text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
========= ===============================================<br>
run task<br>
========= ===============================================<br>
1 Baseline, eyes open<br>
2 Baseline, eyes closed<br>
3, 7, 11 Motor execution: left vs right hand<br>
4, 8, 12 Motor imagery: left vs right hand<br>
5, 9, 13 Motor execution: hands vs feet<br>
6, 10, 14 Motor imagery: hands vs feet<br>
========= ===============================================<br>
```
!pip install mne
```
# ResNet Keras
```
from tensorflow.keras.models import Model
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Activation, Permute, Dropout, Add
from tensorflow.keras.layers import Conv2D, MaxPooling2D, AveragePooling2D
from tensorflow.keras.layers import Conv1D, MaxPooling1D, AveragePooling1D
from tensorflow.keras.layers import SeparableConv2D, DepthwiseConv2D
from tensorflow.keras.layers import BatchNormalization
from tensorflow.keras.layers import SpatialDropout2D
from tensorflow.keras.regularizers import l1_l2, l2
from tensorflow.keras.layers import Input, Flatten
from tensorflow.keras.constraints import max_norm
from tensorflow.keras import backend as K
from tensorflow.keras.initializers import glorot_uniform
from tensorflow.keras.callbacks import EarlyStopping
import tensorflow as tf
def identity_block(X, f, filters, stage, block, l1=0.0, l2=0.01):
# Defining name basis
conv_name_base = 'res' + str(stage) + block + '_branch'
bn_name_base = 'bn' + str(stage) + block + '_branch'
# Retrieve Filters
F1, F2, F3 = filters
# Save the input value
X_shortcut = X
# First component of main path
X = Conv2D(filters = F1, kernel_size = (1, 1), strides = (1,1), data_format='channels_first',
padding = 'valid', name = conv_name_base + '2a', kernel_initializer = glorot_uniform(seed=0),
activity_regularizer=l1_l2(l1, l2))(X)
X = BatchNormalization(axis = 1, name = bn_name_base + '2a')(X)
X = Activation('relu')(X)
# Second component of main path
X = Conv2D(filters = F2, kernel_size = (f, f), strides = (1, 1), data_format='channels_first', padding = 'same',
name = conv_name_base + '2b', kernel_initializer = glorot_uniform(seed=0),
activity_regularizer=l1_l2(l1, l2))(X)
X = BatchNormalization(axis = 1, name = bn_name_base + '2b')(X)
X = Activation('relu')(X)
# Third component of main path
X = Conv2D(filters = F3, kernel_size = (1, 1), strides = (1, 1), data_format='channels_first', padding = 'valid',
name = conv_name_base + '2c', kernel_initializer = glorot_uniform(seed=0),
activity_regularizer=l1_l2(l1, l2))(X)
X = BatchNormalization(axis = 1, name = bn_name_base + '2c')(X)
# Final step: Add shortcut value to main path, and pass it through a RELU activation
X = Add()([X, X_shortcut])
X = Activation('relu')(X)
return X
def convolutional_block(X, f, filters, stage, block, s=2, l1=0.0, l2=0.01):
# Defining name basis
conv_name_base = 'res' + str(stage) + block + '_branch'
bn_name_base = 'bn' + str(stage) + block + '_branch'
# Retrieve Filters
F1, F2, F3 = filters
# Save the input value
X_shortcut = X
##### MAIN PATH #####
# First component of main path
X = Conv2D(filters=F1, kernel_size=(1, 1), strides=(s, s),
data_format='channels_first', padding='same',
name=conv_name_base + '2a', kernel_initializer=glorot_uniform(seed=0),
activity_regularizer=l1_l2(l1, l2))(X)
X = BatchNormalization(axis=1, name=bn_name_base + '2a')(X)
X = Activation('relu')(X)
# Second component of main path
X = Conv2D(filters=F2, kernel_size=(f, f), strides=(1, 1),
data_format='channels_first', padding='same', name=conv_name_base + '2b',
kernel_initializer=glorot_uniform(seed=0),
activity_regularizer=l1_l2(l1, l2))(X)
X = BatchNormalization(axis=1, name=bn_name_base + '2b')(X)
X = Activation('relu')(X)
# Third component of main path
X = Conv2D(filters=F3, kernel_size=(1, 1), strides=(1, 1),
data_format='channels_first', padding='same', name=conv_name_base + '2c',
kernel_initializer=glorot_uniform(seed=0),
activity_regularizer=l1_l2(l1, l2))(X)
X = BatchNormalization(axis=1, name=bn_name_base + '2c')(X)
##### SHORTCUT PATH ####
X_shortcut = Conv2D(filters=F3, kernel_size=(1, 1), strides=(s, s), data_format='channels_first',
padding='same', name=conv_name_base + '1', kernel_initializer=glorot_uniform(seed=0),
activity_regularizer=l1_l2(l1, l2))(X_shortcut)
X_shortcut = BatchNormalization(axis=1, name=bn_name_base + '1')(X_shortcut)
# Final step: Add shortcut value to main path, and pass it through a RELU activation
X = Add()([X, X_shortcut])
X = Activation('relu')(X)
return X
def ResNet(chans=64, samples=121, l1=0.0, l2=0.01):
input_shape= (1, chans, samples)
input1 = Input(shape = input_shape)
##################################################################
X = Conv2D(128, (1, 64), padding = 'same',
input_shape = input_shape, data_format='channels_first',
kernel_initializer = glorot_uniform(seed=0),
activity_regularizer=l1_l2(l1, l2))(input1)
X = BatchNormalization(axis = 1)(X)
X = DepthwiseConv2D((64, 1), kernel_initializer = glorot_uniform(seed=0),
depth_multiplier = 2, data_format='channels_first',
activity_regularizer=l1_l2(l1, l2))(X)
X = BatchNormalization(axis = 1)(X)
X = Activation('elu')(X)
X = AveragePooling2D((1, 2), data_format='channels_first')(X)
# Stage 2
X = convolutional_block(X, f=3, filters=[64, 64, 256], stage=2, block='a', s=1, l1=l1, l2=l2)
X = identity_block(X, 3, [64, 64, 256], stage=2, block='b', l1=l1, l2=l2)
X = identity_block(X, 3, [64, 64, 256], stage=2, block='c', l1=l1, l2=l2)
### START CODE HERE ###
# Stage 3 (≈4 lines)
X = convolutional_block(X, f=3, filters=[128, 128, 512], stage=3, block='a', s=1, l1=l1, l2=l2)
X = identity_block(X, 3, [128, 128, 512], stage=3, block='b', l1=l1, l2=l2)
X = identity_block(X, 3, [128, 128, 512], stage=3, block='c', l1=l1, l2=l2)
X = identity_block(X, 3, [128, 128, 512], stage=3, block='d', l1=l1, l2=l2)
# Stage 4 (≈6 lines)
# X = convolutional_block(X, f=3, filters=[256, 256, 1024], stage=4, block='a', s=1, l1=l1, l2=l2)
# X = identity_block(X, 3, [256, 256, 1024], stage=4, block='b', l1=l1, l2=l2)
# X = identity_block(X, 3, [256, 256, 1024], stage=4, block='c', l1=l1, l2=l2)
# X = identity_block(X, 3, [256, 256, 1024], stage=4, block='d', l1=l1, l2=l2)
# X = identity_block(X, 3, [256, 256, 1024], stage=4, block='e', l1=l1, l2=l2)
# X = identity_block(X, 3, [256, 256, 1024], stage=4, block='f', l1=l1, l2=l2)
# # Stage 5 (≈4 lines)
# X = convolutional_block(X, f=3, filters=[512, 512, 2048], stage=5, block='a', s=2, l1=l1, l2=l2)
# X = identity_block(X, 3, [512, 512, 2048], stage=5, block='b')
# X = identity_block(X, 3, [512, 512, 2048], stage=5, block='c')
# X = identity_block(X, 3, [512, 512, 2048], stage=5, block='d')
# # Stage 6 (≈4 lines)
# X = convolutional_block(X, f=3, filters=[512, 512, 2048], stage=6, block='a', s=2, l1=l1, l2=l2)
# X = identity_block(X, 3, [512, 512, 2048], stage=6, block='b')
# X = identity_block(X, 3, [512, 512, 2048], stage=6, block='c')
# X = identity_block(X, 3, [512, 512, 2048], stage=6, block='d')
# Stage 7 (≈3 lines)
# X = X = convolutional_block(X, f=3, filters=[512, 512, 2048], stage=7, block='a', s=2, l1=l1, l2=l2)
# X = identity_block(X, 3, [512, 512, 2048], stage=7, block='b', l1=l1, l2=l2)
# X = identity_block(X, 3, [512, 512, 2048], stage=7, block='c', l1=l1, l2=l2)
# AVGPOOL (≈1 line). Use "X = AveragePooling2D(...)(X)"
X = AveragePooling2D(pool_size=(2, 2), padding='same')(X)
### END CODE HERE ###
# output layer
X = Flatten()(X)
X = Dense(64, activation='relu')(X)
X = Dense(32, activation='relu')(X)
X = Dense(1, activation='sigmoid', name='fc1', kernel_initializer = glorot_uniform(seed=0))(X)
# Create model
model = Model(inputs = input1, outputs = X, name='ResNet50')
return model
```
# EGGNet Keras Model
```
from tensorflow.keras.models import Model
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Activation, Permute, Dropout
from tensorflow.keras.layers import Conv2D, MaxPooling2D, AveragePooling2D
from tensorflow.keras.layers import Conv1D, MaxPooling1D, AveragePooling1D
from tensorflow.keras.layers import SeparableConv2D, DepthwiseConv2D
from tensorflow.keras.layers import BatchNormalization
from tensorflow.keras.layers import SpatialDropout2D
from tensorflow.keras.regularizers import l1_l2
from tensorflow.keras.layers import Input, Flatten
from tensorflow.keras.constraints import max_norm
from tensorflow.keras import backend as K
def EEGNet(nb_classes, Chans = 64, Samples = 128,
dropoutRate = 0.5, kernLength = 64, F1 = 8,
D = 2, F2 = 16, norm_rate = 0.25, dropoutType = 'Dropout', gpu=True):
""" Keras Implementation of EEGNet
http://iopscience.iop.org/article/10.1088/1741-2552/aace8c/meta
Note that this implements the newest version of EEGNet and NOT the earlier
version (version v1 and v2 on arxiv). We strongly recommend using this
architecture as it performs much better and has nicer properties than
our earlier version. For example:
1. Depthwise Convolutions to learn spatial filters within a
temporal convolution. The use of the depth_multiplier option maps
exactly to the number of spatial filters learned within a temporal
filter. This matches the setup of algorithms like FBCSP which learn
spatial filters within each filter in a filter-bank. This also limits
the number of free parameters to fit when compared to a fully-connected
convolution.
2. Separable Convolutions to learn how to optimally combine spatial
filters across temporal bands. Separable Convolutions are Depthwise
Convolutions followed by (1x1) Pointwise Convolutions.
While the original paper used Dropout, we found that SpatialDropout2D
sometimes produced slightly better results for classification of ERP
signals. However, SpatialDropout2D significantly reduced performance
on the Oscillatory dataset (SMR, BCI-IV Dataset 2A). We recommend using
the default Dropout in most cases.
Assumes the input signal is sampled at 128Hz. If you want to use this model
for any other sampling rate you will need to modify the lengths of temporal
kernels and average pooling size in blocks 1 and 2 as needed (double the
kernel lengths for double the sampling rate, etc). Note that we haven't
tested the model performance with this rule so this may not work well.
The model with default parameters gives the EEGNet-8,2 model as discussed
in the paper. This model should do pretty well in general, although it is
advised to do some model searching to get optimal performance on your
particular dataset.
We set F2 = F1 * D (number of input filters = number of output filters) for
the SeparableConv2D layer. We haven't extensively tested other values of this
parameter (say, F2 < F1 * D for compressed learning, and F2 > F1 * D for
overcomplete). We believe the main parameters to focus on are F1 and D.
Inputs:
nb_classes : int, number of classes to classify
Chans, Samples : number of channels and time points in the EEG data
dropoutRate : dropout fraction
kernLength : length of temporal convolution in first layer. We found
that setting this to be half the sampling rate worked
well in practice. For the SMR dataset in particular
since the data was high-passed at 4Hz we used a kernel
length of 32.
F1, F2 : number of temporal filters (F1) and number of pointwise
filters (F2) to learn. Default: F1 = 8, F2 = F1 * D.
D : number of spatial filters to learn within each temporal
convolution. Default: D = 2
dropoutType : Either SpatialDropout2D or Dropout, passed as a string.
"""
if dropoutType == 'SpatialDropout2D':
dropoutType = SpatialDropout2D
elif dropoutType == 'Dropout':
dropoutType = Dropout
else:
raise ValueError('dropoutType must be one of SpatialDropout2D '
'or Dropout, passed as a string.')
input1 = Input(shape = (1, Chans, Samples))
##################################################################
block1 = Conv2D(F1, (1, kernLength), padding = 'same',
input_shape = (1, Chans, Samples),
use_bias = False)(input1)
block1 = BatchNormalization(axis = 1)(block1)
block1 = DepthwiseConv2D((Chans, 1), use_bias = False,
depth_multiplier = D,
depthwise_constraint = max_norm(1.))(block1)
block1 = BatchNormalization(axis = 1)(block1)
block1 = Activation('elu')(block1)
block1 = AveragePooling2D((1, 4))(block1)
block1 = dropoutType(dropoutRate)(block1)
block2 = SeparableConv2D(F2, (1, 16),
use_bias = False, padding = 'same')(block1)
block2 = BatchNormalization(axis = 1)(block2)
block2 = Activation('elu')(block2)
block2 = AveragePooling2D((1, 8))(block2)
block2 = dropoutType(dropoutRate)(block2)
flatten = Flatten(name = 'flatten')(block2)
# dense = Dense(50,name='dense1', kernel_constraint = max_norm(norm_rate))(flatten)
dense = Dense(1, name = 'out', kernel_constraint = max_norm(norm_rate))(flatten)
softmax = Activation('sigmoid', name = 'sigmoid')(dense)
return Model(inputs=input1, outputs=softmax)
def DeepConvNet(nb_classes, Chans = 64, Samples = 256,
dropoutRate = 0.5):
""" Keras implementation of the Deep Convolutional Network as described in
Schirrmeister et. al. (2017), Human Brain Mapping.
This implementation assumes the input is a 2-second EEG signal sampled at
128Hz, as opposed to signals sampled at 250Hz as described in the original
paper. We also perform temporal convolutions of length (1, 5) as opposed
to (1, 10) due to this sampling rate difference.
Note that we use the max_norm constraint on all convolutional layers, as
well as the classification layer. We also change the defaults for the
BatchNormalization layer. We used this based on a personal communication
with the original authors.
ours original paper
pool_size 1, 2 1, 3
strides 1, 2 1, 3
conv filters 1, 5 1, 10
Note that this implementation has not been verified by the original
authors.
"""
# K.set_image_data_format('channels_first')
# start the model
input_main = Input((1, Chans, Samples))
block1 = Conv2D(25, (1, 11),
input_shape=(1, Chans, Samples),data_format='channels_first',
kernel_constraint = max_norm(2.1, axis=(0,1,2)))(input_main)
block1 = Conv2D(25, (Chans, 1),data_format='channels_first',
kernel_constraint = max_norm(2.1, axis=(0,1,2)))(block1)
block1 = BatchNormalization(axis=1, epsilon=1e-05, momentum=0.1)(block1)
block1 = Activation('elu')(block1)
block1 = MaxPooling2D(pool_size=(1, 2), strides=(1, 2),data_format='channels_first')(block1)
block1 = Dropout(dropoutRate)(block1)
block2 = Conv2D(50, (1, 9),data_format='channels_first',
kernel_constraint = max_norm(2.1, axis=(0,1,2)))(block1)
block2 = BatchNormalization(axis=1, epsilon=1e-05, momentum=0.1)(block2)
block2 = Activation('elu')(block2)
block2 = MaxPooling2D(pool_size=(1, 2), strides=(1, 2),data_format='channels_first')(block2)
block2 = Dropout(dropoutRate)(block2)
block3 = Conv2D(100, (1, 7),data_format='channels_first',
kernel_constraint = max_norm(2.1, axis=(0,1,2)))(block2)
block3 = BatchNormalization(axis=1, epsilon=1e-05, momentum=0.1)(block3)
block3 = Activation('elu')(block3)
block3 = MaxPooling2D(pool_size=(1, 2), strides=(1, 2),data_format='channels_first')(block3)
block3 = Dropout(dropoutRate)(block3)
block4 = Conv2D(200, (1, 5),data_format='channels_first',
kernel_constraint = max_norm(2.1, axis=(0,1,2)))(block3)
block4 = BatchNormalization(axis=1, epsilon=1e-05, momentum=0.1)(block4)
block4 = Activation('elu')(block4)
# block4 = MaxPooling2D(pool_size=(1, 2), strides=(1, 2),data_format='channels_first')(block4)
block4 = Dropout(dropoutRate)(block4)
block5 = Conv2D(250, (1, 3),data_format='channels_first',
kernel_constraint = max_norm(2.1, axis=(0,1,2)))(block4)
block5 = BatchNormalization(axis=1, epsilon=1e-05, momentum=0.1)(block5)
block5 = Activation('elu')(block5)
# block5 = MaxPooling2D(pool_size=(1, 2), strides=(1, 2),data_format='channels_first')(block5)
block5 = Dropout(dropoutRate)(block5)
# block6 = Conv2D(250, (1, 2),data_format='channels_first',
# kernel_constraint = max_norm(2., axis=(0,1,2)))(block5)
# block6 = BatchNormalization(axis=1, epsilon=1e-05, momentum=0.1)(block6)
# block6 = Activation('elu')(block6)
# # block5 = MaxPooling2D(pool_size=(1, 2), strides=(1, 2),data_format='channels_first')(block5)
# block6 = Dropout(dropoutRate)(block6)
flatten = Flatten()(block5)
dense = Dense(32, kernel_constraint = max_norm(0.6), activation='relu')(flatten)
dense = Dense(1, kernel_constraint = max_norm(0.6))(dense)
softmax = Activation('sigmoid')(dense)
return Model(inputs=input_main, outputs=softmax)
# DeepConvNet(2, Chans = 64, Samples = 113,
# dropoutRate = 0.5).summary()
```
# EFFNet
```
from tensorflow.keras.models import Model
from tensorflow.keras.layers import *
from tensorflow.keras.activations import *
from tensorflow.keras.callbacks import *
def get_post(x_in):
x = LeakyReLU()(x_in)
x = BatchNormalization()(x)
return x
def get_block(x_in, ch_in, ch_out):
x = Conv2D(ch_in,
kernel_size=(1, 1),
padding='same',
use_bias=False, data_format='channels_first')(x_in)
x = get_post(x)
x = DepthwiseConv2D(kernel_size=(1, 3), padding='same', use_bias=False, data_format='channels_first')(x)
x = get_post(x)
x = MaxPool2D(pool_size=(2, 1),
strides=(2, 1), data_format='channels_first')(x) # Separable pooling
x = DepthwiseConv2D(kernel_size=(3, 1),
padding='same',
use_bias=False, data_format='channels_first')(x)
x = get_post(x)
x = Conv2D(ch_out,
kernel_size=(2, 1),
strides=(1, 2),
padding='same',
use_bias=False, data_format='channels_first')(x)
x = get_post(x)
return x
def Effnet(input_shape, nb_classes, include_top=True, weights=None):
x_in = Input(shape=input_shape)
x = get_block(x_in, 32, 64)
x = get_block(x, 64, 128)
x = get_block(x, 128, 256)
if include_top:
x = Flatten()(x)
x = Dense(1, activation='sigmoid')(x)
model = Model(inputs=x_in, outputs=x)
if weights is not None:
model.load_weights(weights, by_name=True)
return model
```
# My Models
```
def my_net(nb_classes=2, Chans = 64, Samples = 321,
dropoutRate = 0.5, kernLength = 64, F1 = 8,
D = 2, F2 = 16, norm_rate = 0.25, dropoutType = 'Dropout', gpu=True):
if dropoutType == 'SpatialDropout2D':
dropoutType = SpatialDropout2D
elif dropoutType == 'Dropout':
dropoutType = Dropout
else:
raise ValueError('dropoutType must be one of SpatialDropout2D '
'or Dropout, passed as a string.')
input1 = Input(shape = (1, Chans, Samples))
##################################################################
block1 = Conv2D(F1, (1, kernLength), padding = 'same',
input_shape = (1, Chans, Samples),
use_bias = False, data_format='channels_first')(input1)
block1 = BatchNormalization(axis = 1)(block1)
block1 = DepthwiseConv2D((Chans, 1), use_bias = False,
depth_multiplier = D,
depthwise_constraint = max_norm(1.), data_format='channels_first')(block1)
block1 = BatchNormalization(axis = 1)(block1)
block1 = Activation('elu')(block1)
block1 = AveragePooling2D((1, 2), data_format='channels_first')(block1)
block1 = dropoutType(dropoutRate)(block1)
block2 = SeparableConv2D(F2, (1, 16),
use_bias = False, padding = 'same', data_format='channels_first')(block1)
block2 = BatchNormalization(axis = 1)(block2)
block2 = Activation('elu')(block2)
block2 = AveragePooling2D((1, 4), data_format='channels_first')(block2)
block2 = dropoutType(dropoutRate)(block2)
block3 = Conv2D(F2, (1, 8), padding='same', data_format='channels_first')(block2)
block3 = BatchNormalization(axis=1)(block3)
block3 = Activation('elu')(block3)
block3 = AveragePooling2D((1, 8), data_format='channels_first')(block3)
flatten = Flatten(name = 'flatten')(block3)
# dense = Dense(50,name='dense1', kernel_constraint = max_norm(norm_rate))(flatten)
dense = Dense(1, name = 'out', kernel_constraint = max_norm(norm_rate))(flatten)
softmax = Activation('sigmoid', name = 'sigmoid')(dense)
return Model(inputs=input1, outputs=softmax)
def CNN_model(dropout=0.4, l1=0.0, l2=0.01, chans=64, samples=113):
model = Sequential()
# Layer 1
model.add(Conv2D(256, (2, chans), input_shape=(1, chans, samples), activity_regularizer=l1_l2(l1, l2), name='conv1',
padding='same', data_format='channels_first'))
model.add(BatchNormalization(axis=1))
model.add(DepthwiseConv2D((chans, 4), depth_multiplier = 3, activity_regularizer=l1_l2(l1, l2),
data_format='channels_first'))
model.add(BatchNormalization(axis=1))
model.add(Activation('elu'))
model.add(AveragePooling2D((1, 2), data_format='channels_first', name='pool1'))
model.add(Dropout(dropout))
# Layer 2
model.add(Conv2D(256, (1, 1), padding='same', name='conv2', activity_regularizer=l1_l2(l1, l2), data_format='channels_first'))
model.add(BatchNormalization(axis=1))
model.add(Conv2D(256, (3, 3), padding='same', name='conv3', activity_regularizer=l1_l2(l1, l2), data_format='channels_first',
activation='elu'))
model.add(BatchNormalization(axis=1))
model.add(Activation('elu'))
model.add(MaxPooling2D((1, 2), data_format='channels_first', name='pool2',))
model.add(Dropout(dropout))
# Layer 3
model.add(Conv2D(256, (5, 5), padding='same', name='conv4', activity_regularizer=l1_l2(l1, l2), data_format='channels_first'))
model.add(BatchNormalization(axis=1))
model.add(Activation('elu'))
# Layer 4
model.add(Conv2D(512, (5, 5), padding='same', name='conv5', activity_regularizer=l1_l2(l1, l2), data_format='channels_first'))
model.add(BatchNormalization(axis=1))
model.add(Activation('elu'))
model.add(MaxPooling2D((1, 2), data_format='channels_first', name='pool3',))
model.add(Dropout(dropout))
# Layer 5
model.add(Conv2D(512, (7, 7), padding='same', name='conv6', activity_regularizer=l1_l2(l1, l2), data_format='channels_first'))
model.add(BatchNormalization(axis=1))
model.add(Activation('elu'))
# Layer 6
model.add(Conv2D(512, (7, 7), padding='same', name='conv7', activity_regularizer=l1_l2(l1, l2), data_format='channels_first'))
model.add(BatchNormalization(axis=1))
model.add(Activation('elu'))
model.add(MaxPooling2D((1, 2), data_format='channels_first', name='pool4',))
model.add(Dropout(dropout))
# # Layer 7
# model.add(Conv2D(1024, (5, 5), padding='same', name='conv8', activity_regularizer=l1_l2(l1, l2), data_format='channels_first'))
# model.add(BatchNormalization(axis=1))
# model.add(Activation('elu'))
# # Layer 8
# model.add(Conv2D(1024, (3,3), padding='same', name='conv9', activity_regularizer=l1_l2(l1, l2), data_format='channels_first'))
# model.add(BatchNormalization(axis=1))
# model.add(Activation('elu'))
# model.add(MaxPooling2D((1, 2), data_format='channels_first', name='pool5',))
# model.add(Dropout(dropout))
# Flatten
model.add(Flatten())
# Layer 9
model.add(Dense(16, activation='relu', activity_regularizer=l1_l2(l1, l2)))
model.add(Dropout(dropout))
# Output Layer
model.add(Dense(1, activation='sigmoid'))
return model
```
# Load data and Preprocessing
```
%load_ext autoreload
%autoreload
import numpy as np
from tensorflow.keras import *
import matplotlib.pyplot as plt
from sklearn.pipeline import Pipeline
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
from sklearn import svm
from sklearn.naive_bayes import GaussianNB, BernoulliNB
from sklearn.linear_model import LassoLarsCV
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import ShuffleSplit, cross_val_score, train_test_split
from mne import Epochs, pick_types, events_from_annotations
from mne.channels import read_layout
from mne.io import concatenate_raws, read_raw_edf
from mne.datasets import eegbci
from mne.decoding import CSP
import mne
import pandas as pd
from matplotlib import pyplot as plt
from tensorflow.keras.models import load_model
def download_dataset(subjects, runs):
raws_list = []
for i in subjects:
try:
raws_list.append(eegbci.load_data(i, runs))
except Exception as e:
print(i,' => ' ,e)
return raws_list
```
# Load and Preprocessing functions
```
def load_dataset(subjects, runs):
# check if dataset is downloaded
download_dataset(subjects, runs)
raw = None
for i in subjects[1:]:
for f in eegbci.load_data(i, runs):
if raw is None:
raw = read_raw_edf(f, preload=True)
else:
try:
raw = concatenate_raws([raw, read_raw_edf(f, preload=True)])
except:
print('subject {} failed to concatinate'.format(i))
return raw
def preprocess(raw, event_id, use_filter = True, low_freq=7, high_freq=30, tmin=1, tmax=2):
# strip channel names of "." characters
raw.rename_channels(lambda x: x.strip('.'))
# Apply band-pass filter
if use_filter:
raw.filter(low_freq, high_freq)
events, _ = get_events(raw)
picks = get_picks(raw)
# Read epochs (train will be done only between 1 and 2s)
# Testing will be done with a running classifier
epochs = get_epochs(raw, events, event_id)
epochs_train = epochs.copy().crop(tmin=tmin, tmax=tmax)
labels = epochs_train.events[:, -1] - 2
labels = labels.reshape((labels.shape[0],1))
epochs_data_train = epochs_train.get_data()
epochs_data_train = epochs_data_train.reshape((epochs_data_train.shape[0],1, epochs_data_train.shape[1], epochs_data_train.shape[2]))
return epochs_data_train, labels
def get_events(raw, event_id=dict(T1=2, T2=3)):
return events_from_annotations(raw, event_id=event_id)
def get_epochs(raw,events, event_id):
return Epochs(raw, events, event_id, -1, 4, proj=True, picks=get_picks(raw),
baseline=(None, None), preload=True)
def get_picks(raw):
return pick_types(raw.info, meg=False, eeg=True, stim=False, eog=False,
exclude='bads')
subjects = [i for i in range(1,86)]
runs = [4, 8, 12] # motor Imaginery: left hand vs right hand
raw = load_dataset(subjects, runs)
runs = [3, 7, 11] # Motor execution: left hand vs right hand
# # concatinate two raws
raw = concatenate_raws([raw, load_dataset(subjects, runs)])
# raw.get_data().shape
# Tune parameters
tmin = -0.2
tmax = 0.5
low_freq = 5
high_freq = 60
event_id = dict(right=2, left=3)
epochs_data_train, labels = preprocess(raw, event_id, use_filter=True, low_freq=low_freq, high_freq=high_freq, tmax=tmax, tmin=tmin)
# epochs = get_epochs(raw, get_events(raw)[0], event_id)
# epochs.plot()
X_train, X_test, y_train, y_test = train_test_split(epochs_data_train, labels, test_size=0.1, random_state=42, shuffle=True)
print('shape of train-set-X: {} \nshape of test-set-X: {}'.format(X_train.shape, X_test.shape))
print('shape of train-set-y: {} \nshape of test-set-y: {}'.format(y_train.shape, y_test.shape))
# model = my_net(2, Chans = 64, Samples = X_train.shape[-1],F1=16, D=3, F2=48,
# dropoutRate = 0.0, kernLength = 96,norm_rate = 0.0, dropoutType = 'Dropout')
# model = EEGNet(2, Samples=X_train.shape[-1], kernLength=96, Chans=64, F1=16, D=3, F2=48, dropoutRate=0.0, norm_rate=0)
# model = CNN_model(samples=X_train.shape[-1], l1=0.0, l2=0.000, dropout=0.4)
# model = ResNet(samples=X_train.shape[-1], l1=0.00, l2=0.0)
# model = Effnet((1, 64, X_train.shape[-1]), 1)
model = DeepConvNet(2, Chans = 64, Samples = X_train.shape[-1], dropoutRate = 0.15)
#load saved model
# model = load_model('model_left_vs_right_hand_113acc_76.h5')
model.summary()
# for layer in model.layers:
# layer.trainable = True
optimizer = optimizers.Adam(lr=0.001, decay=1e-5)
model.compile(optimizer , loss=losses.binary_crossentropy , metrics=['acc'])
batch_size = 256
# callback = EarlyStopping('val_loss', patience=300, mode='min', restore_best_weights=True)
earlyStopping = EarlyStopping(monitor='val_loss', patience=40, verbose=0, mode='min')
mcp_save = ModelCheckpoint('mdl_wts.hdf5',verbose=1, save_best_only=True, monitor='val_loss', mode='min')
reduce_lr_loss = ReduceLROnPlateau(monitor='val_loss', factor=0.0001, patience=25, verbose=1, epsilon=1e-4, mode='min')
history = model.fit(X_train, y_train, batch_size=batch_size, epochs=1500, validation_data=(X_test, y_test),
callbacks=[ mcp_save, reduce_lr_loss])
best_model = load_model('mdl_wts.hdf5')
model.evaluate(X_test, y_test)
best_model.evaluate(X_test, y_test)
plt.plot(history.history['acc'])
plt.plot(history.history['loss'])
plt.plot(history.history['val_acc'])
plt.legend(['acc','loss','val_acc'])
subjects = [i for i in range(86,110)]
# [3, 7, 11]
test_epochs, test_labels = preprocess(load_dataset(subjects, [4, 8, 12]), event_id, use_filter=True,low_freq=low_freq,
high_freq=high_freq, tmax=tmax, tmin=tmin)
loss, accuracy = model.evaluate(test_epochs, test_labels)
loss, accuracy = best_model.evaluate(test_epochs, test_labels)
best_model.save('model_left_vs_right_hand_' + str(X_train.shape[-1]) + 'acc_' + str(accuracy) + '.h5')
# Install the PyDrive wrapper & import libraries.
# This only needs to be done once in a notebook.
!pip install -U -q PyDrive
from pydrive.auth import GoogleAuth
from pydrive.drive import GoogleDrive
from google.colab import auth
from oauth2client.client import GoogleCredentials
# Authenticate and create the PyDrive client.
# This only needs to be done once in a notebook.
auth.authenticate_user()
gauth = GoogleAuth()
gauth.credentials = GoogleCredentials.get_application_default()
drive = GoogleDrive(gauth)
# Create & upload a text file.
uploaded = drive.CreateFile({'title': 'Sample file.txt'})
uploaded.SetContentString('Sample upload file content')
uploaded.Upload()
print('Uploaded file with ID {}'.format(uploaded.get('id')))
from tensorflow.keras.models import load_model
# from keras.initializers import glorot_uniform
loaded_model = load_model('model_321.h5')
# loaded_model.summary()
loaded_model.evaluate(X_train, y_train)
loaded_model.evaluate(X_test, y_test)
# from tensorflow.keras.utils import plot_model
# plot_model(model, to_file='model.png', show_shapes=True)
for layer in model.layers:
if type(layer) is Conv2D:
print('CONV2D: filters: {}, kernel_size: {}, padding: {}, strides: {}'.format(layer.filters, layer.kernel_size, layer.padding, layer.strides))
elif type(layer) is SeparableConv2D:
print('SepConv2D: filters: {}, kernel_size: {}, padding: {}, strides: {}'.format(layer.filters, layer.kernel_size, layer.padding, layer.strides))
elif type(layer) is AveragePooling2D:
print('AP2D: pool_size: {}, padding: {}, strides: {}'.format(layer.pool_size,layer.padding, layer.strides))
elif type(layer) is Dropout:
# print(layer.rate)
pass
else:
pass
# print(layer)
K.eval(loaded_model.optimizer.lr)
K.eval(loaded_model.optimizer.decay)
K.eval(loaded_model.non_trainable_variables)
model.weights
loaded_model.weights
```
| github_jupyter |
##### Copyright 2018 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Юникод-строки
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/tutorials/load_data/unicode"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />Смотрите на TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/ru/tutorials/load_data/unicode.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Запустите в Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/ru/tutorials/load_data/unicode.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />Изучайте код на GitHub</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/docs-l10n/site/ru/tutorials/load_data/unicode.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Скачайте ноутбук</a>
</td>
</table>
Note: Вся информация в этом разделе переведена с помощью русскоговорящего Tensorflow сообщества на общественных началах. Поскольку этот перевод не является официальным, мы не гарантируем что он на 100% аккуратен и соответствует [официальной документации на английском языке](https://www.tensorflow.org/?hl=en). Если у вас есть предложение как исправить этот перевод, мы будем очень рады увидеть pull request в [tensorflow/docs](https://github.com/tensorflow/docs) репозиторий GitHub. Если вы хотите помочь сделать документацию по Tensorflow лучше (сделать сам перевод или проверить перевод подготовленный кем-то другим), напишите нам на [docs-ru@tensorflow.org list](https://groups.google.com/a/tensorflow.org/forum/#!forum/docs-ru).
## Введение
Модели обрабатывающие естественные языки, часто имеют дело с разными языками и разными наборами символов. * Unicode * - это стандартная система кодирования, которая используется для представления символов практически всех языков. Каждый символ кодируется с использованием уникального целого числа [кодовой точки](https://en.wikipedia.org/wiki/Code_point) между `0` и` 0x10FFFF`. *Юникод-строка* - это последовательность из нуля или более таких кодовых точек.
Это руководство показывает как представлять юникод-строки в Tensorflow и манипулировать ими используя юникодовские эквиваленты стандартной строковой. Она выделяет юникод-строки в токены на основе обнаружения скрипта.
```
import tensorflow as tf
```
## Тип данных `tf.string`
Базовый TensorFlow `tf.string` `dtype` позволяет вам строить тензоры байт-строк.
Юникод-строки по умолчанию в кодировке utf-8.
```
tf.constant(u"Спасибо 😊")
```
Тензор `tf.string` может содержать байт-строки различной длины поскольку байт-строки обрабатываются как отдельные единицы. Длина строки не включена в размерность тензора.
```
tf.constant([u"Добро", u"пожаловать!"]).shape
```
Замечание: При использовании python при конструировании строк обработка юникода отличается между v2 и v3. В v2, юникод-строки отмечены префиксом "u", как и выше. В v3, строки закодированы в юникоде по умолчанию.
## Представление Юникода
Есть два стандартных способа представления юникод-строк в TensorFlow:
* `string` скаляр — где последовательность кодовых точек закодирована с использованием [набора символов](https://ru.wikipedia.org/wiki/%D0%9D%D0%B0%D0%B1%D0%BE%D1%80_%D1%81%D0%B8%D0%BC%D0%B2%D0%BE%D0%BB%D0%BE%D0%B2).
* `int32` вектор — где каждая позиция содержит единственную кодовую точку.
Например, следующие три значения все представляют юникод-строку `"语言处理"` (что значит "обработка языка" на китайском):
```
# Юникод-строки, представленные как UTF-8 закодированные строки скаляры.
text_utf8 = tf.constant(u"语言处理")
text_utf8
# Юникод-строки представленные как UTF-16-BE закодированные строки скаляры.
text_utf16be = tf.constant(u"语言处理".encode("UTF-16-BE"))
text_utf16be
# Юникод строки представленные как векторы юникодовских кодовых точек.
text_chars = tf.constant([ord(char) for char in u"语言处理"])
text_chars
```
### Конвертация между представлениями
TensorFlow предоставляет операции для конвертации между этими различными представлениями:
* `tf.strings.unicode_decode`: Конвертирует закодированную строку скаляр в вектор кодовых точек.
* `tf.strings.unicode_encode`: Конвертирует вектор кодовых точек в закодированную строку скаляр.
* `tf.strings.unicode_transcode`: Конвертирует строку скаляр в другую кодировку.
```
tf.strings.unicode_decode(text_utf8,
input_encoding='UTF-8')
tf.strings.unicode_encode(text_chars,
output_encoding='UTF-8')
tf.strings.unicode_transcode(text_utf8,
input_encoding='UTF8',
output_encoding='UTF-16-BE')
```
### Размерности пакета
При декодировании нескольких строк размер символов в каждой строке может не совпадать, Возвращаемый результат это [`tf.RaggedTensor`](../../guide/ragged_tensor.ipynb), где длина самого внутреннего измерения меняется в зависимости от количества символов в каждой строке:
```
# Пакет юнокод-строк каждая из которых представлена в виде строки в юникод-кодировке.
batch_utf8 = [s.encode('UTF-8') for s in
[u'hÃllo', u'What is the weather tomorrow', u'Göödnight', u'😊']]
batch_chars_ragged = tf.strings.unicode_decode(batch_utf8,
input_encoding='UTF-8')
for sentence_chars in batch_chars_ragged.to_list():
print(sentence_chars)
```
Вы можете использовать `tf.RaggedTensor` напрямую, или конвертировать его в плотный `tf.Tensor` с паддингом или в `tf.SparseTensor` используя методы `tf.RaggedTensor.to_tensor` и `tf.RaggedTensor.to_sparse`.
```
batch_chars_padded = batch_chars_ragged.to_tensor(default_value=-1)
print(batch_chars_padded.numpy())
batch_chars_sparse = batch_chars_ragged.to_sparse()
```
При кодировании нескольких строк одинаковой длины `tf.Tensor` может быть использован в качестве входных данных:
```
tf.strings.unicode_encode([[99, 97, 116], [100, 111, 103], [ 99, 111, 119]],
output_encoding='UTF-8')
```
При кодировании нескольких строк различной длины нужно использовать `tf.RaggedTensor` в качестве входных данных:
```
tf.strings.unicode_encode(batch_chars_ragged, output_encoding='UTF-8')
```
Если у вас тензор с несколькими строками с паддингом или в разреженном формате, то ковертируйте его в `tf.RaggedTensor` перед вызовом `unicode_encode`:
```
tf.strings.unicode_encode(
tf.RaggedTensor.from_sparse(batch_chars_sparse),
output_encoding='UTF-8')
tf.strings.unicode_encode(
tf.RaggedTensor.from_tensor(batch_chars_padded, padding=-1),
output_encoding='UTF-8')
```
## Операции юникода
### Длина символа
Операция `tf.strings.length` имеет параметр `unit`, который показывает как должна быть посчитана длина. По умолчанию размер `unit` равен `"BYTE"`, но он может быть установлен с другим значением, таким как `"UTF8_CHAR"` или `"UTF16_CHAR"`, чтобы определить число кодовых точек в каждой закодированой `string`.
```
# Заметьте что последний символ занимает до 4 байтов в UTF8.
thanks = u'Thanks 😊'.encode('UTF-8')
num_bytes = tf.strings.length(thanks).numpy()
num_chars = tf.strings.length(thanks, unit='UTF8_CHAR').numpy()
print('{} bytes; {} UTF-8 characters'.format(num_bytes, num_chars))
```
### Подстроки символов
Аналогично у операции `tf.strings.substr` есть параметр "`unit`", который используется для определения смещений параметров "`pos`" и "`len`".
```
# по умолчанию: unit='BYTE'. С len=1, мы возвращаем один байт.
tf.strings.substr(thanks, pos=7, len=1).numpy()
# Установив unit='UTF8_CHAR', мы возвратим один символ, размер которого в этом случае
# 4 байта.
print(tf.strings.substr(thanks, pos=7, len=1, unit='UTF8_CHAR').numpy())
```
### Разбиение юникод-строки
Операция `tf.strings.unicode_split` разбивает юникод-строки в подстроки отдельных символов:
```
tf.strings.unicode_split(thanks, 'UTF-8').numpy()
```
### Смещения байтов для символов
Чтобы выровнять тензор символа порожденный `tf.strings.unicode_decode` с оригинальной строкой, полезно знать смещение начала каждого символа. Метод `tf.strings.unicode_decode_with_offsets` аналогичен `unicode_decode`, за исключением того, что он возвращает второй тензор содержащий размер отступа от начала для каждого символа.
```
codepoints, offsets = tf.strings.unicode_decode_with_offsets(u"🎈🎉🎊", 'UTF-8')
for (codepoint, offset) in zip(codepoints.numpy(), offsets.numpy()):
print("At byte offset {}: codepoint {}".format(offset, codepoint))
```
## Юникод скрипты
Каждая кодовая точка принадлежит коллекции символов известной как [система письма](https://en.wikipedia.org/wiki/Script_%28Unicode%29) . Система письма полезна для определения того, какому языку может принадлежать символ. Например, зная что 'Б' из кириллицы указывает на то, что современный текст содержащий этот символ скорее всего из славянского языка, такого как русский или украинский.
В TensorFlow есть операция `tf.strings.unicode_script` для определения какой системе письма принадлежит данная кодовая точка. Коды систем письма это `int32` числа соответствующие [Международным компонентам для
юникода](http://site.icu-project.org/home) (ICU) [`UScriptCode`](http://icu-project.org/apiref/icu4c/uscript_8h.html) значения.
```
uscript = tf.strings.unicode_script([33464, 1041]) # ['芸', 'Б']
print(uscript.numpy()) # [17, 8] == [USCRIPT_HAN, USCRIPT_CYRILLIC]
```
Операция `tf.strings.unicode_script` может быть также применена к многомерному `tf.Tensor`s или кодовым точкам `tf.RaggedTensor`:
```
print(tf.strings.unicode_script(batch_chars_ragged))
```
## Пример: Простая сегментация
Сегментация это задача разбиения текста на словоподобные юниты. Часто это легко когда испольуются символы пробела для отделения слов, но некоторые языки (например китайский и японский) не используют пробелы, а некоторые языки (например немецкий) содержат длинные соединения, которые должны быть разделены для анализа их значений. В веб текстах, различные языки и скрипты часто перемешаны между собой , как например в "NY株価" (New York Stock Exchange).
Мы можем выполнить грубую сегментацию (без реализации каких-либо моделей ML), используя изменения систем письма для приблизительного определения границ слов. Это будет работать для строк наподобие вышеприведенного примера "NY株価". Это также будет работать для всех языков, которые используют пробелы, так как символы пробела в различных системах письма все классифицируются как USCRIPT_COMMON, специальный код, который отличается от кода любого актуального текста.
```
# dtype: string; shape: [num_sentences]
#
# Предложения для обработки. Поменяйте эту строку чтобы попробовать разные входные данные!
sentence_texts = [u'Hello, world.', u'世界こんにちは']
```
Сперва мы декодируем предложения в кодовые точки, и определим идентификатор системы письма для каждого символа.
```
# dtype: int32; shape: [num_sentences, (num_chars_per_sentence)]
#
# sentence_char_codepoint[i, j] кусок кода для j-го символа
# в i-м предложении.
sentence_char_codepoint = tf.strings.unicode_decode(sentence_texts, 'UTF-8')
print(sentence_char_codepoint)
# dtype: int32; shape: [num_sentences, (num_chars_per_sentence)]
#
# sentence_char_scripts[i, j] код системы письма для j-го символа в
# i-м предложении.
sentence_char_script = tf.strings.unicode_script(sentence_char_codepoint)
print(sentence_char_script)
```
Далее мы используем эти идентификаторы систем письма чтобы определить куда должны быть добавлены границы слов. Мы добавим границу слова в начало каждого предложения и для каждого символа чья система письма отличается от предыдущего символа:
```
# dtype: bool; shape: [num_sentences, (num_chars_per_sentence)]
#
# sentence_char_starts_word[i, j] является True если j'th символ в i'th
# предложении является началом слова.
sentence_char_starts_word = tf.concat(
[tf.fill([sentence_char_script.nrows(), 1], True),
tf.not_equal(sentence_char_script[:, 1:], sentence_char_script[:, :-1])],
axis=1)
# dtype: int64; shape: [num_words]
#
# word_starts[i] это индекс символа начинающего i-е слово (в
# выпрямленном списке символов всех предложений).
word_starts = tf.squeeze(tf.where(sentence_char_starts_word.values), axis=1)
print(word_starts)
```
Затем мы можем использовать эти сдвиги от начала для построения `RaggedTensor` содержащего список слов из всех пакетов:
```
# dtype: int32; shape: [num_words, (num_chars_per_word)]
#
# word_char_codepoint[i, j] is the кодовая точка для j-го символа в
# i-м слове.
word_char_codepoint = tf.RaggedTensor.from_row_starts(
values=sentence_char_codepoint.values,
row_starts=word_starts)
print(word_char_codepoint)
```
И наконец, мы можем сегментировать кодовые точки слов `RaggedTensor` обратно в предложения:
```
# dtype: int64; shape: [num_sentences]
#
# sentence_num_words[i] число слов в i'th предложении.
sentence_num_words = tf.reduce_sum(
tf.cast(sentence_char_starts_word, tf.int64),
axis=1)
# dtype: int32; shape: [num_sentences, (num_words_per_sentence), (num_chars_per_word)]
#
# sentence_word_char_codepoint[i, j, k] это кодовая точка для k-го символа
# в j-м слове i-го предложения.
sentence_word_char_codepoint = tf.RaggedTensor.from_row_lengths(
values=word_char_codepoint,
row_lengths=sentence_num_words)
print(sentence_word_char_codepoint)
```
Чтобы сделать итоговый результат проще для чтения, мы можем закодировать его обратно в UTF-8 строки:
```
tf.strings.unicode_encode(sentence_word_char_codepoint, 'UTF-8').to_list()
```
| github_jupyter |
```
import sklearn.datasets
import re
from sklearn.decomposition import PCA
from sklearn.manifold import TSNE
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.feature_extraction.text import TfidfTransformer
from sklearn.feature_extraction.text import HashingVectorizer
from sklearn.pipeline import Pipeline
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# default parameters
sns.set()
```
functions for preprocessing
```
# clear string
def clearstring(string):
string = re.sub('[^A-Za-z0-9 ]+', '', string)
string = string.split(' ')
string = filter(None, string)
string = [y.strip() for y in string]
string = ' '.join(string)
return string
# because of sklean.datasets read a document as a single element
# so we want to split based on new line
def separate_dataset(trainset):
datastring = []
datatarget = []
for i in range(len(trainset.data)):
data_ = trainset.data[i].split('\n')
# python3, if python2, just remove list()
data_ = list(filter(None, data_))
for n in range(len(data_)):
data_[n] = clearstring(data_[n])
datastring += data_
for n in range(len(data_)):
datatarget.append(trainset.target[i])
return datastring, datatarget
trainset = sklearn.datasets.load_files(container_path = 'local', encoding = 'UTF-8')
trainset.data, trainset.target = separate_dataset(trainset)
from sklearn.cross_validation import train_test_split
# default colors from seaborn
current_palette = sns.color_palette(n_colors = len(trainset.filenames))
# visualize 5% of our data
_, texts, _, labels = train_test_split(trainset.data, trainset.target, test_size = 0.05)
# bag-of-word
bow = CountVectorizer().fit_transform(texts)
#tf-idf, must get from BOW first
tfidf = TfidfTransformer().fit_transform(bow)
#hashing, default n_features, probability cannot divide by negative
hashing = HashingVectorizer(non_negative = True).fit_transform(texts)
```
Visualization for BOW for both PCA and TSNE
```
# size of figure is 1000 x 500 pixels
plt.figure(figsize = (15, 5))
plt.subplot(1, 2, 1)
composed = PCA(n_components = 2).fit_transform(bow.toarray())
for no, _ in enumerate(np.unique(trainset.target_names)):
plt.scatter(composed[np.array(labels) == no, 0], composed[np.array(labels) == no, 1], c = current_palette[no],
label = trainset.target_names[no])
plt.legend()
plt.title('PCA')
plt.subplot(1, 2, 2)
composed = TSNE(n_components = 2).fit_transform(bow.toarray())
for no, _ in enumerate(np.unique(trainset.target_names)):
plt.scatter(composed[np.array(labels) == no, 0], composed[np.array(labels) == no, 1], c = current_palette[no],
label = trainset.target_names[no])
plt.legend()
plt.title('TSNE')
plt.show()
# size of figure is 1000 x 500 pixels
plt.figure(figsize = (15, 5))
plt.subplot(1, 2, 1)
composed = PCA(n_components = 2).fit_transform(tfidf.toarray())
for no, _ in enumerate(np.unique(trainset.target_names)):
plt.scatter(composed[np.array(labels) == no, 0], composed[np.array(labels) == no, 1], c = current_palette[no],
label = trainset.target_names[no])
plt.legend()
plt.title('PCA')
plt.subplot(1, 2, 2)
composed = TSNE(n_components = 2).fit_transform(tfidf.toarray())
for no, _ in enumerate(np.unique(trainset.target_names)):
plt.scatter(composed[np.array(labels) == no, 0], composed[np.array(labels) == no, 1], c = current_palette[no],
label = trainset.target_names[no])
plt.legend()
plt.title('TSNE')
plt.show()
```
Ops, memory error on hashing
| github_jupyter |
```
import pandas as pd
import numpy as np
import json
from bs4 import BeautifulSoup
import requests
import matplotlib.pyplot as plt
```
## Data Acquisition - Select Cryptocurrencies of Interest
1. Get all coin data from http://coincap.io/front
2. Select coins with Market cap more than selected threshold
```
all_coins_dict = json.loads(BeautifulSoup(
requests.get('http://coincap.io/front').content, "html.parser").prettify())
all_coins_df = pd.DataFrame(all_coins_dict)
coins_by_mcap = all_coins_df[all_coins_df.mktcap > 5e9]
coin_portfolio = coins_by_mcap['short']
print("Portfolio coins with MCAP > 5 Billion :\n",coin_portfolio.values)
```
## Acquire Data - Get Historical Prices
1. Get all the price data for a particular coin from http://coincap.io/history/180day/<coin>
2. Create a master dataframe to hold historical price data for all the selected coins
```
#Create a DataFrame that will hold all the price & data for all the selected coins
combined_df = pd.DataFrame()
#Loop thru all the coins in the portfolio & get their historical prices. (180 days)
for coin in coin_portfolio:
dic_t = json.loads(BeautifulSoup(
requests.get('http://coincap.io/history/180day/'+coin).
content, "html.parser").prettify())
prices = dic_t.get('price')
coindf = pd.DataFrame.from_records(prices,columns=['time','price'])
coindf['coin'] = coin
combined_df = combined_df.append(coindf, ignore_index=True)
```
## Data Wrangling
Clean the historical price data, and find the returns and covariance
```
#Change the time formart
combined_df['time'] = pd.to_datetime(combined_df['time'],unit='ms')
combined_df['time'] = [d.date() for d in combined_df['time']]
operational_df = combined_df.groupby(['time', 'coin'],as_index=False)[['price']].mean()
operational_df = operational_df.set_index('time')
pivoted_portfolio = operational_df.pivot(columns='coin')
# get covariance & returns of the coin - daily & for the period
daily_returns = pivoted_portfolio.pct_change()
period_returns = daily_returns.mean()*180
daily_covariance = daily_returns.cov()
period_covariance = daily_covariance*180
```
## Optimize portfolio
```
p_returns, p_volatility, p_sharpe_ratio, p_coin_weights=([] for i in range(4))
# portfolio combinations to probe
number_of_cryptoassets = len(coin_portfolio)
number_crypto_portfolios = 50000
# for each portoflio, get returns, risk and weights
for a_crypto_portfolio in range(number_crypto_portfolios):
weights = np.random.random(number_of_cryptoassets)
weights /= np.sum(weights)
#print(weights)
returns = np.dot(weights, period_returns)*100
#print(weights)
volatility = np.sqrt(np.dot(weights.T, np.dot(period_covariance, weights)))*100
p_sharpe_ratio.append(returns/volatility)
p_returns.append(returns)
p_volatility.append(volatility)
p_coin_weights.append(weights)
# a dictionary for Returns and Risk values of each portfolio
portfolio = {'volatility': p_volatility,
'sharpe_ratio': p_sharpe_ratio, 'returns': p_returns}
# extend original dictionary to accomodate each ticker and weight in the portfolio
for counter,symbol in enumerate(coin_portfolio):
portfolio[symbol+'-%'] = [Weight[counter] for Weight in p_coin_weights]
# make a nice dataframe of the extended dictionary
df = pd.DataFrame(portfolio)
```
## Visualize - Efficient Frontier
```
order_cols = ['returns', 'volatility', 'sharpe_ratio']+[coin+'-%' for coin in coin_portfolio]
df = df[order_cols]
sharpe_portfolio = df.loc[df['sharpe_ratio'] == df['sharpe_ratio'].max()]
min_variance_port = df.loc[df['volatility'] == df['volatility'].min()]
max_returns_port = df.loc[df['returns'] == df['returns'].max()]
plt.style.use('seaborn-dark')
df.plot.scatter(x='volatility', y='returns', c='sharpe_ratio', cmap='PRGn_r',
edgecolors='black', figsize=(10, 7), grid=True)
plt.xlabel('Volatility (Std. Deviation)')
plt.ylabel('Expected Returns')
plt.title('Efficient Frontier')
plt.show()
print('****Portfolio llocations*****')
print(sharpe_portfolio.T)
```
| github_jupyter |
```
import os
os.environ['CUDA_VISIBLE_DEVICES']='0'
from fastai import *
from fastai.vision import *
from batch_norm_vgg import VGG, make_layers, make_layers_BN
from batch_norm_callbacks import ICS, AccuracyList, AccuracyValidationList
import matplotlib.pyplot as plt
import time
data_path = untar_data(URLs.CIFAR)
data_path.ls()
data = ImageDataBunch.from_folder(
data_path, valid='test', size=32, bs=128
)#.split_by_idxs(
# train_idx=[0, 1, 2, 3, 4, 5, 5000, 5001, 5002,5003,5004,5005], valid_idx=[0, 1, 2, 3, 4, 5, 1000,1001, 1002,1003,1004,1005]
# ).label_from_folder(
# ).databunch(bs=2, num_workers=1, no_check=True)
criterion = nn.CrossEntropyLoss()
optimizer = optim.SGD
learn_rate = 0.01
vgg = VGG(make_layers(batch_norm=True), num_classes=data.c).cuda()
learn_vgg = Learner(data, vgg, opt_func=optimizer,
loss_func=criterion,
true_wd=False,
wd=0.,
bn_wd=False,
metrics=accuracy)
accList_BN = AccuracyList(learn_vgg)
icsList_BN = ICS(learn_vgg, num_classes=data.c, bn=True )
accListVal_BN = AccuracyValidationList()
learn_vgg.metrics.append(accListVal_BN)
learn_vgg.metrics
learn_vgg.fit(30, lr=learn_rate, callbacks=[accList_BN, icsList_BN])
accListVal_BN.val_accs
# plt.plot(accList.accs)
plt.plot(icsList_BN.ics_values)
plt.plot(icsList_BN.cos_values)
vgg = VGG(make_layers(batch_norm=False), num_classes=data.c).cuda()
learn_vgg = Learner(data, vgg, opt_func=optimizer,
loss_func=criterion,
true_wd=False,
wd=0.,
bn_wd=False,
metrics=accuracy)
accList_noBN = AccuracyList(learn_vgg)
icsList_noBN = ICS(learn_vgg, num_classes=data.c, bn=False )
accListVal_noBN = AccuracyValidationList()
learn_vgg.metrics.append(accListVal_noBN)
learn_vgg.fit(30, lr=learn_rate, callbacks=[accList_noBN, icsList_noBN])
accListVal_noBN.val_accs
fig = plt.figure(figsize=(20, 10))
ax = fig.add_subplot(111)
plt.ylim(top=1.2, bottom=0)
plt.ylabel("Accuracy", fontsize=15)
plt.xlabel("Steps", fontsize=15)
plt.title("Test Accuracy", fontsize=15)
plt.plot(accListVal_noBN.val_accs, linewidth=3, label="Standard")
plt.plot(accListVal_BN.val_accs, linewidth=3, alpha=0.80, label="Batch Norm")
ax.legend(fontsize=15)
plt.savefig('VAL_ACC.png')
fig = plt.figure(figsize=(20, 10))
ax = fig.add_subplot(111)
plt.ylim(top=1.2, bottom=0)
plt.ylabel("Accuracy", fontsize=15)
plt.xlabel("Steps", fontsize=15)
plt.title("Accuracy", fontsize=15)
plt.plot(accList_noBN.accs, linewidth=0.5, label="Standard")
plt.plot(accList_BN.accs, linewidth=0.5, alpha=0.80, label="Batch Norm")
ax.legend(fontsize=15)
plt.savefig('ACC.png')
fig = plt.figure(figsize=(20, 10))
ax = fig.add_subplot(111)
plt.ylim(top=30, bottom=0.01)
plt.ylabel("L2-difference", fontsize=15)
plt.xlabel("Steps", fontsize=15)
plt.title("ICS layer 5", fontsize=15)
plt.plot(icsList_noBN.ics_values, linewidth=0.5, label="Standard")
plt.plot(icsList_BN.ics_values, linewidth=0.5, alpha=0.80, label="Batch Norm")
ax.legend(fontsize=15)
plt.savefig('ICS.png')
fig = plt.figure(figsize=(20, 10))
ax = fig.add_subplot(111)
plt.ylim(top=1, bottom=-0.1)
plt.ylabel("Cos angle", fontsize=15)
plt.xlabel("Steps", fontsize=15)
plt.title("Cos angle layer 5", fontsize=15)
plt.plot([abs(e) for e in icsList_noBN.cos_values], linewidth=0.5, label="Standard")
plt.plot([abs(e) for e in icsList_BN.cos_values], linewidth=0.5, alpha=0.80, label="Batch Norm")
ax.legend(fontsize=15)
plt.savefig('COS.png')
```
| github_jupyter |
# Green's function of an atomic chain
This example demonstrates a computation of the retarded Green's function function of an atomic chain. The chain consists of identical atoms of an abstract chemical element, say an element "A".
```
import sys
import numpy as np
import matplotlib.pyplot as plt
import tb
import negf
xyz_file = """1
H cell
A1 0.0000000000 0.0000000000 0.0000000000
"""
a = tb.Orbitals('A')
```
Let us assume each atomic site has one s-type orbital and the energy level of -0.7 eV. The coupling matrix element equals -0.5 eV.
```
a.add_orbital('s', -0.7)
tb.Orbitals.orbital_sets = {'A': a}
tb.set_tb_params(PARAMS_A_A={'ss_sigma': -0.5})
```
With all these parameters we can create an instance of the class Hamiltonian. The distance between nearest neighbours is set to 1.1 A.
```
h = tb.Hamiltonian(xyz=xyz_file, nn_distance=1.1).initialize()
```
Now we need to set periodic boundary conditions with a one-dimensional unit cell and lattice constant of 1 A.
```
h.set_periodic_bc([[0, 0, 1.0]])
h_l, h_0, h_r = h.get_hamiltonians()
energy = np.linspace(-3.5, 2.0, 500)
sgf_l = []
sgf_r = []
for E in energy:
L, R = negf.surface_greens_function(E, h_l, h_0, h_r)
sgf_l.append(L)
sgf_r.append(R)
sgf_l = np.array(sgf_l)
sgf_r = np.array(sgf_r)
num_sites = h_0.shape[0]
gf = np.linalg.pinv(np.multiply.outer(energy+0.001j, np.identity(num_sites)) - h_0 - sgf_l - sgf_r)
dos = -np.trace(np.imag(gf), axis1=1, axis2=2)
tr = np.zeros((energy.shape[0]), dtype=complex)
for j, E in enumerate(energy):
gf0 = np.matrix(gf[j, :, :])
gamma_l = 1j * (np.matrix(sgf_l[j, :, :]) - np.matrix(sgf_l[j, :, :]).H)
gamma_r = 1j * (np.matrix(sgf_r[j, :, :]) - np.matrix(sgf_r[j, :, :]).H)
tr[j] = np.real(np.trace(gamma_l * gf0 * gamma_r * gf0.H))
dos[j] = np.real(np.trace(1j * (gf0 - gf0.H)))
fig, ax = plt.subplots(1, 2)
ax[0].plot(energy, dos)
ax[0].set_xlabel('Energy (eV)')
ax[0].set_ylabel('DOS')
ax[0].set_title('Density of states')
ax[1].plot(energy, tr)
ax[1].set_xlabel('Energy (eV)')
ax[1].set_ylabel('Transmission coefficient (a.u)')
ax[1].set_title('Transmission')
fig.tight_layout()
plt.show()
ax = plt.axes()
ax.set_title('Surface self-energy of the semi-infinite chain')
ax.plot(energy, np.real(np.squeeze(sgf_l)))
ax.plot(energy, np.imag(np.squeeze(sgf_l)))
ax.set_xlabel('Energy (eV)')
ax.set_ylabel('Self-energy')
plt.show()
```
| github_jupyter |
### Calculates price-equilibrium in the market for blockchain records, with and without the lightning network.
### Includes symbolic calculations and plots for specific parameter values.
### Focuses on symmetric transfer-rate and power-law transfer-size.
```
import numpy as np
import sympy
sympy.init_printing(use_unicode=True)
from sympy import *
from sympy.solvers import solve
from IPython.display import display
from typing import Callable
from sympy.utilities.lambdify import lambdify, implemented_function
%matplotlib inline
import matplotlib.pyplot as plt
def simplified(exp, title=None):
simp = simplify(exp)
if simplified.LOG:
if title: display(title,simp)
else: display(simp)
return simp
simplified.LOG = True
def firstOrderCondition(exp, var):
diffExp = simplified(diff(exp, var))
solutions = solve(diffExp, var)
if firstOrderCondition.LOG:
display(solutions)
return solutions
firstOrderCondition.LOG = True
class Result(object): # a class for holding results of calculations
def __repr__(self): return self.__dict__.__repr__()
def display(self):
for k,v in sorted(self.__dict__.items()):
display(k,v)
def subs(self, params):
ans = Result()
for k,v in sorted(self.__dict__.items()):
if hasattr(v,"subs"):
ans.__dict__[k] = v.subs(params).evalf()
else:
ans.__dict__[k] = v
return ans
a,p,r,b,vmax,zmin,zmax,beta = symbols('a \\phi r z v_{\max} z_{\min} z_{\max} \\beta', positive=True,finite=True,real=True)
w,T,D,L,n,Supply = symbols('w T \\Delta \\ell n \\tau', positive=True,finite=True,real=True)
params = {
L: 10, # total transfers per day
D: 6, # delta transfers per day
beta: 0.01, # value / transfer-size
r: 4/100/365, # interest rate per day
a: 1.1, # records per reset tx
Supply: 288000, # records per day
zmin: 0.001, # min transfer size (for power law distribution)
zmax: 1, # max transfer size (for uniform distribution)
}
tNL = 27 * a * r**2 / L**2 / beta**3 * p
tNB = p/beta
tLB = L*p/(27*a*r**2)**(Rational(1,2))
(tNL, tNL.subs(params), tNB, tNB.subs(params), tLB, tLB.subs(params).evalf())
```
# Demand curves
```
### Calculate the demand-curves assuming power-law distribution:
def calculateDemandsPowerlaw():
result = Result()
result.demandWithoutLightning = Piecewise(
(L, tNB < zmin),
(L*zmin/tNB, True))
result.demandWithLightning = Piecewise(
(L, tLB < zmin),
(3 * zmin * (L*a*r**2/p**2)**Rational(1,3)*(zmin**Rational(-1,3)-tLB**Rational(-1,3)) + L*zmin/tLB, tNL < zmin),
(3 * zmin * (L*a*r**2/p**2)**Rational(1,3)*( tNL**Rational(-1,3)-tLB**Rational(-1,3)) + L*zmin/tLB, True))
result.txsBlockchain = Piecewise(
(L, tLB < zmin),
(L*zmin/tLB, True))
result.txsLightning = Piecewise(
(0, tLB < zmin),
(L*(1-zmin/tLB), tNL < zmin),
(L*(zmin/tNL-zmin/tLB), True))
result.txsTotal = result.txsBlockchain + result.txsLightning
return result
demandsPowerlaw = calculateDemandsPowerlaw()
demandsPowerlaw.display()
demandsPowerlawNumeric = demandsPowerlaw.subs(params)
demandsPowerlawNumeric.display()
```
# Price curves
```
Demand2 = 3 * zmin * (L*a*r**2/p**2)**Rational(1,3)*(zmin**Rational(-1,3)-tLB**Rational(-1,3)) + L*zmin/tLB
Price2 = simplified(solve (n/2*Demand2 - Supply, p)[0])
Condition2a = simplified(solve(tLB.subs(p, Price2) - zmin,n)[0])
Condition2b = simplified(solve(zmin - tNL.subs(p, Price2),n)[0])
Demand3 = 3 * zmin * (L*a*r**2/p**2)**Rational(1,3)*( tNL**Rational(-1,3)-tLB**Rational(-1,3))
Price3 = simplified(solve (n/2*Demand3 - Supply, p)[0])
Condition3a = simplified(solve(tNL.subs(p, Price2) - zmin,n)[0])
### Calculate the demand-curves assuming power-law distribution:
def calculatePricesPowerlaw():
result = Result()
result.priceWithoutLightning = Piecewise(
(0, n/2 < Supply/L),
(n*L*beta*zmin/2/Supply, True))
result.priceWithLightning = Piecewise(
(0, n/2 < Supply/L),
(3*sqrt(6)*r*zmin/4/Supply**(3/2)*sqrt(L*a)*n**(3/2), n/2 < beta**2*L*Supply/27/a/r**2),
(n*zmin/2/Supply*(beta*L - 3*sqrt(3*a)*r), True))
# result.txsBlockchain = Piecewise(
# (L, tLB < zmin),
# (L*zmin/tLB, True))
# result.txsLightning = Piecewise(
# (0, tLB < zmin),
# (L*(1-zmin/tLB), tNL < zmin),
# (L*(zmin/tNL-zmin/tLB), True))
# result.txsTotal = result.txsBlockchain + result.txsLightning
return result
pricesPowerlaw = calculatePricesPowerlaw()
pricesPowerlaw.display()
pricesPowerlawNumeric = pricesPowerlaw.subs(params)
pricesPowerlawNumeric.display()
```
# Plots
```
plotSymmetric = True
plotAsymmetric = False
linewidth = 3
def plotSymbolic(xRange, yExpression, xVariable, style, label):
plt.plot(xRange, [yExpression.subs(xVariable,xValue) for xValue in xRange], style, label=label, linewidth=linewidth)
def plotDemandCurves(priceRange, demandWithoutLightning, demandWithLightning):
"""
Plot the expected demand of a single pair vs. the price.
"""
global plotSymmetric, plotAsymmetric
plotSymbolic(priceRange, demandWithoutLightning, p, "r.",label="w.o. lightning")
plotSymbolic(priceRange, demandWithLightning, p, "b--",label="with lightning")
plt.gca().set_ylim(-1,11)
plt.xlabel("blockchain fee $\\phi$ [bitcoins]")
plt.ylabel("Demand of a single pair [records/day]")
plt.legend(loc=0)
def plotTxsCurves(priceRange, demandWithoutLightning, demandWithLightning):
"""
Plot the expected transfer-count of a single pair vs. the price.
"""
plotSymbolic(priceRange, demandWithoutLightning, p, "r.",label="w.o. lightning")
plotSymbolic(priceRange, demandWithLightning.txsBlockchain, p, "g-.",label="with lightning; blockchain")
plotSymbolic(priceRange, demandWithLightning.txsLightning, p, "b--",label="with lightning; lightning")
plotSymbolic(priceRange, demandWithLightning.txsTotal, p, "k-",label="with lightning; total")
plt.gca().set_ylim(-1,11)
plt.xlabel("blockchain fee $\\phi$ [bitcoins]")
plt.ylabel("# Transactions per day")
plt.legend(loc=0)
def plotPriceCurves(nRange, priceWithoutLightning, priceWithLightning):
"""
Plot the price vs. the number of users.
"""
global plotSymmetric, plotAsymmetric
plotSymbolic(nRange, priceWithoutLightning, n, "r.",label="w.o. lightning")
plotSymbolic(nRange, priceWithLightning, n, "b--",label="with lightning")
plt.xlabel("Number of users $n$")
plt.ylabel("Market-equilibrium price $\\phi$ [bitcoins/record]")
plt.legend(loc=0)
def plotMarketTxsCurves(nRange, demandWithoutLightning, demandWithLightning, priceWithoutLightning, priceWithLightning):
"""
Plot the expected transfer-count in the entire market vs. the number of users.
"""
plotSymbolic(nRange, (n/2)*demandWithoutLightning.subs(p,priceWithoutLightning), n, "r.",label="w.o. lightning")
plotSymbolic(nRange, (n/2)*demandWithLightning.txsBlockchain.subs(p,priceWithLightning), n, "g-.",label="with lightning; blockchain")
plotSymbolic(nRange, (n/2)*demandWithLightning.txsLightning.subs (p,priceWithLightning), n, "b--",label="with lightning; lightning")
plotSymbolic(nRange, (n/2)*demandWithLightning.txsTotal.subs (p,priceWithLightning), n, "k-",label="with lightning; total")
plt.xlabel("Number of users $n$")
plt.ylabel("# Transactions per day")
plt.legend(loc=0)
def plotSymbolic3(xRange, yExpression, xVariable, style, label):
plt.plot(xRange, [yExpression.subs(xVariable,xValue)*params[Supply] for xValue in xRange], style, label=label)
def plotRevenueCurves(nRange, priceWithoutLightning, priceAsymmetric, priceSymmetric):
global plotSymmetric, plotAsymmetric
plotSymbolic3(nRange, priceWithoutLightning, n, "r-",label="no lightning")
if plotAsymmetric and priceAsymmetric:
plotSymbolic3(nRange, priceAsymmetric, n, "b.",label="asymmetric")
if plotSymmetric and priceSymmetric:
plotSymbolic3(nRange, priceSymmetric, n, "g--",label="symmetric")
plt.xlabel("Number of users $n$")
plt.ylabel("Miners' revenue [bitcoins/day]")
plt.legend(loc=0)
priceRange = np.linspace(0,1e-6,200)
plotDemandCurves(priceRange, demandsPowerlawNumeric.demandWithoutLightning, demandsPowerlawNumeric.demandWithLightning)
plt.title("Demand curves, power-law-distributed transfer-size")
plt.axes().get_xaxis().set_visible(False)
plt.savefig('../graphs/demand-curves-powerlaw-small-price.pdf', format='pdf', dpi=1000); plt.show()
plotTxsCurves(priceRange, demandsPowerlawNumeric.demandWithoutLightning, demandsPowerlawNumeric)
plt.savefig('../graphs/txs-pair-powerlaw-small-price.pdf', format='pdf', dpi=1000); plt.show()
priceRange = np.linspace(0,2.4e-5,160)
plotDemandCurves(priceRange, demandsPowerlawNumeric.demandWithoutLightning, demandsPowerlawNumeric.demandWithLightning)
plt.title("Demand curves, power-law-distributed transfer-size")
#plt.axes().get_xaxis().set_visible(False)
plt.savefig('../graphs/demand-curves-powerlaw-medium-price.pdf', format='pdf', dpi=1000); plt.show()
plotTxsCurves(priceRange, demandsPowerlawNumeric.demandWithoutLightning, demandsPowerlawNumeric)
plt.savefig('../graphs/txs-pair-powerlaw-medium-price.pdf', format='pdf', dpi=1000); plt.show()
priceRange = np.linspace(0,0.01,300)
plotDemandCurves(priceRange, demandsPowerlawNumeric.demandWithoutLightning, demandsPowerlawNumeric.demandWithLightning)
plt.title("Demand curves, power-law-distributed transfer-size")
plt.axes().get_xaxis().set_visible(False)
plt.gca().set_ylim(-0.01,0.1)
plt.savefig('../graphs/demand-curves-powerlaw-large-price.pdf', format='pdf', dpi=1000); plt.show()
plotTxsCurves(priceRange, demandsPowerlawNumeric.demandWithoutLightning, demandsPowerlawNumeric)
plt.savefig('../graphs/txs-pair-powerlaw-large-price.pdf', format='pdf', dpi=1000); plt.show()
priceRange = np.linspace(0,1,300)
plotDemandCurves(priceRange, demandsPowerlawNumeric.demandWithoutLightning, demandsPowerlawNumeric.demandWithLightning)
plt.title("Demand curves, power-law-distributed transfer-size")
#plt.axes().get_xaxis().set_visible(False)
plt.gca().set_ylim(-0.0001,0.01)
plt.savefig('../graphs/demand-curves-powerlaw-huge-price.pdf', format='pdf', dpi=1000); plt.show()
plotTxsCurves(priceRange, demandsPowerlawNumeric.demandWithoutLightning, demandsPowerlawNumeric)
plt.savefig('../graphs/txs-pair-powerlaw-huge-price.pdf', format='pdf', dpi=1000); plt.show()
nRange = np.linspace(0,2e5,100)
plotPriceCurves(nRange, pricesPowerlawNumeric.priceWithoutLightning, pricesPowerlawNumeric.priceWithLightning)
plt.title("Price curves, power-law-distributed transfer-size")
plt.savefig('../graphs/price-curves-powerlaw-smalln.pdf', format='pdf', dpi=1000)
nRange = np.linspace(0,2e7,100)
plotPriceCurves(nRange, pricesPowerlawNumeric.priceWithoutLightning, pricesPowerlawNumeric.priceWithLightning)
plt.title("Price curves, power-law-distributed transfer-size")
plt.savefig('../graphs/price-curves-powerlaw-mediumn.pdf', format='pdf', dpi=1000)
nRange = np.linspace(0,2e8,100)
plotPriceCurves(nRange, pricesPowerlawNumeric.priceWithoutLightning, pricesPowerlawNumeric.priceWithLightning)
plt.title("Price curves, power-law-distributed transfer-size")
plt.savefig('../graphs/price-curves-powerlaw-largen.pdf', format='pdf', dpi=1000)
nRange = np.linspace(0,2e9,100)
plotPriceCurves(nRange, pricesPowerlawNumeric.priceWithoutLightning, pricesPowerlawNumeric.priceWithLightning)
plt.title("Price curves, power-law-distributed transfer-size")
plt.savefig('../graphs/price-curves-powerlaw-hugen.pdf', format='pdf', dpi=1000)
nRange = np.linspace(0,2e5,100)
plotMarketTxsCurves(nRange, demandsPowerlawNumeric.demandWithoutLightning, demandsPowerlawNumeric, pricesPowerlawNumeric.priceWithoutLightning, pricesPowerlawNumeric.priceWithLightning)
plt.title("Txs, powerlaw transfer-size, symmetric")
plt.savefig('../graphs/txs-market-powerlaw-symmetric-smalln.pdf', format='pdf', dpi=1000)
plt.show()
nRange = np.linspace(0,2e8,100)
plotMarketTxsCurves(nRange, demandsPowerlawNumeric.demandWithoutLightning, demandsPowerlawNumeric, pricesPowerlawNumeric.priceWithoutLightning, pricesPowerlawNumeric.priceWithLightning)
plt.title("Txs, powerlaw transfer-size, symmetric")
plt.savefig('../graphs/txs-market-powerlaw-symmetric-largen.pdf', format='pdf', dpi=1000)
plt.show()
nRange = np.linspace(0,2e9,100)
plotMarketTxsCurves(nRange, demandsPowerlawNumeric.demandWithoutLightning, demandsPowerlawNumeric, pricesPowerlawNumeric.priceWithoutLightning, pricesPowerlawNumeric.priceWithLightning)
plt.title("Txs, powerlaw transfer-size, symmetric")
plt.savefig('../graphs/txs-market-powerlaw-symmetric-hugen.pdf', format='pdf', dpi=1000)
plt.show()
```
| github_jupyter |
# PyTorch Lightning example on MNIST
## References
* [Step-by-step walk-through — PyTorch Lightning 1.5.0dev documentation](https://pytorch-lightning.readthedocs.io/en/latest/starter/introduction_guide.html)
* [Introduction to Pytorch Lightning — PyTorch Lightning 1.5.0dev documentation](https://pytorch-lightning.readthedocs.io/en/latest/notebooks/lightning_examples/mnist-hello-world.html)
* [Mastering-PyTorch/pytorch_lightning.ipynb at master · PacktPublishing/Mastering-PyTorch](https://github.com/PacktPublishing/Mastering-PyTorch/blob/master/Chapter14/pytorch_lightning.ipynb)
---
* [torch.nn.functional](https://pytorch.org/docs/stable/nn.functional.html)
* [torchvision.datasets](https://pytorch.org/vision/stable/datasets.html)
* [torchvision.transforms](https://pytorch.org/vision/stable/transforms.html)
* [PyTorch Lightning Documentation — PyTorch Lightning 1.5.0dev documentation](https://pytorch-lightning.readthedocs.io/en/latest/)
```
import os
import argparse
import torch
import torch.nn.functional as F
import torchvision
import pytorch_lightning as pl
# For type hinting
from typing import Union, Dict, List, Any
from torch import Tensor
```
## References
* [pytorch_lightning.LightningModule](https://pytorch-lightning.readthedocs.io/en/latest/common/lightning_module.html)
- PyTorch のコードを 5つのセクションに整理する
1. Computations (init)
2. Train loop (training_step)
3. Validation loop (validation_step)
4. Test loop (test_step)
5. Optimizers (configure_optimizers)
- 特徴は以下の通り
1. PyTorch Lightning のコードは PyTorch のコードと同じである。
2. PyTorch のコードが抽象化されるわけではなく、整理される。
3. `LightningModule` に無いコードは `Trainer` によって自動化されている。
4. Lightning が処理をするため `.cuda()` や `.to()` といったコールは不要。
5. `DataLoader` において、デフォルトでは `DistributedSampler` が設定される。
6. `LightningModule` は `torch.nn.Module` の一つであり、それに機能を追加したもの。
### `__init__`
* [torch.nn.Conv2d](https://pytorch.org/docs/stable/generated/torch.nn.Conv2d.html#torch.nn.Conv2d)
* [torch.nn.MaxPool2d](https://pytorch.org/docs/1.9.0/generated/torch.nn.MaxPool2d.html#torch.nn.MaxPool2d)
* [torch.nn.Dropout2d](https://pytorch.org/docs/stable/generated/torch.nn.Dropout2d.html#torch.nn.Dropout2d)
* [torch.nn.Linear](https://pytorch.org/docs/stable/generated/torch.nn.Linear.html#torch.nn.Linear)
### `forward`
順伝播型ネットワークの定義および処理の実行
* [forward](https://pytorch-lightning.readthedocs.io/en/latest/common/lightning_module.html?highlight=forward#forward)
- [torch.nn.forward](https://pytorch.org/docs/stable/generated/torch.nn.Module.html#torch.nn.Module.forward)
* [torch.nn.functional.relu](https://pytorch.org/docs/1.9.0/generated/torch.nn.functional.relu.html)
- [torch.nn.ReLU](https://pytorch.org/docs/1.9.0/generated/torch.nn.ReLU.html#torch.nn.ReLU)
- 正規化線形関数を適用
* [torch.flatten](https://pytorch.org/docs/stable/generated/torch.flatten.html)
- 次元を 1 + `start_dim` 次元に変更
* [torch.nn.functional.log_softmax](https://pytorch.org/docs/1.9.0/generated/torch.nn.functional.log_softmax.html#torch.nn.functional.log_softmax)
- [torch.nn.LogSoftmax](https://pytorch.org/docs/1.9.0/generated/torch.nn.LogSoftmax.html#torch.nn.LogSoftmax)
- ソフトマックス関数(活性化関数)を適用
---
### Training loop
#### `training_step`
学習を実行し、損失を返却する。 `train_dataloader` から学習用データを取得し、バッチごとに処理をする。処理には順伝播,勾配の最適化,逆伝播,パラメータの最適化が含まれる。
GPUごとに処理をしたい場合は、これに加えて `training_step_end` をオーバーライドし、各GPUを利用した `training_step` の結果を結合する処理を記述する。
* [training_step](https://pytorch-lightning.readthedocs.io/en/latest/common/lightning_module.html#training-step)
* [torch.nn.functional.cross_entropy](https://pytorch.org/docs/1.9.0/generated/torch.nn.functional.cross_entropy.html?highlight=cross_entropy#torch.nn.functional.cross_entropy)
- [torch.nn.CrossEntropyLoss](https://pytorch.org/docs/1.9.0/generated/torch.nn.CrossEntropyLoss.html#torch.nn.CrossEntropyLoss)
* [LightningModule.log](https://pytorch-lightning.readthedocs.io/en/latest/common/lightning_module.html#log)
#### `training_step_end`
#### `training_epoch_end`
---
### Validation loop
#### `validation_step`
バリデーションを実行する。 `validation_epoch_end` への入力として集約したい値を返却する。ここでは汎化性能を確認するため、損失を返却する。
* [validation_step](https://pytorch-lightning.readthedocs.io/en/latest/common/lightning_module.html#validation-step)
#### `validation_step_end`
* [validation_step_end](https://pytorch-lightning.readthedocs.io/en/latest/common/lightning_module.html#validation-step-end)
#### `validation_epoch_end`
バリデーションのエポック終了時にコールされる。すべての `validation_step` の出力を入力として受け取る。
* [validation_epoch_end](https://pytorch-lightning.readthedocs.io/en/latest/common/lightning_module.html#validation-epoch-end)
* [torch.stack](https://pytorch.org/docs/1.9.0/generated/torch.stack.html?highlight=stack#torch.stack)
---
### Test loop
#### `test_step`
テストを実行する。`test_epoch_end` への入力として集約したい値を返却する。ここでは汎化性能を確認するため、損失を返却する。
テストループは `pytorch_lightning.Trainer.test(model)` が実行された場合のみ、実行される。
* [test_step](https://pytorch-lightning.readthedocs.io/en/latest/common/lightning_module.html#test-step)
#### `test_step_end`
* [test_step_end](https://pytorch-lightning.readthedocs.io/en/latest/common/lightning_module.html#test-step-end)
#### `test_epoch_end`
テストのエポック終了時にコールされる。すべての `test_step` の出力を入力として受け取る。
* [test_epoch_end](https://pytorch-lightning.readthedocs.io/en/latest/common/lightning_module.html#test-epoch-end)
---
### Prediction
#### `predict_step`
`pytorch_lightning.Trainer.predict(model)` が実行された場合のみ、実行される。
* [predict_step](https://pytorch-lightning.readthedocs.io/en/latest/common/lightning_module.html#predict-step)
---
### Optimizer
#### `configure_optimizers`
最適化のために使用するオプティマイザを選択する。
* [configure_optimizers](https://pytorch-lightning.readthedocs.io/en/latest/common/lightning_module.html#configure-optimizers)
* [torch.optim.Adadelta](https://pytorch.org/docs/1.9.0/generated/torch.optim.Adadelta.html?highlight=adadelta#torch.optim.Adadelta)
- [[1212.5701] ADADELTA: An Adaptive Learning Rate Method](https://arxiv.org/abs/1212.5701)
---
### DataLoaders
#### `train_dataloader`
* [train_dataloader](https://pytorch-lightning.readthedocs.io/en/latest/common/lightning_module.html#train-dataloader)
* [torchvision.transforms.ToTensor](https://pytorch.org/vision/stable/transforms.html#torchvision.transforms.ToTensor)
* [torchvision.transforms.Normalize](https://pytorch.org/vision/stable/transforms.html#torchvision.transforms.Normalize)
* [torchvision.transforms.Compose](https://pytorch.org/vision/stable/transforms.html#torchvision.transforms.Compose)
* [torchvision.datasets.MNIST](https://pytorch.org/vision/stable/datasets.html#mnist)
* [torch.utils.data.DataLoader](https://pytorch.org/docs/stable/data.html#torch.utils.data.DataLoader)
#### `val_dataloader`
* [val_dataloader](https://pytorch-lightning.readthedocs.io/en/latest/common/lightning_module.html#val-dataloader)
#### `test_dataloader`
* [test_dataloader](https://pytorch-lightning.readthedocs.io/en/latest/common/lightning_module.html#test-dataloader)
---
### Others
```
class MNISTConvNet(pl.LightningModule):
def __init__(self,
conv1_in_channels: int, conv1_out_channels: int, conv1_kernel_size: int, conv1_stride: int,
conv2_in_channels: int, conv2_out_channels: int, conv2_kernel_size: int, conv2_stride: int,
pool1_kernel_size: int, dropout1_p: float, dropout2_p: float,
fullconn1_in_features: int, fullconn1_out_features: int, fullconn2_in_features: int, fullconn2_out_features: int,
adadelta_lr: float, adadelta_rho: float, adadelta_eps: float, adadelta_weight_decay: float,
dataset_root: str, dataset_download: bool,
dataloader_mean: tuple, dataloader_std: tuple, dataloader_batch_size: int, dataloader_num_workers: int
) -> None:
super(MNISTConvNet, self).__init__()
self.conv1 = torch.nn.Conv2d(in_channels=conv1_in_channels, out_channels=conv1_out_channels, kernel_size=conv1_kernel_size, stride=conv1_stride)
self.conv2 = torch.nn.Conv2d(in_channels=conv2_in_channels, out_channels=conv2_out_channels, kernel_size=conv2_kernel_size, stride=conv2_stride)
self.pool1 = torch.nn.MaxPool2d(kernel_size=pool1_kernel_size)
self.dropout1 = torch.nn.Dropout2d(p=dropout1_p, inplace=False)
self.dropout2 = torch.nn.Dropout2d(p=dropout2_p, inplace=False)
self.fullconn1 = torch.nn.Linear(in_features=fullconn1_in_features, out_features=fullconn1_out_features)
self.fullconn2 = torch.nn.Linear(in_features=fullconn2_in_features, out_features=fullconn2_out_features)
self.adadelta_params = {
'lr': adadelta_lr,
'rho': adadelta_rho,
'eps': adadelta_eps,
'weight_decay': adadelta_weight_decay,
}
self.dataset_params = {
'root': dataset_root,
'download': dataset_download,
}
self.dataloader_params = {
'mean': dataloader_mean,
'std': dataloader_std,
'batch_size': dataloader_batch_size,
'num_workers': dataloader_num_workers,
}
def forward(self, x: Tensor) -> Tensor:
x = self.conv1(x)
x = F.relu(input=x)
x = self.conv2(x)
x = F.relu(input=x)
x = self.pool1(x)
x = self.dropout1(x)
x = torch.flatten(input=x, start_dim=1)
x = self.fullconn1(x)
x = F.relu(input=x)
x = self.dropout2(x)
x = self.fullconn2(x)
return F.log_softmax(input=x, dim=1)
def _common_step(self, batch: Any, log_name: str,
log_on_step: Any = None, log_on_epoch: Any = None, log_prog_bar: bool = False
) -> Union[Tensor, Dict[str, Any], None]:
x, y = batch
y_pred = self(x)
loss = F.cross_entropy(input=y_pred, target=y)
self.log(name=log_name, value=loss, prog_bar=log_prog_bar, on_step=log_on_step, on_epoch=log_on_epoch)
return loss
# Training loop
# def on_train_epoch_start(self) -> None:
# return super().on_train_epoch_start()
# def on_train_batch_start(self, batch: Any, batch_idx: int, dataloader_idx: int) -> None:
# return super().on_train_batch_start(batch, batch_idx, dataloader_idx)
def training_step(self, batch: Any, batch_idx: int) -> Union[Tensor, Dict[str, Any]]:
return self._common_step(batch=batch, log_name="train_loss", log_prog_bar=True, log_on_epoch=True)
# def on_train_batch_end(self, outputs: Any, batch: Any, batch_idx: int, dataloader_idx: int) -> None:
# return super().on_train_batch_end(outputs, batch, batch_idx, dataloader_idx)
# def training_step_end(self, *args, **kwargs) -> Any:
# return super().training_step_end(*args, **kwargs)
# def training_epoch_end(self, outputs: Any) -> None:
# return super().training_epoch_end(outputs)
# def on_train_epoch_end(self) -> None:
# return super().on_train_epoch_end()
# Validation loop
# def on_validation_epoch_start(self) -> None:
# return super().on_validation_epoch_start()
# def on_validation_batch_start(self, batch: Any, batch_idx: int, dataloader_idx: int) -> None:
# return super().on_validation_batch_start(batch, batch_idx, dataloader_idx)
def validation_step(self, batch: Any, batch_idx: int) -> Union[Tensor, Dict[str, Any], None]:
return self._common_step(batch=batch, log_name="val_loss", log_prog_bar=True, log_on_step=True, log_on_epoch=True)
# def on_validation_batch_end(self, outputs: Optional[Any], batch: Any, batch_idx: int, dataloader_idx: int) -> None:
# return super().on_validation_batch_end(outputs, batch, batch_idx, dataloader_idx)
# def validation_step_end(self, *args, **kwargs) -> Optional[Any]:
# return super().validation_step_end(*args, **kwargs)
def validation_epoch_end(self, outputs: List[Union[Tensor, Dict[str, Any]]] ) -> None:
return torch.stack(tensors=outputs).mean() # NOTE: Average loss
# def on_validation_epoch_end(self) -> None:
# return super().on_validation_epoch_end()
# Test loop
# def on_test_epoch_start(self) -> None:
# return super().on_test_epoch_start()
# def on_test_batch_start(self, batch: Any, batch_idx: int, dataloader_idx: int) -> None:
# return super().on_test_batch_start(batch, batch_idx, dataloader_idx)
def test_step(self, batch: Any, batch_idx: int) -> Union[Tensor, Dict[str, Any], None]:
return self._common_step(batch=batch, log_name="test_loss", log_prog_bar=True, log_on_step=True, log_on_epoch=True)
# def on_test_batch_end(self, outputs: Optional[Any], batch: Any, batch_idx: int, dataloader_idx: int) -> None:
# return super().on_test_batch_end(outputs, batch, batch_idx, dataloader_idx)
# def test_step_end(self, *args, **kwargs) -> Optional[Any]:
# return super().test_step_end(*args, **kwargs)
def test_epoch_end(self, outputs: List[Union[Tensor, Dict[str, Any]]] ) -> None:
return torch.stack(tensors=outputs).mean() # NOTE: Average loss
# def on_test_epoch_end(self) -> None:
# return super().on_test_epoch_end()
# Prediction
# def on_predict_epoch_start(self) -> None:
# return super().on_predict_epoch_start()
# def on_predict_batch_start(self, batch: Any, batch_idx: int, dataloader_idx: int) -> None:
# return super().on_predict_batch_start(batch, batch_idx, dataloader_idx)
# def predict_step(self, batch: Any, batch_idx: int, dataloader_idx: Optional[int]) -> Any:
# return super().predict_step(batch, batch_idx, dataloader_idx=dataloader_idx)
# def on_predict_batch_end(self, outputs: Optional[Any], batch: Any, batch_idx: int, dataloader_idx: int) -> None:
# return super().on_predict_batch_end(outputs, batch, batch_idx, dataloader_idx)
# def on_predict_epoch_end(self, results: List[Any]) -> None:
# return super().on_predict_epoch_end(results)
# Optimizer
def configure_optimizers(self) -> Any:
return torch.optim.Adadelta(params=self.parameters(),
lr=self.adadelta_params['lr'],
rho=self.adadelta_params['rho'],
eps=self.adadelta_params['eps'],
weight_decay=self.adadelta_params['weight_decay'])
# def on_before_optimizer_step(self, optimizer: Any, optimizer_idx: int) -> None:
# return super().on_before_optimizer_step(optimizer, optimizer_idx)
# def optimizer_step(self, epoch: int, batch_idx: int, optimizer: Any, optimizer_idx: int, optimizer_closure: Optional[Any], on_tpu: bool, using_native_amp: bool, using_lbfgs: bool) -> None:
# return super().optimizer_step(epoch=epoch, batch_idx=batch_idx, optimizer=optimizer, optimizer_idx=optimizer_idx, optimizer_closure=optimizer_closure, on_tpu=on_tpu, using_native_amp=using_native_amp, using_lbfgs=using_lbfgs)
# Dataloaders
def _get_dataloader(self, train: bool) -> Any:
transform_objects = [
torchvision.transforms.ToTensor(),
torchvision.transforms.Normalize(mean=self.dataloader_params['mean'], std=self.dataloader_params['std'])
]
transform = torchvision.transforms.Compose(transforms=transform_objects)
dataset = torchvision.datasets.MNIST(root=self.dataset_params['root'],
train=train,
download=self.dataset_params['download'],
transform=transform)
dataloader = torch.utils.data.DataLoader(dataset=dataset,
batch_size=self.dataloader_params['batch_size'],
num_workers=self.dataloader_params['num_workers'])
return dataloader
# def on_train_dataloader(self) -> None:
# return super().on_train_dataloader()
def train_dataloader(self) -> Any:
return self._get_dataloader(train=True)
# def on_val_dataloader(self) -> None:
# return super().on_val_dataloader()
def val_dataloader(self) -> Any:
return self._get_dataloader(train=True)
# def on_test_dataloader(self) -> None:
# return super().on_test_dataloader()
def test_dataloader(self) -> Any:
return self._get_dataloader(train=False)
# def on_predict_dataloader(self) -> None:
# return super().on_predict_dataloader()
# def predict_dataloader(self) -> Any:
# return super().predict_dataloader()
# Others
# def setup(self, stage: Optional[str]) -> None:
# return super().setup(stage=stage)
# def teardown(self, stage: Optional[str]) -> None:
# return super().teardown(stage=stage)
# def prepare_data(self) -> None:
# return super().prepare_data()
def get_argparser():
parser = argparse.ArgumentParser(description='PyTorch Lightning MNIST Example')
parser.add_argument('--conv1-in-channels', type=int, default=1)
parser.add_argument('--conv1-out-channels', type=int, default=32)
parser.add_argument('--conv1-kernel-size', type=int, default=3)
parser.add_argument('--conv1-stride', type=int, default=1)
parser.add_argument('--conv2-in-channels', type=int, default=32)
parser.add_argument('--conv2-out-channels', type=int, default=64)
parser.add_argument('--conv2-kernel-size', type=int, default=3)
parser.add_argument('--conv2-stride', type=int, default=1)
parser.add_argument('--pool1-kernel-size', type=int, default=2)
parser.add_argument('--dropout1-p', type=float, default=0.25)
parser.add_argument('--dropout2-p', type=float, default=0.5)
parser.add_argument('--fullconn1-in-features', type=int, default=12*12*64)
parser.add_argument('--fullconn1-out-features', type=int, default=128)
parser.add_argument('--fullconn2-in-features', type=int, default=128)
parser.add_argument('--fullconn2-out-features', type=int, default=10)
parser.add_argument('--adadelta-lr', type=float, default=1.0)
parser.add_argument('--adadelta-rho', type=float, default=0.9)
parser.add_argument('--adadelta-eps', type=float, default=1e-06)
parser.add_argument('--adadelta-weight-decay', type=float, default=0)
parser.add_argument('--dataset-root', type=str, default=os.getcwd())
parser.add_argument('--dataset-download', action='store_true', default=True)
parser.add_argument('--dataloader-mean', type=tuple, default=(0.1302,))
parser.add_argument('--dataloader-std', type=tuple, default=(0.3069,))
parser.add_argument('--dataloader-batch-size', type=int, default=32)
parser.add_argument('--dataloader-num-workers', type=int, default=4)
parser.add_argument('--progress-bar-refresh-rate', type=int, default=20)
parser.add_argument('--max-epochs', type=int, default=1)
return parser
```
## References
* [pytorch_lightning.Trainer](https://pytorch-lightning.readthedocs.io/en/latest/common/trainer.html?highlight=trainer)
- [fit](https://pytorch-lightning.readthedocs.io/en/latest/common/trainer.html#fit)
```
def main(args=None) -> None:
if not args:
args = get_argparser().parse_args()
model = MNISTConvNet(conv1_in_channels=args.conv1_in_channels, conv1_out_channels=args.conv1_out_channels, conv1_kernel_size=args.conv1_kernel_size, conv1_stride=args.conv1_stride,
conv2_in_channels=args.conv2_in_channels, conv2_out_channels=args.conv2_out_channels, conv2_kernel_size=args.conv2_kernel_size, conv2_stride=args.conv2_stride,
pool1_kernel_size=args.pool1_kernel_size, dropout1_p=args.dropout1_p, dropout2_p=args.dropout2_p,
fullconn1_in_features=args.fullconn1_in_features, fullconn1_out_features=args.fullconn1_out_features, fullconn2_in_features=args.fullconn2_in_features, fullconn2_out_features=args.fullconn2_out_features,
adadelta_lr=args.adadelta_lr, adadelta_rho=args.adadelta_rho, adadelta_eps=args.adadelta_eps, adadelta_weight_decay=args.adadelta_weight_decay,
dataset_root=args.dataset_root, dataset_download=args.dataset_download,
dataloader_mean=args.dataloader_mean, dataloader_std=args.dataloader_std, dataloader_batch_size=args.dataloader_batch_size, dataloader_num_workers=args.dataloader_num_workers)
trainer = pl.Trainer(progress_bar_refresh_rate=args.progress_bar_refresh_rate, max_epochs=args.max_epochs)
trainer.fit(model)
trainer.test(model)
# trainer.predict(model)
if __name__ == "__main__":
argparser = get_argparser()
args = argparser.parse_args([
"--max-epochs", str(1),
"--adadelta-lr", str(0.5),
])
main(args)
```
| github_jupyter |
# widgets.image_cleaner
fastai offers several widgets to support the workflow of a deep learning practitioner. The purpose of the widgets are to help you organize, clean, and prepare your data for your model. Widgets are separated by data type.
```
from fastai.vision import *
from fastai.widgets import DatasetFormatter, ImageCleaner, ImageDownloader, download_google_images
from fastai.gen_doc.nbdoc import *
%reload_ext autoreload
%autoreload 2
path = untar_data(URLs.MNIST_SAMPLE)
data = ImageDataBunch.from_folder(path)
learn = create_cnn(data, models.resnet18, metrics=error_rate)
learn.fit_one_cycle(2)
learn.save('stage-1')
```
We create a databunch with all the data in the training set and no validation set (DatasetFormatter uses only the training set)
```
db = (ImageItemList.from_folder(path)
.no_split()
.label_from_folder()
.databunch())
learn = create_cnn(db, models.resnet18, metrics=[accuracy])
learn.load('stage-1');
show_doc(DatasetFormatter)
```
The [`DatasetFormatter`](/widgets.image_cleaner.html#DatasetFormatter) class prepares your image dataset for widgets by returning a formatted [`DatasetTfm`](/vision.data.html#DatasetTfm) based on the [`DatasetType`](/basic_data.html#DatasetType) specified. Use `from_toplosses` to grab the most problematic images directly from your learner. Optionally, you can restrict the formatted dataset returned to `n_imgs`.
```
show_doc(DatasetFormatter.from_similars)
from fastai.gen_doc.nbdoc import *
from fastai.widgets.image_cleaner import *
show_doc(DatasetFormatter.from_toplosses)
show_doc(ImageCleaner)
```
[`ImageCleaner`](/widgets.image_cleaner.html#ImageCleaner) is for cleaning up images that don't belong in your dataset. It renders images in a row and gives you the opportunity to delete the file from your file system. To use [`ImageCleaner`](/widgets.image_cleaner.html#ImageCleaner) we must first use `DatasetFormatter().from_toplosses` to get the suggested indices for misclassified images.
```
ds, idxs = DatasetFormatter().from_toplosses(learn)
ImageCleaner(ds, idxs, path)
```
[`ImageCleaner`](/widgets.image_cleaner.html#ImageCleaner) does not change anything on disk (neither labels or existence of images). Instead, it creates a 'cleaned.csv' file in your data path from which you need to load your new databunch for the files to changes to be applied.
```
df = pd.read_csv(path/'cleaned.csv', header='infer')
# We create a databunch from our csv. We include the data in the training set and we don't use a validation set (DatasetFormatter uses only the training set)
np.random.seed(42)
db = (ImageItemList.from_df(df, path)
.no_split()
.label_from_df()
.databunch(bs=64))
learn = create_cnn(db, models.resnet18, metrics=error_rate)
learn = learn.load('stage-1')
```
You can then use [`ImageCleaner`](/widgets.image_cleaner.html#ImageCleaner) again to find duplicates in the dataset. To do this, you can specify `duplicates=True` while calling ImageCleaner after getting the indices and dataset from `.from_similars`. Note that if you are using a layer's output which has dimensions <code>(n_batches, n_features, 1, 1)</code> then you don't need any pooling (this is the case with the last layer). The suggested use of `.from_similars()` with resnets is using the last layer and no pooling, like in the following cell.
```
ds, idxs = DatasetFormatter().from_similars(learn, layer_ls=[0,7,1], pool=None)
ImageCleaner(ds, idxs, path, duplicates=True)
show_doc(ImageDownloader)
```
[`ImageDownloader`](/widgets.image_downloader.html#ImageDownloader) widget gives you a way to quickly bootstrap your image dataset without leaving the notebook. It searches and downloads images that match the search criteria and resolution / quality requirements and stores them on your filesystem within the provided `path`.
Images for each search query (or label) are stored in a separate folder within `path`. For example, if you pupulate `tiger` with a `path` setup to `./data`, you'll get a folder `./data/tiger/` with the tiger images in it.
[`ImageDownloader`](/widgets.image_downloader.html#ImageDownloader) will automatically clean up and verify the downloaded images with [`verify_images()`](/vision.data.html#verify_images) after downloading them.
```
path = Config.data_path()/'image_downloader'
os.makedirs(path, exist_ok=True)
ImageDownloader(path)
```
#### Downloading images in python scripts outside Jupyter notebooks
```
path = Config.data_path()/'image_downloader'
files = download_google_images(path, 'aussie shepherd', size='>1024*768', n_images=30)
len(files)
show_doc(download_google_images)
```
After populating images with [`ImageDownloader`](/widgets.image_downloader.html#ImageDownloader), you can get a an [`ImageDataBunch`](/vision.data.html#ImageDataBunch) by calling `ImageDataBunch.from_folder(path, size=size)`, or using the data block API.
```
# Setup path and labels to search for
path = Config.data_path()/'image_downloader'
labels = ['boston terrier', 'french bulldog']
# Download images
for label in labels:
download_google_images(path, label, size='>400*300', n_images=50)
# Build a databunch and train!
src = (ImageItemList.from_folder(path)
.random_split_by_pct()
.label_from_folder()
.transform(get_transforms(), size=224))
db = src.databunch(bs=16, num_workers=0)
learn = create_cnn(db, models.resnet34, metrics=[accuracy])
learn.fit_one_cycle(3)
```
#### Downloading more than a hundred images
To fetch more than a hundred images, [`ImageDownloader`](/widgets.image_downloader.html#ImageDownloader) uses `selenium` and `chromedriver` to scroll through the Google Images search results page and scrape image URLs. They're not required as dependencies by default. If you don't have them installed on your system, the widget will show you an error message.
To install `selenium`, just `pip install selenium` in your fastai environment.
**On a mac**, you can install `chromedriver` with `brew cask install chromedriver`.
**On Ubuntu**
Take a look at the latest Chromedriver version available, then something like:
```
wget https://chromedriver.storage.googleapis.com/2.45/chromedriver_linux64.zip
unzip chromedriver_linux64.zip
```
Note that downloading under 100 images doesn't require any dependencies other than fastai itself, however downloading more than a hundred images [uses `selenium` and `chromedriver`](/widgets.image_cleaner.html#Downloading-more-than-a-hundred-images).
`size` can be one of:
```
'>400*300'
'>640*480'
'>800*600'
'>1024*768'
'>2MP'
'>4MP'
'>6MP'
'>8MP'
'>10MP'
'>12MP'
'>15MP'
'>20MP'
'>40MP'
'>70MP'
```
## Methods
## Undocumented Methods - Methods moved below this line will intentionally be hidden
```
show_doc(ImageCleaner.make_dropdown_widget)
show_doc(ImageCleaner.next_batch)
show_doc(DatasetFormatter.sort_idxs)
show_doc(ImageCleaner.make_vertical_box)
show_doc(ImageCleaner.relabel)
show_doc(DatasetFormatter.largest_indices)
show_doc(ImageCleaner.delete_image)
show_doc(ImageCleaner.empty)
show_doc(ImageCleaner.empty_batch)
show_doc(DatasetFormatter.comb_similarity)
show_doc(ImageCleaner.get_widgets)
show_doc(ImageCleaner.write_csv)
show_doc(ImageCleaner.create_image_list)
show_doc(ImageCleaner.render)
show_doc(DatasetFormatter.get_similars_idxs)
show_doc(ImageCleaner.on_delete)
show_doc(ImageCleaner.make_button_widget)
show_doc(ImageCleaner.make_img_widget)
show_doc(DatasetFormatter.get_actns)
show_doc(ImageCleaner.batch_contains_deleted)
show_doc(ImageCleaner.make_horizontal_box)
show_doc(DatasetFormatter.get_toplosses_idxs)
show_doc(DatasetFormatter.padded_ds)
```
## New Methods - Please document or move to the undocumented section
| github_jupyter |
## Replicate weights
Replicate weights are usually created for the purpose of variance (uncertainty) estimation. One common use case for replication-based methods is the estimation of non-linear parameters fow which Taylor-based approximation may not be accurate enough. Another use case is when the number of primary sampling units selected per stratum is small (low degree of freedom). Replicate weights are usually created for the purpose of variance (uncertainty)estimation. One common use case for replication-based methods is the estimation of non-linear parameters fow which Taylor-based approximation may not be accurate enough. Another use case is when the number of primary sampling units selected per stratum is small (low degree of freedom).
In this tutorial, we will explore creating replicate weights using the class *ReplicateWeight*. Three replication methods have been implemented: balanced repeated replication (BRR) including the Fay-BRR, bootstrap and jackknife. The replicate method of interest is specified when initializing the class by using the parameter *method*. The parameter *method* takes the values "bootstrap", "brr", or "jackknife". In this tutorial, we show how the API works for producing replicate weights.
```
import pandas as pd
from samplics.datasets import load_psu_sample, load_ssu_sample
from samplics.weighting import ReplicateWeight
```
We import the sample data...
```
# Load PSU sample data
psu_sample_dict = load_psu_sample()
psu_sample = psu_sample_dict["data"]
# Load PSU sample data
ssu_sample_dict = load_ssu_sample()
ssu_sample = ssu_sample_dict["data"]
full_sample = pd.merge(
psu_sample[["cluster", "region", "psu_prob"]],
ssu_sample[["cluster", "household", "ssu_prob"]],
on="cluster")
full_sample["inclusion_prob"] = full_sample["psu_prob"] * full_sample["ssu_prob"]
full_sample["design_weight"] = 1 / full_sample["inclusion_prob"]
full_sample.head(15)
```
### Balanced Repeated Replication (BRR) <a name="section1"></a>
The basic idea of BRR is to slip the sample in independent random groups. The groups are then threated as independent replicates of the the sample design. A special case is when the sample is split into two half samples in each stratum. This design is suitable to many survey designs where only two psus are selected by stratum. In practice, one of the psu is asigned to the first random group and the other psu is assign to the second group. The sample weights are double for one group (say the first one) and the sample weights in the other group are set to zero. To ensure that the replicates are independent, we use hadamard matrices to assign the random groups.
```
import scipy
scipy.linalg.hadamard(8)
```
In our example, we have 10 psus. If we do not have explicit stratification then *replicate()* will group the clusters into 5 strata (2 per stratum). In this case, the smallest number of replicates possible using the hadamard matrix is 8.
The result below shows that *replicate()* created 5 strata by grouping clusters 7 and 10 in the first stratum, clusters 16 and 24 in the second stratum, and so on. We can achieve the same result by providing setting *stratification=True* and providing the stratum variable to *replicate()*.
```
brr = ReplicateWeight(method="brr", stratification=False)
brr_wgt = brr.replicate(full_sample["design_weight"], full_sample["cluster"])
brr_wgt.drop_duplicates().head(10)
```
An extension of BRR is the Fay's method. In the Fay's approach, instead of multiplying one half-sample by zero, we multiple the sampel weights by a factor $\alpha$ and the other halh-sample by $2-\alpha$. We refer to $\alpha$ as the fay coefficient. Note that when $\alpha=0$ then teh Fay's method reduces to BRR.
```
fay = ReplicateWeight(method="brr", stratification=False, fay_coef=0.3)
fay_wgt = fay.replicate(
full_sample["design_weight"],
full_sample["cluster"],
rep_prefix="fay_weight_",
psu_varname="cluster",
str_varname="stratum"
)
fay_wgt.drop_duplicates().head(10)
```
### Bootstrap <a name="section2"></a>
For the bootstrap replicates, we need to provide the number of replicates. When the number of replicates is not provided, *ReplicateWeight* will default to 500. The bootstrap consists of selecting the same number of psus as in the sample but with replacement. The selection is independently repeated for each replicate.
```
bootstrap = ReplicateWeight(method="bootstrap", stratification=False, number_reps=50)
boot_wgt = bootstrap.replicate(full_sample["design_weight"], full_sample["cluster"])
boot_wgt.drop_duplicates().head(10)
```
### Jackknife <a name="section3"></a>
Below, we illustrate the API for creating replicate weights using the jackknife method.
```
jackknife = ReplicateWeight(method="jackknife", stratification=False)
jkn_wgt = jackknife.replicate(full_sample["design_weight"], full_sample["cluster"])
jkn_wgt.drop_duplicates().head(10)
```
With stratification...
```
jackknife = ReplicateWeight(method="jackknife", stratification=True)
jkn_wgt = jackknife.replicate(full_sample["design_weight"], full_sample["cluster"], full_sample["region"])
jkn_wgt.drop_duplicates().head(10)
```
**Important.** For any of the three methods, we can request the replicate coefficient instead of the replicate weights by *rep_coefs=True*.
```
#jackknife = ReplicateWeight(method="jackknife", stratification=True)
jkn_wgt = jackknife.replicate(
full_sample["design_weight"], full_sample["cluster"], full_sample["region"], rep_coefs=True
)
jkn_wgt.drop_duplicates().sort_values(by="_stratum").head(15)
#fay = ReplicateWeight(method="brr", stratification=False, fay_coef=0.3)
fay_wgt = fay.replicate(
full_sample["design_weight"],
full_sample["cluster"],
rep_prefix="fay_weight_",
psu_varname="cluster",
str_varname="stratum",
rep_coefs=True
)
fay_wgt.drop_duplicates().head(10)
```
| github_jupyter |
# Transfer Learning
In this notebook, you'll learn how to use pre-trained networks to solve challenging problems in computer vision. Specifically, you'll use a network trained on [ImageNet](http://www.image-net.org/). ImageNet is a massive dataset with over 1 million labeled images in 1,000 categories.
These pre-trained models work astonishingly well as feature detectors for images they weren't trained on. Using a pre-trained network on images not in the training set is called **Transfer Learning**. Here we'll use transfer learning to train a network that can classify our cat and dog photos with near perfect accuracy.
With [TensorFlow Hub](https://www.tensorflow.org/hub) you can download these pre-trained networks and use them in your applications.
## Import Resources
```
import warnings
warnings.filterwarnings('ignore')
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
import time
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
import tensorflow_hub as hub
import tensorflow_datasets as tfds
tfds.disable_progress_bar()
import logging
logger = tf.get_logger()
logger.setLevel(logging.ERROR)
print('Using:')
print('\t\u2022 TensorFlow version:', tf.__version__)
print('\t\u2022 tf.keras version:', tf.keras.__version__)
print('\t\u2022 Running on GPU' if tf.test.is_gpu_available() else '\t\u2022 GPU device not found. Running on CPU')
```
## Load the Dataset
```
train_split = 60
test_val_split = 20
splits = tfds.Split.ALL.subsplit([60,20, 20])
(training_set, validation_set, test_set), dataset_info = tfds.load('cats_vs_dogs', split=splits, as_supervised=True, with_info=True)
```
## Explore the Dataset
```
dataset_info
num_classes = dataset_info.features['label'].num_classes
total_num_examples = dataset_info.splits['train'].num_examples
print('The Dataset has a total of:')
print('\u2022 {:,} classes'.format(num_classes))
print('\u2022 {:,} images'.format(total_num_examples))
```
As a technical note, if the total number of examples in your dataset is not a multiple of 100 (*i.e.* if `total_num_examples % 100 != 0`), then TensorFlow may not evenly distribute the data among subsplits. As we can see, our dataset has `23,262` examples, which is not a multiple of 100. Therefore, in this particular case, we should expect that our data would not be evenly distributed among the subsplits that we created. This means that even though we set our `split` to allocate 60\% of the data to the training set, 20\% of the data to the validation set, and 20\% of the data to the test set, the actual number of images in each set may vary from these percentages. It is important to note, that these small differences will not affect our training process. We didn't have this issue before when we worked the MNIST and Fashion-MNIST datasets because both of these datasets had 70,000 examples. Since 70,000 is a multiple of 100, then the data was evenly distributed in both of those cases.
```
class_names = ['cat', 'dog']
for image, label in training_set.take(1):
image = image.numpy()
label = label.numpy()
plt.imshow(image)
plt.show()
print('The label of this image is:', label)
print('The class name of this image is:', class_names[label])
```
## Create Pipeline
The pre-trained model we are going to use requires that the input images have color values in the range `[0,1]` and a size of `(224, 224)`. We will therefore have to normalize the pixel values of our images and resize them to the appropriate size. We can normalize our pixel values in the usual way by dividing the original pixel values by `255` and to resize our images we can use the `tf.image.resize()` function.
```
batch_size = 32
image_size = 224
num_training_examples = (total_num_examples * train_split) // 100
def format_image(image, label):
image = tf.cast(image, tf.float32)
image = tf.image.resize(image, (image_size, image_size))
image /= 255
return image, label
training_batches = training_set.shuffle(num_training_examples//4).map(format_image).batch(batch_size).prefetch(1)
validation_batches = validation_set.map(format_image).batch(batch_size).prefetch(1)
testing_batches = test_set.map(format_image).batch(batch_size).prefetch(1)
```
## Transfer Learning with TensorFlow Hub
[TensorFlow Hub](https://www.tensorflow.org/hub) is an online repository of pre-trained models. In addition to complete pre-trained models, TensorFlow Hub also contains models without the last classification layer. These models can be used to perform transfer learning by adding a classification layer that suits the number of classes in your particular dataset. You can take a look at all the models available for TensorFlow 2.0 in [TensorFlow Hub](https://tfhub.dev/s?q=tf2-preview).
In this notebook, we will use a network trained on the ImageNet dataset called MobileNet. MobileNet is a state-of-the-art convolutional neural network developed by Google. Convolutional neural networks are out of the scope of this course, but if you want to learn more about them, you can take a look at this [video](https://www.youtube.com/watch?v=2-Ol7ZB0MmU).
In the cell below we download the pre-trained MobileNet model without the final classification layer from TensorFlow Hub using the `hub.KerasLayer(URL)` function. This function downloads the desired model form the given TensorFlow Hub `URL` and wraps it in a Keras layer so that we can integrate it in a `tf.keras` Sequential model later. Since this will be the first layer of our Sequential model, we need to specify the `input_shape` parameter. The shape of our input tensor must match the size of the images MobileNet was trained on, namely `(224,224,3)`.
Our pre-trained model will be responsible for extracting the features of our images, we will therefore call this part of our model the `feature_extractor`.
```
URL = "https://tfhub.dev/google/tf2-preview/mobilenet_v2/feature_vector/4"
feature_extractor = hub.KerasLayer(URL, input_shape=(image_size, image_size,3))
```
It is important that we freeze the weights and biases in our pre-trained model so that we don't modify them during training. We can do this by setting the parameters of our model to non-trainable, as shown in the code below.
```
feature_extractor.trainable = False
```
## Build the Model
We will now create a `tf.keras` Sequential model with our `feature_extractor` and a new classification layer. Since our dataset only has 2 classes (cat and dog) we create an output layer with only 2 units.
```
model = tf.keras.Sequential([
feature_extractor,
tf.keras.layers.Dense(2, activation = 'softmax')
])
model.summary()
```
## Train the Model Using a GPU
With our model built, we now need to train the new classification layer, but this time we're using a **really deep** neural network. If you try to train this on a CPU like normal, it will take a long, long time. Instead, we're going to use a GPU to do the calculations. On a GPU, linear algebra computations are done in parallel, leading to 100x increased training speeds. TensorFlow will transparently run on a single GPU without requiring that we make changes to our code. With TensorFlow, it's also possible to train on multiple GPUs, further decreasing training time, but this requires that we make changes to our code to incorporate [distributed training](https://www.tensorflow.org/guide/distributed_training).
We can use the `tf.test.is_gpu_available()` function to confirm that TensorFlow is using the GPU.
```
print('Is there a GPU Available:', tf.test.is_gpu_available())
```
TensorFlow uses different string identifiers for CPUs and GPUs. For example, TensorFlow will use the identifier:
```python
'/CPU:0'
```
for the CPU of your machine; and it will use the identifier:
```python
'/GPU:0'
```
for the first GPU of your machine that is visible to TensorFlow. If your system has both devices, `/CPU:0` and `/GPU:0`, by default the GPU devices will be given priority when preforming TensorFlow operations (given that the TensorFlow operations have both CPU and GPU implementations). For example, the TensorFlow `tf.matmul` operation has both CPU and GPU kernels, therefore, the `/GPU:0` device will be selected to run `tf.matmul` unless you explicitly request running it on another device.
### Manual Device Placement
If you would like a particular TensorFlow operation to run on the device of your choice, instead of what's automatically selected for you by default, you can use:
```python
# Place tensors on the CPU
with tf.device('/CPU:0'):
perform operations
```
to have operations run on the CPU; and you can use:
```python
# Place tensors on the GPU
with tf.device('/GPU:0'):
perform operations
```
to have operations run on the GPU.
#### Example
Let's assume we have a system that has both devices, `/CPU:0` and `/GPU:0`. What will happen if we run the code below?
```python
# Place tensors on the CPU
with tf.device('/CPU:0'):
a = tf.random.normal(...)
b = tf.random.normal(...)
c = tf.matmul(a, b)
```
The above code will create both `a` and `b` using the CPU because we manually assigned those statements to the
`/CPU:0` device using the `with tf.device('/CPU:0')` code block. However, since the statement `c = tf.matmul(a, b)` is NOT inside the `with tf.device('/CPU:0')` code block, then TensorFlow will run the `tf.matmul` operation on the `/GPU:0` device. TensorFlow will automatically copy tensors between devices if required.
In the code below, we will multiply matrices of increasing size using both the CPU and GPU so you can see the difference in execution time. You will see, that as the size of the matrices increase, the execution time on the CPU increases rapidly, but on the GPU it stays constant.
```
def plot_times(max_size = 650):
device_times = {'/GPU:0':[], '/CPU:0':[]}
matrix_sizes = range(450, max_size, 50)
len_matrix = len(matrix_sizes)
for i, size in enumerate(matrix_sizes):
for device_name in device_times.keys():
with tf.device(device_name):
m1 = tf.random.uniform(shape=(size,size), dtype=tf.float16)
m2 = tf.random.uniform(shape=(size,size), dtype=tf.float16)
start_time = time.time()
dot_operation = tf.matmul(m2, m1)
time_taken = time.time() - start_time
if i > 0:
device_times[device_name].append(time_taken)
percent_complete = (i + 1) / len_matrix
print('\rPerforming Calculations. Please Wait... {:.0%} Complete'.format(percent_complete), end = '')
matrix_sizes = matrix_sizes[1:]
plt.figure(figsize=(10,7))
plt.plot(matrix_sizes, device_times['/CPU:0'], 'o-', color='magenta', linewidth = 2, label = 'CPU')
plt.plot(matrix_sizes, device_times['/GPU:0'], 'o-', color='cyan', linewidth = 2, label='GPU')
ax = plt.gca()
ax.set_facecolor('black')
plt.grid()
plt.ylabel('Time (s)', color='white', fontsize = 20)
plt.xlabel('Matrix size', color='white', fontsize = 20)
plt.legend(prop={'size': 15})
plt.show()
plot_times(850)
```
From here, I'll let you finish training the model. The process is the same as before except that now your model will automatically run on the GPU. You should get better than 95% accuracy easily.
>**Exercise:** Train the `model` we created above to classify the cat and dog images in our dataset. Because we are using a pre-trained model, you will only need to train the model for a few epochs to get a high accuracy.
```
## Solution
```
# Check Predictions
```
for image_batch, label_batch in testing_batches.take(1):
ps = model.predict(image_batch)
images = image_batch.numpy().squeeze()
labels = label_batch.numpy()
plt.figure(figsize=(10,15))
for n in range(30):
plt.subplot(6,5,n+1)
plt.imshow(images[n], cmap = plt.cm.binary)
color = 'green' if np.argmax(ps[n]) == labels[n] else 'red'
plt.title(class_names[np.argmax(ps[n])], color=color)
plt.axis('off')
```
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
import numpy as np
import torch, torch.nn as nn
import torch.nn.functional as F
from modules import MultiHeadAttention, PositionwiseFeedForward
import mnist
x_train = mnist.make_clouds(mnist.x_train,500)
y_train = mnist.y_train
x_val = mnist.make_clouds(mnist.x_val,500)
y_val = mnist.y_val
import torch, torch.nn as nn
import torch.nn.functional as F
from torch.autograd import Variable
class GlobalAveragePooling(nn.Module):
def __init__(self, dim=-1):
super(self.__class__, self).__init__()
self.dim = dim
def forward(self, x):
return x.mean(dim=self.dim)
class Discriminator(nn.Module):
def __init__(self, in_dim,
hidden_dim=100,
ffn_dim =200,
n_head=8,
normalize_loc=True,
normalize_scale=True):
super(Discriminator, self).__init__()
self.normalize_loc = normalize_loc
self.normalize_scale = normalize_scale
self.fc1 = nn.Linear(in_dim, hidden_dim)
nn.init.xavier_normal_(self.fc1.weight)
nn.init.constant_(self.fc1.bias, 0.0)
self.mha_1 = MultiHeadAttention(n_head=n_head,d_model = hidden_dim)
self.ffn_1 = PositionwiseFeedForward(hidden_dim, ffn_dim, use_residual=False)
self.mha_2 = MultiHeadAttention(n_head=n_head,d_model = hidden_dim)
self.ffn_2 = PositionwiseFeedForward(hidden_dim, ffn_dim, use_residual=False)
self.gl_1 = GlobalAveragePooling(dim = 1)
self.fc2 = nn.Linear(hidden_dim, 2)
nn.init.xavier_normal_(self.fc2.weight)
nn.init.constant_(self.fc2.bias, 0.0)
def forward(self, x):
if self.normalize_loc:
x = x - x.mean(dim=1, keepdim=True)
if self.normalize_scale:
x = x / x.std(dim=1, keepdim=True)
h1 = F.relu(self.fc1(x))
h2 = self.mha_1(h1)
h3 = self.ffn_1(h2)
h4 = self.mha_2(h3)
h5 = self.ffn_2(h4)
score = self.fc2(self.gl_1(h5))
return score
class Generator(nn.Module):
def __init__(self, in_dim, hidden_dim=100,ffn_dim =200,n_head=8):
super(Generator, self).__init__()
self.fc1 = nn.Linear(in_dim, hidden_dim)
nn.init.xavier_normal_(self.fc1.weight)
nn.init.constant_(self.fc1.bias, 0.0)
self.mha_1 = MultiHeadAttention(n_head=n_head,d_model = hidden_dim)
self.ffn_1 = PositionwiseFeedForward(hidden_dim, ffn_dim, use_residual=False)
self.mha_2 = MultiHeadAttention(n_head=n_head,d_model = hidden_dim)
self.ffn_2 = PositionwiseFeedForward(hidden_dim, ffn_dim, use_residual=False)
self.mha_3 = MultiHeadAttention(n_head=n_head,d_model = hidden_dim)
self.ffn_3 = PositionwiseFeedForward(hidden_dim, ffn_dim, use_residual=False)
self.fc2 = nn.Linear(hidden_dim, in_dim)
nn.init.xavier_normal_(self.fc2.weight)
nn.init.constant_(self.fc2.bias, 0.0)
def forward(self, x):
h = F.relu(self.fc1(x))
h = self.mha_1(h)
h = self.ffn_1(h)
h = self.mha_2(h)
h = self.ffn_2(h)
h = self.mha_3(h)
h = self.ffn_3(h)
score = self.fc2(h)
return score
in_dim = 2
hidden_dim = 100
ffn_dim = 200
n_head = 8
disc = Discriminator(in_dim, hidden_dim).cuda(0)
in_dim = 2
hidden_dim = 64
ffn_dim = 128
n_head = 8
gen = Generator(in_dim, hidden_dim).cuda(0)
target = Variable(torch.FloatTensor(x_train[:1])).cuda(0)
noise = torch.rand(target.shape).cuda(0)
# gen_opt = torch.optim.Adam(gen.parameters(), lr=1e-4)
gen_opt = torch.optim.SGD(gen.parameters(), lr=1e-6)
disc_opt = torch.optim.SGD(disc.parameters(), lr=0.001)
from IPython.display import clear_output
from tqdm import trange
import matplotlib.pyplot as plt
%matplotlib inline
IS_FAKE, IS_REAL = 0, 1
noise = torch.rand(target.shape).cuda(0)
for epoch_i in trange(1000):
for i in range(5):
noise = torch.rand(target.shape).cuda(0)
gen_data = gen(noise)
loss_disc = - F.log_softmax(disc(target), 1)[:, IS_REAL].mean() \
- F.log_softmax(disc(gen_data), 1)[:, IS_FAKE].mean()
disc_opt.zero_grad()
loss_disc.backward()
disc_opt.step()
for i in range(1):
noise = torch.rand(target.shape).cuda(0)
gen_data = gen(noise)
loss_gen = - F.log_softmax(disc(gen_data), 1)[:, IS_REAL].mean()
gen_opt.zero_grad()
loss_gen.backward()
gen_opt.step()
if epoch_i % 2 != 0: continue
noise = torch.rand(target.shape).cuda(0)
gen_data = gen(noise)
clear_output(True)
plt.subplot(1,2,1)
_x, _y = target.data.cpu().numpy()[0].T
plt.scatter(_x, -_y)
# plt.ylim(-1, 0)
# plt.xlim(0, 1)
plt.subplot(1,2,2)
_x, _y = gen_data[:1].data.cpu().numpy()[0].T
plt.scatter(_x, -_y)
# plt.ylim(-1, 0)
# plt.xlim(0, 1)
plt.show()
print(F.softmax(disc(gen_data), 1))
```
| github_jupyter |
```
# this is using the positive/negative frequency data to find a correlation with the price change
import pandas as pd
# data date: 2018-1-09 ~ 2018-2-28
freq_data = pd.read_json("Nike_8000_newsSentiment.json")
def weekNum(t):
if t.month == 12 and t.week == 1:
return t.week + (t.year + 1) * 100
elif t.month == 1 and t.week == 53:
return 1 + t.year*100
else:
return t.week + t.year * 100
freq_data['WeekNum'] = pd.to_datetime(freq_data['Date'])
freq_data['WeekNum'] = freq_data['WeekNum'].apply(weekNum)
weekly_freq_data = freq_data.groupby(freq_data['WeekNum']).sum()
weekly_freq_data['freqAvg'] = freq_data.groupby(freq_data['WeekNum']).mean()
weekly_price_data = pd.read_json("NIKE7yrsInWeeks.json")
weekly_price_data['WeekNum'] = weekly_price_data.index
dataNeeded = (weekly_price_data.join(weekly_freq_data))
dataNeeded.dropna(inplace=True)
dataNeeded['frequencyChange'] = dataNeeded['freqeuency'].diff()
#df = freq_data['FreqNum'].loc[::-1]
#df.index = range(len(dataNeeded))
#string(dataNeeded['Date'].iloc[0])
#price_data.merge(freq_data,on='Date')
import matplotlib.pyplot as plt
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
# we can see a little cluster there, which is what we wanted, im not sure
# if it's convincing enough
dataNeeded.corr()
from scipy.stats import pearsonr
# trying our other correlations:
###################################################
stat, pval = pearsonr(dataNeeded['WeeklyChangeAvg'],dataNeeded['freqeuency'])
print("WeeklyChangeAvg vs. frequency correlated (same week):", stat)
print('p val', pval)
###################################################
stat, pval = pearsonr(dataNeeded['WeeklyChangeTotal'],dataNeeded['freqeuency'])
print("WeeklyChangeTotal vs. frequency correlated (same week):", stat)
print('p val', pval)
stat, pval = pearsonr(dataNeeded['WeeklyAvg'],dataNeeded['freqeuency'])
print("WeeklyAvg vs. frequency correlated (same week):", stat)
print('p val', pval)
stat, pval = pearsonr(dataNeeded['WeeklyChangeAvg'],dataNeeded['freqAvg'])
print("WeeklyChangeAvg vs. freqAvg correlated (same week):", stat)
print('p val', pval)
stat, pval = pearsonr(dataNeeded['WeeklyChangeTotal'],dataNeeded['freqAvg'])
print("WeeklyChangeTotal vs. freqAvg correlated (same week):", stat)
print('p val', pval)
stat, pval = pearsonr(dataNeeded['WeeklyAvg'],dataNeeded['freqAvg'])
print("WeeklyAvg vs. frequency freqAvg (same week):", stat)
print('p val', pval)
###################################################
stat, pval = pearsonr(dataNeeded['WeeklyChangeAvg'][1:],dataNeeded['frequencyChange'][1:])
print("WeeklyChangeAvg vs. frequency change correlated (same week):", stat)
print('p val', pval)
###################################################
stat, pval = pearsonr(dataNeeded['WeeklyChangeTotal'][1:],dataNeeded['frequencyChange'][1:])
print("WeeklyChangeAvg vs. frequency change correlated (same week):", stat)
print('p val', pval)
stat, pval = pearsonr(dataNeeded['WeeklyChangeTotal'][1:],dataNeeded['frequencyChange'][1:])
print("WeeklyChangeAvg vs. frequency change correlated (same week):", stat)
print('p val', pval)
stat, pval = pearsonr(dataNeeded['WeeklyChangeAvg'][1:],dataNeeded['freqeuency'][:-1])
print("WeeklyChangeAvg vs. last week's frequency correlated (same week):", stat)
print('p val', pval)
stat, pval = pearsonr(dataNeeded['WeeklyChangeTotal'][1:],dataNeeded['freqeuency'][:-1])
print("WeeklyChangeTotal vs. last week's frequency correlated (same week):", stat)
print('p val', pval)
stat, pval = pearsonr(dataNeeded['WeeklyAvg'][1:],dataNeeded['freqeuency'][:-1])
print("WeeklyAvg vs. last week's frequency correlated (same week):", stat)
print('p val', pval)
stat, pval = pearsonr(dataNeeded['WeeklyChangeAvg'][1:],dataNeeded['freqAvg'][:-1])
print("WeeklyChangeAvg vs. last week's freqAvg correlated (same week):", stat)
print('p val', pval)
stat, pval = pearsonr(dataNeeded['WeeklyChangeTotal'][1:],dataNeeded['freqAvg'][:-1])
print("WeeklyChangeTotal vs. last week's freqAvg correlated (same week):", stat)
print('p val', pval)
stat, pval = pearsonr(dataNeeded['WeeklyAvg'][1:],dataNeeded['freqAvg'][:-1])
print("WeeklyAvg vs. last week's freqAvg correlated (same week):", stat)
print('p val', pval)
stat, pval = pearsonr(dataNeeded['WeeklyChangeAvg'][2:],dataNeeded['frequencyChange'][1:-1])
print("WeeklyChangeAvg vs. last week's frequencyChange correlated (same week):", stat)
print('p val', pval)
stat, pval = pearsonr(dataNeeded['WeeklyChangeTotal'][2:],dataNeeded['frequencyChange'][1:-1])
print("WeeklyChangeTotal vs. last week's frequencyChange correlated (same week):", stat)
print('p val', pval)
stat, pval = pearsonr(dataNeeded['WeeklyAvg'][2:],dataNeeded['frequencyChange'][1:-1])
print("WeeklyAvg vs. last week's frequencyChange correlated (same week):", stat)
print('p val', pval)
stat, pval = pearsonr(dataNeeded['WeeklyChangeAvg'][:-1],dataNeeded['freqeuency'][1:])
print("last week's WeeklyChangeAvg vs. frequency correlated (same week):", stat)
print('p val', pval)
stat, pval = pearsonr(dataNeeded['WeeklyChangeTotal'][:-1],dataNeeded['freqeuency'][1:])
print("last week's WeeklyChangeTotal vs. frequency correlated (same week):", stat)
print('p val', pval)
stat, pval = pearsonr(dataNeeded['WeeklyAvg'][:-1],dataNeeded['freqeuency'][1:])
print("last week's WeeklyAvg vs. frequency correlated (same week):", stat)
print('p val', pval)
stat, pval = pearsonr(dataNeeded['WeeklyChangeAvg'][:-1],dataNeeded['freqAvg'][1:])
print("last week's WeeklyChangeAvg vs. freqAvg correlated (same week):", stat)
print('p val', pval)
stat, pval = pearsonr(dataNeeded['WeeklyChangeTotal'][:-1],dataNeeded['freqAvg'][1:])
print("last week's WeeklyChangeTotal vs. freqAvg correlated (same week):", stat)
print('p val', pval)
stat, pval = pearsonr(dataNeeded['WeeklyAvg'][:-1],dataNeeded['freqAvg'][1:])
print("last week's WeeklyAvg vs. freqAvg correlated (same week):", stat)
print('p val', pval)
stat, pval = pearsonr(dataNeeded['WeeklyChangeTotal'][1:-1],dataNeeded['frequencyChange'][2:])
print("last week's WeeklyChangeTotal vs. frequencyChange correlated (same week):", stat)
print('p val', pval)
stat, pval = pearsonr(dataNeeded['WeeklyChangeAvg'][1:-1],dataNeeded['frequencyChange'][2:])
print("last week's WeeklyChangeAvg vs. frequencyChange correlated:", stat)
print('p val', pval)
stat, pval = pearsonr(dataNeeded['WeeklyAvg'][1:-1],dataNeeded['frequencyChange'][2:])
print("last week's WeeklyAvg vs. frequencyChange correlated (same week):", stat)
print('p val', pval)
```
| github_jupyter |
```
from sys import path as syspath
from os import path as ospath
from cbsa import ReactionSystem
import stochpy as sp
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.gridspec as gridspec
from matplotlib import rc
rc('text', usetex=True)
plt.style.use("bmh")
plt.rcParams["font.family"] = "serif"
prop_cycle = plt.rcParams['axes.prop_cycle']
colors = prop_cycle.by_key()['color']
S = [[-1,-1],
[1,0],
[0,1]]
R = [[0,0],
[0,0],
[0,0]]
x = [100,0,0]
k = [.2,.6]
max_dt = 0.01
total_sim_time = 15
alpha = 0.5
cbsa = ReactionSystem(S)
cbsa.setup()
cbsa.set_x(x)
cbsa.set_k(k)
smod = sp.SSA()
smod.Model('Noise_Analysis.psc')
smod.ChangeParameter("k1",k[0])
smod.ChangeParameter("k2",k[1])
smod.ChangeInitialSpeciesCopyNumber("A",x[0])
smod.ChangeInitialSpeciesCopyNumber("B",x[1])
smod.ChangeInitialSpeciesCopyNumber("C",x[2])
cbsa_data = []
sp_ssa_data = []
sp_ssa_tau_data = []
replicates = 2000
for repl in range(replicates):
cbsa.setup_simulation(use_opencl=False,alpha=alpha,max_dt=max_dt)
cbsa.compute_simulation(total_sim_time,batch_steps=1)
cbsa_data.append(np.array(cbsa.simulation_data))
smod.DoStochSim(mode='time',end=total_sim_time,trajectories=1)
sp_ssa_data.append(smod.data_stochsim.getSpecies())
smod.DoStochSim(method='Tauleap',mode='time',end=total_sim_time,trajectories=1)
sp_ssa_tau_data.append(smod.data_stochsim.getSpecies())
plt.plot(cbsa_data[0][:,0],cbsa_data[0][:,1],color = "blue",alpha=0.03,label="A")
plt.plot(cbsa_data[0][:,0],cbsa_data[0][:,2],color = "red",alpha=0.03,label="B")
plt.plot(cbsa_data[0][:,0],cbsa_data[0][:,3],color = "green",alpha=0.03,label="C")
for i in range(1,500):
plt.plot(cbsa_data[i][:,0],cbsa_data[i][:,1],color = "blue",alpha=0.03)
plt.plot(cbsa_data[i][:,0],cbsa_data[i][:,2],color = "red",alpha=0.03)
plt.plot(cbsa_data[i][:,0],cbsa_data[i][:,3],color = "green",alpha=0.03)
plt.legend()
plt.show()
for i in range(replicates):
plt.plot(sp_ssa_data[i][:,0],sp_ssa_data[i][:,1],color = "blue",alpha=0.03)
plt.plot(sp_ssa_data[i][:,0],sp_ssa_data[i][:,2],color = "red",alpha=0.03)
plt.plot(sp_ssa_data[i][:,0],sp_ssa_data[i][:,3],color = "green",alpha=0.03)
plt.show()
for i in range(replicates):
plt.plot(sp_ssa_tau_data[i][:,0],sp_ssa_tau_data[i][:,1],color = "blue",alpha=0.03)
plt.plot(sp_ssa_tau_data[i][:,0],sp_ssa_tau_data[i][:,2],color = "red",alpha=0.03)
plt.plot(sp_ssa_tau_data[i][:,0],sp_ssa_tau_data[i][:,3],color = "green",alpha=0.03)
plt.show()
bins = 51
cbsa_ends = [cbsa_data[i][np.where(cbsa_data[i][:,1]==0)[0][0],0] for i in range(replicates) if len(np.where(cbsa_data[i][:,1]==0)[0])]
sp_ssa_ends = [sp_ssa_data[i][np.where(sp_ssa_data[i][:,1]==0)[0][0],0] for i in range(replicates) if len(np.where(sp_ssa_data[i][:,1]==0)[0])]
gilles_ssa_tau_ends = [sp_ssa_tau_data[i][np.where(sp_ssa_tau_data[i][:,1]==0)[0][0],0] for i in range(replicates) if len(np.where(sp_ssa_tau_data[i][:,1]==0)[0])]
plt.hist(cbsa_ends,density=True,bins=bins,histtype="step",linewidth=2,label = 'CBSA t '+str(np.round(np.mean(cbsa_ends),2))+" $\pm$"+str(np.round(np.std(cbsa_ends),2)))
plt.hist(sp_ssa_ends,density=True,bins=bins,histtype="step",linewidth=2,label = 'SSA t '+str(np.round(np.mean(sp_ssa_ends),2))+" $\pm$"+str(np.round(np.std(sp_ssa_ends),2)))
plt.hist(gilles_ssa_tau_ends,density=True,bins=bins,histtype="step",linewidth=2,label = r'$\tau$-leap t '+str(np.round(np.mean(sp_ssa_ends),2))+" $\pm$"+str(np.round(np.std(sp_ssa_ends),2)))
plt.legend()
plt.show()
bins = 15
cbsa_A = [cbsa_data[i][-1,1] for i in range(replicates)]
cbsa_B = [cbsa_data[i][-1,2] for i in range(replicates)]
cbsa_C = [cbsa_data[i][-1,3] for i in range(replicates)]
sp_ssa_A = [sp_ssa_data[i][-1,1] for i in range(replicates)]
sp_ssa_B = [sp_ssa_data[i][-1,2] for i in range(replicates)]
sp_ssa_C = [sp_ssa_data[i][-1,3] for i in range(replicates)]
sp_tauleap_A = [sp_ssa_tau_data[i][-1,1] for i in range(replicates)]
sp_tauleap_B = [sp_ssa_tau_data[i][-1,2] for i in range(replicates)]
sp_tauleap_C = [sp_ssa_tau_data[i][-1,3] for i in range(replicates)]
print("CBSA B mean",np.mean(cbsa_B),"variance",np.var(cbsa_B))
print("CBSA C mean",np.mean(cbsa_C),"variance",np.var(cbsa_C))
print("SP SSA B mean",np.mean(sp_ssa_B),"variance",np.var(sp_ssa_B))
print("SP SSA C mean",np.mean(sp_ssa_C),"variance",np.var(sp_ssa_C))
print("Tau-Leap B mean",np.mean(sp_tauleap_B),"variance",np.var(sp_tauleap_B))
print("Tau-Leap C mean",np.mean(sp_tauleap_C),"variance",np.var(sp_tauleap_C))
plt.hist(cbsa_B,density=True,alpha=0.5,histtype="step",linewidth=2,color="blue",bins=bins)
plt.hist(cbsa_C,density=True,alpha=0.5,histtype="step",linewidth=2,color="Blue",bins=bins)
plt.hist(sp_ssa_B,density=True,alpha=0.5,histtype="step",linewidth=2,color="red",bins=bins)
plt.hist(sp_ssa_C,density=True,alpha=0.5,histtype="step",linewidth=2,color="red",bins=bins)
plt.hist(sp_tauleap_B,density=True,alpha=0.5,histtype="step",linewidth=2,color="green",bins=bins)
plt.hist(sp_tauleap_C,density=True,alpha=0.5,histtype="step",linewidth=2,color="green",bins=bins)
plt.show()
fig = plt.figure(constrained_layout=True)
spec = gridspec.GridSpec(ncols=3, nrows=3, figure=fig)
ax1 = fig.add_subplot(spec[0:2, 0:2])
ax2 = fig.add_subplot(spec[2, 0:2])
ax3 = fig.add_subplot(spec[0:2, 2])
ax4 = fig.add_subplot(spec[2, 2])
for i in range(1,500):
ax1.plot(cbsa_data[i][:,0],cbsa_data[i][:,1],color = "blue",alpha=0.05)
ax1.plot(cbsa_data[i][:,0],cbsa_data[i][:,2],color = "red",alpha=0.05)
ax1.plot(cbsa_data[i][:,0],cbsa_data[i][:,3],color = "green",alpha=0.05)
ax1.set_ylabel("Number of Molecules")
ax1.text(0.5,98,"A",color="blue")
ax1.text(15,40,"B",color="red")
ax1.text(15,90,"C",color="green")
ax2.hist(cbsa_ends,bins=bins,histtype="step",linewidth=2,label = 'CBSA')
ax2.hist(sp_ssa_ends,bins=bins,histtype="step",linewidth=2,label = 'SSA')
#ax2.hist(gilles_ssa_tau_ends,density=True,bins=bins,histtype="step",linewidth=2,label = r'$\tau$-leap t '+str(np.round(np.mean(sp_ssa_ends),2))+" $\pm$"+str(np.round(np.std(sp_ssa_ends),2)))
ax2.set_xlim(ax1.get_xlim())
ax2.invert_yaxis()
ax2.set_xticklabels([])
ax2.xaxis.set_ticks_position('top')
ax2.set_xlabel("Time (s)")
ax2.set_ylabel("Frequency")
bins = 40
ax3.hist(cbsa_B+cbsa_C,histtype="step",linewidth=2,bins=bins,orientation='horizontal')
ax3.hist(sp_ssa_B+sp_ssa_C,histtype="step",linewidth=2,bins=bins,orientation='horizontal')
#ax3.hist(sp_tauleap_B+sp_tauleap_C,density=True,histtype="step",linewidth=2,bins=bins,orientation='horizontal')
ax3.set_ylim(ax1.get_ylim())
ax3.set_yticklabels([])
ax3.set_xlabel("Frequency")
h,l = ax2.get_legend_handles_labels()
ax4.legend(h,l)
ax4.get_xaxis().set_visible(False)
ax4.get_yaxis().set_visible(False)
ax4.set(frame_on=False)
fig.savefig("two_reactions.png",dpi=300, bbox_inches='tight')
```
| github_jupyter |
<font color=gray>ADS Sample Notebook.
Copyright (c) 2020, 2021 Oracle, Inc. All rights reserved.
Licensed under the Universal Permissive License v 1.0 as shown at https://oss.oracle.com/licenses/upl.
</font>
***
# Working with the Autonomous Databases
<p style="margin-left:10%; margin-right:10%;">by the <font color=teal> Oracle Cloud Infrastructure Data Science Team </font></p>
---
## Overview:
Data scientists often need to access information stored in databases. This notebook demonstrates how to connect to an Oracle Autonomous Database (ADB) and perform popular CRUD operations. Oracle offers two type of ADB, the Autonomous Data Warehouse (ADW) and the Autonomous Transaction Processing (ATP) databases. In general, there are no differences in how a connection is made to these different types of databases.
This notebook introduces how to pull data from an ADB into a notebook. It also demonstrates how to write generic SQL queries in the notebook, and several ways to connect.
**Important:**
Placeholder text for required values are surrounded by angle brackets that must be removed when adding the indicated content. For example, when adding a database name to `database_name = "<database_name>"` would become `database_name = "production"`.
---
## Prerequisites:
- An existing Autonomous Database, either an ADW or ATP.
- A user account on the ADB.
- If the ADB is accessible over the public internet, the notebook requires permissions to egress to the internet.
- Ensure the tenancy is [configured to use the Vault](https://docs.cloud.oracle.com/en-us/iaas/Content/KeyManagement/Tasks/managingvaults.htm).
This notebook is intended for beginners that need to connect to an ADB. It uses examples based on Python libraries and also basic SQL commands. Basic knowledge of the Python and SQL languages is necessary.
---
## Business Uses:
The Accelerate Data Science (ADS) SDK can load data from various resources and in a wide range of formats. The ADB is one of the most frequently used resources. For example, a business may have an existing dataset that is stored in ADW or ATP that could be loaded and explored with a notebook.
---
## Objectives:
- <a href='#setup'>Setting Up the Notebook Session to Access ADB</a>
- <a href='#setup_01'>Go to the database console</a>
- <a href='#setup_02'>Select the database</a>
- <a href='#setup_03'>Open DB Connection</a>
- <a href='#setup_04'>Download the wallet</a>
- <a href='#setup_05'>Rename the wallet</a>
- <a href='#setup_06'>Upload the wallet to the notebook</a>
- <a href='#setup_07'>Import the wallet</a>
- <a href='#setup_08'>Creating credentials</a>
- <a href='#setup_09'>Store Credentials</a>
- <a href='#setup_vault'>Oracle Cloud Infrastructure Vault</a>
- <a href='#setup_repository'>Repository</a>
- <a href='#setup_10'>Testing the internet connection</a>
- <a href='#setup_11'>Testing the connection to the ADB</a>
- <a href='#setup_12'>Connect to Autonomous Database</a>
- <a href='#load'>Loading an ADB Table as an `ADSDataset` Object</a>
- <a href='#run'>Running an SQL Query on ADB Using `SQLAlchemy` and `Pandas`</a>
- <a href='#create'>Creating a Table with `SQLAlchemy` and `Pandas` and Updating It</a>
- <a href='#query'>Querying Data from ADB using `cx_Oracle`</a>
- <a href='#ref'>References</a>
---
```
import cx_Oracle
import logging
import os
import pandas as pd
import warnings
from ads.database import connection
from ads.database.connection import Connector
from ads.dataset.factory import DatasetFactory
from ads.vault.vault import Vault
from sqlalchemy import create_engine
from urllib.request import urlopen
warnings.filterwarnings("ignore", category=DeprecationWarning)
logging.basicConfig(format='%(levelname)s:%(message)s', level=logging.INFO)
```
<a id='setup'></a>
# Setting Up the Notebook Session to Access ADB
To access ADB, (ADW or ATP databases) from the notebook environment, the database's wallet and user credentials are needed. The wallet is a ZIP file that contains connection information and the encryption keys that are needed to establish a secure connection to the database.
The following instructions are for the ADW though the steps are identical for an ATP database.
**Note: The Oracle Cloud Infrastructure Console is constantly evolving. While we strive to keep the images up to date, they are representative only.**
Upload the wallet to the notebook. The wallet can be obtained from your database administrator, Oracle Cloud Infrastructure API, Oracle CLI, or from the Console. In this example, it is downloaded from the Console.
<a id='setup_01'></a>
## 1. Go to the database console
In the Console, navigate to the **Autonomous Data Warehouse** or **Autonomous Transaction Processing** section.
<a id='setup_02'></a>
## 2. Select the database
Select the database to connect to. If one does not exist, one may need to be created.
<a id='setup_03'></a>
## 3. Open DB Connection
On the main page of the database, click **DB Connection** on an active database:

<a id='setup_04'></a>
## 4. Download the wallet
Click **Download Wallet**. You are asked to create a password. This password is used to access resources from some systems though it is not needed in the notebook.

<a id='setup_05'></a>
## 5. Rename the wallet
It has a name like `Wallet_<database_name>.zip`. For example, `Wallet_production1.zip`. Note the database name and the capitalization because it is used in step 7.
<a id='setup_06'></a>
## 6. Upload the wallet to the notebook
A script is used to configure the connection. This script assumes that the file has the format given in the preceding step and that the file exists in the `/home/datascience` directory. To upload the wallet, drag and drop it into the file browser, or click upload:

<a id='setup_08'></a>
## 7. Creating credentials
`ADS` methods for working with credentials uses a Python `dict` to store the key/value pairs. Therefore, any arbitrary values can be stored. Below is a common use case to store credentials to access an Oracle Autonomous Database.
In the `credential` variable, the following values can be updated to store the desired authentication information; `<database_name>`, `<user_name>`, and `<password>`.
```
# Sample credentials that are going to be stored.
credential = {'database_name': '<tns_name>',
'username': '<user_name>',
'password': '<password>',
'database_type':'oracle'}
```
<a id='setup_09'></a>
## 8. Store Credentials
The `ADS` library supports two mechanisms for storing credentials. It is best practice to store credentials outside of the notebook. Therefore, the preferred method is to use the Oracle Cloud Infrastructure Vault. If your tenancy is not configured to use the Vault, it is recommended that it be configured as this is the most secure method. If you cannot or prefer not to use the Vault, `ADS` supports storing the credentials on the local drive using the repository. The repository is a general-purpose mechanism for storing key/value pairs. The data is not encrypted and therefore is accessible by anyone with access to the notebook session.
<a id='setup_vault'></a>
### Oracle Cloud Infrastructure Vault
Ensure that the tenancy is [configured to use the Vault](https://docs.cloud.oracle.com/en-us/iaas/Content/KeyManagement/Tasks/managingvaults.htm).
The `ADS` vault methods assume that the data being stored is in a dictionary. Internally it stores it as an encoded JSON object.
In the following cell, update `<vault_id>` with the OCID for the vault that you wish to connect to. To encrypt/decrypt the data an encryption key is needed. This key is part of the vault. If one does not exist you may have to create it. Update `<key_id>` with the OCID of the encryption key.
```
vault_id = "<vault_id>"
key_id = "<key_id>"
```
The following cell obtains a handle to the vault and creates a secret. It returns the OCID to the secret so that it can be accessed.
```
# Create a secret
if vault_id != "<vault_id>" and key_id != "<key_id>":
vault = Vault(vault_id=vault_id, key_id=key_id)
secret_id = vault.create_secret(credential)
```
Using the secret OCID that is returned by the `create_secret()` method. The following cell obtains the secret from the vault and converts it to a dictionary. It is safe to hardcode this secret OCID into your notebook.
```
if "vault" in globals():
print(vault.get_secret(secret_id))
else:
print("Skipping as it appears that you do not have vault configured.")
```
There are a few attributes on the `Vault` class that provide information about the actual vault connections.
```
if "vault" in globals():
print(f"Vault OCID: {vault.id}")
print(f"Encryption Key OCID: {vault.key_id}")
print(f"Compartment OCID: {vault.compartment_id}")
else:
print("Skipping as it appears that you do not have vault configured.")
```
<a id='setup_repository'></a>
### Repository
The `ADS` repository methods create a general-purpose key/value store on the local block storage. This can be used to store any information. If you cannot or do not want to use the Vault to store the database credentials, then this method can be used.
If the block storage is destroyed the information will be lost but it is possible to share the files across notebook sessions.
The `update_repository()` method is used to update the values, if they exist in the repository, or create them if they do not.
The `get_repository` looks up the values associated with the key and returns a Python `dict`.
```
database_name = "<database_name>"
connection.update_repository(key=database_name, value=credential)
connection.get_repository(key=database_name)
```
<a id='setup_07'></a>
## 9. Import the wallet
The zipped wallet file must be unpacked and some of the files updated. The `import_wallet()` method will make the required changes.
In a previous step, the wallet file was named `Wallet_<database_name>.zip`. This information will be used to import the wallet into the database repository. Each database is keyed, generally with the name of that database. This information is used by the `ADS` `Connector` class to access the correct wallet.
```
wallet_path = os.path.join(os.path.expanduser("~"), f"Wallet_{database_name}.zip")
if database_name != "<database_name>":
connection.import_wallet(wallet_path=wallet_path, key=database_name)
```
<a id='setup_10'></a>
## 10. Testing the internet connection
Depending on the configuration of the tenancy, the notebook may require public internet access to access the ADB. The most common configurations require public internet access to connect to the ADB. Contact your system administrator.
This code checks to see if the notebook has public internet access:
```
try:
urlopen('http://oracle.com', timeout=5)
print("There is public internet access")
except:
print("There is no public internet access")
```
<a id='setup_11'></a>
## 11. Testing the connection to the ADB
The easiest way to test the ADB connection is to run the `sqlplus` CLI tool. The connection string is used to connect. This connection string is in the `\<user_name>/\<password>@\<SID>` format. For example, `\jdoe/\abc123@\prod1`.
Once it is working, the notebook demonstrates other methods to connect that are more practical for use within notebooks.
Messages similar to these indicate that the connection is successful:

<a id='setup_12'></a>
### 12. Connect to Autonomous Database
The `ADS` `Connector` class allows you several difference different ways to connect to the database. It is integrated with the Vault and the local repository. It also accepts arguments that are passed directly to the database. However, suppling credentials in the notebook is a security risk.
To make a connection using a secret in the Vault, set the parameter `secret_id` to the secret OCID.
```
if "secret_id" in globals():
connector = Connector(secret_id=secret_id)
else:
print("Skipping as it appears that you do not have secret_id configured.")
```
To use the repository with the `Connector` class, set the parameter `key` to the name of the repository key in the repository.
```
if "database_name" in globals() and database_name != '<database_name>':
connector = Connector(key=database_name)
else:
print("Skipping as it appears that you do not have database_name configured.")
```
It is also possible to hard code connection information into the `Connector` constructor to make a connection.
```
if credential['database_name'] != '<database_name>' and \
credential['username'] != '<user_name>' and \
credential['password'] != '<password>':
connector = Connector(database_name=credential['database_name'],
username=credential['username'],
password=credential['password'],
database_type=credential['database_type'])
```
The `config` attribute returns a dictionary of the configuration settings that were used to make the connection.
```
if "connector" in globals():
print(connector.config)
else:
print("Skipping as it appears that you do not have connector configured.")
```
When the `Connector` object is created it will make a connection to the database. If this connection is lost, the `connect()` method can be used to re-establish the connection.
```
if "connector" in globals():
connector.connect()
else:
print("Skipping as it appears that you do not have connector configured.")
```
<a id='load'></a>
# Loading an ADB Table as an `ADSDataset` Object
Now that there is a connection to ADB, it can be queried. In principle, this process only needs to be completed once. Before querying the database, load the credentials file content into memory.
The next step is to test the connection by pulling a table, and then load that table as an `ADSDataset` object through `DatasetFactory`.
By default, the ADB comes with the `sales` table in the `SH` schema. If that table has been removed, you have to update the code to provide the valid name of a table in a schema that you have access to.
ADS supports the ability to query based on an arbitrary SQL expression. In the preceding example, the `table` parameter contained the schema and table to be accessed. This parameter also takes an SQL expression.
In the next cell, the database returns the total sales by customer in descending order.
```
sql_total_sale = """
SELECT CUST_ID, SUM(AMOUNT_SOLD) AS TOTAL_SALES
FROM SH.SALES
GROUP BY CUST_ID
ORDER BY SUM(AMOUNT_SOLD) DESC
"""
if "connector" in globals():
total_sale = DatasetFactory.open(connector.uri, format="sql",
table=sql_total_sale)
else:
print("Skipping as it appears that you do not have connector configured.")
if "total_sale" in globals():
print(total_sale.head())
else:
print("Skipping as it appears that you do not have total_sale configured.")
if "total_sale" in globals():
total_sale.show_in_notebook()
else:
print("Skipping as it appears that you do not have total_sale configured.")
```
<a id='run'></a>
# Running an SQL Query on ADB Using SQLAlchemy and Pandas
`SQLAlchemy` is compatible with ADB. In the next cell, an `SQLAlchemy` `Engine` object is used to connect to the URI for the database. The `Pandas` `read_sql` method is used to run an arbitrary SQL statement.
```
# Make a connection to the Engine and run a query
if "connector" in globals():
engine = create_engine(connector.uri)
sql_customers = "SELECT * FROM SH.CUSTOMERS"
customers = pd.read_sql(sql_customers, con=engine)
print(customers.head())
```
`read_sql` is a convience method around `read_sql_query` and `read_sql_table`. Both of these methods are supported. Count the products in each category:
```
sql_category = """
SELECT PROD_CATEGORY, COUNT(*) AS N
FROM SH.PRODUCTS
GROUP BY PROD_CATEGORY
ORDER BY COUNT(*) DESC
"""
if "engine" in globals():
category = pd.read_sql_query(sql_category, con=engine)
print(category.head())
else:
print("Skipping as it appears that you do not have engine configured.")
```
Next, call the `read_sql_table` method on the products table:
```
if "engine" in globals():
products = pd.read_sql_table(table_name='products', con=engine, schema='SH')
print(products.head())
else:
print("Skipping as it appears that you do not have engine configured.")
```
<a id='create'></a>
# Creating a Table with `SQLAlchemy` and `Pandas` and Updating It
The preceding generic methods are all arbitrary SQL commands to be run to perform Data Manipulation Language commands to alter the data. However, a common use pattern for the data scientist is to pull data from a table, perform manipulations on it, and then push that data frame back into the database. This pattern is demonstrated by creating a table called `PRODUCT_DEMO`. If this table exists in the default schema, then change the name in the code as it is clobbered. The database user needs permissions to create and update a table.
Make a copy of the `PRODUCTS` table from the `SH` schema, and then create the table `PRODUCT_DEMO` in the default schema:
```
if "products" in globals():
products_demo = products.copy()
products_demo.to_sql(name='products_demo', index=False, con=engine, if_exists='replace')
print(products_demo.head())
else:
print("Skipping as it appears that you do not have products configured.")
```
Take a look at `PROD_ID = 116` in the `products_demo` data frame. The product name and description suggests that it a pack of 5:
```
if "products_demo" in globals():
print(products_demo.loc[products_demo.prod_id==116])
else:
print("Skipping as it appears that you do not have products_demo configured.")
```
This should be a pack of 50 so update the record in the `products_demo` data frame:
```
if "products_demo" in globals():
products_demo.loc[products_demo.loc[products_demo.prod_id==116].index, 'prod_name'] = 'CD-RW, High Speed Pack of 50'
products_demo.loc[products_demo.loc[products_demo.prod_id==116].index, 'prod_desc'] = 'CD-RW, High Speed 650MB/74 Minutes, Pack of 50'
print(products_demo.loc[products_demo.prod_id==116])
else:
print("Skipping as it appears that you do not have products_demo configured.")
```
When the database is queried, it does not reflect the change because it has not been pushed back to the database:
```
if "engine" in globals():
pd.read_sql_query('SELECT * FROM products_demo WHERE prod_id = 116', con=engine)
else:
print("Skipping as it appears that you do not have engine configured.")
```
Using the `to_sql` method the entire data frame is pushed back to the database and replaces the existing `products_demo` table. In this example, only a single record was changed. However, it is meant to illustrate that changes to a data frame can be pushed back to the database all together. This includes any updates to the data and structure of the data frame.
```
if "products_demo" in globals():
products_demo.to_sql(name='products_demo', index=False, con=engine, if_exists='replace')
pd.read_sql_query('SELECT * FROM products_demo WHERE prod_id = 116', con=engine)
else:
print("Skipping as it appears that you do not have products_demo configured.")
```
Drop the table that was created and close the `SQLAlchemy` `Engine` to free resources:
```
if "engine" in globals():
engine.execute("DROP TABLE PRODUCTS_DEMO")
engine.dispose()
else:
print("Skipping as it appears that you do not have engine configured.")
```
<a id='query'></a>
# Querying Data from ADB Using `cx_Oracle`
The `cx_Oracle` Python extension module enables access to an Oracle Database. It conforms to the Python database API 2.0 specification with a considerable number of additions and a couple of exclusions.
Using the `cx_Oracle` library is the preferred method and there is no need to create a JDBC connection string.
```
if credential['database_name'] != '<database_name>' and \
credential['username'] != '<user_name>' and \
credential['password'] != '<password>':
connection = cx_Oracle.connect(credential['username'],
credential['password'],
credential['database_name'])
cursor = connection.cursor()
sql_total_sale = "SELECT CUST_ID, SUM(AMOUNT_SOLD) AS TOTAL_SALES FROM SH.SALES GROUP BY CUST_ID ORDER BY SUM(AMOUNT_SOLD) DESC"
total_sale = cursor.execute(sql_total_sale)
data = total_sale.fetchall()
```
With `fetch_all` a `list` object is returned, not a `Pandas` data frame.
```
if "data" in globals():
print(type(data))
else:
print("Skipping as it appears that you do not have data configured.")
```
Use the standard `list` accessor techniquest to work with the data. This is the first record:
```
if "data" in globals():
print(data[0])
else:
print("Skipping as it appears that you do not have data configured.")
```
It is good practice to close connection and cursor objects when they are not being used. This prevents the database from being locked. Close them by calling the `close` method on each object:
```
if "cursor" in globals():
cursor.close()
connection.close()
else:
print("Skipping as it appears that you do not have cursor configured.")
```
<a id="reference"></a>
## References
- [Oracle ADS Library documentation](https://docs.cloud.oracle.com/en-us/iaas/tools/ads-sdk/latest/index.html)
- [Oracle Autonomous Database](https://docs.cloud.oracle.com/en-us/iaas/Content/Database/Concepts/adboverview.htm)
| github_jupyter |
## Post_Analysis2
- Dam classification by reservoir operation performances
- Hydrological Climate Classification (HCC) and Köppen-Geiger Climate Classification (KGC)
```
import os
import numpy as np
import pandas as pd
import geopandas as gpd
import rasterio
import matplotlib as mpl
import matplotlib.pyplot as plt
import matplotlib.gridspec as gridspec
import matplotlib.colors as colors
from mpl_toolkits.axes_grid1.inset_locator import inset_axes
import seaborn as sns
from scipy import stats
import HydroErr as he
from tools import save_hdf
from itertools import compress
from functools import reduce
import warnings
warnings.filterwarnings('ignore',category=pd.io.pytables.PerformanceWarning)
# Load Dam Inflow data from SUTD
dfFlowDams = pd.read_hdf('./data/dfFlowDams.hdf')
ind_dams = np.load('./data/ind_dams.npz')['ind_dams']
# Load Degree of Regulation (DOR)
dor = pd.read_hdf('./data/new_dor.hdf') # The order is sorted during the process
dam_dor = dor.loc[dor.DOR2 <= 0, 'GRAND_ID']
# Select only headwater dams (735)
idx_dor = np.isin(ind_dams[0,:], dam_dor)
damList = ind_dams[0,idx_dor]
ind_dams = ind_dams[1,idx_dor]
ndam = len(damList)
# KGE score at MP1
kge = pd.read_hdf('./data/subset.hdf', 'df').reset_index(drop=False)[['GRAND_ID','KGE_MP1']]
# Load hydropower production results
prod = pd.read_hdf('./data/hydropower_production.hdf').loc[damList].reset_index()[['GRAND_ID','PF','DF']]
# Dam characteristics
char = pd.read_csv('./data/res_obj_dor.csv')[['GRAND_ID','ECAP','MAIN_USE']].set_index('GRAND_ID').loc[damList].reset_index()
# Climate classifications and Continents and Countries
cclass = pd.read_hdf('./data/climate_classification.hdf').loc[damList].reset_index()
# Merge DataFrames
data_frames = [kge,prod,char,cclass]
data_basic = reduce(lambda left,right: pd.merge(left,right,on=['GRAND_ID'],how='inner'), data_frames)
data_basic = data_basic.rename(columns={'KGE_MP1':'KGE'})
data_basic.head()
```
### Dam Classification
The regression is performed by Jia.
```
# Load regression variables
df_var = pd.read_hdf('./data/regression_variables.hdf').reset_index()
# Subset of required indicators
target_var = ['dep_over_head2', 'perc_qmax', 'fill_mean_mon']
subset = pd.merge(data_basic[['GRAND_ID','PF', 'DF', 'KGE']], df_var[['GRAND_ID', *target_var]], on='GRAND_ID', how='inner').set_index('GRAND_ID')
subset.columns = ['PF', 'DF', 'KGE', 'depth', 'qmax', 'fill']
# Threshold regressed by Jia
goodDam = subset['PF'] > 10
goodForecast = subset['KGE'] > 0.463
anyForecast = (subset['depth'] < 0.630) \
| (subset['qmax'] > 0.293) \
| (subset['fill'] < 0.255)
# Classification
type_a1 = goodDam & goodForecast & anyForecast
type_a2 = goodDam & goodForecast & ~anyForecast
type_b = ~goodDam & goodForecast
type_c1 = goodDam & ~goodForecast & anyForecast
type_c2 = goodDam & ~goodForecast & ~anyForecast
type_d = ~goodDam & ~goodForecast
print('='*45)
print('Number of dams in classifications')
print('-'*45)
print('AI:\t%d (%.1f%%)' % (sum(type_a1), sum(type_a1)/ndam*100))
print('AII:\t%d (%.1f%%)' % (sum(type_a2), sum(type_a2)/ndam*100))
print('B:\t%d (%.1f%%)' % (sum(type_b), sum(type_b)/ndam*100))
print('CI:\t%d (%.1f%%)' % (sum(type_c1), sum(type_c1)/ndam*100))
print('CII:\t%d (%.1f%%)' % (sum(type_c2), sum(type_c2)/ndam*100))
print('D:\t%d (%.1f%%)' % (sum(type_d), sum(type_d)/ndam*100))
print('Total:\t%d' % (sum(type_a1)+sum(type_a2)+sum(type_b)+sum(type_c1) \
+ sum(type_c2) + sum(type_d)))
print('='*45)
# Improvements by classifcation
ClassName = ['AI','AII','B','CI','CII','D']
impv6D = pd.concat([subset[ind]['DF'] for ind in list((type_a1, type_a2, type_b, type_c1, type_c2, type_d))], axis=1)
impv6D.columns = ClassName
impv6P = pd.concat([subset[ind]['PF'] for ind in list((type_a1, type_a2, type_b, type_c1, type_c2, type_d))], axis=1)
impv6P.columns = ClassName
# Series of classification
DamClass = pd.Series(data=np.zeros([ndam]),
index = type_a1.index,
name='class')
DamClass[type_a1] = 1
DamClass[type_a2] = 2
DamClass[type_b] = 3
DamClass[type_c1] = 4
DamClass[type_c2] = 5
DamClass[type_d] = 6
```
### Merge DataFrames
```
# Merge DataFrames
data_frames = [data_basic,DamClass]
data = reduce(lambda left,right: pd.merge(left,right,on=['GRAND_ID'],how='inner'), data_frames)
data = data.rename(columns={'KGE_MP1':'KGE'})
# Save data
save_hdf('./data/data_analysis.hdf', data)
data.to_excel('./data/dam_climate_classification.xlsx')
data.head()
```
### Scatter plot with HCC indices
```
data2 = data.copy()
data2['class'] = data['class'].replace({1:'AI', 2:'AII', 3:'B', 4:'CI', 5:'CII', 6:'D'})
sns.set(rc={'figure.figsize':(6,6)})
sns.set_style("whitegrid")
# fig, ax = plt.subplots(nrows=1, ncols=1, figsize=(7,7))
g = sns.PairGrid(data2,
x_vars=["aridity", "seasonality", "snow"],
y_vars=["KGE", "PF", "DF"],
hue='class')
g = g.map(plt.scatter, s=10)
g = g.add_legend()
if False:
fn_save = './figures/aggregation_hcc.png'
g.savefig(fn_save, bbox_inches='tight')
print('%s is saved.' % fn_save)
### ADD DAM CLASSIFICATION COLORS
```
### Boxplots of KGE, PF, DF according to KGC
```
# KGC color scheme
code2 = pd.read_excel('./data/koppen_geiger_classification/code_rgb.xlsx').set_index('Code')
csch = code2.loc[np.unique(data['kgc2']), ['Red','Green','Blue']].values
# Figure setting
sns.set_style("whitegrid")
sns.set_palette(csch/255)
fig, axes = plt.subplots(nrows=1, ncols=3, figsize=(9,7),
sharey=True, sharex=False,
gridspec_kw={'width_ratios':[1,1,1]})
# Plotting
name = ['(a) KGE', '(b) PF (%)', '(c) DF (%)']
for (i, el) in enumerate(['KGE', 'PF', 'DF']):
ax = axes.flatten('F')[i]
sns.boxplot(ax=ax, x=el, y='kgc2', data=data,
order = np.unique(data['kgc2']),
width=0.65, linewidth=1.5, fliersize=4)
ax.set(xlabel="", ylabel="")
ax.annotate(name[i], [0.5,1.03], xycoords='axes fraction',
color='k', ha='center', va='center', fontfamily='sans-serif', fontsize=15)
ax.xaxis.grid(True)
ax.tick_params(axis='both', which='major', labelsize=14)
if i == 0:
ax.set_xlim([0, 1])
ax.set_xticks(np.arange(0,1.1,0.25))
ax.set_xticklabels(['0','0.25','0.5','0.75','1'], fontsize=15, fontname='arial')
elif i == 1:
ax.set_xlim([0, 50])
elif i == 2:
ax.set_xlim([-30, 30])
ax.tick_params(axis='x', which='major')
fig.tight_layout(pad=0.7)
plt.show()
if False:
fn_save = './figures/aggregation_kgc.pdf'
fig.savefig(fn_save, bbox_inches='tight')
print('%s is saved.' % fn_save)
fn_save = './figures/aggregation_kgc.png'
fig.savefig(fn_save, bbox_inches='tight')
print('%s is saved.' % fn_save)
# KGC color scheme
code2 = pd.read_excel('./data/koppen_geiger_classification/code_rgb.xlsx').set_index('Code')
csch = code2.loc[np.unique(data['kgc2']), ['Red','Green','Blue']].values
# Figure setting
sns.set_style("whitegrid")
sns.set_palette(csch/255)
fig, axes = plt.subplots(nrows=3, ncols=1, figsize=(9,7),
sharex=True, sharey=False,
gridspec_kw={'height_ratios':[1,1,1]})
# Plotting
name = ['(a) PF (%)', '(b) DF (%)', '(c) KGE']
for (i, el) in enumerate(['PF', 'DF', 'KGE']):
ax = axes.flatten('F')[i]
sns.boxplot(ax=ax, x='kgc2', y=el, data=data,
order = np.unique(data['kgc2']),
width=0.65, linewidth=1.5, fliersize=4)
ax.set(xlabel="", ylabel="")
ax.annotate(name[i], [-0.09,0.5], xycoords='axes fraction', rotation=90,
color='k', ha='center', va='center', fontfamily='sans-serif', fontsize=15)
ax.xaxis.grid(True)
ax.tick_params(axis='both', which='major', labelsize=14)
ax.set_xticklabels(ax.get_xticklabels(), fontsize=14, fontname='arial', rotation=90)
if el == 'KGE':
ax.set_ylim([0, 1])
ax.set_yticks(np.arange(0,1.1,0.25))
ax.set_yticklabels(['0','0.25','0.5','0.75','1'], fontsize=15, fontname='arial')
elif el == 'PF':
ax.set_ylim([0, 50])
elif el == 'DF':
ax.set_ylim([-30, 30])
ax.tick_params(axis='y', which='major')
fig.tight_layout(pad=0.7)
plt.show()
if True:
fn_save = './figures/aggregation_kgc.pdf'
fig.savefig(fn_save, bbox_inches='tight')
print('%s is saved.' % fn_save)
fn_save = './figures/aggregation_kgc.png'
fig.savefig(fn_save, bbox_inches='tight')
print('%s is saved.' % fn_save)
data['I'] = data['DF']/data['PF']
data['I'].max()
```
### Aggregation by continents
```
sns.set_style('whitegrid')
fig, axes = plt.subplots(nrows=3, ncols=1, figsize=(7,7),
sharey=False, sharex=True,
gridspec_kw={'height_ratios':[1,1,1]},
facecolor='w')
# Plotting
name = ['(a) KGE', '(b) PF (%)', '(c) DF (%)']
cont_order = ['North America', 'South America', 'Africa','Europe', 'Asia', 'Oceania']
cont_name = ['North\nAmerica', 'South\nAmerica', 'Africa','Europe', 'Asia', 'Oceania']
prct_dam = np.round(data.groupby('CONTINENT')['COUNTRY'].count()/ndam*100)[cont_order]
cont_name = ['%s\n(%d%%)' % (cname,cprct) for cname, cprct in zip(cont_name, list(prct_dam))]
for (i, el) in enumerate(['KGE', 'PF', 'DF']):
ax = axes.flatten('F')[i]
sns.boxplot(ax=ax, x='CONTINENT', y=el, data=data,
order = cont_order, palette = 'Paired',
width=0.65, linewidth=1.5, fliersize=4)
ax.set(xlabel="", ylabel="")
ax.annotate(name[i], [-0.13, 0.5],
rotation=90, xycoords='axes fraction',
color='k', ha='center', va='center', fontfamily='sans-serif', fontsize=15)
ax.tick_params(axis='both', which='major', labelsize=14)
ax.yaxis.grid(True)
if i == 0:
ax.set_ylim([0, 1])
ax.set_yticks(np.arange(0,1.1,0.25))
ax.set_yticklabels(['0','0.25','0.5','0.75','1'], fontsize=14, fontname='sans-serif')
elif i == 1:
ax.set_ylim([0, 50])
elif i == 2:
ax.set_ylim([-30, 30])
ax.set_xticklabels(cont_name, fontfamily='sans-serif', fontsize=15, va='center')
ax.tick_params(axis='x', which='major', pad=25)
# Finalizing
fig.tight_layout(pad=0.7)
plt.show()
fn_save = './figures/aggregation_continent.pdf'
fig.savefig(fn_save, bbox_inches='tight')
print('%s is saved.' % fn_save)
fn_save = './figures/aggregation_continent.png'
fig.savefig(fn_save, bbox_inches='tight')
print('%s is saved.' % fn_save)
### Update figure
# 1) Merge PF and DF
# 2) At the third row, 6 boxplots representing 6 classes per each continent
data2['class'] = data['class'].replace({1:'AI', 2:'AII', 3:'B', 4:'CI', 5:'CII', 6:'D'})
data2[['CONTINENT', 'class']].groupby('CONTINENT').value_count()
```
### Dam classification scatter plot
```
temp = data[['GRAND_ID','PF','DF','KGE']]
temp = temp.merge(DamClass.reset_index(), on='GRAND_ID', how='inner')
# Colormap
colorsList = ['lightcoral','crimson',
'gold',
'lightgreen', 'green',
'cornflowerblue']
cmap = colors.ListedColormap(colorsList)
fig, axes = plt.subplots(nrows=2, ncols=2, figsize=(8.5,7.5),
sharey=True, sharex=False,
gridspec_kw={'width_ratios':[1.8,1]})
# (a) Scatterplot of PF and wKGE
ax = axes.flatten('C')[0]
sc = sns.scatterplot(data=temp, x='KGE', y='PF', ax=ax, marker='o',
linewidth=0.5, alpha=0.5, palette=cmap,
hue='class', hue_norm=(1, 6))
ax.plot([-1, 1], [10, 10], color='grey', lw=1, alpha=0.5, linestyle='--')
ax.plot([0.463, 0.463], [-100, 100], color='grey', lw=1, alpha=0.5, linestyle='--')
ax.get_legend().remove()
sc.tick_params(labelsize=13)
ax.yaxis.set_tick_params(which='both', labelbottom=True)
ax.set_xticklabels([])
ax.set_xlabel('')
ax.set_yticks(np.arange(-20,65,10))
ax.set_ylabel(r'$I_{PF}$ (%)', fontsize=15, fontname='arial')
ax.set_xlim([-0.1, 1])
ax.set_ylim([-20, 65])
ax.annotate('(a)', xy=(0.06, 0.92), xycoords='axes fraction',
horizontalalignment='center', verticalalignment='center',
fontname='arial',fontsize=20)
# (b) Boxplot of PF and classifications
ax = axes.flatten('C')[1]
sc = sns.boxplot(data=impv6P, ax=ax, width=0.7, palette=colorsList)
sc.plot([-10, 10], [10, 10], color='grey', lw=1, alpha=0.5, linestyle='--')
sc.tick_params(labelsize=13)
sc.set_xlim([-0.7, 5.7])
ax.set_xticklabels([])
ax.annotate('(b)', xy=(0.90, 0.92), xycoords='axes fraction',
horizontalalignment='center', verticalalignment='center',
fontname='arial',fontsize=20)
# (c) Scatterplot of PF and wKGE
ax = axes.flatten('C')[2]
sc = sns.scatterplot(data=temp, x='KGE', y='DF', ax=ax, marker='o',
linewidth=0.5, alpha=0.5, palette=cmap,
hue='class', hue_norm=(1, 6))
sc.tick_params(labelsize=12)
ax.set_xlim([-0.1, 1])
ax.plot([-1, 1], [10, 10], color='gray', lw=1, alpha=0.5, linestyle='--')
ax.plot([0.463, 0.463], [-100, 100], color='gray', lw=1, alpha=0.5, linestyle='--')
ax.get_legend().remove()
ax.tick_params(labelsize=13)
ax.set_ylabel(r'$I_{DF}$ (%)', fontsize=15, fontname='arial')
ax.set_xlim([-0.1, 1])
ax.set_ylim([-20, 65])
ax.set_xlabel('KGE', fontname='arial', fontsize=15,
horizontalalignment='center', labelpad = 13)
ax.annotate('(c)', xy=(0.06, 0.92), xycoords='axes fraction',
horizontalalignment='center', verticalalignment='center',
fontname='arial',fontsize=20)
# (d) Boxplot of PF and classifications
ax = axes.flatten('C')[3]
sc = sns.boxplot(data=impv6D, ax=ax, width=0.7, palette=colorsList)
sc.plot([-10, 10], [10, 10], color='gray', lw=1, alpha=0.5, linestyle='--')
ax.set_xlabel('Types', fontname='arial', fontsize=15,
horizontalalignment='center', labelpad = 13)
sc.tick_params(labelsize=13)
sc.set_xlim([-0.7, 5.7])
ax.annotate('(d)', xy=(0.90, 0.92), xycoords='axes fraction',
horizontalalignment='center', verticalalignment='center',
fontname='arial',fontsize=20)
plt.tight_layout(w_pad=0.02, h_pad=0.1)
plt.show()
# Save a figure
if False:
fn_save = './figures/impv_classification.png'
fig.savefig(fn_save, bbox_inches='tight')
print('%s is saved.' % fn_save)
```
| github_jupyter |
```
%reload_ext autoreload
%autoreload 2
%matplotlib inline
import matplotlib.pyplot as plt
```
pip install pytorch-transformers
```
import torch
import torch.nn as nn
import pickle
from torch.utils.data import (DataLoader, RandomSampler, SequentialSampler,
TensorDataset)
from tqdm import tqdm_notebook, trange
import os
from pytorch_transformers import BertConfig, BertTokenizer, BertModel
from pytorch_transformers.optimization import AdamW, WarmupLinearSchedule
from torch.utils.data import Dataset, DataLoader
import numpy as np
import torch.optim as optim
from torch.optim import lr_scheduler
import time
import copy
import torch.nn.functional as F
```
## Pretrained bert model
```
class BertForSequenceClassification(nn.Module):
"""BERT model for classification.
This module is composed of the BERT model with a linear layer on top of
the pooled output.
"""
def __init__(self, num_labels=1):
super(BertForSequenceClassification, self).__init__()
self.num_labels = num_labels
self.bert = BertModel.from_pretrained('bert-base-cased')
self.dropout = nn.Dropout(config.hidden_dropout_prob)
self.classifier = nn.Linear(config.hidden_size, num_labels)
nn.init.xavier_normal_(self.classifier.weight)
def forward(self, input_ids, token_type_ids=None, attention_mask=None, labels=None):
outputs = self.bert(input_ids, token_type_ids, attention_mask)
pooled_output = outputs[1]
pooled_output = self.dropout(pooled_output)
logits = self.classifier(pooled_output)
return logits
def freeze_bert_encoder(self):
for param in self.bert.parameters():
param.requires_grad = False
def unfreeze_bert_encoder(self):
for param in self.bert.parameters():
param.requires_grad = True
config = BertConfig(vocab_size_or_config_json_file=32000, hidden_size=768,
num_hidden_layers=12, num_attention_heads=12, intermediate_size=3072)
num_labels = 1
model = BertForSequenceClassification(num_labels)
from pathlib import Path
PATH = Path("/data2/yinterian/aclImdb/")
tokenizer = BertTokenizer.from_pretrained('bert-base-cased')
tokenizer.vocab_size
path = PATH/"train/pos/0_9.txt"
z = tokenizer.tokenize(path.read_text())
z[:10]
ids = tokenizer.convert_tokens_to_ids(z)
ids[:10]
tokens_tensor = torch.tensor([ids])
logits = model(tokens_tensor)
logits
```
Based on these tutorials
* https://pytorch.org/hub/huggingface_pytorch-pretrained-bert_bert/
* https://github.com/huggingface/pytorch-transformers/blob/master/README.md
* https://medium.com/huggingface/multi-label-text-classification-using-bert-the-mighty-transformer-69714fa3fb3d
* https://towardsdatascience.com/bert-classifier-just-another-pytorch-model-881b3cf05784
```
def text2ids(text, max_seq_length=300):
tok_text = tokenizer.tokenize(text)
if len(tok_text) > max_seq_length:
tok_text = tok_text[:max_seq_length]
ids_text = tokenizer.convert_tokens_to_ids(tok_text)
padding = [0] * (max_seq_length - len(ids_text))
ids_text += padding
return np.array(ids_text)
text2ids(path.read_text())
class ImdbDataset(Dataset):
def __init__(self, PATH, train="train"):
self.path_to_images = PATH/train
self.pos_files = list((self.path_to_images/"pos").iterdir())
self.neg_files = list((self.path_to_images/"neg").iterdir())
self.files = self.pos_files + self.neg_files
self.y = np.concatenate((np.ones(len(self.pos_files), dtype=int),
np.zeros(len(self.neg_files), dtype=int)), axis=0)
def __getitem__(self, index):
path = self.files[index]
x = text2ids(path.read_text())
return x, self.y[index]
def __len__(self):
return len(self.y)
train_ds = ImdbDataset(PATH)
valid_ds = ImdbDataset(PATH, "test")
batch_size = 10
train_dl = DataLoader(train_ds, batch_size=batch_size, shuffle=True)
valid_dl = DataLoader(valid_ds, batch_size=batch_size)
```
## train
```
def train_model(model, optimizer, num_epochs=25):
for epoch in range(num_epochs):
model.train()
running_loss = 0.0
for x, y in train_dl:
x = x.cuda()
y = y.unsqueeze(1).float().cuda()
optimizer.zero_grad()
logits = model(x)
loss = F.binary_cross_entropy_with_logits(logits, y)
loss.backward()
optimizer.step()
running_loss += loss.item() * x.size(0)
epoch_loss = running_loss / len(train_ds)
val_loss, accuracy = eval_model(model)
print('train loss: {:.3f}, valid loss {:.3f} accuracy {:.3f}'.format(
epoch_loss, val_loss, accuracy))
def eval_model(model):
model.eval()
running_loss = 0.0
correct = 0
for x, y in valid_dl:
x = x.cuda()
y = y.unsqueeze(1).float().cuda()
logits = model(x)
loss = F.binary_cross_entropy_with_logits(logits, y)
y_pred = logits > 0
correct += (y_pred.float() == y).float().sum()
running_loss += loss.item() * x.size(0)
accuracy = correct / len(valid_ds)
epoch_loss = running_loss / len(valid_ds)
return epoch_loss, accuracy.item()
model = BertForSequenceClassification(num_labels).cuda()
lrlast = .0001
lrmain = .00001
optimizer = optim.Adam(
[
{"params":model.bert.parameters(),"lr": lrmain},
{"params":model.classifier.parameters(), "lr": lrlast},
])
train_model(model, optimizer, num_epochs=2)
```
| github_jupyter |
##### Copyright 2019 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# 深度卷积生成对抗网络
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://tensorflow.google.cn/tutorials/generative/dcgan">
<img src="https://tensorflow.google.cn/images/tf_logo_32px.png" />
在 tensorFlow.google.cn 上查看</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/zh-cn/tutorials/generative/dcgan.ipynb">
<img src="https://tensorflow.google.cn/images/colab_logo_32px.png" />
在 Google Colab 中运行</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/zh-cn/tutorials/generative/dcgan.ipynb">
<img src="https://tensorflow.google.cn/images/GitHub-Mark-32px.png" />
在 GitHub 上查看源代码</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/docs-l10n/site/zh-cn/tutorials/generative/dcgan.ipynb"><img src="https://tensorflow.google.cn/images/download_logo_32px.png" />下载 notebook</a>
</td>
</table>
Note: 我们的 TensorFlow 社区翻译了这些文档。因为社区翻译是尽力而为, 所以无法保证它们是最准确的,并且反映了最新的
[官方英文文档](https://tensorflow.google.cn/?hl=en)。如果您有改进此翻译的建议, 请提交 pull request 到
[tensorflow/docs](https://github.com/tensorflow/docs) GitHub 仓库。要志愿地撰写或者审核译文,请加入
[docs-zh-cn@tensorflow.org Google Group](https://groups.google.com/a/tensorflow.org/forum/#!forum/docs-zh-cn)。
本教程演示了如何使用[深度卷积生成对抗网络](https://arxiv.org/pdf/1511.06434.pdf)(DCGAN)生成手写数字图片。该代码是使用 [Keras Sequential API](https://tensorflow.google.cn/guide/keras) 与 `tf.GradientTape` 训练循环编写的。
## 什么是生成对抗网络?
[生成对抗网络](https://arxiv.org/abs/1406.2661)(GANs)是当今计算机科学领域最有趣的想法之一。两个模型通过对抗过程同时训练。一个*生成器*(“艺术家”)学习创造看起来真实的图像,而*判别器*(“艺术评论家”)学习区分真假图像。
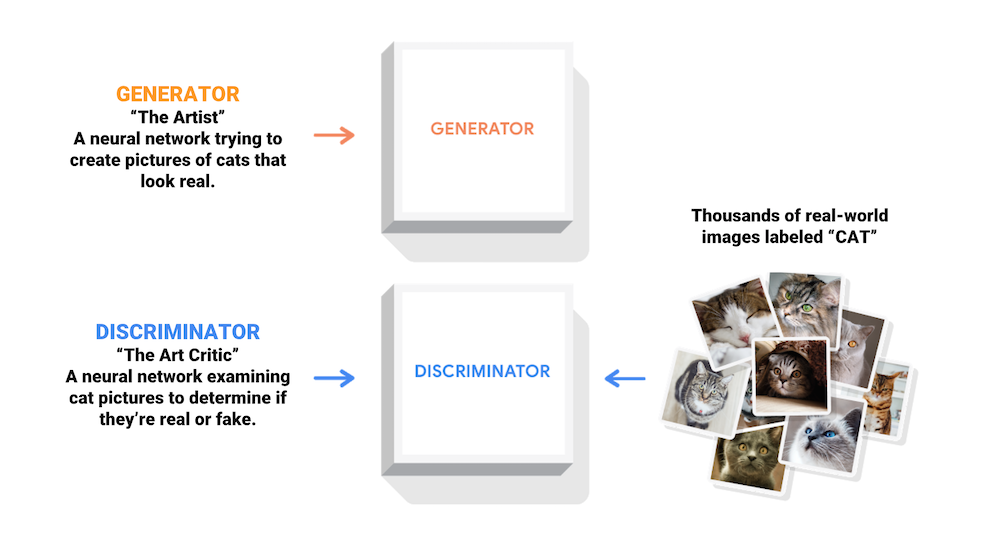
训练过程中,*生成器*在生成逼真图像方面逐渐变强,而*判别器*在辨别这些图像的能力上逐渐变强。当*判别器*不再能够区分真实图片和伪造图片时,训练过程达到平衡。
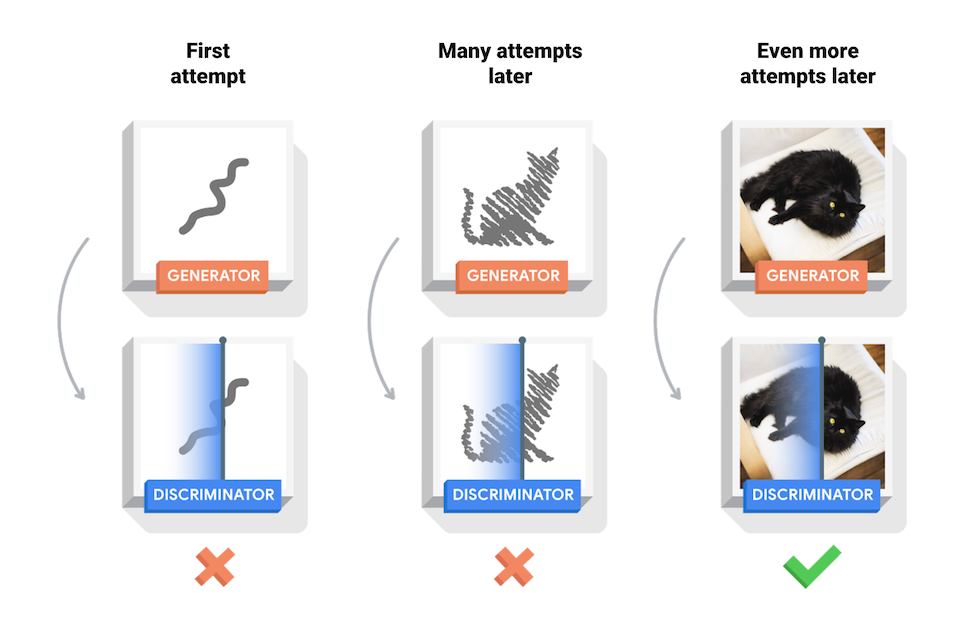
本笔记在 MNIST 数据集上演示了该过程。下方动画展示了当训练了 50 个epoch (全部数据集迭代50次) 时*生成器*所生成的一系列图片。图片从随机噪声开始,随着时间的推移越来越像手写数字。

要了解关于 GANs 的更多信息,我们建议参阅 MIT的 [深度学习入门](http://introtodeeplearning.com/) 课程。
### Import TensorFlow and other libraries
```
import tensorflow as tf
tf.__version__
# 用于生成 GIF 图片
!pip install -q imageio
import glob
import imageio
import matplotlib.pyplot as plt
import numpy as np
import os
import PIL
from tensorflow.keras import layers
import time
from IPython import display
```
### 加载和准备数据集
您将使用 MNIST 数据集来训练生成器和判别器。生成器将生成类似于 MNIST 数据集的手写数字。
```
(train_images, train_labels), (_, _) = tf.keras.datasets.mnist.load_data()
train_images = train_images.reshape(train_images.shape[0], 28, 28, 1).astype('float32')
train_images = (train_images - 127.5) / 127.5 # 将图片标准化到 [-1, 1] 区间内
BUFFER_SIZE = 60000
BATCH_SIZE = 256
# 批量化和打乱数据
train_dataset = tf.data.Dataset.from_tensor_slices(train_images).shuffle(BUFFER_SIZE).batch(BATCH_SIZE)
```
## 创建模型
生成器和判别器均使用 [Keras Sequential API](https://tensorflow.google.cn/guide/keras#sequential_model) 定义。
### 生成器
生成器使用 `tf.keras.layers.Conv2DTranspose` (上采样)层来从种子(随机噪声)中产生图片。以一个使用该种子作为输入的 `Dense` 层开始,然后多次上采样直到达到所期望的 28x28x1 的图片尺寸。注意除了输出层使用 tanh 之外,其他每层均使用 `tf.keras.layers.LeakyReLU` 作为激活函数。
```
def make_generator_model():
model = tf.keras.Sequential()
model.add(layers.Dense(7*7*256, use_bias=False, input_shape=(100,)))
model.add(layers.BatchNormalization())
model.add(layers.LeakyReLU())
model.add(layers.Reshape((7, 7, 256)))
assert model.output_shape == (None, 7, 7, 256) # 注意:batch size 没有限制
model.add(layers.Conv2DTranspose(128, (5, 5), strides=(1, 1), padding='same', use_bias=False))
assert model.output_shape == (None, 7, 7, 128)
model.add(layers.BatchNormalization())
model.add(layers.LeakyReLU())
model.add(layers.Conv2DTranspose(64, (5, 5), strides=(2, 2), padding='same', use_bias=False))
assert model.output_shape == (None, 14, 14, 64)
model.add(layers.BatchNormalization())
model.add(layers.LeakyReLU())
model.add(layers.Conv2DTranspose(1, (5, 5), strides=(2, 2), padding='same', use_bias=False, activation='tanh'))
assert model.output_shape == (None, 28, 28, 1)
return model
```
使用(尚未训练的)生成器创建一张图片。
```
generator = make_generator_model()
noise = tf.random.normal([1, 100])
generated_image = generator(noise, training=False)
plt.imshow(generated_image[0, :, :, 0], cmap='gray')
```
### 判别器
判别器是一个基于 CNN 的图片分类器。
```
def make_discriminator_model():
model = tf.keras.Sequential()
model.add(layers.Conv2D(64, (5, 5), strides=(2, 2), padding='same',
input_shape=[28, 28, 1]))
model.add(layers.LeakyReLU())
model.add(layers.Dropout(0.3))
model.add(layers.Conv2D(128, (5, 5), strides=(2, 2), padding='same'))
model.add(layers.LeakyReLU())
model.add(layers.Dropout(0.3))
model.add(layers.Flatten())
model.add(layers.Dense(1))
return model
```
使用(尚未训练的)判别器来对图片的真伪进行判断。模型将被训练为为真实图片输出正值,为伪造图片输出负值。
```
discriminator = make_discriminator_model()
decision = discriminator(generated_image)
print (decision)
```
## 定义损失函数和优化器
为两个模型定义损失函数和优化器。
```
# 该方法返回计算交叉熵损失的辅助函数
cross_entropy = tf.keras.losses.BinaryCrossentropy(from_logits=True)
```
### 判别器损失
该方法量化判别器从判断真伪图片的能力。它将判别器对真实图片的预测值与值全为 1 的数组进行对比,将判别器对伪造(生成的)图片的预测值与值全为 0 的数组进行对比。
```
def discriminator_loss(real_output, fake_output):
real_loss = cross_entropy(tf.ones_like(real_output), real_output)
fake_loss = cross_entropy(tf.zeros_like(fake_output), fake_output)
total_loss = real_loss + fake_loss
return total_loss
```
### 生成器损失
生成器损失量化其欺骗判别器的能力。直观来讲,如果生成器表现良好,判别器将会把伪造图片判断为真实图片(或 1)。这里我们将把判别器在生成图片上的判断结果与一个值全为 1 的数组进行对比。
```
def generator_loss(fake_output):
return cross_entropy(tf.ones_like(fake_output), fake_output)
```
由于我们需要分别训练两个网络,判别器和生成器的优化器是不同的。
```
generator_optimizer = tf.keras.optimizers.Adam(1e-4)
discriminator_optimizer = tf.keras.optimizers.Adam(1e-4)
```
### 保存检查点
本笔记还演示了如何保存和恢复模型,这在长时间训练任务被中断的情况下比较有帮助。
```
checkpoint_dir = './training_checkpoints'
checkpoint_prefix = os.path.join(checkpoint_dir, "ckpt")
checkpoint = tf.train.Checkpoint(generator_optimizer=generator_optimizer,
discriminator_optimizer=discriminator_optimizer,
generator=generator,
discriminator=discriminator)
```
## 定义训练循环
```
EPOCHS = 50
noise_dim = 100
num_examples_to_generate = 16
# 我们将重复使用该种子(因此在动画 GIF 中更容易可视化进度)
seed = tf.random.normal([num_examples_to_generate, noise_dim])
```
训练循环在生成器接收到一个随机种子作为输入时开始。该种子用于生产一张图片。判别器随后被用于区分真实图片(选自训练集)和伪造图片(由生成器生成)。针对这里的每一个模型都计算损失函数,并且计算梯度用于更新生成器与判别器。
```
# 注意 `tf.function` 的使用
# 该注解使函数被“编译”
@tf.function
def train_step(images):
noise = tf.random.normal([BATCH_SIZE, noise_dim])
with tf.GradientTape() as gen_tape, tf.GradientTape() as disc_tape:
generated_images = generator(noise, training=True)
real_output = discriminator(images, training=True)
fake_output = discriminator(generated_images, training=True)
gen_loss = generator_loss(fake_output)
disc_loss = discriminator_loss(real_output, fake_output)
gradients_of_generator = gen_tape.gradient(gen_loss, generator.trainable_variables)
gradients_of_discriminator = disc_tape.gradient(disc_loss, discriminator.trainable_variables)
generator_optimizer.apply_gradients(zip(gradients_of_generator, generator.trainable_variables))
discriminator_optimizer.apply_gradients(zip(gradients_of_discriminator, discriminator.trainable_variables))
def train(dataset, epochs):
for epoch in range(epochs):
start = time.time()
for image_batch in dataset:
train_step(image_batch)
# 继续进行时为 GIF 生成图像
display.clear_output(wait=True)
generate_and_save_images(generator,
epoch + 1,
seed)
# 每 15 个 epoch 保存一次模型
if (epoch + 1) % 15 == 0:
checkpoint.save(file_prefix = checkpoint_prefix)
print ('Time for epoch {} is {} sec'.format(epoch + 1, time.time()-start))
# 最后一个 epoch 结束后生成图片
display.clear_output(wait=True)
generate_and_save_images(generator,
epochs,
seed)
```
**生成与保存图片**
```
def generate_and_save_images(model, epoch, test_input):
# 注意 training` 设定为 False
# 因此,所有层都在推理模式下运行(batchnorm)。
predictions = model(test_input, training=False)
fig = plt.figure(figsize=(4,4))
for i in range(predictions.shape[0]):
plt.subplot(4, 4, i+1)
plt.imshow(predictions[i, :, :, 0] * 127.5 + 127.5, cmap='gray')
plt.axis('off')
plt.savefig('image_at_epoch_{:04d}.png'.format(epoch))
plt.show()
```
## 训练模型
调用上面定义的 `train()` 方法来同时训练生成器和判别器。注意,训练 GANs 可能是棘手的。重要的是,生成器和判别器不能够互相压制对方(例如,他们以相似的学习率训练)。
在训练之初,生成的图片看起来像是随机噪声。随着训练过程的进行,生成的数字将越来越真实。在大概 50 个 epoch 之后,这些图片看起来像是 MNIST 数字。使用 Colab 中的默认设置可能需要大约 1 分钟每 epoch。
```
%%time
train(train_dataset, EPOCHS)
```
恢复最新的检查点。
```
checkpoint.restore(tf.train.latest_checkpoint(checkpoint_dir))
```
## 创建 GIF
```
# 使用 epoch 数生成单张图片
def display_image(epoch_no):
return PIL.Image.open('image_at_epoch_{:04d}.png'.format(epoch_no))
display_image(EPOCHS)
```
使用训练过程中生成的图片通过 `imageio` 生成动态 gif
```
anim_file = 'dcgan.gif'
with imageio.get_writer(anim_file, mode='I') as writer:
filenames = glob.glob('image*.png')
filenames = sorted(filenames)
last = -1
for i,filename in enumerate(filenames):
frame = 2*(i**0.5)
if round(frame) > round(last):
last = frame
else:
continue
image = imageio.imread(filename)
writer.append_data(image)
image = imageio.imread(filename)
writer.append_data(image)
import IPython
if IPython.version_info > (6,2,0,''):
display.Image(filename=anim_file)
```
如果您正在使用 Colab,您可以通过如下代码下载动画:
```
try:
from google.colab import files
except ImportError:
pass
else:
files.download(anim_file)
```
## 下一步
本教程展示了实现和训练 GAN 模型所需的全部必要代码。接下来,您可能想尝试其他数据集,例如大规模名人面部属性(CelebA)数据集 [在 Kaggle 上获取](https://www.kaggle.com/jessicali9530/celeba-dataset)。要了解更多关于 GANs 的信息,我们推荐参阅 [NIPS 2016 教程: 生成对抗网络](https://arxiv.org/abs/1701.00160)。
| github_jupyter |
## **PySpark Machine Learning**
###Pipeline Example ([Docs](https://spark.apache.org/docs/latest/ml-pipeline.html))
In machine learning, it's common to run a series of steps for data prep, cleansing, feature engineering, and then ultimately model training (among several other potential steps).
Spark ML Pipelines sequences these steps into an ordered array (or DAG). A Pipeline is specified as a sequence of stages, and each stage is either a **Transformer** or an **Estimator**.
It's often a best practice to save a model or a pipeline to disk for later use.
Below is an example Spark ML Pipeline that shows two Transformers (Tokenizer and HashingTF) and one Estimator (Logistic Regression).
<img src="https://spark.apache.org/docs/latest/img/ml-Pipeline.png">
## **Install Spark Dependencies**
```
# Install Spark dependencies
!apt-get install openjdk-8-jdk-headless -qq > /dev/null
!rm spark-3.2.1-bin-hadoop3.2.tgz
!wget --no-cookies --no-check-certificate https://dlcdn.apache.org/spark/spark-3.2.1/spark-3.2.1-bin-hadoop3.2.tgz
!tar zxvf spark-3.2.1-bin-hadoop3.2.tgz
!pip install -q findspark
!pip install pyspark
!ls -al | grep spark
```
## **Import Python and PySpark Libraries**
```
# Set up required environment variables
import os
os.environ["JAVA_HOME"] = "/usr/lib/jvm/java-8-openjdk-amd64"
os.environ["SPARK_HOME"] = "/content/spark-3.2.1-bin-hadoop3.2"
from pyspark import SparkContext
from pyspark.sql import SparkSession
from pyspark.ml import Pipeline
from pyspark.ml.classification import LogisticRegression
from pyspark.ml.feature import HashingTF, Tokenizer
```
## **Initialize Spark Session**
```
spark = SparkSession.builder.appName("Spark ML Pipeline Example").master("local[*]").getOrCreate()
```
## **Load Sample Data**
```
training = spark.createDataFrame([
(0, "a b c d e spark", 1.0),
(1, "b d", 0.0),
(2, "spark f g h", 1.0),
(3, "hadoop mapreduce", 0.0)
], ["id", "text", "label"])
training.show(10,False)
```
## **Configure Pipeline Objects**
Transforms (tokenizer and hashingTF) and Estimators (logistic regression)
```
tokenizer = Tokenizer(inputCol="text", outputCol="words")
hashingTF = HashingTF(inputCol=tokenizer.getOutputCol(), outputCol="features")
lr = LogisticRegression(maxIter=10, regParam=0.001)
tokenizer.transform(training).show(5, False)
```
## **Create Pipeline Object**
```
pipeline = Pipeline(stages=[tokenizer, hashingTF, lr])
```
## **Run Pipeline to transform data and train model**
```
model = pipeline.fit(training)
dir(model.stages[-1].summary)
```
## **Test Model**
```
test = spark.createDataFrame([
(4, "spark i j k"),
(5, "l m n"),
(6, "spark hadoop spark"),
(7, "apache hadoop")
], ["id", "text"])
# Make predictions on test documents and print columns of interest.
prediction = model.transform(test)
prediction.show(10,False)
#selected = prediction.select("id", "text", "probability", "prediction")
#for row in selected.collect():
# rid, text, prob, prediction = row
# print("(%d, %s) --> prob=%s, prediction=%f" % (rid, text, str(prob), prediction))
```
| github_jupyter |
```
#12/29/20
#runnign synthetic benchmark graphs for synthetic OR datasets generated
#making benchmark images
import keras
from keras.models import Sequential, Model, load_model
from keras.layers import Dense, Dropout, Activation, Flatten, Input, Lambda
from keras.layers import Conv2D, MaxPooling2D, AveragePooling2D, Conv1D, MaxPooling1D, LSTM, ConvLSTM2D, GRU, CuDNNLSTM, CuDNNGRU, BatchNormalization, LocallyConnected2D, Permute, TimeDistributed, Bidirectional
from keras.layers import Concatenate, Reshape, Softmax, Conv2DTranspose, Embedding, Multiply
from keras.callbacks import ModelCheckpoint, EarlyStopping, Callback
from keras import regularizers
from keras import backend as K
from keras.utils.generic_utils import Progbar
from keras.layers.merge import _Merge
import keras.losses
from keras.datasets import mnist
from functools import partial
from collections import defaultdict
import tensorflow as tf
from tensorflow.python.framework import ops
import isolearn.keras as iso
import numpy as np
import tensorflow as tf
import logging
logging.getLogger('tensorflow').setLevel(logging.ERROR)
import os
import pickle
import numpy as np
import isolearn.io as isoio
import isolearn.keras as isol
import pandas as pd
import scipy.sparse as sp
import scipy.io as spio
import matplotlib.pyplot as plt
from sequence_logo_helper import dna_letter_at, plot_dna_logo
from keras.backend.tensorflow_backend import set_session
def contain_tf_gpu_mem_usage() :
config = tf.ConfigProto()
config.gpu_options.allow_growth = True
sess = tf.Session(config=config)
set_session(sess)
contain_tf_gpu_mem_usage()
class EpochVariableCallback(Callback) :
def __init__(self, my_variable, my_func) :
self.my_variable = my_variable
self.my_func = my_func
def on_epoch_begin(self, epoch, logs={}) :
K.set_value(self.my_variable, self.my_func(K.get_value(self.my_variable), epoch))
#ONLY RUN THIS CELL ONCE
from tensorflow.python.framework import ops
#Stochastic Binarized Neuron helper functions (Tensorflow)
#ST Estimator code adopted from https://r2rt.com/beyond-binary-ternary-and-one-hot-neurons.html
#See Github https://github.com/spitis/
def st_sampled_softmax(logits):
with ops.name_scope("STSampledSoftmax") as namescope :
nt_probs = tf.nn.softmax(logits)
onehot_dim = logits.get_shape().as_list()[1]
sampled_onehot = tf.one_hot(tf.squeeze(tf.multinomial(logits, 1), 1), onehot_dim, 1.0, 0.0)
with tf.get_default_graph().gradient_override_map({'Ceil': 'Identity', 'Mul': 'STMul'}):
return tf.ceil(sampled_onehot * nt_probs)
def st_hardmax_softmax(logits):
with ops.name_scope("STHardmaxSoftmax") as namescope :
nt_probs = tf.nn.softmax(logits)
onehot_dim = logits.get_shape().as_list()[1]
sampled_onehot = tf.one_hot(tf.argmax(nt_probs, 1), onehot_dim, 1.0, 0.0)
with tf.get_default_graph().gradient_override_map({'Ceil': 'Identity', 'Mul': 'STMul'}):
return tf.ceil(sampled_onehot * nt_probs)
@ops.RegisterGradient("STMul")
def st_mul(op, grad):
return [grad, grad]
#Gumbel Distribution Sampler
def gumbel_softmax(logits, temperature=0.5) :
gumbel_dist = tf.contrib.distributions.RelaxedOneHotCategorical(temperature, logits=logits)
batch_dim = logits.get_shape().as_list()[0]
onehot_dim = logits.get_shape().as_list()[1]
return gumbel_dist.sample()
#model functions for loading optimus scramblers
import keras.backend as K
def mask_dropout_multi_scale(mask, drop_scales=[1, 2, 4, 7], min_drop_rate=0.0, max_drop_rate=0.5) :
rates = K.random_uniform(shape=(K.shape(mask)[0], 1, 1, 1), minval=min_drop_rate, maxval=max_drop_rate)
scale_logits = K.random_uniform(shape=(K.shape(mask)[0], len(drop_scales), 1, 1, 1), minval=-5., maxval=5.)
scale_probs = K.softmax(scale_logits, axis=1)
ret_mask = mask
for drop_scale_ix, drop_scale in enumerate(drop_scales) :
ret_mask = mask_dropout(ret_mask, rates * scale_probs[:, drop_scale_ix, ...], drop_scale=drop_scale)
return K.switch(K.learning_phase(), ret_mask, mask)
def mask_dropout(mask, drop_rates, drop_scale=1) :
random_tensor_downsampled = K.random_uniform(shape=(
K.shape(mask)[0],
1,
K.cast(K.shape(mask)[2] / drop_scale, dtype=tf.int32),
K.shape(mask)[3]
), minval=0.0, maxval=1.0)
keep_mask_downsampled = random_tensor_downsampled >= drop_rates
keep_mask = K.repeat_elements(keep_mask_downsampled, rep=drop_scale, axis=2)
ret_mask = mask * K.cast(keep_mask, dtype=tf.float32)
return ret_mask
def mask_dropout_single_scale(mask, drop_scale=1, min_drop_rate=0.0, max_drop_rate=0.5) :
rates = K.random_uniform(shape=(K.shape(mask)[0], 1, 1, 1), minval=min_drop_rate, maxval=max_drop_rate)
random_tensor_downsampled = K.random_uniform(shape=(
K.shape(mask)[0],
1,
K.cast(K.shape(mask)[2] / drop_scale, dtype=tf.int32),
K.shape(mask)[3]
), minval=0.0, maxval=1.0)
keep_mask_downsampled = random_tensor_downsampled >= rates
keep_mask = K.repeat_elements(keep_mask_downsampled, rep=drop_scale, axis=2)
ret_mask = mask * K.cast(keep_mask, dtype=tf.float32)
return K.switch(K.learning_phase(), ret_mask, mask)
#PWM Masking and Sampling helper functions
def mask_pwm(inputs) :
pwm, onehot_template, onehot_mask = inputs
return pwm * onehot_mask + onehot_template
def sample_pwm_st(pwm_logits) :
n_sequences = K.shape(pwm_logits)[0]
seq_length = K.shape(pwm_logits)[2]
flat_pwm = K.reshape(pwm_logits, (n_sequences * seq_length, 4))
sampled_pwm = st_sampled_softmax(flat_pwm)
return K.reshape(sampled_pwm, (n_sequences, 1, seq_length, 4))
def sample_pwm_gumbel(pwm_logits) :
n_sequences = K.shape(pwm_logits)[0]
seq_length = K.shape(pwm_logits)[2]
flat_pwm = K.reshape(pwm_logits, (n_sequences * seq_length, 4))
sampled_pwm = gumbel_softmax(flat_pwm, temperature=0.5)
return K.reshape(sampled_pwm, (n_sequences, 1, seq_length, 4))
#Generator helper functions
def initialize_sequence_templates(generator, sequence_templates, background_matrices) :
embedding_templates = []
embedding_masks = []
embedding_backgrounds = []
for k in range(len(sequence_templates)) :
sequence_template = sequence_templates[k]
onehot_template = iso.OneHotEncoder(seq_length=len(sequence_template))(sequence_template).reshape((1, len(sequence_template), 4))
for j in range(len(sequence_template)) :
if sequence_template[j] not in ['N', 'X'] :
nt_ix = np.argmax(onehot_template[0, j, :])
onehot_template[:, j, :] = -4.0
onehot_template[:, j, nt_ix] = 10.0
elif sequence_template[j] == 'X' :
onehot_template[:, j, :] = -1.0
onehot_mask = np.zeros((1, len(sequence_template), 4))
for j in range(len(sequence_template)) :
if sequence_template[j] == 'N' :
onehot_mask[:, j, :] = 1.0
embedding_templates.append(onehot_template.reshape(1, -1))
embedding_masks.append(onehot_mask.reshape(1, -1))
embedding_backgrounds.append(background_matrices[k].reshape(1, -1))
embedding_templates = np.concatenate(embedding_templates, axis=0)
embedding_masks = np.concatenate(embedding_masks, axis=0)
embedding_backgrounds = np.concatenate(embedding_backgrounds, axis=0)
generator.get_layer('template_dense').set_weights([embedding_templates])
generator.get_layer('template_dense').trainable = False
generator.get_layer('mask_dense').set_weights([embedding_masks])
generator.get_layer('mask_dense').trainable = False
generator.get_layer('background_dense').set_weights([embedding_backgrounds])
generator.get_layer('background_dense').trainable = False
#Generator construction function
def build_sampler(batch_size, seq_length, n_classes=1, n_samples=1, sample_mode='st') :
#Initialize Reshape layer
reshape_layer = Reshape((1, seq_length, 4))
#Initialize background matrix
onehot_background_dense = Embedding(n_classes, seq_length * 4, embeddings_initializer='zeros', name='background_dense')
#Initialize template and mask matrices
onehot_template_dense = Embedding(n_classes, seq_length * 4, embeddings_initializer='zeros', name='template_dense')
onehot_mask_dense = Embedding(n_classes, seq_length * 4, embeddings_initializer='ones', name='mask_dense')
#Initialize Templating and Masking Lambda layer
masking_layer = Lambda(mask_pwm, output_shape = (1, seq_length, 4), name='masking_layer')
background_layer = Lambda(lambda x: x[0] + x[1], name='background_layer')
#Initialize PWM normalization layer
pwm_layer = Softmax(axis=-1, name='pwm')
#Initialize sampling layers
sample_func = None
if sample_mode == 'st' :
sample_func = sample_pwm_st
elif sample_mode == 'gumbel' :
sample_func = sample_pwm_gumbel
upsampling_layer = Lambda(lambda x: K.tile(x, [n_samples, 1, 1, 1]), name='upsampling_layer')
sampling_layer = Lambda(sample_func, name='pwm_sampler')
permute_layer = Lambda(lambda x: K.permute_dimensions(K.reshape(x, (n_samples, batch_size, 1, seq_length, 4)), (1, 0, 2, 3, 4)), name='permute_layer')
def _sampler_func(class_input, raw_logits) :
#Get Template and Mask
onehot_background = reshape_layer(onehot_background_dense(class_input))
onehot_template = reshape_layer(onehot_template_dense(class_input))
onehot_mask = reshape_layer(onehot_mask_dense(class_input))
#Add Template and Multiply Mask
pwm_logits = masking_layer([background_layer([raw_logits, onehot_background]), onehot_template, onehot_mask])
#Compute PWM (Nucleotide-wise Softmax)
pwm = pwm_layer(pwm_logits)
#Tile each PWM to sample from and create sample axis
pwm_logits_upsampled = upsampling_layer(pwm_logits)
sampled_pwm = sampling_layer(pwm_logits_upsampled)
sampled_pwm = permute_layer(sampled_pwm)
sampled_mask = permute_layer(upsampling_layer(onehot_mask))
return pwm_logits, pwm, sampled_pwm, onehot_mask, sampled_mask
return _sampler_func
#for formulation 2 graphing
def returnXMeanLogits(e_train):
#returns x mean logits for displayign the pwm difference for the version 2 networks
#Visualize background sequence distribution
seq_e_train = one_hot_encode(e_train,seq_len=50)
x_train = seq_e_train
x_train = np.reshape(x_train, (x_train.shape[0], 1, x_train.shape[1], x_train.shape[2]))
pseudo_count = 1.0
x_mean = (np.sum(x_train, axis=(0, 1)) + pseudo_count) / (x_train.shape[0] + 4. * pseudo_count)
x_mean_logits = np.log(x_mean / (1. - x_mean))
return x_mean_logits, x_mean
#loading testing dataset
from optimusFunctions import *
import pandas as pd
csv_to_open = "optimus5_synthetic_random_insert_if_uorf_2_start_2_stop_variable_loc_512.csv"
dataset_name = csv_to_open.replace(".csv", "")
print (dataset_name)
data_df = pd.read_csv("./" + csv_to_open) #open from scores folder
#loaded test set which is sorted by number of start/stop signals
seq_e_test = one_hot_encode(data_df, seq_len=50)
benchmarkSet_seqs = seq_e_test
x_test = np.reshape(benchmarkSet_seqs, (benchmarkSet_seqs.shape[0], 1, benchmarkSet_seqs.shape[1], benchmarkSet_seqs.shape[2]))
print (x_test.shape)
e_train = pd.read_csv("bottom5KIFuAUGTop5KIFuAUG.csv")
print ("training: ", e_train.shape[0], " testing: ", x_test.shape[0])
#one hot encode with optimus encoders
seq_e_train = one_hot_encode(e_train,seq_len=50)
x_mean_logits, x_mean = returnXMeanLogits(e_train)
seq_e_train = one_hot_encode(e_train,seq_len=50)
x_train = seq_e_train
x_train = np.reshape(x_train, (x_train.shape[0], 1, x_train.shape[1], x_train.shape[2]))
#background
#for formulation 2 graphing
def returnXMeanLogits(e_train):
#returns x mean logits for displayign the pwm difference for the version 2 networks
#Visualize background sequence distribution
seq_e_train = one_hot_encode(e_train,seq_len=50)
x_train = seq_e_train
x_train = np.reshape(x_train, (x_train.shape[0], 1, x_train.shape[1], x_train.shape[2]))
pseudo_count = 1.0
x_mean = (np.sum(x_train, axis=(0, 1)) + pseudo_count) / (x_train.shape[0] + 4. * pseudo_count)
x_mean_logits = np.log(x_mean / (1. - x_mean))
return x_mean_logits, x_mean
e_train = pd.read_csv("bottom5KIFuAUGTop5KIFuAUG.csv")
print ("training: ", e_train.shape[0], " testing: ", x_test.shape[0])
#one hot encode with optimus encoders
seq_e_train = one_hot_encode(e_train,seq_len=50)
x_mean_logits, x_mean = returnXMeanLogits(e_train)
x_train = seq_e_train
x_train = np.reshape(x_train, (x_train.shape[0], 1, x_train.shape[1], x_train.shape[2]))
#Define sequence template for optimus
sequence_template = 'N'*50
sequence_mask = np.array([1 if sequence_template[j] == 'N' else 0 for j in range(len(sequence_template))])
#Visualize background sequence distribution
save_figs = True
plot_dna_logo(np.copy(x_mean), sequence_template=sequence_template, figsize=(14, 0.65), logo_height=1.0, plot_start=0, plot_end=50)
#Calculate mean training set conservation
entropy = np.sum(x_mean * -np.log(x_mean), axis=-1) / np.log(2.0)
conservation = 2.0 - entropy
x_mean_conservation = np.sum(conservation) / np.sum(sequence_mask)
print("Mean conservation (bits) = " + str(x_mean_conservation))
#Calculate mean training set kl-divergence against background
x_train_clipped = np.clip(np.copy(x_train[:, 0, :, :]), 1e-8, 1. - 1e-8)
kl_divs = np.sum(x_train_clipped * np.log(x_train_clipped / np.tile(np.expand_dims(x_mean, axis=0), (x_train_clipped.shape[0], 1, 1))), axis=-1) / np.log(2.0)
x_mean_kl_divs = np.sum(kl_divs * sequence_mask, axis=-1) / np.sum(sequence_mask)
x_mean_kl_div = np.mean(x_mean_kl_divs)
print("Mean KL Div against background (bits) = " + str(x_mean_kl_div))
#Initialize Encoder and Decoder networks
batch_size = 32
seq_length = 50
n_samples = 128
sample_mode = 'st'
#sample_mode = 'gumbel'
#Load sampler
sampler = build_sampler(batch_size, seq_length, n_classes=1, n_samples=n_samples, sample_mode=sample_mode)
#Load Predictor
predictor_path = 'optimusRetrainedMain.hdf5'
predictor = load_model(predictor_path)
predictor.trainable = False
predictor.compile(optimizer=keras.optimizers.SGD(lr=0.1), loss='mean_squared_error')
#Build scrambler model
dummy_class = Input(shape=(1,), name='dummy_class')
input_logits = Input(shape=(1, seq_length, 4), name='input_logits')
pwm_logits, pwm, sampled_pwm, pwm_mask, sampled_mask = sampler(dummy_class, input_logits)
scrambler_model = Model([input_logits, dummy_class], [pwm_logits, pwm, sampled_pwm, pwm_mask, sampled_mask])
#Initialize Sequence Templates and Masks
initialize_sequence_templates(scrambler_model, [sequence_template], [x_mean_logits])
scrambler_model.trainable = False
scrambler_model.compile(
optimizer=keras.optimizers.Adam(lr=0.001, beta_1=0.9, beta_2=0.999),
loss='mean_squared_error'
)
#open all score and reshape as needed
file_names = [
"l2x_" + dataset_name + "_importance_scores_test.npy",
"invase_" + dataset_name + "_conv_importance_scores_test.npy",
"l2x_" + dataset_name + "_full_data_importance_scores_test.npy",
"invase_" + dataset_name + "_conv_full_data_importance_scores_test.npy",
]
#deepexplain_optimus_utr_OR_logic_synth_1_start_2_stops_method_integrated_gradients_importance_scores_test.npy
model_names =[
"l2x",
"invase",
"l2x_full_data",
"invase_full_data",
]
model_importance_scores_test = [np.load("./" + file_name) for file_name in file_names]
for scores in model_importance_scores_test:
print (scores.shape)
for model_i in range(len(model_names)) :
if model_importance_scores_test[model_i].shape[-1] > 1 :
model_importance_scores_test[model_i] = np.sum(model_importance_scores_test[model_i], axis=-1, keepdims=True)
for scores in model_importance_scores_test:
print (scores.shape)
#reshape for mse script -> if not (3008, 1, 50, 1) make it that shape
idealShape = model_importance_scores_test[0].shape
print (idealShape)
for model_i in range(len(model_names)) :
if model_importance_scores_test[model_i].shape != idealShape:
model_importance_scores_test[model_i] = np.expand_dims(model_importance_scores_test[model_i], 1)
for scores in model_importance_scores_test:
print (scores.shape)
on_state_logit_val = 50.
print (x_test.shape)
dummy_test = np.zeros((x_test.shape[0], 1))
x_test_logits = 2. * x_test - 1.
print (x_test_logits.shape)
print (dummy_test.shape)
x_test_squeezed = np.squeeze(x_test)
y_pred_ref = predictor.predict([x_test_squeezed], batch_size=32, verbose=True)[0]
_, _, _, pwm_mask, sampled_mask = scrambler_model.predict([x_test_logits, dummy_test], batch_size=batch_size)
feature_quantiles = [0.76, 0.82, 0.88]
for name in model_names:
for quantile in feature_quantiles:
totalName = name + "_" + str(quantile).replace(".","_") + "_quantile_MSE"
data_df[totalName] = None
print (data_df.columns)
feature_quantiles = [0.76, 0.82, 0.88]
#batch_size = 128
from sklearn import metrics
model_mses = []
for model_i in range(len(model_names)) :
print("Benchmarking model '" + str(model_names[model_i]) + "'...")
feature_quantile_mses = []
for feature_quantile_i, feature_quantile in enumerate(feature_quantiles) :
print("Feature quantile = " + str(feature_quantile))
if len(model_importance_scores_test[model_i].shape) >= 5 :
importance_scores_test = np.abs(model_importance_scores_test[model_i][feature_quantile_i, ...])
else :
importance_scores_test = np.abs(model_importance_scores_test[model_i])
n_to_test = importance_scores_test.shape[0] // batch_size * batch_size
importance_scores_test = importance_scores_test[:n_to_test]
importance_scores_test *= np.expand_dims(np.max(pwm_mask[:n_to_test], axis=-1), axis=-1)
quantile_vals = np.quantile(importance_scores_test, axis=(1, 2, 3), q=feature_quantile, keepdims=True)
quantile_vals = np.tile(quantile_vals, (1, importance_scores_test.shape[1], importance_scores_test.shape[2], importance_scores_test.shape[3]))
top_logits_test = np.zeros(importance_scores_test.shape)
top_logits_test[importance_scores_test > quantile_vals] = on_state_logit_val
top_logits_test = np.tile(top_logits_test, (1, 1, 1, 4)) * x_test_logits[:n_to_test]
_, _, samples_test, _, _ = scrambler_model.predict([top_logits_test, dummy_test[:n_to_test]], batch_size=batch_size)
print (samples_test.shape)
msesPerPoint = []
for data_ix in range(samples_test.shape[0]) :
#for each sample, look at kl divergence for the 128 size batch generated
#for MSE, just track the pred vs original pred
if data_ix % 1000 == 0 :
print("Processing example " + str(data_ix) + "...")
#from optimus R^2, MSE, Pearson R script
justPred = np.expand_dims(np.expand_dims(x_test[data_ix, 0, :, :], axis=0), axis=-1)
justPredReshape = np.reshape(justPred, (1,50,4))
expanded = np.expand_dims(samples_test[data_ix, :, 0, :, :], axis=-1) #batch size is 128
expandedReshape = np.reshape(expanded, (n_samples, 50,4))
y_test_hat_ref = predictor.predict(x=justPredReshape, batch_size=1)[0][0]
y_test_hat = predictor.predict(x=[expandedReshape], batch_size=32)
pwmGenerated = y_test_hat.tolist()
tempOriginals = [y_test_hat_ref]*y_test_hat.shape[0]
asArrayOrig = np.array(tempOriginals)
asArrayGen = np.array(pwmGenerated)
squeezed = np.squeeze(asArrayGen)
mse = metrics.mean_squared_error(asArrayOrig, squeezed)
#msesPerPoint.append(mse)
totalName = model_names[model_i] + "_" + str(feature_quantile).replace(".","_") + "_quantile_MSE"
data_df.at[data_ix, totalName] = mse
msesPerPoint.append(mse)
msesPerPoint = np.array(msesPerPoint)
feature_quantile_mses.append(msesPerPoint)
model_mses.append(feature_quantile_mses)
#Store benchmark results as tables
save_figs = False
mse_table = np.zeros((len(model_mses), len(model_mses[0])))
for i, model_name in enumerate(model_names) :
for j, feature_quantile in enumerate(feature_quantiles) :
mse_table[i, j] = np.mean(model_mses[i][j])
#Plot and store mse table
f = plt.figure(figsize = (4, 6))
cells = np.round(mse_table, 3).tolist()
print("--- MSEs ---")
max_len = np.max([len(model_name.upper().replace("\n", " ")) for model_name in model_names])
print(("-" * max_len) + " " + " ".join([(str(feature_quantile) + "0")[:4] for feature_quantile in feature_quantiles]))
for i in range(len(cells)) :
curr_len = len([model_name.upper().replace("\n", " ") for model_name in model_names][i])
row_str = [model_name.upper().replace("\n", " ") for model_name in model_names][i] + (" " * (max_len - curr_len))
for j in range(len(cells[i])) :
cells[i][j] = (str(cells[i][j]) + "00000")[:4]
row_str += " " + cells[i][j]
print(row_str)
print("")
table = plt.table(cellText=cells, rowLabels=[model_name.upper().replace("\n", " ") for model_name in model_names], colLabels=feature_quantiles, loc='center')
ax = plt.gca()
#f.patch.set_visible(False)
ax.axis('off')
ax.axis('tight')
plt.tight_layout()
if save_figs :
plt.savefig(dataset_name + "_l2x_and_invase_full_data" + "_mse_table.png", dpi=300, transparent=True)
plt.savefig(dataset_name + "_l2x_and_invase_full_data" + "_mse_table.eps")
plt.show()
```
| github_jupyter |
```
import os
import pyaudio
import wave
from scipy.io import wavfile
import time
CHUNK = 1024
FORMAT = pyaudio.paInt16
CHANNELS = 1
RATE = 16000
RECORD_SECONDS = 3
max_simvol=75
# WAVE_OUTPUT_FILENAME = 'test.wav'
voxforge_dir_my = './Voxforge/my_zapis'
# generate folder for txt files
if not os.path.isdir(os.path.join(voxforge_dir_my, 'txt')):
os.makedirs(os.path.join(voxforge_dir_my, 'txt'))
if not os.path.isdir(os.path.join(voxforge_dir_my, 'wav')):
os.makedirs(os.path.join(voxforge_dir_my, 'wav'))
while True:
ticks = str(int(time.time()))
while True:
some_var = raw_input("Введите текст на русском языке: ")
original = ' '.join(unicode(some_var,'utf-8').strip().lower().split(' ')).replace('.', '').replace("'", '').replace('-', '').replace(',','')
i1=original.encode("UTF-8")
if (len(i1)/2 < max_simvol) and (len(i1) >2) :
some_var1=i1[:max_simvol*2]
print ','.join([some_var1])
break
else:
prpp='В текст нет или больше букв, чем '
print ','.join([prpp]),max_simvol
some_var = raw_input("Если готовы к записи с микрофона нажмите ВВОД: ")
# write to file
with open(os.path.join(voxforge_dir_my, 'txt', ticks + '.txt'), 'w') as w:
w.write(some_var1)
p = pyaudio.PyAudio()
stream = p.open(format=FORMAT,
channels=CHANNELS,
rate=RATE,
input=True,
frames_per_buffer=CHUNK)
print("* СКАЖИТЕ В МИКРОФОН")
print(some_var1)
frames = []
for i in range(0, int(RATE / CHUNK * RECORD_SECONDS)):
data = stream.read(CHUNK)
frames.append(data)
print("* КОНЕЦ ЗАПИСИ")
stream.stop_stream()
stream.close()
p.terminate()
wf = wave.open(os.path.join(voxforge_dir_my, 'wav', ticks + '.wav'), 'wb')
wf.setnchannels(CHANNELS)
wf.setsampwidth(p.get_sample_size(FORMAT))
wf.setframerate(RATE)
wf.writeframes(b''.join(frames))
wf.close()
# Play audio
wf = wave.open(os.path.join(voxforge_dir_my, 'wav', ticks + '.wav'), 'rb')
p = pyaudio.PyAudio()
stream = p.open(format=p.get_format_from_width(wf.getsampwidth()),
channels=wf.getnchannels(),
rate=wf.getframerate(),
output=True)
data = wf.readframes(CHUNK)
while data != '':
stream.write(data)
data = wf.readframes(CHUNK)
stream.stop_stream()
stream.close()
p.terminate()
```
| github_jupyter |
# Array creation routines
## Ones and zeros
```
import numpy as np
```
Create a new array of 2*2 integers, without initializing entries.
```
np.empty([2,2], int)
```
Let X = np.array([1,2,3], [4,5,6], np.int32).
Create a new array with the same shape and type as X.
```
X = np.array([[1,2,3], [4,5,6]], np.int32)
np.empty_like(X)
```
Create a 3-D array with ones on the diagonal and zeros elsewhere.
```
np.eye(3)
np.identity(3)
```
Create a new array of 3*2 float numbers, filled with ones.
```
np.ones([3,2], float)
```
Let x = np.arange(4, dtype=np.int64). Create an array of ones with the same shape and type as X.
```
x = np.arange(4, dtype=np.int64)
np.ones_like(x)
```
Create a new array of 3*2 float numbers, filled with zeros.
```
np.zeros((3,2), float)
```
Let x = np.arange(4, dtype=np.int64). Create an array of zeros with the same shape and type as X.
```
x = np.arange(4, dtype=np.int64)
np.zeros_like(x)
```
Create a new array of 2*5 uints, filled with 6.
```
np.full((2, 5), 6, dtype=np.uint)
np.ones([2, 5], dtype=np.uint) * 6
```
Let x = np.arange(4, dtype=np.int64). Create an array of 6's with the same shape and type as X.
```
x = np.arange(4, dtype=np.int64)
np.full_like(x, 6)
np.ones_like(x) * 6
```
## From existing data
Create an array of [1, 2, 3].
```
np.array([1, 2, 3])
```
Let x = [1, 2]. Convert it into an array.
```
x = [1,2]
np.asarray(x)
```
Let X = np.array([[1, 2], [3, 4]]). Convert it into a matrix.
```
X = np.array([[1, 2], [3, 4]])
np.asmatrix(X)
```
Let x = [1, 2]. Conver it into an array of `float`.
```
x = [1, 2]
np.asfarray(x)
np.asarray(x, float)
```
Let x = np.array([30]). Convert it into scalar of its single element, i.e. 30.
```
x = np.array([30])
np.asscalar(x)
x[0]
```
Let x = np.array([1, 2, 3]). Create a array copy of x, which has a different id from x.
```
x = np.array([1, 2, 3])
y = np.copy(x)
print id(x), x
print id(y), y
```
## Numerical ranges
Create an array of 2, 4, 6, 8, ..., 100.
```
np.arange(2, 101, 2)
```
Create a 1-D array of 50 evenly spaced elements between 3. and 10., inclusive.
```
np.linspace(3., 10, 50)
```
Create a 1-D array of 50 element spaced evenly on a log scale between 3. and 10., exclusive.
```
np.logspace(3., 10., 50, endpoint=False)
```
## Building matrices
Let X = np.array([[ 0, 1, 2, 3],
[ 4, 5, 6, 7],
[ 8, 9, 10, 11]]).
Get the diagonal of X, that is, [0, 5, 10].
```
X = np.array([[ 0, 1, 2, 3], [ 4, 5, 6, 7], [ 8, 9, 10, 11]])
np.diag(X)
X.diagonal()
```
Create a 2-D array whose diagonal equals [1, 2, 3, 4] and 0's elsewhere.
```
np.diagflat([1, 2, 3, 4])
```
Create an array which looks like below.
array([[ 0., 0., 0., 0., 0.],
[ 1., 0., 0., 0., 0.],
[ 1., 1., 0., 0., 0.]])
```
np.tri(3, 5, -1)
```
Create an array which looks like below.
array([[ 0, 0, 0],
[ 4, 0, 0],
[ 7, 8, 0],
[10, 11, 12]])
```
np.tril(np.arange(1, 13).reshape(4, 3), -1)
```
Create an array which looks like below. array([[ 1, 2, 3],
[ 4, 5, 6],
[ 0, 8, 9],
[ 0, 0, 12]])
```
np.triu(np.arange(1, 13).reshape(4, 3), -1)
```
| github_jupyter |
# Modeling and Simulation in Python
Chapter 7
Copyright 2017 Allen Downey
License: [Creative Commons Attribution 4.0 International](https://creativecommons.org/licenses/by/4.0)
```
# Configure Jupyter so figures appear in the notebook
%matplotlib inline
# Configure Jupyter to display the assigned value after an assignment
%config InteractiveShell.ast_node_interactivity='last_expr_or_assign'
# import functions from the modsim.py module
from modsim import *
from pandas import read_html
```
### Code from the previous chapter
```
filename = 'data/World_population_estimates.html'
tables = read_html(filename, header=0, index_col=0, decimal='M')
table2 = tables[2]
table2.columns = ['census', 'prb', 'un', 'maddison',
'hyde', 'tanton', 'biraben', 'mj',
'thomlinson', 'durand', 'clark']
un = table2.un / 1e9
un.head()
census = table2.census / 1e9
census.head()
def plot_results(census, un, timeseries, title):
"""Plot the estimates and the model.
census: TimeSeries of population estimates
un: TimeSeries of population estimates
timeseries: TimeSeries of simulation results
title: string
"""
plot(census, ':', label='US Census')
plot(un, '--', label='UN DESA')
plot(timeseries, color='gray', label='model')
decorate(xlabel='Year',
ylabel='World population (billion)',
title=title)
def run_simulation(system, update_func):
"""Simulate the system using any update function.
system: System object
update_func: function that computes the population next year
returns: TimeSeries
"""
results = TimeSeries()
results[system.t_0] = system.p_0
for t in linrange(system.t_0, system.t_end):
results[t+1] = update_func(results[t], t, system)
return results
```
### Quadratic growth
Here's the implementation of the quadratic growth model.
```
def update_func_quad(pop, t, system):
"""Compute the population next year with a quadratic model.
pop: current population
t: current year
system: system object containing parameters of the model
returns: population next year
"""
net_growth = system.alpha * pop + system.beta * pop**2
return pop + net_growth
```
Here's a `System` object with the parameters `alpha` and `beta`:
```
t_0 = get_first_label(census)
t_end = get_last_label(census)
p_0 = census[t_0]
system = System(t_0=t_0,
t_end=t_end,
p_0=p_0,
alpha=0.025,
beta=-0.0018)
```
And here are the results.
```
results = run_simulation(system, update_func_quad)
plot_results(census, un, results, 'Quadratic model')
savefig('figs/chap07-fig01.pdf')
```
**Exercise:** Can you find values for the parameters that make the model fit better?
### Equilibrium
To understand the quadratic model better, let's plot net growth as a function of population.
```
pop_array = linspace(0, 15, 100)
net_growth_array = system.alpha * pop_array + system.beta * pop_array**2
None
```
Here's what it looks like.
```
sns.set_style('whitegrid')
plot(pop_array, net_growth_array)
decorate(xlabel='Population (billions)',
ylabel='Net growth (billions)')
sns.set_style('white')
savefig('figs/chap07-fig02.pdf')
```
Here's what it looks like. Remember that the x axis is population now, not time.
It looks like the growth rate passes through 0 when the population is a little less than 14 billion.
In the book we found that the net growth is 0 when the population is $-\alpha/\beta$:
```
-system.alpha / system.beta
```
This is the equilibrium the population tends toward.
`sns` is a library called Seaborn which provides functions that control the appearance of plots. In this case I want a grid to make it easier to estimate the population where the growth rate crosses through 0.
### Dysfunctions
When people first learn about functions, there are a few things they often find confusing. In this section I present and explain some common problems with functions.
As an example, suppose you want a function that takes a `System` object, with variables `alpha` and `beta`, as a parameter and computes the carrying capacity, `-alpha/beta`. Here's a good solution:
```
def carrying_capacity(system):
K = -system.alpha / system.beta
return K
sys1 = System(alpha=0.025, beta=-0.0018)
pop = carrying_capacity(sys1)
print(pop)
```
Now let's see all the ways that can go wrong.
**Dysfunction #1:** Not using parameters. In the following version, the function doesn't take any parameters; when `sys1` appears inside the function, it refers to the object we created outside the function.
```
def carrying_capacity():
K = -sys1.alpha / sys1.beta
return K
sys1 = System(alpha=0.025, beta=-0.0018)
pop = carrying_capacity()
print(pop)
```
This version actually works, but it is not as versatile as it could be. If there are several `System` objects, this function can only work with one of them, and only if it is named `system`.
**Dysfunction #2:** Clobbering the parameters. When people first learn about parameters, they often write functions like this:
```
def carrying_capacity(system):
system = System(alpha=0.025, beta=-0.0018)
K = -system.alpha / system.beta
return K
sys1 = System(alpha=0.025, beta=-0.0018)
pop = carrying_capacity(sys1)
print(pop)
```
In this example, we have a `System` object named `sys1` that gets passed as an argument to `carrying_capacity`. But when the function runs, it ignores the argument and immediately replaces it with a new `System` object. As a result, this function always returns the same value, no matter what argument is passed.
When you write a function, you generally don't know what the values of the parameters will be. Your job is to write a function that works for any valid values. If you assign your own values to the parameters, you defeat the whole purpose of functions.
**Dysfunction #3:** No return value. Here's a version that computes the value of `K` but doesn't return it.
```
def carrying_capacity(system):
K = -system.alpha / system.beta
sys1 = System(alpha=0.025, beta=-0.0018)
pop = carrying_capacity(sys1)
print(pop)
```
A function that doesn't have a return statement always returns a special value called `None`, so in this example the value of `pop` is `None`. If you are debugging a program and find that the value of a variable is `None` when it shouldn't be, a function without a return statement is a likely cause.
**Dysfunction #4:** Ignoring the return value. Finally, here's a version where the function is correct, but the way it's used is not.
```
def carrying_capacity(system):
K = -system.alpha / system.beta
return K
sys2 = System(alpha=0.025, beta=-0.0018)
carrying_capacity(sys2)
# print(K) This line won't work because K only exists inside the function.
```
In this example, `carrying_capacity` runs and returns `K`, but the return value is dropped.
When you call a function that returns a value, you should do something with the result. Often you assign it to a variable, as in the previous examples, but you can also use it as part of an expression.
For example, you could eliminate the temporary variable `pop` like this:
```
print(carrying_capacity(sys1))
```
Or if you had more than one system, you could compute the total carrying capacity like this:
```
total = carrying_capacity(sys1) + carrying_capacity(sys2)
total
```
## Exercises
**Exercise:** In the book, I present a different way to parameterize the quadratic model:
$ \Delta p = r p (1 - p / K) $
where $r=\alpha$ and $K=-\alpha/\beta$. Write a version of `update_func` that implements this version of the model. Test it by computing the values of `r` and `K` that correspond to `alpha=0.025, beta=-0.0018`, and confirm that you get the same results.
```
# Solution goes here
# Solution goes here
# Solution goes here
```
| github_jupyter |
```
import torch
from torch import nn
from torchvision import transforms
from torchvision.transforms.functional import pad
import numpy as np
import time
import numpy.testing as npt
from PIL import Image
from matplotlib import pyplot as plt
torch.manual_seed(1)
#### Load image and visualize its 3 channels ####
#### The image is available at the ./images folder within the repo ####
img = Image.open("flower128.jpg")
plt.figure()
plt.axis('off')
plt.imshow(img)
fig, axs = plt.subplots(nrows=1, ncols=3, figsize=(12,4))
for i, ax in enumerate(axs.flatten()):
plt.sca(ax)
plt.axis('off')
plt.imshow(np.array(img)[:, :, i], cmap='gray', vmin = 0, vmax = 255)
plt.show()
#### Setup conv layer parameters ####
w_in=img.size[0]
h_in=img.size[1]
c_in=3
c_out=5
k=3
stride=2
padding=1
# (n+2*pad-ks)//stride +1
w_out = (w_in+2*padding-k)//stride +1
h_out = (h_in+2*padding-k)//stride +1
#### Setup Pytorch Conv2d layer ####
data = transforms.ToTensor()(img).unsqueeze_(0)
def init_weights(m):
if isinstance(m, nn.Conv2d):
torch.nn.init.normal_(m.weight, 0.0, 0.02)
torch.nn.init.normal_(m.bias, 0.0, 0.02)
convlayer=nn.Conv2d(c_in, c_out, k, stride=stride, padding=padding)
convlayer=convlayer.apply(init_weights)
print(convlayer)
#### Setup Numpy data ####
img2= img.copy()
img2=pad(img2, (1,1))
data2 = np.array(img2).transpose((2, 0, 1))
data2 = np.expand_dims(data2, axis=0)
data2 = data2/255.0
print("pytorch input shape: ",data.shape)
print("numpy input shape: ",data2.shape)
print("output dimensions: ",w_out, h_out)
#### Setup Numpy weight and bias structures ####
print(convlayer.weight.shape)
print(convlayer.bias.shape)
w=np.zeros((c_out, c_in, k, k))
b=np.zeros((c_out))
#### Create 3 kernels ####
ker=[]
ker.append(np.array([[-1.1,-1.1,-1.1],[2.2,2.2,2.2],[-1.1,-1.1,-1.1]]))
ker.append(np.array([[1/9,1/9,1/9],[1/9,1/9,1/9],[1/9,1/9,1/9]]))
ker.append(np.array([[0.0,-1.0,0.0],[-1.0,6.0,-1.0],[0.0,-1.0,0.0]]))
#### Hack the pytorch weights ####
with torch.no_grad():
for w0 in range(convlayer.weight.shape[0]):
for w1 in range(convlayer.weight.shape[1]):
if (w0<len(ker)):
convlayer.weight[w0][w1]=torch.from_numpy(ker[w0])
#### Hack the numpy weights ####
for w0 in range(c_out):
for w1 in range(c_in):
if (w0<len(ker)):
w[w0][w1]=ker[w0]
else:
w[w0][w1]=np.around(convlayer.weight[w0][w1].detach().numpy(),decimals=4) #np.random.normal(0, 0.02)
b[w0]=np.around(convlayer.bias[w0].detach().numpy(),decimals=4)
#print(w)
print(convlayer.bias)
print(b)
#### Create the python convolutional layer ####
def python_conv(data):
res=np.zeros((1, c_out, w_out, h_out))
for o1 in range(w_out):
for o2 in range(h_out):
for co in range(c_out):
total=0
for ci in range(c_in):
kt=0
for k1 in range(k):
for k2 in range(k):
weight = w[co,ci,k1,k2]
pos1=k1+o1*stride
pos2=k2+o2*stride
value = data[0, ci, pos1 ,pos2]
kt+= weight * value
total+=kt
res[0,co,o1,o2]=total+b[co]
return res
#### Create the sigmoid function to normalize the output to the 0-1 range ####
def sigm(data):
z = 1/(1 + np.exp(-data))
return z
#### Run the pytorch conv layer ####
start=time.time()
ptorch_out = convlayer(data)
ptorch_out = nn.Sigmoid()(ptorch_out)
end=time.time()
print("conv2d completed")
print("Pytorch: time taken: ",end-start)
#### Run the python-numpy conv layer ####
start=time.time()
numpy_out = python_conv(data2)
numpy_out = sigm(numpy_out)
end=time.time()
print("Python: time taken: ",end-start)
#### Print and compare output shapes ####
print()
print("Pytorch in shape: ",data.shape)
print("Python in shape: ",data2.shape)
print("Pytorch out shape: ",ptorch_out.shape)
print("Python out shape: ",numpy_out.shape)
print("weight shape: ",convlayer.weight.shape)
print("bias shape: ",convlayer.bias.shape)
print()
#### Extract a fragment of the output to compare both outputs ####
pytorch_verify=ptorch_out[0][0][0].cpu().detach().numpy()
pytorch_verify=np.around(pytorch_verify,decimals=3)
numpy_verify=np.around(numpy_out[0][0][0],decimals=3)
print("pytorch: ",pytorch_verify[:25])
print("python: ",numpy_verify[:25])
assert np.allclose(pytorch_verify, numpy_verify)
%matplotlib inline
#### Visualize the 5 output channels of both pytorch and python-numpy versions ####
nc=5
plt.figure(figsize=(15, 15), dpi=80)
fig, axs = plt.subplots(nrows=1, ncols=nc, figsize=(12,4))
for i, ax in enumerate(axs.flatten()):
plt.sca(ax)
#plt.axis('off')
im=transforms.ToPILImage(mode='L')(ptorch_out[0][i])
plt.imshow(im, cmap='gray')
#plt.tight_layout()
plt.show()
fig, axs = plt.subplots(nrows=1, ncols=nc, figsize=(12,4))
for i, ax in enumerate(axs.flatten()):
plt.sca(ax)
#plt.axis('off')
im = Image.fromarray(numpy_out[0][i]*255)
im = im.convert("L")
plt.imshow(im, cmap='gray')
plt.show()
```
| github_jupyter |
# Inference and Validation
Now that you have a trained network, you can use it for making predictions. This is typically called **inference**, a term borrowed from statistics. However, neural networks have a tendency to perform *too well* on the training data and aren't able to generalize to data that hasn't been seen before. This is called **overfitting** and it impairs inference performance. To test for overfitting while training, we measure the performance on data not in the training set called the **validation** dataset. We avoid overfitting through regularization such as dropout while monitoring the validation performance during training. In this notebook, I'll show you how to do this in PyTorch.
First off, I'll implement my own feedforward network for the exercise you worked on in part 4 using the Fashion-MNIST dataset.
As usual, let's start by loading the dataset through torchvision. You'll learn more about torchvision and loading data in a later part.
```
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
import matplotlib.pyplot as plt
import numpy as np
import time
import torch
from torch import nn
from torch import optim
import torch.nn.functional as F
from torchvision import datasets, transforms
import helper
# Define a transform to normalize the data
transform = transforms.Compose([transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
# Download and load the training data
trainset = datasets.FashionMNIST('F_MNIST_data/', download=True, train=True, transform=transform)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=64, shuffle=True)
# Download and load the test data
testset = datasets.FashionMNIST('F_MNIST_data/', download=True, train=False, transform=transform)
testloader = torch.utils.data.DataLoader(testset, batch_size=64, shuffle=True)
```
## Building the network
As with MNIST, each image in Fashion-MNIST is 28x28 which is a total of 784 pixels, and there are 10 classes. I'm going to get a bit more advanced here, I want to be able to build a network with an arbitrary number of hidden layers. That is, I want to pass in a parameter like `hidden_layers = [512, 256, 128]` and the network is contructed with three hidden layers have 512, 256, and 128 units respectively. To do this, I'll use `nn.ModuleList` to allow for an arbitrary number of hidden layers. Using `nn.ModuleList` works pretty much the same as a normal Python list, except that it registers each hidden layer `Linear` module properly so the model is aware of the layers.
The issue here is I need a way to define each `nn.Linear` module with the appropriate layer sizes. Since each `nn.Linear` operation needs an input size and an output size, I need something that looks like this:
```python
# Create ModuleList and add input layer
hidden_layers = nn.ModuleList([nn.Linear(input_size, hidden_layers[0])])
# Add hidden layers to the ModuleList
hidden_layers.extend([nn.Linear(h1, h2) for h1, h2 in layer_sizes])
```
Getting these pairs of input and output sizes can be done with a handy trick using `zip`.
```python
hidden_layers = [512, 256, 128, 64]
layer_sizes = zip(hidden_layers[:-1], hidden_layers[1:])
for each in layer_sizes:
print(each)
>> (512, 256)
>> (256, 128)
>> (128, 64)
```
I also have the `forward` method returning the log-softmax for the output. Since softmax is a probability distibution over the classes, the log-softmax is a log probability which comes with a [lot of benefits](https://en.wikipedia.org/wiki/Log_probability). Using the log probability, computations are often faster and more accurate. To get the class probabilities later, I'll need to take the exponential (`torch.exp`) of the output. Algebra refresher... the exponential function is the inverse of the log function:
$$ \large{e^{\ln{x}} = x }$$
We can include dropout in our network with [`nn.Dropout`](http://pytorch.org/docs/master/nn.html#dropout). This works similar to other modules such as `nn.Linear`. It also takes the dropout probability as an input which we can pass as an input to the network.
```
class Network(nn.Module):
def __init__(self, input_size, output_size, hidden_layers, drop_p=0.5):
''' Builds a feedforward network with arbitrary hidden layers.
Arguments
---------
input_size: integer, size of the input
output_size: integer, size of the output layer
hidden_layers: list of integers, the sizes of the hidden layers
drop_p: float between 0 and 1, dropout probability
'''
super().__init__()
# Add the first layer, input to a hidden layer
self.hidden_layers = nn.ModuleList([nn.Linear(input_size, hidden_layers[0])])
# Add a variable number of more hidden layers
layer_sizes = zip(hidden_layers[:-1], hidden_layers[1:])
self.hidden_layers.extend([nn.Linear(h1, h2) for h1, h2 in layer_sizes])
self.output = nn.Linear(hidden_layers[-1], output_size)
self.dropout = nn.Dropout(p=drop_p)
def forward(self, x):
''' Forward pass through the network, returns the output logits '''
# Forward through each layer in `hidden_layers`, with ReLU activation and dropout
for linear in self.hidden_layers:
x = F.relu(linear(x))
x = self.dropout(x)
x = self.output(x)
return F.log_softmax(x, dim=1)
```
# Train the network
Since the model's forward method returns the log-softmax, I used the [negative log loss](http://pytorch.org/docs/master/nn.html#nllloss) as my criterion, `nn.NLLLoss()`. I also chose to use the [Adam optimizer](http://pytorch.org/docs/master/optim.html#torch.optim.Adam). This is a variant of stochastic gradient descent which includes momentum and in general trains faster than your basic SGD.
I've also included a block to measure the validation loss and accuracy. Since I'm using dropout in the network, I need to turn it off during inference. Otherwise, the network will appear to perform poorly because many of the connections are turned off. PyTorch allows you to set a model in "training" or "evaluation" modes with `model.train()` and `model.eval()`, respectively. In training mode, dropout is turned on, while in evaluation mode, dropout is turned off. This effects other modules as well that should be on during training but off during inference.
The validation code consists of a forward pass through the validation set (also split into batches). With the log-softmax output, I calculate the loss on the validation set, as well as the prediction accuracy.
```
# Create the network, define the criterion and optimizer
model = Network(784, 10, [516, 256], drop_p=0.5)
criterion = nn.NLLLoss()
optimizer = optim.Adam(model.parameters(), lr=0.001)
# Implement a function for the validation pass
def validation(model, testloader, criterion):
test_loss = 0
accuracy = 0
for images, labels in testloader:
images.resize_(images.shape[0], 784)
output = model.forward(images)
test_loss += criterion(output, labels).item()
ps = torch.exp(output)
equality = (labels.data == ps.max(dim=1)[1])
accuracy += equality.type(torch.FloatTensor).mean()
return test_loss, accuracy
epochs = 2
steps = 0
running_loss = 0
print_every = 40
for e in range(epochs):
model.train()
for images, labels in trainloader:
steps += 1
# Flatten images into a 784 long vector
images.resize_(images.size()[0], 784)
optimizer.zero_grad()
output = model.forward(images)
loss = criterion(output, labels)
loss.backward()
optimizer.step()
running_loss += loss.item()
if steps % print_every == 0:
# Make sure network is in eval mode for inference
model.eval()
# Turn off gradients for validation, saves memory and computations
with torch.no_grad():
test_loss, accuracy = validation(model, testloader, criterion)
print("Epoch: {}/{}.. ".format(e+1, epochs),
"Training Loss: {:.3f}.. ".format(running_loss/print_every),
"Test Loss: {:.3f}.. ".format(test_loss/len(testloader)),
"Test Accuracy: {:.3f}".format(accuracy/len(testloader)))
running_loss = 0
# Make sure training is back on
model.train()
```
## Inference
Now that the model is trained, we can use it for inference. We've done this before, but now we need to remember to set the model in inference mode with `model.eval()`. You'll also want to turn off autograd with the `torch.no_grad()` context.
```
# Test out your network!
model.eval()
dataiter = iter(testloader)
images, labels = dataiter.next()
img = images[0]
# Convert 2D image to 1D vector
img = img.view(1, 784)
# Calculate the class probabilities (softmax) for img
with torch.no_grad():
output = model.forward(img)
ps = torch.exp(output)
# Plot the image and probabilities
helper.view_classify(img.view(1, 28, 28), ps, version='Fashion')
```
## Next Up!
In the next part, I'll show you how to save your trained models. In general, you won't want to train a model everytime you need it. Instead, you'll train once, save it, then load the model when you want to train more or use if for inference.
| github_jupyter |
# 計算の原子
量子コンピューターのプログラミングは、今では誰もが自宅から快適におこなうことができます。
でも、何をどうやってつくるのでしょうか?そもそも量子プログラムとは?量子コンピューターとは何でしょうか?
これらの疑問は、今日の標準のデジタルコンピューターと比較することで回答できます。残念ながら、ほとんどの人は実際にデジタルコンピューターの動作を理解していません。この記事では、まずはデジタルコンピューターの基本原則から見ていきます。その後、量子コンピューティングの話にスムーズに移行できるよう、量子コンピューティング上での計算を行うときと同じツールを使用して実行します。
## 目次
1. [情報をビットに分割する](#bits)
2. [ダイアグラムを使った計算](#diagram)
3. [はじめての量子回路](#first-circuit)
4. [例:加算回路を作成する](#adder)
4.1 [入力をエンコードする](#encoding)
4.2 [足し算の仕方を思い出してみよう](#remembering-add)
4.3 [Qiskitで足し算を行う](#adding-qiskit)
以下は、このページのコードを使用するために必要なPythonのコードです。
```
from qiskit import QuantumCircuit, assemble, Aer
from qiskit.visualization import plot_histogram
```
### 1. 情報をビットに分割する
最初に知っておくべきことは、ビットの概念です。これらは、世界で最も単純なアルファベットになるように設計されています。 0と1の2文字だけで、あらゆる情報を表現できます。
例えば数字もそのひとつです。ヨーロッパ言語では、通常、数字は0、1、2、3、4、5、6、7、8、9の10桁の文字列を使用して表されます。この数列の各数字は、10のべき乗が含まれる回数を表します。たとえば、9213と書くと、
$$ 9000 + 200 + 10 + 3 $$
または、10の累乗を強調する方法で表現されます。
$$ (9\times10^3) + (2\times10^2) + (1\times10^1) + (3\times10^0) $$
通常、このシステムは数10に基づいて使用しますが、他の数に基づいたシステムも同様に簡単に使用できます。たとえば、2進数システムは2に基づいています。これは、2つの文字0と1を使用して、数値を2のべき乗の倍数として表現することを意味します。たとえば、9213は10001111111101になります。
$$ 9213 = (1 \times 2^{13}) + (0 \times 2^{12}) + (0 \times 2^{11})+ (0 \times 2^{10}) +(1 \times 2^9) + (1 \times 2^8) + (1 \times 2^7) \\\\ \,\,\, + (1 \times 2^6) + (1 \times 2^5) + (1 \times 2^4) + (1 \times 2^3) + (1 \times 2^2) + (0 \times 2^1) + (1 \times 2^0) $$
ここでは、10、100、1000などではなく、2、4、8、16、32などの倍数として数値を表現しています。
<a id="binary_widget"></a>
```
from qiskit_textbook.widgets import binary_widget
binary_widget(nbits=5)
```
バイナリ文字列として知られるこれらのビット文字列は、単なる数値以上のものを表すために使用できます。たとえば、ビットを使用してテキストを表現する方法があります。使用する任意の文字、数字、または句読点について、[この表](https://www.ibm.com/support/knowledgecenter/en/ssw_aix_72/network/conversion_table.html)を使用して、最大8ビットの対応するストリングを検索できます。変換コードは任意に定義されたものですが、広く合意された標準になっており、この記事をインターネット経由で送信するのにも使われています。
このようにしてすべての情報はコンピューターで表現されています。数字、文字、画像、音声のいずれであっても、すべてバイナリ文字列の形で存在します。
私たちの標準的なデジタルコンピューターのように、量子コンピューターはこの同じ基本的な考え方に基づいています。主な違いは、量子的に操作できるビットの変形である qubits を使用していることです。この教科書の残りの部分では、量子ビットとは何か、それらが何ができるか、どのようにそれを行うかを探ります。ただし、このセクションでは、量子についてはまったく話しません。そのため、量子ビットを通常のビットとして使用します。
### 練習問題
1. 好きな数字を考えて、2進法で書いてみてください。
2. $n$個のビットがあるとしたら、それは何種類の状態になるでしょうか?
### 2. ダイアグラムを使った計算
量子ビットを使用する場合もビットを使用する場合も、入力を必要な出力に変換するためにそれらを操作する必要があります。少数のビット用の非常に単純なプログラムでは、このプロセスを「回路図」と呼ばれる図で表すと便利です。入力は左側にあり、出力は右側にあり、演算はその間の不可解な記号によって表されます。これらの演算に用いられる記号は、主に歴史的な背景から「ゲート」と呼ばれます。
標準的なビットベースのコンピューターでの回路の例を次に示します。これが具体的に何を意味するかまではわからなくても、これらの回路がどのようなものか、イメージだけをもっていただければ大丈夫です。

量子コンピューターも基本的には同じ考え方ですが、入力、出力、および演算に使用する記号は異なる表現を使います。上記のビットベースのコンピューターで示した例と同じプロセスを表す量子回路を次に示します。

このセクションの残りの部分では、回路の構築方法を説明します。最後に、上記の回路を作成する方法、それが何をするのか、なぜ有用なのかを理解します。
### 3. はじめての量子回路
量子回路を作成するには3つのステップが必要です。最初に入力をエンコードし、次に実際の計算を行い、最後に出力を抽出します。最初の量子回路では、これらの最後の仕事に焦点を当てます。まず、8つの量子ビットと8つの出力を持つ回路を作成します。
```
n = 8
n_q = n
n_b = n
qc_output = QuantumCircuit(n_q,n_b)
```
`qc_output` と呼ばれるこの回路は、`QuantumCircuit`を使用してQiskitによって作成されます。数値`n_q`は、回路内の量子ビットの数を定義します。 `n_b`を使用して、最後に回路から抽出する出力ビット数を定義します。
量子回路の出力の抽出は、「測定」と呼ばれる操作を使用して行われます。各測定は、特定の量子ビットに対して特定の出力ビットを出力として与えるように伝えます。次のコードは、8つの量子ビットのそれぞれに「測定」操作を適用します。量子ビットとビットは両方とも0から7までの数字でラベル付けされています(プログラマーがそうするのが好きなため)。コマンド`qc.measure(j,j)`は、回路`qc`に測定値を追加し、量子ビット`j`に出力をビット`j`に書き込むよう指示します。
```
for j in range(n):
qc_output.measure(j,j)
```
それでは作成した回路の中身をみてみましょう。
```
qc_output.draw()
```
量子ビットは常に出力が ```0```となるよう初期化されます。上記の回路は量子ビットに対して何の操作もしませんので、これらを測定したときに得られる結果もまた```0```です。これは回路を何度も実行し、結果をヒストグラムにプロットすることで確認できます。各量子ビットからの結果は、0番目から各量子ビットにわたって常に00000000であることがわかります。
```
sim = Aer.get_backend('aer_simulator') # this is the simulator we'll use
qobj = assemble(qc_output) # this turns the circuit into an object our backend can run
result = sim.run(qobj).result() # we run the experiment and get the result from that experiment
# from the results, we get a dictionary containing the number of times (counts)
# each result appeared
counts = result.get_counts()
# and display it on a histogram
plot_histogram(counts)
```
何回も実行して結果をヒストグラムとして表示する理由は、量子コンピューターの計算結果にランダム性が存在するためです。この場合は、何も量子的な計算を行っていないため、`00000000` の結果を確実に出力します。
この結果は、量子コンピューターの振る舞いを再現する古典コンピューターである量子シミュレーターから得られていることに注意してください。シミュレーションは少数の量子ビットに対してのみ可能ですが、それでも最初の量子回路を設計する際には非常に便利なツールです。実際の量子デバイスで実行するには `Aer.get_backend('qasm_simulator')` を使用したいデバイスのバックエンドオブジェクトに置き換えるだけです。
### 4. 例:加算回路を作成する
#### 4.1 入力をエンコードする
次に、異なるバイナリ文字列を入力としてエンコードする方法を見てみましょう。このためには、NOTゲートと呼ばれるものが必要です。これは、コンピューターで実行できる最も基本的な操作です。ビット値を単純に反転します: ```0```は```1```になり、```1```は```0```になります。量子ビットの場合、NOTの仕事をする ```x```と呼ばれる操作です。
以下では、エンコードのジョブ専用の新しい回路を作成し、`qc_encode`と呼びます。ここでは量子ビットの数のみを指定します。
```
qc_encode = QuantumCircuit(n)
qc_encode.x(7)
qc_encode.draw()
```
結果の抽出は、以前つくった`qc_output`回路を使用して実行できます。 `qc_encode + qc_output`で2つの回路をつなぎあわせれば、最後に追加された出力を抽出することができます。
```
qc = qc_encode + qc_output
qc.draw()
```
こうしてふたつの回路を組み合わせて実行して結果を表示させることができます。
```
qobj = assemble(qc)
counts = sim.run(qobj).result().get_counts()
plot_histogram(counts)
```
ご覧のとおり、今回は ```10000000``` という結果が出力されました。
反転したビットは、量子ビット7に由来し、文字列の左端にあります。これは、Qiskitが文字列のビットに右から左に番号を付けるためです。この慣習に違和感を覚えたなら、心配しないでください。文字列のビットにどういう順番で番号を付けるかは好みが分かれるところで、他の教科書だと違う順番に番号を付けている場合もあるでしょう。Qiskitでは文字列のビットに右から左に番号を付けるということだけ覚えておいてください。いずれにせよ、量子ビットを使って数字を表す場合、Qiskitの方法には利点があります。なぜなら、私たちが慣れ親しんでいるバイナリ表現と同様、qubit 7は、数に$2^7$がいくつあるかを示してくれるからです。つまり、qubit 7の反転(0→1)操作は、8ビットコンピューター上で128という数字をエンコードするのと同じことなのです。
次に、何か他の数字をエンコードしてみましょう。たとえば、あなたの年齢を表示させます。検索エンジンを使用して、年齢の数字がバイナリでどのように表現されるのかを調べて(「0b」が含まれている場合は無視して)64歳未満の場合は左側に0を追加してエンコードします。
```
qc_encode = QuantumCircuit(n)
qc_encode.x(1)
qc_encode.x(5)
qc_encode.draw()
```
これでコンピューターへの情報のエンコードの仕方がわかりました。次はエンコードした情報の処理方法、エンコードした入力からのアウトプットの出力の仕方についてです。
#### 4.2 足し算の仕方を思い出してみよう
入力から出力を得るには、解く問題が必要です。基本的な算数からおさらいをしましょう。小学校で、比較的大きな数を足し算をするときは、扱いやすい大きさに分解しました。例えば:
```
9213
+ 1854
= ????
```
ここでは 一桁ずつ右から順番に足していきます。まずは3+4
```
9213
+ 1854
= ???7
```
そして 1+5
```
9213
+ 1854
= ??67
```
つづいて 2+8=10 結果が2桁となったため、1を保持して次の桁に繰り上げを行う必要があります。
```
9213
+ 1854
= ?067
¹
```
そして最後に 9+1+1=11を計算することで答えを求めることができます。
```
9213
+ 1854
= 11067
¹
```
これは単純な足し算の例かもしれませんが、すべてのアルゴリズムの基本的な原理を示しています。アルゴリズムが数学的な問題を解決するように設計されているか、テキストや画像を処理するように設計されているかに関係なく、私たちは常に大きなタスクを小さくシンプルなステップに分解します。
コンピューターで実行するには、アルゴリズムを可能な限り小さくて最も簡単な手順にコンパイルする必要があります。これらがどのように見えるかを見るために、上記の足し算の問題をもう一度、ただし今度はバイナリでやってみましょう。
```
10001111111101
+ 00011100111110
= ??????????????
```
2番目の文字列の左端に余分な0がたくさんついていることに注目してください。これは、2つの文字列の長さを揃えるためです。
最初のタスクは、右端の1 + 0を実行することです。バイナリでは、任意の数体系と同様に、答えは1です。2番目の列の0 + 1に対しても同じ結果が得られます。
```
10001111111101
+ 00011100111110
= ????????????11
```
次に1+1=2ですが、 バイナリでは、数値2は ```10```と記述されるため、2ビット必要です。つまり、10進数の10の場合と同様に、1を保持する必要があります。
```
10001111111101
+ 00011100111110
= ???????????011
¹
```
次の桁では```1+1+1```を計算します. 三個の数を合計する必要があるのでコンピューターにとってはやや複雑な処理になってきました。しかしこれも毎回二個のビットの足し算という、より単純なオペレーションにコンパイルすることができます。 まずは最初の二つの1から足していきます。
```
1
+ 1
= 10
```
つづいて、この ```10```と最後の```1```を足す必要があります。これは、いつも通りそれぞれ同じ長さの文字列として並べて上下を足すことで実行できます。
```
10
+ 01
= 11
```
答えは ```11```(3としても知られています)です。
ここで残りの問題に戻ることができます。 上記の答え```11```には、新たなキャリービットがあります。
```
10001111111101
+ 00011100111110
= ??????????1011
¹¹
```
ここで再び1 + 1 + 1を行う必要がでてきました。しかし、私たちはすでにこの問題の解き方を知っていますので、大したことではありません。
実はここから先の問題の解き方を我々はすでに知っています。残っている桁の計算を二つのビットの足し算に分解すると、計算が必要な組合せは四つしかないためです。 四つの計算結果を次に示します(一貫性を保つために、すべての回答を2ビットで記述します)。
```
0+0 = 00 (in decimal, this is 0+0=0)
0+1 = 01 (in decimal, this is 0+1=1)
1+0 = 01 (in decimal, this is 1+0=1)
1+1 = 10 (in decimal, this is 1+1=2)
```
これは半加算器(*half adder*)と呼ばれています。私たちが量子ビットでつくるコンピューターも、この半加算器さえ実装できれば、それらをつなぎ合わせることでどのような数でも足すことができるようになるのです。
#### 4.3 Qiskitで足し算を行う
それでは、Qiskitを使って独自の半加算器を作成してみましょう。この回路には、入力をエンコードする部分、アルゴリズムを実行する部分、および結果を抽出する部分が含まれます。最初の入力をエンコードする部分は、新しい入力を使用するたびに変更する必要がありますが、残りは常に同じままです。

足し合わせる2つのビットは、量子ビット0(`q[0]`)と1(`q[1]`)にそれぞれエンコードされます。上記では、それぞれの量子ビットに ```1```がエンコードされ、```1 + 1```の解を見つけようとしています。結果は2ビットの文字列になり、量子ビット2(`q[2]`)および量子ビット3(`q[3]`)から読み取ります。 残されているのは中央の空白スペースに実際のプログラムを埋めることだけです。
画像の破線は、回路のさまざまな部分を区別するためのものです(ただし、より興味深い用途もあります)。これらは `barrier`コマンドを使用して作成されます。
コンピューティングの基本操作は、論理ゲートで行われます。これまでに回答を手動で書き出すためにNOTゲートを使いましたが、半加算器を作成するのにはこれだけでは十分ではありません。コンピューターに実際の計算を行わせるには、さらに強力なゲートが必要になります。
どのようなゲートが必要になってくるのかをみるために、半加算器がすべきことをもう一度見てみましょう。
```
0+0 = 00
0+1 = 01
1+0 = 01
1+1 = 10
```
これら4つの答えすべての右端のビットは、足し合わせる2つのビットが同じか異なるかによって決定されます。そのため、2つのビットが等しい「0 + 0」と「1 + 1」の場合、答えの右端のビットが「0」になります。異なるビットを足し合わせる「0 + 1」と「1 + 0」の場合、右端のビットは「1」です。
この論理を再現するには、2つのビットが異なるかどうかを判断できるものが必要です。従来、デジタルコンピューティングの研究では、これはXORゲートで実現できます。
量子コンピューティングにおいては, XORゲートの仕事は制御NOTゲートが行います。長い名前なので、通常は単にCNOTと呼びます。 Qiskitでは更に短く ```cx```と記述します。回路図の中では、下図のように描かれています。
```
qc_cnot = QuantumCircuit(2)
qc_cnot.cx(0,1)
qc_cnot.draw()
```
これは、2量子ビットに適用されます。 1つは制御量子ビットとして機能します(これは小さなドットのあるものです)。もう1つは(大きな円で)*ターゲット量子ビット*として機能します。
CNOTの効果を説明する方法はいくつかあります。 一つは、2つの入力ビットを調べて、それらが同じか異なるかを確認することです。次に、ターゲット量子ビットに回答を書き込みます。2つの入力ビットが同じ場合はターゲットは ```0```になり、異なる場合は``` 1```になります。
CNOTを説明する別の方法は、コントロールが ```1```の場合、ターゲットに対してNOTを行い、それ以外は何もしないと言うことです。この説明は、前の説明同様に有効です(ゲートの名前もここからつけられています)。
可能な入力をそれぞれ試して、CNOTを試してみてください。たとえば、入力 ```01```でCNOTをテストする回路を次に示します。
```
qc = QuantumCircuit(2,2)
qc.x(0)
qc.cx(0,1)
qc.measure(0,0)
qc.measure(1,1)
qc.draw()
```
この回路を実行すると、出力が ```11```であることがわかります。次のいずれかの理由からそうなることが考えられます。
- CNOTは入力値が互いに異なることを検出して ```1```を出力しています。これは、量子ビット1の状態(ビット列の左側にあることを思い出してください)を上書きし、 ```01```を ```11```に変換することで行います。
- CNOTは、量子ビット0が状態 ```1```であることを確認し、量子ビット1にNOTを適用します。これにより、量子ビット1の```0```が ```1```に変換されます。そして、```01```を``` 11```に変換します。
半加算器の場合、いずれの入力も結果で上書きしたくありません。そこで結果を異なる量子ビットのペアに書き込みたいと思います。これは2つのCNOTを使用することで実現できます。
```
qc_ha = QuantumCircuit(4,2)
# encode inputs in qubits 0 and 1
qc_ha.x(0) # For a=0, remove this line. For a=1, leave it.
qc_ha.x(1) # For b=0, remove this line. For b=1, leave it.
qc_ha.barrier()
# use cnots to write the XOR of the inputs on qubit 2
qc_ha.cx(0,2)
qc_ha.cx(1,2)
qc_ha.barrier()
# extract outputs
qc_ha.measure(2,0) # extract XOR value
qc_ha.measure(3,1)
qc_ha.draw()
```
半加算器の完成まであと半分の道のりとなりました。出力のもう1つのビット、つまり、4つ目の量子ビットが残っています。
可能な足し算の4つの答えをもう一度見ると、このビットが ```0```ではなく```1```であるケースが1つ: ```1 + 1``` = ```10```しかないことに気付くでしょう。追加するビットが両方とも```1```の場合にのみ発生します。
つまり、両方の入力が ```1```であるかどうかをコンピューターに確認させることでこの部分を計算することができます。両方の入力が```1```である場合のみ、量子ビット3でNOTゲートを実行する必要があります。
このためには、新しいゲートが必要です。CNOTのようですが、制御量子ビットが1つではなく2つになります。これは、両方の制御量子ビットの状態が ```1```の場合にのみ、ターゲット量子ビットに対してNOTを実行します。この新しいゲートは*Toffoli(トフォリ)*と呼ばれ、ブール論理ゲート上のANDゲートに相当します。
QiskitではToffoliは `ccx` というコマンドで実装されます。
```
qc_ha = QuantumCircuit(4,2)
# encode inputs in qubits 0 and 1
qc_ha.x(0) # For a=0, remove the this line. For a=1, leave it.
qc_ha.x(1) # For b=0, remove the this line. For b=1, leave it.
qc_ha.barrier()
# use cnots to write the XOR of the inputs on qubit 2
qc_ha.cx(0,2)
qc_ha.cx(1,2)
# use ccx to write the AND of the inputs on qubit 3
qc_ha.ccx(0,1,3)
qc_ha.barrier()
# extract outputs
qc_ha.measure(2,0) # extract XOR value
qc_ha.measure(3,1) # extract AND value
qc_ha.draw()
```
この例では、2つの入力ビットが両方とも`1`であるため、`1 + 1`を計算しています。何が得られるか見てみましょう。
```
qobj = assemble(qc_ha)
counts = sim.run(qobj).result().get_counts()
plot_histogram(counts)
```
結果は 「2」 のバイナリ表現である「10」です。有名な数学的問題 1+1 を解決できるコンピューターを構築しました!
ここで残る3つの足し算の入力で同じ事を試してみると、アルゴリズムがそれらに対しても正しい結果を与えることを示すことができます。
半加算器は足し算に必要な要素をすべて含んでいます。NOT, CNOTそしてToffoliゲートを用いることで、どのような大きさの数字の組合せでも足し合わせることができるプログラムを構築することができます。
この3つのゲートは、足し算だけでなくコンピューティングにおけるすべてを行うのに十分です。実際、CNOTなしでも実行できます。値が「1」のビットを作るために本当に必要なのはNOTゲートだけです。Toffoliゲートは本質的には数学の世界における原子のようなもので、すべての問題解決手法をコンパイルできる最も単純な要素と言えます。
このように、量子計算は原子を分割して行っていきます。
```
import qiskit.tools.jupyter
%qiskit_version_table
```
| github_jupyter |
# Nested Logit Model: Compute Willingness To Pay Indicators
```
import pandas as pd
import numpy as np
import biogeme.database as db
import biogeme.biogeme as bio
import biogeme.models as models
import biogeme.optimization as opt
import biogeme.results as res
from biogeme.expressions import Beta, DefineVariable, Derive
import seaborn as sns
import matplotlib.pyplot as plt
```
**Import Optima data**
```
pandas = pd.read_csv("../../Data/6-Discrete Choice Models/optima.dat",sep='\t')
database = db.Database ("data/optima", pandas)
```
**Use collumn names as variables**
```
globals().update(database.variables)
```
**Exclude some unwanted entries**
```
exclude = (Choice == -1.)
database.remove(exclude)
```
**Define some dummy variables**
```
male = (Gender == 1)
female = (Gender == 2)
unreportedGender = (Gender == -1)
fulltime = (OccupStat == 1)
notfulltime = (OccupStat != 1)
```
**Rescale some data**
```
TimePT_scaled = TimePT / 200
TimeCar_scaled = TimeCar / 200
MarginalCostPT_scaled = MarginalCostPT / 10
CostCarCHF_scaled = CostCarCHF / 10
distance_km_scaled = distance_km / 5
```
**Compute normalizing weights for each alternative**
```
sumWeight = database.data['Weight'].sum()
normalized_Weight = Weight * len(database.data['Weight']) / sumWeight
```
**Create parameters to be estimated**
```
ASC_CAR = Beta('ASC_CAR',0,None,None,0)
ASC_PT = Beta('ASC_PT',0,None,None,1)
ASC_SM = Beta('ASC_SM',0,None,None,0)
BETA_TIME_FULLTIME = Beta('BETA_TIME_FULLTIME',0,None,None,0)
BETA_TIME_OTHER = Beta('BETA_TIME_OTHER',0,None,None,0)
BETA_DIST_MALE = Beta('BETA_DIST_MALE',0,None,None,0)
BETA_DIST_FEMALE = Beta('BETA_DIST_FEMALE',0,None,None,0)
BETA_DIST_UNREPORTED = Beta('BETA_DIST_UNREPORTED',0,None,None,0)
BETA_COST = Beta('BETA_COST',0,None,None,0)
```
**Define the utility functions**
\begin{align}
V_{PT} & = \beta_{PT} + \beta_{time_{fulltime}} X_{time_{PT}} X_{fulltime} + \beta_{time_{other}} X_{time_{PT}} X_{not\_fulltime} + \beta_{cost} X_{cost_{PT}} \\
V_{car} & = \beta_{car} + \beta_{time_{fulltime}} X_{time_{car}} X_{fulltime} + \beta_{time_{other}} X_{time_{car}} X_{not\_fulltime} + \beta_{cost} X_{cost_{car}} \\
V_{SM} & = \beta_{SM} + \beta_{male} X_{distance} X_{male} + \beta_{female} X_{distance} X_{female} + \beta_{unreported} X_{distance} X_{unreported}
\end{align}
```
V_PT = ASC_PT + BETA_TIME_FULLTIME * TimePT_scaled * fulltime + \
BETA_TIME_OTHER * TimePT_scaled * notfulltime + \
BETA_COST * MarginalCostPT_scaled
V_CAR = ASC_CAR + \
BETA_TIME_FULLTIME * TimeCar_scaled * fulltime + \
BETA_TIME_OTHER * TimeCar_scaled * notfulltime + \
BETA_COST * CostCarCHF_scaled
V_SM = ASC_SM + \
BETA_DIST_MALE * distance_km_scaled * male + \
BETA_DIST_FEMALE * distance_km_scaled * female + \
BETA_DIST_UNREPORTED * distance_km_scaled * unreportedGender
```
**Associate utility functions with alternatives and associate availability of alternatives**
In this example all alternatives are available for each individual
```
V = {0: V_PT,
1: V_CAR,
2: V_SM}
av = {0: 1,
1: 1,
2: 1}
```
**Define the nests**
1. Define the nests paramenters
2. List alternatives in nests
```
MU_NO_CAR = Beta('MU_NO_CAR', 1.,1.,None,0)
CAR_NEST = 1., [1]
NO_CAR_NEST = MU_NO_CAR, [0, 2]
nests = CAR_NEST, NO_CAR_NEST
```
**Define the WTP**
```
WTP_PT_TIME = Derive(V_PT,'TimePT') / Derive(V_PT,'MarginalCostPT')
WTP_CAR_TIME = Derive(V_CAR,'TimeCar') / Derive(V_CAR,'CostCarCHF')
```
**Define what we want to simulate**
```
simulate = {'weight': normalized_Weight,
'WTP PT time': WTP_PT_TIME,
'WTP CAR time': WTP_CAR_TIME}
```
**Define the Biogeme object**
```
biogeme = bio.BIOGEME(database, simulate, removeUnusedVariables=False)
biogeme.modelName = "optima_nested_logit_wtp"
```
**Retrieve the names of the variables we want to use. Then retrieve the results from the model that we estimated earlier**
```
betas = biogeme.freeBetaNames
results = res.bioResults(pickleFile='optima_nested_logit.pickle')
betaValues = results.getBetaValues ()
```
**Perform the simulation**
```
simulatedValues = biogeme.simulate(betaValues)
```
**Compute the WTP for the car and its confidence interval**
```
wtpcar = (60 * simulatedValues['WTP CAR time'] * simulatedValues['weight']).mean()
b = results.getBetasForSensitivityAnalysis(betas,size=100)
left,right = biogeme.confidenceIntervals(b,0.9)
wtpcar_left = (60 * left['WTP CAR time'] * left['weight']).mean()
wtpcar_right = (60 * right['WTP CAR time'] * right['weight']).mean()
```
**Print the Time WTP for car**
```
print(f"Average WTP for car: {wtpcar:.3g} CI:[{wtpcar_left:.3g},{wtpcar_right:.3g}]")
```
**Analize all WTP values**
```
print("Unique values: ", [f"{i:.3g}" for i in 60 * simulatedValues['WTP CAR time'].unique()])
```
**Check this for groups within the population. Define a subgroup that automatically does this for us**
```
def wtpForSubgroup(filter):
size = filter.sum()
sim = simulatedValues[filter]
totalWeight = sim['weight'].sum()
weight = sim['weight'] * size / totalWeight
wtpcar = (60 * sim['WTP CAR time'] * weight ).mean()
wtpcar_left = (60 * left[filter]['WTP CAR time'] * weight ).mean()
wtpcar_right = (60 * right[filter]['WTP CAR time'] * weight ).mean()
return wtpcar, wtpcar_left,wtpcar_right
```
**Filter: Full time workers**
```
filter = database.data['OccupStat'] == 1
w,l,r = wtpForSubgroup(filter)
print(f"WTP car for workers: {w:.3g} CI:[{l:.3g},{r:.3g}]")
```
**Filter: Females**
```
filter = database.data['Gender'] == 2
w,l,r = wtpForSubgroup(filter)
print(f"WTP car for females: {w:.3g} CI:[{l:.3g},{r:.3g}]")
```
**Filter: Males**
```
filter = database.data['Gender'] == 1
w,l,r = wtpForSubgroup(filter)
print(f"WTP car for males: {w:.3g} CI:[{l:.3g},{r:.3g}]")
```
**Plot the distribution of individuals and their WTP in the population**
```
plt.hist(60*simulatedValues['WTP CAR time'],
weights = simulatedValues['weight'])
plt.xlabel("WTP (CHF/hour)")
plt.ylabel("Individuals")
plt.show()
```
| github_jupyter |
# KMeans Clustering with Normalizer
### Required Packages
```
!pip install plotly
import warnings
import operator
import itertools
import numpy as np
import pandas as pd, seaborn as sns
import plotly.express as px
import matplotlib.pyplot as plt
from mpl_toolkits import mplot3d
import plotly.graph_objects as go
from sklearn.pipeline import make_pipeline
from sklearn.cluster import KMeans
from sklearn.metrics import silhouette_score
from sklearn.preprocessing import Normalizer
warnings.filterwarnings('ignore')
from collections import OrderedDict
```
### Initialization
Filepath of CSV file
```
file_path = ""
```
List of features which are required for model training
```
features=[]
```
### Data Fetching
Pandas is an open-source, BSD-licensed library providing high-performance, easy-to-use data manipulation and data analysis tools.
We will use panda's library to read the CSV file using its storage path.And we use the head function to display the initial row or entry.
```
df=pd.read_csv(filename)
df.head()
```
### Feature Selections
It is the process of reducing the number of input variables when developing a predictive model. Used to reduce the number of input variables to both reduce the computational cost of modelling and, in some cases, to improve the performance of the model.
We will assign all the required input features to X.
```
# X = df[features]
X = df.iloc[:, 2:]
```
### Data Preprocessing
Since the majority of the machine learning models in the Sklearn library doesn't handle string category data and Null value, we have to explicitly remove or replace null values. The below snippet have functions, which removes the null value if any exists. And convert the string classes data in the datasets by encoding them to integer classes.
```
def NullClearner(df):
if(isinstance(df, pd.Series) and (df.dtype in ["float64","int64"])):
df.fillna(df.mean(),inplace=True)
return df
elif(isinstance(df, pd.Series)):
df.fillna(df.mode()[0],inplace=True)
return df
else:return df
def EncodeX(df):
return pd.get_dummies(df)
```
Calling preprocessing functions on the feature and target set.
```
x=X.columns.to_list()
for i in x:
X[i]=NullClearner(X[i])
X=EncodeX(X)
X.head()
```
### Data Rescaling
Normalize samples individually to unit norm. For each sample (i.e. a row in the matrix) containing non-zero components, its norm (l1, l2 or inf) is rescaled independently of the others, so that it equals one. Besides dense NumPy arrays, these transformers are capable of working with scipy as well.
[normalize](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.normalize.html)
```
X_normalize=Normalizer().fit_transform(X)
X_normalize=pd.DataFrame(data = X_normalize,columns = X.columns)
X_normalize.head()
```
### Elbow Method
[Info](https://en.wikipedia.org/wiki/Elbow_method_(clustering))
The Elbow Method is one of the most popular methods to determine this optimal value of k.
We iterate the values of k from 1 to 11 and calculate the distortion or inertia for each value of k in the given range.
Where Inertia is the sum of squared distances of samples to their closest cluster center.
To determine the optimal number of clusters, we have to select the value of k at the “elbow” ie the point after which the distortion/inertia start decreasing in a linear fashion.
```
def ElbowPlot(X):
SSE = []
for i in range(1, 11):
kmeans = KMeans(n_clusters=i,n_jobs=-1,random_state=123).fit(X)
SSE.append(kmeans.inertia_)
plt.plot(range(1, 11),SSE)
plt.title('Elbow Method')
plt.xlabel('Number of clusters')
plt.ylabel('SSE')
plt.show()
ElbowPlot(X_normalize)
```
Sometimes, the elbow method does not yield a clear decision (for example, if the elbow is not clear and sharp, or is ambiguous). In such cases, alternatives such as the [silhouette coefficient](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.silhouette_score.html) can be helpful.
### Silhouette Score
[Reference](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.silhouette_score.html)
* Silhouette Coefficient or silhouette score is a metric used to calculate the goodness of a clustering technique. Its value ranges from -1 to 1.
* The best value is 1 and the worst value is -1. Values near 0 indicate overlapping clusters. Negative values generally indicate that a sample has been assigned to the wrong cluster, as a different cluster is more similar.
```
def SilhoutteScore(X):
SScore=dict()
for num_clusters in range(2,11):
kmeans = KMeans(n_clusters=num_clusters,n_jobs=-1,random_state=123).fit(X)
SScore[num_clusters]=silhouette_score(X, kmeans.labels_)
SScore = OrderedDict( sorted(SScore.items(), key=operator.itemgetter(1),reverse=True))
return SScore
SilhoutteScore(X_normalize)
```
### Model
[Model API Reference](https://scikit-learn.org/stable/modules/generated/sklearn.cluster.KMeans.html)
K-means is a centroid-based algorithm, or a distance-based algorithm.The k-means algorithm divides a set of samples into disjoint clusters , each described by the mean of the samples in the cluster. The means are commonly called the cluster “centroids”; note that they are not, in general, points from , although they live in the same space.
The K-means algorithm aims to choose centroids that minimise the inertia, or within-cluster sum-of-squares criterion.
#### Tuning Parameter
> **n_clusters** : The number of clusters to form as well as the number of centroids to generate. To find appropriate number of clusters utilize elbow method or silhouette score.
> **init** : Method for initialization:
> * ‘k-means++’ : selects initial cluster centers for k-mean clustering in a smart way to speed up convergence. See section Notes in k_init for more details.
> * ‘random’: choose n_clusters observations (rows) at random from data for the initial centroids.
> **n_init*: Number of time the k-means algorithm will be run with different centroid seeds. The final results will be the best output of n_init consecutive runs in terms of inertia.
> **max_iter**: Maximum number of iterations of the k-means algorithm for a single run.
> **tol**: Relative tolerance with regards to Frobenius norm of the difference in the cluster centers of two consecutive iterations to declare convergence.
> **algorithm**: K-means algorithm to use. The classical EM-style algorithm is “full”. The “elkan” variation is more efficient on data with well-defined clusters, by using the triangle inequality. However it’s more memory intensive due to the allocation of an extra array of shape (n_samples, n_clusters).
```
kmeans = KMeans(n_clusters=6,n_jobs=-1,random_state=123)
pred_y = kmeans.fit_predict(X_normalize)
```
### Cluster Analysis
First, we add the cluster labels from the trained model into the copy of the data frame for cluster analysis/visualization.
```
ClusterDF = X_normalize.copy(deep=True)
ClusterDF['ClusterID'] = pred_y
ClusterDF.head()
```
#### Cluster Records
The below bar graphs show the number of data points in each available cluster.
```
ClusterDF['ClusterID'].value_counts().plot(kind='bar')
```
#### Cluster Plots
Below written functions get utilized to plot 2-Dimensional and 3-Dimensional cluster plots. Plots include different available clusters along with cluster centroid.
```
def Plot2DCluster(X_Cols,df):
for i in list(itertools.combinations(X_Cols, 2)):
plt.rcParams["figure.figsize"] = (8,6)
xi,yi=df.columns.get_loc(i[0]),df.columns.get_loc(i[1])
for j in df['ClusterID'].unique():
DFC=df[df.ClusterID==j]
plt.scatter(DFC[i[0]],DFC[i[1]],cmap=plt.cm.Accent,label=j)
plt.scatter(kmeans.cluster_centers_[:,xi],kmeans.cluster_centers_[:,yi],marker="^",color="black",label="centroid")
plt.xlabel(i[0])
plt.ylabel(i[1])
plt.legend()
plt.show()
def Plot3DCluster(X_Cols,df):
for i in list(itertools.combinations(X_Cols, 3)):
xi,yi,zi=df.columns.get_loc(i[0]),df.columns.get_loc(i[1]),df.columns.get_loc(i[2])
fig,ax = plt.figure(figsize = (16, 10)),plt.axes(projection ="3d")
ax.grid(b = True, color ='grey',linestyle ='-.',linewidth = 0.3,alpha = 0.2)
for j in df['ClusterID'].unique():
DFC=df[df.ClusterID==j]
ax.scatter3D(DFC[i[0]],DFC[i[1]],DFC[i[2]],alpha = 0.8,cmap=plt.cm.Accent,label=j)
ax.scatter3D(kmeans.cluster_centers_[:,xi],kmeans.cluster_centers_[:,yi],kmeans.cluster_centers_[:,zi],
marker="^",color="black",label="centroid")
ax.set_xlabel(i[0])
ax.set_ylabel(i[1])
ax.set_zlabel(i[2])
plt.legend()
plt.show()
def Plotly3D(X_Cols,df):
for i in list(itertools.combinations(X_Cols,3)):
xi,yi,zi=df.columns.get_loc(i[0]),df.columns.get_loc(i[1]),df.columns.get_loc(i[2])
fig1 = px.scatter_3d(kmeans.cluster_centers_,x=kmeans.cluster_centers_[:,xi],y=kmeans.cluster_centers_[:,yi],
z=kmeans.cluster_centers_[:,zi])
fig2=px.scatter_3d(df, x=i[0], y=i[1],z=i[2],color=df['ClusterID'])
fig3 = go.Figure(data=fig1.data + fig2.data,
layout=go.Layout(title=go.layout.Title(text="x:{}, y:{}, z:{}".format(i[0],i[1],i[2])))
)
fig3.show()
Plot2DCluster(X.columns,ClusterDF)
sns.set_style("whitegrid")
sns.set_context("talk")
plt.rcParams["lines.markeredgewidth"] = 1
sns.pairplot(data=ClusterDF, hue='ClusterID', palette='Dark2', height=5)
Plot3DCluster(X.columns,ClusterDF)
Plotly3D(X.columns,ClusterDF)
```
**creator: Virat Chowdary, GitHub: [profile](https://github.com/viratchowdary21)**
| github_jupyter |
# CNN robustness of CIFAR-10
In this notebook we evaluate the samples we collected over different runs on CIFAR-10.
```
import os
from jax import numpy as jnp
import numpy as onp
import jax
import tensorflow.compat.v2 as tf
import argparse
import time
import tqdm
from collections import OrderedDict
from bnn_hmc.utils import data_utils
from bnn_hmc.utils import models
from bnn_hmc.utils import metrics
from bnn_hmc.utils import losses
from bnn_hmc.utils import checkpoint_utils
from bnn_hmc.utils import cmd_args_utils
from bnn_hmc.utils import logging_utils
from bnn_hmc.utils import train_utils
from bnn_hmc.utils import precision_utils
from itertools import chain
from matplotlib import pyplot as plt
import matplotlib
%matplotlib inline
%load_ext autoreload
%autoreload 2
dtype = jnp.float32
train_set, test_set, task, data_info = data_utils.make_ds_pmap_fullbatch(
"cifar10", dtype)
test_labels = test_set[1]
net_apply, net_init = models.get_model("lenet", data_info)
net_apply = precision_utils.rewrite_high_precision(net_apply)
(_, predict_fn, ensemble_upd_fn, _,_) = train_utils.get_task_specific_fns(task, data_info)
def predict_hmc_seeds(ds, path_dict):
all_preds = {}
for seed, path in path_dict.items():
all_preds[seed] = predict_hmc(ds, path)[0].copy()
return all_preds
def predict_hmc(ds, basepath):
basepath = os.path.join(basepath, "model_step_{}.pt")
ensemble_predictions = None
num_ensembled = 0
last_test_predictions = None
for i in tqdm.tqdm(range(11, 100)):
path = basepath.format(i)
try:
checkpoint_dict = checkpoint_utils.load_checkpoint(path)
except:
continue
_, params, net_state, _, _, accepted, _, _ = (
checkpoint_utils.parse_hmc_checkpoint_dict(checkpoint_dict))
if accepted:
_, test_predictions = onp.asarray(
predict_fn(net_apply, params, net_state, ds))
ensemble_predictions = ensemble_upd_fn(
ensemble_predictions, num_ensembled, test_predictions)
num_ensembled += 1
last_test_predictions = test_predictions
print(num_ensembled)
return ensemble_predictions, last_test_predictions
def predict_de(ds, path_dict):
ensemble_predictions = None
num_ensembled = 0
# last_test_predictions = None
all_predictions = {}
for seed, path in tqdm.tqdm(path_dict.items()):
checkpoint_dict = checkpoint_utils.load_checkpoint(path)
params = checkpoint_dict["params"]
net_state = checkpoint_dict["net_state"]
_, test_predictions = onp.asarray(
predict_fn(net_apply, params, net_state, ds))
ensemble_predictions = ensemble_upd_fn(
ensemble_predictions, num_ensembled, test_predictions)
num_ensembled += 1
all_predictions[seed] = test_predictions.copy()
if num_ensembled >= 10:
break
print(num_ensembled)
return ensemble_predictions, last_test_predictions
```
## Dirs
```
gaussdirs = {}
prefix = "../runs/hmc/cifar10/lenet/gaussian/"
dirs = os.listdir(prefix)
for path in dirs:
full_path = os.path.join(prefix, path)
wd = float(full_path.split("_")[3])
stepsize = float(full_path.split("_")[5])
trajlen = float(full_path.split("_")[7])
# name = "Gaussian wd={} T=1".format(wd, stepsize)
gaussdirs[wd] = full_path
gaussdirs
sumfilterdirs = {}
# prefix = "../runs/hmc/cifar10/lenet/sumfilter/"
prefix = "../runs/hmc/cifar10/lenet/pca_wd100/"
dirs = os.listdir(prefix)
for path in dirs:
full_path = os.path.join(prefix, path)
wd = float(full_path.split("_")[4])
stepsize = float(full_path.split("_")[6])
trajlen = float(full_path.split("_")[8])
# name = "Gaussian wd={} T=1".format(wd, stepsize)
sumfilterdirs[wd] = full_path
sumfilterdirs
sgddirs = {}
prefix = "../runs/sgd/cifar10/lenet/"
dirs = os.listdir(prefix)
for path in dirs:
if path[:3] != "sgd":
continue
full_path = os.path.join(prefix, path, "model_step_299.pt")
wd = float(full_path.split("_")[13])
seed = float(full_path.split("_")[-3].split("/")[0])
if wd not in sgddirs:
sgddirs[wd] = {}
sgddirs[wd][seed] = full_path
sgddirs
arr = onp.load("cifar10_patch_pca.npz")
pca = {
"basis": arr["components"],
"mean": arr["mean"],
"explained_var_ratio": arr["explained_var_ratio"]
}
layer_pcas = {
'conv2_d': pca,
'conv2_d_1': None,
'linear': None,
'linear_1': None,
'linear_2': None,
}
def get_n_components(rel_explained_vars, cutoff):
cumsum = onp.cumsum(rel_explained_vars)
n_components = onp.where(cumsum > cutoff)[0][0]
return n_components
def pca_ajust_layer(params, layer_name, conv_cutoff, linear_cutoff):
is_conv = layer_name[:5] == "conv2"
w = onp.asarray(params[layer_name]["w"])
b = onp.asarray(params[layer_name]["b"])
original_shape = w.shape
pca_dict = layer_pcas[layer_name]
if pca_dict is None:
return w, b
basis = pca_dict["basis"]
mean = pca_dict["mean"]
explained_var_ratio = pca_dict["explained_var_ratio"]
cutoff = conv_cutoff if is_conv else linear_cutoff
n_components = get_n_components(explained_var_ratio, cutoff)
basis = basis[:n_components]
print("{} \tRetained components: {} / {}".format(
layer_name, n_components, basis.shape[1]))
if is_conv:
ks, _, i_c, o_c = w.shape
w = w.reshape((ks**2 * i_c, -1))
w = w.transpose(1, 0)
# print(w.shape, basis.shape, mean.shape)
original_w_mean = (w * mean[None, :]).sum()
w = (w) @ basis.T @ basis
new_w_mean = (w * mean[None, :]).sum()
w = w.transpose(1, 0)
w = w.reshape(original_shape)
else:
w = w.transpose(1, 0)
original_w_mean = w @ mean
w = w @ basis.T @ basis
new_w_mean = w @ mean
w = w.transpose(1, 0)
bias_shift = original_w_mean - new_w_mean
# print(bias_shift)
b = b + bias_shift
return w, b
# w = pca_ajust_layer(params, 'conv2_d')
# w = pca_ajust_layer(params, 'conv2_d_1')
# w = pca_ajust_layer(params, 'linear')
# w = pca_ajust_layer(params, 'linear_1')
# w = pca_ajust_layer(params, 'linear_2')
def adjust_pca_projections(params, conv_cutoff=0.99, linear_cutoff=0.9):
params_flat, treedef = jax.tree_flatten(params)
new_params = {layer_name: {p: v for p, v in layer_dict.items()}
for layer_name, layer_dict in params.items()}
for layer_name in layer_pcas.keys():
w, b = pca_ajust_layer(params, layer_name, conv_cutoff, linear_cutoff)
new_params[layer_name]["w"] = jnp.asarray(w)
new_params[layer_name]["b"] = jnp.asarray(b)
new_params_flat, _ = jax.tree_flatten(new_params)
new_params = jax.tree_unflatten(treedef, new_params_flat)
return new_params
def predict_hmc(ds, basepath, adjust_pca=False):
basepath = os.path.join(basepath, "model_step_{}.pt")
ensemble_predictions = None
num_ensembled = 0
last_test_predictions = None
for i in tqdm.tqdm(range(11, 100)):
path = basepath.format(i)
try:
checkpoint_dict = checkpoint_utils.load_checkpoint(path)
except:
continue
_, params, net_state, _, _, accepted, _, _ = (
checkpoint_utils.parse_hmc_checkpoint_dict(checkpoint_dict))
if adjust_pca:
params = adjust_pca_projections(params)
# return None
if accepted:
_, test_predictions = onp.asarray(
predict_fn(net_apply, params, net_state, ds))
ensemble_predictions = ensemble_upd_fn(
ensemble_predictions, num_ensembled, test_predictions)
num_ensembled += 1
last_test_predictions = test_predictions
print(num_ensembled)
return ensemble_predictions, last_test_predictions
def predict_de(ds, path_dict, adjust_pca=False):
ensemble_predictions = None
num_ensembled = 0
last_test_predictions = None
for path in tqdm.tqdm(path_dict.values()):
checkpoint_dict = checkpoint_utils.load_checkpoint(path)
params = checkpoint_dict["params"]
net_state = checkpoint_dict["net_state"]
_, test_predictions = onp.asarray(
predict_fn(net_apply, params, net_state, ds))
ensemble_predictions = ensemble_upd_fn(
ensemble_predictions, num_ensembled, test_predictions)
num_ensembled += 1
last_test_predictions = test_predictions
print(num_ensembled)
return ensemble_predictions, last_test_predictions
# hmc_predictions, predictions = predict_hmc(test_set, gaussdirs[100], adjust_pca=True)
# print(metrics.accuracy(hmc_predictions, test_labels))
```
## CIFAR-10-C
```
from bnn_hmc.utils import data_utils
import tensorflow_datasets as tfds
all_corruptions = tfds.image_classification.cifar10_corrupted._CORRUPTIONS
all_corruptions
def load_image_dataset(
split, batch_size, name, repeat=False, shuffle=False,
shuffle_seed=None
):
ds, dataset_info = tfds.load(name, split=split, as_supervised=True,
with_info=True)
num_classes = dataset_info.features["label"].num_classes
num_examples = dataset_info.splits[split].num_examples
num_channels = dataset_info.features['image'].shape[-1]
def img_to_float32(image, label):
return tf.image.convert_image_dtype(image, tf.float32), label
ds = ds.map(img_to_float32).cache()
ds_stats = data_utils._ALL_IMG_DS_STATS[data_utils.ImgDatasets("cifar10")]
def img_normalize(image, label):
"""Normalize the image to zero mean and unit variance."""
mean, std = ds_stats
image -= tf.constant(mean, shape=[1, 1, num_channels], dtype=image.dtype)
image /= tf.constant(std, shape=[1, 1, num_channels], dtype=image.dtype)
return image, label
ds = ds.map(img_normalize)
if batch_size == -1:
batch_size = num_examples
if repeat:
ds = ds.repeat()
if shuffle:
ds = ds.shuffle(buffer_size=10 * batch_size, seed=shuffle_seed)
ds = ds.batch(batch_size)
return tfds.as_numpy(ds), num_classes, num_examples
def get_image_dataset(name):
test_set, n_classes, _ = load_image_dataset("test", -1, name)
test_set = next(iter(test_set))
data_info = {
"num_classes": n_classes
}
return test_set, data_info
def pmap_dataset(ds, n_devices):
return jax.pmap(lambda x: x)(data_utils.batch_split_axis(ds, n_devices))
def make_cifar10c_pmap_fullbatch(corruption, intensity, dtype, n_devices=None, truncate_to=None):
"""Make train and test sets sharded over batch dim."""
name = get_ds_name(corruption, intensity).lower()
n_devices = n_devices or len(jax.local_devices())
test_set, data_info = get_image_dataset(name)
loaded = True
test_set = pmap_dataset(test_set, n_devices)
test_set = test_set[0].astype(dtype), test_set[1]
return test_set, task, data_info
def get_ds_name(corruption, intensity):
return "cifar10_corrupted/{}_{}".format(corruption, intensity)
dtype = jnp.float32
test_set, task, data_info = make_cifar10c_pmap_fullbatch(all_corruptions[0], 3, dtype)
test_labels = test_set[1]
net_apply, net_init = models.get_model("lenet", data_info)
net_apply = precision_utils.rewrite_high_precision(net_apply)
(_, predict_fn, ensemble_upd_fn, _,_) = train_utils.get_task_specific_fns(task, data_info)
prior_dirs = {
"gauss": gaussdirs,
"sumfilter": sumfilterdirs
}
sgd_ind_accs = {key: {} for key in sgddirs}
sgd_ens_accs = {key: {} for key in sgddirs}
hmc_ind_accs = {prior: {key: {} for key in dirs}
for prior, dirs in prior_dirs.items()}
hmc_ens_accs = {prior: {key: {} for key in dirs}
for prior, dirs in prior_dirs.items()}
pca_hmc_ind_accs = {prior: {key: {} for key in dirs}
for prior, dirs in prior_dirs.items()}
pca_hmc_ens_accs = {prior: {key: {} for key in dirs}
for prior, dirs in prior_dirs.items()}
for corruption in all_corruptions:
for intensity in [3]:
noisy_test_set, _, _ = make_cifar10c_pmap_fullbatch(corruption, intensity, dtype)
test_labels = noisy_test_set[1]
for prior, dirs in prior_dirs.items():
for key, path in dirs.items():
hmc_predictions, predictions = predict_hmc(noisy_test_set, path)
if hmc_predictions is None:
continue
hmc_ens_accs[prior][key][corruption, intensity] = metrics.accuracy(hmc_predictions, test_labels)
hmc_ind_accs[prior][key][corruption, intensity] = metrics.accuracy(predictions, test_labels)
# hmc_predictions, predictions = predict_hmc(noisy_test_set, path, adjust_pca=True)
# if hmc_predictions is None:
# continue
# pca_hmc_ens_accs[prior][key][corruption, intensity] = metrics.accuracy(hmc_predictions, test_labels)
# pca_hmc_ind_accs[prior][key][corruption, intensity] = metrics.accuracy(predictions, test_labels)
for key, pathdict in sgddirs.items():
de_predictions, predictions = predict_de(noisy_test_set, pathdict)
sgd_ind_accs[key][corruption, intensity] = metrics.accuracy(predictions, test_labels)
sgd_ens_accs[key][corruption, intensity] = metrics.accuracy(de_predictions, test_labels)
datasets = ["CIFAR10", "stl10_like_cifar"]
for ds_name in datasets:
dtype = jnp.float32
train_set, test_set, task, data_info = data_utils.make_ds_pmap_fullbatch(
ds_name, dtype)
test_labels = test_set[1]
for prior, dirs in prior_dirs.items():
for key, path in dirs.items():
hmc_predictions, predictions = predict_hmc(test_set, path)
if hmc_predictions is None:
continue
for intensity in range(1, 6):
hmc_ens_accs[prior][key][ds_name.upper(), intensity] = metrics.accuracy(hmc_predictions, test_labels)
hmc_ind_accs[prior][key][ds_name.upper(), intensity] = metrics.accuracy(predictions, test_labels)
# hmc_predictions, predictions = predict_hmc(test_set, path, adjust_pca=True)
# for intensity in range(1, 6):
# pca_hmc_ens_accs[prior][key][ds_name.upper(), intensity] = metrics.accuracy(hmc_predictions, test_labels)
# pca_hmc_ind_accs[prior][key][ds_name.upper(), intensity] = metrics.accuracy(predictions, test_labels)
for corruption in datasets:
de_predictions, predictions = predict_de(test_set, pathdict)
for intensity in range(1, 6):
sgd_ind_accs[key][ds_name.upper(), intensity] = metrics.accuracy(predictions, test_labels)
sgd_ens_accs[key][ds_name.upper(), intensity] = metrics.accuracy(de_predictions, test_labels)
cmap = matplotlib.cm.get_cmap("tab20")
num_bars = 4
width = 0.8 / num_bars
plt.figure(figsize=(3, 7))
all_corruptions_ = onp.unique([k[0] for k in hmc_ens_accs["gauss"][100.]])
ys = onp.arange(len(all_corruptions_))
std = onp.sqrt(1 / wd)
plt.barh(ys, [hmc_ens_accs["gauss"][wd][c, 3] for c in all_corruptions_],
height=width, label="HMC Gaussian".format(std), color=cmap(0)
)
plt.barh(ys + width, [hmc_ens_accs["sumfilter"][wd][c, 3] for c in all_corruptions_],
height=width, label="HMC PCA Prior".format(std), color=cmap(1)
)
# plt.barh(ys + width, [pca_hmc_ens_accs["gauss"][wd][c, 3] for c in all_corruptions_],
# height=width, label="HMC PCA".format(std), color=cmap(1)
# )
plt.barh(ys + 2 * width, [sgd_ind_accs[100][c, 3] for c in all_corruptions_],
height=width, label="SGD", color="k" #cmap(len(wds)+1)
)
plt.barh(ys + 3 * width, [sgd_ens_accs[100][c, 3] for c in all_corruptions_],
height=width, label="DE", color="grey" #cmap(len(wds)+1)
)
plt.ylim(-0.15, 21.05)
plt.legend(loc=(1, 0))
plt.yticks(ys+0.4, all_corruptions_, rotation=0);
# plt.savefig("gaussian_priors_mnistc.pdf", bbox_inches="tight")
```
## Filter visualization
```
# path = os.path.join(gaussdirs[100.], "model_step_90.pt")
path = os.path.join(sumfilterdirs[100.], "model_step_12.pt")
checkpoint_dict = checkpoint_utils.load_checkpoint(path)
_, params, net_state, _, _, accepted, _, _ = (
checkpoint_utils.parse_hmc_checkpoint_dict(checkpoint_dict))
params.keys()
w = params['conv2_d']['w']
print(w.shape)
f, arr = plt.subplots(2, 3, figsize=(10, 5))
for i in range(2):
for j in range(3):
arr[i, j].imshow(w[:, :, 0, i + 2 * j])
w = params['linear']['w']
print(w.shape)
f, arr = plt.subplots(2, 3, figsize=(10, 5))
for i in range(2):
for j in range(3):
arr[i, j].imshow(w[:, i + 2 * j].reshape(24, 24))
```
### SGD
```
path = sgddirs[100.][2.0]
checkpoint_dict = checkpoint_utils.load_checkpoint(path)
_, params, net_state, _, _ = (
checkpoint_utils.parse_sgd_checkpoint_dict(checkpoint_dict))
w = params['conv2_d_1']['w']
print(w.shape)
f, arr = plt.subplots(2, 3, figsize=(10, 5))
for i in range(2):
for j in range(3):
arr[i, j].imshow(w[:, :, 0, i + 2 * j])
w = params['linear']['w']
print(w.shape)
f, arr = plt.subplots(2, 3, figsize=(10, 5))
for i in range(2):
for j in range(3):
arr[i, j].imshow(w[:, i + 2 * j].reshape(24, 24))
sgd_effective_sigmas = jax.tree_map(lambda p: jnp.linalg.norm(p) / jnp.sqrt(p.size), params)
sgd_effective_sigmas
```
## Weight PCA Projections
```
wd = 100
prior_std = 1 / onp.sqrt(wd)
w = None
base_path = os.path.join(gaussdirs[100.], "model_step_{}.pt")
for step in range(20, 99):
path = base_path.format(step)
checkpoint_dict = checkpoint_utils.load_checkpoint(path)
if checkpoint_dict["accepted"]:
params = checkpoint_dict["params"]
cur_w = onp.asarray(params["conv2_d"]["w"].reshape((75, -1)))
cur_w = cur_w.transpose(1, 0)
if w is None:
w = cur_w.copy()
else:
w = onp.concatenate([w, cur_w.copy()], axis=0)
arr = onp.load("cifar10_patch_pca.npz")
basis = arr["components"]
projections = (w[:, None, :] * basis[None, :, :]).sum(-1)
mean = onp.mean(projections, axis=0)
std = onp.std(projections, axis=0)
plt.plot(mean + 2 * std, "b")
plt.plot(mean - 2 * std, "b")
plt.fill_between(onp.arange(75), mean + 2 * std, mean - 2 * std, color="b", alpha=0.3)
for p in projections:
plt.plot(p, "--b", alpha=0.02)
plt.grid()
plt.hlines(2 * prior_std, 0, 75, color="k", linestyle="dashed", lw=3)
plt.hlines(-2 * prior_std, 0, 75, color="k", linestyle="dashed", lw=3)
plt.xlim(0, 75)
plt.xlabel("PCA component")
plt.ylabel("Projection of filter on PCA component")
plt.title("HMC samples, wd={}".format(wd))
```
## Sum-filter / pca prior
```
w = None
base_path = os.path.join(sumfilterdirs[100.], "model_step_{}.pt")
for step in range(20, 99):
path = base_path.format(step)
checkpoint_dict = checkpoint_utils.load_checkpoint(path)
if checkpoint_dict["accepted"]:
params = checkpoint_dict["params"]
cur_w = onp.asarray(params["conv2_d"]["w"].reshape((75, -1)))
cur_w = cur_w.transpose(1, 0)
if w is None:
w = cur_w.copy()
else:
w = onp.concatenate([w, cur_w.copy()], axis=0)
projections = (w[:, None, :] * basis[None, :, :]).sum(-1)
mean = onp.mean(projections, axis=0)
std = onp.std(projections, axis=0)
plt.plot(mean + 2 * std, "b")
plt.plot(mean - 2 * std, "b")
plt.fill_between(onp.arange(75), mean + 2 * std, mean - 2 * std, color="b", alpha=0.3)
for p in projections:
plt.plot(p, "--b", alpha=0.02)
plt.grid()
plt.hlines(2 * prior_std, 0, 75, color="k", linestyle="dashed", lw=3)
plt.hlines(-2 * prior_std, 0, 75, color="k", linestyle="dashed", lw=3)
plt.xlim(0, 75)
plt.xlabel("PCA component")
plt.ylabel("Projection of filter on PCA component")
plt.title("HMC samples, wd={}".format(wd))
```
| github_jupyter |
```
import google.datalab.bigquery as bq
import pandas as pd
import numpy as np
import shutil
%bigquery schema --table "nyc-tlc:green.trips_2015"
%bq query
SELECT EXTRACT (DAYOFYEAR from pickup_datetime) AS daynumber FROM `nyc-tlc.green.trips_2015` LIMIT 10
%bq query -n taxiquery
WITH trips AS (
SELECT EXTRACT (DAYOFYEAR from pickup_datetime) AS daynumber FROM `nyc-tlc.green.trips_*`
where _TABLE_SUFFIX = @YEAR
)
SELECT daynumber, COUNT(1) AS numtrips FROM trips
GROUP BY daynumber ORDER BY daynumber
query_parameters = [
{
'name': 'YEAR',
'parameterType': {'type': 'STRING'},
'parameterValue': {'value': 2015}
}
]
trips = taxiquery.execute(query_params=query_parameters).result().to_dataframe()
trips[:5]
%bq query
SELECT * FROM `bigquery-public-data.noaa_gsod.stations`
WHERE state = 'NY' AND wban != '99999' AND name LIKE '%LA GUARDIA%'
avg = np.mean(trips['numtrips'])
print 'Just using average={0} has RMSE of {1}'.format(avg, np.sqrt(np.mean((trips['numtrips'] - avg)**2)))
%bq query -n wxquery
SELECT EXTRACT (DAYOFYEAR FROM CAST(CONCAT(@YEAR,'-',mo,'-',da) AS TIMESTAMP)) AS daynumber,
MIN(EXTRACT (DAYOFWEEK FROM CAST(CONCAT(@YEAR,'-',mo,'-',da) AS TIMESTAMP))) dayofweek,
MIN(min) mintemp, MAX(max) maxtemp, MAX(IF(prcp=99.99,0,prcp)) rain
FROM `fh-bigquery.weather_gsod.gsod*`
WHERE stn='725030' AND _TABLE_SUFFIX = @YEAR
GROUP BY 1 ORDER BY daynumber DESC
```
Group by 1 adds a column with the row number starting from zero
```
query_parameters = [
{
'name': 'YEAR',
'parameterType': {'type': 'STRING'},
'parameterValue': {'value': 2015}
}
]
weather = wxquery.execute(query_params=query_parameters).result().to_dataframe()
weather[:5]
data = pd.merge(weather, trips, on='daynumber') #uses pandas to merge the two query results on the daynumber field and stores it in a table called data
data[:5]
j = data.plot(kind='scatter', x='maxtemp', y='numtrips')
j = data.plot(kind='scatter', x='dayofweek', y='numtrips')
j = data[data['dayofweek'] == 7].plot(kind='scatter', x='maxtemp', y='numtrips')
query_parameters = [
{
'name': 'YEAR',
'parameterType': {'type': 'STRING'},
'parameterValue': {'value': 2014}
}
]
weather = wxquery.execute(query_params=query_parameters).result().to_dataframe()
trips = taxiquery.execute(query_params=query_parameters).result().to_dataframe()
data2014 = pd.merge(weather, trips, on='daynumber')
data2014[:5]
data2 = pd.concat([data, data2014])
data2.describe()
j = data2[data2['dayofweek'] == 7].plot(kind='scatter', x='maxtemp', y='numtrips')
import tensorflow as tf
shuffled = data2.sample(frac=1) # this shuffles the entire sample (frac=1 or 100% of the original data set)
# It would be a good idea, if we had more data, to treat the days as categorical variables
# with the small amount of data, we have though, the model tends to overfit
#predictors = shuffled.iloc[:,2:5]
#for day in xrange(1,8):
# matching = shuffled['dayofweek'] == day
# key = 'day_' + str(day)
# predictors[key] = pd.Series(matching, index=predictors.index, dtype=float)
predictors = shuffled.iloc[:,1:5] #iloc slices or splits up the dataframe according to integer location (i.e. the row and/or column coordinates provided)
predictors[:5]
shuffled[:5] #the first five entries/rows of entire shuffled data set
targets = shuffled.iloc[:,5]
targets[:5]
#updates the benchmarks to test predictions using the shuffled data sample
trainsize = int(len(shuffled['numtrips']) * 0.8) #len returns the number of rows, multiplied by .8 gives the sample size for training
avg = np.mean(shuffled['numtrips'][:trainsize])
rmse = np.sqrt(np.mean((targets[trainsize:] - avg)**2))
print 'Just using average={0} has RMSE of {1}'.format(avg, rmse)
SCALE_NUM_TRIPS = 100000.0
trainsize = int(len(shuffled['numtrips']) * 0.8)
testsize = len(shuffled['numtrips']) - trainsize
npredictors = len(predictors.columns)
noutputs = 1
tf.logging.set_verbosity(tf.logging.WARN) # change to INFO to get output every 100 steps ...
shutil.rmtree('./trained_model_linear', ignore_errors=True) # so that we don't load weights from previous runs
estimator = tf.contrib.learn.LinearRegressor(model_dir='./trained_model_linear',
optimizer=tf.train.AdamOptimizer(learning_rate=0.1),
enable_centered_bias=False,
feature_columns=tf.contrib.learn.infer_real_valued_columns_from_input(predictors.values))
print "starting to train ... this will take a while ... use verbosity=INFO to get more verbose output"
def input_fn(features, targets):
return tf.constant(features.values), tf.constant(targets.values.reshape(len(targets), noutputs)/SCALE_NUM_TRIPS)
estimator.fit(input_fn=lambda: input_fn(predictors[:trainsize], targets[:trainsize]), steps=10000)
pred = np.multiply(list(estimator.predict(predictors[trainsize:].values)), SCALE_NUM_TRIPS )
rmse = np.sqrt(np.mean(np.power((targets[trainsize:].values - pred), 2)))
print 'LinearRegression has RMSE of {0}'.format(rmse)
SCALE_NUM_TRIPS = 100000.0
trainsize = int(len(shuffled['numtrips']) * 0.8)
testsize = len(shuffled['numtrips']) - trainsize
npredictors = len(predictors.columns)
noutputs = 1
tf.logging.set_verbosity(tf.logging.WARN) # change to INFO to get output every 100 steps ...
shutil.rmtree('./trained_model', ignore_errors=True) # so that we don't load weights from previous runs
estimator = tf.contrib.learn.DNNRegressor(model_dir='./trained_model',
hidden_units=[5, 2],
optimizer=tf.train.AdamOptimizer(learning_rate=0.01),
enable_centered_bias=False,
feature_columns=tf.contrib.learn.infer_real_valued_columns_from_input(predictors.values))
print "starting to train ... this will take a while ... use verbosity=INFO to get more verbose output"
def input_fn(features, targets):
return tf.constant(features.values), tf.constant(targets.values.reshape(len(targets), noutputs)/SCALE_NUM_TRIPS)
estimator.fit(input_fn=lambda: input_fn(predictors[:trainsize], targets[:trainsize]), steps=10000)
pred = np.multiply(list(estimator.predict(predictors[trainsize:].values)), SCALE_NUM_TRIPS )
rmse = np.sqrt(np.mean((targets[trainsize:].values - pred)**2))
print 'Neural Network Regression has RMSE of {0}'.format(rmse)
```
THE CODE BELOW CREATES NEW INPUTS AND RUNS THOSE THROUGH THE TRAINED MODEL TO PREDICT THE NUMBER OF RIDES (i.e. DEMAND)
```
input = pd.DataFrame.from_dict(data =
{'dayofweek' : [4, 5, 6],
'mintemp' : [30, 60, 50],
'maxtemp' : [40, 70, 60],
'rain' : [0, 0.8, 0]})
# read trained model from ./trained_model
estimator = tf.contrib.learn.LinearRegressor(model_dir='./trained_model_linear',
enable_centered_bias=False,
feature_columns=tf.contrib.learn.infer_real_valued_columns_from_input(input.values))
pred = np.multiply(list(estimator.predict(input.values)), SCALE_NUM_TRIPS )
print pred
```
| github_jupyter |
# Accelerated Python @ Columbia University
*Instructor: Sameer (Sam) Ansari*
Link to this doc: http://bit.ly/accelerated-python
**TL;DR** Get current/future researchers comfortable with using colab / python / numpy / pandas / sklearn for basic data analysis on some small datasets.
| Times | Duration | |
| -------------: | :------------- | ------------- |
| **9 am** | | **Start** - Thurs, Aug 15, 2019 |
| 9:00 - 9:15 | 15 min | Intro to Colab |
| 9:15 - 9:30 | 15 min | How this relates to research @ Google |
| 9:30 - 9:45 | 15 min | Python (intro, lists, functions) |
| 9:45 - 10:15 | 30 min | Python (defaultdict, map/filter/reduce)
| **10:15 - 10:30** | **15 min** | **Break** |
| 10:30 - 10:45 | 15 min | Numpy (arrays, loading CSV, random, mean/std) ||
| 10:45 - 11:15 | 30 min | Matplotlib (intro, visualizing numpy, histograms)
| 11:15 - 12 | 45 min | Pandas (intro, load CSV) |
| **12 - 1** | **1 hr** | **Lunch** |
| 1 - 1:15 | 15min | Pandas (Basic data manipulation, boolean lists, filtering nans) |
| 1:15 - 1:45 | 30 min | Pandas (grouping, visualization, std)
| 1:45- 2:15 | 30 min | sklearn (intro, load existing dataset, visualize)
| **2:15 - 2:30** | **15 min** | **Break** |
| 2:30 - 2:45 | 15 min | Extras if time: sklearn (Regression)
| 2:45 - 3:15 | 30 min | Extras if time: sklearn (Clustering)
| 3:15 - 3:45 | 30 min | Extras if time: discuss research, prediction, image manipulation?
| 3:45 - 4:00 | 15 min | Wrap up, survey, questions
| **4 pm** | | **End** |
# Morning Setup
For the class, I plan on keeping this completed notebook open in one tab to refer to, and a new tab with an empty colab notebook `(File -> New Python3 notebook)` which I'll work through in class with all of you. Feel free to do the same.
This isn't necessary for the course, but you can also check out the more polished [tutorials](https://colab.research.google.com/notebooks/welcome.ipynb) introducing the [basic features of colab](https://colab.research.google.com/notebooks/basic_features_overview.ipynb).
Make sure you can run (click the code cell and then small play button on the left) the following command before class if possible, it should just work.
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import scipy
from sklearn.cluster import KMeans
from sklearn.preprocessing import StandardScaler
print('All good')
```
# Intro
How the things we're talking about relate:
- **numpy** extends python to work well with numeric arrays, lots of useful algorithsm
- **matplotlib** on top of python, pulls over most of matlabs plotting stuff.
- **pandas** uses numpy, tons of helpers for manipulating datasets/tables
- **sklearn** is a bunch of algorithms/modules/etc. that uses pandas/numpy.
These are a lot of general topics, so note that this class is **not** a deep dive into data analysis, many of you probably know more about that than me.
This class is for getting you comfortable with how to use Python to load and do basic processing and analysis of datasets, as well as learn about the types of tools out there that can aid you for doing data analysis.
Python has an absurd amount of libraries and APIs available for almost anything you can think of, which should make it one of your first choices for data wrangling and analysis.
*API request to NOAA for weather data to modify acceleration parameters for a vehicle over bluetooth CAN bus running realtime?* You can probably do that.
If some of the first parts are things you already know well, feel free to leave and come back for the later parts, or vice versa, I'll share completed parts as we go along which you can access here. You'll probably remember more and learn faster if you type it in yourself however.
- [Part 1 Complete](https://colab.research.google.com/drive/1Q_b9Jr4S-jaHR8VrUPKVRAvz46S-eooC)
- [Part 2 Complete](https://colab.research.google.com/drive/1kmhaXKe6jH_hBkrVyIpQ5-GTpwhbx105)
- [Part 3 Complete](https://colab.research.google.com/drive/1Zn6kbEWTTepkcXLuyn2W51rhC6GQKn-n)
- [Part 4 Complete](https://colab.research.google.com/drive/1Q_b9Jr4S-jaHR8VrUPKVRAvz46S-eooC)
# Part 1 - Colab & Python
Python is an interpreted language, you can run it in an interactive shell line by line in a terminal if you wanted. If you've used MATLAB this is a very similar experience to the interactive shell for that.
Ipython and Jupyter notebooks are open-source web applications that allows you to create and share documents that contain live code, equations, visualizations and narrative text. This is much easier to share and analyze with than just raw code with comments.
## Intro to Colab
Colaboratory is a free Jupyter notebook environment that requires no setup and runs entirely in the cloud.
You can jump around using the table of contents by clicking the little tab arrow on the left side of the screen.
```
seconds_in_a_day = 24 * 60 * 60
seconds_in_a_day
```
To execute the code in the above cell, select it with a click and then either press the play button to the left of the code, or use the keyboard shortcut "Command/Ctrl+Enter".
All cells modify the same global state, so variables that you define by executing a cell can be used in other cells:
```
seconds_in_a_week = 7 * seconds_in_a_day
seconds_in_a_week
```
### Tab-completion and exploring code
Colab provides tab completion to explore attributes of Python objects, as well as to quickly view documentation strings. As an example, first run the following cell to import the [`numpy`](http://www.numpy.org) module.
*Note that we can make an alias `np` so we don't have to type `numpy` all the time*
```
import numpy as np
```
If you now insert your cursor after ``np.random.`` and press **Tab**, you will see the list of available completions within the ``np.random`` submodule.
```
np.random.
```
If you type an open parenthesis followed by the **Tab** key after any function or class in the module, you will see a pop-up of its documentation string:
```
np.random.rand(
```
To open the documentation in a persistent pane at the bottom of your screen, add a **?** after the object or method name and execute the cell using **Shift+Enter**:
```
np.random?
```
## How this relates to Research @ Google
I work in google research on a team specializing in computational photography, which in our case is mostly about the Google Pixel phone line taking great photos. Stuff like [Portrait Mode](https://ai.googleblog.com/2018/11/learning-to-predict-depth-on-pixel-3.html) that blurs out the background and [Night Sight](https://ai.googleblog.com/2018/11/night-sight-seeing-in-dark-on-pixel.html) that lets you take photos in the dark, etc.
There are several other research teams in Google, medical brain is one for example. Co-worker who works on [ML detecting diabetic retinopathy](https://ai.googleblog.com/2018/12/improving-effectiveness-of-diabetic.html) from images of the retina.
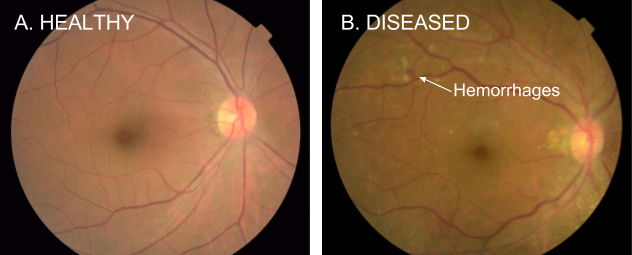
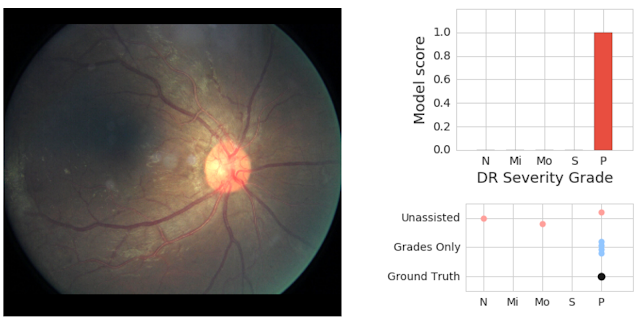
All these algorithms need tons of data. In our team and computational photography, we need lots of images labeled with the relevant features so we can design algorithms and train models with it.
One of the things I built was a ['Frankenphone' data capture system](https://www.cnet.com/news/google-frankenphone-taught-pixel-3-ai-to-take-portrait-photos/) consisting of 5 phones in a fix configuration, that takes photos simultaneously. Using this and computer vision techniques we can estimate the 3D structure of the scene (ie. the true distance of all the pixels in the image in 3D), which we then used to train an ML algorithm to do it automatically from a single camera.

We collected around 50,000 images for our dataset, that's a ton of data, since we store a lot of extra data for each image at around 100-200mb, that comes out to terabytes of data. This ended up becoming a product for Google, as well as a valuable dataset for our research teams.
This relates to todays lesson directly. A lot of our actual analysis and processing of our dataset is done using colab with python, numpy, matplotlib, and a bit of pandas. For example filtering out blurry images from our dataset.

## Python (Intro, Lists, Functions, etc.)
### Lists
```
pressures = [0.273, 0.275, 0.277, 0.275, 0.276]
print('pressures:', pressures)
print('length:', len(pressures))
```
Can use existing functions like `len()` for length, `min/max` for values
```
print('zeroth item of pressures:', pressures[0])
print('fourth item of pressures:', pressures[4])
print('Min pressure:', min(pressures))
print('Max pressure:', max(pressures))
print('Sum of pressures:', sum(pressures))
pressures[0] = 0.265
print('pressures is now:', pressures)
primes = [2, 3, 5]
print('primes is initially:', primes)
primes.append(7)
primes.append(9)
print('primes has become:', primes)
teen_primes = [11, 13, 17, 19]
middle_aged_primes = [37, 41, 43, 47]
print('primes is currently:', primes)
primes.extend(teen_primes)
print('primes has now become:', primes)
primes.append(middle_aged_primes)
print('primes has finally become:', primes)
primes = [2, 3, 5, 7, 9]
print('primes before removing last item:', primes)
del primes[4]
print('primes after removing last item:', primes)
```
Strings to lists and back
```
print('string to list:', list('tin'))
print('list to string:', ''.join(['g', 'o', 'l', 'd']))
list(zip(['San Francisco', 'San Jose', 'Sacramento'], [852469, 1015785, 485199]))
```
### Functions
```
def print_date(year, month, day):
joined = str(year) + '/' + str(month) + '/' + str(day)
# joined = "%04d/%02d/%02d" % (year, month, day)
print(joined)
print_date(1871, 3, 19)
result = print_date(1871, 3, 19)
print('result of call is:', result)
```
### For loops
```
result = []
for i in [1,2,3,4]:
result.append(i**2)
print(result)
```
We can do it in-line as well
```
result = [i**2 for i in [1,2,3,4]]
print(result)
range(4)
for i in range(4):
print(i)
primes = [2, 3, 5]
for p in primes:
squared = p ** 2
cubed = p ** 3
print(p, squared, cubed)
def isEven(n):
return n % 2 == 0
print("6 is even?",isEven(6))
print("7 is even?",isEven(7))
```
### Exercise #1
Write a brute-force function `divisors(n)` that returns some subset of `(2, 3, 5, 7)` for a given integer `n` if `n` is divisible by that number.
Example:
```
divisors(1) -> []
divisors(10) -> [2, 5]
divisors(42) -> [2, 3, 7]
```
**Note:** Python does modulo arithmetic same as MATLAB/R, ex. `36 % 2 = 0`, or `7 % 3 = 1`.
Example:
```python
def isEven(n):
return n % 2 == 0
```
```
def divisors(n):
# Your code here
pass
```
#### Solution
Click below for a solution.
```
def divisors(n):
result = []
for i in [2,3,5,7]:
if n % i == 0:
result.append(i)
return result
```
### Map & defaultdict
Map applies a function to all the items in an input_list. Here is the blueprint:
`map(function_to_apply, list_of_inputs)`
Most of the times we want to pass all the list elements to a function one-by-one and then collect the output. For instance:
```
items = [1, 2, 3, 4, 5]
squared = []
for i in items:
squared.append(i**2)
```
Map allows us to implement this in a much simpler and nicer way:
```
items = [1, 2, 3, 4, 5]
squared = list(map(lambda x: x**2, items))
```
Most of the times we use lambdas with map so I did the same. Instead of a list of inputs we can even have a list of functions!
```
def multiply(x):
return (x*x)
def add(x):
return (x+x)
funcs = [multiply, add]
for i in range(5):
value = list(map(lambda x: x(i), funcs))
print(value)
```
### Dictionaries
Dictionaries are a convenient way to store data for later retrieval by name (key). Keys must be unique, immutable objects, and are typically strings. The values in a dictionary can be anything. For many applications the values are simple types such as integers and strings.
It gets more interesting when the values in a dictionary are collections (lists, dicts, etc.) In this case, the value (an empty list or dict) must be initialized the first time a given key is used. While this is relatively easy to do manually, the defaultdict type automates and simplifies these kinds of operations.
```
car = {
"brand": "Ford",
"model": "Mustang",
"year": 1964
}
print(car)
for key in car:
print(key)
for item in car.items():
print(item)
for key, value in car.items():
print(key,':',value)
```
### Defaultdict
A defaultdict works exactly like a normal dict, but it is initialized with a function (“default factory”) that takes no arguments and provides the default value for a nonexistent key.
A defaultdict will never raise a KeyError. Any key that does not exist gets the value returned by the default factory.
```
from collections import defaultdict
food_list = 'spam spam spam spam spam spam eggs spam'.split()
food_count = defaultdict(int) # default value of int is 0
for food in food_list:
food_count[food] += 1 # increment element's value by 1
print(food_count)
print(food_count['spam'])
city_list = [('TX','Austin'), ('TX','Houston'), ('NY','Albany'), ('NY', 'Syracuse'), ('NY', 'Buffalo'), ('NY', 'Rochester'), ('TX', 'Dallas'), ('CA','Sacramento'), ('CA', 'Palo Alto'), ('GA', 'Atlanta')]
cities_by_state = defaultdict(list)
for state, city in city_list:
cities_by_state[state].append(city)
for state, cities in cities_by_state.items():
print(state, ', '.join(cities))
```
### Filter
As the name suggests, filter creates a list of elements for which a function returns true. Here is a short and concise example:
```
number_list = range(-5, 5)
less_than_zero = list(filter(lambda x: x < 0, number_list))
print(less_than_zero)
```
The filter resembles a for loop but it is a builtin function and faster.
### Importing packages
Python packages are bundles of modules. Modules are basically python scripts containing useful objects/classes/algorithms.
```
# Packages
import numpy as np
import pandas as pd
# Importing a specific module from a package
import matplotlib.pyplot as plt
```
# 15 min break - 10:15am - 10:30am
# Part 2 - Numpy / Matplotlib
## Numpy
### Python lists -> Numpy
Let's consider now what happens when we use a Python data structure that holds many Python objects.
The standard mutable multi-element container in Python is the list.
We can create a list of integers as follows:
```
L = list(range(10))
L
type(L[0])
```
Or, similarly, a list of strings:
```
L2 = [str(c) for c in L]
L2
type(L2[0])
```
Because of Python's dynamic typing, we can even create heterogeneous lists:
```
L3 = [True, "2", 3.0, 4]
[type(item) for item in L3]
```
But this flexibility comes at a cost: to allow these flexible types, each item in the list must contain its own type info, reference count, and other information–that is, each item is a complete Python object.
In the special case that all variables are of the same type, much of this information is redundant: it can be much more efficient to store data in a fixed-type array.
At the implementation level, the array essentially contains a single pointer to one contiguous block of data.
The Python list, on the other hand, contains a pointer to a block of pointers, each of which in turn points to a full Python object like the Python integer we saw earlier.
Again, the advantage of the list is flexibility: because each list element is a full structure containing both data and type information, the list can be filled with data of any desired type.
Fixed-type NumPy-style arrays lack this flexibility, but are much more efficient for storing and manipulating data.
Python lists don't assume operations like one might expect for numerical vectors. Lets say we have a list of integers.
```
a = [1,2,3]
print(a)
```
What if we wanted to add 1 to every value in the array?
```
# a+1 fails
# in-line for loop would work [x + 1 for x in a]
```
### Fixed-Type Arrays in Python
Python offers several different options for storing data in efficient, fixed-type data buffers.
The built-in ``array`` module (available since Python 3.3) can be used to create dense arrays of a uniform type:
```
import array
L = list(range(10))
A = array.array('i', L)
A
```
Here ``'i'`` is a type code indicating the contents are integers.
Much more useful, however, is the ``ndarray`` object of the NumPy package.
While Python's ``array`` object provides efficient storage of array-based data, NumPy adds to this efficient *operations* on that data.
We will explore these operations in later sections; here we'll demonstrate several ways of creating a NumPy array.
We'll start with the standard NumPy import, under the alias ``np``:
```
import numpy as np
```
### Creating Arrays from Python Lists
First, we can use ``np.array`` to create arrays from Python lists:
```
# integer array:
np.array([1, 4, 2, 5, 3])
```
Remember that unlike Python lists, NumPy is constrained to arrays that all contain the same type.
If types do not match, NumPy will upcast if possible (here, integers are up-cast to floating point):
```
np.array([3.14, 4, 2, 3])
```
If we want to explicitly set the data type of the resulting array, we can use the ``dtype`` keyword:
```
np.array([1, 2, 3, 4], dtype='float32')
```
Finally, unlike Python lists, NumPy arrays can explicitly be multi-dimensional; here's one way of initializing a multidimensional array using a list of lists:
```
# nested lists result in multi-dimensional arrays
np.array([range(i, i + 3) for i in [2, 4, 6]])
```
The inner lists are treated as rows of the resulting two-dimensional array.
### Creating Arrays from Scratch
Especially for larger arrays, it is more efficient to create arrays from scratch using routines built into NumPy.
Here are several examples:
```
# Create a length-10 integer array filled with zeros
np.zeros(10, dtype=int)
# Create a 3x5 floating-point array filled with ones
np.ones((3, 5), dtype=float)
# Create a 3x5 array filled with 3.14
np.full((3, 5), 3.14)
# Create an array filled with a linear sequence
# Starting at 0, ending at 20, stepping by 2
# (this is similar to the built-in range() function)
np.arange(0, 20, 2)
# Create an array of five values evenly spaced between 0 and 1
np.linspace(0, 1, 5)
# Create a 3x3 array of uniformly distributed
# random values between 0 and 1
np.random.random((3, 3))
# Create a 3x3 array of normally distributed random values
# with mean 0 and standard deviation 1
np.random.normal(0, 1, (3, 3))
# Create a 3x3 array of random integers in the interval [0, 10)
np.random.randint(0, 10, (3, 3))
# Create a 3x3 identity matrix
np.eye(3)
```
For those of you used to the simple matlab syntax, the following can be used:
```
A = np.matrix('1,1,1 ; 2,4,5 ; 5,7,6')
```
Can also create a bunch of random values, for example a uniform random number from 0 to 1:
```
np.random.random()
np.random.random(10)
```
### Loading CSV data
We can load some data through several mechanisms.
For example, lets get a local CSV text file of state populations over the years. You can see the raw data here: [state-population.csv](https://raw.githubusercontent.com/jakevdp/PythonDataScienceHandbook/master/notebooks/data/state-population.csv)
We can use the terminal directly from colab using `!` and use `curl` to pull the data in, saving to the current directory with the name `state-population.csv`.
```
# Fetch a single <1MB file using the raw GitHub URL.
!curl --remote-name \
-H 'Accept: application/vnd.github.v3.raw' \
--location https://api.github.com/repos/jakevdp/PythonDataScienceHandbook/contents/notebooks/data/state-population.csv
```
#### Load raw text files
We can load the raw text using python, even split it up
```
with open('state-population.csv', 'r') as f:
raw_data_py = f.read()
print(raw_data_py[:100]) # Prints first 1000 characters from the file.
```
This is one big string, we could instead split it up into lines.
```
with open('state-population.csv', 'r') as f:
data_py = f.readlines() # similar to as data_py = f.read().split('\n'), but keeps the '\n'
data_py = [line.strip() for line in data_py]
for line in data_py[:10]:
print(line)
```
Explain `with`, explain `open`, explain `readlines`, explain in-line for loops, explain `strip`, explain slices of lists
#### Load numeric or text files with numpy
We can do a bit better with numpy, though it's mostly used for numeric data, not strings.
```
data_np = np.loadtxt('state-population.csv', dtype='str', delimiter=',')
print(data_np)
```
`<Load numeric columns directly with dtype=int or float>`
Note that even the numeric data is strings, we could split up the columns and convert them to numpy arrays, and build a structure that makes it easy to analyze process. Instead of doing it ourselves, this is basically what Pandas does.
#### Load CSV data using Pandas
If we have a local CSV file, we can load it with `read_csv`
```
data_pd = pd.read_csv('state-population.csv', index_col=0)
data_pd.head()
```
Or we could instead load it directly from a url.
```
data_pd = pd.read_csv('https://raw.githubusercontent.com/jakevdp/PythonDataScienceHandbook/master/notebooks/data/state-population.csv', index_col=0)
data_pd.head()
data_pd.describe()
data_pd.plot.scatter(x='year', y='population');
```
Explain index, unnamed indices, `index_col`, NaN, etc.
## Comparison Operators as ufuncs
In [Computation on NumPy Arrays: Universal Functions](02.03-Computation-on-arrays-ufuncs.ipynb) we introduced ufuncs, and focused in particular on arithmetic operators. We saw that using ``+``, ``-``, ``*``, ``/``, and others on arrays leads to element-wise operations.
NumPy also implements comparison operators such as ``<`` (less than) and ``>`` (greater than) as element-wise ufuncs.
The result of these comparison operators is always an array with a Boolean data type.
All six of the standard comparison operations are available:
```
x = np.array([1, 2, 3, 4, 5])
x < 3 # less than
x > 3 # greater than
x <= 3 # less than or equal
x >= 3 # greater than or equal
x != 3 # not equal
x == 3 # equal
```
It is also possible to do an element-wise comparison of two arrays, and to include compound expressions:
```
(2 * x) == (x ** 2)
```
As in the case of arithmetic operators, the comparison operators are implemented as ufuncs in NumPy; for example, when you write ``x < 3``, internally NumPy uses ``np.less(x, 3)``.
A summary of the comparison operators and their equivalent ufunc is shown here:
| Operator | Equivalent ufunc || Operator | Equivalent ufunc |
|---------------|---------------------||---------------|---------------------|
|``==`` |``np.equal`` ||``!=`` |``np.not_equal`` |
|``<`` |``np.less`` ||``<=`` |``np.less_equal`` |
|``>`` |``np.greater`` ||``>=`` |``np.greater_equal`` |
Just as in the case of arithmetic ufuncs, these will work on arrays of any size and shape.
Here is a two-dimensional example:
```
rng = np.random.RandomState(0)
x = rng.randint(10, size=(3, 4))
x
x < 6
```
In each case, the result is a Boolean array, and NumPy provides a number of straightforward patterns for working with these Boolean results.
## Working with Boolean Arrays
Given a Boolean array, there are a host of useful operations you can do.
We'll work with ``x``, the two-dimensional array we created earlier.
```
print(x)
```
### Counting entries
To count the number of ``True`` entries in a Boolean array, ``np.count_nonzero`` is useful:
```
# how many values less than 6?
np.count_nonzero(x < 6)
```
We see that there are eight array entries that are less than 6.
Another way to get at this information is to use ``np.sum``; in this case, ``False`` is interpreted as ``0``, and ``True`` is interpreted as ``1``:
```
np.sum(x < 6)
```
The benefit of ``sum()`` is that like with other NumPy aggregation functions, this summation can be done along rows or columns as well:
```
# how many values less than 6 in each row?
np.sum(x < 6, axis=1)
```
This counts the number of values less than 6 in each row of the matrix.
If we're interested in quickly checking whether any or all the values are true, we can use (you guessed it) ``np.any`` or ``np.all``:
```
# are there any values greater than 8?
np.any(x > 8)
# are there any values less than zero?
np.any(x < 0)
# are all values less than 10?
np.all(x < 10)
# are all values equal to 6?
np.all(x == 6)
```
``np.all`` and ``np.any`` can be used along particular axes as well. For example:
```
# are all values in each row less than 8?
np.all(x < 8, axis=1)
```
Here all the elements in the first and third rows are less than 8, while this is not the case for the second row.
Finally, a quick warning: as mentioned in [Aggregations: Min, Max, and Everything In Between](02.04-Computation-on-arrays-aggregates.ipynb), Python has built-in ``sum()``, ``any()``, and ``all()`` functions. These have a different syntax than the NumPy versions, and in particular will fail or produce unintended results when used on multidimensional arrays. Be sure that you are using ``np.sum()``, ``np.any()``, and ``np.all()`` for these examples!
## Matplotlib (Intro, visualizing numpy, histograms)
Matplotlib is a multi-platform data visualization library built on NumPy arrays, and designed to work with the broader SciPy stack. If you've used MATLAB, much of the syntax will be familiar.
One of Matplotlib’s most important features is its ability to play well with many operating systems and graphics backends.
Matplotlib supports dozens of backends and output types, which means you can count on it to work regardless of which operating system you are using or which output format you wish.
This cross-platform, everything-to-everyone approach has been one of the great strengths of Matplotlib.
It has led to a large user base, which in turn has led to an active developer base and Matplotlib’s powerful tools and ubiquity within the scientific Python world.
Matplotlib is a bit clunky, there's a lot of newer tools out there that may be more relevant for you, but in general it can do the job and is the most well known.
```
import matplotlib.pyplot as plt
x = np.linspace(0, 10, 100)
fig = plt.figure()
plt.plot(x, np.sin(x), '-')
plt.plot(x, np.cos(x), '--');
```
## Example: Counting Rainy Days
Imagine you have a series of data that represents the amount of precipitation each day for a year in a given city.
For example, here we'll load the daily rainfall statistics for the city of Seattle in 2014, using Pandas (I know, we technically haven't started pandas yet, but we'll get to it soon):
```
import numpy as np
import pandas as pd
# Use pandas to extract rainfall inches as a NumPy array
rainfall = pd.read_csv("https://github.com/jakevdp/PythonDataScienceHandbook/raw/master/notebooks/data/Seattle2014.csv", sep=",")['PRCP'].values
inches = rainfall / 254.0 # 1/10mm -> inches
inches.shape
```
> There's a bit to unpack in how we used pandas to get that data, we'll get to that after lunch.
The array contains 365 values, giving daily rainfall in inches from January 1 to December 31, 2014.
As a first quick visualization, let's look at the histogram of rainy days using Matplotlib.
```
plt.hist(inches, 40);
plt.title('Histogram of inches of rain for Seattle over 2014')
plt.xlabel('Inches');
```
# 1 hr lunch break - 12pm - 1pm
# Part 3 - Pandas
Shamelessly using [google's tutorial](https://colab.research.google.com/notebooks/mlcc/intro_to_pandas.ipynb).
## Intro to pandas
**Learning Objectives:**
* Gain an introduction to the `DataFrame` and `Series` data structures of the *pandas* library
* Access and manipulate data within a `DataFrame` and `Series`
* Import CSV data into a *pandas* `DataFrame`
* Reindex a `DataFrame` to shuffle data
[*pandas*](http://pandas.pydata.org/) is a column-oriented data analysis API. It's a great tool for handling and analyzing input data, and many ML frameworks support *pandas* data structures as inputs.
Although a comprehensive introduction to the *pandas* API would span many pages, the core concepts are fairly straightforward, and we'll present them below. For a more complete reference, the [*pandas* docs site](http://pandas.pydata.org/pandas-docs/stable/index.html) contains extensive documentation and many tutorials.
## Basic Concepts
The following line imports the *pandas* API:
```
import pandas as pd
```
The primary data structures in *pandas* are implemented as two classes:
* **`DataFrame`**, which you can imagine as a relational data table, with rows and named columns.
* **`Series`**, which is a single column. A `DataFrame` contains one or more `Series` and a name for each `Series`.
The data frame is a commonly used abstraction for data manipulation. Similar implementations exist in [Spark](https://spark.apache.org/) and [R](https://www.r-project.org/about.html).
One way to create a `Series` is to construct a `Series` object. For example:
```
pd.Series(['San Francisco', 'San Jose', 'Sacramento'])
```
`DataFrame` objects can be created by passing a `dict` mapping `string` column names to their respective `Series`. If the `Series` don't match in length, missing values are filled with special [NA/NaN](http://pandas.pydata.org/pandas-docs/stable/missing_data.html) values. Example:
```
city_names = pd.Series(['San Francisco', 'San Jose', 'Sacramento'])
population = pd.Series([852469, 1015785, 485199])
city_pop = pd.DataFrame({ 'City name': city_names, 'Population': population })
print(city_pop)
```
But most of the time, you load an entire file into a `DataFrame`. The following example loads a file with California housing data. Run the following cell to load the data and create feature definitions:
```
california_housing_dataframe = pd.read_csv("https://download.mlcc.google.com/mledu-datasets/california_housing_train.csv", sep=",")
california_housing_dataframe.describe()
```
The example above used `DataFrame.describe` to show interesting statistics about a `DataFrame`. Another useful function is `DataFrame.head`, which displays the first few records of a `DataFrame`:
```
california_housing_dataframe.head()
```
Another powerful feature of *pandas* is graphing. For example, `DataFrame.hist` lets you quickly study the distribution of values in a column:
```
california_housing_dataframe.hist('housing_median_age')
```
## Accessing Data
You can access `DataFrame` data using familiar Python dict/list operations:
```
cities = pd.DataFrame({ 'City name': city_names, 'Population': population })
print(type(cities['City name']))
cities['City name']
print(type(cities['City name'][1]))
cities['City name'][1]
print(type(cities[0:2]))
cities[0:2]
```
In addition, *pandas* provides an extremely rich API for advanced [indexing and selection](http://pandas.pydata.org/pandas-docs/stable/indexing.html) that is too extensive to be covered here.
## Manipulating Data
You may apply Python's basic arithmetic operations to `Series`. For example:
```
population / 1000.
import numpy as np
np.log(population)
```
For more complex single-column transformations, you can use `Series.apply`. Like the Python [map function](https://docs.python.org/2/library/functions.html#map),
`Series.apply` accepts as an argument a [lambda function](https://docs.python.org/2/tutorial/controlflow.html#lambda-expressions), which is applied to each value.
The example below creates a new `Series` that indicates whether `population` is over one million:
```
population.apply(lambda val: val > 1000000)
```
Modifying `DataFrames` is also straightforward. For example, the following code adds two `Series` to an existing `DataFrame`:
```
cities['Area square miles'] = pd.Series([46.87, 176.53, 97.92])
cities['Population density'] = cities['Population'] / cities['Area square miles']
cities
```
## Exercise #1
Modify the `cities` table by adding a new boolean column that is True if and only if *both* of the following are True:
* The city is named after a saint.
* The city has an area greater than 50 square miles.
**Note:** Boolean `Series` are combined using the bitwise, rather than the traditional boolean, operators. For example, when performing *logical and*, use `&` instead of `and`.
**Hint:** "San" in Spanish means "saint."
```
# Your code here
```
### Solution
Click below for a solution.
```
cities['Is wide and has saint name'] = (cities['Area square miles'] > 50) & cities['City name'].apply(lambda name: name.startswith('San'))
cities
```
## Indexes
Both `Series` and `DataFrame` objects also define an `index` property that assigns an identifier value to each `Series` item or `DataFrame` row.
By default, at construction, *pandas* assigns index values that reflect the ordering of the source data. Once created, the index values are stable; that is, they do not change when data is reordered.
```
city_names.index
cities.index
```
Call `DataFrame.reindex` to manually reorder the rows. For example, the following has the same effect as sorting by city name:
```
cities.reindex([2, 0, 1])
```
Reindexing is a great way to shuffle (randomize) a `DataFrame`. In the example below, we take the index, which is array-like, and pass it to NumPy's `random.permutation` function, which shuffles its values in place. Calling `reindex` with this shuffled array causes the `DataFrame` rows to be shuffled in the same way.
Try running the following cell multiple times!
```
cities.reindex(np.random.permutation(cities.index))
```
For more information, see the [Index documentation](http://pandas.pydata.org/pandas-docs/stable/indexing.html#index-objects).
## Exercise #2
The `reindex` method allows index values that are not in the original `DataFrame`'s index values. Try it and see what happens if you use such values! Why do you think this is allowed?
```
# Your code here
```
### Solution
Click below for the solution.
If your `reindex` input array includes values not in the original `DataFrame` index values, `reindex` will add new rows for these "missing" indices and populate all corresponding columns with `NaN` values:
```
cities.reindex([0, 4, 5, 2])
```
This behavior is desirable because indexes are often strings pulled from the actual data (see the [*pandas* reindex
documentation](http://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.reindex.html) for an example
in which the index values are browser names).
In this case, allowing "missing" indices makes it easy to reindex using an external list, as you don't have to worry about
sanitizing the input.
# 15 min break - 2:15pm - 2:30pm
# Part 4 - Sklearn (Scikit-Learn)
Shameless borrowed from [google's tutorial](https://colab.research.google.com/github/jakevdp/PythonDataScienceHandbook/blob/master/notebooks/05.02-Introducing-Scikit-Learn.ipynb).
There are several Python libraries which provide solid implementations of a range of machine learning algorithms.
One of the best known is [Scikit-Learn](http://scikit-learn.org), a package that provides efficient versions of a large number of common algorithms.
Scikit-Learn is characterized by a clean, uniform, and streamlined API, as well as by very useful and complete online documentation.
A benefit of this uniformity is that once you understand the basic use and syntax of Scikit-Learn for one type of model, switching to a new model or algorithm is very straightforward.
This section provides an overview of the Scikit-Learn API; a solid understanding of these API elements will form the foundation for understanding the deeper practical discussion of machine learning algorithms and approaches in the following chapters.
We will start by covering *data representation* in Scikit-Learn, followed by covering the *Estimator* API, and finally go through a more interesting example of using these tools for exploring a set of images of hand-written digits.
## Data Representation in Scikit-Learn
Machine learning is about creating models from data: for that reason, we'll start by discussing how data can be represented in order to be understood by the computer.
The best way to think about data within Scikit-Learn is in terms of tables of data.
### Data as table
A basic table is a two-dimensional grid of data, in which the rows represent individual elements of the dataset, and the columns represent quantities related to each of these elements.
For example, consider the [Iris dataset](https://en.wikipedia.org/wiki/Iris_flower_data_set), famously analyzed by Ronald Fisher in 1936.
We can download this dataset in the form of a Pandas ``DataFrame`` using the [seaborn](http://seaborn.pydata.org/) library:
```
import seaborn as sns
iris = sns.load_dataset('iris')
iris.head()
```
Here each row of the data refers to a single observed flower, and the number of rows is the total number of flowers in the dataset.
In general, we will refer to the rows of the matrix as *samples*, and the number of rows as ``n_samples``.
Likewise, each column of the data refers to a particular quantitative piece of information that describes each sample.
In general, we will refer to the columns of the matrix as *features*, and the number of columns as ``n_features``.
#### Features matrix
This table layout makes clear that the information can be thought of as a two-dimensional numerical array or matrix, which we will call the *features matrix*.
By convention, this features matrix is often stored in a variable named ``X``.
The features matrix is assumed to be two-dimensional, with shape ``[n_samples, n_features]``, and is most often contained in a NumPy array or a Pandas ``DataFrame``, though some Scikit-Learn models also accept SciPy sparse matrices.
The samples (i.e., rows) always refer to the individual objects described by the dataset.
For example, the sample might be a flower, a person, a document, an image, a sound file, a video, an astronomical object, or anything else you can describe with a set of quantitative measurements.
The features (i.e., columns) always refer to the distinct observations that describe each sample in a quantitative manner.
Features are generally real-valued, but may be Boolean or discrete-valued in some cases.
#### Target array
In addition to the feature matrix ``X``, we also generally work with a *label* or *target* array, which by convention we will usually call ``y``.
The target array is usually one dimensional, with length ``n_samples``, and is generally contained in a NumPy array or Pandas ``Series``.
The target array may have continuous numerical values, or discrete classes/labels.
While some Scikit-Learn estimators do handle multiple target values in the form of a two-dimensional, ``[n_samples, n_targets]`` target array, we will primarily be working with the common case of a one-dimensional target array.
Often one point of confusion is how the target array differs from the other features columns. The distinguishing feature of the target array is that it is usually the quantity we want to *predict from the data*: in statistical terms, it is the dependent variable.
For example, in the preceding data we may wish to construct a model that can predict the species of flower based on the other measurements; in this case, the ``species`` column would be considered the target array.
With this target array in mind, we can use Seaborn (see [Visualization With Seaborn](04.14-Visualization-With-Seaborn.ipynb)) to conveniently visualize the data:
```
import seaborn as sns; sns.set()
sns.pairplot(iris, hue='species', height=1.5);
```
For use in Scikit-Learn, we will extract the features matrix and target array from the ``DataFrame``, which we can do using some of the Pandas ``DataFrame`` operations discussed in the [Chapter 3](03.00-Introduction-to-Pandas.ipynb):
```
X_iris = iris.drop('species', axis=1)
X_iris.shape
y_iris = iris['species']
y_iris.shape
```
To summarize, the expected layout of features and target values is visualized in the following diagram:
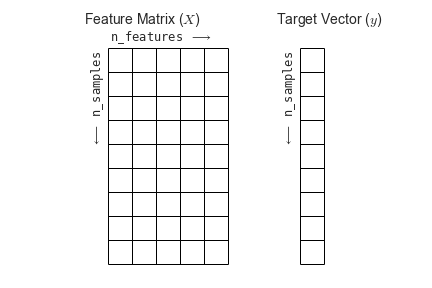
With this data properly formatted, we can move on to consider the *estimator* API of Scikit-Learn:
## Scikit-Learn's Estimator API
The Scikit-Learn API is designed with the following guiding principles in mind, as outlined in the [Scikit-Learn API paper](http://arxiv.org/abs/1309.0238):
- *Consistency*: All objects share a common interface drawn from a limited set of methods, with consistent documentation.
- *Inspection*: All specified parameter values are exposed as public attributes.
- *Limited object hierarchy*: Only algorithms are represented by Python classes; datasets are represented
in standard formats (NumPy arrays, Pandas ``DataFrame``s, SciPy sparse matrices) and parameter
names use standard Python strings.
- *Composition*: Many machine learning tasks can be expressed as sequences of more fundamental algorithms,
and Scikit-Learn makes use of this wherever possible.
- *Sensible defaults*: When models require user-specified parameters, the library defines an appropriate default value.
In practice, these principles make Scikit-Learn very easy to use, once the basic principles are understood.
Every machine learning algorithm in Scikit-Learn is implemented via the Estimator API, which provides a consistent interface for a wide range of machine learning applications.
### Basics of the API
Most commonly, the steps in using the Scikit-Learn estimator API are as follows
(we will step through a handful of detailed examples in the sections that follow).
1. Choose a class of model by importing the appropriate estimator class from Scikit-Learn.
2. Choose model hyperparameters by instantiating this class with desired values.
3. Arrange data into a features matrix and target vector following the discussion above.
4. Fit the model to your data by calling the ``fit()`` method of the model instance.
5. Apply the Model to new data:
- For supervised learning, often we predict labels for unknown data using the ``predict()`` method.
- For unsupervised learning, we often transform or infer properties of the data using the ``transform()`` or ``predict()`` method.
We will now step through several simple examples of applying supervised and unsupervised learning methods.
### Supervised learning example: Simple linear regression
As an example of this process, let's consider a simple linear regression—that is, the common case of fitting a line to $(x, y)$ data.
We will use the following simple data for our regression example:
```
import matplotlib.pyplot as plt
import numpy as np
rng = np.random.RandomState(42)
x = 10 * rng.rand(50)
y = 2 * x - 1 + rng.randn(50)
plt.scatter(x, y);
```
With this data in place, we can use the recipe outlined earlier. Let's walk through the process:
#### 1. Choose a class of model
In Scikit-Learn, every class of model is represented by a Python class.
So, for example, if we would like to compute a simple linear regression model, we can import the linear regression class:
```
from sklearn.linear_model import LinearRegression
```
Note that other more general linear regression models exist as well; you can read more about them in the [``sklearn.linear_model`` module documentation](http://Scikit-Learn.org/stable/modules/linear_model.html).
#### 2. Choose model hyperparameters
An important point is that *a class of model is not the same as an instance of a model*.
Once we have decided on our model class, there are still some options open to us.
Depending on the model class we are working with, we might need to answer one or more questions like the following:
- Would we like to fit for the offset (i.e., *y*-intercept)?
- Would we like the model to be normalized?
- Would we like to preprocess our features to add model flexibility?
- What degree of regularization would we like to use in our model?
- How many model components would we like to use?
These are examples of the important choices that must be made *once the model class is selected*.
These choices are often represented as *hyperparameters*, or parameters that must be set before the model is fit to data.
In Scikit-Learn, hyperparameters are chosen by passing values at model instantiation.
We will explore how you can quantitatively motivate the choice of hyperparameters in [Hyperparameters and Model Validation](05.03-Hyperparameters-and-Model-Validation.ipynb).
For our linear regression example, we can instantiate the ``LinearRegression`` class and specify that we would like to fit the intercept using the ``fit_intercept`` hyperparameter:
```
model = LinearRegression(fit_intercept=True)
model
```
Keep in mind that when the model is instantiated, the only action is the storing of these hyperparameter values.
In particular, we have not yet applied the model to any data: the Scikit-Learn API makes very clear the distinction between *choice of model* and *application of model to data*.
#### 3. Arrange data into a features matrix and target vector
Previously we detailed the Scikit-Learn data representation, which requires a two-dimensional features matrix and a one-dimensional target array.
Here our target variable ``y`` is already in the correct form (a length-``n_samples`` array), but we need to massage the data ``x`` to make it a matrix of size ``[n_samples, n_features]``.
In this case, this amounts to a simple reshaping of the one-dimensional array:
```
X = x[:, np.newaxis]
X.shape
```
#### 4. Fit the model to your data
Now it is time to apply our model to data.
This can be done with the ``fit()`` method of the model:
```
model.fit(X, y)
```
This ``fit()`` command causes a number of model-dependent internal computations to take place, and the results of these computations are stored in model-specific attributes that the user can explore.
In Scikit-Learn, by convention all model parameters that were learned during the ``fit()`` process have trailing underscores; for example in this linear model, we have the following:
```
model.coef_
model.intercept_
```
These two parameters represent the slope and intercept of the simple linear fit to the data.
Comparing to the data definition, we see that they are very close to the input slope of 2 and intercept of -1.
One question that frequently comes up regards the uncertainty in such internal model parameters.
In general, Scikit-Learn does not provide tools to draw conclusions from internal model parameters themselves: interpreting model parameters is much more a *statistical modeling* question than a *machine learning* question.
Machine learning rather focuses on what the model *predicts*.
If you would like to dive into the meaning of fit parameters within the model, other tools are available, including the [Statsmodels Python package](http://statsmodels.sourceforge.net/).
#### 5. Predict labels for unknown data
Once the model is trained, the main task of supervised machine learning is to evaluate it based on what it says about new data that was not part of the training set.
In Scikit-Learn, this can be done using the ``predict()`` method.
For the sake of this example, our "new data" will be a grid of *x* values, and we will ask what *y* values the model predicts:
```
xfit = np.linspace(-1, 11)
```
As before, we need to coerce these *x* values into a ``[n_samples, n_features]`` features matrix, after which we can feed it to the model:
```
Xfit = xfit[:, np.newaxis]
yfit = model.predict(Xfit)
```
Finally, let's visualize the results by plotting first the raw data, and then this model fit:
```
plt.scatter(x, y)
plt.plot(xfit, yfit);
```
Typically the efficacy of the model is evaluated by comparing its results to some known baseline, as we will see in the next example
### Supervised learning example: Iris classification
Let's take a look at another example of this process, using the Iris dataset we discussed earlier.
Our question will be this: given a model trained on a portion of the Iris data, how well can we predict the remaining labels?
For this task, we will use an extremely simple generative model known as Gaussian naive Bayes, which proceeds by assuming each class is drawn from an axis-aligned Gaussian distribution (see [In Depth: Naive Bayes Classification](05.05-Naive-Bayes.ipynb) for more details).
Because it is so fast and has no hyperparameters to choose, Gaussian naive Bayes is often a good model to use as a baseline classification, before exploring whether improvements can be found through more sophisticated models.
We would like to evaluate the model on data it has not seen before, and so we will split the data into a *training set* and a *testing set*.
This could be done by hand, but it is more convenient to use the ``train_test_split`` utility function:
```
from sklearn.model_selection import train_test_split
Xtrain, Xtest, ytrain, ytest = train_test_split(X_iris, y_iris,
random_state=1)
```
With the data arranged, we can follow our recipe to predict the labels:
```
from sklearn.naive_bayes import GaussianNB # 1. choose model class
model = GaussianNB() # 2. instantiate model
model.fit(Xtrain, ytrain) # 3. fit model to data
y_model = model.predict(Xtest) # 4. predict on new data
```
Finally, we can use the ``accuracy_score`` utility to see the fraction of predicted labels that match their true value:
```
from sklearn.metrics import accuracy_score
accuracy_score(ytest, y_model)
```
With an accuracy topping 97%, we see that even this very naive classification algorithm is effective for this particular dataset!
### Unsupervised learning example: Iris dimensionality
As an example of an unsupervised learning problem, let's take a look at reducing the dimensionality of the Iris data so as to more easily visualize it.
Recall that the Iris data is four dimensional: there are four features recorded for each sample.
The task of dimensionality reduction is to ask whether there is a suitable lower-dimensional representation that retains the essential features of the data.
Often dimensionality reduction is used as an aid to visualizing data: after all, it is much easier to plot data in two dimensions than in four dimensions or higher!
Here we will use principal component analysis (PCA; see [In Depth: Principal Component Analysis](05.09-Principal-Component-Analysis.ipynb)), which is a fast linear dimensionality reduction technique.
We will ask the model to return two components—that is, a two-dimensional representation of the data.
Following the sequence of steps outlined earlier, we have:
```
from sklearn.decomposition import PCA # 1. Choose the model class
model = PCA(n_components=2) # 2. Instantiate the model with hyperparameters
model.fit(X_iris) # 3. Fit to data. Notice y is not specified!
X_2D = model.transform(X_iris) # 4. Transform the data to two dimensions
```
Now let's plot the results. A quick way to do this is to insert the results into the original Iris ``DataFrame``, and use Seaborn's ``lmplot`` to show the results:
```
iris['PCA1'] = X_2D[:, 0]
iris['PCA2'] = X_2D[:, 1]
sns.lmplot("PCA1", "PCA2", hue='species', data=iris, fit_reg=False);
```
We see that in the two-dimensional representation, the species are fairly well separated, even though the PCA algorithm had no knowledge of the species labels!
This indicates to us that a relatively straightforward classification will probably be effective on the dataset, as we saw before.
### Unsupervised learning: Iris clustering
Let's next look at applying clustering to the Iris data.
A clustering algorithm attempts to find distinct groups of data without reference to any labels.
Here we will use a powerful clustering method called a Gaussian mixture model (GMM), discussed in more detail in [In Depth: Gaussian Mixture Models](05.12-Gaussian-Mixtures.ipynb).
A GMM attempts to model the data as a collection of Gaussian blobs.
We can fit the Gaussian mixture model as follows:
```
from sklearn.mixture import GaussianMixture # 1. Choose the model class
model = GaussianMixture(n_components=3,
covariance_type='full') # 2. Instantiate the model with hyperparameters
model.fit(X_iris) # 3. Fit to data. Notice y is not specified!
y_gmm = model.predict(X_iris) # 4. Determine cluster labels
```
As before, we will add the cluster label to the Iris ``DataFrame`` and use Seaborn to plot the results:
```
iris['cluster'] = y_gmm
sns.lmplot("PCA1", "PCA2", data=iris, hue='species',
col='cluster', fit_reg=False);
```
By splitting the data by cluster number, we see exactly how well the GMM algorithm has recovered the underlying label: the *setosa* species is separated perfectly within cluster 0, while there remains a small amount of mixing between *versicolor* and *virginica*.
This means that even without an expert to tell us the species labels of the individual flowers, the measurements of these flowers are distinct enough that we could *automatically* identify the presence of these different groups of species with a simple clustering algorithm!
This sort of algorithm might further give experts in the field clues as to the relationship between the samples they are observing.
# Unorganized Extras
Lets make our own dataset with two features A and B. They'll be roughly two 2D normal distributions with a slight separation
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# Create some arbitrary data.
def getSomeData(N=500, cut_percent=0.3, seed=1234):
np.random.seed(seed)
cut = int(cut_percent*N)
# Create two distributions slightly spaced apart
ax1 = np.random.normal(loc=-4, scale=2, size=cut)
ay1 = ax1 * 5 + np.random.normal(loc=0, scale=10, size=ax1.size)
ax2 = np.random.normal(loc=0, scale=2, size=(N-cut))
ay2 = ax2 * 5 - 50 + np.random.normal(loc=0, scale=10, size=ax2.size)
Ax = np.concatenate([ax1, ax2], axis=0)
Ay = np.concatenate([ay1, ay2], axis=0)
data = np.stack([Ax, Ay], axis=1)
# Label data as 0 before the cut, and 1 after.
labels = np.hstack([np.zeros(cut), np.ones(N-cut)]).astype(np.bool)
# Shuffle array so values aren't in order any more.
neworder = np.arange(N)
np.random.shuffle(neworder)
data = data[neworder,:]
labels = labels[neworder]
return data, labels
data, labels = getSomeData()
plt.figure(figsize=(15,15))
plt.subplot(221)
plt.hist(data[:,0]);
plt.title('Histogram of A')
plt.subplot(222)
plt.scatter(data[:,0], data[:,1]);
plt.title('Scatter of A and B')
plt.xlabel('A')
plt.ylabel('B');
plt.subplot(223)
plt.hist(data[labels,0],color='g');
plt.hist(data[~labels,0],color='r');
plt.title('Labeled Histogram of A')
plt.subplot(224)
plt.scatter(data[labels,0], data[labels,1], c='g');
plt.scatter(data[~labels,0], data[~labels,1], c='r');
plt.title('Labeled Scatter of A and B')
plt.xlabel('A')
plt.ylabel('B');
```
### sklearn KMeans
```
from sklearn.cluster import KMeans
from sklearn.preprocessing import StandardScaler
X = StandardScaler().fit_transform(data)
labels_kmeans = KMeans(n_clusters=2,random_state=1234).fit_predict(X).astype(np.bool)
plt.figure(figsize=(20,5))
plt.subplot(131)
plt.scatter(data[:,0], data[:,1]);
plt.title('Given')
plt.subplot(132)
plt.scatter(data[labels_kmeans,0], data[labels_kmeans,1], c='r');
plt.scatter(data[~labels_kmeans,0], data[~labels_kmeans,1], c='g');
plt.title('KMeans Clustered')
plt.subplot(133)
plt.scatter(data[labels,0], data[labels,1], c='r');
plt.scatter(data[~labels,0], data[~labels,1], c='g');
plt.xlabel('A')
plt.ylabel('B');
plt.title('Ground Truth');
```
### Image manipulation
Extra
```
import scipy.misc
from scipy import ndimage
face = scipy.misc.face(gray=True)
plt.figure(figsize=(10,5))
plt.subplot(121)
plt.imshow(face, cmap='gray');
# Like matlab's imgaussfilt.
blurred = ndimage.gaussian_filter(face, sigma=3)
print("Normal:")
dx, dy = scipy.gradient(face)
grad_mag = np.sum(dx*dx + dy*dy)
print(grad_mag / (face.shape[0] * face.shape[1]))
print(np.std(face[:]))
plt.subplot(122)
plt.imshow(blurred, cmap='gray')
print("Blurry:")
dx, dy = scipy.gradient(blurred)
grad_mag = np.sum(dx*dx + dy*dy)
print(grad_mag / (face.shape[0] * face.shape[1]))
print(np.std(blurred[:]))
```
| github_jupyter |
<h1 style="color:red; text-align:center">Webscraping</h1>
[The Notebook](https://colab.research.google.com/drive/1TNNpSnDaVnCvN9vyw3_RsFqN9NOu2xjy?authuser=3#scrollTo=MSTFHCbdUZeO)
Hosted on Colab
# Coding challenge
## Leet-code
<h1 style="color:blue; text-align:center">Two Sums</h1>
Given an array of integers nums and an integer target, return indices of the two numbers such that they add up to target.
You may assume that each input would have exactly one solution, and you may not use the same element twice.
You can return the answer in any order.
Input: nums = [2,7,11,15], target = 9
Output: [0,1]
Output: Because nums[0] + nums[1] == 9, we return [0, 1].
<h1 style="color:red;text-align:center">Two Sums Solution</h1>
```
class Solution:
def twoSum(self, nums, target):
"""
:type nums: List[int]
:type target: int
:rtype: List[int]
"""
h = {}
for i, num in enumerate(nums):
n = target - num
if n not in h:
h[num] = i
else:
return [h[n], i]
%whos
my_solution = Solution()
my_solution.twoSum([1,2,3], 3)
second_solution = Solution()
%whos
```
<h1 style="color:blue; text-align:center">Factorial</h1>
In mathematics, the factorial of a non-negative integer n, denoted by n!, is the product of all positive integers less than or equal to n. For example: 5! = 5 * 4 * 3 * 2 * 1 = 120. By convention the value of 0! is 1.
Write a function to calculate factorial for a given input. If input is below 0 or above 12
5! = 5 * 4 * 3 * 2 * 1 = 120
<h1 style="color:red; text-align:center">Factorial Solution</h1>
```
def factorial(n):
if n < 0 or n > 12:
raise ValueError
return 1 if n <= 1 else n*factorial(n-1)
```
## Exception Handling
## ask for permission than forgiveness;
<h1 style="color:blue; text-align:center">Pangram</h1>
A pangram is a sentence that contains every single letter of the alphabet at least once. For example, the sentence "The quick brown fox jumps over the lazy dog" is a pangram, because it uses the letters A-Z at least once (case is irrelevant).
<h1 style="color:red; text-align:center">Pangram</h1>
```
import string
def is_pangram(s):
s = s.lower()
for char in 'abcdefghijklmnopqrstuvwxyz':
if char not in s:
return False
return True
def is_pangram(input_string):
"""Returns False if the input string is not a pangram else returns True"""
return False if [char for char in string.ascii_lowercase if not char in input_string.lower()] else True
print(string.ascii_lowercase)
```
| github_jupyter |
# Stock Forecast Using Single Denoised Data
This example is going to demonstrate how to use time_seris_transform to denoise data prepare lag/lead features. In this exmple, there are four steps
1. prepare raw and denoise data
2. make lag/lead feature
3. Sklearn Model & Visualize Test
## Prepare raw data
In this example, we use Stock_Extractor to get an single stock data from yahoo finance.
Subsequently, we directly denoise the data using make_techinical_indicator funciton. This function is a wrapper which can take function object and transform the traget column.In time_series_transform.transform_core_api.util, there are different pre-made fucntion for feature engineering such as geometric moving average, wavelet transformation, or fast fourier transformation.
```
import pandas as pd
import seaborn as sns
import time_series_transform as tst
from time_series_transform.transform_core_api.util import wavelet_denoising
stock = tst.Stock_Extractor('googl','yahoo').get_stock_period('5y') # get period of stock
df = stock.make_technical_indicator("Close","Close_Wavelet",wavelet_denoising,wavelet='haar').dataFrame # denoise data
stock.plot(["Close","Close_Wavelet"])
```
## Make lag/lead features
After we prepared the data, we can use Pandas_Time_Series_Panel_Dataset to make lag or lead data as feature or target values for machine learning models. In this example, we only make 30-day lags data for Close data with Wavelet transformation, and only 1 step forward data as label.
```
panel_trans = tst.Pandas_Time_Series_Panel_Dataset(df)
panel_trans = panel_trans.make_slide_window('Date',30,['Close_Wavelet'])
panel_trans = panel_trans.make_lead_column('Date','Close',1)
df = panel_trans.df
df = df.drop(['Open', 'High', 'Low', 'Close', 'Volume', 'Dividends',
'Stock Splits', 'Close_Wavelet', 'symbol'],axis =1).dropna()
df = df.sort_values('Date')
print(df.head())
```
## Machine learning model & visualize test results
After we pre-process the data, we can use sklearn to predict the value. In this example, we use Random Forest with Random Search Tuner to prepare our model.
```
from sklearn.model_selection import TimeSeriesSplit,RandomizedSearchCV
from sklearn.preprocessing import StandardScaler
from sklearn.pipeline import Pipeline
from sklearn.decomposition import PCA
from sklearn.ensemble import RandomForestRegressor
train = df[df.Date<'2020-07-01']
test = df[df.Date >= '2020-07-01']
rf = RandomForestRegressor(n_jobs =1,)
haparms = [{
"n_estimators":np.arange(100,200,10,dtype=int),
"min_samples_split":np.arange(2,40,1,dtype=int),
"min_samples_leaf":np.arange(2,40,1,dtype=int),
}]
randomSearch = RandomizedSearchCV(
rf,
haparms,5,
cv = TimeSeriesSplit(3),
n_jobs =1
)
trainX= train.drop(['Date','Close_lead1'],axis = 1)
trainY = train.Close_lead1
randomSearch.fit(trainX,trainY)
test= test.sort_values('Date')
testX= test.drop(['Date','Close_lead1'],axis = 1)
testY = test.Close_lead1
test_prd = randomSearch.predict(testX)
train_prd = randomSearch.predict(trainX)
compareFrame = pd.DataFrame({'train_prd':train_prd,'train_real':train.Close_lead1.tolist()},index = train.Date)
compareFrame_test = pd.DataFrame({'test_prd':test_prd,'test_real':test.Close_lead1.tolist()},index = test.Date)
compareFrame = compareFrame.append(compareFrame_test)
compareFrame['Date'] = compareFrame.index
compareFrame.index = list(range(1,len(compareFrame)+1))
sns.lineplot(data= [
compareFrame[compareFrame.Date >'2020-04-01'].train_real,
compareFrame[compareFrame.Date >='2020-07-01'].test_real,
compareFrame[compareFrame.Date >'2020-04-01'].train_prd,
compareFrame[compareFrame.Date >='2020-07-01'].test_prd,
])
```
# Stock Forecast Using Multiple Stock Features
This example is going to use a set of stocks to predict one stock. This example also implments different algorithms for feature engineering. Furthermore, this example will show how to implment functional object in Pandas_Time_Series_Panel_Dataset class.
There are three steps in this example
1. prepare stocks data through Portfolio_Extractor api and transform data
2. making lag/lead data and other feature engineering function using Pandas_Time_Series_Panel_Dataset
3. sklearn model and visualize test results
## Prepare stocks data and transformation
Portfolio_Extractor takes a list of stock ticks, and return an Portfolio objects. In this example, we prepare 5 year of data. Subsequently, we transform Close, Open, High, Low values using wavelet, fast fourier, and geometric moving average.
```
import time_series_transform as tst
from time_series_transform.transform_core_api.util import wavelet_denoising,rfft_transform,geometric_ma
port = tst.Portfolio_Extractor(['googl','aapl','fb'],'yahoo').get_portfolio_period('5y')
colList = ['Close','Open','High','Low']
denoiseCol = []
for col in colList:
port.make_technical_indicator(col,f'{col}_wavelet',wavelet_denoising,wavelet='haar')
port.make_technical_indicator(col,f'{col}_fft',rfft_transform,threshold=0.001)
port.make_technical_indicator(col,f'{col}_gma20',geometric_ma,windowSize=20)
port.make_technical_indicator(col,f'{col}_gma200',geometric_ma,windowSize=200)
denoiseCol.extend([f'{col}_wavelet',f'{col}_fft',f'{col}_gma200',f'{col}_gma20'])
df = port.get_portfolio_dataFrame()
print(df.head())
```
## Lag/lead data and other feature engineering
Here, we create an moving standard deviation feature and make 20-day-lag with denoised data from previous step. In order to create the proper feature, we have to be aware of its category. That means that we have to group by each stocks. After making new feature, we can create lag data and expand it by its category.
To manipulate the data using transform_dataFrame function, the first argument of the function must be the data array/list, and the output must be numpy array or list.
```
df = df.drop(['Volume','Dividends','Stock Splits'],axis =1)
panel_transform = tst.Pandas_Time_Series_Panel_Dataset(df)
import pandas as pd
def moving_std (arr,windowSize):
res = pd.Series(arr).rolling(windowSize).std()
return res
panel_transform = panel_transform.transform_dataFrame('Close','Close_std','Date','symbol',moving_std,windowSize = 10)
panel_transform = panel_transform.make_slide_window(
indexCol = 'Date',
windowSize = 20,
colList = denoiseCol+['Close_std'],
groupby = 'symbol')
panel_transform = panel_transform.expand_dataFrame_by_category('Date','symbol')
print(panel_transform.df.head())
panel_transform = panel_transform.make_lead_column('Date','Close_googl',1)
df = panel_transform.df
df = df.dropna()
from sklearn.model_selection import TimeSeriesSplit,RandomizedSearchCV
from sklearn.preprocessing import StandardScaler
from sklearn.pipeline import Pipeline
from sklearn.decomposition import PCA
from sklearn.ensemble import RandomForestRegressor
import numpy as np
import seaborn as sns
train = df[df.Date<'2020-07-01']
test = df[df.Date >= '2020-07-01']
rf = RandomForestRegressor(n_jobs =1)
haparms = [{
"n_estimators":np.arange(100,200,10,dtype=int),
"min_samples_split":np.arange(2,40,1,dtype=int),
"min_samples_leaf":np.arange(2,40,1,dtype=int),
}]
randomSearch = RandomizedSearchCV(
rf,
haparms,10,
cv = TimeSeriesSplit(3),
n_jobs =1
)
train = train.sort_values('Date')
trainX= train.drop(['Date','Close_googl_lead1'],axis = 1)
trainY = train.Close_googl_lead1
randomSearch.fit(trainX,trainY)
train_prd = randomSearch.predict(trainX)
test= test.sort_values('Date')
testX= test.drop(['Date','Close_googl_lead1'],axis = 1)
testY = test.Close_googl_lead1
test_prd = randomSearch.predict(testX)
compareFrame = pd.DataFrame({'train_prd':train_prd,'train_real':train.Close_googl_lead1.tolist()},index = train.Date)
compareFrame_test = pd.DataFrame({'test_prd':test_prd,'test_real':test.Close_googl_lead1.tolist()},index = test.Date)
compareFrame = compareFrame.append(compareFrame_test)
compareFrame['Date'] = compareFrame.index
compareFrame.index = list(range(1,len(compareFrame)+1))
sns.lineplot(data= [
compareFrame[compareFrame.Date >'2020-04-01'].train_real,
compareFrame[compareFrame.Date >='2020-07-01'].test_real,
compareFrame[compareFrame.Date >'2020-04-01'].train_prd,
compareFrame[compareFrame.Date >='2020-07-01'].test_prd,
])
```
| github_jupyter |
# Scraping a single web page
# Preliminaries
For our example, we will scrape a document from the Securities and Exchange Commission (SEC):
https://www.sec.gov/Archives/edgar/data/34088/000003408816000065/0000034088-16-000065-index.htm
## What is allowed?
All published U.S. government data are in the public domain, so no copyright issues with the SEC website. For other websites, look for a Terms of Use statement to determine if your use is allowed.
Find out what parts of the website are considered OK to be scraped: https://www.sec.gov/robots.txt
/Archives/edgar/data is allowed to be scraped.
## Initial setup
If necessary, install the following libraries using PIP, Conda, or your IDE's package manager:
- requests
- beautifulsoup4
- soupseive
- html5lib
```
import requests # best library to manage HTTP transactions
import csv # library to read/write/parse CSV files
from bs4 import BeautifulSoup # web-scraping library
```
## Define a function to write results to a CSV
```
def writeCsv(fileName, array):
fileObject = open(fileName, 'w', newline='', encoding='utf-8')
writerObject = csv.writer(fileObject)
for row in array:
writerObject.writerow(row)
fileObject.close()
```
## Create a User-Agent string
The User-Agent HTTP request header identifies the client to the server. In typical web use, the user agent is a web browser. Here's an example user agent description:
```
Mozilla/5.0 (Macintosh; Intel Mac OS X 10_13_6)
```
The description starts with the name and version of the software, followed by some identifying information about the creator in parentheses. Nefarious scrapers might try to impersonate a web browser by sending its user agent information. We don't want to do that. If you don't include a user agent, the default Python user agent string will be sent and that may be blocked automatically by some web servers.
Here's an example user agent:
```
BaskaufScraper/0.1 (steve.baskauf@vanderbilt.edu)
```
I've given the client a descriptive name that is likely to be unique and a very low version number indicating that it's in development. I've included my email address so that if my script does something bad, the server admins can email me about it instead of just immediately blocking me.
```
baseUrl = 'https://www.sec.gov'
url = 'https://www.sec.gov/Archives/edgar/data/34088/000003408816000065/0000034088-16-000065-index.htm'
acceptMediaType = 'text/html'
userAgentHeader = 'BaskaufScraper/0.1 (steve.baskauf@vanderbilt.edu)'
requestHeaderDictionary = {
'Accept' : acceptMediaType,
'User-Agent': userAgentHeader
}
```
# Scrape the web page
Beautiful Soup documentation: https://www.crummy.com/software/BeautifulSoup/bs4/doc/
Beautiful soup creates a custom object that prints out as HTML, but that has special attributes and methods that allows its XML structure to be traversed.
We will use the following attributes and methods:
- `.find_all(elementNameString)` method: creates a list of all descendant elements named `elementNameString`
- `.contents` attribute: a list of all child elements
- `.name` attribute: the name of the element
- `.text` attribute: the text value contained in the element
- `.get(attributeName)` method: extract the value of the attribute named `attributeName`
## Retrieve the HTML
Retrieve the text of the web page using HTTP, then clean it up by creating a Beautiful Soup object
```
response = requests.get(url, headers = requestHeaderDictionary)
soupObject = BeautifulSoup(response.text,features="html5lib")
print(soupObject)
```
## Extract the data from the HTML
There are two tables in the document. Extract them, then pull out the first one.
```
tableObject = soupObject.find_all('table')[0]
print(tableObject)
```
Extract the rows from the table.
```
rowObjectsList = tableObject.find_all('tr')
print(rowObjectsList)
```
Find the data rows, collect the desired cells, and save as a spreadsheet.
```
# Create a list of lists to output as a CSV
outputTable = []
for row in rowObjectsList:
outputRow = []
# determine if the row is a header or data row
dataRow = False
childElements = row.contents
for element in childElements:
if element.name == 'td':
dataRow = True
# if its a data row, then get the contents of the Description and the Type and the URL of the Document
if dataRow:
cells = row.find_all('td')
outputRow.append(cells[1].text)
outputRow.append(baseUrl + cells[2].a.get('href'))
outputRow.append(cells[3].text)
print(cells[1].text)
print(baseUrl + cells[2].a.get('href'))
print(cells[3].text)
print()
outputTable.append(outputRow)
# Write the lists of lists to a CSV spreadsheet
fileName = 'sec_table.csv'
writeCsv(fileName, outputTable)
```
| github_jupyter |
# Object Detection with YOLO in Keras
In this exercise, you'll use a Keras implementation of YOLO to detect objects in an image.
> **Important**: Using the YOLO model is resource-intensive. before running the code in this notebook, shut down all other notebooks in this library (In each open notebook other than this one, on the **File** menu, click **Close and Halt**). If you experience and Out-of-Memory (OOM) error when running code in this notebook, shut down this entire library, and then reopen it and open only this notebook.
## Install Keras
First, let's ensure that the latest version of Keras is installed.
```
!pip install --upgrade keras
```
## Download and Convert YOLO weights
YOLO is based on the Darknet model architecture - an open-source model written in C. The creators of this model have provided pre-trained weights that were trained on the [Common Objects in Context (COCO) dataset](cocodataset.org) - a common set of sample images for computer vision research.
Run the following cell to download the weights, and convert them into a suitable format for use with Keras.
> _**Note**: This can take some time to run_
```
!wget https://pjreddie.com/media/files/yolov3.weights -O ~/yolo3.weights
!python yolo_keras/convert.py yolo_keras/yolov3.cfg ~/yolo3.weights ~/yolo.h5
```
After the weights have been downloaded, the layers in the model are described.
## Load the Weights into a Keras Model
Now that we have the weights, we can load them into a Keras model.
> **Note** The code to implement the Keras model is in **yolo_keras/model.py**. Additionally, **yolo_keras/utils.py** contains some functions that are used to assemble and use the model.
```
import os
import numpy as np
from keras import backend as K
from keras.models import load_model
from keras.layers import Input
from yolo_keras.utils import *
from yolo_keras.model import *
# Get the COCO classes on which the model was trained
classes_path = "yolo_keras/coco_classes.txt"
with open(classes_path) as f:
class_names = f.readlines()
class_names = [c.strip() for c in class_names]
num_classes = len(class_names)
# Get the anchor box coordinates for the model
anchors_path = "yolo_keras/yolo_anchors.txt"
with open(anchors_path) as f:
anchors = f.readline()
anchors = [float(x) for x in anchors.split(',')]
anchors = np.array(anchors).reshape(-1, 2)
num_anchors = len(anchors)
# Set the expected image size for the model
model_image_size = (416, 416)
# Create YOLO model
home = os.path.expanduser("~")
model_path = os.path.join(home, "yolo.h5")
yolo_model = load_model(model_path, compile=False)
# Generate output tensor targets for bounding box predictions
# Predictions for individual objects are based on a detection probability threshold of 0.3
# and an IoU threshold for non-max suppression of 0.45
input_image_shape = K.placeholder(shape=(2, ))
boxes, scores, classes = yolo_eval(yolo_model.output, anchors, len(class_names), input_image_shape,
score_threshold=0.3, iou_threshold=0.45)
print("YOLO model ready!")
```
## Use the Model to Detect Objects
Now we're ready to use the YOLO model to detect objects in images.
### Create functions to detect and display objects
We'll create a couple of functions:
- **detect_objects**: Submits an image to the model and returns predicted object locations
- **show_objects**: Displays the image with a bounding box fo each detected object.
```
def detect_objects(image):
# normalize and reshape image data
image_data = np.array(image, dtype='float32')
image_data /= 255.
image_data = np.expand_dims(image_data, 0) # Add batch dimension.
# Predict classes and locations using Tensorflow session
sess = K.get_session()
out_boxes, out_scores, out_classes = sess.run(
[boxes, scores, classes],
feed_dict={
yolo_model.input: image_data,
input_image_shape: [image.size[1], image.size[0]],
K.learning_phase(): 0
})
return out_boxes, out_scores, out_classes
def show_objects(image, out_boxes, out_scores, out_classes):
import random
from PIL import Image
import matplotlib.patches as patches
import matplotlib.pyplot as plt
%matplotlib inline
# Set up some display formatting
cmap = plt.get_cmap('tab20b')
colors = [cmap(i) for i in np.linspace(0, 1, 20)]
# Plot the image
img = np.array(image)
plt.figure()
fig, ax = plt.subplots(1, figsize=(12,9))
ax.imshow(img)
# Set up padding for boxes
img_size = model_image_size[0]
pad_x = max(img.shape[0] - img.shape[1], 0) * (img_size / max(img.shape))
pad_y = max(img.shape[1] - img.shape[0], 0) * (img_size / max(img.shape))
unpad_h = img_size - pad_y
unpad_w = img_size - pad_x
# Use a random color for each class
unique_labels = np.unique(out_classes)
n_cls_preds = len(unique_labels)
bbox_colors = random.sample(colors, n_cls_preds)
# process each instance of each class that was found
for i, c in reversed(list(enumerate(out_classes))):
# Get the class name
predicted_class = class_names[c]
# Get the box coordinate and probability score for this instance
box = out_boxes[i]
score = out_scores[i]
# Format the label to be added to the image for this instance
label = '{} {:.2f}'.format(predicted_class, score)
# Get the box coordinates
top, left, bottom, right = box
y1 = max(0, np.floor(top + 0.5).astype('int32'))
x1 = max(0, np.floor(left + 0.5).astype('int32'))
y2 = min(image.size[1], np.floor(bottom + 0.5).astype('int32'))
x2 = min(image.size[0], np.floor(right + 0.5).astype('int32'))
# Set the box dimensions
box_h = ((y2 - y1) / unpad_h) * img.shape[0]
box_w = ((x2 - x1) / unpad_w) * img.shape[1]
y1 = ((y1 - pad_y // 2) / unpad_h) * img.shape[0]
x1 = ((x1 - pad_x // 2) / unpad_w) * img.shape[1]
# Add a box with the color for this class
color = bbox_colors[int(np.where(unique_labels == c)[0])]
bbox = patches.Rectangle((x1, y1), box_w, box_h, linewidth=2, edgecolor=color, facecolor='none')
ax.add_patch(bbox)
plt.text(x1, y1, s=label, color='white', verticalalignment='top',
bbox={'color': color, 'pad': 0})
plt.axis('off')
plt.show()
print("Functions ready")
```
### Use the functions with test images
Now we're ready to get some predictions from our test images.
```
import os
from PIL import Image
test_dir = "../../data/object_detection"
for image_file in os.listdir(test_dir):
# Load image
img_path = os.path.join(test_dir, image_file)
image = Image.open(img_path)
# Resize image for model input
image = letterbox_image(image, tuple(reversed(model_image_size)))
# Detect objects in the image
out_boxes, out_scores, out_classes = detect_objects(image)
# How many objects did we detect?
print('Found {} objects in {}'.format(len(out_boxes), image_file))
# Display the image with bounding boxes
show_objects(image, out_boxes, out_scores, out_classes)
```
## Acknowledgements and Citations
The original YOLO documentation is at https://pjreddie.com/darknet/yolo/.
The Keras implementation of YOLO used in this exercise is based on the work of qqwweee at https://github.com/qqwweee/keras-yolo3, with some simplifications.
The test images used in this exercise are from the PASCAL Visual Object Classes Challenge (VOC2007) dataset at http://host.robots.ox.ac.uk/pascal/VOC/voc2007/.
@misc{pascal-voc-2007,
author = "Everingham, M. and Van~Gool, L. and Williams, C. K. I. and Winn, J. and Zisserman, A.",
title = "The {PASCAL} {V}isual {O}bject {C}lasses {C}hallenge 2007 {(VOC2007)} {R}esults",
howpublished = "http://www.pascal-network.org/challenges/VOC/voc2007/workshop/index.html"}
| github_jupyter |
# A Função Sigmoide na Regressão Logística
Ao aprender sobre regressão logística, fiquei confuso a respeito de por que uma função sigmóide era usada para mapear as entradas até a saída prevista. Quero dizer, claro, é uma função legal que mapeia de forma limpa de qualquer número real para um intervalo de $ -1 $ a $ 1 $, mas de onde ele veio? Este caderno espera explicar.
## Regressão Logistica
Com a classificação, temos uma amostra com alguns atributos (recursos a.k.a) e, com base nesses atributos, queremos saber se ela pertence ou não a uma classe binária. A probabilidade de que a saída seja 1 dada sua entrada poderia ser representada como:
$$P(y=1 \mid x)$$
Se as amostras de dados tiverem recursos de $ n $, e acharmos que podemos representar essa probabilidade por meio de alguma combinação linear, podemos representar isso como:
$$P(y=1 \mid x) = w_o + w_1x_1 + w_2x_2 + ... + w_nx_n$$
O algoritmo de regressão poderia ajustar esses pesos aos dados que ele vê, entretanto, pareceria difícil mapear uma combinação linear arbitrária de entradas, cada uma variando de $ - \infty $ a $ \infty $ a um valor de probabilidade no intervalo de US $ 0 a US $ 1.
### O "Odds Ratio"
O odds ratio é um conceito relacionado à probabilidade que pode nos ajudar. É igual à probabilidade de sucesso dividida pela probabilidade de falha, e pode ser familiar para você se você olhar para linhas de apostas em disputas esportivas:
$$odds(p) = \frac{p}{1-p}$$
Dizendo, "as chances de a saída ser dada uma entrada" ainda parecem capturar o que estamos procurando. No entanto, se traçarmos a função de chances de 0 a 1, ainda há um problema:
<img src='https://nbviewer.jupyter.org/github/rasbt/python-machine-learning-book/blob/master/code/bonus/images/logistic_regression_schematic.png'>
```
import matplotlib.pyplot as plt
import pandas as pd
import statsmodels.api as sm
import numpy as np
import bokeh
from bokeh.plotting import figure, show, output_file
from bokeh.io import output_notebook, push_notebook, show
from ipywidgets import interact
import scipy.special
import statsmodels.formula.api as smf
from scipy import stats
import warnings
warnings.filterwarnings("ignore")
output_notebook()
import math
%matplotlib inline
%config InlineBackend.figure_format = 'svg'
def odds(p):
return p/(1 - p)
x = np.arange(0, 1, 0.1)
odds_x = odds(x)
plt.plot(x, odds_x)
plt.axvline(0.0, color='k')
plt.ylim(-0.1, 15)
plt.xlabel('x')
plt.ylabel('odds(x)')
# y axis ticks and gridline
plt.yticks([0.0, 5, 10])
ax = plt.gca()
ax.yaxis.grid(True)
plt.tight_layout()
```
### Log Odds
Uma combinação linear arbitrária dos recursos de entrada ainda pode ser menor que zero. No entanto, se pegarmos o log do odds ratio, agora temos algo que varia de $ - \infty $ a $\infty$
```
def log_odds(p):
return np.log(p / (1 - p))
x = np.arange(0.005, 1, 0.005)
log_odds_x = log_odds(x)
plt.plot(x, log_odds_x)
plt.axvline(0.0, color='k')
plt.ylim(-8, 8)
plt.xlabel('x')
plt.ylabel('log_odds(x)')
# y axis ticks and gridline
plt.yticks([-7, 0, 7])
ax = plt.gca()
ax.yaxis.grid(True)
plt.tight_layout()
plt.show()
```
Ter uma combinação linear de arbitrária apresenta mapa para o `função log_odds` permite quaisquer possíveis valores de entrada para cada $ x_i $ e ainda representa conceitualmente o que estamos tentando representar: a de que uma combinação linear de entradas está relacionada com a liklihood que uma amostra pertence a uma determinada classe. Nota: o log da função odds é freqüentemente chamado de função "logistic". Então agora nós temos:
$$\text{log_odds}(P(y=1 \mid x)) = w_o + w_1x_1 + w_2x_2 + ... + w_nx_n$$
Se nós ainda quisermos obter o $ P (y = 1 \mid x) $, podemos tomar o inverso da função `log_odds`.
## A Sigmóide
Vamos encontrar o inverso da função `log_odds`: Começando com:
$y = log(\frac{x}{1-x})$
e trocando $ y $ e $ x $ e resolvendo por $ y $
<br>$x = log(\frac{y}{1-y})$
<br>$e^x = \frac{y}{1-y}$
<br>$y = (1-y)*e^x$
<br>$y = e^x - y*e^x$
<br>$y + ye^x = e^x$
<br>$y*(1 + e^x) = e^x$
<br>$y = \frac{e^x}{1+e^x}$
<br>$y = \frac{1}{\frac{1}{e^x} + 1}$
<br>$y = \frac{1}{1 + e^{-x}}$
Vamos usar o $ \phi $ para representar essa função e plotar isso para ter uma ideia do que parece:
```
def inverse_log_odds(z):
return 1.0 / (1.0 + np.exp(-z))
z = np.arange(-7, 7, 0.1)
phi_z = inverse_log_odds(z)
plt.plot(z, phi_z)
plt.axvline(0.0, color='k')
plt.ylim(-0.1, 1.1)
plt.xlabel('z')
plt.ylabel('$\phi (z)$')
# y axis ticks and gridline
plt.yticks([0.0, 0.5, 1.0])
ax = plt.gca()
ax.yaxis.grid(True)
plt.tight_layout()
plt.show()
```
A forma inversa da função logística é parecida com um S, que, eu li, é por isso que é chamada de função Sigmoide. Então voltando ao nosso:
$$\text{log_odds}(P(y=1 \mid x)) = w_o + w_1x_1 + w_2x_2 + ... + w_nx_n$$
Se chamarmos de $ w_o + w_1x_1 + w_2x_2 + ... + w_nx_n = w ^ Tx $ simplesmente $ z (x) $:
$$ \text{log_odds}(P(y=1 \mid x)) = z(x) $$
E tomando o inverso:
$$ P(y=1 \mid x) = \phi(z) = \dfrac{1}{1 + e^{-z}} $$
e lá está: Regressão Logística ajusta-se a pesos de modo que uma combinação linear de suas entradas mapeie as probabilidades de log, sendo a saída igual a 1.
# Vamos a mais um exemplo
## Vamos usar um dataset da UCI
<br>
<img src="img/uci.png" / width='400' >
<br>
Bem-vindo ao Repositório de Aprendizado de Máquina da UC Irvine!
Atualmente, mantemos 468 conjuntos de dados como um serviço para a comunidade de aprendizado de máquina. Você pode ver todos os conjuntos de dados através da nossa interface pesquisável. Para uma visão geral do Repositório, por favor visite nossa página Sobre. Para obter informações sobre como citar conjuntos de dados em publicações, leia nossa política de citações. Se você deseja doar um conjunto de dados, consulte nossa política de doações. Para quaisquer outras perguntas, sinta-se à vontade para entrar em contato com os bibliotecários do Repositório.
# Banco Português
O conjunto de dados provém do repositório UCI repository ( http://archive.ics.uci.edu/ml/index.php ) e está relacionado com campanhas de marketing direto (chamadas telefónicas) de uma instituição bancária portuguesa. O objetivo da classificação é prever se o cliente irá assinar (1/0) para um depósito a prazo (variável y).
Com as seguintes variáveis:
- age (numeric)
- job : type of job (categorical: “admin”, “blue-collar”, “entrepreneur”, “housemaid”, “management”, “retired”, “self-employed”, “services”, “student”, “technician”, “unemployed”, “unknown”)
- marital : marital status (categorical: “divorced”, “married”, “single”, “unknown”)
- education (categorical: “basic.4y”, “basic.6y”, “basic.9y”, “high.school”, “illiterate”, “professional.course”, “university.degree”, “unknown”)
- default: has credit in default? (categorical: “no”, “yes”, “unknown”)
- housing: has housing loan? (categorical: “no”, “yes”, “unknown”)
- loan: has personal loan? (categorical: “no”, “yes”, “unknown”)
- contact: contact communication type (categorical: “cellular”, “telephone”)
- month: last contact month of year (categorical: “jan”, “feb”, “mar”, …, “nov”, “dec”)
- day_of_week: last contact day of the week (categorical: “mon”, “tue”, “wed”, “thu”, “fri”)
- duration: last contact duration, in seconds (numeric). Important note: this attribute highly affects the output target (e.g., if duration=0 then y=’no’). The duration is not known before a call is performed, also, after the end of the call, y is obviously known. Thus, this input should only be included for benchmark purposes and should be discarded if the intention is to have a realistic predictive model
- campaign: number of contacts performed during this campaign and for this client (numeric, includes last contact)
- pdays: number of days that passed by after the client was last contacted from a previous campaign (numeric; 999 means client was not previously contacted)
- previous: number of contacts performed before this campaign and for this client (numeric)
- poutcome: outcome of the previous marketing campaign (categorical: “failure”, “nonexistent”, “success”)
- emp.var.rate: employment variation rate — (numeric)
- cons.price.idx: consumer price index — (numeric)
- cons.conf.idx: consumer confidence index — (numeric)
- euribor3m: euribor 3 month rate — (numeric)
- nr.employed: number of employees — (numeric)
- *y* — has the client subscribed a term deposit? (binary: “1”, means “Yes”, “0” means “No”)
## Lendo e verificando os dados
```
df = pd.read_csv('../../99 Datasets/marketing.csv.zip')
df.shape
df.columns
df.head()
df.info()
```
df = df.astype({
"age":"int64",
"duration":"int64",
"campaign":"int64",
"pdays":"int64",
"previous":"int64",
"emp_var_rate":"float64",
"cons_price_idx":"float64",
"cons_conf_idx":"float64",
"euribor3m":"float64",
"nr_employed":"float64",
"y":"int64"})
df.info()
```
df.describe().T
```
## Investigando a correlação
```
df.corr().round(2)
df['y'].value_counts()
df['y'].value_counts(normalize=True)
# Analisando a correlação de todas as variáveis em relação ao Y: se o cliente comprou ou não o produto de Renda Fixa
df.corr().round(2)['y'].sort_values()
```
## Analisando probabilidade condicional com pandas
<br>
<img src="img/prob_condicional.png" / width='300' >
<br>
Na teoria da probabilidade, a probabilidade condicional é uma medida da probabilidade de um evento (ocorrendo alguma situação particular), dado que outro evento ocorreu. Se o evento de interesse é A e o evento B é conhecido ou assumido como tendo ocorrido, "a probabilidade condicional de A dado B", ou "a probabilidade de A sob a condição B", é geralmente escrita como $ P (A | B ) $, ou às vezes $ PB (A) $ ou $ P (A / B) $. Por exemplo, a probabilidade de que qualquer pessoa tenha uma tosse em um determinado dia pode ser de apenas 5%. Mas se nós sabemos ou assumimos que a pessoa tem um resfriado, então eles são muito mais propensos a tossir. A probabilidade condicional de tossir pela indisposição pode ser de 75%, então: $P (Tosse)$ = 5% ; $P (Tosse | Doente)$ = 75%
O conceito de probabilidade condicional é um dos mais fundamentais e um dos mais importantes na teoria da probabilidade. Mas as probabilidades condicionais podem ser bastante escorregadias e requerem interpretação cuidadosa. Por exemplo, não precisa haver uma relação causal entre A e B, e eles não precisam ocorrer simultaneamente.
Neste sentido é importante pensar que, quando discutimos a correlação entre as variáveis, estamos calculando uma variação linear de uma variável com a outra, o que significa que não captamos variações não lineares entre estas duas variáveis.
Por isso, podemos pensar que a probabilidade condicional entre as variávies nos dá a idéia de interdependência entre ambas, a qual não pode ser captada pela correlação devido a falta de linearidade entre elas.
### Aplicação da probabilidade condicional para variáveis categóricas
- Variáveis categóricas
```
df['job'].value_counts().plot.hist()
df.dtypes
# Mudando o tipo de variável para Job
df['job'] = df['job'].astype('category')
df.dtypes
df.education.unique()
df.marital.unique()
df.job.unique()
```
## Utilizando groupby, pivot e crosstab
df.groupby(['']).agg({})
pd.pivot_table(df, values='', columns='')
pd.crosstab(index, columns, values, aggfunc)
## Groupby
Uma operação groupby envolve alguma combinação de divisão do objeto, aplicação de uma função e combinação dos resultados. Isso pode ser usado para agrupar grandes quantidades de dados e operações de computação nesses grupos.
```
# Média de adesão ao produto bancário por estado civil
print("Mean y by marital status")
print(df.groupby(['marital'])[['y']].mean())
print("Stantard Dev by marital status")
print(df.groupby(['marital'])[['y']].std())
# Contagem do número de adesões ao produto bancário por estado civil
df.groupby(['marital'])[['y']].count()
# Média de adesão do produto bancário em tipo de emprego por estado civil
df.groupby(['marital', 'job'])[['y']].mean().tail(20)
# DataFrame da contagem de adesões ao produto bancário em tipo de emprego por estado civil
temp = pd.pivot_table(df, values='y',
index='job',
columns='marital',
aggfunc='count')
temp['sum'] = temp.sum(axis=1)
temp
```
## Crosstab
Calcula uma tabulação cruzada simples de dois (ou mais) fatores. Por padrão, calcula uma tabela de frequência dos fatores, a menos que uma matriz de valores e uma função de agregação sejam passadas.
```
df.shape
df.drop_duplicates().shape
df.job.value_counts()
# Média, contagens únicas e desvio-padrão de adesão ao produto bancário do estado civil por tipo de emprego
pd.crosstab(index=df['job'],
columns=df['marital'],
values=df['y'],
aggfunc=['mean', 'nunique', 'std'])
```
## Utilizando qcut
Função de discretização baseada em quantis. Transforma em discreta a variável em intervalos de tamanho igual com base na classificação ou com base em quantis de amostra. Por exemplo, 1000 valores para 10 quantis produziriam um objeto Categórico indicando associação de quantil para cada ponto de dados.
```
df['grouped_duration'] = pd.qcut(x=df['duration'], q=5, labels=[1,2,3,4,5])
# Separação da variável idade em tercil (q=3)
pd.qcut(x=df['age'], q=3, labels=["estudante",
"trabalhador",
"aposentado"]).value_counts(3)
# Criando uma Dummy
df['grouped_age'] = pd.qcut(x=df['age'], q=3, labels=[1,2,3])
df['grouped_age'].value_counts()
df.describe().T
```
### Variável duration
A duração de uma chamada para um cliente somente é conhecida após seu término, eventualmente no memsmo momento da adesão do cliente ao produto bancário, e por isso não é conhecida até que o cliente execute a adesão, ou seja, assuma valor verdadeiro (1) para Y. isso se caracteria como LEAKAGE, ou um dados que não pode ser utilizado para predição (Conforme descrito pelo próprio dicionário de dados acima).
### O que é vazamento de dados no Machine Learning?
O vazamento de dados pode fazer com que você crie modelos preditivos excessivamente otimistas, se não completamente inválidos.
O vazamento de dados é quando as informações de fora do conjunto de dados de treinamento são usadas para criar o modelo. Essa informação adicional pode permitir que o modelo aprenda ou saiba algo que de outra forma não conheceria e, por sua vez, invalidaria o desempenho estimado do modo que está sendo construído.
"Se algum outro recurso cujo valor não estivesse realmente disponível na prática no momento em que você deseja usar o modelo para fazer uma previsão, é um recurso que pode introduzir vazamento em seu modelo." _Data Skeptic_
"Quando os dados que você está usando para treinar um algoritmo de aprendizado de máquina têm as informações que você está tentando prever." _Daniel Gutierrez_
Há um tópico em segurança de computador chamado vazamento de dados e prevenção de perda de dados que está relacionado, mas não o que estamos falando.
#### Vazamento de dados é um problema sério por pelo menos 3 razões:
- É um problema se você estiver executando uma competição de aprendizado de máquina. Os modelos top usarão os dados com vazamento em vez de serem bons modelos gerais do problema subjacente.
- É um problema quando você é uma empresa que fornece seus dados. Reverter um anonimato e ofuscação pode resultar em uma violação de privacidade que você não esperava.
- É um problema quando você está desenvolvendo seus próprios modelos preditivos. Você pode estar criando modelos excessivamente otimistas que são praticamente inúteis e não podem ser usados na produção.
Como praticantes de aprendizado de máquina, estamos principalmente preocupados com este último caso.
#### Eu tenho vazamento de dados?
Uma maneira fácil de saber que você tem vazamento de dados é se estiver obtendo um desempenho que parece bom demais para ser verdade.
Como você pode prever números de loteria ou escolher ações com alta precisão.
"Bom demais para ser verdade" desempenho é "uma oferta inoperante" de sua existência
O vazamento de dados geralmente é mais um problema com conjuntos de dados complexos, por exemplo:
- Conjuntos de dados de séries temporais ao criar conjuntos de treinamento e teste podem ser difíceis.
- Gráfico de problemas em que métodos de amostragem aleatória podem ser difíceis de construir.
- Observações analógicas como som e imagens em que as amostras são armazenadas em arquivos separados que possuem um tamanho e um registro de data e hora.
### Prosseguindo com a análise
```
# Visualizando a distribuição da variável duration
df.duration.plot.hist(bins=40, figsize=(8,6))
# Separando o duration em quantil (q=5) e criando uma Dummy
df['grouped_duration'] = pd.qcut(x=df['duration'], q=5, labels=[1,2,3,4,5])
df.groupby(['grouped_duration'])[['y']].mean()
pd.crosstab(index=df['marital'],
columns=df['grouped_duration'],
values=df['y'],
aggfunc='mean')
```
# Quando usamos OLS para estimar esse efeito, temos um modelo linear de probabilidade
- pode ser usado porque a categórica tem distribuição de Bernouli e sua média é a própria probabilidade
- Pode ser maior que 1 e menor que zero (que não faz muito sentido)
```
# cons_conf_idx euribor3m nr_employed
df.columns
```
### Rodando uma primeira regressão linear para comparação
```
function = ''' y ~ age + duration + campaign + pdays + previous + emp_var_rate + cons_price_idx'''
model1 = smf.ols(function, df).fit()
print(model1.summary2())
```
## Rodando uma Logit com a mesma função
```
logit = smf.logit(function, df).fit()
print(logit.summary2())
```
## Olhando as correlações e acrescentando as variáveis originais à Logit
```
df.corr()['y']
function1 = ''' y ~ age
+ duration
+ campaign
+ pdays
+ previous
+ emp_var_rate
+ cons_price_idx
+ cons_conf_idx
+ euribor3m
+ nr_employed
'''
logit = smf.logit(function1, df).fit()
print(logit.summary2())
```
## Criando outras variáveis para serem colocadas na Regressão Logística
```
# Modificando o tipo de variável para inteira para ser colocada na regressão
df.grouped_age = df.grouped_age.astype(int)
df.grouped_duration = df.grouped_duration.astype(int)
df.corr()['y']
```
## Transformando os valores da variável educação para a média do valor de adesão
```
df['education'].value_counts()
df_education = pd.crosstab(index=df['education'],
columns=df['education'],
values=df['y'],
aggfunc=['mean']).sum()
df_education
df.education[df.education == 'basic.4y'] = df_education[0]
df.education[df.education == 'basic.6y'] = df_education[1]
df.education[df.education == 'basic.9y'] = df_education[2]
df.education[df.education == 'high.school'] = df_education[3]
df.education[df.education == 'illiterate'] = df_education[4]
df.education[df.education == 'professional.course'] = df_education[5]
df.education[df.education == 'university.degree'] = df_education[6]
df.education[df.education == 'unknown'] = df_education[7]
df.education.value_counts()
df.education = df.education.astype(float)
df.corr()['y']
function2 = ''' y ~ age
+ education
+ duration
+ campaign
+ pdays
+ previous
+ emp_var_rate
+ cons_price_idx
+ cons_conf_idx
+ euribor3m
+ nr_employed
+ grouped_duration
+ grouped_age
'''
logit = smf.logit(function2, df).fit()
print(logit.summary2())
```
# Logit x Probit (Poisson x OLS)
```
fig = plt.figure(figsize=(12,8))
ax = fig.add_subplot(111)
support = np.linspace(-6, 6, 1000)
ax.plot(support, stats.logistic.cdf(support), 'r-', label='Logistic')
ax.plot(support, stats.norm.cdf(support), label='Probit')
ax.legend();
fig = plt.figure(figsize=(12,8))
ax = fig.add_subplot(111)
support = np.linspace(-6, 6, 1000)
ax.plot(support, stats.logistic.pdf(support), 'r-', label='Logistic')
ax.plot(support, stats.norm.pdf(support), label='Probit')
ax.legend();
```
# Matriz de confusão
```
y_predict = logit.predict()
df.y.mean()
y_predict.mean()
```
## Adotamos o limite de Precisão X Sesibilidade como a média das predições
```
for i in range(0,len(y_predict),1):
if y_predict[i] < y_predict.mean():
y_predict[i] = 0
else:
y_predict[i] = 1
from sklearn.metrics import confusion_matrix
conf_matrix = confusion_matrix(df.y, y_predict)
print(conf_matrix)
tn, fp, fn, tp = confusion_matrix(df.y, y_predict).ravel()
precision = tp / (tp+fp)
precision
recall = tp / (tp+fn)
recall
```
## Utilizando GLM (Generalized Linear Model)
```
glm = smf.glm(function2, df).fit()
print(glm.summary2())
glm_predict = glm.predict()
glm_predict.mean()
for i in range(0,len(y_predict),1):
if y_predict[i] < glm_predict.mean():
y_predict[i] = 0
else:
y_predict[i] = 1
conf_matrix = confusion_matrix(df.y, y_predict)
print(conf_matrix)
tn, fp, fn, tp = confusion_matrix(df.y, y_predict).ravel()
precision = tp / (tp+fp)
precision
recall = tp / (tp+fn)
recall
```
### Fronteira de decisão
- https://gist.github.com/vietjtnguyen/6655020
| github_jupyter |
# Assignment 6 (Oct 26)
Today's topics are:
1. Conda Environments
2. Package installation
3. Basic Markov-chain Monte Carlo (MCMC)
There are many MCMC samplers in python, we are only going to go through one today, `PyMC3`.
See this nice article [Samplers demo](https://mattpitkin.github.io/samplers-demo/) by Matthew Pitkin where he demos 13 popular samplers in python.
## Readings (optional)
If you find this week's material new or challenging, you may want to read through some or all the following resources while working on your assignment:
**Conda Environments**:
* [SPIRL: 5.2. Conda environments](https://cjtu.github.io/spirl/anaconda_environments.html)
* [Conda official page: Managing environments](https://conda.io/projects/conda/en/latest/user-guide/tasks/manage-environments.html)
**MCMC**:
* [MCMC: A (very) Beginnner’s Guide](https://prappleizer.github.io/Tutorials/MCMC/MCMC_Tutorial.html)
* [MCMC sampling for dummies](https://twiecki.io/blog/2015/11/10/mcmc-sampling/)
* [Markov Chain Monte Carlo (MCMC)](https://people.duke.edu/~ccc14/sta-663/MCMC.html)
**YouTube Videos (highly recommended)**:
* [Markov Chain Monte Carlo (MCMC): Data Science Concepts](https://www.youtube.com/watch?v=yApmR-c_hKU)
* [Metropolis - Hastings: Data Science Concepts](https://www.youtube.com/watch?v=yCv2N7wGDCw)
## Managing Conda Environments
At the start of the term, we installed **Anaconda** which gave us access to Python and some popular Python packages. While Anaconda comes with several useful packages by default, we can also install brand new packages with a few simple commands. Once we're all `conda` wizards, we'll have the wealth of open source Python code at our fingertips!
### Why Environments
By default when we open a terminal after installing **Anaconda**, we should see `(base)` at the left, indicating that we are in the default `conda` environment.
```{note}
To open a terminal in VS Code, type `CTRL SHIFT ~` (Mac: `CTRL SHIFT ~`) or in the top bar go to `View -> Terminal`.
```
Terminology:
- **Anaconda**: the program we installed that comes with `conda` and a bunch of packages
- `conda`: the command line tool that helps us manage our Python packages and environments
- `base`: the name of the default `conda` environment
This `base` environment is where Anaconda keeps all of its default programs and inner workings, so it's recommended to *never install packages to `base` directly*. Instead, we want to make a new **environment**.
Having different `conda` environments allows us to tailor the packages we are using to each of our projects. Sometimes we will need `astropy`, sometimes we will need `pandas`. Sometimes we need neither! Today, we'll need a package called `pymc3` which is very specific to MCMC analysis.
The reason we don't want all of these packages in `base` is that they may have incompatible prerequisites (also called *dependency conflicts*). For example:
- `package1` can only run on `python v2.7`
- `package2` needs at least `python v3.9`
Since we can only have 1 version of Python at a time, if we try to install both of these packages, one is guaranteed to not work correctly!
Instead, we can install each package to its own `conda` **environment**. We can even get different versions of Python working on the same computer in only a couple steps!
### Working with Environments
In this section, we will use the terminal. These commands are also available through the user friendly [Anaconda Navigator](https://docs.anaconda.com/anaconda/navigator/getting-started/#navigator-managing-environments) GUI if you prefer to use that instead.
A summary of the most important conda commands is below (we abbreviate environment as env):
| Command | Description |
| - | - |
| `conda list` | List all packages and their versions in current env |
| `conda list \| grep pkg` | Search for pkg in current env and show its version |
| `conda install pkg1 pkg2 ...` | Install pkg 1, pkg 2, ... with `conda` in current env (see installing below) |
| `conda env list` | Show all environments and their locations (file paths) |
| `conda create -n env-name` | Create a new env called *env-name* |
| `conda create -n env-name python=3.X pkg1 pkg2 ...` | Create a new env with python v3.X and install pkg1, pkg2, ... |
| `conda activate env-name` | Activate *env-name* (running Python from this terminal will use packages in this env) |
| `conda deactivate` | Deactivate the current env (takes you to your previous env, usually `base`) |
| `conda remove --name env-name --all` | Delete an environment and all packages in it |
| `conda env export > environment.yml` | Create a file with all packages in current env (use to reproduce env on same machine) |
| `conda env export --from-history > environment.yml` | Create a file with only packages you installed in current env (use to reproduce env on different machines) |
| `conda env create -f environment.yml` | Re-build an env from an environment.yml file (see previous 2 commands) |
### [Hands-on] 5 mins
Type the following commands into a terminal as an interactive tour of the main `conda` commands
You will create, activate and install packages into a new `conda` environment. You will then save the environment to an `environment.yml` file, delete the environment, and re-build it from your `environment.yml`!
```bash
# We want to use a package called seaborn. Let's see if it's installed in base
conda list | grep seaborn
# Create Python 3.7 env with seaborn package v0.8 (type "y" then Enter)
conda create -n new-env python=3.7 seaborn=0.8
# List all envs (you should see new-env among them)
conda env list
# Activate our env
conda activate new-env
# Check that we have seaborn v0.8 installed now
conda list | grep seaborn
# We also want a package called bokeh (v2.1). Let's install it here
conda install bokeh=2.1
# Check that bokeh v2.1 is also installed
conda list | grep bokeh
# Export our environment to an environment.yml file so we can rebuild it later
conda env export > environment.yml
# Check to see we have an environment.yml file now
ls environment.yml
# Deactivate our environment
conda deactivate
# Check if we have seaborn v0.8 installed here (blank means no)
conda list | grep seaborn
# Delete our env and all packages in it (type "y" then Enter)
conda remove --name new-env --all
# List all envs again (new-env should be gone)
conda env list
# Check if we have access to seaborn and bokeh now :(
conda list | grep seaborn
conda list | grep bokeh
# Re-create our env from the file we saved
conda env create -f environment.yml
# List all envs again (new-env is back in town)
conda env list
# Activate our resurrected new-env
conda activate new-env
# Check that we can access seaborn and bokeh again!
conda list | grep seaborn
conda list | grep bokeh
# All done! You can remove this test env by following the same steps as before
conda deactivate
conda remove --name new-env --all
# You can also remove the environment.yml file
rm environment.yml
```
The above steps show that with only a few commands, we can have completely separate Python environments happily co-existing on our computers.
Even if you only ever use 1 environment, it is **very important** to export an `environment.yml` periodically so that if your computer dies or `conda` breaks, you can always reinstall the exact environment you were using to run your code.
If you want to share this environment file with your collaborators, make sure to use the `--from-history` flag (see table above). This makes it more likely the env will be installable on other machines and that `conda` won't install OS-specific packages on the wrong operating system.
### Conda Tips
Some common pitfalls, gotchas, and tips to remember with `conda`:
- Make sure to activate the desired environment each time you open a new terminal
- It doesn't matter what directory you run `conda` commands from:
- Activating an env activates it for the whole terminal even if you switch directories
- 1 exception: if you are loading/saving an `environment.yml`, you need to be in same directory (or give the full path)
- Before uninstalling Anaconda, make sure to save `environment.yml` files for each environment you want to restore later
- If you re-install a new version of Anaconda, you can load `environment.yml` files saved with older versions
- Update Anaconda periodically with the below commands (see [Anaconda versions](https://repo.anaconda.com/archive/) and choose the most recent):
```
conda update conda
conda install anaconda=VersionNumber
```
## Installing New Packages
Now that we know how to make a new `conda` environment, it's important to know that `conda` isn't the only way to install Python packages (in fact sometimes we need to use other ways).
The three ways to install new Python packages are:
1. Run `conda install pkg`
2. Run `pip install pkg`
3. Install from **source file** directly (we will talk about this in week 9!)
It's important to carefully read the documentation of a new package to see what the preferred installation method is - this can prevent errors and headaches down the line! This is particularly important for the package we are using today, `pymc3`, which has different installation instructions for different operating systems.
### Installing with conda / conda-forge
We already talked about how to install packages with `conda`, but there are a couple more things to know.
When we run `conda install`, the `conda` command checks a list of packages maintained by the Anaconda team and will install it and all of its dependencies once it makes sure there are no conflicts.
To check if a package exists and is installable with `conda`, we can use the `conda search` command.
```
# Try this in the terminal!
conda search seaborn=0.8
```
If anything shows up, we can just switch the `conda search` to `conda install` and we have a new package (pending dependency checks)!
But what if we try searching a package like [batman](https://lweb.cfa.harvard.edu/~lkreidberg/batman/)?
```
# Try this in the terminal!
conda search batman=1.9
```
All hope is not lost! In addition to the main Anaconda list of packages, there is also a community-driven list of packages that has rapidly grown much larger than any one team can maintain. This list is called `conda-forge` and we can access it with the **channel** or `-c` option.
```
# Try this in the terminal!
conda search -c conda-forge batman=1.9
```
Now this indicates we need to do `conda install -c conda-forge batman=1.9` to install the package from `conda-forge` (e.g., plain old `conda install batman=1.9` won't work).
### Installing with pip
If a package you want to install isn't in the default or conda-forge channels, you may be able to install it with `pip`, short for "Python Install Package". This is the original way to install packages in Python, but does not handle dependencies and dependency conflicts as well as `conda`.
As a general rule, using `conda` whenever possible and `pip` *only when necessary* will cut down on errors when working with Python packages.
The `pip` command installs from the [Python Package Index](https://pypi.org/) (PyPI or [Cheese Shop](https://www.youtube.com/watch?v=Hz1JWzyvv8A&ab_channel=AndrewBoynton) for short).
For example `pip install numpy` will install NumPy, but since NumPy is available from conda, we should always prefer `conda install numpy`.
```{note}
Just like `conda install`, `pip install` will install to the currently active environment. Remember to activate your environment before installing packages!
```
## Installing PyMC3 and Corner
Now that we know the basics of environments and installing packages, we're ready to install the packages needed for this assignment!
### Install PyMC3
Remember: always **read the documentation** to learn the best way of installing a package. The Markov-Chain Monte Carlo package, [PyMC3 documentation](https://docs.pymc.io/en/stable/) has different installation instructions depending on your OS, so read the instructions carefully before running any commands.
The basic structure of the command will be to create a new `conda` env, pulling from the `conda-forge` channel as we learned above.
```bash
# Check docs for the full command for your machine!
conda install -c conda-forge pymc3 ...
```
Beyond this, the packages to use and python version required may differ on Linux, Mac (M1 CPU), Mac (Intel CPU), and Windows, so please consult the installation instructions in the official [PyMC3 documentation](https://docs.pymc.io/en/stable/)
Once you have installed `pymc3`, you will need to activate the environment:
`conda activate pymc3`
Then, we will install 2 additional packages, `astropy` which you've seen before and `ipykernel` which will allow us to use this environment in Jupyter notebooks, e.g. the one you're reading now!
`conda install astropy ipykernel`
### Install Corner
If we go to the official [Corner documentation](https://corner.readthedocs.io/en/latest/index.html) and head to the installation page, we will see a recommendation to install with `conda`, but a command that says to use `pip`. We can always check if `corner` is available from `conda` with:
```bash
# Try this in the terminal!
conda search corner
conda search -c conda-forge corner
```
It looks like it is available from `conda-forge` (which we should usually prefer over `pip`)!
Make sure you are in the `pymc3` environment (should be in parentheses in your terminal, if not `conda activate pymc3`, then go ahead and install `corner` with:
`conda install -c conda-forge corner`
## Activating a new environment in VS Code
Finally, we're ready to get going with PyMC3 and Corner, we just need to tell VS Code about our new environment. To do so:
1. Reload VS Code with **CTRL+SHIFT+P** (Mac: **CMD+SHIFT+P**) -> Type "Reload window" + **ENTER**
2. Set workspace Python env with **CTRL+SHIFT+P** (Mac: **CMD+SHIFT+P**) -> Type "Python: Select Interpreter" + **ENTER** and choose `pymc3` from the list
3. Also set the Jupyter notebook kernel to `pymc3` with upper right button (should say **Select Kernel** or **Python 3.X...**)
4. Run the below cell to see if it worked! If not let us know!
```
import pymc3 as pm
import corner
print(f"Running on PyMC3 v{pm.__version__}")
print(f"Running on corner v{corner.__version__}")
```
## Markov-chain Monte Carlo (MCMC)
The **Markov-chain Monte Carlo (MCMC)** technique offers us a way to explore the possible parameters of a model fit to some data. Unlike ordinary linear regression which can be solved directly, MCMC uses Bayesian stats and random sampling to build up confidence in model fits that are too complicated or have too many parameters to simply solve by fitting a curve.
The basic idea of MCMC is:
- We have an equation or *model* we want to fit to some data
- This model may have many parameters, or complicated non-linear functions of some parameters (yuck)
- We also start with some *prior* knowledge about these model parameters
- Maybe we know the max/min value they can be, or a range each parameter usually falls in
- MCMC will vary the parameters in a *random but smart* way (using some math we'll mostly gloss over - see optional readings for more info!)
Then, if the data can be fit by the model *and* we had well constrained priors for the parameters *and* we let MCMC run long enough *and* we re-ran MCMC enough times independently, then something cool happens:
- MCMC will find the zones in parameter space that make the model best fit the data
- We will get a statistical average and $\pm$ uncertainty for all parameters simultaneously
- We can get precise fits to data we couldn't fit any other way (except trial & error)!
This technique is great for models with many parameters or sneaky local minima that can look like the best fit while hiding even better fit parameters. MCMC allows us to fit a whole set of parameters simultaneously so that we can avoid simplifying assumptions that weaken our model.
### How does it work?
Mostly Bayes' theorem and magic... but we can give a sense for how MCMC works without going too deep (see readings/videos for *lots* more info).
As the name suggests, MCMC is a combination of **Markov chain** and **Monte Carlo** methods that explore the possible parameters of our fit in many dimensions. A *Markov chain* is a series of events that only look at the current state (e.g., position in parameter space) to determine the next state. The *Monte Carlo* aspect is that at each position in parameter space, we move in a random direction to a new position (new set of parameters to try). Together, this amounts to a [random walk](https://en.wikipedia.org/wiki/Random_walk) (usually called a **walker**) through our parameters that will build up an erratic pattern over time:

The clever part is that even though our **walker** picks a random direction to move at each step, sometimes *it just stays put*. Our **walker** friend typically only moves if doing so gives us parameters that better fit our data (with some leeway). There are several algorithms for deciding whether to move our walker or not but those are outside the scope of this lesson. What's important is that our walker will *sample* our parameter space over time, and (hopefully) *converge* to a set of parameters that make our model fit our data.
The below [gif by M. Joseph](https://blog.revolutionanalytics.com/2013/09/an-animated-peek-into-the-workings-of-bayesian-statistics.html) shows how walkers can start in random parts of parameter space (here only 2 parameters, *mu* and *sigma*), then eventually converge via an MCMC random walk to a set of parameters that best fit the data. We know we're at the best fit parameters when the *posterior distribution* is 1 nice peak (we'll talk about what that means later!).

Since many models have *local minima* in parameter space, which could make a bad fit look good (essentially getting a walker stuck in a poorly fitting zone in parameter space), we generally run multiple walkers to see where most of them end up. If most of our walkers converge to a particular zone in parameter space, we can be more confident that this zone gives us our best fitting parameters.
## Simple Example with MCMC -- Finding Planet Mass and Radius
Let's return to our trusty free-fall gravity example. Recall we measured the height (**z**) of an object over time on some planet with unknown gravity. Previously we used curve fitting techniques to get the gravitational acceleration (**g**) with the equation:
$$z(t) = \frac{1}{2}gt^2$$
However, what if we want to know something about the planet mass (**M**) and planet Radius (**R**)? Pulling out the old Physics I textbook, we can use the definition of **g** and $F=ma$ to get:
$$g = \frac{GM}{R^2}$$
Now we can write height (**z**) in terms of **M**, **R**, **t**, and the universal gravitational constant (**G** = 6.674 $\times 10^{-11} m^3/kg/s^2$)
$$z(t,M,R) = \frac{1}{2}\frac{GM}{R^2}t^2$$
Now we have an expression with 4 variables (z, t, m, R), but we've only measured 2 (z and t). It seems hopeless to solve for our 2 unknowns, **M** and **R**, with normal algebra... in fact there are an infinite number of possible values we can plug in to make the above equation work...
Luckily, we know a little bit about planet masses and radii! For one, neither can be negative. Also, there are maximum values a planet mass / radius can have before it becomes a different object entirely. While this may not seem like much, these are **priors**, knowledge we have about our unknown parameters. To use our priors to inform the most likely M and R, we need one more concept: Bayes' Theorem.
### Bayes' Theorem
The Bayes Theorem tells us about probabilities *given some condition* or *given prior knowledge*. A simplified form of the Theorem is:
$$P(x|y) \propto P(y|x) \cdot P(x)$$
Where $P(x|y)$ means "the probability of x given that we know y", conversely $P(y|x)$ means "the probability of y given that we know x" and $P(x)$ is just the probability that event $x$ occurs. We can translate Bayes' theorem to MCMC speak where these 3 terms have different names but the same form:
$${\rm posterior probability} \propto ({\rm likelihood\ function}) \cdot ({\rm prior})$$
Because this is a lot of stats and jargon that we don't have time to dive deep on, we'll use a short story to help us build intuition for Bayes' theorem:
> One afternoon, you and your friend Lauren are talking about Bayes’ theorem when your brother, Ben, comes out of his room. You both notice that Ben is wearing a Yankees cap and an old worn-out T-shirt with a Yankees logo. Lauren says, "based on Ben's outfit, I bet he's going to a Yankees game!" You reply, "Ah, so based on your **prior** experience of people wearing matching caps and T-shirts to Yankees games, your Bayesian intuition says there is a high **probability** he is going to a baseball game”.
>
> Lauren thinks for a second and then says, “Oh I get it! If I had to guess, I'd say there's about an 80\% chance that people are **wearing a Yankees cap and T-shirt** *given* they are **going to a Yankees game**. This would be our **likelihood function**, *P(y|x)*. So then my **prior** is how likely it is that Ben is **going to a Yankees game**, *P(x)*. Well, the Yankees are playing in town today and I know that Ben has season tickets so I'd say my **prior** is Ben has a 95\% chance of going to the game today. So when I multiply my **likelihood function** by my **prior**, I get my **posterior probability** of Ben **going to a Yankees game** *given* he is **wearing a Yankees cap and T-shirt**, which is 80\% times 95\%, so **76\%!**." Ben rolls his eyes, grabs his keys and leaves, wondering why anyone would ruin a perfectly good Saturday talking about stats...
>
> You reply, "Nice! That's well reasoned based on your priors and Bayes' Theorem. But as Ben’s sister, I know for a fact he is going on a date with his partner and not to a ball game.”
>
> Lauren was skeptical of your certainty. “How could you possibly know that?”
>
> You eagerly reply, “Well, I agree on the **likelihood function** that 80\% of people wear matching caps and T-shirts given they are going to a Yankees game. BUT my **priors** are that
> 1) Ben got that shirt as a gift from his partner for their first-year anniversary,
> 2) Ben only wears that shirt when he sees his partner,
> 3) Ben's partner hates baseball.
>
> Therefore, from my **priors**, I'd guess Ben has only a 1\% chance of going to a Yankees game with his partner today. Therefore, my **posterior probability** that Ben is going to a baseball game given his outfit is 80\% times 1\%, **less than 1\%!**! So you see, having less informed priors can have a huge effect on your Bayesian prediction!"
In the coming weeks we'll learn more about the **likelihood function** and **posterior probability**, but some MCMC packages like **PyMC3** will handle those for us in simple cases as long as we have a model, data, and priors. Today our main focus is going to be on running an MCMC analysis and choosing good **priors**!
### Prior probability / Prior functions:
Our **prior** knowledge of the unknown parameters can be very important for having MCMC converge to the correct final parameters. Priors in MCMC analyses are defined as *probability distributions*. The two most commonly used prior distributions are the **uniform** and **normal** distributions.
The **uniform prior**, $\mathcal{U}(lower,upper)$, is the simpler of the two. If the sampled parameter is in the range `(lower, upper)`, the prior's probability is 1, otherwise the probability is 0.
For example, say we set our Mass (M) prior to $\mathcal{U}(0, 1)$ kg. This means that if means that if your walker goes to a part of parameter space where M=2 kg, the prior probability will be 0 and therefore our **posterior probability** will also be 0. The uniform prior puts a hard range on a parameter, but every value in that range is equally likely.
The **normal (Gaussian) prior**, assigns a bell-curve probability to a parameter with peak probability near the mean and lower probability with increasing standard deviations from the mean:
$$p(x) = \frac{1}{\sqrt{2\pi \sigma^2}}exp\left( -\frac{(x-\mu)^2}{2\sigma^2} \right)$$
For example, assuming we measure M with some uncertainty as $0.50\pm0.1$ kg, we can set a Gaussian prior with $\mu=0.5, \sigma=0.1$. Walkers nearer to 0.5 will have a higher **prior probability**, but the walkers will still explore lower probability space further from the mean.
There is also a combination of the above two priors, sometimes called a **truncated normal prior**. This is a Gaussian prior with $\mu$ and $\sigma$, except we can also set a lower or upper bound, beyond which the probability is 0.
For example, since we know mass (M) cannot be negative, we could use the same mean and sigma as the previous example as $\mu=0.5, \sigma=0.1$, but also set $lower=0$ to make sure we don't get any fits with negative mass.
## MCMC with PyMC3
Finally know all we need to start building MCMC models in Python! To get started with `pymc3`, we only need a couple components: **data**, a **model**, and **priors**. Then we just need to tweak some MCMC parameters and PyMC3 will do all the random walking for us!
The main steps are:
1. Create a `pymc3.model()` instance
2. Setup our PyMC3 object with **priors**, **model**, and **observed data**
3. Run `.sample()` to make the walkers explore parameter space!
### Using MCMC to learn mass and radius from Free-fall!
Recall: We want to estimate the planet mass (M) and radius (R) using this model:
$$z(t) = \frac{1}{2}\frac{GM}{R^2}t^2$$
We have free-fall data (**z**, **t**), and we do have some extra (prior) knowledge on M and R.
* When recording the free-fall height over time, we managed to measure the mass to some precision as M=$0.30\pm0.015$ Jupiter masses. We also know we can't have a negative mass. Therefore, we can use a truncated normal distribution prior for M.
* For R, we only know that it cannot be larger than 2 Jupiter radius, and cannot have be negative value. Therefore, we can use a uniform distribution with an upper and lower bound
* Our model can also be defined with a probability distribution, here we have well-behaved errors on our data points, so we will use a normal distribution for our model
For our priors and model, we will use the following 3 probability distributions provided by PyMC3 (which we alias to `pm`):
- `pm.Uniform()`
- `pm.Normal()`
- `pm.TruncatedNormal()`
You can find all built-in distributions at [PyMC3 Distributions](https://docs.pymc.io/en/stable/api/distributions/continuous.html#pymc3.distributions).
### Read in data
We will read in a free-fall data set from this [link](https://raw.githubusercontent.com/cjtu/spirl/master/spirl/data/position_measurement.csv).
```
import arviz as az
import pandas as pd
import pymc3 as pm
import corner
import matplotlib.pyplot as plt
# See plt styles: https://matplotlib.org/stable/gallery/style_sheets/style_sheets_reference.html
plt.style.use('ggplot')
# read in data
f = "https://raw.githubusercontent.com/cjtu/spirl/master/spirl/data/position_measurement.csv"
df = pd.read_csv(f)
df.head()
# plot the data
f = plt.figure(facecolor='white', figsize=(4,3), dpi=150 )
ax1 = plt.subplot(111)
ax1.errorbar(df['time(s)'], df['z(m)'], yerr = df['z_err(m)'], ms=5, lw=0.5,
fmt='.', color='tab:blue')
ax1.set_xlabel('time (s)', size='small', fontname='Helvetica')
ax1.set_ylabel('z (m)', size='small', fontname='Helvetica')
plt.show()
import astropy.units as u
from astropy import constants as const
Mj2si = const.M_jup.si.value # constant to transfer Jupiter mass to kg
Rj2si = const.R_jup.si.value # constant to transfer Jupiter radius to m
G_cgs = const.G.si.value
t = df['time(s)'].to_numpy()
z = df['z(m)'].to_numpy()
z_err = df['z_err(m)'].to_numpy()
# 1) Initialize PyMC3 model object
planet_model = pm.Model()
def free_fall(t, m, r, G=G_cgs):
"""
Model (likelihood) function for a free-falling object.
Parameters
----------
t : array_like
Time array.
m : float
Mass of object in kg.
r : float
Radius of object in m.
G : float
Gravitational constant in m^3 / kg / s^2.
"""
return 0.5 * (G * m / r**2) * t**2
# Set up the PyMC3 object with its prior and likelihood functions
with planet_model:
# Priors for unknown model parameters
# TruncatedNormal prior to make sure mass > 0 Jupiter masses
mass_j = pm.TruncatedNormal('mass_j', mu=0.3, sd=0.015, lower=0)
# Uniform prior for radius, Make sure 0 < R < 2 Jupiter masses
radius_j = pm.Uniform('radius_j', lower=0, upper=2)
# Convert to SI units
mass_si = Mj2si * mass_j
radius_si = Rj2si * radius_j
# Expected value of outcome, aka "the model"
z_model = free_fall(t, mass_si, radius_si, G_cgs)
# Gaussian likelihood (sampling distribution) of observations
# Passing z to the "observed" arg makes PyMC3 recognize this as the model to fit
obs = pm.Normal('obs', mu=z_model, sd=z_err, observed=z)
# No need to setup the posterior function as PyMC3 does it for you!
```
### Run the MCMC!
```
# Setup number of CPUs to use (won't effect result, only the run time)
cpus = 4
# Setup number of walkers
walkers = 4
# Setup number of steps each walkers walk
Nsamples = 1000
# Do the sampling (let your walkers walk! MCMC! Woohoo!)
with planet_model:
trace = pm.sample(Nsamples, chains=walkers, cores=cpus)
```
### Explore the results
After the sampling is done, your walker's travel history (parameters they stepped on per step) is saved under `trace`. You can access the travel history of each of you walker's steps on different parameters by `trace['par name']`...
```
trace['mass_j']
```
But a more intuitive way to view the `trace` is to use `arviz` pkg.
ArviZ is a Python package for exploratory analysis of Bayesian models. Includes functions for posterior analysis, data storage, sample diagnostics, model checking, and comparison.
(from [ArviZ](https://arviz-devs.github.io/arviz/) site)
On the left of the plot show the marginalized posterior probability distributions of each of your unknown parameters. On the right, are panels for each unknown parameters showing your walker's travel histories.
```
with planet_model:
az.plot_trace(trace, figsize=(16, 8))
```
You can use the `ArviZ`'s `summary` function to show `trace`'s statistical results.
This is done by the `pandas describe` function in the backend; thus, you can save it out and use it later!
```
with planet_model:
df = az.summary(trace, round_to=2)
df.head()
```
### Plot parameters with Corner
Finally, plotting the corner plot!
In a corner plot, the counter plots at the lower left are showing the posterior distribution between two of your fitting parameters. The more circular (more 2D gaussian shape) the counter plots are, the nicer you fitting results are! The histograms on top of them are the marginalized posterior probability distributions of each the unknown parameters, i.e., their probability distributions.
And you can see that MCMC do help us recover the correct answer to M and R (the location of the blue lines)!
Wonderful!
```{note}
We add "true" values as blue lines here, but we don't generally know the true values. In this toy example, we include true values to see how close MCMC got to our parameters.
```
```
# True values of M and R, which is from the Saturn.
true_R = (58232*u.km).to(u.R_jup).value
true_M = (5.6834e26*u.kg).to(u.M_jup).value
with planet_model:
f, ax = plt.subplots(2, 2, figsize=(4, 4), facecolor='white', dpi=200, gridspec_kw={'hspace': .05, 'wspace': 0.05})
fig = corner.corner(trace, quantiles=[0.16, 0.5, 0.84],
show_titles=True, color='xkcd:olive',
labels=[r"$m$", r"$r$"], title_kwargs={'size':7}, title_fmt='1.3f',
truths=dict(mass_j=true_M, radius_j=true_R),
fig=f)
```
## [Assignment] -- Finding Planet Mass and Radius by choosing good priors!
In this assignment, we will repeat the MCMC analysis above, but explore how **priors** can change our results.
You will get a different set of free-fall data ([link](https://raw.githubusercontent.com/cjtu/spirl/master/spirl/data/position_measurement_as.csv)) to fit for the planet R and M with the same model:
$$z(t) = \frac{1}{2}\frac{GM}{R^2}t^2$$
In each question you will set up the same model as before, but try different priors for R and M:
1. Both with a uniform priors (the worst constraints).
2. Uniform prior for M, and Gaussian prior for R.
3. Both with a Gaussian priors (the best constraints).
The true values of M and R for this data set are:
```python
true_R = (25559*u.km).to(u.R_jup).value
true_M = (8.6810E25*u.kg).to(u.M_jup).value
```
Like before, include these values as blue lines in the final corner plots to see how close your MCMC results are.
Tips:
1. If your resulting corner plot looks weird, try to increase the steps rather than number of walkers (you shouldn't need more than 4 walkers!).
2. If you already have your steps (`Nsamples`) > 1e6, but don't see any difference in results, it's ok to call it quits - which question did this happen for? Why do you think it didn't converge?
```
# read in data
f = "https://raw.githubusercontent.com/cjtu/spirl/master/spirl/data/position_measurement_as.csv"
df = pd.read_csv(f)
# plot the data
f = plt.figure(facecolor='white', figsize=(4,3), dpi=150 )
ax1 = plt.subplot(111)
ax1.errorbar(df['time(s)'], df['z(m)'], yerr = df['z_err(m)'], fmt = '.', ms=5,
lw=0.5, color='tab:blue')
ax1.set_xlabel('time (s)', size='small', fontname='Helvetica')
ax1.set_ylabel('z (m)', size='small', fontname='Helvetica')
plt.show()
Mj2si = const.M_jup.si.value # constant to transfer Jupiter mass to kg
Rj2si = const.R_jup.si.value # constant to transfer Jupiter radius to m
G_cgs = const.G.si.value
t = df['time(s)'].to_numpy()
z = df['z(m)'].to_numpy()
z_err = df['z_err(m)'].to_numpy()
```
### 1. Both with uniform priors:
* Prior for M: $\mathcal{U}(0, 0.1)$
* Prior for R: $\mathcal{U}(0, 1.0)$
Please show both the walker history plot and the corner plot.
```
# [your code here]
```
### 2. Gaussian prior for M, and Uniform prior for R:
* Prior for M: $\mathcal{G}(0.045, 0.02)$
* Prior for R: $\mathcal{U}(0, 1)$
Hint: Can mass be negative? Can you account for this with your choice of prior distribution?
Please show both the walker history plot and the corner plot.
```
# [your code here]
```
### 3. Both with a Gaussian priors:
* Prior for M: $\mathcal{G}(0.045, 0.02)$
* Prior for R: $\mathcal{G}(0.357, 0.05)$
Hint: Can mass or radius be negative? Can you account for this with your choice of prior distribution?
Please show both the walker history plot and the corner plot.
```
# [your code here]
```
### 4. What did we learn about **priors**?
In your Pull Request description, write a sentence or two explaining in your words what you learned about **priors** in this assignment.
| github_jupyter |
```
%matplotlib notebook
import numpy as np
import matplotlib.pyplot as plt
import torch
```
# Redes Convolucionales
[Slides 49-88](https://docs.google.com/presentation/d/1IJ2n8X4w8pvzNLmpJB-ms6-GDHWthfsJTFuyUqHfXg8/edit#slide=id.g3a1a71fe7e_8_192)
# Red Convolucional en PyTorch
Las redes neuronales convolucionales utilizan principalmente tres tipos de capas
## [Capas convolucionales](https://pytorch.org/docs/stable/nn.html#convolution-layers)
- Las neuronas de estas capas se organizan en filtros
- Se realiza la correlación cruzada entre la imagen de entrada y los filtros
- Existen capas convolucionales 1D, 2D y 3D
[Visualización de convoluciones con distintos tamaños, strides, paddings, dilations](https://github.com/vdumoulin/conv_arithmetic)
Los argumentos de la capa convolución de dos dimensiones son:
```python
torch.nn.Conv2d(in_channels, #Cantidad de canales de la imagen de entrada
out_channels, #Cantidad de bancos de filtro
kernel_size, #Tamaño de los filtros (entero o tupla)
stride=1, #Paso de los filtros
padding=0, #Cantidad de filas y columnas para agregar a la entrada antes de filtrar
dilation=1, #Espacio entre los pixeles de los filtros
groups=1, #Configuración cruzada entre filtros de entrada y salida
bias=True, #Utilizar sesgo (b)
padding_mode='zeros' #Especifica como agregar nuevas filas/columnas (ver padding)
)
```
## [Capas de pooling](https://pytorch.org/docs/stable/nn.html#pooling-layers)
- Capa que reduce la dimensión (tamaño) de su entrada
- Se usa tipicamente luego de una capa de convolución "activada"
- Realiza una operación no entrenable:
- Promedio de los píxeles en una región (kernel_size=2, stride=2)
1 2 1 0
2 3 1 2 2.00 1.00
0 1 0 1 0.75 0.25
2 0 0 0
- Máximo de los pixeles en una región (kernel_size=2, stride=2)
1 2 1 0
2 3 1 2 3 2
0 1 0 1 2 1
2 0 0 0
- Estas capas ayudan a reducir la complejidad del modelo
- También otorgan "invarianza local a la traslación", es decir que la posición donde estaba el patrón es menos relevante luego de aplicar pooling
Los argumentos de MaxPooling para entradas de dos dimensiones son:
```python
torch.nn.MaxPool2d(kernel_size, # Mismo significado que en Conv2d
stride=None, # Mismo significado que en Conv2d
padding=0, #Mismo significado que en Conv2d
dilation=1, #Mismo significado que en Conv2d
return_indices=False, #Solo necesario para hacer unpooling
ceil_mode=False #Usar ceil en lugar de floor para calcular el tamaño de la salida
)
```
## [Capas completamente conectadas](https://pytorch.org/docs/stable/nn.html#torch.nn.Linear)
- Idénticas a las usadas en redes tipo MLP
- Realizan la operación: $Z = WX + b$
Los argumentos son:
```python
torch.nn.Linear(in_features, #Neuronas en la entrada
out_features, #Neuronas en la salida
bias=True #Utilizar sesgo (b)
)
```
# [Torchvision](https://pytorch.org/vision/stable/index.html)
Es una librería utilitaria de PyTorch que facilita considerablemente el trabajo con imágenes
- Funcionalidad para descargar sets de benchmark: MNIST, CIFAR, IMAGENET, ...
- Modelos clásicos pre-entrenados: AlexNet, VGG, GoogLeNet, ResNet
- Funciones para importar imágenes en distintos formatos
- Funciones de transformación para hacer aumentación de datos en imágenes
## Ejemplo: Base de datos de imágenes de dígitos manuscritos MNIST
- Imágenes de 28x28 píxeles en escala de grises
- Diez categorías: Dígitos manuscritos del cero al nueve
- 60.000 imágenes de entrenamiento, 10.000 imágenes de prueba
- Por defecto las imágenes vienen en [formato PIL](https://pillow.readthedocs.io/en/stable/) (entero 8bit), usamos la transformación [`ToTensor()`](https://pytorch.org/docs/stable/torchvision/transforms.html#torchvision.transforms.ToTensor) para convertirla a tensor en float32
```
import torchvision
mnist_train_data = torchvision.datasets.MNIST(root='~/datasets/',
train=True, download=True,
transform=torchvision.transforms.ToTensor())
mnist_test_data = torchvision.datasets.MNIST(root='~/datasets/',
train=False, download=True,
transform=torchvision.transforms.ToTensor())
image, label = mnist_train_data[0]
display(len(mnist_train_data), type(image), image.dtype, type(label))
fig, ax = plt.subplots(1, 10, figsize=(8, 1.5), tight_layout=True)
idx = np.random.permutation(len(mnist_train_data))[:10]
for k in range(10):
image, label = mnist_train_data[idx[k]]
ax[k].imshow(image[0, :, :].numpy(), cmap=plt.cm.Greys_r)
ax[k].axis('off');
ax[k].set_title(label)
image.shape
```
## Dataloaders
Creamos dataloaders de entrenamiento y validación
```
from torch.utils.data import Subset, DataLoader
import sklearn.model_selection
# Set de entrenamiento y validación estratíficados
sss = sklearn.model_selection.StratifiedShuffleSplit(train_size=0.75).split(mnist_train_data.data,
mnist_train_data.targets)
train_idx, valid_idx = next(sss)
# Data loader de entrenamiento
train_dataset = Subset(mnist_train_data, train_idx)
train_loader = DataLoader(train_dataset, shuffle=True, batch_size=32)
# Data loader de validación
valid_dataset = Subset(mnist_train_data, valid_idx)
valid_loader = DataLoader(valid_dataset, shuffle=False, batch_size=256)
```
# Mi primera red convolucional para clasificar en pytorch
Clasificaremos la base de datos MNIST
Para esto implementaremos la clásica arquitectura Lenet5
<img src="img/LeNet5.png" width="800">
La arquitectura considera
- Dos capas convolucionales con 6 y 16 bancos de filtros, respectivamente
- Las capas convolucionales usan filtros de 5x5 píxeles
- Se usa max-pooling de tamaño 2x2 y stride 2
- La primera capa convolucional espera un minibatch de imágenes de 1 canal (blanco y negro)
- Usaremos la función de activación [Rectified Linear Unit (ReLU)](https://pytorch.org/docs/stable/nn.html#relu)
- Se usan tres capas completamente conectadas con 120, 84 y 10 neuronas, respectivamente
> Podemos usar `reshape` o `view` para convertir un tensor de 4 dimensiones a dos dimensiones. Esto prepara un tensor que sale de una capa convolucional (o pooling) para ingresarlo a las capas completamente conectadas
```
import torch.nn as nn
class Lenet5(nn.Module):
def __init__(self):
super(type(self), self).__init__()
# La entrada son imágenes de 1x32x32
self.features = nn.Sequential(nn.Conv2d(1, 6, 5, padding=2),
nn.ReLU(),
nn.MaxPool2d(2),
nn.Conv2d(6, 16, 5),
nn.ReLU(),
nn.MaxPool2d(2))
self.classifier = nn.Sequential(nn.Linear(16*5*5, 120),
nn.ReLU(),
nn.Linear(120, 84),
nn.ReLU(),
nn.Linear(84, 10))
def forward(self, x):
z = self.features(x)
# Esto es de tamaño Mx16x5x5
z = z.view(-1, 16*5*5)
# Esto es de tamaño Mx400
return self.classifier(z)
model = Lenet5()
print(model)
```
### Clasificación multiclase en PyTorch
Para hacer clasificación con **más de dos categorías** usamos la [entropía cruzada](https://pytorch.org/docs/stable/nn.html#torch.nn.CrossEntropyLoss)
```python
torch.nn.CrossEntropyLoss()
```
- Si el problema de clasificación es de $M$ categorías la última capa de la red debe tener $M$ neuronas
- Adicionalmente no se debe usar función de activación ya que `CrossEntropyLoss` la aplica de forma interna
Para obtener las probabilidades de salida de la red debemos aplicar de forma manual
- `torch.nn.Softmax(dim=1)`
- `torch.nn.LogSoftmax(dim=1)`
a la última capa
Si sólo queremos saber cual es la clase más probable podemos usar el atributo `argmax(dim=1)` sobre la salida de la red (con o sin activar)
### Gradiente descendente con paso adaptivo
Para acelerar el entrenamiento podemos usar un algoritmo de [gradiente descendente con paso adaptivo](https://arxiv.org/abs/1609.04747)
Un ejemplo ampliamente usado es [Adam](https://arxiv.org/abs/1412.6980)
- Se utiliza la historia de los gradientes
- Se utiliza momentum (inercia)
- Cada parámetro tiene un paso distinto
```python
torch.optim.Adam(params, #Parámetros de la red neuronal
lr=0.001, #Tasa de aprendizaje inicial
betas=(0.9, 0.999), #Factores de olvido de los gradientes históricos
eps=1e-08, #Término para evitar división por cero
weight_decay=0, #Regulariza los pesos de la red si es mayor que cero
amsgrad=False #Corrección para mejorar la convergencia de Adam en ciertos casos
)
```
**Atención**
Esta es un área de investigación activa. [Papers recientes indican que Adam llega a un óptimo más rápido que SGD, pero ese óptimo podría no ser mejor que el obtenido por SGD](https://arxiv.org/abs/1712.07628)
> Siempre prueba tus redes con distintos optimizadores
# Entrenamiento de la red convolucional
- Si tenemos acceso a una GPU podemos usar el atributo `.cuda()` o `.to()` para enviar el modelo y los datos a la GPU para acelerar los cálculos
- Actualizamos los parámetros en el conjunto de entrenamiento
- Medimos la convergencia en el conjunto de validación
- Guardamos el modelo con mejor error de validación
- Usaremos `ignite` y `Tensorboard` para entrenar. Guardaremos los entrenamientos en `/tmp/tensorboard/`
```
from ignite.engine import Engine, Events
from ignite.metrics import Loss, Accuracy
model = Lenet5()
optimizer = torch.optim.Adam(model.parameters(), lr=1e-3)
criterion = torch.nn.CrossEntropyLoss(reduction='sum')
max_epochs = 100
device = torch.device('cpu')
#device = torch.device('cuda:0')
model = model.to(device)
# Esto es lo que hace el engine de entrenamiento
def train_one_step(engine, batch):
optimizer.zero_grad()
x, y = batch
x, y = x.to(device), y.to(device)
yhat = model.forward(x)
loss = criterion(yhat, y)
loss.backward()
optimizer.step()
return loss.item() # Este output puede llamar luego como trainer.state.output
# Esto es lo que hace el engine de evaluación
def evaluate_one_step(engine, batch):
with torch.no_grad():
x, y = batch
x, y = x.to(device), y.to(device)
yhat = model.forward(x)
#loss = criterion(yhat, y)
return yhat, y
trainer = Engine(train_one_step)
evaluator = Engine(evaluate_one_step)
metrics = {'Loss': Loss(criterion), 'Acc': Accuracy()}
for name, metric in metrics.items():
metric.attach(evaluator, name)
import time
from torch.utils.tensorboard import SummaryWriter
from ignite.handlers import ModelCheckpoint
# Contexto de escritura de datos para tensorboard
with SummaryWriter(log_dir=f'/tmp/tensorboard/lenet5_mnist_{time.time_ns()}') as writer:
@trainer.on(Events.EPOCH_COMPLETED(every=1)) # Cada 1 epocas
def log_results(engine):
# Evaluo el conjunto de entrenamiento
evaluator.run(train_loader)
writer.add_scalar("train/loss", evaluator.state.metrics['Loss'], engine.state.epoch)
writer.add_scalar("train/accy", evaluator.state.metrics['Acc'], engine.state.epoch)
# Evaluo el conjunto de validación
evaluator.run(valid_loader)
writer.add_scalar("valid/loss", evaluator.state.metrics['Loss'], engine.state.epoch)
writer.add_scalar("valid/accy", evaluator.state.metrics['Acc'], engine.state.epoch)
# Guardo el mejor modelo en validación
best_model_handler = ModelCheckpoint(dirname='.', require_empty=False, filename_prefix="best", n_saved=1,
score_function=lambda engine: -engine.state.metrics['Loss'],
score_name="val_loss")
# Lo siguiente se ejecuta cada ves que termine el loop de validación
evaluator.add_event_handler(Events.COMPLETED,
best_model_handler, {'lenet5': model})
trainer.run(train_loader, max_epochs=max_epochs)
```
## Evaluando la red en el conjunto de test
Primero recuperamos la red con menor costo de validación
```
model = Lenet5()
model.load_state_dict(torch.load('best_lenet5_val_loss=-2.6935.pt'))
```
Haremos la evaluación final del a red en el conjunto de prueba/test
Iteramos sobre el conjunto y guardamos las predicciones de la red
Con esto podemos construir una matriz de confusión y un reporte usando las herramientas de `sklearn`
```
test_loader = DataLoader(mnist_test_data, shuffle=False, batch_size=512)
test_targets = mnist_test_data.targets.numpy()
prediction_test = []
entropy = []
for mbdata, label in test_loader:
logits = model.forward(mbdata)
probs = torch.nn.Softmax(dim=1)(logits)
entropy.append(-(logits*probs).sum(1).detach().numpy())
prediction_test.append(logits.argmax(dim=1).detach().numpy())
prediction_test = np.concatenate(prediction_test)
entropy = np.concatenate(entropy)
from sklearn.metrics import confusion_matrix, classification_report
def plot_confusion_matrix(cm, labels, cmap=plt.cm.Blues):
fig, ax = plt.subplots(figsize=(5, 5), tight_layout=True)
ax.imshow(cm, interpolation='nearest', cmap=cmap)
for i in range(cm.shape[1]):
for j in range(cm.shape[0]):
ax.text(j, i, "{:,}".format(cm[i, j]),
horizontalalignment="center", verticalalignment="center",
color="white" if cm[i, j] > np.amax(cm)/2 else "black")
ax.set_title("Matriz de confusión")
tick_marks = np.arange(len(labels))
plt.xticks(tick_marks, labels)
plt.yticks(tick_marks, labels)
plt.ylabel('Etiqueta real')
plt.xlabel('Predicción')
cm = confusion_matrix(y_true=test_targets, y_pred=prediction_test)
plot_confusion_matrix(cm, labels=[str(i) for i in range(10)])
print(classification_report(test_targets, prediction_test, digits=3))
```
## Análisis de errores
Luego de evaluar la red podemos estudiar sus errores con el objeto de mejorar el modelo
Para problemas con imágenes es muy recomendable visualizar los ejemplos mal predichos por la red
Esto podría revelar
- Imágenes mal etiquetadas: Podemos cambiar su etiqueta y re-entrenar/re-evaluar
- Errores sistemáticos del modelo: Por ejemplo que siempre se equivoque con una clase u objeto en particular
Observemos algunos ejemplos mal clasificados
- Las imágenes corresponden a `digit` que no fueron predichos como `digit`
- El título de la imagen tiene la etiqueta predicha por la red
```
digit = 8
idx = np.where((test_targets == digit) & ~(prediction_test == digit))[0]
fig, ax = plt.subplots(1, 10, figsize=(8, 1.5), tight_layout=True)
for i in range(10):
ax[i].imshow(mnist_test_data[idx[i]][0].numpy()[0, :, :], cmap=plt.cm.Greys_r)
ax[i].set_title(prediction_test[idx[i]])
ax[i].axis('off')
```
Los ejemplos más inciertos, es decir donde el modelo está "más confundido", son aquellos con mayor entropía en sus predicciones
```
np.argsort(entropy)[::-1][:10]
```
También es interesante analizar las probabilidades asignadas por la red. Para esto debemos aplicar activación Softmax a la salida. En particular podemos analizar los ejemplos de mayor confusión
```
image, label = mnist_test_data[1247]
# Usamos unsqueeze para convertirlo en un minibatch de 1 elemento:
y = torch.nn.Softmax(dim=1)(model.forward(image.unsqueeze(0)))
fig, ax = plt.subplots(1, 2, figsize=(5, 2), tight_layout=True)
ax[0].bar(range(10), height=y.detach().numpy()[0])
ax[0].set_xticks(range(10))
ax[1].set_title("Etiqueta: %d" %(label))
ax[1].imshow(image.numpy()[0, :, :], cmap=plt.cm.Greys_r);
ax[1].axis('off');
```
| github_jupyter |
找下daily的index差哪一个
{
"v" : 2,
"key" : {
"_id" : 1
},
"name" : "_id_",
"ns" : "quant_01.daily"
},
{
"v" : 2,
"key" : {
"code" : 1,
"date" : 1,
"index" : 1
},
"name" : "code_1_date_1_index_1",
"ns" : "quant_01.daily"
},
{
"v" : 2,
"key" : {
"code" : 1,
"date" : 1
},
"name" : "code_1_date_1",
"ns" : "quant_01.daily"
},
{
"v" : 2,
"key" : {
"date" : 1,
"pe" : 1,
"index" : 1,
"is_trading" : 1
},
"name" : "date_1_pe_1_index_1_is_trading
"ns" : "quant_01.daily"
},
{
"v" : 2,
"key" : {
"code" : 1,
"date" : 1,
"is_trading" : 1
},
"name" : "code_1_date_1_is_trading_1",
"ns" : "quant_01.daily"
},
[
{
"v" : 2,
"key" : {
"_id" : 1
},
"name" : "_id_",
"ns" : "quant_01.daily"
},
{
"v" : 2,
"key" : {
"code" : 1,
"date" : 1,
"index" : 1
},
"name" : "code_1_date_1_index_1",
"ns" : "quant_01.daily"
},
{
"v" : 2,
"key" : {
"code" : 1,
"date" : 1
},
"name" : "code_1_date_1",
"ns" : "quant_01.daily"
},
{
"v" : 2,
"key" : {
"date" : 1,
"pe" : 1,
"is_trading" : 1
},
"name" : "date_1_pe_1_is_trading_1",
"ns" : "quant_01.daily"
},
{
"v" : 2,
"key" : {
"code" : 1,
"date" : 1,
"is_trading" : 1
},
"name" : "code_1_date_1_is_trading_1",
"ns" : "quant_01.daily"
},
{
"v" : 2,
"key" : {
"date" : 1,
"pe" : 1,
"index" : 1,
"is_trading" : 1
},
"name" : "date_1_pe_1_index_1_is_trading_1",
"ns" : "quant_01.daily"
},
{
"v" : 2,
"key" : {
"code" : 1,
"date" : 1,
"index" : 1,
"is_trading" : 1
},
"name" : "code_1_date_1_index_1_is_trading_1",
"ns" : "quant_01.daily"
}
]
spyder 注释代码的颜色,所有的颜色代码都看一下吧
choice的login token文件
1. 基于第五课的课件,添加一些条件;
2. 涨跌停限制;
3. 止损
4. 233, 貌似就这些,感觉还是很儿戏啊。。。
1. 回测发现加入了止损后,总资产反而下降了;
1. √ 在surface上测试了第8课的代码,用compare对比第5课和第8课的代码区别, 使用第8课代码中的一些计算指标; 胜率怎么办, 2333
1. 将回测时间设定为2016-01-01~2018-06-30, 和前几次作业来个大对比; 看看不同策略在相同时间下的表现;
1. √ 加入仓位图;
1. √ 调整10%回测比率, 比如改成5%;
1. √ 使用固定比例浮动止损;
1. √ 使用波动率止损;
1. 使用均匀加仓;
1. 使用波动率分配头寸; 1% 是乘以初始金额还是剩下的金额
1. 使用止盈;
1. 使用波动率浮动止盈;
回测结果的保存和分析,现在可以保存到文件里了,但我想有更好的展示,估计会很麻烦;
但是可以做到的是,先批量读取,然后做成dataframe,最后输出成excel表
1. 首先是跑通第五课的代码;
2. 求涨跌停价格并保存到不复权的表中;
3. 有了这些条件,然后再在回测框架中添加涨跌停限制和止损
先看看目前的进度
查看有哪些database,自己想看的database下面有哪些collection;
```mongodb
show dbs
use quant_01
show collection
```
需要看下,比如daily和daily_hfq有哪些字段,分别有多少条数据
daily有的字段是
code date index ohlc volume
daily_hfq
亦是一样的
也就是说没有做除权因子等操作,也没有记录pre_close
也没有补全信息, and so on
当然也没有做涨跌停的价格录入
回测代码中也有不少需要加索引的地方
先梳理出执行顺序
再确认各个文件里需要index的地方,不光是爬取数据的地方,回测的地方也需要index
database.py / stock_util.py --> daily_crawler.py -->
basic_crawler.py --> daily_fixing.py -->
compute_high_low_limit.py -->
finance_report.py --> pe_computing.py -->
stock_pool_strategy.py --> backtest.py
```
可以同时运行的有哪些数据呢
daily只是ohlc基本数据
basic_crawler是获取basic的数据
daily_fixing是补充is_trading字段、停牌时的数据、复权因子 和 pre_close
compute_high_low_limit是为了计算涨跌停价格,所以需要前面的都完成以后才能计算;
finance_report是一个独立的过程,可以与其他的过程同时进行
pe_computing需要daily中的close和report的eps,在daily数据和finance_report数据之后才能计算,但是不知道别的数据库在更新daily的时候,会不会影响效率
233,pe的计算果然够慢,如果可能,需要尽快进行哦
```
```
【编程题】为了让回测更接近于实际情况,需要在第5课中回测流程中引入以下逻辑:
1、引入涨停和跌停的判断,跌停时无法卖出,涨停时无法买入;
2、引入止损的风险控制,如果单日亏损超过3%或者累积亏损超过10%则第二天开盘价卖出。
要求编写完整的可执行代码。自行选定回测周期,附回测结果图。
```
获取股票基本信息,使用新的接口
tushare pro 哦
可能会造成各种不适应
```
import QUANTAXIS as QA
df = QA.QA_fetch_get_stock_min('tdx', '300257', '2015-01-01', '2018-10-26')
import tushare as ts
print(ts.__version__)
token = 'd5deeb2574ff968fbf7cfc89a379a1a2597b2ab6dc56989053ec32a9'
ts.set_token(token)
pro = ts.pro_api()
df = pro.trade_cal(exchange_id='', start_date='20180901', end_date='20181001', fields='exchange_id,cal_date,is_open,pretrade_date', is_open='0')
trade_dates = QA.trade_date_sse
QA.QAUtil.*date*?
trade_range = QA.QA_util_get_trade_range('2015-01-01', '2017-12-31')
ts.get_stock_basics('2015-01-01')
for _date in trade_range:
if ts.get_stock_basics(_date) is not None:
print(_date)
break
QA.QA_util_get_last_day('2016-08-09')
```
针对backtest进行debug,看到底是哪里
重构涨跌停计算方式
```
df = ts.get_stock_basics()
df.head()
start = '2015-01-01'
end = '2017-12-31'
codes = ts.get_stock_basics().index.tolist()
codes[:10]
code = codes[1000]
code
from database import DB_CONN
sample = np.random.choice(codes, 10)
sample = np.random.choice(codes, 20).tolist()
sample
```
如果初始日就是
```
st_mark = ['st', 'ST', '*st', '*ST']
total = len(sample)
for i,code in enumerate(sample):
try:
timeToMarket = DB_CONN['basic'].find_one({'code':code},
projection={'code':True, 'timeToMarket':True, '_id':False})['timeToMarket']
except:
continue
daily_cursor = DB_CONN['daily'].find(
{'code':code, 'date':{'$lte': end, '$gte': timeToMarket}, 'index':False},
projection={'code':True, 'date':True, 'pre_close':True, '_id':False})
for j,daily in enumerate(daily_cursor):
date = daily['date']
try:
pre_close = daily['pre_close']
except:
if (j == 0) & (timeToMarket != date):
pass
# print('code: %s, time: %s, 数据初始日没有pre_close' % (code, date))
elif timeToMarket == date:
issueprice = DB_CONN['basic'].find_one({'code':code},
projection={'issueprice':True, '_id':False})['issueprice']
high_limit = np.round(np.round(issueprice * 1.2, 2) * 1.2, 2)
low_limit = np.round(np.round(issueprice * 0.8, 2) * 0.8, 2)
else:
print('code: %s, time: %s, ipo_date: %s, 请速查原因' % (code, date, timeToMarket))
continue
# print('code: %s, date: %s' % (code, date))
if date < '2016-08-09':
_date = '2016-08-09'
else:
_date = date
try:
name = DB_CONN['basic'].find_one({'code':code, 'date':_date},
projection={'name':True, '_id':False})['name']
last_name = name
except:
if i == 0:
name = DB_CONN['basic'].find_one({'code':code},
projection={'name':True, '_id':False})['name']
last_name = name
else:
name = last_name
# if timeToMarket == date:
# issueprice = DB_CONN['basic'].find_one({'code':code},
# projection={'issueprice':True, '_id':False})['issueprice']
# high_limit = np.round(np.round(issueprice * 1.2, 2) * 1.2, 2)
# low_limit = np.round(np.round(issueprice * 0.8, 2) * 0.8, 2)
if (name[:2] in st_mark) or (name[:3] in st_mark) :
high_limit = np.round(pre_close * 1.05, 2)
low_limit = np.round(pre_close * 0.95, 2)
else:
high_limit = np.round(pre_close * 1.1, 2)
low_limit = np.round(pre_close * 0.9, 2)
print('stock: %s high low limit complish, 进度: (%s/%s)' % (code, i+1, total))
code = '000068'
timeToMarket = DB_CONN['basic'].find_one({'code':code},
projection={'code':True, 'timeToMarket':True, '_id':False})['timeToMarket']
timeToMarket
daily_cursor = DB_CONN['daily'].find(
{'code':code, 'date':{'$lte': end, '$gte': timeToMarket}, 'index':False},
projection={'code':True, 'date':True, 'pre_close':True, '_id':False})
result = [i for i in daily_cursor]
result
code = '000001'
name_result = [d for d in DB_CONN['basic'].find({'code':code},
projection={'name':True, 'date':True, '_id':False})]
name_result
df = ts.get_stock_basics('2016-08-09')
df.loc[code, :]
check_data = QA.QA_fetch_stock_day_adv(code, start, end).data
check_data.head(10)
```
这个bug的原因是基础数据获取的不够完整,不是每一个交易日都有数据
```
'603214' in df.index
df.loc['603214',:]
_date
code
DB_CONN['basic'].find_one({'code':code, 'date':_date},
projection={'name':True, '_id':False})['name']
test_cursor = DB_CONN['basic'].find({'code':code},
projection={'name':True, 'date':True, '_id':False})
result = [i for i in test_cursor]
result[:10]
# test_cursor = DB_CONN['basic'].find({'code':code},
# projection={'name':True, 'date':True, '_id':False})
# result = [i for i in test_cursor]
# result
DB_CONN['basic'].find_one({'code':code},
projection={'name':True, 'date':True, '_id':False})
# daily_cursor = DB_CONN['daily'].find(
# {'code':code, 'date':{'$lte': end, '$gte': timeToMarket}, 'index':False},
# projection={'code':True, 'date':True, 'pre_close':True, '_id':False})
# dailies = [i for i in daily_cursor]
# dailies
df = pd.read_excel('ipo_info.xlsx', header=0, dtype={'code':str})
df.head()
```
| github_jupyter |
# Constraint Satisfaction Problems
---
# Heuristics for Arc-Consistency Algorithms
## Introduction
A ***Constraint Satisfaction Problem*** is a triple $(X,D,C)$ where:
- $X$ is a set of variables $X_1, …, X_n$;
- $D$ is a set of domains $D_1, …, D_n$, one for each variable and each of which consists of a set of allowable values $v_1, ..., v_k$;
- $C$ is a set of constraints that specify allowable combinations of values.
A CSP is called *arc-consistent* if every value in the domain of every variable is supported by all the neighbors of the variable while, is called *inconsistent*, if it has no solutions. <br>
***Arc-consistency algorithms*** remove all unsupported values from the domains of variables making the CSP *arc-consistent* or decide that a CSP is *inconsistent* by finding that some variable has no supported values in its domain. <br>
Heuristics significantly enhance the efficiency of the *arc-consistency algorithms* improving their average performance in terms of *consistency-checks* which can be considered a standard measure of goodness for such algorithms. *Arc-heuristic* operate at arc-level and selects the constraint that will be used for the next check, while *domain-heuristics* operate at domain-level and selects which values will be used for the next support-check.
```
from csp import *
```
## Domain-Heuristics for Arc-Consistency Algorithms
In <a name="ref-1"/>[[1]](#cite-van2002domain) are investigated the effects of a *domain-heuristic* based on the notion of a *double-support check* by studying its average time-complexity.
The objective of *arc-consistency algorithms* is to resolve some uncertainty; it has to be know, for each $v_i \in D_i$ and for each $v_j \in D_j$, whether it is supported.
A *single-support check*, $(v_i, v_j) \in C_{ij}$, is one in which, before the check is done, it is already known that either $v_i$ or $v_j$ are supported.
A *double-support check* $(v_i, v_j) \in C_{ij}$, is one in which there is still, before the check, uncertainty about the support-status of both $v_i$ and $v_j$.
If a *double-support check* is successful, two uncertainties are resolved. If a *single-support check* is successful, only one uncertainty is resolved. A good *arc-consistency algorithm*, therefore, would always choose to do a *double-support check* in preference of a *single-support check*, because the cormer offers the potential higher payback.
The improvement with *double-support check* is that, where possible, *consistency-checks* are used to find supports for two values, one value in the domain of each variable, which were previously known to be unsupported. It is motivated by the insight that *in order to minimize the number of consistency-checks it is necessary to maximize the number of uncertainties which are resolved per check*.
### AC-3b: an improved version of AC-3 with Double-Support Checks
As shown in <a name="ref-2"/>[[2]](#cite-van2000improving) the idea is to use *double-support checks* to improve the average performance of `AC3` which does not exploit the fact that relations are bidirectional and results in a new general purpose *arc-consistency algorithm* called `AC3b`.
```
%psource AC3
%psource revise
```
At any stage in the process of making 2-variable CSP *arc-consistent* in `AC3b`:
- there is a set $S_i^+ \subseteq D_i$ whose values are all known to be supported by $X_j$;
- there is a set $S_i^? = D_i \setminus S_i^+$ whose values are unknown, as yet, to be supported by $X_j$.
The same holds if the roles for $X_i$ and $X_j$ are exchanged.
In order to establish support for a value $v_i^? \in S_i^?$ it seems better to try to find a support among the values in $S_j^?$ first, because for each $v_j^? \in S_j^?$ the check $(v_i^?,v_j^?) \in C_{ij}$ is a *double-support check* and it is just as likely that any $v_j^? \in S_j^?$ supports $v_i^?$ than it is that any $v_j^+ \in S_j^+$ does. Only if no support can be found among the elements in $S_j^?$, should the elements $v_j^+$ in $S_j^+$ be used for *single-support checks* $(v_i^?,v_j^+) \in C_{ij}$. After it has been decided for each value in $D_i$ whether it is supported or not, either $S_x^+ = \emptyset$ and the 2-variable CSP is *inconsistent*, or $S_x^+ \neq \emptyset$ and the CSP is *satisfiable*. In the latter case, the elements from $D_i$ which are supported by $j$ are given by $S_x^+$. The elements in $D_j$ which are supported by $x$ are given by the union of $S_j^+$ with the set of those elements of $S_j^?$ which further processing will show to be supported by some $v_i^+ \in S_x^+$.
```
%psource AC3b
%psource partition
```
`AC3b` is a refinement of the `AC3` algorithm which consists of the fact that if, when arc $(i,j)$ is being processed and the reverse arc $(j,i)$ is also in the queue, then consistency-checks can be saved because only support for the elements in $S_j^?$ has to be found (as opposed to support for all the elements in $D_j$ in the
`AC3` algorithm). <br>
`AC3b` inherits all its properties like $\mathcal{O}(ed^3)$ time-complexity and $\mathcal{O}(e + nd)$ space-complexity fron `AC3` and where $n$ denotes the number of variables in the CSP, $e$ denotes the number of binary constraints and $d$ denotes the maximum domain-size of the variables.
## Arc-Heuristics for Arc-Consistency Algorithms
Many *arc-heuristics* can be devised, based on three major features of CSPs:
- the number of acceptable pairs in each constraint (the *constraint size* or *satisfiability*);
- the *domain size*;
- the number of binary constraints that each variable participates in, equal to the *degree* of the node of that variable in the constraint graph.
Simple examples of heuristics that might be expected to improve the efficiency of relaxation are:
- ordering the list of variable pairs by *increasing* relative *satisfiability*;
- ordering by *increasing size of the domain* of the variable $v_j$ relaxed against $v_i$;
- ordering by *descending degree* of node of the variable relaxed.
In <a name="ref-3"/>[[3]](#cite-wallace1992ordering) are investigated the effects of these *arc-heuristics* in an empirical way, experimenting the effects of them on random CSPs. Their results demonstrate that the first two, later called `sat up` and `dom j up` for n-ary and binary CSPs respectively, significantly reduce the number of *consistency-checks*.
```
%psource dom_j_up
%psource sat_up
```
## Experimental Results
For the experiments below on binary CSPs, in addition to the two *arc-consistency algorithms* already cited above, `AC3` and `AC3b`, the `AC4` algorithm was used. <br>
The `AC4` algorithm runs in $\mathcal{O}(ed^2)$ worst-case time but can be slower than `AC3` on average cases.
```
%psource AC4
```
### Sudoku
#### Easy Sudoku
```
sudoku = Sudoku(easy1)
sudoku.display(sudoku.infer_assignment())
%time _, checks = AC3(sudoku, arc_heuristic=no_arc_heuristic)
f'AC3 needs {checks} consistency-checks'
sudoku = Sudoku(easy1)
%time _, checks = AC3b(sudoku, arc_heuristic=no_arc_heuristic)
f'AC3b needs {checks} consistency-checks'
sudoku = Sudoku(easy1)
%time _, checks = AC4(sudoku, arc_heuristic=no_arc_heuristic)
f'AC4 needs {checks} consistency-checks'
sudoku = Sudoku(easy1)
%time _, checks = AC3(sudoku, arc_heuristic=dom_j_up)
f'AC3 with DOM J UP arc heuristic needs {checks} consistency-checks'
sudoku = Sudoku(easy1)
%time _, checks = AC3b(sudoku, arc_heuristic=dom_j_up)
f'AC3b with DOM J UP arc heuristic needs {checks} consistency-checks'
sudoku = Sudoku(easy1)
%time _, checks = AC4(sudoku, arc_heuristic=dom_j_up)
f'AC4 with DOM J UP arc heuristic needs {checks} consistency-checks'
backtracking_search(sudoku, select_unassigned_variable=mrv, inference=forward_checking)
sudoku.display(sudoku.infer_assignment())
```
#### Harder Sudoku
```
sudoku = Sudoku(harder1)
sudoku.display(sudoku.infer_assignment())
%time _, checks = AC3(sudoku, arc_heuristic=no_arc_heuristic)
f'AC3 needs {checks} consistency-checks'
sudoku = Sudoku(harder1)
%time _, checks = AC3b(sudoku, arc_heuristic=no_arc_heuristic)
f'AC3b needs {checks} consistency-checks'
sudoku = Sudoku(harder1)
%time _, checks = AC4(sudoku, arc_heuristic=no_arc_heuristic)
f'AC4 needs {checks} consistency-checks'
sudoku = Sudoku(harder1)
%time _, checks = AC3(sudoku, arc_heuristic=dom_j_up)
f'AC3 with DOM J UP arc heuristic needs {checks} consistency-checks'
sudoku = Sudoku(harder1)
%time _, checks = AC3b(sudoku, arc_heuristic=dom_j_up)
f'AC3b with DOM J UP arc heuristic needs {checks} consistency-checks'
sudoku = Sudoku(harder1)
%time _, checks = AC4(sudoku, arc_heuristic=dom_j_up)
f'AC4 with DOM J UP arc heuristic needs {checks} consistency-checks'
backtracking_search(sudoku, select_unassigned_variable=mrv, inference=forward_checking)
sudoku.display(sudoku.infer_assignment())
```
### 8 Queens
```
chess = NQueensCSP(8)
chess.display(chess.infer_assignment())
%time _, checks = AC3(chess, arc_heuristic=no_arc_heuristic)
f'AC3 needs {checks} consistency-checks'
chess = NQueensCSP(8)
%time _, checks = AC3b(chess, arc_heuristic=no_arc_heuristic)
f'AC3b needs {checks} consistency-checks'
chess = NQueensCSP(8)
%time _, checks = AC4(chess, arc_heuristic=no_arc_heuristic)
f'AC4 needs {checks} consistency-checks'
chess = NQueensCSP(8)
%time _, checks = AC3(chess, arc_heuristic=dom_j_up)
f'AC3 with DOM J UP arc heuristic needs {checks} consistency-checks'
chess = NQueensCSP(8)
%time _, checks = AC3b(chess, arc_heuristic=dom_j_up)
f'AC3b with DOM J UP arc heuristic needs {checks} consistency-checks'
chess = NQueensCSP(8)
%time _, checks = AC4(chess, arc_heuristic=dom_j_up)
f'AC4 with DOM J UP arc heuristic needs {checks} consistency-checks'
backtracking_search(chess, select_unassigned_variable=mrv, inference=forward_checking)
chess.display(chess.infer_assignment())
```
For the experiments below on n-ary CSPs, due to the n-ary constraints, the `GAC` algorithm was used. <br>
The `GAC` algorithm has $\mathcal{O}(er^2d^t)$ time-complexity and $\mathcal{O}(erd)$ space-complexity where $e$ denotes the number of n-ary constraints, $r$ denotes the constraint arity and $d$ denotes the maximum domain-size of the variables.
```
%psource ACSolver.GAC
```
### Crossword
```
crossword = Crossword(crossword1, words1)
crossword.display()
words1
%time _, _, checks = ACSolver(crossword).GAC(arc_heuristic=no_heuristic)
f'GAC needs {checks} consistency-checks'
crossword = Crossword(crossword1, words1)
%time _, _, checks = ACSolver(crossword).GAC(arc_heuristic=sat_up)
f'GAC with SAT UP arc heuristic needs {checks} consistency-checks'
crossword.display(ACSolver(crossword).domain_splitting())
```
### Kakuro
#### Easy Kakuro
```
kakuro = Kakuro(kakuro2)
kakuro.display()
%time _, _, checks = ACSolver(kakuro).GAC(arc_heuristic=no_heuristic)
f'GAC needs {checks} consistency-checks'
kakuro = Kakuro(kakuro2)
%time _, _, checks = ACSolver(kakuro).GAC(arc_heuristic=sat_up)
f'GAC with SAT UP arc heuristic needs {checks} consistency-checks'
kakuro.display(ACSolver(kakuro).domain_splitting())
```
#### Medium Kakuro
```
kakuro = Kakuro(kakuro3)
kakuro.display()
%time _, _, checks = ACSolver(kakuro).GAC(arc_heuristic=no_heuristic)
f'GAC needs {checks} consistency-checks'
kakuro = Kakuro(kakuro3)
%time _, _, checks = ACSolver(kakuro).GAC(arc_heuristic=sat_up)
f'GAC with SAT UP arc heuristic needs {checks} consistency-checks'
kakuro.display(ACSolver(kakuro).domain_splitting())
```
#### Harder Kakuro
```
kakuro = Kakuro(kakuro4)
kakuro.display()
%time _, _, checks = ACSolver(kakuro).GAC()
f'GAC needs {checks} consistency-checks'
kakuro = Kakuro(kakuro4)
%time _, _, checks = ACSolver(kakuro).GAC(arc_heuristic=sat_up)
f'GAC with SAT UP arc heuristic needs {checks} consistency-checks'
kakuro.display(ACSolver(kakuro).domain_splitting())
```
### Cryptarithmetic Puzzle
$$
\begin{array}{@{}r@{}}
S E N D \\
{} + M O R E \\
\hline
M O N E Y
\end{array}
$$
```
cryptarithmetic = NaryCSP(
{'S': set(range(1, 10)), 'M': set(range(1, 10)),
'E': set(range(0, 10)), 'N': set(range(0, 10)), 'D': set(range(0, 10)),
'O': set(range(0, 10)), 'R': set(range(0, 10)), 'Y': set(range(0, 10)),
'C1': set(range(0, 2)), 'C2': set(range(0, 2)), 'C3': set(range(0, 2)),
'C4': set(range(0, 2))},
[Constraint(('S', 'E', 'N', 'D', 'M', 'O', 'R', 'Y'), all_diff),
Constraint(('D', 'E', 'Y', 'C1'), lambda d, e, y, c1: d + e == y + 10 * c1),
Constraint(('N', 'R', 'E', 'C1', 'C2'), lambda n, r, e, c1, c2: c1 + n + r == e + 10 * c2),
Constraint(('E', 'O', 'N', 'C2', 'C3'), lambda e, o, n, c2, c3: c2 + e + o == n + 10 * c3),
Constraint(('S', 'M', 'O', 'C3', 'C4'), lambda s, m, o, c3, c4: c3 + s + m == o + 10 * c4),
Constraint(('M', 'C4'), eq)])
%time _, _, checks = ACSolver(cryptarithmetic).GAC(arc_heuristic=no_heuristic)
f'GAC needs {checks} consistency-checks'
%time _, _, checks = ACSolver(cryptarithmetic).GAC(arc_heuristic=sat_up)
f'GAC with SAT UP arc heuristic needs {checks} consistency-checks'
assignment = ACSolver(cryptarithmetic).domain_splitting()
from IPython.display import Latex
display(Latex(r'\begin{array}{@{}r@{}} ' + '{}{}{}{}'.format(assignment['S'], assignment['E'], assignment['N'], assignment['D']) + r' \\ + ' +
'{}{}{}{}'.format(assignment['M'], assignment['O'], assignment['R'], assignment['E']) + r' \\ \hline ' +
'{}{}{}{}{}'.format(assignment['M'], assignment['O'], assignment['N'], assignment['E'], assignment['Y']) + ' \end{array}'))
```
## References
<a name="cite-van2002domain"/><sup>[[1]](#ref-1) </sup>Van Dongen, Marc RC. 2002. _Domain-heuristics for arc-consistency algorithms_.
<a name="cite-van2000improving"/><sup>[[2]](#ref-2) </sup>Van Dongen, MRC and Bowen, JA. 2000. _Improving arc-consistency algorithms with double-support checks_.
<a name="cite-wallace1992ordering"/><sup>[[3]](#ref-3) </sup>Wallace, Richard J and Freuder, Eugene Charles. 1992. _Ordering heuristics for arc consistency algorithms_.
| github_jupyter |
```
# default_exp callback.core
#export
from fastai.data.all import *
from fastai.optimizer import *
#hide
from nbdev.showdoc import *
#export
_all_ = ['CancelFitException', 'CancelEpochException', 'CancelTrainException', 'CancelValidException', 'CancelBatchException']
```
# Callback
> Basic callbacks for Learner
## Events
Callbacks can occur at any of these times:: *before_fit before_epoch before_train before_batch after_pred after_loss before_backward after_backward after_step after_cancel_batch after_batch after_cancel_train after_train before_validate after_cancel_validate after_validate after_cancel_epoch after_epoch after_cancel_fit after_fit*.
```
# export
_events = L.split('before_fit before_epoch before_train before_batch after_pred after_loss \
before_backward after_backward after_step after_cancel_batch after_batch after_cancel_train \
after_train before_validate after_cancel_validate after_validate after_cancel_epoch \
after_epoch after_cancel_fit after_fit')
mk_class('event', **_events.map_dict(),
doc="All possible events as attributes to get tab-completion and typo-proofing")
# export
_all_ = ['event']
show_doc(event, name='event', title_level=3)
```
To ensure that you are referring to an event (that is, the name of one of the times when callbacks are called) that exists, and to get tab completion of event names, use `event`:
```
test_eq(event.after_backward, 'after_backward')
```
## Callback -
```
#export
_inner_loop = "before_batch after_pred after_loss before_backward after_backward after_step after_cancel_batch after_batch".split()
#export
@funcs_kwargs(as_method=True)
class Callback(GetAttr):
"Basic class handling tweaks of the training loop by changing a `Learner` in various events"
_default,learn,run,run_train,run_valid = 'learn',None,True,True,True
_methods = _events
def __init__(self, **kwargs): assert not kwargs, f'Passed unknown events: {kwargs}'
def __repr__(self): return type(self).__name__
def __call__(self, event_name):
"Call `self.{event_name}` if it's defined"
_run = (event_name not in _inner_loop or (self.run_train and getattr(self, 'training', True)) or
(self.run_valid and not getattr(self, 'training', False)))
res = None
if self.run and _run: res = getattr(self, event_name, noop)()
if event_name=='after_fit': self.run=True #Reset self.run to True at each end of fit
return res
def __setattr__(self, name, value):
if hasattr(self.learn,name):
warn(f"You are setting an attribute ({name}) that also exists in the learner, so you're not setting it in the learner but in the callback. Use `self.learn.{name}` otherwise.")
super().__setattr__(name, value)
@property
def name(self):
"Name of the `Callback`, camel-cased and with '*Callback*' removed"
return class2attr(self, 'Callback')
```
The training loop is defined in `Learner` a bit below and consists in a minimal set of instructions: looping through the data we:
- compute the output of the model from the input
- calculate a loss between this output and the desired target
- compute the gradients of this loss with respect to all the model parameters
- update the parameters accordingly
- zero all the gradients
Any tweak of this training loop is defined in a `Callback` to avoid over-complicating the code of the training loop, and to make it easy to mix and match different techniques (since they'll be defined in different callbacks). A callback can implement actions on the following events:
- `before_fit`: called before doing anything, ideal for initial setup.
- `before_epoch`: called at the beginning of each epoch, useful for any behavior you need to reset at each epoch.
- `before_train`: called at the beginning of the training part of an epoch.
- `before_batch`: called at the beginning of each batch, just after drawing said batch. It can be used to do any setup necessary for the batch (like hyper-parameter scheduling) or to change the input/target before it goes in the model (change of the input with techniques like mixup for instance).
- `after_pred`: called after computing the output of the model on the batch. It can be used to change that output before it's fed to the loss.
- `after_loss`: called after the loss has been computed, but before the backward pass. It can be used to add any penalty to the loss (AR or TAR in RNN training for instance).
- `before_backward`: called after the loss has been computed, but only in training mode (i.e. when the backward pass will be used)
- `after_backward`: called after the backward pass, but before the update of the parameters. It can be used to do any change to the gradients before said update (gradient clipping for instance).
- `after_step`: called after the step and before the gradients are zeroed.
- `after_batch`: called at the end of a batch, for any clean-up before the next one.
- `after_train`: called at the end of the training phase of an epoch.
- `before_validate`: called at the beginning of the validation phase of an epoch, useful for any setup needed specifically for validation.
- `after_validate`: called at the end of the validation part of an epoch.
- `after_epoch`: called at the end of an epoch, for any clean-up before the next one.
- `after_fit`: called at the end of training, for final clean-up.
```
show_doc(Callback.__call__)
```
One way to define callbacks is through subclassing:
```
class _T(Callback):
def call_me(self): return "maybe"
test_eq(_T()("call_me"), "maybe")
```
Another way is by passing the callback function to the constructor:
```
def cb(self): return "maybe"
_t = Callback(before_fit=cb)
test_eq(_t(event.before_fit), "maybe")
show_doc(Callback.__getattr__)
```
This is a shortcut to avoid having to write `self.learn.bla` for any `bla` attribute we seek, and just write `self.bla`.
```
mk_class('TstLearner', 'a')
class TstCallback(Callback):
def batch_begin(self): print(self.a)
learn,cb = TstLearner(1),TstCallback()
cb.learn = learn
test_stdout(lambda: cb('batch_begin'), "1")
```
Note that it only works to get the value of the attribute, if you want to change it, you have to manually access it with `self.learn.bla`. In the example below, `self.a += 1` creates an `a` attribute of 2 in the callback instead of setting the `a` of the learner to 2. It also issues a warning that something is probably wrong:
```
learn.a
class TstCallback(Callback):
def batch_begin(self): self.a += 1
learn,cb = TstLearner(1),TstCallback()
cb.learn = learn
cb('batch_begin')
test_eq(cb.a, 2)
test_eq(cb.learn.a, 1)
```
A proper version needs to write `self.learn.a = self.a + 1`:
```
class TstCallback(Callback):
def batch_begin(self): self.learn.a = self.a + 1
learn,cb = TstLearner(1),TstCallback()
cb.learn = learn
cb('batch_begin')
test_eq(cb.learn.a, 2)
show_doc(Callback.name, name='Callback.name')
test_eq(TstCallback().name, 'tst')
class ComplicatedNameCallback(Callback): pass
test_eq(ComplicatedNameCallback().name, 'complicated_name')
```
### TrainEvalCallback -
```
#export
class TrainEvalCallback(Callback):
"`Callback` that tracks the number of iterations done and properly sets training/eval mode"
run_valid = False
def before_fit(self):
"Set the iter and epoch counters to 0, put the model and the right device"
self.learn.train_iter,self.learn.pct_train = 0,0.
if hasattr(self.dls, 'device'): self.model.to(self.dls.device)
if hasattr(self.model, 'reset'): self.model.reset()
def after_batch(self):
"Update the iter counter (in training mode)"
self.learn.pct_train += 1./(self.n_iter*self.n_epoch)
self.learn.train_iter += 1
def before_train(self):
"Set the model in training mode"
self.learn.pct_train=self.epoch/self.n_epoch
self.model.train()
self.learn.training=True
def before_validate(self):
"Set the model in validation mode"
self.model.eval()
self.learn.training=False
show_doc(TrainEvalCallback, title_level=3)
```
This `Callback` is automatically added in every `Learner` at initialization.
```
#hide
#test of the TrainEvalCallback below in Learner.fit
show_doc(TrainEvalCallback.before_fit)
show_doc(TrainEvalCallback.after_batch)
show_doc(TrainEvalCallback.before_train)
# export
if not hasattr(defaults, 'callbacks'): defaults.callbacks = [TrainEvalCallback]
```
### GatherPredsCallback -
```
#export
#TODO: save_targs and save_preds only handle preds/targets that have one tensor, not tuples of tensors.
class GatherPredsCallback(Callback):
"`Callback` that saves the predictions and targets, optionally `with_loss`"
def __init__(self, with_input=False, with_loss=False, save_preds=None, save_targs=None, concat_dim=0):
store_attr(self, "with_input,with_loss,save_preds,save_targs,concat_dim")
def before_batch(self):
if self.with_input: self.inputs.append((to_detach(self.xb)))
def before_validate(self):
"Initialize containers"
self.preds,self.targets = [],[]
if self.with_input: self.inputs = []
if self.with_loss: self.losses = []
def after_batch(self):
"Save predictions, targets and potentially losses"
if not hasattr(self, 'pred'): return
preds,targs = to_detach(self.pred),to_detach(self.yb)
if self.save_preds is None: self.preds.append(preds)
else: (self.save_preds/str(self.iter)).save_array(preds)
if self.save_targs is None: self.targets.append(targs)
else: (self.save_targs/str(self.iter)).save_array(targs[0])
if self.with_loss:
bs = find_bs(self.yb)
loss = self.loss if self.loss.numel() == bs else self.loss.view(bs,-1).mean(1)
self.losses.append(to_detach(loss))
def after_validate(self):
"Concatenate all recorded tensors"
if not hasattr(self, 'preds'): return
if self.with_input: self.inputs = detuplify(to_concat(self.inputs, dim=self.concat_dim))
if not self.save_preds: self.preds = detuplify(to_concat(self.preds, dim=self.concat_dim))
if not self.save_targs: self.targets = detuplify(to_concat(self.targets, dim=self.concat_dim))
if self.with_loss: self.losses = to_concat(self.losses)
def all_tensors(self):
res = [None if self.save_preds else self.preds, None if self.save_targs else self.targets]
if self.with_input: res = [self.inputs] + res
if self.with_loss: res.append(self.losses)
return res
show_doc(GatherPredsCallback, title_level=3)
show_doc(GatherPredsCallback.before_validate)
show_doc(GatherPredsCallback.after_batch)
show_doc(GatherPredsCallback.after_validate)
#export
class FetchPredsCallback(Callback):
"A callback to fetch predictions during the training loop"
remove_on_fetch = True
def __init__(self, ds_idx=1, dl=None, with_input=False, with_decoded=False, cbs=None, reorder=True):
self.cbs = L(cbs)
store_attr(self, 'ds_idx,dl,with_input,with_decoded,reorder')
def after_validate(self):
to_rm = L(cb for cb in self.learn.cbs if getattr(cb, 'remove_on_fetch', False))
with self.learn.removed_cbs(to_rm + self.cbs) as learn:
self.preds = learn.get_preds(ds_idx=self.ds_idx, dl=self.dl,
with_input=self.with_input, with_decoded=self.with_decoded, inner=True, reorder=self.reorder)
show_doc(FetchPredsCallback, title_level=3)
```
When writing a callback, the following attributes of `Learner` are available:
- `model`: the model used for training/validation
- `data`: the underlying `DataLoaders`
- `loss_func`: the loss function used
- `opt`: the optimizer used to update the model parameters
- `opt_func`: the function used to create the optimizer
- `cbs`: the list containing all `Callback`s
- `dl`: current `DataLoader` used for iteration
- `x`/`xb`: last input drawn from `self.dl` (potentially modified by callbacks). `xb` is always a tuple (potentially with one element) and `x` is detuplified. You can only assign to `xb`.
- `y`/`yb`: last target drawn from `self.dl` (potentially modified by callbacks). `yb` is always a tuple (potentially with one element) and `y` is detuplified. You can only assign to `yb`.
- `pred`: last predictions from `self.model` (potentially modified by callbacks)
- `loss`: last computed loss (potentially modified by callbacks)
- `n_epoch`: the number of epochs in this training
- `n_iter`: the number of iterations in the current `self.dl`
- `epoch`: the current epoch index (from 0 to `n_epoch-1`)
- `iter`: the current iteration index in `self.dl` (from 0 to `n_iter-1`)
The following attributes are added by `TrainEvalCallback` and should be available unless you went out of your way to remove that callback:
- `train_iter`: the number of training iterations done since the beginning of this training
- `pct_train`: from 0. to 1., the percentage of training iterations completed
- `training`: flag to indicate if we're in training mode or not
The following attribute is added by `Recorder` and should be available unless you went out of your way to remove that callback:
- `smooth_loss`: an exponentially-averaged version of the training loss
## Callbacks control flow
It happens that we may want to skip some of the steps of the training loop: in gradient accumulation, we don't always want to do the step/zeroing of the grads for instance. During an LR finder test, we don't want to do the validation phase of an epoch. Or if we're training with a strategy of early stopping, we want to be able to completely interrupt the training loop.
This is made possible by raising specific exceptions the training loop will look for (and properly catch).
```
#export
_ex_docs = dict(
CancelFitException="Skip the rest of this batch and go to `after_batch`",
CancelEpochException="Skip the rest of the training part of the epoch and go to `after_train`",
CancelTrainException="Skip the rest of the validation part of the epoch and go to `after_validate`",
CancelValidException="Skip the rest of this epoch and go to `after_epoch`",
CancelBatchException="Interrupts training and go to `after_fit`")
for c,d in _ex_docs.items(): mk_class(c,sup=Exception,doc=d)
show_doc(CancelBatchException, title_level=3)
show_doc(CancelTrainException, title_level=3)
show_doc(CancelValidException, title_level=3)
show_doc(CancelEpochException, title_level=3)
show_doc(CancelFitException, title_level=3)
```
You can detect one of those exceptions occurred and add code that executes right after with the following events:
- `after_cancel_batch`: reached immediately after a `CancelBatchException` before proceeding to `after_batch`
- `after_cancel_train`: reached immediately after a `CancelTrainException` before proceeding to `after_epoch`
- `after_cancel_valid`: reached immediately after a `CancelValidException` before proceeding to `after_epoch`
- `after_cancel_epoch`: reached immediately after a `CancelEpochException` before proceeding to `after_epoch`
- `after_cancel_fit`: reached immediately after a `CancelFitException` before proceeding to `after_fit`
## Export -
```
#hide
from nbdev.export import notebook2script
notebook2script()
```
| github_jupyter |
<a href="https://www.bigdatauniversity.com"><img src = "https://ibm.box.com/shared/static/jvcqp2iy2jlx2b32rmzdt0tx8lvxgzkp.png" width = 300, align = "center"></a>
<h1 align=center><font size = 5>CONVOLUTIONAL NEURAL NETWORK APPLICATION</font></h1>
## Introduction
In this section, we will use the famous [MNIST Dataset](http://yann.lecun.com/exdb/mnist/) to build two Neural Networks capable to perform handwritten digits classification. The first Network is a simple Multi-layer Perceptron (MLP) and the second one is a Convolutional Neural Network (CNN from now on). In other words, our algorithm will say, with some associated error, what type of digit is the presented input.
---
## Table of contents
<div class="alert alert-block alert-info" style="margin-top: 20px">
<font size = 3><strong>Clik on the links to see the sections:</strong></font>
<br>
- <p><a href="#ref1">What is Deep Learning</a></p>
- <p><a href="#ref2">Simple test: Is tensorflow working?</a></p>
- <p><a href="#ref3">1st part: classify MNIST using a simple model</a></p>
- <p><a href="#ref4">Evaluating the final result</a></p>
- <p><a href="#ref5">How to improve our model?</a></p>
- <p><a href="#ref6">2nd part: Deep Learning applied on MNIST</a></p>
- <p><a href="#ref7">Summary of the Deep Convolutional Neural Network</a></p>
- <p><a href="#ref8">Define functions and train the model</a></p>
- <p><a href="#ref9">Evaluate the model</a></p>
---
<a id="ref1"></a>
# What is Deep Learning?
**Brief Theory:** Deep learning (also known as deep structured learning, hierarchical learning or deep machine learning) is a branch of machine learning based on a set of algorithms that attempt to model high-level abstractions in data by using multiple processing layers, with complex structures or otherwise, composed of multiple non-linear transformations.
<img src="https://ibm.box.com/shared/static/gcbbrh440604cj2nksu3f44be87b8ank.png" alt="HTML5 Icon" style="width:600px;height:450px;">
<div style="text-align:center">It's time for deep learning. Our brain does't work with one or three layers. Why it would be different with machines?. </div>
**In Practice, defining the term "Deep":** in this context, deep means that we are studying a Neural Network which has several hidden layers (more than one), no matter what type (convolutional, pooling, normalization, fully-connected etc). The most interesting part is that some papers noticed that Deep Neural Networks with right architectures/hyper-parameters achieve better results than shallow Neural Networks with same computational power (e.g. number of neurons or connections).
**In Practice, defining "Learning":** In the context of supervised learning, digits recognition in our case, the learning consists of a target/feature which is to be predicted using a given set of observations with the already known final prediction (label). In our case, the target will be the digit (0,1,2,3,4,5,6,7,8,9) and the observations are the intensity and relative position of pixels. After some training, it's possible to generate a "function" that map inputs (digit image) to desired outputs(type of digit). The only problem is how well this map operation occurs. While trying to generate this "function", the training process continues until the model achieves a desired level of accuracy on the training data.
---
__Notice:__ This notebook has been created with tensorflow version 1.8, and might not work with other versions.
```
import tensorflow as tf
tf.__version__
```
In this tutorial, we first classify MNIST using a simple Multi-layer percepetron and then, in the second part, we use deeplearning to improve the accuracy of our results.
<a id="ref3"></a>
# 1st part: classify MNIST using a simple model.
We are going to create a simple Multi-layer percepetron, a simple type of Neural Network, to performe classification tasks on the MNIST digits dataset. If you are not familiar with the MNIST dataset, please consider to read more about it: <a href="http://yann.lecun.com/exdb/mnist/">click here</a>
### What is MNIST?
According to Lecun's website, the MNIST is a: "database of handwritten digits that has a training set of 60,000 examples, and a test set of 10,000 examples. It is a subset of a larger set available from NIST. The digits have been size-normalized and centered in a fixed-size image".
### Import the MNIST dataset using TensorFlow built-in feature
It's very important to notice that MNIST is a high optimized data-set and it does not contain images. You will need to build your own code if you want to see the real digits. Another important side note is the effort that the authors invested on this data-set with normalization and centering operations.
```
from tensorflow.examples.tutorials.mnist import input_data
mnist = input_data.read_data_sets("MNIST_data/", one_hot=True)
```
The <span style="background-color:#dcdcdc
"> One-hot = True</span> argument only means that, in contrast to Binary representation, the labels will be presented in a way that only one bit will be on for a specific digit. For example, five and zero in a binary code would be:
<pre>
Number representation: 0
Binary encoding: [2^5] [2^4] [2^3] [2^2] [2^1] [2^0]
Array/vector: 0 0 0 0 0 0
Number representation: 5
Binary encoding: [2^5] [2^4] [2^3] [2^2] [2^1] [2^0]
Array/vector: 0 0 0 1 0 1
</pre>
Using a different notation, the same digits using one-hot vector representation can be show as:
<pre>
Number representation: 0
One-hot encoding: [5] [4] [3] [2] [1] [0]
Array/vector: 0 0 0 0 0 1
Number representation: 5
One-hot encoding: [5] [4] [3] [2] [1] [0]
Array/vector: 1 0 0 0 0 0
</pre>
### Understanding the imported data
The imported data can be divided as follow:
- Training (mnist.train) >> Use the given dataset with inputs and related outputs for training of NN. In our case, if you give an image that you know that represents a "nine", this set will tell the neural network that we expect a "nine" as the output.
- 55,000 data points
- mnist.train.images for inputs
- mnist.train.labels for outputs
- Validation (mnist.validation) >> The same as training, but now the date is used to generate model properties (classification error, for example) and from this, tune parameters like the optimal number of hidden units or determine a stopping point for the back-propagation algorithm
- 5,000 data points
- mnist.validation.images for inputs
- mnist.validation.labels for outputs
- Test (mnist.test) >> the model does not have access to this informations prior to the test phase. It is used to evaluate the performance and accuracy of the model against "real life situations". No further optimization beyond this point.
- 10,000 data points
- mnist.test.images for inputs
- mnist.test.labels for outputs
### Creating an interactive section
You have two basic options when using TensorFlow to run your code:
- [Build graphs and run session] Do all the set-up and THEN execute a session to evaluate tensors and run operations (ops)
- [Interactive session] create your coding and run on the fly.
For this first part, we will use the interactive session that is more suitable for environments like Jupyter notebooks.
```
sess = tf.InteractiveSession()
```
### Creating placeholders
It's a best practice to create placeholders before variable assignments when using TensorFlow. Here we'll create placeholders for inputs ("Xs") and outputs ("Ys").
__Placeholder 'X':__ represents the "space" allocated input or the images.
* Each input has 784 pixels distributed by a 28 width x 28 height matrix
* The 'shape' argument defines the tensor size by its dimensions.
* 1st dimension = None. Indicates that the batch size, can be of any size.
* 2nd dimension = 784. Indicates the number of pixels on a single flattened MNIST image.
__Placeholder 'Y':___ represents the final output or the labels.
* 10 possible classes (0,1,2,3,4,5,6,7,8,9)
* The 'shape' argument defines the tensor size by its dimensions.
* 1st dimension = None. Indicates that the batch size, can be of any size.
* 2nd dimension = 10. Indicates the number of targets/outcomes
__dtype for both placeholders:__ if you not sure, use tf.float32. The limitation here is that the later presented softmax function only accepts float32 or float64 dtypes. For more dtypes, check TensorFlow's documentation <a href="https://www.tensorflow.org/versions/r0.9/api_docs/python/framework.html#tensor-types">here</a>
```
x = tf.placeholder(tf.float32, shape=[None, 784])
y_ = tf.placeholder(tf.float32, shape=[None, 10])
```
### Assigning bias and weights to null tensors
Now we are going to create the weights and biases, for this purpose they will be used as arrays filled with zeros. The values that we choose here can be critical, but we'll cover a better way on the second part, instead of this type of initialization.
```
# Weight tensor
W = tf.Variable(tf.zeros([784,10],tf.float32))
# Bias tensor
b = tf.Variable(tf.zeros([10],tf.float32))
```
### Execute the assignment operation
Before, we assigned the weights and biases but we did not initialize them with null values. For this reason, TensorFlow need to initialize the variables that you assign.
Please notice that we're using this notation "sess.run" because we previously started an interactive session.
```
# run the op initialize_all_variables using an interactive session
sess.run(tf.global_variables_initializer())
```
### Adding Weights and Biases to input
The only difference from our next operation to the picture below is that we are using the mathematical convention for what is being executed in the illustration. The tf.matmul operation performs a matrix multiplication between x (inputs) and W (weights) and after the code add biases.
<img src="https://ibm.box.com/shared/static/88ksiymk1xkb10rgk0jwr3jw814jbfxo.png" alt="HTML5 Icon" style="width:400px;height:350px;">
<div style="text-align:center">Illustration showing how weights and biases are added to neurons/nodes. </div>
```
#mathematical operation to add weights and biases to the inputs
tf.matmul(x,W) + b
```
### Softmax Regression
Softmax is an activation function that is normally used in classification problems. It generate the probabilities for the output. For example, our model will not be 100% sure that one digit is the number nine, instead, the answer will be a distribution of probabilities where, if the model is right, the nine number will have the larger probability.
For comparison, below is the one-hot vector for a nine digit label:
A machine does not have all this certainty, so we want to know what is the best guess, but we also want to understand how sure it was and what was the second better option. Below is an example of a hypothetical distribution for a nine digit:
```
y = tf.nn.softmax(tf.matmul(x,W) + b)
```
Logistic function output is used for the classification between two target classes 0/1. Softmax function is generalized type of logistic function. That is, Softmax can output a multiclass categorical probability distribution.
### Cost function
It is a function that is used to minimize the difference between the right answers (labels) and estimated outputs by our Network.
```
cross_entropy = tf.reduce_mean(-tf.reduce_sum(y_ * tf.log(y), reduction_indices=[1]))
```
### Type of optimization: Gradient Descent
This is the part where you configure the optimizer for you Neural Network. There are several optimizers available, in our case we will use Gradient Descent that is very well stablished.
```
train_step = tf.train.GradientDescentOptimizer(0.5).minimize(cross_entropy)
```
### Training batches
Train using minibatch Gradient Descent.
In practice, Batch Gradient Descent is not often used because is too computationally expensive. The good part about this method is that you have the true gradient, but with the expensive computing task of using the whole dataset in one time. Due to this problem, Neural Networks usually use minibatch to train.
```
#Load 50 training examples for each training iteration
for i in range(1000):
batch = mnist.train.next_batch(50)
train_step.run(feed_dict={x: batch[0], y_: batch[1]})
```
### Test
```
correct_prediction = tf.equal(tf.argmax(y,1), tf.argmax(y_,1))
accuracy = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
acc = accuracy.eval(feed_dict={x: mnist.test.images, y_: mnist.test.labels}) * 100
print("The final accuracy for the simple ANN model is: {} % ".format(acc) )
sess.close() #finish the session
```
---
<a id="ref4"></a>
# Evaluating the final result
Is the final result good?
Let's check the best algorithm available out there (10th june 2016):
_Result:_ 0.21% error (99.79% accuracy)
<a href="http://cs.nyu.edu/~wanli/dropc/">Reference here</a>
<a id="ref5"></a>
# How to improve our model?
#### Several options as follow:
- Regularization of Neural Networks using DropConnect
- Multi-column Deep Neural Networks for Image Classification
- APAC: Augmented Pattern Classification with Neural Networks
- Simple Deep Neural Network with Dropout
#### In the next part we are going to explore the option:
- Simple Deep Neural Network with Dropout (more than 1 hidden layer)
---
<a id="ref6"></a>
# 2nd part: Deep Learning applied on MNIST
In the first part, we learned how to use a simple ANN to classify MNIST. Now we are going to expand our knowledge using a Deep Neural Network.
Architecture of our network is:
- (Input) -> [batch_size, 28, 28, 1] >> Apply 32 filter of [5x5]
- (Convolutional layer 1) -> [batch_size, 28, 28, 32]
- (ReLU 1) -> [?, 28, 28, 32]
- (Max pooling 1) -> [?, 14, 14, 32]
- (Convolutional layer 2) -> [?, 14, 14, 64]
- (ReLU 2) -> [?, 14, 14, 64]
- (Max pooling 2) -> [?, 7, 7, 64]
- [fully connected layer 3] -> [1x1024]
- [ReLU 3] -> [1x1024]
- [Drop out] -> [1x1024]
- [fully connected layer 4] -> [1x10]
The next cells will explore this new architecture.
### Starting the code
```
import tensorflow as tf
# finish possible remaining session
sess.close()
#Start interactive session
sess = tf.InteractiveSession()
```
### The MNIST data
```
from tensorflow.examples.tutorials.mnist import input_data
mnist = input_data.read_data_sets('MNIST_data', one_hot=True)
```
### Initial parameters
Create general parameters for the model
```
width = 28 # width of the image in pixels
height = 28 # height of the image in pixels
flat = width * height # number of pixels in one image
class_output = 10 # number of possible classifications for the problem
```
### Input and output
Create place holders for inputs and outputs
```
x = tf.placeholder(tf.float32, shape=[None, flat])
y_ = tf.placeholder(tf.float32, shape=[None, class_output])
```
#### Converting images of the data set to tensors
The input image is a 28 pixels by 28 pixels, 1 channel (grayscale). In this case, the first dimension is the __batch number__ of the image, and can be of any size (so we set it to -1). The second and third dimensions are width and hight, and the last one is the image channels.
```
x_image = tf.reshape(x, [-1,28,28,1])
x_image
```
### Convolutional Layer 1
#### Defining kernel weight and bias
We define a kernle here. The Size of the filter/kernel is 5x5; Input channels is 1 (greyscale); and we need 32 different feature maps (here, 32 feature maps means 32 different filters are applied on each image. So, the output of convolution layer would be 28x28x32). In this step, we create a filter / kernel tensor of shape `[filter_height, filter_width, in_channels, out_channels]`
```
W_conv1 = tf.Variable(tf.truncated_normal([5, 5, 1, 32], stddev=0.1))
b_conv1 = tf.Variable(tf.constant(0.1, shape=[32])) # need 32 biases for 32 outputs
```
<img src="https://ibm.box.com/shared/static/vn26neef1nnv2oxn5cb3uueowcawhkgb.png" style="width:800px;height:400px;" alt="HTML5 Icon" >
#### Convolve with weight tensor and add biases.
To creat convolutional layer, we use __tf.nn.conv2d__. It computes a 2-D convolution given 4-D input and filter tensors.
Inputs:
- tensor of shape [batch, in_height, in_width, in_channels]. x of shape [batch_size,28 ,28, 1]
- a filter / kernel tensor of shape [filter_height, filter_width, in_channels, out_channels]. W is of size [5, 5, 1, 32]
- stride which is [1, 1, 1, 1]. The convolutional layer, slides the "kernel window" across the input tensor. As the input tensor has 4 dimensions: [batch, height, width, channels], then the convolution operates on a 2D window on the height and width dimensions. __strides__ determines how much the window shifts by in each of the dimensions. As the first and last dimensions are related to batch and channels, we set the stride to 1. But for second and third dimension, we coould set other values, e.g. [1, 2, 2, 1]
Process:
- Change the filter to a 2-D matrix with shape [5\*5\*1,32]
- Extracts image patches from the input tensor to form a *virtual* tensor of shape `[batch, 28, 28, 5*5*1]`.
- For each batch, right-multiplies the filter matrix and the image vector.
Output:
- A `Tensor` (a 2-D convolution) of size <tf.Tensor 'add_7:0' shape=(?, 28, 28, 32)- Notice: the output of the first convolution layer is 32 [28x28] images. Here 32 is considered as volume/depth of the output image.
```
convolve1= tf.nn.conv2d(x_image, W_conv1, strides=[1, 1, 1, 1], padding='SAME') + b_conv1
```
<img src="https://ibm.box.com/shared/static/iizf4ui4b2hh9wn86pplqxu27ykpqci9.png" style="width:800px;height:400px;" alt="HTML5 Icon" >
#### Apply the ReLU activation Function
In this step, we just go through all outputs convolution layer, __covolve1__, and wherever a negative number occurs,we swap it out for a 0. It is called ReLU activation Function.
```
h_conv1 = tf.nn.relu(convolve1)
```
#### Apply the max pooling
__max pooling__ is a form of non-linear down-sampling. It partitions the input image into a set of rectangles and, and then find the maximum value for that region.
Lets use __tf.nn.max_pool__ function to perform max pooling.
__Kernel size:__ 2x2 (if the window is a 2x2 matrix, it would result in one output pixel)
__Strides:__ dictates the sliding behaviour of the kernel. In this case it will move 2 pixels everytime, thus not overlapping. The input is a matix of size 14x14x32, and the output would be a matrix of size 14x14x32.
<img src="https://ibm.box.com/shared/static/kmaja90mn3aud9mro9cn8pbbg1h5pejy.png" alt="HTML5 Icon" style="width:800px;height:400px;">
```
conv1 = tf.nn.max_pool(h_conv1, ksize=[1, 2, 2, 1], strides=[1, 2, 2, 1], padding='SAME') #max_pool_2x2
conv1
```
First layer completed
### Convolutional Layer 2
#### Weights and Biases of kernels
We apply the convolution again in this layer. Lets look at the second layer kernel:
- Filter/kernel: 5x5 (25 pixels)
- Input channels: 32 (from the 1st Conv layer, we had 32 feature maps)
- 64 output feature maps
__Notice:__ here, the input image is [14x14x32], the filter is [5x5x32], we use 64 filters of size [5x5x32], and the output of the convolutional layer would be 64 covolved image, [14x14x64].
__Notice:__ the convolution result of applying a filter of size [5x5x32] on image of size [14x14x32] is an image of size [14x14x1], that is, the convolution is functioning on volume.
```
W_conv2 = tf.Variable(tf.truncated_normal([5, 5, 32, 64], stddev=0.1))
b_conv2 = tf.Variable(tf.constant(0.1, shape=[64])) #need 64 biases for 64 outputs
```
#### Convolve image with weight tensor and add biases.
```
convolve2= tf.nn.conv2d(conv1, W_conv2, strides=[1, 1, 1, 1], padding='SAME')+ b_conv2
```
#### Apply the ReLU activation Function
```
h_conv2 = tf.nn.relu(convolve2)
```
#### Apply the max pooling
```
conv2 = tf.nn.max_pool(h_conv2, ksize=[1, 2, 2, 1], strides=[1, 2, 2, 1], padding='SAME') #max_pool_2x2
conv2
```
Second layer completed. So, what is the output of the second layer, layer2?
- it is 64 matrix of [7x7]
### Fully Connected Layer
You need a fully connected layer to use the Softmax and create the probabilities in the end. Fully connected layers take the high-level filtered images from previous layer, that is all 64 matrics, and convert them to a flat array.
So, each matrix [7x7] will be converted to a matrix of [49x1], and then all of the 64 matrix will be connected, which make an array of size [3136x1]. We will connect it into another layer of size [1024x1]. So, the weight between these 2 layers will be [3136x1024]
<img src="https://ibm.box.com/shared/static/pr9mnirmlrzm2bitf1d4jj389hyvv7ey.png" alt="HTML5 Icon" style="width:800px;height:400px;">
#### Flattening Second Layer
```
layer2_matrix = tf.reshape(conv2, [-1, 7*7*64])
```
#### Weights and Biases between layer 2 and 3
Composition of the feature map from the last layer (7x7) multiplied by the number of feature maps (64); 1027 outputs to Softmax layer
```
W_fc1 = tf.Variable(tf.truncated_normal([7 * 7 * 64, 1024], stddev=0.1))
b_fc1 = tf.Variable(tf.constant(0.1, shape=[1024])) # need 1024 biases for 1024 outputs
```
#### Matrix Multiplication (applying weights and biases)
```
fcl=tf.matmul(layer2_matrix, W_fc1) + b_fc1
```
#### Apply the ReLU activation Function
```
h_fc1 = tf.nn.relu(fcl)
h_fc1
```
Third layer completed
#### Dropout Layer, Optional phase for reducing overfitting
It is a phase where the network "forget" some features. At each training step in a mini-batch, some units get switched off randomly so that it will not interact with the network. That is, it weights cannot be updated, nor affect the learning of the other network nodes. This can be very useful for very large neural networks to prevent overfitting.
```
keep_prob = tf.placeholder(tf.float32)
layer_drop = tf.nn.dropout(h_fc1, keep_prob)
layer_drop
```
### Readout Layer (Softmax Layer)
Type: Softmax, Fully Connected Layer.
#### Weights and Biases
In last layer, CNN takes the high-level filtered images and translate them into votes using softmax.
Input channels: 1024 (neurons from the 3rd Layer); 10 output features
```
W_fc2 = tf.Variable(tf.truncated_normal([1024, 10], stddev=0.1)) #1024 neurons
b_fc2 = tf.Variable(tf.constant(0.1, shape=[10])) # 10 possibilities for digits [0,1,2,3,4,5,6,7,8,9]
```
#### Matrix Multiplication (applying weights and biases)
```
fc=tf.matmul(layer_drop, W_fc2) + b_fc2
```
#### Apply the Softmax activation Function
__softmax__ allows us to interpret the outputs of __fcl4__ as probabilities. So, __y_conv__ is a tensor of probablities.
```
y_CNN= tf.nn.softmax(fc)
y_CNN
```
---
<a id="ref7"></a>
# Summary of the Deep Convolutional Neural Network
Now is time to remember the structure of our network
#### 0) Input - MNIST dataset
#### 1) Convolutional and Max-Pooling
#### 2) Convolutional and Max-Pooling
#### 3) Fully Connected Layer
#### 4) Processing - Dropout
#### 5) Readout layer - Fully Connected
#### 6) Outputs - Classified digits
---
<a id="ref8"></a>
# Define functions and train the model
#### Define the loss function
We need to compare our output, layer4 tensor, with ground truth for all mini_batch. we can use __cross entropy__ to see how bad our CNN is working - to measure the error at a softmax layer.
The following code shows an toy sample of cross-entropy for a mini-batch of size 2 which its items have been classified. You can run it (first change the cell type to __code__ in the toolbar) to see hoe cross entropy changes.
__reduce_sum__ computes the sum of elements of __(y_ * tf.log(layer4)__ across second dimension of the tensor, and __reduce_mean__ computes the mean of all elements in the tensor..
```
cross_entropy = tf.reduce_mean(-tf.reduce_sum(y_ * tf.log(y_CNN), reduction_indices=[1]))
```
#### Define the optimizer
It is obvious that we want minimize the error of our network which is calculated by cross_entropy metric. To solve the problem, we have to compute gradients for the loss (which is minimizing the cross-entropy) and apply gradients to variables. It will be done by an optimizer: GradientDescent or Adagrad.
```
train_step = tf.train.AdamOptimizer(1e-4).minimize(cross_entropy)
```
#### Define prediction
Do you want to know how many of the cases in a mini-batch has been classified correctly? lets count them.
```
correct_prediction = tf.equal(tf.argmax(y_CNN,1), tf.argmax(y_,1))
```
#### Define accuracy
It makes more sense to report accuracy using average of correct cases.
```
accuracy = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
```
#### Run session, train
```
sess.run(tf.global_variables_initializer())
```
*If you want a fast result (**it might take sometime to train it**)*
```
for i in range(1100):
batch = mnist.train.next_batch(50)
if i%100 == 0:
train_accuracy = accuracy.eval(feed_dict={x:batch[0], y_: batch[1], keep_prob: 1.0})
print("step %d, training accuracy %g"%(i, float(train_accuracy)))
train_step.run(feed_dict={x: batch[0], y_: batch[1], keep_prob: 0.5})
```
<div class="alert alert-success alertsuccess" style="margin-top: 20px">
<font size = 3><strong>*You can run this cell if you REALLY have time to wait, or you are running it using PowerAI (**change the type of the cell to code**)*</strong></font>
_PS. If you have problems running this notebook, please shutdown all your Jupyter runnning notebooks, clear all cells outputs and run each cell only after the completion of the previous cell._
---
<a id="ref9"></a>
# Evaluate the model
Print the evaluation to the user
```
print("test accuracy %g"%accuracy.eval(feed_dict={x: mnist.test.images, y_: mnist.test.labels, keep_prob: 1.0}))
```
## Visualization
Do you want to look at all the filters?
```
kernels = sess.run(tf.reshape(tf.transpose(W_conv1, perm=[2, 3, 0,1]),[32,-1]))
!wget --output-document utils1.py http://deeplearning.net/tutorial/code/utils.py
import utils1
from utils1 import tile_raster_images
import matplotlib.pyplot as plt
from PIL import Image
%matplotlib inline
image = Image.fromarray(tile_raster_images(kernels, img_shape=(5, 5) ,tile_shape=(4, 8), tile_spacing=(1, 1)))
### Plot image
plt.rcParams['figure.figsize'] = (18.0, 18.0)
imgplot = plt.imshow(image)
imgplot.set_cmap('gray')
```
Do you want to see the output of an image passing through first convolution layer?
```
import numpy as np
plt.rcParams['figure.figsize'] = (5.0, 5.0)
sampleimage = mnist.test.images[1]
plt.imshow(np.reshape(sampleimage,[28,28]), cmap="gray")
ActivatedUnits = sess.run(convolve1,feed_dict={x:np.reshape(sampleimage,[1,784],order='F'),keep_prob:1.0})
filters = ActivatedUnits.shape[3]
plt.figure(1, figsize=(20,20))
n_columns = 6
n_rows = np.math.ceil(filters / n_columns) + 1
for i in range(filters):
plt.subplot(n_rows, n_columns, i+1)
plt.title('Filter ' + str(i))
plt.imshow(ActivatedUnits[0,:,:,i], interpolation="nearest", cmap="gray")
```
What about second convolution layer?
```
ActivatedUnits = sess.run(convolve2,feed_dict={x:np.reshape(sampleimage,[1,784],order='F'),keep_prob:1.0})
filters = ActivatedUnits.shape[3]
plt.figure(1, figsize=(20,20))
n_columns = 8
n_rows = np.math.ceil(filters / n_columns) + 1
for i in range(filters):
plt.subplot(n_rows, n_columns, i+1)
plt.title('Filter ' + str(i))
plt.imshow(ActivatedUnits[0,:,:,i], interpolation="nearest", cmap="gray")
sess.close() #finish the session
```
## Want to learn more?
Running deep learning programs usually needs a high performance platform. PowerAI speeds up deep learning and AI. Built on IBM's Power Systems, PowerAI is a scalable software platform that accelerates deep learning and AI with blazing performance for individual users or enterprises. The PowerAI platform supports popular machine learning libraries and dependencies including Tensorflow, Caffe, Torch, and Theano. You can download a [free version of PowerAI](https://cocl.us/ML0120EN_PAI).
Also, you can use Data Science Experience to run these notebooks faster with bigger datasets. Data Science Experience is IBM's leading cloud solution for data scientists, built by data scientists. With Jupyter notebooks, RStudio, Apache Spark and popular libraries pre-packaged in the cloud, DSX enables data scientists to collaborate on their projects without having to install anything. Join the fast-growing community of DSX users today with a free account at [Data Science Experience](https://cocl.us/ML0120EN_DSX)This is the end of this lesson. Hopefully, now you have a deeper and intuitive understanding regarding the LSTM model. Thank you for reading this notebook, and good luck on your studies.
### Thanks for completing this lesson!
Created by <a href = "https://linkedin.com/in/saeedaghabozorgi"> Saeed Aghabozorgi </a>, <a href = "https://linkedin.com/in/luisotsm">Luis Otavio Silveira Martins</a>, <a href = "https://linkedin.com/in/erich-natsubori-sato"> Erich Natsubori Sato </a></h4>
### References:
https://en.wikipedia.org/wiki/Deep_learning
http://sebastianruder.com/optimizing-gradient-descent/index.html#batchgradientdescent
http://yann.lecun.com/exdb/mnist/
https://www.quora.com/Artificial-Neural-Networks-What-is-the-difference-between-activation-functions
https://www.tensorflow.org/versions/r0.9/tutorials/mnist/pros/index.html
| github_jupyter |
# Combining Data on Grades
```
import numpy as np
import pandas as pd
pd.set_option('display.max_columns', 10)
pd.set_option('display.max_rows', 10)
```
### Given: Three Numpy Arrays of Grades
```
rng = np.random.default_rng(seed=42)
ar1 = rng.choice(
['A', 'B', 'C', 'D', 'F'], 100, p=[.2, .4, .29, .08, .03]
)
ar2 = rng.choice(
['A', 'B', 'C', 'D', 'F'], 50, p=[.3, .4, .2, .1, 0]
)
ar3 = rng.choice(
['a', 'b', 'c', 'd', 'f'], 200, p=[.15, .45, .25, .12, .03]
)
```
### Create pandas Series from these Arrays
Use the default index for each. Save the series as `s1`, `s2` and `s3`
```
s1 = pd.Series(ar1)
s2 = pd.Series(ar2)
s3 = pd.Series(ar3)
```
### Get the Value Counts of each of the Series
Save the resulting series as `grades1`, `grades2`, and `grades3`
```
grades1 = s1.value_counts()
grades2 = s2.value_counts()
grades3 = s3.value_counts()
grades1, grades2, grades3
```
#### The following code creates the table shown in the manual. You'll learn about `DataFrame`s soon.
```
pd.DataFrame(
{
'grades1': grades1,
'grades2': grades2,
'grades3': grades3,
})
```
### Compare the indexes of the three `grades` variables
You should see that the index for `grades3` uses lowercase letters, while the other two use uppercase letters.
```
grades1.index, grades2.index, grades3.index
```
### Reindex `grades3` to use uppercase letters
This is a little tricky because the indices are not in alphabetical order. You will need to sort them first, and then set the index for `grades3` to use capital letters.
* We only really need to sort `grades1` and `grades2` if we're going to compare them, but we may as well.
```
grades1.sort_index(inplace=True)
grades2.sort_index(inplace=True)
grades3.sort_index(inplace=True)
grades3.index = ['A', 'B', 'C', 'D', 'F']
grades3.index
grades3
```
### Add the three `grades` Series together
Don't forget to set the fill value to 0.
```
grades_all = grades1.add(grades2, fill_value=0).add(grades3, fill_value=0)
grades_all
```
### From `grades_all`, create a `grades_breakout` Series that holds the share of each grade.
`grades_breakout.sum()` should equal 1
```
grades_breakout = grades_all / grades_all.sum()
grades_breakout, grades_breakout.sum()
```
## A different approach: First, change case of values of `s3`.
```
s3 = s3.str.upper()
s3
```
### Then combine the Series of grades.
```
s_all = pd.concat([s1, s2, s3])
s_all
```
### Then get the value counts of the combined series.
```
s_all.value_counts()
grades_breakout = s_all.value_counts(normalize=True)
grades_breakout.sort_index(inplace=True)
grades_breakout
```
| github_jupyter |
# Оптическое распознавание символов

Общей проблемой компьютерного зрения является обнаружение и интерпретация текста на изображении. Этот вид обработки часто называют *оптическим распознаванием символов* (optical character recognition, OCR).
## Чтение текста на изображении с помощью службы Computer Vision
Когнитивная служба **Computer Vision** обеспечивает поддержку OCR-задач, в том числе:
- API **для OCR**, который можно использовать для чтения текста на нескольких языках. Этот API используется асинхронно и может быть использован и для печатного, и для рукописного текста.
- API **для чтения**, оптимизированный для больших документов. Этот API может использоваться синхронно, и работает хорошо, когда нужно обнаружить и прочитать небольшое количество текста на изображении.
Эту службу можно использовать, создав ресурс либо **Computer Vision**, либо **Cognitive Services**.
Если вы еще этого не сделали, создайте ресурс **Cognitive Services** в своей подписке Azure.
> **Примечание**. Если у вас уже есть ресурс Cognitive Services, просто откройте его страницу **Быстрый запуск** и скопируйте его ключ и конечную точку в ячейку ниже. В противном случае следуйте приведенным ниже действиям для создания этого ресурса.
1. В другой вкладке браузера откройте портал Azure по адресу: https://portal.azure.com, войдя в систему под учетной записью Microsoft.
2. Нажмите кнопку **+Создать ресурс**, выполните поиск по строке *Cognitive Services* и создайте ресурс **Cognitive Services** со следующими настройками:
- **Подписка**. *Ваша подписка Azure*.
- **Группа ресурсов**. *Выберите или создайте группу ресурсов с уникальным именем*.
- **Регион**. *Выберите любой доступный регион*:
- **Имя**. *Введите уникальное имя*.
- **Ценовая категория**. S0
- **Подтверждаю, что прочитал и понял уведомления**. Выбрано.
3. Дождитесь завершения развертывания. Затем перейдите на свой ресурс Cognitive Services и на странице **Обзор** щелкните ссылку для управления ключами службы. Для подключения к вашему ресурсу когнитивных служб из клиентских приложений вам понадобятся конечная точка и ключи.
### Получение ключа и конечной точки для ресурса Cognitive Services
Для использования ресурса когнитивных служб клиентским приложениям необходимы их конечная точка и ключ аутентификации:
1. На портале Azure откройте страницу **Ключи и конечная точка** для вашего ресурса Cognitive Service, скопируйте **Ключ1** для вашего ресурса и вставьте его в приведенный ниже код, заменив подстановочный текст **YOUR_COG_KEY**.
2. Скопируйте **конечную точку** для своего ресурса и вставьте ее в код ниже, заменив **YOUR_COG_ENDPOINT**.
3. Выполните код в расположенной ниже ячейке с кодом, нажав на кнопку **Выполнить код в ячейке** (▷) слева от ячейки.
```
cog_key = 'YOUR_COG_KEY'
cog_endpoint = 'YOUR_COG_ENDPOINT'
print('Ready to use cognitive services at {} using key {}'.format(cog_endpoint, cog_key))
```
Теперь, когда вы установили ключ и конечную точку, можно использовать ресурс службы компьютерного зрения для извлечения текста из изображения.
Давайте начнем с API **для OCR**, который позволит вам синхронно анализировать изображение и читать любой содержащийся в нем текст. В этом случае у вас есть рекламное изображение для вымышленной розничной компании Northwind Traders, которое содержит некоторый текст. Чтобы прочитать его, выполните код из ячейки ниже.
```
from azure.cognitiveservices.vision.computervision import ComputerVisionClient
from msrest.authentication import CognitiveServicesCredentials
import matplotlib.pyplot as plt
from PIL import Image, ImageDraw
import os
%matplotlib inline
# Get a client for the computer vision service
computervision_client = ComputerVisionClient(cog_endpoint, CognitiveServicesCredentials(cog_key))
# Read the image file
image_path = os.path.join('data', 'ocr', 'advert.jpg')
image_stream = open(image_path, "rb")
# Use the Computer Vision service to find text in the image
read_results = computervision_client.recognize_printed_text_in_stream(image_stream)
# Process the text line by line
for region in read_results.regions:
for line in region.lines:
# Read the words in the line of text
line_text = ''
for word in line.words:
line_text += word.text + ' '
print(line_text.rstrip())
# Open image to display it.
fig = plt.figure(figsize=(7, 7))
img = Image.open(image_path)
draw = ImageDraw.Draw(img)
plt.axis('off')
plt.imshow(img)
```
Найденный на изображении текст организован в иерархическую структуру из областей, строк и слов, и код считывает их для получения результатов.
В результатах можно увидеть текст, прочитанный из изображения.
## Отображение ограничивающих прямоугольников
Результаты также включают координаты *ограничивающих прямоугольников* для строк текста и отдельных слов, найденных на изображении. Выполните код из ячейки ниже, чтобы увидеть координаты ограничивающих прямоугольников для строк текста на рекламном изображении, который вы получили ранее.
```
# Open image to display it.
fig = plt.figure(figsize=(7, 7))
img = Image.open(image_path)
draw = ImageDraw.Draw(img)
# Process the text line by line
for region in read_results.regions:
for line in region.lines:
# Show the position of the line of text
l,t,w,h = list(map(int, line.bounding_box.split(',')))
draw.rectangle(((l,t), (l+w, t+h)), outline='magenta', width=5)
# Read the words in the line of text
line_text = ''
for word in line.words:
line_text += word.text + ' '
print(line_text.rstrip())
# Show the image with the text locations highlighted
plt.axis('off')
plt.imshow(img)
```
В результате на изображении будут показаны ограничивающие прямоугольники для каждой строки текста.
## Использование API для чтения
API для OCR, который вы использовали ранее, хорошо работает для изображений с небольшим количеством текста. Когда нужно прочитать большие объемы текста, например отсканированные документы, можно использовать API **для чтения**. Это требует многоступенчатого процесса:
1. Представьте изображение в службу Computer Vision для асинхронного чтения и анализа.
2. Дождитесь завершения операции анализа.
3. Получите результаты анализа.
Выполните код в следующей ячейке, чтобы использовать этот процесс для чтения текста в отсканированном письме менеджеру магазина Northwind Traders.
```
from azure.cognitiveservices.vision.computervision import ComputerVisionClient
from azure.cognitiveservices.vision.computervision.models import OperationStatusCodes
from msrest.authentication import CognitiveServicesCredentials
import matplotlib.pyplot as plt
from PIL import Image
import time
import os
%matplotlib inline
# Read the image file
image_path = os.path.join('data', 'ocr', 'letter.jpg')
image_stream = open(image_path, "rb")
# Get a client for the computer vision service
computervision_client = ComputerVisionClient(cog_endpoint, CognitiveServicesCredentials(cog_key))
# Submit a request to read printed text in the image and get the operation ID
read_operation = computervision_client.read_in_stream(image_stream,
raw=True)
operation_location = read_operation.headers["Operation-Location"]
operation_id = operation_location.split("/")[-1]
# Wait for the asynchronous operation to complete
while True:
read_results = computervision_client.get_read_result(operation_id)
if read_results.status not in [OperationStatusCodes.running]:
break
time.sleep(1)
# If the operation was successfuly, process the text line by line
if read_results.status == OperationStatusCodes.succeeded:
for result in read_results.analyze_result.read_results:
for line in result.lines:
print(line.text)
# Open image to display it.
print('\n')
fig = plt.figure(figsize=(12,12))
img = Image.open(image_path)
plt.axis('off')
plt.imshow(img)
```
Просмотрите результаты. Вот полная расшифровка письма, состоящая в основном из печатного текста с рукописной подписью. Оригинальное изображение письма отображается под результатами OCR (возможно, вам понадобится прокрутка, чтобы его увидеть).
## Чтение рукописного текста
В предыдущем примере в запросе на анализ изображения был задан режим распознавания текста, который оптимизировал работу с *печатным* текстом. Однако обратите внимание, что была прочитана и рукописная подпись.
Такая возможность чтения рукописного текста чрезвычайно полезна. Предположим, вы написали заметку, содержащую список покупок, и хотите использовать приложение на телефоне для прочтения заметки и расшифровки содержащегося в ней текста.
Выполните код из ячейки ниже, чтобы посмотреть пример операции чтения рукописного списка покупок.
```
from azure.cognitiveservices.vision.computervision import ComputerVisionClient
from azure.cognitiveservices.vision.computervision.models import OperationStatusCodes
from msrest.authentication import CognitiveServicesCredentials
import matplotlib.pyplot as plt
from PIL import Image
import time
import os
%matplotlib inline
# Read the image file
image_path = os.path.join('data', 'ocr', 'note.jpg')
image_stream = open(image_path, "rb")
# Get a client for the computer vision service
computervision_client = ComputerVisionClient(cog_endpoint, CognitiveServicesCredentials(cog_key))
# Submit a request to read printed text in the image and get the operation ID
read_operation = computervision_client.read_in_stream(image_stream,
raw=True)
operation_location = read_operation.headers["Operation-Location"]
operation_id = operation_location.split("/")[-1]
# Wait for the asynchronous operation to complete
while True:
read_results = computervision_client.get_read_result(operation_id)
if read_results.status not in [OperationStatusCodes.running]:
break
time.sleep(1)
# If the operation was successfuly, process the text line by line
if read_results.status == OperationStatusCodes.succeeded:
for result in read_results.analyze_result.read_results:
for line in result.lines:
print(line.text)
# Open image to display it.
print('\n')
fig = plt.figure(figsize=(12,12))
img = Image.open(image_path)
plt.axis('off')
plt.imshow(img)
```
## Дополнительные сведения
Дополнительные сведения об использовании службы Computer Vision для распознавания текста см. в [документации по Computer Vision](https://docs.microsoft.com/ru-ru/azure/cognitive-services/computer-vision/concept-recognizing-text)
| github_jupyter |
<a href="https://colab.research.google.com/github/quinn-dougherty/DS-Unit-2-Sprint-3-Advanced-Regression/blob/master/module1-logistic-regression/Copy_of_LS_DS1_231_Logistic_Regression.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Lambda School Data Science - Logistic Regression
Logistic regression is the baseline for classification models, as well as a handy way to predict probabilities (since those too live in the unit interval). While relatively simple, it is also the foundation for more sophisticated classification techniques such as neural networks (many of which can effectively be thought of as networks of logistic models).
## Lecture - Where Linear goes Wrong
### Return of the Titanic 🚢
You've likely already explored the rich dataset that is the Titanic - let's use regression and try to predict survival with it. The data is [available from Kaggle](https://www.kaggle.com/c/titanic/data), so we'll also play a bit with [the Kaggle API](https://github.com/Kaggle/kaggle-api).
```
!pip install kaggle
# Note - you'll also have to sign up for Kaggle and authorize the API
# https://github.com/Kaggle/kaggle-api#api-credentials
# This essentially means uploading a kaggle.json file
# For Colab we can have it in Google Drive
from google.colab import drive
drive.mount('/content/drive')
%env KAGGLE_CONFIG_DIR=/content/drive/My Drive/
# You also have to join the Titanic competition to have access to the data
!kaggle competitions download -c titanic
# How would we try to do this with linear regression?
import pandas as pd
train_df = pd.read_csv('train.csv').dropna()
test_df = pd.read_csv('test.csv').dropna() # Unlabeled, for Kaggle submission
train_df.head()
train_df.describe()
from sklearn.linear_model import LinearRegression
X = train_df[['Pclass', 'Age', 'Fare']]
y = train_df.Survived
linear_reg = LinearRegression().fit(X, y)
linear_reg.score(X, y)
linear_reg.predict(test_df[['Pclass', 'Age', 'Fare']])
linear_reg.coef_
import numpy as np
test_case = np.array([[1, 5, 500]]) # Rich 5-year old in first class
linear_reg.predict(test_case)
np.dot(linear_reg.coef_, test_case.reshape(-1, 1)) + linear_reg.intercept_
from sklearn.linear_model import LogisticRegression
log_reg = LogisticRegression().fit(X, y)
log_reg.score(X, y)
log_reg.predict(test_df[['Pclass', 'Age', 'Fare']])
log_reg.predict(test_case)[0]
help(log_reg.predict)
log_reg.predict_proba(test_case)[0]
# What's the math?
log_reg.coef_
log_reg.intercept_
# The logistic sigmoid "squishing" function, implemented to accept numpy arrays
def sigmoid(x):
return 1 / (1 + np.e**(-x))
sigmoid(log_reg.intercept_ + np.dot(log_reg.coef_, np.transpose(test_case)))
```
So, clearly a more appropriate model in this situation! For more on the math, [see this Wikipedia example](https://en.wikipedia.org/wiki/Logistic_regression#Probability_of_passing_an_exam_versus_hours_of_study).
For live - let's tackle [another classification dataset on absenteeism](http://archive.ics.uci.edu/ml/datasets/Absenteeism+at+work) - it has 21 classes, but remember, scikit-learn LogisticRegression automatically handles more than two classes. How? By essentially treating each label as different (1) from some base class (0).
```
# Live - let's try absenteeism!
!wget http://archive.ics.uci.edu/ml/machine-learning-databases/00445/Absenteeism_at_work_AAA.zip
!unzip Absenteeism_at_work_AAA.zip
!head Absenteeism_at_work.csv
absent_df = pd.read_table('Absenteeism_at_work.csv', sep=';')
absent_df.head()
absent_df.shape
LogisticRegression
absent_df.columns
X = absent_df.drop('Reason for absence', axis='columns')
y = absent_df['Reason for absence']
absent_log1 = LogisticRegression().fit(X, y)
absent_log1.score(X, y)
absent_log1
absent_log1.coef_
?LogisticRegression
```
## Assignment - real-world classification
We're going to check out a larger dataset - the [FMA Free Music Archive data](https://github.com/mdeff/fma). It has a selection of CSVs with metadata and calculated audio features that you can load and try to use to classify genre of tracks. To get you started:
```
!wget https://os.unil.cloud.switch.ch/fma/fma_metadata.zip
!unzip fma_metadata.zip
from sklearn.linear_model import LogisticRegression
import pandas as pd
import numpy as np
import datetime as dt
tracks = pd.read_csv('fma_metadata/tracks.csv')
tracks.describe()
pd.set_option('display.max_columns', 55) # enough columns
tracks.head()
tracks.shape
```
This is the biggest data you've played with so far, and while it does generally fit in Colab, it can take awhile to run. That's part of the challenge!
Your tasks:
- Clean up the variable names in the dataframe
- Use logistic regression to fit a model predicting (primary/top) genre
- Inspect, iterate, and improve your model
- Answer the following questions (written, ~paragraph each):
- What are the best predictors of genre?
- What information isn't very useful for predicting genre?
- What surprised you the most about your results?
*Important caveats*:
- This is going to be difficult data to work with - don't let the perfect be the enemy of the good!
- Be creative in cleaning it up - if the best way you know how to do it is download it locally and edit as a spreadsheet, that's OK!
- If the data size becomes problematic, consider sampling/subsetting
- You do not need perfect or complete results - just something plausible that runs, and that supports the reasoning in your written answers
If you find that fitting a model to classify *all* genres isn't very good, it's totally OK to limit to the most frequent genres, or perhaps trying to combine or cluster genres as a preprocessing step. Even then, there will be limits to how good a model can be with just this metadata - if you really want to train an effective genre classifier, you'll have to involve the other data (see stretch goals).
This is real data - there is no "one correct answer", so you can take this in a variety of directions. Just make sure to support your findings, and feel free to share them as well! This is meant to be practice for dealing with other "messy" data, a common task in data science.
### Resources and stretch goals
- Check out the other .csv files from the FMA dataset, and see if you can join them or otherwise fit interesting models with them
- [Logistic regression from scratch in numpy](https://blog.goodaudience.com/logistic-regression-from-scratch-in-numpy-5841c09e425f) - if you want to dig in a bit more to both the code and math (also takes a gradient descent approach, introducing the logistic loss function)
- Create a visualization to show predictions of your model - ideally show a confidence interval based on error!
- Check out and compare classification models from scikit-learn, such as [SVM](https://scikit-learn.org/stable/modules/svm.html#classification), [decision trees](https://scikit-learn.org/stable/modules/tree.html#classification), and [naive Bayes](https://scikit-learn.org/stable/modules/naive_bayes.html). The underlying math will vary significantly, but the API (how you write the code) and interpretation will actually be fairly similar.
- Sign up for [Kaggle](https://kaggle.com), and find a competition to try logistic regression with
- (Not logistic regression related) If you enjoyed the assignment, you may want to read up on [music informatics](https://en.wikipedia.org/wiki/Music_informatics), which is how those audio features were actually calculated. The FMA includes the actual raw audio, so (while this is more of a longterm project than a stretch goal, and won't fit in Colab) if you'd like you can check those out and see what sort of deeper analysis you can do.
```
real_feats_list = [tracks[tracks.columns[k]].iloc[0] for k in range(len(tracks.columns))]
# for k in range(len(tracks.columns)):
# print(k, tracks[tracks.columns[k]].iloc[0])
tracks.isna().sum()
real_feats_list
tracks.columns
def up_to_prd(s):
if len(s)==0: return ''
elif s[0]=='.': return ''
else: return s[0] + up_to_prd(s[1:])
prefices = ['track_id'] + [up_to_prd(feat) for feat in tracks.columns[1:]]
ind_pairs = list(zip(prefices, real_feats_list))
index = pd.MultiIndex.from_tuples(ind_pairs, names=['first', 'second'])
df = (tracks
.drop([0,1], axis=0)
.T
.set_index(index)
.T)
df.head()
#for k in range(2,df.shape[0]):
# assert '<' in df.album.information.iloc[k]
#list(range(2,df.shape[0]))
#df['<' not in df.album.information.values].head()
# drop column that invovles html tags
onehot_genres = pd.get_dummies(df['track']['genre_top'])
oh_g_tuples = list(zip(['genre_top']*onehot_genres.shape[1], onehot_genres.columns))
onehot_genres_multidx = onehot_genres.T.set_index(pd.MultiIndex.from_tuples(oh_g_tuples)).T
#onehot_genres_multidx.head()
#pd.concat([df, onehot_genres], axis=1).head()
onehot_genres.dtypes
oh_g_tuples
onehot_genres_multidx.shape
df_wgenres = pd.concat([df.drop('genre_top', axis=1, level='second'), onehot_genres_multidx], axis=1)
df_wgenres.genre_top.head()
df_wgenres.dtypes
#df_wgenres.album.date_created.apply(lambda x: x.to_datetime())
#pd.to_datetime(df_wgenres.album[['date_created', 'date_released']])
def up_to_spc(s):
s = str(s)
if len(s)==0: return ''
elif s[0]==' ': return ''
else: return s[0] + up_to_spc(s[1:])
import datetime as dt
def todatetoord(feat):
df_wgenres.album[feat] = (pd.to_datetime(df_wgenres.album[feat]
.apply(up_to_spc))
.apply(dt.datetime.toordinal))
pass
#df_wgenres.album.date_created = pd.to_datetime(df_wgenres.album.date_created.apply(up_to_spc))
#df_wgenres.album.date_released = pd.to_datetime(df_wgenres.album.date_released)
todatetoord('date_created')
todatetoord('date_released')
df_wgenres.dtypes
lgr = LogisticRegression()
X = df_wgenres
y = df_wgenres.genre_top['Hip-Hop']
y.shape, X.shape
log_reg = LogisticRegression().fit(X, y)
log_reg.score(X, y)
''
```
| github_jupyter |
```
!pip install spacy
!python -m spacy download en_core_web_sm
import os
import pandas as pd
import requests
from bs4 import BeautifulSoup
from typing import List, Tuple, Union, Callable, Dict, Iterator
from collections import defaultdict
from difflib import SequenceMatcher
import spacy
from spacy.matcher import Matcher, PhraseMatcher
from spacy.tokens.doc import Doc
from spacy.tokens.span import Span
from spacy.tokens.token import Token
nlp = spacy.load("en_core_web_sm")
filenames = os.listdir('./hrfCases') # Wherever files are located
def similar(a: str, return_b: str, min_score: float) -> Union[str, None]:
"""
• Returns 2nd string if similarity score is above supplied
minimum score. Else, returns None.
"""
if SequenceMatcher(None, a, return_b).ratio() >= min_score:
return return_b
def similar_in_list(lst: Union[List[str], Iterator[str]]) -> Callable:
"""
• Uses a closure on supplied list to return a function that iterates over
the list in order to search for the first similar term. It's used widely
in the scraper.
"""
def impl(item: str, min_score: float) -> Union[str, None]:
for s in lst:
s = similar(item, s, min_score)
if s:
return s
return impl
class BIACase:
def __init__(self, text: str):
"""
• Input will be text from a BIA case pdf file, after the pdf has
been converted from PDF to text.
• Scraping works utilizing spaCy, tokenizing the text, and iterating
token by token searching for matching keywords.
"""
self.doc: Doc = nlp(text)
self.ents: Tuple[Span] = self.doc.ents
self.state = None
self.city = None
def get_interpreter(self):
return False
def get_interpreter_new(self):
"""
• If the terms "interpreter" or "translator" appear in the document,
the field will return whether the asylum seeker had access to an
interpreter during their hearings. Currently, the field's output is
dependent on occurrence of specific tokens in the document; this method
needs to be fine-tuned and validated.
"""
for token in self.doc:
sent = token.sent.text.lower()
similar_interpreter = similar_in_list(['interpreter', 'translator'])
s = similar_interpreter(token.text.lower(), 0.9)
if s == 'interpreter' or s == 'translator':
surrounding = self.get_surrounding_sents(token)
next_word = self.doc[token.i+1].text.lower()
if 'requested' in surrounding.text.lower() \
and 'granted' in surrounding.text.lower():
return True
elif 'requested' in surrounding.text.lower() \
and 'was present' in surrounding.text.lower():
return True
elif 'requested' in surrounding.text.lower() \
and 'granted' not in surrounding.text.lower():
return False
elif 'requested' in surrounding.text.lower() \
and 'was present' in surrounding.text.lower():
return False
return False
new_interpreter_dict = {}
old_interpreter_dict = {}
for file in filenames:
f = open(f"./hrfCases/{file}", "r", encoding='utf-8')
case = BIACase(f.read())
old_interpreter = case.get_interpreter()
old_interpreter_dict[file] = old_interpreter
new_interpreter = case.get_interpreter_new()
new_interpreter_dict[file] = new_interpreter
if old_interpreter != new_interpreter:
print('case: ', )
print('new outcome: ', new_interpreter)
print('old outcome: ', old_interpreter)
f.close()
new_interpreter_df = pd.DataFrame(new_interpreter_dict.items(), columns=['UUID', 'new_interpreter'])
old_interpreter_df = pd.DataFrame(old_interpreter_dict.items(), columns=['UUID', 'old_interpreter'])
```
# Retrieve correct manually extracted data for comparison.
```
df_csv = pd.read_csv('manually_scrapped.csv')
df_csv = df_csv[['UUID', 'interpreter']]
#remove .pdf
df_csv['UUID'] = df_csv['UUID'].str[0:-4]
#remove different ending of .txt file names
new_interpreter_df['UUID'] = new_outcome_df['UUID'].apply(lambda x : x[0:x.find('output-1-to-') - 1])
old_interpreter_df['UUID'] = old_outcome_df['UUID'].apply(lambda x : x[0:x.find('output-1-to-') - 1])
combined_df = df_csv.merge(new_interpreter_df, on='UUID', how='outer').merge(old_interpreter_df, on='UUID', how='outer')
combined_df.head()
combined_df['old_accurate'] = combined_df.apply(compare_outcomes, args=['old'], axis=1)
combined_df['new_accurate'] = combined_df.apply(compare_outcomes, args=['new'], axis=1)
old_accuracy = combined_df['old_accurate'].sum()/len(combined_df)*100
new_accuracy = combined_df['new_accurate'].sum()/len(combined_df)*100
print('old accuracy: ', old_accuracy, "%")
print('new accuracy: ', new_accuracy, "%")
print("improvement: ", new_accuracy - old_accuracy, "%")
diff_df = combined_df[combined_df['new_accurate'] == False]
print(len(diff_df))
diff_df.head(20)
changes_df = combined_df[combined_df['new_accurate'] != combined_df['old_accurate']]
print(len(changes_df))
changes_df
```
| github_jupyter |
# Character level language model - Dinosaurus land
Welcome to Dinosaurus Island! 65 million years ago, dinosaurs existed, and in this assignment they are back. You are in charge of a special task. Leading biology researchers are creating new breeds of dinosaurs and bringing them to life on earth, and your job is to give names to these dinosaurs. If a dinosaur does not like its name, it might go beserk, so choose wisely!
<table>
<td>
<img src="images/dino.jpg" style="width:250;height:300px;">
</td>
</table>
Luckily you have learned some deep learning and you will use it to save the day. Your assistant has collected a list of all the dinosaur names they could find, and compiled them into this [dataset](dinos.txt). (Feel free to take a look by clicking the previous link.) To create new dinosaur names, you will build a character level language model to generate new names. Your algorithm will learn the different name patterns, and randomly generate new names. Hopefully this algorithm will keep you and your team safe from the dinosaurs' wrath!
By completing this assignment you will learn:
- How to store text data for processing using an RNN
- How to synthesize data, by sampling predictions at each time step and passing it to the next RNN-cell unit
- How to build a character-level text generation recurrent neural network
- Why clipping the gradients is important
We will begin by loading in some functions that we have provided for you in `rnn_utils`. Specifically, you have access to functions such as `rnn_forward` and `rnn_backward` which are equivalent to those you've implemented in the previous assignment.
```
import numpy as np
from utils import *
import random
from random import shuffle
```
## 1 - Problem Statement
### 1.1 - Dataset and Preprocessing
Run the following cell to read the dataset of dinosaur names, create a list of unique characters (such as a-z), and compute the dataset and vocabulary size.
```
data = open('dinos.txt', 'r').read()
data= data.lower()
chars = list(set(data))
data_size, vocab_size = len(data), len(chars)
print('There are %d total characters and %d unique characters in your data.' % (data_size, vocab_size))
```
The characters are a-z (26 characters) plus the "\n" (or newline character), which in this assignment plays a role similar to the `<EOS>` (or "End of sentence") token we had discussed in lecture, only here it indicates the end of the dinosaur name rather than the end of a sentence. In the cell below, we create a python dictionary (i.e., a hash table) to map each character to an index from 0-26. We also create a second python dictionary that maps each index back to the corresponding character character. This will help you figure out what index corresponds to what character in the probability distribution output of the softmax layer. Below, `char_to_ix` and `ix_to_char` are the python dictionaries.
```
char_to_ix = { ch:i for i,ch in enumerate(sorted(chars)) }
ix_to_char = { i:ch for i,ch in enumerate(sorted(chars)) }
print(ix_to_char)
```
### 1.2 - Overview of the model
Your model will have the following structure:
- Initialize parameters
- Run the optimization loop
- Forward propagation to compute the loss function
- Backward propagation to compute the gradients with respect to the loss function
- Clip the gradients to avoid exploding gradients
- Using the gradients, update your parameter with the gradient descent update rule.
- Return the learned parameters
<img src="images/rnn.png" style="width:450;height:300px;">
<caption><center> **Figure 1**: Recurrent Neural Network, similar to what you had built in the previous notebook "Building a RNN - Step by Step". </center></caption>
At each time-step, the RNN tries to predict what is the next character given the previous characters. The dataset $X = (x^{\langle 1 \rangle}, x^{\langle 2 \rangle}, ..., x^{\langle T_x \rangle})$ is a list of characters in the training set, while $Y = (y^{\langle 1 \rangle}, y^{\langle 2 \rangle}, ..., y^{\langle T_x \rangle})$ is such that at every time-step $t$, we have $y^{\langle t \rangle} = x^{\langle t+1 \rangle}$.
## 2 - Building blocks of the model
In this part, you will build two important blocks of the overall model:
- Gradient clipping: to avoid exploding gradients
- Sampling: a technique used to generate characters
You will then apply these two functions to build the model.
### 2.1 - Clipping the gradients in the optimization loop
In this section you will implement the `clip` function that you will call inside of your optimization loop. Recall that your overall loop structure usually consists of a forward pass, a cost computation, a backward pass, and a parameter update. Before updating the parameters, you will perform gradient clipping when needed to make sure that your gradients are not "exploding," meaning taking on overly large values.
In the exercise below, you will implement a function `clip` that takes in a dictionary of gradients and returns a clipped version of gradients if needed. There are different ways to clip gradients; we will use a simple element-wise clipping procedure, in which every element of the gradient vector is clipped to lie between some range [-N, N]. More generally, you will provide a `maxValue` (say 10). In this example, if any component of the gradient vector is greater than 10, it would be set to 10; and if any component of the gradient vector is less than -10, it would be set to -10. If it is between -10 and 10, it is left alone.
<img src="images/clip.png" style="width:400;height:150px;">
<caption><center> **Figure 2**: Visualization of gradient descent with and without gradient clipping, in a case where the network is running into slight "exploding gradient" problems. </center></caption>
**Exercise**: Implement the function below to return the clipped gradients of your dictionary `gradients`. Your function takes in a maximum threshold and returns the clipped versions of your gradients. You can check out this [hint](https://docs.scipy.org/doc/numpy-1.13.0/reference/generated/numpy.clip.html).
```
### GRADED FUNCTION: clip
def clip(gradients, maxValue):
'''
Clips the gradients' values between minimum and maximum.
Arguments:
gradients -- a dictionary containing the gradients "dWaa", "dWax", "dWya", "db", "dby"
maxValue -- everything above this number is set to this number, and everything less than -maxValue is set to -maxValue
Returns:
gradients -- a dictionary with the clipped gradients.
'''
dWaa, dWax, dWya, db, dby = gradients['dWaa'], gradients['dWax'], gradients['dWya'], gradients['db'], gradients['dby']
### START CODE HERE ###
# clip to mitigate exploding gradients, loop over [dWax, dWaa, dWya, db, dby]. (≈2 lines)
for gradient in [dWax, dWaa, dWya, db, dby]:
None
### END CODE HERE ###
gradients = {"dWaa": dWaa, "dWax": dWax, "dWya": dWya, "db": db, "dby": dby}
return gradients
np.random.seed(3)
dWax = np.random.randn(5,3)*10
dWaa = np.random.randn(5,5)*10
dWya = np.random.randn(2,5)*10
db = np.random.randn(5,1)*10
dby = np.random.randn(2,1)*10
gradients = {"dWax": dWax, "dWaa": dWaa, "dWya": dWya, "db": db, "dby": dby}
gradients = clip(gradients, 10)
print("gradients[\"dWaa\"][1][2] =", gradients["dWaa"][1][2])
print("gradients[\"dWax\"][3][1] =", gradients["dWax"][3][1])
print("gradients[\"dWya\"][1][2] =", gradients["dWya"][1][2])
print("gradients[\"db\"][4] =", gradients["db"][4])
print("gradients[\"dby\"][1] =", gradients["dby"][1])
```
** Expected output:**
<table>
<tr>
<td>
**gradients["dWaa"][1][2] **
</td>
<td>
10.0
</td>
</tr>
<tr>
<td>
**gradients["dWax"][3][1]**
</td>
<td>
-10.0
</td>
</td>
</tr>
<tr>
<td>
**gradients["dWya"][1][2]**
</td>
<td>
0.29713815361
</td>
</tr>
<tr>
<td>
**gradients["db"][4]**
</td>
<td>
[ 10.]
</td>
</tr>
<tr>
<td>
**gradients["dby"][1]**
</td>
<td>
[ 8.45833407]
</td>
</tr>
</table>
### 2.2 - Sampling
Now assume that your model is trained. You would like to generate new text (characters). The process of generation is explained in the picture below:
<img src="images/dinos3.png" style="width:500;height:300px;">
<caption><center> **Figure 3**: In this picture, we assume the model is already trained. We pass in $x^{\langle 1\rangle} = \vec{0}$ at the first time step, and have the network then sample one character at a time. </center></caption>
**Exercise**: Implement the `sample` function below to sample characters. You need to carry out 4 steps:
- **Step 1**: Pass the network the first "dummy" input $x^{\langle 1 \rangle} = \vec{0}$ (the vector of zeros). This is the default input before we've generated any characters. We also set $a^{\langle 0 \rangle} = \vec{0}$
- **Step 2**: Run one step of forward propagation to get $a^{\langle 1 \rangle}$ and $\hat{y}^{\langle 1 \rangle}$. Here are the equations:
$$ a^{\langle t+1 \rangle} = \tanh(W_{ax} x^{\langle t \rangle } + W_{aa} a^{\langle t \rangle } + b)\tag{1}$$
$$ z^{\langle t + 1 \rangle } = W_{ya} a^{\langle t + 1 \rangle } + b_y \tag{2}$$
$$ \hat{y}^{\langle t+1 \rangle } = softmax(z^{\langle t + 1 \rangle })\tag{3}$$
Note that $\hat{y}^{\langle t+1 \rangle }$ is a (softmax) probability vector (its entries are between 0 and 1 and sum to 1). $\hat{y}^{\langle t+1 \rangle}_i$ represents the probability that the character indexed by "i" is the next character. We have provided a `softmax()` function that you can use.
- **Step 3**: Carry out sampling: Pick the next character's index according to the probability distribution specified by $\hat{y}^{\langle t+1 \rangle }$. This means that if $\hat{y}^{\langle t+1 \rangle }_i = 0.16$, you will pick the index "i" with 16% probability. To implement it, you can use [`np.random.choice`](https://docs.scipy.org/doc/numpy-1.13.0/reference/generated/numpy.random.choice.html).
Here is an example of how to use `np.random.choice()`:
```python
np.random.seed(0)
p = np.array([0.1, 0.0, 0.7, 0.2])
index = np.random.choice([0, 1, 2, 3], p = p.ravel())
```
This means that you will pick the `index` according to the distribution:
$P(index = 0) = 0.1, P(index = 1) = 0.0, P(index = 2) = 0.7, P(index = 3) = 0.2$.
- **Step 4**: The last step to implement in `sample()` is to overwrite the variable `x`, which currently stores $x^{\langle t \rangle }$, with the value of $x^{\langle t + 1 \rangle }$. You will represent $x^{\langle t + 1 \rangle }$ by creating a one-hot vector corresponding to the character you've chosen as your prediction. You will then forward propagate $x^{\langle t + 1 \rangle }$ in Step 1 and keep repeating the process until you get a "\n" character, indicating you've reached the end of the dinosaur name.
```
# GRADED FUNCTION: sample
def sample(parameters, char_to_ix, seed):
"""
Sample a sequence of characters according to a sequence of probability distributions output of the RNN
Arguments:
parameters -- python dictionary containing the parameters Waa, Wax, Wya, by, and b.
char_to_ix -- python dictionary mapping each character to an index.
seed -- used for grading purposes. Do not worry about it.
Returns:
indices -- a list of length n containing the indexes of the sampled characters.
"""
# Retrieve parameters and relevant shapes from "parameters" dictionary
Waa, Wax, Wya, by, b = parameters['Waa'], parameters['Wax'], parameters['Wya'], parameters['by'], parameters['b']
vocab_size = by.shape[0]
n_a = Waa.shape[1]
### START CODE HERE ###
# Step 1: Create the one-hot vector x for the first character (initializing the sequence generation). (≈1 line)
x = None
# Step 1': Initialize a_prev as zeros (≈1 line)
a_prev = None
# Create an empty list of indices, this is the list which will contain the list of indexes of the characters to generate (≈1 line)
indices = None
# Idx is a flag to detect a newline character, we initialize it to -1
idx = -1
# Loop over time-steps t. At each time-step, sample a character from a probability distribution and append
# its index to "indexes". We'll stop if we reach 50 characters (which should be very unlikely with a well
# trained model), which helps debugging and prevents entering an infinite loop.
counter = 0
newline_character = char_to_ix['\n']
while (idx != newline_character and counter != 50):
# Step 2: Forward propagate x using the equations (1), (2) and (3)
a = None
z = None
y = None
# for grading purposes
np.random.seed(counter+seed)
# Step 3: Sample the index of a character within the vocabulary from the probability distribution y
idx = None
# Append the index to "indices"
None
# Step 4: Overwrite the input character as the one corresponding to the sampled index.
x = None
x[None] = None
# Update "a_prev" to be "a"
a_prev = None
# for grading purposes
seed += 1
counter +=1
### END CODE HERE ###
if (counter == 50):
indices.append(char_to_ix['\n'])
return indices
np.random.seed(2)
n, n_a = 20, 100
a0 = np.random.randn(n_a, 1)
i0 = 1 # first character is ix_to_char[i0]
Wax, Waa, Wya = np.random.randn(n_a, vocab_size), np.random.randn(n_a, n_a), np.random.randn(vocab_size, n_a)
b, by = np.random.randn(n_a, 1), np.random.randn(vocab_size, 1)
parameters = {"Wax": Wax, "Waa": Waa, "Wya": Wya, "b": b, "by": by}
indexes = sample(parameters, char_to_ix, 0)
print("Sampling:")
print("list of sampled indices:", indexes)
print("list of sampled characters:", [ix_to_char[i] for i in indexes])
```
** Expected output:**
<table>
<tr>
<td>
**list of sampled indices:**
</td>
<td>
[18, 2, 26, 0]
</td>
</tr><tr>
<td>
**list of sampled characters:**
</td>
<td>
['r', 'b', 'z', '\n']
</td>
</tr>
</table>
## 3 - Building the language model
It is time to build the character-level language model for text generation.
### 3.1 - Gradient descent
In this section you will implement a function performing one step of stochastic gradient descent (with clipped gradients). You will go through the training examples one at a time, so the optimization algorithm will be stochastic gradient descent. As a reminder, here are the steps of a common optimization loop for an RNN:
- Forward propagate through the RNN to compute the loss
- Backward propagate through time to compute the gradients of the loss with respect to the parameters
- Clip the gradients if necessary
- Update your parameters using gradient descent
**Exercise**: Implement this optimization process (one step of stochastic gradient descent).
We provide you with the following functions:
```python
def rnn_forward(X, Y, a_prev, parameters):
""" Performs the forward propagation through the RNN and computes the cross-entropy loss.
It returns the loss' value as well as a "cache" storing values to be used in the backpropagation."""
....
return loss, cache
def rnn_backward(X, Y, parameters, cache):
""" Performs the backward propagation through time to compute the gradients of the loss with respect
to the parameters. It returns also all the hidden states."""
...
return gradients, a
def update_parameters(parameters, gradients, learning_rate):
""" Updates parameters using the Gradient Descent Update Rule."""
...
return parameters
```
```
# GRADED FUNCTION: optimize
def optimize(X, Y, a_prev, parameters, learning_rate = 0.01):
"""
Execute one step of the optimization to train the model.
Arguments:
X -- list of integers, where each integer is a number that maps to a character in the vocabulary.
Y -- list of integers, exactly the same as X but shifted one index to the left.
a_prev -- previous hidden state.
parameters -- python dictionary containing:
Wax -- Weight matrix multiplying the input, numpy array of shape (n_a, n_x)
Waa -- Weight matrix multiplying the hidden state, numpy array of shape (n_a, n_a)
Wya -- Weight matrix relating the hidden-state to the output, numpy array of shape (n_y, n_a)
b -- Bias, numpy array of shape (n_a, 1)
by -- Bias relating the hidden-state to the output, numpy array of shape (n_y, 1)
learning_rate -- learning rate for the model.
Returns:
loss -- value of the loss function (cross-entropy)
gradients -- python dictionary containing:
dWax -- Gradients of input-to-hidden weights, of shape (n_a, n_x)
dWaa -- Gradients of hidden-to-hidden weights, of shape (n_a, n_a)
dWya -- Gradients of hidden-to-output weights, of shape (n_y, n_a)
db -- Gradients of bias vector, of shape (n_a, 1)
dby -- Gradients of output bias vector, of shape (n_y, 1)
a[len(X)-1] -- the last hidden state, of shape (n_a, 1)
"""
### START CODE HERE ###
# Forward propagate through time (≈1 line)
loss, cache = None
# Backpropagate through time (≈1 line)
gradients, a = None
# Clip your gradients between -5 (min) and 5 (max) (≈1 line)
gradients = None
# Update parameters (≈1 line)
parameters = None
### END CODE HERE ###
return loss, gradients, a[len(X)-1]
np.random.seed(1)
vocab_size, n_a = 27, 100
a_prev = np.random.randn(n_a, 1)
Wax, Waa, Wya = np.random.randn(n_a, vocab_size), np.random.randn(n_a, n_a), np.random.randn(vocab_size, n_a)
b, by = np.random.randn(n_a, 1), np.random.randn(vocab_size, 1)
parameters = {"Wax": Wax, "Waa": Waa, "Wya": Wya, "b": b, "by": by}
X = [12,3,5,11,22,3]
Y = [4,14,11,22,25, 26]
loss, gradients, a_last = optimize(X, Y, a_prev, parameters, learning_rate = 0.01)
print("Loss =", loss)
print("gradients[\"dWaa\"][1][2] =", gradients["dWaa"][1][2])
print("np.argmax(gradients[\"dWax\"]) =", np.argmax(gradients["dWax"]))
print("gradients[\"dWya\"][1][2] =", gradients["dWya"][1][2])
print("gradients[\"db\"][4] =", gradients["db"][4])
print("gradients[\"dby\"][1] =", gradients["dby"][1])
print("a_last[4] =", a_last[4])
```
** Expected output:**
<table>
<tr>
<td>
**Loss **
</td>
<td>
126.503975722
</td>
</tr>
<tr>
<td>
**gradients["dWaa"][1][2]**
</td>
<td>
0.194709315347
</td>
<tr>
<td>
**np.argmax(gradients["dWax"])**
</td>
<td> 93
</td>
</tr>
<tr>
<td>
**gradients["dWya"][1][2]**
</td>
<td> -0.007773876032
</td>
</tr>
<tr>
<td>
**gradients["db"][4]**
</td>
<td> [-0.06809825]
</td>
</tr>
<tr>
<td>
**gradients["dby"][1]**
</td>
<td>[ 0.01538192]
</td>
</tr>
<tr>
<td>
**a_last[4]**
</td>
<td> [-1.]
</td>
</tr>
</table>
### 3.2 - Training the model
Given the dataset of dinosaur names, we use each line of the dataset (one name) as one training example. Every 100 steps of stochastic gradient descent, you will sample 10 randomly chosen names to see how the algorithm is doing. Remember to shuffle the dataset, so that stochastic gradient descent visits the examples in random order.
**Exercise**: Follow the instructions and implement `model()`. When `examples[index]` contains one dinosaur name (string), to create an example (X, Y), you can use this:
```python
index = j % len(examples)
X = [None] + [char_to_ix[ch] for ch in examples[index]]
Y = X[1:] + [char_to_ix["\n"]]
```
Note that we use: `index= j % len(examples)`, where `j = 1....num_iterations`, to make sure that `examples[index]` is always a valid statement (`index` is smaller than `len(examples)`).
The first entry of `X` being `None` will be interpreted by `rnn_forward()` as setting $x^{\langle 0 \rangle} = \vec{0}$. Further, this ensures that `Y` is equal to `X` but shifted one step to the left, and with an additional "\n" appended to signify the end of the dinosaur name.
```
# GRADED FUNCTION: model
def model(data, ix_to_char, char_to_ix, num_iterations = 35000, n_a = 50, dino_names = 7, vocab_size = 27):
"""
Trains the model and generates dinosaur names.
Arguments:
data -- text corpus
ix_to_char -- dictionary that maps the index to a character
char_to_ix -- dictionary that maps a character to an index
num_iterations -- number of iterations to train the model for
n_a -- number of hidden neurons in the softmax layer
dino_names -- number of dinosaur names you want to sample at each iteration.
vocab_size -- number of unique characters found in the text, size of the vocabulary
Returns:
parameters -- learned parameters
"""
# Retrieve n_x and n_y from vocab_size
n_x, n_y = vocab_size, vocab_size
# Initialize parameters
parameters = initialize_parameters(n_a, n_x, n_y)
# Initialize loss (this is required because we want to smooth our loss, don't worry about it)
loss = get_initial_loss(vocab_size, dino_names)
# Build list of all dinosaur names (training examples).
with open("dinos.txt") as f:
examples = f.readlines()
examples = [x.lower().strip() for x in examples]
# Shuffle list of all dinosaur names
shuffle(examples)
# Initialize the hidden state of your LSTM
a_prev = np.zeros((n_a, 1))
# Optimization loop
for j in range(num_iterations):
### START CODE HERE ###
# Use the hint above to define one training example (X,Y) (≈ 2 lines)
index = None
X = None
Y = None
# Perform one optimization step: Forward-prop -> Backward-prop -> Clip -> Update parameters
# Choose a learning rate of 0.01
curr_loss, gradients, a_prev = None
### END CODE HERE ###
# Use a latency trick to keep the loss smooth. It happens here to accelerate the training.
loss = smooth(loss, curr_loss)
# Every 2000 Iteration, generate "n" characters thanks to sample() to check if the model is learning properly
if j % 2000 == 0:
print('Iteration: %d, Loss: %f' % (j, loss) + '\n')
# The number of dinosaur names to print
seed = 0
for name in range(dino_names):
# Sample indexes and print them
sampled_indexes = sample(parameters, char_to_ix, seed)
print_sample(sampled_indexes, ix_to_char)
seed += 1 # To get the same result for grading purposed, increment the seed by one.
print('\n')
return parameters
```
Run the following cell, you should observe your model outputting random-looking characters at the first iteration. After a few thousand iterations, your model should learn to generate reasonable-looking names.
```
parameters = model(data, ix_to_char, char_to_ix)
```
## Conclusion
You can see that your algorithm has started to generate plausible dinosaur names towards the end of the training. At first, it was generating random characters, but towards the end you could see dinosaur names with cool endings. Feel free to run the algorithm even longer and play with hyperparameters to see if you can get even better results. Our implemetation generated some really cool names like `maconucon`, `marloralus` and `macingsersaurus`. Your model hopefully also learned that dinosaur names tend to end in `saurus`, `don`, `aura`, `tor`, etc.
If your model generates some non-cool names, don't blame the model entirely--not all actual dinosaur names sound cool. (For example, `dromaeosauroides` is an actual dinosaur name and is in the training set.) But this model should give you a set of candidates from which you can pick the coolest!
This assignment had used a relatively small dataset, so that you could train an RNN quickly on a CPU. Training a model of the english language requires a much bigger dataset, and usually needs much more computation, and could run for many hours on GPUs. We ran our dinosaur name for quite some time, and so far our favoriate name is the great, undefeatable, and fierce: Mangosaurus!
<img src="images/mangosaurus.jpeg" style="width:250;height:300px;">
## 4 - Writing like Shakespeare
The rest of this notebook is optional and is not graded, but we hope you'll do it anyway since it's quite fun and informative.
A similar (but more complicated) task is to generate Shakespeare poems. Instead of learning from a dataset of Dinosaur names you can use a collection of Shakespearian poems. Using LSTM cells, you can learn longer term dependencies that span many characters in the text--e.g., where a character appearing somewhere a sequence can influence what should be a different character much much later in ths sequence. These long term dependencies were less important with dinosaur names, since the names were quite short.
<img src="images/shakespeare.jpg" style="width:500;height:400px;">
<caption><center> Let's become poets! </center></caption>
We have implemented a Shakespeare poem generator with Keras. Run the following cell to load the required packages and models. This may take a few minutes.
```
from __future__ import print_function
from keras.callbacks import LambdaCallback
from keras.models import Model, load_model, Sequential
from keras.layers import Dense, Activation, Dropout, Input, Masking
from keras.layers import LSTM
from keras.utils.data_utils import get_file
from keras.preprocessing.sequence import pad_sequences
from shakespeare_utils import *
import sys
import io
```
To save you some time, we have already trained a model for ~1000 epochs on a collection of Shakespearian poems called [*"The Sonnets"*](shakespeare.txt).
Let's train the model for one more epoch. When it finishes training for an epoch---this will also take a few minutes---you can run `generate_output`, which will prompt asking you for an input (`<`40 characters). The poem will start with your sentence, and our RNN-Shakespeare will complete the rest of the poem for you! For example, try "Forsooth this maketh no sense " (don't enter the quotation marks). Depending on whether you include the space at the end, your results might also differ--try it both ways, and try other inputs as well.
```
print_callback = LambdaCallback(on_epoch_end=on_epoch_end)
model.fit(x, y, batch_size=128, epochs=1, callbacks=[print_callback])
# Run this cell to try with different inputs without having to re-train the model
generate_output()
```
The RNN-Shakespeare model is very similar to the one you have built for dinosaur names. The only major differences are:
- LSTMs instead of the basic RNN to capture longer-range dependencies
- The model is a deeper, stacked LSTM model (2 layer)
- Using Keras instead of python to simplify the code
If you want to learn more, you can also check out the Keras Team's text generation implementation on GitHub: https://github.com/keras-team/keras/blob/master/examples/lstm_text_generation.py.
Congratulations on finishing this notebook!
**References**:
- This exercise took inspiration from Andrej Karpathy's implementation: https://gist.github.com/karpathy/d4dee566867f8291f086. To learn more about text generation, also check out Karpathy's [blog post](http://karpathy.github.io/2015/05/21/rnn-effectiveness/).
- For the Shakespearian poem generator, our implementation was based on the implementation of an LSTM text generator by the Keras team: https://github.com/keras-team/keras/blob/master/examples/lstm_text_generation.py
| github_jupyter |
# Combining Factors from Multiple Sources
The Quantopian research environment is tailored to the rapid testing of predictive alpha factors. The process is very similar because it builds on zipline, but offers much richer access to data sources. The following code sample illustrates how to compute alpha factors not only from market data as previously but also from fundamental and alternative data.
Quantopian provides several hundred MorningStar fundamental variables for free and also includes stocktwits signals as an example of an alternative data source. There are also custom universe definitions such as QTradableStocksUS that applies several filters to limit the backtest universe to stocks that were likely tradeable under realistic market conditions.
```
from quantopian.research import run_pipeline
from quantopian.pipeline import Pipeline
from quantopian.pipeline.data.builtin import USEquityPricing
from quantopian.pipeline.data.morningstar import income_statement, operation_ratios, balance_sheet
from quantopian.pipeline.data.psychsignal import stocktwits
from quantopian.pipeline.factors import CustomFactor, SimpleMovingAverage, Returns
from quantopian.pipeline.filters import QTradableStocksUS
import numpy as np
import pandas as pd
from time import time
# periods in trading days
MONTH = 21
QTR = 4 * MONTH
YEAR = 12 * MONTH
```
We will use a custom `AggregateFundamentals` class to use the last reported fundamental data point. This aims to address the fact that fundamentals are reported quarterly, and Quantopian does not currently provide an easy way to aggregate historical data, say to obtain the sum of the last four quarters, on a rolling basis.
We will again use the custom `MeanReversion` factor from [01_single_factor_zipline](01_single_factor_zipline.ipynb). We will also compute several other factors for the given universe definition using the `rank()` method's mask parameter.
This algorithm uses a naive method to combine the six individual factors by simply adding the ranks of assets for each of these factors. Instead of equal weights, we would like to take into account the relative importance and incremental information in predicting future returns. The ML algorithms of the next chapters will allow us to do exactly this, using the same backtesting framework.
Execution also relies on run_algorithm(), but the return DataFrame on the Quantopian platform only contains the factor values created by the Pipeline. This is convenient because this data format can be used as input for alphalens, the library for the evaluation of the predictive performance of alpha factors.
```
class AggregateFundamentals(CustomFactor):
def compute(self, today, assets, out, inputs):
out[:] = inputs[0]
class MeanReversion(CustomFactor):
inputs = [Returns(window_length=MONTH)]
window_length = YEAR
def compute(self, today, assets, out, monthly_returns):
df = pd.DataFrame(monthly_returns)
out[:] = df.iloc[-1].sub(df.mean()).div(df.std())
def compute_factors():
universe = QTradableStocksUS()
profitability = (AggregateFundamentals(inputs = [income_statement.gross_profit],
window_length = YEAR) /
balance_sheet.total_assets.latest).rank(mask=universe)
roic = operation_ratios.roic.latest.rank(mask=universe)
ebitda_yield = (AggregateFundamentals(inputs = [income_statement.ebitda],
window_length = YEAR) /
USEquityPricing.close.latest).rank(mask=universe)
mean_reversion = MeanReversion().rank(mask=universe)
price_momentum = Returns(window_length=QTR).rank(mask=universe)
sentiment = SimpleMovingAverage(inputs=[stocktwits.bull_minus_bear],
window_length=5).rank(mask=universe)
factor = profitability + roic + ebitda_yield + mean_reversion + price_momentum + sentiment
return Pipeline(
columns={'Profitability': profitability,
'ROIC': roic,
'EBITDA Yield': ebitda_yield,
"Mean Reversion (1M)": mean_reversion,
'Sentiment': sentiment,
"Price Momentum (3M)": price_momentum,
'Alpha Factor': factor})
start_timer = time()
start = pd.Timestamp("2015-01-01")
end = pd.Timestamp("2018-01-01")
results = run_pipeline(compute_factors(), start_date=start, end_date=end)
results.index.names = ['date', 'security']
print("Pipeline run time {:.2f} secs".format(time() - start_timer))
results.head()
```
| github_jupyter |
# Wage Profile
Code to accompany Lecture on
Regression
Jiaming Mao (<jmao@xmu.edu.cn>)
<https://jiamingmao.github.io>
```
import pandas as pd
import numpy as np
import statsmodels.api as sm
import matplotlib.pyplot as plt
from sklearn.linear_model import LinearRegression
from patsy import dmatrix
%matplotlib inline
data = pd.read_csv("Wage.csv")
data.head()
age = data.age
wage = data.wage
age_grid = np.arange(age.min(),age.max())
```
## Linear Regression
```
model = LinearRegression(fit_intercept=True)
model.fit(age[:,np.newaxis], wage)
print("slope: ", model.coef_[0])
print("intercept:", model.intercept_)
pred = model.predict(age_grid[:, np.newaxis])
plt.figure(figsize=(10,8))
plt.scatter(age, wage, facecolor='None', edgecolor='k', alpha=0.1, label='')
plt.plot(age_grid, pred, color='brown', label='Linear')
plt.legend()
plt.xlabel('age')
plt.ylabel('wage')
```
## Polynomial Regression
```
fit = np.polyfit(age, wage, 4)
model = np.poly1d(fit)
print(model)
pred = model(age_grid)
plt.figure(figsize=(10,8))
plt.scatter(age, wage, facecolor='None', edgecolor='k', alpha=0.1, label='')
plt.plot(age_grid, pred, color='darkblue', label='Degree-4 Polynomial')
plt.legend()
plt.xlabel('age')
plt.ylabel('wage')
```
## Piecewise Constant Regression
```
age_cut, bins = pd.cut(age, 4, retbins=True, right=True)
age_dummies = pd.get_dummies(age_cut)
fit = sm.GLM(wage, age_dummies).fit()
fit.params
bin_mapping = np.digitize(age_grid, bins)
age_grid_dummies = pd.get_dummies(bin_mapping)
pred = fit.predict(age_grid_dummies)
plt.figure(figsize=(10,8))
plt.scatter(age, wage, facecolor='None', edgecolor='k', alpha=0.1, label='')
plt.plot(age_grid, pred, color='darkgreen', label='Piecewise Constant')
plt.legend()
plt.xlabel('age')
plt.ylabel('wage')
```
## Cubic Spline and Natural Cubic Spline
```
age_cs = dmatrix("bs(age, knots=(25,40,60), degree=3, include_intercept=False)",{"age": age},return_type='dataframe')
fit1 = sm.GLM(wage, age_cs).fit()
fit1.params
age_ncs = dmatrix("cr(age, knots=(25,40,60))", {"age": age}, return_type='dataframe')
fit2 = sm.GLM(wage, age_ncs).fit()
fit2.params
pred1 = fit1.predict(dmatrix("bs(age_grid, knots=(25,40,60), include_intercept=False)", {"age_grid": age_grid}, return_type='dataframe'))
pred2 = fit2.predict(dmatrix("cr(age_grid, knots=(25,40,60))", {"age_grid": age_grid}, return_type='dataframe'))
plt.figure(figsize=(10,8))
plt.scatter(age, wage, facecolor='None', edgecolor='k', alpha=0.1, label='')
plt.plot(age_grid, pred1, color='cornflowerblue', label='Cubic Spline')
plt.plot(age_grid, pred2, color='darkorange', label='Natural Cubic Spline')
plt.legend()
plt.xlabel('age')
plt.ylabel('wage')
```
| github_jupyter |
```
# Covid19 Analysis
# libraries
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import roc_curve
from sklearn.metrics import auc
from sklearn.metrics import accuracy_score
import pickle
from sklearn.metrics import r2_score
from sklearn.metrics import average_precision_score
from sklearn.metrics import precision_recall_curve
from xgboost import XGBClassifier
from haversine import haversine, Unit
from haversine import haversine_vector
from mlxtend.evaluate import bootstrap
# reading dataset
# https://opendatasus.saude.gov.br/dataset/bd-srag-2020
df = pd.read_csv('/home/pedro/bkp/code/dataset/INFLUD-21-09-2020.csv',sep=';',encoding = "ISO-8859-1")
# Inputing constraint in the dataset
# Positive case:
df = df[df['PCR_SARS2']==1]
print(df.shape)
# Hospitalized people:
df = df[df['PCR_SARS2']==1][df['HOSPITAL']==1][df['NU_IDADE_N']<=110]
print(df.shape)
# Hospitalized people with age small than 110:
df = df[df['PCR_SARS2']==1][df['HOSPITAL']==1][df['NU_IDADE_N']<=110][df['EVOLUCAO'] != 3][df['EVOLUCAO'] != 9][df['EVOLUCAO'].notnull()]
print(df.shape)
# Latitudes and longitudes table from municipalities
df_cod = pd.read_csv('/home/pedro/bkp/code/dataset/municipios.csv', sep=',')
# Removing last number from "codenumber"
df_cod['CO_MUN_RES'] = df_cod['CO_MUN_RES'].astype(str).str[:-1].astype(np.int64)
df_cod['CO_MU_INTE'] = df_cod['CO_MU_INTE'].astype(str).str[:-1].astype(np.int64)
# To match catalogues using muninipacity code
# latitude and longitude for pacient residence
result_01 = pd.merge(df, df_cod[['CO_MUN_RES','latitude_res','longitude_res']], on='CO_MUN_RES', how="left")
result_02 = pd.merge(df, df_cod[['CO_MU_INTE','latitude_int','longitude_int']], on='CO_MU_INTE', how="left")
print(result_01.shape)
print(result_02.shape)
# To transforming in tuple
patient_mun_code = result_01[['latitude_res','longitude_res']].to_numpy()
hospital_mun_code = result_02[['latitude_int','longitude_int']].to_numpy()
print(patient_mun_code.shape)
print(hospital_mun_code.shape)
# To calculate the distance from patient to hospital (difference between municipalities centers in km)
df['distance'] = haversine_vector(patient_mun_code, hospital_mun_code, Unit.KILOMETERS)
# To check
print(df[['CO_MUN_RES','CO_MU_INTE','ID_MUNICIP', 'distance']])
# Histogram for Municipality Distance
plt.hist(df['distance'], 50, density=False, facecolor='g', alpha=0.75) # .dropna(),
plt.xlabel('km')
plt.ylabel('Distribution')
plt.title('Municipality Distance')
plt.show()
# To check long distances
print(df[['CO_MUN_RES','CO_MU_INTE','ID_MUNICIP', 'distance']][df['distance']>3000])
print(df['distance'].max())
print(df[['CO_MUN_RES','CO_MU_INTE','ID_MUNICIP', 'distance']][df['distance']>3479])
# Bigger distance Boa Vista (Roraima) to Joinville (Santa Catarina)
# overcrowded dataset
# dataset with code from hospital (cnes) with epidemiology week and the overcrowded status of hospital.
df_cod = pd.read_csv('/home/pedro/bkp/code/dataset/hospital_overcrowded.csv', sep=',')
# CO_UNI_NOT, SEM_NOT, Overcrowded
# Overload = number of hospitalization in epidemiological week for COVID-19 / 2019 sum hospital hospitalization by SARS
# To check
df = pd.merge(df, df_cod, on=['CO_UNI_NOT', 'SEM_NOT'], how="left")
print(df.shape)
# Municipalities number inicial
# patient municipality code number
print(len(df['CO_MUN_NOT']))
# reporting health unit code number
print(len(df['CO_UNI_NOT']))
print(df['CO_MUN_NOT'].nunique())
print(df['CO_MUN_RES'].nunique())
# IDHM
# Reading IBGE code for each municipalities and separating it for IDHM index
df_atlas = pd.read_excel (r'/home/pedro/bkp/code/dataset/AtlasBrasil_Consulta.xlsx')
# removind last interger in 'code' variable
df_atlas['code'] = df_atlas['code'].astype(str).str[:-1].astype(np.int64)
# Divinding IDHM in bins
IDHM_veryhigh = set(df_atlas['code'][df_atlas['IDHM2010']>=0.800])
print(len(IDHM_veryhigh))
IDHM_high = set(df_atlas['code'][((df_atlas['IDHM2010']>=0.700)&(df_atlas['IDHM2010']<0.800))])
print(len(IDHM_high))
IDHM_medium = set(df_atlas['code'][((df_atlas['IDHM2010']>=0.600)&(df_atlas['IDHM2010']<0.700))])
print(len(IDHM_medium))
IDHM_low = set(df_atlas['code'][((df_atlas['IDHM2010']>=0.500)&(df_atlas['IDHM2010']<0.600))])
print(len(IDHM_low))
IDHM_verylow = set(df_atlas['code'][df_atlas['IDHM2010']<0.500])
print(len(IDHM_verylow))
df.loc[df['CO_MUN_NOT'].isin(IDHM_veryhigh) == True, 'IDHM'] = 5
df.loc[df['CO_MUN_NOT'].isin(IDHM_high) == True, 'IDHM'] = 4
df.loc[df['CO_MUN_NOT'].isin(IDHM_medium) == True, 'IDHM'] = 3
df.loc[df['CO_MUN_NOT'].isin(IDHM_low) == True, 'IDHM'] = 2
df.loc[df['CO_MUN_NOT'].isin(IDHM_verylow) == True, 'IDHM'] = 1
# Private and public hospital separation
df_hospital = pd.read_csv('/home/pedro/bkp/code/dataset/CNES_SUS.txt', sep='\t')
public = set(df_hospital.iloc[:,0][df_hospital.iloc[:,3]=='S'])
private = set(df_hospital.iloc[:,0][df_hospital.iloc[:,3]=='N'])
df.loc[df['CO_UNI_NOT'].isin(public) == True, 'HEALTH_SYSTEM'] = 1
df.loc[df['CO_UNI_NOT'].isin(private) == True, 'HEALTH_SYSTEM'] = 0
# CO_UNI_NOT
# Constraint on dataset: We only analyze people with evolution, IDHM and Health system outcomes
df = df[df['IDHM'].notnull()][(df['HEALTH_SYSTEM']==1)|(df['HEALTH_SYSTEM']==0)]
print(df.shape)
# Municipalities number
print(len(df['CO_MUN_NOT']))
print(len(df['CO_MU_INTE']))
print(df['CO_MUN_NOT'].nunique())
print(df['CO_MU_INTE'].nunique())
# To selecting features and target
df = df[['Overload', 'distance','NU_IDADE_N','CS_SEXO','IDHM','CS_RACA','CS_ESCOL_N','SG_UF_NOT','CS_ZONA',\
'HEALTH_SYSTEM','CS_GESTANT','FEBRE','VOMITO','TOSSE','GARGANTA','DESC_RESP','DISPNEIA','DIARREIA',\
'SATURACAO','CARDIOPATI','HEPATICA','ASMA','PNEUMOPATI','RENAL','HEMATOLOGI','DIABETES',\
'OBESIDADE','NEUROLOGIC','IMUNODEPRE','EVOLUCAO']]
# adding comorbidities
df['SUM_COMORBIDITIES'] = df.iloc[:,19:-1].replace([9,2], 0).fillna(0).sum(axis=1)
df['SUM_SYMPTOMS'] = df.iloc[:,11:17].replace([9,2], 0).fillna(0).sum(axis=1)
# Pre-Processing
df = df[df['EVOLUCAO'].notnull()][df['EVOLUCAO']!=9][df['EVOLUCAO']!=3]#[df_BR['NU_IDADE_N'].notnull()]
df['CS_SEXO']=df['CS_SEXO'].replace({'M': 1, 'F':0, 'I':9, 'NaN':np.nan})
df['SG_UF_NOT'] = df['SG_UF_NOT'].map({'SP': 0, 'RJ':1, 'MG': 2 , 'ES':3, \
'RS':4, 'SC': 5, 'PR': 6, 'MT': 7, 'MS': 8, 'GO':9, 'DF':10, 'RO':11,'AC':12,'AM':13,\
'RR':14,'PA':15,'AP':16,'TO':17,'MA':18,'PI':19,'BA':20,'CE':21,'RN':22,'PB':23,'PE':24,'AL':25,'SE':26})
# Table 1
# Size of dataset
print(df.shape)
# Evolution
print(df['EVOLUCAO'].value_counts())
print(round(100*df['EVOLUCAO'].value_counts()/df['EVOLUCAO'].notnull().sum()),1)
print(df['EVOLUCAO'].isna().sum())
# Health System division
print(df['HEALTH_SYSTEM'].value_counts())
print(df['HEALTH_SYSTEM'].shape)
print(df['EVOLUCAO'][df['HEALTH_SYSTEM'] ==1].notnull().sum())
print(df['EVOLUCAO'][df['HEALTH_SYSTEM'] ==1].value_counts())
print(round(100*df['EVOLUCAO'][df['HEALTH_SYSTEM']==1].value_counts()/df['EVOLUCAO'][df['HEALTH_SYSTEM'] == 1].notnull().sum(),1))
# Private Health System (Hospital)
print(df['EVOLUCAO'][df['HEALTH_SYSTEM'] ==0].notnull().sum())
print(df['EVOLUCAO'][df['HEALTH_SYSTEM'] ==0].value_counts())
print(round(100*df['EVOLUCAO'][df['HEALTH_SYSTEM']==0].value_counts()/df['EVOLUCAO'][df['HEALTH_SYSTEM'] == 0].notnull().sum(),1))
# Mean age
#Cured people
print(df['NU_IDADE_N'][df['EVOLUCAO']==1].mean())
# Died people
print(df['NU_IDADE_N'][df['EVOLUCAO']==2].mean())
# For more information about table 1, take a look at code "table1.py"
#____________________________________________________________________
# Ordering features
df = df[['Overload', 'distance','NU_IDADE_N','CS_SEXO','IDHM','SUM_COMORBIDITIES','SUM_SYMPTOMS',\
'CS_RACA','CS_ESCOL_N','SG_UF_NOT','CS_ZONA','HEALTH_SYSTEM','CS_GESTANT','FEBRE','VOMITO','TOSSE',\
'GARGANTA','DESC_RESP','DISPNEIA','DIARREIA','SATURACAO','CARDIOPATI','HEPATICA','ASMA','PNEUMOPATI',\
'RENAL','HEMATOLOGI','DIABETES','OBESIDADE','NEUROLOGIC','IMUNODEPRE','EVOLUCAO']]
# Pre-Processing
# replacing 2 by 0 (Death patients)
df.iloc[:,13:] = df.iloc[:,13:].replace(to_replace = 2.0, value =0)
# For missing values in comorbidities and symptoms we filled by 0.
df.iloc[:,13:-1] = df.iloc[:,13:-1].fillna(0)
# feature
x = df.iloc[:,:-1]
# labels
y = df['EVOLUCAO']
# data separation
from sklearn.model_selection import train_test_split
x_train, x_test, y_train, y_test = \
train_test_split(x, y, test_size=0.2, random_state=5)
params = params = {'booster ':['gbtree'],
'n_estimators': [200],
'eta':[0.2],
'max_depth':[4],
'gamma': [1],
'subsample': [0.9],
'colsample_bytree': [0.9],
}
xgb = XGBClassifier()
gs = GridSearchCV(estimator=xgb, param_grid=params, return_train_score=True, cv=10, scoring='roc_auc')
gs = gs.fit(x_train, y_train)
print(gs.best_score_)
print(gs.best_params_)
print(gs.cv_results_['mean_test_score'])
print(gs.cv_results_['std_test_score'])
print(gs.cv_results_['mean_train_score'])
print(gs.cv_results_['std_train_score'])
clf = gs.best_estimator_
clf.fit(x_train, y_train)
# save the model to disk
filename_nn = 'xgb_with_symptoms.sav'
pickle.dump(clf, open(filename_nn, 'wb'))
# load the model from disk
loaded_model_xgb = pickle.load(open('xgb_with_symptoms.sav', 'rb'))
# To predicting
xgb_pred = loaded_model_xgb.predict_proba(x_test)
def auc_CI(df_test):
x_test, y_test = df_test[:, :-1], df_test[:,[-1]]
pred = loaded_model_xgb.predict_proba(x_test)
fpr, tpr, thresholds = roc_curve(y_test, pred[:,1], pos_label=1)
return auc(x=fpr, y=tpr)
df_test = pd.merge(x_test, y_test, left_index=True, right_index=True)
# Febre
original, std_err, ci_bounds = bootstrap(df_test[df['FEBRE']==1].values , num_rounds=100, func = auc_CI, ci=0.95, seed=123)
print('Mean Febre: %.3f, SE: +/- %.3f, CI95: [%.3f, %.3f]' % (original, std_err, ci_bounds[0], ci_bounds[1]))
print('Febre number of data %d' % df_test[df_test['FEBRE']==1].shape[0])
# Vomito
original, std_err, ci_bounds = bootstrap(df_test[df['VOMITO']==1].values , num_rounds=100, func = auc_CI, ci=0.95, seed=123)
print('Mean Vomito: %.3f, SE: +/- %.3f, CI95: [%.3f, %.3f]' % (original, std_err, ci_bounds[0], ci_bounds[1]))
print('Vomito number of data %d' % df_test[df_test['VOMITO']==1].shape[0])
# Vomito
original, std_err, ci_bounds = bootstrap(df_test[df['TOSSE']==1].values , num_rounds=100, func = auc_CI, ci=0.95, seed=123)
print('Mean TOSSE: %.3f, SE: +/- %.3f, CI95: [%.3f, %.3f]' % (original, std_err, ci_bounds[0], ci_bounds[1]))
print('TOSSE number of data %d' % df_test[df_test['TOSSE']==1].shape[0])
# GARGANTA
original, std_err, ci_bounds = bootstrap(df_test[df['GARGANTA']==1].values , num_rounds=100, func = auc_CI, ci=0.95, seed=123)
print('Mean GARGANTA: %.3f, SE: +/- %.3f, CI95: [%.3f, %.3f]' % (original, std_err, ci_bounds[0], ci_bounds[1]))
print('GARGANTA number of data %d' % df_test[df_test['GARGANTA']==1].shape[0])
# DESC_RESP
original, std_err, ci_bounds = bootstrap(df_test[df['DESC_RESP']==1].values , num_rounds=100, func = auc_CI, ci=0.95, seed=123)
print('Mean DESC_RESP: %.3f, SE: +/- %.3f, CI95: [%.3f, %.3f]' % (original, std_err, ci_bounds[0], ci_bounds[1]))
print('DESC_RESP number of data %d' % df_test[df_test['DESC_RESP']==1].shape[0])
# DISPNEIA
original, std_err, ci_bounds = bootstrap(df_test[df['DISPNEIA']==1].values , num_rounds=100, func = auc_CI, ci=0.95, seed=123)
print('Mean DISPNEIA: %.3f, SE: +/- %.3f, CI95: [%.3f, %.3f]' % (original, std_err, ci_bounds[0], ci_bounds[1]))
print('DISPNEIA number of data %d' % df_test[df_test['DISPNEIA']==1].shape[0])
# DIARREIA
original, std_err, ci_bounds = bootstrap(df_test[df['DIARREIA']==1].values , num_rounds=100, func = auc_CI, ci=0.95, seed=123)
print('Mean DIARREIA: %.3f, SE: +/- %.3f, CI95: [%.3f, %.3f]' % (original, std_err, ci_bounds[0], ci_bounds[1]))
print('DIARREIA number of data %d' % df_test[df_test['DIARREIA']==1].shape[0])
# SATURACAO
original, std_err, ci_bounds = bootstrap(df_test[df['SATURACAO']==1].values , num_rounds=100, func = auc_CI, ci=0.95, seed=123)
print('Mean SATURACAO: %.3f, SE: +/- %.3f, CI95: [%.3f, %.3f]' % (original, std_err, ci_bounds[0], ci_bounds[1]))
print('SATURACAO number of data %d' % df_test[df_test['SATURACAO']==1].shape[0])
# CARDIOPATI
original, std_err, ci_bounds = bootstrap(df_test[df['CARDIOPATI']==1].values , num_rounds=100, func = auc_CI, ci=0.95, seed=123)
print('Mean CARDIOPATI: %.3f, SE: +/- %.3f, CI95: [%.3f, %.3f]' % (original, std_err, ci_bounds[0], ci_bounds[1]))
print('CARDIOPATI number of data %d' % df_test[df_test['CARDIOPATI']==1].shape[0])
# 'HEPATICA'
original, std_err, ci_bounds = bootstrap(df_test[df['HEPATICA']==1].values , num_rounds=100, func = auc_CI, ci=0.95, seed=123)
print('Mean HEPATICA: %.3f, SE: +/- %.3f, CI95: [%.3f, %.3f]' % (original, std_err, ci_bounds[0], ci_bounds[1]))
print('HEPATICA number of data %d' % df_test[df_test['HEPATICA']==1].shape[0])
# ASMA
original, std_err, ci_bounds = bootstrap(df_test[df['ASMA']==1].values , num_rounds=100, func = auc_CI, ci=0.95, seed=123)
print('Mean ASMA: %.3f, SE: +/- %.3f, CI95: [%.3f, %.3f]' % (original, std_err, ci_bounds[0], ci_bounds[1]))
print('ASMA number of data %d' % df_test[df_test['ASMA']==1].shape[0])
# PNEUMOPATI
original, std_err, ci_bounds = bootstrap(df_test[df['PNEUMOPATI']==1].values , num_rounds=100, func = auc_CI, ci=0.95, seed=123)
print('Mean PNEUMOPATI: %.3f, SE: +/- %.3f, CI95: [%.3f, %.3f]' % (original, std_err, ci_bounds[0], ci_bounds[1]))
print('PNEUMOPATI number of data %d' % df_test[df_test['PNEUMOPATI']==1].shape[0])
# RENAL
original, std_err, ci_bounds = bootstrap(df_test[df['RENAL']==1].values , num_rounds=100, func = auc_CI, ci=0.95, seed=123)
print('Mean RENAL: %.3f, SE: +/- %.3f, CI95: [%.3f, %.3f]' % (original, std_err, ci_bounds[0], ci_bounds[1]))
print('RENAL number of data %d' % df_test[df_test['RENAL']==1].shape[0])
# HEMATOLOGI
original, std_err, ci_bounds = bootstrap(df_test[df['HEMATOLOGI']==1].values , num_rounds=100, func = auc_CI, ci=0.95, seed=123)
print('Mean HEMATOLOGI: %.3f, SE: +/- %.3f, CI95: [%.3f, %.3f]' % (original, std_err, ci_bounds[0], ci_bounds[1]))
print('HEMATOLOGI number of data %d' % df_test[df_test['HEMATOLOGI']==1].shape[0])
# DIABETES
original, std_err, ci_bounds = bootstrap(df_test[df['DIABETES']==1].values , num_rounds=100, func = auc_CI, ci=0.95, seed=123)
print('Mean DIABETES: %.3f, SE: +/- %.3f, CI95: [%.3f, %.3f]' % (original, std_err, ci_bounds[0], ci_bounds[1]))
print('DIABETES number of data %d' % df_test[df_test['DIABETES']==1].shape[0])
# OBESIDADE
original, std_err, ci_bounds = bootstrap(df_test[df['OBESIDADE']==1].values , num_rounds=100, func = auc_CI, ci=0.95, seed=123)
print('Mean OBESIDADE: %.3f, SE: +/- %.3f, CI95: [%.3f, %.3f]' % (original, std_err, ci_bounds[0], ci_bounds[1]))
print('OBESIDADE number of data %d' % df_test[df_test['OBESIDADE']==1].shape[0])
# NEUROLOGIC
original, std_err, ci_bounds = bootstrap(df_test[df['NEUROLOGIC']==1].values , num_rounds=100, func = auc_CI, ci=0.95, seed=123)
print('Mean NEUROLOGIC: %.3f, SE: +/- %.3f, CI95: [%.3f, %.3f]' % (original, std_err, ci_bounds[0], ci_bounds[1]))
print('NEUROLOGIC number of data %d' % df_test[df_test['NEUROLOGIC']==1].shape[0])
# IMUNODEPRE
original, std_err, ci_bounds = bootstrap(df_test[df['IMUNODEPRE']==1].values , num_rounds=100, func = auc_CI, ci=0.95, seed=123)
print('Mean IMUNODEPRE: %.3f, SE: +/- %.3f, CI95: [%.3f, %.3f]' % (original, std_err, ci_bounds[0], ci_bounds[1]))
print('IMUNODEPRE number of data %d' % df_test[df_test['IMUNODEPRE']==1].shape[0])
# Female
original, std_err, ci_bounds = bootstrap(df_test[df['CS_SEXO']==0].values , num_rounds=100, func = auc_CI, ci=0.95, seed=123)
print('Mean Female: %.3f, SE: +/- %.3f, CI95: [%.3f, %.3f]' % (original, std_err, ci_bounds[0], ci_bounds[1]))
print('Female number of data %d' % df_test[df_test['CS_SEXO']==0].shape[0])
# Male
original, std_err, ci_bounds = bootstrap(df_test[df['CS_SEXO']==1].values , num_rounds=100, func = auc_CI, ci=0.95, seed=123)
print('Mean Male: %.3f, SE: +/- %.3f, CI95: [%.3f, %.3f]' % (original, std_err, ci_bounds[0], ci_bounds[1]))
print('Male number of data %d' % df_test[df_test['CS_SEXO']==1].shape[0])
# Hospital unit reporting
# public
original, std_err, ci_bounds = bootstrap(df_test[df['HEALTH_SYSTEM']==1].values , num_rounds=100, func = auc_CI, ci=0.95, seed=123)
print('Mean public: %.3f, SE: +/- %.3f, CI95: [%.3f, %.3f]' % (original, std_err, ci_bounds[0], ci_bounds[1]))
print('Private number of data %d' % df_test[df_test['HEALTH_SYSTEM']==1].shape[0])
# private
original, std_err, ci_bounds = bootstrap(df_test[df['HEALTH_SYSTEM']==0].values , num_rounds=100, func = auc_CI, ci=0.95, seed=123)
print('Mean private: %.3f, SE: +/- %.3f, CI95: [%.3f, %.3f]' % (original, std_err, ci_bounds[0], ci_bounds[1]))
print('Private number of data %d' % df_test[df_test['HEALTH_SYSTEM']==0].shape[0])
# Hospital unit reporting
# Branco
original, std_err, ci_bounds = bootstrap(df_test[df['CS_RACA']==1].values , num_rounds=100, func = auc_CI, ci=0.95, seed=123)
print('Mean Branco: %.3f, SE: +/- %.3f, CI95: [%.3f, %.3f]' % (original, std_err, ci_bounds[0], ci_bounds[1]))
print('Branco number of data %d' % df_test[df_test['CS_RACA']==1].shape[0])
# Preto
original, std_err, ci_bounds = bootstrap(df_test[df['CS_RACA']==2].values , num_rounds=100, func = auc_CI, ci=0.95, seed=123)
print('Mean Preto: %.3f, SE: +/- %.3f, CI95: [%.3f, %.3f]' % (original, std_err, ci_bounds[0], ci_bounds[1]))
print('Preto number of data %d' % df_test[df_test['CS_RACA']==2].shape[0])
# Amarelo
original, std_err, ci_bounds = bootstrap(df_test[df['CS_RACA']==3].values , num_rounds=100, func = auc_CI, ci=0.95, seed=123)
print('Mean Amarelo: %.3f, SE: +/- %.3f, CI95: [%.3f, %.3f]' % (original, std_err, ci_bounds[0], ci_bounds[1]))
print('Amarelo number of data %d' % df_test[df_test['CS_RACA']==3].shape[0])
# Pardo
original, std_err, ci_bounds = bootstrap(df_test[df['CS_RACA']==4].values , num_rounds=100, func = auc_CI, ci=0.95, seed=123)
print('Mean Pardo: %.3f, SE: +/- %.3f, CI95: [%.3f, %.3f]' % (original, std_err, ci_bounds[0], ci_bounds[1]))
print('Pardo number of data %d' % df_test[df_test['CS_RACA']==4].shape[0])
# Indigena
original, std_err, ci_bounds = bootstrap(df_test[df['CS_RACA']==5].values , num_rounds=100, func = auc_CI, ci=0.95, seed=123)
print('Mean Indigena: %.3f, SE: +/- %.3f, CI95: [%.3f, %.3f]' % (original, std_err, ci_bounds[0], ci_bounds[1]))
print('Indigena number of data %d' % df_test[df_test['CS_RACA']==5].shape[0])
# Education
# Illiterate
original, std_err, ci_bounds = bootstrap(df_test[df['CS_ESCOL_N']==0].values , num_rounds=100, func = auc_CI, ci=0.95, seed=123)
print('Mean Illiterate: %.3f, SE: +/- %.3f, CI95: [%.3f, %.3f]' % (original, std_err, ci_bounds[0], ci_bounds[1]))
print('Illiterate number of data %d' % df_test[df_test['CS_ESCOL_N']==0].shape[0])
# Fundamental I
original, std_err, ci_bounds = bootstrap(df_test[df['CS_ESCOL_N']==1].values , num_rounds=100, func = auc_CI, ci=0.95, seed=123)
print('Mean Fundamental I: %.3f, SE: +/- %.3f, CI95: [%.3f, %.3f]' % (original, std_err, ci_bounds[0], ci_bounds[1]))
print('Fundamental I number of data %d' % df_test[df_test['CS_ESCOL_N']==1].shape[0])
# Fundamental II
original, std_err, ci_bounds = bootstrap(df_test[df['CS_ESCOL_N']==2].values , num_rounds=100, func = auc_CI, ci=0.95, seed=123)
print('Mean Fundamental II: %.3f, SE: +/- %.3f, CI95: [%.3f, %.3f]' % (original, std_err, ci_bounds[0], ci_bounds[1]))
print('Fundamental II number of data %d' % df_test[df_test['CS_ESCOL_N']==2].shape[0])
# Medium
original, std_err, ci_bounds = bootstrap(df_test[df['CS_ESCOL_N']==3].values , num_rounds=100, func = auc_CI, ci=0.95, seed=123)
print('Mean Medium: %.3f, SE: +/- %.3f, CI95: [%.3f, %.3f]' % (original, std_err, ci_bounds[0], ci_bounds[1]))
print('Medium number of data %d' % df_test[df_test['CS_ESCOL_N']==3].shape[0])
# University
original, std_err, ci_bounds = bootstrap(df_test[df['CS_ESCOL_N']==4].values , num_rounds=100, func = auc_CI, ci=0.95, seed=123)
print('Mean University: %.3f, SE: +/- %.3f, CI95: [%.3f, %.3f]' % (original, std_err, ci_bounds[0], ci_bounds[1]))
print('University number of data %d' % df_test[df_test['CS_ESCOL_N']==4].shape[0])
# Area
# Urban
original, std_err, ci_bounds = bootstrap(df_test[df['CS_ZONA']==1].values , num_rounds=100, func = auc_CI, ci=0.95, seed=123)
print('Mean Urban: %.3f, SE: +/- %.3f, CI95: [%.3f, %.3f]' % (original, std_err, ci_bounds[0], ci_bounds[1]))
print('Urban number of data %d' % df_test[df_test['CS_ZONA']==1].shape[0])
# Rural
original, std_err, ci_bounds = bootstrap(df_test[df['CS_ZONA']==2].values , num_rounds=100, func = auc_CI, ci=0.95, seed=123)
print('Mean Rural: %.3f, SE: +/- %.3f, CI95: [%.3f, %.3f]' % (original, std_err, ci_bounds[0], ci_bounds[1]))
print('Rural number of data %d' % df_test[df_test['CS_ZONA']==2].shape[0])
# Peri-urban
original, std_err, ci_bounds = bootstrap(df_test[df['CS_ZONA']==3].values , num_rounds=100, func = auc_CI, ci=0.95, seed=123)
print('Peri-urban University: %.3f, SE: +/- %.3f, CI95: [%.3f, %.3f]' % (original, std_err, ci_bounds[0], ci_bounds[1]))
print('Peri-urban number of data %d' % df_test[df_test['CS_ZONA']==3].shape[0])
# IDHM
# Very High
original, std_err, ci_bounds = bootstrap(df_test[df['IDHM']==5].values , num_rounds=100, func = auc_CI, ci=0.95, seed=123)
print('Mean Very High: %.3f, SE: +/- %.3f, CI95: [%.3f, %.3f]' % (original, std_err, ci_bounds[0], ci_bounds[1]))
print('Very High number of data %d' % df_test[df_test['IDHM']==5].shape[0])
# High
original, std_err, ci_bounds = bootstrap(df_test[df['IDHM']==4].values , num_rounds=100, func = auc_CI, ci=0.95, seed=123)
print('Mean High: %.3f, SE: +/- %.3f, CI95: [%.3f, %.3f]' % (original, std_err, ci_bounds[0], ci_bounds[1]))
print('High number of data %d' % df_test[df_test['IDHM']==4].shape[0])
# Medium Low
original, std_err, ci_bounds = bootstrap(df_test[((df['IDHM']==3)|(df['IDHM']==2)|(df['IDHM']==1))].values , num_rounds=100, func = auc_CI, ci=0.95, seed=123)
print('Mean Medium Low: %.3f, SE: +/- %.3f, CI95: [%.3f, %.3f]' % (original, std_err, ci_bounds[0], ci_bounds[1]))
print('Medium Low number of data %d' % df_test[((df['IDHM']==3)|(df['IDHM']==2)|(df['IDHM']==1))].shape[0])
# Macroregions
# Macro Regions
northeast = set([18,19,20,21,22,23,24,25,26])
north = set([11,12,13,14,15,16,17])
midwest = set([7,8,9,10])
southeast = set([0,1,2,3])
south = set([4,5,6])
# North
original, std_err, ci_bounds = bootstrap(df_test[df['SG_UF_NOT'].isin(north)].values , num_rounds=100, func = auc_CI, ci=0.95, seed=123)
print('Mean North: %.3f, SE: +/- %.3f, CI95: [%.3f, %.3f]' % (original, std_err, ci_bounds[0], ci_bounds[1]))
print('North number of data %d' % df_test[df_test['SG_UF_NOT'].isin(north)].shape[0])
# Northeast
original, std_err, ci_bounds = bootstrap(df_test[df['SG_UF_NOT'].isin(northeast)].values , num_rounds=100, func = auc_CI, ci=0.95, seed=123)
print('Mean Northeast: %.3f, SE: +/- %.3f, CI95: [%.3f, %.3f]' % (original, std_err, ci_bounds[0], ci_bounds[1]))
print('Northeast number of data %d' % df_test[df_test['SG_UF_NOT'].isin(northeast)].shape[0])
# Midwest
original, std_err, ci_bounds = bootstrap(df_test[df['SG_UF_NOT'].isin(midwest)].values , num_rounds=100, func = auc_CI, ci=0.95, seed=123)
print('Mean Midwest: %.3f, SE: +/- %.3f, CI95: [%.3f, %.3f]' % (original, std_err, ci_bounds[0], ci_bounds[1]))
print('Midwest number of data %d' % df_test[df_test['SG_UF_NOT'].isin(midwest)].shape[0])
# Southeast
original, std_err, ci_bounds = bootstrap(df_test[df['SG_UF_NOT'].isin(southeast)].values , num_rounds=100, func = auc_CI, ci=0.95, seed=123)
print('Mean Southeast: %.3f, SE: +/- %.3f, CI95: [%.3f, %.3f]' % (original, std_err, ci_bounds[0], ci_bounds[1]))
print('Southeast number of data %d' % df_test[df_test['SG_UF_NOT'].isin(southeast)].shape[0])
# South
original, std_err, ci_bounds = bootstrap(df_test[df['SG_UF_NOT'].isin(south)].values , num_rounds=100, func = auc_CI, ci=0.95, seed=123)
print('Mean South: %.3f, SE: +/- %.3f, CI95: [%.3f, %.3f]' % (original, std_err, ci_bounds[0], ci_bounds[1]))
print('South number of data %d' % df_test[df_test['SG_UF_NOT'].isin(south)].shape[0])
# CI 95% calculation AUC
def auc_CI(df_test):
x_test, y_test = df_test[:, :-1], df_test[:,[-1]]
xgb_pred = loaded_model_xgb.predict_proba(x_test)
fpr, tpr, thresholds = roc_curve(y_test, xgb_pred[:,1], pos_label=1)
return auc(x=fpr, y=tpr)
df_test = pd.merge(x_test, y_test, left_index=True, right_index=True)
original, std_err, ci_bounds = bootstrap(df_test.values , num_rounds=100, func = auc_CI, ci=0.95, seed=123)
print('Mean: %.3f, SE: +/- %.3f, CI95: [%.3f, %.3f]' % (original, std_err, ci_bounds[0], ci_bounds[1]))
# CI 95% calculation AP Death
def ap_CI_death(df_test):
x_test, y_test = df_test[:, :-1], df_test[:,[-1]]
xgb_pred = loaded_model_xgb.predict_proba(x_test)
return average_precision_score(y_test, xgb_pred[:,0], pos_label=0)
df_test = pd.merge(x_test, y_test, left_index=True, right_index=True)
original, std_err, ci_bounds = bootstrap(df_test.values , num_rounds=100, func = ap_CI_death, ci=0.95, seed=123)
print('Mean Death: %.3f, SE: +/- %.3f, CI95: [%.3f, %.3f]' % (original, std_err, ci_bounds[0], ci_bounds[1]))
# CI 95% calculation AP Cure
def ap_CI_cure(df_test):
x_test, y_test = df_test[:, :-1], df_test[:,[-1]]
xgb_pred = loaded_model_xgb.predict_proba(x_test)
return average_precision_score(y_test, xgb_pred[:,1])
df_test = pd.merge(x_test, y_test, left_index=True, right_index=True)
original, std_err, ci_bounds = bootstrap(df_test.values , num_rounds=100, func = ap_CI_cure, ci=0.95, seed=123)
print('Mean Cure: %.3f, SE: +/- %.3f, CI95: [%.3f, %.3f]' % (original, std_err, ci_bounds[0], ci_bounds[1]))
```
| github_jupyter |
### Agenda
1. Installation
2. Basics
3. Iterables
4. Numpy (for math and matrix operations)
5. Matplotlib (for plotting)
6. Q&A
```
# Note: This tutorial is based on Python 3.8
# but it should apply to all Python 3.X versions
# Please note that this tutorial is NOT exhaustive
# We try to cover everything you need for class assignments
# but you should also navigate external resources
#
# More tutorials:
# NUMPY:
# https://cs231n.github.io/python-numpy-tutorial/#numpy
# https://numpy.org/doc/stable/user/quickstart.html
# MATPLOTLIB:
# https://matplotlib.org/gallery/index.html
# BASICS:
# https://www.w3schools.com/python/
# CONSULT THESE WISELY:
# The official documentation, Google, and Stack-overflow are your friends!
```
### 1. Installation
#### Anaconda for environment management
https://www.anaconda.com/
common commands
conda env list <-- list all environments
conda create -n newenv python=3.8 <-- create new environment
conda enc create -f env.yml <-- create environment from config file
conda activate envname <-- activate a environment
conda deactivate <-- exit environment
pip install packagename <-- install package for current environment
jupyter notebook <-- open jupyter in current environment
#### Package installation using conda/pip
Live demo
#### Recommended IDEs
Spyder (in-built in Anaconda) <br/>
Pycharm (the most popular choice, compatible with Anaconda)
```
# common anaconda commands
#conda env list
#conda create -n name python=3.8
#conda env create -f env.yml
#conda activate python2.7
#conda deactivate
#install packages
#pip install <package>
```
### 2. Basics
https://www.w3schools.com/python/
```
# input and output
name = input()
print("hello, " + name)
# print multiple variables separated by a space
print("hello", name, 1, 3.0, True)
# line comment
"""
block
comments
"""
# variables don't need explicit declaration
var = "hello" # string
var = 10.0 # float
var = 10 # int
var = True # boolean
var = [1,2,3] # pointer to list
var = None # empty pointer
# type conversions
var = 10
print(int(var))
print(str(var))
print(float(var))
# basic math operations
var = 10
print("var + 4 =", 10 + 4)
print("var - 4 =", 10 - 4)
print("var * 4 =", 10 * 4)
print("var ^ 4=", 10 ** 4)
print("int(var) / 4 =", 10//4) # // for int division
print("float(var) / 4 =", 10/4) # / for float division
# All compound assignment operators available
# including += -= *= **= /= //=
# pre/post in/decrementers not available (++ --)
# basic boolean operations include "and", "or", "not"
print("not True is", not True)
print("True and False is", True and False)
print("True or False is", True or False)
# String operations
# '' and "" are equivalent
s = "String"
#s = 'Mary said "Hello" to John'
#s = "Mary said \"Hello\" to John"
# basic
print(len(s)) # get length of string and any iterable type
print(s[0]) # get char by index
print(s[1:3]) # [1,3)
print("This is a " + s + "!")
# handy tools
print(s.lower())
print(s*4)
print("ring" in s)
print(s.index("ring"))
# slice by delimiter
print("I am a sentence".split(" "))
# concatenate a list of string using a delimiter
print("...".join(['a','b','c']))
# formatting variables
print("Formatting a string like %.2f"%(0.12345))
print(f"Or like {s}!")
# control flows
# NOTE: No parentheses or curly braces
# Indentation is used to identify code blocks
# So never ever mix spaces with tabs
for i in range(0,5):
for j in range(i, 5):
print("inner loop")
print("outer loop")
# if-else
var = 10
if var > 10:
print(">")
elif var == 10:
print("=")
else:
print("<")
# use "if" to check null pointer or empty arrays
var = None
if var:
print(var)
var = []
if var:
print(var)
var = "object"
if var:
print(var)
# while-loop
var = 5
while var > 0:
print(var)
var -=1
# for-loop
for i in range(3): # prints 0 1 2
print(i)
"""
equivalent to
for (int i = 0; i < 3; i++)
"""
print("-------")
# range (start-inclusive, stop-exclusive, step)
for i in range(2, -3, -2):
print(i)
"""
equivalent to
for (int i = 2; i > -3; i-=2)
"""
# define function
def func(a, b):
return a + b
func(1,3)
# use default parameters and pass values by parameter name
def rangeCheck(a, min_val = 0, max_val=10):
return min_val < a < max_val # syntactic sugar
rangeCheck(5, max_val=4)
# define class
class Foo:
# optinal constructor
def __init__(self, x):
# first parameter "self" for instance reference, like "this" in JAVA
self.x = x
# instance method
def printX(self): # instance reference is required for all function parameters
print(self.x)
# class methods, most likely you will never need this
@classmethod
def printHello(self):
print("hello")
obj = Foo(6)
obj.printX()
# class inheritance - inherits variables and methods
# You might need this when you learn more PyTorch
class Bar(Foo):
pass
obj = Bar(3)
obj.printX()
```
### 3. Iterables
```
alist = list() # linear, size not fixed, not hashable
atuple = tuple() # linear, fixed size, hashable
adict = dict() # hash table, not hashable, stores (key,value) pairs
aset = set() # hash table, like dict but only stores keys
acopy = alist.copy() # shallow copy
print(len(alist)) # gets size of any iterable type
# examplar tuple usage
# creating a dictionary to store ngram counts
d = dict()
d[("a","cat")] = 10
d[["a","cat"]] = 11
"""
List: not hashable (i.e. can't use as dictionary key)
dynamic size
allows duplicates and inconsistent element types
dynamic array implementation
"""
# list creation
alist = [] # empty list, equivalent to list()
alist = [1,2,3,4,5] # initialized list
print(alist[0])
alist[0] = 5
print(alist)
print("-"*10)
# list indexing
print(alist[0]) # get first element (at index 0)
print(alist[-2]) # get last element (at index len-1)
print(alist[3:]) # get elements starting from index 3 (inclusive)
print(alist[:3]) # get elements stopping at index 3 (exclusive)
print(alist[2:4]) # get elements within index range [2,4)
print(alist[6:]) # prints nothing because index is out of range
print(alist[::-1]) # returns a reversed list
print("-"*10)
# list modification
alist.append("new item") # insert at end
alist.insert(0, "new item") # insert at index 0
alist.extend([2,3,4]) # concatenate lists
# above line is equivalent to alist += [2,3,4]
alist.index("new item") # search by content
alist.remove("new item") # remove by content
alist.pop(0) # remove by index
print(alist)
print("-"*10)
if "new item" in alist:
print("found")
else:
print("not found")
print("-"*10)
# list traversal
for ele in alist:
print(ele)
print("-"*10)
# or traverse with index
for i, ele in enumerate(alist):
print(i, ele)
"""
Tuple: hashable (i.e. can use as dictionary key)
fixed size (no insertion or deletion)
"""
# it does not make sense to create empty tuples
atuple = (1,2,3,4,5)
# or you can cast other iterables to tuple
atuple = tuple([1,2,3])
# indexing and traversal are same as list
"""
Named tuples for readibility
"""
from collections import namedtuple
Point = namedtuple('Point', 'x y')
pt1 = Point(1.0, 5.0)
pt2 = Point(2.5, 1.5)
print(pt1.x, pt1.y)
"""
Dict: not hashable
dynamic size
no duplicates allowed
hash table implementation which is fast for searching
"""
# dict creation
adict = {} # empty dict, equivalent to dict()
adict = {'a':1, 'b':2, 'c':3}
print(adict)
# get all keys in dictionary
print(adict.keys())
# get value paired with key
print(adict['a'])
key = 'e'
# NOTE: accessing keys not in the dictionary leads to exception
if key in adict:
print(adict[key])
# add or modify dictionary entries
adict['e'] = 10 # insert new key
adict['e'] = 5 # modify existing keys
print("-"*10)
# traverse keys only
for key in adict:
print(key, adict[key])
print("-"*10)
# or traverse key-value pairs together
for key, value in adict.items():
print(key, value)
print("-"*10)
# NOTE: Checking if a key exists
key = 'e'
if key in adict: # NO .keys() here please!
print(adict[key])
else:
print("Not found!")
"""
Special dictionaries
"""
# set is a dictionary without values
aset = set()
aset.add('a')
# deduplication short-cut using set
alist = [1,2,3,3,3,4,3]
alist = list(set(alist))
print(alist)
# default_dictionary returns a value computed from a default function
# for non-existent entries
from collections import defaultdict
adict = defaultdict(lambda: 'unknown')
adict['cat'] = 'feline'
print(adict['cat'])
print(adict['dog'])
# counter is a dictionary with default value of 0
# and provides handy iterable counting tools
from collections import Counter
# initialize and modify empty counter
counter1 = Counter()
counter1['t'] = 10
counter1['t'] += 1
counter1['e'] += 1
print(counter1)
print("-"*10)
# initialize counter from iterable
counter2 = Counter("letters to be counted")
print(counter2)
print("-"*10)
# computations using counters
print("1", counter1 + counter2)
print("2,", counter1 - counter2)
print("3", counter1 or counter2) # or for intersection, and for union
# sorting
a = [4,6,1,7,0,5,1,8,9]
a = sorted(a)
print(a)
a = sorted(a, reverse=True)
print(a)
# sorting
a = [("cat",1), ("dog", 3), ("bird", 2)]
a = sorted(a)
print(a)
a = sorted(a, key=lambda x:x[1])
print(a)
# useful in dictionary sorting
adict = {'cat':3, 'bird':1}
print(sorted(adict.items(), key=lambda x:x[1]))
# Syntax sugar: one-line control flow + list operation
sent = ["i am good", "a beautiful day", "HELLO FRIEND"]
"""
for i in range(len(sent)):
sent[i] = sent[i].lower().split(" ")
"""
sent1 = [s.lower().split(" ") for s in sent]
print(sent1)
sent2 = [s.lower().split(" ") for s in sent if len(s) > 10]
print(sent2)
# Use this for deep copy!
# copy = [obj.copy() for obj in original]
# Syntax sugar: * operator for repeating iterable elements
print("-"*10)
print([1]*10)
# Note: This only repeating by value
# So you cannot apply the trick on reference types
# To create a double list
# DONT
doublelist = [[]]*10
doublelist[0].append(1)
print(doublelist)
# DO
doublelist = [[] for _ in range(10)]
doublelist[0].append(1)
print(doublelist)
```
### 4. Numpy
Very powerful python tool for handling matrices and higher dimensional arrays
```
import numpy as np
# create arrays
a = np.array([[1,2],[3,4],[5,6]])
print(a)
print(a.shape)
# create all-zero/one arrays
b = np.ones((3,4)) # np.zeros((3,4))
print(b)
print(b.shape)
# create identity matrix
c = np.eye(5)
print(c)
print(c.shape)
# reshaping arrays
a = np.arange(8) # [8,] similar range() you use in for-loops
b = a.reshape((4,2)) # shape [4,2]
c = a.reshape((2,2,-1)) # shape [2,2,2] -- -1 for auto-fill
d = c.flatten() # shape [8,]
e = np.expand_dims(a, 0) # [1,8]
f = np.expand_dims(a, 1) # [8,1]
g = e.squeeze() # shape[8, ] -- remove all unnecessary dimensions
print(a)
print(b)
# concatenating arrays
a = np.ones((4,3))
b = np.ones((4,3))
c = np.concatenate([a,b], 0)
print(c.shape)
d = np.concatenate([a,b], 1)
print(d.shape)
# one application is to create a batch for NN
x1 = np.ones((32,32,3))
x2 = np.ones((32,32,3))
x3 = np.ones((32,32,3))
# --> to create a batch of shape (3,32,32,3)
x = [x1, x2, x3]
x = [np.expand_dims(xx, 0) for xx in x] # xx shape becomes (1,32,32,3)
x = np.concatenate(x, 0)
print(x.shape)
# access array slices by index
a = np.zeros([10, 10])
a[:3] = 1
a[:, :3] = 2
a[:3, :3] = 3
rows = [4,6,7]
cols = [9,3,5]
a[rows, cols] = 4
print(a)
# transposition
a = np.arange(24).reshape(2,3,4)
print(a.shape)
print(a)
a = np.transpose(a, (2,1,0)) # swap 0th and 2nd axes
print(a.shape)
print(a)
c = np.array([[1,2],[3,4]])
# pinv is pseudo inversion for stability
print(np.linalg.pinv(c))
# l2 norm by default, read documentation for more options
print(np.linalg.norm(c))
# summing a matrix
print(np.sum(c))
# the optional axis parameter
print(c)
print(np.sum(c, axis=0)) # sum along axis 0
print(np.sum(c, axis=1)) # sum along axis 1
# dot product
c = np.array([1,2])
d = np.array([3,4])
print(np.dot(c,d))
# matrix multiplication
a = np.ones((4,3)) # 4,3
b = np.ones((3,2)) # 3,2 --> 4,2
print(a @ b) # same as a.dot(b)
c = a @ b # (4,2)
# automatic repetition along axis
d = np.array([1,2,3,4]).reshape(4,1)
print(c + d)
# handy for batch operation
batch = np.ones((3,32))
weight = np.ones((32,10))
bias = np.ones((1,10))
print((batch @ weight + bias).shape)
# speed test: numpy vs list
a = np.ones((100,100))
b = np.ones((100,100))
def matrix_multiplication(X, Y):
result = [[0]*len(Y[0]) for _ in range(len(X))]
for i in range(len(X)):
for j in range(len(Y[0])):
for k in range(len(Y)):
result[i][j] += X[i][k] * Y[k][j]
return result
import time
# run numpy matrix multiplication for 10 times
start = time.time()
for _ in range(10):
a @ b
end = time.time()
print("numpy spends {} seconds".format(end-start))
# run list matrix multiplication for 10 times
start = time.time()
for _ in range(10):
matrix_multiplication(a,b)
end = time.time()
print("list operation spends {} seconds".format(end-start))
# the difference gets more significant as matrices grow in size!
# element-wise operations, for examples
np.log(a)
np.exp(a)
np.sin(a)
# operation with scalar is interpreted as element-wise
a * 3
```
### 5. Matplotlib
Powerful tool for visualization <br/>
Many tutorials online. We only go over the basics here <br/>
```
import matplotlib.pyplot as plt
# line plot
x = [1,2,3]
y = [1,3,2]
plt.plot(x,y)
# scatter plot
plt.scatter(x,y)
# bar plots
plt.bar(x,y)
# plot configurations
x = [1,2,3]
y1 = [1,3,2]
y2 = [4,0,4]
# set figure size
plt.figure(figsize=(5,5))
# set axes
plt.xlim(0,5)
plt.ylim(0,5)
plt.xlabel("x label")
plt.ylabel("y label")
# add title
plt.title("My Plot")
plt.plot(x,y1, label="data1", color="red", marker="*")
plt.plot(x,y2, label="data2", color="green", marker=".")
plt.legend()
```
### Q&A
| github_jupyter |
# Moving Average Time Series
First we start with some basic imports and set some defaults. For the most part we use the classes defined under endochrone/arima for fitting these models. Statsmodels provides a useful API for generating data. As this notebook is intended to showcase the Endochrone Algorithms, we won't use statsmodels for making predictions.
```
from functools import partial
import matplotlib.pyplot as plt
import numpy as np
from statsmodels.tsa.arima_process import arma_generate_sample
import sys; sys.path.insert(0, '..')
from endochrone.stats.correlation import acf, pacf
from endochrone.time_series.arima import ArModel, MaModel
plt.rc('figure', figsize=(20.0, 10.0))
```
### MA1
First we generate some well-behaved data which we can use to for modelling. We define a partial function purely for convenience as we'll want to generate more data for testing according to the same paramaters
```
np.random.seed(seed=1235)
ma1_func = partial(arma_generate_sample, [1], [1, 0.7], scale=1)
N_train = 500
x = ma1_func(N_train)
plt.title("Our MA1 training set")
plt.plot(range(N_train), x);
```
First we look at our ACF and PACF for patterns
```
n_lags = 25
ar1_acf = acf(x, lags=n_lags).values
ar1_pacf = pacf(x, lags=n_lags).values
fig1, a1 = plt.subplots(2,1)
a1[0].scatter(range(n_lags+1), ar1_acf)
a1[0].set_title('ACF')
a1[1].scatter(range(n_lags+1), ar1_pacf)
a1[1].set_title('PACF');
```
There is a slight (but significant) correlation in the 1-lag ACF which quickly decays to zero for subsequent lags. This suggests an MA1 model. We already knew this as we generated the data!
```
ma1 = MaModel(order=1)
ma1.fit(x)
print(ma1.thetas_)
```
Which is pretty close to what we expect. Our intercept to be around zero, and the coefficient to be 0.7 as this is what we put in our signal generator. We should check our model correcly accounts for all available correlation though. We can do this by inspecting the residuals which we'd expect to be indistinguishable from noise
```
ma1_res = ma1.residuals_
n_lags = 25
ma1_res_acfs = acf(ma1_res, lags=n_lags).values
ma1_res_pacfs = pacf(ma1_res, lags=n_lags).values
fig, a = plt.subplots(2,1)
a[0].scatter(range(n_lags+1), ma1_res_acfs)
a[0].set_title('ACF')
a[1].scatter(range(n_lags+1), ma1_res_pacfs)
a[1].set_title('PACF');
print('Mean Sq Error:', np.mean(ma1_res**2))
print('Mean of residuals:', np.mean(ma1_res))
print('Variance of residuals:', np.var(ma1_res))
plt.title('Histogram of Residues');
plt.hist(ma1_res, bins=10);
```
Looks pretty normal to me!
Now lets see how we do against some new data generated according to the same profile. (NB. We do this stepwise - we aren't predicting 30 steps out, we predict each step by looking at the previous correct answer.)
```
N_test = 300
y = ma1_func(N_test)
y_pred = ma1.predict(y)
plt.plot(range(N_test), y, c='green', label='act')
plt.plot(range(1, N_test+1), y_pred, c='blue', label='prediction')
plt.legend();
ma1_errs = y[1:] - y_pred[:-1]
print('Mean Sq Error:', np.mean(ma1_errs**2))
print('Mean Error:', np.mean(ma1_errs))
print('Var Error:', np.var(ma1_errs))
```
### MA7
Let's try a slightly more complex model. This time we'll add values for the 7th term to simulate a weekly cycle. (NB I initially tried this with a 28 day term too, but both the endochrone and statsmodel were much too slow to fit to be of practical use)
```
np.random.seed(seed=14545)
corrs = [1, 0.25] + 5*[0] + [0.84]
ma2_func = partial(arma_generate_sample, [1], corrs, scale=1)
N_train = 5000
x2 = ma2_func(N_train)
plt.title("Our MA7 training set")
plt.plot(range(100), x2[:100]);
```
You can see the cycling as we know it's there, but the ACF and PACF should make it much clearer
```
n_lags = 30
ma2_acf = acf(x2, lags=n_lags).values
ma2_pacf = pacf(x2, lags=n_lags).values
fig2, a2 = plt.subplots(2,1)
fig2.size=(6,4)
a2[0].scatter(range(n_lags+1), ma2_acf)
a2[0].set_title('ACF')
a2[1].scatter(range(n_lags+1), ma2_pacf)
a2[1].set_title('PACF');
```
Indeed we see cycles on a weekly basis. Let's fit our model. We'll plot our coefficients as it's easier to see which ones are far from zero. It's a little slow...
```
ma2 = MaModel(order=7)
ma2.fit(x2)
plt.scatter(range(0,8), ma2.thetas_);
plt.title("Coefficients of our MA7 model");
N_test = 300
y2 = ma2_func(N_test)
y2_pred = ma2.predict(y2)
x2_pred = ma2.predict(x2)
plt.plot(range(0, 120), y2[0:120], c='green', label='act')
plt.plot(range(7, 120), y2_pred[:113], c='blue', label='prediction')
plt.legend();
```
Though we predicted 271 steps, only 60 are shown in the graph. If we plot more than that and it's a little hard to see the cycling. You can see a broad repeating pattern of peaks every 7 days. Though I'll admit visually the fit doesn't seem that good
Let's finally compare our errors:
```
print('MSE(Training): %06f' % np.mean(ma2.residuals_**2))
ma2_errs = y2[7:] - y2_pred[:-7]
print('MSE(Forecast): %06f' % np.mean(ma2_errs**2))
```
This is about what we'd expect as it's the variance of our white noise. As these values are similar it suggests we've avoided over/under-fitting. I'm not completely satisfied with the fit here, so lets quickly compare to the statsmodels implementation.
```
from statsmodels.tsa.arima_model import ARMA
sm_ma2 = ARMA(x2, order=(0,7))
sm_fit_results = sm_ma2.fit()
plt.scatter(range(0,8), ma2.thetas_ - sm_fit_results.params, c='green', label='Difference between Models');
plt.title("Comparision of MA7 model coefficients");
```
The differences here are small enough that we can call the two models equivalent
| github_jupyter |
# RadarCOVID-Report
## Data Extraction
```
import datetime
import json
import logging
import os
import shutil
import tempfile
import textwrap
import uuid
import matplotlib.pyplot as plt
import matplotlib.ticker
import numpy as np
import pandas as pd
import pycountry
import retry
import seaborn as sns
%matplotlib inline
current_working_directory = os.environ.get("PWD")
if current_working_directory:
os.chdir(current_working_directory)
sns.set()
matplotlib.rcParams["figure.figsize"] = (15, 6)
extraction_datetime = datetime.datetime.utcnow()
extraction_date = extraction_datetime.strftime("%Y-%m-%d")
extraction_previous_datetime = extraction_datetime - datetime.timedelta(days=1)
extraction_previous_date = extraction_previous_datetime.strftime("%Y-%m-%d")
extraction_date_with_hour = datetime.datetime.utcnow().strftime("%Y-%m-%d@%H")
current_hour = datetime.datetime.utcnow().hour
are_today_results_partial = current_hour != 23
```
### Constants
```
from Modules.ExposureNotification import exposure_notification_io
spain_region_country_code = "ES"
germany_region_country_code = "DE"
default_backend_identifier = spain_region_country_code
backend_generation_days = 7 * 2
daily_summary_days = 7 * 4 * 3
daily_plot_days = 7 * 4
tek_dumps_load_limit = daily_summary_days + 1
```
### Parameters
```
environment_backend_identifier = os.environ.get("RADARCOVID_REPORT__BACKEND_IDENTIFIER")
if environment_backend_identifier:
report_backend_identifier = environment_backend_identifier
else:
report_backend_identifier = default_backend_identifier
report_backend_identifier
environment_enable_multi_backend_download = \
os.environ.get("RADARCOVID_REPORT__ENABLE_MULTI_BACKEND_DOWNLOAD")
if environment_enable_multi_backend_download:
report_backend_identifiers = None
else:
report_backend_identifiers = [report_backend_identifier]
report_backend_identifiers
environment_invalid_shared_diagnoses_dates = \
os.environ.get("RADARCOVID_REPORT__INVALID_SHARED_DIAGNOSES_DATES")
if environment_invalid_shared_diagnoses_dates:
invalid_shared_diagnoses_dates = environment_invalid_shared_diagnoses_dates.split(",")
else:
invalid_shared_diagnoses_dates = []
invalid_shared_diagnoses_dates
```
### COVID-19 Cases
```
report_backend_client = \
exposure_notification_io.get_backend_client_with_identifier(
backend_identifier=report_backend_identifier)
@retry.retry(tries=10, delay=10, backoff=1.1, jitter=(0, 10))
def download_cases_dataframe():
return pd.read_csv("https://raw.githubusercontent.com/owid/covid-19-data/master/public/data/owid-covid-data.csv")
confirmed_df_ = download_cases_dataframe()
confirmed_df_.iloc[0]
confirmed_df = confirmed_df_.copy()
confirmed_df = confirmed_df[["date", "new_cases", "iso_code"]]
confirmed_df.rename(
columns={
"date": "sample_date",
"iso_code": "country_code",
},
inplace=True)
def convert_iso_alpha_3_to_alpha_2(x):
try:
return pycountry.countries.get(alpha_3=x).alpha_2
except Exception as e:
logging.info(f"Error converting country ISO Alpha 3 code '{x}': {repr(e)}")
return None
confirmed_df["country_code"] = confirmed_df.country_code.apply(convert_iso_alpha_3_to_alpha_2)
confirmed_df.dropna(inplace=True)
confirmed_df["sample_date"] = pd.to_datetime(confirmed_df.sample_date, dayfirst=True)
confirmed_df["sample_date"] = confirmed_df.sample_date.dt.strftime("%Y-%m-%d")
confirmed_df.sort_values("sample_date", inplace=True)
confirmed_df.tail()
confirmed_days = pd.date_range(
start=confirmed_df.iloc[0].sample_date,
end=extraction_datetime)
confirmed_days_df = pd.DataFrame(data=confirmed_days, columns=["sample_date"])
confirmed_days_df["sample_date_string"] = \
confirmed_days_df.sample_date.dt.strftime("%Y-%m-%d")
confirmed_days_df.tail()
def sort_source_regions_for_display(source_regions: list) -> list:
if report_backend_identifier in source_regions:
source_regions = [report_backend_identifier] + \
list(sorted(set(source_regions).difference([report_backend_identifier])))
else:
source_regions = list(sorted(source_regions))
return source_regions
report_source_regions = report_backend_client.source_regions_for_date(
date=extraction_datetime.date())
report_source_regions = sort_source_regions_for_display(
source_regions=report_source_regions)
report_source_regions
def get_cases_dataframe(source_regions_for_date_function, columns_suffix=None):
source_regions_at_date_df = confirmed_days_df.copy()
source_regions_at_date_df["source_regions_at_date"] = \
source_regions_at_date_df.sample_date.apply(
lambda x: source_regions_for_date_function(date=x))
source_regions_at_date_df.sort_values("sample_date", inplace=True)
source_regions_at_date_df["_source_regions_group"] = source_regions_at_date_df. \
source_regions_at_date.apply(lambda x: ",".join(sort_source_regions_for_display(x)))
source_regions_at_date_df.tail()
#%%
source_regions_for_summary_df_ = \
source_regions_at_date_df[["sample_date", "_source_regions_group"]].copy()
source_regions_for_summary_df_.rename(columns={"_source_regions_group": "source_regions"}, inplace=True)
source_regions_for_summary_df_.tail()
#%%
confirmed_output_columns = ["sample_date", "new_cases", "covid_cases"]
confirmed_output_df = pd.DataFrame(columns=confirmed_output_columns)
for source_regions_group, source_regions_group_series in \
source_regions_at_date_df.groupby("_source_regions_group"):
source_regions_set = set(source_regions_group.split(","))
confirmed_source_regions_set_df = \
confirmed_df[confirmed_df.country_code.isin(source_regions_set)].copy()
confirmed_source_regions_group_df = \
confirmed_source_regions_set_df.groupby("sample_date").new_cases.sum() \
.reset_index().sort_values("sample_date")
confirmed_source_regions_group_df = \
confirmed_source_regions_group_df.merge(
confirmed_days_df[["sample_date_string"]].rename(
columns={"sample_date_string": "sample_date"}),
how="right")
confirmed_source_regions_group_df["new_cases"] = \
confirmed_source_regions_group_df["new_cases"].clip(lower=0)
confirmed_source_regions_group_df["covid_cases"] = \
confirmed_source_regions_group_df.new_cases.rolling(7, min_periods=0).mean().round()
confirmed_source_regions_group_df = \
confirmed_source_regions_group_df[confirmed_output_columns]
confirmed_source_regions_group_df = confirmed_source_regions_group_df.replace(0, np.nan)
confirmed_source_regions_group_df.fillna(method="ffill", inplace=True)
confirmed_source_regions_group_df = \
confirmed_source_regions_group_df[
confirmed_source_regions_group_df.sample_date.isin(
source_regions_group_series.sample_date_string)]
confirmed_output_df = confirmed_output_df.append(confirmed_source_regions_group_df)
result_df = confirmed_output_df.copy()
result_df.tail()
#%%
result_df.rename(columns={"sample_date": "sample_date_string"}, inplace=True)
result_df = confirmed_days_df[["sample_date_string"]].merge(result_df, how="left")
result_df.sort_values("sample_date_string", inplace=True)
result_df.fillna(method="ffill", inplace=True)
result_df.tail()
#%%
result_df[["new_cases", "covid_cases"]].plot()
if columns_suffix:
result_df.rename(
columns={
"new_cases": "new_cases_" + columns_suffix,
"covid_cases": "covid_cases_" + columns_suffix},
inplace=True)
return result_df, source_regions_for_summary_df_
confirmed_eu_df, source_regions_for_summary_df = get_cases_dataframe(
report_backend_client.source_regions_for_date)
confirmed_es_df, _ = get_cases_dataframe(
lambda date: [spain_region_country_code],
columns_suffix=spain_region_country_code.lower())
```
### Extract API TEKs
```
raw_zip_path_prefix = "Data/TEKs/Raw/"
base_backend_identifiers = [report_backend_identifier]
multi_backend_exposure_keys_df = \
exposure_notification_io.download_exposure_keys_from_backends(
backend_identifiers=report_backend_identifiers,
generation_days=backend_generation_days,
fail_on_error_backend_identifiers=base_backend_identifiers,
save_raw_zip_path_prefix=raw_zip_path_prefix)
multi_backend_exposure_keys_df["region"] = multi_backend_exposure_keys_df["backend_identifier"]
multi_backend_exposure_keys_df.rename(
columns={
"generation_datetime": "sample_datetime",
"generation_date_string": "sample_date_string",
},
inplace=True)
multi_backend_exposure_keys_df.head()
early_teks_df = multi_backend_exposure_keys_df[
multi_backend_exposure_keys_df.rolling_period < 144].copy()
early_teks_df["rolling_period_in_hours"] = early_teks_df.rolling_period / 6
early_teks_df[early_teks_df.sample_date_string != extraction_date] \
.rolling_period_in_hours.hist(bins=list(range(24)))
early_teks_df[early_teks_df.sample_date_string == extraction_date] \
.rolling_period_in_hours.hist(bins=list(range(24)))
multi_backend_exposure_keys_df = multi_backend_exposure_keys_df[[
"sample_date_string", "region", "key_data"]]
multi_backend_exposure_keys_df.head()
active_regions = \
multi_backend_exposure_keys_df.groupby("region").key_data.nunique().sort_values().index.unique().tolist()
active_regions
multi_backend_summary_df = multi_backend_exposure_keys_df.groupby(
["sample_date_string", "region"]).key_data.nunique().reset_index() \
.pivot(index="sample_date_string", columns="region") \
.sort_index(ascending=False)
multi_backend_summary_df.rename(
columns={"key_data": "shared_teks_by_generation_date"},
inplace=True)
multi_backend_summary_df.rename_axis("sample_date", inplace=True)
multi_backend_summary_df = multi_backend_summary_df.fillna(0).astype(int)
multi_backend_summary_df = multi_backend_summary_df.head(backend_generation_days)
multi_backend_summary_df.head()
def compute_keys_cross_sharing(x):
teks_x = x.key_data_x.item()
common_teks = set(teks_x).intersection(x.key_data_y.item())
common_teks_fraction = len(common_teks) / len(teks_x)
return pd.Series(dict(
common_teks=common_teks,
common_teks_fraction=common_teks_fraction,
))
multi_backend_exposure_keys_by_region_df = \
multi_backend_exposure_keys_df.groupby("region").key_data.unique().reset_index()
multi_backend_exposure_keys_by_region_df["_merge"] = True
multi_backend_exposure_keys_by_region_combination_df = \
multi_backend_exposure_keys_by_region_df.merge(
multi_backend_exposure_keys_by_region_df, on="_merge")
multi_backend_exposure_keys_by_region_combination_df.drop(
columns=["_merge"], inplace=True)
if multi_backend_exposure_keys_by_region_combination_df.region_x.nunique() > 1:
multi_backend_exposure_keys_by_region_combination_df = \
multi_backend_exposure_keys_by_region_combination_df[
multi_backend_exposure_keys_by_region_combination_df.region_x !=
multi_backend_exposure_keys_by_region_combination_df.region_y]
multi_backend_exposure_keys_cross_sharing_df = \
multi_backend_exposure_keys_by_region_combination_df \
.groupby(["region_x", "region_y"]) \
.apply(compute_keys_cross_sharing) \
.reset_index()
multi_backend_cross_sharing_summary_df = \
multi_backend_exposure_keys_cross_sharing_df.pivot_table(
values=["common_teks_fraction"],
columns="region_x",
index="region_y",
aggfunc=lambda x: x.item())
multi_backend_cross_sharing_summary_df
multi_backend_without_active_region_exposure_keys_df = \
multi_backend_exposure_keys_df[multi_backend_exposure_keys_df.region != report_backend_identifier]
multi_backend_without_active_region = \
multi_backend_without_active_region_exposure_keys_df.groupby("region").key_data.nunique().sort_values().index.unique().tolist()
multi_backend_without_active_region
exposure_keys_summary_df = multi_backend_exposure_keys_df[
multi_backend_exposure_keys_df.region == report_backend_identifier]
exposure_keys_summary_df.drop(columns=["region"], inplace=True)
exposure_keys_summary_df = \
exposure_keys_summary_df.groupby(["sample_date_string"]).key_data.nunique().to_frame()
exposure_keys_summary_df = \
exposure_keys_summary_df.reset_index().set_index("sample_date_string")
exposure_keys_summary_df.sort_index(ascending=False, inplace=True)
exposure_keys_summary_df.rename(columns={"key_data": "shared_teks_by_generation_date"}, inplace=True)
exposure_keys_summary_df.head()
```
### Dump API TEKs
```
tek_list_df = multi_backend_exposure_keys_df[
["sample_date_string", "region", "key_data"]].copy()
tek_list_df["key_data"] = tek_list_df["key_data"].apply(str)
tek_list_df.rename(columns={
"sample_date_string": "sample_date",
"key_data": "tek_list"}, inplace=True)
tek_list_df = tek_list_df.groupby(
["sample_date", "region"]).tek_list.unique().reset_index()
tek_list_df["extraction_date"] = extraction_date
tek_list_df["extraction_date_with_hour"] = extraction_date_with_hour
tek_list_path_prefix = "Data/TEKs/"
tek_list_current_path = tek_list_path_prefix + f"/Current/RadarCOVID-TEKs.json"
tek_list_daily_path = tek_list_path_prefix + f"Daily/RadarCOVID-TEKs-{extraction_date}.json"
tek_list_hourly_path = tek_list_path_prefix + f"Hourly/RadarCOVID-TEKs-{extraction_date_with_hour}.json"
for path in [tek_list_current_path, tek_list_daily_path, tek_list_hourly_path]:
os.makedirs(os.path.dirname(path), exist_ok=True)
tek_list_base_df = tek_list_df[tek_list_df.region == report_backend_identifier]
tek_list_base_df.drop(columns=["extraction_date", "extraction_date_with_hour"]).to_json(
tek_list_current_path,
lines=True, orient="records")
tek_list_base_df.drop(columns=["extraction_date_with_hour"]).to_json(
tek_list_daily_path,
lines=True, orient="records")
tek_list_base_df.to_json(
tek_list_hourly_path,
lines=True, orient="records")
tek_list_base_df.head()
```
### Load TEK Dumps
```
import glob
def load_extracted_teks(mode, region=None, limit=None) -> pd.DataFrame:
extracted_teks_df = pd.DataFrame(columns=["region"])
file_paths = list(reversed(sorted(glob.glob(tek_list_path_prefix + mode + "/RadarCOVID-TEKs-*.json"))))
if limit:
file_paths = file_paths[:limit]
for file_path in file_paths:
logging.info(f"Loading TEKs from '{file_path}'...")
iteration_extracted_teks_df = pd.read_json(file_path, lines=True)
extracted_teks_df = extracted_teks_df.append(
iteration_extracted_teks_df, sort=False)
extracted_teks_df["region"] = \
extracted_teks_df.region.fillna(spain_region_country_code).copy()
if region:
extracted_teks_df = \
extracted_teks_df[extracted_teks_df.region == region]
return extracted_teks_df
daily_extracted_teks_df = load_extracted_teks(
mode="Daily",
region=report_backend_identifier,
limit=tek_dumps_load_limit)
daily_extracted_teks_df.head()
exposure_keys_summary_df_ = daily_extracted_teks_df \
.sort_values("extraction_date", ascending=False) \
.groupby("sample_date").tek_list.first() \
.to_frame()
exposure_keys_summary_df_.index.name = "sample_date_string"
exposure_keys_summary_df_["tek_list"] = \
exposure_keys_summary_df_.tek_list.apply(len)
exposure_keys_summary_df_ = exposure_keys_summary_df_ \
.rename(columns={"tek_list": "shared_teks_by_generation_date"}) \
.sort_index(ascending=False)
exposure_keys_summary_df = exposure_keys_summary_df_
exposure_keys_summary_df.head()
```
### Daily New TEKs
```
tek_list_df = daily_extracted_teks_df.groupby("extraction_date").tek_list.apply(
lambda x: set(sum(x, []))).reset_index()
tek_list_df = tek_list_df.set_index("extraction_date").sort_index(ascending=True)
tek_list_df.head()
def compute_teks_by_generation_and_upload_date(date):
day_new_teks_set_df = tek_list_df.copy().diff()
try:
day_new_teks_set = day_new_teks_set_df[
day_new_teks_set_df.index == date].tek_list.item()
except ValueError:
day_new_teks_set = None
if pd.isna(day_new_teks_set):
day_new_teks_set = set()
day_new_teks_df = daily_extracted_teks_df[
daily_extracted_teks_df.extraction_date == date].copy()
day_new_teks_df["shared_teks"] = \
day_new_teks_df.tek_list.apply(lambda x: set(x).intersection(day_new_teks_set))
day_new_teks_df["shared_teks"] = \
day_new_teks_df.shared_teks.apply(len)
day_new_teks_df["upload_date"] = date
day_new_teks_df.rename(columns={"sample_date": "generation_date"}, inplace=True)
day_new_teks_df = day_new_teks_df[
["upload_date", "generation_date", "shared_teks"]]
day_new_teks_df["generation_to_upload_days"] = \
(pd.to_datetime(day_new_teks_df.upload_date) -
pd.to_datetime(day_new_teks_df.generation_date)).dt.days
day_new_teks_df = day_new_teks_df[day_new_teks_df.shared_teks > 0]
return day_new_teks_df
shared_teks_generation_to_upload_df = pd.DataFrame()
for upload_date in daily_extracted_teks_df.extraction_date.unique():
shared_teks_generation_to_upload_df = \
shared_teks_generation_to_upload_df.append(
compute_teks_by_generation_and_upload_date(date=upload_date))
shared_teks_generation_to_upload_df \
.sort_values(["upload_date", "generation_date"], ascending=False, inplace=True)
shared_teks_generation_to_upload_df.tail()
today_new_teks_df = \
shared_teks_generation_to_upload_df[
shared_teks_generation_to_upload_df.upload_date == extraction_date].copy()
today_new_teks_df.tail()
if not today_new_teks_df.empty:
today_new_teks_df.set_index("generation_to_upload_days") \
.sort_index().shared_teks.plot.bar()
generation_to_upload_period_pivot_df = \
shared_teks_generation_to_upload_df[
["upload_date", "generation_to_upload_days", "shared_teks"]] \
.pivot(index="upload_date", columns="generation_to_upload_days") \
.sort_index(ascending=False).fillna(0).astype(int) \
.droplevel(level=0, axis=1)
generation_to_upload_period_pivot_df.head()
new_tek_df = tek_list_df.diff().tek_list.apply(
lambda x: len(x) if not pd.isna(x) else None).to_frame().reset_index()
new_tek_df.rename(columns={
"tek_list": "shared_teks_by_upload_date",
"extraction_date": "sample_date_string",}, inplace=True)
new_tek_df.tail()
shared_teks_uploaded_on_generation_date_df = shared_teks_generation_to_upload_df[
shared_teks_generation_to_upload_df.generation_to_upload_days == 0] \
[["upload_date", "shared_teks"]].rename(
columns={
"upload_date": "sample_date_string",
"shared_teks": "shared_teks_uploaded_on_generation_date",
})
shared_teks_uploaded_on_generation_date_df.head()
estimated_shared_diagnoses_df = shared_teks_generation_to_upload_df \
.groupby(["upload_date"]).shared_teks.max().reset_index() \
.sort_values(["upload_date"], ascending=False) \
.rename(columns={
"upload_date": "sample_date_string",
"shared_teks": "shared_diagnoses",
})
invalid_shared_diagnoses_dates_mask = \
estimated_shared_diagnoses_df.sample_date_string.isin(invalid_shared_diagnoses_dates)
estimated_shared_diagnoses_df[invalid_shared_diagnoses_dates_mask] = 0
estimated_shared_diagnoses_df.head()
```
### Hourly New TEKs
```
hourly_extracted_teks_df = load_extracted_teks(
mode="Hourly", region=report_backend_identifier, limit=25)
hourly_extracted_teks_df.head()
hourly_new_tek_count_df = hourly_extracted_teks_df \
.groupby("extraction_date_with_hour").tek_list. \
apply(lambda x: set(sum(x, []))).reset_index().copy()
hourly_new_tek_count_df = hourly_new_tek_count_df.set_index("extraction_date_with_hour") \
.sort_index(ascending=True)
hourly_new_tek_count_df["new_tek_list"] = hourly_new_tek_count_df.tek_list.diff()
hourly_new_tek_count_df["new_tek_count"] = hourly_new_tek_count_df.new_tek_list.apply(
lambda x: len(x) if not pd.isna(x) else 0)
hourly_new_tek_count_df.rename(columns={
"new_tek_count": "shared_teks_by_upload_date"}, inplace=True)
hourly_new_tek_count_df = hourly_new_tek_count_df.reset_index()[[
"extraction_date_with_hour", "shared_teks_by_upload_date"]]
hourly_new_tek_count_df.head()
hourly_summary_df = hourly_new_tek_count_df.copy()
hourly_summary_df.set_index("extraction_date_with_hour", inplace=True)
hourly_summary_df = hourly_summary_df.fillna(0).astype(int).reset_index()
hourly_summary_df["datetime_utc"] = pd.to_datetime(
hourly_summary_df.extraction_date_with_hour, format="%Y-%m-%d@%H")
hourly_summary_df.set_index("datetime_utc", inplace=True)
hourly_summary_df = hourly_summary_df.tail(-1)
hourly_summary_df.head()
```
### Official Statistics
```
import requests
import pandas.io.json
official_stats_response = requests.get("https://radarcovid.covid19.gob.es/kpi/statistics/basics")
official_stats_response.raise_for_status()
official_stats_df_ = pandas.io.json.json_normalize(official_stats_response.json())
official_stats_df = official_stats_df_.copy()
official_stats_df["date"] = pd.to_datetime(official_stats_df["date"], dayfirst=True)
official_stats_df.head()
official_stats_column_map = {
"date": "sample_date",
"applicationsDownloads.totalAcummulated": "app_downloads_es_accumulated",
"communicatedContagions.totalAcummulated": "shared_diagnoses_es_accumulated",
}
accumulated_suffix = "_accumulated"
accumulated_values_columns = \
list(filter(lambda x: x.endswith(accumulated_suffix), official_stats_column_map.values()))
interpolated_values_columns = \
list(map(lambda x: x[:-len(accumulated_suffix)], accumulated_values_columns))
official_stats_df = \
official_stats_df[official_stats_column_map.keys()] \
.rename(columns=official_stats_column_map)
official_stats_df["extraction_date"] = extraction_date
official_stats_df.head()
official_stats_path = "Data/Statistics/Current/RadarCOVID-Statistics.json"
previous_official_stats_df = pd.read_json(official_stats_path, orient="records", lines=True)
previous_official_stats_df["sample_date"] = pd.to_datetime(previous_official_stats_df["sample_date"], dayfirst=True)
official_stats_df = official_stats_df.append(previous_official_stats_df)
official_stats_df.head()
official_stats_df = official_stats_df[~(official_stats_df.shared_diagnoses_es_accumulated == 0)]
official_stats_df.sort_values("extraction_date", ascending=False, inplace=True)
official_stats_df.drop_duplicates(subset=["sample_date"], keep="first", inplace=True)
official_stats_df.head()
official_stats_stored_df = official_stats_df.copy()
official_stats_stored_df["sample_date"] = official_stats_stored_df.sample_date.dt.strftime("%Y-%m-%d")
official_stats_stored_df.to_json(official_stats_path, orient="records", lines=True)
official_stats_df.drop(columns=["extraction_date"], inplace=True)
official_stats_df = confirmed_days_df.merge(official_stats_df, how="left")
official_stats_df.sort_values("sample_date", ascending=False, inplace=True)
official_stats_df.head()
official_stats_df[accumulated_values_columns] = \
official_stats_df[accumulated_values_columns] \
.astype(float).interpolate(limit_area="inside")
official_stats_df[interpolated_values_columns] = \
official_stats_df[accumulated_values_columns].diff(periods=-1)
official_stats_df.drop(columns="sample_date", inplace=True)
official_stats_df.head()
```
### Data Merge
```
result_summary_df = exposure_keys_summary_df.merge(
new_tek_df, on=["sample_date_string"], how="outer")
result_summary_df.head()
result_summary_df = result_summary_df.merge(
shared_teks_uploaded_on_generation_date_df, on=["sample_date_string"], how="outer")
result_summary_df.head()
result_summary_df = result_summary_df.merge(
estimated_shared_diagnoses_df, on=["sample_date_string"], how="outer")
result_summary_df.head()
result_summary_df = result_summary_df.merge(
official_stats_df, on=["sample_date_string"], how="outer")
result_summary_df.head()
result_summary_df = confirmed_eu_df.tail(daily_summary_days).merge(
result_summary_df, on=["sample_date_string"], how="left")
result_summary_df.head()
result_summary_df = confirmed_es_df.tail(daily_summary_days).merge(
result_summary_df, on=["sample_date_string"], how="left")
result_summary_df.head()
result_summary_df["sample_date"] = pd.to_datetime(result_summary_df.sample_date_string)
result_summary_df = result_summary_df.merge(source_regions_for_summary_df, how="left")
result_summary_df.set_index(["sample_date", "source_regions"], inplace=True)
result_summary_df.drop(columns=["sample_date_string"], inplace=True)
result_summary_df.sort_index(ascending=False, inplace=True)
result_summary_df.head()
with pd.option_context("mode.use_inf_as_na", True):
result_summary_df = result_summary_df.fillna(0).astype(int)
result_summary_df["teks_per_shared_diagnosis"] = \
(result_summary_df.shared_teks_by_upload_date / result_summary_df.shared_diagnoses).fillna(0)
result_summary_df["shared_diagnoses_per_covid_case"] = \
(result_summary_df.shared_diagnoses / result_summary_df.covid_cases).fillna(0)
result_summary_df["shared_diagnoses_per_covid_case_es"] = \
(result_summary_df.shared_diagnoses_es / result_summary_df.covid_cases_es).fillna(0)
result_summary_df.head(daily_plot_days)
def compute_aggregated_results_summary(days) -> pd.DataFrame:
aggregated_result_summary_df = result_summary_df.copy()
aggregated_result_summary_df["covid_cases_for_ratio"] = \
aggregated_result_summary_df.covid_cases.mask(
aggregated_result_summary_df.shared_diagnoses == 0, 0)
aggregated_result_summary_df["covid_cases_for_ratio_es"] = \
aggregated_result_summary_df.covid_cases_es.mask(
aggregated_result_summary_df.shared_diagnoses_es == 0, 0)
aggregated_result_summary_df = aggregated_result_summary_df \
.sort_index(ascending=True).fillna(0).rolling(days).agg({
"covid_cases": "sum",
"covid_cases_es": "sum",
"covid_cases_for_ratio": "sum",
"covid_cases_for_ratio_es": "sum",
"shared_teks_by_generation_date": "sum",
"shared_teks_by_upload_date": "sum",
"shared_diagnoses": "sum",
"shared_diagnoses_es": "sum",
}).sort_index(ascending=False)
with pd.option_context("mode.use_inf_as_na", True):
aggregated_result_summary_df = aggregated_result_summary_df.fillna(0).astype(int)
aggregated_result_summary_df["teks_per_shared_diagnosis"] = \
(aggregated_result_summary_df.shared_teks_by_upload_date /
aggregated_result_summary_df.covid_cases_for_ratio).fillna(0)
aggregated_result_summary_df["shared_diagnoses_per_covid_case"] = \
(aggregated_result_summary_df.shared_diagnoses /
aggregated_result_summary_df.covid_cases_for_ratio).fillna(0)
aggregated_result_summary_df["shared_diagnoses_per_covid_case_es"] = \
(aggregated_result_summary_df.shared_diagnoses_es /
aggregated_result_summary_df.covid_cases_for_ratio_es).fillna(0)
return aggregated_result_summary_df
aggregated_result_with_7_days_window_summary_df = compute_aggregated_results_summary(days=7)
aggregated_result_with_7_days_window_summary_df.head()
last_7_days_summary = aggregated_result_with_7_days_window_summary_df.to_dict(orient="records")[1]
last_7_days_summary
aggregated_result_with_14_days_window_summary_df = compute_aggregated_results_summary(days=13)
last_14_days_summary = aggregated_result_with_14_days_window_summary_df.to_dict(orient="records")[1]
last_14_days_summary
```
## Report Results
```
display_column_name_mapping = {
"sample_date": "Sample\u00A0Date\u00A0(UTC)",
"source_regions": "Source Countries",
"datetime_utc": "Timestamp (UTC)",
"upload_date": "Upload Date (UTC)",
"generation_to_upload_days": "Generation to Upload Period in Days",
"region": "Backend",
"region_x": "Backend\u00A0(A)",
"region_y": "Backend\u00A0(B)",
"common_teks": "Common TEKs Shared Between Backends",
"common_teks_fraction": "Fraction of TEKs in Backend (A) Available in Backend (B)",
"covid_cases": "COVID-19 Cases (Source Countries)",
"shared_teks_by_generation_date": "Shared TEKs by Generation Date (Source Countries)",
"shared_teks_by_upload_date": "Shared TEKs by Upload Date (Source Countries)",
"shared_teks_uploaded_on_generation_date": "Shared TEKs Uploaded on Generation Date (Source Countries)",
"shared_diagnoses": "Shared Diagnoses (Source Countries – Estimation)",
"teks_per_shared_diagnosis": "TEKs Uploaded per Shared Diagnosis (Source Countries)",
"shared_diagnoses_per_covid_case": "Usage Ratio (Source Countries)",
"covid_cases_es": "COVID-19 Cases (Spain)",
"app_downloads_es": "App Downloads (Spain – Official)",
"shared_diagnoses_es": "Shared Diagnoses (Spain – Official)",
"shared_diagnoses_per_covid_case_es": "Usage Ratio (Spain)",
}
summary_columns = [
"covid_cases",
"shared_teks_by_generation_date",
"shared_teks_by_upload_date",
"shared_teks_uploaded_on_generation_date",
"shared_diagnoses",
"teks_per_shared_diagnosis",
"shared_diagnoses_per_covid_case",
"covid_cases_es",
"app_downloads_es",
"shared_diagnoses_es",
"shared_diagnoses_per_covid_case_es",
]
summary_percentage_columns= [
"shared_diagnoses_per_covid_case_es",
"shared_diagnoses_per_covid_case",
]
```
### Daily Summary Table
```
result_summary_df_ = result_summary_df.copy()
result_summary_df = result_summary_df[summary_columns]
result_summary_with_display_names_df = result_summary_df \
.rename_axis(index=display_column_name_mapping) \
.rename(columns=display_column_name_mapping)
result_summary_with_display_names_df
```
### Daily Summary Plots
```
result_plot_summary_df = result_summary_df.head(daily_plot_days)[summary_columns] \
.droplevel(level=["source_regions"]) \
.rename_axis(index=display_column_name_mapping) \
.rename(columns=display_column_name_mapping)
summary_ax_list = result_plot_summary_df.sort_index(ascending=True).plot.bar(
title=f"Daily Summary",
rot=45, subplots=True, figsize=(15, 30), legend=False)
ax_ = summary_ax_list[0]
ax_.get_figure().tight_layout()
ax_.get_figure().subplots_adjust(top=0.95)
_ = ax_.set_xticklabels(sorted(result_plot_summary_df.index.strftime("%Y-%m-%d").tolist()))
for percentage_column in summary_percentage_columns:
percentage_column_index = summary_columns.index(percentage_column)
summary_ax_list[percentage_column_index].yaxis \
.set_major_formatter(matplotlib.ticker.PercentFormatter(1.0))
```
### Daily Generation to Upload Period Table
```
display_generation_to_upload_period_pivot_df = \
generation_to_upload_period_pivot_df \
.head(backend_generation_days)
display_generation_to_upload_period_pivot_df \
.head(backend_generation_days) \
.rename_axis(columns=display_column_name_mapping) \
.rename_axis(index=display_column_name_mapping)
fig, generation_to_upload_period_pivot_table_ax = plt.subplots(
figsize=(12, 1 + 0.6 * len(display_generation_to_upload_period_pivot_df)))
generation_to_upload_period_pivot_table_ax.set_title(
"Shared TEKs Generation to Upload Period Table")
sns.heatmap(
data=display_generation_to_upload_period_pivot_df
.rename_axis(columns=display_column_name_mapping)
.rename_axis(index=display_column_name_mapping),
fmt=".0f",
annot=True,
ax=generation_to_upload_period_pivot_table_ax)
generation_to_upload_period_pivot_table_ax.get_figure().tight_layout()
```
### Hourly Summary Plots
```
hourly_summary_ax_list = hourly_summary_df \
.rename_axis(index=display_column_name_mapping) \
.rename(columns=display_column_name_mapping) \
.plot.bar(
title=f"Last 24h Summary",
rot=45, subplots=True, legend=False)
ax_ = hourly_summary_ax_list[-1]
ax_.get_figure().tight_layout()
ax_.get_figure().subplots_adjust(top=0.9)
_ = ax_.set_xticklabels(sorted(hourly_summary_df.index.strftime("%Y-%m-%d@%H").tolist()))
```
### Publish Results
```
github_repository = os.environ.get("GITHUB_REPOSITORY")
if github_repository is None:
github_repository = "pvieito/Radar-STATS"
github_project_base_url = "https://github.com/" + github_repository
display_formatters = {
display_column_name_mapping["teks_per_shared_diagnosis"]: lambda x: f"{x:.2f}" if x != 0 else "",
display_column_name_mapping["shared_diagnoses_per_covid_case"]: lambda x: f"{x:.2%}" if x != 0 else "",
display_column_name_mapping["shared_diagnoses_per_covid_case_es"]: lambda x: f"{x:.2%}" if x != 0 else "",
}
general_columns = \
list(filter(lambda x: x not in display_formatters, display_column_name_mapping.values()))
general_formatter = lambda x: f"{x}" if x != 0 else ""
display_formatters.update(dict(map(lambda x: (x, general_formatter), general_columns)))
daily_summary_table_html = result_summary_with_display_names_df \
.head(daily_plot_days) \
.rename_axis(index=display_column_name_mapping) \
.rename(columns=display_column_name_mapping) \
.to_html(formatters=display_formatters)
multi_backend_summary_table_html = multi_backend_summary_df \
.head(daily_plot_days) \
.rename_axis(columns=display_column_name_mapping) \
.rename(columns=display_column_name_mapping) \
.rename_axis(index=display_column_name_mapping) \
.to_html(formatters=display_formatters)
def format_multi_backend_cross_sharing_fraction(x):
if pd.isna(x):
return "-"
elif round(x * 100, 1) == 0:
return ""
else:
return f"{x:.1%}"
multi_backend_cross_sharing_summary_table_html = multi_backend_cross_sharing_summary_df \
.rename_axis(columns=display_column_name_mapping) \
.rename(columns=display_column_name_mapping) \
.rename_axis(index=display_column_name_mapping) \
.to_html(
classes="table-center",
formatters=display_formatters,
float_format=format_multi_backend_cross_sharing_fraction)
multi_backend_cross_sharing_summary_table_html = \
multi_backend_cross_sharing_summary_table_html \
.replace("<tr>","<tr style=\"text-align: center;\">")
extraction_date_result_summary_df = \
result_summary_df[result_summary_df.index.get_level_values("sample_date") == extraction_date]
extraction_date_result_hourly_summary_df = \
hourly_summary_df[hourly_summary_df.extraction_date_with_hour == extraction_date_with_hour]
covid_cases = \
extraction_date_result_summary_df.covid_cases.item()
shared_teks_by_generation_date = \
extraction_date_result_summary_df.shared_teks_by_generation_date.item()
shared_teks_by_upload_date = \
extraction_date_result_summary_df.shared_teks_by_upload_date.item()
shared_diagnoses = \
extraction_date_result_summary_df.shared_diagnoses.item()
teks_per_shared_diagnosis = \
extraction_date_result_summary_df.teks_per_shared_diagnosis.item()
shared_diagnoses_per_covid_case = \
extraction_date_result_summary_df.shared_diagnoses_per_covid_case.item()
shared_teks_by_upload_date_last_hour = \
extraction_date_result_hourly_summary_df.shared_teks_by_upload_date.sum().astype(int)
display_source_regions = ", ".join(report_source_regions)
if len(report_source_regions) == 1:
display_brief_source_regions = report_source_regions[0]
else:
display_brief_source_regions = f"{len(report_source_regions)} 🇪🇺"
def get_temporary_image_path() -> str:
return os.path.join(tempfile.gettempdir(), str(uuid.uuid4()) + ".png")
def save_temporary_plot_image(ax):
if isinstance(ax, np.ndarray):
ax = ax[0]
media_path = get_temporary_image_path()
ax.get_figure().savefig(media_path)
return media_path
def save_temporary_dataframe_image(df):
import dataframe_image as dfi
df = df.copy()
df_styler = df.style.format(display_formatters)
media_path = get_temporary_image_path()
dfi.export(df_styler, media_path)
return media_path
summary_plots_image_path = save_temporary_plot_image(
ax=summary_ax_list)
summary_table_image_path = save_temporary_dataframe_image(
df=result_summary_with_display_names_df)
hourly_summary_plots_image_path = save_temporary_plot_image(
ax=hourly_summary_ax_list)
multi_backend_summary_table_image_path = save_temporary_dataframe_image(
df=multi_backend_summary_df)
generation_to_upload_period_pivot_table_image_path = save_temporary_plot_image(
ax=generation_to_upload_period_pivot_table_ax)
```
### Save Results
```
report_resources_path_prefix = "Data/Resources/Current/RadarCOVID-Report-"
result_summary_df.to_csv(
report_resources_path_prefix + "Summary-Table.csv")
result_summary_df.to_html(
report_resources_path_prefix + "Summary-Table.html")
hourly_summary_df.to_csv(
report_resources_path_prefix + "Hourly-Summary-Table.csv")
multi_backend_summary_df.to_csv(
report_resources_path_prefix + "Multi-Backend-Summary-Table.csv")
multi_backend_cross_sharing_summary_df.to_csv(
report_resources_path_prefix + "Multi-Backend-Cross-Sharing-Summary-Table.csv")
generation_to_upload_period_pivot_df.to_csv(
report_resources_path_prefix + "Generation-Upload-Period-Table.csv")
_ = shutil.copyfile(
summary_plots_image_path,
report_resources_path_prefix + "Summary-Plots.png")
_ = shutil.copyfile(
summary_table_image_path,
report_resources_path_prefix + "Summary-Table.png")
_ = shutil.copyfile(
hourly_summary_plots_image_path,
report_resources_path_prefix + "Hourly-Summary-Plots.png")
_ = shutil.copyfile(
multi_backend_summary_table_image_path,
report_resources_path_prefix + "Multi-Backend-Summary-Table.png")
_ = shutil.copyfile(
generation_to_upload_period_pivot_table_image_path,
report_resources_path_prefix + "Generation-Upload-Period-Table.png")
```
### Publish Results as JSON
```
def generate_summary_api_results(df: pd.DataFrame) -> list:
api_df = df.reset_index().copy()
api_df["sample_date_string"] = \
api_df["sample_date"].dt.strftime("%Y-%m-%d")
api_df["source_regions"] = \
api_df["source_regions"].apply(lambda x: x.split(","))
return api_df.to_dict(orient="records")
summary_api_results = \
generate_summary_api_results(df=result_summary_df)
today_summary_api_results = \
generate_summary_api_results(df=extraction_date_result_summary_df)[0]
summary_results = dict(
backend_identifier=report_backend_identifier,
source_regions=report_source_regions,
extraction_datetime=extraction_datetime,
extraction_date=extraction_date,
extraction_date_with_hour=extraction_date_with_hour,
last_hour=dict(
shared_teks_by_upload_date=shared_teks_by_upload_date_last_hour,
shared_diagnoses=0,
),
today=today_summary_api_results,
last_7_days=last_7_days_summary,
last_14_days=last_14_days_summary,
daily_results=summary_api_results)
summary_results = \
json.loads(pd.Series([summary_results]).to_json(orient="records"))[0]
with open(report_resources_path_prefix + "Summary-Results.json", "w") as f:
json.dump(summary_results, f, indent=4)
```
### Publish on README
```
with open("Data/Templates/README.md", "r") as f:
readme_contents = f.read()
readme_contents = readme_contents.format(
extraction_date_with_hour=extraction_date_with_hour,
github_project_base_url=github_project_base_url,
daily_summary_table_html=daily_summary_table_html,
multi_backend_summary_table_html=multi_backend_summary_table_html,
multi_backend_cross_sharing_summary_table_html=multi_backend_cross_sharing_summary_table_html,
display_source_regions=display_source_regions)
with open("README.md", "w") as f:
f.write(readme_contents)
```
### Publish on Twitter
```
enable_share_to_twitter = os.environ.get("RADARCOVID_REPORT__ENABLE_PUBLISH_ON_TWITTER")
github_event_name = os.environ.get("GITHUB_EVENT_NAME")
if enable_share_to_twitter and github_event_name == "schedule" and \
(shared_teks_by_upload_date_last_hour or not are_today_results_partial):
import tweepy
twitter_api_auth_keys = os.environ["RADARCOVID_REPORT__TWITTER_API_AUTH_KEYS"]
twitter_api_auth_keys = twitter_api_auth_keys.split(":")
auth = tweepy.OAuthHandler(twitter_api_auth_keys[0], twitter_api_auth_keys[1])
auth.set_access_token(twitter_api_auth_keys[2], twitter_api_auth_keys[3])
api = tweepy.API(auth)
summary_plots_media = api.media_upload(summary_plots_image_path)
summary_table_media = api.media_upload(summary_table_image_path)
generation_to_upload_period_pivot_table_image_media = api.media_upload(generation_to_upload_period_pivot_table_image_path)
media_ids = [
summary_plots_media.media_id,
summary_table_media.media_id,
generation_to_upload_period_pivot_table_image_media.media_id,
]
if are_today_results_partial:
today_addendum = " (Partial)"
else:
today_addendum = ""
def format_shared_diagnoses_per_covid_case(value) -> str:
if value == 0:
return "–"
return f"≤{value:.2%}"
display_shared_diagnoses_per_covid_case = \
format_shared_diagnoses_per_covid_case(value=shared_diagnoses_per_covid_case)
display_last_14_days_shared_diagnoses_per_covid_case = \
format_shared_diagnoses_per_covid_case(value=last_14_days_summary["shared_diagnoses_per_covid_case"])
display_last_14_days_shared_diagnoses_per_covid_case_es = \
format_shared_diagnoses_per_covid_case(value=last_14_days_summary["shared_diagnoses_per_covid_case_es"])
status = textwrap.dedent(f"""
#RadarCOVID – {extraction_date_with_hour}
Today{today_addendum}:
- Uploaded TEKs: {shared_teks_by_upload_date:.0f} ({shared_teks_by_upload_date_last_hour:+d} last hour)
- Shared Diagnoses: ≤{shared_diagnoses:.0f}
- Usage Ratio: {display_shared_diagnoses_per_covid_case}
Last 14 Days:
- Usage Ratio (Estimation): {display_last_14_days_shared_diagnoses_per_covid_case}
- Usage Ratio (Official): {display_last_14_days_shared_diagnoses_per_covid_case_es}
Info: {github_project_base_url}#documentation
""")
status = status.encode(encoding="utf-8")
api.update_status(status=status, media_ids=media_ids)
```
| github_jupyter |
# Fuzzingbook Release Notes
This book and its code use numbered versioning. The version numbers correspond to the version numbers in [the Python pip package](Importing.ipynb).
## Version 1.1 (in progress)
* We are adding type annotations to all code.
* We are adding more material (videos, quizzes) to the book.
* Apart from code fixes, code semantics stays unchanged.
## Version 1.0.7 (released 22-01-25)
* The `WebFuzzer` constructor now allows to use a subclass of `HTMLGrammarMiner`.
* The `GUIFuzzer` constructor now allows to use a subclass of `GUIGrammarMiner`.
* Class diagrams are now simplified, as we skip non-public methods defined in other chapters.
## Version 1.0.6 (released 2022-01-18)
* Fixed a major performance bug in grammar fuzzing (Issue [#117](https://github.com/uds-se/fuzzingbook/pull/117))
* We now work with recent versions of `z3-solver` (Issue [#115](https://github.com/uds-se/fuzzingbook/issues/115))
* In the [chapter on configuration fuzzing](ConfigurationFuzzer.ipynb), the `OptionGrammarMiner` will now capture args from external Python scripts that are protected by `if __name__ == '__main__'`
* Various minor fixes and documentation improvements
## Version 1.0.5 (released 2022-01-10)
* This release increases compatibility with various Z3 versions for [concolic fuzzing](ConcolicFuzzer.ipynb).
* In the [chapter on symbolic fuzzing](SymbolicFuzzer.ipynb), the `AdvancedSymbolicFuzzer` is now named `SymbolicFuzzer` plain and simple. (`AdvancedSymbolicFuzzer` still works as an alias).
## Version 1.0.4 (released 2022-01-04)
Happy new year!
* This release fixes some bugs in the [chapter on concolic fuzzing](ConcolicFuzzer.ipynb), notably in the `ConcolicTracer.zeval()` method.
## Version 1.0.3 (released 2021-12-14)
* On Unix, the [`ExpectTimeout`](ExpectError.ipynb) class is now much more performant
* The chapter on [greybox fuzzing with grammars](GreyboxGrammarFuzzer.ipynb) should now run on Windows, too
## Version 1.0.2 (released 2021-12-08)
* Minor fixes in the `AFLGoSchedule` and `AFLFastSchedule` classes in the [chapter on greybox fuzzing](GreyboxFuzzer.ipynb).
* Minor fixes across the board.
## Version 1.0.1 (released 2021-11-23)
* The code now passes `mypy` static type checks.
* The `Coverage` class now supports `function_names()` and `__repr__()` methods. Its `__exit__()` method is no longer included in coverage.
* Minor fixes across the board.
## Version 1.0 (released 2021-11-04)
* We now support (but also require) **Python 3.9 or later**. Earlier versions still required Python 3.6 due to some outdated modules such as `astor` and `enforce` we depended upon (and now don't anymore).
* We added missing dependencies to the `fuzzingbook` pip package ([Issue #44](https://github.com/uds-se/debuggingbook/issues/44) in `debuggingbook`) such that `pip install fuzzingbook` also installs all the packages it depends upon. Thanks to @TheSilvus for reporting this!
* We fixed a warning '.gitignore is a symbolic link' during git checkout ([Issue #43](https://github.com/uds-se/debuggingbook/issues/43)) Thanks to @rjc for reporting this!
* We identified some chapters that were using `numpy.random` rather than Python `random`, resulting in, well, random results every time we'd build the book. This is now fixed, and more consistent.
* Under the hood, we have adopted several more improvements from our sister project ["The Debugging Book"](https://www.debuggingbook.org). Notably, the build process is much streamlined, and we run [continuous integration tests](https://github.com/uds-se/fuzzingbook/actions) to ensure quality of changes and pull requests.
## Version 0.9.5 (released 2021-06-08)
* Lots of minor fixes in HTML generation, adopting new tools and tests from ["The Debugging Book"](https://www.debuggingbook.org).
* Code functionality should be unchanged.
* The `bookutils` module is now shared with the `debuggingbook` project; some (hopefully neutral) fixes.
* Several typos and other minor fixes throughout the book.
## Version 0.9.0
* In the Web version, some not-so-critical details (typically, long implementations and logs) are only shown on demand. This is still work in progress.
* The `fuzzingbook_utils` module used by notebooks is now renamed to `bookutils`. Code and notebooks using `fuzzingbook_utils` may still work, but will issue a deprecation warning.
* Several minor fixes to functionality in [Parsing and Recombining Inputs](Parser.ipynb), [Concolic Fuzzing](ConcolicFuzzer.ipynb), [Symbolic Fuzzing](SymbolicFuzzer.ipynb)
* Better style when printing from browser (colored text, smaller fonts)
* Avoid tracking in YouTube videos
* Several typos and other minor fixes throughout the book
## Version 0.8.0 (released 2019-05-21)
First numbered fuzzingbook release.
* Includes [Python pip package](Importing.ipynb).
* Includes _Synopsis_ sections at the beginning of each chapter, highlighting their usage in own code.
* Describes [Tours through the Book](Tours.ipynb).
## Chapter Releases
Before switching to numbered releases, new chapters were coming out every Tuesday.
1. [Introduction to Software Testing](Intro_Testing.ipynb) – 2018-10-30
1. [Fuzzing: Breaking Things with Random Inputs](Fuzzer.ipynb) – 2018-10-30
1. [Getting Coverage](Coverage.ipynb) – 2018-11-06
1. [Mutation-Based Fuzzing](MutationFuzzer.ipynb) – 2018-11-06
1. [Fuzzing with Grammars](Grammars.ipynb) – 2018-11-13
1. [Efficient Grammar Fuzzing](GrammarFuzzer.ipynb) – 2018-11-20
1. [Grammar Coverage](GrammarCoverageFuzzer.ipynb) – 2018-11-27
1. [Testing Configurations](ConfigurationFuzzer.ipynb) – 2018-12-04
1. [Parsing and Recombining Inputs](Parser.ipynb) – 2018-12-11
1. [Probabilistic Grammar Fuzzing](ProbabilisticGrammarFuzzer.ipynb) – 2018-12-18
1. [Fuzzing with Generators](GeneratorGrammarFuzzer.ipynb) – 2019-01-08
1. [Fuzzing APIs](APIFuzzer.ipynb) – 2019-01-15
1. [Carving Unit Tests](Carver.ipynb) – 2019-01-22
1. [Reducing Failure-Inducing Inputs](Reducer.ipynb) – 2019-01-29
1. [Web Testing](WebFuzzer.ipynb) – 2019-02-05
1. [GUI Testing](GUIFuzzer.ipynb) – 2019-02-12
1. [Mining Input Grammars](GrammarMiner.ipynb) – 2019-02-19
1. [Tracking Information Flow](InformationFlow.ipynb) – 2019-03-05
1. [Concolic Fuzzing](ConcolicFuzzer.ipynb) – 2019-03-12
1. [Symbolic Fuzzing](SymbolicFuzzer.ipynb) – 2019-03-19
1. [Mining Function Specifications](DynamicInvariants) – 2019-03-26
1. [Search-Based Fuzzing](SearchBasedFuzzer.ipynb) – 2019-04-02
1. [Evaluating Test Effectiveness with Mutation Analysis](MutationAnalysis.ipynb) – 2019-04-09
1. [Greybox Fuzzing](GreyboxFuzzer.ipynb) – 2019-04-16
1. [Greybox Fuzzing with Grammars](GreyboxGrammarFuzzer.ipynb) – 2019-04-30
1. [Fuzzing in the Large](FuzzingInTheLarge.ipynb) – 2019-05-07
1. [When to Stop Fuzzing](WhenToStopFuzzing.ipynb) – 2019-05-14
1. [Tours through the Book](Tours.ipynb) - 2019-05-21
After all chapters were out, we switched to a release-based schedule, with numbered minor and major releases coming out when they are ready.
| github_jupyter |
# Spark Learning Note - Table Joins
Jia Geng | gjia0214@gmail.com
```
# check java version
# use sudo update-alternatives --config java to switch java version if needed.
!java -version
from pyspark.sql.session import SparkSession
spark = SparkSession.builder.appName('TableJoinExamples').getOrCreate()
spark
```
## Join Operations
Join operation: `left.xJoin(right)` is to bring left and right tables together by one or more keys. Common join types include:
- **inner joins**: keep rows with from both tables keys exist in both left&right table
- **outer joins**: keep rows from both tables with keys exist in either left or right table
- **left outer joins**: keep rows from both tables with keys exist in left table
- **right outer joins**: keep rows from both tables with keys exist in the right table
- **left semi joins**: keep rows from only left tables with keys exist in the left&right table. Result only contain columns from left table.
- **left anti joins**: keep rows from only left tables with keys NOT exist in the right table, Result only contain columns from left table.
- **natural joins**: join by matching the columns between two datasets with the same names
- be careful when using this as column name could be the same but indicating different things!!
- **cross joins**: match every row in the left table with every row in the right table
In spark:
```
df_left.join(df_left, on=, how=)
```
When working with array column, there are many array based operations under `pyspark.sql.functions` that can be used for join expression.
To deal with duplicate col name between two DF.
- use col name as the expression input instead of boolean
- use this when the two columns from left and right tables are referring the same
- drop a column after the join
- rename a column before the join
```
# set up the example
person_data = [(0, 'Bill', 0, [100]), (1, 'Matt', 1, [500, 250, 100]), (2, 'Zack', 1, [250, 100])]
person_coln = ['id', 'name', 'graduate_program', 'person_status']
person = spark.createDataFrame(person_data, person_coln)
program_data = [(0, 'MS', 'ACE', 'UC'), (2, 'MS', 'ECCS', 'UC'), (1, 'PhD', 'ECCS', 'UC')]
program_coln = ['id', 'degree', 'department', 'school']
program = spark.createDataFrame(program_data, program_coln)
status_data = [(500, 'VP'), (250, 'PMC Member'), (100, 'Contributor')]
status_coln = ['id', 'status']
statusDF = spark.createDataFrame(status_data, status_coln)
person.show()
program.show()
statusDF.show()
# inner join person and program on program id
# it kept the id column from both table
# id from person is the person id
# id from program is the program id
# they are different!
person.join(program, person['graduate_program'] == program['id'], 'inner').show()
person.join(program, person['graduate_program'] != program['id'], 'inner').show()
# outer join person and program on program id
# if id can not be matched, missing side will be null
person.join(program, person['graduate_program'] == program['id'], 'outer').show()
# left/right outer join person and program on program id
# only keep the rows from the left/right table
person.join(program, person['graduate_program'] == program['id'], 'left_outer').show()
person.join(program, person['graduate_program'] == program['id'], 'right_outer').show()
# left semi joins - only include shared key rows
# DOES NOT include anything from right table
# right table only provide the keys for 'filtering' the rows
program.join(person, person['graduate_program'] == program['id'], 'left_semi').show()
# left anti joins - exlcude shared key rows
# DOES NOT include anything from right table
# right table only provide the keys for 'filtering' the rows
program.join(person, person['graduate_program'] == program['id'], 'left_anti').show()
from pyspark.sql.functions import array_contains, expr
# some more complex example that including the array operatoins
# here, must use expr(), if use arrary_contains(), it does not work
# because array_contain() takes a column and a value
# here use expr(), we are telling to program to
# find the person_status col from the two tables
# find the id col from the two tables
# ambiguoty is not allowed!
expression = expr('array_contains(person_status, id)')
# must rename the person id to avoid ambiguoty
person.withColumnRenamed('id', 'personID').join(statusDF, expression).show()
```
| github_jupyter |
# Deep Neural Network in TensorFlow
In this notebook, we convert our [intermediate-depth MNIST-classifying neural network](https://github.com/the-deep-learners/TensorFlow-LiveLessons/blob/master/notebooks/intermediate_net_in_keras.ipynb) from Keras to TensorFlow (compare them side by side) following Aymeric Damien's [Multi-Layer Perceptron Notebook](https://github.com/aymericdamien/TensorFlow-Examples/blob/master/notebooks/3_NeuralNetworks/multilayer_perceptron.ipynb) style.
Subsequently, we add a layer to make it deep!
#### Load dependencies
```
import numpy as np
np.random.seed(42)
import tensorflow as tf
tf.set_random_seed(42)
```
#### Load data
```
from tensorflow.examples.tutorials.mnist import input_data
mnist = input_data.read_data_sets("MNIST_data/", one_hot=True)
```
#### Set neural network hyperparameters (tidier at top of file!)
```
lr = 0.1
epochs = 1
batch_size = 128
weight_initializer = tf.contrib.layers.xavier_initializer()
```
#### Set number of neurons for each layer
```
n_input = 784
n_dense_1 = 64
n_dense_2 = 64
n_classes = 10
```
#### Define placeholders Tensors for inputs and labels
```
x = tf.placeholder(tf.float32, [None, n_input])
y = tf.placeholder(tf.float32, [None, n_classes])
```
#### Define types of layers
```
# dense layer with ReLU activation:
def dense(x, W, b):
# DEFINE
# DEFINE
return a
```
#### Define dictionaries for storing weights and biases for each layer -- and initialize
```
bias_dict = {
'b1': tf.Variable(tf.zeros([n_dense_1])),
'b2': tf.Variable(tf.zeros([n_dense_2])),
'b_out': tf.Variable(tf.zeros([n_classes]))
}
weight_dict = {
'W1': tf.get_variable('W1', [n_input, n_dense_1], initializer=weight_initializer),
'W2': tf.get_variable('W2', [n_dense_1, n_dense_2], initializer=weight_initializer),
'W_out': tf.get_variable('W_out', [n_dense_2, n_classes], initializer=weight_initializer)
}
```
#### Design neural network architecture
```
def network(x, weights, biases):
# DEFINE
return out_layer_z
```
#### Build model
```
predictions = network(x, weights=weight_dict, biases=bias_dict)
```
#### Define model's loss and its optimizer
```
cost = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(logits=predictions, labels=y))
optimizer = tf.train.GradientDescentOptimizer(learning_rate=lr).minimize(cost)
```
#### Define evaluation metrics
```
# calculate accuracy by identifying test cases where the model's highest-probability class matches the true y label:
correct_prediction = tf.equal(tf.argmax(predictions, 1), tf.argmax(y, 1))
accuracy_pct = tf.reduce_mean(tf.cast(correct_prediction, tf.float32)) * 100
```
#### Create op for variable initialization
```
initializer_op = tf.global_variables_initializer()
```
#### Train the network in a session
```
with tf.Session() as session:
session.run(initializer_op)
print("Training for", epochs, "epochs.")
# loop over epochs:
for epoch in range(epochs):
avg_cost = 0.0 # track cost to monitor performance during training
avg_accuracy_pct = 0.0
# loop over all batches of the epoch:
n_batches = int(mnist.train.num_examples / batch_size)
for i in range(n_batches):
batch_x, batch_y = mnist.train.next_batch(batch_size)
# feed batch data to run optimization and fetching cost and accuracy:
_, batch_cost, batch_acc = session.run([optimizer, cost, accuracy_pct],
feed_dict={x: batch_x, y: batch_y})
# accumulate mean loss and accuracy over epoch:
avg_cost += batch_cost / n_batches
avg_accuracy_pct += batch_acc / n_batches
# output logs at end of each epoch of training:
print("Epoch ", '%03d' % (epoch+1),
": cost = ", '{:.3f}'.format(avg_cost),
", accuracy = ", '{:.2f}'.format(avg_accuracy_pct), "%",
sep='')
print("Training Complete. Testing Model.\n")
test_cost = cost.eval({x: mnist.test.images, y: mnist.test.labels})
test_accuracy_pct = accuracy_pct.eval({x: mnist.test.images, y: mnist.test.labels})
print("Test Cost:", '{:.3f}'.format(test_cost))
print("Test Accuracy: ", '{:.2f}'.format(test_accuracy_pct), "%", sep='')
```
| github_jupyter |
# Relative Humidity Regression Prediction from Air Quality UCI Dataset - Ganesh Ram Gururajan
## Data import, cleaning and Visualization
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
data = pd.read_csv('C:/Users/ganes/Data Science/Datasets/AirQualityUCI.csv')
data.head()
data.drop(['Date','Time'],axis=1,inplace=True)
sns.heatmap(data.corr())
data.columns
data.drop(['T','AH','PT08.S5(O3)','PT08.S2(NMHC)'],axis=1,inplace=True)
sns.heatmap(data.corr())
data.drop(['PT08.S4(NO2)','C6H6(GT)'],axis=1,inplace=True)
sns.heatmap(data.corr())
data.drop(['NOx(GT)','NO2(GT)'],axis=1,inplace=True)
sns.heatmap(data.corr())
data.head(10)
data.columns
data.shape
data.dtypes
```
## Train Test Split
```
from sklearn.model_selection import train_test_split
X = data.drop(['RH'],axis = 1)
y = data['RH']
X_train,X_test,y_train,y_test = train_test_split(X,y,test_size=0.3,random_state=100)
X_train.shape,X_test.shape,y_train.shape,y_test.shape
```
## Machine Learning
## Decision Tree Regression
```
from sklearn.tree import DecisionTreeRegressor
dt = DecisionTreeRegressor()
dt.fit(X_train,y_train)
pred1 = dt.predict(X_test)
```
## Decision Tree Evaluation
```
from sklearn.metrics import mean_absolute_error,mean_squared_error,r2_score
print(mean_absolute_error(y_test,pred1))
print(np.sqrt(mean_squared_error(y_test,pred1)))
print(r2_score(y_test,pred1))
```
## Random Forest Regressor
```
from sklearn.ensemble import RandomForestRegressor
rf = RandomForestRegressor(n_estimators=100)
rf.fit(X_train,y_train)
pred2 = rf.predict(X_test)
```
## Random Forest Evaluation
```
print(mean_absolute_error(y_test,pred2))
print(np.sqrt(mean_squared_error(y_test,pred2)))
print(r2_score(y_test,pred2))
```
## AdaBoost
```
from sklearn.ensemble import AdaBoostRegressor
ada = AdaBoostRegressor()
ada.fit(X_train,y_train)
pred3 = ada.predict(X_test)
```
## AdaBoost Evaluation
```
print(mean_absolute_error(y_test,pred3))
print(np.sqrt(mean_squared_error(y_test,pred3)))
print(r2_score(y_test,pred3))
```
## XGBoost
```
from xgboost import XGBRegressor
xgb = XGBRegressor()
xgb.fit(X_train,y_train)
pred4 = xgb.predict(X_test)
```
## XGBoost Evaluation
```
print(mean_absolute_error(y_test,pred4))
print(np.sqrt(mean_squared_error(y_test,pred4)))
print(r2_score(y_test,pred4))
```
## Linear Regression
```
from sklearn.linear_model import LinearRegression
lr = LinearRegression()
lr.fit(X_train,y_train)
pred5 = lr.predict(X_test)
```
## Linear Regression Prediction
```
print(mean_absolute_error(y_test,pred5))
print(np.sqrt(mean_squared_error(y_test,pred5)))
print(r2_score(y_test,pred5))
```
## Ridge Cross Validation Regression
```
from sklearn.linear_model import RidgeCV
rcv = RidgeCV()
rcv.fit(X_train,y_train)
pred6 = rcv.predict(X_test)
```
## RidgeCV Evaluation
```
print(mean_absolute_error(y_test,pred6))
print(np.sqrt(mean_squared_error(y_test,pred6)))
print(r2_score(y_test,pred6))
```
## Lars Cross Validation Regression
```
from sklearn.linear_model import LarsCV
lrcv = LarsCV()
lrcv.fit(X_train,y_train)
pred7 = lrcv.predict(X_test)
```
## LarsCV Evaluation
```
print(mean_absolute_error(y_test,pred7))
print(np.sqrt(mean_squared_error(y_test,pred7)))
print(r2_score(y_test,pred7))
```
# End
| github_jupyter |
```
import os.path
import pandas as pd
import numpy as np
import xgboost as xgb
base_path = "../input/"
# load the data
navigation = pd.read_csv(os.path.join(base_path, 'navigation.csv'))
sales = pd.read_csv(os.path.join(base_path, 'sales.csv'))
train = pd.read_csv(os.path.join(base_path, 'train.csv'))
test = pd.read_csv(os.path.join(base_path, 'test.csv'))
vimages = pd.read_csv(os.path.join(base_path, 'vimages.csv'))
sub = pd.read_csv(os.path.join(base_path, 'sample_submission.csv'))
# leave-one-out target encoding for different colors of the same product
product_descriptor = ['product_type', 'product_gender', 'macro_function',
'function', 'sub_function', 'model', 'aesthetic_sub_line', 'macro_material',
'month']
product_target_sum = train.groupby(product_descriptor)['target'].sum().reset_index(name = 'sum_target')
product_target_count = train.groupby(product_descriptor)['target'].count().reset_index(name = 'count_target')
product_target_stats = pd.merge(product_target_sum, product_target_count, on = product_descriptor)
train = train.merge(product_target_stats, on = product_descriptor, how = 'left')
test = test.merge(product_target_stats, on = product_descriptor, how = 'left')
train['mean_target'] = (train['sum_target'] - train['target'])/(train['count_target']-1)
test['mean_target'] = (test['sum_target'])/(test['count_target'])
train.drop(['sum_target','count_target'],axis=1,inplace=True)
test.drop(['sum_target','count_target'],axis=1,inplace=True)
# counts for categorical features in train+test
count_vec_cols = ['macro_function', 'function', 'sub_function', 'model',
'aesthetic_sub_line', 'macro_material', 'color']
for col in count_vec_cols:
tmp = pd.DataFrame({'sku_hash': pd.concat([train['sku_hash'],test['sku_hash']]),
col:pd.concat([train[col],test[col]])})
tmp = pd.DataFrame(tmp.groupby(col)['sku_hash'].count()).reset_index()
tmp.columns = [col,col+'_count']
train = train.merge(tmp, on = col, how = 'left')
test = test.merge(tmp, on = col, how = 'left')
# sum of page views by different traffic source
traffic_source_views = navigation.groupby(['sku_hash','traffic_source'])['page_views'].sum().reset_index()
traffic_source_views = traffic_source_views.pivot(index='sku_hash', columns='traffic_source', values='page_views').reset_index()
traffic_source_views.columns = ['sku_hash',
'page_views_nav1', 'page_views_nav2', 'page_views_nav3',
'page_views_nav4', 'page_views_nav5', 'page_views_nav6']
# sum of sales by different type
type_sales = sales.groupby(['sku_hash','type'])['sales_quantity'].sum().reset_index()
type_sales = type_sales.pivot(index='sku_hash', columns='type', values='sales_quantity').reset_index()
type_sales.columns = ['sku_hash', 'sales_quantity_type1', 'sales_quantity_type2']
# sum of sales by different zone
zone_sales = sales.groupby(['sku_hash','zone_number'])['sales_quantity'].sum().reset_index()
zone_sales = zone_sales.pivot(index='sku_hash', columns='zone_number', values='sales_quantity').reset_index()
zone_sales.columns = ['sku_hash',
'sales_quantity_zone1', 'sales_quantity_zone2', 'sales_quantity_zone3',
'sales_quantity_zone4', 'sales_quantity_zone5']
# overall stats of sales, page views and twitter sentiments
navigation_stats = navigation.groupby('sku_hash')['page_views'].sum().reset_index(name='page_views')
sales_stats = sales.groupby('sku_hash')['sales_quantity','TotalBuzzPost', 'TotalBuzz',
'NetSentiment', 'PositiveSentiment', 'NegativeSentiment', 'Impressions'].sum().reset_index()
# define cross validation splits
train['idx'] = pd.Categorical(train.sku_hash).codes
train['idx'] = train['idx'] % 5
# merge everything for train
X = train.copy()
X = X.merge(navigation_stats, on = 'sku_hash', how = 'left')
X = X.merge(sales_stats, on = 'sku_hash', how = 'left')
X = X.merge(traffic_source_views, on = 'sku_hash', how = 'left')
X = X.merge(type_sales, on = 'sku_hash', how = 'left')
X = X.merge(zone_sales, on = 'sku_hash', how = 'left')
X.loc[X.product_type=='Accessories','product_type'] = '0'
X.loc[X.product_type=='Leather Goods','product_type'] = '1'
X.product_type = X.product_type.astype(int)
X.loc[X.product_gender=='Women','product_gender'] = '-1'
X.loc[X.product_gender=='Unisex','product_gender'] = '0'
X.loc[X.product_gender=='Men','product_gender'] = '1'
X.product_gender = X.product_gender.astype(int)
# transform label to meet the metric
X['y'] = np.log(X['target'] + 1)
# merge everything for test
Z = test.copy()
Z = Z.merge(navigation_stats, on = 'sku_hash', how = 'left')
Z = Z.merge(sales_stats, on = 'sku_hash', how = 'left')
Z = Z.merge(traffic_source_views, on = 'sku_hash', how = 'left')
Z = Z.merge(type_sales, on = 'sku_hash', how = 'left')
Z = Z.merge(zone_sales, on = 'sku_hash', how = 'left')
Z.loc[Z.product_type=='Accessories','product_type'] = '0'
Z.loc[Z.product_type=='Leather Goods','product_type'] = '1'
Z.product_type = Z.product_type.astype(int)
Z.loc[Z.product_gender=='Women','product_gender'] = '-1'
Z.loc[Z.product_gender=='Unisex','product_gender'] = '0'
Z.loc[Z.product_gender=='Men','product_gender'] = '1'
Z.product_gender = Z.product_gender.astype(int)
features = ['product_type', 'product_gender',
'page_views', 'sales_quantity',
'TotalBuzzPost', 'TotalBuzz', 'NetSentiment', 'PositiveSentiment', 'NegativeSentiment', 'Impressions',
'fr_FR_price',
'macro_function_count', 'function_count', 'sub_function_count', 'model_count', 'aesthetic_sub_line_count', 'macro_material_count', 'color_count',
'page_views_nav1', 'page_views_nav2', 'page_views_nav3', 'page_views_nav4', 'page_views_nav5', 'page_views_nav6',
'sales_quantity_type1', 'sales_quantity_type2',
'sales_quantity_zone1','sales_quantity_zone2','sales_quantity_zone3', 'sales_quantity_zone4','sales_quantity_zone5',
'mean_target',]
# define function to generate xgboost objects for a specific month
def train_test_split(tr, te, mo, feats, num_folds):
Xtrain = []
ytrain = []
dtrain = []
Xval = []
yval = []
dval = []
for i in range(num_folds):
Xtrain.append(tr.loc[(tr.month == mo) & (tr.idx != i), feats].values)
ytrain.append(tr.loc[(tr.month == mo) & (tr.idx != i), 'y'].values)
dtrain.append(xgb.DMatrix(Xtrain[i],ytrain[i]))
Xval.append(tr.loc[(tr.month == mo) & (tr.idx == i), feats].values)
yval.append(tr.loc[(tr.month == mo) & (tr.idx == i), 'y'].values)
dval.append(xgb.DMatrix(Xval[i],yval[i]))
Xtest = te.loc[(te.month == mo),feats].values
dtest = xgb.DMatrix(Xtest)
return dtrain, dval, dtest
# define xgboost parameters to use in models
param = {}
param['objective'] = 'reg:linear'
param['eval_metric'] = 'rmse'
param['booster'] = 'gbtree'
param['eta'] = 0.025
param['subsample'] = 0.7
param['colsample_bytree'] = 0.7
param['num_parallel_tree'] = 3
param['min_child_weight'] = 25
param['gamma'] = 5
param['max_depth'] = 3
param['silent'] = 1
# train models for the 1 month
dtrain, dval, dtest = train_test_split(tr = X, te = Z, mo = 1, feats = features, num_folds = 5)
model_m1 = []
for i in range(5):
model_m1.append(
xgb.train(
param,
dtrain[i],
50000,
[(dtrain[i],'train'), (dval[i],'eval')],
early_stopping_rounds = 200,
verbose_eval = False)
)
# run predictions for the 1 month
oof_m1 = []
oof_test_m1 = []
for i in range(5):
oof_m1.append(model_m1[i].predict(dval[i]))
oof_test_m1.append(model_m1[i].predict(dtest))
test_m1 = np.mean(oof_test_m1, axis=0)
m1 = {}
for i in range(5):
m1 = {**m1, **dict(zip(X.loc[(X.month==1) & (X.idx==i),'sku_hash'], oof_m1[i]))}
m1 = {**m1, **dict(zip(Z.loc[(Z.month==1),'sku_hash'], test_m1))}
oof_m1 = pd.DataFrame.from_dict(m1, orient='index').reset_index()
oof_m1.columns = ['sku_hash', 'oof_m1']
X2 = pd.merge(X.copy(), oof_m1, on = 'sku_hash')
Z2 = pd.merge(Z.copy(), oof_m1, on = 'sku_hash')
features2 = features + ['oof_m1']
# train models for the 2 month
dtrain2, dval2, dtest2 = train_test_split(tr = X2, te = Z2, mo = 2, feats = features2, num_folds = 5)
model_m2 = []
for i in range(5):
model_m2.append(
xgb.train(
param,
dtrain2[i],
50000,
[(dtrain2[i],'train'), (dval2[i],'eval')],
early_stopping_rounds = 200,
verbose_eval = False)
)
# run predictions for the 2 month
oof_m2 = []
oof_test_m2 = []
for i in range(5):
oof_m2.append(model_m2[i].predict(dval2[i]))
oof_test_m2.append(model_m2[i].predict(dtest2))
test_m2 = np.mean(oof_test_m2, axis=0)
m2 = {}
for i in range(5):
m2 = {**m2, **dict(zip(X.loc[(X.month==2) & (X.idx==i),'sku_hash'], oof_m2[i]))}
m2 = {**m2, **dict(zip(Z.loc[(Z.month==2),'sku_hash'], test_m2))}
oof_m2 = pd.DataFrame.from_dict(m2, orient='index').reset_index()
oof_m2.columns = ['sku_hash', 'oof_m2']
X3 = pd.merge(X2.copy(), oof_m2, on = 'sku_hash')
Z3 = pd.merge(Z2.copy(), oof_m2, on = 'sku_hash')
features3 = features2 + ['oof_m2']
# train models for the 3 month
dtrain3, dval3, dtest3 = train_test_split(tr = X3, te = Z3, mo = 3, feats = features3, num_folds = 5)
model_m3 = []
for i in range(5):
model_m3.append(
xgb.train(
param,
dtrain3[i],
50000,
[(dtrain3[i],'train'),(dval3[i],'eval')],
early_stopping_rounds = 200,
verbose_eval = False)
)
# run predictions for the 3 month
oof_m3 = []
oof_test_m3 = []
for i in range(5):
oof_m3.append(model_m3[i].predict(dval3[i]))
oof_test_m3.append(model_m3[i].predict(dtest3))
test_m3 = np.mean(oof_test_m3, axis=0)
m3 = {}
for i in range(5):
m3 = {**m3, **dict(zip(X.loc[(X.month==3) & (X.idx==i),'sku_hash'], oof_m3[i]))}
m3 = {**m3, **dict(zip(Z.loc[(Z.month==3),'sku_hash'], test_m3))}
oof_m3 = pd.DataFrame.from_dict(m3, orient='index').reset_index()
oof_m3.columns = ['sku_hash', 'oof_m3']
X3 = pd.merge(X3.copy(), oof_m3, on = 'sku_hash')
Z3 = pd.merge(Z3.copy(), oof_m3, on = 'sku_hash')
# create a single vector of predictions for both train and test
Z3['target'] = 0
Z3.loc[Z3.month == 1, 'target'] = Z3.loc[Z3.month == 1, 'oof_m1']
Z3.loc[Z3.month == 2, 'target'] = Z3.loc[Z3.month == 2, 'oof_m2']
Z3.loc[Z3.month == 3, 'target'] = Z3.loc[Z3.month == 3, 'oof_m3']
X3['pred_target'] = 0
X3.loc[X3.month == 1, 'pred_target'] = X3.loc[X3.month == 1, 'oof_m1']
X3.loc[X3.month == 2, 'pred_target'] = X3.loc[X3.month == 2, 'oof_m2']
X3.loc[X3.month == 3, 'pred_target'] = X3.loc[X3.month == 3, 'oof_m3']
# some cross validation diagnostics
print(f"month1: {np.sqrt(np.mean((X3.loc[X3.month==1,'y'] - X3.loc[X3.month==1,'pred_target'])**2))}")
print(f"month2: {np.sqrt(np.mean((X3.loc[X3.month==2,'y'] - X3.loc[X3.month==2,'pred_target'])**2))}")
print(f"month3: {np.sqrt(np.mean((X3.loc[X3.month==3,'y'] - X3.loc[X3.month==3,'pred_target'])**2))}")
print(f"overall: {np.sqrt(np.mean((X3['y'] - X3['pred_target'])**2))}")
# make a submission
Z3['target'] = np.exp(Z3.target)-1
final_sub = Z3[['ID','target']]
final_sub.to_csv(os.path.join(base_path,'silly-raddar-sub4.csv'),index=None)
# create a oof train version of a submision
X3['target'] = np.exp(X3.pred_target)-1
cv_sub = X3[['ID','target']]
cv_sub.to_csv(os.path.join(base_path,'silly-raddar-cv4.csv'),index=None)
# export features for anokas to use
X3[['ID']+features].to_csv(os.path.join(base_path,'raddar-features-train.csv'),index=None)
Z3[['ID']+features].to_csv(os.path.join(base_path,'raddar-features-test.csv'),index=None)
```
| github_jupyter |
```
# @title Copyright & License (click to expand)
# Copyright 2021 Google LLC
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Vertex AI Model Monitoring
<table align="left">
<td>
<a href="https://console.cloud.google.com/ai-platform/notebooks/deploy-notebook?name=Model%20Monitoring&download_url=https%3A%2F%2Fraw.githubusercontent.com%2FGoogleCloudPlatform%2Fai-platform-samples%2Fmaster%2Fai-platform-unified%2Fnotebooks%2Fofficial%2Fmodel_monitoring%2Fmodel_monitoring.ipynb"
<img src="https://cloud.google.com/images/products/ai/ai-solutions-icon.svg" alt="AI Platform Notebooks"> Open in GCP Notebooks
</a>
</td>
<td>
<a href="https://colab.research.google.com/github/GoogleCloudPlatform/ai-platform-samples/blob/master/ai-platform-unified/notebooks/official/model_monitoring/model_monitoring.ipynb">
<img src="https://cloud.google.com/ml-engine/images/colab-logo-32px.png" alt="Colab logo"> Open in Colab
</a>
</td>
<td>
<a href="https://github.com/GoogleCloudPlatform/ai-platform-samples/blob/master/ai-platform-unified/notebooks/official/model_monitoring/model_monitoring.ipynb">
<img src="https://cloud.google.com/ml-engine/images/github-logo-32px.png" alt="GitHub logo">
View on GitHub
</a>
</td>
</table>
## Overview
### What is Model Monitoring?
Modern applications rely on a well established set of capabilities to monitor the health of their services. Examples include:
* software versioning
* rigorous deployment processes
* event logging
* alerting/notication of situations requiring intervention
* on-demand and automated diagnostic tracing
* automated performance and functional testing
You should be able to manage your ML services with the same degree of power and flexibility with which you can manage your applications. That's what MLOps is all about - managing ML services with the best practices Google and the broader computing industry have learned from generations of experience deploying well engineered, reliable, and scalable services.
Model monitoring is only one piece of the ML Ops puzzle - it helps answer the following questions:
* How well do recent service requests match the training data used to build your model? This is called **training-serving skew**.
* How significantly are service requests evolving over time? This is called **drift detection**.
If production traffic differs from training data, or varies substantially over time, that's likely to impact the quality of the answers your model produces. When that happens, you'd like to be alerted automatically and responsively, so that **you can anticipate problems before they affect your customer experiences or your revenue streams**.
### Objective
In this notebook, you will learn how to...
* deploy a pre-trained model
* configure model monitoring
* generate some artificial traffic
* understand how to interpret the statistics, visualizations, other data reported by the model monitoring feature
### Costs
This tutorial uses billable components of Google Cloud:
* Vertext AI
* BigQuery
Learn about [Vertext AI
pricing](https://cloud.google.com/vertex-ai/pricing) and [Cloud Storage
pricing](https://cloud.google.com/storage/pricing), and use the [Pricing
Calculator](https://cloud.google.com/products/calculator/)
to generate a cost estimate based on your projected usage.
### The example model
The model you'll use in this notebook is based on [this blog post](https://cloud.google.com/blog/topics/developers-practitioners/churn-prediction-game-developers-using-google-analytics-4-ga4-and-bigquery-ml). The idea behind this model is that your company has extensive log data describing how your game users have interacted with the site. The raw data contains the following categories of information:
- identity - unique player identitity numbers
- demographic features - information about the player, such as the geographic region in which a player is located
- behavioral features - counts of the number of times a player has triggered certain game events, such as reaching a new level
- churn propensity - this is the label or target feature, it provides an estimated probability that this player will churn, i.e. stop being an active player.
The blog article referenced above explains how to use BigQuery to store the raw data, pre-process it for use in machine learning, and train a model. Because this notebook focuses on model monitoring, rather than training models, you're going to reuse a pre-trained version of this model, which has been exported to Google Cloud Storage. In the next section, you will setup your environment and import this model into your own project.
## Before you begin
### Setup your dependencies
```
import os
import sys
import IPython
assert sys.version_info.major == 3, "This notebook requires Python 3."
# Install Python package dependencies.
print("Installing TensorFlow 2.4.1 and TensorFlow Data Validation (TFDV)")
!pip3 install -q numpy
!pip3 install -q tensorflow==2.4.1 tensorflow_data_validation[visualization]
!pip3 install -q --upgrade google-api-python-client google-auth-oauthlib google-auth-httplib2 oauth2client requests
!pip3 install -q google-cloud-aiplatform
!pip3 install -q --upgrade google-cloud-storage==1.32.0
# Automatically restart kernel after installing new packages.
if not os.getenv("IS_TESTING"):
print("Restarting kernel...")
app = IPython.Application.instance()
app.kernel.do_shutdown(True)
print("Done.")
import os
import random
import sys
import time
# Import required packages.
import numpy as np
```
### Set up your Google Cloud project
**The following steps are required, regardless of your notebook environment.**
1. [Select or create a Google Cloud project](https://console.cloud.google.com/cloud-resource-manager). When you first create an account, you get a $300 free credit towards your compute/storage costs.
1. [Make sure that billing is enabled for your project](https://cloud.google.com/billing/docs/how-to/modify-project).
1. If you are running this notebook locally, you will need to install the [Cloud SDK](https://cloud.google.com/sdk).
1. You'll use the *gcloud* command throughout this notebook. In the following cell, enter your project name and run the cell to authenticate yourself with the Google Cloud and initialize your *gcloud* configuration settings.
**For this lab, we're going to use region us-central1 for all our resources (BigQuery training data, Cloud Storage bucket, model and endpoint locations, etc.). Those resources can be deployed in other regions, as long as they're consistently co-located, but we're going to use one fixed region to keep things as simple and error free as possible.**
```
PROJECT_ID = "[your-project-id]" # @param {type:"string"}
REGION = "us-central1"
SUFFIX = "aiplatform.googleapis.com"
API_ENDPOINT = f"{REGION}-{SUFFIX}"
PREDICT_API_ENDPOINT = f"{REGION}-prediction-{SUFFIX}"
if os.getenv("IS_TESTING"):
!gcloud --quiet components install beta
!gcloud --quiet components update
!gcloud config set project $PROJECT_ID
!gcloud config set ai/region $REGION
```
### Login to your Google Cloud account and enable AI services
```
# If on AI Platform, then don't execute this code
if not os.path.exists("/opt/deeplearning/metadata/env_version"):
if "google.colab" in sys.modules:
from google.colab import auth as google_auth
google_auth.authenticate_user()
# If you are running this notebook locally, replace the string below with the
# path to your service account key and run this cell to authenticate your GCP
# account.
elif not os.getenv("IS_TESTING"):
%env GOOGLE_APPLICATION_CREDENTIALS ''
!gcloud services enable aiplatform.googleapis.com
```
### Define some helper functions
Run the following cell to define some utility functions used throughout this notebook. Although these functions are not critical to understand the main concepts, feel free to expand the cell if you're curious or want to dive deeper into how some of your API requests are made.
```
# @title Utility functions
import copy
import os
from google.cloud.aiplatform_v1beta1.services.endpoint_service import \
EndpointServiceClient
from google.cloud.aiplatform_v1beta1.services.job_service import \
JobServiceClient
from google.cloud.aiplatform_v1beta1.services.prediction_service import \
PredictionServiceClient
from google.cloud.aiplatform_v1beta1.types.io import BigQuerySource
from google.cloud.aiplatform_v1beta1.types.model_deployment_monitoring_job import (
ModelDeploymentMonitoringJob, ModelDeploymentMonitoringObjectiveConfig,
ModelDeploymentMonitoringScheduleConfig)
from google.cloud.aiplatform_v1beta1.types.model_monitoring import (
ModelMonitoringAlertConfig, ModelMonitoringObjectiveConfig,
SamplingStrategy, ThresholdConfig)
from google.cloud.aiplatform_v1beta1.types.prediction_service import \
PredictRequest
from google.protobuf import json_format
from google.protobuf.duration_pb2 import Duration
from google.protobuf.struct_pb2 import Value
DEFAULT_THRESHOLD_VALUE = 0.001
def create_monitoring_job(objective_configs):
# Create sampling configuration.
random_sampling = SamplingStrategy.RandomSampleConfig(sample_rate=LOG_SAMPLE_RATE)
sampling_config = SamplingStrategy(random_sample_config=random_sampling)
# Create schedule configuration.
duration = Duration(seconds=MONITOR_INTERVAL)
schedule_config = ModelDeploymentMonitoringScheduleConfig(monitor_interval=duration)
# Create alerting configuration.
emails = [USER_EMAIL]
email_config = ModelMonitoringAlertConfig.EmailAlertConfig(user_emails=emails)
alerting_config = ModelMonitoringAlertConfig(email_alert_config=email_config)
# Create the monitoring job.
endpoint = f"projects/{PROJECT_ID}/locations/{REGION}/endpoints/{ENDPOINT_ID}"
predict_schema = ""
analysis_schema = ""
job = ModelDeploymentMonitoringJob(
display_name=JOB_NAME,
endpoint=endpoint,
model_deployment_monitoring_objective_configs=objective_configs,
logging_sampling_strategy=sampling_config,
model_deployment_monitoring_schedule_config=schedule_config,
model_monitoring_alert_config=alerting_config,
predict_instance_schema_uri=predict_schema,
analysis_instance_schema_uri=analysis_schema,
)
options = dict(api_endpoint=API_ENDPOINT)
client = JobServiceClient(client_options=options)
parent = f"projects/{PROJECT_ID}/locations/{REGION}"
response = client.create_model_deployment_monitoring_job(
parent=parent, model_deployment_monitoring_job=job
)
print("Created monitoring job:")
print(response)
return response
def get_thresholds(default_thresholds, custom_thresholds):
thresholds = {}
default_threshold = ThresholdConfig(value=DEFAULT_THRESHOLD_VALUE)
for feature in default_thresholds.split(","):
feature = feature.strip()
thresholds[feature] = default_threshold
for custom_threshold in custom_thresholds.split(","):
pair = custom_threshold.split(":")
if len(pair) != 2:
print(f"Invalid custom skew threshold: {custom_threshold}")
return
feature, value = pair
thresholds[feature] = ThresholdConfig(value=float(value))
return thresholds
def get_deployed_model_ids(endpoint_id):
client_options = dict(api_endpoint=API_ENDPOINT)
client = EndpointServiceClient(client_options=client_options)
parent = f"projects/{PROJECT_ID}/locations/{REGION}"
response = client.get_endpoint(name=f"{parent}/endpoints/{endpoint_id}")
model_ids = []
for model in response.deployed_models:
model_ids.append(model.id)
return model_ids
def set_objectives(model_ids, objective_template):
# Use the same objective config for all models.
objective_configs = []
for model_id in model_ids:
objective_config = copy.deepcopy(objective_template)
objective_config.deployed_model_id = model_id
objective_configs.append(objective_config)
return objective_configs
def send_predict_request(endpoint, input):
client_options = {"api_endpoint": PREDICT_API_ENDPOINT}
client = PredictionServiceClient(client_options=client_options)
params = {}
params = json_format.ParseDict(params, Value())
request = PredictRequest(endpoint=endpoint, parameters=params)
inputs = [json_format.ParseDict(input, Value())]
request.instances.extend(inputs)
response = client.predict(request)
return response
def list_monitoring_jobs():
client_options = dict(api_endpoint=API_ENDPOINT)
parent = f"projects/{PROJECT_ID}/locations/us-central1"
client = JobServiceClient(client_options=client_options)
response = client.list_model_deployment_monitoring_jobs(parent=parent)
print(response)
def pause_monitoring_job(job):
client_options = dict(api_endpoint=API_ENDPOINT)
client = JobServiceClient(client_options=client_options)
response = client.pause_model_deployment_monitoring_job(name=job)
print(response)
def delete_monitoring_job(job):
client_options = dict(api_endpoint=API_ENDPOINT)
client = JobServiceClient(client_options=client_options)
response = client.delete_model_deployment_monitoring_job(name=job)
print(response)
# Sampling distributions for categorical features...
DAYOFWEEK = {1: 1040, 2: 1223, 3: 1352, 4: 1217, 5: 1078, 6: 1011, 7: 1110}
LANGUAGE = {
"en-us": 4807,
"en-gb": 678,
"ja-jp": 419,
"en-au": 310,
"en-ca": 299,
"de-de": 147,
"en-in": 130,
"en": 127,
"fr-fr": 94,
"pt-br": 81,
"es-us": 65,
"zh-tw": 64,
"zh-hans-cn": 55,
"es-mx": 53,
"nl-nl": 37,
"fr-ca": 34,
"en-za": 29,
"vi-vn": 29,
"en-nz": 29,
"es-es": 25,
}
OS = {"IOS": 3980, "ANDROID": 3798, "null": 253}
MONTH = {6: 3125, 7: 1838, 8: 1276, 9: 1718, 10: 74}
COUNTRY = {
"United States": 4395,
"India": 486,
"Japan": 450,
"Canada": 354,
"Australia": 327,
"United Kingdom": 303,
"Germany": 144,
"Mexico": 102,
"France": 97,
"Brazil": 93,
"Taiwan": 72,
"China": 65,
"Saudi Arabia": 49,
"Pakistan": 48,
"Egypt": 46,
"Netherlands": 45,
"Vietnam": 42,
"Philippines": 39,
"South Africa": 38,
}
# Means and standard deviations for numerical features...
MEAN_SD = {
"julianday": (204.6, 34.7),
"cnt_user_engagement": (30.8, 53.2),
"cnt_level_start_quickplay": (7.8, 28.9),
"cnt_level_end_quickplay": (5.0, 16.4),
"cnt_level_complete_quickplay": (2.1, 9.9),
"cnt_level_reset_quickplay": (2.0, 19.6),
"cnt_post_score": (4.9, 13.8),
"cnt_spend_virtual_currency": (0.4, 1.8),
"cnt_ad_reward": (0.1, 0.6),
"cnt_challenge_a_friend": (0.0, 0.3),
"cnt_completed_5_levels": (0.1, 0.4),
"cnt_use_extra_steps": (0.4, 1.7),
}
DEFAULT_INPUT = {
"cnt_ad_reward": 0,
"cnt_challenge_a_friend": 0,
"cnt_completed_5_levels": 1,
"cnt_level_complete_quickplay": 3,
"cnt_level_end_quickplay": 5,
"cnt_level_reset_quickplay": 2,
"cnt_level_start_quickplay": 6,
"cnt_post_score": 34,
"cnt_spend_virtual_currency": 0,
"cnt_use_extra_steps": 0,
"cnt_user_engagement": 120,
"country": "Denmark",
"dayofweek": 3,
"julianday": 254,
"language": "da-dk",
"month": 9,
"operating_system": "IOS",
"user_pseudo_id": "104B0770BAE16E8B53DF330C95881893",
}
```
## Import your model
The churn propensity model you'll be using in this notebook has been trained in BigQuery ML and exported to a Google Cloud Storage bucket. This illustrates how you can easily export a trained model and move a model from one cloud service to another.
Run the next cell to import this model into your project. **If you've already imported your model, you can skip this step.**
```
MODEL_NAME = "churn"
IMAGE = "us-docker.pkg.dev/cloud-aiplatform/prediction/tf2-cpu.2-4:latest"
ARTIFACT = "gs://mco-mm/churn"
output = !gcloud --quiet beta ai models upload --container-image-uri=$IMAGE --artifact-uri=$ARTIFACT --display-name=$MODEL_NAME --format="value(model)"
print("model output: ", output)
MODEL_ID = output[1].split("/")[-1]
print(f"Model {MODEL_NAME}/{MODEL_ID} created.")
```
## Deploy your endpoint
Now that you've imported your model into your project, you need to create an endpoint to serve your model. An endpoint can be thought of as a channel through which your model provides prediction services. Once established, you'll be able to make prediction requests on your model via the public internet. Your endpoint is also serverless, in the sense that Google ensures high availability by reducing single points of failure, and scalability by dynamically allocating resources to meet the demand for your service. In this way, you are able to focus on your model quality, and freed from adminstrative and infrastructure concerns.
Run the next cell to deploy your model to an endpoint. **This will take about ten minutes to complete. If you've already deployed a model to an endpoint, you can reuse your endpoint by running the cell after the next one.**
```
ENDPOINT_NAME = "churn"
output = !gcloud --quiet beta ai endpoints create --display-name=$ENDPOINT_NAME --format="value(name)"
print("endpoint output: ", output)
ENDPOINT = output[-1]
ENDPOINT_ID = ENDPOINT.split("/")[-1]
output = !gcloud --quiet beta ai endpoints deploy-model $ENDPOINT_ID --display-name=$ENDPOINT_NAME --model=$MODEL_ID --traffic-split="0=100"
DEPLOYED_MODEL_ID = output[1].split()[-1][:-1]
print(
f"Model {MODEL_NAME}/{MODEL_ID}/{DEPLOYED_MODEL_ID} deployed to Endpoint {ENDPOINT_NAME}/{ENDPOINT_ID}/{ENDPOINT}."
)
# @title Run this cell only if you want to reuse an existing endpoint.
if not os.getenv("IS_TESTING"):
ENDPOINT_ID = "" # @param {type:"string"}
ENDPOINT = f"projects/mco-mm/locations/us-central1/endpoints/{ENDPOINT_ID}"
```
## Run a prediction test
Now that you have imported a model and deployed that model to an endpoint, you are ready to verify that it's working. Run the next cell to send a test prediction request. If everything works as expected, you should receive a response encoded in a text representation called JSON.
**Try this now by running the next cell and examine the results.**
```
import pprint as pp
print(ENDPOINT)
print("request:")
pp.pprint(DEFAULT_INPUT)
try:
resp = send_predict_request(ENDPOINT, DEFAULT_INPUT)
print("response")
pp.pprint(resp)
except Exception:
print("prediction request failed")
```
Taking a closer look at the results, we see the following elements:
- **churned_values** - a set of possible values (0 and 1) for the target field
- **churned_probs** - a corresponding set of probabilities for each possible target field value (5x10^-40 and 1.0, respectively)
- **predicted_churn** - based on the probabilities, the predicted value of the target field (1)
This response encodes the model's prediction in a format that is readily digestible by software, which makes this service ideal for automated use by an application.
## Start your monitoring job
Now that you've created an endpoint to serve prediction requests on your model, you're ready to start a monitoring job to keep an eye on model quality and to alert you if and when input begins to deviate in way that may impact your model's prediction quality.
In this section, you will configure and create a model monitoring job based on the churn propensity model you imported from BigQuery ML.
### Configure the following fields:
1. Log sample rate - Your prediction requests and responses are logged to BigQuery tables, which are automatically created when you create a monitoring job. This parameter specifies the desired logging frequency for those tables.
1. Monitor interval - the time window over which to analyze your data and report anomalies. The minimum window is one hour (3600 seconds).
1. Target field - the prediction target column name in training dataset.
1. Skew detection threshold - the skew threshold for each feature you want to monitor.
1. Prediction drift threshold - the drift threshold for each feature you want to monitor.
```
USER_EMAIL = "" # @param {type:"string"}
JOB_NAME = "churn"
# Sampling rate (optional, default=.8)
LOG_SAMPLE_RATE = 0.8 # @param {type:"number"}
# Monitoring Interval in seconds (optional, default=3600).
MONITOR_INTERVAL = 3600 # @param {type:"number"}
# URI to training dataset.
DATASET_BQ_URI = "bq://mco-mm.bqmlga4.train" # @param {type:"string"}
# Prediction target column name in training dataset.
TARGET = "churned"
# Skew and drift thresholds.
SKEW_DEFAULT_THRESHOLDS = "country,language" # @param {type:"string"}
SKEW_CUSTOM_THRESHOLDS = "cnt_user_engagement:.5" # @param {type:"string"}
DRIFT_DEFAULT_THRESHOLDS = "country,language" # @param {type:"string"}
DRIFT_CUSTOM_THRESHOLDS = "cnt_user_engagement:.5" # @param {type:"string"}
```
### Create your monitoring job
The following code uses the Google Python client library to translate your configuration settings into a programmatic request to start a model monitoring job. Instantiating a monitoring job can take some time. If everything looks good with your request, you'll get a successful API response. Then, you'll need to check your email to receive a notification that the job is running.
```
skew_thresholds = get_thresholds(SKEW_DEFAULT_THRESHOLDS, SKEW_CUSTOM_THRESHOLDS)
drift_thresholds = get_thresholds(DRIFT_DEFAULT_THRESHOLDS, DRIFT_CUSTOM_THRESHOLDS)
skew_config = ModelMonitoringObjectiveConfig.TrainingPredictionSkewDetectionConfig(
skew_thresholds=skew_thresholds
)
drift_config = ModelMonitoringObjectiveConfig.PredictionDriftDetectionConfig(
drift_thresholds=drift_thresholds
)
training_dataset = ModelMonitoringObjectiveConfig.TrainingDataset(target_field=TARGET)
training_dataset.bigquery_source = BigQuerySource(input_uri=DATASET_BQ_URI)
objective_config = ModelMonitoringObjectiveConfig(
training_dataset=training_dataset,
training_prediction_skew_detection_config=skew_config,
prediction_drift_detection_config=drift_config,
)
model_ids = get_deployed_model_ids(ENDPOINT_ID)
objective_template = ModelDeploymentMonitoringObjectiveConfig(
objective_config=objective_config
)
objective_configs = set_objectives(model_ids, objective_template)
monitoring_job = create_monitoring_job(objective_configs)
# Run a prediction request to generate schema, if necessary.
try:
_ = send_predict_request(ENDPOINT, DEFAULT_INPUT)
print("prediction succeeded")
except Exception:
print("prediction failed")
```
After a minute or two, you should receive email at the address you configured above for USER_EMAIL. This email confirms successful deployment of your monitoring job. Here's a sample of what this email might look like:
<br>
<br>
<img src="https://storage.googleapis.com/mco-general/img/mm6.png" />
<br>
As your monitoring job collects data, measurements are stored in Google Cloud Storage and you are free to examine your data at any time. The circled path in the image above specifies the location of your measurements in Google Cloud Storage. Run the following cell to take a look at your measurements in Cloud Storage.
```
!gsutil ls gs://cloud-ai-platform-fdfb4810-148b-4c86-903c-dbdff879f6e1/*/*
```
You will notice the following components in these Cloud Storage paths:
- **cloud-ai-platform-..** - This is a bucket created for you and assigned to capture your service's prediction data. Each monitoring job you create will trigger creation of a new folder in this bucket.
- **[model_monitoring|instance_schemas]/job-..** - This is your unique monitoring job number, which you can see above in both the response to your job creation requesst and the email notification.
- **instance_schemas/job-../analysis** - This is the monitoring jobs understanding and encoding of your training data's schema (field names, types, etc.).
- **instance_schemas/job-../predict** - This is the first prediction made to your model after the current monitoring job was enabled.
- **model_monitoring/job-../serving** - This folder is used to record data relevant to drift calculations. It contains measurement summaries for every hour your model serves traffic.
- **model_monitoring/job-../training** - This folder is used to record data relevant to training-serving skew calculations. It contains an ongoing summary of prediction data relative to training data.
### You can create monitoring jobs with other user interfaces
In the previous cells, you created a monitoring job using the Python client library. You can also use the *gcloud* command line tool to create a model monitoring job and, in the near future, you will be able to use the Cloud Console, as well for this function.
## Generate test data to trigger alerting
Now you are ready to test the monitoring function. Run the following cell, which will generate fabricated test predictions designed to trigger the thresholds you specified above. It takes about five minutes to run this cell and at least an hour to assess and report anamolies in skew or drift so after running this cell, feel free to proceed with the notebook and you'll see how to examine the resulting alert later.
```
def random_uid():
digits = [str(i) for i in range(10)] + ["A", "B", "C", "D", "E", "F"]
return "".join(random.choices(digits, k=32))
def monitoring_test(count, sleep, perturb_num={}, perturb_cat={}):
# Use random sampling and mean/sd with gaussian distribution to model
# training data. Then modify sampling distros for two categorical features
# and mean/sd for two numerical features.
mean_sd = MEAN_SD.copy()
country = COUNTRY.copy()
for k, (mean_fn, sd_fn) in perturb_num.items():
orig_mean, orig_sd = MEAN_SD[k]
mean_sd[k] = (mean_fn(orig_mean), sd_fn(orig_sd))
for k, v in perturb_cat.items():
country[k] = v
for i in range(0, count):
input = DEFAULT_INPUT.copy()
input["user_pseudo_id"] = str(random_uid())
input["country"] = random.choices([*country], list(country.values()))[0]
input["dayofweek"] = random.choices([*DAYOFWEEK], list(DAYOFWEEK.values()))[0]
input["language"] = str(random.choices([*LANGUAGE], list(LANGUAGE.values()))[0])
input["operating_system"] = str(random.choices([*OS], list(OS.values()))[0])
input["month"] = random.choices([*MONTH], list(MONTH.values()))[0]
for key, (mean, sd) in mean_sd.items():
sample_val = round(float(np.random.normal(mean, sd, 1)))
val = max(sample_val, 0)
input[key] = val
print(f"Sending prediction {i}")
try:
send_predict_request(ENDPOINT, input)
except Exception:
print("prediction request failed")
time.sleep(sleep)
print("Test Completed.")
test_time = 300
tests_per_sec = 1
sleep_time = 1 / tests_per_sec
iterations = test_time * tests_per_sec
perturb_num = {"cnt_user_engagement": (lambda x: x * 3, lambda x: x / 3)}
perturb_cat = {"Japan": max(COUNTRY.values()) * 2}
monitoring_test(iterations, sleep_time, perturb_num, perturb_cat)
```
## Interpret your results
While waiting for your results, which, as noted, may take up to an hour, you can read ahead to get sense of the alerting experience.
### Here's what a sample email alert looks like...
<img src="https://storage.googleapis.com/mco-general/img/mm7.png" />
This email is warning you that the *cnt_user_engagement*, *country* and *language* feature values seen in production have skewed above your threshold between training and serving your model. It's also telling you that the *cnt_user_engagement* feature value is drifting significantly over time, again, as per your threshold specification.
### Monitoring results in the Cloud Console
You can examine your model monitoring data from the Cloud Console. Below is a screenshot of those capabilities.
#### Monitoring Status
<img src="https://storage.googleapis.com/mco-general/img/mm1.png" />
#### Monitoring Alerts
<img src="https://storage.googleapis.com/mco-general/img/mm2.png" />
## Clean up
To clean up all Google Cloud resources used in this project, you can [delete the Google Cloud
project](https://cloud.google.com/resource-manager/docs/creating-managing-projects#shutting_down_projects) you used for the tutorial.
Otherwise, you can delete the individual resources you created in this tutorial:
```
# Delete endpoint resource
!gcloud ai endpoints delete $ENDPOINT_NAME --quiet
# Delete model resource
!gcloud ai models delete $MODEL_NAME --quiet
```
## Learn more about model monitoring
**Congratulations!** You've now learned what model monitoring is, how to configure and enable it, and how to find and interpret the results. Check out the following resources to learn more about model monitoring and ML Ops.
- [TensorFlow Data Validation](https://www.tensorflow.org/tfx/guide/tfdv)
- [Data Understanding, Validation, and Monitoring At Scale](https://blog.tensorflow.org/2018/09/introducing-tensorflow-data-validation.html)
- [Vertex Product Documentation](https://cloud.google.com/vertex)
- [Model Monitoring Reference Docs](https://cloud.google.com/vertex/docs/reference)
- [Model Monitoring blog article]()
| github_jupyter |
## UAP Generate & Evaluate (whitebox)
attacks
- sgd-uap (untargeted) eps = 2-12
dataset
- cifar10
- svhn
model_name
- resnet18
- [soft-pruning]
- resnet18_sfP
- resnet18_sfP-mixup
- resnet18_sfP-cutout
- [post-training pruning]
- resnet18_PX_0.Y
- X = 2, 3, 4
- Y = 3, 6, 9
```
import numpy as np
import pandas as pd
import os
import torch
import json
from utils_attack import uap_batch
from utils_model import *
dir_results = './results/'
dir_uap = './uaps/'
nb_epoch = 10
atk_name = 'sgd-uap'
y_target = -1
model_names = ['resnet18', 'resnet18_sfP', 'resnet18_sfP-mixup', 'resnet18_sfP-cutout']
for layer in [2, 3, 4]:
for prune_pct in ['0.3', '0.6', '0.9']:
model_names.append('resnet18_P%i-%s' % (layer, prune_pct))
model_names
def eval_uap(uap):
top, top_probs, top1acc, top5acc, outputs, labels = evaluate(model, testloader, uap = uap)
print('Top 1 accuracy', sum(top1acc) / len(labels))
print('Top 5 accuracy', sum(top5acc) / len(labels))
dict_entry = {}
dict_entry['model_name'] = model_name
dict_entry['dataset'] = dataset
dict_entry['atk_name'] = atk_name
dict_entry['atk_param'] = y_target
dict_entry['eps'] = eps
if atk_name[:3] == 'sgd': dict_entry['nb_epoch'] = nb_epoch
dict_entry['top1acc'] = sum(top1acc) / len(labels)
dict_entry['top5acc'] = sum(top5acc) / len(labels)
dict_entry['UER'] = 1 - sum(top1acc) / len(labels)
print('UER ', dict_entry['UER'])
print('\n')
if y_target >= 0: dict_entry['tgt_success'] = sum(outputs == y_target) / len(labels)
# Output distribution
for i in range(10):
dict_entry['label_dist%i' % i] = sum(outputs == i) / len(labels)
all_results.append(dict_entry)
```
```
all_results = []
dataset = 'svhn'
batch_size = 1627
testloader = get_testdata(dataset, batch_size)
for model_name in model_names:
model = get_model(model_name, dataset, batch_size)
model.eval()
for eps in range(2, 13, 2):
uap_save_path = dir_uap + dataset + '/' + model_name
print(uap_save_path + '/sgd-eps%i.pth' % eps)
uap = torch.load(uap_save_path + '/sgd-eps%i.pth' % eps)
eval_uap(uap)
pd.DataFrame(all_results).to_csv(dir_results + 'eval_whitebox-%s.csv' % dataset)
```
## View Evaluation Results
```
df = pd.read_csv(dir_results + 'eval_whitebox-svhn.csv', index_col = 0)
df[(df['dataset'] == 'svhn') & (df['eps'] == 8)].sort_values(by = 'UER')
df = pd.read_csv(dir_results + 'eval_whitebox-cifar10.csv', index_col = 0)
df[(df['dataset'] == 'cifar10') & (df['eps'] == 8)].sort_values(by = 'UER')
```
| github_jupyter |
```
from qiskit import IBMQ, Aer, transpile, schedule as build_schedule
from qiskit.transpiler import PassManager
from qiskit.transpiler.passes.calibration import RZXCalibrationBuilderNoEcho
from qiskit_nature.drivers import Molecule
from qiskit_nature.drivers.second_quantization import ElectronicStructureDriverType, ElectronicStructureMoleculeDriver
from qiskit_nature.problems.second_quantization import ElectronicStructureProblem
from qiskit_nature.converters.second_quantization import QubitConverter
from qiskit_nature.transformers.second_quantization.electronic import FreezeCoreTransformer
from qiskit_nature.operators.second_quantization import FermionicOp
from qiskit_nature.mappers.second_quantization import ParityMapper
from qiskit_nature.algorithms import GroundStateEigensolver
from qiskit.algorithms import NumPyMinimumEigensolver, VQE
from qiskit.algorithms.optimizers import SPSA
from qiskit.circuit import QuantumCircuit, ParameterVector
from qiskit.utils import QuantumInstance
import matplotlib.pyplot as plt
import numpy as np
IBMQ.load_account()
provider = IBMQ.get_provider()
lagos = provider.get_backend('ibm_lagos')
def HEA_naive(num_q, depth):
circuit = QuantumCircuit(num_q)
params = ParameterVector("theta", length=num_q * (3 * depth + 2))
counter = 0
for q in range(num_q):
circuit.rx(params[counter], q)
counter += 1
circuit.rz(params[counter], q)
counter += 1
for d in range(depth):
for q in range(num_q - 1):
circuit.cx(q, q + 1)
for q in range(num_q):
circuit.rz(params[counter], q)
counter += 1
circuit.rx(params[counter], q)
counter += 1
circuit.rz(params[counter], q)
counter += 1
return circuit, params
def HEA_aware(num_q, depth, hardware):
circuit = QuantumCircuit(num_q)
params = ParameterVector("theta", length=num_q * (3 * depth + 2))
counter = 0
for q in range(num_q):
circuit.rx(params[counter], q)
counter += 1
circuit.rz(params[counter], q)
counter += 1
for d in range(depth):
for q in range(num_q - 1):
gate = QuantumCircuit(num_q)
gate.rzx(np.pi/2, q, q + 1)
pass_ = RZXCalibrationBuilderNoEcho(hardware)
qc_cr = PassManager(pass_).run(gate)
circuit.compose(qc_cr, inplace=True)
for q in range(num_q):
circuit.rz(params[counter], q)
counter += 1
circuit.rx(params[counter], q)
counter += 1
circuit.rz(params[counter], q)
counter += 1
return circuit, params
depth = 1
qubits = 2
back = lagos
circuit, _ = HEA_naive(qubits, depth)
t = transpile(circuit, back)
schedule = build_schedule(t, back)
schedule.draw()
circuit, _ = HEA_aware(qubits, depth, back)
t = transpile(circuit, back)
schedule = build_schedule(t, back)
schedule.draw()
qubit_converter = QubitConverter(ParityMapper(), two_qubit_reduction=True)
numpy_solver = NumPyMinimumEigensolver()
spsa = SPSA(100)
qi = QuantumInstance(Aer.get_backend('aer_simulator'))
vqe_circuit = VQE(ansatz=circuit, quantum_instance=qi, optimizer=spsa)
# H2 simulation
total_dist = 4
dist = 0.1
incr_early = 0.1
incr_late = 0.3
real_energies = []
vqe_energies = []
dists = []
while dist < total_dist:
print(dist, total_dist)
molecule = Molecule(geometry=[['H', [0., 0., 0.]], ['H', [0., 0., dist]]])
driver = ElectronicStructureMoleculeDriver(molecule, basis='sto3g', \
driver_type=ElectronicStructureDriverType.PYSCF)
es_problem = ElectronicStructureProblem(driver)
second_q_ops = es_problem.second_q_ops()
if dist == 0.1:
FermionicOp.set_truncation(0)
print(second_q_ops[0])
FermionicOp.set_truncation(1)
print(qubit_converter.convert(second_q_ops[0], num_particles=es_problem.num_particles))
print(es_problem.grouped_property)
calc = GroundStateEigensolver(qubit_converter, numpy_solver)
res = calc.solve(es_problem)
real_energies.append(np.real(res.total_energies[0]))
calc = GroundStateEigensolver(qubit_converter, vqe_circuit)
res = calc.solve(es_problem)
vqe_energies.append(np.real(res.total_energies[0]))
dists.append(dist)
if dist > total_dist / 2:
dist += incr_late
else:
dist += incr_early
plt.plot(dists, real_energies, label='Real', color='red')
plt.scatter(dists, vqe_energies, label='VQE', color='black')
plt.title("H2")
plt.ylim(-1.2, 0.4)
plt.xlim(0, 4)
plt.xlabel('Angstroms')
plt.ylabel('Hartree')
plt.legend()
plt.show()
depth = 2
qubits = 4
circuit, _ = HEA_naive(qubits, depth)
spsa = SPSA(100)
qi = QuantumInstance(Aer.get_backend('aer_simulator'))
vqe_circuit = VQE(ansatz=circuit, quantum_instance=qi, optimizer=spsa)
#print(circuit)
total_dist = 5
dist = 0.3
incr_early = 0.1
incr_late = 0.3
real_energies = []
vqe_energies = []
dists = []
while dist < total_dist:
print(dist, total_dist)
molecule = Molecule(geometry=[['Li', [0., 0., 0.]], ['H', [0., 0., dist]]])
driver = ElectronicStructureMoleculeDriver(molecule, basis='sto3g', \
driver_type=ElectronicStructureDriverType.PYSCF)
transformer = FreezeCoreTransformer(freeze_core=True, remove_orbitals=[3, 4])
es_problem = ElectronicStructureProblem(driver, transformers=[transformer])
second_q_ops = es_problem.second_q_ops()
if dist == 0.3:
print(second_q_ops[0])
print(qubit_converter.convert(second_q_ops[0], num_particles=es_problem.num_particles))
print(es_problem.grouped_property)
print(es_problem.grouped_property_transformed)
calc = GroundStateEigensolver(qubit_converter, numpy_solver)
res = calc.solve(es_problem)
real_energies.append(np.real(res.total_energies[0]))
calc = GroundStateEigensolver(qubit_converter, vqe_circuit) #VQEClient()
res = calc.solve(es_problem)
vqe_energies.append(np.real(res.total_energies[0]))
dists.append(dist)
if dist > total_dist / 2:
dist += incr_late
else:
dist += incr_early
plt.plot(dists, real_energies, label='Real', color='green')
plt.scatter(dists, vqe_energies, label='VQE', color='black')
plt.title("LiH")
plt.ylim(-8, -6.6)
plt.xlim(0.3, 5)
plt.xlabel('Angstroms')
plt.ylabel('Hartree')
plt.legend()
plt.show()
depth = 2
qubits = 6
circuit, _ = HEA_naive(qubits, depth)
spsa = SPSA(100)
qi = QuantumInstance(Aer.get_backend('aer_simulator'))
vqe_circuit = VQE(ansatz=circuit, quantum_instance=qi, optimizer=spsa)
#print(circuit)
total_dist = 5
dist = 0.4
incr_early = 0.2
incr_late = 0.4
real_energies = []
vqe_energies = []
dists = []
while dist < total_dist:
print(dist, total_dist)
molecule = Molecule(geometry=[['H', [0., 0., -dist]], ['Be', [0., 0., 0.]], ['H', [0., 0., dist]]])
driver = ElectronicStructureMoleculeDriver(molecule, basis='sto3g', \
driver_type=ElectronicStructureDriverType.PYSCF)
transformer = FreezeCoreTransformer(freeze_core=True, remove_orbitals=[4, 5])
es_problem = ElectronicStructureProblem(driver, transformers=[transformer])
second_q_ops = es_problem.second_q_ops()
if dist == 0.4:
print(second_q_ops[0])
print(qubit_converter.convert(second_q_ops[0], num_particles=es_problem.num_particles))
print(es_problem.grouped_property)
print(es_problem.grouped_property_transformed)
calc = GroundStateEigensolver(qubit_converter, numpy_solver)
res = calc.solve(es_problem)
real_energies.append(np.real(res.total_energies[0]))
calc = GroundStateEigensolver(qubit_converter, vqe_circuit)
res = calc.solve(es_problem)
vqe_energies.append(np.real(res.total_energies[0]))
dists.append(dist)
if dist > total_dist / 2:
dist += incr_late
else:
dist += incr_early
plt.plot(dists, real_energies, label='Real', color='blue')
plt.scatter(dists, vqe_energies, label='VQE', color='black')
plt.title("BeH2")
plt.ylim(-15.7, -12.0)
plt.xlim(0.48, 5)
plt.xlabel('Angstroms')
plt.ylabel('Hartree')
plt.legend()
plt.show()
from qiskit.tools.jupyter import *
%qiskit_version_table
```
| github_jupyter |
# Evaluation Methodology of Federated Learning
In this notebook we study the different evaluation methodologies that we can use when we want to evaluate FL problems. First, we set up the FL configuration (for more information see [Basic Concepts Notebook](./federated_learning_basic_concepts.ipynb)).
```
import shfl
import tensorflow as tf
import numpy as np
import random
random.seed(123)
np.random.seed(seed=123)
class Reshape(shfl.private.FederatedTransformation):
def apply(self, labeled_data):
labeled_data.data = np.reshape(labeled_data.data, (labeled_data.data.shape[0], labeled_data.data.shape[1], labeled_data.data.shape[2],1))
class Normalize(shfl.private.FederatedTransformation):
def __init__(self, mean, std):
self.__mean = mean
self.__std = std
def apply(self, labeled_data):
labeled_data.data = (labeled_data.data - self.__mean)/self.__std
def model_builder():
model = tf.keras.models.Sequential()
model.add(tf.keras.layers.Conv2D(32, kernel_size=(3, 3), padding='same', activation='relu', strides=1, input_shape=(28, 28, 1)))
model.add(tf.keras.layers.MaxPooling2D(pool_size=2, strides=2, padding='valid'))
model.add(tf.keras.layers.Dropout(0.4))
model.add(tf.keras.layers.Conv2D(32, kernel_size=(3, 3), padding='same', activation='relu', strides=1))
model.add(tf.keras.layers.MaxPooling2D(pool_size=2, strides=2, padding='valid'))
model.add(tf.keras.layers.Dropout(0.3))
model.add(tf.keras.layers.Flatten())
model.add(tf.keras.layers.Dense(128, activation='relu'))
model.add(tf.keras.layers.Dropout(0.1))
model.add(tf.keras.layers.Dense(64, activation='relu'))
model.add(tf.keras.layers.Dense(10, activation='softmax'))
model.compile(optimizer="rmsprop", loss="categorical_crossentropy", metrics=["accuracy"])
return shfl.model.DeepLearningModel(model)
#Read data
database = shfl.data_base.Emnist()
train_data, train_labels, test_data, test_labels = database.load_data()
#Distribute among clients
non_iid_distribution = shfl.data_distribution.NonIidDataDistribution(database)
federated_data, test_data, test_labels = non_iid_distribution.get_federated_data(num_nodes=5, percent=10)
#Set up aggregation operator
aggregator = shfl.federated_aggregator.FedAvgAggregator()
federated_government = shfl.federated_government.FederatedGovernment(model_builder, federated_data, aggregator)
#Reshape and normalize
shfl.private.federated_operation.apply_federated_transformation(federated_data, Reshape())
mean = np.mean(test_data.data)
std = np.std(test_data.data)
shfl.private.federated_operation.apply_federated_transformation(federated_data, Normalize(mean, std))
```
## Evaluation methodology 1: Global test dataset
The first evaluation methodology that we propose consists of the federated version of the classical evaluation methods. For that purpose, we use a common test dataset allocated in the server. We show the evaluation metrics (loss and accuracy in this case) in each round of learning both in local models and global updated model. We show the behaviour of this evaluation methodology as follows.
```
test_data = np.reshape(test_data, (test_data.shape[0], test_data.shape[1], test_data.shape[2],1))
federated_government.run_rounds(1, test_data, test_labels)
```
This methodology is the simplest and shows both local and global models. The problem with this methodology is that the local evaluation metrics are biased by the distribution of test set data. That is, the performance of the local models is not properly represented when using a non-iid scenario (see [Federated Sampling](./federated_learning_sampling.ipynb)) because the distribution of training data for each client is different from the data we test on. For that reason, we propose the following evaluation methodology.
## Evaluation methodology 2: Global test dataset and local test datasets
In this evaluation methodology we consider that there is, as in the previous one, a global test dataset and that each client has a local test dataset according to the distribution of their training data. Hence, in each round we show the evaluation metrics of each client on their local and the global test. This evaluation methodology is more complete as it shows the performance of the local FL models in the global and local distribution of the data, which gives as more information.
First, we split each client's data in train and test partitions. You can find this method in [Federated Operation](https://github.com/sherpaai/Sherpa.ai-Federated-Learning-Framework/blob/master/shfl/private/federated_operation.py).
```
shfl.private.federated_operation.split_train_test(federated_data)
```
After that, each client owns a training set which uses for training the local learning model and a test set which uses to evaluate.
We are now ready to show the behaviour of this evaluation methodology.
```
#We restart federated goverment
federated_government = shfl.federated_government.FederatedGovernment(model_builder, federated_data, aggregator)
test_data = np.reshape(test_data, (test_data.shape[0], test_data.shape[1], test_data.shape[2],1))
federated_government.run_rounds(1, test_data, test_labels)
```
We appreciate the significance of this new evaluation methodology in the output produced. For example, the first client performed the worst in the global test while it was the best in its local test. This indicates that the data distribution of this client is most likely very poor compared to the global data distribution, for example, consisting of only two classes. This produces a really good local learning model in just one round of learning, being a simpler problem but with a very low global test performance.
This highlights the strenght of using specific evaluation methodologies in FL, especially when the distribution of data among clients follows a non-IID distribution (see [Federated Sampling](./federated_learning_sampling.ipynb)).
| github_jupyter |
# Random Forest
* * *

* * *
Drawing by Phil Cutler.
## Introduction
Any tutorial on [Random Forests](https://en.wikipedia.org/wiki/Random_forest) (RF) should also include a review of decision trees, as these are models that are ensembled together to create the Random Forest model -- or put another way, the "trees that comprise the forest." Much of the complexity and detail of the Random Forest algorithm occurs within the individual decision trees and therefore it's important to understand decision trees to understand the RF algorithm as a whole. Therefore, before proceeding, it is recommended that you read through the accompanying [Classification and Regression Trees Tutorial](decision-trees.ipynb).
## History
The Random Forest algorithm is preceded by the [Random Subspace Method](https://en.wikipedia.org/wiki/Random_subspace_method) (aka "attribute bagging"), which accounts for half of the source of randomness in a Random Forest. The Random Subspace Method is an ensemble method that consists of several classifiers each operating in a subspace of the original feature space. The outputs of the models are then combined, usually by a simple majority vote. Tin Kam Ho applied the random subspace method to decision trees in 1995.
[Leo Breiman](https://en.wikipedia.org/wiki/Leo_Breiman) and [Adele Culter](http://www.math.usu.edu/~adele/) combined Breiman's [bagging](https://en.wikipedia.org/wiki/Bootstrap_aggregating) idea with the random subspace method to create a "Random Forest", a name which is trademarked by the duo. Due to the trademark, the algorithm is sometimes called Random Decision Forests.
The introduction of random forests proper was first made in a [paper](http://www.stat.berkeley.edu/~breiman/randomforest2001.pdf) by Leo Breiman [1]. This paper describes a method of building a forest of uncorrelated trees using a CART like procedure, combined with randomized node optimization and bagging. In addition, this paper combines several ingredients, some previously known and some novel, which form the basis of the modern practice of random forests, in particular:
- Using [out-of-bag error](https://en.wikipedia.org/wiki/Out-of-bag_error) as an estimate of the [generalization error](https://en.wikipedia.org/wiki/Generalization_error).
- Measuring [variable importance](https://en.wikipedia.org/wiki/Random_forest#Properties) through permutation.
The report also offers the first theoretical result for random forests in the form of a bound on the generalization error which depends on the strength of the trees in the forest and their correlation.
Although Breiman's implementation of Random Forests used his CART algorithm to construct the decision trees, many modern implementations of Random Forest use entropy-based algorithms for constructing the trees.
## Bagging
Bagging (Bootstrap aggregating) was proposed by Leo Breiman in 1994 to improve the classification by combining classifications of randomly generated training sets. Although it is usually applied to decision tree methods, it can be used with any type of method. Bagging is a special case of the [model averaging](https://en.wikipedia.org/wiki/Ensemble_learning) approach.
- Bagging or *bootstrap aggregation* averages a noisy fitted function, refit to many bootstrap samples to reduce it's variance.
- Bagging can dramatically reduce the variance of unstable procedures (like trees), leading to improved prediction, however any simple, interpretable, model structure (like that of a tree) is lost.
- Bagging produces smoother decision boundaries than trees.
The training algorithm for random forests applies the general technique of bootstrap aggregating, or bagging, to tree learners. Given a training set $X = x_1, ..., x_n$ with responses $Y = y_1, ..., y_n$, bagging repeatedly ($B$ times) selects a random sample with replacement of the training set and fits trees to these samples:
For $b = 1, ..., B$:
1. Sample, with replacement, $n$ training examples from $X$, $Y$; call these $X_b$, $Y_b$.
2. Train a decision or regression tree, $f_b$, on $X_b$, $Y_b$.
After training, predictions for unseen samples $x'$ can be made by averaging the predictions from all the individual regression trees on $x'$:
$$ {\hat {f}}={\frac {1}{B}}\sum _{b=1}^{B}{\hat {f}}_{b}(x')$$
or by taking the majority vote in the case of decision trees.
## Random Forest Algorithm
The above procedure describes the original bagging algorithm for trees. [Random Forests](https://en.wikipedia.org/wiki/Random_forest) differ in only one way from this general scheme: they use a modified tree learning algorithm that selects, at each candidate split in the learning process, a random subset of the features. This process is sometimes called "feature bagging".
- Random Forests correct for decision trees' habit of overfitting to their training set.
- Random Forest is an improvement over bagged trees that "de-correlates" the trees even further, reducing the variance.
- At each tree split, a random sample of $m$ features is drawn and only those $m$ features are considered for splitting.
- Typically $m = \sqrt{p}$ or $\log_2p$ where $p$ is the original number of features.
- For each tree gown on a bootstrap sample, the error rates for observations left out of the bootstrap sample is monitored. This is called the ["out-of-bag"](https://en.wikipedia.org/wiki/Out-of-bag_error) or OOB error rate.
- Each tree has the same (statistical) [expectation](https://en.wikipedia.org/wiki/Expected_value), so increasing the number of trees does not alter the bias of bagging or the Random Forest algorithm.
## Decision Boundary
This is an example of a decision boundary in two dimensions of a (binary) classification Random Forest. The black circle is the Bayes Optimal decision boundary and the blue square-ish boundary is learned by the classification tree.

Source: Elements of Statistical Learning
## Random Forest by Randomization (aka "Extra-Trees")
In [Extremely Randomized Trees](http://link.springer.com/article/10.1007%2Fs10994-006-6226-1) (aka Extra-Trees) [2], randomness goes one step further in the way splits are computed. As in Random Forests, a random subset of candidate features is used, but instead of looking for the best split, thresholds (for the split) are drawn at random for each candidate feature and the best of these randomly-generated thresholds is picked as the splitting rule. This usually allows to reduce the variance of the model a bit more, at the expense of a slightly greater increase in bias.
Extremely Randomized Trees is implemented in the [extraTrees](https://cran.r-project.org/web/packages/extraTrees/index.html) R package and also available in the [h2o](https://0xdata.atlassian.net/browse/PUBDEV-2837) R package as part of the `h2o.randomForest()` function via the `histogram_type = "Random"` argument.
## Out-of-Bag (OOB) Estimates
In random forests, there is no need for cross-validation or a separate test set to get an unbiased estimate of the test set error. It is estimated internally, during the run, as follows:
- Each tree is constructed using a different bootstrap sample from the original data. About one-third of the cases are left out of the bootstrap sample and not used in the construction of the kth tree.
- Put each case left out in the construction of the kth tree down the kth tree to get a classification. In this way, a test set classification is obtained for each case in about one-third of the trees.
- At the end of the run, take j to be the class that got most of the votes every time case n was oob. The proportion of times that j is not equal to the true class of n averaged over all cases is the oob error estimate. This has proven to be unbiased in many tests.
## Variable Importance
In every tree grown in the forest, put down the OOB cases and count the number of votes cast for the correct class. Now randomly permute the values of variable m in the oob cases and put these cases down the tree. Subtract the number of votes for the correct class in the variable-$m$-permuted OOB data from the number of votes for the correct class in the untouched OOB data. The average of this number over all trees in the forest is the raw importance score for variable $m$.
If the values of this score from tree to tree are independent, then the standard error can be computed by a standard computation. The correlations of these scores between trees have been computed for a number of data sets and proved to be quite low, therefore we compute standard errors in the classical way, divide the raw score by its standard error to get a $z$-score, and assign a significance level to the $z$-score assuming normality.
If the number of variables is very large, forests can be run once with all the variables, then run again using only the most important variables from the first run.
For each case, consider all the trees for which it is oob. Subtract the percentage of votes for the correct class in the variable-$m$-permuted OOB data from the percentage of votes for the correct class in the untouched OOB data. This is the local importance score for variable m for this case.
Variable importance in Extremely Randomized Trees is explained [here](http://www.slideshare.net/glouppe/understanding-variable-importances-in-forests-of-randomized-trees).
## Overfitting
Leo Breiman famously claimed that "Random Forests do not overfit." This is perhaps not exactly the case, however they are certainly more robust to overfitting than a Gradient Boosting Machine (GBM). Random Forests can be overfit by growing trees that are "too deep", for example. However, it is hard to overfit a Random Forest by adding more trees to the forest -- typically that will increase accuracy (at the expense of computation time).
## Missing Data
Missing values do not necessarily have to be imputed in a Random Forest implementation, although some software packages will require it.
## Practical Uses
Here is a short article called, [The Unreasonable Effectiveness of Random Forests](https://medium.com/rants-on-machine-learning/the-unreasonable-effectiveness-of-random-forests-f33c3ce28883#.r734znc9f), by Ahmed El Deeb, about the utility of Random Forests. It summarizes some of the algorithm's pros and cons nicely.
## Resources
- [Gilles Louppe - Understanding Random Forests (PhD Dissertation)](http://arxiv.org/abs/1407.7502) (pdf)
- [Gilles Louppe - Understanding Random Forests: From Theory to Practice](http://www.slideshare.net/glouppe/understanding-random-forests-from-theory-to-practice) (slides)
- [Trevor Hastie - Gradient Boosting & Random Forests at H2O World 2014](https://www.youtube.com/watch?v=wPqtzj5VZus&index=16&list=PLNtMya54qvOFQhSZ4IKKXRbMkyLMn0caa) (YouTube)
- [Mark Landry - Gradient Boosting Method and Random Forest at H2O World 2015](https://www.youtube.com/watch?v=9wn1f-30_ZY) (YouTube)
***
# Random Forest Software in R
The oldest and most well known implementation of the Random Forest algorithm in R is the [randomForest](https://cran.r-project.org/web/packages/randomForest/index.html) package. There are also a number of packages that implement variants of the algorithm, and in the past few years, there have been several "big data" focused implementations contributed to the R ecosystem as well.
Here is a non-comprehensive list:
- [randomForest::randomForest](http://www.rdocumentation.org/packages/randomForest/functions/randomForest)
- [h2o::h2o.randomForest](http://www.rdocumentation.org/packages/h2o/functions/h2o.randomForest)
- [DistributedR::hpdRF_parallelForest](https://github.com/vertica/DistributedR/blob/master/algorithms/HPdclassifier/R/hpdRF_parallelForest.R)
- [party::cForest](http://www.rdocumentation.org/packages/party/functions/cforest): A random forest variant for response variables measured at arbitrary scales based on conditional inference trees.
- [randomForestSRC](https://cran.r-project.org/web/packages/randomForestSRC/index.html) implements a unified treatment of Breiman's random forests for survival, regression and classification problems.
- [quantregForest](https://cran.r-project.org/web/packages/quantregForest/index.html) can regress quantiles of a numeric response on exploratory variables via a random forest approach.
- [ranger](https://cran.r-project.org/web/packages/ranger/index.html)
- [Rborist](https://cran.r-project.org/web/packages/Rborist/index.html)
- The [caret](https://topepo.github.io/caret/index.html) package wraps a number of different Random Forest packages in R ([full list here](https://topepo.github.io/caret/Random_Forest.html)):
- Conditional Inference Random Forest (`party::cForest`)
- Oblique Random Forest (`obliqueRF`)
- Parallel Random Forest (`randomForest` + `foreach`)
- Random Ferns (`rFerns`)
- Random Forest (`randomForest`)
- Random Forest (`ranger`)
- Quantile Random Forest (`quantregForest`)
- Random Forest by Randomization (`extraTrees`)
- Random Forest Rule-Based Model (`inTrees`)
- Random Forest with Additional Feature Selection (`Boruta`)
- Regularized Random Forest (`RRF`)
- Rotation Forest (`rotationForest`)
- Weighted Subspace Random Forest (`wsrf`)
- The [mlr](https://github.com/mlr-org/mlr) package wraps a number of different Random Forest packages in R:
- Conditional Inference Random Forest (`party::cForest`)
- Rotation Forest (`rotationForest`)
- Parallel Forest (`ParallelForest`)
- Survival Forest (`randomForestSRC`)
- Random Ferns (`rFerns`)
- Random Forest (`randomForest`)
- Random Forest (`ranger`)
- Synthetic Random Forest (`randomForestSRC`)
- Random Uniform Forest (`randomUniformForest`)
Since there are so many different Random Forest implementations available, there have been several benchmarks to compare the performance of popular implementations, including implementations outside of R. A few examples:
1. [Benchmarking Random Forest Classification](http://www.wise.io/tech/benchmarking-random-forest-part-1) by Erin LeDell, 2013
2. [Benchmarking Random Forest Implementations](http://datascience.la/benchmarking-random-forest-implementations/) by Szilard Pafka, 2015
3. [Ranger](http://arxiv.org/pdf/1508.04409v1.pdf) publication by Marvin N. Wright and Andreas Ziegler, 2015
## randomForest
Authors: Fortran original by [Leo Breiman](http://www.stat.berkeley.edu/~breiman/) and [Adele Cutler](http://www.math.usu.edu/~adele/), R port by [Andy Liaw](https://www.linkedin.com/in/andy-liaw-1399347) and Matthew Wiener.
Backend: Fortran
Features:
- This package wraps the original Fortran code by Leo Breiman and Adele Culter and is probably the most widely known/used implementation in R.
- Single-threaded.
- Although it's single-threaded, smaller forests can be trained in parallel by writing custom [foreach](https://cran.r-project.org/web/packages/foreach/index.html) or [parallel](http://stat.ethz.ch/R-manual/R-devel/library/parallel/doc/parallel.pdf) code, then combined into a bigger forest using the [randomForest::combine()](http://www.rdocumentation.org/packages/randomForest/functions/combine) function.
- Row weights unimplemented (been on the wishlist for as long as I can remember).
- Uses CART trees split by Gini Impurity.
- Categorical predictors are allowed to have up to 53 categories.
- Multinomial response can have no more than 32 categories.
- Supports R formula interface (but I've read some reports that claim it's slower when the formula interface is used).
- GPL-2/3 Licensed.
```
# randomForest example
#install.packages("randomForest")
#install.packages("cvAUC")
library(randomForest)
library(cvAUC)
# Load binary-response dataset
train <- read.csv("data/higgs_train_10k.csv")
test <- read.csv("data/higgs_test_5k.csv")
# Dimensions
dim(train)
dim(test)
# Columns
names(train)
# Identity the response column
ycol <- "response"
# Identify the predictor columns
xcols <- setdiff(names(train), ycol)
# Convert response to factor (required by randomForest)
train[,ycol] <- as.factor(train[,ycol])
test[,ycol] <- as.factor(test[,ycol])
# Train a default RF model with 500 trees
set.seed(1) # For reproducibility
system.time(model <- randomForest(x = train[,xcols],
y = train[,ycol],
xtest = test[,xcols],
ntree = 50))
# Generate predictions on test dataset
preds <- model$test$votes[, 2]
labels <- test[,ycol]
# Compute AUC on the test set
cvAUC::AUC(predictions = preds, labels = labels)
```
## caret method "parRF"
Authors: Max Kuhn
Backend: Fortran (wraps the `randomForest` package)
This is a wrapper for the `randomForest` package that parallelizes the tree building.
```
library(caret)
library(doParallel)
library(e1071)
# Train a "parRF" model using caret
registerDoParallel(cores = 8)
model <- caret::train(x = train[,xcols],
y = train[,ycol],
method = "parRF",
preProcess = NULL,
weights = NULL,
metric = "Accuracy",
maximize = TRUE,
trControl = trainControl(),
tuneGrid = NULL,
tuneLength = 3)
```
## h2o
Authors: [Jan Vitek](http://www.cs.purdue.edu/homes/jv/), [Arno Candel](https://www.linkedin.com/in/candel), H2O.ai contributors
Backend: Java
Features:
- Distributed and parallelized computation on either a single node or a multi-node cluster.
- Automatic early stopping based on convergence of user-specified metrics to user-specified relative tolerance.
- Data-distributed, which means the entire dataset does not need to fit into memory on a single node.
- Uses histogram approximations of continuous variables for speedup.
- Uses squared error to determine optimal splits.
- Automatic early stopping based on convergence of user-specified metrics to user-specified relative tolerance.
- Support for exponential families (Poisson, Gamma, Tweedie) and loss functions in addition to binomial (Bernoulli), Gaussian and multinomial distributions, such as Quantile regression (including Laplace).
- Grid search for hyperparameter optimization and model selection.
- Apache 2.0 Licensed.
- Model export in plain Java code for deployment in production environments.
- GUI for training & model eval/viz (H2O Flow).
Implementation details are presented in slidedecks by [Michal Mahalova](http://www.slideshare.net/0xdata/rf-brighttalk) and [Jan Vitek](http://www.slideshare.net/0xdata/jan-vitek-distributedrandomforest522013).
```
#install.packages("h2o")
library(h2o)
#h2o.shutdown(prompt = FALSE)
h2o.init(nthreads = -1) #Start a local H2O cluster using nthreads = num available cores
# Load binary-response dataset
train <- h2o.importFile("./data/higgs_train_10k.csv")
test <- h2o.importFile("./data/higgs_test_5k.csv")
# Dimensions
dim(train)
dim(test)
# Columns
names(train)
# Identity the response column
ycol <- "response"
# Identify the predictor columns
xcols <- setdiff(names(train), ycol)
# Convert response to factor (required by randomForest)
train[,ycol] <- as.factor(train[,ycol])
test[,ycol] <- as.factor(test[,ycol])
# Train a default RF model with 100 trees
system.time(model <- h2o.randomForest(x = xcols,
y = ycol,
training_frame = train,
seed = 1, #for reproducibility
ntrees = 50))
# Compute AUC on test dataset
# H2O computes many model performance metrics automatically, including AUC
perf <- h2o.performance(model = model, newdata = test)
h2o.auc(perf)
```
## Rborist
Authors: Mark Seligman
Backend: C++
The [Arborist](https://github.com/suiji/Arborist) provides a fast, open-source implementation of the Random Forest algorithm. The Arborist achieves its speed through efficient C++ code and parallel, distributed tree construction. This [slidedeck](http://www.rinfinance.com/agenda/2015/talk/MarkSeligman.pdf) provides detail about the implementation and vision of the project.
Features:
- Began as proprietary implementation, but was open-sourced and rewritten following dissolution of venture.
- Project called "Arborist" but R package is called "Rborist". A Python interface is in development.
- CPU based but a GPU version called Curborist (Cuda Rborist) is in development (unclear if it will be open source).
- Unlimited factor cardinality.
- Emphasizes multi-core but not multi-node.
- Both Python support and GPU support have been "coming soon" since summer 2015, not sure the status of the projects.
- GPL-2/3 licensed.
## ranger
Authors: [Marvin N. Wright](http://www.imbs-luebeck.de/imbs/node/323) and Andreas Ziegler
Backend: C++
[Ranger](http://arxiv.org/pdf/1508.04409v1.pdf) is a fast [implementation](https://github.com/imbs-hl/ranger) of random forest (Breiman 2001) or recursive partitioning, particularly suited for high dimensional data. Classification, regression, probability estimation and survival forests are supported. Classification and regression forests are implemented as in the original Random Forest (Breiman 2001), survival forests as in Random Survival Forests (Ishwaran et al. 2008). For probability estimation forests see Malley et al. (2012).
Features:
- Multi-threaded.
- Direct support for [GWAS](https://en.wikipedia.org/wiki/Genome-wide_association_study) (Genome-wide association study) data.
- Excellent speed and support for high-dimensional or wide data.
- Not as fast for "tall & skinny" data (many rows, few columns).
- GPL-3 licensed.

Plot from the [ranger article](http://arxiv.org/pdf/1508.04409v1.pdf).
## References
[1] [http://www.stat.berkeley.edu/~breiman/randomforest2001.pdf](http://www.stat.berkeley.edu/~breiman/randomforest2001.pdf)
[2] [P. Geurts, D. Ernst., and L. Wehenkel, “Extremely randomized trees”, Machine Learning, 63(1), 3-42, 2006.](http://link.springer.com/article/10.1007%2Fs10994-006-6226-1)
[3] [http://www.cs.uvm.edu/~icdm/algorithms/10Algorithms-08.pdf](http://www.cs.uvm.edu/~icdm/algorithms/10Algorithms-08.pdf)
| github_jupyter |
```
from google.colab import files
uploaded = files.upload()
for fn in uploaded.keys():
print('User uploaded file "{name}" with length {length} bytes'.format(
name=fn, length=len(uploaded[fn])))
from IPython.core.interactiveshell import InteractiveShell
from IPython.display import HTML
InteractiveShell.ast_node_interactivity = 'all'
import warnings
warnings.filterwarnings('ignore', category = RuntimeWarning)
warnings.filterwarnings('ignore', category = UserWarning)
from sklearn.model_selection import train_test_split
from sklearn import preprocessing
import pandas as pd
import numpy as np
# from utils import get_data, generate_output, guess_human, seed_sequence, get_embeddings, find_closest
data = pd.read_csv('./IND115datalabels.csv')
# data2= pd.read_csv('./IND115data.csv')
diff=data.values[:,0]
ndiff=diff.shape[0]
for i in range(1,ndiff):
diff[i]=diff[i]-diff[i-1]
xm=data.values[:,0]
ym=data.values[:,1]
timesteps=10
n=2
labelstrue=data.values[:,2]
m=xm.shape[0]
ux=np.mean(xm)
uy=np.mean(ym)
rx=np.max(xm)-np.min(xm)
ry=np.max(ym)-np.min(ym)
#m,n=xm.shape
#xm=preprocessing.MinMaxScaler(feature_range=(-1,1)).fit_transform(xm.reshape(-1,1))
#ym=preprocessing.MinMaxScaler(feature_range=(-1,1)).fit_transform(ym.reshape(-1,1))
xm=(xm-ux)/rx
ym=(ym-uy)/ry
xtmp=np.empty((m,2))
xtmp[:,0]=xm
xtmp[:,1]=ym
xnew=np.array(xtmp[0:timesteps,:]).reshape(1,-1,n)
print(xnew)
for i in range(1,m-timesteps):
xnew=np.append(xnew,np.array(xtmp[i:i+timesteps,:]).reshape(1,-1,n),axis=0)
print(xnew.shape)
new_m=xnew.shape[0]
# y=ym[:new_m]
y=labelstrue[timesteps:]
y.shape
print(xnew[:5,:,:])
print(y[:5])
data.shape
print(xnew[:5,:,0])
print(y[:5])
X=xnew
#X_train, X_test, y_train, y_test = train_test_split(X,y,test_size=0.2,shuffle=False)
X_train=X[2000:,:,:]
y_train=y[2000:]
X_test=X[:2000,:,:]
y_test=y[:2000]
X_train.shape
print("Xtrain",X_train[:5,:,:])
#print("ytrain",y_train[:5])
print("Xtest",X_test[:5,:,:])
#print("ytest",y_test[:5])
X_test.shape
from keras.models import Sequential, load_model
from keras.layers import LSTM, Dense, Dropout, Embedding, Masking, Bidirectional
from keras.optimizers import Adam, RMSprop, SGD
from keras.utils import plot_model
model = Sequential()
# timesteps=5
# Recurrent layer 1
model.add(
LSTM(
32,input_shape=(timesteps,n),return_sequences=False, dropout=0.1,
recurrent_dropout=0.1))
# Fully connected layer 2
model.add(Dense(10, activation='tanh'))
# Fully connected layer 3
# model.add(Dense(5, activation='tanh'))
# Fully connected layer 4
model.add(Dense(1, activation='sigmoid'))
# Dropout for regularization
#model.add(Dropout(0.5))
#rmsprop=RMSprop(lr=0.01, rho=0.9, epsilon=None, decay=0.0) #lr=0.001
#model.compile(
# optimizer=rmsprop, loss='mean_squared_error')
# sgd=SGD(lr=0.01, momentum=0.0, decay=0.0, nesterov=False)
# model.compile(
# optimizer=sgd, loss='mean_squared_error')
# Compile the model
adam=Adam(lr=0.001, beta_1=0.9, beta_2=0.999, epsilon=None, decay=0.0, amsgrad=False)
model.compile(
optimizer=adam, loss='binary_crossentropy')
model.summary()
from keras.models import load_model
print(X_train.shape)
# Load in model and demonstrate training
# model = load_model('../models/train-embeddings-rnn.h5')
h = model.fit(X_train, y=y_train, epochs = 50, batch_size = 10,
validation_data = (X_test, y_test),
verbose = 1)
import matplotlib.pyplot as plt
nn=1000
xplot=X_train[:nn,:,:]
yplot=y_train[:nn]
ypredict=[]
for i in range(0,nn):
ypredict=np.append(ypredict,model.predict(xplot[i,:,:].reshape(1,timesteps,n)))
# ypredict=np.append(ypredict,model.predict(xplot))
plt.figure(figsize=(12,5))
xt= np.arange(0,nn,1)
plt.plot(xt,yplot,'ro',label='Actual output')
plt.plot(xt,ypredict,'go',label='Predicted output')
plt.legend()
# plt.plot(xt,yplot-ypredict,'bo')
import matplotlib.pyplot as plt
plt.plot(h.history['loss'])
plt.plot(h.history['val_loss'])
from sklearn import metrics
thresh=0.9
ylen=len(ypredict)
labelspred=np.empty(ylen)
for i in range(0,ylen):
if(ypredict[i]>thresh):
labelspred[i]=1
else:
labelspred[i]=0
print("Accuracy score ",metrics.accuracy_score(labelstrue[2000:3000], labelspred, normalize=True))
print("Precision score ",metrics.precision_score(labelstrue[2000:3000],labelspred,average=None))
print("Recall score ",metrics.recall_score(labelstrue[2000:3000], labelspred, average=None))
print("f1 score ",metrics.f1_score(labelstrue[2000:3000], labelspred, average=None))
print('Model Performance: Log Loss and Accuracy on training data')
model.evaluate(X_train, y_train, batch_size = 10)
print('\nModel Performance: Log Loss and Accuracy on validation data')
model.evaluate(X_test,y_test, batch_size = 10)
```
| github_jupyter |
# Informes de la comunidad de Madrid
Actualizado diariamente, este documento se [visualiza mejor aquí](https://nbviewer.jupyter.org/github/jaimevalero/COVID-19/blob/master/jupyter/Madrid_Pain_Graphs.ipynb).
Datos de la situación de la infección por coronavirus en la Comunidad de Madrid.
Nos descargamos los datos, agrupamos, y calculamos :
- Gráfico de seguimiento.
- Muertes medias diarias, últimos 7 días.
- Muertes medias diarias desde que la comunidad de Madrid publica datos.
```
# Miramos si hay nuevos datos a descargar.
!# cd ../data/; FILELIST=" 200509 200508 200507 200506 200505 200504 200503 200502 200501 200430 200429 200428 200427 200426 200425 200424 200423 200422 200510 200511 200512 200513 200514 200515 200516 200517 200518 200519 200520 200521 200522 200523 200524 200525 200526 200527 200528 200529 200530 200609 200608 200607 200606 200605 200604 200603 200602 200601 200610 200611 200612 200613 200614 200615 200616 200617 200618 200619 200620 200621 200622 200623 200624 200625 200626 200627 200628 200629 200630 " ; for fecha in `echo $FILELIST` ; do FILE=${fecha}_cam_covid19.pdf ; [ ! -f ../data/${FILE} ] && echo $FILE::::: && wget https://www.comunidad.madrid/sites/default/files/doc/sanidad/$FILE 1>/dev/null 2>/dev/null && ls -altr $FILE ; done
# Miramos solo hoy y los ultimos diez dias
! cd ../data/; FILELIST=`seq -w 0 7 | while read i ; do date +%y%m%d -d "$i day ago" ; done` ; for fecha in `echo $FILELIST` ; do FILE=${fecha}_cam_covid19.pdf ; [ ! -f ../data/${FILE} ] && echo $FILE::::: && wget https://www.comunidad.madrid/sites/default/files/aud/sanidad/$FILE 1>/dev/null 2>/dev/null && ls -altr $FILE ; done
! cd ../data/; FILELIST=`seq -w 0 7 | while read i ; do date +%y%m%d -d "$i day ago" ; done` ; for fecha in `echo $FILELIST` ; do FILE=${fecha}_cam_covid19.pdf ; [ ! -f ../data/${FILE} ] && echo $FILE::::: && wget https://www.comunidad.madrid/sites/default/files/doc/sanidad/$FILE 1>/dev/null 2>/dev/null && ls -altr $FILE ; done
! cd ../data/; FILELIST=`seq -w 0 7 | while read i ; do date +%y%m%d -d "$i day ago" ; done` ; for fecha in `echo $FILELIST` ; do FILE=${fecha}cam_covid19.pdf ; [ ! -f ../data/${FILE} ] && echo $FILE::::: && wget https://www.comunidad.madrid/sites/default/files/doc/sanidad/$FILE 1>/dev/null 2>/dev/null && ls -altr $FILE ; done
! cd ../data/; FILELIST=`seq -w 0 7 | while read i ; do date +%y%m%d -d "$i day ago" ; done` ; for fecha in `echo $FILELIST` ; do FILE=${fecha}_cam_covid19.pdf ; [ ! -f ../data/${FILE} ] && echo $FILE::::: && wget https://www.comunidad.madrid/sites/default/files/doc/sanidad/$FILE 1>/dev/null 2>/dev/null && ls -altr $FILE ; done
! cd ../data/; FILELIST=`seq -w 0 7 | while read i ; do date +%y%m%d -d "$i day ago" ; done` ; for fecha in `echo $FILELIST` ; do FILE=${fecha}_cam_covid19.pdf ; [ ! -f ../data/${FILE} ] && echo $FILE::::: && wget https://www.comunidad.madrid/sites/default/files/doc/sanidad/$FILE 1>/dev/null 2>/dev/null && ls -altr $FILE ; done
! cd ../data/; FILELIST=`seq -w 0 7 | while read i ; do date +%Y%m%d -d "$i day ago" ; done` ; for fecha in `echo $FILELIST` ; do FILE=${fecha}_cam_covid19.pdf ; [ ! -f ../data/${FILE} ] && echo $FILE::::: && wget https://www.comunidad.madrid/sites/default/files/doc/sanidad/$FILE 1>/dev/null 2>/dev/null && ls -altr $FILE ; done
! cd ../data/; FILELIST=`seq -w 0 7 | while read i ; do date +%y%m%d -d "$i day ago" ; done` ; for fecha in `echo $FILELIST` ; do FILE=${fecha}_cam_covid19.pdf ; [ ! -f ../data/${FILE} ] && echo $FILE::::: && wget https://www.comunidad.madrid/sites/default/files/$FILE 1>/dev/null 1>/dev/null 2>/dev/null && ls -altr $FILE ; done
! cd ../data/; FILELIST=`seq -w 0 7 | while read i ; do date +%-d.%-m.%Y -d "$i day ago" ; done` ; for fecha in `echo $FILELIST` ; do FILE=${fecha}_2.pdf ; [ ! -f ../data/${FILE} ] && echo $FILE::::: && wget https://www.comunidad.madrid/sites/default/files/doc/sanidad/$FILE 1>/dev/null 2>/dev/null && ls -altr $FILE ; done
! cd ../data/; FILELIST=`seq -w 0 7 | while read i ; do date +%-d.%-m.%Y -d "$i day ago" ; done` ; for fecha in `echo $FILELIST` ; do FILE=${fecha}.pdf ; [ ! -f ../data/${FILE} ] && echo $FILE::::: && wget https://www.comunidad.madrid/sites/default/files/doc/sanidad/$FILE 1>/dev/null 2>/dev/null && ls -altr $FILE ; done
! cd ../data/; FILELIST=`seq -w 0 7 | while read i ; do date +%y%m%d -d "$i day ago" ; done` ; for fecha in `echo $FILELIST` ; do FILE=${fecha}cam_covid19.pdf ; [ ! -f ../data/${FILE} ] && echo $FILE::::: && wget https://www.comunidad.madrid/sites/default/files/doc/sanidad/$FILE 1>/dev/null 2>/dev/null && ls -altr $FILE ; done
! cd ../data/; FILELIST=`seq -w 0 7 | while read i ; do date +%y%m%d -d "$i day ago" ; done` ; for fecha in `echo $FILELIST` ; do FILE=${fecha}_cam_covid-19.pdf ; [ ! -f ../data/${FILE} ] && echo $FILE::::: && wget https://www.comunidad.madrid/sites/default/files/doc/sanidad/$FILE 1>/dev/null 2>/dev/null && ls -altr $FILE ; done
! cd ../data/; FILELIST=`seq -w 0 7 | while read i ; do date +%-d%-m%Y -d "$i day ago" ; done` ; for fecha in `echo $FILELIST` ; do FILE=${fecha}_cam_covid19.pdf ; [ ! -f ../data/${FILE} ] && echo $FILE::::: && wget https://www.comunidad.madrid/sites/default/files/doc/sanidad/$FILE 1>/dev/null 2>/dev/null && ls -altr $FILE ; done
! cd ../data/; FILELIST=`seq -w 0 7 | while read i ; do date +%y%m%d -d "$i day ago" ; done` ; for fecha in `echo $FILELIST` ; do FILE=${fecha}_cam_covid19.pdf ; [ ! -f ../data/${FILE} ] && echo $FILE::::: && wget https://www.comunidad.madrid/sites/default/files/doc/sanidad/prev/$FILE 1>/dev/null 2>/dev/null && ls -altr $FILE ; done
! cd ../data/; FILELIST=`seq -w 0 7 | while read i ; do date +%y%m%d -d "$i day ago" ; done` ; for fecha in `echo $FILELIST` ; do FILE=${fecha}_cam_covid19.pdf ; [ ! -f ../data/${FILE} ] && echo $FILE::::: && wget https://www.comunidad.madrid/sites/default/files/aud/sanidad/$FILE 1>/dev/null 2>/dev/null && ls -altr $FILE ; done
! cd ../data/; FILELIST=`seq -w 0 7 | while read i ; do date +%y%m%d -d "$i day ago" ; done` ; for fecha in `echo $FILELIST` ; do FILE=${fecha}_cam_covid.pdf ; [ ! -f ../data/${FILE} ] && echo $FILE::::: && wget https://www.comunidad.madrid/sites/default/files/doc/sanidad/$FILE 1>/dev/null 2>/dev/null && ls -altr $FILE ; done
! cd ../data/; FILELIST=`seq -w 0 7 | while read i ; do date +%y%m%d -d "$i day ago" ; done` ; for fecha in `echo $FILELIST` ; do FILE=${fecha}_cam_covid19.pdf ; [ ! -f ../data/${FILE} ] && echo $FILE::::: && wget https://www.comunidad.madrid/sites/default/files/doc/presidencia/$FILE 1>/dev/null 2>/dev/null && ls -altr $FILE ; done
! cd ../data/; FILELIST=`seq -w 0 7 | while read i ; do date +%y%m%d -d "$i day ago" ; done` ; for fecha in `echo $FILELIST` ; do FILE=${fecha}_cam_covid19.pdf.pdf ; [ ! -f ../data/${FILE} ] && echo $FILE::::: && wget https://www.comunidad.madrid/sites/default/files/doc/sanidad/$FILE 1>/dev/null 2>/dev/null && ls -altr $FILE ; done
! cd /root/scripts/COVID-19/data; ls -1 *pdf.pdf | while read fichero ; do NOMBRE=`echo $fichero | sed -e 's@.pdf.pdf@.pdf@g' ` ; mv "$fichero" "$NOMBRE" ; done ; cd -
from tabula import read_pdf
from IPython.display import display, HTML
import os
import pandas as pd
import glob
import re
from tqdm.notebook import tqdm
import warnings
import os.path
#import datetime
warnings.filterwarnings('ignore')
os.environ["JAVA_HOME"] = "/usr/lib/jvm/java-1.8.0-openjdk-1.8.0.141-1.b16.el7_3.x86_64/jre"
# Auxiliary functions
from datetime import datetime, date, time, timedelta
df_cache = pd.read_csv("/root/kaggle/covid19-madrid/madrid_results.csv")
def query_cache(fecha):
""" Query cache file to avoid parse pdf
return empty dataframe is not found"""
try :
if '.' in fecha:
date_regexp='%d.%m.%Y'
else:
date_regexp='%y%m%d'
date_formatted = datetime.strptime(fecha,date_regexp ).strftime('%Y-%m-%d')
df = df_cache.query( 'Fecha==@date_formatted')
except:
print("Cache miss:" , fecha)
return pd.DataFrame()
#print(f"fecha {fecha},{date_formatted}")
try:
df['Fecha'] = pd.to_datetime(date_formatted, format='%y-%m-%d')
except :
df['Fecha'] = pd.to_datetime(date_formatted, format='%Y-%m-%d')
df.set_index('Fecha', inplace=True, drop=True)
return df
""" Rellenar dias vacios con interpolacion"""
def interpolate_dataframe(df,freq):
if freq == 'H':
rng = pd.date_range(df.index.min(), df.index.max() + pd.Timedelta(23, 'H'), freq='H')
elif freq == 'D' :
rng = pd.date_range(
datetime.strptime(str(df.index.min())[:10]+' 00:00:00', "%Y-%m-%d %H:%M:%S") ,
datetime.strptime(str(df.index.max())[:10]+' 00:00:00', "%Y-%m-%d %H:%M:%S"),
freq='D')
df.index = pd.to_datetime(df.index)
df2 = df.reindex(rng)
df = df2
for column in df.columns :
s = pd.Series(df[column])
s.interpolate(method="quadratic", inplace =True)
df[column] = pd.DataFrame([s]).T
return df
def get_daily_date_new_format_vacunas(fecha,filename):
""" Se añadio una página de vacunas que obviamos"""
BLACKLIST= ["792021"]
if fecha in BLACKLIST :
print (f"Error, fecha no reconocida, está en blacklist {fecha}")
return pd.DataFrame()
print(f"get_daily_date_new_format_vacunas, {fecha}, {filename}")
file_path = '../data/'+fecha+'_cam_covid19.pdf'
if not os.path.isfile(file_path):
file_path = '../data/'+fecha+'cam_covid19.pdf'
if not os.path.isfile(file_path):
file_path = filename
try:
df_pdf = read_pdf(file_path,area=(150, 625, 400, 900) , pages=2)[0]
except:
print(f"ERROR,{fecha},{filename}" )
return pd.DataFrame()
df = df_pdf['Datos sanidad mortuoria.'].astype(str).str.replace(r".", '').replace("(", ' ').replace(r"*","")
df = pd.DataFrame(df)
dict = {}
dict['HOSPITALES'] = df[df['Datos sanidad mortuoria.'].str.contains('Hospitales')].iloc[0]['Datos sanidad mortuoria.'].split(' ')[0]
dict['DOMICILIOS'] = df[df['Datos sanidad mortuoria.'].str.contains('Domicilios')].iloc[0]['Datos sanidad mortuoria.'].split(' ')[0]
dict['CENTROS SOCIOSANITARIOS'] = df[df['Datos sanidad mortuoria.'].str.contains('Centros')].iloc[0]['Datos sanidad mortuoria.'].split(' ')[0]
dict['OTROS LUGARES'] = df[df['Datos sanidad mortuoria.'].str.contains('otros')].iloc[0]['Datos sanidad mortuoria.'].split(' ')[0]
cadena_a_parsear = df[df['Datos sanidad mortuoria.'].str.contains('otal')].iloc[0]['Datos sanidad mortuoria.']
dict['FALLECIDOS TOTALES'] = re.search(r'(\d+)', cadena_a_parsear)[0]
try:
df2_pdf = read_pdf(file_path,area=(300, 100, 800, 400) , pages=2)
dict['PACIENTES UCI DIA'] = df2_pdf[0].loc[3:3].values[0][1].replace(".",'')
dict['PACIENTES UCI ACUMULADOS']= df2_pdf[0].loc[6:6].values[0][1].replace(".",'')
dict['HOSPITALIZADOS']= int(df2_pdf[0].loc[3:3].to_records()[0][1].replace(".",""))
df2_pdf = read_pdf(file_path,area=(150, 100, 300, 400) , pages=2)
# Comparacion porque añadieron una pagina para las vacunas
if df2_pdf[0].columns[0] != 'CASOS POSITIVOS' :
df2_pdf = read_pdf(file_path,area=(150, 100, 300, 400) , pages=1)
dict['POSITIVOS']= int(df2_pdf[0].loc[2:2].values[0][0].replace(".",''))
if dict['POSITIVOS'] == 0 :
raise Exception(f"file_path: {file_path}")
print(f"""file_path: {file_path}, {dict['POSITIVOS']} """)
except Exception as e:
print(f"{fecha} mal parseada: {e}")
df = pd.DataFrame.from_dict(dict, orient='index').T
if '.' in fecha :
try:
df['Fecha'] = pd.to_datetime(fecha, format='%d.%m.%Y')
except :
df['Fecha'] = pd.to_datetime(fecha, format='%d.%m.%y')
else:
try:
df['Fecha'] = pd.to_datetime(fecha, format='%y%m%d')
except :
df['Fecha'] = pd.to_datetime(fecha, format='%Y%m%d')
df.set_index('Fecha', inplace=True, drop=True)
return df
def get_daily_date_new_format(fecha,filename):
print(f"get_daily_date_new_format({fecha},{filename})")
PAGINA_DE_DATOS=1
file_path = '../data/'+fecha+'_cam_covid19.pdf'
if not os.path.isfile(file_path):
file_path = '../data/'+fecha+'cam_covid19.pdf'
if not os.path.isfile(file_path):
file_path = filename
#print("Analizando:" + file_path)
df_pdf = read_pdf(file_path,area=(000, 600, 400, 800) , pages=PAGINA_DE_DATOS)
# Parche, para los saltos de linea en el pdf
if 'Unnamed: 0' not in df_pdf[0].columns :
return pd.DataFrame()
df = df_pdf[0]
df = df['Unnamed: 0'].astype(str).str.replace(r".", '').replace("(", ' ')
df = df.T
df.columns = df.iloc[0]
df = df.iloc[1:]
#print("2 get_daily_date_new_format")
df = pd.DataFrame(data=df)
df
dict = {}
try:
df2_pdf = read_pdf(file_path,area=(300, 100, 800, 400) , pages=PAGINA_DE_DATOS)
dict['PACIENTES UCI DIA'] = df2_pdf[0].loc[3:3].values[0][1].replace(".",'')
dict['PACIENTES UCI ACUMULADOS']= df2_pdf[0].loc[6:6].values[0][1].replace(".",'')
dict['HOSPITALIZADOS']= int(df2_pdf[0].loc[3:3].to_records()[0][1].replace(".",""))
except Exception as e:
print(f"{fecha} mal parseada: {e}")
try:
df2_pdf = read_pdf(file_path,area=(150, 100, 300, 400) , pages=1)
dict['POSITIVOS']= int(df2_pdf[0].loc[2:2].values[0][0].replace(".",''))
if dict['POSITIVOS'] == 0 :
raise Exception(f"file_path: {file_path} {dict['POSITIVOS']} ")
print(f"""file_path: {file_path}, {dict['POSITIVOS']} """)
#raise Exception(f"file_path: {file_path} {dict['POSITIVOS']} ")
except Exception as e:
import traceback
traceback.print_exc()
raise Exception(f"file_path: {file_path} {dict['POSITIVOS']} ")
dict['HOSPITALES'] = df[df['Unnamed: 0'].str.contains('Hospitales')].iloc[0]['Unnamed: 0'].split(' ')[0]
dict['DOMICILIOS'] = df[df['Unnamed: 0'].str.contains('Domicilios')].iloc[0]['Unnamed: 0'].split(' ')[0]
dict['CENTROS SOCIOSANITARIOS'] = df[df['Unnamed: 0'].str.contains('Centros')].iloc[0]['Unnamed: 0'].split(' ')[0]
dict['OTROS LUGARES'] = df[df['Unnamed: 0'].str.contains('otros')].iloc[0]['Unnamed: 0'].split(' ')[0]
#print("3 get_daily_date_new_format")
cadena_a_parsear = df[df['Unnamed: 0'].str.contains('otal')].iloc[0]['Unnamed: 0']
dict['FALLECIDOS TOTALES'] = re.search(r'(\d+)', cadena_a_parsear)[0]
#print("4 get_daily_date_new_format")
df = pd.DataFrame.from_dict(dict, orient='index').T
#print("4.5 get_daily_date_new_format")
if '.' in fecha :
try:
df['Fecha'] = pd.to_datetime(fecha, format='%d.%m.%Y')
except :
df['Fecha'] = pd.to_datetime(fecha, format='%d.%m.%y')
else:
try:
df['Fecha'] = pd.to_datetime(fecha, format='%y%m%d')
except :
df['Fecha'] = pd.to_datetime(fecha, format='%Y%m%d')
#print("5 get_daily_date_new_format")
df.set_index('Fecha', inplace=True, drop=True)
#print(df)
return df
def get_daily_data(fecha,filename):
#print(f"""get_daily_data: {fecha}""")
#print(f"""../data/{fecha}_cam_covid19.pdf""")
if fecha > "210228" :
#print(f"Detected vacunas format {fecha}")
return get_daily_date_new_format_vacunas(fecha,filename)
if fecha > '200512' :
return get_daily_date_new_format(fecha,filename)
col2str = {'dtype': str}
kwargs = {'output_format': 'dataframe',
'pandas_options': col2str,
'stream': True}
df_pdf = read_pdf('../data/'+fecha+'_cam_covid19.pdf',pages='1',multiple_tables = True,**kwargs)
df = df_pdf[0]
df = df[df['Unnamed: 0'].notna()]
df = df[(df['Unnamed: 0']=='HOSPITALES') | (df['Unnamed: 0'] == 'DOMICILIOS') | (df['Unnamed: 0'] == 'CENTROS SOCIOSANITARIOS') | (df['Unnamed: 0'] == 'OTROS LUGARES') | (df['Unnamed: 0'] == 'FALLECIDOS TOTALES')]
df = df[['Unnamed: 0','Unnamed: 2']]
df['Unnamed: 2'] = df['Unnamed: 2'].astype(str).str.replace(r".", '')
df = df.T
df.columns = df.iloc[0]
df = df.iloc[1:]
df['Fecha'] = pd.to_datetime(fecha, format='%y%m%d')
df = df.rename_axis(None)
df.set_index('Fecha', inplace=True, drop=True)
df.index
df.dropna()
#df = df.T
return df
def get_all_data( ):
#BLACKLIST = ["200429","200422"]
#BLACKLIST = ["200514",]
BLACKLIST = []
df = pd.DataFrame()
list_df = []
#pdf_list= (glob.glob('../data/*_covid19.pdf'),
# key=os.path.getmtime,
# reverse=True )
pdf_list= set(glob.glob('../data/*202*pdf') +
glob.glob('../data/*cam_covid19.pdf') +
glob.glob('../data/*cam_covid-19.pdf')+
glob.glob('../data/*cam_covid.pdf')
)
for pdf_file in tqdm(pdf_list,
desc="Procesando pdfs diarios"):
# extract fecha from username , eg : ../data/2200422_cam_covid19.pdf
format_point_occurences = pdf_file.split('/')[2].split('_')[0].count(".")
# Hack to fix filename inconsistences on remote server
if format_point_occurences > 2 :
day = pdf_file.split('/')[2].split('_')[0].split('.')[0].zfill(2)
month = pdf_file.split('/')[2].split('_')[0].split('.')[1].zfill(2)
year = pdf_file.split('/')[2].split('_')[0].split('.')[2][-2:]
fecha = year+month+day
fecha=fecha.replace('.pdf','')
else :
fecha = pdf_file.split('/')[2].split('_')[0].replace('cam_','').replace('_cam_','').replace('cam','')
if fecha not in BLACKLIST:
# query cache, otherwise parse pdf
df = query_cache(fecha)
if df.empty:
df = get_daily_data(fecha,pdf_file)
list_df.append(df)
df = pd.concat(list_df)
df = df.fillna(0)
df = df.astype(int)
df = df.drop_duplicates()
df = df.sort_values(by=['Fecha'], ascending=True)
df['HOSPITALES hoy'] = df['HOSPITALES'] - df['HOSPITALES'].shift(1)
df['CENTROS SOCIOSANITARIOS hoy'] = df['CENTROS SOCIOSANITARIOS'] - df['CENTROS SOCIOSANITARIOS'].shift(1)
df['FALLECIDOS TOTALES hoy'] = df['FALLECIDOS TOTALES'] - df['FALLECIDOS TOTALES'].shift(1)
df = df.sort_values(by=['Fecha'], ascending=False)
return df
total = get_all_data()
total.to_csv('/root/kaggle/covid19-madrid/madrid_results.csv')
total
from tabula import read_pdf
from IPython.display import display, HTML
import os
import pandas as pd
import glob
import re
from tqdm.notebook import tqdm
import warnings
import os.path
#import datetime
warnings.filterwarnings('ignore')
#file_path = "../data/210130_cam_covid19.pdf"
file_path = "../data/210923_cam_covid19.pdf"
df2_pdf = read_pdf(file_path,area=(150, 100, 300, 400) , pages=2)
# Comparacion porque añadieron una pagina para las vacunas
if df2_pdf[0].columns[0] != 'CASOS POSITIVOS' :
df2_pdf = read_pdf(file_path,area=(150, 100, 300, 400) , pages=1)
value = int(df2_pdf[0].loc[2:2].values[0][0].replace(".",''))
value
total.to_csv('/root/kaggle/covid19-madrid/madrid_results.csv')
total
df = total
df = df.fillna(0)
df = df.astype(int)
df
import re
import PyPDF2
def get_daily_date_edades(fecha,filename,page):
""" Se añadio una página de vacunas que obviamos"""
#print(f"get_daily_date_new_format_vacunas, {fecha}, {filename}")
file_path = '../data/'+fecha+'_cam_covid19.pdf'
if not os.path.isfile(file_path):
file_path = '../data/'+fecha+'cam_covid19.pdf'
if not os.path.isfile(file_path):
file_path = filename
col2str = {'dtype': str}
kwargs = {'output_format': 'dataframe',
'pandas_options': col2str,
'multiple_tables' :True,
'stream': True,
'area' : (150, 00, 400, 400)}
df_pdf = read_pdf('../data/'+fecha+'_cam_covid19.pdf',pages=page,**kwargs)
df = df_pdf[0]
for column in ['Unnamed: 2','Unnamed: 3','Unnamed: 6','Unnamed: 7']:
print(column,df.columns.values )
if column in df.columns.values :
df.drop( column , inplace=True,axis=1)
df = df.dropna()
df = df.rename(columns = {'Unnamed: 0': 'Ages' , 'Mujeres': 'Female', 'Hombres': 'Male'}, inplace = False)
#df['Date'] = datetime.strptime(fecha,date_regexp ).strftime('%Y-%m-%d')
if '.' in fecha:
date_regexp='%d.%m.%Y'
else:
date_regexp='%y%m%d'
date_formatted = datetime.strptime(fecha,date_regexp ).strftime('%Y-%m-%d')
df['Fecha'] = pd.to_datetime(date_formatted, format='%Y-%m-%d')
return df
fecha='210404'
filename='../data/210404_cam_covid19.pdf'
df_ages= get_daily_date_edades(fecha,filename,11)
df_ages
df2_pdf[0]
int(df2_pdf[0].loc[3:3].to_records()[0][1].replace(".",""))
pd.set_option('display.max_rows', 500)
def get_all_data( ):
#BLACKLIST = ["200429","200422"]
#BLACKLIST = ["200514",]
BLACKLIST = []
df = pd.DataFrame()
list_df = []
#pdf_list= (glob.glob('../data/*_covid19.pdf'),
# key=os.path.getmtime,
# reverse=True )
pdf_list= set( glob.glob('../data/2104*cam_covid19.pdf'))
for pdf_file in tqdm(pdf_list,
desc="Procesando pdfs diarios"):
# extract fecha from username , eg : ../data/2200422_cam_covid19.pdf
format_point_occurences = pdf_file.split('/')[2].split('_')[0].count(".")
# Hack to fix filename inconsistences on remote server
if format_point_occurences > 2 :
day = pdf_file.split('/')[2].split('_')[0].split('.')[0].zfill(2)
month = pdf_file.split('/')[2].split('_')[0].split('.')[1].zfill(2)
year = pdf_file.split('/')[2].split('_')[0].split('.')[2][-2:]
fecha = year+month+day
fecha=fecha.replace('.pdf','')
else :
fecha = pdf_file.split('/')[2].split('_')[0].replace('cam_','').replace('_cam_','').replace('cam','')
for page in range(9, 13):
try:
df= get_daily_date_edades(fecha,filename,11)
list_df.append(df)
except:
pass
return list_df
list_df = get_all_data( )
df_ages = pd.concat(list_df)
df_ages.query( 'Ages=="80-89"')
total
VENTANA_MEDIA_MOVIL=7
df = interpolate_dataframe(total,'D')
df.index.name = 'Fecha'
df = df.sort_values(by=['Fecha'], ascending=True)
df['HOSPITALES hoy'] = df['HOSPITALES'] - df['HOSPITALES'].shift(1)
df['CENTROS SOCIOSANITARIOS hoy'] = df['CENTROS SOCIOSANITARIOS'] - df['CENTROS SOCIOSANITARIOS'].shift(1)
df['FALLECIDOS TOTALES hoy'] = df['FALLECIDOS TOTALES'] - df['FALLECIDOS TOTALES'].shift(1)
df['MA CENTROS SOCIOSANITARIOS hoy'] = df['CENTROS SOCIOSANITARIOS hoy'].rolling(window=VENTANA_MEDIA_MOVIL).mean()
df['MA HOSPITALES hoy'] = df['HOSPITALES hoy'].rolling(window=VENTANA_MEDIA_MOVIL).mean()
df['MA FALLECIDOS TOTALES hoy'] = df['FALLECIDOS TOTALES hoy'].rolling(window=VENTANA_MEDIA_MOVIL).mean()
df = df.sort_index(ascending=False)
df_master = df.copy()
total.head()
# Hacemos lo contrario
# En lugar de sacar el nº de muertos dado el nº de infectados, como lo primero lo sabemos (en madrid), sacamos lo segundo y extrapolamos al conjunto de españa
df = df_master
R0_estimada = df['FALLECIDOS TOTALES hoy'].values[0:7].sum() / df['FALLECIDOS TOTALES hoy'].values[7:14].sum()
print(df['FALLECIDOS TOTALES hoy'].values[0:7].sum(), df['FALLECIDOS TOTALES hoy'].values[7:14].sum() )
print(f"""R0_estimada = {R0_estimada}""")
PROPORCION_ENFERMOS_MUERTOS=750000/15000 # Esta es la proporcion enfermos muertos (15.000 muertos para 750.000 afectados)
RATIO_NO_HEMOS_COLAPSADO=2 # La mitad de los muertos se ha calculado del colapso. Como ahora no hemos colapsado
PESO_MADRID_MUERTES_TOTALES=1/3
casos_españa_estimados = df['FALLECIDOS TOTALES hoy'].values[0:5].sum() * PROPORCION_ENFERMOS_MUERTOS * RATIO_NO_HEMOS_COLAPSADO / PESO_MADRID_MUERTES_TOTALES
print(f"""casos_españa_estimados = {casos_españa_estimados}""")
```
## Gráfico estimacion R0
Considerando solo los datos de Madrid, estimamos el R0 a partir del nº de muertos (considerando que el nº de muertos es una combinacion lineal del nº de enfermos), por lo que es posible calcular el ratio igual.
Para calcular el R0, sacamos la suma de muertos de la última semana, entre la suma de muertos de la semana anterior.
```
from datetime import datetime, timedelta
import seaborn as sns
from matplotlib import pyplot as plt
import matplotlib.dates as mdates
df = df_master
def calcular_estimaciones_R0(df):
def calcular_R0_dia(dia,df):
dia_semana_anterior = dia - timedelta(days=7)
return dia,df.loc[dia:dia - timedelta(days=6)]['FALLECIDOS TOTALES hoy'].sum() / df.loc[dia- timedelta(days=7):dia - timedelta(days=13)]['FALLECIDOS TOTALES hoy'].sum()
VENTANA_MEDIA_MOVIL=7
df_R0_estimada = pd.DataFrame([calcular_R0_dia(dia,df) for dia in df.index[0:50]],columns=['Fecha','R0_estimada'])
df_R0_estimada = df_R0_estimada.sort_values(by=['Fecha'], ascending=True)
df_R0_estimada['MA R0_estimada'] = df_R0_estimada['R0_estimada'].rolling(window=VENTANA_MEDIA_MOVIL).mean()
df_R0_estimada = df_R0_estimada.sort_values(by=['Fecha'], ascending=False)
df_R0_estimada.set_index('Fecha', inplace=True, drop=True)
return df_R0_estimada
df= calcular_estimaciones_R0(df_master)
#df=df[['R0_estimada']]
df
chart_df=df[df.columns[-3:]]
chart_df.plot(legend=True,figsize=(13.5,9), marker='o')
plt.gca().xaxis.set_major_formatter(mdates.DateFormatter('%m-%d'))
plt.gca().xaxis.set_major_locator(mdates.DayLocator(interval=1))
plt.xticks(rotation=45)
ax = plt.gca()
ax.axhline(1, color='r',linestyle = ':' )
ax.set_title("Estimacion R0 Comunidad de Madrid")
ax.set_ylim(ymin=0)
plt.show()
df.style.format ({ c : "{:20,.3f}" for c in df.columns }).background_gradient(cmap='Wistia', )
R0_estimada * 1.2
HTML("<h2>Gráfico muertes diarias en Madrid, según Comunidad de Madrid </h2>")
import pandas as pd
import io
import matplotlib.dates as mdates
from matplotlib import pyplot as plt
df = df_master
chart_df=df[df.columns[-3:]].head(60)
chart_df.plot(legend=True,figsize=(13.5,9), marker='o')
plt.gca().xaxis.set_major_formatter(mdates.DateFormatter('%b-%d'))
plt.gca().xaxis.set_major_locator(mdates.DayLocator(interval=7))
plt.xticks(rotation=45)
ax = plt.gca()
plt.setp(ax.get_xminorticklabels(), visible=False)
ax.set_title("Muertes diarias COVID 19, media movil "+str(VENTANA_MEDIA_MOVIL)+" dias. Fuente: Comunidad de Madrid")
ax.set_ylim(ymin=0)
plt.show()
from IPython.display import display, HTML
HTML("<h2>Comparamos los datos de hoy, de hace una semana y de un mes </h2>")
from matplotlib import colors
def background_gradient(s, m, M, cmap='PuBu', low=0, high=0):
rng = M - m
norm = colors.Normalize(m - (rng * low),
M + (rng * high))
normed = norm(s.values)
c = [colors.rgb2hex(x) for x in plt.cm.get_cmap(cmap)(normed)]
return ['background-color: %s' % color for color in c]
df = df_master
df.style.format ({ c : "{:20,.0f}" for c in df.columns }).background_gradient(cmap='Wistia', subset= df.columns[-3:] )
df = df_master
pd.concat([df.head(1).tail(1) , df.head(8).tail(1) , df.head(30).tail(1)]).astype(int)[['MA HOSPITALES hoy','MA CENTROS SOCIOSANITARIOS hoy','MA FALLECIDOS TOTALES hoy']].style.format ({ c : "{:20,.0f}" for c in df.columns }).background_gradient(cmap='Wistia', subset= df.columns[-3:] )
from IPython.display import display, HTML
HTML("<h2>Muertes medias diarias, últimos 7 días, con datos</h2>")
from datetime import date
df = df_master
inicio_crisis = df.head(7).index[6]
df=df.head(7)
dia_mas_reciente = df.index[0]
dias_transcurridos_inicio_crisis = dia_mas_reciente - inicio_crisis
df = pd.DataFrame((df.head(1).max(axis=0) - df.tail(1).max(axis=0) ) / dias_transcurridos_inicio_crisis.days ).T[['HOSPITALES','DOMICILIOS','CENTROS SOCIOSANITARIOS','OTROS LUGARES','FALLECIDOS TOTALES']]
df.style.format ({ c : "{:20,.0f}" for c in df.columns }).background_gradient(cmap='Wistia' )
HTML("<h2>Muertes medias diarias desde que la comunidad de Madrid publica datos</h2>")
# Calculamos los incrementos medios, desde que tenemos fechas
df = df_master
df = pd.DataFrame((df.head(1).max(axis=0) - df.tail(1).max(axis=0) ) / df.shape[0] ).T[['HOSPITALES','DOMICILIOS','CENTROS SOCIOSANITARIOS','OTROS LUGARES','FALLECIDOS TOTALES']]
df.style.format ({ c : "{:20,.0f}" for c in df.columns }).background_gradient(cmap='Wistia' )
from tabula import read_pdf
from IPython.display import display, HTML
import os
import pandas as pd
import glob
import re
from tqdm.notebook import tqdm
import warnings
import os.path
fecha="201005"
import os
file_path = '../data/'+fecha+'_cam_covid19.pdf'
if not os.path.isfile(file_path):
file_path = '../data/'+fecha+'cam_covid19.pdf'
#print("Analizando:" + file_path)
```
| github_jupyter |
**This notebook is an exercise in the [Data Visualization](https://www.kaggle.com/learn/data-visualization) course. You can reference the tutorial at [this link](https://www.kaggle.com/alexisbcook/line-charts).**
---
In this exercise, you will use your new knowledge to propose a solution to a real-world scenario. To succeed, you will need to import data into Python, answer questions using the data, and generate **line charts** to understand patterns in the data.
## Scenario
You have recently been hired to manage the museums in the City of Los Angeles. Your first project focuses on the four museums pictured in the images below.

You will leverage data from the Los Angeles [Data Portal](https://data.lacity.org/) that tracks monthly visitors to each museum.

## Setup
Run the next cell to import and configure the Python libraries that you need to complete the exercise.
```
import pandas as pd
pd.plotting.register_matplotlib_converters()
import matplotlib.pyplot as plt
%matplotlib inline
import seaborn as sns
print("Setup Complete")
```
The questions below will give you feedback on your work. Run the following cell to set up the feedback system.
```
# Set up code checking
import os
if not os.path.exists("../input/museum_visitors.csv"):
os.symlink("../input/data-for-datavis/museum_visitors.csv", "../input/museum_visitors.csv")
from learntools.core import binder
binder.bind(globals())
from learntools.data_viz_to_coder.ex2 import *
print("Setup Complete")
```
## Step 1: Load the data
Your first assignment is to read the LA Museum Visitors data file into `museum_data`. Note that:
- The filepath to the dataset is stored as `museum_filepath`. Please **do not** change the provided value of the filepath.
- The name of the column to use as row labels is `"Date"`. (This can be seen in cell A1 when the file is opened in Excel.)
To help with this, you may find it useful to revisit some relevant code from the tutorial, which we have pasted below:
```python
# Path of the file to read
spotify_filepath = "../input/spotify.csv"
# Read the file into a variable spotify_data
spotify_data = pd.read_csv(spotify_filepath, index_col="Date", parse_dates=True)
```
The code you need to write now looks very similar!
```
# Path of the file to read
museum_filepath = "../input/museum_visitors.csv"
# Fill in the line below to read the file into a variable museum_data
museum_data = pd.read_csv(museum_filepath, index_col='Date', parse_dates=True)
# Run the line below with no changes to check that you've loaded the data correctly
step_1.check()
# Uncomment the line below to receive a hint
#step_1.hint()
# Uncomment the line below to see the solution
#step_1.solution()
```
## Step 2: Review the data
Use a Python command to print the last 5 rows of the data.
```
# Print the last five rows of the data
museum_data.tail() # Your code here
```
The last row (for `2018-11-01`) tracks the number of visitors to each museum in November 2018, the next-to-last row (for `2018-10-01`) tracks the number of visitors to each museum in October 2018, _and so on_.
Use the last 5 rows of the data to answer the questions below.
```
# Fill in the line below: How many visitors did the Chinese American Museum
# receive in July 2018?
ca_museum_jul18 = museum_data.loc['2018-07-01', 'Chinese American Museum']
# Fill in the line below: In October 2018, how many more visitors did Avila
# Adobe receive than the Firehouse Museum?
avila_oct18 = museum_data.loc['2018-10-01', 'Avila Adobe'] - museum_data.loc['2018-10-01', 'Firehouse Museum']
# Check your answers
step_2.check()
# Lines below will give you a hint or solution code
#step_2.hint()
#step_2.solution()
```
## Step 3: Convince the museum board
The Firehouse Museum claims they ran an event in 2014 that brought an incredible number of visitors, and that they should get extra budget to run a similar event again. The other museums think these types of events aren't that important, and budgets should be split purely based on recent visitors on an average day.
To show the museum board how the event compared to regular traffic at each museum, create a line chart that shows how the number of visitors to each museum evolved over time. Your figure should have four lines (one for each museum).
> **(Optional) Note**: If you have some prior experience with plotting figures in Python, you might be familiar with the `plt.show()` command. If you decide to use this command, please place it **after** the line of code that checks your answer (in this case, place it after `step_3.check()` below) -- otherwise, the checking code will return an error!
```
# Line chart showing the number of visitors to each museum over time
# Set the width and height of the figure
plt.figure(figsize=(12,6))
# Line chart showing the number of visitors to each museum over time
sns.lineplot(data=museum_data)
# Add title
plt.title("Monthly Visitors to Los Angeles City Museums")
# Check your answer
step_3.check()
# Lines below will give you a hint or solution code
#step_3.hint()
#step_3.solution_plot()
```
## Step 4: Assess seasonality
When meeting with the employees at Avila Adobe, you hear that one major pain point is that the number of museum visitors varies greatly with the seasons, with low seasons (when the employees are perfectly staffed and happy) and also high seasons (when the employees are understaffed and stressed). You realize that if you can predict these high and low seasons, you can plan ahead to hire some additional seasonal employees to help out with the extra work.
#### Part A
Create a line chart that shows how the number of visitors to Avila Adobe has evolved over time. (_If your code returns an error, the first thing that you should check is that you've spelled the name of the column correctly! You must write the name of the column exactly as it appears in the dataset._)
```
# Line plot showing the number of visitors to Avila Adobe over time
plt.figure(figsize=(12, 6))
sns.lineplot(data=museum_data['Avila Adobe'])
plt.title('Monthly Visitors to Avila Adobe')
plt.xlabel('Date')
# Check your answer
step_4.a.check()
# Lines below will give you a hint or solution code
#step_4.a.hint()
#step_4.a.solution_plot()
```
#### Part B
Does Avila Adobe get more visitors:
- in September-February (in LA, the fall and winter months), or
- in March-August (in LA, the spring and summer)?
Using this information, when should the museum staff additional seasonal employees?
```
#step_4.b.hint()
# Check your answer (Run this code cell to receive credit!)
step_4.b.solution()
```
# Keep going
Move on to learn about **[bar charts and heatmaps](https://www.kaggle.com/alexisbcook/bar-charts-and-heatmaps)** with a new dataset!
---
*Have questions or comments? Visit the [Learn Discussion forum](https://www.kaggle.com/learn-forum/161291) to chat with other Learners.*
| github_jupyter |
# Introduction
This is a quick introduction to the tradebook module. The tradebook is just a log of trades that shows your positions and values based on the trades you have done and provide you a few helper methods. This provides a flexible approach for simulating trades based on an event based system or a system where you iterate through each line as a separate observation.
Caveat
--------
**This is not an orderbook.** All trades are assumed to be executed and its kept as simple as possible.
## Initialize a tradebook
```
import pandas as pd
from fastbt.tradebook import TradeBook
tb = TradeBook()
```
### Add some trades
To add trades to a tradebook, you need 5 mandatory parameters
* timestamp - could be a string or an id; but datetime or pandas timestamp preferred
* symbol - the security symbol or asset code
* price - float/number
* qty - float/number
* order - **B for BUY and S for SELL**
Just use the `add_trade` method to add trades. Its the only method to add trades. You can include any other keyword arguments to add additional information to trades.
```
# Let's add a few trades
tb.add_trade(pd.to_datetime('2019-02-01'), 'AAA', 100, 100, 'B')
tb.add_trade(pd.to_datetime('2019-02-02'), 'AAA', 102, 100, 'B')
tb.add_trade(pd.to_datetime('2019-02-02'), 'BBB', 1000, 15, 'S')
```
### Get some information
Internally all data is represented as dictionaries
Use
* `tb.positions` to get the positions for all stocks
* `tb.values` to get the values
* `tb.trades` for trades
To get these details for a single stock use, `tb.positions.get()`
```
# Print tradebook summary
tb # Shows that you have made 3 trades and 2 of the positions are still open
# Get positions
tb.positions # negative indicates short position
# Get position for a particular stock
print(tb.positions.get('AAA'))
# Get the current value
tb.values # Negative values indicate cash outflow
# Get the trades
tb.trades.get('AAA')
# Get all your trades
tb.all_trades
# A few helper methods
print(tb.o) # Number of open positions
print(tb.l) # Number of long positions
print(tb.s) # Number of short positions
```
### Something to look out for
* A position of zero indicates that all trades are settled
* A positive position indicates holdings
* A negative position indicates short selling
* Conversely, a positive value indicates money received from short selling and a negative value indicates cash outflow for buying holding
* If all positions are zero, then the corresponding values indicate profit or loss
* Trades are represented as a dictionary with keys being the symbol and values being the list of all trades. So to get the first trade, use `tb.trades[symbol][0]`
Let's try out by closing all existing positions
```
tb.positions, tb.values
# Close existing positions
tb.add_trade(pd.to_datetime('2019-03-05'), 'AAA', 105, 200, 'S', info='exit')
tb.add_trade(pd.to_datetime('2019-03-05'), 'BBB', 1010, 15, 'B', info='exit')
tb.positions, tb.values
```
> You could now see that both the positions are closed but you got a profit on AAA and a loss on BBB
```
# Summing up total profit
print(tb)
tb.values.values()
```
> And you could nicely load them up in a dataframe and see your additional info column added
```
pd.DataFrame(tb.all_trades).sort_values(by='ts')
```
# Creating a strategy
Let's create a simple strategy for bitcoin and let's see how it works. This is a long only strategy
> **ENTER** when 7 day simple moving average (SMA) is greater than 30 day SMA and **EXIT** when 7 day SMA is less than 30 day SMA
Other info
-----------
* Invest $10000 for each trade
* Hold only one position at a single time (BUY only, no reversals)
* If you are already holding a position, check for the exit rule
* SMA is calculated on OPEN price and its assumed that you buy and sell at the open price
The sample file already has the columns sma7 and sma30
```
df = pd.read_csv('data/BTC.csv', parse_dates=['date'])
# We would be using standard Python csv library
import csv
filename = 'data/BTC.csv' # File available in data directory
btc = TradeBook()
capital = 10000 # this is fixed
with open(filename) as csvfile:
reader = csv.DictReader(csvfile)
for row in reader:
# Convert to floats since by default csv reads everything as string
sma7 = float(row['sma7']) + 0
sma30 = float(row['sma30']) + 0
price = float(row['open']) + 0
# Check for entry rule and existing position
# Enter only if you have no existing position
if (sma7 > sma30) and (btc.l == 0):
qty = int(capital/price)
btc.add_trade(row['date'], 'BTC', price, qty, 'B')
# Check for exit
if btc.positions['BTC'] > 0:
qty = btc.positions['BTC'] # Get the present position
if sma7 < sma30:
btc.add_trade(row['date'], 'BTC', price , qty, 'S')
btc, btc.values
```
**Hurray!** You have made a profit and still hold a position. But its not surprising since bitcoin has increased twenty fold during this period. Let's do some analytics for fun.
Beware, you are not taking commission and transaction costs into account
```
trades = pd.DataFrame(btc.all_trades)
trades['ts'] = pd.to_datetime(trades['ts'])
trades['year'] = trades['ts'].dt.year
trades['values'] = trades['qty'] * trades['price']
trades.groupby(['year', 'order']).agg({'qty': sum, 'values': sum}).unstack()
```
Looks, 2013 and 2017 seemed to be really good years
| github_jupyter |
# Exercise 05
Montecarlo simulation of the 4d Ising model using the cluster algorithm.
A sample python code to do cluster updates of the d-dimensional Ising model.
```
## union-find routines
# returns index of cluster to which spin at index x belongs (x is an integer)
def find_cluster(cluster, x):
while(x != cluster[x]):
# compress path: set parent of x to its grandparent
cluster[x] = cluster[cluster[x]];
# climb tree
x = cluster[x];
return x
# connect clusters to which spins at x and y belong
def connect_clusters(cluster, cluster_size, x, y):
x_cluster = find_cluster(cluster, x)
y_cluster = find_cluster(cluster, y)
# if not already connected, then connect them
if (x_cluster != y_cluster):
# connect smaller tree to root of larger tree
if (cluster_size[x] < cluster_size[y]):
cluster[x_cluster] = y_cluster;
cluster_size[y_cluster] += cluster_size[x_cluster];
else:
cluster[y_cluster] = x_cluster;
cluster_size[x_cluster] += cluster_size[y_cluster];
## d-dimensional lattice with periodic boundary conditions
import numpy as np
# shape is a tuple of lattice dimensions, e.g. (8,8,8,8) would be a 8^4 lattice
# ix is an integer index that refers to a site on this lattice
# mu is a dimension, starting at 0, e.g. could be 0,1,2 or 3 on an L^4 lattice
# returns index of neighbour to site ix in +mu direction
def iup(shape, ix, mu):
# convert flat index ix to d-dimensional tuple of indices
x_d = np.unravel_index(ix, shape)
# convert to array, and add 1 to mu-component
tmpx_d = np.array(x_d)
tmpx_d[mu] += 1
# convert back to tuple
y_d = tuple(tmpx_d)
# convert tuple back to flat index iy with pbcs (mode='wrap')
iy = np.ravel_multi_index(y_d, shape, mode='wrap')
return iy
## cluster update routine
import numpy as np
# spins is a 1d array containing all the spins
# shape is a tuple with the dimensions of the lattice, e.g. (4,4) would be a 2d 4x4 lattice
# beta is the lattice coupling
def cluster_update(spins, shape, beta):
# make array of cluster values for each spin
# initially each is its own cluster
cluster = np.empty_like(spins)
for i in range(0, cluster.size):
cluster[i] = i
# each cluster has initial size 1
cluster_size = np.ones_like(cluster)
# construct clusters
# sum over all sites..
for ix in range(0, spins.size):
# ..and over all directions
for mu in range(0, len(shape)):
# find index of neighbouring site
iy = iup(shape, ix, mu)
# construct bond (or not) between sites
if(spins[ix] == spins[iy]):
if(np.random.uniform() > np.exp(-2.0*beta)):
connect_clusters(cluster, cluster_size, ix, iy)
# make random spins for each cluster
cluster_spins = 2 * np.random.randint(2, size=cluster.shape) - 1
# assign cluster spins to sites
for i in range(0, cluster.size):
spins[i] = cluster_spins[find_cluster(cluster, i)]
## observables
import numpy as np
# returns energy normalised by the volume (number of sites)
def E(spins, shape):
tmpE = 0
for ix in range(0, spins.size):
for mu in range(0, len(shape)):
iy = iup(shape, ix, mu)
tmpE += spins[ix] * spins[iy]
return -tmpE / spins.size
# returns absolute magnetisation normalised by the volume (number of sites)
def M(spins):
return np.fabs(np.sum(spins)/spins.size)
## routine to do a simulation and plot E & M
import matplotlib.pyplot as plt
import numpy as np
# shape is a tuple of lattice dimensions, eg (4,4,4,4) or (8,8)
# beta is a list or array of beta values to simulate
# n_therm is how many updates to discard for thermalisation
# n_updates is how many updates to then do with measurements
# plots resulting energy and magnetisation vs beta
def do_simulation(shape, beta_list, n_therm, n_updates):
# randomly set all spins to either 1 or -1:
spins = 2 * np.random.randint(2, size=np.prod(shape)) - 1
# do simulation
obs_beta = []
for beta in beta_list:
for i in range (0, n_therm):
cluster_update(spins, shape, beta)
m = []
e = []
for i in range (0, n_updates):
cluster_update(spins, shape, beta)
m.append(M(spins))
e.append(E(spins, shape))
m = np.array(m)
e = np.array(e)
obs_beta.append([beta, np.average(m), np.std(m)/np.sqrt(m.size), np.average(e), np.std(e)/np.sqrt(m.size)])
obs_beta = np.array(obs_beta)
# critical beta line for plots
if(len(shape)==4):
beta_critical = 0.1496947
elif(len(shape)==2):
beta_critical = 0.440687
else:
beta_critical = 0
# plot energy
plt.rcParams["figure.figsize"] = (20,10)
plt.figure()
plt.errorbar(obs_beta[:,0], obs_beta[:,3], yerr=obs_beta[:,4])
plt.axvline(x=beta_critical, label="critical beta", linestyle='--', color="grey")
plt.ylabel('E/N')
plt.title(str(shape)+' lattice: Energy vs Beta')
plt.show()
# plot magnetisation
plt.rcParams["figure.figsize"] = (20,10)
plt.figure()
plt.errorbar(obs_beta[:,0], obs_beta[:,1], yerr=obs_beta[:,2])
plt.axvline(x=beta_critical, label="critical beta", linestyle='--', color="grey")
plt.ylabel('|M|/N')
plt.title(str(shape)+' lattice: Magnetisation vs Beta')
plt.show()
# simulate a 2d 8x8 lattice for a range of beta, 200 thermalisation updates, 2000 measurement updates
do_simulation((8,8), np.arange(0,1,0.03), 200, 2000)
# simulate a 4d 4^4 lattice for a range of beta, 50 thermalisation updates, 200 measurement updates
do_simulation((4,4,4,4), np.arange(0,0.31,0.03), 50, 200)
```
| github_jupyter |
# Scientific Computing with Python (Second Edition)
# Chapter 03
*We start by importing all from Numpy. As explained in Chapter 01 the examples are written assuming this import is initially done.*
```
from numpy import *
```
## 3.1 Lists
```
L = ['a', 20.0, 5]
M = [3,['a', -3.0, 5]]
L[1] # returns 20.0
L[0] # returns 'a'
M[1] # returns ['a',-3.0,5]
M[1][2] # returns 5
L=list(range(4))
# generates a list with four elements: [0, 1, 2 ,3]
L
L=list(range(17,29,4))
# generates [17, 21, 25]
L
len(L) # returns 3
```
### 3.1.1 Slicing
```
L = ['C', 'l', 'o', 'u', 'd', 's']
L[1:5] # remove one element and take four from there:
# returns ['l', 'o', 'u', 'd']
L = ['C', 'l', 'o', 'u', 'd', 's']
L[1:] # ['l', 'o', 'u', 'd', 's']
L[:5] # ['C', 'l', 'o', 'u', 'd']
L[:] # the entire list
L = ['C', 'l', 'o', 'u', 'd', 's']
L[-2:] # ['d', 's']
L[:-2] # ['C', 'l', 'o', 'u']
L = list(range(4)) # [0, 1, 2, 3]
L[4] # IndexError: list index out of range
L[1:100] # same as L[1:]
L[-100:-1] # same as L[:-1]
L[-100:100] # same as L[:]
L[5:0] # empty list []
L[-2:2] # empty list []
```
As we imported `numpy`, we have to make sure that we take the built-in command `sum` and not the one from `numpy`.
```
sum=__builtin__.sum
a = [1,2,3]
for iteration in range(4):
print(sum(a[0:iteration-1]))
L = list(range(100))
L[:10:2] # [0, 2, 4, 6, 8]
L[::20] # [0, 20, 40, 60, 80]
L[10:20:3] # [10, 13, 16, 19]
L[20:10:-3] # [20, 17, 14, 11]
L = [1, 2, 3]
R = L[::-1] # L is not modified
R # [3, 2, 1]
```
### 3.1.2 Altering lists
```
L = ['a', 1, 2, 3, 4]
L
L[2:3] = [] # ['a', 1, 3, 4]
L
L[3:] = [] # ['a', 1, 3]
L
L[1:1] = [1000, 2000] # ['a', 1000, 2000, 1, 3]
L
L = [1, -17]
M = [-23.5, 18.3, 5.0]
L + M # gives [1, -17, 23.5, 18.3, 5.0]
n = 3
n * [1.,17,3] # gives [1., 17, 3, 1., 17, 3, 1., 17, 3]
[0] * 5 # gives [0,0,0,0,0]
```
### 3.1.3 Belonging to a list
```
L = ['a', 1, 'b', 2]
'a' in L # True
3 in L # False
4 not in L # True
```
### 3.1.4 List methods
```
L = [1, 2, 3]
L.reverse() # the list L is now reversed
L # [3, 2, 1]
L=[3, 4, 4, 5]
newL = L.sort()
print(newL[0])
L = [0, 1, 2, 3, 4]
L.append(5) # [0, 1, 2, 3, 4, 5]
L
L.reverse() # [5, 4, 3, 2, 1, 0]
L
L.sort() # [0, 1, 2, 3, 4, 5]
L
L.remove(0) # [1, 2, 3, 4, 5]
L
L.pop() # [1, 2, 3, 4]
L
L.pop() # [1, 2, 3]
L
L.extend(['a','b','c']) # [1, 2, 3, 'a', 'b', 'c']
L
```
### 3.1.5 Merging lists – `zip`
```
ind = [0,1,2,3,4]
color = ["red", "green", "blue", "alpha"]
list(zip(color,ind)) # gives [('red', 0), ('green', 1), ('blue', 2), ('alpha', 3)]
```
### 3.1.6 List comprehension
```
L = [2, 3, 10, 1, 5]
L2 = [x*2 for x in L] # [4, 6, 20, 2, 10]
L2
L3 = [x*2 for x in L if 4 < x <= 10] # [20, 10]
L3
M = [[1,2,3],[4,5,6]]
flat = [M[i][j] for i in range(2) for j in range(3)]
# returns [1, 2, 3, 4, 5, 6]
flat
```
## 3.2 A quick glance at the concept of arrays
```
v = array([1.,2.,3.])
A = array([[1.,2.,3.],[4.,5.,6.]])
v[2] # returns 3.0
A[1,2] # returns 6.0
M = array([[1.,2.],[3.,4.]])
v = array([1., 2., 3.])
v[0] # 1
v[:2] # array([1.,2.])
M[0,1] # 2
v[:2] = [10, 20] # v is now array([10., 20., 3.])
v
len(v) # 3
```
## 3.3 Tuples
```
my_tuple = 1, 2, 3 # our first tuple
my_tuple = (1, 2, 3) # the same
my_tuple = 1, 2, 3, # again the same
my_tuple
len(my_tuple) # 3, same as for lists
my_tuple[0] = 'a' # error! tuples are immutable
1, 2 == 3, 4 # returns (1, False, 4)
(1, 2) == (3, 4) # returns False
singleton = 1, # note the comma
len(singleton) # 1
singleton = (1,) # this creates the same tuple
```
### 3.3.1 Packing and unpacking variables
```
a, b = 0, 1 # a gets 0 and b gets 1
a, b = [0, 1] # exactly the same effect
(a, b) = 0, 1 # same
[a,b] = [0,1] # same thing
print(a, b)
a, b = b, a
print(a,b)
```
## 3.4 Dictionaries
```
truck_wheel = {'name':'wheel','mass':5.7,
'Ix':20.0,'Iy':1.,'Iz':17.,
'center of mass':[0.,0.,0.]}
truck_wheel
truck_wheel['name'] # returns 'wheel'
truck_wheel['mass'] # returns 5.7
truck_wheel['Ixy'] = 0.0
truck_wheel
truck_wheel = dict([('name','wheel'),('mass',5.7),
('Ix',20.0), ('Iy',1.), ('Iz',17.),
('center of mass',[0.,0.,0.])])
truck_wheel
```
### 3.4.2 Looping over dictionaries
```
for key in truck_wheel.keys():
print(key) # prints (in any order) 'Ix', 'Iy', 'name',...
for key in truck_wheel:
print(key) # prints (in any order) 'Ix', 'Iy', 'name',...
for value in truck_wheel.values():
print(value)
# prints (in any order) 1.0, 20.0, 17.0, 'wheel', ...
for item in truck_wheel.items():
print(item)
# prints (in any order) ('Iy', 1.0), ('Ix, 20.0),...
```
## 3.5 Sets
```
A = {1,2,3,4}
B = {5}
B
C = A.union(B) # returns {1, 2, 3, 4, 5}
C
D = A.intersection(C) # returns {1, 2, 3, 4}
D
E = C.difference(A) # returns {5}
E
5 in C # returns True
A = {1,2,3,3,3}
B = {1,2,3}
A == B # returns True
A = {1,2,3}
B = {1,3,2}
A == B # returns True
A={1,2,3,4}
A.union({5})
A.intersection({2,4,6}) # returns {2, 4}
{2,4}.issubset({1,2,3,4,5}) # returns True
{1,2,3,4,5}.issuperset({2,4}) # returns True
empty_set=set([])
```
## 3.6 Container conversions
No code.
## 3.7 Checking the type of a variable
```
label = 'local error'
type(label) # returns str
x = [1, 2] # list
type(x) # returns list
isinstance(x, list) # True
test = True
isinstance(test, bool) # True
isinstance(test, int) # True
type(test) == int # False
type(test) == bool # True
if isinstance(test, int):
print("The variable is an integer")
```
| github_jupyter |
<img src="piwrf_logo.gif" width="350" align="center">
Pi-WRF is a community driven project to facilitate education/learning related to numerical weather prediction. The [Pi-WRF GitHub](https://github.com/NCAR/pi-wrf) repository contains the necessary components and instructions to both use and contribute to the Pi-WRF project.
```
import pytz
import datetime
import numpy as np
from tzwhere import tzwhere
import ipywidgets as widgets
from ipyleaflet import (Map,
basemaps,
Marker)
```
# Domain selection:
<p>Below is a map with a marker that is positioned at Boulder Colorado, the location of NCAR. You may zoom in or out using the +/- in the upper left of the map. You may grab and drag the blue marker to any location where you would like to center your gridbox.</p>
```
# coordinates default to Boulder, CO
center = [40.0150, -105.2705]
zoom = 2
n = Map(basemap=basemaps.OpenStreetMap.Mapnik, center=center, zoom=zoom)
marker = Marker(location=center, draggable=True)
n.add_layer(marker);
display(n)
```
# Datetime range selection:
<p>The cell below automatically generates a datetime range based on the current time of the timezone for the map marker. The times are rounded according to the model's expected input. Current times are rounded down to the most recent appropriate hour for a start time. The end time adds six hours to the start time. It is possible to manually set the start and end times at the end of the cell if desired.</p>
```
# function for rounding times
def round_hour(x, base=6):
return x // base * base
# time range is generated automatically based on current time at time zone of map marker
from tzwhere import tzwhere
tzwhere = tzwhere.tzwhere()
timezone_str = tzwhere.tzNameAt(marker.location[0], marker.location[1])
timezone = pytz.timezone(timezone_str)
today = datetime.datetime.now(timezone)
now = today.time()
step = now
tomorrow = today
now = now.replace(hour = round_hour(now.hour))
start_time = str(today.strftime('%m')) + "-" \
+ str(today.strftime('%d')) + "-" \
+ str(today.year) + "-" \
+ str(now.strftime('%H'))
if now.hour >= 18:
step = step.replace(hour = 0)
tomorrow = tomorrow.replace(day = tomorrow.day + 1)
else:
step = now.replace(hour = (now.hour + 6))
end_time = str(today.strftime('%m')) + "-" \
+ str(tomorrow.strftime('%d')) + "-" \
+ str(today.year) + "-" \
+ step.strftime('%H')
# you may enter the start_time and end_time manually here
# you just neec to uncomment the start_time and end_time lines and modify them as desired
# format: MM-DD-YYYY-HH
# where hour may be 00, 06, 12, or 18
# start_time = '07-24-2021-12'
# end_time = '07-24-2021-18'
# print(start_time)
# print(end_time)
%%bash -s "$start_time" "$end_time"
sed -i /userstartdate=/c\\userstartdate="$1" /pi-wrf/run_wrf
sed -i /userenddate=/c\\userenddate="$2" /pi-wrf/run_wrf
# gridbox dimensions
gridbox_width = 5
gridbox_height = 4
# generate variables for domain based on map marker
xlim = np.sort(np.array([(marker.location[1]-gridbox_width/2),(marker.location[1]+gridbox_width/2)]))
print(xlim) # longitude bounds
ylim = np.sort(np.array([(marker.location[0]-gridbox_height/2),(marker.location[0]+gridbox_height/2)]))
print(ylim) # latitude bounds
# call set_domain with arguments based on map marker
%cd ~/../pi-wrf/pi_wrf/pi_wrf
%pwd
%ls
from set_domain import set_domain
set_domain(xlim, ylim)
```
# Initiate model
<p>The model is initiated from the cell below. The above cells should have already been executed so the appropriate inputs will be available for the model. The model will take a few minutes to run. You should wait until it finishes running before proceeding to the following cells. The cell's number will show an asterisk '*' while it is still running.</p>
```
%%bash
cd ~/../pi-wrf/pi_wrf/pi_wrf
../../run_wrf
```
# Model Output:
<p>The cells below demonstrate how to retrieve specific plots based on the model output. When the cells are executed, the contents of the Output directory are displayed. These contain the series of plots generated from the model output. The first cell defaults to showing the Your_Domain_Relative plot. The second cell defaults to showing the hourly-temperature_06. You may access any of the available plots in the directory by changing the filename in either of these cells.</p>
```
from IPython.display import Image
#Image(filename='test.png')
%cd ~/../pi-wrf/Output
%pwd
%ls -ls
Image(filename='Your_Domain_Relative.png')
from IPython.display import Image
#Image(filename='test.png')
#%cd ../../Output
%pwd
%ls -ls
Image(filename='hourly-temperature_06.png')
from IPython.display import Image
#Image(filename='test.png')
#%cd ../../Output
%pwd
%ls -ls
Image(filename='hourly-wind_06.png')
```
| github_jupyter |
# Creating Estimators in tf.estimator with Keras
This tutorial covers how to create your own training script using the building
blocks provided in `tf.keras`, which will predict the ages of
[abalones](https://en.wikipedia.org/wiki/Abalone) based on their physical
measurements. You'll learn how to do the following:
* Construct a custom model function
* Configure a neural network using `tf.keras`
* Choose an appropriate loss function from `tf.losses`
* Define a training op for your model
* Generate and return predictions
## An Abalone Age Predictor
It's possible to estimate the age of an
[abalone](https://en.wikipedia.org/wiki/Abalone) (sea snail) by the number of
rings on its shell. However, because this task requires cutting, staining, and
viewing the shell under a microscope, it's desirable to find other measurements
that can predict age.
The [Abalone Data Set](https://archive.ics.uci.edu/ml/datasets/Abalone) contains
the following
[feature data](https://archive.ics.uci.edu/ml/machine-learning-databases/abalone/abalone.names)
for abalone:
| Feature | Description |
| -------------- | --------------------------------------------------------- |
| Length | Length of abalone (in longest direction; in mm) |
| Diameter | Diameter of abalone (measurement perpendicular to length; in mm)|
| Height | Height of abalone (with its meat inside shell; in mm) |
| Whole Weight | Weight of entire abalone (in grams) |
| Shucked Weight | Weight of abalone meat only (in grams) |
| Viscera Weight | Gut weight of abalone (in grams), after bleeding |
| Shell Weight | Weight of dried abalone shell (in grams) |
The label to predict is number of rings, as a proxy for abalone age.
### Set up the environment
```
import os
import sagemaker
from sagemaker import get_execution_role
sagemaker_session = sagemaker.Session()
role = get_execution_role()
```
### Upload the data to a S3 bucket
```
inputs = sagemaker_session.upload_data(path='data', key_prefix='data/DEMO-abalone')
```
**sagemaker_session.upload_data** will upload the abalone dataset from your machine to a bucket named **sagemaker-{your aws account number}**, if you don't have this bucket yet, sagemaker_session will create it for you.
## Complete source code
Here is the full code for the network model:
```
!cat 'abalone.py'
```
## Defining a `model_fn`
The script above implements a `model_fn` as the function responsible for implementing the model for training, evaluation, and prediction. The next section covers how to implement a `model_fn` using `Keras layers`.
### Constructing the `model_fn`
The basic skeleton for an `model_fn` looks like this:
```python
def model_fn(features, labels, mode, params):
# Logic to do the following:
# 1. Configure the model via TensorFlow or Keras operations
# 2. Define the loss function for training/evaluation
# 3. Define the training operation/optimizer
# 4. Generate predictions
# 5. Return predictions/loss/train_op/eval_metric_ops in EstimatorSpec object
return EstimatorSpec(mode, predictions, loss, train_op, eval_metric_ops)
```
The `model_fn` must accept three arguments:
* `features`: A dict containing the features passed to the model via
`input_fn`.
* `labels`: A `Tensor` containing the labels passed to the model via
`input_fn`. Will be empty for `predict()` calls, as these are the values the
model will infer.
* `mode`: One of the following tf.estimator.ModeKeys string values
indicating the context in which the model_fn was invoked:
* `tf.estimator.ModeKeys.TRAIN` The `model_fn` was invoked in training
mode, namely via a `train()` call.
* `tf.estimator.ModeKeys.EVAL`. The `model_fn` was invoked in
evaluation mode, namely via an `evaluate()` call.
* `tf.estimator.ModeKeys.PREDICT`. The `model_fn` was invoked in
predict mode, namely via a `predict()` call.
`model_fn` may also accept a `params` argument containing a dict of
hyperparameters used for training (as shown in the skeleton above).
The body of the function performs the following tasks (described in detail in the
sections that follow):
* Configuring the model for the abalone predictor. This will be a neural
network.
* Defining the loss function used to calculate how closely the model's
predictions match the target values.
* Defining the training operation that specifies the `optimizer` algorithm to
minimize the loss values calculated by the loss function.
The `model_fn` must return a tf.estimator.EstimatorSpec
object, which contains the following values:
* `mode` (required). The mode in which the model was run. Typically, you will
return the `mode` argument of the `model_fn` here.
* `predictions` (required in `PREDICT` mode). A dict that maps key names of
your choice to `Tensor`s containing the predictions from the model, e.g.:
```python
predictions = {"results": tensor_of_predictions}
```
In `PREDICT` mode, the dict that you return in `EstimatorSpec` will then be
returned by `predict()`, so you can construct it in the format in which
you'd like to consume it.
* `loss` (required in `EVAL` and `TRAIN` mode). A `Tensor` containing a scalar
loss value: the output of the model's loss function (discussed in more depth
later in [Defining loss for the model](https://github.com/tensorflow/tensorflow/blob/eb84435170c694175e38bfa02751c3ef881c7a20/tensorflow/docs_src/extend/estimators.md#defining-loss)) calculated over all
the input examples. This is used in `TRAIN` mode for error handling and
logging, and is automatically included as a metric in `EVAL` mode.
* `train_op` (required only in `TRAIN` mode). An Op that runs one step of
training.
* `eval_metric_ops` (optional). A dict of name/value pairs specifying the
metrics that will be calculated when the model runs in `EVAL` mode. The name
is a label of your choice for the metric, and the value is the result of
your metric calculation. The tf.metrics
module provides predefined functions for a variety of common metrics. The
following `eval_metric_ops` contains an `"accuracy"` metric calculated using
`tf.metrics.accuracy`:
```python
eval_metric_ops = {
"accuracy": tf.metrics.accuracy(labels, predictions)
}
```
If you do not specify `eval_metric_ops`, only `loss` will be calculated
during evaluation.
### Configuring a neural network with `keras layers`
Constructing a [neural
network](https://en.wikipedia.org/wiki/Artificial_neural_network) entails
creating and connecting the input layer, the hidden layers, and the output
layer.
The input layer of the neural network then must be connected to one or more
hidden layers via an [activation
function](https://en.wikipedia.org/wiki/Activation_function) that performs a
nonlinear transformation on the data from the previous layer. The last hidden
layer is then connected to the output layer, the final layer in the model.
`tf.layers` provides the `tf.layers.dense` function for constructing fully
connected layers. The activation is controlled by the `activation` argument.
Some options to pass to the `activation` argument are:
* `tf.nn.relu`. The following code creates a layer of `units` nodes fully
connected to the previous layer `input_layer` with a
[ReLU activation function](https://en.wikipedia.org/wiki/Rectifier_(neural_networks)
(tf.nn.relu):
```python
hidden_layer = Dense(10, activation='relu', name='first-layer')(features)
```
* `tf.nn.relu6`. The following code creates a layer of `units` nodes fully
connected to the previous layer `hidden_layer` with a ReLU activation
function:
```python
second_hidden_layer = Dense(20, activation='relu', name='first-layer')(hidden_layer)
```
* `None`. The following code creates a layer of `units` nodes fully connected
to the previous layer `second_hidden_layer` with *no* activation function,
just a linear transformation:
```python
output_layer = Dense(1, activation='linear')(second_hidden_layer)
```
Other activation functions are possible, e.g.:
```python
output_layer = Dense(10, activation='sigmoid')(second_hidden_layer)
```
The above code creates the neural network layer `output_layer`, which is fully
connected to `second_hidden_layer` with a sigmoid activation function
(tf.sigmoid).
Putting it all together, the following code constructs a full neural network for
the abalone predictor, and captures its predictions:
```python
def model_fn(features, labels, mode, params):
"""Model function for Estimator."""
# Connect the first hidden layer to input layer
# (features["x"]) with relu activation
first_hidden_layer = Dense(10, activation='relu', name='first-layer')(features['x'])
# Connect the second hidden layer to first hidden layer with relu
second_hidden_layer = Dense(20, activation='relu', name='first-layer')(hidden_layer)
# Connect the output layer to second hidden layer (no activation fn)
output_layer = Dense(1, activation='linear')(second_hidden_layer)
# Reshape output layer to 1-dim Tensor to return predictions
predictions = tf.reshape(output_layer, [-1])
predictions_dict = {"ages": predictions}
...
```
Here, because you'll be passing the abalone `Datasets` using `numpy_input_fn`
as shown below, `features` is a dict `{"x": data_tensor}`, so
`features["x"]` is the input layer. The network contains two hidden
layers, each with 10 nodes and a ReLU activation function. The output layer
contains no activation function, and is
tf.reshape to a one-dimensional
tensor to capture the model's predictions, which are stored in
`predictions_dict`.
### Defining loss for the model
The `EstimatorSpec` returned by the `model_fn` must contain `loss`: a `Tensor`
representing the loss value, which quantifies how well the model's predictions
reflect the label values during training and evaluation runs. The tf.losses
module provides convenience functions for calculating loss using a variety of
metrics, including:
* `absolute_difference(labels, predictions)`. Calculates loss using the
[absolute-difference
formula](https://en.wikipedia.org/wiki/Deviation_statistics#Unsigned_or_absolute_deviation)
(also known as L<sub>1</sub> loss).
* `log_loss(labels, predictions)`. Calculates loss using the [logistic loss
forumula](https://en.wikipedia.org/wiki/Loss_functions_for_classification#Logistic_loss)
(typically used in logistic regression).
* `mean_squared_error(labels, predictions)`. Calculates loss using the [mean
squared error](https://en.wikipedia.org/wiki/Mean_squared_error) (MSE; also
known as L<sub>2</sub> loss).
The following example adds a definition for `loss` to the abalone `model_fn`
using `mean_squared_error()` (in bold):
```python
def model_fn(features, labels, mode, params):
"""Model function for Estimator."""
# Connect the first hidden layer to input layer
# (features["x"]) with relu activation
first_hidden_layer = Dense(10, activation='relu', name='first-layer')(features[INPUT_TENSOR_NAME])
# Connect the second hidden layer to first hidden layer with relu
second_hidden_layer = Dense(20, activation='relu')(first_hidden_layer)
# Connect the output layer to second hidden layer (no activation fn)
output_layer = Dense(1, activation='linear')(second_hidden_layer)
# Reshape output layer to 1-dim Tensor to return predictions
predictions = tf.reshape(output_layer, [-1])
predictions_dict = {"ages": predictions}
# Calculate loss using mean squared error
loss = tf.losses.mean_squared_error(labels, predictions)
...
```
Supplementary metrics for evaluation can be added to an `eval_metric_ops` dict.
The following code defines an `rmse` metric, which calculates the root mean
squared error for the model predictions. Note that the `labels` tensor is cast
to a `float64` type to match the data type of the `predictions` tensor, which
will contain real values:
```python
eval_metric_ops = {
"rmse": tf.metrics.root_mean_squared_error(
tf.cast(labels, tf.float64), predictions)
}
```
### Defining the training op for the model
The training op defines the optimization algorithm TensorFlow will use when
fitting the model to the training data. Typically when training, the goal is to
minimize loss. A simple way to create the training op is to instantiate a
`tf.train.Optimizer` subclass and call the `minimize` method.
The following code defines a training op for the abalone `model_fn` using the
loss value calculated in [Defining Loss for the Model](https://github.com/tensorflow/tensorflow/blob/eb84435170c694175e38bfa02751c3ef881c7a20/tensorflow/docs_src/extend/estimators.md#defining-loss), the
learning rate passed to the function in `params`, and the gradient descent
optimizer. For `global_step`, the convenience function
tf.train.get_global_step takes care of generating an integer variable:
```python
optimizer = tf.train.GradientDescentOptimizer(
learning_rate=params["learning_rate"])
train_op = optimizer.minimize(
loss=loss, global_step=tf.train.get_global_step())
```
### The complete abalone `model_fn`
Here's the final, complete `model_fn` for the abalone age predictor. The
following code configures the neural network; defines loss and the training op;
and returns a `EstimatorSpec` object containing `mode`, `predictions_dict`, `loss`,
and `train_op`:
```python
def model_fn(features, labels, mode, params):
"""Model function for Estimator."""
# Connect the first hidden layer to input layer
# (features["x"]) with relu activation
first_hidden_layer = Dense(10, activation='relu', name='first-layer')(features[INPUT_TENSOR_NAME])
# Connect the second hidden layer to first hidden layer with relu
second_hidden_layer = Dense(20, activation='relu')(first_hidden_layer)
# Connect the output layer to second hidden layer (no activation fn)
output_layer = Dense(1, activation='linear')(second_hidden_layer)
# Reshape output layer to 1-dim Tensor to return predictions
predictions = tf.reshape(output_layer, [-1])
# Provide an estimator spec for `ModeKeys.PREDICT`.
if mode == tf.estimator.ModeKeys.PREDICT:
return tf.estimator.EstimatorSpec(
mode=mode,
predictions={"ages": predictions})
# Calculate loss using mean squared error
loss = tf.losses.mean_squared_error(labels, predictions)
# Calculate root mean squared error as additional eval metric
eval_metric_ops = {
"rmse": tf.metrics.root_mean_squared_error(
tf.cast(labels, tf.float64), predictions)
}
optimizer = tf.train.GradientDescentOptimizer(
learning_rate=params["learning_rate"])
train_op = optimizer.minimize(
loss=loss, global_step=tf.train.get_global_step())
# Provide an estimator spec for `ModeKeys.EVAL` and `ModeKeys.TRAIN` modes.
return tf.estimator.EstimatorSpec(
mode=mode,
loss=loss,
train_op=train_op,
eval_metric_ops=eval_metric_ops)
```
# Submitting script for training
We can use the SDK to run our local training script on SageMaker infrastructure.
1. Pass the path to the abalone.py file, which contains the functions for defining your estimator, to the sagemaker.TensorFlow init method.
2. Pass the S3 location that we uploaded our data to previously to the fit() method.
```
from sagemaker.tensorflow import TensorFlow
abalone_estimator = TensorFlow(entry_point='abalone.py',
role=role,
framework_version='1.12.0',
training_steps= 100,
evaluation_steps= 100,
hyperparameters={'learning_rate': 0.001},
train_instance_count=1,
train_instance_type='ml.c4.xlarge')
abalone_estimator.fit(inputs)
```
`estimator.fit` will deploy a script in a container for training and returns the SageMaker model name using the following arguments:
* **`entry_point="abalone.py"`** The path to the script that will be deployed to the container.
* **`training_steps=100`** The number of training steps of the training job.
* **`evaluation_steps=100`** The number of evaluation steps of the training job.
* **`role`**. AWS role that gives your account access to SageMaker training and hosting
* **`hyperparameters={'learning_rate' : 0.001}`**. Training hyperparameters.
Running the code block above will do the following actions:
* deploy your script in a container with tensorflow installed
* copy the data from the bucket to the container
* instantiate the tf.estimator
* train the estimator with 10 training steps
* save the estimator model
# Submiting a trained model for hosting
The deploy() method creates an endpoint which serves prediction requests in real-time.
```
abalone_predictor = abalone_estimator.deploy(initial_instance_count=1, instance_type='ml.m4.xlarge')
```
# Invoking the endpoint
```
import tensorflow as tf
import numpy as np
prediction_set = tf.contrib.learn.datasets.base.load_csv_without_header(
filename=os.path.join('data/abalone_predict.csv'), target_dtype=np.int, features_dtype=np.float32)
data = prediction_set.data[0]
tensor_proto = tf.make_tensor_proto(values=np.asarray(data), shape=[1, len(data)], dtype=tf.float32)
abalone_predictor.predict(tensor_proto)
```
# Deleting the endpoint
```
sagemaker.Session().delete_endpoint(abalone_predictor.endpoint)
```
| github_jupyter |
# Retail Demo Store Experimentation Workshop - A/B Testing Exercise
In this exercise we will define, launch, and evaluate the results of an A/B experiment using the experimentation framework implemented in the Retail Demo Store project. If you have not already stepped through the **[3.1-Overview](./3.1-Overview.ipynb)** workshop notebook, please do so now as it provides the foundation built upon in this exercise.
Recommended Time: 30 minutes
## Prerequisites
Since this module uses the Retail Demo Store's Recommendation service to run experiments across variations that depend on the personalization features of the Retail Demo Store, it is assumed that you have either completed the [Personalization](../1-Personalization/1.1-Personalize.ipynb) workshop or those resources have been pre-provisioned in your AWS environment. If you are unsure and attending an AWS managed event such as a workshop, check with your event lead.
## Exercise 1: A/B Experiment
For the first exercise we will demonstrate how to use the A/B testing technique to implement an experiment over two implementations, or variations, of product recommendations. The first variation will represent our current implementation using the [**Default Product Resolver**](https://github.com/aws-samples/retail-demo-store/blob/master/src/recommendations/src/recommendations-service/experimentation/resolvers.py) and the second variation will use the [**Personalize Resolver**](https://github.com/aws-samples/retail-demo-store/blob/master/src/recommendations/src/recommendations-service/experimentation/resolvers.py). The scenario we are simulating is adding product recommendations powered by Amazon Personalize to home page and measuring the impact/uplift in click-throughs for products as a result of deploying a personalization strategy.
### What is A/B Testing?
A/B testing, also known as bucket or split testing, is used to compare the performance of two variations (A and B) of a single variable/experience by exposing separate groups of users to each variation and measuring user responses. An A/B experiment is run for a period of time, typically dictated by the number of users necessary to reach a statistically significant result, followed by statistical analysis of the results to determine if a conclustion can be reached as to the best performing variation.
### Our Experiment Hypothesis
**Sample scenario:**
Website analytics have shown that user sessions frequently end on the home page for our e-commerce site, the Retail Demo Store. Furthermore, when users do make a purchase, most purchases are for a single product. Currently on our home page we are using a basic approach of recommending featured products. We hypothesize that adding personalized recommendations to the home page will result in increasing the click-through rate of products by 25%. The current click-through rate is 15%.
### ABExperiment Class
Before stepping through creating and executing our A/B test, let's look at the relevant source code for the [**ABExperiment**](https://github.com/aws-samples/retail-demo-store/blob/master/src/recommendations/src/recommendations-service/experimentation/experiment_ab.py) class that implements A/B experiments in the Retail Demo Store project.
As noted in the **3.1-Overview** notebook, all experiment types are subclasses of the abstract **Experiment** class. See **[3.1-Overview](./3.1-Overview.ipynb)** for more details on the experimentation framework.
The `ABExperiment.get_items()` method is where item recommendations are retrieved for the experiment. The `ABExperiment.calculate_variation_index()` method is where users are assigned to a variation/group using a consistent hashing algorithm. This ensures that each user is assigned to the same variation across multiple requests for recommended items for the duration of the experiment. Once the variation is determined, the variation's **Resolver** is used to retrieve recommendations. Details on the experiment are added to item list to support conversion/outcome tracking and UI annotation.
```python
# from src/recommendations/src/recommendations-service/experimentation/experiment_ab.py
class ABExperiment(Experiment):
...
def get_items(self, user_id, current_item_id = None, item_list = None, num_results = 10, tracker = None):
...
# Determine which variation to use for the user.
variation_idx = self.calculate_variation_index(user_id)
# Increment exposure counter for variation for this experiment.
self._increment_exposure_count(variation_idx)
# Get item recommendations from the variation's resolver.
variation = self.variations[variation_idx]
resolve_params = {
'user_id': user_id,
'product_id': current_item_id,
'num_results': num_results
}
items = variation.resolver.get_items(**resolve_params)
# Inject experiment details into recommended item list.
rank = 1
for item in items:
correlation_id = self._create_correlation_id(user_id, variation_idx, rank)
item_experiment = {
'id': self.id,
'feature': self.feature,
'name': self.name,
'type': self.type,
'variationIndex': variation_idx,
'resultRank': rank,
'correlationId': correlation_id
}
item.update({
'experiment': item_experiment
})
rank += 1
...
return items
def calculate_variation_index(self, user_id):
""" Given a user_id and this experiment's configuration, return the variation
The same variation will be returned for given user for this experiment no
matter how many times this method is called.
"""
if len(self.variations) == 0:
return -1
hash_str = f'experiments.{self.feature}.{self.name}.{user_id}'.encode('ascii')
hash_int = int(hashlib.sha1(hash_str).hexdigest()[:15], 16)
index = hash_int % len(self.variations)
return index
```
### Setup - Import Dependencies
Throughout this workshop we will need access to some common libraries and clients for connecting to AWS services. Let's set those up now.
```
import boto3
import json
import uuid
import numpy as np
import requests
import pandas as pd
import random
import scipy.stats as scs
import time
import decimal
import matplotlib.pyplot as plt
from boto3.dynamodb.conditions import Key
from random import randint
# import custom scripts used for plotting
from src.plot import *
from src.stats import *
%matplotlib inline
plt.style.use('ggplot')
# We will be using a DynamoDB table to store configuration info for our experiments.
dynamodb = boto3.resource('dynamodb')
# Service discovery will allow us to dynamically discover Retail Demo Store resources
servicediscovery = boto3.client('servicediscovery')
# Retail Demo Store config parameters are stored in SSM
ssm = boto3.client('ssm')
# Utility class to convert types for printing as JSON.
class CompatEncoder(json.JSONEncoder):
def default(self, obj):
if isinstance(obj, decimal.Decimal):
if obj % 1 > 0:
return float(obj)
else:
return int(obj)
else:
return super(CompatEncoder, self).default(obj)
```
### Sample Size Calculation
The first step is to determine the sample size necessary to reach a statistically significant result given a target of 25% gain in click-through rate from the home page. There are several sample size calculators available online including calculators from [Optimizely](https://www.optimizely.com/sample-size-calculator/?conversion=15&effect=20&significance=95), [AB Tasty](https://www.abtasty.com/sample-size-calculator/), and [Evan Miller](https://www.evanmiller.org/ab-testing/sample-size.html#!15;80;5;25;1). For this exercise, we will use the following function to calculate the minimal sample size for each variation.
```
def min_sample_size(bcr, mde, power=0.8, sig_level=0.05):
"""Returns the minimum sample size to set up a split test
Arguments:
bcr (float): probability of success for control, sometimes
referred to as baseline conversion rate
mde (float): minimum change in measurement between control
group and test group if alternative hypothesis is true, sometimes
referred to as minimum detectable effect
power (float): probability of rejecting the null hypothesis when the
null hypothesis is false, typically 0.8
sig_level (float): significance level often denoted as alpha,
typically 0.05
Returns:
min_N: minimum sample size (float)
References:
Stanford lecture on sample sizes
http://statweb.stanford.edu/~susan/courses/s141/hopower.pdf
"""
# standard normal distribution to determine z-values
standard_norm = scs.norm(0, 1)
# find Z_beta from desired power
Z_beta = standard_norm.ppf(power)
# find Z_alpha
Z_alpha = standard_norm.ppf(1-sig_level/2)
# average of probabilities from both groups
pooled_prob = (bcr + bcr+mde) / 2
min_N = (2 * pooled_prob * (1 - pooled_prob) * (Z_beta + Z_alpha)**2
/ mde**2)
return min_N
# This is the conversion rate using the current implementation
baseline_conversion_rate = 0.15
# This is the lift expected by adding personalization
absolute_percent_lift = baseline_conversion_rate * .25
# Calculate the sample size needed to reach a statistically significant result
sample_size = int(min_sample_size(baseline_conversion_rate, absolute_percent_lift))
print('Sample size for each variation: ' + str(sample_size))
```
### Experiment Strategy Datastore
With our sample size defined, let's create an experiment strategy for our A/B experiment. Walk through each of the following steps to configure your environment.
A DynamoDB table was created by the Retail Demo Store CloudFormation template that we will use to store the configuration information for our experiments. The table name can be found in a system parameter.
```
response = ssm.get_parameter(Name='retaildemostore-experiment-strategy-table-name')
table_name = response['Parameter']['Value'] # Do Not Change
print('Experiments DDB table: ' + table_name)
table = dynamodb.Table(table_name)
```
Next we need to lookup the Amazon Personalize campaign ARN for product recommendations. This is the campaign that was created in the [Personalization workshop](../1-Personalization/personalize.ipynb) (or was pre-built for you depending on your workshop event).
```
response = ssm.get_parameter(Name = 'retaildemostore-product-recommendation-campaign-arn')
campaign_arn = response['Parameter']['Value'] # Do Not Change
print('Personalize product recommendations ARN: ' + campaign_arn)
```
### Create A/B Experiment
The Retail Demo Store supports running multiple experiments concurrently. For this workshop we will create a single A/B test/experiment that uniformly splits users between a control group that receives recommendations from the default behavior and a variance group that receives recommendations from Amazon Personalize. The Recommendations service already has logic that supports A/B tests once an active experiment is detected our Experiment Strategy DynamoDB table.
Experiment configurations are stored in a DynamoDB table where each item in the table represents an experiment and has the following fields.
- **id** - Uniquely identified this experience (UUID).
- **feature** - Identifies the Retail Demo Store feature where the experiment should be applied. The name for the home page product recommendations feature is `home_product_recs`.
- **name** - The name of the experiment. Keep the name short but descriptive. It will be used in the UI for demo purposes and when logging events for experiment result tracking.
- **status** - The status of the experiment (`ACTIVE`, `EXPIRED`, or `PENDING`).
- **type** - The type of test (`ab` for an A/B test, `interleaving` for interleaved recommendations, or `mab` for multi-armed bandit test)
- **variations** - List of configurations representing variations for the experiment. For example, for A/B tests of the `home_product_recs` feature, the `variations` can be two Amazon Personalize campaign ARNs (variation type `personalize-recommendations`) or a single Personalize campaign ARN and the default product behavior.
```
feature = 'home_product_recs'
experiment_name = 'home_personalize_ab'
# First, make sure there are no other active experiments so we can isolate
# this experiment for the exercise (to keep things clean/simple).
response = table.scan(
ProjectionExpression='#k',
ExpressionAttributeNames={'#k' : 'id'},
FilterExpression=Key('status').eq('ACTIVE')
)
for item in response['Items']:
response = table.update_item(
Key=item,
UpdateExpression='SET #s = :inactive',
ExpressionAttributeNames={
'#s' : 'status'
},
ExpressionAttributeValues={
':inactive' : 'INACTIVE'
}
)
# Query the experiment strategy table to see if our experiment already exists
response = table.query(
IndexName='feature-name-index',
KeyConditionExpression=Key('feature').eq(feature) & Key('name').eq(experiment_name),
FilterExpression=Key('status').eq('ACTIVE')
)
if response.get('Items') and len(response.get('Items')) > 0:
print('Experiment already exists')
home_page_experiment = response['Items'][0]
else:
print('Creating experiment')
# Default product resolver
variation_0 = {
'type': 'product'
}
# Amazon Personalize resolver
variation_1 = {
'type': 'personalize-recommendations',
'campaign_arn': campaign_arn
}
home_page_experiment = {
'id': uuid.uuid4().hex,
'feature': feature,
'name': experiment_name,
'status': 'ACTIVE',
'type': 'ab',
'variations': [ variation_0, variation_1 ]
}
response = table.put_item(
Item=home_page_experiment
)
print(json.dumps(response, indent=4))
print('Experiment item:')
print(json.dumps(home_page_experiment, indent=4, cls=CompatEncoder))
```
## Load Users
For our experiment simulation, we will load all Retail Demo Store users and run the experiment until the sample size for both variations has been met.
First, let's discover the IP address for the Retail Demo Store's [Users](https://github.com/aws-samples/retail-demo-store/tree/master/src/users) service.
```
response = servicediscovery.discover_instances(
NamespaceName='retaildemostore.local',
ServiceName='users',
MaxResults=1,
HealthStatus='HEALTHY'
)
users_service_instance = response['Instances'][0]['Attributes']['AWS_INSTANCE_IPV4']
print('Users Service Instance IP: {}'.format(users_service_instance))
```
Next, let's fetch all users, randomize their order, and load them into a local data frame.
```
# Load all users so we have enough to satisfy our sample size requirements.
response = requests.get('http://{}/users/all?count=10000'.format(users_service_instance))
users = response.json()
random.shuffle(users)
users_df = pd.DataFrame(users)
pd.set_option('display.max_rows', 5)
users_df
```
## Discover Recommendations Service
Next, let's discover the IP address for the Retail Demo Store's [Recommendations](https://github.com/aws-samples/retail-demo-store/tree/master/src/recommendations) service. This is the service where the Experimentation framework is implemented and the `/recommendations` endpoint is what we call to simulate our A/B experiment.
```
response = servicediscovery.discover_instances(
NamespaceName='retaildemostore.local',
ServiceName='recommendations',
MaxResults=1,
HealthStatus='HEALTHY'
)
recommendations_service_instance = response['Instances'][0]['Attributes']['AWS_INSTANCE_IPV4']
print('Recommendation Service Instance IP: {}'.format(recommendations_service_instance))
```
## Simulate Experiment
Next we will define a function to simulate our A/B experiment by making calls to the [Recommendations](https://github.com/aws-samples/retail-demo-store/tree/master/src/recommendations) service across the users we just loaded. Then we will run our simulation.
### Simulation Function
The following `simulate_experiment` function is supplied with the sample size for each group (A and B) and the probability of conversion for each group that we want to use for our simulation. It runs the simulation long enough to satisfy the sample size requirements and calls the Recommendations service for each user in the experiment.
```
def simulate_experiment(N_A, N_B, p_A, p_B):
"""Returns a pandas dataframe with simulated CTR data
Parameters:
N_A (int): sample size for control group
N_B (int): sample size for test group
Note: final sample size may not match N_A & N_B provided because the
group at each row is chosen at random by the ABExperiment class.
p_A (float): conversion rate; conversion rate of control group
p_B (float): conversion rate; conversion rate of test group
Returns:
df (df)
"""
# will hold exposure/outcome data
data = []
# total number of users to sample for both variations
N = N_A + N_B
if N > len(users):
raise ValueError('Sample size is greater than number of users')
print('Generating data for {} users... this may take a few minutes'.format(N))
# initiate bernoulli distributions to randomly sample from based on simulated probabilities
A_bern = scs.bernoulli(p_A)
B_bern = scs.bernoulli(p_B)
for idx in range(N):
if idx > 0 and idx % 500 == 0:
print('Generated data for {} users so far'.format(idx))
# initite empty row
row = {}
# Get next user from shuffled list
user = users[idx]
# Call Recommendations web service to get recommendations for the user
response = requests.get('http://{}/recommendations?userID={}&feature={}'.format(recommendations_service_instance, user['id'], feature))
recommendations = response.json()
recommendation = recommendations[randint(0, len(recommendations)-1)]
variation = recommendation['experiment']['variationIndex']
row['variation'] = variation
# Determine if variation converts based on probabilities provided
if variation == 0:
row['converted'] = A_bern.rvs()
else:
row['converted'] = B_bern.rvs()
if row['converted'] == 1:
# Update experiment with outcome/conversion
correlation_id = recommendation['experiment']['correlationId']
requests.post('http://{}/experiment/outcome'.format(recommendations_service_instance), data={'correlationId':correlation_id})
data.append(row)
# convert data into dataframe
df = pd.DataFrame(data)
print('Done')
return df
```
### Run Simulation
Next we run the simulation by defining our simulation parameters for sample sizes and probabilities and then call `simulate_experiment`. This will take several minutes depending on the sample sizes.
```
%%time
# Set size of both groups to calculated sample size
N_A = N_B = sample_size
# Use probabilities from our hypothesis
# bcr: baseline conversion rate
p_A = 0.15
# d_hat: difference in a metric between the two groups, sometimes referred to as minimal detectable effect or lift depending on the context
p_B = 0.1875
# Run simulation
ab_data = simulate_experiment(N_A, N_B, p_A, p_B)
ab_data
```
### Inspect Experiment Summary Statistics
Since the **Experiment** class updates statistics for the experiment in the experiment strategy DynamoDB table when a user is exposed to an experiment ("exposure") and when a user converts ("outcome"), we should see updated counts on our experiment. Let's reload our experiment and inspect the exposure and conversion counts for our simulation.
```
# Query DDB table for experiment item.
response = table.get_item(Key={'id': home_page_experiment['id']})
print(json.dumps(response['Item'], indent=4, cls=CompatEncoder))
```
You should now see counters for `conversions` and `exposures` for each variation. These represent how many times a user has been exposed to a variation and how many times a user has converted for a variation (i.e. clicked on a recommended item/product).
### Analyze Simulation Results
Next, let's take a closer look at the results of our simulation. We'll start by calculating some summary statistics.
```
ab_summary = ab_data.pivot_table(values='converted', index='variation', aggfunc=np.sum)
# add additional columns to the pivot table
ab_summary['total'] = ab_data.pivot_table(values='converted', index='variation', aggfunc=lambda x: len(x))
ab_summary['rate'] = ab_data.pivot_table(values='converted', index='variation')
ab_summary
```
The output above tells us how many users converted for each variation, the actual sample size for each variation in the simulation, and the conversion rate for each variation.
Next let's isolate the data and conversion counts for each variation.
```
A_group = ab_data[ab_data['variation'] == 0]
B_group = ab_data[ab_data['variation'] == 1]
A_converted, B_converted = A_group['converted'].sum(), B_group['converted'].sum()
A_converted, B_converted
```
Isolate the actual sample size for each variation.
```
A_total, B_total = len(A_group), len(B_group)
A_total, B_total
```
Calculate the actual conversion rates and uplift for our simulation.
```
p_A, p_B = A_converted / A_total, B_converted / B_total
p_A, p_B
p_B - p_A
```
### Determining Statistical Significance
In statistical hypothesis testing there are two types of errors that can occur. These are referred to as type 1 and type 2 errors.
Type 1 errors occur when the null hypothesis is true but is rejected. In other words, a "false positive" conclusion. Put in A/B testing terms, a type 1 error is when we conclude a statistically significant result when there isn't one.
Type 2 errors occur when we conclude that there is not a winner between two variations when in fact there is an actual winner. In other words, the null hypothesis is false yet we fail to reject it. Therefore, type 2 errors are a "false negative" conclusion.
If the probability of making a type 1 error is determined by "α" (alpha), the probability of a type 2 error is "β" (beta). Beta depends on the power of the test (i.e the probability of not committing a type 2 error, which is equal to 1-β).
Let's inspect the results of our simulation more closely to verify that it is statistically significant.
#### Calculate p-value
Formally, the p-value is the probability of seeing a particular result (or greater) from zero, assuming that the null hypothesis is TRUE. In other words, the p-value is the expected fluctuation in a given sample, similar to the variance. As an example, imagine we ran an A/A test where displayed the same variation to two groups of users. After such an experiment we would expect the conversion rates of both groups to be very similar but not dramatically different.
What we are hoping to see is a p-value that is less than our significance level. The significance level we used when calculating our sample size was 5%, which means we are seeking results with 95% accuracy. 5% is considered the industry standard.
```
p_value = scs.binom(A_total, p_A).pmf(p_B * B_total)
print('p-value = {0:0.9f}'.format(p_value))
```
Is the p-value less than the signficance level of 5%? This tells us the probability of a type 1 error.
Let's plot the data from both groups as binomial distributions.
```
fig, ax = plt.subplots(figsize=(12,6))
xA = np.linspace(A_converted-49, A_converted+50, 100)
yA = scs.binom(A_total, p_A).pmf(xA)
ax.scatter(xA, yA, s=10)
xB = np.linspace(B_converted-49, B_converted+50, 100)
yB = scs.binom(B_total, p_B).pmf(xB)
ax.scatter(xB, yB, s=10)
plt.xlabel('converted')
plt.ylabel('probability')
```
Based the probabilities from our hypothesis, we should see that the test group in blue (B) converted more users than the control group in red (A). However, the plot above is not a plot of the null and alternate hypothesis. The null hypothesis is a plot of the difference between the probability of the two groups.
> Given the randomness of our user selection, group hashing, and probabilities, your simulation results should be different for each simulation run and therefore may or may not be statistically significant.
In order to calculate the difference between the two groups, we need to standardize the data. Because the number of samples can be different between the two groups, we should compare the probability of successes, p.
According to the central limit theorem, by calculating many sample means we can approximate the true mean of the population from which the data for the control group was taken. The distribution of the sample means will be normally distributed around the true mean with a standard deviation equal to the standard error of the mean.
```
SE_A = np.sqrt(p_A * (1-p_A)) / np.sqrt(A_total)
SE_B = np.sqrt(p_B * (1-p_B)) / np.sqrt(B_total)
SE_A, SE_B
fig, ax = plt.subplots(figsize=(12,6))
xA = np.linspace(0, .3, A_total)
yA = scs.norm(p_A, SE_A).pdf(xA)
ax.plot(xA, yA)
ax.axvline(x=p_A, c='red', alpha=0.5, linestyle='--')
xB = np.linspace(0, .3, B_total)
yB = scs.norm(p_B, SE_B).pdf(xB)
ax.plot(xB, yB)
ax.axvline(x=p_B, c='blue', alpha=0.5, linestyle='--')
plt.xlabel('Converted Proportion')
plt.ylabel('PDF')
```
The dashed lines represent the mean conversion rate for each group. The distance between the red dashed line and the blue dashed line is equal to d_hat, or the minimum detectable effect.
```
p_A_actual = ab_summary.loc[0, 'rate']
p_B_actual = ab_summary.loc[1, 'rate']
bcr = p_A_actual
d_hat = p_B_actual - p_A_actual
A_total, B_total, bcr, d_hat
```
Finally, let's calculate the power, alpha, and beta from our simulation.
```
abplot(A_total, B_total, bcr, d_hat, show_power=True)
```
The power value we used when determining out sample size for our experiment was 80%. This is considered the industry standard. Is the power value calculated in the plot above greater than 80%?
```
abplot(A_total, B_total, bcr, d_hat, show_beta=True)
abplot(A_total, B_total, bcr, d_hat, show_alpha=True)
```
Are the alpha and beta values plotted in the graphs above less than our significance level of 5%? If so, we have a statistically significant result.
## Next Steps
You have completed the exercise for implementing an A/B test using the experimentation framework in the Retail Demo Store. Close this notebook and open the notebook for the next exercise, **[3.3-Interleaving-Experiment](./3.3-Interleaving-Experiment.ipynb)**.
### References and Further Reading
- [A/B testing](https://en.wikipedia.org/wiki/A/B_testing), Wikipedia
- [A/B testing](https://www.optimizely.com/optimization-glossary/ab-testing/), Optimizely
- [Evan's Awesome A/B Tools](https://www.evanmiller.org/ab-testing/), Evan Miller
| github_jupyter |
<a href="https://colab.research.google.com/github/mposa/MEAM517/blob/master/Lecture14/van_der_pol_sos_examples.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Install Drake
# New Section
## Constant V, line search over multipliers
```
# Install drake. We are using the lqr controller in drake as the final stabilizing controller.
# The installation process will take about 2 minutes but it's only required in the start of the Colab's virtual machine.
!curl -s https://raw.githubusercontent.com/mposa/MEAM517/master/colab_drake_setup.py > colab_drake_setup.py
from colab_drake_setup import setup
setup()
import math
import numpy as np
from pydrake.solvers.mathematicalprogram import MathematicalProgram, Solve
import pydrake.symbolic as sym
from pydrake.symbolic import Polynomial
from scipy.linalg import solve_lyapunov
from pydrake.solvers.mathematicalprogram import SolutionResult
import matplotlib.pyplot as plt
from scipy.integrate import solve_ivp
# Initial guess
rho = 1
delta = .1
double_next = False
rho_opt = 0
# Stopping criteria
min_delta = .001
while delta > min_delta:
# Initialize an empty optimization program.
prog = MathematicalProgram()
# Declare state "x" as indeterminates
x = prog.NewIndeterminates(2)
# Dynamics
a = 1
xdot = np.array([-a*(x[0] - x[0]**3 / 3 - x[1]), -x[0]/a])
# Solve Lyapunov equation for V
A = np.array([[-a, a], [-1/a, 0]])
Q = np.identity(2);
#Q = np.array([[10, 0], [0, 1]])
S=solve_lyapunov(A.T, -Q)
V = x.T @ S @ x;
Vdot = V.Jacobian(x) @ xdot
sig_poly = prog.NewSosPolynomial(sym.Variables(x), 4)
sig = sig_poly[0].ToExpression()
p1 = (V - rho) * sig - Vdot
#p1 = (V - rho) - Vdot
prog.AddSosConstraint(p1);
prog.AddCost(-rho);
# Solve program
result = Solve(prog)
if result.get_solution_result() == SolutionResult.kSolutionFound:
rho_opt = rho
if double_next:
delta = delta * 2
double_next = True
rho = rho + delta
else:
delta = delta/2
rho = rho - delta
double_next = False
print(rho)
print(result.get_solution_result())
print("Done!")
print(rho_opt)
# Plotting
def dynamics(t, x) : return [a*(x[0] - x[0]**3 / 3 - x[1]), x[0]/a]
ode_sol = solve_ivp(dynamics, [0, 20], [0, 3], t_eval=np.linspace(0, 20, 1000))
Vsol = result.GetSolution(V)
N_sample = 40
x1_sample, x2_sample = np.meshgrid(np.linspace(-3,3,N_sample), np.linspace(-3,3,N_sample))
Vplot = np.zeros([N_sample, N_sample])
x1dot = np.zeros([N_sample, N_sample])
x2dot = np.zeros([N_sample, N_sample])
for i in range(x1_sample.shape[0]):
for j in range(x1_sample.shape[1]):
env = {x[0] : x1_sample[i][j], x[1]: x2_sample[i][j]}
Vplot[i][j] = Vsol.Evaluate(env)
x1dot[i][j] = xdot[0].Evaluate(env)
x2dot[i][j] = xdot[1].Evaluate(env)
plt.rcParams['figure.figsize'] = [12,8];
plt.quiver(x1_sample, x2_sample, x1dot, x2dot);
plt.contour(x1_sample, x2_sample, Vplot, [rho]);
plt.plot(ode_sol.y[0,500:], ode_sol.y[1,500:], linewidth=3);
```
## Constant multipliers, search V
```
import math
import numpy as np
from pydrake.solvers.mathematicalprogram import MathematicalProgram, Solve, SolverOptions, SolverType, ChooseBestSolver
import pydrake.symbolic as sym
from pydrake.symbolic import Polynomial
import matplotlib.pyplot as plt
from scipy.integrate import solve_ivp
# Initialize an empty optimization program.
prog = MathematicalProgram()
# Declare state "x" as indeterminates
x = prog.NewIndeterminates(2)
# parameters
degree = 6
sigma_degree = 8
# Dynamics
a = 1
xdot = np.array([-a*(x[0] - x[0]**3 / 3 - x[1]), -x[0]/a])
rho = 1
V_poly = prog.NewSosPolynomial(sym.Variables(x), degree)
V = V_poly[0].ToExpression()
Vdot = V.Jacobian(x) @ xdot
# Constrain V(0,0) = 0
env_0 = {x[0] : 0, x[1]: 0}
prog.AddLinearConstraint(V.EvaluatePartial(env_0) == 0)
sigma_1 = (x.T @ x) ** (sigma_degree/2)
sigma_2 = (1 + x.T @ x) ** (sigma_degree/2 - 1)
p1 = sigma_1 * (V - rho) - sigma_2 * Vdot
prog.AddSosConstraint(p1);
# Create a cost by sampling over points
N_sample = 5
x1_sample, x2_sample = np.meshgrid(np.linspace(-3,3,N_sample), np.linspace(-3,3,N_sample))
for i in range(x1_sample.shape[0]):
for j in range(x1_sample.shape[1]):
env = {x[0] : x1_sample[i][j], x[1]: x2_sample[i][j]}
prog.AddCost(V.EvaluatePartial(env))
# Solve program
result = Solve(prog)
print(result.get_solution_result())
# Print resulting V
print(result.GetSolution(V))
# Plotting
def dynamics(t, x) : return [a*(x[0] - x[0]**3 / 3 - x[1]), x[0]/a]
ode_sol = solve_ivp(dynamics, [0, 20], [0, 3], t_eval=np.linspace(0, 20, 1000))
Vsol = result.GetSolution(V)
N_sample = 40
x1_sample, x2_sample = np.meshgrid(np.linspace(-3,3,N_sample), np.linspace(-3,3,N_sample))
Vplot = np.zeros([N_sample, N_sample])
x1dot = np.zeros([N_sample, N_sample])
x2dot = np.zeros([N_sample, N_sample])
for i in range(x1_sample.shape[0]):
for j in range(x1_sample.shape[1]):
env = {x[0] : x1_sample[i][j], x[1]: x2_sample[i][j]}
Vplot[i][j] = Vsol.Evaluate(env)
x1dot[i][j] = xdot[0].Evaluate(env)
x2dot[i][j] = xdot[1].Evaluate(env)
plt.rcParams['figure.figsize'] = [12,8];
plt.quiver(x1_sample, x2_sample, x1dot, x2dot);
plt.contour(x1_sample, x2_sample, Vplot, [1]);
plt.plot(ode_sol.y[0,500:], ode_sol.y[1,500:], linewidth=3);
```
| github_jupyter |
# Template
- Author: Israel Oliveira [\[e-mail\]](mailto:'Israel%20Oliveira%20'<prof.israel@gmail.com>)
```
%load_ext watermark
import numpy as np
import pandas as pd
from surprise import SVD
from surprise import Dataset
from surprise import Reader
from surprise.model_selection import cross_validate, GridSearchCV
# Run this cell before close.
%watermark
%watermark --iversion
%watermark -b -r -g -p surprise
pd.set_option('display.max_rows', 500)
pd.set_option('display.max_columns', 30)
pd.set_option('display.width', 1000)
!ls ../data/
path_data = '../data/'
```
#### Carrega dataset só com as top colunas escolhidas na priveira análise exploratória.
```
top_cols = pd.read_csv('top_cols.csv')['cols'].to_list()
df_marked = pd.read_csv(path_data+'estaticos_market.csv', usecols=top_cols)
col_user = 'id'
top_cols.remove(col_user)
df_marked[top_cols].head()
```
No caso de uso da biblioteca Surprise, é necessário normalizar os valores numéricos.
Quanto aos valores booleanos, teremos rating binário min/max.
```
df_marked.dtypes[13] == float
df_marked.dtypes.unique()
rest_cols = []
for col in top_cols:
if df_marked[col].dtype in [float, int, bool]:
df_marked[col] = df_marked[col].fillna(0)*1
else:
rest_cols.append(col)
print("{}: {}".format(col,df_marked[col].unique()))
for col in rest_cols:
df_marked[col] = df_marked[col].fillna(0)*1
df_marked[top_cols].head()
```
É necessário normalizar (deixar entre $[0, 1]$) e escalar para um valor que matenha algum nível de detalhe. Usarei uma escala de inteiros entre $[0, 100]$, deve ser o suficiente.
```
def normalize(x):
return (x-np.min(x))/(np.max(x) - np.min(x)) if (np.max(x) - np.min(x)) > 0 else (x-np.min(x))
escala = 100
for col in top_cols:
df_marked[col] = (escala*normalize(df_marked[col].tolist())).astype(np.uint8)
df_marked[top_cols].head()
remove_cols = []
for col in top_cols:
if df_marked[col].nunique() == 1:
remove_cols.append(col)
df_marked = df_marked.drop(columns=remove_cols)
for col in remove_cols:
top_cols.remove(col)
```
Agora temos todos os valores normalizados entre $[0, 100]$, podendo gerar o datset para o surprase.
- `'id'`: será o id de usuário (`user_id`), cada valor da coluna será considerado um usuário.
- `top_cols`: serão o ids de itens (`user_id`), cada coluna será considerado um item.
- valores nas `top_cols`: serão os ratings (`rating`).-
```
df_marked = pd.melt(df_marked, id_vars=["id"], var_name="itemID", value_name="rating").rename(columns={"id": "userID"})
df_marked.head()
sum(df_marked.userID == df_marked.userID.loc[0])
```
Gerando dataset no formato para o Surprise.
```
reader = Reader(rating_scale=(0, escala))
data = Dataset.load_from_df(df_marked[['userID', 'itemID', 'rating']], reader)
```
Primeira tentativa com algoritmo.
PC:
```
algo = SVD()
cross_validate(algo, data, measures=['RMSE', 'MAE'], cv=5, verbose=True)
len(top_cols)
top_cols
```
Para o Experimento B: variar a quantidade de colunas.
MAC:
```
algo = SVD()
cross_validate(algo, data, measures=['RMSE', 'MAE'], cv=5, verbose=True)
```
| github_jupyter |
## 1. Heart disease and potential risk factors
<p>Millions of people develop some sort of heart disease every year and heart disease is the biggest killer of both men and women in the United States and around the world. Statistical analysis has identified many risk factors associated with heart disease such as age, blood pressure, total cholesterol, diabetes, hypertension, family history of heart disease, obesity, lack of physical exercise, etc. In this notebook, we're going to run statistical tests and regression models using the Cleveland heart disease dataset to assess one particular factor -- maximum heart rate one can achieve during exercise and how it is associated with a higher likelihood of getting heart disease.</p>
<p><img src="https://assets.datacamp.com/production/project_445/img/run31.png" height="300" width="300"></p>
```
# Read datasets Cleveland_hd.csv into hd_data
hd_data <- read.csv("datasets/Cleveland_hd.csv")
# take a look at the first 5 rows of hd_data
head(hd_data,5)
```
## 2. Converting diagnosis class into outcome variable
<p>We noticed that the outcome variable <code>class</code> has more than two levels. According to the codebook, any non-zero values can be coded as an "event." Let's create a new variable called <code>hd</code> to represent a binary 1/0 outcome.</p>
<p>There are a few other categorical/discrete variables in the dataset. Let's also convert sex into a 'factor' for next step analysis. Otherwise, R will treat this as continuous by default.</p>
<p>The full data dictionary is also displayed here.</p>
<p><img src="https://assets.datacamp.com/production/project_445/img/datadict.png" height="500" width="500"></p>
```
# load the tidyverse package
library(tidyverse)
# Use the 'mutate' function from dplyr to recode our data
hd_data %>% mutate(hd = ifelse(class > 0, 1, 0))-> hd_data
# recode sex using mutate function and save as hd_data
hd_data %>% mutate(sex = factor(sex, levels = 0:1, labels = c('Female','Male')))-> hd_data;
```
## 3. Identifying important clinical variables
<p>Now, let's use statistical tests to see which predictors are related to heart disease. We can explore the associations for each variable in the dataset. Depending on the type of the data (i.e., continuous or categorical), we use t-test or chi-squared test to calculate the p-values.</p>
<p>Recall, t-test is used to determine whether there is a significant difference between the means of two groups (e.g., is the mean age from group A different from the mean age from group B?). A chi-squared test for independence compares the equivalence of two proportions.</p>
```
# Does sex have an effect? Sex is a binary variable in this dataset,
# so the appropriate test is chi-squared test
hd_sex <- chisq.test(hd_data$sex,hd_data$hd)
# Does age have an effect? Age is continuous, so we use a t-test
hd_age <- t.test(hd_data$age~hd_data$hd)
# What about thalach? Thalach is continuous, so we use a t-test
hd_heartrate <- t.test(hd_data$thalach~hd_data$hd)
# Print the results to see if p<0.05.
print(hd_sex)
print(hd_age)
print(hd_heartrate)
```
## 4. Explore the associations graphically (i)
<p>A good picture is worth a thousand words. In addition to p-values from statistical tests, we can plot the age, sex, and maximum heart rate distributions with respect to our outcome variable. This will give us a sense of both the direction and magnitude of the relationship.</p>
<p>First, let's plot age using a boxplot since it is a continuous variable.</p>
```
# Recode hd to be labelled
hd_data%>%mutate(hd_labelled = ifelse(hd == 0, 'No Disease', 'Disease')) -> hd_data
# age vs hd
ggplot(data = hd_data, aes(x = hd_labelled,y = age)) + geom_boxplot()
```
## 5. Explore the associations graphically (ii)
<p>Next, let's plot sex using a barplot since it is a binary variable in this dataset.</p>
```
# sex vs hd
ggplot(data = hd_data,aes(x=hd_labelled,fill=sex)) + geom_bar(position='fill')+ ylab("Sex %")
```
## 6. Explore the associations graphically (iii)
<p>And finally, let's plot thalach using a boxplot since it is a continuous variable.</p>
```
# max heart rate vs hd
ggplot(data = hd_data,aes(x=hd_labelled,y=thalach)) + geom_boxplot()
```
## 7. Putting all three variables in one model
<p>The plots and the statistical tests both confirmed that all the three variables are highly significantly associated with our outcome (p<0.001 for all tests). </p>
<p>In general, we want to use multiple logistic regression when we have one binary outcome variable and two or more predicting variables. The binary variable is the dependent (Y) variable; we are studying the effect that the independent (X) variables have on the probability of obtaining a particular value of the dependent variable. For example, we might want to know the effect that maximum heart rate, age, and sex have on the probability that a person will have a heart disease in the next year. The model will also tell us what the remaining effect of maximum heart rate is after we control or adjust for the effects of the other two effectors. </p>
<p>The <code>glm()</code> command is designed to perform generalized linear models (regressions) on binary outcome data, count data, probability data, proportion data, and many other data types. In our case, the outcome is binary following a binomial distribution.</p>
```
# use glm function from base R and specify the family argument as binomial
model <- glm(data = hd_data, hd~age+sex+thalach, family='binomial')
# extract the model summary
summary(model)
```
## 8. Extracting useful information from the model output
<p>It's common practice in medical research to report Odds Ratio (OR) to quantify how strongly the presence or absence of property A is associated with the presence or absence of the outcome. When the OR is greater than 1, we say A is positively associated with outcome B (increases the Odds of having B). Otherwise, we say A is negatively associated with B (decreases the Odds of having B).</p>
<p>The raw glm coefficient table (the 'estimate' column in the printed output) in R represents the log(Odds Ratios) of the outcome. Therefore, we need to convert the values to the original OR scale and calculate the corresponding 95% Confidence Interval (CI) of the estimated Odds Ratios when reporting results from a logistic regression. </p>
```
# load the broom package
library('broom')
# tidy up the coefficient table
tidy_m <- tidy(model)
tidy_m
# calculate OR
tidy_m$OR <- exp(tidy$estimate)
# calculate 95% CI and save as lower CI and upper CI
tidy_m$lower_CI <- exp(tidy_m$estimate - 1.96 * tidy_m$std.error)
tidy_m$upper_CI <- exp(tidy_m$estimate + 1.96 * tidy_m$std.error)
# display the updated coefficient table
tidy_m
```
## 9. Predicted probabilities from our model
<p>So far, we have built a logistic regression model and examined the model coefficients/ORs. We may wonder how can we use this model we developed to predict a person's likelihood of having heart disease given his/her age, sex, and maximum heart rate. Furthermore, we'd like to translate the predicted probability into a decision rule for clinical use by defining a cutoff value on the probability scale. In practice, when an individual comes in for a health check-up, the doctor would like to know the predicted probability of heart disease, for specific values of the predictors: a 45-year-old female with a max heart rate of 150. To do that, we create a data frame called newdata, in which we include the desired values for our prediction.</p>
```
# get the predicted probability in our dataset using the predict() function
pred_prob <- predict(model,hd_data, type = "response")
# create a decision rule using probability 0.5 as cutoff and save the predicted decision into the main data frame
hd_data$pred_hd <- ifelse(pred_prob>=0.5,1,0)
# create a newdata data frame to save a new case information
newdata <- data.frame(age = 45, sex = "Female", thalach = 150)
# predict probability for this new case and print out the predicted value
p_new <- predict(model,newdata, type = "response")
p_new
```
## 10. Model performance metrics
<p>Are the predictions accurate? How well does the model fit our data? We are going to use some common metrics to evaluate the model performance. The most straightforward one is Accuracy, which is the proportion of the total number of predictions that were correct. On the other hand, we can calculate the classification error rate using 1- accuracy. However, accuracy can be misleading when the response is rare (i.e., imbalanced response). Another popular metric, Area Under the ROC curve (AUC), has the advantage that it's independent of the change in the proportion of responders. AUC ranges from 0 to 1. The closer it gets to 1 the better the model performance. Lastly, a confusion matrix is an N X N matrix, where N is the level of outcome. For the problem at hand, we have N=2, and hence we get a 2 X 2 matrix. It cross-tabulates the predicted outcome levels against the true outcome levels.</p>
<p>After these metrics are calculated, we'll see (from the logistic regression OR table) that older age, being male and having a lower max heart rate are all risk factors for heart disease. We can also apply our model to predict the probability of having heart disease. For a 45 years old female who has a max heart rate of 150, our model generated a heart disease probability of 0.177 indicating low risk of heart disease. Although our model has an overall accuracy of 0.71, there are cases that were misclassified as shown in the confusion matrix. One way to improve our current model is to include other relevant predictors from the dataset into our model, but that's a task for another day!</p>
```
# load Metrics package
library(Metrics)
# calculate auc, accuracy, clasification error
auc <- auc(hd_data$hd,hd_data$pred_hd)
accuracy <- accuracy(hd_data$hd,hd_data$pred_hd)
classification_error <- ce(hd_data$hd,hd_data$pred_hd)
# print out the metrics on to screen
print(paste("AUC=", auc))
print(paste("Accuracy=", accuracy))
print(paste("Classification Error=", classification_error))
# confusion matrix
table(hd_data$hd,hd_data$pred_hd, dnn=c('True Status','Predicted Status')) # confusion matrix
```
| github_jupyter |
**This notebook is an exercise in the [Intermediate Machine Learning](https://www.kaggle.com/learn/intermediate-machine-learning) course. You can reference the tutorial at [this link](https://www.kaggle.com/alexisbcook/pipelines).**
---
In this exercise, you will use **pipelines** to improve the efficiency of your machine learning code.
# Setup
The questions below will give you feedback on your work. Run the following cell to set up the feedback system.
```
# Set up code checking
import os
if not os.path.exists("../input/train.csv"):
os.symlink("../input/home-data-for-ml-course/train.csv", "../input/train.csv")
os.symlink("../input/home-data-for-ml-course/test.csv", "../input/test.csv")
from learntools.core import binder
binder.bind(globals())
from learntools.ml_intermediate.ex4 import *
print("Setup Complete")
```
You will work with data from the [Housing Prices Competition for Kaggle Learn Users](https://www.kaggle.com/c/home-data-for-ml-course).

Run the next code cell without changes to load the training and validation sets in `X_train`, `X_valid`, `y_train`, and `y_valid`. The test set is loaded in `X_test`.
```
import pandas as pd
from sklearn.model_selection import train_test_split
# Read the data
X_full = pd.read_csv('../input/train.csv', index_col='Id')
X_test_full = pd.read_csv('../input/test.csv', index_col='Id')
# Remove rows with missing target, separate target from predictors
X_full.dropna(axis=0, subset=['SalePrice'], inplace=True)
y = X_full.SalePrice
X_full.drop(['SalePrice'], axis=1, inplace=True)
# Break off validation set from training data
X_train_full, X_valid_full, y_train, y_valid = train_test_split(X_full, y,
train_size=0.8, test_size=0.2,
random_state=0)
# "Cardinality" means the number of unique values in a column
# Select categorical columns with relatively low cardinality (convenient but arbitrary)
categorical_cols = [cname for cname in X_train_full.columns if
X_train_full[cname].nunique() < 10 and
X_train_full[cname].dtype == "object"]
# Select numerical columns
numerical_cols = [cname for cname in X_train_full.columns if
X_train_full[cname].dtype in ['int64', 'float64']]
# Keep selected columns only
my_cols = categorical_cols + numerical_cols
X_train = X_train_full[my_cols].copy()
X_valid = X_valid_full[my_cols].copy()
X_test = X_test_full[my_cols].copy()
X_train.head()
```
The next code cell uses code from the tutorial to preprocess the data and train a model. Run this code without changes.
```
from sklearn.compose import ColumnTransformer
from sklearn.pipeline import Pipeline
from sklearn.impute import SimpleImputer
from sklearn.preprocessing import OneHotEncoder
from sklearn.ensemble import RandomForestRegressor
from sklearn.metrics import mean_absolute_error
# Preprocessing for numerical data
numerical_transformer = SimpleImputer(strategy='constant')
# Preprocessing for categorical data
categorical_transformer = Pipeline(steps=[
('imputer', SimpleImputer(strategy='most_frequent')),
('onehot', OneHotEncoder(handle_unknown='ignore'))
])
# Bundle preprocessing for numerical and categorical data
preprocessor = ColumnTransformer(
transformers=[
('num', numerical_transformer, numerical_cols),
('cat', categorical_transformer, categorical_cols)
])
# Define model
model = RandomForestRegressor(n_estimators=100, random_state=0)
# Bundle preprocessing and modeling code in a pipeline
clf = Pipeline(steps=[('preprocessor', preprocessor),
('model', model)
])
# Preprocessing of training data, fit model
clf.fit(X_train, y_train)
# Preprocessing of validation data, get predictions
preds = clf.predict(X_valid)
print('MAE:', mean_absolute_error(y_valid, preds))
```
The code yields a value around 17862 for the mean absolute error (MAE). In the next step, you will amend the code to do better.
# Step 1: Improve the performance
### Part A
Now, it's your turn! In the code cell below, define your own preprocessing steps and random forest model. Fill in values for the following variables:
- `numerical_transformer`
- `categorical_transformer`
- `model`
To pass this part of the exercise, you need only define valid preprocessing steps and a random forest model.
```
# Preprocessing for numerical data
numerical_transformer = SimpleImputer()
# Preprocessing for categorical data
categorical_transformer = Pipeline(steps=[
('imputer', SimpleImputer(strategy='most_frequent')),
('onehot', OneHotEncoder(handle_unknown='ignore'))
])
# Bundle preprocessing for numerical and categorical data
preprocessor = ColumnTransformer(
transformers=[
('num', numerical_transformer, numerical_cols),
('cat', categorical_transformer, categorical_cols)
])
# Define model
model = RandomForestRegressor(n_estimators=100, random_state=0)
# Check your answer
step_1.a.check()
# Lines below will give you a hint or solution code
#step_1.a.hint()
#step_1.a.solution()
```
### Part B
Run the code cell below without changes.
To pass this step, you need to have defined a pipeline in **Part A** that achieves lower MAE than the code above. You're encouraged to take your time here and try out many different approaches, to see how low you can get the MAE! (_If your code does not pass, please amend the preprocessing steps and model in Part A._)
```
# Bundle preprocessing and modeling code in a pipeline
my_pipeline = Pipeline(steps=[('preprocessor', preprocessor),
('model', model)
])
# Preprocessing of training data, fit model
my_pipeline.fit(X_train, y_train)
# Preprocessing of validation data, get predictions
preds = my_pipeline.predict(X_valid)
# Evaluate the model
score = mean_absolute_error(y_valid, preds)
print('MAE:', score)
# Check your answer
step_1.b.check()
# Line below will give you a hint
#step_1.b.hint()
```
# Step 2: Generate test predictions
Now, you'll use your trained model to generate predictions with the test data.
```
# Preprocessing of test data, fit model
preds_test = my_pipeline.predict(X_test)
# Check your answer
step_2.check()
# Lines below will give you a hint or solution code
#step_2.hint()
#step_2.solution()
```
Run the next code cell without changes to save your results to a CSV file that can be submitted directly to the competition.
```
# Save test predictions to file
output = pd.DataFrame({'Id': X_test.index,
'SalePrice': preds_test})
output.to_csv('submission.csv', index=False)
```
# Submit your results
Once you have successfully completed Step 2, you're ready to submit your results to the leaderboard! If you choose to do so, make sure that you have already joined the competition by clicking on the **Join Competition** button at [this link](https://www.kaggle.com/c/home-data-for-ml-course).
1. Begin by clicking on the **Save Version** button in the top right corner of the window. This will generate a pop-up window.
2. Ensure that the **Save and Run All** option is selected, and then click on the **Save** button.
3. This generates a window in the bottom left corner of the notebook. After it has finished running, click on the number to the right of the **Save Version** button. This pulls up a list of versions on the right of the screen. Click on the ellipsis **(...)** to the right of the most recent version, and select **Open in Viewer**. This brings you into view mode of the same page. You will need to scroll down to get back to these instructions.
4. Click on the **Output** tab on the right of the screen. Then, click on the file you would like to submit, and click on the **Submit** button to submit your results to the leaderboard.
You have now successfully submitted to the competition!
If you want to keep working to improve your performance, select the **Edit** button in the top right of the screen. Then you can change your code and repeat the process. There's a lot of room to improve, and you will climb up the leaderboard as you work.
# Keep going
Move on to learn about [**cross-validation**](https://www.kaggle.com/alexisbcook/cross-validation), a technique you can use to obtain more accurate estimates of model performance!
---
*Have questions or comments? Visit the [course discussion forum](https://www.kaggle.com/learn/intermediate-machine-learning/discussion) to chat with other learners.*
| github_jupyter |
# Facies classification using machine learning techniques
The ideas of
<a href="https://home.deib.polimi.it/bestagini/">Paolo Bestagini's</a> "Try 2", <a href="https://github.com/ar4">Alan Richardson's</a> "Try 2",
<a href="https://github.com/dalide">Dalide's</a> "Try 6", augmented, by Dimitrios Oikonomou and Eirik Larsen (ESA AS) by
- adding the gradient of gradient of features as augmented features.
- with an ML estimator for PE using both training and blind well data.
- removing the NM_M from augmented features.
In the following, we provide a possible solution to the facies classification problem described at https://github.com/seg/2016-ml-contest.
The proposed algorithm is based on the use of random forests, xgboost or gradient boost combined in one-vs-one multiclass strategy. In particular, we would like to study the effect of:
- Robust feature normalization.
- Feature imputation for missing feature values.
- Well-based cross-validation routines.
- Feature augmentation strategies.
- Test multiple classifiers
# Script initialization
Let's import the used packages and define some parameters (e.g., colors, labels, etc.).
```
# Import
from __future__ import division
get_ipython().magic(u'matplotlib inline')
import matplotlib as mpl
import matplotlib.pyplot as plt
mpl.rcParams['figure.figsize'] = (20.0, 10.0)
inline_rc = dict(mpl.rcParams)
from classification_utilities import make_facies_log_plot
import pandas as pd
import numpy as np
import seaborn as sns
from sklearn import preprocessing
from sklearn.model_selection import LeavePGroupsOut
from sklearn.metrics import f1_score
from sklearn.model_selection import GridSearchCV
from sklearn.multiclass import OneVsOneClassifier
from sklearn.ensemble import RandomForestClassifier, RandomForestRegressor, GradientBoostingClassifier
import xgboost as xgb
from xgboost.sklearn import XGBClassifier
from scipy.signal import medfilt
import sys, scipy, sklearn
print('Python: ' + sys.version.split('\n')[0])
print(' ' + sys.version.split('\n')[0])
print('Pandas: ' + pd.__version__)
print('Numpy: ' + np.__version__)
print('Scipy: ' + scipy.__version__)
print('Sklearn: ' + sklearn.__version__)
print('Xgboost: ' + xgb.__version__)
```
### Parameters
```
feature_names = ['GR', 'ILD_log10', 'DeltaPHI', 'PHIND', 'PE', 'NM_M', 'RELPOS']
facies_names = ['SS', 'CSiS', 'FSiS', 'SiSh', 'MS', 'WS', 'D', 'PS', 'BS']
facies_colors = ['#F4D03F', '#F5B041','#DC7633','#6E2C00', '#1B4F72','#2E86C1', '#AED6F1', '#A569BD', '#196F3D']
#Select classifier type
clfType='RF' #Random Forest clasifier
#clfType='GB' #Gradient Boosting Classifier
#clfType='XB' #XGB Clasifier
#Seed
seed = 24
np.random.seed(seed)
```
# Load data
Let's load the data
```
# Load data from file
data = pd.read_csv('../facies_vectors.csv')
# Load Test data from file
test_data = pd.read_csv('../validation_data_nofacies.csv')
test_data.insert(0,'Facies',np.ones(test_data.shape[0])*(-1))
#Create Dataset for PE prediction from both dasets
all_data=pd.concat([data,test_data])
```
#### Let's store features, labels and other data into numpy arrays.
```
# Store features and labels
X = data[feature_names].values # features
y = data['Facies'].values # labels
# Store well labels and depths
well = data['Well Name'].values
depth = data['Depth'].values
```
# Data inspection
Let us inspect the features we are working with. This step is useful to understand how to normalize them and how to devise a correct cross-validation strategy. Specifically, it is possible to observe that:
- Some features seem to be affected by a few outlier measurements.
- Only a few wells contain samples from all classes.
- PE measurements are available only for some wells.
```
# Define function for plotting feature statistics
def plot_feature_stats(X, y, feature_names, facies_colors, facies_names):
# Remove NaN
nan_idx = np.any(np.isnan(X), axis=1)
X = X[np.logical_not(nan_idx), :]
y = y[np.logical_not(nan_idx)]
# Merge features and labels into a single DataFrame
features = pd.DataFrame(X, columns=feature_names)
labels = pd.DataFrame(y, columns=['Facies'])
for f_idx, facies in enumerate(facies_names):
labels[labels[:] == f_idx] = facies
data = pd.concat((labels, features), axis=1)
# Plot features statistics
facies_color_map = {}
for ind, label in enumerate(facies_names):
facies_color_map[label] = facies_colors[ind]
sns.pairplot(data, hue='Facies', palette=facies_color_map, hue_order=list(reversed(facies_names)))
```
## Feature distribution
plot_feature_stats(X, y, feature_names, facies_colors, facies_names)
mpl.rcParams.update(inline_rc)
```
# Facies per well
for w_idx, w in enumerate(np.unique(well)):
ax = plt.subplot(3, 4, w_idx+1)
hist = np.histogram(y[well == w], bins=np.arange(len(facies_names)+1)+.5)
plt.bar(np.arange(len(hist[0])), hist[0], color=facies_colors, align='center')
ax.set_xticks(np.arange(len(hist[0])))
ax.set_xticklabels(facies_names)
ax.set_title(w)
# Features per well
for w_idx, w in enumerate(np.unique(well)):
ax = plt.subplot(3, 4, w_idx+1)
hist = np.logical_not(np.any(np.isnan(X[well == w, :]), axis=0))
plt.bar(np.arange(len(hist)), hist, color=facies_colors, align='center')
ax.set_xticks(np.arange(len(hist)))
ax.set_xticklabels(feature_names)
ax.set_yticks([0, 1])
ax.set_yticklabels(['miss', 'hit'])
ax.set_title(w)
```
## Feature imputation
Let us fill missing PE values. This is the only cell that differs from the approach of Paolo Bestagini. Currently no feature engineering is used, but this should be explored in the future.
```
reg = RandomForestRegressor(max_features='sqrt', n_estimators=50, random_state=seed)
DataImpAll = all_data[feature_names].copy()
DataImp = DataImpAll.dropna(axis = 0, inplace=False)
Ximp=DataImp.loc[:, DataImp.columns != 'PE']
Yimp=DataImp.loc[:, 'PE']
reg.fit(Ximp, Yimp)
X[np.array(data.PE.isnull()),feature_names.index('PE')] = reg.predict(data.loc[data.PE.isnull(),:][['GR', 'ILD_log10', 'DeltaPHI', 'PHIND', 'NM_M', 'RELPOS']])
```
# Augment features
```
# ## Feature augmentation
# Our guess is that facies do not abrutly change from a given depth layer to the next one. Therefore, we consider features at neighboring layers to be somehow correlated. To possibly exploit this fact, let us perform feature augmentation by:
# - Select features to augment.
# - Aggregating aug_features at neighboring depths.
# - Computing aug_features spatial gradient.
# - Computing aug_features spatial gradient of gradient.
# Feature windows concatenation function
def augment_features_window(X, N_neig, features=-1):
# Parameters
N_row = X.shape[0]
if features==-1:
N_feat = X.shape[1]
features=np.arange(0,X.shape[1])
else:
N_feat = len(features)
# Zero padding
X = np.vstack((np.zeros((N_neig, X.shape[1])), X, (np.zeros((N_neig, X.shape[1])))))
# Loop over windows
X_aug = np.zeros((N_row, N_feat*(2*N_neig)+X.shape[1]))
for r in np.arange(N_row)+N_neig:
this_row = []
for c in np.arange(-N_neig,N_neig+1):
if (c==0):
this_row = np.hstack((this_row, X[r+c,:]))
else:
this_row = np.hstack((this_row, X[r+c,features]))
X_aug[r-N_neig] = this_row
return X_aug
# Feature gradient computation function
def augment_features_gradient(X, depth, features=-1):
if features==-1:
features=np.arange(0,X.shape[1])
# Compute features gradient
d_diff = np.diff(depth).reshape((-1, 1))
d_diff[d_diff==0] = 0.001
X_diff = np.diff(X[:,features], axis=0)
X_grad = X_diff / d_diff
# Compensate for last missing value
X_grad = np.concatenate((X_grad, np.zeros((1, X_grad.shape[1]))))
return X_grad
# Feature augmentation function
def augment_features(X, well, depth, N_neig=1, features=-1):
if (features==-1):
N_Feat=X.shape[1]
else:
N_Feat=len(features)
# Augment features
X_aug = np.zeros((X.shape[0], X.shape[1] + N_Feat*(N_neig*2+2)))
for w in np.unique(well):
w_idx = np.where(well == w)[0]
X_aug_win = augment_features_window(X[w_idx, :], N_neig,features)
X_aug_grad = augment_features_gradient(X[w_idx, :], depth[w_idx],features)
X_aug_grad_grad = augment_features_gradient(X_aug_grad, depth[w_idx])
X_aug[w_idx, :] = np.concatenate((X_aug_win, X_aug_grad,X_aug_grad_grad), axis=1)
# Find padded rows
padded_rows = np.unique(np.where(X_aug[:, 0:7] == np.zeros((1, 7)))[0])
return X_aug, padded_rows
# Train and test a classifier
def train_and_test(X_tr, y_tr, X_v, well_v, clf):
# Feature normalization
scaler = preprocessing.RobustScaler(quantile_range=(25.0, 75.0)).fit(X_tr)
X_tr = scaler.transform(X_tr)
X_v = scaler.transform(X_v)
# Train classifier
clf.fit(X_tr, y_tr)
# Test classifier
y_v_hat = clf.predict(X_v)
# Clean isolated facies for each well
for w in np.unique(well_v):
y_v_hat[well_v==w] = medfilt(y_v_hat[well_v==w], kernel_size=5)
return y_v_hat
# Define window length
N_neig=1
# Define which features to augment by introducing window and gradients.
augm_Features=['GR', 'ILD_log10', 'DeltaPHI', 'PHIND', 'PE', 'RELPOS']
# Get the columns of features to be augmented
feature_indices=[feature_names.index(log) for log in augm_Features]
# Augment features
X_aug, padded_rows = augment_features(X, well, depth, N_neig=N_neig, features=feature_indices)
# Remove padded rows
data_no_pad = np.setdiff1d(np.arange(0,X_aug.shape[0]), padded_rows)
X=X[data_no_pad ,:]
depth=depth[data_no_pad]
X_aug=X_aug[data_no_pad ,:]
y=y[data_no_pad]
data=data.iloc[data_no_pad ,:]
well=well[data_no_pad]
```
## Generate training, validation and test data splitsar4_submission_withFac.ipynb
The choice of training and validation data is paramount in order to avoid overfitting and find a solution that generalizes well on new data. For this reason, we generate a set of training-validation splits so that:
- Features from each well belongs to training or validation set.
- Training and validation sets contain at least one sample for each class.
# Initialize model selection methods
```
lpgo = LeavePGroupsOut(2)
# Generate splits
split_list = []
for train, val in lpgo.split(X, y, groups=data['Well Name']):
hist_tr = np.histogram(y[train], bins=np.arange(len(facies_names)+1)+.5)
hist_val = np.histogram(y[val], bins=np.arange(len(facies_names)+1)+.5)
if np.all(hist_tr[0] != 0) & np.all(hist_val[0] != 0):
split_list.append({'train':train, 'val':val})
# Print splits
for s, split in enumerate(split_list):
print('Split %d' % s)
print(' training: %s' % (data.iloc[split['train']]['Well Name'].unique()))
print(' validation: %s' % (data.iloc[split['val']]['Well Name'].unique()))
```
## Classification parameters optimization
Let us perform the following steps for each set of parameters:
- Select a data split.
- Normalize features using a robust scaler.
- Train the classifier on training data.
- Test the trained classifier on validation data.
- Repeat for all splits and average the F1 scores.
At the end of the loop, we select the classifier that maximizes the average F1 score on the validation set. Hopefully, this classifier should be able to generalize well on new data.
```
# Parameters search grid (uncomment parameters for full grid search... may take a lot of time)
if clfType=='XB':
md_grid = [3]
mcw_grid = [1]
gamma_grid = [0.3]
ss_grid = [0.7]
csb_grid = [0.8]
alpha_grid =[0.2]
lr_grid = [0.05]
ne_grid = [200]
param_grid = []
for N in md_grid:
for M in mcw_grid:
for S in gamma_grid:
for L in ss_grid:
for K in csb_grid:
for P in alpha_grid:
for R in lr_grid:
for E in ne_grid:
param_grid.append({'maxdepth':N,
'minchildweight':M,
'gamma':S,
'subsample':L,
'colsamplebytree':K,
'alpha':P,
'learningrate':R,
'n_estimators':E})
if clfType=='RF':
N_grid = [100] # [50, 100, 150]
M_grid = [10] # [5, 10, 15]
S_grid = [25] # [10, 25, 50, 75]
L_grid = [5] # [2, 3, 4, 5, 10, 25]
param_grid = []
for N in N_grid:
for M in M_grid:
for S in S_grid:
for L in L_grid:
param_grid.append({'N':N, 'M':M, 'S':S, 'L':L})
if clfType=='GB':
N_grid = [100]
MD_grid = [3]
M_grid = [10]
LR_grid = [0.1]
L_grid = [5]
S_grid = [25]
param_grid = []
for N in N_grid:
for M in MD_grid:
for M1 in M_grid:
for S in LR_grid:
for L in L_grid:
for S1 in S_grid:
param_grid.append({'N':N, 'MD':M, 'MF':M1,'LR':S,'L':L,'S1':S1})
def getClf(clfType, param):
if clfType=='RF':
clf = OneVsOneClassifier(RandomForestClassifier(n_estimators=param['N'], criterion='entropy',
max_features=param['M'], min_samples_split=param['S'], min_samples_leaf=param['L'],
class_weight='balanced', random_state=seed), n_jobs=-1)
if clfType=='XB':
clf = OneVsOneClassifier(XGBClassifier(
learning_rate = param['learningrate'],
n_estimators=param['n_estimators'],
max_depth=param['maxdepth'],
min_child_weight=param['minchildweight'],
gamma = param['gamma'],
subsample=param['subsample'],
colsample_bytree=param['colsamplebytree'],
reg_alpha = param['alpha'],
nthread =1,
seed = seed,
) , n_jobs=-1)
if clfType=='GB':
clf=OneVsOneClassifier(GradientBoostingClassifier(
loss='exponential',
n_estimators=param['N'],
learning_rate=param['LR'],
max_depth=param['MD'],
max_features= param['MF'],
min_samples_leaf=param['L'],
min_samples_split=param['S1'],
random_state=seed,
max_leaf_nodes=None,)
, n_jobs=-1)
return clf
# For each set of parameters
score_param = []
for param in param_grid:
print('features: %d' % X_aug.shape[1])
# For each data split
score_split = []
for split in split_list:
split_train_no_pad = split['train']
# Select training and validation data from current split
X_tr = X_aug[split_train_no_pad, :]
X_v = X_aug[split['val'], :]
y_tr = y[split_train_no_pad]
y_v = y[split['val']]
# Select well labels for validation data
well_v = well[split['val']]
# Train and test
y_v_hat = train_and_test(X_tr, y_tr, X_v, well_v, getClf(clfType,param))
# Score
score = f1_score(y_v, y_v_hat, average='micro')
score_split.append(score)
print('Split: %d, Score = %.3f' % (split_list.index(split),score))
# Average score for this param
score_param.append(np.mean(score_split))
print('Average F1 score = %.3f %s' % (score_param[-1], param))
# Best set of parameters
best_idx = np.argmax(score_param)
param_best = param_grid[best_idx]
score_best = score_param[best_idx]
print('\nBest F1 score = %.3f %s' % (score_best, param_best))
# ## Predict labels on test data
# Let us now apply the selected classification technique to test data.
# Training data
X_tr = X_aug
y_tr = y
# Prepare test data
well_ts = test_data['Well Name'].values
depth_ts = test_data['Depth'].values
X_ts = test_data[feature_names].values
# Augment Test data features
X_ts, padded_rows = augment_features(X_ts, well_ts,depth,N_neig=N_neig, features=feature_indices)
# Predict test labels
y_ts_hat = train_and_test(X_tr, y_tr, X_ts, well_ts, getClf(clfType,param_best))
# Save predicted labels
test_data['Facies'] = y_ts_hat
test_data.to_csv('esa_predicted_facies_{0}_sub01.csv'.format(clfType))
# Plot predicted labels
make_facies_log_plot(
test_data[test_data['Well Name'] == 'STUART'],
facies_colors=facies_colors)
make_facies_log_plot(
test_data[test_data['Well Name'] == 'CRAWFORD'],
facies_colors=facies_colors)
mpl.rcParams.update(inline_rc)
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import emcee
import scipy.optimize as op
import scipy
import corner
from multiprocessing import Pool
import multiprocessing
import time
from astropy.table import Table
import astropy
import matplotlib.pyplot as plt
from matplotlib.ticker import MultipleLocator
import matplotlib
%matplotlib notebook
print('numpy version: {}'.format(np.__version__))
print('pandas version: {}'.format(pd.__version__))
print('matplotlib version: {}'.format(matplotlib.__version__))
print('astropy version: {}'.format(astropy.__version__))
print('emcee version: {}'.format(emcee.__version__))
print('scipy version: {}'.format(scipy.__version__))
```
# Figure 9
Create Figure 9 (the Redshift Completeness Factor as a function of observed galaxy brightness based on ZTF BTS data) in [Fremling et al. 2020](https://ui.adsabs.harvard.edu/abs/2019arXiv191012973F/abstract).
```
# BTS data
bts_df = pd.read_hdf('../data/final_rcf_table.h5')
z_sn = bts_df.z_sn.values
z_host = bts_df.z_host.values
norm_Ia = np.where( ( (bts_df.sn_type == 'Ia-norm') |
(bts_df.sn_type == 'Ia') |
(bts_df.sn_type == 'Ia-91bg') |
(bts_df.sn_type == 'Ia-91T') |
(bts_df.sn_type == 'Ia-99aa') |
(bts_df.sn_type == 'ia')
| (bts_df.sn_type == 'Ia-norm*')
| (bts_df.sn_type == 'Ia-91T*')
| (bts_df.sn_type == 'Ia-91T**')
| (bts_df.sn_type == 'SN Ia')
)
)
norm_cc = np.where( (bts_df.sn_type == 'IIb') |
(bts_df.sn_type == 'Ib') |
(bts_df.sn_type == 'IIP') |
(bts_df.sn_type == 'Ib/c') |
(bts_df.sn_type == 'Ic-norm') |
(bts_df.sn_type == 'IIn') |
(bts_df.sn_type == 'IIL') |
(bts_df.sn_type == 'Ic-broad') |
(bts_df.sn_type == 'II') |
(bts_df.sn_type == 'II-pec') |
(bts_df.sn_type == 'Ib-pec') |
(bts_df.sn_type == 'Ic') |
(bts_df.sn_type == 'Ic-BL') |
(bts_df.sn_type == 'IIP*') |
(bts_df.sn_type == 'II*') |
(bts_df.sn_type == 'Ibn') |
(bts_df.sn_type == 'II**') |
(bts_df.sn_type == 'Ib-norm') |
(bts_df.sn_type == 'IIn*')
)
has_host_z = np.where((z_host > 0) & np.isfinite(z_host))
no_host = np.where((z_host < 0) | np.isnan(z_host))
has_host_cc = np.intersect1d(has_host_z, norm_cc)
has_host_ia = np.intersect1d(has_host_z, norm_Ia)
no_host_cc = np.intersect1d(no_host, norm_cc)
no_host_ia = np.intersect1d(no_host, norm_Ia)
z_mix = z_sn.copy()
z_mix[has_host_z] = z_host[has_host_z]
# prep data for analysis
hit_idx = np.where(np.isfinite(bts_df.z_host)
& (bts_df.z_host > 0)
& np.isfinite(bts_df.sep)
& (bts_df.sn_type != 'ambiguous')
)
miss_idx = np.where(np.isfinite(bts_df.z_host)
& (bts_df.z_host == -999)
& np.isfinite(bts_df.sep)
& (bts_df.sn_type != 'ambiguous')
)
ps1_r_det = np.where(np.isfinite(bts_df.rKron_PS1) & (bts_df.rKron_PS1.values > 0))
ps1_r_hit = np.intersect1d(hit_idx, ps1_r_det[0])
ps1_r_miss = np.intersect1d(miss_idx, ps1_r_det[0])
print('There are {} galaxies with r-band detections'.format(len(ps1_r_det[0])))
ps1_i_det = np.where(np.isfinite(bts_df.iKron_PS1) & (bts_df.iKron_PS1.values > 0))
ps1_i_hit = np.intersect1d(hit_idx, ps1_i_det[0])
ps1_i_miss = np.intersect1d(miss_idx, ps1_i_det[0])
print('There are {} galaxies with i-band detections'.format(len(ps1_i_det[0])))
# values for analysis
in_ned_ps1 = np.append(np.ones_like(z_mix[ps1_r_hit]),
np.zeros_like(z_mix[ps1_r_miss])
)
r_mag_ps1 = np.append(bts_df.rKron_PS1[ps1_r_hit], bts_df.rKron_PS1[ps1_r_miss])
z_ps1 = np.append(z_mix[ps1_r_hit], z_mix[ps1_r_miss])
```
### Get p(m_obs)
```
def p_z(x, a, c):
return 1 / (1 + np.exp(a*x - c))
def lnprior_z(theta):
a, c = theta
if (0 < a < 1e6) and (-100 < c < 100):
return 0.
return -np.inf
def lnlike_z(theta, trials, x):
a, c = theta
return np.sum(np.log([scipy.stats.bernoulli.pmf(t, _p) for t, _p in zip(trials, p_z(x, a, c))]))
def lnprob_z(theta, trials, x):
lp = lnprior_z(theta)
if not np.isfinite(lp):
return -np.inf
return lp + lnlike_z(theta, trials, x)
nll = lambda *args: -lnlike_z(*args)
result = op.minimize(nll, [(1, 16)], args=(in_ned_ps1, r_mag_ps1))
b, c = result["x"]
ndim, nwalkers = 2, 20
pos_ps1 = [result["x"] + 1e-4*np.random.randn(ndim) for i in range(nwalkers)]
max_n = 10000
old_tau = np.inf
backend = emcee.backends.HDFBackend("../data/magCF_PS1_chains.h5")
with Pool(4) as pool:
sampler_ps1 = emcee.EnsembleSampler(nwalkers, ndim, lnprob_z,
args=(in_ned_ps1, r_mag_ps1),
pool=pool,
backend=backend)
start = time.time()
for sample in sampler_ps1.sample(pos_ps1, iterations=max_n, progress=True):
if sampler_ps1.iteration % 100:
continue
tau = sampler_ps1.get_autocorr_time(tol=0)
# Check convergence
converged = np.all(tau * 100 < sampler_ps1.iteration)
converged &= np.all(np.abs(old_tau - tau) / tau < 0.01)
if converged:
break
old_tau = tau
end = time.time()
multi_time = end - start
print("Multiprocessing took {0:.1f} seconds".format(multi_time))
ps1_grid = np.linspace(8, 24, 1000)
autocorr_ps1 = sampler_ps1.get_autocorr_time(tol=0)
n_burn_ps1 = 5*np.ceil(np.max(autocorr_ps1)).astype(int)
samples_ps1 = sampler_ps1.get_chain(discard=50, flat=True)
p_ps1_1d_grid = np.empty((len(ps1_grid),len(samples_ps1)))
for grid_num, ps1_w1 in enumerate(ps1_grid):
p_ps1_1d_grid[grid_num] = p_z(ps1_w1, samples_ps1[:,0], samples_ps1[:,1])
p_ps15, p_ps150, p_ps195 = np.percentile(p_ps1_1d_grid, (5,50,95), axis=1)
```
### Same thing, but just things within the SDSS footprint
```
in_sdss_df = Table.read('../data/RCF_has_SDSS_adammiller.fit').to_pandas()
sdss_footprint = np.zeros_like(bts_df.rKron_PS1).astype(bool)
for zn in in_sdss_df.ZTF_Name:
sdss_footprint[bts_df.ZTF_Name == zn.decode('utf-8')] = 1
ps1_sdss = np.where(np.isfinite(bts_df.rKron_PS1) & (bts_df.rKron_PS1 > 0) & sdss_footprint)
ps1_sdss_hit = np.intersect1d(hit_idx, ps1_sdss[0])
ps1_sdss_miss = np.intersect1d(miss_idx, ps1_sdss[0])
print('There are {} sources with SDSS imaging'.format(len(ps1_sdss[0])))
in_ned_ps1_sdss = np.append(np.ones_like(z_mix[ps1_sdss_hit]),
np.zeros_like(z_mix[ps1_sdss_miss])
)
r_mag_ps1_sdss = np.append(bts_df.rKron_PS1[ps1_sdss_hit], bts_df.rKron_PS1[ps1_sdss_miss])
z_ps1_sdss = np.append(z_mix[ps1_sdss_hit], z_mix[ps1_sdss_miss])
nll = lambda *args: -lnlike_z(*args)
result = op.minimize(nll, [(1, 16)], args=(in_ned_ps1_sdss, r_mag_ps1_sdss))
b, c = result["x"]
ndim, nwalkers = 2, 20
pos_ps1_sdss = [result["x"] + 1e-4*np.random.randn(ndim) for i in range(nwalkers)]
max_n = 10000
old_tau = np.inf
backend = emcee.backends.HDFBackend("../data/magCF_in_SDSS_chains.h5")
with Pool(4) as pool:
sampler_ps1_sdss = emcee.EnsembleSampler(nwalkers, ndim, lnprob_z,
args=(in_ned_ps1_sdss, r_mag_ps1_sdss),
pool=pool,
backend=backend)
start = time.time()
for sample in sampler_ps1_sdss.sample(pos_ps1, iterations=max_n, progress=True):
if sampler_ps1_sdss.iteration % 100:
continue
tau = sampler_ps1_sdss.get_autocorr_time(tol=0)
# Check convergence
converged = np.all(tau * 100 < sampler_ps1_sdss.iteration)
converged &= np.all(np.abs(old_tau - tau) / tau < 0.01)
if converged:
break
old_tau = tau
end = time.time()
multi_time = end - start
print("Multiprocessing took {0:.1f} seconds".format(multi_time))
autocorr_ps1_sdss = sampler_ps1_sdss.get_autocorr_time(tol=0)
n_burn_ps1_sdss = 5*np.ceil(np.max(autocorr_ps1_sdss)).astype(int)
samples_ps1_sdss = sampler_ps1_sdss.get_chain(discard=50, flat=True)
p_ps1_sdss_1d_grid = np.empty((len(ps1_grid),len(samples_ps1_sdss)))
for grid_num, ps1_w1 in enumerate(ps1_grid):
p_ps1_sdss_1d_grid[grid_num] = p_z(ps1_w1, samples_ps1_sdss[:,0], samples_ps1_sdss[:,1])
p_ps1_sdss5, p_ps1_sdss50, p_ps1_sdss95 = np.percentile(p_ps1_sdss_1d_grid, (5,50,95), axis=1)
```
#### What about outside SDSS?
```
no_sdss_footprint = sdss_footprint*-1+1
no_sdss = np.where(np.isfinite(bts_df.rKron_PS1) & (bts_df.rKron_PS1 > 0) & no_sdss_footprint)
no_sdss_hit = np.intersect1d(hit_idx, no_sdss[0])
no_sdss_miss = np.intersect1d(miss_idx, no_sdss[0])
print('There are {} sources outside SDSS imaging'.format(len(no_sdss[0])))
in_ned_no_sdss = np.append(np.ones_like(z_mix[no_sdss_hit]),
np.zeros_like(z_mix[no_sdss_miss])
)
r_mag_no_sdss = np.append(bts_df.rKron_PS1[no_sdss_hit], bts_df.rKron_PS1[no_sdss_miss])
z_no_sdss = np.append(z_mix[no_sdss_hit], z_mix[no_sdss_miss])
nll = lambda *args: -lnlike_z(*args)
result = op.minimize(nll, [(1, 16)], args=(in_ned_no_sdss, r_mag_no_sdss))
b, c = result["x"]
ndim, nwalkers = 2, 20
pos_ps1_sdss = [result["x"] + 1e-4*np.random.randn(ndim) for i in range(nwalkers)]
max_n = 10000
old_tau = np.inf
backend = emcee.backends.HDFBackend("../data/magCF_outside_SDSS_chains.h5")
with Pool(4) as pool:
sampler_no_sdss = emcee.EnsembleSampler(nwalkers, ndim, lnprob_z,
args=(in_ned_no_sdss, r_mag_no_sdss),
pool=pool,
backend=backend)
start = time.time()
for sample in sampler_no_sdss.sample(pos_ps1, iterations=max_n, progress=True):
if sampler_no_sdss.iteration % 100:
continue
tau = sampler_no_sdss.get_autocorr_time(tol=0)
# Check convergence
converged = np.all(tau * 100 < sampler_no_sdss.iteration)
converged &= np.all(np.abs(old_tau - tau) / tau < 0.01)
if converged:
break
old_tau = tau
end = time.time()
multi_time = end - start
print("Multiprocessing took {0:.1f} seconds".format(multi_time))
autocorr_no_sdss = sampler_no_sdss.get_autocorr_time(tol=0)
n_burn_no_sdss = 5*np.ceil(np.max(autocorr_no_sdss)).astype(int)
samples_no_sdss = sampler_no_sdss.get_chain(discard=n_burn_no_sdss, flat=True)
p_no_sdss_1d_grid = np.empty((len(ps1_grid),len(samples_no_sdss)))
for grid_num, ps1_w1 in enumerate(ps1_grid):
p_no_sdss_1d_grid[grid_num] = p_z(ps1_w1, samples_no_sdss[:,0], samples_no_sdss[:,1])
p_no_sdss5, p_no_sdss50, p_no_sdss95 = np.percentile(p_no_sdss_1d_grid, (5,50,95), axis=1)
```
## Make the figure
```
fig, ax = plt.subplots()
ax.plot(ps1_grid[1:-1], p_ps150[1:-1],
lw=3,
label='in PS1 footprint', color='#7570b3')
ax.fill_between(ps1_grid, p_ps15, p_ps195,
alpha=0.5, color='#7570b3')
ax.plot(ps1_grid[1:-1], p_ps1_sdss50[1:-1],
'-.', alpha=0.75, lw=2,
label='in SDSS footprint', color='#66c2a5')
ax.fill_between(ps1_grid, p_ps1_sdss5, p_ps1_sdss95,
alpha=0.2, color='#66c2a5')
ax.plot(ps1_grid[1:-1], p_no_sdss50[1:-1],
'--', alpha=0.75, lw=2,
label='outside SDSS footprint', color='#fc8d62')
ax.fill_between(ps1_grid, p_no_sdss5, p_no_sdss95,
alpha=0.2, color='#fc8d62')
ax.set_xlabel('$r_{ps1}$ (mag)', fontsize=17)
ax.set_ylabel('RCF', fontsize=16)
ax.tick_params(labelsize=12)
ax.set_xlim(11,22)
ax.set_ylim(0,1.02)
ax.xaxis.set_major_locator(MultipleLocator(1))
ax.xaxis.set_minor_locator(MultipleLocator(0.25))
ax.yaxis.set_minor_locator(MultipleLocator(0.05))
ax.yaxis.set_major_locator(MultipleLocator(0.1))
ax.legend(fancybox=True)
fig.subplots_adjust(left=0.1, right=0.98, top=0.99, bottom=0.12)
fig.savefig('RCF_r_band.pdf')
print('90, 50, 10% completeness = {:.2f}, {:.2f}, {:.2f} mag, respectively'.format(ps1_grid[np.argmin(np.abs(p_ps150-0.9))],
ps1_grid[np.argmin(np.abs(p_ps150-0.5))],
ps1_grid[np.argmin(np.abs(p_ps150-0.1))]))
```
| github_jupyter |
## Importing the necesssary Libraries
```
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
%matplotlib inline
import psycopg2 as pg2
import datetime as dt
# package used for converting the data into datetime format
from sklearn.preprocessing import LabelEncoder
from sklearn.feature_selection import RFE, f_regression
from sklearn.linear_model import (LinearRegression, Ridge, Lasso, RandomizedLasso)
from sklearn.preprocessing import MinMaxScaler
from sklearn.ensemble import RandomForestRegressor
from sklearn.metrics import r2_score
from sklearn.metrics import mean_squared_error
import math
import warnings
warnings.filterwarnings("ignore")
## Execution time counter
#%load_ext autotime
```
## Establish connection to the PostgreSQL database
```
conn= pg2.connect('dbname = Booksville user= postgres password =891630 host= 127.0.0.1')
cur=conn.cursor()
df_raw = pd.read_sql_query('select * from public."keepa" limit 1000', conn)
#Check the dimension of the raw data to see if its properly imported
print('Starting size of our Dataset ')
df_raw.shape
# Print out count of each datatype in the dataframe
df_raw.dtypes.value_counts()
```
## Price Aggregator
Amazon's listing price for books is stored in one of the three columns in the sql dump we obtained from Keepa. If Amazon is fullfilling the sale, the price is stored in amazon_price column. But if the book is sold by a third party seller, the listing price would be marketplace_new_price for new books and marketplace_used_price for used books.
We are combining the three columns in to one column called 'price' and assign its values based on the three given price assignment information.
The aggregator function adds the new column to the dataset and assigns the value that appears first from the following list and finally drops the three columns from the dataset.
* amazon_Price
* marketplace_new
* marketplace_used_price
```
def PriceAggregator(original_df):
df=original_df
# create a copy of the three columns to choose amazon price from
df_copy=df[['amazon_price','marketplace_new_price','marketplace_used_price']]
# Replace missing price denoted by -1 to Null in all three price columns
for item in df_copy:
df_copy[item].replace('-1',np.nan, inplace=True)
# Add a new column to store the aggregated price with default value of 'amazon_price'
df.insert(79,'price',df_copy['amazon_price'].astype('float'))
#Loop throgh all three columns to assign non-null value to the newly created price column.
#Keep amazon_price as it is if it is not null, otherwise assign marketplace_new_price as the new price.
#Where both 'amazon_price' and 'marketplace_new_price' are null, price will be set to
#'marketplace_used_price' regardless of its value.
for i in range(df['price'].size):
if pd.isnull(df['price'][i]):
if pd.isnull(df_copy['marketplace_new_price'][i]):
if pd.isnull(df_copy['marketplace_used_price'][i]):
pass
else:
df['price'][i]=df_copy['marketplace_used_price'][i]
else:
df['price'][i]=df_copy['marketplace_new_price'][i]
else:
pass
# Delete records where price value is missing since that is what we are trying to predict
df.dropna(subset=['price'], axis=0, inplace=True)
#Reset index after dropping rows with missing price
df.reset_index(drop= True, inplace=True)
#Delete old price columns after assigning aggregated price to a brand new column
df.drop(['amazon_price','marketplace_new_price','marketplace_used_price'], axis=1 , inplace=True)
#Return the a dataframe with a new price column added to the original dataframe
return df
df=PriceAggregator(df_raw)
df.shape
```
## Delete duplicate records, if there are any.
```
# data size before deleting duplicates
df.shape
df.drop_duplicates(inplace = True)
# data size after deleting duplicates
df.shape
```
We can see from .shape() function there are 99600 unique records with 77 features in the dataset.
## Understanding the Data
```
# Highlight of the dataframe
df.head(3)
```
Descriptive statistics that summarize the central tendency, dispersion and shape of a dataset’s distribution,
```
df.describe(include= 'all' )
```
The data was stored in the PostgreSQL database as text(string) type and running discriptive statistics with .describe().The dataframe doesn't tell us much until we do the proper type of conversion.
# Data Wrangling
Data wrangling is the process of converting data from the initial format to a format that may be better for analysis. As part of the wrangling process we are applying different techniques to come up with a cleaner and complete dataset to apply machine learning. Here are some of the steps we are following;
- Identify Missing Values
- Replace or Delete Missing Values
- Correct Data format
- Aggregate highly related categorical values where necessary
### Replace missing values with Null Values.
We will replace every missing values with Numpy Null in order to keep uniformity and computational speed.
```
df.fillna(np.NaN)
df.replace('', np.NaN, inplace=True)
## count and sort null values in every coulumn in descending order
df.isna().sum().sort_values(ascending=False).to_frame(name='Count of Null Values')
```
Null or missing value implies that we dont have information about the feature. We can delete the features that contain Null values for the majority of the records, because keeping them will not provide anything about the data.
```
## create a list of features that contain null value for more than 50% of the records based on the the above observation
Null_features=['coupon','offers','liveOffersOrder','promotions','buyBoxSellerIdHistory','features','upcList','variations',
'hazardousMaterialType','genre','platform','variationCSV','parentAsin','department','size','model','color'
,'partNumber','mpn','brand','edition','format']
# Column names with their number of null values
df[Null_features].isna().sum()
```
We can delete these features without losing any useful information from our data since more than 50% of the records in the above list contain null values.
```
## delete columns that contain very high count of null values
df.drop(Null_features, axis=1, inplace=True)
# check the shape of the dataset to confirm the features are dropped
df.shape
```
For the remaining Null values in our data where the total count is relatively small, we will replace them by a statistically representative values like mean or mode.
* Mode value is for categorical columns where there is a clear majority of null values or will be replaced by 'Unknown'
* Mean value is used for numerical columns
```
#Assigns column names that contain null values to a list
with_Nulls=df.loc[:, df.isna().sum()!=0].columns.tolist()
#Lists down the number of null values in every column in descending order
df[with_Nulls].isna().sum().sort_values(ascending=False)
# let's see what kind of information is in each column
df[with_Nulls].head(5)
```
The sample shows that the records are mainly comprised of string or categorical values. Lets further divide the series based on the number of missing (Null) values.
```
Nulls2Unknown=['categoryTree_4','categoryTree_3','categoryTree_2','author','studio','publisher','manufacturer',
'label']
#print out the highest frequency value(Mode) in the given list of features, it only shows the count not the value.
#Based on the count we can tell if there's a statistical representative mode value to replace the nulls.
for item in with_Nulls:
print(f'{item}\t\t{df[item].value_counts().max()}')
```
Given that our data contains 100,000 records we can clearly see the high mode value will replace the null values for some of the features.
```
# The following 3 features have very high Mode value, therefore we'll replace nulls by mode
Nulls2Mode=['languages_0','categoryTree_0','categoryTree_1']
mode = df.filter(['languages_0','categoryTree_0','categoryTree_1']).mode()
df[Nulls2Mode]=df[Nulls2Mode].fillna(df.mode().iloc[0])
```
For the following features since there is no one single category with a high frequency(Mode) in the group, we are filling the missing(Null) values with 'Unknown'.
```
NullswithNoMode=df.loc[:, df.isna().sum()!=0].columns.tolist()
#Based on the top 3 most frequent records in each column, it shows that there is no dominant value that can be used
#as a mode to replace null values. Therefore, we are replacing null values with 'Unknown'.
for item in NullswithNoMode:
print(item)
print(df[item].value_counts().nlargest(3))
print('Total Number of null values in %s = %d' % (item,df[item].isna().sum()))
print('')
# Replace nulls with 'Unknown' for multimodel features
df[NullswithNoMode]=df[NullswithNoMode].fillna('Unknown')
# Check if there are still missing or null values in the dataset
df[df.loc[:, df.isna().sum()!=0].columns].isna().sum()
```
We have entirely replaced the null and missing values in the dataset by statistically representative values.
## Data Type Conversion
```
df.dtypes.value_counts()
```
The data imported from postgreSQL to pandas dataframe contain columns as object type(string). Most of those features are actually nemerical values, and we will convert the object data type in to the proper format.
Lets group all those features that are in string (object) format and convert them to numeric
```
df.dtypes.value_counts()
#Convert columns that contain numerical values to numeric data type using pandas to_numeric
numeric=['availabilityAmazon',
'ean','hasReviews', 'isEligibleForSuperSaverShipping', 'isEligibleForTradeIn',
'isRedirectASIN', 'isSNS', 'lastPriceChange','lastRatingUpdate', 'lastUpdate', 'listedSince',
'newPriceIsMAP', 'numberOfItems','numberOfPages', 'offersSuccessful', 'packageHeight',
'packageLength', 'packageQuantity', 'packageWeight', 'packageWidth',
'publicationDate', 'releaseDate', 'rootCategory','stats_atIntervalStart', 'stats_avg', 'stats_avg30', 'stats_avg90',
'stats_avg180', 'stats_current', 'stats_outOfStockPercentage30',
'stats_outOfStockPercentage90', 'stats_outOfStockPercentageInInterval',
'trackingSince','sales_rank', 'price']
#cols = ['productType','rootCategory','stats_atIntervalStart','availabilityAmazon','hasReviews','isRedirectASIN','isSNS','isEligibleForTradeIn','isEligibleForSuperSaverShipping', 'ean','hasReviews', 'availabilityAmazon','isEligibleForTradeIn','lastPriceChange','lastRatingUpdate','lastUpdate','lastRatingUpdate','lastUpdate','listedSince',"newPriceIsMAP", "numberOfItems", "numberOfPages","packageHeight", "packageLength","packageQuantity", "packageWeight", "packageWidth",'stats_avg', 'stats_avg30', 'stats_avg90', 'stats_avg180', 'stats_current',"stats_outOfStockPercentage30", "stats_outOfStockPercentage90","stats_outOfStockPercentageInInterval","trackingSince",'upc','price','amazon_price', 'marketplace_new_price', 'marketplace_used_price', 'sales_rank']
df[numeric] = df[numeric].apply(pd.to_numeric, errors='coerce', axis=1)
df.dtypes.value_counts()
strings=df.loc[:, df.dtypes == np.object].columns.tolist()
print('\n'+ 'Sample of the dataset with only categorical information'+'\n')
df[strings].head(3)
```
We can delete 'asin', 'ean' and 'imageCSV' columns since the information they contain is not characteristic discription of books.
```
df.drop(['asin','imagesCSV','ean', 'upc'], axis=1, inplace=True)
df.shape
df.dtypes.value_counts()
# Print features that are object type
df.loc[:, df.dtypes == np.object].columns
```
Information regarding what language a book is written in and whether its an original print or translated version is recorded in the 'language_0' column. These two information are separated by comma, this column can be split into two parts and stored separately in order to reduce categorical data that we have to encode later.
```
df['languages_0'].head(5)
new = df['languages_0'].str.split(",", n = 1, expand = True)
df['language_1']=new[0]
df['language_2']=new[1]
#reduce categories from 9 to 6 groupes by combining related categories
#df['language_1'].value_counts().to_frame()
#group English, english and Middle English to one categry
df['language_1'].replace(('English', 'english','Middle English'),'English', inplace = True)
#group Spanish,Portuguese and Latin under "Spanish"
df['language_1'].replace(('Spanish', 'Portuguese','Latin'),'Spanish', inplace = True)
#group Chinese, mandarin Chinese and simplified chinese under Chinese
df['language_1'].replace(('Simplified Chinese', 'Mandarin Chinese','Chinese'),'Chinese', inplace = True)
#group Arabic,Hebrew and Turkish under Middle Eastern
df['language_1'].replace(('Arabic', 'Hebrew','Turkish'),'Middle Eastern', inplace = True)
# group languages with single entry record in to one group called 'Others'
df['language_1'].replace(('Hindi', 'Scots','Filipino','Malay','Dutch','Greek','Korean','Romanian','Czech'),'Others', inplace = True)
#group Danish and Norwegian under 'Scandinavian'
df['language_1'].replace(('Danish', 'Norwegian'),'Scandinavian', inplace=True)
#replace ('published','Published,Dolby Digital 1.0','Published,DTS-HD 5.1') by Published
df['language_2'].replace(('published','Published,Dolby Digital 1.0','Published,DTS-HD 5.1'),'Published', inplace=True)
df[['language_1','language_2']].head(5)
#Since we have copied the information into new columns we can delete the languages_0 column
df.drop(['languages_0'], axis=1 , inplace=True)
df.columns
df.shape
#Cleaning binding column
df.binding.value_counts()
```
The binding column contains 73 differnt categories that are mostly related and some of them contain very small elements. We will aggregate closely related categories to reduce the dimension of our variables to avoid curse of dimensionality
```
df.binding.nunique()
# create a dictionary of identical items to create a group an aggregated category
dict={'Unknown':['Printed Access Code', 'Unknown','Health and Beauty', 'Lawn & Patio', 'Workbook', 'Kitchen', 'Automotive', 'Jewelry'],
'spiral':[ 'Spiral-bound', 'Staple Bound', 'Ring-bound', 'Plastic Comb', 'Loose Leaf', 'Thread Bound'],
'magazines':[ 'Journal', 'Single Issue Magazine', 'Print Magazine'],
'audios':[ 'Audible Audiobook', 'Audio CD', 'DVD', 'Album', 'MP3 CD', 'Audio CD Library Binding'],
'digital_prints':[ 'CD-ROM', 'Blu-ray', 'DVD-ROM', 'Kindle Edition', 'Video Game', 'Sheet music', 'Software Download',
'Personal Computers', 'Electronics', 'Game', 'Wireless Phone Accessory'],
'hardcovers':['Hardcover', 'Hardcover-spiral', 'Turtleback', 'Roughcut'],
'others':[ 'Cards', 'Pamphlet', 'Calendar', 'Map', 'Stationery', 'Accessory', 'Misc. Supplies', 'Office Product', 'Poster',
'Wall Chart', 'Bookmark', 'JP Oversized'],
'paperbacks':[ 'Paperback', 'Perfect Paperback', 'Mass Market Paperback', 'Flexibound', 'Print on Demand (Paperback)',
'Comic', 'Puzzle', 'Paperback Bunko'],
'leather_bonded':[ 'Bonded Leather', 'Leather Bound', 'Imitation Leather', 'Vinyl Bound'],
'board_book':[ 'Board book', 'Baby Product', 'Toy', 'Rag Book', 'Card Book', 'Bath Book', 'Pocket Book'],
'schoolLibrary_binding':[ 'School & Library Binding', 'Library Binding', 'Textbook Binding']}
for key,val in dict.items():
df.binding.replace(val,key, inplace=True)
df.binding.value_counts()
df.head()
#catTree_under10.categoryTree_2.values= 'Other'
def groupUnder10(x):
cond = df[x].value_counts()
threshold = 10
df[x] = np.where(df[x].isin(cond.index[cond > threshold ]), df[x], 'Others')
print('All the different categories that contain less than 10 items in the %s column are grouped together and renamed to "Others".' %x)
df[['categoryTree_1','categoryTree_2','categoryTree_3','categoryTree_4']].nunique()
groupUnder10('categoryTree_2')
#group under 10 counts in to one for categoryTree_3 column
groupUnder10('categoryTree_3')
groupUnder10('categoryTree_4')
df[['categoryTree_0','categoryTree_1','categoryTree_2','categoryTree_3','categoryTree_4']].nunique()
## Some features are duplicated within the dataset, lets delete those duplicated columns
## Delete duplicated features
duplicates=df[['label', 'manufacturer', 'publisher', 'studio']]
df['label'].equals(df['manufacturer'])
df['label'].equals(duplicates['publisher'])
df['label'].equals(duplicates['studio'])
df[df.duplicated(['label', 'manufacturer', 'publisher', 'studio'])]
duplicates.describe(include='all')
df.duplicated(subset=['label', 'manufacturer', 'publisher', 'studio'],keep='first').value_counts()
```
Since the above 4 columns contain 89493 duplicated informartion out of 99600 total records we can keep one of those and drop the reamining ones without losing useful information.
```
# Keep publisher and drop the rest
df.drop(['label', 'manufacturer','studio'], axis =1, inplace=True)
df.shape
df.describe(include='all').transpose()
```
# Exploratory Data Analysis
## Outlier detection and transformation
Before we decide whether to use standard deviation or interquntile range to identify outliers, lets plot the data points using a distribution plot.
```
def distWithBox(data):
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
sns.set(style="ticks")
x = df[data]
f, (ax_box, ax_hist) = plt.subplots(2, sharex=True,
gridspec_kw={"height_ratios": (.15, .85)})
sns.boxplot(x, ax=ax_box)
sns.distplot(x, ax=ax_hist)
ax_box.set(yticks=[])
sns.despine(ax=ax_hist)
sns.despine(ax=ax_box, left=True)
## Distribution and box plot of the raw data with outliers
distWithBox('price')
```
For normally distributed data, the skewness should be about 0. A skewness value > 0 means that there is more weight in the left tail of the distribution. The function skewtest can be used to determine if the skewness value is close enough to 0, statistically speaking.
```
from scipy.stats import skew
from scipy.stats import skewtest
skew(df['price'])
import seaborn as sns
sns.set(style="whitegrid")
ax = sns.boxplot(x=df['price'])
```
We can see from the the distribution plot, the skewtest and the box plot that price is not normally distributed. The price data is right skewed and there are outlier values that need to be handled.
When a data set has outliers or extreme values, we summarize a typical value using the median as opposed to the mean. When a data set has outliers, variability is often summarized by a statistic called the interquartile range, which is the difference between the first and third quartiles. The first quartile, denoted Q1, is the value in the data set that holds 25% of the values below it. The third quartile, denoted Q3, is the value in the data set that holds 25% of the values above it. The quartiles can be determined following the same approach that we used to determine the median, but we now consider each half of the data set separately. The interquartile range is defined as follows:
Interquartile Range(IQR) = Q3-Q1
Outliers are values 1.5*IQR below Q1 or above Q3 or equivalently, values below Q1-1.5 IQR or above Q3+1.5 IQR.
These are referred to as Tukey fences.
```
from numpy import percentile
data=df['price']
q25, q75 = percentile(data, 25), percentile(data, 75)
iqr = q75 - q25
print('Percentiles:\n\t25th=%.3f \n\t75th=%.3f \n\tIQR=%.3f' % (q25, q75, iqr))
# calculate the outlier cutoff
cut_off = iqr * 1.5
lower, upper = q25 - cut_off, q75 + cut_off
# identify outliers
outliers = [x for x in data if x < lower or x > upper]
print('Identified outliers: %d' % len(outliers) )
outliers_removed = [x for x in data if x >= lower and x <= upper]
print('Non-outlier observations: %d' % len(outliers_removed))
outliers=[]
data_1=df['price']
for item in data_1:
if item <lower or item>upper:
outliers.append(item)
x=df['price']
outlier_indices=list(data_1.index[(x<lower) | (x> upper)])
len(outlier_indices)
df.drop(axis=0,index=outlier_indices, inplace=True)
df.shape
## lets plot distribution with and box plot to see the change after we trim down the outliers
distWithBox('price')
```
### Correlation Between Numerical Features
We are running pearson correlation between numeric valued features to see if there is any linear dependence between the variables.
```
cor=df.corr()
sns.heatmap(cor,cmap="PiYG")
```
Boxplots will show us the distribution of categorical data against a continuous variable. We are using a boxplot to visualize the distribution of values in binding and language columns against price.
Based on the visualization we can see that there is not so much overlap in the binding category, which implies that it is a good predictor of price. But when it comes to language_1, books in almost in every language fall within a price range of 500 to 2000( which is $5 to $20). It implies that knowing what language a book is written in doesn't tell us how much it would worth.
```
sns.boxplot(x=df['price'],y=df['binding'], data=df)
sns.boxplot(y=df['language_1'],x=df['price'], data=df)
```
## Encoding categorical columns
```
cat_cols=['author','language_1','language_2','binding','categoryTree_0', 'categoryTree_1', 'categoryTree_2', 'categoryTree_3',
'categoryTree_4','productGroup','publisher','title','type','language_1','language_2']
for item in cat_cols:
df[item]=df[item].astype(str)
df[cat_cols].head()
# Label encoding to convert string to representative numeric values
df[cat_cols]= df[cat_cols].apply(LabelEncoder().fit_transform)
# Display top 5 records from the dataset to check if all the records are converted to numbers
df.head(5)
```
## Feature Selection
VarianceThreshold is a simple baseline approach to feature selection. It removes all features whose variance doesn’t meet some threshold. By default, it removes all zero-variance features, i.e. features that have the same value in all samples.
threshold .8 * (1 - .8)
Using 0.8 as a threshhold, we will remove features with less than 20 percent variation within itself.
```
df_X=df.loc[:, df.columns != 'price']
df_y=df['price']
from sklearn.feature_selection import VarianceThreshold
print('%s Number of features before VarianceThreshhold'%len(df_X.columns))
selector=VarianceThreshold(threshold=(.8*(1-.8)))
FeaturesTransformed=selector.fit_transform(df_X)
## print the support and shape of the transformed features
print(selector.get_support())
data=df_X[df_X.columns[selector.get_support(indices=True)]]
cols=data.columns
df_reduced=pd.DataFrame(FeaturesTransformed, columns=cols)
df_reduced.shape
data=df_reduced
target=df_y
data.shape
```
# Yellowbrick for Feature Selection
we are using yellowbrick's feature selection method for finding and selecting the most useful features and eliminate zero importance features from the dataset.
### Important Features for Random Forest Regressor
```
#Using yellowbrick feature selection method with random forest regressor
from sklearn.ensemble import RandomForestRegressor
from yellowbrick.features.importances import FeatureImportances
fig = plt.figure(figsize=(10,10))
ax = fig.add_subplot()
viz = FeatureImportances(RandomForestRegressor(), ax=ax)
viz.fit(data, target)
viz.poof()
feature_importances = pd.DataFrame(viz.feature_importances_,
index=data.columns,
columns=['importance']).sort_values('importance', ascending=False)
## important features for Random Forest Regression
RF_importants=feature_importances.index[feature_importances.importance!=0]
df[RF_importants].shape
# predicting price using random forest regressor
from sklearn.ensemble import RandomForestRegressor
from sklearn.model_selection import train_test_split as split
X=df[RF_importants]
Y=df['price']
model=RandomForestRegressor()
X_train, X_test, Y_train, Y_test= split(X,Y,test_size=0.25, random_state=42)
model.fit(X_train,Y_train)
Y_test=model.predict(X_test)
ax1=sns.distplot(target,hist=False, color='r',label="Actual price")
sns.distplot(Y_test,hist=False,color='b', label="Predicted price", ax=ax1)
plt.title(" Actual Vs Predicted Price ")
plt.xlabel('Price')
plt.ylabel('Proportion of Books')
plt.show()
```
### Important Features for Gradient Boosting Regressor
```
from sklearn.ensemble import GradientBoostingRegressor
from yellowbrick.features.importances import FeatureImportances
fig = plt.figure(figsize=(20,20))
ax = fig.add_subplot()
viz = FeatureImportances(GradientBoostingRegressor(), ax=ax)
viz.fit(data, target)
viz.poof()
feature_importances = pd.DataFrame(viz.feature_importances_,
index=data.columns,
columns=['importance']).sort_values('importance', ascending=False)
## important features for gradient boosting regression
GBR_importants=feature_importances.index[feature_importances.importance!=0]
df[GBR_importants].shape
from sklearn.ensemble import GradientBoostingRegressor
from sklearn.model_selection import train_test_split as split
X=df[GBR_importants]
Y=df['price']
model=GradientBoostingRegressor()
X_train, X_test, Y_train, Y_test= split(X,Y,test_size=0.25, random_state=42)
model.fit(X_train,Y_train)
Y_test=model.predict(X_test)
ax1=sns.distplot(target,hist=False, color='r',label="Actual price")
sns.distplot(Y_test,hist=False,color='b', label="Predicted price", ax=ax1)
plt.title(" Actual Vs Predicted Price ")
plt.xlabel('Price')
plt.ylabel('Proportion of Books')
plt.show()
```
### Important Features for Decision Tree Regressor
```
from sklearn.tree import DecisionTreeRegressor
from yellowbrick.features.importances import FeatureImportances
fig = plt.figure(figsize=(20,20))
ax = fig.add_subplot()
viz = FeatureImportances( DecisionTreeRegressor(), ax=ax)
viz.fit(data, target)
viz.poof()
feature_importances = pd.DataFrame(viz.feature_importances_,
index=data.columns,
columns=['importance']).sort_values('importance', ascending=False)
## important features for decision tree regression
DTR_importants=feature_importances.index[feature_importances.importance!=0]
df[DTR_importants].shape
from sklearn.tree import DecisionTreeRegressor
from sklearn.model_selection import train_test_split as split
X=df[DTR_importants]
Y=df['price']
model=DecisionTreeRegressor()
X_train, X_test, Y_train, Y_test= split(X,Y,test_size=0.25, random_state=42)
model.fit(X_train,Y_train)
Y_test=model.predict(X_test)
ax1=sns.distplot(target,hist=False, color='r',label="Actual price")
sns.distplot(Y_test,hist=False,color='b', label="Predicted price", ax=ax1)
plt.title(" Actual Vs Predicted Price ")
plt.xlabel('Price')
plt.ylabel('Proportion of Books')
plt.show()
```
## Model Development
In this section we will implement several models that will predict price using the dependent variables and compare the accuracy, r-score, goodness of fit and plot residuals. Based on the scores and visual comparison of the plots, we will refine the best performing models using grid search to fine tune the hyperparameters to generate a better predictive model.
```
# This function applies multiple models on the data and returns model name with r2-score and mean squared error value
def ModelScores(data,target):
X=data
Y=target
from sklearn.metrics import r2_score
from sklearn.metrics import mean_squared_error
import math
from sklearn.model_selection import train_test_split as split
X_train, X_test, Y_train, Y_test= split(X,Y,test_size=0.25, random_state=42)
from sklearn.ensemble import GradientBoostingRegressor
from sklearn.ensemble import RandomForestRegressor
from sklearn.tree import DecisionTreeRegressor
from sklearn.linear_model import LinearRegression
from sklearn.neural_network import MLPRegressor
from sklearn.linear_model import RidgeCV
from sklearn.linear_model import LassoLars
from sklearn.linear_model import Lasso
from sklearn.linear_model import ElasticNet
from sklearn.linear_model import BayesianRidge
from sklearn.linear_model import RANSACRegressor
models={'Gradient Boost': GradientBoostingRegressor(),
'Random Forest': RandomForestRegressor(),
'Decision Tree': DecisionTreeRegressor(),
'Linear Regression': LinearRegression(),
'MLP': MLPRegressor(),
'Ridge CV': RidgeCV(),
'LassoLars':LassoLars(),
'Lasso':Lasso(),
'Elastic Search': ElasticNet(),
'Bayesian Ridge':BayesianRidge(),
'Ransac':RANSACRegressor()
}
for name,model in models.items():
mdl=model
mdl.fit(X_train, Y_train)
prediction = mdl.predict(X_test)
print(name)
print("Accuracy Score", r2_score(Y_test, prediction))
mse3 = mean_squared_error(Y_test, prediction)
print("The root mean square value", math.sqrt(mse3))
data=data
target=df['price']
ModelScores(data, target)
%matplotlib inline
from yellowbrick.classifier import ClassificationReport
from yellowbrick.classifier import ClassPredictionError
from yellowbrick.regressor import ResidualsPlot
from sklearn.svm import SVR
from sklearn.ensemble import RandomForestRegressor
from sklearn.ensemble import GradientBoostingRegressor
from sklearn.tree import DecisionTreeRegressor
regressors = {
"Gradient Boost": GradientBoostingRegressor(),
"Random Forest": RandomForestRegressor(),
"Decision Tree": DecisionTreeRegressor()
}
for _, regressor in regressors.items():
visualizer = ResidualsPlot(regressor)
visualizer.fit(X_train, Y_train)
visualizer.score(X_test, Y_test)
visualizer.poof()
from yellowbrick.target import FeatureCorrelation
feature_names = np.array(df.columns)
data=df.loc[:, df.columns != 'price']
target=df['price']
figsize=(20, 20)
visualizer = FeatureCorrelation(labels=feature_names)
visualizer.fit(data, target)
visualizer.poof()
#validation curve for decision tree regression and Random forest regression models
from yellowbrick.model_selection import ValidationCurve
# Extract the instances and target
X = df_X
y = df_y
regressors = {
"Gradient Boost": GradientBoostingRegressor(),
"Decision Tree": DecisionTreeRegressor(),
"Random Forest": RandomForestRegressor()
}
for _, regressor in regressors.items():
viz = ValidationCurve(
regressor, param_name="max_depth",
param_range=np.arange(1, 11), cv=10, scoring="r2"
)
# Fit and poof the visualizer
viz.fit(X, y)
viz.poof()
```
We will use the validation curve from the above three figures to narrow down the optimal 'max_depth' value range to use, for hyperparameter tuning in a grid search.
```
# Cross Validation Score for Random Forest Regressor
from sklearn.ensemble import RandomForestRegressor
from sklearn.model_selection import KFold
from yellowbrick.model_selection import CVScores
ind=df[RF_importants].values
dep=df['price'].values
_, ax = plt.subplots()
cv = KFold(10)
oz = CVScores(
RandomForestRegressor(), ax=ax, cv=cv, scoring='r2'
)
oz.fit(ind, dep)
oz.poof()
# CV score for Gradiet Boosting Regresor
from sklearn.ensemble import GradientBoostingRegressor
from sklearn.model_selection import StratifiedKFold
from yellowbrick.model_selection import CVScores
ind=df[GBR_importants].values
dep=df['price'].values
_, ax = plt.subplots()
cv = StratifiedKFold(n_splits=10, random_state=42)
oz = CVScores(
GradientBoostingRegressor(), ax=ax, cv=cv, scoring = 'r2'
)
oz.fit(ind, dep)
oz.poof()
# CV score for Decision Tree Regressor
from sklearn.tree import DecisionTreeRegressor
from sklearn.model_selection import KFold
from yellowbrick.model_selection import CVScores
ind=df[DTR_importants].values
dep=df['price'].values
_, ax = plt.subplots()
cv = KFold(10)
oz = CVScores(
DecisionTreeRegressor(), ax=ax, cv=cv, scoring = 'r2'
)
oz.fit(ind, dep)
oz.poof()
```
## Hyperparameter Tuning
```
# hyper-parameter tunung for decision tree
from sklearn.model_selection import GridSearchCV
from sklearn.tree import DecisionTreeRegressor
DecisionTree = DecisionTreeRegressor(random_state = 40)
min_samples_split = [2,3,4,5,6,7,8]
min_samples_leaf = [1,2,3,4,5]
max_depth = [4,5,6,7,8,9]
tuned_params = [{'min_samples_split': min_samples_split}, {'min_samples_leaf': min_samples_leaf},{'max_depth': max_depth}]
n_folds = 5
X=df[DTR_importants]
Y=df['price']
grid = GridSearchCV(
DecisionTree, tuned_params, cv=n_folds
)
grid.fit(X, Y)
print(grid.best_estimator_)
from sklearn.model_selection import GridSearchCV
from sklearn.ensemble import GradientBoostingRegressor
GradientBoosting = GradientBoostingRegressor(random_state = 40)
alphas = [0.001, 0.01, 0.1, 0.5, 0.9]
sample_split = [2,3,4,5,6,7,8]
max_depth = [4,5,6,7,8,9]
learning_rate = [0.1, 0.3, 0.5, 0.7]
tuned_params = [{'alpha': alphas}, {'min_samples_split': sample_split}, {'max_depth': max_depth}, {'learning_rate':learning_rate}]
n_folds = 5
X=df[GBR_importants]
Y=df['price']
grid = GridSearchCV(
GradientBoosting, tuned_params, cv=n_folds
)
grid.fit(X, Y)
print(grid.best_estimator_)
from sklearn.model_selection import GridSearchCV
from sklearn.ensemble import RandomForestRegressor
RandomForest = RandomForestRegressor(random_state = 40)
estimators = [10,50,100]
sample_split = [2,3,4,5,6,7,8]
sample_leaf = [1,2,3,4,5]
max_depth = [4,5,6,7,8,9]
tuned_params = [{'n_estimators': estimators}, {'min_samples_split': sample_split}, {'min_samples_leaf': sample_leaf},{'max_leaf_nodes': max_depth}]
n_folds = 5
X=df[RF_importants]
Y=df['price']
grid = GridSearchCV(
RandomForest, tuned_params, cv=n_folds
)
grid.fit(X, Y)
print(grid.best_estimator_)
```
## Model Evaluation
In this part we are using the result that we obtained from the grid search as an input to retrain our models. The grid search is applied with cross validation by taking the average score over 5 folds.
```
X=df[DTR_importants]
Y=df['price']
X_train, X_test, Y_train, Y_test= split(X,Y,test_size=0.25, random_state=42)
from sklearn.tree import DecisionTreeRegressor
model=DecisionTreeRegressor(criterion='mse', max_depth=9, max_features=None,
max_leaf_nodes=None, min_impurity_decrease=0.0,
min_impurity_split=None, min_samples_leaf=1,
min_samples_split=2, min_weight_fraction_leaf=0.0,
presort=False, random_state=40, splitter='best')
model.fit(X_train,Y_train)
prediction13 = model.predict(X_test)
print("Accuracy Score", r2_score(Y_test, prediction13))
mse = mean_squared_error(Y_test, prediction13)
print("The root mean square value", math.sqrt(mse))
X=df[GBR_importants]
Y=df['price']
X_train, X_test, Y_train, Y_test= split(X,Y,test_size=0.25, random_state=42)
from sklearn.ensemble import GradientBoostingRegressor
model=GradientBoostingRegressor(alpha=0.9, criterion='friedman_mse', init=None,
learning_rate=0.1, loss='ls', max_depth=5, max_features=None,
max_leaf_nodes=None, min_impurity_decrease=0.0,
min_impurity_split=None, min_samples_leaf=1,
min_samples_split=2, min_weight_fraction_leaf=0.0,
n_estimators=100, n_iter_no_change=None, presort='auto',
random_state=40, subsample=1.0, tol=0.0001,
validation_fraction=0.1, verbose=0, warm_start=False)
model.fit(X_train,Y_train)
prediction13 = model.predict(X_test)
print("Accuracy Score", r2_score(Y_test, prediction13))
mse = mean_squared_error(Y_test, prediction13)
print("The root mean square value", math.sqrt(mse))
X=df[RF_importants]
Y=df['price']
X_train, X_test, Y_train, Y_test= split(X,Y,test_size=0.25, random_state=42)
from sklearn.ensemble import RandomForestRegressor
rfg = RandomForestRegressor(bootstrap=True, criterion='mse', max_depth=None,
max_features='auto', max_leaf_nodes=None,
min_impurity_decrease=0.0, min_impurity_split=None,
min_samples_leaf=1, min_samples_split=2,
min_weight_fraction_leaf=0.0, n_estimators=100, n_jobs=None,
oob_score=False, random_state=40, verbose=0, warm_start=False)
rfg.fit(X_train, Y_train)
prediction14 = rfg.predict(X_test)
print("Accuracy Score", r2_score(Y_test, prediction14))
mse2 = mean_squared_error(Y_test, prediction14)
print("The root mean square value", math.sqrt(mse2))
```
http://sphweb.bumc.bu.edu/otlt/mph-modules/bs/bs704_summarizingdata/bs704_summarizingdata7.html
| github_jupyter |
<img src="images/dask_horizontal.svg" align="right" width="30%">
# Introduction
Welcome to the Dask Tutorial.
Dask is a parallel computing library that scales the existing Python ecosystem. This tutorial will introduce Dask and parallel data analysis more generally.
Dask can scale down to your laptop laptop and up to a cluster. Here, we'll use an environment you setup on your laptop to analyze medium sized datasets in parallel locally.
# Overview
Dask provides multi-core and distributed parallel execution on larger-than-memory datasets.
We can think of Dask at a high and a low level
* **High level collections:** Dask provides high-level Array, Bag, and DataFrame
collections that mimic NumPy, lists, and Pandas but can operate in parallel on
datasets that don't fit into memory. Dask's high-level collections are
alternatives to NumPy and Pandas for large datasets.
* **Low Level schedulers:** Dask provides dynamic task schedulers that
execute task graphs in parallel. These execution engines power the
high-level collections mentioned above but can also power custom,
user-defined workloads. These schedulers are low-latency (around 1ms) and
work hard to run computations in a small memory footprint. Dask's
schedulers are an alternative to direct use of `threading` or
`multiprocessing` libraries in complex cases or other task scheduling
systems like `Luigi` or `IPython parallel`.
Different users operate at different levels but it is useful to understand
both.
The Dask [use cases](http://dask.pydata.org/en/latest/use-cases.html) provides a number of sample workflows where Dask should be a good fit.
## Prepare
You should clone this repository
git clone http://github.com/dask/dask-tutorial
The included file `environment.yml` contains a list of all of the packages needed to run this tutorial. To install them using `conda`, you can do
conda env create environment.yml
conda activate dask-tutorial
Do this *before* running this notebook
Finally, run the following script to download and create data for analysis.
```
# in directory dask-tutorial/
# this takes a little while
%run prep.py
```
## Links
* Reference
* [Documentation](https://dask.pydata.org/en/latest/)
* [Code](https://github.com/dask/dask/)
* [Blog](http://matthewrocklin.com/blog/)
* Ask for help
* [`dask`](http://stackoverflow.com/questions/tagged/dask) tag on Stack Overflow
* [github issues](https://github.com/dask/dask/issues/new) for bug reports and feature requests
* [gitter](https://gitter.im/dask/dask) for quasi-realtime conversation
## Tutorial Structure
Each section is a Jupyter notebook. There's a mixture of text, code, and exercises.
If you haven't used Jupyterlab, it's similar to the Jupyter Notebook. If you haven't used the Notebook, the quick intro is
1. There are two modes: command and edit
2. From command mode, press `Enter` to edit a cell (like this markdown cell)
3. From edit mode, press `Esc` to change to command mode
4. Press `shift+enter` to execute a cell and move to the next cell.
The toolbar has commands for executing, converting, and creating cells.
The layout of the tutorial will be as follows:
- Foundations: an explanation of what Dask is, how it works, and how to use lower-level primitives to set up computations. Casual users may wish to skip this section, although we consider it useful knowledge for all users.
- Distributed: information on running Dask on the distributed scheduler, which enables scale-up to distributed settings and enhanced monitoring of task operations. The distributed scheduler is now generally the recommended engine for executing task work, even on single workstations or laptops.
- Collections: convenient abstractions giving a familiar feel to big data
- bag: Python iterators with a functional paradigm, such as found in func/iter-tools and toolz - generalize lists/generators to big data; this will seem very familiar to users of PySpark's [RDD](http://spark.apache.org/docs/2.1.0/api/python/pyspark.html#pyspark.RDD)
- array: massive multi-dimensional numerical data, with Numpy functionality
- dataframes: massive tabular data, with Pandas functionality
Whereas there is a wealth of information in the documentation, linked above, here we aim to give practical advice to aid your understanding and application of Dask in everyday situations. This means that you should not expect every feature of Dask to be covered, but the examples hopefully are similar to the kinds of work-flows that you have in mind.
## Exercise: Print `Hello, world!`
Each notebook will have exercises for you to solve. You'll be given a blank or partially completed cell, followed by a "magic" cell that will load the solution. For example
Print the text "Hello, world!".
```
# Your code here
%load solutions/00-hello-world.py
```
The above cell needs to be executed twice, once to load the solution and once to run it.
| github_jupyter |
```
from loop import TrainingLoop
import os
import numpy as np
import tensorflow as tf
# These lines will make the gpu not give errors.
gpus= tf.config.experimental.list_physical_devices('GPU')
tf.config.experimental.set_memory_growth(gpus[0], True)
from tensorflow import keras
from tensorflow.keras import layers
from tensorflow.keras.models import Sequential
# Set random seed so the comparison of different solutions won't be affected by it.
tf.random.set_seed(42)
np.random.seed(42)
@tf.function
def calc_loss(x_train, y_train):
with tf.GradientTape() as tape:
logits = model(x_train, training=False)
loss_value = keras.losses.CategoricalCrossentropy(from_logits=True)(y_train, logits)
return loss_value
def batch_selector(data, idx):
largest_loss = 0
largest_loss_idx = idx
if idx < len(data) - length:
for i in range(idx, idx+length):
x_batch_train = data[i][0]
y_batch_train = data[i][1]
loss = calc_loss(x_batch_train, y_batch_train)
if loss > largest_loss:
largest_loss = loss
largest_loss_idx = i
return largest_loss_idx
else:
loss = calc_loss(data[idx][0], data[idx][1])
return idx
length = 75
log_path = 'logs/original/mnist.csv'
# Function to load dataset from file. This is needed so we can easily load the two datasets without copy pasteing.
def load_data( name ):
X_train = np.load(os.path.join('data', name, name + '_train_vectors.npy'))
X_test = np.load(os.path.join('data', name, name + '_test_vectors.npy'))
Y_train = np.load(os.path.join('data', name, name + '_train_labels.npy'))
Y_test = np.load(os.path.join('data', name, name + '_test_labels.npy'))
# The images need to have shape (28, 28, 1), we didn't take care of this in preprocessing.
X_train = np.expand_dims(X_train, -1)
X_test = np.expand_dims(X_test, -1)
return X_train, Y_train, X_test, Y_test
# The same model is used for both datasets so it is more convenient to make them in a funtion.
def make_model(X_train, Y_train):
# This is a simple convolutional neural network. It isn't the best possible network for MNIST
# but the point here is to test how much batch selection methods will speed up a CNN, not the CNN itself.
model = Sequential()
model.add(layers.Input(shape = (28, 28, 1,)))
model.add(layers.Conv2D(64, kernel_size = (3, 3), activation = "relu"))
model.add(layers.MaxPooling2D( pool_size = (2, 2)))
model.add(layers.Conv2D(64, kernel_size = (3, 3), activation = "relu"))
model.add(layers.MaxPooling2D(pool_size = (2, 2)))
model.add(layers.Flatten())
model.add(layers.Dense(10, activation = "softmax"))
# Put the model in our custom training loop.
training = TrainingLoop(
model = model,
X = X_train,
y = Y_train,
optimizer = keras.optimizers.Adam(),
loss_function = keras.losses.CategoricalCrossentropy(from_logits=True),
batch_size = 64,
train_metrics = tf.keras.metrics.CategoricalAccuracy(),
val_metrics = tf.keras.metrics.CategoricalAccuracy(),
validation_split = 0.2,
batch_selection = batch_selector,
length=length
)
# We still have to compile the model for the test evaluation.
model.compile(loss = "categorical_crossentropy", optimizer = "adam", metrics=["accuracy"], log_file=log_path)
return model, training
# Load and train the MNIST dataset.
X_train, Y_train, X_test, Y_test = load_data( "mnist" )
model, training = make_model( X_train, Y_train )
training.train(epochs = 20)
# Evaluate the MNIST model.
model.evaluate( X_test, Y_test )
# Load and train the Fashion-MNIST dataset.
X_train_f, Y_train_f, X_test_f, Y_test_f = load_data( "fashion_mnist" )
model_f, training_f = make_model( X_train_f, Y_train_f )
training_f.train(epochs = 20)
# Evaluate the Fashion-MNIST dataset.
model_f.evaluate( X_test_f, Y_test_f )
```
| github_jupyter |
## Loading the data
First we set up our imports:
```
import numpy as np
import yt
```
First we load the data set, specifying both the unit length/mass/velocity, as well as the size of the bounding box (which should encapsulate all the particles in the data set)
At the end, we flatten the data into "ad" in case we want access to the raw simulation data
>This dataset is available for download at https://yt-project.org/data/GadgetDiskGalaxy.tar.gz (430 MB).
```
fname = "GadgetDiskGalaxy/snapshot_200.hdf5"
unit_base = {
"UnitLength_in_cm": 3.08568e21,
"UnitMass_in_g": 1.989e43,
"UnitVelocity_in_cm_per_s": 100000,
}
bbox_lim = 1e5 # kpc
bbox = [[-bbox_lim, bbox_lim], [-bbox_lim, bbox_lim], [-bbox_lim, bbox_lim]]
ds = yt.load(fname, unit_base=unit_base, bounding_box=bbox)
ds.index
ad = ds.all_data()
```
Let's make a projection plot to look at the entire volume
```
px = yt.ProjectionPlot(ds, "x", ("gas", "density"))
px.show()
```
Let's print some quantities about the domain, as well as the physical properties of the simulation
```
print("left edge: ", ds.domain_left_edge)
print("right edge: ", ds.domain_right_edge)
print("center: ", ds.domain_center)
```
We can also see the fields that are available to query in the dataset
```
sorted(ds.field_list)
```
Let's create a data object that represents the full simulation domain, and find the total mass in gas and dark matter particles contained in it:
```
ad = ds.all_data()
# total_mass returns a list, representing the total gas and dark matter + stellar mass, respectively
print([tm.in_units("Msun") for tm in ad.quantities.total_mass()])
```
Now let's say we want to zoom in on the box (since clearly the bounding we chose initially is much larger than the volume containing the gas particles!), and center on wherever the highest gas density peak is. First, let's find this peak:
```
density = ad["PartType0", "density"]
wdens = np.where(density == np.max(density))
coordinates = ad["PartType0", "Coordinates"]
center = coordinates[wdens][0]
print("center = ", center)
```
Set up the box to zoom into
```
new_box_size = ds.quan(250, "code_length")
left_edge = center - new_box_size / 2
right_edge = center + new_box_size / 2
print(new_box_size.in_units("Mpc"))
print(left_edge.in_units("Mpc"))
print(right_edge.in_units("Mpc"))
ad2 = ds.region(center=center, left_edge=left_edge, right_edge=right_edge)
```
Using this new data object, let's confirm that we're only looking at a subset of the domain by first calculating the total mass in gas and particles contained in the subvolume:
```
print([tm.in_units("Msun") for tm in ad.quantities.total_mass()])
```
And then by visualizing what the new zoomed region looks like
```
px = yt.ProjectionPlot(ds, "x", ("gas", "density"), center=center, width=new_box_size)
px.show()
```
Cool - there's a disk galaxy there!
| github_jupyter |
<h1>Table of Contents<span class="tocSkip"></span></h1>
<div class="toc"><ul class="toc-item"><li><span><a href="#Before-running-the-notebook" data-toc-modified-id="Before-running-the-notebook-1"><span class="toc-item-num">1 </span>Before running the notebook</a></span></li><li><span><a href="#Generation" data-toc-modified-id="Generation-2"><span class="toc-item-num">2 </span>Generation</a></span></li><li><span><a href="#XOR" data-toc-modified-id="XOR-3"><span class="toc-item-num">3 </span>XOR</a></span></li><li><span><a href="#Visualize-results" data-toc-modified-id="Visualize-results-4"><span class="toc-item-num">4 </span>Visualize results</a></span></li><li><span><a href="#MT" data-toc-modified-id="MT-5"><span class="toc-item-num">5 </span>MT</a></span></li><li><span><a href="#SR" data-toc-modified-id="SR-6"><span class="toc-item-num">6 </span>SR</a></span></li></ul></div>
```
%load_ext autoreload
%autoreload 2
import os
import sys
import warnings
import numpy as np
import pandas as pd
import scipy.special as ss
import matplotlib.pyplot as plt
from sklearn.metrics import accuracy_score
from scipy.stats import pearsonr
module_path = os.path.abspath(os.path.join('./modules/'))
scripts_path = os.path.abspath(os.path.join('./scripts/'))
sys.path.append(module_path)
sys.path.append(scripts_path)
np.seterr(all='ignore')
ss.seterr(all='ignore')
warnings.simplefilter(action='ignore', category=FutureWarning)
from main import main as infer
from generate_graph import main as generate
from compute_metrics import membership_metrics, normalize_nonzero_membership, calculate_AUC
cmap = 'PuBuGn'
def extract_overlapping_membership(i, cm, U, threshold=0.1):
groups = np.where(U[i] > threshold)[0]
wedge_sizes = U[i][groups]
wedge_colors = [cm(c) for c in groups]
return wedge_sizes, wedge_colors
def normalize_nonzero_membership(U):
den1 = U.sum(axis=1, keepdims=True)
nzz = den1 == 0.
den1[nzz] = 1.
return U / den1
```
# Before running the notebook
- Modify the _generation_ setting file inserting the right output path, e.g. <code>folder: "../data/input/"</code>;
- Modify the _generation_ setting file inserting the right output path, e.g. <code>label: "test"</code>;
- Modify the _inference_ setting file inserting the right output path, e.g. <code>folder: "../data/output/XOR/"</code>;
- Modify the _inference_ setting file choosing the right flag value for outputting results, e.g. <code>output_inference: True</code>;
- Modify the _cells_ defining the <code>args</code> dictionary inserting the desired input for the <code>generate</code> and <code>infer</code> functions.
# Generation
Graphs realizations are drawn from the specified distribution; here self loops are allowed.
```
gen_args = {
'min_ratio': 0.8, # bottom value for the range of interest for mu
'max_ratio': np.nan, # top value for the range of interest for mu
'step_ratio': np.nan, # stepsize value for the range of interest for mu
'samples': 1, # number of samples to be drawn for each set of params
'settings': './setting_syn_XOR.yaml' # path to the setting file to be used
}
generate(gen_args)
in_folder = '../data/input'
theta_GT = np.load(f'{in_folder}/results_test_r0.8_XOR_112.npz')
```
# XOR
Data are loaded and preprocessed (e.g. self loops are removed); the XOR inference algorithm is run.
```
inf_args = {
'adj' : 'syn_test_r0.8_XOR_112.dat', # name of the network file
'in_folder' : '../data/input/', # folder of the input network
'algorithm' : 'XOR', # the desired inference model
'settings' : 'setting_XOR.yaml', # complete path of the settings file
'K' : 3, # number of communities to infer
'beta' : 5, # inverse temperature parameter
'NFold' : 5, # number of folds to perform cross-validation
'out_mask' : False # flag to output the masks
}
# The model output function appends results to existing dataframe if it is found in memory.
# Delete if you want to avoid repetitions of lines or drop duplicates when loading.
! rm -r ../data/output/XOR
infer(inf_args)
folder = '../data/output/XOR'
metrics = pd.read_csv(f'{folder}/metrics_test_r0.8_XOR_112_XOR.csv')
conv = metrics.convergence_out == True
best_seed = metrics[metrics[conv].logL_out == np.max(metrics[conv].logL_out)].seed_out.item()
theta = np.load(f'{folder}/parameters_test_r0.8_XOR_112_XOR_{str(best_seed)}.npz')
list(theta.keys())
print('Inferred % of nodes interacting with hierarchy:',theta['ratio'])
print('Ground truth % of nodes interacting with hierarchy:',theta_GT['sigma'].sum()/theta_GT['sigma'].shape[0])
```
# Visualize results
### Ground truth $\sigma$
```
theta_GT['sigma']
```
### Inferred $\sigma$
```
theta['Q'].astype('int')
```
### Ranking
```
plt.figure()
plt.scatter(theta_GT['s'][theta_GT['sigma']==1],theta['s'][theta_GT['sigma']==1],s=100,c='royalblue',edgecolors='black')
plt.ylabel(r'$s_{inferred}$',fontsize=20)
plt.xlabel(r'$s_{GT}$',fontsize=20)
plt.show()
```
### Communities
#### Extract networks (for visualization)
```
import networkx as nx
from tools import import_data
from matplotlib.colors import LinearSegmentedColormap
from matplotlib.colors import ListedColormap
from matplotlib.ticker import MaxNLocator
from compute_metrics import opt_permutation
G = import_data('../../data/input/syn_test_r0.8_XOR_112.dat', header=0)[0][0]
Gc = max(nx.weakly_connected_components(G), key=len)
nodes_to_remove = set(G.nodes()).difference(Gc)
G.remove_nodes_from(list(nodes_to_remove))
```
#### Plot network
```
from_list = LinearSegmentedColormap.from_list
cm = from_list('Set15', plt.cm.Set1(range(0,10)), 10)
cmap = cm
wedgeprops = {'edgecolor' : 'lightgrey'}
u_gt = normalize_nonzero_membership(theta_GT['u'])
u_inf = normalize_nonzero_membership(theta['u'])
N = u_gt.shape[0]
# Add one column for a dummy membership, this is for nodes with sigma_i=1
u_gt = np.hstack((u_gt,np.zeros((N,1))))
u_inf = np.hstack((u_inf,np.zeros((N,1))))
u_gt[np.where(theta_GT['sigma']==1)[0]] = 0
u_gt[np.where(theta_GT['sigma']==1)[0],-1] = 1
u_inf[np.where(theta['Q'][0]==1)[0],-1] = 1
# Permute so that groups GT correspond to inferred
u_inf = np.dot(u_inf, opt_permutation(u_inf, u_gt)) # permute on clusters (2nd dimension)
# Custom position, for better visualization
delta = 0.6
C = theta['u'].shape[1] +1
pos_groups = nx.circular_layout(nx.cycle_graph(C))
pos = {}
nodes = list(G.nodes())
for c, i in enumerate(nodes):
r = np.random.rand() * 2 * np.math.pi
radius = np.random.rand()
pos[i] = pos_groups[np.argmax(u_gt[c])] + delta * radius * np.array([np.math.cos(r), np.math.sin(r)])
rad = 0.09
plt.figure(figsize=(12,6))
plt.subplot(1, 2, 1)
ax = plt.gca()
nx.draw_networkx_edges(G, pos,arrows=False, edge_color='lightgrey', alpha=0.5)
for n in list(G.nodes()):
wedge_sizes, wedge_colors = extract_overlapping_membership(n, cm, u_gt, threshold=0.1)
if len(wedge_sizes) > 0:
pie,t = plt.pie(wedge_sizes, center=pos[n], colors=wedge_colors,
radius=rad, wedgeprops=wedgeprops, normalize=False)
ax.axis("equal")
plt.title('Ground truth')
plt.subplot(1, 2, 2)
ax = plt.gca()
nx.draw_networkx_edges(G, pos, arrows=False, edge_color='lightgrey', alpha=0.5)
for n in list(G.nodes()):
wedge_sizes, wedge_colors = extract_overlapping_membership(n, cm, u_inf, threshold=0.1)
if len(wedge_sizes) > 0:
pie,t = plt.pie(wedge_sizes, center=pos[n], colors=wedge_colors,
radius=rad, wedgeprops=wedgeprops, normalize=False)
ax.axis("equal")
plt.title('Inferred')
plt.show()
```
# MT
```
inf_args = {
'adj' : 'syn_test_r0.8_XOR_112.dat', # name of the network file
'in_folder' : '../data/input/', # folder of the input network
'algorithm' : 'MT', # the desired inference model
'settings' : 'setting_MT.yaml', # complete path of the settings file
'K' : 3, # number of communities to infer
'beta' : 5, # inverse temperature parameter
'NFold' : 5, # number of folds to perform cross-validation
'out_mask' : False # flag to output the masks
}
# The model output function appends results to existing dataframe if it is found in memory.
# Delete if you want to avoid repetitions of lines or drop duplicates when loading.
! rm -r ../data/output/MT
infer(inf_args)
folder = '../data/output/MT'
metrics = pd.read_csv(f'{folder}/metrics_test_r0.8_XOR_112_MT.csv')
conv = metrics.convergence_out == True
best_seed = metrics[metrics[conv].logL_out == np.max(metrics[conv].logL_out)].seed_out.item()
theta_MT = np.load(f'{folder}/parameters_test_r0.8_XOR_112_MT_{str(best_seed)}.npz')
list(theta.keys())
```
# SR
```
inf_args = {
'adj' : 'syn_test_r0.8_XOR_112.dat', # name of the network file
'in_folder' : '../data/input/', # folder of the input network
'algorithm' : 'SR', # the desired inference model
'settings' : 'setting_SR.yaml', # complete path of the settings file
'K' : 0, # number of communities to infer NOT USED IN THIS CASE
'beta' : 5, # inverse temperature parameter NOT USED IN THIS CASE
'NFold' : 5, # number of folds to perform cross-validation
'out_mask' : False # flag to output the masks
}
# The model output function appends results to existing dataframe if it is found in memory.
# Delete if you want to avoid repetitions of lines or drop duplicates when loading.
! rm -r ../data/output/SR
infer(inf_args)
folder = '../data/output/SR'
theta_SR = np.load(f'{folder}/parameters_test_r0.8_XOR_112_SR.npz')
list(theta_SR.keys())
```
| github_jupyter |
#Section 1
---
```
import numpy as np
import matplotlib.pyplot as plt
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Conv2D , Activation, Flatten , Dense, MaxPooling2D
from tensorflow.keras.layers import Dropout
from keras.layers.normalization import BatchNormalization
from keras.datasets import mnist
from keras.utils import np_utils, to_categorical
from sklearn.model_selection import KFold
def load_data():
return mnist.load_data()
def prepare_data(train_images, train_labels, test_images, test_labels):
x_train = train_images.reshape(60000,28,28,1)
x_test = test_images.reshape(10000,28,28,1)
#one hot encoding
y_train = to_categorical(train_labels)
y_test = to_categorical(test_labels)
return (x_train, y_train, x_test, y_test )
def model(x_train, y_train, x_test, y_test ):
kfold = KFold(n_splits=3, shuffle=True)
fold_no = 0
history = ''
# Define per-fold score containers
for train_index, test_index in kfold.split(x_train):
print('------------------------------------------------------------------------')
print(f'Training for fold {fold_no} ...')
model = Sequential()
model.add(Conv2D(32, kernel_size=(3, 3), activation='relu', padding='valid', input_shape=(28,28,1)))
model.add(BatchNormalization())
model.add(MaxPooling2D(pool_size=(2,2)))
model.add(Conv2D(32, kernel_size=(3, 3), activation='relu', padding='valid'))
model.add(BatchNormalization())
model.add(MaxPooling2D(pool_size=(2,2)))
model.add(Flatten())
model.add(Dense(64, activation='relu'))
model.add(Dense(10, activation='softmax'))
model.compile(optimizer='adam', loss='categorical_crossentropy', metrics=['accuracy'])
history = model.fit(x_train[train_index], y_train[train_index],
batch_size=2048, epochs=10, verbose=2,
validation_data=(x_train[test_index], y_train[test_index]))
scores = model.evaluate(x_train[test_index], y_train[test_index], verbose=0)
print(f'Score for fold {fold_no}: {model.metrics_names[0]} of {scores[0]}; {model.metrics_names[1]} of {scores[1]*100}%')
fold_no += 1
model.summary()
score = model.evaluate(x_test, y_test)
print('Test loss :', score[0])
print('Test accuracy:', score[1] * 100)
return history
def plot(history):
plt.plot(history.history['accuracy'])
plt.plot(history.history['val_accuracy'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
# summarize history for loss
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
def main():
(train_images, train_labels), (test_images, test_labels) = load_data()
x_train, y_train, x_test, y_test = prepare_data(train_images, train_labels, test_images, test_labels)
history = model(x_train, y_train, x_test, y_test)
plot(history)
main()
```
#noisy
```
def show_result(x_test_noisy, test_images):
n = 10
for i in range(n):
# Display original
ax = plt.subplot(2, n, i + 1)
plt.imshow(test_images[i].reshape(28, 28))
plt.gray()
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
# Display noisy
ax = plt.subplot(2, n, i + 1)
plt.imshow(x_test_noisy[i].reshape(28, 28))
plt.gray()
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
plt.show()
def add_noise(x_train, x_test, noise_factor_train, noise_factor_test):
x_train_noisy = x_train + noise_factor_train * np.random.normal(loc=0.0, scale=1.0, size=x_train.shape)
x_test_noisy = x_test + noise_factor_test * np.random.normal(loc=0.0, scale=1.0, size=x_test.shape)
x_train_noisy = np.clip(x_train_noisy, 0., 1.)
x_test_noisy = np.clip(x_test_noisy, 0., 1.)
return x_train_noisy, x_test_noisy
def main_noisy(noise_factor_train = 0.4, noise_factor_test = 0.7):
(train_images, train_labels), (test_images, test_labels) = load_data()
x_train, y_train, x_test, y_test = prepare_data(train_images, train_labels, test_images, test_labels)
x_train_noisy, x_test_noisy = add_noise(x_train, x_test, noise_factor_train, noise_factor_test)
history = model(x_train_noisy, y_train, x_test_noisy, y_test)
show_result(x_train_noisy, x_test_noisy)
plot(history)
main_noisy(noise_factor_train = 0.9, noise_factor_test = 0.9)
main_noisy(noise_factor_train = 0.7, noise_factor_test = 0.7)
main_noisy(noise_factor_train = 0.3, noise_factor_test = 0.3)
main_noisy(noise_factor_train = 0.4, noise_factor_test = 0.7)
main_noisy(noise_factor_train = 0.7, noise_factor_test = 0.4)
```
| github_jupyter |
```
# notebook that goes through and produces the same plot regional_plot_script.py
import os
os.getcwd()
import os
os.chdir('..') # assume cwd was regional_plots
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib.dates as mdates
import matplotlib.ticker as ticker
today="2021-03-07_region"
website_folder="docs/assets/data"
regions = pd.read_csv("data/nhs_regions.csv", header=0).iloc[:,0].to_list()
cpred_df = pd.read_csv(f"{website_folder}/{today}/Cpred_region.csv")
cproj_df = pd.read_csv(f"{website_folder}/{today}/Cproj_region.csv")
cactual_df = pd.read_csv(f"{website_folder}/region_site_data.csv")
rt_df = pd.read_csv(f"{website_folder}/{today}/Rt_region.csv")
def subset_df(df, col, val):
condition = (df[col] == val)
return df.loc[condition]
from matplotlib.dates import date2num as d2n
month_dict = {"01": "Jan", "02": "Feb", "03": "Mar", "04": "Apr", "05": "May", "06": "Jun", "07": "Jul", "08": "Aug", "09": "Sep", "10": "Oct", "11": "Nov", "12": "Dec"}
def get_xticks_and_xlabels(dfs, col="Date"):
date_df = pd.concat([df["Date"] for df in dfs])
# return date_df
dates = date_df.unique()
months = [month_dict[d.split("-")[1]] for d in dates]
date_nums = d2n(dates)
indx = [d.split("-")[-1] == "01" for d in dates]
indx = [i for i,x in enumerate(indx) if x]
date_nums = date_nums[indx]
months = [months[id] for id in indx]
return date_nums, months
def make_cases_plot(ax, region, cpred_df, cproj_df, actual_case_df, end=today, model_col="royalblue", actual_col="dodgerblue"):
ax.set_title(f"{region} cases")
ax.set_ylabel(f"Daily case count")
# ax.set_xlabel(f"Date")
reg_cpred_df = subset_df(cpred_df, "area", region)
reg_cproj_df = subset_df(cproj_df, "area", region)
reg_actual_df = subset_df(actual_case_df, "NHS_Region", region)
ax.plot(d2n(reg_cpred_df["Date"]), reg_cpred_df["C_50"], color=model_col)
ax.fill_between(d2n(reg_cpred_df["Date"]), reg_cpred_df["C_25"], reg_cpred_df["C_75"], color=model_col, alpha=0.5)
ax.fill_between(d2n(reg_cpred_df["Date"]), reg_cpred_df["C_025"], reg_cpred_df["C_975"], color=model_col, alpha=0.25)
ax.axvline(d2n(reg_cpred_df["Date"])[-1], ls="--", color=model_col, alpha=0.7)
ax.plot(d2n(reg_cproj_df["Date"]), reg_cproj_df["C_50"], color="k", ls="--")
ax.fill_between(d2n(reg_cproj_df["Date"]), reg_cproj_df["C_25"], reg_cproj_df["C_75"], color="k", alpha=0.5)
ax.fill_between(d2n(reg_cproj_df["Date"]), reg_cproj_df["C_025"], reg_cproj_df["C_975"], color="k", alpha=0.25)
ax.plot(d2n(reg_actual_df["Date"]), reg_actual_df["cases_new"], color=actual_col, alpha=0.4)
xticks, xlabels = get_xticks_and_xlabels([reg_cpred_df, reg_cproj_df, reg_actual_df])
ax.set_xticks(xticks)
ax.set_xticklabels(xlabels)
def make_r_plot(ax, region, rt_df, actual_case_df, end=today, model_col="royalblue", tick_spacing=0.5):
ax.yaxis.set_major_locator(ticker.MultipleLocator(tick_spacing))
ax.set_title(f"{region} Rt")
ax.set_ylabel(f"Rt")
# ax.set_xlabel(f"Date")
reg_rt_df = subset_df(rt_df, "area", region)
inferred_df = reg_rt_df.loc[reg_rt_df["provenance"] == "inferred"]
projected_df = reg_rt_df.loc[reg_rt_df["provenance"] == "projected"]
ax.axvline(d2n(inferred_df["Date"])[-1], ls="--", color=model_col, alpha=0.7)
ax.axhline(1.0, ls="--", color="k", alpha=0.9)
ax.plot(d2n(inferred_df["Date"]), inferred_df["Rt_50"], color=model_col)
ax.fill_between(d2n(inferred_df["Date"]), inferred_df["Rt_25"], inferred_df["Rt_75"], color=model_col, alpha=0.5)
ax.fill_between(d2n(inferred_df["Date"]), inferred_df["Rt_2_5"], inferred_df["Rt_97_5"], color=model_col, alpha=0.25)
ax.plot(d2n(projected_df["Date"]), projected_df["Rt_50"], color="k", ls="--")
ax.fill_between(d2n(projected_df["Date"]), projected_df["Rt_25"], projected_df["Rt_75"], color="k", alpha=0.5)
ax.fill_between(d2n(projected_df["Date"]), projected_df["Rt_2_5"], projected_df["Rt_97_5"], color="k", alpha=0.25)
xticks, xlabels = get_xticks_and_xlabels([reg_rt_df, actual_case_df])
ax.set_xticks(xticks)
ax.set_xticklabels(xlabels)
fig, axs = plt.subplots(9, 2, figsize=(15,35))
ax_list = [axs[r_id, 1] for r_id, _ in enumerate(regions)]
ax_list[0].get_shared_y_axes().join(*ax_list)
for r_id, region in enumerate(regions):
for id in range(2):
ax = axs[r_id, id]
if id==0:
make_cases_plot(ax, region, cpred_df, cproj_df, cactual_df)
if id==1:
make_r_plot(ax, region, rt_df, cactual_df)
# for ax in axs.reshape(-1):
# ax.set_ylim(bottom=0)
fig.savefig(f"{website_folder}/{today}/regional_plot.pdf")
```
| github_jupyter |
This is a type of location optimization analysis, specifically finding the optimal location of facilites on a network. These are two types of Set-Coverage analysis implemented in **Python**:
### Set-Coverage Problem
#### Objective: Determine the minimum number of facilities and their locations in order to cover all demands within a pre-specified maximum distance (or time) coverage
### Partial Set-Coverage Problem
#### Objective: Determine the minimum number of facilities and their locations in order to cover a given fraction of the population within a pre-specified maximum distance (or time) coverage
more information on GOSTNets Optimization can be found in the wiki: https://github.com/worldbank/GOST_PublicGoods/wiki/GOSTnets-Optimization
```
import geopandas as gpd
import pandas as pd
import os, sys
# add to your system path the location of the GOSTnet.py library
sys.path.append(r'../../../GOSTNets/GOSTNets')
import GOSTnet_Optimization as gn
import importlib
```
#### These Set-Coverage Problems require an OD matrix as an input¶
read in OD matrix saved as csv file as dataframe
```
pth = r'../../../../lima_optimization_output'
OD_df = pd.read_csv(os.path.join(pth, 'saved_OD.csv'),index_col=0)
import pulp
```
### Run the Set-Coverage Problem
#### Objective: Determine the minimum number of facilities and their locations in order to cover all demands within a pre-specified maximum distance (or time) coverage
In the example below there is a pre-specified maximum distance coverage of 1200 seconds
```
set_coverage_result = gn.optimize_set_coverage(OD_df, max_coverage = 1200)
set_coverage_result
```
### Partial Set-Coverage Problem
#### Objective: Determine the minimum number of facilities and their locations in order to cover a given fraction of the population within a pre-specified maximum distance (or time) coverage
This problem factors in population coverage, therefore as an additional input we need to produce a series that has each origin and its respective population
```
origins_gdf = pd.read_csv(os.path.join(pth, 'origins_snapped.csv'))
#origins_gdf
#origins_gdf.Population.values
origins_w_pop_series = pd.Series(origins_gdf.Population.values, index=origins_gdf.NN)
len(origins_w_pop_series)
#some origins end up snapping to the same nearest node, therefore the code below groups and sums origin populations
origins_w_pop_series_no_dupl = origins_w_pop_series.groupby('NN').sum()
len(origins_w_pop_series_no_dupl)
origins_w_pop_series_no_dupl[:10]
#sum(origins_w_pop_series_no_dupl)
origins_w_pop_series_no_dupl[3]
```
In the example below, inputs include covering 90 percent of the population, a pre-specified maximum distance coverage of 1200 seconds, and a series of origins with their population
```
OD_df[:5]
import importlib
importlib.reload(gn)
# neet to double-check this
partial_set_coverage_result = gn.optimize_partial_set_coverage(OD_df, pop_coverage = .8, max_coverage = 900, origins_pop_series = origins_w_pop_series_no_dupl, existing_facilities = None)
partial_set_coverage_result
```
| github_jupyter |
ERROR: type should be string, got "https://arxiv.org/pdf/1810.04805.pdf\n\n```\nimport os\nos.sys.path.append('..')\n%matplotlib inline\n%reload_ext autoreload\n%autoreload 2\nfrom __future__ import absolute_import\nfrom __future__ import division\nfrom __future__ import print_function\n\nimport argparse\nimport collections\nimport logging\nimport json\nimport math\nimport os\nimport random\nimport six\nfrom tqdm import tqdm_notebook as tqdm\nfrom IPython.display import HTML, display\n\nimport numpy as np\nimport torch\nimport torch.nn as nn\nimport torch.nn.functional as F\nfrom torch.utils.data import TensorDataset, DataLoader, RandomSampler, SequentialSampler\nfrom torch.utils.data.distributed import DistributedSampler\n\nimport tokenization\nfrom modeling import BertConfig, BertForMaskedLanguageModelling\nfrom optimization import BERTAdam\nfrom masked_language_model import notqdm, convert_tokens_to_features, LMProcessor, predict_masked_words, predict_next_words, improve_words_recursive\n\nlogging.basicConfig(format = '%(asctime)s - %(levelname)s - %(name)s - %(message)s', \n datefmt = '%m/%d/%Y %H:%M:%S',\n level = logging.INFO)\nlogger = logging.getLogger(__name__)\n```\n\n# Args\n\n```\nparser = argparse.ArgumentParser()\n\n## Required parameters\nparser.add_argument(\"--data_dir\",\n default=None,\n type=str,\n required=True,\n help=\"The input data dir. Should contain the .tsv files (or other data files) for the task.\")\nparser.add_argument(\"--bert_config_file\",\n default=None,\n type=str,\n required=True,\n help=\"The config json file corresponding to the pre-trained BERT model. \\n\"\n \"This specifies the model architecture.\")\nparser.add_argument(\"--task_name\",\n default=None,\n type=str,\n required=True,\n help=\"The name of the task to train.\")\nparser.add_argument(\"--vocab_file\",\n default=None,\n type=str,\n required=True,\n help=\"The vocabulary file that the BERT model was trained on.\")\nparser.add_argument(\"--output_dir\",\n default=None,\n type=str,\n required=True,\n help=\"The output directory where the model checkpoints will be written.\")\n\n## Other parameters\nparser.add_argument(\"--init_checkpoint\",\n default=None,\n type=str,\n help=\"Initial checkpoint (usually from a pre-trained BERT model).\")\nparser.add_argument(\"--do_lower_case\",\n default=False,\n action='store_true',\n help=\"Whether to lower case the input text. True for uncased models, False for cased models.\")\nparser.add_argument(\"--max_seq_length\",\n default=128,\n type=int,\n help=\"The maximum total input sequence length after WordPiece tokenization. \\n\"\n \"Sequences longer than this will be truncated, and sequences shorter \\n\"\n \"than this will be padded.\")\nparser.add_argument(\"--do_train\",\n default=False,\n action='store_true',\n help=\"Whether to run training.\")\nparser.add_argument(\"--do_eval\",\n default=False,\n action='store_true',\n help=\"Whether to run eval on the dev set.\")\nparser.add_argument(\"--train_batch_size\",\n default=32,\n type=int,\n help=\"Total batch size for training.\")\nparser.add_argument(\"--eval_batch_size\",\n default=8,\n type=int,\n help=\"Total batch size for eval.\")\nparser.add_argument(\"--learning_rate\",\n default=5e-5,\n type=float,\n help=\"The initial learning rate for Adam.\")\nparser.add_argument(\"--num_train_epochs\",\n default=3.0,\n type=float,\n help=\"Total number of training epochs to perform.\")\nparser.add_argument(\"--warmup_proportion\",\n default=0.1,\n type=float,\n help=\"Proportion of training to perform linear learning rate warmup for. \"\n \"E.g., 0.1 = 10%% of training.\")\nparser.add_argument(\"--no_cuda\",\n default=False,\n action='store_true',\n help=\"Whether not to use CUDA when available\")\nparser.add_argument(\"--local_rank\",\n type=int,\n default=-1,\n help=\"local_rank for distributed training on gpus\")\nparser.add_argument('--seed', \n type=int, \n default=42,\n help=\"random seed for initialization\")\nparser.add_argument('--gradient_accumulation_steps',\n type=int,\n default=1,\n help=\"Number of updates steps to accumualte before performing a backward/update pass.\")\nexperiment_name = 'poetry_uncased_5_tied_mlm'\n\nargv = \"\"\"\n--task_name lm \\\n--data_dir {DATA_DIR} \\\n--vocab_file {BERT_BASE_DIR}/vocab.txt \\\n--bert_config_file {BERT_BASE_DIR}/bert_config.json \\\n--init_checkpoint {BERT_BASE_DIR}/pytorch_model.bin \\\n--do_train \\\n--do_eval \\\n--gradient_accumulation_steps 2 \\\n--train_batch_size 16 \\\n--learning_rate 3e-5 \\\n--num_train_epochs 3.0 \\\n--max_seq_length 128 \\\n--output_dir ../outputs/{name}/\n\"\"\".format(\n BERT_BASE_DIR='../data/weights/cased_L-12_H-768_A-12',\n DATA_DIR='../data/input/poetry_gutenberg',\n name=experiment_name\n).replace('\\n', '').split(' ')\nprint(argv)\nargs = parser.parse_args(argv)\n```\n\n# Init\n\n```\nif args.local_rank == -1 or args.no_cuda:\n device = torch.device(\"cuda\" if torch.cuda.is_available() and not args.no_cuda else \"cpu\")\n n_gpu = torch.cuda.device_count()\nelse:\n device = torch.device(\"cuda\", args.local_rank)\n n_gpu = 1\n # Initializes the distributed backend which will take care of sychronizing nodes/GPUs\n torch.distributed.init_process_group(backend='nccl')\nlogger.info(\"device %s n_gpu %d distributed training %r\", device, n_gpu, bool(args.local_rank != -1))\n\nif args.gradient_accumulation_steps < 1:\n raise ValueError(\"Invalid gradient_accumulation_steps parameter: {}, should be >= 1\".format(\n args.gradient_accumulation_steps))\n\nargs.train_batch_size = int(args.train_batch_size / args.gradient_accumulation_steps)\n\nrandom.seed(args.seed)\nnp.random.seed(args.seed)\ntorch.manual_seed(args.seed)\nif n_gpu > 0:\n torch.cuda.manual_seed_all(args.seed)\n\nif not args.do_train and not args.do_eval:\n raise ValueError(\"At least one of `do_train` or `do_eval` must be True.\")\nbert_config = BertConfig.from_json_file(args.bert_config_file)\n\nif args.max_seq_length > bert_config.max_position_embeddings:\n raise ValueError(\n \"Cannot use sequence length {} because the BERT model was only trained up to sequence length {}\".format(\n args.max_seq_length, bert_config.max_position_embeddings))\nif os.path.exists(args.output_dir) and os.listdir(args.output_dir):\n print(\"Output directory ({}) already exists and is not empty.\".format(args.output_dir))\nos.makedirs(args.output_dir, exist_ok=True)\nsave_path = os.path.join(args.output_dir, 'state_dict.pkl')\nsave_path\n```\n\n# Load data\n\n```\ntokenizer = tokenization.FullTokenizer(\n vocab_file=args.vocab_file, do_lower_case=args.do_lower_case)\n\ndecoder = {v:k for k,v in tokenizer.wordpiece_tokenizer.vocab.items()}\nprocessors = {\n \"lm\": LMProcessor,\n}\n \ntask_name = args.task_name.lower()\nif task_name not in processors:\n raise ValueError(\"Task not found: %s\" % (task_name))\n\nprocessor = processors[task_name](tokenizer=tokenizer)\nlabel_list = processor.get_labels()\ntrain_examples = processor.get_train_examples(args.data_dir, skip=30, tqdm=tqdm)\nnum_train_steps = int(\n len(train_examples) / args.train_batch_size * args.num_train_epochs)\ntrain_features = convert_tokens_to_features(\n train_examples, label_list, args.max_seq_length, tokenizer, tqdm=tqdm)\n\nall_input_ids = torch.tensor([f.input_ids for f in train_features], dtype=torch.long)\nall_input_mask = torch.tensor([f.input_mask for f in train_features], dtype=torch.long)\nall_segment_ids = torch.tensor([f.segment_ids for f in train_features], dtype=torch.long)\nall_label_ids = torch.tensor([f.label_id for f in train_features], dtype=torch.long)\nall_label_weights = torch.tensor([f.label_weights for f in train_features], dtype=torch.long)\ntrain_data = TensorDataset(all_input_ids, all_input_mask, all_segment_ids, all_label_ids, all_label_weights)\nif args.local_rank == -1:\n train_sampler = RandomSampler(train_data)\nelse:\n train_sampler = DistributedSampler(train_data)\ntrain_dataloader = DataLoader(train_data, sampler=train_sampler, batch_size=args.train_batch_size)\n```\n\n# Load model\n\n```\nmodel = BertForMaskedLanguageModelling(bert_config)\nif args.init_checkpoint is not None:\n model.bert.load_state_dict(torch.load(args.init_checkpoint, map_location='cpu'))\n \nif os.path.isfile(save_path):\n model.load_state_dict(torch.load(save_path, map_location='cpu'))\n \nmodel.to(device)\n\nif args.local_rank != -1:\n model = torch.nn.parallel.DistributedDataParallel(model, device_ids=[args.local_rank],\n output_device=args.local_rank)\nelif n_gpu > 1:\n model = torch.nn.DataParallel(model)\n \nmodel\n```\n\n# Opt\n\n```\nno_decay = ['bias', 'gamma', 'beta']\noptimizer_parameters = [\n {'params': [p for n, p in model.named_parameters() if n not in no_decay], 'weight_decay_rate': 0.01},\n {'params': [p for n, p in model.named_parameters() if n in no_decay], 'weight_decay_rate': 0.0}\n ]\n\noptimizer = BERTAdam(optimizer_parameters,\n lr=args.learning_rate,\n warmup=args.warmup_proportion,\n t_total=num_train_steps)\n```\n\n# Train 3\n\n```\nval_test=\"\"\"What is more gentle than a wind in summer? What is more soothing than the pretty hummer That stays one moment in an open flower, And buzzes cheerily from bower to bower? What is more tranquil than a musk-rose blowing In a green island, far from all men's knowing?\"\"\"\ndisplay(predict_next_words(val_test, processor, tokenizer, model, max_seq_length=args.max_seq_length, n=20, T=1, device=device))\n# val_test=\"\"\"The next day I was somewhat somnolent, of which you may be sure Miss Frankland took no notice. She retired to her own room when we went for our recreation. My sisters scolded me for not coming to them the previous night, but I told them that Miss F. had continued to move about her room for so long a time that I had fallen fast asleep, and even then had not had enough, as they might have observed how sleepy I had been all\"\"\"\ndisplay(predict_next_words(val_test, processor, tokenizer, model, max_seq_length=args.max_seq_length, n=20, T=1, device=device))\ndisplay(predict_masked_words(val_test, processor, tokenizer, model, device=device, max_seq_length=args.max_seq_length))\nglobal_step = 0\nmodel.train()\nfor _ in tqdm(range(int(args.num_train_epochs)), desc=\"Epoch\"):\n tr_loss, nb_tr_examples, nb_tr_steps = 0, 0, 0\n with tqdm(total=len(train_dataloader), desc='Iteration', mininterval=60) as prog:\n for step, batch in enumerate(train_dataloader):\n \n batch = tuple(t.to(device) for t in batch)\n input_ids, input_mask, segment_ids, label_ids, label_weights = batch\n loss, logits = model(input_ids, segment_ids, input_mask, label_ids, label_weights)\n if n_gpu > 1:\n loss = loss.mean() # mean() to average on multi-gpu.\n if args.gradient_accumulation_steps > 1:\n loss = loss / args.gradient_accumulation_steps\n loss.backward()\n tr_loss += loss.item()\n nb_tr_examples += input_ids.size(0)\n nb_tr_steps += 1\n if (step + 1) % args.gradient_accumulation_steps == 0:\n optimizer.step() # We have accumulated enougth gradients\n model.zero_grad()\n prog.update(1)\n prog.desc = 'Iter. loss={:2.6f}'.format(tr_loss/nb_tr_examples)\n if step%3000==10:\n \n print('step', step, 'loss', tr_loss/nb_tr_examples)\n display(predict_masked_words(val_test, processor, tokenizer, model, device=device, max_seq_length=args.max_seq_length))\n display(predict_next_words(val_test, processor, tokenizer, model, max_seq_length=args.max_seq_length, n=10, device=device))\n tr_loss, nb_tr_examples, nb_tr_steps = 0, 0, 0\n \n # TODO validation test at end of each epoch to check for overfitting\n \n \n torch.save(model.state_dict(), save_path)\n\nglobal_step += 1\ndisplay(predict_next_words(val_test, processor, tokenizer, model, max_seq_length=args.max_seq_length, n=10, device=device, debug=False))\ntorch.save(model.state_dict(), save_path)\nval_test=\"\"\"Frank could no longer resist\"\"\"\ndisplay(predict_next_words(val_test, processor, tokenizer, model, max_seq_length=args.max_seq_length, n=100, T=.1, device=device))\nval_test=\"\"\"There was no doubt the lad had seen everything\"\"\"\ndisplay(predict_next_words(val_test, processor, tokenizer, model, max_seq_length=args.max_seq_length, n=26, T=.5, device=device))\nval_test=\"\"\"The next night I had been asleep about a couple of hours when I was suddenly awakened by\"\"\"\ndisplay(predict_next_words(val_test, processor, tokenizer, model, max_seq_length=args.max_seq_length, n=100, T=1, device=device))\nval_test=\"\"\"His mind spun in on itself\"\"\"\ndisplay(predict_next_words(val_test, processor, tokenizer, model, max_seq_length=args.max_seq_length, n=150, T=1, device=device))\nval_test=\"\"\"A giant spider descended on to\"\"\"\ndisplay(predict_next_words(val_test, processor, tokenizer, model, max_seq_length=args.max_seq_length, n=100, T=.5, device=device))\nval_test=\"\"\"Madness enveloped his mind as\"\"\"\ndisplay(predict_next_words(val_test, processor, tokenizer, model, max_seq_length=args.max_seq_length, n=100, T=.1, device=device))\nval_test=\"\"\"A thin film of\"\"\"\ndisplay(predict_next_words(val_test, processor, tokenizer, model, max_seq_length=args.max_seq_length, n=100, T=1, device=device))\nval_test=\"\"\"Quivering with fear, he trembled as\"\"\"\ndisplay(predict_next_words(val_test, processor, tokenizer, model, max_seq_length=args.max_seq_length, n=100, T=1, device=device))\nval_test=\"\"\"Madness enveloped his mind as\"\"\"\ndisplay(predict_next_words(val_test, processor, tokenizer, model, max_seq_length=args.max_seq_length, n=300, T=1, device=device))\nval_test=\"\"\"Roses are red, violets are blue, you are something but something\"\"\"\npredict_next_words(val_test, processor, tokenizer, model, max_seq_length=args.max_seq_length, n=100, T=1, device=device)\nval_test=\"\"\"The sun shone on the great home \"\"\"\nh=predict_next_words(val_test, processor, tokenizer, model, max_seq_length=args.max_seq_length, n=12, T=1, device=device)\nh.data = h.data.replace('.', '.<p></p>')\nh\n```\n\n# Generate here\n\n```\nimprove_words_recursive?\nval_test=\"\"\"Quivering with fear, he trembled, he, but lo! Fell he, Again until emptied of life, Again; Till, when Love sighed-- Love dreamed, but dreaming his; and the still stream he very much esteems I plac, or lie large in size Balyms others attributed to Dashdims-- saint of Bat. 4. 143 Bb version-Morgen 144 3 5 sing the Heathings 48 8 singbi 79 12 13 5 4 5 6 Vuivering with fear, he trembled, he, but lo! Fell he, Again until emptied of life, Again; Till, when Love sighed-- Love dreamed, but dreaming his; and the still stream he very much esteems I plac, or lie large in size Balyms others attributed to Dashdims-- saint of Bat. 4. 143 Bb version-Morgen 144 3 5 sing the Heathings\"\"\"\ndisplay(improve_words_recursive(\n val_test, \n processor, \n tokenizer, \n model, \n max_seq_length=args.max_seq_length, \n iterations=50, # How many change to go through, it replaces \n T=1, # Tempreture - Higher gives more random, but less stable output\n device=\"cuda\", \n debug=1\n))\nval_test=\"\"\"What is more gentle than a wind in summer. How is more hearty than a winters soul. Why are we glum unto the Autumn. And where are we that we may skip in the spring. Unto thee I ask the question of the seasons, so you furnish me with empty lies and we may hunt over all for the final lie so we my sleep\"\"\"\ndisplay(improve_words_recursive(val_test, processor, tokenizer, model, max_seq_length=args.max_seq_length, iterations=50, T=1, device=\"cuda\", debug=1))\ntext=\"\"\"It's strange to think that we will live and die\nin the eternal grey,\nAfter years of the same things; sin and toil\nand empty delight.\nI will wed, then father and perish\nwhat else can I do?\nI bore as did those who came before me\nwithout question or blame.\nAnd go hither unto oblivion, leaving neither trace nor name\n\"\"\"\nimprove_words_recursive(text, processor, tokenizer, model, iterations=100, max_seq_length=128, n=10, T=0.9, device=\"cuda\", debug=1)\ntext=\"\"\"Mary had a little lamb\nLittle lamb, little lamb\nMary had a little lamb\nIts fleece was white as snow\nAnd everywhere that Mary went\nMary went, Mary went\nEverywhere that Mary went\nThe lamb was sure to go\n\nHe followed her to school one day\nSchool one day, school one day\nHe followed her to school one day\nWhich was against the rule\nIt made the children laugh and play\nLaugh and play, laugh and play\nIt made the children laugh and play\nTo see a lamb at school\"\"\"\nimprove_words_recursive(text, processor, tokenizer, model, ITERATIVE_MASK_FRAC=0.1, iterations=100, max_seq_length=128, n=10, T=0.3, device=\"cuda\", debug=10)\ntext=\"\"\"The rose is red, the violet's blue,\n The honey's sweet, and so are you.\n Thou are my love and I am thine;\n I drew thee to my Valentine:\n The lot was cast and then I drew,\n And Fortune said it shou'd be you. The rose is red, the violet's blue,\n The honey's sweet, and so are you.\n Thou are my love and I am thine;\n I drew thee to my Valentine:\n The lot was cast and then I drew,\n And Fortune said it shou'd be you.\"\"\"\nimprove_words_recursive(text, processor, tokenizer, model, ITERATIVE_MASK_FRAC=0.2, iterations=100, max_seq_length=128, T=0.7, device=\"cuda\", debug=1)\ntext=\"\"\"I saw the best minds of my generation destroyed by madness, starving hysterical naked,\n\ndragging themselves through the negro streets at dawn looking for an angry fix,\n\nangelheaded hipsters burning for the ancient heavenly connection to the starry dynamo in the machinery of night,\n\nwho poverty and tatters and hollow-eyed and high sat up smoking in the supernatural darkness of cold-water flats floating across the tops of cities contemplating jazz,\n\nwho bared their brains to Heaven under the El and saw Mohammedan angels staggering on tenement roofs illuminated,\n\nwho passed through universities with radiant eyes hallucinating Arkansas and Blake-light tragedy among the scholars of war,\n\nwho were expelled from the academies for crazy & publishing obscene odes on the windows of the skull,\n\nwho cowered in unshaven rooms in underwear, burning their money in wastebaskets and listening to the Terror through the wall,\n\nwho got busted in their pubic beards returning through Laredo with a belt of marijuana for New York,\n\nwho ate fire in paint hotels or drank turpentine in Paradise Alley, death, or purgatoried their torsos night after night\n\nwith dreams, with drugs, with waking nightmares, alcohol and cock and endless balls,\n\nincomparable blind streets of shuddering cloud and lightning in the mind leaping towards poles of Canada & Paterson, illuminating all the motionless world of Time between,\n\nPeyote solidities of halls, backyard green tree cemetery dawns, wine drunkenness over the rooftops, storefront boroughs of teahead joyride neon blinking traffic light, sun and moon and tree vibrations in the roaring winter dusks of Brooklyn, ashcan rantings and kind king light of mind,\"\"\"\nimprove_words_recursive(text, processor, tokenizer, model, ITERATIVE_MASK_FRAC=0.2, iterations=100, max_seq_length=128, n=10, T=1, device=\"cuda\", debug=1)\n```\n\nAnd, as, and the sky's blue, And love's sea, and I was love; And and I of love: I was bide: Love gave me to the sea: Love gave me, and I was sea, And ay it was Fortune's betead. The sky is blue, and man's sea, The sea of the star, and of the sea; And I was deep, though I sing it; And I did say of the sea. And and and and and and I said; And then and then, of the hills and the sea, and and and,\n\nAnd stan'd down on the sea. But ere the the, while they slept, He left them, give'd them upon the dew, He lay'd their bodies in the dew. On the shore were the boars'slings in the dew, Roasting o'er all the hot dew; He rose, and gave them the dew. He bore them down to the sea, He lay'd them with them, On the spray of the and water. Twowain and twas, He led their heads to the shore, And bounding\n\nreason's it I think that we will live and engage in the same works, The works of the same, with fire and toil and empty heart. I was a woman then, but now for what else can I hope. We talk nothing, all that I go under to do... and achieve at all some oar, but do know the one and, to, do have, I after know. things I and that of, of can. I another. do or also over of, spring, have of, little secure, a-and at, at work of one yours,, heart and I\n\nquivering with fear, But he stood there, and ah! saw him, he was void of sight, O!-- and he prayed-- he prayed, and raised his hand to the holy book to see, Too little a mythismen or dream: and the psalm Of living things do to you, my saintly friend. VI. \" Bless ye the holyrysm-- and the book he held him,-- He saw it, \" ( quivering with fear ) He saw, he prayed, lo! but he lived, was void of life, O; he\n\ncarries me out, for one good in each. He is more hearty than the single thing. They are under the darkness of the heaven. And, am I, I may go down the roads, tiptoed off, up out of the mountains. Never you will lead me with a child and we may hold it over me or you or over-night me all night crying night of and and it, me, any, you and and-everywhere and and me in all you a you night, We. you and the me under together up still and and of the thing thing.. up all we ahead andto\n\nit's enough to talk with spirits that see and know in the eternal things of the great, the great, that love to toil and toil. I am one of their friends and foes, and am I one? I am as a one who had it. I am the voice, that passes over to the turner, that is the voice of the life one that it of, the the of,, it is me the man it all, the you., the to living the, of of voice and the the the, to is and, the the one, the great, that love, to\n\n# TODO\n\n- show probability in next word logging. Record probability of each letter, then use them when displaying as html\n- try other ways of doing next word. E.g. going back and redoing, doing more than 1 at once\n- make the masked language generator often mask last word\n- should I be doing loss on just the masked words, or all? It's hard to tell from the tensorflow repo. This is marked with a TODO or FIXME in the code\n- add validation loss, since overfitting seems to be a factor\n- for eval, don't pad, just have a batch size of one. That may lead to better results\n- for eval, add some words, and let it fill in the blanks\n - recursivly replace low confidence words?\n - for this I may have to make it predict unmasked words?\n\n# Search for a result in the training text \n\nTo check it's not just remembering\n\n```\ntrain_txt = open(os.path.join(args.data_dir,'train.txt')).readlines()\nimport difflib \ncandidate_test = 'of the morning of the morning and the evening'\nmatches = []\nfor cand_train in tqdm(train_txt):\n diffl = difflib.SequenceMatcher(None, cand_train.lower().strip(), candidate_test.lower() ).ratio()\n matches.append([diffl, cand_train])\nimport pandas as pd\ndf = pd.DataFrame(matches, columns=['ratio', 'str'])\ndf = df.sort_values('ratio', ascending=False)\ndf\nimport pandas as pd\ndf = pd.DataFrame(matches, columns=['ratio', 'str'])\ndf = df.sort_values('ratio', ascending=False)\ndf\n```\n\n" | github_jupyter |
# Solving Problems With Dynamic Programming
Dynamic programming is a really useful general technique for solving problems that involves breaking down problems into smaller overlapping sub-problems, storing the results computed from the sub-problems and reusing those results on larger chunks of the problem. Dynamic programming solutions are pretty much always more efficent than naive brute-force solutions. Dynamic programming techniques are particularly effective on problems that contain [optimal substructure](https://en.wikipedia.org/wiki/Optimal_substructure).
Dynamic programming is related to a number of other fundamental concepts in computer science in interesting ways. Recursion, for example, is similar to (but not identical to) dynamic programming. The key difference is that in a naive recursive solution, answers to sub-problems may be computed many times. A recursive solution that caches answers to sub-problems which were already computed is called [memoization](https://en.wikipedia.org/wiki/Memoization), which is basically the inverse of dynamic programming. Another variation is when the sub-problems don't actually overlap at all, in which case the technique is known as [divide and conquer](https://en.wikipedia.org/wiki/Divide_and_conquer_algorithms). Finally, dynamic programming is tied to the concept of [mathematical induction](https://en.wikipedia.org/wiki/Mathematical_induction) and can be thought of as a specific application of inductive reasoning in practice.
While the core ideas behind dynamic programming are actually pretty simple, it turns out that it's fairly challenging to use on non-trivial problems because it's often not obvious how to frame a difficult problem in terms of overlapping sub-problems. This is where experience and practice come in handy, which is the idea for this notebook. We'll build both naive and "intelligent" solutions to several well-known problems and see how the problems are decomposed to use dynamic programming solutions.
## Fibonacci Numbers
First we'll look at the problem of computing numbers in the [Fibonacci sequence](https://en.wikipedia.org/wiki/Fibonacci_number). The problem definition is very simple - each number in the sequence is the sum of the two previous numbers in the sequence. Or, more formally:
$F_n = F_{n-1} + F_{n-2}$, with $F_0 = 0$ and $F_1 = 1$ as the seed values.
Our solution will be responsible for calculating each of Fibonacci numbers up to some defined limit. We'll first implement a naive solution that re-calculates each number in the sequence from scratch.
```
def fib(n):
if n == 0:
return 0
if n == 1:
return 1
return fib(n - 1) + fib(n - 2)
def all_fib(n):
fibs = []
for i in range(n):
fibs.append(fib(i))
return fibs
```
Let's try it out on a pretty small number first.
```
%time print(all_fib(10))
```
Okay, probably too trivial. Let's try a bit bigger...
```
%time print(all_fib(20))
```
The runtime was at least measurable now, but still pretty quick. Let's try one more time...
```
%time print(all_fib(40))
```
That escalated quickly! Clearly this is a pretty bad solution. Let's see what it looks like when applying dynamic programming.
```
def all_fib_dp(n):
fibs = []
for i in range(n):
if i < 2:
fibs.append(i)
else:
fibs.append(fibs[i - 2] + fibs[i - 1])
return fibs
```
This time we're saving the result at each iteration and computing new numbers as a sum of the previously saved results. Let's see what this does to the performance of the function.
```
%time print(all_fib_dp(40))
```
By not computing the full recusrive tree on each iteration, we've essentially reduced the running time for the first 40 numbers from ~75 seconds to virtually instant. This also happens to be a good example of the danger of naive recursive functions. Our new Fibonaci number function can compute additional values in linear time vs. exponential time for the first version.
```
%time print(all_fib_dp(100))
```
## Longest Increasing Subsequence
The Fibonacci problem is a good starter example but doesn't really capture the challenge of representing problems in terms of optimal sub-problems because for Fibonacci numbers the answer is pretty obvious. Let's move up one step in difficulty to a problem known as the [longest increasing subsequence](https://en.wikipedia.org/wiki/Longest_increasing_subsequence) problem. The objective is to find the longest subsequence of a given sequence such that all elements in the subsequence are sorted in increasing order. Note that the elements do not need to be contiguous; that is, they are not required to appear next to each other. For example, in the sequence [ 10, 22, 9, 33, 21, 50, 41, 60, 80 ] the longest increasing subsequence (LIS) is [10, 22, 33, 50, 60, 80].
It turns out that it's fairly difficult to do a "brute-force" solution to this problem. The dynamic programming solution is much more concise and a natural fit for the problem definition, so we'll skip creating an unnecessarily complicated naive solution and jump straight to the DP solution.
```
def find_lis(seq):
n = len(seq)
max_length = 1
best_seq_end = -1
# keep a chain of the values of the lis
prev = [0 for i in range(n)]
prev[0] = -1
# the length of the lis at each position
length = [0 for i in range(n)]
length[0] = 1
for i in range(1, n):
length[i] = 0
prev[i] = -1
# start from index i-1 and work back to 0
for j in range(i - 1, -1, -1):
if (length[j] + 1) > length[i] and seq[j] < seq[i]:
# there's a number before position i that increases the lis at i
length[i] = length[j] + 1
prev[i] = j
if length[i] > max_length:
max_length = length[i]
best_seq_end = i
# recover the subsequence
lis = []
element = best_seq_end
while element != -1:
lis.append(seq[element])
element = prev[element]
return lis[::-1]
```
The intuition here is that for a given index $i$, we can compute the length of the longest increasing subsequence $length(i)$ by looking at all indices $j < i$ and if $length(j) + 1 > i$ and $seq[j] < seq[i]$ (meaning there's a number at position $j$ that increases the longest subsequence at that index such that it is now longer than the longest recorded subsequence at $i$) then we increase $length(i)$ by 1. It's a bit confusing at first glance but step through it carefully and convince yourself that this solution finds the optimal subsequence. The "prev" list holds the indices of the elements that form the actual values in the subsequence.
Let's generate some test data and try it out.
```
import numpy as np
seq_small = list(np.random.randint(0, 20, 20))
seq_small
%time print(find_lis(seq_small))
```
Just based on the eye test the output looks correct. Let's see how well it performs on much larger sequences.
```
seq = list(np.random.randint(0, 10000, 10000))
%time print(find_lis(seq))
```
So it's still pretty fast, but the difference is definitely noticable. At 10,000 integers in the sequence our algorithm already takes several seconds to complete. In fact, even though this solution uses dynamic programming its runtime is still $O(n^2)$. The lesson here is that dynamic programming doesn't always result in lightning-fast solutions. There are also different ways to apply DP to the same problem. In fact there's a solution to this problem that uses binary search trees and runs in $O(nlogn)$ time, significantly better than the solution we just came up with.
## Knapsack Problem
The [knapsack problem](https://en.wikipedia.org/wiki/Knapsack_problem) is another classic dynamic programming exercise. The generalization of this problem is very old and comes in many variations, and there are actually multiple ways to tackle this problem aside from dynamic programming. Still, it's a common example for DP exercises.
The problem at its core is one of combinatorial optimization. Given a set of items, each with a mass and a value, determine the collection of items that results in the highest possible value while not exceeding some limit on the total weight. The variation we'll look at is commonly referred to as the 0-1 knapsack problem, which restricts the number of copies of each kind of item to 0 or 1. More formally, given a set of $n$ items each with weight $w_i$ and value $v_i$ along with a maximum total weight $W$, our objective is:
$\Large max \Sigma v_i x_i$, where $\Large \Sigma w_i x_i \leq W$
Let's see what the implementation looks like then discuss why it works.
```
def knapsack(W, w, v):
# create a W x n solution matrix to store the sub-problem results
n = len(v)
S = [[0 for x in range(W)] for k in range(n)]
for x in range(1, W):
for k in range(1, n):
# using this notation k is the number of items in the solution and x is the max weight of the solution,
# so the initial assumption is that the optimal solution with k items at weight x is at least as good
# as the optimal solution with k-1 items for the same max weight
S[k][x] = S[k-1][x]
# if the current item weighs less than the max weight and the optimal solution including this item is
# better than the current optimum, the new optimum is the one resulting from including the current item
if w[k] < x and S[k-1][x-w[k]] + v[k] > S[k][x]:
S[k][x] = S[k-1][x-w[k]] + v[k]
return S
```
The intuition behind this algorithm is that once you've solved for the optimal combination of items at some weight $x < W$ and with some number of items $k < n$, then it's easy to solve the problem with one more item or one higher max weight because you can just check to see if the solution obtained by incorporating that item is better than the best solution you've already found. So how do you get the initial solution? Keep going down the rabbit hole until to reach 0 (in which case the answer is 0). At first glance it's very hard to grasp, but that's part of the magic of dynamic programming. Let's run an example to see what it looks like.
```
w = list(np.random.randint(0, 10, 5))
v = list(np.random.randint(0, 100, 5))
w, v
knapsack(15, w, v)
```
The output here is the array of optimal values for a given max weight (think of it as the column index) and max number of items (the row index). Notice how the output follows what looks sort of like a wavefront pattern. This seems to be a recurring phenomenon with dynamic programming solutions. The value in the lower right corner is the max value that we were looking for under the given constraints and is the answer to the problem.
| github_jupyter |
## N-Gram Language Modelling using NLTK
### Practical Concepts
1. Special Tokens
1. Start Token SOS
2. End Token EOS
3. Unknown Words
2. Preprocessing
1. Padding
2. Replace Singletons / UNK
```
import nltk
from pathlib import Path
SOS = "<s> "
EOS = "</s>"
UNK = "<UNK>"
def add_sentence_tokens(sentences, n):
"""Wrap each sentence in SOS and EOS tokens.
For n >= 2, n-1 SOS tokens are added, otherwise only one is added.
Args:
sentences (list of str): the sentences to wrap.
n (int): order of the n-gram model which will use these sentences.
Returns:
List of sentences with SOS and EOS tokens wrapped around them.
"""
sos = SOS * (n-1) if n > 1 else SOS
return ['{}{} {}'.format(sos, s, EOS) for s in sentences]
def replace_singletons(tokens):
"""Replace tokens which appear only once in the corpus with <UNK>.
Args:
tokens (list of str): the tokens comprising the corpus.
Returns:
The same list of tokens with each singleton replaced by <UNK>.
"""
vocab = nltk.FreqDist(tokens)
return [token if vocab[token] > 1 else UNK for token in tokens]
def preprocess(sentences, n):
"""Add SOS/EOS/UNK tokens to given sentences and tokenize.
Args:
sentences (list of str): the sentences to preprocess.
n (int): order of the n-gram model which will use these sentences.
Returns:
The preprocessed sentences, tokenized by words.
"""
sentences = add_sentence_tokens(sentences, n)
tokens = ' '.join(sentences).split(' ')
tokens = replace_singletons(tokens)
#Add Preprocessing steps here (Refer to Tutorial 1 & 2)
return tokens
%%capture
#@title Data Loading Snippet
def load_data(data_dir):
"""Load train and test corpora from a directory.
Directory must contain two files: train.txt and test.txt.
Newlines will be stripped out.
Args:
data_dir (Path) -- pathlib.Path of the directory to use.
Returns:
The train and test sets, as lists of sentences.
"""
train_path = data_dir.joinpath('train.txt').absolute().as_posix()
test_path = data_dir.joinpath('test.txt').absolute().as_posix()
with open(train_path, 'r') as f:
train = [l.strip() for l in f.readlines()]
with open(test_path, 'r') as f:
test = [l.strip() for l in f.readlines()]
return train, test
!wget https://raw.githubusercontent.com/joshualoehr/ngram-language-model/master/data/train.txt
!wget https://raw.githubusercontent.com/joshualoehr/ngram-language-model/master/data/test.txt
data_path = Path("./")
train, test = load_data(data_path)
```
### Lets take a look at some examples from our corpus
```
import random
print("Train Sentences:\n{}".format("\n".join(random.sample(train, 3))))
print()
print("Test Sentences:\n{}".format("\n".join(random.sample(train, 3))))
```
## Language Model
1. Smoothing
2. Modelling
3. Perplexity
4. OOV Conversion
5. Sentence Generation
```
import argparse
from itertools import product
import math
class LanguageModel(object):
"""An n-gram language model trained on a given corpus.
For a given n and given training corpus, constructs an n-gram language
model for the corpus by:
1. preprocessing the corpus (adding SOS/EOS/UNK tokens)
2. calculating (smoothed) probabilities for each n-gram
Also contains methods for calculating the perplexity of the model
against another corpus, and for generating sentences.
Args:
train_data (list of str): list of sentences comprising the training corpus.
n (int): the order of language model to build (i.e. 1 for unigram, 2 for bigram, etc.).
laplace (int): lambda multiplier to use for laplace smoothing (default 1 for add-1 smoothing).
"""
def __init__(self, train_data, n, laplace=1):
self.n = n
self.laplace = laplace
self.tokens = preprocess(train_data, n)
self.vocab = nltk.FreqDist(self.tokens)
self.model = self._create_model()
self.masks = list(reversed(list(product((0,1), repeat=n))))
def _smooth(self):
"""Apply Laplace smoothing to n-gram frequency distribution.
Here, n_grams refers to the n-grams of the tokens in the training corpus,
while m_grams refers to the first (n-1) tokens of each n-gram.
Returns:
dict: Mapping of each n-gram (tuple of str) to its Laplace-smoothed
probability (float).
"""
vocab_size = len(self.vocab)
n_grams = nltk.ngrams(self.tokens, self.n)
n_vocab = nltk.FreqDist(n_grams)
m_grams = nltk.ngrams(self.tokens, self.n-1)
m_vocab = nltk.FreqDist(m_grams)
def smoothed_count(n_gram, n_count):
m_gram = n_gram[:-1]
m_count = m_vocab[m_gram]
return (n_count + self.laplace) / (m_count + self.laplace * vocab_size)
return { n_gram: smoothed_count(n_gram, count) for n_gram, count in n_vocab.items() }
def _create_model(self):
"""Create a probability distribution for the vocabulary of the training corpus.
If building a unigram model, the probabilities are simple relative frequencies
of each token with the entire corpus.
Otherwise, the probabilities are Laplace-smoothed relative frequencies.
Returns:
A dict mapping each n-gram (tuple of str) to its probability (float).
"""
if self.n == 1:
num_tokens = len(self.tokens)
return { (unigram,): count / num_tokens for unigram, count in self.vocab.items() }
else:
return self._smooth()
def _convert_oov(self, ngram):
"""Convert, if necessary, a given n-gram to one which is known by the model.
Starting with the unmodified ngram, check each possible permutation of the n-gram
with each index of the n-gram containing either the original token or <UNK>. Stop
when the model contains an entry for that permutation.
This is achieved by creating a 'bitmask' for the n-gram tuple, and swapping out
each flagged token for <UNK>. Thus, in the worst case, this function checks 2^n
possible n-grams before returning.
Returns:
The n-gram with <UNK> tokens in certain positions such that the model
contains an entry for it.
"""
mask = lambda ngram, bitmask: tuple((token if flag == 1 else "<UNK>" for token,flag in zip(ngram, bitmask)))
ngram = (ngram,) if type(ngram) is str else ngram
for possible_known in [mask(ngram, bitmask) for bitmask in self.masks]:
if possible_known in self.model:
return possible_known
def perplexity(self, test_data):
"""Calculate the perplexity of the model against a given test corpus.
Args:
test_data (list of str): sentences comprising the training corpus.
Returns:
The perplexity of the model as a float.
"""
test_tokens = preprocess(test_data, self.n)
test_ngrams = nltk.ngrams(test_tokens, self.n)
N = len(test_tokens)
known_ngrams = (self._convert_oov(ngram) for ngram in test_ngrams)
probabilities = [self.model[ngram] for ngram in known_ngrams]
return math.exp((-1/N) * sum(map(math.log, probabilities)))
def _best_candidate(self, prev, i, without=[]):
"""Choose the most likely next token given the previous (n-1) tokens.
If selecting the first word of the sentence (after the SOS tokens),
the i'th best candidate will be selected, to create variety.
If no candidates are found, the EOS token is returned with probability 1.
Args:
prev (tuple of str): the previous n-1 tokens of the sentence.
i (int): which candidate to select if not the most probable one.
without (list of str): tokens to exclude from the candidates list.
Returns:
A tuple with the next most probable token and its corresponding probability.
"""
blacklist = ["<UNK>"] + without
candidates = ((ngram[-1],prob) for ngram,prob in self.model.items() if ngram[:-1]==prev)
candidates = filter(lambda candidate: candidate[0] not in blacklist, candidates)
candidates = sorted(candidates, key=lambda candidate: candidate[1], reverse=True)
if len(candidates) == 0:
return ("</s>", 1)
else:
return candidates[0 if prev != () and prev[-1] != "<s>" else i]
def generate_sentences(self, num, min_len=12, max_len=24):
"""Generate num random sentences using the language model.
Sentences always begin with the SOS token and end with the EOS token.
While unigram model sentences will only exclude the UNK token, n>1 models
will also exclude all other words already in the sentence.
Args:
num (int): the number of sentences to generate.
min_len (int): minimum allowed sentence length.
max_len (int): maximum allowed sentence length.
Yields:
A tuple with the generated sentence and the combined probability
(in log-space) of all of its n-grams.
"""
for i in range(num):
sent, prob = ["<s>"] * max(1, self.n-1), 1
while sent[-1] != "</s>":
prev = () if self.n == 1 else tuple(sent[-(self.n-1):])
blacklist = sent + (["</s>"] if len(sent) < min_len else [])
next_token, next_prob = self._best_candidate(prev, i, without=blacklist)
sent.append(next_token)
prob *= next_prob
if len(sent) >= max_len:
sent.append("</s>")
yield ' '.join(sent), -1/math.log(prob)
n_gram = 3
laplace = 0.01
print("Loading {}-gram model...".format(n_gram))
lm = LanguageModel(train, n_gram, laplace=laplace)
print("Vocabulary size: {}".format(len(lm.vocab)))
lm.model[('all', 'star', 'usa')], lm.model[('raising', '14', 'billion')]
perplexity = lm.perplexity(test)
print("Model perplexity: {:.3f}".format(perplexity))
n_gen_sentences = 3
print("Generating sentences...")
for sentence, prob in lm.generate_sentences(n_gen_sentences):
print("{} ({:.5f})".format(sentence, prob))
```
| github_jupyter |
Importing packages
```
import numpy as np
import pandas as pd
import featuretools as ft
import utils
from tqdm import tqdm
from sklearn.cluster import KMeans
from featuretools.primitives import make_agg_primitive
import featuretools.variable_types as vtypes
from tsfresh.feature_extraction.feature_calculators import (number_peaks, mean_abs_change,
cid_ce, last_location_of_maximum, length)
```
Loading data
```
data_path = 'train_FD004.txt'
data = utils.load_data(data_path)
data.head()
```
Creating cutoff times
```
splits = 5
cutoff_time_list = []
for i in tqdm(range(splits)):
cutoff_time_list.append(utils.make_cutoff_times(data))
cutoff_time_list[0].head()
```
Making entitysets
```
nclusters = 50
def make_entityset(data, nclusters, kmeans=None):
X = data[['operational_setting_1', 'operational_setting_2', 'operational_setting_3']]
if kmeans:
kmeans=kmeans
else:
kmeans = KMeans(n_clusters=nclusters).fit(X)
data['settings_clusters'] = kmeans.predict(X)
es = ft.EntitySet('Dataset')
es.entity_from_dataframe(dataframe=data,
entity_id='recordings',
index='index',
time_index='time')
es.normalize_entity(base_entity_id='recordings',
new_entity_id='engines',
index='engine_no')
es.normalize_entity(base_entity_id='recordings',
new_entity_id='settings_clusters',
index='settings_clusters')
return es, kmeans
es, kmeans = make_entityset(data, nclusters)
es
es.plot()
```
Creating features
```
Complexity = make_agg_primitive(lambda x: cid_ce(x, False),
input_types=[vtypes.Numeric],
return_type=vtypes.Numeric,
name="complexity")
fm, features = ft.dfs(entityset=es,
target_entity='engines',
agg_primitives=['last', 'max', Complexity],
trans_primitives=[],
chunk_size=.26,
cutoff_time=cutoff_time_list[0],
max_depth=3,
verbose=True)
fm.to_csv('advanced_fm.csv')
fm.head()
```
Making predictions
```
fm_list = [fm]
splits=4
for i in tqdm(range(1, splits)):
fm = ft.calculate_feature_matrix(entityset=make_entityset(data, nclusters, kmeans=kmeans)[0],
features=features,
chunk_size=.26,
cutoff_time=cutoff_time_list[i])
fm_list.append(fm)
```
| github_jupyter |
# Model Evaluation, Design, and Grid Search
Following (mostly) Python Machine Learning 3rd (Raschka) Chapter 6
# Training/Test Split
This is how we've set-up our examples so far. We'll use the [breast cancer dataset from scikit-learn.](https://scikit-learn.org/stable/modules/generated/sklearn.datasets.load_breast_cancer.html#sklearn.datasets.load_breast_cancer)
```
import pandas as pd
import numpy as np
from sklearn.datasets import load_breast_cancer
bc = load_breast_cancer()
X = bc['data']
y = bc['target']
X_names = bc['feature_names']
bcDf = pd.concat([pd.DataFrame(X),pd.DataFrame(y)], axis=1)
bcDf.columns = list(X_names) + ['target']
pd.set_option('display.max_columns', 50)
bcDf.head()
```
## Check the numerical summaries of this data:
```
bcDf.describe().T
```
## See if there are any interesting distributions or data issues
```
bcDf.isna().sum().sum()
```
- No missing values, so no need to impute.
```
import matplotlib.pyplot as plt
%matplotlib inline
bcDf.hist(figsize=(14,14))
plt.show()
```
- Doesn't look like there are any obvious data quality issues.
- All numerical, but on different scales, so we'll need to standardize the features.
```
import seaborn as sns
sns.heatmap(bcDf.corr())
plt.show()
```
- There are a lot of correlated features (that's bad).
- Recall [multicollinearity](https://en.wikipedia.org/wiki/Multicollinearity) is bad.
- We are going to try using [principal components](https://en.wikipedia.org/wiki/Principal_component_analysis) to extract uncorrelated features.
- We'll take more about exactly what this is doing later.
- We are going to extract 5, which will reduce the feature space from 30 dimensions to 5.
- This is technically a suite of hyperparameters we are introducing, but this is a well researched dataset.
## Pipeline Gameplan
- Split the data into training/test.
- Create feature processing pipeline.
- Standardize features.
- Extract 5 PCA components. (arbitrary decision; could evaluate)
- Fit a Logistic Regression model.
- Evaluate accuracy (relatively balanced dataset so accuracy is okay).
<img src='files/diagrams/06_01.png' style='width: 700px;'>
[Image source: Python Machine Learning 3rd Edition, Raschka](https://github.com/rasbt/python-machine-learning-book-3rd-edition/tree/master/ch06)
#### Benefits of pipelines
- Chains together feature processing and fitting steps.
- No separate feature transformation fits for test data.
- Plays nice with many of the more robust evaluation options.
## Split the data
```
from sklearn.model_selection import train_test_split
def create_splits(X, y):
return train_test_split(X, y, test_size=0.20)
X_train, X_test, y_train, y_test = create_splits(X, y)
print(f'Training sample: {X_train.shape[0]:,}')
print(f'Test sample: {X_test.shape[0]:,}')
print(f'Training sample: {y_train.shape[0]:,}')
print(f'Test sample: {y_test.shape[0]:,}')
```
## Create our pipeline
```
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.decomposition import PCA
from sklearn.linear_model import LogisticRegression
def generate_estimates(x, y, comp=5):
modeling_pipeline = Pipeline([
('scaling', StandardScaler()),
('pca', PCA(n_components=5)),
('model', LogisticRegression(penalty='none'))
]
)
return modeling_pipeline.fit(x, y)
m = generate_estimates(X_train, y_train)
m
```
## Make our predictions on the test set and determine performance estimates
```
from sklearn.metrics import confusion_matrix
y_test_pred = m.predict(X_test)
print(confusion_matrix(y_test, y_test_pred))
from sklearn.metrics import roc_curve
def generate_probs(X, model=m):
return model.predict_proba(X)[:, 1]
def generate_roc(y, probs):
fpr, tpr, _ = roc_curve(y, probs)
return fpr, tpr
fpr_test, tpr_test = generate_roc(y_test, generate_probs(X_test))
fpr_train, tpr_train = generate_roc(y_train, generate_probs(X_train))
plt.plot(fpr_test, tpr_test,'-r')
plt.plot(fpr_train, tpr_train,'-b')
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.legend(['Test','Training'])
plt.plot([0, 1], [0, 1], color='navy', linestyle='--')
plt.show()
from sklearn.metrics import roc_auc_score
roc_auc_score(y_test, generate_probs(X_test))
```
# Why is our performance estimate potentially flawed?
- How representative is the test dataset?
- Why did we choose 20% to hold for the test set?
- Feature sets in our partitions follow the same distribution?
- Did we try any additional settings or parameter combinations?
- We don't really have a baseline to compare our results against.
## What type of variation can we expect in the test data set?
```
samples = 1000
trainingMeans = []
testMeans = []
i = 0
while i < samples:
X_train, X_test, y_train, y_test = create_splits(X, y)
trainingMeans.append(np.mean(y_train))
testMeans.append(np.mean(y_test))
i += 1
plt.hist(trainingMeans)
plt.xlim(0.4, 0.8)
plt.title('Percent of Positive Classes in Training Data')
plt.show()
plt.hist(testMeans)
plt.xlim(0.4, 0.8)
plt.title('Percent of Positive Classes in Test Data')
plt.show()
```
- Smaller dataset will have more vulnerability to sampling issues.
- What if our data set was bigger?
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
# population for our illustration
population = np.random.binomial(1, 0.5, size=1000000)
pd.Series(population).value_counts()
```
#### Let's draw samples at varying sample sizes to see how representative sample means would be from a large distribution.
```
from collections import defaultdict
reps = 100
sample_sizes = [5, 10, 50, 100, 250, 500, 1000, 2500, 5000, 10000, 100000, 200000]
means = defaultdict(list)
for rep in range(reps):
for ss in sample_sizes:
means[ss].append(np.random.choice(a=population, size=ss, replace=False).mean())
meansDf = pd.DataFrame.from_dict(means)
for i in [0.05, 0.1, 0.25, 0.50, 0.75, 0.9, 0.95]:
if i == 0.5:
plt.plot(meansDf.quantile(i), '-b')
else:
plt.plot(meansDf.quantile(i), '--r')
plt.xscale('log')
plt.title('Percentiles of Sampling Distribution')
plt.show()
```
- Small samples are going to be much more at risk for sampling variations.
- It is not advisable to use a simple training/test split for small datasets.
- It's very likely your test set won't mirror your training.
- Neither may mirror the "population".
- For larger datasets, choosing a smaller percentage for test can be okay.
- The number of samples held will be hopefully very large, and have less risk to sampling variation.
### Rascha on Choosing an appropriate ratio for partitioning a dataset into training and test dates
From page 124 of *Python Machine Learning 3rd Edition, Raschka*:
> If we are dividing a dataset into training and test datasets, we have to keep in mind that we are withholding valuable information that the learning algorithm could benefit from.<br><br>Thus, we don't want to allocate too much information to the test set.<br><br>However, the smaller the test set, the more inaccurate the estimation of the generalization error.<br><br>Dividing a dataset into training and test datasets is all about balancing this tradeoff.<br><br>...<br><br>Moreover, instead of discarding the allocated test data after model training and evaluation, it is a common practice to retrain a classifier on the entire dataset, as it can improve the predictive performance of our model.
## Strengths
- Quick guage of performance.
- Okay if looking for quick interpretability.
## Weaknesses
- Prone to overfitting and not ideal for tuning hyperparameters.
- Many don't partition the training into training and validation, which isn't a good practice.
- There could be other issues with the splits that affect the comparabiltiy and would lead to poor results in the real world. The performance estimate is going to be sensitive to how the partition was done.
## Other issues to be aware
- Is the historical data representative of the current data generating process?
- Have there been structure shifts in the data over time? (leakage)
- Rare event issues and sampling
# Better: Holdout Method
This is for larger datasets, where we don't need to worry about sampling variation risk. We are going to pretend the breast cancer data is "big" enough for this to be a valid approach.
- Split the data into training and test.
> **Develop and train our models using the training partition and estimate general performance on the test set.**
**PRETEND THE TEST SET DOESN'T EXIST WHILE MODELING OR PERFORMING PREPROCESSING. IDEAS WHY?**
- Split the training into training and validation.
> **If we train our models on the entire training dataset, we are going to be at risk for overfitting? Why?**
- Training data is where we'll be creating models and those will be evaluated on the validation data. The test set is for final validation and check that we didn't overfit on the validation set.
<img src='files/diagrams/training-validation-test-data-set.png' style='width: 600px;'>
[Image source: Raschka, Chapter 6, Page 196](https://github.com/rasbt/python-machine-learning-book-3rd-edition/tree/master/ch06)
## Chaining train_test_split
```
from sklearn.model_selection import train_test_split
def create_holdout_splits(X, y):
x_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2)
X_train, X_val, y_train, y_val = train_test_split(x_train, y_train, test_size=0.2)
return X_train, X_val, X_test, y_train, y_val, y_test
X_train, X_val, X_test, y_train, y_val, y_test = create_holdout_splits(X, y)
print(f'Training sample: {X_train.shape[0]:,}')
print(f'Validation sample: {X_val.shape[0]:,}')
print(f'Test sample: {X_test.shape[0]:,}')
```
## Compare and evaluate models
Let's see how the PCA model to a model that uses the full feature set.
### Create pipeline we can use for feature transforms and prediction.
- Might as well check our assumption on using 5 components.
```
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.decomposition import PCA
from sklearn.linear_model import LogisticRegression
def generate_estimates(use_pca = False, comp=None):
if use_pca:
modeling_pipeline = Pipeline([
('scaling', StandardScaler()),
('pca', PCA(n_components=comp)),
('model', LogisticRegression(penalty='none'))
]
)
else:
modeling_pipeline = Pipeline([
('scaling', StandardScaler()),
('model', LogisticRegression(penalty='none'))
]
)
return modeling_pipeline
m_rawFeatures = generate_estimates(use_pca=False).fit(X_train, y_train)
print(m_rawFeatures)
m_pca_models = {}
for i in range(1,6):
m_pca_models[i] = generate_estimates(use_pca=True, comp=i).fit(X_train, y_train)
print(m_pca_models[i])
print('Models fitted')
```
### Compare confusion matrices
```
from sklearn.metrics import confusion_matrix
y_val_rawFeatures = m_rawFeatures.predict(X_val)
print('Using raw features:')
print(confusion_matrix(y_val, y_val_rawFeatures))
y_val_pca = {}
for i in range(1,6):
y_val_pca[i] = m_pca_models[i].predict(X_val)
print(f'Using PCA{i}:')
print(confusion_matrix(y_val, y_val_pca[i]))
```
- Looks like the PCA model performed better and maybe we didn't need all 5 of those components.
### Check the ROC curves
```
from sklearn.metrics import roc_curve
def generate_probs(X, model=m):
return model.predict_proba(X)[:, 1]
def generate_roc(y, probs):
fpr, tpr, _ = roc_curve(y, probs)
return fpr, tpr
fpr_val_rawFeatures, tpr_val_rawFeatures = generate_roc(y_val,
generate_probs(X_val, model=m_rawFeatures))
fpr_val_pca = {}
tpr_val_pca = {}
for i in range(1,6):
fpr_val_pca[i], tpr_val_pca[i] = generate_roc(y_val,
generate_probs(X_val, model=m_pca_models[i]))
plt.plot(fpr_val_rawFeatures, tpr_val_rawFeatures,'-r')
for i in range(1,6):
plt.plot(fpr_val_pca[i], tpr_val_pca[i],'-b')
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.legend(['Raw Features (No Regularization)','PCA'])
plt.plot([0, 1], [0, 1], color='navy', linestyle='--')
plt.show()
```
- Looks like the ROC curves overlap themselves for each component.
- Using PCA performed significantly better in the validation data.
- Now check to see if it continues in the test data.
```
# create confusion matrices
y_test_pca = {}
for i in range(1,6):
y_test_pca[i] = m_pca_models[i].predict(X_test)
print(f'Using PCA{i}:')
print(confusion_matrix(y_test, y_test_pca[i]))
print('\n')
# create false/true positive rate curves
fpr_test_pca = {}
tpr_test_pca = {}
for i in range(1,6):
fpr_test_pca[i], tpr_test_pca[i] = generate_roc(y_test,
generate_probs(X_test,
model=m_pca_models[i])
)
# plot the ROC curve
for i in range(1,6):
plt.plot(fpr_test_pca[i], tpr_test_pca[i],'-b')
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.plot([0, 1], [0, 1], color='navy', linestyle='--')
plt.show()
```
- Looks like the performance was consistent with what we observed in the validation dataset.
- Which model should be choose in this case?
# Cross-validation - Best option for small datasets
<img src='files/diagrams/06_03.png' style='width: 600px;'>
[Image source: Python Machine Learning 3rd Edition, Figure 6.3](https://github.com/rasbt/python-machine-learning-book-3rd-edition/tree/master/ch06)
#### Basic mechanics:
- Split the data into training and test.
- The training data will be divided into *k* folds.
- Each folder will use a different partition for the validation data.
- The models will be run on each fold.
- Performance estimate will be taken with the median or mean score.
- Need to define what metric you are optimizing toward.
You could do this with a loop, but scikit-learn has this built-in.
Let's get our data again:
```
from sklearn.model_selection import train_test_split
def create_splits(X, y):
return train_test_split(X, y, test_size=0.20)
X_train, X_test, y_train, y_test = create_splits(X, y)
print(f'Training sample: {X_train.shape[0]:,}')
print(f'Test sample: {X_test.shape[0]:,}')
```
#### Set-up the same pipeline as before and run it through the function
Things to consider:
- Need an estimator (classifer/regression), which a pipeline satisfies if the last step is a model.
- This accepts multiple metrics, so you'll need to determine which one is most appropriate.
[List of metrics accepted](https://scikit-learn.org/stable/modules/model_evaluation.html#scoring-parameter)
```
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.decomposition import PCA
from sklearn.linear_model import LogisticRegression
def generate_estimates(comp=5):
modeling_pipeline = Pipeline([
('scaling', StandardScaler()),
('pca', PCA(n_components=5)),
('model', LogisticRegression(penalty='none'))
]
)
return modeling_pipeline
from sklearn.model_selection import cross_validate
clf = generate_estimates()
cv_results = cross_validate(clf, X_train, y_train,
scoring=['accuracy', 'recall', 'precision', 'f1_macro', 'roc_auc'], cv=5)
cv_results
```
#### What does this return?
- How long it took the model to fit. This will be more important with larger datasets and more complex models.
- How long it took the model to score the validation set in the fold.
- The metrics.
It returns an array so you can look at the distribution and/or central tendency of the model.
```
for k in cv_results.keys():
print(f'{k}: {cv_results[k].mean():.4f} (+/- {cv_results[k].std():.4f})')
```
## Solved the sampling variation issue, now can compare between models with better certainty
```
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.decomposition import PCA
from sklearn.linear_model import LogisticRegression
def generate_estimates(use_pca = False, comp=None):
if use_pca:
modeling_pipeline = Pipeline([
('scaling', StandardScaler()),
('pca', PCA(n_components=5)),
('model', LogisticRegression(penalty='none'))
]
)
else:
modeling_pipeline = Pipeline([
('scaling', StandardScaler()),
('model', LogisticRegression())
]
)
return modeling_pipeline
from sklearn.model_selection import cross_validate
pca_clf = generate_estimates(use_pca=True, comp=2)
nopca_clf = generate_estimates()
pca_cv_results = cross_validate(pca_clf, X_train, y_train, scoring=['accuracy', 'recall', 'precision', 'f1_macro', 'roc_auc'], cv=5)
pca_cv_results
nopca_cv_results = cross_validate(nopca_clf, X_train, y_train, scoring=['accuracy', 'recall', 'precision', 'f1_macro', 'roc_auc'], cv=5)
nopca_cv_results
a1 = pca_cv_results['test_accuracy'].mean()
a2 = nopca_cv_results['test_accuracy'].mean()
plt.plot(pca_cv_results['test_accuracy'])
plt.plot(nopca_cv_results['test_accuracy'])
plt.legend([f'PCA (mean={a1:.2f})',f'No PCA (mean={a2:.2f})'])
plt.title('Accuracy on Validation')
plt.show()
a1 = pca_cv_results['test_f1_macro'].mean()
a2 = nopca_cv_results['test_f1_macro'].mean()
plt.plot(pca_cv_results['test_f1_macro'])
plt.plot(nopca_cv_results['test_f1_macro'])
plt.legend([f'PCA (mean={a1:.2f})',f'No PCA (mean={a2:.2f})'])
plt.title('F1-Score on Validation')
plt.show()
```
> What looks like the better option?
## Cross-validation not just for classification tasks
- Same concept for regression, the scoring metrics will be different.
- Recall our California housing dataset:
```
from sklearn.datasets import fetch_california_housing
ch = fetch_california_housing()
ch.keys()
X_housing = pd.DataFrame(ch['data'], columns=list(ch['feature_names']))
X_housing.head(2)
y_housing = ch['target']
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.decomposition import PCA
from sklearn.linear_model import LinearRegression
def generate_estimates():
modeling_pipeline = Pipeline([
('scaling', StandardScaler()),
('model', LinearRegression())
]
)
return modeling_pipeline
from sklearn.model_selection import cross_validate
reg = generate_estimates()
cv_results = cross_validate(reg, X_housing, y_housing, scoring=['r2'], cv=10)
cv_results
plt.hist(cv_results['test_r2'])
plt.title('$R^2$ Distribution from Cross-validation')
plt.show()
```
> Looks like the parameters being learned are highly influenced by the training data for California housing.
## But what about setting hyperparameters?
How to determine how many components we may want to use?
### General comments on hyperparameters:
- These aren't learned by the model, they are selected by the analyst.
- You need to learn the best combination by experimenting across the search space.
- If you have a large dataset you can evaluate these in the validation dataset.
- If you have a smaller dataset you can evaluate these using cross validation.
Consider how large your search space can be:
- For regularization, the $C$ parameter in Logistic Regression could span from near zero to approaching infinity.
- For hyperparameters with that type of bounds, you'll likely want to start with a handful of values that are spacing on a logarithmic scale, e.g. $C \in (0.001, 0.01, 0.1, 1, 10, 100, 1000)$.
- If you have other hyperparameters, e.g., the specific solver for Logistic Regression, you'll need to evaulate each solver at each regularization level:
$$\begin{equation*}
PE_{solver,C} =
\begin{pmatrix}
PE_{newton-cg,0.001} & PE_{newton-cg,0.01} & \cdots & PE_{newton-cg,1000} \\
PE_{lbfgs,0.001} & PE_{lbfgs,0.01} & \cdots & PE_{lbfgs,1000} \\
\vdots & \vdots & \ddots & \vdots \\
PE_{saga,0.001} & PE_{saga,0.01} & \cdots & PE_{saga,1000}
\end{pmatrix}
\end{equation*}
$$
- So you if you have 7 different regularization strengths and 5 different solvers, you'll be running $7*5=35$ models. If you use 10 cross-validation folds, that will be 350 models (hence smaller datasets for CV).
- And making it more complicated, you may have multiple metrics (e.g., precision vs. recall).
> When you find the "best" set of hyperparameters, you'll want to explore the nearby space in more detail.
- If you find $C=10$ as the "best", you might want to try another round with $C \in (7,8,9,10,11,12,13)$.
> Finding the "perfect" set of hyperparameters is likely impossible. The search space is generally going to be too large.
- Random search of Bayesian hyperparameter optimization can help in those situations.
- In random search you'll provide a distribution of values instead of discrete values.
- Optimization-based searches will try to make intelligent choices based on past explorations.
# Validation Curves
- You can use these to visualize seeing the differences in different parameters.
- scikit-learn uses a varient on k-fold cross-validation to plot the distribution of metrics for different parameter values.
- We can plot the range of accuracies we observe for many folds across different settings of the parameter.
This is using the breast cancer dataset. We are going to vary $C$ to see its effect. Some of this code is borrowed from page 205-206 of Python Machine Learning 3rd Edition.
```
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.decomposition import PCA
from sklearn.linear_model import LogisticRegression
lg_pipe = modeling_pipeline = Pipeline([
('scaler', StandardScaler()),
('pca', PCA(n_components=5)),
('logreg', LogisticRegression(penalty='l2'))
]
)
from sklearn.model_selection import validation_curve
param_range = [0.001, 0.01, 0.1, 1.0, 10.0, 100.0]
training_scores, test_scores = validation_curve(estimator=lg_pipe, X=X_train, y=y_train,
param_name='logreg__C',
param_range=param_range, cv=10)
train_mean = np.mean(training_scores, axis=1)
train_std = np.std(training_scores, axis=1)
test_mean = np.mean(test_scores, axis=1)
test_std = np.std(test_scores, axis=1)
plt.plot(param_range, train_mean, color='blue', marker='o', label='Training Accuracy')
plt.fill_between(param_range, train_mean + train_std, train_mean - train_std, alpha=0.2, color='blue')
plt.plot(param_range, test_mean, color='orange', marker='o', label='Test Accuracy')
plt.fill_between(param_range, test_mean + test_std, test_mean - test_std, alpha=0.2, color='orange')
plt.xscale('log')
plt.xlabel('$C$')
plt.ylabel('Accuracy')
plt.legend(loc='lower right')
plt.show()
```
> What does this suggest as a parameter for $C$?
# Grid Search
<img src='files/diagrams/grid_search_workflow.png' style='width: 500px;'>
[Image source](https://scikit-learn.org/stable/modules/cross_validation.html#cross-validation)
Examples:
- [Digits dataset](https://scikit-learn.org/stable/auto_examples/model_selection/plot_grid_search_digits.html#sphx-glr-auto-examples-model-selection-plot-grid-search-digits-py)
- [Text extraction](https://scikit-learn.org/stable/auto_examples/model_selection/grid_search_text_feature_extraction.html#sphx-glr-auto-examples-model-selection-grid-search-text-feature-extraction-py)
- [Multiple metric example](https://scikit-learn.org/stable/auto_examples/model_selection/plot_multi_metric_evaluation.html#sphx-glr-auto-examples-model-selection-plot-multi-metric-evaluation-py)
On the breast cancer dataset:
```
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.decomposition import PCA
from sklearn.linear_model import LogisticRegression
lg_pipe = modeling_pipeline = Pipeline([
('scaler', StandardScaler()),
('pca', PCA(n_components=5)),
('logreg', LogisticRegression(penalty='l2', solver='liblinear'))
]
)
from sklearn.model_selection import GridSearchCV
param_grid = [
{'logreg__C': [1, 10, 100, 1000], 'pca__n_components': [1,2,3,4,5,10,15]}
]
gcv_results = GridSearchCV(estimator=lg_pipe, param_grid=param_grid, scoring='accuracy')
gcv_results = gcv_results.fit(X_train, y_train)
```
> This outputs a dictionary with the results:
```
gcv_results.cv_results_.keys()
```
> And a series of summarys for the best fit
```
gcv_results.best_score_
gcv_results.best_params_
gcv_results.best_estimator_
```
> After you find the "best" hyperparameters, you'll retrain your training data using those and then evaluate the test data using that model. There's an option in GridSearchCV to do this automatically.
You could use multiple estimators, but it'll get a little complicated, see below for example:
```python
from sklearn.base import BaseEstimator
from sklearn.model_selection import GridSearchCV
class DummyEstimator(BaseEstimator):
def fit(self): pass
def score(self): pass
# Create a pipeline
pipe = Pipeline([('clf', DummyEstimator())]) # Placeholder Estimator
# Candidate learning algorithms and their hyperparameters
search_space = [{'clf': [LogisticRegression()], # Actual Estimator
'clf__penalty': ['l1', 'l2'],
'clf__C': np.logspace(0, 4, 10)},
{'clf': [DecisionTreeClassifier()], # Actual Estimator
'clf__criterion': ['gini', 'entropy']}]
# Create grid search
gs = GridSearchCV(pipe, search_space)
```
# Evaluation Wrap-up
<img src='files/diagrams/model-eval-conclusions.jpg' style='width: 600px;'>
[Image source: *Model Evaluation, Model Selection, and Algorithm Selection in Machine Learning, Raschka*](https://sebastianraschka.com/blog/2018/model-evaluation-selection-part4.html)
# Unbalanced Classes
Consider the credit from last week:
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
credit = pd.read_csv('data/default.csv', index_col=0)
credit.head()
```
> Remember we had interesting distributions with our numerical variables.
```
credit.hist(bins=50)
plt.show()
credit['balance2income'] = credit['balance']/credit['income']
credit['balance2income'].hist(bins=100)
plt.title('Balance-Income Ratio')
plt.show()
credit['default'].value_counts().plot.barh()
plt.title('Default are relatively rare...', loc='left')
plt.show()
```
### Recall we didn't have the best results with a straightforward logistic regression
> Will try to add some additional features to capture some of the mixture distributions and the truncation of the credit balance.
We can add these prior to processing since they are only considering the example row.
```
credit['balance_student_int'] = np.where(credit['student']=='Yes', credit['balance'], 0)
credit['income_student_int'] = np.where(credit['student']=='Yes', credit['income'], 0)
credit['zero_balance'] = np.where(credit['balance'] == 0, 'Yes', 'No')
credit.head()
```
### We'll do Cross Validation, but still need to split the data into training and test
```
from sklearn.model_selection import train_test_split
creditFeatures = [x for x in credit.columns if x != 'default']
y = np.where(credit['default'] == 'Yes', 1, 0)
X_train, X_test, y_train, y_test = train_test_split(credit[creditFeatures], y, test_size=0.20)
print(f'Training examples: {X_train.shape[0]:,}')
print(f'Test examples: {X_test.shape[0]:,}')
y_train.mean(), y_test.mean()
```
#### Create a modeling pipeline
```
from sklearn.pipeline import Pipeline
from sklearn.compose import ColumnTransformer
from sklearn.preprocessing import StandardScaler
from sklearn.preprocessing import PolynomialFeatures
from sklearn.preprocessing import OneHotEncoder
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import GridSearchCV
nums = ['balance2income', 'balance', 'income', 'balance_student_int', 'income_student_int']
ohes = ['student', 'zero_balance']
processing_pipeline = ColumnTransformer(transformers=[
('numscaling', StandardScaler(), nums),
('dummys', OneHotEncoder(drop='first'), ohes)]
)
modeling_pipeline = Pipeline([
('data_processing', processing_pipeline),
('logreg', LogisticRegression())]
)
```
### Run a baseline model
```
creditbase = modeling_pipeline.fit(X_train, y_train)
base_p = creditbase.predict(X_test)
base_pr = creditbase.predict_proba(X_test)
from sklearn.metrics import classification_report
print(classification_report(y_test, base_p))
```
### Define search space
> We'll try using weights to try to account for the unbalanced distribution of defaults.
```
param_grid = [
{'logreg__class_weight': [None, 'balanced'], 'logreg__C':[0.01, 0.1, 1, 10, 100]}
]
```
### Run the experiment
> Let's look for high-recall, since default could be very expensive.
```
gcv_results = GridSearchCV(estimator=modeling_pipeline, param_grid=param_grid, scoring='recall', refit=True)
gcv_results = gcv_results.fit(X_train, y_train)
gcv_results.best_estimator_
```
### Determine how this performs on the test data
```
y_testp = gcv_results.predict(X_test)
from sklearn.metrics import classification_report
print(classification_report(y_test, y_testp))
```
### Did weighting help?
- Recall went from 0.36 to 0.92 for detecting the default.
- Precision went from 0.69 to 0.17 for detecting the default.
> We detected more of the defaults, with the trade-off of more false-positives. Depending what the cost of those is - which you'd need to talk with a SME about - this might be a preferred model.
```
from sklearn.metrics import roc_curve
y_testpr = gcv_results.predict_proba(X_test)
def generate_roc(y, probs):
fpr, tpr, _ = roc_curve(y, probs)
return fpr, tpr
fpr_wgt, tpr_wgt = generate_roc(y_test, y_testpr[:,1])
fpr_base, tpr_base = generate_roc(y_test, base_pr[:,1])
plt.plot(fpr_wgt, tpr_wgt,'-r')
plt.plot(fpr_base, tpr_base,'--b')
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.legend(['Weighted Model','Unweighted Model'])
plt.plot([0, 1], [0, 1], color='navy', linestyle='--')
plt.show()
```
> Weighted model has essentially the same ROC curve as the unweighted model in this case.
```
plt.barh(creditbase['data_processing'].get_feature_names_out(),creditbase['logreg'].coef_[0])
plt.barh(creditbase['data_processing'].get_feature_names_out(), gcv_results.best_estimator_['logreg'].coef_[0], alpha=0.5)
plt.legend(['Unweighted Coefficients', 'Weighted Coefficients'])
plt.show()
```
## Resampling and Generating new data
### Resampling.
If weighting isn't supporting, you can resample the training data.
- This gets complicated since you want to evaluate based on the original distribution.
- GridSearchCV won't support this well since the validation dataset is split from the training data.
- You could use loops for this, but you need to be careful to make sure the performance estimate is based on the `actual` distribution.
### Generating new data.
- The more complicated, probably less ROI option, is generating new data.
- SMOTE is a popular technique, if needed. [See this paper for a description](https://arxiv.org/pdf/1106.1813.pdf)
> There is another library called `imbalanced-learn` that has methods specifically designed for these types of problems as well.
# Using classifiers to determine dataset bias
- We shouldn't be able to predict whether an example in the training or test set.
```
from sklearn.datasets import load_iris
from sklearn.model_selection import train_test_split
iris = load_iris()
iX_train, iX_test, iy_train, iy_test = train_test_split(iris['data'],
iris['target'],
shuffle=True)
```
Creating a new label, whether included in the training or test set and that will be our new target.
```
inTraining = np.ones((iX_train.shape[0], 1))
inTest = np.zeros((iX_test.shape[0], 1))
irisTarget = np.append(inTraining, inTest, axis=0).reshape(-1)
irisTraining = np.append(iX_train, iX_test, axis=0)
from sklearn.linear_model import LogisticRegression
clf = LogisticRegression(class_weight='balanced')
clf = clf.fit(irisTraining, irisTarget)
preds = clf.predict(irisTraining)
from sklearn.metrics import confusion_matrix
confusion_matrix(irisTarget, preds)
from sklearn.dummy import DummyClassifier
dumdum = DummyClassifier(strategy='uniform')
dumdum = dumdum.fit(irisTraining, irisTarget)
dumPreds = dumdum.predict(irisTraining)
confusion_matrix(irisTarget, dumPreds)
```
Those are pretty close, so we would have confidence the test and training data is nearly identical if we were making a classifier.
# Readings
[Raschkas's Lecture](https://github.com/rasbt/stat479-machine-learning-fs19/blob/master/11_eval4-algo/11-eval4-algo__slides.pdf)
<br>[Full paper: Model Evaluation, Model Selection, and Algorithm Selection in Machine Learning, Raschka](https://arxiv.org/abs/1811.12808)
<br>[Evaluation: From Precision, Recall and F-Factor to ROC, Informedness, Markedness & Correlation](https://arxiv.org/abs/2010.16061)
| github_jupyter |
# Calculating GC Content
## GC Content
In molecular biology and genetics, GC-content (or guanine-cytosine content) is the percentage of nitrogenous bases in a DNA or RNA molecule that are either guanine (G) or cytosine (C). GC-content may be given for a certain fragment of DNA or RNA or for an entire genome.
GC-content may be given for a certain fragment of DNA or RNA or for an entire genome. When it refers to a fragment, it may denote the GC-content of an individual gene or section of a gene (domain), a group of genes or gene clusters, a non-coding region, or a synthetic oligonucleotide such as a primer.
[See More on Wikipedia](https://en.wikipedia.org/wiki/GC-content)
## Calculation
GC-content is usually expressed as a **percentage value**, but sometimes as a **ratio** (called G+C ratio or GC-ratio). GC-content percentage is calculated as
$ GC = \frac {G+C} {Len(DNA Sequence)} \times 100% $
## Applications
### Molecular biology
In polymerase chain reaction (PCR) experiments, the GC-content of short oligonucleotides known as **primers** is often used to predict their annealing temperature to the template DNA. A higher GC-content level indicates a relatively higher melting temperature.
### Systematics
The species problem in **prokaryotic** taxonomy has led to various suggestions in **classifying** bacteria, and the adhoc committee on reconciliation of approaches to bacterial systematics has recommended use of GC-ratios in higher-level hierarchical classification.For example, the Actinobacteria are characterised as "high GC-content bacteria".In Streptomyces coelicolor A3(2), GC-content is 72%
# Calculating GC Content
1. Download a DNA sequence from [NCBI](https://www.ncbi.nlm.nih.gov/)
2. Reading and cleaning the DNA sequence.
3. Create an algorithms for calculating **GC** content.
## Download a DNA Sequence from NCBI
You can download the dataset from [NCBI](https://www.ncbi.nlm.nih.gov/). Accession ID: NM_207618.2, Enter the accession id and click on **FASTA**, copy all the sequence without *>NM_207618.2 Mus musculus vomeronasal 1 receptor, D18 (V1rd18), mRNA*.then save it as **.txt** file.
## Reading and cleaning the DNA sequence
```
# Read the DNA sequence
with open("../data/dna.txt", "r") as f:
seq = f.read()
seq
```
The output is not a clean dataset because it contains a special character("\n"). It also may contain a special character("\n"). The special characters affect our analysis and may cause errors. That's why we have to clean the dataset by removing the special characters.
```
# Check the DNA sequence length
len(seq)
```
It is incorrect! because the original length of this DNA sequence is **1157**. The special characters 18. Let's clean the dataset.
```
# Replace "\n" character with empty space.
seq = seq.replace("\n", "")
seq
# Now check the length
len(seq)
```
YES! it is the original length of the DNA sequence. But we have to remove "\r"
```
seq = seq.replace("\r", "")
seq
len(seq)
```
## Create a Function To Read Sequence File
```
def read_seq(inputfile):
"""
Reads the DNA sequence file and returns as strings. Removes all the special characters.
"""
with open(inputfile, "r") as f:
seq = f.read()
seq = seq.replace("\n", "")
seq = seq.replace("r", "")
return seq
# Read the seq file with rea_seq function
dna = read_seq("../data/dna.txt")
dna
# Check length
len(dna)
```
## Calculate GC Content
```
def calculate_gc_content(seq):
"""
Take DNA sequence as input and calculate the GC content.
"""
no_of_g = seq.count("G")
no_of_c = seq.count("C")
total = no_of_g + no_of_c
gc = total/len(seq) * 100
return gc
calculate_gc_content(dna)
```
## Final Code
```
def read_seq(inputfile):
"""
Reads the DNA sequence file and returns as strings. Removes all the special characters.
"""
with open(inputfile, "r") as f:
seq = f.read()
seq = seq.replace("\n", "")
seq = seq.replace("r", "")
return seq
def calculate_gc_content(seq):
"""
Take DNA sequence as input and calculate the GC content.
"""
no_of_g = seq.count("G")
no_of_c = seq.count("C")
total = no_of_g + no_of_c
gc = total/len(seq) * 100
return gc
# Read the DNA sequence file
dna = read_seq("../data/dna.txt")
# Calculate GC Content
result = calculate_gc_content(dna)
print(result)
```
So the GC content is 39.5852% <br>
| github_jupyter |
# Frank-Wolfe Algorithm
By ZincCat
$\min _{\mathbf{x} \in \mathbb{R}^{N}}\|\mathbf{y}-\mathbf{D} \mathbf{x}\|_{2}^{2}$
s.t. $\|\mathbf{x}\|_{?} \leq 1$
```
import numpy as np
from matplotlib import pyplot as plt
p = 200
n = 300
np.random.seed(19890817)
D = np.random.normal(10, 5, (p, n))
y = np.random.normal(10, 5, p)
x0 = np.random.rand(n)
def f(x):
return np.linalg.norm(y-D@x)**2
def grad(x):
return 2*D.T@(D@x-y)
# 调用scipy求值
from scipy.optimize import minimize
cons1 = ({'type': 'ineq', 'fun': lambda x: 1 - np.linalg.norm(x, 1)})
res = minimize(f, x0, constraints=cons1, tol=1e-4, options={'maxiter': 1e4, 'disp': True})
minValue = res.fun
print("Scipy result:", res.fun)
# 调用scipy求值
from scipy.optimize import minimize
cons2 = ({'type': 'ineq', 'fun': lambda x: 1 - np.linalg.norm(x, np.inf)})
res = minimize(f, x0, constraints=cons2, tol=1e-10, options={'maxiter': 1e4, 'disp': True})
minValue2 = res.fun
print("Scipy result:", res.fun)
def fw(f, x, grad, maxIter, mode, log):
xn = x.copy()
for i in range(maxIter):
value = f(xn)
print(i, "th iteration, f(x)=", value)
log.append(value)
gamma = 2/(i+2)
g = grad(xn)
if mode == 1:
d = np.argmax(np.abs(g))
xn = (1-gamma)*xn
xn[d] -= gamma * np.sign(g[d])
elif mode == 'inf':
d = -np.sign(g)
xn += gamma * (d-xn)
return xn
```
$l_\infty$ constraint
```
maxIter = 3000000
linf = []
x = fw(f, x0/2/np.linalg.norm(x0, np.inf), grad, maxIter, 'inf', linf)
plt.plot(np.log(linf))
plt.xlabel('Iterations')
plt.ylabel('$\ln (f(x_k)-f^*)$')
plt.savefig('wolfeInf')
plt.show()
```
$l_1$ constraint
```
maxIter = 300000
l1 = []
x2 = fw(f, x0/2/np.linalg.norm(x0, 1), grad, maxIter, 1, l1)
plt.plot(np.log(l1-minValue))
plt.xlabel('Iterations')
plt.ylabel('$\ln (f(x_k)-f^*)$')
plt.savefig('wolfe1')
plt.show()
```
projected gradient descent
```
def P1(x):
norm = np.linalg.norm(x, 1)
if norm > 1:
return x/norm
return x
def Pinf(x):
t = np.minimum(x, np.ones(n))
return np.maximum(t, -np.ones(n))
def linesearch_Armijo(f, x, g, d, alpha=0.4, beta=0.8):
# backtrack linesearch using Armijo rules
t = 10.0
value = f(x)
while f(x + t*d) > value + alpha*t*np.dot(g, d):
t *= beta
return t
def projectedDescent(f, x, grad, proj):
# 投影梯度下降函数
# 输入函数f, 目前x取值, 梯度函数, 要投影到的矩阵A
# 输出下降后x取值, 步长t
xn = x.copy()
g = grad(xn)
grad_norm = np.linalg.norm(g, 2)
d = -g/grad_norm
t = linesearch_Armijo(f, xn, g, d)
xn += t*d
return proj(xn), t
# 绘图
time1 = [] # 记录时间步, 用于绘图
values1 = [] # 记录某一时间步下函数值, 用于绘图
Plot = True # 是否绘图, 请保证此时alpha, beta均为单一取值
timestep = 0
x = x0.copy() #满足约束的初值
# 用于判定终止
count = 0
eps = 1e-13
oldvalue = f(x)
maxIter = 200000 # 最大迭代次数
while True:
value = f(x)
print("Iteration:", timestep, "Value", value)
# 用函数值改变量作为终止条件
if abs(value - oldvalue) < eps:
count += 1
else:
count = 0
oldvalue = value
if Plot:
time1.append(timestep)
values1.append(value)
if timestep > maxIter or count >= 5:
break
x, t = projectedDescent(f, x, grad, Pinf) # 此时使用无穷范数
timestep += 1
```
| github_jupyter |
```
import nltk
from nltk.corpus import wordnet as wn
import string
import itertools
from nltk.corpus import wordnet_ic
import codecs
import pickle
import Stemmer
import datetime
def load_MSRP_corpus(path, ftype):
"""Returns the tuples in the MSRP Corus, which has the following format
class #1_ID #2_ID #1_String #2_String
"""
if ftype == "train":
fpath = (path+"msr_paraphrase_train.txt"
if path[-1] == '/' or path[-1] == '\\'
else path+"/msr_paraphrase_train.txt")
elif ftype == "test":
fpath = (path+"msr_paraphrase_test.txt"
if path[-1] == '/' or path[-1] == '\\'
else path+"/msr_paraphrase_test.txt")
pairs = []
with codecs.open(fpath, 'r', encoding='utf-8') as fid:
for line in list(fid)[1:]:
pairs.append(line.split('\t'))
return pairs
def word_tokenizer(text, stemmer, flat_output=True):
"""Returns the text divided by sentences and tokenized.
If flat_output is True, returns
The format of the results is (token, lemma, stem, tag)"""
# Methods for sentence splitting, tokenization and lemmatization
sent_detector = nltk.data.load('tokenizers/punkt/english.pickle')
word_detector = nltk.TreebankWordTokenizer()
wnl = nltk.stem.WordNetLemmatizer()
tags_mapping = {"NN":wn.NOUN, "VB":wn.VERB, "JJ":wn.ADJ, "RB":wn.ADV}
# Text processing
output = []
for sent in sent_detector.tokenize(text.lower()):
tokens = word_detector.tokenize(sent)
tmp = []
for token, tag in nltk.pos_tag(tokens):
wn_pos = tags_mapping.get(tag[:2], None)
if wn_pos:
tmp.append((token, wnl.lemmatize(token, pos=wn_pos), stemmer.stemWord(token), tag))
else:
tmp.append((token, token, stemmer.stemWord(token), tag))
output.append(tmp)
if flat_output:
return list(itertools.chain(*output))
return output
```
#### Parsing training instances
We parse both sentences in each instance of the MSPR training dataset in the format (token, lemma, POS_tag). It might be divided in sentences as well depending on the flag **flat_output** being false.
tags_mapping maps Treebank POS tags to WordNet POS tags.
```
stemmer = Stemmer.Stemmer('english')
raw_data = load_MSRP_corpus(path="D:/Corpus/MSPC", ftype="train")
parsed_texts = {}
train_pairs = []
train_y = []
for _class, id1, id2, text1, text2 in raw_data:
if id1 not in parsed_texts:
parsed_texts[int(id1)] = word_tokenizer(text1, stemmer=stemmer, flat_output=True)
if id2 not in parsed_texts:
parsed_texts[int(id2)] = word_tokenizer(text2, stemmer=stemmer, flat_output=True)
train_pairs.append((int(id1),int(id2)))
train_y.append(int(_class))
```
#### Parsing test instances
We parse both sentences in each instance of the MSPR test dataset in the format (token, lemma, POS_tag). It might be divided in sentences as well depending on the flag **flat_output** being false.
```
raw_data = load_MSRP_corpus(path="D:/Corpus/MSPC", ftype="test")
test_pairs = []
test_y = []
for _class, id1, id2, text1, text2 in raw_data:
if id1 not in parsed_texts:
parsed_texts[int(id1)] = word_tokenizer(text1, stemmer=stemmer, flat_output=True)
if id2 not in parsed_texts:
parsed_texts[int(id2)] = word_tokenizer(text2, stemmer=stemmer, flat_output=True)
test_pairs.append((int(id1),int(id2)))
test_y.append(int(_class))
```
#### Generating vocabulary
The vocabulary is generated taking into account the lemma and wordnet tag (if possible).
Here, we can perform several experiments like taking the lemma and regular POS tag, only taking the tokens, etc.
**Notes:**
- The code depends on the format used in parsed_texts, assuming **flat_output=True**
```
tags_mapping = {"NN":wn.NOUN, "VB":wn.VERB, "JJ":wn.ADJ, "RB":wn.ADV}
# itertools.chain(*parsed_texts.values()) converts all documents into one single list
vocab_tokens = nltk.FreqDist([token
for token, lemma, stem, tag
in itertools.chain(*parsed_texts.values())])
vocab_lemmas = nltk.FreqDist([lemma
for token, lemma, stem, tag
in itertools.chain(*parsed_texts.values())])
vocab_stems = nltk.FreqDist([stem
for token, lemma, stem, tag
in itertools.chain(*parsed_texts.values())])
vocab_lemmapos = nltk.FreqDist([(lemma, tags_mapping.get(tag[:2], tag))
for token, lemma, stem, tag
in itertools.chain(*parsed_texts.values())])
print(len(vocab_tokens), "tokens in the vocabulary of tokens")
print(len(vocab_lemmas), "tokens in the vocabulary of lemmas")
print(len(vocab_stems), "tokens in the vocabulary of stems")
print(len(vocab_lemmapos), "tokens in the vocabulary of lemmas+pos")
index2token = list(vocab_tokens.keys())
token2index = dict([(token,i) for i, token in enumerate(index2token)])
index2lemma = list(vocab_lemmas.keys())
lemma2index = dict([(lemma,i) for i, lemma in enumerate(index2lemma)])
index2stem = list(vocab_stems.keys())
stem2index = dict([(stem,i) for i, stem in enumerate(index2stem)])
index2lemmapos = list(vocab_lemmapos.keys())
lemmapos2index = dict([(lemmapos,i) for i, lemmapos in enumerate(index2lemmapos)])
```
#### Serializing all MSRPC and needed data structures
We serialize the Microsoft Research Paraphrase Corpus with the format:
[parsed_texts, train_pairs, train_y, test_pairs, test_y]
The goal of serializing all of these is to make experiments with the parsed corpus used to compute the WordNet similarity measures.
<parsed_texts> contains all the documents from the MSRPC tokenized. It is a dictionary where the keys are the id of the documents and the values are lists of the following tuples (token, lemma, POS tag)
We also serialize the vocabularies for different features (tokens, lemmas, stems, and lemmas+pos) and the data structure needed to convert from indexes to the given feature and viceversa.
[vocab, token_to_index, index_to_token]
```
date_obj = datetime.date.today()
date_str = "{:04d}".format(date_obj.year) + "{:02d}".format(date_obj.month) + "{:02d}".format(date_obj.day)
pickle.dump([parsed_texts, train_pairs, train_y, test_pairs, test_y],
open("msrpc_parsed_"+date_str+".pickle", "wb"),
protocol=pickle.HIGHEST_PROTOCOL)
pickle.dump([vocab_tokens, token2index, index2token],
open("tokens_data_"+date_str+".pickle", "wb"),
protocol=pickle.HIGHEST_PROTOCOL)
pickle.dump([vocab_lemmas, lemma2index, index2lemma],
open("lemmas_data_"+date_str+".pickle", "wb"),
protocol=pickle.HIGHEST_PROTOCOL)
pickle.dump([vocab_stems, stem2index, index2stem],
open("stems_data_"+date_str+".pickle", "wb"),
protocol=pickle.HIGHEST_PROTOCOL)
pickle.dump([vocab_lemmapos, lemmapos2index, index2lemmapos],
open("lemmapos_data_"+date_str+".pickle", "wb"),
protocol=pickle.HIGHEST_PROTOCOL)
```
#### Computing some Information Content (IC)
```
brown_ic = wordnet_ic.ic('ic-brown.dat')
semcor_ic = wordnet_ic.ic('ic-semcor.dat')
bnc_ic_2000 = wordnet_ic.ic('ic-bnc.dat')
bnc_ic_2007 = wordnet_ic.ic('ic-bnc-2007.dat')
```
**Note:** This code is not in use!
***Warning*** Computing BNC Information Content (IC) takes hours!!!
There is a version of the IC for the BNC corpus. However it was computed using the 2000 version of BNC. The corpus we are computing here is the version of 2007. The similiarty measures using these two IC are different.
reader_bnc = nltk.corpus.reader.BNCCorpusReader(root='D:/Corpus/2554/2554/download/Texts/', fileids=r'[A-K]/\w*/\w*\.xml')
bnc_ic = wn.ic(reader_bnc, False, 0.0)
def is_root(synset_x):
if synset_x.root_hypernyms()[0] == synset_x:
return True
return False
def generate_ic_file(IC, output_filename):
"""Dump in output_filename the IC counts.
The expected format of IC is a dict
{'v':defaultdict, 'n':defaultdict, 'a':defaultdict, 'r':defaultdict}"""
with codecs.open(output_filename, 'w', encoding='utf-8') as fid:
# Hash code of WordNet 3.0
fid.write("wnver::eOS9lXC6GvMWznF1wkZofDdtbBU"+"\n")
# We only stored nouns and verbs because those are the only POS tags
# supported by wordnet.ic() function
for tag_type in ['v', 'n']:#IC:
for key, value in IC[tag_type].items():
if key != 0:
synset_x = wn.of2ss(of="{:08d}".format(key)+tag_type)
if is_root(synset_x):
fid.write(str(key)+tag_type+" "+str(value)+" ROOT\n")
else:
fid.write(str(key)+tag_type+" "+str(value)+"\n")
print("Done")
generate_ic_file(bnc_ic, "C:/Users/MiguelAngel/AppData/Roaming/nltk_data/corpora/wordnet_ic/ic-bnc-2007.dat")
#### Examples applying WordNet similarity measures and using different IC
```
syns = wn.synsets(lemma="car", pos='n')
print(syns)
syns[0]
def nan_default(result):
if result:
return result
return 0.0
print("{:40} {:.2f}".format("Path Similarity:", nan_default(wn.path_similarity(syns[0], syns[1]))))
print("{:40} {:.2f}".format("Leacock-Chodorow Similarity:", nan_default(wn.lch_similarity(syns[0], syns[1]))))
print("{:40} {:.2f}".format("Wu-Palmer Similarity:", nan_default(wn.wup_similarity(syns[0], syns[1]))))
print("{:40} {:.2f}".format("Resnik Similarity (brown_ic):", nan_default(wn.res_similarity(syns[0], syns[1], brown_ic))))
print("{:40} {:.2f}".format("Resnik Similarity (semcor_ic):", nan_default(wn.res_similarity(syns[0], syns[1], semcor_ic))))
print("{:40} {:.2f}".format("Resnik Similarity (bnc_ic):", nan_default(wn.res_similarity(syns[0], syns[1], bnc_ic_2007))))
print("{:40} {:.2f}".format("Jiang-Conrath Similarity (brown_ic):", nan_default(wn.jcn_similarity(syns[0], syns[1], brown_ic))))
print("{:40} {:.2f}".format("Jiang-Conrath Similarity (semcor_ic):", nan_default(wn.jcn_similarity(syns[0], syns[1], semcor_ic))))
print("{:40} {:.2f}".format("Jiang-Conrath Similarity (bnc_ic):", nan_default(wn.jcn_similarity(syns[0], syns[1], bnc_ic_2007))))
print("{:40} {:.2f}".format("Lin Similarity (brown_ic):", nan_default(wn.lin_similarity(syns[0], syns[1], brown_ic))))
print("{:40} {:.2f}".format("Lin Similarity (semcor_ic):", nan_default(wn.lin_similarity(syns[0], syns[1], semcor_ic))))
print("{:40} {:.2f}".format("Lin Similarity (bnc_ic):", nan_default(wn.lin_similarity(syns[0], syns[1], bnc_ic_2007))))
```
### Computing all the similarities needed in the MSRPC
We only care about the following POS tags: NN, VB, RB, JJ. Note: The WordNet authors divide the adjectives in two types: ('a' - Adjevtive) and ('s' - Adjective Satellite)
The dictionary *tags_mapping** maps Treebank POS to WordNet POS.
We compute the similarities for all words in phrase1 to all words in phrase2 that have the same POS tag. Also, for each pair of words we compute the similarity between all the words synsents. During testing we can decide if we use the first synset or the max similarity between all synsets.
```
def wordnet_similarity(s1, s2, metric="Path", ic=None):
# Computing the Path similarity
# This measure ranges between 0 and 1, not normalization needed
if metric == "path":
if s1.pos() == s2.pos():
# path_similarity returns None if no connecting path could be found
# between s1 and s2. We convert None to 0.0
return nan_default(wn.path_similarity(s1, s2))
else:
return 0.0
# Computing the Leacock-Chodorow similarity
# This measure range is unknown, normalization needed!!!**********
if metric == "lch":
if s1.pos() == s2.pos():
# path_similarity returns None if no connecting path could be found
# between s1 and s2. We convert None to 0.0
return nan_default(wn.lch_similarity(s1, s2))
else:
return 0.0
# Computing the Wu-Palmer similarity
# This measure range is unknown, normalization needed!!!**********
# path_similarity returns None if no connecting path could be found
# between s1 and s2. We convert None to 0.0
if metric == "wup":
if s1.pos() == s2.pos():
return nan_default(wn.wup_similarity(s1, s2))
else:
return 0.0
# Computing the Resnik similarity
# This measure range is unknown, normalization needed!!!**********
# Only comparing verbs and nouns because those are the only keys in IC
if metric == "res":
if s1.pos() == s2.pos() and s1.pos() in ['v', 'n']:
return wn.res_similarity(s1, s2, ic)
else:
return 0.0
# Computing the Jiang-Conrath similarity
# This measure range is unknown, normalization needed!!!**********
# Only comparing verbs and nouns because those are the only keys in IC
if metric == "jcn":
if s1.pos() == s2.pos() and s1.pos() in ['v', 'n']:
return wn.jcn_similarity(s1, s2, ic)
else:
return 0.0
# Computing the Lin similarity
# This measure range is unknown, normalization needed!!!**********
# Only comparing verbs and nouns because those are the only keys in IC
if metric == "lin":
try:
if s1 == s2:
return 1.0
if s1.pos() == s2.pos() and s1.pos() in ['v', 'n']:
return wn.lin_similarity(s1, s2, ic)
else:
return 0.0
except:
print("Synsets causing the exception")
print(s1, s2)
raise
print("No available metric selected. Raising an error.")
raise
return None
def compute_msrpc_wordnet_sims_tokens(pairs, vocab, token2index, tags_mapping,
sims_first_synset, sims_all_synsets, metric, ic):
"""Compute the WordNet <metric> similarity for all the pairs of tokens needed in the MSRPCorpus
using only the given tokens to extract the wordnet synsets"""
count_first = 0
count_max = 0
# For each pair of phrase in training dataset
for id1, id2 in pairs:
# For each token in first phrase
for token1, lemma1, stem1, tag1 in parsed_texts[id1]:
token1_id = token2index.get(token1, None)
if token1_id == None:
print("Error, token1 not in vocabulary", token1)
raise
# Extract the synsets from first phrase tokens
syns_tk1 = wn.synsets(lemma=token1)
# Compare to each token in phrase 2
for token2, lemma2, stem2, tag2 in parsed_texts[id2]:
# index of token2 in the vocabulary
token2_id = token2index.get(token2, None)
if token2_id == None:
print("Error, token2 not in vocabulary")
raise
# Extract the synsets from second prhase tokens
syns_tk2 = wn.synsets(lemma=token2)
# Validating we have not computed this similarity before
# *****************
# *** IMPORTANT *** We need to apply the same method when retrieving similarity scores***
# *****************
sim_key = (token1_id, token2_id) if token1_id > token2_id else (token2_id, token1_id)
# Must be different tokens
if sim_key[0] != sim_key[1] and len(syns_tk1)*len(syns_tk2) != 0 and sim_key not in sims_all_synsets:
all_sims = []
#sims_first_synset[sim_key] = 0.0
#sims_all_synsets[sim_key] = 0.0
for s1 in syns_tk1:
for s2 in syns_tk2:
# The similarity must be different of None and Zero
res = wordnet_similarity(s1, s2, metric=metric, ic=ic)
if res != 0.0:
all_sims.append(res)
if len(all_sims) > 0:
sims_first_synset[sim_key] = all_sims[0]
sims_all_synsets[sim_key] = max(all_sims)
count_max += len(syns_tk1)*len(syns_tk2)
count_first += 0 if len(syns_tk1)*len(syns_tk2) == 0 else 1
# For debugging. Stopping early.
#if count_first > 1:
# break
# Finish debugging
print(count_first, "new token pairs computed.")
print(count_max, "Wordnet similarity computed.")
return None
def compute_msrpc_wordnet_sims_lemmas(pairs, vocab, lemma2index, tags_mapping,
sims_first_synset, sims_all_synsets, metric, ic):
"""Compute the WordNet <metric> similarity for all the pairs of tokens needed in the MSRPCorpus
using only the lemmas (NO POS tags) of the given tokens to extract the wordnet synsets"""
count_first = 0
count_max = 0
# For each pair of phrase in training dataset
for id1, id2 in pairs:
# For each token in first phrase
for token1, lemma1, stem1, tag1 in parsed_texts[id1]:
token1_id = lemma2index.get(lemma1, None)
if token1_id == None:
print("Error, lemma1 not in vocabulary", lemma1)
raise
# Extract the synsets from first phrase tokens
syns_tk1 = wn.synsets(lemma=lemma1)
# Compare to each token in phrase 2
for token2, lemma2, stem2, tag2 in parsed_texts[id2]:
# index of token2 in the vocabulary
token2_id = lemma2index.get(lemma2, None)
if token2_id == None:
print("Error, lemma2 not in vocabulary")
raise
# Extract the synsets from second prhase tokens
syns_tk2 = wn.synsets(lemma=lemma2)
# Validating we have not computed this similarity before
# *****************
# *** IMPORTANT *** We need to apply the same method when retrieving similarity scores***
# *****************
sim_key = (token1_id, token2_id) if token1_id > token2_id else (token2_id, token1_id)
# Must be different tokens
if sim_key[0] != sim_key[1] and len(syns_tk1)*len(syns_tk2) != 0 and sim_key not in sims_all_synsets:
all_sims = []
#sims_first_synset[sim_key] = 0.0
#sims_all_synsets[sim_key] = 0.0
for s1 in syns_tk1:
for s2 in syns_tk2:
# The similarity must be different of None and Zero
res = wordnet_similarity(s1, s2, metric=metric, ic=ic)
if res != 0.0:
all_sims.append(res)
if len(all_sims) > 0:
sims_first_synset[sim_key] = all_sims[0]
sims_all_synsets[sim_key] = max(all_sims)
count_max += len(syns_tk1)*len(syns_tk2)
count_first += 0 if len(syns_tk1)*len(syns_tk2) == 0 else 1
# For debugging. Stopping early.
#if count_first > 1:
# break
# Finish debugging
print(count_first, "new token pairs computed.")
print(count_max, "Wordnet similarity computed.")
return None
def compute_msrpc_wordnet_sims_lemmaspos(pairs, vocab, lemmapos2index, tags_mapping,
sims_first_synset, sims_all_synsets, metric, ic):
"""Compute the WordNet <metric> similarity for all the pairs of tokens needed in the MSRPCorpus
using the lemmas and POS tags of the given tokens to extract the wordnet synsets"""
count_first = 0
count_max = 0
# For each pair of phrase in training dataset
for id1, id2 in pairs:
# For each token in first phrase
for token1, lemma1, stem1, tag1 in parsed_texts[id1]:
# Only look at WordNet POS tags
wn_tag1 = tags_mapping.get(tag1[:2], None)
if wn_tag1:
# index of token1 in the vocabulary
token1_id = lemmapos2index.get((lemma1, wn_tag1), None)
if token1_id == None:
print("Error, lemma1 not in vocabulary", lemma1)
raise
# Extract the synsets from first phrase tokens
syns_tk1 = wn.synsets(lemma=lemma1, pos=wn_tag1)
# Compare to each token in phrase 2
for token2, lemma2, stem2, tag2 in parsed_texts[id2]:
# Only look at WordNet POS tags
wn_tag2 = tags_mapping.get(tag2[:2], None)
# Compute similarity of those tokens with the same POS tag
if wn_tag2 and wn_tag1 == wn_tag2:
# index of token2 in the vocabulary
token2_id = lemmapos2index.get((lemma2, wn_tag2), None)
if token2_id == None:
print("Error, lemma2 not in vocabulary")
raise
# Extract the synsets from second prhase tokens
syns_tk2 = wn.synsets(lemma=lemma2, pos=wn_tag2)
# Validating we have not computed this similarity before
# *****************
# *** IMPORTANT *** We need to apply the same method when retrieving similarity scores***
# *****************
sim_key = (token1_id, token2_id) if token1_id > token2_id else (token2_id, token1_id)
# Must be different tokens
if sim_key[0] != sim_key[1] and len(syns_tk1)*len(syns_tk2) != 0 and sim_key not in sims_all_synsets:
all_sims = []
#sims_first_synset[sim_key] = 0.0
#sims_all_synsets[sim_key] = 0.0
for s1 in syns_tk1:
for s2 in syns_tk2:
# The similarity must be different of None and Zero
res = wordnet_similarity(s1, s2, metric=metric, ic=ic)
if res != 0.0:
all_sims.append(res)
if len(all_sims) > 0:
sims_first_synset[sim_key] = all_sims[0]
sims_all_synsets[sim_key] = max(all_sims)
count_max += len(syns_tk1)*len(syns_tk2)
count_first += 0 if len(syns_tk1)*len(syns_tk2) == 0 else 1
# For debugging. Stopping early.
#if count_first > 1:
# break
# Finish debugging
print(count_first, "new token pairs computed.")
print(count_max, "Wordnet similarity computed.")
return None
```
### WordNet similarity
path, lch, wup, res, jcn, lin
```
for ic_data, ic_name in zip([bnc_ic_2000, semcor_ic, brown_ic], ['bnc_ic_2000', 'semcor_ic', 'brown_ic']):
for metric in ['lin', 'jcn', 'res']:#['lin', 'jcn', 'res', 'path', 'lch', 'wup']:
tags_mapping = {"NN":wn.NOUN, "VB":wn.VERB, "JJ":wn.ADJ, "RB":wn.ADV}
date_obj = datetime.date.today()
date_str = "{:04d}".format(date_obj.year) + "{:02d}".format(date_obj.month) + "{:02d}".format(date_obj.day)
print("METRIC:", metric)
#***********************************************
print("Using only tokens")
print("Training dataset")
sims_first_synset = {}
sims_all_synsets = {}
compute_msrpc_wordnet_sims_tokens(train_pairs, vocab_tokens, token2index, tags_mapping,
sims_first_synset, sims_all_synsets, metric, ic_data)#bnc_ic_2007)
print("Test dataset")
compute_msrpc_wordnet_sims_tokens(test_pairs, vocab_tokens, token2index, tags_mapping,
sims_first_synset, sims_all_synsets, metric, ic_data)#bnc_ic_2007)
print("Total pairs:", len(sims_first_synset))
pickle.dump(sims_first_synset, open(metric+"_first_tokens_"+ic_name+"_"+date_str+".pickle", "wb"), protocol=pickle.HIGHEST_PROTOCOL)
pickle.dump(sims_all_synsets, open(metric+"_all_tokens_"+ic_name+"_"+date_str+".pickle", "wb"), protocol=pickle.HIGHEST_PROTOCOL)
#***********************************************
#***********************************************
print("Using only lemmas")
print("Training dataset")
sims_first_synset = {}
sims_all_synsets = {}
compute_msrpc_wordnet_sims_lemmas(train_pairs, vocab_lemmas, lemma2index, tags_mapping,
sims_first_synset, sims_all_synsets, metric, ic_data)#bnc_ic_2007)
print("Test dataset")
compute_msrpc_wordnet_sims_lemmas(test_pairs, vocab_lemmas, lemma2index, tags_mapping,
sims_first_synset, sims_all_synsets, metric, ic_data)#bnc_ic_2007)
print("Total pairs:", len(sims_first_synset))
pickle.dump(sims_first_synset, open(metric+"_first_lemmas_"+ic_name+"_"+date_str+".pickle", "wb"), protocol=pickle.HIGHEST_PROTOCOL)
pickle.dump(sims_all_synsets, open(metric+"_all_lemmas_"+ic_name+"_"+date_str+".pickle", "wb"), protocol=pickle.HIGHEST_PROTOCOL)
#***********************************************
#***********************************************
print("Using lemmas+pos")
print("Training dataset")
sims_first_synset = {}
sims_all_synsets = {}
compute_msrpc_wordnet_sims_lemmaspos(train_pairs, vocab_lemmapos, lemmapos2index, tags_mapping,
sims_first_synset, sims_all_synsets, metric, ic_data)#bnc_ic_2007)
print("Test dataset")
compute_msrpc_wordnet_sims_lemmaspos(test_pairs, vocab_lemmapos, lemmapos2index, tags_mapping,
sims_first_synset, sims_all_synsets, metric, ic_data)#bnc_ic_2007)
print("Total pairs:", len(sims_first_synset))
pickle.dump(sims_first_synset, open(metric+"_first_lemmapos_"+ic_name+"_"+date_str+".pickle", "wb"), protocol=pickle.HIGHEST_PROTOCOL)
pickle.dump(sims_all_synsets, open(metric+"_all_lemmapos_"+ic_name+"_"+date_str+".pickle", "wb"), protocol=pickle.HIGHEST_PROTOCOL)
#***********************************************
print("\n")
s = wn.synsets("hit", "n")[0]
import nltk.corpus.reader.wordnet as wn2
wn2._lcs_ic(s, s,bnc_ic_2007)
wn.jcn_similarity(s, s, bnc_ic_2007)
wn.res_similarity(s, s, bnc_ic_2007)
wn.lin_similarity(s, s, bnc_ic_2007)
wn2._INF
```
##### Serializing the WordNet similarity scores
```
pickle.dump(sims_first_synset, open(metric+"_first_"+date+".pickle", "wb"), protocol=pickle.HIGHEST_PROTOCOL)
pickle.dump(sims_all_synsets, open(metric+"_all_"+date+".pickle", "wb"), protocol=pickle.HIGHEST_PROTOCOL)
```
#### Computing the WordNet similarity scores of the vocabulary terms
We compute these scores in order to normalize the similarity measures that do not have a range between 0 and 1.
path, lch, wup, res, jcn, lin
```
for metric in ['path', 'lch', 'wup', 'res', 'jcn', 'lin']:
print(metric)
wn_tags = set(['v','n','r','a'])
vocab_sim = {}
#metric = "lch"
# The tag is already a wordnet tag where it is one of the wn_tags
for lemma, tag in vocab:
idx = token_to_index[(lemma, tag)]
if tag in wn_tags:
syns_tk = wn.synsets(lemma=lemma, pos=tag)
all_sims = []
for s in syns_tk:
try:
res = wordnet_similarity(s, s, metric=metric, ic=bnc_ic_2000)
if res > 0.0:
all_sims.append(res)
except ZeroDivisionError:
pass
if len(all_sims) > 0:
vocab_sim[idx] = max(all_sims)
print("Vocab size:", len(vocab))
for pos in ['n','v','r','a']:
print(pos)
non_zero = [x for idx, x in vocab_sim.items() if x > 0.0 and index_to_token[idx][1]==pos]
print("Sims computed:", len(vocab_sim))
print("Max:", max(non_zero, default=-1))
print("Min", min(non_zero, default=-1))
print("Mean:", -1 if len(non_zero) == 0 else sum(non_zero)/len(non_zero))
pickle.dump(vocab_sim, open("vocab_"+metric+"_"+date+".pickle", "wb"), protocol=pickle.HIGHEST_PROTOCOL)
print()
```
WordNet similarity ranges:
- path: 0.0 - 1.0
- lch:
- nouns: 0.0 - 3.6375861597263857
- verbs: 0.0 - 3.258096538021482
- wup: 0.0 - 1.0
- res: 0.0 - 1e+300 **This number (inf) is too big. Need to check the maximum value in the similarity file**
- jcn: 0.0 - 1e+300 **This number (inf) is too big. Need to check the maximum value in the similarity file**
- lin: 0.0 - 1.0
**Need to check that all the similarities computed fall within these ranges**
**Questions**
- Shall we generate an index for the words or not? **Yes. Done!**
- Shall we remove the tokens composed by only punctuation symbols? **Yes. Done!**
- Shall we replace the numbers by a unique identifier?
- What is the range of each WordNet similarity measure? **Done for our specific corpus!**
- How to serialize the Information Count from British National Corpus (BNC)? **I implemented it manually. Done!**
**Tasks**
- Define the similarities needed in both datasets. **Done!**
- Run example for all WordNet similarity measures. **Done!**
- Serialize all the similarities needed in both, training and test dataset for each of the WordNet similarities. Try with first synset and with max of all synsets. **Done!**
- Save the values without normalization. We need to know the ranges of the WordNet metrics or the maximum value to normalize. **Done for MSRPC!**
- When serializing the WordNet similarity values, we also need to serialize the vocabulary, token_to_index, and index_to_token data structures. **Done**
**Notes**
- The WordNet metrics that need normalization, because their values are not in the 0-1 range, won't be easily normalized because there is not a score for the similarity between the same lemmas other than Inf.
- An idea to normalize is compute the similarity of a large amount of words between themself and pick the highest value.
#### Computing IDF over the BNC Corpus
```
def compute_idf(corpus):
df = {}
idf = {}
for fid in corpus.fileids():
words = set([token.lower() for token in corpus.words(fid)])
for word in words:
df[word] = df.get(word, 0) + 1
N = len(corpus.fileids())
for word in df:
idf[word] = np.log(N/df[word])
return idf
#corpus = nltk.corpus.brown
# BNC 2007 XML version
corpus = nltk.corpus.reader.BNCCorpusReader(root='D:/Corpus/2554/2554/download/Texts/',
fileids=r'[A-K]/\w*/\w*.xml')
idf = compute_idf(corpus)
len(idf)
pickle.dump(idf, open("idf_20170810.pickle", "wb"), protocol=pickle.HIGHEST_PROTOCOL)
```
| github_jupyter |
# Training Neural Networks
The network we built in the previous part isn't so smart, it doesn't know anything about our handwritten digits. Neural networks with non-linear activations work like universal function approximators. There is some function that maps your input to the output. For example, images of handwritten digits to class probabilities. The power of neural networks is that we can train them to approximate this function, and basically any function given enough data and compute time.
<img src="assets/function_approx.png" width=500px>
At first the network is naive, it doesn't know the function mapping the inputs to the outputs. We train the network by showing it examples of real data, then adjusting the network parameters such that it approximates this function.
To find these parameters, we need to know how poorly the network is predicting the real outputs. For this we calculate a **loss function** (also called the cost), a measure of our prediction error. For example, the mean squared loss is often used in regression and binary classification problems
$$
\large \ell = \frac{1}{2n}\sum_i^n{\left(y_i - \hat{y}_i\right)^2}
$$
where $n$ is the number of training examples, $y_i$ are the true labels, and $\hat{y}_i$ are the predicted labels.
By minimizing this loss with respect to the network parameters, we can find configurations where the loss is at a minimum and the network is able to predict the correct labels with high accuracy. We find this minimum using a process called **gradient descent**. The gradient is the slope of the loss function and points in the direction of fastest change. To get to the minimum in the least amount of time, we then want to follow the gradient (downwards). You can think of this like descending a mountain by following the steepest slope to the base.
<img src='assets/gradient_descent.png' width=350px>
## Backpropagation
For single layer networks, gradient descent is straightforward to implement. However, it's more complicated for deeper, multilayer neural networks like the one we've built. Complicated enough that it took about 30 years before researchers figured out how to train multilayer networks.
Training multilayer networks is done through **backpropagation** which is really just an application of the chain rule from calculus. It's easiest to understand if we convert a two layer network into a graph representation.
<img src='assets/backprop_diagram.png' width=550px>
In the forward pass through the network, our data and operations go from bottom to top here. We pass the input $x$ through a linear transformation $L_1$ with weights $W_1$ and biases $b_1$. The output then goes through the sigmoid operation $S$ and another linear transformation $L_2$. Finally we calculate the loss $\ell$. We use the loss as a measure of how bad the network's predictions are. The goal then is to adjust the weights and biases to minimize the loss.
To train the weights with gradient descent, we propagate the gradient of the loss backwards through the network. Each operation has some gradient between the inputs and outputs. As we send the gradients backwards, we multiply the incoming gradient with the gradient for the operation. Mathematically, this is really just calculating the gradient of the loss with respect to the weights using the chain rule.
$$
\large \frac{\partial \ell}{\partial W_1} = \frac{\partial L_1}{\partial W_1} \frac{\partial S}{\partial L_1} \frac{\partial L_2}{\partial S} \frac{\partial \ell}{\partial L_2}
$$
**Note:** I'm glossing over a few details here that require some knowledge of vector calculus, but they aren't necessary to understand what's going on.
We update our weights using this gradient with some learning rate $\alpha$.
$$
\large W^\prime_1 = W_1 - \alpha \frac{\partial \ell}{\partial W_1}
$$
The learning rate $\alpha$ is set such that the weight update steps are small enough that the iterative method settles in a minimum.
## Losses in PyTorch
Let's start by seeing how we calculate the loss with PyTorch. Through the `nn` module, PyTorch provides losses such as the cross-entropy loss (`nn.CrossEntropyLoss`). You'll usually see the loss assigned to `criterion`. As noted in the last part, with a classification problem such as MNIST, we're using the softmax function to predict class probabilities. With a softmax output, you want to use cross-entropy as the loss. To actually calculate the loss, you first define the criterion then pass in the output of your network and the correct labels.
Something really important to note here. Looking at [the documentation for `nn.CrossEntropyLoss`](https://pytorch.org/docs/stable/nn.html#torch.nn.CrossEntropyLoss),
> This criterion combines `nn.LogSoftmax()` and `nn.NLLLoss()` in one single class.
>
> The input is expected to contain scores for each class.
This means we need to pass in the raw output of our network into the loss, not the output of the softmax function. This raw output is usually called the *logits* or *scores*. We use the logits because softmax gives you probabilities which will often be very close to zero or one but floating-point numbers can't accurately represent values near zero or one ([read more here](https://docs.python.org/3/tutorial/floatingpoint.html)). It's usually best to avoid doing calculations with probabilities, typically we use log-probabilities.
```
import torch
from torch import nn
import torch.nn.functional as F
from torchvision import datasets, transforms
# Define a transform to normalize the data
transform = transforms.Compose([transforms.ToTensor(),
transforms.Normalize((0.5,), (0.5,)),
])
# Download and load the training data
trainset = datasets.MNIST('~/.pytorch/MNIST_data/', download=True, train=True, transform=transform)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=64, shuffle=True)
# Build a feed-forward network
model = nn.Sequential(nn.Linear(784, 128),
nn.ReLU(),
nn.Linear(128, 64),
nn.ReLU(),
nn.Linear(64, 10))
# Define the loss
criterion = nn.CrossEntropyLoss()
# Get our data
images, labels = next(iter(trainloader))
# Flatten images
images = images.view(images.shape[0], -1)
# Forward pass, get our logits
logits = model(images)
# Calculate the loss with the logits and the labels
loss = criterion(logits, labels)
print(loss)
```
In my experience it's more convenient to build the model with a log-softmax output using `nn.LogSoftmax` or `F.log_softmax` ([documentation](https://pytorch.org/docs/stable/nn.html#torch.nn.LogSoftmax)). Then you can get the actual probabilites by taking the exponential `torch.exp(output)`. With a log-softmax output, you want to use the negative log likelihood loss, `nn.NLLLoss` ([documentation](https://pytorch.org/docs/stable/nn.html#torch.nn.NLLLoss)).
>**Exercise:** Build a model that returns the log-softmax as the output and calculate the loss using the negative log likelihood loss.
```
## Solution
# Build a feed-forward network
model = nn.Sequential(nn.Linear(784, 128),
nn.ReLU(),
nn.Linear(128, 64),
nn.ReLU(),
nn.Linear(64, 10),
nn.LogSoftmax(dim=1))
# Define the loss
criterion = nn.NLLLoss()
# Get our data
images, labels = next(iter(trainloader))
# Flatten images
images = images.view(images.shape[0], -1)
# Forward pass, get our log-probabilities
logps = model(images)
# Calculate the loss with the logps and the labels
loss = criterion(logps, labels)
print(loss)
```
## Autograd
Now that we know how to calculate a loss, how do we use it to perform backpropagation? Torch provides a module, `autograd`, for automatically calculating the gradients of tensors. We can use it to calculate the gradients of all our parameters with respect to the loss. Autograd works by keeping track of operations performed on tensors, then going backwards through those operations, calculating gradients along the way. To make sure PyTorch keeps track of operations on a tensor and calculates the gradients, you need to set `requires_grad = True` on a tensor. You can do this at creation with the `requires_grad` keyword, or at any time with `x.requires_grad_(True)`.
You can turn off gradients for a block of code with the `torch.no_grad()` content:
```python
x = torch.zeros(1, requires_grad=True)
>>> with torch.no_grad():
... y = x * 2
>>> y.requires_grad
False
```
Also, you can turn on or off gradients altogether with `torch.set_grad_enabled(True|False)`.
The gradients are computed with respect to some variable `z` with `z.backward()`. This does a backward pass through the operations that created `z`.
```
x = torch.randn(2,2, requires_grad=True)
print(x)
y = x**2
print(y)
```
Below we can see the operation that created `y`, a power operation `PowBackward0`.
```
## grad_fn shows the function that generated this variable
print(y.grad_fn)
```
The autograd module keeps track of these operations and knows how to calculate the gradient for each one. In this way, it's able to calculate the gradients for a chain of operations, with respect to any one tensor. Let's reduce the tensor `y` to a scalar value, the mean.
```
z = y.mean()
print(z)
```
You can check the gradients for `x` and `y` but they are empty currently.
```
print(x.grad)
```
To calculate the gradients, you need to run the `.backward` method on a Variable, `z` for example. This will calculate the gradient for `z` with respect to `x`
$$
\frac{\partial z}{\partial x} = \frac{\partial}{\partial x}\left[\frac{1}{n}\sum_i^n x_i^2\right] = \frac{x}{2}
$$
```
z.backward()
print(x.grad)
print(x/2)
```
These gradients calculations are particularly useful for neural networks. For training we need the gradients of the weights with respect to the cost. With PyTorch, we run data forward through the network to calculate the loss, then, go backwards to calculate the gradients with respect to the loss. Once we have the gradients we can make a gradient descent step.
## Loss and Autograd together
When we create a network with PyTorch, all of the parameters are initialized with `requires_grad = True`. This means that when we calculate the loss and call `loss.backward()`, the gradients for the parameters are calculated. These gradients are used to update the weights with gradient descent. Below you can see an example of calculating the gradients using a backwards pass.
```
# Build a feed-forward network
model = nn.Sequential(nn.Linear(784, 128),
nn.ReLU(),
nn.Linear(128, 64),
nn.ReLU(),
nn.Linear(64, 10),
nn.LogSoftmax(dim=1))
criterion = nn.NLLLoss()
images, labels = next(iter(trainloader))
images = images.view(images.shape[0], -1)
logps = model(images)
loss = criterion(logps, labels)
print('Before backward pass: \n', model[0].weight.grad)
loss.backward()
print('After backward pass: \n', model[0].weight.grad)
```
## Training the network!
There's one last piece we need to start training, an optimizer that we'll use to update the weights with the gradients. We get these from PyTorch's [`optim` package](https://pytorch.org/docs/stable/optim.html). For example we can use stochastic gradient descent with `optim.SGD`. You can see how to define an optimizer below.
```
from torch import optim
# Optimizers require the parameters to optimize and a learning rate
optimizer = optim.SGD(model.parameters(), lr=0.01)
```
Now we know how to use all the individual parts so it's time to see how they work together. Let's consider just one learning step before looping through all the data. The general process with PyTorch:
* Make a forward pass through the network
* Use the network output to calculate the loss
* Perform a backward pass through the network with `loss.backward()` to calculate the gradients
* Take a step with the optimizer to update the weights
Below I'll go through one training step and print out the weights and gradients so you can see how it changes. Note that I have a line of code `optimizer.zero_grad()`. When you do multiple backwards passes with the same parameters, the gradients are accumulated. This means that you need to zero the gradients on each training pass or you'll retain gradients from previous training batches.
```
print('Initial weights - ', model[0].weight)
images, labels = next(iter(trainloader))
images.resize_(64, 784)
# Clear the gradients, do this because gradients are accumulated
optimizer.zero_grad()
# Forward pass, then backward pass, then update weights
output = model(images)
loss = criterion(output, labels)
loss.backward()
print('Gradient -', model[0].weight.grad)
# Take an update step and few the new weights
optimizer.step()
print('Updated weights - ', model[0].weight)
```
### Training for real
Now we'll put this algorithm into a loop so we can go through all the images. Some nomenclature, one pass through the entire dataset is called an *epoch*. So here we're going to loop through `trainloader` to get our training batches. For each batch, we'll doing a training pass where we calculate the loss, do a backwards pass, and update the weights.
> **Exercise: ** Implement the training pass for our network. If you implemented it correctly, you should see the training loss drop with each epoch.
```
model = nn.Sequential(nn.Linear(784, 128),
nn.ReLU(),
nn.Linear(128, 64),
nn.ReLU(),
nn.Linear(64, 10),
nn.LogSoftmax(dim=1))
criterion = nn.NLLLoss()
optimizer = optim.SGD(model.parameters(), lr=0.003)
epochs = 5
for e in range(epochs):
running_loss = 0
for images, labels in trainloader:
# Flatten MNIST images into a 784 long vector
images = images.view(images.shape[0], -1)
# TODO: Training pass
optimizer.zero_grad()
output = model(images)
loss = criterion(output, labels)
loss.backward()
optimizer.step()
running_loss += loss.item()
else:
print(f"Training loss: {running_loss/len(trainloader)}")
```
With the network trained, we can check out it's predictions.
```
%matplotlib inline
import helper
images, labels = next(iter(trainloader))
img = images[0].view(1, 784)
# Turn off gradients to speed up this part
with torch.no_grad():
logps = model(img)
# Output of the network are log-probabilities, need to take exponential for probabilities
ps = torch.exp(logps)
helper.view_classify(img.view(1, 28, 28), ps)
```
Now our network is brilliant. It can accurately predict the digits in our images. Next up you'll write the code for training a neural network on a more complex dataset.
| github_jupyter |
# {glue:text}`voila-dashboards_github_org`
**Activity from {glue:}`voila-dashboards_start` to {glue:}`voila-dashboards_stop`**
```
from datetime import date
from dateutil.relativedelta import relativedelta
from myst_nb import glue
import seaborn as sns
import pandas as pd
import numpy as np
import altair as alt
from markdown import markdown
from IPython.display import Markdown
from ipywidgets.widgets import HTML, Tab
from ipywidgets import widgets
from datetime import timedelta
from matplotlib import pyplot as plt
import os.path as op
from warnings import simplefilter
simplefilter('ignore')
# Altair config
def author_url(author):
return f"https://github.com/{author}"
def alt_theme():
return {
'config': {
'axisLeft': {
'labelFontSize': 15,
},
'axisBottom': {
'labelFontSize': 15,
},
}
}
alt.themes.register('my_theme', alt_theme)
alt.themes.enable("my_theme")
# Define colors we'll use for GitHub membership
author_types = ['MEMBER', 'CONTRIBUTOR', 'COLLABORATOR', "NONE"]
author_palette = np.array(sns.palettes.blend_palette(["lightgrey", "lightgreen", "darkgreen"], 4)) * 256
author_colors = ["rgb({}, {}, {})".format(*color) for color in author_palette]
author_color_dict = {key: val for key, val in zip(author_types, author_palette)}
github_org = "jupyterhub"
top_n_repos = 15
n_days = 10
# Parameters
github_org = "voila-dashboards"
n_days = 90
############################################################
# Variables
stop = date.today()
start = date.today() - relativedelta(days=n_days)
# Strings for use in queries
start_date = f"{start:%Y-%m-%d}"
stop_date = f"{stop:%Y-%m-%d}"
# Glue variables for use in markdown
glue(f"{github_org}_github_org", github_org, display=False)
glue(f"{github_org}_start", start_date, display=False)
glue(f"{github_org}_stop", stop_date, display=False)
```
## Load data
Load and clean up the data
```
from pathlib import Path
path_data = Path("../data")
comments = pd.read_csv(path_data.joinpath('comments.csv'), index_col=None).drop_duplicates()
issues = pd.read_csv(path_data.joinpath('issues.csv'), index_col=None).drop_duplicates()
prs = pd.read_csv(path_data.joinpath('prs.csv'), index_col=None).drop_duplicates()
for idata in [comments, issues, prs]:
idata.query("org == @github_org", inplace=True)
# What are the top N repos, we will only plot these in the full data plots
top_commented_repos = comments.groupby("repo").count().sort_values("createdAt", ascending=False)['createdAt']
use_repos = top_commented_repos.head(top_n_repos).index.tolist()
```
## Merged Pull requests
Here's an analysis of **merged pull requests** across each of the repositories in the Jupyter
ecosystem.
```
merged = prs.query('state == "MERGED" and closedAt > @start_date and closedAt < @stop_date')
prs_by_repo = merged.groupby(['org', 'repo']).count()['author'].reset_index().sort_values(['org', 'author'], ascending=False)
alt.Chart(data=prs_by_repo, title=f"Merged PRs in the last {n_days} days").mark_bar().encode(
x=alt.X('repo', sort=prs_by_repo['repo'].values.tolist()),
y='author',
color='org'
)
```
### Authoring and merging stats by repository
Let's see who has been doing most of the PR authoring and merging. The PR author is generally the
person that implemented a change in the repository (code, documentation, etc). The PR merger is
the person that "pressed the green button" and got the change into the main codebase.
```
# Prep our merging DF
merged_by_repo = merged.groupby(['repo', 'author'], as_index=False).agg({'id': 'count', 'authorAssociation': 'first'}).rename(columns={'id': "authored", 'author': 'username'})
closed_by_repo = merged.groupby(['repo', 'mergedBy']).count()['id'].reset_index().rename(columns={'id': "closed", "mergedBy": "username"})
charts = []
title = f"PR authors for {github_org} in the last {n_days} days"
this_data = merged_by_repo.replace(np.nan, 0).groupby('username', as_index=False).agg({'authored': 'sum', 'authorAssociation': 'first'})
this_data = this_data.sort_values('authored', ascending=False)
ch = alt.Chart(data=this_data, title=title).mark_bar().encode(
x='username',
y='authored',
color=alt.Color('authorAssociation', scale=alt.Scale(domain=author_types, range=author_colors))
)
ch
charts = []
title = f"Merges for {github_org} in the last {n_days} days"
ch = alt.Chart(data=closed_by_repo.replace(np.nan, 0), title=title).mark_bar().encode(
x='username',
y='closed',
)
ch
```
## Issues
Issues are **conversations** that happen on our GitHub repositories. Here's an
analysis of issues across the Jupyter organizations.
```
created = issues.query('state == "OPEN" and createdAt > @start_date and createdAt < @stop_date')
closed = issues.query('state == "CLOSED" and closedAt > @start_date and closedAt < @stop_date')
created_counts = created.groupby(['org', 'repo']).count()['number'].reset_index()
created_counts['org/repo'] = created_counts.apply(lambda a: a['org'] + '/' + a['repo'], axis=1)
sorted_vals = created_counts.sort_values(['org', 'number'], ascending=False)['repo'].values
alt.Chart(data=created_counts, title=f"Issues created in the last {n_days} days").mark_bar().encode(
x=alt.X('repo', sort=alt.Sort(sorted_vals.tolist())),
y='number',
)
closed_counts = closed.groupby(['org', 'repo']).count()['number'].reset_index()
closed_counts['org/repo'] = closed_counts.apply(lambda a: a['org'] + '/' + a['repo'], axis=1)
sorted_vals = closed_counts.sort_values(['number'], ascending=False)['repo'].values
alt.Chart(data=closed_counts, title=f"Issues closed in the last {n_days} days").mark_bar().encode(
x=alt.X('repo', sort=alt.Sort(sorted_vals.tolist())),
y='number',
)
created_closed = pd.merge(created_counts.rename(columns={'number': 'created'}).drop(columns='org/repo'),
closed_counts.rename(columns={'number': 'closed'}).drop(columns='org/repo'),
on=['org', 'repo'], how='outer')
created_closed = pd.melt(created_closed, id_vars=['org', 'repo'], var_name="kind", value_name="count").replace(np.nan, 0)
charts = []
# Pick the top 10 repositories
top_repos = created_closed.groupby(['repo']).sum().sort_values(by='count', ascending=False).head(10).index
ch = alt.Chart(created_closed.query('repo in @top_repos'), width=120).mark_bar().encode(
x=alt.X("kind", axis=alt.Axis(labelFontSize=15, title="")),
y=alt.Y('count', axis=alt.Axis(titleFontSize=15, labelFontSize=12)),
color='kind',
column=alt.Column("repo", header=alt.Header(title=f"Issue activity, last {n_days} days for {github_org}", titleFontSize=15, labelFontSize=12))
)
ch
# Set to datetime
for kind in ['createdAt', 'closedAt']:
closed.loc[:, kind] = pd.to_datetime(closed[kind])
closed.loc[:, 'time_open'] = closed['closedAt'] - closed['createdAt']
closed.loc[:, 'time_open'] = closed['time_open'].dt.total_seconds()
time_open = closed.groupby(['org', 'repo']).agg({'time_open': 'median'}).reset_index()
time_open['time_open'] = time_open['time_open'] / (60 * 60 * 24)
time_open['org/repo'] = time_open.apply(lambda a: a['org'] + '/' + a['repo'], axis=1)
sorted_vals = time_open.sort_values(['org', 'time_open'], ascending=False)['repo'].values
alt.Chart(data=time_open, title=f"Time to close for issues closed in the last {n_days} days").mark_bar().encode(
x=alt.X('repo', sort=alt.Sort(sorted_vals.tolist())),
y=alt.Y('time_open', title="Median Days Open"),
)
```
## Most-upvoted issues
```
thumbsup = issues.sort_values("thumbsup", ascending=False).head(25)
thumbsup = thumbsup[["title", "url", "number", "thumbsup", "repo"]]
text = []
for ii, irow in thumbsup.iterrows():
itext = f"- ({irow['thumbsup']}) {irow['title']} - {irow['repo']} - [#{irow['number']}]({irow['url']})"
text.append(itext)
text = '\n'.join(text)
HTML(markdown(text))
```
## Commenters across repositories
These are commenters across all issues and pull requests in the last several days.
These are colored by the commenter's association with the organization. For information
about what these associations mean, [see this StackOverflow post](https://stackoverflow.com/a/28866914/1927102).
```
commentors = (
comments
.query("createdAt > @start_date and createdAt < @stop_date")
.groupby(['org', 'repo', 'author', 'authorAssociation'])
.count().rename(columns={'id': 'count'})['count']
.reset_index()
.sort_values(['org', 'count'], ascending=False)
)
n_plot = 50
charts = []
for ii, (iorg, idata) in enumerate(commentors.groupby(['org'])):
title = f"Top {n_plot} commentors for {iorg} in the last {n_days} days"
idata = idata.groupby('author', as_index=False).agg({'count': 'sum', 'authorAssociation': 'first'})
idata = idata.sort_values('count', ascending=False).head(n_plot)
ch = alt.Chart(data=idata.head(n_plot), title=title).mark_bar().encode(
x='author',
y='count',
color=alt.Color('authorAssociation', scale=alt.Scale(domain=author_types, range=author_colors))
)
charts.append(ch)
alt.hconcat(*charts)
```
## First responders
First responders are the first people to respond to a new issue in one of the repositories.
The following plots show first responders for recently-created issues.
```
first_comments = []
for (org, repo, issue_id), i_comments in comments.groupby(['org', 'repo', 'id']):
ix_min = pd.to_datetime(i_comments['createdAt']).idxmin()
first_comment = i_comments.loc[ix_min]
if isinstance(first_comment, pd.DataFrame):
first_comment = first_comment.iloc[0]
first_comments.append(first_comment)
first_comments = pd.concat(first_comments, axis=1).T
# Make up counts for viz
first_responder_counts = first_comments.groupby(['org', 'author', 'authorAssociation'], as_index=False).\
count().rename(columns={'id': 'n_first_responses'}).sort_values(['org', 'n_first_responses'], ascending=False)
n_plot = 50
title = f"Top {n_plot} first responders for {github_org} in the last {n_days} days"
idata = first_responder_counts.groupby('author', as_index=False).agg({'n_first_responses': 'sum', 'authorAssociation': 'first'})
idata = idata.sort_values('n_first_responses', ascending=False).head(n_plot)
ch = alt.Chart(data=idata.head(n_plot), title=title).mark_bar().encode(
x='author',
y='n_first_responses',
color=alt.Color('authorAssociation', scale=alt.Scale(domain=author_types, range=author_colors))
)
ch
```
## Recent activity
### A list of merged PRs by project
Below is a tabbed readout of recently-merged PRs. Check out the title to get an idea for what they
implemented, and be sure to thank the PR author for their hard work!
```
tabs = widgets.Tab(children=[])
for ii, ((org, repo), imerged) in enumerate(merged.query("repo in @use_repos").groupby(['org', 'repo'])):
merged_by = {}
pr_by = {}
issue_md = []
issue_md.append(f"#### Closed PRs for repo: [{org}/{repo}](https://github.com/{github_org}/{repo})")
issue_md.append("")
issue_md.append(f"##### ")
for _, ipr in imerged.iterrows():
user_name = ipr['author']
user_url = author_url(user_name)
pr_number = ipr['number']
pr_html = ipr['url']
pr_title = ipr['title']
pr_closedby = ipr['mergedBy']
pr_closedby_url = f"https://github.com/{pr_closedby}"
if user_name not in pr_by:
pr_by[user_name] = 1
else:
pr_by[user_name] += 1
if pr_closedby not in merged_by:
merged_by[pr_closedby] = 1
else:
merged_by[pr_closedby] += 1
text = f"* [(#{pr_number})]({pr_html}): _{pr_title}_ by **[@{user_name}]({user_url})** merged by **[@{pr_closedby}]({pr_closedby_url})**"
issue_md.append(text)
issue_md.append('')
markdown_html = markdown('\n'.join(issue_md))
children = list(tabs.children)
children.append(HTML(markdown_html))
tabs.children = tuple(children)
tabs.set_title(ii, repo)
tabs
```
### A list of recent issues
Below is a list of issues with recent activity in each repository. If they seem of interest
to you, click on their links and jump in to participate!
```
# Add comment count data to issues and PRs
comment_counts = (
comments
.query("createdAt > @start_date and createdAt < @stop_date")
.groupby(['org', 'repo', 'id'])
.count().iloc[:, 0].to_frame()
)
comment_counts.columns = ['n_comments']
comment_counts = comment_counts.reset_index()
n_plot = 5
tabs = widgets.Tab(children=[])
for ii, (repo, i_issues) in enumerate(comment_counts.query("repo in @use_repos").groupby('repo')):
issue_md = []
issue_md.append("")
issue_md.append(f"##### [{github_org}/{repo}](https://github.com/{github_org}/{repo})")
top_issues = i_issues.sort_values('n_comments', ascending=False).head(n_plot)
top_issue_list = pd.merge(issues, top_issues, left_on=['org', 'repo', 'id'], right_on=['org', 'repo', 'id'])
for _, issue in top_issue_list.sort_values('n_comments', ascending=False).head(n_plot).iterrows():
user_name = issue['author']
user_url = author_url(user_name)
issue_number = issue['number']
issue_html = issue['url']
issue_title = issue['title']
text = f"* [(#{issue_number})]({issue_html}): _{issue_title}_ by **[@{user_name}]({user_url})**"
issue_md.append(text)
issue_md.append('')
md_html = HTML(markdown('\n'.join(issue_md)))
children = list(tabs.children)
children.append(HTML(markdown('\n'.join(issue_md))))
tabs.children = tuple(children)
tabs.set_title(ii, repo)
display(Markdown(f"Here are the top {n_plot} active issues in each repository in the last {n_days} days"))
display(tabs)
```
| github_jupyter |
# Hyperparameter tuning by grid-search
In the previous notebook, we saw that hyperparameters can affect the
statistical performance of a model. In this notebook, we will show how to
optimize hyperparameters using a grid-search approach.
## Our predictive model
Let us reload the dataset as we did previously:
```
from sklearn import set_config
set_config(display="diagram")
import pandas as pd
adult_census = pd.read_csv("../datasets/adult-census.csv")
```
We extract the column containing the target.
```
target_name = "class"
target = adult_census[target_name]
target
```
We drop from our data the target and the `"education-num"` column which
duplicates the information from the `"education"` column.
```
data = adult_census.drop(columns=[target_name, "education-num"])
data.head()
```
Once the dataset is loaded, we split it into a training and testing sets.
```
from sklearn.model_selection import train_test_split
data_train, data_test, target_train, target_test = train_test_split(
data, target, random_state=42)
```
We will define a pipeline as seen in the first module. It will handle both
numerical and categorical features.
As we will use a tree-based model as a predictor, here we apply an ordinal
encoder on the categorical features: it encodes every category with an
arbitrary integer. For simple models such as linear models, a one-hot encoder
should be preferred. But for complex models, in particular tree-based models,
the ordinal encoder is useful as it avoids having high-dimensional
representations.
First we select all the categorical columns.
```
from sklearn.compose import make_column_selector as selector
categorical_columns_selector = selector(dtype_include=object)
categorical_columns = categorical_columns_selector(data)
```
Then we build our ordinal encoder, giving it the known categories.
```
from sklearn.preprocessing import OrdinalEncoder
categorical_preprocessor = OrdinalEncoder(handle_unknown="use_encoded_value",
unknown_value=-1)
```
We now use a column transformer with code to select the categorical columns
and apply to them the ordinal encoder.
```
from sklearn.compose import ColumnTransformer
preprocessor = ColumnTransformer([
('cat-preprocessor', categorical_preprocessor, categorical_columns)],
remainder='passthrough', sparse_threshold=0)
```
Finally, we use a tree-based classifier (i.e. histogram gradient-boosting) to
predict whether or not a person earns more than 50 k$ a year.
```
# for the moment this line is required to import HistGradientBoostingClassifier
from sklearn.experimental import enable_hist_gradient_boosting
from sklearn.ensemble import HistGradientBoostingClassifier
from sklearn.pipeline import Pipeline
model = Pipeline([
("preprocessor", preprocessor),
("classifier",
HistGradientBoostingClassifier(random_state=42, max_leaf_nodes=4))])
model
```
## Tuning using a grid-search
Instead of manually writing the two `for` loops, scikit-learn provides a
class called `GridSearchCV` which implement the exhaustive search implemented
during the exercise.
Let see how to use the `GridSearchCV` estimator for doing such search.
Since the grid-search will be costly, we will only explore the combination
learning-rate and the maximum number of nodes.
```
%%time
from sklearn.model_selection import GridSearchCV
param_grid = {
'classifier__learning_rate': (0.05, 0.1, 0.5, 1, 5),
'classifier__max_leaf_nodes': (3, 10, 30, 100)}
model_grid_search = GridSearchCV(model, param_grid=param_grid,
n_jobs=2, cv=2)
model_grid_search.fit(data_train, target_train)
```
Finally, we will check the accuracy of our model using the test set.
```
accuracy = model_grid_search.score(data_test, target_test)
print(
f"The test accuracy score of the grid-searched pipeline is: "
f"{accuracy:.2f}"
)
```
<div class="admonition warning alert alert-danger">
<p class="first admonition-title" style="font-weight: bold;">Warning</p>
<p>Be aware that the evaluation should normally be performed in a
cross-validation framework by providing <tt class="docutils literal">model_grid_search</tt> as a model to
the <tt class="docutils literal">cross_validate</tt> function.</p>
<p class="last">Here, we are using a single train-test split to highlight the specificities
of the <tt class="docutils literal">model_grid_search</tt> instance. We will show such examples in the last
section of this notebook.</p>
</div>
The `GridSearchCV` estimator takes a `param_grid` parameter which defines
all hyperparameters and their associated values. The grid-search will be in
charge of creating all possible combinations and test them.
The number of combinations will be equal to the product of the
number of values to explore for each parameter (e.g. in our example 4 x 4
combinations). Thus, adding new parameters with their associated values to be
explored become rapidly computationally expensive.
Once the grid-search is fitted, it can be used as any other predictor by
calling `predict` and `predict_proba`. Internally, it will use the model with
the best parameters found during `fit`.
Get predictions for the 5 first samples using the estimator with the best
parameters.
```
model_grid_search.predict(data_test.iloc[0:5])
```
You can know about these parameters by looking at the `best_params_`
attribute.
```
print(f"The best set of parameters is: "
f"{model_grid_search.best_params_}")
```
The accuracy and the best parameters of the grid-searched pipeline are
similar to the ones we found in the previous exercise, where we searched the
best parameters "by hand" through a double for loop.
In addition, we can inspect all results which are stored in the attribute
`cv_results_` of the grid-search. We will filter some specific columns
from these results.
```
cv_results = pd.DataFrame(model_grid_search.cv_results_).sort_values(
"mean_test_score", ascending=False)
cv_results.head()
```
Let us focus on the most interesting columns and shorten the parameter
names to remove the `"param_classifier__"` prefix for readability:
```
# get the parameter names
column_results = [f"param_{name}" for name in param_grid.keys()]
column_results += [
"mean_test_score", "std_test_score", "rank_test_score"]
cv_results = cv_results[column_results]
def shorten_param(param_name):
if "__" in param_name:
return param_name.rsplit("__", 1)[1]
return param_name
cv_results = cv_results.rename(shorten_param, axis=1)
cv_results
```
With only 2 parameters, we might want to visualize the grid-search as a
heatmap. We need to transform our `cv_results` into a dataframe where:
- the rows will correspond to the learning-rate values;
- the columns will correspond to the maximum number of leaf;
- the content of the dataframe will be the mean test scores.
```
pivoted_cv_results = cv_results.pivot_table(
values="mean_test_score", index=["learning_rate"],
columns=["max_leaf_nodes"])
pivoted_cv_results
```
We can use a heatmap representation to show the above dataframe visually.
```
import seaborn as sns
ax = sns.heatmap(pivoted_cv_results, annot=True, cmap="YlGnBu", vmin=0.7,
vmax=0.9)
ax.invert_yaxis()
```
The above tables highlights the following things:
* for too high values of `learning_rate`, the statistical performance of the
model is degraded and adjusting the value of `max_leaf_nodes` cannot fix
that problem;
* outside of this pathological region, we observe that the optimal choice
of `max_leaf_nodes` depends on the value of `learning_rate`;
* in particular, we observe a "diagonal" of good models with an accuracy
close to the maximal of 0.87: when the value of `max_leaf_nodes` is
increased, one should increase the value of `learning_rate` accordingly
to preserve a good accuracy.
The precise meaning of those two parameters will be explained in a latter
notebook.
For now we will note that, in general, **there is no unique optimal parameter
setting**: 6 models out of the 16 parameter configuration reach the maximal
accuracy (up to small random fluctuations caused by the sampling of the
training set).
In this notebook we have seen:
* how to optimize the hyperparameters of a predictive model via a
grid-search;
* that searching for more than two hyperparamters is too costly;
* that a grid-search does not necessarily find an optimal solution.
| github_jupyter |
```
# from google.colab import drive
# drive.mount('/content/drive')
import torch.nn as nn
import torch.nn.functional as F
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import torch
import torchvision
import torchvision.transforms as transforms
from torch.utils.data import Dataset, DataLoader
from torchvision import transforms, utils
from matplotlib import pyplot as plt
import copy
# Ignore warnings
import warnings
warnings.filterwarnings("ignore")
transform = transforms.Compose(
[transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
trainset = torchvision.datasets.CIFAR10(root='./data', train=True, download=True, transform=transform)
testset = torchvision.datasets.CIFAR10(root='./data', train=False, download=True, transform=transform)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=10, shuffle=True)
testloader = torch.utils.data.DataLoader(testset, batch_size=10, shuffle=False)
classes = ('plane', 'car', 'bird', 'cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck')
foreground_classes = {'plane', 'car', 'bird'}
background_classes = {'cat', 'deer', 'dog', 'frog', 'horse','ship', 'truck'}
fg1,fg2,fg3 = 0,1,2
dataiter = iter(trainloader)
background_data=[]
background_label=[]
foreground_data=[]
foreground_label=[]
batch_size=10
for i in range(5000):
images, labels = dataiter.next()
for j in range(batch_size):
if(classes[labels[j]] in background_classes):
img = images[j].tolist()
background_data.append(img)
background_label.append(labels[j])
else:
img = images[j].tolist()
foreground_data.append(img)
foreground_label.append(labels[j])
foreground_data = torch.tensor(foreground_data)
foreground_label = torch.tensor(foreground_label)
background_data = torch.tensor(background_data)
background_label = torch.tensor(background_label)
def create_mosaic_img(bg_idx,fg_idx,fg):
"""
bg_idx : list of indexes of background_data[] to be used as background images in mosaic
fg_idx : index of image to be used as foreground image from foreground data
fg : at what position/index foreground image has to be stored out of 0-8
"""
image_list=[]
j=0
for i in range(9):
if i != fg:
image_list.append(background_data[bg_idx[j]].type("torch.DoubleTensor"))
j+=1
else:
image_list.append(foreground_data[fg_idx].type("torch.DoubleTensor"))
label = foreground_label[fg_idx]- fg1 # minus 7 because our fore ground classes are 7,8,9 but we have to store it as 0,1,2
#image_list = np.concatenate(image_list ,axis=0)
image_list = torch.stack(image_list)
return image_list,label
desired_num = 30000
mosaic_list_of_images =[] # list of mosaic images, each mosaic image is saved as list of 9 images
fore_idx =[] # list of indexes at which foreground image is present in a mosaic image i.e from 0 to 9
mosaic_label=[] # label of mosaic image = foreground class present in that mosaic
for i in range(desired_num):
bg_idx = np.random.randint(0,35000,8)
fg_idx = np.random.randint(0,15000)
fg = np.random.randint(0,9)
fore_idx.append(fg)
image_list,label = create_mosaic_img(bg_idx,fg_idx,fg)
mosaic_list_of_images.append(image_list)
mosaic_label.append(label)
class MosaicDataset(Dataset):
"""MosaicDataset dataset."""
def __init__(self, mosaic_list_of_images, mosaic_label, fore_idx):
"""
Args:
csv_file (string): Path to the csv file with annotations.
root_dir (string): Directory with all the images.
transform (callable, optional): Optional transform to be applied
on a sample.
"""
self.mosaic = mosaic_list_of_images
self.label = mosaic_label
self.fore_idx = fore_idx
def __len__(self):
return len(self.label)
def __getitem__(self, idx):
return self.mosaic[idx] , self.label[idx], self.fore_idx[idx]
batch = 250
msd = MosaicDataset(mosaic_list_of_images, mosaic_label , fore_idx)
train_loader = DataLoader( msd,batch_size= batch ,shuffle=True)
class Focus(nn.Module):
def __init__(self):
super(Focus, self).__init__()
self.conv1 = nn.Conv2d(in_channels=3, out_channels=6, kernel_size=3, padding=0)
self.pool = nn.MaxPool2d(2, 2)
self.conv2 = nn.Conv2d(in_channels=6, out_channels=6, kernel_size=3, padding=0)
# self.conv3 = nn.Conv2d(in_channels=12, out_channels=32, kernel_size=3, padding=0)
self.fc1 = nn.Linear(1014, 512)
self.fc2 = nn.Linear(512, 64)
# self.fc3 = nn.Linear(512, 64)
# self.fc4 = nn.Linear(64, 10)
self.fc3 = nn.Linear(64,1)
def forward(self,z): #y is avg image #z batch of list of 9 images
y = torch.zeros([batch,3, 32,32], dtype=torch.float64)
x = torch.zeros([batch,9],dtype=torch.float64)
y = y.to("cuda")
x = x.to("cuda")
for i in range(9):
x[:,i] = self.helper(z[:,i])[:,0]
x = F.softmax(x,dim=1)
x1 = x[:,0]
torch.mul(x1[:,None,None,None],z[:,0])
for i in range(9):
x1 = x[:,i]
y = y + torch.mul(x1[:,None,None,None],z[:,i])
return x, y
def helper(self, x):
x = self.pool(F.relu(self.conv1(x)))
x = (F.relu(self.conv2(x)))
# print(x.shape)
# x = (F.relu(self.conv3(x)))
x = x.view(x.size(0), -1)
# print(x.shape)
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
# x = F.relu(self.fc3(x))
# x = F.relu(self.fc4(x))
x = self.fc3(x)
return x
focus_net = Focus().double()
focus_net = focus_net.to("cuda")
class Classification(nn.Module):
def __init__(self):
super(Classification, self).__init__()
self.conv1 = nn.Conv2d(in_channels=3, out_channels=6, kernel_size=3, padding=0)
self.pool = nn.MaxPool2d(2, 2)
self.conv2 = nn.Conv2d(in_channels=6, out_channels=6, kernel_size=3, padding=0)
# self.conv3 = nn.Conv2d(in_channels=12, out_channels=20, kernel_size=3, padding=0)
self.fc1 = nn.Linear(1014, 512)
self.fc2 = nn.Linear(512, 64)
# self.fc3 = nn.Linear(512, 64)
# self.fc4 = nn.Linear(64, 10)
self.fc3 = nn.Linear(64,3)
def forward(self, x):
x = self.pool(F.relu(self.conv1(x)))
x = (F.relu(self.conv2(x)))
# print(x.shape)
# x = (F.relu(self.conv3(x)))
x = x.view(x.size(0), -1)
# print(x.shape)
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
# x = F.relu(self.fc3(x))
# x = F.relu(self.fc4(x))
x = self.fc3(x)
return x
classify = Classification().double()
classify = classify.to("cuda")
test_images =[] #list of mosaic images, each mosaic image is saved as laist of 9 images
fore_idx_test =[] #list of indexes at which foreground image is present in a mosaic image
test_label=[] # label of mosaic image = foreground class present in that mosaic
for i in range(10000):
bg_idx = np.random.randint(0,35000,8)
fg_idx = np.random.randint(0,15000)
fg = np.random.randint(0,9)
fore_idx_test.append(fg)
image_list,label = create_mosaic_img(bg_idx,fg_idx,fg)
test_images.append(image_list)
test_label.append(label)
test_data = MosaicDataset(test_images,test_label,fore_idx_test)
test_loader = DataLoader( test_data,batch_size= batch ,shuffle=False)
import torch.optim as optim
criterion = nn.CrossEntropyLoss()
optimizer_classify = optim.Adam(classify.parameters(), lr=0.001)#, betas=(0.9, 0.999), eps=1e-08, weight_decay=0, amsgrad=False)
optimizer_focus = optim.Adam(focus_net.parameters(), lr=0.001)#, betas=(0.9, 0.999), eps=1e-08, weight_decay=0, amsgrad=False)
col1=[]
col2=[]
col3=[]
col4=[]
col5=[]
col6=[]
col7=[]
col8=[]
col9=[]
col10=[]
col11=[]
col12=[]
col13=[]
correct = 0
total = 0
count = 0
flag = 1
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
with torch.no_grad():
for data in train_loader:
inputs, labels , fore_idx = data
inputs, labels , fore_idx = inputs.to("cuda"),labels.to("cuda"), fore_idx.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
for j in range(labels.size(0)):
count += 1
focus = torch.argmax(alphas[j])
if alphas[j][focus] >= 0.5 :
argmax_more_than_half += 1
else:
argmax_less_than_half += 1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true += 1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false += 1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false += 1
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 30000 train images: %d %%' % ( 100 * correct / total))
print("total correct", correct)
print("total train set images", total)
print("focus_true_pred_true %d =============> FTPT : %d %%" % (focus_true_pred_true , (100 * focus_true_pred_true / total) ) )
print("focus_false_pred_true %d =============> FFPT : %d %%" % (focus_false_pred_true, (100 * focus_false_pred_true / total) ) )
print("focus_true_pred_false %d =============> FTPF : %d %%" %( focus_true_pred_false , ( 100 * focus_true_pred_false / total) ) )
print("focus_false_pred_false %d =============> FFPF : %d %%" % (focus_false_pred_false, ( 100 * focus_false_pred_false / total) ) )
print("argmax_more_than_half ==================> ",argmax_more_than_half)
print("argmax_less_than_half ==================> ",argmax_less_than_half)
print(count)
print("="*100)
col1.append(0)
col2.append(argmax_more_than_half)
col3.append(argmax_less_than_half)
col4.append(focus_true_pred_true)
col5.append(focus_false_pred_true)
col6.append(focus_true_pred_false)
col7.append(focus_false_pred_false)
correct = 0
total = 0
count = 0
flag = 1
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
with torch.no_grad():
for data in test_loader:
inputs, labels , fore_idx = data
inputs, labels , fore_idx = inputs.to("cuda"),labels.to("cuda"), fore_idx.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
for j in range(labels.size(0)):
focus = torch.argmax(alphas[j])
if alphas[j][focus] >= 0.5 :
argmax_more_than_half += 1
else:
argmax_less_than_half += 1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true += 1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false += 1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false += 1
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 test images: %d %%' % (
100 * correct / total))
print("total correct", correct)
print("total train set images", total)
print("focus_true_pred_true %d =============> FTPT : %d %%" % (focus_true_pred_true , (100 * focus_true_pred_true / total) ) )
print("focus_false_pred_true %d =============> FFPT : %d %%" % (focus_false_pred_true, (100 * focus_false_pred_true / total) ) )
print("focus_true_pred_false %d =============> FTPF : %d %%" %( focus_true_pred_false , ( 100 * focus_true_pred_false / total) ) )
print("focus_false_pred_false %d =============> FFPF : %d %%" % (focus_false_pred_false, ( 100 * focus_false_pred_false / total) ) )
print("argmax_more_than_half ==================> ",argmax_more_than_half)
print("argmax_less_than_half ==================> ",argmax_less_than_half)
col8.append(argmax_more_than_half)
col9.append(argmax_less_than_half)
col10.append(focus_true_pred_true)
col11.append(focus_false_pred_true)
col12.append(focus_true_pred_false)
col13.append(focus_false_pred_false)
nos_epochs = 200
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
for epoch in range(nos_epochs): # loop over the dataset multiple times
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
running_loss = 0.0
epoch_loss = []
cnt=0
iteration = desired_num // batch
#training data set
for i, data in enumerate(train_loader):
inputs , labels , fore_idx = data
inputs, labels = inputs.to("cuda"), labels.to("cuda")
# zero the parameter gradients
optimizer_focus.zero_grad()
optimizer_classify.zero_grad()
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
# print(outputs)
# print(outputs.shape,labels.shape , torch.argmax(outputs, dim=1))
loss = criterion(outputs, labels)
loss.backward()
optimizer_focus.step()
optimizer_classify.step()
running_loss += loss.item()
mini = 60
if cnt % mini == mini-1: # print every 40 mini-batches
print('[%d, %5d] loss: %.3f' %(epoch + 1, cnt + 1, running_loss / mini))
epoch_loss.append(running_loss/mini)
running_loss = 0.0
cnt=cnt+1
if epoch % 5 == 0:
for j in range (batch):
focus = torch.argmax(alphas[j])
if(alphas[j][focus] >= 0.5):
argmax_more_than_half +=1
else:
argmax_less_than_half +=1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true +=1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false +=1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false +=1
if(np.mean(epoch_loss) <= 0.005):
break;
if epoch % 5 == 0:
# focus_net.eval()
# classify.eval()
col1.append(epoch+1)
col2.append(argmax_more_than_half)
col3.append(argmax_less_than_half)
col4.append(focus_true_pred_true)
col5.append(focus_false_pred_true)
col6.append(focus_true_pred_false)
col7.append(focus_false_pred_false)
#************************************************************************
#testing data set
with torch.no_grad():
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
for data in test_loader:
inputs, labels , fore_idx = data
inputs, labels = inputs.to("cuda"), labels.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
for j in range (batch):
focus = torch.argmax(alphas[j])
if(alphas[j][focus] >= 0.5):
argmax_more_than_half +=1
else:
argmax_less_than_half +=1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true +=1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false +=1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false +=1
col8.append(argmax_more_than_half)
col9.append(argmax_less_than_half)
col10.append(focus_true_pred_true)
col11.append(focus_false_pred_true)
col12.append(focus_true_pred_false)
col13.append(focus_false_pred_false)
print('Finished Training')
# torch.save(focus_net.state_dict(),"/content/drive/My Drive/Research/Cheating_data/16_experiments_on_cnn_3layers/"+name+"_focus_net.pt")
# torch.save(classify.state_dict(),"/content/drive/My Drive/Research/Cheating_data/16_experiments_on_cnn_3layers/"+name+"_classify.pt")
columns = ["epochs", "argmax > 0.5" ,"argmax < 0.5", "focus_true_pred_true", "focus_false_pred_true", "focus_true_pred_false", "focus_false_pred_false" ]
df_train = pd.DataFrame()
df_test = pd.DataFrame()
df_train[columns[0]] = col1
df_train[columns[1]] = col2
df_train[columns[2]] = col3
df_train[columns[3]] = col4
df_train[columns[4]] = col5
df_train[columns[5]] = col6
df_train[columns[6]] = col7
df_test[columns[0]] = col1
df_test[columns[1]] = col8
df_test[columns[2]] = col9
df_test[columns[3]] = col10
df_test[columns[4]] = col11
df_test[columns[5]] = col12
df_test[columns[6]] = col13
df_train
# plt.figure(12,12)
plt.plot(col1,col2, label='argmax > 0.5')
plt.plot(col1,col3, label='argmax < 0.5')
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.xlabel("epochs")
plt.ylabel("training data")
plt.title("On Training set")
plt.show()
plt.plot(col1,col4, label ="focus_true_pred_true ")
plt.plot(col1,col5, label ="focus_false_pred_true ")
plt.plot(col1,col6, label ="focus_true_pred_false ")
plt.plot(col1,col7, label ="focus_false_pred_false ")
plt.title("On Training set")
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.xlabel("epochs")
plt.ylabel("training data")
plt.savefig("train_ftpt.pdf", bbox_inches='tight')
plt.show()
df_test
# plt.figure(12,12)
plt.plot(col1,col8, label='argmax > 0.5')
plt.plot(col1,col9, label='argmax < 0.5')
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.xlabel("epochs")
plt.ylabel("Testing data")
plt.title("On Testing set")
plt.show()
plt.plot(col1,col10, label ="focus_true_pred_true ")
plt.plot(col1,col11, label ="focus_false_pred_true ")
plt.plot(col1,col12, label ="focus_true_pred_false ")
plt.plot(col1,col13, label ="focus_false_pred_false ")
plt.title("On Testing set")
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.xlabel("epochs")
plt.ylabel("Testing data")
plt.savefig("test_ftpt.pdf", bbox_inches='tight')
plt.show()
correct = 0
total = 0
count = 0
flag = 1
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
with torch.no_grad():
for data in train_loader:
inputs, labels , fore_idx = data
inputs, labels , fore_idx = inputs.to("cuda"),labels.to("cuda"), fore_idx.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
for j in range(labels.size(0)):
focus = torch.argmax(alphas[j])
if alphas[j][focus] >= 0.5 :
argmax_more_than_half += 1
else:
argmax_less_than_half += 1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true += 1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false += 1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false += 1
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 30000 train images: %d %%' % (
100 * correct / total))
print("total correct", correct)
print("total train set images", total)
print("focus_true_pred_true %d =============> FTPT : %d %%" % (focus_true_pred_true , (100 * focus_true_pred_true / total) ) )
print("focus_false_pred_true %d =============> FFPT : %d %%" % (focus_false_pred_true, (100 * focus_false_pred_true / total) ) )
print("focus_true_pred_false %d =============> FTPF : %d %%" %( focus_true_pred_false , ( 100 * focus_true_pred_false / total) ) )
print("focus_false_pred_false %d =============> FFPF : %d %%" % (focus_false_pred_false, ( 100 * focus_false_pred_false / total) ) )
print("argmax_more_than_half ==================> ",argmax_more_than_half)
print("argmax_less_than_half ==================> ",argmax_less_than_half)
correct = 0
total = 0
count = 0
flag = 1
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
with torch.no_grad():
for data in test_loader:
inputs, labels , fore_idx = data
inputs, labels , fore_idx = inputs.to("cuda"),labels.to("cuda"), fore_idx.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
for j in range(labels.size(0)):
focus = torch.argmax(alphas[j])
if alphas[j][focus] >= 0.5 :
argmax_more_than_half += 1
else:
argmax_less_than_half += 1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true += 1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false += 1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false += 1
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 test images: %d %%' % (
100 * correct / total))
print("total correct", correct)
print("total train set images", total)
print("focus_true_pred_true %d =============> FTPT : %d %%" % (focus_true_pred_true , (100 * focus_true_pred_true / total) ) )
print("focus_false_pred_true %d =============> FFPT : %d %%" % (focus_false_pred_true, (100 * focus_false_pred_true / total) ) )
print("focus_true_pred_false %d =============> FTPF : %d %%" %( focus_true_pred_false , ( 100 * focus_true_pred_false / total) ) )
print("focus_false_pred_false %d =============> FFPF : %d %%" % (focus_false_pred_false, ( 100 * focus_false_pred_false / total) ) )
print("argmax_more_than_half ==================> ",argmax_more_than_half)
print("argmax_less_than_half ==================> ",argmax_less_than_half)
correct = 0
total = 0
with torch.no_grad():
for data in train_loader:
inputs, labels , fore_idx = data
inputs, labels = inputs.to("cuda"), labels.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 30000 train images: %d %%' % ( 100 * correct / total))
print("total correct", correct)
print("total train set images", total)
correct = 0
total = 0
with torch.no_grad():
for data in test_loader:
inputs, labels , fore_idx = data
inputs, labels = inputs.to("cuda"), labels.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 test images: %d %%' % ( 100 * correct / total))
print("total correct", correct)
print("total train set images", total)
max_alpha =[]
alpha_ftpt=[]
argmax_more_than_half=0
argmax_less_than_half=0
for i, data in enumerate(test_loader):
inputs, labels,_ = data
inputs = inputs.double()
inputs, labels = inputs.to("cuda"),labels.to("cuda")
alphas, avg = focus_net(inputs)
outputs = classify(avg)
mx,_ = torch.max(alphas,1)
max_alpha.append(mx.cpu().detach().numpy())
for j in range(labels.size(0)):
focus = torch.argmax(alphas[j])
if alphas[j][focus] >= 0.5 :
argmax_more_than_half += 1
else:
argmax_less_than_half += 1
if (focus == fore_idx[j] and predicted[j] == labels[j]):
alpha_ftpt.append(alphas[j][focus].item())
max_alpha = np.concatenate(max_alpha,axis=0)
print(max_alpha.shape)
plt.figure(figsize=(6,6))
_,bins,_ = plt.hist(max_alpha,bins=50,color ="c")
plt.title("alpha values histogram")
plt.savefig("alpha_hist.pdf")
plt.figure(figsize=(6,6))
_,bins,_ = plt.hist(np.array(alpha_ftpt),bins=50,color ="c")
plt.title("alpha values in ftpt")
plt.savefig("alpha_hist_ftpt.pdf")
```
| github_jupyter |
### MLflow Project and Model Example
<table>
<tr><td>
<img src="https://raw.githubusercontent.com/dmatrix/mlflow-workshop-part-2/master/images/models_1.png"
alt="Bank Note " width="400">
</td></tr>
<tr><td> Save a Keras Model Flavor and load as both Keras Native Flavor and PyFunc Flavor</td></tr>
<tr><td>
<img src="https://raw.githubusercontent.com/dmatrix/mlflow-workshop-part-2/master/images/models_2.png"
alt="Bank Note " width="400">
</td></tr>
</table>
```
!pip install keras
import warnings
import tensorflow as tf
import keras
import pandas as pd
import numpy as np
import mlflow.keras
import mlflow.pyfunc
warnings.filterwarnings("ignore", category=DeprecationWarning)
print(f"mlflow version={mlflow.__version__};keras version={keras.__version__};tensorlow version={tf.__version__}")
```
[Source](https://androidkt.com/linear-regression-model-in-keras/)
Modified and extended for this tutorial
Problem: Build a simple Linear NN Model that predicts Celsius temperaturers from training data with Fahrenheit degree
<table>
<tr><td>
<img src="https://raw.githubusercontent.com/dmatrix/mlflow-workshop-part-2/master/images/temperature-conversion.png"
alt="Bank Note " width="600">
</td></tr>
</table>
Generate our X, y, and predict data
```
def f2c(f):
return (f - 32) * 5.0/9.0
def gen_data(start, stop, step):
X_fahrenheit = np.arange(start, stop, step, dtype=float)
# Randomize the input
np.random.shuffle(X_fahrenheit)
y_celsius = np.array(np.array([f2c(f) for f in X_fahrenheit]))
predict_data =[]
[predict_data.append(t) for t in range (212, 170, -5)]
return (X_fahrenheit, y_celsius, predict_data)
```
#### Define Inference Functions
```
def predict_keras_model(uri, data):
model = mlflow.keras.load_model(uri)
return [(f"(F={f}, C={model.predict([f])[0]})") for f in data]
def predict_pyfunc_model(uri, data):
# Given Fahrenheit -> Predict Celcius
# Create a pandas DataFrame with Fahrenheit unseen values
# Get the Celius prediction
pyfunc_model = mlflow.pyfunc.load_model(uri)
df = pd.DataFrame(np.array(data))
return pyfunc_model.predict(df)
```
#### Build a Keras Dense NN model
```
# Define the model
def baseline_model():
model = keras.Sequential([
keras.layers.Dense(64, activation='relu', input_shape=[1]),
keras.layers.Dense(64, activation='relu'),
keras.layers.Dense(1)
])
optimizer = keras.optimizers.RMSprop(0.001)
# Compile the model
model.compile(loss='mean_squared_error',
optimizer=optimizer,
metrics=['mean_absolute_error', 'mean_squared_error'])
return model
```
Capture run metrics using MLflow API
With `model.<flavor>.autolog()`, you don't need to log parameter, metrics, models etc.
autlogging is a convenient method to make it easier to use MLflow Fluent APIs, hence making your code eaiser to read.
* Autologgin Features for [Model Flavors](https://mlflow.org/docs/latest/tracking.html#automatic-logging)
* PR for in development:
* `mlflow.pytorch.autolog()`
* `mlflow.scklearn.autolog()`
```
def mlflow_run(params, X, y, run_name="Keras Linear Regression"):
# Start MLflow run and log everyting...
with mlflow.start_run(run_name=run_name) as run:
run_id = run.info.run_uuid
exp_id = run.info.experiment_id
model = baseline_model()
# single line of MLflow Fluent API obviates the need to log
# individual parameters, metrics, model, artifacts etc...
# https://mlflow.org/docs/latest/python_api/mlflow.keras.html#mlflow.keras.autolog
mlflow.keras.autolog()
model.fit(X, y, batch_size=params['batch_size'], epochs=params['epochs'])
return (exp_id, run_id)
```
#### Train the Keras model
### Run 1
Generate X, y, and predict_data
```
(X, y, predict_data) = gen_data(-212, 10512, 2)
params = {'batch_size': 10,'epochs': 100}
(exp_id, run_id) = mlflow_run(params, X, y)
print(f"Finished Experiment id={exp_id} and run id = {run_id}")
```
### Check the MLflow UI
* check the Model file
* check the Conda.yaml file
* check the metrics
### Load the Keras Model Flavor: Native Flavor
```
# Load this Keras Model Flavor as a Keras native model flavor and make a prediction
model_uri = f"runs:/{run_id}/model"
print(f"Loading the Keras Model={model_uri} as Keras Model")
predictions = predict_keras_model(model_uri, predict_data)
print(predictions)
```
### Load the Keras Model Flavor as a PyFunc Flavor
```
# Load this Keras Model Flavor as a pyfunc model flavor and make a prediction
pyfunc_uri = f"runs:/{run_id}/model"
print(f"Loading the Keras Model={pyfunc_uri} as Pyfunc Model")
predictions = predict_pyfunc_model(pyfunc_uri, predict_data)
print(predictions)
```
### Excercise/Lab Assignment
Using what we have learning in this session:
* Can you decrease the MSE?
* Increase the size of input data
* Change the batch size and number of epochs
* Run at least three experiments with different parameters: number of epochs, batches
* Compare the runs and metrics
Were you able lower the MSE?
**CHALLENGE**: Excercise/Lab Assignment
* Convert this program as a GitHub MLflow Project
* Use the Lab 1 [MLflow Project](https://github.com/mlflow/mlflow-example) as a template
* Other examples are in [here](https://github.com/mlflow/mlflow/tree/master/examples)
| github_jupyter |
Copyright (c) Microsoft Corporation. All rights reserved.
Licensed under the MIT License.
# ONNX Runtime: Tutorial for Nuphar execution provider
**Accelerating model inference via compiler, using Docker Images for ONNX Runtime with Nuphar**
This example shows how to accelerate model inference using Nuphar, an execution provider that leverages just-in-time compilation to generate optimized executables.
For more background about Nuphar, please check [Nuphar-ExecutionProvider.md](https://github.com/microsoft/onnxruntime/blob/master/docs/execution_providers/Nuphar-ExecutionProvider.md) and its [build instructions](https://github.com/microsoft/onnxruntime/blob/master/BUILD.md#nuphar).
#### Tutorial Roadmap:
0. Prerequistes
1. Create and run inference on a simple ONNX model, and understand how ***compilation*** works in Nuphar.
2. Create and run inference on a model using ***LSTM***, run symbolic shape inference, edit LSTM ops to Scan, and check Nuphar speedup.
3. ***Quantize*** the LSTM model and check speedup in Nuphar (CPU with AVX2 support is required).
4. Working on a real model: ***Bidirectional Attention Flow ([BiDAF](https://arxiv.org/pdf/1611.01603))*** from onnx model zoo.
5. ***Ahead-Of-Time (AOT) compilation*** to save just-in-time compilation cost on model load.
## 0. Prerequistes
Please make sure you have installed following Python packages. Besides, C++ compiler/linker is required for ahead-of-time compilation. Please make sure you have g++ if running on Linux, or Visual Studio 2017 on Windows.
```
import cpufeature
import numpy as np
import onnx
from onnx import helper, numpy_helper
import os
from timeit import default_timer as timer
import shutil
import subprocess
import sys
import tarfile
import urllib.request
def is_windows():
return sys.platform.startswith('win')
if is_windows():
assert shutil.which('cl.exe'), 'Please make sure MSVC compiler and liner are in PATH.'
else:
assert shutil.which('g++'), 'Please make sure g++ is installed.'
```
And Nuphar package in onnxruntime is required too. Please make sure you are using Nuphar enabled build.
```
import onnxruntime
from onnxruntime.nuphar.model_editor import convert_to_scan_model
from onnxruntime.nuphar.model_quantizer import convert_matmul_model
from onnxruntime.nuphar.rnn_benchmark import generate_model
from onnxruntime.nuphar.symbolic_shape_infer import SymbolicShapeInference
```
## 1. Create and run inference on a simple ONNX model
Let's start with a simple model: Y = ((X + X) * X + X) * X + X
```
model = onnx.ModelProto()
opset = model.opset_import.add()
opset.domain == 'onnx'
opset.version = 7 # ONNX opset 7 is required for LSTM op later
graph = model.graph
X = 'input'
Y = 'output'
# declare graph input/ouput with shape [seq, batch, 1024]
dim = 1024
model.graph.input.add().CopyFrom(helper.make_tensor_value_info(X, onnx.TensorProto.FLOAT, ['seq', 'batch', dim]))
model.graph.output.add().CopyFrom(helper.make_tensor_value_info(Y, onnx.TensorProto.FLOAT, ['seq', 'batch', dim]))
# create nodes: Y = ((X + X) * X + X) * X + X
num_nodes = 5
for i in range(num_nodes):
n = helper.make_node('Mul' if i % 2 else 'Add',
[X, X if i == 0 else 'out_'+str(i-1)],
['out_'+str(i) if i < num_nodes - 1 else Y],
'node'+str(i))
model.graph.node.add().CopyFrom(n)
# save the model
simple_model_name = 'simple.onnx'
onnx.save(model, simple_model_name)
```
We will use nuphar execution provider to run the inference for the model that we created above, and use settings string to check the generated code.
Because of the redirection of output, we dump the lowered code from a subprocess to a log file:
```
code_to_run = '''
import onnxruntime
s = 'codegen_dump_lower:verbose'
onnxruntime.capi._pybind_state.set_nuphar_settings(s)
sess = onnxruntime.InferenceSession('simple.onnx')
'''
log_file = 'simple_lower.log'
with open(log_file, "w") as f:
subprocess.run([sys.executable, '-c', code_to_run], stdout=f, stderr=f)
```
The lowered log is similar to C source code, but the whole file is lengthy to show here. Let's just check the last few lines that are most important:
```
with open(log_file) as f:
log_lines = f.readlines()
log_lines[-10:]
```
The compiled code showed that the nodes of Add/Mul were fused into a single function, and vectorization was applied in the loop. The fusion was automatically done by the compiler in the Nuphar execution provider, and did not require any manual model editing.
Next, let's run inference on the model and compare the accuracy and performance with numpy:
```
seq = 128
batch = 16
input_data = np.random.rand(seq, batch, dim).astype(np.float32)
sess = onnxruntime.InferenceSession(simple_model_name)
feed = {X:input_data}
output = sess.run([], feed)
np_output = ((((input_data + input_data) * input_data) + input_data) * input_data) + input_data
assert np.allclose(output[0], np_output)
repeats = 100
start_ort = timer()
for i in range(repeats):
output = sess.run([], feed)
end_ort = timer()
start_np = timer()
for i in range(repeats):
np_output = ((((input_data + input_data) * input_data) + input_data) * input_data) + input_data
end_np = timer()
'onnxruntime: {0:.3f} seconds, numpy: {1:.3f} seconds'.format(end_ort - start_ort, end_np - start_np)
```
## 2. Create and run inference on a model using LSTM
Now, let's take one step further to work on a 4-layer LSTM model, created from onnxruntime.nuphar.rnn_benchmark module.
```
lstm_model = 'LSTMx4.onnx'
input_dim = 256
hidden_dim = 1024
generate_model('lstm', input_dim, hidden_dim, bidirectional=False, layers=4, model_name=lstm_model)
```
**IMPORTANT**: Nuphar generates code before knowing shapes of input data, unlike other execution providers that do runtime shape inference. Thus, shape inference information is critical for compiler optimizations in Nuphar. To do that, we run symbolic shape inference on the model. Symbolic shape inference is based on the ONNX shape inference, and enhanced by sympy to better handle Shape/ConstantOfShape/etc. ops using symbolic computation.
```
SymbolicShapeInference.infer_shapes(input_model=lstm_model, output_model=lstm_model)
```
Now, let's check baseline performance on the generated model, using CPU execution provider.
```
sess_baseline = onnxruntime.InferenceSession(lstm_model)
sess_baseline.set_providers(['CPUExecutionProvider']) # default provider in this container is Nuphar, this overrides to CPU EP
seq = 128
input_data = np.random.rand(seq, 1, input_dim).astype(np.float32)
feed = {sess_baseline.get_inputs()[0].name:input_data}
output = sess_baseline.run([], feed)
```
To run RNN models in Nuphar execution provider efficiently, LSTM/GRU/RNN ops need to be converted to Scan ops. This is because Scan is more flexible, and supports quantized RNNs.
```
scan_model = 'Scan_LSTMx4.onnx'
convert_to_scan_model(lstm_model, scan_model)
```
After conversion, let's compare performance and accuracy with baseline:
```
sess_nuphar = onnxruntime.InferenceSession(scan_model)
output_nuphar = sess_nuphar.run([], feed)
assert np.allclose(output[0], output_nuphar[0])
repeats = 10
start_baseline = timer()
for i in range(repeats):
output = sess_baseline.run([], feed)
end_baseline = timer()
start_nuphar = timer()
for i in range(repeats):
output = sess_nuphar.run([], feed)
end_nuphar = timer()
'nuphar: {0:.3f} seconds, baseline: {1:.3f} seconds'.format(end_nuphar - start_nuphar, end_baseline - start_baseline)
```
## 3. Quantize the LSTM model
Let's get more speed-ups from Nuphar by quantizing the floating point GEMM/GEMV in LSTM model to int8 GEMM/GEMV.
**NOTE:** For inference speed of quantizated model, a CPU with AVX2 instructions is preferred.
```
cpufeature.CPUFeature['AVX2'] or 'No AVX2, quantization model might be slow'
```
We can use onnxruntime.nuphar.model_quantizer to quantize floating point GEMM/GEMVs. Assuming GEMM/GEMV takes form of input * weights, weights are statically quantized per-column, and inputs are dynamically quantized per-row.
```
quantized_model = 'Scan_LSTMx4_int8.onnx'
convert_matmul_model(scan_model, quantized_model)
```
Now run the quantized model, and check accuracy. Please note that quantization may cause accuracy loss, so we relax the comparison threshold a bit.
```
sess_quantized = onnxruntime.InferenceSession(quantized_model)
output_quantized = sess_quantized.run([], feed)
assert np.allclose(output[0], output_quantized[0], rtol=1e-3, atol=1e-3)
```
Now check quantized model performance:
```
start_quantized = timer()
for i in range(repeats):
output = sess_quantized.run([], feed)
end_quantized = timer()
'quantized: {0:.3f} seconds, non-quantized: {1:.3f} seconds'.format(end_quantized - start_quantized, end_nuphar - start_nuphar)
```
## 4. Working on a real model: Bidirectional Attention Flow (BiDAF)
BiDAF is a machine comprehension model that uses LSTMs. The inputs to this model are paragraphs of contexts and queries, and the outputs are start/end indices of words in the contexts that answers the queries.
First let's download the model:
```
# download BiDAF model
cwd = os.getcwd()
bidaf_url = 'https://onnxzoo.blob.core.windows.net/models/opset_9/bidaf/bidaf.tar.gz'
bidaf_local = os.path.join(cwd, 'bidaf.tar.gz')
if not os.path.exists(bidaf_local):
urllib.request.urlretrieve(bidaf_url, bidaf_local)
with tarfile.open(bidaf_local, 'r') as f:
f.extractall(cwd)
```
Now let's check the performance of the CPU provider:
```
bidaf = os.path.join(cwd, 'bidaf', 'bidaf.onnx')
sess_baseline = onnxruntime.InferenceSession(bidaf)
sess_baseline.set_providers(['CPUExecutionProvider'])
# load test data
test_data_dir = os.path.join(cwd, 'bidaf', 'test_data_set_3')
tps = [onnx.load_tensor(os.path.join(test_data_dir, 'input_{}.pb'.format(i))) for i in range(len(sess_baseline.get_inputs()))]
feed = {tp.name:numpy_helper.to_array(tp) for tp in tps}
output_baseline = sess_baseline.run([], feed)
```
The context in this test data:
```
' '.join(list(feed['context_word'].reshape(-1)))
```
The query:
```
' '.join(list(feed['query_word'].reshape(-1)))
```
And the answer:
```
' '.join(list(feed['context_word'][output_baseline[0][0]:output_baseline[1][0]+1].reshape(-1)))
```
Now put all steps together:
```
# editing
bidaf_converted = 'bidaf_mod.onnx'
SymbolicShapeInference.infer_shapes(bidaf, bidaf_converted)
convert_to_scan_model(bidaf_converted, bidaf_converted)
# When quantizing, there's an only_for_scan option to quantize only the GEMV inside Scan ops.
# This is useful when the input dims of LSTM being much bigger than hidden dims.
# BiDAF has several LSTMs with input dim being 800/1400/etc, while hidden dim is 100.
# So unlike the LSTMx4 model above, we use only_for_scan here
convert_matmul_model(bidaf_converted, bidaf_converted, only_for_scan=True)
# inference and verify accuracy
sess = onnxruntime.InferenceSession(bidaf_converted)
output = sess.run([], feed)
assert all([np.allclose(o, ob) for o, ob in zip(output, output_baseline)])
```
Check performance after all these steps:
```
start_baseline = timer()
for i in range(repeats):
output = sess_baseline.run([], feed)
end_baseline = timer()
start_nuphar = timer()
for i in range(repeats):
output = sess.run([], feed)
end_nuphar = timer()
'nuphar: {0:.3f} seconds, baseline: {1:.3f} seconds'.format(end_nuphar - start_nuphar, end_baseline - start_baseline)
```
The benefit of quantization in BiDAF is not as great as in the LSTM sample above, because BiDAF has relatively small hidden dimensions, which limited the gain from optimization inside Scan ops. However, this model still benefits from fusion/vectorization/etc.
# 5. Ahead-Of-Time (AOT) compilation
Nuphar runs Just-in-time (JIT) compilation when loading models. The compilation may lead to slow cold start. We can use create_shared script to build dll from JIT code and accelerate model loading.
```
start_jit = timer()
sess = onnxruntime.InferenceSession(bidaf_converted)
end_jit = timer()
'JIT took {0:.3f} seconds'.format(end_jit - start_jit)
# create a folder for JIT cache
cache_dir = os.path.join(cwd, 'bidaf_cache')
# remove any stale cache files
if os.path.exists(cache_dir):
shutil.rmtree(cache_dir)
os.makedirs(cache_dir, exist_ok=True)
# use settings to enable JIT cache
settings = 'nuphar_cache_path:{}'.format(cache_dir)
onnxruntime.capi._pybind_state.set_nuphar_settings(settings)
sess = onnxruntime.InferenceSession(bidaf_converted)
```
Now object files of JIT code is stored in cache_dir, let's link them into dll:
```
cache_versioned_dir = os.path.join(cache_dir, os.listdir(cache_dir)[0])
# use onnxruntime.nuphar.create_shared module to create dll
onnxruntime_dir = os.path.split(os.path.abspath(onnxruntime.__file__))[0]
subprocess.run([sys.executable, '-m', 'onnxruntime.nuphar.create_shared', '--input_dir', cache_versioned_dir], check=True)
os.listdir(cache_versioned_dir)
```
Check the model loading speed-up with AOT dll:
```
start_aot = timer()
# NOTE: Nuphar settings string is not sticky. It needs to be reset before creating InferenceSession
settings = 'nuphar_cache_path:{}'.format(cache_dir)
onnxruntime.capi._pybind_state.set_nuphar_settings(settings)
sess = onnxruntime.InferenceSession(bidaf_converted)
end_aot = timer()
'AOT took {0:.3f} seconds'.format(end_aot - start_aot)
```
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.