCRYS.head()
df_COCRYS.loc['STU', 'mol_rdk']
```
## Merge all dataframes
```
input_file = './rdKit_db_molecules.obj'
df_mols = pd.read_pickle(input_file)
df_mols.keys()
list_dfs = [df_COCRYS, df_DEKOIS, df_DUD]
list_dfs_names = ['COCRYS', 'DEKOIS2', 'DUD']
# Create the final dataframe
df_all_libraries = pd.concat(list_dfs, keys = list_dfs_names)
df_all_libraries['Activity'] = df_all_libraries['Activity']\
.replace({'active': 1, 'inactive': 0})
df_all_libraries
# Save the dataframe
output_file = './rdKit_db_molecules.obj'
if not Path(output_file).exists():
df_all_libraries.to_pickle(output_file)
```
| github_jupyter |
# Spelling correcting using Levenshtein Distance
```
import numpy
import os
filepath = os.path.split(os.path.realpath('__file__'))[0]
#reading dictionary of words
with open(os.path.join(filepath,'data/dict.txt'), 'r', encoding='utf-8') as f:
word_list = [words.strip() for words in f.readlines()]
#Function to calculate Levenshtein Distance
dist_dict = {}
def LevDistance(orig_string, corr_string):
'''
Function to calculate Levenshtein Distance
orig_string : Original string which needs to be corrected
corr_string : Potential corrected version of the string
'''
if min(len(orig_string), len(corr_string)) == 0:
#If one either of the string is empty, distance is the length
#of the non-empty string
return max(len(orig_string), len(corr_string))
elif (orig_string==corr_string):
#If both the strings are exactly same, distance is zero
return 0
if orig_string[-1] == corr_string[-1]:
cost = 0
else:
cost = 1
# print(orig_string," + ",corr_string, " : cost - ",str(cost))
l1 = (orig_string[:-1], corr_string)
if not l1 in dist_dict:
dist_dict[l1] = LevDistance(*l1)
l2 = (orig_string, corr_string[:-1])
if not l2 in dist_dict:
dist_dict[l2] = LevDistance(*l2)
l3 = (orig_string[:-1], corr_string[:-1])
if not l3 in dist_dict:
dist_dict[l3] = LevDistance(*l3)
res = min([dist_dict[l1]+1, dist_dict[l2]+1, dist_dict[l3]+cost])
return res
LevDistance("Akshau","akshay")
import os
import random
from edit_distance import EditDistance
import re
import time
class SpellChecker:
def __init__(self,text_path):
dir_name = os.path.dirname(os.path.realpath('__file__'))
self.dictionary_file = dir_name+"/data/dict.txt"
self.corpus_file = dir_name+text_path
pass
def read_dictionary(self):
self.dictionary_words = set(line.strip() for line in open(self.dictionary_file))
def corpus_word_freq(self, print_results=False, ignore_higfreq_words = True, eval_lines =50):
'''
Function to generate dictionary for correct and corrupted words in the corpus.
Generating dictionary for correct words in the corpus by comparing if they
exists in the available dictionary 'corpus_word_dict' of words and if they
don't, adding them the to the dictionary 'invalid_corpus_tokens' of
corrupted words.
'''
# initiating dictionary for words
self.corpus_word_dict = {}
self.invalid_corpus_tokens = {}
lines_eval = 0
with open(self.corpus_file,'r') as f:
for line in f:
word_list = re.findall(r'\b[a-zA-Z]+\b', line)
for word in word_list:
#checking if the word exists in the given dictionary
if word.lower() in self.dictionary_words:
self.corpus_word_dict[word.lower()] = self.corpus_word_dict.get(word.lower(), 0) + 1
# if it doesn't exists, assuming it to be corrupted word and adding to other dictionary
# However, if the words start with a Upper case alphabet, assuming it to be a proper noun and
# ignoring those words to be added to corrupted dict
elif(lines_eval < eval_lines and not (word[0].isupper() and word[1].islower())):
self.invalid_corpus_tokens[word.lower()] = self.invalid_corpus_tokens.get(word.lower(),0) + 1
lines_eval += 1
'''
Since some words won't be the part of correct words dict, and if a word
in corrupted dict appears over a certain number of times, it is
considered as not corrupted and moved to the correct word dict.
'''
if ignore_higfreq_words:
self.ignored_words = set()
for key,val in self.invalid_corpus_tokens.items():
if int(val) > 25:
self.ignored_words.add(key)
for word in self.ignored_words:
self.corpus_word_dict[word] = self.invalid_corpus_tokens[word]
del self.invalid_corpus_tokens[word]
if print_results:
print(str(len(self.corpus_word_dict))," correct words added to the corpus dictionary")
print(str(len(self.invalid_corpus_tokens))," corrupted words exists in the corpus")
def print_top_n_line(self, n = 50, ignore_higfreq_words= True, calc_word_dict = True):
'''
Function to print first n lines from the corpus with underlined
corrupted words
'''
line_count = 0
if calc_word_dict:
self.corpus_word_freq(ignore_higfreq_words = ignore_higfreq_words, eval_lines=n)
with open(self.corpus_file, 'r') as f:
for line in f:
word_list = re.findall(r'\b[a-zA-Z]+\b', line)
for word in word_list:
if word.lower() in self.invalid_corpus_tokens:
line = re.sub(word,"\033[4m"+word+"\033[0m",line)
print(line)
line_count += 1
if line_count >= n:
break
def closest_replacement(self, possible_opt_dict):
'''
Function to resolve conflict in case of multiple replacements
with same edit distance available.
Replacement with the max occurance in the corpus will be selected
'''
inv_dict = {}
for key, value in possible_opt_dict.items():
if key in self.corpus_word_dict.keys():
possible_opt_dict[key] = self.corpus_word_dict[key]
inv_dict[possible_opt_dict[key]] = key
if len(inv_dict) > 0 :
return inv_dict[max(inv_dict.keys())]
else:
return None
def spell_check_first_n_lines(self, n=50, ignore_higfreq_words=True, min_distance = 4,
show_status = False, show_options = True, print_results = False):
'''
Function to spell check first n lines and replace the corrupted
words by the closest replacement in the dictionary.
Closest replacement of the word is defined using Levenshtein Distance
between the corrupted word and the words in dictonary. If multiple
replacement words with shortest distance are found, the replacement option
with the highest freqeuncy in the corpus is used.
Output:- Output of the function will be saved in "Output.txt" file in the current directory
n : Number of lines to be evaluated
ignore_higfreq_words : default:- True; Flag to ignore high frequency word from corrupted word dict
min_distance : default :- 4; Min edit distance to be used.
show_status : defualt :- False; Flag to enable printing of status with every correction
show_options : default :- True; Flag to enable printing of options for each corrupted words
print_results: default :- False; Flag to enable priting of the corrected text after spell check
'''
self.corpus_word_freq(ignore_higfreq_words = ignore_higfreq_words, eval_lines = n)
self.replacement_dict = {}
for index, invalid_word in enumerate(self.invalid_corpus_tokens):
if show_status:
print("Processing word "+str(index)+" of "+str(len(self.invalid_corpus_tokens))+" word "+invalid_word)
distance_dict = {}
min_dist = min_distance
for valid_word in self.dictionary_words:
# Only calculating edit distance with the correct words whose lenght is +/- 1 length of
# corrupted words
if (len(valid_word) >= len(invalid_word)-1) and (len(valid_word) <= len(invalid_word)+1):
#edit_dist = EditDistance(invalid_word, valid_word).calculate()
edit_dist = LevDistance(valid_word,invalid_word)
if edit_dist < min_dist:
min_dist = edit_dist
distance_dict = {}
if edit_dist <= min_dist:
distance_dict[valid_word] = edit_dist
#print(distance_dict)
# In case only one replacement found for the corrupted word with the minimum possible distance
# then, it will be selected as the replacement option.
if len(distance_dict) == 1:
self.replacement_dict[invalid_word] = list(distance_dict.keys())[0]
# In case multiple replacement options are available with the minimum edit distance, the word
# having highest frequency in corpus will be selected as the replacement option.
elif len(distance_dict) > 1 and self.closest_replacement(distance_dict) is not None:
self.replacement_dict[invalid_word] = self.closest_replacement(distance_dict)
else:
# In case no replacement word is found within the minimum distance limit, the corrupted word
# will be assigned as the replacement option.
self.replacement_dict[invalid_word] = invalid_word
if show_options:
print("Found the following replacements : ")
print(self.replacement_dict)
self.correct_text(replace_lines = n, print_res=print_results)
def correct_text(self, replace_lines = 50, print_res=False):
line_count = 0
os.remove("Output.txt")
print("\n")
with open(self.corpus_file, 'r') as f:
for line in f:
word_list = re.findall(r'\b[a-zA-Z]+\b', line)
for word in word_list:
if word.lower() in self.invalid_corpus_tokens:
line = re.sub(word,self.replacement_dict[word.lower()],line)
if print_res:
print(line)
with open('Output.txt', 'a') as w:
print(line, file=w)
line_count += 1
if line_count >= replace_lines:
print("\n---Corrected text written to Output.txt---")
break
```
## Applying Spell Check on Jane Austin
```
sc = SpellChecker("/data/Corrupted_Jane_Austin.txt")
sc.read_dictionary()
sc.print_top_n_line(50)
sc.corpus_word_freq(eval_lines=500)
starttime = time.time()
sc.spell_check_first_n_lines(500, show_status=True,show_options=False, print_results= True)
print("\n\nExecution time - %s seconds"% round(time.time()-starttime,3))
sc.replacement_dict
```
| github_jupyter |
```
# run this code to login to https://okpy.org/ and setup the assignment for submission
from ist256 import okclient
ok = okclient.Lab()
```
# In-Class Coding Lab: Web Services and APIs
### Overview
The web has long evolved from user-consumption to device consumption. In the early days of the web when you wanted to check the weather, you opened up your browser and visited a website. Nowadays your smart watch / smart phone retrieves the weather for you and displays it on the device. Your device can't predict the weather. It's simply consuming a weather based service.
The key to making device consumption work are API's (Application Program Interfaces). Products we use everyday like smartphones, Amazon's Alexa, and gaming consoles all rely on API's. They seem "smart" and "powerful" but in actuality they're only interfacing with smart and powerful services in the cloud.
API consumption is the new reality of programming; it is why we cover it in this course. Once you undersand how to conusme API's you can write a program to do almost anything and harness the power of the internet to make your own programs look "smart" and "powerful."
This lab covers how to properly use consume web service API's with Python. Here's what we will cover.
1. Understading requests and responses
1. Proper error handling
1. Parameter handling
1. Refactoring as a function
```
# Run this to make sure you have the pre-requisites!
!pip install -q requests
```
## Part 1: Understanding Requests and responses
In this part we learn about the Python requests module. http://docs.python-requests.org/en/master/user/quickstart/
This module makes it easy to write code to send HTTP requests over the internet and handle the responses. It will be the cornerstone of our API consumption in this course. While there are other modules which accomplish the same thing, `requests` is the most straightforward and easiest to use.
We'll begin by importing the modules we will need. We do this here so we won't need to include these lines in the other code we write in this lab.
```
# start by importing the modules we will need
import requests
import json
```
### The request
As you learned in class and your assigned readings, the HTTP protocol has **verbs** which consititue the type of request you will send to the remote resource, or **url**. Based on the url and request type, you will get a **response**.
The following line of code makes a **get** request (that's the HTTP verb) to Google's Geocoding API service. This service attempts to convert the address (in this case `Syracuse University`) into a set of coordinates global coordinates (Latitude and Longitude), so that location can be plotted on a map.
```
url = 'https://nominatim.openstreetmap.org/search?q=Hinds+Hall+Syracuse+University&format=json'
response = requests.get(url)
```
### The response
The `get()` method returns a `Response` object variable. I called it `response` in this example but it could be called anything.
The HTTP response consists of a *status code* and *body*. The status code lets you know if the request worked, while the body of the response contains the actual data.
```
response.ok # did the request work?
response.text # what's in the body of the response, as a raw string
```
### Converting responses into Python object variables
In the case of **web site url's** the response body is **HTML**. This should be rendered in a web browser. But we're dealing with Web Service API's so...
In the case of **web API url's** the response body could be in a variety of formats from **plain text**, to **XML** or **JSON**. In this course we will only focus on JSON format because as we've seen these translate easily into Python object variables.
Let's convert the response to a Python object variable. I this case it will be a Python dictionary
```
geodata = response.json() # try to decode the response from JSON format
geodata # this is now a Python object variable
```
With our Python object, we can now walk the list of dictionary to retrieve the latitude and longitude
```
lat = geodata[0]['lat']
lon =geodata[0]['lon']
print(lat, lon)
```
In the code above we "walked" the Python list of dictionary to get to the location
- `geodata` is a list
- `geodata[0]` is the first item in that list, a dictionary
- `geodata[0]['lat']` is a dictionary key which represents the latitude
- `geodata[0]['lon']` is a dictionary key which represents the longitude
It should be noted that this process will vary for each API you call, so its important to get accustomed to performing this task. You'll be doing it quite often.
One final thing to address. What is the type of `lat` and `lon`?
```
type(lat), type(lon)
```
Bummer they are strings. we want them to be floats so we will need to parse the strings with the `float()` function:
```
lat = float(geodata[0]['lat'])
lon = float(geodata[0]['lon'])
print("Latitude: %f, Longitude: %f" % (lat, lon))
```
### Now You Try It!
Walk the `geodata` object variable and reteieve the value under the key `display_name` and the key `bounding_box`
```
# todo:
# retrieve the place_id put in a variable
# retrieve the formatted_address put it in a variable
# print both of them out
```
## Part 2: Parameter Handling
In the example above we hard-coded "Hinds Hall Syracuse University" into the request:
```
url = 'https://nominatim.openstreetmap.org/search?q=Hinds+Hall+Syracuse+University&format=json'
```
A better way to write this code is to allow for the input of any location and supply that to the service. To make this work we need to send parameters into the request as a dictionary. This way we can geolocate any address!
You'll notice that on the url, we are passing **key-value pairs** the key is `q` and the value is `Hinds+Hall+Syracuse+University`. The other key is `format` and the value is `json`. Hey, Python dictionaries are also key-value pairs so:
```
url = 'https://nominatim.openstreetmap.org/search' # base URL without paramters after the "?"
search = 'Hinds Hall Syracuse University'
options = { 'q' : search, 'format' : 'json'}
response = requests.get(url, params = options)
geodata = response.json()
coords = { 'lat' : float(geodata[0]['lat']), 'lng' : float(geodata[0]['lon']) }
print("Search for:", search)
print("Coordinates:", coords)
print("%s is located at (%f,%f)" %(search, coords['lat'], coords['lng']))
```
### Looking up any address
RECALL: For `requests.get(url, params = options)` the part that says `params = options` is called a **named argument**, which is Python's way of specifying an optional function argument.
With our parameter now outside the url, we can easily re-write this code to work for any location! Go ahead and execute the code and input `Queens, NY`. This will retrieve the coordinates `(40.728224,-73.794852)`
```
url = 'https://nominatim.openstreetmap.org/search' # base URL without paramters after the "?"
search = input("Enter a loacation to Geocode: ")
options = { 'q' : search, 'format' : 'json'}
response = requests.get(url, params = options)
geodata = response.json()
coords = { 'lat' : float(geodata[0]['lat']), 'lng' : float(geodata[0]['lon']) }
print("Search for:", search)
print("Coordinates:", coords)
print("%s is located at (%f,%f)" %(search, coords['lat'], coords['lng']))
```
### So useful, it should be a function!
One thing you'll come to realize quickly is that your API calls should be wrapped in functions. This promotes **readability** and **code re-use**. For example:
```
def get_coordinates(search):
url = 'https://nominatim.openstreetmap.org/search' # base URL without paramters after the "?"
options = { 'q' : search, 'format' : 'json'}
response = requests.get(url, params = options)
geodata = response.json()
coords = { 'lat' : float(geodata[0]['lat']), 'lng' : float(geodata[0]['lon']) }
return coords
# main program here:
location = input("Enter a location: ")
coords = get_coordinates(location)
print("%s is located at (%f,%f)" %(location, coords['lat'], coords['lng']))
```
### Other request methods
Not every API we call uses the `get()` method. Some use `post()` because the amount of data you provide it too large to place on the url.
An example of this is the **Text-Processing.com** sentiment analysis service. http://text-processing.com/docs/sentiment.html This service will detect the sentiment or mood of text. You give the service some text, and it tells you whether that text is positive, negative or neutral.
```
# 'you suck' == 'negative'
url = 'http://text-processing.com/api/sentiment/'
options = { 'text' : 'you suck'}
response = requests.post(url, data = options)
sentiment = response.json()
sentiment
# 'I love cheese' == 'positive'
url = 'http://text-processing.com/api/sentiment/'
options = { 'text' : 'I love cheese'}
response = requests.post(url, data = options)
sentiment = response.json()
sentiment
```
In the examples provided we used the `post()` method instead of the `get()` method. the `post()` method has a named argument `data` which takes a dictionary of data. The key required by **text-processing.com** is `text` which hold the text you would like to process for sentiment.
We use a post in the event the text we wish to process is very long. Case in point:
```
tweet = "Arnold Schwarzenegger isn't voluntarily leaving the Apprentice, he was fired by his bad (pathetic) ratings, not by me. Sad end to a great show"
url = 'http://text-processing.com/api/sentiment/'
options = { 'text' : tweet }
response = requests.post(url, data = options)
sentiment = response.json()
sentiment
```
### Now You Try It!
Use the above example to write a program which will input any text and print the sentiment using this API!
```
# todo write code here
```
## Part 3: Proper Error Handling (In 3 Simple Rules)
When you write code that depends on other people's code from around the Internet, there's a lot that can go wrong. Therefore we perscribe the following advice:
```
Assume anything that CAN go wrong WILL go wrong
```
### Rule 1: Don't assume the internet 'always works'
The first rule of programming over a network is to NEVER assume the network is available. You need to assume the worst. No WiFi, user types in a bad url, the remote website is down, etc.
We handle this in the `requests` module by catching the `requests.exceptions.RequestException` Here's an example:
```
url = "http://this is not a website"
try:
response = requests.get(url) # throws an exception when it cannot connect
# internet is broken
except requests.exceptions.RequestException as e:
print("ERROR: Cannot connect to ", url)
print("DETAILS:", e)
```
### Rule 2: Don't assume the response you get back is valid
Assuming the internet is not broken (Rule 1) You should now check for HTTP response 200 which means the url responded successfully. Other responses like 404 or 501 indicate an error occured and that means you should not keep processing the response.
Here's one way to do it:
```
url = 'http://www.syr.edu/mikeisawesum' # this should 404
try:
response = requests.get(url)
if response.ok: # same as response.status_code == 200
data = response.text
else: # Some other non 200 response code
print("There was an Error requesting:", url, " HTTP Response Code: ", response.status_code)
# internet is broken
except requests.exceptions.RequestException as e:
print("ERROR: Cannot connect to ", url)
print("DETAILS:", e)
```
### Rule 2a: Use exceptions instead of if else in this case
Personally I don't like to use `if ... else` to handle an error. Instead, I prefer to instruct `requests` to throw an exception of `requests.exceptions.HTTPError` whenever the response is not ok. This makes the code you write a little cleaner.
Errors are rare occurences, and so I don't like error handling cluttering up my code.
```
url = 'http://www.syr.edu/mikeisawesum' # this should 404
try:
response = requests.get(url) # throws an exception when it cannot connect
response.raise_for_status() # throws an exception when not 'ok'
data = response.text
# response not ok
except requests.exceptions.HTTPError as e:
print("ERROR: Response from ", url, 'was not ok.')
print("DETAILS:", e)
# internet is broken
except requests.exceptions.RequestException as e:
print("ERROR: Cannot connect to ", url)
print("DETAILS:", e)
```
### Rule 3: Don't assume the data you get back is the data you expect.
And finally, do not assume the data arriving the the `response` is the data you expected. Specifically when you try and decode the `JSON` don't assume that will go smoothly. Catch the `json.decoder.JSONDecodeError`.
```
url = 'http://www.syr.edu' # this is HTML, not JSON
try:
response = requests.get(url) # throws an exception when it cannot connect
response.raise_for_status() # throws an exception when not 'ok'
data = response.json() # throws an exception when cannot decode json
# cannot decode json
except json.decoder.JSONDecodeError as e:
print("ERROR: Cannot decode the response into json")
print("DETAILS", e)
# response not ok
except requests.exceptions.HTTPError as e:
print("ERROR: Response from ", url, 'was not ok.')
print("DETAILS:", e)
# internet is broken
except requests.exceptions.RequestException as e:
print("ERROR: Cannot connect to ", url)
print("DETAILS:", e)
```
### Now You try it!
Using the last example above, write a program to input a location, call the `get_coordinates()` function, then print the coordindates. Make sure to handle all three types of exceptions!!!
```
# todo write code here to input a location, look up coordinates, and print
# it should handle errors!!!
```
## Metacognition
Please answer the following questions. This should be a personal narrative, in your own voice. Answer the questions by double clicking on the question and placing your answer next to the Answer: prompt.
1. Record any questions you have about this lab that you would like to ask in recitation. It is expected you will have questions if you did not complete the code sections correctly. Learning how to articulate what you do not understand is an important skill of critical thinking.
Answer:
2. What was the most difficult aspect of completing this lab? Least difficult?
Answer:
3. What aspects of this lab do you find most valuable? Least valuable?
Answer:
4. Rate your comfort level with this week's material so far.
1 ==> I can do this on my own and explain how to do it.
2 ==> I can do this on my own without any help.
3 ==> I can do this with help or guidance from others. If you choose this level please list those who helped you.
4 ==> I don't understand this at all yet and need extra help. If you choose this please try to articulate that which you do not understand.
Answer:
```
# to save and turn in your work, execute this cell. Your latest submission will be graded.
ok.submit()
```
| github_jupyter |
##### Copyright 2020 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Data augmentation
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/tutorials/images/data_augmentation"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/tutorials/images/data_augmentation.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/tutorials/images/data_augmentation.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/tutorials/images/data_augmentation.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
</td>
</table>
## Overview
This tutorial demonstrates data augmentation: a technique to increase the diversity of your training set by applying random (but realistic) transformations such as image rotation. You will learn how to apply data augmentation in two ways. First, you will use [Keras Preprocessing Layers](https://www.tensorflow.org/api_docs/python/tf/keras/layers/experimental/preprocessing/). Next, you will use `tf.image`.
## Setup
```
import matplotlib.pyplot as plt
import numpy as np
import tensorflow as tf
import tensorflow_datasets as tfds
from tensorflow.keras import layers
```
## Download a dataset
This tutorial uses the [tf_flowers](https://www.tensorflow.org/datasets/catalog/tf_flowers) dataset. For convenience, download the dataset using [TensorFlow Datasets](https://www.tensorflow.org/datasets). If you would like to learn about others ways of importing data, see the [load images](https://www.tensorflow.org/tutorials/load_data/images) tutorial.
```
(train_ds, val_ds, test_ds), metadata = tfds.load(
'tf_flowers',
split=['train[:80%]', 'train[80%:90%]', 'train[90%:]'],
with_info=True,
as_supervised=True,
)
```
The flowers dataset has five classes.
```
num_classes = metadata.features['label'].num_classes
print(num_classes)
```
Let's retrieve an image from the dataset and use it to demonstrate data augmentation.
```
get_label_name = metadata.features['label'].int2str
image, label = next(iter(train_ds))
_ = plt.imshow(image)
_ = plt.title(get_label_name(label))
```
## Use Keras preprocessing layers
Note: The [Keras Preprocesing Layers](https://www.tensorflow.org/api_docs/python/tf/keras/layers/experimental/preprocessing) introduced in this section are currently experimental.
### Resizing and rescaling
You can use preprocessing layers to [resize](https://www.tensorflow.org/api_docs/python/tf/keras/layers/experimental/preprocessing/Resizing) your images to a consistent shape, and to [rescale](https://www.tensorflow.org/api_docs/python/tf/keras/layers/experimental/preprocessing/Rescaling) pixel values.
```
IMG_SIZE = 180
resize_and_rescale = tf.keras.Sequential([
layers.experimental.preprocessing.Resizing(IMG_SIZE, IMG_SIZE),
layers.experimental.preprocessing.Rescaling(1./255)
])
```
Note: the rescaling layer above standardizes pixel values to `[0,1]`. If instead you wanted `[-1,1]`, you would write `Rescaling(1./127.5, offset=-1)`.
You can see the result of applying these layers to an image.
```
result = resize_and_rescale(image)
_ = plt.imshow(result)
```
You can verify the pixels are in `[0-1]`.
```
print("Min and max pixel values:", result.numpy().min(), result.numpy().max())
```
### Data augmentation
You can use preprocessing layers for data augmentation as well.
Let's create a few preprocessing layers and apply them repeatedly to the same image.
```
data_augmentation = tf.keras.Sequential([
layers.experimental.preprocessing.RandomFlip("horizontal_and_vertical"),
layers.experimental.preprocessing.RandomRotation(0.2),
])
# Add the image to a batch
image = tf.expand_dims(image, 0)
plt.figure(figsize=(10, 10))
for i in range(9):
augmented_image = data_augmentation(image)
ax = plt.subplot(3, 3, i + 1)
plt.imshow(augmented_image[0])
plt.axis("off")
```
There are a variety of preprocessing [layers](https://www.tensorflow.org/api_docs/python/tf/keras/layers/experimental/preprocessing) you can use for data augmentation including `layers.RandomContrast`, `layers.RandomCrop`, `layers.RandomZoom`, and others.
### Two options to use the preprocessing layers
There are two ways you can use these preprocessing layers, with important tradeoffs.
#### Option 1: Make the preprocessing layers part of your model
```
model = tf.keras.Sequential([
resize_and_rescale,
data_augmentation,
layers.Conv2D(16, 3, padding='same', activation='relu'),
layers.MaxPooling2D(),
# Rest of your model
])
```
There are two important points to be aware of in this case:
* Data augmentation will run on-device, synchronously with the rest of your layers, and benefit from GPU acceleration.
* When you export your model using `model.save`, the preprocessing layers will be saved along with the rest of your model. If you later deploy this model, it will automatically standardize images (according to the configuration of your layers). This can save you from the effort of having to reimplement that logic server-side.
Note: Data augmentation is inactive at test time so input images will only be augmented during calls to `model.fit` (not `model.evaluate` or `model.predict`).
#### Option 2: Apply the preprocessing layers to your dataset
```
aug_ds = train_ds.map(
lambda x, y: (resize_and_rescale(x, training=True), y))
```
With this approach, you use `Dataset.map` to create a dataset that yields batches of augmented images. In this case:
* Data augmentation will happen asynchronously on the CPU, and is non-blocking. You can overlap the training of your model on the GPU with data preprocessing, using `Dataset.prefetch`, shown below.
* In this case the prepreprocessing layers will not be exported with the model when you call `model.save`. You will need to attach them to your model before saving it or reimplement them server-side. After training, you can attach the preprocessing layers before export.
You can find an example of the first option in the [image classification](https://www.tensorflow.org/tutorials/images/classification) tutorial. Let's demonstrate the second option here.
### Apply the preprocessing layers to the datasets
Configure the train, validation, and test datasets with the preprocessing layers you created above. You will also configure the datasets for performance, using parallel reads and buffered prefetching to yield batches from disk without I/O become blocking. You can learn more dataset performance in the [Better performance with the tf.data API](https://www.tensorflow.org/guide/data_performance) guide.
Note: data augmentation should only be applied to the training set.
```
batch_size = 32
AUTOTUNE = tf.data.AUTOTUNE
def prepare(ds, shuffle=False, augment=False):
# Resize and rescale all datasets
ds = ds.map(lambda x, y: (resize_and_rescale(x), y),
num_parallel_calls=AUTOTUNE)
if shuffle:
ds = ds.shuffle(1000)
# Batch all datasets
ds = ds.batch(batch_size)
# Use data augmentation only on the training set
if augment:
ds = ds.map(lambda x, y: (data_augmentation(x, training=True), y),
num_parallel_calls=AUTOTUNE)
# Use buffered prefecting on all datasets
return ds.prefetch(buffer_size=AUTOTUNE)
train_ds = prepare(train_ds, shuffle=True, augment=True)
val_ds = prepare(val_ds)
test_ds = prepare(test_ds)
```
### Train a model
For completeness, you will now train a model using these datasets. This model has not been tuned for accuracy (the goal is to show you the mechanics).
```
model = tf.keras.Sequential([
layers.Conv2D(16, 3, padding='same', activation='relu'),
layers.MaxPooling2D(),
layers.Conv2D(32, 3, padding='same', activation='relu'),
layers.MaxPooling2D(),
layers.Conv2D(64, 3, padding='same', activation='relu'),
layers.MaxPooling2D(),
layers.Flatten(),
layers.Dense(128, activation='relu'),
layers.Dense(num_classes)
])
model.compile(optimizer='adam',
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=['accuracy'])
epochs=5
history = model.fit(
train_ds,
validation_data=val_ds,
epochs=epochs
)
loss, acc = model.evaluate(test_ds)
print("Accuracy", acc)
```
### Custom data augmentation
You can also create custom data augmenation layers. This tutorial shows two ways of doing so. First, you will create a `layers.Lambda` layer. This is a good way to write concise code. Next, you will write a new layer via [subclassing](https://www.tensorflow.org/guide/keras/custom_layers_and_models), which gives you more control. Both layers will randomly invert the colors in an image, according to some probability.
```
def random_invert_img(x, p=0.5):
if tf.random.uniform([]) < p:
x = (255-x)
else:
x
return x
def random_invert(factor=0.5):
return layers.Lambda(lambda x: random_invert_img(x, factor))
random_invert = random_invert()
plt.figure(figsize=(10, 10))
for i in range(9):
augmented_image = random_invert(image)
ax = plt.subplot(3, 3, i + 1)
plt.imshow(augmented_image[0].numpy().astype("uint8"))
plt.axis("off")
```
Next, implement a custom layer by [subclassing](https://www.tensorflow.org/guide/keras/custom_layers_and_models).
```
class RandomInvert(layers.Layer):
def __init__(self, factor=0.5, **kwargs):
super().__init__(**kwargs)
self.factor = factor
def call(self, x):
return random_invert_img(x)
_ = plt.imshow(RandomInvert()(image)[0])
```
Both of these layers can be used as described in options 1 and 2 above.
## Using tf.image
The above `layers.preprocessing` utilities are convenient. For finer control, you can write your own data augmentation pipelines or layers using `tf.data` and `tf.image`. You may also want to check out [TensorFlow Addons Image: Operations](https://www.tensorflow.org/addons/tutorials/image_ops) and [TensorFlow I/O: Color Space Conversions](https://www.tensorflow.org/io/tutorials/colorspace)
Since the flowers dataset was previously configured with data augmentation, let's reimport it to start fresh.
```
(train_ds, val_ds, test_ds), metadata = tfds.load(
'tf_flowers',
split=['train[:80%]', 'train[80%:90%]', 'train[90%:]'],
with_info=True,
as_supervised=True,
)
```
Retrieve an image to work with.
```
image, label = next(iter(train_ds))
_ = plt.imshow(image)
_ = plt.title(get_label_name(label))
```
Let's use the following function to visualize and compare the original and augmented images side-by-side.
```
def visualize(original, augmented):
fig = plt.figure()
plt.subplot(1,2,1)
plt.title('Original image')
plt.imshow(original)
plt.subplot(1,2,2)
plt.title('Augmented image')
plt.imshow(augmented)
```
### Data augmentation
#### Flipping the image
Flip the image either vertically or horizontally.
```
flipped = tf.image.flip_left_right(image)
visualize(image, flipped)
```
#### Grayscale the image
Grayscale an image.
```
grayscaled = tf.image.rgb_to_grayscale(image)
visualize(image, tf.squeeze(grayscaled))
_ = plt.colorbar()
```
#### Saturate the image
Saturate an image by providing a saturation factor.
```
saturated = tf.image.adjust_saturation(image, 3)
visualize(image, saturated)
```
#### Change image brightness
Change the brightness of image by providing a brightness factor.
```
bright = tf.image.adjust_brightness(image, 0.4)
visualize(image, bright)
```
#### Center crop the image
Crop the image from center up to the image part you desire.
```
cropped = tf.image.central_crop(image, central_fraction=0.5)
visualize(image,cropped)
```
#### Rotate the image
Rotate an image by 90 degrees.
```
rotated = tf.image.rot90(image)
visualize(image, rotated)
```
### Random transformations
Warning: There are two sets of random image operations: `tf.image.random*` and `tf.image.stateless_random*`. Using `tf.image.random*` operations is strongly discouraged as they use the old RNGs from TF 1.x. Instead, please use the random image operations introduced in this tutorial. For more information, please refer to [Random number generation](https://www.tensorflow.org/guide/random_numbers).
Applying random transformations to the images can further help generalize and expand the dataset. Current `tf.image` API provides 8 such random image operations (ops):
* [`tf.image.stateless_random_brightness`](https://www.tensorflow.org/api_docs/python/tf/image/stateless_random_brightness)
* [`tf.image.stateless_random_contrast`](https://www.tensorflow.org/api_docs/python/tf/image/stateless_random_contrast)
* [`tf.image.stateless_random_crop`](https://www.tensorflow.org/api_docs/python/tf/image/stateless_random_crop)
* [`tf.image.stateless_random_flip_left_right`](https://www.tensorflow.org/api_docs/python/tf/image/stateless_random_flip_left_right)
* [`tf.image.stateless_random_flip_up_down`](https://www.tensorflow.org/api_docs/python/tf/image/stateless_random_flip_up_down)
* [`tf.image.stateless_random_hue`](https://www.tensorflow.org/api_docs/python/tf/image/stateless_random_hue)
* [`tf.image.stateless_random_jpeg_quality`](https://www.tensorflow.org/api_docs/python/tf/image/stateless_random_jpeg_quality)
* [`tf.image.stateless_random_saturation`](https://www.tensorflow.org/api_docs/python/tf/image/stateless_random_saturation)
These random image ops are purely functional: the ouput only depends on the input. This makes them simple to use in high performance, deterministic input pipelines. They require a `seed` value be input each step. Given the same `seed`, they return the same results independent of how many times they are called.
Note: `seed` is a `Tensor` of shape `(2,)` whose values are any integers.
In the following sections, we will:
1. Go over examples of using random image operations to transform an image, and
2. Demonstrate how to apply random transformations to a training dataset.
#### Randomly change image brightness
Randomly change the brightness of `image` by providing a brightness factor and `seed`. The brightness factor is chosen randomly in the range `[-max_delta, max_delta)` and is associated with the given `seed`.
```
for i in range(3):
seed = (i, 0) # tuple of size (2,)
stateless_random_brightness = tf.image.stateless_random_brightness(
image, max_delta=0.95, seed=seed)
visualize(image, stateless_random_brightness)
```
#### Randomly change image contrast
Randomly change the contrast of `image` by providing a contrast range and `seed`. The contrast range is chosen randomly in the interval `[lower, upper]` and is associated with the given `seed`.
```
for i in range(3):
seed = (i, 0) # tuple of size (2,)
stateless_random_contrast = tf.image.stateless_random_contrast(
image, lower=0.1, upper=0.9, seed=seed)
visualize(image, stateless_random_contrast)
```
#### Randomly crop an image
Randomly crop `image` by providing target `size` and `seed`. The portion that gets cropped out of `image` is at a randomly chosen offet and is associated with the given `seed`.
```
for i in range(3):
seed = (i, 0) # tuple of size (2,)
stateless_random_crop = tf.image.stateless_random_crop(
image, size=[210, 300, 3], seed=seed)
visualize(image, stateless_random_crop)
```
### Apply augmentation to a dataset
Let's first download the image dataset again in case they are modified in the previous sections.
```
(train_datasets, val_ds, test_ds), metadata = tfds.load(
'tf_flowers',
split=['train[:80%]', 'train[80%:90%]', 'train[90%:]'],
with_info=True,
as_supervised=True,
)
```
Let's define a utility function for resizing and rescaling the images. This function will be used in unifying the size and scale of images in the dataset:
```
def resize_and_rescale(image, label):
image = tf.cast(image, tf.float32)
image = tf.image.resize(image, [IMG_SIZE, IMG_SIZE])
image = (image / 255.0)
return image, label
```
Let's also define `augment` function that can apply the random transformations to the images. This function will be used on the dataset in the next step.
```
def augment(image_label, seed):
image, label = image_label
image, label = resize_and_rescale(image, label)
image = tf.image.resize_with_crop_or_pad(image, IMG_SIZE + 6, IMG_SIZE + 6)
# Make a new seed
new_seed = tf.random.experimental.stateless_split(seed, num=1)[0, :]
# Random crop back to the original size
image = tf.image.stateless_random_crop(
image, size=[IMG_SIZE, IMG_SIZE, 3], seed=seed)
# Random brightness
image = tf.image.stateless_random_brightness(
image, max_delta=0.5, seed=new_seed)
image = tf.clip_by_value(image, 0, 1)
return image, label
```
#### Option 1: Using `tf.data.experimental.Counter()`
Create a `tf.data.experimental.Counter()` object (let's call it `counter`) and `zip` the dataset with `(counter, counter)`. This will ensure that each image in the dataset gets associated with a unique value (of shape `(2,)`) based on `counter` which later can get passed into the `augment` function as the `seed` value for random transformations.
```
# Create counter and zip together with train dataset
counter = tf.data.experimental.Counter()
train_ds = tf.data.Dataset.zip((train_datasets, (counter, counter)))
```
Map the `augment` function to the training dataset.
```
train_ds = (
train_ds
.shuffle(1000)
.map(augment, num_parallel_calls=AUTOTUNE)
.batch(batch_size)
.prefetch(AUTOTUNE)
)
val_ds = (
val_ds
.map(resize_and_rescale, num_parallel_calls=AUTOTUNE)
.batch(batch_size)
.prefetch(AUTOTUNE)
)
test_ds = (
test_ds
.map(resize_and_rescale, num_parallel_calls=AUTOTUNE)
.batch(batch_size)
.prefetch(AUTOTUNE)
)
```
#### Option 2: Using `tf.random.Generator`
Create a `tf.random.Generator` object with an intial `seed` value. Calling `make_seeds` function on the same generator object returns a new, unique `seed` value always. Define a wrapper function that 1) calls `make_seeds` function and that 2) passes the newly generated `seed` value into the `augment` function for random transformations.
Note: `tf.random.Generator` objects store RNG state in a `tf.Variable`, which means it can be saved as a [checkpoint](https://www.tensorflow.org/guide/checkpoint) or in a [SavedModel](https://www.tensorflow.org/guide/saved_model). For more details, please refer to [Random number generation](https://www.tensorflow.org/guide/random_numbers).
```
# Create a generator
rng = tf.random.Generator.from_seed(123, alg='philox')
# A wrapper function for updating seeds
def f(x, y):
seed = rng.make_seeds(2)[0]
image, label = augment((x, y), seed)
return image, label
```
Map the wrapper function `f` to the training dataset.
```
train_ds = (
train_datasets
.shuffle(1000)
.map(f, num_parallel_calls=AUTOTUNE)
.batch(batch_size)
.prefetch(AUTOTUNE)
)
val_ds = (
val_ds
.map(resize_and_rescale, num_parallel_calls=AUTOTUNE)
.batch(batch_size)
.prefetch(AUTOTUNE)
)
test_ds = (
test_ds
.map(resize_and_rescale, num_parallel_calls=AUTOTUNE)
.batch(batch_size)
.prefetch(AUTOTUNE)
)
```
These datasets can now be used to train a model as shown previously.
## Next steps
This tutorial demonstrated data augmentation using [Keras Preprocessing Layers](https://www.tensorflow.org/api_docs/python/tf/keras/layers/experimental/preprocessing/) and `tf.image`. To learn how to include preprocessing layers inside your model, see the [Image classification](https://www.tensorflow.org/tutorials/images/classification) tutorial. You may also be interested in learning how preprocessing layers can help you classify text, as shown in the [Basic text classification](https://www.tensorflow.org/tutorials/keras/text_classification) tutorial. You can learn more about `tf.data` in this [guide](https://www.tensorflow.org/guide/data), and you can learn how to configure your input pipelines for performance [here](https://www.tensorflow.org/guide/data_performance).
| github_jupyter |
```
import os
import csv
from predictor import Predictor
def read_csv(path):
with open(path, 'r') as f:
reader = csv.reader(f)
data_list = list(reader)
return data_list
file_path = '/root/data/pm25_data/datas/Taiwan_data/201703_Taiwan_sol.csv'
dev_path = '/root/data/pm25_data/datas/sensors/test/'
model_path = 'models/muti_lstm_20_plus.ckpt'
predictor = Predictor(model_path , network = 'muti_lstm')
all_data = read_csv(file_path)
#
datas = dict()
result = ['PM2.5'] + ['' for i in range(len(all_data[1:]))]
for i , row in enumerate(all_data[1:]):
data = [a.strip() for a in row]
dev = data[2]
if dev in datas:
datas[dev].append([i,data])
else:
datas.update({dev : [[i,data]]})
print ('Finish , total dev : {}'.format(len(datas.keys())))
# predict
import datetime
import config
date_format = '%Y-%m-%d %H:%M:%S'
for a , (key , values) in enumerate(datas.items()):
dev = os.path.join(dev_path , key + '.csv')
dev_data = read_csv(dev)
date_strings = [ i[0] for i in dev_data]
dates = [datetime.datetime.strptime(i,date_format) for i in date_strings]
print (dev , len(datas.items()) , a)
for source_idx , item in values:
date , time , dev , *content = item[:]
# round data time
date_time = date + ' ' + time
today = datetime.datetime.strptime(date_time,date_format)
future_dates = [date
for date in dates
if date >= today]
if future_dates:
# next_closest_date = min(future_dates)
next_closest_date = future_dates[0]
else:
next_closest_date = dates[-1]
next_closest_date = next_closest_date.strftime(date_format)
cur = [i[0] for i in dev_data].index(next_closest_date)
if cur - config.sample_feq < 0:
pm25 = dev_data[cur][1]
else:
row_seq = [i for i in range(cur - config.sample_feq -1 , cur -1 , config.sample_skip)]
data = [[float(j)/3000 for j in dev_data[i][1:8]] for i in row_seq]
pred = predictor([data])[0][0] * 3000
pm25 = "{:0.2f}".format(pred)
result[source_idx +1] = pm25
print (len(all_data) , len(result))
# dump result to csv
out_path = '/root/data/pm25_data/datas/Taiwan_data/'
model_name = model_path.split('/')[-1].replace('.ckpt','_result.csv')
# 1d to 2d
result = [[i] for i in result]
with open(out_path + '/' + model_name, "w", encoding = "utf-8") as f:
writer = csv.writer(f)
writer.writerows(result)
# filter all attribute only pm2.5 exist
import csv
out = []
path = '/root/data/pm25_data/datas/Taiwan_data/201703_Taiwan_sol.csv'
with open(path, newline='', encoding = "utf-8") as csvfile:
rows = csv.reader(csvfile)
for i,row in enumerate(rows):
data = [a.strip() for a in row]
out.append([data[3]])
out_path = '/root/data/pm25_data/datas/Taiwan_data/201703_Taiwan_sol_filtered.csv'
with open(out_path, "w", encoding = "utf-8") as f:
writer = csv.writer(f)
writer.writerows(out)
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import urllib2
import webbrowser
import os
import bs4, re
from bs4 import BeautifulSoup
import requests
import math
import nltk
from nltk.tokenize import word_tokenize
import matplotlib.pyplot as plt
%matplotlib inline
from datetime import datetime
```
### The cell below opens (and converts to a dataframe) a csv file containing a URL that, when opened, will automatically download a csv file tuned to the prescribed search. For example, the first search performed looked (1) myeloma, (2) USA, (3) Recruiting AND Non yet recruiting AND Active, but no longer recruiting, (4) Car-T. The URL syntax can be found at ClinicalTrials.gov site.
```
df = pd.read_csv('D:\Python_Database\Myeloma\MM Trials\keyword_search.csv')
df1 = pd.read_csv('D:\Python_Database\uscities.csv')
df.head()
```
### The cells below open the url stored in the dataframe, and then moves the csv file from download to the desired file location
```
chrome_path = 'C:/Program Files (x86)/Google/Chrome/Application/chrome.exe %s'
url = df['url'].tolist()
webbrowser.get(using=chrome_path).open(url[0])
os.rename("C:/Users/robin/Downloads/SearchResults.csv", "D:/Python_Database/Myeloma/MM Trials/CART_Results_" + str(datetime.now())[:10] +".csv")
df = pd.read_csv('D:\Python_Database\Myeloma\MM Trials\CART_Results_' + str(datetime.now())[:10] + '.csv')
df_ = df
df = df[['NCT Number','Title','Locations','Phases', 'Status', 'Interventions', 'Last Update Posted','Sponsor/Collaborators']]
color = ['#0000ff','#0e0dff','#1716ff','#1e1dff','#2323ff','#2828ff','#2c2dff','#2f32ff','#3336ff','#353aff',
'#383eff','#3b42ff','#3d46ff','#3f49ff','#414dff','#4351ff','#4554ff','#4757ff','#485bff','#4a5eff',
'#4b61ff','#4d65ff','#4e68ff','#4f6bff','#506eff','#5172ff','#5275ff','#5378ff','#547bff','#547eff',
'#5581ff','#5684ff','#5687ff','#568bff','#578eff','#5791ff','#5794ff','#5897ff','#589aff','#589dff',
'#58a0ff','#58a3ff','#57a6ff','#57a9ff','#57acff','#56afff','#56b2ff','#55b5ff','#55b8ff','#54bbff',
'#53bfff','#52c2ff','#51c5ff','#50c8ff','#4ecbff','#4dceff','#4bd1ff','#4ad4ff','#48d7ff','#46daff',
'#44ddff','#41e0ff','#3ee3ff','#3be6ff','#38e9ff','#34ecff','#30f0ff','#2bf3ff','#25f6ff','#1df9ff',
'#13fcff','#00ffff','#08fdfb','#0efbf8','#13f9f4','#17f7f1','#1af6ed','#1df4ea','#1ff2e6','#22f0e3',
'#23eedf','#25ecdc','#27ead8','#28e8d5','#29e6d1','#2ae5ce','#2be3ca','#2ce1c7','#2ddfc4','#2eddc0',
'#2fdbbd','#2fdab9','#30d8b6','#30d6b3','#30d4af','#31d2ac','#31d0a8','#31cfa5','#31cda2','#31cb9e',
'#31c99b','#31c798','#31c594','#31c491','#31c28e','#31c08a','#31be87','#30bc84','#30bb81','#30b97d',
'#2fb77a','#2fb577','#2eb473','#2eb270','#2db06d','#2dae6a','#2cac66','#2bab63','#2ba960','#2aa75d',
'#29a559','#28a456','#27a253','#26a04f','#259f4c','#249d49','#239b45','#229942','#20983f','#1f963b',
'#1e9438','#1c9334','#1a9131','#198f2d','#178d2a','#158c26','#138a22','#10881d','#0e8719','#0a8514',
'#07830e','#038207','#008000','#0c8200','#158400','#1c8500','#228700','#278900','#2c8b00','#308c00',
'#358e00','#399000','#3d9200','#419300','#449500','#489700','#4b9900','#4f9a00','#529c00','#569e00',
'#59a000','#5da100','#60a300','#63a500','#66a700','#6aa900','#6daa00','#70ac00','#73ae00','#76b000',
'#79b100','#7db300','#80b500','#83b700','#86b900','#89ba00','#8cbc00','#8fbe00','#92c000','#95c200',
'#99c300','#9cc500','#9fc700','#a2c900','#a5ca00','#a8cc00','#abce00','#aed000','#b1d200','#b4d300',
'#b7d500','#bad700','#bed900','#c1db00','#c4dc00','#c7de00','#cae000','#cde200','#d0e400','#d3e600',
'#d6e700','#d9e900','#ddeb00','#e0ed00','#e3ef00','#e6f000','#e9f200','#ecf400','#eff600','#f2f800',
'#f6fa00','#f9fb00','#fcfd00','#ffff00','#fffc00','#fffa00','#fff700','#fff500','#fff200','#fff000',
'#ffed00','#ffeb00','#ffe800','#ffe500','#ffe300','#ffe000','#ffde00','#ffdb00','#ffd800','#ffd600',
'#ffd300','#ffd100','#ffce00','#ffcb00','#ffc900','#ffc600','#ffc300','#ffc100','#ffbe00','#ffbb00',
'#ffb900','#ffb600','#ffb300','#ffb100','#ffae00','#ffab00','#ffa800','#ffa600','#ffa300','#ffa000',
'#ff9d00','#ff9a00','#ff9800','#ff9500','#ff9200','#ff8f00','#ff8c00','#ff8900','#ff8600','#ff8300',
'#ff8000','#ff7d00','#ff7a00','#ff7600','#ff7300','#ff7000','#ff6c00','#ff6900','#ff6500','#ff6200',
'#ff5e00','#ff5a00','#ff5700','#ff5200','#ff4e00','#ff4a00','#ff4500','#ff4000','#ff3b00','#ff3500',
'#ff2f00','#ff2800','#ff1f00','#ff1400','#ff0000']
color = color[::len(color)/len(df_)]
# len(color)
mask = df['Status'].str.contains('Recruiting', case=True)
colors = color[:len(df['NCT Number'])]
colors = pd.DataFrame(colors)
colors.columns = ['Color']
df = pd.concat([df, colors], axis=1)
pd.set_option('max_colwidth', 800)
temp = df.reindex(index=df.index[::-1])
temp[['NCT Number','Title', 'Status']]
test_data = df['Locations'].tolist()
city, state, city_state = [], [], []
for i in range(len(df1['city'])):
city.append(df1['city'][i])
state.append(df1['state_name'][i])
city_state.append(city[i] + ', ' + state[i])
results = []
for i in range(len(test_data)):
for j in range(len(city_state)):
if str(test_data[i]).find(city_state[j]) >=0:
results.append(city_state[j])
temp = []
temp1 = []
for i in range(len(test_data)):
for j in range(len(city_state)):
if str(test_data[i]).find(city_state[j]) >= 0:
temp1.append(city_state[j])
temp.append(temp1)
temp1=[]
location_of_study=[]
for i in range(len(temp)):
names = set(temp[i])
names = list(names)
location_of_study.append(names)
num=[]
for i in range(len(location_of_study)):
num.append(len(location_of_study[i]))
ind, pos = [], []
for i in range(len(num)):
if num[i] > 1:
ind.append(num[i])
pos.append(i)
df = df.reset_index(drop=True)
index = df.index.tolist()
temp = [x*1000 for x in index]
df = df.set_index([temp])
for i in range(len(pos)):
k=0
while k < ind[i]:
df.loc[(pos[i]*1000)+k] = df.loc[pos[i]*1000]
k=k+1
df = df.sort_index()
new_column = []
for i in range(len(location_of_study)):
for j in range(len(location_of_study[i])):
new_column.append(location_of_study[i][j])
City_State = pd.DataFrame(new_column, columns=['City, State'])
df_new = pd.concat([df.reset_index(drop=True), City_State], axis=1)
df_new.head()
cit = df1['city'].tolist()
state = df1['state_name'].tolist()
loc_db = []
for i in range(len(cit)):
loc_db.append(cit[i] + ', ' + state[i])
lat_e, lng_e = [],[]
citystate = df_new['City, State'].tolist()
for i in range(len(citystate)):
for j in range(len(loc_db)):
if citystate[i] == loc_db[j]:
lng_e.append(df1['lng'][j])
lat_e.append(df1['lat'][j])
lyo = df_new['Last Update Posted'].tolist()
lyo = [2000 + int(x[-2:]) for x in lyo]
LYO = pd.DataFrame(lyo, columns=['Year'])
Lat = pd.DataFrame(lat_e, columns=['Lat'])
Lng = pd.DataFrame(lng_e, columns=['Lng'])
Lat = Lat.reset_index(drop=True)
Lng = Lng.reset_index(drop=True)
LYO = LYO.reset_index(drop=True)
df_new = pd.concat([df_new, Lat, Lng, LYO], axis=1)
df_new.head()
df_new['Status'].unique().tolist()
df_new.head(1)
NCT_no = df_new['NCT Number'].unique().tolist()
NCT_no[1]
df_new_ = df_new[['NCT Number','Title', 'City, State', 'Lat', 'Lng', 'Color']]
mask2 = []
for i in range(len(NCT_no)):
mask2.append(df_new_.mask(df_new_['NCT Number'] != NCT_no[i]).dropna(axis=0, inplace=False).reset_index(drop=True))
mask2[1]
city_state = df_new['City, State'].unique().tolist()
mask3 = []
for i in range(len(city_state)):
mask3.append(df_new_.mask(df_new_['City, State'] != city_state[i]).dropna(axis=0, inplace=False).reset_index(drop=True))
len(mask3[0])
city_site = []
for i in range(len(mask3)):
city_site.append(mask3[i].iloc[0])
city_site[0]
for i in range(len(mask3)):
for j in range(len(mask3[i])):
if len(mask3[i]) > 1:
mask3[i]['Lat'][j] = mask3[i]['Lat'][j] + 1* math.cos(j*math.pi/((7+1)/2.))
mask3[i]['Lng'][j] = mask3[i]['Lng'][j] + 1* math.sin(j*math.pi/((7+1)/2.))
else:
mask3[i]['Lat'][j] = mask3[i]['Lat'][j]
mask3[i]['Lng'][j] = mask3[i]['Lng'][j]
mask3[0]
import cartopy.crs as ccrs
import cartopy.feature as cfeature
plt.figure(figsize=(18,16))
states_provinces = cfeature.NaturalEarthFeature(
category='cultural',
name='admin_1_states_provinces_lines',
scale='50m',
facecolor='none')
ax = plt.axes(projection=ccrs.PlateCarree())
ax.set_extent([-125, -66.5, 20, 50], ccrs.Geodetic())
ax.coastlines()
ax.add_feature(cfeature.BORDERS)
ax.add_feature(states_provinces, edgecolor='gray')
ax.background_patch.set_visible(False)
ax.outline_patch.set_visible(False)
for i in range(len(mask3)):
for j in range(len(mask3[i])):
plt.plot([city_site[i]['Lng'],mask3[i].iloc[j]['Lng']], [city_site[i]['Lat'], mask3[i].iloc[j]['Lat']], c='k', alpha=.3 )
plt.scatter(city_site[i]['Lng'], city_site[i]['Lat'], c='orange', alpha=0.2)
plt.scatter(mask3[i].iloc[j]['Lng'], mask3[i].iloc[j]['Lat'], c=mask3[i].iloc[j]['Color'], alpha=1)
plt.savefig('MM Trials_CART_' + str(datetime.now())[:10] + '.png', format='png', dpi=600, transparent=True)
plt.title('MM Trials_CART_' + str(datetime.now())[:10] + '\n')
plt.show()
mask2[1].iloc[0]['NCT Number']
city_lng, city_lat, city_name = [],[],[]
for i in range(len(city_site)):
city_lng.append(city_site[i]['Lng'].tolist())
city_lat.append(city_site[i]['Lat'].tolist())
city_name.append(city_site[i]['City, State'])
jit_lng, jit_lat, jit_trial_no, jit_trial_name = [],[],[],[]
for i in range(len(mask3)):
for j in range(len(mask3[i])):
jit_lng.append(mask3[i].iloc[j]['Lng'])
jit_lat.append(mask3[i].iloc[j]['Lat'])
jit_trial_no.append(mask3[i].iloc[j]['NCT Number'])
jit_trial_name.append(mask3[i].iloc[j]['Title'])
import folium
colors = []
for i in range(len(mask3)):
for j in range(len(mask3[i])):
colors.append(mask3[i].iloc[j]['Color'])
map = folium.Map(location = [38.58, -99.09], zoom_start=3.5, prefer_canvas=True, tiles = 'cartodbdark_matter', control_scale=True)
fg = folium.FeatureGroup(name = "My Map")
for lat, lng, number, name, col in zip(jit_lat, jit_lng, jit_trial_no, jit_trial_name, colors):
fg.add_child(folium.CircleMarker(location = [lat, lng], popup = number + ', ' + name, radius = 3, color = col))
for lat, lng, city in zip(city_lat, city_lng, city_name):
fg.add_child(folium.CircleMarker(location = [lat, lng], popup = city, radius = .2, color = "magenta"))
segments = []
for i in range(len(city_site)):
for j in range(len(mask3[i])):
segments.append(tuple([[city_site[i]['Lat'], city_site[i]['Lng']], [mask3[i].iloc[j]['Lat'], mask3[i].iloc[j]['Lng']]]))
for i in range(len(segments)):
fg.add_child(folium.PolyLine(locations=segments[i], color="white", weight=.10, opacity=20))
print 'CAR-T Trials in US as of ' + str(datetime.now())[:10]
map.add_child(fg)
map.save("Map1" + str(datetime.now())[:10] + ".html")
map
NCT_No = []
length = []
for i in range(len(mask2)):
NCT_No.append(mask2[i].iloc[0]['NCT Number'])
length.append(len(mask2[i]))
plt.figure(figsize=(7,10))
ax1 = plt.axes(frameon=False)
barlist = plt.barh(df_['NCT Number'].tolist(), length, alpha = 0.8)
for i in range(len(barlist)):
barlist[i].set_color(color[i])
plt.grid()
plt.title('Legend and Number of Sites per Trials \n')
plt.savefig('Legend_CART ' + str(datetime.now())[:10] + '.png', format='png', dpi=600, bbox_inches="tight", transparent=True)
plt.show()
from matplotlib.pyplot import figure
import mpld3
fig = plt.figure(figsize=(7,10))
plt.gca()
plt.axes(frameon=False)
barlist = plt.barh(df_['NCT Number'].tolist(), length, alpha = 0.8)
for i in range(len(barlist)):
barlist[i].set_color(color[i])
plt.grid()
plt.title('Legend and Number of Sites per Trials as of ' + str(datetime.now())[:10] + '\n')
mpld3.display()
mpld3.save_html(fig,'Legend_' + str(datetime.now())[:10] +'.html')
df_['NCT Number'].tolist()
import pygal
# from pygal.style import Style
# custom_style = Style(
# background='transparent',
# plot_background='transparent',
# foreground='#53E89B',
# foreground_strong='#53A0E8',
# foreground_subtle='#630C0D',
# opacity='.6',
# opacity_hover='.9',
# transition='400ms ease-in',
# colors=colors)
# bar_chart = pygal.HorizontalBar()
# for i in range(len(length)):
# bar_chart.add(df_['NCT Number'].tolist()[i], length[i])
# bar_chart.render_to_file('bar_chart.svg')
import pygal
from IPython.display import SVG, display
from pygal.style import Style
custom_style = Style(
background='transparent',
plot_background='transparent',
# foreground='#53E89B',
# foreground_strong='#53A0E8',
# foreground_subtle='#630C0D',
opacity='.4',
opacity_hover='.5',
transition='400ms ease-in',
colors=(color))
# chart = pygal.StackedLine(fill=True, interpolate='cubic', )
bar_chart = pygal.HorizontalBar(show_legend=False, height = 1000, spacing = 1, style=custom_style)
for i in range(len(length)):
# bar_chart.add(df_['NCT Number'].tolist()[i], )
bar_chart.add(df_['NCT Number'].tolist()[i], [{'value': length[i], 'label': df_['Title'].tolist()[i]}])
# chart.add('A', [1, 3, 5, 16, 13, 3, 7])
# chart.add('B', [5, 2, 3, 2, 5, 7, 17])
# chart.add('C', [6, 10, 9, 7, 3, 1, 0])
# chart.add('D', [2, 3, 5, 9, 12, 9, 5])
# chart.add('E', [7, 4, 2, 1, 2, 10, 0])
display({'image/svg+xml': bar_chart.render()}, raw=True)
bar_chart.render_to_file('bar_chart1' + str(datetime.now())[:10] + '.svg')
```
| github_jupyter |
```
import io
import pandas as pd
import matplotlib.pyplot as plt
import requests
from datetime import datetime, timedelta
import ipywidgets as widgets
from IPython.display import display
%matplotlib widget
class ShortDataDash:
def __init__(self):
self.df = pd.DataFrame()
self.days_slider = widgets.IntSlider(
value=5,
min=5,
max=252,
step=1,
description="Days Back",
style={"description_width": "initial"},
)
self.count_slider = widgets.IntSlider(
value=10,
min=1,
max=25,
step=1,
description="Number to show.",
style={"description_width": "initial"},
)
self.output1 = widgets.Output()
self.output2 = widgets.Output()
self.load_button = widgets.Button(
description="Load Data", layout=widgets.Layout(width="200px", height="40px")
)
self.load_button.on_click(self.load_button_click)
self.show_button = widgets.Button(
description="Change Number Shown", layout=self.load_button.layout
)
self.show_button.on_click(self.show_button_click)
self.slider_box = widgets.HBox([self.days_slider, self.count_slider])
self.button_box = widgets.VBox([self.load_button, self.show_button])
self.stock_input = widgets.Text(
value="GME",
placeholder="GME",
description="Ticker:",
)
self.ticker_button = widgets.Button(description="Plot Ticker")
self.ticker_button.on_click(self.ticker_button_click)
def show_button_click(self, b):
self.output1.clear_output()
with self.output1:
self.update()
def load_button_click(self, b):
self.output1.clear_output()
self.output2.clear_output()
with self.output1:
print(f"Data Loading for {self.days_slider.value} days")
self.fetch_new_data()
self.update()
def ticker_button_click(self, b):
self.output2.clear_output()
with self.output2:
self.ticker_plot()
def fetch_new_data(self):
self.df = pd.DataFrame()
today = datetime.now().date()
idx = 0
len_df = 0
while len_df < self.days_slider.value:
date = today - timedelta(days=idx)
r = requests.get(
f"https://cdn.finra.org/equity/regsho/daily/CNMSshvol{date.strftime('%Y%m%d')}.txt"
)
if r.status_code == 200:
self.df = pd.concat(
[self.df, pd.read_csv(io.StringIO(r.text), sep="|")], axis=0
)
len_df += 1
idx += 1
self.df = self.df[self.df.Date > 20100101]
self.df.Date = self.df["Date"].apply(
lambda x: datetime.strptime(str(x), "%Y%m%d")
)
def update(self):
if not self.df.empty:
temp = (
self.df.groupby("Symbol")[["ShortVolume", "TotalVolume"]]
.agg("sum")
.sort_values(by="ShortVolume", ascending=False)
.head(self.count_slider.value)[::-1]
)
self.fig, self.ax = plt.subplots(figsize=(6, 6))
self.ax.barh(temp.index, temp.TotalVolume, alpha=0.4, label="Total Volume")
self.ax.barh(temp.index, temp.ShortVolume, label="Short Volume")
self.ax.set_title(
f"Top {self.count_slider.value} Short Volume in Last {self.days_slider.value} Days"
)
self.ax.legend()
self.fig.tight_layout()
plt.show()
def ticker_plot(self):
stock_data = self.df.copy().loc[
self.df.Symbol == self.stock_input.value,
["Date", "ShortVolume", "TotalVolume"],
]
self.fig2, self.ax2 = plt.subplots(figsize=(6, 6))
self.ax2.plot(
stock_data.Date, stock_data.TotalVolume, alpha=0.4, label="Total Volume"
)
self.ax2.plot(stock_data.Date, stock_data.ShortVolume, label="Short Volume")
self.ax2.set_title(
f"Stock Volume and Short Volume for {self.stock_input.value.upper()}"
)
self.ax2.legend()
self.fig2.autofmt_xdate()
self.fig2.tight_layout()
plt.show()
def build_app(self):
title_html = """
<h2>Finra Short Data</h2>
<p>This widget downloads the consolidated NMS short data from FINRA and aggregates the data by summing over the entire time period.</p>
<p>Note that clicking the 'Load Data' button will reload all data. This can get time consuming, so if you pick a few hundred days, expect a few minutes for loading time.</p>
"""
middle_html = """
Here we allow the user to query for a single stock. This will work with the loaded data. Note that if you want to reload the data, this will once again take some time.
"""
return [
widgets.HTML(
title_html, layout=widgets.Layout(margin="0 0 3em 0", max_width="800px")
),
self.slider_box,
self.button_box,
self.output1,
widgets.HTML(
middle_html,
layout=widgets.Layout(margin="0 0 3em 0", max_width="800px"),
),
self.stock_input,
self.ticker_button,
self.output2,
]
dash = ShortDataDash()
app = widgets.VBox(
dash.build_app(), layout=widgets.Layout(max_width="1024px", margin="0 auto 0 auto")
)
display(app)
```
| github_jupyter |
Deep Learning Models -- A collection of various deep learning architectures, models, and tips for TensorFlow and PyTorch in Jupyter Notebooks.
- Author: Sebastian Raschka
- GitHub Repository: https://github.com/rasbt/deeplearning-models
```
%load_ext watermark
%watermark -a 'Sebastian Raschka' -v -p torch
```
- Runs on CPU or GPU (if available)
# Model Zoo -- Convolutional Neural Network
## Imports
```
import time
import numpy as np
import torch
import torch.nn.functional as F
from torchvision import datasets
from torchvision import transforms
from torch.utils.data import DataLoader
if torch.cuda.is_available():
torch.backends.cudnn.deterministic = True
```
## Settings and Dataset
```
##########################
### SETTINGS
##########################
# Device
device = torch.device("cuda:3" if torch.cuda.is_available() else "cpu")
# Hyperparameters
random_seed = 1
learning_rate = 0.05
num_epochs = 10
batch_size = 128
# Architecture
num_classes = 10
##########################
### MNIST DATASET
##########################
# Note transforms.ToTensor() scales input images
# to 0-1 range
train_dataset = datasets.MNIST(root='data',
train=True,
transform=transforms.ToTensor(),
download=True)
test_dataset = datasets.MNIST(root='data',
train=False,
transform=transforms.ToTensor())
train_loader = DataLoader(dataset=train_dataset,
batch_size=batch_size,
shuffle=True)
test_loader = DataLoader(dataset=test_dataset,
batch_size=batch_size,
shuffle=False)
# Checking the dataset
for images, labels in train_loader:
print('Image batch dimensions:', images.shape)
print('Image label dimensions:', labels.shape)
break
```
## Model
```
##########################
### MODEL
##########################
class ConvNet(torch.nn.Module):
def __init__(self, num_classes):
super(ConvNet, self).__init__()
# calculate same padding:
# (w - k + 2*p)/s + 1 = o
# => p = (s(o-1) - w + k)/2
# 28x28x1 => 28x28x8
self.conv_1 = torch.nn.Conv2d(in_channels=1,
out_channels=8,
kernel_size=(3, 3),
stride=(1, 1),
padding=1) # (1(28-1) - 28 + 3) / 2 = 1
# 28x28x8 => 14x14x8
self.pool_1 = torch.nn.MaxPool2d(kernel_size=(2, 2),
stride=(2, 2),
padding=0) # (2(14-1) - 28 + 2) = 0
# 14x14x8 => 14x14x16
self.conv_2 = torch.nn.Conv2d(in_channels=8,
out_channels=16,
kernel_size=(3, 3),
stride=(1, 1),
padding=1) # (1(14-1) - 14 + 3) / 2 = 1
# 14x14x16 => 7x7x16
self.pool_2 = torch.nn.MaxPool2d(kernel_size=(2, 2),
stride=(2, 2),
padding=0) # (2(7-1) - 14 + 2) = 0
self.linear_1 = torch.nn.Linear(7*7*16, num_classes)
# optionally initialize weights from Gaussian;
# Guassian weight init is not recommended and only for demonstration purposes
for m in self.modules():
if isinstance(m, torch.nn.Conv2d) or isinstance(m, torch.nn.Linear):
m.weight.data.normal_(0.0, 0.01)
m.bias.data.zero_()
if m.bias is not None:
m.bias.detach().zero_()
def forward(self, x):
out = self.conv_1(x)
out = F.relu(out)
out = self.pool_1(out)
out = self.conv_2(out)
out = F.relu(out)
out = self.pool_2(out)
logits = self.linear_1(out.view(-1, 7*7*16))
probas = F.softmax(logits, dim=1)
return logits, probas
torch.manual_seed(random_seed)
model = ConvNet(num_classes=num_classes)
model = model.to(device)
optimizer = torch.optim.SGD(model.parameters(), lr=learning_rate)
```
## Training
```
def compute_accuracy(model, data_loader):
correct_pred, num_examples = 0, 0
for features, targets in data_loader:
features = features.to(device)
targets = targets.to(device)
logits, probas = model(features)
_, predicted_labels = torch.max(probas, 1)
num_examples += targets.size(0)
correct_pred += (predicted_labels == targets).sum()
return correct_pred.float()/num_examples * 100
start_time = time.time()
for epoch in range(num_epochs):
model = model.train()
for batch_idx, (features, targets) in enumerate(train_loader):
features = features.to(device)
targets = targets.to(device)
### FORWARD AND BACK PROP
logits, probas = model(features)
cost = F.cross_entropy(logits, targets)
optimizer.zero_grad()
cost.backward()
### UPDATE MODEL PARAMETERS
optimizer.step()
### LOGGING
if not batch_idx % 50:
print ('Epoch: %03d/%03d | Batch %03d/%03d | Cost: %.4f'
%(epoch+1, num_epochs, batch_idx,
len(train_loader), cost))
model = model.eval()
print('Epoch: %03d/%03d training accuracy: %.2f%%' % (
epoch+1, num_epochs,
compute_accuracy(model, train_loader)))
print('Time elapsed: %.2f min' % ((time.time() - start_time)/60))
print('Total Training Time: %.2f min' % ((time.time() - start_time)/60))
```
## Evaluation
```
with torch.set_grad_enabled(False): # save memory during inference
print('Test accuracy: %.2f%%' % (compute_accuracy(model, test_loader)))
%watermark -iv
```
| github_jupyter |
## Visualize image-specific class saliency with backpropagation
---
A quick demo of creating saliency maps for CNNs using [FlashTorch 🔦](https://github.com/MisaOgura/flashtorch).
❗This notebook is for those who are using this notebook in **Google Colab**.
If you aren't on Google Colab already, please head to the Colab version of this notebook **[here](https://colab.research.google.com/github/MisaOgura/flashtorch/blob/master/examples/visualise_saliency_with_backprop_colab.ipynb)** to execute.
---
The gradients obtained can be used to visualise an image-specific class saliency map, which can gives some intuition on regions within the input image that contribute the most (and least) to the corresponding output.
More details on saliency maps: [Deep Inside Convolutional Networks: Visualising Image Classification Models and Saliency Maps](https://arxiv.org/pdf/1312.6034.pdf).
### 0. Set up
A GPU runtime is available on Colab for free, from the `Runtime` tab on the top menu bar.
It is **recommended to use GPU** as a runtime for the enhanced speed of computation.
```
# Install flashtorch
!pip install flashtorch torch==1.5.0 torchvision==0.6.0 -U
# Download example images
!mkdir -p images
!wget -nv \
https://github.com/MisaOgura/flashtorch/raw/master/examples/images/great_grey_owl.jpg \
https://github.com/MisaOgura/flashtorch/raw/master/examples/images/peacock.jpg \
https://github.com/MisaOgura/flashtorch/raw/master/examples/images/toucan.jpg \
-P /content/images
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
import matplotlib.pyplot as plt
import torchvision.models as models
from flashtorch.utils import apply_transforms, load_image
from flashtorch.saliency import Backprop
```
### 1. Load an image
```
image = load_image('/content/images/great_grey_owl.jpg')
plt.imshow(image)
plt.title('Original image')
plt.axis('off');
```
### 2. Load a pre-trained Model
```
model = models.alexnet(pretrained=True)
```
### 3. Create an instance of Backprop with the model
```
backprop = Backprop(model)
```
### 4. Visualize saliency maps
```
# Transform the input image to a tensor
owl = apply_transforms(image)
# Set a target class from ImageNet task: 24 in case of great gray owl
target_class = 24
# Ready to roll!
backprop.visualize(owl, target_class, guided=True, use_gpu=True)
```
### 5. What about other birds?
What makes peacock a peacock...?
```
peacock = apply_transforms(load_image('/content/images/peacock.jpg'))
backprop.visualize(peacock, 84, guided=True, use_gpu=True)
```
Or a toucan?
```
toucan = apply_transforms(load_image('/content/images/toucan.jpg'))
backprop.visualize(toucan, 96, guided=True, use_gpu=True)
```
Please try out other models/images too!
| github_jupyter |
[this doc on github](https://github.com/dotnet/interactive/tree/master/samples/notebooks/csharp/Docs)
# Formatting Outputs
## HTML Formatting
When you return a value or a display a value in a .NET notebook, the default formatting behavior is normally uses HTML to try to provide some useful information about the object.
### Enumerables
If it's an array or other type implementing `IEnumerable`, that might look like this:
```
display(new [] {"hello", "world"} );
Enumerable.Range(1, 5)
```
As you can see, the same basic structure is used whether you pass the object to the `display` method or return it as the cell's value.
### Objects
Similarly to the behavior for `IEnumerable` objects, you'll also see table output for dictionaries, but for each value in the dictionary, the key is provided rather than the index within the collection.
```
var dictionary = new Dictionary<string, int>
{
["zero"] = 0,
["one"] = 1,
["two"] = 2
};
dictionary
```
The default formatting behavior for other types of objects is to produce a table showing their properties and the values of those properties.
```
class Person
{
public string FirstName { get; set; }
public string LastName { get; set; }
public int Age { get; set; }
}
display(new Person { FirstName = "Mitch", LastName = "Buchannon", Age = 42} );
```
When you have a collection of such objects, you can see the values listed for each item in the collection:
```
var groupOfPeople = new []
{
new Person { FirstName = "Mitch", LastName = "Buchannon", Age = 42 },
new Person { FirstName = "Hobie ", LastName = "Buchannon", Age = 23 },
new Person { FirstName = "Summer", LastName = "Quinn", Age = 25 },
new Person { FirstName = "C.J.", LastName = "Parker", Age = 23 },
};
display(groupOfPeople);
```
### Dictionaries
Displaying a dictionary will show the items by key rather than index.
```
display(groupOfPeople.ToDictionary(p => $"{p.FirstName}"));
```
Now let's try something a bit more complex. Let's look at a graph of objects.
We'll redefine the `Person` class to allow a reference to a collection of other `Person` instances.
```
class Person
{
public string FirstName { get; set; }
public string LastName { get; set; }
public int Age { get; set; }
public List<Person> Friends { get; } = new List<Person>();
}
var mitch = new Person { FirstName = "Mitch", LastName = "Buchannon", Age = 42 };
var hobie = new Person { FirstName = "Hobie ", LastName = "Buchannon", Age = 23 };
var summer = new Person { FirstName = "Summer", LastName = "Quinn", Age = 25 };
var cj = new Person { FirstName = "C.J.", LastName = "Parker", Age = 23 };
mitch.Friends.AddRange(new [] { hobie, summer, cj });
hobie.Friends.AddRange(new [] { mitch, summer, cj });
summer.Friends.AddRange(new [] { mitch, hobie, cj });
cj.Friends.AddRange(new [] { mitch, hobie, summer });
var groupOfPeople = new List<Person> { mitch, hobie, summer, cj };
display(groupOfPeople);
```
That's a bit hard to read, right? The defaut formatting behaviors are not always as useful as they might be. In order to give you more control object formatters can be customized from within the .NET notebook.
## Customization
## Registering plain text formatters
Let's clean up the output above by customizing the formatter for the `Person.Friends` property, which is creating a lot of noise.
The way to do this is to use the `Formatter` API. This API lets you customize the formatting for a specific type. For example:
```
Formatter.Register<Person>((person, writer) => {
writer.Write("person");
}, mimeType: "text/plain");
groupOfPeople
```
With that in mind, we can make it even more concise by registering a good formatter for `Person`:
```
Formatter.ResetToDefault();
Formatter.Register<Person>((person, writer) => {
writer.Write(person.FirstName);
}, mimeType: "text/plain");
groupOfPeople
```
### Registering HTML formatters
To replace the default HTML table view, you can register a formatter for the `"text/html"` mime type. Let's do that, and write some HTML using PocketView.
```
Formatter.ResetToDefault();
Formatter.Register<List<Person>>((people, writer) =>
{
foreach (var person in people)
{
writer.Write(
span(
b(person.FirstName),
" ",
i($"({person.Age} years old and has {person.Friends.Count} friends)"),
br));
}
}, mimeType: "text/html");
groupOfPeople
```
---
**_See also_**
* [Displaying output](Displaying%20output.ipynb)
* [HTML](HTML.ipynb)
| github_jupyter |
<script async src="https://www.googletagmanager.com/gtag/js?id=UA-59152712-8"></script>
<script>
window.dataLayer = window.dataLayer || [];
function gtag(){dataLayer.push(arguments);}
gtag('js', new Date());
gtag('config', 'UA-59152712-8');
</script>
# Adding Unit Tests to the NRPy+ Unit Testing Infrastructure
## Author: Kevin Lituchy
## Introduction:
The goal of this module is to give the user an overview/understanding of NRPy+'s Unit Testing framework, which will give the user enough information to begin creating their own unit tests. We will begin by giving an overview of the important prerequisite knowledge to make the most out of unit tests. Next, we give an explanation for the user interaction within the unit testing framework; this will give the user the ability to create tests for themselves. Then we give the user some insight into interpreting the output from their unit tests. Finally, a full example using a test module will be run through in full, both with and without errors, to give the user a realistic sense of what unit testing entails.
For in-depth explanations of all subfunctions (not user-interactable), see the [UnitTesting Function Reference](./UnitTesting/UnitTesting_Function_Reference.ipynb). This may not be essential to get unit tests up and running, but it will be invaluable information if the user ever wants to make modifications to the unit testing code.
<a id='toc'></a>
# Table of Contents
$$\label{toc}$$
This notebook is organized as follows:
1. [Step 1](#motivation): Motivation and Prerequisite Knowledge
1. [Step 1.a](#dicts): Dictionaries
1. [Step 1.b](#logging): Logging
1. [Step 2](#interaction): User Interaction
1. [Step 2.a](#testfile): Test File
1. [Step 2.b](#trustedvaluesdict): trusted_values_dict
1. [Step 2.c](#bash): Bash Script
1. [Step 3](#output): Interpreting output
1. [Step 4](#checklist): Checklist
1. [Step 4.a](#4a): Directory Creation
1. [Step 4.b](#4b): Test File Creation
1. [Step 4.c](#4c): Input Parameters
1. [Step 4.d](#4d): Fill-in bash script and run
1. [Step 5](#latex_pdf_output): Output this notebook to $\LaTeX$-formatted PDF
<a id='motivation'></a>
# Step 1: Motivation and Prerequisite Knowledge \[Back to [top](#toc)\]
$$\label{motivation}$$
What is the purpose of unit testing, and why should you do it? To begin
thinking about that, consider what subtleties can occur within your code
that are almost unnoticeable to the eye, but wind up giving you an
incorrect result. You could make a small optimization, and not notice
any change in your result. However, maybe the optimization you made only
works on Python 3 and not Python 2, or it changes a value by some tiny
amount -- too small to be noticeable at a simple glance, but enough to
make a difference in succeeding calculations.
This is where unit testing comes in. By initially calculating values for
the globals of your modules in a **trusted** version of your code and
storing those values in a dictionary, you can then easily check if
something stopped working correctly by comparing your newly calculated
values to the ones you've stored. On the frontend, there are four
concepts essential to understand to get your unit tests up and running:
`trusted_values_dict`, `create_test`, your testing module (which
will simply be referred to as `test_file`), and a bash script (which
will simply be referred to as `bash_script`). There is also some
important prerequisite knowledge that may be helpful to grasp before
beginning your testing. There are many functions at play in the backend
as well, all of which are described in the Function Reference. Mastery of these functions may not be
essential to get your tests up-and-running, but some basic understanding
of them with undoubtedly help with debugging.
An important caveat is that the unit testing does not test the
**correctness** of your code or your variables. The unit tests act as a
protective measure to ensure that nothing was broken between versions of
your code; it gets its values by running _your_ code, so if something
starts out incorrect, it will be stored as incorrect in the system.
<a id='dicts'></a>
## Step 1.a: Dictionaries \[Back to [top](#toc)\]
$$\label{dicts}$$
Dictionaries are used throughout the unit testing infrastructure. The user must create simple dictionaries to pass to our testing functions. If you know nothing about dictionaries, we recommend [this](https://www.w3schools.com/python/python_dictionaries.asp) article; it will get you up to speed for simple dictionary creation.
<a id='logging'></a>
## Step 1.b: Logging \[Back to [top](#toc)\]
$$\label{logging}$$
Logging is a python module that allows the user to specify their desired level of output by modifying a parameter, rather than having to use if-statements and print-statements. We allow the user to change the level of output through a parameter `logging_level`, in which we support the following levels:
`ERROR`: Only print when an error occurs
`INFO`: Print general information about test beginning, completion, major function calls, etc., as well as everything above. (Recommended)
`DEBUG`: Print maximum amount of information -- every comparison, as well as everything above.
A good way to think of these logging levels is that `INFO` is the default, `ERROR` is similar to a non-verbose mode, and `DEBUG` is similar to a verbose mode.
<a id='interaction'></a>
# Step 2: User Interaction \[Back to [top](#toc)\]
$$\label{interaction}$$
Within the module the user is intending to test, a directory named `tests` should be created. This will house the test file for the given module and its associated `trusted_values_dict`. For example, if I intend to test `BSSN`, I will create a new directory `BSSN/tests`. Within the `tests` directory, the user should create a file called `test_(module).py` -- or `test_BSSN.py` for the given example.
<a id='testfile'></a>
## Step 2.a: Test File \[Back to [top](#toc)\]
$$\label{testfile}$$
The test file is how the user inputs their module, functions, and globals information to the testing suite. For the purpose of consistency, we've created a skeleton for the test file (found [here](../edit/UnitTesting/test_skeleton.py)) that contains all information the user must specify. The user should change the name of the function to something relevant to their test. However, note that the name of the function must begin with `test_` in order for the bash script to successfully run the test -- this is the default naming scheme for most test suites/software. Inside the function, multiple fields are required to be filled out by the user; these fields are `module`, `module_name`, and `function_and_global_dict`. Below the function there is some code that begins with `if __name__ == '__main__':`. The user can ignore this code as it does backend work and makes sure to pass the proper information for the test.
`module` is a string representing the module to be tested.
`module_name` is a string representing the name of the module
`function_and_global_dict` is a dictionary whose keys are string representations of functions that the user would like to be called on `module` and whose values are lists of string representations of globals that can be acquired by running their respective functions on `module`
Example:
```
def test_BrillLindquist():
module = 'BSSN.BrillLindquist'
module_name = 'bl'
function_and_global_dict = {'BrillLindquist(ComputeADMGlobalsOnly = True)':
['alphaCart', 'betaCartU', 'BCartU', 'gammaCartDD', 'KCartDD']}
create_test(module, module_name, function_and_global_dict)
```
In most cases, this simple structure is enough to do exactly what the user wants. Sometimes, however, there is other information that needs to be passed into the test -- this is where optional arguments come in.
The tests can take two optional arguments, `logging_level` and `initialization_string_dict`
`logging_level` follows the same scheme as described [above](#logging).
`initialization_string_dict` is a dictionary whose keys are functions that **must** also be in `function_and_global_dict` and whose values are strings containing well-formed Python code. The strings are executed as Python code before their respective function is called on the module. The purpose of this argument is to allow the user to do any necessary NRPy+ setup before the call their function.
Example:
```
def test_quantities():
module = 'BSSN.BSSN_quantities'
module_name = 'BSSN_quantities'
function_and_global_dict = {'BSSN_basic_tensors()': ['gammabarDD', 'AbarDD', 'LambdabarU', 'betaU', 'BU']}
logging_level = 'DEBUG'
initialization_string = '''
import reference_metric as rfm
rfm.reference_metric()
rfm.ref_metric__hatted_quantities()
'''
initialization_string_dict = {'BSSN_basic_tensors()': initialization_string}
create_test(module, module_name, function_and_global_dict, logging_level=logging_level,
initialization_string_dict=initialization_string_dict)
```
An important thing to note is that even though `initialization_string` looks odd with its indentation, this is necessary for Python to interpret it correctly. If it was indented, Python would think you were trying to indent that code when it shouldn't be, and an error will occur.
A question you may be wondering is why we need to create a new dictionary for the intialization string, insted of just passing it as its own argument. This is because the testing suite can accept multiple function calls, each with their own associated global list, in one function. It then naturally follows that we need `initailization_string_dict` to allow each function call to have its own code that runs before its function call. In the following example, the function `BSSN_basic_tensors()` has an initialization string, but the function `declare_BSSN_gridfunctions_if_not_declared_already()` doesn't. You can also clearly see they each have their own associated globals.
Example:
```
def test_quantities():
module = 'BSSN.BSSN_quantities'
module_name = 'BSSN_quantities'
function_and_global_dict = {'declare_BSSN_gridfunctions_if_not_declared_already()':
['hDD', 'aDD', 'lambdaU', 'vetU', 'betU', 'trK', 'cf', 'alpha'],
'BSSN_basic_tensors()': ['gammabarDD', 'AbarDD', 'LambdabarU', 'betaU', 'BU']}
logging_level = 'DEBUG'
initialization_string = '''
import reference_metric as rfm
rfm.reference_metric()
rfm.ref_metric__hatted_quantities()
'''
initialization_string_dict = {'BSSN_basic_tensors()': initialization_string}
create_test(module, module_name, function_and_global_dict, logging_level=logging_level,
initialization_string_dict=initialization_string_dict)
```
Lastly, within a single test file, you can define multiple test functions. It's as simple as defining a new function whose name starts with `test_` in the file and making sure to fill out the necessary fields.
<a id='trustedvaluesdict'></a>
## Step 2.b: trusted_values_dict \[Back to [top](#toc)\]
$$\label{trustedvaluesdict}$$
At this point, it's should be understood that our test suite will compare trusted values of your variables to newly calculated values to ensure that no variables were unknowingly modified. The `trusted_values_dict` acts as the means of storing the trusted value for each variable with the purpose of future comparison. A new `trusted_values_dict` is created by default when a test file is run for the first time -- it's visible in `tests/trusted_values_dict.py`. Note that if you run your code but can't see the file, refresh your IDE -- it's there, sometimes IDE's just get confused when you create a file within Python. The default structure of all `trusted_value_dict` files is as follows:
```
from mpmath import mpf, mp, mpc
from UnitTesting.standard_constants import precision
mp.dps = precision
trusted_values_dict = {}
```
The proper code to copy into this file will be printed to the console when a test is run. The test suite will also automatically write its calculated globals' values for a given function to this file in the proper format; make sure to check that things seem correct though! Remember that the `trusted_values_dict` stores **trusted**, not necessarily **correct**, values for each global.
<a id='bash'></a>
## Step 2.c: Bash Script \[Back to [top](#toc)\]
$$\label{bash}$$
In order to successfully run all the user's unit tests and properly integrate testing with TravisCI, we use a bash script as the 'hub' of all the tests to be run. This makes it easy for the user to comment out tests they don't want to run, add new tests to be automatically run with one line, etc.
We offer a skeleton file, [`run_NRPy_UnitTests`](../edit/UnitTesting/run_NRPy_UnitTests.sh), which contains all the proper code to be easily run with minimum user interaction. All the user must do is call the `add_test` function on the test file they'd like to be run underneath the `TODO` comment. There are many examples in the file that show exactly how to create a new test. Then to add more tests, simply go to the next line and add another test. It's as simple as that!
To run the bash script, open up a terminal, type in the path of the bash script, and then pick the Python interpreter to run the code -- for example, `./UnitTesting/run_NRPy_UnitTests.sh python` or `./UnitTesting/run_NRPy_UnitTests.sh python3`.
There's an additional field in the bash script called `rerun_if_fail`. It is a boolean that, if true, will automatically rerun the tests that failed with their `logging_level` set to `DEBUG`. This gives the user a plethora of debugging information that should make it much easier to figure out the issue. We'd recommend enabling it if there are only a couple modules that failed, as there is a very large amount of information printed for each failing module. However, it is an invaluable resource for figuring out a bug in your code, so keep it in mind when tests are failing.
<a id='output'></a>
# Step 3: Interpreting Output \[Back to [top](#toc)\]
$$\label{output}$$
Once a user's tests are fully set up, they need to be able to interpret the output of their tests; doing this allows the user to easily figure out what went wrong, why, and how to fix it. The amount of output for a given module is of course dependent on its logging level. For the purposes of this tutorial, we will assume that `logging_level` is set to `INFO`.
While running a test, output is printed to the console that tells the user what is occurring at what point in time.
Example successful test run console output:
```
Testing test_u0_smallb_Poynting__Cartesian...
INFO:root: Creating file /home/kevin/virtpypy/nrpyunittesting/u0_smallb_Poynting__Cartesian/tests/u0sbPoyn__compute_u0_smallb_Poynting__Cartesian__test.py...
INFO:root: ...Success: File created.
INFO:root: Currently working on function compute_u0_smallb_Poynting__Cartesian() in module u0sbPoyn...
INFO:root: Importing trusted_values_dict...
INFO:root: ...Success: Imported trusted_values_dict.
INFO:root: Calling evaluate_globals...
INFO:root: ...Success: evaluate_globals ran without errors.
INFO:root: Calling cse_simplify_and_evaluate_sympy_expressions...
INFO:root: ...Success: cse_simplify_and_evaluate_sympy_expressions ran without errors.
INFO:root: Calling calc_error...
INFO:root: ...Success: calc_error ran without errors.
.
----------------------------------------------------------------------
Ran 1 test in 3.550s
OK
INFO:root: Test for function compute_u0_smallb_Poynting__Cartesian() in module u0sbPoyn passed! Deleting test file...
INFO:root: ...Deletion successful. Test complete.
----------------------------------------------------------------------
----------------------------------------------------------------------
All tests passed!
```
Step-by-step, how do we interpret this output?
The first line tells the user what is being called -- this is the function that they define in their current test file being run; this is why a descriptive function name is important.
Next, a file is created in the same directory as the current test file that actually runs the tests -- a new file is created to ensure that a clean Python environment is used for each test.
Each function test within the test file is then called successively -- in this example, that is `compute_u0_smallb_Poynting__Cartesian()` in module `u0sbPoyn`.
The function's associated `trusted_values_dict` is imported.
Each global in the function's global list is then evaluated to a SymPy expression in `evaluate_globals`.
Each global's SymPy expression is then evaluated into a numerical value in `cse_simplify_and_evaluate_sympy_expressions`.
Each global is compared to its trusted value in `calc_error`.
In this example, since no values differed, the test passed -- there is nothing the user has to do.
The purpose of giving the user this output is to make it as easy as possible, when something fails, to figure out why and how to fix it. Say the user wasn't given the above output -- instead, after `INFO:root: Calling calc_error...` is printed, an error is printed and the program quits -- then the user knows the error occurred somewhere in `calc_error`, and it's easy to figure out what. If this output wasn't given, it would be extremely difficult to bugfix.
Now let's consider example output when the trusted values for a couple globals differ from the newly calculated values:
```
Testing function test_u0_smallb_Poynting__Cartesian...
INFO:root: Creating file /home/kevin/virtpypy/nrpyunittesting/u0_smallb_Poynting__Cartesian/tests/u0sbPoyn__compute_u0_smallb_Poynting__Cartesian__test.py...
INFO:root: ...Success: File created.
INFO:root: Currently working on function compute_u0_smallb_Poynting__Cartesian() in module u0sbPoyn...
INFO:root: Importing trusted_values_dict...
INFO:root: ...Success: Imported trusted_values_dict.
INFO:root: Calling evaluate_globals...
INFO:root: ...Success: evaluate_globals ran without errors.
INFO:root: Calling cse_simplify_and_evaluate_sympy_expressions...
INFO:root: ...Success: cse_simplify_and_evaluate_sympy_expressions ran without errors.
INFO:root: Calling calc_error...
ERROR:root:
Variable(s) ['g4DD[0][0]', 'g4DD[0][1]'] in module u0sbPoyn failed. Please check values.
If you are confident that the newly calculated values are correct, comment out the old trusted values for
u0sbPoyn__compute_u0_smallb_Poynting__Cartesian__globals in your trusted_values_dict and copy the following code between the ##### into your trusted_values_dict.
Make sure to fill out the TODO comment describing why the values had to be changed. Then re-run test script.
#####
# Generated on: 2019-08-14
# Reason for changing values: TODO
trusted_values_dict['u0sbPoyn__compute_u0_smallb_Poynting__Cartesian__globals'] = {'g4DD[0][0]': mpf('1.42770464273047624140299713523'), 'g4DD[0][1]': mpf('0.813388473397507463814385913308'), 'g4DD[0][2]': mpf('0.652706348793296836714132090803'), 'g4DD[0][3]': mpf('1.22429414375154980405074869244'), 'g4DD[1][0]': mpf('0.813388473397507463814385913308'), 'g4DD[1][1]': mpf('0.657497767033916602485987823457'), 'g4DD[1][2]': mpf('0.057738705167452830657737194997'), 'g4DD[1][3]': mpf('0.391026617743468030141684721457'), 'g4DD[2][0]': mpf('0.652706348793296836714132090803'), 'g4DD[2][1]': mpf('0.057738705167452830657737194997'), 'g4DD[2][2]': mpf('0.142350778742078798444481435581'), 'g4DD[2][3]': mpf('0.723120760610660329170684690325'), 'g4DD[3][0]': mpf('1.22429414375154980405074869244'), 'g4DD[3][1]': mpf('0.391026617743468030141684721457'), 'g4DD[3][2]': mpf('0.723120760610660329170684690325'), 'g4DD[3][3]': mpf('0.919283767179900235255729512573'), 'g4UU[0][0]': mpf('-3.03008926847944211197781568781'), 'g4UU[0][1]': mpf('2.25487680174746330097911429618'), 'g4UU[0][2]': mpf('0.883964088219310673292773829627'), 'g4UU[0][3]': mpf('2.38097417962378184338842987037'), 'g4UU[1][0]': mpf('2.25487680174746330097911429618'), 'g4UU[1][1]': mpf('-0.109478866681308257858746913533'), 'g4UU[1][2]': mpf('-1.57673807708112257475456728614'), 'g4UU[1][3]': mpf('-1.71617440778374167644096697698'), 'g4UU[2][0]': mpf('0.883964088219310673292773829627'), 'g4UU[2][1]': mpf('-1.57673807708112257475456728614'), 'g4UU[2][2]': mpf('-2.06437348264308527564608946534'), 'g4UU[2][3]': mpf('1.11728919891515454886216209391'), 'g4UU[3][0]': mpf('2.38097417962378184338842987037'), 'g4UU[3][1]': mpf('-1.71617440778374167644096697698'), 'g4UU[3][2]': mpf('1.11728919891515454886216209391'), 'g4UU[3][3]': mpf('-2.23203972375107587678882970577'), 'PoynSU[0]': mpf('0.103073801363157111172177901697'), 'PoynSU[1]': mpf('0.11100316917740755485837448786'), 'PoynSU[2]': mpf('-0.00451075406485067218999293829888'), 'smallb2etk': mpf('0.164454779456120937541853683919'), 'smallb4D[0]': mpf('0.567950228622914592095169687713'), 'smallb4D[1]': mpf('0.286535540626704686153523625219'), 'smallb4D[2]': mpf('0.10714698030450909234705495631'), 'smallb4D[3]': mpf('0.455828728852934996540499932291'), 'smallb4U[0]': mpf('0.105192967134481035810628308481'), 'smallb4U[1]': mpf('0.298063886336868595407751154048'), 'smallb4U[2]': mpf('0.338357239072152954142217077157'), 'smallb4U[3]': mpf('-0.0371837983175520496394928051176'), 'u0': mpf('0.751914772923022001194226504595'), 'uBcontraction': mpf('0.214221928967111307128784385185'), 'uD[0]': mpf('0.216251888123560253383388291707'), 'uD[1]': mpf('0.167535113620266400428280039145'), 'uD[2]': mpf('0.232536332826570514618343904792'), 'uU[0]': mpf('-0.364066660324468905733189036042'), 'uU[1]': mpf('-0.0378849772494775056256865716175'), 'uU[2]': mpf('-0.476480636313712572229280970632')}
#####
.
----------------------------------------------------------------------
Ran 1 test in 3.481s
OK
ERROR:root: Test for function compute_u0_smallb_Poynting__Cartesian() in module u0sbPoyn failed! Please examine test file.
----------------------------------------------------------------------
----------------------------------------------------------------------
Tests failed!
Failures:
u0_smallb_Poynting__Cartesian/tests/test_u0_smallb_Poynting__Cartesian.py: ['test_u0_smallb_Poynting__Cartesian']
----------------------------------------------------------------------
```
This seems like a lot to take-in, but it's not too difficult to understand once fairly well acquainted with the output. The beginning is identical to the output from the successful test run, up until `calc_error` is called. This gives the user some insight that there was an error during `calc_error`, which gives an indication that at least one global had a different calculated and trusted value.
The next line confirms this suspicion: globals `g4DD[0][0]` and `g4DD[0][1]` had differing trusted and calculated values. This gives the user the **exact** information they're looking for. `g4DD` is a rank-2 tensor as per NRPy+ naming convention, and the user can very easily see the indices that failed. If the user expected this -- say they found a bug in their code that generated this global -- they can then copy the new `trusted_values_dict` entry into their `trusted_values_dict` and comment-out/delete the old entry. Then by re-running the test, there should no longer be an error -- the trusted and calculated values should be the same.
The next output tells the user that the test failed, and to examine the test file. This is necessary if theres an unexpected failure -- it will help the user figure out why it occurred.
Finally, additional output is given that tells the user all the test files and their respective functions that failed. This may seem repetitive, but there's a good reason for it. Say 20 modules were being tested, 10 of which had failures. Then to figure out what failed, the user would have to scroll through all the output and keep a mental note of what failed. By printing everything that failed at the very end, the user gets instant insight into what failed; this may help the user figure out why.
<a id='checklist'></a>
# Step 4: Checklist for adding a new test \[Back to [top](#toc)\]
$$\label{checklist}$$
<a id='4a'></a>
## Step 4.a: Directory Creation \[Back to [top](#toc)\]
$$\label{4a}$$
Create a `tests` directory in the directory of the module being tested.
<a id='4b'></a>
## Step 4.b: Test File Creation \[Back to [top](#toc)\]
$$\label{4b}$$
Create a `test_file.py` in the `tests` directory based off [UnitTesting/test_skeleton.py](../edit/UnitTesting/test_skeleton.py)
<a id='4c'></a>
## Step 4c: Input Parameters \[Back to [top](#toc)\]
$$\label{4c}$$
Change the name of `test_your_module()` to whatever you're testing, making sure the name starts with `test_`. Fill in the following paremeters in `test_file.py`: `module`, `module_name`, `function_and_global_dict`, and if need be, `logging_level` and `initialization_string`.
<a id='4d'></a>
## Step 4d: Fill-in bash script and run \[Back to [top](#toc)\]
$$\label{4d}$$
Use the `add_test` function in the [run_NRPy_UnitTests.sh](../edit/UnitTesting/run_NRPy_UnitTests.sh) to create your test below the `TODO` line. Run the bash script to run your test!
<a id='latex_pdf_output'></a>
# Step 5: Output this notebook to $\LaTeX$-formatted PDF file \[Back to [top](#toc)\]
$$\label{latex_pdf_output}$$
The following code cell converts this Jupyter notebook into a proper, clickable $\LaTeX$-formatted PDF file. After the cell is successfully run, the generated PDF may be found in the root NRPy+ tutorial directory, with filename [Tutorial-UnitTesting.pdf](Tutorial-UnitTesting.pdf) (Note that clicking on this link may not work; you may need to open the PDF file through another means.)
```
import cmdline_helper as cmd # NRPy+: Multi-platform Python command-line interface
cmd.output_Jupyter_notebook_to_LaTeXed_PDF("Tutorial-UnitTesting")
```
| github_jupyter |
#### New to Plotly?
Plotly's Python library is free and open source! [Get started](https://plot.ly/python/getting-started/) by downloading the client and [reading the primer](https://plot.ly/python/getting-started/).
<br>You can set up Plotly to work in [online](https://plot.ly/python/getting-started/#initialization-for-online-plotting) or [offline](https://plot.ly/python/getting-started/#initialization-for-offline-plotting) mode, or in [jupyter notebooks](https://plot.ly/python/getting-started/#start-plotting-online).
<br>We also have a quick-reference [cheatsheet](https://images.plot.ly/plotly-documentation/images/python_cheat_sheet.pdf) (new!) to help you get started!
#### Defining and plotting triangulated surfaces
#### with Plotly `Mesh3d`
A triangulation of a compact surface is a finite collection of triangles that cover the surface in such a way that every point on the surface is in a triangle, and the intersection of any two triangles is either void, a common edge or a common vertex. A triangulated surface is called tri-surface.
The triangulation of a surface defined as the graph of a continuous function, $z=f(x,y), (x,y)\in D\subset\mathbb{R}^2$ or in a parametric form:
$$x=x(u,v), y=y(u,v), z=z(u,v), (u,v)\in U\subset\mathbb{R}^2,$$
is the image through $f$,respectively through the parameterization, of the Delaunay triangulation or an user defined triangulation of the planar domain $D$, respectively $U$.
The Delaunay triangulation of a planar region is defined and illustrated in a Python Plotly tutorial posted [here](https://plot.ly/python/alpha-shapes/).
If the planar region $D$ ($U$) is rectangular, then one defines a meshgrid on it, and the points
of the grid are the input points for the `scipy.spatial.Delaunay` function that defines the planar triangulation of $D$, respectively $U$.
### Triangulation of the Moebius band ###
The Moebius band is parameterized by:
$$\begin{align*}
x(u,v)&=(1+0.5 v\cos(u/2))\cos(u)\\
y(u,v)&=(1+0.5 v\cos(u/2))\sin(u)\quad\quad u\in[0,2\pi],\: v\in[-1,1]\\
z(u,v)&=0.5 v\sin(u/2)
\end{align*}
$$
Define a meshgrid on the rectangle $U=[0,2\pi]\times[-1,1]$:
```
import plotly.plotly as py
import plotly.graph_objs as go
import numpy as np
import matplotlib.cm as cm
from scipy.spatial import Delaunay
u=np.linspace(0,2*np.pi, 24)
v=np.linspace(-1,1, 8)
u,v=np.meshgrid(u,v)
u=u.flatten()
v=v.flatten()
#evaluate the parameterization at the flattened u and v
tp=1+0.5*v*np.cos(u/2.)
x=tp*np.cos(u)
y=tp*np.sin(u)
z=0.5*v*np.sin(u/2.)
#define 2D points, as input data for the Delaunay triangulation of U
points2D=np.vstack([u,v]).T
tri = Delaunay(points2D)#triangulate the rectangle U
```
`tri.simplices` is a `np.array` of integers, of shape (`ntri`,3), where `ntri` is the number of triangles generated by `scipy.spatial.Delaunay`.
Each row in this array contains three indices, i, j, k, such that points2D[i,:], points2D[j,:], points2D[k,:] are vertices of a triangle in the Delaunay triangularization of the rectangle $U$.
```
print tri.simplices.shape, '\n', tri.simplices[0]
```
The images of the `points2D` through the surface parameterization are 3D points. The same simplices define the triangles on the surface.
Setting a combination of keys in `Mesh3d` leads to generating and plotting of a tri-surface, in the same way as `plot_trisurf` in matplotlib or `trisurf` in Matlab does.
We note that `Mesh3d` with different combination of keys can generate [alpha-shapes](https://plot.ly/python/alpha-shapes/).
In order to plot a tri-surface, we choose a colormap, and associate to each triangle on the surface, the color in colormap, corresponding to the normalized mean value of z-coordinates of the triangle vertices.
Define a function that maps a mean z-value to a matplotlib color, converted to a Plotly color:
```
def map_z2color(zval, colormap, vmin, vmax):
#map the normalized value zval to a corresponding color in the colormap
if vmin>vmax:
raise ValueError('incorrect relation between vmin and vmax')
t=(zval-vmin)/float((vmax-vmin))#normalize val
R, G, B, alpha=colormap(t)
return 'rgb('+'{:d}'.format(int(R*255+0.5))+','+'{:d}'.format(int(G*255+0.5))+\
','+'{:d}'.format(int(B*255+0.5))+')'
```
To plot the triangles on a surface, we set in Plotly `Mesh3d` the lists of x, y, respectively z- coordinates of the vertices, and the lists of indices, i, j, k, for x, y, z coordinates of all vertices:
```
def tri_indices(simplices):
#simplices is a numpy array defining the simplices of the triangularization
#returns the lists of indices i, j, k
return ([triplet[c] for triplet in simplices] for c in range(3))
def plotly_trisurf(x, y, z, simplices, colormap=cm.RdBu, plot_edges=None):
#x, y, z are lists of coordinates of the triangle vertices
#simplices are the simplices that define the triangularization;
#simplices is a numpy array of shape (no_triangles, 3)
#insert here the type check for input data
points3D=np.vstack((x,y,z)).T
tri_vertices=map(lambda index: points3D[index], simplices)# vertices of the surface triangles
zmean=[np.mean(tri[:,2]) for tri in tri_vertices ]# mean values of z-coordinates of
#triangle vertices
min_zmean=np.min(zmean)
max_zmean=np.max(zmean)
facecolor=[map_z2color(zz, colormap, min_zmean, max_zmean) for zz in zmean]
I,J,K=tri_indices(simplices)
triangles=go.Mesh3d(x=x,
y=y,
z=z,
facecolor=facecolor,
i=I,
j=J,
k=K,
name=''
)
if plot_edges is None:# the triangle sides are not plotted
return [triangles]
else:
#define the lists Xe, Ye, Ze, of x, y, resp z coordinates of edge end points for each triangle
#None separates data corresponding to two consecutive triangles
lists_coord=[[[T[k%3][c] for k in range(4)]+[ None] for T in tri_vertices] for c in range(3)]
Xe, Ye, Ze=[reduce(lambda x,y: x+y, lists_coord[k]) for k in range(3)]
#define the lines to be plotted
lines=go.Scatter3d(x=Xe,
y=Ye,
z=Ze,
mode='lines',
line=dict(color= 'rgb(50,50,50)', width=1.5)
)
return [triangles, lines]
```
Call this function for data associated to Moebius band:
```
data1=plotly_trisurf(x,y,z, tri.simplices, colormap=cm.RdBu, plot_edges=True)
```
Set the layout of the plot:
```
axis = dict(
showbackground=True,
backgroundcolor="rgb(230, 230,230)",
gridcolor="rgb(255, 255, 255)",
zerolinecolor="rgb(255, 255, 255)",
)
layout = go.Layout(
title='Moebius band triangulation',
width=800,
height=800,
scene=dict(
xaxis=dict(axis),
yaxis=dict(axis),
zaxis=dict(axis),
aspectratio=dict(
x=1,
y=1,
z=0.5
),
)
)
fig1 = go.Figure(data=data1, layout=layout)
py.iplot(fig1, filename='Moebius-band-trisurf')
```
### Triangularization of the surface $z=\sin(-xy)$, defined over a disk ###
We consider polar coordinates on the disk, $D(0, 1)$, centered at origin and of radius 1, and define
a meshgrid on the set of points $(r, \theta)$, with $r\in[0,1]$ and $\theta\in[0,2\pi]$:
```
n=12 # number of radii
h=1.0/(n-1)
r = np.linspace(h, 1.0, n)
theta= np.linspace(0, 2*np.pi, 36)
r,theta=np.meshgrid(r,theta)
r=r.flatten()
theta=theta.flatten()
#Convert polar coordinates to cartesian coordinates (x,y)
x=r*np.cos(theta)
y=r*np.sin(theta)
x=np.append(x, 0)# a trick to include the center of the disk in the set of points. It was avoided
# initially when we defined r=np.linspace(h, 1.0, n)
y=np.append(y,0)
z = np.sin(-x*y)
points2D=np.vstack([x,y]).T
tri=Delaunay(points2D)
```
Plot the surface with a modified layout:
```
data2=plotly_trisurf(x,y,z, tri.simplices, colormap=cm.cubehelix, plot_edges=None)
fig2 = go.Figure(data=data2, layout=layout)
fig2['layout'].update(dict(title='Triangulated surface',
scene=dict(camera=dict(eye=dict(x=1.75,
y=-0.7,
z= 0.75)
)
)))
py.iplot(fig2, filename='trisurf-cubehx')
```
This example is also given as a demo for matplotlib [`plot_trisurf`](http://matplotlib.org/examples/mplot3d/trisurf3d_demo.html).
### Plotting tri-surfaces from data stored in ply-files ###
A PLY (Polygon File Format or Stanford Triangle Format) format is a format for storing graphical objects
that are represented by a triangulation of an object, resulted usually from scanning that object. A Ply file contains the coordinates of vertices, the codes for faces (triangles) and other elements, as well as the color for faces or the normal direction to faces.
In the following we show how we can read a ply file via the Python package, `plyfile`. This package can be installed with `pip`.
We choose a ply file from a list provided [here](http://people.sc.fsu.edu/~jburkardt/data/ply/ply.html).
```
!pip install plyfile
from plyfile import PlyData, PlyElement
import urllib2
req = urllib2.Request('http://people.sc.fsu.edu/~jburkardt/data/ply/chopper.ply')
opener = urllib2.build_opener()
f = opener.open(req)
plydata = PlyData.read(f)
```
Read the file header:
```
for element in plydata.elements:
print element
nr_points=plydata.elements[0].count
nr_faces=plydata.elements[1].count
```
Read the vertex coordinates:
```
points=np.array([plydata['vertex'][k] for k in range(nr_points)])
points[0]
x,y,z=zip(*points)
faces=[plydata['face'][k][0] for k in range(nr_faces)]
faces[0]
```
Now we can get data for a Plotly plot of the graphical object read from the ply file:
```
data3=plotly_trisurf(x,y,z, faces, colormap=cm.RdBu, plot_edges=None)
title="Trisurf from a PLY file<br>"+\
"Data Source:<a href='http://people.sc.fsu.edu/~jburkardt/data/ply/airplane.ply'> [1]</a>"
noaxis=dict(showbackground=False,
showline=False,
zeroline=False,
showgrid=False,
showticklabels=False,
title=''
)
fig3 = go.Figure(data=data3, layout=layout)
fig3['layout'].update(dict(title=title,
width=1000,
height=1000,
scene=dict(xaxis=noaxis,
yaxis=noaxis,
zaxis=noaxis,
aspectratio=dict(x=1, y=1, z=0.4),
camera=dict(eye=dict(x=1.25, y=1.25, z= 1.25)
)
)
))
py.iplot(fig3, filename='Chopper-Ply-cls')
```
This a version of the same object plotted along with triangle edges:
```
from IPython.display import HTML
HTML('<iframe src=https://plot.ly/~empet/13734/trisurf-from-a-ply-file-data-source-1/ \
width=800 height=800></iframe>')
```
#### Reference
See https://plot.ly/python/reference/ for more information and chart attribute options!
```
from IPython.display import display, HTML
display(HTML('<link href="//fonts.googleapis.com/css?family=Open+Sans:600,400,300,200|Inconsolata|Ubuntu+Mono:400,700" rel="stylesheet" type="text/css" />'))
display(HTML('<link rel="stylesheet" type="text/css" href="http://help.plot.ly/documentation/all_static/css/ipython-notebook-custom.css">'))
! pip install git+https://github.com/plotly/publisher.git --upgrade
import publisher
publisher.publish(
'triangulation.ipynb', 'python/surface-triangulation/', 'Surface Triangulation',
'How to make Tri-Surf plots in Python with Plotly.',
title = 'Python Surface Triangulation | plotly',
name = 'Surface Triangulation',
has_thumbnail='true', thumbnail='thumbnail/trisurf.jpg',
language='python',
display_as='3d_charts', order=11,
ipynb= '~notebook_demo/71')
```
| github_jupyter |
# Dictionaries
Dictionaries are another type of Python container. Instead of storing values by index, they store them associated with a key.
You create dictionaries using curly brackets, assiging values to their keys using a colon, e.g.
```python
a = { "cat" : "mieow", "dog" : "woof", "horse" : "neigh" }
```
```
a = { "cat" : "mieow", "dog" : "woof", "horse" : "neigh"}
a
```
You can look up values in the dictionary by placing the key in square brackets. For example, we can look up the value associated with the key "cat" using `a["cat"]`.
```
a["cat"]
```
What happens if the key does not exist?
```
a['fish']
```
You insert items into the dictionary by assigning values to keys, e.g.
```
a["fish"] = "bubble"
a
```
You can list all of the keys or values of a dictionary using the `keys` or `values` functions (which you can find using tab completion and Python help)
```
help(a.values)
a.keys()
a.values()
```
You can loop over the dictionary by looping over the keys and looking up the values in a for loop, e.g.
```
for key in a.keys():
print("A %s goes %s" % (key, a[key]))
```
You can put anything as a value into a dictionary, including other dictionaries and even lists. The keys should be either numbers or strings.
```
b = { "a" : ["aardvark", "anteater", "antelope"], "b" : ["badger", "beetle"], 26.5: a}
```
What do you think is at `b["a"][-1]`? What about `b[26.5]["fish"]`?
```
b[26.5]["fish"]
```
# Exercise
Below you have a dictionary that contains the full mapping of every letter to its Morse-code equivalent.
```
letter_to_morse = {'a':'.-', 'b':'-...', 'c':'-.-.', 'd':'-..', 'e':'.', 'f':'..-.',
'g':'--.', 'h':'....', 'i':'..', 'j':'.---', 'k':'-.-', 'l':'.-..', 'm':'--',
'n':'-.', 'o':'---', 'p':'.--.', 'q':'--.-', 'r':'.-.', 's':'...', 't':'-',
'u':'..-', 'v':'...-', 'w':'.--', 'x':'-..-', 'y':'-.--', 'z':'--..',
'0':'-----', '1':'.----', '2':'..---', '3':'...--', '4':'....-',
'5':'.....', '6':'-....', '7':'--...', '8':'---..', '9':'----.',
' ':'/' }
```
## Exercise 1
Use the morse code dictionary to look up the morse code for the letters "s" and "o". What is the morse code for "SOS" (the international emergency distress signal)?
```
print(letter_to_morse["s"], letter_to_morse["o"], letter_to_morse["s"])
```
## Exercise 2
Here is a string that contains a message that must be converted to morse code. Write a loop that converts each letter into the morse code equivalent, and stores it into a list. Print the list out to see the full morse code message that must be sent. Note that you will need to use the `.lower()` function to get the lower case of capital letters.
```
message = "SOS We have hit an iceberg and need help quickly"
morse = []
for letter in message:
morse.append( letter_to_morse[letter.lower()] )
print(morse)
```
## Exercise 3
The inverted form of a dictionary is one where the keys are now looked up by value. For example, the below code inverts `letter_to_morse` such that the morse code is the key, and the letter is the value.
```
morse_to_letter = {}
for letter in letter_to_morse.keys():
morse_to_letter[ letter_to_morse[letter] ] = letter
```
Check that this code works by verifying that `morse_to_letter["..."]` equals "s".
Next, loop through the morse code message you created in exercise 2 and see if you can convert it back to english. Note that you can join a list of letters together into a string using the code `"".join(letters)`.
```
english = []
for code in morse:
english.append( morse_to_letter[code] )
print("".join(english))
```
| github_jupyter |

# Quickstart: Fraud Classification using Automated ML
In this quickstart, you use automated machine learning in Azure Machine Learning service to train a classification model on an associated fraud credit card dataset. This process accepts training data and configuration settings, and automatically iterates through combinations of different feature normalization/standardization methods, models, and hyperparameter settings to arrive at the best model.
You will learn how to:
> * Download a dataset and look at the data
> * Train a machine learning classification model using autoML
> * Explore the results
### Connect to your workspace and create an experiment
You start with importing some libraries and creating an experiment to track the runs in your workspace. A workspace can have multiple experiments, and all the users that have access to the workspace can collaborate on them.
```
import logging
from matplotlib import pyplot as plt
import pandas as pd
import numpy as np
import azureml.core
from azureml.core.experiment import Experiment
from azureml.core.workspace import Workspace
from azureml.core.dataset import Dataset
from azureml.train.automl import AutoMLConfig
ws = Workspace.from_config()
# choose a name for your experiment
experiment_name = "fraud-classification-automl-tutorial"
experiment = Experiment(ws, experiment_name)
```
### Load Data
Load the credit card dataset from a csv file containing both training features and labels. The features are inputs to the model, while the training labels represent the expected output of the model. Next, we'll split the data using random_split and extract the training data for the model.
Follow this [how-to](https://aka.ms/azureml/howto/createdatasets) if you want to learn more about Datasets and how to use them.
```
data = "https://automlsamplenotebookdata.blob.core.windows.net/automl-sample-notebook-data/creditcard.csv"
dataset = Dataset.Tabular.from_delimited_files(data)
training_data, validation_data = dataset.random_split(percentage=0.8, seed=223)
label_column_name = "Class"
```
## Train
When you use automated machine learning in Azure ML, you input training data and configuration settings, and the process automatically iterates through combinations of different feature normalization/standardization methods, models, and hyperparameter settings to arrive at the best model.
Learn more about how you configure automated ML [here](https://docs.microsoft.com/azure/machine-learning/how-to-configure-auto-train).
Instantiate an [AutoMLConfig](https://docs.microsoft.com/python/api/azureml-train-automl-client/azureml.train.automl.automlconfig.automlconfig?view=azure-ml-py) object. This defines the settings and data used to run the experiment.
|Property|Description|
|-|-|
|**task**|classification or regression|
|**primary_metric**|This is the metric that you want to optimize.
|**enable_early_stopping** | Stop the run if the metric score is not showing improvement.|
|**n_cross_validations**|Number of cross validation splits.|
|**training_data**|Input dataset, containing both features and label column.|
|**label_column_name**|The name of the label column.|
You can find more information about primary metrics [here](https://docs.microsoft.com/azure/machine-learning/service/how-to-configure-auto-train#primary-metric)
```
automl_settings = {
"n_cross_validations": 3,
"primary_metric": "average_precision_score_weighted",
"experiment_timeout_hours": 0.25, # This is a time limit for testing purposes, remove it for real use cases, this will drastically limit ability to find the best model possible
"verbosity": logging.INFO,
"enable_stack_ensemble": False,
}
automl_config = AutoMLConfig(
task="classification",
debug_log="automl_errors.log",
training_data=training_data,
label_column_name=label_column_name,
**automl_settings,
)
```
Call the `submit` method on the experiment object and pass the run configuration.
**Note: Depending on the data and the number of iterations an AutoML run can take a while to complete.**
In this example, we specify `show_output = True` to print currently running iterations to the console. It is also possible to navigate to the experiment through the **Experiment** activity tab in the left menu, and monitor the run status from there.
```
local_run = experiment.submit(automl_config, show_output=True)
local_run
```
### Analyze results
Below we select the best model from our iterations. The `get_output` method on `automl_classifier` returns the best run and the model for the run.
```
best_run, best_model = local_run.get_output()
best_model
```
## Tests
Now that the model is trained, split the data in the same way the data was split for training (The difference here is the data is being split locally) and then run the test data through the trained model to get the predicted values.
```
# convert the test data to dataframe
X_test_df = validation_data.drop_columns(
columns=[label_column_name]
).to_pandas_dataframe()
y_test_df = validation_data.keep_columns(
columns=[label_column_name], validate=True
).to_pandas_dataframe()
# call the predict functions on the model
y_pred = best_model.predict(X_test_df)
y_pred
```
### Calculate metrics for the prediction
Now visualize the data to show what our truth (actual) values are compared to the predicted values
from the trained model that was returned.
```
from sklearn.metrics import confusion_matrix
import numpy as np
import itertools
cf = confusion_matrix(y_test_df.values, y_pred)
plt.imshow(cf, cmap=plt.cm.Blues, interpolation="nearest")
plt.colorbar()
plt.title("Confusion Matrix")
plt.xlabel("Predicted")
plt.ylabel("Actual")
class_labels = ["False", "True"]
tick_marks = np.arange(len(class_labels))
plt.xticks(tick_marks, class_labels)
plt.yticks([-0.5, 0, 1, 1.5], ["", "False", "True", ""])
# plotting text value inside cells
thresh = cf.max() / 2.0
for i, j in itertools.product(range(cf.shape[0]), range(cf.shape[1])):
plt.text(
j,
i,
format(cf[i, j], "d"),
horizontalalignment="center",
color="white" if cf[i, j] > thresh else "black",
)
plt.show()
```
## Control cost and further exploration
If you want to control cost you can stop the compute instance this notebook is running on by clicking the "Stop compute" button next to the status dropdown in the menu above.
If you want to run more notebook samples, you can click on **Sample Notebooks** next to the **Files** view and explore the notebooks made available for you there.
| github_jupyter |
<div style="float: right; color:red; font-weight:bold;">Rename this file before you work on it!</div>
# Pandas Exercise
Use the Adventure Works dataset to create the following reports. The dataset is availablt for download in it's original format at https://msdn.microsoft.com/en-us/library/hh403424.aspx (follow instructions to download).
## Task
1. write the Python Pandas expression to produce a table as described in the problem statements.
2. The SQL expression may give you a hint. It also allows you to see both systems side-by-side.
3. If you don't know SQL just ignore the SQL code.
```
import pandas as pd
import numpy as np
%%time
Empoyees = pd.read_excel('/home/data/AdventureWorks/Employees.xls')
%%time
Territory = pd.read_excel('/home/data/AdventureWorks/SalesTerritory.xls')
%%time
Customers = pd.read_excel('/home/data/AdventureWorks/Customers.xls')
%%time
Orders = pd.read_excel('/home/data/AdventureWorks/ItemsOrdered.xls')
```
# Filtering (with)
### 1. Provide a list of employees that are married.
### 2a. Show me a list of employees that have a lastname that begins with "R".
### 2b. Show me a list of employees that have a lastname that ends with "r"
### 2c. Provide a list of employees that have a hyphenated lastname.
### 3a. Provide a list of employees that are on salary and have more than 35 vacation hours left.
### 3b. Show the same as above but limit it to American employees.
### 3c. Show the same as above but limit it to non-American employees.
### 4a. List the married employees with more than 35 vacation hours, only ones living in Washington state.
### 4b. Change the logic to include anyone who meets any of the 3 conditions (i.e., people who are either married, live in Washington state, or have more than 35 vacation hours left)
### 4c. Show the same as above, but only for Production Technicians
### 5a. List all employees living in Redmond, Seattle, and Bellevue, showing EmployeeID, FirstName, LastName, and City. Sort the list alphabetically by city.
### 5b. For the list above, make sure these are only in the state of Washington, just to be careful.
### 6. Provide a list of employees who have no title, whether it's a NULL or empty string.
### 7a. Provide a list of employees who have at least 60 vacation hours left.
### 7b. Provide a list of employees who have less than 60 vacation hours left.
```
SELECT e.EmployeeID, e.Title, e.FirstName, e.LastName, e.VacationHours, e.SickLeaveHours
FROM dbo.Employees AS e
WHERE e.VacationHours < 60
;
```
### 7c. Show me employees who have more than 20 and less than 60 vacation hours left.
### 7d. If you did not use BETWEEN for 7c, do the same but use BETWEEN. If you did use BETWEEN for 7c, do it another way.
# Grouping
### 1a. What is the earliest birthdate for all employees?
### 1b. Add to the above, the most recent birthdate for all employees
### 1c. Show the above results broken down by gender
### 1d. Show the above results broken down by gender, and salaried/hourly
### 2a. What are the average vacation hours for all employees?
### 2b. Add to the above, the minimum vacation hours for all employees
### 2c. Show the above results broken down and ordered by job title
### 2d. Show the above results broken down by job title, and married/single employees
### 2e. Add to the above, the maximum vacation hours per group
### 2f. Show the above results broken down by job title, married/single employees, and State
### 2g. Show the above results but only for American employees
### 2h. Change the grouping above so it's broken down by married/single and State, no more job title
### 2i. Limit the results above to States where the average vacation hours is greater than 30
### 2j. Limit the results above to States where the average vacation hours is greater than 30 and the maximum vacation hours is less than 50
### 2k. Show the same results but only for non-American employees
### 3a. Report how many employees are in the company
### 3b. For the above report, show the number of employees per manager (hint: use ManagerID)
### 3c. Remove any manager ID's that are NULL from the results above
### 3d. Show the same results as above, but only for managers who have at least 5 employees
### 4a. List the average vacation hours of all employees
### 4b. Break down the results by State
### 4c. Break down the results by city and State
### 4d. Add something that shows the number of employees per city
### 4e. Sort the results by the city and state
### 4f. Make city and State a single column in the format of "City, State"
### 4g. Add a column that shows the difference between the maximum vacation hours and minimum vacation hours for each city
### 4h. Now sort the results by the new column created above
### 4i. Limit the results to cities that have more than 1 employee
### 4j. Limit the results to non-U.S. cities
| github_jupyter |
# INFO 3350/6350
## Lecture 07: Vectorization, distance metrics, and regression
## To do
* Read HDA ch. 5 and Grimmer and Stewart for Monday (a lot of reading)
* HW3 (gender and sentiment; dictionary methods) due by Thursday night at 11:59.
* Extra credit for good, consistent answers on Ed
* Study groups are great for homeworks.
* Questions?
## Definitions
* What is a **vector**?
* An ordered collection of numbers that locate a point in space relative to a shared reference point (called the *origin*).
* We can also think of vectors as representing the quantified *features* of an object.
* Vectors are usually written as *row matrices*, or just as lists: $vec = [1.0, 0.5, 3.0, 1.2]$
* Vectors have as many *dimensions* as there are features of the object to represent.
* The number of features to represent is a choice of the experiment. There is no correct choice, though some choices are better than others for a given purpose.
* What is **vectorization**?
* The process of transforming an object into its vector representation, typically by measuring some of the object's properties.
## Why would we want to do this?
One goal of humanistic inquiry and of scientific research is to compare objects, so that we can gather them into types and compare any one object to others that we observe. Think of biological species or literary genres or historical eras. But how can we measure the difference or similarity between objects that are, after all, always necessarily individual and unique?
* Measuring the *properties* of objects lets us compare those objects to one another.
* But ... *which* properties?
* Example: We counted words by type to compare gender and sentiment in novels.
* Establishing a vector representation allows us to define a **distance metric** between objects that aren't straightforwardly spatial.
* "Distance" is a metaphor. Ditto "similarity."
* Nothing is, in itself, like or unlike anything else.
* We sometimes seek to assert that objects are similar by erasing aspects of their particularity.
* Measuring similarity and difference are (always and only) interpretive interventions.
## A spatial example
Consider this map of central campus:

**How far apart are Gates Hall (purple star) and the clock tower (orange star)?**
What do we need to know or define in order to answer this question?
* Where is each building in physical space.
* Latitude/longitude; meters north/south and east/west of the book store; etc.
* How do we want to measure the distance between them (walking, driving, flying, tunneling, ...). Minutes or miles?
Normal, boring answer: about 0.4 miles on foot via Campus Rd and Ho Plaza, or a bit less if you cut some corners, or less than 0.3 miles if you can fly.
| Clock tower | Gates Hall |
| --- | --- |
|  |  |
More interesting version: How far apart are these buildings conceptually? Architecturally? Historically?
* What are the features and metrics you would use to answer this question?
* This is a lot more like the problem of comparing texts.
## A textual example
```
text = '''\
My cat likes water.
The dog eats food.
The dog and the cat play together.
A dog and a cat meet another dog and cat.
The end.'''
# Print with sentence numbers
for line in enumerate(text.split('\n')):
print(line)
```
Let us stipulate that we want to compare these five sentences according to their "*dogness*" and "*catness*." We care about those two aspects alone, nothing else.
Let's develop some intuitions here:
* Sentences 0 and 1 are as far apart as can be: 0 is about cats, 1 is about dogs.
* Sentence 2 lies between 0 and 1. It contains a mix of dogness and catness.
* Sentence 3 is kind of like sentence 2, but it has twice as much of both dogness and catness.
* How different are sentences 2 and 3? (There's no objectively correct answer.)
* Sentence 4 is a zero point. It has no dogness or catness.
### Count relevant words
||**cat**|**dog**|
|---|---|---|
|**sent**| | |
|0|1|0|
|1|0|1|
|2|1|1|
|3|2|2|
|4|0|0|
The **vector representation** of sentence 0 is `[1, 0]`. The vector representation of sentence 3 is `[2, 2]`. And so on ...
### Visualize (scatter plot)
Sketch this by hand ...
### Distance measures
How far apart are sentences 0 and 1 (and all the rest)?
#### Manhattan distance
* Also called "city block" distance.
* Not much used, but easy to understand and to compute (which matters for very large data sets).
* Sum of the absolute difference in each dimension.
For **sentences 0 and 1**, the Manhattan distance = |1| + |-1| = 2.
#### Euclidean distance
* Straight-line or "as the crow flies" distance.
* Widely used in data science, but not always the best choice for textual data.
Recall the Pythagorean theorem for the hypotenuse of a triangle: $a^2 = b^2 + c^2$ or $a = \sqrt{b^2 +c^2}$.
For **sentences 0 and 1**, the Euclidean distance = $\sqrt{1^2 + 1^2} = \sqrt{2} = 1.414$.
OK, but what about the Euclidean distance between **sentence 0 and sentence 3**? Well, that distance = $\sqrt{1^2 + 2^2} = \sqrt{5} = 2.24$.
And between **sentences 2 and 3** (both balanced 50:50 between dogs and cats)? That's 1.4 again, the same as the distance between sentences 0 and 1 (which, recall, are totally divergent in dog/cat content).
An obvious improvement in this case would be to **normalize word counts by document length**.
#### Cosine distance
Maybe instead of distance, we could measure the difference in **direction** from the origin between points.
* **Sentences 0 and 1** are 90 degrees apart.
* **Sentences 2 and 3** are 0 degrees apart.
* **Sentences 0 and 1** are each 45 degrees away from **sentences 2 and 3**.
Now, recall the values of the **cosine** of an angle between 0 and 90 degrees. (Sketch by hand)
So, the cosines of the angles between sentences are:
sentences|angle|cosine
---|---|---
0 and 1|90|0
2 and 3|0|1
0 and 2|45|0.707
0 and 3|45|0.707
1 and 2|45|0.707
We could then transform these cosine **similarities** into **distances** by subtracting them from 1, so that the most *dissimilar* sentences (like 0 and 1) have the greatest distance between them.
The big advantage here is that we don't need to worry about getting length normalization right. Cosine distance is often a good choice for text similarity tasks.
#### Higher dimensions
All of these metrics can be calculated in arbitrarily many dimensions. Which is good, because textual data is often very high-dimensional. Imagine counting the occurrences of each word type in a large corpus of novels or historical documents. Can easily be tens of thousands of dimensions.
## In the real world
* There's nothing wrong with any of these vectorizations and distance metrics, exactly, but they're not state of the art.
* If you've done some recent NLP work, you'll know that, at the very least, you'd want to use static word embeddings in place of raw tokens.
* This allows you to capture the similarity of meaning between, e.g., "cat" and "kitten."
* If you were especially ambitious, you'd be looking at something like BERT or ELMo or GPT-2/3, etc.
* These transformer-based methods allow for *contextual* embeddings, that is, they represent a word token differently depending on the context in which it appears, so that the representation of "bank" in "my money is in the bank" is different from the the representation of "bank" in "we walked along the bank of the river."
* We'll touch on contextual embeddings near the end of the semester.
* And then you might want features that correspond to aspects of a text other than the specific words it contains.
* When was it written?
* By *whom* was it written?
* How long is it?
* In what style is it written?
* Who read it?
* How much did it cost?
* How many people read or reviewed it?
* What else did its readers also read?
* And so on ...
Here, though, we're trying to grasp the *idea* behind document similarity, on which all of these methods depend: transform text into a numeric representation of its features (often, a representation of its content or meaning), then quantify the difference or similarity between those numeric representations.
## In the problem set world
We'll dig into how, as a practical matter, we can vectorize texts and calclulate distance metrics in this week's problem set.
We'll use `scikit-learn` to implement vectorization and distance metrics. The `scikit-learn` API almost always involves *three* steps:
1. Instantiate a learning object (such as a vectorizer, regressor, classifier, etc.). This is the object that will hold the parameters of your fitted model.
1. Call the instantiated learning object's `.fit()` method, passing in your data. This allows the model to learn the optimal parameters from your data.
1. Call the fitted model's `.transform()` or `.predict()` method, passing in either the same data from the `fit` step or new data. This step uses the fitted model to generate outputs given the input data you supply.
For example:
```
from sklearn.feature_extraction.text import CountVectorizer
# get example text as one doc per line
docs = [sent for sent in text.split('\n')]
# instantiate vectorizer object
# note setup options
vectorizer = CountVectorizer(
vocabulary=['cat', 'dog']
)
# fit to data
vectorizer.fit(docs)
# transform docs to features
features = vectorizer.transform(docs)
# print output feature matrix
print(vectorizer.get_feature_names_out())
print(features.toarray())
# calculate distances
from sklearn.metrics.pairwise import euclidean_distances, cosine_distances, cosine_similarity
import numpy as np
print("Euclidean distances")
print(np.round(euclidean_distances(features),2))
print("\nCosine distances")
print(np.round(cosine_distances(features),2))
print("\nCosine **similarities**")
print(np.round(cosine_similarity(features),2))
# FYI, a heatmap vis
import seaborn as sns
print("Euclidean distances")
sns.heatmap(
euclidean_distances(features),
annot=True,
square=True
);
```
## Regression
We are often interested in the relationships between measured properties of texts, or between a textual property and some other variable (year of publication, number of sales, and so on).
Maybe the most basic way to measure the relationship between two variables is to use **linear regression**. The idea is to calculate a straight line through your data such that the average distance between the observed data points and the line is as small as possible.
(Sketch what this looks like)
You can then calculate the **coefficient of determination**, written $r^2$ ("r squared"), which measures the fraction of the variation in the dependent (y) variable that is predictable from the independent (x) variable.
$r^2$ = 1 - (sum of squared residuals)/(sum of squared values)
An $r^2$ value of 1 indicates perfect correlation between the variables; zero means no correlation.
* There's a *lot* more to this. We'll spend some time on it later in the semester.
* For now, focus on the fact that regression is a way to calculate a line of best fit through a data set.
* Notice that we could also try to find something like a "line of *worst* fit," which we could think of as the dividing line between two regions of feature space. This would be something like the line on which we are least likely to encounter any actual data points.
* Think about what use-value such a dividing line might have ...
| github_jupyter |
```
import numpy as np
```
The OneR algorithm is quite simple but can be quite effective, showing the power of using even basic statistics in many applications.
The algorithm is:
* For each variable
* For each value of the variable
* The prediction based on this variable goes the most frequent class
* Compute the error of this prediction
* Sum the prediction errors for all values of the variable
* Use the variable with the lowest error
```
# Load our dataset
from sklearn.datasets import load_iris
#X, y = np.loadtxt("X_classification.txt"), np.loadtxt("y_classification.txt")
dataset = load_iris()
X = dataset.data
y = dataset.target
print(dataset.DESCR)
n_samples, n_features = X.shape
```
Our attributes are continuous, while we want categorical features to use OneR. We will perform a *preprocessing* step called discretisation. At this stage, we will perform a simple procedure: compute the mean and determine whether a value is above or below the mean.
```
# Compute the mean for each attribute
attribute_means = X.mean(axis=0)
assert attribute_means.shape == (n_features,)
X_d = np.array(X >= attribute_means, dtype='int')
# Now, we split into a training and test set
from sklearn.cross_validation import train_test_split
# Set the random state to the same number to get the same results as in the book
random_state = 14
X_train, X_test, y_train, y_test = train_test_split(X_d, y, random_state=random_state)
print("There are {} training samples".format(y_train.shape))
print("There are {} testing samples".format(y_test.shape))
from collections import defaultdict
from operator import itemgetter
def train(X, y_true, feature):
"""Computes the predictors and error for a given feature using the OneR algorithm
Parameters
----------
X: array [n_samples, n_features]
The two dimensional array that holds the dataset. Each row is a sample, each column
is a feature.
y_true: array [n_samples,]
The one dimensional array that holds the class values. Corresponds to X, such that
y_true[i] is the class value for sample X[i].
feature: int
An integer corresponding to the index of the variable we wish to test.
0 <= variable < n_features
Returns
-------
predictors: dictionary of tuples: (value, prediction)
For each item in the array, if the variable has a given value, make the given prediction.
error: float
The ratio of training data that this rule incorrectly predicts.
"""
# Check that variable is a valid number
n_samples, n_features = X.shape
assert 0 <= feature < n_features
# Get all of the unique values that this variable has
values = set(X[:,feature])
# Stores the predictors array that is returned
predictors = dict()
errors = []
for current_value in values:
most_frequent_class, error = train_feature_value(X, y_true, feature, current_value)
predictors[current_value] = most_frequent_class
errors.append(error)
# Compute the total error of using this feature to classify on
total_error = sum(errors)
return predictors, total_error
# Compute what our predictors say each sample is based on its value
#y_predicted = np.array([predictors[sample[feature]] for sample in X])
def train_feature_value(X, y_true, feature, value):
# Create a simple dictionary to count how frequency they give certain predictions
class_counts = defaultdict(int)
# Iterate through each sample and count the frequency of each class/value pair
for sample, y in zip(X, y_true):
if sample[feature] == value:
class_counts[y] += 1
# Now get the best one by sorting (highest first) and choosing the first item
sorted_class_counts = sorted(class_counts.items(), key=itemgetter(1), reverse=True)
most_frequent_class = sorted_class_counts[0][0]
# The error is the number of samples that do not classify as the most frequent class
# *and* have the feature value.
n_samples = X.shape[1]
error = sum([class_count for class_value, class_count in class_counts.items()
if class_value != most_frequent_class])
return most_frequent_class, error
# Compute all of the predictors
all_predictors = {variable: train(X_train, y_train, variable) for variable in range(X_train.shape[1])}
errors = {variable: error for variable, (mapping, error) in all_predictors.items()}
# Now choose the best and save that as "model"
# Sort by error
best_variable, best_error = sorted(errors.items(), key=itemgetter(1))[0]
print("The best model is based on variable {0} and has error {1:.2f}".format(best_variable, best_error))
# Choose the bset model
model = {'variable': best_variable,
'predictor': all_predictors[best_variable][0]}
print(model)
def predict(X_test, model):
variable = model['variable']
predictor = model['predictor']
y_predicted = np.array([predictor[int(sample[variable])] for sample in X_test])
return y_predicted
y_predicted = predict(X_test, model)
print(y_predicted)
# Compute the accuracy by taking the mean of the amounts that y_predicted is equal to y_test
accuracy = np.mean(y_predicted == y_test) * 100
print("The test accuracy is {:.1f}%".format(accuracy))
from sklearn.metrics import classification_report
print(classification_report(y_test, y_predicted))
```
| github_jupyter |
# Geocube Data Indexation Tutorial
-------
**Short description**
This notebook introduces you to the Geocube Python Client. You will learn how to index images in the Geocube.
-------
**Requirements**
-------
- Python 3.7
- The Geocube Python Client library : https://github.com/airbusgeo/geocube-client-python.git
- The Geocube Server & Client ApiKey (for the purpose of this notebook, GEOCUBE_SERVER and GEOCUBE_CLIENTAPIKEY environment variable)
-------
**Installation**
-------
`pip install --user git+https://github.com/airbusgeo/geocube-client-python.git`
## 1 - Connect to the Geocube
```
import glob
import math
import os
from datetime import datetime
import numpy as np
from matplotlib import pyplot as plt
%matplotlib inline
plt.rcParams['figure.figsize'] = [20, 16]
from geocube import Client, entities, utils
# Define the connection to the server
secure = False # in local, or true
geocube_client_server = os.environ.get('GEOCUBE_SERVER')
geocube_client_api_key = os.environ.get('GEOCUBE_CLIENTAPIKEY')
# Connect to the server
client = Client(geocube_client_server, secure, geocube_client_api_key)
```
## 2 - Indexation in a nutshell
In the Geocube, an image (a **dataset**) is indexed by a **record** and an **instance** of a **variable**.
These concepts will soon be defined in details, but in short, a record defines the data-take and the variable definies the kind of data.
<img src="GetImage.png" width=400>
Adding an image in the Geocube is a process called *indexation*.
```
print("It's very easy to add an image in the Geocube:")
try:
print("Create the AOI of the record")
aoi_id = client.create_aoi(utils.read_aoi('inputs/UTM32UNG.json'))
print("Create the record")
record = client.create_record(aoi_id, "MyFirstRecord", {"source":"tutorial"}, datetime.now())
print("Create the variable and instantiate it")
instance = client.create_variable("MyFirstVariable", "i2,-1,0,255", ['']).instantiate("AnInstance", {})
print("And finally, index the image")
client.index_dataset(os.getcwd()+'/inputs/myFirstImage.tif', record, instance, instance.dformat, bands=[1])
print("You did it !")
except utils.GeocubeError as e:
print(e)
print("But it only works once ! (I'm kidding... let's go to the next step !)")
```
## 3 - Records
A record defines a data-take by its geometry, its sensing time and user-defined tags that describe the context in more detail.
A record is usually linked to an image of a satellite. For example, the image taken by S2A over the 31TDL tile on the 1st of April 2018 is described by the record:
- **S2A_MSIL1C_20180401T105031_N0206_R051_T31TDL_20180401T144530**
* **AOI** : _31TDL tile (POLYGON ((2.6859855651855 45.680294036865, 2.811126708984324 45.680294036865, 2.811126708984324 45.59909820556617, 2.6859855651855 45.59909820556617, 2.6859855651855 45.680294036865)))_
* **DateTime** : _2018-04-01 10:50:31_
But a record can describe any product like a mosaic over a country, or a decade mosaic :
- **Mosaic of France January 2020**
* **AOI** : _France_
* **DateTime** : _2020-01-31 00:00:00_
<img src="RecordsSeveralLayers.png" width=400>
### Create an AOI
A record is linked to an AOI (that can be shared between several records). Before creating the record, its AOI must be created with the function `create_aoi` taking a geometry in **geographic coordinates** as input.
If the aoi already exists, `create_aoi` raises an error. Its ID can be retrieved from the details of the error.
```
aoi = utils.read_aoi('inputs/UTM32UNG.json')
try:
aoi_id = client.create_aoi(aoi)
print("AOI created with id "+aoi_id)
except utils.GeocubeError as e:
aoi_id = e.details[e.details.rindex(' ')+1:]
print("AOI already exists with id: "+aoi_id)
```
### Create a record
The `create_records` function is used to create new records. Records are uniquely defined by :
- `name`
- `tags` : user-defined depending on the project (currently, no standard are implemented).
- `datetime`
```
name = "S2B_MSIL1C_20190118T104359_N0207_R008_T32UNG_20190118T123528"
tags = {"source":"tutorial", "constellation":"SENTINEL2", "satellite":"SENTINEL2B", "user-defined-tag": "whatever is necessary to search for this record"}
date = datetime(2019, 1, 18, 10, 43, 59, 0, None)
try:
record_id = client.create_record(aoi_id, name, tags, date)
print("Record created with id: ", record_id)
except utils.GeocubeError as e:
record_id = client.list_records(name, tags=tags, from_time=date, to_time=date)[0].id
print(f'{e.codename} {e.details} id={record_id}')
```
## 4 - Variables
A variable describes the kind of data stored in a product, for example _a spectral band, the NDVI, an RGB image, the backscatter, the coherence_...
This entity has what is needed to **describe**, **process** and **visualize** the product.
In particular, the variable has a `dformat` (for _data format_ ):
- dformat.dtype : _data type_
- dformat.min : theoretical minimum value
- dformat.max : theoretical maximum value
- dformat.no_data : the NoData value
In the Geocube Database, the (internal) data format of an image indexed in the Geocube may be different (for exemple, in order to optimize storage costs), but when the data is retrieved, the Geocube maps the internal format to the data format of the variable. This process may map the data below the minimum or above the maximum value. In that case, no crop is performed.
### Create a variable
```
print("Create a variable that describes an RGB product")
variable_name = "RGB"
try:
variable = client.create_variable(
name=variable_name,
dformat={"dtype":"i2", "no_data": -1, "min_value": 0, "max_value": 255},
bands=['R', 'G', 'B'],
description="",
unit="",
resampling_alg=entities.Resampling.bilinear)
except utils.GeocubeError as e:
print(e.codename + " " + e.details)
variable = client.variable(name=variable_name)
print(variable)
print("Create a variable that describes an NDVI product")
variable_name = "NDVI"
try:
variable = client.create_variable(
name=variable_name,
dformat={"dtype":"f4", "no_data": np.nan, "min_value": -1, "max_value": 1},
bands=[''],
description="Normalized Difference Vegetation Index",
resampling_alg=entities.Resampling.bilinear)
except utils.GeocubeError as e:
print(e.codename + " " + e.details)
variable = client.variable(name=variable_name)
print(variable)
```
### Instantiate a variable
An instance is a declination of a variable with different processing parameters.
For example, an RGB variable can be defined with different spectral bands (RGB bands of Sentinel-2 are not the same as LANDSAT's), a Label variable can have a different mapping. The SAR products can be processed with different processing graphs or softwares, but they all belongs to the same variable.
The processing parameters can be provided in the metadata field of the instance.
`client.variable("RGB").instantiate("Sentinel2-Raw-Bands", {"R":"664.6", "G":...})`
`client.variable("LandUseLabels").instantiate("v1", {"0":"Undefined","1":"Urban", "2":...})`
`client.variable("Sigma0VV").instantiate("terrain-corrected", {"snap_graph_name":"mygraph.xml", ...})`
```
try:
instance=client.variable("RGB").instantiate("master", {"any-metadata": "(for information purpose)"})
except utils.GeocubeError as e:
print(e.codename + " " + e.details)
instance = variable.instance("master")
print(instance)
try:
instance=client.variable("NDVI").instantiate("master", {"any-metadata": "(for information purpose)"})
except utils.GeocubeError as e:
print(e.codename + " " + e.details)
instance = variable.instance("master")
print(instance)
```
## 5 - Dataset
As we saw in introduction, we have all we need to index an image in the Geocube. Such an image is called a **dataset**.
Actually, to index an image, we also have to define :
- which band(s) are indexed (usually all the bands, but it can be a subset)
- how to map the value of its pixels to the dataformat of the variable.
For the second point, we will define :
- the dataformat of the dataset (`dformat.[no_data, min, max]`) that describes the pixel of the image
- the mapping from each pixel to the data format of the variable (`variable.dformat`). This mapping is defined as `[MinOut, MaxOut, Exponent]`. See the diagram below:
NB:
- **`dataset.Min` and `dataset.max` are NOT necessarily the minimum and maximum values of the pixels but the minimum and maximum possible values.**
- `index_dataset()` **does not perform any transformation on the image** (all the information provided during the indexation is for the interpretation of the image - by the Geocube or the user) and is idempotent.
<a name="diagram"></a><img src="InternalDFormatToVariableDFormat.png" width=800>
### Index a dataset (common case)
The dataformat of the dataset is generally the same as the one of the variable.
```
# Define URI, record and variable.instance
uri = os.getcwd() + "/inputs/S2B_MSIL1C_20190118T104359_N0207_R008_T32UNG_20190118T123528.tif"
record = client.list_records("S2B_MSIL1C_20190118T104359_N0207_R008_T32UNG_20190118T123528")[0]
instance = client.variable(name="RGB").instance("master")
# In that case, the dformat of the dataset is the same as the one of the variable
dataset_dformat = instance.dformat
client.index_dataset(uri, record, instance, dataset_dformat, bands=[1,2,3])
print("Done !")
```
### Index a dataset (Storage optimisation)
In order to optimize the storage of a large volume of data, it can be decided to reduce the size of the data type (for example from float32 to int16) and/or scale the data.
So, the dataformat of the dataset can be different from the variable in some ways:
- **For compression purpose** :
1. the data type is smaller. For example data is encoded in byte [0, 255] that maps to float [0, 1] in the variable.
- **To optimize accuracy** : the range of values is smaller than the one of the variable. Two examples :
2. Given a variable between -1 and 1, the data in a given image is known to be in [0, 1] instead of [-1, 1]. To optimize accuracy, the data is encoded between 0 and 255 and min/max_out are [0, 1].
3. Given a variable between 0 and 100, 90% of the data is known to be between 0 and 10. To optimize accuracy, the data is encoded between 0 and 255, using a non-linear mapping to [0, 100] using an exponent=2. Data is scaled according to the non-linear scaling in the [diagram](#diagram):
<img src="DataFormatExample.png" width=800>
NB: below : dformat.dtype is retrieved from the file, hence the "auto" keyword.
```
print("NDVI variable is defined in the range [-1, 1]")
instance = client.variable(name="NDVI").instance("master")
print("Example 1: the datatype of the dataset has been encoded in int16, mapping [-10000, 10000] to [-1, 1]")
internal_dformat = {"dtype":"auto", "no_data": -10001, "min_value": -10000, "max_value": 10000}
try:
tags = {"source":"tutorial", "constellation":"SENTINEL2", "satellite":"SENTINEL2A"}
date = datetime(2019, 2, 24, 10, 30, 19, 0, None)
client.create_record(aoi_id, "S2A_MSIL1C_20190224T103019_N0207_R108_T32UNG_20190224T141253", tags, date)
except utils.GeocubeError:
pass
uri = os.getcwd() + "/inputs/ndviS2A_MSIL1C_20190224T103019_N0207_R108_T32UNG_20190224T141253.tif"
record = client.list_records("S2A_MSIL1C_20190224T103019_N0207_R108_T32UNG_20190224T141253")[0]
client.index_dataset(uri, record, instance, internal_dformat, bands=[1])
plt.imshow
print("Done !")
print("Example 2: this NDVI dataset is known to have no value below 0. Therefore, it has been encoded in uint8, mapping [0, 255] to [0, 1]")
internal_dformat = {"dtype":"auto", "no_data": 0, "min_value": 0, "max_value": 255}
min_out, max_out = 0, 1
uri = os.getcwd() + "/inputs/ndviS2B_MSIL1C_20190118T104359_N0207_R008_T32UNG_20190118T123528.tif"
record = client.list_records("S2B_MSIL1C_20190118T104359_N0207_R008_T32UNG_20190118T123528")[0]
client.index_dataset(uri, record, instance, internal_dformat, bands=[1], min_out=min_out, max_out=max_out)
print("Done !")
print("Example 3: this NDVI dataset has most of its value in [0, 0.1]. Therefore, it has been encoded in uint8, mapping [0, 255] to [0, 1] with an exponent=2")
internal_dformat = {"dtype":"auto", "no_data": 0, "min_value": 0, "max_value": 255}
min_out, max_out, exponent = 0, 1, 2
uri = os.getcwd() + "/inputs/ndviS2A_MSIL1C_20190224T103019_N0207_R108_T32UNG_20190224T141253_2.tif"
record = client.list_records("S2A_MSIL1C_20190224T103019_N0207_R108_T32UNG_20190224T141253")[0]
client.index_dataset(uri, record, instance, internal_dformat, bands=[1], min_out=min_out, max_out=max_out, exponent=2)
print("Done !")
```
## 6 - Index a list of datasets
```
filepaths = list(glob.glob(os.getcwd() + "/inputs/S2B_MSIL1C*.tif"))
print("Create all records")
records_name = []
records_tags = []
records_date = []
records_aoi = []
for filepath in filepaths:
# This record already exists
if 'S2B_MSIL1C_20190118T104359_N0207_R008_T32UNG_20190118T123528' in filepath:
continue
record_name = os.path.basename(filepath).strip(".tif")
records_name.append(record_name)
records_date.append(datetime(int(record_name[11:15]), int(record_name[15:17]), int(record_name[17:19]), int(record_name[20:22]), int(record_name[22:24]), int(record_name[24:26]), 0))
records_aoi.append(aoi_id)
records_tags.append({"source":"tutorial", "constellation":"SENTINEL2", "satellite":"SENTINEL2B"})
try:
record_ids = client.create_records(records_aoi, records_name, records_tags, records_date)
print(f"{len(record_ids)} records added")
except utils.GeocubeError as e:
print(e.codename + " " + e.details)
print("Index all datasets")
records = client.list_records(tags={"source":"tutorial", "constellation":"SENTINEL2", "satellite":"SENTINEL2B"})
record_map = {record.name: record.id for record in records}
instance = client.variable(name="RGB").instance("master")
containers = []
for filepath in filepaths:
# Find the record
record_id = record_map[os.path.basename(filepath).strip(".tif")]
# Create the container (dformat is the one of the variable)
containers.append(entities.Container.new(filepath, record_id,
instance = instance.instance_id,
bands=[1, 2, 3],
dformat=instance.dformat,
min_out=0, max_out=255))
client.index(containers)
print("Done !")
```
## 7 - Conclusion
In this notebook, you have learnt to create aois, records and variables, instantiate a variable and index a dataset.
To populate the Geocube for the next tutorial, we will index some additional datasets :
```
instance = client.variable("NDVI").instance("master")
try:
aoi_id = client.create_aoi(utils.read_aoi('inputs/UTM32VNH.json'))
record = client.create_record(aoi_id, "S2B_MSIL1C_20190105T103429_N0207_R108_T32VNH_20190105T122413", {"source":"tutorial", "constellation":"SENTINEL2", "satellite":"SENTINEL2B"}, datetime(2019,1,5,10,34,29))
client.index_dataset(os.getcwd()+'/inputs/ndviS2B_MSIL1C_20190105T103429_N0207_R108_T32VNH_20190105T122413.tif', record, instance, "auto,-1001,-1,1", bands=[1])
except utils.GeocubeError as e:
print("It seems that you already did it !")
try:
aoi_id = client.create_aoi(utils.read_aoi('inputs/UTM32VNJ.json'))
record = client.create_record(aoi_id, "S2B_MSIL1C_20190105T103429_N0207_R108_T32VNJ_20190105T122413", {"source":"tutorial", "constellation":"SENTINEL2", "satellite":"SENTINEL2B"}, datetime(2019,1,5,10,34,29))
client.index_dataset(os.getcwd()+'/inputs/ndviS2B_MSIL1C_20190105T103429_N0207_R108_T32VNJ_20190105T122413.tif', record, instance, "auto,-1001,-1,1", bands=[1])
except utils.GeocubeError as e:
print("It seems that you already did it !")
```
| github_jupyter |
# Pedestrian Tracking with YOLOv3 and DeepSORT
This is a pedestrian tracking demo using the open source project [ZQPei/deep_sort_pytorch](https://github.com/ZQPei/deep_sort_pytorch) which combines [DeepSORT](https://github.com/nwojke/deep_sort) with [YOLOv3](https://pjreddie.com/darknet/yolo/).
For other deep-learning Colab notebooks, visit [tugstugi/dl-colab-notebooks](https://github.com/tugstugi/dl-colab-notebooks).
## Install ZQPei/deep_sort_pytorch
```
import os
from os.path import exists, join, basename
project_name = "deep_sort_pytorch"
if not exists(project_name):
# clone and install
!git clone -q --recursive https://github.com/ZQPei/deep_sort_pytorch.git
import sys
sys.path.append(project_name)
import IPython
from IPython.display import clear_output
```
## Download pretrained weights
```
yolo_pretrained_weight_dir = join(project_name, 'detector/YOLOv3/weight/')
if not exists(join(yolo_pretrained_weight_dir, 'yolov3.weights')):
!cd {yolo_pretrained_weight_dir} && wget -q https://pjreddie.com/media/files/yolov3.weights
deepsort_pretrained_weight_dir = join(project_name, 'deep_sort/deep/checkpoint')
if not exists(join(deepsort_pretrained_weight_dir, 'ckpt.t7')):
file_id = '1_qwTWdzT9dWNudpusgKavj_4elGgbkUN'
!cd {deepsort_pretrained_weight_dir} && curl -Lb ./cookie "https://drive.google.com/uc?export=download&id={file_id}" -o ckpt.t7
```
## Track pedestrians
First, download a test video from internet and show in the notebook:
```
VIDEO_URL = 'http://www.robots.ox.ac.uk/ActiveVision/Research/Projects/2009bbenfold_headpose/Datasets/TownCentreXVID.avi'
DURATION_S = 20 # process only the first 20 seconds
video_file_name = 'video.mp4'
if not exists(video_file_name):
!wget -q $VIDEO_URL
dowloaded_file_name = basename(VIDEO_URL)
# convert to MP4, because we can show only MP4 videos in the colab noteook
!ffmpeg -y -loglevel info -t $DURATION_S -i $dowloaded_file_name $video_file_name
def show_local_mp4_video(file_name, width=640, height=480):
import io
import base64
from IPython.display import HTML
video_encoded = base64.b64encode(io.open(file_name, 'rb').read())
return HTML(data='''<video width="{0}" height="{1}" alt="test" controls>
<source src="data:video/mp4;base64,{2}" type="video/mp4" />
</video>'''.format(width, height, video_encoded.decode('ascii')))
clear_output()
show_local_mp4_video('video.mp4')
```
Now, track the pedestrians on the downloaded video:
```
!cd {project_name} && python yolov3_deepsort.py --ignore_display ../video.mp4 --save_path ../output.avi
```
Finally, we can visualize the result:
```
# first convert to mp4 to show in a Colab notebook
!ffmpeg -y -loglevel panic -i output.avi output.mp4
show_local_mp4_video('output.mp4', width=960, height=720)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/tangyubin/movie_recommend/blob/master/movie_recommend.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
%matplotlib inline
import seaborn as sns
import tensorflow as tf
from tensorflow import keras
import os
import subprocess
import tempfile
import shutil
print(tf.__version__)
sess = tf.InteractiveSession()
a = tf.Variable(0.5)
b = tf.Variable(2.0)
x_1 = tf.placeholder(tf.float32, [None, 1], name = 'x_1')
x_2 = tf.placeholder(tf.float32, [None, 1], name = 'x_2')
y = tf.add(tf.multiply(a, x_1), tf.multiply(b, x_2))
sess.run(tf.global_variables_initializer())
sess.run(y, feed_dict={x_1:[[1]], x_2:[[1]]})
def save_model(sess, ii, oo, version=1):
MODEL_DIR = tempfile.gettempdir()
export_path = os.path.join(MODEL_DIR, str(version))
print('export_path = {}\n'.format(export_path))
if os.path.isdir(export_path):
print('\nAlready save a model, cleaning up\n')
shutil.rmtree(export_path, True)
print('\nDelete {} completed!\n'.format(export_path))
tf.saved_model.simple_save(
sess,
export_path,
inputs = ii,
outputs = oo
)
print('\nSave model:\n')
print(os.listdir(export_path))
save_model(sess, {'x_1':x_1, 'x_2':x_2}, {'y':y})
os.listdir('/tmp/1/variables')
!saved_model_cli show --dir /tmp/1 --all
!echo "deb http://storage.googleapis.com/tensorflow-serving-apt stable tensorflow-model-server tensorflow-model-server-universal" | tee /etc/apt/sources.list.d/tensorflow-serving.list && \
curl https://storage.googleapis.com/tensorflow-serving-apt/tensorflow-serving.release.pub.gpg | apt-key add -
!apt update
!apt-get install tensorflow-model-server
os.environ["MODEL_DIR"] = tempfile.gettempdir()
%%bash --bg
nohup tensorflow_model_server \
--rest_api_port=8501 \
--model_name=fashion_model \
--model_base_path="${MODEL_DIR}" >server.log 2>&1
!tail server.log
import json
req = {
'signature_name':'serving_default',
'instances':[
{
'x_1':1,
'x_2':1
}
]
}
data = json.dumps(req)
!pip install -q requests
import requests
headers = {"content-type":"application/json"}
json_response = requests.post('http://localhost:8501/v1/models/fashion_model:predict',
data=data, headers=headers)
predictions = json.loads(json_response.text)['predictions']
predictions
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import os
pd.set_option('display.max_columns', 100000)
pd.set_option('display.max_row', 1000000)
%matplotlib inline
%matplotlib inline
import numpy as np
import pandas as pd
from pandas.plotting import scatter_matrix
from imblearn.over_sampling import SMOTE
from imblearn.under_sampling import RandomUnderSampler
import yellowbrick as yb
from yellowbrick.features import RadViz
from yellowbrick.model_selection import LearningCurve
import pylab as pl
import seaborn as sns
from matplotlib import cm
import matplotlib.pyplot as plt
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.impute import SimpleImputer
from sklearn.metrics import f1_score
from sklearn.pipeline import Pipeline
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
from sklearn.naive_bayes import MultinomialNB
from sklearn.svm import LinearSVC, NuSVC, SVC
from sklearn.ensemble import AdaBoostClassifier
from sklearn.model_selection import StratifiedKFold
from sklearn.neighbors import KNeighborsClassifier
from sklearn.model_selection import train_test_split as tts
from sklearn.linear_model import LogisticRegressionCV, LogisticRegression, SGDClassifier
from sklearn.ensemble import BaggingClassifier, ExtraTreesClassifier, RandomForestClassifier
from sklearn.preprocessing import OneHotEncoder, LabelEncoder
from sklearn.preprocessing import StandardScaler, MinMaxScaler
```
### AirBNB
Hypothesis - The more Reviews the higher the price the listings
# Read CSV to Pandas Dataframe
```
# Congress Soars to New Heights on Social Media - https://www.pewresearch.org/internet/2020/07/16/congress-soars-to-new-heights-on-social-media/
abnb_listing_bc =pd.read_csv('/Users/ellemafa/Downloads/listings.csv')
abnb_listing_bc.head()
```
### Drop Columns of No Use
```
abnb_listing_bc = abnb_listing_bc[['host_id', 'neighbourhood_group', 'price', 'room_type', 'number_of_reviews', 'reviews_per_month', 'availability_365']]
abnb_listing_bc.neighbourhood_group.value_counts().plot.barh()
abnb_listing_bc.room_type.value_counts().plot.barh()
abnb_listing_bc.describe(include='all')
abnb_listing_bc.isna().sum()
abnb_listing_bc['availability_365']
```
## Remove all rows within availability_365 column that == 0
```
abnb_listing_bc_365 = abnb_listing_bc[~(abnb_listing_bc.availability_365 == 0)]
abnb_listing_bc_365[abnb_listing_bc_365.availability_365 == '0']
```
## Remove all rows within reviews_per_month column that are null
```
df = abnb_listing_bc_365.dropna()
len(df)
abnb_listing_bc_365.isna().sum()
df.head()
#historgram of all numerical data
df.hist(bins=30, figsize=(12,12))
pl.suptitle("Histogram for each numeric input variable: AirBNB Barcelona")
plt.show()
```
### Set X, y for deeper feature evaluation
```
df.columns
# label encoding the data
from sklearn.preprocessing import LabelEncoder
le = LabelEncoder()
df['neighbourhood_group']= le.fit_transform(df['neighbourhood_group'])
df['room_type']= le.fit_transform(df['room_type'])
#feature we will go with
feature_names = ['host_id', 'price', 'neighbourhood_group',
'number_of_reviews', 'reviews_per_month', 'availability_365']
X = df[feature_names]
y = df['room_type']
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=123)
import statsmodels.api as sm
#fit linear regression model
model = sm.OLS(y, X).fit()
#view model summary
print(model.summary())
from yellowbrick.classifier import ConfusionMatrix, ROCAUC, PrecisionRecallCurve, ClassificationReport
import pandas as pd # To read data
from sklearn.linear_model import LinearRegression
linear_regressor = LinearRegression() # create object for the class
linear_regressor.fit(X, y) # perform linear regression
Y_pred = linear_regressor.predict(X) # make predictions
print(np.sqrt(mean_squared_error(y,Y_pred)))
print(r2_score(y, Y_pred))
from sklearn.linear_model import Ridge
from sklearn.ensemble import RandomForestRegressor
from sklearn.svm import SVR
from sklearn.metrics import r2_score
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_squared_error
from sklearn.feature_selection import SelectFromModel
from yellowbrick.regressor import PredictionError, ResidualsPlot
from yellowbrick.features import FeatureImportances
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import train_test_split
#using CV
regressor = Ridge(alpha=0.5)
regressor = regressor.fit(X_train, y_train)
#cross validation, 10 fold to try next
scores = cross_val_score(estimator=regressor, X=X_train, y=y_train, cv=10, scoring="neg_mean_squared_error")
rmse_scores = np.sqrt(-scores)
print(rmse_scores)
pred_train = regressor.predict(X_test)
print(rmse_scores.mean())
print(r2_score(y_test, pred_train))
model = Ridge(random_state=0)
#labels = ['age', 'absences', 'famrel', 'freetime', 'goout', 'Dalc', 'Walc', 'health', 'traveltime', 'studytime', 'failures', 'school_GP', 'school_MS', 'sex_F', 'sex_M', 'famsize_GT3', 'famsize_LE3', 'Pstatus_A', 'Pstatus_T', 'Mjob_at_home', 'Mjob_health', 'Mjob_other', 'Mjob_services', 'Mjob_teacher', 'Fjob_at_home', 'Fjob_health', 'Fjob_other', 'Fjob_services', 'Fjob_teacher', 'reason_course', 'reason_home', 'reason_other', 'reason_reputation', 'guardian_father', 'guardian_mother', 'guardian_other','G1', 'G2']
viz = FeatureImportances(model, labels=feature_names, size=(1080, 720))
viz.fit(X_train, y_train)
# Note: the FeatureImportances visualizer is a model visualizer,
# not a feature visualizer, so it doesn't have a transform method!
viz.show()
```
## Random Forest
```
regressor = RandomForestRegressor(n_estimators=100, random_state=None, min_samples_split=100)
regressor = regressor.fit(X, y)
#cross validation, 10 fold to try next
scores = cross_val_score(estimator=regressor,X = X, y = y, cv=10, scoring="neg_mean_squared_error")
rmse_scores = np.sqrt(-scores)
print(rmse_scores)
pred_test= regressor.predict(X)
print(np.sqrt(mean_squared_error(y,pred_test)))
print(r2_score(y, pred_test))
```
# SVM
```
regressor = SVR(kernel = 'poly')
regressor = regressor.fit(X, y)
#cross validation, 10 fold to try next
scores = cross_val_score(estimator=regressor, X=X, y=y, cv=10, scoring="neg_mean_squared_error")
rmse_scores = np.sqrt(-scores)
print(rmse_scores)
pred_test= regressor.predict(X)
print(np.sqrt(mean_squared_error(y,pred_test)))
print(r2_score(y, pred_test))
```
| github_jupyter |
```
%reload_ext nb_black
import json
import pandas as pd
with open("../secrets.json", "r") as f:
secrets = json.load(f)
import spotipy
import spotipy.util as util
from spotipy.oauth2 import SpotifyClientCredentials
import spotipy.oauth2 as oauth2
CLIENT_ID = secrets["spotify_client_id"]
CLIENT_SECRET = secrets["spotify_client_secret"]
credentials = oauth2.SpotifyClientCredentials(
client_id=CLIENT_ID, client_secret=CLIENT_SECRET
)
token = credentials.get_access_token()
sp = spotipy.Spotify(auth=token)
# track = "coldplay yellow"
# res = spotify.search(track, type="track", market="US", limit=1)
# print(res)
s_limit = 10
res = sp.search("study", limit=s_limit, type="playlist", market="US")
playlist_info = {}
max_results = 30
offset = 0
while offset < max_results:
playlists = res["playlists"]
for i, item in enumerate(playlists["items"]):
# Store the name as a key. Get the number of tracks and uri
playlist_info[item["name"]] = [item["tracks"]["total"], item["uri"]]
if playlists["next"]:
res = sp.next(playlists)
offset += s_limit
else:
res = None
playlist_info
#pd.DataFrame(playlist_info).T.to_csv("../data/study_playlist_names.csv")
all_track_ids = []
for p in list(playlist_info.items()):
pl_id = p[1][1]
offset = 0
track_ids = []
while True:
response = sp.playlist_tracks(pl_id,
offset=offset,
fields='items.track.id,total',
additional_types=['track'])
#store the track ids from the playlist
for item in response['items']:
try:
track_ids.append( item['track']['id'])
except:
pass
offset = offset + len(response['items'])
print(offset, "/", response['total'])
if len(response['items']) == 0:
break
#Add the per_playlist ids to the master list
all_track_ids.extend(track_ids)
#track_ids.append(response['items'][0]['track']['id'])
print(json.dumps(response, indent=4, sort_keys=True))
len(all_track_ids)
track_ids.append(1)
pd.DataFrame(all_track_ids).to_csv("study_playlists_track_ids.csv")
all_track_ids[0]
track_info = []
import time
st = 0
end = len(all_track_ids)
# end = 200
step = 50
list(range(st, end, step))
for i in range(st, end, step):
print(i)
if len(all_track_ids) - i >= step - 1:
response = sp.tracks(all_track_ids[i : i + step])
else:
response = sp.tracks(all_track_ids[i:])
time.sleep(2)
track_info.append(response)
info_dict = {}
for batch in track_info:
type(batch)
for ind in batch["tracks"]:
track_artist = ind["artists"][0]["name"]
track_name = ind["name"]
track_album = ind["album"]["name"]
track_popularity = ind["popularity"]
track_id = ind["id"]
info_dict[track_id] = [track_artist, track_name, track_album, track_popularity]
len(info_dict)
tdf = pd.DataFrame(info_dict).T
tdf = tdf.reset_index()
# tdf.columns
tdf = tdf.rename(
columns={"index": "id", 0: "artist", 1: "title", 2: "album", 3: "popularity"}
)
tdf.head()
tdf.to_csv("../data/study_track_info.csv")
track_features = []
import time
st = 0
end = len(all_track_ids)
# end = 200
step = 100
list(range(st, end, step))
for i in range(st, end, step):
print(i)
if len(all_track_ids) - i >= step - 1:
response = sp.audio_features(all_track_ids[i : i + step])
else:
response = sp.audio_features(all_track_ids[i:])
time.sleep(0.3)
track_features.append(response)
dfs = []
for item in track_features:
df = pd.DataFrame.from_dict(item)
dfs.append(df)
study_track_features = pd.concat(dfs)
study_track_features.head()
study_track_features.to_csv("../data/study_track_features.csv")
study_track_info = tdf.copy()
study_track_features.drop(columns=["track_href", "analysis_url", "uri", "type"], inplace=True)
full_df = study_track_info.merge(study_track_features, on="id", how="outer")
full_df.shape
full_df.to_csv("../data/study_playlist_tracks.csv")
```
| github_jupyter |
# SageMakerCV PyTorch Tutorial
SageMakerCV is a collection of computer vision tools developed to take full advantage of Amazon SageMaker by providing state of the art model accuracy, training speed, and training cost reductions. SageMakerCV is based on the lessons we learned from developing the record breaking computer vision models we announced at Re:Invent in 2019 and 2020, along with talking to our customers and understanding the challenges they faced in training their own computer vision models.
The tutorial in this notebook walks through using SageMakerCV to train Mask RCNN on the COCO dataset. The only prerequisite is to setup SageMaker studio, the instructions for which can be found in [Onboard to Amazon SageMaker Studio Using Quick Start](https://docs.aws.amazon.com/sagemaker/latest/dg/onboard-quick-start.html). Everything else, from getting the COCO data to launching a distributed training cluster, is included here.
## Setup and Roadmap
Before diving into the tutorial itself, let's take a minute to discuss the various tools we'll be using.
#### SageMaker Studio
[SageMaker Studio](https://aws.amazon.com/sagemaker/studio/) is a machine learning focused IDE where you can interactively develop models and launch SageMaker training jobs all in one place. SageMaker Studio provides a Jupyter Lab like environment, but with a number of enhancements. We'll just scratch the surface here. See the [SageMaker Studio Documentation](https://docs.aws.amazon.com/sagemaker/latest/dg/studio.html) for more details.
For our purposes, the biggest difference from regular Jupyter Lab is that SageMaker Studio allows you to change your compute resources as needed, by connecting notebooks to Docker containers on different ML instances. This is a little confusing to just describe, so let's walk through an example.
Once you've completed the setup on [Onboard to Amazon SageMaker Studio Using Quick Start](https://docs.aws.amazon.com/sagemaker/latest/dg/onboard-quick-start.html), go to the [SageMaker Console](https://us-west-2.console.aws.amazon.com/sagemaker) and click `Open SageMaker Studio` near the top right of the page.
<img src="../assets/SageMaker_console.png" style="width: 600px">
If you haven't yet created a user, do so via the link at the top left of the page. Give it any name you like. For execution role, you can either use an existing SageMaker role, or create a new one. If you're unsure, create a new role. On the `Create IAM Role` window, make sure to select `Any S3 Bucket`.
<img src="../assets/Create_IAM_role.png" style="width: 600px">
Back on the SageMaker Studio page, select `Open Studio` next to the user you just created.
<img src="../assets/Studio_domain.png" style="width: 600px">
This will take a couple minutes to start up the first time. Once it starts, you'll have a Jupyter Lab like interface running on a small instance with an attached EBS volume. Let's start by taking a look at the `Launcher` tab.
<img src="../assets/Studio_launcher.png" style="width: 750px">
If you don't see the `Launcher`, you can bring one up by clicking the `+` on the menu bar in the upper left corner.
<img src="../assets/Studio_menu_bar.png" style="width: 600px">
The `Launcher` gives you access to all kinds of tools. This is where you can create new notebooks, text files, or get a terminal for your instance. Try the `System Terminal`. This gives you a new terminal tab for your Studio instance. It's useful for things like downloading data or cloning github repos into studio. For example, you can run `aws s3 ls` to browse your current S3 buckets. Go ahead and clone this repo onto Studio with
`git clone https://github.com/aws-samples/amazon-sagemaker-cv`
Let's look at the launcher one more time. Bring another one up with the `+`. Notice you have an option for `Select a SageMaker image` above the button to launch a notebook. This allows you to select a Docker image that will launch on a new instance. The notebook you create will be attached to that new instance, along with the EBS volume on your Studio instance. Let's try it out. On the `Launcher` page, click the drop down menu next to `Select a SageMaker Image` and select `PyTorch 1.6 Python 3.6 (Optimzed for GPU)`, then click the `Notebook` button below the dropdown.
<img src="../assets/Select_pytorch_image.png" style="width: 600px">
Take a look at the upper righthand corner of the notebook.
<img src="../assets/notebook_pytorch_kernel.png" style="width: 600px">
The `Ptyhon 3 (PyTorch 1.6 Python 3.6 GPU Optimized)` refers to the kernel associated with this notebook. The `Unknown` refers to the current instance type. Click `Unknown` and select `ml.g4dn.xlarge`.
<img src="../assets/instance_types.png" style="width: 600px">
This will launch a `ml.g4dn.xlarge` instance and attach this notebook to it. This will take a couple of minutes, because Studio needs to download the PyTorch Docker image to the new instance. Once an instance has started, launching new notebooks with the same instance type and kernel is immediate. You'll also see the `Unknown` replaced with and instance description `4 vCPU + 16 GiB + 1 GPU`. You can also change instance as needed. Say you want to run your notebook on a `ml.p3dn.24xlarge` to get 8 GPUs. To change instances, just click the instance description. To get more instances in the menu, deselect `Fast launch only`.
Once your notebook is up and running, you can also get a terminal into your new instance.
<img src="../assets/Launch_terminal.png" style="width: 600px">
This can be useful for customizing your image with setup scripts, pip installing new packages, or using mpi to launch multi GPU training jobs. Click to get a terminal and run `ls`. Note that you have the same directories as your main Studio instance. Studio will attach the same EBS volume to all the instances you start, so all your files and data are shared across any notebooks you start. This means that you can prototype a model on a single GPU instance, then switch to a multi GPU instance while still having access to all of your data and scripts.
Finally, when you want to shut down instances, click the circle with a square in it on the left hand side.
<img src="../assets/running_instances.png" style="width: 600px">
This shows your current running instances, and the Docker containers attached to those instances. To shut them down, just click the power button to their right.
Now that we've explored studio a bit, let's get started with SageMakerCV. If you followed the instructions above to clone the repo, you should have `amazon-sagemaker-cv` in the file browser on the left. Navigate to `amazon-sagemaker-cv/pytorch/tutorial.ipynb` to open this notebook on your instance. If you still have a `g4dn` running, it should automatically attach to it.
The rest of this notebook is broken into 4 sections.
- Installing SageMakerCV and Downloading the COCO Data
Since we're using the base AWS Deep Learning Container image, we need to add the SageMakerCV tools. Then we'll download the COCO dataset and upload it to S3.
- Prototyping in Studio
We'll walk through how to train a model on Studio, how SageMakerCV is structured, and how you can add your own models and features.
- Launching a SageMaker Training Job
There's lots of bells and whistles available to train your models fast, an on large datasets. We'll put a lot of those together to launch a high performance training job. Specifically, we'll create a training job with 4 P4d.24xlarge instances connected with 400 GB EFA, and streaming our training data from S3, so we don't have to load the dataset onto the instances before training. You could even use this same configuration to train on a dataset that wouldn't fit on the instances. If you'd rather only launch a smaller (or larger) training cluster, we'll discuss how to modify configuration.
- Testing Our Model
Finally, we'll take the output trained Mask RCNN model and visualize its performance in Studio.
#### Installing SageMakerCV
To install SageMakerCV on the PyTorch Studio Docker, just run `pip install -e .` in the `amazon-sagemaker-cv/pytorch` directory. You can do this with either an image terminal, or by running the paragraph below. Note that we use the `-e` option. This will keep the SageMakerCV modules editable, so any changes you make will be launched on your training job.
```
!pip install -e .
```
***
### Setup on S3 and Download COCO data
Next we need to setup an S3 bucket for all our data and results. Enter a name for your S3 bucket below. You can either create a new bucket, or use an existing bucket. If you use an existing bucket, make sure it's in the same region where you plan to run training. For new buckets, we'll specify that it needs to be in the current SageMaker region. By default we'll put everything in an S3 location on your bucket named `smcv-tutorial`, and locally in `/root/smcv-tutorial`, but you can change these locations.
```
S3_BUCKET = 'sagemaker-smcv-tutorial' # Don't include s3:// in your bucket name
S3_DIR = 'smcv-pytorch-tutorial'
LOCAL_DATA_DIR = '/root/smcv-pytorch-tutorial' #for reasons detailed in Destributed Training, do not put this dir in your source dir
import os
import zipfile
from pathlib import Path
from s3fs import S3FileSystem
from concurrent.futures import ThreadPoolExecutor
import boto3
from botocore.client import ClientError
from tqdm import tqdm
s3 = boto3.resource('s3')
boto_session = boto3.session.Session()
region = boto_session.region_name
# Check if bucket exists. If it doesn't, create it.
try:
bucket = s3.meta.client.head_bucket(Bucket=S3_BUCKET)
print(f"S3 Bucket {S3_BUCKET} Exists")
except ClientError:
print(f"Creating Bucket {S3_BUCKET}")
bucket = s3.create_bucket(Bucket=S3_BUCKET, CreateBucketConfiguration={'LocationConstraint': region})
```
***
Next we'll download the COCO data to Studio, unzip the files, and upload to S3. The reason we want the data in two places is that it's convenient to have the data locally on Studio for prototyping. We also want to unarchive the data before moving it to S3 so that we can stream it to our training instances instead of downloading it all at once.
Once this is finished, you'll have copies of the COCO data on your Studio instance, and in S3. Be careful not to open the `data/coco/train2017` dir in the Studio file browser. It contains 118287 images, and can cause your web browser to crash. If you need to browse these files, use the terminal.
This only needs to be done once, and only if you don't already have the data. The COCO 2017 dataset is about 20GB, so this step takes around 30 minutes to complete. The next paragraph sets up all the file directories we'll use for downloading, and later in training.
```
COCO_URL="http://images.cocodataset.org"
ANNOTATIONS_ZIP="annotations_trainval2017.zip"
TRAIN_ZIP="train2017.zip"
VAL_ZIP="val2017.zip"
COCO_DIR=os.path.join(LOCAL_DATA_DIR, 'data', 'coco')
os.makedirs(COCO_DIR, exist_ok=True)
S3_DATA_LOCATION=os.path.join("s3://", S3_BUCKET, S3_DIR, "data", "coco")
S3_WEIGHTS_LOCATION=os.path.join("s3://", S3_BUCKET, S3_DIR, "data", "weights")
WEIGHTS_DIR=os.path.join(LOCAL_DATA_DIR, 'data', 'weights')
os.makedirs(WEIGHTS_DIR, exist_ok=True)
R50_WEIGHTS="resnet50.pkl"
R50_WEIGHTS_SRC="https://sagemakercv.s3.us-west-2.amazonaws.com/weights/pytorch"
```
***
And this paragraph will download everything, and take around 30 minutes to complete.
```
print("Downloading annotations")
!wget -O $COCO_DIR/$ANNOTATIONS_ZIP $COCO_URL/annotations/$ANNOTATIONS_ZIP
!unzip $COCO_DIR/$ANNOTATIONS_ZIP -d $COCO_DIR
!rm $COCO_DIR/$ANNOTATIONS_ZIP
!aws s3 cp --recursive $COCO_DIR/annotations $S3_DATA_LOCATION/annotations
print("Downloading COCO training data")
!wget -O $COCO_DIR/$TRAIN_ZIP $COCO_URL/zips/$TRAIN_ZIP
# train data has ~128000 images. Unzip is too slow, about 1.5 hours beceause of disk read and write speed on the EBS volume.
# This technique is much faster because it grabs all the zip metadata at once, then uses threading to unzip multiple files at once.
print("Unzipping COCO training data")
train_zip = zipfile.ZipFile(os.path.join(COCO_DIR, TRAIN_ZIP))
jpeg_files = [image.filename for image in train_zip.filelist if image.filename.endswith('.jpg')]
os.makedirs(os.path.join(COCO_DIR, 'train2017'))
with ThreadPoolExecutor() as executor:
threads = list(tqdm(executor.map(lambda x: train_zip.extract(x, COCO_DIR), jpeg_files), total=len(jpeg_files)))
# same issue for uploading to S3. this uploads in parallel, and is faster than using aws cli in this case.
print("Uploading COCO training data to S3")
train_images = [i for i in Path(os.path.join(COCO_DIR, 'train2017')).glob("*.jpg")]
s3fs = S3FileSystem()
with ThreadPoolExecutor() as executor:
threads = list(tqdm(executor.map(lambda image: s3fs.put(image.as_posix(), os.path.join(S3_DATA_LOCATION, 'train2017', image.name)),
train_images), total=len(train_images)))
# !rm $COCO_DIR/$TRAIN_ZIP
print("Downloading COCO validation data")
!wget -O $COCO_DIR/$VAL_ZIP $COCO_URL/zips/$VAL_ZIP
# switch to also threading
!unzip -q $COCO_DIR/$VAL_ZIP -d $COCO_DIR
val_images = [i for i in Path(os.path.join(COCO_DIR, 'val2017')).glob("*.jpg")]
with ThreadPoolExecutor() as executor:
threads = list(tqdm(executor.map(lambda image: s3fs.put(image.as_posix(), os.path.join(S3_DATA_LOCATION, 'val2017', image.name)),
val_images), total=len(val_images)))
# !rm $COCO_DIR/$VAL_ZIP
# grab resnet backbone from public S3 bucket
!wget -O $WEIGHTS_DIR/$R50_WEIGHTS $R50_WEIGHTS_SRC/$R50_WEIGHTS
!aws s3 cp $WEIGHTS_DIR/$R50_WEIGHTS $S3_WEIGHTS_LOCATION/$R50_WEIGHTS
print("FINISHED!")
```
***
### Training on Studio
Now that we have the data, we can get to training a Mask RCNN model to detect objects in the COCO dataset images.
Since training on a single GPU can take days, we'll just train for a couple thousands steps, and run a single evaluation to make sure our model is at least starting to learn something. We'll train a full model on a larger cluster of GPUs in a SageMaker training job.
The reason we first want to train in Studio is that we want to dig a bit into the SageMakerCV framework, and talk about the model architecture, since we expect many users will want to modify models for their own use cases.
#### Mask RCNN
First, just a very brief overview of Mask RCNN. If you would like a more in depth examination, we recommend taking a look at the [original paper](https://arxiv.org/abs/1703.06870), the [feature pyramid paper](https://arxiv.org/abs/1612.03144) which describes a popular architectural change we'll use in our model, and blog posts from [viso.ai](https://viso.ai/deep-learning/mask-r-cnn/), [tryo labs](https://tryolabs.com/blog/2018/01/18/faster-r-cnn-down-the-rabbit-hole-of-modern-object-detection/), [Jonathan Hui](https://jonathan-hui.medium.com/image-segmentation-with-mask-r-cnn-ebe6d793272), and [Lilian Weng](https://lilianweng.github.io/lil-log/2017/12/31/object-recognition-for-dummies-part-3.html).
Mask RCNN is a two stage object detection model that locates objects in images by places bounding boxes around, and segmentation masks over, any object for which the model is trained to find. It also provides classifcations for each object.
<img src="../assets/traffic.png" style="width: 1200px">
Mask RCNN is called a two stage model because it performs detection in two steps. The first identified any objects in the image, versus background. The second stage determines the specific class of each object, and applies the segmentation mask. Below is an architectural diagram of the model. Let's walk through each step.
<img src="../assets/mask_rcnn_arch.jpeg" style="width: 1200px">
Credit: Jonathan Hui
The `Convolution Network` is often referred to as the model backbone. This is a pretrained image classification model, commonly ResNet, which has been trained on a large image classification dataset, like ImageNet. The classification layer is removed, and instead the backbone outputs a set of convolution feature maps. The idea is, the classification model learned to identify objects in the process of classifying images, and now we can use that information to build a more complex model that can find those objects in the image. We want to pretrain because training the backbone at the same time as training the object detector tends to be very unstable.
One additional component that is sometimes added to the backbone is a `Fearure Pyramid Network`. This take the outputs of the backbone, and combines them to together into a new set of feature maps by perform both up and down convolutions. The idea is that the different sized feature maps will help the model detect images of different sizes. The feature pyramid also helps with this, by allowing the different feature maps to share information with each other.
The outputs of the feature pyramid are then passed to the `Region Proposal Network` which is responsible for finding regions of the image that might contain an object (this is the first of the two stages). The RPN will output several hundred thousand regions, each with a probability of containing an object. We'll typically take the top few thousand most likely regions. Because these several thousand regions will usually have a lot of overlap, we perform [non-max supression](https://towardsdatascience.com/non-maximum-suppression-nms-93ce178e177c), which removed regions with large areas of overlap. This gives us a set of `regions of interest` regions of the image that we think might contain an image.
Next, we use those regions to crop out the corresponding sections of the feature maps that came from the feature pyramid network using a technique called [ROI align](https://firiuza.medium.com/roi-pooling-vs-roi-align-65293ab741db).
We pass our cropped feature maps to the `box head` which classifies each region into either a specific object category, or as background. It also refines the position of the bounding box. In Mask RCNN, we also pass the feature maps to a `mask head` which produces a segmentation mask over the object.
#### SageMakerCV Internals
An important feature of Mask RCNN is its multiple heads. One head constructs a bounding box, while another creates a mask. These are referred to as the `ROI heads`. It's common for users to extend this and other two stage models by adding their own ROI heads. For example, a keypoint head it common. Doing so means modifying SageMakerCV's internals, so let's talk about those for a second.
The high level Mask RCNN model can be found in `amazon-sageamaker-cv/pytorch/sagemakercv/detection/detector/generatlized_rcnn.py`. If you trace through the forward function, you'll see that the model first passes an image through the backbone (which also contains the feature pyramid), then the RPN in the graphable module. Then results are then passed through non-max suppression, and into the roi heads.
Probably the most important feature to be aware of are the `build` imports at the top. Each section of the model has an associated build function `(build_backbone, build_rpn, build_roi_heads)`. These functions simplify building the model by letting us pass in a single configuration file for building all the different pieces.
For example, if you open `amazon-sageamaker-cv/pytorch/sagemakercv/detection/roi_heads/roi_heads.py`, you'll find the `build_roi_heads` function at the bottom. To add a new head, you would write a torch module with its own build function, and call the build function from here.
For example, say you want to add a keypoint head to the model. An example keypoint module and associate build function is in `amazon-sageamaker-cv/pytorch/sagemakercv/detection/roi_heads/keypoint_head/keypoint_head.py`. To enable the keypoint head, you would set `cfg.MODEL.KEYPOINY_ON=True` and add the keypoint parameters to your configuration yaml file.
SageMakerCV uses similar build functions for the optimizers and schedulers, which you can add to or modify in the `amazon-sageamaker-cv/pytorch/sagemakercv/training/optimizers/` directory.
Finally, data loading tools are located in `amazon-sagemaker-cv/pytorch/data/`. Here you can add a new dataset, sampler, and preprocessing data transformations. Data loaders are constructed in the `build.py` file. Notice the `@DATASETS.register("COCO")` decorator at the top of the COCO `make_coco_dataloader` function. This adds the function to a dictionary of datasets, so that when you specify `COCO` in yout configuration file, the `make_data_loader` knows which data loader to create.
#### Setting Up Training
Let's actually use some of these functions to train a model.
Start by importing the default configuration file.
```
from configs import cfg
```
***
We use the [yacs](https://github.com/rbgirshick/yacs) format for configuration files. If you want to see the entire config, run `print(cfg.dump())` but this prints out a lot, and to not overwhelm you with too much information, we'll just focus on the bits we want to change for this model.
***
First, let's put in all the file directories for the data and weights we downloaded in the previous section, as well as an output directory for the model results.
```
cfg.INPUT.TRAIN_INPUT_DIR = os.path.join(COCO_DIR, "train2017")
cfg.INPUT.VAL_INPUT_DIR = os.path.join(COCO_DIR, "val2017")
cfg.INPUT.TRAIN_ANNO_DIR = os.path.join(COCO_DIR, "annotations", "instances_train2017.json")
cfg.INPUT.VAL_ANNO_DIR = os.path.join(COCO_DIR, "annotations", "instances_val2017.json")
cfg.OUTPUT_DIR = os.path.join(LOCAL_DATA_DIR, "model-output")
# create output dir if it doesn't exist
os.makedirs(cfg.OUTPUT_DIR, exist_ok=True)
# backbone weights file
cfg.MODEL.WEIGHT=os.path.join(WEIGHTS_DIR, R50_WEIGHTS)
```
****
Next we need to setup our data loader. The data loader is a tool in PyTorch that handles loading and applying transformations on our dataset. For more information see the [PyTorch Dataset and Data Loader Documentation](https://pytorch.org/tutorials/beginner/basics/data_tutorial.html).
For our purposes, we need to set two parameters. The size divisibility has to do with downsampling that occurs in the backbone and feature pyramid. The feature maps generated by these layers must evenly divide into the original image size. For example, if the input image is 1344x1344, the smallest feature map will be 42x42, which means the input image must be divisible by 32 `(42x32=1344)`. The `SIZE_DIVISIBLITY` parameter makes sure all images are resized to be multiples of 32.
The number of workers has to do with the number of background processes that will run the data loader in parallel. This can be useful for really high performance systems, to make sure our data loader is feeding data to the GPUs fast enough. However, for prototyping, these background processes can cause memory problems, so we'll turn it off when running in a notebook. We'll switch them back on later for our larger training job.
```
# dataloader settings
cfg.DATALOADER.SIZE_DIVISIBILITY=32
cfg.DATALOADER.NUM_WORKERS=0
```
***
This section specifies model details, including the type of model, and internal hyperparameters. We wont cover the details of all of these, but more information can be found in this blog posts listed above, as well as the original paper.
```
cfg.MODEL.META_ARCHITECTURE="GeneralizedRCNN" # The type of model we're training. found in amazon-sagemaker-cv/pytorch/sagemakercv/detection/detector/generalized_rcnn.py
cfg.MODEL.RESNETS.TRANS_FUNC="BottleneckWithFixedBatchNorm" # Type of bottleneck function in the Resnet50 backbone. see https://arxiv.org/abs/1512.03385
cfg.MODEL.BACKBONE.CONV_BODY="R-50-FPN" # Type of backbone, Resnet50 with feature pyramid network
cfg.MODEL.BACKBONE.OUT_CHANNELS=256 # number of channels on the output feature maps from the backbone
cfg.MODEL.RPN.USE_FPN=True # Use Feature Pyramid. RPN needs to know this since FPN adds an extra feature map
cfg.MODEL.RPN.ANCHOR_STRIDE=(4, 8, 16, 32, 64) # positions of anchors, see blog posts for details
cfg.MODEL.RPN.PRE_NMS_TOP_N_TRAIN=2000 # top N anchors to keep before non-max suppression during training
cfg.MODEL.RPN.PRE_NMS_TOP_N_TEST=1000 # top N anchors to keep before non-max suppression during testing
cfg.MODEL.RPN.POST_NMS_TOP_N_TEST=1000 # top N anchors to keep after non-max suppression during testing
cfg.MODEL.RPN.FPN_POST_NMS_TOP_N_TRAIN=1000 # top N anchors to keep after non-max suppression during training
cfg.MODEL.RPN.FPN_POST_NMS_TOP_N_TEST=1000 # top N anchors to keep before non-max suppression during training
cfg.MODEL.RPN.FPN_POST_NMS_TOP_N_PER_IMAGE=True # Run NMS per FPN level
cfg.MODEL.RPN.LS=0.1 # label smoothing improves performance on less common categories
# ROI Heads
cfg.MODEL.ROI_HEADS.USE_FPN=True # Use Feature Pyramid. ROI needs to know this since FPN adds an extra feature map
cfg.MODEL.ROI_HEADS.BBOX_REG_WEIGHTS=(10., 10., 5., 5.) # Regression wieghts for bounding boxes, see blog posts
cfg.MODEL.ROI_BOX_HEAD.POOLER_RESOLUTION=7 # Pixel size of region cropped from feature map
cfg.MODEL.ROI_BOX_HEAD.POOLER_SCALES=(0.25, 0.125, 0.0625, 0.03125) # Pooling for ROI align
cfg.MODEL.ROI_BOX_HEAD.POOLER_SAMPLING_RATIO=2 # Sampling for ROI Align
cfg.MODEL.ROI_BOX_HEAD.FEATURE_EXTRACTOR="FPN2MLPFeatureExtractor" # Type of ROI feature extractor found in SageMakerCV core utils
cfg.MODEL.ROI_BOX_HEAD.PREDICTOR="FPNPredictor" # Predictor type used for inference found in SageMakerCV core utils
cfg.MODEL.ROI_BOX_HEAD.LOSS="GIoULoss" # Use GIoU loss, improves box performance https://giou.stanford.edu/GIoU.pdf
cfg.MODEL.ROI_BOX_HEAD.DECODE=True # Convert boxes to pixel positions
cfg.MODEL.ROI_BOX_HEAD.CARL=True # Use carl loss https://arxiv.org/pdf/1904.04821.pdf
cfg.MODEL.ROI_MASK_HEAD.POOLER_SCALES=(0.25, 0.125, 0.0625, 0.03125) # Mask head ROI align
cfg.MODEL.ROI_MASK_HEAD.FEATURE_EXTRACTOR="MaskRCNNFPNFeatureExtractor" # Mask feature extractor type in SageMakerCV core utils
cfg.MODEL.ROI_MASK_HEAD.PREDICTOR="MaskRCNNC4Predictor" # Predictor used for inference
cfg.MODEL.ROI_MASK_HEAD.POOLER_RESOLUTION=14 # Pixel size of region cropped from feature map
cfg.MODEL.ROI_MASK_HEAD.POOLER_SAMPLING_RATIO=2 # ROI align sampling ratio
cfg.MODEL.ROI_MASK_HEAD.RESOLUTION=28 # output resolution of mask
cfg.MODEL.ROI_MASK_HEAD.SHARE_BOX_FEATURE_EXTRACTOR=False # share feature extractor between box and mask heads
cfg.MODEL.MASK_ON=True # use mask head
```
***
Next we set up the configuration for training, including the optimizer, hyperparameters, batch size, and training length. Batch size is global, so if you set a batch size of 64 across 8 GPUs, it will be a batch size of 8 per GPU. SageMakerCV currently supports the following optimizere: SGD (stochastic gradient descent), Adam, Lamb, and NovoGrad [link - this speeds up training by allowing increased batch sizes], and the following learning rate schedulers: stepwise and cosine decay. New, custom optimizers and schedulers can be added by modifying the `sagemakercv/training/build.py` file.
For training on Studio, we'll just run for a few hundred steps. We'll be using SageMaker training instances for the full training on multiple GPUs.
We also set the mixed precision optimization level. This is a value between O0-O4. O0 means no optimization, and training purely in FP32. O1-O3 are varying degrees of mixed precision, explained in the [Nvidia Apex documentation](https://nvidia.github.io/apex/amp.html). O4 is a special optimization level that combines pure FP16 training with channel last memory optimizations. This is the optimization we used to achieve the performance we announced at Re:Invent in 2020. However, optimization levels O2-O4 introduce some numerical instability, and can take a lot of tuning to train properly. For most users, we recommend starting on O1, since this provides most performance benefits of mixed precision, while retaining good numeric stability.
```
cfg.SOLVER.OPTIMIZER="NovoGrad" # Type of optimizer, NovoGrad, Adam, SGD, Lamb
cfg.SOLVER.BASE_LR=0.004 # Learning rate after warmup [Suggested values]
cfg.SOLVER.BETA1=0.9 # Beta value for Novograd, Adam, and Lamb
cfg.SOLVER.BETA2=0.4 # Beta value for Novograd, Adam, and Lamb
cfg.SOLVER.ALPHA=.1 # Alpha for final value of cosine decay
cfg.SOLVER.LR_SCHEDULE="COSINE" # Decay type, COSINE or MULTISTEP
cfg.SOLVER.IMS_PER_BATCH=8 # Global training batch size, must be a multiple of the number of GPUs
cfg.SOLVER.WEIGHT_DECAY=0.0005 # Training weight decay applied as decoupled weight decay on optimizer. [Paper link]
cfg.SOLVER.MAX_ITER=2500 # Total number of training steps for local training
cfg.SOLVER.WARMUP_FACTOR=.01 # Starting learning rate as a multiple of the BASE_LR
cfg.SOLVER.WARMUP_ITERS=100 # Number of warmup steps to reach BASE_LR
cfg.SOLVER.GRADIENT_CLIPPING=0.0 # Gradient clipping norm, leave as 0.0 to disable gradient clipping
cfg.OPT_LEVEL="O1" # Mixed precision optimization level
cfg.TEST.IMS_PER_BATCH=16 # Evaluation batch size, must be a multiple of the number of GPUs
cfg.TEST.PER_EPOCH_EVAL=True # Eval after every epoch or only at the end of training for local training
```
***
Finally, SageMakerCV includes a number of training hooks, ie. tools that will trigger on certain events during training.
For example, the `DetectronCheckpointHook` tells the trainer to read and write model checkpoints in the [Detectron](https://github.com/facebookresearch/detectron2) format. This will read in a checkpoint at the beginning of training, if one is provided, and write a checkpoint after each epoch. In this case, our backbone weights will be the starting checkpoint.
The `AMP_Hook` applies automatic mixed precision to the model, based on the optimization level we set.
The `IterTimerHook` and `TextLoggerHook` record information about training step time and loss values at each iteration, and format the results to be easy to read in AWS CloudWatch.
The `COCOEvaluation` run evaluation either after each epoch, or at the end of training.
We can supply these hooks to the configuration as a list of strings.
```
cfg.HOOKS=["DetectronCheckpointHook",
"AMP_Hook",
"IterTimerHook",
"TextLoggerHook",
"COCOEvaluation"]
```
***
Let's save this configuration file as a yaml, so we have a record of how we created this training.
```
import yaml
from contextlib import redirect_stdout
from datetime import datetime
local_config_file = f"configs/local-config-studio.yaml"
with open(local_config_file, 'w') as outfile:
with redirect_stdout(outfile): print(cfg.dump())
```
***
Here's how you can load this saved yaml file and map it to the configuration for future training sessions.
```
cfg.merge_from_file(local_config_file)
```
***
Now let's actually run some training. Since we expect users will want to modify the training for their own cases, we'll use some of the training tools more directly than normal.
explain why we want to do this interactively all in train.py
```
import torch
from sagemakercv.detection.detector import build_detection_model # takes cfg builds model
from sagemakercv.training import make_optimizer, make_lr_scheduler # takes cfg builds opt sch
from sagemakercv.data import make_data_loader, Prefetcher # takes cfg builds data loader
from sagemakercv.utils.runner import build_hooks, Runner # model trainer
from sagemakercv.training.trainers import train_step # actual train step
from sagemakercv.utils.runner.hooks.checkpoint import DetectronCheckpointHook # need to run this hook first, explain why
```
***
Create our dataloader
`num_iterations` is the number of expected steps per epoch. It's the size of the dataset divided by the global batch size. It's used so that we can keep track of when we've reached the end of an epoch. For this small local training, we'll actually be training for fewer steps than a full epoch.
```
train_coco_loader, num_iterations = make_data_loader(cfg, is_distributed=False) #local traiining explain num_iterations
```
***
The prefetcher boosts performance by asynchronously grabbing the next training element, and sending it to the GPU before it's needed.
```
device = torch.device(cfg.MODEL.DEVICE) # tell torch to use the GPU not the CPU as in default config
train_iterator = Prefetcher(iter(train_coco_loader), device)
```
***
Build the model, optimizer, and scheduler.
```
model = build_detection_model(cfg)
model.to(device) #send to GPU
optimizer = make_optimizer(cfg, model)
scheduler = make_lr_scheduler(cfg, optimizer)
```
***
Build the hooks we set in the config.
```
hooks = build_hooks(cfg)
```
***
Now we're to build a runner, which is a training tool designed for specifically for SageMaker. It manages your training steps, hooks, and logs, making it easier to track models on CloudWatch as well as managing sending data back and forth from S3. To build the runner, we need to define a `train_step`, which is just the standard pytorch training step. This is similar to Keras in TensorFlow but more customizable. For example, a very basic train_step would be something like:
```
def train_step(inputs, model, optimizer, scheduler):
optimizer.zero_grad()
loss = model(inputs)
loss.backward()
optimizer.step()
scheduler.step()
return loss
```
In this case, the `train_step` can be found in `sagemakercv/training/trainers.py` and includes all steps needed for Mask RCNN.
```
runner = Runner(model, train_step, cfg, device, optimizer, scheduler)
```
***
Register hooks with the runner so it knows when to trigger them. [explain detectron]
```
for hook in hooks:
runner.register_hook(hook, priority='HIGHEST' if isinstance(hook, DetectronCheckpointHook) else 'NORMAL')
```
***
And finally we're ready to run training. This will print a lot of info as it's setting up the model. When training starts, you'll sometimes see a few `Gradient Overflow` warnings. This is fine, it's just the mixed precision adjusting the loss scaling. After about a minute you should start seeing loss values in step increments of 50. At 2500 steps on a G4dn instane, this takes about an hour. You can speed this up by reducing the number of training steps, or using a P3.2xlarge instance. You should get a [MaP score](https://towardsdatascience.com/map-mean-average-precision-might-confuse-you-5956f1bfa9e2) of around .1 for both box and mask after 2500 steps.
```
runner.run(train_iterator, num_iterations)
```
***
Okay, so we printed a lot of numbers. Before moving on, let's first visualize what our model learned so we can make sure it's working how we expect.
SageMakerCV includes some simple visualization tools.
```
import os
import torch
import gc
from sagemakercv.data.datasets.evaluation.coco.coco_labels import coco_categories
from sagemakercv.utils.visualize import Visualizer
viz = Visualizer(model, cfg, temp_dir=cfg.OUTPUT_DIR, categories=coco_categories)
```
The visualizer can take a local file path, S3 location, or web address for an image. It will run the image through the model you just trained, and output it's predictions for objects it finds.
You can use any images you want, either from the Coco data we downloaded earlier, or your own. For example, [pixalbay](https://pixabay.com/) has lots of free use images. Just copy the image address into the `image_src` below.
```
image_src = 'https://cdn.pixabay.com/photo/2021/07/29/14/48/new-york-6507350_1280.jpg'
#'https://cdn.pixabay.com/photo/2020/05/12/11/39/cat-5162540__480.jpg'
viz(image_src, threshold=0.9) # Threshold is the minimum probability to display. Usually set around .75 or .9 so we don't get a bunch of spots where the model says there's a 2% chance of hot dogs.
```
***
Note that running interactively in a notebook can build of GPU memory pressure over time. If you run inference a lot of times, you might eventually get a cuda out of memory error. If this happens, just run the garbage collection in the cell below.
```
torch.cuda.empty_cache()
gc.collect()
```
***
Before moving on, let's delete our model and related training tools so we don't keep too much in memory. The model you just trained is saved in your `cfg.OUTPUT_DIR` directory, so you can reload it later. We'll cover how to import a saved model in the final section.
```
del model, optimizer, scheduler, runner
torch.cuda.empty_cache()
gc.collect()
```
***
### Distributed Training
Great! You've managed to train a model (albeit for just a few thousand steps) on your studio instance. For many users this will be all they need. If you only intend to deal with small amounts of data (<10G) that can be trained on a single GPU, you can do everything you need with what we've run so far.
However, most practical applications of Mask RCNN require training on huge datasets across many GPUs or nodes. For that, we'll need to launch a SageMaker training job. Here we can train a model on as many as 512 [A100 GPUs](https://www.nvidia.com/en-us/data-center/a100/). We won't go quite that far. Instead, let's try training on 32 A100 GPUs across 4 P4d nodes. We'll also cover how to modify the configuration for different GPU types and counts.
The section below is also replicated in the `SageMaker.ipynb` notebook for future training once all the above setup is complete.
Before we get started, a few notes about how SageMaker training instances work. SageMaker takes care of a lot of setup for you, but it's important to understand a little of what's happening under the hood so you can customize training to your own needs.
First we're going to look at a toy estimator to explain what's happening:
```
from sagemaker import get_execution_role
from sagemaker.pytorch import PyTorch
estimator = PyTorch(
entry_point='train.py',
source_dir='.',
py_version='py3',
framework_version='1.8.1',
role=get_execution_role(),
instance_count=4,
instance_type='ml.p4d.24xlarge',
distribution=distribution,
output_path='s3://my-bucket/my-output/',
checkpoint_s3_uri='s3://my-bucket/my-checkpoints/',
model_dir='s3://my-bucket/my-model/',
hyperparameters={'config': 'my-config.yaml'},
volume_size=500,
code_location='s3://my-bucket/my-code/',
)
```
The estimator forms the basic configuration of your training job.
SageMaker will first launch `instance_count=4` `instance_type=ml.p4d.24xlarge` instances. The `role` is an IAM role that SageMaker will use to launch instances on your behalf. SageMaker includes a `get_execution_role` function which grabs the execution role of your current instance. Each instance will have a `volume_size=500` EBS volume attached for your model and data. On `ml.p4d.24xlarge` and `ml.p3dn.24xlarge` instance types, SageMaker will automatically set up the [Elastic Fabric Adapter](https://aws.amazon.com/hpc/efa/). EFA provides up to 400 GB/s communication between your training nodes, as well as [GPU Direct RDMA](https://aws.amazon.com/about-aws/whats-new/2020/11/efa-supports-nvidia-gpudirect-rdma/) on `ml.p4d.24xlarge`, which allows your GPUs to bypass the host and communicate directly with each other across nodes.
Next, SageMaker we copy all the contents of `source_dir='.'` first to the `code_location='s3://my-bucket/my-code/'` S3 location, then to each of your instances. One common mistake is to leave large files or data in this directory or its subdirectories. This will slow down your launch times, or can even cause the launch to hang. Make sure to keep your working data and model artifacts elsewhere on your Studio instance so you don't accidently copy them to your training instance. You should instead use `Channels` to copy data and model artifacts, which we'll cover shortly.
SageMaker will then download the training Docker image to all your instances. Which container you download is determined by `py_version='py3'` and `framework_version='1.8.1'`. You can also use your own [custom Docker image](https://aws.amazon.com/blogs/machine-learning/bringing-your-own-custom-container-image-to-amazon-sagemaker-studio-notebooks/) by specifying an ECR address with the `image_uri` option. When building a custom container, we recommend building the [CUDA utilities](https://github.com/aws-samples/amazon-sagemaker-cv/tree/main/pytorch/cuda_utils) from source within the container, to ensure compatibility with your PyTorch version. SageMakerCV currently include prebuilt whls for CUDA utilities for versions 1.6-1.9.
Before starting training, SageMaker will check your source directory for a `setup.py` file, and install if one is present. Then SageMaker will actually launch training, via `entry_point='train.py'`. Anything in `hyperparameters={'config': 'my-config.yaml'}` will be passed to the training script as a command line argument (ie `python train.py --config my-config.yaml`). The distribution will determine what form of distributed training to launch. This will be covered in more detail later.
During training, anything written to `/opt/ml/checkpoints` on your training instances will be synced to `checkpoint_s3_uri='s3://my-bucket/my-checkpoints/'` at the same time. This can be useful for checkpointing a model you might want to restart later, or for writting Tensorboard logs to monitor your training.
When training complets, you can write your model artifacats to `/opt/ml/model` and it will save to `model_dir='s3://my-bucket/my-model/'`. Another option is to also write model artifacts to your checkpoints file.
Training logs, and any failure messages will to written to `/opt/ml/output` and saved to `output_path='s3://my-bucket/my-output/'`.
```
from sagemaker import get_execution_role
from sagemaker.pytorch import PyTorch
```
First we need to set some names. You want `AWS_DEFAULT_REGION` to be the same region as the S3 bucket your created earlier, to ensure your training jobs are reading from nearby S3 buckets.
Next, set a `user_id`. This is just for naming your training job so it's easier to find later. This can be anything you like. We also get the current date and time to make organizing training jobs a little easier.
```
# explain region. Don't launch a training job in VA with S3 bucket in OR
os.environ['AWS_DEFAULT_REGION'] = region # This is the region we set at the beginning, when creating the S3 bucket for our data
# this is all for naming
user_id="username-smcv-tutorial" # This is used for naming your training job, and organizing your results on S3. It can be anything you like.
date_str=datetime.now().strftime("%d-%m-%Y")
time_str=datetime.now().strftime("%d-%m-%Y-%H-%M-%S")
```
For instance type, we'll use an `ml.p4d.24xlarge`. We recommend this instance type for large training. It includes the latest A100 Nvidia GPUs, which can train several times faster than the previous generation. If you would rather train part way on smaller instanes, `ml.p3.2xlarge, ml.p3.8xlarge, ml.p3.16xlarge, ml.p3dn.24xlarge, ml.g4dn.12xlarge` are all good options. In particular, if you're looking for a low cost way to try a short distributed training, but aren't worried about the model fully converging, we recommend the `ml.g4dn.12xlarge` which uses 4 Nvidia T4 GPUs per node.
`s3_location` will be the base S3 storage location we used earlier for the COCO data. For `role` we get the execution role from our studio instance. For `source_dir` we use the current directory. Again, make sure you haven't accidently written any large files to this directory.
```
# specify training type, s3 src and nodes
instance_type="ml.p4d.24xlarge" # This can be any of 'ml.p3dn.24xlarge', 'ml.p4d.24xlarge', 'ml.p3.16xlarge', 'ml.p3.8xlarge', 'ml.p3.2xlarge', 'ml.g4dn.12xlarge'
nodes=1 # 4
s3_location=os.path.join("s3://", S3_BUCKET, S3_DIR)
role=get_execution_role() #give Sagemaker permission to launch nodes on our behalf
source_dir='.'
```
***
Let's modify our previous training configuration for multinode. We don't need to change much. We'll increase the batch size since we have more and large GPUs. For A100 GPUs a batch size of 12 per GPU works well. For V100 and T4 GPUs, a batch size of 6 per GPU is recommended. Make sure to lower the learning rate and increase your number of training steps if you decrease the batch size. For example, if you want to train on 2 `ml.g4dn.12xlarge` instances, you'll have 8 T4 GPUs. A batch size of `cfg.SOLVER.IMS_PER_BATCH=48`, with inference batch size of `cfg.TEST.IMS_PER_BATCH=32`, learning rate of `cfg.SOLVER.BASE_LR=0.006`, and training steps of `cfg.SOLVER.MAX_ITER=25000` is probably about right.
The configuration below has been tested to converge to better than MLPerf accuracy.
```
cfg.SOLVER.OPTIMIZER="NovoGrad" # Type of optimizer, NovoGrad, Adam, SGD, Lamb
cfg.SOLVER.BASE_LR=0.016 # 0.042 # Learning rate after warmup
cfg.SOLVER.BETA1=0.9 # Beta value for Novograd, Adam, and Lamb
cfg.SOLVER.BETA2=0.3 # Beta value for Novograd, Adam, and Lamb
cfg.SOLVER.ALPHA=.001 # Alpha for final value of cosine decay
cfg.SOLVER.LR_SCHEDULE="COSINE" # Decay type, COSINE or MULTISTEP
cfg.SOLVER.IMS_PER_BATCH=96 #384 # Global training batch size, must be a multiple of the number of GPUs
cfg.SOLVER.WEIGHT_DECAY=0.001 # Training weight decay applied as decoupled weight decay on optimizer
cfg.SOLVER.MAX_ITER=15000 #5000 # Total number of training steps
cfg.SOLVER.WARMUP_FACTOR=.01 # Starting learning rate as a multiple of the BASE_LR
cfg.SOLVER.WARMUP_ITERS=625 # Number of warmup steps to reach BASE_LR
cfg.SOLVER.GRADIENT_CLIPPING=0.0 # Gradient clipping norm, leave as 0.0 to disable gradient clipping
cfg.OPT_LEVEL="O1" # Mixed precision optimization level
cfg.TEST.IMS_PER_BATCH=64 # 128 # Evaluation batch size, must be a multiple of the number of GPUs
cfg.TEST.PER_EPOCH_EVAL=True # False # Eval after every epoch or only at the end of training
```
***
Earlier we mentioned the `distrbution` strategy in SageMaker. Distributed training can be either multi GPU single node (ie training on 8 GPU in a single ml.p4d.24xlarge) or mutli GPU multi node (ie training on 32 GPUs across 4 ml.p4d.24xlarges). For PyTorch you can use either [PyTorch's built in DDP](https://pytorch.org/tutorials/intermediate/ddp_tutorial.html) or [SageMaker Distributed Data Parallel](https://docs.aws.amazon.com/sagemaker/latest/dg/data-parallel.html) (SMDDP). For single node multi GPU we recommend PyTorch DDP, while for multinode we recommend SMDDP. SMDDP is built to fully utilize AWS network topology, and EFA, providing a speed boost on multinode.
To enable SMDDP, set `distribution = { "smdistributed": { "dataparallel": { "enabled": True } } }`. SageMakerCV already has SMDDP integrated. To implement SMDDP for your own models, follow [these instructions](https://docs.aws.amazon.com/sagemaker/latest/dg/data-parallel-intro.html). SMDDP will launch training from the first node in your cluster using [MPI](https://www.open-mpi.org/).
For PyTorch DDP we'll actually disable SageMaker's distribution tool, and set it up manually. When you disable distribution, SageMaker will simply launch the same `entry_point` script on each node. The setup below is to run PyTorch DDP for multi GPU on a single node, but you can use the same setup to run PyTorch DDP multinode as well. Manually setting up your own distribution can be useful when you want to do complex custom distribution strategies. For example, you can use the `SM_CURRENT_HOST` environment variable on each node to set node specific parameters. PyTorch DDP requires running the `torch.distributed.launch` on each training node. The `launch_ddp.py` script grabs the environment variables, then launches `train.py` with `torch.distributed.launch`.
```
if nodes>1 and instance_type in ['ml.p3dn.24xlarge', 'ml.p4d.24xlarge', 'ml.p3.16xlarge']:
distribution = { "smdistributed": { "dataparallel": { "enabled": True } } }
entry_point = "train.py"
else:
distribution = None
entry_point = "launch_ddp.py"
```
***
We'll set a job name based on the user name and time. We'll then set output directories on S3 using the date and job name.
For this training, we'll use the same S3 location for all 3 SageMaker model outputs `/opt/ml/checkpoint`, `/opt/ml/model`, and `/opt/ml/output`.
```
job_name = f'{user_id}-{time_str}'
output_path = os.path.join(s3_location, "sagemaker-output", date_str, job_name)
code_location = os.path.join(s3_location, "sagemaker-code", date_str, job_name)
```
***
Next we need to add our data sources to our configuration file, but first let's talk a little more about how SageMaker gets data to your instance.
The most straightforward way to get your data is using "Channels." These are S3 locations you specify in a dictionary when you launch a training job. For example, let's say you launch a training job with:
```
channels = {'train': 's3://my-bucket/data/train/',
'test': 's3://my-bucket/data/test/',
'weights': 's3://my-bucket/data/weights/',
'dave': 's3://my-bucket/data/daves_weird_data/'}
pytorch_estimator.fit(channels)
```
At the start of training, SageMaker will create a set of corresponding directories on each training node:
```
/opt/ml/input/data/train/
/opt/ml/input/data/test/
/opt/ml/input/data/weights/
/opt/ml/input/data/dave/
```
SageMaker will then copy all the contents of the corresponding S3 locations to these directories, which you can then access in training.
One downside of setting up channels like this is that it requires all the data to be downloaded to your instance at the start of of training, which can delay the training launch if you're dealing with a large dataset.
We have two ways to speed up launch. The first is [Fast File Mode](https://aws.amazon.com/about-aws/whats-new/2021/10/amazon-sagemaker-fast-file-mode/) which downloads data from S3 as it's requested by the training model, speeding up your launch time. You can use fast file mode by sepcifying `TrainingInputMode='FastFile'` in your SageMaker estimator configuration.
If you're dealing with really huge data, on the order of several terabytes, you might not want to keep if on the instance at all, and just stream it directly from S3 into your model. The [PyTorch S3 plugin](https://aws.amazon.com/blogs/machine-learning/announcing-the-amazon-s3-plugin-for-pytorch/) provides just this capability. To use the PyTorch S3 plugin, you need to build a PyTorch dataset using the S3 plugin base class. S3 plugin support is already built into SageMakerCV. If you want to see how it's implemented, the dataset can be found in `sagemakercv/data/datasets/coco.py`.
In our case, we'll use a mix of channels and the S3 plugin. We'll download the smaller pieces at the start of training (the validation data, pretrained weights, and image annotations), and we'll use the S3 plugin to stream the training data, since it's large. SageMakerCV will automatically switch to the S3 plugin when you supply an S3 location in the configuration file. So all we need to do is setup our channels for the data we want to download, then give the config file the locations either on the instance or on S3.
We also want to set the output dir to `/opt/ml/checkpoints`. SageMaker will sync the contents of this directory back to S3. Let's also turn up the number of workers so the data loads fast enough for the bigger GPUs we're using.
First, we setup our training channels.
```
channels = {'validation': os.path.join(S3_DATA_LOCATION, 'val2017'),
'weights': S3_WEIGHTS_LOCATION,
'annotations': os.path.join(S3_DATA_LOCATION, 'annotations')}
```
***
Next, in the configuration file, we need to provide the corresponding location on each training node for these channels. We need to specify locations for the validation data, train and validation annotations, and the backbone weights.
```
CHANNELS_DIR='/opt/ml/input/data/' # on node
cfg.INPUT.VAL_INPUT_DIR = os.path.join(CHANNELS_DIR, 'validation') # Corresponds to the vdalidation key in the channels
cfg.INPUT.TRAIN_ANNO_DIR = os.path.join(CHANNELS_DIR, 'annotations', 'instances_train2017.json')
cfg.INPUT.VAL_ANNO_DIR = os.path.join(CHANNELS_DIR, 'annotations', 'instances_val2017.json')
cfg.MODEL.WEIGHT=os.path.join(CHANNELS_DIR, 'weights', R50_WEIGHTS) # backbone weights file
```
For our training data location, we'll instead point it directly to S3 so it streams the data in.
```
cfg.INPUT.TRAIN_INPUT_DIR = os.path.join(S3_DATA_LOCATION, "train2017") # Set to S3 location so we use the S3 plugin
```
Set the output directory to the SageMaker's checkpoint directory. This way all the files our model writes out will be immediately copied to S3.
```
cfg.OUTPUT_DIR = '/opt/ml/checkpoints/'
```
Turn up the number of workers for the dataloader. This is especially important when using the S3 plugin. The plugin is fast, but there's still a bit of network overhead. Having more dataloaders grabbing elements from S3 simultaneously will mean our model isn't waiting around for data to load. A good rule of thunb is to set the number of dataloader workers equal to the number of vCPUs divded by the number of GPUs per instance. For example, on the `ml.p4d.24xlarge` there are 96 vCPUs and 8 GPUs, so we'll use 12 workers.
```
cfg.DATALOADER.NUM_WORKERS=12
cfg.HOOKS=["DetectronCheckpointHook",
"AMP_Hook",
"IterTimerHook",
"TextLoggerHook",
"COCOEvaluation"]
```
Like we did for the local training, we'll save the configuration file so we can replicate this same training later.
```
dist_config_file = f"configs/dist-training-config.yaml"
with open(dist_config_file, 'w') as outfile:
with redirect_stdout(outfile): print(cfg.dump())
```
***
And now we can launch our training job. Let's set the config file we just created as a hyperparameter for the estimator. This will get passed to our training script as a command line argument. For example, SageMaker would launch training with `python train.py --config configs/dist-training-config.yaml`
Using 4 P4d nodes, training takes about 45 minutes. This section will also print a lot of output logs. By setting `wait=False` you can avoid printing logs in the notebook. This setting will just launch the job then return, and is useful for when you want to launch several jobs at the same time. You can then montior each job from the [SageMaker Training Console](https://us-west-2.console.aws.amazon.com/sagemaker).
```
hyperparameters = {"config": dist_config_file}
estimator = PyTorch(
entry_point=entry_point,
source_dir=source_dir,
py_version='py3',
framework_version='1.8.1',
role=role,
instance_count=nodes,
instance_type=instance_type,
distribution=distribution,
output_path=output_path,
checkpoint_s3_uri=output_path,
model_dir=output_path,
hyperparameters=hyperparameters,
volume_size=500,
code_location=code_location,
disable_profiler=True, # Reduce number of logs since we don't need profiler or debugger for this training
debugger_hook_config=False,
)
estimator.fit(channels, wait=True, job_name=job_name)
```
***
### Part 5: Visualizing Results
And there you have it, a fully trained Mask RCNN model in under an hour. Now let's see how our model does on prediction by actually visualizing the output.
Our model is stored at the S3 location we gave to the training job in `output_path`. We'll need to grab the results and store them on our studio instance so we can check performance, and visualize the output.
```
import os
from s3fs import S3FileSystem
from configs import cfg
import torch
import gc
from sagemakercv.detection.detector import build_detection_model
from sagemakercv.utils.model_serialization import load_state_dict
from sagemakercv.data.datasets.evaluation.coco.coco_labels import coco_categories
from sagemakercv.utils.visualize import Visualizer
# Turn down logging, we don't need all the info about loading the state dict
import logging
logger = logging.getLogger()
logger.setLevel(logging.CRITICAL)
# Reuse the local configuration file we made earlier
cfg.merge_from_file(local_config_file)
s3fs = S3FileSystem()
TRAINING_OUTPUT=os.path.join(LOCAL_DATA_DIR, 'training_output') # Make a new local directory for our training results and new images
os.makedirs(TRAINING_OUTPUT, exist_ok=True)
# Grab the filename of the last checkpoint our training job wrote
s3fs.get(os.path.join(output_path, 'last_checkpoint'), os.path.join(TRAINING_OUTPUT, 'last_checkpoint'))
with open(os.path.join(TRAINING_OUTPUT, 'last_checkpoint'), 'r') as f:
checkpoint_name=f.readline().split('/')[-1]
# Copy the saved weights to our local directory
s3fs.get(os.path.join(output_path, checkpoint_name), os.path.join(TRAINING_OUTPUT, checkpoint_name))
```
***
To run inference, we first build a blank model and then add weights from the distributed training job.
We can build a new model the same way we built one for local training. This time, instead of using the checkpointer to load the backbone weights, we'll directly load all the saved weights from the trained model.
```
device = torch.device(cfg.MODEL.DEVICE)
model = build_detection_model(cfg)
_ = model.to(device)
# Load weights file as a dictionary
weights = torch.load(os.path.join(TRAINING_OUTPUT, checkpoint_name))['model']
# Map saved weights to our model
load_state_dict(model, weights, False)
```
***
Like we did with the local training, we can pass the model to the visualization tool, and give it an image either from local, S3, or a web address.
```
torch.cuda.empty_cache()
gc.collect()
viz = Visualizer(model, cfg, temp_dir=TRAINING_OUTPUT, categories=coco_categories)
image_src = 'https://cdn.pixabay.com/photo/2021/07/29/14/48/new-york-6507350_1280.jpg'
#'https://cdn.pixabay.com/photo/2020/05/12/11/39/cat-5162540__480.jpg'
viz(image_src, threshold=0.9) #explain threshold
```
#### Conclusion
In this notebook, we've walked through the entire process of training Mask RCNN on SageMaker. We've implemented several of SageMaker's more advanced features, such as distributed training, EFA, and streaming data directly from S3. From here you can use the provided template datasets to train on your own data, or modify the framework with your own object detection model.
When you're done, make sure to check that all of your SageMaker training jobs have stopped by checking the [SageMaker Training Console](https://us-west-2.console.aws.amazon.com/sagemaker). Also check that you've stopped any Studio instance you have running by selecting the session monitor on the left (the circle with a square in it), and clicking the power button next to any running instances. Your files will still be saved on the Studio EBS volume.
<img src="../assets/running_instances.png" style="width: 600px">
| github_jupyter |
```
# Data sources:
# 1. boston_road_massdot2015_MA.dbf, given by CoB
# 2. massdot_crashes_12_14_MA.dbf, given by CoB after associating each accident co-ordinate to the road segment (FID)
# 3. crashes_12-14.csv, Original data from MassDOT aggregated and by Josiah
# Convert dbf to csv
import pandas as pd
%matplotlib inline
import matplotlib as plt
from simpledbf import Dbf5
dbf_incident_data = Dbf5('massdot_crashes_12_14_MA.dbf',codec='utf-8')
incident_data = dbf_incident_data.to_dataframe()
# CoB data has truncated columns names from ArcGIS , fix that using Josiah's dataset
original = pd.read_csv('crashes_12-14.csv')
full_column_name = original.columns.tolist()
trunc_column_name = incident_data.columns.tolist()
full_column_name.extend(trunc_column_name[-5:])
incident_data.columns = full_column_name
# Convert column names into lowercase
incident_data.columns = [x.lower().strip() for x in incident_data.columns.tolist()]
# A single accident can have multiple entries, one for each vehicle involved in the crash
# Clean massdot crash data file to make single crash record for each row
# remove vehicle related details as its not possible to retain multiple vehicle details in a single row
incident_data.drop('vehicle_number',axis=1,inplace=True)
incident_data.drop('vehicle_configuration',axis=1,inplace=True)
incident_data.drop('vehicle_travel_directions',axis=1,inplace=True)
incident_data.drop('vehicle_action_prior_to_crash',axis=1,inplace=True)
# remove byte order mark from crash_number column
incident_data.rename(columns={'\xef\xbb\xbfcrash_number':'crash_number'}, inplace=True)
#Each incident is reported as multiple crashes. If one crash involves 2 vehicles, 2 crash reports are filed
#Making each incident has one entry
incident_data.drop_duplicates('crash_number', inplace=True)
# remove records without location records
incident_data = incident_data[incident_data['near_fid'] != -1]
#convert date to three new columns 'year', 'day' and 'hour' of crash report
from datetime import datetime
date_to_dayofweek_converter = lambda x: datetime.strptime(str(x), '%Y-%m-%d').strftime('%a')
date_to_year_converter = lambda x: datetime.strptime(str(x), '%Y-%m-%d').strftime('%y')
def time_to_hour_converter(x):
if x == '0S':
return -1
else:
return datetime.strptime(str(x), '%HH %MM %SS').strftime('%H')
incident_data['day'] = incident_data['crash_date'].apply(date_to_dayofweek_converter)
incident_data['year'] = incident_data['crash_date'].apply(date_to_year_converter)
incident_data['hour'] = incident_data['crash_time'].apply(time_to_hour_converter)
# read road data from csv to create a dataframe and convert column names to lowercase
dbf_roads_data = Dbf5('boston_road_massdot2015_MA.dbf',codec='utf-8')
road_data = dbf_roads_data.to_dataframe()
road_data.columns = road_data.columns.str.lower()
print road_data.columns
# Join crash_data and road_data on near_fid and strsegfid
crash_road_data = incident_data.merge(road_data, left_on='near_fid', right_on='strsegfid', how='left')
#drop zero information columms or duplicates
crash_road_data.drop('city_town_name',axis=1, inplace='True')
crash_road_data.drop('city_x',axis=1, inplace='True')
crash_road_data.drop('city_y',axis=1, inplace='True')
crash_road_data.drop('date_time',axis=1, inplace='True')
crash_road_data.drop('near_fid_y',axis=1, inplace='True')
crash_road_data.drop('near_fid_1',axis=1, inplace='True')
crash_road_data.drop('strsegfid',axis=1, inplace='True')
crash_road_data.drop('mpo',axis=1, inplace='True')
# save merged dataset
crash_road_data.to_csv('crash_road_12_14_MA_unique.csv')
#basic analysis
#Total accident reports from 2012 to 2014 with valid FID data : 11559
print len(crash_road_data)
#Total number of street segments in boston : 14,622
print len(road_data)
#Number of street segments that has seen a crash : 2,932
print len(crash_road_data.near_fid_x.unique())
#number of crashes every weekday
ax = crash_road_data.day.value_counts().plot(kind='bar')
ax.set_xlabel('Day of week')
ax.set_ylabel('Crash Incident Count (2012 to 2014)')
#number of crashes vs hour of day (-1 where the data has no time entry)
# not sure only 12 hours, there was not AM PM details in the data
ax = crash_road_data.hour.value_counts().plot(kind='bar')
ax.set_xlabel('Hour of day')
ax.set_ylabel('Crash incident count')
#number of crashes each year
ax = crash_road_data.year.value_counts().plot(kind='bar')
ax.set_xlabel('Year')
ax.set_ylabel('Crash incident count')
#number of crashes vs ambient light
ax = crash_road_data.ambient_light.value_counts().plot(kind='bar')
ax.set_xlabel('Ambient light')
ax.set_ylabel('Crash incident count')
#number of crashes vs road surface
ax = crash_road_data.road_surface_condition.value_counts().plot(kind='bar')
ax.set_xlabel('Road Surface')
ax.set_ylabel('Crash incident counts')
#number of crashes vs facility type
ax = crash_road_data.facility.value_counts().plot(kind='bar')
ax.set_xlabel('Road Type (1:Mainline roadway, 3:Tunnel, 5:Ramp to Tunnel, 7:Simple Ramp)')
ax.set_ylabel('Crash incident count')
print crash_road_data.facility.value_counts(normalize=True)
# the top 3 dangerous facilities are (1,3,7) i.e (Mainline roadway, Tunnel, Simple Ramp)
ax = road_data.facility.value_counts().plot(kind='bar')
ax.set_xlabel('Road Type (1:Mainline roadway, 3:Tunnel, 5:Ramp to Tunnel, 7:Simple Ramp)')
ax.set_ylabel('Road count in Boston')
print road_data.facility.value_counts(normalize=True)
# Tunnels are 1.9% of all roads but , 9.5% of all crashes happened in tunnels. Maybe tunnels have high AADT?
# Average tunnel AADT vs Average total road AADT
total_tunnels = len(road_data[road_data['facility'] == 3])
tunnels_with_aadt = len(road_data[ (road_data['facility'] == 3) & (road_data['aadt'] != 0)])
total_aadt = sum(road_data[(road_data['facility'] == 3) & (road_data['aadt'] != 0)].aadt.tolist())
average_aadt_in_tunnels = total_aadt/tunnels_with_aadt
print average_aadt_in_tunnels
# Average aadt in all roads
total_roads = len(road_data)
roads_with_aadt = len(road_data[road_data['aadt'] > 1000])
total_aadt_roads = sum(road_data[road_data['aadt'] > 1000].aadt.tolist())
average_aadt = total_aadt_roads/roads_with_aadt
print average_aadt
# Total fatal crashes
import numpy as np
injury_road_data = crash_road_data[(crash_road_data['total_fatal_injuries'] > 0) | (crash_road_data['total_nonfatal_injuries'] > 0)]
len(injury_road_data)
grouped_facility = crash_road_data.groupby('facility')
print grouped_facility['total_fatal_injuries'].agg(np.sum)
print grouped_facility['total_nonfatal_injuries'].agg(np.sum)
#number of fatal crashes vs facility type
ax = injury_road_data.facility.value_counts().plot(kind='bar')
ax.set_xlabel('Road Type (1:Mainline roadway, 3:Tunnel, 5:Ramp to Tunnel, 7:Simple Ramp)')
print injury_road_data.facility.value_counts(normalize=True)
# Tunnels are 1.9% of all roads and cause 6.6 percent of all injury crashes
#number of crashes vs median type
ax = crash_road_data.med_type.value_counts().plot(kind='bar')
ax.set_xlabel('Median Type')
ax.set_ylabel('Crash incident count')
print crash_road_data.med_type.value_counts(normalize=True)
ax = road_data.med_type.value_counts().plot(kind='bar')
ax.set_xlabel('Median Type')
ax.set_ylabel('Roads count')
print road_data.med_type.value_counts(normalize=True)
#High crash streets
ax = crash_road_data.st_name.value_counts().iloc[1:20].plot(kind='bar')
ax.set_xlabel('Street names')
ax.set_ylabel('Crash incident counts')
#High crash route ids
ax = crash_road_data.route_id.value_counts().iloc[1:20].plot(kind='bar')
ax.set_xlabel('Route ids')
ax.set_ylabel('Crash incident counts')
# number of crashes at intersection vs segments
intersection_crashes = crash_road_data[crash_road_data['at_roadway_intersection'] != 'NA']
intersection_crashes['intersection_road1'], intersection_crashes['intersection_road2'] = intersection_crashes['at_roadway_intersection'].str.split(' / ',1).str
# 3391 crashes out of 11920 crashes happened at intersections ~ 28% of crashes happenned at intersections
intersection_crashes.to_csv("intersection_crashes.csv")
print "Intersection "
print "Total incidents : ", len(intersection_crashes)
print "Total nonfatal injuries : ", sum(intersection_crashes['total_nonfatal_injuries'].tolist())
print "Total fatal injuries : ", sum(intersection_crashes['total_fatal_injuries'].tolist())
print "All roads "
print "Total incidents : " , len(crash_road_data)
print "Total nonfatal injuries : ", sum(crash_road_data['total_nonfatal_injuries'].tolist())
print "Total fatal injuries : ", sum(crash_road_data['total_fatal_injuries'].tolist())
#High crash intersections
ax = intersection_crashes.at_roadway_intersection.value_counts().iloc[1:20].plot(kind='bar')
ax.set_xlabel('Intersection name')
ax.set_ylabel('Number of crash incidents')
grouped_intersections = intersection_crashes.groupby('at_roadway_intersection')
intersection_injuries = grouped_intersections['total_nonfatal_injuries'].agg(np.sum).reset_index()
print intersection_injuries.sort('total_nonfatal_injuries', ascending=False)[:20]
```
| github_jupyter |
## Structured and Time Series Data
```
%matplotlib inline
%reload_ext autoreload
%autoreload 2
from fastai.structured import *
from fastai.column_data import *
np.set_printoptions(threshold=50, edgeitems=20)
PATH='data/rossmann/'
!ls {PATH}
```
### Create datasets
In addition to the provided data, we will be using external datasets put together by participants in the Kaggle competition. You can download all of them here.
For completeness, the implementation used to put them together is included below.
```
def concat_csvs(dirname):
path = f'{PATH}{dirname}'
filenames=glob(f"{PATH}/*.csv")
wrote_header = False
with open(f"{path}.csv","w") as outputfile:
for filename in filenames:
name = filename.split(".")[0]
with open(filename) as f:
line = f.readline()
if not wrote_header:
wrote_header = True
outputfile.write("file,"+line)
for line in f:
outputfile.write(name + "," + line)
outputfile.write("\n")
# concat_csvs('googletrend')
# concat_csvs('weather')
```
Feature Space:
* train: Training set provided by competition
* store: List of stores
* store_states: mapping of store to the German state they are in
* List of German state names
* googletrend: trend of certain google keywords over time, found by users to correlate well w/ given data
* weather: weather
* test: testing set
```
table_names = ['train', 'store', 'store_states', 'state_names',
'googletrend', 'weather', 'test']
```
We'll be using the popular data manipulation framework `pandas`. Among other things, pandas allows you to manipulate tables/data frames in python as one would in a database.
We're going to go ahead and load all of our csv's as dataframes into the list `tables`.
```
tables = [pd.read_csv(f'{PATH}{fname}.csv', low_memory=False) for fname in table_names]
from IPython.display import HTML, display
```
We can use `head()` to get a quick look at the contents of each table:
* train: Contains store information on a daily basis, tracks things like sales, customers, whether that day was a holdiay, etc.
* store: general info about the store including competition, etc.
* store_states: maps store to state it is in
* state_names: Maps state abbreviations to names
* googletrend: trend data for particular week/state
* weather: weather conditions for each state
* test: Same as training table, w/o sales and customers
```
for t in tables: display(t.head())
```
This is very representative of a typical industry dataset.
The following returns summarized aggregate information to each table accross each field.
```
for t in tables: display(DataFrameSummary(t).summary())
```
| github_jupyter |
# Regression modeling
A general, primary goal of many statistical data analysis tasks is to relate the influence of one variable on another. For example, we may wish to know how different medical interventions influence the incidence or duration of disease, or perhaps a how baseball player's performance varies as a function of age.
```
%matplotlib inline
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
sns.set()
from scipy.optimize import fmin
x = np.array([2.2, 4.3, 5.1, 5.8, 6.4, 8.0])
y = np.array([0.4, 10.1, 14.0, 10.9, 15.4, 18.5])
plt.plot(x,y,'ro')
```
We can build a model to characterize the relationship between $X$ and $Y$, recognizing that additional factors other than $X$ (the ones we have measured or are interested in) may influence the response variable $Y$.
<div style="font-size: 150%;">
$y_i = f(x_i) + \epsilon_i$
</div>
where $f$ is some function, for example a linear function:
<div style="font-size: 150%;">
$y_i = \beta_0 + \beta_1 x_i + \epsilon_i$
</div>
and $\epsilon_i$ accounts for the difference between the observed response $y_i$ and its prediction from the model $\hat{y_i} = \beta_0 + \beta_1 x_i$. This is sometimes referred to as **process uncertainty**.
We would like to select $\beta_0, \beta_1$ so that the difference between the predictions and the observations is zero, but this is not usually possible. Instead, we choose a reasonable criterion: ***the smallest sum of the squared differences between $\hat{y}$ and $y$***.
<div style="font-size: 120%;">
$$R^2 = \sum_i (y_i - [\beta_0 + \beta_1 x_i])^2 = \sum_i \epsilon_i^2 $$
</div>
Squaring serves two purposes: (1) to prevent positive and negative values from cancelling each other out and (2) to strongly penalize large deviations. Whether the latter is a good thing or not depends on the goals of the analysis.
In other words, we will select the parameters that minimize the squared error of the model.
```
sum_of_squares = lambda theta, x, y: np.sum((y - theta[0] - theta[1]*x) ** 2)
sum_of_squares([0,1],x,y)
b0,b1 = fmin(sum_of_squares, [0,1], args=(x,y))
b0,b1
plt.plot(x, y, 'ro')
plt.plot([0,10], [b0, b0+b1*10])
plt.plot(x, y, 'ro')
plt.plot([0,10], [b0, b0+b1*10])
for xi, yi in zip(x,y):
plt.plot([xi]*2, [yi, b0+b1*xi], 'k:')
plt.xlim(2, 9); plt.ylim(0, 20)
```
Minimizing the sum of squares is not the only criterion we can use; it is just a very popular (and successful) one. For example, we can try to minimize the sum of absolute differences:
```
sum_of_absval = lambda theta, x, y: np.sum(np.abs(y - theta[0] - theta[1]*x))
b0,b1 = fmin(sum_of_absval, [0,1], args=(x,y))
print('\nintercept: {0:.2}, slope: {1:.2}'.format(b0,b1))
plt.plot(x, y, 'ro')
plt.plot([0,10], [b0, b0+b1*10])
```
We are not restricted to a straight-line regression model; we can represent a curved relationship between our variables by introducing **polynomial** terms. For example, a cubic model:
<div style="font-size: 150%;">
$y_i = \beta_0 + \beta_1 x_i + \beta_2 x_i^2 + \epsilon_i$
</div>
```
sum_squares_quad = lambda theta, x, y: np.sum((y - theta[0] - theta[1]*x - theta[2]*(x**2)) ** 2)
b0,b1,b2 = fmin(sum_squares_quad, [1,1,-1], args=(x,y))
print('\nintercept: {0:.2}, x: {1:.2}, x2: {2:.2}'.format(b0,b1,b2))
plt.plot(x, y, 'ro')
xvals = np.linspace(0, 10, 100)
plt.plot(xvals, b0 + b1*xvals + b2*(xvals**2))
```
Although polynomial model characterizes a nonlinear relationship, it is a linear problem in terms of estimation. That is, the regression model $f(y | x)$ is linear in the parameters.
For some data, it may be reasonable to consider polynomials of order>2. For example, consider the relationship between the number of home runs a baseball player hits and the number of runs batted in (RBI) they accumulate; clearly, the relationship is positive, but we may not expect a linear relationship.
```
sum_squares_cubic = lambda theta, x, y: np.sum((y - theta[0] - theta[1]*x - theta[2]*(x**2)
- theta[3]*(x**3)) ** 2)
bb = pd.read_csv("../data/baseball.csv", index_col=0)
plt.plot(bb.hr, bb.rbi, 'r.')
b0,b1,b2,b3 = fmin(sum_squares_cubic, [0,1,-1,0], args=(bb.hr, bb.rbi))
xvals = np.arange(40)
plt.plot(xvals, b0 + b1*xvals + b2*(xvals**2) + b3*(xvals**3))
```
### Exercise: Polynomial function
Write a function that specifies a polynomial of arbitrary degree.
```
# Write your answer here
```
In practice, we need not fit least squares models by hand because they are implemented generally in packages such as [`scikit-learn`](http://scikit-learn.org/) and [`statsmodels`](https://github.com/statsmodels/statsmodels/). For example, `scikit-learn` package implements least squares models in its `LinearRegression` class:
```
from sklearn import linear_model
straight_line = linear_model.LinearRegression()
straight_line.fit(x.reshape(-1, 1), y)
straight_line.coef_
plt.plot(x, y, 'ro')
plt.plot(x, straight_line.predict(x[:, np.newaxis]), color='blue',
linewidth=3)
```
For more general regression model building, its helpful to use a tool for describing statistical models, called `patsy`. With `patsy`, it is easy to specify the desired combinations of variables for any particular analysis, using an "R-like" syntax. `patsy` parses the formula string, and uses it to construct the approriate *design matrix* for the model.
For example, the quadratic model specified by hand above can be coded as:
```
from patsy import dmatrix
X = dmatrix('x + I(x**2)')
np.asarray(X)
```
The `dmatrix` function returns the design matrix, which can be passed directly to the `LinearRegression` fitting method.
```
poly_line = linear_model.LinearRegression(fit_intercept=False)
poly_line.fit(X, y)
poly_line.coef_
plt.plot(x, y, 'ro')
plt.plot(x, poly_line.predict(X), color='blue',
linewidth=3)
```
## Model Selection
How do we choose among competing models for a given dataset? More parameters are not necessarily better, from the standpoint of model fit. For example, fitting a 9-th order polynomial to the sample data from the above example certainly results in an overfit.
```
def calc_poly(params, data):
x = np.c_[[data**i for i in range(len(params))]]
return np.dot(params, x)
sum_squares_poly = lambda theta, x, y: np.sum((y - calc_poly(theta, x)) ** 2)
betas = fmin(sum_squares_poly, np.zeros(10), args=(x,y), maxiter=1e6)
plt.plot(x, y, 'ro')
xvals = np.linspace(0, max(x), 100)
plt.plot(xvals, calc_poly(betas, xvals))
```
One approach is to use an information-theoretic criterion to select the most appropriate model. For example **Akaike's Information Criterion (AIC)** balances the fit of the model (in terms of the likelihood) with the number of parameters required to achieve that fit. We can easily calculate AIC as:
$$AIC = n \log(\hat{\sigma}^2) + 2p$$
where $p$ is the number of parameters in the model and $\hat{\sigma}^2 = RSS/(n-p-1)$.
Notice that as the number of parameters increase, the residual sum of squares goes down, but the second term (a penalty) increases.
To apply AIC to model selection, we choose the model that has the **lowest** AIC value.
```
n = len(x)
aic = lambda rss, p, n: n * np.log(rss/(n-p-1)) + 2*p
RSS1 = sum_of_squares(fmin(sum_of_squares, [0,1], args=(x,y)), x, y)
RSS2 = sum_squares_quad(fmin(sum_squares_quad, [1,1,-1], args=(x,y)), x, y)
print('\nModel 1: {0}\nModel 2: {1}'.format(aic(RSS1, 2, n), aic(RSS2, 3, n)))
```
Hence, on the basis of "information distance", we would select the 2-parameter (linear) model.
## Logistic Regression
Fitting a line to the relationship between two variables using the least squares approach is sensible when the variable we are trying to predict is continuous, but what about when the data are dichotomous?
- male/female
- pass/fail
- died/survived
Let's consider the problem of predicting survival in the Titanic disaster, based on our available information. For example, lets say that we want to predict survival as a function of the fare paid for the journey.
```
titanic = pd.read_excel("../data/titanic.xls", "titanic")
titanic.name
jitter = np.random.normal(scale=0.02, size=len(titanic))
plt.scatter(np.log(titanic.fare), titanic.survived + jitter, alpha=0.3)
plt.yticks([0,1])
plt.ylabel("survived")
plt.xlabel("log(fare)")
```
I have added random jitter on the y-axis to help visualize the density of the points, and have plotted fare on the log scale.
Clearly, fitting a line through this data makes little sense, for several reasons. First, for most values of the predictor variable, the line would predict values that are not zero or one. Second, it would seem odd to choose least squares (or similar) as a criterion for selecting the best line.
```
x = np.log(titanic.fare[titanic.fare>0])
y = titanic.survived[titanic.fare>0]
betas_titanic = fmin(sum_of_squares, [1,1], args=(x,y))
jitter = np.random.normal(scale=0.02, size=len(titanic))
plt.scatter(np.log(titanic.fare), titanic.survived + jitter, alpha=0.3)
plt.yticks([0,1])
plt.ylabel("survived")
plt.xlabel("log(fare)")
plt.plot([0,7], [betas_titanic[0], betas_titanic[0] + betas_titanic[1]*7.])
```
If we look at this data, we can see that for most values of `fare`, there are some individuals that survived and some that did not. However, notice that the cloud of points is denser on the "survived" (`y=1`) side for larger values of fare than on the "died" (`y=0`) side.
### Stochastic model
Rather than model the binary outcome explicitly, it makes sense instead to model the *probability* of death or survival in a **stochastic** model. Probabilities are measured on a continuous [0,1] scale, which may be more amenable for prediction using a regression line. We need to consider a different probability model for this exerciese however; let's consider the **Bernoulli** distribution as a generative model for our data:
<div style="font-size: 120%;">
$$f(y|p) = p^{y} (1-p)^{1-y}$$
</div>
where $y = \{0,1\}$ and $p \in [0,1]$. So, this model predicts whether $y$ is zero or one as a function of the probability $p$. Notice that when $y=1$, the $1-p$ term disappears, and when $y=0$, the $p$ term disappears.
So, the model we want to fit should look something like this:
<div style="font-size: 120%;">
$$p_i = \beta_0 + \beta_1 x_i + \epsilon_i$$
</div>
However, since $p$ is constrained to be between zero and one, it is easy to see where a linear (or polynomial) model might predict values outside of this range. We can modify this model sligtly by using a **link function** to transform the probability to have an unbounded range on a new scale. Specifically, we can use a **logit transformation** as our link function:
<div style="font-size: 120%;">
$$\text{logit}(p) = \log\left[\frac{p}{1-p}\right] = x$$
</div>
Here's a plot of $p/(1-p)$
```
logit = lambda p: np.log(p/(1.-p))
unit_interval = np.linspace(0,1)
plt.plot(unit_interval/(1-unit_interval), unit_interval)
```
And here's the logit function:
```
plt.plot(logit(unit_interval), unit_interval)
```
The inverse of the logit transformation is:
<div style="font-size: 150%;">
$$p = \frac{1}{1 + \exp(-x)}$$
</div>
```
invlogit = lambda x: 1. / (1 + np.exp(-x))
```
So, now our model is:
<div style="font-size: 120%;">
$$\text{logit}(p_i) = \beta_0 + \beta_1 x_i + \epsilon_i$$
</div>
We can fit this model using maximum likelihood. Our likelihood, again based on the Bernoulli model is:
<div style="font-size: 120%;">
$$L(y|p) = \prod_{i=1}^n p_i^{y_i} (1-p_i)^{1-y_i}$$
</div>
which, on the log scale is:
<div style="font-size: 120%;">
$$l(y|p) = \sum_{i=1}^n y_i \log(p_i) + (1-y_i)\log(1-p_i)$$
</div>
We can easily implement this in Python, keeping in mind that `fmin` minimizes, rather than maximizes functions:
```
def logistic_like(theta, x, y):
p = invlogit(theta[0] + theta[1] * x)
# Return negative of log-likelihood
return -np.sum(y * np.log(p) + (1-y) * np.log(1 - p))
```
Remove null values from variables
```
x, y = titanic[titanic.fare.notnull()][['fare', 'survived']].values.T
```
... and fit the model.
```
b0, b1 = fmin(logistic_like, [0.5,0], args=(x,y))
b0, b1
jitter = np.random.normal(scale=0.01, size=len(x))
plt.plot(x, y+jitter, 'r.', alpha=0.3)
plt.yticks([0,.25,.5,.75,1])
xvals = np.linspace(0, 600)
plt.plot(xvals, invlogit(b0+b1*xvals))
```
As with our least squares model, we can easily fit logistic regression models in `scikit-learn`, in this case using the `LogisticRegression`.
```
logistic = linear_model.LogisticRegression()
logistic.fit(x[:, np.newaxis], y)
logistic.coef_
```
### Exercise: multivariate logistic regression
Which other variables might be relevant for predicting the probability of surviving the Titanic? Generalize the model likelihood to include 2 or 3 other covariates from the dataset.
```
# Write your answer here
```
| github_jupyter |
```
%matplotlib inline
%load_ext line_profiler
import numpy as np
import os
import time
import keras
import pandas as pd
import matplotlib.pyplot as plt
from scipy.stats import logistic
from scipy.special import softmax
from IPython.display import display
from keras.layers import (Input, Dense, Lambda, Flatten, Reshape, BatchNormalization,
Activation, Dropout, Conv2D, Conv2DTranspose,
Concatenate, Add, Multiply)
from keras.engine import InputSpec
from keras.optimizers import Adam
from keras.models import Model
from keras.utils import to_categorical
from keras import metrics
from keras import backend as K
from keras_tqdm import TQDMNotebookCallback
from keras.datasets import cifar10
import tensorflow as tf
from pixelcnn_helpers import pixelcnn_loss, sigmoid, compute_pvals, compute_mixture, \
PixelConv2D, conv_block, resnet_block, final_block
```
# Parameters
```
img_rows, img_cols, img_chns = 32, 32, 3
original_img_size = (img_rows, img_cols, img_chns)
# Debug params
num_sample = 50000
# Hyper params
batch_size = int(os.environ.get('BATCH_SIZE', 16))
epochs = int(os.environ.get('EPOCHS', 1000))
learning_rate = float(os.environ.get('LEARNING_RATE', 0.002))
mixture_components = int(os.environ.get('MIXTURE_COMPONENTS', 5))
if K.image_data_format() == 'channels_last':
bn_axis = 3
else:
bn_axis = 1
```
# Extract Test Image
```
(X_train, y_train), (X_test, y_test) = cifar10.load_data()
X_train = X_train[:num_sample, :img_rows, :img_cols, :img_chns].astype(float)
X_train_orig = X_train
y_train = y_train[:num_sample]
print(X_train.shape)
print(X_test.shape)
# Shift/scale to [-1, 1] interval
X_train = (X_train - 127.5) / 127.5
X_test = (X_test - 127.5) / 127.5
y_train = to_categorical(y_train, num_classes=10)
y_test = to_categorical(y_test, num_classes=10)
print(X_train.shape, y_train.shape)
print(X_test.shape, y_test.shape)
```
# Model
```
def build_outputs(input_tensor, final_filters):
m_outs = []
invs_outs = []
weights = []
x = input_tensor
for t in ['rb', 'gb', 'bb']:
# Only allow things behind it...
decoder_out_m = PixelConv2D(t, name='x_m' + str(t),
filters=mixture_components,
kernel_size=1,
strides=1)(x)
decoder_out_invs = PixelConv2D(t, name='x_s' + str(t),
filters=mixture_components,
kernel_size=1,
strides=1,
activation='softplus')(x)
mixture_weights = PixelConv2D(t, name='weights' + str(t),
filters=mixture_components,
kernel_size=1,
strides=1)(x)
m_outs.append(decoder_out_m)
invs_outs.append(decoder_out_invs)
weights.append(mixture_weights)
out_m = Concatenate()(m_outs)
out_invs = Concatenate()(invs_outs)
out_mixture = Concatenate()(weights)
return Concatenate()([out_m, out_invs, out_mixture])
# Work around Keras/tensorboard bug: https://github.com/keras-team/keras/issues/10074
K.clear_session()
print("Building conv layers...")
main_input = Input(shape=original_img_size, name='main_input')
# Same architecture as "PixelRNN" paper (except for the outputs/loss, which are from the PixelCNN++ paper)
x = conv_block(main_input, 256, (5, 5), name='conv1', is_first=True)
resnet_depth = 5
for stage in range(resnet_depth):
x = resnet_block(x, [128, 128, 256], stage, 'a', kernel=3)
resnet_depth = 5
for stage in range(resnet_depth):
x = resnet_block(x, [128, 128, 256], stage + 5, 'a', kernel=3)
resnet_depth = 5
for stage in range(resnet_depth):
x = resnet_block(x, [128, 128, 256], stage + 10, 'a', kernel=3)
final_width = 1024
x = final_block(x, final_width, 256, '1x1_1000_1')
x = final_block(x, final_width, final_width, '1x1_1000_2')
print("Building output layers...")
outputs = build_outputs(x, final_width)
main_output = outputs
print("Building model...")
model = Model(inputs=main_input, outputs=main_output)
optimizer = Adam(lr=learning_rate)
model.compile(optimizer=optimizer,
loss=lambda x, y: pixelcnn_loss(x, y, img_rows, img_cols, img_chns, mixture_components))
model.summary()
early_stopping = keras.callbacks.EarlyStopping('val_loss', min_delta=50.0, patience=5)
reduce_lr = keras.callbacks.ReduceLROnPlateau(monitor='val_loss', factor=0.5, patience=2, min_lr=0.0001)
history = model.fit(
X_train, X_train,
batch_size=batch_size,
epochs=50,
callbacks=[TQDMNotebookCallback(), early_stopping, reduce_lr],
validation_data=(X_test, X_test),
verbose=0
)
df = pd.DataFrame(history.history)
display(df.describe(percentiles=[0.25 * i for i in range(4)] + [0.95, 0.99]))
col = 'val_loss' if 'val_loss' in df else 'loss'
df[col][-25:].plot(figsize=(8, 6))
debug = False
def gen_image(model, num_samples=batch_size):
x_sample = np.zeros((num_samples, img_rows, img_cols, img_chns))
# Iteratively generate each conditional pixel P(x_i | x_{1,..,i-1})
for i in range(img_rows):
for j in range(img_cols):
for k in range(img_chns):
# =======================================================
#x_out = model.predict(X_train, num_samples)
x_out = model.predict(x_sample, num_samples)
for n in range(num_samples):
offset = k * mixture_components
x_ms = x_out[n, i, j, offset:offset + mixture_components]
offset = mixture_components * img_chns + k * mixture_components
x_invs = x_out[n, i, j, offset:offset + mixture_components]
offset = 2 * mixture_components * img_chns + k * mixture_components
weights = softmax(x_out[n, i, j, offset:offset + mixture_components])
pvals = compute_mixture(x_ms, x_invs, weights, mixture_components)
pvals /= (np.sum(pvals) + 1e-5)
pixel_val = np.argmax(np.random.multinomial(1, pvals))
x_sample[n, i, j, k] = (pixel_val - 127.5) / 127.5
if 0 <= i <= 3 and 0 <= j <= 3 and n == 0 and debug:
print("====", i, j, k)
print("X_train[0/1", i, j, k, "] = ", X_train[0, i, j, k], ', ', X_train[1, i, j, k],
" (%s, %s)" % (X_train[0, i, j, k] * 127.5 + 127.5, X_train[1, i, j, k] * 127.5 + 127.5))
print(" m: ", x_ms, "(%s)" % (x_ms * 127.5 + 127.5))
print(" E[m]: ", (x_ms * weights).sum(), "(%s)" % (x_ms * 127.5 + 127.5))
print(" invs: ", x_invs)
print(" weights: ", weights, " (", x_out[n, i, j, offset:offset + mixture_components] ,")")
s = pd.Series(pvals)
print(" pvals: ", s[s>1e-2])
print(" pixel_val: ", pixel_val)
samples = pd.Series(np.random.choice(len(pvals), 1000, p=pvals / pvals.sum()))
samples.hist(bins=128, alpha=0.5, label='generated')
print("row", i)
return (x_sample * 127.5 + 127.5)
start = time.time()
n = 1 if debug else batch_size
num_samples = min(n, batch_size)
rows = int(n // np.ceil(np.sqrt(num_samples)))
cols = int(n // np.ceil(np.sqrt(num_samples)))
figure = np.zeros((img_rows * rows, img_cols * cols, img_chns))
print(figure.shape)
samples = gen_image(model, num_samples=num_samples)
for i in range(rows):
for j in range(cols):
img = samples[i * rows + j] / 255.
d_x = (i * img_rows)
d_y = (j * img_cols)
figure[d_x:d_x + img_rows, d_y:d_y + img_cols, :] = img
done = time.time()
elapsed = done - start
print("Elapsed: ", elapsed)
plt.figure(figsize=(12, 12))
plt.imshow(figure)
plt.show()
```
| github_jupyter |
# Introduction
The goal of this document is to construct a computer from logical gates. The basic logical gates that are used to construct the computer are: `And`, `Or`, and `Not`. The major outline of this document is followed from the book _The Elements of Computing Systems, Noam Nisan and Shimon Shocken (2008)_.
The book is followed from Boolean Logic (chapter 1) to Assembler (chapter 6). There are a total of 13 chapters. The construction of the Virtual Machine, and eventually an Operating System, is left out of scope. The main reason for this is because Python does not allow an easy 'screen' interface. The entire emulator will be written as a `REPL` application, which allows the user excessive control of the entire emulator. The cost of this, however, is that everything is done at a very low level.
The goal is to create a computer that can run Assembly code, and that we have an assembler. I haven't followed the Assembly implementation in the book, instead, I used it as a baseline, and transformed it into my own preferences.
Website's that can be useful for further understanding:
* http://fourier.eng.hmc.edu/e85_old/lectures/digital_logic/node6.html
# Boolean logic
Every Boolean function, no matter how complex, can be expressed using three Boolean operators only: `And`, `Or`, and `Not`.
## Gate logic
A _gate_ is a physical device that implements a Boolean function. If a Boolean function $f$ operates on $n$ variables and returns $m$ binary results, the gate that implements $f$ will have _$n$ input pins and $m$ output pins_. Today, most gates are implemented as transistors etched in silicon, packaged as _chips_.
There are the following three _primitive gates_, which will be used to create all the other components. The gates are: `And`, `Or`, and `Not`. The following symbols are used to indicate each gate:

We can define their Boolean functions mathematically as:
1. $\text{And}(a,b)=a\land b$
2. $\text{Or}(a,b)=a\lor b$
3. $\text{Not}(a)=\lnot a$
For ease of writing, we will also adopt the convention to write a $1$ when the value is `True`, and $0$ if the value is `False`.
## Primitive gates
We will now implement the primitive logic gates in Python. The input variables can be either a Boolean value, or a function. The input will be called by the `boolf(x)` function, which returns a Boolean value, or executes the function. Allowing functions gives us the option to easily chain multiple gates together with references.
```
def boolf(x):
if callable(x): return x()
if type(x) is bool: return x
raise ValueError('Unsupported type {} for boolf()'.format(type(x)))
class And():
def __init__(self, a, b): self.a = a; self.b = b
def out(self): return boolf(self.a) and boolf(self.b)
def __repr__(self): return '{}'.format(self.out())
class Or():
def __init__(self, a, b): self.a = a; self.b = b
def out(self): return boolf(self.a) or boolf(self.b)
def __repr__(self): return '{}'.format(self.out())
class Not():
def __init__(self, a): self.a = a
def out(self): return not boolf(self.a)
def __repr__(self): return '{}'.format(self.out())
```
To test a simple scenario, let $G(a,b,c) = \text{Not}(\text{Or}(a, \text{And}(b, c)))$. Because this is a composite gate, we are only using primitive gates. If $(a,b,c)=(0,1,1)$, the resulting output should be $0$.
```
a=False;b=c=True
G = Not(Or(lambda: a, And(lambda: b, lambda: c).out).out)
print(G)
b=False
print(G)
```
**Reference vs. value types**
Notice that primitive Boolean values should be the result of a function. If a Boolean value is the input for a gate, Python will reference to it by value, which is something we do not want. Thus, we input a function into the gate, and Python passes the function as a reference. Then we use `boolf(x)` to either get the variable, or return the function result. This allows us to change the input values, after the gate has been constructed, and keep calling `.out()` on the last gate to execute all the logic in the intermediate gates.
## Composite gates
We start with primitive gates and design more complicated functionality by interconnecting them, leading to the construction of _composite gates_.
### Three-way And gate
The first gate we can construct from two And gates, is a three-way And gate. It can be defined in the following way: $\text{And3W}(a,b,c)= \text{And}(\text{And}(a,b),c)$.

```
class And3W():
def __init__(self, a, b, c): self.a = a; self.b = b; self.c = c
def out(self):
and1 = And(boolf(self.a), boolf(self.b))
and2 = And(and1.out, boolf(self.c))
return and2.out()
def __repr__(self): return '{}'.format(self.out())
And3W(True, True, True)
```
### Xor gate
Let us consider another logic design example -- that of a `Xor` gate. The gate $\text{Xor}(a,b)$ is $1$ exactly when $a=1$ and $b=0$, or $a=0$ and $b=1$. In the case where $a=b$ the gate is $0$. It can be defined in the following way: $\text{Xor(a,b)}=\text{Or}(\text{And}(a,\text{Not}(b)), \text{And}(\text{Not}(a),b))$. This definition leads to the logic design shown in the figure below:
```
class Xor():
def __init__(self, a, b): self.a = a; self.b = b
def out(self): return Or(And(boolf(self.a), Not(boolf(self.b)).out).out,\
And(Not(boolf(self.a)).out, boolf(self.b)).out\
).out()
def __repr__(self): return '{}'.format(self.out())
Xor(False, True).out() and Xor(True, False).out() \
and not Xor(True, True).out() and not Xor(False, False).out()
```
## Basic Logic Gates
In this section we will define all the basic logical gates that are used as the building blocks to create more advanced circuits.
### Not
A not gate has a single input, and inverts that input as output. It is defined as $\text{Not}(a)=\lnot a$. The following truth table applies for a Not gate:
|in|out|
|--|--|
|0|1|
|1|0|
The gate has already been implemented earlier in _Primitive gates_.
### And
An And gate has two inputs $a, b$, and only outputs a $1$ if exacty $a=b=1$. It is defined as $\text{And}(a,b)=a \land b$. The following truth table applies for an And gate:
|a|b|out|
|--|--|--|
|0|0|0|
|0|1|0|
|1|0|0|
|1|1|1|
The gate has already been implemented earlier in _Primitive gates_.
### Or
An Or gate has two inputs $a, b$, and only outputs a $0$ if exacty $a=b=0$, otherwise it will output $1$. It is defined as $\text{Or}(a,b)=a \lor b$. The following truth table applies for an Or gate:
|a|b|out|
|--|--|--|
|0|0|0|
|0|1|1|
|1|0|1|
|1|1|1|
The gate has already been implemented earlier in _Primitive gates_.
### Xor
An Xor gate has two inputs $a,b$, its output is $1$ exactly when $a=1$ and $b=0$, or $a=0$ and $b=1$. Otherwise, the output is $0$. It is defined as $\text{Xor(a,b)}=\text{Or}(\text{And}(a,\text{Not}(b)), \text{And}(\text{Not}(a),b))$. The following thruth table applies for a Xor gate:
|a|b|out|
|--|--|--|
|0|0|0|
|0|1|1|
|1|0|1|
|1|1|0|
### Nand
A Nand gate has two input $a,b$ and its output is the inverse of an And gate. It's value is always $1$, except when $a=b=1$ for which the output is $0$. It is defined as $\text{Nand}(a,b)=\text{Not}(\text{And}(a,b))$. The following truth table applies for a Nand gate:
|a|b|out|
|--|--|--|
|0|0|1|
|0|1|1|
|1|0|1|
|1|1|0|
In Python we can implement the gate in the following way:
```
class Nand():
def __init__(self,a,b): self.a=a; self.b=b
def out(self): return Not(And(boolf(self.a), boolf(self.b)).out).out()
def __repr__(self): return '{}'.format(self.out())
Nand(False, False).out() and Nand(True, False).out()\
and Nand(False, True).out() and not Nand(True, True).out()
```
### Multiplexor
A multiplexor is a three-input gate that uses one of the inputs, called _selection bit_, to select and output one of the other two inputs, called _data bits_. It is defined as $\text{Mux}(a,b,sel)=\text{Or}(\text{And}(a, \text{Not}(sel), \text{And}(b,sel))$. The following truth table applies for a multiplexor:
|a|b|sel|out|
|--|--|--|--|
|0|0|0|0|
|0|1|0|0|
|1|0|0|1|
|1|1|0|1|
|0|0|1|0|
|0|1|1|1|
|1|0|1|0|
|1|1|1|1|
Which can also be expressed in a more simple form, such as:
|sel|out|
|--|--|
|0|a|
|1|b|
Using the definition, we can implement it in the following way:
```
class Mux():
def __init__(self,a,b,sel): self.a=a; self.b=b; self.sel=sel
def out(self): return Or(And(boolf(self.a), Not(boolf(self.sel)).out).out,\
And(boolf(self.b), boolf(self.sel)).out
).out()
def __repr__(self): return '{}'.format(self.out())
Mux(True, False, False).out() and Mux(False, True, True).out()\
and not Mux(False, False, False).out()
```
### Demultiplexor
A demultiplexor performs the opposite function of a multiplexor: It takes a single input and channels it to one of two possible outputs according to a selector bit that specifies which output to choose. It is defined for each output bit as:
$$ \text{DMux}(in,sel) = \begin{cases}\begin{aligned} \text{And}(in,\text{Not}(sel)) \quad &\text{output bit } a \\ \text{And}(in,sel) \quad &\text{output bit } b \end{aligned}\end{cases} $$
It can be represented with the following table:
|sel|a|b|
|--|--|--|
|0|in|0|
|1|0|in|
Using the definition, it can be implement in Python in the following way. Notice that $z=in$ because `in` is a reserved keyword in Python.
```
class DMux():
def __init__(self, z, sel): self.z=z; self.sel=sel;
def a(self): return And(boolf(self.z), Not(boolf(self.sel)).out).out()
def b(self): return And(boolf(self.z), boolf(self.sel)).out()
def __repr__(self): return 'a: {}, b: {}'.format(self.a(), self.b())
DMux(True, False)
DMux(True, True)
```
## Multi-Bit Versions of Basic Gates
Computer hardware is typically designed to operate on multi-bit arrays called "buses". For example, a basic requirement of a 32-bit computer is to be able to compute (bit-wise) an And function of two given 32-bit buses. To implement this operation, we can build an array of 32 binary And gates, each operating seperately on a pair of bits. In order to enclose all this logic in one package, we can encapsulate the gates array in a single chip interface consistring of two 32-bit input buses and one 32-bit output bus.
This section describes a typical set of such multi-bit logic gates, as needed for the construction of a typical 16-bit computer. We note in passing that the architecture of _n_-bit logic gates is basically the same irrespective of _n_'s values.
Also, we will adopt the notation to refer to the individual bits. For example, to refer to individual bits in a 16-bit bus named `data`, we use the notation `data[0], data[1], ..., data[15]`.
We will also define a helper function `bstr(L)`, that transforms a list of Boolean values into a binary string. For example, the list `[True, True, False]` gives the binary string: `110`. This is useful because it gives a compact representation of each individual bit.
```
def bstr(L):
out='';
for i in L: out += '1' if i else '0'
return out
bstr([False,True,False,False,False,True]+10*[False])
```
### Not16
```
class Not16():
def __init__(self, in16): self.in16=in16
def out(self): return [not self.in16[i] for i in range(16)]
def __repr__(self): return bstr(self.out())
Not16([False] * 16)
```
### And16
```
class And16():
def __init__(self, a, b): self.a=a; self.b=b
def out(self): return [self.a[i] and self.b[i] for i in range(16)]
def __repr__(self): return bstr(self.out())
And16([True]*16, [False]*15+[True])
```
### Or16
```
class Or16():
def __init__(self, a, b): self.a=a; self.b=b
def out(self): return [self.a[i] or self.b[i] for i in range(16)]
def __repr__(self): return bstr(self.out())
Or16([False]+15*[True], [False]*15+[True])
bits=16
x = [False]*bits
y = [False]*bits
x[1]=y[1]=True
F = And16(Or16(x,y).out(), y)
print(bstr(F.a))
x[2]=y[2]=True
#Required to manually update a, b (Boooh!)
F.a=x;F.b=y;
print(bstr(F.a))
a=x
print(bstr(a))
# Now we have reference types, because a and x are lists.
# Lists are classes, and those are passed on by reference.
x[2]=True
print(bstr(a))
```
### Mux16
## Multi-Way Versions of Basic Gates
### Or8Way
### Multi-Way/Multi-Bit Multiplexor
#### Mux4Way16
#### Mux8Way16
#### DMux4Way
#### DMux8Way
# Boolean Arithmetic
# Sequential Logic
# Machine Language
# Computer Architecture
# Assembler
| github_jupyter |
# Classifying movie reviews [Binary classification]
## Loading the IMDB dataset
We'll be working with "IMDB dataset", a set of 50,000 highly-polarized reviews from the Internet Movie Database. We will only keep the top 10,000 most frequently occurring words in the training data. Rare words will be discarded.
```
from keras.datasets import imdb
(train_data, train_labels), (test_data, test_labels) = imdb.load_data(num_words=10000)
```
Let's have a closer look at the training data:
**train_data** and **test_data** are lists of reviews, each review being a list of word indices (encoding a sequence of words):
```
train_data[0]
```
**train_labels** and **test_labels** are lists of 0s and 1s, where 0 stands for "negative" and 1 stands for "positive":
```
train_labels[0]
```
For kicks, here's how you can quickly decode one of these reviews back to English words:
```
# word_index is a dictionary mapping words to an integer index
word_index = imdb.get_word_index()
# We reverse it, mapping integer indices to words
reverse_word_index = dict([(value, key) for (key, value) in word_index.items()])
# We decode the review; note that our indices were offset by 3
# because 0, 1 and 2 are reserved indices for "padding", "start of sequence", and "unknown".
decoded_review = ' '.join([reverse_word_index.get(i - 3, '?') for i in train_data[0]])
decoded_review
```
## Encoding integer sequences in a binary matrix
We cannot feed lists of integers into a neural network. We have to turn our lists into tensors.
We will one-hot-encode our lists to turn them into vectors of 0s and 1s turning the sequence [3, 5] into a 10,000-dimensional vector that would be all-zeros except for indices 3 and 5, which would be ones. Then we could use as first layer in our network a Dense layer, capable of handling floating point vector data.
```
import numpy as np
def vectorize_sequences(sequences, dimension=10000):
# Create an all-zero matrix of shape (len(sequences), dimension)
results = np.zeros((len(sequences), dimension))
for i, sequence in enumerate(sequences):
results[i, sequence] = 1. # set specific indices of results[i] to 1s
return results
# Our vectorized training data
x_train = vectorize_sequences(train_data)
# Our vectorized test data
x_test = vectorize_sequences(test_data)
```
Here's what our samples look like now:
```
x_train[0]
```
We will also vectorize our labels:
```
# Our vectorized labels
y_train = np.asarray(train_labels).astype('float32')
y_test = np.asarray(test_labels).astype('float32')
y_train[0]
```
## Model definition
The intermediate layers will use relu as their "activation function", and the final layer will use a sigmoid activation so as to output a probability (a score between 0 and 1, indicating how likely the sample is to have the target "1", i.e. how likely the review is to be positive). A **relu (rectified linear unit)** is a function meant to **zero-out negative values**, while a **sigmoid** "squashes" arbitrary values into the **[0, 1] interval**, thus outputting something that can be interpreted as a probability.
```
from keras import models
from keras import layers
model = models.Sequential()
model.add(layers.Dense(16, activation='relu', input_shape=(10000,)))
model.add(layers.Dense(16, activation='relu'))
model.add(layers.Dense(1, activation='sigmoid'))
```
## Compilling the model
Since we are facing a binary classification problem and the output of our network is a probability (we end our network with a single-unit layer with a sigmoid activation), is it best to use the **binary_crossentropy** loss. It isn't the only viable choice: you could use, for instance, mean_squared_error.
Crossentropy is a quantity from the field of Information Theory, that measures the "distance" between probability distributions, or in our case, between the ground-truth distribution and our predictions.
```
model.compile(optimizer='rmsprop',
loss='binary_crossentropy',
metrics=['accuracy'])
```
## Configuring the optimizer
We are passing our optimizer, loss function and metrics as strings, which is possible because rmsprop, binary_crossentropy and accuracy are packaged as part of Keras. Sometimes you may want to configure the parameters of your optimizer, or pass a custom loss function or metric function.
## Using custom losses and metrics
```
from keras import losses
from keras import metrics
from keras import optimizers
model.compile(optimizer=optimizers.RMSprop(lr=0.001),
loss=losses.binary_crossentropy,
metrics=[metrics.binary_accuracy])
```
## Setting aside a validation set
In order to monitor during training the accuracy of the model on data that it has never seen before, we will create a "validation set" by setting apart 10,000 samples from the original training data.
```
x_val = x_train[:10000]
partial_x_train = x_train[10000:]
y_val = y_train[:10000]
partial_y_train = y_train[10000:]
```
## Training the model
We will now train our model for 20 epochs (20 iterations over all samples in the x_train and y_train tensors), in mini-batches of 512 samples. At this same time we will monitor loss and accuracy on the 10,000 samples that we set apart. This is done by passing the validation data as the validation_data argument.
```
history = model.fit(partial_x_train,
partial_y_train,
epochs=20,
batch_size=512,
validation_data=(x_val, y_val))
```
Note that the call to model.fit() returns a **History object**. This object has a member history, which is a dictionary containing data about everything that happened during training. **It contains 4 entries**: one per metric that was being monitored, during training and during validation. Let's take a look at it:
```
history_dict = history.history
history_dict.keys()
```
## Plotting training and validation loss
Let's use Matplotlib to plot the training and validation loss side by side, as well as the training and validation accuracy.
```
import matplotlib.pyplot as plt
acc = history.history['binary_accuracy']
val_acc = history.history['val_binary_accuracy']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(1, len(acc) + 1)
# "bo" is for "blue dot"
plt.plot(epochs, loss, 'bo', label='Training loss')
# b is for "solid blue line"
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
```
## Plotting training and validation accuracy
The dots are the training loss and accuracy, while the solid lines are the validation loss and accuracy. Note that your own results may vary slightly due to a different random initialization of your network.
```
plt.clf() # clear figure
acc_values = history_dict['binary_accuracy']
val_acc_values = history_dict['val_binary_accuracy']
plt.plot(epochs, acc, 'bo', label='Training acc')
plt.plot(epochs, val_acc, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
```
## Retraining a new model from scratch
Let's train a new network from scratch for 4 epochs, then evaluate it on our test data.
```
model = models.Sequential()
model.add(layers.Dense(16, activation='relu', input_shape=(10000,)))
model.add(layers.Dense(16, activation='relu'))
model.add(layers.Dense(1, activation='sigmoid'))
model.compile(optimizer='rmsprop',
loss='binary_crossentropy',
metrics=['accuracy'])
model.fit(x_train, y_train, epochs=4, batch_size=512)
results = model.evaluate(x_test, y_test)
results
```
## Using a trained network to generate predictions on new data
After having trained a network, you will want to use it in a practical setting. You can generate the likelihood of reviews being positive by using the predict method.
```
model.predict(x_test)
```
As you can see, the network is very confident for some samples (0.99 or more, or 0.01 or less) but less confident for others (0.6, 0.4).
## Conclusion
As you can see, the training loss decreases with every epoch and the training accuracy increases with every epoch. **That's what you would expect when running gradient descent optimization -- the quantity you are trying to minimize should get lower with every iteration**.
But that isn't the case for the validation loss and accuracy: they seem to peak at the fourth epoch. This is an example of what we were warning against earlier: a model that performs better on the training data isn't necessarily a model that will do better on data it has never seen before.
In precise terms, what you are seeing is **"overfitting"**: after the second epoch, we are over-optimizing on the training data, and **we ended up learning representations that are specific to the training data and do not generalize to data outside of the training set**.
| github_jupyter |
# Mask R-CNN Demo
A quick intro to using the pre-trained model to detect and segment objects.
```
import os
import sys
import random
import math
import numpy as np
import skimage.io
import matplotlib
import matplotlib.pyplot as plt
# Root directory of the project
ROOT_DIR = os.path.abspath("../")
# Import Mask RCNN
sys.path.append(ROOT_DIR) # To find local version of the library
from mrcnn import utils
import mrcnn.model as modellib
from mrcnn import visualize
# Import COCO config
sys.path.append(os.path.join(ROOT_DIR, "samples/coco/")) # To find local version
import coco
%matplotlib inline
# Directory to save logs and trained model
MODEL_DIR = os.path.join(ROOT_DIR, "logs")
WEIGHTS_DIR = os.path.join(ROOT_DIR, "weights")
# Local path to trained weights file
COCO_MODEL_PATH = os.path.join(WEIGHTS_DIR, "ResNet-101", "mask_rcnn_coco.h5")
# Directory of images to run detection on
IMAGE_DIR = os.path.join(ROOT_DIR, "images")
```
## Configurations
We'll be using a model trained on the MS-COCO dataset. The configurations of this model are in the ```CocoConfig``` class in ```coco.py```.
For inferencing, modify the configurations a bit to fit the task. To do so, sub-class the ```CocoConfig``` class and override the attributes you need to change.
```
class InferenceConfig(coco.CocoConfig):
# Set batch size to 1 since we'll be running inference on
# one image at a time. Batch size = GPU_COUNT * IMAGES_PER_GPU
GPU_COUNT = 1
IMAGES_PER_GPU = 1
config = InferenceConfig()
config.display()
```
## Create Model and Load Trained Weights
```
# Create model object in inference mode.
model = modellib.MaskRCNN(mode="inference", model_dir=MODEL_DIR, config=config)
# Load weights trained on MS-COCO
model.load_weights(COCO_MODEL_PATH, by_name=True)
```
## Class Names
The model classifies objects and returns class IDs, which are integer value that identify each class. Some datasets assign integer values to their classes and some don't. For example, in the MS-COCO dataset, the 'person' class is 1 and 'teddy bear' is 88. The IDs are often sequential, but not always. The COCO dataset, for example, has classes associated with class IDs 70 and 72, but not 71.
To improve consistency, and to support training on data from multiple sources at the same time, our ```Dataset``` class assigns it's own sequential integer IDs to each class. For example, if you load the COCO dataset using our ```Dataset``` class, the 'person' class would get class ID = 1 (just like COCO) and the 'teddy bear' class is 78 (different from COCO). Keep that in mind when mapping class IDs to class names.
To get the list of class names, you'd load the dataset and then use the ```class_names``` property like this.
```
# Load COCO dataset
dataset = coco.CocoDataset()
dataset.load_coco(COCO_DIR, "train")
dataset.prepare()
# Print class names
print(dataset.class_names)
```
We don't want to require you to download the COCO dataset just to run this demo, so we're including the list of class names below. The index of the class name in the list represent its ID (first class is 0, second is 1, third is 2, ...etc.)
```
# COCO Class names
# Index of the class in the list is its ID. For example, to get ID of
# the teddy bear class, use: class_names.index('teddy bear')
class_names = ['BG', 'person', 'bicycle', 'car', 'motorcycle', 'airplane',
'bus', 'train', 'truck', 'boat', 'traffic light',
'fire hydrant', 'stop sign', 'parking meter', 'bench', 'bird',
'cat', 'dog', 'horse', 'sheep', 'cow', 'elephant', 'bear',
'zebra', 'giraffe', 'backpack', 'umbrella', 'handbag', 'tie',
'suitcase', 'frisbee', 'skis', 'snowboard', 'sports ball',
'kite', 'baseball bat', 'baseball glove', 'skateboard',
'surfboard', 'tennis racket', 'bottle', 'wine glass', 'cup',
'fork', 'knife', 'spoon', 'bowl', 'banana', 'apple',
'sandwich', 'orange', 'broccoli', 'carrot', 'hot dog', 'pizza',
'donut', 'cake', 'chair', 'couch', 'potted plant', 'bed',
'dining table', 'toilet', 'tv', 'laptop', 'mouse', 'remote',
'keyboard', 'cell phone', 'microwave', 'oven', 'toaster',
'sink', 'refrigerator', 'book', 'clock', 'vase', 'scissors',
'teddy bear', 'hair drier', 'toothbrush']
```
## Run Object Detection
```
# Load a random image from the images folder
file_names = next(os.walk(IMAGE_DIR))[2]
image = skimage.io.imread(os.path.join(IMAGE_DIR, random.choice(file_names)))
# Run detection
results = model.detect([image], verbose=1)
# Visualize results
r = results[0]
visualize.display_instances(image, r['rois'], r['masks'], r['class_ids'],
class_names, r['scores'])
```
| github_jupyter |
###### Copyright © Anand B Pillai, Anvetsu Technologies Pvt. Ltd (2015)
# Data Structures & Types
## 1. Mutable data structures as default arguments
### 1.1. Show me the Code!
```
def add_name(names=[]):
""" Add a name to a list """
name=raw_input("Enter name: ").strip()
names.append(name)
return names
print add_name()
print add_name()
print add_name()
```
## Gotcha !!!
#### This is because,
1. The function definition line is evaluated just once and not every time the function is called.
2. This is because Python functions are first class objects.
### 1.2. Show me the Fix !
#### 1.2.1. Use a place-holder value instead of modifying the default value directly.
```
def add_name(names=None):
if names==None:
names = []
name=raw_input("Enter name: ").strip()
names.append(name)
return names
print add_name()
print add_name()
names = ['Appu','Dhruv']
print add_name(names)
```
##### 1.2.2. Use a sentinel object.
```
sentinel = object()
def add_name(names=sentinel):
if names==sentinel:
names = []
name=raw_input("Enter name: ").strip()
names.append(name)
return names
names = ['Appu','Dhruv']
print add_name(names)
```
##### 1.2.3. Use an inner function, which is always evaluated from the context of the outer function.
```
def add_name():
def inner(names=[]):
name=raw_input("Enter name: ").strip()
names.append(name)
return names
return inner()
add_name()
add_name()
add_name()
```
#### 1.3. Valid uses of this behavior
##### 1.3.1. A caching memoizer pattern
```
def fibonacci(n, memo={}):
""" Return n'th fibonacci number """
# Uses an inline caching dictionary
# as a memoizing data structure
if n in memo:
print '*** memoized data ***'
return memo[n]
a, b, c = 0, 1, 1
for i in range(n-1):
c = a + b
a, b = b, c
memo[n] = c
return c
fibonacci(1)
fibonacci(2); fibonacci(2)
fibonacci(3); fibonacci(3)
```
#### 1.4. More Reading
1. https://stackoverflow.com/questions/1132941/least-astonishment-in-python-the-mutable-default-argument
1. http://effbot.org/zone/default-values.htm
## 2. Mutable Argument Modification / Name Binding
### 2.1. Show me the Code!
```
def f(x, l):
""" A function taking a list as argument """
# This is a silly function, really.
if len(l)<5: # L1
l = g(x) # L2
# L3
l.append(x) # L4
def g(x):
""" A functon """
return [x*x]*5
nums = range(5)
f(10, nums)
print nums
nums=range(3)
f(10, nums)
print nums # 'nums' remains the same. Not surprised ? Good, Surprised - Not so good :)
```
## Gotcha !!!
#### This is because,
1. The __nums__ that is replaced in line #2 is a __new__ object recieved from g(...). It doesn't
replace the original object.
2. This is because in Python objects are bound to variables by name. Names _refer_ to objects, they don't bind strongly to them.
3. In order to modify a mutable, you need to call methods on it that modifies it. In case of list, these are _append_, _extend_, _remove_, _pop_ etc.
### 2.2. Show me the Fix !
```
def f(x, l):
""" A function taking a list as argument """
if len(l)<5: # L1
l = g(x) # L2
# L3
l.append(x) # L4
# Return it
return l
def g(x):
""" A functon """
return ([x]*5)
nums=range(3)
nums = f(10, nums)
print nums
```
## 3. Immutable Variable Comparison
### 3.1. Show me the Code!
```
def greet(greeting, default_value="Hi"):
""" Greet someone with a greeting """
if greeting is not default_value:
greeting = default_value + ", " + greeting
print greeting
# Test 1
greet("Hi")
# Test 2
greet("Good Morning")
# Test 3
greet("Good Morning", "Hello there")
# Test 4
greet("Hello there", "Hello there") # Fine
# Test 5
greet("Hi, how do you do!", "Hi, how do you do!")
# Test 6
greeting="Hello there"
greet(greeting, default_value="Hello there") # Hmmm, not what you expected ?
```
## Gotcha !!!
#### This is because,
1. You used __is__, the identity comparison operator instead of __!=__, the equality comparison operator.
1. However, the code still works as expected in __Test 4__ above because Python optimizes string memory for literal strings. Since
both arguments are passed as literal strings and their value is the same, Python creates the object just once for both arguments,
so the _is_ comparison works.
1. In __Test 6__, we use a separate name _greeting_ for the first argument and the literal string for the second. Hence Python doesn't
get a chance to optimize in this case and the _is_ comparison fails.
### 3.2. Show me the Fix !
```
def greet(greeting, default_value="Hi"):
""" Greet someone with a greeting """
# Simple: Use == or != operator always
if greeting != default_value:
greeting = default_value + ", " + greeting
print greeting
# Test 4
greet("Hello there", "Hello there")
# Test 6
greeting="Hello there"
greet(greeting, default_value="Hello there")
```
## 4. Integer vs Float Division
#### Integer division in Python always produces an integer result, ignoring the fractional part. Moreover, it __floors__ the result which can sometimes be a little confusing.
### 4.1. Show me the Code!
```
5/2 # Not 2.5, but 2, i.e the answer rounded off
-5/2 # Prints -3, not -2, i.e answer is floored away from zero
```
### 4.2. Notes
This is pretty well known behaviour of Python. It is not exactly a Gotcha, but newbie programmers are caught off-guard when
they encounter this behaviour for the first time. It does take a while to get used to it.
### 4.3. Workarounds
#### 4.3.2 Workaround #1 - Specifically use float division
```
# Just remember to convert one of the numbers to float, typically multiplying by 1.0.
# This is what I do.
x=5
y=1.0*x/2
print y
x=5
y=x/2.0
print y
```
#### 4.3.2 Workaround #2 - Backported from future
```
from __future__ import division
print "True division =>", 5 / 2
print "Floor division =>", 5 // 2
```
For Python 2.x, import __division__ from the future (means a feature backported from Python 3.x). Then
you get two division operators, __/__ performing true division and the new __//__ performing floor division.
#### 4.3.3 Workaround #3 - Use decimal module
```
import decimal
x=decimal.Decimal(5)
y=decimal.Decimal(2)
z=x/y
print z
```
__NOTE__ - Above is overkill for such a simple example. __Decimal__ types are more useful to get absolute precision for your floating point numbers. We will see another example below.
#### 4.4. More Reading
1. http://python-history.blogspot.in/2010/08/why-pythons-integer-division-floors.html
1. https://stackoverflow.com/questions/183853/in-python-what-is-the-difference-between-and-when-used-for-division
## 5. Floating Point Precision & Round-Off
#### Floating point numbers are always represented as a round-off to their actual internal value. In Python, sometimes these can cause some unexpected results. These are not a bug in the language or your code, but simply some interesting results of the way programming languages represent floating point numbers and display them.
### 5.1. Precision
#### 5.1.1. Show me the Code!
```
x=0.1
y=0.2
z=x+y
print z # All good
# However,
z
```
#### 5.1.2. Notes
##### What is happening here ?
When you print the variable z, print takes care to represent the number rounded off to the closest
value. However when you inspect the number by not printing it, you get to see the actual number internally represented.
Technically this is called a __Representation Error__ .
### 5.2 Round-off
#### 5.2.1. Show me the Code!
```
x = 0.325
print round(x, 2) # Good
x = 0.365
print round(x, 2) # What the ...!!!
```
#### 5.2.2. Notes
##### What is happening here ?
Since the decimal fraction 0.365 is exactly half-way between 0.37 and 0.38, sometimes it could be represented by a binary
fraction which is closer to 0.36 than it is closer to 0.37. But how to find the exact precision of a float in Python ?
```
x=0.365
x # Doesn't help!
# Solution - Use decimal module
import decimal
decimal.Decimal(0.365)
decimal.Decimal(0.325) # Now you understand why 0.325 nicely rounds to 0.33
```
As you can see, 0.365 is internally represented by 0.3649999999999999911182158029987476766109466552734375 which is closer to 0.36 when rounded off to 2 decimal places. Which is why round(0.365) produces 0.36.
#### 5.2.3. Workarounds
##### 5.2.3.1 Use ceil for rounding up
```
import math
x=0.365
# Multiple and divide by power of 10 equal to precision required
math.ceil(pow(10,2)*x)/pow(10,2)
```
##### 5.2.3.2 Use decimal module
```
from decimal import *
x=Decimal(0.365).quantize(Decimal('0.01'), rounding=ROUND_UP)
y=round(x, 2)
print y
```
#### 5.3. More Reading
1. https://docs.python.org/2/tutorial/floatingpoint.html
1. https://stackoverflow.com/questions/4518641/how-to-round-off-a-floating-number-in-python
# 6. Modifying Mutables inside Immutables
#### When you have mutables (lists, dictionaries) as elements inside immutables (tuples here) you can have some unexpected results when trying to modify the former.
### 6.1. Show me the Code!
```
def make_shipment(container, items, index=0):
""" Modify objects to be shipped in 'container' by adding
objects from 'items' into it at index 'index' """
# container is a tuple containing lists
container[index] += items
# Real-life example - container of items to be exported
container = (['apples','mangoes','oranges'], ['silk','cotton','wool'])
make_shipment(container, ['papayas'])
```
#### However,
```
print container # But container is modified as well!
```
## Gotcha !!!
#### This is because,
1. For mutable types in Python,
>>> x += [y]
is not exactly the same as,
>>> x = x + [y]
2. In the first one, __x__ remains the same, but in second case, a new object is created and assigned to __x__ .
3. Hence when,
container[index] += items
is performed, the referenced list changes in-place. The item assignment doesn't work, but when the exception
occurs, the item has already been changed in place.
### 6.2. Show me the Fix!
```
def make_shipment(container, items, index=0):
""" Modify objects to be shipped in 'container' by adding
objects from 'items' into it at index 'index' """
# container is a tuple containing lists
# Use .extend(...)
container[index].extend(items)
# Real-life example - container of items to be exported
container = (['apples','mangoes','oranges'], ['silk','cotton','wool'])
make_shipment(container, ['papayas'])
print container
def make_shipment(container, items, index=0):
""" Modify objects to be shipped in 'container' by adding
objects from 'items' into it at index 'index' """
# container is a tuple containing lists
# Or retrieve the item at index to a variable
item = container[index]
# Then add to it.
item += items
# Real-life example - container of items to be exported
container = (['apples','mangoes','oranges'], ['silk','cotton','wool'])
make_shipment(container, ['papayas'])
print container
```
#### 6.3. More Reading
1. http://web.archive.org/web/20031203024741/http://zephyrfalcon.org/labs/python_pitfalls.html
# 7. Boolean Type Fallacy
#### Python doesn't respect its own boolean types. In fact, the two boolean types __True__ and __False__ can be quite flexible if you chose them to be. A developer can (often accidentally) overwrite Python's boolean types causing all kinds of problems and in this case, a bit of fun :)
### 7.1. Show me the Fun!
#### This show is named __"The Blind Truthness of Falsehood"__
```
print True
print False
x='blind'
True=x
## Fun
print 'Love is',True
print 'Hate is',not x
# Now watch the fun!
# Python allows you to overwrite its default boolean types.
False=True # Yes you can do this in #Python.
print 'Love is',x # What do you expect to get printed ?
print 'Love is',True,'as well as',False
print 'Hate is',False # What do you expect to get printed ?
print 'Hate is',False,'as well as',False
print
# REAL-LIFE, NEAR-DEATH EXAMPLE
# Point-blank situation
no_bullet_in_gun = False
if no_bullet_in_gun:
print "GO AHEAD, SHOOT ME IN THE HEAD !" # Goes ahead... your life ends here.
True='dead'
else:
print "NO PLEASE... I BEG YOU TO SPARE ME...!"
True='alive'
print 'I am',True
# Reset our world to sanity
True, False=bool(1), bool(0)
no_bullet_in_gun=False
if no_bullet_in_gun:
print "GO AHEAD, SHOOT ME IN THE HEAD !"
x='dead'
else:
print "NO PLEASE... I BEG YOU TO SPARE ME...!" # Spares you, you live to write more code in Python, but
x='alive' # hopefully not like the one above.
print 'I am',x
```
### 7.2. Show me the Fix !
#### You are kidding right ?
Word of advice - Don't overwrite your boolean types though Python allows it. It is harmful to health.
###### Copyright © Anand B Pillai, Anvetsu Technologies Pvt. Ltd (2015)
| github_jupyter |
# User Quick Start
<sub>Note if you are viewing this on github, you may need to view it on Google Colab [](https://colab.research.google.com/github/Ahuge/sept/blob/release/docs/introduction/non-developer.ipynb)</sub>
## Introduction
This `User Quick Start` guide tries to demonstrate
- How to get started modifying your own ``SEPT`` templates
- Why you should care and flexibility you can gain from using ``SEPT``
### Installation
Users can easily install ``SEPT`` according to the following steps:
Installing ``SEPT`` from the python packaging index is as simple as executing the following command:
```bash
pip install sept
```
## Getting Started
This tutorial is aimed at someone that is not a developer and instead will be a user of some program that a developer writes that takes advantage of ``SEPT``
For this tutorial, we are going to learn how to write ``SEPT`` templates using the simple application that was created from the [Developer Sample Application](./developer.ipynb#sample-qt-application) tutorial.
The finished product from that tutorial will look similar to the following:

This application is a program that will prepare several "Version" entities from your internal `Shotgun <https://shotgunsoftware.com>`_ website. If you haven't worked with Shotgun before, don't worry. You should be able to substitute Shotgun for any production tracking software, and a "Version" entity for any rendered image sequence that can is approved to send back to the movie studio.
There are 2 main components to this application.
On the left side of the dialog you can see the editor window where we can test our ``SEPT`` template.

On the right side is the help window that shows all of the ``Token`` and ``Operator`` keywords that you have access to.

## Modifying Templates
The following are some examples of path templates that you may write.
For simplicity's sake, the examples are going to work off of a single input file, however in reality you may be working with tens or even hundreds of input files at once.
The quicktime file will have the following information about it
```yaml
project: HeroJourney
sequence: Battle
shot: 001
step: comp
version: 2
extension: mov
```
Within your studio, the filepath looks like `HeroJourney_Battle_001_comp_v1.mov`.
```
!pip install sept
from sept import PathTemplateParser
data = {
"project": "HeroJourney",
"sequence": "Battle",
"shot": "001",
"step": "comp",
"version": 2,
"extension": "mov",
}
parser = PathTemplateParser()
```
### First Example
In this example, your client expects the movie file to no longer have the project code when you deliver it to them.
For example, we want the filename to be `Battle_001_comp_v2.mov`.
This means we need to write a custom template to remove the project code.
The following code block allows you to write a `SEPT` template and execute it to see the result.
```
# Type your SEPT template here:
template_str = "{{sequence}}_{{shot}}_{{step}}_v{{version}}.{{extension}}"
result = parser.parse(template_str, data)
print(result)
```
#### Breaking It Down
The template above takes the `sequence`, `shot`, and `step` tokens and joins them with an underscore.
It then adds "_v" and the `version` token to the end before adding the file `extension` token at the end.
To put a `Token` in your template you can place the name of your `Token` between two sets of curly brackets and `SEPT` will write out whatever value is in the `data` with that name.
```
{{tokenName}}
```
You can put any characters you would like outside of the curly brackets and they will be rendered exactly as you have written them.
## Introduction To Operators
There are times when the client requires naming that cannot be created by just adding tokens together from your Shotgun data.
In these cases you may need to apply an `Operator` to a `Token` to change it in some way.
`SEPT` provides several common operations out of the box but a developer can write custom ones that may apply better to your specific use case.
If there is functionality that `SEPT` does not provide out of the box that you think it should, please reach out and let me know what you think it should provide!
### Using An Operator
To use an `Operator` with your `Token` you need to modify how you write the expression.
Instead of `{{tokenName}}`, you should instead write `{{operatorName:tokenName}}`.
The syntax extends the syntax you already have learned by adding an `operatorName` followed by a full colon `:` and then the `tokenName`.
### Lowercase Template Example
In this example, our client has requested that everything in our filename is lowercase.
Without using an `Operator`, there is no easy way to achieve this, you would need to request that a producer on the show changes the name of the sequence from "Battle" to "battle". If this is at the start of the project, it may not be a huge deal, but as soon as work has started, this becomes nearly impossible to achieve without having to redo work.
But not to worry! Operators are here!
To create a filename that looks like `battle_001_comp_v2.mov`, we just need to apply a "lower" `Operator` on the sequence `Token`.
```
# Type your SEPT template here:
template_str = "{{lower:sequence}}_{{shot}}_{{step}}_v{{version}}.{{extension}}"
result = parser.parse(template_str, data)
print(result)
```
#### Breaking It Down
Luckily for us, we know that all of the shots are numbers and lowercasing a number doesn't change it so all we have to apply `lower` to is the "sequence" `Token`.
If you compare this to the previous template that we wrote, you will see that the only change is the addition of our `Operator` right at the beginning.
### Operators With Inputs
There are some advanced `Operator` types that require additional information from you to do their work.
A good example of one of these would be the `replace` `Operator`. This allows you to find and replace characters in your Token.
It needs to know what you want to find and what you want to replace it with.
These are called "Operator Inputs" and any `Operator` that requires them should provide a description of what it expects and some examples of using it.
To set the value of an input, we need to surround it in square brackets directly after the name of our `Operator`.
```
{{operatorName[inputValue]:tokenName}}
```
Some operators may expect multiple input values, the syntax for this is very similar, you just need to separate the input values with a comma.
```
{{operatorName[inputValue1,inputValue2]:tokenName}}
```
Below is the description from `replace`:
```
replace Operator
Operator Inputs
Find String: The characters that you want to search for and replace
Required: True
Replace String: The characters that you want to replace the "Find String" with.
Required: True
The replace Operator allows you to find and replace characters in your Token.
Examples (name = "alex"):
{{replace[ex,an]:name}} -> "alan"
{{replace[kite,dog:name}} -> "alex"
```
#### Replace Sequence Example
In this example, our client has renamed the "Battle" sequence and decided that it is now called "Conflict".
This messes us up because we have been working with it as "Battle" and we don't want to redo any work.
Not to worry! Operators are here!
To create a filename that looks like `Conflict_001_comp_v2.mov`, we just need to apply a "replace" `Operator` on the sequence `Token`.
```
# Type your SEPT template here:
template_str = "{{replace[Battle,Conflict]:sequence}}_{{shot}}_{{step}}_v{{version}}.{{extension}}"
result = parser.parse(template_str, data)
print(result)
```
#### Breaking It Down
Because we know that our sequence is called "Battle", we can search for the word "Battle" and replace it with "Conflict".
This is preferable to just writing the word "Conflict" in there because now our template will work even if the input file is from a different sequence because we only want to replace the "Battle" sequence.
This (again) is an extension of our syntax, compared to our "lower" example, you can see we have added `[Battle,Conflict]` in our template.
If we refer to the "replace" documentation above we can see that "replace" takes two inputs, `Find String` and `Replace String`.
In our example we have set the `Find String` equal to "Battle" and the `Replace String` equal to "Conflict". This means that any time it finds "Battle" as the sequence, it will replace it with "Conflict".
### Nested Operators
There may be certain times when you need to apply more than one `Operator` to a `Token` in order to get exactly what you want.
`SEPT` fully supports this by nesting an `Operator` within another `Operator`.
The syntax for this should be an extension of everything you have learned already. You can take an entire Token Expression and use it as the `Token` value for a separate Token Expression. This allows you to apply more than one `Operator` to a `Token`.
```
{{operatorName2:{{operatorName1:tokenName}}}}
```
`SEPT` will apply each `Operator` one at a time inside out.
#### Nested Operator Example
In this example we need to return only the first 4 characters from our sequence and then make sure that they are all in uppercase.
This will introduce you to two new operators that we need to use to achieve our goal.
The first is the opposite to the "lower" `Operator` that we saw earlier, "upper", and the second is "substr" which allows us to return a subset of the `Token`.
"upper" doesn't take any inputs and "substr" takes a `Start Location` and optionally a `End Location`.
"substr" is a bit special in that it will only accept certain values as inputs, it takes numbers for the location in the `Token` as well as "start" and "end".
To create a filename that looks like `BATT_001_comp_v2.mov`, we can use the following expression.
```
# Type your SEPT template here:
template_str = "{{substr[start,4]:{{upper:sequence}}}}_{{shot}}_{{step}}_v{{version}}.{{extension}}"
result = parser.parse(template_str, data)
print(result)
```
#### Breaking It Down
In the above example, we are applying the "upper" and then the "substr", in this specific case it doesn't matter which one happens first but it is important to keep in mind which one is being applied first.
We are passing the "substr" `Operator` two inputs, the first is "start" and the second is "4".
The value "start" will always equal "0" but for people that are not familiar with zero based index collections "start" is clearer.
## Conclusion
You've reached the end of our interactive tutorial, this should have taught you the basics of writing a `SEPT` template.
- You learned how to write a `Token`
- Apply an `Operator` to a `Token`
- Customize how the `Operator` works by passing it input values
- Apply more than one `Operator` to a `Token` by nesting the operators.
_____
If you are interested in a more technical understanding of how `SEPT` works and how you can customize it to work better for your company, you should check out the [Developer Introduction](./developer.ipynb)
| github_jupyter |
<header class="w3-container w3-teal">
<img src="images/utfsm.png" alt="" height="100px" align="left"/>
<img src="images/mat.png" alt="" height="100px" align="right"/>
</header>
<br/><br/><br/><br/><br/>
# MAT281
## Laboratorio Aplicaciones de la Matemática en la Ingeniería
### Regresión Lineal
## INSTRUCCIONES
* Anoten su nombre y rol en la celda siguiente.
* Desarrollen los problemas de manera secuencial.
* Guarden constantemente con ***Ctr-S*** para evitar sorpresas.
* Reemplacen en las celdas de código donde diga #FIX_ME por el código correspondiente.
* Ejecuten cada celda de código utilizando ***Ctr-Enter***
```
# Configuracion para recargar módulos y librerías
%reload_ext autoreload
%autoreload 2
# Si quieren graficos dentro del ipython notebook, sacar comentario a linea siguiente.
#%matplotlib inline
from IPython.core.display import HTML
HTML(open("style/mat281.css", "r").read())
from mat281_code.lab import greetings
alumno_1 = ("Sebastian Flores", "2004001-7")
alumno_2 = ("Maria Jose Vargas", "2004007-8")
HTML(greetings(alumno_1, alumno_2))
```
## Observación
Este laboratorio utiliza la librería sklearn (oficialmente llamada [scikit learn](http://scikit-learn.org/stable/)), puesto que buscamos aplicar la técnica de regresión lineal a datos tal como se haría en una aplicación real.
El laboratorio consiste de 2 secciones:
1. Explicación de uso de regresión lineal en sklearn y análisis de residuos.
3. Aplicación a problema real.
## 1. Regresión Lineal con sklearn
Generemos un conjunt de datos sintéticos para aplicar regresión lineal, utilizando numpy y sklearn. El modelo será inicialmente muy simple, del tipo
$$ y = \theta_0 + \theta_1 \ x_1 + \theta_2 x_2 + \varepsilon $$
con $x_i$, $\theta_i$ e $y$ valores reales, y $\varepsilon$ el error de medición.
```
# Generando los datos
import numpy as np
# Model
m = 1000 # Number of data examples
true_coefficients = np.array([27., 5.])
true_intercept = 1982.
n = len(true_coefficients) + 1
# Data
nx1, nx2 = 21, 26
x1, x2 = np.meshgrid(np.linspace(-1.,1.,nx1), np.linspace(80.,120.,nx2))
X = np.array([x1.flatten(), x2.flatten()]).T
Y = true_intercept + np.dot(X, true_coefficients)
# Measurement contamination
measurement_error = np.random.normal(size=Y.shape)
# Mesured data
Y = Y + 200 * measurement_error/measurement_error.max()
```
Visualicemos los datos que hemos generado.
```
# Visualización de los datos
from matplotlib import pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
# Plot of data
fig = plt.figure(figsize=(16,8))
ax = fig.add_subplot(111, projection='3d')
ax.scatter(X[:,0], X[:,1], Y, 'rs')
plt.xlabel("X[:,0]")
plt.ylabel("X[:,1]")
plt.title("y")
plt.show()
```
Apliquemos un modelo de regresión lineal, obteniendo el intercepto y coeficientes. Utilizando sklearn podemos obtener además la predicción bajo el modelo lineal de manera extremadamente sencilla.
```
# Aplicando regresion lineal con sklearn a los datos
from sklearn import linear_model
regr = linear_model.LinearRegression(fit_intercept=True, normalize=False)
regr.fit(X, Y) # No es necesario agregar la columna de 1s, el algooritmo lo hace por nosotros.
print "Intercepto hallado: ", regr.intercept_
print "Coeficientes hallados: ", regr.coef_
Y_pred = regr.predict(X) # Puedo verificar que valores daría el modelo en los puntos conocidos.
```
Visualicemos simultáneamente los datos y la superficie generada con la regresión lineal.
```
from matplotlib import pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
# Plot of data
fig = plt.figure(figsize=(16,8))
ax = fig.add_subplot(111, projection='3d')
ax.scatter(X[:,0], X[:,1], Y, 'rs')
ax.plot_trisurf(X[:,0], X[:,1], Y_pred, color="r", alpha=0.5, lw=0)
plt.xlabel("X[:,0]")
plt.ylabel("X[:,1]")
plt.title("y")
plt.show()
```
Hmm, se ve bastante bien. ¿Qué tal los residuos? ¿Responden a nuestra hipótesis de distribución normal? Realicemos una exploración visual.
```
# Analisando gráfico de los residuos
res = Y_pred - Y
plt.figure(figsize=(16,8))
plt.hist(res)
plt.show()
```
El gráfico anterior parece suficientemente normal. Utilicemos un [test de normalidad](http://docs.scipy.org/doc/scipy-0.16.1/reference/generated/scipy.stats.normaltest.html) para verificar que efectivamente los residuos tengan una distribución normal.
```
# Analisis estadístico de residuos
import scipy.stats as stats
z, pval = stats.normaltest(res)
if pval < 0.05:
print "Not normal distribution"
print "p-value =", pval
```
Calculemos el error promedio en varias normas
```
print "Error promedio en norma 1:\t", np.linalg.norm(res,1)/len(res)
print "Error promedio en norma 2:\t", np.linalg.norm(res,2)/len(res)
print "Error promedio en norma inf:\t", np.linalg.norm(res,np.inf)
```
## Desafío 1
#### [10%] Desafio 1.1
Utilice el código anterior para generar los mismos datos, pero esta vez no contamine los datos con errores gausianos, sino con elija utilizar un error de tipo gamma, uniforme o poisson (o bien, alguna otra de las [posibles](http://docs.scipy.org/doc/numpy-1.10.0/reference/routines.random.html)). Utilice alguno de los siguientes códigos:
```Python
gamma_measurement_error = np.random.gamma(1, size=Y.shape)
uniform_measurement_error = np.random.uniform(size=Y.shape)-0.5
weibull_measurement_error = np.random.weibull(1, size=Y.shape)
```
```
# 1.1 Desafio
# Codigo para generar datos con otro tipo de error de medicion
# y para realizar regresión lineal y análisis de los datos.
```
## Desafío 1: Continuación
#### [10%] 1.2 Indique el tipo de error introducido, y los valores obtenidos para el intercepto y los coeficientes.
***FIX ME: COMENTARIO AQUI ***
#### [10%] 1.3 ¿Cómo se comparan los valores anteriores a los valores reales (conocidos) y los valores obtenidos para error normal?
***FIX ME: COMENTARIO AQUI ***
#### [10%] 1.4 ¿Se logra reconocer que los residuos no corresponden a una distibución normal? ¿Cómo?
***FIX ME: COMENTARIO AQUI ***
#### [10%] 1.5 Compare el error promedio (residuos) del modelo 1 (gausiano) con el modelo 2 (no gausiano), utilizando norma 1, norma 2 y norma infinito. ¿Que observa?
***FIX ME: COMENTARIO AQUI ***
## Aplicación a datos reales
Un conjunto de datos bastante interesante es el [Motor Dataset](https://archive.ics.uci.edu/ml/datasets/Auto+MPG), que contiene características de 392 automobiles, entre los años 1970 y 1982.
<img src="images/delorean.jpg" alt="" width="600px" align="middle"/>
Este conjunto de datos tiene los siguientes valores:
1. Rendimiento en millas por galon, mpg (real).
2. Cilindros del motor, cylinders (entero positivo).
3. Displazamiento del motor en pulgadas cúbicas, displacement (real).
4. Caballos de fueza, horsepower (real).
5. Peso en libras, weight (real).
6. Tiempo de aceleración de 0 a 100 km/h, acceleration (real).
7. Año del modelo, model year (entero positivo).
8. Pais de origen, origin (entero positivo, categórico).
9. Nombre del Auto, car name (string, unico para cada dato).
Como ya es tradición, exploraremos los datos utilizando head (o tail, o cat).
```
%%bash
head data/auto-mpg.data.txt
```
## Desafío 2
#### [10 %] Desafio 2.1
Realice una regresión lineal a los datos, buscando predecir el rendimiento (en mpg) utilizando como predictores: el desplazamiento, los caballos de fuerza, el peso y la aceleración, correspondientes a las columnas 2, 3, 4 y 5. Utilice el código provisto anteriormente para ajustar un modelo de regresión lineal, obtener los coeficientes del modelo, realizar predicciones y analice los residuos.
No es necesario incluir gráficos (pero tampoco está prohibido).
```
import numpy as np
# Lectura de datos
Y = np.loadtxt("data/auto-mpg.data.txt", delimiter=" ", usecols=(0,))
X = np.loadtxt("data/auto-mpg.data.txt", delimiter=" ", usecols=(2,3,4,5))
# Procesamiento de datos
```
## Desafío 2 : Continuación
#### 2.2 [10%] ¿Qué valores se obtienen para el intercepto y los coeficientes de los datos?
***FIX ME: COMENTARIO AQUI ***
#### 2.3 [10%] ¿Cuál es el error de entrenamiento en las distintas normas?
***FIX ME: COMENTARIO AQUI ***
#### 2.4 [10%] ¿Qué tipo de residuo se obtienen? ¿Son gaussianos?
***FIX ME: COMENTARIO AQUI ***
Calculo el rendimiento en mpg tendría el Delorean, si en base a [cierta](http://www.time-traveler.org/delorean/technical.htm) [información](https://en.wikipedia.org/wiki/DeLorean_DMC-12) sabemos que posee:
1. 6 cilindros
2. 174 pulgadas cúbicas desplazamiento.
3. 200 Caballos de fuerza.
4. 2712 libras de peso.
5. Una aceleración de 10.5.
6. Se construyó el año 1981.
**Consejo**: Utilice el metodo .predict() apropiadamente.
```
# Calcular el mpg aqui
mpg = 0
print mpg
```
## Desafío 3
#### [10%] 3.1 Indique el rendimiento del delorean. ¿Se corresponde con el [valor medido](https://upload.wikimedia.org/wikipedia/commons/7/7b/DMC_Sticker.jpg) de 21 mpg?
***FIX ME: COMENTARIO AQUI ***
| github_jupyter |
# DeepPurpose Deep Dive
## Tutorial 1: Training a Drug-Target Interaction Model from Scratch
#### [@KexinHuang5](https://twitter.com/KexinHuang5)
In this tutorial, we take a deep dive into DeepPurpose and show how it builds a drug-target interaction model from scratch.
Agenda:
- Part I: Overview of DeepPurpose and Data
- Part II: Drug Target Interaction Prediction
- DeepPurpose Framework
- Applications to Drug Repurposing and Virtual Screening
- Pretrained Models
- Hyperparameter Tuning
- Model Robustness Evaluation
Let's start!
```
from DeepPurpose import utils, dataset
from DeepPurpose import DTI as models
import warnings
warnings.filterwarnings("ignore")
```
## Part I: Overview of DeepPurpose and Data
Drug-target interaction measures the binding of drug molecules to the protein targets. Accurate identification of DTI is fundamental for drug discovery and supports many downstream tasks. Among others, drug screening and repurposing are two main applications based on DTI. Drug screening helps identify ligand candidates that can bind to the protein of interest, whereas drug repurposing finds new therapeutic purposes for existing drugs. Both tasks could alleviate the costly, time-consuming, and labor-intensive process of synthesis and analysis, which is extremely important, especially in the cases of hunting effective and safe treatments for COVID-19.
DeepPurpose is a pytorch-based deep learning framework that is initiated to provide a simple but powerful toolkit for drug-target interaction prediction and its related applications. We see many exciting recent works in this direction, but to leverage these models, it takes lots of efforts due to the esoteric instructions and interface. DeepPurpose is designed to make things as simple as possible using a unified framework.
DeepPurpose uses an encoder-decoder framework. Drug repurposing and screening are two applications after we obtain DTI models. The input to the model is a drug target pair, where drug uses the simplified molecular-input line-entry system (SMILES) string and target uses the amino acid sequence. The output is a score indicating the binding activity of the drug target pair. Now, we begin talking about the data format expected.
(**Data**) DeepPurpose takes into an array of drug's SMILES strings (**d**), an array of target protein's amino acid sequence (**t**), and an array of label (**y**), which can either be binary 0/1 indicating interaction outcome or a real number indicating affinity value. The input drug and target arrays should be paired, i.e. **y**\[0\] is the score for **d**\[0\] and **t**\[0\].
Besides transforming into numpy arrays through some data wrangling on your own, DeepPurpose also provides two ways to help data preparation.
The first way is to read from local files. For example, to load drug target pairs, we expect a file.txt where each line is a drug SMILES string, followed by a protein sequence, and an affinity score or 0/1 label:
```CC1=C...C4)N MKK...LIDL 7.365``` \
```CC1=C...C4)N QQP...EGKH 4.999```
Then, we use ```dataset.read_file_training_dataset_drug_target_pairs``` to load it.
```
X_drugs, X_targets, y = dataset.read_file_training_dataset_drug_target_pairs('./toy_data/dti.txt')
print('Drug 1: ' + X_drugs[0])
print('Target 1: ' + X_targets[0])
print('Score 1: ' + str(y[0]))
```
Many method researchers want to test on benchmark datasets such as KIBA/DAVIS/BindingDB, DeepPurpose also provides data loaders to ease preprocessing. For example, we want to load the DAVIS dataset, we can use ```dataset.load_process_DAVIS```. It will download, preprocess to the designated data format. It supports label log-scale transformation for easier regression and also allows label binarization given a customized threshold.
```
X_drugs, X_targets, y = dataset.load_process_DAVIS(path = './data', binary = False, convert_to_log = True, threshold = 30)
print('Drug 1: ' + X_drugs[0])
print('Target 1: ' + X_targets[0])
print('Score 1: ' + str(y[0]))
```
For more detailed examples and tutorials of data loading, checkout this [tutorial](./DEMO/load_data_tutorial.ipynb).
## Part II: Drug Target Interaction Prediction Framework
DeepPurpose provides a simple framework to conduct DTI research using 8 encoders for drugs and 7 for proteins. It basically consists of the following steps, where each step corresponds to one line of code:
- Encoder specification
- Data encoding and split
- Model configuration generation
- Model initialization
- Model Training
- Model Prediction and Repuposing/Screening
- Model Saving and Loading
Let's start with data encoding!
(**Encoder specification**) After we obtain the required data format from Part I, we need to prepare them for the encoders. Hence, we first specify the encoder to use for drug and protein. Here we try MPNN for drug and CNN for target.
If you find MPNN and CNN are too large for the CPUs, you can try smaller encoders by uncommenting the last line:
```
drug_encoding, target_encoding = 'MPNN', 'CNN'
#drug_encoding, target_encoding = 'Morgan', 'Conjoint_triad'
```
Note that you can switch encoder just by changing the encoding name above. The full list of encoders are listed [here](https://github.com/kexinhuang12345/DeepPurpose#encodings). Here, we are using the message passing neural network encoder for drug and convolutional neural network encoder for protein.
(**Data encoding and split**) Now, we encode the data into the specified format, using ```utils.data_process``` function. It specifies train/validation/test split fractions, and random seed to ensure same data splits for reproducibility. This function also support data splitting methods such as ```cold_drug``` and ```cold_protein```, which splits on drug/proteins for model robustness evaluation to test on unseen drug/proteins.
The function outputs train, val, test pandas dataframes.
```
train, val, test = utils.data_process(X_drugs, X_targets, y,
drug_encoding, target_encoding,
split_method='random',frac=[0.7,0.1,0.2],
random_seed = 1)
train.head(1)
```
(**Model configuration generation**) Now, we initialize a model with its configuration. You can modify almost any hyper-parameters (e.g., learning rate, epoch, batch size), model parameters (e.g. hidden dimensions, filter size) and etc in this function. The supported configurations are listed here in this [link](https://github.com/kexinhuang12345/DeepPurpose/blob/e169e2f550694145077bb2af95a4031abe400a77/DeepPurpose/utils.py#L486).
For the sake of example, we specify the epoch size to be 3, and set the model parameters to be small so that you can run on both CPUs & GPUs quickly and can proceed to the next steps. For a reference parameters, checkout the notebooks in the DEMO folder.
```
config = utils.generate_config(drug_encoding = drug_encoding,
target_encoding = target_encoding,
cls_hidden_dims = [1024,1024,512],
train_epoch = 5,
LR = 0.001,
batch_size = 128,
hidden_dim_drug = 128,
mpnn_hidden_size = 128,
mpnn_depth = 3,
cnn_target_filters = [32,64,96],
cnn_target_kernels = [4,8,12]
)
```
(**Model initialization**) Next, we initialize a model using the above configuration.
```
model = models.model_initialize(**config)
model
```
(**Model Training**) Next, it is ready to train, using the ```model.train``` function! If you do not have test set, you can just use ```model.train(train, val)```.
```
model.train(train, val, test)
```
We see that the model will automatically generate and plot the training process, along with the validation result and test result.
(**Model Prediction and Repuposing/Screening**) Next, we see how we can predict affinity scores on new data. Suppose the new data is a drug-target pair below.
```
X_drug = ['CC1=C2C=C(C=CC2=NN1)C3=CC(=CN=C3)OCC(CC4=CC=CC=C4)N']
X_target = ['MKKFFDSRREQGGSGLGSGSSGGGGSTSGLGSGYIGRVFGIGRQQVTVDEVLAEGGFAIVFLVRTSNGMKCALKRMFVNNEHDLQVCKREIQIMRDLSGHKNIVGYIDSSINNVSSGDVWEVLILMDFCRGGQVVNLMNQRLQTGFTENEVLQIFCDTCEAVARLHQCKTPIIHRDLKVENILLHDRGHYVLCDFGSATNKFQNPQTEGVNAVEDEIKKYTTLSYRAPEMVNLYSGKIITTKADIWALGCLLYKLCYFTLPFGESQVAICDGNFTIPDNSRYSQDMHCLIRYMLEPDPDKRPDIYQVSYFSFKLLKKECPIPNVQNSPIPAKLPEPVKASEAAAKKTQPKARLTDPIPTTETSIAPRQRPKAGQTQPNPGILPIQPALTPRKRATVQPPPQAAGSSNQPGLLASVPQPKPQAPPSQPLPQTQAKQPQAPPTPQQTPSTQAQGLPAQAQATPQHQQQLFLKQQQQQQQPPPAQQQPAGTFYQQQQAQTQQFQAVHPATQKPAIAQFPVVSQGGSQQQLMQNFYQQQQQQQQQQQQQQLATALHQQQLMTQQAALQQKPTMAAGQQPQPQPAAAPQPAPAQEPAIQAPVRQQPKVQTTPPPAVQGQKVGSLTPPSSPKTQRAGHRRILSDVTHSAVFGVPASKSTQLLQAAAAEASLNKSKSATTTPSGSPRTSQQNVYNPSEGSTWNPFDDDNFSKLTAEELLNKDFAKLGEGKHPEKLGGSAESLIPGFQSTQGDAFATTSFSAGTAEKRKGGQTVDSGLPLLSVSDPFIPLQVPDAPEKLIEGLKSPDTSLLLPDLLPMTDPFGSTSDAVIEKADVAVESLIPGLEPPVPQRLPSQTESVTSNRTDSLTGEDSLLDCSLLSNPTTDLLEEFAPTAISAPVHKAAEDSNLISGFDVPEGSDKVAEDEFDPIPVLITKNPQGGHSRNSSGSSESSLPNLARSLLLVDQLIDL']
y = [7.365]
X_pred = utils.data_process(X_drug, X_target, y,
drug_encoding, target_encoding,
split_method='no_split')
y_pred = model.predict(X_pred)
print('The predicted score is ' + str(y_pred))
```
We can also do repurposing and screening using the trained model. Basically, for repurposing a set of existing drugs (**r**) for a single new target (*t*), we run the above prediction function after pairing each repurposing drug with the target. Similarly, for screening, we instead have a set of drug-target pairs (**d**, **t**). We wrap the operation into a ```models.repurpose``` and ```models.virtual_screening``` methods.
For example, suppose we want to do repurposing from a set of antiviral drugs for a COVID-19 target 3CL protease. The corresponding data can be retrieved using ```dataset``` functions.
```
t, t_name = dataset.load_SARS_CoV2_Protease_3CL()
print('Target Name: ' + t_name)
print('Amino Acid Sequence: '+ t)
r, r_name, r_pubchem_cid = dataset.load_antiviral_drugs()
print('Repurposing Drug 1 Name: ' + r_name[0])
print('Repurposing Drug 1 SMILES: ' + r[0])
print('Repurposing Drug 1 Pubchem CID: ' + str(r_pubchem_cid[0]))
```
Now, we can call the ```repurpose``` function. After feeding the necessary inputs, it will print a list of repurposed drugs ranked on its affinity to the target protein. The ```convert_y``` parameter should be set to be ```False``` when the ranking is ascending (i.e. lower value -> higher affinity) due to the log transformation, vice versus.
```
y_pred = models.repurpose(X_repurpose = r, target = t, model = model, drug_names = r_name, target_name = t_name,
result_folder = "./result/", convert_y = True)
```
Now, let's move on to showcase how to do virtual screening. We first load a sample of data from BindingDB dataset.
```
t, d = dataset.load_IC50_1000_Samples()
```
We can then use the ```virtual_screening``` function to generate a list of drug-target pairs that have high binding affinities. If no drug/target names are provided, the index of the drug/target list is used instead.
```
y_pred = models.virtual_screening(d, t, model)
```
Saving and loading models are also really easy. The loading function also automatically detects if the model is trained on multiple GPUs. To save a model:
```
model.save_model('./tutorial_model')
```
To load a saved/pretrained model:
```
model = models.model_pretrained(path_dir = './tutorial_model')
model
```
We have also provided a list of pretrained model, you can find all available ones under the [list](https://github.com/kexinhuang12345/DeepPurpose#pretrained-models). For example, to load a MPNN+CNN model pretrained on BindingDB Kd dataset:
```
model = models.model_pretrained(model = 'MPNN_CNN_BindingDB')
model
```
We also provided many more functionalities for DTI research purposes.
For example, this [demo](https://github.com/kexinhuang12345/DeepPurpose/blob/master/DEMO/Drug_Property_Pred-Ax-Hyperparam-Tune.ipynb) shows how to use Ax platform to do some latest hyperparameter tuning methods such as Bayesian Optimization on DeepPurpose.
Model robustness is very important for DTI task. One way to measure is to see how the model can predict drug or protein that do not exist in the training set, i.e., cold drug/target setting. You can achieve this by modifying the ```split_method``` parameter in the ```data_process``` function:
```
X_drugs, X_targets, y = dataset.load_process_DAVIS(path = './data', binary = False, convert_to_log = True, threshold = 30)
train, val, test = utils.data_process(X_drugs, X_targets, y,
drug_encoding, target_encoding,
split_method='cold_drug',frac=[0.7,0.1,0.2],
random_seed = 1)
```
That wraps up our tutorials on the main functionalities of DeepPurpose's Drug Target Interaction Prediction framework!
Do checkout the upcoming tutorials:
Tutorial 2: Drug Property Prediction using DeepPurpose
Tutorial 3: Repurposing and Virtual Screening Using One Line of Code
**Star & watch & contribute to DeepPurpose's [github repository](https://github.com/kexinhuang12345/DeepPurpose)!**
Feedbacks would also be appreciated and you can send me an email (kexinhuang@hsph.harvard.edu)!
| github_jupyter |
```
import numpy as np
import pandas as pd
import torch
import torchvision
from torch.utils.data import Dataset, DataLoader
from torchvision import transforms, utils
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from matplotlib import pyplot as plt
%matplotlib inline
torch.backends.cudnn.deterministic = True
torch.backends.cudnn.benchmark = False
transform = transforms.Compose(
[transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
trainset = torchvision.datasets.CIFAR10(root='./data', train=True, download=True, transform=transform)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=10, shuffle=False)
classes = ('plane', 'car', 'bird', 'cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck')
foreground_classes = {'plane', 'car', 'bird'}
#foreground_classes = {'bird', 'cat', 'deer'}
background_classes = {'cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck'}
#background_classes = {'plane', 'car', 'dog', 'frog', 'horse','ship', 'truck'}
fg1,fg2,fg3 = 0,1,2
dataiter = iter(trainloader)
background_data=[]
background_label=[]
foreground_data=[]
foreground_label=[]
batch_size=10
for i in range(5000):
images, labels = dataiter.next()
for j in range(batch_size):
if(classes[labels[j]] in background_classes):
img = images[j].tolist()
background_data.append(img)
background_label.append(labels[j])
else:
img = images[j].tolist()
foreground_data.append(img)
foreground_label.append(labels[j])
foreground_data = torch.tensor(foreground_data)
foreground_label = torch.tensor(foreground_label)
background_data = torch.tensor(background_data)
background_label = torch.tensor(background_label)
def create_mosaic_img(bg_idx,fg_idx,fg):
"""
bg_idx : list of indexes of background_data[] to be used as background images in mosaic
fg_idx : index of image to be used as foreground image from foreground data
fg : at what position/index foreground image has to be stored out of 0-8
"""
image_list=[]
j=0
for i in range(9):
if i != fg:
image_list.append(background_data[bg_idx[j]])#.type("torch.DoubleTensor"))
j+=1
else:
image_list.append(foreground_data[fg_idx])#.type("torch.DoubleTensor"))
label = foreground_label[fg_idx]-fg1 # minus 7 because our fore ground classes are 7,8,9 but we have to store it as 0,1,2
#image_list = np.concatenate(image_list ,axis=0)
image_list = torch.stack(image_list)
return image_list,label
desired_num = 30000
mosaic_list_of_images =[] # list of mosaic images, each mosaic image is saved as list of 9 images
fore_idx =[] # list of indexes at which foreground image is present in a mosaic image i.e from 0 to 9
mosaic_label=[] # label of mosaic image = foreground class present in that mosaic
for i in range(desired_num):
np.random.seed(i)
bg_idx = np.random.randint(0,35000,8)
fg_idx = np.random.randint(0,15000)
fg = np.random.randint(0,9)
fore_idx.append(fg)
image_list,label = create_mosaic_img(bg_idx,fg_idx,fg)
mosaic_list_of_images.append(image_list)
mosaic_label.append(label)
class MosaicDataset(Dataset):
"""MosaicDataset dataset."""
def __init__(self, mosaic_list_of_images, mosaic_label, fore_idx):
"""
Args:
csv_file (string): Path to the csv file with annotations.
root_dir (string): Directory with all the images.
transform (callable, optional): Optional transform to be applied
on a sample.
"""
self.mosaic = mosaic_list_of_images
self.label = mosaic_label
self.fore_idx = fore_idx
def __len__(self):
return len(self.label)
def __getitem__(self, idx):
return self.mosaic[idx] , self.label[idx], self.fore_idx[idx]
batch = 250
msd = MosaicDataset(mosaic_list_of_images, mosaic_label , fore_idx)
train_loader = DataLoader( msd,batch_size= batch ,shuffle=False,num_workers=0,)
data,labels,fg_index = iter(train_loader).next()
bg = []
for i in range(120):
torch.manual_seed(i)
betag = torch.ones((250,9))/9 #torch.randn(250,9)
a=bg.append( betag.requires_grad_() )
class Module2(nn.Module):
def __init__(self):
super(Module2, self).__init__()
self.conv1 = nn.Conv2d(3, 6, 5)
self.pool = nn.MaxPool2d(2, 2)
self.conv2 = nn.Conv2d(6, 16, 5)
self.fc1 = nn.Linear(16 * 5 * 5, 120)
self.fc2 = nn.Linear(120, 84)
self.fc3 = nn.Linear(84, 10)
self.fc4 = nn.Linear(10,3)
def forward(self,y): #z batch of list of 9 images
y1 = self.pool(F.relu(self.conv1(y)))
y1 = self.pool(F.relu(self.conv2(y1)))
y1 = y1.view(-1, 16 * 5 * 5)
y1 = F.relu(self.fc1(y1))
y1 = F.relu(self.fc2(y1))
y1 = F.relu(self.fc3(y1))
y1 = self.fc4(y1)
return y1
torch.manual_seed(1234)
what_net = Module2().double()
#what_net.load_state_dict(torch.load("simultaneous_what.pt"))
what_net = what_net.to("cuda")
def attn_avg(x,beta):
y = torch.zeros([batch,3, 32,32], dtype=torch.float64)
y = y.to("cuda")
alpha = F.softmax(beta,dim=1) # alphas
for i in range(9):
alpha1 = alpha[:,i]
y = y + torch.mul(alpha1[:,None,None,None],x[:,i])
return y,alpha
def calculate_attn_loss(dataloader,what,criter):
what.eval()
r_loss = 0
alphas = []
lbls = []
pred = []
fidices = []
correct = 0
tot = 0
with torch.no_grad():
for i, data in enumerate(dataloader, 0):
inputs, labels,fidx= data
lbls.append(labels)
fidices.append(fidx)
inputs = inputs.double()
beta = bg[i] # alpha for ith batch
inputs, labels,beta = inputs.to("cuda"),labels.to("cuda"),beta.to("cuda")
avg,alpha = attn_avg(inputs,beta)
alpha = alpha.to("cuda")
outputs = what(avg)
_, predicted = torch.max(outputs.data, 1)
correct += sum(predicted == labels)
tot += len(predicted)
pred.append(predicted.cpu().numpy())
alphas.append(alpha.cpu().numpy())
loss = criter(outputs, labels)
r_loss += loss.item()
alphas = np.concatenate(alphas,axis=0)
pred = np.concatenate(pred,axis=0)
lbls = np.concatenate(lbls,axis=0)
fidices = np.concatenate(fidices,axis=0)
#print(alphas.shape,pred.shape,lbls.shape,fidices.shape)
analysis = analyse_data(alphas,lbls,pred,fidices)
return r_loss/(i+1),analysis,correct.item(),tot,correct.item()/tot
# for param in what_net.parameters():
# param.requires_grad = False
def analyse_data(alphas,lbls,predicted,f_idx):
'''
analysis data is created here
'''
batch = len(predicted)
amth,alth,ftpt,ffpt,ftpf,ffpf = 0,0,0,0,0,0
for j in range (batch):
focus = np.argmax(alphas[j])
if(alphas[j][focus] >= 0.5):
amth +=1
else:
alth +=1
if(focus == f_idx[j] and predicted[j] == lbls[j]):
ftpt += 1
elif(focus != f_idx[j] and predicted[j] == lbls[j]):
ffpt +=1
elif(focus == f_idx[j] and predicted[j] != lbls[j]):
ftpf +=1
elif(focus != f_idx[j] and predicted[j] != lbls[j]):
ffpf +=1
#print(sum(predicted==lbls),ftpt+ffpt
return [ftpt,ffpt,ftpf,ffpf,amth,alth]
optim1 = []
for i in range(120):
optim1.append(optim.RMSprop([bg[i]], lr=0.01))
# instantiate optimizer
optimizer_what = optim.RMSprop(what_net.parameters(), lr=0.001)#, momentum=0.9)#,nesterov=True)
criterion = nn.CrossEntropyLoss()
acti = []
analysis_data_tr = []
analysis_data_tst = []
loss_curi_tr = []
loss_curi_tst = []
epochs = 100
# calculate zeroth epoch loss and FTPT values
running_loss,anlys_data,correct,total,accuracy = calculate_attn_loss(train_loader,what_net,criterion)
print('training epoch: [%d ] loss: %.3f correct: %.3f, total: %.3f, accuracy: %.3f' %(0,running_loss,correct,total,accuracy))
loss_curi_tr.append(running_loss)
analysis_data_tr.append(anlys_data)
# training starts
for epoch in range(epochs): # loop over the dataset multiple times
ep_lossi = []
running_loss = 0.0
what_net.train()
for i, data in enumerate(train_loader, 0):
# get the inputs
inputs, labels,_ = data
inputs = inputs.double()
beta = bg[i] # alpha for ith batch
inputs, labels,beta = inputs.to("cuda"),labels.to("cuda"),beta.to("cuda")
# zero the parameter gradients
optimizer_what.zero_grad()
optim1[i].zero_grad()
# forward + backward + optimize
avg,alpha = attn_avg(inputs,beta)
outputs = what_net(avg)
loss = criterion(outputs, labels)
# print statistics
running_loss += loss.item()
#alpha.retain_grad()
loss.backward(retain_graph=False)
optimizer_what.step()
optim1[i].step()
running_loss_tr,anls_data,correct,total,accuracy = calculate_attn_loss(train_loader,what_net,criterion)
analysis_data_tr.append(anls_data)
loss_curi_tr.append(running_loss_tr) #loss per epoch
print('training epoch: [%d ] loss: %.3f correct: %.3f, total: %.3f, accuracy: %.3f' %(epoch+1,running_loss_tr,correct,total,accuracy))
if running_loss_tr<=0.08:
break
print('Finished Training run ')
analysis_data_tr = np.array(analysis_data_tr)
columns = ["epochs", "argmax > 0.5" ,"argmax < 0.5", "focus_true_pred_true", "focus_false_pred_true", "focus_true_pred_false", "focus_false_pred_false" ]
df_train = pd.DataFrame()
df_test = pd.DataFrame()
df_train[columns[0]] = np.arange(0,epoch+2)
df_train[columns[1]] = analysis_data_tr[:,-2]/300
df_train[columns[2]] = analysis_data_tr[:,-1]/300
df_train[columns[3]] = analysis_data_tr[:,0]/300
df_train[columns[4]] = analysis_data_tr[:,1]/300
df_train[columns[5]] = analysis_data_tr[:,2]/300
df_train[columns[6]] = analysis_data_tr[:,3]/300
df_train
fig= plt.figure(figsize=(6,6))
plt.plot(df_train[columns[0]],df_train[columns[3]], label ="focus_true_pred_true ")
plt.plot(df_train[columns[0]],df_train[columns[4]], label ="focus_false_pred_true ")
plt.plot(df_train[columns[0]],df_train[columns[5]], label ="focus_true_pred_false ")
plt.plot(df_train[columns[0]],df_train[columns[6]], label ="focus_false_pred_false ")
plt.title("On Train set")
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.xticks([0,5,10,15,20])
plt.xlabel("epochs")
plt.ylabel("percentage of data")
#plt.vlines(vline_list,min(min(df_train[columns[3]]/300),min(df_train[columns[4]]/300),min(df_train[columns[5]]/300),min(df_train[columns[6]]/300)), max(max(df_train[columns[3]]/300),max(df_train[columns[4]]/300),max(df_train[columns[5]]/300),max(df_train[columns[6]]/300)),linestyles='dotted')
plt.show()
fig.savefig("train_analysis.pdf")
fig.savefig("train_analysis.png")
aph = []
for i in bg:
aph.append(F.softmax(i,dim=1).detach().numpy())
aph = np.concatenate(aph,axis=0)
torch.save({
'epoch': 500,
'model_state_dict': what_net.state_dict(),
#'optimizer_state_dict': optimizer_what.state_dict(),
"optimizer_alpha":optim1,
"FTPT_analysis":analysis_data_tr,
"alpha":aph
}, "cifar_what_net_500.pt")
aph
running_loss_tr,anls_data,correct,total,accuracy = calculate_attn_loss(train_loader,what_net,criterion)
print("argmax>0.5",anls_data[-2])
```
| github_jupyter |
```
import pandas as pd
import numpy as np
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.feature_extraction.text import TfidfTransformer
from sklearn.naive_bayes import MultinomialNB
from sklearn.model_selection import train_test_split
from sklearn.metrics import classification_report
import seaborn as sns
import re
import random as rd
import tensorflow as tf
from tensorflow.keras.layers.experimental.preprocessing import TextVectorization
from nltk import word_tokenize, RegexpTokenizer
from nltk.corpus import stopwords, wordnet
from sklearn.preprocessing import OneHotEncoder
import matplotlib.pyplot as plt
import autokeras as ak
from sklearn.model_selection import KFold
import nltk
from sklearn.svm import SVC
from sklearn.linear_model import LogisticRegression
nltk.download('stopwords')
nltk.download('wordnet')
def Tokenize(f): ## Pre-processando a frase
## Colocando em minusculo
## Retirando a pontuaçao
## Retirando as StopWords
f = f.lower().replace('\n', '').replace('-','').replace('#','').replace('.','').replace(',','').replace('!','').replace('r\n','').replace(' ','').replace('https','').replace('rt','').replace('rn','')
token = RegexpTokenizer(r"\w+")
f = token.tokenize(f)
stop_words = set(stopwords.words('portuguese'))
new_word = [word for word in f if not word in stop_words]
return ' '.join(new_word)
def remove_user(frase):
frase = re.sub('@\w+','',frase)
frase = re.sub('{https}[^ ]+','',frase)
frase = re.sub('https\w+','',frase)
# re.sub('#\w+','',frase)
return frase
def pre_X(frases):
lista = []
for frase in frases:
lista.append(frase)
return lista
def pre_Y(number):
lista = []
for numb in number:
lista.append(numb)
return lista
def set_array(frases):
vocab = []
palavras = []
for frase in frases:
text_array = remove_user(frase)
text_array = Tokenize(text_array)
text_array = text_array.split(' ')
for i in range(len(text_array)):
vocab.append(text_array[i])
return vocab
```
## KANSOAN
```
df = pd.read_csv("data.txt", sep=',', header=None, names=['text','sentiment'])
df_remove = df[df['sentiment'] == '#Inveja'].index
df = df.drop(df_remove)
df_remove = df[df['sentiment'] == '#Raiva'].index
df = df.drop(df_remove)
df_remove = df[df['sentiment'] == '#Ironia'].index
df = df.drop(df_remove)
def binario(termo):
if termo == '#Feliz' or termo == '#Amor':
return 1
elif termo == '#Triste' or termo == '#Chateado':
return 0
df['sentiment'] = df['sentiment'].apply(binario)
df['text'] = df['text'].apply(remove_user)
df['text'] = df['text'].apply(Tokenize)
text_KANSOAN = df['text']
sentiment = np.asarray(df['sentiment'])
count_vect = CountVectorizer()
tfidf_transformer = TfidfTransformer()
X_train_KANSOAN = count_vect.fit_transform(text_KANSOAN)
X_train_KANSOAN = tfidf_transformer.fit_transform(X_train_KANSOAN) # Aplicando o TF-IDF
X_train_KANSOAN.shape
```
## TWEETSENT
```
df_tweet= pd.read_csv('export_TweetSentBR.csv')
df_tweet
df_remove = df_tweet[df_tweet['sentiment'] == '-']
df_tweet = df_tweet.drop(df_remove.index)
df_remove = df_tweet[df_tweet['sentiment'] == '0']
df_tweet = df_tweet.drop(df_remove.index)
df_tweet['text'] = df_tweet['text'].apply(remove_user)
df_tweet['text'] = df_tweet['text'].apply(Tokenize)
df_tweet['sentiment'] = df_tweet['sentiment'].apply(lambda x: int(x))
text_Tweetsent = df_tweet['text']
polarity_ = np.asarray(df_tweet['sentiment'])
X_text = count_vect.transform(text_Tweetsent)
X_test_TWEET = tfidf_transformer.transform(X_text) # Aplicando o TF-IDF
X_test_TWEET.shape
```
## TASH
```
df1 = pd.read_csv("tash-pt.csv")
df1 = df1.dropna()
df1 = df1.drop(columns=['id_twitter'])
df1['text'] = df1['text'].apply(remove_user)
df1['text'] = df1['text'].apply(Tokenize)
df1_remove = df1.drop(df1[df1['sentiment'] == 0].index)
sentiment_ = np.asarray(df1_remove['sentiment'])
text_TASH = df1_remove['text']
X_text_TASH = count_vect.transform(text_TASH)
X_test_TASH_ = tfidf_transformer.transform(X_text_TASH) # Aplicando o TF-IDF
X_test_TASH_.shape
```
## UNILEX
```
df1_ = pd.read_excel('./rotulação/TweetsPolitical01 OK OK.xlsx')
df2 = pd.read_excel('./rotulação/TweetsPolitical02 OK OK.xlsx')
df3 = pd.read_excel('./rotulação/TweetsPolitical03 OK OK.xlsx')
df4 = pd.read_excel('./rotulação/TweetsPolitical04 OK OK.xlsx')
df5 = pd.read_excel('./rotulação/TweetsPolitical05 OK OK.xlsx')
df6 = pd.read_excel('./rotulação/TweetsPolitical06 OK OK.xlsx')
df7 = pd.read_excel('./rotulação/TweetsPolitical07 OK OK.xlsx')
df8 = pd.read_excel('./rotulação/TweetsPolitical08 OK OK.xlsx')
df9 = pd.read_excel('./rotulação/TweetsPolitical09 OK OK.xlsx')
df10 = pd.read_excel('./rotulação/TweetsPolitical10 OK OK.xlsx')
df11 = pd.read_excel('./rotulação/TweetsPolitical11 OK OK.xlsx')
df12 = pd.read_excel('./rotulação/TweetsPolitical12 OK OK.xlsx')
df13 = pd.read_excel('./rotulação/TweetsPolitical13 OK OK.xlsx')
df14 = pd.read_excel('./rotulação/TweetsPolitical14 OK OK.xlsx')
df15 = pd.read_excel('./rotulação/TweetsPolitical15 OK OK.xlsx')
lista = [df1_,df2,df3,df4,df5,df6,df7,df8,df9,df10,df11,df12,df13,df14,df15]
for df_ in lista:
df_.drop(df_[df_['Polaridade']== 0.0].index, inplace=True)
df_all = pd.concat(lista)
df_all = df_all.reset_index()
df_all
df_all = df_all.dropna()
df_all
df_all['Polaridade'].value_counts()
remove1 = df_all[df_all['Polaridade'] == 11].index
df_all = df_all.drop(remove1).reset_index()
df_all = df_all.drop(columns=['index','level_0'])
remove2 = df_all[df_all['Polaridade'] == -2].index
df_all = df_all.drop(remove2).reset_index()
df_all = df_all.drop(columns=['index'])
remove3 = df_all[df_all['Polaridade'] == 10].index
df_all = df_all.drop(remove3).reset_index()
df_all = df_all.drop(columns=['index'])
df_all
df_all['Polaridade'].value_counts()
df_all
df_all['Tweet'] = df_all['Tweet'].apply(remove_user)
df_all['Tweet'] = df_all['Tweet'].apply(Tokenize)
polarity = np.asarray(df_all['Polaridade'])
Tweet = df_all['Tweet']
Tweet = count_vect.transform(Tweet)
X_test_UNILEX = tfidf_transformer.transform(Tweet) # Aplicando o TF-IDF
X_test_UNILEX.shape
def func(num):
if num == 0:
return -1
else:
return 1
sentiment = df['sentiment'].apply(func)
```
## NB - TRAIN -> KANSOAN- TEST -> TWEET
```
## TIRAR OS SPLITS , até no treino
clf = MultinomialNB()
clf.fit(X_train_KANSOAN, sentiment)
y_pred = clf.predict(X_test_TWEET)
print(classification_report(polarity_,y_pred) )
```
## NB - TRAIN -> KANSOAN - TEST -> UNILEX
```
clf = MultinomialNB()
clf.fit(X_train_KANSOAN, sentiment)
y_pred = clf.predict(X_test_UNILEX)
print(classification_report(polarity,y_pred))
```
## NB - TRAIN -> KANSOAN - TEST -> TASH
```
clf = MultinomialNB()
clf.fit(X_train_KANSOAN, sentiment)
y_pred = clf.predict(X_test_TASH_)
print(classification_report(sentiment_,y_pred))
```
## SVM - TRAIN->KANSOAN - TEST-> TWEET
```
clf = SVC().fit(X_train_KANSOAN, sentiment)
y_pred = clf.predict(X_test_TWEET)
print(classification_report(polarity_,y_pred))
```
## SVM T KANSOAN T UNILEX
```
clf = SVC().fit(X_train_KANSOAN, sentiment)
y_pred = clf.predict(X_test_UNILEX)
print(classification_report(polarity,y_pred))
```
## SVM T KANSOAN T TASH
```
clf = SVC().fit(X_train_KANSOAN, sentiment)
y_pred = clf.predict(X_test_TASH_)
print(classification_report(sentiment_,y_pred, zero_division=True))
```
## RL T KANSOAN T TWEET
```
clf = LogisticRegression(max_iter=1000).fit(X_train_KANSOAN, sentiment)
y_pred = clf.predict(X_test_TWEET)
print(classification_report(polarity_,y_pred))
```
## RL T KANSOAN T UNILEX
```
clf = LogisticRegression(max_iter=1000).fit(X_train_KANSOAN, sentiment)
y_pred = clf.predict(X_test_UNILEX)
print(classification_report(polarity,y_pred))
```
## RL T KANSOAN T TASH
```
clf = LogisticRegression(max_iter=1000).fit(X_train_KANSOAN, sentiment)
y_pred = clf.predict(X_test_TASH_)
print(classification_report(sentiment_,y_pred))
one = OneHotEncoder(sparse=False)
teste_ = one.fit_transform(np.asarray(sentiment).reshape(-1,1))
nome = pd.DataFrame(teste_)
nome['coluna'] = 0
nome.rename(columns={1: 2, 'coluna':1} , inplace=True)
Y_KANSOAN = nome[[0,1,2]].to_numpy()
def func(num):
if num == -1:
return 0
else:
return 1
sentiment = df['sentiment']
polarity_ = np.asarray(df_tweet['sentiment'].apply(func))
polarity = np.asarray(df_all['Polaridade'].apply(func))
sentiment_ = np.asarray(df1_remove['sentiment'].apply(func))
np.unique(sentiment_)
```
## TOTALMENT.C T KANSOAN T TWEET
```
model = tf.keras.Sequential([
tf.keras.layers.Dense(50, activation='relu'),
tf.keras.layers.Dropout(0.3),
tf.keras.layers.Dense(25, activation='relu'),
tf.keras.layers.Dropout(0.3),
tf.keras.layers.Dense(10, activation='tanh'),
tf.keras.layers.Dropout(0.3),
tf.keras.layers.Dense(1 , activation='sigmoid')
])
model.compile(
optimizer='adam',
loss=tf.keras.losses.binary_crossentropy,
metrics=['accuracy']
)
y_one = np.asarray(sentiment)
fit = model.fit(X_train_KANSOAN.todense(), y_one, epochs=5, validation_data=(X_test_TWEET.todense(),polarity_))
predicted = model.predict(X_test_TWEET.todense())
def ajeita(predicted):
for i in range(len(predicted)):
if predicted[i,0] >= 0.5:
predicted[i,0] = 1
else:
predicted[i,0] = 0
return predicted
predicted = ajeita(predicted)
print(classification_report(polarity_, predicted))
```
## UNILEX
```
predicted = model.predict(X_test_UNILEX.todense())
predicted = ajeita(predicted)
print(classification_report(polarity, predicted))
```
## TASH
```
predicted = model.predict(X_test_TASH_.todense())
predicted = ajeita(predicted)
print(classification_report(sentiment_, predicted))
```
## LSTM T KANSOAN T UNILEX
```
vectorize_layer = TextVectorization(
max_tokens=15000,
output_mode='int',
output_sequence_length=len(max(text_KANSOAN )))
vocab = set_array(text_KANSOAN )
vectorize_layer.adapt(np.unique(vocab))
len(vectorize_layer.get_vocabulary())
model = tf.keras.Sequential([
vectorize_layer,
tf.keras.layers.Embedding(
input_dim=len(vectorize_layer.get_vocabulary()),
output_dim=64,mask_zero=True),
tf.keras.layers.LSTM(50, activation='relu' ,return_sequences=True),
tf.keras.layers.Dropout(0.3),
tf.keras.layers.LSTM(25 , activation='tanh', return_sequences=True),
tf.keras.layers.Dropout(0.3),
tf.keras.layers.LSTM(10 , activation='tanh'),
tf.keras.layers.Dropout(0.2),
tf.keras.layers.Dense(1, activation='sigmoid')
])
model.compile(
optimizer= tf.keras.optimizers.Adam(),
loss=tf.keras.losses.binary_crossentropy,
metrics=['accuracy']
)
y_one = np.asarray(sentiment)
fit = model.fit(np.asarray(pre_X(text_KANSOAN)), y_one, epochs=5, batch_size=128,validation_data=(np.asarray(pre_X(df_all['Tweet'])),polarity))
predicted = model.predict(np.asarray(pre_X(df_all['Tweet'])))
predicted = ajeita(predicted)
print(classification_report(polarity,predicted))
```
## TWEET
```
predicted = model.predict(np.asarray(pre_X(text_Tweetsent)))
predicted = ajeita(predicted)
print(classification_report(polarity_, predicted))
```
## TASH
```
predicted = model.predict(np.asarray(pre_X(text_TASH)))
predicted = ajeita(predicted)
print(classification_report(sentiment_, predicted))
```
## CONV1D T KANSOAN T TASH
```
model = tf.keras.Sequential([
vectorize_layer,
tf.keras.layers.Embedding(
input_dim=len(vectorize_layer.get_vocabulary()),
output_dim=64,
mask_zero=True),
tf.keras.layers.Conv1D(32,6, activation='relu'),
tf.keras.layers.MaxPooling1D(2),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(16, activation='relu'),
tf.keras.layers.Dense(1, activation='sigmoid')
])
model.compile(
optimizer= tf.keras.optimizers.Adam(),
loss=tf.keras.losses.binary_crossentropy,
metrics=['accuracy']
)
y_one = np.asarray(sentiment)
fit = model.fit(np.asarray(pre_X(text_KANSOAN)), y_one, epochs=5, batch_size=128, validation_data=(np.asarray(pre_X(text_TASH)),sentiment_))
predicted = model.predict(np.asarray(pre_X(text_TASH)))
predicted = ajeita(predicted)
print(classification_report(sentiment_,predicted))
```
## TWEET
```
predicted = model.predict(np.asarray(pre_X(text_Tweetsent)))
predicted = ajeita(predicted)
print(classification_report(polarity_, predicted))
```
## UNILEX
```
predicted = model.predict(np.asarray(pre_X(df_all['Tweet'])))
predicted = ajeita(predicted)
print(classification_report(polarity, predicted))
```
## BDR T KANSOAN T TWEET
```
model = tf.keras.Sequential([
vectorize_layer,
tf.keras.layers.Embedding(
input_dim=len(vectorize_layer.get_vocabulary()),
output_dim=64,
mask_zero=True),
tf.keras.layers.Bidirectional(tf.keras.layers.LSTM(64)),
tf.keras.layers.Dense(16, activation='relu'),
tf.keras.layers.Dense(1, activation='sigmoid')
])
model.compile(
optimizer= tf.keras.optimizers.Adam(),
loss=tf.keras.losses.binary_crossentropy,
metrics=['accuracy']
)
y_one = np.asarray(sentiment)
fit = model.fit(np.asarray(pre_X(text_KANSOAN)), y_one, epochs=5, batch_size=128,validation_data=(np.asarray(pre_X(text_Tweetsent)),polarity_))
predicted = model.predict(np.asarray(pre_X(text_Tweetsent)))
predicted = ajeita(predicted)
print(classification_report(polarity_, predicted))
```
## UNILEX
```
predicted = model.predict(np.asarray(pre_X(df_all['Tweet'])))
predicted = ajeita(predicted)
print(classification_report(polarity,predicted))
```
## TASH
```
predicted = model.predict(np.asarray(pre_X(text_TASH)))
predicted = ajeita(predicted)
print(classification_report(sentiment_, predicted))
```
| github_jupyter |
# Assignment 2
Before working on this assignment please read these instructions fully. In the submission area, you will notice that you can click the link to **Preview the Grading** for each step of the assignment. This is the criteria that will be used for peer grading. Please familiarize yourself with the criteria before beginning the assignment.
An NOAA dataset has been stored in the file `data/C2A2_data/BinnedCsvs_d400/fb441e62df2d58994928907a91895ec62c2c42e6cd075c2700843b89.csv`. The data for this assignment comes from a subset of The National Centers for Environmental Information (NCEI) [Daily Global Historical Climatology Network](https://www1.ncdc.noaa.gov/pub/data/ghcn/daily/readme.txt) (GHCN-Daily). The GHCN-Daily is comprised of daily climate records from thousands of land surface stations across the globe.
Each row in the assignment datafile corresponds to a single observation.
The following variables are provided to you:
* **id** : station identification code
* **date** : date in YYYY-MM-DD format (e.g. 2012-01-24 = January 24, 2012)
* **element** : indicator of element type
* TMAX : Maximum temperature (tenths of degrees C)
* TMIN : Minimum temperature (tenths of degrees C)
* **value** : data value for element (tenths of degrees C)
For this assignment, you must:
1. Read the documentation and familiarize yourself with the dataset, then write some python code which returns a line graph of the record high and record low temperatures by day of the year over the period 2005-2014. The area between the record high and record low temperatures for each day should be shaded.
2. Overlay a scatter of the 2015 data for any points (highs and lows) for which the ten year record (2005-2014) record high or record low was broken in 2015.
3. Watch out for leap days (i.e. February 29th), it is reasonable to remove these points from the dataset for the purpose of this visualization.
4. Make the visual nice! Leverage principles from the first module in this course when developing your solution. Consider issues such as legends, labels, and chart junk.
The data you have been given is near **Ann Arbor, Michigan, United States**, and the stations the data comes from are shown on the map below.
```
import matplotlib.pyplot as plt
import mplleaflet
import pandas as pd
def leaflet_plot_stations(binsize, hashid):
df = pd.read_csv('data/C2A2_data/BinSize_d{}.csv'.format(binsize))
station_locations_by_hash = df[df['hash'] == hashid]
lons = station_locations_by_hash['LONGITUDE'].tolist()
lats = station_locations_by_hash['LATITUDE'].tolist()
plt.figure(figsize=(8,8))
plt.scatter(lons, lats, c='r', alpha=0.7, s=200)
return mplleaflet.display()
leaflet_plot_stations(400,'fb441e62df2d58994928907a91895ec62c2c42e6cd075c2700843b89')
df = pd.read_csv('data/C2A2_data/BinnedCsvs_d100/4e86d2106d0566c6ad9843d882e72791333b08be3d647dcae4f4b110.csv')
df.head()
# to show the head of the df after sorting
df.sort(['ID','Date']).head()
# to take the year and df from the tuple of the zip function
df['Year'], df['Month-Date'] = zip(*df['Date'].apply(lambda x: (x[:4], x[5:])))
# to opt out the leap year days
df = df[df['Month-Date'] != '02-29']
import numpy as np
# to get the max and min of the temperatures
t_min1 = df[(df['Element'] == 'TMIN') & (df['Year'] != '2015')].groupby('Month-Date').aggregate({'Data_Value':np.min})
t_max1 = df[(df['Element'] == 'TMAX') & (df['Year'] != '2015')].groupby('Month-Date').aggregate({'Data_Value':np.max})
t_min1.head()
# to get the minimum of temperatures
t_min_15 = df[(df['Element'] == 'TMIN') & (df['Year'] == '2015')].groupby('Month-Date').aggregate({'Data_Value':np.min})
t_max_15 = df[(df['Element'] == 'TMAX') & (df['Year'] == '2015')].groupby('Month-Date').aggregate({'Data_Value':np.max})
b_min1 = np.where(t_min_15['Data_Value'] < t_min1['Data_Value'])[0]
b_max1 = np.where(t_max_15['Data_Value'] > t_max1['Data_Value'])[0]
b_max1, b_min1
t_min_15.head()
plt.figure()
plt.plot(t_min1.values, 'b', label = 'record low')
plt.plot(t_max1.values, 'r', label = 'record high')
plt.scatter(b_min1, t_min_15.iloc[b_min1], s = 10, c = 'g', label = 'broken low')
plt.scatter(b_max1, t_max_15.iloc[b_max1], s = 10, c = 'm', label = 'broken high')
plt.gca().axis([-5, 370, -150, 650])
plt.xticks(range(0, len(t_min1), 20), t_min1.index[range(0, len(t_min1), 20)], rotation = '45')
plt.xlabel('Day of the Year')
plt.ylabel('Temperature (Tenths of Degrees C)')
plt.title('Temperature Summary Plot near Singapore')
plt.legend(loc = 4, frameon = False)
plt.gca().fill_between(range(len(t_min1)), t_min1['Data_Value'], t_max1['Data_Value'], facecolor = 'yellow', alpha = 0.5)
plt.gca().spines['top'].set_visible(False)
plt.gca().spines['right'].set_visible(False)
plt.show()
```
| github_jupyter |
##### Copyright 2020 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Better performance with tf.function
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/guide/function"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/guide/function.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/guide/function.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/guide/function.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
</td>
</table>
In TensorFlow 2, eager execution is turned on by default. The user interface is intuitive and flexible (running one-off operations is much easier
and faster), but this can come at the expense of performance and deployability.
You can use `tf.function` to make graphs out of your programs. It is a transformation tool that creates Python-independent dataflow graphs out of your Python code. This will help you create performant and portable models, and it is required to use `SavedModel`.
This guide will help you conceptualize how `tf.function` works under the hood so you can use it effectively.
The main takeaways and recommendations are:
- Debug in eager mode, then decorate with `@tf.function`.
- Don't rely on Python side effects like object mutation or list appends.
- `tf.function` works best with TensorFlow ops; NumPy and Python calls are converted to constants.
## Setup
```
import tensorflow as tf
```
Define a helper function to demonstrate the kinds of errors you might encounter:
```
import traceback
import contextlib
# Some helper code to demonstrate the kinds of errors you might encounter.
@contextlib.contextmanager
def assert_raises(error_class):
try:
yield
except error_class as e:
print('Caught expected exception \n {}:'.format(error_class))
traceback.print_exc(limit=2)
except Exception as e:
raise e
else:
raise Exception('Expected {} to be raised but no error was raised!'.format(
error_class))
```
## Basics
### Usage
A `Function` you define (for example by applying the `@tf.function` decorator) is just like a core TensorFlow operation: You can execute it eagerly; you can compute gradients; and so on.
```
@tf.function # The decorator converts `add` into a `Function`.
def add(a, b):
return a + b
add(tf.ones([2, 2]), tf.ones([2, 2])) # [[2., 2.], [2., 2.]]
v = tf.Variable(1.0)
with tf.GradientTape() as tape:
result = add(v, 1.0)
tape.gradient(result, v)
```
You can use `Function`s inside other `Function`s.
```
@tf.function
def dense_layer(x, w, b):
return add(tf.matmul(x, w), b)
dense_layer(tf.ones([3, 2]), tf.ones([2, 2]), tf.ones([2]))
```
`Function`s can be faster than eager code, especially for graphs with many small ops. But for graphs with a few expensive ops (like convolutions), you may not see much speedup.
```
import timeit
conv_layer = tf.keras.layers.Conv2D(100, 3)
@tf.function
def conv_fn(image):
return conv_layer(image)
image = tf.zeros([1, 200, 200, 100])
# warm up
conv_layer(image); conv_fn(image)
print("Eager conv:", timeit.timeit(lambda: conv_layer(image), number=10))
print("Function conv:", timeit.timeit(lambda: conv_fn(image), number=10))
print("Note how there's not much difference in performance for convolutions")
```
### Tracing
This section exposes how `Function` works under the hood, including implementation details *which may change in the future*. However, once you understand why and when tracing happens, it's much easier to use `tf.function` effectively!
#### What is "tracing"?
A `Function` runs your program in a [TensorFlow Graph](https://www.tensorflow.org/guide/intro_to_graphs#what_are_graphs). However, a `tf.Graph` cannot represent all the things that you'd write in an eager TensorFlow program. For instance, Python supports polymorphism, but `tf.Graph` requires its inputs to have a specified data type and dimension. Or you may perform side tasks like reading command-line arguments, raising an error, or working with a more complex Python object; none of these things can run in a `tf.Graph`.
`Function` bridges this gap by separating your code in two stages:
1) In the first stage, referred to as "**tracing**", `Function` creates a new `tf.Graph`. Python code runs normally, but all TensorFlow operations (like adding two Tensors) are *deferred*: they are captured by the `tf.Graph` and not run.
2) In the second stage, a `tf.Graph` which contains everything that was deferred in the first stage is run. This stage is much faster than the tracing stage.
Depending on its inputs, `Function` will not always run the first stage when it is called. See ["Rules of tracing"](#rules_of_tracing) below to get a better sense of how it makes that determination. Skipping the first stage and only executing the second stage is what gives you TensorFlow's high performance.
When `Function` does decide to trace, the tracing stage is immediately followed by the second stage, so calling the `Function` both creates and runs the `tf.Graph`. Later you will see how you can run only the tracing stage with [`get_concrete_function`](#obtaining_concrete_functions).
When we pass arguments of different types into a `Function`, both stages are run:
```
@tf.function
def double(a):
print("Tracing with", a)
return a + a
print(double(tf.constant(1)))
print()
print(double(tf.constant(1.1)))
print()
print(double(tf.constant("a")))
print()
```
Note that if you repeatedly call a `Function` with the same argument type, TensorFlow will skip the tracing stage and reuse a previously traced graph, as the generated graph would be identical.
```
# This doesn't print 'Tracing with ...'
print(double(tf.constant("b")))
```
You can use `pretty_printed_concrete_signatures()` to see all of the available traces:
```
print(double.pretty_printed_concrete_signatures())
```
So far, you've seen that `tf.function` creates a cached, dynamic dispatch layer over TensorFlow's graph tracing logic. To be more specific about the terminology:
- A `tf.Graph` is the raw, language-agnostic, portable representation of a TensorFlow computation.
- A `ConcreteFunction` wraps a `tf.Graph`.
- A `Function` manages a cache of `ConcreteFunction`s and picks the right one for your inputs.
- `tf.function` wraps a Python function, returning a `Function` object.
- **Tracing** creates a `tf.Graph` and wraps it in a `ConcreteFunction`, also known as a **trace.**
#### Rules of tracing
A `Function` determines whether to reuse a traced `ConcreteFunction` by computing a **cache key** from an input's args and kwargs. A **cache key** is a key that identifies a `ConcreteFunction` based on the input args and kwargs of the `Function` call, according to the following rules (which may change):
- The key generated for a `tf.Tensor` argument is its shape and dtype.
- Starting in TensorFlow 2.3, the key generated for a `tf.Variable` argument is its `id()`.
- The key generated for a Python primitive is its value. The key generated for nested `dict`s, `list`s, `tuple`s, `namedtuple`s, and [`attr`](https://www.attrs.org/en/stable/)s is the flattened tuple. (As a result of this flattening, calling a concrete function with a different nesting structure than the one used during tracing will result in a TypeError).
- For all other Python types, the keys are based on the object `id()` so that methods are traced independently for each instance of a class.
Note: Cache keys are based on the `Function` input parameters so changes to global and [free variables](https://docs.python.org/3/reference/executionmodel.html#binding-of-names) alone will not create a new trace. See [this section](#depending_on_python_global_and_free_variables) for recommended practices when dealing with Python global and free variables.
#### Controlling retracing
Retracing, which is when your `Function` creates more than one trace, helps ensures that TensorFlow generates correct graphs for each set of inputs. However, tracing is an expensive operation! If your `Function` retraces a new graph for every call, you'll find that your code executes more slowly than if you didn't use `tf.function`.
To control the tracing behavior, you can use the following techniques:
- Specify `input_signature` in `tf.function` to limit tracing.
```
@tf.function(input_signature=(tf.TensorSpec(shape=[None], dtype=tf.int32),))
def next_collatz(x):
print("Tracing with", x)
return tf.where(x % 2 == 0, x // 2, 3 * x + 1)
print(next_collatz(tf.constant([1, 2])))
# We specified a 1-D tensor in the input signature, so this should fail.
with assert_raises(ValueError):
next_collatz(tf.constant([[1, 2], [3, 4]]))
# We specified an int32 dtype in the input signature, so this should fail.
with assert_raises(ValueError):
next_collatz(tf.constant([1.0, 2.0]))
```
- Specify a \[None\] dimension in `tf.TensorSpec` to allow for flexibility in trace reuse.
Since TensorFlow matches tensors based on their shape, using a `None` dimension as a wildcard will allow `Function`s to reuse traces for variably-sized input. Variably-sized input can occur if you have sequences of different length, or images of different sizes for each batch (See [Transformer](../tutorials/text/transformer.ipynb) and [Deep Dream](../tutorials/generative/deepdream.ipynb) tutorials for example).
```
@tf.function(input_signature=(tf.TensorSpec(shape=[None], dtype=tf.int32),))
def g(x):
print('Tracing with', x)
return x
# No retrace!
print(g(tf.constant([1, 2, 3])))
print(g(tf.constant([1, 2, 3, 4, 5])))
```
- Cast Python arguments to Tensors to reduce retracing.
Often, Python arguments are used to control hyperparameters and graph constructions - for example, `num_layers=10` or `training=True` or `nonlinearity='relu'`. So if the Python argument changes, it makes sense that you'd have to retrace the graph.
However, it's possible that a Python argument is not being used to control graph construction. In these cases, a change in the Python value can trigger needless retracing. Take, for example, this training loop, which AutoGraph will dynamically unroll. Despite the multiple traces, the generated graph is actually identical, so retracing is unnecessary.
```
def train_one_step():
pass
@tf.function
def train(num_steps):
print("Tracing with num_steps = ", num_steps)
tf.print("Executing with num_steps = ", num_steps)
for _ in tf.range(num_steps):
train_one_step()
print("Retracing occurs for different Python arguments.")
train(num_steps=10)
train(num_steps=20)
print()
print("Traces are reused for Tensor arguments.")
train(num_steps=tf.constant(10))
train(num_steps=tf.constant(20))
```
If you need to force retracing, create a new `Function`. Separate `Function` objects are guaranteed not to share traces.
```
def f():
print('Tracing!')
tf.print('Executing')
tf.function(f)()
tf.function(f)()
```
### Obtaining concrete functions
Every time a function is traced, a new concrete function is created. You can directly obtain a concrete function, by using `get_concrete_function`.
```
print("Obtaining concrete trace")
double_strings = double.get_concrete_function(tf.constant("a"))
print("Executing traced function")
print(double_strings(tf.constant("a")))
print(double_strings(a=tf.constant("b")))
# You can also call get_concrete_function on an InputSpec
double_strings_from_inputspec = double.get_concrete_function(tf.TensorSpec(shape=[], dtype=tf.string))
print(double_strings_from_inputspec(tf.constant("c")))
```
Printing a `ConcreteFunction` displays a summary of its input arguments (with types) and its output type.
```
print(double_strings)
```
You can also directly retrieve a concrete function's signature.
```
print(double_strings.structured_input_signature)
print(double_strings.structured_outputs)
```
Using a concrete trace with incompatible types will throw an error
```
with assert_raises(tf.errors.InvalidArgumentError):
double_strings(tf.constant(1))
```
You may notice that Python arguments are given special treatment in a concrete function's input signature. Prior to TensorFlow 2.3, Python arguments were simply removed from the concrete function's signature. Starting with TensorFlow 2.3, Python arguments remain in the signature, but are constrained to take the value set during tracing.
```
@tf.function
def pow(a, b):
return a ** b
square = pow.get_concrete_function(a=tf.TensorSpec(None, tf.float32), b=2)
print(square)
assert square(tf.constant(10.0)) == 100
with assert_raises(TypeError):
square(tf.constant(10.0), b=3)
```
### Obtaining graphs
Each concrete function is a callable wrapper around a `tf.Graph`. Although retrieving the actual `tf.Graph` object is not something you'll normally need to do, you can obtain it easily from any concrete function.
```
graph = double_strings.graph
for node in graph.as_graph_def().node:
print(f'{node.input} -> {node.name}')
```
### Debugging
In general, debugging code is easier in eager mode than inside `tf.function`. You should ensure that your code executes error-free in eager mode before decorating with `tf.function`. To assist in the debugging process, you can call `tf.config.run_functions_eagerly(True)` to globally disable and reenable `tf.function`.
When tracking down issues that only appear within `tf.function`, here are some tips:
- Plain old Python `print` calls only execute during tracing, helping you track down when your function gets (re)traced.
- `tf.print` calls will execute every time, and can help you track down intermediate values during execution.
- `tf.debugging.enable_check_numerics` is an easy way to track down where NaNs and Inf are created.
- `pdb` can help you understand what's going on during tracing. (Caveat: PDB will drop you into AutoGraph-transformed source code.)
## AutoGraph Transformations
AutoGraph is a library that is on by default in `tf.function`, and transforms a subset of Python eager code into graph-compatible TensorFlow ops. This includes control flow like `if`, `for`, `while`.
TensorFlow ops like `tf.cond` and `tf.while_loop` continue to work, but control flow is often easier to write and understand when written in Python.
```
# Simple loop
@tf.function
def f(x):
while tf.reduce_sum(x) > 1:
tf.print(x)
x = tf.tanh(x)
return x
f(tf.random.uniform([5]))
```
If you're curious you can inspect the code autograph generates.
```
print(tf.autograph.to_code(f.python_function))
```
### Conditionals
AutoGraph will convert some `if <condition>` statements into the equivalent `tf.cond` calls. This substitution is made if `<condition>` is a Tensor. Otherwise, the `if` statement is executed as a Python conditional.
A Python conditional executes during tracing, so exactly one branch of the conditional will be added to the graph. Without AutoGraph, this traced graph would be unable to take the alternate branch if there is data-dependent control flow.
`tf.cond` traces and adds both branches of the conditional to the graph, dynamically selecting a branch at execution time. Tracing can have unintended side effects; see [AutoGraph tracing effects](https://github.com/tensorflow/tensorflow/blob/master/tensorflow/python/autograph/g3doc/reference/control_flow.md#effects-of-the-tracing-process) for more.
```
@tf.function
def fizzbuzz(n):
for i in tf.range(1, n + 1):
print('Tracing for loop')
if i % 15 == 0:
print('Tracing fizzbuzz branch')
tf.print('fizzbuzz')
elif i % 3 == 0:
print('Tracing fizz branch')
tf.print('fizz')
elif i % 5 == 0:
print('Tracing buzz branch')
tf.print('buzz')
else:
print('Tracing default branch')
tf.print(i)
fizzbuzz(tf.constant(5))
fizzbuzz(tf.constant(20))
```
See the [reference documentation](https://github.com/tensorflow/tensorflow/blob/master/tensorflow/python/autograph/g3doc/reference/control_flow.md#if-statements) for additional restrictions on AutoGraph-converted if statements.
### Loops
AutoGraph will convert some `for` and `while` statements into the equivalent TensorFlow looping ops, like `tf.while_loop`. If not converted, the `for` or `while` loop is executed as a Python loop.
This substitution is made in the following situations:
- `for x in y`: if `y` is a Tensor, convert to `tf.while_loop`. In the special case where `y` is a `tf.data.Dataset`, a combination of `tf.data.Dataset` ops are generated.
- `while <condition>`: if `<condition>` is a Tensor, convert to `tf.while_loop`.
A Python loop executes during tracing, adding additional ops to the `tf.Graph` for every iteration of the loop.
A TensorFlow loop traces the body of the loop, and dynamically selects how many iterations to run at execution time. The loop body only appears once in the generated `tf.Graph`.
See the [reference documentation](https://github.com/tensorflow/tensorflow/blob/master/tensorflow/python/autograph/g3doc/reference/control_flow.md#while-statements) for additional restrictions on AutoGraph-converted `for` and `while` statements.
#### Looping over Python data
A common pitfall is to loop over Python/Numpy data within a `tf.function`. This loop will execute during the tracing process, adding a copy of your model to the `tf.Graph` for each iteration of the loop.
If you want to wrap the entire training loop in `tf.function`, the safest way to do this is to wrap your data as a `tf.data.Dataset` so that AutoGraph will dynamically unroll the training loop.
```
def measure_graph_size(f, *args):
g = f.get_concrete_function(*args).graph
print("{}({}) contains {} nodes in its graph".format(
f.__name__, ', '.join(map(str, args)), len(g.as_graph_def().node)))
@tf.function
def train(dataset):
loss = tf.constant(0)
for x, y in dataset:
loss += tf.abs(y - x) # Some dummy computation.
return loss
small_data = [(1, 1)] * 3
big_data = [(1, 1)] * 10
measure_graph_size(train, small_data)
measure_graph_size(train, big_data)
measure_graph_size(train, tf.data.Dataset.from_generator(
lambda: small_data, (tf.int32, tf.int32)))
measure_graph_size(train, tf.data.Dataset.from_generator(
lambda: big_data, (tf.int32, tf.int32)))
```
When wrapping Python/Numpy data in a Dataset, be mindful of `tf.data.Dataset.from_generator` versus ` tf.data.Dataset.from_tensors`. The former will keep the data in Python and fetch it via `tf.py_function` which can have performance implications, whereas the latter will bundle a copy of the data as one large `tf.constant()` node in the graph, which can have memory implications.
Reading data from files via TFRecordDataset/CsvDataset/etc. is the most effective way to consume data, as then TensorFlow itself can manage the asynchronous loading and prefetching of data, without having to involve Python. To learn more, see the [tf.data guide](../../guide/data).
#### Accumulating values in a loop
A common pattern is to accumulate intermediate values from a loop. Normally, this is accomplished by appending to a Python list or adding entries to a Python dictionary. However, as these are Python side effects, they will not work as expected in a dynamically unrolled loop. Use `tf.TensorArray` to accumulate results from a dynamically unrolled loop.
```
batch_size = 2
seq_len = 3
feature_size = 4
def rnn_step(inp, state):
return inp + state
@tf.function
def dynamic_rnn(rnn_step, input_data, initial_state):
# [batch, time, features] -> [time, batch, features]
input_data = tf.transpose(input_data, [1, 0, 2])
max_seq_len = input_data.shape[0]
states = tf.TensorArray(tf.float32, size=max_seq_len)
state = initial_state
for i in tf.range(max_seq_len):
state = rnn_step(input_data[i], state)
states = states.write(i, state)
return tf.transpose(states.stack(), [1, 0, 2])
dynamic_rnn(rnn_step,
tf.random.uniform([batch_size, seq_len, feature_size]),
tf.zeros([batch_size, feature_size]))
```
## Limitations
TensorFlow `Function` has a few limitations by design that you should be aware of when converting a Python function to a `Function`.
### Executing Python side effects
Side effects, like printing, appending to lists, and mutating globals, can behave unexpectedly inside a `Function`, sometimes executing twice or not all. They only happen the first time you call a `Function` with a set of inputs. Afterwards, the traced `tf.Graph` is reexecuted, without executing the Python code.
The general rule of thumb is to avoid relying on Python side effects in your logic and only use them to debug your traces. Otherwise, TensorFlow APIs like `tf.data`, `tf.print`, `tf.summary`, `tf.Variable.assign`, and `tf.TensorArray` are the best way to ensure your code will be executed by the TensorFlow runtime with each call.
```
@tf.function
def f(x):
print("Traced with", x)
tf.print("Executed with", x)
f(1)
f(1)
f(2)
```
If you would like to execute Python code during each invocation of a `Function`, `tf.py_function` is an exit hatch. The drawback of `tf.py_function` is that it's not portable or particularly performant, cannot be saved with SavedModel, and does not work well in distributed (multi-GPU, TPU) setups. Also, since `tf.py_function` has to be wired into the graph, it casts all inputs/outputs to tensors.
#### Changing Python global and free variables
Changing Python global and [free variables](https://docs.python.org/3/reference/executionmodel.html#binding-of-names) counts as a Python side effect, so it only happens during tracing.
```
external_list = []
@tf.function
def side_effect(x):
print('Python side effect')
external_list.append(x)
side_effect(1)
side_effect(1)
side_effect(1)
# The list append only happened once!
assert len(external_list) == 1
```
You should avoid mutating containers like lists, dicts, other objects that live outside the `Function`. Instead, use arguments and TF objects. For example, the section ["Accumulating values in a loop"](#accumulating_values_in_a_loop) has one example of how list-like operations can be implemented.
You can, in some cases, capture and manipulate state if it is a [`tf.Variable`](https://www.tensorflow.org/guide/variable). This is how the weights of Keras models are updated with repeated calls to the same `ConcreteFunction`.
#### Using Python iterators and generators
Many Python features, such as generators and iterators, rely on the Python runtime to keep track of state. In general, while these constructs work as expected in eager mode, they are examples of Python side effects and therefore only happen during tracing.
```
@tf.function
def buggy_consume_next(iterator):
tf.print("Value:", next(iterator))
iterator = iter([1, 2, 3])
buggy_consume_next(iterator)
# This reuses the first value from the iterator, rather than consuming the next value.
buggy_consume_next(iterator)
buggy_consume_next(iterator)
```
Just like how TensorFlow has a specialized `tf.TensorArray` for list constructs, it has a specialized `tf.data.Iterator` for iteration constructs. See the section on [AutoGraph Transformations](#autograph_transformations) for an overview. Also, the [`tf.data`](https://www.tensorflow.org/guide/data) API can help implement generator patterns:
```
@tf.function
def good_consume_next(iterator):
# This is ok, iterator is a tf.data.Iterator
tf.print("Value:", next(iterator))
ds = tf.data.Dataset.from_tensor_slices([1, 2, 3])
iterator = iter(ds)
good_consume_next(iterator)
good_consume_next(iterator)
good_consume_next(iterator)
```
### Deleting tf.Variables between `Function` calls
Another error you may encounter is a garbage-collected variable. `ConcreteFunction`s only retain [WeakRefs](https://docs.python.org/3/library/weakref.html) to the variables they close over, so you must retain a reference to any variables.
```
external_var = tf.Variable(3)
@tf.function
def f(x):
return x * external_var
traced_f = f.get_concrete_function(4)
print("Calling concrete function...")
print(traced_f(4))
# The original variable object gets garbage collected, since there are no more
# references to it.
external_var = tf.Variable(4)
print()
print("Calling concrete function after garbage collecting its closed Variable...")
with assert_raises(tf.errors.FailedPreconditionError):
traced_f(4)
```
## Known Issues
If your `Function` is not evaluating correctly, the error may be explained by these known issues which are planned to be fixed in the future.
### Depending on Python global and free variables
`Function` creates a new `ConcreteFunction` when called with a new value of a Python argument. However, it does not do that for the Python closure, globals, or nonlocals of that `Function`. If their value changes in between calls to the `Function`, the `Function` will still use the values they had when it was traced. This is different from how regular Python functions work.
For that reason, we recommend a functional programming style that uses arguments instead of closing over outer names.
```
@tf.function
def buggy_add():
return 1 + foo
@tf.function
def recommended_add(foo):
return 1 + foo
foo = 1
print("Buggy:", buggy_add())
print("Correct:", recommended_add(foo))
print("Updating the value of `foo` to 100!")
foo = 100
print("Buggy:", buggy_add()) # Did not change!
print("Correct:", recommended_add(foo))
```
You can close over outer names, as long as you don't update their values.
#### Depending on Python objects
The recommendation to pass Python objects as arguments into `tf.function` has a number of known issues, that are expected to be fixed in the future. In general, you can rely on consistent tracing if you use a Python primitive or `tf.nest`-compatible structure as an argument or pass in a *different* instance of an object into a `Function`. However, `Function` will *not* create a new trace when you pass **the same object and only change its attributes**.
```
class SimpleModel(tf.Module):
def __init__(self):
# These values are *not* tf.Variables.
self.bias = 0.
self.weight = 2.
@tf.function
def evaluate(model, x):
return model.weight * x + model.bias
simple_model = SimpleModel()
x = tf.constant(10.)
print(evaluate(simple_model, x))
print("Adding bias!")
simple_model.bias += 5.0
print(evaluate(simple_model, x)) # Didn't change :(
```
Using the same `Function` to evaluate the updated instance of the model will be buggy since the updated model has the [same cache key](#rules_of_tracing) as the original model.
For that reason, we recommend that you write your `Function` to avoid depending on mutable object attributes or create new objects.
If that is not possible, one workaround is to make new `Function`s each time you modify your object to force retracing:
```
def evaluate(model, x):
return model.weight * x + model.bias
new_model = SimpleModel()
evaluate_no_bias = tf.function(evaluate).get_concrete_function(new_model, x)
# Don't pass in `new_model`, `Function` already captured its state during tracing.
print(evaluate_no_bias(x))
print("Adding bias!")
new_model.bias += 5.0
# Create new Function and ConcreteFunction since you modified new_model.
evaluate_with_bias = tf.function(evaluate).get_concrete_function(new_model, x)
print(evaluate_with_bias(x)) # Don't pass in `new_model`.
```
As [retracing can be expensive](https://www.tensorflow.org/guide/intro_to_graphs#tracing_and_performance), you can use `tf.Variable`s as object attributes, which can be mutated (but not changed, careful!) for a similar effect without needing a retrace.
```
class BetterModel:
def __init__(self):
self.bias = tf.Variable(0.)
self.weight = tf.Variable(2.)
@tf.function
def evaluate(model, x):
return model.weight * x + model.bias
better_model = BetterModel()
print(evaluate(better_model, x))
print("Adding bias!")
better_model.bias.assign_add(5.0) # Note: instead of better_model.bias += 5
print(evaluate(better_model, x)) # This works!
```
### Creating tf.Variables
`Function` only supports creating variables once, when first called, and then reusing them. You cannot create `tf.Variables` in new traces. Creating new variables in subsequent calls is currently not allowed, but will be in the future.
Example:
```
@tf.function
def f(x):
v = tf.Variable(1.0)
return v
with assert_raises(ValueError):
f(1.0)
```
You can create variables inside a `Function` as long as those variables are only created the first time the function is executed.
```
class Count(tf.Module):
def __init__(self):
self.count = None
@tf.function
def __call__(self):
if self.count is None:
self.count = tf.Variable(0)
return self.count.assign_add(1)
c = Count()
print(c())
print(c())
```
#### Using with multiple Keras optimizers
You may encounter `ValueError: tf.function-decorated function tried to create variables on non-first call.` when using more than one Keras optimizer with a `tf.function`. This error occurs because optimizers internally create `tf.Variables` when they apply gradients for the first time.
```
opt1 = tf.keras.optimizers.Adam(learning_rate = 1e-2)
opt2 = tf.keras.optimizers.Adam(learning_rate = 1e-3)
@tf.function
def train_step(w, x, y, optimizer):
with tf.GradientTape() as tape:
L = tf.reduce_sum(tf.square(w*x - y))
gradients = tape.gradient(L, [w])
optimizer.apply_gradients(zip(gradients, [w]))
w = tf.Variable(2.)
x = tf.constant([-1.])
y = tf.constant([2.])
train_step(w, x, y, opt1)
print("Calling `train_step` with different optimizer...")
with assert_raises(ValueError):
train_step(w, x, y, opt2)
```
If you need to change the optimizer during training, a workaround is to create a new `Function` for each optimizer, calling the [`ConcreteFunction`](#obtaining_concrete_functions) directly.
```
opt1 = tf.keras.optimizers.Adam(learning_rate = 1e-2)
opt2 = tf.keras.optimizers.Adam(learning_rate = 1e-3)
# Not a tf.function.
def train_step(w, x, y, optimizer):
with tf.GradientTape() as tape:
L = tf.reduce_sum(tf.square(w*x - y))
gradients = tape.gradient(L, [w])
optimizer.apply_gradients(zip(gradients, [w]))
w = tf.Variable(2.)
x = tf.constant([-1.])
y = tf.constant([2.])
# Make a new Function and ConcreteFunction for each optimizer.
train_step_1 = tf.function(train_step).get_concrete_function(w, x, y, opt1)
train_step_2 = tf.function(train_step).get_concrete_function(w, x, y, opt2)
for i in range(10):
if i % 2 == 0:
train_step_1(w, x, y) # `opt1` is not used as a parameter.
else:
train_step_2(w, x, y) # `opt2` is not used as a parameter.
```
#### Using with multiple Keras models
You may also encounter `ValueError: tf.function-decorated function tried to create variables on non-first call.` when passing different model instances to the same `Function`.
This error occurs because Keras models (which [do not have their input shape defined](https://www.tensorflow.org/guide/keras/custom_layers_and_models#best_practice_deferring_weight_creation_until_the_shape_of_the_inputs_is_known)) and Keras layers create `tf.Variables`s when they are first called. You may be attempting to initialize those variables inside a `Function`, which has already been called. To avoid this error, try calling `model.build(input_shape)` to initialize all the weights before training the model.
## Further reading
To learn about how to export and load a `Function`, see the [SavedModel guide](../../guide/saved_model). To learn more about graph optimizations that are performed after tracing, see the [Grappler guide](../../guide/graph_optimization). To learn how to optimize your data pipeline and profile your model, see the [Profiler guide](../../guide/profiler.md).
| github_jupyter |
```
import requests
from bs4 import BeautifulSoup as soup
import re
import urllib.parse
from urllib.request import Request, urlopen
from urllib.parse import urlparse
import random
import json
def randomAgent():
randomAgents = [
'Mozilla/5.0 (Macintosh; U; Intel Mac OS X 10.5; en-US; rv:1.9.1b3) Gecko/20090305 Firefox/3.1b3 GTB5',
'Mozilla/5.0 (Macintosh; U; Intel Mac OS X 10.5; ko; rv:1.9.1b2) Gecko/20081201 Firefox/3.1b2' ,
'Mozilla/5.0 (X11; U; SunOS sun4u; en-US; rv:1.9b5) Gecko/2008032620 Firefox/3.0b5',
'Mozilla/5.0 (Windows NT 6.1; WOW64; rv:2.0b8pre) Gecko/20101114 Firefox/4.0b8pre',
'Mozilla/5.0 (X11; Linux x86_64; rv:2.0b9pre) Gecko/20110111 Firefox/4.0b9pre'
'Mozilla/5.0 (Windows NT 6.1; Win64; x64; rv:2.0b9pre) Gecko/20101228 Firefox/4.0b9pre'
'Mozilla/5.0 (Windows NT 6.1; Win64; x64; rv:2.2a1pre) Gecko/20110324 Firefox/4.2a1pre'
'Mozilla/5.0 (X11; U; Linux amd64; rv:5.0) Gecko/20100101 Firefox/5.0 (Debian)'
'Mozilla/5.0 (Windows NT 6.1; WOW64; rv:6.0a2) Gecko/20110613 Firefox/6.0a2'
'Mozilla/5.0 (X11; Linux i686 on x86_64; rv:12.0) Gecko/20100101 Firefox/12.0'
]
return random.choice(randomAgents)
def scrape(url):
req = Request(url, headers={'User-Agent' : randomAgent()})
webpage = urlopen(req).read()
return webpage
wiki_skeletal_muscles = 'https://en.wikipedia.org/wiki/List_of_skeletal_muscles_of_the_human_body'
page = scrape(wiki_skeletal_muscles)
def scrapeByTag(webpage, tag):
page_soup = soup(webpage,'html.parser')
taglist = page_soup.findAll(tag)
return taglist
def scrapeByClassName(webpage, tag, classname):
page_soup = soup(webpage,'html.parser')
classlist = page_soup.findAll(tag, class_=classname)
return classlist
def formatText(text, i):
text = str(text)
cleaned = re.sub(r"<{}>|<{}>", "", text).format(i)
return cleaned
def toJSON(title, obj, location):
with open(location + '/' + title + '.json', 'w', encoding='utf-8') as f:
json.dump(obj, f, ensure_ascii=False, indent=4)
return
tables = scrapeByClassName(page, 'table', 'wikitable')
def getHeaders(head):
headings = []
for item in head.find_all("th"):
item = (item.text).rstrip("\n")
headings.append(item)
return headings
def getRows(rows):
all_rows = []
for row_num in range(len(rows)):
row = []
for row_item in rows[row_num].find_all("td"):
aa = re.sub("(\xa0)|(\n)|,","",row_item.text)
row.append(aa)
all_rows.append(row)
return all_rows
def createObjects(headers, rows):
objects = []
for row in rows:
if len(row) == len(headers):
obj = {}
for i in range(len(headers)):
title = headers[i]
data = row[i]
obj[title] = data
objects.append(obj)
else:
continue
return objects
def extractMusclesData(page):
muscles = []
tables = scrapeByClassName(page, 'table', 'wikitable')
for table in tables:
body = table.find_all("tr")
muscles += createObjects(getHeaders(body[0]), getRows(body[1:]))
return muscles
toJSON('skeletalMuscles', extractMusclesData(page), 'Muscles')
```
| github_jupyter |
```
%matplotlib inline
```
# Dimensionality Reduction with Neighborhood Components Analysis
Sample usage of Neighborhood Components Analysis for dimensionality reduction.
This example compares different (linear) dimensionality reduction methods
applied on the Digits data set. The data set contains images of digits from
0 to 9 with approximately 180 samples of each class. Each image is of
dimension 8x8 = 64, and is reduced to a two-dimensional data point.
Principal Component Analysis (PCA) applied to this data identifies the
combination of attributes (principal components, or directions in the
feature space) that account for the most variance in the data. Here we
plot the different samples on the 2 first principal components.
Linear Discriminant Analysis (LDA) tries to identify attributes that
account for the most variance *between classes*. In particular,
LDA, in contrast to PCA, is a supervised method, using known class labels.
Neighborhood Components Analysis (NCA) tries to find a feature space such
that a stochastic nearest neighbor algorithm will give the best accuracy.
Like LDA, it is a supervised method.
One can see that NCA enforces a clustering of the data that is visually
meaningful despite the large reduction in dimension.
Adapted from `<https://scikit-learn.org/stable/auto_examples/neighbors/plot_nca_dim_reduction.html>`_
```
# License: BSD 3 clause
import numpy as np
import matplotlib.pyplot as plt
from sklearn import datasets
from sklearn.model_selection import train_test_split
from sklearn.decomposition import PCA
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import StandardScaler
from skhubness.neighbors import (KNeighborsClassifier,
NeighborhoodComponentsAnalysis)
print(__doc__)
n_neighbors = 3
random_state = 0
# Load Digits dataset
digits = datasets.load_digits()
X, y = digits.data, digits.target
# Split into train/test
X_train, X_test, y_train, y_test = \
train_test_split(X, y, test_size=0.5, stratify=y,
random_state=random_state)
dim = len(X[0])
n_classes = len(np.unique(y))
# Reduce dimension to 2 with PCA
pca = make_pipeline(StandardScaler(),
PCA(n_components=2, random_state=random_state))
# Reduce dimension to 2 with LinearDiscriminantAnalysis
lda = make_pipeline(StandardScaler(),
LinearDiscriminantAnalysis(n_components=2))
# Reduce dimension to 2 with NeighborhoodComponentAnalysis
nca = make_pipeline(StandardScaler(),
NeighborhoodComponentsAnalysis(n_components=2,
random_state=random_state))
# Use a nearest neighbor classifier to evaluate the methods
knn = KNeighborsClassifier(n_neighbors=n_neighbors)
# Make a list of the methods to be compared
dim_reduction_methods = [('PCA', pca), ('LDA', lda), ('NCA', nca)]
# plt.figure()
for i, (name, model) in enumerate(dim_reduction_methods):
plt.figure()
# plt.subplot(1, 3, i + 1, aspect=1)
# Fit the method's model
model.fit(X_train, y_train)
# Fit a nearest neighbor classifier on the embedded training set
knn.fit(model.transform(X_train), y_train)
# Compute the nearest neighbor accuracy on the embedded test set
acc_knn = knn.score(model.transform(X_test), y_test)
# Embed the data set in 2 dimensions using the fitted model
X_embedded = model.transform(X)
# Plot the projected points and show the evaluation score
plt.scatter(X_embedded[:, 0], X_embedded[:, 1], c=y, s=30, cmap='Set1')
plt.title("{}, KNN (k={})\nTest accuracy = {:.2f}".format(name,
n_neighbors,
acc_knn))
plt.show()
```
| github_jupyter |
# Logistic Regression with MinMaxScaler
### Required Packages
```
!pip install imblearn
import warnings
import numpy as np
import pandas as pd
import seaborn as se
import matplotlib.pyplot as plt
from imblearn.over_sampling import RandomOverSampler
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import LabelEncoder,MinMaxScaler
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import classification_report,plot_confusion_matrix
warnings.filterwarnings('ignore')
```
### Initialization
Filepath of CSV file
```
#filepath
file_path= ""
```
List of features which are required for model training .
```
#x_values
features=[]
```
Target feature for prediction.
```
#y_value
target=''
```
### Data Fetching
Pandas is an open-source, BSD-licensed library providing high-performance, easy-to-use data manipulation and data analysis tools.
We will use panda's library to read the CSV file using its storage path.And we use the head function to display the initial row or entry.
```
df=pd.read_csv(file_path)
df.head()
```
### Feature Selections
It is the process of reducing the number of input variables when developing a predictive model. Used to reduce the number of input variables to both reduce the computational cost of modelling and, in some cases, to improve the performance of the model.
We will assign all the required input features to X and target/outcome to Y.
```
X = df[features]
Y = df[target]
```
### Data Preprocessing
Since the majority of the machine learning models in the Sklearn library doesn't handle string category data and Null value, we have to explicitly remove or replace null values. The below snippet have functions, which removes the null value if any exists. And convert the string classes data in the datasets by encoding them to integer classes.
```
def NullClearner(df):
if(isinstance(df, pd.Series) and (df.dtype in ["float64","int64"])):
df.fillna(df.mean(),inplace=True)
return df
elif(isinstance(df, pd.Series)):
df.fillna(df.mode()[0],inplace=True)
return df
else:return df
def EncodeX(df):
return pd.get_dummies(df)
def EncodeY(df):
if len(df.unique())<=2:
return df
else:
un_EncodedT=np.sort(pd.unique(df), axis=-1, kind='mergesort')
df=LabelEncoder().fit_transform(df)
EncodedT=[xi for xi in range(len(un_EncodedT))]
print("Encoded Target: {} to {}".format(un_EncodedT,EncodedT))
return df
x=X.columns.to_list()
for i in x:
X[i]=NullClearner(X[i])
X=EncodeX(X)
Y=EncodeY(NullClearner(Y))
X.head()
```
#### Correlation Map
In order to check the correlation between the features, we will plot a correlation matrix. It is effective in summarizing a large amount of data where the goal is to see patterns.
```
f,ax = plt.subplots(figsize=(18, 18))
matrix = np.triu(X.corr())
se.heatmap(X.corr(), annot=True, linewidths=.5, fmt= '.1f',ax=ax, mask=matrix)
plt.show()
```
#### Distribution Of Target Variable
```
plt.figure(figsize = (10,6))
se.countplot(Y)
```
### Data Splitting
The train-test split is a procedure for evaluating the performance of an algorithm. The procedure involves taking a dataset and dividing it into two subsets. The first subset is utilized to fit/train the model. The second subset is used for prediction. The main motive is to estimate the performance of the model on new data.
```
x_train,x_test,y_train,y_test=train_test_split(X,Y,test_size=0.2,random_state=123)
```
#### Handling Target Imbalance
The challenge of working with imbalanced datasets is that most machine learning techniques will ignore, and in turn have poor performance on, the minority class, although typically it is performance on the minority class that is most important.
One approach to addressing imbalanced datasets is to oversample the minority class. The simplest approach involves duplicating examples in the minority class.We will perform overspampling using imblearn library.
```
x_train,y_train = RandomOverSampler(random_state=123).fit_resample(x_train, y_train)
```
### Data Rescaling
Transform features by scaling each feature to a given range.
This estimator scales and translates each feature individually such that it is in the given range on the training set, e.g. between zero and one.
[More on MinMaxScaler module and parameters](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.MinMaxScaler.html)
### Model
Logistic regression is a statistical model that in its basic form uses a logistic function to model a binary dependent variable, although many more complex extensions exist. In regression analysis, logistic regression (or logit regression) is estimating the parameters of a logistic model (a form of binary regression). This can be extended to model several classes of events.
#### Model Tuning Parameters
1. penalty : {‘l1’, ‘l2’, ‘elasticnet’, ‘none’}, default=’l2’
> Used to specify the norm used in the penalization. The ‘newton-cg’, ‘sag’ and ‘lbfgs’ solvers support only l2 penalties. ‘elasticnet’ is only supported by the ‘saga’ solver. If ‘none’ (not supported by the liblinear solver), no regularization is applied.
2. C : float, default=1.0
> Inverse of regularization strength; must be a positive float. Like in support vector machines, smaller values specify stronger regularization.
3. tol : float, default=1e-4
> Tolerance for stopping criteria.
4. solver : {‘newton-cg’, ‘lbfgs’, ‘liblinear’, ‘sag’, ‘saga’}, default=’lbfgs’
> Algorithm to use in the optimization problem.
For small datasets, ‘<code>liblinear</code>’ is a good choice, whereas ‘<code>sag</code>’ and ‘<code>saga</code>’ are faster for large ones.
For multiclass problems, only ‘<code>newton-cg</code>’, ‘<code>sag</code>’, ‘<code>saga</code>’ and ‘<code>lbfgs</code>’ handle multinomial loss; ‘<code>liblinear</code>’ is limited to one-versus-rest schemes.
* ‘<code>newton-cg</code>’, ‘<code>lbfgs</code>’, ‘<code>sag</code>’ and ‘<code>saga</code>’ handle L2 or no penalty.
* ‘<code>liblinear</code>’ and ‘<code>saga</code>’ also handle L1 penalty.
* ‘<code>saga</code>’ also supports ‘<code>elasticnet</code>’ penalty.
* ‘<code>liblinear</code>’ does not support setting <code>penalty='none'</code>.
5. random_state : int, RandomState instance, default=None
> Used when <code>solver</code> == ‘sag’, ‘saga’ or ‘liblinear’ to shuffle the data.
6. max_iter : int, default=100
> Maximum number of iterations taken for the solvers to converge.
7. multi_class : {‘auto’, ‘ovr’, ‘multinomial’}, default=’auto’
> If the option chosen is ‘<code>ovr</code>’, then a binary problem is fit for each label. For ‘<code>multinomial</code>’ the loss minimised is the multinomial loss fit across the entire probability distribution, even when the data is binary. ‘<code>multinomial</code>’ is unavailable when <code>solver</code>=’<code>liblinear</code>’. ‘auto’ selects ‘ovr’ if the data is binary, or if <code>solver</code>=’<code>liblinear</code>’, and otherwise selects ‘<code>multinomial</code>’.
8. verbose : int, default=0
> For the liblinear and lbfgs solvers set verbose to any positive number for verbosity.
9. n_jobs : int, default=None
> Number of CPU cores used when parallelizing over classes if multi_class=’ovr’”. This parameter is ignored when the <code>solver</code> is set to ‘liblinear’ regardless of whether ‘multi_class’ is specified or not. <code>None</code> means 1 unless in a joblib.parallel_backend context. <code>-1</code> means using all processors
```
# Build Model here
model = make_pipeline(MinMaxScaler(),LogisticRegression())
model.fit(x_train, y_train)
```
#### Model Accuracy
score() method return the mean accuracy on the given test data and labels.
In multi-label classification, this is the subset accuracy which is a harsh metric since you require for each sample that each label set be correctly predicted.
```
print("Accuracy score {:.2f} %\n".format(model.score(x_test,y_test)*100))
```
#### Confusion Matrix
A confusion matrix is utilized to understand the performance of the classification model or algorithm in machine learning for a given test set where results are known.
```
plot_confusion_matrix(model,x_test,y_test,cmap=plt.cm.Blues)
```
#### Classification Report
A Classification report is used to measure the quality of predictions from a classification algorithm. How many predictions are True, how many are False.
* **where**:
- Precision:- Accuracy of positive predictions.
- Recall:- Fraction of positives that were correctly identified.
- f1-score:- percent of positive predictions were correct
- support:- Support is the number of actual occurrences of the class in the specified dataset.
```
print(classification_report(y_test,model.predict(x_test)))
```
#### Creator: Nikhil Shrotri , Github: [Profile](https://github.com/nikhilshrotri/)
| github_jupyter |
```
import time
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits.axes_grid1 import make_axes_locatable
from matplotlib.ticker import MaxNLocator
plt.rcParams['mathtext.fontset'] = 'stix'
import functools
import re
from multiprocessing import Pool
import itertools
```
### Defining labels and corresponding state positions
```
# states
L = -5.4
R = 5.4
# index
bins=np.arange(-15., 17, 1)
idx_L=np.digitize(L, bins)
idx_R=np.digitize(R, bins)
```
### Defining functions for counting transitions
```
def within(coordinate, center, radius):
"""
Determine whether a coordinate is within the predefined center.
"""
return np.linalg.norm(coordinate - center) <= radius
def classify(coordinate, radius):
"""
Classify coordinates as labels.
"""
if within(coordinate, idx_L, radius):
return "L"
elif within(coordinate, idx_R, radius):
return "R"
else:
return "N"
def count(data, commit):
# Convert 1*commit A,B,C by a,b,c
data = data.replace('L' * commit, 'l')
data = data.replace('R' * commit, 'r')
# Remove remained A,B,C, and N.
data = data.replace('L', "") # Remove all A's
data = data.replace('R', '')
data = data.replace('N', '')
# Convert a,b,c back to A,B,C. I think it should be okay to process with just X,Y,Z
data = data.replace('l', 'L')
data = data.replace('r', 'R')
# Now note here that each A,B,C are actually represent 1*commit A,B,C. AA+ means there are at least 2*commit A.
# Here replacing AA+ by A.
data = re.sub('LL+', 'L', data)
data = re.sub('RR+', 'R', data)
n1 = data.count("LR")
n2 = data.count("RL")
return n1, n2
def find_plateau(count_arr):
"""
The input should be just one count arr, e.g. res[i].
How do I determine a plateau?
1. Plateau with commit time > 10,
2. Plateau should be at least 2-step long.
"""
for i in range(5,len(count_arr)-1):
if count_arr[i+1]==count_arr[i]:
return count_arr[i]
return count_arr[-1]
def count_arr(data_array):
"""
Counts vs commit times from the input data_array.
"""
# Choose the range of commit time. L=10 and R=21, so transition must be at least 5 steps.
commits = list(range(5,20, 1))
# Convert data_array to "L" and "R"
data = "".join(list(map(functools.partial(classify, radius=1), data_array)))
# Count transitions vs commit times
count_partial = functools.partial(count, data)
res = list(map(count_partial, commits))
res = list(zip(*res))
return res
```
### Read data
```
pred2=[]
for i in range(10):
pdfile = './N_mean0_42/Output-conc/{}/prediction.npy'.format(i)
prediction2 = np.load(pdfile)
pred2.append(prediction2)
```
### Analyze
```
# Pick one segment from data
i=17
segment_len=200000
data_array=pred2[0][i*segment_len:(i+1)*segment_len]
# Choose the range of commit time. L=10 and R=21, so transition must be at least 5 steps.
commits = list(range(5,20, 1))
# Convert data_array to "L" and "R"
data = "".join(list(map(functools.partial(classify, radius=1), data_array)))
# Count transitions vs commit times
count2 = functools.partial(count, data)
p = Pool(4)
res = list(p.map(count2, commits))
res = list(zip(*res))
```
### Make commit time plot
```
plt.rcParams["figure.figsize"] = (15,3)
plt.plot(data_array);
fig, ax=plt.subplots(figsize=(12,4), nrows=1, ncols=2)
ax=ax.flatten()
pairs = ['LR', 'RL']
ax[0].plot(commits, res[0], label='{}'.format(pairs[0]))
ax[1].plot(commits, res[1], label='{}'.format(pairs[1]))
ax[0].tick_params(axis='both', which='both', direction='in', labelsize=16)
ax[0].set_xlabel('Commit time (steps)', size=16)
ax[0].legend(loc='upper right', fontsize=16)
ax[0].set_ylabel('Count', size=16)
ax[0].set_yticks(np.arange(20))
ax[0].set_ylim(0,15)
ax[0].grid()
ax[1].tick_params(axis='both', which='both', direction='in', labelsize=16)
ax[1].set_xlabel('Commit time (steps)', size=16)
ax[1].legend(loc='upper right', fontsize=16)
ax[1].set_ylabel('Count', size=16)
ax[1].set_yticks(np.arange(20))
ax[1].set_ylim(0,15)
ax[1].grid()
fig.tight_layout()
plt.show()
```
### Compute for one trajectory
```
N_val = {}
# This for loop can be paralleled.
for i in range(20):
# Pick one segment from data
segment_len=200000
data_array = pred2[0][i*segment_len:(i+1)*segment_len]
N_val[i] = list(map(find_plateau, count_arr(data_array)))
N_arr = np.array(list(N_val.values()))
t_arr = segment_len/N_arr
# Mean transition time
mean_t = len(pred2[0])*0.05/np.sum(N_arr, axis=0)
stdv_t = np.std(t_arr*0.05, axis=0)/np.sqrt(len(N_arr))
```
### Transition time ($ps$)
```
mean_t, stdv_t
```
### Calculate averaged over 10 independent training and runs
```
def traj_i(traj):
N_val = {}
# This for loop can be paralleled.
for i in range(20):
# Pick one segment from data
segment_len=200000
data_array = traj[i*segment_len:(i+1)*segment_len]
N_val[i] = list(map(find_plateau, count_arr(data_array)))
N_arr = np.array(list(N_val.values()))
t_arr = segment_len/N_arr
# Mean transition time
mean_t = len(traj)*0.05/np.sum(N_arr, axis=0)
stdv_t = np.std(t_arr*0.05, axis=0)/np.sqrt(len(N_arr))
return mean_t, stdv_t
mean_t_arr = []
stdv_t_arr = []
for i in range(10):
traj = pred2[i]
mean_t_i, stdv_t_i = traj_i(traj)
mean_t_arr.append(mean_t_i)
stdv_t_arr.append(stdv_t_i)
print(i)
mean_t_arr = np.array(mean_t_arr)
mean_t_arr
mean_t = np.mean(mean_t_arr,axis=0)
mean_t
stdv_t_arr = np.array(stdv_t_arr)
stdv_t_arr
stdv_t = np.sqrt(np.mean(stdv_t_arr**2, axis=0))
stdv_t
```
| github_jupyter |
```
import pickle
from cbsa import ReactionSystem
import cbsa2models
import numpy as np
import matplotlib.pyplot as plt
def switch_time_gaps(data):
tf_str = "".join([str(int(data[i,1])) for i in range(data.shape[0])])
tf_steps = [step.split("10") for step in tf_str.split("01")]
tf_steps = [len(item)+2 for sublist in tf_steps for item in sublist]
tf_steps[0] -= 1
tf_steps[-1] -= 1
cumsum = [0]+list(np.cumsum(tf_steps))
time_steps = np.diff(data[:,0])
tf_times = {0:[],1:[]}
value = 0
for i in range(1,len(cumsum)):
time = np.sum(time_steps[cumsum[i-1]:cumsum[i]])
tf_times[value].append(time)
if value:
value = 0
else:
value = 1
return tf_times
def mean_value(data,index,time=None):
time_index = data.shape[0]
if time:
time_index = np.searchsorted(data[:,0],time,side="left")
time_steps = np.diff(data[:time_index,0])
sum_ts = np.sum(time_steps)
if not sum_ts: return 0.
return np.sum(time_steps*data[:time_index,index][0:-1])/sum_ts
total_sim_time = 100000
cbsa = cbsa2models.generate_cbsa_burst_model(total_sim_time=total_sim_time)
cbsa.setup_simulation(use_opencl=False)
cbsa.compute_simulation(total_sim_time,batch_steps=1)
cbsa_data = np.array(cbsa.simulation_data)
with open("cbsa_burst_0.5_0.5_200_0.5.pdata","wb") as f:
pickle.dump(cbsa_data,f)
cbsa_data = pickle.load( open( "cbsa_burst_0.5_0.5_200_0.5.pdata", "rb" ) )
cbsa_switch_times = switch_time_gaps(cbsa_data)
cbsa_times = np.linspace(0.,cbsa_data[-1,0],5000)
cbsa_mean_values_Eact = [mean_value(cbsa_data,1,i) for i in cbsa_times]
cbsa_mean_values_Mol = [mean_value(cbsa_data,3,i) for i in cbsa_times]
from matplotlib import rc
fontsize = 14
rc('text', usetex=True)
plt.style.use("bmh")
plt.rcParams["font.family"] = "serif"
plt.rcParams["xtick.labelsize"] = 12
plt.rcParams["ytick.labelsize"] = 12
prop_cycle = plt.rcParams['axes.prop_cycle']
colors = prop_cycle.by_key()['color']
fig = plt.figure(figsize=(10, 4),constrained_layout=True)
gs = fig.add_gridspec(nrows=10, ncols=2)
ax0 = fig.add_subplot(gs[0:6,0:1])
ax1 = fig.add_subplot(gs[6:10,0:1])
ax2 = fig.add_subplot(gs[0:4,1:2])
ax3 = fig.add_subplot(gs[4:7,1:2])
ax4 = fig.add_subplot(gs[7:10,1:2])
ax0.plot(cbsa_data[:,0],cbsa_data[:,3],color=colors[0])
ax0.set_xlim(-1,400)
ax0.set_xticklabels([])
ax0.set_ylabel("$P$",fontsize=fontsize)
ax0.text(-0.16,0.9,r'\textbf{a)}', transform=ax0.transAxes,fontsize=18)
ax1.plot(cbsa_data[:,0],cbsa_data[:,1],color=colors[4])
ax1.set_xlim(-1,400)
ax1.set_xlabel("Time (s)",fontsize=fontsize)
ax1.set_ylabel("$Tf_{act}$",fontsize=fontsize)
ax2.hist(cbsa_switch_times[0]+cbsa_switch_times[1],bins=35,color=colors[4])
ax2.set_xlabel("Switching Times (s)",fontsize=fontsize)
ax2.set_ylabel("Frequency",fontsize=fontsize)
ax2.text(85,500,"Mean: "+str(np.round(np.mean(cbsa_switch_times[0]+cbsa_switch_times[1]),2))+"\nStd: "+str(np.round(np.std(cbsa_switch_times[0]+cbsa_switch_times[1]),2)),fontsize=fontsize)
ax2.text(-0.22,0.8,r'\textbf{b)}', transform=ax2.transAxes,fontsize=18)
ax3.plot(cbsa_times,cbsa_mean_values_Mol,color=colors[0])
ax3.set_xticklabels([])
ax3.set_ylabel("$<P>$",fontsize=fontsize)
ax3.text(-0.22,0.8,r'\textbf{c)}', transform=ax3.transAxes,fontsize=18)
ax3.set_xlim(-300,6000)
#ax3.set_ylim(0,np.max(np.array(cbsa_mean_values_Mol))*1.1)
ax4.plot(cbsa_times,cbsa_mean_values_Eact,color=colors[4])
ax4.set_xlabel("Time (s)",fontsize=fontsize)
ax4.set_ylabel("$<Tf_{act}>$",fontsize=fontsize)
ax4.set_xlim(-300,6000)
#ax4.set_ylim(0,np.max(np.array(cbsa_mean_values_Eact))*1.1)
#ax4.set_yticks([0.,0.5])
plt.savefig("burst.png",dpi=300, bbox_inches='tight')
plt.show()
def run_stochpy(cbsa_model,method="Direct"):
smod = cbsa2models.cbsa2stochpy(cbsa_model)
smod.DoStochSim(method=method,mode='time',end=cbsa_model.bench_total_sim_time,trajectories=1)
return smod.data_stochsim.getSpecies()
def run_stochpy_ssa(cbsa_model):
return run_stochpy(cbsa_model)
def run_stochpy_tauleap(cbsa_model):
return run_stochpy(cbsa_model,method="Tauleap")
ssa_data = run_stochpy_ssa(cbsa)
with open("stochpy_ssa_burst_0.05_0.05_200_0.5.pdata","wb") as f:
pickle.dump(ssa_data,f)
tauleap_data = run_stochpy_tauleap(cbsa)
with open("stochpy_tauleap_burst_0.05_0.05_200_0.5.pdata","wb") as f:
pickle.dump(tauleap_data,f)
ssa_data = pickle.load( open( "stochpy_ssa_burst_0.05_0.05_200_0.5.pdata", "rb" ) )
ssa_switch_times = switch_time_gaps(ssa_data)
ssa_times = np.linspace(0.,ssa_data[-1,0],5000)
ssa_mean_values_Eact = [mean_value(ssa_data,1,i) for i in ssa_times]
ssa_mean_values_Mol = [mean_value(ssa_data,3,i) for i in ssa_times]
tauleap_data = pickle.load( open( "stochpy_tauleap_burst_0.05_0.05_200_0.5.pdata", "rb" ) )
tauleap_switch_times = switch_time_gaps(tauleap_data)
tauleap_times = np.linspace(0.,tauleap_data[-1,0],5000)
tauleap_mean_values_Eact = [mean_value(tauleap_data,1,i) for i in tauleap_times]
tauleap_mean_values_Mol = [mean_value(tauleap_data,3,i) for i in tauleap_times]
fig = plt.figure(figsize=(4, 5),constrained_layout=True)
gs = fig.add_gridspec(nrows=4, ncols=1)
ax0 = fig.add_subplot(gs[0:2])
ax1 = fig.add_subplot(gs[2])
ax2 = fig.add_subplot(gs[3])
bins = 35
ax0.hist(cbsa_switch_times[0]+cbsa_switch_times[1],bins=bins,alpha=0.9,histtype='step',label=r'CBSA $'+str(np.round(np.mean(cbsa_switch_times[0]+cbsa_switch_times[1]),2))+r'\pm$'+str(np.round(np.std(cbsa_switch_times[0]+cbsa_switch_times[1]),2)),linewidth=2)
ax0.hist(ssa_switch_times[0]+ssa_switch_times[1],bins=bins,alpha=0.9,histtype='step',label=r'SSA $'+str(np.round(np.mean(ssa_switch_times[0]+ssa_switch_times[1]),2))+r'\pm$'+str(np.round(np.std(ssa_switch_times[0]+ssa_switch_times[1]),2)),linewidth=2)
ax0.hist(tauleap_switch_times[0]+tauleap_switch_times[1],bins=bins,alpha=0.9,histtype='step',label=r'$\tau$-leap $'+str(np.round(np.mean(tauleap_switch_times[0]+tauleap_switch_times[1]),2))+r'\pm$'+str(np.round(np.std(tauleap_switch_times[0]+tauleap_switch_times[1]),2)),linewidth=2)
ax0.set_ylabel("Frequency",fontsize=fontsize)
ax0.set_xlabel("Switching Times (s)",fontsize=fontsize)
ax0.text(-0.2,0.9,r'\textbf{a)}', transform=ax0.transAxes,fontsize=17)
ax0.legend(loc="upper right")
ax1.plot(cbsa_times,cbsa_mean_values_Mol,alpha=0.9)
ax1.plot(ssa_times,ssa_mean_values_Mol,alpha=0.9)
ax1.plot(tauleap_times,tauleap_mean_values_Mol,alpha=0.9)
ax1.set_xticklabels([])
ax1.set_ylabel("$<P>$",fontsize=fontsize)
ax1.text(-0.2,1,r'\textbf{b)}', transform=ax1.transAxes,fontsize=17)
ax1.set_xlim(-400,10000)
ax2.plot(cbsa_times,cbsa_mean_values_Eact,alpha=0.9)
ax2.plot(ssa_times,ssa_mean_values_Eact,alpha=0.9)
ax2.plot(tauleap_times,tauleap_mean_values_Eact,alpha=0.9)
ax2.set_xlabel("Time (s)",fontsize=fontsize)
ax2.set_ylabel("$<Tf_{act}>$",fontsize=fontsize)
ax2.set_xlim(-400,10000)
plt.savefig("burst_ssa_validation.png",dpi=300, bbox_inches='tight')
plt.show()
```
| github_jupyter |
```
!ls
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import _init_paths
import os
import sys
import numpy as np
import argparse
import pprint
import pdb
import time
import cv2
import xml.dom.minidom as minidom
from roi_data_layer.roidb import combined_roidb
from roi_data_layer.roibatchLoader import roibatchLoader
from datasets.imagenet import imagenet
dataset = 'imagenet'
imdb_name = 'imagenet_train'
imdbval_name = 'imagenet_val'
split = 'train'#train val val1 val2 test
name = 'imagenet_{}'.format(split)
devkit_path = 'data/imagenet/ILSVRC/devkit'
data_path = 'data/imagenet/ILSVRC'
db_handle = (lambda split=split, devkit_path=devkit_path, data_path=data_path: imagenet(split,devkit_path,data_path))
print(type(db_handle))
import scipy.io as sio
synsets_image = sio.loadmat(os.path.join(devkit_path,'data','meta_det.mat'))
synsets_video = sio.loadmat(os.path.join(devkit_path,'data','meta_vid.mat'))
print(synsets_image.keys())
print(synsets_image)
syn_I = synsets_image['synsets']
syn_V = synsets_video['synsets']
print(type(syn_I))
print(syn_I.shape)
```
#### synsets are in order of: ID(int), WNID(string), Name(string), Description(string).
```
print(syn_I[0,0])
print(syn_V[0,0])
```
#### find the wnids in both VID and DET
```
vid_wnids = [syn_V[0,i][1] for i in range(30)]
print(vid_wnids)
used_wnid_name = []
for i in range(0,200):
wnid = syn_I[0,i][1]
name = syn_I[0,i][2]
if wnid in vid_wnids:
#print(wnid,name)
used_wnid_name.append((wnid,name))
print(len(used_wnid_name))
```
#### text file for different classes.
```
vtmp_index = [] # label as 1.
vtmp_index0 = [] # label as 0.
vtmp_index_1 = [] # label as -1.
for i in range(1,201):
image_set_file = os.path.join(data_path,'ImageSets','DET','train_'+str(i)+'.txt')
print(image_set_file,os.path.exists(image_set_file))
with open(image_set_file,'r') as f:
tmp_index = [x for x in f.readlines()]
vtmp = []
for line in tmp_index:
line = line.strip()
line = line.split(' ')
if line[1] == '1':
vtmp_index.append(data_path+'/Data/DET/train/'+line[0])
elif line[1] == '0':
vtmp_index0.append(data_path+'/Data/DET/train/'+line[0])
elif line[1] == '-1':
vtmp_index_1.append(data_path+'/Data/DET/train/'+line[0])
else:
raise Exception('Error.')
print(len(vtmp_index),len(vtmp_index0),len(vtmp_index_1),'sum:',len(vtmp_index)+len(vtmp_index0)+len(vtmp_index_1))
```
#### There are still some overlaps between input files.
```
len(np.unique(vtmp_index))
```
##### what about trian.txt val.txt and test.txt. It is < image_index + id_number >
```
print('Training set:')
image_set_file = os.path.join(data_path,'ImageSets','DET','train.txt')
print(image_set_file,os.path.exists(image_set_file))
with open(image_set_file,'r') as f:
tmp_index = [x for x in f.readlines()]
print(len(tmp_index))
print(tmp_index[0])
print('Val set:')
image_set_file = os.path.join(data_path,'ImageSets','DET','val.txt')
print(image_set_file,os.path.exists(image_set_file))
with open(image_set_file,'r') as f:
tmp_index = [x for x in f.readlines()]
print(len(tmp_index))
```
### Examine Video set
```
vtmp_index = [] # label as 1.
vtmp_index0 = [] # label as 0.
vtmp_index_1 = [] # label as -1.
for i in range(1,31):
image_set_file = os.path.join(data_path,'ImageSets','VID','train_'+str(i)+'.txt')
print(image_set_file,os.path.exists(image_set_file))
with open(image_set_file,'r') as f:
tmp_index = [x for x in f.readlines()]
vtmp = []
for line in tmp_index:
line = line.strip()
line = line.split(' ')
assert os.path.exists(data_path+'/Data/VID/train/'+line[0])
if line[1] == '1':
vtmp_index.append(data_path+'/Data/VID/train/'+line[0])
elif line[1] == '0':
vtmp_index0.append(data_path+'/Data/VID/train/'+line[0])
elif line[1] == '-1':
vtmp_index_1.append(data_path+'/Data/VID/train/'+line[0])
else:
raise Exception('Error.')
```
#### All the images in video set are fully annotated.
```
print(len(vtmp_index),len(vtmp_index0),len(vtmp_index_1),'sum:',len(vtmp_index)+len(vtmp_index0)+len(vtmp_index_1))
print(vtmp_index[0])
#def get_vid_train_data_statistics(homePath = 'data/imagenet/ILSVRC/'):
homePath = 'data/imagenet/ILSVRC/'
dataPath = os.path.join(homePath,'Data','VID','train')
setPath = os.path.join(homePath,'ImageSets','VID')
filenames_all = []
vid_num = 0
vid_counter = np.zeros(30)
for vid_idx in range(1,31):
image_set_file = os.path.join(homePath,'ImageSets','VID','train_'+str(vid_idx)+'.txt')
filenames = []
with open(image_set_file) as f:
items = [x.split(' ')[0] for x in f.readlines()]
for idx in range(len(items)):
vid_num += 1
vid_counter[vid_idx-1] += 1
item = items[idx]
item_dir = os.path.join(dataPath, item)
fns = [fn for fn in os.listdir(item_dir) if 'JPEG' in fn]
assert fns>0
filenames.extend(fns)
assert len(filenames)>0
print(syn_V[0,vid_idx-1][2],len(filenames))
filenames_all.extend(filenames)
print('total vid number:',vid_num)
print('total images:',len(filenames_all))
print('number of videos for each category:',vid_counter)
get_vid_train_data_statistics()
def cal_vid_number(vid_counter, frames_per_vid = 15, max_vid_per_category = 100):
return np.minimum(vid_counter, max_vid_per_category)*frames_per_vid
print(cal_vid_number(vid_counter))
print(np.sum(cal_vid_number(vid_counter)))
####VID object info####
_wnid = (0,)
_classes = ('__background__',)
for i in xrange(30):
_classes = _classes + (synsets_video['synsets'][0][i][2][0],)
_wnid = _wnid + (synsets_video['synsets'][0][i][1][0],)
_wnid_to_ind = dict(zip(_wnid, xrange(31)))
_class_to_ind = dict(zip(_classes, xrange(31)))
#######################
def read_annotation(dataPath):
filename = dataPath.replace('Data','Annotations').replace('JPEG','xml')
assert os.path.exists(filename),'%s'%(filename)
# print 'Loading: {}'.format(filename)
def get_data_from_tag(node, tag):
return node.getElementsByTagName(tag)[0].childNodes[0].data
with open(filename) as f:
data = minidom.parseString(f.read())
objs = data.getElementsByTagName('object')
num_objs = len(objs)
boxes = np.zeros((num_objs, 4), dtype=np.int32)
gt_classes = np.zeros(num_objs,dtype=np.int32)
# Load object bounding boxes into a data frame.
for ix, obj in enumerate(objs):
x1 = float(get_data_from_tag(obj, 'xmin'))
y1 = float(get_data_from_tag(obj, 'ymin'))
x2 = float(get_data_from_tag(obj, 'xmax'))
y2 = float(get_data_from_tag(obj, 'ymax'))
cls = _wnid_to_ind[
str(get_data_from_tag(obj, "name")).lower().strip()]
boxes[ix, :] = [x1, y1, x2, y2]
gt_classes[ix] = cls
return boxes, gt_classes
def watch_vid_train_data(vid_idx, idx, homePath = 'data/imagenet/ILSVRC/', show_class=True, save_vid=False, dest_vid_path = 'video.avi'):
dataPath = os.path.join(homePath,'Data','VID','train')
setPath = os.path.join(homePath,'ImageSets','VID')
assert vid_idx>0 and vid_idx<31
image_set_file = os.path.join(homePath,'ImageSets','VID','train_'+str(vid_idx)+'.txt')
with open(image_set_file) as f:
items = [x.split(' ')[0] for x in f.readlines()]
assert idx>=0 and idx<len(items), print(len(items))
item = items[idx]
item_dir = os.path.join(dataPath, item)
filenames = [fn for fn in os.listdir(item_dir) if 'JPEG' in fn]
assert len(filenames)>0
print(len(filenames))
height , width , _ = cv2.imread(os.path.join(item_dir,filenames[0])).shape
while height>800 or width > 800:
height = int(height/2)
width = int(width/2)
if save_vid:
fourcc = cv2.VideoWriter_fourcc(*'XVID')
video = cv2.VideoWriter(dest_vid_path, fourcc,20.0,(width,height))
for i in range(len(filenames)):
sys.stdout.write('%d/%d \r'%(i+1,len(filenames)))
sys.stdout.flush()
img = cv2.imread(os.path.join(item_dir,'%06d.JPEG'%(i)))
boxes, gt_classes = read_annotation(os.path.join(item_dir,'%06d.JPEG'%(i)))
for obj_id in range(len(gt_classes)):
bbox = boxes[obj_id,:]
class_name = _classes[gt_classes[obj_id]]
cv2.rectangle(img, (bbox[0],bbox[1]), (bbox[2],bbox[3]), (0, 204, 0), 2)
cv2.putText(img, '%s' % (class_name), (bbox[0], bbox[1] + 15), cv2.FONT_HERSHEY_PLAIN,
1.0, (0, 0, 255), thickness=1)
if img.shape[0]!=height:
img = cv2.resize(img,(width,height))
if save_vid:
video.write(img)
else:
cv2.imshow('image',img)
cv2.waitKey(50)
video.release()
cv2.destroyAllWindows()
watch_vid_train_data(1,35,save_vid=True,dest_vid_path='video.avi')
def getEgTrainDataVids():
save_dir = 'eg_train_vid'
if not os.path.isdir(save_dir):
os.mkdir(save_dir)
for i in range(1,31):
try:
for j in range(0,1000,10):
watch_vid_train_data(i,j,save_vid=True,dest_vid_path=os.path.join(save_dir,'%d-%d.avi'%(i,j)))
except:
print('j out of range.')
getEgTrainDataVids()
```
#### check val set.
```
vtmp_index = [] # label as 1.
vtmp_index0 = [] # label as 0.
vtmp_index_1 = [] # label as -1.
for i in range(1,31):
image_set_file = os.path.join(data_path,'ImageSets','VID','train_'+str(i)+'.txt')
print(image_set_file,os.path.exists(image_set_file))
with open(image_set_file,'r') as f:
tmp_index = [x for x in f.readlines()]
vtmp = []
for line in tmp_index:
line = line.strip()
line = line.split(' ')
if line[1] == '1':
vtmp_index.append(data_path+'/Data/DET/train/'+line[0])
elif line[1] == '0':
vtmp_index0.append(data_path+'/Data/DET/train/'+line[0])
elif line[1] == '-1':
vtmp_index_1.append(data_path+'/Data/DET/train/'+line[0])
else:
raise Exception('Error.')
```
## Examine the annotation files.
| github_jupyter |
https://github.com/ustroetz/python-osrm
OSMR API documentation: http://project-osrm.org/docs/v5.24.0/api/?language=cURL#
Выжимка:
**Выбирается самый быстрый путь, а не минимальное расстояние между двумя координатами!**
**Расстояние указывается в метрах**
**Open Source OSMR API поддерживает только пути для машин. Есть попытаться указать другие опции, результат тот же.**
**route** — посчитает путь со всеми указанными координатами.
options:
* skip_waypoints=true — не будет выводить информацию об указанных координатах. Позволяет экономить место;
* alternatives=n — показать альтернативные пути;
* geometries=geojson — возвращает список пар координат для построения пути;
* overview=full — полный, simplified — не подробный, уменьшает количество пар координат, false — без геометрии;
response:
* code — обозначает успешность запроса (200 успешно, 400 — ошибка);
* waypoints — точки на пути, исчезнут при skip_waypoints=true;
* routes — пути, отранжированные от самого быстрого к самому медленному.
**table** — посчитает пути между всеми парами координат, переданными в запрос.
options:
* skip_waypoints=true — не будет выводить информацию об указанных координатах. Позволяет экономить место, но также обнулит sources и destinations;
* sources — координаты, используемые в качестве отправной точки, вернутся в качестве waypoint объекта
* destinations — координаты, используемые в качестве пункта назначения, вернутся в качестве waypoint объекта
* annotations — duration (default), distance , or duration,distance — что возвращается;
* fallback_speed — в случае, если между точкам не удается найти путь, будет посчитано кратчайшее расстояние, для которого необходимо указать скорость.
response:
* code
* durations — null, если путь не удалось построить;
* distances — null, если путь не удалось построить;
* sources
* destinations
* fallback_speed_cells — если был использован fallback_speed, то вернется массив, указывающий, какие пары были рассчитаны подобным образом.
Макет ссылки:
http://router.project-osrm.org/route/v1/driving/59.98259700478887,30.4297523923562;55.75269662241035,37.64085841204411?overview=false
```
import requests
import json
import folium
response = requests.get('http://router.project-osrm.org/route/v1/driving/59.98259700478887,30.4297523923562;55.75269662241035,37.64085841204411?overview=simplified&geometries=geojson')
route = json.loads(response.text)
route['routes'][0]['geometry']['coordinates']
route['routes'][0]['distance']
r = route['routes'][0]['geometry']['coordinates']
m = folium.Map(location=[59.98259700478887,30.4297523923562],
zoom_start=5)
folium.PolyLine(r,
color='red',
weight=15,
opacity=0.8).add_to(m)
m
```
# Пробуем на датасете
```
import psycopg2
import numpy as np
from scipy.cluster.hierarchy import fclusterdata
from scipy.spatial.distance import pdist
from math import pi, sin, cos, atan2
import matplotlib.pyplot as plt
import pandas as pd
from itertools import combinations
from tqdm import tqdm
conn = psycopg2.connect(dbname='sociolinguistic', user='app',
password='rfrfyl.babaloos', host='gisly.net')
cursor = conn.cursor()
cursor.execute(""" select place_born.longitude, place_born.latitude
from sociolinguistic_place place_born
where place_born.longitude is not null
and place_born.latitude is not null""")
X = np.array(cursor.fetchall())
cursor.close()
conn.close()
X[0]
table = list(combinations(X, 2))
longitude_x = [x[0][0] for x in table]
latitude_x = [x[0][1] for x in table]
longitude_y = [x[1][0] for x in table]
latitude_y = [x[1][1] for x in table]
df = pd.DataFrame()
df['longitude_x'] = longitude_x
df['latitude_x'] = latitude_x
df['longitude_y'] = longitude_y
df['latitude_y'] = latitude_y
df['distance'] = np.nan
df['duration'] = np.nan
df['geometry'] = np.nan
def calculate_dist(place1, place2):
place1 = str(place1[0]) + ',' + str(place1[1])
place2 = str(place2[0]) + ',' + str(place2[1])
try:
response = requests.get(
'http://router.project-osrm.org/route/v1/driving/{0};{1}?overview=simplified&geometries=geojson'.format(
place1, place2))
response = json.loads(response.text)
distance = response['routes'][0]['distance']
duration = response['routes'][0]['duration']
geometry = json.dumps(response['routes'][0]['geometry']['coordinates'])
except Exception as e:
distance = np.nan
duration = np.nan
geometry = np.nan
return distance, duration, geometry
for num, row in tqdm(df.iterrows()):
distance, duration, geometry = calculate_dist([row['longitude_x'], row['latitude_x']],
[row['longitude_y'], row['latitude_y']])
df.loc[num, 'distance'] = distance
df.loc[num, 'duration'] = duration
df.loc[num, 'geometry'] = geometry
```
| github_jupyter |
# User characterization
In this notebook, we we analyze the traces from the users’ perspective
```
"""
Note that only the `Venus` trace is public available now.
Other traces are being censored. We will release them as soon as possible.
"""
# cluster_list = ["Venus", "Earth", "Saturn", "Uranus"]
cluster_list = ["Venus"]
from datetime import timedelta
import matplotlib
import matplotlib.patches as mpatches
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
from matplotlib import rc
from pylab import *
matplotlib.font_manager._rebuild()
sns.set_style("ticks")
font = {
"font.family": "Roboto",
"font.size": 12,
}
sns.set_style(font)
paper_rc = {
"lines.linewidth": 3,
"lines.markersize": 10,
}
sns.set_context("paper", font_scale=1.6, rc=paper_rc)
current_palette = sns.color_palette()
def load_data(cluster, opt):
"""Data Loading & Processing"""
if opt == "user":
df = pd.read_pickle(f"../../data/{cluster}/cluster_user.pkl")
elif opt == "log":
df = pd.read_csv(
f"../../data/{cluster}/cluster_log.csv",
parse_dates=["submit_time", "start_time", "end_time"],
)
else:
raise ValueError("Please check opt")
return df
```
## User Runtime
The CDFs of users that consume the cluster resources
(a) GPU Time
(b) CPU Time
```
def plot_cdf_time_user(cluster_list, save=False):
fig, (ax1, ax2) = plt.subplots(
ncols=2, nrows=1, constrained_layout=True, figsize=(8, 3)
)
linestyles = ["--", "-.", ":", "--"]
for k in range(0, len(cluster_list)):
g_time, c_time = [], []
udf = load_data(cluster_list[k], "user")
gpu_time = udf["total_gpu_time"].copy()
gpu_time.sort_values(ascending=False, inplace=True)
gpu_time_sum = gpu_time.sum()
cpu_time = udf["total_cpu_only_time"].copy()
cpu_time.sort_values(ascending=False, inplace=True)
cpu_time_sum = cpu_time.sum()
for i in range(len(udf)):
g_time.append(gpu_time.iloc[: i + 1].sum() / gpu_time_sum * 100)
c_time.append(cpu_time.iloc[: i + 1].sum() / cpu_time_sum * 100)
ax1.plot(
np.linspace(0, 100, len(udf)),
g_time,
linestyles[k],
alpha=0.9,
label=cluster_list[k],
)
ax2.plot(
np.linspace(0, 100, len(udf)),
c_time,
linestyles[k],
alpha=0.9,
label=cluster_list[k],
)
ax1.set_xlabel(f"Fraction of Users (%)")
ax1.set_ylabel(f"Fraction of Total\nGPU Time (%)")
ax1.set_xlim(0, 100)
ax1.set_ylim(0, 101)
ax1.legend(loc="lower right")
ax1.grid(linestyle=":")
ax1.text(0.45, -0.4, "(a)", transform=ax1.transAxes, size=14)
ax2.set_xlabel(f"Fraction of Users (%)")
ax2.set_ylabel(f"Fraction of Total\nCPU Time (%)")
ax2.set_xlim(0, 100)
ax2.set_ylim(0, 101)
ax2.legend(loc="lower right")
ax2.grid(linestyle=":")
ax2.text(0.45, -0.4, "(b)", transform=ax2.transAxes, size=14)
if save:
fig.savefig(
f"./user_cdf_time.pdf", bbox_inches="tight", dpi=600,
)
plot_cdf_time_user(cluster_list, save=True)
```
## User Queuing Time
(a) CDFs of users w.r.t. GPU job queuing delay.
(b) Distributions of user GPU job completion ratios.
```
def plot_cdf_pend_user(cluster_list, save=False):
fig, (ax1, ax2) = plt.subplots(
ncols=2, nrows=1, constrained_layout=True, figsize=(8, 3)
)
linestyles = ["--", "-.", ":", "--"]
for k in range(0, len(cluster_list)):
g_num = []
udf = load_data(cluster_list[k], "user")
gpu_time = udf["total_gpu_pend_time"].copy()
gpu_time.sort_values(ascending=False, inplace=True)
gpu_time_sum = gpu_time.sum()
for i in range(len(udf)):
g_num.append(gpu_time.iloc[: i + 1].sum() / gpu_time_sum * 100)
ax1.plot(
np.linspace(0, 100, len(udf)),
g_num,
linestyles[k],
alpha=0.9,
label=cluster_list[k],
)
udf = pd.read_pickle(f"../../data/cluster_user.pkl")
state = udf["completed_gpu_percent"].copy() * 100
ax2 = sns.histplot(
state, bins=10, stat="count", kde=False, alpha=0.9, label="Completed"
)
ax1.set_xlabel(f"Fraction of Users (%)")
ax1.set_ylabel(f"Fraction of Total\nQueuing Time (%)")
ax1.set_xlim(0, 100)
ax1.set_ylim(0, 101)
ax1.legend(loc="lower right")
ax1.grid(linestyle=":")
ax1.text(0.45, -0.4, "(a)", transform=ax1.transAxes, size=14)
ax2.set_xlabel(f"GPU Job Completion Rate (%)")
ax2.set_ylabel(f"User Number")
ax2.set_xlim(0, 100)
ax2.grid(linestyle=":")
ax2.text(0.45, -0.4, "(b)", transform=ax2.transAxes, size=14)
if save:
fig.savefig(
f"./user_cdf_pend.pdf", bbox_inches="tight", dpi=600,
)
plot_cdf_pend_user(cluster_list, save=True)
```
| github_jupyter |
<img src="../../../images/qiskit-heading.gif" alt="Note: In order for images to show up in this jupyter notebook you need to select File => Trusted Notebook" width="500 px" align="left">
## _*The Vaidman Detection Test: Interaction Free Measurement*_
The latest version of this notebook is available on https://github.com/Qiskit/qiskit-tutorial.
***
### Contributors
Alex Breitweiser
## Introduction
One surprising result of quantum mechanics is the ability to measure something without ever directly "observing" it. This interaction-free measurement cannot be reproduced in classical mechanics. The prototypical example is the [Elitzur–Vaidman Bomb Experiment](https://en.wikipedia.org/wiki/Elitzur%E2%80%93Vaidman_bomb_tester) - in which one wants to test whether bombs are active without detonating them. In this example we will test whether an unknown operation is null (the identity) or an X gate, corresponding to a dud or a live bomb.
### The Algorithm
The algorithm will use two qubits, $q_1$ and $q_2$, as well as a small parameter, $\epsilon = \frac{\pi}{n}$ for some integer $n$. Call the unknown gate, which is either the identity or an X gate, $G$, and assume we have it in a controlled form. The algorithm is then:
1. Start with both $q_1$ and $q_2$ in the $|0\rangle$ state
2. Rotate $q_1$ by $\epsilon$ about the Y axis
3. Apply a controlled $G$ on $q_2$, conditioned on $q_1$
4. Measure $q_2$
5. Repeat (2-4) $n$ times
6. Measure $q_1$

### Explanation and proof of correctness
There are two cases: Either the gate is the identity (a dud), or it is an X gate (a live bomb).
#### Case 1: Dud
After rotation, $q_1$ is now approximately
$$q_1 \approx |0\rangle + \frac{\epsilon}{2} |1\rangle$$
Since the unknown gate is the identity, the controlled gate leaves the two qubit state separable,
$$q_1 \times q_2 \approx (|0\rangle + \frac{\epsilon}{2} |1\rangle) \times |0\rangle$$
and measurement is trivial (we will always measure $|0\rangle$ for $q_2$).
Repetition will not change this result - we will always keep separability and $q_2$ will remain in $|0\rangle$.
After n steps, $q_1$ will flip by $\pi$ to $|1\rangle$, and so measuring it will certainly yield $1$. Therefore, the output register for a dud bomb will read:
$$000...01$$
#### Case 2: Live
Again, after rotation, $q_1$ is now approximately
$$q_1 \approx |0\rangle + \frac{\epsilon}{2} |1\rangle$$
But, since the unknown gate is now an X gate, the combined state after $G$ is now
$$q_1 \times q_2 \approx |00\rangle + \frac{\epsilon}{2} |11\rangle$$
Measuring $q_2$ now might yield $1$, in which case we have "measured" the live bomb (obtained a result which differs from that of a dud) and it explodes. However, this only happens with a probability proportional to $\epsilon^2$. In the vast majority of cases, we will measure $0$ and the entire system will collapse back to
$$q_1 \times q_2 = |00\rangle$$
After every step, the system will most likely return to the original state, and the final measurement of $q_1$ will yield $0$. Therefore, the most likely outcome of a live bomb is
$$000...00$$
which will identify a live bomb without ever "measuring" it. If we ever obtain a 1 in the bits preceding the final bit, we will have detonated the bomb, but this will only happen with probability of order
$$P \propto n \epsilon^2 \propto \epsilon$$
This probability may be made arbitrarily small at the cost of an arbitrarily long circuit.
## Generating Random Bombs
A test set must be generated to experiment on - this can be done by classical (pseudo)random number generation, but as long as we have access to a quantum computer we might as well take advantage of the ability to generate true randomness.
```
# useful additional packages
import matplotlib.pyplot as plt
%matplotlib inline
import numpy as np
from collections import Counter #Use this to convert results from list to dict for histogram
# importing QISKit
from qiskit import QuantumCircuit, QuantumRegister, ClassicalRegister
from qiskit import execute, Aer, IBMQ
from qiskit.backends.ibmq import least_busy
# import basic plot tools
from qiskit.tools.visualization import plot_histogram
# To use IBMQ Quantum Experience
IBMQ.load_accounts()
```
We will generate a test set of 50 "bombs", and each "bomb" will be run through a 20-step measurement circuit. We set up the program as explained in previous examples.
```
# Use local qasm simulator
backend = Aer.get_backend('qasm_simulator')
# Use the IBMQ Quantum Experience
# backend = least_busy(IBMQ.backends())
N = 50 # Number of bombs
steps = 20 # Number of steps for the algorithm, limited by maximum circuit depth
eps = np.pi / steps # Algorithm parameter, small
# Prototype circuit for bomb generation
q_gen = QuantumRegister(1, name='q_gen')
c_gen = ClassicalRegister(1, name='c_gen')
IFM_gen = QuantumCircuit(q_gen, c_gen, name='IFM_gen')
# Prototype circuit for bomb measurement
q = QuantumRegister(2, name='q')
c = ClassicalRegister(steps+1, name='c')
IFM_meas = QuantumCircuit(q, c, name='IFM_meas')
```
Generating a random bomb is achieved by simply applying a Hadamard gate to a $q_1$, which starts in $|0\rangle$, and then measuring. This randomly gives a $0$ or $1$, each with equal probability. We run one such circuit for each bomb, since circuits are currently limited to a single measurement.
```
# Quantum circuits to generate bombs
qc = []
circuits = ["IFM_gen"+str(i) for i in range(N)]
# NB: Can't have more than one measurement per circuit
for circuit in circuits:
IFM = QuantumCircuit(q_gen, c_gen, name=circuit)
IFM.h(q_gen[0]) #Turn the qubit into |0> + |1>
IFM.measure(q_gen[0], c_gen[0])
qc.append(IFM)
_ = [i.qasm() for i in qc] # Suppress the output
```
Note that, since we want to measure several discrete instances, we do *not* want to average over multiple shots. Averaging would yield partial bombs, but we assume bombs are discretely either live or dead.
```
result = execute(qc, backend=backend, shots=1).result() # Note that we only want one shot
bombs = []
for circuit in qc:
for key in result.get_counts(circuit): # Hack, there should only be one key, since there was only one shot
bombs.append(int(key))
#print(', '.join(('Live' if bomb else 'Dud' for bomb in bombs))) # Uncomment to print out "truth" of bombs
plot_histogram(Counter(('Live' if bomb else 'Dud' for bomb in bombs))) #Plotting bomb generation results
```
## Testing the Bombs
Here we implement the algorithm described above to measure the bombs. As with the generation of the bombs, it is currently impossible to take several measurements in a single circuit - therefore, it must be run on the simulator.
```
# Use local qasm simulator
backend = Aer.get_backend('qasm_simulator')
qc = []
circuits = ["IFM_meas"+str(i) for i in range(N)]
#Creating one measurement circuit for each bomb
for i in range(N):
bomb = bombs[i]
IFM = QuantumCircuit(q, c, name=circuits[i])
for step in range(steps):
IFM.ry(eps, q[0]) #First we rotate the control qubit by epsilon
if bomb: #If the bomb is live, the gate is a controlled X gate
IFM.cx(q[0],q[1])
#If the bomb is a dud, the gate is a controlled identity gate, which does nothing
IFM.measure(q[1], c[step]) #Now we measure to collapse the combined state
IFM.measure(q[0], c[steps])
qc.append(IFM)
_ = [i.qasm() for i in qc] # Suppress the output
result = execute(qc, backend=backend, shots=1, max_credits=5).result()
def get_status(counts):
# Return whether a bomb was a dud, was live but detonated, or was live and undetonated
# Note that registers are returned in reversed order
for key in counts:
if '1' in key[1:]:
#If we ever measure a '1' from the measurement qubit (q1), the bomb was measured and will detonate
return '!!BOOM!!'
elif key[0] == '1':
#If the control qubit (q0) was rotated to '1', the state never entangled because the bomb was a dud
return 'Dud'
else:
#If we only measured '0' for both the control and measurement qubit, the bomb was live but never set off
return 'Live'
results = {'Live': 0, 'Dud': 0, "!!BOOM!!": 0}
for circuit in qc:
status = get_status(result.get_counts(circuit))
results[status] += 1
plot_histogram(results)
```
| github_jupyter |
# GPU
```
gpu_info = !nvidia-smi
gpu_info = '\n'.join(gpu_info)
print(gpu_info)
```
# CFG
```
CONFIG_NAME = 'config37.yml'
debug = False
from google.colab import drive, auth
# ドライブのマウント
drive.mount('/content/drive')
# Google Cloudの権限設定
auth.authenticate_user()
def get_github_secret():
import json
with open('/content/drive/MyDrive/config/github.json') as f:
github_config = json.load(f)
return github_config
github_config = get_github_secret()
! rm -r kaggle-cassava
user_name = github_config["user_name"]
password = github_config["password"]
! git clone https://{user_name}:{password}@github.com/raijin0704/kaggle-cassava.git
import sys
sys.path.append('./kaggle-cassava')
from src.utils.envs.main import create_env
env_dict = create_env()
env_dict
# ====================================================
# CFG
# ====================================================
import yaml
CONFIG_PATH = f'./kaggle-cassava/config/{CONFIG_NAME}'
with open(CONFIG_PATH) as f:
config = yaml.load(f)
INFO = config['info']
TAG = config['tag']
CFG = config['cfg']
DATA_PATH = env_dict["data_path"]
env = env_dict["env"]
NOTEBOOK_PATH = env_dict["notebook_dir"]
OUTPUT_DIR = env_dict["output_dir"]
TITLE = env_dict["title"]
CFG['train'] = True
CFG['inference'] = False
CFG['debug'] = debug
if CFG['debug']:
CFG['epochs'] = 1
# 環境変数
import os
os.environ["GCLOUD_PROJECT"] = INFO['PROJECT_ID']
# 間違ったバージョンを実行しないかチェック
assert INFO['TITLE'] == TITLE, f'{TITLE}, {INFO["TITLE"]}'
TITLE = INFO["TITLE"]
import os
if env=='colab':
!rm -r /content/input
! cp /content/drive/Shareddrives/便利用/kaggle/cassava/input.zip /content/input.zip
! unzip input.zip > /dev/null
! rm input.zip
train_num = len(os.listdir(DATA_PATH+"/train_images"))
assert train_num == 21397
```
# install apex
```
if CFG['apex']:
try:
import apex
except Exception:
! git clone https://github.com/NVIDIA/apex.git
% cd apex
!pip install --no-cache-dir --global-option="--cpp_ext" --global-option="--cuda_ext" .
%cd ..
```
# Library
```
# ====================================================
# Library
# ====================================================
import os
import datetime
import math
import time
import random
import glob
import shutil
from pathlib import Path
from contextlib import contextmanager
from collections import defaultdict, Counter
import scipy as sp
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
import seaborn as sns
from sklearn import preprocessing
from sklearn.metrics import accuracy_score
from sklearn.model_selection import StratifiedKFold
from tqdm.auto import tqdm
from functools import partial
import cv2
from PIL import Image
import torch
import torch.nn as nn
import torch.nn.functional as F
from torch.optim import Adam, SGD
import torchvision.models as models
from torch.nn.parameter import Parameter
from torch.utils.data import DataLoader, Dataset
from torch.optim.lr_scheduler import CosineAnnealingWarmRestarts, CosineAnnealingLR, ReduceLROnPlateau
from albumentations import (
Compose, OneOf, Normalize, Resize, RandomResizedCrop, RandomCrop, HorizontalFlip, VerticalFlip,
RandomBrightness, RandomContrast, RandomBrightnessContrast, Rotate, ShiftScaleRotate, Cutout,
IAAAdditiveGaussianNoise, Transpose, CenterCrop
)
from albumentations.pytorch import ToTensorV2
from albumentations import ImageOnlyTransform
import timm
import mlflow
import warnings
warnings.filterwarnings('ignore')
if CFG['apex']:
from apex import amp
if CFG['debug']:
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
else:
device = torch.device('cuda')
from src.utils.logger import init_logger
from src.utils.utils import seed_torch, EarlyStopping
from src.utils.loss.bi_tempered_logistic_loss import bi_tempered_logistic_loss
from src.utils.augments.randaugment import RandAugment
from src.utils.augments.augmix import RandomAugMix
start_time = datetime.datetime.now()
start_time_str = start_time.strftime('%m%d%H%M')
```
# Directory settings
```
# ====================================================
# Directory settings
# ====================================================
if os.path.exists(OUTPUT_DIR):
shutil.rmtree(OUTPUT_DIR)
if not os.path.exists(OUTPUT_DIR):
os.makedirs(OUTPUT_DIR)
```
# save basic files
```
# with open(f'{OUTPUT_DIR}/{start_time_str}_TAG.json', 'w') as f:
# json.dump(TAG, f, indent=4)
# with open(f'{OUTPUT_DIR}/{start_time_str}_CFG.json', 'w') as f:
# json.dump(CFG, f, indent=4)
import shutil
notebook_path = f'{OUTPUT_DIR}/{start_time_str}_{TITLE}.ipynb'
shutil.copy2(NOTEBOOK_PATH, notebook_path)
```
# Data Loading
```
train = pd.read_csv(f'{DATA_PATH}/train.csv')
test = pd.read_csv(f'{DATA_PATH}/sample_submission.csv')
label_map = pd.read_json(f'{DATA_PATH}/label_num_to_disease_map.json',
orient='index')
if CFG['debug']:
train = train.sample(n=1000, random_state=CFG['seed']).reset_index(drop=True)
```
# Utils
```
# ====================================================
# Utils
# ====================================================
def get_score(y_true, y_pred):
return accuracy_score(y_true, y_pred)
logger_path = OUTPUT_DIR+f'{start_time_str}_train.log'
LOGGER = init_logger(logger_path)
seed_torch(seed=CFG['seed'])
def remove_glob(pathname, recursive=True):
for p in glob.glob(pathname, recursive=recursive):
if os.path.isfile(p):
os.remove(p)
def rand_bbox(size, lam):
W = size[2]
H = size[3]
cut_rat = np.sqrt(1. - lam)
cut_w = np.int(W * cut_rat)
cut_h = np.int(H * cut_rat)
# uniform
cx = np.random.randint(W)
cy = np.random.randint(H)
bbx1 = np.clip(cx - cut_w // 2, 0, W)
bby1 = np.clip(cy - cut_h // 2, 0, H)
bbx2 = np.clip(cx + cut_w // 2, 0, W)
bby2 = np.clip(cy + cut_h // 2, 0, H)
return bbx1, bby1, bbx2, bby2
```
# CV split
```
folds = train.copy()
Fold = StratifiedKFold(n_splits=CFG['n_fold'], shuffle=True, random_state=CFG['seed'])
for n, (train_index, val_index) in enumerate(Fold.split(folds, folds[CFG['target_col']])):
folds.loc[val_index, 'fold'] = int(n)
folds['fold'] = folds['fold'].astype(int)
print(folds.groupby(['fold', CFG['target_col']]).size())
```
# Dataset
```
# ====================================================
# Dataset
# ====================================================
class TrainDataset(Dataset):
def __init__(self, df, transform=None):
self.df = df
self.file_names = df['image_id'].values
self.labels = df['label'].values
self.transform = transform
def __len__(self):
return len(self.df)
def __getitem__(self, idx):
file_name = self.file_names[idx]
file_path = f'{DATA_PATH}/train_images/{file_name}'
image = cv2.imread(file_path)
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
if self.transform:
augmented = self.transform(image=image)
image = augmented['image']
label = torch.tensor(self.labels[idx]).long()
return image, label
class TestDataset(Dataset):
def __init__(self, df, transform=None):
self.df = df
self.file_names = df['image_id'].values
self.transform = transform
def __len__(self):
return len(self.df)
def __getitem__(self, idx):
file_name = self.file_names[idx]
file_path = f'{DATA_PATH}/test_images/{file_name}'
image = cv2.imread(file_path)
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
if self.transform:
augmented = self.transform(image=image)
image = augmented['image']
return image
# train_dataset = TrainDataset(train, transform=None)
# for i in range(1):
# image, label = train_dataset[i]
# plt.imshow(image)
# plt.title(f'label: {label}')
# plt.show()
```
# Transforms
```
def _get_train_augmentations(aug_list):
process = []
for aug in aug_list:
if aug == 'Resize':
process.append(Resize(CFG['size'], CFG['size']))
elif aug == 'RandomResizedCrop':
process.append(RandomResizedCrop(CFG['size'], CFG['size']))
elif aug =='CenterCrop':
process.append(CenterCrop(CFG['size'], CFG['size']))
elif aug == 'Transpose':
process.append(Transpose(p=0.5))
elif aug == 'HorizontalFlip':
process.append(HorizontalFlip(p=0.5))
elif aug == 'VerticalFlip':
process.append(VerticalFlip(p=0.5))
elif aug == 'ShiftScaleRotate':
process.append(ShiftScaleRotate(p=0.5))
elif aug == 'RandomBrightness':
process.append(RandomBrightness(limit=(-0.2,0.2), p=1))
elif aug == 'Cutout':
process.append(Cutout(max_h_size=CFG['CutoutSize'], max_w_size=CFG['CutoutSize'], p=0.5))
elif aug == 'RandAugment':
process.append(RandAugment(CFG['RandAugmentN'], CFG['RandAugmentM'], p=0.5))
elif aug == 'RandomAugMix':
process.append(RandomAugMix(severity=CFG['AugMixSeverity'],
width=CFG['AugMixWidth'],
alpha=CFG['AugMixAlpha'], p=0.5))
elif aug == 'Normalize':
process.append(Normalize(
mean=[0.485, 0.456, 0.406],
std=[0.229, 0.224, 0.225],
))
elif aug in ['mixup', 'cutmix', 'fmix']:
pass
else:
raise ValueError(f'{aug} is not suitable')
process.append(ToTensorV2())
return process
def _get_valid_augmentations(aug_list):
process = []
for aug in aug_list:
if aug == 'Resize':
process.append(Resize(CFG['size'], CFG['size']))
elif aug == 'RandomResizedCrop':
process.append(OneOf(
[RandomResizedCrop(CFG['size'], CFG['size'], p=0.5),
Resize(CFG['size'], CFG['size'], p=0.5)], p=1))
elif aug =='CenterCrop':
process.append(OneOf(
[CenterCrop(CFG['size'], CFG['size'], p=0.5),
Resize(CFG['size'], CFG['size'], p=0.5)], p=1))
# process.append(
# CenterCrop(CFG['size'], CFG['size'], p=1.))
elif aug == 'Transpose':
process.append(Transpose(p=0.5))
elif aug == 'HorizontalFlip':
process.append(HorizontalFlip(p=0.5))
elif aug == 'VerticalFlip':
process.append(VerticalFlip(p=0.5))
elif aug == 'ShiftScaleRotate':
process.append(ShiftScaleRotate(p=0.5))
elif aug == 'RandomBrightness':
process.append(RandomBrightness(limit=(-0.2,0.2), p=1))
elif aug == 'Cutout':
process.append(Cutout(max_h_size=CFG['CutoutSize'], max_w_size=CFG['CutoutSize'], p=0.5))
elif aug == 'RandAugment':
process.append(RandAugment(CFG['RandAugmentN'], CFG['RandAugmentM'], p=0.5))
elif aug == 'RandomAugMix':
process.append(RandomAugMix(severity=CFG['AugMixSeverity'],
width=CFG['AugMixWidth'],
alpha=CFG['AugMixAlpha'], p=0.5))
elif aug == 'Normalize':
process.append(Normalize(
mean=[0.485, 0.456, 0.406],
std=[0.229, 0.224, 0.225],
))
elif aug in ['mixup', 'cutmix', 'fmix']:
pass
else:
raise ValueError(f'{aug} is not suitable')
process.append(ToTensorV2())
return process
# ====================================================
# Transforms
# ====================================================
def get_transforms(*, data):
if data == 'train':
return Compose(
_get_train_augmentations(TAG['augmentation'])
)
elif data == 'valid':
try:
augmentations = TAG['valid_augmentation']
except KeyError:
augmentations = ['Resize', 'Normalize']
return Compose(
_get_valid_augmentations(augmentations)
)
num_fig = 5
train_dataset = TrainDataset(train, transform=get_transforms(data='train'))
valid_dataset = TrainDataset(train, transform=get_transforms(data='valid'))
origin_dataset = TrainDataset(train, transform=None)
fig, ax = plt.subplots(num_fig, 3, figsize=(10, num_fig*3))
for j, dataset in enumerate([train_dataset, valid_dataset, origin_dataset]):
for i in range(num_fig):
image, label = dataset[i]
if j < 2:
ax[i,j].imshow(image.transpose(0,2).transpose(0,1))
else:
ax[i,j].imshow(image)
ax[i,j].set_title(f'label: {label}')
```
# MODEL
```
# ====================================================
# MODEL
# ====================================================
class CustomModel(nn.Module):
def __init__(self, model_name, pretrained=False):
super().__init__()
self.model = timm.create_model(model_name, pretrained=pretrained)
if hasattr(self.model, 'classifier'):
n_features = self.model.classifier.in_features
self.model.classifier = nn.Linear(n_features, CFG['target_size'])
elif hasattr(self.model, 'fc'):
n_features = self.model.fc.in_features
self.model.fc = nn.Linear(n_features, CFG['target_size'])
elif hasattr(self.model, 'head'):
n_features = self.model.head.in_features
self.model.head = nn.Linear(n_features, CFG['target_size'])
def forward(self, x):
x = self.model(x)
return x
model = CustomModel(model_name=TAG['model_name'], pretrained=False)
train_dataset = TrainDataset(train, transform=get_transforms(data='train'))
train_loader = DataLoader(train_dataset, batch_size=4, shuffle=True,
num_workers=4, pin_memory=True, drop_last=True)
for image, label in train_loader:
output = model(image)
print(output)
break
```
# Helper functions
```
# ====================================================
# Helper functions
# ====================================================
class AverageMeter(object):
"""Computes and stores the average and current value"""
def __init__(self):
self.reset()
def reset(self):
self.val = 0
self.avg = 0
self.sum = 0
self.count = 0
def update(self, val, n=1):
self.val = val
self.sum += val * n
self.count += n
self.avg = self.sum / self.count
def asMinutes(s):
m = math.floor(s / 60)
s -= m * 60
return '%dm %ds' % (m, s)
def timeSince(since, percent):
now = time.time()
s = now - since
es = s / (percent)
rs = es - s
return '%s (remain %s)' % (asMinutes(s), asMinutes(rs))
# ====================================================
# loss
# ====================================================
def get_loss(criterion, y_preds, labels):
if TAG['criterion']=='CrossEntropyLoss':
loss = criterion(y_preds, labels)
elif TAG['criterion'] == 'bi_tempered_logistic_loss':
loss = criterion(y_preds, labels, t1=CFG['bi_tempered_loss_t1'], t2=CFG['bi_tempered_loss_t2'])
return loss
# ====================================================
# Helper functions
# ====================================================
def train_fn(train_loader, model, criterion, optimizer, epoch, scheduler, device):
batch_time = AverageMeter()
data_time = AverageMeter()
losses = AverageMeter()
scores = AverageMeter()
# switch to train mode
model.train()
start = end = time.time()
global_step = 0
for step, (images, labels) in enumerate(train_loader):
# measure data loading time
data_time.update(time.time() - end)
images = images.to(device)
labels = labels.to(device)
batch_size = labels.size(0)
r = np.random.rand(1)
is_aug = r < 0.5 # probability of augmentation
if is_aug & ('cutmix' in TAG['augmentation']) & (epoch+1>=CFG['heavy_aug_start_epoch']):
# generate mixed sample
# inference from https://github.com/clovaai/CutMix-PyTorch/blob/master/train.py
lam = np.random.beta(CFG['CutmixAlpha'], CFG['CutmixAlpha'])
rand_index = torch.randperm(images.size()[0]).to(device)
labels_a = labels
labels_b = labels[rand_index]
bbx1, bby1, bbx2, bby2 = rand_bbox(images.size(), lam)
images[:, :, bbx1:bbx2, bby1:bby2] = images[rand_index, :, bbx1:bbx2, bby1:bby2]
# adjust lambda to exactly match pixel ratio
lam = 1 - ((bbx2 - bbx1) * (bby2 - bby1) / (images.size()[-1] * images.size()[-2]))
# compute output
y_preds = model(images)
loss = get_loss(criterion, y_preds, labels_a) * lam + \
get_loss(criterion, y_preds, labels_b) * (1. - lam)
else:
y_preds = model(images)
loss = get_loss(criterion, y_preds, labels)
# record loss
losses.update(loss.item(), batch_size)
if CFG['gradient_accumulation_steps'] > 1:
loss = loss / CFG['gradient_accumulation_steps']
if CFG['apex']:
with amp.scale_loss(loss, optimizer) as scaled_loss:
scaled_loss.backward()
else:
loss.backward()
# clear memory
del loss, y_preds
torch.cuda.empty_cache()
grad_norm = torch.nn.utils.clip_grad_norm_(model.parameters(), CFG['max_grad_norm'])
if (step + 1) % CFG['gradient_accumulation_steps'] == 0:
optimizer.step()
optimizer.zero_grad()
global_step += 1
# measure elapsed time
batch_time.update(time.time() - end)
end = time.time()
if step % CFG['print_freq'] == 0 or step == (len(train_loader)-1):
print('Epoch: [{0}][{1}/{2}] '
'Data {data_time.val:.3f} ({data_time.avg:.3f}) '
'Elapsed {remain:s} '
'Loss: {loss.val:.4f}({loss.avg:.4f}) '
'Grad: {grad_norm:.4f} '
#'LR: {lr:.6f} '
.format(
epoch+1, step, len(train_loader), batch_time=batch_time,
data_time=data_time, loss=losses,
remain=timeSince(start, float(step+1)/len(train_loader)),
grad_norm=grad_norm,
#lr=scheduler.get_lr()[0],
))
return losses.avg
def valid_fn(valid_loader, model, criterion, device):
batch_time = AverageMeter()
data_time = AverageMeter()
losses = AverageMeter()
scores = AverageMeter()
# switch to evaluation mode
model.eval()
preds = []
start = end = time.time()
for step, (images, labels) in enumerate(valid_loader):
# measure data loading time
data_time.update(time.time() - end)
images = images.to(device)
labels = labels.to(device)
batch_size = labels.size(0)
# compute loss
with torch.no_grad():
y_preds = model(images)
loss = get_loss(criterion, y_preds, labels)
losses.update(loss.item(), batch_size)
# record accuracy
preds.append(y_preds.softmax(1).to('cpu').numpy())
if CFG['gradient_accumulation_steps'] > 1:
loss = loss / CFG['gradient_accumulation_steps']
# measure elapsed time
batch_time.update(time.time() - end)
end = time.time()
if step % CFG['print_freq'] == 0 or step == (len(valid_loader)-1):
print('EVAL: [{0}/{1}] '
'Data {data_time.val:.3f} ({data_time.avg:.3f}) '
'Elapsed {remain:s} '
'Loss: {loss.val:.4f}({loss.avg:.4f}) '
.format(
step, len(valid_loader), batch_time=batch_time,
data_time=data_time, loss=losses,
remain=timeSince(start, float(step+1)/len(valid_loader)),
))
predictions = np.concatenate(preds)
return losses.avg, predictions
def inference(model, states, test_loader, device):
model.to(device)
tk0 = tqdm(enumerate(test_loader), total=len(test_loader))
probs = []
for i, (images) in tk0:
images = images.to(device)
avg_preds = []
for state in states:
# model.load_state_dict(state['model'])
model.load_state_dict(state)
model.eval()
with torch.no_grad():
y_preds = model(images)
avg_preds.append(y_preds.softmax(1).to('cpu').numpy())
avg_preds = np.mean(avg_preds, axis=0)
probs.append(avg_preds)
probs = np.concatenate(probs)
return probs
```
# Train loop
```
# ====================================================
# scheduler
# ====================================================
def get_scheduler(optimizer):
if TAG['scheduler']=='ReduceLROnPlateau':
scheduler = ReduceLROnPlateau(optimizer, mode='min', factor=CFG['factor'], patience=CFG['patience'], verbose=True, eps=CFG['eps'])
elif TAG['scheduler']=='CosineAnnealingLR':
scheduler = CosineAnnealingLR(optimizer, T_max=CFG['T_max'], eta_min=CFG['min_lr'], last_epoch=-1)
elif TAG['scheduler']=='CosineAnnealingWarmRestarts':
scheduler = CosineAnnealingWarmRestarts(optimizer, T_0=CFG['T_0'], T_mult=1, eta_min=CFG['min_lr'], last_epoch=-1)
return scheduler
# ====================================================
# criterion
# ====================================================
def get_criterion():
if TAG['criterion']=='CrossEntropyLoss':
criterion = nn.CrossEntropyLoss()
elif TAG['criterion'] == 'bi_tempered_logistic_loss':
criterion = bi_tempered_logistic_loss
return criterion
# ====================================================
# Train loop
# ====================================================
def train_loop(folds, fold):
LOGGER.info(f"========== fold: {fold} training ==========")
if not CFG['debug']:
mlflow.set_tag('running.fold', str(fold))
# ====================================================
# loader
# ====================================================
trn_idx = folds[folds['fold'] != fold].index
val_idx = folds[folds['fold'] == fold].index
train_folds = folds.loc[trn_idx].reset_index(drop=True)
valid_folds = folds.loc[val_idx].reset_index(drop=True)
train_dataset = TrainDataset(train_folds,
transform=get_transforms(data='train'))
valid_dataset = TrainDataset(valid_folds,
transform=get_transforms(data='valid'))
train_loader = DataLoader(train_dataset,
batch_size=CFG['batch_size'],
shuffle=True,
num_workers=CFG['num_workers'], pin_memory=True, drop_last=True)
valid_loader = DataLoader(valid_dataset,
batch_size=CFG['batch_size'],
shuffle=False,
num_workers=CFG['num_workers'], pin_memory=True, drop_last=False)
# ====================================================
# model & optimizer & criterion
# ====================================================
best_model_path = OUTPUT_DIR+f'{TAG["model_name"]}_fold{fold}_best.pth'
latest_model_path = OUTPUT_DIR+f'{TAG["model_name"]}_fold{fold}_latest.pth'
model = CustomModel(TAG['model_name'], pretrained=True)
model.to(device)
# 学習途中の重みがあれば読み込み
if os.path.isfile(latest_model_path):
state_latest = torch.load(latest_model_path)
state_best = torch.load(best_model_path)
model.load_state_dict(state_latest['model'])
epoch_start = state_latest['epoch']+1
# er_best_score = state_latest['score']
er_counter = state_latest['counter']
er_best_score = state_best['best_score']
val_loss_history = state_latest['val_loss_history']
LOGGER.info(f'Load training model in epoch:{epoch_start}, best_score:{er_best_score:.3f}, counter:{er_counter}')
# 学習済みモデルを再学習する場合
elif os.path.isfile(best_model_path):
state_best = torch.load(best_model_path)
model.load_state_dict(state_best['model'])
epoch_start = 0 # epochは0からカウントしなおす
er_counter = 0
er_best_score = state_best['best_score']
val_loss_history = [] # 過去のval_lossも使用しない
LOGGER.info(f'Retrain model, best_score:{er_best_score:.3f}')
else:
epoch_start = 0
er_best_score = None
er_counter = 0
val_loss_history = []
optimizer = Adam(model.parameters(), lr=CFG['lr'], weight_decay=CFG['weight_decay'], amsgrad=False)
scheduler = get_scheduler(optimizer)
criterion = get_criterion()
# 再開時のepochまでschedulerを進める
# assert len(range(epoch_start)) == len(val_loss_history)
for _, val_loss in zip(range(epoch_start), val_loss_history):
if isinstance(scheduler, ReduceLROnPlateau):
scheduler.step(val_loss)
elif isinstance(scheduler, CosineAnnealingLR):
scheduler.step()
elif isinstance(scheduler, CosineAnnealingWarmRestarts):
scheduler.step()
# ====================================================
# apex
# ====================================================
if CFG['apex']:
model, optimizer = amp.initialize(model, optimizer, opt_level='O1', verbosity=0)
# ====================================================
# loop
# ====================================================
# best_score = 0.
# best_loss = np.inf
early_stopping = EarlyStopping(
patience=CFG['early_stopping_round'],
eps=CFG['early_stopping_eps'],
verbose=True,
save_path=best_model_path,
counter=er_counter, best_score=er_best_score,
val_loss_history = val_loss_history,
save_latest_path=latest_model_path)
for epoch in range(epoch_start, CFG['epochs']):
start_time = time.time()
# train
avg_loss = train_fn(train_loader, model, criterion, optimizer, epoch, scheduler, device)
# eval
avg_val_loss, preds = valid_fn(valid_loader, model, criterion, device)
valid_labels = valid_folds[CFG['target_col']].values
# scoring
score = get_score(valid_labels, preds.argmax(1))
# get learning rate
if hasattr(scheduler, 'get_last_lr'):
last_lr = scheduler.get_last_lr()[0]
else:
# ReduceLROnPlateauには関数get_last_lrがない
last_lr = optimizer.param_groups[0]['lr']
# log mlflow
if not CFG['debug']:
mlflow.log_metric(f"fold{fold} avg_train_loss", avg_loss, step=epoch)
mlflow.log_metric(f"fold{fold} avg_valid_loss", avg_val_loss, step=epoch)
mlflow.log_metric(f"fold{fold} score", score, step=epoch)
mlflow.log_metric(f"fold{fold} lr", last_lr, step=epoch)
# early stopping
early_stopping(avg_val_loss, model, preds, epoch)
if early_stopping.early_stop:
print(f'Epoch {epoch+1} - early stopping')
break
if isinstance(scheduler, ReduceLROnPlateau):
scheduler.step(avg_val_loss)
elif isinstance(scheduler, CosineAnnealingLR):
scheduler.step()
elif isinstance(scheduler, CosineAnnealingWarmRestarts):
scheduler.step()
elapsed = time.time() - start_time
LOGGER.info(f'Epoch {epoch+1} - avg_train_loss: {avg_loss:.4f} avg_val_loss: {avg_val_loss:.4f} time: {elapsed:.0f}s')
LOGGER.info(f'Epoch {epoch+1} - Accuracy: {score}')
# log mlflow
if not CFG['debug']:
mlflow.log_artifact(best_model_path)
if os.path.isfile(latest_model_path):
mlflow.log_artifact(latest_model_path)
check_point = torch.load(best_model_path)
valid_folds[[str(c) for c in range(5)]] = check_point['preds']
valid_folds['preds'] = check_point['preds'].argmax(1)
return valid_folds
def get_trained_fold_preds(folds, fold, best_model_path):
val_idx = folds[folds['fold'] == fold].index
valid_folds = folds.loc[val_idx].reset_index(drop=True)
check_point = torch.load(best_model_path)
valid_folds[[str(c) for c in range(5)]] = check_point['preds']
valid_folds['preds'] = check_point['preds'].argmax(1)
return valid_folds
def save_confusion_matrix(oof):
from sklearn.metrics import confusion_matrix
cm_ = confusion_matrix(oof['label'], oof['preds'], labels=[0,1,2,3,4])
label_name = ['0 (CBB)', '1 (CBSD)', '2 (CGM)', '3 (CMD)', '4 (Healthy)']
cm = pd.DataFrame(cm_, index=label_name, columns=label_name)
cm.to_csv(OUTPUT_DIR+'oof_confusion_matrix.csv', index=True)
# ====================================================
# main
# ====================================================
def get_result(result_df):
preds = result_df['preds'].values
labels = result_df[CFG['target_col']].values
score = get_score(labels, preds)
LOGGER.info(f'Score: {score:<.5f}')
return score
def main():
"""
Prepare: 1.train 2.test 3.submission 4.folds
"""
if CFG['train']:
# train
oof_df = pd.DataFrame()
for fold in range(CFG['n_fold']):
best_model_path = OUTPUT_DIR+f'{TAG["model_name"]}_fold{fold}_best.pth'
if fold in CFG['trn_fold']:
_oof_df = train_loop(folds, fold)
elif os.path.exists(best_model_path):
_oof_df = get_trained_fold_preds(folds, fold, best_model_path)
else:
_oof_df = None
if _oof_df is not None:
oof_df = pd.concat([oof_df, _oof_df])
LOGGER.info(f"========== fold: {fold} result ==========")
_ = get_result(_oof_df)
# CV result
LOGGER.info(f"========== CV ==========")
score = get_result(oof_df)
# save result
oof_df.to_csv(OUTPUT_DIR+'oof_df.csv', index=False)
save_confusion_matrix(oof_df)
# log mlflow
if not CFG['debug']:
mlflow.log_metric('oof score', score)
mlflow.delete_tag('running.fold')
mlflow.log_artifact(OUTPUT_DIR+'oof_df.csv')
if CFG['inference']:
# inference
model = CustomModel(TAG['model_name'], pretrained=False)
states = [torch.load(OUTPUT_DIR+f'{TAG["model_name"]}_fold{fold}_best.pth') for fold in CFG['trn_fold']]
test_dataset = TestDataset(test, transform=get_transforms(data='valid'))
test_loader = DataLoader(test_dataset, batch_size=CFG['batch_size'], shuffle=False,
num_workers=CFG['num_workers'], pin_memory=True)
predictions = inference(model, states, test_loader, device)
# submission
test['label'] = predictions.argmax(1)
test[['image_id', 'label']].to_csv(OUTPUT_DIR+'submission.csv', index=False)
```
# rerun
```
def _load_save_point(run_id):
# どこで中断したか取得
stop_fold = int(mlflow.get_run(run_id=run_id).to_dictionary()['data']['tags']['running.fold'])
# 学習対象のfoldを変更
CFG['trn_fold'] = [fold for fold in CFG['trn_fold'] if fold>=stop_fold]
# 学習済みモデルがあれば.pthファイルを取得(学習中も含む)
client = mlflow.tracking.MlflowClient()
artifacts = [artifact for artifact in client.list_artifacts(run_id) if ".pth" in artifact.path]
for artifact in artifacts:
client.download_artifacts(run_id, artifact.path, OUTPUT_DIR)
def check_have_run():
results = mlflow.search_runs(INFO['EXPERIMENT_ID'])
run_id_list = results[results['tags.mlflow.runName']==TITLE]['run_id'].tolist()
# 初めて実行する場合
if len(run_id_list) == 0:
run_id = None
# 既に実行されている場合
else:
assert len(run_id_list)==1
run_id = run_id_list[0]
_load_save_point(run_id)
return run_id
def push_github():
! cp {NOTEBOOK_PATH} kaggle-cassava/notebook/{TITLE}.ipynb
!git config --global user.email "raijin.1059@gmail.com"
! git config --global user.name "Raijin Shibata"
!cd kaggle-cassava ;git add .; git commit -m {TITLE}; git remote set-url origin https://{user_name}:{password}@github.com/raijin0704/kaggle-cassava.git; git push origin master
if __name__ == '__main__':
if CFG['debug']:
main()
else:
mlflow.set_tracking_uri(INFO['TRACKING_URI'])
mlflow.set_experiment('single model')
# 既に実行済みの場合は続きから実行する
run_id = check_have_run()
with mlflow.start_run(run_id=run_id, run_name=TITLE):
if run_id is None:
mlflow.log_artifact(CONFIG_PATH)
mlflow.log_param('device', device)
mlflow.set_tag('env', env)
mlflow.set_tags(TAG)
mlflow.log_params(CFG)
mlflow.log_artifact(notebook_path)
main()
mlflow.log_artifacts(OUTPUT_DIR)
remove_glob(f'{OUTPUT_DIR}/*latest.pth')
push_github()
if env=="kaggle":
shutil.copy2(CONFIG_PATH, f'{OUTPUT_DIR}/{CONFIG_NAME}')
! rm -r kaggle-cassava
elif env=="colab":
shutil.copytree(OUTPUT_DIR, f'{INFO["SHARE_DRIVE_PATH"]}/{TITLE}')
shutil.copy2(CONFIG_PATH, f'{INFO["SHARE_DRIVE_PATH"]}/{TITLE}/{CONFIG_NAME}')
```
| github_jupyter |
##### Copyright 2018 The TF-Agents Authors.
### Get Started
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/agents/blob/master/tf_agents/colabs/1_dqn_tutorial.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/agents/blob/master/tf_agents/colabs/1_dqn_tutorial.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
</table>
```
# Note: If you haven't installed the following dependencies, run:
!apt-get install xvfb
!pip install 'gym==0.10.11'
!pip install 'imageio==2.4.0'
!pip install PILLOW
!pip install pyglet
!pip install pyvirtualdisplay
!pip install tf-agents-nightly
!pip install tf-nightly
```
## Introduction
This example shows how to train a [DQN (Deep Q Networks)](https://storage.googleapis.com/deepmind-media/dqn/DQNNaturePaper.pdf) agent on the Cartpole environment using the TF-Agents library.

We will walk you through all the components in a Reinforcement Learning (RL) pipeline for training, evaluation and data collection.
## Setup
```
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import base64
import imageio
import IPython
import matplotlib
import matplotlib.pyplot as plt
import PIL.Image
import pyvirtualdisplay
import tensorflow as tf
from tf_agents.agents.dqn import dqn_agent
from tf_agents.drivers import dynamic_step_driver
from tf_agents.environments import suite_gym
from tf_agents.environments import tf_py_environment
from tf_agents.eval import metric_utils
from tf_agents.metrics import tf_metrics
from tf_agents.networks import q_network
from tf_agents.policies import random_tf_policy
from tf_agents.replay_buffers import tf_uniform_replay_buffer
from tf_agents.trajectories import trajectory
from tf_agents.utils import common
tf.compat.v1.enable_v2_behavior()
# Set up a virtual display for rendering OpenAI gym environments.
display = pyvirtualdisplay.Display(visible=0, size=(1400, 900)).start()
```
## Hyperparameters
```
env_name = 'CartPole-v0' # @param
num_iterations = 20000 # @param
initial_collect_steps = 1000 # @param
collect_steps_per_iteration = 1 # @param
replay_buffer_capacity = 100000 # @param
fc_layer_params = (100,)
batch_size = 64 # @param
learning_rate = 1e-3 # @param
log_interval = 200 # @param
num_eval_episodes = 10 # @param
eval_interval = 1000 # @param
```
## Environment
Environments in RL represent the task or problem that we are trying to solve. Standard environments can be easily created in TF-Agents using `suites`. We have different `suites` for loading environments from sources such as the OpenAI Gym, Atari, DM Control, etc., given a string environment name.
Now let us load the CartPole environment from the OpenAI Gym suite.
```
env = suite_gym.load(env_name)
```
We can render this environment to see how it looks. A free-swinging pole is attached to a cart. The goal is to move the cart right or left in order to keep the pole pointing up.
```
#@test {"skip": true}
env.reset()
PIL.Image.fromarray(env.render())
```
The `time_step = environment.step(action)` statement takes `action` in the environment. The `TimeStep` tuple returned contains the environment's next observation and reward for that action. The `time_step_spec()` and `action_spec()` methods in the environment return the specifications (types, shapes, bounds) of the `time_step` and `action` respectively.
```
print('Observation Spec:')
print(env.time_step_spec().observation)
print('Action Spec:')
print(env.action_spec())
```
So, we see that observation is an array of 4 floats: the position and velocity of the cart, and the angular position and velocity of the pole. Since only two actions are possible (move left or move right), the `action_spec` is a scalar where 0 means "move left" and 1 means "move right."
```
time_step = env.reset()
print('Time step:')
print(time_step)
action = 1
next_time_step = env.step(action)
print('Next time step:')
print(next_time_step)
```
Usually we create two environments: one for training and one for evaluation. Most environments are written in pure python, but they can be easily converted to TensorFlow using the `TFPyEnvironment` wrapper. The original environment's API uses numpy arrays, the `TFPyEnvironment` converts these to/from `Tensors` for you to more easily interact with TensorFlow policies and agents.
```
train_py_env = suite_gym.load(env_name)
eval_py_env = suite_gym.load(env_name)
train_env = tf_py_environment.TFPyEnvironment(train_py_env)
eval_env = tf_py_environment.TFPyEnvironment(eval_py_env)
```
## Agent
The algorithm that we use to solve an RL problem is represented as an `Agent`. In addition to the DQN agent, TF-Agents provides standard implementations of a variety of `Agents` such as [REINFORCE](http://www-anw.cs.umass.edu/~barto/courses/cs687/williams92simple.pdf), [DDPG](https://arxiv.org/pdf/1509.02971.pdf), [TD3](https://arxiv.org/pdf/1802.09477.pdf), [PPO](https://arxiv.org/abs/1707.06347) and [SAC](https://arxiv.org/abs/1801.01290).
The DQN agent can be used in any environment which has a discrete action space. To create a DQN Agent, we first need a `Q-Network` that can learn to predict `Q-Values` (expected return) for all actions given an observation from the environment.
We can easily create a `Q-Network` using the specs of the observations and actions. We can specify the layers in the network which, in this example, is the `fc_layer_params` argument set to a tuple of `ints` representing the sizes of each hidden layer (see the Hyperparameters section above).
```
q_net = q_network.QNetwork(
train_env.observation_spec(),
train_env.action_spec(),
fc_layer_params=fc_layer_params)
```
We also need an `optimizer` to train the network we just created, and a `train_step_counter` variable to keep track of how many times the network was updated.
```
optimizer = tf.compat.v1.train.AdamOptimizer(learning_rate=learning_rate)
train_step_counter = tf.compat.v2.Variable(0)
tf_agent = dqn_agent.DqnAgent(
train_env.time_step_spec(),
train_env.action_spec(),
q_network=q_net,
optimizer=optimizer,
td_errors_loss_fn=dqn_agent.element_wise_squared_loss,
train_step_counter=train_step_counter)
tf_agent.initialize()
```
## Policies
In TF-Agents, policies represent the standard notion of policies in RL: given a `time_step` produce an action or a distribution over actions. The main method is `policy_step = policy.step(time_step)` where `policy_step` is a named tuple `PolicyStep(action, state, info)`. The `policy_step.action` is the `action` to be applied to the environment, `state` represents the state for stateful (RNN) policies and `info` may contain auxiliary information such as log probabilities of the actions.
Agents contain two policies: the main policy that is used for evaluation/deployment (agent.policy) and another policy that is used for data collection (agent.collect_policy).
```
eval_policy = tf_agent.policy
collect_policy = tf_agent.collect_policy
```
We can also independently create policies that are not part of an agent. For example, a random policy:
```
random_policy = random_tf_policy.RandomTFPolicy(train_env.time_step_spec(),
train_env.action_spec())
```
## Metrics and Evaluation
The most common metric used to evaluate a policy is the average return. The return is the sum of rewards obtained while running a policy in an environment for an episode, and we usually average this over a few episodes. We can compute the average return metric as follows.
```
#@test {"skip": true}
def compute_avg_return(environment, policy, num_episodes=10):
total_return = 0.0
for _ in range(num_episodes):
time_step = environment.reset()
episode_return = 0.0
while not time_step.is_last():
action_step = policy.action(time_step)
time_step = environment.step(action_step.action)
episode_return += time_step.reward
total_return += episode_return
avg_return = total_return / num_episodes
return avg_return.numpy()[0]
compute_avg_return(eval_env, random_policy, num_eval_episodes)
# Please also see the metrics module for standard implementations of different
# metrics.
```
## Replay Buffer
In order to keep track of the data collected from the environment, we will use the TFUniformReplayBuffer. This replay buffer is constructed using specs describing the tensors that are to be stored, which can be obtained from the agent using `tf_agent.collect_data_spec`.
```
replay_buffer = tf_uniform_replay_buffer.TFUniformReplayBuffer(
data_spec=tf_agent.collect_data_spec,
batch_size=train_env.batch_size,
max_length=replay_buffer_capacity)
```
For most agents, the `collect_data_spec` is a `Trajectory` named tuple containing the observation, action, reward etc.
## Data Collection
Now let us execute the random policy in the environment for a few steps and record the data (observations, actions, rewards etc) in the replay buffer.
```
#@test {"skip": true}
def collect_step(environment, policy):
time_step = environment.current_time_step()
action_step = policy.action(time_step)
next_time_step = environment.step(action_step.action)
traj = trajectory.from_transition(time_step, action_step, next_time_step)
# Add trajectory to the replay buffer
replay_buffer.add_batch(traj)
for _ in range(initial_collect_steps):
collect_step(train_env, random_policy)
# This loop is so common in RL, that we provide standard implementations of
# these. For more details see the drivers module.
```
In order to sample data from the replay buffer, we will create a `tf.data` pipeline which we can feed to the agent for training later. We can specify the `sample_batch_size` to configure the number of items sampled from the replay buffer. We can also optimize the data pipline using parallel calls and prefetching.
In order to save space, we only store the current observation in each row of the replay buffer. But since the DQN Agent needs both the current and next observation to compute the loss, we always sample two adjacent rows for each item in the batch by setting `num_steps=2`.
```
# Dataset generates trajectories with shape [Bx2x...]
dataset = replay_buffer.as_dataset(
num_parallel_calls=3, sample_batch_size=batch_size, num_steps=2).prefetch(3)
iterator = iter(dataset)
```
## Training the agent
The training loop involves both collecting data from the environment and optimizing the agent's networks. Along the way, we will occasionally evaluate the agent's policy to see how we are doing.
The following will take ~5 minutes to run.
```
#@test {"skip": true}
%%time
# (Optional) Optimize by wrapping some of the code in a graph using TF function.
tf_agent.train = common.function(tf_agent.train)
# Reset the train step
tf_agent.train_step_counter.assign(0)
# Evaluate the agent's policy once before training.
avg_return = compute_avg_return(eval_env, tf_agent.policy, num_eval_episodes)
returns = [avg_return]
for _ in range(num_iterations):
# Collect a few steps using collect_policy and save to the replay buffer.
for _ in range(collect_steps_per_iteration):
collect_step(train_env, tf_agent.collect_policy)
# Sample a batch of data from the buffer and update the agent's network.
experience, unused_info = next(iterator)
train_loss = tf_agent.train(experience)
step = tf_agent.train_step_counter.numpy()
if step % log_interval == 0:
print('step = {0}: loss = {1}'.format(step, train_loss.loss))
if step % eval_interval == 0:
avg_return = compute_avg_return(eval_env, tf_agent.policy, num_eval_episodes)
print('step = {0}: Average Return = {1}'.format(step, avg_return))
returns.append(avg_return)
```
## Visualization
### Plots
We can plot return vs global steps to see the performance of our agent. In `Cartpole-v0`, the environment gives a reward of +1 for every time step the pole stays up, and since the maximum number of steps is 200, the maximum possible return is also 200.
```
#@test {"skip": true}
steps = range(0, num_iterations + 1, eval_interval)
plt.plot(steps, returns)
plt.ylabel('Average Return')
plt.xlabel('Step')
plt.ylim(top=250)
```
### Videos
It is helpful to visualize the performance of an agent by rendering the environment at each step. Before we do that, let us first create a function to embed videos in this colab.
```
def embed_mp4(filename):
"""Embeds an mp4 file in the notebook."""
video = open(filename,'rb').read()
b64 = base64.b64encode(video)
tag = '''
<video width="640" height="480" controls>
<source src="data:video/mp4;base64,{0}" type="video/mp4">
Your browser does not support the video tag.
</video>'''.format(b64.decode())
return IPython.display.HTML(tag)
```
The following code visualizes the agent's policy for a few episodes:
```
num_episodes = 3
video_filename = 'imageio.mp4'
with imageio.get_writer(video_filename, fps=60) as video:
for _ in range(num_episodes):
time_step = eval_env.reset()
video.append_data(eval_py_env.render())
while not time_step.is_last():
action_step = tf_agent.policy.action(time_step)
time_step = eval_env.step(action_step.action)
video.append_data(eval_py_env.render())
embed_mp4(video_filename)
```
| github_jupyter |
<img src="./pictures/logo_sizinglab.png" style="float:right; max-width: 60px; display: inline" alt="SizingLab" /></a>
## Dynamic models for propellers
*Written by Marc Budinger, Aitor Ochotorena(INSA Toulouse) and Scott Delbecq (ISAE-SUPAERO), Toulouse, France.*
The complexity of the propellers characteristics increases as they work in dynamic scenarios. So far, we have seen how $C_T$ and $C_P$ coefficients can be well expressed in terms of geometric ratio (pitch/diameter). In these dynamic scenarios, these coefficients depend also on the speed. Based on dimensional analysis and the polynomial regression of suppliers data, adequate prediction models for $C_T$ and $C_P$ are generated.
*Flow distribution in multirotor vehicle for climbing*

1. [Propeller regression](#section_1)
2. [Pareto filtering](#section_2)
<a id='section_1'></a>
### Propeller regression
### Data analysis
Based on the datasheets of different models of *APC propellers MR*, estimation models are developed to characterize the thrust and power coefficients of the propellers for dynamic scenarios. In the next DataFrame, an extra column is added to calculate the value of the dimensionless air compressibility estimation or B coefficient.
```
import pandas as pd
import math
# Read the .csv file with bearing data
path='./docs/'
df = pd.read_csv(path+'Propeller_Data.csv', sep=';')
K=101000 #[Pa] bulk modulus air
df['B coef']=1.18*(df['RPM']/60*2*3.14)**(2)*(df['DIAMETER_in']*0.0254)**(2)/K
# Print the head (first lines of the file)
df.head()
```
### Heatmap
The use of Heatmaps enables to visualize numerically the dependence of the parameters on each others to describe their behaviour. A value close to '1' will mean a very strong dependence, while a value close to '0' means a negligible effect. We see how $Cp$ and $Ct$ do not longer depend only on beta to describe their behaviour. A tutorial of seaborn heatmap can be found [here](https://likegeeks.com/seaborn-heatmap-tutorial/).
```
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
%matplotlib inline
f, axes = plt.subplots(1,2,figsize=(12,4))
df_Cp=df[['Cp','BETA','J', 'B coef']]
df_Ct=df[['Ct','BETA','J', 'B coef']]
# heatmap
sns.heatmap(df_Cp.corr(method='pearson')**2,annot=True,cmap='coolwarm',square=True, ax=axes[0])
sns.heatmap(df_Ct.corr(method='pearson')**2,annot=True,cmap='coolwarm',square=True, ax=axes[1])
```
### Scatter matrix
Another technique very similar to the heatmap to observe dependencies is the scatter matrix, in which we can represent in a matrix small subplots. Here we are interested in studying the trend of the parameter rather than knowing the exact value of the magnitude.
```
pd.plotting.scatter_matrix(df_Cp)
pd.plotting.scatter_matrix(df_Ct)
plt.tick_params(axis = 'both', labelsize = 14)
plt.rcParams.update({'font.size': 14})
```
### Surrogate modeling techniques
In this part, data are fitted using a non-linear polynomial model regression. For this purpose, a polynomial $(beta+ J)^n$ is developed using the package `PolynomialFeatures` of [scikit learn](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.PolynomialFeatures.html) to a degree n, whose value of determination coefficient ($R^2$) is higher than required.
As a condition, we set that the value of R squared required must be $R_{est}=0.98$
At the end, data and the surrogate model are plotted in a 3D graph.
Steps:
1. Read the dependent variable (in our case $C_T$ or $C_P$) and non-dependent variables ($\beta$ and J) from the dataframe.
2. We set up a minimum coefficient of determination $R^2$, for which the mathematical model must fit the data.
3. A binomial expansion in the form of $(\beta +J )^n$ is developed to fit the data values. The polynomial will raise until the R-squared value reaches the desired limit.
4. We plot together the values and the equation found in a 3D-graph and we print the expression.
- $C_p=f(\beta,J)$
```
from sklearn.preprocessing import PolynomialFeatures
from sklearn.metrics import mean_squared_error, r2_score
from mpl_toolkits.mplot3d import Axes3D
import matplotlib.pyplot as plt
import statsmodels.api as sm
import numpy as np
%matplotlib widget
R_est=0.98 # minimum R-squared value to achieve
# read dataframe values
X = df[['J', 'BETA']].values
y1 = df['Cp'].values
# extracting columns from X
mu_col=X[:,0]
beta_col=X[:,1]
#a matrix of mu and beta is made
XX = np.concatenate((mu_col.reshape(np.size(y1),1),beta_col.reshape(np.size(y1),1)),axis=1)
# in this while we increase the polynomial binomium until R2 reaches the limit
i=1 # start by 1.Grade polynomium
while True:
poly = PolynomialFeatures(i) # development of i-grade polynomium
x_poly = poly.fit_transform(XX)
model = sm.OLS(y1, x_poly) #fit regression model to the data
result= model.fit() # save value of coeficients
y_poly_pred = result.predict(x_poly) # according to the regression model found, a value of y is generated
rmse = np.sqrt(mean_squared_error(y1,y_poly_pred)) #Root mean square error
r2 = r2_score(y1,y_poly_pred) #R2 correlation value
i +=1 # we increase the polynomium degre
if r2>R_est: #if R2 calculated is greater than
break
print("RMSE=",rmse)
print("R2=",r2)
#here we expand the formula of the i-degree polynomium
data = pd.DataFrame.from_dict({
'J': np.random.randint(low=1, high=10, size=5),
'BETA': np.random.randint(low=-1, high=1, size=5),
})
p = PolynomialFeatures(degree=i-1).fit(data)
#concatenate the OLS regression parameters with variables
string1=['+({:.3f})*'.format(x) for x in result.params]
string2=p.get_feature_names(data.columns)
Result=""
for i in range(len(string1)):
Result += string1[i]
Result += string2[i]
print('Cp=',Result)
#meshgrid 3D for plot
xx1, xx2 = np.meshgrid(np.linspace(mu_col.min(), mu_col.max(), len(y1)),
np.linspace(beta_col.min(), beta_col.max(), len(y1)))
#create the 3D axis
fig = plt.figure(figsize=(10, 8))
ax = Axes3D(fig, azim=-210, elev=15)
ax.scatter(mu_col, beta_col, y1, color='black', alpha=1.0, facecolor='white')
# reshape data to suit in the grid. Here a x b = len(y1)
xp = np.reshape(mu_col, (94, 30))
yp = np.reshape(beta_col, (94, 30))
zp = np.reshape(y_poly_pred, (94, 30))
# plotting the predicted surface
ax.plot_surface(xp, yp, zp, cmap=plt.cm.RdBu_r, alpha=0.3, linewidth=0)
#axis labels
ax.set_xlabel('J (Advance ratio) [-]',fontsize=14)
ax.set_ylabel('angle (pitch/diameter) [-]',fontsize=14)
ax.set_zlabel(r'$C_P [-]$',fontsize=14)
plt.grid()
plt.tick_params(axis = 'both', labelsize = 14)
```
- $C_t=f(\beta,J)$
```
R_est=0.98 # minimum R-squared value to achieve
# read dataframe values
X = df[['J', 'BETA']].values
y1 = df['Ct'].values
# extracting columns from X
mu_col=X[:,0]
beta_col=X[:,1]
#a matrix of mu and beta is made
XX = np.concatenate((mu_col.reshape(np.size(y1),1),beta_col.reshape(np.size(y1),1)),axis=1)
# in this while we increase the polynomial binomium until R2 reaches the limit
i=1 # start by 1.Grade polynomium
while True:
poly = PolynomialFeatures(i) # development of i-grade polynomium
x_poly = poly.fit_transform(XX)
model = sm.OLS(y1, x_poly) #fit regression model to the data
result= model.fit() # save value of coeficients
y_poly_pred = result.predict(x_poly) # according to the regression model found, a value of y is generated
rmse = np.sqrt(mean_squared_error(y1,y_poly_pred)) #Root mean square error
r2 = r2_score(y1,y_poly_pred) #R2 correlation value
i +=1 # we increase the polynomium degre
if r2>R_est: #if R2 calculated is greater than
break
print("RMSE=",rmse)
print("R2=",r2)
#here we expand the formula of the i-degree polynomium
data = pd.DataFrame.from_dict({
'mu': np.random.randint(low=1, high=10, size=5),
'beta': np.random.randint(low=-1, high=1, size=5),
})
p = PolynomialFeatures(degree=i-1).fit(data)
#concatenate the OLS regression parameters with variables
string1=['+({:.3f})*'.format(x) for x in result.params]
string2=p.get_feature_names(data.columns)
Result=""
for i in range(len(string1)):
Result += string1[i]
Result += string2[i]
print('Ct=',Result)
#meshgrid 3D for plot
xx1, xx2 = np.meshgrid(np.linspace(mu_col.min(), mu_col.max(), len(y1)),
np.linspace(beta_col.min(), beta_col.max(), len(y1)))
#create the 3D axis
fig = plt.figure(figsize=(10, 8))
ax = Axes3D(fig, azim=-210, elev=15)
ax.scatter(mu_col, beta_col, y1, color='black', alpha=1.0, facecolor='white')
# reshape data to suit in the grid. Here a x b = len(y1)
xp = np.reshape(mu_col, (94, 30))
yp = np.reshape(beta_col, (94, 30))
zp = np.reshape(y_poly_pred, (94, 30))
# plotting the predicted surface
ax.plot_surface(xp, yp, zp, cmap=plt.cm.RdBu_r, alpha=0.3, linewidth=0)
#axis labels
ax.set_xlabel('J (Advance ratio) [-]',fontsize=14)
ax.set_ylabel('angle (pitch/diameter) [-]',fontsize=14)
ax.set_zlabel(r'$C_T [-]$',fontsize=14)
plt.grid()
plt.tick_params(axis = 'both', labelsize = 14)
```
<a id='section_4'></a>
<a id='section_2'></a>
### Pareto filtering: High thrust and small diameter
```
# This function tests if a component is dominated
# return 0 if non dominated, the number of domination other else
# inputs :
# x_,y_ : the component's characteristics to test
# X_,Y_ : the data set characteristics
def dominated(x_,y_,X_,Y_):
compteur=0
for a,b in zip(X_,Y_):
# a>x_ for high torque and b<y_ for small weight
if (a>x_) and (b<y_):
compteur +=1
return compteur
import matplotlib.pyplot as plt
import pandas as pd
path='./data/'
df_pro = pd.read_csv(path+'Propeller_Data.csv', sep=';')
df_pro['Dominated']=0
df_pro = df_pro.reset_index(drop=True)# clear row index after ignoring one family battery
for row in range(len(df_pro['Ct'])):
if dominated(df_pro.loc[row,'Thrust'], df_pro.loc[row,'DIAMETER'],df_pro['Thrust'].values,df_pro['DIAMETER'].values)>0:
df_pro.loc[row,'Dominated']=1
color_wheel = {0: 'r',
1: 'b'}
linewidth = {0: 300,
1: 100}
colors = df_pro['Dominated'].map(lambda x: color_wheel.get(x))
s = df_pro['Dominated'].map(lambda x: linewidth.get(x))
df_pro[df_pro['Dominated']==0].to_csv(r'./data/Non-Dominated-Propeller.csv',sep=';')#We save the non-dominated series to process after.
pd.plotting.scatter_matrix(df_pro[['BETA','DIAMETER','Ct','Cp']], color=colors, figsize=[15,10], s=s);
plt.tick_params(axis = 'both', labelsize = 14)
plt.rcParams.update({'font.size': 14})
```
### References
- Principles of Helicopter Aerodynamics. Leishman
- [MR Propellers](https://www.masterairscrew.com/pages/mr-drone-propellers)
- [Drones](https://www.droneomega.com/quadcopter-propeller/)
| github_jupyter |
# Pivot review data with Elastic data frames
This notebook shows how data can be pivoted with [Elastic data frames](https://www.elastic.co/guide/en/elastic-stack-overview/master/ml-dataframes.html) to reveal insights into the behaviour of reviewers. The use case and data is from Mark Harwood's talk on [entity-centric indexing](https://www.elastic.co/videos/entity-centric-indexing-mark-harwood).
An alternative version of this notebook uses python [pandas](https://pandas.pydata.org/) to create the same results.
```
import bz2
import matplotlib.pyplot as plt
import csv
import time
import pandas as pd
from elasticsearch import helpers
from elasticsearch import Elasticsearch
from elasticsearch.exceptions import NotFoundError
```
## Connect to Elasticsearch
First connect to Elasticsearch. This assumes access is via `localhost:9200`, change next line to change connection parameters (see https://elasticsearch-py.readthedocs.io/en/master/api.html).
```
es = Elasticsearch()
```
## Read data to Elasticsearch
Note this deletes and creates indices.
```
index_name = "anonreviews"
index_name_pivot = 'anonreviews_pivot'
index_settings = { "settings": { "number_of_shards": 1, "number_of_replicas": 0 }, "mappings": { "properties": { "reviewerId": { "type": "keyword" }, "vendorId": { "type": "keyword" }, "date": { "type": "date", "format" : "yyyy-MM-dd HH:mm" }, "rating": { "type": "integer" } } } }
es.indices.delete(index=index_name, ignore=[400, 404])
es.indices.create(index=index_name, body=index_settings)
actions = []
bulk_batch_size = 10000
n = 0
csv_handle = bz2.open('./anonreviews.csv.bz2', 'rt')
csv_reader = csv.DictReader(csv_handle)
for row in csv_reader:
n += 1
action = { "_index": index_name, "_source": row }
actions.append(action)
if n % bulk_batch_size == 0:
helpers.bulk(es, actions)
actions = []
if len(actions) > 0:
helpers.bulk(es, actions)
```
Flush to ensure all docs are indexed, then summarise indexing.
```
es.indices.flush(index=index_name)
es.count(index=index_name)
```
## Aggregate and Pivot data
Pivot data so we get summaries for each reviewer.
In pandas, we do the following:
```
aggregations = {
'rating':'mean',
'vendorId':'nunique',
'reviewerId':'count'
}
grouped = reviews.groupby('reviewerId').agg(aggregations)
grouped.columns=['avg_rating', 'dc_vendorId', 'count']
```
In Elasticsearch we can use data frames to achieve the same transformation. First preview the transformation:
```
request = {
"source": {
"index": index_name
},
"dest": {
"index": index_name_pivot
},
"pivot": {
"group_by": {
"reviewerId": {
"terms": {
"field": "reviewerId"
}
}
},
"aggregations": {
"avg_rating": {
"avg": {
"field": "rating"
}
},
"dc_vendorId": {
"cardinality": {
"field": "vendorId"
}
},
"count": {
"value_count": {
"field": "_id"
}
}
}
}
}
response = es.transport.perform_request('POST', '/_data_frame/transforms/_preview', body=request)
response['preview'][0:10]
```
### Create Elastic Data Frame
```
# First delete old index if it exists
es.indices.delete(index='anonreviews_pivot', ignore=[400, 404])
# Stop and delete any old jobs (ignore if they don't exist)
try:
es.transport.perform_request('POST', '/_data_frame/transforms/anonreviews_pivot/_stop')
es.transport.perform_request('DELETE', '/_data_frame/transforms/anonreviews_pivot')
except NotFoundError:
pass
# Now create data frame job (called anonreviews_pivot)
es.transport.perform_request('PUT', '/_data_frame/transforms/anonreviews_pivot', body=request)
# Start job
es.transport.perform_request('POST', '/_data_frame/transforms/anonreviews_pivot/_start')
# Poll for progress
while True:
response = es.transport.perform_request('GET', '/_data_frame/transforms/anonreviews_pivot/_stats')
if response['transforms'][0]['state']['task_state'] == 'stopped':
print(response['transforms'][0]['state']['progress'])
break
if 'progress' in response['transforms'][0]['state']:
print(response['transforms'][0]['state']['progress'])
time.sleep(5)
def hits_to_df(response, create_index=True):
hits = []
index = []
for hit in response['hits']['hits']:
hits.append(hit['_source'])
index.append(hit['_source']['reviewerId'])
if create_index:
return pd.DataFrame(hits, index=index)
else:
return pd.DataFrame(hits)
```
### Find 'haters'
Reviewers that give more than five zero star reviews to one vendor
```
q = "dc_vendorId:1 AND count :>5 AND avg_rating:0"
sort = "count:desc"
response = es.search(index='anonreviews_pivot', q=q, sort=sort, size=100)
hits_to_df(response)
```
For example, reviewer 10392 gives 94 zero star reviews to vendor 122
```
q = "reviewerId:10392"
response = es.search(index='anonreviews', q=q, size=5) # top 5 only
hits_to_df(response, False)
```
### Find 'fanboys'
Reviewers that give more than five five star reviews to one vendor
```
q = "dc_vendorId:1 AND count :>5 AND avg_rating:5"
sort = "count:desc"
response = es.search(index='anonreviews_pivot', q=q, sort=sort, size=100)
hits_to_df(response)
```
Reviewer 183751 gives 73 five star reviews to vendor 190
```
q = "reviewerId:183751"
response = es.search(index='anonreviews', q=q, size=5) # top 5 only
hits_to_df(response, False)
```
| github_jupyter |
# Module 3.1
Goal: make code
- **readable**
- **reuseable**
- testable
Approach: **modularization** (create small reuseable pieces of code)
- functions
- modules
## Unit 5: Functions
Python functions
* `len(a)`
* `max(a)`
* `sum(a)`
* `range(start, stop, step)` or `range(stop)`
* `print(s)` or `print(s1, s2, ...)` or `print(s, end=" ")`
Functions have
* **name**
* **arguments** (in **parentheses**) — optional (but _always_ parentheses)
* **return value** — (can be `None`)
```
arg = ['a', 'b', 'c']
retval = len(arg)
retval
retval = print("hello")
print(retval)
retval is None
```
### Defining functions
```python
def func_name(arg1, arg2, ...):
"""documentation string (optional)"""
# body
...
return results
```
[**Heaviside** step function](http://mathworld.wolfram.com/HeavisideStepFunction.html) (again):
$$
\Theta(x) = \begin{cases}
0 & x < 0 \\
\frac{1}{2} & x = 0\\
1 & x > 0
\end{cases}
$$
```
x = 1.23
theta = None
if x < 0:
theta = 0
elif x > 0:
theta = 1
else:
theta = 0.5
print("theta({0}) = {1}".format(x, theta))
```
[**Heaviside** step function](http://mathworld.wolfram.com/HeavisideStepFunction.html) (again):
$$
\Theta(x) = \begin{cases}
0 & x < 0 \\
\frac{1}{2} & x = 0\\
1 & x > 0
\end{cases}
$$
```
def Heaviside(x):
theta = None
if x < 0:
theta = 0
elif x > 0:
theta = 1
else:
theta = 0.5
return theta
def Heaviside(x):
theta = None
if x < 0:
theta = 0
elif x > 0:
theta = 1
else:
theta = 0.5
return theta
```
Now **call** the function:
```
Heaviside(0)
Heaviside(1.2)
```
Add doc string:
```
def Heaviside(x):
"""Heaviside step function
Parameters
----------
x : float
Returns
-------
float
"""
theta = None
if x < 0:
theta = 0
elif x > 0:
theta = 1
else:
theta = 0.5
return theta
help(Heaviside)
```
Make code more concise:
```
def Heaviside(x):
"""Heaviside step function
Parameters
----------
x : float
Returns
-------
float
"""
if x < 0:
return 0
elif x > 0:
return 1
return 0.5
X = [i*0.5 for i in range(-3, 4)]
Y = [Heaviside(x) for x in X]
print(list(zip(X, Y)))
```
### Multiple return values
Functions always return a single object.
- `None`
- basic data type (float, int, str, ...)
- container data type, e.g. a list or a **tuple**
- _any_ object
Move a particle at coordinate `r = [x, y]` by a translation vector `[tx, ty]`:
```
def translate(r, t):
"""Return r + t for 2D vectors r, t"""
x1 = r[0] + t[0]
y1 = r[1] + t[1]
return [x1, y1]
pos = [1, -1]
tvec = [9, 1]
new_pos = translate(pos, tvec)
new_pos
```
[Metal umlaut](https://en.wikipedia.org/wiki/Metal_umlaut) search and replace: replace all "o" with "ö" and "u" with "ü": return the new string and the number of replacements.
```
def metal_umlaut_search_replace(name):
new_name = ""
counter = 0
for char in name:
if char == "o":
char = "ö"
counter += 1
elif char == "u":
char = "ü"
counter += 1
new_name += char
return new_name, counter # returns the tuple (new_name, counter)
metal_umlaut_search_replace("Motely Crue")
retval = metal_umlaut_search_replace("Motely Crue")
type(retval)
retval[0]
```
Use *tuple unpacking* to get the returned values:
```
name, n = metal_umlaut_search_replace("Motely Crue")
print(name, "rocks", n, "times harder now!")
```
### Variable argument lists
Functions have arguments in their _call signature_, e.g., the `x` in `def Heaviside(x)` or `x` and `y` in a function `area()`:
```
def area(x, y):
"""Calculate area of rectangle with lengths x and y"""
return x*y
```
Add functionality: calculate the area when you scale the rectangle with a factor `scale`.
```
def area(x, y, scale):
"""Calculate scaled area of rectangle with lengths x and y and scale factor scale"""
return scale*x*y
```
**Inconvenience**: even for unscaled rectangles I always have to provide `scale=1`, i.e.,
```python
area(Lx, Ly, 1)
```
**Optional argument** with **default value**:
```
def area(x, y, scale=1):
"""Calculate scaled area of rectangle with lengths `x` and `y`.
scale factor `scale` defaults to 1
"""
return scale*x*y
Lx, Ly = 2, 10.5
print(area(Lx, Ly)) # uses scale=1
print(area(Lx, Ly, scale=0.5))
print(area(Lx, Ly, scale=2))
print(area(Lx, Ly, 2)) # DISCOURAGED, use scale=2
```
#### Variable arguments summary
```python
def func(arg1, arg2, kwarg1="abc", kwarg2=None, kwarg3=1):
...
```
* *positional arguments* (`arg1`, `arg2`):
* all need to be provided when function is called: `func(a, b)`
* must be in given order
* *keyword arguments* `kwarg1="abc" ...`:
* optional; set to default if not provided
* no fixed order: `func(a, b, kwarg2=1000, kwarg1="other")`
See more under [More on Defining Functions](https://docs.python.org/3/tutorial/controlflow.html#more-on-defining-functions) in the Python Tutorial.
## Unit 6: Modules and packages
_Modules_ (and _packages_) are **libraries** of reuseable code blocks (e.g. functions).
Example: `math` module
```
import math
math.sin(math.pi/3)
```
### Creating a module
A module is just a file with Python code.
Create `physics.py` with content
```python
# PHY194 physics module
pi = 3.14159
h = 6.62606957e-34
def Heaviside(x):
"""Heaviside function Theta(x)"""
if x < 0:
return 0
elif x > 0:
return 1
return 0.5
```
### Importing
Import it
```
import physics
```
*Note*: `physics.py` must be in the same directory!
Access contents with the dot `.` operator:
```
two_pi = 2 * physics.pi
h_bar = physics.h / two_pi
print(h_bar)
physics.Heaviside(2)
```
**Direct import** (use sparingly as it can become messy)
```
from physics import h, pi
h_bar = h / (2*pi)
print(h_bar)
```
**Aliased** import:
```
import physics as phy
h_bar = phy.h / (2*phy.pi)
print(h_bar)
```
### The Standard Library
See [The Python Standard Library](https://docs.python.org/3/library/index.html).
| github_jupyter |
FaceRec Evaluation 3: MeMAD - YLE Dataset
====================================
#### NOT COMPLETED
This notebook serves as evaluation for the face recognition system.
It contains the code for 1. **create a ground truth** and 2. **test the system on it**.
```
import os
import sys
sys.path.insert(0, os.path.abspath('../'))
import re
from tqdm.notebook import tqdm
import numpy as np
import pandas as pd
from SPARQLWrapper import SPARQLWrapper, JSON
from frame_collector import FrameCollector
from src.utils.media_fragment import convert_to_seconds_npt
from src.connectors import memad_connector as memad
from src.connectors import limecraft_connector as limecraft
```
## Part 1: Create a Ground Truth
The [MeMAD Knowledge Graph](http://data.memad.eu) contains information about the presence of people in particular video segments.
The following cells retrieve the 6 people with the highest number of appearence, for then collecting all segments (+ media uri and start/end time) involving them.
For clarity, we will call _media_ an entire video resource (e.g. a mpeg4 file), while with _segment_ a temporal fragment of variable length, to not be confused with _shots_. See definitions of MediaResource and Part in the [ebucore ontology](https://www.ebu.ch/metadata/ontologies/ebucore/)
Problem: Yle has no parts
```
ENDPOINT = "http://data.memad.eu/sparql-endpoint"
sparql = SPARQLWrapper(ENDPOINT)
sparql.setReturnFormat(JSON)
with open('memad_people.rq') as file:
query = file.read()
sparql.setQuery(query)
people = sparql.query().convert()["results"]["bindings"]
for x in people:
for k in x:
x[k] = x[k]['value']
df_people = pd.DataFrame.from_dict(people)
df_people['nb'] = df_people['nb'].apply(pd.to_numeric)
df_people
```
Select top 10 celebrities
```
people_uri = []
people_labels = []
iterator = df_people.itertuples()
while len(people_uri) < 10:
p = next(iterator)
if p.c not in people_uri:
people_uri.append(p.c)
people_labels.append(p.label)
list(zip(people_uri, people_labels))
with open('memad_program_by_person.rq') as file:
query = file.read()
from src.utils.media_fragment import convert_to_seconds_npt
def pad_spaces(text):
pad = 25 - len(text)
return text + ''.join([' ' for _ in range(0,pad)])
def get_segments(person, name):
q = query.replace('?person', '<%s>' % person)
sparql.setQuery(q)
results = sparql.query().convert()["results"]["bindings"]
for r in results:
for p in r:
value = r[p]['value']
if p in ['start', 'end']:
value = convert_to_seconds_npt(re.sub(r"T(\d{2}:\d{2}:\d{2}).+", "\g<1>", value))
if p == 'prop':
value = re.split(r"[/#]", value)[-1]
r[p] = value
r['person'] = name
r['person_uri'] = person
results = [r for r in results if 'start' in r and r['start'] < r['end']]
print('- %s\t%s'%(pad_spaces(name), len(results)))
return pd.DataFrame.from_dict(results)
print('Num. results per person:')
df = pd.concat([get_segments(uri, name)
for uri, name in tqdm(zip(people_uri, people_labels), total=len(people_uri))],
ignore_index=True)
df
df['media'].value_counts()
df['p'].value_counts()
df['duration'] = df['end'] - df['start']
df['duration'].describe()
df_subset = df[df['duration'] < 50]
df_subset = df_subset[df_subset['duration'] > 20]
print('Subset length: %d' %len(df_subset))
df_subset['person_uri'].value_counts()
from src.connectors import memad_connector as memad
from src.connectors import limecraft_connector as limecraft
old_loc = ''
fc = None
df_subset.sort_values(by='media', ascending=False)
table = df_subset
def ex_frame(row):
global fc
global old_loc
loc = memad.get_locator_for(row['media'])
if not loc:
print('No loc for: '+ row['media'])
return 0
loc = limecraft.locator2video(loc['locator']['value'])
if loc != old_loc:
fc = FrameCollector(loc, 'memad_gt2', id=row['media'])
old_loc = loc
start = row['start']
end = row['end']
duration = end - start
quarter = int(duration / 4)
quarter1 = start + quarter
center = start + quarter * 2
quarter2 = start + quarter * 3
try:
x = fc.run(frame_no=quarter1)
if x == 0:
x = fc.run(frame_no=center)
if x == 0:
x = fc.run(frame_no=quarter2)
return x
except:
print('Error: ' + loc)
return 0
frames = [ex_frame(row) for idx, row in tqdm(table.iterrows(), total=len(table))]
frames = [l if l is not None else 0 for l in frames]
df_subset = df_subset.reset_index(drop=True)
df_subset['frame'] = frames
df_extracted = df_subset[df_subset['frame'] != 0]
df_extracted
df_extracted['person'].value_counts()
df_extracted.to_csv(r'memad_parts2.csv', index = False)
```
### Plan
1. Extract 1 folders for each person
2. Manually remove the ones not containing the person
3. Re-import in this notebook and fix the table
##### 1. Extract 3 folders for each person
```
from shutil import copyfile
for p in df_extracted['person'].unique():
out_path = './dataset_memad2/%s' % p.replace(' ', '_')
os.makedirs(out_path, exist_ok=True)
for i, row in df_extracted[df_extracted['person'] == p].iterrows():
x = row['frame']
fname = x.split('/')[-1]
copyfile(x, os.path.join(out_path, fname))
```
##### 2. Manually move those to 2 groups: containing and not containing the person
_(This action is made offline)_
##### 3. Re-import in this notebook and fix the table
```
manual_good = []
manual_bad = []
for p in os.listdir('./dataset_memad'):
if p == '.DS_Store':
continue
for f in os.listdir(os.path.join('./dataset_memad', p)):
if f == '.DS_Store':
continue
if p == 'unknown':
manual_bad.append('./frames/memad_gt/'+f)
else:
manual_good.append((p,'./frames/memad_gt/'+f))
unique, counts = np.unique(np.array(manual_good)[:,0], return_counts=True)
for a,b in zip(unique,counts):
print(a, b)
```
| github_jupyter |
```
from __future__ import print_function
from matplotlib import pyplot as plt
import matplotlib.patches as mpatches
%matplotlib inline
import sys
sys.path.insert(0, '../py')
from graviti import *
import numpy as np
import scipy as sp
from scipy.sparse import coo_matrix
import scipy.ndimage as ndi
from skimage.draw import polygon
from skimage import io
from skimage.measure import label, regionprops
import skimage.io
import skimage.measure
import skimage.color
import glob
import pickle
import pandas as pd
import os
import timeit
import random
import pyvips
import timeit
import multiprocessing
from joblib import Parallel, delayed
from datetime import datetime
from tqdm import tqdm
import umap
import seaborn as sns; sns.set()
def get_LogCovMat_parallel(featureMat):
out = np.real(sp.linalg.logm(np.cov(featureMat,rowvar=False))).flatten()
return out
def get_LogCorrMat_parallel(featureMat):
return np.real(sp.linalg.logm(np.corrcoef(featureMat,rowvar=False))).flatten()
def get_position_parallel(featureMat):
pos = featureMat[0]
return pos
patches = glob.glob('/home/garner1/pipelines/nucleAI/data/TCGA-05-4402-01Z-00-DX1/*.pkl')
intensity_features = []
for patch in patches[:]:
#print(patch)
infile = open(patch,'rb')
lista = pickle.load(infile)
intensity_features.extend(lista)
infile.close()
print(len(intensity_features))
list_of_features = random.sample(intensity_features,3000)
num_cores = multiprocessing.cpu_count() # numb of cores
pos = Parallel(n_jobs=num_cores)( delayed(get_position_parallel)(f) for f in tqdm(intensity_features) if f is not None)
data_cov = Parallel(n_jobs=num_cores)( delayed(get_LogCovMat_parallel)(f[1]) for f in tqdm(intensity_features) if f is not None)
data_corrcoef = Parallel(n_jobs=num_cores)( delayed(get_LogCorrMat_parallel)(f[1]) for f in tqdm(intensity_features) if f is not None)
#Get data in a serial way
#pos = np.array([f[0] for f in list_of_features if f is not None])
#data_cov = np.array([np.real(sp.linalg.logm(np.cov(f[1],rowvar=False))).flatten() for f in list_of_features if f is not None])
#data_corrcoef = np.array([np.real(sp.linalg.logm(np.corrcoef(f[1],rowvar=False))).flatten() for f in list_of_features if f is not None])
reducer = umap.UMAP(n_components=2,min_dist=0,n_neighbors=10)
embedding = reducer.fit_transform(data_cov)
# Define dataframe in lowD
df = pd.DataFrame(dict(x=embedding[:,0], y=embedding[:,1]))
import seaborn
ax = seaborn.FacetGrid(data=df,height=10, aspect=1)
ax.map(plt.scatter, 'x', 'y',s=20).add_legend()
plt.gca().invert_yaxis()
barycenter_cov = np.mean(data_cov,axis=0)
dist = sp.linalg.norm(data_cov-barycenter_cov,axis=1)
print(np.mean(dist),np.std(dist),np.cov(dist))
ax = seaborn.FacetGrid(data=df_plot[['x','y','cluster']],
hue='cluster',
height=10, aspect=1)
ax.map(plt.scatter, 'x', 'y',s=20).add_legend()
plt.gca().invert_yaxis()
# K-means classification
from sklearn.cluster import KMeans, DBSCAN
import hdbscan
df = df_plot[df_plot['cluster']==3]
db = hdbscan.HDBSCAN(min_cluster_size=10).fit(df[['x_umap','y_umap']]
)
#kmeans = KMeans(n_clusters=2, random_state=0).fit(embedding)
df['cluster_B'] = db.labels_
# Show the cluster to study
import seaborn
seaborn.set(style='white')
fg = seaborn.FacetGrid(data=df[['x_umap','y_umap','cluster_B']],
height=10, aspect=1,
hue='cluster_B')
fg.map(plt.scatter, 'x_umap', 'y_umap',s=10).add_legend()
x = pos[:,0]
y = pos[:,1]
df_plot = pd.DataFrame(dict(x=x, y=y))
fig, ax = plt.subplots(figsize=(10,10))
ax = sns.scatterplot(x="x", y="y", s=10,data=df_plot)
ax.invert_yaxis()
embedding = reducer.fit_transform(data_cov)
df_plot['x_umap'] = embedding[:,0]
df_plot['y_umap'] = embedding[:,1]
fig, ax = plt.subplots(figsize=(10,10))
ax = sns.scatterplot(x="x_umap", y="y_umap", s=10, data=df_plot)
# K-means classification
from sklearn.cluster import KMeans, DBSCAN
import hdbscan
db = hdbscan.HDBSCAN(min_cluster_size=100).fit(embedding)
#kmeans = KMeans(n_clusters=2, random_state=0).fit(embedding)
df_plot['cluster'] = db.labels_
# Show the cluster to study
import seaborn
seaborn.set(style='white')
fg = seaborn.FacetGrid(data=df_plot[['x_umap','y_umap','cluster']],
height=10, aspect=1,
hue='cluster')
fg.map(plt.scatter, 'x_umap', 'y_umap',s=10).add_legend()
ax = seaborn.FacetGrid(data=df_plot[['x','y','cluster']],
hue='cluster',
height=10, aspect=1)
ax.map(plt.scatter, 'x', 'y',s=20).add_legend()
plt.gca().invert_yaxis()
# K-means classification
from sklearn.cluster import KMeans, DBSCAN
import hdbscan
df = df_plot[df_plot['cluster']==3]
db = hdbscan.HDBSCAN(min_cluster_size=10).fit(df[['x_umap','y_umap']]
)
#kmeans = KMeans(n_clusters=2, random_state=0).fit(embedding)
df['cluster_B'] = db.labels_
# Show the cluster to study
import seaborn
seaborn.set(style='white')
fg = seaborn.FacetGrid(data=df[['x_umap','y_umap','cluster_B']],
height=10, aspect=1,
hue='cluster_B')
fg.map(plt.scatter, 'x_umap', 'y_umap',s=10).add_legend()
ax = seaborn.FacetGrid(data=df[['x','y','cluster_B']],
hue='cluster_B',
height=10, aspect=1)
ax.map(plt.scatter, 'x', 'y',s=20).add_legend()
plt.gca().invert_yaxis()
```
| github_jupyter |
# Performance Metric Investigation
```
import pandas as pd, numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import time
import os
from tqdm import tqdm
sns.set(style='darkgrid')
%matplotlib inline
def clean_ohlc(ohlc_df):
ohlc_df.loc[ohlc_df['Open'] > ohlc_df['High'], 'High'] = ohlc_df['Open']
ohlc_df.loc[ohlc_df['Open'] < ohlc_df['Low'], 'Low'] = ohlc_df['Open']
ohlc_df.loc[ohlc_df['Close'] > ohlc_df['High'], 'High'] = ohlc_df['Close']
ohlc_df.loc[ohlc_df['Close'] < ohlc_df['Low'], 'Low'] = ohlc_df['Close']
return ohlc_df
def resample_ohlc(ohlc_df, freq='D'):
resample_df = pd.DataFrame()
resample_df['Open'] = ohlc_df['Open'].resample(freq).first().copy()
resample_df['High'] = ohlc_df['High'].resample(freq).max().copy()
resample_df['Low'] = ohlc_df['Low'].resample(freq).min().copy()
resample_df['Close'] = ohlc_df['Close'].resample(freq).last().copy()
resample_df['Volume'] = ohlc_df['Volume'].resample(freq).sum().copy()
return resample_df
```
## Load the Data
```
# Create a blank list of returns
returns_list = []
# Load some bitcoin data
xbt_1d = pd.read_csv('data/XBTUSD_1d.csv', parse_dates=True, index_col=0)
# Add the returns to the list
xbt_1d['returns'] = xbt_1d['Close'].pct_change().dropna()
returns_list.append(xbt_1d['returns'])
# Load in some altcoin data
for altcoin in ['BCH', 'ETH', 'LTC', 'XRP']:
filepath = 'data/' + altcoin + 'XBT_1d.csv'
df = pd.read_csv(filepath, parse_dates=True, index_col=0)
returns = df['Close'].pct_change().dropna()
returns_list.append(returns)
# Load in some ETF data
path = '/Users/bblandin/Dropbox/Ground-Up Development/Data/Flat Files (Raw)/ETFs'
data_list = os.listdir(path)
for file in data_list:
file_path = path + '/' + file
df = pd.read_csv(file_path, parse_dates=True, index_col=0)
returns = df['Adj Close'].pct_change().dropna()
returns_list.append(returns)
```
### Equity Curve Metrics
These are path-dependent and require Monte Carlo simulation to assess accurately
```
def equity_curve(returns):
eq = (1 + returns).cumprod(axis=0)
if len(eq.shape) > 1:
return pd.DataFrame(eq)
else:
return pd.Series(eq)
# return eq
def drawdown(equity_curve):
_drawdown = equity_curve / equity_curve.cummax() - 1
return _drawdown
def max_drawdown(equity_curve, percent=True):
abs_drawdown = np.abs(drawdown(equity_curve))
_max_drawdown = np.max(abs_drawdown)
if percent == True:
return _max_drawdown * 100
else:
return _max_drawdown
def ulcer_index(equity_curve):
_drawdown = drawdown(equity_curve)
_ulcer_index = np.sqrt(np.mean(_drawdown**2)) * 100
return _ulcer_index
def twr(equity_curve):
eq_arr = np.array(equity_curve)
_twr = eq_arr[-1] / eq_arr[0]
return _twr
def ghpr(equity_curve):
_twr = twr(equity_curve)
_ghpr = _twr ** (1 / len(equity_curve)) - 1
return _ghpr
```
### Returns Distribution Metrics
These are path-independent and remain the same regardless of which order returns occur in
```
def sharpe_ratio(returns):
return np.mean(returns) / np.std(returns)
def sortino_ratio(returns):
losses = returns[returns <= 0]
downside_deviation = np.std(losses)
return np.mean(returns) / downside_deviation
def mean_return(returns):
return np.mean(returns)
def win_percent(returns):
wins = returns[returns > 0]
return len(wins) / len(returns)
def win_loss_ratio(returns):
wins = returns[returns > 0]
losses = returns[returns <= 0]
avg_win = np.mean(wins)
avg_loss = np.mean(np.abs(losses))
return avg_win / avg_loss
def profit_factor(returns):
wins = returns[returns > 0]
losses = returns[returns <= 0]
sum_wins = np.sum(wins)
sum_losses = np.sum(np.abs(losses))
return sum_wins / sum_losses
def cpc_index(returns):
_profit_factor = profit_factor(returns)
_win_percent = win_percent(returns)
_win_loss_ratio = win_loss_ratio(returns)
return _profit_factor * _win_percent * _win_loss_ratio
def tail_ratio(returns):
percentile_5 = np.abs(np.percentile(returns, 5))
percentile_95 = np.abs(np.percentile(returns, 95))
return percentile_95 / percentile_5
def common_sense_ratio(returns):
_common_sense_ratio = profit_factor(returns) * tail_ratio(returns)
return _common_sense_ratio
def outlier_win_ratio(returns):
wins = returns[returns > 0]
mean_win = np.mean(wins)
outlier_win = np.percentile(returns, 99)
return outlier_win / mean_win
def outlier_loss_ratio(returns):
losses = returns[returns <= 0]
mean_loss = np.mean(np.abs(losses))
outlier_loss = np.abs(np.percentile(returns, 1))
return outlier_loss / mean_loss
def ideal_f(returns, time_horizon=250, n_curves=1000, drawdown_limit=20, certainty_level=95):
"""
Calculates ideal fraction to stake on an investment with given return distribution
Args:
returns: (array-like) distribution that's representative of future returns
time_horizon: (integer) the number of returns to sample for each curve
n_curves: (integer) the number of equity curves to generate on each iteration of f
drawdown_limit: (real) user-specified value for drawdown which must not be exceeded
certainty_level: (real) the level of confidence that drawdown
limit will not be exceeded
Returns:
'f_curve': calculated drawdown and ghpr value at each value of f
'optimal_f': the ideal fraction of one's account to stake on an investment
'max_loss': the maximum loss sustained in the provided returns distribution
"""
# print('Calculating ideal f...')
start = time.time()
f_values = np.linspace(0, 0.99, 200)
max_loss = np.abs(np.min(returns))
bounded_f = f_values / max_loss
f_curve = pd.DataFrame(columns=['ghpr', 'drawdown'])
for f in bounded_f:
# Generate n_curves number of random equity curves
reordered_returns = np.random.choice(f * returns, size=(time_horizon, n_curves))
curves = equity_curve(reordered_returns)
curves_df = pd.DataFrame(curves)
# Calculate GHPR and Maximum Drawdown for each equity curve
curves_drawdown = max_drawdown(curves_df)
curves_ghpr = ghpr(curves_df)
# Calculate drawdown at our certainty level
drawdown_percentile = np.percentile(curves_drawdown, certainty_level)
# Calculate median ghpr value
ghpr_median = np.median(curves_ghpr)
if drawdown_percentile <= drawdown_limit:
_ghpr = ghpr_median
else:
_ghpr = 0
f_curve.loc[f, 'ghpr'] = _ghpr * 100
f_curve.loc[f, 'drawdown'] = drawdown_percentile
try:
optimal_f = f_curve['ghpr'].astype(float).idxmax()
max_ghpr = f_curve.loc[optimal_f, 'ghpr']
return {'f_curve':f_curve, 'optimal_f':optimal_f, 'max_loss':max_loss, 'max_ghpr':max_ghpr}
except(TypeError):
print('something went wrong')
return {'f_curve':f_curve, 'optimal_f':0, 'max_loss':0, 'max_ghpr':0}
def get_performance_metrics(returns):
metric_funcs = [
sharpe_ratio, sortino_ratio, win_percent,
win_loss_ratio, profit_factor, cpc_index,
tail_ratio, common_sense_ratio, outlier_win_ratio,
outlier_loss_ratio
]
metric_dict = {
metric_func.__name__:metric_func(returns)
for metric_func in metric_funcs
}
ideal_f_results = ideal_f(returns)
metric_dict['optimal_f'] = ideal_f_results['optimal_f']
metric_dict['ghpr'] = ideal_f_results['max_ghpr']
return pd.Series(metric_dict)
# Gather all the metrics we want to track on each run
metric_funcs = [
sharpe_ratio, sortino_ratio, win_percent,
win_loss_ratio, profit_factor, cpc_index,
tail_ratio, common_sense_ratio, outlier_win_ratio,
outlier_loss_ratio
]
metric_columns = [
metric_func.__name__ for metric_func in metric_funcs
]
metric_columns += ['optimal_f', 'ghpr']
metric_df = pd.DataFrame(columns=metric_columns)
# We'll say that our strategies are in the market 25% of the time
exposure = 0.25
for i in tqdm(range(1000)):
returns = np.random.choice(returns_list)
sample_size = max(250, int(len(returns) * exposure))
sample = np.random.choice(returns, sample_size)
metric_row = get_performance_metrics(sample)
metric_df = metric_df.append(metric_row, ignore_index=True)
metric_df = metric_df[metric_df['ghpr'] > 0]
# Plot the results
fig, axes = plt.subplots(4, 3, figsize=(9,12))
metric_df_col_num = 0
for row in range(4):
axes[row, 0].set_ylabel('ghpr')
for col in range(3):
metric_column = metric_df.columns[metric_df_col_num]
axes[row, col].scatter(metric_df[metric_column], metric_df['ghpr'])
axes[row, col].set_xlabel(metric_column)
metric_df_col_num += 1
fig.suptitle('Performance Metric Correlation', size=16)
plt.tight_layout()
fig.subplots_adjust(top=0.95)
plt.savefig('Performance Metric Correlation')
from sklearn.linear_model import LinearRegression
from sklearn.model_selection import train_test_split
from sklearn.metrics import r2_score
from sklearn.preprocessing import PolynomialFeatures
# Single feature is sharpe ratio of returns
X = metric_df['sharpe_ratio'].values.reshape(-1,1)
# Target is median GHPR
y = metric_df['ghpr']
# Split out our training and test sets
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3)
# Use polynomial features to account for non-linear curve
poly = PolynomialFeatures(2)
X_train = poly.fit_transform(X_train)
X_test = poly.fit_transform(X_test)
# Train our model
model = LinearRegression()
model.fit(X_train, y_train)
# Make some predictions
y_pred = model.predict(X_test)
# How'd we do?
r2 = r2_score(y_test, y_pred)
print('RSQ Achieved: {}%'.format(np.round(r2 * 100, 2)))
X = metric_df.drop(columns=['optimal_f', 'ghpr'])
y = metric_df['ghpr']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3)
model = LinearRegression()
model.fit(X_train, y_train)
y_pred = model.predict(X_test)
r2 = r2_score(y_test, y_pred)
print('RSQ Achieved: {}%'.format(np.round(r2 * 100, 2)))
X = metric_df.drop(columns=['optimal_f', 'ghpr'])
y = metric_df['ghpr']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3)
poly = PolynomialFeatures(2)
X_train = poly.fit_transform(X_train)
X_test = poly.fit_transform(X_test)
model = LinearRegression()
model.fit(X_train, y_train)
y_pred = model.predict(X_test)
r2 = r2_score(y_test, y_pred)
print('RSQ Achieved: {}%'.format(np.round(r2 * 100, 2)))
```
| github_jupyter |
# You Only Look Once (YOLO)
Este componente utiliza o modelo YOLO para classificação usando [Yolov4](https://pypi.org/project/yolov4/). <br>
Está é uma biblioteca que encapsula o modelo yolov4 com algumas variações, podendo utilizar o modelo completo, como também,a versão reduzida.
Este notebook apresenta:
- como usar o [SDK](https://platiagro.github.io/sdk/) para carregar datasets, salvar modelos e outros artefatos.
- como declarar parâmetros e usá-los para criar componentes reutilizáveis.
```
# Download weigths
import gdown
gdown.download('https://drive.google.com/u/0/uc?id=1GJwGiR7rizY_19c_czuLN8p31BwkhWY5', 'yolov4-tiny.weights', quiet=False)
gdown.download('https://drive.google.com/uc?id=1L-SO373Udc9tPz5yLkgti5IAXFboVhUt', 'yolov4-full.weights', quiet=False)
%%writefile Model.py
from typing import List, Iterable, Dict, Union
import numpy as np
import cv2
import tensorflow as tf
from yolov4.tf import YOLOv4
from yolo_utils import decode_yolo_bbox
import joblib
class Model:
def __init__(self):
self.loaded = False
def load(self):
# Carrega artefatos: estimador, etc
artifacts = joblib.load("/tmp/data/yolo.joblib")
self.names = artifacts["names"]
self.inference_parameters = artifacts["inference_parameters"]
# Load Model
is_tiny = self.inference_parameters['yolo_weight_type'] == 'tiny'
self.yolo = YOLOv4(tiny=is_tiny)
self.yolo.classes = "coco.names"
self.yolo.make_model()
# Download and load weigths
if is_tiny:
self.yolo.load_weights("yolov4-tiny.weights", weights_type="yolo")
else:
self.yolo.load_weights("yolov4-full.weights", weights_type="yolo")
# Health check validation
random_img = np.random.randint(0, 255, size=(100, 100, 3), dtype=np.uint8)
_ = self.yolo.predict(random_img)
self.loaded = True
def class_names(self):
return ['x_min', 'y_min', 'x_max', 'y_max', 'class', 'probability']
def predict(self, X: np.ndarray, feature_names: Iterable[str], meta: Dict = None) -> Union[np.ndarray, List, str, bytes]:
if not self.loaded:
self.load()
# Check if data is a bytes
if isinstance(X, bytes):
im_bytes = X # Get image bytes
# If not, should be a list or ndarray
else:
# Garantee is a ndarray
X = np.array(X)
# Seek for extra dimension
if len(X.shape) == 2:
im_bytes = X[0,0] # Get image bytes
else:
im_bytes = X[0] # Get image bytes
# Preprocess img bytes to img_arr
im_arr = np.frombuffer(im_bytes, dtype=np.uint8)
img = cv2.imdecode(im_arr, flags=cv2.IMREAD_COLOR)
img = cv2.cvtColor(img, cv2.COLOR_BGR2RGB)
frame = np.array(img).astype(np.uint8)
# How to interpret YOLO bbox: https://stackoverflow.com/questions/52455429/what-does-the-coordinate-output-of-yolo-algorithm-represent
predictions = self.yolo.predict(frame,
score_threshold=self.inference_parameters['score_threshold'],
iou_threshold=self.inference_parameters['iou_threshold'])
# Compile results
results = []
for i, prediction in enumerate(predictions):
result = []
# Decode yolo bbox
encoded_bbox = prediction[:4]
decoded_bbox = decode_yolo_bbox(frame, encoded_bbox)
name = self.names[int(prediction[4])]
prob = prediction[5]
# Check if prediction is null
if prob < 1e-6:
decoded_bbox = [None]*4
name = None
prob = None
result += list(decoded_bbox)
# Get class name
result.append(name)
# Get probability
result.append(prob)
# Compile result
results.append(result)
return np.array(results).astype(str)
```
| github_jupyter |
# Digit Recognition
## 1. Introduction
In this analysis, the handwritten digits are identified using support vector machines and radial basis functions.
### 1.1 Libraries
The essential libraries used here are numpy, matplotlib, and scikit-learn. For convenience, pandas and IPython.display are used for displaying tables, and tqdm is used for progress bars.
```
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
from itertools import product
from sklearn.svm import SVC
from sklearn.model_selection import cross_val_score, cross_val_predict, ShuffleSplit, KFold
from tqdm import tqdm
from IPython.display import display, Math, Latex, HTML
%matplotlib inline
np.set_printoptions(precision=4,threshold=200)
tqdm_bar_fmt='{percentage:3.0f}%|{bar}|'
```
### 1.2 Dataset
The US Postal Service Zip Code dataset is used, which contains handwritten digits zero to nine. The data has been preprocessed, whereby features of intensity and symmetry are extracted.
```
def download_data():
train_url = "http://www.amlbook.com/data/zip/features.train"
test_url = "http://www.amlbook.com/data/zip/features.test"
column_names = ['digit','intensity','symmetry']
train = pd.read_table(train_url,names=column_names,header=None,delim_whitespace=True)
test = pd.read_table(test_url,names=column_names,header=None,delim_whitespace=True)
train.digit = train.digit.astype(int)
test.digit = test.digit.astype(int)
return train,test
def process_data(train,test):
X_train = train.iloc[:,1:].values
y_train = train.iloc[:,0].values
X_test = test.iloc[:,1:].values
y_test = test.iloc[:,0].values
return X_train,y_train,X_test,y_test
train,test = download_data()
X_train,y_train,X_test,y_test = process_data(train,test)
```
## 2. Support Vector Machines for Digit Recognition
### 2.1 Polynomial Kernels
We wish to implement the following polynomial kernel for our support vector machine:
$$K\left(\mathbf{x_n,x_m}\right) = \left(1+\mathbf{x_n^Tx_m}\right)^Q$$
This is implemented in scikit-learn in the subroutine [sklearn.svm.SVC](http://scikit-learn.org/stable/modules/svm.html), where the kernel function takes the form:
$$\left(\gamma \langle x,x' \rangle + r\right)^d$$
where $d$ is specified by the keyword `degree`, and $r$ by `coef0`.
### 2.1.1 One vs Rest Classification
In the following subroutine, the data is split into "one-vs-rest", where $y=1$ corresponds to a match to the digit, and $y=0$ corresponds to all the other digits. The training step is implemented in the call to `clf.fit()`.
```
def get_misclassification_ovr(X_train,y_train,X_test,y_test,digit,
Q=2,r=1.0,C=0.01,kernel='poly',verbose=False):
clf = SVC(C=C, kernel=kernel, degree=Q, coef0 = r, gamma = 1.0,
decision_function_shape='ovr', verbose=False)
y_in = (y_train==digit).astype(int)
y_out = (y_test==digit).astype(int)
model = clf.fit(X_train,y_in) # print(model)
E_in = np.mean(y_in != clf.predict(X_train))
E_out = np.mean(y_out != clf.predict(X_test))
n_support_vectors = len(clf.support_vectors_)
if verbose is True:
print()
print("Q = {}, C = {}: Support vectors: {}".format(Q, C, n_support_vectors))
print("{} vs all: E_in = {}".format(digit,E_in))
print("{} vs all: E_out = {}".format(digit,E_out))
return E_in,E_out,n_support_vectors
```
The following code trains on the data for the cases: 0 vs all, 1 vs all, ..., 9 vs all. For each of the digits, 0 to 9, the errors $E_{in}, E_{out}$ and the number of support vectors are recorded and stored in a pandas dataframe.
```
results = pd.DataFrame()
i=0
for digit in tqdm(range(10),bar_format=tqdm_bar_fmt):
ei, eo, n = get_misclassification_ovr(X_train,y_train,X_test,y_test,digit)
df = pd.DataFrame({'digit': digit, 'E_in': ei, 'E_out': eo, 'n': n}, index=[i])
results = results.append(df)
i += 1
display(HTML(results[['digit','E_in','E_out','n']].iloc[::2].to_html(index=False)))
display(HTML(results[['digit','E_in','E_out','n']].iloc[1::2].to_html(index=False)))
from tabulate import tabulate
print(tabulate(results, headers='keys', tablefmt='simple'))
```
### 2.1.2 One vs One Classification
One vs one classification makes better use of the data, but is more computatationally expensive. The following subroutine splits the data so that $y=0$ for the first digit, and $y=1$ for the second digit. The rows of data corresponding to all other digits are removed.
```
def get_misclassification_ovo(X_train,y_train,X_test,y_test,digit1,digit2,
Q=2,r=1.0,C=0.01,kernel='poly'):
clf = SVC(C=C, kernel=kernel, degree=Q, coef0 = r, gamma = 1.0,
decision_function_shape='ovo', verbose=False)
select_in = np.logical_or(y_train==digit1,y_train==digit2)
y_in = (y_train[select_in]==digit1).astype(int)
X_in = X_train[select_in]
select_out = np.logical_or(y_test==digit1,y_test==digit2)
y_out = (y_test[select_out]==digit1).astype(int)
X_out = X_test[select_out]
model = clf.fit(X_in,y_in)
E_in = np.mean(y_in != clf.predict(X_in))
E_out = np.mean(y_out != clf.predict(X_out))
n_support_vectors = len(clf.support_vectors_)
return E_in,E_out,n_support_vectors
```
In the following code, a 1-vs-5 classifier is tested for $Q=2,5$ and $C=0.001,0.01,0.1,1$.
```
C_arr = [0.0001, 0.001, 0.01, 1]
Q_arr = [2, 5]
CQ_arr = list(product(C_arr,Q_arr))
results = pd.DataFrame()
i=0
for C, Q in tqdm(CQ_arr,bar_format=tqdm_bar_fmt):
ei, eo, n = get_misclassification_ovo(X_train,y_train,X_test,y_test,
digit1=1,digit2=5,Q=Q,r=1.0,C=C)
df = pd.DataFrame({'C': C, 'Q': Q, 'E_in': ei, 'E_out': eo, 'n': n}, index=[i])
results = results.append(df)
i += 1
display(HTML(results[['C','Q','E_in','E_out','n']].to_html(index=False)))
```
### 2.1.3 Polynomial Kernel with Cross-Validation
For k-fold cross-validation, the subroutine [`sklearn.model_selection.KFold`](http://scikit-learn.org/stable/modules/generated/sklearn.model_selection.KFold.html#sklearn.model_selection.KFold)`()` is employed to split the data into folds. Random shuffling is enabled with the `shuffle=True` option.
```
def select_digits_ovo(X,y,digit1,digit2):
subset = np.logical_or(y==digit1,y==digit2)
X_in = X[subset].copy()
y_in = (y[subset]==digit1).astype(int).copy()
return X_in,y_in
def get_misclassification_ovo_cv(X_in,y_in,Q=2,r=1.0,C=0.01,kernel='poly'):
kf = KFold(n_splits=10, shuffle=True)
kf.get_n_splits(X_in)
E_cv = []
for train_index, test_index in kf.split(X_in):
X_trn, X_tst = X_in[train_index], X_in[test_index]
y_trn, y_tst = y_in[train_index], y_in[test_index]
clf = SVC(C=C, kernel=kernel, degree=Q, coef0 = r, gamma = 1.0,
decision_function_shape='ovo', verbose=False)
model = clf.fit(X_trn,y_trn)
E_cv.append(np.mean(y_tst != clf.predict(X_tst)))
return E_cv
```
In this example, our parameters are $Q=2, C \in \{0.0001,0.001,0.01,0.1,1\}$ and we are considering the 1-vs-5 classifier.
```
C_arr = [0.0001, 0.001, 0.01, 0.1, 1]; d1 = 1; d2 = 5
Q=2
X_in,y_in = select_digits_ovo(X_train,y_train,d1,d2)
```
Due to the effect of random shuffling, 100 runs are carried out, and the results tabulated.
```
E_cv_arr = []
count_arr = []
for n in tqdm(range(100),bar_format=tqdm_bar_fmt):
counts = np.zeros(len(C_arr),int)
kf = KFold(n_splits=10, shuffle=True)
kf.get_n_splits(X_in)
for train_index, test_index in kf.split(X_in):
X_trn, X_tst = X_in[train_index], X_in[test_index]
y_trn, y_tst = y_in[train_index], y_in[test_index]
E_cv = []
for C in C_arr:
clf = SVC(C=C, kernel='poly', degree=Q, coef0=1.0, gamma=1.0,
decision_function_shape='ovo', verbose=False)
model = clf.fit(X_trn,y_trn)
E_cv.append(np.mean(y_tst != clf.predict(X_tst)))
counts[np.argmin(E_cv)] += 1
E_cv_arr.append(np.array(E_cv))
count_arr.append(counts)
```
The number of times each particular value of $C$ is picked (for having the smallest $E_{cv}$) is calculated, as well as the mean cross validation error. The data is tabulated as follows:
```
df = pd.DataFrame({'C': C_arr})
df['count'] = np.sum(np.array(count_arr),axis=0)
df['E_cv'] = np.mean(np.array(E_cv_arr),axis=0)
display(HTML(df.to_html(index=False)))
fig = plt.figure(figsize=(10,5))
ax1 = fig.add_subplot(121)
_ = df.plot(y=['count'],ax=ax1,marker='x',color='red',markersize=10,grid=True)
ax2 = fig.add_subplot(122)
_ = df.plot(y=['E_cv'],ax=ax2,marker='o',color='green',markersize=7,grid=True)
```
### 2.2 Scaling Law for Support Vector Machines
This is based on [scikit-learn example code](http://scikit-learn.org/stable/tutorial/basic/tutorial.html), but modified to demonstrate how the training time scales with the size of the data. The dataset in this example is the original MNIST data.
```
from sklearn.datasets import fetch_mldata
from sklearn import svm
import timeit
digits = fetch_mldata('MNIST original') # stores data in ~/scikit_learn_data by default
n_max = len(digits.target)
selection = np.random.permutation(n_max)
n_arr = [500, 1000, 1500, 2000, 5000, 10000]
t_arr = []
for n in n_arr:
sel = selection[:n]
X = digits.data[sel]
y = digits.target[sel]
clf = svm.SVC(gamma=0.001, C=100.)
t0 = timeit.default_timer()
clf.fit(X,y)
t_arr.append(timeit.default_timer() - t0)
print("n = {}, time = {}".format(n, t_arr[-1]))
plt.plot(np.array(n_arr),np.array(t_arr),'bx-')
plt.xlabel('no. of samples')
plt.ylabel('time (s)')
plt.grid()
```
## 3. Radial Basis Functions
### 3.1 Background
The hypothesis is given by:
$$h\left(\mathbf{x}\right) = \sum\limits_{k=1}^K w_k \exp\left(-\gamma \Vert \mathbf{x} - \mu_k \Vert^2\right)$$
This is implemented in the subroutine [`sklearn.svm.SVC`](http://scikit-learn.org/stable/modules/svm.html)`(..., kernel='rbf', ...)` as shown in the code below.
```
def get_misclassification_rbf(X_train,y_train,X_test,y_test,C,digit1=1,digit2=5):
clf = SVC(C=C, kernel='rbf', gamma = 1.0,
decision_function_shape='ovo', verbose=False)
select_in = np.logical_or(y_train==digit1,y_train==digit2)
y_in = (y_train[select_in]==digit1).astype(float)
X_in = X_train[select_in]
select_out = np.logical_or(y_test==digit1,y_test==digit2)
y_out = (y_test[select_out]==digit1).astype(float)
X_out = X_test[select_out]
model = clf.fit(X_in,y_in)
E_in = np.mean(y_in != clf.predict(X_in))
E_out = np.mean(y_out != clf.predict(X_out))
n_support_vectors = len(clf.support_vectors_)
return E_in,E_out,n_support_vectors
```
For $C \in \{0.01, 1, 100, 10^4, 10^6\}$, the in-sample and out-of-sample errors, as well as the number of support vectors are tabulated as follows:
```
C_arr = [0.01, 1., 100., 1e4, 1e6]
results = pd.DataFrame()
i=0
for C in tqdm(C_arr,bar_format=tqdm_bar_fmt):
ei, eo, n = get_misclassification_rbf(X_train,y_train,X_test,y_test,
C,digit1=1,digit2=5)
df = pd.DataFrame({'C': C, 'E_in': ei, 'E_out': eo, 'n': n}, index=[i])
results = results.append(df)
i += 1
display(HTML(results.to_html(index=False)))
```
The table is also plotted graphically below:
```
def plot_Ein_Eout(df):
fig = plt.figure(figsize=(12,5))
ax1 = fig.add_subplot(121)
df.plot(ax=ax1, x='C', y='E_in', kind='line', marker='o', markersize=7, logx=True)
df.plot(ax=ax1, x='C', y='E_out', kind='line', marker='x', markersize=7, logx=True)
ax1.legend(loc='best',frameon=False)
ax1.grid(True)
ax1.set_title('In-Sample vs Out-of-Sample Errors')
ax2 = fig.add_subplot(122)
df.plot(ax=ax2, x='C', y='n', kind='line', marker='+', markersize=10, logx=True)
ax2.legend(loc='best',frameon=False)
ax2.grid(True)
ax2.set_title('Number of Support Vectors')
plot_Ein_Eout(results)
```
| github_jupyter |
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
<a href="https://colab.research.google.com/github/lmoroney/dlaicourse/blob/master/TensorFlow%20In%20Practice/Course%204%20-%20S%2BP/S%2BP%20Week%204%20Lesson%201.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
try:
# %tensorflow_version only exists in Colab.
%tensorflow_version 2.x
except Exception:
pass
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
print(tf.__version__)
def plot_series(time, series, format="-", start=0, end=None):
plt.plot(time[start:end], series[start:end], format)
plt.xlabel("Time")
plt.ylabel("Value")
plt.grid(True)
def trend(time, slope=0):
return slope * time
def seasonal_pattern(season_time):
"""Just an arbitrary pattern, you can change it if you wish"""
return np.where(season_time < 0.4,
np.cos(season_time * 2 * np.pi),
1 / np.exp(3 * season_time))
def seasonality(time, period, amplitude=1, phase=0):
"""Repeats the same pattern at each period"""
season_time = ((time + phase) % period) / period
return amplitude * seasonal_pattern(season_time)
def noise(time, noise_level=1, seed=None):
rnd = np.random.RandomState(seed)
return rnd.randn(len(time)) * noise_level
time = np.arange(4 * 365 + 1, dtype="float32")
baseline = 10
series = trend(time, 0.1)
baseline = 10
amplitude = 40
slope = 0.05
noise_level = 5
# Create the series
series = baseline + trend(time, slope) + seasonality(time, period=365, amplitude=amplitude)
# Update with noise
series += noise(time, noise_level, seed=42)
split_time = 1000
time_train = time[:split_time]
x_train = series[:split_time]
time_valid = time[split_time:]
x_valid = series[split_time:]
window_size = 20
batch_size = 32
shuffle_buffer_size = 1000
def windowed_dataset(series, window_size, batch_size, shuffle_buffer):
series = tf.expand_dims(series, axis=-1)
ds = tf.data.Dataset.from_tensor_slices(series)
ds = ds.window(window_size + 1, shift=1, drop_remainder=True)
ds = ds.flat_map(lambda w: w.batch(window_size + 1))
ds = ds.shuffle(shuffle_buffer)
ds = ds.map(lambda w: (w[:-1], w[1:]))
return ds.batch(batch_size).prefetch(1)
def model_forecast(model, series, window_size):
ds = tf.data.Dataset.from_tensor_slices(series)
ds = ds.window(window_size, shift=1, drop_remainder=True)
ds = ds.flat_map(lambda w: w.batch(window_size))
ds = ds.batch(32).prefetch(1)
forecast = model.predict(ds)
return forecast
tf.keras.backend.clear_session()
tf.random.set_seed(51)
np.random.seed(51)
window_size = 30
train_set = windowed_dataset(x_train, window_size, batch_size=128, shuffle_buffer=shuffle_buffer_size)
model = tf.keras.models.Sequential([
tf.keras.layers.Conv1D(filters=32, kernel_size=5,
strides=1, padding="causal",
activation="relu",
input_shape=[None, 1]),
tf.keras.layers.Bidirectional(tf.keras.layers.LSTM(32, return_sequences=True)),
tf.keras.layers.Bidirectional(tf.keras.layers.LSTM(32, return_sequences=True)),
tf.keras.layers.Dense(1),
tf.keras.layers.Lambda(lambda x: x * 200)
])
lr_schedule = tf.keras.callbacks.LearningRateScheduler(
lambda epoch: 1e-8 * 10**(epoch / 20))
optimizer = tf.keras.optimizers.SGD(lr=1e-8, momentum=0.9)
model.compile(loss=tf.keras.losses.Huber(),
optimizer=optimizer,
metrics=["mae"])
history = model.fit(train_set, epochs=100, callbacks=[lr_schedule])
plt.semilogx(history.history["lr"], history.history["loss"])
plt.axis([1e-8, 1e-4, 0, 30])
tf.keras.backend.clear_session()
tf.random.set_seed(51)
np.random.seed(51)
#batch_size = 16
dataset = windowed_dataset(x_train, window_size, batch_size, shuffle_buffer_size)
model = tf.keras.models.Sequential([
tf.keras.layers.Conv1D(filters=32, kernel_size=3,
strides=1, padding="causal",
activation="relu",
input_shape=[None, 1]),
tf.keras.layers.LSTM(32, return_sequences=True),
tf.keras.layers.LSTM(32, return_sequences=True),
tf.keras.layers.Dense(1),
tf.keras.layers.Lambda(lambda x: x * 200)
])
optimizer = tf.keras.optimizers.SGD(lr=1e-5, momentum=0.9)
model.compile(loss=tf.keras.losses.Huber(),
optimizer=optimizer,
metrics=["mae"])
history = model.fit(dataset,epochs=500)
rnn_forecast = model_forecast(model, series[..., np.newaxis], window_size)
rnn_forecast = rnn_forecast[split_time - window_size:-1, -1, 0]
plt.figure(figsize=(10, 6))
plot_series(time_valid, x_valid)
plot_series(time_valid, rnn_forecast)
tf.keras.metrics.mean_absolute_error(x_valid, rnn_forecast).numpy()
import matplotlib.image as mpimg
import matplotlib.pyplot as plt
#-----------------------------------------------------------
# Retrieve a list of list results on training and test data
# sets for each training epoch
#-----------------------------------------------------------
mae=history.history['mae']
loss=history.history['loss']
epochs=range(len(loss)) # Get number of epochs
#------------------------------------------------
# Plot MAE and Loss
#------------------------------------------------
plt.plot(epochs, mae, 'r')
plt.plot(epochs, loss, 'b')
plt.title('MAE and Loss')
plt.xlabel("Epochs")
plt.ylabel("Accuracy")
plt.legend(["MAE", "Loss"])
plt.figure()
epochs_zoom = epochs[200:]
mae_zoom = mae[200:]
loss_zoom = loss[200:]
#------------------------------------------------
# Plot Zoomed MAE and Loss
#------------------------------------------------
plt.plot(epochs_zoom, mae_zoom, 'r')
plt.plot(epochs_zoom, loss_zoom, 'b')
plt.title('MAE and Loss')
plt.xlabel("Epochs")
plt.ylabel("Accuracy")
plt.legend(["MAE", "Loss"])
plt.figure()
```
| github_jupyter |
## Practice 02: Dealing with texts using CNN
Today we're gonna apply the newly learned tools for the task of predicting job salary.
<img src="https://storage.googleapis.com/kaggle-competitions/kaggle/3342/logos/front_page.png" width=400px>
Based on YSDA [materials](https://github.com/yandexdataschool/nlp_course/blob/master/week02_classification/seminar.ipynb). _Special thanks to [Oleg Vasilev](https://github.com/Omrigan/) for the core assignment idea._
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
%matplotlib inline
```
### About the challenge
For starters, let's download and unpack the data from [here](https://www.dropbox.com/s/5msc5ix7ndyba10/Train_rev1.csv.tar.gz?dl=0).
You can also get it from [yadisk url](https://yadi.sk/d/vVEOWPFY3NruT7) the competition [page](https://www.kaggle.com/c/job-salary-prediction/data) (pick `Train_rev1.*`).
```
# Do this only once
!curl -L "https://www.dropbox.com/s/5msc5ix7ndyba10/Train_rev1.csv.tar.gz?dl=1" -o Train_rev1.csv.tar.gz
!tar -xvzf ./Train_rev1.csv.tar.gz
data = pd.read_csv("./Train_rev1.csv", index_col=None)
data.shape
```
One problem with salary prediction is that it's oddly distributed: there are many people who are paid standard salaries and a few that get tons o money. The distribution is fat-tailed on the right side, which is inconvenient for MSE minimization.
There are several techniques to combat this: using a different loss function, predicting log-target instead of raw target or even replacing targets with their percentiles among all salaries in the training set. We gonna use logarithm for now.
_You can read more [in the official description](https://www.kaggle.com/c/job-salary-prediction#description)._
```
data['Log1pSalary'] = np.log1p(data['SalaryNormalized']).astype('float32')
plt.figure(figsize=[8, 4])
plt.subplot(1, 2, 1)
plt.hist(data["SalaryNormalized"], bins=20);
plt.subplot(1, 2, 2)
plt.hist(data['Log1pSalary'], bins=20);
```
Our task is to predict one number, __Log1pSalary__.
To do so, our model can access a number of features:
* Free text: __`Title`__ and __`FullDescription`__
* Categorical: __`Category`__, __`Company`__, __`LocationNormalized`__, __`ContractType`__, and __`ContractTime`__.
```
text_columns = ["Title", "FullDescription"]
categorical_columns = ["Category", "Company", "LocationNormalized", "ContractType", "ContractTime"]
target_column = "Log1pSalary"
data[categorical_columns] = data[categorical_columns].fillna('NaN') # cast missing values to string "NaN"
data.sample(3)
```
### Preprocessing text data
Just like last week, applying NLP to a problem begins from tokenization: splitting raw text into sequences of tokens (words, punctuation, etc).
__Your task__ is to lowercase and tokenize all texts under `Title` and `FullDescription` columns. Store the tokenized data as a __space-separated__ string of tokens for performance reasons.
It's okay to use nltk tokenizers. Assertions were designed for WordPunctTokenizer, slight deviations are okay.
```
print("Raw text:")
print(data["FullDescription"][2::100000])
import nltk
tokenizer = nltk.tokenize.WordPunctTokenizer()
# see task above
def normalize(text):
text = str(text).lower()
return ' '.join(tokenizer.tokenize(text))
data[text_columns] = data[text_columns].applymap(normalize)
```
Now we can assume that our text is a space-separated list of tokens:
```
print("Tokenized:")
print(data["FullDescription"][2::100000])
assert data["FullDescription"][2][:50] == 'mathematical modeller / simulation analyst / opera'
assert data["Title"][54321] == 'international digital account manager ( german )'
```
Not all words are equally useful. Some of them are typos or rare words that are only present a few times.
Let's count how many times is each word present in the data so that we can build a "white list" of known words.
```
# Count how many times does each token occur in both "Title" and "FullDescription" in total
# build a dictionary { token -> it's count }
from collections import Counter
from tqdm import tqdm as tqdm
token_counts = Counter() # <YOUR CODE HERE>
# hint: you may or may not want to use collections.Counter
for row in tqdm(data[text_columns].values.flatten()):
token_counts.update(row.split(' '))
token_counts.most_common(1)[0][1]
print("Total unique tokens :", len(token_counts))
print('\n'.join(map(str, token_counts.most_common(n=5))))
print('...')
print('\n'.join(map(str, token_counts.most_common()[-3:])))
assert token_counts.most_common(1)[0][1] in range(2600000, 2700000)
assert len(token_counts) in range(200000, 210000)
print('Correct!')
# Let's see how many words are there for each count
plt.hist(list(token_counts.values()), range=[0, 10**4], bins=50, log=True)
plt.xlabel("Word counts");
```
Now filter tokens a list of all tokens that occur at least 10 times.
```
min_count = 10
# tokens from token_counts keys that had at least min_count occurrences throughout the dataset
tokens = [token for token, count in token_counts.items() if count >= min_count]
# Add a special tokens for unknown and empty words
UNK, PAD = "UNK", "PAD"
tokens = [UNK, PAD] + sorted(tokens)
print("Vocabulary size:", len(tokens))
assert type(tokens) == list
assert len(tokens) in range(32000, 35000)
assert 'me' in tokens
assert UNK in tokens
print("Correct!")
```
Build an inverse token index: a dictionary from token(string) to it's index in `tokens` (int)
```
# You have already done that ;)
token_to_id = {token: idx for idx, token in enumerate(tokens) }
assert isinstance(token_to_id, dict)
assert len(token_to_id) == len(tokens)
for tok in tokens:
assert tokens[token_to_id[tok]] == tok
print("Correct!")
```
And finally, let's use the vocabulary you've built to map text lines into neural network-digestible matrices.
```
UNK_IX, PAD_IX = map(token_to_id.get, [UNK, PAD])
def as_matrix(sequences, max_len=None):
""" Convert a list of tokens into a matrix with padding """
if isinstance(sequences[0], str):
sequences = list(map(str.split, sequences))
max_len = min(max(map(len, sequences)), max_len or float('inf'))
matrix = np.full((len(sequences), max_len), np.int32(PAD_IX)) # Return a new array of given shape and type, filled with fill_value.
for i,seq in enumerate(sequences):
row_ix = [token_to_id.get(word, UNK_IX) for word in seq[:max_len]]
matrix[i, :len(row_ix)] = row_ix
return matrix
print("Lines:")
print('\n'.join(data["Title"][::100000].values), end='\n\n')
print("Matrix:")
print(as_matrix(data["Title"][::100000]))
```
Now let's encode the categirical data we have.
As usual, we shall use one-hot encoding for simplicity. Kudos if you implement more advanced encodings: tf-idf, pseudo-time-series, etc.
```
!pip install feature_engine
from sklearn.feature_extraction import DictVectorizer
from sklearn.feature_extraction.text import TfidfVectorizer
from feature_engine import categorical_encoders as ce
# we only consider top-1k most frequent companies to minimize memory usage
top_companies, top_counts = zip(*Counter(data['Company']).most_common(1000))
recognized_companies = set(top_companies)
data["Company"] = data["Company"].apply(lambda comp: comp if comp in recognized_companies else "Other")
# encoder = ce.CountFrequencyCategoricalEncoder(encoding_method='frequency',
# variables=categorical_columns)
# encoder.fit(data[categorical_columns])
# data[categorical_columns] = encoder.transform(data[categorical_columns])
# categorical_vectorizer = TfidfVectorizer(dtype=np.float32)
# categorical_vectorizer.fit(data[categorical_columns])
categorical_vectorizer = DictVectorizer(dtype=np.float32, sparse=False)
categorical_vectorizer.fit(data[categorical_columns].apply(dict, axis=1))
# np.sum(categorical_vectorizer.transform(data[categorical_columns][:1].apply(dict, axis=1))[0])
```
### The deep learning part
Once we've learned to tokenize the data, let's design a machine learning experiment.
As before, we won't focus too much on validation, opting for a simple train-test split.
__To be completely rigorous,__ we've comitted a small crime here: we used the whole data for tokenization and vocabulary building. A more strict way would be to do that part on training set only. You may want to do that and measure the magnitude of changes.
```
from sklearn.model_selection import train_test_split
data_train, data_val = train_test_split(data, test_size=0.2, random_state=42)
data_train.index = range(len(data_train))
data_val.index = range(len(data_val))
print("Train size = ", len(data_train))
print("Validation size = ", len(data_val))
def make_batch(data, max_len=None, word_dropout=0):
"""
Creates a keras-friendly dict from the batch data.
:param word_dropout: replaces token index with UNK_IX with this probability
:returns: a dict with {'title' : int64[batch, title_max_len]
"""
batch = {}
batch["Title"] = as_matrix(data["Title"].values, max_len)
batch["FullDescription"] = as_matrix(data["FullDescription"].values, max_len)
batch['Categorical'] = categorical_vectorizer.transform(data[categorical_columns].apply(dict, axis=1))
if word_dropout != 0:
batch["FullDescription"] = apply_word_dropout(batch["FullDescription"], 1. - word_dropout)
if target_column in data.columns:
batch[target_column] = data[target_column].values
return batch
def apply_word_dropout(matrix, keep_prop, replace_with=UNK_IX, pad_ix=PAD_IX,):
dropout_mask = np.random.choice(2, np.shape(matrix), p=[keep_prop, 1 - keep_prop])
dropout_mask &= matrix != pad_ix
return np.choose(dropout_mask, [matrix, np.full_like(matrix, replace_with)])
a = make_batch(data_train[:3], max_len=10)
a
```
#### Architecture
Our main model consists of three branches:
* Title encoder
* Description encoder
* Categorical features encoder
We will then feed all 3 branches into one common network that predicts salary.
<img src="https://github.com/yandexdataschool/nlp_course/raw/master/resources/w2_conv_arch.png" width=600px>
This clearly doesn't fit into PyTorch __Sequential__ interface. To build such a network, one will have to use [__PyTorch nn.Module API__](https://pytorch.org/docs/stable/nn.html#torch.nn.Module).
But to start with let's build the simple model using only the part of the data. Let's create the baseline solution using only the description part (so it should definetely fit into the Sequential model).
```
import torch
from torch import nn
import torch.nn.functional as F
# You will need these to make it simple
class Flatten(nn.Module):
def forward(self, input):
return input.view(input.size(0), -1)
class Reorder(nn.Module):
def forward(self, input):
return input.permute((0, 2, 1))
```
To generate minibatches we will use simple pyton generator.
```
def iterate_minibatches(data, batch_size=256, shuffle=True, cycle=False, **kwargs):
""" iterates minibatches of data in random order """
while True:
indices = np.arange(len(data))
if shuffle:
indices = np.random.permutation(indices)
for start in range(0, len(indices), batch_size):
batch = make_batch(data.iloc[indices[start : start + batch_size]], **kwargs)
target = batch.pop(target_column)
yield batch, target
if not cycle: break
iterator = iterate_minibatches(data_train, 3)
batch, target = next(iterator)
batch, target
# Here is some startup code:
n_tokens = len(tokens)
n_cat_features = len(categorical_vectorizer.vocabulary_)
hid_size = 64
n_maximums = 2
simple_model = nn.Sequential()
simple_model.add_module('emb', nn.Embedding(num_embeddings=n_tokens, embedding_dim=hid_size))
simple_model.add_module('reorder', Reorder())
# <YOUR CODE HERE>
simple_model.add_module('conv1', nn.Conv1d(
in_channels=hid_size,
out_channels=2 * hid_size,
kernel_size=3
))
simple_model.add_module('relu1', nn.ReLU())
simple_model.add_module('conv2', nn.Conv1d(
in_channels=2 * hid_size,
out_channels=2 * hid_size,
kernel_size=3
))
simple_model.add_module('relu2', nn.ReLU())
simple_model.add_module('bn1', nn.BatchNorm1d(hid_size*2))
simple_model.add_module('adaptive_pool', nn.AdaptiveMaxPool1d(n_maximums))
simple_model.add_module('flatten', nn.Flatten())
simple_model.add_module('linear_out', nn.Linear(hid_size*2 * n_maximums, 1))
```
__Remember!__ We are working with regression problem and predicting only one number.
```
# Try this to check your model. `torch.long` tensors are required for nn.Embedding layers.
simple_model(torch.tensor(batch['FullDescription'], dtype=torch.long))
device = torch.device('cuda:0') if torch.cuda.is_available() else torch.device('cpu')
# device = torch.device('cpu')
```
And now simple training pipeline:
```
from IPython.display import clear_output
from random import sample
epochs = 1
model = simple_model
opt = torch.optim.Adam(model.parameters())
loss_func = nn.MSELoss() # <YOUR CODE HERE>
model.to(device)
history = []
model.train()
for epoch_num in range(epochs):
for idx, (batch, target) in enumerate(iterate_minibatches(data_train)):
# Preprocessing the batch data and target
batch = torch.tensor(batch['FullDescription'], dtype=torch.long).to(device)
target = torch.tensor(target).to(device)
predictions = model(batch)
predictions = predictions.view(predictions.size(0))
loss = loss_func(predictions, target) # <YOUR CODE HERE>
# train with backprop
# <YOUR CODE HERE>
loss.backward()
opt.step()
opt.zero_grad()
history.append(loss.item())
if (idx+1)%10==0:
clear_output(True)
plt.plot(history,label='loss')
plt.yscale('log')
plt.legend()
plt.show()
```
To evaluate the model it can be switched to `eval` state.
```
simple_model.eval()
```
Let's check the model quality.
```
from tqdm import tqdm, tqdm_notebook
def print_metrics(model, data, batch_size=256, name="", **kw):
squared_error = abs_error = num_samples = 0.0
for batch_x, batch_y in tqdm(iterate_minibatches(data, batch_size=batch_size, shuffle=False, **kw)):
batch = torch.tensor(batch_x['FullDescription'], dtype=torch.long).to(device)
batch_pred = model(batch)[:, 0].detach().cpu().numpy()
squared_error += np.sum(np.square(batch_pred - batch_y))
abs_error += np.sum(np.abs(batch_pred - batch_y))
num_samples += len(batch_y)
print("%s results:" % (name or ""))
print("Mean square error: %.5f" % (squared_error / num_samples))
print("Mean absolute error: %.5f" % (abs_error / num_samples))
return squared_error, abs_error
print_metrics(simple_model, data_train, name='Train')
print_metrics(simple_model, data_val, name='Val');
```
### Bonus area: three-headed network.
Now you can try to implement the network we've discussed above. Use [__PyTorch nn.Module API__](https://pytorch.org/docs/stable/nn.html#torch.nn.Module).
```
len(categorical_vectorizer.vocabulary_)
class ThreeInputsNet(nn.Module):
def __init__(self, n_tokens=len(tokens), n_cat_features=len(categorical_vectorizer.vocabulary_), hid_size=64):
super(ThreeInputsNet, self).__init__()
n_max = 4 # hid_size // 4 if hid_size // 4 > 4 else 2
self.title_emb = nn.Embedding(num_embeddings=n_tokens, embedding_dim=hid_size)
# <YOUR CODE HERE>
self.conv1_title = nn.Conv1d(in_channels=hid_size, out_channels=hid_size*2, kernel_size=2, padding=1)
self.conv2_title = nn.Conv1d(in_channels=hid_size*2, out_channels=hid_size*2, kernel_size=2, padding=1)
self.bn1_title = nn.BatchNorm1d(hid_size*2)
self.adaptive_pool_title = nn.AdaptiveMaxPool1d(n_max)
# self.linear_title = nn.Linear(2*n_max*hid_size)
self.full_emb = nn.Embedding(num_embeddings=n_tokens, embedding_dim=hid_size)
# <YOUR CODE HERE>
self.conv1_full= nn.Conv1d(in_channels=hid_size, out_channels=hid_size*2, kernel_size=3)
self.conv2_full = nn.Conv1d(in_channels=hid_size*2, out_channels=hid_size*2, kernel_size=3)
self.bn1_full = nn.BatchNorm1d(hid_size*2)
self.adaptive_pool_full = nn.AdaptiveMaxPool1d(n_max)
# self.linear_full = nn.Linear(, 1)
self.category_out = nn.Linear(n_cat_features, n_cat_features * 2) # <YOUR CODE HERE>
self.final_linear_1 = nn.Linear(n_cat_features*2 + 2*n_max*hid_size * 2, 64)
self.final_linear_2 = nn.Linear(64, 1)
self.ReLU = nn.ReLU()
self.dropout = nn.Dropout()
def forward(self, whole_input):
input1, input2, input3 = whole_input
# <YOUR CODE HERE>
title_beg = self.title_emb(input1).permute((0, 2, 1))
x = self.ReLU(self.conv1_title(title_beg))
x = self.ReLU(self.conv2_title(x))
x = self.bn1_title(x)
title = self.adaptive_pool_title(x)
# <YOUR CODE HERE>
full_beg = self.full_emb(input2).permute((0, 2, 1))
x = self.ReLU(self.conv1_full(full_beg))
x = self.ReLU(self.conv2_full(x))
x = self.bn1_full(x)
full = self.adaptive_pool_full(x)
category = self.category_out(input3)
concatenated = torch.cat(
[
title.view(title.size(0), -1),
full.view(full.size(0), -1),
category.view(category.size(0), -1)
],
dim=1)
# <YOUR CODE HERE>
x = self.ReLU(self.final_linear_1(concatenated))
# x = self.dropout(x)
out = self.final_linear_2(x)
return out
iterator = iterate_minibatches(data_train, 3)
batch, target = next(iterator)
device = torch.device('cuda:0') if torch.cuda.is_available() else torch.device('cpu')
# device = torch.device('cpu')
model = ThreeInputsNet()
model.to(device)
model([torch.tensor(batch['Title'], dtype=torch.long, device=device),
torch.tensor(batch['FullDescription'], dtype=torch.long, device=device),
torch.tensor(batch['Categorical'], dtype=torch.float, device=device)])
opt = torch.optim.Adam(model.parameters())
loss_func = nn.MSELoss() # <YOUR CODE HERE>
epochs = 1
history = []
model.train()
for epoch_num in range(epochs):
for idx, (batch, target) in enumerate(iterate_minibatches(data_train)):
# Preprocessing the batch data and target
batch = [torch.tensor(batch['Title'], dtype=torch.long, device=device),
torch.tensor(batch['FullDescription'], dtype=torch.long, device=device),
torch.tensor(batch['Categorical'], dtype=torch.float, device=device)]
target = torch.tensor(target).to(device)
predictions = model(batch)
predictions = predictions.view(predictions.size(0))
loss = loss_func(predictions, target) # <YOUR CODE HERE>
# train with backprop
# <YOUR CODE HERE>
loss.backward()
opt.step()
opt.zero_grad()
history.append(loss.item())
if (idx+1)%10==0:
clear_output(True)
plt.plot(history, label='loss')
plt.yscale('log')
plt.legend()
plt.show()
def print_metrics(model, data, batch_size=256, name="", **kw):
squared_error = abs_error = num_samples = 0.0
for batch_x, batch_y in tqdm(iterate_minibatches(data, batch_size=batch_size, shuffle=False, **kw)):
batch = [torch.tensor(batch_x['Title'], dtype=torch.long, device=device),
torch.tensor(batch_x['FullDescription'], dtype=torch.long, device=device),
torch.tensor(batch_x['Categorical'], dtype=torch.float, device=device)]
batch_pred = model(batch)[:, 0].detach().cpu().numpy()
squared_error += np.sum(np.square(batch_pred - batch_y))
abs_error += np.sum(np.abs(batch_pred - batch_y))
num_samples += len(batch_y)
print("%s results:" % (name or ""))
print("Mean square error: %.5f" % (squared_error / num_samples))
print("Mean absolute error: %.5f" % (abs_error / num_samples))
return squared_error, abs_error
model.eval()
print_metrics(model, data_train, name='Train')
print_metrics(model, data_val, name='Val');
```
**JUST FOR COMPARING!**
Val results **Simple Model**:<br>
+ Mean square error: 0.32914<br>
+ Mean absolute error: 0.44004<br>
Val results **ThreeInputModel**:<br>
+ Mean square error: 0.11429<br>
+ Mean absolute error: 0.25679<br>
### Bonus area 2: comparing RNN to CNN
Try implementing simple RNN (or LSTM) and applying it to this task. Compare the quality/performance of these networks.
*Hint: try to build networks with ~same number of paremeters.*
```
class ThreeInputsNetLSTM(nn.Module):
def __init__(self, n_tokens=len(tokens), n_cat_features=len(categorical_vectorizer.vocabulary_), hid_size=64, num_layers=2):
super(ThreeInputsNetLSTM, self).__init__()
self.n_max = 4
self.num_layers = num_layers
self.n_tokens = n_tokens
self.n_cat_features = n_cat_features
self.hid_size = hid_size
self.title_emb = nn.Embedding(num_embeddings=self.n_tokens, embedding_dim=self.hid_size)
# <YOUR CODE HERE>
self.conv1_title = nn.Conv1d(in_channels=self.hid_size, out_channels=self.hid_size*2, kernel_size=2, padding=1)
self.conv2_title = nn.Conv1d(in_channels=self.hid_size*2, out_channels=self.hid_size*2, kernel_size=2, padding=1)
self.bn1_title = nn.BatchNorm1d(self.hid_size*2)
self.adaptive_pool_title = nn.AdaptiveMaxPool1d(self.n_max)
# self.linear_title = nn.Linear(2*n_max*hid_size)
self.full_emb = nn.Embedding(num_embeddings=self.n_tokens, embedding_dim=self.hid_size)
# <YOUR CODE HERE>
self.conv1_full= nn.Conv1d(in_channels=self.hid_size, out_channels=self.hid_size*2, kernel_size=3)
self.conv2_full = nn.Conv1d(in_channels=self.hid_size*2, out_channels=self.hid_size*2, kernel_size=3)
self.bn1_full = nn.BatchNorm1d(self.hid_size*2)
self.adaptive_pool_full = nn.AdaptiveMaxPool1d(self.n_max)
self.lstm1_full = nn.LSTM(self.hid_size, self.hid_size, self.num_layers, batch_first=True, dropout=0.5) #bidirectional=True)
self.linear1_full = nn.Linear(self.hid_size, self.hid_size*2)
# self.linear_full = nn.Linear(, 1)
self.category_out = nn.Linear(self.n_cat_features, self.n_cat_features * 2) # <YOUR CODE HERE>
self.final_linear_1 = nn.Linear(self.n_cat_features*2 + 2*self.n_max*self.hid_size*2 + self.hid_size*2, 64)
self.final_linear_2 = nn.Linear(64, 1)
self.ReLU = nn.ReLU()
self.dropout = nn.Dropout()
def forward(self, whole_input):
input1, input2, input3 = whole_input
# <YOUR CODE HERE>
title_beg = self.title_emb(input1).permute((0, 2, 1))
x = self.ReLU(self.conv1_title(title_beg))
x = self.ReLU(self.conv2_title(x))
x = self.bn1_title(x)
title = self.adaptive_pool_title(x)
# <YOUR CODE HERE>
full_beg = self.full_emb(input2).permute((0, 2, 1))
x = self.ReLU(self.conv1_full(full_beg))
x = self.ReLU(self.conv2_full(x))
x = self.bn1_full(x)
full = self.adaptive_pool_full(x)
hidden, cell = self.init_hidden(x.shape[0])
lstm1, (hidden, cell) = self.lstm1_full(full_beg.permute((0, 2, 1)), (hidden, cell))
# lstm1 = self.linear1_full(torch.cat((hidden[-2,:,:], hidden[-1,:,:]), dim = 1))
lstm1 = self.linear1_full(hidden[-1,:,:])
category = self.category_out(input3)
concatenated = torch.cat(
[
title.view(title.size(0), -1),
lstm1.view(lstm1.size(0), -1),
full.view(full.size(0), -1),
category.view(category.size(0), -1)
],
dim=1)
# <YOUR CODE HERE>
# print(concatenated.shape)
x = self.ReLU(self.final_linear_1(concatenated))
# x = self.dropout(x)
out = self.final_linear_2(x)
return out
def init_hidden(self, batch_size):
hidden = torch.randn(self.num_layers, batch_size, self.hid_size).to(device)
cell = torch.randn(self.num_layers, batch_size, self.hid_size).to(device)
return hidden, cell
iterator = iterate_minibatches(data_train, 3)
batch, target = next(iterator)
model = ThreeInputsNetLSTM()
model.to(device)
model([torch.tensor(batch['Title'], dtype=torch.long, device=device),
torch.tensor(batch['FullDescription'], dtype=torch.long, device=device),
torch.tensor(batch['Categorical'], dtype=torch.float, device=device)])
opt = torch.optim.Adam(model.parameters())
loss_func = nn.MSELoss() # <YOUR CODE HERE>
epochs = 1
history = []
model.train()
batch_size = 256
for epoch_num in range(epochs):
for idx, (batch, target) in enumerate(iterate_minibatches(data_train, batch_size=batch_size)):
# Preprocessing the batch data and target
batch = [torch.tensor(batch['Title'], dtype=torch.long, device=device),
torch.tensor(batch['FullDescription'], dtype=torch.long, device=device),
torch.tensor(batch['Categorical'], dtype=torch.float, device=device)]
target = torch.tensor(target).to(device)
predictions = model(batch)
predictions = predictions.view(predictions.size(0))
loss = loss_func(predictions, target) # <YOUR CODE HERE>
# train with backprop
# <YOUR CODE HERE>
loss.backward()
opt.step()
opt.zero_grad()
history.append(loss.item())
if (idx+1)%10==0:
clear_output(True)
plt.plot(history, label='loss')
plt.yscale('log')
plt.legend()
plt.show()
def print_metrics(model, data, batch_size=256, name="", **kw):
squared_error = abs_error = num_samples = 0.0
for batch_x, batch_y in tqdm(iterate_minibatches(data, batch_size=batch_size, shuffle=False, **kw)):
batch = [torch.tensor(batch_x['Title'], dtype=torch.long, device=device),
torch.tensor(batch_x['FullDescription'], dtype=torch.long, device=device),
torch.tensor(batch_x['Categorical'], dtype=torch.float, device=device)]
batch_pred = model(batch)[:, 0].detach().cpu().numpy()
squared_error += np.sum(np.square(batch_pred - batch_y))
abs_error += np.sum(np.abs(batch_pred - batch_y))
num_samples += len(batch_y)
print("%s results:" % (name or ""))
print("Mean square error: %.5f" % (squared_error / num_samples))
print("Mean absolute error: %.5f" % (abs_error / num_samples))
return squared_error, abs_error
model.eval()
print_metrics(model, data_train, name='Train')
print_metrics(model, data_val, name='Val');
```
**JUST FOR COMPARING!**
Val results **Simple Model**:<br>
+ Mean square error: 0.32914<br>
+ Mean absolute error: 0.44004<br>
Val results **ThreeInputModel**:<br>
+ Mean square error: 0.11429<br>
+ Mean absolute error: 0.25679<br>
Val results **ThreeInputModelLSTM**:<br>
+ Mean square error: 0.10456<br>
+ Mean absolute error: 0.24456<br>
### Bonus area 3: fixing the data leaks
Fix the data leak we ignored in the beginning of the __Deep Learning part__. Compare results with and without data leaks using same architectures and training time.
```
# <YOUR CODE HERE>
```
__Terrible start-up idea #1962:__ make a tool that automaticaly rephrases your job description (or CV) to meet salary expectations :)
| github_jupyter |
```
import matplotlib.pyplot as plt
from rdkit.Chem import MolFromSmiles
from rdkit.Chem.Descriptors import ExactMolWt
import math
import pandas as pd
import numpy as np
df = pd.read_csv("39_Formose reaction_MeOH.csv")#39_Formose reaction_MeOH.csv
print(df.columns)
# first get rid of empty lines in the mass list by replacing with ''
df.replace('', np.nan, inplace=True)
# also, some 'Mass' values are not numbers
df.dropna(subset=['Mass'], inplace=True)
# now replace NaNs with '' to avoid weird errors
df.fillna('', inplace=True)
df.shape
df.head()
# make a list of exact mass and relative abundance.
mass_list = []
rel_abundance = []
for i in range(len(df)):
if float(df['Mass'].iloc[i]) < 200 and "No Hit" not in df['Molecular Formula'].iloc[i]:
mass_list.append(float(df['Mass'].iloc[i]))
rel_abundance.append(float(df['Rel. Abundance'].iloc[i]))
# now, "renormalize" the relative abundance.
highest = max(rel_abundance)
norm_factor = 100.0/highest
normalized_abun = []
for ab in rel_abundance:
normalized_abun.append(norm_factor*ab)
print(f'{len(mass_list)} items in {mass_list}')
```
It turns out that all weights below MW < 160 had no hit for a molecular formula by the spectrometer.
Now check for any peaks in the spectrum that match our model
```
# for some flexibility, create a container for the figure
fig = plt.figure(figsize=(8,8)) # create a figure object
ax = fig.add_subplot(111) # create an axis object
# first, draw the experimental spectrum
plt.vlines(x=mass_list, ymin=0, ymax=normalized_abun, color='gray')
plt.yscale('log')
#plt.ylim([0.875, 125])
# now, look at the simulated network for matches. Calculate mass using RDKit
wt_gen_dict = {} # a dictionary {weight:generation it first appeared in}
with open("../main/formose/formose_output.txt") as out:
lines = out.readlines()
# a dictionary in which we'll store the smiles string
# and the generation in which it first appeared
smiles_gen_dict = {}
for line in lines:
line_comps = line.split('\t') #items in the line are separated by a tabspace
smiles_gen_dict[line_comps[1]] = line_comps[0][1] #2nd character of component 0 is the generation number
for smiles, gen in smiles_gen_dict.items():
#print(gen+"--->"+smiles)
mol = MolFromSmiles(smiles)
wt = ExactMolWt(mol)
if wt not in wt_gen_dict:
wt_gen_dict[wt]=gen
proton_mass = 1.007276
matching_mass_list = []
match_gen_list = []
# check every peak in the exp spectrum
for obs_mass in mass_list:
for sim_weight, gen in wt_gen_dict.items():
if abs((sim_weight - proton_mass) - obs_mass) < 0.0001:
print(f'{obs_mass} got matched in gen {gen}. It corresponds to MW {obs_mass + proton_mass}')
matching_mass_list.append(obs_mass)
match_gen_list.append(int(gen))
break
# it will be convenient to plot these by generation if I separated them by gen into a map
round_match_dict = {}
for i in range(len(match_gen_list)):
# if this gen id is in the map,
if match_gen_list[i] in round_match_dict.keys():
round_match_dict[match_gen_list[i]].append(matching_mass_list[i])
else:
# each value corresponding to a gen is supposed to be a list of masses (same as those in exp spectrum)
round_match_dict[match_gen_list[i]] = [matching_mass_list[i]]
# we want to color the markers by generation
colors = ['r', 'c', 'g', 'b', 'm', 'k']
for gen_app, matches_list in round_match_dict.items():
# fetch the abundances of each subset of matching peaks
mass_abuns = []
for mass in matches_list:
mass_abuns.append(normalized_abun[mass_list.index(mass)])
#print(f"Matches list: {matches_list}")
#print(f"Abundances: {mass_abuns}")
plt.scatter(matches_list, mass_abuns, marker='x', color=colors[gen_app-2], label=f'Generation {gen_app}')
# also, check for matches with 13-C isotope peaks.
c13_matching_masses = []
c13_match_gens = []
# need to add this to get the mass of 13-C carbon
c13_correction = 1.003355
# now repeat the above exercise
# check every peak in the exp spectrum
for obs_mass in mass_list:
for sim_weight, gen in wt_gen_dict.items():
if abs((sim_weight - proton_mass + c13_correction) - obs_mass) < 0.0001:
print(f'{obs_mass} with 13-C got matched in gen {gen}. It corresponds to MW {obs_mass - c13_correction + proton_mass} w/o C-13')
c13_matching_masses.append(obs_mass)
c13_match_gens.append(int(gen))
break
c13_gen_match_dict = {}
for i in range(len(c13_matching_masses)):
# if this gen id is in the map,
if c13_match_gens[i] in c13_gen_match_dict.keys():
c13_gen_match_dict[c13_match_gens[i]].append(c13_matching_masses[i])
else:
# each value corresponding to a gen is supposed to be a list of masses (same as those in exp spectrum)
c13_gen_match_dict[c13_match_gens[i]] = [c13_matching_masses[i]]
for gen_app, matches_list in c13_gen_match_dict.items():
# fetch the abundances of each subset of matching peaks
mass_abuns = []
for mass in matches_list:
mass_abuns.append(normalized_abun[mass_list.index(mass)])
plt.scatter(matches_list, mass_abuns, marker='o', s=20, color=colors[gen_app-2], label=f'Generation {gen_app} ' + '(C$^{13}$)')
plt.xlabel('Exact Mass')
plt.ylabel('Relative Abundance')
plt.title('Matching peaks in the experimental FT-ICR-MS spectrum of the formose reaction.')
# add a legend
# make the entries in the legend order by generation in ascending order
handles, labels = ax.get_legend_handles_labels()
hl = sorted(zip(handles, labels), key=lambda list: list[1])
h, l = zip(*hl)
plt.legend(h, l, loc='upper left')
plt.savefig('formose_6g_ms.jpg', dpi=300)
plt.show()
print(f'There are total {len(matching_mass_list) + len(c13_matching_masses)} matches out of {len(mass_list)} total peaks in this range in the mass spectra')
# This is old code I was using before. Should be ignored
'''# open the output.txt
weights_dict = {} # a dictionary {weight:generation it first appeared in}
with open("formose_output.txt") as out:
lines = out.readlines()
# a dictionary in which we'll store the smiles string
# and the generation in which it first appeared
smiles_gen_dict = {}
for line in lines:
line_comps = line.split('\t') #items in the line are separated by a tabspace
smiles_gen_dict[line_comps[1]] = line_comps[0][1] #2nd character of component 0 is the generation number
for smiles, gen in smiles_gen_dict.items():
#print(gen+"--->"+smiles)
mol = MolFromSmiles(smiles)
wt = ExactMolWt(mol)
if wt not in weights_dict:
weights_dict[wt]=gen # This part only see the last matching in generation for a specific mass
espectro = open('formose_ms.csv','r')
lines = espectro.readlines()
#plt.ylim([0, 100])
#obs_mass = [] # list of masses in the glucose.csv
#rel_abundance = [] # rel abundance
match_gen = [] # entry is 0 if not matched, else is set equal generation of first appearance
# check for matches with MOD output
proton_mass = 1.007276
for mass in mass_list:
found = False
gen_app = 0
#print(weights_dict)
for sim_mass, gen in weights_dict.items():
if abs((mass + proton_mass) - sim_mass) <= 0.0001:
found = True
gen_app=gen
print(f"{mass} (intensity: {rel_abundance[obs_mass.index(mass)]}) appeared in {gen_app}")
if found == True:
match = int(gen_app)
break
else:
match = 0
match_gen.append(match)
# Now, mark the matching peaks and color them by generation
# diff color for each generation, add a legend
# to fix issues with legend, I had to make a dictionary for matches by generation
gen_mass_match = {}
for i in range(len(mass_list)):
if match_gen[i] in gen_mass_match.keys():
gen_mass_match[match_gen[i]].append(mass_list[i])
else:
gen_mass_match[match_gen[i]] = [mass_list[i]]
# Now, place the markers, labelled by generation
colors= ['r','c','g','b','m','k']
for match_gen, mass_list in gen_mass_match.items():
# I needed the index of the mass in the list obs_mass
intensities = [rel_abundance[mass_list.index(mass)] for mass in mass_list]
if match_gen != 0:
plt.scatter(mass_list, intensities, marker='x', color=colors[match_gen-2], label = f"Generation {match_gen}")
#Glucose info
mod_glucose_smiles="C(C1C(C(C(C(O)O1)O)O)O)O"
mod_glucose_mass= 180.063388104
a = mod_glucose_mass - proton_mass
print(f'Calculated mass of glucose: {a}')
plt.xlabel("Exact Mass")
plt.ylabel("Relative Abundance")
plt.yscale("log")
#legend should be ordered by generation number
import operator
handles, labels = ax.get_legend_handles_labels()
hl = sorted(zip(handles, labels), key= operator.itemgetter(1))
h,l = zip(*hl)
ax.legend(h, l, loc="upper left")
plt.title("Matching peaks in experimental FT-ICR-MS data for the formose reaction")
plt.savefig("formose_mass_spectra6.jpg", dpi=300, transparent=False)
plt.show()'''
# There are values with negative KMD assigned by the spectrometer??
for i in range(len(df['KMD'])):
if float(df['KMD'].iloc[i]) < 0 and "No Hit" not in df['Molecular Formula'].iloc[i]:
print(df.iloc[i])
# What mass does RDKit say, glucose has?
glucose_mol = MolFromSmiles("C(C1C(C(C(C(O)O1)O)O)O)O")
glucose_wt = ExactMolWt(glucose_mol)
print("RDKit calculated glucose weight", glucose_wt)
```
| github_jupyter |
<script async src="https://www.googletagmanager.com/gtag/js?id=UA-59152712-8"></script>
<script>
window.dataLayer = window.dataLayer || [];
function gtag(){dataLayer.push(arguments);}
gtag('js', new Date());
gtag('config', 'UA-59152712-8');
</script>
# Testing Piecewise Polytropic EOS
## Author: Leo Werneck
<a id='toc'></a>
# Table of Contents
$$\label{toc}$$
This module is organized as follows
1. [Step 1](#introduction): **Introduction**
1. [Step 2](#plugging_in_values): **Plugging in some values**
1. [Step 3](#p_cold): **Taking a look at $P_{\rm cold}$**
1. [Step 3.a](#p_cold__computing): *A function to evaluate $P_{\rm cold}$ based on `IllinoisGRMHD`*
1. [Step 3.b](#p_cold__plotting): *Plotting $P_{\rm cold}\left(\rho\right)$*
1. [Step 4](#eps_cold): **Taking a look at $\epsilon_{\rm cold}$**
1. [Step 4.a](#eps_cold__computing): *A function to evaluate $\epsilon_{\rm cold}$ based on `IllinoisGRMHD`*
1. [Step 4.b](#eps_cold__plotting): *Plotting $\epsilon_{\rm cold}\left(\rho\right)$*
1. [Step 5](#ppeos__c_code): **The piecewise polytrope EOS C code**
1. [Step 5.a](#ppeos__c_code__prelim): *Preliminary treatment of the input*
1. [Step 5.a.i](#ppeos__c_code__prelim__computing_ktab): Determining $\left\{K_{1},K_{2},\ldots,K_{\rm neos}\right\}$
1. [Step 5.a.ii](#ppeos__c_code__prelim__computing_eps_integ_consts): Determining $\left\{C_{0},C_{1},C_{2},\ldots,C_{\rm neos}\right\}$
1. [Step 5.b](#ppeos__c_code__eos_struct_setup) *Setting up the `eos_struct`*
1. [Step 5.c](#ppeos__c_code__find_polytropic_k_and_gamma_index) *The `find_polytropic_K_and_Gamma_index()` function*
1. [Step 5.d](#ppeos__c_code__compute_P_cold__eps_cold__dPcold_drho__eps_th__h__Gamma_cold): *The new `compute_P_cold__eps_cold__dPcold_drho__eps_th__h__Gamma_cold()` function*
1. [Step n](#latex_pdf_output): **Output this notebook to $\LaTeX$-formatted PDF file**
<a id='introduction'></a>
# Step 1: Introduction \[Back to [top](#toc)\]
$$\label{introduction}$$
We will test here a few piecewise polytrope (PP) equation of state (EOS) relations. The main reference we will be following is [J.C. Read *et al.* (2008)](https://arxiv.org/pdf/0812.2163.pdf), but we will also try out [the approach of `Whisky`, presented in Takami *et al.* (2014)](https://arxiv.org/pdf/1412.3240v2.pdf)
First we will start with the original one implemented by IllinoisGRMHD:
$$
\boxed{
P_{\rm cold} =
\left\{
\begin{matrix}
K_{0}\rho^{\Gamma_{0}} & , & \rho \leq \rho_{0}\\
K_{1}\rho^{\Gamma_{1}} & , & \rho_{0} \leq \rho \leq \rho_{1}\\
\vdots & & \vdots\\
K_{j}\rho^{\Gamma_{j}} & , & \rho_{j-1} \leq \rho \leq \rho_{j}\\
\vdots & & \vdots\\
K_{N-1}\rho^{\Gamma_{N-1}} & , & \rho_{N-2} \leq \rho \leq \rho_{N-1}\\
K_{N}\rho^{\Gamma_{N}} & , & \rho \geq \rho_{N-1}
\end{matrix}
\right.
}\ .
$$
Notice that we have the following sets of variables:
$$
\left\{\underbrace{\rho_{0},\rho_{1},\ldots,\rho_{N-1}}_{N\ {\rm values}}\right\}\ ;\
\left\{\underbrace{K_{0},K_{1},\ldots,K_{N}}_{N+1\ {\rm values}}\right\}\ ;\
\left\{\underbrace{\Gamma_{0},\Gamma_{1},\ldots,\Gamma_{N}}_{N+1\ {\rm values}}\right\}\ .
$$
Also, notice that $K_{0}$ and the entire sets $\left\{\rho_{0},\rho_{1},\ldots,\rho_{N-1}\right\}$ and $\left\{\Gamma_{0},\Gamma_{1},\ldots,\Gamma_{N}\right\}$ must be specified by the user. The values of $\left\{K_{1},\ldots,K_{N}\right\}$, on the other hand, are determined by imposing that $P_{\rm cold}$ be continuous, i.e.
$$
P_{\rm cold}\left(\rho_{0}\right) = K_{0}\rho_{0}^{\Gamma_{0}} = K_{1}\rho_{0}^{\Gamma_{1}} \implies
\boxed{K_{1} = K_{0}\rho_{0}^{\Gamma_{0}-\Gamma_{1}}}\ .
$$
Analogously,
$$
\boxed{K_{j} = K_{j-1}\rho_{j-1}^{\Gamma_{j-1}-\Gamma_{j}}\ ,\ j\in\left[1,N\right]}\ .
$$
<a id='plugging_in_values'></a>
# Step 2: Plugging in some values \[Back to [top](#toc)\]
$$\label{plugging_in_values}$$
Just so that we work with realistic values (i.e. values actually used by researchers), we will implement a simple check using the values from [Table II in J.C. Read *et al.* (2008)](https://arxiv.org/pdf/0812.2163.pdf):
| $\rho_{i}$ | $\Gamma_{i}$ | $K_{\rm expected}$ |
|------------|--------------|--------------------|
|2.44034e+07 | 1.58425 | 6.80110e-09 |
|3.78358e+11 | 1.28733 | 1.06186e-06 |
|2.62780e+12 | 0.62223 | 5.32697e+01 |
| $-$ | 1.35692 | 3.99874e-08 |
```
# Determining K_{i} i != 0
# We start by setting up all the values of rho_{i}, Gamma_{i}, and K_{0}
import numpy as np
rho_ppoly_tab = [2.44034e+07,3.78358e+11,2.62780e+12]
Gamma_ppoly_tab = [1.58425,1.28733,0.62223,1.35692]
K_ppoly_tab = [6.80110e-09,0,0,0]
K_expected = [6.80110e-09,1.06186e-06,5.32697e+01,3.99874e-08]
NEOS = len(rho_ppoly_tab)+1
for j in range(1,NEOS):
K_ppoly_tab[j] = K_ppoly_tab[j-1] * rho_ppoly_tab[j-1]**(Gamma_ppoly_tab[j-1] - Gamma_ppoly_tab[j])
print("K_ppoly_tab["+str(j)+"] = "+str(K_ppoly_tab[j])+\
" | K_expected["+str(j)+"] = "+str(K_expected[j])+\
" | Diff = "+str(np.fabs(K_ppoly_tab[j] - K_expected[j])))
```
<a id='p_cold'></a>
# Step 3: Taking a look at $P_{\rm cold}$ \[Back to [top](#toc)\]
$$\label{p_cold}$$
<a id='p_cold__computing'></a>
## Step 3.a: A function to evaluate $P_{\rm cold}$ based on `IllinoisGRMHD` \[Back to [top](#toc)\]
$$\label{p_cold__computing}$$
The results above look reasonable to the expected ones. Let us then see what the plot of $P_{\rm cold}\times\rho$ looks like. For the case of our specific example, which has $N_{\rm EOS}=4$ (where $N_{\rm EOS}$ stands for the number of polytropic EOSs used), we have
$$
\boxed{
P_{\rm cold} =
\left\{
\begin{matrix}
K_{0}\rho^{\Gamma_{0}} & , & \rho \leq \rho_{0}\\
K_{1}\rho^{\Gamma_{1}} & , & \rho_{0} \leq \rho \leq \rho_{1}\\
K_{2}\rho^{\Gamma_{2}} & , & \rho_{1} \leq \rho \leq \rho_{2}\\
K_{3}\rho^{\Gamma_{3}} & , & \rho \geq \rho_{2}
\end{matrix}
\right.
}\ .
$$
```
# Set the pressure through the polytropic EOS: P_cold = K * rho^{Gamma}
def P_cold(rho,rho_ppoly_tab,Gamma_ppoly_tab,K_ppoly_tab,NEOS):
if rho < rho_ppoly_tab[0] or NEOS==1:
return K_ppoly_tab[0] * rho**Gamma_ppoly_tab[0]
if NEOS > 2:
for j in range(1,NEOS-1):
if rho_ppoly_tab[j-1] <= rho and rho < rho_ppoly_tab[j]:
return K_ppoly_tab[j] * rho**Gamma_ppoly_tab[j]
if rho >= rho_ppoly_tab[NEOS-2]:
return K_ppoly_tab[NEOS-1] * rho**Gamma_ppoly_tab[NEOS-1]
```
<a id='p_cold__plotting'></a>
## Step 3.b: Plotting $P_{\rm cold}\left(\rho\right)$ \[Back to [top](#toc)\]
$$\label{p_cold__plotting}$$
To create a reasonably interesting plot, which will also test all regions of our piecewise polytrope EOS, let us plot $P_{\rm cold}\left(\rho\right)$ with $\rho\in\left[\frac{\rho_{0}}{10},10\rho_{2}\right]$.
**Remember**: We *don't* expect $P_{\rm cold}$ to be *smooth*, but we *do* expect it to be *continuous* (by construction).
```
# Set rho in [rho_ppoly_tab[0]/10,rho_ppoly_tab[2]*10]
rho_for_plot = np.linspace(rho_ppoly_tab[0]/10,rho_ppoly_tab[NEOS-2]*10,1000)
# Set P_{cold}(rho)
P_for_plot = np.linspace(0,0,1000)
for i in range(len(P_for_plot)):
P_for_plot[i] = P_cold(rho_for_plot[i],rho_ppoly_tab,Gamma_ppoly_tab,K_ppoly_tab,NEOS)
# Plot P_{cold}(rho) x rho
import matplotlib.pyplot as plt
f = plt.figure(figsize=(10,4))
ax1 = f.add_subplot(121)
ax1.set_title(r"Plot #1: $P_{\rm cold}\times\rho_{b}$",fontsize='16')
ax1.set_xlabel(r"$\rho_{b}$",fontsize='14')
ax1.set_ylabel(r"$P_{\rm cold}$",fontsize='14')
ax1.grid()
ax1.plot(rho_for_plot,P_for_plot)
ax2 = f.add_subplot(122)
ax2.set_title(r"Plot #2: $\log_{10}\left(P_{\rm cold}\right)\times\log_{10}\left(\rho_{b}\right)$",fontsize='16')
ax2.set_xlabel(r"$\log_{10}\left(\rho_{b}\right)$",fontsize='14')
ax2.set_ylabel(r"$\log_{10}\left(P_{\rm cold}\right)$",fontsize='14')
ax2.grid()
ax2.plot(np.log10(rho_for_plot),np.log10(P_for_plot))
```
<a id='eps_cold'></a>
# Step 4: Taking a look at $\epsilon_{\rm cold}$ \[Back to [top](#toc)\]
$$\label{eps_cold}$$
$\epsilon_{\rm cold}$ is determined via
$$
\epsilon_{\rm cold} = \int d\rho\, \frac{P_{\rm cold}}{\rho^{2}} \ ,
$$
for some integration constant $C$. **Notation alert**: in the literature, the integration constants $C_{j}$ below are usually called $\epsilon_{j}$. We will keep them as $C_{j}$ for a clearer exposition.
In the case of a piecewise polytropic EOS, we then must have
$$
\boxed{
\epsilon_{\rm cold} =
\left\{
\begin{matrix}
\frac{K_{0}\rho^{\Gamma_{0}-1}}{\Gamma_{0}-1} + C_{0} & , & \rho \leq \rho_{0}\\
\frac{K_{1}\rho^{\Gamma_{1}-1}}{\Gamma_{1}-1} + C_{1} & , & \rho_{0} \leq \rho \leq \rho_{1}\\
\vdots & & \vdots\\
\frac{K_{j}\rho^{\Gamma_{j}-1}}{\Gamma_{j}-1} + C_{j} & , & \rho_{j-1} \leq \rho \leq \rho_{j}\\
\vdots & & \vdots\\
\frac{K_{N-1}\rho^{\Gamma_{N-1}-1}}{\Gamma_{N-1}-1} + C_{N-1} & , & \rho_{N-2} \leq \rho \leq \rho_{N-1}\\
\frac{K_{N}\rho^{\Gamma_{N}-1}}{\Gamma_{N}-1} + C_{N} & , & \rho \geq \rho_{N-1}
\end{matrix}
\right.
}\ .
$$
We fix $C_{0}$ by demanding that $\epsilon_{\rm cold}\left(\rho=0\right) = 0$. Then, continuity of $\epsilon_{\rm cold}$ imposes that
$$
\frac{K_{0}\rho_{0}^{\Gamma_{0}-1}}{\Gamma_{0}-1} = \frac{K_{1}\rho_{0}^{\Gamma_{1}-1}}{\Gamma_{1}-1} + C_{1}\implies \boxed{C_{1} = \frac{K_{0}\rho_{0}^{\Gamma_{0}-1}}{\Gamma_{0}-1} - \frac{K_{1}\rho_{0}^{\Gamma_{1}-1}}{\Gamma_{1}-1}}\ ,
$$
for $C_{1}$ and
$$
\boxed{C_{j} = C_{j-1} + \frac{K_{j-1}\rho_{j-1}^{\Gamma_{j-1}-1}}{\Gamma_{j-1}-1} - \frac{K_{j}\rho_{j-1}^{\Gamma_{j}-1}}{\Gamma_{j}-1}\ ,\ j\geq1}\ ,
$$
generically.
<a id='eps_cold__computing'></a>
## Step 4.a: *A function to evaluate $\epsilon_{\rm cold}$ based on `IllinoisGRMHD`* \[Back to [top](#toc)\]
$$\label{eps_cold__computing}$$
We continue with our previous values of the piecewise polytrope EOS, for which we will have
$$
\boxed{
\epsilon_{\rm cold} =
\left\{
\begin{matrix}
\frac{K_{0}\rho^{\Gamma_{0}-1}}{\Gamma_{0}-1} & , & \rho \leq \rho_{0}\\
\frac{K_{1}\rho^{\Gamma_{1}-1}}{\Gamma_{1}-1} + C_{1} & , & \rho_{0} \leq \rho \leq \rho_{1}\\
\frac{K_{2}\rho^{\Gamma_{2}-1}}{\Gamma_{2}-1} + C_{2} & , & \rho_{1} \leq \rho \leq \rho_{2}\\
\frac{K_{3}\rho^{\Gamma_{3}-1}}{\Gamma_{3}-1} + C_{3} & , & \rho \geq \rho_{2}
\end{matrix}
\right.
}\ ,
$$
remembering that the integration constants are set via
$$
\boxed{
C_{j} =
\left\{
\begin{matrix}
0 & , & {\rm if}\ j=0\\
C_{j-1} + \frac{K_{j-1}\rho_{j-1}^{\Gamma_{j-1}-1}}{\Gamma_{j-1}-1} - \frac{K_{j}\rho_{j-1}^{\Gamma_{j}-1}}{\Gamma_{j}-1} & , & {\rm otherwise}
\end{matrix}
\right.
}\ .
$$
```
# Setting eps_cold
def eps_cold__integration_constants(rho_ppoly_tab,Gamma_ppoly_tab,K_ppoly_tab,NEOS):
if NEOS == 1:
eps_integ_consts[0] = 0
return eps_integ_consts
eps_integ_consts = [0 for i in range(NEOS)]
for j in range(1,NEOS):
aux_jm1 = K_ppoly_tab[j-1] * rho_ppoly_tab[j-1]**(Gamma_ppoly_tab[j-1]-1) / (Gamma_ppoly_tab[j-1]-1)
aux_jp0 = K_ppoly_tab[j+0] * rho_ppoly_tab[j-1]**(Gamma_ppoly_tab[j+0]-1) / (Gamma_ppoly_tab[j+0]-1)
eps_integ_consts[j] = eps_integ_consts[j-1] + aux_jm1 - aux_jp0
return eps_integ_consts
def eps_cold(rho,rho_ppoly_tab,Gamma_ppoly_tab,K_ppoly_tab,NEOS):
if rho < rho_ppoly_tab[0] or NEOS==1:
return K_ppoly_tab[0] * rho**(Gamma_ppoly_tab[0]-1) / (Gamma_ppoly_tab[0]-1)
# Compute C_{j}
eps_integ_consts = eps_cold__integration_constants(rho_ppoly_tab,Gamma_ppoly_tab,K_ppoly_tab,NEOS)
# Compute eps_cold for rho >= rho_{0}
if NEOS>2:
for j in range(1,NEOS-1):
if rho >= rho_ppoly_tab[j-1] and rho < rho_ppoly_tab[j]:
return eps_integ_consts[j] + K_ppoly_tab[j] * rho**(Gamma_ppoly_tab[j]-1) / (Gamma_ppoly_tab[j]-1)
if rho >= rho_ppoly_tab[NEOS-2]:
return eps_integ_consts[NEOS-1] + K_ppoly_tab[NEOS-1] * rho**(Gamma_ppoly_tab[NEOS-1]-1) / (Gamma_ppoly_tab[NEOS-1]-1)
```
<a id='eps_cold__plotting'></a>
## Step 4.b: Plotting $\epsilon_{\rm cold}$ \[Back to [top](#toc)\]
$$\label{eps_cold__plotting}$$
To create a reasonably interesting plot, which will also test all regions of our piecewise polytrope EOS, let us plot $\epsilon_{\rm cold}\left(\rho\right)$ with $\rho\in\left[\frac{\rho_{0}}{10},10\rho_{2}\right]$.
**Remember**: We *don't* expect $\epsilon_{\rm cold}$ to be *smooth*, but we *do* expect it to be *continuous* (by construction).
```
# Set rho in [rho_ppoly_tab[0]/10,rho_ppoly_tab[2]*10]
rho_for_plot = np.linspace(rho_ppoly_tab[0]/10,rho_ppoly_tab[NEOS-2]*10,1000)
# Set P_{cold}(rho)
eps_for_plot = np.linspace(0,0,1000)
for i in range(len(eps_for_plot)):
eps_for_plot[i] = eps_cold(rho_for_plot[i],rho_ppoly_tab,Gamma_ppoly_tab,K_ppoly_tab,NEOS)
# Plot P_{cold}(rho) x rho
import matplotlib.pyplot as plt
f = plt.figure(figsize=(12,4))
ax1 = f.add_subplot(121)
ax1.set_title(r"Plot #1: $\varepsilon_{\rm cold}\times\rho_{b}$",fontsize='16')
ax1.set_xlabel(r"$\rho_{b}$",fontsize='14')
ax1.set_ylabel(r"$\varepsilon_{\rm cold}$",fontsize='14')
ax1.grid()
ax1.plot(rho_for_plot,eps_for_plot)
ax2 = f.add_subplot(122)
ax2.set_title(r"Plot #2: $\log_{10}\left(\varepsilon_{\rm cold}\right)\times\log_{10}\left(\rho_{b}\right)$",fontsize='16')
ax2.set_xlabel(r"$\log_{10}\left(\rho_{b}\right)$",fontsize='14')
ax2.set_ylabel(r"$\log_{10}\left(\varepsilon_{\rm cold}\right)$",fontsize='14')
ax2.grid()
ax2.plot(np.log10(rho_for_plot),np.log10(eps_for_plot))
```
<a id='ppeos__c_code'></a>
# Step 5: The piecewise polytrope EOS C code \[Back to [top](#toc)\]
$$\label{ppeos__c_code}$$
Now we will begin implementing a C code to handle all the topics we discussed here.
<a id='ppeos__c_code__prelim'></a>
## Step 5.a: Preliminary treatment of the input \[Back to [top](#toc)\]
$$\label{ppeos__c_code__prelim}$$
Here we assume that we get as input the following values:
1. neos: the number of polytropic EOSs we will be handling (equivalent to $N+1$ in the discussion above)
1. rho_ppoly_tab: an array containing the values $\left\{\rho_{0},\rho_{1},\ldots,\rho_{{\rm neos}-1}\right\}$. Notice that for both neos=1 and neos=2, the size of this array is $1$.
1. K_ppoly_tab: an array of size neos containing only the value of $K_{0}$ in the entry K_ppoly_tab[0].
1. Gamma_ppoly_tab: an array of size neos containing the values $\left\{\Gamma_{0},\Gamma_{1},\ldots,\Gamma_{N}\right\}$.
```
import os,sys
IGM_src_dir = os.path.join("..","src")
Piecewise_Polytrope_EOS__C = os.path.join(IGM_src_dir,"IllinoisGRMHD_EoS_lowlevel_functs.C")
%%writefile $Piecewise_Polytrope_EOS__C
#ifndef ILLINOISGRMHD_EOS_FUNCTS_C_
#define ILLINOISGRMHD_EOS_FUNCTS_C_
/* Function : setup_K_ppoly_tab__and__eps_integ_consts()
* Authors : Leo Werneck
* Description : For a given set of EOS inputs, determine
* values of K_ppoly_tab that will result in a
* everywhere continuous P_cold function.
* Dependencies: none
*
* Inputs : eos - a struct containing the following
* relevant quantities:
* : neos - number of polytropic EOSs used
* : rho_ppoly_tab - array of rho values that determine
* the polytropic EOS to be used.
* : Gamma_ppoly_tab - array of Gamma_cold values to be
* used in each polytropic EOS.
* : K_ppoly_tab - array of K_ppoly_tab values to be used
* in each polytropic EOS. Only K_ppoly_tab[0]
* is known prior to the function call
* : eps_integ_const - array of C_{j} values, which are the
* integration constants that arrise when
* determining eps_{cold} for a piecewise
* polytropic EOS. This array should be
* uninitialized or contain absurd values
* prior to this function call.
*
* Outputs : K_ppoly_tab - fully populated array of K_ppoly_tab
* to be used in each polytropic EOS.
* : eps_integ_const - fully populated array of C_{j}'s,
* used to compute eps_cold for
* a piecewise polytropic EOS.
*/
static void setup_K_ppoly_tab__and__eps_integ_consts(eos_struct &eos){
```
<a id='ppeos__c_code__prelim__computing_ktab'></a>
### Step 5.a.i: Determining $\left\{K_{1},K_{2},\ldots,K_{\rm neos}\right\}$ \[Back to [top](#toc)\]
$$\label{ppeos__c_code__prelim__computing_ktab}$$
We start by computing the values $\left\{K_{1},K_{2},\ldots,K_{\rm neos}\right\}$ from the input values. Remember that the values of $K_{j}$ are obtained by demanding that the $P_{\rm cold}$ function be everywhere continuous, resulting in the relation (see [the discussion above for the details](#introduction))
$$
\boxed{K_{j} = K_{j-1}\rho_{j-1}^{\Gamma_{j-1}-\Gamma_{j}}\ ,\ j\in\left[1,N\right]}\ .
$$
**Implementation note**: Because the case neos=1 is handled first, the attentive reader will notice that we already set $C_{0}=0$ below, *before* discussing its implementation. For more on that, please refer to the [next subsection, where we discuss determining $\left\{C_{0},C_{1},C_{2},\ldots,C_{\rm neos}\right\}$](#ppeos__c_code__prelim__computing_eps_integ_consts).
```
%%writefile -a $Piecewise_Polytrope_EOS__C
/* When neos = 1, we will only need the value K_ppoly_tab[0] and eps_integ_const[0].
* Since our only polytropic EOS is given by
* -----------------------------------
* | P_{0} = K_{0} * rho ^ (Gamma_{0}) | ,
* -----------------------------------
* and, therefore,
* ---------------------------------------------------------------
* | eps_{0} = K_{0} * rho ^ (Gamma_{0}-1) / (Gamma_{0}-1) + C_{0} | ,
* ---------------------------------------------------------------
* we only need to set up K_{0} := K_ppoly_tab[0] and C_{0} := eps_integ_const[0].
* K_{0} is a user input, so we need to do nothing. C_{0}, on the other hand,
* is fixed by demanding that eps(rho) -> 0 as rho -> 0. Thus, C_{0} = 0.
*/
eos.eps_integ_const[0] = 0.0;
if(eos.neos==1) return;
/********************
* Setting up K_{j} *
********************/
/* When neos > 1, we have the following structure
*
* / K_{0} * rho^(Gamma_{0}) , rho < rho_{0}
* | K_{1} * rho^(Gamma_{1}) , rho_{0} <= rho < rho_{1}
* | ...
* P(rho) = < K_{j} * rho^(Gamma_{j}) , rho_{j-1} <= rho < rho_{j}
* | ...
* | K_{neos-2} * rho^(Gamma_{neos-2}) , rho_{neos-3} <= rho < rho_{neos-2}
* \ K_{neos-1} * rho^(Gamma_{neos-1}) , rho >= rho_{neos-2}
*
* Imposing that P(rho) be everywhere continuous, we have
* -------------------------------------------------
* | K_{j} = K_{j-1} * rho^(Gamma_{j-1} - Gamma_{j}) |
* -------------------------------------------------
*/
for(int j=1; j<eos.neos; j++){
// Set a useful auxiliary variable to keep things more compact:
// First, (Gamma_{j-1} - Gamma_{j}):
CCTK_REAL Gamma_diff = eos.Gamma_ppoly_tab[j-1] - eos.Gamma_ppoly_tab[j];
// Implement the boxed equation above, using our auxiliary variable:
eos.K_ppoly_tab[j] = eos.K_ppoly_tab[j-1] * pow(eos.rho_ppoly_tab[j-1],Gamma_diff);
}
```
<a id='ppeos__c_code__prelim__computing_eps_integ_consts'></a>
### Step 5.a.ii: Determining $\left\{C_{0},C_{1},C_{2},\ldots,C_{\rm neos}\right\}$ \[Back to [top](#toc)\]
$$\label{ppeos__c_code__prelim__computing_eps_integ_consts}$$
We now focus our attention to the computation of $\left\{C_{0},C_{1},C_{2},\ldots,C_{\rm neos}\right\}$, which are the integration constants that emerge when computing $\epsilon_{\rm cold}$ for a piecewise polytrope EOS (see [the discussion above for the details](#eps_cold)). Here we set them so that $\epsilon_{\rm cold}$ is also everywhere continuous, i.e.
$$
\boxed{
C_{j} =
\left\{
\begin{matrix}
0 & , & {\rm if}\ j=0\\
C_{j-1} + \frac{K_{j-1}\rho_{j-1}^{\Gamma_{j-1}-1}}{\Gamma_{j-1}-1} - \frac{K_{j}\rho_{j-1}^{\Gamma_{j}-1}}{\Gamma_{j}-1} & , & {\rm otherwise}
\end{matrix}
\right.
}\ .
$$
```
%%writefile -a $Piecewise_Polytrope_EOS__C
/********************
* Setting up C_{j} *
********************/
/* When neos > 1, we have the following structure (let neos->N):
*
* / K_{0}*rho^(Gamma_{0}-1)/(Gamma_{0}-1) + C_{0}, rho < rho_{0}
* | K_{1}*rho^(Gamma_{1}-1)/(Gamma_{1}-1) + C_{1}, rho_{0} <= rho < rho_{1}
* | ...
* eps(rho) = < K_{j}*rho^(Gamma_{j}-1)/(Gamma_{j}-1) + C_{j}, rho_{j-1} <= rho < rho_{j}
* | ...
* | K_{N-2}*rho^(Gamma_{N-2}-1)/(Gamma_{N-2}-1) + C_{N-2}, rho_{N-3} <= rho < rho_{N-2}
* \ K_{N-1}*rho^(Gamma_{N-1}-1)/(Gamma_{N-1}-1) + C_{N-1}, rho >= rho_{N-2}
*
* Imposing that eps_{cold}(rho) be everywhere continuous, we have
* ---------------------------------------------------------------
* | C_{j} = C_{j-1} |
* | + ( K_{j-1}*rho_{j-1}^(Gamma_{j-1}-1) )/(Gamma_{j-1}-1) |
* | - ( K_{j+0}*rho_{j-1}^(Gamma_{j+0}-1) )/(Gamma_{j+0}-1) |
* ---------------------------------------------------------------
*/
for(int j=1; j<eos.neos; j++){
// Set a few useful auxiliary variables to keep things more compact:
// First, (Gamma_{j-1}-1):
CCTK_REAL Gammajm1m1 = eos.Gamma_ppoly_tab[j-1] - 1.0;
// Then, (Gamma_{j+0}-1):
CCTK_REAL Gammajp0m1 = eos.Gamma_ppoly_tab[j+0] - 1.0;
// Next, ( K_{j-1}*rho_{j-1}^(Gamma_{j-1}-1) )/(Gamma_{j-1}-1):
CCTK_REAL aux_epsm1 = eos.K_ppoly_tab[j-1]*pow(eos.rho_ppoly_tab[j-1],Gammajm1m1)/Gammajm1m1;
// Finally, ( K_{j+0}*rho_{j+0}^(Gamma_{j+0}-1) )/(Gamma_{j+0}-1):
CCTK_REAL aux_epsp0 = eos.K_ppoly_tab[j+0]*pow(eos.rho_ppoly_tab[j-1],Gammajp0m1)/Gammajp0m1;
// Implement the boxed equation above, using our auxiliary variables:
eos.eps_integ_const[j] = eos.eps_integ_const[j-1] + aux_epsm1 - aux_epsp0;
}
}
```
<a id='ppeos__c_code__eos_struct_setup'></a>
## Step 5.b: Setting up the `eos_struct` \[Back to [top](#toc)\]
$$\label{ppeos__c_code__eos_struct_setup}$$
We will now set up the `eos_struct` from input. The `eos_struct` declaration can be found inside the `IllinoisGRMHD_headers.h` source file, but we will repeat it here for the sake of the reader:
```c
#define MAX_EOS_PARAMS 10
struct eos_struct {
int neos;
CCTK_REAL rho_ppoly_tab[MAX_EOS_PARAMS-1];
CCTK_REAL eps_integ_const[MAX_EOS_PARAMS],K_ppoly_tab[MAX_EOS_PARAMS],Gamma_ppoly_tab[MAX_EOS_PARAMS];
CCTK_REAL Gamma_th;
};
```
```
%%writefile -a $Piecewise_Polytrope_EOS__C
/* Function : initialize_EOS_struct_from_input()
* Authors : Leo Werneck
* Description : Initialize the eos struct from user
* input
* Dependencies: setup_K_ppoly_tab__and__eps_integ_consts()
* : cctk_parameters.h (FIXME)
*
* Inputs : eos - a struct containing the following
* relevant quantities:
* : neos - number of polytropic EOSs used
* : rho_ppoly_tab - array of rho values that determine
* the polytropic EOS to be used.
* : Gamma_ppoly_tab - array of Gamma_cold values to be
* used in each polytropic EOS.
* : K_ppoly_tab - array of K_ppoly_tab values to be used
* in each polytropic EOS.
* : eps_integ_const - array of C_{j} values, which are the
* integration constants that arrise when
* determining eps_{cold} for a piecewise
* polytropic EOS.
*
* Outputs : eos - fully initialized EOS struct
*/
static void initialize_EOS_struct_from_input(eos_struct &eos){
/* We start by setting up the eos_struct
* with the inputs given by the user at
* the start of the simulation. Keep in
* mind that these parameters are found
* in the "cctk_parameters.h" header file.
* ^^^^^^^FIXME^^^^^^^
*/
// Initialize: neos, {rho_{j}}, {K_{0}}, and {Gamma_{j}}
#ifndef ENABLE_STANDALONE_IGM_C2P_SOLVER
DECLARE_CCTK_PARAMETERS;
#endif
eos.neos = neos;
eos.K_ppoly_tab[0] = K_ppoly_tab0;
for(int j=0; j<=neos-2; j++) eos.rho_ppoly_tab[j] = rho_ppoly_tab_in[j];
for(int j=0; j<=neos-1; j++) eos.Gamma_ppoly_tab[j] = Gamma_ppoly_tab_in[j];
// Initialize {K_{j}}, j>=1, and {eps_integ_const_{j}}
setup_K_ppoly_tab__and__eps_integ_consts(eos);
}
```
<a id='ppeos__c_code__find_polytropic_k_and_gamma_index'></a>
## Step 5.c: The `find_polytropic_K_and_Gamma_index()` function \[Back to [top](#toc)\]
$$\label{ppeos__c_code__find_polytropic_k_and_gamma_index}$$
```
%%writefile -a $Piecewise_Polytrope_EOS__C
/* Function : find_polytropic_K_and_Gamma_index()
* Authors : Leo Werneck & Zach Etienne
* Description : For a given value of rho, find the
* appropriate values of Gamma_ppoly_tab
* and K_ppoly_tab by determining the appropriate
* index
* Dependencies: initialize_EOS_struct_from_input()
* : cctk_parameters.h (FIXME)
*
* Inputs : eos - a struct containing the following
* relevant quantities:
* : neos - number of polytropic EOSs used
* : rho_ppoly_tab - array of rho values that determine
*
* Outputs : index - the appropriate index for the K_ppoly_tab
* and Gamma_ppoly_tab array
*/
static inline int find_polytropic_K_and_Gamma_index(eos_struct eos, CCTK_REAL rho_in) {
/* We want to find the appropriate polytropic EOS for the
* input value rho_in. Remember that:
*
* if rho < rho_{0}: P_{0} , index: 0
* if rho >= rho_{0} but < rho_{1}: P_{1} , index: 1
* if rho >= rho_{1} but < rho_{2}: P_{2} , index: 2
* ...
* if rho >= rho_{j-1} but < rho_{j}: P_{j} , index: j
*
* Then, a simple way of determining the index is through
* the formula:
* ---------------------------------------------------------------------------
* | index = (rho >= rho_{0}) + (rho >= rho_{1}) + ... + (rho >= rho_{neos-2}) |
* ---------------------------------------------------------------------------
*/
if(eos.neos == 1) return 0;
int polytropic_index = 0;
for(int j=0; j<=eos.neos-2; j++) polytropic_index += (rho_in >= eos.rho_ppoly_tab[j]);
return polytropic_index;
}
%%writefile -a $Piecewise_Polytrope_EOS__C
/* Function : compute_P_cold__eps_cold()
* Authors : Leo Werneck
* Description : Computes P_cold and eps_cold.
* Dependencies: initialize_EOS_struct_from_input()
* : find_polytropic_K_and_Gamma_index()
*
* Inputs : P_cold - cold pressure
* : eps_cold - cold specific internal energy
* : eos - a struct containing the following
* relevant quantities:
* : neos - number of polytropic EOSs used
* : rho_ppoly_tab - array of rho values that determine
* the polytropic EOS to be used.
* : Gamma_ppoly_tab - array of Gamma_cold values to be
* used in each polytropic EOS.
* : K_ppoly_tab - array of K_ppoly_tab values to be used
* in each polytropic EOS.
* : eps_integ_const - array of C_{j} values, which are the
* integration constants that arrise when
* determining eps_{cold} for a piecewise
* polytropic EOS.
*
* Outputs : P_cold - cold pressure (supports SPEOS and PPEOS)
* : eps_cold - cold specific internal energy (supports SPEOS and PPEOS)
* : polytropic_index - polytropic index used for P_cold and eps_cold
*
* SPEOS: Single-Polytrope Equation of State
* PPEOS: Piecewise Polytrope Equation of State
*/
static inline void compute_P_cold__eps_cold(eos_struct eos, CCTK_REAL rho_in,
CCTK_REAL &P_cold,CCTK_REAL &eps_cold) {
// This code handles equations of state of the form defined
// in Eqs 13-16 in http://arxiv.org/pdf/0802.0200.pdf
if(rho_in==0) {
P_cold = 0.0;
eps_cold = 0.0;
return;
}
/* --------------------------------------------------
* | Single and Piecewise Polytropic EOS modification |
* --------------------------------------------------
*
* We now begin our modifications to this function so that
* it supports both single and piecewise polytropic equations
* of state.
*
* The modifications below currently assume that the user
* has called the recently added function
*
* - initialize_EOS_struct_from_input()
*
* *before* this function is called. We can add some feature
* to check this automatically as well, but we'll keep that as
* a TODO/FIXME for now.
*/
/* First, we compute the pressure, which in the case of a
* piecewise polytropic EOS is given by
*
* / P_{1} / K_{1} * rho^(Gamma_{1}) , rho_{0} <= rho < rho_{1}
* | ... | ...
* P(rho) = < P_{j} = < K_{j} * rho^(Gamma_{j}) , rho_{j-1} <= rho < rho_{j}
* | ... | ...
* \ P_{neos-2} \ K_{neos-2} * rho^(Gamma_{neos-2}), rho_{neos-3} <= rho < rho_{neos-2}
*
* The index j is determined by the find_polytropic_K_and_Gamma_index() function.
*/
// Set up useful auxiliary variables
int polytropic_index = find_polytropic_K_and_Gamma_index(eos,rho_in);
CCTK_REAL K_ppoly_tab = eos.K_ppoly_tab[polytropic_index];
CCTK_REAL Gamma_ppoly_tab = eos.Gamma_ppoly_tab[polytropic_index];
CCTK_REAL eps_integ_const = eos.eps_integ_const[polytropic_index];
// Then compute P_{cold}
P_cold = K_ppoly_tab*pow(rho_in,Gamma_ppoly_tab);
/* Then we compute the cold component of the specific internal energy,
* which in the case of a piecewise polytropic EOS is given by (neos -> N)
*
* / P_{1}/(rho*(Gamma_{1}-1)) + C_{1} , rho_{0} <= rho < rho_{1}
* | ...
* eps(rho) = < P_{j}/(rho*(Gamma_{j}-1)) + C_{j} , rho_{j-1} <= rho < rho_{j}
* | ...
* \ P_{N-2}/(rho*(Gamma_{N-2}-1)) + C_{N-2}, rho_{N-3} <= rho < rho_{N-2}
*/
eps_cold = P_cold/(rho_in*(Gamma_ppoly_tab-1.0)) + eps_integ_const;
}
```
<a id='ppeos__c_code__compute_P_cold__eps_cold__dPcold_drho__eps_th__h__Gamma_cold'></a>
## Step 5.d: The new `compute_P_cold__eps_cold__dPcold_drho__eps_th__h__Gamma_cold()` function \[Back to [top](#toc)\]
$$\label{ppeos__c_code__compute_P_cold__eps_cold__dPcold_drho__eps_th__h__Gamma_cold}$$
We now write down the updated version of the `compute_P_cold__eps_cold__dPcold_drho__eps_th__h__Gamma_cold()` function, which is found in the `inlined_functions.C` source file of `IllinoisGRMHD`, and documented in the [`IllinoisGRMHD`'s inlined functions NRPy+ tutorial module](Tutorial-IllinoisGRMHD__inlined_functions.ipynb).
```
%%writefile -a $Piecewise_Polytrope_EOS__C
/* Function : compute_P_cold__eps_cold__dPcold_drho__eps_th__h__Gamma_cold()
* Authors : Leo Werneck & Zach Etienne
* Description : Compute basic quantities related to
* : the EOS, namely: P_cold, eps_cold,
* : dPcold/drho, eps_th, h, and Gamma_cold
* Dependencies: initialize_EOS_struct_from_input()
*
* Inputs : U - array containing primitives {rho,P,v^{i},B^i}
* : P_cold - cold pressure
* : eps_cold - cold specific internal energy
* : dPcold_drho - derivative of P_cold
* : eps_th - thermal specific internal energy
* : h - enthalpy
* : Gamma_cold - cold polytropic Gamma
* : eos - a struct containing the following
* relevant quantities:
* : neos - number of polytropic EOSs used
* : rho_ppoly_tab - array of rho values that determine
* the polytropic EOS to be used.
* : Gamma_ppoly_tab - array of Gamma_cold values to be
* used in each polytropic EOS.
* : K_ppoly_tab - array of K_ppoly_tab values to be used
* in each polytropic EOS.
* : eps_integ_const - array of C_{j} values, which are the
* integration constants that arrise when
* determining eps_{cold} for a piecewise
* polytropic EOS.
*
* Outputs : P_cold - cold pressure (supports SPEOS and PPEOS)
* : eps_cold - cold specific internal energy (supports SPEOS and PPEOS)
* : dPcold_drho - derivative of P_cold (supports SPEOS and PPEOS)
* : eps_th - thermal specific internal energy (supports SPEOS and PPEOS)
* : h - enthalpy (supports SPEOS and PPEOS)
* : Gamma_cold - cold polytropic Gamma (supports SPEOS and PPEOS)
*
* SPEOS: Single-Polytrope Equation of State
* PPEOS: Piecewise Polytrope Equation of State
*/
static inline void compute_P_cold__eps_cold__dPcold_drho__eps_th__h__Gamma_cold(CCTK_REAL *U, eos_struct &eos, CCTK_REAL Gamma_th,
CCTK_REAL &P_cold,CCTK_REAL &eps_cold,CCTK_REAL &dPcold_drho,CCTK_REAL &eps_th,CCTK_REAL &h,
CCTK_REAL &Gamma_cold) {
// This code handles equations of state of the form defined
// in Eqs 13-16 in http://arxiv.org/pdf/0802.0200.pdf
if(U[RHOB]==0) {
P_cold = 0.0;
eps_cold = 0.0;
dPcold_drho = 0.0;
eps_th = 0.0;
h = 0.0;
Gamma_cold = eos.Gamma_ppoly_tab[0];
return;
}
int polytropic_index = find_polytropic_K_and_Gamma_index(eos, U[RHOB]);
compute_P_cold__eps_cold(eos,U[RHOB], P_cold,eps_cold);
CCTK_REAL Gamma_ppoly_tab = eos.Gamma_ppoly_tab[polytropic_index];
// Set auxiliary variable rho_b^{-1}
CCTK_REAL U_RHOB_inv = 1.0/U[RHOB];
// Next compute dP/drho = Gamma * P / rho
dPcold_drho = Gamma_ppoly_tab*P_cold*U_RHOB_inv;
// Then we compute eps_th, h, and set Gamma_cold = Gamma_ppoly_tab[j].
eps_th = (U[PRESSURE] - P_cold)/(Gamma_th-1.0)*U_RHOB_inv;
h = 1.0 + eps_cold + eps_th + U[PRESSURE]*U_RHOB_inv;
Gamma_cold = Gamma_ppoly_tab;
}
```
<a id='ppeos__c_code__print_eos_table'></a>
## Step 5.e: The `print_EOS_table()` function \[Back to [top](#toc)\]
$$\label{ppeos__c_code__print_eos_table}$$
This function, which is called for diagnostic purposes and for user convenience, prints out the most relevant EOS table components to the user at the *beginning* of the run. Notice that the function call happens only once, in the [`driver_conserv_to_prims.C`](Tutorial-IllinoisGRMHD__the_conservative_to_primitive_algorithm.ipynb) file.
```
%%writefile -a $Piecewise_Polytrope_EOS__C
/* Function : print_EOS_table()
* Authors : Leo Werneck
* Description : Prints out the EOS table, for diagnostic purposes
*
* Dependencies: initialize_EOS_struct_from_input()
*
* Inputs : eos - a struct containing the following
* relevant quantities:
* : neos - number of polytropic EOSs used
* : rho_ppoly_tab - array of rho values that determine
* the polytropic EOS to be used.
* : Gamma_ppoly_tab - array of Gamma_cold values to be
* used in each polytropic EOS.
*
* Outputs : CCTK_VInfo string with the EOS table used by IllinoisGRMHD
*/
static inline void print_EOS_table( eos_struct eos ) {
/* Start by printint a header t the table */
#ifndef ENABLE_STANDALONE_IGM_C2P_SOLVER
CCTK_VInfo(CCTK_THORNSTRING,"\n"
#else
printf("\n"
#endif
".--------------------------------------------.\n"
"| EOS Table |\n"
".--------------------------------------------.");
printf("| rho_ppoly_tab[j] |\n"
".--------------------------------------------.\n");
/* Adjust the maximum index of rhob to
* allow for single polytropes as well
*/
int max_rho_index;
if( eos.neos==1 ) {
max_rho_index = 0;
}
else {
max_rho_index = eos.neos-2;
}
/* Print out rho_pppoly_tab */
for(int jj=0; jj<=max_rho_index; jj++) {
printf("| rho_ppoly_tab[%d] = %.15e |\n",jj,eos.rho_ppoly_tab[jj]);
if(jj == eos.neos-2) {
printf(".--------------------------------------------.\n");
}
}
/* Print out Gamma_ppoly_tab */
printf("| Gamma_ppoly_tab[j] |\n"
".--------------------------------------------.\n");
for(int jj=0; jj<=eos.neos-1; jj++) {
printf("| Gamma_ppoly_tab[%d] = %.15e |\n",jj,eos.Gamma_ppoly_tab[jj]);
if(jj == eos.neos-1) {
printf(".--------------------------------------------.\n");
}
}
/* Print out K_ppoly_tab */
printf("| K_ppoly_tab[j] |\n"
".--------------------------------------------.\n");
for(int jj=0; jj<=eos.neos-1; jj++) {
printf("| K_ppoly_tab[%d] = %.15e |\n",jj,eos.K_ppoly_tab[jj]);
if(jj == eos.neos-1) {
printf(".--------------------------------------------.\n\n");
}
}
}
#endif // ILLINOISGRMHD_EOS_FUNCTS_C_
```
<a id='latex_pdf_output'></a>
# Step n: Output this notebook to $\LaTeX$-formatted PDF file \[Back to [top](#toc)\]
$$\label{latex_pdf_output}$$
The following code cell converts this Jupyter notebook into a proper, clickable $\LaTeX$-formatted PDF file. After the cell is successfully run, the generated PDF may be found in the root NRPy+ tutorial directory, with filename
[Tutorial-IllinoisGRMHD__piecewise_polytrope_tests.pdf](Tutorial-IllinoisGRMHD__piecewise_polytrope_tests.pdf) (Note that clicking on this link may not work; you may need to open the PDF file through another means).
```
import os
nrpy_dir_path = os.path.join("..","..")
latex_nrpy_style_path = os.path.join(nrpy_dir_path,"latex_nrpy_style.tplx")
#!jupyter nbconvert --to latex --template $latex_nrpy_style_path --log-level='WARN' Tutorial-IllinoisGRMHD__Piecewise_Polytrope_EOS.ipynb
#!pdflatex -interaction=batchmode Tutorial-IllinoisGRMHD__Piecewise_Polytrope_EOS.tex
#!pdflatex -interaction=batchmode Tutorial-IllinoisGRMHD__Piecewise_Polytrope_EOS.tex
#!pdflatex -interaction=batchmode Tutorial-IllinoisGRMHD__Piecewise_Polytrope_EOS.tex
!rm -f Tut*.out Tut*.aux Tut*.log
```
| github_jupyter |
##### Copyright 2019 The TensorFlow Hub Authors.
Licensed under the Apache License, Version 2.0 (the "License");
```
# Copyright 2018 The TensorFlow Hub Authors. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ==============================================================================
```
# 最近傍とテキスト埋め込みによるセマンティック検索
<table class="tfo-notebook-buttons" align="left">
<td> <a target="_blank" href="https://www.tensorflow.org/hub/tutorials/semantic_approximate_nearest_neighbors"><img src="https://www.tensorflow.org/images/tf_logo_32px.png">TensorFlow.org で表示</a> </td>
<td> <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/ja/hub/tutorials/semantic_approximate_nearest_neighbors.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png">Google Colab で実行</a> </td>
<td><a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/ja/hub/tutorials/semantic_approximate_nearest_neighbors.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png">GitHub で表示</a></td>
<td> <a href="https://storage.googleapis.com/tensorflow_docs/docs-l10n/site/ja/hub/tutorials/semantic_approximate_nearest_neighbors.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png">ノートブックをダウンロード</a> </td>
<td> <a href="https://tfhub.dev/google/universal-sentence-encoder/2"><img src="https://www.tensorflow.org/images/hub_logo_32px.png">TF Hub モデルを参照</a> </td>
</table>
このチュートリアルでは、[TensorFlow Hub](https://tfhub.dev)(TF-Hub)が提供する入力データから埋め込みを生成し、抽出された埋め込みを使用して最近傍(ANN)インデックスを構築する方法を説明します。構築されたインデックスは、リアルタイムに類似性の一致と検索を行うために使用できます。
大規模なコーパスのデータを取り扱う場合、特定のクエリに対して最も類似するアイテムをリアルタイムで見つけるために、レポジトリ全体をスキャンして完全一致を行うというのは、効率的ではありません。そのため、おおよその類似性一致アルゴリズムを使用することで、正確な最近傍の一致を見つける際の精度を少しだけ犠牲にし、速度を大幅に向上させることができます。
このチュートリアルでは、ニュースの見出しのコーパスに対してリアルタイムテキスト検索を行い、クエリに最も類似する見出しを見つけ出す例を示します。この検索はキーワード検索とは異なり、テキスト埋め込みにエンコードされた意味的類似性をキャプチャします。
このチュートリアルの手順は次のとおりです。
1. サンプルデータをダウンロードする。
2. TF-Hub モジュールを使用して、データの埋め込みを生成する。
3. 埋め込みの ANN インデックスを構築する。
4. インデックスを使って、類似性の一致を実施する。
TF-Hub モジュールから埋め込みを生成するには、[TensorFlow Transform](https://beam.apache.org/documentation/programming-guide/)(TF-Transform)を使った [Apache Beam](https://www.tensorflow.org/tfx/tutorials/transform/simple) を使用します。また、最近傍インデックスの構築には、Spotify の [ANNOY](https://github.com/spotify/annoy) ライブラリを使用します。ANN フレームワークのベンチマークは、こちらの [Github リポジトリ](https://github.com/erikbern/ann-benchmarks)をご覧ください。
このチュートリアルでは TensorFlow 1.0 を使用し、TF1 の [Hub モジュール](https://www.tensorflow.org/hub/tf1_hub_module)のみと連携します。更新版は、[このチュートリアルの TF2 バージョン](https://github.com/tensorflow/hub/blob/master/examples/colab/tf2_semantic_approximate_nearest_neighbors.ipynb)をご覧ください。
## セットアップ
必要なライブラリをインストールします。
```
!pip install -q apache_beam
!pip install -q 'scikit_learn~=0.23.0' # For gaussian_random_matrix.
!pip install -q annoy
```
必要なライブラリをインポートします。
```
import os
import sys
import pathlib
import pickle
from collections import namedtuple
from datetime import datetime
import numpy as np
import apache_beam as beam
import annoy
from sklearn.random_projection import gaussian_random_matrix
import tensorflow.compat.v1 as tf
import tensorflow_hub as hub
# TFT needs to be installed afterwards
!pip install -q tensorflow_transform==0.24
import tensorflow_transform as tft
import tensorflow_transform.beam as tft_beam
print('TF version: {}'.format(tf.__version__))
print('TF-Hub version: {}'.format(hub.__version__))
print('TF-Transform version: {}'.format(tft.__version__))
print('Apache Beam version: {}'.format(beam.__version__))
```
## 1. サンプルデータをダウンロードする
[A Million News Headlines](https://dataverse.harvard.edu/dataset.xhtml?persistentId=doi:10.7910/DVN/SYBGZL#) データセットには、15 年にわたって発行されたニュースの見出しが含まれます。出典は、有名なオーストラリア放送協会(ABC)です。このニュースデータセットは、2003 年の始めから 2017 年の終わりまでの特筆すべき世界的なイベントについて、オーストラリアにより焦点を当てた記録が含まれます。
**形式**: 1)発行日と 2)見出しのテキストの 2 列をタブ区切りにしたデータ。このチュートリアルで関心があるのは、見出しのテキストのみです。
```
!wget 'https://dataverse.harvard.edu/api/access/datafile/3450625?format=tab&gbrecs=true' -O raw.tsv
!wc -l raw.tsv
!head raw.tsv
```
単純化するため、見出しのテキストのみを維持し、発行日は削除します。
```
!rm -r corpus
!mkdir corpus
with open('corpus/text.txt', 'w') as out_file:
with open('raw.tsv', 'r') as in_file:
for line in in_file:
headline = line.split('\t')[1].strip().strip('"')
out_file.write(headline+"\n")
!tail corpus/text.txt
```
## TF-Hub モジュールを読み込むためのヘルパー関数
```
def load_module(module_url):
embed_module = hub.Module(module_url)
placeholder = tf.placeholder(dtype=tf.string)
embed = embed_module(placeholder)
session = tf.Session()
session.run([tf.global_variables_initializer(), tf.tables_initializer()])
print('TF-Hub module is loaded.')
def _embeddings_fn(sentences):
computed_embeddings = session.run(
embed, feed_dict={placeholder: sentences})
return computed_embeddings
return _embeddings_fn
```
## 2. データの埋め込みを生成する
このチュートリアルでは、[ユニバーサルセンテンスエンコーダ](https://tfhub.dev/google/universal-sentence-encoder/2)を使用して、見出しデータの埋め込みを生成します。その後で、文章レベルの意味の類似性を計算するために、文章埋め込みを簡単に使用することが可能となります。埋め込み生成プロセスは、Apache Beam と TF-Transform を使用して実行します。
### 埋め込み抽出メソッド
```
encoder = None
def embed_text(text, module_url, random_projection_matrix):
# Beam will run this function in different processes that need to
# import hub and load embed_fn (if not previously loaded)
global encoder
if not encoder:
encoder = hub.Module(module_url)
embedding = encoder(text)
if random_projection_matrix is not None:
# Perform random projection for the embedding
embedding = tf.matmul(
embedding, tf.cast(random_projection_matrix, embedding.dtype))
return embedding
```
### TFT preprocess_fn メソッドの作成
```
def make_preprocess_fn(module_url, random_projection_matrix=None):
'''Makes a tft preprocess_fn'''
def _preprocess_fn(input_features):
'''tft preprocess_fn'''
text = input_features['text']
# Generate the embedding for the input text
embedding = embed_text(text, module_url, random_projection_matrix)
output_features = {
'text': text,
'embedding': embedding
}
return output_features
return _preprocess_fn
```
### データセットのメタデータの作成
```
def create_metadata():
'''Creates metadata for the raw data'''
from tensorflow_transform.tf_metadata import dataset_metadata
from tensorflow_transform.tf_metadata import schema_utils
feature_spec = {'text': tf.FixedLenFeature([], dtype=tf.string)}
schema = schema_utils.schema_from_feature_spec(feature_spec)
metadata = dataset_metadata.DatasetMetadata(schema)
return metadata
```
### Beam パイプライン
```
def run_hub2emb(args):
'''Runs the embedding generation pipeline'''
options = beam.options.pipeline_options.PipelineOptions(**args)
args = namedtuple("options", args.keys())(*args.values())
raw_metadata = create_metadata()
converter = tft.coders.CsvCoder(
column_names=['text'], schema=raw_metadata.schema)
with beam.Pipeline(args.runner, options=options) as pipeline:
with tft_beam.Context(args.temporary_dir):
# Read the sentences from the input file
sentences = (
pipeline
| 'Read sentences from files' >> beam.io.ReadFromText(
file_pattern=args.data_dir)
| 'Convert to dictionary' >> beam.Map(converter.decode)
)
sentences_dataset = (sentences, raw_metadata)
preprocess_fn = make_preprocess_fn(args.module_url, args.random_projection_matrix)
# Generate the embeddings for the sentence using the TF-Hub module
embeddings_dataset, _ = (
sentences_dataset
| 'Extract embeddings' >> tft_beam.AnalyzeAndTransformDataset(preprocess_fn)
)
embeddings, transformed_metadata = embeddings_dataset
# Write the embeddings to TFRecords files
embeddings | 'Write embeddings to TFRecords' >> beam.io.tfrecordio.WriteToTFRecord(
file_path_prefix='{}/emb'.format(args.output_dir),
file_name_suffix='.tfrecords',
coder=tft.coders.ExampleProtoCoder(transformed_metadata.schema))
```
### ランダムプロジェクションの重み行列を生成する
[ランダムプロジェクション](https://en.wikipedia.org/wiki/Random_projection)は、ユークリッド空間に存在する一連の点の次元を縮小するために使用される、単純でありながら高性能のテクニックです。理論的背景については、[Johnson-Lindenstrauss の補題](https://en.wikipedia.org/wiki/Johnson%E2%80%93Lindenstrauss_lemma)をご覧ください。
ランダムプロジェクションを使用して埋め込みの次元を縮小するということは、ANN インデックスの構築とクエリに必要となる時間を短縮できるということです。
このチュートリアルでは、[Scikit-learn](https://en.wikipedia.org/wiki/Random_projection#Gaussian_random_projection) ライブラリの[ガウスランダムプロジェクションを使用します。](https://scikit-learn.org/stable/modules/random_projection.html#gaussian-random-projection)
```
def generate_random_projection_weights(original_dim, projected_dim):
random_projection_matrix = None
if projected_dim and original_dim > projected_dim:
random_projection_matrix = gaussian_random_matrix(
n_components=projected_dim, n_features=original_dim).T
print("A Gaussian random weight matrix was creates with shape of {}".format(random_projection_matrix.shape))
print('Storing random projection matrix to disk...')
with open('random_projection_matrix', 'wb') as handle:
pickle.dump(random_projection_matrix,
handle, protocol=pickle.HIGHEST_PROTOCOL)
return random_projection_matrix
```
### パラメータの設定
ランダムプロジェクションを使用せずに、元の埋め込み空間を使用してインデックスを構築する場合は、`projected_dim` パラメータを `None` に設定します。これにより、高次元埋め込みのインデックス作成ステップが減速することに注意してください。
```
module_url = 'https://tfhub.dev/google/universal-sentence-encoder/2' #@param {type:"string"}
projected_dim = 64 #@param {type:"number"}
```
### パイプラインの実行
```
import tempfile
output_dir = pathlib.Path(tempfile.mkdtemp())
temporary_dir = pathlib.Path(tempfile.mkdtemp())
g = tf.Graph()
with g.as_default():
original_dim = load_module(module_url)(['']).shape[1]
random_projection_matrix = None
if projected_dim:
random_projection_matrix = generate_random_projection_weights(
original_dim, projected_dim)
args = {
'job_name': 'hub2emb-{}'.format(datetime.utcnow().strftime('%y%m%d-%H%M%S')),
'runner': 'DirectRunner',
'batch_size': 1024,
'data_dir': 'corpus/*.txt',
'output_dir': output_dir,
'temporary_dir': temporary_dir,
'module_url': module_url,
'random_projection_matrix': random_projection_matrix,
}
print("Pipeline args are set.")
args
!rm -r {output_dir}
!rm -r {temporary_dir}
print("Running pipeline...")
%time run_hub2emb(args)
print("Pipeline is done.")
!ls {output_dir}
```
生成された埋め込みをいくつか読み取ります。
```
import itertools
embed_file = os.path.join(output_dir, 'emb-00000-of-00001.tfrecords')
sample = 5
record_iterator = tf.io.tf_record_iterator(path=embed_file)
for string_record in itertools.islice(record_iterator, sample):
example = tf.train.Example()
example.ParseFromString(string_record)
text = example.features.feature['text'].bytes_list.value
embedding = np.array(example.features.feature['embedding'].float_list.value)
print("Embedding dimensions: {}".format(embedding.shape[0]))
print("{}: {}".format(text, embedding[:10]))
```
## 3. 埋め込みの ANN インデックスを構築する
[ANNOY](https://github.com/spotify/annoy)(Approximate Nearest Neighbors Oh Yeah)は、特定のクエリ点に近い空間内のポイントを検索するための、Python バインディングを使った C++ ライブラリです。メモリにマッピングされた、大規模な読み取り専用ファイルベースのデータ構造も作成します。[Spotify](https://www.spotify.com) が構築したもので、おすすめの音楽に使用されています。
```
def build_index(embedding_files_pattern, index_filename, vector_length,
metric='angular', num_trees=100):
'''Builds an ANNOY index'''
annoy_index = annoy.AnnoyIndex(vector_length, metric=metric)
# Mapping between the item and its identifier in the index
mapping = {}
embed_files = tf.gfile.Glob(embedding_files_pattern)
print('Found {} embedding file(s).'.format(len(embed_files)))
item_counter = 0
for f, embed_file in enumerate(embed_files):
print('Loading embeddings in file {} of {}...'.format(
f+1, len(embed_files)))
record_iterator = tf.io.tf_record_iterator(
path=embed_file)
for string_record in record_iterator:
example = tf.train.Example()
example.ParseFromString(string_record)
text = example.features.feature['text'].bytes_list.value[0].decode("utf-8")
mapping[item_counter] = text
embedding = np.array(
example.features.feature['embedding'].float_list.value)
annoy_index.add_item(item_counter, embedding)
item_counter += 1
if item_counter % 100000 == 0:
print('{} items loaded to the index'.format(item_counter))
print('A total of {} items added to the index'.format(item_counter))
print('Building the index with {} trees...'.format(num_trees))
annoy_index.build(n_trees=num_trees)
print('Index is successfully built.')
print('Saving index to disk...')
annoy_index.save(index_filename)
print('Index is saved to disk.')
print("Index file size: {} GB".format(
round(os.path.getsize(index_filename) / float(1024 ** 3), 2)))
annoy_index.unload()
print('Saving mapping to disk...')
with open(index_filename + '.mapping', 'wb') as handle:
pickle.dump(mapping, handle, protocol=pickle.HIGHEST_PROTOCOL)
print('Mapping is saved to disk.')
print("Mapping file size: {} MB".format(
round(os.path.getsize(index_filename + '.mapping') / float(1024 ** 2), 2)))
embedding_files = "{}/emb-*.tfrecords".format(output_dir)
embedding_dimension = projected_dim
index_filename = "index"
!rm {index_filename}
!rm {index_filename}.mapping
%time build_index(embedding_files, index_filename, embedding_dimension)
!ls
```
## 4. インデックスを使って、類似性の一致を実施する
ANN インデックスを使用して、入力クエリに意味的に近いニュースの見出しを検索できるようになりました。
### インデックスとマッピングファイルを読み込む
```
index = annoy.AnnoyIndex(embedding_dimension)
index.load(index_filename, prefault=True)
print('Annoy index is loaded.')
with open(index_filename + '.mapping', 'rb') as handle:
mapping = pickle.load(handle)
print('Mapping file is loaded.')
```
### 類似性の一致メソッド
```
def find_similar_items(embedding, num_matches=5):
'''Finds similar items to a given embedding in the ANN index'''
ids = index.get_nns_by_vector(
embedding, num_matches, search_k=-1, include_distances=False)
items = [mapping[i] for i in ids]
return items
```
### 特定のクエリから埋め込みを抽出する
```
# Load the TF-Hub module
print("Loading the TF-Hub module...")
g = tf.Graph()
with g.as_default():
embed_fn = load_module(module_url)
print("TF-Hub module is loaded.")
random_projection_matrix = None
if os.path.exists('random_projection_matrix'):
print("Loading random projection matrix...")
with open('random_projection_matrix', 'rb') as handle:
random_projection_matrix = pickle.load(handle)
print('random projection matrix is loaded.')
def extract_embeddings(query):
'''Generates the embedding for the query'''
query_embedding = embed_fn([query])[0]
if random_projection_matrix is not None:
query_embedding = query_embedding.dot(random_projection_matrix)
return query_embedding
extract_embeddings("Hello Machine Learning!")[:10]
```
### クエリを入力して、類似性の最も高いアイテムを検索する
```
#@title { run: "auto" }
query = "confronting global challenges" #@param {type:"string"}
print("Generating embedding for the query...")
%time query_embedding = extract_embeddings(query)
print("")
print("Finding relevant items in the index...")
%time items = find_similar_items(query_embedding, 10)
print("")
print("Results:")
print("=========")
for item in items:
print(item)
```
## 今後の学習
[tensorflow.org/hub](https://www.tensorflow.org/) では、TensorFlow についてさらに学習し、TF-Hub API ドキュメントを確認することができます。また、[tfhub.dev](https://www.tensorflow.org/hub/) では、その他のテキスト埋め込みモジュールや画像特徴量ベクトルモジュールなど、利用可能な TensorFlow Hub モジュールを検索することができます。
さらに、Google の [Machine Learning Crash Course](https://developers.google.com/machine-learning/crash-course/) もご覧ください。機械学習の実用的な導入をテンポよく学習できます。
| github_jupyter |
```
%matplotlib inline
from __future__ import division
import math
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
pd.set_option('display.max_columns', 150)
pd.set_option('display.max_rows', 25)
pd.set_option('display.width', 100)
pd.set_option('display.max_colwidth', 1024)
#wget http://www.mathstat.strath.ac.uk/outreach/nessie/datasets/whiskies.txt
df = pd.read_csv('whiskies.txt')
df.head(1)
import sklearn.datasets
import sklearn.metrics as metrics
from sklearn.decomposition import PCA
from sklearn.cluster import KMeans, DBSCAN, MeanShift
from sklearn.preprocessing import StandardScaler
X = df.drop(['RowID', 'Distillery', 'Postcode', ' Latitude', ' Longitude'], axis=1)
X.describe()
# X_std = StandardScaler().fit_transform(X)
# pd.DataFrame(X_std, columns=X.columns).describe()
n_components = 5
pca = PCA(n_components=n_components)
X_pca = pca.fit_transform(X)
X.shape, X_pca.shape
weights = np.round(pca.components_, 3)
ev = np.round(pca.explained_variance_ratio_, 3)
ev
pca_df = pd.DataFrame(weights, columns=X.columns)
pca_df
import warnings
warnings.simplefilter(action = "ignore")
component_names = []
pca_df_t = pca_df.T
for col in pca_df_t:
component = pca_df_t[col]
order = component.abs()#.order(ascending=False)
top = order.head(3)
component_name = [name if component[name] > 0 else ('neg-' + name) for name in top.index]
component_names.append('/'.join(component_name))
pca_df.index = component_names
pca_df
inertia = [np.NaN]
for i in range(1,20):
kmeans = KMeans(n_clusters=i)
kmeans.fit(X)
inertia.append(kmeans.inertia_)
ax = plt.subplot(111)
ax.plot(inertia, 'o-')
ax.set_ylabel('inertia')
ax.set_xlabel('# clusters')
n_clusters = 4
kmeans = KMeans(n_clusters=n_clusters)
kmeans.fit(X)
X['cluster'] = kmeans.labels_
X.cluster.value_counts()
X_pca_clustered = np.insert(X_pca, n_components, values=kmeans.labels_, axis=1)
pca_cluster_df = pd.DataFrame(X_pca_clustered, columns=component_names + ['cluster'])
g = sns.PairGrid(pca_cluster_df, hue='cluster', vars=component_names, size=3)
g.map_diag(plt.hist)
g.map_offdiag(plt.scatter)
g.add_legend()
dist = sklearn.metrics.pairwise.euclidean_distances(X)
sim = sklearn.metrics.pairwise.cosine_similarity(X)
dist_df = pd.DataFrame(dist, columns=df.Distillery, index=df.Distillery)
sim_df = pd.DataFrame(sim, columns=df.Distillery, index=df.Distillery)
cluster_s = X.cluster
cluster_s.index = df.Distillery
sim_df['Ardbeg']#.order(ascending=False)
features_df = df.ix[:,1:13].set_index('Distillery')
#Add the cluster info to the features DataFrame so we only have to persist one file for both.
features_df['cluster'] = cluster_s
features_df.to_pickle('features.dataframe')
sim_df.to_pickle('sims.dataframe')
```
| github_jupyter |
## Introduction to Classical Natural Language Processing ##
This notebook is a hands-on introduction to Classical NLP, that is, non-deep learning techniques of NLP. It is designed to be used with the edX course on Natural Language Processing, from Microsoft.
The topics covered align with the NLP tasks related to the various stages of the NLP pipeline: text processing, text exploration, building features, and application level tasks.
## 1. Introduction ##
** 1.1 NLTK Setup **
- NLTK is included with the Anaconda Distribution of Python, or can be downloaded directly from nltk.org.
- Once NLTK is installed, the text data files (corpora) should be downloaded. See the following cell to start the download.
```
import nltk
# uncomment the line below to download NLTK resources the first time NLTK is used and RUN this cell.
# when the "NLTK Downloader" dialog appears (takes 10-20 seconds), click on the "download" button
#nltk.download()
```
** 1.2 Crash Course in Regular Expressions **
If you are new to using regular expressions, or would like a quick refresher, you can study the examples
and resulting output in the code cell below.
Here is a cheat sheet for the SEARCH BASICS (code examples follow below):
Operator Meaning Example Example meaning
+ one or more a+ look for 1 or more "a" characters
* zero or more a* look for 0 or more "a" characters
? optional a? look for 0 or 1 "a" characters
[] choose 1 [abc] look for "a" or "b" or "c"
[-] range [a-z] look for any character between "a" and "z"
[^] not [^a] look for character that is not "a"
() grouping (a-z)+ look for one of more occurences of chars between "a" and "z"
(|) or operator (ey|ax) look for strings "ey" or "ax"
ab follow ab look for character "a" followed by character "b"
^ start ^a look for character "a" at start of string/line
$ end a$ look for character "a" at end of string/line
\s whitespace \sa look for whitespace character followed by "a"
. any character a.b look for "a" followed by any char followed by "b"
Common Uses:
- re.search finds first matching object
- re.findall returns all matching objects
- re.sub replaces matches with replacement string
```
import re
# search for single char
re.search(r"x", "this is an extra helping")
# search for single char
re.search(r"x", "this is an extra helping").group(0) # gives easier-to-read output
# find all occurences of any character between "a" and "z"
re.findall(r"[a-z]", "$34.33 cash.")
# find all occurences of either "name:" or "phone:"
re.findall(r"(name|phone):", "My name: Joe, my phone: (312)555-1212")
# find "lion", "lions" or "Lion", or "Lions"
re.findall(r"([Ll]ion)s?", "Give it to the Lions or the lion.")
# replace allll lowercase letters with "x"
re.sub("[a-z]", "x", "Hey. I know this regex stuff...")
```
## 2. Text Processing ##
This section introduces some of the tasks and techniques used to acquire, clean, and normalize the text data.
** 2.1 Data Acquisition **
Issues:
- how do I find the data I need?
- is it already in digital form, or will it need OCR?
- how much will it cost?
- will it be updated/expanded over time? More costs?
- (if CUSTOMER DATA), do I have the legal / privacy rights needed to use the data in the way I need for my application?
- do I have the safeguards needed to securely store the data?
```
import nltk
# shows how to access one of the gutenberg books included in NLTK
print("gutenberg book ids=", nltk.corpus.gutenberg.fileids())
# load words from "Alice in Wonderland"
alice = nltk.corpus.gutenberg.words("carroll-alice.txt")
print("len(alice)=", len(alice))
print(alice[:100])
# load words from "Monty Python and the Holy Grail"
grail = nltk.corpus.webtext.words("grail.txt")
print("len(grail)=", len(grail))
print(grail[:100])
```
** 2.2 Plain Text Extraction **
If your text data lives in a non-plain text file (WORD, POWERPOINT, PDF, HTML, etc.), you will need to use a “filter” to extract the plain text from the file.
Python has a number of libraries to extract plain text from popular file formats, but they are take searching and supporting code to use.
** 2.3 Word and Sentence Segmentation (Tokenization) **
Word Segmentation Issues:
- Some languages don’t white space characters
- Words with hyphens or apostrophes (Who’s at the drive-in?)
- Numbers, currency, percentages, dates, times (04/01/2018, $55,000.00)
- Ellipses, special characters
Sentence Segmentation Issues:
- Quoted speech within a sentence
- Abbreviations with periods (The Ph.D. was D.O.A)
Tokenization Techniques
- Perl script (50 lines) with RegEx (Grefenstette, 1999)
- maxmatch Algorithm:
themanranafterit -> the man ran after it
thetabledownthere -> theta bled own there (Palmer, 2000)
```
# code example: simple version of maxmatch algorithm for tokenization (word segmentation)
def tokenize(str, dict):
s = 0
words = []
while (s < len(str)):
found = False
# find biggest word in dict that matches str[s:xxx]
for word in dict:
lw = len(word)
if (str[s:s+lw] == word):
words.append(word)
s += lw
found = True
break
if (not found):
words.append(str[s])
s += 1
print(words)
#return words
# small dictionary of known words, longest words first
dict = ["before", "table", "theta", "after", "where", "there", "bled", "said", "lead", "man", "her", "own", "the", "ran", "it"]
# this algorithm is designed to work with languages that don't have whitespace characters
# so simulate that in our test
tokenize("themanranafterit", dict) # works!
tokenize("thetabledownthere", dict) # fails!
# NLTK example: WORD segmentation
nltk.word_tokenize("the man, he ran after it's $3.23 dog on 03/23/2016.")
# NLTK example: SENTENCE segmentation
nltk.sent_tokenize('The man ran after it. The table down there? Yes, down there!')
```
** 2.4 Stopword Removal **
Stopwords are common words that are "not interesting" for the app/task at hand.
Easy part – removing words that appear in list.
Tricky part – what to use for stop words? App-dependent. Standard lists, high-frequency words in your text, …
```
# code example: simple algorithm for removing stopwords
stoppers = "a is of the this".split()
def removeStopWords(stopWords, txt):
newtxt = ' '.join([word for word in txt.split() if word not in stopWords])
return newtxt
removeStopWords(stoppers, "this is a test of the stop word removal code.")
# NLTK example: removing stopwords
from nltk.corpus import stopwords
stops = stopwords.words("English")
print("len(stops)=", len(stops))
removeStopWords(stops, "this is a test of the stop word removal code.")
```
** 2.5 Case Removal **
Case removal is part of a larger task called *Text Normalization*, which includes:
- case removal
- stemming (covered in next section)
Goal of Case removal – converting all text to, for example, lower case
```
# code example: case removal
str = 'The man ran after it. The table down there? Yes, down there!'
str.lower()
```
** 2.6 Stemming **
Goal of Stemming:
– stripping off endings and other pieces, called AFFIXES
– for English, this is prefixes and suffixes.
- convert word to its base word, called the LEMMA / STEM (e.g., foxes -> fox)
Porter Stemmer
- 100+ cascading “rewrite” rules
ational -> ate (e.g., relational -> relate)
ing -> <null> (e.g., playing -> play)
sess -> ss (e.g., grasses -> grass)
```
# NLTK example: stemming
def stem_with_porter(words):
porter = nltk.PorterStemmer()
new_words = [porter.stem(w) for w in words]
return new_words
def stem_with_lancaster(words):
porter = nltk.LancasterStemmer()
new_words = [porter.stem(w) for w in words]
return new_words
str = "Please don't unbuckle your seat-belt while I am driving, he said"
print("porter:", stem_with_porter(str.split()))
print()
print("lancaster:", stem_with_lancaster(str.split()))
```
## 3. Text Exploration ##
** 3.1 Frequency Analysis **
- Frequency Analysis
- Letter
- Word
- Bigrams
- Plots
```
# NLTK example: frequence analysis
import nltk
from nltk.corpus import gutenberg
from nltk.probability import FreqDist
# get raw text from "Sense and Sensibility" by Jane Austen
raw = gutenberg.raw("austen-sense.txt")
fd_letters = FreqDist(raw)
words = gutenberg.words("austen-sense.txt")
fd_words = FreqDistist(words)
sas = nltk.Text(words)
# these 2 lines let us size the freq dist plot
import matplotlib.pyplot as plt
plt.figure(figsize=(20, 5))
# frequency plot for letters from SAS
fd_letters.plot(100)
# these 2 lines let us size the freq dist plot
import matplotlib.pyplot as plt
plt.figure(figsize=(20, 5))
# frequency plot for words from SAS
fd_words.plot(50)
```
** 3.2 Collocations **
These are interesting word pairs, usually formed by the most common bigrams. Bigrams are collections of word pairs that occur together in the text.
```
# let's look at collocations for our "Sense and Sensibility" text
sas.collocations()
```
Nice!
Now we are getting a feel for the language and subjects of the text.
# ** 3.3 Long words **
Sometimes looking at the long words in a text can be revealing. Let's try it on sas.
```
# let's look at long words in the text
longWords = [w for w in set(words) if len(w) > 13]
longWords[:15]
```
** 3.3 Concordance Views **
Concordance views, also called Keywords in Context (KWIC), show the specifed word with the words that surround it in text. These views can be helpful in understaning how the words are being used in the text.
```
# Let's try looking at some of these recent words in a Concordance view
sas.concordance("affectionately")
print()
sas.concordance("correspondence")
print()
sas.concordance("dare")
print()
```
** 3.4 Other Exploration Task/Views **
```
# look at words similiar to a word
sas.similar("affection")
# these 2 lines let us size the freq dist plot
import matplotlib.pyplot as plt
plt.figure(figsize=(15, 4))
# look at words as they appear over time in the book/document
sas.dispersion_plot(["sense", "love", "heart", "listen", "man", "woman"])
```
## 4. Building Features ##
** 4.1 Bag-of-Words (BOW) **
One of the simplest features when dealing with multiple texts (like multiple documents, or multiple sentences within a document), is called Bag-of-Words. It builds a vocabular from each word in the set of texts, and then a feature for each word, indicate the presence/absence of that word within each text. Sometimes, the count of the word is used in place of a presence flag.
A common way to represent a set of features like this is called a One-Hot vector. For example, lets say our vocabular from our set of texts is:
today, here, I, a, fine, sun, moon, bird, saw
The sentence we want to build a BOW for is:
I saw a bird today.
Using a 1/0 for each word in the vocabulary, our BOW encoded as a one-hot vector would be:
1 0 1 1 0 0 1 1
** 4.2 N-Grams **
N-grams represent the sequence of N words that are found in a text. They are commonly used as a model of the text language since they represent the frequence of words/phrases appearing in the text.
Common types of N-grams:
unigrams - these are the set of single words appearing in the text
bigrams - these are the set of word pairs, like "good day" or "big deal", from the text
trigrams - these are the set of word triples, like "really good food", from the text
To build bigrams for a text, you need to extract all possible word pairs from the text and count how many times each pair occurs. Then, you can use the top N words (1000), or the top percent (50%) as your language model.
** 4.3 Morphological Parsing **
**Goal**: convert input word into its morphological parts. For example: “geese” would return goose + N + PL
Morphological Parsing:
geese -> goose + N + PL
caught -> catch + V + PastPart
Morphological parsing is related to stemming, but instead of mapping the word variants to a stem word, it labels
the stem word and its affixes.
Morphological parsing, even for English, is quite involved
** 4.4 TD/IDF **
TD/IDF stands for Term Document Inverse Document Frequency. "Term" here can be thought of as a word. This is a measure of the relative importance of a word within a document, in the context of multiple documents.
We start with the TD part - this is simply a normalized frequency of the word in the document:
- (word count in document) / (total words in document)
The IDF is a weighting of the uniquess of the word across all of the documents. Here is the complete formula of TD/IDF:
- td_idf(t,d) = wc(t,d)/wc(d) / dc(t)/dc()
where:
- wc(t,d) = # of occurrences of term t in doc d
- wc(d) = # of words in doc d
- dc(t) = # of docs that contain at least 1 occurrence of term t
- dc() = # of docs in collection
** 4.5 Word Sense Disambiguation (WSD) **
Related to POS tagging, WSD is use to distingish between difference senses of a word. Each sense of the word
uses the same POS tag, but means something different. For example:
- she served the King
- she served the ball
- he took his money to the bank
- he took his canoe to the bank
- I play bass guitar
- I fish for bass
** 4.6 Anaphora Resolution ** \
Examples:
- Sam and Bill left with the toys. They were later found.
Who does "they" refer to in the above sentence? Sam and Bill, or the toys?
** 4.7 Part-of-speech (POS) Tagging **
- Verb, noun, adjective, etc.
- Simple tag set: 19 word classes
“They refuse to permit us to obtain a refuse permit”
What tagset to use?
- Brown Corpus (87 tags)
- C5 tagset (61 tags)
- Penn Treebank (45 tags)
Types of Taggers
- Rule-based (e.g., regular expression)
- Lookup (Unigram)
- N-Gram
- Hybrid and Backoff
- Brill Tagger (learns rules)
- HMM Tagger
** HMM Tagger **
Previosly, we introduce Hidden Markov Models with a weather example. Here, we will show an example of how we can use a HMM to create a POS tagger.
```
# before building the HMM Tagger, let's warm up with implementing our HMM weather example here
# states
start = -1; cold = 0; normal = 1; hot = 2; stateCount = 3
stateNames = ["cold", "normal", "hot"]
# outputs
hotChoc = 0; soda=1; iceCream = 2
timeSteps = 7
# state transition probabilities
trans = {}
trans[(start, cold)] = .1
trans[(start, normal)] = .8
trans[(start, hot)] = .1
trans[(cold, cold)] = .7
trans[(cold, normal)] = .1
trans[(cold, hot)] = .2
trans[(normal, cold)] = .3
trans[(normal, normal)] = .4
trans[(normal, hot)] = .3
trans[(hot, cold)] = .2
trans[(hot, normal)] = .4
trans[(hot, hot)] = .4
# state outputs
output = {}
output[(cold, hotChoc)] = .7
output[(cold, soda)] = .3
output[(cold, iceCream)] = 0
output[(normal, hotChoc)] = .1
output[(normal, soda)] = .7
output[(normal, iceCream)] = .2
output[(hot, hotChoc)] = 0
output[(hot, soda)] = .6
output[(hot, iceCream)] = .4
diary = [soda, soda, hotChoc, iceCream, soda, soda, iceCream]
# manage cell values and back pointers
cells = {}
backStates = {}
def computeMaxPrev(t, sNext):
maxValue = 0
maxState = 0
for s in range(stateCount):
value = cells[t, s] * trans[(s, sNext)]
if (s == 0 or value > maxValue):
maxValue = value
maxState = s
return (maxValue, maxState)
def viterbi(trans, output, diary):
# special handling for t=0 which have no prior states)
for s in range(stateCount):
cells[(0, s)] = trans[(start, s)] * output[(s, diary[0])]
# handle rest of time steps
for t in range(1, timeSteps):
for s in range(stateCount):
maxValue, maxState = computeMaxPrev(t-1, s)
backStates[(t,s)] = maxState
cells[(t, s)] = maxValue * output[(s, diary[t])]
#print("t=", t, "s=", s, "maxValue=", maxValue, "maxState=", maxState, "output=", output[(s, diary[t])], "equals=", cells[(t, s)])
# walk thru cells backwards to get most probable path
path = []
for tt in range(timeSteps):
t = timeSteps - tt - 1 # step t backwards over timesteps
maxValue = 0
maxState = 0
for s in range(stateCount):
value = cells[t, s]
if (s == 0 or value > maxValue):
maxValue = value
maxState = s
path.insert(0, maxState)
return path
# test our algorithm on the weather problem
path = viterbi(trans, output, diary)
print("Weather by days:")
for i in range(timeSteps):
state = path[i]
print(" day=", i+1, stateNames[state])
```
** HMM Tagger Overview**
We are going to use a Hidden Markov Model to help us assign Part-of-Speech tags (like noun, verb, adjective, etc.) to words in a sentence. We treat the human author of the sentence as moving between different meaning states (POS tags) as they compose the sentence. Those state are hidden from us, but we observe the words of the sentence (the output of the meaning states).
In our example here, we will use 4 POS tags from the 87 tag Brown corpus:
- VB (verb, base form)
- TO (infinitive marker)
- NN (common singular noun)
- PPSS (other nominative pronoun)
We are given the state-to-state transition probabilities and the state-output probabilities (see next code cell). We are also given the sentence to decode: "I WANT TO RACE".
```
# OK, here is our HMM POS Tagger for this example
# states
start = -1; VB = 0; TO = 1; NN = 2; PPSS = 3; stateCount = 4
stateNames = ["VB", "TO", "NN", "PPSS"]
# outputs
I = 0; WANT = 1; To = 2; RACE=3
timeSteps = 4
# state transition probabilities
trans = {}
trans[(start, VB)] = .19
trans[(start, TO)] = .0043
trans[(start, NN)] = .041
trans[(start, PPSS)] = .067
trans[(VB, VB)] = .0038
trans[(VB, TO)] = .035
trans[(VB, NN)] = .047
trans[(VB, PPSS)] = .0070
trans[(TO, VB)] = .83
trans[(TO, TO)] = 0
trans[(TO, NN)] = .00047
trans[(TO, PPSS)] = 0
trans[(NN, VB)] = .0040
trans[(NN, TO)] = .016
trans[(NN, NN)] = .087
trans[(NN, PPSS)] = .0045
trans[(PPSS, VB)] = .23
trans[(PPSS, TO)] = .00079
trans[(PPSS, NN)] = .0012
trans[(PPSS, PPSS)] = .00014
# state outputs
output = {}
output[(VB, I)] = 0
output[(VB, WANT)] = .0093
output[(VB, To)] = 0
output[(VB, RACE)] = .00012
output[(TO, I)] = 0
output[(TO, WANT)] = 0
output[(TO, To)] = .99
output[(TO, RACE)] = 0
output[(NN, I)] = 0
output[(NN, WANT)] = .000054
output[(NN, To)] = 0
output[(NN, RACE)] = .00057
output[(PPSS, I)] = .37
output[(PPSS, WANT)] = 0
output[(PPSS, To)] = 0
output[(PPSS, RACE)] = 0
sentence = [I, WANT, To, RACE]
words = ["I", "WANT", "TO", "RACE"]
# manage cell values and back pointers
cells = {}
backStates = {}
def computeMaxPrev(t, sNext):
maxValue = 0
maxState = 0
for s in range(stateCount):
value = cells[t, s] * trans[(s, sNext)]
if (s == 0 or value > maxValue):
maxValue = value
maxState = s
return (maxValue, maxState)
def viterbi(trans, output, sentence):
# special handling for t=0 which have no prior states)
for s in range(stateCount):
cells[(0, s)] = trans[(start, s)] * output[(s, sentence[0])]
# handle rest of time steps
for t in range(1, timeSteps):
for s in range(stateCount):
maxValue, maxState = computeMaxPrev(t-1, s)
backStates[(t,s)] = maxState
cells[(t, s)] = maxValue * output[(s, sentence[t])]
#print("t=", t, "s=", s, "maxValue=", maxValue, "maxState=", maxState, "output=", output[(s, sentence[t])], "equals=", cells[(t, s)])
# walk thru cells backwards to get most probable path
path = []
for tt in range(timeSteps):
t = timeSteps - tt - 1 # step t backwards over timesteps
maxValue = 0
maxState = 0
for s in range(stateCount):
value = cells[t, s]
if (s == 0 or value > maxValue):
maxValue = value
maxState = s
path.insert(0, maxState)
return path
# test our algorithm on the POS TAG data
path = viterbi(trans, output, sentence)
print("Tagged Sentence:")
for i in range(timeSteps):
state = path[i]
print(" word=", words[i], "\ttag=", stateNames[state])
# Here is an example of using the NLTK POS tagger
import nltk
nltk.pos_tag("they refuse to permit us to obtain the refuse permit".split())
# POS tagging with supervised learning, using word suffix parts as features
import nltk
# start by finding the most common 1, 2, and 3 character suffixes of words (using Brown corpus of 1.1 million words)
from nltk.corpus import brown
fd = nltk.FreqDist() # create an empty one that we will count with
for word in brown.words():
wl = word.lower()
fd[wl[-1:]] += 1
fd[wl[-2:]] += 1
fd[wl[-3:]] += 1
topSuffixes = [ key for (key,value) in fd.most_common(30)]
print(topSuffixes[:40])
def pos_features(word):
features = {}
for suffix in topSuffixes:
features[suffix] = word.lower().endswith(suffix)
return features
#pos_features("table")
tagWords = brown.tagged_words(categories="news")
data = [(pos_features(word), tag) for (word,tag) in tagWords]
print("len(data)=", len(data))
dataCount = len(data)
trainCount = int(.8*dataCount)
trainData = data[:trainCount]
testData = data[trainCount:]
dtree = nltk.DecisionTreeClassifier.train(trainData)
#dtree = nltk.NaiveBayesClassifier.train(trainData)
print("train accuracy=", nltk.classify.accuracy(dtree, trainData))
print("test accuracy=", nltk.classify.accuracy(dtree, testData))
print(dtree.classify(pos_features("cats")))
print(dtree.classify(pos_features("house")))
print(dtree.pseudocode(depth=4))
```
## 5. Classical NLP Applications ##
** 5.1 Name Gender Classifier **
```
# code to build a classifier to classify names as male or female
# demonstrates the basics of feature extraction and model building
names = [(name, 'male') for name in nltk.corpus.names.words("male.txt")]
names += [(name, 'female') for name in nltk.corpus.names.words("female.txt")]
def extract_gender_features(name):
name = name.lower()
features = {}
features["suffix"] = name[-1:]
features["suffix2"] = name[-2:] if len(name) > 1 else name[0]
features["suffix3"] = name[-3:] if len(name) > 2 else name[0]
#features["suffix4"] = name[-4:] if len(name) > 3 else name[0]
#features["suffix5"] = name[-5:] if len(name) > 4 else name[0]
#features["suffix6"] = name[-6:] if len(name) > 5 else name[0]
features["prefix"] = name[:1]
features["prefix2"] = name[:2] if len(name) > 1 else name[0]
features["prefix3"] = name[:3] if len(name) > 2 else name[0]
features["prefix4"] = name[:4] if len(name) > 3 else name[0]
features["prefix5"] = name[:5] if len(name) > 4 else name[0]
#features["wordLen"] = len(name)
#for letter in "abcdefghijklmnopqrstuvwyxz":
# features[letter + "-count"] = name.count(letter)
return features
data = [(extract_gender_features(name), gender) for (name,gender) in names]
import random
random.shuffle(data)
#print(data[:10])
#print()
#print(data[-10:])
dataCount = len(data)
trainCount = int(.8*dataCount)
trainData = data[:trainCount]
testData = data[trainCount:]
bayes = nltk.NaiveBayesClassifier.train(trainData)
def classify(name):
label = bayes.classify(extract_gender_features(name))
print("name=", name, "classifed as=", label)
print("trainData accuracy=", nltk.classify.accuracy(bayes, trainData))
print("testData accuracy=", nltk.classify.accuracy(bayes, testData))
bayes.show_most_informative_features(25)
# print gender classifier errors so we can design new features to identify the cases
errors = []
for (name,label) in names:
if bayes.classify(extract_gender_features(name)) != label:
errors.append({"name": name, "label": label})
#errors
```
** 5.2 Sentiment Analysis **
```
# movie reviews / sentiment analysis - part #1
from nltk.corpus import movie_reviews as reviews
import random
docs = [(list(reviews.words(id)), cat) for cat in reviews.categories() for id in reviews.fileids(cat)]
random.shuffle(docs)
#print([ (len(d[0]), d[0][:2], d[1]) for d in docs[:10]])
fd = nltk.FreqDist(word.lower() for word in reviews.words())
topKeys = [ key for (key,value) in fd.most_common(2000)]
# movie reviews sentiment analysis - part #2
import nltk
def review_features(doc):
docSet = set(doc)
features = {}
for word in topKeys:
features[word] = (word in docSet)
return features
#review_features(reviews.words("pos/cv957_8737.txt"))
data = [(review_features(doc), label) for (doc,label) in docs]
dataCount = len(data)
trainCount = int(.8*dataCount)
trainData = data[:trainCount]
testData = data[trainCount:]
bayes2 = nltk.NaiveBayesClassifier.train(trainData)
print("train accuracy=", nltk.classify.accuracy(bayes2, trainData))
print("test accuracy=", nltk.classify.accuracy(bayes2, testData))
bayes2.show_most_informative_features(20)
```
** 5.3 Named Entity Recognition (NER) **
Popular Named Entity Types:
- ORGANIZATION
- PERSON
- LOCATION
- GPE
- DATE
Named Entity Extraction Techniques:
- Chunking (tag pattern to group)
- Chinking (tag pattern to omit)
- Nested chunks (recursion)
- Hand-crafted rules
- Rules learned from data
```
# Named Entity Regcognition (NER)
# processes sentences and produces (entity, relation, entity) triples!
import nltk
# first, process the document by separating the text into sentences, then words within sentences, then tag words by sentence
def preprocess(doc):
sents = nltk.sent_tokenize(doc)
sents2 = [nltk.word_tokenize(sent) for sent in sents]
sents3 = [nltk.pos_tag(sent) for sent in sents2]
# we are going to use a technique called CHUNKING where we label sequences of POS tags as a high level tag, like a
# noun phrase (NP).
# here we test our idea with a simple sentence, and a grammar for detectecting NP. The grammar:
# <NP> ::= [ <DT> ] [ <JJ list> ] <NN>
tagged_sent = [("the", "DT"), ("little", "JJ"), ("yellow", "JJ"), ("dog", "NN"), ("barked", "VBD"), ("at", "IN"),
("the", "DT"), ("cat", "NN")]
#np_grammar = "NP: {<DT>?<JJ>*<NN>}"
#np_grammar = "NP: {<DT>?<JJ.*>*<NN.*>*}"
np_grammar = r"""
NP:
{<DT|PP\$>?<JJ>*<NN>}
{<NPP>+}
"""
parser = nltk.RegexpParser(np_grammar)
result = parser.parse(tagged_sent)
print(result.__repr__())
```
| github_jupyter |
```
import os
os.environ['CUDA_VISIBLE_DEVICES'] = ''
!wget https://f000.backblazeb2.com/file/malaya-model/v39/summarization/base.pb
!wget https://f000.backblazeb2.com/file/malaya-model/v39/summarization/small.pb
import tensorflow as tf
from tensorflow.tools.graph_transforms import TransformGraph
from glob import glob
tf.set_random_seed(0)
pbs = glob('*.pb')
pbs
import tensorflow_text
import tf_sentencepiece
transforms = ['add_default_attributes',
'remove_nodes(op=Identity, op=CheckNumerics, op=Dropout)',
'fold_constants(ignore_errors=true)',
'fold_batch_norms',
'fold_old_batch_norms',
'quantize_weights(fallback_min=-10, fallback_max=10)',
'strip_unused_nodes',
'sort_by_execution_order']
for pb in pbs:
input_graph_def = tf.GraphDef()
with tf.gfile.FastGFile(pb, 'rb') as f:
input_graph_def.ParseFromString(f.read())
print(pb)
transformed_graph_def = TransformGraph(input_graph_def,
['Placeholder', 'Placeholder_2'],
['greedy', 'beam', 'nucleus'], transforms)
with tf.gfile.GFile(f'{pb}.quantized', 'wb') as f:
f.write(transformed_graph_def.SerializeToString())
def load_graph(frozen_graph_filename, **kwargs):
with tf.gfile.GFile(frozen_graph_filename, 'rb') as f:
graph_def = tf.GraphDef()
graph_def.ParseFromString(f.read())
# https://github.com/onnx/tensorflow-onnx/issues/77#issuecomment-445066091
# to fix import T5
for node in graph_def.node:
if node.op == 'RefSwitch':
node.op = 'Switch'
for index in xrange(len(node.input)):
if 'moving_' in node.input[index]:
node.input[index] = node.input[index] + '/read'
elif node.op == 'AssignSub':
node.op = 'Sub'
if 'use_locking' in node.attr:
del node.attr['use_locking']
elif node.op == 'AssignAdd':
node.op = 'Add'
if 'use_locking' in node.attr:
del node.attr['use_locking']
elif node.op == 'Assign':
node.op = 'Identity'
if 'use_locking' in node.attr:
del node.attr['use_locking']
if 'validate_shape' in node.attr:
del node.attr['validate_shape']
if len(node.input) == 2:
node.input[0] = node.input[1]
del node.input[1]
with tf.Graph().as_default() as graph:
tf.import_graph_def(graph_def)
return graph
import malaya
model = malaya.summarization.abstractive.transformer(model = 'small')
string = 'KUALA LUMPUR: Presiden Perancis Emmanuel Macron tidak menampakkan beliau seorang sosok yang bertamadun, selar Tun Dr Mahathir Mohamad menerusi kemas kini terbaharu di blognya. Bekas Perdana Menteri itu mendakwa, pemerintah tertinggi Perancis itu bersikap primitif kerana menuduh orang Islam terlibat dalam pembunuhan guru yang menghina Islam, malah menegaskan tindakan membunuh bukan ajaran Islam. Jelas Dr Mahathir, sejarah membuktikan bahawa orang Perancis pernah membunuh jutaan manusia, yang ramai mangsanya terdiri dari orang Islam.'
model.summarize([string])
e = model._tokenizer.encode(f'ringkasan: {string}')
e = e + [1]
g = load_graph('small.pb.quantized')
x = g.get_tensor_by_name('import/Placeholder:0')
logits = g.get_tensor_by_name('import/greedy:0')
test_sess = tf.InteractiveSession(graph = g)
l = test_sess.run(logits, feed_dict = {x: [e]})
model._tokenizer.decode(l[0].tolist())
quantized = glob('*.pb.quantized')
quantized
from b2sdk.v1 import *
info = InMemoryAccountInfo()
b2_api = B2Api(info)
application_key_id = 'd3c416cf4cb1'
application_key = '0007c73b0ef09cbff76ebdd5b14f2e0044d6d44b74'
b2_api.authorize_account("production", application_key_id, application_key)
file_info = {'how': 'good-file'}
b2_bucket = b2_api.get_bucket_by_name('malaya-model')
for file in quantized:
print(file)
key = file
outPutname = f"v40/summarize/transformer-{file}"
b2_bucket.upload_local_file(
local_file=key,
file_name=outPutname,
file_infos=file_info,
)
!rm *.pb*
# converter = tf.compat.v1.lite.TFLiteConverter.from_frozen_graph(
# graph_def_file='test.pb',
# input_arrays=['Placeholder', 'Placeholder_1'],
# input_shapes={'Placeholder' : [None, 512], 'Placeholder_1': [None, 512]},
# output_arrays=['logits'],
# )
# # converter.allow_custom_ops=True
# converter.target_spec.supported_ops = [tf.lite.OpsSet.TFLITE_BUILTINS, tf.lite.OpsSet.SELECT_TF_OPS]
# converter.target_spec.supported_types = [tf.float16]
# converter.optimizations = [tf.lite.Optimize.DEFAULT]
# converter.experimental_new_converter = True
# tflite_model = converter.convert()
# converter.target_spec.supported_ops = [tf.lite.OpsSet.TFLITE_BUILTINS,
# tf.lite.OpsSet.SELECT_TF_OPS]
# converter.target_spec.supported_types = [tf.float16]
# converter.optimizations = [tf.lite.Optimize.DEFAULT]
# tflite_model = converter.convert()
# with open('tiny-bert-sentiment-float16.tflite', 'wb') as f:
# f.write(tflite_model)
# converter.target_spec.supported_ops = [tf.lite.OpsSet.TFLITE_BUILTINS,
# tf.lite.OpsSet.SELECT_TF_OPS]
# converter.optimizations = [tf.lite.Optimize.OPTIMIZE_FOR_SIZE]
# tflite_model = converter.convert()
# with open('tiny-bert-sentiment-hybrid.tflite', 'wb') as f:
# f.write(tflite_model)
# interpreter = tf.lite.Interpreter(model_path='tiny-bert-sentiment-hybrid.tflite')
# interpreter.allocate_tensors()
```
| github_jupyter |
<i>Copyright (c) Microsoft Corporation. All rights reserved.</i>
<i>Licensed under the MIT License.</i>
# Variational Autoencoders for Collaborative Filtering on MovieLens dataset.
This notebook accompanies the paper "*Variational autoencoders for collaborative filtering*" by Dawen Liang, Rahul G. Krishnan, Matthew D. Hoffman, and Tony Jebara, in The Web Conference (aka WWW) 2018 [[Liang, Dawen, et al,2018]](https://arxiv.org/pdf/1802.05814.pdf). In this original paper, the public dataset "20M-MovieLens" is used. However, in our notebook, we used the "1M-MovieLens" dataset because it takes less time for the model to train.
In this notebook, we will show a complete self-contained example of training a Multinomial Variational Autoencoder (described in the original paper) on the public "1M-Movielens" dataset, including data preprocessing, model training and model evaluation. In the whole notebook we assume that the reader has basic knowledge about VAE [[Kingma et al, 2013]](https://arxiv.org/pdf/1312.6114.pdf).
# 0 Global Settings and Imports
```
# download the necessary libraries
! pip install tensorflow==2.2.0-rc1
! pip install keras==2.3.1
! pip install papermill
import sys
import os
import numpy as np
import pandas as pd
import papermill as pm
import matplotlib.pyplot as plt
%matplotlib inline
import seaborn as sns
sns.set()
import tensorflow as tf
import keras
from recommenders.utils.timer import Timer
from recommenders.datasets import movielens
from recommenders.datasets.split_utils import min_rating_filter_pandas
from recommenders.datasets.python_splitters import numpy_stratified_split
from recommenders.evaluation.python_evaluation import map_at_k, ndcg_at_k, precision_at_k, recall_at_k
from recommenders.datasets.sparse import AffinityMatrix
from recommenders.utils.python_utils import binarize
from recommenders.models.vae.multinomial_vae import Mult_VAE
from tempfile import TemporaryDirectory
print("System version: {}".format(sys.version))
print("Pandas version: {}".format(pd.__version__))
print("Tensorflow version: {}".format(tf.__version__))
print("Keras version: {}".format(keras.__version__))
# top k items to recommend
TOP_K = 100
# Select MovieLens data size: 100k, 1m, 10m, or 20m
MOVIELENS_DATA_SIZE = '1m'
# Model parameters
HELDOUT_USERS = 600 # CHANGE FOR DIFFERENT DATASIZE
INTERMEDIATE_DIM = 200
LATENT_DIM = 70
EPOCHS = 400
BATCH_SIZE = 100
# temporary Path to save the optimal model's weights
tmp_dir = TemporaryDirectory()
WEIGHTS_PATH = os.path.join(tmp_dir, "mvae_weights.hdf5")
SEED = 98765
```
# 1 Multi-VAE algorithm
__Notations__: The notation used is described below.
$u \in \{1,\dots,U\}$ to index users and $i \in \{1,\dots,I\}$ to index items. The user-by-item interaction matrix is the click matrix $\mathbf{X} \in \mathbb{N}^{U\times I}$, which is given as an input to the model. The lower case $\mathbf{x}_u =[X_{u1},\dots,X_{uI}]^\top \in \mathbb{N}^I$ is a bag-of-words vector with the number of clicks for each item from user u. The click matrix is binarized. It is straightforward to extend it to general count data.
__Multi-VAE Model__:
For each user u, the model starts by sampling a $K$-dimensional latent representation $\mathbf{z}_u$ from a standard Gaussian prior. The latent representation $\mathbf{z}_u$ is transformed via a non-linear function $f_\theta (\cdot) \in \mathbb{R}^I$ to produce a probability distribution over $I$ items $\pi (\mathbf{z}_u)$ from which the click history $\mathbf{x}_u$ is assumed to have been drawn:
$$
\mathbf{z}_u \sim \mathcal{N}(0, \mathbf{I}_K), \pi(\mathbf{z}_u) = softmax\{f_\theta (\mathbf{z}_u\},\\
\mathbf{x}_u \sim \mathrm{Mult}(N_u, \pi(\mathbf{z}_u))
$$
$\mathbf{z}_u$ needs to be sampled from an approximate posterior $q_\phi (\mathbf{z}_u | \mathbf{x}_u )$, which is assumed to be a Gaussian. To compute the gradients the reparametrization trick is used and $\mathbf{z}_u$ is calculated by the formula
$$
\mathbf{z}_u = \mathbf\mu(x_u)+\mathbf\sigma(x_u) \cdot \mathbf\epsilon
$$
where $\mathbf\epsilon \sim \mathcal{N}(0, \mathbf{I})$ and $ \mathbf\mu(x_u), \sigma(x_u)$ are calculated in encoder.
The objective of Multi-VAE for a single user $u$ is:
$$
\mathcal{L}_u(\theta, \phi) = \mathbb{E}_{q_\phi(z_u | x_u)}[\log p_\theta(x_u | z_u)] - \beta \cdot KL(q_\phi(z_u | x_u) \| p(z_u))
$$
where $q_\phi$ is the approximating variational distribution (inference model/ encoder), and $p_\theta$ refers to generative model/decoder. The first term is the log-likelohood and the second term is the Kullback-Leibler divergence term.
Regarding the first term, we use the multinomial log-likelihood formula as proposed in the original paper:
$$\log p_\theta(\mathbf{x}_u | \mathbf{z}_u) = \sum_{i} \mathbf{x}_{ui}\log \mathbf{\pi}_i (z_u) $$
The authors use multinomial likelihood which is intuitively better for modeling implicit feedback and as it is proven by their research work performs better than logistic or gaussian likelihoods, which are typically more popular in the literature. By using multinomial likelihood, they are treating the problem as multi-class classification. As the authors discuss in the paper in section 2.1: "this likelihood rewards the model for putting probability mass on the non-zero entries in $x_u$. But the model has a limited budget of probability mass, since $\pi(z_u)$ must sum to 1; the items must compete for this limited budget. The model should therefore assign more probability mass to items that are more likely to be clicked."
Also, $\mathbf KL-divergence$ is treated as a regularization term as described from[[Higgins et al, 2016]](https://openreview.net/pdf?id=Sy2fzU9gl), [[Burgess et al, 2018]](https://arxiv.org/pdf/1804.03599.pdf) :
1. When $\mathbf \beta = 1$ corresponds to the original VAE formulation of [[Kingma et al, 2013]](https://arxiv.org/pdf/1312.6114.pdf)
2. Setting $\mathbf \beta > 1$, provides a stronger constraint on the latent bottleneck than in the original VAE formulation. These constraints limit the capacity of $\mathbf z$, which, combined with the pressure to maximise the log likelihood of the training data $\mathbf x$ under the model, should encourage the model to learn the most efficient representation of the data.
3. Setting $\mathbf \beta<1$. In case of recommendation systems our goal is to make good recommendations and not reconstruct the input as close as possible to the initial input, or in other words to maximize the log likelihood. So if we set β<1, then we are weakening the influence of the prior. Futher details can be found in section 2.2 of [[Liang, Dawen, et al,2018]](https://arxiv.org/pdf/1802.05814.pdf).
As a result, for the reasons explained above in 3., the KL-divergence is multiplied by the parameter $\beta \in [0,1]$.
__Selecting β:__ As proposed from the original paper, a simple heuristic methodology is used for selecting $\mathbf \beta$ . The training process starts with $\mathbf \beta = 0$, and gradually increase $\mathbf \beta$ to 1. The $\mathbf KL$ term is slowly annealed over a large number of gradient updates to $\mathbf \theta , \phi $. The optimal $\mathbf \beta$ is recorded when the performance of the model reaches the peak. Then using this optimal beta the model is retrained using the same annealing procedure, but $\mathbf \beta$ stops increasing when it reaches the optimal value found in the previous step.
This method works well and with the benefit that it does not require training multiple models with different values of $\mathbf \beta$, which can be time-consuming. This idea is based on the paper
[[Bowman et al, 2015]](https://arxiv.org/pdf/1511.06349.pdf). This methodology is explained in more detail in section (4).
# 2 Keras implementation of Multi-VAE
For the implementation of the model, Keras package is used.
For the scope of the project, the MovieLens dataset is used, which is composed of user-to-item interactions and integer ratings from 1 to 5. We convert MovieLens into binarized clicked matrix ( 1: the user liked this movie , 0: the user did NOT like or did NOT watch/rate this movie), and evaluate based on heldout users data.
# 3 Data Preparation
### 3.1 Load data and split
The data are loaded and splitted into train / validation / test sets following strong generalization:
- All unique users are splitted into training users and heldout users (i.e. validation and test users)
- By using the lists of these users, the corresponding training data and heldout data are obtained, which are converted to click matrices
- Models are trained using the entire click history of the training users.
- To evaluate them, part of the click history from heldout data(validation and test) is used to learn the necessary user-level representations for the model. Then, evaluation metrics are computed by looking at how well the model ranks the rest of the unseen click history from the heldout data
#### 3.1.1 Load data
```
df = movielens.load_pandas_df(
size=MOVIELENS_DATA_SIZE,
header=["userID", "itemID", "rating", "timestamp"]
)
df.head()
df.shape
```
#### 3.1.2 Data Filtering
For the data filtering we are using the below 3 steps as recommended from the original paper [[Liang, Dawen, et al,2018]](https://dl.acm.org/doi/pdf/10.1145/3178876.3186150?casa_token=zul5haircsAAAAAA:iIKn7y-xWwSeqaP-MmmyUaJoJuNZX9Fx1aXeFJwkwtMpVDCrPMW3kZjuYo1LKhSuMeUMNf1mbP2o).
So, we have to make sure that :
- user-to-movie interactions with rating <=3.5 are filtered out. Applying this filtering we make sure that if a movie is rated less than 3.5 from the users that they watched this movie, it will not be contained in the final click matrix. If we do not apply this filter, the final click matrix will be even sparser.
- the users who clicked less than 5 movies are filtered out.
- the movies which are not clicked by any user are filtered out.
```
# Binarize the data (only keep ratings >= 4)
df_preferred = df[df['rating'] > 3.5]
print (df_preferred.shape)
df_low_rating = df[df['rating'] <= 3.5]
# df.head()
df_preferred.head(10)
# Keep users who clicked on at least 5 movies
df = min_rating_filter_pandas(df_preferred, min_rating=5, filter_by="user")
# Keep movies that were clicked on by at least on 1 user
df = min_rating_filter_pandas(df, min_rating=1, filter_by="item")
# Obtain both usercount and itemcount after filtering
usercount = df[['userID']].groupby('userID', as_index = False).size()
itemcount = df[['itemID']].groupby('itemID', as_index = False).size()
# Compute sparsity after filtering
sparsity = 1. * df.shape[0] / (usercount.shape[0] * itemcount.shape[0])
print("After filtering, there are %d watching events from %d users and %d movies (sparsity: %.3f%%)" %
(df.shape[0], usercount.shape[0], itemcount.shape[0], sparsity * 100))
```
#### 3.1.3 Split data
For data slitting we use:
- 600 (~ 10%) users in validation set
- 600 (~ 10%) users in testing set
- the rest of them (~ 80%) in training set
Since the model is trained using the click history of the training users, we have to make sure that the movies that exist in the validation and test sets are the movies that exist in the train set. In other words, validation and test set should not contain movies that do not exist in the train set.
```
unique_users = sorted(df.userID.unique())
np.random.seed(SEED)
unique_users = np.random.permutation(unique_users)
# Create train/validation/test users
n_users = len(unique_users)
print("Number of unique users:", n_users)
train_users = unique_users[:(n_users - HELDOUT_USERS * 2)]
print("\nNumber of training users:", len(train_users))
val_users = unique_users[(n_users - HELDOUT_USERS * 2) : (n_users - HELDOUT_USERS)]
print("\nNumber of validation users:", len(val_users))
test_users = unique_users[(n_users - HELDOUT_USERS):]
print("\nNumber of test users:", len(test_users))
# For training set keep only users that are in train_users list
train_set = df.loc[df['userID'].isin(train_users)]
print("Number of training observations: ", train_set.shape[0])
# For validation set keep only users that are in val_users list
val_set = df.loc[df['userID'].isin(val_users)]
print("\nNumber of validation observations: ", val_set.shape[0])
# For test set keep only users that are in test_users list
test_set = df.loc[df['userID'].isin(test_users)]
print("\nNumber of test observations: ", test_set.shape[0])
# train_set/val_set/test_set contain user - movie interactions with rating 4 or 5
# Obtain list of unique movies used in training set
unique_train_items = pd.unique(train_set['itemID'])
print("Number of unique movies that rated in training set", unique_train_items.size)
# For validation set keep only movies that used in training set
val_set = val_set.loc[val_set['itemID'].isin(unique_train_items)]
print("Number of validation observations after filtering: ", val_set.shape[0])
# For test set keep only movies that used in training set
test_set = test_set.loc[test_set['itemID'].isin(unique_train_items)]
print("\nNumber of test observations after filtering: ", test_set.shape[0])
```
## 3.2 Click matrix generation
From section 3.1 we end up with 3 datasets train_set, val_set and test_set. As the authors indicate, a click matrix, that contains only 0-s and 1-s, is provided as an input to the model, where each row represents a user and each column represents a movie.
So, the click matrix contains the preferences of the user, marking each cell with 0 when the user did not enjoy (ratings below 3.5) or did not watch a movie and with 1 when the user enjoyed a movie (ratings above 3.5).
The training set will be a click matrix containing full historicity of all training users. However, the test set and validation set should be splitted into train and test parts. As a result, we get 4 datasets:
- val_data_tr
- val_data_te
- test_data_tr
- test_data_te
'val_data_tr' contains 75% of the the preferred movies (movies marked as 1 in the click matrix) per user.
The rest 25% of the preffered movies are contained into the 'val_data_te'. The same splitting is followed for test set.
The 'val_data_tr' is given as an input for our model at the end of each epoch. The result of the model is a 'reconstructed_val_data_tr', which contains the movies recommended for each user by the model. In order to evaluate the performance of the model, at the end of each epoch, we compare the 'reconstructed_val_data_tr' (predicted recommendations by the model) with the 'val_data_te' (true movie preferences of each user) using NDCG@k metric.
For the final evaluation of the model the 'test_data_tr' and 'test_data_te' are being used. As it is described before, the 'test_data_tr' is given as an input for the model and returns the 'reconstructed_test_data_tr' dataset with the recommendations made by the model. Then, the 'reconstructed_test_data_tr' is compared with 'test_data_te' through different metrics:
- MAP
- NDCG@k
- Recall@k
- Precision@k
```
# Instantiate the sparse matrix generation for train, validation and test sets
# use list of unique items from training set for all sets
am_train = AffinityMatrix(df=train_set, items_list=unique_train_items)
am_val = AffinityMatrix(df=val_set, items_list=unique_train_items)
am_test = AffinityMatrix(df=test_set, items_list=unique_train_items)
# Obtain the sparse matrix for train, validation and test sets
train_data, _, _ = am_train.gen_affinity_matrix()
print(train_data.shape)
val_data, val_map_users, val_map_items = am_val.gen_affinity_matrix()
print(val_data.shape)
test_data, test_map_users, test_map_items = am_test.gen_affinity_matrix()
print(test_data.shape)
# Split validation and test data into training and testing parts
val_data_tr, val_data_te = numpy_stratified_split(val_data, ratio=0.75, seed=SEED)
test_data_tr, test_data_te = numpy_stratified_split(test_data, ratio=0.75, seed=SEED)
# Binarize train, validation and test data
train_data = binarize(a=train_data, threshold=3.5)
val_data = binarize(a=val_data, threshold=3.5)
test_data = binarize(a=test_data, threshold=3.5)
# Binarize validation data: training part
val_data_tr = binarize(a=val_data_tr, threshold=3.5)
# Binarize validation data: testing part (save non-binary version in the separate object, will be used for calculating NDCG)
val_data_te_ratings = val_data_te.copy()
val_data_te = binarize(a=val_data_te, threshold=3.5)
# Binarize test data: training part
test_data_tr = binarize(a=test_data_tr, threshold=3.5)
# Binarize test data: testing part (save non-binary version in the separate object, will be used for calculating NDCG)
test_data_te_ratings = test_data_te.copy()
test_data_te = binarize(a=test_data_te, threshold=3.5)
# retrieve real ratings from initial dataset
test_data_te_ratings=pd.DataFrame(test_data_te_ratings)
val_data_te_ratings=pd.DataFrame(val_data_te_ratings)
for index,i in df_low_rating.iterrows():
user_old= i['userID'] # old value
item_old=i['itemID'] # old value
if (test_map_users.get(user_old) is not None) and (test_map_items.get(item_old) is not None) :
user_new=test_map_users.get(user_old) # new value
item_new=test_map_items.get(item_old) # new value
rating=i['rating']
test_data_te_ratings.at[user_new,item_new]= rating
if (val_map_users.get(user_old) is not None) and (val_map_items.get(item_old) is not None) :
user_new=val_map_users.get(user_old) # new value
item_new=val_map_items.get(item_old) # new value
rating=i['rating']
val_data_te_ratings.at[user_new,item_new]= rating
val_data_te_ratings=val_data_te_ratings.to_numpy()
test_data_te_ratings=test_data_te_ratings.to_numpy()
# test_data_te_ratings
# Just checking
print(np.sum(val_data))
print(np.sum(val_data_tr))
print(np.sum(val_data_te))
# Just checking
print(np.sum(test_data))
print(np.sum(test_data_tr))
print(np.sum(test_data_te))
```
# 4 Train Multi-VAE using Keras
__Model Architecture:__
For "MovieLens-1M dataset", we set both the generative function $f_\theta(\cdot)$ and the inference model $g_\phi(\cdot)$ to be 3-layer multilayer perceptron (MLP) with symmetrical architecture.
The generative function is a [70 -> 200 -> n_items] MLP, which means the inference function is a [n_items -> 200 -> 70] MLP. Thus the overall architecture for the Multi-VAE is [n_items -> 200 -> 70 -> 200 -> n_items].
Also, Dropout is applied both in encoder and decoder to avoid overfitting.
Note that this architecture, illustrated in the graph below, is the one that resulted optimal results after investigating multiple architectures both for encoder and decoder.

__Model Training:__
One of the most interesting parts of the original paper [[Liang, Dawen, et al,2018]](https://arxiv.org/pdf/1802.05814.pdf) is the tuning of $\mathbf \beta$ using annealing. As it is described in section 1, $\mathbf KL- divergence$ is handled as regularization term. $\mathbf KL-divergence$ term is linearly annealed by increasing $\mathbf \beta$ from 0 to 1 slowly over a large number of training steps. From the original paper is proposed that __$\mathbf \beta$ should reach $\mathbf \beta = 1$ at around the 80% of the epochs__ and then $\mathbf \beta$ remains stable at the value of 1. Then optimal beta is found based on the peak value of NDCG@k metric of the validation set.
After finding the optimal beta, we retrain the model with the same annealing but we stop the increase of $\mathbf \beta$ after reaching the value of optimal $\mathbf \beta$ found in the previous step. Alternatively, we can say that we anneal $\mathbf \beta$ until it reaches the anneal_cap (anneal cap = optimal $\mathbf \beta$) at 80% of epochs.
## 4.1 Multi-VAE without annealing (initial approach)
Firstly, for comparison reasons Multi-VAE model is trained using constant $\mathbf \beta$= 1.
```
model_without_anneal = Mult_VAE(n_users=train_data.shape[0], # Number of unique users in the training set
original_dim=train_data.shape[1], # Number of unique items in the training set
intermediate_dim=INTERMEDIATE_DIM,
latent_dim=LATENT_DIM,
n_epochs=EPOCHS,
batch_size=BATCH_SIZE,
k=TOP_K,
verbose=0,
seed=SEED,
save_path=WEIGHTS_PATH,
drop_encoder=0.5,
drop_decoder=0.5,
annealing=False,
beta=1.0
)
with Timer() as t:
model_without_anneal.fit(x_train=train_data,
x_valid=val_data,
x_val_tr=val_data_tr,
x_val_te=val_data_te_ratings,
mapper=am_val
)
print("Took {} seconds for training.".format(t))
model_without_anneal.display_metrics()
ndcg_val_without_anneal = model_without_anneal.ndcg_per_epoch()
```
#### Prediction and Evaluation of Multi-VAE model using constant 𝛽 = 1.
Evaluate with recommending 10 items.
```
# Use k = 10
with Timer() as t:
# Model prediction on the training part of test set
top_k = model_without_anneal.recommend_k_items(x=test_data_tr,
k=10,
remove_seen=True
)
# Convert sparse matrix back to df
top_k_df = am_test.map_back_sparse(top_k, kind='prediction')
test_df = am_test.map_back_sparse(test_data_te_ratings, kind='ratings') # use test_data_te_, with the original ratings
print("Took {} seconds for prediction.".format(t))
# Use the ranking metrics for evaluation
eval_map_1 = map_at_k(test_df, top_k_df, col_prediction='prediction', k=10)
eval_ndcg_1 = ndcg_at_k(test_df, top_k_df, col_prediction='prediction', k=10)
eval_precision_1 = precision_at_k(test_df, top_k_df, col_prediction='prediction', k=10)
eval_recall_1 = recall_at_k(test_df, top_k_df, col_prediction='prediction', k=10)
print("MAP@10:\t\t%f" % eval_map_1,
"NDCG@10:\t%f" % eval_ndcg_1,
"Precision@10:\t%f" % eval_precision_1,
"Recall@10: \t%f" % eval_recall_1, sep='\n')
```
Evaluate with recommending 100 items.
```
# Use k = TOP_K
with Timer() as t:
# Model prediction on the training part of test set
top_k = model_without_anneal.recommend_k_items(x=test_data_tr,
k=TOP_K,
remove_seen=True
)
# Convert sparse matrix back to df
top_k_df = am_test.map_back_sparse(top_k, kind='prediction')
test_df = am_test.map_back_sparse(test_data_te_ratings, kind='ratings') # use test_data_te_, with the original ratings
print("Took {} seconds for prediction.".format(t))
# Use the ranking metrics for evaluation
eval_map_2 = map_at_k(test_df, top_k_df, col_prediction='prediction', k=TOP_K)
eval_ndcg_2 = ndcg_at_k(test_df, top_k_df, col_prediction='prediction', k=TOP_K)
eval_precision_2 = precision_at_k(test_df, top_k_df, col_prediction='prediction', k=TOP_K)
eval_recall_2 = recall_at_k(test_df, top_k_df, col_prediction='prediction', k=TOP_K)
print("MAP@100:\t%f" % eval_map_2,
"NDCG@100:\t%f" % eval_ndcg_2,
"Precision@100:\t%f" % eval_precision_2,
"Recall@100: \t%f" % eval_recall_2, sep='\n')
```
## 4.2 Multi-VAE with annealing
Now the annealing procedure for finding the optimal $\mathbf \beta$ is used.
In order to find the optimal $\beta$, the model is trained using annealing with anneal_cap equal 1.0.
```
model_with_anneal = Mult_VAE(n_users=train_data.shape[0], # Number of unique users in the training set
original_dim=train_data.shape[1], # Number of unique items in the training set
intermediate_dim=INTERMEDIATE_DIM,
latent_dim=LATENT_DIM,
n_epochs=EPOCHS,
batch_size=BATCH_SIZE,
k=TOP_K,
verbose=0,
seed=SEED,
save_path=WEIGHTS_PATH,
drop_encoder=0.5,
drop_decoder=0.5,
annealing=True,
anneal_cap=1.0,
)
with Timer() as t:
model_with_anneal.fit(x_train=train_data,
x_valid=val_data,
x_val_tr=val_data_tr,
x_val_te=val_data_te_ratings,
mapper=am_val
)
print("Took {} seconds for training.".format(t))
model_with_anneal.display_metrics()
ndcg_val_with_anneal = model_with_anneal.ndcg_per_epoch()
```
Using the optimal beta as anneal cap , we retrain our model.
When NDCG@k of validation set reach a peak, the weights of the model are saved. Using this model we evaluate the test set.
```
# Get optimal beta
optimal_beta = model_with_anneal.get_optimal_beta()
print( "The optimal beta is: ", optimal_beta)
model_optimal_beta = Mult_VAE(n_users=train_data.shape[0], # Number of unique users in the training set
original_dim=train_data.shape[1], # Number of unique items in the training set
intermediate_dim=INTERMEDIATE_DIM,
latent_dim=LATENT_DIM,
n_epochs=EPOCHS,
batch_size=BATCH_SIZE,
k=TOP_K,
verbose=0,
seed=SEED,
save_path=WEIGHTS_PATH,
drop_encoder=0.5,
drop_decoder=0.5,
annealing=True,
anneal_cap=optimal_beta,
)
with Timer() as t:
model_optimal_beta.fit(x_train=train_data,
x_valid=val_data,
x_val_tr=val_data_tr,
x_val_te=val_data_te_ratings,
mapper=am_val
)
print("Took {} seconds for training.".format(t))
model_optimal_beta.display_metrics()
ndcg_val_optimal_beta = model_optimal_beta.ndcg_per_epoch()
```
#### Prediction and Evaluation of Standard-VAE model using the optimal $\beta$ with annealing.
Evaluate with recommending 10 items.
```
# Use k = 10
with Timer() as t:
# Model prediction on the training part of test set
top_k = model_optimal_beta.recommend_k_items(x=test_data_tr,
k=10,
remove_seen=True
)
# Convert sparse matrix back to df
top_k_df = am_test.map_back_sparse(top_k, kind='prediction')
test_df = am_test.map_back_sparse(test_data_te_ratings, kind='ratings') # use test_data_te_, with the original ratings
print("Took {} seconds for prediction.".format(t))
# Use the ranking metrics for evaluation
eval_map_3 = map_at_k(test_df, top_k_df, col_prediction='prediction', k=10)
eval_ndcg_3 = ndcg_at_k(test_df, top_k_df, col_prediction='prediction', k=10)
eval_precision_3 = precision_at_k(test_df, top_k_df, col_prediction='prediction', k=10)
eval_recall_3 = recall_at_k(test_df, top_k_df, col_prediction='prediction', k=10)
print("MAP@10:\t\t%f" % eval_map_3,
"NDCG@10:\t%f" % eval_ndcg_3,
"Precision@10:\t%f" % eval_precision_3,
"Recall@10: \t%f" % eval_recall_3, sep='\n')
```
Evaluate with recommending 100 items.
```
# Use k = 100s
with Timer() as t:
# Model prediction on the training part of test set
top_k = model_optimal_beta.recommend_k_items(x=test_data_tr,
k=TOP_K,
remove_seen=True
)
# Convert sparse matrix back to df
top_k_df = am_test.map_back_sparse(top_k, kind='prediction')
test_df = am_test.map_back_sparse(test_data_te_ratings, kind='ratings') # use test_data_te_, with the original ratings
print("Took {} seconds for prediction.".format(t))
# Use the ranking metrics for evaluation
eval_map_4 = map_at_k(test_df, top_k_df, col_prediction='prediction', k=TOP_K)
eval_ndcg_4 = ndcg_at_k(test_df, top_k_df, col_prediction='prediction', k=TOP_K)
eval_precision_4 = precision_at_k(test_df, top_k_df, col_prediction='prediction', k=TOP_K)
eval_recall_4 = recall_at_k(test_df, top_k_df, col_prediction='prediction', k=TOP_K)
print("MAP@100:\t%f" % eval_map_4,
"NDCG@100:\t%f" % eval_ndcg_4,
"Precision@100:\t%f" % eval_precision_4,
"Recall@100: \t%f" % eval_recall_4, sep='\n')
```
# 5 Conclusion
Through this notebook is proven that Mult-VAE has competitive performance.
From the plot below, we can see that the model using annealing outperforms the model without annealing. Specifically, the results of evaluating the test set, for the the 2 different approaches, are:
| Model | NDCG@100 | NDCG@10 |
| --- | --- | --- |
| Mult-VAE (wihtout annealing, β=1)| 0.384 | 0.430 |
| Mult-VAE (with annealing, optimal β)| 0.430 | 0.479 |
This annealing procedure is used as an efficient way to tune the parameter $\mathbf \beta$. Otherwise, training multiple models using different values of $\mathbf \beta$ can be really time consuming.
```
# Plot setup
plt.figure(figsize=(15, 5))
sns.set(style='whitegrid')
# Plot NDCG@k of validation sets for three models
plt.plot(ndcg_val_without_anneal, color='b', linestyle='-', label='without anneal')
plt.plot(ndcg_val_with_anneal, color='g', linestyle='-', label='with anneal at β=1')
plt.plot(ndcg_val_optimal_beta, color='r', linestyle='-', label='with anneal at optimal β')
# Add plot title and axis names
plt.title('VALIDATION NDCG@100 FOR DIFFERENT MODELS \n', size=16)
plt.xlabel('Epochs', size=14)
plt.ylabel('NDCG@100', size=14)
plt.legend(loc='lower right')
plt.show()
```
Both for choosing the optimal $\mathbf \beta$ and the optimal model weights the NDCG@100 evaluation metric is used for the validation set (section [4], Paragraph: __Model Training__) as proposed from [[Liang, Dawen, et al,2018]](https://arxiv.org/pdf/1802.05814.pdf).
Last but not least, it is worth mentioning that for different shuffling of input data, the model may need again hyperparameter tuning or may not work properly since the training dataset may not be indicative for the test set.
# 6 References
[Liang, Dawen, et al, 2018] [Liang, Dawen, et al. "Variational autoencoders for collaborative filtering." Proceedings of the 2018 World Wide Web Conference. 2018.](https://dl.acm.org/doi/pdf/10.1145/3178876.3186150?casa_token=zul5haircsAAAAAA:iIKn7y-xWwSeqaP-MmmyUaJoJuNZX9Fx1aXeFJwkwtMpVDCrPMW3kZjuYo1LKhSuMeUMNf1mbP2o)
[Kingma et al, 2013] [Kingma, Diederik P., and Max Welling. "Auto-encoding variational bayes." (2013).](https://arxiv.org/pdf/1312.6114.pdf)
[Burgess et al, 2018] [Burgess, Christopher P., et al. "Understanding disentangling in $\beta $-VAE." (2018)](https://arxiv.org/pdf/1804.03599.pdf)
[Higgins et al, 2016] [Higgins, Irina, et al. "beta-vae: Learning basic visual concepts with a constrained variational framework." (2016).](https://openreview.net/pdf?id=Sy2fzU9gl)
[Bowman et al, 2015] [Samuel R. Bowman, Luke Vilnis, Oriol Vinyals, Andrew M. Dai, Rafal Jozefowicz,
and Samy Bengio. 2015. Generating sentences from a continuous space. (2015).](https://arxiv.org/pdf/1511.06349.pdf)
| github_jupyter |
# Demo
The parameters used in this script are defined in `params_demo.py`.
```
import warnings; warnings.simplefilter('ignore')
import os, sys
parentdir = os.path.dirname(os.path.dirname(os.path.realpath("./DeepHalos")))
sys.path.append(parentdir)
import numpy as np
from dlhalos_code import data_processing as tn
from dlhalos_code import CNN
import params_demo as params
import dlhalos_code.data_processing as tn
from pickle import dump
import matplotlib.pyplot as plt
from plots import plot_violins as pv
from plots import plots_for_predictions as pp
path = "./demo-data/"
```
### Prepare the data
```
# Pre-process the simulation
sim_id = ["2"]
s = tn.SimulationPreparation(sim_id, path=path)
# Create a training set by drawing randomly from the simulation 11400 particles
training_set = tn.InputsPreparation(sim_id, shuffle=True, scaler_type="minmax", return_rescaled_outputs=True,
output_range=(-1, 1), log_high_mass_limit=13.4,
load_ids=False, random_style="random", random_subset_all=21400,
path=path)
training_ids = training_set.particle_IDs[:6400]
validation_ids = training_set.particle_IDs[6400:11400]
test_ids = training_set.particle_IDs[11400:]
labels = training_set.labels_particle_IDS
scaler = training_set.scaler_output
# Create the data generators which are used as input to the model
generator_training = tn.DataGenerator(training_ids, labels, s.sims_dic, shuffle=True, **params.params_tr)
generator_validation = tn.DataGenerator(validation_ids, labels, s.sims_dic, shuffle=False, **params.params_val)
generator_test = tn.DataGenerator(test_ids, labels, s.sims_dic, shuffle=False, **params.params_val)
```
### Train the model using our custom Cauchy loss function
```
# Train the model using our custom Cauchy loss function for 30 epochs.
num_epochs = 30
Model = CNN.CNNCauchy(params.param_conv, params.param_fcc, model_type="regression",
training_generator=generator_training, validation_generator=generator_validation,
shuffle=True, num_epochs=num_epochs, dim=generator_training.dim, initialiser="Xavier_uniform",
steps_per_epoch=len(generator_training), validation_steps=len(generator_validation),
max_queue_size=10, use_multiprocessing=False, workers=0, verbose=0, num_gpu=1,
lr=params.lr, save_summary=True, path_summary=params.saving_path, validation_freq=1,
train=True, compile=True, initial_epoch=None, lr_scheduler=False, seed=params.seed)
# Check out what your model looks like
print(Model.model.summary())
```
### Predict the final halo mass label for the validation set
Note: To speed up training, we have selected (a) a small volume for the input box and (b) a small training set size. These choices will affect the accuracy of the final halo mass predictions. For more realistic cases, see the parameter files used for the paper in `DeepHalos/scripts`.
```
# Predict the labels for the validation set from the model after 30 epochs
pred = Model.model.predict_generator(generator_test, use_multiprocessing=False, workers=0, verbose=1)
truth_rescaled = np.array([labels[ID] for ID in test_ids])
h_m_pred = scaler.inverse_transform(pred.reshape(-1, 1)).flatten()
true = scaler.inverse_transform(truth_rescaled.reshape(-1, 1)).flatten()
# Plot the predictions for the validation set as a scatter plot
f, axes = plt.subplots(1, 1, figsize=(8, 6))
axes.plot([11, 13.5], [11, 13.5], color="k")
axes.scatter(true[true>=11], h_m_pred[true>=11], s=1)
axes.set_xlabel(r"$\log \left( M_\mathrm{true}/\mathrm{M}_\odot\right)$")
axes.set_ylabel(r"$\log \left( M_\mathrm{predicted}/\mathrm{M}_\odot\right)$")
# Plot the predictions for the validation set as a violin plot
f1 = pv.plot_violin(true[true>=11], h_m_pred[true>=11], bins_violin=None,
return_stats=None, box=False, alpha=0.5, vert=True, col="C0", figsize=(8, 6))
# Plot the residuals for different mass bins of halos
f1, a, m = pp.plot_histogram_predictions(h_m_pred[true>=11], true[true>=11], radius_bins=False, particle_ids=None, errorbars=False,
label="Raw density", color="C0")
# Plot the likelihood fit
# To run this cell, set "local_machine=True" in line 1 in `dlhalos_code/loss_functions.py`
# and reload the module as here:
import importlib
from dlhalos_code import loss_functions as lf
importlib.reload(lf)
slices = [-0.85, -0.6, 0, 0.5, 0.75, 0.95]
gamma = Model.model.layers[-1].get_weights()[0][0]
f, a = pp.plot_likelihood_distribution(h_m_pred, true, gamma, scaler, bins=np.linspace(-1,1,50), fig=None, axes=None,
title=None, legend=True, slices=slices)
```
| github_jupyter |
<p><font size="6"><b>03 - Pandas: Indexing and selecting data - part I</b></font></p>
> *DS Data manipulation, analysis and visualization in Python*
> *May/June, 2021*
>
> *© 2021, Joris Van den Bossche and Stijn Van Hoey (<mailto:jorisvandenbossche@gmail.com>, <mailto:stijnvanhoey@gmail.com>). Licensed under [CC BY 4.0 Creative Commons](http://creativecommons.org/licenses/by/4.0/)*
---
```
import pandas as pd
# redefining the example DataFrame
data = {'country': ['Belgium', 'France', 'Germany', 'Netherlands', 'United Kingdom'],
'population': [11.3, 64.3, 81.3, 16.9, 64.9],
'area': [30510, 671308, 357050, 41526, 244820],
'capital': ['Brussels', 'Paris', 'Berlin', 'Amsterdam', 'London']}
countries = pd.DataFrame(data)
countries
```
# Subsetting data
## Subset variables (columns)
For a DataFrame, basic indexing selects the columns (cfr. the dictionaries of pure python)
Selecting a **single column**:
```
countries['area'] # single []
```
Remember that the same syntax can also be used to *add* a new columns: `df['new'] = ...`.
We can also select **multiple columns** by passing a list of column names into `[]`:
```
countries[['area', 'population']] # double [[]]
```
## Subset observations (rows)
Using `[]`, slicing or boolean indexing accesses the **rows**:
### Slicing
```
countries[0:4]
```
### Boolean indexing (filtering)
Often, you want to select rows based on a certain condition. This can be done with *'boolean indexing'* (like a where clause in SQL) and comparable to numpy.
The indexer (or boolean mask) should be 1-dimensional and the same length as the thing being indexed.
```
countries['area'] > 100000
countries[countries['area'] > 100000]
countries[countries['population'] > 50]
```
An overview of the possible comparison operations:
Operator | Description
------ | --------
== | Equal
!= | Not equal
\> | Greater than
\>= | Greater than or equal
\< | Lesser than
<= | Lesser than or equal
and to combine multiple conditions:
Operator | Description
------ | --------
& | And (`cond1 & cond2`)
\| | Or (`cond1 \| cond2`)
<div class="alert alert-info" style="font-size:120%">
<b>REMEMBER</b>: <br><br>
So as a summary, `[]` provides the following convenience shortcuts:
* **Series**: selecting a **label**: `s[label]`
* **DataFrame**: selecting a single or multiple **columns**:`df['col']` or `df[['col1', 'col2']]`
* **DataFrame**: slicing or filtering the **rows**: `df['row_label1':'row_label2']` or `df[mask]`
</div>
## Some other useful methods: `isin` and `string` methods
The `isin` method of Series is very useful to select rows that may contain certain values:
```
s = countries['capital']
s.isin?
s.isin(['Berlin', 'London'])
```
This can then be used to filter the dataframe with boolean indexing:
```
countries[countries['capital'].isin(['Berlin', 'London'])]
```
Let's say we want to select all data for which the capital starts with a 'B'. In Python, when having a string, we could use the `startswith` method:
```
string = 'Berlin'
string.startswith('B')
```
In pandas, these are available on a Series through the `str` namespace:
```
countries['capital'].str.startswith('B')
```
For an overview of all string methods, see: https://pandas.pydata.org/pandas-docs/stable/reference/series.html#string-handling
# Exercises using the Titanic dataset
```
df = pd.read_csv("data/titanic.csv")
df.head()
```
<div class="alert alert-success">
<b>EXERCISE</b>:
<ul>
<li>Select all rows for male passengers and calculate the mean age of those passengers. Do the same for the female passengers.</li>
</ul>
</div>
```
# %load _solutions/pandas_03a_selecting_data1.py
# %load _solutions/pandas_03a_selecting_data2.py
# %load _solutions/pandas_03a_selecting_data3.py
```
We will later see an easier way to calculate both averages at the same time with groupby.
<div class="alert alert-success">
<b>EXERCISE</b>:
<ul>
<li>How many passengers older than 70 were on the Titanic?</li>
</ul>
</div>
```
# %load _solutions/pandas_03a_selecting_data4.py
# %load _solutions/pandas_03a_selecting_data5.py
```
<div class="alert alert-success">
<b>EXERCISE</b>:
<ul>
<li>Select the passengers that are between 30 and 40 years old?</li>
</ul>
</div>
```
# %load _solutions/pandas_03a_selecting_data6.py
```
<div class="alert alert-success">
<b>EXERCISE</b>:
Split the 'Name' column on the `,` extract the first part (the surname), and add this as new column 'Surname'.
* Get the first value of the 'Name' column.
* Split this string (check the `split()` method of a string) and get the first element of the resulting list.
* Write the previous step as a function, and 'apply' this function to each element of the 'Name' column (check the `apply()` method of a Series).
</div>
```
# %load _solutions/pandas_03a_selecting_data7.py
# %load _solutions/pandas_03a_selecting_data8.py
# %load _solutions/pandas_03a_selecting_data9.py
# %load _solutions/pandas_03a_selecting_data10.py
# %load _solutions/pandas_03a_selecting_data11.py
# %load _solutions/pandas_03a_selecting_data12.py
# %load _solutions/pandas_03a_selecting_data13.py
# %load _solutions/pandas_03a_selecting_data14.py
```
<div class="alert alert-success">
<b>EXERCISE</b>:
<ul>
<li>Select all passenger that have a surname starting with 'Williams'.</li>
</ul>
</div>
```
# %load _solutions/pandas_03a_selecting_data15.py
```
<div class="alert alert-success">
<b>EXERCISE</b>:
<ul>
<li>Select all rows for the passengers with a surname of more than 15 characters.</li>
</ul>
</div>
```
# %load _solutions/pandas_03a_selecting_data16.py
```
# [OPTIONAL] more exercises
For the quick ones among you, here are some more exercises with some larger dataframe with film data. These exercises are based on the [PyCon tutorial of Brandon Rhodes](https://github.com/brandon-rhodes/pycon-pandas-tutorial/) (so all credit to him!) and the datasets he prepared for that. You can download these data from here: [`titles.csv`](https://drive.google.com/open?id=0B3G70MlBnCgKajNMa1pfSzN6Q3M) and [`cast.csv`](https://drive.google.com/open?id=0B3G70MlBnCgKal9UYTJSR2ZhSW8) and put them in the `/notebooks/data` folder.
```
cast = pd.read_csv('data/cast.csv')
cast.head()
titles = pd.read_csv('data/titles.csv')
titles.head()
```
<div class="alert alert-success">
<b>EXERCISE</b>:
<ul>
<li>How many movies are listed in the titles dataframe?</li>
</ul>
</div>
```
# %load _solutions/pandas_03a_selecting_data17.py
```
<div class="alert alert-success">
<b>EXERCISE</b>:
<ul>
<li>What are the earliest two films listed in the titles dataframe?</li>
</ul>
</div>
```
# %load _solutions/pandas_03a_selecting_data18.py
```
<div class="alert alert-success">
<b>EXERCISE</b>:
<ul>
<li>How many movies have the title "Hamlet"?</li>
</ul>
</div>
```
# %load _solutions/pandas_03a_selecting_data19.py
```
<div class="alert alert-success">
<b>EXERCISE</b>:
<ul>
<li>List all of the "Treasure Island" movies from earliest to most recent.</li>
</ul>
</div>
```
# %load _solutions/pandas_03a_selecting_data20.py
```
<div class="alert alert-success">
<b>EXERCISE</b>:
<ul>
<li>How many movies were made from 1950 through 1959?</li>
</ul>
</div>
```
# %load _solutions/pandas_03a_selecting_data21.py
# %load _solutions/pandas_03a_selecting_data22.py
```
<div class="alert alert-success">
<b>EXERCISE</b>:
<ul>
<li>How many roles in the movie "Inception" are NOT ranked by an "n" value?</li>
</ul>
</div>
```
# %load _solutions/pandas_03a_selecting_data23.py
# %load _solutions/pandas_03a_selecting_data24.py
# %load _solutions/pandas_03a_selecting_data25.py
```
<div class="alert alert-success">
<b>EXERCISE</b>:
<ul>
<li>But how many roles in the movie "Inception" did receive an "n" value?</li>
</ul>
</div>
```
# %load _solutions/pandas_03a_selecting_data26.py
```
<div class="alert alert-success">
<b>EXERCISE</b>:
<ul>
<li>Display the cast of the "Titanic" (the most famous 1997 one) in their correct "n"-value order, ignoring roles that did not earn a numeric "n" value.</li>
</ul>
</div>
```
# %load _solutions/pandas_03a_selecting_data27.py
```
<div class="alert alert-success">
<b>EXERCISE</b>:
<ul>
<li>List the supporting roles (having n=2) played by Brad Pitt in the 1990s, in order by year.</li>
</ul>
</div>
```
# %load _solutions/pandas_03a_selecting_data28.py
```
# Acknowledgement
> The optional exercises are based on the [PyCon tutorial of Brandon Rhodes](https://github.com/brandon-rhodes/pycon-pandas-tutorial/) (so all credit to him!) and the datasets he prepared for that.
---
| github_jupyter |
# Introduction to Data Science – Practical Natural Language Processing (NLP)
*COMP 5360 / MATH 4100, University of Utah, http://datasciencecourse.net/*
In this lecture, we'll do some practical NLP following up on last week's theoretical lecture. We will do some basic text processing followed by a sentiment analysis for movie reviews. For this purpose, we'll introduce the [Natural Language Toolkit (NLTK)](http://www.nltk.org/), a Python library for Natural Language Processing.
We won't cover NLTK or NLP extensively here – this lecture is meant to give you a few pointers if you want to use NLP in the future, e.g., for your project.
Also, there is a well-regarded alternative to NLTK: [Spacy](https://spacy.io/). If you're planning to use a lot of NLP in your project, that might be worth checking out.
**Reading:**
[S. Bird, E. Klein, and E. Loper, *Natural Language Processing with Python – Analyzing Text with the Natural Language Toolkit*](http://www.nltk.org/book/).
[C. Manning and H. Schütze, *Foundations of Statistical Natural Language Processing* (1999).](http://nlp.stanford.edu/fsnlp/)
[D. Jurafsky and J. H. Martin, *Speech and Language Processing* (2016).](https://web.stanford.edu/~jurafsky/slp3/)
**In a prior lecture,** guest lecturer Vivek Srikumar gave a nice overview of Natural Language Processing (NLP). He gave several examples of NLP tasks:
* Part of speech tagging (what are the nouns, verbs, adjectives, prepositions).
+ Information Extraction
+ Sentiment Analysis (determine the attitude of text, e.g., is it positive or negative).
+ Semantic Parsing (translate natural language into a formal meaning representation).
One of the major takeaways from his talk is that the current state-of-the-art for many NLP tasks is to find a good way to represent the text ("extract features") and then to use machine learning / statistics tools, such as classification or clustering.
Our goal today is to use NLTK + scikit-learn to do some basic NLP tasks.
### Install datasets and models
To use NLTK, you must first download and install the datasets and models. Run the following:
```
import nltk
nltk.download('all')
# imports and setup
import numpy as np
from sklearn.neighbors import NearestNeighbors
from sklearn.neighbors import KNeighborsClassifier
from sklearn import svm
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import train_test_split
import matplotlib.pyplot as plt
%matplotlib inline
plt.rcParams['figure.figsize'] = (15, 9)
plt.style.use('ggplot')
```
## Basics of NLTK
We have downloaded a set of text corpora above. Here is a list of these texts:
```
from nltk.book import *
```
Let's look at the first 20 words of text1 – Moby Dick:
```
text1[0:20]
```
### Text Statistics
We can check the length of a text. The text of Moby Dick is 26,0819 words, whereas Monty Python and the Holy Grail has 16,967 words.
```
len(text1)
len(text6)
```
We can check for the frequency of a word. The word "swallow" appears 10 times in Monty Python.
```
text6.count("swallow")
```
We might want to know the context in which "swallow" appears in the text
"You shall know a word by the company it keeps." – John Firth
Use the [`concordance`](http://www.nltk.org/api/nltk.html#nltk.text.Text.concordance) function to print out the words just before and after all occurrences of the word "swallow".
```
text6.concordance("swallow")
```
Words that occur with notable frequencey are "fly" or "flight", "unladen", "air", "African", "European". We can learn about what a swallow can do or properties of a swallow by this.
And if we look for Ishmael in Moby Dick:
```
text1.concordance("Ishmael")
```
Here, we see a lot of "I"s. We could probably infer that it's a person based on that.
We can see what other words frequently appear in the same context using the [`similar`](http://www.nltk.org/api/nltk.html#nltk.text.Text.similar) function.
```
text6.similar("swallow")
text6.similar("african")
text6.similar("coconut")
```
This means that 'african' and 'unladen' both appeared in the text with the same word just before and just after. To see what the phrase is, we can use the [`common_contexts`](http://www.nltk.org/api/nltk.html#nltk.text.Text.concordance) function.
```
text6.common_contexts(["African", "unladen"])
```
We see that both "an unladen swallow" and "an african swallow" appear in the text.
```
text6.concordance("unladen")
print()
text6.concordance("african")
```
### Dispersion plot
`text4` is the Inaugural Address Corpus which includes inaugural addresses going back to 1789.
We can use a dispersion plot to see where in a text certain words appear, and hence how the language of the address has changed over time.
```
text4.dispersion_plot(["citizens", "democracy", "freedom", "duty", "America", "nation", "God"])
```
### Exploring texts using statistics
We'll explore a text by counting the frequency of different words.
The total number of words ("outcomes") in Moby Dick is 260,819 and the number of different words is 19,317.
```
frequency_dist = FreqDist(text1)
print(frequency_dist)
# find 50 most common words
print('\n',frequency_dist.most_common(50))
# not suprisingly, whale occurs quite frequently (906 times!)
print('\n', frequency_dist['whale'])
```
We can find all the words in Moby Dick with more than 15 characters
```
unique_words = set(text1)
long_words = [w.lower() for w in unique_words if len(w) > 15]
long_words
```
### Stopword Removal
Sometimes, it is useful to ignore frequently used words, to concentrate on the meaning of the remaining words. These are referred to as *stopwords*. Examples are "the", "was", "is", etc.
NLTK comes with a stopword corpus.
```
from nltk.corpus import stopwords
stopwords = nltk.corpus.stopwords.words('english')
print(stopwords)
```
Depending on the task, these stopwords are important modifiers, or superfluous content.
### Exercise 1.1: Frequent Words
Find the most frequently used words in Moby Dick that are not stopwords and not punctuation. Hint: [`str.isalpha()`](https://docs.python.org/3/library/stdtypes.html#str.isalpha) could be useful here.
```
# your code here
```
### Stopwords in different corpora
Is there a difference between the frequency in which stopwords appear in the different texts?
```
def content_fraction(text):
stopwords = nltk.corpus.stopwords.words('english')
content = [w for w in text if w.lower() not in stopwords]
return len(content) / len(text)
for i,t in enumerate([text1,text2,text3,text4,text5,text6,text7,text8,text9]):
print(i+1,content_fraction(t))
```
Apparently, "text8: Personals Corpus" has the most content.
### Collocations
A *collocation* is a sequence of words that occur together unusually often, we can retreive these using the [`collocations()`](http://www.nltk.org/api/nltk.html#nltk.text.Text.collocations) function.
```
text1.collocations()
```
## Sentiment analysis for movie reviews
We ask the simple question: Is the attitude of a movie review positive or negative?
How can we approach this question?
Our data is a corpus consisting of 2000 movie reviews together with the user's sentiment polarity (positive or negative). More information about this dataset is available [from this website](https://www.cs.cornell.edu/people/pabo/movie-review-data/).
Our goal is to predict the sentiment polarity from just the review.
Of course, this is something that we can do very easily:
1. That movie was terrible. -> negative
+ That movie was great! -> positive
```
from nltk.corpus import movie_reviews as reviews
```
The datset contains 1000 positive and 1000 negative movie reviews.
The paths to / IDs for the individual reviews are accessible via the fileids() call:
```
reviews.fileids()[0:5]
```
We can access the positives or negatives explicitly:
```
reviews.fileids('pos')[0:5]
```
There are in fact 1000 positive and 1000 negative reviews:
```
num_reviews = len(reviews.fileids())
print(num_reviews)
print(len(reviews.fileids('pos')),len(reviews.fileids('neg')))
```
Let's see the review for the third movie. Its a negative review for [The Mod Squad](https://www.rottentomatoes.com/m/mod_squad/), which has a "rotten" rating on rotten tomatoes.

```
# the name of the file
fid = reviews.fileids()[2]
print(fid)
print('\n', reviews.raw(fid))
print('\n', "The Category:", reviews.categories(fid) )
print('\n', "Individual Words:",reviews.words(fid))
```
Let's look at some sentences that indicate that this is a negative review:
* "it is movies like these that make a jaded movie viewer thankful for the invention of the timex indiglo watch"
* "sounds like a cool movie , does it not ? after the first fifteen minutes , it quickly becomes apparent that it is not ."
* "nothing spectacular"
* "avoid this film at all costs"
### A Custom Algorithm
We'll build a sentiment classifier using methods we already know to predicts the label ['neg', 'pos'] from the review text
`reviews.categories(file_id)` returns the label ['neg', 'pos'] for that movie
```
categories = [reviews.categories(fid) for fid in reviews.fileids()]
labels = {'pos':1, 'neg':0}
# create the labels - 1 for positive, 0 for negative
y = [labels[x[0]] for x in categories]
y[0], y[1000]
```
Here, we collect all words into a nested array datastructure:
```
doc_words = [list(reviews.words(fid)) for fid in reviews.fileids()]
# first 10 words of the third document - mod squad
doc_words[2][1:10]
```
Here we get all of the words in the reviews and make a FreqDist, pick the most common 2000 words and remove the stopwords.
```
# get the 2000 most common words in lowercase
most_common = nltk.FreqDist(w.lower() for w in reviews.words()).most_common(2000)
# remove stopwords
filtered_words = [word_tuple for word_tuple in most_common if word_tuple[0].lower() not in stopwords]
# remove punctuation marks
filtered_words = [word_tuple for word_tuple in filtered_words if word_tuple[0].isalpha()]
print(len(filtered_words))
filtered_words[0:50]
```
We extract this word list from the frequency tuple.
```
word_features = [word_tuple[0] for word_tuple in filtered_words]
len(word_features)
```
We define a function that takes a document and returns a list of zeros and ones indicating which of the words in `word_features` appears in that document.
```
def document_features(document):
document_words = set(document)
features = np.zeros(len(word_features))
for i, word in enumerate(word_features):
features[i] = (word in document_words)
return features
```
Let's just focus on the third document. Which words from `word_features` are in this document?
```
words_in_doc_2 = document_features(doc_words[2])
print(words_in_doc_2)
inds = np.where(words_in_doc_2 == 1)[0]
print('\n', [word_features[i] for i in inds])
```
Now we build our feature set for all the reviews.
```
X = np.zeros([num_reviews,len(word_features)])
for i in range(num_reviews):
X[i,:] = document_features(doc_words[i])
X[0:5]
```
The result is a feature vector for each of these reviews that we can use in classification.
Now that we have features for each document and labels, **we have a classification problem!**
NLTK has a built-in classifier, but we'll use the scikit-learn classifiers we're already familiar with.
Let's try k-nearest neighbors:
```
k = 30
model = KNeighborsClassifier(n_neighbors=k)
scores = cross_val_score(model, X, y, cv=10)
print(scores)
```
And SVM:
```
model = svm.SVC(kernel='rbf', C=30, gamma="auto")
scores = cross_val_score(model, X, y, cv=10)
print(scores)
```
Here we can see that kNN with these parameters is less accurate than SVM, which is about 80% accurate. Of course, we could now use cross validation to find the optimal parameters, `k` and `C`
So, let's see what our algorithm things about the Mod Squad!
```
XTrain, XTest, yTrain, yTest = train_test_split(X, y, random_state=1, test_size=0.2)
model.fit(XTrain, yTrain)
mod_squad = [X[2]]
mod_squad
model.predict(mod_squad)
```
Our model says 0 - so a bad review! We have succesfully build a classifier that can detect the Mod Squad review as a bad review!
Let's take a look at a mis-classified movie. Remember, that the first 1000 movies are negative reviews, so we can just look for the first negative one:
```
model.predict(X[0:10])
```
Review 9, which was misclassified, is for Aberdeen, which has [generally favorable reviews](https://www.rottentomatoes.com/m/aberdeen/) with about 80% positive. Let's looks at the review:
```
fid = reviews.fileids()[8]
print('\n', reviews.raw(fid))
print('\n', reviews.categories(fid) )
```
So if we read this, we can see that this is a negative review, but not a terrible review. Take this sentence for example:
* "if signs & wonders sometimes feels overloaded with ideas , at least it's willing to stretch beyond what we've come to expect from traditional drama"
* "yet this ever-reliable swedish actor adds depth and significance to the otherwise plodding and forgettable aberdeen , a sentimental and painfully mundane european drama"
## We could have also used the Classifier from the NLTK library
Below is the sentiment analysis from [Ch. 6 of the NLTK book](http://www.nltk.org/book/ch06.html).
```
documents = [(list(reviews.words(fileid)), category)
for category in reviews.categories()
for fileid in reviews.fileids(category)]
```
This list contains tuples where the review, stored as an array of words, is the first item in the tuple and the category is the second.
```
documents[1]
```
Extract the features from all of the documents
```
def document_features(document):
document_words = set(document)
features = {}
for word in word_features:
features['contains({})'.format(word)] = (word in document_words)
return features
featuresets = [(document_features(d), c) for (d,c) in documents]
featuresets[2]
```
Split into train_set, test_set and perform classification
```
train_set, test_set = featuresets[100:], featuresets[:100]
classifier = nltk.NaiveBayesClassifier.train(train_set)
print(nltk.classify.accuracy(classifier, test_set))
classifier.show_most_informative_features(10)
```
NLTK gives us 88% accuracy, which isn't bad, but our home-made naive algorithm also achieved a respectable 80%.
What improvements could we have made? Obviously, we could have used more data, or – in our home-grown model select words that discriminate between good and bad reviews. We could have used n-grams, e.g., to catch "not bad" as a postitive sentiment.
| github_jupyter |

[](https://colab.research.google.com/github/JohnSnowLabs/spark-nlp-workshop/blob/master/tutorials/streamlit_notebooks/healthcare/RE_CLINICAL.ipynb)
# **Detect causality between symptoms and treatment**
To run this yourself, you will need to upload your license keys to the notebook. Just Run The Cell Below in order to do that. Also You can open the file explorer on the left side of the screen and upload `license_keys.json` to the folder that opens.
Otherwise, you can look at the example outputs at the bottom of the notebook.
## 1. Colab Setup
Import license keys
```
import os
import json
from google.colab import files
license_keys = files.upload()
with open(list(license_keys.keys())[0]) as f:
license_keys = json.load(f)
sparknlp_version = license_keys["PUBLIC_VERSION"]
jsl_version = license_keys["JSL_VERSION"]
print ('SparkNLP Version:', sparknlp_version)
print ('SparkNLP-JSL Version:', jsl_version)
```
Install dependencies
```
%%capture
for k,v in license_keys.items():
%set_env $k=$v
!wget https://raw.githubusercontent.com/JohnSnowLabs/spark-nlp-workshop/master/jsl_colab_setup.sh
!bash jsl_colab_setup.sh
# Install Spark NLP Display for visualization
!pip install --ignore-installed spark-nlp-display
```
Import dependencies into Python
```
import pandas as pd
from pyspark.ml import Pipeline
from pyspark.sql import SparkSession
import pyspark.sql.functions as F
from tabulate import tabulate
import sparknlp
from sparknlp.annotator import *
from sparknlp_jsl.annotator import *
from sparknlp.base import *
import sparknlp_jsl
```
Start the Spark session
```
spark = sparknlp_jsl.start(license_keys['SECRET'])
# manually start session
# params = {"spark.driver.memory" : "16G",
# "spark.kryoserializer.buffer.max" : "2000M",
# "spark.driver.maxResultSize" : "2000M"}
# spark = sparknlp_jsl.start(license_keys['SECRET'],params=params)
```
## 2. Select the Relation Extraction model and construct the pipeline
Select the models:
* Clinical Relation Extraction models: **re_clinical**
For more details: https://github.com/JohnSnowLabs/spark-nlp-models#pretrained-models---spark-nlp-for-healthcare
```
# Change this to the model you want to use and re-run the cells below.
RE_MODEL_NAME = "re_clinical"
NER_MODEL_NAME = "ner_clinical"
```
Create the pipeline
```
document_assembler = DocumentAssembler() \
.setInputCol('text')\
.setOutputCol('document')
sentence_detector = SentenceDetector() \
.setInputCols(['document'])\
.setOutputCol('sentences')
tokenizer = Tokenizer()\
.setInputCols(['sentences']) \
.setOutputCol('tokens')
pos_tagger = PerceptronModel()\
.pretrained("pos_clinical", "en", "clinical/models") \
.setInputCols(["sentences", "tokens"])\
.setOutputCol("pos_tags")
dependency_parser = DependencyParserModel()\
.pretrained("dependency_conllu", "en")\
.setInputCols(["sentences", "pos_tags", "tokens"])\
.setOutputCol("dependencies")
embeddings = WordEmbeddingsModel.pretrained('embeddings_clinical', 'en', 'clinical/models')\
.setInputCols(["sentences", "tokens"])\
.setOutputCol("embeddings")
clinical_ner_model = MedicalNerModel.pretrained(NER_MODEL_NAME, "en", "clinical/models") \
.setInputCols(["sentences", "tokens", "embeddings"])\
.setOutputCol("clinical_ner_tags")
clinical_ner_chunker = NerConverter()\
.setInputCols(["sentences", "tokens", "clinical_ner_tags"])\
.setOutputCol("clinical_ner_chunks")
clinical_re_Model = RelationExtractionModel()\
.pretrained(RE_MODEL_NAME, 'en', 'clinical/models')\
.setInputCols(["embeddings", "pos_tags", "clinical_ner_chunks", "dependencies"])\
.setOutputCol("clinical_relations")\
.setMaxSyntacticDistance(4)
#.setRelationPairs()#["problem-test", "problem-treatment"]) # we can set the possible relation pairs (if not set, all the relations will be calculated)
pipeline = Pipeline(stages=[
document_assembler,
sentence_detector,
tokenizer,
pos_tagger,
dependency_parser,
embeddings,
clinical_ner_model,
clinical_ner_chunker,
clinical_re_Model])
empty_df = spark.createDataFrame([['']]).toDF("text")
pipeline_model = pipeline.fit(empty_df)
light_pipeline = LightPipeline(pipeline_model)
```
## 3. Create example inputs
```
# Enter examples as strings in this array
input_list = [
"""She is followed by Dr. X in our office and has a history of severe tricuspid regurgitation with mild elevation and PA pressure. On 05/12/08, preserved left and right ventricular systolic function, aortic sclerosis with apparent mild aortic stenosis, and bi-atrial enlargement. She has previously had a Persantine Myoview nuclear rest-stress test scan completed at ABCD Medical Center in 07/06 that was negative. She has had significant mitral valve regurgitation in the past being moderate, but on the most recent echocardiogram on 05/12/08, that was not felt to be significant. She has a history of hypertension and EKGs in our office show normal sinus rhythm with frequent APCs versus wandering atrial pacemaker. She does have a history of significant hypertension in the past. She has had dizzy spells and denies clearly any true syncope. She has had bradycardia in the past from beta-blocker therapy."""
]
```
# 4. Run the pipeline
```
df = spark.createDataFrame(pd.DataFrame({"text": input_list}))
result = pipeline_model.transform(df)
light_result = light_pipeline.fullAnnotate(input_list[0])
```
# 5. Visualize
```
from sparknlp_display import RelationExtractionVisualizer
vis = RelationExtractionVisualizer()
vis.display(light_result[0], 'clinical_relations', show_relations=True) # default show_relations: True
```
| github_jupyter |
## ANALYSING RETURNS - Module 1 Graded Quiz
##### salimt
```
import pandas as pd
import numpy as np
import sys
!{sys.executable} -m pip install matplotlib
```
## Annualizing Returns
To annualize a return for a period, you compound the return for as many times as there are periods in a year. For instance, to annualize a monthly return you compund that return 12 times. The formula to annualize a monthly return $R_m$ is:
$$ (1+R_m)^{12} - 1$$
To annualize a quarterly return $R_q$ you would get:
$$ (1+R_q)^{4} - 1$$
And finally, to annualize a daily return $R_d$ you would get:
$$ (1+R_d)^{252} - 1$$
**The Annualized Return and Annualized Volatility of the Lo 20 AND Hi 20 portfolios over the entire period**
```
prices = pd.read_csv("data\\Portfolios_Formed_on_ME_monthly_EW.csv",header = 0, index_col=0, parse_dates=True,
na_values=-99.99)
prices.index = pd.to_datetime(prices.index, format="%Y%m")
prices.index = prices.index.to_period('M')
prices.head()
returns = prices[['Lo 20', 'Hi 20']]
returns = returns/100
returns.head()
n_months = returns.shape[0]
return_per_month = (returns+1).prod()**(1/n_months) - 1
annualized_return = (return_per_month + 1)**12-1
annualized_return
annualized_vol = returns.std()*np.sqrt(12)
annualized_vol
```
**The Annualized Return and Annualized Volatility of the Lo 20 AND Hi20 portfolios over the period 1999 - 2015 (both inclusive)**
```
return9915 = returns["1999":"2015"]
n_months = return9915.shape[0]
return_per_month = (return9915+1).prod()**(1/n_months) - 1
annualized_return9915 = (return_per_month + 1)**12-1
annualized_return9915
annualized_vol9915 = return9915.std()*np.sqrt(12)
annualized_vol9915
return9915.plot()
```
**The Max Drawdown (expressed as a positive number) experienced over the 1999-2015 period in the SmallCap (Lo 20) portfolio**
```
wealth_index = 1000*(1+return9915).cumprod()
wealth_index_max = wealth_index.min()
wealth_index.plot()
def drawdown(return_series: pd.Series):
"""Takes a time series of asset returns.
returns a DataFrame with columns for
the wealth index,
the previous peaks, and
the percentage drawdown
"""
wealth_index = 1000*(1+return_series).cumprod()
previous_peaks = wealth_index.cummax()
drawdowns = (wealth_index - previous_peaks)/previous_peaks
return pd.DataFrame({"Wealth": wealth_index,
"Previous Peak": previous_peaks,
"Drawdown": drawdowns})
drawdown(return9915["Lo 20"])["Drawdown"].min()
drawdown(return9915["Lo 20"])["Drawdown"].idxmin()
drawdown(return9915["Hi 20"])["Drawdown"].min()
drawdown(return9915["Hi 20"])["Drawdown"].idxmin()
```
**Since 2009 (including all of 2009) through 2018 which Hedge Fund Index has exhibited the highest semideviation**
```
prices = pd.read_csv("data\\edhec-hedgefundindices.csv",header = 0, index_col=0, parse_dates=True,
na_values=-99.99)
prices.index = pd.to_datetime(prices.index, format="%Y%m")
prices.index = prices.index.to_period('M')
prices.head()
prices09 = prices["2009":]
prices09.head()
def semideviation(r):
"""
Returns the semideviation aka negative semideviation of r
r must be a Series or a DataFrame, else raises a TypeError
"""
is_negative = r < 0
return r[is_negative].std(ddof=0)
semidev = semideviation(prices09)
semidev.sort_values(ascending=False)
semidev.idxmax()
semidev.idxmin()
def skewness_kurtosis(r, deg):
"""
Alternative to scipy.stats.skew()
Computes the skewness of the supplied Series or DataFrame
Returns a float or a Series
"""
demeaned_r = r - r.mean()
# use the population standard deviation, so set dof=0
sigma_r = r.std(ddof=0)
exp = (demeaned_r**deg).mean()
return exp/sigma_r**deg
skew = skewness_kurtosis(prices09,3)
skew.sort_values(ascending=False)
kurtosis_d = skewness_kurtosis(prices00,4)
kurtosis_d.sort_values(ascending=False)
prices00 = prices["2000":]
prices00.head()
```
| github_jupyter |
# TensorFlow Tutorial
Welcome to this week's programming assignment. Until now, you've always used numpy to build neural networks. Now we will step you through a deep learning framework that will allow you to build neural networks more easily. Machine learning frameworks like TensorFlow, PaddlePaddle, Torch, Caffe, Keras, and many others can speed up your machine learning development significantly. All of these frameworks also have a lot of documentation, which you should feel free to read. In this assignment, you will learn to do the following in TensorFlow:
- Initialize variables
- Start your own session
- Train algorithms
- Implement a Neural Network
Programing frameworks can not only shorten your coding time, but sometimes also perform optimizations that speed up your code.
## 1 - Exploring the Tensorflow Library
To start, you will import the library:
```
import math
import numpy as np
import h5py
import matplotlib.pyplot as plt
import tensorflow as tf
from tensorflow.python.framework import ops
from tf_utils import load_dataset, random_mini_batches, convert_to_one_hot, predict
%matplotlib inline
np.random.seed(1)
```
Now that you have imported the library, we will walk you through its different applications. You will start with an example, where we compute for you the loss of one training example.
$$loss = \mathcal{L}(\hat{y}, y) = (\hat y^{(i)} - y^{(i)})^2 \tag{1}$$
```
y_hat = tf.constant(36, name='y_hat') # Define y_hat constant. Set to 36.
y = tf.constant(39, name='y') # Define y. Set to 39
loss = tf.Variable((y - y_hat)**2, name='loss') # Create a variable for the loss
init = tf.global_variables_initializer() # When init is run later (session.run(init)),
# the loss variable will be initialized and ready to be computed
with tf.Session() as session: # Create a session and print the output
session.run(init) # Initializes the variables
print(session.run(loss)) # Prints the loss
```
Writing and running programs in TensorFlow has the following steps:
1. Create Tensors (variables) that are not yet executed/evaluated.
2. Write operations between those Tensors.
3. Initialize your Tensors.
4. Create a Session.
5. Run the Session. This will run the operations you'd written above.
Therefore, when we created a variable for the loss, we simply defined the loss as a function of other quantities, but did not evaluate its value. To evaluate it, we had to run `init=tf.global_variables_initializer()`. That initialized the loss variable, and in the last line we were finally able to evaluate the value of `loss` and print its value.
Now let us look at an easy example. Run the cell below:
```
a = tf.constant(2)
b = tf.constant(10)
c = tf.multiply(a,b)
print(c)
```
As expected, you will not see 20! You got a tensor saying that the result is a tensor that does not have the shape attribute, and is of type "int32". All you did was put in the 'computation graph', but you have not run this computation yet. In order to actually multiply the two numbers, you will have to create a session and run it.
```
sess = tf.Session()
print(sess.run(c))
```
Great! To summarize, **remember to initialize your variables, create a session and run the operations inside the session**.
Next, you'll also have to know about placeholders. A placeholder is an object whose value you can specify only later.
To specify values for a placeholder, you can pass in values by using a "feed dictionary" (`feed_dict` variable). Below, we created a placeholder for x. This allows us to pass in a number later when we run the session.
```
# Change the value of x in the feed_dict
x = tf.placeholder(tf.int64, name = 'x')
print(sess.run(2 * x, feed_dict = {x: 7}))
sess.close()
```
When you first defined `x` you did not have to specify a value for it. A placeholder is simply a variable that you will assign data to only later, when running the session. We say that you **feed data** to these placeholders when running the session.
Here's what's happening: When you specify the operations needed for a computation, you are telling TensorFlow how to construct a computation graph. The computation graph can have some placeholders whose values you will specify only later. Finally, when you run the session, you are telling TensorFlow to execute the computation graph.
### 1.1 - Linear function
Lets start this programming exercise by computing the following equation: $Y = WX + b$, where $W$ and $X$ are random matrices and b is a random vector.
**Exercise**: Compute $WX + b$ where $W, X$, and $b$ are drawn from a random normal distribution. W is of shape (4, 3), X is (3,1) and b is (4,1). As an example, here is how you would define a constant X that has shape (3,1):
```python
X = tf.constant(np.random.randn(3,1), name = "X")
```
You might find the following functions helpful:
- tf.matmul(..., ...) to do a matrix multiplication
- tf.add(..., ...) to do an addition
- np.random.randn(...) to initialize randomly
```
# GRADED FUNCTION: linear_function
def linear_function():
"""
Implements a linear function:
Initializes W to be a random tensor of shape (4,3)
Initializes X to be a random tensor of shape (3,1)
Initializes b to be a random tensor of shape (4,1)
Returns:
result -- runs the session for Y = WX + b
"""
np.random.seed(1)
### START CODE HERE ### (4 lines of code)
X = tf.constant(np.random.randn(3,1), name = "X")
W = tf.constant(np.random.randn(4,3), name = "W")
b = tf.constant(np.random.randn(4,1), name = "b")
Y = tf.add(tf.matmul(W, X), b)
### END CODE HERE ###
# Create the session using tf.Session() and run it with sess.run(...) on the variable you want to calculate
### START CODE HERE ###
sess = tf.Session()
result = sess.run(Y)
### END CODE HERE ###
# close the session
sess.close()
return result
print( "result = " + str(linear_function()))
```
*** Expected Output ***:
<table>
<tr>
<td>
**result**
</td>
<td>
[[-2.15657382]
[ 2.95891446]
[-1.08926781]
[-0.84538042]]
</td>
</tr>
</table>
### 1.2 - Computing the sigmoid
Great! You just implemented a linear function. Tensorflow offers a variety of commonly used neural network functions like `tf.sigmoid` and `tf.softmax`. For this exercise lets compute the sigmoid function of an input.
You will do this exercise using a placeholder variable `x`. When running the session, you should use the feed dictionary to pass in the input `z`. In this exercise, you will have to (i) create a placeholder `x`, (ii) define the operations needed to compute the sigmoid using `tf.sigmoid`, and then (iii) run the session.
** Exercise **: Implement the sigmoid function below. You should use the following:
- `tf.placeholder(tf.float32, name = "...")`
- `tf.sigmoid(...)`
- `sess.run(..., feed_dict = {x: z})`
Note that there are two typical ways to create and use sessions in tensorflow:
**Method 1:**
```python
sess = tf.Session()
# Run the variables initialization (if needed), run the operations
result = sess.run(..., feed_dict = {...})
sess.close() # Close the session
```
**Method 2:**
```python
with tf.Session() as sess:
# run the variables initialization (if needed), run the operations
result = sess.run(..., feed_dict = {...})
# This takes care of closing the session for you :)
```
```
# GRADED FUNCTION: sigmoid
def sigmoid(z):
"""
Computes the sigmoid of z
Arguments:
z -- input value, scalar or vector
Returns:
results -- the sigmoid of z
"""
### START CODE HERE ### ( approx. 4 lines of code)
# Create a placeholder for x. Name it 'x'.
x = tf.placeholder(tf.float32, name = "x")
# compute sigmoid(x)
sigmoid = tf.sigmoid(x)
# Create a session, and run it. Please use the method 2 explained above.
# You should use a feed_dict to pass z's value to x.
with tf.Session() as sess :
# Run session and call the output "result"
result = sess.run(sigmoid, feed_dict = {x: z})
### END CODE HERE ###
return result
print ("sigmoid(0) = " + str(sigmoid(0)))
print ("sigmoid(12) = " + str(sigmoid(12)))
```
*** Expected Output ***:
<table>
<tr>
<td>
**sigmoid(0)**
</td>
<td>
0.5
</td>
</tr>
<tr>
<td>
**sigmoid(12)**
</td>
<td>
0.999994
</td>
</tr>
</table>
<font color='blue'>
**To summarize, you how know how to**:
1. Create placeholders
2. Specify the computation graph corresponding to operations you want to compute
3. Create the session
4. Run the session, using a feed dictionary if necessary to specify placeholder variables' values.
### 1.3 - Computing the Cost
You can also use a built-in function to compute the cost of your neural network. So instead of needing to write code to compute this as a function of $a^{[2](i)}$ and $y^{(i)}$ for i=1...m:
$$ J = - \frac{1}{m} \sum_{i = 1}^m \large ( \small y^{(i)} \log a^{ [2] (i)} + (1-y^{(i)})\log (1-a^{ [2] (i)} )\large )\small\tag{2}$$
you can do it in one line of code in tensorflow!
**Exercise**: Implement the cross entropy loss. The function you will use is:
- `tf.nn.sigmoid_cross_entropy_with_logits(logits = ..., labels = ...)`
Your code should input `z`, compute the sigmoid (to get `a`) and then compute the cross entropy cost $J$. All this can be done using one call to `tf.nn.sigmoid_cross_entropy_with_logits`, which computes
$$- \frac{1}{m} \sum_{i = 1}^m \large ( \small y^{(i)} \log \sigma(z^{[2](i)}) + (1-y^{(i)})\log (1-\sigma(z^{[2](i)})\large )\small\tag{2}$$
```
# GRADED FUNCTION: cost
def cost(logits, labels):
"""
Computes the cost using the sigmoid cross entropy
Arguments:
logits -- vector containing z, output of the last linear unit (before the final sigmoid activation)
labels -- vector of labels y (1 or 0)
Note: What we've been calling "z" and "y" in this class are respectively called "logits" and "labels"
in the TensorFlow documentation. So logits will feed into z, and labels into y.
Returns:
cost -- runs the session of the cost (formula (2))
"""
### START CODE HERE ###
# Create the placeholders for "logits" (z) and "labels" (y) (approx. 2 lines)
z = tf.placeholder(tf.float32, name = "z")
y = tf.placeholder(tf.float32, name = "y")
# Use the loss function (approx. 1 line)
cost = tf.nn.sigmoid_cross_entropy_with_logits(logits = z, labels = y)
# Create a session (approx. 1 line). See method 1 above.
sess = tf.Session()
# Run the session (approx. 1 line).
cost = sess.run(cost, feed_dict = {z : logits, y : labels})
# Close the session (approx. 1 line). See method 1 above.
sess.close()
### END CODE HERE ###
return cost
logits = sigmoid(np.array([0.2,0.4,0.7,0.9]))
cost = cost(logits, np.array([0,0,1,1]))
print ("cost = " + str(cost))
```
** Expected Output** :
<table>
<tr>
<td>
**cost**
</td>
<td>
[ 1.00538719 1.03664088 0.41385433 0.39956614]
</td>
</tr>
</table>
### 1.4 - Using One Hot encodings
Many times in deep learning you will have a y vector with numbers ranging from 0 to C-1, where C is the number of classes. If C is for example 4, then you might have the following y vector which you will need to convert as follows:
<img src="images/onehot.png" style="width:600px;height:150px;">
This is called a "one hot" encoding, because in the converted representation exactly one element of each column is "hot" (meaning set to 1). To do this conversion in numpy, you might have to write a few lines of code. In tensorflow, you can use one line of code:
- tf.one_hot(labels, depth, axis)
**Exercise:** Implement the function below to take one vector of labels and the total number of classes $C$, and return the one hot encoding. Use `tf.one_hot()` to do this.
```
# GRADED FUNCTION: one_hot_matrix
def one_hot_matrix(labels, C):
"""
Creates a matrix where the i-th row corresponds to the ith class number and the jth column
corresponds to the jth training example. So if example j had a label i. Then entry (i,j)
will be 1.
Arguments:
labels -- vector containing the labels
C -- number of classes, the depth of the one hot dimension
Returns:
one_hot -- one hot matrix
"""
### START CODE HERE ###
# Create a tf.constant equal to C (depth), name it 'C'. (approx. 1 line)
C = tf.constant(C, name = "C")
# Use tf.one_hot, be careful with the axis (approx. 1 line)
one_hot_matrix = tf.one_hot(indices = labels, depth = C, axis = 0)
# Create the session (approx. 1 line)
sess = tf.Session()
# Run the session (approx. 1 line)
one_hot = sess.run(one_hot_matrix)
# Close the session (approx. 1 line). See method 1 above.
sess.close()
### END CODE HERE ###
return one_hot
labels = np.array([1,2,3,0,2,1])
one_hot = one_hot_matrix(labels, C = 4)
print ("one_hot = " + str(one_hot))
```
**Expected Output**:
<table>
<tr>
<td>
**one_hot**
</td>
<td>
[[ 0. 0. 0. 1. 0. 0.]
[ 1. 0. 0. 0. 0. 1.]
[ 0. 1. 0. 0. 1. 0.]
[ 0. 0. 1. 0. 0. 0.]]
</td>
</tr>
</table>
### 1.5 - Initialize with zeros and ones
Now you will learn how to initialize a vector of zeros and ones. The function you will be calling is `tf.ones()`. To initialize with zeros you could use tf.zeros() instead. These functions take in a shape and return an array of dimension shape full of zeros and ones respectively.
**Exercise:** Implement the function below to take in a shape and to return an array (of the shape's dimension of ones).
- tf.ones(shape)
```
# GRADED FUNCTION: ones
def ones(shape):
"""
Creates an array of ones of dimension shape
Arguments:
shape -- shape of the array you want to create
Returns:
ones -- array containing only ones
"""
### START CODE HERE ###
# Create "ones" tensor using tf.ones(...). (approx. 1 line)
ones = tf.ones(shape)
# Create the session (approx. 1 line)
sess = tf.Session()
# Run the session to compute 'ones' (approx. 1 line)
ones = sess.run(ones)
# Close the session (approx. 1 line). See method 1 above.
sess.close()
### END CODE HERE ###
return ones
print ("ones = " + str(ones([3])))
```
**Expected Output:**
<table>
<tr>
<td>
**ones**
</td>
<td>
[ 1. 1. 1.]
</td>
</tr>
</table>
# 2 - Building your first neural network in tensorflow
In this part of the assignment you will build a neural network using tensorflow. Remember that there are two parts to implement a tensorflow model:
- Create the computation graph
- Run the graph
Let's delve into the problem you'd like to solve!
### 2.0 - Problem statement: SIGNS Dataset
One afternoon, with some friends we decided to teach our computers to decipher sign language. We spent a few hours taking pictures in front of a white wall and came up with the following dataset. It's now your job to build an algorithm that would facilitate communications from a speech-impaired person to someone who doesn't understand sign language.
- **Training set**: 1080 pictures (64 by 64 pixels) of signs representing numbers from 0 to 5 (180 pictures per number).
- **Test set**: 120 pictures (64 by 64 pixels) of signs representing numbers from 0 to 5 (20 pictures per number).
Note that this is a subset of the SIGNS dataset. The complete dataset contains many more signs.
Here are examples for each number, and how an explanation of how we represent the labels. These are the original pictures, before we lowered the image resolutoion to 64 by 64 pixels.
<img src="images/hands.png" style="width:800px;height:350px;"><caption><center> <u><font color='purple'> **Figure 1**</u><font color='purple'>: SIGNS dataset <br> <font color='black'> </center>
Run the following code to load the dataset.
```
# Loading the dataset
X_train_orig, Y_train_orig, X_test_orig, Y_test_orig, classes = load_dataset()
```
Change the index below and run the cell to visualize some examples in the dataset.
```
# Example of a picture
index = 0
plt.imshow(X_train_orig[index])
print ("y = " + str(np.squeeze(Y_train_orig[:, index])))
```
As usual you flatten the image dataset, then normalize it by dividing by 255. On top of that, you will convert each label to a one-hot vector as shown in Figure 1. Run the cell below to do so.
```
# Flatten the training and test images
X_train_flatten = X_train_orig.reshape(X_train_orig.shape[0], -1).T
X_test_flatten = X_test_orig.reshape(X_test_orig.shape[0], -1).T
# Normalize image vectors
X_train = X_train_flatten/255.
X_test = X_test_flatten/255.
# Convert training and test labels to one hot matrices
Y_train = convert_to_one_hot(Y_train_orig, 6)
Y_test = convert_to_one_hot(Y_test_orig, 6)
print ("number of training examples = " + str(X_train.shape[1]))
print ("number of test examples = " + str(X_test.shape[1]))
print ("X_train shape: " + str(X_train.shape))
print ("Y_train shape: " + str(Y_train.shape))
print ("X_test shape: " + str(X_test.shape))
print ("Y_test shape: " + str(Y_test.shape))
```
**Note** that 12288 comes from $64 \times 64 \times 3$. Each image is square, 64 by 64 pixels, and 3 is for the RGB colors. Please make sure all these shapes make sense to you before continuing.
**Your goal** is to build an algorithm capable of recognizing a sign with high accuracy. To do so, you are going to build a tensorflow model that is almost the same as one you have previously built in numpy for cat recognition (but now using a softmax output). It is a great occasion to compare your numpy implementation to the tensorflow one.
**The model** is *LINEAR -> RELU -> LINEAR -> RELU -> LINEAR -> SOFTMAX*. The SIGMOID output layer has been converted to a SOFTMAX. A SOFTMAX layer generalizes SIGMOID to when there are more than two classes.
### 2.1 - Create placeholders
Your first task is to create placeholders for `X` and `Y`. This will allow you to later pass your training data in when you run your session.
**Exercise:** Implement the function below to create the placeholders in tensorflow.
```
# GRADED FUNCTION: create_placeholders
def create_placeholders(n_x, n_y):
"""
Creates the placeholders for the tensorflow session.
Arguments:
n_x -- scalar, size of an image vector (num_px * num_px = 64 * 64 * 3 = 12288)
n_y -- scalar, number of classes (from 0 to 5, so -> 6)
Returns:
X -- placeholder for the data input, of shape [n_x, None] and dtype "float"
Y -- placeholder for the input labels, of shape [n_y, None] and dtype "float"
Tips:
- You will use None because it let's us be flexible on the number of examples you will for the placeholders.
In fact, the number of examples during test/train is different.
"""
### START CODE HERE ### (approx. 2 lines)
X = tf.placeholder(tf.float32, [n_x, None], name = "X")
Y = tf.placeholder(tf.float32, [n_y, None], name = "Y")
### END CODE HERE ###
return X, Y
X, Y = create_placeholders(12288, 6)
print ("X = " + str(X))
print ("Y = " + str(Y))
```
**Expected Output**:
<table>
<tr>
<td>
**X**
</td>
<td>
Tensor("Placeholder_1:0", shape=(12288, ?), dtype=float32) (not necessarily Placeholder_1)
</td>
</tr>
<tr>
<td>
**Y**
</td>
<td>
Tensor("Placeholder_2:0", shape=(10, ?), dtype=float32) (not necessarily Placeholder_2)
</td>
</tr>
</table>
### 2.2 - Initializing the parameters
Your second task is to initialize the parameters in tensorflow.
**Exercise:** Implement the function below to initialize the parameters in tensorflow. You are going use Xavier Initialization for weights and Zero Initialization for biases. The shapes are given below. As an example, to help you, for W1 and b1 you could use:
```python
W1 = tf.get_variable("W1", [25,12288], initializer = tf.contrib.layers.xavier_initializer(seed = 1))
b1 = tf.get_variable("b1", [25,1], initializer = tf.zeros_initializer())
```
Please use `seed = 1` to make sure your results match ours.
```
# GRADED FUNCTION: initialize_parameters
def initialize_parameters():
"""
Initializes parameters to build a neural network with tensorflow. The shapes are:
W1 : [25, 12288]
b1 : [25, 1]
W2 : [12, 25]
b2 : [12, 1]
W3 : [6, 12]
b3 : [6, 1]
Returns:
parameters -- a dictionary of tensors containing W1, b1, W2, b2, W3, b3
"""
tf.set_random_seed(1) # so that your "random" numbers match ours
### START CODE HERE ### (approx. 6 lines of code)
W1 = tf.get_variable("W1", [25,12288], initializer = tf.contrib.layers.xavier_initializer(seed = 1))
b1 = tf.get_variable("b1", [25,1], initializer = tf.zeros_initializer())
W2 = tf.get_variable("W2", [12,25], initializer = tf.contrib.layers.xavier_initializer(seed = 1))
b2 = tf.get_variable("b2", [12,1], initializer = tf.zeros_initializer())
W3 = tf.get_variable("W3", [6,12], initializer = tf.contrib.layers.xavier_initializer(seed = 1))
b3 = tf.get_variable("b3", [6,1], initializer = tf.zeros_initializer())
### END CODE HERE ###
parameters = {"W1": W1,
"b1": b1,
"W2": W2,
"b2": b2,
"W3": W3,
"b3": b3}
return parameters
tf.reset_default_graph()
with tf.Session() as sess:
parameters = initialize_parameters()
print("W1 = " + str(parameters["W1"]))
print("b1 = " + str(parameters["b1"]))
print("W2 = " + str(parameters["W2"]))
print("b2 = " + str(parameters["b2"]))
```
**Expected Output**:
<table>
<tr>
<td>
**W1**
</td>
<td>
< tf.Variable 'W1:0' shape=(25, 12288) dtype=float32_ref >
</td>
</tr>
<tr>
<td>
**b1**
</td>
<td>
< tf.Variable 'b1:0' shape=(25, 1) dtype=float32_ref >
</td>
</tr>
<tr>
<td>
**W2**
</td>
<td>
< tf.Variable 'W2:0' shape=(12, 25) dtype=float32_ref >
</td>
</tr>
<tr>
<td>
**b2**
</td>
<td>
< tf.Variable 'b2:0' shape=(12, 1) dtype=float32_ref >
</td>
</tr>
</table>
As expected, the parameters haven't been evaluated yet.
### 2.3 - Forward propagation in tensorflow
You will now implement the forward propagation module in tensorflow. The function will take in a dictionary of parameters and it will complete the forward pass. The functions you will be using are:
- `tf.add(...,...)` to do an addition
- `tf.matmul(...,...)` to do a matrix multiplication
- `tf.nn.relu(...)` to apply the ReLU activation
**Question:** Implement the forward pass of the neural network. We commented for you the numpy equivalents so that you can compare the tensorflow implementation to numpy. It is important to note that the forward propagation stops at `z3`. The reason is that in tensorflow the last linear layer output is given as input to the function computing the loss. Therefore, you don't need `a3`!
```
# GRADED FUNCTION: forward_propagation
def forward_propagation(X, parameters):
"""
Implements the forward propagation for the model: LINEAR -> RELU -> LINEAR -> RELU -> LINEAR -> SOFTMAX
Arguments:
X -- input dataset placeholder, of shape (input size, number of examples)
parameters -- python dictionary containing your parameters "W1", "b1", "W2", "b2", "W3", "b3"
the shapes are given in initialize_parameters
Returns:
Z3 -- the output of the last LINEAR unit
"""
# Retrieve the parameters from the dictionary "parameters"
W1 = parameters['W1']
b1 = parameters['b1']
W2 = parameters['W2']
b2 = parameters['b2']
W3 = parameters['W3']
b3 = parameters['b3']
### START CODE HERE ### (approx. 5 lines) # Numpy Equivalents:
Z1 = tf.add(tf.matmul(W1, X), b1) # Z1 = np.dot(W1, X) + b1
A1 = tf.nn.relu(Z1) # A1 = relu(Z1)
Z2 = tf.add(tf.matmul(W2, A1), b2) # Z2 = np.dot(W2, a1) + b2
A2 = tf.nn.relu(Z2) # A2 = relu(Z2)
Z3 = tf.add(tf.matmul(W3, A2), b3) # Z3 = np.dot(W3,Z2) + b3
### END CODE HERE ###
return Z3
tf.reset_default_graph()
with tf.Session() as sess:
X, Y = create_placeholders(12288, 6)
parameters = initialize_parameters()
Z3 = forward_propagation(X, parameters)
print("Z3 = " + str(Z3))
```
**Expected Output**:
<table>
<tr>
<td>
**Z3**
</td>
<td>
Tensor("Add_2:0", shape=(6, ?), dtype=float32)
</td>
</tr>
</table>
You may have noticed that the forward propagation doesn't output any cache. You will understand why below, when we get to brackpropagation.
### 2.4 Compute cost
As seen before, it is very easy to compute the cost using:
```python
tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(logits = ..., labels = ...))
```
**Question**: Implement the cost function below.
- It is important to know that the "`logits`" and "`labels`" inputs of `tf.nn.softmax_cross_entropy_with_logits` are expected to be of shape (number of examples, num_classes). We have thus transposed Z3 and Y for you.
- Besides, `tf.reduce_mean` basically does the summation over the examples.
```
# GRADED FUNCTION: compute_cost
def compute_cost(Z3, Y):
"""
Computes the cost
Arguments:
Z3 -- output of forward propagation (output of the last LINEAR unit), of shape (6, number of examples)
Y -- "true" labels vector placeholder, same shape as Z3
Returns:
cost - Tensor of the cost function
"""
# to fit the tensorflow requirement for tf.nn.softmax_cross_entropy_with_logits(...,...)
logits = tf.transpose(Z3)
labels = tf.transpose(Y)
### START CODE HERE ### (1 line of code)
cost = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(logits = logits, labels = labels))
### END CODE HERE ###
return cost
tf.reset_default_graph()
with tf.Session() as sess:
X, Y = create_placeholders(12288, 6)
parameters = initialize_parameters()
Z3 = forward_propagation(X, parameters)
cost = compute_cost(Z3, Y)
print("cost = " + str(cost))
```
**Expected Output**:
<table>
<tr>
<td>
**cost**
</td>
<td>
Tensor("Mean:0", shape=(), dtype=float32)
</td>
</tr>
</table>
### 2.5 - Backward propagation & parameter updates
This is where you become grateful to programming frameworks. All the backpropagation and the parameters update is taken care of in 1 line of code. It is very easy to incorporate this line in the model.
After you compute the cost function. You will create an "`optimizer`" object. You have to call this object along with the cost when running the tf.session. When called, it will perform an optimization on the given cost with the chosen method and learning rate.
For instance, for gradient descent the optimizer would be:
```python
optimizer = tf.train.GradientDescentOptimizer(learning_rate = learning_rate).minimize(cost)
```
To make the optimization you would do:
```python
_ , c = sess.run([optimizer, cost], feed_dict={X: minibatch_X, Y: minibatch_Y})
```
This computes the backpropagation by passing through the tensorflow graph in the reverse order. From cost to inputs.
**Note** When coding, we often use `_` as a "throwaway" variable to store values that we won't need to use later. Here, `_` takes on the evaluated value of `optimizer`, which we don't need (and `c` takes the value of the `cost` variable).
### 2.6 - Building the model
Now, you will bring it all together!
**Exercise:** Implement the model. You will be calling the functions you had previously implemented.
```
def model(X_train, Y_train, X_test, Y_test, learning_rate = 0.0001,
num_epochs = 1500, minibatch_size = 32, print_cost = True):
"""
Implements a three-layer tensorflow neural network: LINEAR->RELU->LINEAR->RELU->LINEAR->SOFTMAX.
Arguments:
X_train -- training set, of shape (input size = 12288, number of training examples = 1080)
Y_train -- test set, of shape (output size = 6, number of training examples = 1080)
X_test -- training set, of shape (input size = 12288, number of training examples = 120)
Y_test -- test set, of shape (output size = 6, number of test examples = 120)
learning_rate -- learning rate of the optimization
num_epochs -- number of epochs of the optimization loop
minibatch_size -- size of a minibatch
print_cost -- True to print the cost every 100 epochs
Returns:
parameters -- parameters learnt by the model. They can then be used to predict.
"""
ops.reset_default_graph() # to be able to rerun the model without overwriting tf variables
tf.set_random_seed(1) # to keep consistent results
seed = 3 # to keep consistent results
(n_x, m) = X_train.shape # (n_x: input size, m : number of examples in the train set)
n_y = Y_train.shape[0] # n_y : output size
costs = [] # To keep track of the cost
# Create Placeholders of shape (n_x, n_y)
### START CODE HERE ### (1 line)
X, Y = create_placeholders(n_x, n_y)
### END CODE HERE ###
# Initialize parameters
### START CODE HERE ### (1 line)
parameters = initialize_parameters()
### END CODE HERE ###
# Forward propagation: Build the forward propagation in the tensorflow graph
### START CODE HERE ### (1 line)
Z3 = forward_propagation(X, parameters)
### END CODE HERE ###
# Cost function: Add cost function to tensorflow graph
### START CODE HERE ### (1 line)
cost = compute_cost(Z3, Y)
### END CODE HERE ###
# Backpropagation: Define the tensorflow optimizer. Use an AdamOptimizer.
### START CODE HERE ### (1 line)
optimizer = tf.train.AdamOptimizer(learning_rate = learning_rate).minimize(cost)
### END CODE HERE ###
# Initialize all the variables
init = tf.global_variables_initializer()
# Start the session to compute the tensorflow graph
with tf.Session() as sess:
# Run the initialization
sess.run(init)
# Do the training loop
for epoch in range(num_epochs):
epoch_cost = 0. # Defines a cost related to an epoch
num_minibatches = int(m / minibatch_size) # number of minibatches of size minibatch_size in the train set
seed = seed + 1
minibatches = random_mini_batches(X_train, Y_train, minibatch_size, seed)
for minibatch in minibatches:
# Select a minibatch
(minibatch_X, minibatch_Y) = minibatch
# IMPORTANT: The line that runs the graph on a minibatch.
# Run the session to execute the "optimizer" and the "cost", the feedict should contain a minibatch for (X,Y).
### START CODE HERE ### (1 line)
_ , minibatch_cost = sess.run([optimizer, cost], feed_dict={X: minibatch_X, Y: minibatch_Y})
### END CODE HERE ###
epoch_cost += minibatch_cost / num_minibatches
# Print the cost every epoch
if print_cost == True and epoch % 100 == 0:
print ("Cost after epoch %i: %f" % (epoch, epoch_cost))
if print_cost == True and epoch % 5 == 0:
costs.append(epoch_cost)
# plot the cost
plt.plot(np.squeeze(costs))
plt.ylabel('cost')
plt.xlabel('iterations (per tens)')
plt.title("Learning rate =" + str(learning_rate))
plt.show()
# lets save the parameters in a variable
parameters = sess.run(parameters)
print ("Parameters have been trained!")
# Calculate the correct predictions
correct_prediction = tf.equal(tf.argmax(Z3), tf.argmax(Y))
# Calculate accuracy on the test set
accuracy = tf.reduce_mean(tf.cast(correct_prediction, "float"))
print ("Train Accuracy:", accuracy.eval({X: X_train, Y: Y_train}))
print ("Test Accuracy:", accuracy.eval({X: X_test, Y: Y_test}))
return parameters
```
Run the following cell to train your model! On our machine it takes about 5 minutes. Your "Cost after epoch 100" should be 1.016458. If it's not, don't waste time; interrupt the training by clicking on the square (⬛) in the upper bar of the notebook, and try to correct your code. If it is the correct cost, take a break and come back in 5 minutes!
```
parameters = model(X_train, Y_train, X_test, Y_test)
```
**Expected Output**:
<table>
<tr>
<td>
**Train Accuracy**
</td>
<td>
0.999074
</td>
</tr>
<tr>
<td>
**Test Accuracy**
</td>
<td>
0.716667
</td>
</tr>
</table>
Amazing, your algorithm can recognize a sign representing a figure between 0 and 5 with 71.7% accuracy.
**Insights**:
- Your model seems big enough to fit the training set well. However, given the difference between train and test accuracy, you could try to add L2 or dropout regularization to reduce overfitting.
- Think about the session as a block of code to train the model. Each time you run the session on a minibatch, it trains the parameters. In total you have run the session a large number of times (1500 epochs) until you obtained well trained parameters.
### 2.7 - Test with your own image (optional / ungraded exercise)
Congratulations on finishing this assignment. You can now take a picture of your hand and see the output of your model. To do that:
1. Click on "File" in the upper bar of this notebook, then click "Open" to go on your Coursera Hub.
2. Add your image to this Jupyter Notebook's directory, in the "images" folder
3. Write your image's name in the following code
4. Run the code and check if the algorithm is right!
```
import scipy
from PIL import Image
from scipy import ndimage
## START CODE HERE ## (PUT YOUR IMAGE NAME)
my_image = "thumbs_up.jpg"
## END CODE HERE ##
# We preprocess your image to fit your algorithm.
fname = "images/" + my_image
image = np.array(ndimage.imread(fname, flatten=False))
my_image = scipy.misc.imresize(image, size=(64,64)).reshape((1, 64*64*3)).T
my_image_prediction = predict(my_image, parameters)
plt.imshow(image)
print("Your algorithm predicts: y = " + str(np.squeeze(my_image_prediction)))
```
You indeed deserved a "thumbs-up" although as you can see the algorithm seems to classify it incorrectly. The reason is that the training set doesn't contain any "thumbs-up", so the model doesn't know how to deal with it! We call that a "mismatched data distribution" and it is one of the various of the next course on "Structuring Machine Learning Projects".
<font color='blue'>
**What you should remember**:
- Tensorflow is a programming framework used in deep learning
- The two main object classes in tensorflow are Tensors and Operators.
- When you code in tensorflow you have to take the following steps:
- Create a graph containing Tensors (Variables, Placeholders ...) and Operations (tf.matmul, tf.add, ...)
- Create a session
- Initialize the session
- Run the session to execute the graph
- You can execute the graph multiple times as you've seen in model()
- The backpropagation and optimization is automatically done when running the session on the "optimizer" object.
| github_jupyter |
```
import os, platform, pprint, sys
import fastai
import matplotlib as mpl
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import sklearn
import yellowbrick as yb
from fastai.tabular.data import TabularDataLoaders, TabularPandas
from fastai.tabular.all import FillMissing, Categorify, Normalize, tabular_learner, accuracy, ClassificationInterpretation, ShowGraphCallback, RandomSplitter, range_of
from sklearn.base import BaseEstimator
from sklearn.metrics import accuracy_score, classification_report
from sklearn.neighbors import KNeighborsClassifier
from lightgbm import LGBMClassifier
from yellowbrick.model_selection import CVScores, LearningCurve, ValidationCurve
seed: int = 14
# set up pretty printer for easier data evaluation
pretty = pprint.PrettyPrinter(indent=4, width=30).pprint
# declare file paths for the data we will be working on
data_path_1: str = '../../data/prepared/baseline/'
data_path_2: str = '../../data/prepared/timebased/'
modelPath : str = '../models'
# list the names of the datasets we will be using
attacks : list = [ 'DNS', 'LDAP', 'MSSQL', 'NetBIOS', 'NTP', 'Portmap', 'SNMP', 'SSDP', 'Syn', 'TFTP', 'UDP', 'UDPLag' ]
datasets: list = [
"DNS_vs_benign.csv" , "LDAP_vs_benign.csv" , "MSSQL_vs_benign.csv" , "NetBIOS_vs_benign.csv" ,
"NTP_vs_benign.csv" , "Portmap_vs_benign.csv" , "SNMP_vs_benign.csv" , "SSDP_vs_benign.csv" ,
"Syn_vs_benign.csv" , "TFTP_vs_benign.csv" , "UDP_vs_benign.csv" , "UDPLag_vs_benign.csv" ,
]
# set up enumeration of experiment types
Baseline : int = 0
Timebased: int = 1
# print library and python versions for reproducibility
print(
f'''
python:\t{platform.python_version()}
\tfastai:\t\t{fastai.__version__}
\tmatplotlib:\t{mpl.__version__}
\tnumpy:\t\t{np.__version__}
\tpandas:\t\t{pd.__version__}
\tsklearn:\t{sklearn.__version__}
\tyellowbrick:\t{yb.__version__}
'''
)
def load_data(filePath: str) -> pd.DataFrame:
'''
Loads the Dataset from the given filepath and caches it for quick access in the future
Function will only work when filepath is a .csv file
'''
# slice off the ./CSV/ from the filePath
if filePath[0] == '.' and filePath[1] == '.':
filePathClean: str = filePath[17::]
pickleDump: str = f'../../data/cache/{filePathClean}.pickle'
else:
pickleDump: str = f'../../data/cache/{filePath}.pickle'
print(f'Loading Dataset: {filePath}')
print(f'\tTo Dataset Cache: {pickleDump}')
# check if data already exists within cache
if os.path.exists(pickleDump):
df = pd.read_pickle(pickleDump)
# if not, load data and cache it
else:
df = pd.read_csv(filePath, low_memory=True)
df.to_pickle(pickleDump)
return df
def run_experiment(df: pd.DataFrame, name: str) -> tuple:
'''
Run Classification experiment on the given dataset using LGBM
returns the 7-tuple with the following indicies:
results: tuple = (name, model, classes, X_train, y_train, X_test, y_test)
'''
# First we split the features into the dependent variable and
# continous and categorical features
dep_var: str = 'Label'
if 'Protocol' in df.columns:
categorical_features: list = ['Protocol']
else:
categorical_features: list = []
continuous_features = list(set(df) - set(categorical_features) - set([dep_var]))
# Next, we set up the feature engineering pipeline, namely filling missing values
# encoding categorical features, and normalizing the continuous features
# all within a pipeline to prevent the normalization from leaking details
# about the test sets through the normalized mapping of the training sets
procs = [FillMissing, Categorify, Normalize]
splits = RandomSplitter(valid_pct=0.2, seed=seed)(range_of(df))
# The dataframe is loaded into a fastai datastructure now that
# the feature engineering pipeline has been set up
to = TabularPandas(
df , y_names=dep_var ,
splits=splits , cat_names=categorical_features ,
procs=procs , cont_names=continuous_features ,
)
# We use fastai to quickly extract the names of the classes as they are mapped to the encodings
dls = to.dataloaders(bs=64)
mds = tabular_learner(dls)
classes : list = list(mds.dls.vocab)
# We extract the training and test datasets from the dataframe
X_train = to.train.xs.reset_index(drop=True)
X_test = to.valid.xs.reset_index(drop=True)
y_train = to.train.ys.values.ravel()
y_test = to.valid.ys.values.ravel()
# Now that we have the train and test datasets, we set up a Support Vector Machine
# using SciKitLearn and print the results
model = LGBMClassifier()
# model = SVC(C=1, gamma=0.1, kernel='rbf', random_state=seed)
model.fit(X_train, y_train)
prediction = model.predict(X_test)
print(f'\tAccuracy: {accuracy_score(y_test, prediction)}\n')
report = classification_report(y_test, prediction)
print(report)
# we add a target_type_ attribute to our model so yellowbrick knows how to make the visualizations
if len(classes) == 2:
model.target_type_ = 'binary'
elif len(classes) > 2:
model.target_type_ = 'multiclass'
else:
print('Must be more than one class to perform classification')
raise ValueError('Wrong number of classes')
# Now that the classifier has been created and trained, we pass out our training values
# so that yellowbrick can use them to create various visualizations
results: tuple = (name, model, classes, X_train, y_train, X_test, y_test)
return results
def visualize_learning_curve_train(results: tuple) -> None:
'''
Takes a 7-tuple from the run_experiments function and creates a learning curve
results: tuple = (name, model, classes, X_train, y_train, X_test, y_test)
'''
# Track the learning curve of the classifier, here we want the
# training and validation scores to approach 1
visualizer = LearningCurve(results[1], scoring='f1_weighted')
visualizer.fit(results[3], results[4])
visualizer.show()
def visualize_learning_curve_test(results: tuple) -> None:
'''
Takes a 7-tuple from the run_experiments function and creates a learning curve
results: tuple = (name, model, classes, X_train, y_train, X_test, y_test)
'''
# Track the learning curve of the classifier, here we want the
# training and validation scores to approach 1
visualizer = LearningCurve(results[1], scoring='f1_weighted')
visualizer.fit(results[5], results[6])
visualizer.show()
def visualize_confusion_matrix(results: tuple) -> None:
'''
Takes a 7-tuple from the run_experiments function and creates a confusion matrix
results: tuple = (name, model, classes, X_train, y_train, X_test, y_test)
'''
visualizer = yb.classifier.ConfusionMatrix(results[1], classes=results[2], title=results[0])
visualizer.score(results[5], results[6])
visualizer.show()
def visualize_roc(results: tuple) -> None:
'''
Takes a 7-tuple from the run_experiments function and creates a
Receiver Operating Characteristic (ROC) Curve
results: tuple = (name, model, classes, X_train, y_train, X_test, y_test)
'''
visualizer = yb.classifier.ROCAUC(results[1], classes=results[2], title=results[0])
visualizer.score(results[5], results[6])
visualizer.poof()
def visualize_pr_curve(results: tuple) -> None:
'''
Takes a 7-tuple from the run_experiments function and creates a
Precision-Recall Curve
results: tuple = (name, model, classes, X_train, y_train, X_test, y_test)
'''
visualizer = yb.classifier.PrecisionRecallCurve(results[1], title=results[0])
visualizer.score(results[5], results[6])
visualizer.poof()
def visualize_report(results: tuple) -> None:
'''
Takes a 7-tuple from the run_experiments function and creates a report
detailing the Precision, Recall, f1, and Support scores for all
classification outcomes
results: tuple = (name, model, classes, X_train, y_train, X_test, y_test)
'''
visualizer = yb.classifier.ClassificationReport(results[1], classes=results[2], title=results[0], support=True)
visualizer.score(results[5], results[6])
visualizer.poof()
def visualize_class_balance(results: tuple) -> None:
'''
Takes a 7-tuple from the run_experiments function and creates a histogram
detailing the balance between classification outcomes
results: tuple = (name, model, classes, X_train, y_train, X_test, y_test)
'''
visualizer = yb.target.ClassBalance(labels=results[0])
visualizer.fit(results[4], results[6])
visualizer.show()
def get_file_path(directory: str):
'''
Closure that will return a function that returns the filepath to the directory given to the closure
'''
def func(file: str) -> str:
return os.path.join(directory, file)
return func
# use the get_file_path closure to create a function that will return the path to a file
baseline_path = get_file_path(data_path_1)
timebased_path = get_file_path(data_path_2)
# create a list of the paths to all of the dataset files
baseline_files : list = list(map(baseline_path , datasets))
timebased_files : list = list(map(timebased_path , datasets))
baseline_dfs : map = map(load_data , baseline_files )
timebased_dfs : map = map(load_data , timebased_files)
experiments : zip = zip(baseline_dfs , timebased_dfs , attacks)
def experiment_runner():
'''
A generator that handles running the experiments
'''
num = 1
for baseline, timebased, info in experiments:
print(f'Running experiment #{num}:\t{info}')
print('Baseline results')
baseline_results = run_experiment(baseline, f'{info}_vs_benign_baseline')
print('\nTime-based results')
timebased_results = run_experiment(timebased, f'{info}_vs_benign_timebased')
num += 1
yield (baseline_results, timebased_results, info, num)
def do_experiment(num: int) -> tuple:
'''
A function that runs the specific experiment specified
'''
index = num - 1
baseline = load_data(baseline_files[index])
timebased = load_data(timebased_files[index])
info = attacks[index]
print(f'Running experiment #{num}:\t{info}')
print('Baseline results\n')
baseline_results = run_experiment(baseline, f'{info}_vs_benign_baseline')
print('Time-based results\n')
timebased_results = run_experiment(timebased, f'{info}_vs_benign_timebased')
return (baseline_results, timebased_results, info, num)
experiment = experiment_runner()
results = next(experiment)
visualize_report(results[Baseline])
visualize_report(results[Timebased])
visualize_confusion_matrix(results[Baseline])
visualize_confusion_matrix(results[Timebased])
visualize_pr_curve(results[Baseline])
visualize_pr_curve(results[Timebased])
results = next(experiment)
visualize_report(results[Baseline])
visualize_report(results[Timebased])
visualize_confusion_matrix(results[Baseline])
visualize_confusion_matrix(results[Timebased])
visualize_pr_curve(results[Baseline])
visualize_pr_curve(results[Timebased])
results = next(experiment)
visualize_report(results[Baseline])
visualize_report(results[Timebased])
visualize_confusion_matrix(results[Baseline])
visualize_confusion_matrix(results[Timebased])
visualize_pr_curve(results[Baseline])
visualize_pr_curve(results[Timebased])
results = next(experiment)
visualize_report(results[Baseline])
visualize_report(results[Timebased])
visualize_confusion_matrix(results[Baseline])
visualize_confusion_matrix(results[Timebased])
visualize_pr_curve(results[Baseline])
visualize_pr_curve(results[Timebased])
results = next(experiment)
visualize_report(results[Baseline])
visualize_report(results[Timebased])
visualize_confusion_matrix(results[Baseline])
visualize_confusion_matrix(results[Timebased])
visualize_pr_curve(results[Baseline])
visualize_pr_curve(results[Timebased])
results = next(experiment)
visualize_report(results[Baseline])
visualize_report(results[Timebased])
visualize_confusion_matrix(results[Baseline])
visualize_confusion_matrix(results[Timebased])
visualize_pr_curve(results[Baseline])
visualize_pr_curve(results[Timebased])
results = next(experiment)
visualize_report(results[Baseline])
visualize_report(results[Timebased])
visualize_confusion_matrix(results[Baseline])
visualize_confusion_matrix(results[Timebased])
visualize_pr_curve(results[Baseline])
visualize_pr_curve(results[Timebased])
results = next(experiment)
visualize_report(results[Baseline])
visualize_report(results[Timebased])
visualize_confusion_matrix(results[Baseline])
visualize_confusion_matrix(results[Timebased])
visualize_pr_curve(results[Baseline])
visualize_pr_curve(results[Timebased])
results = next(experiment)
visualize_report(results[Baseline])
visualize_report(results[Timebased])
visualize_confusion_matrix(results[Baseline])
visualize_confusion_matrix(results[Timebased])
visualize_pr_curve(results[Baseline])
visualize_pr_curve(results[Timebased])
results = next(experiment)
visualize_report(results[Baseline])
visualize_report(results[Timebased])
visualize_confusion_matrix(results[Baseline])
visualize_confusion_matrix(results[Timebased])
visualize_pr_curve(results[Baseline])
visualize_pr_curve(results[Timebased])
results = next(experiment)
visualize_report(results[Baseline])
visualize_report(results[Timebased])
visualize_confusion_matrix(results[Baseline])
visualize_confusion_matrix(results[Timebased])
visualize_pr_curve(results[Baseline])
visualize_pr_curve(results[Timebased])
results = next(experiment)
visualize_report(results[Baseline])
visualize_report(results[Timebased])
visualize_confusion_matrix(results[Baseline])
visualize_confusion_matrix(results[Timebased])
visualize_pr_curve(results[Baseline])
visualize_pr_curve(results[Timebased])
```
| github_jupyter |
### Reference
* http://users.cecs.anu.edu.au/~u5098633/papers/www15.pdf
* https://grouplens.org/datasets/movielens/1m/
* https://vitobellini.github.io/posts/2018/01/03/how-to-build-a-recommender-system-in-tensorflow.html
* https://github.com/npow/AutoRec
```
import random
import numpy as np
import pandas as pd
import tensorflow as tf
print('version info:')
print('Numpy\t', np.__version__)
print('Pandas\t', pd.__version__)
print('TF\t', tf.__version__)
BATCH_SIZE = 256
WEIGHT_DECAY = 5e-4
```
### [Option] download MovieLens 1M dataset
```
import os
import wget
import zipfile
dirPath = '../dataset'
zipFilePath = os.path.join(dirPath, 'ml-1m.zip')
remoteRrl = 'https://files.grouplens.org/datasets/movielens/ml-1m.zip'
if not os.path.exists(dirPath):
os.makedirs(dirPath)
# download
wget.download(remoteRrl, zipFilePath)
# unzip files
with zipfile.ZipFile(zipFilePath, 'r') as zipRef:
zipRef.extractall(dirPath)
```
## load dataset
```
df = pd.read_csv('../dataset/ml-1m/ratings.dat', sep='::', engine='python', names=['UserID', 'MovieID', 'Rating', 'Timestamp'], header=None)
df = df.drop('Timestamp', axis=1)
numOfUsers = df.UserID.nunique()
numOfItems = df.MovieID.nunique()
df.head()
# Normalize rating in [0, 1]
ratings = df.Rating.values.astype(np.float)
scaledRatings = (ratings - min(ratings)) / (max(ratings) - min(ratings))
df.Rating = pd.DataFrame(scaledRatings)
df.head()
# user-item rating matrix
## U-AutoRec (users-based)
userItemRatingMatrix = df.pivot(index='UserID', columns='MovieID', values='Rating')
## I-AutoRec (items-based)
# userItemRatingMatrix = df.pivot(index='MovieID', columns='UserID', values='Rating')
userItemRatingMatrix.fillna(-1, inplace=True)
userItemRatingMatrix
# create tf.dataset
def getDataset(userItemRatingMatrix):
userItemRatingMatrix_np = userItemRatingMatrix.to_numpy(dtype=np.float32)
random.shuffle(userItemRatingMatrix_np)
# [train : valid : test] = [0.7 : 0.15 : 0.15]
numOfTrainSet = int(numOfUsers * 0.7)
numOfValidSet = int(numOfUsers * 0.15)
numOfTestSet = numOfUsers - numOfTrainSet - numOfValidSet
trainSet_np = userItemRatingMatrix_np[0:numOfTrainSet]
validSet_np = userItemRatingMatrix_np[numOfTrainSet:numOfTrainSet+numOfValidSet]
testSet_np = userItemRatingMatrix_np[numOfTrainSet+numOfValidSet:]
trainSet = tf.data.Dataset.from_tensor_slices(trainSet_np)
validSet = tf.data.Dataset.from_tensor_slices(validSet_np)
testSet = tf.data.Dataset.from_tensor_slices(testSet_np)
trainSet = trainSet.shuffle(buffer_size=BATCH_SIZE*8).batch(BATCH_SIZE)
validSet = validSet.batch(BATCH_SIZE)
testSet = testSet.batch(BATCH_SIZE)
return trainSet, validSet, testSet
trainSet, validSet, testSet = getDataset(userItemRatingMatrix)
```
## build model
```
# build model
## tf.keras.Dense: https://www.tensorflow.org/api_docs/python/tf/keras/layers/Dense
regularizer = tf.keras.regularizers.L2(WEIGHT_DECAY)
def getEncoder(numOfInput, numOfHidden1, numOfHidden2):
x = tf.keras.Input(shape=(numOfInput,))
out = tf.keras.layers.Dense(units=numOfHidden1, activation='sigmoid', kernel_regularizer=regularizer) (x)
out = tf.keras.layers.Dense(units=numOfHidden2, activation='sigmoid', kernel_regularizer=regularizer) (out)
return tf.keras.Model(inputs=[x], outputs=[out])
def getDecoder(numOfInput, numOfHidden1, numOfHidden2):
x = tf.keras.Input(shape=(numOfHidden2,))
out = tf.keras.layers.Dense(units=numOfHidden1, activation='sigmoid', kernel_regularizer=regularizer) (x)
out = tf.keras.layers.Dense(units=numOfInput, activation='sigmoid', kernel_regularizer=regularizer) (out)
return tf.keras.Model(inputs=[x], outputs=[out])
def getAutoEncoder(numOfInput, numOfHidden1, numOfHidden2):
encoder = getEncoder(numOfInput, numOfHidden1, numOfHidden2)
decoder = getDecoder(numOfInput, numOfHidden1, numOfHidden2)
return encoder, decoder
encoder, decoder = getAutoEncoder(numOfInput=userItemRatingMatrix.shape[-1], numOfHidden1=10, numOfHidden2=5)
```
## training
```
# optimizer
optimizer = tf.keras.optimizers.SGD(learning_rate=5e-1, momentum=0.9)
# loss function
def getLoss(pred, gt, mask):
reconstructionLoss = tf.reduce_sum(tf.pow(pred - gt, 2) * mask, axis=-1) / tf.reduce_sum(mask, -1)
reconstructionLoss = tf.reduce_mean(reconstructionLoss)
return reconstructionLoss
# training with tf.GradientTape
from collections import defaultdict
weights = encoder.trainable_weights + decoder.trainable_weights
records = defaultdict(list)
numOfEpochs = 100
for epoch in range(numOfEpochs):
trainLosses = []
for step, batch in enumerate(trainSet):
mask = tf.cast(batch != -1, dtype=tf.float32)
# replace unrated value with mean of ratings
mean = tf.reduce_sum(batch * mask, axis=-1, keepdims=True) / tf.reduce_sum(mask, axis=-1, keepdims=True)
x = mask * batch + (1 - mask) * mean
with tf.GradientTape() as tape:
embedding = encoder(x, training=True)
pred = decoder(embedding, training=True)
loss = getLoss(pred=pred, gt=x, mask=mask)
grads = tape.gradient(loss, weights)
optimizer.apply_gradients(zip(grads, weights))
trainLosses.append(loss)
if epoch%5 == 0:
# calculate reconstruction loss from validation dataset
validLosses = []
for batch in validSet:
mask = tf.cast(batch != -1, dtype=tf.float32)
# replace unrated value with mean of ratings
mean = tf.reduce_sum(batch * mask, axis=-1, keepdims=True) / tf.reduce_sum(mask, axis=-1, keepdims=True)
x = mask * batch + (1 - mask) * mean
embedding = encoder(x, training=False)
pred = decoder(embedding, training=False)
validLoss = getLoss(pred=pred, gt=x, mask=mask)
validLosses.append(validLoss)
records['train'].append(tf.reduce_mean(trainLosses).numpy())
records['valid'].append(tf.reduce_mean(validLosses).numpy())
print(f'epoch:{epoch}, trainLoss:{records['train'][-1]}, validLoss:{records['valid'][-1]}')
import matplotlib.pyplot as plt
plt.plot(records['train'], label='train')
plt.plot(records['valid'], label='valid')
plt.legend()
plt.ylabel('Loss(RMSE)')
plt.xlabel('Epochs')
```
## testing
calculate the RMSE of ratings
```
testLosses = []
for batch in testSet:
mask = tf.cast(batch != -1, dtype=tf.float32)
# replace unrated value with mean of ratings
mean = tf.reduce_sum(batch * mask, axis=-1, keepdims=True) / tf.reduce_sum(mask, axis=-1, keepdims=True)
x = mask * batch + (1 - mask) * mean
embedding = encoder.predict(x)
pred = decoder.predict(embedding)
testLoss = getLoss(pred=pred, gt=x, mask=mask)
testLosses.append(testLoss)
print(f'RMSE: {tf.reduce_mean(testLosses)}')
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
synthetic = pd.read_csv('../results/results_henao/synthetic_1580965994.csv')
synthetic = synthetic.rename(columns=lambda x: x.strip())
mimic = pd.read_csv('../results/results_henao/mimic_1580877802.csv')
mimic = mimic.iloc[np.array(range(0, 20, 2)), :]
mimic = mimic.rename(columns=lambda x: x.strip())
reddit = pd.read_csv('../results/results_henao/reddit_1580933275.csv')
reddit = reddit.iloc[np.array(range(0, 20, 2)), :]
reddit = reddit.rename(columns=lambda x: x.strip())
synthetic_baselines = pd.read_csv('../results/results_henao/synthetic_baselines_1580966689.csv')
synthetic_baselines = synthetic_baselines.rename(columns=lambda x: x.strip())
reddit.columns
mimic_baselines = pd.read_csv('../results/results_rapidshare/mimic_baselines_1580932980.csv')[:30]
mimic_baselines = mimic_baselines.rename(columns=lambda x: x.strip())
reddit_baselines = pd.read_csv('../results/results_rapidshare/reddit_baselines_1580932800.csv')[:30]
reddit_baselines = reddit_baselines.rename(columns=lambda x: x.strip())
def stack_metric(df, metric, num, model, dataset):
dfs = df[['%s%i' % (metric, i) for i in range(num)]].stack().reset_index()[['level_1', 0]]
dfs.columns = ['task', metric]
dfs[metric] = dfs[metric].astype(float)
dfs['model'] = model
dfs['dataset'] = dataset
return dfs
import seaborn as sns
fig, ax = plt.subplots(1, 2, figsize=(8, 4))
sns.set_style("whitegrid")
data = pd.concat([stack_metric(synthetic, 'auc', 2, 'CET', 'Synthetic'),
stack_metric(synthetic_baselines[synthetic_baselines['model_type'] == ' survival'], 'auc', 2, 'ET', 'Synthetic'),
stack_metric(synthetic_baselines[synthetic_baselines['model_type'] == ' s_mlp'], 'auc', 2, 'BC', 'Synthetic'),
stack_metric(mimic, 'auc', 10, 'CET', 'MIMIC'),
stack_metric(mimic_baselines[mimic_baselines['model_type'] == ' survival'], 'auc', 10, 'ET', 'MIMIC'),
stack_metric(mimic_baselines[mimic_baselines['model_type'] == ' s_mlp'], 'auc', 10, 'BC', 'MIMIC'),
stack_metric(reddit, 'auc', 9, 'CET', 'reddit'),
stack_metric(reddit_baselines[reddit_baselines['model_type'] == ' survival'], 'auc', 9, 'ET', 'reddit'),
stack_metric(reddit_baselines[reddit_baselines['model_type'] == ' s_mlp'], 'auc', 9, 'BC', 'reddit')],
axis=0)
sns.boxplot(x='dataset', y='auc', hue='model', data=data, ax=ax[0], linewidth=0.5, width=.6)
ax[0].set_ylabel('AUC', fontsize=12)
ax[0].set_xlabel('')
ax[0].set_ylim([.5, 1.])
data = pd.concat([stack_metric(synthetic, 'raem', 2, 'CET', 'Synthetic'),
stack_metric(synthetic_baselines[synthetic_baselines['model_type'] == ' survival'], 'raem', 2, 'ET', 'Synthetic'),
stack_metric(mimic, 'raem', 10, 'CET', 'MIMIC'),
stack_metric(mimic_baselines[mimic_baselines['model_type'] == ' survival'], 'raem', 10, 'ET', 'MIMIC'),
stack_metric(reddit, 'raem', 9, 'CET', 'reddit'),
stack_metric(reddit_baselines[reddit_baselines['model_type'] == ' survival'], 'raem', 9, 'ET', 'reddit')],
axis=0)
sns.boxplot(x='dataset', y='raem', hue='model', data=data, ax=ax[1], linewidth=0.5, width=.4)
ax[1].set_ylim([1e-3, 1e7])
ax[1].set_yscale('log')
ax[1].set_ylabel('MRAE', fontsize=12)
ax[1].set_xlabel('')
ax[1].set_yticks([1e-2, 1e0, 1e2, 1e4, 1e6])
plt.tight_layout()
plt.savefig('/Users/mme/Downloads/fig3.pdf')
plt.show()
data = pd.concat([stack_metric(synthetic, 'raem', 2, 'CET', 'Synth'),
stack_metric(synthetic_baselines[synthetic_baselines['model_type'] == ' survival'], 'raem', 2, 'ET', 'Synth'),
stack_metric(mimic, 'raem', 10, 'CET', 'MIMIC'),
stack_metric(mimic_baselines[mimic_baselines['model_type'] == ' survival'], 'raem', 10, 'ET', 'MIMIC'),
stack_metric(reddit, 'raem', 9, 'CET', 'reddit'),
stack_metric(reddit_baselines[reddit_baselines['model_type'] == ' survival'], 'raem', 9, 'ET', 'reddit')],
axis=0)
data.values
stack_metric(mimic_baselines[mimic_baselines['model_type'] == 's_mlp'], 'auc', 10, 'BC', 'M')
mimic_baselines['model_type'].value_counts().index[1]
```
| github_jupyter |
# Problem Statement
Given a dataset of the daily car count in a parking lot, a corresponding weather index and a indicator of sky conditions (clear / cloudy) along with the corresponding day of the week, we need to provide a daily forecast for the car count for the next month.
## Approach
Given any problem, the first step is visualization. Then we start with a simple linear regression model, and examine correlations between different features. We will then move to a more probabilistic approach.
Let's load the packages and the dataset first, and mark the input (independent) and the output (dependent) variables.
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn import linear_model
from sklearn.model_selection import cross_val_predict
Data = pd.read_csv('data.csv')
inputVariables=list(Data)
#print inputVariables
del inputVariables[2]
outputVariables=list(Data)[2]
inputData=Data[inputVariables]
#print inputData
```
We see that there are we have a few quantitative (numerical) variables (car.count and weather) and a few qualitative (categorical) variables (Date, day.of.week and cloud.indicator).
Before we start to develop any model for the problem at hand, or do any data processing, we should first visualize the available numerical data.
```
plt.figure(figsize=(16,4))
plt.subplot(1,2,1)
plt.plot(Data[outputVariables])
plt.ylabel('Car Count')
plt.xlabel('Time (Day)')
plt.subplot(1,2,2)
plt.plot(inputData['weather'])
plt.ylabel('Weather')
plt.xlabel('Time (Day)')
```
These two plots tell us that the car count data does not appear to be stationary, while the weather data does. To see this in a more quantitative fashion, we analyze the data using a multifractal method (see details in Agarwal et. al., PRSA, 2012). Below I plot the second moment of the fluctuation functions for the Car Count and the Weather data.

The slopes of the blue and purple curves denote the statistical dynamics for the car count and weather data, respectively. The dashed lines orange and yellow denote the white noise and pink noise, respectively. The slope of these fluctuation functions give us the dynamics, while the points where the slope changes are the dominant timescales in the data.
#### Car Count
It has a multifractal structure, with Gaussian white noise dynamics on timescales from a day to upto 200 days, and approximate pink noise dynamics on longer timescales. This tells us that given our problem for prediction of the next 30 days, a white noise structure might work.
#### Weather
It has a fractal structure, with white noise on all timescales.
Let us now visualze the correlation between the weather and the car count data.
```
plt.figure()
plt.plot(inputData['weather'],Data[outputVariables],'o')
plt.xlabel('Weather')
plt.ylabel('Car Count')
```
Fitting a straight line (Linear Regression for Car Count vs Weather) through this gives us the following model:
Using normal equation formulation:
y = Car Count;
X = [1 Weather]; (1 is for the bias in the model)
w = inv(X' X) * (X'y);
```
Y = Data[outputVariables]
Y = np.reshape(Y,(len(Y),1))
X = np.ones((len(inputData['weather']), 1))
C = inputData['weather']
C = np.reshape(C,(len(C),1))
X = np.concatenate((X, C), 1)
#w = (np.linalg.inv((X.T).dot(X))).dot(X.T).dot(Y) ## Normal Equation Form
model_1 = linear_model.LinearRegression(fit_intercept=True, normalize=False, copy_X=True, n_jobs=1)
model_1.fit(C,Y)
c = model_1.intercept_[0]
w = model_1.coef_[0]
predicted = cross_val_predict(model_1, C, Y, cv=10)
print('Mean squared error:',np.mean((predicted - Y) ** 2))
print('R-squared:',model_1.score(C, Y))
plt.figure()
plt.plot(inputData['weather'],Data[outputVariables],'o')
x = np.linspace(min(C), max(C), 1000)
y = c + w * x
plt.plot(x, y, linewidth = 4)
plt.xlabel('Weather')
plt.ylabel('Car Count')
equation = 'y = ' + str(w) + 'x' ' + ' + str(c)
plt.text(10, 240,equation, horizontalalignment='center', verticalalignment='center')
```
This is a simple model and we see that the fit is not really great.
We need to use the information from the categorical variables and see how they affect the car count.
Given all the variables, we can possibly have the following dependencies:
1) Car Count depends on day of week
2) Car Count depends on the month of year (e.g. holiday season)
3) Car Count depends on sky conditions (Cloudy vs Clear)
4) Sky condition itself has a monthly dependence (as would be expected from the Earth climate sciences)
Before we check for the above dependencies, we need to modify the categorical variables to a numeric form. We parse the date first to compute the month of the year and replace the 'date' with the 'month'.
```
Date = inputData['date']
Month = Date.copy()
from dateutil.parser import parse
current = 0
while current < len(Date):
temp = parse(Date[current])
Month[current] = temp.month
current += 1
Month.name = 'month'
inputData=inputData.join(Month)
del inputData['date']
```
We then change the categorical variables to dummy variables.
```
for column in inputData.columns:
if inputData[column].dtype==object:
dummyCols=pd.get_dummies(inputData[column])
inputData=inputData.join(dummyCols)
del inputData[column]
```
We first check the car count dependency on the day of the week, and plot the car count distributions for each day of the week. The variation in the distributions will tell us if the car count is correlated with the day of the week.
```
ID = np.asarray(inputData)
plt.figure(figsize=(12,12))
for i in range(1,8):
A = ID[:,i]
B = Y[A==1]
plt.subplot(2,4,i)
plt.hist(B)
plt.axis([0, 240, 0, 80])
plt.title((str(i)))
```
These histograms show that the distribution for all days of the week (1 = Friday, 2 = Monday, 3 = Saturday, 4 = Sunday, 5 = Thursday, 6 = Tuesday, 7 = Wednesday) are almost similar, with only very slight variations. Therefore, we ignore this dependency.
We now check the monthly dependencies, and plot the car count distributions for each month. As with the day of the week, the variation in these distributions will tell us if the car count is correlated with the month.
```
plt.figure(figsize=(12,12))
for i in range(1,13):
A = inputData[i]
B = Y[A==1]
plt.subplot(3,4,i)
plt.hist(B)
plt.axis([0, 240, 0, 55])
plt.title(str(i))
```
These histograms show that the distribution for all the months (1 = January...12 = December) are also similar, with small inter-month variations. We will discuss these variations later.
We now look at the last categorical variable in the data, the sky conditions (clear / cloudy).
```
Clear = inputData['clear']
Weather = inputData['weather']
Weather_Cloudy = Weather[Clear==0]
Weather_Clear = Weather[Clear==1]
Car = Data[outputVariables]
Car_Cloudy = Car[Clear==0]
Car_Clear = Car[Clear==1]
plt.figure()
plt.subplot(1,2,1)
plt.hist(Car_Cloudy)
plt.axis([0, 300, 0, 320])
plt.title('cloudy')
plt.subplot(1,2,2)
plt.hist(Car_Clear)
plt.axis([0, 300, 0, 320])
plt.title('clear')
plt.figure()
plt.plot(Weather_Cloudy,Car_Cloudy,'o',label='cloudy')
plt.plot(Weather_Clear,Car_Clear,'o',label='clear')
plt.xlabel('weather')
plt.legend()
```
Here we see a clear distinction between the 'cloudy' and the 'clear' sky conditions. The car count in the parking lot is high when the sky is clear and relatively low when there are cloudy conditions (People like to go out when it is clear outside!!!). Therefore, we start with developing models for the car count with respect to sky conditions.
```
Weather_Cloudy = np.reshape(Weather_Cloudy,(len(Weather_Cloudy),1))
Car_Cloudy = np.reshape(Car_Cloudy,(len(Car_Cloudy),1))
model_1 = linear_model.LinearRegression(fit_intercept=True, normalize=False, copy_X=True, n_jobs=1)
model_1.fit(Weather_Cloudy,Car_Cloudy)
c1 = model_1.intercept_[0]
w1 = model_1.coef_[0]
predicted_1 = cross_val_predict(model_1, Weather_Cloudy, Car_Cloudy, cv=10)
print('Mean squared error (Cloudy):',np.mean((predicted_1 - Car_Cloudy) ** 2))
print('R-squared (Cloudy):',model_1.score(Weather_Cloudy,Car_Cloudy))
Weather_Clear = np.reshape(Weather_Clear,(len(Weather_Clear),1))
Car_Clear = np.reshape(Car_Clear,(len(Car_Clear),1))
model_1 = linear_model.LinearRegression(fit_intercept=True, normalize=False, copy_X=True, n_jobs=1)
model_1.fit(Weather_Clear,Car_Clear)
c2 = model_1.intercept_[0]
w2 = model_1.coef_[0]
predicted_2 = cross_val_predict(model_1, Weather_Clear, Car_Clear, cv=10)
print('Mean squared error (Clear):',np.mean((predicted_2 - Car_Clear) ** 2))
print('R-squared (Clear):',model_1.score(Weather_Clear, Car_Clear))
print('Mean squared error (Total):',(np.add(np.sum((predicted_2 - Car_Clear) ** 2), np.sum((predicted_1 - Car_Cloudy) ** 2)))/(len(predicted_1)+len(predicted_2)))
plt.figure()
plt.plot(Weather_Cloudy,Car_Cloudy,'o',label='cloudy')
plt.plot(Weather_Clear,Car_Clear,'o',label='clear')
plt.xlabel('weather')
plt.legend()
x = np.linspace(min(Weather_Cloudy), max(Weather_Cloudy), 1000)
y = c1 + w1 * x
plt.plot(x, y, linewidth = 4)
x = np.linspace(min(Weather_Clear), max(Weather_Clear), 1000)
y = c2 + w2 * x
plt.plot(x, y, linewidth = 4)
plt.xlabel('Weather')
plt.ylabel('Car Count')
equation = 'For Cloudy Skies: y = ' + str(w1) + 'x' ' + ' + str(c1)
plt.text(10, 0,equation, horizontalalignment='center',
verticalalignment='center')
equation = 'For Clear Skies: y = ' + str(w2) + 'x' ' + ' + str(c2)
plt.text(10, 240,equation, horizontalalignment='center',
verticalalignment='center')
```
The plot above clearly shows that the car count is high for clear skies compared to the cloudy conditions, with the model having a 'positive' slope and a high intercept for the clear sky conditions and 'negative' slope and a relatively low intercept for the cloudy sky conditions. The combined Mean Squared Error of the two models is also considerable lower than before where we had separated sky conditions.
We would now like to investigate if the weather index and the sky conditions are somehow related, for which we look at the cross-correlation coefficients between the respective data.
```
plt.figure(figsize=(10,4))
plt.subplot(1,2,1)
plt.xcorr(ID[:,0],ID[:,8],maxlags=None)
plt.title('Weather vs. Clear')
plt.ylabel('Correlation Coefficient')
plt.xlabel('Lag')
plt.subplot(1,2,2)
plt.xcorr(ID[:,0],ID[:,9],maxlags=None)
plt.title('Weather vs. Cloudy')
plt.xlabel('Lag')
```
The two plots above show that the weather index and the sky conditions have really low correlations (if at all). Just to reconfirm our hypothesis that the car count in the parking lot is correlated with the sky conditions, we also plot the cross-correlation coefficients between the car count and clear sky conditions, and see that it supports our argument.
```
plt.xcorr(Data[outputVariables],ID[:,8],maxlags=None)
plt.title('Car Count vs. Clear')
plt.ylabel('Correlation Coefficient')
plt.xlabel('Lag')
```
### Naive Bayes
Given that the maximum correlation exists between the car count and tke sky conditions, we will compute the fraction of 'clear' days in a given month, using the Bayes Rule, as follows:
P(clear|Month) = P(Month|clear) * P(clear) / P(Month)
```
Cp = np.zeros((12,1))
Clr = ID[:,8]
for i in range(1,13):
A = ID[:,i+9]
B = Clr[A==1]
Cp[i-1] = float(len(np.where(B==1)[0]))/len(B) ## Computing P(clear|Month)
x = np.arange(1,13)
x = np.reshape((x - np.mean(x))/np.std(x),(len(x),1)) # Scaling the independent variable (Month Index)
x = np.concatenate((x,x**2,x**3,x**4,x**5),1)
model_1=linear_model.LinearRegression(fit_intercept=True, normalize=False, copy_X=True, n_jobs=1) # Ordinary Linear Regression
model_2=linear_model.Ridge(alpha=0.1,fit_intercept=True, normalize=False, copy_X=True) # Using L2 Regularization
model_1.fit(x,Cp)
model_2.fit(x,Cp)
print model_1.coef_
print model_2.coef_
p1 = model_1.coef_[0]
c1 = model_1.intercept_
p2 = model_2.coef_[0]
c2 = model_2.intercept_
x = np.arange(1,12.1,0.1)
x2 = (x - np.mean(range(1,13)))/np.std(range(1,13))
y1 = x2**5 * p1[4] + x2**4 * p1[3] + x2**3 * p1[2] + x2**2 * p1[1] + x2 * p1[0] + c1
y2 = x2**5 * p2[4] + x2**4 * p2[3] + x2**3 * p2[2] + x2**2 * p2[1] + x2 * p2[0] + c2
plt.plot(range(1,13),Cp,'-o')
plt.plot(x,y1)
plt.plot(x,y2)
plt.xlabel('Month')
plt.ylabel('Clear Days Fraction')
```
Above, we fitted the monthly variation in the clear day fraction using Ordinary Least Squares, without and with L2 regularization by a 5th order polynomial to look at the inter-month variations.
We can use the computed P(clear|Month) values as a scaling between the clear and cloudy sky conditions, for each month as described below.
```
model_1=linear_model.LinearRegression(fit_intercept=True, normalize=False, copy_X=True, n_jobs=1)
P = np.zeros((5,1))
plt.figure(figsize=(18,13))
Clr = inputData['clear']
Cnt = Data[outputVariables]
Wtr = inputData['weather']
for i in range(1,13):
A = inputData[i]
Ct = Cnt[A==1]
Wt = Wtr[A==1]
Cl = Clr[A==1]
B = np.reshape(Wt[Cl==0],(len(Wt[Cl==0]),1))
C = np.reshape(Ct[Cl==0],(len(Ct[Cl==0]),1))
model_1.fit(B,C)
x1 = np.linspace(min(B), max(B), 1000)
y1 = model_1.intercept_ + model_1.coef_[0] * x1
P[0] = model_1.coef_[0]
P[1] = model_1.intercept_
D = np.reshape(Wt[Cl==1],(len(Wt[Cl==1]),1))
E = np.reshape(Ct[Cl==1],(len(Ct[Cl==1]),1))
model_1.fit(D,E)
x2 = np.linspace(min(D), max(D), 1000)
y2 = model_1.intercept_ + model_1.coef_[0] * x2
P[2] = model_1.coef_[0]
P[3] = model_1.intercept_
model_1.fit(np.reshape(Wt,(len(Wt),1)),np.reshape(Ct,(len(Ct),1)))
x3 = np.linspace(min(Wt), max(Wt), 1000)
y3 = model_1.intercept_ + model_1.coef_[0] * x3
plt.subplot(3,4,i)
plt.plot(B,C,'o',label='cloudy')
plt.plot(D,E,'o',label='clear')
plt.plot(x1, y1, linewidth = 4,label='cloudy')
plt.plot(x2, y2, linewidth = 4,label='clear')
plt.ylabel('Car Count')
plt.xlabel('weather')
plt.title(str(i))
x2 = np.linspace(min(D), max(D), 1000)
P[4] = Cp[i-1] # Using the clear day fraction for a given month to scale the two separate models
y = (P[4]*P[3] + (1-P[4])*P[1]) + (P[4]*P[2] + (1-P[4])*P[0]) * x2
plt.plot(x2, y, linewidth = 4,label='Weigthed Sky Cond.')
plt.plot(x3, y3, linewidth = 4,label='Unweigthed Sky Cond.')
plt.legend()
```
The plots above for each month, while displaying a lot of variation, do not necessarily capture the complete dynamics / variance of the given dataset.
## Probabilistic Models
We know there exists a high correlation between the sky conditions and the car count in the parking lot. To start with this, we can bulid Gaussian probabilistic models, based on P(clear|Month) as computed above and the distribution of car counts for a given month and given sky condition.
#### Model 1
For a given month we use the P(clear|Month) computed above, and then based on that, sample the car counts. As a first step in this model, we need to check if the distribution of car counts for a given month and a given sky condition has Gaussian structure. Lets try this below:
```
Clr = inputData['clear']
Cnt = Data[outputVariables]
Wtr = inputData['weather']
plt.figure(figsize=(15,20))
for i in range(1,13):
A = inputData[i]
Ct = Cnt[A==1]
Wt = Wtr[A==1]
Cl = Clr[A==1]
C = np.reshape(Ct[Cl==0],(len(Ct[Cl==0]),1))
E = np.reshape(Ct[Cl==1],(len(Ct[Cl==1]),1))
plt.subplot(6,4,2*i-1)
plt.hist(C)
plt.axis([0, 300, 0, 30])
plt.title(str(i) + ' - Cloudy')
plt.subplot(6,4,2*i)
plt.hist(E)
plt.axis([0, 300, 0, 30])
plt.title(str(i) + ' - Clear')
```
While the histograms for the car count on clear days has an approximate Gaussian structure (bell shaped), that for cloudy days does not. Therefore, we append the cloudy day data with its negative self, which should then give us an approximate bell curve, as shown below.
```
Clr = inputData['clear']
Cnt = Data[outputVariables]
Wtr = inputData['weather']
plt.figure(figsize=(15,20))
for i in range(1,13):
A = inputData[i]
Ct = Cnt[A==1]
Wt = Wtr[A==1]
Cl = Clr[A==1]
C = np.reshape(Ct[Cl==0],(len(Ct[Cl==0]),1))
C = np.concatenate((C,np.multiply(C,-1)))
E = np.reshape(Ct[Cl==1],(len(Ct[Cl==1]),1))
plt.subplot(6,4,2*i-1)
plt.hist(C)
plt.axis([-300, 300, 0, 40])
plt.title(str(i) + ' - Cloudy')
plt.subplot(6,4,2*i)
plt.hist(E)
plt.axis([0, 300, 0, 40])
plt.title(str(i) + ' - Clear')
```
Now that the Car Count distributions are approximate Gaussians for each month and sky condition, we can move to the next step and build our first model as follows:
1) For each month the number of clear days is P(clear|Month)* (Number of Days in that Month)
2) For each clear / cloudy day in that month, sample from the corresponding car count distribution.
```
meanCnt = np.zeros((12,2))
stdCnt = np.zeros((12,2))
Clr = inputData['clear']
Cnt = Data[outputVariables]
Wtr = inputData['weather']
MonthDay = [31,28,31,30,31,30,31,31,30,31,30,31] # Actual Days in a month
MonthDay = [200,200,200,200,200,200,200,200,200,200,200,200] # To produce distribution
plt.figure(figsize=(12,20))
np.random.seed(1)
for i in range(1,13):
A = inputData[i]
Ct = Cnt[A==1]
Wt = Wtr[A==1]
Cl = Clr[A==1]
C = np.reshape(Ct[Cl==0],(len(Ct[Cl==0]),1))
C = np.concatenate((C,np.multiply(C,-1)))
E = np.reshape(Ct[Cl==1],(len(Ct[Cl==1]),1))
meanCnt[i-1,0] = np.nanmean(C)
stdCnt[i-1,0] = np.nanstd(C)
meanCnt[i-1,1] = np.nanmean(E)
stdCnt[i-1,1] = np.nanstd(E)
Cnts = np.zeros((MonthDay[i-1],1))
clearDays = np.floor(Cp[i-1] * MonthDay[i-1])
Cnts[0:int(clearDays)] = np.floor(abs((np.random.normal(meanCnt[i-1,1],stdCnt[i-1,1],(int(clearDays),1)))))
Cnts[int(clearDays):MonthDay[i-1]] = np.floor(abs(np.random.normal(meanCnt[i-1,0],stdCnt[i-1,0],(int(MonthDay[i-1]-clearDays),1))))
plt.subplot(6,4,2*i-1)
plt.hist(Cnts)
plt.title(str(i) + '-Simulated')
plt.axis([0, 250, 0, 50])
plt.subplot(6,4,2*i)
plt.hist(Ct)
plt.title(str(i) + '-Actual')
plt.axis([0, 250, 0, 50])
```
#### Model 2
1) For each month, instead of using a fixed P(clear|Month), sample the P(clear|Month) from the given distribution of P(clear|Month) for each month of the record and then probabilistically compute the number of clear days in that month.
2) For each clear / cloudy day in that month, sample from the corresponding car count distribution.
We first compute the distribution of P(clear|Month) below:
```
D = np.zeros((7,12))
for i in range(10,22):
A = ID[:,i]
J = np.where((np.ediff1d(A))>0)[0]
D[0:len(J),i-10] = J
D[1:7,0] = D[0:6,0]
D[0,0] = -1
M = np.zeros((7,12))
for i in range(0,7):
for j in range(0,11):
if D[i,j+1] != 0:
J = ID[range(int(D[i,j])+1,int(D[i,j+1])+1),8];
M[i,j] = float(len(np.where(J==1)[0]))/len(J);
for i in range(0,6):
J = ID[range(int(D[i,11])+1,int(D[i+1,0])+1),8];
M[i,11] = float(len(np.where(J==1)[0]))/len(J);
J = ID[range(2343,2373),8]
M[6,5] = float(len(np.where(J==1)[0]))/len(J);
M1 = np.reshape(range(1,13),(12,1))
M1 = (np.concatenate((M1,M1,M1,M1,M1,M1,M1),1)).T
M[6,6:12] = float('nan')
plt.figure()
plt.plot(M1,M,'o')
plt.ylabel('Clear Day Fraction')
plt.xlabel('Month')
```
We see that there is a huge inter-annual as well as inter-month variation for the clear day fraction. We will now sample from this given distribution for each month for P(clear|month) and then continue as before to sample from the clear / cloudy distribution.
```
meanClear = np.zeros((12,1))
stdClear = np.zeros((12,1))
meanCnt = np.zeros((12,2))
stdCnt = np.zeros((12,2))
Clr = inputData['clear']
Cnt = Data[outputVariables]
Wtr = inputData['weather']
MonthDay = [31,28,31,30,31,30,31,31,30,31,30,31] # Actual Days in a month
MonthDay = [200,200,200,200,200,200,200,200,200,200,200,200] # To produce distribution
np.random.seed(1)
plt.figure(figsize=(12,20))
for i in range(1,13):
A = inputData[i]
Ct = Cnt[A==1]
Wt = Wtr[A==1]
Cl = Clr[A==1]
C = np.reshape(Ct[Cl==0],(len(Ct[Cl==0]),1))
C = np.concatenate((C,np.multiply(C,-1)))
E = np.reshape(Ct[Cl==1],(len(Ct[Cl==1]),1))
meanClear[i-1] = np.nanmean(M[:,i-1])
stdClear[i-1] = np.nanstd(M[:,i-1])
meanCnt[i-1,0] = np.nanmean(C)
stdCnt[i-1,0] = np.nanstd(C)
meanCnt[i-1,1] = np.nanmean(E)
stdCnt[i-1,1] = np.nanstd(E)
Cnts = np.zeros((MonthDay[i-1],1))
clearDays = np.floor(np.random.normal(meanClear[i-1],stdClear[i-1]) * MonthDay[i-1])
Cnts[0:int(clearDays)] = np.floor(abs((np.random.normal(meanCnt[i-1,1],stdCnt[i-1,1],(int(clearDays),1)))))
Cnts[int(clearDays):MonthDay[i-1]] = np.floor(abs(np.random.normal(meanCnt[i-1,0],stdCnt[i-1,0],(int(MonthDay[i-1]-clearDays),1))))
plt.subplot(6,4,2*i-1)
plt.hist(Cnts)
plt.axis([0, 250, 0, 50])
plt.title(str(i) + '-Simulated')
plt.subplot(6,4,2*i)
plt.hist(Ct)
plt.axis([0, 250, 0, 50])
plt.title(str(i) + '-Actual')
```
## Markov Chain for Sky Condition
We can now move to even more complex models, where similar to a Recurrent Neural Network, we build a Markov Chain for the sky condition using the transition probabilities from the given data, and then sample the car count from the complete clear / cloudy condition.
```
Clr = ID[:,8]
#### 1 = clear, 0 = cloudy; p11 = clear->clear, p10 = clear->cloudy, p01 = cloudy->clear, p00 = cloudy->cloudy
p10,p11,p01,p00 = 0,0,0,0
for i in range(0,ID.shape[0]-1):
if Clr[i]==1 and Clr[i+1]==1:
p11 = p11+1;
elif Clr[i]==1 and Clr[i+1]==0:
p10 = p10+1;
elif Clr[i]==0 and Clr[i+1]==1:
p01 = p01+1;
elif Clr[i]==0 and Clr[i+1]==0:
p00 = p00+1;
# Transition Probabilities
P11 = float(p11)/(p11+p10)
P10 = float(p10)/(p11+p10)
P01 = float(p01)/(p01+p00)
P00 = float(p00)/(p01+p00)
print p10,p11,p01,p00
print P10,P11,P01,P00
A = inputData['clear']
C = Data[outputVariables]
E = C[A==1]
C = C[A==0]
C = np.concatenate((C,np.multiply(C,-1)))
PDF = np.zeros((2,2))
PDF[0,0] = np.nanmean(C)
PDF[0,1] = np.nanmean(E)
PDF[1,0] = np.nanstd(C)
PDF[1,1] = np.nanstd(E)
T = 2000 # Number of Days to simulate
Cr_Cy = np.zeros((T,1)) # Sky Condition
MC_Cnt = np.zeros((T,1)) # Corresponding Car Count
Cr_Cy[0] = ID[-1,8] # Initial Sky Condition as the last data point given to us
Cnt = Data[outputVariables]
Cnt = np.reshape(Cnt,(len(Cnt),1))
MC_Cnt[0] = Cnt[-1] # Last car count data point given to us
for i in range(1,T):
a = np.random.uniform()
if Cr_Cy[i-1]==1 and a < P11:
Cr_Cy[i] = 1
MC_Cnt[i] = np.floor(abs((np.random.normal(PDF[0,1],PDF[1,1]))))
elif Cr_Cy[i-1]==1 and a > P11:
Cr_Cy[i] = 0
MC_Cnt[i] = np.floor(abs((np.random.normal(PDF[0,0],PDF[1,0]))))
if Cr_Cy[i-1]==0 and a < P00:
Cr_Cy[i] = 0
MC_Cnt[i] = np.floor(abs((np.random.normal(PDF[0,0],PDF[1,0]))))
if Cr_Cy[i-1]==0 and a > P00:
Cr_Cy[i] = 1
MC_Cnt[i] = np.floor(abs((np.random.normal(PDF[0,1],PDF[1,1]))))
#plt.figure()
#plt.plot(Cr_Cy)
#plt.figure()
#plt.plot(MC_Cnt)
plt.figure()
plt.subplot(1,2,1)
plt.hist(MC_Cnt)
plt.title('Simulated')
plt.axis([0, 300, 0, 600])
plt.subplot(1,2,2)
plt.hist(Cnt)
plt.title('Actual')
plt.axis([0, 300, 0, 600])
df = pd.DataFrame(MC_Cnt)
df.to_csv('Next_'+str(T)+'_days.csv') # Continuous Days
```
This Markov Chain model gives an excellent fit to the statistical characteristics of the actual data, and therefore can be used to predict the next 30 days (change T in above code).
The actual car count had a non-stationary structure, with a increasing trend first and then a decreasing trend. The above model does not capture that but since we only need to predict for the next month, the effect of trend can be ignored up to 1st order.
Next, instead of computing the annual transition probabilities, we do it monthwise:
```
Clr = ID[:,8]
#### 1 = clear, 0 = cloudy; p11 = clear->clear, p10 = clear->cloudy, p01 = cloudy->clear, p00 = cloudy->cloudy
P = np.zeros((12,4))
A = inputData['clear']
Cnt = Data[outputVariables]
PDF = np.zeros((12,4))
MonthDay = [31,28,31,30,31,30,31,31,30,31,30,31]
MonthDay = [200,200,200,200,200,200,200,200,200,200,200,200]
Cr_Cy = np.zeros((12,200))
MC_Cnt = np.zeros((12,200))
plt.figure(figsize=(20,20))
for j in range(1,13):
Month = inputData[j]
Car = np.reshape(Cnt[Month==1],(len(Cnt[Month==1]),1))
p10,p11,p01,p00 = 0,0,0,0
Month = np.reshape(Clr[Month==1],(len(Clr[Month==1]),1))
for i in range(0,len(Month)-1):
if Month[i]==1 and Month[i+1]==1:
p11 = p11+1;
elif Month[i]==1 and Month[i+1]==0:
p10 = p10+1;
elif Month[i]==0 and Month[i+1]==1:
p01 = p01+1;
elif Month[i]==0 and Month[i+1]==0:
p00 = p00+1;
P[j-1,0] = float(p11)/(p11+p10)
P[j-1,1] = float(p10)/(p11+p10)
P[j-1,2] = float(p01)/(p01+p00)
P[j-1,3] = float(p00)/(p01+p00)
#print Month.shape
#print Car.shape
E = Car[Month==1]
C = Car[Month==0]
C = np.concatenate((C,np.multiply(C,-1)))
PDF[j-1,0] = np.nanmean(C)
PDF[j-1,1] = np.nanmean(E)
PDF[j-1,2] = np.nanstd(C)
PDF[j-1,3] = np.nanstd(E)
T = MonthDay[j-1]
Cr_Cy[j-1,0] = Month[-1]
#C = np.reshape(Car[Month==1],(len(Car[Month==1]),1))
MC_Cnt[j-1,0] = Car[-1]
for i in range(1,T):
a = np.random.uniform()
if Cr_Cy[j-1,i-1]==1 and a < P[j-1,0]:
Cr_Cy[j-1,i] = 1
MC_Cnt[j-1,i] = np.floor(abs((np.random.normal(PDF[j-1,1],PDF[j-1,3]))))
elif Cr_Cy[j-1,i-1]==1 and a > P[j-1,0]:
Cr_Cy[j-1,i] = 0
MC_Cnt[j-1,i] = np.floor(abs((np.random.normal(PDF[j-1,0],PDF[j-1,2]))))
if Cr_Cy[j-1,i-1]==0 and a < P[j-1,3]:
Cr_Cy[j-1,i] = 0
MC_Cnt[j-1,i] = np.floor(abs((np.random.normal(PDF[j-1,0],PDF[j-1,2]))))
if Cr_Cy[j-1,i-1]==0 and a > P[j-1,3]:
Cr_Cy[j-1,i] = 1
MC_Cnt[j-1,i] = np.floor(abs((np.random.normal(PDF[j-1,1],PDF[j-1,3]))))
#plt.figure()
#plt.plot(Cr_Cy)
#plt.figure()
#plt.plot(MC_Cnt)
plt.subplot(6,4,2*j-1)
plt.hist(MC_Cnt[j-1,:])
plt.title(str(j) + '-simulated')
plt.axis([0, 250, 0, 50])
plt.subplot(6,4,2*j)
plt.hist(Car)
plt.title(str(j) + '-actual')
plt.axis([0, 250, 0, 50])
df = pd.DataFrame(MC_Cnt.T)
df.to_csv('Next_'+str(200)+'_days_Month.csv') # 0 - January ----> 11 - December
```
Even this model produces good statistics and can be used for prediction of car count for any given month.
# Results
The first step is to identify the independent variables. The dependent variable is the Car Count, but how does it depend on other variables!!
To answer this question, we looked at the correlations between different independent (numerical and categorical variables) and the car count.
1) I started with visualization and a linear regression model for the car count and weather index. No clear dependency between the value of the weather index and the car count was apparent.
2) I then studied if the car count varied with the day of the week. To do this we looked at the distribution of the car count for each day of the week for the complete record. These distributions showed that only slighest of dependencies (if any) exist and therefore we ignore this variable in future models.
3) Next I examined the correlation between the month of the year and the car count, and again utilized the distributions of the car count with respect to the month. These distribution had slight variations.
4) Lastly, I looked at how the car count in the parking lot varies with the sky conditions. Both simple data visualization and histrograms showed a high correlation between the car count and the sky condition. Clear sky led to more cars, while cloudy skies led to a decrease in the car count. I developed two separate linear models for clear / cloudy skies, and saw that the combined mean squared error was lower than for the single model.
5) I then verified the correlation between the weather index and the sky conditions to see if they were related, but found otherwise, while the the car count was highly correlated with the sky conditions supported by the cross-correlation function.
6) Using the Bayes Rule, I then computed the clear days fraction in any given month. Then for each month, I developed two separate models for clear / cloudy skies, and then using the clear day fraction, scaled them. This scaled model was similar to using just one model without any regard to the sky conditions.
7) Since the only correlation being observed was between the car count and sky conditions, I then moved to probabilistic models. Using the clear day fraction for a given month computed in the previous step, I first computed the clear days and cloudy days in a month and then sampled the car count from its distribution for that month and sky condition. To perform this, I had to first modify the "cloudy" distribution to make it Gaussian. This sampling provided a better model than the linear models above, but there is no interannual variation.
8) To account for the interannual variation, I now also sample the clear day fraction from the clear day fraction for each month and year. And then proceed as before to compute the total clear days in a month and then sampling from the respective distribution. This model also has good statistics and can be used for prediction when one is only interesed in the distribution of car counts in a month, and not the day-to-day variation.
9) To add complexity towards day-to-day variation, I built a Markov Chain model, similar to a Recurrent Neural Network, where I computed the transition probabilities for the sky conditions. Then combining a simulation of this Markov Chain model with the sampling of car count from the complete car count distribution w.r.t. sky conditions (ignoring the month), gave a good enough match between the observed and the simulated car count distribution. This gives a prediction for any number of continuous days in a year.
10) Lastly, I modified the previous model to make it dependent on the month and computed the transition probabilities separately for each month. This gives us a prediction for any number of days for a given month.
## Future Work
1) I used the complete data set for training the Markov chain models, but given more time, it can be trained iteratively starting from a smaller training set and increasing its size and checking for errors.
2) Markov chain models only look at the previous state. Since we know that there is a long term trend in the data, the Markov Chain model cannon capture it. To do this we can build more complex models that also than into account long time memory.
3) The transition probabilities in the Markov Chain models can be updated with each time step of prediction, rather than fixed probabilities.
4) The two Markov Chain models have different dynamics: the first ignores the monthly dependencies, while the second does not take into account the continuity of probabilities between different months. A more complex model can be generated as some combination of these two, where both the monthly dependencies and continuity exist.
| github_jupyter |
# Tables
```
cd ..
import NotebookImport
from DX_screen import *
from metaPCNA import *
```
# GSEA
### Gene level fraction upregulated statistic
```
gs2 = gene_sets.ix[dx_rna.index].fillna(0)
rr = screen_feature(dx_rna.frac, rev_kruskal, gs2.T,
align=False)
fp = (1.*gene_sets.T * dx_rna.frac).T.dropna().replace(0, np.nan).mean().order()
fp.name = 'mean frac'
rr.join(fp).to_csv(FIGDIR + 'f_up_gene_sets.csv')
ff_u = filter_pathway_hits(rr.ix[ti(fp>.5)].p.order(), gs2)
ff_p = filter_pathway_hits(rr.ix[ti(fp<.5)].p.order(), gs2)
ff = ff_u.append(ff_p)
selected = rr.ix[ff[ff < .00001].index].join(fp)
selected.sort('p')
selected.to_csv(FIGDIR + 'f_up_gene_sets_selected.csv')
```
Looking for subsets of the cell-cycle pathway
```
d = pd.DataFrame({g: gs2['REACTOME_CELL_CYCLE'] for g in gs2.columns})
a,b = odds_ratio_df(d.T>0, gs2.T>0)
dd = rr.ix[ti((a > 100) & (rr.q < 10e-15))].join(fp).sort(fp.name, ascending=False)
filter_pathway_hits(dd, gs2)
(combine(gs2['REACTOME_M_G1_TRANSITION']>0,
gs2['REACTOME_DEPOSITION_OF_NEW_CENPA_CONTAINING_NUCLEOSOMES_AT_THE_CENTROMERE']>0)
).value_counts()
f2 = fp.ix[ti(rr.q < .00001)]
ff_u = filter_pathway_hits(fp.ix[ti(f2>.5)].order()[::-1], gs2)
ff_p = filter_pathway_hits(fp.ix[ti(f2<.5)].order(), gs2)
ff = ff_u.append(ff_p)
selected = rr.ix[ff.index].join(f2)
selected.ix[(f2 - .5).abs().order().index[::-1]].dropna()
selected.to_csv(FIGDIR + 'f_up_gene_sets_selected_fc.csv')
```
### Gene level proliferation statistic
```
gs2 = gene_sets.ix[pcna_corr.dropna().index].fillna(0)
rr = screen_feature(pcna_corr, rev_kruskal, gs2.T,
align=False)
fp = (1.*gene_sets.T * pcna_corr).T.dropna().replace(0, np.nan).mean().order()
fp.name = 'mean score'
rr.join(fp).to_csv(FIGDIR + 'pcna_gene_sets.csv')
ff_u = filter_pathway_hits(rr.ix[ti(fp>0)].p.order(), gs2)
ff_p = filter_pathway_hits(rr.ix[ti(fp<0)].p.order(), gs2)
ff = ff_u.append(ff_p)
selected = rr.ix[ff[ff < .00001].index].join(fp)
selected.sort('p')
selected.to_csv(FIGDIR + 'pcna_gene_sets_selected.csv')
f2 = fp.ix[ti(rr.q < .0001)]
ff_u = filter_pathway_hits(fp.ix[ti(f2>0)].order()[::-1], gs2)
ff_p = filter_pathway_hits(fp.ix[ti(f2<0)].order(), gs2)
ff = ff_u.append(ff_p)
selected = rr.ix[ff.index].join(f2)
selected.sort('p')
selected.to_csv(FIGDIR + 'pcna_gene_sets_selected_fc.csv')
```
### Gene level detrended fraction upregulated statistic
```
gs2 = gene_sets.ix[f_win.dropna().index].fillna(0)
rr = screen_feature(f_win, rev_kruskal, gs2.T,
align=False)
fp = (1.*gene_sets.T * f_win).T.dropna().replace(0, np.nan).mean().order()
fp.name = 'mean score'
(rr.q < .00001).value_counts()
rr.join(fp).to_csv(FIGDIR + 'detrended_fup_sets.csv')
f2 = fp.ix[ti(rr.q < .0001)]
ff_u = filter_pathway_hits(rr.ix[ti(f2>0)].p.order(), gs2)
ff_p = filter_pathway_hits(rr.ix[ti(f2<0)].p.order(), gs2)
ff = ff_u.append(ff_p)
selected = rr.ix[ff[ff < .00001].index].join(fp)
selected.sort('p')
selected.to_csv(FIGDIR + 'detrended_fup_sets_selected.csv')
f2 = fp.ix[ti(rr.q < .0001)]
ff_u = filter_pathway_hits(fp.ix[ti(f2>0)].order()[::-1], gs2)
ff_p = filter_pathway_hits(fp.ix[ti(f2<0)].order(), gs2)
ff = ff_u.append(ff_p)
selected = rr.ix[ff.index].join(f2)
selected.sort('p')
selected.to_csv(FIGDIR + 'detrended_fup_sets_selected_fc.csv')
```
### Diggining into the different telomere gene subsets
```
[g for g in gs2 if 'TELO' in g]
a = gs2['REACTOME_PACKAGING_OF_TELOMERE_ENDS']>0
b = gs2['REACTOME_EXTENSION_OF_TELOMERES']>0
a.name = 'end packaging'
b.name = 'extension'
cc = combine(a,b).replace('neither', np.nan).dropna()
fig, ax = subplots(figsize=(4,4))
series_scatter(dx_rna.frac, pcna_corr.ix[ti(cc == 'end packaging')],
ax=ax, color=colors[0], ann=None, alpha=1, s=30)
series_scatter(dx_rna.frac, pcna_corr.ix[ti(cc == 'extension')],
ax=ax, color=colors[1], ann=None, alpha=1, s=30)
ax.set_xlabel('Fraction Upregulated')
ax.set_ylabel('Proliferation Score')
ax.legend(['end packaging','extension'], loc='upper left', frameon=True)
prettify_ax(ax)
fig.tight_layout()
fig.savefig(FIGDIR + 'S3_Fig.pdf')
violin_plot_pandas(cc, f_win)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/NeuromatchAcademy/course-content/blob/master/tutorials/W3D5_NetworkCausality/student/W3D5_Tutorial3.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Tutorial 3: Simultaneous fitting/regression
**Week 3, Day 5: Network Causality**
**By Neuromatch Academy**
**Content creators**: Ari Benjamin, Tony Liu, Konrad Kording
**Content reviewers**: Mike X Cohen, Madineh Sarvestani, Yoni Friedman, Ella Batty, Michael Waskom
**Our 2021 Sponsors, including Presenting Sponsor Facebook Reality Labs**
<p align='center'><img src='https://github.com/NeuromatchAcademy/widgets/blob/master/sponsors.png?raw=True'/></p>
---
# Tutorial objectives
*Estimated timing of tutorial: 20 min*
This is tutorial 3 on our day of examining causality. Below is the high level outline of what we'll cover today, with the sections we will focus on in this notebook in bold:
1. Master definitions of causality
2. Understand that estimating causality is possible
3. Learn 4 different methods and understand when they fail
1. perturbations
2. correlations
3. **simultaneous fitting/regression**
4. instrumental variables
### Notebook 3 objectives
In tutorial 2 we explored correlation as an approximation for causation and learned that correlation $\neq$ causation for larger networks. However, computing correlations is a rather simple approach, and you may be wondering: will more sophisticated techniques allow us to better estimate causality? Can't we control for things?
Here we'll use some common advanced (but controversial) methods that estimate causality from observational data. These methods rely on fitting a function to our data directly, instead of trying to use perturbations or correlations. Since we have the full closed-form equation of our system, we can try these methods and see how well they work in estimating causal connectivity when there are no perturbations. Specifically, we will:
- Learn about more advanced (but also controversial) techniques for estimating causality
- conditional probabilities (**regression**)
- Explore limitations and failure modes
- understand the problem of **omitted variable bias**
```
# @title Tutorial slides
# @markdown These are the slides for the videos in all tutorials today
from IPython.display import IFrame
IFrame(src=f"https://mfr.ca-1.osf.io/render?url=https://osf.io/gp4m9/?direct%26mode=render%26action=download%26mode=render", width=854, height=480)
```
---
# Setup
```
# Imports
import numpy as np
import matplotlib.pyplot as plt
from sklearn.multioutput import MultiOutputRegressor
from sklearn.linear_model import Lasso
# @title Figure Settings
import ipywidgets as widgets # interactive display
%config InlineBackend.figure_format = 'retina'
plt.style.use("https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/nma.mplstyle")
# @title Plotting Functions
def see_neurons(A, ax, ratio_observed=1, arrows=True):
"""
Visualizes the connectivity matrix.
Args:
A (np.ndarray): the connectivity matrix of shape (n_neurons, n_neurons)
ax (plt.axis): the matplotlib axis to display on
Returns:
Nothing, but visualizes A.
"""
n = len(A)
ax.set_aspect('equal')
thetas = np.linspace(0, np.pi * 2, n, endpoint=False)
x, y = np.cos(thetas), np.sin(thetas),
if arrows:
for i in range(n):
for j in range(n):
if A[i, j] > 0:
ax.arrow(x[i], y[i], x[j] - x[i], y[j] - y[i], color='k', head_width=.05,
width = A[i, j] / 25,shape='right', length_includes_head=True,
alpha = .2)
if ratio_observed < 1:
nn = int(n * ratio_observed)
ax.scatter(x[:nn], y[:nn], c='r', s=150, label='Observed')
ax.scatter(x[nn:], y[nn:], c='b', s=150, label='Unobserved')
ax.legend(fontsize=15)
else:
ax.scatter(x, y, c='k', s=150)
ax.axis('off')
def plot_connectivity_matrix(A, ax=None):
"""Plot the (weighted) connectivity matrix A as a heatmap
Args:
A (ndarray): connectivity matrix (n_neurons by n_neurons)
ax: axis on which to display connectivity matrix
"""
if ax is None:
ax = plt.gca()
lim = np.abs(A).max()
ax.imshow(A, vmin=-lim, vmax=lim, cmap="coolwarm")
# @title Helper Functions
def sigmoid(x):
"""
Compute sigmoid nonlinearity element-wise on x.
Args:
x (np.ndarray): the numpy data array we want to transform
Returns
(np.ndarray): x with sigmoid nonlinearity applied
"""
return 1 / (1 + np.exp(-x))
def create_connectivity(n_neurons, random_state=42, p=0.9):
"""
Generate our nxn causal connectivity matrix.
Args:
n_neurons (int): the number of neurons in our system.
random_state (int): random seed for reproducibility
Returns:
A (np.ndarray): our 0.1 sparse connectivity matrix
"""
np.random.seed(random_state)
A_0 = np.random.choice([0, 1], size=(n_neurons, n_neurons), p=[p, 1 - p])
# set the timescale of the dynamical system to about 100 steps
_, s_vals, _ = np.linalg.svd(A_0)
A = A_0 / (1.01 * s_vals[0])
# _, s_val_test, _ = np.linalg.svd(A)
# assert s_val_test[0] < 1, "largest singular value >= 1"
return A
def simulate_neurons(A, timesteps, random_state=42):
"""
Simulates a dynamical system for the specified number of neurons and timesteps.
Args:
A (np.array): the connectivity matrix
timesteps (int): the number of timesteps to simulate our system.
random_state (int): random seed for reproducibility
Returns:
- X has shape (n_neurons, timeteps).
"""
np.random.seed(random_state)
n_neurons = len(A)
X = np.zeros((n_neurons, timesteps))
for t in range(timesteps - 1):
# solution
epsilon = np.random.multivariate_normal(np.zeros(n_neurons), np.eye(n_neurons))
X[:, t + 1] = sigmoid(A.dot(X[:, t]) + epsilon)
assert epsilon.shape == (n_neurons,)
return X
def get_sys_corr(n_neurons, timesteps, random_state=42, neuron_idx=None):
"""
A wrapper function for our correlation calculations between A and R.
Args:
n_neurons (int): the number of neurons in our system.
timesteps (int): the number of timesteps to simulate our system.
random_state (int): seed for reproducibility
neuron_idx (int): optionally provide a neuron idx to slice out
Returns:
A single float correlation value representing the similarity between A and R
"""
A = create_connectivity(n_neurons, random_state)
X = simulate_neurons(A, timesteps)
R = correlation_for_all_neurons(X)
return np.corrcoef(A.flatten(), R.flatten())[0, 1]
def correlation_for_all_neurons(X):
"""Computes the connectivity matrix for the all neurons using correlations
Args:
X: the matrix of activities
Returns:
estimated_connectivity (np.ndarray): estimated connectivity for the selected neuron, of shape (n_neurons,)
"""
n_neurons = len(X)
S = np.concatenate([X[:, 1:], X[:, :-1]], axis=0)
R = np.corrcoef(S)[:n_neurons, n_neurons:]
return R
```
The helper functions defined above are:
- `sigmoid`: computes sigmoid nonlinearity element-wise on input, from Tutorial 1
- `create_connectivity`: generates nxn causal connectivity matrix., from Tutorial 1
- `simulate_neurons`: simulates a dynamical system for the specified number of neurons and timesteps, from Tutorial 1
- `get_sys_corr`: a wrapper function for correlation calculations between A and R, from Tutorial 2
- `correlation_for_all_neurons`: computes the connectivity matrix for the all neurons using correlations, from Tutorial 2
---
# Section 1: Regression: recovering connectivity by model fitting
```
# @title Video 1: Regression approach
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = 'https://player.bilibili.com/player.html?bvid={0}&page={1}'.format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id="BV1m54y1q78b", width=854, height=480, fs=1)
print('Video available at https://www.bilibili.com/video/{0}'.format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="Av4LaXZdgDo", width=854, height=480, fs=1, rel=0)
print('Video available at https://youtube.com/watch?v=' + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
```
You may be familiar with the idea that correlation only implies causation when there no hidden *confounders*. This aligns with our intuition that correlation only implies causality when no alternative variables could explain away a correlation.
**A confounding example**:
Suppose you observe that people who sleep more do better in school. It's a nice correlation. But what else could explain it? Maybe people who sleep more are richer, don't work a second job, and have time to actually do homework. If you want to ask if sleep *causes* better grades, and want to answer that with correlations, you have to control for all possible confounds.
A confound is any variable that affects both the outcome and your original covariate. In our example, confounds are things that affect both sleep and grades.
**Controlling for a confound**:
Confonds can be controlled for by adding them as covariates in a regression. But for your coefficients to be causal effects, you need three things:
1. **All** confounds are included as covariates
2. Your regression assumes the same mathematical form of how covariates relate to outcomes (linear, GLM, etc.)
3. No covariates are caused *by* both the treatment (original variable) and the outcome. These are [colliders](https://en.wikipedia.org/wiki/Collider_(statistics)); we won't introduce it today (but Google it on your own time! Colliders are very counterintuitive.)
In the real world it is very hard to guarantee these conditions are met. In the brain it's even harder (as we can't measure all neurons). Luckily today we simulated the system ourselves.
```
# @title Video 2: Fitting a GLM
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = 'https://player.bilibili.com/player.html?bvid={0}&page={1}'.format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id="BV16p4y1S7yE", width=854, height=480, fs=1)
print('Video available at https://www.bilibili.com/video/{0}'.format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="GvMj9hRv5Ak", width=854, height=480, fs=1, rel=0)
print('Video available at https://youtube.com/watch?v=' + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
```
Recall that in our system each neuron effects every other via:
$$
\vec{x}_{t+1} = \sigma(A\vec{x}_t + \epsilon_t),
$$
where $\sigma$ is our sigmoid nonlinearity from before: $\sigma(x) = \frac{1}{1 + e^{-x}}$
Our system is a closed system, too, so there are no omitted variables. The regression coefficients should be the causal effect. Are they?
We will use a regression approach to estimate the causal influence of all neurons to neuron #1. Specifically, we will use linear regression to determine the $A$ in:
$$
\sigma^{-1}(\vec{x}_{t+1}) = A\vec{x}_t + \epsilon_t ,
$$
where $\sigma^{-1}$ is the inverse sigmoid transformation, also sometimes referred to as the **logit** transformation: $\sigma^{-1}(x) = \log(\frac{x}{1-x})$.
Let $W$ be the $\vec{x}_t$ values, up to the second-to-last timestep $T-1$:
$$
W =
\begin{bmatrix}
\mid & \mid & ... & \mid \\
\vec{x}_0 & \vec{x}_1 & ... & \vec{x}_{T-1} \\
\mid & \mid & ... & \mid
\end{bmatrix}_{n \times (T-1)}
$$
Let $Y$ be the $\vec{x}_{t+1}$ values for a selected neuron, indexed by $i$, starting from the second timestep up to the last timestep $T$:
$$
Y =
\begin{bmatrix}
x_{i,1} & x_{i,2} & ... & x_{i, T} \\
\end{bmatrix}_{1 \times (T-1)}
$$
You will then fit the following model:
$$
\sigma^{-1}(Y^T) = W^TV
$$
where $V$ is the $n \times 1$ coefficient matrix of this regression, which will be the estimated connectivity matrix between the selected neuron and the rest of the neurons.
**Review**: As you learned in Week 1, *lasso* a.k.a. **$L_1$ regularization** causes the coefficients to be sparse, containing mostly zeros. Think about why we want this here.
## Coding Exercise 1: Use linear regression plus lasso to estimate causal connectivities
You will now create a function to fit the above regression model and V. We will then call this function to examine how close the regression vs the correlation is to true causality.
**Code**:
You'll notice that we've transposed both $Y$ and $W$ here and in the code we've already provided below. Why is that?
This is because the machine learning models provided in scikit-learn expect the *rows* of the input data to be the observations, while the *columns* are the variables. We have that inverted in our definitions of $Y$ and $W$, with the timesteps of our system (the observations) as the columns. So we transpose both matrices to make the matrix orientation correct for scikit-learn.
- Because of the abstraction provided by scikit-learn, fitting this regression will just be a call to initialize the `Lasso()` estimator and a call to the `fit()` function
- Use the following hyperparameters for the `Lasso` estimator:
- `alpha = 0.01`
- `fit_intercept = False`
- How do we obtain $V$ from the fitted model?
We will use the helper function `logit`.
```
# @markdown Execute this cell to enable helper function `logit`
def logit(x):
"""
Applies the logit (inverse sigmoid) transformation
Args:
x (np.ndarray): the numpy data array we want to transform
Returns
(np.ndarray): x with logit nonlinearity applied
"""
return np.log(x/(1-x))
def get_regression_estimate(X, neuron_idx):
"""
Estimates the connectivity matrix using lasso regression.
Args:
X (np.ndarray): our simulated system of shape (n_neurons, timesteps)
neuron_idx (int): a neuron index to compute connectivity for
Returns:
V (np.ndarray): estimated connectivity matrix of shape (n_neurons, n_neurons).
if neuron_idx is specified, V is of shape (n_neurons,).
"""
# Extract Y and W as defined above
W = X[:, :-1].transpose()
Y = X[[neuron_idx], 1:].transpose()
# Apply inverse sigmoid transformation
Y = logit(Y)
############################################################################
## TODO: Insert your code here to fit a regressor with Lasso. Lasso captures
## our assumption that most connections are precisely 0.
## Fill in function and remove
raise NotImplementedError("Please complete the regression exercise")
############################################################################
# Initialize regression model with no intercept and alpha=0.01
regression = ...
# Fit regression to the data
regression.fit(...)
V = regression.coef_
return V
# Set parameters
n_neurons = 50 # the size of our system
timesteps = 10000 # the number of timesteps to take
random_state = 42
neuron_idx = 1
# Set up system and simulate
A = create_connectivity(n_neurons, random_state)
X = simulate_neurons(A, timesteps)
# Estimate causality with regression
V = get_regression_estimate(X, neuron_idx)
print("Regression: correlation of estimated connectivity with true connectivity: {:.3f}".format(np.corrcoef(A[neuron_idx, :], V)[1, 0]))
print("Lagged correlation of estimated connectivity with true connectivity: {:.3f}".format(get_sys_corr(n_neurons, timesteps, random_state, neuron_idx=neuron_idx)))
```
[*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W3D5_NetworkCausality/solutions/W3D5_Tutorial3_Solution_17d9b1a7.py)
You should find that using regression, our estimated connectivity matrix has a correlation of 0.865 with the true connectivity matrix. With correlation, our estimated connectivity matrix has a correlation of 0.703 with the true connectivity matrix.
We can see from these numbers that multiple regression is better than simple correlation for estimating connectivity.
---
# Section 2: Partially Observed Systems
*Estimated timing to here from start of tutorial: 10 min*
If we are unable to observe the entire system, **omitted variable bias** becomes a problem. If we don't have access to all the neurons, and so therefore can't control for them, can we still estimate the causal effect accurately?
```
# @title Video 3: Omitted variable bias
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = 'https://player.bilibili.com/player.html?bvid={0}&page={1}'.format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id="BV1ov411i7dc", width=854, height=480, fs=1)
print('Video available at https://www.bilibili.com/video/{0}'.format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="5CCib6CTMac", width=854, height=480, fs=1, rel=0)
print('Video available at https://youtube.com/watch?v=' + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
```
**Video correction**: the labels "connectivity from"/"connectivity to" are swapped in the video but fixed in the figures/demos below
We first visualize different subsets of the connectivity matrix when we observe 75% of the neurons vs 25%.
Recall the meaning of entries in our connectivity matrix: $A[i,j] = 1$ means a connectivity **from** neuron $i$ **to** neuron $j$ with strength $1$.
```
#@markdown Execute this cell to visualize subsets of connectivity matrix
# Run this cell to visualize the subsets of variables we observe
n_neurons = 25
A = create_connectivity(n_neurons)
fig, axs = plt.subplots(2, 2, figsize=(10, 10))
ratio_observed = [0.75, 0.25] # the proportion of neurons observed in our system
for i, ratio in enumerate(ratio_observed):
sel_idx = int(n_neurons * ratio)
offset = np.zeros((n_neurons, n_neurons))
axs[i,1].title.set_text("{}% neurons observed".format(int(ratio * 100)))
offset[:sel_idx, :sel_idx] = 1 + A[:sel_idx, :sel_idx]
im = axs[i, 1].imshow(offset, cmap="coolwarm", vmin=0, vmax=A.max() + 1)
axs[i, 1].set_xlabel("Connectivity from")
axs[i, 1].set_ylabel("Connectivity to")
plt.colorbar(im, ax=axs[i, 1], fraction=0.046, pad=0.04)
see_neurons(A,axs[i, 0],ratio)
plt.suptitle("Visualizing subsets of the connectivity matrix", y = 1.05)
plt.show()
```
### Interactive Demo 3: Regression performance as a function of the number of observed neurons
We will first change the number of observed neurons in the network and inspect the resulting estimates of connectivity in this interactive demo. How does the estimated connectivity differ?
```
# @markdown Execute this cell to get helper functions `get_regression_estimate_full_connectivity` and `get_regression_corr_full_connectivity`
def get_regression_estimate_full_connectivity(X):
"""
Estimates the connectivity matrix using lasso regression.
Args:
X (np.ndarray): our simulated system of shape (n_neurons, timesteps)
neuron_idx (int): optionally provide a neuron idx to compute connectivity for
Returns:
V (np.ndarray): estimated connectivity matrix of shape (n_neurons, n_neurons).
if neuron_idx is specified, V is of shape (n_neurons,).
"""
n_neurons = X.shape[0]
# Extract Y and W as defined above
W = X[:, :-1].transpose()
Y = X[:, 1:].transpose()
# apply inverse sigmoid transformation
Y = logit(Y)
# fit multioutput regression
reg = MultiOutputRegressor(Lasso(fit_intercept=False,
alpha=0.01, max_iter=250 ), n_jobs=-1)
reg.fit(W, Y)
V = np.zeros((n_neurons, n_neurons))
for i, estimator in enumerate(reg.estimators_):
V[i, :] = estimator.coef_
return V
def get_regression_corr_full_connectivity(n_neurons, A, X, observed_ratio, regression_args):
"""
A wrapper function for our correlation calculations between A and the V estimated
from regression.
Args:
n_neurons (int): number of neurons
A (np.ndarray): connectivity matrix
X (np.ndarray): dynamical system
observed_ratio (float): the proportion of n_neurons observed, must be betweem 0 and 1.
regression_args (dict): dictionary of lasso regression arguments and hyperparameters
Returns:
A single float correlation value representing the similarity between A and R
"""
assert (observed_ratio > 0) and (observed_ratio <= 1)
sel_idx = np.clip(int(n_neurons*observed_ratio), 1, n_neurons)
sel_X = X[:sel_idx, :]
sel_A = A[:sel_idx, :sel_idx]
sel_V = get_regression_estimate_full_connectivity(sel_X)
return np.corrcoef(sel_A.flatten(), sel_V.flatten())[1,0], sel_V
# @markdown Execute this cell to enable demo. the plots will take a few seconds to update after moving the slider.
n_neurons = 50
A = create_connectivity(n_neurons, random_state=42)
X = simulate_neurons(A, 4000, random_state=42)
reg_args = {
"fit_intercept": False,
"alpha": 0.001
}
@widgets.interact(n_observed = widgets.IntSlider(min = 5, max = 45, step = 5, continuous_update=False))
def plot_observed(n_observed):
to_neuron = 0
fig, axs = plt.subplots(1, 3, figsize=(15, 5))
sel_idx = n_observed
ratio = (n_observed) / n_neurons
offset = np.zeros((n_neurons, n_neurons))
axs[0].title.set_text("{}% neurons observed".format(int(ratio * 100)))
offset[:sel_idx, :sel_idx] = 1 + A[:sel_idx, :sel_idx]
im = axs[1].imshow(offset, cmap="coolwarm", vmin=0, vmax=A.max() + 1)
plt.colorbar(im, ax=axs[1], fraction=0.046, pad=0.04)
see_neurons(A,axs[0], ratio, False)
corr, R = get_regression_corr_full_connectivity(n_neurons,
A,
X,
ratio,
reg_args)
#rect = patches.Rectangle((-.5,to_neuron-.5),n_observed,1,linewidth=2,edgecolor='k',facecolor='none')
#axs[1].add_patch(rect)
big_R = np.zeros(A.shape)
big_R[:sel_idx, :sel_idx] = 1 + R
#big_R[to_neuron, :sel_idx] = 1 + R
im = axs[2].imshow(big_R, cmap="coolwarm", vmin=0, vmax=A.max() + 1)
plt.colorbar(im, ax=axs[2],fraction=0.046, pad=0.04)
c = 'w' if n_observed<(n_neurons-3) else 'k'
axs[2].text(0,n_observed+3,"Correlation : {:.2f}".format(corr), color=c, size=15)
#axs[2].axis("off")
axs[1].title.set_text("True connectivity")
axs[1].set_xlabel("Connectivity from")
axs[1].set_ylabel("Connectivity to")
axs[2].title.set_text("Estimated connectivity")
axs[2].set_xlabel("Connectivity from")
#axs[2].set_ylabel("Connectivity to")
```
Next, we will inspect a plot of the correlation between true and estimated connectivity matrices vs the percent of neurons observed over multiple trials.
What is the relationship that you see between performance and the number of neurons observed?
**Note:** the cell below will take about 25-30 seconds to run.
```
# @markdown Plot correlation vs. subsampling
import warnings
warnings.filterwarnings('ignore')
# we'll simulate many systems for various ratios of observed neurons
n_neurons = 50
timesteps = 5000
ratio_observed = [1, 0.75, 0.5, .25, .12] # the proportion of neurons observed in our system
n_trials = 3 # run it this many times to get variability in our results
reg_args = {
"fit_intercept": False,
"alpha": 0.001
}
corr_data = np.zeros((n_trials, len(ratio_observed)))
for trial in range(n_trials):
A = create_connectivity(n_neurons, random_state=trial)
X = simulate_neurons(A, timesteps)
print("simulating trial {} of {}".format(trial + 1, n_trials))
for j, ratio in enumerate(ratio_observed):
result,_ = get_regression_corr_full_connectivity(n_neurons,
A,
X,
ratio,
reg_args)
corr_data[trial, j] = result
corr_mean = np.nanmean(corr_data, axis=0)
corr_std = np.nanstd(corr_data, axis=0)
plt.plot(np.asarray(ratio_observed) * 100, corr_mean)
plt.fill_between(np.asarray(ratio_observed) * 100,
corr_mean - corr_std,
corr_mean + corr_std,
alpha=.2)
plt.xlim([100, 10])
plt.xlabel("Percent of neurons observed")
plt.ylabel("connectivity matrices correlation")
plt.title("Performance of regression as a function of the number of neurons observed");
```
---
# Summary
*Estimated timing of tutorial: 20 min*
```
# @title Video 4: Summary
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = 'https://player.bilibili.com/player.html?bvid={0}&page={1}'.format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id="BV1bh411o73r", width=854, height=480, fs=1)
print('Video available at https://www.bilibili.com/video/{0}'.format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="T1uGf1H31wE", width=854, height=480, fs=1, rel=0)
print('Video available at https://youtube.com/watch?v=' + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
```
In this tutorial, we explored:
1) Using regression for estimating causality
2) The problem of ommitted variable bias, and how it arises in practice
| github_jupyter |
```
import os
import cv2
import math
import warnings
import numpy as np
import pandas as pd
import seaborn as sns
import tensorflow as tf
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix, fbeta_score
from keras import optimizers
from keras import backend as K
from keras.models import Sequential, Model
from keras import applications
from keras.preprocessing.image import ImageDataGenerator
from keras.callbacks import LearningRateScheduler, EarlyStopping, ReduceLROnPlateau
from keras.layers import Dense, Dropout, Flatten, Conv2D, MaxPool2D, Activation, BatchNormalization, GlobalAveragePooling2D, Input
# Set seeds to make the experiment more reproducible.
from tensorflow import set_random_seed
from numpy.random import seed
set_random_seed(0)
seed(0)
%matplotlib inline
sns.set(style="whitegrid")
warnings.filterwarnings("ignore")
train = pd.read_csv('../input/imet-2019-fgvc6/train.csv')
labels = pd.read_csv('../input/imet-2019-fgvc6/labels.csv')
test = pd.read_csv('../input/imet-2019-fgvc6/sample_submission.csv')
train["attribute_ids"] = train["attribute_ids"].apply(lambda x:x.split(" "))
train["id"] = train["id"].apply(lambda x: x + ".png")
test["id"] = test["id"].apply(lambda x: x + ".png")
print('Number of train samples: ', train.shape[0])
print('Number of test samples: ', test.shape[0])
print('Number of labels: ', labels.shape[0])
display(train.head())
display(labels.head())
```
### Model parameters
```
# Model parameters
BATCH_SIZE = 64
EPOCHS = 30
WARMUP_EPOCHS = 2
LEARNING_RATE = 0.001
WARMUP_LEARNING_RATE = 0.001
HEIGHT = 156
WIDTH = 156
CANAL = 3
N_CLASSES = labels.shape[0]
ES_PATIENCE = 4
RLROP_PATIENCE = 2
DECAY_DROP = 0.5
DECAY_EPOCHS = 10
def f2_score_thr(threshold=0.5):
def f2_score(y_true, y_pred):
beta = 2
y_pred = K.cast(K.greater(K.clip(y_pred, 0, 1), threshold), K.floatx())
true_positives = K.sum(K.clip(y_true * y_pred, 0, 1), axis=1)
predicted_positives = K.sum(K.clip(y_pred, 0, 1), axis=1)
possible_positives = K.sum(K.clip(y_true, 0, 1), axis=1)
precision = true_positives / (predicted_positives + K.epsilon())
recall = true_positives / (possible_positives + K.epsilon())
return K.mean(((1+beta**2)*precision*recall) / ((beta**2)*precision+recall+K.epsilon()))
return f2_score
def custom_f2(y_true, y_pred):
beta = 2
tp = np.sum((y_true == 1) & (y_pred == 1))
tn = np.sum((y_true == 0) & (y_pred == 0))
fp = np.sum((y_true == 0) & (y_pred == 1))
fn = np.sum((y_true == 1) & (y_pred == 0))
p = tp / (tp + fp + K.epsilon())
r = tp / (tp + fn + K.epsilon())
f2 = (1+beta**2)*p*r / (p*beta**2 + r + 1e-15)
return f2
def step_decay(epoch):
initial_lrate = LEARNING_RATE
drop = DECAY_DROP
epochs_drop = DECAY_EPOCHS
lrate = initial_lrate * math.pow(drop, math.floor((1+epoch)/epochs_drop))
return lrate
train_datagen=ImageDataGenerator(rescale=1./255, validation_split=0.25,
horizontal_flip=True)
train_generator=train_datagen.flow_from_dataframe(
dataframe=train,
directory="../input/imet-2019-fgvc6/train",
x_col="id",
y_col="attribute_ids",
batch_size=BATCH_SIZE,
shuffle=True,
class_mode="categorical",
target_size=(HEIGHT, WIDTH),
subset='training')
valid_generator=train_datagen.flow_from_dataframe(
dataframe=train,
directory="../input/imet-2019-fgvc6/train",
x_col="id",
y_col="attribute_ids",
batch_size=BATCH_SIZE,
shuffle=True,
class_mode="categorical",
target_size=(HEIGHT, WIDTH),
subset='validation')
test_datagen = ImageDataGenerator(rescale=1./255)
test_generator = test_datagen.flow_from_dataframe(
dataframe=test,
directory = "../input/imet-2019-fgvc6/test",
x_col="id",
target_size=(HEIGHT, WIDTH),
batch_size=1,
shuffle=False,
class_mode=None)
```
### Model
```
def create_model(input_shape, n_out):
input_tensor = Input(shape=input_shape)
base_model = applications.Xception(weights=None, include_top=False,
input_tensor=input_tensor)
base_model.load_weights('../input/xception/xception_weights_tf_dim_ordering_tf_kernels_notop.h5')
x = GlobalAveragePooling2D()(base_model.output)
x = Dense(1024, activation='relu')(x)
x = Dropout(0.5)(x)
final_output = Dense(n_out, activation='sigmoid', name='final_output')(x)
model = Model(input_tensor, final_output)
return model
model = create_model(input_shape=(HEIGHT, WIDTH, CANAL), n_out=N_CLASSES)
for layer in model.layers:
layer.trainable = False
for i in range(-5,0):
model.layers[i].trainable = True
metrics = ["accuracy", "categorical_accuracy"]
optimizer = optimizers.Adam(lr=WARMUP_LEARNING_RATE)
model.compile(optimizer=optimizer, loss="binary_crossentropy", metrics=metrics)
model.summary()
```
#### Train top layers
```
STEP_SIZE_TRAIN = train_generator.n//train_generator.batch_size
STEP_SIZE_VALID = valid_generator.n//valid_generator.batch_size
history = model.fit_generator(generator=train_generator,
steps_per_epoch=STEP_SIZE_TRAIN,
validation_data=valid_generator,
validation_steps=STEP_SIZE_VALID,
epochs=WARMUP_EPOCHS,
verbose=2)
```
#### Fine-tune the complete model
```
for layer in model.layers:
layer.trainable = True
# lrate = LearningRateScheduler(step_decay)
es = EarlyStopping(monitor='val_loss', mode='min', patience=ES_PATIENCE, restore_best_weights=True, verbose=1)
rlrop = ReduceLROnPlateau(monitor='val_loss', mode='min', patience=RLROP_PATIENCE, factor=DECAY_DROP, min_lr=1e-6, verbose=1)
callbacks = [es, rlrop]
optimizer = optimizers.Adam(lr=LEARNING_RATE)
metrics = ["accuracy"]
model.compile(optimizer=optimizer, loss="binary_crossentropy", metrics=metrics)
model.summary()
STEP_SIZE_TRAIN = train_generator.n//train_generator.batch_size
STEP_SIZE_VALID = valid_generator.n//valid_generator.batch_size
history = model.fit_generator(generator=train_generator,
steps_per_epoch=STEP_SIZE_TRAIN,
validation_data=valid_generator,
validation_steps=STEP_SIZE_VALID,
epochs=EPOCHS,
callbacks=callbacks,
verbose=2)
```
### Complete model graph loss
```
sns.set_style("whitegrid")
fig, (ax1, ax2) = plt.subplots(1, 2, sharex='col', figsize=(20,7))
ax1.plot(history.history['loss'], label='Train loss')
ax1.plot(history.history['val_loss'], label='Validation loss')
ax1.legend(loc='best')
ax1.set_title('Loss')
ax2.plot(history.history['acc'], label='Train Accuracy')
ax2.plot(history.history['val_acc'], label='Validation accuracy')
ax2.legend(loc='best')
ax2.set_title('Accuracy')
plt.xlabel('Epochs')
sns.despine()
plt.show()
```
### Find best threshold value
```
lastFullValPred = np.empty((0, N_CLASSES))
lastFullValLabels = np.empty((0, N_CLASSES))
for i in range(STEP_SIZE_VALID+1):
im, lbl = next(valid_generator)
scores = model.predict(im, batch_size=valid_generator.batch_size)
lastFullValPred = np.append(lastFullValPred, scores, axis=0)
lastFullValLabels = np.append(lastFullValLabels, lbl, axis=0)
print(lastFullValPred.shape, lastFullValLabels.shape)
def find_best_fixed_threshold(preds, targs, do_plot=True):
score = []
thrs = np.arange(0, 0.5, 0.01)
for thr in thrs:
score.append(custom_f2(targs, (preds > thr).astype(int)))
score = np.array(score)
pm = score.argmax()
best_thr, best_score = thrs[pm], score[pm].item()
print(f'thr={best_thr:.3f}', f'F2={best_score:.3f}')
if do_plot:
plt.plot(thrs, score)
plt.vlines(x=best_thr, ymin=score.min(), ymax=score.max())
plt.text(best_thr+0.03, best_score-0.01, f'$F_{2}=${best_score:.3f}', fontsize=14);
plt.show()
return best_thr, best_score
threshold, best_score = find_best_fixed_threshold(lastFullValPred, lastFullValLabels, do_plot=True)
```
### Apply model to test set and output predictions
```
test_generator.reset()
STEP_SIZE_TEST = test_generator.n//test_generator.batch_size
preds = model.predict_generator(test_generator, steps=STEP_SIZE_TEST)
predictions = []
for pred_ar in preds:
valid = []
for idx, pred in enumerate(pred_ar):
if pred > threshold:
valid.append(idx)
if len(valid) == 0:
valid.append(np.argmax(pred_ar))
predictions.append(valid)
filenames = test_generator.filenames
label_map = {valid_generator.class_indices[k] : k for k in valid_generator.class_indices}
results = pd.DataFrame({'id':filenames, 'attribute_ids':predictions})
results['id'] = results['id'].map(lambda x: str(x)[:-4])
results['attribute_ids'] = results['attribute_ids'].apply(lambda x: list(map(label_map.get, x)))
results["attribute_ids"] = results["attribute_ids"].apply(lambda x: ' '.join(x))
results.to_csv('submission.csv',index=False)
results.head(10)
```
| github_jupyter |
```
import os
import numpy as np
import prepare_data
from sklearn import metrics
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import GradientBoostingClassifier
import seaborn as sns
data_path = os.path.join("statistic_features", "stat_features_60s.csv")
patient_list = ['002','003','005','007','08a','08b','09a','09b', '10a','011','013','014','15a','15b','016',
'017','018','019','020','021','022','023','025','026','027','028','029','030','031','032',
'033','034','035','036','037','038','040','042','043','044','045','047','048','049','051']
statistics_list=["std_x", "std_y", "std_z"]
train_patient_list, test_patient_list = train_test_split(patient_list, random_state=152, test_size=0.3)
#prepare_data.save_statistic_features(patient_list, sorce_path="ICHI14_dataset\data",
# save_path=data_path, window_len=60)
X_train, y_train = prepare_data.load_statistic_features(train_patient_list,
data_path=data_path,
statistics_list=statistics_list)
print(X_train.shape)
print(y_train.shape)
X_test, y_test = prepare_data.load_statistic_features(test_patient_list,
data_path=data_path,
statistics_list=statistics_list)
print(X_test.shape)
print(y_test.shape)
sns.countplot(y_train)
sns.countplot(y_test)
```
### 1. Lin. Reg, only one window
```
%%time
model1 = LogisticRegression()
model1.fit(X_train, y_train)
y_predict = model1.predict(X_train)
print("\nTrain set result: ")
print(metrics.classification_report(y_train, y_predict))
print("Confussion matrix: \n", metrics.confusion_matrix(y_train, y_predict))
accuracy = metrics.accuracy_score(y_train, y_predict)
print("\nAccuracy on train set: ", accuracy)
y_predict = model1.predict(X_test)
print("\nTrain set result: ")
print(metrics.classification_report(y_test, y_predict))
print("Confussion matrix: \n", metrics.confusion_matrix(y_test, y_predict))
accuracy = metrics.accuracy_score(y_test, y_predict)
print("\nAccuracy on train set: ", accuracy)
```
### 2. Lin. Reg, several windows features for one window
```
X_train_new, y_train_new = prepare_data.load_stat_features_others_windows(train_patient_list,
data_path=data_path,
n_others_windows=16)
X_test_new, y_test_new = prepare_data.load_stat_features_others_windows(test_patient_list,
data_path=data_path,
n_others_windows=16)
print(X_train_new.shape)
print(y_train_new.shape)
print(X_test_new.shape)
print(y_test_new.shape)
%%time
model3 = LogisticRegression()
model3.fit(X_train_new, y_train_new)
y_predict = model3.predict(X_train_new)
print("\nTrain set result: ")
print(metrics.classification_report(y_train_new, y_predict))
print("Confussion matrix: \n", metrics.confusion_matrix(y_train_new, y_predict))
accuracy = metrics.accuracy_score(y_train_new, y_predict)
print("\nAccuracy on train set: ", accuracy)
y_predict = model3.predict(X_test_new)
print("\nTest set result: ")
print(metrics.classification_report(y_test_new, y_predict))
print("Confussion matrix: \n", metrics.confusion_matrix(y_test_new, y_predict))
accuracy = metrics.accuracy_score(y_test_new, y_predict)
print("\nAccuracy on test set: ", accuracy)
%%time
model4 = GradientBoostingClassifier(n_estimators=30, max_depth=4)
model4.fit(X_train_new, y_train_new)
y_predict = model4.predict(X_train_new)
print("\nTrain set result: ")
print(metrics.classification_report(y_train_new, y_predict))
print("Confussion matrix: \n", metrics.confusion_matrix(y_train_new, y_predict))
accuracy = metrics.accuracy_score(y_train_new, y_predict)
print("\nAccuracy on train set: ", accuracy)
y_predict = model4.predict(X_test_new)
print("\nTest set result: ")
print(metrics.classification_report(y_test_new, y_predict))
print("Confussion matrix: \n", metrics.confusion_matrix(y_test_new, y_predict))
accuracy = metrics.accuracy_score(y_test_new, y_predict)
print("\nAccuracy on test set: ", accuracy)
```
### 3. RNN
```
from keras.preprocessing import sequence
from keras.layers import Dense, Activation, Embedding
from keras.layers import LSTM, Bidirectional, Dropout
from keras.layers import SpatialDropout1D
from keras.models import Sequential
from keras.optimizers import SGD, Adam
from keras.layers.normalization import BatchNormalization
from keras.regularizers import l2
from keras.callbacks import ModelCheckpoint, EarlyStopping
test_patient_list, valid_patient_list = train_test_split(test_patient_list, random_state=151, test_size=0.5)
X_train_new, y_train_new = prepare_data.load_stat_features_others_windows_rnn(train_patient_list,
data_path=data_path,
n_others_windows=16)
X_test_new, y_test_new = prepare_data.load_stat_features_others_windows_rnn(test_patient_list,
data_path=data_path,
n_others_windows=16)
X_valid_new, y_valid_new = prepare_data.load_stat_features_others_windows_rnn(valid_patient_list,
data_path=data_path,
n_others_windows=16)
print(X_train_new.shape)
print(y_train_new.shape)
print(X_valid_new.shape)
print(y_valid_new.shape)
print(X_test_new.shape)
print(y_test_new.shape)
RNN = Sequential()
RNN.add(LSTM(10, dropout=0.1, recurrent_dropout=0.1, input_shape=(17, 3)))
#RNN.add(Bidirectional(layer=LSTM(10, dropout=0.1, recurrent_dropout=0.1) ,input_shape=(17, 3)))
RNN.add(Dense(1, activation="sigmoid", kernel_initializer="glorot_uniform", kernel_regularizer=l2(0.01)))
RNN.compile(loss="binary_crossentropy", optimizer="adam", metrics=["accuracy"])
print(RNN.summary())
callbacks = [ModelCheckpoint('saved_models/RNN2_std_16win_weights.hdf5', monitor='val_acc', save_best_only=True),
EarlyStopping(monitor='val_acc', patience=5)]
%%time
RNN.fit(X_train_new, y_train_new,
batch_size=32,
epochs=15,
validation_data=(X_valid_new, y_valid_new),
callbacks=callbacks,
verbose=1)
scores = RNN.evaluate(X_test_new, y_test_new)
print("Test accuracy =", scores[1])
RNN.save_weights("saved_models/RNN2.2_std_16win_weights.hdf5")
```
### Results of model with best valid acc
```
RNN.load_weights("saved_models/RNN2_std_16win_weights.hdf5")
scores = RNN.evaluate(X_test_new, y_test_new)
print("Test accuracy =", scores[1])
scores = RNN.evaluate(X_valid_new, y_valid_new)
print("Valid accuracy =", scores[1])
scores = RNN.evaluate(X_train_new, y_train_new)
print("Train accuracy =", scores[1])
```
#### Results:
simple rnn(10), EarlyStopping = 5, STD-features:
4 windows: max test acc = 0.7419, <10m
8 windows: max test acc = 0.75089, 0.7473, 0.7652, val_acc: 0.7291, train_acc: 0.7404, ~24 epoch, <15m,
10 windows: max test acc = 0.7471, valid = 0.7548, <10m
12 windows: max test acc = 0.0.7487, valid = 0.7462, ~15 epoch, <10m
16 windows: test accuracy = 0.7567, valid accuracy = 0.7552, time = 7 min - rnn 2.2
Bidirectional (random states for spliting data ~ 152, 151):
8 windows: test accuracy = 0.75, valid accuracy = 0.7466, time = 5min
16 windows: test accuracy = 0.7508, valid accuracy = 0.7633, time = 10 min - rnn 1.2
```
saved_model = RNN.to_json()
with open("saved_models/RNN2_std_16win.json", "w") as json_file:
json_file.write(saved_model)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_01_1_overview.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# T81-558: Applications of Deep Neural Networks
**Module 1: Python Preliminaries**
* Instructor: [Jeff Heaton](https://sites.wustl.edu/jeffheaton/), McKelvey School of Engineering, [Washington University in St. Louis](https://engineering.wustl.edu/Programs/Pages/default.aspx)
* For more information visit the [class website](https://sites.wustl.edu/jeffheaton/t81-558/).
# Module 1 Material
* **Part 1.1: Course Overview** [[Video]](https://www.youtube.com/watch?v=IzZSwS45vt4&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_01_1_overview.ipynb)
* Part 1.2: Introduction to Python [[Video]](https://www.youtube.com/watch?v=czq5d53vKvo&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_01_2_intro_python.ipynb)
* Part 1.3: Python Lists, Dictionaries, Sets and JSON [[Video]](https://www.youtube.com/watch?v=kcGx2I5akSs&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_01_3_python_collections.ipynb)
* Part 1.4: File Handling [[Video]](https://www.youtube.com/watch?v=FSuSLCMgCZc&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_01_4_python_files.ipynb)
* Part 1.5: Functions, Lambdas, and Map/Reduce [[Video]](https://www.youtube.com/watch?v=jQH1ZCSj6Ng&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_01_5_python_functional.ipynb)
Watch one (or more) of these depending on how you want to setup your Python TensorFlow environment:
* [How to Submit a Module Assignment locally](https://www.youtube.com/watch?v=hmCGjCVhYNc)
* [How to Use Google CoLab and Submit Assignment](https://www.youtube.com/watch?v=Pt-Od-oBgOM)
* [Installing TensorFlow, Keras, and Python in Windows CPU or GPU](https://www.youtube.com/watch?v=PnK1jO2kXOQ&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN)
* [Installing TensorFlow, Keras, and Python for an Intel Mac](https://www.youtube.com/watch?v=LnzgQr14p7s&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN)
* [Installing TensorFlow, Keras, and Python for an M1 Mac](https://www.youtube.com/watch?v=_CO-ND1FTOU&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN)
# Google CoLab Instructions
The following code ensures that Google CoLab is running the correct version of TensorFlow.
```
try:
from google.colab import drive
%tensorflow_version 2.x
COLAB = True
print("Note: using Google CoLab")
except:
print("Note: not using Google CoLab")
COLAB = False
```
# Part 1.1: Course Overview
Deep learning is a group of exciting new technologies for neural networks. [[Cite:lecun2015deep]](https://www.nature.com/articles/nature14539) By using a combination of advanced training techniques neural network architectural components, it is now possible to train neural networks of much greater complexity. This course introduces the student to deep belief neural networks, regularization units (ReLU), convolution neural networks, and recurrent neural networks. High-performance computing (HPC) aspects demonstrate how deep learning can be leveraged both on graphical processing units (GPUs), as well as grids. Deep learning allows a model to learn hierarchies of information in a way that is similar to the function of the human brain. The focus is primarily upon the application of deep learning, with some introduction to the mathematical foundations of deep learning. Students make use of the Python programming language to architect a deep learning model for several real-world data sets and interpret the results of these networks. [[Cite:goodfellow2016deep]](https://www.deeplearningbook.org/)
# Assignments
Your grade is calculated according to the following assignments:
Assignment |Weight|Description
--------------------|------|-------
Ice Breaker | 5%|Post a short get to know you discussion topic (individual)
Class Assignments | 50%|10 small programming assignments (5% each, individual)
Kaggle Project | 25%|"Kaggle In-Class" competition (team)
Final Project | 20%|Deep Learning Implementation Report (team)
The 10 class assignments correspond with each of the first 10 modules. Generally, each module assignment is due just before the following module date. Refer to the syllabus for exact due dates. The 10 class assignments are submitted using the Python submission script. Refer to assignment 1 for details.
The Kaggle and Final Projects are completed in teams. The same teams will complete each of these.
* **Module 1 Assignment**: [How to Submit an Assignment](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/assignments/assignment_yourname_class1.ipynb)
* **Module 2 Assignment**: [Creating Columns in Pandas](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/assignments/assignment_yourname_class2.ipynb)
* **Module 3 Assignment**: [Data Preparation in Pandas](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/assignments/assignment_yourname_class3.ipynb)
* **Module 4 Assignment**: [Classification and Regression Neural Networks](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/assignments/assignment_yourname_class4.ipynb)
* **Module 5 Assignment**: [Predict Home Price](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/assignments/assignment_yourname_class5.ipynb)
* **Module 6 Assignment**: [Image Processing](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/assignments/assignment_yourname_class6.ipynb)
* **Module 7 Assignment**: [Computer Vision](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/assignments/assignment_yourname_class7.ipynb)
* **Module 8 Assignment**: [Feature Engineering](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/assignments/assignment_yourname_class8.ipynb)
* **Module 9 Assignment**: [Transfer Learning](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/assignments/assignment_yourname_class9.ipynb)
* **Module 10 Assignment**: [Time Series Neural Network](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/assignments/assignment_yourname_class10.ipynb)
# Your Instructor: Jeff Heaton
Jeff Heaton, pictured in Figure 1.JH is the author of this book and developer of this course. A brief summary of his credentials is given here:
* Master of Information Management (MIM), Washington University in St. Louis, MO
* PhD in Computer Science, Nova Southeastern University in Ft. Lauderdale, FL
* [Vice President and Data Scientist](http://www.rgare.com/knowledge-center/media/articles/rga-where-logic-meets-curiosity), Reinsurance Group of America (RGA)
* Senior Member, IEEE
* jtheaton at domain name of this university
* Other industry certifications: FLMI, ARA, ACS
Social media:
* [Homepage](http://www.heatonresearch.com) - My home page. Includes my research interests and publications.
* [YouTube Channel](https://www.youtube.com/user/HeatonResearch) - My YouTube Channel. Subscribe for my videos on AI and updates to this class.
* [Discord Server](https://discord.gg/3bjthYv) - To discuss this class and AI topics.
* [GitHub](https://github.com/jeffheaton) - My GitHub repositories.
* [LinkedIn](https://www.linkedin.com/in/jeffheaton) - My Linked In profile.
* [Twitter](https://twitter.com/jeffheaton) - My Twitter feed.
* [Google Scholar](https://scholar.google.com/citations?user=1jPGeg4AAAAJ&hl=en) - My citations on Google Scholar.
* [Research Gate](https://www.researchgate.net/profile/Jeff_Heaton) - My profile/research at Research Gate.
* [Orcid](https://orcid.org/0000-0003-1496-4049)
* [Others](http://www.heatonresearch.com/about/) - About me and other social media sites that I am a member of.
**Figure 1.JH: Jeff Heaton Recording a Video**

My PhD dissertation explored new methods for feature engineering in deep learning. [[Cite:heaton2017automated]](https://nsuworks.nova.edu/gscis_etd/994/) I also created the Encog Machine Learning Framework in Java and C#. [[Cite:heaton2015encog]](https://arxiv.org/pdf/1506.04776.pdf) I've also conducted research on artificial life [[Cite:heaton2019evolving]](https://link.springer.com/article/10.1007/s10710-018-9336-1).
# Course Resources
* [Google CoLab](https://colab.research.google.com/) - Free web-based platform that includes Python, Juypter Notebooks, and TensorFlow [[Cite:GoogleTensorFlow]](http://download.tensorflow.org/paper/whitepaper2015.pdf). No setup needed.
* [Python Anaconda](https://www.continuum.io/downloads) - Python distribution that includes many data science packages, such as Numpy, Scipy, Scikit-Learn, Pandas, and much more.
* [Juypter Notebooks](http://jupyter.org/) - Easy to use environment that combines Python, Graphics and Text.
* [TensorFlow](https://www.tensorflow.org/) - Google's mathematics package for deep learning.
* [Kaggle](https://www.kaggle.com/) - Competitive data science. Good source of sample data.
* [Course GitHub Repository](https://github.com/jeffheaton/t81_558_deep_learning) - All of the course notebooks will be published here.
# What is Deep Learning
The focus of this class is deep learning, which is a prevalent type of machine learning that builds upon the original neural networks popularized in the 1980s. There is very little difference between how a deep neural network is calculated compared with the first neural network. We've always been able to create and calculate deep neural networks. A deep neural network is nothing more than a neural network with many layers. While we've always been able to create/calculate deep neural networks, we've lacked an effective means of training them. Deep learning provides an efficient means to train deep neural networks.
## What is Machine Learning
If deep learning is a type of machine learning, this begs the question, "What is machine learning?" Figure 1.ML-DEV illustrates how machine learning differs from traditional software development.
**Figure 1.ML-DEV: ML vs Traditional Software Development**

* **Traditional Software Development** - Programmers create programs that specify how to transform input into the desired output.
* **Machine Learning** - Programmers create models that can learn to produce the desired output for given input. This learning fills the traditional role of the computer program.
Researchers have applied machine learning to many different areas. This class explores three specific domains for the application of deep neural networks, as illustrated in Figure 1.ML-DOM.
**Figure 1.ML-DOM: Application of Machine Learning**

* **Predictive Modeling** - Several named input values allow the neural network to predict another named value that becomes the output. For example, using four measurements of iris flowers to predict the species. This type of data is often called tabular data.
* **Computer Vision** - The use of machine learning to detect patterns in visual data. For example, is an image a picture of a cat or a dog.
* **Time Series** - The use of machine learning to detect patterns in time. Typical applications of time series are financial applications, speech recognition, and even natural language processing (NLP).
### Regression
Regression is when a model, such as a neural network, accepts input, and produces a numeric output. Consider if you must to write a program that predicted how many miles per gallon (MPG) a car could achieve. For the inputs, you would probably want such features as the weight of the car, the horsepower, how large the engine is, and other values. Your program would be a combination of math and if-statements.
Machine learning lets the computer learn the "formula" for calculating the MPG of a car, using data. Consider [this](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/data/auto-mpg.csv) dataset. We can use regression machine learning models to study this data and learn how to predict the MPG for a car.
### Classification
The output of a classification model is what class the input most closely resembles. For example, consider using four measurements of an iris flower to determine the likely species that the flower. The [iris dataset](https://data.heatonresearch.com/data/t81-558/iris.csv) is a typical dataset for machine learning examples.
### Beyond Classification and Regression
One of the most potent aspects of neural networks is that a neural network can be both regression and classification at the same time. The output from a neural network could be any number of the following:
* An image
* A series of numbers that could be interpreted as text, audio, or another time series
* A regression number
* A classification class
## What are Neural Networks
Neural networks are one of the earliest examples of a machine learning model. Neural networks were initially introduced in the 1940s and have risen and fallen several times from popularity. The current generation of deep learning begain in 2006 with an improved training algorithm by Geoffrey Hinton. [[Cite:hinton2006fast]](https://www.mitpressjournals.org/doi/abs/10.1162/neco.2006.18.7.1527) This technique finally allowed neural networks with many layers (deep neural networks) to be efficiently trained. Four researchers have contributed significantly to the development of neural networks. They have consistently pushed neural network research, both through the ups and downs. These four luminaries are shown in Figure 1.LUM.
**Figure 1.LUM: Neural Network Luminaries**

The current luminaries of artificial neural network (ANN) research and ultimately deep learning, in order as appearing in the above picture:
* [Yann LeCun](http://yann.lecun.com/), Facebook and New York University - Optical character recognition and computer vision using convolutional neural networks (CNN). The founding father of convolutional nets.
* [Geoffrey Hinton](http://www.cs.toronto.edu/~hinton/), Google and University of Toronto. Extensive work on neural networks. Creator of deep learning and early adapter/creator of backpropagation for neural networks.
* [Yoshua Bengio](http://www.iro.umontreal.ca/~bengioy/yoshua_en/index.html), University of Montreal. Extensive research into deep learning, neural networks, and machine learning. He has so far remained entirely in academia.
* [Andrew Ng](http://www.andrewng.org/), Badiu and Stanford University. Extensive research into deep learning, neural networks, and application to robotics.
Geoffrey Hinton, Yann LeCun, and Yoshua Bengio won the [Turing Award](https://www.acm.org/media-center/2019/march/turing-award-2018) for their contributions to deep learning.
## Why Deep Learning?
For predictive modeling, neural networks are not that different than other models, such as:
* Support Vector Machines
* Random Forests
* Gradient Boosted Machines
Like these other models, neural networks can perform both **classification** and **regression**. When applied to relatively low-dimensional predictive modeling tasks, deep neural networks do not necessarily add significant accuracy over other model types. Andrew Ng describes the advantage of deep neural networks over traditional model types as illustrated by Figure 1.DL-VS.
**Figure 1.DL-VS: Why Deep Learning?**

Neural networks also have two additional significant advantages over other machine learning models:
* **Convolutional Neural Networks** - Can scan an image for patterns within the image.
* **Recurrent Neural Networks** - Can find patterns across several inputs, not just within a single input.
Neural networks are also very flexible in the types of data that are compatible with the input and output layers. A neural network can take tabular data, images, audio sequences, time series tabular data, and text as its input or output.
# Python for Deep Learning
Python 3.x is the programming language that will be used for this class. Python, as a programming language, has the widest support for deep learning. The most popular frameworks for deep learning in Python are:
* [TensorFlow/Keras](https://www.tensorflow.org/) (Google)
* [PyTorch](https://pytorch.org/) (Facebook)
# Software Installation
This class is technically oriented. A successful student needs to be able to compile and execute Python code that makes use of TensorFlow for deep learning. There are two options for you to accomplish this:
* Install Python, TensorFlow and some IDE (Jupyter, TensorFlow, and others)
* Use Google CoLab in the cloud
Near the top of this document, there are links to videos that describe how to use Google CoLab. There are also videos explaining how to install Python on your local computer. The following sections take you through the process of installing Python on your local computer. This process is essentially the same on Windows, Linux, or Mac. For specific OS instructions, refer to one of the tutorial YouTube videos earlier in this document.
To install Python on your computer complete the following instructions:
* [Installing Python and TensorFlow - Windows/Linux](./install/tensorflow-install-jul-2020.ipynb)
* [Installing Python and TensorFlow - Mac Intel](./install/tensorflow-install-mac-jan-2021.ipynb)
* [Installing Python and TensorFlow - Mac M1](./install/tensorflow-install-mac-metal-jul-2021.ipynb)
# Python Introduction
Some important links for Python and Python packages.
* [Anaconda v3.x](https://www.continuum.io/downloads) Scientific Python Distribution, including: [Scikit-Learn](http://scikit-learn.org/), [Pandas](http://pandas.pydata.org/), and others: csv, json, numpy, scipy
* [Jupyter Notebooks](http://jupyter.readthedocs.io/en/latest/install.html)
* [PyCharm IDE](https://www.jetbrains.com/pycharm/)
* [MatPlotLib](http://matplotlib.org/)
## Jupyter Notebooks
Space matters in Python, indent code to define blocks
Jupyter Notebooks Allow Python and Markdown to coexist.
Even LaTeX math:
$ f'(x) = \lim_{h\to0} \frac{f(x+h) - f(x)}{h}. $
## Python Versions
* If you see `xrange` instead of `range`, you are dealing with Python 2
* If you see `print x` instead of `print(x)`, you are dealing with Python 2
* This class uses Python 3.x!
```
# What version of Python do you have?
import sys
import tensorflow.keras
import pandas as pd
import sklearn as sk
import tensorflow as tf
check_gpu = len(tf.config.list_physical_devices('GPU'))>0
print(f"Tensor Flow Version: {tf.__version__}")
print(f"Keras Version: {tensorflow.keras.__version__}")
print()
print(f"Python {sys.version}")
print(f"Pandas {pd.__version__}")
print(f"Scikit-Learn {sk.__version__}")
print("GPU is", "available" if check_gpu \
else "NOT AVAILABLE")
```
Software used in this class:
* **Python** - The programming language.
* **TensorFlow** - Googles deep learning framework, must have the version specified above.
* **Keras** - [Keras](https://github.com/fchollet/keras) is a high-level neural networks API, written in Python and capable of running on top of TensorFlow, CNTK, or Theano. [[Cite:franccois2017deep]](https://www.manning.com/books/deep-learning-with-python)
* **Pandas** - Allows for data preprocessing. Tutorial [here](http://pandas.pydata.org/pandas-docs/version/0.18.1/tutorials.html)
* **Scikit-Learn** - Machine learning framework for Python. Tutorial [here](http://scikit-learn.org/stable/tutorial/basic/tutorial.html).
# Module 1 Assignment
You can find the first assignment here: [assignment 1](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/assignments/assignment_yourname_class1.ipynb)
| github_jupyter |
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
import tensorflow as tf
print(tf.__version__)
import numpy as np
import matplotlib.pyplot as plt
def plot_series(time, series, format="-", start=0, end=None):
plt.plot(time[start:end], series[start:end], format)
plt.xlabel("Time")
plt.ylabel("Value")
plt.grid(True)
!wget --no-check-certificate \
https://raw.githubusercontent.com/jbrownlee/Datasets/master/daily-min-temperatures.csv \
-O /tmp/daily-min-temperatures.csv
import csv
time_step = []
temps = []
with open('/tmp/daily-min-temperatures.csv') as csvfile:
reader = csv.reader(csvfile, delimiter=',')
next(reader)
step = 0
for row in reader:
temps.append(float(row[1]))
time_step.append(step)
step = step + 1
# YOUR CODE HERE. READ TEMPERATURES INTO TEMPS
# HAVE TIME STEPS BE A SIMPLE ARRAY OF 1, 2, 3, 4 etc
series = np.array(temps)
time = np.array(time_step)
plt.figure(figsize=(10, 6))
plot_series(time, series)
split_time = 2500
time_train = time[ : split_time]
x_train = series[ : split_time]
time_valid = time[split_time : ]
x_valid = series[split_time : ]
window_size = 30
batch_size = 32
shuffle_buffer_size = 1000
def windowed_dataset(series, window_size, batch_size, shuffle_buffer):
return tf.data.Dataset.from_tensor_slices(
tf.expand_dims(
series,
axis=-1)
).window(
window_size,
shift = 1,
drop_remainder = True
).flat_map(
lambda w : w.batch(window_size + 1)
).shuffle(
shuffle_buffer
).map(
lambda w : (w[ : -1], w[1 : ])
).batch(batch_size).prefetch(1)
def model_forecast(model, series, window_size):
return model.predict(
tf.data.Dataset.from_tensor_slices(
series
).window(
window_size,
shift=1,
drop_remainder = True
).flat_map(
lambda w : w.batch(window_size)
).batch(
batch_size
).prefetch(1))
tf.keras.backend.clear_session()
tf.random.set_seed(51)
np.random.seed(51)
window_size = 64
batch_size = 256
train_set = windowed_dataset(x_train, window_size, batch_size, shuffle_buffer_size)
print(train_set)
print(x_train.shape)
model = tf.keras.models.Sequential([
tf.keras.layers.Conv1D(filters=32, kernel_size=5,
strides=1, padding="causal",
activation="relu",
input_shape=[None, 1]),
tf.keras.layers.LSTM(64, return_sequences=True),
tf.keras.layers.LSTM(64, return_sequences=True),
tf.keras.layers.Dense(30, activation="relu"),
tf.keras.layers.Dense(10, activation="relu"),
tf.keras.layers.Dense(1),
tf.keras.layers.Lambda(lambda x: x * 400)
])
lr_schedule = tf.keras.callbacks.LearningRateScheduler(
lambda epoch: 1e-8 * 10**(epoch / 20))
optimizer = tf.keras.optimizers.SGD(lr=1e-8, momentum=0.9)
model.compile(loss=tf.keras.losses.Huber(),
optimizer=optimizer,
metrics=["mae"])
history = model.fit(train_set, epochs=100, callbacks=[lr_schedule])
plt.semilogx(history.history["lr"], history.history["loss"])
plt.axis([1e-8, 1e-4, 0, 60])
tf.keras.backend.clear_session()
tf.random.set_seed(51)
np.random.seed(51)
train_set = windowed_dataset(x_train, window_size=60, batch_size=100, shuffle_buffer=shuffle_buffer_size)
model = tf.keras.models.Sequential([
tf.keras.layers.Conv1D(filters=60, kernel_size=5,
strides=1, padding="causal",
activation="relu",
input_shape=[None, 1]),
tf.keras.layers.LSTM(60, return_sequences=True),
tf.keras.layers.LSTM(60, return_sequences=True),
tf.keras.layers.Dense(30, activation="relu"),
tf.keras.layers.Dense(10, activation="relu"),
tf.keras.layers.Dense(1),
tf.keras.layers.Lambda(lambda x: x * 400)
])
optimizer = tf.keras.optimizers.SGD(lr=1e-5, momentum=0.9)
model.compile(loss=tf.keras.losses.Huber(),
optimizer=optimizer,
metrics=["mae"])
history = model.fit(train_set,epochs=150)
# EXPECTED OUTPUT SHOULD SEE AN MAE OF <2 WITHIN ABOUT 30 EPOCHS
rnn_forecast = model_forecast(model, series[..., np.newaxis], window_size)
rnn_forecast = rnn_forecast[split_time - window_size:-1, -1, 0]
plt.figure(figsize=(10, 6))
plot_series(time_valid, x_valid)
plot_series(time_valid, rnn_forecast)
# EXPECTED OUTPUT. PLOT SHOULD SHOW PROJECTIONS FOLLOWING ORIGINAL DATA CLOSELY
tf.keras.metrics.mean_absolute_error(x_valid, rnn_forecast).numpy()
# EXPECTED OUTPUT MAE < 2 -- I GOT 1.789626
print(rnn_forecast)
# EXPECTED OUTPUT -- ARRAY OF VALUES IN THE LOW TEENS
```
| github_jupyter |
# Rasterio Tutorial
[FOSS4G UK 2019](https://uk.osgeo.org/foss4guk2019/) - [Workshop 'Geoprocessing with Jupyter Notebooks'](https://github.com/samfranklin/foss4guk19-jupyter)
[Sam Franklin](https://github.com/samfranklin) - [SCISYS](https://www.scisys.co.uk/)
## Introduction
Geographic information systems use GeoTIFF and other formats to organize and store gridded raster datasets such as satellite imagery and terrain models. The `Rasterio` package reads and writes these formats and provides a Python API based on Numpy N-dimensional arrays and GeoJSON.
### Objectives:
* Read multiple raster datasets
* Inspect properties of the raster, e.g. get the bounds, cordinate reference system, etc ...
* Use matplotlib library to make plots including a 2D map, contour plot, histogram of values
* Merge multiple ASCIIgrid files to single, compressed GeoTIFF
* Read a vector linestring using the geopandas library
* Create a "profile" or transect of elevation along a route and visualise.
* export the 2D plot to a csv file
#### References:
* [Rasterio Documentation](https://rasterio.readthedocs.io/en/stable/) - Read the docs!
* [Ordnance Survey Open Data - OS Terrain 50](https://www.ordnancesurvey.co.uk/business-and-government/help-and-support/products/terrain-50.html) - for open UK Digital Elevation Model data
* https://overpass-turbo.eu - an interface for extracting data from [OpenStreetMap](https://www.openstreetmap.org/)
#### Acknowledgements:
* Contains OS data © Crown copyright and database right (2019)
* "© OpenStreetMap contributors", further copyright information is available [here](https://www.openstreetmap.org/copyright)
```
import rasterio as rs
from rasterio import merge
import os
import zipfile
from matplotlib import pyplot
from rasterio.plot import show
from rasterio.plot import show_hist
from os import listdir
from os.path import isfile, join
from osgeo import gdal
import importlib
import geopandas
import pandas
import numpy as np
from datetime import datetime
import utilities
from utilities import make_equidistant_points_from_linstring
import matplotlib.pyplot as plt
```
# Part 1. Read multiple DEM files data with rasterio + simple plotting
Note, the data for this part of the tutorial all sits in this repo's `./tutorial/data/dem-raw/` directory
```
# get current directory using a notebook "magic"
!pwd
```
<div class="alert alert-warning">
<strong>NOTE</strong>
<br>This is a good time to talk about Jupyter Notebook magics which is what "!pwd"
<br> Magics are special built-in functions.
<br> A full list of supported magics are in <a href="https://ipython.org/ipython-doc/3/interactive/magics.html">the docs</a>
</div>
Let's start by making some local directories for data if they don't exist already
```
root = os.getcwd()
pth_raw = os.path.join(os.path.join(root,'data'),'dem-raw')
if not os.path.exists(pth_raw):
os.makedirs(pth_raw)
print('raw data dir\t\t=' + pth_raw)
pth_cwd = os.path.join(os.path.join(root,'data'),'dem-working')
if not os.path.exists(pth_cwd):
os.makedirs(pth_cwd)
print('working data dir\t=' + pth_cwd)
```
Next, we need to make a list of all the 'raw' dem, we can do this with a for loop and list comprehension
```
onlyfiles = [f for f in listdir(pth_raw) if isfile(join(pth_raw, f))]
```
Check the list's first entry, you should have `nt00_OST50GRID_20170713.zip`
```
onlyfiles[0]
```
next, extract the all the .asc and .prj files from the .zip file and extract to different directory, this cell does this for you
```
raster_list = []
for ifile in onlyfiles:
input_zip = os.path.join(pth_raw,ifile)
zipobj = zipfile.ZipFile(input_zip)
for i in zipobj.filelist:
if i.filename.endswith('.asc'):
zipobj.extract(i.filename,pth_cwd)
raster_list.append(os.path.join(pth_cwd,i.filename))
print('extracting :: ' + i.filename)
elif i.filename.endswith('.prj'):
zipobj.extract(i.filename,pth_cwd)
print('raster list completed')
```
Check the list's first entry, you should have a path that looks roughly like `./foss4guk19-jupyter/tutorials/data/dem-working/NT00.asc`
```
raster_list[:1]
```
Open the the first raster dataset in the list with rasterio with the `open` method
Note, this doesn't read the actual pixel values of the raster data, we'll do this later
```
src = rs.open(raster_list[0])
```
Now we have opened the raster as a rasterio dataset, we can start to get more information about it, try several of the following attributes:
```python
src.meta # to retrieve metadata, similar to a 'gdalinfo'
src.bounds # to retrieve bounding box info
src.crs # prints the cordinate reference system info held
src.transform # print raster transform from pixel to raster coords
```
```
src.meta
src.bounds
src.crs
src.transform
```
_Enough_ with metadata, let's **READ** the first band of the DEM and plot.
```
# plot the data (no geographic info)
pyplot.imshow(src.read(1), cmap='pink')
pyplot.show()
```
<div class="alert alert-warning">
<strong>NOTE</strong>
<br>The above image is okay, but note the units of the plot. These are pixel units, and not geographic coordinates.
<br> plot.show() is a simple image viewer method.
<br> to plot geographically, use the inbuilt rasterio 'show' function, below
</div>
```
show(src.read(), transform=src.transform, cmap='jet', title='ouch. rainbow palette!');
```
<div class="alert alert-info">
<strong>ACTIVITY</strong>
<br>Right, let's not use the 'jet' colormap palette (see 'cmap=' above).
<br>Go over to Matplotlib's <a href="https://matplotlib.org/tutorials/colors/colormaps.html">cmap documentation</a> (scroll down) and select a different cmap name, then rerun the cell above with your update
</div>
The `rasterio` plot method, is just a wrapper for the highly flexible `matplotlib` library, below we're using a custom function to make a 3-panel plot.
On the centre plot, the `contour=True` option is used
On the right plot, again, using an inbuilt function `show_hist` conveniently plots the histogram of values of the single raster
```
def make_nice_single_raster_plot(input_raster):
fig, (ax1, ax2, ax3) = pyplot.subplots(1,3, figsize=(21,7))
show(input_raster.read(), ax=ax1, transform=input_raster.transform, cmap='BrBG', title='#NoRainbow');
show(input_raster.read(), ax=ax2, transform=input_raster.transform, contour=True, title='contours');
show_hist(input_raster, bins=50, lw=0.0, stacked=False, histtype='stepfilled', title="Histogram of height vals")
make_nice_single_raster_plot(src)
```
# Part 2. Doing some processing, i.e. a merge
Next we're going to merge all rasters in the `raster_list` python list that we made earlier
<div class="alert alert-warning">
<strong>Merging with rasterio</strong>
<br> The merge module copies a list of actual raster data to an output file
<br>Feel free to refer to rasterio merge <a href="https://rasterio.readthedocs.io/en/stable/api/rasterio.merge.html">documentation</a>
</div>
The below cell creats the output raster in memory as the `dest` object
However, `dest` just holds the data, and not the geographic meta
```
raster_obj_list = [rs.open(res) for res in raster_list]
dest, output_trans = merge.merge(raster_obj_list,bounds=None, res=None, nodata=None, precision=7)
```
Next, we need to burn the 'profile' information, i.e. the meta to the output file
First we need to get a copy of one of the input rasters, using ...
```
raster_obj_list[0].profile
```
The final step of the merge, is to write the output file to disk using the `open` method with a `"w"` command
```
# Register GDAL format drivers and configuration options with a
# context manager.
with rs.Env():
# Write an array as a raster band to a new file. For
# the new file's profile, we start with the profile of the source
output_profile = raster_obj_list[0].profile
# And then change the band count to 1, and set LZW compression.
output_profile.update(
{
"driver": "GTiff",
"height": dest.shape[1],
"width": dest.shape[2],
"transform": output_trans,
"count": 1,
"compress": 'lzw'
}
)
with rs.open(os.path.join(pth_cwd,"merge.tif"), "w", **output_profile) as dest1:
dest1.write(dest)
# At the end of the ``with rasterio.Env()`` block, context
# manager exits and all drivers are de-registered.
```
<div class="alert alert-info">
<strong>ACTIVITY</strong>
<br> to check your output directory which should be ./tutorial/data/dem-working
<br>Feel free to open the 'merge.tif' file in a desktop GIS
</div>
For completeness, we will open and read the merge.tif from disk
```
mrg = rs.open(os.path.join(pth_cwd,"merge.tif"))
```
Let's plot it
```
show(mrg.read(), transform=mrg.transform, title='merged');
```
To double check let's view the profile (meta data)
```
mrg.profile
```
The next cell reimport a custom utilities module (you can edit seperate modules but need to reload themn when to avoid a kernel restart)
The module holds a copy of the make single raster 3-panel plot function that we used above
```
importlib.reload(utilities)
from utilities import make_nice_single_raster_plot
# plot with the latest merged raster
make_nice_single_raster_plot(mrg,10)
```
# Part 2. Sampling the Raster
### Sampling two locations
Grabbing coordinates from the lowerleft corner and upper right corner of the merged raster
```
locations = [(mrg.bounds.left + 1, mrg.bounds.bottom + 1),(mrg.bounds.right - 1, mrg.bounds.top - 1)];locations
```
sample the raster with the list of tuple coordinates
```
res = [val.item(0) for val in mrg.sample(locations)];res
```
### Sample the raster with a vector file to create an Elevation Profile
* For this, we will make use of the `geopandas` python package
* First we'll read in the vector linstring from the geojson file.
* Then, create equidistance points along that linestring.
* Then, sample the raster (meaning get the pixel value) for all points in the list
* Then graph the points
```
# run custom function to generate equi-distance points along the linestring
# these points will be used for sampling the raster DEM
path_to_linestring = os.path.join(os.path.join(root,'data'),'cycleway.geojson')
crs = 27700
interval_m = 1000
path_to_sampling_points = os.path.join(pth_cwd,'sampling-points.geojson')
utilities.make_equidistant_points_from_linstring(path_to_linestring,interval_m,crs,path_to_sampling_points)
gdf = geopandas.read_file(path_to_sampling_points)
# print the top five rows of the geo dataframe to review the structure of the data
gdf.head(5)
```
## Note on Formats
The geometry for each record in the dataframe is in the `Well Known Text` format, i.e.
```python
POINT (310978.417847056 700000)
```
However, the rasterio `sample` method requires a list of all XY points used for sampling as a list of python tuples, i.e.
```python
[(x_point1, y_point1), (x_point2, y_point2), (x_point3, y_point3)]
```
Therefore some we need to do some data manipulation to fit to the rasterio arguments.
```
# burn the X and Y values to specific columns in the geodataframe
gdf['x'] = gdf['geometry'].apply(lambda p: p.x)
gdf['y'] = gdf['geometry'].apply(lambda p: p.y)
gdf.head(5)
# create a list of tuples of the X and Y values
geom_as_tuples = list(zip(gdf.x, gdf.y))
geom_as_tuples[:5]
# sample the merged raster with each xy point in the list
list_elevations = []
for val in mrg.sample(geom_as_tuples):
list_elevations.append(val.item(0))
list_elevations[:5]
# OR write the above loop as a python List Comprehension
list_elevations = [val.item(0) for val in mrg.sample(geom_as_tuples)]
list_elevations[:5]
# add the elevations as a column to the sampling geodataframe,
# Why? Because our geodataframe has a bunch of useful methods for plotting
gdf['elev'] = pandas.Series(list_elevations)
gdf.head(5)
# plot the elevation "profile"
gdf['elev'].plot()
# create a slope GeoTIFF using the GDAL DEM Processing tool
dest_dataset = os.path.join(pth_cwd, 'slope.tif')
src_dataset = os.path.join(pth_cwd, 'merge.tif')
try:
gdal.DEMProcessing(dest_dataset,src_dataset,'slope',format='GTiff')
except Exception as e:
print(e)
# open the slope dataset as a rasterio dataset
slope_data = rs.open(os.path.join(pth_cwd,"slope.tif"))
list_slope_vals = [val.item(0) for val in slope_data.sample(geom_as_tuples)]#;list_slope_vals
gdf['slope'] = pandas.Series(list_slope_vals)
fig, ax = plt.subplots()
fig.suptitle('Cycle Route Elevation and Slope Profiles', fontsize=20)
ax1 = gdf['elev'].iloc[1:].plot(ax=ax)
ax2 = gdf['slope'].iloc[1:].plot(ax=ax, secondary_y=True)
ax1.set_xlabel('Distance along line (km)',fontsize=18)
ax1.set_ylabel('Elevation',fontsize=18)
ax2.set_ylabel('Slope',fontsize=18)
ax.legend([ax.get_lines()[0], ax.right_ax.get_lines()[0]], ['Elevation (m)','Slope (Degrees)'])
```
| github_jupyter |
```
import pandas as pd
import numpy as np
train = pd.read_csv( "../data/processed/train_1.csv")
test = pd.read_csv("../data/processed/test_1.csv")
validation = pd.read_csv("../data/processed/validation_1.csv")
from sklearn.model_selection import train_test_split
X = train['review']
y = train['sentiment']
X_train, X_test, y_train, y_test = train_test_split(X,y,test_size=0.2, random_state=42)
print("Creating the bag of words...")
from sklearn.feature_extraction.text import CountVectorizer
# Initialize the "CountVectorizer" object, which is scikit-learn's
# bag of words tool.
vectorizer = CountVectorizer(analyzer = "word",
tokenizer = None,
preprocessor = None,
stop_words = None,
max_features = 5000)
# fit_transform() does two functions: First, it fits the model
# and learns the vocabulary; second, it transforms our training data
# into feature vectors. The input to fit_transform should be a list of
# strings.
%time train_data_features = vectorizer.fit_transform(X_train)
# Numpy arrays are easy to work with, so convert the result to an
# array
train_data_features = train_data_features.toarray()
print("Training the random forest...")
from sklearn.ensemble import RandomForestClassifier
# Initialize a Random Forest classifier with 100 trees
forest = RandomForestClassifier(n_estimators = 100)
# Fit the forest to the training set, using the bag of words as
# features and the sentiment labels as the response variable
#
# This may take a few minutes to run
%time forest = forest.fit(train_data_features, y_train)
test_data_features = vectorizer.transform(X_test)
test_data_features = test_data_features.toarray()
pred = forest.predict(test_data_features)
from sklearn.metrics import accuracy_score, log_loss,confusion_matrix, roc_curve, roc_auc_score, precision_score, recall_score, f1_score
import matplotlib.pyplot as plt
acc = accuracy_score(y_test,pred)
cm = confusion_matrix(y_test,pred)
print("Accuracy Score: " + str(acc))
print("Confusion Matrix: "+ str(cm))
# 1. import
from sklearn.naive_bayes import MultinomialNB
# 2. instantiate a Multinomial Naive Bayes model
nb = MultinomialNB()
%time nb.fit(train_data_features, y_train)
pred = nb.predict(test_data_features)
acc = accuracy_score(y_test, pred)
cm = confusion_matrix(y_test,pred)
print("Accuracy Score: " + str(acc))
print("Confusion Matrix: "+ str(cm))
print("Precision Score: "+ str(precision_score(y_test,pred)))
print("Recall Score: "+ str(recall_score(y_test,pred)))
print("F1 Score: "+ str(f1_score(y_test,pred)))
null_ = []
for i in range(0,len(y_test)):
null_.append(1)
null_accuracy = accuracy_score(y_test, null_)
print('Null accuracy:', null_accuracy)
# 1. import
import lightgbm as lgb
# 2. instantiate a Multinomial Naive Bayes model
lgbm = lgb.LGBMClassifier()
%time lgbm.fit(train_data_features, y_train)
pred = lgbm.predict(test_data_features)
acc = accuracy_score(y_test, pred)
print("Accuracy Score: " + str(acc))
output = pd.DataFrame({"review":X_test, "actual":y_test, "pred":pred})
wrong = output[output['actual'] != output['pred']]
wrong.to_csv("wrong_predictions.csv",index=False,quoting=3)
import xgboost as xgb
xgbo = xgb.XGBClassifier()
%time xgbo.fit(train_data_features, y_train)
predic = xgbo.predict(test_data_features)
acc = accuracy_score(y_test, predic)
print("Accuracy Score: " + str(acc))
from sklearn.linear_model import LogisticRegression
logistic = LogisticRegression()
%time logistic.fit(train_data_features, y_train)
predic = logistic.predict(test_data_features)
acc = accuracy_score(y_test, predic)
cm = confusion_matrix(y_test,predic)
print("Accuracy Score: " + str(acc))
print("Confusion Matrix: "+ str(cm))
print("Precision Score: "+ str(precision_score(y_test,predic)))
print("Recall Score: "+ str(recall_score(y_test,predic)))
print("F1 Score: "+ str(f1_score(y_test,predic)))
X_test = validation['review']
y_test = validation['sentiment']
test_data_features = vectorizer.transform(X_test)
test_data_features = test_data_features.toarray()
pred = nb.predict(test_data_features)
acc = accuracy_score(y_test, pred)
cm = confusion_matrix(y_test,pred)
print("Accuracy Score: " + str(acc))
print("Confusion Matrix: "+ str(cm))
print("Precision Score: "+ str(precision_score(y_test,pred)))
print("Recall Score: "+ str(recall_score(y_test,pred)))
print("F1 Score: "+ str(f1_score(y_test,pred)))
predic = logistic.predict(test_data_features)
acc = accuracy_score(y_test, predic)
cm = confusion_matrix(y_test,predic)
print("Accuracy Score: " + str(acc))
print("Confusion Matrix: "+ str(cm))
print("Precision Score: "+ str(precision_score(y_test,predic)))
print("Recall Score: "+ str(recall_score(y_test,predic)))
print("F1 Score: "+ str(f1_score(y_test,predic)))
tes = ['barang bagus sekali saya suka saya suka',
'penipu saya beli telepon genggam yang sampai di rumah saya malah sabun batang awas ya',
'apa apaan ini sudah sampai lama barang rusak lagi',
'awal saya khawatir karena penjual belum punya reputasi yang bagus, tapi ternyata barang cepat sekali sampai packing tebal dan rapi barang sampai dengan selamat.']
tes_features = vectorizer.transform(tes).toarray()
print(nb.predict(tes_features))
print(logistic.predict(tes_features))
```
| github_jupyter |
```
## This can leave open processes if you don't keep track of them, be sure to clean up after
import numpy as np
import torch
import gym
import pybullet_envs
import os
import time
import sys
from pathlib import Path
sys.path.append(str(Path().resolve().parent))
import utils
import TD3
from numpngw import write_apng
from gym.envs.registration import registry, make, spec
def register(id, *args, **kvargs):
if id in registry.env_specs:
return
else:
return gym.envs.registration.register(id, *args, **kvargs)
register(id='MyAntBulletEnv-v0',
entry_point='override_ant_random:MyAntBulletEnv',
max_episode_steps=2000,
reward_threshold=2500.0)
def eval_policy_render(policy, env_name, seed, eval_episodes=5):
eval_env = gym.make(env_name, render=True)
eval_env.seed(seed + 100)
avg_reward = 0.
for i in range(eval_episodes):
r = np.linalg.norm([20,20])
rand_deg = np.random.randint(0,360) # degrees here for reader clarity, rather than directly in 2pi
rand_x = r*np.cos(np.pi/180 * rand_deg)
rand_y = r*np.sin(np.pi/180 * rand_deg)
eval_env.robot.walk_target_x = rand_x
eval_env.robot.walk_target_y = rand_y
state, done = eval_env.reset(), False
images = [eval_env.render('rgb_array')]
time_step_counter = 1
while not done:
if time_step_counter % 500 == 0:
rand_deg = np.random.randint(0,360) # degrees here for reader clarity, rather than directly in 2pi
rand_x = r*np.cos(np.pi/180 * rand_deg)
rand_y = r*np.sin(np.pi/180 * rand_deg)
eval_env.robot.walk_target_x = rand_x
eval_env.robot.walk_target_y = rand_y
time.sleep(1. / 60.)
action = policy.select_action(np.array(state))
state, reward, done, _ = eval_env.step(action)
avg_reward += reward
images.append(eval_env.render('rgb_array'))
time_step_counter = time_step_counter + 1
print(f'Saving animation: anim_{i}.png, lenght: {len(images)} frames.')
#write_apng(f'anim_{i}.png', images[::2], delay=50) #uncomment this line to save animations
print('Save file complete')
avg_reward /= eval_episodes
return avg_reward
def load_policy(env_name_var):
args = {
"policy" : "TD3", # Policy name (TD3, DDPG or OurDDPG)
"env" : env_name_var, # OpenAI gym environment name
"seed" : 0, # Sets Gym, PyTorch and Numpy seeds
"start_timesteps" : 25e3, # Time steps initial random policy is used
"eval_freq" : 5e3, # How often (time steps) we evaluate
"max_timesteps" : 2e6, # Max time steps to run environment
"expl_noise" : 0.1, # Std of Gaussian exploration noise
"batch_size" : 256, # Batch size for both actor and critic
"discount" : 0.99, # Discount factor
"tau" : 0.007, # Target network update rate
"policy_noise" : 0.2, # Noise added to target policy during critic update
"noise_clip" : 0.5, # Range to clip target policy noise
"policy_freq" : 2, # Frequency of delayed policy updates
"save_model" : "store_true", # Save model and optimizer parameters
"load_model" : "default", # Model load file name, "" doesn't load, "default" uses file_name
}
file_name = f"{args['policy']}_{args['env']}_{args['seed']}_{args['tau']}"
print("---------------------------------------")
print(f"Policy: {args['policy']}, Env: {args['env']}, Seed: {args['seed']}")
print("---------------------------------------")
if not os.path.exists("./results"):
os.makedirs("./results")
if args['save_model'] and not os.path.exists("./models"):
os.makedirs("./models")
env = gym.make(args['env'])
# Set seeds
env.seed(args['seed'])
env.action_space.seed(args['seed'])
torch.manual_seed(args['seed'])
np.random.seed(args['seed'])
state_dim = env.observation_space.shape[0]
action_dim = env.action_space.shape[0]
max_action = float(env.action_space.high[0])
kwargs = {
"state_dim": state_dim,
"action_dim": action_dim,
"max_action": max_action,
"discount": args['discount'],
"tau": args['tau'],
}
# Initialize policy
if args['policy'] == "TD3":
# Target policy smoothing is scaled wrt the action scale
kwargs["policy_noise"] = args['policy_noise'] * max_action
kwargs["noise_clip"] = args['noise_clip'] * max_action
kwargs["policy_freq"] = args['policy_freq']
policy = TD3.TD3(**kwargs)
if args['load_model'] != "":
policy_file = file_name if args['load_model'] == "default" else args['load_model']
policy.load(f"./models/{policy_file}")
return policy
policy = load_policy("MyAntBulletEnv-v0")
eval_policy_render(policy, "MyAntBulletEnv-v0", 0)
```
| github_jupyter |
# FloPy
### MT3D-USGS Example
Demonstrates functionality of the flopy MT3D-USGS module using the 'Crank-Nicolson' example distributed with MT3D-USGS.
#### Problem description:
* Grid dimensions: 1 Layer, 3 Rows, 650 Columns
* Stress periods: 3
* Units are in seconds and meters
* Flow package: UPW
* Stress packages: SFR, GHB
* Solvers: NWT, GCG
```
import sys
import os
import platform
import string
from io import StringIO, BytesIO
import math
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
# run installed version of flopy or add local path
try:
import flopy
except:
fpth = os.path.abspath(os.path.join('..', '..'))
sys.path.append(fpth)
import flopy
print(sys.version)
print('numpy version: {}'.format(np.__version__))
print('matplotlib version: {}'.format(mpl.__version__))
print('flopy version: {}'.format(flopy.__version__))
modelpth = os.path.join('data')
modelname = 'CrnkNic'
mfexe = 'mfnwt'
mtexe = 'mt3dusgs'
if platform.system() == 'Windows':
mfexe += '.exe'
mtexe += '.exe'
# Make sure modelpth directory exists
if not os.path.exists(modelpth):
os.mkdir(modelpth)
# Instantiate MODFLOW object in flopy
mf = flopy.modflow.Modflow(modelname=modelname, exe_name=mfexe, model_ws=modelpth, version='mfnwt')
```
Set up model discretization
```
Lx = 650.0
Ly = 15
nrow = 3
ncol = 650
nlay = 1
delr = Lx / ncol
delc = Ly / nrow
xmax = ncol * delr
ymax = nrow * delc
X, Y = np.meshgrid(np.linspace(delr / 2, xmax - delr / 2, ncol),
np.linspace(ymax - delc / 2, 0 + delc / 2, nrow))
```
Instantiate output control (oc) package for MODFLOW-NWT
```
# Output Control: Create a flopy output control object
oc = flopy.modflow.ModflowOc(mf)
```
Instantiate solver package for MODFLOW-NWT
```
# Newton-Raphson Solver: Create a flopy nwt package object
headtol = 1.0E-4
fluxtol = 5
maxiterout = 5000
thickfact = 1E-06
linmeth = 2
iprnwt = 1
ibotav = 1
nwt = flopy.modflow.ModflowNwt(mf, headtol=headtol, fluxtol=fluxtol, maxiterout=maxiterout,
thickfact=thickfact, linmeth=linmeth, iprnwt=iprnwt, ibotav=ibotav,
options='SIMPLE')
```
Instantiate discretization (DIS) package for MODFLOW-NWT
```
# The equations for calculating the ground elevation in the 1 Layer CrnkNic model.
# Although Y isn't used, keeping it here for symetry
def topElev(X, Y):
return 100. - (np.ceil(X)-1) * 0.03
grndElev = topElev(X, Y)
bedRockElev = grndElev - 3.
Steady = [False, False, False]
nstp = [1, 1, 1]
tsmult = [1., 1., 1.]
# Stress periods extend from (12AM-8:29:59AM); (8:30AM-11:30:59AM); (11:31AM-23:59:59PM)
perlen = [30600, 10800, 45000]
# Create the discretization object
# itmuni = 1 (seconds); lenuni = 2 (meters)
dis = flopy.modflow.ModflowDis(mf, nlay, nrow, ncol, nper=3, delr=delr, delc=delc,
top=grndElev, botm=bedRockElev, laycbd=0, itmuni=1, lenuni=2,
steady=Steady, nstp=nstp, tsmult=tsmult, perlen=perlen)
```
Instantiate upstream weighting (UPW) flow package for MODFLOW-NWT
```
# UPW parameters
# UPW must be instantiated after DIS. Otherwise, during the mf.write_input() procedures,
# flopy will crash.
laytyp = 1
layavg = 2
chani = 1.0
layvka = 1
iphdry = 0
hk = 0.1
hani = 1
vka = 1.
ss = 0.000001
sy = 0.20
hdry = -888
upw = flopy.modflow.ModflowUpw(mf, laytyp=laytyp, layavg=layavg, chani=chani, layvka=layvka,
ipakcb=53, hdry=hdry, iphdry=iphdry, hk=hk, hani=hani,
vka=vka, ss=ss, sy=sy)
```
Instantiate basic (BAS or BA6) package for MODFLOW-NWT
```
# Create a flopy basic package object
def calc_strtElev(X, Y):
return 99.5 - (np.ceil(X)-1) * 0.0001
ibound = np.ones((nlay, nrow, ncol))
ibound[:,0,:] *= -1
ibound[:,2,:] *= -1
strtElev = calc_strtElev(X, Y)
bas = flopy.modflow.ModflowBas(mf, ibound=ibound, hnoflo=hdry, strt=strtElev)
```
Instantiate streamflow routing (SFR2) package for MODFLOW-NWT
```
# Streamflow Routing Package: Try and set up with minimal options in use
# 9 11 IFACE # Data Set 1: ISTCB1 ISTCB2
nstrm = ncol
nss = 6
const = 1.0
dleak = 0.0001
istcb1 = -10
istcb2 = 11
isfropt = 1
segment_data = None
channel_geometry_data = None
channel_flow_data = None
dataset_5 = None
reachinput = True
# The next couple of lines set up the reach_data for the 30x100 hypothetical model.
# Will need to adjust the row based on which grid discretization we're doing.
# Ensure that the stream goes down one of the middle rows of the model.
strmBed_Elev = 98.75 - (np.ceil(X[1,:])-1) * 0.0001
s1 = 'k,i,j,iseg,ireach,rchlen,strtop,slope,strthick,strhc1\n'
iseg = 0
irch = 0
for y in range(ncol):
if y <= 37:
if iseg == 0:
irch = 1
else:
irch += 1
iseg = 1
strhc1 = 1.0e-10
elif y <= 104:
if iseg == 1:
irch = 1
else:
irch += 1
iseg = 2
strhc1 = 1.0e-10
elif y <= 280:
if iseg == 2:
irch = 1
else:
irch += 1
iseg = 3
strhc1 = 2.946219199e-6
elif y <= 432:
if iseg == 3:
irch = 1
else:
irch += 1
iseg = 4
strhc1 = 1.375079882e-6
elif y <= 618:
if iseg == 4:
irch = 1
else:
irch += 1
iseg = 5
strhc1 = 1.764700062e-6
else:
if iseg == 5:
irch = 1
else:
irch += 1
iseg = 6
strhc1 = 1e-10
# remember that lay, row, col need to be zero-based and are adjusted accordingly by flopy
# layer + row + col + iseg + irch + rchlen + strtop + slope + strthick + strmbed K
s1 += '0,{}'.format(1)
s1 += ',{}'.format(y)
s1 += ',{}'.format(iseg)
s1 += ',{}'.format(irch)
s1 += ',{}'.format(delr)
s1 += ',{}'.format(strmBed_Elev[y])
s1 += ',{}'.format(0.0001)
s1 += ',{}'.format(0.50)
s1 += ',{}\n'.format(strhc1)
if not os.path.exists('data'):
os.mkdir('data')
fpth = os.path.join('data', 's1.csv')
f = open(fpth, 'w')
f.write(s1)
f.close()
dtype = [('k', '<i4'), ('i', '<i4'), ('j', '<i4'), ('iseg', '<i4'),
('ireach', '<f8'), ('rchlen', '<f8'), ('strtop', '<f8'),
('slope', '<f8'), ('strthick', '<f8'), ('strhc1', '<f8')]
if (sys.version_info > (3, 0)):
f = open(fpth, 'rb')
else:
f = open(fpth, 'r')
reach_data = np.genfromtxt(f, delimiter=',', names=True, dtype=dtype)
f.close()
s2 = "nseg,icalc,outseg,iupseg,nstrpts, flow,runoff,etsw,pptsw, roughch, roughbk,cdpth,fdpth,awdth,bwdth,width1,width2\n \
1, 1, 2, 0, 0, 0.0125, 0.0, 0.0, 0.0, 0.082078856000, 0.082078856000, 0.0, 0.0, 0.0, 0.0, 1.5, 1.5\n \
2, 1, 3, 0, 0, 0.0, 0.0, 0.0, 0.0, 0.143806300000, 0.143806300000, 0.0, 0.0, 0.0, 0.0, 1.5, 1.5\n \
3, 1, 4, 0, 0, 0.0, 0.0, 0.0, 0.0, 0.104569661821, 0.104569661821, 0.0, 0.0, 0.0, 0.0, 1.5, 1.5\n \
4, 1, 5, 0, 0, 0.0, 0.0, 0.0, 0.0, 0.126990045841, 0.126990045841, 0.0, 0.0, 0.0, 0.0, 1.5, 1.5\n \
5, 1, 6, 0, 0, 0.0, 0.0, 0.0, 0.0, 0.183322283828, 0.183322283828, 0.0, 0.0, 0.0, 0.0, 1.5, 1.5\n \
6, 1, 0, 0, 0, 0.0, 0.0, 0.0, 0.0, 0.183322283828, 0.183322283828, 0.0, 0.0, 0.0, 0.0, 1.5, 1.5"
fpth = os.path.join('data', 's2.csv')
f = open(fpth, 'w')
f.write(s2)
f.close()
if (sys.version_info > (3, 0)):
f = open(fpth, 'rb')
else:
f = open(fpth, 'r')
segment_data = np.genfromtxt(f, delimiter=',',names=True)
f.close()
# Be sure to convert segment_data to a dictionary keyed on stress period.
segment_data = np.atleast_1d(segment_data)
segment_data = {0: segment_data,
1: segment_data,
2: segment_data}
# There are 3 stress periods
dataset_5 = {0: [nss, 0, 0],
1: [nss, 0, 0],
2: [nss, 0, 0]}
sfr = flopy.modflow.ModflowSfr2(mf, nstrm=nstrm, nss=nss, const=const, dleak=dleak, isfropt=isfropt, istcb2=0,
reachinput=True, reach_data=reach_data, dataset_5=dataset_5,
segment_data=segment_data, channel_geometry_data=channel_geometry_data)
```
Instantiate gage package for use with MODFLOW-NWT package
```
gages = [[1,38,61,1],[2,67,62,1], [3,176,63,1], [4,152,64,1], [5,186,65,1], [6,31,66,1]]
files = ['CrnkNic.gage','CrnkNic.gag1','CrnkNic.gag2','CrnkNic.gag3','CrnkNic.gag4','CrnkNic.gag5',
'CrnkNic.gag6']
gage = flopy.modflow.ModflowGage(mf, numgage=6, gage_data=gages, filenames = files)
```
Instantiate linkage with mass transport routing (LMT) package for MODFLOW-NWT (generates linker file)
```
lmt = flopy.modflow.ModflowLmt(mf, output_file_name='CrnkNic.ftl', output_file_header='extended',
output_file_format='formatted', package_flows = ['sfr'])
```
Write the MODFLOW input files
```
pth = os.getcwd()
print(pth)
mf.write_input()
# run the model
mf.run_model()
```
Now draft up MT3D-USGS input files.
```
# Instantiate MT3D-USGS object in flopy
mt = flopy.mt3d.Mt3dms(modflowmodel=mf, modelname=modelname, model_ws=modelpth,
version='mt3d-usgs', namefile_ext='mtnam', exe_name=mtexe,
ftlfilename='CrnkNic.ftl', ftlfree=True)
```
Instantiate basic transport (BTN) package for MT3D-USGS
```
btn = flopy.mt3d.Mt3dBtn(mt, sconc=3.7, ncomp=1, prsity=0.2, cinact=-1.0,
thkmin=0.001, nprs=-1, nprobs=10, chkmas=True,
nprmas=10, dt0=180, mxstrn=2500)
```
Instantiate advection (ADV) package for MT3D-USGS
```
adv = flopy.mt3d.Mt3dAdv(mt, mixelm=0, percel=1.00, mxpart=5000, nadvfd=1)
```
Instatiate generalized conjugate gradient solver (GCG) package for MT3D-USGS
```
rct = flopy.mt3d.Mt3dRct(mt,isothm=0,ireact=100,igetsc=0,rc1=0.01)
gcg = flopy.mt3d.Mt3dGcg(mt, mxiter=10, iter1=50, isolve=3, ncrs=0,
accl=1, cclose=1e-6, iprgcg=1)
```
Instantiate source-sink mixing (SSM) package for MT3D-USGS
```
# For SSM, need to set the constant head boundary conditions to the ambient concentration
# for all 1,300 constant head boundary cells.
itype = flopy.mt3d.Mt3dSsm.itype_dict()
ssm_data = {}
ssm_data[0] = [(0, 0, 0, 3.7, itype['CHD'])]
ssm_data[0].append((0, 2, 0, 3.7, itype['CHD']))
for i in [0,2]:
for j in range(1, ncol):
ssm_data[0].append((0, i, j, 3.7, itype['CHD']))
ssm = flopy.mt3d.Mt3dSsm(mt, stress_period_data=ssm_data)
```
Instantiate streamflow transport (SFT) package for MT3D-USGS
```
dispsf = []
for y in range(ncol):
if y <= 37:
dispsf.append(0.12)
elif y <= 104:
dispsf.append(0.15)
elif y <= 280:
dispsf.append(0.24)
elif y <= 432:
dispsf.append(0.31)
elif y <= 618:
dispsf.append(0.40)
else:
dispsf.append(0.40)
# Enter a list of the observation points
# Each observation is taken as the last reach within the first 5 segments
seg_len = np.unique(reach_data['iseg'], return_counts=True)
obs_sf = np.cumsum(seg_len[1])
obs_sf = obs_sf.tolist()
# The last reach is not an observation point, therefore drop
obs_sf.pop(-1)
# In the first and last stress periods, concentration at the headwater is 3.7
sf_stress_period_data = {0: [0, 0, 3.7],
1: [0, 0, 11.4],
2: [0, 0, 3.7]}
gage_output = [None, None, 'CrnkNic.sftobs']
sft = flopy.mt3d.Mt3dSft(mt, nsfinit=650, mxsfbc=650, icbcsf=81, ioutobs=82,
isfsolv=1, cclosesf=1.0E-6, mxitersf=10, crntsf=1.0, iprtxmd=0,
coldsf=3.7, dispsf=dispsf, nobssf=5, obs_sf=obs_sf,
sf_stress_period_data = sf_stress_period_data,
filenames=gage_output)
sft.dispsf[0].format.fortran = "(10E15.6)"
```
Write the MT3D-USGS input files
```
mt.write_input()
# run the model
mt.run_model()
```
# Compare mt3d-usgs results to an analytical solution
```
# Define a function to read SFT output file
def load_ts_from_SFT_output(fname, nd=1):
f=open(fname, 'r')
iline=0
lst = []
for line in f:
if line.strip().split()[0].replace(".", "", 1).isdigit():
l = line.strip().split()
t = float(l[0])
loc = int(l[1])
conc = float(l[2])
if(loc == nd):
lst.append( [t,conc] )
ts = np.array(lst)
f.close()
return ts
# Also define a function to read OTIS output file
def load_ts_from_otis(fname, iobs=1):
f = open(fname,'r')
iline = 0
lst = []
for line in f:
l = line.strip().split()
t = float(l[0])
val = float(l[iobs])
lst.append( [t, val] )
ts = np.array(lst)
f.close()
return ts
```
Load output from SFT as well as from the OTIS solution
```
# Model output
fname_SFTout = os.path.join('data', 'CrnkNic.sftcobs.out')
# Loading MT3D-USGS output
ts1_mt3d = load_ts_from_SFT_output(fname_SFTout, nd=38)
ts2_mt3d = load_ts_from_SFT_output(fname_SFTout, nd=105)
ts3_mt3d = load_ts_from_SFT_output(fname_SFTout, nd=281)
ts4_mt3d = load_ts_from_SFT_output(fname_SFTout, nd=433)
ts5_mt3d = load_ts_from_SFT_output(fname_SFTout, nd=619)
# OTIS results located here
fname_OTIS = os.path.join('..', 'data', 'mt3d_test', 'mfnwt_mt3dusgs', 'sft_crnkNic', 'OTIS_solution.out')
# Loading OTIS output
ts1_Otis = load_ts_from_otis(fname_OTIS, 1)
ts2_Otis = load_ts_from_otis(fname_OTIS, 2)
ts3_Otis = load_ts_from_otis(fname_OTIS, 3)
ts4_Otis = load_ts_from_otis(fname_OTIS, 4)
ts5_Otis = load_ts_from_otis(fname_OTIS, 5)
```
Set up some plotting functions
```
def set_plot_params():
import matplotlib as mpl
from matplotlib.font_manager import FontProperties
mpl.rcParams['font.sans-serif'] = 'Arial'
mpl.rcParams['font.serif'] = 'Times'
mpl.rcParams['font.cursive'] = 'Zapf Chancery'
mpl.rcParams['font.fantasy'] = 'Comic Sans MS'
mpl.rcParams['font.monospace'] = 'Courier New'
mpl.rcParams['pdf.compression'] = 0
mpl.rcParams['pdf.fonttype'] = 42
ticksize = 10
mpl.rcParams['legend.fontsize'] = 7
mpl.rcParams['axes.labelsize'] = 12
mpl.rcParams['xtick.labelsize'] = ticksize
mpl.rcParams['ytick.labelsize'] = ticksize
return
def set_sizexaxis(a,fmt,sz):
success = 0
x = a.get_xticks()
# print x
xc = np.chararray(len(x), itemsize=16)
for i in range(0,len(x)):
text = fmt % ( x[i] )
xc[i] = string.strip(string.ljust(text,16))
# print xc
a.set_xticklabels(xc, size=sz)
success = 1
return success
def set_sizeyaxis(a,fmt,sz):
success = 0
y = a.get_yticks()
# print y
yc = np.chararray(len(y), itemsize=16)
for i in range(0,len(y)):
text = fmt % ( y[i] )
yc[i] = string.strip(string.ljust(text,16))
# print yc
a.set_yticklabels(yc, size=sz)
success = 1
return success
```
Compare output:
```
#set up figure
try:
plt.close('all')
except:
pass
set_plot_params()
fig = plt.figure(figsize=(6, 4), facecolor='w')
ax = fig.add_subplot(1, 1, 1)
ax.plot(ts1_Otis[:,0], ts1_Otis[:,1], 'k-', linewidth=1.0)
ax.plot(ts2_Otis[:,0], ts2_Otis[:,1], 'b-', linewidth=1.0)
ax.plot(ts3_Otis[:,0], ts3_Otis[:,1], 'r-', linewidth=1.0)
ax.plot(ts4_Otis[:,0], ts4_Otis[:,1], 'g-', linewidth=1.0)
ax.plot(ts5_Otis[:,0], ts5_Otis[:,1], 'c-', linewidth=1.0)
ax.plot((ts1_mt3d[:,0])/3600, ts1_mt3d[:,1], 'kD', markersize=2.0, mfc='none',mec='k')
ax.plot((ts2_mt3d[:,0])/3600, ts2_mt3d[:,1], 'b*', markersize=3.0, mfc='none',mec='b')
ax.plot((ts3_mt3d[:,0])/3600, ts3_mt3d[:,1], 'r+', markersize=3.0)
ax.plot((ts4_mt3d[:,0])/3600, ts4_mt3d[:,1], 'g^', markersize=2.0, mfc='none',mec='g')
ax.plot((ts5_mt3d[:,0])/3600, ts5_mt3d[:,1], 'co', markersize=2.0, mfc='none',mec='c')
#customize plot
ax.set_xlabel('Time, hours')
ax.set_ylabel('Concentration, mg L-1')
ax.set_ylim([3.5,13])
ticksize = 10
#legend
leg = ax.legend(
(
'Otis, Site 1', 'Otis, Site 2', 'Otis, Site 3', 'Otis, Site 4', 'Otis, Site 5',
'MT3D-USGS, Site 1', 'MT3D-USGS, Site 2', 'MT3D-USGS, Site 3', 'MT3D-USGS, Site 4', 'MT3D-USGS, Site 5',
),
loc='upper right', labelspacing=0.25, columnspacing=1,
handletextpad=0.5, handlelength=2.0, numpoints=1)
leg._drawFrame = False
plt.show()
```
| github_jupyter |
# Self Organising Map Challenge
## The Kohonen Network
The Kohonen Self Organizing Map (SOM) provides a data visualization technique which helps to understand high dimensional data by reducing the dimensions of data to a map. SOM also represents clustering concept by grouping similar data together.
Unlike other learning technique in neural networks, training a SOM requires no target vector. A SOM learns to classify the training data without any external supervision.

### Structure
A network has a width and a height that descibes the grid of nodes. For example, the grid may be 4x4, and so there would be 16 nodes.
Each node has a weight for each value in the input vector. A weight is simply a float value that the node multiplies the input value by to determine how influential it is (see below)
Each node has a set of weights that match the size of the input vector. For example, if the input vector has 10 elements, each node would have 10 weights.
### Training
To train the network
1. Each node's weights are initialized.
2. We enumerate through the training data for some number of iterations (repeating if necessary). The current value we are training against will be referred to as the `current input vector`
3. Every node is examined to calculate which one's weights are most like the input vector. The winning node is commonly known as the Best Matching Unit (BMU).
4. The radius of the neighbourhood of the BMU is now calculated. This is a value that starts large, typically set to the 'radius' of the lattice, but diminishes each time-step. Any nodes found within this radius are deemed to be inside the BMU's neighbourhood.
5. Each neighbouring node's (the nodes found in step 4) weights are adjusted to make them more like the input vector. The closer a node is to the BMU, the more its weights get altered.
6. Go to step 2 until we've completed N iterations.
### Calculating the Best Matching Unit (BMU)
To determine the best matching unit, one method is to iterate through all the nodes and calculate the Euclidean distance between each node's weight vector and the current input vector. The node with a weight vector closest to the input vector is tagged as the BMU.
The Euclidean distance $\mathsf{distance}_{i}$ (from the input vector $V$ to the $i$th node's weights $W_i$)is given as (using Pythagoras):
$$ \mathsf{distance}_{i}=\sqrt{\sum_{k=0}^{k=n}(V_k - W_{i_k})^2}$$
where V is the current input vector and $W_i$ is the node's weight vector. $n$ is the size of the input & weight vector.
*Note*: $V$ and $W$ are vectors. $V$ is the input vector, and $W_i$ is the weight vector of the $i$th node. $V_k$ and $W_{i_k}$ represent the $k$'th value within those vectors.
The BMU is the node with the minimal distance for the current input vector
### Calculating the Neighbourhood Radius
The next step is to calculate which of the other nodes are within the BMU's neighbourhood. All these nodes will have their weight vectors altered.
First we calculate what the radius of the neighbourhood should be and then use Pythagoras to determine if each node is within the radial distance or not.
A unique feature of the Kohonen learning algorithm is that the area of the neighbourhood shrinks over time. To do this we use the exponential decay function:
Given a desired number of training iterations $n$:
$$n_{\mathsf{max iterations}} = 100$$
Calculate the radius $\sigma_t$ at iteration number $t$:
$$\sigma_t = \sigma_0 \exp\left(- \frac{t}{\lambda} \right) \qquad t = 1,2,3,4... $$
Where $\sigma_0$ denotes the neighbourhood radius at iteration $t=0$, $t$ is the current iteration. We define $\sigma_0$ (the initial radius) and $\lambda$ (the time constant) as below:
$$\sigma_0 = \frac{\max(width,height)}{2} \qquad \lambda = \frac{n_{\mathsf{max iterations}}}{\log(\sigma_0)} $$
Where $width$ & $height$ are the width and height of the grid.
### Calculating the Learning Rate
We define the initial leanring rate $\alpha_0$ at iteration $t = 0$ as:
$$\alpha_0 = 0.1$$
So, we can calculate the learning rate at a given iteration t as:
$$\alpha_t = \alpha_0 \exp \left(- \frac{t}{\lambda} \right) $$
where $t$ is the iteration number, $\lambda$ is the time constant (calculated above)
### Calculating the Influence
As well as the learning rate, we need to calculate the influence $\theta_t$ of the learning/training at a given iteration $t$.
So for each node, we need to caclulate the euclidean distance $d_i$ from the BMU to that node. Similar to when we calculate the distance to find the BMU, we use Pythagoras. The current ($i$th) node's x position is given by $x(W_i)$, and the BMU's x position is, likewise, given by $x(Z)$. Similarly, $y()$ returns the y position of a node.
$$ d_{i}=\sqrt{(x(W_i) - x(Z))^2 + (y(W_i) - y(Z))^2} $$
Then, the influence decays over time according to:
$$\theta_t = \exp \left( - \frac{d_{i}^2}{2\sigma_t^2} \right) $$
Where $\sigma_t$ is the neighbourhood radius at iteration $t$ as calculated above.
Note: You will need to come up with an approach to x() and y().
### Updating the Weights
To update the weights of a given node, we use:
$$W_{i_{t+1}} = W_{i_t} + \alpha_t \theta_t (V_t - W_{i_t})$$
So $W_{i_{t+1}}$ is the new value of the weight for the $i$th node, $V_t$ is the current value of the training data, $W_{i_t}$ is the current weight and $\alpha_t$ and $\theta_t$ are the learning rate and influence calculated above.
*Note*: the $W$ and $V$ are vectors
## Challenge
In this challenge, you need to implement a SOM such that it can categorise a set of random 3-dimensional colours. The output should be an image of the organised data.
### Expectations and Criteria
- You are expected to detail your decisions & thinking in comments
- You may use any language you like (although the examples given below are using Python)
- Avoid using too many libraries that would do too much of the work for you. For example, in Python a good approach would be to use only functions from `numpy` (together with in-built libs if needed)
- Try and make your code as efficient as possible. Feel free to deviate from a literal interpretation of the instructions given under the **Training** heading above in the interest of code efficiency
- Consider the best way to structure your code and code base for later use by other developers or in anticipation of productionisation (you don't need to use Jupyter Notebook if it doesn't fit)
- Include any tests you may have used (tests not required however)
- Package your code into a docker image with a `Dockerfile`
- Discuss your approach and process to productionise this application
- You can use code below to generate the trainig data of 10 colours:
---
---
## 1. Response
#### Project Packaging & Structure
<div style='color:#FF4500;'>
- ```numpy``` is used as primary library to hold and manipluate data.
- ```fastAPI``` is used to package the application and expose ```/train/```, ```/atrain/```, ```/list-of-models/```, ```/download/```, ```/predict/``` endpoints. ```fastAPI``` was chosen due to its high performance and auto tuned for number of CPU cores for handling high request load.
- The solution is packaged as a production ready server application & containerised. The solution files reside under ```app/``` folder:
- `kohonen.py` : <i>file contains ```Kohonen``` that implements the algorithm.</i>
- `main.py` : <i>file act as entry point for ```fastAPI```, where all REST API requests will fall.</i>
- `settings.py` : <i>file contain settings, contant values etc used in the project.</i>
- `utils/utils.py` : <i>file contains helper methods that are used in the project.</i>
- `saved_models/` : <i>folder which (can) contain saved models (weights) post training. Ideally, models/weights could be saved on a blob storage for better scalablity.</i>
- <i>There are already saved grids/model files (.npy) under following names/configurations:</i>
- ```10X10 100N```
- ```10X10 200N```
- ```10X10 500N```
- ```100X100 1000N```
- `exceptions.py` : <i>file implementing kohonen algorithm exceptions.</i>
- `logs/logging.log` : <i>file contains logs that are captured throughout the appplication.</i>
- `api_params` : <i> file contains multiple classes that are responsible for parsing ```fastAPI``` requests (body parameters).</i>
- `app/test.py` : <i> Please look for <a href='app/test.py'>test.py</a> file for triggering tests. Uncomment lines, to run via command line ```python test.py```</i>
- `saved_plots/` : <i>folder to save any plot images (used in test.py) for analysis/code-testing purposes</i>
- `saved_train_inputs/` : <i>folder to save any training data (used in test.py) as .npy file for analysis/code-testing purposes</i>
- `configs/` : <i>folder contains python package requiements files.</i>
- ```prod.requirements.text``` is used to install requirements when packaging, containerising.
- ```requirements.text``` has to be used to create local envrionment for running, testing the application via command line, jupyter lab etc.
- (Core Libs)
- ```! pip install numpy==1.20.1```
- ```! pip install matplotlib==3.3.4```
- (Packaging/Productioning Libs)
- ```! pip install fastapi==0.63.0```
- ```! pip install aiofiles==0.6.0```
- ```! pip install uvicorn==0.13.4```
- (For making HTTP requests to running application)
- ```! pip install requests==2.25.1```
- `Dockerfile` : <i>to package this project as REST API</i>
- This is based on ```tiangolo/uvicorn-gunicorn-fastapi:latest``` image which has ```python==3.8.6``` preinstalled.
</div>
---
# Application Packaging
<label style='color:#FF4500;'> There are many ways to productionise a machine learning model. Here, I have approached it as a dedicated server (depends on usage and consumption scenarios) using light-weight high performant ```fastAPI``` framework containerised using ```Docker```. Model can also be deployed as ```Cloud function``` for minimising cost.</label>
<label style='color:#FF4500;'>Further, the current deployment strategy allows to synchronously as well asynchronously train the model. Async, one can send/submit a traning job to sever, which in response sends a operation-location. This operation-location can be later accessed to retrive model's weights in form of ```.npy``` file. This means one do not wait for the response of your ```train``` request to server. Opposite to that, using synchronous endpoint, you have to wait till server returns the response, thus making it suitable for smaller grid resolution. You will see this is the approach I have taken in demonstrating the use of application seamlessly working with smaller and larger grid resolutions. You will notice that smaller grids such as `10x10` are run `Synchronously` whereas the `100x100` grid is demonstrated using `Asynchronous` approach.
</label>
# Navigating this notebook
This notebook is created to demonstrate the Application Packaging strategy along with show the training results. The following is repeated for all different endpoints of application.
- Consuming Endpoint Examples
- endpoint name
- method / cell demonstrating the use of endpoint
- result from calling the endpoint
This section examplifies saved models already packaged within the docker image.
- Sample Examples
- 10x10 100N
- 10X10 200N
- 10X10 500N
- 100X100 1000N
---
Build/Running containised (on 8000/your choice port):
-----------------------------------------------------
1. ```docker build . -t kohonen``` (assuming your current directory is this project's directory)
2. ```docker run -d --name kohonen -p 8000:80 kohonen:latest```
---
Running from source code:
------------------------
#### As local server:
1. ```cd app/```
2. ```uvicorn main:app``` (default port is 8000)
#### As local via python file:
- Look for ```app/test.py``` <a href='app/test.py'>test.py</a> file
- It has sample tests written for running the network over following configurations:
1. 10X10 for 100N
2. 10X10 for 200N
3. 10X10 for 500N
4. 100X100 for 1000N
----
Productioning approach as REST Service
---------------------
<h2 style='color:#FF4500;'>Using '/atrain/'</h2>
Recommended way is to perform training by performing a POST request at ```/atrain/``` with payload as following:
```
payload = {
"grid_shape" : [10, 10],
"max_iterations" : 100,
"learning_rate" : 0.1,
"training_data" : input_sample.tolist(),
"training_data_shape" : input_sample.shape
}
```
```input_sample``` is ```numpy.array``` what you want to project over grid of shape ```grid_shape``` for dimentiality reduction.
The response of that would be a json object that would contain a key called ```operation-location```. The value of that is a unique identifier (or ```model_name```) to name your training job.
Once the training has completed, you would be able to download ```model_name``` file (model_name.npy) by performing a GET request at ```/download/{model_name}``` and getting status code of 200 i.e. the model has been trained and saved under ```model_name``` name. The downloaded ```.npy``` file can be therefore consumed for analysis, plotting etc.
<label style='color:#FF4500;'>Using ```/atrain/``` is ideal especially when training on over large grids (order of size 100X100 or higher)</label>
<h2 style='color:#FF4500;'> Using '/train/' </h2>
This is a synchronous way to send a training request and receiving response ONLY after model is trained and saved. The response will be same as of ```/atrain/``` containing ```operation-location``` key and value as trained model's unique identifier. Further you can follow same process to download it and consume it.
<h2 style='color:#FF4500;'> Using '/list-of-models/' </h2>
Allows you to get list of all preexisting trained model.
<h2 style='color:#FF4500;'>Using '/predict/'</h2>
Allows to map test input vector to gird nodes i.e. which test input color is closest to which node in the grid. Returns list of `x`, `y` coordinates of grid nodes which can be plotted over grid for visualising & labelling test input using a trained model.
<h2 style='color:#FF4500;'>Using '/download/{model_name}/'</h2>
This is useful endpoint which works in conjuction with ```/train``` and ```/atrain/``` api to get the trained model file. Additionally, we can build a UI that can facilitate model visualisation post training. I have not gone to the extend of building the UI to visualise the trained grid, however it can be a consumer facing application for clustering high dimentional data such as geographical data etc.
----
Saving Trained Model
---------------------
All trained models are saved using unique identifier of format ```{d-m-Y_HhMmSs}T_{grid_x}X{grid_y}_{max_iterations}N_{learning_rate}LR``` and as ```.npy``` file under ```saved_models``` folder.
For example: ```05-03-2021_23h35m57sT_10X10_500N_0.1LR``` indicates that the model training started at `05th` of `March` `2021` at `23hrs` `35mins` `57seconds` (UTC) on grid of size `10X10` for `500` iterations and initial learning rate of `0.1`
---
---
# Application Endpoints
#### Installing required packages
```
# Core Libs
# ! pip install numpy==1.20.1
# ! pip install matplotlib==3.3.4
# Package Libs
# ! pip install fastapi==0.63.0
# ! pip install aiofiles==0.6.0
# ! pip install uvicorn==0.13.4
# For making HTTP requests to running applicaiton
# ! pip install requests==2.25.1
# OR via requirements file
# 1. cd ```configs/```
# 2. run pip install -r prod.requirements.txt
import requests as req
import numpy as np
import matplotlib.pyplot as plt
import json
```
### Input Data of Shape (10,3)
```
shape = (10,3)
# Sample Data
data_sample = np.array(
[[0.09803922, 0.78823529, 0.27058824],
[0.17254902, 0.8745098 , 0.18823529],
[0.82352941, 0.6 , 0.49411765],
[0.83529412, 0.56078431, 0.65098039],
[0.85098039, 0.74509804, 0.54509804],
[0.06666667, 0.3254902 , 0.63529412],
[0.1254902 , 0.56470588, 0.09411765],
[0.82352941, 0.00392157, 0.61176471],
[0.91764706, 0.85882353, 0.70980392],
[0.38039216, 0.68627451, 0.70980392]]
)
# Random Data
# data_sample = np.random.randint(0, 255, size=(shape[0], shape[1])) / 255.0
def scatter_input_data(input_data):
"""
Helper method to scatter list of 3D colors.
"""
_total = len(input_data)
x = range(_total),
y = np.ones(_total)
plt.scatter(x,y, c=input_data, s=100)
scatter_input_data(data_sample)
# Saves training input data
# np.save("temp/data_sample.npy", data_sample)
```
Above uses ```temp/``` folder to save training sample
---
---
# Consuming Endpoints (Examples)
```
port = 8000 # Set your port here, if you are running Docker or Uvicorn on different port
# endpoint = f"http://localhost:{port}" # Docker
endpoint = f"http://127.0.0.1:{port}" # Uvicorn
```
## GET /download/{model_name}/ : Download trained model
This is useful endpoint which works in conjuction with ```/train``` and ```/atrain/``` api to get the trained model.
Additionally, we can build a UI that can facilitate model visualisation post training. I have not gone to the extend of building the UI to visualise the trained grid, however it can be a consumer facing application for clustering high dimentional data such as geographical data etc.
#### Save downloaded model and view grid
This will examplfied later when consuming ```/train/``` and ```/atrain/```
```
def download(model_name:str)->bytes:
"""
Examplifies how to download a model (weights) via name.
Do not use .npy extension in the name.
Parameters
----------
model_name : str
name of model / operation-location returned by /train/, /atrain/
Returns
-------
file bytes : bytes
Downloads the model file (.npy)
Example
--------
model_name = '10x10 100N' instead of '10x10 100N.npy'
file_bytes = download("10x10 100N")
"""
download_endpoint = f"{endpoint}/download/{model_name}/"
try:
response = req.get(url=download_endpoint)
if response.status_code == 200:
return response.content
except:
print("Could not download model file. Response content: ", response.content)
print(f"Status code: {response.status_code}")
return None
def load_weights(file_bytes):
"""
Helper method to save the downloaded model file bytes
into a temporary sample .npy file at 'temp/' folder
Parameters
----------
file_bytes : bytes
file bytes of model wieghts downloaded using 'def download' above.
Returns
-------
weights : numpy.array
"""
temp_path = "temp/sample_weights.npy"
with open(temp_path, 'wb') as f:
f.write(file_bytes)
return np.load(temp_path)
def view(file_bytes):
"""
For simplicity 'sample.npy' is same name for downloading the temporary file,
So, immediately after downloading the model file, you can use this to view/plot.
Parameters
----------
file_bytes :
Example
-------
response = response from /train/ , /atrain/ ...
model_name = response.json()["operation-location"]
model_weights = download(model_name) # download file bytes
view(model_weights) # converts bytes to .npy file, saves it and shows the grid.
"""
if file_bytes is None:
return "File bytes is None! Please check if downloaded the file properly!"
weights = load_weights(file_bytes)
plt.imshow(weights)
```
Above uses ```temp/``` folder to save model weights under ```sample_weights.npy``` name. If you are running view multiple times, make sure ```download``` was run before it with correct model name else, ```view``` will try to load last saved model
---
## GET /list-of-models/ : List of already trained models.
Allows you to know what existing models are trained.
```
req.get(url=f"{endpoint}/list-of-models/").json()
```
---
## POST /train/ : Synchronous Training
Allows you to train the grid Synchronously. That means you have to wait till the grid completes its training to get REST response.
## 2.1 Train a 10x10 network over 100 iterations
```
payload = {
"grid_shape" : [10,10],
"max_iterations" : 100,
"learning_rate" : 0.1,
"training_data" : data_sample.tolist(),
"training_data_shape" : data_sample.shape,
"debug_mode": False,
"verbose" : False,
}
response = req.post(url=f"{endpoint}/train/", data=json.dumps(payload))
response.json()
```
##### INFO: Try running the above GET again to see the model saved .
#### Download/View
```
model_name_10x10_100N = response.json()["operation-location"]
model_weights_10x10_100N = download(model_name_10x10_100N)
view(model_weights_10x10_100N)
```
<h2> NOTE : <i> Around 10 runs were performed for calculating average times indendently (i.e. only single model was trained at a time).
All of the runs are only for training set of 10 RGB colors with initial learning rate of 0.1.</i>
</h2>
<p style='color:#FF4500'>
<b>Dockerised</b> : indicates running the project from docker image as local server.<br>
<b>Not Dockerised</b> : indicates running the project from command line as local server.
</p>
- How long does this take?
<p style="color:#FF4500">Running this training independantly (i.e. server is only performing this task), it was observed that it takes less than <b>5 seconds</b> to train.</p>
<table style="width:20%;text-align: center">
<tr>
<th>Type</th>
<th>Average Time</th>
</tr>
<tr>
<td>Dockerised</td>
<td>~ 3-4 seconds</td>
</tr>
<tr>
<td>Dockerised with Vectorisation</td>
<td style="color:#FF4500">~ 0.2-0.3 seconds</td>
</tr>
<tr>
<td>Not Dockerised</td>
<td>~ 10 seconds</td>
</tr>
</table>
- What does the map look like? (You will need to translate the weights of each node in the map to pixel data)
<p style='color:#FF4500;'> Influence of training items can been seen on the map where similar colors/shades grouped together towards same area and different colors (RGB values) move away from each other. Since resolution is very low 10X10, the is overlapping of different groups at boundaries. In order to determine more groups (if exists), it is essential to either increase number of iterations or to increase the grid size.</p>
## 2.2 Train a 10x10 network for 200 iterations
```
payload["max_iterations"] = 200
payload
response = req.post(url=f"{endpoint}/train/", data=json.dumps(payload))
response.content
```
#### Download/View
```
model_name_10x10_200N = response.json()["operation-location"]
model_weights_10x10_200N = download(model_name_10x10_200N)
view(model_weights_10x10_200N)
```
- How long does this take?
<p style="color:#FF4500">Running this training independantly (i.e. server is only performing this task), it was observed that it takes less than <b>10 seconds</b> to train.</p>
<table style="width:20%;text-align: center">
<tr>
<th>Type</th>
<th>Average Time</th>
</tr>
<tr>
<td>Dockerised</td>
<td>~ 7.3 seconds</td>
</tr>
<tr>
<td>EDIT: Dockerised with Vectorization</td>
<td style="color:#FF4500">~ 0.3-0.4 seconds</td>
</tr>
<tr>
<td>Not Dockerised</td>
<td>~ 20 seconds</td>
</tr>
</table>
- What does the map look like?
<p style='color:#FF4500;'> The map looks very same as 10x10 100N run with change in absolute positions. A light color is associated with colors that have higher average eucledian distance from BMU. For example, BMU for a green input-training-item have a cluster with the lightest-green being out far. BMU must be at the centre of a particular group, and the more closer a node is to BMU, the more similar/dark the color of node becomes.</p>
## 2.3 Train a 10x10 network for 500 iterations
```
payload["max_iterations"] = 500
payload
response = req.post(url=f"{endpoint}/train/", data=json.dumps(payload))
response.content
```
#### Download/View
```
model_name_10x10_500N = response.json()["operation-location"]
model_weights_10x10_500N = download(model_name_10x10_500N)
view(model_weights_10x10_500N)
```
- How long does this take?
<p style="color:#FF4500">Running this training independantly (i.e. server is only performing this task), it was observed that it takes less than <b>20 seconds</b> to train.</p>
<table style="width:20%;text-align: center">
<tr>
<th>Type</th>
<th>Average Time</th>
</tr>
<tr>
<td>Dockerised</td>
<td>~ 20 seconds</td>
</tr>
<tr>
<td>EDIT: Dockerised with Vectorization</td>
<td style="color:#FF4500">~ 1.2 - 1.5 seconds</td>
</tr>
<tr>
<td>Not Dockerised</td>
<td>~ 35 seconds</td>
</tr>
</table>
- What does the map look like?
<p style='color:#FF4500;'> Model is better able to move similar group of colours to a area on grid than previous run. With more iterations (500 > 100,200), the algorithm gets more oppurtunities to map the training input to lower dimensions - making a color/node more closer to the BMU.</p>
---
## POST /atrain/ : Asynchronous Training
Allows you to train the grid Asynchronously. That means you dont have to wait till the grid completes its training to get REST response. Both ```/train/```, ```/atrain/``` will have ```operation-location``` key:value pair in the response. You can futher use it to download the trained model using ```/download/```
<p style='color:#FF4500;'>Useful and practical for training large grids of sizes of order >= 100X100</p>
## 3. Train a 100x100 network over 1000 iterations
#### Training using the SYNCHRONOUS /train/
```
payload["grid_shape"] = [100, 100]
payload["max_iterations"] = 1000
response = req.post(url=f"{endpoint}/train/", data=json.dumps(payload))
response.content
model_name_100x100_1000N = response.json()["operation-location"]
model_weights_100x100_1000N = download(model_name_100x100_1000N)
view(model_weights_100x100_1000N)
```
# ---
```
payload["debug_mode"] = False
payload["grid_shape"] = [100, 100]
payload["max_iterations"] = 1000
```
##### Below are optional settings (default to False) which allow the algorithm to run multiprocessing at various targets mentioned above in optimisation section.
```
payload["pp_FIND_BMU"] = False
payload["pp_INF_BMU_W"] = False
payload["pp_INF_BMU_POS"] = False
payload
```
#### Training using the ASYNCHRONOUS /atrain/
```
# POST Request asynchronous
aresponse = req.post(url=f"{endpoint}/atrain/", data=json.dumps(payload))
aresponse_body = aresponse.json()
a_model_name_100X100_1000N = aresponse_body["operation-location"]
print(f"Training for '{a_model_name_100X100_1000N}' has started. You can run below 2-cells to wait and download the model")
```
Since it is asynchronous call, your model might not be trained just yet. So, if you try to download file at ```operation-location```, you will probably receive 404 status code in response.
An approach to would be trying to get the file at regular intervals and when you receive 200 status code in response, you will be downloading the file.
```
import timeit
import time
async def async_download(model_name, wait_between_calls=20):
"""
Helper method that faciliates downloading trained model by looking for its saved file
with time iterval.
Parameters
----------
model_name (operation-location) : str
The operation location that was provided by API under
which the model file will be saved.
wait_between_calls : int
Seconds to wait between each try to download the file from application sever.
"""
while True:
download_endpoint = f"{endpoint}/download/{model_name}/"
response = req.get(url=download_endpoint)
if response.status_code == 404:
time.sleep(wait_between_calls)
elif response.status_code == 200:
view(response.content)
break
else:
raise Exception("Exception at server.")
break
```
#### Execute cell below to wait and try download the model.
<p style='color:#FF4500;'> <b>NOTE:</b> ONLY run cell below, if you have POSTed a asynchronous requests to server to train. The executing of below cell will await for the model to finish train. Otherwise, you can skip it anyway and continue with other cells and come back later to this one (as running this cell is a blocking execution) </p>
```
print(f"Starting to wait for '{a_model_name_100X100_1000N}' model!")
start_time = timeit.default_timer()
await async_download(a_model_name_100X100_1000N)
print("{:.2f}".format(timeit.default_timer() - start_time))
```
---
## GET /predict/ : To predict similarity from a trained model
Allows to map test input vector to gird nodes i.e. which test input color is closest to which node in the grid.
Returns list of (x,y) coordinates of grid nodes which can be used to plot test input over gird plot for visualising/analysing/labelling test input using a trained model.
```
def predict(data, my_model_name):
"""
Examplifies how to send test data for mapping
it to model using '/predict/' endpoint.
Parameters
----------
data : numpy.array
Testing items (colors) to be clustered.
my_model_name : str
Name that identifies model traning.
This string is retured by /atrain/, /train/ endpoints as `operation-location`
Returns
-------
response : json, dict
Dictionary object containing predicted x,y coordinates (indicies) of grid nodes,
that are closest to testing item.
"""
payload = {
"test_data" : data.tolist(),
"test_data_shape" : data.shape,
"model_name" : my_model_name
}
response = req.get(url=f"{endpoint}/predict/", data=json.dumps(payload))
return response.json()
def plot_predictions(file_bytes:bytes, test_data, predicted_grid_idxs):
"""
Helper method to plot input data (colours) over trained grid (@weights) using @predicted_grid_idxs.
For anaylsis, index of input data (individual colors) is also plotted.
Parameters
---------
file_bytes : bytes
A model's weights file bytes,
which are response of /download/.
test_data: list, numpy.array
Similar to training data, it is list of 3D colors.
predicted_grid_idxs : list
Response from /predict/ endpoint.
"""
plt.imshow(load_weights(file_bytes))
indicies = np.array(predicted_grid_idxs)
y, x = indicies.T
plt.scatter(x, y, c=test_data, s=40, edgecolors='b')
for i, _xy in enumerate(predicted_grid_idxs):
plt.text(_xy[1],_xy[0], i, ha='left', va="top" )
plt.show()
```
<label style="color:#FF4500;"> <b>NOTE:</b>You can set ```model_name``` to previous async run's model (```a_model_name_100X100_1000N```).</label> <br>
<label style="color:#FF4500;"> <b>NOTE:</b>You should set it to (```model_name_10x10_500N```) in case you have skipped async training, model has not YET been trainied (how you know? : if you have run the blocking cell ```await asynchronous_download``` and its execution has not yet finished!). </label>
```
model_name = model_name_10x10_500N
# model_name = a_model_name_100X100_1000N
model_weights = download(model_name)
# Getting Predictions
print(f"Predicting from {model_name} model.")
response = predict(data_sample, model_name)
predictions = response['predictions']
print(predictions)
```
### Remember the input?
```
scatter_input_data(data_sample)
```
#### Testing the imput again on the trained grid.
#### Here the following, I am scattering the same input vector and we can only see the boundaries as the pixel/node color has been matched with input training-item.
```
plot_predictions(model_weights, data_sample, predictions)
```
As assumed before, when input data itself is mapped to the grid via predicting, the BMUs stand straight in the middle of respective cluster.
---
---
# Optimisations
### Rule of Thumb of Optimisations
1. Avoided ```for``` loops for manipluating numpy arrays, calculations.
2. Used list comphresenshion instead of ```for``` loop whereever possible.
3. Used numpy vector operations such as ```.multiply```, ```.square```, ```.apply_along_axis``` etc were given more precedance over ```for``` loop.
### What could you do to improve performance?
#### Specific optimisations
To improve performance, it was discovered that main target area is executions per iteration. Overall algorithm is in its nature is iterative therefore, following target areas were discovered with possibilities to improve Time Complexity:
1. Finding BMU node, where all node weights of grid are compared with current input vector. (`pp_FIND_BMU`)
2. Calculating Influence:
- 2.1 Where all node weights are compared with BMU weights. (`pp_INF_BMU_W`)
- 2.2 Where all node positions are compared with BMU position. (`pp_INF_BMU_POS`)
<label style='color:#FF4500;'><b>When a function is applied to large numpy array and the function calculations are independant of other elements in array, an intuitive way is to partition the large array into chunks and use multiple processes that utilise (multiple) number of CPU cores available to allocate mapping thus parallelising the computation. Later, combine all the result from each process and pass it over.</b> </label>
### Observations
There are essentially following combinations of parallel processing that be applied.
1. Parallel Run Type-1 :
- A. when multiprocessing is ON ONLY at `pp_FIND_BMU`.
- B. when multiprocessing is ON ONLY at `pp_INF_BMU_W`.
- C. when multiprocessing is ON ONLY at `pp_INF_BMU_POS`.
2. Parallel Run Type-2 :
- A. when multiprocessing is ON at both `pp_FIND_BMU` and `pp_INF_BMU_W`.
- B. when multiprocessing is ON at both `pp_FIND_BMU` and `pp_INF_BMU_POS`.
- C. when multiprocessing is ON at both `pp_INF_BMU_W` and `pp_INF_BMU_POS`.
3. Parallel Run Type-3 : when multiprocessing is ON at all `pp_FIND_BMU` and `pp_INF_BMU_W` and `pp_INF_BMU_POS`
#### Smaller vs Larger Grid - multiprocessing overhead!
<p style='color:#FF4500;'>It was also found that when any type of multiprocessing is applied while training smaller grids like (10x10=100 nodes for 100N) there is more overhead and significant delay as compared to sequential execution. It was also found that multiprocessing is useful in case of large grids which are of order 100x100 or more.</p>
---
### What does the network look like after 1000 iterations?
<div style='color:#FF4500;'>
Yes, the network was significantly slower as both resolution of the grid and iterations are increased 10X than first run.
The input mapped in this higher resolution grid with large iterations, shows clear demarcations of group of classes that exist in input vector. Higher resolution shows intricate relationships among the input feature vectors which makes it much clearer representation than previous low resolution grids.
</div>
### Sequential execution, for case 100X100 1000N.
<table style='width:40%; text-align:center'>
<tr>
<th>Type</th>
<th>Average Time</th>
</tr>
<tr>
<td>Dockerised</td>
<td>~ 1-1.5 hours with 4-6 seconds per iteration</td>
</tr>
<tr>
<td>EDIT: Dockerised with Vectorization</td>
<td style="color:#FF4500">~ 24-26 seconds overall </td>
</tr>
<tr>
<td>Not Dockerised</td>
<td>~ 1.5-3 hours with 9-12 seconds per iteration</td>
</tr>
</table>
### Using ```multiprocessing```, for case 100X100 1000N.
<h4 style='color:#FF4500;'>Dockerised/production envrionment</h4>
<table style='width:60%;text-align: center'>
<tr>
<th>Type</th>
<th>Average Time (Dockerised)</th>
</tr>
<tr>
<td>Parallel Run Type 1 (A) - (pp_FIND_BMU)</td>
<td style="color:#FF4500">~ 0.9-1 hours</td>
</tr>
<tr>
<td>Parallel Run Type 1 (B) - (pp_INF_BMU_W)</td>
<td style="color:#FF4500">~ 0.9-1 hours</td>
</tr>
<tr>
<td>Parallel Run Type 1 (C) - (pp_INF_BMU_POS)</td>
<td>~ 1-1.2 hours</td>
</tr>
<tr>
<td>-</td>
</tr>
<tr>
<td>Parallel Run Type 2 (A) - (pp_FIND_BMU and pp_INF_BMU_W)</td>
<td>~ 1.5 hours</td>
</tr>
<tr>
<td>Parallel Run Type 2 (B) - (pp_FIND_BMU and pp_INF_BMU_POS)</td>
<td>more than 2 hours</td>
</tr>
<tr>
<td>Parallel Run Type 2 (C) - (pp_INF_BMU_W and pp_INF_BMU_POS)</td>
<td>more than 3 hours</td>
</tr>
<tr>
<td>-</td>
</tr>
<tr>
<td>Parallel Run Type 3 - (pp_FIND_BMU and pp_INF_BMU_W and pp_INF_BMU_POS)</td>
<td>more than 3 hours</td>
</tr>
</table>
---
<h2 style='color:#FF4500;'>Conclusion on how to improve efficiency of algorithm </h2>
- NOTE: ```multiprocessing``` used in the project, means the use of python's inbuilt ```multiprocessing``` module, and specifically using ```Pool``` class to divide numerical numpy calculations to CPU cores.
- I found for smaller grids such as 10x10 for x iterations, running the training without using ```multiprocessing``` yeilded quicker results i.e. training sequentially was faster.
- Using ```multiprocessing```
- Total training time during BMU node search (`Parallel Run Type 1 (A) pp_FIND_BMU`) was just shy of an hour (~ 0.95 hours).
- Similar training time in case of `Parallel Run Type 1 (B) pp_INF_BMU_W` as previous.
- `Parallel Run Type 1 (C) pp_INF_BMU_POS` took more time than both previous cases.
- Combination of Type 1 (A, B C) always took more than 1.5 hours
- Lastly, Type 3, where ```multiprocessing``` was used at each target area, takes more 3 hours to finish.
<label style='color:#FF4500;'><b> Therefore, to improve from 1-1.5 hours sequential training for 100x100 1000N configuration, I would select ```Parallel Run Type 1 (A)``` or ```(B)``` for ```multiprocessing```. The above were tested on Windows i7 4-core Intel processors. If I would have more resources, I would like to test these configuration on different hardware CPU settings and see what results I get.</b></label>
---
### Prospective Improvements
There are other packages that I have read such as ```Dask```, or ```Ray```, which I would like to try and see any improvements than my current approach. I didn't went on to examine further using these two packages, as not recommeded as per the guidelines for this challenge i.e. to only use numpy and python in-built packages.
### Algorithm Time Complexity
Overall algorithm goes through N iterations O(N), but for each iteration it has to compare BMU for O(S) (Point 1) size of nodes and then calculuate influence adding `O(I)`+`O(I)` (Point 2.1 and 2.2) = `O(N* (S+I_1+I_2))` = `O(M^2)` where `M` is highly influenced by size of grid.
<h1 style='color:#FF4500;'>
EDIT : Optimisation :
Dockerised with Vectorization WINS
<h1>
---
# Download/View (Saved) Trained Examples
# 10 x 10 for 100N
```
model_name = "10X10_100N"
model_weights = download(model_name)
view(model_weights)
```
# 10 x 10 for 200N
```
model_name = "10X10_200N"
model_weights = download(model_name)
view(model_weights)
```
# 10 x 10 for 500N
```
model_name = "10X10_500N"
model_weights = download(model_name)
view(model_weights)
```
# 100 x 100 for 1000N
```
model_name = "100X100_1000N"
model_weights = download(model_name)
view(model_weights)
```
| github_jupyter |

# Circuit Rewriting using the Transpiler
Previously we have performed basic operations on circuits, and ran those circuits on real quantum devices using the `execute` function. `execute` is a helper function that performs three tasks for the user:
1) Circuits are rewritten to match the constraints of a given backend and optimized.
2) The rewritten circuits are packaged for submission.
3) The packaged circuits are submitted to the device.
The first step is called transpilation, and is the fundamental step required for running circuits on real quantum devices. The qiskit function that does this is `transpile`, which takes a single or list of input circuits, as well as a collection of parameters, and returns a modified list of circuits. Using this `transpile` function is the goal of this tutorial.
For those wishing to bypass this in-depth discussion of what happens during the transpilation process, we note that the standard way of calling the `transpile` function is:
```python
new_circuits = transpile(circuits, backend, optimization_level=1)
```
where `circuits` is a single or list of input circuits, `backend` is the target device, and `optimization_level` selects between four different `[0,1,2,3]` preset methods of circuit rewriting and optimization. By default, `optimization_level=1`.
# Table of contents
1) [Introduction](#introduction)
2) [Basis Gates](#basis)
3) [Initial Layout](#layout)
4) [Mapping Circuits to Hardware](#mapping)
5) [Optimizing Gates](#optimizing)
6) [User Defined Topologies](#topology)
7) [Passing Transpiled Circuits to Devices](#passing)
```
import numpy as np
from qiskit import *
from qiskit.visualization import plot_histogram, plot_gate_map, plot_circuit_layout
from qiskit.tools.monitor import job_monitor
import matplotlib.pyplot as plt
%matplotlib inline
IBMQ.load_account()
provider = IBMQ.get_provider(group='open')
```
## Introduction <a name='introduction'></a>
Consider the following circuit that creates a five-qubit GHZ state:
```
ghz = QuantumCircuit(5, 5)
ghz.h(0)
for idx in range(1,5):
ghz.cx(0,idx)
ghz.barrier(range(5))
ghz.measure(range(5), range(5))
ghz.draw(output='mpl')
```
We would like to take this circuit and run it on an IBM Q quantum device. However, in its original form, this circuit cannot be run. This circuit, as is the case with most circuits, must undergo a series of transformations that make it compatible with a given target device, and optimize it to reduce the effects of noise on the resulting outcomes.
As we will see, rewriting quantum circuits to match hardware constraints and optimizing for performance can be far from trivial. The flow of logic in the rewriting tool chain need not be linear, and can often have iterative sub-loops, conditional branches, and other complex behaviors. That being said, the basic building blocks follow the structure given below.

In what follows, we highlight the key steps in this workflow, and show how they modify circuits based on the value of keyword arguments passed to the `transpile` function. The `transpile` function has many input arguments:
```
import inspect
inspect.signature(transpile)
```
Here we focus only on the most important ones: `basis_gates`, `initial_layout` `optimization_level` and `coupling_map`.
## Basis Gates <a name='basis'></a>
When writing a quantum circuit you are free to use any quantum gate (unitary operator) that you like, along with a collection of non-gate operations such as qubit measurements and reset operations. However, when running a circuit on a real quantum device one no longer has this flexibility. Due to limitations in, for example, the physical interactions between qubits, difficulty in implementing multi-qubit gates, control electronics etc, a quantum computing device can only natively support a handful of quantum gates and non-gate operations. In the present case of IBM Q devices, the native gate set can be found by querying the devices themselves, and looking for the corresponding attribute in their configuration:
```
provider = IBMQ.get_provider(group='open')
provider.backends(simulator=False)
backend = provider.get_backend('ibmqx2')
backend.configuration().basis_gates
```
We see that the `ibmqx2` device supports five native gates: four single-qubit gates (`u1`, `u2`, `u3`, and `id`) and one two-qubit entangling gate `cx`. In addition, the device supports qubit measurements (otherwise we can not read out an answer!). Although we have queried only a single device, let us note that all IBM Q devices support this gate set.
The `u*` gates represent arbitrary single-qubit rotations of one, two, and three angles. The `u1` gates are single-parameter rotations that represent generalized phase gates of the form
$$
U_{1}(\lambda) = \begin{bmatrix}
1 & 0 \\
0 & e^{i\lambda}
\end{bmatrix}
$$
This set includes common gates such as $Z$, $T$, $T^{\dagger}$, $S$, and $S^{\dagger}$. It turns out that these gates do not actually need to be performed on hardware, but instead, can be implemented in software as "virtual gates". These virtual gates are called "frame changes" and take zero time, and have no associated error; they are free gates on hardware.
Two-angle rotations, $U_{2}(\phi,\lambda)$, are actually two frame changes with a single $X_{\pi/2}$ gate in between them, and can be used to synthesize gates like the Hadamard ($U_{2}(0,\pi)$) gate. As the only actual gate performed is the $X_{\pi/2}$ gate, the error and gate time associated with any $U_{2}$ gate is the same as an $X_{\pi/2}$ gate. Similarly, $U_{3}(\theta,\phi,\lambda)$ gates are formed from three frame changes with two $X_{\pi/2}$ gates in between them. The errors and gate times are twice those of a single $X_{\pi/2}$. The identity gate, $id$, is straightforward, and is a placeholder gate with a fixed time-interval.
The only entangling gate supported by the IBM Q devices is the CNOT gate (`cx`) which, in the computational basis, can be written as:
$$
\mathrm{CNOT}(0,1) = \begin{bmatrix}
1 & 0 & 0 & 0 \\
0 & 0 & 0 & 1 \\
0 & 0 & 1 & 0 \\
0 & 1 & 0 & 0
\end{bmatrix}
$$
where the matrix form above follows from the specific bit-ordering convention used in Qiskit. (This was discussed in [Part 1: Getting Started with Qiskit](1_getting_started_with_qiskit.ipynb#Statevector-backend).)
Every quantum circuit run on an IBM Q device must be expressed using only these basis gates. For example, suppose one wants to run a simple phase estimation circuit:
```
qc = QuantumCircuit(2, 1)
qc.h(0)
qc.x(1)
qc.cu1(np.pi/4, 0, 1)
qc.h(0)
qc.measure([0], [0])
qc.draw(output='mpl')
```
We have $H$, $X$, and controlled-$U_{1}$ gates, all of which are not in our devices basis gate set, and must be expanded. This expansion is taken care of for us in the `execute` function. However, we can decompose the circuit to show what it would look like in the native gate set of the IBM Q devices
```
qc_basis = qc.decompose()
qc_basis.draw(output='mpl')
```
A few things to highlight. One, as mentioned in [Part 4: Quantum Circuit Properties](4_quantum_circuit_properties.ipynb), the circuit has gotten longer with respect to the initial one. This can be verified by checking the depth of the circuits:
```
print(qc.depth(), ',', qc_basis.depth())
```
Two, although we had a single controlled gate, the fact that it was not in the basis set means that, when expanded, it requires more than a single `cx` gate to implement. All said, unrolling to the basis set of gates leads to an increase in the depth of a quantum circuit and the number of gates.
It is important to highlight two special cases:
### SWAP Gate Decomposition
A SWAP gate is not a native gate on the IBM Q devices, and must be decomposed into three CNOT gates:
```
swap_circ = QuantumCircuit(2)
swap_circ.swap(0, 1)
swap_circ.decompose().draw(output='mpl')
```
As a product of three CNOT gates, SWAP gates are expensive operations to perform on a noisy quantum devices. However, such operations are usually necessary for embedding a circuit into the limited entangling gate connectivities of actual devices. Thus, minimizing the number of SWAP gates in a circuit is a primary goal in the transpilation process.
### Toffoli Gate Decomposition
A Toffoli, or controlled-controlled-not gate, is a three qubit gate. Given that our basis gate set includes only single- and two-qubit gates, it is obvious that this gate must be decomposed. This decomposition is quite costly.
```
ccx_circ = QuantumCircuit(3)
ccx_circ.ccx(0, 1, 2)
ccx_circ.decompose().draw(output='mpl')
```
For every Toffoli gate in a quantum circuit, the IBM Q hardware may execute up to six CNOT gates, and a handful of single-qubit gates. From this example, it should be clear that any algorithm that makes use of multiple Toffoli gates will end up as a circuit with large depth and will therefore be appreciably affected by noise and gate errors.
## Initial Layout <a name='layout'></a>
Quantum circuits are abstract entities whose qubits are "virtual" representations of actual qubits used in computations. We need to be able to map these virtual qubits in a one-to-one manner to the "physical" qubits in an actual quantum device.

By default, qiskit will do this mapping for you. The choice of mapping depends on the properties of the circuit, the particular device you are targeting, and the optimization level that is chosen. The basic mapping strategies are the following:
- **Trivial layout**: Map virtual qubits to the same numbered physical qubit on the device, i.e. `[0,1,2,3,4]` -> `[0,1,2,3,4]` (default in `optimization_level=0`).
- **Dense layout**: Find the sub-graph of the device with same number of qubits as the circuit with the greatest connectivity (default in `optimization_level=1`).
- **Noise adaptive layout**: Uses the noise properties of the device, in concert with the circuit properties, to generate the layout with the best noise properties (default in `optimization_level=2` and `optimization_level=3`).
The choice of initial layout is extremely important when:
1) Computing the number of SWAP operations needed to map the input circuit onto the device topology.
2) Taking into account the noise properties of the device.
As we will see, the choice of `initial_layout` can mean the difference between getting a result, and getting nothing but noise.
To begin, lets see what layouts are automatically picked at various optimization levels. The modified circuits returned by `transpile` have this initial layout information in them, and we can view this layout selection graphically using `plot_circuit_layout`. Let us pick the `ibmq_16_melbourne` device:
```
backend = provider.get_backend('ibmq_16_melbourne')
plot_gate_map(backend, plot_directed=True)
```
### Layout using `optimization_level=0`
```
backend = provider.get_backend('ibmq_16_melbourne')
new_circ_lv0 = transpile(ghz, backend=backend, optimization_level=0)
plot_circuit_layout(new_circ_lv0, backend)
```
### Layout using `optimization_level=3`
```
from qiskit.visualization import plot_circuit_layout
backend = provider.get_backend('ibmq_16_melbourne')
new_circ_lv3 = transpile(ghz, backend=backend, optimization_level=3)
print('Depth:', new_circ_lv3.depth())
plot_circuit_layout(new_circ_lv3, backend)
```
Let us now execute this level 3 circuit, and get the counts. Note that the circuit has already been rewritten to match the target backend. Therefore, the circuit will go unmodified by `execute` and be directly packaged and sent to the device.
```
job1 = execute(new_circ_lv3, backend)
job_monitor(job1)
```
### Specifying an Initial Layout
It is completely possible to specify your own initial layout. To do so we can pass a list of integers to `transpile` via the `initial_layout` keyword argument, where the index labels the virtual qubit in the circuit and the corresponding value is the label for the physical qubit to map onto. For example, lets map our GHZ circuit onto `ibmq_16_melbourne` in two different ways:
#### Good choice
```
# Virtual -> physical
# 0 -> 11
# 1 -> 12
# 2 -> 10
# 3 -> 2
# 4 -> 4
good_ghz = transpile(ghz, backend, initial_layout=[11,12,10,2,4])
print('Depth:', good_ghz.depth())
plot_circuit_layout(good_ghz, backend)
job2 = execute(good_ghz, backend)
job_monitor(job2)
```
#### Bad choice
```
# Virtual -> physical
# 0 -> 0
# 1 -> 6
# 2 -> 10
# 3 -> 13
# 4 -> 7
bad_ghz = transpile(ghz, backend, initial_layout=[0,6,10,13,7])
print('Depth:', bad_ghz.depth())
plot_circuit_layout(bad_ghz, backend)
job3 = execute(bad_ghz, backend)
job_monitor(job3)
counts1 = job1.result().get_counts()
counts2 = job2.result().get_counts()
counts3 = job3.result().get_counts()
plot_histogram([counts1, counts2, counts3],
figsize=(15,6),
legend=['level3', 'good', 'bad'])
```
From the figure it is clear that the choice of initial layout is an extremely important step, and can mean the difference between a good answer and one dominated by noise. Although the 'good' circuit has a smaller depth, the results generated by the "level 3"-circuit are more in line with the expected answer. This is because our manual layout selection did not take into account the noise properties of the device. Noise affects each qubit differently, and qubit measurement errors can also corrupt the answer. The noise adaptive layout in `optimization_level=2` and `optimization_level=3` takes this into account, and the benefits are obvious.
### Being a bit Smarter
Picking a good `initial_layout` is critical, but it is not the only step in optimizing a circuit on a quantum device. Often it is beneficial to rewrite a circuit, with the goal of targeting a specific device. Our GHZ circuit can be perfectly mapped to the `ibmq_16_melbourne` device provided that we reformulate the gate sequence as:
```
ghz2 = QuantumCircuit(5, 5)
ghz2.h(2)
ghz2.cx(2, 1)
ghz2.cx(1, 0)
ghz2.cx(2, 3)
ghz2.cx(3, 4)
ghz2.barrier(range(5))
ghz2.measure(range(5), range(5))
ghz2.draw(output='mpl')
```
This circuit can now be mapped exactly onto the device topology using `initial_layout=[10,4,5,6,8]`. This can be verified by once again computing the depth.
```
exact_ghz = transpile(ghz2, backend, initial_layout=[10,4,5,6,8])
print('Depth:', exact_ghz.depth())
plot_circuit_layout(exact_ghz, backend)
job4 = execute(exact_ghz, backend)
job_monitor(job4)
counts4 = job4.result().get_counts()
plot_histogram([counts1, counts4],
figsize=(15,6),
legend=['level3', 'exact'])
```
## Mapping Circuits to Hardware Topology <a name='mapping'></a>
Our original GHZ circuit consists of a single qubit (`0`) coupled to the others via CNOT gates. If implemented directly, this would require hardware that has a single qubit coupled to four other qubits. Looking at the gate maps of the public IBM Q hardware we see that there is one device that fits this description.
```
backend = provider.get_backend('ibmqx2')
plot_gate_map(backend)
```
and five that do not. Four that have the following topology:
```
backend = provider.get_backend('ibmq_ourense')
plot_gate_map(backend)
```
and one ladder-type configuration:
```
backend = provider.get_backend('ibmq_16_melbourne')
plot_gate_map(backend)
```
It seems our circuit is a good match to the `ibmqx2` topology provided that we map virtual qubit `0` to physical qubit `2`. The arrows on the graphs above indicate that CNOT gates are only allowed in one direction. However, flipping the direction of a CNOT gate is relatively trivial, and does not greatly affect the fidelity of the computation.
`ibmq_ourense`, `ibmq_vigo`, and `ibmq_16_melbourne` represent a problem. There is no qubit anywhere that is connected to four others via CNOT gates. In order to implement a CNOT gate between qubits that are not directly connected, one or more SWAP gates must be inserted into the circuit to move the qubit states around until they are adjacent on the device gate map. Our choice of `initial_layout` in `job3` above was purposely designed to take many SWAP gates to make adjacent. As we have seen previously, each SWAP gate is decomposed into three CNOT gates on the IBM Q devices, and represents an expensive and noisy operation to perform. Thus, finding the minimum number of SWAP gates needed to map a circuit onto a given device, is an important step (if not the most important) in the whole execution process.
As with many important things in life, finding the optimal SWAP mapping is hard. In fact it is in a class of problems called NP-Hard, and is thus prohibitively expensive to compute for all but the smallest quantum devices and input circuits. To get around this, by default Qiskit uses a stochastic heuristic algorithm called `StochasticSwap` to compute a good, but not necessarily minimal SWAP count. The use of a stochastic method means the circuits generated by `transpile` (or `execute` that calls `transpile` internally) are not guaranteed to be the same over repeated runs. Indeed, running the same circuit repeatedly will in general result in a distribution of circuit depths and gate counts at the output.
In order to highlight this, we run the GHZ circuit 100 times, using the "bad" (disconnected) `initial_layout` from the previous section:
```
bad_circs = transpile([ghz]*100, backend, initial_layout=[0,6,10,13,7])
depths = [circ.depth() for circ in bad_circs]
plt.figure(figsize=(8,6))
plt.hist(depths, bins=list(range(34,50)), align='left', color='#648fff')
plt.xlabel('Depth', fontsize=14)
plt.ylabel('Counts', fontsize=14);
```
This distribution is quite wide, signaling the difficultly the SWAP mapper is having in computing the best mapping. Most circuits will have a distribution of depths, perhaps not as wide as this one, due to the stochastic nature of the default SWAP mapper. Of course, we want the best circuit we can get, especially in cases where the depth is critical to success or failure. In cases like this, it is best to `transpile` a circuit several times, e.g. 10, and take the one with the lowest depth. The `transpile` function will automatically run in parallel mode, making this procedure relatively speedy in most cases.
Just to highlight the difference when running on a different device, we show the same distribution when using the `ibmqx2` backend with the default settings:
```
backend = provider.get_backend('ibmqx2')
qx2_circs = transpile([ghz]*100, backend)
depths = [circ.depth() for circ in qx2_circs]
plt.figure(figsize=(8,6))
plt.hist(depths, bins=list(range(6,15)), align='left', color='#648fff')
plt.xlabel('Depth', fontsize=14)
plt.ylabel('Counts', fontsize=14);
```
## Optimizing Single- and Multi-Qubit Gates <a name='optimizing'></a>
Decomposing quantum circuits into the basis gate set of the IBM Q devices, and the addition of SWAP gates needed to match hardware topology, conspire to increase the depth and gate count of quantum circuits. Fortunately many routines for optimizing circuits by combining or eliminating gates exist. In some cases these methods are so effective the output circuits have lower depth than the inputs. In other cases, not much can be done, and the computation may be difficult to perform on noisy devices. Different gate optimizations are turned on with different `optimization_level` values. Below we show the benefits gained from setting the optimization level higher:
<div class="alert alert-block alert-success">
<b>Remember:</b> The output from <code>transpile</code> varies due to the stochastic swap mapper. So the numbers below will likely change each time you run the code.
</div>
```
backend = provider.get_backend('ibmq_16_melbourne')
circ0 = transpile(ghz, backend, optimization_level=0)
print('Depth:', circ0.depth())
print('Gate counts:', circ0.count_ops())
circ1 = transpile(ghz, backend, optimization_level=1)
print('Depth:', circ1.depth())
print('Gate counts:', circ1.count_ops())
circ2 = transpile(ghz, backend, optimization_level=2)
print('Depth:', circ2.depth())
print('Gate counts:', circ2.count_ops())
circ3 = transpile(ghz, backend, optimization_level=3)
print('Depth:', circ3.depth())
print('Gate counts:', circ3.count_ops())
```
## Transpiling for User Defined Topologies <a name='topology'></a>
Up to now, we have focused on rewriting and optimizing circuits for the IBM Q set of devices. However, `transpile` is much more general than that, and can be used to map circuits onto user defined device topologies. The topology is encoded in what qiskit calls the `coupling_map`, and the coupling map for a device can be obtained from its `configuration`:
```
backend = provider.get_backend('ibmqx2')
backend.configuration().coupling_map
```
The elements of this list are `[control, target]` pairs that indicate that a CNOT gate can be performed using those qubits. The coupling map is in essence an adjacency matrix characterizing the entangling gate coupling of the device:
```
from scipy.sparse import coo_matrix
cmap = backend.configuration().coupling_map
rows = [c[0] for c in cmap]
cols = [c[1] for c in cmap]
data = np.ones_like(rows)
adj_matrix = coo_matrix((data,(rows,cols)),shape=(5,5))
plt.spy(adj_matrix);
```
If it is possible to perform CNOT gates in both directions on all pairs of qubits, then this matrix is symmetric.
We are free to construct our own device topology by defining our own `coupling_map` and using it in `transpile`. For example, a five qubit linear nearest-neighbor (LNN) topology supporting bi-directional CNOT gates is written as:
```
lnn5 = [[0,1], [1,0], [1,2], [2,1], [2,3], [3,2], [3,4], [4,3]]
rows = [c[0] for c in lnn5]
cols = [c[1] for c in lnn5]
data = np.ones_like(rows)
adj_matrix = coo_matrix((data,(rows,cols)),shape=(5,5))
plt.spy(adj_matrix);
```
We can then use this `coupling_map` in place of an actual backend as follows:
```
lnn5_circ = transpile(ghz, backend=None, coupling_map=lnn5)
lnn5_circ.draw(output='mpl')
```
Or we can write a directional five qubit '+' shaped topology, with all directional CNOT gates pointing to the center as:
```
plus = [[0,2], [1,2], [3,2], [4,2]]
rows = [c[0] for c in plus]
cols = [c[1] for c in plus]
data = np.ones_like(rows)
adj_matrix = coo_matrix((data,(rows,cols)),shape=(5,5))
plt.spy(adj_matrix);
plus_circ = transpile(ghz, backend=None, coupling_map=plus)
plus_circ.draw(output='mpl')
```
In the above examples we can see that a Hadamard gate is still in the circuit. This is because with `backend=None` we also need to specify the `basis_gates` if we want a specific gate set at the end.
## Passing Transpiled Circuits to a Backend <a name='passing'></a>
Now that we have transpiled circuits, the final step is to pass them to a backend for execution. There are two ways of doing this:
1) Pass the circuits to `execute` as done before.
2) Use the `assemble` function to package them, and then call the backend directly.
### Using `execute` with transpiled circuits
Typically one is interested in transpiling circuits for the same backend that they intend to run their circuits on. In this case, the circuits have already been mapped to the backend and optimized, so calling:
```
job = execute(transpiled_circuits, backend, **kwargs)
```
will simply package the circuits up, and send them to the target backend. The exception to this rule is if the level of optimization is different than what the circuits were originally transpiled with. That is to say setting `optimization_level=3` in `execute` while passing circuits transpiled under `optimization_level=1` (the default) will undergo further optimization before being sent to the device.
### Packaging Circuits with `assemble`
It is also possible to skip the `execute` function and bundle the transpiled circuits using the `assemble` function. The `assemble` function is designed to take both, circuits and pulse schedules, and has lots of input arguments:
```
inspect.signature(assemble)
```
many of which overlap with those in `execute`. Here `experiments` is the circuit or list of circuits to be run, and `backend` is of course the backend you are targeting. The other important ones are:
- **shots** - Number of times to execute each circuit.
- **memory** - Return the resulting bitstring for each shot rather than collecting counts.
A standard call to `assemble` would look like:
```
qobj = assemble(transpiled_circuits, backend, shots=2048)
```
where `qobj` is the container format used in sending circuits (and pulse experiments) to the devices. This `qobj` can be run by calling the `backend` directly:
```
job = backend.run(qobj)
```
which will return a `job` just like `execute`.
```
import qiskit.tools.jupyter
%qiskit_version_table
%qiskit_copyright
```
| github_jupyter |
ERROR: type should be string, got "https://colab.research.google.com/drive/11iBiBM58MXyrHedrGo1pqTucmlBDPbjn\n\n```\nfrom matplotlib import pyplot as plt\n%matplotlib inline\nimport numpy as np\nfrom keras.utils import np_utils\nfrom keras.models import Sequential\nfrom keras.layers.core import Dense, Dropout, Activation, Flatten\nfrom keras.callbacks import EarlyStopping\nfrom keras.layers import Conv2D, MaxPooling2D\nfrom keras.layers.normalization import BatchNormalization\nfrom keras import regularizers\nfrom keras.datasets import cifar10\n(X_train, y_train), (X_val, y_val) = cifar10.load_data()\nX_train = X_train.astype('float32')/255.\nX_val = X_val.astype('float32')/255.\nn_classes = 10\ny_train = np_utils.to_categorical(y_train, n_classes)\ny_val = np_utils.to_categorical(y_val, n_classes)\nimport matplotlib.pyplot as plt\n%matplotlib inline\nplt.subplot(221)\nplt.imshow(X_train[0].reshape(32,32,3), cmap=plt.get_cmap('gray'))\nplt.grid('off')\nplt.subplot(222)\nplt.imshow(X_train[1].reshape(32,32,3), cmap=plt.get_cmap('gray'))\nplt.grid('off')\nplt.subplot(223)\nplt.imshow(X_train[2].reshape(32,32,3), cmap=plt.get_cmap('gray'))\nplt.grid('off')\nplt.subplot(224)\nplt.imshow(X_train[3].reshape(32,32,3), cmap=plt.get_cmap('gray'))\nplt.grid('off')\nplt.show()\ninput_shape = X_train[0].shape\n\nweight_decay = 1e-4\nmodel = Sequential()\nmodel.add(Conv2D(32, (3,3), padding='same', kernel_regularizer=regularizers.l2(weight_decay), input_shape=X_train.shape[1:]))\nmodel.add(Activation('relu'))\nmodel.add(BatchNormalization())\nmodel.add(Conv2D(32, (3,3), padding='same', kernel_regularizer=regularizers.l2(weight_decay)))\nmodel.add(Activation('relu'))\nmodel.add(BatchNormalization())\nmodel.add(MaxPooling2D(pool_size=(2,2)))\nmodel.add(Dropout(0.2))\n \nmodel.add(Conv2D(64, (3,3), padding='same', kernel_regularizer=regularizers.l2(weight_decay)))\nmodel.add(Activation('relu'))\nmodel.add(BatchNormalization())\nmodel.add(Conv2D(64, (3,3), padding='same', kernel_regularizer=regularizers.l2(weight_decay)))\nmodel.add(Activation('relu'))\nmodel.add(BatchNormalization())\nmodel.add(MaxPooling2D(pool_size=(2,2)))\nmodel.add(Dropout(0.3))\n \nmodel.add(Conv2D(128, (3,3), padding='same', kernel_regularizer=regularizers.l2(weight_decay)))\nmodel.add(Activation('relu'))\nmodel.add(BatchNormalization())\nmodel.add(Conv2D(128, (3,3), padding='same', kernel_regularizer=regularizers.l2(weight_decay)))\nmodel.add(Activation('relu'))\nmodel.add(BatchNormalization())\nmodel.add(MaxPooling2D(pool_size=(2,2)))\nmodel.add(Dropout(0.4))\n \nmodel.add(Flatten())\nmodel.add(Dense(10, activation='softmax'))\nfrom keras.optimizers import Adam\nadam = Adam(lr = 0.01)\nmodel.compile(loss='categorical_crossentropy', optimizer=adam,metrics=['accuracy'])\nhistory = model.fit(X_train, y_train, batch_size=32,epochs=10, verbose=1, validation_data=(X_val, y_val))\nhistory_dict = history.history\nloss_values = history_dict['loss']\nval_loss_values = history_dict['val_loss']\nacc_values = history_dict['acc']\nval_acc_values = history_dict['val_acc']\nepochs = range(1, len(val_loss_values) + 1)\n\nimport matplotlib.pyplot as plt\n%matplotlib inline\nplt.subplot(211)\nplt.plot(epochs, history.history['loss'], 'bo', label='Training loss')\nplt.plot(epochs, val_loss_values, 'r', label='Test loss')\nplt.title('Training and test loss without data augmentation')\nplt.xlabel('Epochs')\nplt.ylabel('Loss')\nplt.legend()\nplt.grid('off')\nplt.show()\nplt.subplot(212)\nplt.plot(epochs, history.history['acc'], 'bo', label='Training accuracy')\nplt.plot(epochs, val_acc_values, 'r', label='Test accuracy')\nplt.title('Training and test accuracy without data augmentation')\nplt.xlabel('Epochs')\nplt.ylabel('Accuracy')\nplt.gca().set_yticklabels(['{:.0f}%'.format(x*100) for x in plt.gca().get_yticks()]) \nplt.legend()\nplt.grid('off')\nplt.show()\n```\n\n# Data augmentation\n\n```\nfrom keras.preprocessing.image import ImageDataGenerator\ndatagen = ImageDataGenerator(\n rotation_range=15,\n width_shift_range=0.1,\n height_shift_range=0.1,\n horizontal_flip=True,\n )\ndatagen.fit(X_train)\nmodel = Sequential()\nmodel.add(Conv2D(32, (3,3), padding='same', kernel_regularizer=regularizers.l2(weight_decay), input_shape=X_train.shape[1:]))\nmodel.add(Activation('elu'))\nmodel.add(BatchNormalization())\nmodel.add(Conv2D(32, (3,3), padding='same', kernel_regularizer=regularizers.l2(weight_decay)))\nmodel.add(Activation('elu'))\nmodel.add(BatchNormalization())\nmodel.add(MaxPooling2D(pool_size=(2,2)))\nmodel.add(Dropout(0.2))\n \nmodel.add(Conv2D(64, (3,3), padding='same', kernel_regularizer=regularizers.l2(weight_decay)))\nmodel.add(Activation('elu'))\nmodel.add(BatchNormalization())\nmodel.add(Conv2D(64, (3,3), padding='same', kernel_regularizer=regularizers.l2(weight_decay)))\nmodel.add(Activation('elu'))\nmodel.add(BatchNormalization())\nmodel.add(MaxPooling2D(pool_size=(2,2)))\nmodel.add(Dropout(0.3))\n \nmodel.add(Conv2D(128, (3,3), padding='same', kernel_regularizer=regularizers.l2(weight_decay)))\nmodel.add(Activation('elu'))\nmodel.add(BatchNormalization())\nmodel.add(Conv2D(128, (3,3), padding='same', kernel_regularizer=regularizers.l2(weight_decay)))\nmodel.add(Activation('elu'))\nmodel.add(BatchNormalization())\nmodel.add(MaxPooling2D(pool_size=(2,2)))\nmodel.add(Dropout(0.4))\n \nmodel.add(Flatten())\nmodel.add(Dense(10, activation='softmax'))\nfrom keras.optimizers import Adam\nadam = Adam(lr = 0.01)\nmodel.compile(loss='categorical_crossentropy', optimizer='adam',metrics=['accuracy'])\n# Defining batch size\nbatch_size=32\nhistory = model.fit_generator(datagen.flow(X_train, y_train, batch_size=batch_size),steps_per_epoch=X_train.shape[0] // batch_size, epochs=10,validation_data=(X_val,y_val))\nhistory_dict = history.history\nloss_values = history_dict['loss']\nval_loss_values = history_dict['val_loss']\nacc_values = history_dict['acc']\nval_acc_values = history_dict['val_acc']\nepochs = range(1, len(val_loss_values) + 1)\n\nimport matplotlib.pyplot as plt\n%matplotlib inline\nplt.subplot(211)\nplt.plot(epochs, history.history['loss'], 'bo', label='Training loss')\nplt.plot(epochs, val_loss_values, 'r', label='Test loss')\nplt.title('Training and test loss with data augmentation')\nplt.xlabel('Epochs')\nplt.ylabel('Loss')\nplt.legend()\nplt.grid('off')\nplt.show()\nplt.subplot(212)\nplt.plot(epochs, history.history['acc'], 'bo', label='Training accuracy')\nplt.plot(epochs, val_acc_values, 'r', label='Test accuracy')\nplt.title('Training and test accuracy with data augmentation')\nplt.xlabel('Epochs')\nplt.ylabel('Accuracy')\nplt.gca().set_yticklabels(['{:.0f}%'.format(x*100) for x in plt.gca().get_yticks()]) \nplt.legend()\nplt.grid('off')\nplt.show()\n```\n\n" | github_jupyter |
```
# Copyright 2019 Google LLC
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Getting started: Training and prediction with Keras in AI Platform
<img src="https://storage.googleapis.com/cloud-samples-data/ai-platform/census/keras-tensorflow-cmle.png" alt="Keras, TensorFlow, and AI Platform logos" width="300px">
<table align="left">
<td>
<a href="https://colab.research.google.com/github/GoogleCloudPlatform/ai-platform-samples/blob/master/notebooks/samples/tensorflow/keras/getting_started_keras.ipynb">
<img src="https://cloud.google.com/ml-engine/images/colab-logo-32px.png" alt="Colab logo"> Run in Colab
</a>
</td>
<td>
<a href="https://github.com/GoogleCloudPlatform/ai-platform-samples/blob/master/notebooks/samples/tensorflow/keras/getting_started_keras.ipynb">
<img src="https://cloud.google.com/ml-engine/images/github-logo-32px.png" alt="GitHub logo">
View on GitHub
</a>
</td>
</table>
## Overview
This tutorial shows how to train a neural network on AI Platform
using the Keras sequential API and how to serve predictions from that
model.
Keras is a high-level API for building and training deep learning models.
[tf.keras](https://www.tensorflow.org/guide/keras) is TensorFlow’s
implementation of this API.
The first two parts of the tutorial walk through training a model on Cloud
AI Platform using prewritten Keras code, deploying the trained model to
AI Platform, and serving online predictions from the deployed model.
The last part of the tutorial digs into the training code used for this model and ensuring it's compatible with AI Platform. To learn more about building
machine learning models in Keras more generally, read [TensorFlow's Keras
tutorials](https://www.tensorflow.org/tutorials/keras).
### Dataset
This tutorial uses the [United States Census Income
Dataset](https://archive.ics.uci.edu/ml/datasets/census+income) provided by the
[UC Irvine Machine Learning
Repository](https://archive.ics.uci.edu/ml/index.php). This dataset contains
information about people from a 1994 Census database, including age, education,
marital status, occupation, and whether they make more than $50,000 a year.
### Objective
The goal is to train a deep neural network (DNN) using Keras that predicts
whether a person makes more than $50,000 a year (target label) based on other
Census information about the person (features).
This tutorial focuses more on using this model with AI Platform than on
the design of the model itself. However, it's always important to think about
potential problems and unintended consequences when building machine learning
systems. See the [Machine Learning Crash Course exercise about
fairness](https://developers.google.com/machine-learning/crash-course/fairness/programming-exercise)
to learn about sources of bias in the Census dataset, as well as machine
learning fairness more generally.
### Costs
This tutorial uses billable components of Google Cloud Platform (GCP):
* AI Platform
* Cloud Storage
Learn about [AI Platform
pricing](https://cloud.google.com/ml-engine/docs/pricing) and [Cloud Storage
pricing](https://cloud.google.com/storage/pricing), and use the [Pricing
Calculator](https://cloud.google.com/products/calculator/)
to generate a cost estimate based on your projected usage.
## Before you begin
You must do several things before you can train and deploy a model in
AI Platform:
* Set up your local development environment.
* Set up a GCP project with billing and the necessary
APIs enabled.
* Authenticate your GCP account in this notebook.
* Create a Cloud Storage bucket to store your training package and your
trained model.
### Set up your local development environment
**If you are using Colab or AI Platform Notebooks**, your environment already meets
all the requirements to run this notebook. You can skip this step.
**Otherwise**, make sure your environment meets this notebook's requirements.
You need the following:
* The Google Cloud SDK
* Git
* Python 3
* virtualenv
* Jupyter notebook running in a virtual environment with Python 3
The Google Cloud guide to [Setting up a Python development
environment](https://cloud.google.com/python/setup) and the [Jupyter
installation guide](https://jupyter.org/install) provide detailed instructions
for meeting these requirements. The following steps provide a condensed set of
instructions:
1. [Install and initialize the Cloud SDK.](https://cloud.google.com/sdk/docs/)
2. [Install Python 3.](https://cloud.google.com/python/setup#installing_python)
3. [Install
virtualenv](https://cloud.google.com/python/setup#installing_and_using_virtualenv)
and create a virtual environment that uses Python 3.
4. Activate that environment and run `pip install jupyter` in a shell to install
Jupyter.
5. Run `jupyter notebook` in a shell to launch Jupyter.
6. Open this notebook in the Jupyter Notebook Dashboard.
### Set up your GCP project
**The following steps are required, regardless of your notebook environment.**
1. [Select or create a GCP project.](https://console.cloud.google.com/cloud-resource-manager)
2. [Make sure that billing is enabled for your project.](https://cloud.google.com/billing/docs/how-to/modify-project)
3. [Enable the AI Platform ("Cloud Machine Learning Engine") and Compute Engine APIs.](https://console.cloud.google.com/flows/enableapi?apiid=ml.googleapis.com,compute_component)
4. Enter your project ID in the cell below. Then run the cell to make sure the
Cloud SDK uses the right project for all the commands in this notebook.
**Note**: Jupyter runs lines prefixed with `!` as shell commands, and it interpolates Python variables prefixed with `$` into these commands.
```
PROJECT_ID = '[your-project-id]' #@param {type:"string"}
! gcloud config set project $PROJECT_ID
```
### Authenticate your GCP account
**If you are using AI Platform Notebooks**, your environment is already
authenticated. Skip this step.
**If you are using Colab**, run the cell below and follow the instructions
when prompted to authenticate your account via oAuth.
**Otherwise**, follow these steps:
1. In the GCP Console, go to the [**Create service account key**
page](https://console.cloud.google.com/apis/credentials/serviceaccountkey).
2. From the **Service account** drop-down list, select **New service account**.
3. In the **Service account name** field, enter a name.
4. From the **Role** drop-down list, select
**Machine Learning Engine > AI Platform Admin** and
**Storage > Storage Object Admin**.
5. Click *Create*. A JSON file that contains your key downloads to your
local environment.
6. Enter the path to your service account key as the
`GOOGLE_APPLICATION_CREDENTIALS` variable in the cell below and run the cell.
```
import sys
# If you are running this notebook in Colab, run this cell and follow the
# instructions to authenticate your GCP account. This provides access to your
# Cloud Storage bucket and lets you submit training jobs and prediction
# requests.
if 'google.colab' in sys.modules:
from google.colab import auth as google_auth
google_auth.authenticate_user()
# If you are running this notebook locally, replace the string below with the
# path to your service account key and run this cell to authenticate your GCP
# account.
else:
%env GOOGLE_APPLICATION_CREDENTIALS ''
```
### Create a Cloud Storage bucket
**The following steps are required, regardless of your notebook environment.**
When you submit a training job using the Cloud SDK, you upload a Python package
containing your training code to a Cloud Storage bucket. AI Platform runs
the code from this package. In this tutorial, AI Platform also saves the
trained model that results from your job in the same bucket. You can then
create an AI Platform model version based on this output in order to serve
online predictions.
Set the name of your Cloud Storage bucket below. It must be unique across all
Cloud Storage buckets.
You may also change the `REGION` variable, which is used for operations
throughout the rest of this notebook. Make sure to [choose a region where Cloud
AI Platform services are
available](https://cloud.google.com/ml-engine/docs/tensorflow/regions).
```
BUCKET_NAME = '[your-bucket-name]' #@param {type:"string"}
REGION = 'us-central1' #@param {type:"string"}
```
**Only if your bucket doesn't already exist**: Run the following cell to create your Cloud Storage bucket.
```
! gsutil mb -l $REGION gs://$BUCKET_NAME
```
Finally, validate access to your Cloud Storage bucket by examining its contents:
```
! gsutil ls -al gs://$BUCKET_NAME
```
## Part 1. Quickstart for training in AI Platform
This section of the tutorial walks you through submitting a training job to Cloud
AI Platform. This job runs sample code that uses Keras to train a deep neural
network on the United States Census data. It outputs the trained model as a
[TensorFlow SavedModel
directory](https://www.tensorflow.org/guide/saved_model#save_and_restore_models)
in your Cloud Storage bucket.
### Get training code and dependencies
First, download the training code and change the notebook's working directory:
```
# Clone the repository of AI Platform samples
! git clone --depth 1 https://github.com/GoogleCloudPlatform/ai-platform-samples
# Set the working directory to the sample code directory
%cd ai-platform-samples/training/tensorflow/census/tf-keras
```
Notice that the training code is structured as a Python package in the
`trainer/` subdirectory:
```
# `ls` shows the working directory's contents. The `p` flag adds trailing
# slashes to subdirectory names. The `R` flag lists subdirectories recursively.
! ls -pR
```
Run the following cell to install Python dependencies needed to train the model locally. When you run the training job in AI Platform,
dependencies are preinstalled based on the [runtime
version](https://cloud.google.com/ml-engine/docs/tensorflow/runtime-version-list)
you choose.
```
! pip install -r requirements.txt --user
```
### Train your model locally
Before training on AI Platform, train the job locally to verify the file
structure and packaging is correct.
For a complex or resource-intensive job, you
may want to train locally on a small sample of your dataset to verify your code.
Then you can run the job on AI Platform to train on the whole dataset.
This sample runs a relatively quick job on a small dataset, so the local
training and the AI Platform job run the same code on the same data.
Run the following cell to train a model locally:
```
# Explicitly tell `gcloud ai-platform local train` to use Python 3
! gcloud config set ml_engine/local_python $(which python3)
# This is similar to `python -m trainer.task --job-dir local-training-output`
# but it better replicates the AI Platform environment, especially for
# distributed training (not applicable here).
! gcloud ai-platform local train \
--package-path trainer \
--module-name trainer.task \
--job-dir local-training-output
```
### Train your model using AI Platform
Next, submit a training job to AI Platform. This runs the training module
in the cloud and exports the trained model to Cloud Storage.
First, give your training job a name and choose a directory within your Cloud
Storage bucket for saving intermediate and output files:
```
import uuid
JOB_NAME = "my_first_keras_job_" + uuid.uuid4().hex[:10]
JOB_DIR = 'gs://' + BUCKET_NAME + '/keras-job-dir'
```
Run the following command to package the `trainer/` directory, upload it to the
specified `--job-dir`, and instruct AI Platform to run the
`trainer.task` module from that package.
The `--stream-logs` flag lets you view training logs in the cell below. You can
also see logs and other job details in the GCP Console.
### Hyperparameter tuning
You can optionally perform hyperparameter tuning by using the included
`hptuning_config.yaml` configuration file. This file tells AI Platform to tune the batch size and learning rate for training over multiple trials to maximize accuracy.
In this example, the training code uses a [TensorBoard
callback](https://www.tensorflow.org/api_docs/python/tf/keras/callbacks/TensorBoard),
which [creates TensorFlow `Summary`
`Event`s](https://www.tensorflow.org/api_docs/python/tf/summary/FileWriter#add_summary)
during training. AI Platform uses these events to track the metric you want to
optimize. Learn more about [hyperparameter tuning in
AI Platform Training](https://cloud.google.com/ml-engine/docs/tensorflow/hyperparameter-tuning-overview).
```
! gcloud ai-platform jobs submit training $JOB_NAME \
--package-path trainer/ \
--module-name trainer.task \
--region $REGION \
--python-version 3.7 \
--runtime-version 2.1 \
--job-dir $JOB_DIR \
--stream-logs
```
## Part 2. Quickstart for online predictions in AI Platform
This section shows how to use AI Platform and your trained model from Part 1
to predict a person's income bracket from other Census information about them.
### Create model and version resources in AI Platform
To serve online predictions using the model you trained and exported in Part 1,
create a *model* resource in AI Platform and a *version* resource
within it. The version resource is what actually uses your trained model to
serve predictions. This structure lets you adjust and retrain your model many times and
organize all the versions together in AI Platform. Learn more about [models
and
versions](https://cloud.google.com/ml-engine/docs/tensorflow/projects-models-versions-jobs).
First, name and create the model resource:
```
MODEL_NAME = 'my_first_keras_model'
MODEL_VERSION = 'v1'
# Delete model version resource
! gcloud ai-platform versions delete $MODEL_VERSION --quiet --model $MODEL_NAME
# Delete model resource
! gcloud ai-platform models delete $MODEL_NAME --quiet
# Create Model
! gcloud ai-platform models create $MODEL_NAME --regions $REGION
```
Next, create the model version. The training job from Part 1 exported a timestamped
[TensorFlow SavedModel
directory](https://www.tensorflow.org/guide/saved_model#structure_of_a_savedmodel_directory)
to your Cloud Storage bucket. AI Platform uses this directory to create a
model version. Learn more about [SavedModel and
AI Platform](https://cloud.google.com/ml-engine/docs/tensorflow/deploying-models).
You may be able to find the path to this directory in your training job's logs.
Look for a line like:
```
Model exported to: gs://<your-bucket-name>/keras-job-dir/keras_export/1545439782
```
Execute the following command to identify your SavedModel directory and use it to create a model version resource:
```
# Get a list of directories in the `keras_export` parent directory
! gsutil ls $JOB_DIR/keras_export/
# Pick the directory with the latest timestamp, in case you've trained
# multiple times
SAVED_MODEL_PATH = 'gs://{}/keras-job-dir/keras_export/'.format(BUCKET_NAME)
# Create model version based on that SavedModel directory
! gcloud ai-platform versions create $MODEL_VERSION \
--model $MODEL_NAME \
--runtime-version 2.1 \
--python-version 3.7 \
--framework tensorflow \
--origin $SAVED_MODEL_PATH
```
### Prepare input for prediction
To receive valid and useful predictions, you must preprocess input for prediction in the same way that training data was preprocessed. In a production
system, you may want to create a preprocessing pipeline that can be used identically at training time and prediction time.
For this exercise, use the training package's data-loading code to select a random sample from the evaluation data. This data is in the form that was used to evaluate accuracy after each epoch of training, so it can be used to send test predictions without further preprocessing:
```
from trainer import util
_, _, eval_x, eval_y = util.load_data()
prediction_input = eval_x.sample(20)
prediction_targets = eval_y[prediction_input.index]
prediction_input
```
Notice that categorical fields, like `occupation`, have already been converted to integers (with the same mapping that was used for training). Numerical fields, like `age`, have been scaled to a
[z-score](https://developers.google.com/machine-learning/crash-course/representation/cleaning-data). Some fields have been dropped from the original
data. Compare the prediction input with the raw data for the same examples:
```
import pandas as pd
_, eval_file_path = util.download(util.DATA_DIR)
raw_eval_data = pd.read_csv(eval_file_path,
names=util._CSV_COLUMNS,
na_values='?')
raw_eval_data.iloc[prediction_input.index]
```
Export the prediction input to a newline-delimited JSON file:
```
import json
with open('prediction_input.json', 'w') as json_file:
for row in prediction_input.values.tolist():
json.dump(row, json_file)
json_file.write('\n')
! cat prediction_input.json
```
The `gcloud` command-line tool accepts newline-delimited JSON for online
prediction, and this particular Keras model expects a flat list of
numbers for each input example.
AI Platform requires a different format when you make online prediction requests to the REST API without using the `gcloud` tool. The way you structure
your model may also change how you must format data for prediction. Learn more
about [formatting data for online
prediction](https://cloud.google.com/ml-engine/docs/tensorflow/prediction-overview#prediction_input_data).
### Submit the online prediction request
Use `gcloud` to submit your online prediction request.
```
! gcloud ai-platform predict \
--model $MODEL_NAME \
--version $MODEL_VERSION \
--json-instances prediction_input.json
```
Since the model's last layer uses a [sigmoid function](https://developers.google.com/machine-learning/glossary/#sigmoid_function) for its activation, outputs between 0 and 0.5 represent negative predictions ("<=50K") and outputs between 0.5 and 1 represent positive ones (">50K").
Do the predicted income brackets match the actual ones? Run the following cell
to see the true labels.
```
prediction_targets
```
## Part 3. Developing the Keras model from scratch
At this point, you have trained a machine learning model on AI Platform, deployed the trained model as a version resource on AI Platform, and received online predictions from the deployment. The next section walks through recreating the Keras code used to train your model. It covers the following parts of developing a machine learning model for use with AI Platform:
* Downloading and preprocessing data
* Designing and training the model
* Visualizing training and exporting the trained model
While this section provides more detailed insight to the tasks completed in previous parts, to learn more about using `tf.keras`, read [TensorFlow's guide to Keras](https://www.tensorflow.org/tutorials/keras). To learn more about structuring code as a training packge for AI Platform, read [Packaging a training application](https://cloud.google.com/ml-engine/docs/tensorflow/packaging-trainer) and reference the [complete training code](https://github.com/GoogleCloudPlatform/ai-platform-samples/tree/master/training/tensorflow/census/tf-keras), which is structured as a Python package.
### Import libraries and define constants
First, import Python libraries required for training:
```
import os
from six.moves import urllib
import tempfile
import numpy as np
import pandas as pd
import tensorflow as tf
# Examine software versions
print(__import__('sys').version)
print(tf.__version__)
print(tf.keras.__version__)
```
Then, define some useful constants:
* Information for downloading training and evaluation data
* Information required for Pandas to interpret the data and convert categorical fields into numeric features
* Hyperparameters for training, such as learning rate and batch size
```
### For downloading data ###
# Storage directory
DATA_DIR = os.path.join(tempfile.gettempdir(), 'census_data')
# Download options.
DATA_URL = 'https://storage.googleapis.com/cloud-samples-data/ai-platform' \
'/census/data'
TRAINING_FILE = 'adult.data.csv'
EVAL_FILE = 'adult.test.csv'
TRAINING_URL = '%s/%s' % (DATA_URL, TRAINING_FILE)
EVAL_URL = '%s/%s' % (DATA_URL, EVAL_FILE)
### For interpreting data ###
# These are the features in the dataset.
# Dataset information: https://archive.ics.uci.edu/ml/datasets/census+income
_CSV_COLUMNS = [
'age', 'workclass', 'fnlwgt', 'education', 'education_num',
'marital_status', 'occupation', 'relationship', 'race', 'gender',
'capital_gain', 'capital_loss', 'hours_per_week', 'native_country',
'income_bracket'
]
_CATEGORICAL_TYPES = {
'workclass': pd.api.types.CategoricalDtype(categories=[
'Federal-gov', 'Local-gov', 'Never-worked', 'Private', 'Self-emp-inc',
'Self-emp-not-inc', 'State-gov', 'Without-pay'
]),
'marital_status': pd.api.types.CategoricalDtype(categories=[
'Divorced', 'Married-AF-spouse', 'Married-civ-spouse',
'Married-spouse-absent', 'Never-married', 'Separated', 'Widowed'
]),
'occupation': pd.api.types.CategoricalDtype([
'Adm-clerical', 'Armed-Forces', 'Craft-repair', 'Exec-managerial',
'Farming-fishing', 'Handlers-cleaners', 'Machine-op-inspct',
'Other-service', 'Priv-house-serv', 'Prof-specialty', 'Protective-serv',
'Sales', 'Tech-support', 'Transport-moving'
]),
'relationship': pd.api.types.CategoricalDtype(categories=[
'Husband', 'Not-in-family', 'Other-relative', 'Own-child', 'Unmarried',
'Wife'
]),
'race': pd.api.types.CategoricalDtype(categories=[
'Amer-Indian-Eskimo', 'Asian-Pac-Islander', 'Black', 'Other', 'White'
]),
'native_country': pd.api.types.CategoricalDtype(categories=[
'Cambodia', 'Canada', 'China', 'Columbia', 'Cuba', 'Dominican-Republic',
'Ecuador', 'El-Salvador', 'England', 'France', 'Germany', 'Greece',
'Guatemala', 'Haiti', 'Holand-Netherlands', 'Honduras', 'Hong', 'Hungary',
'India', 'Iran', 'Ireland', 'Italy', 'Jamaica', 'Japan', 'Laos', 'Mexico',
'Nicaragua', 'Outlying-US(Guam-USVI-etc)', 'Peru', 'Philippines', 'Poland',
'Portugal', 'Puerto-Rico', 'Scotland', 'South', 'Taiwan', 'Thailand',
'Trinadad&Tobago', 'United-States', 'Vietnam', 'Yugoslavia'
]),
'income_bracket': pd.api.types.CategoricalDtype(categories=[
'<=50K', '>50K'
])
}
# This is the label (target) we want to predict.
_LABEL_COLUMN = 'income_bracket'
### Hyperparameters for training ###
# This the training batch size
BATCH_SIZE = 128
# This is the number of epochs (passes over the full training data)
NUM_EPOCHS = 20
# Define learning rate.
LEARNING_RATE = .01
```
### Download and preprocess data
#### Download the data
Next, define functions to download training and evaluation data. These functions also fix minor irregularities in the data's formatting.
```
def _download_and_clean_file(filename, url):
"""Downloads data from url, and makes changes to match the CSV format.
The CSVs may use spaces after the comma delimters (non-standard) or include
rows which do not represent well-formed examples. This function strips out
some of these problems.
Args:
filename: filename to save url to
url: URL of resource to download
"""
temp_file, _ = urllib.request.urlretrieve(url)
with tf.io.gfile.GFile(temp_file, 'r') as temp_file_object:
with tf.io.gfile.GFile(filename, 'w') as file_object:
for line in temp_file_object:
line = line.strip()
line = line.replace(', ', ',')
if not line or ',' not in line:
continue
if line[-1] == '.':
line = line[:-1]
line += '\n'
file_object.write(line)
tf.io.gfile.remove(temp_file)
def download(data_dir):
"""Downloads census data if it is not already present.
Args:
data_dir: directory where we will access/save the census data
"""
tf.io.gfile.makedirs(data_dir)
training_file_path = os.path.join(data_dir, TRAINING_FILE)
if not tf.io.gfile.exists(training_file_path):
_download_and_clean_file(training_file_path, TRAINING_URL)
eval_file_path = os.path.join(data_dir, EVAL_FILE)
if not tf.io.gfile.exists(eval_file_path):
_download_and_clean_file(eval_file_path, EVAL_URL)
return training_file_path, eval_file_path
```
Use those functions to download the data for training and verify that you have CSV files for training and evaluation:
```
training_file_path, eval_file_path = download(DATA_DIR)
# You should see 2 files: adult.data.csv and adult.test.csv
!ls -l $DATA_DIR
```
Next, load these files using Pandas and examine the data:
```
# This census data uses the value '?' for fields (column) that are missing data.
# We use na_values to find ? and set it to NaN values.
# https://pandas.pydata.org/pandas-docs/stable/generated/pandas.read_csv.html
train_df = pd.read_csv(training_file_path, names=_CSV_COLUMNS, na_values='?')
eval_df = pd.read_csv(eval_file_path, names=_CSV_COLUMNS, na_values='?')
# Here's what the data looks like before we preprocess the data.
train_df.head()
```
#### Preprocess the data
The first preprocessing step removes certain features from the data and
converts categorical features to numerical values for use with Keras.
Learn more about [feature engineering](https://developers.google.com/machine-learning/crash-course/representation/feature-engineering) and [bias in data](https://developers.google.com/machine-learning/crash-course/fairness/types-of-bias).
```
UNUSED_COLUMNS = ['fnlwgt', 'education', 'gender']
def preprocess(dataframe):
"""Converts categorical features to numeric. Removes unused columns.
Args:
dataframe: Pandas dataframe with raw data
Returns:
Dataframe with preprocessed data
"""
dataframe = dataframe.drop(columns=UNUSED_COLUMNS)
# Convert integer valued (numeric) columns to floating point
numeric_columns = dataframe.select_dtypes(['int64']).columns
dataframe[numeric_columns] = dataframe[numeric_columns].astype('float32')
# Convert categorical columns to numeric
cat_columns = dataframe.select_dtypes(['object']).columns
dataframe[cat_columns] = dataframe[cat_columns].apply(lambda x: x.astype(
_CATEGORICAL_TYPES[x.name]))
dataframe[cat_columns] = dataframe[cat_columns].apply(lambda x: x.cat.codes)
return dataframe
prepped_train_df = preprocess(train_df)
prepped_eval_df = preprocess(eval_df)
```
Run the following cell to see how preprocessing changed the data. Notice in particular that `income_bracket`, the label that you're training the model to predict, has changed from `<=50K` and `>50K` to `0` and `1`:
```
prepped_train_df.head()
```
Next, separate the data into features ("x") and labels ("y"), and reshape the label arrays into a format for use with `tf.data.Dataset` later:
```
# Split train and test data with labels.
# The pop() method will extract (copy) and remove the label column from the dataframe
train_x, train_y = prepped_train_df, prepped_train_df.pop(_LABEL_COLUMN)
eval_x, eval_y = prepped_eval_df, prepped_eval_df.pop(_LABEL_COLUMN)
# Reshape label columns for use with tf.data.Dataset
train_y = np.asarray(train_y).astype('float32').reshape((-1, 1))
eval_y = np.asarray(eval_y).astype('float32').reshape((-1, 1))
```
Scaling training data so each numerical feature column has a mean of 0 and a standard deviation of 1 [can improve your model](https://developers.google.com/machine-learning/crash-course/representation/cleaning-data).
In a production system, you may want to save the means and standard deviations from your training set and use them to perform an identical transformation on test data at prediction time. For convenience in this exercise, temporarily combine the training and evaluation data to scale all of them:
```
def standardize(dataframe):
"""Scales numerical columns using their means and standard deviation to get
z-scores: the mean of each numerical column becomes 0, and the standard
deviation becomes 1. This can help the model converge during training.
Args:
dataframe: Pandas dataframe
Returns:
Input dataframe with the numerical columns scaled to z-scores
"""
dtypes = list(zip(dataframe.dtypes.index, map(str, dataframe.dtypes)))
# Normalize numeric columns.
for column, dtype in dtypes:
if dtype == 'float32':
dataframe[column] -= dataframe[column].mean()
dataframe[column] /= dataframe[column].std()
return dataframe
# Join train_x and eval_x to normalize on overall means and standard
# deviations. Then separate them again.
all_x = pd.concat([train_x, eval_x], keys=['train', 'eval'])
all_x = standardize(all_x)
train_x, eval_x = all_x.xs('train'), all_x.xs('eval')
```
Finally, examine some of your fully preprocessed training data:
```
# Verify dataset features
# Note how only the numeric fields (not categorical) have been standardized
train_x.head()
```
### Design and train the model
#### Create training and validation datasets
Create an input function to convert features and labels into a
[`tf.data.Dataset`](https://www.tensorflow.org/guide/datasets) for training or evaluation:
```
def input_fn(features, labels, shuffle, num_epochs, batch_size):
"""Generates an input function to be used for model training.
Args:
features: numpy array of features used for training or inference
labels: numpy array of labels for each example
shuffle: boolean for whether to shuffle the data or not (set True for
training, False for evaluation)
num_epochs: number of epochs to provide the data for
batch_size: batch size for training
Returns:
A tf.data.Dataset that can provide data to the Keras model for training or
evaluation
"""
if labels is None:
inputs = features
else:
inputs = (features, labels)
dataset = tf.data.Dataset.from_tensor_slices(inputs)
if shuffle:
dataset = dataset.shuffle(buffer_size=len(features))
# We call repeat after shuffling, rather than before, to prevent separate
# epochs from blending together.
dataset = dataset.repeat(num_epochs)
dataset = dataset.batch(batch_size)
return dataset
```
Next, create these training and evaluation datasets.Use the `NUM_EPOCHS`
and `BATCH_SIZE` hyperparameters defined previously to define how the training
dataset provides examples to the model during training. Set up the validation
dataset to provide all its examples in one batch, for a single validation step
at the end of each training epoch.
```
# Pass a numpy array by using DataFrame.values
training_dataset = input_fn(features=train_x.values,
labels=train_y,
shuffle=True,
num_epochs=NUM_EPOCHS,
batch_size=BATCH_SIZE)
num_eval_examples = eval_x.shape[0]
# Pass a numpy array by using DataFrame.values
validation_dataset = input_fn(features=eval_x.values,
labels=eval_y,
shuffle=False,
num_epochs=NUM_EPOCHS,
batch_size=num_eval_examples)
```
#### Design a Keras Model
Design your neural network using the [Keras Sequential API](https://www.tensorflow.org/guide/keras#sequential_model).
This deep neural network (DNN) has several hidden layers, and the last layer uses a sigmoid activation function to output a value between 0 and 1:
* The input layer has 100 units using the ReLU activation function.
* The hidden layer has 75 units using the ReLU activation function.
* The hidden layer has 50 units using the ReLU activation function.
* The hidden layer has 25 units using the ReLU activation function.
* The output layer has 1 units using a sigmoid activation function.
* The optimizer uses the binary cross-entropy loss function, which is appropriate for a binary classification problem like this one.
Feel free to change these layers to try to improve the model:
```
def create_keras_model(input_dim, learning_rate):
"""Creates Keras Model for Binary Classification.
Args:
input_dim: How many features the input has
learning_rate: Learning rate for training
Returns:
The compiled Keras model (still needs to be trained)
"""
Dense = tf.keras.layers.Dense
model = tf.keras.Sequential(
[
Dense(100, activation=tf.nn.relu, kernel_initializer='uniform',
input_shape=(input_dim,)),
Dense(75, activation=tf.nn.relu),
Dense(50, activation=tf.nn.relu),
Dense(25, activation=tf.nn.relu),
Dense(1, activation=tf.nn.sigmoid)
])
# Custom Optimizer:
# https://www.tensorflow.org/api_docs/python/tf/train/RMSPropOptimizer
optimizer = tf.keras.optimizers.RMSprop(
lr=learning_rate)
# Compile Keras model
model.compile(
loss='binary_crossentropy', optimizer=optimizer, metrics=['accuracy'])
return model
```
Next, create the Keras model object and examine its structure:
```
num_train_examples, input_dim = train_x.shape
print('Number of features: {}'.format(input_dim))
print('Number of examples: {}'.format(num_train_examples))
keras_model = create_keras_model(
input_dim=input_dim,
learning_rate=LEARNING_RATE)
# Take a detailed look inside the model
keras_model.summary()
```
#### Train and evaluate the model
Define a learning rate decay to encourage model paramaters to make smaller
changes as training goes on:
```
class CustomCallback(tf.keras.callbacks.TensorBoard):
"""Callback to write out a custom metric used by CAIP for HP Tuning."""
def on_epoch_end(self, epoch, logs=None): # pylint: disable=no-self-use
"""Write tf.summary.scalar on epoch end."""
tf.summary.scalar('epoch_accuracy', logs['accuracy'], epoch)
# Setup TensorBoard callback.
tensorboard_cb = CustomCallback(os.path.join(JOB_DIR, 'keras_tensorboard'),
histogram_freq=1)
# Setup Learning Rate decay.
lr_decay_cb = tf.keras.callbacks.LearningRateScheduler(
lambda epoch: LEARNING_RATE + 0.02 * (0.5 ** (1 + epoch)),
verbose=True)
```
Finally, train the model. Provide the appropriate `steps_per_epoch` for the
model to train on the entire training dataset (with `BATCH_SIZE` examples per step) during each epoch. And instruct the model to calculate validation
accuracy with one big validation batch at the end of each epoch.
```
history = keras_model.fit(training_dataset,
epochs=NUM_EPOCHS,
steps_per_epoch=int(num_train_examples/BATCH_SIZE),
validation_data=validation_dataset,
validation_steps=1,
callbacks=[tensorboard_cb, lr_decay_cb],
verbose=1)
```
### Visualize training and export the trained model
#### Visualize training
Import `matplotlib` to visualize how the model learned over the training period.
```
from matplotlib import pyplot as plt
%matplotlib inline
```
Plot the model's loss (binary cross-entropy) and accuracy, as measured at the
end of each training epoch:
```
# Visualize History for Loss.
plt.title('Keras model loss')
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['training', 'validation'], loc='upper right')
plt.show()
# Visualize History for Accuracy.
plt.title('Keras model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.plot(history.history['accuracy'])
plt.plot(history.history['val_accuracy'])
plt.legend(['training', 'validation'], loc='lower right')
plt.show()
```
Over time, loss decreases and accuracy increases. But do they converge to a
stable level? Are there big differences between the training and validation
metrics (a sign of overfitting)?
Learn about [how to improve your machine learning
model](https://developers.google.com/machine-learning/crash-course/). Then, feel
free to adjust hyperparameters or the model architecture and train again.
#### Export the model for serving
AI Platform requires when you [create a model version
resource](https://cloud.google.com/ml-engine/docs/tensorflow/deploying-models#create_a_model_version).
Since not all optimizers can be exported to the SavedModel format, you may see
warnings during the export process. As long you successfully export a serving
graph, AI Platform can used the SavedModel to serve predictions.
```
# Export the model to a local SavedModel directory
tf.keras.models.save_model(keras_model, 'keras_export')
```
You may export a SavedModel directory to your local filesystem or to Cloud
Storage, as long as you have the necessary permissions. In your current
environment, you granted access to Cloud Storage by authenticating your GCP account and setting the `GOOGLE_APPLICATION_CREDENTIALS` environment variable.
AI Platform training jobs can also export directly to Cloud Storage, because
AI Platform service accounts [have access to Cloud Storage buckets in their own
project](https://cloud.google.com/ml-engine/docs/tensorflow/working-with-cloud-storage).
Try exporting directly to Cloud Storage:
```
# Export the model to a SavedModel directory in Cloud Storage
export_path = os.path.join(JOB_DIR, 'keras_export')
tf.keras.models.save_model(keras_model, export_path)
print('Model exported to: ', export_path)
```
You can now deploy this model to AI Platform and serve predictions by
following the steps from Part 2.
## Cleaning up
To clean up all GCP resources used in this project, you can [delete the GCP
project](https://cloud.google.com/resource-manager/docs/creating-managing-projects#shutting_down_projects) you used for the tutorial.
Alternatively, you can clean up individual resources by running the following
commands:
```
# Delete model version resource
! gcloud ai-platform versions delete $MODEL_VERSION --quiet --model $MODEL_NAME
# Delete model resource
! gcloud ai-platform models delete $MODEL_NAME --quiet
# Delete Cloud Storage objects that were created
! gsutil -m rm -r $JOB_DIR
# If the training job is still running, cancel it
! gcloud ai-platform jobs cancel $JOB_NAME --quiet --verbosity critical
```
If your Cloud Storage bucket doesn't contain any other objects and you would like to delete it, run `gsutil rm -r gs://$BUCKET_NAME`.
## What's next?
* View the [complete training
code](https://github.com/GoogleCloudPlatform/ai-platform-samples/tree/master/training/tensorflow/census/tf-keras) used in this guide, which structures the code to accept custom
hyperparameters as command-line flags.
* Read about [packaging
code](https://cloud.google.com/ml-engine/docs/tensorflow/packaging-trainer) for an AI Platform training job.
* Read about [deploying a
model](https://cloud.google.com/ml-engine/docs/tensorflow/deploying-models) to serve predictions.
| github_jupyter |
# 1. Quick start: read csv and flatten json fields + smart dump
Hi! This notebook is a derivative of https://www.kaggle.com/ogrellier/create-extracted-json-fields-dataset. I also tried to use [this kernel by julian3833](https://www.kaggle.com/julian3833/1-quick-start-read-csv-and-flatten-json-fields), but failed to execute `json_normalise` on a dask DataFrame. It extends the original code by using `dask.DataFrame`, see the docs [here](https://docs.dask.org/en/latest/dataframe.html). This allows to process data **with pandas-like interface in parallel threads and in chunks**. This allows to run faster and to work around the RAM limit.
# Main goals
1. **Process dataset, that can not fit into memory.**
2. **Use dask toolkit that allows to scale data processing to a cluster instead of a single core.**
3. **Store pre-processed flat data**
The output is stored in gziped csv file to reduce the file size.
```
import os
import json
import numpy as np
import pandas as pd
from pandas.io.json import json_normalize
import pyarrow as pa
import dask
import dask.dataframe as dd
# Set up a logger to dump messages to both log file and notebook
import logging as logging
def ini_log(filename):
logger = logging.getLogger(__name__)
logger.setLevel(logging.DEBUG)
handlers = [logging.StreamHandler(None), logging.FileHandler(filename, 'a')]
fmt=logging.Formatter('%(asctime)-15s: %(levelname)s %(message)s')
for h in handlers:
h.setFormatter(fmt)
logger.addHandler(h)
return logger
log = ini_log('out.log')
#log.basicConfig(filename='out.log',level=log.DEBUG, format='%(asctime)-15s: %(levelname)s %(message)s')
import gc
gc.enable()
```
The original functions (from the aforementioned kernel) with updates to run with `dask`
```
def_num = np.nan
def_str = 'NaN'
def get_keys_for_field(field=None):
the_dict = {
'device': [
'browser', 'object',
'deviceCategory',
('isMobile', False, bool),
'operatingSystem'
],
'geoNetwork': [
'city',
'continent',
'country',
'metro',
'networkDomain',
'region',
'subContinent'
],
'totals': [
('pageviews', 0, np.int16),
('hits', def_num, np.int16),
('bounces', 0, np.int8),
('newVisits', 0, np.int16),
('transactionRevenue', 0, np.int64),
('visits', -1, np.int16),
('timeOnSite', -1, np.int32),
('sessionQualityDim', -1, np.int8),
],
'trafficSource': [
'adContent',
#'adwordsClickInfo',
'campaign',
('isTrueDirect', False, bool),
#'keyword', #can not be saved in train (utf-8 symbols left)
'medium',
'referralPath',
'source'
],
}
return the_dict[field]
def convert_to_dict(x):
#print(x, type(x))
return eval(x.replace('false', 'False')
.replace('true', 'True')
.replace('null', 'np.nan'))
def develop_json_fields(fin, json_fields=['totals'], bsize=1e8, cols_2drop=[]):
df = dd.read_csv(fin, blocksize=bsize,
#converters={column: json.loads for column in JSON_COLUMNS},
dtype={'fullVisitorId': 'str', # Important!!
#usecols=lambda c: c not in cols_2drop,
'date': 'str',
**{c: 'str' for c in json_fields}
},
parse_dates=['date'],)#.head(10000, 100)
df = df.drop(cols_2drop, axis=1)
# Get the keys
for json_field in json_fields:
log.info('Doing Field {}'.format(json_field))
# Get json field keys to create columns
the_keys = get_keys_for_field(json_field)
# Replace the string by a dict
log.info('Transform string to dict')
df[json_field] = df[json_field].apply(lambda x: convert_to_dict(x), meta=('','object'))
log.info('{} converted to dict'.format(json_field))
#display(df.head())
for k in the_keys:
if isinstance(k, str):
t_ = def_str
k_ = k
else:
t_ = k[1]
k_ = k[0]
df[json_field + '_' + k_] = df[json_field].to_bag().pluck(k_, default=t_).to_dataframe().iloc[:,0]
if not isinstance(k, str) and len(k)>2:
df[json_field + '_' + k_] = df[json_field + '_' + k_].astype(k[2])
del df[json_field]
gc.collect()
log.info('{} fields extracted'.format(json_field))
return df
print(os.listdir("../input"))
!head ../input/train_v2.csv
```
## Let's load the original data with pre-processing
```
JSON_COLUMNS = ['device', 'geoNetwork', 'totals', 'trafficSource']
DROP_COLUMNS = ['customDimensions', 'hits', 'socialEngagementType']
def measure_memory(df, name):
size_df = df.memory_usage(deep=True)
log.info('{} size: {:.2f} MB'.format(name, size_df.sum().compute()/ 1024**2))
def read_parse_store(fin, label='XXX', bsize=1e9):
log.debug('Start with {}'.format(label))
df_ = develop_json_fields(fin, bsize=bsize, json_fields=JSON_COLUMNS, cols_2drop=DROP_COLUMNS)
#some stats
measure_memory(df_, label)
log.info('Number of partitions in {}: {}'.format(label, df_.npartitions))
#visualize a few rows
display(df_.head())
#reduce var size
df_['visitNumber'] = df_['visitNumber'].astype(np.uint16)
#read the whole dataset into pd.DataFrame in memory and store into a single file
#otherwise dask.DataFrame would be stored into multiple files- 1 per partition
df_.compute().to_csv("{}-flat.csv.gz".format(label), index=False , compression='gzip')
```
Process training data
```
%%time
read_parse_store('../input/train_v2.csv', 'train')
```
Process test data
```
%%time
read_parse_store('../input/test_v2.csv', 'test')
!ls -l
```
| github_jupyter |
# Taller 1: Básico de Python
+ Funciones
+ Listas
+ Diccionarios
Este taller es para resolver problemas básicos de python. Manejo de listas, diccionarios, etc.
El taller debe ser realizado en un Notebook de Jupyter en la carpeta de cada uno. Debe haber commits con el avance del taller. Debajo de cada pregunta hay una celda para el código.
## Basico de Python
### 1. Qué versión de python está corriendo?
```
import sys
print('{0[0]}.{0[1]}'.format(sys.version_info))
```
### 2. Calcule el área de un circulo de radio 5
```
pi = 3.1416
radio = 5
area= pi * radio**2
print(area)
```
### 3. Escriba código que imprima todos los colores de que están en color_list_1 y no estan presentes en color_list_2
Resultado esperado :
{'Black', 'White'}
```
color_list_1 = set(["White", "Black", "Red"])
color_list_2 = set(["Red", "Green"])
color_list_1 - color_list_2
```
### 4 Imprima una línea por cada carpeta que compone el Path donde se esta ejecutando python
e.g. C:/User/sergio/code/programación
Salida Esperada:
+ User
+ sergio
+ code
+ programacion
```
path = 'C:/Users/Margarita/Documents/Mis_documentos/Biologia_EAFIT/Semestre_IX/Programacion/'
size = len (path)
guardar = ""
for i in range(3,size):
if path[i] != '/':
guardar = guardar + path[i]
else:
print(guardar)
guardar = ""
```
## Manejo de Listas
### 5. Imprima la suma de números de my_list
```
my_list = [5,7,8,9,17]
sum_list = sum (my_list)
print(sum_list)
```
### 6. Inserte un elemento_a_insertar antes de cada elemento de my_list
```
elemento_a_insertar = 'E'
my_list = [1, 2, 3, 4]
```
La salida esperada es una lista así: [E, 1, E, 2, E, 3, E, 4]
```
elemento_a_insertar = 'E'
my_list = [1, 2, 3, 4]
size = len (my_list)
carpeta = []
for i in range(size):
carpeta = carpeta + [elemento_a_insertar,my_list[i]]
my_list = carpeta
print (my_list)
```
### 7. Separe my_list en una lista de lista cada N elementos
```
N = 3
my_list = ['a', 'b', 'c', 'd', 'e', 'f', 'g', 'h', 'i', 'j', 'k', 'l', 'm', 'n']
```
Salida Epserada: [['a', 'd', 'g', 'j', 'm'], ['b', 'e', 'h', 'k', 'n'], ['c', 'f', 'i', 'l']]
```
N=3
lista=[]
listaa = []
my_list = ['a', 'b', 'c', 'd', 'e', 'f', 'g', 'h', 'i', 'j', 'k', 'l', 'm', 'n']
size = len(my_list)
for i in range(N):
lista = lista + [listaa]
for i in range (size):
lista[i%N] = lista[i%N] + [my_list[i]]
print(lista)
```
### 8. Encuentra la lista dentro de list_of_lists que la suma de sus elementos sea la mayor
```
list_of_lists = [ [1,2,3], [4,5,6], [10,11,12], [7,8,9] ]
```
Salida Esperada: [10, 11, 12]
```
list_of_lists = [ [1,2,3], [4,5,6], [10,11,12], [7,8,9] ]
size = len(list_of_lists)
carpeta = list_of_lists[1]
for i in range(size):
if sum(list_of_lists[i]) > sum(carpeta):
carpeta = list_of_lists[i]
print(carpeta)
```
## Manejo de Diccionarios
### 9. Cree un diccionario que para cada número de 1 a N de llave tenga como valor N al cuadrado
```
N = 5
```
Salida Esperada: {1:1, 2:4, 3:9, 4:16, 5:25}
```
N = 5
diccio = {}
for i in range(1,N+1):
diccio [i]= i**2
print(diccio)
```
### 10. Concatene los diccionarios en dictionary_list para crear uno nuevo
```
dictionary_list=[{1:10, 2:20} , {3:30, 4:40}, {5:50,6:60}]
```
Salida Esperada: {1: 10, 2: 20, 3: 30, 4: 40, 5: 50, 6: 60}
```
dictionary_list=[{1:10, 2:20} , {3:30, 4:40}, {5:50,6:60}]
final= {}
for i in dictionary_list:
for k in i:
final[k] = i[k]
print(final)
```
### 11. Añada un nuevo valor "cuadrado" con el valor de "numero" de cada diccionario elevado al cuadrado
```
dictionary_list=[{'numero': 10, 'cantidad': 5} , {'numero': 12, 'cantidad': 3}, {'numero': 5, 'cantidad': 45}]
```
Salida Esperada: [{'numero': 10, 'cantidad': 5, 'cuadrado': 100} , {'numero': 12, 'cantidad': 3, , 'cuadrado': 144}, {'numero': 5, 'cantidad': 45, , 'cuadrado': 25}]
```
dictionary_list
```
## Manejo de Funciones
### 12. Defina y llame una función que reciva 2 parametros y solucione el problema __3__
```
def diferencia_conjuntos(color_list_1, color_list_2):
print (color_list_1 - color_list_2)
# Implementar la función
diferencia_conjuntos(
color_list_1 = set(["White", "Black", "Red"]) ,
color_list_2 = set(["Red", "Green"]))
```
### 13. Defina y llame una función que reciva de parametro una lista de listas y solucione el problema 8
```
def max_list_of_lists(list_of_lists):
size = len(list_of_lists)
carpeta = list_of_lists[1]
for i in range(size):
if sum(list_of_lists[i]) > sum(carpeta):
carpeta = list_of_lists[i]
print(carpeta)
# Implementar la función
list_of_lists = [ [1,2,3], [4,5,6], [10,11,12], [7,8,9] ]
max_list_of_lists (list_of_lists)
```
### 14. Defina y llame una función que reciva un parametro N y resuleva el problema 9
```
def diccionario_cuadradovalor(N):
diccio = {}
final = {}
for i in range(1,N+1):
final = diccio [i]= i**2
print(diccio)
#Implementar la función:
N = 5
diccionario_cuadradovalor(N)
```
| github_jupyter |
# Mask R-CNN - Inspect Balloon Training Data
Inspect and visualize data loading and pre-processing code.
```
import os
import sys
import itertools
import math
import logging
import json
import re
import random
from collections import OrderedDict
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
import matplotlib.patches as patches
import matplotlib.lines as lines
from matplotlib.patches import Polygon
# Root directory of the project
ROOT_DIR = os.path.abspath("../../")
# Import Mask RCNN
sys.path.append(ROOT_DIR) # To find local version of the library
from mrcnn import utils
from mrcnn import visualize
from mrcnn.visualize import display_images
import mrcnn.model as modellib
from mrcnn.model import log
from samples.balloon import balloon
%matplotlib inline
```
## Configurations
Configurations are defined in balloon.py
```
config = balloon.BalloonConfig()
BALLOON_DIR = os.path.join(ROOT_DIR, "datasets/balloon")
```
## Dataset
```
# Load dataset
# Get the dataset from the releases page
# https://github.com/matterport/Mask_RCNN/releases
dataset = balloon.BalloonDataset()
dataset.load_balloon(BALLOON_DIR, "train")
# Must call before using the dataset
dataset.prepare()
print("Image Count: {}".format(len(dataset.image_ids)))
print("Class Count: {}".format(dataset.num_classes))
for i, info in enumerate(dataset.class_info):
print("{:3}. {:50}".format(i, info['name']))
```
## Display Samples
Load and display images and masks.
```
# Load and display random samples
image_ids = np.random.choice(dataset.image_ids, 4)
for image_id in image_ids:
image = dataset.load_image(image_id)
mask, class_ids = dataset.load_mask(image_id)
visualize.display_top_masks(image, mask, class_ids, dataset.class_names)
```
## Bounding Boxes
Rather than using bounding box coordinates provided by the source datasets, we compute the bounding boxes from masks instead. This allows us to handle bounding boxes consistently regardless of the source dataset, and it also makes it easier to resize, rotate, or crop images because we simply generate the bounding boxes from the updates masks rather than computing bounding box transformation for each type of image transformation.
```
# Load random image and mask.
image_id = random.choice(dataset.image_ids)
image = dataset.load_image(image_id)
mask, class_ids = dataset.load_mask(image_id)
# Compute Bounding box
bbox = utils.extract_bboxes(mask)
# Display image and additional stats
print("image_id ", image_id, dataset.image_reference(image_id))
log("image", image)
log("mask", mask)
log("class_ids", class_ids)
log("bbox", bbox)
# Display image and instances
visualize.display_instances(image, bbox, mask, class_ids, dataset.class_names)
```
## Resize Images
To support multiple images per batch, images are resized to one size (1024x1024). Aspect ratio is preserved, though. If an image is not square, then zero padding is added at the top/bottom or right/left.
```
# Load random image and mask.
image_id = np.random.choice(dataset.image_ids, 1)[0]
image = dataset.load_image(image_id)
mask, class_ids = dataset.load_mask(image_id)
original_shape = image.shape
# Resize
image, window, scale, padding, _ = utils.resize_image(
image,
min_dim=config.IMAGE_MIN_DIM,
max_dim=config.IMAGE_MAX_DIM,
mode=config.IMAGE_RESIZE_MODE)
mask = utils.resize_mask(mask, scale, padding)
# Compute Bounding box
bbox = utils.extract_bboxes(mask)
# Display image and additional stats
print("image_id: ", image_id, dataset.image_reference(image_id))
print("Original shape: ", original_shape)
log("image", image)
log("mask", mask)
log("class_ids", class_ids)
log("bbox", bbox)
# Display image and instances
visualize.display_instances(image, bbox, mask, class_ids, dataset.class_names)
```
## Mini Masks
Instance binary masks can get large when training with high resolution images. For example, if training with 1024x1024 image then the mask of a single instance requires 1MB of memory (Numpy uses bytes for boolean values). If an image has 100 instances then that's 100MB for the masks alone.
To improve training speed, we optimize masks by:
* We store mask pixels that are inside the object bounding box, rather than a mask of the full image. Most objects are small compared to the image size, so we save space by not storing a lot of zeros around the object.
* We resize the mask to a smaller size (e.g. 56x56). For objects that are larger than the selected size we lose a bit of accuracy. But most object annotations are not very accuracy to begin with, so this loss is negligable for most practical purposes. Thie size of the mini_mask can be set in the config class.
To visualize the effect of mask resizing, and to verify the code correctness, we visualize some examples.
```
image_id = np.random.choice(dataset.image_ids, 1)[0]
image, image_meta, class_ids, bbox, mask = modellib.load_image_gt(
dataset, config, image_id, use_mini_mask=False)
log("image", image)
log("image_meta", image_meta)
log("class_ids", class_ids)
log("bbox", bbox)
log("mask", mask)
display_images([image]+[mask[:,:,i] for i in range(min(mask.shape[-1], 7))])
visualize.display_instances(image, bbox, mask, class_ids, dataset.class_names)
# Add augmentation and mask resizing.
image, image_meta, class_ids, bbox, mask = modellib.load_image_gt(
dataset, config, image_id, augment=True, use_mini_mask=True)
log("mask", mask)
display_images([image]+[mask[:,:,i] for i in range(min(mask.shape[-1], 7))])
mask = utils.expand_mask(bbox, mask, image.shape)
visualize.display_instances(image, bbox, mask, class_ids, dataset.class_names)
```
## Anchors
The order of anchors is important. Use the same order in training and prediction phases. And it must match the order of the convolution execution.
For an FPN network, the anchors must be ordered in a way that makes it easy to match anchors to the output of the convolution layers that predict anchor scores and shifts.
* Sort by pyramid level first. All anchors of the first level, then all of the second and so on. This makes it easier to separate anchors by level.
* Within each level, sort anchors by feature map processing sequence. Typically, a convolution layer processes a feature map starting from top-left and moving right row by row.
* For each feature map cell, pick any sorting order for the anchors of different ratios. Here we match the order of ratios passed to the function.
**Anchor Stride:**
In the FPN architecture, feature maps at the first few layers are high resolution. For example, if the input image is 1024x1024 then the feature meap of the first layer is 256x256, which generates about 200K anchors (256*256*3). These anchors are 32x32 pixels and their stride relative to image pixels is 4 pixels, so there is a lot of overlap. We can reduce the load significantly if we generate anchors for every other cell in the feature map. A stride of 2 will cut the number of anchors by 4, for example.
In this implementation we use an anchor stride of 2, which is different from the paper.
```
# Generate Anchors
backbone_shapes = modellib.compute_backbone_shapes(config, config.IMAGE_SHAPE)
anchors = utils.generate_pyramid_anchors(config.RPN_ANCHOR_SCALES,
config.RPN_ANCHOR_RATIOS,
backbone_shapes,
config.BACKBONE_STRIDES,
config.RPN_ANCHOR_STRIDE)
# Print summary of anchors
num_levels = len(backbone_shapes)
anchors_per_cell = len(config.RPN_ANCHOR_RATIOS)
print("Count: ", anchors.shape[0])
print("Scales: ", config.RPN_ANCHOR_SCALES)
print("ratios: ", config.RPN_ANCHOR_RATIOS)
print("Anchors per Cell: ", anchors_per_cell)
print("Levels: ", num_levels)
anchors_per_level = []
for l in range(num_levels):
num_cells = backbone_shapes[l][0] * backbone_shapes[l][1]
anchors_per_level.append(anchors_per_cell * num_cells // config.RPN_ANCHOR_STRIDE**2)
print("Anchors in Level {}: {}".format(l, anchors_per_level[l]))
```
Visualize anchors of one cell at the center of the feature map of a specific level.
```
## Visualize anchors of one cell at the center of the feature map of a specific level
# Load and draw random image
image_id = np.random.choice(dataset.image_ids, 1)[0]
image, image_meta, _, _, _ = modellib.load_image_gt(dataset, config, image_id)
fig, ax = plt.subplots(1, figsize=(10, 10))
ax.imshow(image)
levels = len(backbone_shapes)
for level in range(levels):
colors = visualize.random_colors(levels)
# Compute the index of the anchors at the center of the image
level_start = sum(anchors_per_level[:level]) # sum of anchors of previous levels
level_anchors = anchors[level_start:level_start+anchors_per_level[level]]
print("Level {}. Anchors: {:6} Feature map Shape: {}".format(level, level_anchors.shape[0],
backbone_shapes[level]))
center_cell = backbone_shapes[level] // 2
center_cell_index = (center_cell[0] * backbone_shapes[level][1] + center_cell[1])
level_center = center_cell_index * anchors_per_cell
center_anchor = anchors_per_cell * (
(center_cell[0] * backbone_shapes[level][1] / config.RPN_ANCHOR_STRIDE**2) \
+ center_cell[1] / config.RPN_ANCHOR_STRIDE)
level_center = int(center_anchor)
# Draw anchors. Brightness show the order in the array, dark to bright.
for i, rect in enumerate(level_anchors[level_center:level_center+anchors_per_cell]):
y1, x1, y2, x2 = rect
p = patches.Rectangle((x1, y1), x2-x1, y2-y1, linewidth=2, facecolor='none',
edgecolor=(i+1)*np.array(colors[level]) / anchors_per_cell)
ax.add_patch(p)
```
## Data Generator
```
# Create data generator
random_rois = 2000
g = modellib.data_generator(
dataset, config, shuffle=True, random_rois=random_rois,
batch_size=4,
detection_targets=True)
# Uncomment to run the generator through a lot of images
# to catch rare errors
# for i in range(1000):
# print(i)
# _, _ = next(g)
# Get Next Image
if random_rois:
[normalized_images, image_meta, rpn_match, rpn_bbox, gt_class_ids, gt_boxes, gt_masks, rpn_rois, rois], \
[mrcnn_class_ids, mrcnn_bbox, mrcnn_mask] = next(g)
log("rois", rois)
log("mrcnn_class_ids", mrcnn_class_ids)
log("mrcnn_bbox", mrcnn_bbox)
log("mrcnn_mask", mrcnn_mask)
else:
[normalized_images, image_meta, rpn_match, rpn_bbox, gt_boxes, gt_masks], _ = next(g)
log("gt_class_ids", gt_class_ids)
log("gt_boxes", gt_boxes)
log("gt_masks", gt_masks)
log("rpn_match", rpn_match, )
log("rpn_bbox", rpn_bbox)
image_id = modellib.parse_image_meta(image_meta)["image_id"][0]
print("image_id: ", image_id, dataset.image_reference(image_id))
# Remove the last dim in mrcnn_class_ids. It's only added
# to satisfy Keras restriction on target shape.
mrcnn_class_ids = mrcnn_class_ids[:,:,0]
b = 0
# Restore original image (reverse normalization)
sample_image = modellib.unmold_image(normalized_images[b], config)
# Compute anchor shifts.
indices = np.where(rpn_match[b] == 1)[0]
refined_anchors = utils.apply_box_deltas(anchors[indices], rpn_bbox[b, :len(indices)] * config.RPN_BBOX_STD_DEV)
log("anchors", anchors)
log("refined_anchors", refined_anchors)
# Get list of positive anchors
positive_anchor_ids = np.where(rpn_match[b] == 1)[0]
print("Positive anchors: {}".format(len(positive_anchor_ids)))
negative_anchor_ids = np.where(rpn_match[b] == -1)[0]
print("Negative anchors: {}".format(len(negative_anchor_ids)))
neutral_anchor_ids = np.where(rpn_match[b] == 0)[0]
print("Neutral anchors: {}".format(len(neutral_anchor_ids)))
# ROI breakdown by class
for c, n in zip(dataset.class_names, np.bincount(mrcnn_class_ids[b].flatten())):
if n:
print("{:23}: {}".format(c[:20], n))
# Show positive anchors
fig, ax = plt.subplots(1, figsize=(16, 16))
visualize.draw_boxes(sample_image, boxes=anchors[positive_anchor_ids],
refined_boxes=refined_anchors, ax=ax)
# Show negative anchors
visualize.draw_boxes(sample_image, boxes=anchors[negative_anchor_ids])
# Show neutral anchors. They don't contribute to training.
visualize.draw_boxes(sample_image, boxes=anchors[np.random.choice(neutral_anchor_ids, 100)])
```
## ROIs
```
if random_rois:
# Class aware bboxes
bbox_specific = mrcnn_bbox[b, np.arange(mrcnn_bbox.shape[1]), mrcnn_class_ids[b], :]
# Refined ROIs
refined_rois = utils.apply_box_deltas(rois[b].astype(np.float32), bbox_specific[:,:4] * config.BBOX_STD_DEV)
# Class aware masks
mask_specific = mrcnn_mask[b, np.arange(mrcnn_mask.shape[1]), :, :, mrcnn_class_ids[b]]
visualize.draw_rois(sample_image, rois[b], refined_rois, mask_specific, mrcnn_class_ids[b], dataset.class_names)
# Any repeated ROIs?
rows = np.ascontiguousarray(rois[b]).view(np.dtype((np.void, rois.dtype.itemsize * rois.shape[-1])))
_, idx = np.unique(rows, return_index=True)
print("Unique ROIs: {} out of {}".format(len(idx), rois.shape[1]))
if random_rois:
# Dispalay ROIs and corresponding masks and bounding boxes
ids = random.sample(range(rois.shape[1]), 8)
images = []
titles = []
for i in ids:
image = visualize.draw_box(sample_image.copy(), rois[b,i,:4].astype(np.int32), [255, 0, 0])
image = visualize.draw_box(image, refined_rois[i].astype(np.int64), [0, 255, 0])
images.append(image)
titles.append("ROI {}".format(i))
images.append(mask_specific[i] * 255)
titles.append(dataset.class_names[mrcnn_class_ids[b,i]][:20])
display_images(images, titles, cols=4, cmap="Blues", interpolation="none")
# Check ratio of positive ROIs in a set of images.
if random_rois:
limit = 10
temp_g = modellib.data_generator(
dataset, config, shuffle=True, random_rois=10000,
batch_size=1, detection_targets=True)
total = 0
for i in range(limit):
_, [ids, _, _] = next(temp_g)
positive_rois = np.sum(ids[0] > 0)
total += positive_rois
print("{:5} {:5.2f}".format(positive_rois, positive_rois/ids.shape[1]))
print("Average percent: {:.2f}".format(total/(limit*ids.shape[1])))
```
| github_jupyter |
```
#hide
#skip
! [ -e /content ] && pip install -Uqq fastai # upgrade fastai on colab
# default_exp interpret
#export
from fastai.data.all import *
from fastai.optimizer import *
from fastai.learner import *
import sklearn.metrics as skm
#hide
from fastai.test_utils import *
```
# Interpretation
> Classes to build objects to better interpret predictions of a model
```
#export
@typedispatch
def plot_top_losses(x, y, *args, **kwargs):
raise Exception(f"plot_top_losses is not implemented for {type(x)},{type(y)}")
#export
_all_ = ["plot_top_losses"]
#export
class Interpretation():
"Interpretation base class, can be inherited for task specific Interpretation classes"
def __init__(self, dl, inputs, preds, targs, decoded, losses):
store_attr("dl,inputs,preds,targs,decoded,losses")
@classmethod
def from_learner(cls, learn, ds_idx=1, dl=None, act=None):
"Construct interpretation object from a learner"
if dl is None: dl = learn.dls[ds_idx]
return cls(dl, *learn.get_preds(dl=dl, with_input=True, with_loss=True, with_decoded=True, act=None))
def top_losses(self, k=None, largest=True):
"`k` largest(/smallest) losses and indexes, defaulting to all losses (sorted by `largest`)."
return self.losses.topk(ifnone(k, len(self.losses)), largest=largest)
def plot_top_losses(self, k, largest=True, **kwargs):
losses,idx = self.top_losses(k, largest)
if not isinstance(self.inputs, tuple): self.inputs = (self.inputs,)
if isinstance(self.inputs[0], Tensor): inps = tuple(o[idx] for o in self.inputs)
else: inps = self.dl.create_batch(self.dl.before_batch([tuple(o[i] for o in self.inputs) for i in idx]))
b = inps + tuple(o[idx] for o in (self.targs if is_listy(self.targs) else (self.targs,)))
x,y,its = self.dl._pre_show_batch(b, max_n=k)
b_out = inps + tuple(o[idx] for o in (self.decoded if is_listy(self.decoded) else (self.decoded,)))
x1,y1,outs = self.dl._pre_show_batch(b_out, max_n=k)
if its is not None:
plot_top_losses(x, y, its, outs.itemgot(slice(len(inps), None)), self.preds[idx], losses, **kwargs)
#TODO: figure out if this is needed
#its None means that a batch knows how to show itself as a whole, so we pass x, x1
#else: show_results(x, x1, its, ctxs=ctxs, max_n=max_n, **kwargs)
learn = synth_learner()
interp = Interpretation.from_learner(learn)
x,y = learn.dls.valid_ds.tensors
test_eq(interp.inputs, x)
test_eq(interp.targs, y)
out = learn.model.a * x + learn.model.b
test_eq(interp.preds, out)
test_eq(interp.losses, (out-y)[:,0]**2)
#export
class ClassificationInterpretation(Interpretation):
"Interpretation methods for classification models."
def __init__(self, dl, inputs, preds, targs, decoded, losses):
super().__init__(dl, inputs, preds, targs, decoded, losses)
self.vocab = self.dl.vocab
if is_listy(self.vocab): self.vocab = self.vocab[-1]
def confusion_matrix(self):
"Confusion matrix as an `np.ndarray`."
x = torch.arange(0, len(self.vocab))
d,t = flatten_check(self.decoded, self.targs)
cm = ((d==x[:,None]) & (t==x[:,None,None])).long().sum(2)
return to_np(cm)
def plot_confusion_matrix(self, normalize=False, title='Confusion matrix', cmap="Blues", norm_dec=2,
plot_txt=True, **kwargs):
"Plot the confusion matrix, with `title` and using `cmap`."
# This function is mainly copied from the sklearn docs
cm = self.confusion_matrix()
if normalize: cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
fig = plt.figure(**kwargs)
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
tick_marks = np.arange(len(self.vocab))
plt.xticks(tick_marks, self.vocab, rotation=90)
plt.yticks(tick_marks, self.vocab, rotation=0)
if plot_txt:
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
coeff = f'{cm[i, j]:.{norm_dec}f}' if normalize else f'{cm[i, j]}'
plt.text(j, i, coeff, horizontalalignment="center", verticalalignment="center", color="white" if cm[i, j] > thresh else "black")
ax = fig.gca()
ax.set_ylim(len(self.vocab)-.5,-.5)
plt.tight_layout()
plt.ylabel('Actual')
plt.xlabel('Predicted')
plt.grid(False)
def most_confused(self, min_val=1):
"Sorted descending list of largest non-diagonal entries of confusion matrix, presented as actual, predicted, number of occurrences."
cm = self.confusion_matrix()
np.fill_diagonal(cm, 0)
res = [(self.vocab[i],self.vocab[j],cm[i,j])
for i,j in zip(*np.where(cm>=min_val))]
return sorted(res, key=itemgetter(2), reverse=True)
def print_classification_report(self):
"Print scikit-learn classification report"
d,t = flatten_check(self.decoded, self.targs)
print(skm.classification_report(t, d, labels=list(self.vocab.o2i.values()), target_names=[str(v) for v in self.vocab]))
```
## Export -
```
#hide
from nbdev.export import notebook2script
notebook2script()
```
| github_jupyter |
# MNIST uDNN Model Pre-training
Credit: this notebook is based on Keras official documents https://keras.io/examples/vision/mnist_convnet/
[](https://colab.research.google.com/drive/1-Bn1Z7XItLokDJanKX9I3KlBWWjXI-RE?usp=sharing)
```
import numpy as np
from tensorflow import keras
from tensorflow.keras import layers
import tensorflow as tf
import matplotlib.pyplot as plt
```
## Use Google Colab Pro GPU (Optional)
```
device_name = tf.test.gpu_device_name()
if device_name != '/device:GPU:0':
raise SystemError('GPU device not found')
print('Found GPU at: {}'.format(device_name))
```
## Import MNIST Dataset
```
# Model / data parameters
num_classes = 10
input_shape = (28, 28, 1)
# the data, split between train and test sets
(trainX, trainY), (testX, testY) = keras.datasets.mnist.load_data()
# Scale images to the [0, 1] range
trainX = trainX.astype("float32") / 255
testX = testX.astype("float32") / 255
# Make sure images have shape (28, 28, 1)
trainX = np.expand_dims(trainX, -1)
testX = np.expand_dims(testX, -1)
print("trainX shape:", trainX.shape)
print(trainX.shape[0], "train samples")
print(testX.shape[0], "test samples")
# convert class vectors to binary class matrices
trainY = keras.utils.to_categorical(trainY, num_classes)
testY = keras.utils.to_categorical(testY, num_classes)
```
## Define uDNN Model
```
model = keras.Sequential([
keras.Input(shape=input_shape),
layers.Conv2D(16, kernel_size=(3,3), activation="relu"),
layers.MaxPooling2D(pool_size=(2,2)),
layers.Conv2D(32, kernel_size=(3,3), activation="relu"),
layers.MaxPooling2D(pool_size=(2,2)),
layers.Flatten(),
layers.Dropout(0.5),
layers.Dense(num_classes, activation="softmax"),
])
model.summary()
```
## Compile and Train Model
```
model.compile(loss="categorical_crossentropy", optimizer="adam", metrics=["accuracy"])
model.fit(trainX, trainY, epochs=15, batch_size=128, validation_split=0.1)
model.evaluate(testX, testY)
```
## Prepare to Export Model
```
! git clone https://github.com/leleonardzhang/uDNN-tf2msp.git
! pip install fxpmath
import sys
from fxpmath import Fxp
sys.path.insert(0, '/content/uDNN-tf2msp')
import encoder
```
## Export Model to Header File
```
encoder.export_model(model)
```
## Get Sample Input and Output (Optional)
```
test_n = 1 # the nth data in test dataset
print("Fixed Point Input")
print(Fxp(testX[test_n].transpose(2,0,1), signed = True, n_word = 16, n_frac = 10).val.flatten().tolist())
print("Output Label")
print(np.argmax(testY[test_n]))
```
## Next Step
1. Download header file `neural_network_parameters.h` and copy the header file into the root directory of MSP implementation `uDNN-tf2msp-msp_impl/`.
2. Modify the pointer `input_buffer` in `neural_network_parameters.h` and point to desired data input.
3. Compile and run the program on MSP.
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.