text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
# Atelier web avec Python
L'objectif de cet atelier est de travailler sur un cas concret. Il s'agit de trouver une solution à un petit problème et de le mettre en oeuvre avec Python, ce n'est pas une présentation générale du web avec Python ou même d'un framework.
Quel est ce problème ? Guersande a bossé sur les partitifs en finnois à partir du corpus [Europarl](http://www.statmt.org/europarl/). Le résultat de son travail est présenté dans une page web : [apps.lattice.cnrs.fr/partitif](http://apps.lattice.cnrs.fr/partitif)
Rien à dire sur le travail mené, ni même sur la présentation des résultats mais la page web générée pèse 319 Mo 😨😨😨. C'est trop, beaucoup trop.
Notre problème à résoudre est : **comment réduire la taille du fichier html sans trop modifier sa structure et sa présentation ?**
Une solution aurait pu être de paginer les résultats, ce qui reviendrait à avoir plusieurs pages html statiques au lieu d'une. C'est facile à mettre en oeuvre et cela résoud le problème mais l'utilisateur perd la vue sur l'intégralité des résultats. Et puis avec cette solution ce notebook n'aurait pas lieu d'être.
Celle que l'on va essayer de mettre en oeuvre ne change rien pour l'utilisateur : il a tous les lemmes sur une seule page et il peut afficher les exemples à la demande.
Les exemples c'est précisement là-dessus que l'on va opérer le changement. Dans la version actuelle les exemples sont cachés/montrés avec le clic de l'utilisateur mais ils sont inclus dans le code html, c'est même eux qui sont en grande partie responsables de la taille de la page. L'idée ici est de les extraire de la page afin de l'alléger et de les inclure uniquement lors du clic de l'utilisateur.
## Extraction des exemples
Le premier truc qu'on va devoir faire sur la page est d'extraire les exemples de la page web et les stocker autre part, on verra où après.
Pour s'exercer on va travailler sur un fichier réduit, on passera au gros fichier une fois que tout sera au point. Le fichier de travail est `test.html`.
Je donne la trame, à vous de compléter :
```
from bs4 import BeautifulSoup
import json
soup = BeautifulSoup(open("test.html"), 'html.parser')
divs = soup.find_all('div', class_="structure")
verbs = dict()
for div in divs:
verbs[div['id']] = list()
examples = div.find_all('div', class_="exemple")
print("{} examples divs found for verb {}".format(len(examples), div['id']))
```
## HTML + JSON : et maintenant ?
On a une page html avec une taille décente d'un côté (476 Ko) et un fichier json (230 Mo) avec les exemples de l'autre. Super. Notez qu'un passage on a bien élagué en supprimant le balisage html.
Maintenant ben il va falloir adapter le "voir/masquer les exemples" de la page originale, le rendre dynamique en incluant le code html des exemples à la volée. Ça se fait en Javascript grâce à la techno [AJAX](https://fr.wikipedia.org/wiki/Ajax_%28informatique%29), et pour se simplifier la vie on va utiliser la bibliothèque JQuery.
### Modification du DOM
Pour jouer avec JS le plus pratique est d'utiliser les outils inclus dans le navigateur. Dans Chrome ça s'appelle 'Developper tools' (ctrl+maj+I ou clic droit 'Inspect'), allez dans l'onglet 'Sources', ctrl+P et sélectionnez le fichier à éditer. Pour nous ce sera `flipflop.js`
Deux fonctions nous intéressent dans ce fichier :
* `flipflipON` est appelée par un clic sur 'voir les exemples'
* `flipflopOFF` est appelée par un clic sur 'cacher les exemples'
Nous allons modifier la fonction `flipflopON` et utliser la fonction JQuery `append` pour ajouter du contenu à l'élément sélectionné. Modifiez le code directement dans le naivigateur pour voir ce que ça donne.
```javascript
function flipflopON(id1,id2)
{
if (document.getElementById(id1).style.display == "none") {
document.getElementById(id1).style.display = "block";
document.getElementById(id2).style.display = "none";
$('#'+id1).append("<div class=\"exemple\">Youhou !</div>")
}
else {document.getElementById(id1).style.display = "none";
document.getElementById(id2).style.display = "block";
}
}
```
C'est cool ça marche mais si vous "voir/masquer" plusieurs fois vous verrez que les éléments ajoutés sont conservés. Il va falloir modifier aussi la fonction `flipflopOFF` pour faire le ménage.
```javascript
function flipflopOFF(id1,id2)
{
if (document.getElementById(id1).style.display == "none") {
document.getElementById(id1).style.display = "block";
document.getElementById(id2).style.display = "none";
$('#'+id2+">div.exemple").remove();
}
else {document.getElementById(id1).style.display = "block";
document.getElementById(id2).style.display = "none";
}
}
```
### Conversion du json en html
C'est bien c'est bien mais nous on a du json en entrée. Il nous faut une fonction qui transforme du json dans l'html désiré. Au travail.
Voici un exemple d'entrée JSON pour 3 exemples du verbe 'keskittää' :
```json
[{"sp": " keskittää + keskustelu (Nom\n)", "phrase": "Olisi siis virhe keskittää Euroopan parlamentin vuoden 2001 talousarviota koskeva keskustelu pelkästään tähän kysymykseen. "},{"sp": " keskittää + kaikki (Nom\n)", "phrase":"Keskititte kaikki tietenkin puheenvuoronne rauhanprosessiin ja neuvotteluihin, jotka ovat epäilemättä edistymisen avaintekijöitä tällä Lähi-idän alueella. "}, {"sp": " keskittää + puheen#vuoro (Nom\n)", "phrase":"Keskititte kaikki tietenkin puheenvuoronne rauhanprosessiin ja neuvotteluihin, jotka ovat epäilemättä edistymisen avaintekijöitä tällä Lähi-idän alueella. "}]
```
Qui devra donner en html :
```
%%html
<div class="exemple">
<div class="sp">keskittää + keskustelu (Nom\n)</div>
<div class="phrase">Olisi siis virhe keskittää Euroopan parlamentin vuoden 2001 talousarviota koskeva keskustelu pelkästään tähän kysymykseen.</div>
</div>
```
Pour cela nous allons utiliser la fonction [each](http://api.jquery.com/jquery.each/) de JQuery qui permet d'itérer sur des objets ou des tableaux.
Copiez-collez dans Developper Tools dans le navigateur et regardez ce que ça donne.
```javascript
function flipflopON(id1,id2)
{
examples = [{"sp": " keskittää + keskustelu (Nom\n)", "phrase": "Olisi siis virhe keskittää Euroopan parlamentin vuoden 2001 talousarviota koskeva keskustelu pelkästään tähän kysymykseen. "},{"sp": " keskittää + kaikki (Nom\n)", "phrase":"Keskititte kaikki tietenkin puheenvuoronne rauhanprosessiin ja neuvotteluihin, jotka ovat epäilemättä edistymisen avaintekijöitä tällä Lähi-idän alueella. "}, {"sp": " keskittää + puheen#vuoro (Nom\n)", "phrase":"Keskititte kaikki tietenkin puheenvuoronne rauhanprosessiin ja neuvotteluihin, jotka ovat epäilemättä edistymisen avaintekijöitä tällä Lähi-idän alueella. "}]
if (document.getElementById(id1).style.display == "none") {
document.getElementById(id1).style.display = "block";
document.getElementById(id2).style.display = "none";
$.each(examples, function(index, example){
ex_elem = $('<div/>', {'class': 'exemple'}).append(
$('<div/>', {'class': 'sp', 'text': example.sp}),
$('<div/>', {'class': 'phrase', 'text': example.phrase})
);
$('#'+id1).append(ex_elem);
});
}
else {document.getElementById(id1).style.display = "none";
document.getElementById(id2).style.display = "block";
}
}
```
À ce stade nous sommes capables d'afficher les exemples à partir d'une donnée JSON : parcourir le JSON, générer le code html pour emballer les données et modifier le DOM. Tout ça piloté par les clics de l'utilisateur.
### Service Python
On avance mais notre exemple fonctionne avec une donnée en dur. Ce qu'il nous faut maintenant c'est récupérer le tableau d'exemples correspondants à un verbe donné. Cela signifie :
1. Charger le fichier `verbs.json`, trouver le tableau des exemples pour un verbe donné et le renvoyer. Fonction Python, facile.
2. "Servir" le résultat en json. Avec Flask, pas bien compliqué.
3. Appeler le serveur flask en JS côté client
Un peu à l'arrache on peut faire un truc comme ça avec Flask :
```
from flask import Flask, jsonify
import json
examples = json.load(open("examples.json"))
app = Flask(__name__)
@app.route('/<verb>')
def hello_world(verb):
if verb in examples:
res = examples[verb]
else:
res = ""
return jsonify(res)
if __name__ == '__main__':
app.run()
```
C'est tout ? Oui (enfin presque). Essayez avec les url http://localhost:5000/pilata, http://localhost:5000/keskittää, http://localhost:5000/machin
Ce n'est pas vraiment la solution idéale, dans la vraie vie on pourra utiliser [redis](https://redis.io/) pour éviter de charger les 230 Mo de json en mémoire avec Flask mais ça ajoute (encore) une techno. Ça se discute, c'est selon les compétences et les habitudes du dév.
### Et côté client ?
JQuery nous facilite bien la vie, encore une fois on y trouve la fonction qu'il nous faut : [ajax](http://api.jquery.com/jquery.ajax/)
```javascript
function flipflopON(id1,id2)
{
if (document.getElementById(id1).style.display == "none") {
document.getElementById(id1).style.display = "block";
document.getElementById(id2).style.display = "none";
verb = id1.substring(7) //contenupilata -> pilata
$.ajax({
url: 'http://localhost:5000/'+verb,
type: "GET",
success: function(examples){
$.each(examples, function(index, example){
ex_elem = $('<div/>', {'class': 'exemple'}).append(
$('<div/>', {'class': 'sp', 'text': example.sp}),
$('<div/>', {'class': 'phrase', 'text': example.phrase})
);
$('#'+id1).append(ex_elem);
});
}
});
}
else{
document.getElementById(id1).style.display = "none";
document.getElementById(id2).style.display = "block";
}
}
```
Et ça marche !!
Ça marche en local en tout cas. Le plus compliqué dans tout ça est sans doute le déploiement sur un serveur de production. Flask ne **doit pas** être utilisé en prod directement, voir http://flask.pocoo.org/docs/0.12/deploying/#deployment
Si on a la main sur le serveur Gunicorn est une option fiable et facile à mettre en oeuvre.
Il reste d'autres questions relatives au déploiement mais elles sont un peu hors sujet ici.
| github_jupyter |
# Analyzing a double hanger resonator (S Param)
### Prerequisite
You must have a working local installation of Ansys.
```
%load_ext autoreload
%autoreload 2
import qiskit_metal as metal
from qiskit_metal import designs, draw
from qiskit_metal import MetalGUI, Dict, Headings
import pyEPR as epr
```
## Create the design in Metal
Set up a design of a given dimension. Dimensions will be respected in the design rendering.
<br>
Note the chip design is centered at origin (0,0).
```
design = designs.DesignPlanar({}, True)
design.chips.main.size['size_x'] = '2mm'
design.chips.main.size['size_y'] = '2mm'
#Reference to Ansys hfss QRenderer
hfss = design.renderers.hfss
gui = MetalGUI(design)
```
Perform the necessary imports.
```
from qiskit_metal.qlibrary.couplers.coupled_line_tee import CoupledLineTee
from qiskit_metal.qlibrary.tlines.meandered import RouteMeander
from qiskit_metal.qlibrary.qubits.transmon_pocket import TransmonPocket
from qiskit_metal.qlibrary.tlines.straight_path import RouteStraight
from qiskit_metal.qlibrary.terminations.open_to_ground import OpenToGround
```
Add 2 transmons to the design.
```
options = dict(
# Some options we want to modify from the deafults
# (see below for defaults)
pad_width = '425 um',
pocket_height = '650um',
# Adding 4 connectors (see below for defaults)
connection_pads=dict(
a = dict(loc_W=+1,loc_H=+1),
b = dict(loc_W=-1,loc_H=+1, pad_height='30um'),
c = dict(loc_W=+1,loc_H=-1, pad_width='200um'),
d = dict(loc_W=-1,loc_H=-1, pad_height='50um')
)
)
## Create 2 transmons
q1 = TransmonPocket(design, 'Q1', options = dict(
pos_x='+1.4mm', pos_y='0mm', orientation = '90', **options))
q2 = TransmonPocket(design, 'Q2', options = dict(
pos_x='-0.6mm', pos_y='0mm', orientation = '90', **options))
gui.rebuild()
gui.autoscale()
```
Add 2 hangers consisting of capacitively coupled transmission lines.
```
TQ1 = CoupledLineTee(design, 'TQ1', options=dict(pos_x='1mm',
pos_y='3mm',
coupling_length='200um'))
TQ2 = CoupledLineTee(design, 'TQ2', options=dict(pos_x='-1mm',
pos_y='3mm',
coupling_length='200um'))
gui.rebuild()
gui.autoscale()
```
Add 2 meandered CPWs connecting the transmons to the hangers.
```
ops=dict(fillet='90um')
design.overwrite_enabled = True
options1 = Dict(
total_length='8mm',
hfss_wire_bonds = True,
pin_inputs=Dict(
start_pin=Dict(
component='TQ1',
pin='second_end'),
end_pin=Dict(
component='Q1',
pin='a')),
lead=Dict(
start_straight='0.1mm'),
**ops
)
options2 = Dict(
total_length='9mm',
hfss_wire_bonds = True,
pin_inputs=Dict(
start_pin=Dict(
component='TQ2',
pin='second_end'),
end_pin=Dict(
component='Q2',
pin='a')),
lead=Dict(
start_straight='0.1mm'),
**ops
)
meanderQ1 = RouteMeander(design, 'meanderQ1', options=options1)
meanderQ2 = RouteMeander(design, 'meanderQ2', options=options2)
gui.rebuild()
gui.autoscale()
```
Add 2 open to grounds at the ends of the horizontal CPW.
```
otg1 = OpenToGround(design, 'otg1', options = dict(pos_x='3mm',
pos_y='3mm'))
otg2 = OpenToGround(design, 'otg2', options = dict(pos_x = '-3mm',
pos_y='3mm',
orientation='180'))
gui.rebuild()
gui.autoscale()
```
Add 3 straight CPWs that comprise the long horizontal CPW.
```
ops_oR = Dict(hfss_wire_bonds = True,
pin_inputs=Dict(
start_pin=Dict(
component='TQ1',
pin='prime_end'),
end_pin=Dict(
component='otg1',
pin='open')))
ops_mid = Dict(hfss_wire_bonds = True,
pin_inputs=Dict(
start_pin=Dict(
component='TQ1',
pin='prime_start'),
end_pin=Dict(
component='TQ2',
pin='prime_end')))
ops_oL = Dict(hfss_wire_bonds = True,
pin_inputs=Dict(
start_pin=Dict(
component='TQ2',
pin='prime_start'),
end_pin=Dict(
component='otg2',
pin='open')))
cpw_openRight = RouteStraight(design, 'cpw_openRight', options=ops_oR)
cpw_middle = RouteStraight(design, 'cpw_middle', options=ops_mid)
cpw_openLeft = RouteStraight(design, 'cpw_openLeft', options=ops_oL)
gui.rebuild()
gui.autoscale()
```
## Render the qubit from Metal into the HangingResonators design in Ansys. <br>
Open a new Ansys window, connect to it, and add a driven modal design called HangingResonators to the currently active project.<br>
If Ansys is already open, you can skip `hfss.open_ansys()`. <br>
**Wait for Ansys to fully open before proceeding.**<br> If necessary, also close any Ansys popup windows.
```
#hfss.open_ansys()
hfss.connect_ansys()
hfss.activate_drivenmodal_design("HangingResonators")
```
Set the buffer width at the edge of the design to be 0.5 mm in both directions.
```
hfss.options['x_buffer_width_mm'] = 0.5
hfss.options['y_buffer_width_mm'] = 0.5
```
Here, pin cpw_openRight_end and cpw_openLeft_end are converted into lumped ports, each with an impedance of 50 Ohms. <br>
Neither of the junctions in Q1 or Q2 are rendered. <br>
As a reminder, arguments are given as <br><br>
First parameter: List of components to render (empty list if rendering whole Metal design) <br>
Second parameter: List of pins (qcomp, pin) with open endcaps <br>
Third parameter: List of pins (qcomp, pin, impedance) to render as lumped ports <br>
Fourth parameter: List of junctions (qcomp, qgeometry_name, impedance, draw_ind) to render as lumped ports or as lumped port in parallel with a sheet inductance <br>
Fifth parameter: List of junctions (qcomp, qgeometry_name) to omit altogether during rendering
Sixth parameter: Whether to render chip via box plus buffer or fixed chip size
```
hfss.render_design([],
[],
[('cpw_openRight', 'end', 50), ('cpw_openLeft', 'end', 50)],
[],
[('Q1', 'rect_jj'), ('Q2', 'rect_jj')],
True)
hfss.save_screenshot()
hfss.add_sweep(setup_name="Setup",
name="Sweep",
start_ghz=4.0,
stop_ghz=8.0,
count=2001,
type="Interpolating")
hfss.analyze_sweep('Sweep', 'Setup')
```
Plot S, Y, and Z parameters as a function of frequency. <br>
The left and right plots display the magnitude and phase, respectively.
```
hfss.plot_params(['S11', 'S21'])
hfss.plot_params(['Y11', 'Y21'])
hfss.plot_params(['Z11', 'Z21'])
hfss.disconnect_ansys()
gui.main_window.close()
```
| github_jupyter |
```
epochs = 20
n_train_items = 6000
train_item_per_client = 600
rounds = 1
C = 0.1
total_client = 100
n_workers = int(total_client * C)
epsilon = 2
delta = 0.001
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from torchvision import datasets, transforms
import numpy as np
# from torch.utils.tensorboard import SummaryWriter
import random
import syft as sy # <-- NEW: import the Pysyft library
hook = sy.TorchHook(torch) # <-- NEW: hook PyTorch ie add extra functionalities to support Federated Learning
# simulation functions
def connect_to_workers(n_workers):
return [
sy.VirtualWorker(hook, id=f"worker{i+1}")
for i in range(n_workers)
]
workers = connect_to_workers(n_workers=n_workers)
class Arguments():
def __init__(self):
self.batch_size = 10
self.test_batch_size = 60
self.epochs = epochs
self.rounds = rounds
self.lr = 0.02
self.momentum = 0.5
self.no_cuda = False
self.seed = 0
self.log_interval = 4
self.save_model = False
self.n_train_items = n_train_items
self.train_item_per_client = train_item_per_client
args = Arguments()
use_cuda = not args.no_cuda and torch.cuda.is_available()
torch.manual_seed(args.seed)
device = torch.device("cuda" if use_cuda else "cpu")
kwargs = {'num_workers': 1, 'pin_memory': True} if use_cuda else {}
# federated_train_loader = sy.FederatedDataLoader( # <-- this is now a FederatedDataLoader
# datasets.MNIST('../data', train=True, download=True,
# transform=transforms.Compose([
# transforms.ToTensor(),
# transforms.Normalize((0.1307,), (0.3081,))
# ]))
# .federate(workers), # <-- NEW: we distribute the dataset across all the workers, it's now a FederatedDataset
# batch_size=args.batch_size, shuffle=True, **kwargs)
# test_loader = torch.utils.data.DataLoader(
# datasets.MNIST('../data', train=False, transform=transforms.Compose([
# transforms.ToTensor(),
# transforms.Normalize((0.1307,), (0.3081,))
# ])),
# batch_size=args.test_batch_size, shuffle=True, **kwargs)
train_loader = torch.utils.data.DataLoader(
datasets.MNIST('../data', train=True, download=True, transform=transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.1307,), (0.3081,))
])),
batch_size=args.batch_size
)
test_loader = torch.utils.data.DataLoader(
datasets.MNIST('../data', train=False, download=True, transform=transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.1307,), (0.3081,))
])),
batch_size=args.test_batch_size
)
#---
def create_dataset_for_client(train_loader):
train_dataloader_client = []
tmp = []
for count, batch_data in enumerate(train_loader):
if (count * args.batch_size) % (args.train_item_per_client) == 0:
#Data in tmp is equal to the train_item_per_client
train_dataloader_client.append(tmp)
tmp = []
else :
#Keep appending the data into tmp
tmp.append(batch_data)
return train_dataloader_client
train_dataloader_client = create_dataset_for_client(train_loader)
# less_train_dataloader[0][0].shape
len(train_dataloader_client)
# from PIL import Image
# import numpy
# #mnist_dataset.__getitem__(2)[1]
# a = (mnist_dataset.__getitem__(0)[0]).numpy()
# a.dtype = 'uint8'
# print(a)
# Image.fromarray(a[0], mode= 'P')
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
self.conv1 = nn.Conv2d(1, 32, 5, 1)
self.conv2 = nn.Conv2d(32, 64, 5, 1)
self.fc1 = nn.Linear(3136, 512)
self.fc2 = nn.Linear(512, 10)
self.same_padding = nn.ReflectionPad2d(2)
# self.batch32 = nn.BatchNorm2d(32)
# self.batch64 = nn.BatchNorm2d(64)
def forward(self, x):
x = self.same_padding(x)
x = F.relu(self.conv1(x))
x = F.max_pool2d(x, 2, 2)
# x = self.batch32(x)
x = self.same_padding(x)
x = F.relu(self.conv2(x))
# x = self.batch64(x)
x = F.max_pool2d(x, 2, 2)
x = x.view(-1, 3136)
x = F.relu(self.fc1(x))
x = self.fc2(x)
return F.log_softmax(x, dim=1)
def model_init(workers, Net):
model_list = list()
for worker in workers:
model_list.append(Net)
return model_list
def opt_init(model_list):
opt_list = list()
for model in model_list:
opt_list.append(optim.SGD(model.parameters(), lr=args.lr))
return opt_list
def random_sample(train_dataloader):
choice_list = sorted(random.sample(range(100), 10))
count = 0
tmp = []
for i, data in enumerate(train_dataloader):
if i == choice_list[count]:
tmp.append(data)
if count == 9:
pass
else:
count += 1
return tmp
def train(args, device, train_loader, opt_list, workers):
global model_list
## start training and record the model into model_list
less_train_dataloader = random_sample(train_loader)
for epoch in range(args.epochs):
for client_data in less_train_dataloader:
for batch_idx, (data, target) in enumerate(client_data): # <-- now it is a distributed dataset
model_on_worker = model_list[batch_idx%len(workers)]
model_on_worker.train()
model_on_worker.send(workers[batch_idx%len(workers)]) # <-- NEW: send the model to the right location
data_on_worker = data.send(workers[batch_idx%len(workers)])
target_on_worker = target.send(workers[batch_idx%len(workers)])
data_on_worker, target_on_worker = data_on_worker.to(device), target_on_worker.to(device)
opt_list[batch_idx%len(workers)].zero_grad()
output = model_on_worker(data_on_worker)
loss = F.cross_entropy(output, target_on_worker)
loss.backward()
opt_list[batch_idx%len(workers)].step()
model_on_worker.get() # <-- NEW: get the model back
model_list[batch_idx%len(workers)] = model_on_worker #When len(dataloader) is longer than the len(worker) send and get must be modified
#model_list here is full of the model which has trained on the workers, there are all different now.
if epoch % args.log_interval == 0:
loss = loss.get() # <-- NEW: get the loss back
print('Train Epoch: {}/{} ({:.0f}%)\tLoss: {:.6f}'.format(
epoch, args.epochs ,
100. * epoch / args.epochs, loss.item()))
##Aggregation time
new_model = []
tmp_model = Net().to(device)
with torch.no_grad():
for p in model_list[0].parameters():
new_model.append(0)
for m in model_list:
# We will sum all the model in model)list
for par_idx, par in enumerate(m.parameters()):
#average the model_list
new_model[par_idx] = new_model[par_idx]+par.data
# we get new model in list format and need to set_ to model
# add gussian noise before testing
for param in new_model.parameters():
param.add_(torch.normal(0,1.3*1.5,param.size(),device=device)*args.lr/args.batch_size)
param.add_(torch.normal(0,1.3*1.5,param.size(),device=device)*args.lr/args.batch_size)
# param.add_(torch.from_numpy(np.random.normal(0,1.3*1.5,param.size())*args.lr).to(device))
for worker in range(len(workers)):
for par_idx in range(len(new_model)):
list(model_list[worker].parameters())[par_idx].set_(new_model[par_idx]/len(workers))
#init model with new_model
def test(args, model, device, test_loader, r):C_wn
model.eval()
test_loss = 0
correct = 0
with torch.no_grad():
for data, target in test_loader:
data, target = data.to(device), target.to(device)
output = model(data)
test_loss += F.cross_entropy(output, target, reduction='sum').item() # sum up batch loss
pred = output.argmax(1, keepdim=True) # get the index of the max log-probability
correct += pred.eq(target.view_as(pred)).sum().item()
test_loss /= len(test_loader)*(args.test_batch_size)
accuracy = 100. * correct / (len(test_loader)*args.test_batch_size)
#Since the test loader here is a list, we can get the len by * it with batch.size
# writer.add_scalar('Accuracy', accuracy,r)
# writer.add_scalar('Loss', test_loss, r)
print('\nTest set round{}: Average loss: {:.4f}, Accuracy: {}/{} ({:.2f}%)\n'.format(
r, test_loss, correct, len(test_loader)* (args.test_batch_size),
accuracy))
# logdir = '/root/notebooks/tensorflow/logs/pysyft_shuffle6000_crossE'
# writer = SummaryWriter(logdir)
#test
# model_test = Net()
# output_test = model_test(less_train_dataloader[0][0])
# loss_test = F.cross_entropy(output_test, less_train_dataloader[0][1])
# print(loss_test)
%%time
#optimizer = optim.SGD(model.parameters(), lr=args.lr) # TODO momentum is not supported at the moment
model_list = []
model_list = model_init(workers, Net().to(device))
opt_list = opt_init(model_list)
# not finish in train, finish latter
pars = [list(model.parameters()) for model in model_list]
for r in range(1, args.rounds + 1):
train(args, device, train_dataloader_client, opt_list, workers)
print("After training")
test(args, model_list[0], device, test_loader, r)
if (args.save_model):
torch.save(model.state_dict(), "mnist_cnn.pt")
```
| github_jupyter |
```
cd ..
#source: https://www.kaggle.com/bhaveshsk/getting-started-with-titanic-dataset/data
#data analysis and wrangling
import pandas as pd
import numpy as np
import random as rnd
#data visualization
import seaborn as sns
import matplotlib.pyplot as plt
%matplotlib inline
#machine learning packages
from sklearn.linear_model import LogisticRegression
from sklearn.svm import SVC
from sklearn.ensemble import RandomForestClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.naive_bayes import GaussianNB
from sklearn.linear_model import Perceptron
from sklearn.linear_model import SGDClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn import metrics
train_df = pd.read_csv("./input/train.csv")
test_df = pd.read_csv("./input/test.csv")
df = pd.concat([train_df,test_df], sort=True)
df.head()
from src.preprocessing import add_derived_title
df = add_derived_title(df)
df['Title'] = df['Title'].map({"Mr": 1, "Miss": 2, "Mrs": 3, "Master": 4, "Rare": 5}).fillna(0)
freq_port = df.Embarked.dropna().mode()[0]
df['Embarked'] = df['Embarked'].fillna(freq_port)
# EXERCISE 2: Write a unit test and extract the following implementation into a function:
# df = impute_nans(df, columns)
# 'Fare' column
df['Fare'] = df['Fare'].fillna(df['Fare'].dropna().median())
# 'Age' column
df['Age'] = df['Age'].fillna(df['Age'].dropna().median())
df['Sex'] = df['Sex'].map( {'female': 1, 'male': 0} ).astype(int)
df['AgeBand'] = pd.cut(df['Age'], 5)
df.loc[ df['Age'] <= 16, 'Age'] = 0
df.loc[(df['Age'] > 16) & (df['Age'] <= 32), 'Age'] = 1
df.loc[(df['Age'] > 32) & (df['Age'] <= 48), 'Age'] = 2
df.loc[(df['Age'] > 48) & (df['Age'] <= 64), 'Age'] = 3
df = df.drop(['AgeBand'], axis=1)
# EXERCISE 3: Write a unit test and extract the following implementation into a function:
# df = add_is_alone_column(df)
df['FamilySize'] = df['SibSp'] + df['Parch'] + 1
df['IsAlone'] = 0
df.loc[df['FamilySize'] == 1, 'IsAlone'] = 1
# drop unused columns
df = df.drop(['Parch', 'SibSp', 'FamilySize'], axis=1)
df = df.drop(['Ticket', 'Cabin'], axis=1)
df = df.drop(['Name', 'PassengerId'], axis=1)
df['Age*Class'] = df.Age * df.Pclass
df['Embarked'] = df['Embarked'].map( {'S': 0, 'C': 1, 'Q': 2} ).astype(int)
df['FareBand'] = pd.qcut(df['Fare'], 4)
df.loc[ df['Fare'] <= 7.91, 'Fare'] = 0
df.loc[(df['Fare'] > 7.91) & (df['Fare'] <= 14.454), 'Fare'] = 1
df.loc[(df['Fare'] > 14.454) & (df['Fare'] <= 31), 'Fare'] = 2
df.loc[ df['Fare'] > 31, 'Fare'] = 3
df['Fare'] = df['Fare'].astype(int)
df = df.drop(['FareBand'], axis=1)
train_df = df[-df['Survived'].isna()]
test_df = df[df['Survived'].isna()]
test_df = test_df.drop('Survived', axis=1)
X_train = train_df.drop("Survived", axis=1)
Y_train = train_df["Survived"]
X_test = test_df.copy()
# EXERCISE 1: Create a function, train_model(...), to eliminate the duplication in the next few cells
from src.preprocessing import train_model
_, acc_svc = train_model(SVC, X_train, Y_train, gamma='auto')
_, acc_knn = train_model(KNeighborsClassifier, X_train, Y_train, n_neighbors=3)
_, acc_gaussian = train_model(GaussianNB, X_train, Y_train)
_, acc_perceptron = train_model(Perceptron, X_train, Y_train)
_, acc_sgd = train_model(SGDClassifier, X_train, Y_train)
_, acc_decision_tree = train_model(DecisionTreeClassifier, X_train, Y_train)
_, acc_random_forest = train_model(RandomForestClassifier, X_train, Y_train, n_estimators=100)
models = pd.DataFrame({
'Model': ['Support Vector Machines', 'KNN',
'Random Forest', 'Naive Bayes', 'Perceptron',
'Stochastic Gradient Decent',
'Decision Tree'],
'Score': [acc_svc, acc_knn,
acc_random_forest, acc_gaussian, acc_perceptron,
acc_sgd, acc_decision_tree]})
models.sort_values(by='Score', ascending=False)
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from gaussian_processes_util import plot_gp_2D
noise_2D = 0.1
rx, ry = np.arange(-5, 5, 0.3), np.arange(-5, 5, 0.3)
gx, gy = np.meshgrid(rx, rx)
X_2D = np.c_[gx.ravel(), gy.ravel()]
Y_2D_truth = np.sin(0.5 * np.linalg.norm(X_2D, axis=1))
X_2D_train = np.random.uniform(-4, 4, (100, 2))
Y_2D_train = np.sin(0.5 * np.linalg.norm(X_2D_train, axis=1)) + \
noise_2D * np.random.randn(len(X_2D_train))
sinewave1 = pd.DataFrame(X_2D_train, columns=['x1', 'x2'])
sinewave2 = pd.DataFrame(Y_2D_train, columns=['y'])
sinewave =pd.concat([sinewave1, sinewave2], axis=1)
plt.figure(figsize=(14,7))
plot_gp_2D(gx, gy, Y_2D_truth, X_2D_train, Y_2D_train,
f'3D Sinewave and Noisy Samples', 1)
import sys
sys.path.insert(0, '\\\\newwinsrc\\sasgen\\dev\\mva-vb023\\GTKWX6ND\\misc\\python') # location of src
import swat as sw
s = sw.CAS('rdcgrd327.unx.sas.com', 29640, nworkers=2)
s.sessionprop.setsessopt(caslib='CASUSER',timeout=31535000)
if s.tableexists('sinewave').exists:
s.CASTable('sinewave').droptable()
dataset = s.upload_frame(sinewave,
importoptions=dict(vars=[dict(type='double'),
dict(type='double'),
dict(type='double')
]),
casout=dict(name='sinewave', promote=True))
s.loadactionset(actionset="nonParametricBayes")
s.gpreg(
table={"name":"sinewave"},
inputs={"x1","x2"},
target="y",
seed=1234,
nInducingPoints=7,
fixInducingPoints=False,
kernel="RBF",
partbyfrac={"valid":0, "test":0, "seed":1235},
nloOpts={"algorithm":"ADAM",
"optmlOpt":{"maxIters":91},
"sgdOpt":{"learningRate":0.15,
"momentum":0.8,
"adaptiveRate":True,
"adaptiveDecay":0.9,
"miniBatchSize":100
},
"printOpt":{"printFreq":10}
},
output={"casout":{"name":"GpReg_Pred", "replace":True}, "copyvars":"ALL"},
outInducingPoints={"name":"GpReg_inducing", "replace":True},
outVariationalCov={"name":"GpReg_S", "replace":True},
saveState={"name":"gpregStore", "replace":True}
)
s.gpreg(
table={"name":"sinewave"},
inputs={"x1","x2"},
target="y",
seed=1234,
nInducingPoints=25,
fixInducingPoints=True,
kernel="RBF",
partbyfrac={"valid":0, "test":0, "seed":1235},
nloOpts={"algorithm":"ADAM",
"optmlOpt":{"maxIters":1},
"sgdOpt":{"learningRate":0.15,
"momentum":0.8,
"adaptiveRate":True,
"adaptiveDecay":0.9,
"miniBatchSize":100
},
"printOpt":{"printFreq":1}
},
output={"casout":{"name":"GpReg_Pred1", "replace":True}, "copyvars":"ALL"},
outInducingPoints={"name":"GpReg_inducing1", "replace":True},
outVariationalCov={"name":"GpReg_S1", "replace":True},
saveState={"name":"gpregStore1", "replace":True}
)
sinewave_test = pd.DataFrame(X_2D, columns=['x1', 'x2'])
if s.tableexists('sinewave_test').exists:
s.CASTable('sinewave_test').droptable()
dataset = s.upload_frame(sinewave_test,
importoptions=dict(vars=[dict(type='double'),
dict(type='double')
]),
casout=dict(name='sinewave_test', promote=True))
if s.tableexists('test_pred').exists:
s.CASTable('test_pred').droptable()
if s.tableexists('test_pred1').exists:
s.CASTable('test_pred1').droptable()
s.loadactionset('aStore')
s.score(
table='sinewave_test',
out='test_pred',
rstore='gpregStore',
)
pred = s.CASTable("test_pred").to_frame()
s.loadactionset('aStore')
s.score(
table='sinewave_test',
out='test_pred1',
rstore='gpregStore1',
)
pred1 = s.CASTable("test_pred1").to_frame()
mu = pred['P_yMEAN'].values
mu1 = pred1['P_yMEAN'].values
plt.figure(figsize=(14,7))
plot_gp_2D(gx, gy, mu1, X_2D_train, Y_2D_train,
f'Prediction after 1st Iteration', 1)
plot_gp_2D(gx, gy, mu, X_2D_train, Y_2D_train,
f'Prediction After 201 Iterations', 2)
```
| github_jupyter |
# Multiclass classification with Amazon SageMaker XGBoost algorithm
_**Single machine and distributed training for multiclass classification with Amazon SageMaker XGBoost algorithm**_
---
---
## Contents
1. [Introduction](#Introduction)
2. [Prerequisites and Preprocessing](#Prequisites-and-Preprocessing)
1. [Permissions and environment variables](#Permissions-and-environment-variables)
2. [Data ingestion](#Data-ingestion)
3. [Data conversion](#Data-conversion)
3. [Training the XGBoost model](#Training-the-XGBoost-model)
1. [Training on a single instance](#Training-on-a-single-instance)
2. [Training on multiple instances](#Training-on-multiple-instances)
4. [Set up hosting for the model](#Set-up-hosting-for-the-model)
1. [Import model into hosting](#Import-model-into-hosting)
2. [Create endpoint configuration](#Create-endpoint-configuration)
3. [Create endpoint](#Create-endpoint)
5. [Validate the model for use](#Validate-the-model-for-use)
---
## Introduction
This notebook demonstrates the use of Amazon SageMaker’s implementation of the XGBoost algorithm to train and host a multiclass classification model. The MNIST dataset is used for training. It has a training set of 60,000 examples and a test set of 10,000 examples. To illustrate the use of libsvm training data format, we download the dataset and convert it to the libsvm format before training.
To get started, we need to set up the environment with a few prerequisites for permissions and configurations.
---
## Prequisites and Preprocessing
### Permissions and environment variables
Here we set up the linkage and authentication to AWS services.
1. The roles used to give learning and hosting access to your data. See the documentation for how to specify these.
2. The S3 bucket that you want to use for training and model data.
```
%%time
import os
import boto3
import re
import copy
import time
from time import gmtime, strftime
from sagemaker import get_execution_role
role = get_execution_role()
region = boto3.Session().region_name
bucket='<bucket-name>' # put your s3 bucket name here, and create s3 bucket
prefix = 'sagemaker/DEMO-xgboost-multiclass-classification'
# customize to your bucket where you have stored the data
bucket_path = 'https://s3-{}.amazonaws.com/{}'.format(region,bucket)
```
### Data ingestion
Next, we read the dataset from the existing repository into memory, for preprocessing prior to training. This processing could be done *in situ* by Amazon Athena, Apache Spark in Amazon EMR, Amazon Redshift, etc., assuming the dataset is present in the appropriate location. Then, the next step would be to transfer the data to S3 for use in training. For small datasets, such as this one, reading into memory isn't onerous, though it would be for larger datasets.
```
%%time
import pickle, gzip, numpy, urllib.request, json
# Load the dataset
urllib.request.urlretrieve("http://deeplearning.net/data/mnist/mnist.pkl.gz", "mnist.pkl.gz")
f = gzip.open('mnist.pkl.gz', 'rb')
train_set, valid_set, test_set = pickle.load(f, encoding='latin1')
f.close()
```
### Data conversion
Since algorithms have particular input and output requirements, converting the dataset is also part of the process that a data scientist goes through prior to initiating training. In this particular case, the data is converted from pickle-ized numpy array to the libsvm format before being uploaded to S3. The hosted implementation of xgboost consumes the libsvm converted data from S3 for training. The following provides functions for data conversions and file upload to S3 and download from S3.
```
%%time
import struct
import io
import boto3
def to_libsvm(f, labels, values):
f.write(bytes('\n'.join(
['{} {}'.format(label, ' '.join(['{}:{}'.format(i + 1, el) for i, el in enumerate(vec)])) for label, vec in
zip(labels, values)]), 'utf-8'))
return f
def write_to_s3(fobj, bucket, key):
return boto3.Session().resource('s3').Bucket(bucket).Object(key).upload_fileobj(fobj)
def get_dataset():
import pickle
import gzip
with gzip.open('mnist.pkl.gz', 'rb') as f:
u = pickle._Unpickler(f)
u.encoding = 'latin1'
return u.load()
def upload_to_s3(partition_name, partition):
labels = [t.tolist() for t in partition[1]]
vectors = [t.tolist() for t in partition[0]]
num_partition = 5 # partition file into 5 parts
partition_bound = int(len(labels)/num_partition)
for i in range(num_partition):
f = io.BytesIO()
to_libsvm(f, labels[i*partition_bound:(i+1)*partition_bound], vectors[i*partition_bound:(i+1)*partition_bound])
f.seek(0)
key = "{}/{}/examples{}".format(prefix,partition_name,str(i))
url = 's3n://{}/{}'.format(bucket, key)
print('Writing to {}'.format(url))
write_to_s3(f, bucket, key)
print('Done writing to {}'.format(url))
def download_from_s3(partition_name, number, filename):
key = "{}/{}/examples{}".format(prefix,partition_name, number)
url = 's3n://{}/{}'.format(bucket, key)
print('Reading from {}'.format(url))
s3 = boto3.resource('s3')
s3.Bucket(bucket).download_file(key, filename)
try:
s3.Bucket(bucket).download_file(key, 'mnist.local.test')
except botocore.exceptions.ClientError as e:
if e.response['Error']['Code'] == "404":
print('The object does not exist at {}.'.format(url))
else:
raise
def convert_data():
train_set, valid_set, test_set = get_dataset()
partitions = [('train', train_set), ('validation', valid_set), ('test', test_set)]
for partition_name, partition in partitions:
print('{}: {} {}'.format(partition_name, partition[0].shape, partition[1].shape))
upload_to_s3(partition_name, partition)
%%time
convert_data()
```
## Training the XGBoost model
Now that we have our data in S3, we can begin training. We'll use Amazon SageMaker XGboost algorithm, and will actually fit two models in order to demonstrate the single machine and distributed training on SageMaker. In the first job, we'll use a single machine to train. In the second job, we'll use two machines and use the ShardedByS3Key mode for the train channel. Since we have 5 part file, one machine will train on three and the other on two part files. Note that the number of instances should not exceed the number of part files.
First let's setup a list of training parameters which are common across the two jobs.
```
containers = {'us-west-2': '433757028032.dkr.ecr.us-west-2.amazonaws.com/xgboost:latest',
'us-east-1': '811284229777.dkr.ecr.us-east-1.amazonaws.com/xgboost:latest',
'us-east-2': '825641698319.dkr.ecr.us-east-2.amazonaws.com/xgboost:latest',
'eu-west-1': '685385470294.dkr.ecr.eu-west-1.amazonaws.com/xgboost:latest'}
container = containers[boto3.Session().region_name]
#Ensure that the train and validation data folders generated above are reflected in the "InputDataConfig" parameter below.
common_training_params = \
{
"AlgorithmSpecification": {
"TrainingImage": container,
"TrainingInputMode": "File"
},
"RoleArn": role,
"OutputDataConfig": {
"S3OutputPath": bucket_path + "/"+ prefix + "/xgboost"
},
"ResourceConfig": {
"InstanceCount": 1,
"InstanceType": "ml.m4.10xlarge",
"VolumeSizeInGB": 5
},
"HyperParameters": {
"max_depth":"5",
"eta":"0.2",
"gamma":"4",
"min_child_weight":"6",
"silent":"0",
"objective": "multi:softmax",
"num_class": "10",
"num_round": "10"
},
"StoppingCondition": {
"MaxRuntimeInSeconds": 86400
},
"InputDataConfig": [
{
"ChannelName": "train",
"DataSource": {
"S3DataSource": {
"S3DataType": "S3Prefix",
"S3Uri": bucket_path + "/"+ prefix+ '/train/',
"S3DataDistributionType": "FullyReplicated"
}
},
"ContentType": "libsvm",
"CompressionType": "None"
},
{
"ChannelName": "validation",
"DataSource": {
"S3DataSource": {
"S3DataType": "S3Prefix",
"S3Uri": bucket_path + "/"+ prefix+ '/validation/',
"S3DataDistributionType": "FullyReplicated"
}
},
"ContentType": "libsvm",
"CompressionType": "None"
}
]
}
```
Now we'll create two separate jobs, updating the parameters that are unique to each.
### Training on a single instance
```
#single machine job params
single_machine_job_name = 'DEMO-xgboost-classification' + strftime("%Y-%m-%d-%H-%M-%S", gmtime())
print("Job name is:", single_machine_job_name)
single_machine_job_params = copy.deepcopy(common_training_params)
single_machine_job_params['TrainingJobName'] = single_machine_job_name
single_machine_job_params['OutputDataConfig']['S3OutputPath'] = bucket_path + "/"+ prefix + "/xgboost-single"
single_machine_job_params['ResourceConfig']['InstanceCount'] = 1
```
### Training on multiple instances
You can also run the training job distributed over multiple instances. For larger datasets with multiple partitions, this can significantly boost the training speed. Here we'll still use the small/toy MNIST dataset to demo this feature.
```
#distributed job params
distributed_job_name = 'DEMO-xgboost-distrib-classification' + strftime("%Y-%m-%d-%H-%M-%S", gmtime())
print("Job name is:", distributed_job_name)
distributed_job_params = copy.deepcopy(common_training_params)
distributed_job_params['TrainingJobName'] = distributed_job_name
distributed_job_params['OutputDataConfig']['S3OutputPath'] = bucket_path + "/"+ prefix + "/xgboost-distributed"
#number of instances used for training
distributed_job_params['ResourceConfig']['InstanceCount'] = 2 # no more than 5 if there are total 5 partition files generated above
# data distribution type for train channel
distributed_job_params['InputDataConfig'][0]['DataSource']['S3DataSource']['S3DataDistributionType'] = 'ShardedByS3Key'
# data distribution type for validation channel
distributed_job_params['InputDataConfig'][1]['DataSource']['S3DataSource']['S3DataDistributionType'] = 'ShardedByS3Key'
```
Let's submit these jobs, taking note that the first will be submitted to run in the background so that we can immediately run the second in parallel.
```
%%time
region = boto3.Session().region_name
sm = boto3.Session().client('sagemaker')
sm.create_training_job(**single_machine_job_params)
sm.create_training_job(**distributed_job_params)
status = sm.describe_training_job(TrainingJobName=distributed_job_name)['TrainingJobStatus']
print(status)
sm.get_waiter('training_job_completed_or_stopped').wait(TrainingJobName=distributed_job_name)
status = sm.describe_training_job(TrainingJobName=distributed_job_name)['TrainingJobStatus']
print("Training job ended with status: " + status)
if status == 'Failed':
message = sm.describe_training_job(TrainingJobName=distributed_job_name)['FailureReason']
print('Training failed with the following error: {}'.format(message))
raise Exception('Training job failed')
```
Let's confirm both jobs have finished.
```
print('Single Machine:', sm.describe_training_job(TrainingJobName=single_machine_job_name)['TrainingJobStatus'])
print('Distributed:', sm.describe_training_job(TrainingJobName=distributed_job_name)['TrainingJobStatus'])
```
# Set up hosting for the model
In order to set up hosting, we have to import the model from training to hosting. The step below demonstrated hosting the model generated from the distributed training job. Same steps can be followed to host the model obtained from the single machine job.
### Import model into hosting
Next, you register the model with hosting. This allows you the flexibility of importing models trained elsewhere.
```
%%time
import boto3
from time import gmtime, strftime
model_name=distributed_job_name + '-mod'
print(model_name)
info = sm.describe_training_job(TrainingJobName=distributed_job_name)
model_data = info['ModelArtifacts']['S3ModelArtifacts']
print(model_data)
primary_container = {
'Image': container,
'ModelDataUrl': model_data
}
create_model_response = sm.create_model(
ModelName = model_name,
ExecutionRoleArn = role,
PrimaryContainer = primary_container)
print(create_model_response['ModelArn'])
```
### Create endpoint configuration
SageMaker supports configuring REST endpoints in hosting with multiple models, e.g. for A/B testing purposes. In order to support this, customers create an endpoint configuration, that describes the distribution of traffic across the models, whether split, shadowed, or sampled in some way. In addition, the endpoint configuration describes the instance type required for model deployment.
```
from time import gmtime, strftime
endpoint_config_name = 'DEMO-XGBoostEndpointConfig-' + strftime("%Y-%m-%d-%H-%M-%S", gmtime())
print(endpoint_config_name)
create_endpoint_config_response = sm.create_endpoint_config(
EndpointConfigName = endpoint_config_name,
ProductionVariants=[{
'InstanceType':'ml.m4.xlarge',
'InitialVariantWeight':1,
'InitialInstanceCount':1,
'ModelName':model_name,
'VariantName':'AllTraffic'}])
print("Endpoint Config Arn: " + create_endpoint_config_response['EndpointConfigArn'])
```
### Create endpoint
Lastly, the customer creates the endpoint that serves up the model, through specifying the name and configuration defined above. The end result is an endpoint that can be validated and incorporated into production applications. This takes 9-11 minutes to complete.
```
%%time
import time
endpoint_name = 'DEMO-XGBoostEndpoint-' + strftime("%Y-%m-%d-%H-%M-%S", gmtime())
print(endpoint_name)
create_endpoint_response = sm.create_endpoint(
EndpointName=endpoint_name,
EndpointConfigName=endpoint_config_name)
print(create_endpoint_response['EndpointArn'])
resp = sm.describe_endpoint(EndpointName=endpoint_name)
status = resp['EndpointStatus']
print("Status: " + status)
while status=='Creating':
time.sleep(60)
resp = sm.describe_endpoint(EndpointName=endpoint_name)
status = resp['EndpointStatus']
print("Status: " + status)
print("Arn: " + resp['EndpointArn'])
print("Status: " + status)
```
## Validate the model for use
Finally, the customer can now validate the model for use. They can obtain the endpoint from the client library using the result from previous operations, and generate classifications from the trained model using that endpoint.
```
runtime_client = boto3.client('runtime.sagemaker')
```
In order to evaluate the model, we'll use the test dataset previously generated. Let us first download the data from S3 to the local host.
```
download_from_s3('test', 0, 'mnist.local.test') # reading the first part file within test
```
Start with a single prediction. Lets use the first record from the test file.
```
!head -1 mnist.local.test > mnist.single.test
%%time
import json
file_name = 'mnist.single.test' #customize to your test file 'mnist.single.test' if use the data above
with open(file_name, 'r') as f:
payload = f.read()
response = runtime_client.invoke_endpoint(EndpointName=endpoint_name,
ContentType='text/x-libsvm',
Body=payload)
result = response['Body'].read().decode('ascii')
print('Predicted label is {}.'.format(result))
```
OK, a single prediction works.
Let's do a whole batch and see how good is the predictions accuracy.
```
import sys
def do_predict(data, endpoint_name, content_type):
payload = '\n'.join(data)
response = runtime_client.invoke_endpoint(EndpointName=endpoint_name,
ContentType=content_type,
Body=payload)
result = response['Body'].read().decode('ascii')
preds = [float(num) for num in result.split(',')]
return preds
def batch_predict(data, batch_size, endpoint_name, content_type):
items = len(data)
arrs = []
for offset in range(0, items, batch_size):
arrs.extend(do_predict(data[offset:min(offset+batch_size, items)], endpoint_name, content_type))
sys.stdout.write('.')
return(arrs)
```
The following function helps us calculate the error rate on the batch dataset.
```
%%time
import json
file_name = 'mnist.local.test'
with open(file_name, 'r') as f:
payload = f.read().strip()
labels = [float(line.split(' ')[0]) for line in payload.split('\n')]
test_data = payload.split('\n')
preds = batch_predict(test_data, 100, endpoint_name, 'text/x-libsvm')
print ('\nerror rate=%f' % ( sum(1 for i in range(len(preds)) if preds[i]!=labels[i]) /float(len(preds))))
```
Here are a few predictions
```
preds[0:10]
```
and the corresponding labels
```
labels[0:10]
```
The following function helps us create the confusion matrix on the labeled batch test dataset.
```
import numpy
def error_rate(predictions, labels):
"""Return the error rate and confusions."""
correct = numpy.sum(predictions == labels)
total = predictions.shape[0]
error = 100.0 - (100 * float(correct) / float(total))
confusions = numpy.zeros([10, 10], numpy.int32)
bundled = zip(predictions, labels)
for predicted, actual in bundled:
confusions[int(predicted), int(actual)] += 1
return error, confusions
```
The following helps us visualize the erros that the XGBoost classifier is making.
```
import matplotlib.pyplot as plt
%matplotlib inline
NUM_LABELS = 10 # change it according to num_class in your dataset
test_error, confusions = error_rate(numpy.asarray(preds), numpy.asarray(labels))
print('Test error: %.1f%%' % test_error)
plt.xlabel('Actual')
plt.ylabel('Predicted')
plt.grid(False)
plt.xticks(numpy.arange(NUM_LABELS))
plt.yticks(numpy.arange(NUM_LABELS))
plt.imshow(confusions, cmap=plt.cm.jet, interpolation='nearest');
for i, cas in enumerate(confusions):
for j, count in enumerate(cas):
if count > 0:
xoff = .07 * len(str(count))
plt.text(j-xoff, i+.2, int(count), fontsize=9, color='white')
```
### Delete Endpoint
Once you are done using the endpoint, you can use the following to delete it.
```
sm.delete_endpoint(EndpointName=endpoint_name)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/TeachingTextMining/TextClassification/blob/main/06-SA-AutoGOAL/06.2.0-TextClassification-with-AutoGOAL-End2End.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
## Clasificación de textos utilizando AutoML
La clasificación de textos consiste en, dado un texto, asignarle una entre varias categorías. Algunos ejemplos de esta tarea son:
- dado un tweet, categorizar su connotación como positiva, negativa o neutra.
- dado un post de Facebook, clasificarlo como portador de un lenguaje ofensivo o no.
En la actividad exploraremos cómo utilizar la librería [AutoGOAL](https://github.com/autogoal/autogoal) para obtener una solución end-to-end a esta tarea y su aplicación para clasificar reviews de [IMDB](https://www.imdb.com/) sobre películas en las categorías \[$positive$, $negative$\].
**Instrucciones:**
- siga las indicaciones y comentarios en cada apartado.
**Después de esta actividad nos habremos familiarizado con:**
- cómo modelar un problema de clasificación con AutoGOAL
- cómo utilizar AutoGOAL para buscar automáticamente un *pipeline* para clasificación de textos.
- utilizar este *pipeline* para clasificar nuevos textos.
**Requerimientos**
- python 3.6.12 - 3.8
- tensorflow==2.3.0
- autogoal==0.4.4
- pandas==1.1.5
- plotly==4.13.0
- tqdm==4.56.0
<a name="sec:setup"></a>
### Instalación de librerías e importación de dependencias.
Para comenzar, es preciso instalar e incluir las librerías necesarias. En este caso, el entorno de Colab incluye las necesarias.
Ejecute la siguiente casilla prestando atención a las explicaciones dadas en los comentarios.
```
# instalar librerías. Esta casilla es últil por ejemplo si se ejecuta el cuaderno en Google Colab
# Note que existen otras dependencias como tensorflow, etc. que en este caso se encontrarían ya instaladas
%%capture
!pip install autogoal[contrib]==0.4.4
print('Done!')
# temporal cell, just to test AutoGOAL install...
%%capture
#!python -m site
#!ls /usr/local/lib/python3.7/dist-packages
#!pip install rich
#!unzip autogoal.zip
#!mv /usr/local/lib/python3.7/dist-packages/autogoal/ /usr/local/lib/python3.7/dist-packages/autogoal.bak
#!cp -r autogoal /usr/local/lib/python3.7/dist-packages/
print('Done!')
# reset environment
%reset -f
# para construir gráficas y realizar análisis exploratorio de los datos
import plotly.graph_objects as go
import plotly.figure_factory as ff
import plotly.express as px
# para cargar datos y realizar pre-procesamiento básico
import pandas as pd
from collections import Counter
from sklearn.preprocessing import LabelEncoder
# para evaluar los modelos
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score, classification_report, confusion_matrix, roc_curve, auc, f1_score
from sklearn.utils.multiclass import unique_labels
# para configurar AutoGOAL
from autogoal.ml import AutoML
from autogoal.search import (Logger, PESearch, ConsoleLogger, ProgressLogger, MemoryLogger)
from autogoal.kb import Seq, Sentence, VectorCategorical, Supervised
from autogoal.contrib import find_classes
# para guardar el modelo
import pickle
import datetime
print('Done!')
```
#### Definición de funciones y variables necesarias para el pre-procesamiento de datos
Antes de definir el pipeline definiremos algunas variables útiles como el listado de stop words y funciones para cargar los datos, entrenar el modelo etc.
```
# función auxiliar para realizar predicciones con el modelo
def predict_model(model, cfg, data, pref='m'):
"""
data: list of the text to predict
pref: identificador para las columnas (labels_[pref], scores_[pref]_[class 1], etc.)
"""
res = {}
scores = None
labels = model.predict(data)
if hasattr(model, 'predict_proba'):
scores = model.predict_proba(data)
# empaquetar scores dentro de un diccionario que contiene labels, scores clase 1, scores clase 2, .... El nombre de la clase se normaliza a lowercase
res = {f'scores_{pref}_{cls.lower()}':score for cls, score in zip(model.classes_, [col for col in scores.T])}
# añadir datos relativos a la predicción
res[f'labels_{pref}'] = cfg['label_encoder'].inverse_transform(labels)
# convertir a dataframe ordenando las columnas primero el label y luego los scores por clase, las clases ordenadas alfabeticamente.
res = pd.DataFrame(res, columns=sorted(list(res.keys())))
return res
# función auxiliar que evalúa los resultados de una clasificación
def evaluate_model(y_true, y_pred, y_score=None, pos_label='positive'):
"""
data: list of the text to predict
pref: identificador para las columnas (labels_[pref], scores_[pref]_[class 1], etc.)
"""
print('==== Sumario de la clasificación ==== ')
print(classification_report(y_true, y_pred))
print('Accuracy -> {:.2%}\n'.format(accuracy_score(y_true, y_pred)))
# graficar matriz de confusión
display_labels = sorted(unique_labels(y_true, y_pred), reverse=True)
cm = confusion_matrix(y_true, y_pred, labels=display_labels)
z = cm[::-1]
x = display_labels
y = x[::-1].copy()
z_text = [[str(y) for y in x] for x in z]
fig_cm = ff.create_annotated_heatmap(z, x=x, y=y, annotation_text=z_text, colorscale='Viridis')
fig_cm.update_layout(
height=400, width=400,
showlegend=True,
margin={'t':150, 'l':0},
title={'text' : 'Matriz de Confusión', 'x':0.5, 'xanchor': 'center'},
xaxis = {'title_text':'Valor Real', 'tickangle':45, 'side':'top'},
yaxis = {'title_text':'Valor Predicho', 'tickmode':'linear'},
)
fig_cm.show()
# curva roc (definido para clasificación binaria)
fig_roc = None
if y_score is not None:
fpr, tpr, thresholds = roc_curve(y_true, y_score, pos_label=pos_label)
fig_roc = px.area(
x=fpr, y=tpr,
title={'text' : f'Curva ROC (AUC={auc(fpr, tpr):.4f})', 'x':0.5, 'xanchor': 'center'},
labels=dict(x='Ratio Falsos Positivos', y='Ratio Verdaderos Positivos'),
width=400, height=400
)
fig_roc.add_shape(type='line', line=dict(dash='dash'), x0=0, x1=1, y0=0, y1=1)
fig_roc.update_yaxes(scaleanchor="x", scaleratio=1)
fig_roc.update_xaxes(constrain='domain')
fig_roc.show()
# custom logger
# - imprime y guarda el mejor pipeline cada vez que se encuentre una nueva solución candidad
# - imprime pipelines cuya evaluación falló
class CustomLogger(Logger):
def __init__(self, classifier, save_model=True, check_folder="."):
self.save_model = save_model
self.check_folder = check_folder
self.classifier = classifier
def error(self, e: Exception, solution):
if e and solution:
with open("reviews_errors.log", "a") as fp:
fp.write(f"solution={repr(solution)}\nerror={repr(e)}\n\n")
def update_best(self, new_best, new_fn, *args):
pipecode = datetime.datetime.now(datetime.timezone.utc).strftime("reviews--%Y-%m-%d--%H-%M-%S--{0}".format(hex(id(new_best))))
with open("reviews_update_best.log", "a") as fp:
fp.write(f"\n{pipecode}\nsolution={repr(new_best)}\nfitness={new_fn}\n\n")
if(self.save_model):
fp = open('{1}.pkl'.format(self.check_folder,pipecode), 'wb')
new_best.sampler_.replay().save(fp)
pickle.Pickler(fp).dump((self.classifier.input, self.classifier.output))
fp.close()
print('Done!')
```
<a name="sec:load-data"></a>
### Carga de datos y análisis exploratorio
Antes de entrenar el pipeline, es necesario cargar los datos. Existen diferentes opciones, entre estas:
- montar nuestra partición de Google Drive y leer un fichero desde esta.
- leer los datos desde un fichero en una carpeta local.
- leer los datos directamente de un URL.
Ejecute la siguiente casilla prestando atención a las instrucciones adicionales en los comentarios.
```
# descomente las siguientes 3 líneas para leer datos desde Google Drive, asumiendo que se trata de un fichero llamado review.csv localizado dentro de una carpeta llamada 'Datos' en su Google Drive
#from google.colab import drive
#drive.mount('/content/drive')
#path = '/content/drive/MyDrive/Datos/ejemplo_review_train.csv'
# descomente la siguiente línea para leer los datos desde un archivo local, por ejemplo, asumiendo que se encuentra dentro de un directorio llamado sample_data
#path = './sample_data/ejemplo_review_train.csv'
# descomente la siguiente línea para leer datos desde un URL
path = 'https://github.com/TeachingTextMining/TextClassification/raw/main/06-SA-AutoGOAL/sample_data/ejemplo_review_train.csv'
# leer los datos
data = pd.read_csv(path, sep=',')
print('Done!')
```
Una vez leídos los datos, ejecute la siguiente casilla para construir una gráfica que muestra la distribución de clases en el corpus.
```
text_col = 'Phrase' # columna del dataframe que contiene el texto (depende del formato de los datos)
class_col = 'Sentiment' # columna del dataframe que contiene la clase (depende del formato de los datos)
# obtener algunas estadísticas sobre los datos
categories = sorted(data[class_col].unique(), reverse=False)
hist= Counter(data[class_col])
print(f'Total de instancias -> {data.shape[0]}')
print(f'Distribución de clases -> {{item[0]:round(item[1]/len(data[class_col]), 3) for item in sorted(hist.items(), key=lambda x: x[0])}}')
print(f'Categorías -> {categories}')
print(f'Comentario de ejemplo -> {data[text_col][0]}')
print(f'Categoría del comentario -> {data[class_col][0]}')
fig = go.Figure(layout=go.Layout(height=400, width=600))
fig.add_trace(go.Bar(x=categories, y=[hist[cat] for cat in categories]))
fig.show()
print('Done!')
```
Finalmente, ejecute la siguiente casilla para crear los conjuntos de entrenamiento y validación que se utilizarán para entrenar y validar los modelos.
```
# obtener conjuntos de entrenamiento (90%) y validación (10%)
seed = 0 # fijar random_state para reproducibilidad
train, val = train_test_split(data, test_size=.1, stratify=data[class_col], random_state=seed)
print('Done!')
```
### Implementación y configuración del modelo
Con AutoGOAL podemos configurar el modelo facilmente pues solo necesitamos instanciar la clase AutomML. Lo más importante es elegir los tipos adecuados para datos de entrada y salida en nuestro modelo y la métrica de evaluación. En este caso:
- entrada (input), una tupla de:
- Seq(Sentence()) -> una lista (Seq) con cada una de las instancias (Sentence)
- Supervised[VectorCategorical]) -> indica se trata de aprendizaje supervisado.
- salida (output): VectorCategorical -> el elemento *i* representa la categoría asociada a la instancia *i*.
Ejecute la siguiente casilla prestando atención a los comentarios adicionales.
```
# configuraciones
cfg = {}
cfg['iterations'] = 1 # cantidad de iteraciones a realizar
cfg['popsize'] = 50 # tamaño de la población
cfg['search_timeout'] = 120 # tiempo máximo de búsqueda en segundos
cfg['evaluation_timeout'] = 60 # tiempo máximo que empleará evaluando un pipeline en segundos
cfg['memory'] = 20 # cantidad máxima de memoria a utilizar
cfg['score_metric'] = f1_score # métrica de evaluación
search_kwargs=dict(
pop_size=cfg['popsize'],
search_timeout=cfg['search_timeout'],
evaluation_timeout=cfg['evaluation_timeout'],
memory_limit=cfg['memory'] * 1024 ** 3,
)
model = AutoML(
input=(Seq[Sentence], Supervised[VectorCategorical]), # tipo datos de entrada
output=VectorCategorical, # tipo datos de salida
score_metric=cfg['score_metric'],
search_algorithm=PESearch, # algoritmo de búsqueda
registry=None, # para incluir clases adicionales
search_iterations=cfg['iterations'],
include_filter=".*", # indica qué módulos pueden incluirse en los pipelines evaluados
exclude_filter=None, # indica módulos a excluir de los pipelines evaluados
validation_split=0.3, # porción de los datos de entrenamiento que AutoGOAL tomará para evaluar cada pipeline
cross_validation_steps=3, # cantidad de particiones en la crossvalidación
cross_validation="mean", # tipo de agregación para los valores de la métrica en cada partición de la crossvalidación (promedio, mediana, etc.)
random_state=None, # semilla para el generador de números aleatorios
errors="warn", # tratamiento ante errores
**search_kwargs
)
# configurar loggers
loggers = [ProgressLogger(), ConsoleLogger(), MemoryLogger(), CustomLogger(model, save_model=False, check_folder=".")]
print('Done!')
```
<a name="sec:pre-proc"></a>
### Pre-procesamiento de los datos
Notar que en este caso AutoGOAL trabajará directamente con el texto, decidiendo si aplica algún algoritmo de extracción de rasgos o pre-procesamiento. En este caso, solo necesitaremos codificar las categorías como números utilizando [LabelEncoder](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.LabelBinarizer.html) de [scikit-learn](https://scikit-learn.org/stable/).
#### Instanciar LabelEncoder
```
# instanciar LabelEncoder
cfg['label_encoder'] = LabelEncoder()
print('Done!')
```
#### Pre-procesamiento
```
# entrenar LabelEncoder
cfg['label_encoder'].fit(train[class_col])
# guardar LabelEncoder entrenado para su posterior uso (codificar nuevos datos).
with open('label_encoder_reviews.pkl', 'wb') as f:
pickle.dump(cfg['label_encoder'], f)
# codificar labels
train_labels = cfg['label_encoder'].transform(train[class_col])
val_labels = cfg['label_encoder'].transform(val[class_col])
print('Done!')
```
### Entrenamiento del modelo
Por último es necesario "entrenar el modelo", que en este caso significa iniciar la búsqueda.
```
model.fit(train[text_col].to_list(), train_labels, logger=loggers)
print(model.best_pipeline_)
print(model.best_score_)
print('Done!')
```
### Evaluación del modelo
Luego de entrenado el modelo, podemos evaluar su desempeño en los conjuntos de entrenamiento y validación.
Ejecute la siguiente casilla para evaluar el modelo en el conjunto de entrenamiento.
```
# predecir y evaluar el modelo en el conjunto de entrenamiento
print('==== Evaluación conjunto de entrenamiento ====')
data = train
true_labels = data[class_col]
m_pred = predict_model(model, cfg, data[text_col].to_list(), pref='m')
# el nombre de los campos dependerá de pref al llamar a predic_model y las clas|es. Ver comentarios en la definición de la función
evaluate_model(true_labels, m_pred['labels_m'])
print('Done!')
```
Ejecute la siguiente casilla para evaluar el modelo en el conjunto de validación. Compare los resultados.
```
# predecir y evaluar el modelo en el conjunto de validación
print('==== Evaluación conjunto de validacióm ====')
data = val
true_labels = data[class_col]
m_pred = predict_model(model, cfg, data[text_col].to_list(), pref='m')
# el nombre de los campos dependerá de pref al llamar a predic_model y las clases. Ver comentarios en la definición de la función
evaluate_model(true_labels, m_pred['labels_m'])
print('Done!')
```
## Predicción de nuevos datos
Una vez entrenado el modelo, podemos evaluar su rendimiento en datos no utilizados durante el entrenamiento o emplearlo para predecir nuevas instancias. En cualquier caso, se debe cuidar realizar los pasos de pre-procesamiento necesarios según el caso. En el ejemplo, utilizaremos la porción de prueba preparada inicialmente.
**Notar que**:
- se cargará el modelo previamente entrenado y guardado, estableciendo las configuraciones pertinentes.
- si disponemos de un modelo guardado, podremos ejecutar directamente esta parte del cuaderno. Sin embargo, será necesario al menos ejecutar previamente la sección [Instalación de librerías...](#sec:setup)
### Cargar otros elementos necesarios
Antes de predecir nuevos datos, también es preciso cargar otros elementos necesarios como el codificador para las etiquetas, etc.
Ejecute la siguiente casilla.
```
# configuraciones
text_col = 'Phrase' # columna del dataframe que contiene el texto (depende del formato de los datos)
class_col = 'Sentiment' # columna del dataframe que contiene la clase (depende del formato de los datos)
cfg = {} # diccionario para agrupar configuraciones y variables para su posterior uso
# cargar el LabelEncoder
with open('label_encoder_reviews.pkl', 'rb') as f:
cfg['label_encoder'] = pickle.load(f)
```
### Leer datos de entrenamiento y pre-procesarlos
Antes de entrenar el modelo, debemos leer los datos de entrenamiento. Podemos recordar los detalles de [Pre-procesamiento de los datos](#sec:pre-proc).
Ejecute las siguientes casillas.
```
# descomente las siguientes 3 líneas para leer datos desde Google Drive, asumiendo que se trata de un fichero llamado review.csv localizado dentro de una carpeta llamada 'Datos' en su Google Drive
#from google.colab import drive
#drive.mount('/content/drive')
#path = '/content/drive/MyDrive/Datos/ejemplo_review_train.csv'
# descomente la siguiente línea para leer los datos desde un archivo local, por ejemplo, asumiendo que se encuentra dentro de un directorio llamado sample_data
#path = './sample_data/ejemplo_review_train.csv'
# descomente la siguiente línea para leer datos desde un URL
path = 'https://github.com/TeachingTextMining/TextClassification/raw/main/06-SA-AutoGOAL/sample_data/ejemplo_review_train.csv'
# leer los datos
data = pd.read_csv(path, sep=',')
print('Done!')
```
### Entrenar mejor pipeline
En este caso, podemos entrenar con todos los datos.
```
# codificar labels
train_labels = cfg['label_encoder'].transform(data[class_col])
model.fit_pipeline(data[text_col].to_list(), train_labels)
print('Done!')
```
### Predecir nuevos datos
Con el modelo cargado, es posible utilizarlo para analizar nuevos datos.
Ejecute las siguientes casillas para:
(a) categorizar un texto de muestra.
(b) cargar nuevos datos, categorizarlos y mostrar algunas estadísticas sobre el corpus.
```
# ejemplo de texto a clasificar en formato [text 1, text 2, ..., text n]
text = ['Brian De Palma\'s undeniable virtuosity can\'t really camouflage the fact that his plot here is a thinly disguised\
\"Psycho\" carbon copy, but he does provide a genuinely terrifying climax. His "Blow Out", made the next year, was an improvement.']
# predecir los nuevos datos.
m_pred = predict_model(model, cfg, text, pref='m')
# el nombre de los campos dependerá de pref al llamar a predic_model y las clases. Ver comentarios en la definición de la función
pred_labels = m_pred['labels_m'].values[0]
print(f'La categoría del review es -> {pred_labels}')
print('Done!')
```
También podemos predecir nuevos datos cargados desde un fichero.
Ejecute la siguiente casilla, descomentando las instrucciones necesarias según sea el caso.
```
# descomente las siguientes 3 líneas para leer datos desde Google Drive, asumiendo que se trata de un fichero llamado review.csv localizado dentro de una carpeta llamada 'Datos' en su Google Drive
#from google.colab import drive
#drive.mount('/content/drive')
#path = '/content/drive/MyDrive/Datos/ejemplo_review_train.csv'
# descomente la siguiente línea para leer los datos desde un archivo local, por ejemplo, asumiendo que se encuentra dentro de un directorio llamado sample_data
#path = './sample_data/ejemplo_review_train.csv'
# descomente la siguiente línea para leer datos desde un URL
path = 'https://github.com/TeachingTextMining/TextClassification/raw/main/01-SA-Pipeline/sample_data/ejemplo_review_test.csv'
# leer los datos
new_data = pd.read_csv(path, sep=',')
print('Done!')
```
Ejecute la siguiente celda para predecir los datos y mostrar algunas estadísticas sobre el análisis realizado.
```
# predecir los datos de prueba
m_pred = predict_model(model, cfg, new_data[text_col].to_list(), pref='m')
pred_labels = m_pred['labels_m']
# obtener algunas estadísticas sobre la predicción en el conjunto de pruebas
categories = sorted(pred_labels.unique(), reverse=False)
hist = Counter(pred_labels.values)
fig = go.Figure(layout=go.Layout(height=400, width=600))
fig.add_trace(go.Bar(x=categories, y=[hist[cat] for cat in categories]))
fig.show()
print('Done!')
```
| github_jupyter |
Inspired and based on:
- https://shiny.rstudio.com/gallery/movie-explorer.html
- https://demo.bokeh.org/movies
- https://github.com/bokeh/bokeh/tree/master/examples/app/movies
```
flex_subtitle = "built using jupyter-flex"
flex_external_link = "https://github.com/danielfrg/jupyter-flex/blob/master/examples/movie-explorer.ipynb"
flex_title = "Movie explorer"
flex_show_source = True
import sqlite3 as sql
from os.path import dirname, join
import numpy as np
import pandas.io.sql as psql
import plotly.graph_objects as go
from bokeh.sampledata.movies_data import movie_path
import ipywidgets as widgets
from IPython.display import display
```
# Sidebar
```
axis_map = {
"Tomato Meter": "Meter",
"Numeric Rating": "numericRating",
"Number of Reviews": "Reviews",
"Box Office (dollars)": "BoxOffice",
"Length (minutes)": "Runtime",
"Year": "Year",
}
genres = """
All
Action
Adventure
Animation
Biography
Comedy
Crime
Documentary
Drama
Family
Fantasy
History
Horror
Music
Musical
Mystery
Romance
Sci-Fi
Short
Sport
Thriller
War
Western
"""
```
### Filter
```
components = []
reviews_label = widgets.HTML(value="Minimum number of reviews on Rotten Tomatoes:")
reviews = widgets.IntSlider(value=80, min=10, max=300, step=1)
components.extend([reviews_label, reviews])
released_year_label = widgets.HTML(value="Year released:")
released_year = widgets.IntRangeSlider(value=[1970, 2014], min=1940, max=2014, step=1)
components.extend([released_year_label, released_year])
oscars_label = widgets.HTML(value="Minimum number of Oscar wins (all categories):")
oscars = widgets.IntSlider(value=0, min=0, max=4, step=1)
components.extend([oscars_label, oscars])
boxoffice_label = widgets.HTML(value="Dollars at Box Office (millions):")
boxoffice = widgets.IntSlider(value=0, min=0, max=800, step=1)
components.extend([boxoffice_label, boxoffice])
genre_label = widgets.HTML(value="Genre (a movie can have multiple genres):")
genre = widgets.Dropdown(options=genres.split("\n"), value="All")
components.extend([genre_label, genre])
director_label = widgets.HTML(value="Director name contains (e.g., Miyazaki):")
director = widgets.Text()
components.extend([director_label, director])
cast_label = widgets.HTML(value="Cast names contains (e.g. Tom Hanks):")
cast = widgets.Text()
components.extend([cast_label, cast])
all_widgets = widgets.VBox(components)
all_widgets
```
### Variables
```
components = []
x_axis_label = widgets.HTML(value="X-axis variable:")
x_axis = widgets.Dropdown(options=list(axis_map.items()), value="Meter")
components.extend([x_axis_label, x_axis])
y_axis_label = widgets.HTML(value="Y-axis variable:")
y_axis = widgets.Dropdown(options=list(axis_map.items()), value="Reviews")
components.extend([y_axis_label, y_axis])
note = "Note: The Tomato Meter is the proportion of positive reviews (as judged by the Rotten Tomatoes staff), and the Numeric rating is a normalized 1-10 score of those reviews which have star ratings (for example, 3 out of 4 stars)."
note_label = widgets.HTML(value=note)
components.append(note_label)
all_widgets = widgets.VBox(components)
all_widgets
```
# Movies
### Movie explorer
```
conn = sql.connect(movie_path)
query = """
SELECT omdb.ID,
imdbID,
Title,
Year,
omdb.Rating as mpaaRating,
Runtime,
Genre,
Released,
Director,
Writer,
omdb.Cast,
imdbRating,
imdbVotes,
Language,
Country,
Oscars,
tomatoes.Rating as numericRating,
Meter,
Reviews,
Fresh,
Rotten,
userMeter,
userRating,
userReviews,
BoxOffice,
Production
FROM omdb, tomatoes
WHERE omdb.ID = tomatoes.ID AND Reviews >= 10
"""
movies = psql.read_sql(query, conn)
movies["color"] = np.where(movies["Oscars"] > 0, "orange", "grey")
movies["alpha"] = np.where(movies["Oscars"] > 0, 0.9, 0.25)
movies.fillna(0, inplace=True) # just replace missing values with zero
movies["revenue"] = movies.BoxOffice.apply(lambda x: '{:,d}'.format(int(x)))
fig = go.FigureWidget()
plot = go.Scatter(x=[], y=[], mode="markers", text=[], hoverinfo="text", marker=dict(color=[], opacity=[], size=7))
fig.add_trace(plot)
margin = go.layout.Margin(l=20, r=20, b=20, t=30)
fig = fig.update_layout(margin=margin)
def select_movies():
selected = movies[
(movies.Reviews >= reviews.value) &
(movies.BoxOffice >= (boxoffice.value * 1e6)) &
(movies.Year >= released_year.value[0]) &
(movies.Year <= released_year.value[1]) &
(movies.Oscars >= oscars.value)
]
if (genre.value != "All"):
selected = selected[selected.Genre.str.contains(genre.value)==True]
if (director.value != ""):
selected = selected[selected.Director.str.contains(director.value)==True]
if (cast.value != ""):
selected = selected[selected.Cast.str.contains(cast.value)==True]
return selected
def on_value_change(change):
df = select_movies()
x_name = x_axis.value
y_name = y_axis.value
fig.data[0]['x'] = df[x_name]
fig.data[0]['y'] = df[y_name]
fig.data[0]['marker']['color'] = df["color"]
fig.data[0]['marker']['opacity'] = df["alpha"]
fig.data[0]['text'] = df["Title"] + "<br>" + df["Year"].astype(str) + "<br>" + df["BoxOffice"].astype(str)
fig.update_xaxes(title_text=x_axis.label)
fig.update_yaxes(title_text=y_axis.label)
fig.update_layout(title="%d movies selected" % len(df))
controls = [reviews, boxoffice, released_year, oscars, genre, director, cast, x_axis, y_axis]
for control in controls:
control.observe(on_value_change, names="value")
on_value_change(None)
fig
```
| github_jupyter |
# Creating Simulated PyVISA Instruments
When developing stuff in a large codebase like QCoDeS, it is often uncanningly easy to submit a change that breaks stuff. Therefore, _continuous integration_ is performed in the form of automated tests that run before new code is allowed into the codebase. The many tests of QCoDeS can be found in `qcodes.tests`.
But how about drivers? They constitute the majority of the codebase, but how can we test them? Wouldn't that require a physical copy each instrument to be present on the California server where we run our tests? It used to be so, but not anymore! For drivers utilising PyVISA (i.e. `VisaInstrument` drivers), we may create simulated instruments to which the drivers may connect.
## What?
This way, we may instantiate drivers and run simple tests on them. Tests like:
* Can the driver even instantiate? This is very relevant when underlying APIs change.
* Is the drivers (e.g.) "voltage-to-bytecode" converter working properly?
## Not!
It is not feasible to simulate any but the most trivial features of the instrument. Simulated instruments can not and should not perform tests like:
* Do we wait sufficiently long for this oscilloscope's trace to be acquired?
* Does our driver handle overlapping commands of this AWG correctly?
## How?
The basic scheme goes as follows:
* Write a `.yaml` file for the simulated instrument. The instructions for that may be found here: https://pyvisa-sim.readthedocs.io/en/latest/ and specifically here: https://pyvisa-sim.readthedocs.io/en/latest/definitions.html#definitions
* Then write a test for your instrument and put it in `qcodes/tests/drivers`. The file should have the name `test_<nameofyourdriver>.py`.
* Check that all is well by running `$ pytest test_<nameofyourdriver>.py`.
Below is an example.
## Example: Weinschel_8320
The Weinschel 8320 is a very simple driver.
```
from qcodes.instrument.visa import VisaInstrument
import qcodes.utils.validators as vals
import numpy as np
class Weinschel_8320(VisaInstrument):
"""
QCoDeS driver for the stepped attenuator
Weinschel is formerly known as Aeroflex/Weinschel
"""
def __init__(self, name, address, **kwargs):
super().__init__(name, address, terminator='\r', **kwargs)
self.add_parameter('attenuation', unit='dB',
set_cmd='ATTN ALL {:02.0f}',
get_cmd='ATTN? 1',
vals=vals.Enum(*np.arange(0, 60.1, 2).tolist()),
get_parser=float)
self.connect_message()
```
### The `.yaml` file
The simplest `.yaml` file that is still useful, reads, in all its glory:
```
spec: "1.0"
devices:
device 1:
eom:
GPIB INSTR:
q: "\r" # MAKE SURE! that this matches the terminator of the driver!
r: "\r"
error: ERROR
dialogues:
- q: "*IDN?"
r: "QCoDeS, Weinschel 8320 (Simulated), 1337, 0.0.01"
resources:
GPIB::1::INSTR:
device: device 1
```
Note that since no physical connection is made, it doesn't matter what interface we pretend to use (GPIB, USB, ethernet, serial, ...). As a convention, we always write GPIB in the `.yaml` files.
We save the above file as `qcodes/instrument/sims/Weinschel_8320.yaml`. This simulates an instrument with no settable parameter; only an `*IDN?` response. This is enough to instantiate the instrument.
Then we may connect to the simulated instrument.
```
import qcodes.instrument.sims as sims
# path to the .yaml file containing the simulated instrument
visalib = sims.__file__.replace('__init__.py', 'Weinschel_8320.yaml@sim')
wein_sim = Weinschel_8320('wein_sim',
address='GPIB::1::INSTR', # This matches the address in the .yaml file
visalib=visalib
)
```
### The test
Now we can write a useful test!
```
import pytest
from qcodes.instrument_drivers.weinschel.Weinschel_8320 import Weinschel_8320
import qcodes.instrument.sims as sims
visalib = sims.__file__.replace('__init__.py', 'Weinschel_8320.yaml@sim')
# The following decorator makes the driver
# available to all the functions in this module
@pytest.fixture(scope='function')
def driver():
wein_sim = Weinschel_8320('wein_sim',
address='GPIB::1::65535::INSTR',
visalib=visalib
)
yield wein_sim
wein_sim.close()
def test_init(driver):
"""
Test that simple initialisation works
"""
# There is not that much to do, really.
# We can check that the IDN string reads back correctly
idn_dict = driver.IDN()
assert idn_dict['vendor'] == 'QCoDeS'
```
Save the test as `qcodes/tests/drivers/test_weinschel_8320.py`.
Open a command line/console/terminal, navigate to the `qcodes/tests/drivers/` folder and run
```
>> pytest test_weinschel_8320.py
```
This should give you an output similar to
```
========================================= 1 passed in 0.73 seconds ==========================================
```
## Congratulations! That was it.
## Bonus example: including parameters in the simulated instrument
It is also possible to add queriable parameters to the `.yaml` file, but testing that you can read those back is of limited value. You should only add them if your driver needs them to instantiate, e.g. if it checks that some range or impedance is configured correctly on startup, or - more generally - if a part of your driver code that you'd like to test needs it to run.
For the sake of this example, let us add a test that the driver's parameter's validator will reject an attenuation of less than 0 dBm. Note that this concrete test is redundant, since we have separate tests for validators. It is, however, an excellent example to learn from.
First we update the `.yaml` file to contain a property matching the parameter.
```
spec: "1.0"
devices:
device 1:
eom:
GPIB INSTR:
q: "\r" # MAKE SURE! that this matches the terminator of the driver!
r: "\r"
error: ERROR
dialogues:
- q: "*IDN?"
r: "QCoDeS, Weinschel 8320 (Simulated), 1337, 0.0.01"
properties:
attenuation:
default: 0
getter:
q: "ATTN? 1" # the set/get commands have to simply be copied over from the driver
r: "{:02.0f}"
setter:
q: "ATTN ALL {:02.0f}"
resources:
GPIB::1::INSTR:
device: device 1
```
Notice that we don't include the the
```r: OK```
as the response of setting a property. This is in contrast to what https://pyvisa-sim.readthedocs.io/en/latest/definitions.html#properties does. The response of a successful setting of a parameter will not return 'OK'.
Next we update the test script.
```
import pytest
from qcodes.instrument_drivers.weinschel.Weinschel_8320 import Weinschel_8320
import qcodes.instrument.sims as sims
visalib = sims.__file__.replace('__init__.py', 'Weinschel_8320.yaml@sim')
# The following decorator makes the driver
# available to all the functions in this module
@pytest.fixture(scope='function')
def driver():
wein_sim = Weinschel_8320('wein_sim',
address='GPIB::1::INSTR',
visalib=visalib
)
yield wein_sim
wein_sim.close()
def test_init(driver):
"""
Test that simple initialisation works
"""
# There is not that much to do, really.
# We can check that the IDN string reads back correctly
idn_dict = driver.IDN()
assert idn_dict['vendor'] == 'QCoDeS'
def test_attenuation_validation(driver):
"""
Test that incorrect values are rejected
"""
bad_values = [-1, 1, 1.5]
for bv in bad_values:
with pytest.raises(ValueError):
driver.attenuation(bv)
```
Open a command line/console/terminal, navigate to the `qcodes/tests/drivers/` folder and run
```
>> pytest test_weinschel_8320.py
```
This should give you an output similar to
```
========================================= 2 passed in 0.73 seconds ==========================================
```
## That's it!
| github_jupyter |
# Segment 2
## Regression but Not Backwards
```
# loading libraries
import numpy as np
import pandas as pd
from sklearn.metrics import mean_squared_error
from sklearn.model_selection import train_test_split, cross_val_score
from sklearn.preprocessing import LabelBinarizer
# define column names
names = ['sepal_length', 'sepal_width', 'petal_length', 'petal_width', 'species']
# loading training data
iris = pd.read_csv('iris.data', header=None, names=names)
enc = LabelBinarizer().fit_transform(iris['species']) # one hot encode categorical species feature
X_reg = np.concatenate([iris[['sepal_length', "sepal_width", "petal_width"]],enc[:,:2]], axis=1) # end index is exclusive
y_reg = np.array(iris['petal_length']) # column name is another way of indexing df
# split into train and test
X_train_reg, X_test_reg, y_train_reg, y_test_reg = train_test_split(X_reg, y_reg, test_size=0.33, random_state=42)
X_train_reg[:5,:5]
```
### Linear Regression
```
from sklearn import linear_model
import statsmodels.api as sm
from IPython.core.display import HTML
def short_summary(est):
return HTML(est.summary().tables[1].as_html())
```
#### Using Sklearn
Train Model
```
# Create linear regression object
regr = linear_model.LinearRegression()
# Train the model using the training sets
regr.fit(X_train_reg, y_train_reg)
# The coefficients
print('Sklearn Coefficients: \n', regr.coef_)
print('Sklearn Intercept: \n', regr.intercept_)
```
Test Model
```
# Make predictions using the testing set
y_pred = regr.predict(X_test_reg)
print("Sklearn MSE: ", mean_squared_error(y_test_reg, y_pred))
```
#### Using Statsmodels
Train Model
```
X_train_regSM = sm.add_constant(X_train_reg)
# Fit and summarize OLS model
mod = sm.OLS(y_train_reg, X_train_regSM)
res = mod.fit()
print("Statmodels Coefficients:")
short_summary(res)
```
Test Model
```
X_test_regSM = sm.add_constant(X_test_reg)
y_pred = res.predict(X_test_regSM)
print("Statsmodels MSE: ", mean_squared_error(y_test_reg, y_pred))
#add plots? normality etc, fitted line 3D
```
### Multivariate Regression
```
X_MVreg = np.concatenate([iris[['sepal_length']], enc[:,:2]], axis=1) # , "sepal_width" missing
y_MVreg = np.array(iris[['petal_length', 'petal_width']]) # 2 response vars
# split into train and test
X_train_MVreg, X_test_MVreg, y_train_MVreg, y_test_MVreg = train_test_split(X_MVreg, y_MVreg, test_size=0.33, random_state=42)
print("Regression X_train:")
print(X_train_MVreg[1:5,])
print("\nRegression y_train:")
print(y_train_MVreg[1:5])
```
#### Sklearn
Train Model
```
# Create linear regression object
regr_MV = linear_model.LinearRegression()
# Train the model using the training sets
regr_MV.fit(X_train_MVreg, y_train_MVreg)
# The coefficients
print('Sklearn Coefficients: \n', regr_MV.coef_, "\n")
print('Sklearn Intercepts: \n', regr_MV.intercept_)
```
Same as fitting linear model 2 seperate times for each y variable.
```
# petal_length
regr_MV1 = linear_model.LinearRegression()
# Train the model using the training sets
regr_MV1.fit(X_train_MVreg, y_train_MVreg[:, 0])
print('Sklearn Coefficients: \n', regr_MV1.coef_)
print('Sklearn Intercept: \n', regr_MV1.intercept_)
# petal_width
regr_MV2 = linear_model.LinearRegression()
# Train the model using the training sets
regr_MV2.fit(X_train_MVreg, y_train_MVreg[:, 1])
print('Sklearn Coefficients: \n', regr_MV2.coef_)
print('Sklearn Intercept: \n', regr_MV2.intercept_)
```
Test Model
```
# Make predictions using the testing set
y_pred_MV = regr_MV.predict(X_test_MVreg)
print("Sklearn MSE: ", mean_squared_error(y_test_MVreg, y_pred_MV))
```
#### Statsmodels
Using Statsmodels to fit 2 seperate models. Does not allow for multivariate regression.
```
X_train_MVreg_SM = sm.add_constant(X_train_MVreg)
# Fit and summarize OLS model
mod_MV1 = sm.OLS(y_train_MVreg[:,0], X_train_MVreg_SM).fit()
print("Statmodels Coefficients:")
short_summary(mod_MV1)
mod_MV2 = sm.OLS(y_train_MVreg[:,1], X_train_MVreg_SM).fit()
print("Statmodels Coefficients:")
short_summary(mod_MV2)
```
#### R for proper Multivariate Regression
Using R within python to provide proper context on statistical tests in multivariate regression since sklearn does not provide hypothesis test results for multivariate regression and statsmodel does not have multivariate regression functionality at all.
The _pyper_ library enables this and can be installed using `pip install pyper`. to install
```
import pyper as pr
# CREATE A R INSTANCE WITH PYPER
r=pr.R(RCMD="C:/Program Files/R/R-4.0.0/bin/R", use_pandas = True) #R version >3.5
#convert arrays to dataframes
X_train_MVreg_df=pd.DataFrame(X_train_MVreg, columns=['sepal_length', 'Iris-virginica', 'Iris-versicolor'])
X_test_MVreg_df=pd.DataFrame(X_test_MVreg, columns=['sepal_length', 'Iris-virginica', 'Iris-versicolor'])
y_train_MVreg_df=pd.DataFrame(y_train_MVreg, columns=['petal_length', 'petal_width'])
y_test_MVreg_df=pd.DataFrame(y_test_MVreg, columns=['petal_length', 'petal_width'])
X_train_MVreg_df.head()
y_train_MVreg_df.head()
# PASS DATA FROM PYTHON TO R
r.assign("X_Train", X_train_MVreg_df)
r.assign("Y_Train", y_train_MVreg_df)
r.assign("X_Test", X_test_MVreg_df)
r.assign("Y_Test", y_test_MVreg_df)
r("Y_Train <- as.matrix(Y_Train)")
r("mvmod <- lm(Y_Train ~ sepal_length + Iris.virginica + Iris.versicolor, data=X_Train)")
print(r("summary(mvmod)"))
print(r("library(car)"))
print(r("Anova(mvmod)"))
r("ypred=predict(mvmod, X_Test) ")
print(r('ypred'))
```
| github_jupyter |
# Centroid digits of MNIST
```
import numpy as np
import matplotlib.pyplot as plt
from keras.datasets import mnist # Importing mnist dataset from keras(Tensorflow)
# loading mnist dataset and split it into train_x, train_y, test_x, test_y
(train_X, train_y), (test_X, test_y) = mnist.load_data()
# Printing shape of dataset
print('X_train: ' + str(train_X.shape))
print('Y_train: ' + str(train_y.shape))
print('X_test: ' + str(test_X.shape))
print('Y_test: ' + str(test_y.shape))
import matplotlib.pyplot as plt
# Plotting the first 100 samples at train_X series
for i in range(100):
plt.imshow(train_X[i])
plt.show()
# Declaring 2 lists : The first for train and the second for test only for features X
train_features = []
test_features = []
# Here we'll split each image into 4-grids(2x2) grid to get centriod of each image.So we'll create the function called Get_centriod
# which hold 3 paramater (curr_image,w,h) current image, it's width and finally it's height.
# Cause we've determined to split into 4-grids(2x2) so we'll iterate 4 times
# #1#2
# #3#4
def Get_centroid(curr_image,w,h):
feature_vector = []
x1 = y1 = x2 = y2 = 0
for item in range(4):
if item == 0: # If we're at first grid(#1), we'll set x1,y1 to 0 & x2,y2 to 14
x1 = y1 = 0
x2 = y2 = 14
elif item == 1: # If we're at first grid(#2), we'll set x1 to 14, y1 to 0 & x2 to 28 & y2 to 14
x1 = 14
y1 = 0
x2 = 2*14 # 28
y2 = 14
elif item == 2: # If we're at first grid(#3), we'll update x1 to 0, y1 to 14 & x2 to 14, y2 to 28
x1 = 0
y1 = 14
x2 = 14
y2 = 2*14
else : # If we're at first grid(#4), we'll update x1,y1 to 14 & x2,y2 to 28
x1 = y1 = 14
x2 = y2 = 2*14
sum_x = 0
sum_y = 0
total_pixels = 0
for i in range(x1,x2): # Iterate through each determined grid
for j in range(y1,y2):
sum_x = sum_x + (i-x1)*curr_image[i][j] # iterate through each row to get sum the pixel of current image
sum_y = sum_y + (j-y1)*curr_image[i][j] # iterate through each column to get sum the pixel of current image
total_pixels = total_pixels+curr_image[i][j] # To get total sum of pixels of each grid.
if total_pixels != 0: # To avoid divided by zero
feature_vector.append((sum_x/total_pixels)) # get centroid of x at each grid
feature_vector.append((sum_y/total_pixels)) # get centroid of y at each grid
else:
# if total_pixels is equal to zero then add zero
feature_vector.append(0)
feature_vector.append(0)
return feature_vector
for i in range(10000): # We'll iterate through 10000; and Getting centriod of each element of train_x then add it into train_features list to fit it into model later.
train_features.append(Get_centroid(train_X[i],28,28))
for i in range(1000): # We'll iterate through 1000; and Getting centriod of each element of test_x then add it into train_features list to test it into model later.
test_features.append(Get_centroid(test_X[i],28,28))
from sklearn.neighbors import KNeighborsClassifier # Importing KNN model
knn_model = KNeighborsClassifier(n_neighbors=8)
knn_model.fit(train_features,train_y[:10000]) # Fit the model with train_features & the first 10000 samples train_y
predicted_y = knn_model.predict(test_features)
from sklearn.metrics import accuracy_score # to get accuracy
print("Accuracy of KNN model: "+str(accuracy_score(test_y[:1000], predicted_y))) # Predict the model with the first 1000 samples of test_y & predicted_y1
```
| github_jupyter |
<a href="https://colab.research.google.com/github/vasudevgupta7/gsoc-wav2vec2/blob/main/notebooks/wav2vec2_onnx.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Wav2Vec2 ONNX
In this notebook, we will be exporting TF Wav2Vec2 model into ONNX and will compare ONNX exported and TF model latency on CPU.
```
!pip3 install -qU tf2onnx onnxruntime
!pip3 install -q git+https://github.com/vasudevgupta7/gsoc-wav2vec2@main
```
## Exporting TF model to ONNX
Exporting to ONNX is quite straight forward. One can use `tf2onnx.convert.from_keras(...)` method.
```
import tensorflow as tf
from wav2vec2 import Wav2Vec2ForCTC
model_id = "vasudevgupta/gsoc-wav2vec2-960h"
model = Wav2Vec2ForCTC.from_pretrained(model_id)
AUDIO_MAXLEN = 50000
ONNX_PATH = "onnx-wav2vec2.onnx"
import tf2onnx
input_signature = (tf.TensorSpec((None, AUDIO_MAXLEN), tf.float32, name="speech"),)
_ = tf2onnx.convert.from_keras(model, input_signature=input_signature, output_path=ONNX_PATH)
ls
```
## Inference using ONNX exported model
For running inference with the onnx-exported model, we will first download some speech sample and then apply some pre-processing.
```
!wget https://github.com/vasudevgupta7/gsoc-wav2vec2/raw/main/data/sample.wav
from wav2vec2 import Wav2Vec2Processor
processor = Wav2Vec2Processor(is_tokenizer=False)
```
Instance of `Wav2Vec2Processor(is_tokenizer=False)` is going to normalize the speech along the time axis. This preprocessing was applied during training also.
```
import soundfile as sf
FILENAME = "sample.wav"
speech, _ = sf.read(FILENAME)
speech = tf.constant(speech, dtype=tf.float32)
speech = processor(speech)[None]
padding = tf.zeros((speech.shape[0], AUDIO_MAXLEN - speech.shape[1]))
speech = tf.concat([speech, padding], axis=-1)
speech.shape
```
Now we will initiate ONNX runtime session and use that session to make predictions.
```
import onnxruntime as rt
session = rt.InferenceSession(ONNX_PATH)
@tf.function(jit_compile=True)
def jitted_forward(speech):
return model(speech)
import numpy as np
onnx_outputs = session.run(None, {"speech": speech.numpy()})[0]
tf_outputs = jitted_forward(speech)
assert np.allclose(onnx_outputs, tf_outputs.numpy(), atol=1e-2)
tokenizer = Wav2Vec2Processor(is_tokenizer=True)
prediction = np.argmax(onnx_outputs, axis=-1)
prediction = tokenizer.decode(prediction.squeeze().tolist())
```
Instance of `Wav2Vec2Processor(is_tokenizer=True)` is used for decoding model outputs to string.
```
from IPython.display import Audio
print("prediction:", prediction)
Audio(filename=FILENAME)
```
## Comparing latency of TF model & ONNX exported model
Now we will be comparing latency for jitted model & ONNX exported model.
```
import time
from contextlib import contextmanager
@contextmanager
def timeit(prefix="Time taken:"):
start = time.time()
yield
time_taken = time.time() - start
print(prefix, time_taken, "seconds")
with timeit(prefix="JIT Compiled Wav2vec2 time taken:"):
jitted_forward(speech)
with timeit(prefix="Eager mode time taken:"):
model(speech)
with timeit(prefix="ONNX-Wav2Vec2 time taken:"):
session.run(None, {"speech": speech.numpy()})
```
| github_jupyter |
```
import tensorflow as tf
import numpy as np
import copy
import time
from sklearn.cross_validation import train_test_split
import os
import pickle
def layer_norm_all(h, base, num_units, scope):
with tf.variable_scope(scope):
h_reshape = tf.reshape(h, [-1, base, num_units])
mean = tf.reduce_mean(h_reshape, [2], keep_dims=True)
var = tf.reduce_mean(tf.square(h_reshape - mean), [2], keep_dims=True)
epsilon = tf.constant(1e-3)
rstd = tf.rsqrt(var + epsilon)
h_reshape = (h_reshape - mean) * rstd
h = tf.reshape(h_reshape, [-1, base * num_units])
alpha = tf.get_variable('layer_norm_alpha', [4 * num_units],
initializer=tf.constant_initializer(1.0), dtype=tf.float32)
bias = tf.get_variable('layer_norm_bias', [4 * num_units],
initializer=tf.constant_initializer(0.0), dtype=tf.float32)
return (h * alpha) + bias
def layer_norm(x, scope="layer_norm", alpha_start=1.0, bias_start=0.0):
with tf.variable_scope(scope):
num_units = x.get_shape().as_list()[1]
alpha = tf.get_variable('alpha', [num_units],
initializer=tf.constant_initializer(alpha_start), dtype=tf.float32)
bias = tf.get_variable('bias', [num_units],
initializer=tf.constant_initializer(bias_start), dtype=tf.float32)
mean, variance = moments_for_layer_norm(x)
y = (alpha * (x - mean)) / (variance) + bias
return y
def moments_for_layer_norm(x, axes=1, name=None):
epsilon = 1e-3
if not isinstance(axes, list): axes = [axes]
mean = tf.reduce_mean(x, axes, keep_dims=True)
variance = tf.sqrt(tf.reduce_mean(tf.square(x - mean), axes, keep_dims=True) + epsilon)
return mean, variance
def zoneout(new_h, new_c, h, c, h_keep, c_keep, is_training):
mask_c = tf.ones_like(c)
mask_h = tf.ones_like(h)
if is_training:
mask_c = tf.nn.dropout(mask_c, c_keep)
mask_h = tf.nn.dropout(mask_h, h_keep)
mask_c *= c_keep
mask_h *= h_keep
h = new_h * mask_h + (-mask_h + 1.) * h
c = new_c * mask_c + (-mask_c + 1.) * c
return h, c
class LN_LSTMCell(tf.contrib.rnn.RNNCell):
def __init__(self, num_units, f_bias=1.0, use_zoneout=False,
zoneout_keep_h = 0.9, zoneout_keep_c = 0.5,
is_training = True,reuse=None, name=None):
super(LN_LSTMCell, self).__init__(_reuse=reuse, name=name)
self.num_units = num_units
self.f_bias = f_bias
self.use_zoneout = use_zoneout
self.zoneout_keep_h = zoneout_keep_h
self.zoneout_keep_c = zoneout_keep_c
self.is_training = is_training
def build(self, inputs_shape):
w_init = tf.orthogonal_initializer(1.0)
h_init = tf.orthogonal_initializer(1.0)
b_init = tf.constant_initializer(0.0)
h_size = self.num_units
self.W_xh = tf.get_variable('W_xh',[inputs_shape[1], 4 * h_size], initializer=w_init, dtype=tf.float32)
self.W_hh = tf.get_variable('W_hh',[h_size, 4 * h_size], initializer=h_init, dtype=tf.float32)
self.bias = tf.get_variable('bias', [4 * h_size], initializer=b_init, dtype=tf.float32)
def call(self, x, state):
h, c = state
h_size = self.num_units
concat = tf.concat(axis=1, values=[x, h])
W_full = tf.concat(axis=0, values=[self.W_xh, self.W_hh])
concat = tf.matmul(concat, W_full) + self.bias
concat = layer_norm_all(concat, 4, h_size, 'ln')
i, j, f, o = tf.split(axis=1, num_or_size_splits=4, value=concat)
new_c = c * tf.sigmoid(f + self.f_bias) + tf.sigmoid(i) * tf.tanh(j)
new_h = tf.tanh(layer_norm(new_c, 'ln_c')) * tf.sigmoid(o)
if self.use_zoneout:
new_h, new_c = zoneout(new_h, new_c, h, c, self.zoneout_keep_h,
self.zoneout_keep_c, self.is_training)
return new_h, new_c
def zero_state(self, batch_size, dtype):
h = tf.zeros([batch_size, self.num_units], dtype=dtype)
c = tf.zeros([batch_size, self.num_units], dtype=dtype)
return (h, c)
@property
def state_size(self):
return self.num_units
@property
def output_size(self):
return self.num_units
class Model:
def __init__(self, num_layers, size_layer, dimension_input, dimension_output, learning_rate):
def lstm_cell():
return tf.contrib.rnn.LayerNormBasicLSTMCell(size_layer)
self.rnn_cells = tf.nn.rnn_cell.MultiRNNCell([lstm_cell() for _ in range(num_layers)])
self.X = tf.placeholder(tf.float32, [None, None, dimension_input])
self.Y = tf.placeholder(tf.float32, [None, dimension_output])
drop = tf.contrib.rnn.DropoutWrapper(self.rnn_cells, output_keep_prob = 0.5)
self.outputs, self.last_state = tf.nn.dynamic_rnn(drop, self.X, dtype = tf.float32)
self.rnn_W = tf.Variable(tf.random_normal((size_layer, dimension_output)))
self.rnn_B = tf.Variable(tf.random_normal([dimension_output]))
self.logits = tf.matmul(self.outputs[:, -1], self.rnn_W) + self.rnn_B
self.cost = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(logits = self.logits, labels = self.Y))
l2 = sum(0.0005 * tf.nn.l2_loss(tf_var) for tf_var in tf.trainable_variables())
self.cost += l2
self.optimizer = tf.train.AdamOptimizer(learning_rate = learning_rate).minimize(self.cost)
self.correct_pred = tf.equal(tf.argmax(self.logits, 1), tf.argmax(self.Y, 1))
self.accuracy = tf.reduce_mean(tf.cast(self.correct_pred, tf.float32))
maxlen = 20
location = os.getcwd()
num_layers = 2
size_layer = 256
learning_rate = 1e-7
batch = 100
with open('dataset-emotion.p', 'rb') as fopen:
df = pickle.load(fopen)
with open('vector-emotion.p', 'rb') as fopen:
vectors = pickle.load(fopen)
with open('dataset-dictionary.p', 'rb') as fopen:
dictionary = pickle.load(fopen)
label = np.unique(df[:,1])
train_X, test_X, train_Y, test_Y = train_test_split(df[:,0], df[:, 1].astype('int'), test_size = 0.2)
tf.reset_default_graph()
sess = tf.InteractiveSession()
model = Model(num_layers, size_layer, vectors.shape[1], label.shape[0], learning_rate)
sess.run(tf.global_variables_initializer())
dimension = vectors.shape[1]
saver = tf.train.Saver(tf.global_variables())
EARLY_STOPPING, CURRENT_CHECKPOINT, CURRENT_ACC, EPOCH = 10, 0, 0, 0
while True:
lasttime = time.time()
if CURRENT_CHECKPOINT == EARLY_STOPPING:
print('break epoch:', EPOCH)
break
train_acc, train_loss, test_acc, test_loss = 0, 0, 0, 0
for i in range(0, (train_X.shape[0] // batch) * batch, batch):
batch_x = np.zeros((batch, maxlen, dimension))
batch_y = np.zeros((batch, len(label)))
for k in range(batch):
tokens = train_X[i + k].split()[:maxlen]
emb_data = np.zeros((maxlen, dimension), dtype = np.float32)
for no, text in enumerate(tokens[::-1]):
try:
emb_data[-1 - no, :] += vectors[dictionary[text], :]
except Exception as e:
print(e)
continue
batch_y[k, int(train_Y[i + k])] = 1.0
batch_x[k, :, :] = emb_data[:, :]
loss, _ = sess.run([model.cost, model.optimizer], feed_dict = {model.X : batch_x, model.Y : batch_y})
train_loss += loss
train_acc += sess.run(model.accuracy, feed_dict = {model.X : batch_x, model.Y : batch_y})
for i in range(0, (test_X.shape[0] // batch) * batch, batch):
batch_x = np.zeros((batch, maxlen, dimension))
batch_y = np.zeros((batch, len(label)))
for k in range(batch):
tokens = test_X[i + k].split()[:maxlen]
emb_data = np.zeros((maxlen, dimension), dtype = np.float32)
for no, text in enumerate(tokens[::-1]):
try:
emb_data[-1 - no, :] += vectors[dictionary[text], :]
except:
continue
batch_y[k, int(test_Y[i + k])] = 1.0
batch_x[k, :, :] = emb_data[:, :]
loss, acc = sess.run([model.cost, model.accuracy], feed_dict = {model.X : batch_x, model.Y : batch_y})
test_loss += loss
test_acc += acc
train_loss /= (train_X.shape[0] // batch)
train_acc /= (train_X.shape[0] // batch)
test_loss /= (test_X.shape[0] // batch)
test_acc /= (test_X.shape[0] // batch)
if test_acc > CURRENT_ACC:
print('epoch:', EPOCH, ', pass acc:', CURRENT_ACC, ', current acc:', test_acc)
CURRENT_ACC = test_acc
CURRENT_CHECKPOINT = 0
saver.save(sess, os.getcwd() + "/model-rnn-vector.ckpt")
else:
CURRENT_CHECKPOINT += 1
EPOCH += 1
print('time taken:', time.time()-lasttime)
print('epoch:', EPOCH, ', training loss:', train_loss, ', training acc:', train_acc, ', valid loss:', test_loss, ', valid acc:', test_acc)
```
| github_jupyter |
```
import serial, time, glob
ports = glob.glob('/dev/tty[A-Za-z]*') # Modify if on a platform other than Linux
ports
ser = serial.Serial(ports[0], 115200)
def read_all():
r = ser.read_all() # clear buffer
ser.write(b'a')
while ser.in_waiting < 1:
pass # wait for a response
time.sleep(0.05)
r = ser.read_all()
t = str(r).split('A')[-1].strip()
r = [[int(s.strip('b\'')) for s in str(l).split(',')[:8]] for l in str(r).split('A')[:-1]]
return(r, t)
import numpy as np
r, t = read_all()
np.asarray(r)
import glob, serial, time, math
import numpy as np
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.neural_network import MLPClassifier, MLPRegressor
from sklearn.preprocessing import StandardScaler
# Load data and train a model
df1 = pd.read_csv('demoq.csv')
# Train a model
X_train, X_test, y_train, y_test = train_test_split(df1[[str(i) for i in range(8,72)]], df1['q'])
scaler = StandardScaler()
scaler.fit(X_train)
X_train = scaler.transform(X_train)
X_test = scaler.transform(X_test)
mlpc = MLPClassifier(hidden_layer_sizes=(20, 20, 20), max_iter=1000)
mlpc.fit(X_train, y_train)
print('Score:',mlpc.score(X_test, y_test))
in1 = read_all()[0] #df1[[str(i) for i in range(72)]].iloc[200]
X = scaler.transform([np.asarray(in1).flatten()])
print(mlpc.predict(X))
len(np.asarray(in1).flatten())
X = []
y = []
q = []
readings = read_all()
X.append(np.asarray(readings[0]).flatten())
q.append(4)
y.append(1)# 0 empty, 1 finger, 2 pen
print(len(X))
from matplotlib import pyplot as plt
%matplotlib inline
plt.plot(X[0])
import glob, serial, time, math
import numpy as np
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.neural_network import MLPClassifier, MLPRegressor
from sklearn.preprocessing import StandardScaler
# Train a model
# X2 = [np.asarray(x[0]).flatten() for x in X]
X_train, X_test, y_train, y_test = train_test_split(X, y)
scaler = StandardScaler()
scaler.fit(X_train)
X_train = scaler.transform(X_train)
X_test = scaler.transform(X_test)
mlpc = MLPClassifier(hidden_layer_sizes=(20, 20, 20), max_iter=500)
mlpc.fit(X_train, y_train)
print('Score:',mlpc.score(X_test, y_test))
t1 = time.time()
in1 = read_all()[0]
x = scaler.transform([np.asarray(in1).flatten()])
t2 = time.time()
print('Prediction: ', mlpc.predict(x))
print('Time: ',t2-t1)
xd = pd.DataFrame(X)
xd.head()
xd['y'] = y
xd['q'] = q
xd.head()
xd.to_csv('quad_small3.csv', index=False)
from IPython.display import clear_output
for i in range(100):
r, t = read_all()
print(np.asarray(r))
time.sleep(0.1)
clear_output(wait=True)
def read_av():
read = np.asarray(read_all()[0]).flatten()
for i in range(10):
read = read+(np.asarray(read_all()[0]).flatten())
return read/10
base = read_all()[0]
from matplotlib import pyplot as plt
%matplotlib inline
read = read_all()[0]
plt.imshow(np.asarray(base)-np.asarray(read))
base
```
# Image recon
```
# Import required libraries
from image_util import *
import skimage.filters
from matplotlib import pyplot as plt
import cairocffi as cairo
import math, random
import numpy as np
import pandas as pd
from IPython.display import Image
from scipy.interpolate import interp1d
import astra
%matplotlib inline
def r8_to_sino(readings):
sino = []
for e in range(8):
start = e*8 + (e+2)%8
end = e*8 + (e+6)%8
if end-start == 4:
sino.append(readings[start : end])
else:
r = readings[start : (e+1)*8]
for p in readings[e*8 : end]:
r.append(p)
sino.append(r)
return np.asarray(sino)
nviews = 8
ndetectors = 4
nvdetectors = 8
IMSIZE = 50
R = IMSIZE/2
D = IMSIZE/2
# Transforming from a round fan-beam to a fan-flat projection (See diagram)
beta = np.linspace(math.pi/8, 7*math.pi/8, ndetectors)
alpha = np.asarray([R*math.sin(b-math.pi/2)/(R**2 + D**2)**0.5 for b in beta])
tau = np.asarray([(R+D)*math.tan(a) for a in alpha])
tau_new = np.linspace(-(max(tau)/2), max(tau)/2, nvdetectors)
vol_geom = astra.create_vol_geom(IMSIZE, IMSIZE)
angles = np.linspace(0,2*math.pi,nviews);
d_size = (tau[-1]-tau[0])/nvdetectors
proj_geom= astra.create_proj_geom('fanflat', d_size, nvdetectors, angles, D, R);
proj_id = astra.create_projector('line_fanflat', proj_geom, vol_geom)
base = read_av()
np.asarray(base).reshape(8,8)
%%time
for i in range(1):
print(i)
r2 = read_av()
readings = (np.asarray(base)-np.asarray(r2))# - base
readings = r8_to_sino(readings.tolist()) # Get important ones and reorder
readings2 = []
for r in readings:
f = interp1d(tau, r, kind='cubic') # Can change to linear
readings2.append(f(tau_new))
sinogram_id = astra.data2d.create('-sino', proj_geom, np.asarray(readings2))
# Plotting sinogram - new (transformed) set of readings
plt.figure(num=None, figsize=(16, 10), dpi=80, facecolor='w', edgecolor='k')
ax1 = plt.subplot(1, 3, 1)
ax1.imshow(readings2) #<< Set title
# Doing the reconstruction, in this case with FBP
rec_id = astra.data2d.create('-vol', vol_geom)
cfg = astra.astra_dict('FBP')
cfg['ReconstructionDataId'] = rec_id
cfg['ProjectionDataId'] = sinogram_id
cfg['ProjectorId'] = proj_id
# Create the algorithm object from the configuration structure
alg_id = astra.algorithm.create(cfg)
astra.algorithm.run(alg_id, 1)
# Get the result
rec = astra.data2d.get(rec_id)
ax2 = plt.subplot(1, 3, 2)
ax2.imshow(rec)
norm_rec = rec/(np.amax(np.abs(rec)))
blurred = skimage.filters.gaussian(norm_rec, 3)
ax3 = plt.subplot(1, 3, 3)
ax3.imshow(blurred)
plt.savefig('r8s'+str(i) + '.png')
print(max(np.asarray(readings2).flatten()))
# Clean up.
astra.algorithm.delete(alg_id)
astra.data2d.delete(rec_id)
astra.data2d.delete(sinogram_id)
astra.projector.delete(proj_id)
np.linspace(math.pi/8, 7*math.pi/8, ndetectors)
np.linspace(0, math.pi, ndetectors)
r = []
y = []
y2 = []
for i in range(50):
r.append(np.asarray(read_all()[0]).flatten())
y.append(0)
y2.append(0)
for i in range(50):
r.append(np.asarray(read_all()[0]).flatten())
y.append(2)
y2.append(2)
from sklearn.model_selection import train_test_split
from sklearn.ensemble import RandomForestRegressor
from sklearn.ensemble import RandomForestClassifier
X_train, X_test, y_train, y_test = train_test_split(r, y)
regr = RandomForestClassifier(max_depth=5, random_state=0)
regr.fit(X_train, y_train)
regr.score(X_test, y_test)
regr.predict([np.asarray(read_all()[0]).flatten()])
df = pd.read_csv('r8_small_rotation.csv')
r = df[[str(i) for i in range(64)]]
y = df['Y']
from sklearn.neural_network import MLPClassifier, MLPRegressor
X_train, X_test, y_train, y_test = train_test_split(r, y)
scaler = StandardScaler()
scaler.fit(X_train)
X_train = scaler.transform(X_train)
X_test = scaler.transform(X_test)
mlpc = MLPClassifier(hidden_layer_sizes=(20, 20, 20), max_iter=400)
mlpc.fit(X_train, y_train)
print(mlpc.score(X_test, y_test))
from IPython.display import clear_output
av = 0
while True:
read = [np.asarray(read_all()[0]).flatten()]
read = scaler.transform(read)
print(mlpc.predict(read))
time.sleep(0.1)
clear_output(wait=True)
ser.read_all()
import pandas as pd
df1 = pd.DataFrame(r)
df1.head()
df1['Y'] = y
df1.head()
df1.to_csv('r8_small_rotation.csv', index=False)
```
| github_jupyter |
# Bar data
```
from ib_insync import *
util.startLoop()
ib = IB()
ib.connect('127.0.0.1', 7497, clientId=14)
```
## Historical data
To get the earliest date of available bar data the "head timestamp" can be requested:
```
contract = Stock('TSLA', 'SMART', 'USD')
ib.reqHeadTimeStamp(contract, whatToShow='TRADES', useRTH=True)
```
To request hourly data of the last 60 trading days:
```
bars = ib.reqHistoricalData(
contract,
endDateTime='',
durationStr='60 D',
barSizeSetting='1 hour',
whatToShow='TRADES',
useRTH=True,
formatDate=1)
bars[0]
```
Convert the list of bars to a data frame and print the first and last rows:
```
df = util.df(bars)
display(df.head())
display(df.tail())
```
Instruct the notebook to draw plot graphics inline:
```
%matplotlib inline
```
Plot the close data
```
df.plot(y='close');
```
There is also a utility function to plot bars as a candlestick plot. It can accept either a DataFrame or a list of bars. Here it will print the last 100 bars:
```
util.barplot(bars[-100:], title=contract.symbol);
```
## Historical data with realtime updates
A new feature of the API is to get live updates for historical bars. This is done by setting `endDateTime` to an empty string and the `keepUpToDate` parameter to `True`.
Let's get some bars with an keepUpToDate subscription:
```
contract = Forex('EURUSD')
bars = ib.reqHistoricalData(
contract,
endDateTime='',
durationStr='900 S',
barSizeSetting='10 secs',
whatToShow='MIDPOINT',
useRTH=True,
formatDate=1,
keepUpToDate=True)
```
Replot for every change of the last bar:
```
from IPython.display import display, clear_output
import matplotlib.pyplot as plt
def onBarUpdate(bars, hasNewBar):
plt.close()
plot = util.barplot(bars)
clear_output(wait=True)
display(plot)
ib.setCallback('barUpdate', onBarUpdate)
ib.sleep(60)
ib.cancelHistoricalData(bars)
```
Realtime bars
------------------
With ``reqRealTimeBars`` a subscription is started that sends a new bar every 5 seconds.
First we'll set up a callback for bar updates:
```
def onBarUpdate(bars, hasNewBar):
print(bars[-1])
ib.setCallback('barUpdate', onBarUpdate)
```
Then do the real request,
```
bars = ib.reqRealTimeBars(contract, 5, 'MIDPOINT', False)
```
let it run for half a minute and then cancel the realtime bars.
```
ib.sleep(30)
ib.cancelRealTimeBars(bars)
```
The advantage of reqRealTimeBars is that it behaves more robust when the connection to the IB server farms is interrupted. After the connection is restored, the bars from during the network outage will be backfilled and the live bars will resume.
reqHistoricalData + keepUpToDate will, at the moment of writing, leave the whole API inoperable after a network interruption.
```
ib.disconnect()
```
| github_jupyter |
```
%matplotlib inline
from matplotlib import pyplot as plt
import pandas as pd
import numpy as np
import psycopg2
from datetime import date
pd.options.display.max_columns = 40
pd.options.mode.chained_assignment = None # default='warn'
```
## Hack for Heat #8: Complaint resolution time revisited
In this post, I'm going to explore a simple way of dealing with the censoring issue in complaint resolution time: Basically, because we only care about heat complaints during the heat season, we can constrain our cases of interest to the heating season specifically (1<sup>st</sup> October to May 31<sup>st</sup>).
```
#pull from our database again:
connection = psycopg2.connect('dbname = threeoneone user= threeoneoneadmin password = threeoneoneadmin')
cursor = connection.cursor()
cursor.execute('''SELECT createddate, closeddate, borough, complainttype FROM service;''')
data = cursor.fetchall()
data = pd.DataFrame(data)
data.columns = ['createddate', 'closeddate', 'borough', 'complainttype']
data = data.loc[(data['complainttype'] == 'HEATING') | (data['complainttype'] == 'HEAT/HOT WATER') ]
```
So far, we've subsetted the data to only include the heating complaints. Now let's create a datetime mask. What we want is to have subsets of data for each of the heating seasons:
```
heatmonths = range(1, 6) + [10,11,12]
heatmonths
createdmask = data['createddate'].map(lambda x: (x.month in heatmonths))
closedmask = data['closeddate'].map(lambda x: (x.month in heatmonths) if x != None else False)
mask = createdmask & closedmask
heatseasondata = data.loc[mask]
```
How many heat/hot water complaints are created and closed inside vs. outside of heating season?
```
len(data.loc[mask]) #inside heating season
len(data.loc[~mask])#outside heating season
```
The next thing we want to do is ignore cases where the complaint was resolved in the *next* heating season:
```
prevmonths = range(1, 6)
nextmonths = [10,11,12]
heatseasondata['createdheatseason'] = [x.year if (x.month in prevmonths) else (x.year-1) for x in heatseasondata['createddate']]
heatseasondata.head()
heatseasondata['closedheatseason'] = [x.year if (x.month in prevmonths) else (x.year-1) for x in heatseasondata['closeddate']]
```
Now that we've done this, we can select only the cases where the closed date was in the same season as the created date:
```
heatseasondata = heatseasondata.loc[heatseasondata['createdheatseason'] == heatseasondata['closedheatseason']]
```
Okay, now we can calculate some average resolution times:
```
heatseasondata['resolutiontime'] = heatseasondata['closeddate'] - heatseasondata['createddate']
heatseasondata['resolutiontimeint'] = heatseasondata.resolutiontime.astype('timedelta64[D]')
resolutiontimedata = heatseasondata.groupby(by='createdheatseason').mean()['resolutiontimeint']
resolutiontimedata.to_csv('resolutiontimebyyear.csv')
resolutiontimedata
```
## Resolution times by year:
```
x = resolutiontimedata.index.values
y = resolutiontimedata.values
plt.figure(figsize=(12,10));
plt.plot(x,y);
```
## Resolution time by borough:
```
restimebyboro = heatseasondata.groupby(by=['borough', 'createdheatseason']).mean()['resolutiontimeint']
restimebyboro.to_csv('restimebyboro.csv')
restimebyboro = restimebyboro.loc[[x in range(2010,2017) for x in restimebyboro.index.get_level_values('createdheatseason')]]
boros = heatseasondata.borough.unique()
boroplots = {x:[] for x in boros}
for boro in boros:
boroplots[boro] = restimebyboro.xs(boro).values
boroplots.pop('Unspecified')
x = range(2010,2017)
plt.figure(figsize=(12,10));
for boro in boroplots:
plt.plot(x,boroplots[boro]);
plt.legend(boroplots.keys());
plt.xticks(x, [str(label) for label in x]);
```
I was told beforehand that the average resolution time from a similar analysis last year was about 3-5 days, so this looks about right.
| github_jupyter |
```
# basics
import os
import time
import numpy as np
import pandas as pd
# scipy and sklearn
from scipy.stats import entropy
from sklearn.metrics import balanced_accuracy_score
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import OneHotEncoder, StandardScaler
# plotting
import matplotlib
import matplotlib.pyplot as plt
import matplotlib.patches as mpatches
from matplotlib.collections import PatchCollection
import seaborn as sns
from IPython.display import Image
from matplotlib.patches import Rectangle
# CMDGB
import graphviz
from CMGDB import PlotMorseSets, PlotMorseGraph
# local
from data_loaders import retrieve_predictions
from models import *
from utils import get_model_weights, convert_weight_dict_to_dataframe, compute_morse_graph, \
compute_morse_graph_with_gpflow_gp, compute_order_retraction, PlotOrderRetraction
sns.set()
plt.style.use('ggplot')
%matplotlib inline
%load_ext autoreload
%autoreload 2
# suppress warnings
import warnings
warnings.filterwarnings('ignore')
```
# IRIS Morse Graphs
**Config:**
** Make sure to restart kernel before each new config run! Easiest way is to click on "Restart & Run All"
```
# Pick interesting_iris from: ["iris_baseline", "iris_layers_2", "iris_epoch_450"]
interesting_iris = "iris_epoch_450"
# Pick smallest_or_largest from: ["smallest", "largest"]
smallest_or_largest = "largest"
```
**Code:**
```
weights = get_model_weights("./data/{}".format(interesting_iris))
weights = convert_weight_dict_to_dataframe(weights)
final_weights = weights.loc[weights["epoch"] != "1"]
weight_cols = [c for c in list(final_weights.columns) if c not in ["model_id", "epoch", "val_loss"]]
std_df = pd.DataFrame(columns=["weight_name", "std"],
data=list(zip(weight_cols, [np.std(final_weights[col]) for col in weight_cols])))
std_df = std_df.sort_values(by="std")
if smallest_or_largest == "smallest":
two_weights = list(std_df["weight_name"])[:2]
elif smallest_or_largest == "largest":
two_weights = list(std_df["weight_name"])[-2:]
else:
raise ValueError()
morseg, mapg = compute_morse_graph(weights[['epoch'] + two_weights], phase_subdiv=15)
compute_order_retraction(morseg, mapg, title="{}_{}_2".format(interesting_iris, smallest_or_largest))
PlotMorseGraph(morseg)
graphviz.Source.from_file('Hasse.dot')
# This part of the code is to get the correct colors to appear in order retraction plot
# MUST BE SET MANUALLY FOR EACH RUN BASED ON ABOVE MORSE GRAPHS
# map the numbers in the colored morse graph to the numbers in the uncolored morse graph:
morse_nodes_map = {1: 0, 0:1}
with open('CMGDB_retract.txt','r') as infile:
retract_indices = []
retract_tiles = []
for i in range(mapg.num_vertices()):
index, tile = [int(x) for x in next(infile).split()]
retract_indices.append(index)
retract_tiles.append(tile)
PlotOrderRetraction(morseg, mapg, retract_tiles, retract_indices, morse_nodes_map)
```
| github_jupyter |
# Experiment Initialization
Here, I define the terms of my experiment, among them the location of the files in S3 (bucket and folder name), and each of the video prefixes (everything before the file extension) that I want to track.
Note that these videos should be similar-ish: while we can account for differences in mean intensities between videos, particle sizes should be approximately the same, and (slightly less important) particles should be moving at about the same order of magnitude speed. In this experiment, these videos were taken in 0.4% agarose gel at 100x magnification and 100.02 fps shutter speeds with nanoparticles of about 100nm in diameter.
```
to_track = []
result_futures = {}
start_knot = 29 #Must be unique number for every run on Cloudknot.
remote_folder = 'Gel_Studies/09_19_18_NP_concentration' #Folder in AWS S3 containing files to be analyzed
bucket = 'ccurtis.data'
vids = 10
concs = ['1', 'pt5', 'pt1', 'pt05']
for conc in concs:
for num in range(1, vids+1):
#to_track.append('100x_0_4_1_2_gel_{}_bulk_vid_{}'.format(vis, num))
to_track.append('{}uL_XY{}'.format(conc, '%02d' % num))
to_track
```
The videos used with this analysis are fairly large (2048 x 2048 pixels and 651 frames), and in cases like this, the tracking algorithm can quickly eat up RAM. In this case, we chose to crop the videos to 512 x 512 images such that we can run our jobs on smaller EC2 instances with 16GB of RAM.
Note that larger jobs can be made with user-defined functions such that splitting isn't necessary-- or perhaps an intermediate amount of memory that contains splitting, tracking, and msd calculation functions all performed on a single EC2 instance.
The compiled functions in the knotlets module require access to buckets on AWS. In this case, we will be using a publicly (read-only) bucket. If users want to run this notebook on their own, will have to transfer files from nancelab.publicfiles to their own bucket, as it requires writing to S3 buckets.
```
import diff_classifier.knotlets as kn
for prefix in to_track[8:]:
kn.split(prefix, remote_folder=remote_folder, bucket=bucket)
```
## Tracking predictor
Tracking normally requires user input in the form of tracking parameters e.g. particle radius, linking max distance, max frame gap etc. When large datasets aren't required, each video can be manageably manually tracked using the TrackMate GUI. However, when datasets get large e.g. >20 videos, this can become extremely arduous. For videos that are fairly similar, you can get away with using similar tracking parameters across all videos. However, one parameter that is a little more noisy that the others is the quality filter value. Quality is a numerical value that approximate how likely a particle is to be "real."
In this case, I built a predictor that estimates the quality filter value based on intensity distributions from the input images. Using a relatively small training dataset (5-20 videos), users can get fairly good estimates of quality filter values that can be used in parallelized tracking workflows.
Note: in the current setup, the predictor should be run in Python 3. While the code will run in Python 3, there are differences between the random number generators in Python2 and Python3 that I was not able to control for.
```
import os
import diff_classifier.imagej as ij
import boto3
import os.path as op
import diff_classifier.aws as aws
import diff_classifier.knotlets as kn
import numpy as np
from sklearn.externals import joblib
```
The regress_sys function should be run twice. When have_output is set to False, it generates a list of files that the user should manually track using Trackmate. Once the quality filter values are found, they can be used as input (y) to generate a regress object that can predict quality filter values for additional videos. Once y is assigned, set have_output to True and re-run the cell.
```
tnum=10 #number of training datasets
pref = []
for num in to_track:
for row in range(0, 4):
for col in range(0, 4):
pref.append("{}_{}_{}".format(num, row, col))
y = np.array([6.91, 6.06, 3.34, 3.5, 2.22, 2.8, 5.68, 5.28, 3.2, 1.4])
# Creates regression object based of training dataset composed of input images and manually
# calculated quality cutoffs from tracking with GUI interface.
regress = ij.regress_sys(remote_folder, pref, y, tnum, randselect=True,
have_output=True, bucket_name=bucket)
#Read up on how regress_sys works before running.
#Pickle object
filename = 'regress.obj'
with open(filename,'wb') as fp:
joblib.dump(regress,fp)
import boto3
s3 = boto3.client('s3')
aws.upload_s3(filename, remote_folder+'/'+filename, bucket_name=bucket)
```
Users should input all tracking parameters into the tparams object. Note that the quality value will be overwritten by values found using the quality predictor found above.
```
tparams1 = {'radius': 5.0, 'threshold': 0.0, 'do_median_filtering': False,
'quality': 5.0, 'xdims': (0, 511), 'ydims': (1, 511),
'median_intensity': 300.0, 'snr': 0.0, 'linking_max_distance': 12.0,
'gap_closing_max_distance': 18.0, 'max_frame_gap': 8,
'track_duration': 20.0}
# tparams2 = {'radius': 4.0, 'threshold': 0.0, 'do_median_filtering': False,
# 'quality': 10.0, 'xdims': (0, 511), 'ydims': (1, 511),
# 'median_intensity': 300.0, 'snr': 0.0, 'linking_max_distance': 8.0,
# 'gap_closing_max_distance': 12.0, 'max_frame_gap': 6,
# 'track_duration': 20.0}
```
## Cloudknot setup
Cloudknot requires the user to define a function that will be sent to multiple computers to run. In this case, the function knotlets.tracking will be used. We create a docker image that has the required installations (defined by the requirements.txt file from diff_classifier on Github, and the base Docker Image below that has Fiji pre-installed in the correct location.
Note that I modify the Docker image below such that the correct version of boto3 is installed. For some reason, versions later than 1.5.28 error out, so I specified 5.28 as the correct version. Run my_image.build below to double-check that the Docker image is successfully built prior to submitting the job to Cloudknot.
```
import cloudknot as ck
import os.path as op
github_installs=('https://github.com/ccurtis7/diff_classifier.git@Chad')
#my_image = ck.DockerImage(func=kn.tracking, base_image='arokem/python3-fiji:0.3', github_installs=github_installs)
my_image = ck.DockerImage(func=kn.assemble_msds, base_image='arokem/python3-fiji:0.3', github_installs=github_installs)
docker_file = open(my_image.docker_path)
docker_string = docker_file.read()
docker_file.close()
req = open(op.join(op.split(my_image.docker_path)[0], 'requirements.txt'))
req_string = req.read()
req.close()
new_req = req_string[0:req_string.find('\n')-4]+'5.28'+ req_string[req_string.find('\n'):]
req_overwrite = open(op.join(op.split(my_image.docker_path)[0], 'requirements.txt'), 'w')
req_overwrite.write(new_req)
req_overwrite.close()
my_image.build("0.1", image_name="test_image")
```
The object all_maps is an iterable containing all the inputs sent to Cloudknot. This is useful, because if the user needs to modify some of the tracking parameters for a single video, this can be done prior to submission to Cloudknot.
```
names = []
all_maps = []
for prefix in to_track:
for i in range(0, 4):
for j in range(0, 4):
names.append('{}_{}_{}'.format(prefix, i, j))
all_maps.append(('{}_{}_{}'.format(prefix, i, j), remote_folder, bucket, 'regress.obj',
4, 4, (512, 512), tparams))
all_maps
```
The Cloudknot knot object sets up the compute environment which will run the code. Note that the name must be unique. Every time you submit a new knot, you should change the name. I do this with the variable start_knot, which I vary for each run.
If larger jobs are anticipated, users can adjust both RAM and storage with the memory and image_id variables. Memory specifies the amount of RAM to be used. Users can build a customized AMI with as much space as they need, and enter the ID into image_ID. Read the Cloudknot documentation for more details.
```
knot = ck.Knot(name='download_and_track_{}_b{}'.format('chad22', start_knot),
docker_image = my_image,
memory = 16000,
resource_type = "SPOT",
bid_percentage = 100,
#image_id = 'ami-015a1b4cd3895860b', #May need to change this line
pars_policies=('AmazonS3FullAccess',),
)
result_futures = knot.map(all_maps, starmap=True)
result_futures2 = knot.map(all_maps3, starmap=True)
knot.clobber()
knot2 = ck.Knot(name='download_and_track_{}_b{}'.format('chad30', start_knot),
docker_image = my_image,
memory = 16000,
resource_type = "SPOT",
bid_percentage = 100,
#image_id = 'ami-015a1b4cd3895860b', #May need to change this line
pars_policies=('AmazonS3FullAccess',),
)
result_futures3 = knot2.map(all_maps3, starmap=True)
knot2.clobber()
knot4 = ck.Knot(name='download_and_track_{}_b{}'.format('chad31', start_knot),
docker_image = my_image,
memory = 16000,
resource_type = "SPOT",
bid_percentage = 100,
#image_id = 'ami-015a1b4cd3895860b', #May need to change this line
pars_policies=('AmazonS3FullAccess',),
)
result_futures4 = knot4.map(all_maps3, starmap=True)
knot4.clobber()
ck.aws.get_region()
names = []
all_maps2 = []
for prefix in to_track:
all_maps2.append((prefix, remote_folder, bucket, (512, 512), 651, 4, 4))
all_maps2
knot3 = ck.Knot(name='download_and_track_{}_b{}'.format('chad33', start_knot),
docker_image = my_image,
memory = 64000,
resource_type = "SPOT",
bid_percentage = 100,
#image_id = 'ami-015a1b4cd3895860b', #May need to change this line
pars_policies=('AmazonS3FullAccess',),
)
result_futures5 = knot3.map(all_maps2, starmap=True)
knot3.clobber()
ck.aws.get_region()
ck.aws.set_region('us-east-1')
ck.aws.get_region()
knot2 = ck.Knot(name='download_and_track_{}_b{}'.format('chad2', start_knot),
docker_image = my_image,
memory = 144000,
resource_type = "SPOT",
bid_percentage = 100,
#image_id = 'ami-0e00afdf500081a0d', #May need to change this line
pars_policies=('AmazonS3FullAccess',))
result_futures2 = knot2.map(all_maps, starmap=True)
knot3 = ck.Knot(name='download_and_track_{}_b{}'.format('chad3', start_knot),
docker_image = my_image,
memory = 144000,
resource_type = "SPOT",
bid_percentage = 100,
#image_id = 'ami-0e00afdf500081a0d', #May need to change this line
pars_policies=('AmazonS3FullAccess',))
result_futures3 = knot3.map(all_maps, starmap=True)
knot4 = ck.Knot(name='download_and_track_{}_b{}'.format('chad4', start_knot),
docker_image = my_image,
memory = 144000,
resource_type = "SPOT",
bid_percentage = 100,
#image_id = 'ami-0e00afdf500081a0d', #May need to change this line
pars_policies=('AmazonS3FullAccess',))
result_futures4 = knot4.map(all_maps, starmap=True)
knot5 = ck.Knot(name='download_and_track_{}_b{}'.format('chad5', start_knot),
docker_image = my_image,
memory = 144000,
resource_type = "SPOT",
bid_percentage = 100,
#image_id = 'ami-0e00afdf500081a0d', #May need to change this line
pars_policies=('AmazonS3FullAccess',))
result_futures5 = knot5.map(all_maps, starmap=True)
ck.aws.set_region('eu-west-1')
knot5.clobber()
tparams2 = {'radius': 3.5, 'threshold': 0.0, 'do_median_filtering': False,
'quality': 10.0, 'xdims': (0, 511), 'ydims': (1, 511),
'median_intensity': 300.0, 'snr': 0.0, 'linking_max_distance': 15.0,
'gap_closing_max_distance': 22.0, 'max_frame_gap': 5,
'track_duration': 20.0}
missing = []
all_maps3 = []
import boto3
import botocore
s3 = boto3.resource('s3')
for name in names:
try:
s3.Object(bucket, '{}/Traj_{}.csv'.format(remote_folder, name)).load()
except botocore.exceptions.ClientError as e:
if e.response['Error']['Code'] == "404":
missing.append(name)
all_maps3.append((name, remote_folder, bucket, 'regress.obj',
4, 4, (512, 512), tparams1))
else:
print('Something else has gone wrong')
all_maps3
import diff_classifier.aws as aws
old_folder = 'Gel_Studies/08_14_18_gel_validation/old_msds2'
for name in missing:
filename = 'Traj_{}.csv'.format(name)
aws.download_s3('{}/{}'.format(old_folder, filename), filename, bucket_name=bucket)
aws.upload_s3(filename, '{}/{}'.format(remote_folder, filename), bucket_name=bucket)
```
Users can monitor the progress of their job in the Batch interface. Once the code is complete, users should clobber their knot to make sure that all AWS resources are removed.
```
knot.clobber()
```
## Downstream analysis and visualization
The knotlet.assemble_msds function (which can also potentially be submitted to Cloudknot as well for large jobs) calculates the mean squared displacements and trajectory features from the raw trajectory csv files found from the Cloudknot submission. It accesses them from the S3 bucket to which they were saved.
```
for prefix in to_track:
kn.assemble_msds(prefix, remote_folder, bucket=bucket)
print('Successfully output msds for {}'.format(prefix))
for prefix in to_track[5:7]:
kn.assemble_msds(prefix, remote_folder, bucket='ccurtis.data')
print('Successfully output msds for {}'.format(prefix))
all_maps2 = []
for prefix in to_track:
all_maps2.append((prefix, remote_folder, bucket, 'regress100.obj',
4, 4, (512, 512), tparams))
knot = ck.Knot(name='download_and_track_{}_b{}'.format('chad', start_knot),
docker_image = my_image,
memory = 16000,
resource_type = "SPOT",
bid_percentage = 100,
#image_id = 'ami-0e00afdf500081a0d', #May need to change this line
pars_policies=('AmazonS3FullAccess',))
```
Diff_classifier includes some useful imaging tools as well, including checking trajectories, plotting heatmaps of trajectory features, distributions of diffusion coefficients, and MSD plots.
```
import diff_classifier.heatmaps as hm
import diff_classifier.aws as aws
prefix = to_track[1]
msds = 'msd_{}.csv'.format(prefix)
feat = 'features_{}.csv'.format(prefix)
aws.download_s3('{}/{}'.format(remote_folder, msds), msds, bucket_name=bucket)
aws.download_s3('{}/{}'.format(remote_folder, feat), feat, bucket_name=bucket)
hm.plot_trajectories(prefix, upload=False, figsize=(8, 8))
geomean, geoSEM = hm.plot_individual_msds(prefix, x_range=10, y_range=300, umppx=1, fps=1, upload=False)
hm.plot_heatmap(prefix, upload=False)
hm.plot_particles_in_frame(prefix, y_range=6000, upload=False)
missing = []
all_maps2 = []
import boto3
import botocore
s3 = boto3.resource('s3')
for name in names:
try:
s3.Object(bucket, '{}/Traj_{}.csv'.format(remote_folder, name)).load()
except botocore.exceptions.ClientError as e:
if e.response['Error']['Code'] == "404":
missing.append(name)
#all_maps2.append((name, remote_folder, bucket, 'regress.obj',
# 4, 4, (512, 512), tparams2))
else:
print('Something else has gone wrong')
missing = ['PS_COOH_2mM_XY05_1_1', 'PS_NH2_2mM_XY04_2_1']
all_maps
kn.tracking(missing[0], remote_folder, bucket=bucket, tparams=tparams1)
kn.tracking(missing[1], remote_folder, bucket=bucket, tparams=tparams1)
```
| github_jupyter |
```
from molmap import model as molmodel
import molmap
import matplotlib.pyplot as plt
import pandas as pd
from tqdm import tqdm
from joblib import load, dump
tqdm.pandas(ascii=True)
import numpy as np
import tensorflow as tf
import os
os.environ["CUDA_VISIBLE_DEVICES"]="5"
np.random.seed(123)
tf.compat.v1.set_random_seed(123)
#tmp_feature_dir = './tmpignore'
tmp_feature_dir = '/raid/shenwanxiang/tempignore'
if not os.path.exists(tmp_feature_dir):
os.makedirs(tmp_feature_dir)
mp1 = molmap.loadmap('../descriptor.mp')
mp2 = molmap.loadmap('../fingerprint.mp')
task_name = 'PCBA'
from chembench import load_data
df, induces = load_data(task_name)
print(len(induces[0][0]), len(induces[0][1]), len(induces[0][2]), df.shape)
nan_idx = df[df.smiles.isna()].index.to_list()
nan_idx
MASK = -1
smiles_col = df.columns[0]
values_col = df.columns[1:]
Y = df[values_col].astype('float32').fillna(MASK).values
if Y.shape[1] == 0:
Y = Y.reshape(-1, 1)
Y = Y.astype('float32')
Y.shape
```
### batch extract features
```
batch = 30
xs = np.array_split(df.smiles.to_list(), batch)
X1_name_all = os.path.join(tmp_feature_dir, 'X1_%s.data' % (task_name))
X2_name_all = os.path.join(tmp_feature_dir, 'X2_%s.data' % (task_name))
if os.path.exists(X1_name_all) & os.path.exists(X2_name_all):
X1 = load(X1_name_all)
X2 = load(X2_name_all)
else:
## descriptors
X1s = []
for i, batch_smiles in tqdm(enumerate(xs), ascii=True):
ii = str(i).zfill(2)
X1_name = os.path.join(tmp_feature_dir, 'X1_%s_%s.data' % (task_name, ii))
print('save to %s' % X1_name)
if not os.path.exists(X1_name):
X1 = mp1.batch_transform(batch_smiles, n_jobs = 8)
X1 = X1.astype('float32')
dump(X1, X1_name)
else:
X1 = load(X1_name)
X1s.append(X1)
del X1
X1 = np.concatenate(X1s)
del X1s
dump(X1, X1_name_all)
## fingerprint
X2s = []
for i, batch_smiles in tqdm(enumerate(xs), ascii=True):
ii = str(i).zfill(2)
X2_name = os.path.join(tmp_feature_dir, 'X2_%s_%s.data' % (task_name, ii))
if not os.path.exists(X2_name):
X2 = mp2.batch_transform(batch_smiles, n_jobs = 8)
X2 = X2.astype('float32')
dump(X2, X2_name)
else:
X2 = load(X2_name)
X2s.append(X2)
del X2
X2 = np.concatenate(X2s)
del X2s
dump(X2, X2_name_all)
molmap1_size = X1.shape[1:]
molmap2_size = X2.shape[1:]
X1.shape
def get_pos_weights(trainY):
"""pos_weights: neg_n / pos_n """
dfY = pd.DataFrame(trainY)
pos = dfY == 1
pos_n = pos.sum(axis=0)
neg = dfY == 0
neg_n = neg.sum(axis=0)
pos_weights = (neg_n / pos_n).values
neg_weights = (pos_n / neg_n).values
return pos_weights, neg_weights
prcs_metrics = ['MUV', 'PCBA'] #
epochs = 800
patience = 20 #early stopping, dual to large computation cost, the larger dataset set small waitig patience for early stopping
dense_layers = [512] #128 outputs
batch_size = 128
lr = 1e-4
weight_decay = 0
monitor = 'val_auc'
dense_avf = 'relu'
last_avf = None #sigmoid in loss
if task_name in prcs_metrics:
metric = 'PRC'
else:
metric = 'ROC'
results = []
for i, split_idxs in enumerate(induces):
train_idx, valid_idx, test_idx = split_idxs
train_idx = list(set(train_idx) - set(nan_idx))
valid_idx = list(set(valid_idx) - set(nan_idx))
test_idx = list(set(test_idx) - set(nan_idx))
print(len(train_idx), len(valid_idx), len(test_idx))
trainX = (X1[train_idx], X2[train_idx])
trainY = Y[train_idx]
validX = (X1[valid_idx], X2[valid_idx])
validY = Y[valid_idx]
testX = (X1[test_idx], X2[test_idx])
testY = Y[test_idx]
pos_weights, neg_weights = get_pos_weights(trainY)
loss = lambda y_true, y_pred: molmodel.loss.weighted_cross_entropy(y_true,y_pred, neg_weights, MASK = MASK)
#loss = molmodel.loss.cross_entropy
model = molmodel.net.DoublePathNet(molmap1_size, molmap2_size,
n_outputs=Y.shape[-1],
dense_layers=dense_layers,
dense_avf = dense_avf,
last_avf=last_avf)
opt = tf.keras.optimizers.Adam(learning_rate=lr, beta_1=0.9, beta_2=0.999, epsilon=1e-08, decay=0.0) #
#import tensorflow_addons as tfa
#opt = tfa.optimizers.AdamW(weight_decay = 0.001,learning_rate=0.001)
model.compile(optimizer = opt, loss = loss)
performance = molmodel.cbks.CLA_EarlyStoppingAndPerformance((trainX, trainY),
(validX, validY),
patience = patience,
criteria = monitor,
metric = metric,
)
model.fit(trainX, trainY, batch_size=batch_size,
epochs=epochs, verbose= 0, shuffle = True,
validation_data = (validX, validY),
callbacks=[performance])
best_epoch = performance.best_epoch
trainable_params = model.count_params()
train_aucs = performance.evaluate(trainX, trainY)
valid_aucs = performance.evaluate(validX, validY)
test_aucs = performance.evaluate(testX, testY)
final_res = {
'task_name':task_name,
'train_auc':np.nanmean(train_aucs),
'valid_auc':np.nanmean(valid_aucs),
'test_auc':np.nanmean(test_aucs),
'metric':metric,
'# trainable params': trainable_params,
'best_epoch': best_epoch,
'batch_size':batch_size,
'lr': lr,
'weight_decay':weight_decay
}
print(final_res)
del model, trainX, validX, testX
results.append(final_res)
pd.DataFrame(results)
pd.DataFrame(results).test_auc.mean()
pd.DataFrame(results).test_auc.std()
pd.DataFrame(results).to_csv('./results/%s.csv' % task_name)
```
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
import os
import json
import numpy as np, pandas as pd
import matplotlib.pyplot as plt, seaborn as sns
from tqdm import tqdm, tqdm_notebook
from pathlib import Path
from collections import Counter
from sklearn.decomposition import PCA
from sklearn.preprocessing import StandardScaler
from ast import literal_eval
from functools import partial
import pickle
sns.set()
DATA = Path('../../data')
RAW = DATA/'raw'
PROCESSED = DATA/'processed'
SUBMISSIONS = DATA/'submissions'
%%time
product = pd.read_csv(RAW/'productid_category.csv', low_memory=False)
train_tracking = pd.read_csv(RAW/'train_tracking.csv', low_memory=False)
test_tracking = pd.read_csv(RAW/'test_tracking.csv', low_memory=False)
train_session = pd.read_csv(RAW/'train_session.csv', low_memory=False)
test_session = pd.read_csv(RAW/'random_submission.csv', low_memory=False)
train_features = train_session.copy()
test_features = test_session.copy()
```
# Mapped actions
```
def set_event_types(tracking):
def extract_page(x):
pages_types = ['_LR', '_PA', '_LP', '_CAROUSEL', '_SHOW_CASE']
pages = ['CAROUSEL', 'PA', 'SHOW_CASE']
pages_map = [['PURCHASE_PRODUCT_UNKNOW_ORIGIN', 'UNKNOWN'], ['LIST_PRODUCT', 'LP'], ['SEARCH', 'LR']]
for pages_type in pages_types:
if x.endswith(pages_type):
return x[-len(pages_type)+1:]
for page in pages:
if x == page:
return x
for page_map in pages_map:
if x == page_map[0]:
return page_map[1]
return '::' + x
def extract_event(x):
page, _type, type_s = x
concatenated = _type
if page == 'UNKNOWN':
return page
actions = ['PRODUCT', 'ADD_TO_BASKET', 'PURCHASE_PRODUCT']
for action in actions:
if action in concatenated:
return page + '_' + action
return page
def extract_action(x):
actions = ['PRODUCT', 'ADD_TO_BASKET', 'PURCHASE_PRODUCT']
event_type = x
for action in actions:
if action in event_type:
return action
return 'None'
tracking['page_n_type'] = list(zip(tracking.type.apply(extract_page), tracking.type, tracking.type_simplified))
tracking['event_type'] = tracking.page_n_type.apply(extract_event)
tracking['action_type'] = tracking.event_type.apply(extract_action)
return tracking
def map_actions(tracking):
tracking = tracking.copy()
columns = list(tracking.columns.values)
tracking = set_event_types(tracking)
event_list = ['CAROUSEL', 'PA', 'LR', 'LR_ADD_TO_BASKET', 'LR_PRODUCT',
'SHOW_CASE', 'UNKNOWN', 'PA_PRODUCT', 'CAROUSEL_PRODUCT',
'CAROUSEL_ADD_TO_BASKET', 'LP_PRODUCT', 'SHOW_CASE_PRODUCT',
'LP_ADD_TO_BASKET', 'PA_ADD_TO_BASKET', 'SHOW_CASE_ADD_TO_BASKET']
event_dict = dict(zip(sorted(event_list), range(len(event_list)+1)))
actions = ['None', 'PRODUCT', 'ADD_TO_BASKET', 'PURCHASE_PRODUCT']
action_dict = dict(zip(sorted(actions), range(len(actions))))
tracking['event_id'] = tracking.event_type.apply(lambda x: event_dict[x])
tracking['action_id'] = tracking.action_type.apply(lambda x: action_dict[x])
return tracking[columns + ['event_id', 'action_id']]
tracking = train_tracking.copy()
```
**Algorithm:**
```
columns = list(tracking.columns.values)
tracking = set_event_types(tracking)
event_list = ['CAROUSEL', 'PA', 'LR', 'LR_ADD_TO_BASKET', 'LR_PRODUCT',
'SHOW_CASE', 'UNKNOWN', 'PA_PRODUCT', 'CAROUSEL_PRODUCT',
'CAROUSEL_ADD_TO_BASKET', 'LP_PRODUCT', 'SHOW_CASE_PRODUCT',
'LP_ADD_TO_BASKET', 'PA_ADD_TO_BASKET', 'SHOW_CASE_ADD_TO_BASKET']
event_dict = dict(zip(sorted(event_list), range(len(event_list))))
tracking['event_id'] = tracking.event_type.apply(lambda x: event_dict[x])
tracking = tracking[columns + ['event_id']]
```
**Test:**
```
str(len(tracking.head().event_id.values == map_actions(train_tracking).head().event_id.values)/len(train_tracking)*100) + '%'
```
# Timestamp series in session
```
def duration_to_seconds2(tracking):
if not 'timestamp' in tracking.columns:
tracking['timestamp'] = pd.to_timedelta(tracking.duration).dt.total_seconds()
return tracking
def duration_timestamp(features, tracking):
return features
def events_timeseries(features, tracking):
tracking = map_actions(duration_to_seconds2(tracking))
group = tracking.sort_values(['timestamp']).groupby('sid')
eventseries = group.event_id.agg(list)
features = test_features
tracking = test_tracking
```
**Dimensiones:** Sesiones, ventana (0, 30), features <br>
**Output:** mapa de sesiones, cubo de features
```
tracking = map_actions(duration_to_seconds2(tracking))
tracking.event_id.unique()
def to_timeseries_features(tstamps):
series = pd.Series(list(tstamps))
ts = list(tstamps)
dts = series.diff().values
dts[0] = 0
max_dts = max(max(dts),1)
dts_perc = dts/max_dts
return [ts, dts_perc]
def to_eventseries(evs):
return list(evs)
group = tracking.sort_values(['timestamp']).groupby('sid')
timeseries = group.timestamp.agg(partial(to_timeseries_features))
eventseries = group.event_id.agg(to_eventseries)
sessions = tracking.sid.unique()
sessions_map = dict(zip(sorted(sessions), range(len(sessions))))
len(sessions_map), len(sessions)
WINDOW_SIZE = 15
# Building 3D variable
frames = np.zeros((len(sessions), WINDOW_SIZE, 2))
for session in timeseries.index:
session_id = sessions_map[session]
n_actions = len(timeseries[session])
featblock = np.array(timeseries[session]).T
if featblock.shape[0] == WINDOW_SIZE:
pass
elif featblock.shape[0] < WINDOW_SIZE:
zeros = np.zeros((WINDOW_SIZE - featblock.shape[0], 2))
featblock = np.concatenate((zeros, featblock))
else:
featblock = featblock[-WINDOW_SIZE:]
frames[session_id] = featblock
shape = frames[:,:,0].shape
frames[:,:,0].mean()
scaler.fit(frames[:,:,0].reshape((-1, 1)))
ts_norm = scaler.transform(frames[:,:,0].reshape((-1, 1))).reshape(shape)
zero_pos = scaler.transform(np.zeros(shape).reshape((-1, 1)))[0][0]
zero_pos
frames[:,:,0] = ts_norm-zero_pos
np.save(PROCESSED/'f_duration_test.npy', frames)
with open(PROCESSED/'f_sessionsmap_test.pkl', 'wb') as handle:
# json.dump(sessions_map, handle)
pickle.dump(sessions_map, handle, protocol=pickle.HIGHEST_PROTOCOL)
WINDOW_SIZE = 15
# Building 3D variable
frames_events = np.zeros((len(sessions), WINDOW_SIZE, 1))
print(frames_events.shape)
for session in eventseries.index:
session_id = sessions_map[session]
n_actions = len(eventseries[session])
featblock = np.array([eventseries[session]]).T
if featblock.shape[0] == WINDOW_SIZE:
pass
elif featblock.shape[0] < WINDOW_SIZE:
zeros = np.zeros((WINDOW_SIZE - featblock.shape[0], 1))
featblock = np.concatenate((zeros, featblock))
else:
featblock = featblock[-WINDOW_SIZE:]
frames_events[session_id] = featblock
```
# Number of actions series
### ADD_TO_BASKET
```
features = test_features
tracking = test_tracking
WINDOW_SIZE = 30
with open(PROCESSED/'f_sessionsmap_test.pkl', 'rb') as f:
sessions_map = pickle.load(f)
# sessions = tracking.sid.unique()
# sessions_map = dict(zip(sorted(sessions), range(len(sessions))))
# len(sessions_map), len(sessions)
dict(zip(sorted(tracking.type.unique()), range(tracking.type.nunique())))
actions = ['None', 'PRODUCT', 'ADD_TO_BASKET', 'PURCHASE_PRODUCT']
action_dict = dict(zip(sorted(actions), range(len(actions))))
tracking = map_actions(duration_to_seconds2(tracking))
ADD_TO_BASKET = action_dict['ADD_TO_BASKET']
PRODUCT = action_dict['PRODUCT']
PURCHASE_PRODUCT = action_dict['PURCHASE_PRODUCT']
def to_action_series(x):
actions = np.array(list(x))
if len(actions) > WINDOW_SIZE:
actions = actions[-WINDOW_SIZE:]
default_len = np.array([
np.cumsum(actions == ADD_TO_BASKET),
np.cumsum(actions == PRODUCT),
np.cumsum(actions == PURCHASE_PRODUCT)
])
if default_len.shape[1] <= WINDOW_SIZE:
result = np.zeros((3, WINDOW_SIZE))
result[:,-default_len.shape[1]:] = default_len
else:
result = default_len[:,-WINDOW_SIZE:]
return result
group = tracking.sort_values(['timestamp']).groupby('sid')
action_series = group.action_id.apply(to_action_series)
df = action_series.to_frame().copy()
df['sid'] = pd.Series(df.index.values).apply(lambda x: sessions_map[x]).values
df
action_series2 = df.set_index('sid').sort_index()['action_id'].values
action_seq = np.stack(action_series2).reshape((-1, WINDOW_SIZE, 3))
shape = action_seq.shape
pd.Series(action_seq[:,:,0].reshape(-1, 1).T[0]).describe()
# action_seq[:,:,0].T.shape
action_seq = action_seq/WINDOW_SIZE
np.save(PROCESSED/'f_actions_test.npy', action_seq)
# with open(PROCESSED/'f_sessionsmap_test.pkl', 'wb') as handle:
# pickle.dump(sessions_map, handle, protocol=pickle.HIGHEST_PROTOCOL)
```
# Number of products added to basket
```
features = train_features
tracking = train_tracking
WINDOW_SIZE = 30
with open(PROCESSED/'f_sessionsmap_train.pkl', 'rb') as f:
sessions_map = pickle.load(f)
tracking = map_actions(duration_to_seconds2(tracking))
def to_q_series(x):
qs = np.array(list(x))
if len(qs) > WINDOW_SIZE:
qs = qs[-WINDOW_SIZE:]
default_len = np.array([
np.cumsum(qs)
])
if default_len.shape[1] <= WINDOW_SIZE:
result = np.zeros((1, WINDOW_SIZE))
result[:,-default_len.shape[1]:] = default_len
else:
result = default_len[:,-WINDOW_SIZE:]
return result
# tracking['new_id'] = tracking.sid.apply(lambda x: sessions_map[x])
# tracking = tracking.reset_index('new_id', drop=True)
tracking['filled_q'] = tracking.quantity.fillna(0)
group = tracking.sort_values(['timestamp']).groupby('sid')
q_series = group.filled_q.apply(to_q_series)
df = q_series.to_frame().copy()
df['sid'] = df.index
df['sid'] = df.sid.apply(lambda x: sessions_map[x])
q_seq = df.set_index('sid').values
q_seq.shape
q_series2 = df.set_index('sid').sort_index()['filled_q'].values
q_seq = np.stack(q_series2).reshape((-1, WINDOW_SIZE, 1))
shape = q_seq.shape
shape
scaler = StandardScaler()
scaler.fit(q_seq.reshape(-1, 1))
# action_seq[:,:,0].T.shape
new_q = scaler.transform(q_seq.reshape(-1, 1)).reshape(shape)
new_q[:,-1,0]
np.save(PROCESSED/'f_quantity_train.npy', new_q)
# with open(PROCESSED/'f_sessionsmap_test.pkl', 'wb') as handle:
# pickle.dump(sessions_map, handle, protocol=pickle.HIGHEST_PROTOCOL)
```
| github_jupyter |
# **YOLO V3 OBJECT DETECTOR**
```
!git clone https://github.com/pjreddie/darknet
!cd darknet
!make
! wget https://pjreddie.com/media/files/yolov3.weights
! ./darknet detector test cfg/coco.data cfg/yolov3.cfg yolov3.weights ./darknet/data/horses.jpg
!git clone https://github.com/pjreddie/darknet
import os
import re
import cv2 # opencv library
import numpy as np
from os.path import isfile, join
import matplotlib.pyplot as plt
import zipfile
import skimage.io as skio
import skimage.transform as sktr
# get file names of the frames
img_dir = '../input/fames-1/frames'
col_frames = os.listdir(img_dir)
# empty list to store the frames
col_images=[]
for idx,item in enumerate(col_frames):
#print(idx,item)
col_images.append(sktr.resize(skio.imread(img_dir+'/'+str(item)),(224,224)))
# read the frames
#img = cv2.imread('../input/fames-1/frames'+item)
# append the frames to the list
#col_images.append(img)
print((col_images))
import skimage.io as skio
import skimage.transform as sktr
img_dir = '../input/fames-1/frames'
i = 0
col_images = []
for filename in os.listdir(img_dir):
if(i>=116):
filepath = os.path.join(img_dir,filename)
col_images.append(sktr.resize(skio.imread(filepath),(224,224)))
else:
i = i+1
col_img_arr = np.array(col_images)
print(col_img_arr.shape)
# plot 13th frame
i = 13
for frame in [i, i+1]:
plt.imshow(col_images[i])
plt.title("frame: "+str(frame))
plt.show()
# convert the frames to grayscale
from skimage.color import rgb2gray
grayA = rgb2gray(col_images[i])
grayB = rgb2gray(col_images[i+1])
# plot the image after frame differencing
plt.imshow(cv2.absdiff(grayB, grayA), cmap = 'gray')
plt.show()
diff_image = cv2.absdiff(grayB, grayA)
# perform image thresholding
ret, thresh = cv2.threshold(diff_image,60/255,1, cv2.THRESH_BINARY)
# plot image after thresholding
plt.imshow(thresh, cmap = 'gray')
plt.show()
# apply image dilation
kernel = np.ones((3,3),np.uint8)
dilated = cv2.dilate(thresh,kernel,iterations = 1)
# plot dilated image
plt.imshow(dilated, cmap = 'gray')
plt.show()
# plot vehicle detection zone
#plt.imshow(dilated)
dil = (cv2.line(dilated,(0,125),(256,125),(100/255,0,0)))
plt.imshow(dil)
plt.show()
# find contours
thresh = cv2.convertScaleAbs(thresh)
contours, hierarchy = cv2.findContours(thresh,cv2.RETR_TREE,cv2.CHAIN_APPROX_NONE)
valid_cntrs = []
for i,cntr in enumerate(contours):
x,y,w,h = cv2.boundingRect(cntr)
if (x <= 200) & (y >= 80) & (cv2.contourArea(cntr) >= 25):
valid_cntrs.append(cntr)
# count of discovered contours
len(valid_cntrs)
dmy = col_images[13].copy()
cv2.drawContours(dmy, valid_cntrs, -1, (127,200,0), 2)
cv2.line(dmy, (0,125),(256,125),(100, 255, 255))
plt.imshow(dmy)
plt.show()
# kernel for image dilation
kernel = np.ones((4,4),np.uint8)
# font style
font = cv2.FONT_HERSHEY_SIMPLEX
# directory to save the ouput frames
pathIn = "../input/output/"
frame_out = []
for i in range(len(col_images)-1):
# frame differencing
grayA = rgb2gray(col_images[i])
grayB = rgb2gray(col_images[i])
diff_image = cv2.absdiff(grayB, grayA)
diff_image = cv2.convertScaleAbs(diff_image)
# image thresholding
ret, thresh = cv2.threshold(diff_image, 30, 255, cv2.THRESH_BINARY)
# image dilation
dilated = cv2.dilate(thresh,kernel,iterations = 1)
# find contours
contours, hierarchy = cv2.findContours(dilated.copy(), cv2.RETR_TREE,cv2.CHAIN_APPROX_NONE)
# shortlist contours appearing in the detection zone
valid_cntrs = []
for cntr in contours:
x,y,w,h = cv2.boundingRect(cntr)
if (x <= 200) & (y >= 125) & (cv2.contourArea(cntr) >= 25):
if (y >= 90) & (cv2.contourArea(cntr) < 40):
break
valid_cntrs.append(cntr)
# add contours to original frames
dmy = col_images[i].copy()
cv2.drawContours(dmy, valid_cntrs, -1, (127,200,0), 2)
cv2.putText(dmy, "vehicles detected: " + str(len(valid_cntrs)), (55, 15), font, 0.6, (0, 180, 0), 2)
cv2.line(dmy, (0, 125),(256,125),(100, 255, 255))
cv2.imwrite(pathIn+str(i)+'.png',dmy)
# specify video name
pathOut = 'vehicle_detection.mp4'
# specify frames per second
fps = 14.0
frame_array = []
files = [f for f in os.listdir(pathIn) if isfile(join(pathIn, f))]
for i in range(len(files)):
filename=pathIn +'/'+str(files[i])
#read frames
img = cv2.imread(filename)
height, width, layers = img.shape
size = (width,height)
#inserting the frames into an image array
frame_array.append(img)
out = cv2.VideoWriter(pathOut,cv2.VideoWriter_fourcc(*'DIVX'), fps, (224,224))
for i in range(len(frame_array)):
# writing to a image array
out.write(frame_array[i])
out.release()
```
| github_jupyter |
# Lecture 3: Logistic Regression and Text Models
***
<img src="figs/logregwordcloud.png",width=1000,height=50>
### Problem 1: Logistic Regression for 2D Continuous Features
***
In the video lecture you saw some examples of using logistic regression to do binary classification on text data (SPAM vs HAM) and on 1D continuous data. In this problem we'll look at logistic regression for 2D continuous data. The data we'll use are <a href="https://www.math.umd.edu/~petersd/666/html/iris_with_labels.jpg">sepal</a> measurements from the ubiquitous *iris* dataset.
<!---
<img style="float:left; width:450px" src="https://upload.wikimedia.org/wikipedia/commons/9/9f/Iris_virginica.jpg",width=300,height=50>
-->
<img style="float:left; width:450px" src="http://www.twofrog.com/images/iris38a.jpg",width=300,height=50>
<!---
<img style="float:right; width:490px" src="https://upload.wikimedia.org/wikipedia/commons/4/41/Iris_versicolor_3.jpg",width=300,height=50>
-->
<img style="float:right; width:490px" src="http://blazingstargardens.com/wp-content/uploads/2016/02/Iris-versicolor-Blue-Flag-Iris1.jpg",width=300,height=62>
The two features of our model will be the **sepal length** and **sepal width**. Execute the following cell to see a plot of the data. The blue points correspond to the sepal measurements of the Iris Setosa (left) and the red points correspond to the sepal measurements of the Iris Versicolour (right).
```
import matplotlib.pyplot as plt
%matplotlib inline
from sklearn import datasets
iris = datasets.load_iris()
X_train = iris.data[iris.target != 2, :2] # first two features and
y_train = iris.target[iris.target != 2] # first two labels only
fig = plt.figure(figsize=(8,8))
mycolors = {"blue": "steelblue", "red": "#a76c6e", "green": "#6a9373"}
plt.scatter(X_train[:, 0], X_train[:, 1], s=100, alpha=0.9, c=[mycolors["red"] if yi==1 else mycolors["blue"] for yi in y_train])
plt.xlabel('sepal length', fontsize=16)
plt.ylabel('sepal width', fontsize=16);
```
We'll train a logistic regression model of the form
$$
p(y = 1 ~|~ {\bf x}; {\bf w}) = \frac{1}{1 + \textrm{exp}[-(w_0 + w_1x_1 + w_2x_2)]}
$$
using **sklearn**'s logistic regression classifier as follows
```
from sklearn.linear_model import LogisticRegression # import from sklearn
logreg = LogisticRegression() # initialize classifier
logreg.fit(X_train, y_train); # train on training data
```
**Q**: Determine the parameters ${\bf w}$ fit by the model. It might be helpful to consult the documentation for the classifier on the <a href="http://scikit-learn.org/stable/modules/generated/sklearn.linear_model.LogisticRegression.html">sklearn website</a>. **Hint**: The classifier stores the coefficients and bias term separately.
**Q**: In general, what does the Logistic Regression decision boundary look like for data with two features?
**Q**: Modify the code below to plot the decision boundary along with the data.
```
import numpy as np
fig = plt.figure(figsize=(8,8))
plt.scatter(X_train[:, 0], X_train[:, 1], s=100, c=[mycolors["red"] if yi==1 else mycolors["blue"] for yi in y_train])
plt.xlabel('Sepal length')
plt.ylabel('Sepal width')
x_min, x_max = np.min(X_train[:,0])-0.1, np.max(X_train[:,0])+0.1
y_min, y_max = np.min(X_train[:,1])-0.1, np.max(X_train[:,1])+0.1
plt.xlim(x_min, x_max)
plt.ylim(y_min, y_max)
x1 = np.linspace(x_min, x_max, 100)
w0 = logreg.intercept_
w1 = logreg.coef_[0][0]
w2 = logreg.coef_[0][1]
x2 = #TODO
plt.plot(x1, x2, color="gray");
```
### Problem 2: The Bag-of-Words Text Model
***
The remainder of today's exercise will consider the problem of predicting the semantics of text. In particular, later we'll look at predicting whether movie reviews are positive or negative just based on their text.
Before we can utilize text as features in a learning model, we need a concise mathematical way to represent things like words, phrases, sentences, etc. The most common text models are based on the so-called <a href="https://en.wikipedia.org/wiki/Vector_space_model">Vector Space Model</a> (VSM) where individual words in a document are associated with entries of a vector:
$$
\textrm{"The sky is blue"} \quad \Rightarrow \quad
\left[
\begin{array}{c}
0 \\
1 \\
0 \\
0 \\
1
\end{array}
\right]
$$
The first step in creating a VSM is to define a vocabulary, $V$, of words that you will include in your model. This vocabulary can be determined by looking at all (or most) of the words in the training set, or even by including a fixed vocabulary based on the english language. A vector representation of a document like a movie review is then a vector with length $|V|$ where each entry in the vector maps uniquely to a word in the vocabulary. A vector encoding of a document would then be a vector that is nonzero in positions corresponding to words present in the document and zero everywhere else. How you fill in the nonzero entries depends on the model you're using. Two simple conventions are the **Bag-of-Words** model and the **binary** model.
In the binary model we simply set an entry of the vector to $1$ if the associate word appears at least once in the document. In the more common Bag-of-Words model we set an entry of the vector equal to the frequency with which the word appears in the document. Let's see if we can come up with a simple implementation of the Bag-of-Words model in Python, and then later we'll see how sklearn can do the heavy lifting for us.
Consider a training set containing three documents, specified as follows
$\texttt{Training Set}:$
$\texttt{d1}: \texttt{new york times}$
$\texttt{d2}: \texttt{new york post}$
$\texttt{d3}: \texttt{los angeles times}$
First we'll define the vocabulary based on the words in the test set. It is $V = \{ \texttt{angeles}, \texttt{los}, \texttt{new}, \texttt{post}, \texttt{times}, \texttt{york}\}$.
We need to define an association between the particular words in the vocabulary and the specific entries in our vectors. Let's define this association in the order that we've listed them above. We can store this mapping as a Python dictionary as follows:
```
V = {"angeles": 0, "los": 1, "new": 2, "post": 3, "times": 4, "york": 5}
```
Let's also store the documents in a list as follows:
```
D = ["new york times", "new york post", "los angeles times"]
```
To be consistent with sklearn conventions, we'll encode the documents as *row-vectors* stored in a matrix. In this case, each row of the matrix corresponds to a document, and each column corresponds to a term in the vocabulary. For our example this gives us a matrix $M$ of shape $3 \times 6$. The $(d,t)$-entry in $M$ is then the number of times the term $t$ appears in document $d$
**Q**: Your first task is to write some simple Python code to construct the *term-frequency* matrix $M$
```
M = np.zeros((len(D),len(V)))
for ii, doc in enumerate(D):
for term in doc.split():
#TODO
print M
```
Hopefully your code returns the matrix
$$M =
\left[
\begin{array}{ccccccc}
0 & 0 & 1 & 0 & 1 & 1 \\
0 & 0 & 1 & 1 & 0 & 1 \\
1 & 1 & 0 & 0 & 1 & 0 \\
\end{array}
\right]$$.
Note that the entry in the (2,0) position is $1$ because the first word (angeles) appears once in the third document.
OK, let's see how we can construct the same term-frequency matrix in sklearn. We will use something called the <a href="http://scikit-learn.org/stable/modules/generated/sklearn.feature_extraction.text.CountVectorizer.html">CountVectorizer</a> to accomplish this. Let's see some code and then we'll explain how it functions.
```
from sklearn.feature_extraction.text import CountVectorizer # import CountVectorizer
vectorizer = CountVectorizer() # initialize the vectorizer
X = vectorizer.fit_transform(D) # fit to training data and transform to matrix
```
The $\texttt{fit_transform}$ method actually does two things. It fits the model to the training data by building a vocabulary. It then transforms the text in $D$ into matrix form.
If we wish to see the vocabulary you can do it like so
```
print vectorizer.vocabulary_
```
Note that this is the same vocabulary and indexing that we definfed ourselves. Hopefully that means we'll get the same term-frequency matrix. We can print $X$ and check
```
print X.todense()
```
Yep, they're the same! Notice that we had to convert $X$ to a dense matrix for printing. This is because CountVectorizer actually returns a sparse matrix. This is a very good thing since most vectors in a text model will be **extremely** sparse, since most documents will only contain a handful of words from the vocabulary.
OK, let's see how we can use the CountVectorizer to transform the test documents into their own term-frequency matrix. Removing the stop words we have
OK, now suppose that we have a query document not included in the training set that we want to vectorize.
```
d4 = ["new york new tribune"]
```
We've already fit the CountVectorizer to the training set, so all we need to do is transform the test set documents into a term-frequency vector using the same conventions. Since we've already fit the model, we do the transformation with the $\texttt{transform}$ method:
```
x4 = vectorizer.transform(d4)
```
Let's print it and see what it looks like
```
print x4.todense()
```
Notice that the query document included the word $\texttt{new}$ twice, which corresponds to the entry in the $(0,2)$-position.
**Q**: What's missing from $x4$ that we might expect to see from the query document?
<br>
### Problem 3: Term Frequency - Inverse Document Frequency
***
The Bag-of-Words model for text classification is very popular, but let's see if we can do better. Currently we're weighting every word in the corpus by it's frequency. It turns out that in text classification there are often features that are not particularly useful predictors for the document class, either because they are too common or too uncommon. **Stop-words** are extremely common, low-information words like "a", "the", "as", etc. Removing these from documents is typically the first thing done in peparing data for document classification.
**Q**: Can you think of a situation where it might be useful to keep stop words in the corpus?
Other words that tend to be uninformative predictors are words that appear very very rarely. In particular, if they do not appear frequently enough in the training data then it is difficult for a classification algorithm to weight them heavily in the classification process.
In general, the words that tend to be useful predictors are the words that appear frequently, but not too frequently. Consider the following frequency graph for a corpus.
<img src="figs/feat_freq.png",width=400,height=50>
The features in column A appear too frequently to be very useful, and the features in column C appear too rarely. One first-pass method of feature selection in text classification would be to discard the words from columns A and C, and build a classifier with only features from column B.
Another common model for identifying the useful terms in a document is the Term Frequency - Inverse Document Frequency (tf-idf) model. Here we won't throw away any terms, but we'll replace their Bag-of-Words frequency counts with tf-idf scores which we describe below.
The tf-idf score is the product of two statistics, *term frequency* and *inverse document frequency*
$$\texttt{tfidf(d,t)} = \texttt{tf(d,t)} \times \texttt{idf(t)}$$
The term frequency $\texttt{tf(d,t)}$ is a measure of the frequency with which term $t$ appears in document $d$. The inverse document frequency $\texttt{idf(t)}$ is a measure of how much information the word provides, that is, whether the term is common or rare across all documents. By multiplying the two quantities together, we obtain a representation of term $t$ in document $d$ that weighs how common the term is in the document with how common the word is in the entire corpus. You can imagine that the words that get the highest associated values are terms that appear many times in a small number of documents.
There are many ways to compute the composite terms $\texttt{tf}$ and $\texttt{idf}$. For simplicity, we'll define $\texttt{tf(d,t)}$ to be the number of times term $t$ appears in document $d$ (i.e., Bag-of-Words). We will define the inverse document frequency as follows:
$$
\texttt{idf(t)} = \ln ~ \frac{\textrm{total # documents}}{\textrm{1 + # documents with term }t}
= \ln ~ \frac{|D|}{|d: ~ t \in d |}
$$
Note that we could have a potential problem if a term comes up that is not in any of the training documents, resulting in a divide by zero. This might happen if you use a canned vocabulary instead of constructing one from the training documents. To guard against this, many implementations will use add-one smoothing in the denominator (this is what sklearn does).
$$
\texttt{idf(t)} = \ln ~ \frac{\textrm{total # documents}}{\textrm{1 + # documents with term }t}
= \ln ~ \frac{|D|}{1 + |d: ~ t \in d |}
$$
**Q**: Compute $\texttt{idf(t)}$ (without smoothing) for each of the terms in the training documents from the previous problem
**Q**: Compute the td-ifd matrix for the training set
Hopefully you got something like the following:
$$
X_{tfidf} =
\left[
\begin{array}{ccccccccc}
0. & 0. & 0.40546511 & 0. & 0.40546511 & 0.40546511 \\
0. & 0. & 0.40546511 & 1.09861229 & 0. & 0.40546511 \\
1.09861229 & 1.09861229 & 0. & 0. & 0.40546511 & 0.
\end{array}
\right]
$$
The final step in any VSM method is the normalization of the vectors. This is done so that very long documents to not completely overpower the small and medium length documents.
```
row_norms = np.array([np.linalg.norm(row) for row in Xtfidf])
X_tfidf_n = np.dot(np.diag(1./row_norms), Xtfidf)
print X_tfidf_n
```
Let's see what we get when we use sklearn. Sklearn has a vectorizer called TfidfVectorizer which is similar to CountVectorizer, but it computes tf-idf scores.
```
from sklearn.feature_extraction.text import TfidfVectorizer
tfidf = TfidfVectorizer()
Y = tfidf.fit_transform(D)
print Y.todense()
```
Note that these are not quite the same, becuase sklearn's implementation of tf-idf uses the add-one smoothing in the denominator for idf.
<br>
### Problem 4: Classifying Semantics in Movie Reviews
***
> The data for this problem was taken from the <a href="https://www.kaggle.com/c/word2vec-nlp-tutorial/details/part-1-for-beginners-bag-of-words">Bag of Words Meets Bag of Popcorn</a> Kaggle competition
In this problem you will use the text from movie reviews to predict whether the reviewer felt positively or negatively about the movie using Bag-of-Words and tf-idf. I've partially cleaned the data and stored it in files called $\texttt{labeledTrainData.tsv}$ and $\texttt{labeledTestData.tsv}$ in the data directory.
```
import csv
def read_and_clean_data(fname, remove_stops=True):
with open('data/stopwords.txt', 'rt') as f:
stops = [line.rstrip('\n') for line in f]
with open(fname,'rt') as tsvin:
reader = csv.reader(tsvin, delimiter='\t')
labels = []; text = []
for ii, row in enumerate(reader):
labels.append(int(row[0]))
words = row[1].lower().split()
words = [w for w in words if not w in stops] if remove_stops else words
text.append(" ".join(words))
return text, labels
text_train, labels_train = read_and_clean_data('data/labeledTrainData.tsv', remove_stops=True)
text_test, labels_test = read_and_clean_data('data/labeledTestData.tsv', remove_stops=True)
```
The current parameters are set to not remove stop words from the text so that it's a bit easier to explore.
Look at a few of the reviews stored in $\texttt{text_train}$ as well as their associated labels in $\texttt{labels_train}$. Can you figure out which label refers to a positive review and which refers to a negative review?
```
labels_train[:4]
```
The first review is labeled $1$ and has the following text:
```
text_train[3]
```
The fourth review is labeled $0$ and has the following text:
```
text_train[0]
```
Hopefully it's obvious that label 1 corresponds to positive reviews and label 0 to negative reviews!
OK, the first thing we'll do is train a logistic regression classifier using the Bag-of-Words model, and see what kind of accuracy we can get. To get started, we need to vectorize the text into mathematical features that we can use. We'll use CountVectorizer to do the job. (Before starting, I'm going to reload the data and remove the stop words this time)
```
text_train, labels_train = read_and_clean_data('data/labeledTrainData.tsv', remove_stops=True)
text_test, labels_test = read_and_clean_data('data/labeledTestData.tsv', remove_stops=True)
cvec = CountVectorizer()
X_bw_train = cvec.fit_transform(text_train)
y_train = np.array(labels_train)
X_bw_test = cvec.transform(text_test)
y_test = np.array(labels_test)
```
**Q**: How many different words are in the vocabulary?
OK, now we'll train a logistic regression classifier on the training set, and test the accuracy on the test set. To do this we'll need to load some kind of accuracy metric from sklearn.
```
from sklearn.metrics import accuracy_score
bwLR = LogisticRegression()
bwLR.fit(X_bw_train, y_train)
pred_bwLR = bwLR.predict(X_bw_test)
print "Logistic Regression accuracy with Bag-of-Words: ", accuracy_score(y_test, pred_bwLR)
```
OK, so we got an accuracy of around 81% using Bag-of-Words. Now lets do the same tests but this time with tf-idf features.
```
tvec = TfidfVectorizer()
X_tf_train = tvec.fit_transform(text_train)
X_tf_test = tvec.transform(text_test)
tfLR = LogisticRegression()
tfLR.fit(X_tf_train, y_train)
pred_tfLR = tfLR.predict(X_tf_test)
print "Logistic Regression accuracy with tf-idf: ", accuracy_score(y_test, pred_tfLR)
```
**WOOHOO**! With tf-idf features we got around 85% accuracy, which is a 4% improvement. (If you're scoffing at this, wait until you get some more experience working with real-world data. 4% improvement is pretty awesome).
**Q**: Which words are the strongest predictors for a positive review and which words are the strongest predictors for negative reviews? I'm not going to give you the answer to this one because it's the same question we'll ask on the next homework assignment. But if you figure this out you'll have a great head start!
<br><br><br>
<br><br><br>
<br><br><br>
<br><br><br>
<br><br><br>
<br><br><br>
<br><br><br>
<br><br><br>
<br><br><br>
<br><br><br>
<br><br><br>
<br><br><br>
<br><br><br>
<br><br><br>
<br><br><br>
<br><br><br>
## Notebook Solutions
***
<br><br><br>
### Problem 1: Logistic Regression for 2D Continuous Features
***
In the video lecture you saw some examples of using logistic regression to do binary classification on text data (SPAM vs HAM) and on 1D continuous data. In this problem we'll look at logistic regression for 2D continuous data. The data we'll use are <a href="https://www.math.umd.edu/~petersd/666/html/iris_with_labels.jpg">sepal</a> measurements from the ubiquitous *iris* dataset.
<!---
<img style="float:left; width:450px" src="https://upload.wikimedia.org/wikipedia/commons/9/9f/Iris_virginica.jpg",width=300,height=50>
-->
<img style="float:left; width:450px" src="http://www.twofrog.com/images/iris38a.jpg",width=300,height=50>
<!---
<img style="float:right; width:490px" src="https://upload.wikimedia.org/wikipedia/commons/4/41/Iris_versicolor_3.jpg",width=300,height=50>
-->
<img style="float:right; width:490px" src="http://blazingstargardens.com/wp-content/uploads/2016/02/Iris-versicolor-Blue-Flag-Iris1.jpg",width=300,height=62>
The two features of our model will be the **sepal length** and **sepal width**. Execute the following cell to see a plot of the data. The blue points correspond to the sepal measurements of the Iris Setosa (left) and the red points correspond to the sepal measurements of the Iris Versicolour (right).
```
import matplotlib.pyplot as plt
%matplotlib inline
from sklearn import datasets
iris = datasets.load_iris()
X_train = iris.data[iris.target != 2, :2] # first two features and
y_train = iris.target[iris.target != 2] # first two labels only
fig = plt.figure(figsize=(8,8))
mycolors = {"blue": "steelblue", "red": "#a76c6e", "green": "#6a9373"}
plt.scatter(X_train[:, 0], X_train[:, 1], s=100, alpha=0.9, c=[mycolors["red"] if yi==1 else mycolors["blue"] for yi in y_train])
plt.xlabel('sepal length', fontsize=16)
plt.ylabel('sepal width', fontsize=16);
```
We'll train a logistic regression model of the form
$$
p(y = 1 ~|~ {\bf x}; {\bf w}) = \frac{1}{1 + \textrm{exp}[-(w_0 + w_1x_1 + w_2x_2)]}
$$
using **sklearn**'s logistic regression classifier as follows
```
from sklearn.linear_model import LogisticRegression # import from sklearn
logreg = LogisticRegression() # initialize classifier
logreg.fit(X_train, y_train); # train on training data
```
**Q**: Determine the parameters ${\bf w}$ fit by the model. It might be helpful to consult the documentation for the classifier on the <a href="http://scikit-learn.org/stable/modules/generated/sklearn.linear_model.LogisticRegression.html">sklearn website</a>. **Hint**: The classifier stores the coefficients and bias term separately.
**A**: The bias term is stored in logreg.intercept\_ . The remaining coefficients are stored in logreg.coef\_ . For this problem we have
$$
w_0 =-0.599, \quad w_1 = 2.217, \quad \textrm{and} \quad w_2 = -3.692
$$
**Q**: In general, what does the Logistic Regression decision boundary look like for data with two features?
**A**: The decision boundary for Logistic Regresion for data with two features is a line. To see this, remember that the decision boundary is made up of $(x_1, x_2)$ points such that $\textrm{sigm}({\bf w}^T{\bf x}) = 0.5$. We then have
$$
\frac{1}{1 + \textrm{exp}[-(w_0 + w_1x_1 + w_2x_2)]} = \frac{1}{2} ~~\Rightarrow ~~ w_0 + w_1x_1 + w_2x_2 = 0 ~~\Rightarrow~~ x_2 = -\frac{w_1}{w_2}x_1 - \frac{w_0}{w_2}
$$
So the decision boundary is a line with slope $-w_1/w_2$ and intercept $-w_0/w_2$.
**Q**: Modify the code below to plot the decision boundary along with the data.
```
import numpy as np
fig = plt.figure(figsize=(8,8))
plt.scatter(X_train[:, 0], X_train[:, 1], s=100, c=[mycolors["red"] if yi==1 else mycolors["blue"] for yi in y_train])
plt.xlabel('Sepal length')
plt.ylabel('Sepal width')
x_min, x_max = np.min(X_train[:,0])-0.1, np.max(X_train[:,0])+0.1
y_min, y_max = np.min(X_train[:,1])-0.1, np.max(X_train[:,1])+0.1
plt.xlim(x_min, x_max)
plt.ylim(y_min, y_max)
x1 = np.linspace(x_min, x_max, 100)
w0 = logreg.intercept_
w1 = logreg.coef_[0][0]
w2 = logreg.coef_[0][1]
x2 = -(w0/w2) - (w1/w2)*x1 #TODO
plt.plot(x1, x2, color="gray");
```
### Problem 2: The Bag-of-Words Text Model
***
The remainder of today's exercise will consider the problem of predicting the semantics of text. In particular, later we'll look at predicting whether movie reviews are positive or negative just based on their text.
Before we can utilize text as features in a learning model, we need a concise mathematical way to represent things like words, phrases, sentences, etc. The most common text models are based on the so-called <a href="https://en.wikipedia.org/wiki/Vector_space_model">Vector Space Model</a> (VSM) where individual words in a document are associated with entries of a vector:
$$
\textrm{"The sky is blue"} \quad \Rightarrow \quad
\left[
\begin{array}{c}
0 \\
1 \\
0 \\
0 \\
1
\end{array}
\right]
$$
The first step in creating a VSM is to define a vocabulary, $V$, of words that you will include in your model. This vocabulary can be determined by looking at all (or most) of the words in the training set, or even by including a fixed vocabulary based on the english language. A vector representation of a document like a movie review is then a vector with length $|V|$ where each entry in the vector maps uniquely to a word in the vocabulary. A vector encoding of a document would then be a vector that is nonzero in positions corresponding to words present in the document and zero everywhere else. How you fill in the nonzero entries depends on the model you're using. Two simple conventions are the **Bag-of-Words** model and the **binary** model.
In the binary model we simply set an entry of the vector to $1$ if the associate word appears at least once in the document. In the more common Bag-of-Words model we set an entry of the vector equal to the frequency with which the word appears in the document. Let's see if we can come up with a simple implementation of the Bag-of-Words model in Python, and then later we'll see how sklearn can do the heavy lifting for us.
Consider a training set containing three documents, specified as follows
$\texttt{Training Set}:$
$\texttt{d1}: \texttt{new york times}$
$\texttt{d2}: \texttt{new york post}$
$\texttt{d3}: \texttt{los angeles times}$
First we'll define the vocabulary based on the words in the test set. It is $V = \{ \texttt{angeles}, \texttt{los}, \texttt{new}, \texttt{post}, \texttt{times}, \texttt{york}\}$.
We need to define an association between the particular words in the vocabulary and the specific entries in our vectors. Let's define this association in the order that we've listed them above. We can store this mapping as a Python dictionary as follows:
```
V = {"angeles": 0, "los": 1, "new": 2, "post": 3, "times": 4, "york": 5}
```
Let's also store the documents in a list as follows:
```
D = ["new york times", "new york post", "los angeles times"]
```
To be consistent with sklearn conventions, we'll encode the documents as *row-vectors* stored in a matrix. In this case, each row of the matrix corresponds to a document, and each column corresponds to a term in the vocabulary. For our example this gives us a matrix $M$ of shape $3 \times 6$. The $(d,t)$-entry in $M$ is then the number of times the term $t$ appears in document $d$
**Q**: Your first task is to write some simple Python code to construct the *term-frequency* matrix $M$
```
M = np.zeros((len(D),len(V)))
for ii, doc in enumerate(D):
for term in doc.split():
M[ii, V[term]] += 1
print M
```
Hopefully your code returns the matrix
$$M =
\left[
\begin{array}{ccccccc}
0 & 0 & 1 & 0 & 1 & 1 \\
0 & 0 & 1 & 1 & 0 & 1 \\
1 & 1 & 0 & 0 & 1 & 0 \\
\end{array}
\right]$$.
Note that the entry in the (2,0) position is $1$ because the first word (angeles) appears once in the third document.
OK, let's see how we can construct the same term-frequency matrix in sklearn. We will use something called the <a href="http://scikit-learn.org/stable/modules/generated/sklearn.feature_extraction.text.CountVectorizer.html">CountVectorizer</a> to accomplish this. Let's see some code and then we'll explain how it functions.
```
from sklearn.feature_extraction.text import CountVectorizer # import CountVectorizer
vectorizer = CountVectorizer() # initialize the vectorizer
X = vectorizer.fit_transform(D) # fit to training data and transform to matrix
```
The $\texttt{fit_transform}$ method actually does two things. It fits the model to the training data by building a vocabulary. It then transforms the text in $D$ into matrix form.
If we wish to see the vocabulary you can do it like so
```
print vectorizer.vocabulary_
```
Note that this is the same vocabulary and indexing that we definfed ourselves. Hopefully that means we'll get the same term-frequency matrix. We can print $X$ and check
```
print X.todense()
```
Yep, they're the same! Notice that we had to convert $X$ to a dense matrix for printing. This is because CountVectorizer actually returns a sparse matrix. This is a very good thing since most vectors in a text model will be **extremely** sparse, since most documents will only contain a handful of words from the vocabulary.
OK, now suppose that we have a query document not included in the training set that we want to vectorize.
```
d4 = ["new york new tribune"]
```
We've already fit the CountVectorizer to the training set, so all we need to do is transform the test set documents into a term-frequency vector using the same conventions. Since we've already fit the model, we do the transformation with the $\texttt{transform}$ method:
```
x4 = vectorizer.transform(d4)
```
Let's print it and see what it looks like
```
print x4.todense()
```
Notice that the query document included the word $\texttt{new}$ twice, which corresponds to the entry in the $(0,2)$-position.
**Q**: What's missing from $x4$ that we might expect to see from the query document?
**A**: The word $\texttt{tribune}$ do not appear in vector $x4$ at all. This is because it did not occur in the training set, which means it is not present in the VSM vocabulary. This should not bother us too much. Most reasonable text data sets will have most of the important words present in the training set and thus in the vocabulary. On the other hand, the throw-away words that are present only in the test set are probably useless anyway, since the learning model is trained based on the text in the training set, and thus won't be able to do anything intelligent with words the model hasn't seen yet.
<br>
### Problem 3: Term Frequency - Inverse Document Frequency
***
The Bag-of-Words model for text classification is very popular, but let's see if we can do better. Currently we're weighting every word in the corpus by it's frequency. It turns out that in text classification there are often features that are not particularly useful predictors for the document class, either because they are too common or too uncommon. **Stop-words** are extremely common, low-information words like "a", "the", "as", etc. Removing these from documents is typically the first thing done in peparing data for document classification.
**Q**: Can you think of a situation where it might be useful to keep stop words in the corpus?
**A**: If you plan to use bi-grams or tri-grams as features. Bi-grams are pairs of words that appear side-by-side in a document, e.g. "he went", "went to", "to the", "the store".
Other words that tend to be uninformative predictors are words that appear very very rarely. In particular, if they do not appear frequently enough in the training data then it is difficult for a classification algorithm to weight them heavily in the classification process.
In general, the words that tend to be useful predictors are the words that appear frequently, but not too frequently. Consider the following frequency graph for a corpus.
<img src="figs/feat_freq.png",width=400,height=50>
The features in column A appear too frequently to be very useful, and the features in column C appear too rarely. One first-pass method of feature selection in text classification would be to discard the words from columns A and C, and build a classifier with only features from column B.
Another common model for identifying the useful terms in a document is the Term Frequency - Inverse Document Frequency (tf-idf) model. Here we won't throw away any terms, but we'll replace their Bag-of-Words frequency counts with tf-idf scores which we describe below.
The tf-idf score is the product of two statistics, *term frequency* and *inverse document frequency*
$$\texttt{tfidf(d,t)} = \texttt{tf(d,t)} \times \texttt{idf(t)}$$
The term frequency $\texttt{tf(d,t)}$ is a measure of the frequency with which term $t$ appears in document $d$. The inverse document frequency $\texttt{idf(t)}$ is a measure of how much information the word provides, that is, whether the term is common or rare across all documents. By multiplying the two quantities together, we obtain a representation of term $t$ in document $d$ that weighs how common the term is in the document with how common the word is in the entire corpus. You can imagine that the words that get the highest associated values are terms that appear many times in a small number of documents.
There are many ways to compute the composite terms $\texttt{tf}$ and $\texttt{idf}$. For simplicity, we'll define $\texttt{tf(d,t)}$ to be the number of times term $t$ appears in document $d$ (i.e., Bag-of-Words). We will define the inverse document frequency as follows:
$$
\texttt{idf(t)} = \ln ~ \frac{\textrm{total # documents}}{\textrm{# documents with term }t}
= \ln ~ \frac{|D|}{|d: ~ t \in d |}
$$
Note that we could have a potential problem if a term comes up that is not in any of the training documents, resulting in a divide by zero. This might happen if you use a canned vocabulary instead of constructing one from the training documents. To guard against this, many implementations will use add-one smoothing in the denominator (this is what sklearn does).
$$
\texttt{idf(t)} = \ln ~ \frac{\textrm{total # documents}}{\textrm{1 + # documents with term }t}
= \ln ~ \frac{|D|}{1 + |d: ~ t \in d |}
$$
**Q**: Compute $\texttt{idf(t)}$ (without smoothing) for each of the terms in the training documents from the previous problem
**A**:
$
\texttt{idf}(\texttt{angeles}) = \ln ~ \frac{3}{1} = \ln ~ \frac{3}{1} = 1.10
$
$
\texttt{idf}(\texttt{los}) = \ln ~ \frac{3}{1} = \ln ~ \frac{3}{1} = 1.10
$
$
\texttt{idf}(\texttt{new}) = \ln ~ \frac{3}{2} = \ln ~ \frac{3}{2} = 0.41
$
$
\texttt{idf}(\texttt{post}) = \ln ~ \frac{3}{1} = \ln ~ \frac{3}{1} = 1.10
$
$
\texttt{idf}(\texttt{times}) = \ln ~ \frac{3}{2} = \ln ~ \frac{3}{2} = 0.41
$
$
\texttt{idf}(\texttt{york}) = \ln ~ \frac{3}{2} = \ln ~ \frac{3}{2} = 0.41
$
**Q**: Compute the td-ifd matrix for the training set
**A**: There are several ways to do this. One way would be to multiply the term-frequency matrix on the right with a diagonal matrix with the idf-values on the main diagonal
```
idf = np.array([np.log(3), np.log(3), np.log(3./2), np.log(3), np.log(3./2), np.log(3./2)])
Xtfidf = np.dot(X.todense(), np.diag(idf))
print Xtfidf
```
Hopefully you got something like the following:
$$
X_{tfidf} =
\left[
\begin{array}{ccccccccc}
0. & 0. & 0.40546511 & 0. & 0.40546511 & 0.40546511 \\
0. & 0. & 0.40546511 & 1.09861229 & 0. & 0.40546511 \\
1.09861229 & 1.09861229 & 0. & 0. & 0.40546511 & 0.
\end{array}
\right]
$$
The final step in any VSM method is the normalization of the vectors. This is done so that very long documents to not completely overpower the small and medium length documents.
```
row_norms = np.array([np.linalg.norm(row) for row in Xtfidf])
X_tfidf_n = np.dot(np.diag(1./row_norms), Xtfidf)
print X_tfidf_n
```
Let's see what we get when we use sklearn. Sklearn has a vectorizer called TfidfVectorizer which is similar to CountVectorizer, but it computes tf-idf scores.
```
from sklearn.feature_extraction.text import TfidfVectorizer
tfidf = TfidfVectorizer()
Y = tfidf.fit_transform(D)
print Y.todense()
```
Note that these are not quite the same, becuase sklearn's implementation of tf-idf uses the add-one smoothing in the denominator for idf.
<br>
### Problem 4: Classifying Semantics in Movie Reviews
***
> The data for this problem was taken from the <a href="https://www.kaggle.com/c/word2vec-nlp-tutorial/details/part-1-for-beginners-bag-of-words">Bag of Words Meets Bag of Popcorn</a> Kaggle competition
In this problem you will use the text from movie reviews to predict whether the reviewer felt positively or negatively about the movie using Bag-of-Words and tf-idf. I've partially cleaned the data and stored it in files called $\texttt{labeledTrainData.tsv}$ and $\texttt{labeledTestData.tsv}$ in the data directory.
```
import csv
def read_and_clean_data(fname, remove_stops=True):
with open('data/stopwords.txt', 'r') as f:
stops = [line.rstrip('\n') for line in f]
with open(fname,'rb') as tsvin:
reader = csv.reader(tsvin, delimiter='\t')
labels = []; text = []
for ii, row in enumerate(reader):
labels.append(int(row[0]))
words = row[1].lower().split()
words = [w for w in words if not w in stops] if remove_stops else words
text.append(" ".join(words))
return text, labels
text_train, labels_train = read_and_clean_data('data/labeledTrainData.tsv', remove_stops=False)
text_test, labels_test = read_and_clean_data('data/labeledTestData.tsv', remove_stops=False)
```
The current parameters are set to not remove stop words from the text so that it's a bit easier to explore.
**Q**: Look at a few of the reviews stored in $\texttt{text_train}$ as well as their associated labels in $\texttt{labels_train}$. Can you figure out which label refers to a positive review and which refers to a negative review?
**A**:
```
labels_train[:4]
```
The first review is labeled $1$ and has the following text:
```
text_train[0]
```
The fourth review is labeled $0$ and has the following text:
```
text_train[3]
```
Hopefully it's obvious that label 1 corresponds to positive reviews and label 0 to negative reviews!
OK, the first thing we'll do is train a logistic regression classifier using the Bag-of-Words model, and see what kind of accuracy we can get. To get started, we need to vectorize the text into mathematical features that we can use. We'll use CountVectorizer to do the job. (Before starting, I'm going to reload the data and remove the stop words this time)
```
text_train, labels_train = read_and_clean_data('data/labeledTrainData.tsv', remove_stops=True)
text_test, labels_test = read_and_clean_data('data/labeledTestData.tsv', remove_stops=True)
cvec = CountVectorizer()
X_bw_train = cvec.fit_transform(text_train)
y_train = np.array(labels_train)
X_bw_test = cvec.transform(text_test)
y_test = np.array(labels_test)
```
**Q**: How many different words are in the vocabulary?
```
X_bw_train.shape
```
**A**: It looks like around 17,800 distinct words
OK, now we'll train a logistic regression classifier on the training set, and test the accuracy on the test set. To do this we'll need to load some kind of accuracy metric from sklearn.
```
from sklearn.metrics import accuracy_score
bwLR = LogisticRegression()
bwLR.fit(X_bw_train, y_train)
pred_bwLR = bwLR.predict(X_bw_test)
print "Logistic Regression accuracy with Bag-of-Words: ", accuracy_score(y_test, pred_bwLR)
```
OK, so we got an accuracy of around 81% using Bag-of-Words. Now lets do the same tests but this time with tf-idf features.
```
tvec = TfidfVectorizer()
X_tf_train = tvec.fit_transform(text_train)
X_tf_test = tvec.transform(text_test)
tfLR = LogisticRegression()
tfLR.fit(X_tf_train, y_train)
pred_tfLR = tfLR.predict(X_tf_test)
print "Logistic Regression accuracy with tf-idf: ", accuracy_score(y_test, pred_tfLR)
```
**WOOHOO**! With tf-idf features we got around 85% accuracy, which is a 4% improvement. (If you're scoffing at this, wait until you get some more experience working with real-world data. 4% improvement is pretty awesome).
**Q**: Which words are the strongest predictors for a positive review and which words are the strongest predictors for negative reviews? I'm not going to give you the answer to this one because it's the same question we'll ask on the next homework assignment. But if you figure this out you'll have a great head start!
<br><br><br>
<br><br><br>
<br><br><br>
<br><br><br>
<br><br><br>
```
from IPython.core.display import HTML
HTML("""
<style>
.MathJax nobr>span.math>span{border-left-width:0 !important};
</style>
""")
```
| github_jupyter |
# ML with TensorFlow Extended (TFX) -- Part 1
The puprpose of this tutorial is to show how to do end-to-end ML with TFX libraries on Google Cloud Platform. This tutorial covers:
1. Data analysis and schema generation with **TF Data Validation**.
2. Data preprocessing with **TF Transform**.
3. Model training with **TF Estimator**.
4. Model evaluation with **TF Model Analysis**.
This notebook has been tested in Jupyter on the Deep Learning VM.
## Setup Cloud environment
```
import tensorflow as tf
import tensorflow_data_validation as tfdv
print('TF version: {}'.format(tf.__version__))
print('TFDV version: {}'.format(tfdv.__version__))
PROJECT = 'cloud-training-demos' # Replace with your PROJECT
BUCKET = 'cloud-training-demos-ml' # Replace with your BUCKET
REGION = 'us-central1' # Choose an available region for Cloud MLE
import os
os.environ['PROJECT'] = PROJECT
os.environ['BUCKET'] = BUCKET
os.environ['REGION'] = REGION
%%bash
gcloud config set project $PROJECT
gcloud config set compute/region $REGION
## ensure we predict locally with our current Python environment
gcloud config set ml_engine/local_python `which python`
```
<img valign="middle" src="images/tfx.jpeg">
### UCI Adult Dataset: https://archive.ics.uci.edu/ml/datasets/adult
Predict whether income exceeds $50K/yr based on census data. Also known as "Census Income" dataset.
```
DATA_DIR='gs://cloud-samples-data/ml-engine/census/data'
import os
TRAIN_DATA_FILE = os.path.join(DATA_DIR, 'adult.data.csv')
EVAL_DATA_FILE = os.path.join(DATA_DIR, 'adult.test.csv')
!gsutil ls -l $TRAIN_DATA_FILE
!gsutil ls -l $EVAL_DATA_FILE
```
## 1. Data Analysis
For data analysis, visualization, and schema generation, we use [TensorFlow Data Validation](https://www.tensorflow.org/tfx/guide/tfdv) to perform the following:
1. **Analyze** the training data and produce **statistics**.
2. Generate data **schema** from the produced statistics.
3. **Configure** the schema.
4. **Validate** the evaluation data against the schema.
5. **Save** the schema for later use.
### 1.1 Compute and visualise statistics
```
HEADER = ['age', 'workclass', 'fnlwgt', 'education', 'education_num',
'marital_status', 'occupation', 'relationship', 'race', 'gender',
'capital_gain', 'capital_loss', 'hours_per_week',
'native_country', 'income_bracket']
TARGET_FEATURE_NAME = 'income_bracket'
TARGET_LABELS = [' <=50K', ' >50K']
WEIGHT_COLUMN_NAME = 'fnlwgt'
# This is a convenience function for CSV. We can write a Beam pipeline for other formats.
# https://www.tensorflow.org/tfx/data_validation/api_docs/python/tfdv/generate_statistics_from_csv
train_stats = tfdv.generate_statistics_from_csv(
data_location=TRAIN_DATA_FILE,
column_names=HEADER,
stats_options=tfdv.StatsOptions(
weight_feature=WEIGHT_COLUMN_NAME,
sample_rate=1.0
)
)
tfdv.visualize_statistics(train_stats)
```
### 1.2 Infer Schema
```
schema = tfdv.infer_schema(statistics=train_stats)
tfdv.display_schema(schema=schema)
print(tfdv.get_feature(schema, 'age'))
```
### 1.3 Configure Schema
```
# Relax the minimum fraction of values that must come from the domain for feature occupation.
occupation = tfdv.get_feature(schema, 'occupation')
occupation.distribution_constraints.min_domain_mass = 0.9
# Add new value to the domain of feature native_country, assuming that we start receiving this
# we won't be able to make great predictions of course, because this country is not part of our
# training data.
native_country_domain = tfdv.get_domain(schema, 'native_country')
native_country_domain.value.append('Egypt')
# All features are by default in both TRAINING and SERVING environments.
schema.default_environment.append('TRAINING')
schema.default_environment.append('EVALUATION')
schema.default_environment.append('SERVING')
# Specify that the class feature is not in SERVING environment.
tfdv.get_feature(schema, TARGET_FEATURE_NAME).not_in_environment.append('SERVING')
tfdv.display_schema(schema=schema)
```
### 1.4 Validate evaluation data
```
eval_stats = tfdv.generate_statistics_from_csv(
EVAL_DATA_FILE,
column_names=HEADER,
stats_options=tfdv.StatsOptions(
weight_feature=WEIGHT_COLUMN_NAME)
)
eval_anomalies = tfdv.validate_statistics(eval_stats, schema, environment='EVALUATION')
tfdv.display_anomalies(eval_anomalies)
```
### 1.5 Freeze the schema
```
RAW_SCHEMA_LOCATION = 'raw_schema.pbtxt'
from tensorflow.python.lib.io import file_io
from google.protobuf import text_format
tfdv.write_schema_text(schema, RAW_SCHEMA_LOCATION)
!cat {RAW_SCHEMA_LOCATION}
```
## License
Copyright 2019 Google LLC
Licensed under the Apache License, Version 2.0 (the "License");
you may not use this file except in compliance with the License.
You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0.
Unless required by applicable law or agreed to in writing, software
distributed under the License is distributed on an "AS IS" BASIS,
WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
See the License for the specific language governing permissions and
limitations under the License.
---
**Disclaimer**: This is not an official Google product. The sample code provided for an educational purpose.
---
| github_jupyter |
```
# Copyright 2020 Google LLC
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Vertex client library: AutoML image object detection model for online prediction
<table align="left">
<td>
<a href="https://colab.research.google.com/github/GoogleCloudPlatform/vertex-ai-samples/blob/master/notebooks/community/gapic/automl/showcase_automl_image_object_detection_online.ipynb">
<img src="https://cloud.google.com/ml-engine/images/colab-logo-32px.png" alt="Colab logo"> Run in Colab
</a>
</td>
<td>
<a href="https://github.com/GoogleCloudPlatform/vertex-ai-samples/blob/master/notebooks/community/gapic/automl/showcase_automl_image_object_detection_online.ipynb">
<img src="https://cloud.google.com/ml-engine/images/github-logo-32px.png" alt="GitHub logo">
View on GitHub
</a>
</td>
</table>
<br/><br/><br/>
## Overview
This tutorial demonstrates how to use the Vertex client library for Python to create image object detection models and do online prediction using Google Cloud's [AutoML](https://cloud.google.com/vertex-ai/docs/start/automl-users).
### Dataset
The dataset used for this tutorial is the Salads category of the [OpenImages dataset](https://www.tensorflow.org/datasets/catalog/open_images_v4) from [TensorFlow Datasets](https://www.tensorflow.org/datasets/catalog/overview). This dataset does not require any feature engineering. The version of the dataset you will use in this tutorial is stored in a public Cloud Storage bucket. The trained model predicts the bounding box locations and corresponding type of salad items in an image from a class of five items: salad, seafood, tomato, baked goods, or cheese.
### Objective
In this tutorial, you create an AutoML image object detection model and deploy for online prediction from a Python script using the Vertex client library. You can alternatively create and deploy models using the `gcloud` command-line tool or online using the Google Cloud Console.
The steps performed include:
- Create a Vertex `Dataset` resource.
- Train the model.
- View the model evaluation.
- Deploy the `Model` resource to a serving `Endpoint` resource.
- Make a prediction.
- Undeploy the `Model`.
### Costs
This tutorial uses billable components of Google Cloud (GCP):
* Vertex AI
* Cloud Storage
Learn about [Vertex AI
pricing](https://cloud.google.com/vertex-ai/pricing) and [Cloud Storage
pricing](https://cloud.google.com/storage/pricing), and use the [Pricing
Calculator](https://cloud.google.com/products/calculator/)
to generate a cost estimate based on your projected usage.
## Installation
Install the latest version of Vertex client library.
```
import os
import sys
# Google Cloud Notebook
if os.path.exists("/opt/deeplearning/metadata/env_version"):
USER_FLAG = "--user"
else:
USER_FLAG = ""
! pip3 install -U google-cloud-aiplatform $USER_FLAG
```
Install the latest GA version of *google-cloud-storage* library as well.
```
! pip3 install -U google-cloud-storage $USER_FLAG
```
### Restart the kernel
Once you've installed the Vertex client library and Google *cloud-storage*, you need to restart the notebook kernel so it can find the packages.
```
if not os.getenv("IS_TESTING"):
# Automatically restart kernel after installs
import IPython
app = IPython.Application.instance()
app.kernel.do_shutdown(True)
```
## Before you begin
### GPU runtime
*Make sure you're running this notebook in a GPU runtime if you have that option. In Colab, select* **Runtime > Change Runtime Type > GPU**
### Set up your Google Cloud project
**The following steps are required, regardless of your notebook environment.**
1. [Select or create a Google Cloud project](https://console.cloud.google.com/cloud-resource-manager). When you first create an account, you get a $300 free credit towards your compute/storage costs.
2. [Make sure that billing is enabled for your project.](https://cloud.google.com/billing/docs/how-to/modify-project)
3. [Enable the Vertex APIs and Compute Engine APIs.](https://console.cloud.google.com/flows/enableapi?apiid=ml.googleapis.com,compute_component)
4. [The Google Cloud SDK](https://cloud.google.com/sdk) is already installed in Google Cloud Notebook.
5. Enter your project ID in the cell below. Then run the cell to make sure the
Cloud SDK uses the right project for all the commands in this notebook.
**Note**: Jupyter runs lines prefixed with `!` as shell commands, and it interpolates Python variables prefixed with `$` into these commands.
```
PROJECT_ID = "[your-project-id]" # @param {type:"string"}
if PROJECT_ID == "" or PROJECT_ID is None or PROJECT_ID == "[your-project-id]":
# Get your GCP project id from gcloud
shell_output = !gcloud config list --format 'value(core.project)' 2>/dev/null
PROJECT_ID = shell_output[0]
print("Project ID:", PROJECT_ID)
! gcloud config set project $PROJECT_ID
```
#### Region
You can also change the `REGION` variable, which is used for operations
throughout the rest of this notebook. Below are regions supported for Vertex. We recommend that you choose the region closest to you.
- Americas: `us-central1`
- Europe: `europe-west4`
- Asia Pacific: `asia-east1`
You may not use a multi-regional bucket for training with Vertex. Not all regions provide support for all Vertex services. For the latest support per region, see the [Vertex locations documentation](https://cloud.google.com/vertex-ai/docs/general/locations)
```
REGION = "us-central1" # @param {type: "string"}
```
#### Timestamp
If you are in a live tutorial session, you might be using a shared test account or project. To avoid name collisions between users on resources created, you create a timestamp for each instance session, and append onto the name of resources which will be created in this tutorial.
```
from datetime import datetime
TIMESTAMP = datetime.now().strftime("%Y%m%d%H%M%S")
```
### Authenticate your Google Cloud account
**If you are using Google Cloud Notebook**, your environment is already authenticated. Skip this step.
**If you are using Colab**, run the cell below and follow the instructions when prompted to authenticate your account via oAuth.
**Otherwise**, follow these steps:
In the Cloud Console, go to the [Create service account key](https://console.cloud.google.com/apis/credentials/serviceaccountkey) page.
**Click Create service account**.
In the **Service account name** field, enter a name, and click **Create**.
In the **Grant this service account access to project** section, click the Role drop-down list. Type "Vertex" into the filter box, and select **Vertex Administrator**. Type "Storage Object Admin" into the filter box, and select **Storage Object Admin**.
Click Create. A JSON file that contains your key downloads to your local environment.
Enter the path to your service account key as the GOOGLE_APPLICATION_CREDENTIALS variable in the cell below and run the cell.
```
# If you are running this notebook in Colab, run this cell and follow the
# instructions to authenticate your GCP account. This provides access to your
# Cloud Storage bucket and lets you submit training jobs and prediction
# requests.
# If on Google Cloud Notebook, then don't execute this code
if not os.path.exists("/opt/deeplearning/metadata/env_version"):
if "google.colab" in sys.modules:
from google.colab import auth as google_auth
google_auth.authenticate_user()
# If you are running this notebook locally, replace the string below with the
# path to your service account key and run this cell to authenticate your GCP
# account.
elif not os.getenv("IS_TESTING"):
%env GOOGLE_APPLICATION_CREDENTIALS ''
```
### Set up variables
Next, set up some variables used throughout the tutorial.
### Import libraries and define constants
#### Import Vertex client library
Import the Vertex client library into our Python environment.
```
import time
from google.cloud.aiplatform import gapic as aip
from google.protobuf import json_format
from google.protobuf.json_format import MessageToJson, ParseDict
from google.protobuf.struct_pb2 import Struct, Value
```
#### Vertex constants
Setup up the following constants for Vertex:
- `API_ENDPOINT`: The Vertex API service endpoint for dataset, model, job, pipeline and endpoint services.
- `PARENT`: The Vertex location root path for dataset, model, job, pipeline and endpoint resources.
```
# API service endpoint
API_ENDPOINT = "{}-aiplatform.googleapis.com".format(REGION)
# Vertex location root path for your dataset, model and endpoint resources
PARENT = "projects/" + PROJECT_ID + "/locations/" + REGION
```
#### AutoML constants
Set constants unique to AutoML datasets and training:
- Dataset Schemas: Tells the `Dataset` resource service which type of dataset it is.
- Data Labeling (Annotations) Schemas: Tells the `Dataset` resource service how the data is labeled (annotated).
- Dataset Training Schemas: Tells the `Pipeline` resource service the task (e.g., classification) to train the model for.
```
# Image Dataset type
DATA_SCHEMA = "gs://google-cloud-aiplatform/schema/dataset/metadata/image_1.0.0.yaml"
# Image Labeling type
LABEL_SCHEMA = "gs://google-cloud-aiplatform/schema/dataset/ioformat/image_bounding_box_io_format_1.0.0.yaml"
# Image Training task
TRAINING_SCHEMA = "gs://google-cloud-aiplatform/schema/trainingjob/definition/automl_image_object_detection_1.0.0.yaml"
```
# Tutorial
Now you are ready to start creating your own AutoML image object detection model.
## Set up clients
The Vertex client library works as a client/server model. On your side (the Python script) you will create a client that sends requests and receives responses from the Vertex server.
You will use different clients in this tutorial for different steps in the workflow. So set them all up upfront.
- Dataset Service for `Dataset` resources.
- Model Service for `Model` resources.
- Pipeline Service for training.
- Endpoint Service for deployment.
- Prediction Service for serving.
```
# client options same for all services
client_options = {"api_endpoint": API_ENDPOINT}
def create_dataset_client():
client = aip.DatasetServiceClient(client_options=client_options)
return client
def create_model_client():
client = aip.ModelServiceClient(client_options=client_options)
return client
def create_pipeline_client():
client = aip.PipelineServiceClient(client_options=client_options)
return client
def create_endpoint_client():
client = aip.EndpointServiceClient(client_options=client_options)
return client
def create_prediction_client():
client = aip.PredictionServiceClient(client_options=client_options)
return client
clients = {}
clients["dataset"] = create_dataset_client()
clients["model"] = create_model_client()
clients["pipeline"] = create_pipeline_client()
clients["endpoint"] = create_endpoint_client()
clients["prediction"] = create_prediction_client()
for client in clients.items():
print(client)
```
## Dataset
Now that your clients are ready, your first step in training a model is to create a managed dataset instance, and then upload your labeled data to it.
### Create `Dataset` resource instance
Use the helper function `create_dataset` to create the instance of a `Dataset` resource. This function does the following:
1. Uses the dataset client service.
2. Creates an Vertex `Dataset` resource (`aip.Dataset`), with the following parameters:
- `display_name`: The human-readable name you choose to give it.
- `metadata_schema_uri`: The schema for the dataset type.
3. Calls the client dataset service method `create_dataset`, with the following parameters:
- `parent`: The Vertex location root path for your `Database`, `Model` and `Endpoint` resources.
- `dataset`: The Vertex dataset object instance you created.
4. The method returns an `operation` object.
An `operation` object is how Vertex handles asynchronous calls for long running operations. While this step usually goes fast, when you first use it in your project, there is a longer delay due to provisioning.
You can use the `operation` object to get status on the operation (e.g., create `Dataset` resource) or to cancel the operation, by invoking an operation method:
| Method | Description |
| ----------- | ----------- |
| result() | Waits for the operation to complete and returns a result object in JSON format. |
| running() | Returns True/False on whether the operation is still running. |
| done() | Returns True/False on whether the operation is completed. |
| canceled() | Returns True/False on whether the operation was canceled. |
| cancel() | Cancels the operation (this may take up to 30 seconds). |
```
TIMEOUT = 90
def create_dataset(name, schema, labels=None, timeout=TIMEOUT):
start_time = time.time()
try:
dataset = aip.Dataset(
display_name=name, metadata_schema_uri=schema, labels=labels
)
operation = clients["dataset"].create_dataset(parent=PARENT, dataset=dataset)
print("Long running operation:", operation.operation.name)
result = operation.result(timeout=TIMEOUT)
print("time:", time.time() - start_time)
print("response")
print(" name:", result.name)
print(" display_name:", result.display_name)
print(" metadata_schema_uri:", result.metadata_schema_uri)
print(" metadata:", dict(result.metadata))
print(" create_time:", result.create_time)
print(" update_time:", result.update_time)
print(" etag:", result.etag)
print(" labels:", dict(result.labels))
return result
except Exception as e:
print("exception:", e)
return None
result = create_dataset("salads-" + TIMESTAMP, DATA_SCHEMA)
```
Now save the unique dataset identifier for the `Dataset` resource instance you created.
```
# The full unique ID for the dataset
dataset_id = result.name
# The short numeric ID for the dataset
dataset_short_id = dataset_id.split("/")[-1]
print(dataset_id)
```
### Data preparation
The Vertex `Dataset` resource for images has some requirements for your data:
- Images must be stored in a Cloud Storage bucket.
- Each image file must be in an image format (PNG, JPEG, BMP, ...).
- There must be an index file stored in your Cloud Storage bucket that contains the path and label for each image.
- The index file must be either CSV or JSONL.
#### CSV
For image object detection, the CSV index file has the requirements:
- No heading.
- First column is the Cloud Storage path to the image.
- Second column is the label.
- Third/Fourth columns are the upper left corner of bounding box. Coordinates are normalized, between 0 and 1.
- Fifth/Sixth/Seventh columns are not used and should be 0.
- Eighth/Ninth columns are the lower right corner of the bounding box.
#### Location of Cloud Storage training data.
Now set the variable `IMPORT_FILE` to the location of the CSV index file in Cloud Storage.
```
IMPORT_FILE = "gs://cloud-samples-data/vision/salads.csv"
```
#### Quick peek at your data
You will use a version of the Salads dataset that is stored in a public Cloud Storage bucket, using a CSV index file.
Start by doing a quick peek at the data. You count the number of examples by counting the number of rows in the CSV index file (`wc -l`) and then peek at the first few rows.
```
if "IMPORT_FILES" in globals():
FILE = IMPORT_FILES[0]
else:
FILE = IMPORT_FILE
count = ! gsutil cat $FILE | wc -l
print("Number of Examples", int(count[0]))
print("First 10 rows")
! gsutil cat $FILE | head
```
### Import data
Now, import the data into your Vertex Dataset resource. Use this helper function `import_data` to import the data. The function does the following:
- Uses the `Dataset` client.
- Calls the client method `import_data`, with the following parameters:
- `name`: The human readable name you give to the `Dataset` resource (e.g., salads).
- `import_configs`: The import configuration.
- `import_configs`: A Python list containing a dictionary, with the key/value entries:
- `gcs_sources`: A list of URIs to the paths of the one or more index files.
- `import_schema_uri`: The schema identifying the labeling type.
The `import_data()` method returns a long running `operation` object. This will take a few minutes to complete. If you are in a live tutorial, this would be a good time to ask questions, or take a personal break.
```
def import_data(dataset, gcs_sources, schema):
config = [{"gcs_source": {"uris": gcs_sources}, "import_schema_uri": schema}]
print("dataset:", dataset_id)
start_time = time.time()
try:
operation = clients["dataset"].import_data(
name=dataset_id, import_configs=config
)
print("Long running operation:", operation.operation.name)
result = operation.result()
print("result:", result)
print("time:", int(time.time() - start_time), "secs")
print("error:", operation.exception())
print("meta :", operation.metadata)
print(
"after: running:",
operation.running(),
"done:",
operation.done(),
"cancelled:",
operation.cancelled(),
)
return operation
except Exception as e:
print("exception:", e)
return None
import_data(dataset_id, [IMPORT_FILE], LABEL_SCHEMA)
```
## Train the model
Now train an AutoML image object detection model using your Vertex `Dataset` resource. To train the model, do the following steps:
1. Create an Vertex training pipeline for the `Dataset` resource.
2. Execute the pipeline to start the training.
### Create a training pipeline
You may ask, what do we use a pipeline for? You typically use pipelines when the job (such as training) has multiple steps, generally in sequential order: do step A, do step B, etc. By putting the steps into a pipeline, we gain the benefits of:
1. Being reusable for subsequent training jobs.
2. Can be containerized and ran as a batch job.
3. Can be distributed.
4. All the steps are associated with the same pipeline job for tracking progress.
Use this helper function `create_pipeline`, which takes the following parameters:
- `pipeline_name`: A human readable name for the pipeline job.
- `model_name`: A human readable name for the model.
- `dataset`: The Vertex fully qualified dataset identifier.
- `schema`: The dataset labeling (annotation) training schema.
- `task`: A dictionary describing the requirements for the training job.
The helper function calls the `Pipeline` client service'smethod `create_pipeline`, which takes the following parameters:
- `parent`: The Vertex location root path for your `Dataset`, `Model` and `Endpoint` resources.
- `training_pipeline`: the full specification for the pipeline training job.
Let's look now deeper into the *minimal* requirements for constructing a `training_pipeline` specification:
- `display_name`: A human readable name for the pipeline job.
- `training_task_definition`: The dataset labeling (annotation) training schema.
- `training_task_inputs`: A dictionary describing the requirements for the training job.
- `model_to_upload`: A human readable name for the model.
- `input_data_config`: The dataset specification.
- `dataset_id`: The Vertex dataset identifier only (non-fully qualified) -- this is the last part of the fully-qualified identifier.
- `fraction_split`: If specified, the percentages of the dataset to use for training, test and validation. Otherwise, the percentages are automatically selected by AutoML.
```
def create_pipeline(pipeline_name, model_name, dataset, schema, task):
dataset_id = dataset.split("/")[-1]
input_config = {
"dataset_id": dataset_id,
"fraction_split": {
"training_fraction": 0.8,
"validation_fraction": 0.1,
"test_fraction": 0.1,
},
}
training_pipeline = {
"display_name": pipeline_name,
"training_task_definition": schema,
"training_task_inputs": task,
"input_data_config": input_config,
"model_to_upload": {"display_name": model_name},
}
try:
pipeline = clients["pipeline"].create_training_pipeline(
parent=PARENT, training_pipeline=training_pipeline
)
print(pipeline)
except Exception as e:
print("exception:", e)
return None
return pipeline
```
### Construct the task requirements
Next, construct the task requirements. Unlike other parameters which take a Python (JSON-like) dictionary, the `task` field takes a Google protobuf Struct, which is very similar to a Python dictionary. Use the `json_format.ParseDict` method for the conversion.
The minimal fields you need to specify are:
- `budget_milli_node_hours`: The maximum time to budget (billed) for training the model, where 1000 = 1 hour. For image object detection, the budget must be a minimum of 20 hours.
- `model_type`: The type of deployed model:
- `CLOUD_HIGH_ACCURACY_1`: For deploying to Google Cloud and optimizing for accuracy.
- `CLOUD_LOW_LATENCY_1`: For deploying to Google Cloud and optimizing for latency (response time),
- `MOBILE_TF_HIGH_ACCURACY_1`: For deploying to the edge and optimizing for accuracy.
- `MOBILE_TF_LOW_LATENCY_1`: For deploying to the edge and optimizing for latency (response time).
- `MOBILE_TF_VERSATILE_1`: For deploying to the edge and optimizing for a trade off between latency and accuracy.
- `disable_early_stopping`: Whether True/False to let AutoML use its judgement to stop training early or train for the entire budget.
Finally, create the pipeline by calling the helper function `create_pipeline`, which returns an instance of a training pipeline object.
```
PIPE_NAME = "salads_pipe-" + TIMESTAMP
MODEL_NAME = "salads_model-" + TIMESTAMP
task = json_format.ParseDict(
{
"budget_milli_node_hours": 20000,
"model_type": "CLOUD_HIGH_ACCURACY_1",
"disable_early_stopping": False,
},
Value(),
)
response = create_pipeline(PIPE_NAME, MODEL_NAME, dataset_id, TRAINING_SCHEMA, task)
```
Now save the unique identifier of the training pipeline you created.
```
# The full unique ID for the pipeline
pipeline_id = response.name
# The short numeric ID for the pipeline
pipeline_short_id = pipeline_id.split("/")[-1]
print(pipeline_id)
```
### Get information on a training pipeline
Now get pipeline information for just this training pipeline instance. The helper function gets the job information for just this job by calling the the job client service's `get_training_pipeline` method, with the following parameter:
- `name`: The Vertex fully qualified pipeline identifier.
When the model is done training, the pipeline state will be `PIPELINE_STATE_SUCCEEDED`.
```
def get_training_pipeline(name, silent=False):
response = clients["pipeline"].get_training_pipeline(name=name)
if silent:
return response
print("pipeline")
print(" name:", response.name)
print(" display_name:", response.display_name)
print(" state:", response.state)
print(" training_task_definition:", response.training_task_definition)
print(" training_task_inputs:", dict(response.training_task_inputs))
print(" create_time:", response.create_time)
print(" start_time:", response.start_time)
print(" end_time:", response.end_time)
print(" update_time:", response.update_time)
print(" labels:", dict(response.labels))
return response
response = get_training_pipeline(pipeline_id)
```
# Deployment
Training the above model may take upwards of 60 minutes time.
Once your model is done training, you can calculate the actual time it took to train the model by subtracting `end_time` from `start_time`. For your model, you will need to know the fully qualified Vertex Model resource identifier, which the pipeline service assigned to it. You can get this from the returned pipeline instance as the field `model_to_deploy.name`.
```
while True:
response = get_training_pipeline(pipeline_id, True)
if response.state != aip.PipelineState.PIPELINE_STATE_SUCCEEDED:
print("Training job has not completed:", response.state)
model_to_deploy_id = None
if response.state == aip.PipelineState.PIPELINE_STATE_FAILED:
raise Exception("Training Job Failed")
else:
model_to_deploy = response.model_to_upload
model_to_deploy_id = model_to_deploy.name
print("Training Time:", response.end_time - response.start_time)
break
time.sleep(60)
print("model to deploy:", model_to_deploy_id)
```
## Model information
Now that your model is trained, you can get some information on your model.
## Evaluate the Model resource
Now find out how good the model service believes your model is. As part of training, some portion of the dataset was set aside as the test (holdout) data, which is used by the pipeline service to evaluate the model.
### List evaluations for all slices
Use this helper function `list_model_evaluations`, which takes the following parameter:
- `name`: The Vertex fully qualified model identifier for the `Model` resource.
This helper function uses the model client service's `list_model_evaluations` method, which takes the same parameter. The response object from the call is a list, where each element is an evaluation metric.
For each evaluation -- you probably only have one, we then print all the key names for each metric in the evaluation, and for a small set (`evaluatedBoundingBoxCount` and `boundingBoxMeanAveragePrecision`) you will print the result.
```
def list_model_evaluations(name):
response = clients["model"].list_model_evaluations(parent=name)
for evaluation in response:
print("model_evaluation")
print(" name:", evaluation.name)
print(" metrics_schema_uri:", evaluation.metrics_schema_uri)
metrics = json_format.MessageToDict(evaluation._pb.metrics)
for metric in metrics.keys():
print(metric)
print("evaluatedBoundingBoxCount", metrics["evaluatedBoundingBoxCount"])
print(
"boundingBoxMeanAveragePrecision",
metrics["boundingBoxMeanAveragePrecision"],
)
return evaluation.name
last_evaluation = list_model_evaluations(model_to_deploy_id)
```
## Deploy the `Model` resource
Now deploy the trained Vertex `Model` resource you created with AutoML. This requires two steps:
1. Create an `Endpoint` resource for deploying the `Model` resource to.
2. Deploy the `Model` resource to the `Endpoint` resource.
### Create an `Endpoint` resource
Use this helper function `create_endpoint` to create an endpoint to deploy the model to for serving predictions, with the following parameter:
- `display_name`: A human readable name for the `Endpoint` resource.
The helper function uses the endpoint client service's `create_endpoint` method, which takes the following parameter:
- `display_name`: A human readable name for the `Endpoint` resource.
Creating an `Endpoint` resource returns a long running operation, since it may take a few moments to provision the `Endpoint` resource for serving. You call `response.result()`, which is a synchronous call and will return when the Endpoint resource is ready. The helper function returns the Vertex fully qualified identifier for the `Endpoint` resource: `response.name`.
```
ENDPOINT_NAME = "salads_endpoint-" + TIMESTAMP
def create_endpoint(display_name):
endpoint = {"display_name": display_name}
response = clients["endpoint"].create_endpoint(parent=PARENT, endpoint=endpoint)
print("Long running operation:", response.operation.name)
result = response.result(timeout=300)
print("result")
print(" name:", result.name)
print(" display_name:", result.display_name)
print(" description:", result.description)
print(" labels:", result.labels)
print(" create_time:", result.create_time)
print(" update_time:", result.update_time)
return result
result = create_endpoint(ENDPOINT_NAME)
```
Now get the unique identifier for the `Endpoint` resource you created.
```
# The full unique ID for the endpoint
endpoint_id = result.name
# The short numeric ID for the endpoint
endpoint_short_id = endpoint_id.split("/")[-1]
print(endpoint_id)
```
### Compute instance scaling
You have several choices on scaling the compute instances for handling your online prediction requests:
- Single Instance: The online prediction requests are processed on a single compute instance.
- Set the minimum (`MIN_NODES`) and maximum (`MAX_NODES`) number of compute instances to one.
- Manual Scaling: The online prediction requests are split across a fixed number of compute instances that you manually specified.
- Set the minimum (`MIN_NODES`) and maximum (`MAX_NODES`) number of compute instances to the same number of nodes. When a model is first deployed to the instance, the fixed number of compute instances are provisioned and online prediction requests are evenly distributed across them.
- Auto Scaling: The online prediction requests are split across a scaleable number of compute instances.
- Set the minimum (`MIN_NODES`) number of compute instances to provision when a model is first deployed and to de-provision, and set the maximum (`MAX_NODES) number of compute instances to provision, depending on load conditions.
The minimum number of compute instances corresponds to the field `min_replica_count` and the maximum number of compute instances corresponds to the field `max_replica_count`, in your subsequent deployment request.
```
MIN_NODES = 1
MAX_NODES = 1
```
### Deploy `Model` resource to the `Endpoint` resource
Use this helper function `deploy_model` to deploy the `Model` resource to the `Endpoint` resource you created for serving predictions, with the following parameters:
- `model`: The Vertex fully qualified model identifier of the model to upload (deploy) from the training pipeline.
- `deploy_model_display_name`: A human readable name for the deployed model.
- `endpoint`: The Vertex fully qualified endpoint identifier to deploy the model to.
The helper function calls the `Endpoint` client service's method `deploy_model`, which takes the following parameters:
- `endpoint`: The Vertex fully qualified `Endpoint` resource identifier to deploy the `Model` resource to.
- `deployed_model`: The requirements specification for deploying the model.
- `traffic_split`: Percent of traffic at the endpoint that goes to this model, which is specified as a dictionary of one or more key/value pairs.
- If only one model, then specify as **{ "0": 100 }**, where "0" refers to this model being uploaded and 100 means 100% of the traffic.
- If there are existing models on the endpoint, for which the traffic will be split, then use `model_id` to specify as **{ "0": percent, model_id: percent, ... }**, where `model_id` is the model id of an existing model to the deployed endpoint. The percents must add up to 100.
Let's now dive deeper into the `deployed_model` parameter. This parameter is specified as a Python dictionary with the minimum required fields:
- `model`: The Vertex fully qualified model identifier of the (upload) model to deploy.
- `display_name`: A human readable name for the deployed model.
- `disable_container_logging`: This disables logging of container events, such as execution failures (default is container logging is enabled). Container logging is typically enabled when debugging the deployment and then disabled when deployed for production.
- `automatic_resources`: This refers to how many redundant compute instances (replicas). For this example, we set it to one (no replication).
#### Traffic Split
Let's now dive deeper into the `traffic_split` parameter. This parameter is specified as a Python dictionary. This might at first be a tad bit confusing. Let me explain, you can deploy more than one instance of your model to an endpoint, and then set how much (percent) goes to each instance.
Why would you do that? Perhaps you already have a previous version deployed in production -- let's call that v1. You got better model evaluation on v2, but you don't know for certain that it is really better until you deploy to production. So in the case of traffic split, you might want to deploy v2 to the same endpoint as v1, but it only get's say 10% of the traffic. That way, you can monitor how well it does without disrupting the majority of users -- until you make a final decision.
#### Response
The method returns a long running operation `response`. We will wait sychronously for the operation to complete by calling the `response.result()`, which will block until the model is deployed. If this is the first time a model is deployed to the endpoint, it may take a few additional minutes to complete provisioning of resources.
```
DEPLOYED_NAME = "salads_deployed-" + TIMESTAMP
def deploy_model(
model, deployed_model_display_name, endpoint, traffic_split={"0": 100}
):
deployed_model = {
"model": model,
"display_name": deployed_model_display_name,
"automatic_resources": {
"min_replica_count": MIN_NODES,
"max_replica_count": MAX_NODES,
},
}
response = clients["endpoint"].deploy_model(
endpoint=endpoint, deployed_model=deployed_model, traffic_split=traffic_split
)
print("Long running operation:", response.operation.name)
result = response.result()
print("result")
deployed_model = result.deployed_model
print(" deployed_model")
print(" id:", deployed_model.id)
print(" model:", deployed_model.model)
print(" display_name:", deployed_model.display_name)
print(" create_time:", deployed_model.create_time)
return deployed_model.id
deployed_model_id = deploy_model(model_to_deploy_id, DEPLOYED_NAME, endpoint_id)
```
## Make a online prediction request
Now do a online prediction to your deployed model.
### Get test item
You will use an arbitrary example out of the dataset as a test item. Don't be concerned that the example was likely used in training the model -- we just want to demonstrate how to make a prediction.
```
test_items = !gsutil cat $IMPORT_FILE | head -n1
cols = str(test_items[0]).split(",")
if len(cols) == 11:
test_item = str(cols[1])
test_label = str(cols[2])
else:
test_item = str(cols[0])
test_label = str(cols[1])
print(test_item, test_label)
```
### Make a prediction
Now you have a test item. Use this helper function `predict_item`, which takes the following parameters:
- `filename`: The Cloud Storage path to the test item.
- `endpoint`: The Vertex fully qualified identifier for the `Endpoint` resource where the `Model` resource was deployed.
- `parameters_dict`: Additional filtering parameters for serving prediction results.
This function calls the prediction client service's `predict` method with the following parameters:
- `endpoint`: The Vertex fully qualified identifier for the `Endpoint` resource where the `Model` resource was deployed.
- `instances`: A list of instances (encoded images) to predict.
- `parameters`: Additional parameters for serving.
- `confidence_threshold`: The threshold for returning predictions. Must be between 0 and 1.
- `max_predictions`: The maximum number of predictions per object to return, sorted by confidence.
You might ask, how does confidence_threshold affect the model accuracy? The threshold won't change the accuracy. What it changes is recall and precision.
- Precision: The higher the precision the more likely what is predicted is the correct prediction, but return fewer predictions. Increasing the confidence threshold increases precision.
- Recall: The higher the recall the more likely a correct prediction is returned in the result, but return more prediction with incorrect prediction. Decreasing the confidence threshold increases recall.
In this example, you will predict for precision. You set the confidence threshold to 0.5 and the maximum number of predictions for an object to two. Since, all the confidence values across the classes must add up to one, there are only two possible outcomes:
1. There is a tie, both 0.5, and returns two predictions.
2. One value is above 0.5 and all the rest are below 0.5, and returns one prediction.
#### Request
Since in this example your test item is in a Cloud Storage bucket, you will open and read the contents of the image using `tf.io.gfile.Gfile()`. To pass the test data to the prediction service, we will encode the bytes into base 64 -- This makes binary data safe from modification while it is transferred over the Internet.
The format of each instance is:
{ 'content': { 'b64': [base64_encoded_bytes] } }
Since the `predict()` method can take multiple items (instances), you send our single test item as a list of one test item. As a final step, you package the instances list into Google's protobuf format -- which is what we pass to the `predict()` method.
#### Response
The `response` object returns a list, where each element in the list corresponds to the corresponding image in the request. You will see in the output for each prediction -- in our case there is just one:
- `confidences`: Confidence level in the prediction.
- `displayNames`: The predicted label.
- `bboxes`: The bounding box for the label.
```
import base64
import tensorflow as tf
def predict_item(filename, endpoint, parameters_dict):
parameters = json_format.ParseDict(parameters_dict, Value())
with tf.io.gfile.GFile(filename, "rb") as f:
content = f.read()
# The format of each instance should conform to the deployed model's prediction input schema.
instances_list = [{"content": base64.b64encode(content).decode("utf-8")}]
instances = [json_format.ParseDict(s, Value()) for s in instances_list]
response = clients["prediction"].predict(
endpoint=endpoint, instances=instances, parameters=parameters
)
print("response")
print(" deployed_model_id:", response.deployed_model_id)
predictions = response.predictions
print("predictions")
for prediction in predictions:
print(" prediction:", dict(prediction))
predict_item(test_item, endpoint_id, {"confidenceThreshold": 0.5, "maxPredictions": 2})
```
## Undeploy the `Model` resource
Now undeploy your `Model` resource from the serving `Endpoint` resoure. Use this helper function `undeploy_model`, which takes the following parameters:
- `deployed_model_id`: The model deployment identifier returned by the endpoint service when the `Model` resource was deployed to.
- `endpoint`: The Vertex fully qualified identifier for the `Endpoint` resource where the `Model` is deployed to.
This function calls the endpoint client service's method `undeploy_model`, with the following parameters:
- `deployed_model_id`: The model deployment identifier returned by the endpoint service when the `Model` resource was deployed.
- `endpoint`: The Vertex fully qualified identifier for the `Endpoint` resource where the `Model` resource is deployed.
- `traffic_split`: How to split traffic among the remaining deployed models on the `Endpoint` resource.
Since this is the only deployed model on the `Endpoint` resource, you simply can leave `traffic_split` empty by setting it to {}.
```
def undeploy_model(deployed_model_id, endpoint):
response = clients["endpoint"].undeploy_model(
endpoint=endpoint, deployed_model_id=deployed_model_id, traffic_split={}
)
print(response)
undeploy_model(deployed_model_id, endpoint_id)
```
# Cleaning up
To clean up all GCP resources used in this project, you can [delete the GCP
project](https://cloud.google.com/resource-manager/docs/creating-managing-projects#shutting_down_projects) you used for the tutorial.
Otherwise, you can delete the individual resources you created in this tutorial:
- Dataset
- Pipeline
- Model
- Endpoint
- Batch Job
- Custom Job
- Hyperparameter Tuning Job
- Cloud Storage Bucket
```
delete_dataset = True
delete_pipeline = True
delete_model = True
delete_endpoint = True
delete_batchjob = True
delete_customjob = True
delete_hptjob = True
delete_bucket = True
# Delete the dataset using the Vertex fully qualified identifier for the dataset
try:
if delete_dataset and "dataset_id" in globals():
clients["dataset"].delete_dataset(name=dataset_id)
except Exception as e:
print(e)
# Delete the training pipeline using the Vertex fully qualified identifier for the pipeline
try:
if delete_pipeline and "pipeline_id" in globals():
clients["pipeline"].delete_training_pipeline(name=pipeline_id)
except Exception as e:
print(e)
# Delete the model using the Vertex fully qualified identifier for the model
try:
if delete_model and "model_to_deploy_id" in globals():
clients["model"].delete_model(name=model_to_deploy_id)
except Exception as e:
print(e)
# Delete the endpoint using the Vertex fully qualified identifier for the endpoint
try:
if delete_endpoint and "endpoint_id" in globals():
clients["endpoint"].delete_endpoint(name=endpoint_id)
except Exception as e:
print(e)
# Delete the batch job using the Vertex fully qualified identifier for the batch job
try:
if delete_batchjob and "batch_job_id" in globals():
clients["job"].delete_batch_prediction_job(name=batch_job_id)
except Exception as e:
print(e)
# Delete the custom job using the Vertex fully qualified identifier for the custom job
try:
if delete_customjob and "job_id" in globals():
clients["job"].delete_custom_job(name=job_id)
except Exception as e:
print(e)
# Delete the hyperparameter tuning job using the Vertex fully qualified identifier for the hyperparameter tuning job
try:
if delete_hptjob and "hpt_job_id" in globals():
clients["job"].delete_hyperparameter_tuning_job(name=hpt_job_id)
except Exception as e:
print(e)
if delete_bucket and "BUCKET_NAME" in globals():
! gsutil rm -r $BUCKET_NAME
```
| github_jupyter |
##### Copyright 2019 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# 利用 Keras 来训练多工作器(worker)
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://tensorflow.google.cn/tutorials/distribute/multi_worker_with_keras"><img src="https://tensorflow.google.cn/images/tf_logo_32px.png" />在 tensorflow.google.cn 上查看</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/zh-cn/tutorials/distribute/multi_worker_with_keras.ipynb"><img src="https://tensorflow.google.cn/images/colab_logo_32px.png" />在 Google Colab 运行</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/zh-cn/tutorials/distribute/multi_worker_with_keras.ipynb"><img src="https://tensorflow.google.cn/images/GitHub-Mark-32px.png" />在 Github 上查看源代码</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/docs-l10n/site/zh-cn/tutorials/distribute/multi_worker_with_keras.ipynb"><img src="https://tensorflow.google.cn/images/download_logo_32px.png" />下载此 notebook</a>
</td>
</table>
Note: 我们的 TensorFlow 社区翻译了这些文档。因为社区翻译是尽力而为, 所以无法保证它们是最准确的,并且反映了最新的
[官方英文文档](https://www.tensorflow.org/?hl=en)。如果您有改进此翻译的建议, 请提交 pull request 到
[tensorflow/docs](https://github.com/tensorflow/docs) GitHub 仓库。要志愿地撰写或者审核译文,请加入
[docs-zh-cn@tensorflow.org Google Group](https://groups.google.com/a/tensorflow.org/forum/#!forum/docs-zh-cn)。
## 概述
本教程使用 `tf.distribute.Strategy` API 演示了使用 Keras 模型的多工作器(worker)分布式培训。借助专为多工作器(worker)训练而设计的策略,设计在单一工作器(worker)上运行的 Keras 模型可以在最少的代码更改的情况下无缝地处理多个工作器。
[TensorFlow 中的分布式培训](../../guide/distribute_strategy.ipynb)指南可用于概述TensorFlow支持的分布式策略,并想要更深入理解`tf.distribute.Strategy` API 感兴趣的人。
## 配置
首先,设置 TensorFlow 和必要的导入。
```
from __future__ import absolute_import, division, print_function, unicode_literals
try:
# % tensorflow_version仅存在于Colab中。
%tensorflow_version 2.x
except Exception:
pass
import tensorflow_datasets as tfds
import tensorflow as tf
tfds.disable_progress_bar()
```
## 准备数据集
现在,让我们从 [TensorFlow 数据集](https://tensorflow.google.cn/datasets) 中准备MNIST数据集。 [MNIST 数据集](http://yann.lecun.com/exdb/mnist/) 包括60,000个训练样本和10,000个手写数字0-9的测试示例,格式为28x28像素单色图像。
```
BUFFER_SIZE = 10000
BATCH_SIZE = 64
# 将 MNIST 数据从 (0, 255] 缩放到 (0., 1.]
def scale(image, label):
image = tf.cast(image, tf.float32)
image /= 255
return image, label
datasets, info = tfds.load(name='mnist',
with_info=True,
as_supervised=True)
train_datasets_unbatched = datasets['train'].map(scale).cache().shuffle(BUFFER_SIZE)
train_datasets = train_datasets_unbatched.batch(BATCH_SIZE)
```
## 构建 Keras 模型
在这里,我们使用`tf.keras.Sequential` API来构建和编译一个简单的卷积神经网络 Keras 模型,用我们的 MNIST 数据集进行训练。
注意:有关构建 Keras 模型的详细训练说明,请参阅[TensorFlow Keras 指南](https://tensorflow.google.cn/guide/keras#sequential_model)。
```
def build_and_compile_cnn_model():
model = tf.keras.Sequential([
tf.keras.layers.Conv2D(32, 3, activation='relu', input_shape=(28, 28, 1)),
tf.keras.layers.MaxPooling2D(),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(64, activation='relu'),
tf.keras.layers.Dense(10, activation='softmax')
])
model.compile(
loss=tf.keras.losses.sparse_categorical_crossentropy,
optimizer=tf.keras.optimizers.SGD(learning_rate=0.001),
metrics=['accuracy'])
return model
```
让我们首先尝试用少量的 epoch 来训练模型,并在单个工作器(worker)中观察结果,以确保一切正常。 随着训练的迭代,您应该会看到损失(loss)下降和准确度(accuracy)接近1.0。
```
single_worker_model = build_and_compile_cnn_model()
single_worker_model.fit(x=train_datasets, epochs=3)
```
## 多工作器(worker)配置
现在让我们进入多工作器(worker)训练的世界。在 TensorFlow 中,需要 `TF_CONFIG` 环境变量来训练多台机器,每台机器可能具有不同的角色。 `TF_CONFIG`用于指定作为集群一部分的每个 worker 的集群配置。
`TF_CONFIG` 有两个组件:`cluster` 和 `task` 。 `cluster` 提供有关训练集群的信息,这是一个由不同类型的工作组成的字典,例如 `worker` 。在多工作器(worker)培训中,除了常规的“工作器”之外,通常还有一个“工人”承担更多责任,比如保存检查点和为 TensorBoard 编写摘要文件。这样的工作器(worker)被称为“主要”工作者,习惯上`worker` 中 `index` 0被指定为主要的 `worker`(事实上这就是`tf.distribute.Strategy`的实现方式)。 另一方面,`task` 提供当前任务的信息。
在这个例子中,我们将任务 `type` 设置为 `"worker"` 并将任务 `index` 设置为 `0` 。这意味着具有这种设置的机器是第一个工作器,它将被指定为主要工作器并且要比其他工作器做更多的工作。请注意,其他机器也需要设置 `TF_CONFIG` 环境变量,它应该具有相同的 `cluster` 字典,但是不同的任务`type` 或 `index` 取决于这些机器的角色。
为了便于说明,本教程展示了如何在 `localhost` 上设置一个带有2个工作器的`TF_CONFIG`。 实际上,用户会在外部IP地址/端口上创建多个工作器,并在每个工作器上适当地设置`TF_CONFIG`。
警告:不要在 Colab 中执行以下代码。TensorFlow 的运行时将尝试在指定的IP地址和端口创建 gRPC 服务器,这可能会失败。
```
os.environ['TF_CONFIG'] = json.dumps({
'cluster': {
'worker': ["localhost:12345", "localhost:23456"]
},
'task': {'type': 'worker', 'index': 0}
})
```
注意,虽然在该示例中学习速率是固定的,但是通常可能需要基于全局批量大小来调整学习速率。
## 选择正确的策略
在 TensorFlow 中,分布式训练包括同步训练(其中训练步骤跨工作器和副本同步)、异步训练(训练步骤未严格同步)。
`MultiWorkerMirroredStrategy` 是同步多工作器训练的推荐策略,将在本指南中进行演示。
要训练模型,请使用 `tf.distribute.experimental.MultiWorkerMirroredStrategy` 的实例。 `MultiWorkerMirroredStrategy` 在所有工作器的每台设备上创建模型层中所有变量的副本。 它使用 `CollectiveOps` ,一个用于集体通信的 TensorFlow 操作,来聚合梯度并使变量保持同步。 [`tf.distribute.Strategy`指南](../../guide/distribute_strategy.ipynb)有关于此策略的更多详细信息。
```
strategy = tf.distribute.experimental.MultiWorkerMirroredStrategy()
```
注意:解析 `TF_CONFIG` 并且在调用 `MultiWorkerMirroredStrategy.__init__()` 时启动 TensorFlow 的 GRPC 服务器,因此必须在创建`tf.distribute.Strategy`实例之前设置 `TF_CONFIG` 环境变量。
`MultiWorkerMirroredStrategy` 通过[`CollectiveCommunication`](https://github.com/tensorflow/tensorflow/blob/a385a286a930601211d78530734368ccb415bee4/tensorflow/python/distribute/cross_device_ops.py#L928)参数提供多个实现。`RING` 使用 gRPC 作为跨主机通信层实现基于环的集合。`NCCL` 使用[Nvidia 的 NCCL](https://developer.nvidia.com/nccl)来实现集体。 `AUTO` 将选择推迟到运行时。 集体实现的最佳选择取决于GPU的数量和种类以及群集中的网络互连。
## 使用 MultiWorkerMirroredStrategy 训练模型
通过将 `tf.distribute.Strategy` API集成到 `tf.keras` 中,将训练分发给多人的唯一更改就是将模型进行构建和 `model.compile()` 调用封装在 `strategy.scope()` 内部。 分发策略的范围决定了如何创建变量以及在何处创建变量,对于 MultiWorkerMirroredStrategy 而言,创建的变量为 MirroredVariable ,并且将它们复制到每个工作器上。
注意:在此Colab中,以下代码可以按预期结果运行,但是由于未设置`TF_CONFIG`,因此这实际上是单机训练。 在您自己的示例中设置了 `TF_CONFIG` 后,您应该期望在多台机器上进行培训可以提高速度。
```
NUM_WORKERS = 2
# 由于`tf.data.Dataset.batch`需要全局的批处理大小,
# 因此此处的批处理大小按工作器数量增加。
# 以前我们使用64,现在变成128。
GLOBAL_BATCH_SIZE = 64 * NUM_WORKERS
train_datasets = train_datasets_unbatched.batch(GLOBAL_BATCH_SIZE)
with strategy.scope():
multi_worker_model = build_and_compile_cnn_model()
multi_worker_model.fit(x=train_datasets, epochs=3)
```
### 数据集分片和批(batch)大小
在多工作器训练中,需要将数据分片为多个部分,以确保融合和性能。 但是,请注意,在上面的代码片段中,数据集直接发送到`model.fit()`,而无需分片; 这是因为`tf.distribute.Strategy` API在多工作器训练中会自动处理数据集分片。
如果您喜欢手动分片进行训练,则可以通过`tf.data.experimental.DistributeOptions` API关闭自动分片。
```
options = tf.data.Options()
options.experimental_distribute.auto_shard = False
train_datasets_no_auto_shard = train_datasets.with_options(options)
```
要注意的另一件事是 `datasets` 的批处理大小。 在上面的代码片段中,我们使用 `GLOBAL_BATCH_SIZE = 64 * NUM_WORKERS` ,这是单个工作器的大小的 `NUM_WORKERS` 倍,因为每个工作器的有效批量大小是全局批量大小(参数从 `tf.data.Dataset.batch()` 传入)除以工作器的数量,通过此更改,我们使每个工作器的批处理大小与以前相同。
## 性能
现在,您已经有了一个Keras模型,该模型全部通过 `MultiWorkerMirroredStrategy` 运行在多个工作器中。 您可以尝试以下技术来调整多工作器训练的效果。
* `MultiWorkerMirroredStrategy` 提供了多个[集体通信实现][collective communication implementations](https://github.com/tensorflow/tensorflow/blob/master/tensorflow/python/distribute/cross_device_ops.py). `RING` 使用gRPC作为跨主机通信层实现基于环的集合。 `NCCL` 使用 [Nvidia's NCCL](https://developer.nvidia.com/nccl) 来实现集合。 `AUTO` 将推迟到运行时选择。集体实施的最佳选择取决于GPU的数量和种类以及集群中的网络互连。 要覆盖自动选择,请为 `MultiWorkerMirroredStrategy` 的构造函数的 `communication` 参数指定一个有效值,例如: `communication=tf.distribute.experimental.CollectiveCommunication.NCCL`.
* 如果可能的话,将变量强制转换为 `tf.float`。ResNet 的官方模型包括如何完成此操作的[示例](https://github.com/tensorflow/models/blob/8367cf6dabe11adf7628541706b660821f397dce/official/resnet/resnet_model.py#L466)。
## 容错能力
在同步训练中,如果其中一个工作器出现故障并且不存在故障恢复机制,则集群将失败。 在工作器退出或不稳定的情况下,将 Keras 与 `tf.distribute.Strategy` 一起使用会具有容错的优势。 我们通过在您选择的分布式文件系统中保留训练状态来做到这一点,以便在重新启动先前失败或被抢占的实例后,将恢复训练状态。
由于所有工作器在训练 epochs 和 steps 方面保持同步,因此其他工作器将需要等待失败或被抢占的工作器重新启动才能继续。
### ModelCheckpoint 回调
要在多工作器训练中利用容错功能,请在调用 `tf.keras.Model.fit()` 时提供一个 `tf.keras.callbacks.ModelCheckpoint` 实例。 回调会将检查点和训练状态存储在与 `ModelCheckpoint` 的 `filepath` 参数相对应的目录中。
```
# 将`filepath`参数替换为文件系统中的路径
# 使所有工作器能够访问到
callbacks = [tf.keras.callbacks.ModelCheckpoint(filepath='/tmp/keras-ckpt')]
with strategy.scope():
multi_worker_model = build_and_compile_cnn_model()
multi_worker_model.fit(x=train_datasets, epochs=3, callbacks=callbacks)
```
如果某个工作线程被抢占,则整个集群将暂停,直到重新启动被抢占的工作线程为止。工作器重新加入集群后,其他工作器也将重新启动。 现在,每个工作器都将读取先前保存的检查点文件,并获取其以前的状态,从而使群集能够恢复同步,然后继续训练。
如果检查包含在`ModelCheckpoint` 中指定的 `filepath` 的目录,则可能会注意到一些临时生成的检查点文件。 这些文件是恢复以前丢失的实例所必需的,并且在成功退出多工作器训练后,这些文件将在 `tf.keras.Model.fit()` 的末尾被库删除。
## 您可以查阅
1. [Distributed Training in TensorFlow](https://www.tensorflow.org/guide/distribute_strategy) 该指南概述了可用的分布式策略。
2. [ResNet50](https://github.com/tensorflow/models/blob/master/official/resnet/imagenet_main.py) 官方模型,该模型可以使用 `MirroredStrategy` 或 `MultiWorkerMirroredStrategy` 进行训练
| github_jupyter |
# Notebook for testing the TensorFlow 2.0 setup
This netbook is for testing the [TensorFlow](https://www.tensorflow.org/) setup using the [Keras API](https://keras.io/). Below is a set of required imports.
Run the cell, and no error messages should appear. In particular, **TensorFlow 2 is required**.
Some warnings may appear, this should be fine.
```
%matplotlib inline
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers
from tensorflow.keras.utils import plot_model, to_categorical
from tensorflow.keras.datasets import mnist, fashion_mnist, imdb
import os
if not os.path.isfile('pml_utils.py'):
!wget https://raw.githubusercontent.com/csc-training/intro-to-dl/master/day1/pml_utils.py
from pml_utils import show_failures
from sklearn.model_selection import train_test_split
from distutils.version import LooseVersion as LV
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
sns.set()
print('Using Tensorflow version: {}, and Keras version: {}.'.format(tf.__version__, tf.keras.__version__))
assert(LV(tf.__version__) >= LV("2.0.0"))
```
Let's check if we have GPU available.
```
gpus = tf.config.list_physical_devices('GPU')
if len(gpus) > 0:
for gpu in gpus:
tf.config.experimental.set_memory_growth(gpu, True)
from tensorflow.python.client import device_lib
for d in device_lib.list_local_devices():
if d.device_type == 'GPU':
print('GPU', d.physical_device_desc)
else:
print('No GPU, using CPU instead.')
```
## Getting started: 30 seconds to Keras
(This section is adapted from https://keras.io/)
The core data structure of Keras is a *Model*, a way to organize layers. While there are several ways to create Models in Keras, we will be using the [*functional* API](https://keras.io/guides/functional_api/).
We start by creating an input layer:
```
inputs = keras.Input(shape=(100,))
```
We create further layers by calling a specific layer on its input object:
```
x = layers.Dense(units=64, activation="relu")(inputs)
outputs = layers.Dense(units=10, activation="softmax")(x)
```
Then we can create a Model by specifying its inputs and outputs:
```
model = keras.Model(inputs=inputs, outputs=outputs, name="test_model")
```
A summary of the model:
```
print(model.summary())
```
Let's draw a fancier graph of our model:
```
plot_model(model, show_shapes=True)
```
Once your model looks good, configure its learning process with `.compile()`:
```
model.compile(loss='categorical_crossentropy',
optimizer='sgd',
metrics=['accuracy'])
```
You can now begin training your model with `.fit()`. Let's generate some random data and use it to train the model:
```
X_train = np.random.rand(128, 100)
Y_train = to_categorical(np.random.randint(10, size=128))
model.fit(X_train, Y_train, epochs=5, batch_size=32, verbose=2);
```
Evaluate your performance on test data with `.evaluate():`
```
X_test = np.random.rand(64, 100)
Y_test = to_categorical(np.random.randint(10, size=64))
loss, acc = model.evaluate(X_test, Y_test, batch_size=32)
print()
print('loss:', loss, 'acc:', acc)
```
---
*Run this notebook in Google Colaboratory using [this link](https://colab.research.google.com/github/csc-training/intro-to-dl/blob/master/day1/01-tf2-test-setup.ipynb).*
| github_jupyter |
```
import os
import numpy as np
import h5py
import gdal
from osgeo import gdal, gdalconst, osr
import time
import torch
import matplotlib.pyplot as plt
from sklearn.metrics import confusion_matrix, f1_score, accuracy_score
# !nvidia-smi
experiment_id = 'sentinel_2A_2018_T11SKA_spatio_temporal_attention_segmentation'
quadrant_size = (5490,5490)
grid_cell_size = (1372,1372)
patch_size = (32,32)
input_patch_size = 32
label_patch_size = (16,16)
timestamps = [0,2,3,5,8,11,14,17,19,22,24,26,28,30] # Timestamps to consider
no_timestamps = time_steps = len(timestamps)
no_features = channels = 10 # Number of Features
step_size = 16
output_patch_width = 16
no_of_grid_cells_x = 4
no_of_grid_cells_y = 4
batch_size = 16
diff = 8
total_no_of_grid_cells = no_of_grid_cells_x * no_of_grid_cells_y
grids_train = [0,2,5,7,8,10,13,15]
grids_test = [1,3,4,6,9,11,12,14]
no_of_classes = 20
labels_list = [0,1,2,3,4,5,6,7,8,9,10,11,12,13,14,15,16,17,18,19]
learning_rate = 0.0001
max_accuracy_test = 0
no_of_epochs = 100
quadrant = 4
data_dir = 'data/sentinel/sentinel_2A_2018_T11SKA/numpy_arrays/quadrant_wise/quadrant' + str(quadrant)
numpy_array_prefix = 'sentinel_2A_2018_T11SKA_series_quadrant4_patch'
label_dir = 'data/sentinel/sentinel_2A_2018_T11SKA/labels/quadrant_wise/quadrant' + str(quadrant)
label_array_prefix = 'sentinel_2A_2018_T11SKA_raw_label_quadrant4_patch'
model_folder = 'models_sentinel_2A_2018_T11SKA_spatio_temporal_attention_segmentation/' + experiment_id
if not os.path.exists(model_folder):
os.makedirs(model_folder)
#label conversion from usda raw labels
def convert_to_label_array(raw_label,label):
if(label.shape != raw_label.shape):
print("Shapes not equal")
label[(raw_label==1) | (raw_label==225) | (raw_label==226) | (raw_label==237)] = 1 # Corn
label[(raw_label==2) | (raw_label==238)] = 2 # Cotton
label[(raw_label==4) | (raw_label==236)] = 3 # Sorghum
label[(raw_label==22) | (raw_label==23) | (raw_label==24)] = 4 # Wheat
label[(raw_label==36)] = 5 # Alfa alfa
label[(raw_label==67)] = 6 # Peaches
label[(raw_label==69)] = 7 # Grapes
label[(raw_label==71)] = 8 # Tree crops
label[(raw_label==72)] = 9 # Citrus
label[(raw_label==75)] = 10 # Almonds
label[(raw_label==76)] = 11 # Walnut
label[(raw_label==204)] = 12 # Pistachio
label[(raw_label==212)] = 13 # Oranges
label[(raw_label==218)] = 14 # Nectarines
label[(raw_label==5) | (raw_label==3) | (raw_label==27) | (raw_label==28) | (raw_label==44) | (raw_label==53) | (raw_label==21) | (raw_label==33) |(raw_label==42) | (raw_label==205)] = 15 # Misc Crops and Veg
label[(raw_label==37) | (raw_label==58) | (raw_label==59) | (raw_label==61) | (raw_label==152) | (raw_label==176) | (raw_label==190) | (raw_label==195)] = 16 # Wetlands and Grass
label[(raw_label==61) | (raw_label==131)] = 17 # Barren/Idle land
label[(raw_label==111)] = 18 # Water
label[(raw_label==121) | (raw_label==122) | (raw_label==123) | (raw_label==124)] = 19 # Urban
return label
# Model Architecture
class UNET_LSTM_BIDIRECTIONAL_ATTENTION(torch.nn.Module):
def __init__(self, in_channels, out_channels):
super(UNET_LSTM_BIDIRECTIONAL_ATTENTION,self).__init__()
self.conv1_1 = torch.nn.Conv2d(in_channels, 64, 3, padding=1)
self.conv1_2 = torch.nn.Conv2d(64, 64, 3, padding=1)
self.conv2_1 = torch.nn.Conv2d(64, 128, 3, padding=1)
self.conv2_2 = torch.nn.Conv2d(128, 128, 3, padding=1)
self.conv3_1 = torch.nn.Conv2d(128, 256, 3, padding=1)
self.conv3_2 = torch.nn.Conv2d(256, 256, 3, padding=1)
self.lstm = torch.nn.LSTM(256, 256, batch_first=True, bidirectional=True)
self.attention = torch.nn.Linear(512, 1)
self.unpool2 = torch.nn.ConvTranspose2d(512 , 128, kernel_size=2, stride=2)
self.upconv2_1 = torch.nn.Conv2d(256, 128, 3, padding=1)
self.upconv2_2 = torch.nn.Conv2d(128, 128, 3, padding=1)
self.unpool1 = torch.nn.ConvTranspose2d(128 , 64, kernel_size=2, stride=2)
self.upconv1_1 = torch.nn.Conv2d(128, 64, 3, padding=1)
self.upconv1_2 = torch.nn.Conv2d(64, 64, 3, padding=1)
self.out = torch.nn.Conv2d(64, out_channels, kernel_size=1, padding=0)
self.maxpool = torch.nn.MaxPool2d(2)
self.relu = torch.nn.ReLU(inplace=True)
self.dropout = torch.nn.Dropout(p=0.1)
for m in self.modules():
if isinstance(m, torch.nn.Conv2d) or isinstance(m, torch.nn.Linear):
torch.nn.init.xavier_uniform_(m.weight)
def crop_and_concat(self, x1, x2):
x1_shape = x1.shape
x2_shape = x2.shape
offset_2, offset_3 = (x1_shape[2]-x2_shape[2])//2, (x1_shape[3]-x2_shape[3])//2
x1_crop = x1[:, :, offset_2:offset_2+x2_shape[2], offset_3:offset_3+x2_shape[3]]
return torch.cat([x1_crop, x2], dim=1)
def forward(self,x):
x = x.view(-1, channels, input_patch_size, input_patch_size)
conv1 = self.relu(self.conv1_2(self.relu(self.conv1_1(x))))
maxpool1 = self.maxpool(conv1)
conv2 = self.relu(self.conv2_2(self.relu(self.conv2_1(maxpool1))))
maxpool2 = self.maxpool(conv2)
conv3 = self.relu(self.conv3_2(self.relu(self.conv3_1(maxpool2))))
shape_enc = conv3.shape
conv3 = conv3.view(-1, time_steps, conv3.shape[1], conv3.shape[2]*conv3.shape[3])
conv3 = conv3.permute(0,3,1,2)
conv3 = conv3.reshape(conv3.shape[0]*conv3.shape[1], time_steps, 256)
lstm, _ = self.lstm(conv3)
lstm = self.relu(lstm.reshape(-1, 512))
attention_weights = torch.nn.functional.softmax(torch.squeeze(torch.nn.functional.avg_pool2d(self.attention(torch.tanh(lstm)).view(-1,shape_enc[2],shape_enc[3],time_steps).permute(0,3,1,2), 8)), dim=1)
context = torch.sum((attention_weights.view(-1, 1, 1, time_steps).repeat(1, 8, 8, 1).view(-1, 1)*lstm).view(-1, time_steps, 512), dim=1).view(-1,shape_enc[2],shape_enc[3], 512).permute(0,3,1,2)
attention_weights_fixed = attention_weights.detach()
unpool2 = self.unpool2(context)
agg_conv2 = torch.sum(attention_weights_fixed.view(-1, time_steps, 1, 1, 1) * conv2.view(-1, time_steps, conv2.shape[1], conv2.shape[2], conv2.shape[3]), dim=1)
upconv2 = self.relu(self.upconv2_2(self.relu(self.upconv2_1(self.crop_and_concat(agg_conv2, unpool2)))))
unpool1 = self.unpool1(upconv2)
agg_conv1 = torch.sum(attention_weights_fixed.view(-1, time_steps, 1, 1, 1) * conv1.view(-1, time_steps, conv1.shape[1], conv1.shape[2], conv1.shape[3]), dim=1)
upconv1 = self.relu(self.upconv1_2(self.relu(self.upconv1_1(self.crop_and_concat(agg_conv1, unpool1)))))
out = self.out(upconv1)
return out[:,:,diff:-diff, diff:-diff]
# build model
model = UNET_LSTM_BIDIRECTIONAL_ATTENTION(in_channels=no_features, out_channels=no_of_classes)
model = model.to('cuda')
criterion = torch.nn.CrossEntropyLoss()
optimizer = torch.optim.Adam(model.parameters(), lr=learning_rate)
## train model
train_loss = []
train_accuracy = []
test_loss = []
test_accuracy = []
no_of_patches_x = int((grid_cell_size[0] - (patch_size[0]))/step_size)
no_of_patches_y = int((grid_cell_size[1] - (patch_size[1]))/step_size)
no_of_patches_x_test = int((grid_cell_size[0] - (patch_size[0]))/step_size)
no_of_patches_y_test = int((grid_cell_size[1] - (patch_size[1]))/step_size)
w = int((patch_size[0]-label_patch_size[0])/2)
no_of_batches_y = int(no_of_patches_y/batch_size)
no_of_batches_y_test = int(no_of_patches_y_test/batch_size)
image_batch = np.zeros((batch_size, ) + (no_timestamps, ) + patch_size + (no_features, ))
label_batch = np.zeros((batch_size, ) + label_patch_size)
print(no_of_patches_x,no_of_batches_y, image_batch.shape)
for epoch in range(no_of_epochs):
print('\n## EPOCH ',epoch,' ##')
# Train
print('\tTraining')
model.train()
total_loss = 0
accuracy_grids = 0
start_time = time.time()
for grid in grids_train:
start_grid_time = time.time()
accuracy_rows = 0
train_grid_cell = np.load(os.path.join(data_dir,numpy_array_prefix + str(grid) + '.npy'))
train_grid_cell_raw_label = np.load(os.path.join(label_dir,label_array_prefix + str(grid) + '.npy'))
train_grid_cell_label = np.zeros((grid_cell_size))
train_grid_cell_label = convert_to_label_array(train_grid_cell_raw_label,train_grid_cell_label)
for x in range(no_of_patches_x):
accuracy = 0
for y in range(no_of_batches_y):
for b in range(batch_size):
image_batch[b] = train_grid_cell[timestamps, x*step_size:(x*step_size) + patch_size[0], (((y*batch_size)+b)*step_size):(((y*batch_size)+b)*step_size) + patch_size[1], :]
label_batch[b] = train_grid_cell_label[x*step_size + w:(x*step_size) + w + label_patch_size[0], (((y*batch_size)+b)*step_size) + w:(((y*batch_size)+b)*step_size) + w + label_patch_size[1]]
image_batch_tr = np.transpose(image_batch,(0,1,4,2,3))
image_batch_t = torch.Tensor(image_batch_tr)
label_batch_t = torch.Tensor(label_batch)
optimizer.zero_grad()
patch_out = model(image_batch_t.to('cuda'))
label_batch_t = label_batch_t.type(torch.long).to('cuda')
loss = criterion(patch_out, label_batch_t)
loss.backward()
optimizer.step()
total_loss += loss.item()
patch_out_pred = torch.argmax(torch.nn.functional.softmax(patch_out, dim=1), dim=1)
patch_out_pred = np.reshape(patch_out_pred.cpu().numpy(), (-1))
label_batch_t = np.reshape(label_batch_t.cpu().numpy(), (-1))
accuracy += accuracy_score(patch_out_pred,label_batch_t)
accuracy_rows += accuracy/no_of_batches_y
print('\t\tGrid no: ', grid, '\tAccuracy grid: ',accuracy_rows/no_of_patches_x, '\t Time taken for Grid: ', time.time() - start_grid_time)
accuracy_grids += accuracy_rows/no_of_patches_x
print('\n\tTrain:\t Loss: {1}\t Accuracy: {2}\t Time: {3}'.format(epoch, total_loss/(no_of_patches_x), (accuracy_grids/len(grids_train)), time.time() - start_time))
train_loss.append(total_loss/(no_of_patches_x))
train_accuracy.append(accuracy_grids/len(grids_train))
# Test
model.eval()
print('\n\tTesting')
total_loss_test = 0
accuracy_test_grids = 0
start_time_test = time.time()
pred_list = []
true_list = []
for grid in grids_test:
start_grid_time_test = time.time()
accuracy_test_row = 0
test_grid_cell = np.load(os.path.join(data_dir,numpy_array_prefix + str(grid) + '.npy'))
test_grid_cell_raw_label = np.load(os.path.join(label_dir,label_array_prefix + str(grid) + '.npy'))
test_grid_cell_label = np.zeros((grid_cell_size))
test_grid_cell_label = convert_to_label_array(test_grid_cell_raw_label,test_grid_cell_label)
for x in range(no_of_patches_x_test):
accuracy_test = 0
# print(str(x)+str('-'),end = '')
for y in range(no_of_batches_y_test):
for b in range(batch_size):
image_batch[b] = test_grid_cell[timestamps, x*step_size:(x*step_size) + patch_size[0], (((y*batch_size)+b)*step_size):(((y*batch_size)+b)*step_size) + patch_size[1], :]
label_batch[b] = test_grid_cell_label[x*step_size + w:(x*step_size) + w + label_patch_size[0], (((y*batch_size)+b)*step_size) + w:(((y*batch_size)+b)*step_size) + w + label_patch_size[1]]
image_batch_tr = np.transpose(image_batch,(0,1,4,2,3))
image_batch_t = torch.Tensor(image_batch_tr)
label_batch_t = torch.Tensor(label_batch)
patch_out = model(image_batch_t.to('cuda'))
label_batch_t = label_batch_t.type(torch.long).to('cuda')
loss_test = criterion(patch_out, label_batch_t)
patch_out_pred = torch.argmax(torch.nn.functional.softmax(patch_out, dim=1), dim=1)
total_loss_test += loss_test.item()
patch_out_pred = np.reshape(patch_out_pred.cpu().numpy(), (-1))
label_batch_t = np.reshape(label_batch_t.cpu().numpy(), (-1))
accuracy_test += accuracy_score(patch_out_pred,label_batch_t)
pred_list.append(patch_out_pred)
true_list.append(label_batch_t)
accuracy_test_row += accuracy_test/no_of_batches_y_test
print('\t\tGrid no: ', grid, '\tAccuracy grid: ',accuracy_test_row/no_of_patches_x_test, '\t Time taken for Grid: ', time.time() - start_grid_time_test)
accuracy_test_grids += accuracy_test_row/no_of_patches_x_test
pred_list_arr = np.array(pred_list).reshape(-1)
true_list_arr = np.array(true_list).reshape(-1)
mean_f1_score = np.mean(f1_score(true_list_arr,pred_list_arr,average = None,labels=labels_list))
print('\tTest:\t Loss: {}\t Accuracy: {}\t Time: {}'.format(total_loss_test/(no_of_patches_x_test), (accuracy_test_grids/len(grids_test)), time.time() - start_time_test))
print('\t\t\t Mean F1 Score: {}'.format(mean_f1_score))
test_loss.append(total_loss_test/(no_of_patches_x_test))
test_accuracy.append(accuracy_test_grids/len(grids_test))
model_name = 'state_dict_epoch-'+str(epoch)+'_test_acc-' + str("{:.4f}".format(accuracy_test_grids/len(grids_test))) + '_mean_f1_score-'+ str("{:.4f}".format(mean_f1_score))+'_'+str(experiment_id)+'.pt'
torch.save(model.state_dict(), os.path.join(model_folder, model_name))
print('Saved model at', str(os.path.join(model_folder, model_name)) )
# Plot graphs
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.plot(train_loss, label="train loss")
plt.plot(test_loss, label="test loss")
plt.legend(loc="upper right")
plt.savefig(os.path.join(model_folder, ('loss_'+experiment_id+'_pytorch.png')))
plt.show()
plt.close()
plt.title('model accuracy')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.plot(train_accuracy, label="train acc")
plt.plot(test_accuracy, label="test acc")
plt.legend(loc="lower right")
plt.savefig(os.path.join(model_folder, ('accuracy_'+experiment_id+'_pytorch.png')))
plt.show()
plt.close()
```
| github_jupyter |
# Python for n00bs
## Workshop 2: Advanced Python for coders
Welcome back to programming with Python! My name is Vikram Mark Radhakrishnan. You can find me on [LinkedIn](https://www.linkedin.com/in/vikram-mark-radhakrishnan-90038660/), or reach me via email at radhakrishnan@strw.leidenuniv.nl
Shout-out to the [AI Lab One](https://www.meetup.com/AI-Lab/) and [City AI](https://city.ai/) for making this workshop possible!
<img src="nb_images/AI_Lab.png">
### 1. Here's how we function!
We will begin today by learning how to write our own functions in Python.
Sometimes we need to write a lot of code to do a single, repeatable task. In this case it is useful to write our own function, and replace that block of code with a function call.
A function can accept zero to any number of parameters, and can either return something or not.
```
def bmi_calculator(h, w):
bmi = w / (h ** 2)
return bmi
```
We can set default values for the parameters passed to a function. These default parameters will be overwritten if the user passes different values.
```
def volume_of_cylinder(h=10, r=5):
vol = 3.14159 * r ** 2 * h
return vol
```
Python also allows us to write functions that take in a variable number of arguments. There are two ways you can do this. You can pass a parameter called \*args to the function, or/and a parameter called \*\*kwargs.
The former implies that you are passing a list of arguments, which in the function will be accessed as a list named args. The latter implies that you are passing a dictionary, which in the function will be accessed by kwargs. Let's take a look at some examples.
```
def multiplier(*args):
result = 1
for num in args:
result *= num
print("After multiplying all these numbers together we get: " + str(result))
def display_stats(**kwargs):
for key, value in kwargs.items():
print("The " + key + " is " + str(value))
display_stats(name="Vikram", age="29", job="PhD Student", hobby="Python Instructor")
```
**ToDo:** Write a function that accepts a list of numbers, and takes the mean or median of these numbers, based on a boolean parameter called "mean". If mean is true, which it is by default, the function returns the arithmetic mean of the numbers. If mean is false, the function returns the median, i.e. the middle value of the sorted list.
### 2. Let's file this away...
Python allows for a lot of reusability. You can use codes that other people have written, simply by using import. Libraries and Python packages that are extremely useful for data science, for example, are available to you in Python simply by installing them on your computer and importing them in your code.
<img src="nb_images/python.png">
A useful Python library is the [os](https://docs.python.org/3/library/os.html) library, i.e. the miscellaneous operating system interfaces library.
```
import os
current_directory = os.getcwd()
print("First we were in " + current_directory)
os.mkdir("test_dir")
os.chdir("test_dir")
os.getcwd()
os.listdir("../")
```
We can create new files with Python using its built in file handling capapbility. We open a file to write, read, or append to with the "open" keyword.
```
newfile = open("random.txt", "w")
newfile.write("This is a bunch of generic text. ")
newfile.write("I would like to say a few words, and here they are.\nNitwit! Blubber! Oddment! Tweak!")
newfile.close()
f = open("random.txt", "r")
print(f.readline())
f.close()
```
You don't want to forget to close the file after you are done with it! This is because when the file is opened, it is locked by Python, and cannot be accessed outside of Python, or opened in a differnt mode in the same code. It's better instead to use the "with" statement, which automatically takes care of closing the file, even if there is an exception.
```
with open("random.txt") as f:
read_data = f.read()
print(read_data)
```
**ToDo:** List all the files in the directory one level above your current working directory. Save this list in a file called "index.txt".
### 3. Try-ing to deal with errors
The try... except statement in Python is useful code for handling errors. If you execute a code that you suspect might fail for some particular reason, or if you just want to make sure your code runs through till the end without failing at certain problem spots, you can enclose the tricky code in a try block, and if this code generates an error, the code in the following except block will be executed. Let's look at an example:
```
# Let's try to read a file that doesn't exist:
try:
f = open("imaginaryfile.txt", "r")
f.readline()
except:
print("That didn't work! Maybe this file does not exist?")
```
**ToDo**: Write a snippet of code to enter two floating point numbers from a user and find their product. If the user enters something that is not a floating point number, print an error message.
### 4. Let's put your Python skills in action!
Let's move on to coding a fun little game in Python:
```
# We'll need this for what we are going to do next
from IPython.display import clear_output
```
We will code the game of Hangman together! The objective of the game is to guess a word or phrase by guessing letters. Each time you guess a letter that isn't part of the word, you lose an attempt. You have 10 attempts before the game is over.
```
# This function displays the blanks and letters
def display_word(gl, w, a=10):
# First clear the previous display
clear_output()
for character in w:
if character in gl: #display the letters already guessed
print(character, end = '')
elif character == ' ': #print spaces because we don't guess those
print(character, end = '')
else: #otherwise print a dash
print('_', end = '')
print("\nAttempts left: " + str(a))
# Let's make a list to store the letters that have already been guessed
# We set the number of attempts to guess
# Now player 1 inputs a word or phrase
# Convert it to lower case to reduce complexity
# We make a set out of this word to keep track of all the letters to guess
# We also remove spaces because we don't guess those
# Start a loop where you keep track of attempts
# Player 2 guesses a letter
# Check if this letter has not been guessed before, in which case add it to the list
# Check if the letter is in the actual word or phrase. If not, player 2 loses an attempt.
# Otherwise remove this letter from the list of unique letters
# Use the function written above to display the guessed word
# Check for a win
```
### 5. OOPs I did it again!
We are now going to look at something that can make your code a lot more efficient, modular, and reusable - Object Oriented Programming (OOP). We have already seen examples of "objects" in Python. Strings, lists, dictionaries, etc are all objects, that have associated attributes and methods. We refer to these as primitive data structures. We are now going to look into how we can create our own objects in Python and why this is useful.
First let's understand what a "class" is. Essentially, a class is a blueprint, or prototype of an object.
```
class Store:
# The initializer method. You never explicitly call this method, it runs when an object is instanciated
def __init__(self, money, **items):
self.money = money
self.products = items
def buy(self):
pass
def sell(self):
pass
# A class that inherits from the Store class. It extends the buy() and a sell() method polymorphically
class groceryStore(Store):
def __init__(self, money, **items):
super().__init__(money, **items)
def buy(self, item):
if item in self.products.keys():
self.products[item] += 1
else:
self.products[item] = 1
self.money -= 1
def sell(self, item):
if item in self.products.keys():
self.products[item] -= 1
self.money += 1
else:
print("Item out of stock")
# This class inherits from Store, extends the buy() and sell() method polymorphically, and has a trade() method
class fashionStore(Store):
def __init__(self, money, **items):
super().__init__(money, **items)
def buy(self, item):
if item in self.products.keys():
self.products[item] += 1
else:
self.products[item] = 1
self.money -= 100
def sell(self, item):
if item in self.products.keys():
self.products[item] -= 1
self.money += 100
else:
print("Item out of stock")
def trade(self, item1, item2):
if item1 in self.products.keys():
self.products[item1] -= 1
if item2 in self.products.keys():
self.products[item2] += 1
else:
self.products[item2] = 1
else:
print("Item out of stock")
```
To appreciate the power of OOP, we have to understand its underlying features:
* Encapsulation
* Inheritance
* Polymorphism
* Abstraction
**ToDo:** Let's code the hangman game again, only this time we use object oriented programming!
```
class HangMan:
def __init__(self, word, attempts):
self.word = word.lower()
self.attempts = attempts
self.gl = [] #an empty list initially which will store each guessed character
# Store the unique characters in the word excluding spaces
self.unique_letters = set(self.word)
try:
self.unique_letters.remove(' ')
except:
pass
def display_word(self):
"""Function to display the word with blanks for the not yet guessed letters"""
# First clear the previous display
clear_output()
for character in self.word:
if character in self.gl: #display the letters already guessed
print(character, end = '')
elif character == ' ': #print spaces because we don't guess those
print(character, end = '')
else: #otherwise print a dash
print('_', end = '')
print("\nAttempts left: " + str(self.attempts))
def guess(self, letter):
"""Check if this letter has not been guessed before, in which case add it to the list.
Then check if the letter is in the word."""
if letter in self.gl:
print("You guessed this letter before!")
else:
self.gl.append(letter)
if letter not in self.word:
self.attempts -= 1
else:
self.unique_letters.remove(letter)
def playGame(self):
"""Keep displaying the blanks and guessing letters until the game is won or lost."""
while(self.attempts > 0):
nextguess = input("Guess a letter: ")
self.guess(nextguess)
self.display_word()
if len(self.unique_letters) == 0:
break
if self.attempts > 0:
print("Victory!")
else:
print("You lose!")
```
| github_jupyter |
# 0. Import
```
import torch
from torch import tensor
```
# 1. Element-wise operations
Element-wise Operations หมายถึง ทำ Operations แยกคนละตัวเรียงตามตำแหน่ง ตัวแรกทำกับตัวแรก ตัวที่สองทำกับตัวที่สอง ... โดยอาจจะทำไปพร้อม ๆ กัน ทำขนานกันไปก็ได้
## 1.1 Basic operations
```
a = tensor([1., 2., 3., 4.])
```
บวก ลบ คูณ หาร ยกกำลัง กับ ตัวเลขธรรมดา ใส่ . จุด หลังตัวเลข หมายถึง ให้เป็นเลขแบบ Float จุดทศนิยม รองรับการคำนวนคณิตศาสตร์ เช่น 1. เหมือนกับ 1.0
```
a + 1
2**a
```
บวก ลบ คูณ หาร ยกกำลัง กับ tensor ด้วยกัน
```
b = torch.ones(4) + 1
b
a - b
a * b
j = torch.arange(5)
j
2**(j + 1) - j
```
## 1.2 เปรียบเทียบความเร็ว
Tensor
```
a = torch.arange(10000)
%timeit a + 1
```
วน loop array
```
l = range(10000)
%timeit [i+1 for i in l]
```
tensor เร็วกว่าประมาณเกือบ 40 เท่า เนื่องจากเป็นการประมวลผลแบบขนาน ไม่ได้ทำทีละ item เรียงไปเรื่อย ๆ
## 1.3 Multiplication
```
c = torch.ones((3, 3))
c
```
คูณแบบปกติ จะเป็น คูณแบบ element-wise ไม่ได้คูณแบบ matrix
```
c * c
```
คูณแบบ matrix dot product
```
c.matmul(c)
```
## 1.4 Comparison Operations
Operation เปรียบเทียบ ก็ element-wise
```
a = tensor([1, 2, 3, 4])
b = tensor([4, 2, 2, 4])
```
1 คือ True, 0 คือ False
```
a == b
a > b
```
## 1.5 Transcendental functions
ฟังก์ชันคณิตศาสตร์ ก็ element-wise
```
a = torch.arange(5.)
torch.sin(a)
torch.log(a)
torch.exp(a)
```
# 2. Broadcasting
ในกรณี 2 ฝั่งมีสมาชิกไม่เท่ากัน จะเกิดการ Broadcasting กระจายสมาชิกจาก 1 ให้กลายเป็นเท่ากัน ก่อนทำ Element-wise Operation
ใน Numpy, TensorFlow และ Pytorch มิติแรก 0 จะเป็น Row มิติที่สอง 1 จะเป็น Column
```
a = torch.arange(0, 40, 10).repeat(1, 1, 3).view(3, 4).t()
a
```
## 2.1 Row Broadcasting
กระจาย 1 Row เป็น 4 Row ก่อนบวก
```
b = tensor([0, 1, 2])
b
a + b
```
## 2.2 Column Broadcasting
กระจาย 1 Column เป็น 3 Column ก่อนบวก
```
c = tensor([[0], [1], [2], [3]])
c
a + c
```
## 2.3 Row and Column Broadcasting
```
d = tensor([2])
d
```
กระจายเป็น 4 Row 3 Column ก่อนบวก
```
a+d
```
หมายเหตุ การ Broadcasting ไม่ได้จำกัดอยู่แค่ 2 มิติ เราสามารถ Broadcast ได้ในทุก ๆ มิติ เช่น broardcast มิติที่ 4 ใน tensor 5 มิติ เป็นต้น
# 3. Indexing and Slicing
```
a = torch.ones((4, 5))
a
```
## 3.1 Indexing
Indexing คือ การเลือกเฉพาะ Row, Column ที่ต้องการขึ้นมา เช่น ให้ Row ที่ 0 เป็น ค่า 2
*ไม่ได้ระบุมิติที่ 2 ให้ถือว่าเป็น : คือเอาทุก Column*
```
a[0] = 2
a
```
ให้ Column ที่ 1 (เริ่มต้นที่ 0) เป็น 3
*หมายเหตุ : (เครื่องหมาย colon) แปลว่าทุกอัน ถ้าอยู่ตำแหน่งแรก แปลว่าทุก Row*
```
a[:, 1] = 3
a
```
ให้ Column ที่ 1 จากสุดท้าย เป็น 4
*Index ติดลบ แปลว่า นับจากท้ายสุด*
```
a[:, -1] = 4
a
```
## 3.2 Slicing
Slicing เลือกเอาช่วง Index ในลำดับที่ต้องการ
```
x = tensor([0, 1, 2, 3, 4, 5, 6, 7, 8, 9])
x
```
เริ่มที่ 3 ถึง 6 (ไม่รวม 6)
```
x[3:6]
```
เริ่มที่ 1 ถึง 7 (ไม่รวม 7) ข้ามทีละ 2
```
x[1:7:2]
```
เลือกอันดับ 2 จากสุดท้าย ถึง อันดับ 10 (ไม่รวม 10)
```
x[-2:10]
```
เลือกตั้งแต่ 5 เป็นต้นไป
```
x[5:]
```
เลือกตั้งแต่ต้นจนถึง 5 (ไม่รวม 5)
```
x[:5]
```
# 4. Summation
การหาผลรวมของ Tensor เนื่องจาก Tensor มีหลายมิติ เราจึงต้องกำหนดว่า จะคิดตามมิติไหน ดังตัวอย่าง
```
a = tensor([[1, 1, 1], [2, 2, 2], [3, 3, 3]])
a
```
สมมติ a คือ Tensor 2 มิติ ขนาด 3 x 3
```
a.shape
```
Sum ตามแนวมิติที่ 0 คือ Row
```
s = a.sum(0)
s
s.shape
```
Sum ตามแนวมิติที่ 1 คือ Column
```
s = a.sum(1)
s
s.shape
```
เมื่อมีการ Sum มิตินั้นจะหายไป ทำให้ Tensor 2 มิติ กลายเป็น Tensor 1 มิติ หรือ Vector ถ้าเราต้องการให้มิติคงเดิม ให้เราใส่ keepdim=True
```
s = a.sum(0, keepdim=True)
s
s.shape
s = a.sum(1, keepdim=True)
s
s.shape
```
Sum มิติท้ายสุด
```
a.sum(-1)
```
*มิติ ติดลบ แปลว่า นับจากท้ายสุด*
# Credit
* http://scipy-lectures.org/intro/numpy/operations.html
* https://docs.scipy.org/doc/numpy/reference/arrays.indexing.html
* https://course.fast.ai/videos/?lesson=8
| github_jupyter |
# TensorFlow Tutorial #16
# Reinforcement Learning (Q-Learning)
by [Magnus Erik Hvass Pedersen](http://www.hvass-labs.org/)
/ [GitHub](https://github.com/Hvass-Labs/TensorFlow-Tutorials) / [Videos on YouTube](https://www.youtube.com/playlist?list=PL9Hr9sNUjfsmEu1ZniY0XpHSzl5uihcXZ)
## Introduction
This tutorial is about so-called Reinforcement Learning in which an agent is learning how to navigate some environment, in this case Atari games from the 1970-80's. The agent does not know anything about the game and must learn how to play it from trial and error. The only information that is available to the agent is the screen output of the game, and whether the previous action resulted in a reward or penalty.
This is a very difficult problem in Machine Learning / Artificial Intelligence, because the agent must both learn to distinguish features in the game-images, and then connect the occurence of certain features in the game-images with its own actions and a reward or penalty that may be deferred many steps into the future.
This problem was first solved by the researchers from Google DeepMind. This tutorial is based on the main ideas from their early research papers (especially [this](https://arxiv.org/abs/1312.5602) and [this](http://www.nature.com/nature/journal/v518/n7540/full/nature14236.html)), although we make several changes because the original DeepMind algorithm was awkward and over-complicated in some ways. But it turns out that you still need several tricks in order to stabilize the training of the agent, so the implementation in this tutorial is unfortunately also somewhat complicated.
The basic idea is to have the agent estimate so-called Q-values whenever it sees an image from the game-environment. The Q-values tell the agent which action is most likely to lead to the highest cumulative reward in the future. The problem is then reduced to finding these Q-values and storing them for later retrieval using a function approximator.
This builds on some of the previous tutorials. You should be familiar with TensorFlow and Convolutional Neural Networks from Tutorial #01 and #02. It will also be helpful if you are familiar with one of the builder APIs in Tutorials #03 or #03-B.
## The Problem
This tutorial uses the Atari game Breakout, where the player or agent is supposed to hit a ball with a paddle, thus avoiding death while scoring points when the ball smashes pieces of a wall.
When a human learns to play a game like this, the first thing to figure out is what part of the game environment you are controlling - in this case the paddle at the bottom. If you move right on the joystick then the paddle moves right and vice versa. The next thing is to figure out what the goal of the game is - in this case to smash as many bricks in the wall as possible so as to maximize the score. Finally you need to learn what to avoid - in this case you must avoid dying by letting the ball pass beside the paddle.
Below are shown 3 images from the game that demonstrate what we need our agent to learn. In the image to the left, the ball is going downwards and the agent must learn to move the paddle so as to hit the ball and avoid death. The image in the middle shows the paddle hitting the ball, which eventually leads to the image on the right where the ball smashes some bricks and scores points. The ball then continues downwards and the process repeats.

The problem is that there are 10 states between the ball going downwards and the paddle hitting the ball, and there are an additional 18 states before the reward is obtained when the ball hits the wall and smashes some bricks. How can we teach an agent to connect these three situations and generalize to similar situations? The answer is to use so-called Reinforcement Learning with a Neural Network, as shown in this tutorial.
## Q-Learning
One of the simplest ways of doing Reinforcement Learning is called Q-learning. Here we want to estimate so-called Q-values which are also called action-values, because they map a state of the game-environment to a numerical value for each possible action that the agent may take. The Q-values indicate which action is expected to result in the highest future reward, thus telling the agent which action to take.
Unfortunately we do not know what the Q-values are supposed to be, so we have to estimate them somehow. The Q-values are all initialized to zero and then updated repeatedly as new information is collected from the agent playing the game. When the agent scores a point then the Q-value must be updated with the new information.
There are different formulas for updating Q-values, but the simplest is to set the new Q-value to the reward that was observed, plus the maximum Q-value for the following state of the game. This gives the total reward that the agent can expect from the current game-state and onwards. Typically we also multiply the max Q-value for the following state by a so-called discount-factor slightly below 1. This causes more distant rewards to contribute less to the Q-value, thus making the agent favour rewards that are closer in time.
The formula for updating the Q-value is:
Q-value for state and action = reward + discount * max Q-value for next state
In academic papers, this is typically written with mathematical symbols like this:
$$
Q(s_{t},a_{t}) \leftarrow \underbrace{r_{t}}_{\rm reward} + \underbrace{\gamma}_{\rm discount} \cdot \underbrace{\max_{a}Q(s_{t+1}, a)}_{\rm estimate~of~future~rewards}
$$
Furthermore, when the agent loses a life, then we know that the future reward is zero because the agent is dead, so we set the Q-value for that state to zero.
### Simple Example
The images below demonstrate how Q-values are updated in a backwards sweep through the game-states that have previously been visited. In this simple example we assume all Q-values have been initialized to zero. The agent gets a reward of 1 point in the right-most image. This reward is then propagated backwards to the previous game-states, so when we see similar game-states in the future, we know that the given actions resulted in that reward.
The discounting is an exponentially decreasing function. This example uses a discount-factor of 0.97 so the Q-value for the 3rd image is about $0.885 \simeq 0.97^4$ because it is 4 states prior to the state that actually received the reward. Similarly for the other states. This example only shows one Q-value per state, but in reality there is one Q-value for each possible action in the state, and the Q-values are updated in a backwards-sweep using the formula above. This is shown in the next section.

### Detailed Example
This is a more detailed example showing the Q-values for two successive states of the game-environment and how to update them.

The Q-values for the possible actions have been estimated by a Neural Network. For the action NOOP in state *t* the Q-value is estimated to be 2.900, which is the highest Q-value for that state so the agent takes that action, i.e. the agent does not do anything between state *t* and *t+1* because NOOP means "No Operation".
In state *t+1* the agent scores 4 points, but this is limited to 1 point in this implementation so as to stabilize the training. The maximum Q-value for state *t+1* is 1.830 for the action RIGHTFIRE. So if we select that action and continue to select the actions proposed by the Q-values estimated by the Neural Network, then the discounted sum of all the future rewards is expected to be 1.830.
Now that we know the reward of taking the NOOP action from state *t* to *t+1*, we can update the Q-value to incorporate this new information. This uses the formula above:
$$
Q(state_{t},NOOP) \leftarrow \underbrace{r_{t}}_{\rm reward} + \underbrace{\gamma}_{\rm discount} \cdot \underbrace{\max_{a}Q(state_{t+1}, a)}_{\rm estimate~of~future~rewards} = 1.0 + 0.97 \cdot 1.830 \simeq 2.775
$$
The new Q-value is 2.775 which is slightly lower than the previous estimate of 2.900. This Neural Network has already been trained for 150 hours so it is quite good at estimating Q-values, but earlier during the training, the estimated Q-values would be more different.
The idea is to have the agent play many, many games and repeatedly update the estimates of the Q-values as more information about rewards and penalties becomes available. This will eventually lead to good estimates of the Q-values, provided the training is numerically stable, as discussed further below. By doing this, we create a connection between rewards and prior actions.
## Motion Trace
If we only use a single image from the game-environment then we cannot tell which direction the ball is moving. The typical solution is to use multiple consecutive images to represent the state of the game-environment.
This implementation uses another approach by processing the images from the game-environment in a motion-tracer that outputs two images as shown below. The left image is from the game-environment and the right image is the processed image, which shows traces of recent movements in the game-environment. In this case we can see that the ball is going downwards and has bounced off the right wall, and that the paddle has moved from the left to the right side of the screen.
Note that the motion-tracer has only been tested for Breakout and partially tested for Space Invaders, so it may not work for games with more complicated graphics such as Doom.

## Training Stability
We need a function approximator that can take a state of the game-environment as input and produce as output an estimate of the Q-values for that state. We will use a Convolutional Neural Network for this. Although they have achieved great fame in recent years, they are actually a quite old technologies with many problems - one of which is training stability. A significant part of the research for this tutorial was spent on tuning and stabilizing the training of the Neural Network.
To understand why training stability is a problem, consider the 3 images below which show the game-environment in 3 consecutive states. At state $t$ the agent is about to score a point, which happens in the following state $t+1$. Assuming all Q-values were zero prior to this, we should now set the Q-value for state $t+1$ to be 1.0 and it should be 0.97 for state $t$ if the discount-value is 0.97, according to the formula above for updating Q-values.

If we were to train a Neural Network to estimate the Q-values for the two states $t$ and $t+1$ with Q-values 0.97 and 1.0, respectively, then the Neural Network will most likely be unable to distinguish properly between the images of these two states. As a result the Neural Network will also estimate a Q-value near 1.0 for state $t+2$ because the images are so similar. But this is clearly wrong because the Q-values for state $t+2$ should be zero as we do not know anything about future rewards at this point, and that is what the Q-values are supposed to estimate.
If this is continued and the Neural Network is trained after every new game-state is observed, then it will quickly cause the estimated Q-values to explode. This is an artifact of training Neural Networks which must have sufficiently large and diverse training-sets. For this reason we will use a so-called Replay Memory so we can gather a large number of game-states and shuffle them during training of the Neural Network.
## Flowchart
This flowchart shows roughly how Reinforcement Learning is implemented in this tutorial. There are two main loops which are run sequentially until the Neural Network is sufficiently accurate at estimating Q-values.
The first loop is for playing the game and recording data. This uses the Neural Network to estimate Q-values from a game-state. It then stores the game-state along with the corresponding Q-values and reward/penalty in the Replay Memory for later use.
The other loop is activated when the Replay Memory is sufficiently full. First it makes a full backwards sweep through the Replay Memory to update the Q-values with the new rewards and penalties that have been observed. Then it performs an optimization run so as to train the Neural Network to better estimate these updated Q-values.
There are many more details in the implementation, such as decreasing the learning-rate and increasing the fraction of the Replay Memory being used during training, but this flowchart shows the main ideas.

## Neural Network Architecture
The Neural Network used in this implementation has 3 convolutional layers, all of which have filter-size 3x3. The layers have 16, 32, and 64 output channels, respectively. The stride is 2 in the first two convolutional layers and 1 in the last layer.
Following the 3 convolutional layers there are 4 fully-connected layers each with 1024 units and ReLU-activation. Then there is a single fully-connected layer with linear activation used as the output of the Neural Network.
This architecture is different from those typically used in research papers from DeepMind and others. They often have large convolutional filter-sizes of 8x8 and 4x4 with high stride-values. This causes more aggressive down-sampling of the game-state images. They also typically have only a single fully-connected layer with 256 or 512 ReLU units.
During the research for this tutorial, it was found that smaller filter-sizes and strides in the convolutional layers, combined with several fully-connected layers having more units, were necessary in order to have sufficiently accurate Q-values. The Neural Network architectures originally used by DeepMind appear to distort the Q-values quite significantly. A reason that their approach still worked, is possibly due to their use of a very large Replay Memory with 1 million states, and that the Neural Network did one mini-batch of training for each step of the game-environment, and some other tricks.
The architecture used here is probably excessive but it takes several days of training to test each architecture, so it is left as an exercise for the reader to try and find a smaller Neural Network architecture that still performs well.
## Installation
The [documentation](https://github.com/openai/gym) for OpenAI Gym currently suggests that you need to build it in order to install it. But if you just want to install the Atari games, then you only need to install a single pip-package by typing the following commands in a terminal.
- conda create --name tf-gym --clone tf
- source activate tf-gym
- pip install gym[atari]
This assumes you already have an Anaconda environment named `tf` which has TensorFlow installed, it will then be cloned to another environment named `tf-gym` where OpenAI Gym is also installed. This allows you to easily switch between your normal TensorFlow environment and another one which also contains OpenAI Gym.
You can also have two environments named `tf-gpu` and `tf-gpu-gym` for the GPU versions of TensorFlow.
## Imports
```
%matplotlib inline
import matplotlib.pyplot as plt
import tensorflow as tf
import gym
import numpy as np
import math
```
The main source-code for Reinforcement Learning is located in the following module:
```
import reinforcement_learning as rl
```
This was developed using Python 3.6.0 (Anaconda) with package versions:
```
# TensorFlow
tf.__version__
# OpenAI Gym
gym.__version__
```
## Game Environment
This is the name of the game-environment that we want to use in OpenAI Gym.
```
env_name = 'Breakout-v0'
# env_name = 'SpaceInvaders-v0'
```
This is the base-directory for the TensorFlow checkpoints as well as various log-files.
```
rl.checkpoint_base_dir = 'checkpoints_tutorial16/'
```
Once the base-dir has been set, you need to call this function to set all the paths that will be used. This will also create the checkpoint-dir if it does not already exist.
```
rl.update_paths(env_name=env_name)
```
## Download Pre-Trained Model
You can download a TensorFlow checkpoint which holds all the pre-trained variables for the Neural Network. Two checkpoints are provided, one for Breakout and one for Space Invaders. They were both trained for about 150 hours on a laptop with 2.6 GHz CPU and a GTX 1070 GPU.
#### COMPATIBILITY ISSUES
These TensorFlow checkpoints were developed with OpenAI gym v. 0.8.1 and atari-py v. 0.0.19 which had unused / redundant actions as noted above. There appears to have been a change in the gym API since then, as the unused actions are no longer present. This means the vectors with actions and Q-values now only contain 4 elements instead of the 6 shown here. This also means that the TensorFlow checkpoints cannot be used with newer versions of gym and atari-py, so in order to use these pre-trained checkpoints you need to install the older versions of gym and atari-py - or you can just train a new model yourself so you get a new TensorFlow checkpoint.
#### WARNING!
These checkpoints are 280-360 MB each. They are currently hosted on the webserver I use for [www.hvass-labs.org](www.hvass-labs.org) because it is awkward to automatically download large files on Google Drive. To lower the traffic on my webserver, this line has been commented out, so you have to activate it manually. You are welcome to download it, I just don't want it to download automatically for everyone who only wants to run this Notebook briefly.
```
# rl.maybe_download_checkpoint(env_name=env_name)
```
I believe the webserver is located in Denmark. If you are having problems downloading the files using the automatic function above, then you can try and download the files manually in a webbrowser or using `wget` or `curl`. Or you can download from Google Drive, where you will get an anti-virus warning that is awkward to bypass automatically:
* [Download Breakout Checkpoint from Google Drive](https://drive.google.com/uc?export=download&id=0B2aDiIly76ZvUjZTcXRuRFY0RjQ)
* [Download Space Invaders Checkpoint from Google Drive](https://drive.google.com/uc?export=download&id=0B2aDiIly76ZvWDR4TExwdmw1RVE)
You can use the checksum to ensure the downloaded files are complete:
* [SHA256 Checksum](http://www.hvass-labs.org/projects/tensorflow/tutorial16/sha256sum.txt)
## Create Agent
The Agent-class implements the main loop for playing the game, recording data and optimizing the Neural Network. We create an object-instance and need to set `training=True` because we want to use the replay-memory to record states and Q-values for plotting further below. We disable logging so this does not corrupt the logs from the actual training that was done previously. We can also set `render=True` but it will have no effect as long as `training==True`.
```
agent = rl.Agent(env_name=env_name,
training=True,
render=True,
use_logging=False)
```
The Neural Network is automatically instantiated by the Agent-class. We will create a direct reference for convenience.
```
model = agent.model
```
Similarly, the Agent-class also allocates the replay-memory when `training==True`. The replay-memory will require more than 3 GB of RAM, so it should only be allocated when needed. We will need the replay-memory in this Notebook to record the states and Q-values we observe, so they can be plotted further below.
```
replay_memory = agent.replay_memory
```
## Training
The agent's `run()` function is used to play the game. This uses the Neural Network to estimate Q-values and hence determine the agent's actions. If `training==True` then it will also gather states and Q-values in the replay-memory and train the Neural Network when the replay-memory is sufficiently full. You can set `num_episodes=None` if you want an infinite loop that you would stop manually with `ctrl-c`. In this case we just set `num_episodes=1` because we are not actually interested in training the Neural Network any further, we merely want to collect some states and Q-values in the replay-memory so we can plot them below.
```
agent.run(num_episodes=1)
```
In training-mode, this function will output a line for each episode. The first counter is for the number of episodes that have been processed. The second counter is for the number of states that have been processed. These two counters are stored in the TensorFlow checkpoint along with the weights of the Neural Network, so you can restart the training e.g. if you only have one computer and need to train during the night.
Note that the number of episodes is almost 90k. It is impractical to print that many lines in this Notebook, so the training is better done in a terminal window by running the following commands:
```
source activate tf-gpu-gym # Activate your Python environment with TF and Gym.
python reinforcement-learning.py --env Breakout-v0 --training
```
## Training Progress
Data is being logged during training so we can plot the progress afterwards. The reward for each episode and a running mean of the last 30 episodes are logged to file. Basic statistics for the Q-values in the replay-memory are also logged to file before each optimization run.
This could be logged using TensorFlow and TensorBoard, but they were designed for logging variables of the TensorFlow graph and data that flows through the graph. In this case the data we want logged does not reside in the graph, so it becomes a bit awkward to use TensorFlow to log this data.
We have therefore implemented a few small classes that can write and read these logs.
```
log_q_values = rl.LogQValues()
log_reward = rl.LogReward()
```
We can now read the logs from file:
```
log_q_values.read()
log_reward.read()
```
### Training Progress: Reward
This plot shows the reward for each episode during training, as well as the running mean of the last 30 episodes. Note how the reward varies greatly from one episode to the next, so it is difficult to say from this plot alone whether the agent is really improving during the training, although the running mean does appear to trend upwards slightly.
```
plt.plot(log_reward.count_states, log_reward.episode, label='Episode Reward')
plt.plot(log_reward.count_states, log_reward.mean, label='Mean of 30 episodes')
plt.xlabel('State-Count for Game Environment')
plt.legend()
plt.show()
```
### Training Progress: Q-Values
The following plot shows the mean Q-values from the replay-memory prior to each run of the optimizer for the Neural Network. Note how the mean Q-values increase rapidly in the beginning and then they increase fairly steadily for 40 million states, after which they still trend upwards but somewhat more irregularly.
The fast improvement in the beginning is probably due to (1) the use of a smaller replay-memory early in training so the Neural Network is optimized more often and the new information is used faster, (2) the backwards-sweeping of the replay-memory so the rewards are used to update the Q-values for many of the states, instead of just updating the Q-values for a single state, and (3) the replay-memory is balanced so at least half of each mini-batch contains states whose Q-values have high estimation-errors for the Neural Network.
The [original paper from DeepMind](https://arxiv.org/abs/1312.5602) showed much slower progress in the first phase of training, see Figure 2 in that paper but note that the Q-values are not directly comparable, possibly because they used a higher discount factor of 0.99 while we only used 0.97 here.
```
plt.plot(log_q_values.count_states, log_q_values.mean, label='Q-Value Mean')
plt.xlabel('State-Count for Game Environment')
plt.legend()
plt.show()
```
## Testing
When the agent and Neural Network is being trained, the so-called epsilon-probability is typically decreased from 1.0 to 0.1 over a large number of steps, after which the probability is held fixed at 0.1. This means the probability is 0.1 or 10% that the agent will select a random action in each step, otherwise it will select the action that has the highest Q-value. This is known as the epsilon-greedy policy. The choice of 0.1 for the epsilon-probability is a compromise between taking the actions that are already known to be good, versus exploring new actions that might lead to even higher rewards or might lead to death of the agent.
During testing it is common to lower the epsilon-probability even further. We have set it to 0.01 as shown here:
```
agent.epsilon_greedy.epsilon_testing
```
We will now instruct the agent that it should no longer perform training by setting this boolean:
```
agent.training = False
```
We also reset the previous episode rewards.
```
agent.reset_episode_rewards()
```
We can render the game-environment to screen so we can see the agent playing the game, by setting this boolean:
```
agent.render = True
```
We can now run a single episode by calling the `run()` function again. This should open a new window that shows the game being played by the agent. At the time of this writing, it was not possible to resize this tiny window, and the developers at OpenAI did not seem to care about this feature which should obviously be there.
```
agent.run(num_episodes=1)
```
### Mean Reward
The game-play is slightly random, both with regard to selecting actions using the epsilon-greedy policy, but also because the OpenAI Gym environment will repeat any action between 2-4 times, with the number chosen at random. So the reward of one episode is not an accurate estimate of the reward that can be expected in general from this agent.
We need to run 30 or even 50 episodes to get a more accurate estimate of the reward that can be expected.
We will first reset the previous episode rewards.
```
agent.reset_episode_rewards()
```
We disable the screen-rendering so the game-environment runs much faster.
```
agent.render = False
```
We can now run 30 episodes. This records the rewards for each episode. It might have been a good idea to disable the output so it does not print all these lines - you can do this as an exercise.
```
agent.run(num_episodes=30)
```
We can now print some statistics for the episode rewards, which vary greatly from one episode to the next.
```
rewards = agent.episode_rewards
print("Rewards for {0} episodes:".format(len(rewards)))
print("- Min: ", np.min(rewards))
print("- Mean: ", np.mean(rewards))
print("- Max: ", np.max(rewards))
print("- Stdev: ", np.std(rewards))
```
We can also plot a histogram with the episode rewards.
```
_ = plt.hist(rewards, bins=30)
```
## Example States
We can plot examples of states from the game-environment and the Q-values that are estimated by the Neural Network.
This helper-function prints the Q-values for a given index in the replay-memory.
```
def print_q_values(idx):
"""Print Q-values and actions from the replay-memory at the given index."""
# Get the Q-values and action from the replay-memory.
q_values = replay_memory.q_values[idx]
action = replay_memory.actions[idx]
print("Action: Q-Value:")
print("====================")
# Print all the actions and their Q-values.
for i, q_value in enumerate(q_values):
# Used to display which action was taken.
if i == action:
action_taken = "(Action Taken)"
else:
action_taken = ""
# Text-name of the action.
action_name = agent.get_action_name(i)
print("{0:12}{1:.3f} {2}".format(action_name, q_value,
action_taken))
# Newline.
print()
```
This helper-function plots a state from the replay-memory and optionally prints the Q-values.
```
def plot_state(idx, print_q=True):
"""Plot the state in the replay-memory with the given index."""
# Get the state from the replay-memory.
state = replay_memory.states[idx]
# Create figure with a grid of sub-plots.
fig, axes = plt.subplots(1, 2)
# Plot the image from the game-environment.
ax = axes.flat[0]
ax.imshow(state[:, :, 0], vmin=0, vmax=255,
interpolation='lanczos', cmap='gray')
# Plot the motion-trace.
ax = axes.flat[1]
ax.imshow(state[:, :, 1], vmin=0, vmax=255,
interpolation='lanczos', cmap='gray')
# This is necessary if we show more than one plot in a single Notebook cell.
plt.show()
# Print the Q-values.
if print_q:
print_q_values(idx=idx)
```
The replay-memory has room for 200k states but it is only partially full from the above call to `agent.run(num_episodes=1)`. This is how many states are actually used.
```
num_used = replay_memory.num_used
num_used
```
Get the Q-values from the replay-memory that are actually used.
```
q_values = replay_memory.q_values[0:num_used, :]
```
For each state, calculate the min / max Q-values and their difference. This will be used to lookup interesting states in the following sections.
```
q_values_min = q_values.min(axis=1)
q_values_max = q_values.max(axis=1)
q_values_dif = q_values_max - q_values_min
```
### Example States: Highest Reward
This example shows the states surrounding the state with the highest reward.
During the training we limit the rewards to the range [-1, 1] so this basically just gets the first state that has a reward of 1.
```
idx = np.argmax(replay_memory.rewards)
idx
```
This state is where the ball hits the wall so the agent scores a point.
We can show the surrounding states leading up to and following this state. Note how the Q-values are very close for the different actions, because at this point it really does not matter what the agent does as the reward is already guaranteed. But note how the Q-values decrease significantly after the ball has hit the wall and a point has been scored.
Also note that the agent uses the Epsilon-greedy policy for taking actions, so there is a small probability that a random action is taken instead of the action with the highest Q-value.
```
for i in range(-5, 3):
plot_state(idx=idx+i)
```
### Example: Highest Q-Value
This example shows the states surrounding the one with the highest Q-values. This means that the agent has high expectation that several points will be scored in the following steps. Note that the Q-values decrease significantly after the points have been scored.
```
idx = np.argmax(q_values_max)
idx
for i in range(0, 5):
plot_state(idx=idx+i)
```
### Example: Loss of Life
This example shows the states leading up to a loss of life for the agent.
```
idx = np.argmax(replay_memory.end_life)
idx
for i in range(-10, 0):
plot_state(idx=idx+i)
```
### Example: Greatest Difference in Q-Values
This example shows the state where there is the greatest difference in Q-values, which means that the agent believes one action will be much more beneficial than another. But because the agent uses the Epsilon-greedy policy, it sometimes selects a random action instead.
```
idx = np.argmax(q_values_dif)
idx
for i in range(0, 5):
plot_state(idx=idx+i)
```
### Example: Smallest Difference in Q-Values
This example shows the state where there is the smallest difference in Q-values, which means that the agent believes it does not really matter which action it selects, as they all have roughly the same expectations for future rewards.
The Neural Network estimates these Q-values and they are not precise. The differences in Q-values may be so small that they fall within the error-range of the estimates.
```
idx = np.argmin(q_values_dif)
idx
for i in range(0, 5):
plot_state(idx=idx+i)
```
## Output of Convolutional Layers
The outputs of the convolutional layers can be plotted so we can see how the images from the game-environment are being processed by the Neural Network.
This is the helper-function for plotting the output of the convolutional layer with the given name, when inputting the given state from the replay-memory.
```
def plot_layer_output(model, layer_name, state_index, inverse_cmap=False):
"""
Plot the output of a convolutional layer.
:param model: An instance of the NeuralNetwork-class.
:param layer_name: Name of the convolutional layer.
:param state_index: Index into the replay-memory for a state that
will be input to the Neural Network.
:param inverse_cmap: Boolean whether to inverse the color-map.
"""
# Get the given state-array from the replay-memory.
state = replay_memory.states[state_index]
# Get the output tensor for the given layer inside the TensorFlow graph.
# This is not the value-contents but merely a reference to the tensor.
layer_tensor = model.get_layer_tensor(layer_name=layer_name)
# Get the actual value of the tensor by feeding the state-data
# to the TensorFlow graph and calculating the value of the tensor.
values = model.get_tensor_value(tensor=layer_tensor, state=state)
# Number of image channels output by the convolutional layer.
num_images = values.shape[3]
# Number of grid-cells to plot.
# Rounded-up, square-root of the number of filters.
num_grids = math.ceil(math.sqrt(num_images))
# Create figure with a grid of sub-plots.
fig, axes = plt.subplots(num_grids, num_grids, figsize=(10, 10))
print("Dim. of each image:", values.shape)
if inverse_cmap:
cmap = 'gray_r'
else:
cmap = 'gray'
# Plot the outputs of all the channels in the conv-layer.
for i, ax in enumerate(axes.flat):
# Only plot the valid image-channels.
if i < num_images:
# Get the image for the i'th output channel.
img = values[0, :, :, i]
# Plot image.
ax.imshow(img, interpolation='nearest', cmap=cmap)
# Remove ticks from the plot.
ax.set_xticks([])
ax.set_yticks([])
# Ensure the plot is shown correctly with multiple plots
# in a single Notebook cell.
plt.show()
```
### Game State
This is the state that is being input to the Neural Network. The image on the left is the last image from the game-environment. The image on the right is the processed motion-trace that shows the trajectories of objects in the game-environment.
```
idx = np.argmax(q_values_max)
plot_state(idx=idx, print_q=False)
```
### Output of Convolutional Layer 1
This shows the images that are output by the 1st convolutional layer, when inputting the above state to the Neural Network. There are 16 output channels of this convolutional layer.
Note that you can invert the colors by setting `inverse_cmap=True` in the parameters to this function.
```
plot_layer_output(model=model, layer_name='layer_conv1', state_index=idx, inverse_cmap=False)
```
### Output of Convolutional Layer 2
These are the images output by the 2nd convolutional layer, when inputting the above state to the Neural Network. There are 32 output channels of this convolutional layer.
```
plot_layer_output(model=model, layer_name='layer_conv2', state_index=idx, inverse_cmap=False)
```
### Output of Convolutional Layer 3
These are the images output by the 3rd convolutional layer, when inputting the above state to the Neural Network. There are 64 output channels of this convolutional layer.
All these images are flattened to a one-dimensional array (or tensor) which is then used as the input to a fully-connected layer in the Neural Network.
During the training-process, the Neural Network has learnt what convolutional filters to apply to the images from the game-environment so as to produce these images, because they have proven to be useful when estimating Q-values.
Can you see what it is that the Neural Network has learned to detect in these images?
```
plot_layer_output(model=model, layer_name='layer_conv3', state_index=idx, inverse_cmap=False)
```
## Weights for Convolutional Layers
We can also plot the weights of the convolutional layers in the Neural Network. These are the weights that are being optimized so as to improve the ability of the Neural Network to estimate Q-values. Tutorial #02 explains in greater detail what convolutional weights are.
There are also weights for the fully-connected layers but they are not shown here.
This is the helper-function for plotting the weights of a convoluational layer.
```
def plot_conv_weights(model, layer_name, input_channel=0):
"""
Plot the weights for a convolutional layer.
:param model: An instance of the NeuralNetwork-class.
:param layer_name: Name of the convolutional layer.
:param input_channel: Plot the weights for this input-channel.
"""
# Get the variable for the weights of the given layer.
# This is a reference to the variable inside TensorFlow,
# not its actual value.
weights_variable = model.get_weights_variable(layer_name=layer_name)
# Retrieve the values of the weight-variable from TensorFlow.
# The format of this 4-dim tensor is determined by the
# TensorFlow API. See Tutorial #02 for more details.
w = model.get_variable_value(variable=weights_variable)
# Get the weights for the given input-channel.
w_channel = w[:, :, input_channel, :]
# Number of output-channels for the conv. layer.
num_output_channels = w_channel.shape[2]
# Get the lowest and highest values for the weights.
# This is used to correct the colour intensity across
# the images so they can be compared with each other.
w_min = np.min(w_channel)
w_max = np.max(w_channel)
# This is used to center the colour intensity at zero.
abs_max = max(abs(w_min), abs(w_max))
# Print statistics for the weights.
print("Min: {0:.5f}, Max: {1:.5f}".format(w_min, w_max))
print("Mean: {0:.5f}, Stdev: {1:.5f}".format(w_channel.mean(),
w_channel.std()))
# Number of grids to plot.
# Rounded-up, square-root of the number of output-channels.
num_grids = math.ceil(math.sqrt(num_output_channels))
# Create figure with a grid of sub-plots.
fig, axes = plt.subplots(num_grids, num_grids)
# Plot all the filter-weights.
for i, ax in enumerate(axes.flat):
# Only plot the valid filter-weights.
if i < num_output_channels:
# Get the weights for the i'th filter of this input-channel.
img = w_channel[:, :, i]
# Plot image.
ax.imshow(img, vmin=-abs_max, vmax=abs_max,
interpolation='nearest', cmap='seismic')
# Remove ticks from the plot.
ax.set_xticks([])
ax.set_yticks([])
# Ensure the plot is shown correctly with multiple plots
# in a single Notebook cell.
plt.show()
```
### Weights for Convolutional Layer 1
These are the weights of the first convolutional layer of the Neural Network, with respect to the first input channel of the state. That is, these are the weights that are used on the image from the game-environment. Some basic statistics are also shown.
Note how the weights are more negative (blue) than positive (red). It is unclear why this happens as these weights are found through optimization. It is apparently beneficial for the following layers to have this processing with more negative weights in the first convolutional layer.
```
plot_conv_weights(model=model, layer_name='layer_conv1', input_channel=0)
```
We can also plot the convolutional weights for the second input channel, that is, the motion-trace of the game-environment. Once again we see that the negative weights (blue) have a much greater magnitude than the positive weights (red).
```
plot_conv_weights(model=model, layer_name='layer_conv1', input_channel=1)
```
### Weights for Convolutional Layer 2
These are the weights of the 2nd convolutional layer in the Neural Network. There are 16 input channels and 32 output channels of this layer. You can change the number for the input-channel to see the associated weights.
Note how the weights are more balanced between positive (red) and negative (blue) compared to the weights for the 1st convolutional layer above.
```
plot_conv_weights(model=model, layer_name='layer_conv2', input_channel=0)
```
### Weights for Convolutional Layer 3
These are the weights of the 3rd convolutional layer in the Neural Network. There are 32 input channels and 64 output channels of this layer. You can change the number for the input-channel to see the associated weights.
Note again how the weights are more balanced between positive (red) and negative (blue) compared to the weights for the 1st convolutional layer above.
```
plot_conv_weights(model=model, layer_name='layer_conv3', input_channel=0)
```
## Discussion
We trained an agent to play old Atari games quite well using Reinforcement Learning. Recent improvements to the training algorithm have improved the performance significantly. But is this true human-like intelligence? The answer is clearly NO!
Reinforcement Learning in its current form is a crude numerical algorithm for connecting visual images, actions, rewards and penalties when there is a time-lag between the signals. The learning is based on trial-and-error and cannot do logical reasoning like a human. The agent has no sense of "self" while a human has an understanding of what part of the game-environment it is controlling, so a human can reason logically like this: "(A) I control the paddle, and (B) I must avoid dying which happens when the ball flies past the paddle, so (C) I must move the paddle to hit the ball, and (D) this automatically scores points when the ball smashes bricks in the wall". A human would first learn these basic logical rules of the game - and then try and refine the eye-hand coordination to play the game better. Reinforcement Learning has no real comprehension of what is going on in the game and merely works on improving the eye-hand coordination until it gets lucky and does the right thing to score more points.
Furthermore, the training of the Reinforcement Learning algorithm required almost 150 hours of computation which played the game at high speeds. If the game was played at normal real-time speeds then it would have taken more than 1700 hours to train the agent, which is more than 70 days and nights.
Logical reasoning would allow for much faster learning than Reinforcement Learning, and it would be able to solve much more complicated problems than simple eye-hand coordination. I am skeptical if someone will be able to create true human-like intelligence from Reinforcement Learning algorithms.
Does that mean Reinforcement Learning is completely worthless? No, it has real-world applications that currently cannot be solved by other methods.
Another point of criticism is the use of Neural Networks. The majority of the research in Reinforcement Learning is actually spent on trying to stabilize the training of the Neural Network using various tricks. This is a waste of research time and strongly indicates that Neural Networks may not be a very good Machine Learning model compared to the human brain.
## Exercises & Research Ideas
Below are suggestions for exercises and experiments that may help improve your skills with TensorFlow and Reinforcement Learning. Some of these ideas can easily be extended into full research problems that would help the community if you can solve them.
You should keep a log of your experiments, describing for each experiment the settings you tried and the results. You should also save the source-code and checkpoints / log-files.
It takes so much time to run these experiments, so please share your results with the rest of the community. Even if an experiment failed to produce anything useful, it will be helpful to others so they know not to redo the same experiment.
[Thread on GitHub for discussing these experiments](https://github.com/Hvass-Labs/TensorFlow-Tutorials/issues/32)
You may want to backup this Notebook and the other files before making any changes.
You may find it helpful to add more command-line parameters to `reinforcement_learning.py` so you don't have to edit the source-code for testing other parameters.
* Change the epsilon-probability during testing to e.g. 0.001 or 0.05. Which gives the best results? Could you use this value during training? Why/not?
* Continue training the agent for the Breakout game using the downloaded checkpoint. Does the agent get better or worse the more you train it? Why? (You should run it in a terminal window as described above.)
* Try and change the game-environment to Space Invaders and re-run this Notebook. The checkpoint can be downloaded automatically. It was trained for about 150 hours, which is roughly the same as for Breakout, but note that it has processed far fewer states. The reason is that the hyper-parameters such as the learning-rate were tuned for Breakout. Can you make some kind of adaptive learning-rate that would work better for both Breakout and Space Invaders? What about the other hyper-parameters? What about other games?
* Try different architectures for the Neural Network. You will need to restart the training because the checkpoints cannot be reused for other architectures. You will need to train the agent for several days with each new architecture so as to properly assess its performance.
* The replay-memory throws away all data after optimization of the Neural Network. Can you make it reuse the data somehow? The ReplayMemory-class has the function `estimate_all_q_values()` which may be helpful.
* The reward is limited to -1 and 1 in the function `ReplayMemory.add()` so as to stabilize the training. This means the agent cannot distinguish between small and large rewards. Can you use batch normalization to fix this problem, so you can use the actual reward values?
* Can you improve the training by adding L2-regularization or dropout?
* Try using other optimizers for the Neural Network. Does it help with the training speed or stability?
* Let the agent take up to 30 random actions at the beginning of each new episode. This is used in some research papers to further randomize the game-environment, so the agent cannot memorize the first sequence of actions.
* Try and save the game at regular intervals. If the agent dies, then you can reload the last saved game. Would this help training the agent faster and better, because it does not need to play the game from the beginning?
* There are some invalid actions available to the agent in OpenAI Gym. Does it improve the training if you only allow the valid actions from the game-environment?
* Does the MotionTracer work for other games? Can you improve on the MotionTracer?
* Try and use the last 4 image-frames from the game instead of the MotionTracer.
* Try larger and smaller sizes for the replay memory.
* Try larger and smaller discount rates for updating the Q-values.
* If you look closely in the states and actions that are display above, you will note that the agent has sometimes taken actions that do not correspond to the movement of the paddle. For example, the action might be LEFT but the paddle has either not moved at all, or it has moved right instead. Is this a bug in the source-code for this tutorial, or is it a bug in OpenAI Gym, or is it a bug in the underlying Atari Learning Environment? Does it matter?
## License (MIT)
Copyright (c) 2017 by [Magnus Erik Hvass Pedersen](http://www.hvass-labs.org/)
Permission is hereby granted, free of charge, to any person obtaining a copy of this software and associated documentation files (the "Software"), to deal in the Software without restriction, including without limitation the rights to use, copy, modify, merge, publish, distribute, sublicense, and/or sell copies of the Software, and to permit persons to whom the Software is furnished to do so, subject to the following conditions:
The above copyright notice and this permission notice shall be included in all copies or substantial portions of the Software.
THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY, FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE SOFTWARE.
| github_jupyter |
This notebooks serves as a demo of `TorchModel` capabilities.
Aside from that, it is also used as an interactive test.
```
%load_ext autoreload
%autoreload 2
import os
import sys
import warnings
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import torch
import torch.nn as nn
sys.path.insert(0, "../../..")
from batchflow import *
from batchflow.opensets import MNIST
from batchflow.models.torch import *
from batchflow.models.torch.layers import *
from batchflow.models.torch.callbacks import ReduceLROnPlateau, EarlyStopping
```
# Setup: global parameters and functions
```
# Global parameters
mnist = MNIST(bar=False)
BAR = True
PLOT = False
IMAGE_SHAPE = (1, 28, 28)
BATCH_SIZE = 16
N_ITERS = 10
N_ITERS_LARGE = 100
if __name__ == '__main__':
MICROBATCH = None
DEVICE = 'gpu:0'
BAR = 't'
PLOT = True
def get_classification_config(model_class, config):
default_config = {
# Shapes info. Can be commented
'inputs_shapes': IMAGE_SHAPE,
'classes': 10,
'loss': 'ce',
'microbatch_size': MICROBATCH,
'device': DEVICE,
}
if 'inputs_shapes' in config and isinstance(config['inputs_shapes'], list):
inputs = [B.images for item in config['inputs_shapes']]
else:
inputs = B.images
pipeline_config = {
'model': model_class,
'model_config': {**default_config, **config},
'inputs': inputs,
'targets': B.labels,
'gather': {'metrics_class' : 'classification',
'fmt' : 'logits',
'axis' : 1,
'targets' : B.labels},
'evaluate': 'accuracy',
}
return pipeline_config
def get_segmentation_config(model_class, config):
default_config = {
# Shapes info. Can be commented
'inputs_shapes': IMAGE_SHAPE,
'targets_shapes': IMAGE_SHAPE,
'loss': 'mse',
'microbatch': MICROBATCH,
'device': DEVICE,
}
if 'inputs_shapes' in config and isinstance(config['inputs_shapes'], list):
inputs = [B.images for item in config['inputs_shapes']]
else:
inputs = B.images
pipeline_config = {
'model': model_class,
'model_config': {**default_config, **config},
'inputs': inputs,
'targets': B.images,
'gather': {'metrics_class' : 'segmentation',
'fmt' : 'proba',
'axis' : None,
'targets' : B.images},
'evaluate': 'jaccard',
}
return pipeline_config
def get_pipeline(pipeline_config):
""" Pipeline config must contain 'model', 'model_config', 'feed_dict' keys. """
pipeline = (Pipeline(config=pipeline_config)
.init_variable('loss_history', [])
.to_array(channels='first', dtype='float32')
.multiply(multiplier=1/255., preserve_type=False)
.init_model(name='MODEL', model_class=C('model'), config=C('model_config'))
.train_model('MODEL',
inputs=pipeline_config['inputs'],
targets=pipeline_config['targets'],
outputs='loss',
save_to=V('loss_history', mode='a'))
)
return pipeline
def run(task, model_class, config, description, batch_size=BATCH_SIZE, n_iters=N_ITERS, **kwargs):
if task == 'classification':
pipeline_config = get_classification_config(model_class, config)
elif task == 'segmentation':
pipeline_config = get_segmentation_config(model_class, config)
train_pipeline = get_pipeline(pipeline_config) << mnist.train
_ = train_pipeline.run(batch_size, n_iters=n_iters,
bar={'bar': BAR, 'monitors': 'loss_history'},
**kwargs)
print(f'{task} "{description}" is done! Number of parameters in the model: {train_pipeline.model.num_parameters:,}')
return train_pipeline
def show_some_results(ppl, task, size=10):
batch_ind = np.random.randint(len(ppl.v('targets')))
image_ind = np.random.choice(len(ppl.v('targets')[batch_ind]), size=size, replace=False)
true = ppl.v('targets')[batch_ind]
pred = ppl.v('predictions')[batch_ind]
if task == 'classification':
print(pd.DataFrame({'true': true[image_ind],
'pred': np.argmax(pred[image_ind], axis=1)}).to_string(index=False))
elif task == 'segmentation':
pass # for the sake of parsing by notebooks_test.py
fig, ax = plt.subplots(2, size, figsize=(10, 5))
[axi.set_axis_off() for axi in ax.ravel()]
for plot_num, image_num in enumerate(image_ind):
ax[0][plot_num].imshow(true[image_num][0], cmap='gray', vmin=0, vmax=1)
ax[1][plot_num].imshow(pred[image_num][0], cmap='gray', vmin=0, vmax=1)
def test(pipeline, show_results=PLOT, batch_size=64, n_epochs=1, drop_last=False):
test_pipeline = (mnist.test.p
.import_model('MODEL', pipeline)
.init_variable('targets', default=[])
.init_variable('predictions', default=[])
.init_variable('metrics', default=[])
.to_array(channels='first', dtype='float32')
.multiply(multiplier=1/255., preserve_type=False)
.update(V('targets', mode='a'), pipeline.config['targets'])
.predict_model('MODEL',
inputs=pipeline.config['inputs'],
outputs='predictions',
save_to=V('predictions', mode='a'))
.gather_metrics(**pipeline.config['gather'], predictions=V.predictions[-1],
save_to=V('metrics', mode='a'))
.run(batch_size, shuffle=False, n_epochs=n_epochs, drop_last=drop_last, bar=BAR)
)
if show_results:
show_some_results(test_pipeline, pipeline.config['gather/metrics_class'])
metrics = test_pipeline.get_variable('metrics')
to_evaluate = pipeline.config['evaluate']
evaluated = np.mean([m.evaluate(to_evaluate) for m in metrics])
print(f'{to_evaluate} metrics is: {evaluated:.3}')
return test_pipeline
```
# Tester
Test the simplest possible model and record timings
```
config = {'initial_block': {'layout': 'Vf', 'features': 10}}
ppl = run('classification', TorchModel, config, 'simple fc', n_iters=2, batch_size=2)
%%time
test(ppl, show_results=False, batch_size=BATCH_SIZE, drop_last=True, n_epochs=5);
```
# Classification
```
config = {
'initial_block': {'layout': 'fa'*2,
'features': [64, 128],},
'body': {'layout': 'fa'*2,
'features': [256, 512]},
'head': {'layout': 'faf',
'features': [600, 10]},
}
ppl = run('classification', TorchModel, config, 'simple fc', n_iters=200, batch_size=64)
test(ppl, show_results=False);
config = {
'body': {'type': 'encoder',
'output_type': 'tensor',
'num_stages': 3,
'blocks/channels': '2 * same'},
'head': {'layout': 'f'}
}
ppl = run('classification', TorchModel, config, 'encoder')
# Example with multiple inputs
config = {
'inputs_shapes': [IMAGE_SHAPE, IMAGE_SHAPE],
'initial_block': {'type': 'wrapper',
'input_type': 'list',
'input_index': slice(None),
'module': Combine(op='concat', force_resize=False),},
'body': {'type': 'encoder',
'num_stages': 3,
'output_type': 'tensor',
'blocks/channels': '2 * same'},
'head': {'layout': 'faf', 'features': [50, 10]}
}
ppl = run('classification', TorchModel, config, 'duo input')
ppl.model.repr(1)
```
# Classification: named networks
```
ppl = run('classification', VGG16, {}, 'vgg16', n_iters=100, batch_size=128)
test(ppl, show_results=False);
ppl = run('classification', ResNet18, {}, 'resnet18', n_iters=100, batch_size=128)
test(ppl, show_results=False);
ppl = run('classification', SEResNeXt18, {}, 'SE-ResNeXt18', n_iters=100, batch_size=128)
test(ppl, show_results=False);
ppl = run('classification', DenseNetS, {}, 'DenseNetS', n_iters=100, batch_size=128)
test(ppl, show_results=False);
ppl = run('classification', EfficientNetB0, {}, 'EfficientNetB0', n_iters=100, batch_size=128)
test(ppl, show_results=False);
config = {
'initial_block': {'layout': 'cna', 'channels': 3,
'kernel_size': 5, 'stride': 1, 'padding': 'same'},
'body': {'num_stages': 4,
'blocks': {'n_reps': [1, 1, 2, 1],
'bottleneck': False,
'channels': '2 * same',
'attention': 'se'}
}
}
ppl = run('classification', ResNet, config, 'resnet with config', n_iters=50, batch_size=128)
test(ppl, show_results=False);
# reusing encoder from model from the previous cell
config = {
'initial_block': {'type': 'wrapper', 'module': ppl.model.model.initial_block},
'body': {'type': 'wrapper', 'module': ppl.model.model.body},
'head' : {'type': 'wrapper', 'module': ppl.model.model.head},
}
ppl = run('classification', TorchModel, config, 'reused encoder', n_iters=50, batch_size=32)
test(ppl, show_results=False);
```
# Classification: imported models
```
import torchvision.models as models
resnet18 = models.resnet18(pretrained=True)
resnet18.fc = torch.nn.Identity()
config = {
'initial_block': {'layout': 'cna',
'channels': 3},
'body': {'type': 'wrapper', 'module': resnet18},
'head': {'layout': 'Dnfaf',
'features': [50, 10],
'dropout_rate': 0.3,
'multisample': 0.3},
}
ppl = run('classification', TorchModel, config, 'torchvision resnet', n_iters=100, batch_size=128)
test(ppl, show_results=False);
config = {
'initial_block': {
'layout': 'cna',
'channels': 3,
'output_list': True
},
'body': {
'type': 'timm',
'output_type': 'tensor',
'path': 'resnet34d',
'pretrained': True,
},
'head': {'layout': 'Dnfaf',
'features': [50, 10],
'dropout_rate': 0.3,
'multisample': 0.3},
}
ppl = run('classification', TorchModel, config, 'TIMM-resnet34', n_iters=100, batch_size=128)
# ppl.model.model.body.config
test(ppl, show_results=False);
config = {
'trainable': ['initial_block', 'head'],
**config
}
ppl = run('classification', TorchModel, config, 'TIMM-resnet34 finetune', n_iters=100, batch_size=128)
# ppl.model.model.body.config
test(ppl, show_results=False);
config = {
'initial_block': {
'layout': 'cna',
'channels': 3,
'output_list': True
},
'body': {
'type': 'hugging-face',
'output_type': 'tensor',
'path': 'facebook/convnext-tiny-224',
'num_stages': 2,
'depths': [2, 2],
'num_channels': 3, # number of input channels
'patch_size': 2, # ~stride in the beginning
'hidden_sizes': [48, 64], # ~channels
},
'head': {'layout': 'Dnfaf',
'features': [50, 10],
'dropout_rate': 0.3,
'multisample': 0.3},
}
ppl = run('classification', TorchModel, config, 'HF-ConvNext', n_iters=100, batch_size=128)
# ppl.model.model.body.config
test(ppl, show_results=False);
config = {
'initial_block': {
'layout': 'cna',
'channels': 3,
'output_list': True
},
'body': {
'type': 'hugging-face',
'output_type': 'tensor',
'path': 'nvidia/segformer-b0-finetuned-ade-512-512',
'num_encoder_blocks': 2, # ~num_stages
'strides': [2, 2],
'num_attention_heads': [1, 4],
'hidden_sizes': [48, 64], # ~channels
},
'head': {'layout': 'Dnfaf',
'features': [50, 10],
'dropout_rate': 0.3,
'multisample': 0.3},
}
ppl = run('classification', TorchModel, config, 'HF-SegFormer', n_iters=100, batch_size=128)
# ppl.model.model.body.config
test(ppl, show_results=False);
ppl.model.repr()
```
# Segmentation
```
config = {
'initial_block': {'layout': 'cna', 'channels': 1},
'body': {'type': 'decoder',
'num_stages': 3,
'order': ['block'],}
}
ppl = run('segmentation', TorchModel, config, 'decoder')
ppl.model.repr()
config = {
'initial_block': {
'layout': 'cnaRp cnaRp tna+ tna+ BScna+ cnac',
'channels': [16, 32, 32, 16, 'same', 8, 1],
'custom_padding': False,
'transposed_conv': {'kernel_size': 2, 'stride': 2},
'branch': {'layout': 'ca', 'channels': 'same'}
},
}
ppl = run('segmentation', TorchModel, config, 'hardcoded unet')
config = {
'order': ['initial_block', 'encoder', 'embedding', 'decoder', 'head'],
'initial_block': {
'layout': 'cna',
'channels': 8
},
'encoder': {
'type': 'encoder',
'num_stages': 3,
'blocks/channels': '2 * same',
'skip': {'channels': 'int(1.2 * same)', 'layout': 'cna'}
},
'embedding': {
'layout': 'cna',
'channels': 'same',
'input_type': 'list',
'output_type': 'list',
},
'decoder': {
'type': 'decoder',
'blocks/channels': 'same // 2',
'upsample': {'layout': 'b', 'factor': 2},
},
'head': {
'layout': 'c',
'channels': 1}
}
ppl = run('segmentation', TorchModel, config, 'encoder->embedding->decoder from scratch')
ppl.model.model.repr(2)
config = {
'order': ['initial_block', 'encoder', 'embedding', 'decoder', 'head'],
'encoder': {
'type': 'encoder',
'num_stages': 2,
'blocks/channels': '4 * same'
},
'embedding': {
'base_block': ASPP,
'channels': 4,
'pyramid': (2, 4, 8),
'input_type': 'list',
'output_type': 'list',
},
'decoder': {
'type': 'decoder',
'num_stages': 2,
'blocks': {'base_block': ResBlock, 'channels': 'same'},
'upsample': {'layout': 'b', 'factor': 2}
},
'head': {
'layout': 'c',
'channels': 1
}
}
ppl = run('segmentation', TorchModel, config, 'unet-like with ASPP', n_iters=100, batch_size=128)
```
# Segmentation: named networks
```
config = {
'initial_block': {'layout': 'cna', 'channels': 4},
'encoder': {'blocks/channels': 'int(same * 1.3)'},
'embedding': {'channels': 'same'},
'decoder': {'blocks/channels': 'int(same // 1.3)',
'upsample': {'layout': 'b', 'factor': 2}},
'head': {'layout': 'c', 'channels': 1}
}
ppl = run('segmentation', UNet, config, 'unet')
ppl = run('segmentation', ResUNet, config, 'unet with residual blocks')
config = {
'initial_block': {'layout': 'cna', 'channels': 4},
'encoder/num_stages': 2,
'embedding/channels': 6,
'decoder/blocks/channels': 6,
}
ppl = run('segmentation', DenseUNet, config, 'unet with dense blocks')
```
# Segmentation: imported encoders
```
config = {
'order': ['initial_block', 'encoder', 'decoder', 'head'],
'initial_block': {
'layout': 'cna',
'channels': 3,
'output_type': 'list',
},
'encoder': {
'type': 'hugging-face',
'path': 'nvidia/segformer-b0-finetuned-ade-512-512',
'num_encoder_blocks': 2, # ~num_stages
'strides': [2, 2],
'num_attention_heads': [1, 4],
'hidden_sizes': [48, 64], # ~channels
},
'decoder': {
'type': 'mlpdecoder',
'upsample/features': 16,
'block/channels': 16,
},
'head': {'layout': 'c',
'channels': 1},
}
ppl = run('segmentation', TorchModel, config, 'HF-SegFormer + mlp-decoder', n_iters=100, batch_size=128)
ppl.model.repr(1)
ppl.model.model.repr(2, show_num_parameters=True)
config = {
'order': ['initial_block', 'encoder', 'embedding', 'decoder', 'head'],
'initial_block': {
'layout': 'cna',
'channels': 3,
'output_type': 'list',
},
'encoder': {
'type': 'timm',
'path': 'resnet18',
'pretrained': True,
'out_indices': (0, 1, 2),
},
'embedding': {
'base_block': ResBlock,
'channels': 'same',
'input_type': 'list',
'output_type': 'list',
},
'decoder': {
'type': 'decoder',
'num_stages': 3,
'blocks': {'base_block': ResBlock, 'channels': 'same // 4'},
'upsample': {'layout': 'b', 'factor': 2},
},
'head': {'layout': 'c',
'channels': 1},
}
ppl = run('segmentation', TorchModel, config, 'TIMM-ResNet18 + decoder', n_iters=100, batch_size=128)
ppl.model.repr(1)
ppl.model.model.repr(2, show_num_parameters=True)
```
# Callbacks
```
config = {
'callbacks': [
ReduceLROnPlateau(patience=20, cooldown=20, min_delta=0.1, factor=0.9),
EarlyStopping(patience=100, min_delta=0.01)
],
'decay': {'name': 'exp', 'gamma': 0.8, 'frequency': 20},
}
with Monitor() as monitor:
ppl = run('classification', ResNet34, config, 'resnet34', batch_size=64, n_iters=200)
test(ppl, show_results=False);
```
# Model info
```
ppl.model.show_loss()
# ppl.model.show_lr() # only learning rate
monitor.visualize()
ppl.model.information()
```
# AMP
```
config = {
'amp': False,
}
ppl = run('classification', ResNet34, config, 'resnet34', batch_size=64, n_iters=200)
config = {
'amp': True # default
}
ppl = run('classification', ResNet34, config, 'resnet34', batch_size=64, n_iters=200)
config = {
'amp': True,
'sam_rho': 0.05,
}
ppl = run('classification', ResNet34, config, 'resnet34', batch_size=64, n_iters=200)
```
# Wrappers: TTA + augmentations on train
```
from torchvision import transforms
config = {
'order': ['aug0', 'preprocess', 'initial_block', 'body', 'head'],
'aug0': {
'module': transforms.Compose([
# transforms.RandomAdjustSharpness(0.1, p=1.0),
transforms.RandomVerticalFlip(p=0.5),
transforms.RandomHorizontalFlip(p=0.5),
]),
'disable_at_inference': True,
},
'preprocess': {
'module': transforms.Resize(32),
'disable_at_inference': False,
},
}
ppl = run('classification', ResNet18, config, 'resnet18', n_iters=100, batch_size=128)
print(f'Used modules at train: {ppl.model.model._last_used_modules}')
test(ppl, show_results=False)
print(f'Used modules at inference: {ppl.model.model._last_used_modules}')
ppl.model.repr(2)
import ttach
ppl.model.wrap_tta(wrapper='ClassificationTTAWrapper',
transforms=ttach.aliases.vlip_transform(),
merge_mode='mean')
test(ppl, show_results=False)
print(f'Used modules at inference: {ppl.model.model.model._last_used_modules}')
```
# Wrappers: TRT demo
```
BATCH_SIZE = 128
ppl = run('classification', ResNet34, {}, 'resnet34', batch_size=64, n_iters=200)
test(ppl, show_results=False, batch_size=BATCH_SIZE);
model = ppl.model
%%time
from torch2trt import torch2trt
module = model.model.eval()
inputs = model.make_placeholder_data(batch_size=BATCH_SIZE, unwrap=False)
model_trt = torch2trt(module=module.eval(), inputs=inputs,
fp16_mode=True, use_onnx=True, max_batch_size=BATCH_SIZE)
diff = torch.abs(model_trt(*inputs) - module.eval()(*inputs))
torch.max(diff), torch.mean(diff)
%%timeit -r 20 -n 100
inputs = model.make_placeholder_data(batch_size=BATCH_SIZE)
with torch.no_grad():
output = module.eval()(inputs).cpu()
torch.cuda.synchronize()
del output, inputs
torch.cuda.empty_cache()
%%timeit -r 20 -n 100
inputs = model.make_placeholder_data(batch_size=BATCH_SIZE)
with torch.no_grad():
output = model_trt(inputs).cpu()
torch.cuda.synchronize()
del output, inputs
torch.cuda.empty_cache()
```
# Wrappers: TRT
```
%%time
model.wrap_trt(batch_size=BATCH_SIZE)
%%timeit -r 20 -n 100
inputs = model.make_placeholder_data(batch_size=BATCH_SIZE)
with torch.no_grad():
output = model.model(inputs).cpu()
torch.cuda.synchronize()
del output, inputs
torch.cuda.empty_cache()
inputs = model.make_placeholder_data(batch_size=1)
model.model(inputs)
model.model(torch.tile(inputs, (BATCH_SIZE, 1, 1, 1)))
test(ppl, show_results=False, batch_size=BATCH_SIZE, drop_last=True);
```
# Wrappers: TTA + TRT
```
ppl = run('classification', ResNet34, {}, 'resnet34', batch_size=BATCH_SIZE, n_iters=200)
model = ppl.model
%%time
test(ppl, show_results=False, batch_size=BATCH_SIZE, drop_last=True, n_epochs=5);
%%time
model.wrap_trt(batch_size=BATCH_SIZE, fp16_mode=True)
# import ttach
# ppl.model.wrap_tta(wrapper='ClassificationTTAWrapper',
# transforms=ttach.aliases.vlip_transform(),
# merge_mode='mean')
%%time
test(ppl, show_results=False, batch_size=BATCH_SIZE, drop_last=True, n_epochs=5);
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import plotly.express as px
from fbprophet import Prophet
# praying_df = pd.read_csv(r"C:\Users\Z Dubs\lambda\labs_week1\pop_data\historical_pop_final.csv", encoding='utf-8')
df = pd.read_csv('historical_pop_data_final.csv', encoding='utf-8')
df
```
---
Exploring the data
---
```
city_state = 'San Jose, CA'
sample = df[(df.city_state == city_state)]
sample
import seaborn as sns
import matplotlib.pyplot as plt
%matplotlib inline
# Visualize the target
sns.lineplot(data=sample, x="year", y='total_pop')
plt.show()
# Visualize comparison of target to features
sns.pairplot(sample)
plt.show()
from datetime import datetime
# Converting game_date to datetime
sample['year'] = pd.to_datetime(sample['year'], format='%Y')
sample['year'] = sample['year'].dt.year
# # Altering data to be used as constraints in train/test split
# cutoff = pd.to_datetime('2018')
# # Seperate train and test sets
# train = sample[(sample['year'] <= cutoff)]
# test = sample[(sample['year'] > cutoff)]
# train.shape, test.shape
sample
sample = sample[['year', 'total_pop']]
sample
sample.columns = ['ds', 'y']
sample
sample['ds'] = pd.to_datetime(sample['ds'], format='%Y')
# sample['ds'] = sample['ds'].dt.year
sample.dtypes
m = Prophet()
m.fit(sample)
future = m.make_future_dataframe(periods=5, freq= 'y')
future.tail()
forecast = m.predict(future)
forecast[['ds', 'yhat', 'yhat_lower', 'yhat_upper']].tail()
m.plot(forecast);
from fbprophet.plot import plot_plotly, plot_components_plotly
plot_plotly(m, forecast)
import pickle
# Save the Model to file in the current working directory
Pkl_Filename = "FbProphet.pkl"
with open(Pkl_Filename, 'wb') as file:
pickle.dump(m, file)
# Load in and try out the new pickled model
with open(Pkl_Filename, 'rb') as file:
FbProphet_model = pickle.load(file)
FbProphet_model
future2 = FbProphet_model.make_future_dataframe(periods=5, freq= 'y')
future2.tail()
forecast2 = FbProphet_model.predict(future2)
forecast2[['ds', 'yhat', 'yhat_lower', 'yhat_upper']].tail()
X = sample.iloc[:, 0].values.reshape(-1, 1)
y = sample.iloc[:, 1].values.reshape(-1, 1)
X
from sklearn.linear_model import LinearRegression
model = LinearRegression().fit(X, y)
y_pred = model.predict([[2020]])
y_pred
model.score(X, y)
from statsmodels.tsa.seasonal import seasonal_decompose
# sample.index = sample.year
result = seasonal_decompose(sample['total_pop'].values, freq=3)
result.plot()
# pip install pmdarima
# Import the library
from pmdarima import auto_arima
# Ignore harmless warnings
import warnings
warnings.filterwarnings("ignore")
# Fit auto_arima function to AirPassengers dataset
stepwise_fit = auto_arima(sample['total_pop'], start_p = 1, start_q = 1,
max_p = 3, max_q = 3, m = 12,
start_P = 0, seasonal = True,
d = None, trace = True,
error_action ='ignore', # we don't want to know if an order does not work
suppress_warnings = True, # we don't want convergence warnings
stepwise = True) # set to stepwise
# To print the summary
stepwise_fit.summary()
# Fitting Random Forest Regression to the dataset
# import the regressor
from sklearn.ensemble import RandomForestRegressor
# create regressor object
rfr = RandomForestRegressor(n_estimators = 100, random_state = 7)
# fit the regressor with x and y data
rfr.fit(X, y.ravel())
y_pred2 = rfr.predict([[2020]])
y_pred2
rfr.score(X, y)
# Visualising the Random Forest Regression results
# arange for creating a range of values
# from min value of x to max
# value of x with a difference of 0.01
# between two consecutive values
X_grid = np.arange(min(X), max(X), 0.01)
# reshape for reshaping the data into a len(X_grid)*1 array,
# i.e. to make a column out of the X_grid value
X_grid = X_grid.reshape((len(X_grid), 1))
# Scatter plot for original data
plt.scatter(x, y, color = 'blue')
# plot predicted data
plt.plot(X_grid, rfr.predict(X_grid),
color = 'green')
plt.title('Random Forest Regression')
plt.xlabel('Year')
plt.ylabel('Population')
plt.show()
import xgboost as xgb
xgbr = xgb.XGBRegressor(verbosity=0)
xgbr.fit(X, y)
y_pred3 = xgbr.predict([[2050]])
y_pred3
xgbr.score(X, y)
```
| github_jupyter |
# Chapter 2 - Math Tools II (Linear Operators)
At the end of the last chapter, we showed that the state of a quantum system can be represented with a complex unit vector. Seeing as how a quantum computer is a machine that manipulates the quantum states of its qubits, we need to characterize how the state vector can evolve. Under the framework of quantum mechanics, _Linear Operators_ are used to describe the time evolution and measurement of quantum systems.
In this chapter, we will describe what linear operators are and show how they relate to matrices. <a href="https://www.numpy.org">Numpy</a> will also be used for Python implementations.
## 2.1 Linear Operators
Operators are functions that map vectors to vectors.
$$ A : V \rightarrow W,\quad\text{ where V and W are vector spaces.}$$
If the operator $A$ maps the vector $|v\rangle$ to $|w\rangle$, then we write it as
$$ A|v\rangle = |w\rangle.$$
A linear operator is an operator that satisfies the conditions of lineararity, i.e.
1. $$A \left(|v\rangle + |w\rangle\right) = \left(A|v\rangle\right) + \left(A|w\rangle\right)$$
2. $$A \left(c|v\rangle\right) = c \left(A|v\rangle\right)$$
Since every vector $|v\rangle\in V$ can be written as a linear combination of a set of basis vectors,
$$ |v\rangle = \sum_{j} \alpha_j |v_j\rangle,$$
knowing which vectors each basis vector is mapped to under the action of a linear operator allows us to calculate which vector any input vector is mapped to:
$$ A|v\rangle = A \left(\sum_{j} \alpha_j |v_j\rangle\right) = \sum_j \alpha_j A|v_j\rangle. $$
```
#Importing necessary modules: numpy can be used to describe matrices in two ways: np.array and np.matrix
#Both were used in Chapter 1, but from now on, we will carry on with np.array
#We'll also import a sympy function so that we can visualize matrices in a clearer way. This does not affect any calculation
import numpy as np
from sympy import Matrix, init_printing
#This function uses SymPy to convert a numpy array into a clear matrix image
def view(mat):
display(Matrix(mat))
#Importing necessary modules: numpy can be used to describe matrices in two ways: np.array and np.matrix
#Both were used in Chapter 1, but from now on, we will carry on with np.array
#We'll also import a sympy function so that we can visualize matrices in a clearer way. This does not affect any calculation
import numpy as np
from sympy import Matrix, init_printing
#This function uses SymPy to convert a numpy array into a clear matrix image
def view(mat):
display(Matrix(mat))#Defining a matrix in numpy, as an array
A = np.array([[1,2,3],[4,5,6],[7,8,9]])
print('Matrix A:')
print(A)
#Multiplying two matrices:
B = np.array([[3,2,1],[6,5,4],[9,8,7]])
print('AB: ')
#Printing it
print(A@B)
print('BA:')
#Viewing it... Looks better
view(B@A)
#Let's say we have a vector |u> ε R^3, where |u> = (5,2,1). Let's say we have a vector |v> = 3|u> + (2,2,2)
#In other words, every component of |u>, u_i, is now mapped to 3u_i + 2
import numpy as np
u = np.array([5,2,1])
v = 3*u+2
v
#How do we generally represent this operation, though?
```
## 2.2 Representing Linear Operators with Matrices
A matrix is a two-dimensional array of numbers, and in chapter 1 we saw how to represent vectors as either column or row matrices — matrices with exactly one column or row respectively. A matrix can also be used to numerically represent a linear operator.
Let $V$ be an $n$-dimensional vector space with a basis set $\left\{|v_k\rangle : 1\leq k \leq n \right\}$, $W$ be an $m$-dimensional vector space with a basis set $\left\{|w_j\rangle : 1\leq j \leq m \right\}$ and $A : V \rightarrow W$ be a linear operator such that
$$ A|v_k\rangle = \sum_{j=1}^m a_{jk} |w_j\rangle. $$
Then an $m\times n$ matrix (a matrix with $m$ rows and $n$ columns) can be used to represent $A$:
$$ A \equiv \begin{pmatrix}
a_{11} & a_{12} & \cdots & a_{1n} \\
a_{21} & a_{22} & \cdots & a_{2n} \\
\vdots & \vdots & \ddots & \vdots \\
a_{m1} & a_{m2} & \cdots & a_{mn}
\end{pmatrix} $$
The columns of this matrix are the column vectors $A|v_k\rangle$.
From here onwards, we will assume that our matrix representations use orthonormal basis sets, and for any operator $A: V\rightarrow V$, we use the same basis set for input and output bases.
### 2.2.1 Matrix-Vector Multiplication
A matrix is a numerical representation of an operator, and matrix-vector multiplication is a numerical
method to calculate the action of an operator on a vector.
First, use the same basis of $V$ to represent $|v\rangle$ as a column matrix:
$$|v\rangle = \sum_{k=1}^n \alpha_k |v_k\rangle \equiv \begin{pmatrix} \alpha_1 \\ \alpha_2 \\ \vdots \\ \alpha_n \end{pmatrix}.$$
Now since $A|v\rangle = \sum_{k=1}^n \alpha_k |v_k\rangle$, we can calculate the outcome of applying the operator:
$$
\begin{align}
\begin{pmatrix}
a_{11} & a_{12} & \cdots & a_{1n} \\
a_{21} & a_{22} & \cdots & a_{2n} \\
\vdots & \vdots & \ddots & \vdots \\
a_{m1} & a_{m2} & \cdots & a_{mn}
\end{pmatrix}
\begin{pmatrix} \alpha_1 \\ \alpha_2 \\ \vdots \\ \alpha_n \end{pmatrix}
&=
\alpha_1 \begin{pmatrix} a_{11} \\ a_{21} \\ \vdots \\ a_{m1} \end{pmatrix} +
\alpha_2 \begin{pmatrix} a_{12} \\ a_{22} \\ \vdots \\ a_{m2} \end{pmatrix} +
\cdots +
\alpha_n \begin{pmatrix} a_{1n} \\ a_{2n} \\ \vdots \\ a_{mn} \end{pmatrix} \\
&=
\begin{pmatrix}
\sum_{k=1}^n \alpha_k a_{1k} \\
\sum_{k=1}^n \alpha_k a_{2k} \\
\vdots \\
\sum_{k=1}^n \alpha_k a_{mk}
\end{pmatrix}
\end{align}
$$
### 2.2.2 Composition of Operators and Matrix-Matrix Multiplication
Let $A: V\rightarrow W$ and $B: W \rightarrow X$ be linear operators acting on the vector spaces $V, W$ and $X$. The linear operator $BA : V \rightarrow X$ is defined as the composition of $A$ and $B$:
$$ BA|v\rangle = B(A |v\rangle)$$
Since this composition is also a linear operator, it can also be numerically represented with a matrix. Given basis sets $\{|v_i\rangle\}, \{|w_j\rangle\}$ and $\{|x_k\rangle\}$ for $V,W$ and $X$ respectively, and the matrices $A$ and $B$ written according to those basis sets:
$$
A =
\begin{pmatrix}
| & | & & | \\
A|v_1\rangle & A|v_2\rangle & \cdots & A|v_n\rangle \\
| & | & & | \end{pmatrix}
=
\begin{pmatrix}
a_{11} & a_{12} & \cdots & a_{1n} \\
a_{21} & a_{22} & \cdots & a_{2n} \\
\vdots & \vdots & \ddots & \vdots \\
a_{m1} & a_{m2} & \cdots & a_{mn} \end{pmatrix}
$$
$$
B =
\begin{pmatrix}
| & | & & | \\
B|w_1\rangle & B|w_2\rangle & \cdots & B|w_m\rangle \\
| & | & & | \end{pmatrix}
=
\begin{pmatrix}
b_{11} & b_{12} & \cdots & b_{1m} \\
b_{21} & b_{22} & \cdots & b_{2m} \\
\vdots & \vdots & \ddots & \vdots \\
b_{o1} & b_{o2} & \cdots & b_{om} \end{pmatrix}
$$
$$
\begin{align}
BA &=
\begin{pmatrix}
| & | & & | \\
BA|v_1\rangle & BA|v_2\rangle & \cdots & BA|v_n\rangle \\
| & | & & | \end{pmatrix} \\
&=
\begin{pmatrix}
b_{11} & b_{12} & \cdots & b_{1m} \\
b_{21} & b_{22} & \cdots & b_{2m} \\
\vdots & \vdots & \ddots & \vdots \\
b_{o1} & b_{o2} & \cdots & b_{om} \end{pmatrix}
\begin{pmatrix}
a_{11} & a_{12} & \cdots & a_{1n} \\
a_{21} & a_{22} & \cdots & a_{2n} \\
\vdots & \vdots & \ddots & \vdots \\
a_{m1} & a_{m2} & \cdots & a_{mn} \end{pmatrix} \\
&=
\begin{pmatrix}
c_{11} & c_{12} & \cdots & c_{1n} \\
c_{21} & c_{22} & \cdots & c_{2n} \\
\vdots & \vdots & \ddots & \vdots \\
c_{o1} & c_{o2} & \cdots & c_{on}
\end{pmatrix}, \quad\text{where } c_{ij} = \sum_{k=1}^m b_{ik} a_{kj}
\end{align}
$$
```
#Defining a matrix in numpy, as an array
A = np.array([[1,2,3],[4,5,6],[7,8,9]])
print('Matrix A:')
print(A)
#Multiplying two matrices:
B = np.array([[3,2,1],[6,5,4],[9,8,7]])
print('AB: ')
#Printing it
print(A@B)
print('BA:')
#Viewing it... Looks better. Doesn't affect any calculation
view(B@A)
#Representing a Linear Operator as Matrices
#Application to Quantum Computing
#X|0> = |1>, where, according to section 1.10, |0> = (1,0) and |1> = (0,1)
zero = np.array([1,0])
one = np.array([0,1])
X = np.array([[0,1],[1,0]])
print('X = ')
view(X)
print('')
#Checking to see if X|0>
XZero = X@zero
print('Is X|0> = |1>:')
#Prints true if all elements in both matrices are equal
print(XZero.all() == one.all())
print('')
print('X is a very important operator in quantum computing because it essentially acts as a quantum NOT gate. The implication of this tranformation is taking a qubit which is always 0 and making it always 1 and vice versa (X|1>=|0>).')
```
## 2.3 Adjoint / Hermitian Conjugate
Let $A : V\rightarrow V$ be a linear operator, then there exists a linear operator $A^\dagger : V \rightarrow V$ defined such that for any vectors $|v\rangle, |w\rangle \in V$ the inner product
$$ (|v\rangle, A|w\rangle) = (A^\dagger|v\rangle, |w\rangle)$$
$A^\dagger$ is called the _adjoint_ or _Hermitian conjugate_ of $A$.
Although it may not seem obvious, given a matrix representation of $A$, we can compute the matrix representation of $A^\dagger$ in the same basis by taking the transpose of $A$ and replacing every element with its complex conjugate:
$$
A \equiv \begin{pmatrix}
a_{11} & a_{12} & \cdots & a_{1n} \\
a_{21} & a_{22} & \cdots & a_{2n} \\
\vdots & \vdots & \ddots & \vdots \\
a_{n1} & a_{n2} & \cdots & a_{nn}
\end{pmatrix}
\implies
A^\dagger \equiv \begin{pmatrix}
a_{11}^* & a_{21}^* & \cdots & a_{n1}^* \\
a_{12}^* & a_{22}^* & \cdots & a_{n2}^* \\
\vdots & \vdots & \ddots & \vdots \\
a_{1n}^* & a_{2n}^* & \cdots & a_{nn}^*
\end{pmatrix}
$$
```
# Calculating the adjoint of a matrix
A = np.array([ [0+1j, 1+2j], [3+0j, 0-1j] ])
print('A = ')
view(A)
def adjoint(A):
return A.conj().transpose()
A_dagger = adjoint(A)
print('Adjoint of A = ')
view(A_dagger)
```
## 2.4 Eigenvectors and Eigenvalues
For linear operators that map to the same vector space, $A: V\rightarrow V$, a vector $|v\rangle\neq \mathbf{0}$ that is
mapped to a scaled version of itself
$$ A|v\rangle = \lambda|v\rangle$$
is called an _eigenvector_ of $A$, and the scale factor $\lambda$ is called the _eigenvalue_ of $|v\rangle$.
For example, the matrix
$$ X = \begin{pmatrix} 0 & 1 \\ 1 & 0 \end{pmatrix}$$
has eigenvectors $|+\rangle = \frac{1}{\sqrt2} \binom{1}{1}$ and $|-\rangle = \frac{1}{\sqrt2}\binom{1}{-1}$ with eigenvalues +1 and -1 respectively.
The set of all eigenvectors of an operator, sharing the same eigenvalue $\lambda$, forms a vector space which is a subspace of $V$, we call this the $\lambda$-eigenspace of the operator.
### 2.4.1 Normal Operators
If a set of eigenvectors forms a basis of $V$, we call it an eigenbasis. In the context of quantum computing, we will only consider operators that have at least one orthonormal eigenbasis that spans $V$. We call these operators _Normal operators_. A normal operator $N$ can be written in the form
$$ N = \sum_{k} \lambda_k |v_k\rangle \langle v_k| $$
where $\{|v_k\rangle\}$ forms an orthonormal eigenbasis of $V$, and $\lambda_k$ is the eigenvalue of $|v_k\rangle$. This is called the _spectral decomposition_ of $N$.
The spectral decomposition of $X$ is
$$ X = (+1)|+\rangle \langle +| + (-1)|-\rangle \langle -| $$
### 2.4.2 Trace
The trace of an operator is the sum of all of its eigenvalues. It can be calculated by adding all of the diagonal elements of any matrix representation of the operator.
### 2.4.3 Determinant
The determinant of an operator is the product of all of its eigenvalues. [Click here](https://en.wikipedia.org/wiki/Determinant) to see how to numerically calculate the determinant of a matrix representation of an operator.
Given this definition, you may now understand why solving for $\lambda$ in the equation
$$ \det(A - \lambda I) = 0$$
allows us to calculate the eigenvalues of a matrix.
```
# Verifying the eigenvalues of X
X = np.array([ [0, 1], [1, 0] ])
print('X = ')
view(X)
eigenvalues = np.linalg.eigvals(X)
print(f'X has {len(eigenvalues)} eigenvalues:', eigenvalues)
# Verifying the spectral decomposition of X
# The eigenvectors
plus = np.array([ [np.sqrt(0.5)], [np.sqrt(0.5)] ])
minus = np.array([ [np.sqrt(0.5)], [-np.sqrt(0.5)] ])
# |+><+|
plus_plus = plus @ adjoint(plus)
print('|+><+| = ')
view(plus_plus)
# |-><-|
minus_minus = minus @ adjoint(minus)
print('|-><-| = ')
view(minus_minus)
# the spectral decomposition
sd = plus_plus - minus_minus
print('|+><+| - |-><-| = ')
view(sd)
```
## 2.5 Types of Linear Operators
The following classifications of linear operators will be used all the time in quantum computing:
### 2.5.1 Identity Operator
The identity operator maps all vectors onto themselves.
$$ I |v\rangle = |v\rangle, \forall |v\rangle \in V $$
The matrix representation of the identity operator is a square matrix with 1s in all of the diagonal elements and 0s everywhere else in any choice of basis. So in a 2-dimensional space,
$$ I_2 = \begin{pmatrix} 1 & 0 \\ 0 & 1 \end{pmatrix}. $$
Every vector in $V$ is an eigenvector of the identity, with eigenvalue 1.
### Unitary Operators
A unitary operator $U$ is one whose adjoint is its own inverse:
$$ U U^\dagger = I = U^\dagger U. $$
The eigenvalues of a unitary operator all have length 1 (hence the name unitary).
The columns of a unitary matrix are all mutually orthogonal to eachother, i.e. a unitary operator maps orthogonal bases to orthogonal bases.
Unitary operators preserve inner products, i.e
$$ (|v\rangle, |w\rangle) = (U|v\rangle, U|w\rangle) $$
This can be easily shown using bra-ket notation:
$$ \begin{align}
(U|v\rangle, U|w\rangle) &= (U|v\rangle)^\dagger U|w\rangle \\
&= \langle v | U^\dagger U |w\rangle \\
&= \langle v | I |w\rangle \\
&= \langle v|w \rangle \\
&= (|v\rangle, |w\rangle)
\end{align} $$
Unitary operators are used to describe the time evolution of quantum systems.
### Hermitian Operators
A Hermitian operator $H$ is one that is equal to its adjoint:
$$ H = H^\dagger. $$
The eigenvalues of a Hermitian operator are all real numbers, moreover the matrix representation of a Hermitian operator must have real numbers on the diagonal.
Eigenvectors of a Hermitian operator with different eigenvalues are orthogonal.
Hermitian operators are used to describe measurements of quantum systems.
```
# Unitary operators
#Defining U and U†
#U = 0.5*np.array([[1+1j, 1-1j],[1-1j, 1+1j]])
U = np.sqrt(0.5)*np.array([ [1, 1], [0+1j, 0-1j] ])
UDagger = adjoint(U)
print('U = ')
view(U)
print('U† = ')
view(UDagger)
#|v> and |psi> = U|v>
v = np.array([1,2])
w = np.array([3,4])
# <v|w>
v_w = adjoint(v)@w
# <v| UDagger U |w>
v_U_w = adjoint(v)@UDagger@U@w
#UU† (Supposed to be I_2)
print('UU† = ')
view(U@UDagger)
print('This +/- 4.2*10^(-17)i is 0, so UU† = I')
print('')
print('Is <v|w> = <v| U† U |w>?')
print('Yes' if np.abs(v_w - v_U_w) < 1e-5 else 'No')
print('\nDo the eigenvalues have unit length?')
eigvals = np.linalg.eigvals(U)
for x in eigvals:
print('Yes' if np.abs(np.abs(x) - 1.0 < 1e-5) else 'No')
# Hermitian Operator
Y = np.array([ [0, 0+1j], [0-1j, 0] ])
print('Y = ')
view(Y)
print('Y† = ')
view(adjoint(Y))
print('Are the eigenvalues real?')
for x in np.linalg.eigvals(Y):
print('Yes' if np.abs(x.imag) < 1e-5 else 'No')
```
## 2.6 Tensor Product
A _tensor product_ is a way to "combine" two vector spaces into a larger vector space.
$$\otimes : V \times W \rightarrow V\otimes W$$
### 2.6.1 Kronecker Product
Within the context of quantum computing, the _Kronecker product_ is be used to numerically evaluate the tensor product of vectors and/or matrices.
For a matrix
$$ A = \begin{pmatrix}
a_{11} & a_{12} & \cdots a_{1n} \\
a_{21} & a_{22} & \cdots a_{2n} \\
\vdots & \vdots & \ddots \vdots \\
a_{m1} & a_{m2} & \cdots a_{mn} \\
\end{pmatrix} $$
and any matrix $B$, the Kronecker product can be evaluated as
$$
A\otimes B = \begin{pmatrix}
a_{11} B & a_{12} B & \cdots a_{1n} B \\
a_{21} B & a_{22} B & \cdots a_{2n} B \\
\vdots & \vdots & \ddots \vdots \\
a_{m1} B & a_{m2} B & \cdots a_{mn} B \\
\end{pmatrix} $$
Where each $a_{jk} B$ is a submatrix that is just $B$ scaled by $a_{jk}$.
The outer product $|v\rangle \langle w|$ is implicitly the tensor product $|v\rangle\otimes\langle w|$,
We often drop the $\otimes$ symbol between two kets or two bras,
$$ |v\rangle\otimes|w\rangle \equiv |v\rangle|w\rangle \equiv |vw\rangle $$
### 2.6.2 Properties of the Tensor Product
* $$ (\alpha |v\rangle) \otimes |w\rangle = \alpha (|v\rangle|w\rangle) = |v\rangle \otimes (\alpha |w\rangle) $$
* $$(|v_1\rangle + |v_2\rangle)\otimes |w\rangle = \alpha|v_1\rangle|w\rangle + |v_2\rangle|w\rangle $$
* $$|v\rangle \otimes( |w_1\rangle + |w_2\rangle) = |v\rangle|w_1\rangle + |v\rangle|w_2\rangle $$
```
v = np.array([1,2,3]).transpose()
w = np.array([4,5]).transpose()
print('v = ')
view(v)
print('w = ')
view(w)
print('Tensor product |v>⊗|w> = ')
view(np.kron(v,w))
print('Outer product |v><w| = ')
view(np.outer(v,w))
X = np.array([[0,1],[1,0]])
Z = np.array([[1,0],[0,-1]])
print('X =')
view(X)
print('Z = ')
view(Z)
print('X⊗Z = ')
view(np.kron(X,Z))
print('Z⊗X = ')
view(np.kron(Z,X))
#Excercise. Create the equivelent to np.kron() for ket vectors (i.e two column matrices)
```
| github_jupyter |
# Machine Learning - Fire Prediction
## Preprocessing
Before performing the training, we must merge, encode, and scale the data.
### Data Merge
Create three combinations of datasets by merging on date and County: drought and fire; precipitation and fire; and drought, precipitation, and fire. This will allow different combinations of machine learning.
```
# dependencies
import pandas as pd
# import fire data
fireFile = "./fire_data_clean.csv"
fireData = pd.read_csv(fireFile)
fireData
fireData["County"].unique()
# import drought data
droughtFile = "./drought_data_clean.csv"
droughtData = pd.read_csv(droughtFile)
droughtData
droughtData["County"].unique()
# import precip data
precipFile = "./precip_data_clean.csv"
precipData = pd.read_csv(precipFile)
precipData
# merge the drought and fire data
droughtMerged = pd.merge(droughtData, fireData, on = ["Date", "County"])
droughtMerged
# merge the precipitation and fire data
precipMerged = pd.merge(precipData, fireData, on = ["Date","County"])
precipMerged
# merge the drought, precipitation, and fire data
masterMerge = pd.merge(droughtMerged, precipData, on = ["Date","County"])
masterMerge
```
### Data Encoding
#### Drought Data
```
# check data types
droughtMerged.dtypes
# find null values
for column in droughtMerged.columns:
print(f"Column {column} has {droughtMerged[column].isnull().sum()} null values")
# encode the drought dataframe
droughtML = pd.get_dummies(droughtMerged)
droughtML
```
#### Precipitation Data
```
# check data types
precipMerged.dtypes
# find null values
for column in precipMerged.columns:
print(f"Column {column} has {precipMerged[column].isnull().sum()} null values")
# encode the precipitation dataframe
precipML = pd.get_dummies(precipMerged)
precipML
```
#### Drought & Precipitation Data
```
# check the data types
masterMerge.dtypes
# find null values
for column in masterMerge.columns:
print(f"Column {column} has {masterMerge[column].isnull().sum()} null values")
# encode the drought + precipitation dataframe
masterML = pd.get_dummies(masterMerge)
masterML
masterML.drop(columns='None')
```
### Data Scaling
```
#I tried more models than needed for the project requirements, but left
#the best fitting supervised machine learning models
from sklearn.preprocessing import StandardScaler, MinMaxScaler
# from sklearn.decomposition import PCA
# from sklearn.manifold import TSNE
# from sklearn.cluster import KMeans
from sklearn.ensemble import RandomForestClassifier
from matplotlib import pyplot as plt
from sklearn.datasets import make_regression
from matplotlib.pyplot import figure
import numpy as np
from sklearn.linear_model import LinearRegression, Lasso
from sklearn.model_selection import train_test_split
df = masterML
experiment_df = df[['D0', 'D1', 'D3', 'D4','AcresBurned', 'Duration','Precip']]
experiment_df
X = experiment_df
y = experiment_df['AcresBurned']
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=1)
scaler = StandardScaler().fit(X_train)
X_train_scaled = scaler.transform(X_train)
X_test_scaled = scaler.transform(X_test)
reg = LinearRegression().fit(X_train_scaled, y_train)
print(reg.score(X_test_scaled, y_test))
reg = LinearRegression().fit(X_train_scaled, y_train)
reg.score(X_train_scaled, y_train)
```
## Machine Learning
```
X = df
y = df["AcresBurned"]
# X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=1)
# scaler = StandardScaler().fit(X_train)
# X_train_scaled = scaler.transform(X_train)
# X_test_scaled = scaler.transform(X_test)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=.33, random_state=1)
scaler = StandardScaler().fit(X_train)
X_train_scaled = scaler.transform(X_train)
X_test_scaled = scaler.transform(X_test)
reg = LinearRegression().fit(X_train_scaled, y_train)
reg.score(X_test_scaled, y_test)
reg = LinearRegression().fit(X_train_scaled, y_train)
reg.score(X_train_scaled, y_train)
df = pd.DataFrame(
{"actual": y_train, "predicted" :list(reg.predict(X_train_scaled))}
)
df
df = pd.DataFrame(
{"actual": y_test, "predicted" :list(reg.predict(X_test_scaled)), "net": list(reg.predict(X_test_scaled) - y_test)}
)
df
plt.scatter(list(X_test["Precip"]), list(y_test.values), c="green", label="Training Data")
plt.scatter(list(X_test["Precip"]), reg.predict(X_test), c="red", label="Prediction")
plt.legend()
plt.ylabel('Acres Burned')
plt.xlabel('Precipitation Level by County')
#plt.hlines(y=0, xmin=y.min(), xmax=y.max())
plt.title("Linear Regression Model Based on Precipitation")
plt.show()
plt.scatter(list(X_test["D4"]), list(y_test.values), c="green", label="Training Data")
plt.scatter(list(X_test["D4"]), reg.predict(X_test), c="red", label="Prediction")
plt.legend()
plt.ylabel('Acres Burned')
plt.xlabel('Drought Level 4')
#plt.hlines(y=0, xmin=y.min(), xmax=y.max())
plt.title("Linear Regression Severe Drought Level 4")
plt.show()
plt.scatter(list(X_test["D3"]), list(y_test.values), c="green", label="Training Data")
plt.scatter(list(X_test["D3"]), reg.predict(X_test), c="red", label="Prediction")
plt.legend()
plt.ylabel('Acres Burned')
plt.xlabel('Drought Level 3')
#plt.hlines(y=0, xmin=y.min(), xmax=y.max())
plt.title("Linear Regression Drought Level 3")
plt.show()
from sklearn.ensemble import RandomForestClassifier
clf = RandomForestClassifier()
clf.fit(X_train, y_train)
print(clf.score(X_train, y_train))
print(clf.score(X_test, y_test))
plt.scatter(list(X_test["Precip"]), list(y_test.values), c="green", label="Training Data")
# plt.scatter(list(X_test["Precip"]), clf.predict(X_test), c="red", label="Prediction")
plt.legend()
plt.ylabel('Acres Burned')
plt.xlabel('Precipitation Level by County')
#plt.hlines(y=0, xmin=y.min(), xmax=y.max())
plt.title("Random Forest Training on Precipitation")
plt.show()
plt.scatter(list(X_test["Precip"]), clf.predict(X_test), c="red", label="Prediction")
plt.legend()
plt.ylabel('Acres Burned')
plt.xlabel('Precipitation Level by County')
#plt.hlines(y=0, xmin=y.min(), xmax=y.max())
plt.title("Random Forest Prediction on Precipitation")
plt.show()
plt.scatter(list(X_test["D4"]), list(y_test.values), c="green", label="Training Data")
# plt.scatter(list(X_test["Precip"]), clf.predict(X_test), c="red", label="Prediction")
plt.legend()
plt.ylabel('Acres Burned')
plt.xlabel('Severe Drought Level 4')
#plt.hlines(y=0, xmin=y.min(), xmax=y.max())
plt.title("Random Forest Training on Drought D4")
plt.show()
plt.scatter(list(X_test["D4"]), clf.predict(X_test), c="red", label="Prediction")
plt.legend()
plt.ylabel('Acres Burned')
plt.xlabel('Severe Drought Level 4')
#plt.hlines(y=0, xmin=y.min(), xmax=y.max())
plt.title("Random Forest Testing on Drought D4")
plt.show()
plt.scatter(list(X_test["D3"]), list(y_test.values), c="green", label="Training Data")
# plt.scatter(list(X_test["Precip"]), clf.predict(X_test), c="red", label="Prediction")
plt.legend()
plt.ylabel('Acres Burned')
plt.xlabel('Percent of County in Drought Level 3')
#plt.hlines(y=0, xmin=y.min(), xmax=y.max())
plt.title("Random Forest Training on Drought D3")
plt.show()
plt.scatter(list(X_test["Precip"]), clf.predict(X_test), c="red", label="Prediction")
plt.legend()
plt.ylabel('Acres Burned')
plt.xlabel('Percent of County in Drought Level 3')
#plt.hlines(y=0, xmin=y.min(), xmax=y.max())
plt.title("Random Forest Testing on Drought D3")
plt.show()
lasso = Lasso().fit(X_train_scaled, y_train)
lasso.score(X_test_scaled, y_test)
plt.scatter(list(X_test["Precip"]), list(y_test.values), c="green", label="Training RAW Data")
plt.scatter(list(X_test["Precip"]), lasso.predict(X_test), c="red", label="Training Data")
plt.legend()
#plt.hlines(y=0, xmin=y.min(), xmax=y.max())
plt.ylabel('Acres Burned')
plt.xlabel('Precipitation Level by County')
plt.title("Lasso Model Precipitation")
plt.show()
```
| github_jupyter |
```
# Copyright 2020 Google LLC
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Vertex client library: Local text binary classification model for online prediction
<table align="left">
<td>
<a href="https://colab.research.google.com/github/GoogleCloudPlatform/vertex-ai-samples/blob/master/notebooks/community/gapic/custom/showcase_local_text_binary_classification_online.ipynb">
<img src="https://cloud.google.com/ml-engine/images/colab-logo-32px.png" alt="Colab logo"> Run in Colab
</a>
</td>
<td>
<a href="https://github.com/GoogleCloudPlatform/vertex-ai-samples/blob/master/notebooks/community/gapic/custom/showcase_local_text_binary_classification_online.ipynb">
<img src="https://cloud.google.com/ml-engine/images/github-logo-32px.png" alt="GitHub logo">
View on GitHub
</a>
</td>
</table>
<br/><br/><br/>
## Overview
This tutorial demonstrates how to use the Vertex client library for Python to deploy a locally trained custom text binary classification model for online prediction.
### Dataset
The dataset used for this tutorial is the [IMDB Movie Reviews](https://www.tensorflow.org/datasets/catalog/imdb_reviews) from [TensorFlow Datasets](https://www.tensorflow.org/datasets/catalog/overview). The version of the dataset you will use in this tutorial is stored in a public Cloud Storage bucket. The trained model predicts whether a review is positive or negative in sentiment.
### Objective
In this notebook, you create a custom model locally in the notebook, then learn to deploy the locally trained model to Vertex, and then do a prediction on the deployed model. You can alternatively create and deploy models using the `gcloud` command-line tool or online using the Google Cloud Console.
The steps performed include:
- Create a model locally.
- Train the model locally.
- View the model evaluation.
- Upload the model as a Vertex `Model` resource.
- Deploy the `Model` resource to a serving `Endpoint` resource.
- Make a prediction.
- Undeploy the `Model` resource.
### Costs
This tutorial uses billable components of Google Cloud (GCP):
* Vertex AI
* Cloud Storage
Learn about [Vertex AI
pricing](https://cloud.google.com/vertex-ai/pricing) and [Cloud Storage
pricing](https://cloud.google.com/storage/pricing), and use the [Pricing
Calculator](https://cloud.google.com/products/calculator/)
to generate a cost estimate based on your projected usage.
## Installation
Install the latest version of Vertex client library.
```
import os
import sys
# Google Cloud Notebook
if os.path.exists("/opt/deeplearning/metadata/env_version"):
USER_FLAG = "--user"
else:
USER_FLAG = ""
! pip3 install -U google-cloud-aiplatform $USER_FLAG
```
Install the latest GA version of *google-cloud-storage* library as well.
```
! pip3 install -U google-cloud-storage $USER_FLAG
```
### Restart the kernel
Once you've installed the Vertex client library and Google *cloud-storage*, you need to restart the notebook kernel so it can find the packages.
```
if not os.getenv("IS_TESTING"):
# Automatically restart kernel after installs
import IPython
app = IPython.Application.instance()
app.kernel.do_shutdown(True)
```
## Before you begin
### GPU runtime
*Make sure you're running this notebook in a GPU runtime if you have that option. In Colab, select* **Runtime > Change Runtime Type > GPU**
### Set up your Google Cloud project
**The following steps are required, regardless of your notebook environment.**
1. [Select or create a Google Cloud project](https://console.cloud.google.com/cloud-resource-manager). When you first create an account, you get a $300 free credit towards your compute/storage costs.
2. [Make sure that billing is enabled for your project.](https://cloud.google.com/billing/docs/how-to/modify-project)
3. [Enable the Vertex APIs and Compute Engine APIs.](https://console.cloud.google.com/flows/enableapi?apiid=ml.googleapis.com,compute_component)
4. [The Google Cloud SDK](https://cloud.google.com/sdk) is already installed in Google Cloud Notebook.
5. Enter your project ID in the cell below. Then run the cell to make sure the
Cloud SDK uses the right project for all the commands in this notebook.
**Note**: Jupyter runs lines prefixed with `!` as shell commands, and it interpolates Python variables prefixed with `$` into these commands.
```
PROJECT_ID = "[your-project-id]" # @param {type:"string"}
if PROJECT_ID == "" or PROJECT_ID is None or PROJECT_ID == "[your-project-id]":
# Get your GCP project id from gcloud
shell_output = !gcloud config list --format 'value(core.project)' 2>/dev/null
PROJECT_ID = shell_output[0]
print("Project ID:", PROJECT_ID)
! gcloud config set project $PROJECT_ID
```
#### Region
You can also change the `REGION` variable, which is used for operations
throughout the rest of this notebook. Below are regions supported for Vertex. We recommend that you choose the region closest to you.
- Americas: `us-central1`
- Europe: `europe-west4`
- Asia Pacific: `asia-east1`
You may not use a multi-regional bucket for training with Vertex. Not all regions provide support for all Vertex services. For the latest support per region, see the [Vertex locations documentation](https://cloud.google.com/vertex-ai/docs/general/locations)
```
REGION = "us-central1" # @param {type: "string"}
```
#### Timestamp
If you are in a live tutorial session, you might be using a shared test account or project. To avoid name collisions between users on resources created, you create a timestamp for each instance session, and append onto the name of resources which will be created in this tutorial.
```
from datetime import datetime
TIMESTAMP = datetime.now().strftime("%Y%m%d%H%M%S")
```
### Authenticate your Google Cloud account
**If you are using Google Cloud Notebook**, your environment is already authenticated. Skip this step.
**If you are using Colab**, run the cell below and follow the instructions when prompted to authenticate your account via oAuth.
**Otherwise**, follow these steps:
In the Cloud Console, go to the [Create service account key](https://console.cloud.google.com/apis/credentials/serviceaccountkey) page.
**Click Create service account**.
In the **Service account name** field, enter a name, and click **Create**.
In the **Grant this service account access to project** section, click the Role drop-down list. Type "Vertex" into the filter box, and select **Vertex Administrator**. Type "Storage Object Admin" into the filter box, and select **Storage Object Admin**.
Click Create. A JSON file that contains your key downloads to your local environment.
Enter the path to your service account key as the GOOGLE_APPLICATION_CREDENTIALS variable in the cell below and run the cell.
```
# If you are running this notebook in Colab, run this cell and follow the
# instructions to authenticate your GCP account. This provides access to your
# Cloud Storage bucket and lets you submit training jobs and prediction
# requests.
# If on Google Cloud Notebook, then don't execute this code
if not os.path.exists("/opt/deeplearning/metadata/env_version"):
if "google.colab" in sys.modules:
from google.colab import auth as google_auth
google_auth.authenticate_user()
# If you are running this notebook locally, replace the string below with the
# path to your service account key and run this cell to authenticate your GCP
# account.
elif not os.getenv("IS_TESTING"):
%env GOOGLE_APPLICATION_CREDENTIALS ''
```
### Create a Cloud Storage bucket
**The following steps are required, regardless of your notebook environment.**
When you submit a custom training job using the Vertex client library, you upload a Python package
containing your training code to a Cloud Storage bucket. Vertex runs
the code from this package. In this tutorial, Vertex also saves the
trained model that results from your job in the same bucket. You can then
create an `Endpoint` resource based on this output in order to serve
online predictions.
Set the name of your Cloud Storage bucket below. Bucket names must be globally unique across all Google Cloud projects, including those outside of your organization.
```
BUCKET_NAME = "gs://[your-bucket-name]" # @param {type:"string"}
if BUCKET_NAME == "" or BUCKET_NAME is None or BUCKET_NAME == "gs://[your-bucket-name]":
BUCKET_NAME = "gs://" + PROJECT_ID + "aip-" + TIMESTAMP
```
**Only if your bucket doesn't already exist**: Run the following cell to create your Cloud Storage bucket.
```
! gsutil mb -l $REGION $BUCKET_NAME
```
Finally, validate access to your Cloud Storage bucket by examining its contents:
```
! gsutil ls -al $BUCKET_NAME
```
### Set up variables
Next, set up some variables used throughout the tutorial.
### Import libraries and define constants
#### Import Vertex client library
Import the Vertex client library into our Python environment.
```
import time
from google.cloud.aiplatform import gapic as aip
from google.protobuf import json_format
from google.protobuf.json_format import MessageToJson, ParseDict
from google.protobuf.struct_pb2 import Struct, Value
```
#### Vertex constants
Setup up the following constants for Vertex:
- `API_ENDPOINT`: The Vertex API service endpoint for dataset, model, job, pipeline and endpoint services.
- `PARENT`: The Vertex location root path for dataset, model, job, pipeline and endpoint resources.
```
# API service endpoint
API_ENDPOINT = "{}-aiplatform.googleapis.com".format(REGION)
# Vertex location root path for your dataset, model and endpoint resources
PARENT = "projects/" + PROJECT_ID + "/locations/" + REGION
```
#### Hardware Accelerators
Set the hardware accelerators (e.g., GPU), if any, for prediction.
Set the variable `DEPLOY_GPU/DEPLOY_NGPU` to use a container image supporting a GPU and the number of GPUs allocated to the virtual machine (VM) instance. For example, to use a GPU container image with 4 Nvidia Telsa K80 GPUs allocated to each VM, you would specify:
(aip.AcceleratorType.NVIDIA_TESLA_K80, 4)
For GPU, available accelerators include:
- aip.AcceleratorType.NVIDIA_TESLA_K80
- aip.AcceleratorType.NVIDIA_TESLA_P100
- aip.AcceleratorType.NVIDIA_TESLA_P4
- aip.AcceleratorType.NVIDIA_TESLA_T4
- aip.AcceleratorType.NVIDIA_TESLA_V100
Otherwise specify `(None, None)` to use a container image to run on a CPU.
```
if os.getenv("IS_TESTING_DEPOLY_GPU"):
DEPLOY_GPU, DEPLOY_NGPU = (
aip.AcceleratorType.NVIDIA_TESLA_K80,
int(os.getenv("IS_TESTING_DEPOLY_GPU")),
)
else:
DEPLOY_GPU, DEPLOY_NGPU = (aip.AcceleratorType.NVIDIA_TESLA_K80, 1)
```
#### Container (Docker) image
Next, we will set the Docker container images for prediction
- Set the variable `TF` to the TensorFlow version of the container image. For example, `2-1` would be version 2.1, and `1-15` would be version 1.15. The following list shows some of the pre-built images available:
- TensorFlow 1.15
- `gcr.io/cloud-aiplatform/prediction/tf-cpu.1-15:latest`
- `gcr.io/cloud-aiplatform/prediction/tf-gpu.1-15:latest`
- TensorFlow 2.1
- `gcr.io/cloud-aiplatform/prediction/tf2-cpu.2-1:latest`
- `gcr.io/cloud-aiplatform/prediction/tf2-gpu.2-1:latest`
- TensorFlow 2.2
- `gcr.io/cloud-aiplatform/prediction/tf2-cpu.2-2:latest`
- `gcr.io/cloud-aiplatform/prediction/tf2-gpu.2-2:latest`
- TensorFlow 2.3
- `gcr.io/cloud-aiplatform/prediction/tf2-cpu.2-3:latest`
- `gcr.io/cloud-aiplatform/prediction/tf2-gpu.2-3:latest`
- XGBoost
- `gcr.io/cloud-aiplatform/prediction/xgboost-cpu.1-2:latest`
- `gcr.io/cloud-aiplatform/prediction/xgboost-cpu.1-1:latest`
- `gcr.io/cloud-aiplatform/prediction/xgboost-cpu.0-90:latest`
- `gcr.io/cloud-aiplatform/prediction/xgboost-cpu.0-82:latest`
- Scikit-learn
- `gcr.io/cloud-aiplatform/prediction/sklearn-cpu.0-23:latest`
- `gcr.io/cloud-aiplatform/prediction/sklearn-cpu.0-22:latest`
- `gcr.io/cloud-aiplatform/prediction/sklearn-cpu.0-20:latest`
For the latest list, see [Pre-built containers for prediction](https://cloud.google.com/vertex-ai/docs/predictions/pre-built-containers)
```
if os.getenv("IS_TESTING_TF"):
TF = os.getenv("IS_TESTING_TF")
else:
TF = "2-1"
if TF[0] == "2":
if DEPLOY_GPU:
DEPLOY_VERSION = "tf2-gpu.{}".format(TF)
else:
DEPLOY_VERSION = "tf2-cpu.{}".format(TF)
else:
if DEPLOY_GPU:
DEPLOY_VERSION = "tf-gpu.{}".format(TF)
else:
DEPLOY_VERSION = "tf-cpu.{}".format(TF)
DEPLOY_IMAGE = "gcr.io/cloud-aiplatform/prediction/{}:latest".format(DEPLOY_VERSION)
print("Deployment:", DEPLOY_IMAGE, DEPLOY_GPU)
```
#### Machine Type
Next, set the machine type to use for prediction.
- Set the variable `DEPLOY_COMPUTE` to configure the compute resources for the VM you will use for prediction.
- `machine type`
- `n1-standard`: 3.75GB of memory per vCPU.
- `n1-highmem`: 6.5GB of memory per vCPU
- `n1-highcpu`: 0.9 GB of memory per vCPU
- `vCPUs`: number of \[2, 4, 8, 16, 32, 64, 96 \]
*Note: You may also use n2 and e2 machine types for training and deployment, but they do not support GPUs*
```
if os.getenv("IS_TESTING_DEPLOY_MACHINE"):
MACHINE_TYPE = os.getenv("IS_TESTING_DEPLOY_MACHINE")
else:
MACHINE_TYPE = "n1-standard"
VCPU = "4"
DEPLOY_COMPUTE = MACHINE_TYPE + "-" + VCPU
print("Deploy machine type", DEPLOY_COMPUTE)
```
# Tutorial
Now you are ready to start locally training a custom model IMDB Movie Reviews, and then deploy the model to the cloud.
## Set up clients
The Vertex client library works as a client/server model. On your side (the Python script) you will create a client that sends requests and receives responses from the Vertex server.
You will use different clients in this tutorial for different steps in the workflow. So set them all up upfront.
- Model Service for `Model` resources.
- Endpoint Service for deployment.
- Prediction Service for serving.
```
# client options same for all services
client_options = {"api_endpoint": API_ENDPOINT}
def create_model_client():
client = aip.ModelServiceClient(client_options=client_options)
return client
def create_endpoint_client():
client = aip.EndpointServiceClient(client_options=client_options)
return client
def create_prediction_client():
client = aip.PredictionServiceClient(client_options=client_options)
return client
clients = {}
clients["model"] = create_model_client()
clients["endpoint"] = create_endpoint_client()
clients["prediction"] = create_prediction_client()
for client in clients.items():
print(client)
```
## Train a model locally
In this tutorial, you train a IMDB Movie Reviews model locally.
### Set location to store trained model
You set the variable `MODEL_DIR` for where in your Cloud Storage bucket to save the model in TensorFlow SavedModel format.
Also, you create a local folder for the training script.
```
MODEL_DIR = BUCKET_NAME + "/imdb"
model_path_to_deploy = MODEL_DIR
! rm -rf custom
! mkdir custom
! mkdir custom/trainer
```
#### Task.py contents
In the next cell, you write the contents of the training script task.py. I won't go into detail, it's just there for you to browse. In summary:
- Gets the directory where to save the model artifacts from the command line (`--model_dir`), and if not specified, then from the environment variable `AIP_MODEL_DIR`.
- Loads IMDB Movie Reviews dataset from TF Datasets (tfds).
- Builds a simple RNN model using TF.Keras model API.
- Compiles the model (`compile()`).
- Sets a training distribution strategy according to the argument `args.distribute`.
- Trains the model (`fit()`) with epochs specified by `args.epochs`.
- Saves the trained model (`save(args.model_dir)`) to the specified model directory.
```
%%writefile custom/trainer/task.py
# Single, Mirror and Multi-Machine Distributed Training for IMDB
import tensorflow_datasets as tfds
import tensorflow as tf
from tensorflow.python.client import device_lib
import argparse
import os
import sys
tfds.disable_progress_bar()
parser = argparse.ArgumentParser()
parser.add_argument('--model-dir', dest='model_dir',
default=os.getenv('AIP_MODEL_DIR'), type=str, help='Model dir.')
parser.add_argument('--lr', dest='lr',
default=1e-4, type=float,
help='Learning rate.')
parser.add_argument('--epochs', dest='epochs',
default=20, type=int,
help='Number of epochs.')
parser.add_argument('--steps', dest='steps',
default=100, type=int,
help='Number of steps per epoch.')
parser.add_argument('--distribute', dest='distribute', type=str, default='single',
help='distributed training strategy')
args = parser.parse_args()
print('Python Version = {}'.format(sys.version))
print('TensorFlow Version = {}'.format(tf.__version__))
print('TF_CONFIG = {}'.format(os.environ.get('TF_CONFIG', 'Not found')))
print(device_lib.list_local_devices())
# Single Machine, single compute device
if args.distribute == 'single':
if tf.test.is_gpu_available():
strategy = tf.distribute.OneDeviceStrategy(device="/gpu:0")
else:
strategy = tf.distribute.OneDeviceStrategy(device="/cpu:0")
# Single Machine, multiple compute device
elif args.distribute == 'mirror':
strategy = tf.distribute.MirroredStrategy()
# Multiple Machine, multiple compute device
elif args.distribute == 'multi':
strategy = tf.distribute.experimental.MultiWorkerMirroredStrategy()
# Multi-worker configuration
print('num_replicas_in_sync = {}'.format(strategy.num_replicas_in_sync))
# Preparing dataset
BUFFER_SIZE = 10000
BATCH_SIZE = 64
def make_datasets():
dataset, info = tfds.load('imdb_reviews/subwords8k', with_info=True,
as_supervised=True)
train_dataset, test_dataset = dataset['train'], dataset['test']
encoder = info.features['text'].encoder
padded_shapes = ([None],())
return train_dataset.shuffle(BUFFER_SIZE).padded_batch(BATCH_SIZE, padded_shapes), encoder
train_dataset, encoder = make_datasets()
# Build the Keras model
def build_and_compile_rnn_model(encoder):
model = tf.keras.Sequential([
tf.keras.layers.Embedding(encoder.vocab_size, 64),
tf.keras.layers.Bidirectional(tf.keras.layers.LSTM(64)),
tf.keras.layers.Dense(64, activation='relu'),
tf.keras.layers.Dense(1, activation='sigmoid')
])
model.compile(loss=tf.keras.losses.BinaryCrossentropy(from_logits=True),
optimizer=tf.keras.optimizers.Adam(args.lr),
metrics=['accuracy'])
return model
with strategy.scope():
# Creation of dataset, and model building/compiling need to be within
# `strategy.scope()`.
model = build_and_compile_rnn_model(encoder)
# Train the model
model.fit(train_dataset, epochs=args.epochs, steps_per_epoch=args.steps)
model.save(args.model_dir)
```
### Train the model
```
! python custom/trainer/task.py --epochs=10 --model-dir=$MODEL_DIR
```
## Load the saved model
Your model is stored in a TensorFlow SavedModel format in a Cloud Storage bucket. Now load it from the Cloud Storage bucket, and then you can do some things, like evaluate the model, and do a prediction.
To load, you use the TF.Keras `model.load_model()` method passing it the Cloud Storage path where the model is saved -- specified by `MODEL_DIR`.
```
import tensorflow as tf
model = tf.keras.models.load_model(MODEL_DIR)
```
## Evaluate the model
Now let's find out how good the model is.
### Load evaluation data
You will load the IMDB Movie Review test (holdout) data from `tfds.datasets`, using the method `load()`. This will return the dataset as a tuple of two elements. The first element is the dataset and the second is information on the dataset, which will contain the predefined vocabulary encoder. The encoder will convert words into a numerical embedding, which was pretrained and used in the custom training script.
When you trained the model, you needed to set a fix input length for your text. For forward feeding batches, the `padded_batch()` property of the corresponding `tf.dataset` was set to pad each input sequence into the same shape for a batch.
For the test data, you also need to set the `padded_batch()` property accordingly.
```
import tensorflow_datasets as tfds
dataset, info = tfds.load("imdb_reviews/subwords8k", with_info=True, as_supervised=True)
test_dataset = dataset["test"]
encoder = info.features["text"].encoder
BATCH_SIZE = 64
padded_shapes = ([None], ())
test_dataset = test_dataset.padded_batch(BATCH_SIZE, padded_shapes)
```
### Perform the model evaluation
Now evaluate how well the model in the custom job did.
```
model.evaluate(test_dataset)
```
## Upload the model for serving
Next, you will upload your TF.Keras model from the custom job to Vertex `Model` service, which will create a Vertex `Model` resource for your custom model. During upload, you need to define a serving function to convert data to the format your model expects. If you send encoded data to Vertex, your serving function ensures that the data is decoded on the model server before it is passed as input to your model.
### How does the serving function work
When you send a request to an online prediction server, the request is received by a HTTP server. The HTTP server extracts the prediction request from the HTTP request content body. The extracted prediction request is forwarded to the serving function. For Google pre-built prediction containers, the request content is passed to the serving function as a `tf.string`.
The serving function consists of two parts:
- `preprocessing function`:
- Converts the input (`tf.string`) to the input shape and data type of the underlying model (dynamic graph).
- Performs the same preprocessing of the data that was done during training the underlying model -- e.g., normalizing, scaling, etc.
- `post-processing function`:
- Converts the model output to format expected by the receiving application -- e.q., compresses the output.
- Packages the output for the the receiving application -- e.g., add headings, make JSON object, etc.
Both the preprocessing and post-processing functions are converted to static graphs which are fused to the model. The output from the underlying model is passed to the post-processing function. The post-processing function passes the converted/packaged output back to the HTTP server. The HTTP server returns the output as the HTTP response content.
One consideration you need to consider when building serving functions for TF.Keras models is that they run as static graphs. That means, you cannot use TF graph operations that require a dynamic graph. If you do, you will get an error during the compile of the serving function which will indicate that you are using an EagerTensor which is not supported.
## Get the serving function signature
You can get the signatures of your model's input and output layers by reloading the model into memory, and querying it for the signatures corresponding to each layer.
When making a prediction request, you need to route the request to the serving function instead of the model, so you need to know the input layer name of the serving function -- which you will use later when you make a prediction request.
```
loaded = tf.saved_model.load(model_path_to_deploy)
serving_input = list(
loaded.signatures["serving_default"].structured_input_signature[1].keys()
)[0]
print("Serving function input:", serving_input)
```
### Upload the model
Use this helper function `upload_model` to upload your model, stored in SavedModel format, up to the `Model` service, which will instantiate a Vertex `Model` resource instance for your model. Once you've done that, you can use the `Model` resource instance in the same way as any other Vertex `Model` resource instance, such as deploying to an `Endpoint` resource for serving predictions.
The helper function takes the following parameters:
- `display_name`: A human readable name for the `Endpoint` service.
- `image_uri`: The container image for the model deployment.
- `model_uri`: The Cloud Storage path to our SavedModel artificat. For this tutorial, this is the Cloud Storage location where the `trainer/task.py` saved the model artifacts, which we specified in the variable `MODEL_DIR`.
The helper function calls the `Model` client service's method `upload_model`, which takes the following parameters:
- `parent`: The Vertex location root path for `Dataset`, `Model` and `Endpoint` resources.
- `model`: The specification for the Vertex `Model` resource instance.
Let's now dive deeper into the Vertex model specification `model`. This is a dictionary object that consists of the following fields:
- `display_name`: A human readable name for the `Model` resource.
- `metadata_schema_uri`: Since your model was built without an Vertex `Dataset` resource, you will leave this blank (`''`).
- `artificat_uri`: The Cloud Storage path where the model is stored in SavedModel format.
- `container_spec`: This is the specification for the Docker container that will be installed on the `Endpoint` resource, from which the `Model` resource will serve predictions. Use the variable you set earlier `DEPLOY_GPU != None` to use a GPU; otherwise only a CPU is allocated.
Uploading a model into a Vertex Model resource returns a long running operation, since it may take a few moments. You call response.result(), which is a synchronous call and will return when the Vertex Model resource is ready.
The helper function returns the Vertex fully qualified identifier for the corresponding Vertex Model instance upload_model_response.model. You will save the identifier for subsequent steps in the variable model_to_deploy_id.
```
IMAGE_URI = DEPLOY_IMAGE
def upload_model(display_name, image_uri, model_uri):
model = {
"display_name": display_name,
"metadata_schema_uri": "",
"artifact_uri": model_uri,
"container_spec": {
"image_uri": image_uri,
"command": [],
"args": [],
"env": [{"name": "env_name", "value": "env_value"}],
"ports": [{"container_port": 8080}],
"predict_route": "",
"health_route": "",
},
}
response = clients["model"].upload_model(parent=PARENT, model=model)
print("Long running operation:", response.operation.name)
upload_model_response = response.result(timeout=180)
print("upload_model_response")
print(" model:", upload_model_response.model)
return upload_model_response.model
model_to_deploy_id = upload_model("imdb-" + TIMESTAMP, IMAGE_URI, model_path_to_deploy)
```
### Get `Model` resource information
Now let's get the model information for just your model. Use this helper function `get_model`, with the following parameter:
- `name`: The Vertex unique identifier for the `Model` resource.
This helper function calls the Vertex `Model` client service's method `get_model`, with the following parameter:
- `name`: The Vertex unique identifier for the `Model` resource.
```
def get_model(name):
response = clients["model"].get_model(name=name)
print(response)
get_model(model_to_deploy_id)
```
## Deploy the `Model` resource
Now deploy the trained Vertex custom `Model` resource. This requires two steps:
1. Create an `Endpoint` resource for deploying the `Model` resource to.
2. Deploy the `Model` resource to the `Endpoint` resource.
### Create an `Endpoint` resource
Use this helper function `create_endpoint` to create an endpoint to deploy the model to for serving predictions, with the following parameter:
- `display_name`: A human readable name for the `Endpoint` resource.
The helper function uses the endpoint client service's `create_endpoint` method, which takes the following parameter:
- `display_name`: A human readable name for the `Endpoint` resource.
Creating an `Endpoint` resource returns a long running operation, since it may take a few moments to provision the `Endpoint` resource for serving. You call `response.result()`, which is a synchronous call and will return when the Endpoint resource is ready. The helper function returns the Vertex fully qualified identifier for the `Endpoint` resource: `response.name`.
```
ENDPOINT_NAME = "imdb_endpoint-" + TIMESTAMP
def create_endpoint(display_name):
endpoint = {"display_name": display_name}
response = clients["endpoint"].create_endpoint(parent=PARENT, endpoint=endpoint)
print("Long running operation:", response.operation.name)
result = response.result(timeout=300)
print("result")
print(" name:", result.name)
print(" display_name:", result.display_name)
print(" description:", result.description)
print(" labels:", result.labels)
print(" create_time:", result.create_time)
print(" update_time:", result.update_time)
return result
result = create_endpoint(ENDPOINT_NAME)
```
Now get the unique identifier for the `Endpoint` resource you created.
```
# The full unique ID for the endpoint
endpoint_id = result.name
# The short numeric ID for the endpoint
endpoint_short_id = endpoint_id.split("/")[-1]
print(endpoint_id)
```
### Compute instance scaling
You have several choices on scaling the compute instances for handling your online prediction requests:
- Single Instance: The online prediction requests are processed on a single compute instance.
- Set the minimum (`MIN_NODES`) and maximum (`MAX_NODES`) number of compute instances to one.
- Manual Scaling: The online prediction requests are split across a fixed number of compute instances that you manually specified.
- Set the minimum (`MIN_NODES`) and maximum (`MAX_NODES`) number of compute instances to the same number of nodes. When a model is first deployed to the instance, the fixed number of compute instances are provisioned and online prediction requests are evenly distributed across them.
- Auto Scaling: The online prediction requests are split across a scaleable number of compute instances.
- Set the minimum (`MIN_NODES`) number of compute instances to provision when a model is first deployed and to de-provision, and set the maximum (`MAX_NODES) number of compute instances to provision, depending on load conditions.
The minimum number of compute instances corresponds to the field `min_replica_count` and the maximum number of compute instances corresponds to the field `max_replica_count`, in your subsequent deployment request.
```
MIN_NODES = 1
MAX_NODES = 1
```
### Deploy `Model` resource to the `Endpoint` resource
Use this helper function `deploy_model` to deploy the `Model` resource to the `Endpoint` resource you created for serving predictions, with the following parameters:
- `model`: The Vertex fully qualified model identifier of the model to upload (deploy) from the training pipeline.
- `deploy_model_display_name`: A human readable name for the deployed model.
- `endpoint`: The Vertex fully qualified endpoint identifier to deploy the model to.
The helper function calls the `Endpoint` client service's method `deploy_model`, which takes the following parameters:
- `endpoint`: The Vertex fully qualified `Endpoint` resource identifier to deploy the `Model` resource to.
- `deployed_model`: The requirements specification for deploying the model.
- `traffic_split`: Percent of traffic at the endpoint that goes to this model, which is specified as a dictionary of one or more key/value pairs.
- If only one model, then specify as **{ "0": 100 }**, where "0" refers to this model being uploaded and 100 means 100% of the traffic.
- If there are existing models on the endpoint, for which the traffic will be split, then use `model_id` to specify as **{ "0": percent, model_id: percent, ... }**, where `model_id` is the model id of an existing model to the deployed endpoint. The percents must add up to 100.
Let's now dive deeper into the `deployed_model` parameter. This parameter is specified as a Python dictionary with the minimum required fields:
- `model`: The Vertex fully qualified model identifier of the (upload) model to deploy.
- `display_name`: A human readable name for the deployed model.
- `disable_container_logging`: This disables logging of container events, such as execution failures (default is container logging is enabled). Container logging is typically enabled when debugging the deployment and then disabled when deployed for production.
- `dedicated_resources`: This refers to how many compute instances (replicas) that are scaled for serving prediction requests.
- `machine_spec`: The compute instance to provision. Use the variable you set earlier `DEPLOY_GPU != None` to use a GPU; otherwise only a CPU is allocated.
- `min_replica_count`: The number of compute instances to initially provision, which you set earlier as the variable `MIN_NODES`.
- `max_replica_count`: The maximum number of compute instances to scale to, which you set earlier as the variable `MAX_NODES`.
#### Traffic Split
Let's now dive deeper into the `traffic_split` parameter. This parameter is specified as a Python dictionary. This might at first be a tad bit confusing. Let me explain, you can deploy more than one instance of your model to an endpoint, and then set how much (percent) goes to each instance.
Why would you do that? Perhaps you already have a previous version deployed in production -- let's call that v1. You got better model evaluation on v2, but you don't know for certain that it is really better until you deploy to production. So in the case of traffic split, you might want to deploy v2 to the same endpoint as v1, but it only get's say 10% of the traffic. That way, you can monitor how well it does without disrupting the majority of users -- until you make a final decision.
#### Response
The method returns a long running operation `response`. We will wait sychronously for the operation to complete by calling the `response.result()`, which will block until the model is deployed. If this is the first time a model is deployed to the endpoint, it may take a few additional minutes to complete provisioning of resources.
```
DEPLOYED_NAME = "imdb_deployed-" + TIMESTAMP
def deploy_model(
model, deployed_model_display_name, endpoint, traffic_split={"0": 100}
):
if DEPLOY_GPU:
machine_spec = {
"machine_type": DEPLOY_COMPUTE,
"accelerator_type": DEPLOY_GPU,
"accelerator_count": DEPLOY_NGPU,
}
else:
machine_spec = {
"machine_type": DEPLOY_COMPUTE,
"accelerator_count": 0,
}
deployed_model = {
"model": model,
"display_name": deployed_model_display_name,
"dedicated_resources": {
"min_replica_count": MIN_NODES,
"max_replica_count": MAX_NODES,
"machine_spec": machine_spec,
},
"disable_container_logging": False,
}
response = clients["endpoint"].deploy_model(
endpoint=endpoint, deployed_model=deployed_model, traffic_split=traffic_split
)
print("Long running operation:", response.operation.name)
result = response.result()
print("result")
deployed_model = result.deployed_model
print(" deployed_model")
print(" id:", deployed_model.id)
print(" model:", deployed_model.model)
print(" display_name:", deployed_model.display_name)
print(" create_time:", deployed_model.create_time)
return deployed_model.id
deployed_model_id = deploy_model(model_to_deploy_id, DEPLOYED_NAME, endpoint_id)
```
## Make a online prediction request
Now do a online prediction to your deployed model.
### Prepare the request content
Since the dataset is a `tf.dataset`, which acts as a generator, we must use it as an iterator to access the data items in the test data. We do the following to get a single data item from the test data:
- Set the property for the number of batches to draw per iteration to one using the method `take(1)`.
- Iterate once through the test data -- i.e., we do a break within the for loop.
- In the single iteration, we save the data item which is in the form of a tuple.
- The data item will be the first element of the tuple, which you then will convert from an tensor to a numpy array -- `data[0].numpy()`.
```
import tensorflow_datasets as tfds
dataset, info = tfds.load("imdb_reviews/subwords8k", with_info=True, as_supervised=True)
test_dataset = dataset["test"]
test_dataset.take(1)
for data in test_dataset:
print(data)
break
test_item = data[0].numpy()
```
### Send the prediction request
Ok, now you have a test data item. Use this helper function `predict_data`, which takes the following parameters:
- `data`: The test data item is a 64 padded numpy 1D array.
- `endpoint`: The Vertex AI fully qualified identifier for the endpoint where the model was deployed.
- `parameters_dict`: Additional parameters for serving.
This function uses the prediction client service and calls the `predict` method with the following parameters:
- `endpoint`: The Vertex AI fully qualified identifier for the endpoint where the model was deployed.
- `instances`: A list of instances (data items) to predict.
- `parameters`: Additional parameters for serving.
To pass the test data to the prediction service, you must package it for transmission to the serving binary as follows:
1. Convert the data item from a 1D numpy array to a 1D Python list.
2. Convert the prediction request to a serialized Google protobuf (`json_format.ParseDict()`)
Each instance in the prediction request is a dictionary entry of the form:
{input_name: content}
- `input_name`: the name of the input layer of the underlying model.
- `content`: The data item as a 1D Python list.
Since the `predict()` service can take multiple data items (instances), you will send your single data item as a list of one data item. As a final step, you package the instances list into Google's protobuf format -- which is what we pass to the `predict()` service.
The `response` object returns a list, where each element in the list corresponds to the corresponding image in the request. You will see in the output for each prediction:
- `predictions` -- the predicated binary sentiment between 0 (negative) and 1 (positive).
```
def predict_data(data, endpoint, parameters_dict):
parameters = json_format.ParseDict(parameters_dict, Value())
# The format of each instance should conform to the deployed model's prediction input schema.
instances_list = [{serving_input: data.tolist()}]
instances = [json_format.ParseDict(s, Value()) for s in instances_list]
response = clients["prediction"].predict(
endpoint=endpoint, instances=instances, parameters=parameters
)
print("response")
print(" deployed_model_id:", response.deployed_model_id)
predictions = response.predictions
print("predictions")
for prediction in predictions:
print(" prediction:", prediction)
predict_data(test_item, endpoint_id, None)
```
## Undeploy the `Model` resource
Now undeploy your `Model` resource from the serving `Endpoint` resoure. Use this helper function `undeploy_model`, which takes the following parameters:
- `deployed_model_id`: The model deployment identifier returned by the endpoint service when the `Model` resource was deployed to.
- `endpoint`: The Vertex fully qualified identifier for the `Endpoint` resource where the `Model` is deployed to.
This function calls the endpoint client service's method `undeploy_model`, with the following parameters:
- `deployed_model_id`: The model deployment identifier returned by the endpoint service when the `Model` resource was deployed.
- `endpoint`: The Vertex fully qualified identifier for the `Endpoint` resource where the `Model` resource is deployed.
- `traffic_split`: How to split traffic among the remaining deployed models on the `Endpoint` resource.
Since this is the only deployed model on the `Endpoint` resource, you simply can leave `traffic_split` empty by setting it to {}.
```
def undeploy_model(deployed_model_id, endpoint):
response = clients["endpoint"].undeploy_model(
endpoint=endpoint, deployed_model_id=deployed_model_id, traffic_split={}
)
print(response)
undeploy_model(deployed_model_id, endpoint_id)
```
# Cleaning up
To clean up all GCP resources used in this project, you can [delete the GCP
project](https://cloud.google.com/resource-manager/docs/creating-managing-projects#shutting_down_projects) you used for the tutorial.
Otherwise, you can delete the individual resources you created in this tutorial:
- Dataset
- Pipeline
- Model
- Endpoint
- Batch Job
- Custom Job
- Hyperparameter Tuning Job
- Cloud Storage Bucket
```
delete_dataset = True
delete_pipeline = True
delete_model = True
delete_endpoint = True
delete_batchjob = True
delete_customjob = True
delete_hptjob = True
delete_bucket = True
# Delete the dataset using the Vertex fully qualified identifier for the dataset
try:
if delete_dataset and "dataset_id" in globals():
clients["dataset"].delete_dataset(name=dataset_id)
except Exception as e:
print(e)
# Delete the training pipeline using the Vertex fully qualified identifier for the pipeline
try:
if delete_pipeline and "pipeline_id" in globals():
clients["pipeline"].delete_training_pipeline(name=pipeline_id)
except Exception as e:
print(e)
# Delete the model using the Vertex fully qualified identifier for the model
try:
if delete_model and "model_to_deploy_id" in globals():
clients["model"].delete_model(name=model_to_deploy_id)
except Exception as e:
print(e)
# Delete the endpoint using the Vertex fully qualified identifier for the endpoint
try:
if delete_endpoint and "endpoint_id" in globals():
clients["endpoint"].delete_endpoint(name=endpoint_id)
except Exception as e:
print(e)
# Delete the batch job using the Vertex fully qualified identifier for the batch job
try:
if delete_batchjob and "batch_job_id" in globals():
clients["job"].delete_batch_prediction_job(name=batch_job_id)
except Exception as e:
print(e)
# Delete the custom job using the Vertex fully qualified identifier for the custom job
try:
if delete_customjob and "job_id" in globals():
clients["job"].delete_custom_job(name=job_id)
except Exception as e:
print(e)
# Delete the hyperparameter tuning job using the Vertex fully qualified identifier for the hyperparameter tuning job
try:
if delete_hptjob and "hpt_job_id" in globals():
clients["job"].delete_hyperparameter_tuning_job(name=hpt_job_id)
except Exception as e:
print(e)
if delete_bucket and "BUCKET_NAME" in globals():
! gsutil rm -r $BUCKET_NAME
```
| github_jupyter |
# LightGBM学习示例
姓名:邹子涵
学号:202020085400139
# 建立模型
```
import pandas as pd
import lightgbm as lgb
import numpy as np
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split, StratifiedKFold, GridSearchCV
from sklearn.metrics import mean_squared_error
import warnings
warnings.filterwarnings('ignore')
%matplotlib inline
data = pd.read_csv('dataset/abalone.data',header=None,names=['Sex','Length','Diameter','Height','Whole-weight','Shucked-weight','Viscera-weight','Shell-weight','Rings'])
data.loc[(data['Sex']=="M"),'Sex']=0
data.loc[(data['Sex']=="F"),'Sex']=1
data.loc[(data['Sex']=="I"),'Sex']=2
n = len(data)+1
nlist = range(1,n)
data['id'] = nlist
data['Rings'] = np.log(data['Rings'])
data
train,test = train_test_split(data,test_size=0.2,random_state=0)
train
col_features = ['Length','Diameter','Height','Whole-weight','Shucked-weight','Viscera-weight','Shell-weight']
y_train = train['Rings']
y_test = test['Rings']
X_train = train[col_features]
X_test = test[col_features]
print('开始训练...')
# 直接初始化LGBMRegressor
# 这个LightGBM的Regressor和sklearn中其他Regressor基本是一致的
gbm = lgb.LGBMRegressor(objective='regression',
num_leaves=31,
learning_rate=0.05,
n_estimators=20)
# 使用fit函数拟合
gbm.fit(X_train, y_train,
eval_set=[(X_test, y_test)],
eval_metric='l1',
early_stopping_rounds=5)
# 预测
print('开始预测...')
y_pred = gbm.predict(X_test, num_iteration=gbm.best_iteration_)
# 评估预测结果
print('预测结果的rmse是:')
print(mean_squared_error(y_test, y_pred) ** 0.5)
```
# sklearn与LightGBM配合使用
LightGBM建模,sklearn评估
```
# 配合scikit-learn的网格搜索交叉验证选择最优超参数
estimator = lgb.LGBMRegressor(num_leaves=31)
param_grid = {
'learning_rate': [0.01, 0.1, 1],
'n_estimators': [20, 40]
}
gbm = GridSearchCV(estimator, param_grid)
gbm.fit(X_train, y_train)
print('用网格搜索找到的最优超参数为:')
print(gbm.best_params_)
# 构建lgb中的Dataset数据格式
lgb_train = lgb.Dataset(X_train, y_train)
lgb_test = lgb.Dataset(X_test, y_test, reference=lgb_train)
# 设定参数
params = {
'num_leaves': 5,
'metric': ('l1', 'l2'),
'verbose': 0
}
evals_result = {} # to record eval results for plotting
print('开始训练...')
# 训练
gbm = lgb.train(params,
lgb_train,
num_boost_round=100,
valid_sets=[lgb_train, lgb_test],
feature_name=['f' + str(i + 1) for i in range(7)],
categorical_feature=[21],
evals_result=evals_result,
verbose_eval=10)
# 保存模型
print('保存模型...')
# 保存模型到文件中
gbm.save_model('.\model.txt')
print('在训练过程中绘图...')
ax = lgb.plot_metric(evals_result, metric='l1')
plt.show()
print('画出特征重要度...')
ax = lgb.plot_importance(gbm, max_num_features=10)
plt.show()
```
| github_jupyter |
<a href="https://colab.research.google.com/github/NeuromatchAcademy/course-content-dl/blob/main/tutorials/W1D3_MultiLayerPerceptrons/student/W1D3_Tutorial1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Tutorial 1: Biological vs. Artificial neurons
**Week 1, Day 3: Multi Layer Perceptrons**
**By Neuromatch Academy**
__Content creators:__ Arash Ash, Surya Ganguli
__Content reviewers:__ Saeed Salehi, Felix Bartsch, Yu-Fang Yang, Antoine De Comite, Melvin Selim Atay
__Content editors:__ Gagana B, Spiros Chavlis
__Production editors:__ Anoop Kulkarni, Spiros Chavlis
**Our 2021 Sponsors, including Presenting Sponsor Facebook Reality Labs**
<p align='center'><img src='https://github.com/NeuromatchAcademy/widgets/blob/master/sponsors.png?raw=True'/></p>
---
# Tutorial objectives
In this tutorial we will explore the Multi-leayer Perceptrons (MLPs). MLPs are arguably one of the most tractable models that we can use to study deep learning fundamentals. Here we will learn why they are:
* similar to biological networks
* good at function approximation
* implemented the way they are in PyTorch
```
# @title Video 0: Introduction
# Insert the ID of the corresponding youtube video
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id=f"BV11X4y1c7eK", width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id=f"kemIQ_8AHdA", width=854, height=480, fs=1, rel=0)
print("Video available at https://youtube.com/watch?v=" + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
# @title Tutorial slides
# @markdown These are the slides for the videos in all tutorials today
from IPython.display import IFrame
IFrame(src=f"https://mfr.ca-1.osf.io/render?url=https://osf.io/4ye56/?direct%26mode=render%26action=download%26mode=render", width=854, height=480)
```
---
# Setup
This is a GPU free notebook!
```
# Imports
import random
import torch
import numpy as np
import matplotlib.pyplot as plt
import torch.nn as nn
import torch.optim as optim
from torch.utils.data import DataLoader, TensorDataset
from tqdm.auto import tqdm
from IPython.display import display
# @title Figure settings
import ipywidgets as widgets # interactive display
%config InlineBackend.figure_format = 'retina'
plt.style.use("https://raw.githubusercontent.com/NeuromatchAcademy/content-creation/main/nma.mplstyle")
# @title Plotting functions
def imshow(img):
img = img / 2 + 0.5 # unnormalize
npimg = img.numpy()
plt.imshow(np.transpose(npimg, (1, 2, 0)))
plt.axis(False)
plt.show()
def plot_function_approximation(x, relu_acts, y_hat):
fig, axes = plt.subplots(2, 1)
# Plot ReLU Activations
axes[0].plot(x, relu_acts.T);
axes[0].set(xlabel='x',
ylabel='Activation',
title='ReLU Activations - Basis Functions')
labels = [f"ReLU {i + 1}" for i in range(relu_acts.shape[0])]
axes[0].legend(labels, ncol = 2)
# Plot function approximation
axes[1].plot(x, torch.sin(x), label='truth')
axes[1].plot(x, y_hat, label='estimated')
axes[1].legend()
axes[1].set(xlabel='x',
ylabel='y(x)',
title='Function Approximation')
plt.tight_layout()
plt.show()
# @title Set random seed
# @markdown Executing `set_seed(seed=seed)` you are setting the seed
# for DL its critical to set the random seed so that students can have a
# baseline to compare their results to expected results.
# Read more here: https://pytorch.org/docs/stable/notes/randomness.html
# Call `set_seed` function in the exercises to ensure reproducibility.
import random
import torch
def set_seed(seed=None, seed_torch=True):
if seed is None:
seed = np.random.choice(2 ** 32)
random.seed(seed)
np.random.seed(seed)
if seed_torch:
torch.manual_seed(seed)
torch.cuda.manual_seed_all(seed)
torch.cuda.manual_seed(seed)
torch.backends.cudnn.benchmark = False
torch.backends.cudnn.deterministic = True
print(f'Random seed {seed} has been set.')
# In case that `DataLoader` is used
def seed_worker(worker_id):
worker_seed = torch.initial_seed() % 2**32
np.random.seed(worker_seed)
random.seed(worker_seed)
# @title Set device (GPU or CPU). Execute `set_device()`
# especially if torch modules used.
# inform the user if the notebook uses GPU or CPU.
def set_device():
device = "cuda" if torch.cuda.is_available() else "cpu"
if device != "cuda":
print("GPU is not enabled in this notebook. \n"
"If you want to enable it, in the menu under `Runtime` -> \n"
"`Hardware accelerator.` and select `GPU` from the dropdown menu")
else:
print("GPU is enabled in this notebook. \n"
"If you want to disable it, in the menu under `Runtime` -> \n"
"`Hardware accelerator.` and select `None` from the dropdown menu")
return device
SEED = 2021
set_seed(seed=SEED)
DEVICE = set_device()
```
---
# Section 1: Neuron Physiology and Motivation to Deep Learning
```
# @title Video 1: Biological to Artificial Neurons
# Insert the ID of the corresponding youtube video
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id=f"BV1mf4y157vf", width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id=f"ELAbflymSLo", width=854, height=480, fs=1, rel=0)
print("Video available at https://youtube.com/watch?v=" + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
```
## Section 1.1: Leaky Integrate-and-fire (LIF)
The basic idea of LIF neuron was proposed in 1907 by Louis Édouard Lapicque, long before we understood the electrophysiology of a neuron (see a translation of [Lapicque's paper](https://pubmed.ncbi.nlm.nih.gov/17968583/) ). More details of the model can be found in the book [**Theoretical neuroscience**](http://www.gatsby.ucl.ac.uk/~dayan/book/) by Peter Dayan and Laurence F. Abbott.
The model dynamics is defined with the following formula,
\begin{equation}
\frac{d V}{d t}=\left\{\begin{array}{cc}
\frac{1}{C}\left(-\frac{V}{R}+I \right) & t>t_{r e s t} \\
0 & \text { otherwise }
\end{array}\right.
\end{equation}
Note that $V$, $C$, and $R$ are the membrane voltage, capacitance, and resitance of the neuron respectively and $-\frac{V}{R}$ is the leakage current. When $I$ is sufficiently strong such that $V$ reaches a certain threshold value $V_{\rm th}$, it momentarily spikes and then $V$ is reset to $V_{\rm reset}< V_{\rm th}$, and voltage stays at $V_{\rm reset}$ for $\tau_{\rm ref}$ ms, mimicking the refractoriness of the neuron during an action potential (note that $V_{\rm reset}$ and $\tau_{\rm ref}$ is assumed to be zero in the lecture):
\begin{eqnarray}
V(t)=V_{\rm reset} \text{ for } t\in(t_{\text{sp}}, t_{\text{sp}} + \tau_{\text{ref}}]
\end{eqnarray}
where $t_{\rm sp}$ is the spike time when $V(t)$ just exceeded $V_{\rm th}$.
Thus, the LIF model captures the facts that a neuron:
- performs spatial and temporal integration of synaptic inputs
- generates a spike when the voltage reaches a certain threshold
- goes refractory during the action potential
- has a leaky membrane
For in-depth content on computational models of neurons, follow the [NMA](https://www.neuromatchacademy.org/) tutorial 1 of *Biological Neuron Models*. Specifically, for NMA-CN 2021 follow this [Tutorial](https://github.com/NeuromatchAcademy/course-content/blob/master/tutorials/W2D3_BiologicalNeuronModels/W2D3_Tutorial1.ipynb).
## Section 1.2: Simulating an LIF Neuron
In the cell below is given a function for LIF neuron model with it's arguments described.
Note that we will use Euler's method to make a numerical approximation to a derivative. Hence we will use the following implementation of the model dynamics,
\begin{equation}
V_n=\left\{\begin{array}{cc}
V_{n-1} + \frac{1}{C}\left(-\frac{V}{R}+I \right) \Delta t & t>t_{r e s t} \\
0 & \text { otherwise }
\end{array}\right.
\end{equation}
```
def run_LIF(I, T=50, dt=0.1, t_ref=10,
Rm=1, Cm=10, Vth=1, V_spike=0.5):
"""
Simulate the LIF dynamics with external input current
Args:
I : input current (mA)
T : total time to simulate (msec)
dt : simulation time step (msec)
t_ref : refractory period (msec)
Rm : resistance (kOhm)
Cm : capacitance (uF)
Vth : spike threshold (V)
V_spike : spike delta (V)
Returns:
time : time points
Vm : membrane potentials
"""
# Set up array of time steps
time = torch.arange(0, T+dt, dt)
# Set up array for tracking Vm
Vm = torch.zeros(len(time))
# Iterate over each time step
t_rest = 0
for i, t in enumerate(time):
# If t is after refractory period
if t > t_rest:
Vm[i] = Vm[i-1] + 1/Cm*(-Vm[i-1]/Rm + I) * dt
# If Vm is over the threshold
if Vm[i] >= Vth:
# Increase volatage by change due to spike
Vm[i] += V_spike
# Set up new refactory period
t_rest = t + t_ref
return time, Vm
sim_time, Vm = run_LIF(1.5)
# Plot the membrane voltage across time
plt.plot(sim_time, Vm)
plt.title('LIF Neuron Output')
plt.ylabel('Membrane Potential (V)')
plt.xlabel('Time (msec)')
plt.show()
```
### Interactive Demo 1.2: Neuron's transfer function explorer for different $R_m$ and $t_{ref}$
We know that real neurons communicate by modulating the spike count meaning that more input current causes a neuron to spike more often. Therefore, to find an input-output relationship, it makes sense to characterize their spike count as a function of input current. This is called the neuron's input-output transfer function. Let's plot the neuron's transfer function and see how it changes with respect to the **membrane resistance** and **refractory time**?
```
# @title
# @markdown Make sure you execute this cell to enable the widget!
my_layout = widgets.Layout()
@widgets.interact(Rm=widgets.FloatSlider(1., min=1, max=100.,
step=0.1, layout=my_layout),
t_ref=widgets.FloatSlider(1., min=1, max=100.,
step=0.1, layout=my_layout)
)
def plot_IF_curve(Rm, t_ref):
T = 1000 # total time to simulate (msec)
dt = 1 # simulation time step (msec)
Vth = 1 # spike threshold (V)
Is_max = 2
Is = torch.linspace(0, Is_max, 10)
spike_counts = []
for I in Is:
_, Vm = run_LIF(I, T=T, dt=dt, Vth=Vth, Rm=Rm, t_ref=t_ref)
spike_counts += [torch.sum(Vm > Vth)]
plt.plot(Is, spike_counts)
plt.title('LIF Neuron: Transfer Function')
plt.ylabel('Spike count')
plt.xlabel('I (mA)')
plt.xlim(0, Is_max)
plt.ylim(0, 80)
plt.show()
```
### Think! 1.2: Real and Artificial neuron similarities
What happens at infinite membrane resistance (Rm) and small refactory time (t_ref)? Why?
Take 10 mins to discuss the similarity between a real neuron and an artificial one with your pod.
[*Click for solution*](https://github.com/NeuromatchAcademy/course-content-dl/tree/main//tutorials/W1D3_MultiLayerPerceptrons/solutions/W1D3_Tutorial1_Solution_d58d2933.py)
---
# Section 2: The Need for MLPs
```
# @title Video 2: Universal Approximation Theorem
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id=f"BV1SP4y147Uv", width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id=f"tg8HHKo1aH4", width=854, height=480, fs=1, rel=0)
print("Video available at https://youtube.com/watch?v=" + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
```
## Coding Exercise 2: Function approximation with ReLU
We learned that one hidden layer MLPs are enough to approximate any smooth function! Now let's manually fit a sine function using ReLU activation.
We will approximate the sine function using a linear combination (a weighted sum) of ReLUs with slope 1. We need to determine the bias terms (which determines where the ReLU inflection point from 0 to linear occurs) and how to weight each ReLU. The idea is to set the weights iteratively so that the slope changes in the new sample's direction.
First, we generate our "training data" from a sine function using `torch.sine` function.
```python
>>> import torch
>>> torch.manual_seed(2021)
<torch._C.Generator object at 0x7f8734c83830>
>>> a = torch.randn(5)
>>> print(a)
tensor([ 2.2871, 0.6413, -0.8615, -0.3649, -0.6931])
>>> torch.sin(a)
tensor([ 0.7542, 0.5983, -0.7588, -0.3569, -0.6389])
```
These are the points we will use to learn how to approximate the function. We have 10 training data points so we will have 9 ReLUs (we don't need a ReLU for the last data point as we don't have anything to the right of it to model).
We first need to figure out the bias term for each ReLU and compute the activation of each ReLU where:
\begin{equation}
y(x) = \text{max}(0,x+b)
\end{equation}
We then need to figure out the correct weights on each ReLU so the linear combination approximates the desired function.
```
def approximate_function(x_train, y_train):
####################################################################
# Fill in missing code below (...),
# then remove or comment the line below to test your function
raise NotImplementedError("Complete approximate_function!")
####################################################################
# Number of relus
n_relus = x_train.shape[0] - 1
# x axis points (more than x train)
x = torch.linspace(torch.min(x_train), torch.max(x_train), 1000)
## COMPUTE RELU ACTIVATIONS
# First determine what bias terms should be for each of `n_relus` ReLUs
b = ...
# Compute ReLU activations for each point along the x axis (x)
relu_acts = torch.zeros((n_relus, x.shape[0]))
for i_relu in range(n_relus):
relu_acts[i_relu, :] = torch.relu(x + b[i_relu])
## COMBINE RELU ACTIVATIONS
# Set up weights for weighted sum of ReLUs
combination_weights = torch.zeros((n_relus, ))
# Figure out weights on each ReLU
prev_slope = 0
for i in range(n_relus):
delta_x = x_train[i+1] - x_train[i]
slope = (y_train[i+1] - y_train[i]) / delta_x
combination_weights[i] = ...
prev_slope = slope
# Get output of weighted sum of ReLU activations for every point along x axis
y_hat = ...
return y_hat, relu_acts, x
# Make training data from sine function
N_train = 10
x_train = torch.linspace(0, 2*np.pi, N_train).view(-1, 1)
y_train = torch.sin(x_train)
## Uncomment the lines below to test your function approximation
# y_hat, relu_acts, x = approximate_function(x_train, y_train)
# plot_function_approximation(x, relu_acts, y_hat)
```
[*Click for solution*](https://github.com/NeuromatchAcademy/course-content-dl/tree/main//tutorials/W1D3_MultiLayerPerceptrons/solutions/W1D3_Tutorial1_Solution_3655f212.py)
*Example output:*
<img alt='Solution hint' align='left' width=1120.0 height=832.0 src=https://raw.githubusercontent.com/NeuromatchAcademy/course-content-dl/main/tutorials/W1D3_MultiLayerPerceptrons/static/W1D3_Tutorial1_Solution_3655f212_0.png>
As you see in the top panel, we obtain 10 shifted ReLUs with the same slope. These are the basis functions that MLP uses to span the functional space, i.e., MLP finds a linear combination of these ReLUs.
---
# Section 3: MLPs in Pytorch
```
# @title Video 3: Building MLPs in PyTorch
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id=f"BV1zh411z7LY", width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id=f"XtwLnaYJ7uc", width=854, height=480, fs=1, rel=0)
print("Video available at https://youtube.com/watch?v=" + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
```
In the previous segment, we implemented a function to approximate any smooth function using MLPs. We saw that using Lipschitz continuity; we can prove that our approximation is mathematically correct. MLPs are fascinating, but before we get into the details on designing them, let's familiarize ourselves with some basic terminology of MLPs- layer, neuron, depth, width, weight, bias, and activation function. Armed with these ideas, we can now design an MLP given its input, hidden layers, and output size.
## Coding Exercise 3: Implement a general-purpose MLP in Pytorch
The objective is to design an MLP with these properties:
* works with any input (1D, 2D, etc.)
* construct any number of given hidden layers using `nn.Sequential()` and `add_module()` function
* use the same given activation function (i.e., [Leaky ReLU](https://pytorch.org/docs/stable/generated/torch.nn.LeakyReLU.html)) in all hidden layers.
**Leaky ReLU** is described by the following mathematical formula:
\begin{equation}
\text{LeakyReLU}(x) = \text{max}(0,x) + \text{negative_slope} \cdot \text{min}(0, x) =
\left\{
\begin{array}{ll}
x & ,\; \text{if} \; x \ge 0 \\
\text{negative_slope} \cdot x & ,\; \text{otherwise}
\end{array}
\right.
\end{equation}
```
class Net(nn.Module):
def __init__(self, actv, input_feature_num, hidden_unit_nums, output_feature_num):
super(Net, self).__init__()
self.input_feature_num = input_feature_num # save the input size for reshapinng later
self.mlp = nn.Sequential() # Initialize layers of MLP
in_num = input_feature_num # initialize the temporary input feature to each layer
for i in range(len(hidden_unit_nums)): # Loop over layers and create each one
####################################################################
# Fill in missing code below (...),
# then remove or comment the line below to test your function
raise NotImplementedError("Create MLP Layer")
####################################################################
out_num = hidden_unit_nums[i] # assign the current layer hidden unit from list
layer = ... # use nn.Linear to define the layer
in_num = out_num # assign next layer input using current layer output
self.mlp.add_module('Linear_%d'%i, layer) # append layer to the model with a name
actv_layer = eval('nn.%s'%actv) # Assign activation function (eval allows us to instantiate object from string)
self.mlp.add_module('Activation_%d'%i, actv_layer) # append activation to the model with a name
out_layer = nn.Linear(in_num, output_feature_num) # Create final layer
self.mlp.add_module('Output_Linear', out_layer) # append the final layer
def forward(self, x):
# reshape inputs to (batch_size, input_feature_num)
# just in case the input vector is not 2D, like an image!
x = x.view(-1, self.input_feature_num)
####################################################################
# Fill in missing code below (...),
# then remove or comment the line below to test your function
raise NotImplementedError("Run MLP model")
####################################################################
logits = ... # forward pass of MLP
return logits
input = torch.zeros((100, 2))
## Uncomment below to create network and test it on input
# net = Net(actv='LeakyReLU(0.1)', input_feature_num=2, hidden_unit_nums=[100, 10, 5], output_feature_num=1).to(DEVICE)
# y = net(input.to(DEVICE))
# print(f'The output shape is {y.shape} for an input of shape {input.shape}')
```
[*Click for solution*](https://github.com/NeuromatchAcademy/course-content-dl/tree/main//tutorials/W1D3_MultiLayerPerceptrons/solutions/W1D3_Tutorial1_Solution_cca91a84.py)
```
The output shape is torch.Size([100, 1]) for an input of shape torch.Size([100, 2])
```
## Section 3.1: Classification with MLPs
```
# @title Video 4: Cross Entropy
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id=f"BV1Ag41177mB", width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id=f"N8pVCbTlves", width=854, height=480, fs=1, rel=0)
print("Video available at https://youtube.com/watch?v=" + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
```
The main loss function we could use out of the box for multi-class classification for `N` samples and `C` number of classes is:
* CrossEntropyLoss:
This criterion expects a batch of predictions `x` with shape `(N, C)` and class index in the range $[0, C-1]$ as the target (label) for each `N` samples, hence a batch of `labels` with shape `(N, )`. There are other optional parameters like class weights and class ignores. Feel free to check the PyTorch documentation [here](https://pytorch.org/docs/stable/generated/torch.nn.CrossEntropyLoss.html) for more detail. Additionally, [here](https://sparrow.dev/cross-entropy-loss-in-pytorch/) you can learn where is appropriate to use the CrossEntropyLoss.
To get CrossEntropyLoss of a sample $i$, we could first calculate $-\log(\text{softmax(x}))$ and then take the element corresponding to $\text { labels }_i$ as the loss. However, due to numerical stability, we implement this more stable equivalent form,
\begin{equation}
\operatorname{loss}(x_i, \text { labels }_i)=-\log \left(\frac{\exp (x[\text { labels }_i])}{\sum_{j} \exp (x[j])}\right)=-x_i[\text { labels }_i]+\log \left(\sum_{j=1}^C \exp (x_i[j])\right)
\end{equation}
### Coding Exercise 3.1: Implement Batch Cross Entropy Loss
To recap, since we will be doing batch learning, we'd like a loss function that given:
* a batch of predictions `x` with shape `(N, C)`
* a batch of `labels` with shape `(N, )` that ranges from `0` to `C-1`
returns the average loss $L$ calculated according to:
\begin{align}
loss(x_i, \text { labels }_i) &= -x_i[\text { labels }_i]+\log \left(\sum_{j=1}^C \exp (x_i[j])\right) \\
L &= \frac{1}{N} \sum_{i=1}^{N}{loss(x_i, \text { labels }_i)}
\end{align}
Steps:
1. Use indexing operation to get predictions of class corresponding to the labels (i.e., $x_i[\text { labels }_i]$)
2. Compute $loss(x_i, \text { labels }_i)$ vector (`losses`) using `torch.log()` and `torch.exp()` without Loops!
3. Return the average of the loss vector
```
def cross_entropy_loss(x, labels):
# x is the model predictions we'd like to evaluate using lables
x_of_labels = torch.zeros(len(labels))
####################################################################
# Fill in missing code below (...),
# then remove or comment the line below to test your function
raise NotImplementedError("Cross Entropy Loss")
####################################################################
# 1. prediction for each class corresponding to the label
for i, label in enumerate(labels):
x_of_labels[i] = x[i, label]
# 2. loss vector for the batch
losses = ...
# 3. Return the average of the loss vector
avg_loss = ...
return avg_loss
labels = torch.tensor([0, 1])
x = torch.tensor([[10.0, 1.0, -1.0, -20.0], # correctly classified
[10.0, 10.0, 2.0, -10.0]]) # Not correctly classified
CE = nn.CrossEntropyLoss()
pytorch_loss = CE(x, labels).item()
## Uncomment below to test your function
# our_loss = cross_entropy_loss(x, labels).item()
# print(f'Our CE loss: {our_loss:0.8f}, Pytorch CE loss: {pytorch_loss:0.8f}')
# print(f'Difference: {np.abs(our_loss - pytorch_loss):0.8f}')
```
[*Click for solution*](https://github.com/NeuromatchAcademy/course-content-dl/tree/main//tutorials/W1D3_MultiLayerPerceptrons/solutions/W1D3_Tutorial1_Solution_799fb4d7.py)
```
Our CE loss: 0.34672737, Pytorch CE loss: 0.34672749
Difference: 0.00000012
```
## Section 3.2: Spiral classification dataset
Before we could start optimizing these loss functions, we need a dataset!
Let's turn this fancy-looking equation into a classification dataset
\begin{equation}
\begin{array}{c}
X_{k}(t)=t\left(\begin{array}{c}
\sin \left[\frac{2 \pi}{K}\left(2 t+k-1\right)\right]+\mathcal{N}\left(0, \sigma\right) \\
\cos \left[\frac{2 \pi}{K}\left(2 t+k-1\right)\right]+\mathcal{N}\left(0, \sigma\right)
\end{array}\right)
\end{array}, \quad 0 \leq t \leq 1, \quad k=1, \ldots, K
\end{equation}
```
def create_spiral_dataset(K, sigma, N):
# Initialize t, X, y
t = torch.linspace(0, 1, N)
X = torch.zeros(K*N, 2)
y = torch.zeros(K*N)
# Create data
for k in range(K):
X[k*N:(k+1)*N, 0] = t*(torch.sin(2*np.pi/K*(2*t+k)) + sigma*torch.randn(N))
X[k*N:(k+1)*N, 1] = t*(torch.cos(2*np.pi/K*(2*t+k)) + sigma*torch.randn(N))
y[k*N:(k+1)*N] = k
return X, y
# Set parameters
K = 4
sigma = 0.16
N = 1000
set_seed(seed=SEED)
X, y = create_spiral_dataset(K, sigma, N)
plt.scatter(X[:, 0], X[:, 1], c = y)
plt.show()
```
## Section 3.3: Training and Evaluation
```
# @title Video 5: Training and Evaluating an MLP
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id=f"BV1QV411p7mF", width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id=f"DfXZhRfBEqQ", width=854, height=480, fs=1, rel=0)
print("Video available at https://youtube.com/watch?v=" + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
```
### Coding Exercise 3.3: Implement it for a classfication task
Now that we have the Spiral dataset and a loss function, it's your turn to implement a simple train/test split for training and validation.
Steps to follow:
* Dataset shuffle
* Train/Test split (20% for test)
* Dataloader definition
* Training and Evaluation
```
def shuffle_and_split_data(X, y, seed):
# set seed for reproducibility
torch.manual_seed(seed)
# Number of samples
N = X.shape[0]
####################################################################
# Fill in missing code below (...),
# then remove or comment the line below to test your function
raise NotImplementedError("Shuffle & split data")
####################################################################
# Shuffle data
shuffled_indices = ... # get indices to shuffle data, could use torch.randperm
X = X[shuffled_indices]
y = y[shuffled_indices]
# Split data into train/test
test_size = ... # assign test datset size using 20% of samples
X_test = X[:test_size]
y_test = y[:test_size]
X_train = X[test_size:]
y_train = y[test_size:]
return X_test, y_test, X_train, y_train
## Uncomment below to test your function
# X_test, y_test, X_train, y_train = shuffle_and_split_data(X, y, seed=SEED)
# plt.scatter(X_test[:, 0], X_test[:, 1], c=y_test)
# plt.title('Test data')
# plt.show()
```
[*Click for solution*](https://github.com/NeuromatchAcademy/course-content-dl/tree/main//tutorials/W1D3_MultiLayerPerceptrons/solutions/W1D3_Tutorial1_Solution_40d9ee43.py)
*Example output:*
<img alt='Solution hint' align='left' width=1120.0 height=832.0 src=https://raw.githubusercontent.com/NeuromatchAcademy/course-content-dl/main/tutorials/W1D3_MultiLayerPerceptrons/static/W1D3_Tutorial1_Solution_40d9ee43_0.png>
And we need to make a Pytorch data loader out of it. Data loading in PyTorch can be separated in 2 parts:
* Data must be wrapped on a Dataset parent class where the methods __getitem__ and __len__ must be overrided. Note that at this point the data is not loaded on memory. PyTorch will only load what is needed to the memory. Here `TensorDataset` does this for us directly.
* Use a Dataloader that will actually read the data in batches and put into memory. Also, the option of `num_workers > 0` allows multithreading, which prepares multiple batches in the queue to speed things up.
```
g_seed = torch.Generator()
g_seed.manual_seed(SEED)
batch_size = 128
test_data = TensorDataset(X_test, y_test)
test_loader = DataLoader(test_data, batch_size=batch_size,
shuffle=False, num_workers=2,
worker_init_fn=seed_worker,
generator=g_seed)
train_data = TensorDataset(X_train, y_train)
train_loader = DataLoader(train_data, batch_size=batch_size, drop_last=True,
shuffle=True, num_workers=2,
worker_init_fn=seed_worker,
generator=g_seed)
```
Let's write a general-purpose training and evaluation code and keep it in our pocket for next tutorial as well. So make sure you review it to see what it does.
Note that `model.train()` tells your model that you are training the model. So layers like dropout, batchnorm etc. which behave different on the train and test procedures know what is going on and hence can behave accordingly. And to turn off training mode we set `model.eval()`
```
def train_test_classification(net, criterion, optimizer, train_loader,
test_loader, num_epochs=1, verbose=True,
training_plot=False, device='cpu'):
net.train()
training_losses = []
for epoch in tqdm(range(num_epochs)): # loop over the dataset multiple times
running_loss = 0.0
for i, data in enumerate(train_loader, 0):
# get the inputs; data is a list of [inputs, labels]
inputs, labels = data
inputs = inputs.to(device).float()
labels = labels.to(device).long()
# zero the parameter gradients
optimizer.zero_grad()
# forward + backward + optimize
outputs = net(inputs)
loss = criterion(outputs, labels)
loss.backward()
optimizer.step()
# print statistics
if verbose:
training_losses += [loss.item()]
net.eval()
def test(data_loader):
correct = 0
total = 0
for data in data_loader:
inputs, labels = data
inputs = inputs.to(device).float()
labels = labels.to(device).long()
outputs = net(inputs)
_, predicted = torch.max(outputs, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
acc = 100 * correct / total
return total, acc
train_total, train_acc = test(train_loader)
test_total, test_acc = test(test_loader)
if verbose:
print(f"Accuracy on the {train_total} training samples: {train_acc:0.2f}")
print(f"Accuracy on the {test_total} testing samples: {test_acc:0.2f}")
if training_plot:
plt.plot(training_losses)
plt.xlabel('Batch')
plt.ylabel('Training loss')
plt.show()
return train_acc, test_acc
```
### Think! 3.3.1: What's the point of .eval() and .train()?
Is it necessary to use `net.train()` and `net.eval()` for our MLP model? why?
[*Click for solution*](https://github.com/NeuromatchAcademy/course-content-dl/tree/main//tutorials/W1D3_MultiLayerPerceptrons/solutions/W1D3_Tutorial1_Solution_51988471.py)
Now let's put everything together and train your first deep-ish model!
```
set_seed(SEED)
net = Net('ReLU()', X_train.shape[1], [128], K).to(DEVICE)
criterion = nn.CrossEntropyLoss()
optimizer = optim.Adam(net.parameters(), lr=1e-3)
num_epochs = 100
_, _ = train_test_classification(net, criterion, optimizer, train_loader,
test_loader, num_epochs=num_epochs,
training_plot=True, device=DEVICE)
```
And finally, let's visualize the learned decision-map. We know you're probably running out of time, so we won't make you write code now! But make sure you have reviewed it since we'll start with another visualization technique next time.
```
def sample_grid(M=500, x_max=2.0):
ii, jj = torch.meshgrid(torch.linspace(-x_max, x_max, M),
torch.linspace(-x_max, x_max, M))
X_all = torch.cat([ii.unsqueeze(-1),
jj.unsqueeze(-1)],
dim=-1).view(-1, 2)
return X_all
def plot_decision_map(X_all, y_pred, X_test, y_test,
M=500, x_max=2.0, eps=1e-3):
decision_map = torch.argmax(y_pred, dim=1)
for i in range(len(X_test)):
indices = (X_all[:, 0] - X_test[i, 0])**2 + (X_all[:, 1] - X_test[i, 1])**2 < eps
decision_map[indices] = (K + y_test[i]).long()
decision_map = decision_map.view(M, M)
plt.imshow(decision_map, extent=[-x_max, x_max, -x_max, x_max], cmap='jet')
plt.show()
X_all = sample_grid()
y_pred = net(X_all)
plot_decision_map(X_all, y_pred, X_test, y_test)
```
### Think! 3.3.2: Does it generalize well?
Do you think this model is performing well outside its training distribution? Why?
[*Click for solution*](https://github.com/NeuromatchAcademy/course-content-dl/tree/main//tutorials/W1D3_MultiLayerPerceptrons/solutions/W1D3_Tutorial1_Solution_d90d9e4b.py)
What would be your suggestions to increase models ability to generalize? Think about it and discuss with your pod.
---
# Summary
In this tutorial we have explored the Multi-leayer Perceptrons (MLPs). More specifically, we have discuss the similarities of artificial and biological neural networks, we have learned the Universal Approximation Theorem, and we have implemented MLPs in PyTorch.
| github_jupyter |
# Historical Daily Temperature Analysis
## Objective
>If there is scientific evidence of extreme fluctuations in our weather patterns due to human impact to the environment then we should be able to identify factual examples of increase in the frequency in extreme temperatures.
There has been a great deal of discussion around climate change and global warming. Since NOAA has made their data public, let us explore the data ourselves and see what insights we can discover.
1. How many weather stations in US?
2. For US weather stations, what is the average years of record keeping?
3. For each US weather station, on each day of the year, identify the frequency at which daily High and Low temperature records are broken.
4. Does the historical frequency of daily temperature records (High or Low) in the US provide statistical evidence of dramatic climate change?
5. What is the average life-span of a daily temperature record (High or Low) in the US?
This analytical notebook is a component of a package of notebooks. The package is intended to serve as an exercise in the applicability of IPython/Juypter Notebooks to public weather data for DIY Analytics.
<div class="alert alert-warning">
<strong>WARNING:</strong> This notebook requires a minimum of <font color="red"><strong>12GB</strong></font> of free disk space to <strong>Run All</strong> cells for all processing phases. The time required to run all phases is **1 hr 50 mins**. The notebook supports two approaches for data generation (CSV and HDF). It is recommended that you choose to run either <emphasis>Approach 1</emphasis> or <emphasis>Approach 2</emphasis> and <font color="red"><strong>avoid</strong></font> running all cells.
</div>
## Assumptions
1. We will observe and report everything in Fahrenheit.
2. The data we extract from NOAA may be something to republish as it may require extensive data munging.
### Data
The Global Historical Climatology Network (GHCN) - [Daily dataset](http://gis.ncdc.noaa.gov/all-records/catalog/search/resource/details.page?id=gov.noaa.ncdc:C00861) integrates daily climate observations from approximately 30 different data sources. Over 25,000 worldwide weather stations are regularly updated with observations from within roughly the last month. The dataset is also routinely reconstructed (usually every week) from its roughly 30 data sources to ensure that GHCN-Daily is generally in sync with its growing list of constituent sources. During this process, quality assurance checks are applied to the full dataset. Where possible, GHCN-Daily station data are also updated daily from a variety of data streams. Station values for each daily update also undergo a suite of quality checks.
<div class="alert alert-info">
This notebook was developed using a <strong>March 16, 2015</strong> snapshot of USA-Only daily temperature readings from the Global Historical Climatology Network. The emphasis herein is on the generation of human readable data. Data exploration and analytical exercises are deferred to separate but related notebooks.
</div>
#### Ideal datasets
NOAA's [National Climatic Data Center](http://www.ncdc.noaa.gov/about-ncdc)(NCDC) is responsible for preserving, monitoring, assessing, and providing public access to the USA's climate and historical weather data and information. Since weather is something that can be observed at varying intervals, the process used by NCDC is the best that we have yet it is far from ideal. Optimally, weather metrics should be observed, streamed, stored and analyzed in real-time. Such an approach could offer the data as a service associated with a data lake.
> [Data lakes](http://www.pwc.com/us/en/technology-forecast/2014/cloud-computing/features/data-lakes.jhtml) that can scale at the pace of the cloud remove integration barriers and clear a path for more timely and informed business decisions.
Access to cloud-based data services that front-end a data lake would help to reduce the possibility of human error and divorce us from downstream processing that alters the data from it's native state.
#### Available datasets
NOAA NCDC provides public FTP access to the **GHCN-Daily dataset**, which contains a file for each US weather station. Each file contains historical daily data for the given station since that station began to observe and record. Here are some details about the available data:
* The data is delivered in a **text file based, fixed record machine-readable format**.
* Version 3 of the GHCN-Daily dataset was released in September 2012.
* Changes to the processing system associated with the Version 3 release also allowed for updates to occur 7 days a week rather than only on most weekdays.
* Version 3 contains station-based measurements from well over 90,000 land-based stations worldwide, about two thirds of which are for precipitation measurement only.
* Other meteorological elements include, but are not limited to, daily maximum and minimum temperature, temperature at the time of observation, snowfall and snow depth.
##### Citation Information
* [GHCN-Daily journal article](doi:10.1175/JTECH-D-11-00103.1): Menne, M.J., I. Durre, R.S. Vose, B.E. Gleason, and T.G. Houston, 2012: An overview of the Global Historical Climatology Network-Daily Database. Journal of Atmospheric and Oceanic Technology, 29, 897-910.
* Menne, M.J., I. Durre, B. Korzeniewski, S. McNeal, K. Thomas, X. Yin, S. Anthony, R. Ray, R.S. Vose, B.E.Gleason, and T.G. Houston, 2012: [Global Historical Climatology Network - Daily (GHCN-Daily)](http://doi.org/10.7289/V5D21VHZ), [Version 3.20-upd-2015031605], NOAA National Climatic Data Center [March 16, 2015].
## Getting Started
### Project Setup
This project is comprised of several notebooks that address various stages of the analytical process. A common theme for the project is the enablement of reproducible research. This notebook will focus on the creation of new datasets that will be used for downstream analytics.
#### Analytical Workbench
This notebook is compatible with [Project Jupyter](http://jupyter.org/).
<div class="alert" style="border: 1px solid #aaa; background: radial-gradient(ellipse at center, #ffffff 50%, #eee 100%);">
<div class="row">
<div class="col-sm-1"><img src="https://knowledgeanyhow.org/static/images/favicon_32x32.png" style="margin-top: -6px"/></div>
<div class="col-sm-11">Execution of this notebook depends on one or more features found in [IBM Knowledge Anyhow Workbench (KAWB)](https://marketplace.ibmcloud.com/apps/2149#!overview). To request a free trial account, visit us at https://knowledgeanyhow.org. You can, however, load it and view it on nbviewer or in IPython / Jupyter.</div>
</div>
</div>
```
# Import special helper functions for the IBM Knowledge Anyhow Workbench.
import kawb
```
##### Workarea Folder Structure
The project will be comprised of several file artifacts. This project's file subfolder structure is:
```
noaa_hdta_*.ipynb - Notebooks
noaa-hdta/data/ghcnd_hcn.tar.gz - Obtained from NCDC FTP site.
noaa-hdta/data/usa-daily/15mar2015/*.dly - Daily weather station files
noaa-hdta/data/hdf5/15mar2015/*.h5 - Hierarchical Data Format files
noaa-hdta/data/derived/15mar2015/*.h5 - Comma delimited files
```
**Notes**:
1. The initial project research used the 15-March-2015 snapshot of the GHCN Dataset. The folder structure allows for other snapshots to be used.
2. This notebook can be used to generate both Hierarchical Data Format and comma delimited files. It is recommended to pick one or the other as disk space requirements can be as large as:
* HDF5: <font color="red">8GB</font>
* CSV: <font color="red">4GB</font>
<div class="alert" style="border: 1px solid #aaa; background: radial-gradient(ellipse at center, #ffffff 50%, #eee 100%);">
<div class="row">
<div class="col-sm-1"><img src="https://knowledgeanyhow.org/static/images/favicon_32x32.png" style="margin-top: -6px"/></div>
<div class="col-sm-11">In IBM Knowledge Anyhow Workbench, the user workarea is located under the "resources" folder.</div>
</div>
</div>
```
%%bash
# Create folder structure
mkdir -p noaa-hdta/data noaa-hdta/data/usa-daily/15mar2015
mkdir -p noaa-hdta/data/hdf5
mkdir -p noaa-hdta/data/derived/15mar2015/missing
mkdir -p noaa-hdta/data/derived/15mar2015/summaries
mkdir -p noaa-hdta/data/derived/15mar2015/raw
mkdir -p noaa-hdta/data/derived/15mar2015/station_details
# List all project realted files and folders
!ls -laR noaa-*
```
##### Tags
KAWB allows all files to be tagged for project organization and search. This project will use the following tags.
* ```noaa_data```: Used to tag data files (.dly, .h5, .csv)
* ```noaa_hdta```: Used to tag project notebooks (.ipynb)
The following inline code can be used throughout the project to tag project files.
```
import glob
data_tagdetail = ['noaa_data',
['/resources/noaa-hdta/data/',
'/resources/noaa-hdta/data/usa-daily/15mar2015/',
'/resources/noaa-hdta/data/hdf5/15mar2015/',
'/resources/noaa-hdta/data/derived/15mar2015/'
]]
nb_tagdetail = ['noaa_hdta',['/resources/noaa_hdta_*.ipynb']]
def tag_files(tagdetail):
pathnames = tagdetail[1]
for path in pathnames:
for fname in glob.glob(path):
kawb.set_tag(fname, tagdetail[0])
# Tag Project files
tag_files(data_tagdetail)
tag_files(nb_tagdetail)
```
#### Technical Awareness
If the intended use of this notebook is to generate *Hierarchical Data Formatted* files, then the user of this notebook should be familiar with the following software concepts:
* [HDF5 Files](http://docs.h5py.org/en/latest/)
### Obtain the data
1. Copy URL below to a new browser tab and then download the ```noaa-hdta/data/ghcnd_hcn.tar.gz``` file.
```
ftp://ftp.ncdc.noaa.gov/pub/data/ghcn/daily
```
2. Upload the same file to your workbench.
3. Move and unpack the tarball into the designated project folders
```
%%bash
mv /resources/ghcnd_hcn.tar.gz /resources/noaa-hdta/data/ghcnd_hcn.tar.gz
cd /resources/noaa-hdta/data
tar -xzf ghcnd_hcn.tar.gz
mv ./ghcnd_hcn/*.dly ./usa-daily/15mar2015/
rm -R ghcnd_hcn
ls -la
```
<div class="alert" style="border: 1px solid #aaa; background: radial-gradient(ellipse at center, #ffffff 50%, #eee 100%);">
<div class="row">
<div class="col-sm-1"><img src="https://knowledgeanyhow.org/static/images/favicon_32x32.png" style="margin-top: -6px"/></div>
<div class="col-sm-11">In IBM Knowledge Anyhow Workbench, you can drag/drop the file on your workbench browser tab to simplify the uploading process.</div>
</div></div>
```
%%bash
# Provide the inline code for obtaining the data. See step 3 above.
```
### Dependencies
If the intended use of this notebook is to generate *Hierarchical Data Formatted* files, then this notebook requires the installation of the following software dependencies:
```
$ pip install h5py
```
```
# Provide the inline code necessary for loading any required libraries
```
## Tidy Data Check
Upon review of the NOAA NCDC GHCN Dataset, the data **fails** to meet the requirements of [Tidy Data](http://www.jstatsoft.org/v59/i10/paper). A dataset is tidy if rows, columns and tables are matched up with observations, variables and types. In tidy data:
1. Each variable forms a column.
2. Each observation forms a row.
3. Each type of observational unit forms a table.
Messy data is any other arrangement of the data.
In the case of the GHCN Dataset, we are presented with datasets that contain observations for each day in a month for a given year. Each ".dly" file contains data for one station. The name of the file corresponds to a station's identification code. Each record in a file contains one month of daily data. Each row contains observations for more than 20 different element types. The variables on each line include the following:
```
------------------------------
Variable Columns Type
------------------------------
ID 1-11 Character
YEAR 12-15 Integer
MONTH 16-17 Integer
ELEMENT 18-21 Character
VALUE1 22-26 Integer
MFLAG1 27-27 Character
QFLAG1 28-28 Character
SFLAG1 29-29 Character
VALUE2 30-34 Integer
MFLAG2 35-35 Character
QFLAG2 36-36 Character
SFLAG2 37-37 Character
. . .
. . .
. . .
VALUE31 262-266 Integer
MFLAG31 267-267 Character
QFLAG31 268-268 Character
SFLAG31 269-269 Character
------------------------------
```
A more detailed interpretation of the format of the data is outlined in ```readme.txt``` here ftp://ftp.ncdc.noaa.gov/pub/data/ghcn/daily.
### Sample Preview
The variables of interest to this project have the following definitions:
* **ID** is the station identification code. See "ghcnd-stations.txt" for a complete list of stations and their metadata.
* **YEAR** is the year of the record.
* **MONTH** is the month of the record.
* **ELEMENT** is the element type. There are more than 20 possible elements. The five core elements are:
* *PRCP* = Precipitation (tenths of mm)
* *SNOW* = Snowfall (mm)
* *SNWD* = Snow depth (mm)
* *TMAX* = Maximum temperature (tenths of degrees C)
* *TMIN* = Minimum temperature (tenths of degrees C)
* **VALUE(n)** is element value on the n<sup>th</sup> day of the month (missing = -9999).
* **NAPAD(n)** contains non-applicable fields of interest.
where **n** denotes the day of the month (1-31) If the month has less than 31 days, then the remaining variables are set to missing (e.g., for April, VALUE31 = -9999, NAPAD31 = {MFLAG31 = blank, QFLAG31 = blank, SFLAG31 = blank}).
Here is a snippet depicting how the data is represented:
```
USC00011084201409TMAX 350 H 350 H 344 H 339 H 306 H 333 H 328 H 339 H 339 H 322 H 339 H 339 H 339 H 333 H 339 H 333 H 339 H 328 H 322 H 328 H 283 H 317 H 317 H 272 H 283 H 272 H 272 H 272 H-9999 -9999 -9999
USC00011084201409TMIN 217 H 217 H 228 H 222 H 217 H 217 H 222 H 233 H 233 H 228 H 222 H 222 H 217 H 211 H 217 H 217 H 211 H 206 H 200 H 189 H 172 H 178 H 122 H 139 H 144 H 139 H 161 H 206 H-9999 -9999 -9999
USC00011084201409TOBS 217 H 256 H 233 H 222 H 217 H 233 H 239 H 239 H 233 H 278 H 294 H 256 H 250 H 228 H 222 H 222 H 211 H 206 H 211 H 194 H 217 H 194 H 139 H 161 H 144 H 194 H 217 H 228 H-9999 -9999 -9999
USC00011084201409PRCP 0 H 0 H 0 H 13 H 25 H 8 H 0 H 0 H 0 H 0 H 0 H 0 H 0 H 0 H 0 H 25 H 178 H 0 H 0 H 56 H 0 H 0 H 0 H 0 H 0 H 0 H 0 H 0 H-9999 -9999 -9999
```
### Remove noise
The NOAA NCDC GHCN Dataset includes ```-9999``` as an indicator of missing observation data. We will take this into consideration as we transform the data into a usable format for the project.
## Data Processing Decision
Before we can attempt to answer the questions driving this project, we must first map the data into a more reasonable format. As a result, the focus of this notebook will be the creation of new datasets that can be consumed by other notebooks in the project.
> [Data munging](http://en.wikipedia.org/wiki/Data_wrangling) or data wrangling is loosely the process of manually converting or mapping data from one "raw" form into another format that allows for more convenient consumption of the data."
This data munging endeavor is an undertaking unto itself since our goal here is to extract from the machine-readable content new information and store it in a human-readable format. Essentially, we seek to explode (decompress) the data for general consumption and [normalize](http://en.wikipedia.org/wiki/Database_normalization#Normal_forms) it into a relational model that may be informative to users.
### Desired Data Schema
#### Historical Daily Summary
The goal here is to capture summary information about a given day throughout history at a specific weather station in the US. This dataset contains 365 rows where each row depicts the aggregated low and high temperature data for a specific day throughout the history of the weather station.
Column | Description
--- | ---
Station ID | Name of the US Weather Station
Month |Month of the observations
Day | Day of the observations
FirstYearOfRecord | First year that this weather station started collecting data for this day in in history.
TMin | Current record low temperature (F) for this day in history.
TMinRecordYear | Year in which current record low temperature (F) occurred.
TMax | Current record high temperature (F) for this day in history.
TMaxRecordYear | Year in which current record high temperature occurred.
CurTMinMaxDelta | Difference in degrees F between record high and low records for this day in history.
CurTMinRecordDur | LifeSpan of curent low record temperature.
CurTMaxRecordDur | LifeSpan of current high record temperature.
MaxDurTMinRecord | Maximum years a low temperature record was held for this day in history.
MinDurTMinRecord | Minimum years a low temperature record was held for this day in history.
MaxDurTMaxRecord | Maximum years a high temperature record was held for this day in history.
MinDurTMaxRecord | Minimum years a high temperature record was held for this day in history.
TMinRecordCount | Total number of TMin records set on this day (does not include first since that may not be a record).
TMaxRecordCount | Total number of TMax records set on this day (does not include first since that may not be a record).
#### Historical Daily Detail
The goal here is to capture details for each year that a record has changed for a specific weather station in the US. During the processing of the Historical Daily Summary dataset, we will log each occurrence of a new temperature record. Information in this file can be used to drill-down into and/or validate the summary file.
Column | Description
--- | ---
Station ID | Name of the US Weather Station
Year | Year of the observation
Month |Month of the observation
Day | Day of the observation
Type | Type of temperature record (Low = *TMin*, High = *TMax*)
OldTemp | Temperature (F) prior to change.
NewTemp | New temperature (F) record for this day.
TDelta | Delta between old and new temperatures.
#### Historical Daily Missing Record Detail
The goal here is to capture details pertaining to missing data. Each record in this dataset represents a day in history that a specific weather station in the USA failed to observe a temperature reading.
Column | Description
--- | ---
Station ID | Name of the US Weather Station
Year | Year of the missing observation
Month |Month of the missing observation
Day | Day of the missing observation
Type | Type of temperature missing (Low = *TMin*, High = *TMax*)
#### Historical Raw Detail
The goal here is to capture raw daily details. Each record in this dataset represents a specific temperature observation for a day in history for a specific that a specific weather station.
Column | Description
--- | ---
Station ID | Name of the US Weather Station
Year | Year of the observation
Month |Month of the observation
Day | Day of the observation
Type | Type of temperature reading (Low = *TMin*, High = *TMax*)
FahrenheitTemp | Fahrenheit Temperature
### Derived Datasets
While this notebook is focused on daily temperature data, we could imagine future work associated with other observation types like snow accumulations and precipitation. Therefore, the format we choose to capture and store our desired data should also allow us to organize and append future datasets.
The [HDF5 Python Library](http://docs.h5py.org/en/latest/) provides support for the standard Hierarchical Data Format. This library will allow us to:
1. Create, save and publish derived datasets for reuse.
2. Organize our datasets into groups (folders).
3. Create datasets that can be easily converted to/from dataframes.
However, HDF5 files can be very large which could be a problem if we want to share the data. Alternatively, we could store the information in new collections of CSV files where each ```.csv``` contained weather station specific content for one of our target schemas.
# Extract, Transform and Load (ETL)
The focus of this notebook is to perform the to compute intensive activities that will **explode** the **text file based, fixed record machine-readable format** into the derived data schemas we have specified.
### Gather Phase 1
The purpose of this phase will be to do the following:
* For each daily data file (*.dly) provided by NOAA:
* For each record where ELEMENT == TMAX or TMIN
* Identify missing daily temperature readings, write them to the missing dataset.
* Convert each daily temperature reading from celcius to fahrenheit, write each daily reading to the raw dataset.
#### Approach 1: Comma delimited files
Use the code below to layout your target project environment (if it should differ from what is described herein) and then run the process for *Gather Phase 1*. This will take about 20 minutes to process the **1218** or more weather station files. You should expect to see output like this:
```
>> Processing file 0: USC00207812
Extracting content from file /resources/noaa-hdta/data/usa-daily/15mar2015/USC00207812.dly.
>> Processing Complete: 7024 lines of file /resources/noaa-hdta/data/usa-daily/15mar2015/USC00207812.dly.
>> Elapsed file execution time 0:00:00
>> Processing file 1: USC00164700
Extracting content from file /resources/noaa-hdta/data/usa-daily/15mar2015/USC00164700.dly.
>> Processing Complete: 9715 lines of file /resources/noaa-hdta/data/usa-daily/15mar2015/USC00164700.dly.
.
.
.
>> Processing file 1217: USC00200230
Extracting content from file /resources/noaa-hdta/data/usa-daily/15mar2015/USC00200230.dly.
>> Processing Complete: 10112 lines of file /resources/noaa-hdta/data/usa-daily/15mar2015/USC00200230.dly.
>> Elapsed file execution time 0:00:00
>> Processing Complete.
>> Elapsed corpus execution time 0:19:37
```
<div class="alert" style="border: 1px solid #aaa; background: radial-gradient(ellipse at center, #ffffff 50%, #eee 100%);">
<div class="row">
<div class="col-sm-1"><img src="https://knowledgeanyhow.org/static/images/favicon_32x32.png" style="margin-top: -6px"/></div>
<div class="col-sm-11">[IBM Knowledge Anyhow Workbench (KAWB)](https://marketplace.ibmcloud.com/apps/2149#!overview) provides support for importable notebooks that address the challenges of code reuse. It also supports the concept of code injection from reusable cookbooks. For more details, please refer to the Share and Reuse tutorials that come with your instance of KAWB.</div>
</div>
</div>
```
import mywb.noaa_hdta_etl_csv_tools as csvtools
csvtools.help()
```
%inject csvtools.noaa_run_phase1_approach1
```
# Approach 1 Content Layout for Gather Phases 1 and 2
htda_approach1_content_layout = {
'Content_Version': '15mar2015',
'Daily_Input_Files': '/resources/noaa-hdta/data/usa-daily/15mar2015/*.dly',
'Raw_Details': '/resources/noaa-hdta/data/derived/15mar2015/raw',
'Missing_Details': '/resources/noaa-hdta/data/derived/15mar2015/missing',
'Station_Summary': '/resources/noaa-hdta/data/derived/15mar2015/summaries',
'Station_Details': '/resources/noaa-hdta/data/derived/15mar2015/station_details',
}
# Run Gather Phase 1 for all 1218 files using approach 1 (CSV)
csvtools.noaa_run_phase1_approach1(htda_approach1_content_layout)
```
##### Observe Output
You can compute the disk capacity of your *Gather Phase 1* results.
```
96M /resources/noaa-hdta/data/derived/15mar2015/missing
24M /resources/noaa-hdta/data/derived/15mar2015/summaries
3.2G /resources/noaa-hdta/data/derived/15mar2015/raw
```
```
%%bash
# Compute size of output folders
du -h --max-depth=1 /resources/noaa-hdta/data/derived/15mar2015/
```
#### Approach 2: HDF5 files
Use the code below to layout your target project environment (if it should differ from what is described herein) and then run the process for *Gather Phase 1*. It will take less than 20 minutes to process the **1218** or more weather station files. **Note**: You will need to have room for about <font color="red">**6.5GB**</font>. You should expect to see output like this:
```
>> Processing file 0: USC00207812
Extracting content from file /resources/noaa-hdta/data/usa-daily/15mar2015/USC00207812.dly.
>> Processing Complete: 7024 lines of file /resources/noaa-hdta/data/usa-daily/15mar2015/USC00207812.dly.
>> Elapsed file execution time 0:00:00
>> Processing file 1: USC00164700
Extracting content from file /resources/noaa-hdta/data/usa-daily/15mar2015/USC00164700.dly.
>> Processing Complete: 9715 lines of file /resources/noaa-hdta/data/usa-daily/15mar2015/USC00164700.dly.
.
.
.
>> Processing file 1217: USC00200230
Extracting content from file /resources/noaa-hdta/data/usa-daily/15mar2015/USC00200230.dly.
>> Processing Complete: 10112 lines of file /resources/noaa-hdta/data/usa-daily/15mar2015/USC00200230.dly.
>> Elapsed file execution time 0:00:00
>> Processing Complete.
>> Elapsed corpus execution time 0:17:43
```
```
import mywb.noaa_hdta_etl_hdf_tools as hdftools
hdftools.help()
```
%inject hdftools.noaa_run_phase1_approach2
```
# Approach 2 Content Layout for Gather Phases 1 and 2
htda_approach2_content_layout = {
'Content_Version': '15mar2015',
'Daily_Input_Files': '/resources/noaa-hdta/data/usa-daily/15mar2015/*.dly',
'Raw_Details': '/resources/noaa-hdta/data/hdf5/15mar2015/noaa_ncdc_raw_details.h5',
'Missing_Details': '/resources/noaa-hdta/data/hdf5/15mar2015/noaa_ncdc_missing_details.h5',
'Station_Summary': '/resources/noaa-hdta/data/hdf5/15mar2015/noaa_ncdc_station_summaries.h5',
'Station_Details': '/resources/noaa-hdta/data/hdf5/15mar2015/noaa_ncdc_station_details.h5',
}
# Run Gather Phase 1 for all 1218 files using approach 2 (HDF5)
hdftools.noaa_run_phase1_approach2(htda_approach2_content_layout)
%%bash
# Compute size of output folders
du -h --max-depth=1 /resources/noaa-hdta/data/hdf5/15mar2015/
```
### Gather Phase 2
The purpose of this phase will be to do the following:
* For each daily data file (*.dly) provided by NOAA:
* For each record where ELEMENT == TMAX or TMIN
* Identify missing daily temperature readings, write them to the missing dataset.
* Convert each daily temperature reading from celcius to fahrenheit, write each daily reading to the raw dataset.
* For each raw dataset per weather station that was generated in *Gather Phase 1*:
* For each tuple in a dataset, identify when a new temperature record has occurred (TMIN or TMAX):
* Create a Station Detail tuple and add to list of tuples
* Update the Summary Detail list of tuples for this day in history for this station
* Store the lists to disk
#### Approach 1: Comma delimited files
Use the code below to layout your target project environment (if it should differ from what is described herein) and then run the process for *Gather Phase 2*. This will take about 40 minutes to process the **1218** or more raw weather station files. You should expect to see output like this:
```
Processing dataset 0 - 1218: USC00011084
Processing dataset 1 - 1218: USC00012813
.
.
.
Processing dataset 1216: USC00130133
Processing dataset 1217: USC00204090
>> Processing Complete.
>> Elapsed corpus execution time 0:38:47
```
```
# Decide if we need to generate station detail files.
csvtools.noaa_run_phase2_approach1.help()
# Run Gather Phase 2 for all 1218 raw files using approach 1 (CSV)
csvtools.noaa_run_phase2_approach1(htda_approach1_content_layout, create_details=True)
```
##### Observe Output
You can compute the disk capacity of your *Gather Phase 2* results.
```
96M /resources/noaa-hdta/data/derived/15mar2015/missing
24M /resources/noaa-hdta/data/derived/15mar2015/summaries
3.2G /resources/noaa-hdta/data/derived/15mar2015/raw
129M /resources/noaa-hdta/data/derived/15mar2015/station_details
3.4G /resources/noaa-hdta/data/derived/15mar2015/
```
```
%%bash
# Compute size of output folders
du -h --max-depth=1 /resources/noaa-hdta/data/derived/15mar2015/
```
#### Approach 2: HDF5 files
Use the code below to layout your target project environment (if it should differ from what is described herein) and then run the process for *Gather Phase 2*. This will take about 30 minutes to process the **1218** or more raw weather station files. **Note**: You will need to have room for about <font color="red">**6.5GB**</font>. You should expect to see output like this:
```
Fetching keys for type = raw_detail
>> Fetch Complete.
>> Elapsed key-fetch execution time 0:00:09
Processing dataset 0 - 1218: USC00011084
Processing dataset 1 - 1218: USC00012813
.
.
.
Processing dataset 1216 - 1218: USW00094794
Processing dataset 1217 - 1218: USW00094967
>> Processing Complete.
>> Elapsed corpus execution time 0:28:48
```
%inject hdftools.noaa_run_phase2_approach2
### noaa_run_phase2_approach2
Takes a dictionary of project folder details to drive the processing of *Gather Phase 2 Approach 2* using **HDF files**.
#### Disk Storage Requirements
* This function creates a **Station Summaries** dataset that requires ~2GB of free space.
* This function can also create a **Station Details** dataset. If you require this dataset to be generated, modify the call to ```noaa_run_phase2_approach2()``` with ```create_details=True```. You will need additional free space to support this feature. Estimated requirement: <font color="red">**5GB**</font>
```
# Run Gather Phase 2 for all 1218 files using approach 2 (HDF)
hdftools.noaa_run_phase2_approach2(htda_approach2_content_layout)
```
##### Observe Output TODO
You can compute the disk capacity of your *Gather Phase 2* results.
```
HDF File Usage (Phases 1 & 2) - Per File and Total
4.9G /resources/noaa-hdta/data/hdf5/15mar2015/noaa_ncdc_raw_details.h5
1.4G /resources/noaa-hdta/data/hdf5/15mar2015/noaa_ncdc_missing_details.h5
1.3G /resources/noaa-hdta/data/hdf5/15mar2015/noaa_ncdc_station_summaries.h5
7.4G /resources/noaa-hdta/data/hdf5/15mar2015/
```
```
%%bash
# Compute size of output folders
echo "HDF File Usage (Phases 1 & 2) - Per File and Total"
du -ah /resources/noaa-hdta/data/hdf5/15mar2015/
```
# References
* [Data Analysis Workflow Navigation repository](https://github.com/vinomaster/dawn): This notebook outline was derived from the **Research Analysis Navigation Template**.
* [NOAA National Climatic Data Center](http://www.ncdc.noaa.gov/)
* [NOAA Data Fraud News](https://stevengoddard.wordpress.com/2013/01/11/noaa-temperature-fraud-expands-part-1/)
* [HDF5](http://www.h5py.org/)
* [HDF5 v. Database](http://nbviewer.ipython.org/github/dolaameng/tutorials/blob/master/ml-tutorials/BASIC_pandas_io%28specially%20hdf5%29.ipynb)
# Summary
This notebook provides two approaches in the creation of human readable datasets for historical daily temperature analytics. This analytical notebook is a component of a package of notebooks. The tasks addressed herein focused on data munging activities to produce a desired set of datasets for several predefined schemas. These datasets can now be used in other package notebooks for data exploration activities.
This notebook has embraced the concepts of reproducible research and can be shared with others so that they can recreate the data locally.
# Future Investigations
1. Fix the ordering of the record lifespan calculations. See code and comments for clarity.
2. Fix FTP link so that it can be embedded into notebook.
2. Explore multi-level importable notebooks to allow the tools files to share common code.
<div class="alert" style="border: 1px solid #aaa; background: radial-gradient(ellipse at center, #ffffff 50%, #eee 100%);">
<div class="row">
<div class="col-sm-1"><img src="https://knowledgeanyhow.org/static/images/favicon_32x32.png" style="margin-top: -6px"/></div>
<div class="col-sm-11">This notebook was created using [IBM Knowledge Anyhow Workbench](https://knowledgeanyhow.org). To learn more, visit us at https://knowledgeanyhow.org.</div>
</div>
</div>
| github_jupyter |

### Egeria Hands-On Lab
# Welcome to the Open Lineage Lab
## Introduction
Egeria is an open source project that provides open standards and implementation libraries to connect tools, catalogs and platforms together so they can share information (called metadata) about data and the technology that supports it.
In this hands-on lab you will get a chance to work with Egeria metadata and governance servers and learn how to manually create metadata to describe lineage for data movement processes. For this purpose we use **Open Lineage Services** governance server solution designed to capture and manage a historical warehouse of lineage information.
We will also show how using General **Egeria UI** you can search data assets and visualize lineage previously created.
To read more about lineage concepts in Egeria, see https://egeria.odpi.org/open-metadata-publication/website/lineage/.
## The Scenario
The Egeria team use the personas and scenarios from the fictitious company called Coco Pharmaceuticals. (See https://opengovernance.odpi.org/coco-pharmaceuticals/ for more information).
On their business transformation journey after they successfully created data catalog for the data lake, new challenge emerges. Due to regulatory requirements, business came up with request to improve data traceability. Introducing data lineage for critical data flows was ideal use-case for the next level of maturity in their governance program.
In this lab we discover how to manually catalogue data assets in the data lake and describe data movement for simple data transformation process executed by their in-house built ETL tool. Finally, the users can find data assets and visualize end to end lineage in the web UI.
Peter Profile and Erin Overview got assigned to work on a solution to capture and report data lineage using Egeria.
## Setting up
Coco Pharmaceuticals make widespread use of Egeria for tracking and managing their data and related assets.
Figure 1 below shows their metadata servers and the Open Metadata and Governance (OMAG) Server Platforms that are hosting them. Each metadata server supports a department in the organization. The servers are distributed across the platform to even out the workload. Servers can be moved to a different platform if needed.

> **Figure 1:** Coco Pharmaceuticals' OMAG Server Platforms
For the scope of this lab, we are going to interact with two servers hosted on Data Lake platform:
- `cocoMDS1` as metadata repository to store all the assets;
- `cocoOLS1` as dedicated governance server to enable open lineage services and historical lineage repository;
- `UI platform` running the APIs to support Egeria UI application.
> **Important**: When running this lab using [`kubernetes deployment`](https://odpi.github.io/egeria-docs/guides/admin/kubernetes/chart_lab/) the UI Platform is already configured and started for you.
The code below checks that the platforms are running. It checks that the servers are configured and then if they are running on the platform. If a server is configured, but not running, it will start it.
Look for the "Done." message. This appears when `environment-check` has finished.
```
%run ../common/environment-check.ipynb
```
## Excercise 1
### Capturing lineage manually
In this exercise Peter and Erin will start with minimal use-case and execute steps to create lineage manually. They are looking at simple high level transformation activity implemented using CocoETL, in-house developed ETL tool that uses python scripting language. Files from previous clinical trials are stored on server location accessible by the tool. `ConvertFileToCSV` is script that reads file coming out of legacy system of records and transform it to csv file structure.

> Figure 2: Simple asset lineage
For use-cases like this one, **Data Engine Access Service (OMAS)** API seems perfect match. It enables external data platforms, tools or engines to interact with Egeria and share metadata needed to construct lineage graph.
#### Check if assets are present in the catalog
At first, Erin wants to be sure upfront that the assets are not present in the catalog. She uses Egeria UI Asset Catalog search option but fist she needs to log in.
> **Important:** When running this lab using kubernetes deployment, make sure that you [expose the Egeria UI](https://odpi.github.io/egeria-docs/guides/admin/kubernetes/chart_lab/#accessing-the-egeria-ui) running in the container to your local network and access it via localhost.
To access Egeria UI go to https://localhost:8443/
username: erinoverview
password: secret

> **Figure 3** Log on as Erin Overview
Once in, from the top navigation bar she clicks on `Search` and navigates to *Asset Catalog* search page.

> **Figure 4** Navigate to Asset Catalog search page
Erin already knows the descriptive name of the data file asset in interest so she inputs the text "archive" in the search box and selects type `Asset` from the list.

> **Figure 5** Assets search
The UI responds with message that no assets are found with the input provided. This is expected since at this moment the assets are not yet created.
#### Adding assets in the catalog
Peter is now ready to start creating assets using API calls. He is using Data Engine Access Service (OMAS) REST API available on Data Lake Platform `cocoMDS1` metadata server.
```
platformURL = dataLakePlatformURL
serverName = "cocoMDS1"
```
To ba able to call Data Engine OMAS endpoints, parameters like unique qualified name of the tool and service account are required.
```
cocoETLName = "CocoPharma/DataEngine/CocoETL"
cocoETLUser = "cocoETLnpa"
dataEngineOMASEndpoint = platformURL + '/servers/' + serverName + '/open-metadata/access-services/data-engine/users/' + cocoETLUser
```
##### Step 1 - Register the tool
External systems interacting with Egeria using Data Engine OMAS need to be registered first. This step is required only once as long as the cocoETLName does not change.
In our case, to register the tool properly Peter provides descriptive information that will be useful for others to understand as many details possible about the characteristics of the external source of metadata.
```
url = dataEngineOMASEndpoint + '/registration'
requestBody = {
"dataEngine":
{
"qualifiedName": cocoETLName,
"displayName": "CocoETL",
"description": "Requesting to register external data engine capability for Coco Pharmaceuticals in-house Data Platform ETL tool CocoETL.",
"engineType": "DataEngine",
"engineVersion": "1",
"enginePatchLevel": "0",
"vendor": "Coco Pharmaceuticals",
"version": "1",
"source": "CocoPharma"
}
}
print(requestBody)
postAndPrintResult(url, json=requestBody, headers=None)
```
At this point, the tool is properly registered and its name can be used as *externalSourceName* further on.
> Note: This information gets stored as [`SoftwareServerCapability`](https://egeria.odpi.org/open-metadata-publication/website/open-metadata-types/0042-Software-Server-Capabilities.html) in Egeria.
##### Step 2 - Create file assets
Lets look at the files. They are stored in well know server location defined by the networkAddress and filesystem location.
```
networkAddress = "filesrv01.coco.net"
filesRoot = "file://secured/research/previous-clinical-trials/"
```
Peter onboards the source file `old-archive.dat`. He is using [`DataFile`](https://egeria.odpi.org/open-metadata-publication/website/open-metadata-types/0220-Files-and-Folders.html) as fileType.
```
url = dataEngineOMASEndpoint + '/data-files'
fileName1 = "old-archive.dat"
filePath1 = filesRoot + fileName1
fileQualifiedName1 = filePath1 + "@" + cocoETLName
requestFileBody = {
"externalSourceName": cocoETLName,
"file": {
"fileType": "DataFile",
"qualifiedName": fileQualifiedName1,
"displayName": fileName1,
"pathName": filePath1,
"networkAddress": networkAddress,
"columns": []
}
}
print(requestFileBody)
postAndPrintResult(url, json=requestFileBody, headers=None)
```
Next, he calls the same endpoint but this time for the destination file `old-archive.csv`. He is using [`CSVFile`](https://egeria.odpi.org/open-metadata-publication/website/open-metadata-types/0220-Files-and-Folders.html) as fileType.
```
url = dataEngineOMASEndpoint + '/data-files'
fileName2 = "old-archive.csv"
filePath2 = filesRoot + fileName2
fileQualifiedName2 = filePath2 + "@" + cocoETLName
requestFileBody = {
"externalSourceName": cocoETLName,
"file": {
"fileType": "CSVFile",
"qualifiedName": fileQualifiedName2,
"displayName": fileName2,
"pathName": filePath2,
"networkAddress": networkAddress,
"columns": []
}
}
print(requestFileBody)
postAndPrintResult(url, json=requestFileBody, headers=None)
```
> Note that in both calls, that the columns are not provided because in this exercise we are only focusing on the high level lineage without providing schema level details.
##### Step 3 - Create process assets
Using adequate name and description for the activity, he then requests new asset to represent the process.
```
url = dataEngineOMASEndpoint + '/processes'
activityName = "ConvertFileToCSV"
processQualifiedName = activityName + "@" + cocoETLName
requestProcessBody = {
"process":
{
"qualifiedName": processQualifiedName,
"displayName": activityName,
"name": activityName,
"description": "Process named 'ConvertFileToCSV' representing high level processing activity performed by CocoETL tool.",
"owner": cocoETLUser,
"updateSemantic": "REPLACE"
},
"externalSourceName": cocoETLName
}
print(requestProcessBody)
postAndPrintResult(url, json=requestProcessBody, headers=None)
```
Well done. At this point all the assets are stored in the catalog.
#### Adding lineage mappings in the catalog
Finally, he needs to send the lineage mappings connecting the assets. This is done using their fully qualified names.
```
url = dataEngineOMASEndpoint + '/lineage-mappings'
requestLineageMappingsBody = {
"lineageMappings": [
{
"sourceAttribute": fileQualifiedName1,
"targetAttribute": processQualifiedName
},
{
"sourceAttribute": processQualifiedName,
"targetAttribute": fileQualifiedName2
}
],
"externalSourceName": cocoETLName
}
print(requestLineageMappingsBody)
postAndPrintResult(url, json=requestLineageMappingsBody, headers=None)
```
#### Finding assets in the UI and showing lineage
Erin is ready to inspect the catalog again. She goes back to the search page and searches the text "archive".
This time, she is able to find the file assets Peter created in the previous steps.
> Tip: Once logged on, Erin can directly navigate to the search results using https://localhost:8443/asset-catalog/search?q=archive&types=Asset

Clicking one of the file names, she can access the details page.

To inspect the lineage graph, Erin clicks on `End2End`.

This step completes **Exercise 1**.
| github_jupyter |
# The Marmousi elastic model
Let's make a synthetic dataset and write it out to a SEG-Y file.
```
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
from obspy.io.segy.segy import _read_segy
ls -l ../data/marmousi
!cat ../data/marmousi/README
```
<hr />
We're going to make our own synthetic.
## Read the elastic data
```
velocity = _read_segy('../data/marmousi/velocity.segy')
density = _read_segy('../data/marmousi/density.segy')
velocity.binary_file_header
s = velocity.textual_file_header.decode()
print('\n'.join(map(''.join, zip(*[iter(s)]*80))))
def get_data(section):
nsamples = section.binary_file_header.number_of_samples_per_data_trace
dt = section.binary_file_header.sample_interval_in_microseconds
ntraces = len(section.traces) # Get from actual file contents.
tbase = np.arange(0, nsamples * dt / 1000.0, dt)
vol = np.stack(t.data for t in section.traces)
ma, mi = np.amax(vol), np.amin(vol)
print('number of traces (ntraces) :', ntraces)
print('samples per trace (nsamples) :', nsamples)
print('sample interval :', dt)
print('data range : {} - {}'.format(mi, ma))
return tbase, vol
t, vp = get_data(velocity)
_, rho = get_data(density)
```
## Deal with weirdness
The data are transposed: the so-called traces in the ObsPy stream are really timeslices, not traces. Very strange. We'll fix it.
```
vp.shape
vp = vp.T
rho = rho.T
nsamples = 751
t = np.linspace(0, (nsamples-1)*0.004, nsamples)
vp.shape
```
Now the data are organized correctly. We have to transpose them to display them, but the first dimension is traces.
```
fig = plt.figure(figsize = (20,20))
ax1 = fig.add_subplot(121)
ax1.set_title(r'$V_\mathrm{P}\ \ \mathrm{[m/s]}$')
img = ax1.imshow(vp.T, aspect=2, cmap='viridis')
plt.colorbar(img, shrink=0.2)
ax2 = fig.add_subplot(122)
ax2.set_title(r'$\rho\ \ \mathrm{[kg/m^3]}$')
img = ax2.imshow(rho.T, aspect=2, cmap='viridis')
plt.colorbar(img, shrink=0.2)
plt.show()
```
## Make a synthetic
```
imp = vp * rho
def make_rc(imp):
upper = imp[:, :-1]
lower = imp[:, 1: ]
return (lower - upper) / (lower + upper)
rc = make_rc(imp)
plt.imshow(rc.T, aspect=2, cmap="Greys")
plt.show()
from scipy.signal import ricker
f = 25
wavelet = ricker(100, 1000/(4*f))
plt.plot(wavelet)
plt.show()
def convolve(trace):
return np.convolve(trace, wavelet, mode='same')
synth = np.apply_along_axis(convolve,
axis=-1,
arr=rc)
plt.figure(figsize=(12,6))
plt.imshow(synth.T, cmap="Greys", aspect=2)
plt.colorbar()
plt.show()
np.amin(synth), np.amax(synth)
synth.dtype
```
## Write a SEG-Y file
```
from obspy.core import Trace, Stream, UTCDateTime
from obspy.io.segy.segy import SEGYTraceHeader
from obspy.core import AttribDict
from obspy.io.segy.segy import SEGYBinaryFileHeader
```
If you're on a 64-bit machine and Python environment, the data are currently 64-bit — 8 bytes per sample. We don't want or need this many bytes, so we'll reduce the bit-depth to 32-bit:
```
synth = synth.astype(np.float32)
synth.dtype
stream = Stream()
# Add the traces to the stream.
for i, trace in enumerate(synth):
# Make the trace.
tr = Trace(trace)
# Add required data.
tr.stats.delta = 0.004
tr.stats.starttime = 0 # Not strictly required.
# Add yet more to the header (optional).
tr.stats.segy = {'trace_header': SEGYTraceHeader()}
tr.stats.segy.trace_header.trace_sequence_number_within_line = i + 1
tr.stats.segy.trace_header.receiver_group_elevation = 0
# Append the trace to the stream.
stream.append(tr)
# Add a text header.
stream.stats = AttribDict()
stream.stats.textual_file_header = '{:80s}'.format('This is the textual header.').encode()
stream.stats.textual_file_header += '{:80s}'.format('This file contains seismic data.').encode()
# Add a binary header.
stream.stats.binary_file_header = SEGYBinaryFileHeader()
stream.stats.binary_file_header.trace_sorting_code = 4
stream.stats.binary_file_header.seg_y_format_revision_number = 0x0100
import sys
stream.write('../data/marmousi/synthetic_25Hz.sgy',
format='SEGY',
data_encoding=5,
byteorder=sys.byteorder)
```
<hr />
© Agile Scientific 2017
| github_jupyter |
# Introduction to Qiskit
Welcome to the Quantum Challenge! Here you will be using Qiskit, the open source quantum software development kit developed by IBM Quantum and community members around the globe. The following exercises will familiarize you with the basic elements of Qiskit and quantum circuits.
To begin, let us define what a quantum circuit is:
> **"A quantum circuit is a computational routine consisting of coherent quantum operations on quantum data, such as qubits. It is an ordered sequence of quantum gates, measurements, and resets, which may be conditioned on real-time classical computation."** (https://qiskit.org/textbook/ch-algorithms/defining-quantum-circuits.html)
While this might be clear to a quantum physicist, don't worry if it is not self-explanatory to you. During this exercise you will learn what a qubit is, how to apply quantum gates to it, and how to measure its final state. You will then be able to create your own quantum circuits! By the end, you should be able to explain the fundamentals of quantum circuits to your colleagues.
Before starting with the exercises, please run cell *Cell 1* below by clicking on it and pressing 'shift' + 'enter'. This is the general way to execute a code cell in the Jupyter notebook environment that you are using now. While it is running, you will see `In [*]:` in the top left of that cell. Once it finishes running, you will see a number instead of the star, which indicates how many cells you've run. You can find more information about Jupyter notebooks here: https://qiskit.org/textbook/ch-prerequisites/python-and-jupyter-notebooks.html.
---
For useful tips to complete this exercise as well as pointers for communicating with other participants and asking questions, please take a look at the following [repository](https://github.com/qiskit-community/may4_challenge_exercises). You will also find a copy of these exercises, so feel free to edit and experiment with these notebooks.
---
```
# Cell 1
import numpy as np
from qiskit import Aer, QuantumCircuit, execute
from qiskit.visualization import plot_histogram
from IPython.display import display, Math, Latex
from may4_challenge import plot_state_qsphere
from may4_challenge.ex1 import minicomposer
from may4_challenge.ex1 import check1, check2, check3, check4, check5, check6, check7, check8
from may4_challenge.ex1 import return_state, vec_in_braket, statevec
```
## Exercise I: Basic Operations on Qubits and Measurements
### Writing down single-qubit states
Let us start by looking at a single qubit. The main difference between a classical bit, which can take the values 0 and 1 only, is that a quantum bit, or **qubit**, can be in the states $\vert0\rangle$, $\vert1\rangle$, as well as a linear combination of these two states. This feature is known as superposition, and allows us to write the most general state of a qubit as:
$$\vert\psi\rangle = \sqrt{1-p}\vert0\rangle + e^{i \phi} \sqrt{p} \vert1\rangle$$
If we were to measure the state of this qubit, we would find the result $1$ with probability $p$, and the result $0$ with probability $1-p$. As you can see, the total probability is $1$, meaning that we will indeed measure either $0$ or $1$, and no other outcomes exists.
In addition to $p$, you might have noticed another parameter above. The variable $\phi$ indicates the relative quantum phase between the two states $\vert0\rangle$ and $\vert1\rangle$. As we will discover later, this relative phase is quite important. For now, it suffices to note that the quantum phase is what enables interference between quantum states, resulting in our ability to write quantum algorithms for solving specific tasks.
If you are interested in learning more, we refer you to [the section in the Qiskit textbook on representations of single-qubit states](https://qiskit.org/textbook/ch-states/representing-qubit-states.html).
### Visualizing quantum states
We visualize quantum states throughout this exercise using what is known as a `qsphere`. Here is how the `qsphere` looks for the states $\vert0\rangle$ and $\vert1\rangle$, respectively. Note that the top-most part of the sphere represents the state $\vert0\rangle$, while the bottom represents $\vert1\rangle$.
<img src="qsphere01.png" alt="qsphere with states 0 and 1" style="width: 400px;"/>
It should be no surprise that the superposition state with quantum phase $\phi = 0$ and probability $p = 1/2$ (meaning an equal likelihood of measuring both 0 and 1) is shown on the `qsphere` with two points. However, note also that the size of the circles at the two points is smaller than when we had simply $\vert0\rangle$ and $\vert1\rangle$ above. This is because the size of the circles is proportional to the probability of measuring each one, which is now reduced by half.
<img src="qsphereplus.png" alt="qsphere with superposition 1" style="width: 200px;"/>
In the case of superposition states, where the quantum phase is non-zero, the qsphere allows us to visualize that phase by changing the color of the respective blob. For example, the state with $\phi = 90^\circ$ (degrees) and probability $p = 1/2$ is shown in the `qsphere` below.
<img src="qspherey.png" alt="qsphere with superposition 2" style="width: 200px;"/>
### Manipulating qubits
Qubits are manipulated by applying quantum gates. Let's go through an overview of the different gates that we will consider in the following exercises.
First, let's describe how we can change the value of $p$ for our general quantum state. To do this, we will use two gates:
1. **$X$-gate**: This gate flips between the two states $\vert0\rangle$ and $\vert1\rangle$. This operation is the same as the classical NOT gate. As a result, the $X$-gate is sometimes referred to as a bit flip or NOT gate. Mathematically, the $X$ gate changes $p$ to $1-p$, so in particular from 0 to 1, and vice versa.
2. **$H$-gate**: This gate allows us to go from the state $\vert0\rangle$ to the state $\frac{1}{\sqrt{2}}\left(\vert0\rangle + \vert1\rangle\right)$. This state is also known as the $\vert+\rangle$. Mathematically, this means going from $p=0, \phi=0$ to $p=1/2, \phi=0$. As the final state of the qubit is a superposition of $\vert0\rangle$ and $\vert1\rangle$, the Hadamard gate represents a true quantum operation.
Notice that both gates changed the value of $p$, but not $\phi$. Fortunately for us, it's quite easy to visualize the action of these gates by looking at the figure below.
<img src="quantumgates.png" alt="quantum gates" style="width: 400px;"/>
Once we have the state $\vert+\rangle$, we can then change the quantum phase by applying several other gates. For example, an $S$ gate adds a phase of $90$ degrees to $\phi$, while the $Z$ gate adds a phase of $180$ degrees to $\phi$. To subtract a phase of $90$ degrees, we can apply the $S^\dagger$ gate, which is read as S-dagger, and commonly written as `sdg`. Finally, there is a $Y$ gate which applies a sequence of $Z$ and $X$ gates.
You can experiment with the gates $X$, $Y$, $Z$, $H$, $S$ and $S^\dagger$ to become accustomed to the different operations and how they affect the state of a qubit. To do so, you can run *Cell 2* which starts our circuit widget. After running the cell, choose a gate to apply to a qubit, and then choose the qubit (in the first examples, the only qubit to choose is qubit 0). Watch how the corresponding state changes with each gate, as well as the description of that state. It will also provide you with the code that creates the corresponding quantum circuit in Qiskit below the qsphere.
If you want to learn more about describing quantum states, Pauli operators, and other single-qubit gates, see chapter 1 of our textbook: https://qiskit.org/textbook/ch-states/introduction.html.
```
# Cell 2
# press shift + return to run this code cell
# then, click on the gate that you want to apply to your qubit
# next, you have to choose the qubit that you want to apply it to (choose '0' here)
# click on clear to restart
minicomposer(1, dirac=True, qsphere=True)
```
Here are four small exercises to attain different states on the qsphere. You can either solve them with the widget above and copy paste the code it provides into the respective cells to create the quantum circuits, or you can directly insert a combination of the following code lines into the program to apply the different gates:
qc.x(0) # bit flip
qc.y(0) # bit and phase flip
qc.z(0) # phase flip
qc.h(0) # superpostion
qc.s(0) # quantum phase rotation by pi/2 (90 degrees)
qc.sdg(0) # quantum phase rotation by -pi/2 (90 degrees)
The '(0)' indicates that we apply this gate to qubit 'q0', which is the first (and in this case only) qubit.
Try to attain the given state on the qsphere in each of the following exercises.
### I.i) Let us start by performing a bit flip. The goal is to reach the state $\vert1\rangle$ starting from state $\vert0\rangle$. <img src="state1.png" width="300">
If you have reached the desired state with the widget, copy and paste the code from *Cell 2* into *Cell 3* (where it says "FILL YOUR CODE IN HERE") and run it to check your solution.
```
# Cell 3
def create_circuit():
qc = QuantumCircuit(1)
qc.x(0)
return qc
# check solution
qc = create_circuit()
state = statevec(qc)
check1(state)
plot_state_qsphere(state.data, show_state_labels=True, show_state_angles=True)
```
### I.ii) Next, let's create a superposition. The goal is to reach the state $|+\rangle = \frac{1}{\sqrt{2}}\left(|0\rangle + |1\rangle\right)$. <img src="stateplus.png" width="300">
Fill in the code in the lines indicated in *Cell 4*. If you prefer the widget, you can still copy the code that the widget gives in *Cell 2* and paste it into *Cell 4*.
```
# Cell 4
def create_circuit2():
qc = QuantumCircuit(1)
qc.h(0)
return qc
qc = create_circuit2()
state = statevec(qc)
check2(state)
plot_state_qsphere(state.data, show_state_labels=True, show_state_angles=True)
```
### I.iii) Let's combine those two. The goal is to reach the state $|-\rangle = \frac{1}{\sqrt{2}}\left(|0\rangle - |1\rangle\right)$. <img src="stateminus.png" width="300">
Can you combine the above two tasks to come up with the solution?
```
# Cell 5
def create_circuit3():
qc = QuantumCircuit(1)
qc.h(0)
qc.y(0)
return qc
qc = create_circuit3()
state = statevec(qc)
check3(state)
plot_state_qsphere(state.data, show_state_labels=True, show_state_angles=True)
```
### I.iv) Finally, we move on to the complex numbers. The goal is to reach the state $|\circlearrowleft\rangle = \frac{1}{\sqrt{2}}\left(|0\rangle - i|1\rangle\right)$. <img src="stateleft.png" width="300">
```
# Cell 6
def create_circuit4():
qc = QuantumCircuit(1)
qc.h(0)
qc.y(0)
qc.s(0)
return qc
qc = create_circuit4()
state = statevec(qc)
check4(state)
plot_state_qsphere(state.data, show_state_labels=True, show_state_angles=True)
```
## Exercise II: Quantum Circuits Using Multi-Qubit Gates
Great job! Now that you've understood the single-qubit gates, let us look at gates operating on multiple qubits. The basic gates on two qubits are given by
qc.cx(c,t) # controlled-X (= CNOT) gate with control qubit c and target qubit t
qc.cz(c,t) # controlled-Z gate with control qubit c and target qubit t
qc.swap(a,b) # SWAP gate that swaps the states of qubit a and qubit b
If you'd like to read more about the different multi-qubit gates and their relations, visit chapter 2 of our textbook: https://qiskit.org/textbook/ch-gates/introduction.html.
As before, you can use the two-qubit circuit widget below to see how the combined two qubit state evolves when applying different gates (run *Cell 7*) and get the corresponding code that you can copy and paste into the program. Note that for two qubits a general state is of the form $a|00\rangle + b |01\rangle + c |10\rangle + d|11\rangle$, where $a$, $b$, $c$, and $d$ are complex numbers whose absolute values squared give the probability to measure the respective state; e.g., $|a|^2$ would be the probability to end in state '0' on both qubits. This means we can now have up to four points on the qsphere.
```
# Cell 7
# press shift + return to run this code cell
# then, click on the gate that you want to apply followed by the qubit(s) that you want it to apply to
# for controlled gates, the first qubit you choose is the control qubit and the second one the target qubit
# click on clear to restart
minicomposer(2, dirac = True, qsphere = True)
```
We start with the canonical two qubit gate, the controlled-NOT (also CNOT or CX) gate. Here, as with all controlled two qubit gates, one qubit is labelled as the "control", while the other is called the "target". If the control qubit is in state $|0\rangle$, it applies the identity $I$ gate to the target, i.e., no operation is performed. Instead, if the control qubit is in state $|1\rangle$, an X-gate is performed on the target qubit. Therefore, with both qubits in one of the two classical states, $|0\rangle$ or $|1\rangle$, the CNOT gate is limited to classical operations.
This situation changes dramatically when we first apply a Hadamard gate to the control qubit, bringing it into the superposition state $|+\rangle$. The action of a CNOT gate on this non-classical input can produce highly entangled states between control and target qubits. If the target qubit is initially in the $|0\rangle$ state, the resulting state is denoted by $|\Phi^+\rangle$, and is one of the so-called Bell states.
### II.i) Construct the Bell state $|\Phi^+\rangle = \frac{1}{\sqrt{2}}\left(|00\rangle + |11\rangle\right)$. <img src="phi+.png" width="300">
For this state we would have probability $\frac{1}{2}$ to measure "00" and probability $\frac{1}{2}$ to measure "11". Thus, the outcomes of both qubits are perfectly correlated.
```
# Cell 8
def create_circuit():
qc = QuantumCircuit(2)
qc.h(0)
qc.cx(0, 1)
return qc
qc = create_circuit()
state = statevec(qc) # determine final state after running the circuit
display(Math(vec_in_braket(state.data)))
check5(state)
qc.draw(output='mpl') # we draw the circuit
```
Next, try to create the state of perfectly anti-correlated qubits. Note the minus sign here, which indicates the relative phase between the two states.
### II.ii) Construct the Bell state $\vert\Psi^-\rangle = \frac{1}{\sqrt{2}}\left(\vert01\rangle - \vert10\rangle\right)$. <img src="psi-.png" width="300">
```
# Cell 9
def create_circuit6():
qc = QuantumCircuit(2,2) # this time, we not only want two qubits, but also
# two classical bits for the measurement later
qc.h(0)
qc.cx(0, 1)
qc.x(0)
qc.z(1)
return qc
qc = create_circuit6()
state = statevec(qc) # determine final state after running the circuit
display(Math(vec_in_braket(state.data)))
check6(state)
qc.measure(0, 0) # we perform a measurement on qubit q_0 and store the information on the classical bit c_0
qc.measure(1, 1) # we perform a measurement on qubit q_1 and store the information on the classical bit c_1
qc.draw(output='mpl') # we draw the circuit
```
As you can tell from the circuit (and the code) we have added measurement operators to the circuit. Note that in order to store the measurement results, we also need two classical bits, which we have added when creating the quantum circuit: `qc = QuantumCircuit(num_qubits, num_classicalbits)`.
In *Cell 10* we have defined a function `run_circuit()` that will run a circuit on the simulator. If the right state is prepared, we have probability $\frac{1}{2}$ to measure each of the two outcomes, "01" and "10". However, performing the measurement with 1000 shots does not imply that we will measure exactly 500 times "01" and 500 times "10". Just like flipping a coin multiple times, it is unlikely that one will get exactly a 50/50 split between the two possible output values. Instead, there are fluctuations about this ideal distribution. You can call `run_circuit` multiple times to see the variance in the ouput.
```
# Cell 10
def run_circuit(qc):
backend = Aer.get_backend('qasm_simulator') # we choose the simulator as our backend
result = execute(qc, backend, shots = 1000).result() # we run the simulation
counts = result.get_counts() # we get the counts
return counts
counts = run_circuit(qc)
print(counts)
plot_histogram(counts) # let us plot a histogram to see the possible outcomes and corresponding probabilities
```
### II.iii) You are given the quantum circuit described in the function below. Swap the states of the first and the second qubit.
This should be your final state: <img src="stateIIiii.png" width="300">
```
# Cell 11
def create_circuit7():
qc = QuantumCircuit(2)
qc.rx(np.pi/3,0)
qc.x(1)
return qc
qc = create_circuit7()
qc.swap(1, 0)
state = statevec(qc) # determine final state after running the circuit
display(Math(vec_in_braket(state.data)))
check7(state)
plot_state_qsphere(state.data, show_state_labels=True, show_state_angles=True)
```
### II.iv) Write a program from scratch that creates the GHZ state (on three qubits), $\vert \text{GHZ}\rangle = \frac{1}{\sqrt{2}} \left(|000\rangle + |111 \rangle \right)$, performs a measurement with 2000 shots, and returns the counts. <img src="ghz.png" width="300">
If you want to track the state as it is evolving, you could use the circuit widget from above for three qubits, i.e., `minicomposer(3, dirac=True, qsphere=True)`. For how to get the counts of a measurement, look at the code in *Cell 9* and *Cell 10*.
```
# Cell 12
def create_circuit8():
qc = QuantumCircuit(3,3) # this time, we not only want two qubits, but also
# two classical bits for the measurement later
qc.h(0)
qc.cx(0, 1)
qc.cx(1, 2)
qc.measure([0, 1, 2], [0, 1, 2])
return qc
qc = create_circuit8()
qc.draw('mpl')
backend = Aer.get_backend('qasm_simulator') # we choose the simulator as our backend
result = execute(qc, backend, shots = 2000).result() # we run the simulation
counts = result.get_counts() # we get the counts
print(counts)
check8(counts)
plot_histogram(counts)
```
Congratulations for finishing this introduction to Qiskit! Once you've reached all 8 points, the solution string will be displayed. You need to copy and paste that string on the IBM Quantum Challenge page to complete the exercise and track your progress.
Now that you have created and run your first quantum circuits, you are ready for the next exercise, where we will make use of the actual hardware and learn how to reduce the noise in the outputs.
| github_jupyter |
# Example dot tuning
```
from qcodes import Station
import nanotune as nt
from nanotune.tuningstages.settings import DataSettings, SetpointSettings, Classifiers
from nanotune.tests.functional_tests.sim_tuner import SimDotTuner
from nanotune.drivers.mock_dac import MockDAC, MockDACChannel
from nanotune.tests.mock_classifier import MockClassifer
from sim.qcodes_mocks import MockDoubleQuantumDotInstrument
from load_sim_scenarios import sim_scenario_dottuning
from load_sim_device import load_sim_device
```
# Load instruments and simulator to play back data
We use a MockDAC and a simulator to replace a real setup.
The simulator takes previously measured data and plays it back. It needs to be initialized and added to the station.
```
station = Station()
dac = MockDAC('dac', MockDACChannel)
station.add_component(dac)
qd_mock_instrument = MockDoubleQuantumDotInstrument('qd_mock_instrument')
station.add_component(qd_mock_instrument, name="qd_mock_instrument")
```
Now we load the device and overwrite the channel's voltage parameters with the simulator's pin voltages.
Then the simulation scenario is loaded, i.e. we tell the simulator which data to play back and update the device's normalization constants. In a real measurement, the normalization constants need to be measured. Unfortunately this is currently not supported by the simulator.
```
device = load_sim_device(station, "./chip.yaml")
sim_sceanrio = sim_scenario_dottuning(station)
device.normalization_constants
```
# Initialize tuner
We use the SimDotTuner here, which is a subclass of the DotTuner with the only difference that before each measurement, the data simulated is advanced to replay the next dataset.
```
settings = {
"name": "test_tuner",
"data_settings": DataSettings(
db_name="dot_tuning_example.db",
db_folder='.',
segment_db_name="dot_segments_temp.db",
segment_db_folder='.',
),
"classifiers": Classifiers(
pinchoff=MockClassifer(category="pinchoff"),
singledot=MockClassifer(category="singledot"),
doubledot=MockClassifer(category="doubledot"),
dotregime=MockClassifer(category="dotregime")),
"setpoint_settings": SetpointSettings(
voltage_precision=0.001,
ranges_to_sweep=[(-1, 0)],
safety_voltage_ranges=[(-2, 0)],
),
}
dottuner = SimDotTuner(
**settings,
sim_scenario=sim_sceanrio,
)
dottuner.tune_dot_regime(device)
```
The tuning result is saved in the tuner's tuning history.
```
dottuner.tuning_history.results[device.name].tuningresults.keys()
```
The valid ranges found during tuning can be accessed via the current valid ranges attribute.
```
device.current_valid_ranges()
```
| github_jupyter |
<h1>Table of Contents<span class="tocSkip"></span></h1>
<div class="toc"><ul class="toc-item"><li><span><a href="#Intro-and-setup" data-toc-modified-id="Intro-and-setup-1"><span class="toc-item-num">1 </span>Intro and setup</a></span></li><li><span><a href="#Implementing-a-Neural-Network" data-toc-modified-id="Implementing-a-Neural-Network-2"><span class="toc-item-num">2 </span>Implementing a Neural Network</a></span></li><li><span><a href="#Training-the-network" data-toc-modified-id="Training-the-network-3"><span class="toc-item-num">3 </span>Training the network</a></span><ul class="toc-item"><li><span><a href="#Training-with-ReLu-activation" data-toc-modified-id="Training-with-ReLu-activation-3.1"><span class="toc-item-num">3.1 </span>Training with ReLu activation</a></span></li><li><span><a href="#Training-with-sigmoid-activation" data-toc-modified-id="Training-with-sigmoid-activation-3.2"><span class="toc-item-num">3.2 </span>Training with sigmoid activation</a></span></li><li><span><a href="#Training-with-leaky-ReLu" data-toc-modified-id="Training-with-leaky-ReLu-3.3"><span class="toc-item-num">3.3 </span>Training with leaky ReLu</a></span></li><li><span><a href="#Training-with-tanh" data-toc-modified-id="Training-with-tanh-3.4"><span class="toc-item-num">3.4 </span>Training with <code>tanh</code></a></span></li></ul></li><li><span><a href="#Mini-batches" data-toc-modified-id="Mini-batches-4"><span class="toc-item-num">4 </span>Mini batches</a></span></li><li><span><a href="#Evaluating-models" data-toc-modified-id="Evaluating-models-5"><span class="toc-item-num">5 </span>Evaluating models</a></span></li><li><span><a href="#PyTorch-simple-implementation" data-toc-modified-id="PyTorch-simple-implementation-6"><span class="toc-item-num">6 </span>PyTorch simple implementation</a></span></li><li><span><a href="#Using-my-own-image" data-toc-modified-id="Using-my-own-image-7"><span class="toc-item-num">7 </span>Using my own image</a></span></li></ul></div>
# Building a Neural Network from Scratch
## Intro and setup
The purpose of this notebook is simple. I want to build a simple Artificial Neural Network without using any framework such as `scikit-learn`, `PyTorch` or `TensorFlow`.
I believe this will help me gain a deeper understanding of the mathematics involved and will help that understanding sediment in my memory better.
First of all let's import some libraries. Since we aren't going to use any deep learning or machine learning frameworks, the number of imports will be limited.
```
import numpy as np
import matplotlib.pyplot as plt
from typing import List
import pickle # To store data and trained models
from tqdm import tqdm # To visualize progress bars during training
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
```
We will be training and testing our network using the [MNIST_784 dataset](https://www.openml.org/d/554). We can download that as a `python` dictionary using `scikit-learn`.
```
from sklearn.datasets import fetch_openml
mnist = fetch_openml(name="mnist_784")
with open("mnist.pickle", 'wb') as f:
pickle.dump(mnist, f)
with open("mnist.pickle", 'rb') as f:
mnist = pickle.load(f)
data = mnist.data
labels = mnist.target
mnist.keys()
# Let's select a random datapoint.
# np.random.seed(42)
n = np.random.choice(np.arange(data.shape[0]+1))
print(n)
test_img = data.iloc[n].values
test_label = mnist.target.iloc[n]
print(test_img.shape)
side_length = int(np.sqrt(test_img.shape))
reshaped_test_img = test_img.reshape(side_length, side_length)
print("Image label: " + str(test_label))
plt.imshow(reshaped_test_img, cmap="Greys")
plt.axis('off')
plt.show()
```
So we know that image is a vector of shape 784, which is a flattened 28 x 28 2D matrix.
If we split our dataset into a training set with e.g. 60.000 datapoints and a test set with 10.000 datapoints we will end up with the following: $input\;layer\;size = m*n$, where $m$ is the number of samples and $n$ the number of features. Hence in our case the network's input layer with have size (60.000 x 784).<br>Let's start by coding a simple NN with just one hidden layer.
If the hidden layer has say 4 nodes: how do we control the mapping from the input layer to the hidden layer? In other words, what should be the size of the **weight matrix**?
Each node in the input layer will connect to each input in the hidden layer. So we will need 784 * 4 = 3136 weights. And each node in the hidden layer will add a bias to each weighted sum so we'll need 4 biases.
As a rule: if layer $L_n$ has $s_n$ nodes and layer $L_{n+1}$ has $s_{n+1
}$ nodes then the $\Theta_n$ matrix mapping fron $L_n$ to $L_{n+1}$ will be of size $(s_{n+1},\;s_{n})$.
```
w1 = np.ones((4, 784)) * 0.01
z1 = np.dot(w1, data.T)
print(z1.shape)
w2 = np.ones((10, 4))
z2 = np.dot(w2, z1)
print(z2.shape)
```
(But remember: we need to add *biases* to each node in the hidden layer.)
## Implementing a Neural Network
Let's start by defining some **activation functions**:
```
def sigmoid(z: np.ndarray) -> np.ndarray:
return 1.0 / (1.0 + np.exp(-z))
def relu(z: np.ndarray) -> np.ndarray:
return np.maximum(0, z)
def tanh(z: np.ndarray) -> np.ndarray:
return np.tanh(z)
def leaky_relu(z: np.ndarray) -> np.ndarray:
return np.where(z > 0, z, z * 0.01)
```
For the final layer we will use **softmax** given that this is a multiclass classification problem.
```
def softmax(z: np.ndarray) -> np.ndarray:
e = np.exp(z - np.max(z))
return e / np.sum(e, axis=0, keepdims=True)
```
We are also going to need some pre-processing functions:
```
def normalize(x: np.ndarray) -> np.ndarray:
return (x - np.min(x)) / (np.max(x) - np.min(x))
def one_hot_encode(x: np.ndarray, num_labels: int) -> np.ndarray:
return np.eye(num_labels)[x]
```
Finally, we will need derivatives for the activation functions to perform gradient descent.
```
def derivative(function_name: str, z: np.ndarray) -> np.ndarray:
if function_name == "sigmoid":
return sigmoid(z) * (1 - sigmoid(z))
if function_name == "tanh":
return 1 - np.square(tanh(z))
if function_name == "relu":
y = (z > 0) * 1
return y
if function_name == "leaky_relu":
return np.where(z > 0, 1, 0.01)
return "No such activation"
```
We now have everything we need for our deep neural network.
```
class NN(object):
def __init__(self, X: np.ndarray, y: np.ndarray, X_test: np.ndarray, y_test: np.ndarray, activation: str, num_labels: int, architecture: List[int]):
self.X = normalize(X) # normalize training data in range 0,1
assert np.all((self.X >= 0) | (self.X <= 1)) # test that normalize succeded
self.X, self.X_test = X.copy(), X_test.copy()
self.y, self.y_test = y.copy(), y_test.copy()
self.layers = {} # define dict to store results of activation
self.architecture = architecture # size of hidden layers as array
self.activation = activation # activation function
assert self.activation in ["relu", "tanh", "sigmoid", "leaky_relu"]
self.parameters = {}
self.num_labels = num_labels
self.m = X.shape[1]
self.architecture.append(self.num_labels)
self.num_input_features = X.shape[0]
self.architecture.insert(0, self.num_input_features)
self.L = len(architecture)
assert self.X.shape == (self.num_input_features, self.m)
assert self.y.shape == (self.num_labels, self.m)
def initialize_parameters(self):
for i in range(1, self.L):
print(f"Initializing parameters for layer: {i}.")
self.parameters["w"+str(i)] = np.random.randn(self.architecture[i], self.architecture[i-1]) * 0.01
self.parameters["b"+str(i)] = np.zeros((self.architecture[i], 1))
def forward(self):
params=self.parameters
self.layers["a0"] = self.X
for l in range(1, self.L-1):
self.layers["z" + str(l)] = np.dot(params["w" + str(l)],
self.layers["a"+str(l-1)]) + params["b"+str(l)]
self.layers["a" + str(l)] = eval(self.activation)(self.layers["z"+str(l)])
assert self.layers["a"+str(l)].shape == (self.architecture[l], self.m)
self.layers["z" + str(self.L-1)] = np.dot(params["w" + str(self.L-1)],
self.layers["a"+str(self.L-2)]) + params["b"+str(self.L-1)]
self.layers["a"+str(self.L-1)] = softmax(self.layers["z"+str(self.L-1)])
self.output = self.layers["a"+str(self.L-1)]
assert self.output.shape == (self.num_labels, self.m)
assert all([s for s in np.sum(self.output, axis=1)])
cost = - np.sum(self.y * np.log(self.output + 0.000000001))
return cost, self.layers
def backpropagate(self):
derivatives = {}
dZ = self.output - self.y
assert dZ.shape == (self.num_labels, self.m)
dW = np.dot(dZ, self.layers["a" + str(self.L-2)].T) / self.m
db = np.sum(dZ, axis=1, keepdims=True) / self.m
dAPrev = np.dot(self.parameters["w" + str(self.L-1)].T, dZ)
derivatives["dW" + str(self.L-1)] = dW
derivatives["db" + str(self.L-1)] = db
for l in range(self.L-2, 0, -1):
dZ = dAPrev * derivative(self.activation, self.layers["z" + str(l)])
dW = 1. / self.m * np.dot(dZ, self.layers["a" + str(l-1)].T)
db = 1. / self.m * np.sum(dZ, axis=1, keepdims=True)
if l > 1:
dAPrev = np.dot(self.parameters["w" + str(l)].T, (dZ))
derivatives["dW" + str(l)] = dW
derivatives["db" + str(l)] = db
self.derivatives = derivatives
return self.derivatives
def fit(self, lr=0.01, epochs=1000):
self.costs = []
self.initialize_parameters()
self.accuracies = {"train": [], "test": []}
for epoch in tqdm(range(epochs), colour="BLUE"):
cost, cache = self.forward()
self.costs.append(cost)
derivatives = self.backpropagate()
for layer in range(1, self.L):
self.parameters["w"+str(layer)] = self.parameters["w"+str(layer)] - lr * derivatives["dW" + str(layer)]
self.parameters["b"+str(layer)] = self.parameters["b"+str(layer)] - lr * derivatives["db" + str(layer)]
train_accuracy = self.accuracy(self.X, self.y)
test_accuracy = self.accuracy(self.X_test, self.y_test)
if epoch % 10 == 0:
print(f"Epoch: {epoch:3d} | Cost: {cost:.3f} | Accuracy: {train_accuracy:.3f}")
self.accuracies["train"].append(train_accuracy)
self.accuracies["test"].append(test_accuracy)
print("Training terminated")
def predict(self, x):
params = self.parameters
n_layers = self.L - 1
values = [x]
for l in range(1, n_layers):
z = np.dot(params["w" + str(l)], values[l-1]) + params["b" + str(l)]
a = eval(self.activation)(z)
values.append(a)
z = np.dot(params["w"+str(n_layers)], values[n_layers-1]) + params["b"+str(n_layers)]
a = softmax(z)
if x.shape[1]>1:
ans = np.argmax(a, axis=0)
else:
ans = np.argmax(a)
return ans
def accuracy(self, X, y):
P = self.predict(X)
return sum(np.equal(P, np.argmax(y, axis=0))) / y.shape[1]*100
def pickle_model(self, name: str):
with open("fitted_model_"+ name + ".pickle", "wb") as modelFile:
pickle.dump(self, modelFile)
def plot_counts(self):
counts = np.unique(np.argmax(self.output, axis=0), return_counts=True)
plt.bar(counts[0], counts[1], color="navy")
plt.ylabel("Counts")
plt.xlabel("y_hat")
plt.title("Distribution of predictions")
plt.show()
def plot_cost(self, lr):
plt.figure(figsize=(8, 4))
plt.plot(np.arange(0, len(self.costs)), self.costs, lw=1, color="orange")
plt.title(f"Learning rate: {lr}\nFinal Cost: {self.costs[-1]:.5f}", fontdict={
"family":"sans-serif",
"size": "12"})
plt.xlabel("Epoch")
plt.ylabel("Cost")
plt.show()
def plot_accuracies(self, lr):
acc = self.accuracies
fig = plt.figure(figsize=(6,4))
ax = fig.add_subplot(111)
ax.plot(acc["train"], label="train")
ax.plot(acc["test"], label="test")
plt.legend(loc="lower right")
ax.set_title("Accuracy")
ax.annotate(f"Train: {acc['train'][-1]:.2f}", (len(acc["train"])+4, acc["train"][-1]+2), color="blue")
ax.annotate(f"Test: {acc['test'][-1]:.2f}", (len(acc["test"])+4, acc["test"][-1]-2), color="orange")
ax.spines["right"].set_visible(False)
ax.spines["top"].set_visible(False)
plt.show()
def __str__(self):
return str(self.architecture)
```
## Training the network
Let's split the data into a training and a test set.
```
# reloading data
with open("mnist.pickle", 'rb') as f:
mnist = pickle.load(f)
data = mnist.data
labels = mnist.target
train_test_split_no = 60000
X_train = data.values[:train_test_split_no].T
y_train = labels[:train_test_split_no].values.astype(int)
y_train = one_hot_encode(y_train, 10).T
X_test = data.values[train_test_split_no:].T
y_test = labels[train_test_split_no:].values.astype(int)
y_test = one_hot_encode(y_test, 10).T
X_train.shape, X_test.shape
```
### Training with ReLu activation
```
PARAMS = [X_train, y_train, X_test, y_test, "relu", 10, [512, 300]]
nn_relu = NN(*PARAMS)
epochs_relu = 50
lr_relu = 0.003
nn_relu.fit(lr=lr_relu, epochs=epochs_relu)
nn_relu.plot_cost(lr_relu)
nn_relu.plot_accuracies(lr_relu)
nn_relu.pickle_model("relu")
with open("fitted_model_relu.pickle", "rb") as f:
nn_relu = pickle.load(f)
```
### Training with sigmoid activation
```
PARAMS_sigmoid = [X_train, y_train, X_test, y_test, "sigmoid", 10, [512, 300]]
lr_sigmoid = 0.01
epochs_sigmoid = 150
nn_sigmoid = NN(*PARAMS_sigmoid)
nn_sigmoid.fit(lr=lr_sigmoid, epochs=epochs_sigmoid)
nn_sigmoid.pickle_model("sigmoid")
with open("fitted_model_sigmoid.pickle", "rb") as f:
nn_sigmoid = pickle.load(f)
nn_sigmoid.plot_cost(lr_sigmoid)
nn_sigmoid.plot_accuracies(lr_sigmoid)
```
### Training with leaky ReLu
```
PARAMS_leaky = [X_train, y_train, X_test, y_test, "leaky_relu", 10, [128, 32]]
nn_leaky = NN(*PARAMS_leaky)
lr_leaky = 0.003
epochs_leaky = 200
nn_leaky.fit(X_train, y_train, lr=lr_leaky, epochs=epochs_leaky)
nn_leaky.plot_cost(lr=lr_leaky)
nn_leaky.plot_accuracies(lr_leaky)
nn_leaky.pickle_model("leaky")
with open("fitted_model_leaky.pickle", "rb") as f:
nn_leaky = pickle.load(f)
```
### Training with `tanh`
```
PARAMS_tanh = [X_train, y_train, X_test, y_test, "relu", 10, [64, 10]]
nn_tanh = NN(*PARAMS_tanh)
nn_tanh.fit(X_train, y_train, lr=0.003, epochs=100)
nn_tanh.plot_cost(lr=0.003)
nn_tanh.plot_accuracies(lr=0.003)
nn_tanh.pickle_model("tanh")
with open("fitted_model_tanh.pickle", "rb") as f:
nn_tanh = pickle.load(f)
```
## Mini batches
```
# Not yet implemented
def iterate_mini_batches(X, y, batch_size, shuffle=False):
no_of_samples = X.shape[1]
if shuffle:
ind_list = np.array([i for i in range(no_of_samples)])
np.random.shuffle(ind_list)
X = X[:, ind_list]
y = y[:, ind_list]
for start_idx in range(0, no_of_samples, batch_size):
if start_idx + batch_size <= no_of_samples:
end_idx = start_idx+batch_size -1
else:
end_idx = (no_of_samples) % batch_size + start_idx + 1
print(start_idx, end_idx)
yield X[:, start_idx:end_idx], y[:, start_idx:end_idx]
```
## Evaluating models
```
from tabulate import tabulate
def print_accuracies(models, X_train, y_train, X_test, y_test):
data = [[model.activation, model.architecture[1:-1], model.accuracy(X_train, y_train), model.accuracy(X_test, y_test)] for model in models]
print(tabulate(data, headers=["Activation", "Architecture","Train", "Test"]))
print_accuracies([nn_relu, nn_sigmoid, nn_leaky], X_train, y_train, X_test, y_test)
# Results for nn_relu
example_index = 2132
plt.imshow(X_test[:, example_index].reshape(28,28), cmap="Greys")
prediction = nn_relu.predict(X_test[:, example_index].reshape((X_test.shape[0], 1)))
plt.title("Prediction: " + str(prediction))
plt.axis('off')
plt.show()
X_test[:, example_index].shape
def plot_predictions(model, X_test, labels, rows, cols):
indexes = np.random.choice(range(X_test.shape[1]), rows * cols)
fig, ax = plt.subplots(rows, cols, figsize=(5, 4))
n = 0
for i in range(rows):
for j in range(cols):
ix = indexes[n]
example = X_test[:, ix].reshape(28,28)
prediction = model.predict(X_test[:, ix].reshape((X_test[:, ix].shape[0], 1)))
subplot = ax[i, j]
subplot.set_title(f"Prediction: {prediction}\nLabel: {np.argmax(labels[:, ix])}")
subplot.imshow(example, cmap="Greys")
subplot.axis('off')
n += 1
plt.tight_layout()
plt.show()
plot_predictions(nn_relu, X_test, y_test, 3, 3)
nn_relu.plot_counts()
```
## PyTorch simple implementation
```
import torch
from torch import nn
from torch.utils.data import DataLoader
from torchvision import datasets
from torchvision.transforms import ToTensor, Lambda, Compose
training_data = datasets.MNIST(
root="data",
train=True,
download=True,
transform=ToTensor(),
)
test_data = datasets.MNIST(
root="data",
train=False,
download=True,
transform=ToTensor(),
)
batch_size = 64
# Create data loaders.
train_dataloader = DataLoader(training_data, batch_size=batch_size)
test_dataloader = DataLoader(test_data, batch_size=batch_size)
for X, y in test_dataloader:
print("Shape of X [N, C, H, W]: ", X.shape)
print("Shape of y: ", y.shape, y.dtype)
break
# Get cpu or gpu device for training.
device = "cuda" if torch.cuda.is_available() else "cpu"
print("Using {} device".format(device))
# Define model
class NeuralNetwork(nn.Module):
def __init__(self):
super(NeuralNetwork, self).__init__()
self.flatten = nn.Flatten()
self.linear_relu_stack = nn.Sequential(
nn.Linear(28*28, 512),
nn.ReLU(),
nn.Linear(512, 512),
nn.ReLU(),
nn.Linear(512, 10)
)
def forward(self, x):
x = self.flatten(x)
logits = self.linear_relu_stack(x)
return logits
model = NeuralNetwork().to(device)
print(model)
loss_fn = nn.CrossEntropyLoss()
optimizer = torch.optim.SGD(model.parameters(), lr=1e-3)
def train(dataloader, model, loss_fn, optimizer):
size = len(dataloader.dataset)
model.train()
for batch, (X, y) in enumerate(dataloader):
X, y = X.to(device), y.to(device)
# Compute prediction error
pred = model(X)
loss = loss_fn(pred, y)
# Backpropagation
optimizer.zero_grad()
loss.backward()
optimizer.step()
if batch % 100 == 0:
loss, current = loss.item(), batch * len(X)
print(f"loss: {loss:>7f} [{current:>5d}/{size:>5d}]")
def test(dataloader, model, loss_fn):
size = len(dataloader.dataset)
num_batches = len(dataloader)
model.eval()
test_loss, correct = 0, 0
with torch.no_grad():
for X, y in dataloader:
X, y = X.to(device), y.to(device)
pred = model(X)
test_loss += loss_fn(pred, y).item()
correct += (pred.argmax(1) == y).type(torch.float).sum().item()
test_loss /= num_batches
correct /= size
print(f"Test Error: \n Accuracy: {(100*correct):>0.1f}%, Avg loss: {test_loss:>8f} \n")
epochs = 10
for t in range(epochs):
print(f"Epoch {t+1}\n-------------------------------")
train(train_dataloader, model, loss_fn, optimizer)
test(test_dataloader, model, loss_fn)
print("Done!")
import cv2
def open_as_mnist(image_path):
"""
Assume this is a color or grey scale image of a digit which has not so far been preprocessed
Black and White
Resize to 20 x 20 (digit in center ideally)
Sharpen
Add white border to make it 28 x 28
Convert to white on black
"""
# open as greyscale
image = cv2.imread(image_path, 0)
# crop to contour with largest area
cropped = do_cropping(image)
# resizing the image to 20 x 20
resized20 = cv2.resize(cropped, (20, 20), interpolation=cv2.INTER_CUBIC)
cv2.imwrite('1_resized.jpg', resized20)
# gaussian filtering
blurred = cv2.GaussianBlur(resized20, (3, 3), 0)
# white digit on black background
ret, thresh = cv2.threshold(blurred, 127, 255, cv2.THRESH_BINARY_INV)
padded = to20by20(thresh)
resized28 = padded_image(padded, 28)
# normalize the image values to fit in the range [0,1]
norm_image = np.asarray(resized28, dtype=np.float32) / 255.
# cv2.imshow('image', norm_image)
# cv2.waitKey(0)
# # Flatten the image to a 1-D vector and return
flat = norm_image.reshape(1, 28 * 28)
# return flat
# normalize pixels to 0 and 1. 0 is pure white, 1 is pure black.
tva = [(255 - x) * 1.0 / 255.0 for x in flat]
return tva
def padded_image(image, tosize):
"""
This method adds padding to the image and makes it to a tosize x tosize array,
without losing the aspect ratio.
Assumes desired image is square
:param image: the input image as numpy array
:param tosize: the final dimensions
"""
# image dimensions
image_height, image_width = image.shape
# if not already square then pad to square
if image_height != image_width:
# Add padding
# The aim is to make an image of different width and height to a sqaure image
# For that first the biggest attribute among width and height are determined.
max_index = np.argmax([image_height, image_width])
# if height is the biggest one, then add padding to width until width becomes
# equal to height
if max_index == 0:
#order of padding is: top, bottom, left, right
left = int((image_height - image_width) / 2)
right = image_height - image_width - left
padded_img = cv2.copyMakeBorder(image, 0, 0,
left,
right,
cv2.BORDER_CONSTANT)
# else if width is the biggest one, then add padding to height until height becomes
# equal to width
else:
top = int((image_width - image_height) / 2)
bottom = image_width - image_height - top
padded_img = cv2.copyMakeBorder(image, top, bottom, 0, 0, cv2.BORDER_CONSTANT)
else:
padded_img = image
# now that it's a square, add any additional padding required
image_height, image_width = padded_img.shape
padding = tosize - image_height
# need to handle where padding is not divisiable by 2
left = top = int(padding/2)
right = bottom = padding - left
resized = cv2.copyMakeBorder(image, top, bottom, left, right, cv2.BORDER_CONSTANT)
return resized
nn_relu
```
## Using my own image
```
import cv2
img = cv2.imread("for_mnist2.jpeg", cv2.IMREAD_GRAYSCALE) # flag 0 imports to grayscale
plt.imshow(img, cmap="Greys")
reshaped = np.asarray(img).reshape((X_test.shape[0], 1)) * 1.0
img
reshaped
nn_relu.predict(reshaped)
y_test[0].shape
```
| github_jupyter |
# Running real-life data periodograms
-----------------------------------
### In this notebook we will use the full PDC tests panel to explore real-life observations.
The chosen example is presented here - https://arxiv.org/abs/2111.02383 :
S Mus is a spectroscopic binary Cepheid with 505 d orbital and 9.65996 d pulsational periods.
This notebook shows how to calculate the PDC periodograms for S Mus, distinguishing the two periodic signals.
### 1 Imports & funcs
```
import sys
from sparta.Auxil.PeriodicityDetector import PeriodicityDetector
from sparta.UNICOR.Spectrum import Spectrum
from sparta.UNICOR.Template import Template
from sparta.Auxil.TimeSeries import TimeSeries
from sparta.Observations import Observations
import numpy as np
import random
from scipy import interpolate
from PyAstronomy import pyasl
import matplotlib.pyplot as plt
from scipy import signal
import winsound
from copy import deepcopy
import pandas as pd
```
### 2 Using the PeriodicityDetector class to run PDC on simulated velocity times series
`Observations` class enables one to load observation data from a given folder
and place it into a TimeSeries object.
```
# One has to define how the observation files will be read and processed.
# This can be done using the load_spectrum_from_fits function in the ReadSpec.py file.
obs_data = Observations(survey="CORALIE", sample_rate=1, min_wv=5800, max_wv=6000)
for i in obs_data.observation_TimeSeries.vals:
i = i.SpecPreProccess()
plt.figure(figsize=(7, 3.5))
plt.title("SMUS observations")
plt.ylabel("Flux")
plt.xlabel("Wavelength (Angstrom)")
for s in obs_data.observation_TimeSeries.vals:
plt.plot(s.wv[0], s.sp[0], alpha=0.3)
plt.show()
obs_data.observation_TimeSeries.size
# The observation_TimeSeries instance inside obs_data now contains a list of Spectrum objects
obs_data.observation_TimeSeries.vals
# In this case, we chose to load a list of radial velocities calculated in advance.
# One can also calculate them using the UNICOR_quickstart.ipynb notebook.
columns_to_keep = ['jdb', 'vrad', 'noise']
df = pd.read_table(r"C:\Users\AbrahamBinnfeld\Downloads\all_ready\SMus_RVdata.dat", sep="\s+", usecols=columns_to_keep)
pre_calculated_vrads = df.vrad[1:61].astype(float).values
print(pre_calculated_vrads)
```
Initializing and running PeriodicityDetector
```
# Choosing frequency range and frequency resolution for the periodograms.
obs_data.initialize_periodicity_detector(freq_range=(1 / 5_000, 0.33), periodogram_grid_resolution=5_000)
# Setting the known periods, so they are calculated in the plots
obs_data.periodicity_detector.period = [41, 9.66, 505]
obs_data.observation_TimeSeries.calculated_vrad_list = pre_calculated_vrads
obs_data.periodicity_detector.run_GLS_process()
print("done run_GLS_process")
obs_data.periodicity_detector.run_PDC_process(calc_biased_flag=False, calc_unbiased_flag=True)
print("done run_PDC_process")
obs_data.periodicity_detector.run_USURPER_process(calc_biased_flag=False, calc_unbiased_flag=True)
print("done run_USURPER_process")
obs_data.periodicity_detector.run_Partial_USURPER_process(reversed_flag=True)
print("done run_Partial_USURPER_process")
obs_data.periodicity_detector.run_Partial_USURPER_process(reversed_flag=False)
print("done run_Partial_USURPER_process rev")
obs_data.periodicity_detector.periodogram_plots(velocities_flag=True)
plt.show()
# Activate noise to indicate end of run
frequency = 2500 # Set Frequency To 2500 Hertz
duration = 500 # Set Duration To 1000 ms == 1 second
winsound.Beep(frequency, duration)
```
Note that as they are designed to do, the partial USuRPER presents only the pulsation period, while the partial PDC only present the orbital period.
P-value for the periodograms can be calculated, based on the methodology presented in - https://arxiv.org/abs/1912.12150
```
obs_data.periodicity_detector.calc_pdc_periodograms_pval([0.3, 0.42], plot=True)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/DingLi23/s2search/blob/pipelining/pipelining/exp-cspf/exp-cspf_cspf_1w_ale_plotting.ipynb" target="_blank"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
### Experiment Description
> This notebook is for experiment \<exp-cspf\> and data sample \<cspf\>.
### Initialization
```
%load_ext autoreload
%autoreload 2
import numpy as np, sys, os
in_colab = 'google.colab' in sys.modules
# fetching code and data(if you are using colab
if in_colab:
!rm -rf s2search
!git clone --branch pipelining https://github.com/youyinnn/s2search.git
sys.path.insert(1, './s2search')
%cd s2search/pipelining/exp-cspf/
pic_dir = os.path.join('.', 'plot')
if not os.path.exists(pic_dir):
os.mkdir(pic_dir)
```
### Loading data
```
sys.path.insert(1, '../../')
import numpy as np, sys, os, pandas as pd
from getting_data import read_conf
from s2search_score_pdp import pdp_based_importance
sample_name = 'cspf'
f_list = [
'title', 'abstract', 'venue', 'authors',
'year',
'n_citations'
]
ale_xy = {}
ale_metric = pd.DataFrame(columns=['feature_name', 'ale_range', 'ale_importance', 'absolute mean'])
for f in f_list:
file = os.path.join('.', 'scores', f'{sample_name}_1w_ale_{f}.npz')
if os.path.exists(file):
nparr = np.load(file)
quantile = nparr['quantile']
ale_result = nparr['ale_result']
values_for_rug = nparr.get('values_for_rug')
ale_xy[f] = {
'x': quantile,
'y': ale_result,
'rug': values_for_rug,
'weird': ale_result[len(ale_result) - 1] > 20
}
if f != 'year' and f != 'n_citations':
ale_xy[f]['x'] = list(range(len(quantile)))
ale_xy[f]['numerical'] = False
else:
ale_xy[f]['xticks'] = quantile
ale_xy[f]['numerical'] = True
ale_metric.loc[len(ale_metric.index)] = [f, np.max(ale_result) - np.min(ale_result), pdp_based_importance(ale_result, f), np.mean(np.abs(ale_result))]
# print(len(ale_result))
print(ale_metric.sort_values(by=['ale_importance'], ascending=False))
print()
```
### ALE Plots
```
import matplotlib.pyplot as plt
import seaborn as sns
from matplotlib.ticker import MaxNLocator
categorical_plot_conf = [
{
'xlabel': 'Title',
'ylabel': 'ALE',
'ale_xy': ale_xy['title']
},
{
'xlabel': 'Abstract',
'ale_xy': ale_xy['abstract']
},
{
'xlabel': 'Authors',
'ale_xy': ale_xy['authors'],
# 'zoom': {
# 'inset_axes': [0.3, 0.3, 0.47, 0.47],
# 'x_limit': [89, 93],
# 'y_limit': [-1, 14],
# }
},
{
'xlabel': 'Venue',
'ale_xy': ale_xy['venue'],
# 'zoom': {
# 'inset_axes': [0.3, 0.3, 0.47, 0.47],
# 'x_limit': [89, 93],
# 'y_limit': [-1, 13],
# }
},
]
numerical_plot_conf = [
{
'xlabel': 'Year',
'ylabel': 'ALE',
'ale_xy': ale_xy['year'],
# 'zoom': {
# 'inset_axes': [0.15, 0.4, 0.4, 0.4],
# 'x_limit': [2019, 2023],
# 'y_limit': [1.9, 2.1],
# },
},
{
'xlabel': 'Citations',
'ale_xy': ale_xy['n_citations'],
# 'zoom': {
# 'inset_axes': [0.4, 0.65, 0.47, 0.3],
# 'x_limit': [-1000.0, 12000],
# 'y_limit': [-0.1, 1.2],
# },
},
]
def pdp_plot(confs, title):
fig, axes_list = plt.subplots(nrows=1, ncols=len(confs), figsize=(20, 5), dpi=100)
subplot_idx = 0
plt.suptitle(title, fontsize=20, fontweight='bold')
# plt.autoscale(False)
for conf in confs:
axes = axes if len(confs) == 1 else axes_list[subplot_idx]
sns.rugplot(conf['ale_xy']['rug'], ax=axes, height=0.02)
axes.axhline(y=0, color='k', linestyle='-', lw=0.8)
axes.plot(conf['ale_xy']['x'], conf['ale_xy']['y'])
axes.grid(alpha = 0.4)
# axes.set_ylim([-2, 20])
axes.xaxis.set_major_locator(MaxNLocator(integer=True))
axes.yaxis.set_major_locator(MaxNLocator(integer=True))
if ('ylabel' in conf):
axes.set_ylabel(conf.get('ylabel'), fontsize=20, labelpad=10)
# if ('xticks' not in conf['ale_xy'].keys()):
# xAxis.set_ticklabels([])
axes.set_xlabel(conf['xlabel'], fontsize=16, labelpad=10)
if not (conf['ale_xy']['weird']):
if (conf['ale_xy']['numerical']):
axes.set_ylim([-1.5, 1.5])
pass
else:
axes.set_ylim([-10, 15])
pass
if 'zoom' in conf:
axins = axes.inset_axes(conf['zoom']['inset_axes'])
axins.xaxis.set_major_locator(MaxNLocator(integer=True))
axins.yaxis.set_major_locator(MaxNLocator(integer=True))
axins.plot(conf['ale_xy']['x'], conf['ale_xy']['y'])
axins.set_xlim(conf['zoom']['x_limit'])
axins.set_ylim(conf['zoom']['y_limit'])
axins.grid(alpha=0.3)
rectpatch, connects = axes.indicate_inset_zoom(axins)
connects[0].set_visible(False)
connects[1].set_visible(False)
connects[2].set_visible(True)
connects[3].set_visible(True)
subplot_idx += 1
pdp_plot(categorical_plot_conf, f"ALE for {len(categorical_plot_conf)} categorical features")
# plt.savefig(os.path.join('.', 'plot', f'{sample_name}-1wale-categorical.png'), facecolor='white', transparent=False, bbox_inches='tight')
pdp_plot(numerical_plot_conf, f"ALE for {len(numerical_plot_conf)} numerical features")
# plt.savefig(os.path.join('.', 'plot', f'{sample_name}-1wale-numerical.png'), facecolor='white', transparent=False, bbox_inches='tight')
```
| github_jupyter |
```
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
from tensorflow.keras.layers import Input, Conv2D, Dense, Flatten, Dropout, GlobalMaxPooling2D, MaxPooling2D, BatchNormalization
from tensorflow.keras.models import Model
# Load in the data
cifar10 = tf.keras.datasets.cifar10
(x_train, y_train), (x_test, y_test) = cifar10.load_data()
x_train, x_test = x_train / 255.0, x_test / 255.0
y_train, y_test = y_train.flatten(), y_test.flatten()
print("x_train.shape:", x_train.shape)
print("y_train.shape", y_train.shape)
# number of classes
K = len(set(y_train))
print("number of classes:", K)
# Build the model using the functional API
i = Input(shape=x_train[0].shape)
x = Conv2D(32, (3, 3), activation='relu', padding='same')(i)
x = BatchNormalization()(x)
x = Conv2D(32, (3, 3), activation='relu', padding='same')(x)
x = BatchNormalization()(x)
x = MaxPooling2D((2, 2))(x)
# x = Dropout(0.2)(x)
x = Conv2D(64, (3, 3), activation='relu', padding='same')(x)
x = BatchNormalization()(x)
x = Conv2D(64, (3, 3), activation='relu', padding='same')(x)
x = BatchNormalization()(x)
x = MaxPooling2D((2, 2))(x)
# x = Dropout(0.2)(x)
x = Conv2D(128, (3, 3), activation='relu', padding='same')(x)
x = BatchNormalization()(x)
x = Conv2D(128, (3, 3), activation='relu', padding='same')(x)
x = BatchNormalization()(x)
x = MaxPooling2D((2, 2))(x)
# x = Dropout(0.2)(x)
# x = GlobalMaxPooling2D()(x)
x = Flatten()(x)
x = Dropout(0.2)(x)
x = Dense(1024, activation='relu')(x)
x = Dropout(0.2)(x)
x = Dense(K, activation='softmax')(x)
model = Model(i, x)
# Compile
# Note: make sure you are using the GPU for this!
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
# Fit with data augmentation
batch_size = 32
data_generator = tf.keras.preprocessing.image.ImageDataGenerator(width_shift_range=0.1, height_shift_range=0.1, horizontal_flip=True)
train_generator = data_generator.flow(x_train, y_train, batch_size)
steps_per_epoch = x_train.shape[0] // batch_size
r = model.fit(train_generator, validation_data=(x_test, y_test), steps_per_epoch=steps_per_epoch, epochs=50)
model.save("model1_image recognition_50epoch.h5")
# Plot loss per iteration
import matplotlib.pyplot as plt
plt.plot(r.history['loss'], label='loss')
plt.plot(r.history['val_loss'], label='val_loss')
plt.legend()
# Plot accuracy per iteration
plt.plot(r.history['accuracy'], label='acc')
plt.plot(r.history['val_accuracy'], label='val_acc')
plt.legend()
# Plot confusion matrix
from sklearn.metrics import confusion_matrix
import itertools
def plot_confusion_matrix(cm, classes,
normalize=False,
title='Confusion matrix',
cmap=plt.cm.Blues):
"""
This function prints and plots the confusion matrix.
Normalization can be applied by setting `normalize=True`.
"""
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
print("Normalized confusion matrix")
else:
print('Confusion matrix, without normalization')
print(cm)
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
fmt = '.2f' if normalize else 'd'
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, format(cm[i, j], fmt),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
plt.show()
p_test = model.predict(x_test).argmax(axis=1)
cm = confusion_matrix(y_test, p_test)
plot_confusion_matrix(cm, list(range(10)))
# label mapping
labels = '''airplane
automobile
bird
cat
deer
dog
frog
horse
ship
truck'''.split()
# Show some misclassified examples
misclassified_idx = np.where(p_test != y_test)[0]
i = np.random.choice(misclassified_idx)
plt.imshow(x_test[i], cmap='gray')
plt.title("True label: %s Predicted: %s" % (labels[y_test[i]], labels[p_test[i]]));
# Now that the model is so large, it's useful to summarize it
model.summary()
```
| github_jupyter |
# pMEC1135
Also referred to as pYPK0-C4. This vector is identical to [pMEC1136](pMEC1136.ipynb) but for a point mutation in XYL1 N272D. The systematic name of this vector is :
pYPK0-TEF1-SsXR_N272D-TDH3-SsXDH-PGI1-XK-FBA1-ScTAL1-PDC1
This vector expresses four genes and was assembled from four single gene expression cassettes:
Gene | Enzyme | Acronym | Cassette
-------------------------------------------------- |-------------------|---|-----|
[SsXYL1_N272D](http://www.ncbi.nlm.nih.gov/gene/4839234) |D-xylose reductase |XR | [pYPK0_TEF1_PsXYL1_N272D_TDH3](pYPK0_TEF1_PsXYL1_N272D_TDH3.ipynb)
[SsXYL2](http://www.ncbi.nlm.nih.gov/gene/4852013) |xylitol dehydrogenase |XDH | [pYPK0_TDH3_PsXYL2_PGI1](pYPK0_TDH3_PsXYL2_PGI1.ipynb)
[ScXKS1](http://www.yeastgenome.org/locus/S000003426/overview) |Xylulokinase |XK | [pYPK0_PGI1_ScXKS1_FBA1](pYPK0_PGI1_ScXKS1_FBA1.ipynb)
[ScTAL1](http://www.yeastgenome.org/locus/S000004346/overview) |Transaldolase |tal1p | [pYPK0_FBA1_ScTAL1_PDC1](pYPK0_FBA1_ScTAL1_PDC1.ipynb)
The vector [pMEC1135](pMEC1135.ipynb) is identical to this vector, but has a point mutation in XYL1.
[Yeast Pathway Kit Standard Primers](ypk_std_primers.ipynb)
```
from pydna.all import *
p567,p577,p468,p467,p568,p578,p775,p778,p167,p166 = parse("yeast_pahtway_kit_standard_primers.txt")
pYPK0 =read("pYPK0.gb")
pYPK0.cseguid()
from Bio.Restriction import ZraI, AjiI, EcoRV
p417,p626 =parse(''' >417_ScTEF1tpf (30-mer)
TTAAATAACAATGCATACTTTGTACGTTCA
>626_ScTEF1tpr_PacI (35-mer)
taattaaTTTGTAATTAAAACTTAGATTAGATTGC''', ds=False)
p415,p623 =parse(''' >415_ScTDH3tpf (29-mer)
TTAAATAATAAAAAACACGCTTTTTCAGT
>623_ScTDH3tpr_PacI (33-mer)
taattaaTTTGTTTGTTTATGTGTGTTTATTCG''', ds=False)
p549,p622 =parse(''' >549_ScPGI1tpf (27-mer)
ttaaatAATTCAGTTTTCTGACTGAGT
>622_ScPGI1tpr_PacI (28-mer)
taattaaTTTTAGGCTGGTATCTTGATT''', ds=False)
p409,p624 =parse(''' >409_ScFBA1tpf (37-mer)
TTAAATAATAACAATACTGACAGTACTAAATAATTGC
>624_ScFBA1tpr_PacI (29-mer)
taattaaTTTGAATATGTATTACTTGGTT''', ds=False)
p1 =read("pYPK0_TEF1_PsXYL1_N272D_TDH3.gb")
p2 =read("pYPK0_TDH3_PsXYL2_PGI1.gb")
p3 =read("pYPK0_PGI1_ScXKS1_FBA1.gb")
p4 =read("pYPK0_FBA1_ScTAL1_PDC1.gb")
cas1 =pcr( p167, p623, p1)
cas2 =pcr( p415, p622, p2)
cas3 =pcr( p549, p624, p3)
cas4 =pcr( p409, p166, p4)
pYPK0_E_Z, stuffer = pYPK0.cut((EcoRV, ZraI))
asm =Assembly( [pYPK0_E_Z, cas1, cas2, cas3, cas4] , limit = 61)
candidate = asm.assemble_circular()[0]
candidate.figure()
pw = candidate.synced(pYPK0)
len(pw)
pw.cseguid()
pw.name = "pMEC1135"
pw.description="pYPK0-TEF1-SsXR_N272D-TDH3-SsXDH-PGI1-XK-FBA1-ScTAL1-PDC1 pYPK0_C4"
pw.stamp()
pw.write("pMEC1135.gb")
```
| github_jupyter |
```
df1 <- read.csv('output/gene/metrics_clinical_plus_genes.csv', sep=',')
head(df1)
df2 <- read.csv('output/gene/metrics_clinical_only.csv', sep=',')
head(df2)
require(ggplot2)
mg <- merge(df1, df2, by=c("marker", "experiment"))
df <- data.frame(marker = mg$marker,
diff = mg$accuracy.x - mg$accuracy.y,
fold = mg$experiment)
result <- aggregate(diff ~ marker, data=df, FUN=mean)
colnames(result) <- c("marker", "mean")
tmp <- aggregate(diff ~ marker, data=df, FUN=sd)
colnames(tmp) <- c("marker", "sd")
result <- merge(result, tmp, by="marker")
result$marker = gsub('percent_', '', result$marker)
result <- result[result$mean > 0,]
ticks.line.color='grey30'
p <- ggplot(result) +
geom_bar(aes(marker, mean), stat="identity", position="dodge", alpha=.75) +
# scale_x_discrete(limits = rev(levels(result$marker))) +
# scale_y_log10() +
theme_bw() +
theme(panel.border = element_blank(),
panel.grid.major = element_blank(),
panel.grid.minor = element_blank(),
axis.line = element_line(colour=ticks.line.color),
text = element_text(size=10, family="serif"),
legend.key.size = unit(10, "pt"),
legend.position = 'bottom',
axis.text.x = element_text(size=10, angle=45, hjust=1, colour=ticks.line.color),
axis.text.y = element_text(size=10, colour=ticks.line.color),
axis.title.x = element_text(vjust=1),
axis.ticks = element_line(color=ticks.line.color),
plot.background = element_rect(fill = "transparent", color=NA), # bg of the plot
legend.box.background = element_rect(color=NA, fill = "transparent"), # get rid of legend panel bg
legend.background = element_rect(colour=NA, size=0),
legend.key = element_rect(fill="transparent", colour=NA) # get rid of key legend fill,
#and of the surrounding
) +
xlab("Clinical Marker") + ylab("Accuracy Gain")
print(mean(result$mean))
print(sd(result$mean))
print(min(result$mean))
print(max(result$mean))
ggsave('images/accuracy_gain.pdf', p, units='cm', width=18, height=7)
options(repr.plot.width=8, repr.plot.height=3)
p
# computing final results
df <- data.frame(feature = mg$marker,
diff = 1 - mg$accuracy.y / mg$accuracy.x,
fold = mg$experiment)
result <- aggregate(diff ~ feature, data=df, FUN=mean)
colnames(result) <- c("feature", "mean")
tmp <- aggregate(diff ~ feature, data=df, FUN=sd)
colnames(tmp) <- c("feature", "sd")
result <- merge(result, tmp, by="feature")
result$feature = gsub('percent_', '', result$feature)
result <- result[result$mean > 0,]
# replace "_" by " "
result$feature <- chartr("_", " ", result$feature)
result$feature[grep('cell markers', result$feature, fixed=T)] <- 'cell markers'
result$feature[grep('race', result$feature, fixed=T)] <- 'race'
result$feature[grep('wbc', result$feature, fixed=T)] <- 'wbc'
result <- aggregate(. ~ feature, FUN=mean, data=result)
# creating plot
ticks.line.color='grey40'
p <- ggplot(result) +
geom_bar(aes(feature, mean), fill='grey40', stat="identity", position="dodge", alpha=.75) +
scale_y_continuous(labels = scales::percent) +
xlab("Clinical Marker") +
ylab("Accuracy Gain") +
theme_bw() +
theme(panel.border = element_blank(),
panel.grid.major = element_blank(),
panel.grid.minor = element_blank(),
axis.line = element_line(colour=ticks.line.color),
text = element_text(size=10, family="serif"),
legend.key.size = unit(10, "pt"),
legend.position = 'bottom',
axis.text.x = element_text(size=10, angle=45, hjust=1, colour=ticks.line.color),
axis.text.y = element_text(size=10, colour=ticks.line.color),
axis.title.x = element_text(vjust=1),
axis.ticks = element_line(color=ticks.line.color),
plot.background = element_rect(fill = "transparent", color=NA), # bg of the plot
legend.box.background = element_rect(color=NA, fill = "transparent"), # get rid of legend panel bg
legend.background = element_rect(colour=NA, size=0),
legend.key = element_rect(fill="transparent", colour=NA) # get rid of key legend fill,
#and of the surrounding
)
# showing descriptive statistics
print(mean(result$mean))
print(sd(result$mean))
print(min(result$mean))
print(max(result$mean))
# exporting plot
ggsave('images/accuracy_percentual_gain.pdf', p, units='cm', width=18, height=7)
# showing plot
options(repr.plot.width=8, repr.plot.height=3)
p
t.test(df1$accuracy, df2$accuracy, alternative='greater')
sd(result$mean)
```
| github_jupyter |
```
import pandas as pd
import re
import seaborn as sns
import matplotlib.pyplot as plt
%matplotlib inline
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LinearRegression
from sklearn.feature_selection import RFE
from sklearn import preprocessing
from sklearn.linear_model import RidgeCV, LassoCV, Ridge, Lasso
data = pd.read_csv("train.csv")
print(data.shape)
data.head()
(data.isnull().sum()/data.shape[0])>0.50
for i,j in zip(data.columns,data.isnull().sum()/data.shape[0]>0.50):
if(j==True):
print(i," has been dropped")
data=data.drop(str(i),axis=1)
#These columns appear to be null
#Later implement the datatype matching using regex because int32,64,128 as well as float stuff
alter_list=[]
for i,j in zip(data.columns,data.isnull().sum()):
if(j>0):
if(data[i].dtypes == "float64" or data[i].dtypes == "int64"):
data_type="Numeric"
elif(data[i].dtypes == "object"):
data_type="String"
else:
data_type=""
print(i,j," Data type being : ",data_type)
alter_list.append((i,data_type))
# data['Age'] = data['Age'].fillna(data['Age'].mean())
# data['Embarked'] = data['Embarked'].fillna('S')
for i in alter_list:
if i[1] == "Numeric":
data[i[0]] = data[i[0]].fillna(data[i[0]].mean())
elif i[1]=="String":
print("Unique values in the column",i[0],data[i[0]].unique())
print("Number of unique vlaues in column",i[0], data[i[0]].nunique())
print("Number of each unique value occurances made in column",i[0]+":\n",data[i[0]].value_counts(dropna=False))
import math
toleration_limit = math.floor(data.shape[0]*0.1)
if data[i[0]].nunique()>toleration_limit:
data = data.drop(i[0],axis=1)
else:
maxm_occ = data[i[0]].value_counts(dropna=False)[:1].index.tolist()
print(maxm_occ[0])
data[i[0]].fillna(maxm_occ[0])
# data = data.drop(['Name', 'Ticket', 'Fare', 'PassengerId'], axis=1)
print("Analysing Unique Values in a columns as columns with more unique values are likely not to contribute")
for i in data.columns:
print("Analysing Column",i)
no_uniq = data[i].nunique()
print(data.shape[0])
print(no_uniq/data.shape[0])
if(no_uniq/data.shape[0]>0.1):
data = data.drop(str(i),axis=1)
else:
print("no uniq before",no_uniq)
print("target_col",data[i])
if (no_uniq>(data.shape[0]*(0.1))):
data[i] = pd.cut(x=data[i],bins=math.ceil(math.sqrt(no_uniq)),labels=False)
info_bins = pd.cut(x=data[i],bins=math.ceil(math.sqrt(no_uniq)))
print("info_bins",info_bins)
print("nuniq",data[i].nunique())
print(no_uniq)
target_column = "Pclass"
le = preprocessing.LabelEncoder()
data[target_column] = le.fit_transform(data[target_column])
X= pd.get_dummies(data.drop(target_column,1))
y=data[target_column]
data = pd.get_dummies(data)
print(data.head())
data.isnull().sum()
X
reg = LassoCV()
reg.fit(X, y)
print("Best alpha using built-in LassoCV: %f" % reg.alpha_)
print("Best score using built-in LassoCV: %f" %reg.score(X,y))
coef = pd.Series(reg.coef_, index = X.columns)
print("Lasso picked " + str(sum(coef != 0)) + " variables and eliminated the other " + str(sum(coef == 0)) + " variables")
imp_coef = coef.sort_values()
import matplotlib
matplotlib.rcParams['figure.figsize'] = (8.0, 10.0)
imp_coef.plot(kind = "barh")
plt.title("Feature importance using Lasso Model")
#Using Pearson Correlation
cor = data.corr()
imp_fact = abs(cor[target_column])
matplotlib.rcParams['figure.figsize'] = (8.0, 10.0)
imp_fact.plot(kind = "barh")
plt.show()
plt.figure(figsize=(10,10))
sns.heatmap(cor, annot=True, cmap=plt.cm.Reds)
plt.show()
#Correlation with output variable
cor_target = abs(cor[target_column])
#Selecting highly correlated features
relevant_features = cor_target[cor_target>0.2]
relevant_features
data.to_csv(r'cleaned_train.csv')
from sklearn.linear_model import LogisticRegression
no_uniq = data[target_column].nunique()
# if (no_uniq>2):
# data[target_column] = pd.cut(data[target_column],len(data),labels=False)
# print("nuniq",data[target_column].nunique())
# print("target_col",data[target_column])
survived_train = data[target_column]
data = data.drop(target_column,axis=1)
train_data,eval_data,labels,eval_labels = train_test_split(data, survived_train, random_state = 0)
# train_data = data.values[:600]
# labels = survived_train[:600]
# eval_data = data.values[600:]
# print(train_data)
# print(labels[0])
# # eval_labels = survived_train[600:]
# print(eval_labels)
if(no_uniq==2):
model = LogisticRegression(fit_intercept=True)
model.fit(train_data, labels)
eval_predictions = model.predict(eval_data)
else:
# from sklearn.tree import DecisionTreeClassifier
# model = DecisionTreeClassifier(max_depth = 3).fit(train_data, labels)
# eval_predictions = model.predict(eval_data)
from sklearn.svm import SVC
model= SVC(kernel = 'linear', C = 1).fit(train_data, labels)
eval_predictions = model.predict(eval_data)
print('Accuracy of the model on train data: {0}'.format(model.score(train_data, labels)))
print('Accuracy of the model on eval data: {0}'.format(model.score(eval_data, eval_labels)))
data
model.predict([[1,3,1,0,1,0,1,0,0]])
print(model.predict([[1,23,2,1,0,0,0,0,1]]))
print(eval_predictions)
```
| github_jupyter |
# Titanic - Deep Learning aproach
### obs: this code is from my github(https://github.com/dimitreOliveira/titanicDeepLearning) that's why it's so modular
> ### DEPENDENCIES
```
import csv
import re
import numpy as np
import pandas as pd
import tensorflow as tf
import matplotlib.pyplot as plt
from tensorflow.python.framework import ops
from sklearn import preprocessing
from sklearn.model_selection import train_test_split
```
### DATASET METHODS
```
def load_data(train_path, test_path):
"""
method for data loading
:param train_path: path for the train set file
:param test_path: path for the test set file
:return: a 'pandas' array for each set
"""
train_data = pd.read_csv(train_path)
test_data = pd.read_csv(test_path)
print("number of training examples = " + str(train_data.shape[0]))
print("number of test examples = " + str(test_data.shape[0]))
print("train shape: " + str(train_data.shape))
print("test shape: " + str(test_data.shape))
return train_data, test_data
def pre_process_data(df):
"""
Perform a number of pre process functions on the data set
:param df: pandas data frame
:return: updated data frame
"""
# setting `passengerID` as Index since it wont be necessary for the analysis
df = df.set_index("PassengerId")
# convert 'Sex' values
df['gender'] = df['Sex'].map({'female': 0, 'male': 1}).astype(int)
# We see that 2 passengers embarked data is missing, we fill those in as the most common Embarked value
df.loc[df.Embarked.isnull(), 'Embarked'] = df['Embarked'].mode()[0]
# Replace missing age values with median ages by gender
for gender in df['gender'].unique():
median_age = df[(df['gender'] == gender)].Age.median()
df.loc[(df['Age'].isnull()) & (df['gender'] == gender), 'Age'] = median_age
# convert 'gender' values to new columns
df = pd.get_dummies(df, columns=['gender'])
# convert 'Embarked' values to new columns
df = pd.get_dummies(df, columns=['Embarked'])
# bin Fare into five intervals with equal amount of values
# df['Fare-bin'] = pd.qcut(df['Fare'], 5, labels=[1, 2, 3, 4, 5]).astype(int)
# bin Age into seven intervals with equal amount of values
# ('baby','child','teenager','young','mid-age','over-50','senior')
bins = [0, 4, 12, 18, 30, 50, 65, 100]
age_index = (1, 2, 3, 4, 5, 6, 7)
df['Age-bin'] = pd.cut(df['Age'], bins, labels=age_index).astype(int)
# create a new column 'family' as a sum of 'SibSp' and 'Parch'
df['family'] = df['SibSp'] + df['Parch'] + 1
df['family'] = df['family'].map(lambda x: 4 if x > 4 else x)
# create a new column 'FTicket' as the first character of the 'Ticket'
df['FTicket'] = df['Ticket'].map(lambda x: x[0])
# combine smaller categories into one
df['FTicket'] = df['FTicket'].replace(['W', 'F', 'L', '5', '6', '7', '8', '9'], '4')
# convert 'FTicket' values to new columns
df = pd.get_dummies(df, columns=['FTicket'])
# get titles from the name
df['title'] = df.apply(lambda row: re.split('[,.]+', row['Name'])[1], axis=1)
# convert titles to values
df['title'] = df['title'].map({' Capt': 'Other', ' Master': 'Master', ' Mr': 'Mr', ' Don': 'Other',
' Dona': 'Other', ' Lady': 'Other', ' Col': 'Other', ' Miss': 'Miss',
' the Countess': 'Other', ' Dr': 'Other', ' Jonkheer': 'Other', ' Mlle': 'Other',
' Sir': 'Other', ' Rev': 'Other', ' Ms': 'Other', ' Mme': 'Other', ' Major': 'Other',
' Mrs': 'Mrs'})
# convert 'title' values to new columns
df = pd.get_dummies(df, columns=['title'])
df = df.drop(['Name', 'Ticket', 'Cabin', 'Sex', 'Fare', 'Age'], axis=1)
return df
def mini_batches(train_set, train_labels, mini_batch_size):
"""
Generate mini batches from the data set (data and labels)
:param train_set: data set with the examples
:param train_labels: data set with the labels
:param mini_batch_size: mini batch size
:return: mini batches
"""
set_size = train_set.shape[0]
batches = []
num_complete_minibatches = set_size // mini_batch_size
for k in range(0, num_complete_minibatches):
mini_batch_x = train_set[k * mini_batch_size: (k + 1) * mini_batch_size]
mini_batch_y = train_labels[k * mini_batch_size: (k + 1) * mini_batch_size]
mini_batch = (mini_batch_x, mini_batch_y)
batches.append(mini_batch)
# Handling the end case (last mini-batch < mini_batch_size)
if set_size % mini_batch_size != 0:
mini_batch_x = train_set[(set_size - (set_size % mini_batch_size)):]
mini_batch_y = train_labels[(set_size - (set_size % mini_batch_size)):]
mini_batch = (mini_batch_x, mini_batch_y)
batches.append(mini_batch)
return batches
```
### AUXILIARY MODEL METHODS
```
def create_placeholders(input_size, output_size):
"""
Creates the placeholders for the tensorflow session.
:param input_size: scalar, input size
:param output_size: scalar, output size
:return: X placeholder for the data input, of shape [None, input_size] and dtype "float"
:return: Y placeholder for the input labels, of shape [None, output_size] and dtype "float"
"""
x = tf.placeholder(shape=(None, input_size), dtype=tf.float32, name="X")
y = tf.placeholder(shape=(None, output_size), dtype=tf.float32, name="Y")
return x, y
def forward_propagation(x, parameters, keep_prob=1.0, hidden_activation='relu'):
"""
Implement forward propagation with dropout for the [LINEAR->RELU]*(L-1)->LINEAR-> computation
:param x: data, pandas array of shape (input size, number of examples)
:param parameters: output of initialize_parameters()
:param keep_prob: probability to keep each node of the layer
:param hidden_activation: activation function of the hidden layers
:return: last LINEAR value
"""
a_dropout = x
n_layers = len(parameters) // 2 # number of layers in the neural network
for l in range(1, n_layers):
a_prev = a_dropout
a_dropout = linear_activation_forward(a_prev, parameters['w%s' % l], parameters['b%s' % l], hidden_activation)
if keep_prob < 1.0:
a_dropout = tf.nn.dropout(a_dropout, keep_prob)
al = tf.matmul(a_dropout, parameters['w%s' % n_layers]) + parameters['b%s' % n_layers]
return al
def linear_activation_forward(a_prev, w, b, activation):
"""
Implement the forward propagation for the LINEAR->ACTIVATION layer
:param a_prev: activations from previous layer (or input data): (size of previous layer, number of examples)
:param w: weights matrix: numpy array of shape (size of current layer, size of previous layer)
:param b: bias vector, numpy array of shape (size of the current layer, 1)
:param activation: the activation to be used in this layer, stored as a text string: "sigmoid" or "relu"
:return: the output of the activation function, also called the post-activation value
"""
a = None
if activation == "sigmoid":
z = tf.matmul(a_prev, w) + b
a = tf.nn.sigmoid(z)
elif activation == "relu":
z = tf.matmul(a_prev, w) + b
a = tf.nn.relu(z)
elif activation == "leaky relu":
z = tf.matmul(a_prev, w) + b
a = tf.nn.leaky_relu(z)
return a
def initialize_parameters(layer_dims):
"""
:param layer_dims: python array (list) containing the dimensions of each layer in our network
:return: python dictionary containing your parameters "w1", "b1", ..., "wn", "bn":
Wl -- weight matrix of shape (layer_dims[l], layer_dims[l-1])
bl -- bias vector of shape (layer_dims[l], 1)
"""
parameters = {}
n_layers = len(layer_dims) # number of layers in the network
for l in range(1, n_layers):
parameters['w' + str(l)] = tf.get_variable('w' + str(l), [layer_dims[l - 1], layer_dims[l]],
initializer=tf.contrib.layers.xavier_initializer())
parameters['b' + str(l)] = tf.get_variable('b' + str(l), [layer_dims[l]], initializer=tf.zeros_initializer())
return parameters
def compute_cost(z3, y):
"""
:param z3: output of forward propagation (output of the last LINEAR unit), of shape (6, number of examples)
:param y: "true" labels vector placeholder, same shape as Z3
:return: Tensor of the cost function
"""
cost = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(logits=z3, labels=y))
return cost
def predict(data, parameters):
"""
make a prediction based on a data set and parameters
:param data: based data set
:param parameters: based parameters
:return: array of predictions
"""
init = tf.global_variables_initializer()
with tf.Session() as sess:
sess.run(init)
dataset = tf.cast(tf.constant(data), tf.float32)
fw_prop_result = forward_propagation(dataset, parameters)
fw_prop_activation = tf.nn.softmax(fw_prop_result)
prediction = fw_prop_activation.eval()
return prediction
def accuracy(predictions, labels):
"""
calculate accuracy between two data sets
:param predictions: data set of predictions
:param labels: data set of labels (real values)
:return: percentage of correct predictions
"""
prediction_size = predictions.shape[0]
prediction_accuracy = np.sum(np.argmax(predictions, 1) == np.argmax(labels, 1)) / prediction_size
return 100 * prediction_accuracy
def minibatch_accuracy(predictions, labels):
"""
calculate accuracy between two data sets
:param predictions: data set of predictions
:param labels: data set of labels (real values)
:return: percentage of correct predictions
"""
prediction_size = predictions.shape[0]
prediction_accuracy = np.sum(np.argmax(predictions, 1) == np.argmax(labels, 1)) / prediction_size
return 100 * prediction_accuracy
def l2_regularizer(cost, l2_beta, parameters, n_layers):
"""
Function to apply l2 regularization to the model
:param cost: usual cost of the model
:param l2_beta: beta value used for the normalization
:param parameters: parameters from the model (used to get weights values)
:param n_layers: number of layers of the model
:return: cost updated
"""
regularizer = 0
for i in range(1, n_layers):
regularizer += tf.nn.l2_loss(parameters['w%s' % i])
cost = tf.reduce_mean(cost + l2_beta * regularizer)
return cost
def build_submission_name(layers_dims, num_epochs, lr_decay,
learning_rate, l2_beta, keep_prob, minibatch_size, num_examples):
"""
builds a string (submission file name), based on the model parameters
:param layers_dims: model layers dimensions
:param num_epochs: model number of epochs
:param lr_decay: model learning rate decay
:param learning_rate: model learning rate
:param l2_beta: beta used on l2 normalization
:param keep_prob: keep probability used on dropout normalization
:param minibatch_size: model mini batch size (0 to do not use mini batches)
:param num_examples: number of model examples (training data)
:return: built string
"""
submission_name = 'ly{}-epoch{}.csv' \
.format(layers_dims, num_epochs)
if lr_decay != 0:
submission_name = 'lrdc{}-'.format(lr_decay) + submission_name
else:
submission_name = 'lr{}-'.format(learning_rate) + submission_name
if l2_beta > 0:
submission_name = 'l2{}-'.format(l2_beta) + submission_name
if keep_prob < 1:
submission_name = 'dk{}-'.format(keep_prob) + submission_name
if minibatch_size != num_examples:
submission_name = 'mb{}-'.format(minibatch_size) + submission_name
return submission_name
def plot_model_cost(train_costs, validation_costs, submission_name):
"""
:param train_costs: array with the costs from the model training
:param validation_costs: array with the costs from the model validation
:param submission_name: name of the submission (used for the plot title)
:return:
"""
plt.plot(np.squeeze(train_costs), label='Train cost')
plt.plot(np.squeeze(validation_costs), label='Validation cost')
plt.ylabel('cost')
plt.xlabel('iterations (per tens)')
plt.title("Model: " + submission_name)
plt.legend()
plt.show()
def plot_model_accuracy(train_accuracies, validation_accuracies, submission_name):
"""
:param train_accuracies: array with the accuracies from the model training
:param validation_accuracies: array with the accuracies from the model validation
:param submission_name: name of the submission (used for the plot title)
:return:
"""
plt.plot(np.squeeze(train_accuracies), label='Train accuracy')
plt.plot(np.squeeze(validation_accuracies), label='Validation accuracy')
plt.ylabel('accuracy')
plt.xlabel('iterations (per tens)')
plt.title("Model: " + submission_name)
plt.legend()
plt.show()
```
### MODEL
```
def model(train_set, train_labels, validation_set, validation_labels, layers_dims, learning_rate=0.01, num_epochs=1001,
print_cost=True, plot_cost=True, l2_beta=0., keep_prob=1.0, hidden_activation='relu', return_best=False,
minibatch_size=0, lr_decay=0, print_accuracy=True, plot_accuracy=True):
"""
:param train_set: training set
:param train_labels: training labels
:param validation_set: validation set
:param validation_labels: validation labels
:param layers_dims: array with the layer for the model
:param learning_rate: learning rate of the optimization
:param num_epochs: number of epochs of the optimization loop
:param print_cost: True to print the cost every 500 epochs
:param plot_cost: True to plot the train and validation cost
:param l2_beta: beta parameter for the l2 regularization
:param keep_prob: probability to keep each node of each hidden layer (dropout)
:param hidden_activation: activation function to be used on the hidden layers
:param return_best: True to return the highest params from all epochs
:param minibatch_size: size of th mini batch
:param lr_decay: if != 0, sets de learning rate decay on each epoch
:param print_accuracy: True to print the accuracy every 500 epochs
:param plot_accuracy: True to plot the train and validation accuracy
:return parameters: parameters learnt by the model. They can then be used to predict.
:return submission_name: name for the trained model
"""
ops.reset_default_graph() # to be able to rerun the model without overwriting tf variables
input_size = layers_dims[0]
output_size = layers_dims[-1]
num_examples = train_set.shape[0]
n_layers = len(layers_dims)
train_costs = []
validation_costs = []
train_accuracies = []
validation_accuracies = []
prediction = []
best_iteration = [float('inf'), 0, float('-inf'), 0]
best_params = None
if minibatch_size == 0 or minibatch_size > num_examples:
minibatch_size = num_examples
num_minibatches = num_examples // minibatch_size
if num_minibatches == 0:
num_minibatches = 1
submission_name = build_submission_name(layers_dims, num_epochs, lr_decay, learning_rate, l2_beta, keep_prob,
minibatch_size, num_examples)
x, y = create_placeholders(input_size, output_size)
tf_valid_dataset = tf.cast(tf.constant(validation_set), tf.float32)
parameters = initialize_parameters(layers_dims)
fw_output = forward_propagation(x, parameters, keep_prob, hidden_activation)
train_cost = compute_cost(fw_output, y)
train_prediction = tf.nn.softmax(fw_output)
fw_output_valid = forward_propagation(tf_valid_dataset, parameters, 1, hidden_activation)
validation_cost = compute_cost(fw_output_valid, validation_labels)
valid_prediction = tf.nn.softmax(fw_output_valid)
if l2_beta > 0:
train_cost = l2_regularizer(train_cost, l2_beta, parameters, n_layers)
validation_cost = l2_regularizer(validation_cost, l2_beta, parameters, n_layers)
if lr_decay != 0:
global_step = tf.Variable(0, trainable=False)
learning_rate = tf.train.inverse_time_decay(learning_rate, global_step=global_step, decay_rate=lr_decay,
decay_steps=1)
optimizer = tf.train.AdamOptimizer(learning_rate).minimize(train_cost, global_step=global_step)
else:
optimizer = tf.train.AdamOptimizer(learning_rate).minimize(train_cost)
init = tf.global_variables_initializer()
with tf.Session() as sess:
sess.run(init)
for epoch in range(num_epochs):
train_epoch_cost = 0.
validation_epoch_cost = 0.
minibatches = mini_batches(train_set, train_labels, minibatch_size)
for minibatch in minibatches:
(minibatch_X, minibatch_Y) = minibatch
feed_dict = {x: minibatch_X, y: minibatch_Y}
_, minibatch_train_cost, prediction, minibatch_validation_cost = sess.run(
[optimizer, train_cost, train_prediction, validation_cost], feed_dict=feed_dict)
train_epoch_cost += minibatch_train_cost / num_minibatches
validation_epoch_cost += minibatch_validation_cost / num_minibatches
validation_accuracy = accuracy(valid_prediction.eval(), validation_labels)
train_accuracy = accuracy(prediction, minibatch_Y)
if print_cost is True and epoch % 500 == 0:
print("Train cost after epoch %i: %f" % (epoch, train_epoch_cost))
print("Validation cost after epoch %i: %f" % (epoch, validation_epoch_cost))
if print_accuracy is True and epoch % 500 == 0:
print('Train accuracy after epoch {}: {:.2f}'.format(epoch, train_accuracy))
print('Validation accuracy after epoch {}: {:.2f}'.format(epoch, validation_accuracy))
if plot_cost is True and epoch % 10 == 0:
train_costs.append(train_epoch_cost)
validation_costs.append(validation_epoch_cost)
if plot_accuracy is True and epoch % 10 == 0:
train_accuracies.append(train_accuracy)
validation_accuracies.append(validation_accuracy)
if return_best is True:
if validation_epoch_cost < best_iteration[0]:
best_iteration[0] = validation_epoch_cost
best_iteration[1] = epoch
best_params = sess.run(parameters)
if validation_accuracy > best_iteration[2]:
best_iteration[2] = validation_accuracy
best_iteration[3] = epoch
best_params = sess.run(parameters)
if return_best is True:
parameters = best_params
else:
parameters = sess.run(parameters)
print("Parameters have been trained, getting metrics...")
train_accuracy = accuracy(predict(train_set, parameters), train_labels)
validation_accuracy = accuracy(predict(validation_set, parameters), validation_labels)
print('Train accuracy: {:.2f}'.format(train_accuracy))
print('Validation accuracy: {:.2f}'.format(validation_accuracy))
print('Lowest validation cost: {:.2f} at epoch {}'.format(best_iteration[0], best_iteration[1]))
print('Highest validation accuracy: {:.2f} at epoch {}'.format(best_iteration[2], best_iteration[3]))
submission_name = 'tr_acc-{:.2f}-vd_acc{:.2f}-'.format(train_accuracy, validation_accuracy) + submission_name
if return_best is True:
print('Lowest cost: {:.2f} at epoch {}'.format(best_iteration[0], best_iteration[1]))
if plot_cost is True:
plot_model_cost(train_costs, validation_costs, submission_name)
if plot_accuracy is True:
plot_model_accuracy(train_accuracies, validation_accuracies, submission_name)
return parameters, submission_name
```
### Load data
```
TRAIN_PATH = '../input/train.csv'
TEST_PATH = '../input/test.csv'
train, test = load_data(TRAIN_PATH, TEST_PATH)
CLASSES = 2
train_dataset_size = train.shape[0]
# The labels need to be one-hot encoded
train_raw_labels = pd.get_dummies(train.Survived).as_matrix()
```
### Pre process data
```
train = pre_process_data(train)
test = pre_process_data(test)
# drop unwanted columns
train_pre = train.drop(['Survived'], axis=1).as_matrix().astype(np.float)
test_pre = test.as_matrix().astype(np.float)
```
### Normalize data
```
# scale values
standard_scaler = preprocessing.StandardScaler()
train_pre = standard_scaler.fit_transform(train_pre)
test_pre = standard_scaler.fit_transform(test_pre)
# data split
X_train, X_valid, Y_train, Y_valid = train_test_split(train_pre, train_raw_labels, test_size=0.3, random_state=1)
```
### Model parameters
```
# hyperparameters
input_layer = train_pre.shape[1]
output_layer = 2
num_epochs = 10001
learning_rate = 0.0001
train_size = 0.8
# layers_dims = [input_layer, 256, 128, 64, output_layer]
layers_dims = [input_layer, 512, 128, 64, output_layer]
```
### Train model
```
parameters, submission_name = model(X_train, Y_train, X_valid, Y_valid, layers_dims, num_epochs=num_epochs,
learning_rate=learning_rate, print_cost=False, plot_cost=True, l2_beta=0.1,
keep_prob=0.5, minibatch_size=0, return_best=True, print_accuracy=False,
plot_accuracy=True)
```
### Make predictions
```
final_prediction = predict(test_pre, parameters)
submission = pd.DataFrame({"PassengerId":test.index.values})
submission["Survived"] = np.argmax(final_prediction, 1)
submission.to_csv("submission.csv", index=False)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/jagan-mathematics/NLP-Neural_Architechture_Search/blob/master/Notebooks/Model%20search%20for%20kick%20start%20experiment.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab"/></a>
## Introduction
Model search (MS) is a framework that implements AutoML algorithms for model architecture search at scale. It aims to help researchers speed up their exploration process for finding the right model architecture for their classification problems (i.e., DNNs with different types of layers).
The library enables you to:
- Run many AutoML algorithms out of the box on your data - including automatically searching for the right model architecture, the right ensemble of models and the best distilled models.
- Compare many different models that are found during the search.
- Create you own search space to customize the types of layers in your neural networks.
- The technical description of the capabilities of this framework are found in InterSpeech paper.
While this framework can potentially be used for regression problems, the current version supports classification problems only. Let's start by looking at some classic classification problems and see how the framework can automatically find competitive model architectures.
In this notebook we have provided complete implementation to search the model for text classification task.
```
!pip install -U tensorboard_plugin_profile
!git clone https://github.com/google/model_search.git
%cd model_search/
!pip install -r requirements.txt
%%bash
protoc --python_out=./ model_search/proto/phoenix_spec.proto
protoc --python_out=./ model_search/proto/hparam.proto
protoc --python_out=./ model_search/proto/distillation_spec.proto
protoc --python_out=./ model_search/proto/ensembling_spec.proto
protoc --python_out=./ model_search/proto/transfer_learning_spec.proto
import sys
from absl import app
# Addresses `UnrecognizedFlagError: Unknown command line flag 'f'`
sys.argv = sys.argv[:1]
# `app.run` calls `sys.exit`
try:
app.run(lambda argv: None)
except:
pass
!pwd
import model_search
from model_search import constants
from model_search import single_trainer
from model_search.data import data
import tensorflow as tf
import json
import os
from typing import List, Union
%load_ext tensorboard
```
## Tokenizer
We had faced lot of problems while using keras tokenizer, so we decided to create my own. It will do some basic processing like lowering text and removing special charaters. Below is the implementation of that.
```
class TextProcessing:
def __init__(self, vocab_size=1, max_sequence_length=1, pattern = '[!"#$%&()*,-./:;<=>?@[\\]^_`{|}~\t\n]'):
self._max_sequence_length = max_sequence_length
self.word_to_idx = None
self.idx_to_word = None
self.pattern = pattern
self.vocab_size = vocab_size
self.__types = {'max_sequence_length': tf.io.FixedLenFeature([], tf.int64),
'vocab_size': tf.io.FixedLenFeature([], tf.int64),
'pattern': tf.io.FixedLenFeature([], tf.string),
'words': tf.io.VarLenFeature(tf.string),
'count': tf.io.VarLenFeature(tf.int64),
'words_to_idx_w': tf.io.VarLenFeature(tf.string),
'words_to_idx_i': tf.io.VarLenFeature(tf.int64),
'idx_to_word_w': tf.io.VarLenFeature(tf.string),
'idx_to_word_i': tf.io.VarLenFeature(tf.int64)
}
@tf.function
def __pad_sequence(self, sequence, max_sequence):
sequence = tf.squeeze(sequence)
if tf.size(sequence) > max_sequence:
sequence = tf.slice(sequence, [0], [max_sequence])
axis_1 = tf.zeros_like(sequence, dtype = tf.int32)
idx = tf.stack((axis_1, tf.range(tf.size(axis_1))), axis = -1)
idx = tf.cast(idx, dtype=tf.int64)
return tf.sparse.to_dense(tf.sparse.SparseTensor(indices=idx, values=sequence, dense_shape=[1,max_sequence]))
def __parse_example(self, record):
return tf.io.parse_single_example(record, self.__types)
def from_config(self, filename):
raw_data = tf.data.TFRecordDataset(filename)
raw_record = next(iter(raw_data))
dic = self.__parse_example(raw_record)
for item, value in dic.items():
if isinstance(value, tf.SparseTensor):
dic[item] = tf.sparse.to_dense(value)
elif isinstance(value, tf.Tensor):
dic[item] = value.numpy()
self.words = dic['words']
self.count = dic['count']
self.word_to_idx = tf.lookup.StaticHashTable(
initializer=tf.lookup.KeyValueTensorInitializer(
keys=dic['words_to_idx_w'],
values=tf.cast(dic['words_to_idx_i'], tf.int32),
),
default_value=tf.constant(-1),
name="word_to_idx"
)
self.idx_to_word = tf.lookup.StaticHashTable(
initializer=tf.lookup.KeyValueTensorInitializer(
keys=tf.cast(dic['idx_to_word_i'], tf.int32),
values=dic['idx_to_word_w'],
),
default_value=tf.constant(''),
name="idx_to_word"
)
self._max_sequence_length = dic['max_sequence_length']
self.pattern = dic['pattern'].decode()
self.vocab_size = dic['vocab_size']
def export(self, filename):
words_to_idx = self.word_to_idx.export()
idx_to_word = self.idx_to_word.export()
features = {
'max_sequence_length' : tf.train.Feature(int64_list=tf.train.Int64List(value=[self._max_sequence_length])),
'vocab_size' : tf.train.Feature(int64_list=tf.train.Int64List(value=[self.vocab_size])),
'pattern': tf.train.Feature(bytes_list=tf.train.BytesList(value=[self.pattern.encode()])),
'words': tf.train.Feature(bytes_list=tf.train.BytesList(value=self.words.numpy().tolist())),
'count': tf.train.Feature(int64_list=tf.train.Int64List(value=self.count.numpy().tolist())),
'words_to_idx_w': tf.train.Feature(bytes_list=tf.train.BytesList(value=words_to_idx[0].numpy().tolist())),
'words_to_idx_i': tf.train.Feature(int64_list=tf.train.Int64List(value=words_to_idx[1].numpy().tolist())),
'idx_to_word_w': tf.train.Feature(bytes_list=tf.train.BytesList(value=idx_to_word[1].numpy().tolist())),
'idx_to_word_i': tf.train.Feature(int64_list=tf.train.Int64List(value=idx_to_word[0].numpy().tolist())),
}
example_proto = tf.train.Example(features=tf.train.Features(feature=features))
writer = tf.io.TFRecordWriter(filename)
writer.write(example_proto.SerializeToString())
def adapt(self, dataset):
dataset = dataset = dataset.flat_map(lambda x , y : tf.data.Dataset.from_tensor_slices(self._split(x)))
dataset = dataset.flat_map(lambda x :tf.data.Dataset.from_tensor_slices(x))
words_list = list(map(lambda x : x.decode(), list(dataset.as_numpy_iterator())))
words, idx, count = tf.unique_with_counts(words_list)
self.words = words
self.count = count
top_k_count = tf.math.top_k(count, k = self.vocab_size-1)
sorted_words = tf.gather(words,top_k_count.indices)
word_idx = tf.range(1, self.vocab_size)
self.word_to_idx = tf.lookup.StaticHashTable(
initializer=tf.lookup.KeyValueTensorInitializer(
keys=sorted_words,
values=word_idx,
),
default_value=tf.constant(-1),
name="word_to_idx"
)
self.idx_to_word = tf.lookup.StaticHashTable(
initializer=tf.lookup.KeyValueTensorInitializer(
keys=word_idx,
values=sorted_words,
),
default_value=tf.constant(''),
name="idx_to_word"
)
def _split(self, text):
text = tf.strings.lower(text)
text = tf.strings.regex_replace(text, self.pattern,'')
text = tf.strings.regex_replace(text, ' +',' ')
text = tf.strings.strip(text)
text = tf.sparse.to_dense(tf.compat.v1.strings.split(text, ' '))
return text
def __text_to_sequence(self, text):
text = self._split(text)
text = self.word_to_idx.lookup(text)
text = tf.squeeze(text)
text = tf.gather(text, tf.where(tf.not_equal(text, -1)))
text = tf.squeeze(text)
return text
def encode(self, text):
text = self.__text_to_sequence(text)
return text
def int_vectorize_text(self, text, label):
text = self.encode(text)
text = self.__pad_sequence(text, self._max_sequence_length)
# text = tf.cast(text, dtype = tf.dtypes.float32)
label = tf.cast(label, dtype = tf.dtypes.float32)
text = tf.reshape(text, [self._max_sequence_length, 1])
return text, label
```
## Data Provider
provider class will be utilized by model_search to invoke input function and input layer function.
There are some limitation in the class that I created it only supports TSV or file data with passage at first column and label at second.
```Note : by default it uses tab deliminator you can change that in field_delim parameter```
```
class provider(data.Provider):
"""A tsv data provider."""
def __init__(self, input_file, default_records, feature_idx:List[Union['idx']], batch_size, label,
tokenizer_path=None, field_delim='\t', header=True, testing = False):
self._input_file = input_file
self._default_records = default_records
self._feature_idx = feature_idx
self._field_delim = field_delim
self._header = header
self._batch_size = batch_size
self._label = label
self._testing = testing
self.tokenizer_path = tokenizer_path
self.processor = TextProcessing()
if tokenizer_path is not None:
self.processor.from_config(tokenizer_path)
def get_input_fn(self, hparams, mode, batch_size):
del hparams
def input_fn(params=None):
del params
def encode_pyfn(text, label):
text_encoded, target = tf.py_function(self.processor.int_vectorize_text,
inp=[text, label],
Tout=(tf.int32, tf.float32))
return {'text':text_encoded}, target
dataset = tf.data.experimental.CsvDataset(self._input_file,
record_defaults= self._default_records,
header=self._header,
field_delim=self._field_delim,
select_cols=self._feature_idx)
if mode == tf.estimator.ModeKeys.TRAIN:
dataset = dataset.shuffle(100 * batch_size)
if not self._testing and mode is not None:
assert self.tokenizer_path is not None, 'tokenizer configuration is not found. create tokenizer using "TextProcessing"'
dataset = dataset.map(encode_pyfn, num_parallel_calls=tf.data.experimental.AUTOTUNE)
elif self._testing and mode is not None:
assert self.tokenizer_path is not None, 'tokenizer configuration is not found. create tokenizer using "TextProcessing"'
dataset = dataset.map(self.processor.int_vectorize_text, num_parallel_calls=tf.data.experimental.AUTOTUNE)
if mode == tf.estimator.ModeKeys.TRAIN:
dataset = dataset.batch(batch_size, drop_remainder=True).prefetch(tf.data.experimental.AUTOTUNE)
return dataset
return input_fn
def get_serving_input_fn(self, hparams):
"""Returns an `input_fn` for serving in an exported SavedModel.
Args:
hparams: tf.HParams object.
Returns:
Returns an `input_fn` that takes no arguments and returns a
`ServingInputReceiver`.
"""
tf.compat.v1.disable_eager_execution()
features = {
'text':
tf.compat.v1.placeholder(
tf.int32, [None, self.processor._max_sequence_length, 1],
'text')
}
return tf.estimator.export.build_raw_serving_input_receiver_fn(
features=features)
def number_of_classes(self):
return self._label
def get_input_layer_fn(self, problem_type):
# initialize = embedding_initialize(self.processor.word_to_idx.export()[0], (50, ), '/content/drive/MyDrive/experiment_nlp/glove_initializer/glove.6B/glove.6B.50d.tfrecord-00000-of-00001')
def input_layer_fn(features,
is_training,
scope_name="Phoenix/Input",
lengths_feature_name=None):
input_feature = tf.reshape(features['text'], [-1, self.processor._max_sequence_length, 1])
features = {
'text': tf.sparse.from_dense(input_feature)
}
one_hot_layer = tf.feature_column.sequence_categorical_column_with_identity('text', num_buckets=self.processor.vocab_size)
text_embedding = tf.feature_column.embedding_column(one_hot_layer,
dimension=128)
# initializer = initialize)
columns = [text_embedding]
sequence_input_layer = tf.keras.experimental.SequenceFeatures(columns, name = scope_name)
sequence_input, sequence_length = sequence_input_layer(features, training=is_training)
return sequence_input, sequence_length
return input_layer_fn
```
## parameter setup
```
#@title Parameters
dataset_file = 'dataset/train.tsv' #@param {type:"string"}
root_dir = 'Model searching' #@param {type : "string"}
experiment_name = 'example' #@param {type : "string"}
tokenizer_path = 'tokenizer_config.tfrecord' #@param {type: "string"}
# index of the columns to be selected for the file
selected_columns = [2, 3] #@param {type:"raw"}
default_records = ['', 0.0] #@param {type:"raw"}
batch_size = 32 #@param {type: "integer"}
# no of labels
label = 5 #@param {type: "integer"}
vocab_size = 10000 #@param {type: "integer"}
max_sequence_length = 250 #@param {type: "integer"}
number_models = 5 #@param {type: "integer"}
train_steps = 350 #@param {type: "integer"}
eval_steps = 100 #@param {type: "integer"}
vocab_size += 1
```
## Input pipeline analysis
I had created my own Tokenizer class, it is important to make sure that our pipeline is not spending much time in ```Dataset.map``` which transforms our text to tokens and also our pipeline is not suffering with any bottle-necks.
we found out the shuffling data before transformation decreases data fetching time.
```
if not os.path.exists(tokenizer_path):
data_provider = provider(
input_file = dataset_file,
default_records = default_records,
feature_idx = selected_columns,
batch_size=batch_size,
label = label,
)
input_func = data_provider.get_input_fn(None, None, 1)
dataset = input_func()
tokenizer = TextProcessing(vocab_size = vocab_size, max_sequence_length = max_sequence_length)
print('[INFO] => started adapting')
tokenizer.adapt(dataset)
print('[INFO] => Exporting configs...')
tokenizer.export(tokenizer_path)
del data_provider, input_func, dataset, tokenizer
data_fn = provider(
input_file = dataset_file,
default_records = default_records,
feature_idx = selected_columns,
batch_size=batch_size,
label = label,
tokenizer_path = tokenizer_path,
testing = True
)
input_fn = data_fn.get_input_fn(None, 'train', 32)
train_ds = input_fn(None)
```
Let define dummy model with one lstm layer to profile our input pipeline.
```
model = tf.keras.models.Sequential([
tf.keras.layers.Input((250, 1)),
tf.keras.layers.LSTM(units = 23),
tf.keras.layers.Dense(label, activation='softmax')
])
model.compile(
loss='sparse_categorical_crossentropy',
optimizer=tf.keras.optimizers.Adam(0.001),
metrics=['accuracy']
)
from datetime import datetime
logs = "logs/" + datetime.now().strftime("%Y%m%d-%H%M%S")
tboard_callback = tf.keras.callbacks.TensorBoard(log_dir = logs,
histogram_freq = 1,
profile_batch = '32,64')
model.fit(train_ds, epochs=2,
callbacks = [tboard_callback])
```
Go to the profiler tab and explore the performance of input pipeline
```
%tensorboard --logdir=logs
```
## model_search
Here come the most important part. we going to select the best model for our dataset using 'model_search' by google. You will able to find out best model when number_model parameter value is more higher. but the best model comes at the cost of computation and time.
```
trainer = single_trainer.SingleTrainer(
provider(
input_file = dataset_file,
default_records = default_records,
feature_idx = selected_columns,
batch_size=batch_size,
label = label,
tokenizer_path = tokenizer_path
),
spec='/content/model_search/model_search/configs/rnn_last_config.pbtxt')
trainer.try_models(
number_models=number_models,
train_steps=train_steps,
eval_steps=eval_steps,
root_dir=root_dir,
batch_size=batch_size,
experiment_name=experiment_name,
experiment_owner="model_search_user")
```
## Model search Report
```
path = os.path.join(root_dir, experiment_name)
trial_ids = os.listdir(path)
records = []
for trial in trial_ids:
file = os.path.join(path, trial, 'trial.json')
if os.path.isdir(os.path.join(path, trial)):
with open(file, 'r') as handler:
data = json.load(handler)
temp_ = data['metrics']['metrics']
if temp_:
row = {'accuracy' : temp_['accuracy']['observations'][0]['value'][0],
'global_step' : temp_['global_step']['observations'][0]['value'][0],
'loss': temp_['loss']['observations'][0]['value'][0],
'num_parameters': temp_['loss']['observations'][0]['value'][0],
'score': data['score'],
'status' : data['status'],
'trial_id' : data['trial_id']
}
records.append(row)
import pandas as pd
pd.DataFrame(records)
```
**Summary**
- This notebook provides an complete implementation of model search to find the best fit model for text classification task.
- We have created our own tokenizer class because we had some hard times with keras tokenizer.
- Data provider that we implemented has some limitations like we can only able to use tsv or csv file format, we have to provide file as text first label second format.
**Future work**
- We are working on option to train model with pre-trained embedding layer. eg: glove, bert etc.,
- multiple options to load dataset from provider
- general aproach for text generation and text segmentation tasks.
- adding option to pass preprocessing function and stop words removal property in Tokenizer class
| github_jupyter |
# Basics in Python
* as we remember, python is an interpreted programming language without compiler building process
* python is a multiparadigm language, everything is possible --- but overall its working like an object oriented programming language
* datatypes are dynamically handled
## more datatypes
lets have a look at a few more datatypes and special datatypes
```
# as we remember we dont need to clarify a datatype of a variable
my_number = 42 # is automatically interpreted as integer
# expert knowledge: we generate a object variable from the builtin class Integer it is possible to use functions from Integer class
my_float_number = 42.42 # with the dot identifier it is possible to create floating numbers this variable is interpreted as float
my_text = "example text" # this variable is interpreted as string
# expert knowledge: a string is a chain of chars (list of chars) char is the simple datatype, string is a complex char datatype
my_boolean = True # this variable is interpreted as boolean
# to show the datatype of our variable we can use the builtinfunction type() inside a print statement
print(type(my_number), type(my_float_number), type(my_text), type(my_boolean))
```
As we see in the printout all datatypes are types of classes, that means every datatype comes with class functions. We will see later in this training how this work
```
# in some programs it is possible to enforce a datatype
# because of dynamically handling of datatype variables it could happen that python will change a variable type during program runtime
# in case of datatype changing in special programs, error could appear
# to enforce a datatype we use signal words (builtin functions)
my_variable_1 = int(42) # this variable is an integer
my_variable_2 = int(42.42) # also an integer the floating point part will be delete
my_variable_3 = float(42) # floating point datatype
my_variable_4 = str(42) # a number as a string datatype
my_variable_5 = bool(42) # a boolean datatype if the number is > 0 its always true in Python Boolean statements are True or False (case-sensitive)
my_variable_6 = bool(" ") # whitespace string is True --> behind a whitespace is a character ascii sign which is a True context
my_variable_7 = bool("") # empty string is always False
print( my_variable_1, type(my_variable_1), "\n",
my_variable_2, type(my_variable_2), "\n",
my_variable_3, type(my_variable_3), "\n",
my_variable_4, type(my_variable_4), "\n",
my_variable_5, type(my_variable_5), "\n",
my_variable_6, type(my_variable_6), "\n",
my_variable_7, type(my_variable_7) )
# the naming of a variable does have a huge variance
# it is possible to use "_" underline signs between words
# the possible variable naming styles are:
# --> camelCase
# --> PascalCase
# --> snake_case
# it is also possible to use numbers inside a variable name but not for the first sign
# the first variable letter need to be an underline or a normal letter
# reserved functions or special signs are not allowed
# if u using a reserved function (built-in) or reserved word (str, int, ...) u overwrite the functionallity --- this will devastate your program
# during development most of all programmer using temporary variable names, in production code we need to keep the topic information for our variable
# because of nearly no limitation for the variable name i prefer to use snake_case with lower letter with descritpion of the variable topic
# if the variable does have the topic to keep a sensor gps signal frame we call the variable name
my_gps_sensor_frame = "some gps sensor data" # gps could have diferent datatypes but this is not the topic at the moment
# some developer keep the datatype inside the variable also
str_my_gps_sensor_frame = "..." # depends on you how to style your variables
# capital letter variables are normally cons inside a program
MY_AWESOME_CIRCLE_CONSTANT = 3.1415 # short form of Pi for calculation
WINDOW_WIDTH = 640 # window size paramter in guis
WINDOW_HEIGHT = 480 # window size paramter in guis
# sometimes we dont know how to store or use a variable
# in such a case it is possible to use a placeholder variable instead of a complete variable name
# the common placeholder for variables is "_"
_ = "dont like to use a this variable"
# in python it is also possible to use multiple variable allocations
var_1 = var_2 = var_3 = 42 # all 3 variables are integer with value of 42
# we learned that a variable (a single value without variable storage also!) are objects of classes
# if we keep in mind that everything in python is handled the object oriented way we understand how variables are stored and used
# the interpreter decide what kind of datatype our variable is and create an object from the specific class dynamically
# to learn what kind of class function we could use, we need to have a look at the specific class documentation
# the builtin function for selecting the class function is help()
# it is possible to use help on the class directly, the function, variable or a value
help(int)
# help shows us the docstring documentation of the class/function/variable
# under the method section we see a few methods with syntax "_" "_" "name" "_" "_"
# these double underlined functions are special system relevant functions
# they are activated with special code instructions, for example if we program
# print(my_variable)
# the internal __str__ function is called
#
# my_var1 + my_var2
# the internal __add__ function is called
#
# function names with a start of 2 underline signs are private and hidden functions and not direct callable
# in class context everything with double underline in the beginning is hidden and only accessable inside the class
# functions which we are able to use normally start with a normal letter and always have round brackets (with or without parameters)
#
# functions from an object are callable by typing a dot after the object
# this process could explain as a concatination of objects with functions
# if we concat a object to a function, the return "object" of the function is also able to add a concatination
# this codestyle is typically python in oneliner code instruction
#
# lets have a look to an example
my_string_variable = "this is a test string"
print("Normal: \n", my_string_variable)
print("function call with capital first letter: \n", my_string_variable.capitalize())
print("function call uppercase: \n", my_string_variable.upper())
print("function call to upper, and concat the result of upper with a function call swap which returns upper case to lower case: \n", my_string_variable.upper().swapcase())
print("function call with capital and swap: \n", my_string_variable.capitalize().swapcase())
# concatination of functions are very strong and multiple times usable
# try to find the limitation of concatination :)
```
## a few special datatypes
* we already learned a few basic datatypes, with external modules it is also possible to expand the number of datatypes in all ways
* special datatypes are normally called as complex
* in programming languages complex datatypes are structures with subdatatypes or compound datatypes
```
# type of lists
# if we want to work with lists it is possible to use the list datatype
# in python we have a few list objects which are enforced by different brackets
# the classical list is enforced by a square bracket
# all list keeping objects of other datatypes/variables
my_list_1 = [1,2,3,4] # this variable is a normal list, an array
my_list_2 = [1,2, "text", 42.42] # it is also possible to mix the elements of datatype
print("normal list:", my_list_2, type(my_list_2))
# standard lists are mutable, that means we are able to directly add more values without overwright the variable
# to add a value to the list, we can use .append(value) function
my_list_1.append(42)
print("after append:",my_list_1)
# the second list type we want to have a look on is a tuple
# a tuple list is a immutable list datatype
# it is also possible to mix the sub datatypes
# tuples are enforced by normal brackets
my_list_3 = (42, 55, "GMT", "zone")
print("tuple list", my_list_3, type(my_list_3))
# we could use a tuple datatype if we want to struct different datatypes to a fix variable
# it is also possible to use a tuple for multiple return inside function
# for example a gps sensor dataset consist of longitude, latitude, time and height --- the perfect datatype after sensor readout is tuple
# tuple lists are not changeable --> immutable, we always need to overwrite our variable or convert back to lists with using the list() function
# the third datatype is used in different ways
# a set is a unique value list of possible mixed types
# a set list is immutable and will keep only unique values
# to enforce a set we need to program with curly brackets
my_list_4 = {42,55 , 20, 42, "test", 17, 0, 4} # keep an eye on 42 its 2 times inside the declaration
print("set list", my_list_4, type(my_list_4)) # 42 we see only once
# with sets it is also possible to use mathematically set functions like join, innerjoin a.s.o - try it out by yourself :)
# sets are very special lists it is also possible to use the set property to expand our posibilities
# it is possible to declare a new code block with a ":" sign... in case of lists the code block instruction is interpreted as a new declaration
# inside sets this means it is possible to interprete the value left from the double dot as a unique variable and the part right of the double dot as a value to the variable
# this special code instruction is better known as key value list
# inside a key value list the key need to be unique (set property), the value could be what ever we want (variable property)
my_list_5 = {42: "test value", "banana":[1,2,3], "sth.more":{ "subthing_1": 42.42, "subthing_2":1704}}
print("dictionary list:", my_list_5, type(my_list_5))
# in python programming language a key value list is better known as "Dictionary"
# lists or list variants (datatype that could be a list too) are iterable
# iterable means we are able to call an element of a list directly
# the list call instruction is a square bracket
# with tuples and arrays it is possible to ask for an element by calling the element position
# we start counting with 0 that means the first element of a list is called with [0]
print("element 1 of list 1:", my_list_1[0]) # the first element of list_1 counting from left to right
# it is also possible to call the last element (or first element from the right position)
print("last element of list 3", my_list_3[-1])
# inside the square bracket it is possible to have a range of values with stepping
# for a range of values we need to use the doubledot instruction again
# if we want to have "from value_1 to value_3" we need to call [0:2] left to right orientation
# the stepping is also callable with again a double dot [0:20:2] --> every second value from 0 to 20 element
print("first two elements of list 2", my_list_2[:2]) # first two elements of list 2
# sets are special list, to call an element of a set we need to use a function
# it is only possible to get a set value element by element in order
# the function pop will show us the value elementwise
# after pop using the value is deleted from the set
print("set before pop:", my_list_4)
print("pop value:",my_list_4.pop())
print("set after pop:",my_list_4)
# a dictionary works a little bit different
# we are able to call the key value directly on the dictionary
# if we using the normal iteration counter (0,1,2,3,...) we get an error
# to get values of the dictionary it is necessary to use a function or direct key call
print("dictionary:",my_list_5)
print("keys of dictionary:",my_list_5.keys())
print("call a specific key from dictionary:",my_list_5["sth.more"])
# it is also possible to concat list calls if we have a sublist inside a list
# because of multiple datatype mix and declaration it is possible to have multidimensional lists
# inside our dictionare we have a subdictionary we want to call
print("subdictionary call with listconcat:", my_list_5["sth.more"]["subthing_2"])
# if we want to change a value inside a dictionary we only need to call the keys and declarate new values to the key
# it is also possible to add new keys if we calling a non existing key with a declaration
my_list_5["new_key"] = (1,2,3,4,5)
print("after adding a new key:",my_list_5)
# as we remember strings are a special datatype, list of chars
# that means strings are list like objects and iterable
# we are able to do a list call on a string too, this helps us if we want to work with strings directly for comparison a.s.o.
print("Stringtext:", my_string_variable)
print("first 10 signs:",my_string_variable[:10])
print("first 10 every second sign:", my_string_variable[:10:2])
print("string readorder from left perspective:", my_string_variable[::-1])
# as we said with sets we are able to do mathematical operation
# example to use set functions
my_list_set1 = {1,2,3,4,5,6,7,8}
my_list_set2 = {2,4,6,8,10}
print(my_list_set1.difference(my_list_set2)) # delta set set1
print(my_list_set1.intersection(my_list_set2)) # same in both
# a lot more is possible
```
## palindrom check
```
# with the knowledge of listcalls and class.function calls we are able to develop a short program
# lets develop a short palindrom check
# a palindrom is a word or sentence which is the same from both readorders (left to right == right to left - readorder)
# first we define a palindrom string
my_palin = "A man, a plan, a canal: Panama."
print("original:\n", my_palin)
# inside the string we have special characters we need to replace
# "," - comma; ":" - doubledot; "." - dot; " " - whitespace
# additional to the replacement we need to lower or uppercase all signs
my_corrected_palin = my_palin.replace(":","").replace(",","").replace(".","").replace(" ","").lower()
print("corrected:\n", my_corrected_palin)
# next we need to change the readorder
my_changed_readorder_palin = my_corrected_palin[::-1]
print("change readorder:\n", my_changed_readorder_palin)
# last we check both variables
print("is our string a palindrom?\n", my_changed_readorder_palin == my_corrected_palin)
```
## print syntax
* with the print function it is possible to print to our active terminal variables or text
* the normal structure inside a string is comma separation of variables, this have an effect that between two variable information a whitespace is automatically added
* if we using string concatination we need to ensure that all dataparts are string datatype (need to enforce it to string)
* it is also possible to multiply string with a number, if we want to print 3 times "bla" we need to program: 3*"bla"
* if we using string statements it is also possible to use excape commands like backslash n for newline as we see above
* sometimes we dont want to do the escape command or we want to fill our string statement with variables
* in past python version (< v3.7) it was possible to set placeholder inside a string statement with curly brackets and replace the placeholder with variables and format function usage
* today the format function is used if we have dynamically URL calls which are changed depending on what we want - that means we declare a general URL GET request with placeholder, and replace the placeholder with our variables, for example data from different users is cylce call with changing user information
* if want to print a string as it is - raw text, it is possible to add a small r infront of the string, this formats the string to raw text string
* since python 3.7 it is possible to format a string as a f-string which let us handle the variable replacement more easy
```
# normal string usage with comma separation
print("1234", "\n", "5678")
# a docstring is a string also
my_docstring = """this docstring test is a test
additional line
indent with pressing tabulator key on keyboard
indentation with whitespaces
this is again a new line
"""
print("formatted printout depends on the docstring: \n",my_docstring)
# string concatination with string enforcement and calculation
print("1" + "2" + str(42) * 3)
# string replacement of placeholder with format
print("on my text at position {} we have a variable {} with values".format(42, "testname") )
print("on my text at position {0} we have a variable {0} with values".format(42, "testname") )
print("on my text at position {0} we have a variable {1} with values".format(42, "testname") )
print("on my text at position {1} we have a variable {0} with values".format(42, "testname") )
print("on my text at position {1} we have a variable {1} with values".format(42, "testname") )
# string raw text
print(r"your button expect an integer value for example = {my_int} with value 42.")
print(r"with a \n escape command you add a newline to your print statement")
# f-string direct replacement of variables inside the string
my_int = 99
print(f"value of my variable is {my_int} with datatype: {type(my_int)}")
```
* congratulation you finished this lesson
## what we learned
* datatypes in python with enforcement
* special structured datatypes (lists, dictionary)
* using a class function (function calls)
* list calls
* print syntax
| github_jupyter |
# Webscrapping with Beautiful Soup.
This example shows how to download any ZIP file listed on a given webpage.
```
import requests
#import urllib
import os
from bs4 import BeautifulSoup
import pandas as pd
# Create destination
cwd = os.getcwd()
datadir = cwd +"/data"
print("The current Working directory is " + cwd)
if not os.path.exists(datadir):
os.mkdir(datadir) #, 0777);
#print "Created new directory " + newdir
#newfile = open('zipfiles.txt','w')
def scrape_find_zips(url):
""" scrape the given url for any zip files in href tags """
response = requests.get(url)
#print(response.status_code)
#print(response.text)
soup = BeautifulSoup(response.text, 'html.parser')
all_tags = soup.find_all('a', href=True)
tag_dict = {}
for tag in all_tags:
if '.tar.gz' in tag.get_text() or '.zip' in tag.get_text() or 'py2.py3-none-any.whl' in tag.get_text():
tag_dict[tag.get_text()] = tag['href']
return tag_dict
base = "http://www.nemweb.com.au"
url = "http://www.nemweb.com.au/Reports/Current/Daily_Reports/" #Did previously
url = "http://www.nemweb.com.au/Reports/ARCHIVE/Daily_Reports/"
tag_dict = scrape_find_zips(url)
print(next (iter (tag_dict.values())))
print(next (iter (tag_dict.keys())))
dest=os.getcwd()+'/data/'
def scrape_downloader(url, urls, dest='./data/'):
""" download all files listed in urls {} dict
Input:
url = base url
urls= target paths to append to base url
dest= local path to store files
"""
for k_name, v_path in urls.items():
src = url+v_path
tgt = dest+k_name
r = requests.get(src)
with open(tgt, 'wb') as code:
code.write(r.content)
print(src)
scrape_downloader(base, tag_dict, dest='./data/')
```
#Setup
Setup folder to unzip into
```
# Remember that datadir = cwd +"/data"
unzip_dir=os.getcwd()+'/data/unzip/'
if not os.path.exists(unzip_dir):
os.mkdir(unzip_dir)
print('Created',unzip_dir)
f_zip = [datadir + '/' + x for x in os.listdir('./data/')]
print(f_zip[0])
import zipfile
for f in f_zip[0:3]: #remove [0] to extend loop
print(f)
if f.endswith(".zip"):
#Unzip
zip_ref = zipfile.ZipFile(f, 'r')
zip_ref.extractall(unzip_dir)
zip_ref.close()
print(root)
print(dirs)
print(files)
f_unzip = os.listdir('./data/unzip/')
print(unzip_dir+f_unzip[0])
print(f_unzip)
```
# Process CSV file looking for specific header
```
import csv
#----------------------------------------------------------------------
def csv_reader(file_obj, segment=["I","TUNIT","","1"]):
"""
Read a csv file
scan for segment beginning with I in row n colunm 1 and TUNIT or DREGION etc in column3
e.g. segment=["I","TUNIT",None,"1"] for power generator data
e.g. segment=["I","DREGION",None,"2"] for state totals
"""
#reader = csv.DictReader(file_obj, delimiter=',', dialect=csv.unix_dialect)
reader = csv.reader(file_obj, delimiter=',', dialect=csv.unix_dialect)
lines=[]
#save contents? only when found valid header
valid=False
for row, line in enumerate(reader):
#print(row, line[0:9]) #Uncomment to debug.
#Find "I" NemWeb header rows for 'T-Units'
if line[0]==segment[0] and line[1]==segment[1] and line[3]==segment[3]:
print("Found Start at:",row, line[0:9])
valid=True
start=row
headers = line
if line[0]==segment[0] and valid and row>start: # and line[1]=="TUNIT" and line[3]=="2":
print("Found next header at",row,"exiting with",row-start,"lines")
break
if valid and not line[0]=='I':
# save non header row
lines.append(line)
return lines, headers
# create list of file paths
csv_paths = [unzip_dir+f for f in f_unzip[0:]] # 0:3 for 3 files
print(csv_paths)
#Create empty lists
newlines = lines = header = all_lines = []
for csv_path in csv_paths:
print("Processing",csv_path)
with open(csv_path, "r", newline='\n') as f_obj:
# send file object to reader and extract valid rows of data
newlines, header = csv_reader(f_obj,segment=["I","DREGION",None,"2"])
lines.extend(newlines)
#note lines.append creates a list of lists, rather than just one long list.
# Now we could create new csv, store data in hadoop, or in a dataframe
print(header)
print(lines[0])
df = pd.DataFrame(data=lines, columns=header)
# Define our index as date and time column
df.set_index("SETTLEMENTDATE", inplace=True)
df.index = pd.to_datetime(df.index)
# chart if you must?
#df.pivot(columns='DUID', values='TOTALCLEARED').astype(float).plot(title="Energy",legend=False)
from IPython.display import display
display(df.iloc[0:11,[0,5,6,7,11,12]])
#df.dtypes
#Save to Hadoop File for fast loading later
df.to_hdf('AllRows.hdf5',key='NRG')
nsw = df.loc[df.loc[:,"REGIONID"]=="NSW1",:]
display(nsw["TOTALDEMAND"].astype(float))
daily = nsw["TOTALDEMAND"].astype(float).fillna(0).resample(rule="D").sum().dropna()
daily.head(20)
# Alternatively, for simple csv file formats with only 1 header row we can add all directory contents to a single dataframe
# Combine data into dataframe
def df_from_many(dir_path, file_type=".CSV", date_col='DATE'):
#Setup target pandas dataframe
df = pd.DataFrame()
#Loop over cwd/data zip files
print(os.getcwd())
for root, dirs, files in os.walk(dir_path):
#print(root,dirs,files)
for file in files:
if file.endswith(file_type):
print(file)
tempdf = pd.read_csv(dir_path+file, skiprows=1)
tempdf.index = pd.to_datetime(tempdf.loc[date_col,:])
df = pd.concat([df, tempdf], axis=0)
```
| github_jupyter |
```
import scipy
from scipy.io import arff
import pandas as pd
import numpy as np
from sklearn.metrics import accuracy_score
data, meta = scipy.io.arff.loadarff('yeast/yeast-train.arff')
df = pd.DataFrame(data)
meta
df.head()
df.dtypes
len(df.columns)
labels_df = df.ix[:,103:117]
labels_df.head()
for i in range(1,15):
class_name = 'Class'+str(i)
labels_df[class_name] = labels_df[class_name].astype('float64')
labels_df.dtypes
# Here, I'm trying to check the correlations between labels
labels_df.apply(lambda s: labels_df.corrwith(s))
# you can also generate a random multi-label dataset on your own
from sklearn.datasets import make_multilabel_classification
X, y = make_multilabel_classification(sparse = True, n_labels = 7,
return_indicator = 'sparse', allow_unlabeled = False)
matrix_df = pd.SparseDataFrame([ pd.SparseSeries(X[i].toarray().ravel())
for i in np.arange(X.shape[0])])
matrix_df.head()
# Start Multi-Label Experiments From HERE
training_data, train_meta = scipy.io.arff.loadarff('yeast/yeast-train.arff')
train_df = pd.DataFrame(training_data)
testing_data, test_meta = scipy.io.arff.loadarff('yeast/yeast-test.arff')
test_df = pd.DataFrame(testing_data)
# sparse: If True, returns a sparse matrix, where sparse matrix means a matrix having a large number of zero elements.
# n_labels: The average number of labels for each instance.
# return_indicator: If ‘sparse’ return Y in the sparse binary indicator format.
# allow_unlabeled: If True, some instances might not belong to any class.
train_df.head()
test_df.head()
X_train = train_df.iloc[:,:103]
Y_train = train_df.iloc[:,103:]
X_test = test_df.iloc[:,:103]
Y_test = test_df.iloc[:,103:]
for i in range(1,15):
class_name = 'Class'+str(i)
Y_train[class_name] = Y_train[class_name].astype('float64')
Y_train.dtypes
for i in range(1,15):
class_name = 'Class'+str(i)
Y_test[class_name] = Y_test[class_name].astype('float64')
Y_test.dtypes
X_test.head()
Y_test.head()
# Method 1 - Problem Transformation - Binary Relavance
## Simply treat each label independently
from skmultilearn.problem_transform import BinaryRelevance
from sklearn.naive_bayes import GaussianNB
classifier = BinaryRelevance(GaussianNB())
# train
classifier.fit(X_train, Y_train)
# predict
predictions = classifier.predict(X_test)
accuracy_score(Y_test,predictions) # 0.10359869138495092, very low accuracy
# Method 1 - Problem Transformation - Classifier Chains
## Each round predict 1 label column, and put it in feature columns for the next round of label prediction
## For this method, in order to achive higher accuracy, better to have higher corrlation between labels
## So try the labels correlation check as I did here
Y_train.apply(lambda s: Y_train.corrwith(s))
Y_test.apply(lambda s: Y_test.corrwith(s))
## Not very high correlation, let's check the accuracy from Classifier Chains
from skmultilearn.problem_transform import ClassifierChain
from sklearn.naive_bayes import GaussianNB
classifier = ClassifierChain(GaussianNB())
# train
classifier.fit(X_train, Y_train)
# predict
predictions = classifier.predict(X_test)
accuracy_score(Y_test,predictions) # 0.092693565976008724, even lower...
# Method 1 - Problem Transformation - Label Powerset
## Group rows with exactly same label value set (such as 2 rows all have labels (0,1,0,1)), together
## Then give each label set a value, so that it will become 1-label prediction problem
## But, this method may suffer from data imbalance when it comes to real world dataset
from skmultilearn.problem_transform import LabelPowerset
from sklearn.naive_bayes import GaussianNB
classifier = LabelPowerset(GaussianNB())
# train
classifier.fit(X_train, Y_train)
# predict
predictions = classifier.predict(X_test)
accuracy_score(Y_test,predictions) # 0.18647764449291168, a little higher...
# Method 2 - Adapt Algorithms
## Scikit-Learn package: http://scikit.ml/api/api/skmultilearn.adapt.html#module-skmultilearn.adapt
from skmultilearn.adapt import MLkNN
# Scikit-learn Adapt Algorithms requires Dense/Sparse matrix as data input
X_train_matrix = scipy.sparse.csr_matrix(X_train.values)
Y_train_matrix = scipy.sparse.csr_matrix(Y_train.values)
X_test_matrix = scipy.sparse.csr_matrix(X_test.values)
Y_test_matrix = scipy.sparse.csr_matrix(Y_test.values)
classifier = MLkNN(k=10)
# train
classifier.fit(X_train_matrix, Y_train_matrix)
# predict
predictions = classifier.predict(X_test_matrix)
accuracy_score(Y_test_matrix,predictions) # 0.16684841875681569
# increase k
classifier = MLkNN(k=20)
# train
classifier.fit(X_train_matrix, Y_train_matrix)
# predict
predictions = classifier.predict(X_test_matrix)
accuracy_score(Y_test_matrix,predictions) # 0.18102508178844057
from skmultilearn.adapt import BRkNNaClassifier
classifier = BRkNNaClassifier(k=10)
# train
classifier.fit(X_train_matrix, Y_train_matrix)
# predict
predictions = classifier.predict(X_test_matrix)
accuracy_score(Y_test_matrix,predictions) # 0.10032715376226826
# Method 3 - Ensembling Multi-Label
## Scikit-Learn emsembling: http://scikit.ml/api/classify.html#ensemble-approaches
## tools to install in order to get "graph_tool.all": https://gist.github.com/v-pravin/949fc18d58a560cf85d2
from sklearn.ensemble import RandomForestClassifier
from skmultilearn.problem_transform import LabelPowerset
from skmultilearn.cluster import IGraphLabelCooccurenceClusterer
from skmultilearn.ensemble import LabelSpacePartitioningClassifier
# construct base forest classifier
base_classifier = RandomForestClassifier()
# setup problem transformation approach with sparse matrices for random forest
problem_transform_classifier = LabelPowerset(classifier=base_classifier,
require_dense=[False, False])
# partition the label space using fastgreedy community detection
# on a weighted label co-occurrence graph with self-loops allowed
clusterer = IGraphLabelCooccurenceClusterer('fastgreedy', weighted=True,
include_self_edges=True)
# setup the ensemble metaclassifier
classifier = LabelSpacePartitioningClassifier(problem_transform_classifier, clusterer)
# train
classifier.fit(X_train_matrix, Y_train_matrix)
# predict
predictions = classifier.predict(X_test_matrix)
accuracy_score(Y_test_matrix,predictions)
```
| github_jupyter |
# Introduction to Neural Networks
In this notebook you will learn how to create and use a neural network to classify articles of clothing. To achieve this, we will use a sub module of TensorFlow called *keras*.
*This guide is based on the following TensorFlow documentation.*
https://www.tensorflow.org/tutorials/keras/classification
## Keras
Before we dive in and start discussing neural networks, I'd like to give a breif introduction to keras.
From the keras official documentation (https://keras.io/) keras is described as follows.
"Keras is a high-level neural networks API, written in Python and capable of running on top of TensorFlow, CNTK, or Theano. It was developed with a focus on enabling fast experimentation.
Use Keras if you need a deep learning library that:
- Allows for easy and fast prototyping (through user friendliness, modularity, and extensibility).
- Supports both convolutional networks and recurrent networks, as well as combinations of the two.
- Runs seamlessly on CPU and GPU."
Keras is a very powerful module that allows us to avoid having to build neural networks from scratch. It also hides a lot of mathematical complexity (that otherwise we would have to implement) inside of helpful packages, modules and methods.
In this guide we will use keras to quickly develop neural networks.
## What is a Neural Network
So, what are these magical things that have been beating chess grandmasters, driving cars, detecting cancer cells and winning video games?
A deep neural network is a layered representation of data. The term "deep" refers to the presence of multiple layers. Recall that in our core learning algorithms (like linear regression) data was not transformed or modified within the model, it simply existed in one layer. We passed some features to our model, some math was done, an answer was returned. The data was not changed or transformed throughout this process. A neural network processes our data differently. It attempts to represent our data in different ways and in different dimensions by applying specific operations to transform our data at each layer. Another way to express this is that at each layer our data is transformed in order to learn more about it. By performing these transformations, the model can better understand our data and therefore provide a better prediction.
## How it Works
Before going into too much detail I will provide a very surface level explination of how neural networks work on a mathematical level. All the terms and concepts I discuss will be defined and explained in more detail below.
On a lower level neural networks are simply a combination of elementry math operations and some more advanced linear algebra. Each neural network consists of a sequence of layers in which data passes through. These layers are made up on neurons and the neurons of one layer are connected to the next (see below). These connections are defined by what we call a weight (some numeric value). Each layer also has something called a bias, this is simply an extra neuron that has no connections and holds a single numeric value. Data starts at the input layer and is trasnformed as it passes through subsequent layers. The data at each subsequent neuron is defined as the following.
> $Y =(\sum_{i=0}^n w_i x_i) + b$
> $w$ stands for the weight of each connection to the neuron
> $x$ stands for the value of the connected neuron from the previous value
> $b$ stands for the bias at each layer, this is a constant
> $n$ is the number of connections
> $Y$ is the output of the current neuron
> $\sum$ stands for sum
The equation you just read is called a weighed sum. We will take this weighted sum at each and every neuron as we pass information through the network. Then we will add what's called a bias to this sum. The bias allows us to shift the network up or down by a constant value. It is like the y-intercept of a line.
But that equation is the not complete one! We forgot a crucial part, **the activation function**. This is a function that we apply to the equation seen above to add complexity and dimensionality to our network. Our new equation with the addition of an activation function $F(x)$ is seen below.
> $Y =F((\sum_{i=0}^n w_i x_i) + b)$
Our network will start with predefined activation functions (they may be different at each layer) but random weights and biases. As we train the network by feeding it data it will learn the correct weights and biases and adjust the network accordingly using a technqiue called **backpropagation** (explained below). Once the correct weights and biases have been learned our network will hopefully be able to give us meaningful predictions. We get these predictions by observing the values at our final layer, the output layer.
## Breaking Down The Neural Network!
Before we dive into any code lets break down how a neural network works and what it does.

*Figure 1*
### Data
The type of data a neural network processes varies drastically based on the problem being solved. When we build a neural network, we define what shape and kind of data it can accept. It may sometimes be neccessary to modify our dataset so that it can be passed to our neural network.
Some common types of data a neural network uses are listed below.
- Vector Data (2D)
- Timeseries or Sequence (3D)
- Image Data (4D)
- Video Data (5D)
There are of course many different types or data, but these are the main categories.
### Layers
As we mentioned earlier each neural network consists of multiple layers. At each layer a different transformation of data occurs. Our initial input data is fed through the layers and eventually arrives at the output layer where we will obtain the result.
#### Input Layer
The input layer is the layer that our initial data is passed to. It is the first layer in our neural network.
#### Output Layer
The output layer is the layer that we will retrive our results from. Once the data has passed through all other layers it will arrive here.
#### Hidden Layer(s)
All the other layers in our neural network are called "hidden layers". This is because they are hidden to us, we cannot observe them. Most neural networks consist of at least one hidden layer but can have an unlimited amount. Typically, the more complex the model the more hidden layers.
#### Neurons
Each layer is made up of what are called neurons. Neurons have a few different properties that we will discuss later. The important aspect to understand now is that each neuron is responsible for generating/holding/passing ONE numeric value.
This means that in the case of our input layer it will have as many neurons as we have input information. For example, say we want to pass an image that is 28x28 pixels, thats 784 pixels. We would need 784 neurons in our input layer to capture each of these pixels.
This also means that our output layer will have as many neurons as we have output information. The output is a little more complicated to understand so I'll refrain from an example right now but hopefully you're getting the idea.
But what about our hidden layers? Well these have as many neurons as we decide. We'll discuss how we can pick these values later but understand a hidden layer can have any number of neurons.
####Connected Layers
So how are all these layers connected? Well the neurons in one layer will be connected to neurons in the subsequent layer. However, the neurons can be connected in a variety of different ways.
Take for example *Figure 1* (look above). Each neuron in one layer is connected to every neuron in the next layer. This is called a **dense** layer. There are many other ways of connecting layers but well discuss those as we see them.
### Weights
Weights are associated with each connection in our neural network. Every pair of connected nodes will have one weight that denotes the strength of the connection between them. These are vital to the inner workings of a neural network and will be tweaked as the neural network is trained. The model will try to determine what these weights should be to achieve the best result. Weights start out at a constant or random value and will change as the network sees training data.
### Biases
Biases are another important part of neural networks and will also be tweaked as the model is trained. A bias is simply a constant value associated with each layer. It can be thought of as an extra neuron that has no connections. The purpose of a bias is to shift an entire activation function by a constant value. This allows a lot more flexibllity when it comes to choosing an activation and training the network. There is one bias for each layer.
### Activation Function
Activation functions are simply a function that is applied to the weighed sum of a neuron. They can be anything we want but are typically higher order/degree functions that aim to add a higher dimension to our data. We would want to do this to introduce more comolexity to our model. By transforming our data to a higher dimension, we can typically make better, more complex predictions.
A list of some common activation functions and their graphs can be seen below.
- Relu (Rectified Linear Unit)

- Tanh (Hyperbolic Tangent)
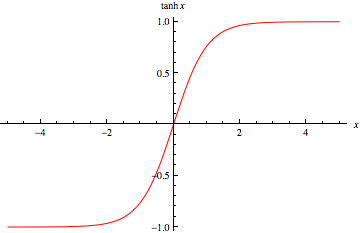
- Sigmoid

### Backpropagation
Backpropagation is the fundemental algorithm behind training neural networks. It is what changes the weights and biases of our network. To fully explain this process, we need to start by discussing something called a cost/loss function.
#### Loss/Cost Function
As we now know our neural network feeds information through the layers until it eventually reaches an output layer. This layer contains the results that we look at to determine the prediciton from our network. In the training phase it is likely that our network will make many mistakes and poor predicitions. In fact, at the start of training our network doesn't know anything (it has random weights and biases)!
We need some way of evaluating if the network is doing well and how well it is doing. For our training data we have the features (input) and the labels (expected output), because of this we can compare the output from our network to the expected output. Based on the difference between these values we can determine if our network has done a good job or poor job. If the network has done a good job, we'll make minor changes to the weights and biases. If it has done a poor job our changes may be more drastic.
So, this is where the cost/loss function comes in. This function is responsible for determining how well the network did. We pass it the output and the expected output, and it returns to us some value representing the cost/loss of the network. This effectively makes the networks job to optimize this cost function, trying to make it as low as possible.
Some common loss/cost functions include.
- Mean Squared Error
- Mean Absolute Error
- Hinge Loss
#### Gradient Descent
Gradient descent and backpropagation are closely related. Gradient descent is the algorithm used to find the optimal paramaters (weights and biases) for our network, while backpropagation is the process of calculating the gradient that is used in the gradient descent step.
Gradient descent requires some pretty advanced calculus and linear algebra to understand so we'll stay away from that for now. Let's just read the formal definition for now.
"Gradient descent is an optimization algorithm used to minimize some function by iteratively moving in the direction of steepest descent as defined by the negative of the gradient. In machine learning, we use gradient descent to update the parameters of our model." (https://ml-cheatsheet.readthedocs.io/en/latest/gradient_descent.html)
And that's all we really need to know for now. I'll direct you to the video for a more in depth explination.

### Optimizer
You may sometimes see the term optimizer or optimization function. This is simply the function that implements the backpropagation algorithm described above. Here's a list of a few common ones.
- Gradient Descent
- Stochastic Gradient Descent
- Mini-Batch Gradient Descent
- Momentum
- Nesterov Accelerated Gradient
- Adam
*This article explains them quite well is where I've pulled this list from.* (https://medium.com/@sdoshi579/optimizers-for-training-neural-network-59450d71caf6)
https://www.tensorflow.org/api_docs/python/tf/keras/optimizers
# Creating a Neural Network
Okay now you have reached the exciting part of this tutorial! No more math and complex explinations. Time to get hands on and train a very basic neural network.
*As stated earlier this guide is based off of the following TensorFlow tutorial.*
https://www.tensorflow.org/tutorials/keras/classification
### Imports
```
# TensorFlow and tf.keras
import tensorflow as tf
# Helper libraries
import numpy as np
import matplotlib.pyplot as plt
```
### Dataset
For this tutorial we will use the MNIST Fashion Dataset. This is a dataset that is included in keras.
This dataset includes 60,000 images for training and 10,000 images for validation/testing.
```
fashion_mnist = tf.keras.datasets.fashion_mnist # load dataset
(train_images, train_labels), (test_images, test_labels) = fashion_mnist.load_data() # split into tetsing and training
```
Let's have a look at this data to see what we are working with.
```
train_images.shape
```
So we've got 60,000 images that are made up of 28x28 pixels (784 in total).
```
train_images[0,23,23] # let's have a look at one pixel
```
Our pixel values are between 0 and 255, 0 being black and 255 being white. This means we have a grayscale image as there are no color channels.
```
train_labels[:10] # let's have a look at the first 10 training labels
```
Our labels are integers ranging from 0 - 9. Each integer represents a specific article of clothing. We'll create an array of label names to indicate which is which.
```
class_names = ['T-shirt/top', 'Trouser', 'Pullover', 'Dress', 'Coat',
'Sandal', 'Shirt', 'Sneaker', 'Bag', 'Ankle boot']
```
Fianlly let's look at what some of these images look like!
```
plt.figure()
plt.imshow(train_images[1])
plt.colorbar()
plt.grid(False)
plt.show()
```
## Data Preprocessing
The last step before creating our model is to *preprocess* our data. This simply means applying some prior transformations to our data before feeding it the model. In this case we will simply scale all our greyscale pixel values (0-255) to be between 0 and 1. We can do this by dividing each value in the training and testing sets by 255.0. We do this because smaller values will make it easier for the model to process our values.
```
train_images = train_images / 255.0
test_images = test_images / 255.0
```
## Building the Model
Now it's time to build the model! We are going to use a keras *sequential* model with three different layers. This model represents a feed-forward neural network (one that passes values from left to right). We'll break down each layer and its architecture below.
```
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Flatten, Dense
model = Sequential()
model.add(Flatten(input_shape=(28, 28), name="flatten_1"))
model.add(Dense(128, activation='relu', name="dense_1"))
model.add(Dense(10, activation='softmax', name="dense_output"))
model.compile(
optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy']
)
model.summary()
```
**Layer 1:** This is our input layer and it will conist of 784 neurons. We use the flatten layer with an input shape of (28, 28) to denote that our input should come in in that shape. The flatten means that our layer will reshape the shape (28, 28) array into a vector of 784 neurons so that each pixel will be associated with one neuron.
**Layer 2:** This is our first and only hidden layer. The *dense* denotes that this layer will be fully connected and each neuron from the previous layer connects to each neuron of this layer. It has 128 neurons and uses the rectify linear unit activation function.
**Layer 3:** This is our output later and is also a dense layer. It has 10 neurons that we will look at to determine our models output. Each neuron represnts the probabillity of a given image being one of the 10 different classes. The activation function *softmax* is used on this layer to calculate a probabillity distribution for each class. This means the value of any neuron in this layer will be between 0 and 1, where 1 represents a high probabillity of the image being that class.
### Compile the Model
The last step in building the model is to define the loss function, optimizer and metrics we would like to track. I won't go into detail about why we chose each of these right now.
## Training the Model
Now it's finally time to train the model. Since we've already done all the work on our data this step is as easy as calling a single method.
```
epochs = 10
validation_split = 0.10
history = model.fit(train_images, train_labels,
epochs=epochs,
validation_split=validation_split) # we pass the data, labels and epochs and watch the magic!
```
## Evaluating the Model
Now it's time to test/evaluate the model. We can do this quite easily using another builtin method from keras.
The *verbose* argument is defined from the keras documentation as:
"verbose: 0 or 1. Verbosity mode. 0 = silent, 1 = progress bar."
(https://keras.io/models/sequential/)
```
test_loss, test_acc = model.evaluate(test_images, test_labels, verbose=1)
print('Test accuracy:', test_acc)
```
You'll likely notice that the accuracy here is lower than when training the model. This difference is reffered to as **overfitting**.
And now we have a trained model that's ready to use to predict some values!
## Making Predictions
To make predictions we simply need to pass an array of data in the form we've specified in the input layer to ```.predict()``` method.
```
predictions = model.predict(test_images)
```
This method returns to us an array of predictions for each image we passed it. Let's have a look at the predictions for image 1.
```
predictions[0]
```
If we wan't to get the value with the highest score we can use a useful function from numpy called ```argmax()```. This simply returns the index of the maximium value from a numpy array.
```
np.argmax(predictions[0])
```
And we can check if this is correct by looking at the value of the cooresponding test label.
```
test_labels[0]
```
## Verifying Predictions
I've written a small function here to help us verify predictions with some simple visuals.
```
COLOR = 'white'
plt.rcParams['text.color'] = COLOR
plt.rcParams['axes.labelcolor'] = COLOR
def predict(model, image, correct_label):
class_names = ['T-shirt/top', 'Trouser', 'Pullover', 'Dress', 'Coat',
'Sandal', 'Shirt', 'Sneaker', 'Bag', 'Ankle boot']
prediction = model.predict(np.array([image]))
predicted_class = class_names[np.argmax(prediction)]
show_image(image, class_names[correct_label], predicted_class, np.max(prediction)*100)
def show_image(img, label, guess, accuracy):
print("Expected: " + label)
plt.figure()
plt.imshow(img, cmap=plt.cm.binary)
plt.colorbar()
plt.grid(False)
plt.show()
print(f'Guess: {guess} - Accuracy: {accuracy:.1f}%')
def get_number():
while True:
num = input("Pick a number: ")
if num.isdigit():
num = int(num)
if 0 <= num <= 1000:
return int(num)
else:
print("Try again...")
num = get_number()
image = test_images[num]
label = test_labels[num]
predict(model, image, label)
```
And that's pretty much it for an introduction to neural networks!
## Sources
1. Doshi, Sanket. “Various Optimization Algorithms For Training Neural Network.” Medium, Medium, 10 Mar. 2019, www.medium.com/@sdoshi579/optimizers-for-training-neural-network-59450d71caf6.
2. “Basic Classification: Classify Images of Clothing : TensorFlow Core.” TensorFlow, www.tensorflow.org/tutorials/keras/classification.
3. “Gradient Descent¶.” Gradient Descent - ML Glossary Documentation, www.ml-cheatsheet.readthedocs.io/en/latest/gradient_descent.html.
4. Chollet François. Deep Learning with Python. Manning Publications Co., 2018.
5. “Keras: The Python Deep Learning Library.” Home - Keras Documentation, www.keras.io/.
| github_jupyter |
# How to Solve Python Coding Questions using Math
[Source](https://towardsdatascience.com/how-to-solve-python-coding-questions-using-math-72d5540b5a24)
- Author: Israel Oliveira [\[e-mail\]](mailto:'Israel%20Oliveira%20'<prof.israel@gmail.com>)
```
%load_ext watermark
%config Completer.use_jedi = False
import pandas as pd
from random import uniform
import math
#import matplotlib.pyplot as plt
#%matplotlib inline
#from IPython.core.pylabtools import figsize
#figsize(12, 8)
#import seaborn as sns
#sns.set_theme()
#pd.set_option("max_columns", None)
#pd.set_option("max_rows", None)
#from IPython.display import Markdown, display
#def md(arg):
# display(Markdown(arg))
#from pandas_profiling import ProfileReport
# report = ProfileReport(#DataFrame here#, minimal=True)
# report.to
#import pyarrow.parquet as pq
# df = pq.ParquetDataset(path_to_folder_with_parquets, filesystem=None).read_pandas().to_pandas()
# Run this cell before close.
%watermark -d --iversion -b -r -g -m -v
!cat /proc/cpuinfo |grep 'model name'|head -n 1 |sed -e 's/model\ name/CPU/'
!free -h |cut -d'i' -f1 |grep -v total
```
# Solution 1: for loop
```
def countOdds(low,high):
count = 0
for i in range(low,high+1):
if i%2 != 0:
count+=1
return count
#test case
countOdds(3,7)
```
# Solution 2: math
```
# calculate the number of odd numbers between 1 and low-1: low//2
# calculate the number of odd numbers between 1 and high: (high+1)//2
# the difference is our result
def countOdds_2(low, high) -> int:
return (high + 1) // 2 - low // 2
#test case
countOdds_2(3,7)
countOdds(3,70000000)
countOdds_2(3,70000000)
```
# Question 2: Arranging Coins, by Bloomberg
Final solution:
```
def arranging(n):
left, right = 0, n
while left<=right:
k = (left+right)//2
current = k*(k+1)//2
if current == n:
return k
elif current >n:
right = k-1
else:
left = k+1
return right
# test case
n= 8
arranging(n)
```
Solution a:
```
def arranging_a(n):
for k in range(1,n//2+2):
current = k*(k+1)//2
if current == n:
return k
elif current >n:
return k-1
return False
# test case
n = 8
arranging_a(n)
```
Validating:
```
s = 0
for n in range(1,100):
s += abs(arranging_a(n)-arranging(n))
s == 0
```
Timing:
```
%%timeit
n = int(uniform(1e7, 1e8))
arranging(n)
%%timeit
n = int(uniform(1e7, 1e8))
arranging_a(n)
```
Solution b
$$
n = \frac{k(k+1)}{2} + \epsilon
$$
$$
8 = \frac{3(3+1)}{2} + \epsilon = 6 + \epsilon, ~~ k = 3
$$
$$
\epsilon \in [0, k+1)
$$
$$
n = f(k) + \epsilon
$$
$$
f(k) = \frac{k(k+1)}{2} = \frac{k^2+k}{2}
$$
$$
k^2+k = 2n
$$
$$
k+1 = \frac{2n}{k}
$$
$$
n = \frac{k^2}{2} + \frac{k}{2} \approx \frac{k^2}{2}, ~~\text{if}~~ n >> k
$$
$$
n > \frac{k^2}{2} ~~\therefore~~ k^2 < 2n
$$
$$
k^2+k = 2n+k > 2n ~~\therefore~~n+\frac{k}{2} > n
$$
$$
n > \frac{k^2}{2} ~~\therefore~~ \sqrt{2n} > k
$$
$$
\sqrt{2n} > k ~~\therefore~~ \sqrt{2(8)} = 4 > 3
$$
$$
k_0 = \lfloor \sqrt{2n} \rfloor \\
k_i = k_{i-1} -1
$$
```
def arranging_b(n):
k = int((2*n)**(0.5))
while k:
current = k*(k+1)//2
if (current == n) or ((current < n) and (n-current < k+1)):
return k
k -= 1
return False
# test case
n = 13
arranging_b(n)
n = 2
k = int(math.sqrt(2*n))
current = k*(k+1)//2
current
s = 0
for n in range(1,100):
d = (arranging_b(n)-arranging(n))
if abs(d) > 0:
print(f'{n}: {d}')
s += d
s == 0
```
Timing
```
%%timeit
n = int(uniform(1e7, 1e8))
arranging(n)
%%timeit
n = int(uniform(1e7, 1e8))
arranging_b(n)
```
# Question 4: Set Mismatch, by Amazon
```
def findErrorNums(nums):
return [sum(nums)-sum(set(nums)),sum(range(1,len(nums)+1)) - sum(set(nums))]# twice & missing value
nums = [1,2,3,4,5,6,7,7,9,10]
findErrorNums(nums)
2**(0.5)
```
| github_jupyter |
# Training the DataSet
The dataset we are using for Emotion Detection from Facial Expressions is "FER2013"
```
import pandas as pd
import cv2
import numpy as np
dataset_path = '/fer2013/fer2013.csv'
image_size=(48,48)
def load_fer2013():
data = pd.read_csv(dataset_path)
pixels = data['pixels'].tolist()
width, height = 48, 48
faces = []
for pixel_sequence in pixels:
face = [int(pixel) for pixel in pixel_sequence.split(' ')]
face = np.asarray(face).reshape(width, height)
face = cv2.resize(face.astype('uint8'),image_size)
faces.append(face.astype('float32'))
faces = np.asarray(faces)
faces = np.expand_dims(faces, -1)
emotions = pd.get_dummies(data['emotion']).as_matrix()
return faces, emotions
def preprocess_input(x, v2=True):
x = x.astype('float32')
x = x / 255.0
if v2:
x = x - 0.5
x = x * 2.0
return x
from keras.callbacks import CSVLogger, ModelCheckpoint, EarlyStopping
from keras.callbacks import ReduceLROnPlateau
from keras.preprocessing.image import ImageDataGenerator
from load_and_process import load_fer2013
from load_and_process import preprocess_input
from models.cnn import mini_XCEPTION
from sklearn.model_selection import train_test_split
# parameters
batch_size = 32
num_epochs = 10000
input_shape = (48, 48, 1)
validation_split = .2
verbose = 1
num_classes = 7
patience = 50
base_path = 'models/'
# data generator
data_generator = ImageDataGenerator(
featurewise_center=False,
featurewise_std_normalization=False,
rotation_range=10,
width_shift_range=0.1,
height_shift_range=0.1,
zoom_range=.1,
horizontal_flip=True)
# model parameters/compilation
model = mini_XCEPTION(input_shape, num_classes)
model.compile(optimizer='adam', loss='categorical_crossentropy',
metrics=['accuracy'])
model.summary()
# callbacks
log_file_path = base_path + '_emotion_training.log'
csv_logger = CSVLogger(log_file_path, append=False)
early_stop = EarlyStopping('val_loss', patience=patience)
reduce_lr = ReduceLROnPlateau('val_loss', factor=0.1,
patience=int(patience/4), verbose=1)
trained_models_path = base_path + '_mini_XCEPTION'
model_names = trained_models_path + '.{epoch:02d}-{val_acc:.2f}.hdf5'
model_checkpoint = ModelCheckpoint(model_names, 'val_loss', verbose=1,
save_best_only=True)
callbacks = [model_checkpoint, csv_logger, early_stop, reduce_lr]
# loading dataset
faces, emotions = load_fer2013()
faces = preprocess_input(faces)
num_samples, num_classes = emotions.shape
xtrain, xtest,ytrain,ytest = train_test_split(faces, emotions,test_size=0.2,shuffle=True)
model.fit_generator(data_generator.flow(xtrain, ytrain,
batch_size),
steps_per_epoch=len(xtrain) / batch_size,
epochs=num_epochs, verbose=1, callbacks=callbacks,
validation_data=(xtest,ytest))
```
| github_jupyter |
# Linear Algebra with examples using Numpy
```
import numpy as np
from numpy.random import randn as randn
from numpy.random import randint as randint
```
## Linear Algebra and Machine Learning
* Ranking web pages in order of importance
* Solved as the problem of finding the eigenvector of the page score matrix
* Dimensionality reduction - Principal Component Analysis
* Movie recommendation
* Use singular value decomposition (SVD) to break down user-movie into user-feature and movie-feature matrices, keeping only the top $k$-ranks to identify the best matches
* Topic modeling
* Extensive use of SVD and matrix factorization can be found in Natural Language Processing, specifically in topic modeling and semantic analysis
## Vectors
A vector can be represented by an array of real numbers
$$\mathbf{x} = [x_1, x_2, \ldots, x_n]$$
Geometrically, a vector specifies the coordinates of the tip of the vector if the tail were placed at the origin
```
from IPython.display import Image
Image('vector.png')
print x
print x.shape
```
The norm of a vector $\mathbf{x}$ is defined by
$$||\boldsymbol{x}|| = \sqrt{x_1^2 + x_2^2 + \cdots + x_n^2}$$
```
print(np.sqrt(np.sum(x**2)))
print(np.linalg.norm(x))
```
Adding a constant to a vector adds the constant to each element
$$a + \boldsymbol{x} = [a + x_1, a + x_2, \ldots, a + x_n]$$
```
a = 4
print(x)
print(x + 4)
```
Multiplying a vector by a constant multiplies each term by the constant.
$$a \boldsymbol{x} = [ax_1, ax_2, \ldots, ax_n]$$
```
print x
print x*4
print np.linalg.norm(x*4)
#print np.linalg.norm(x)
```
If we have two vectors $\boldsymbol{x}$ and $\boldsymbol{y}$ of the same length $(n)$, then the _dot product_ is give by
$$\boldsymbol{x} \cdot \boldsymbol{y} = x_1y_1 + x_2y_2 + \cdots + x_ny_n$$
```
y = np.array([4, 3, 2, 1])
print x
print y
np.dot(x,y)
```
If $\mathbf{x} \cdot \mathbf{y} = 0$ then $x$ and $y$ are *orthogonal* (aligns with the intuitive notion of perpindicular)
```
w = np.array([1, 2])
v = np.array([-2, 1])
np.dot(w,v)
```
The norm squared of a vector is just the vector dot product with itself
$$
||x||^2 = x \cdot x
$$
```
print(x)
print np.linalg.norm(x)**2
print np.dot(x,x)
```
The distance between two vectors is the norm of the difference.
$$
d(x,y) = ||x-y||
$$
```
np.linalg.norm(x-y)
```
_Cosine Similarity_ is the cosine of the angle between the two vectors give by
$$cos(\theta) = \frac{\boldsymbol{x} \cdot \boldsymbol{y}}{||\boldsymbol{x}|| \text{ } ||\boldsymbol{y}||}$$
```
x = np.array([1,2,3,4])
y = np.array([5,6,7,8])
np.dot(x,y)/(np.linalg.norm(x)*np.linalg.norm(y))
```
If both $\boldsymbol{x}$ and $\boldsymbol{y}$ are zero-centered, this calculation is the _correlation_ between $\boldsymbol{x}$ and $\boldsymbol{y}$
```
from scipy.stats import pearsonr
x_centered = x - np.mean(x)
y_centered = y - np.mean(y)
# The following gives the "Centered Cosine Similarity"
# ... which is equivelent to the "Sample Pearson Correlation Coefficient"
# ... (in the correlation case, we're interpreting the vector as a list of samples)
# ... see: https://en.wikipedia.org/wiki/Pearson_product-moment_correlation_coefficient#For_a_sample
r1 = np.dot(x_centered,y_centered)/(np.linalg.norm(x_centered)*np.linalg.norm(y_centered))
r2 = pearsonr(x_centered,y_centered)
print(r1,r2[0])
```
### Linear Combinations of Vectors
If we have two vectors $\boldsymbol{x}$ and $\boldsymbol{y}$ of the same length $(n)$, then
$$\boldsymbol{x} + \boldsymbol{y} = [x_1+y_1, x_2+y_2, \ldots, x_n+y_n]$$
```
x = np.array([1,2,3,4])
y = np.array([5,6,7,8])
print x+y
a=2
x = np.array([1,2,3,4])
print a*x
```
A _linear combination_ of a collection of vectors $(\boldsymbol{x}_1,
\boldsymbol{x}_2, \ldots,
\boldsymbol{x}_m)$
is a vector of the form
$$a_1 \cdot \boldsymbol{x}_1 + a_2 \cdot \boldsymbol{x}_2 +
\cdots + a_m \cdot \boldsymbol{x}_m$$
```
a1=2
x1 = np.array([1,2,3,4])
print a1*x1
a2=4
x2 = np.array([5,6,7,8])
print a2*x2
print a1*x1 + a2*x2
```
# Matrices
An $n \times p$ matrix is an array of numbers with $n$ rows and $p$ columns:
$$
X =
\begin{bmatrix}
x_{11} & x_{12} & \cdots & x_{1p} \\
x_{21} & x_{22} & \cdots & x_{2p} \\
\vdots & \vdots & \ddots & \vdots \\
x_{n1} & x_{n2} & \cdots & x_{np}
\end{bmatrix}
$$
$n$ = the number of subjects
$p$ = the number of features
For the following $2 \times 3$ matrix
$$
X =
\begin{bmatrix}
1 & 2 & 3\\
4 & 5 & 6
\end{bmatrix}
$$
We can create in Python using NumPY
```
X = np.array([[1,2,3],[4,5,6]])
print X[1, 2]
print X
print X.shape
```
### Basic Properties
Let $X$ and $Y$ be matrices **of the dimension $n \times p$**. Let $x_{ij}$ $y_{ij}$ for $i=1,2,\ldots,n$ and $j=1,2,\ldots,p$ denote the entries in these matrices, then
1. $X+Y$ is the matrix whose $(i,j)^{th}$ entry is $x_{ij} + y_{ij}$
2. $X-Y$ is the matrix whose $(i,j)^{th}$ entry is $x_{ij} - y_{ij}$
3. $aX$, where $a$ is any real number, is the matrix whose $(i,j)^{th}$ entry is $ax_{ij}$
```
X = np.array([[1,2,3],[4,5,6]])
print X
Y = np.array([[7,8,9],[10,11,12]])
print Y
print X+Y
X = np.array([[1,2,3],[4,5,6]])
print X
Y = np.array([[7,8,9],[10,11,12]])
print Y
print X-Y
X = np.array([[1,2,3],[4,5,6]])
print X
a=5
print a*X
```
In order to multiply two matrices, they must be _conformable_ such that the number of columns of the first matrix must be the same as the number of rows of the second matrix.
Let $X$ be a matrix of dimension $n \times k$ and let $Y$ be a matrix of dimension $k \times p$, then the product $XY$ will be a matrix of dimension $n \times p$ whose $(i,j)^{th}$ element is given by the dot product of the $i^{th}$ row of $X$ and the $j^{th}$ column of $Y$
$$\sum_{s=1}^k x_{is}y_{sj} = x_{i1}y_{1j} + \cdots + x_{ik}y_{kj}$$
### Note:
$$XY \neq YX$$
If $X$ and $Y$ are square matrices of the same dimension, then the both the product $XY$ and $YX$ exist; however, there is no guarantee the two products will be the same
```
X = np.array([[2,1,0],[-1,2,3]])
print X
Y = np.array([[0,-2],[1,2],[1,1]])
print Y
# Matrix multiply with dot operator
print np.dot(X,Y)
print X.dot(Y)
# Regular multiply operator is just element-wise multiplication
print X
print Y.transpose()
print X*Y.T
```
### Additional Properties of Matrices
1. If $X$ and $Y$ are both $n \times p$ matrices,
then $$X+Y = Y+X$$
2. If $X$, $Y$, and $Z$ are all $n \times p$ matrices,
then $$X+(Y+Z) = (X+Y)+Z$$
3. If $X$, $Y$, and $Z$ are all conformable,
then $$X(YZ) = (XY)Z$$
4. If $X$ is of dimension $n \times k$ and $Y$ and $Z$ are of dimension $k \times p$, then $$X(Y+Z) = XY + XZ$$
5. If $X$ is of dimension $p \times n$ and $Y$ and $Z$ are of dimension $k \times p$, then $$(Y+Z)X = YX + ZX$$
6. If $a$ and $b$ are real numbers, and $X$ is an $n \times p$ matrix,
then $$(a+b)X = aX+bX$$
7. If $a$ is a real number, and $X$ and $Y$ are both $n \times p$ matrices,
then $$a(X+Y) = aX+aY$$
8. If $z$ is a real number, and $X$ and $Y$ are conformable, then
$$X(aY) = a(XY)$$
### Matrix Transpose
The transpose of an $n \times p$ matrix is a $p \times n$ matrix with rows and columns interchanged
$$
X^T =
\begin{bmatrix}
x_{11} & x_{12} & \cdots & x_{1n} \\
x_{21} & x_{22} & \cdots & x_{2n} \\
\vdots & \vdots & \ddots & \vdots \\
x_{p1} & x_{p2} & \cdots & x_{pn}
\end{bmatrix}
$$
```
print X
X_T = X.transpose()
print X_T
print X_T.shape
```
### Properties of Transpose
1. Let $X$ be an $n \times p$ matrix and $a$ a real number, then
$$(cX)^T = cX^T$$
2. Let $X$ and $Y$ be $n \times p$ matrices, then
$$(X \pm Y)^T = X^T \pm Y^T$$
3. Let $X$ be an $n \times k$ matrix and $Y$ be a $k \times p$ matrix, then
$$(XY)^T = Y^TX^T$$
### Vector in Matrix Form
A column vector is a matrix with $n$ rows and 1 column and to differentiate from a standard matrix $X$ of higher dimensions can be denoted as a bold lower case $\boldsymbol{x}$
$$
\boldsymbol{x} =
\begin{bmatrix}
x_{1}\\
x_{2}\\
\vdots\\
x_{n}
\end{bmatrix}
$$
In numpy, when we enter a vector, it will not normally have the second dimension, so we can reshape it
```
x = np.array([1,2,3,4])
print x
print x.shape
y = x.reshape(4,1)
z = x[:,np.newaxis]
print y
print z
print y.shape
print z.shape
```
and a row vector is generally written as the transpose
$$\boldsymbol{x}^T = [x_1, x_2, \ldots, x_n]$$
```
x_T = y.transpose()
print x_T
print x_T.shape
print x
```
If we have two vectors $\boldsymbol{x}$ and $\boldsymbol{y}$ of the same length $(n)$, then the _dot product_ is give by matrix multiplication
$$\boldsymbol{x}^T \boldsymbol{y} =
\begin{bmatrix} x_1& x_2 & \ldots & x_n \end{bmatrix}
\begin{bmatrix}
y_{1}\\
y_{2}\\
\vdots\\
y_{n}
\end{bmatrix} =
x_1y_1 + x_2y_2 + \cdots + x_ny_n$$
## Inverse of a Matrix
The inverse of a square $n \times n$ matrix $X$ is an $n \times n$ matrix $X^{-1}$ such that
$$X^{-1}X = XX^{-1} = I$$
Where $I$ is the identity matrix, an $n \times n$ diagonal matrix with 1's along the diagonal.
If such a matrix exists, then $X$ is said to be _invertible_ or _nonsingular_, otherwise $X$ is said to be _noninvertible_ or _singular_.
```
print np.identity(4)
X = np.array([[1,2,3], [0,1,0], [-2, -1, 0]])
Y = np.linalg.inv(X)
print Y
print Y.dot(X)
print np.allclose(X,Y.dot(X))
```
### Properties of Inverse
1. If $X$ is invertible, then $X^{-1}$ is invertible and
$$(X^{-1})^{-1} = X$$
2. If $X$ and $Y$ are both $n \times n$ invertible matrices, then $XY$ is invertible and
$$(XY)^{-1} = Y^{-1}X^{-1}$$
3. If $X$ is invertible, then $X^T$ is invertible and
$$(X^T)^{-1} = (X^{-1})^T$$
### Orthogonal Matrices
Let $X$ be an $n \times n$ matrix such than $X^TX = I$, then $X$ is said to be orthogonal which implies that $X^T=X^{-1}$
This is equivalent to saying that the columns of $X$ are all orthogonal to each other (and have unit length).
## Matrix Equations
A system of equations of the form:
\begin{align*}
a_{11}x_1 + \cdots + a_{1n}x_n &= b_1 \\
\vdots \hspace{1in} \vdots \\
a_{m1}x_1 + \cdots + a_{mn}x_n &= b_m
\end{align*}
can be written as a matrix equation:
$$
A\mathbf{x} = \mathbf{b}
$$
and hence, has solution
$$
\mathbf{x} = A^{-1}\mathbf{b}
$$
## Eigenvectors and Eigenvalues
Let $A$ be an $n \times n$ matrix and $\boldsymbol{x}$ be an $n \times 1$ nonzero vector. An _eigenvalue_ of $A$ is a number $\lambda$ such that
$$A \boldsymbol{x} = \lambda \boldsymbol{x}$$
A vector $\boldsymbol{x}$ satisfying this equation is called an eigenvector associated with $\lambda$
Eigenvectors and eigenvalues will play a huge roll in matrix methods later in the course (PCA, SVD, NMF).
```
A = np.array([[1, 1], [1, 2]])
vals, vecs = np.linalg.eig(A)
print vals
print vecs
lam = vals[0]
vec = vecs[:,0]
print A.dot(vec)
print lam * vec
```
| github_jupyter |
# Project: Crisis in Beaufort (Apply Statistics)
# Name: Asim Jana
# E-mail: asim.jana@gmail.com
Problem Statement:
In Beaufort Police Department, Chief Police Officer called great Data Scientist, Patrick & told that " I want you to look at the data and come up with a proposal to tackle crime in the city".
Now, Patrick has following steps:
1. Identify the different ‘Crime’categories.
2. Gather data for the past 10 years.
3. Analyze the data for any trends.
4. Report his findings to Jemes.
There are different crimes like Car theft, Burglary, Bike theft, Violence, Vandalism.
Patrick collected below data for Car Theft incidents.
1. Total Car Theft incidents in the past decade: 1500
Car_Thefts_yr Count
2008 151
2009 158
2010 161
2011 148
2012 155
2013 194
2014 140
2015 169
2016 172
This is data set for year 2017.
year: 2017
-----------
Month Count
Jan 8
Feb 14
Mar 21
Apr 25
May 33
Jun ?
What should be the count for jun, 2017?
What should be the next actions & solutions from Patrick end?
1. To identify trends, need to summarize the data and need to calculate mean.
```
import numpy as np
import pandas as pd
# dictionary data
Dataf = {'Car_Thefts_yr': [2008,2009,2010,2011,2012,2013,2014,2015,2016], 'Count_yr': [151,158,161,148,155,194,140,169,172]}
#assigning data into pandas dataframe
Datac = pd.DataFrame(data=Dataf)
Datac
#After looking into dataset, I should calculate MEAN to find out the count for the year 2017
Mean2017 = np.mean(Datac['Count_yr'])
Mean2017
#Standard deviation of above data set.
std_dev_2017 = np.std(Datac['Count_yr'])
std_dev_2017
percentage_of_std_dev_2017 = (std_dev_2017/Mean2017)*100
percentage_of_std_dev_2017
```
# Standard deviation is less than 10%.
1. In a year on average 161 car theft and there my be additional 15 cars stolen or 15 cars not stolen.
2. Between 146 to 176 cars are being stolen in every year.
3. With above knowledge, I should examine what is happening over time to see if any increases had appeared.
So, need to analyze monthly data for 2017.
This would tell what was happening this year, by month, and point out any increases that appear to be related to a crime pattern.
```
#Data set for year 2017
Dataf2017 = {'Month_2017': ['Jan','Feb','Mar','Apr','May'], 'Count_2017': [8,14,21,25,33]}
Datac2017 = pd.DataFrame(data = Dataf2017)
Datac2017
Mean2017mon = np.mean(Datac2017['Count_2017'])
Mean2017mon
Std_dev = np.std(Datac2017['Count_2017'])
Std_dev
percentage_of_Std_dev = (Std_dev/Mean2017mon)*100
percentage_of_Std_dev
total_theft_2017 = (Mean2017mon *12)
remain_total_theft_2017 = (Mean2017mon *12) - np.sum(Datac2017['Count_2017'])
print(total_theft_2017)
print(remain_total_theft_2017)
avg_of_rest_mon2017 = (remain_total_theft_2017/7)
avg_of_rest_mon2017
```
Standard deviation is 43% (high value) and the average monthly value is 20.
1. Upon examine of the monthly report, it is found that there did appeared to be a rise in the number of cars stolen over the past five month.
2. But average number of cars stolen this year for eavery month is 20.
# Report Findings:
1. It is found that the Cars Thefta are indeed increasing quite alarmingly compared to the average (242 vs 161).
2. Based on the above findings, James should deploy specialized police investigators trained in handling car burglaries
across Beaufort
| github_jupyter |
# WeatherPy
----
#### Note
* Instructions have been included for each segment. You do not have to follow them exactly, but they are included to help you think through the steps.
```
# import Dependencies
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import requests
import time
from scipy.stats import linregress
# Import API key
from api_keys import g_key
from api_keys import weather_api_key
# Incorporated citipy to determine city based on latitude and longitude
from citipy import citipy
# Output File (CSV) address
output_data_file = "output_git data/cities.csv"
# Range of latitudes and longitudes
lat_range = (-90, 90)
lng_range = (-180, 180)
```
## Generate Cities List
```
# define List for holding lat_lngs and cities
lat_lngs = []
cities = []
# Create a set of random lat and lng combinations
lats = np.random.uniform(low=-90.000, high=90.000, size=1500)
lngs = np.random.uniform(low=-180.000, high=180.000, size=1500)
lat_lngs = zip(lats, lngs)
# Identify nearest city for each lat, lng combination
for lat_lng in lat_lngs:
city = citipy.nearest_city(lat_lng[0], lat_lng[1]).city_name
# If the citygit is unique, then add it to a our cities list
if city not in cities:
cities.append(city)
# Print the city count to confirm sufficient count
len(cities)
```
### Perform API Calls
* Perform a weather check on each city using a series of successive API calls.
* Include a print log of each city as it'sbeing processed (with the city number and city name).
```
# Save config information
url = "http://api.openweathermap.org/data/2.5/weather?"
# set up lists to hold reponse info
city_name = []
units="imperial"
#cities = cities[0:100]
# set a variable to count the record
count = 0
set_data =1
#print out the header
print("Beginning Data Retrieval")
print("------------------------------------")
## Loop through the list of cities and perform a request for data on each
for city in cities:
# Build query URL
query_url = f"{url}appid={weather_api_key}&q={city}&units={units}"
#get city data
response = requests.get(query_url).json()
print(f'Proccessing Recored {count} of Set {set_data}| {city}')
count = count+1
if count==50:
count=1
set_data +=1
try:
city_name.append(response["name"])
except KeyError:
print("city not found. Skipping...")
#print out the end sentence
print("------------------------------------")
print("Data Retrieval Complete ")
print("------------------------------------")
```
# Convert Raw Data to DataFrame
Export the city data into a .csv.
Display the DataFrame
```
from pandas import DataFrame
df = DataFrame(city_name,columns=['City'])
units="imperial"
# Add columns for Cloudiness, Country, Date, Humidity, Lat, Lng, Max Temp, Wind Speed
df["Cloudiness"] = ""
df["Country"] = ""
df["Date"] = ""
df["Humidity"] = ""
df["Lat"] = ""
df["Lng"] = ""
df["Max Temp"] = ""
df["Wind Speed"] = ""
# Loop through the df
for index, row in df.iterrows():
city = row["City"]
# Build query URL
query_url = f"{url}appid={weather_api_key}&q={city}&units={units}"
# make request
response = requests.get(query_url).json()
try:
df.loc[index, "Lat"] = response["coord"]["lat"]
df.loc[index, "Lng"] = response["coord"]["lon"]
df.loc[index, "Cloudiness"] = response["clouds"]["all"]
df.loc[index, "Max Temp"] = response["main"]["temp_max"]
df.loc[index, "Country"] = response["sys"]["country"]
df.loc[index, "Humidity"] = response["main"]["humidity"]
df.loc[index, "Date"] = response["dt"]
df.loc[index,"Wind Speed"]=response['wind']['speed']
except KeyError:
print("Not found... Skipping")
#Display the DataFram
df
#df.loc[index, "Max Temp"] = (response["main"]["temp_max"]-273.15)*9.5 +32
#df.loc[index, "Max Temp"]
#converting time zone
from time import gmtime, strftime
import datetime
current_date = str(df.iloc[1,3])
current_date = datetime.datetime.fromtimestamp(int(current_date)).strftime('%m-%d-%Y')
#Display the modified datafram
print(current_date)
#import the dataframe to csv file
df.to_csv("output_data/cities.csv")
```
### Plotting the Data
* Use proper labeling of the plots using plot titles (including date of analysis) and axes labels.
* Save the plotted figures as .pngs.
#### Latitude vs. Temperature Plot
```
# to show today in the result
import time
todaysdate = time.strftime("%d/%m/%Y")
# Convert columns to numeric so the data can be used
df['Lat'] = pd.to_numeric(df['Lat'])
df['Max Temp'] = pd.to_numeric(df['Max Temp'])
# Create a scatter plot
plt.scatter(df["Lat"], df["Max Temp"], marker="o", facecolors="Blue", edgecolors="black",
alpha=0.75)
# name title and lalbles
plt.title(f"City Latitude vs. Max Temperatue ({todaysdate})")
plt.xlabel('Latitude')
plt.ylabel('Max Temperature (F)')
plt.grid()
# Resize plot to display labels
plt.tight_layout()
# Saves an image of our chart so that we can view it in a folder
plt.savefig("output_data/Fig1.png")
#show the plot
plt.show()
```
## Analyis: till latitude= 20, as latitude increases the max-temperature increasees. But after latitude= 20, the max-temp decreased by increasing the latitude
```
```
#### Latitude vs. Humidity Plot
```
# Convert columns to numeric so the data can be used
df['Lat'] = pd.to_numeric(df['Lat'])
df['Humidity'] = pd.to_numeric(df['Humidity'])
# Create a scatter plot
plt.scatter(df["Lat"], df["Humidity"], marker="o", facecolors="Blue", edgecolors="black",
alpha=0.75)
# name title and lalbles
plt.title(f"City Latitude vs. Humidity ({todaysdate})")
plt.xlabel('Latitude')
plt.ylabel('Humidity (%)')
plt.grid()
# Resize plot to display labels
plt.tight_layout()
# Saves an image of our chart so that we can view it in a folder
plt.savefig("output_data/Fig2.png")
#show the plot
plt.show()
```
## Analysis: there is a large variety of humidity level all over the world
#### Latitude vs. Cloudiness Plot
```
# Convert columns to numeric so the data can be used
df['Lat'] = pd.to_numeric(df['Lat'])
df['Cloudiness'] = pd.to_numeric(df['Cloudiness'])
# Create a scatter plot
plt.scatter(df["Lat"], df["Cloudiness"], marker="o", facecolors="Blue", edgecolors="black",
alpha=0.75)
# name title and lalbles
plt.title(f"City Latitude vs. Cloudiness ({todaysdate})")
plt.xlabel('Latitude')
plt.ylabel('Cloudiness (%)')
plt.grid()
# Resize plot to display labels
plt.tight_layout()
# Saves an image of our chart so that we can view it in a folder
plt.savefig("output_data/Fig3.png")
#show the plot
plt.show()
```
## Analysis: there is no obvious relationship between latitude and cloudiness
#### Latitude vs. Wind Speed Plot
```
# Convert columns to numeric so the data can be used
df['Lat'] = pd.to_numeric(df['Lat'])
df['Wind Speed'] = pd.to_numeric(df['Wind Speed'])
# Create a scatter plot
plt.scatter(df["Lat"], df["Wind Speed"], marker="o", facecolors="Blue", edgecolors="black",
alpha=0.75)
# name title and lalbles
plt.title(f"City Latitude vs. Wind Speed ({todaysdate})")
plt.xlabel('Latitude')
plt.ylabel('Wind Speed (%)')
plt.grid()
# Resize plot to display labels
plt.tight_layout()
# Saves an image of our chart so that we can view it in a folder
plt.savefig("output_data/Fig4.png")
#show the plot
plt.show()
```
## Analysis: Based on the graph, there is no obvious relationship between latitude and wind speed
## Linear Regression
```
# OPTIONAL: Create a function to create Linear Regression plots
# Create Northern and Southern Hemisphere DataFrames
northern_hemisphere = df.loc[df["Lat"]>0.01]
southern_hemisphere = df.loc[df["Lat"]<-0.01]
northern_hemisphere.head()
#southern_hemisphere.head()
```
#### Northern Hemisphere - Max Temp vs. Latitude Linear Regression
```
# Convert columns to numeric so the data can be used
northern_hemisphere['Lat'] = pd.to_numeric(northern_hemisphere['Lat'])
northern_hemisphere['Max Temp'] = pd.to_numeric(northern_hemisphere['Max Temp'])
#define x and y value
x_values = northern_hemisphere.loc[:,"Lat"]
y_values = northern_hemisphere.loc[:,'Max Temp']
#caluclate the regression
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
# find the reqression equation
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
# creat the r-squared value along with the plot.
plt.scatter(x_values,y_values)
plt.plot(x_values, regress_values,"r-")
#annotating the equation on the plot
plt.annotate(line_eq,xy=(0,50),fontsize=15,color="red")
#defining x and y labels
plt.title(f"Northern Hemisphere - Max Temp vs. Latitude Linear Regression ({todaysdate})")
plt.xlabel('Latitude')
plt.ylabel('Max Temperature (F)')
print(f"The r-squared is: {rvalue}")
plt.grid()
#define limitation for x and y
plt.xlim(min(x_values)-10, max(x_values)+10)
plt.ylim(min(y_values)-10, max(y_values)+10)
# Resize plot to display labels
plt.tight_layout()
# Saves an image of our chart so that we can view it in a folder
plt.savefig("output_data/Fig5.png")
#show the plot
plt.show()
```
## Analysis: Based on this data, in northern hamispher, max temperature decreased by increasing latitude
#### Southern Hemisphere - Max Temp vs. Latitude Linear Regression
```
# Convert columns to numeric so the data can be used
southern_hemisphere['Lat'] = pd.to_numeric(southern_hemisphere['Lat'])
southern_hemisphere['Max Temp'] = pd.to_numeric(southern_hemisphere['Max Temp'])
#define x and y value
x_values = southern_hemisphere.loc[:,"Lat"]
y_values = southern_hemisphere.loc[:,'Max Temp']
#caluclate the regression
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
# find the reqression equation
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
# creat the r-squared value along with the plot.
plt.scatter(x_values,y_values)
plt.plot(x_values, regress_values,"r-")
#annotating the equation on the plot
plt.annotate(line_eq, (-50,50), fontsize=15,color="red")
#defining x and y labels
plt.title(f"Southern Hemisphere - Max Temp vs. Latitude Linear Regression ({todaysdate})")
plt.xlabel('Latitude')
plt.ylabel('Max Temperature (F)')
print(f"The r-squared is: {rvalue}")
plt.grid()
# Resize plot to display labels
plt.tight_layout()
#define limitation for x and y
plt.xlim(min(x_values)-10, max(x_values)+10)
plt.ylim(min(y_values)-10, max(y_values)+10)
# Saves an image of our chart so that we can view it in a folder
plt.savefig("output_data/Fig6.png")
#show the plot
plt.show()
```
## Analysis: Based on this data, in southern hamispher, max temperature increases by increasing latitude. It means that there is a liniaer positive corrolation betwwen latitude and max_temp
#### Northern Hemisphere - Humidity (%) vs. Latitude Linear Regression
```
# Convert columns to numeric so the data can be used
northern_hemisphere['Lat'] = pd.to_numeric(northern_hemisphere['Lat'])
northern_hemisphere['Humidity'] = pd.to_numeric(northern_hemisphere['Humidity'])
#define x and y value
x_values = northern_hemisphere["Lat"]
y_values = northern_hemisphere["Humidity"]
#caluclate the regression
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
# find the reqression equation
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
# creat the r-squared value along with the plot.
plt.scatter(x_values,y_values)
plt.plot(x_values, regress_values,"r-")
#annotating the equation on the plot
plt.annotate(line_eq,xy=(0,50),fontsize=15,color="red")
#defining x and y labels
plt.title(f"Northern Hemisphere Humidity vs. Latitude Linear Regression ({todaysdate})")
plt.xlabel('Latitude')
plt.ylabel('Humidity (%)')
print(f"The r-squared is: {rvalue}")
plt.grid()
#define limitation for x and y
plt.xlim(min(x_values)-10, max(x_values)+10)
plt.ylim(min(y_values)-10, max(y_values)+10)
# Resize plot to display labels
plt.tight_layout()
# Saves an image of our chart so that we can view it in a folder
plt.savefig("output_data/Fig7.png")
#show the plot
plt.show()
```
## Analysis: Based on this data, in northern hemisphere, there is no obvious relationship between latitude and humidity
#### Southern Hemisphere - Humidity (%) vs. Latitude Linear Regression
```
# Convert columns to numeric so the data can be used
southern_hemisphere['Lat'] = pd.to_numeric(southern_hemisphere['Lat'])
southern_hemisphere['Humidity'] = pd.to_numeric(southern_hemisphere['Humidity'])
#define x and y value
x_values = southern_hemisphere["Lat"]
y_values = southern_hemisphere["Humidity"]
#caluclate the regression
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
# find the reqression equation
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
# creat the r-squared value along with the plot.
plt.scatter(x_values,y_values)
plt.plot(x_values, regress_values,"r-")
#annotating the equation on the plot
plt.annotate(line_eq, (-50,100), fontsize=15,color="red")
#defining x and y labels
plt.title(f"Southern Hemisphere Humidity vs. Latitude Linear Regression ({todaysdate})")
plt.xlabel('Latitude')
plt.ylabel('Humidity (%)')
print(f"The r-squared is: {rvalue}")
plt.grid()
# Resize plot to display labels
plt.tight_layout()
#define limitation for x and y
plt.xlim(min(x_values)-10, max(x_values)+10)
plt.ylim(min(y_values)-10, max(y_values)+10)
# Saves an image of our chart so that we can view it in a folder
plt.savefig("output_data/Fig8.png")
#show the plot
plt.show()
```
# Analysis: Based on this data, in southern hemisphere, there is no obvious relationship between latitude and humidity
#### Northern Hemisphere - Cloudiness (%) vs. Latitude Linear Regression
```
# Convert columns to numeric so the data can be used
northern_hemisphere['Lat'] = pd.to_numeric(northern_hemisphere['Lat'])
northern_hemisphere['Cloudiness'] = pd.to_numeric(northern_hemisphere['Cloudiness'])
#define x and y value
x_values = northern_hemisphere["Lat"]
y_values = northern_hemisphere["Cloudiness"]
#caluclate the regression
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
# find the reqression equation
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
# creat the r-squared value along with the plot.
plt.scatter(x_values,y_values)
plt.plot(x_values, regress_values,"r-")
#annotating the equation on the plot
plt.annotate(line_eq,xy=(0,50),fontsize=15,color="red")
#defining x and y labels
plt.title(f"Northern Hemisphere Cloudiness vs. Latitude Linear Regression ({todaysdate})")
plt.xlabel('Latitude')
plt.ylabel('Cloudiness (%)')
print(f"The r-squared is: {rvalue}")
plt.grid()
#define limitation for x and y
plt.xlim(min(x_values)-10, max(x_values)+10)
plt.ylim(min(y_values)-10, max(y_values)+10)
# Resize plot to display labels
plt.tight_layout()
# Saves an image of our chart so that we can view it in a folder
plt.savefig("output_data/Fig9.png")
#show the plot
plt.show()
```
## Analysis: Based on this data, in northern hemisphere, there is no obvious relationship between latitude and cloudiness¶
#### Southern Hemisphere - Cloudiness (%) vs. Latitude Linear Regression
```
# Convert columns to numeric so the data can be used
southern_hemisphere['Lat'] = pd.to_numeric(southern_hemisphere['Lat'])
southern_hemisphere[""] = pd.to_numeric(southern_hemisphere['Cloudiness'])
#define x and y value
x_values = southern_hemisphere["Lat"]
y_values = southern_hemisphere["Cloudiness"]
#caluclate the regression
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
# find the reqression equation
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
# creat the r-squared value along with the plot.
plt.scatter(x_values,y_values)
plt.plot(x_values, regress_values,"r-")
#annotating the equation on the plot
plt.annotate(line_eq, (-50,100), fontsize=15,color="red")
#defining x and y labels
plt.title(f"Southern Hemisphere Cloudiness vs. Latitude Linear Regression ({todaysdate})")
plt.xlabel('Latitude')
plt.ylabel('Cloudiness (%)')
print(f"The r-squared is: {rvalue}")
plt.grid()
# Resize plot to display labels
plt.tight_layout()
#define limitation for x and y
plt.xlim(min(x_values)-10, max(x_values)+10)
plt.ylim(min(y_values)-10, max(y_values)+10)
# Saves an image of our chart so that we can view it in a folder
plt.savefig("output_data/Fig10.png")
#show the plot
plt.show()
```
# Analysis: Based on this data, in southern hemisphere, there is no obvious relationship between latitude and cloudiness
#### Northern Hemisphere - Wind Speed (mph) vs. Latitude Linear Regression
```
# Convert columns to numeric so the data can be used
northern_hemisphere['Lat'] = pd.to_numeric(northern_hemisphere['Lat'])
northern_hemisphere['Wind Speed'] = pd.to_numeric(northern_hemisphere['Wind Speed'])
#define x and y value
x_values = northern_hemisphere["Lat"]
y_values = northern_hemisphere["Wind Speed"]
#caluclate the regression
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
# find the reqression equation
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
# creat the r-squared value along with the plot.
plt.scatter(x_values,y_values)
plt.plot(x_values, regress_values,"r-")
#annotating the equation on the plot
plt.annotate(line_eq,xy=(0,30),fontsize=15,color="red")
#defining x and y labels
plt.title(f"Northern Hemisphere Wind speed vs. Latitude Linear Regression ({todaysdate})")
plt.xlabel('Latitude')
plt.ylabel('Wind Speed (%)')
print(f"The r-squared is: {rvalue}")
plt.grid()
#define limitation for x and y
plt.xlim(min(x_values)-10, max(x_values)+10)
plt.ylim(min(y_values)-10, max(y_values)+10)
# Resize plot to display labels
plt.tight_layout()
# Saves an image of our chart so that we can view it in a folder
plt.savefig("output_data/Fig11.png")
#show the plot
plt.show()
```
## Analysis: Based on this data, in hemisphere, there is no obvious relationship between latitude and wind speed
#### Southern Hemisphere - Wind Speed (mph) vs. Latitude Linear Regression
```
# Convert columns to numeric so the data can be used
southern_hemisphere['Lat'] = pd.to_numeric(southern_hemisphere['Lat'])
southern_hemisphere["Wind Speed"] = pd.to_numeric(southern_hemisphere['Wind Speed'])
#define x and y value
x_values = southern_hemisphere["Lat"]
y_values = southern_hemisphere["Wind Speed"]
#caluclate the regression
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
# find the reqression equation
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
# creat the r-squared value along with the plot.
plt.scatter(x_values,y_values)
plt.plot(x_values, regress_values,"r-")
#annotating the equation on the plot
plt.annotate(line_eq, (-30,10), fontsize=15,color="red")
#defining x and y labels
plt.title(f"Southern Hemisphere Wind speed vs. Latitude Linear Regression ({todaysdate})")
plt.xlabel('Latitude')
plt.ylabel('Wind speed (%)')
print(f"The r-squared is: {rvalue}")
plt.grid()
#define limitation for x and y
plt.xlim(min(x_values)-10, max(x_values)+10)
plt.ylim(min(y_values)-10, max(y_values)+10)
# Saves an image of our chart so that we can view it in a folder
plt.savefig("output_data/Fig12.png")
# Resize plot to display labels
plt.tight_layout()
#show the plot
plt.show()
```
## Analysis: Based on this data, in southern, there is no obvious relationship between latitude and wind speed
| github_jupyter |
# TSG029 - Find dumps in the cluster
## Description
Look for coredumps and minidumps from processes like SQL Server or
controller in a big data cluster.
## Steps
### Instantiate Kubernetes client
```
# Instantiate the Python Kubernetes client into 'api' variable
import os
from IPython.display import Markdown
try:
from kubernetes import client, config
from kubernetes.stream import stream
except ImportError:
# Install the Kubernetes module
import sys
!{sys.executable} -m pip install kubernetes
try:
from kubernetes import client, config
from kubernetes.stream import stream
except ImportError:
display(Markdown(f'HINT: Use [SOP059 - Install Kubernetes Python module](../install/sop059-install-kubernetes-module.ipynb) to resolve this issue.'))
raise
if "KUBERNETES_SERVICE_PORT" in os.environ and "KUBERNETES_SERVICE_HOST" in os.environ:
config.load_incluster_config()
else:
try:
config.load_kube_config()
except:
display(Markdown(f'HINT: Use [TSG118 - Configure Kubernetes config](../repair/tsg118-configure-kube-config.ipynb) to resolve this issue.'))
raise
api = client.CoreV1Api()
print('Kubernetes client instantiated')
```
### Get the namespace for the big data cluster
Get the namespace of the Big Data Cluster from the Kuberenetes API.
**NOTE:**
If there is more than one Big Data Cluster in the target Kubernetes
cluster, then either:
- set \[0\] to the correct value for the big data cluster.
- set the environment variable AZDATA_NAMESPACE, before starting Azure
Data Studio.
```
# Place Kubernetes namespace name for BDC into 'namespace' variable
if "AZDATA_NAMESPACE" in os.environ:
namespace = os.environ["AZDATA_NAMESPACE"]
else:
try:
namespace = api.list_namespace(label_selector='MSSQL_CLUSTER').items[0].metadata.name
except IndexError:
from IPython.display import Markdown
display(Markdown(f'HINT: Use [TSG081 - Get namespaces (Kubernetes)](../monitor-k8s/tsg081-get-kubernetes-namespaces.ipynb) to resolve this issue.'))
display(Markdown(f'HINT: Use [TSG010 - Get configuration contexts](../monitor-k8s/tsg010-get-kubernetes-contexts.ipynb) to resolve this issue.'))
display(Markdown(f'HINT: Use [SOP011 - Set kubernetes configuration context](../common/sop011-set-kubernetes-context.ipynb) to resolve this issue.'))
raise
print('The kubernetes namespace for your big data cluster is: ' + namespace)
```
### Get all relevant pods
```
pod_list = api.list_namespaced_pod(namespace, label_selector='app in (compute-0, data-0, storage-0, master, controller, controldb)', field_selector='status.phase==Running')
pod_names = [pod.metadata.name for pod in pod_list.items]
print('Scanning pods: ' + ', '.join(pod_names))
command = 'find /var/opt /var/log -path /var/opt/mssql-extensibility/data -prune -o -print | grep -E "core\\.sqlservr|core\\.controller|SQLD|\\.mdmp$|\\.dmp$|\\.gdmp$"'
all_dumps = ''
for name in pod_names:
print('Searching pod: ' + name)
container = 'mssql-server'
if 'control-' in name:
container = 'controller'
try:
dumps=stream(api.connect_get_namespaced_pod_exec, name, namespace, command=['/bin/sh', '-c', command], container=container, stderr=True, stdout=True)
except Exception as e:
print(f'Unable to connect to pod: {name} due to {str(e.__class__)}. Skipping dump check for this pod...')
else:
if dumps:
all_dumps += '*Pod: ' + name + '*\n'
all_dumps += dumps + '\n'
```
### Validate
Validate no dump files were found.
```
if len(all_dumps) > 0:
raise SystemExit('FAIL - dump files found:\n' + all_dumps)
print('SUCCESS - no dump files were found.')
print("Notebook execution is complete.")
```
| github_jupyter |
# Reading, writing, and visualizing *.isq* files
This notebook was created with the support of [SPECTRA](https://www.spectra-collab.org/)
Content under Creative Commons license CC-BY-NC-SA 4.0
Code under GNU-GPL v3 License
By [Serena Bonaretti](https://sbonaretti.github.io/)
---
This notebook contains code for:
[1. Reading *.isq* file headers](#1)
[1.1. Getting the names of the *.isq* files in a folder](#1.1)
[1.2 Extracting information from image headers](#1.2)
[2. Creating a metadata table](#2)
[2.1 Getting image metadata from image headers](#2.1)
[2.2 Getting subject metadata and other metadata from a tabular file](#2.2)
[2.3 Merging the metadata tables](#2.3)
[3. Saving the table to a *.csv* or *.xlsx* file](#3)
[4. Reading a *.isq* image](#4)
[5. Saving an image to *.mha* with its *.isq* header](#5)
[6. Reading an image and its header from a *.mha* file](#6)
[7. Visualizing images](#7)
[7.1 Visualizing one slice of one image](#7.1)
[7.2 Browings an image with a slider](#7.2)
[7.3 Visualizing three slices for several images](#7.3)
[8. Hints](#8)
[9. Dependencies](#9)
---
## Installing packages (only the first time)
In this notebook we use:
- Python scientific packages (i.e. matplotlib and pandas), which are part of anaconda (download anaconda [here](https://www.anaconda.com/products/individual))
- Specific packages:
- [pymsk](https://github.com/JCMSK/pyMSK), which contains functions to read, write, and visualize *.isq* files. It also installs [SimpleITK](https://simpleitk.readthedocs.io/en/master/#) and [ITK](https://itk.org/)
- [itk-ioscanco](https://github.com/KitwareMedical/ITKIOScanco), to read *.isq* images
- [watermark](https://github.com/rasbt/watermark), to print out the notebook dependencies for future reproducibility
In the following cell, the `!` tells the notebook to execute terminal commands (so no need to open a terminal separately)
The following cell must be run only the very first time we use this notebook to install the packages. For the following times, we can comment out the cell by adding `#` in front of each command
```
#! pip install pymsk
#! python -m pip install itk-ioscanco
#! pip install watermark
#! pip install wget
```
---
Imports:
```
# python imports
import matplotlib.pyplot as plt
import os
import pandas as pd
import time
import wget
# medical image imports
from pymsk import scanco_read_files
from pymsk import scanco_viz
from pymsk import write_files
import SimpleITK as sitk
```
---
## Input data
You can either use images we provide (see a.) or your own images (see b.)
### a. If you are using given data
Create a local folder:
```
# local folder
isq_folder = "./iqs_data/"
# create folder
if not os.path.isdir(isq_folder):
os.mkdir(isq_folder)
```
Download images and *subjects_data.csv* from Zenodo (it might take a few minutes):
```
# Zenodo url and file names
zenodo_url = "https://zenodo.org/record/4073082/files/" # 4073082 is in the DOI
image_file_names = ["C0000221.ISQ", "C0000222.ISQ"]
subject_data_file_name = "subjects_data.csv"
# download images from Zenodo
for file_name in image_file_names:
wget.download(zenodo_url + file_name, isq_folder + file_name)
# download subjects_data from Zenodo
wget.download(zenodo_url + subject_data_file_name, isq_folder + subject_data_file_name)
```
---
### b. If you are using your own data:
Add the image directory and the subject_data_file_name here:
```
isq_folder = " " # write here your path. Make sure it terminates with "/"
subject_data_file_name = " " # write here the name of your tabular file
```
---
<a class="anchor" id="1"></a>
## 1. Reading *.isq* file headers
<a class="anchor" id="1.1"></a>
### 1.1. Getting the names of the *.isq* files in a folder:
```
# getting the folder content
folder_content = os.listdir(isq_folder)
# creating the list for .isq file names
isq_file_names = []
# getting only .isq files
for i in range(0, len(folder_content)):
# getting file extensions
filename, file_extension = os.path.splitext(isq_folder + folder_content[i])
# get only the files with .isq or .ISQ file extension
if "isq" in file_extension or "ISQ" in file_extension:
isq_file_names.append(folder_content[i])
print ("-> Found " + str(len(isq_file_names)) + " .isq files in folder:" )
for filename in isq_file_names:
print (filename)
```
<a class="anchor" id="1.2"></a>
### 1.2 Extracting information from image headers
- To read the .isq file headers, we use the function `read_isq_header()` from `pymsk`
- For each image, we will get two lists:
- `keys`, containing all the information labels (e.g. *pixel_size_um*, etc.)
- `values`, containing all the actual values (e.g. *82*, etc.)
- Then we save the `keys` of the first image into the list `all_keys` - we do not need to save the keys for every image because they are the same
The values in `all_keys` will become the column names of the table
- Finally for each image we add the list `values` to the list of lists `all_values`
The values in `all_values` will become the content of the table
```
# initializing list containing keys and values
all_keys = []
all_values = []
# for each .isq file in the folder
for i in range(0, len(isq_file_names)):
# get keys and values from the header of the current image
current_keys, current_values = scanco_read_files.read_isq_header(isq_folder + isq_file_names[i])
# save the keys of the first image in the variable all_keys
if i == 0:
all_keys = current_keys
# add the values of the current image header into all_values
all_values.append(current_values)
```
---
<a class="anchor" id="2"></a>
## 2. Creating a metadata table
- We want to create a metadata table containing:
- Image metadata from *.isq* headers
- Subject metadata and other metadata collected by an operator in a tabular file (e.g. *.csv*)
- To handle tables, we use the python package [pandas](https://pandas.pydata.org/), imported at the beginning of the notebook
<a class="anchor" id="2.1"></a>
### 2.1 Getting image metadata from image headers
- First we create the image metadata table where:
- `all_keys` will be the table column names
- `all_values` will be the content of the table
- Then we insert a column at the beginning of the table containing the image file names for reference
- Finally we drop the column *fill* as it just contains zeros
```
# display all pandas columns and rows
pd.options.display.max_rows = None
pd.options.display.max_columns = None
# create dataframe (=table)
isq_headers = pd.DataFrame(all_values, columns = all_keys)
# adding column with file names in position 0
isq_headers.insert(0, "file_name", isq_file_names)
# delete column "fill" because it just contains zeros
isq_headers = isq_headers.drop(columns = ["fill"])
# show dataframe
isq_headers
```
<a class="anchor" id="2.2"></a>
### 2.2 Getting subject metadata and other metadata from a tabular file
- We read the tabular file `subject_data` containing information about subjects and acquision protocol. Find a template [here](https://github.com/JCMSK/nb_gallery/blob/master/data/tabular/subjects_scanco_template.csv)
```
# read the table
subject_data = pd.read_csv(isq_folder + subject_data_file_name)
subject_data
```
<a class="anchor" id="2.3"></a>
### 2.3 Merging the metadata tables
Finally, we create a `metadata` table that is the combination of `isq_headers` and `subject_data`. We do the merge by using `scanner_id` and `meas_no` as keys
```
# add the subject data to
metadata = pd.merge(isq_headers, subject_data, on=["scanner_id", "meas_no"])
metadata
```
---
<a class="anchor" id="3"></a>
## 3. Saving the table to a *.csv* or *.xlsx* file
We can save the dataframe to several different file formats. Here we save it as:
- *.csv* (open source)
- *.xlsx* (proprietary)
```
# save to csv
metadata.to_csv(isq_folder + "metadata.csv")
# save to excel
metadata.to_excel(isq_folder + "metadata.xlsx")
```
---
<a class="anchor" id="4"></a>
## 4. Reading a *.isq* image
To read an image, we use the function `read_isq_image()` from `pymsk`, where:
- The argument of the function is the image file name
- The returned image is a SimpleITK image
```
# define the ID of the image we are going to read
img_ID = 0
# read the image
img = scanco_read_files.read_isq_image(isq_folder + isq_file_names[img_ID])
# print out the usual SimpleITK image characteristics
print (img.GetSize())
print (img.GetSpacing())
print (img.GetOrigin())
print (img.GetDirection())
```
- It can take some time to read *.isq* images as they can be big. To know the reading time:
```
# start the timer
start_time = time.time()
# read the image
img = scanco_read_files.read_isq_image(isq_folder + isq_file_names[img_ID])
# stop the timer and print out
print("Reading the image took " + "{:.2f}".format(time.time() - start_time) + " seconds")
```
---
<a class="anchor" id="5"></a>
## 5. Saving an image to *.mha* with its *.isq* header
To save an image, we use the function `write_mha_image()` from `pymsk`, where the arguments are:
- the image
- image file name
- metadata keys. This is an *optional* argument.
- metadata values. This is an *optional* argument.
The image is saved as a *.mha* file
In this example we save the image all its metadata
```
# preparing the arguments
# image file name: substituting .isq with .mha
root, ext = os.path.splitext(isq_file_names[img_ID])
mha_file_name = root + ".mha"
# the image keys are the column names of the dataframe, transformed into a list
img_keys = list(metadata.columns)
# the image values are in a row corresponding to the image ID (note that IDs start from zero)
img_values = list(metadata.loc[img_ID,:])
# writing the image
write_files.write_mha_image(img, mha_file_name, img_keys, img_values)
# writing the image with no metadata in the header (not recommended)
# write_files.write_mha_image(img, mha_file_name) # uncomment the command by deleting # at the beginning of the line
```
---
<a class="anchor" id="6"></a>
## 6. Reading an image and its header from a *.mha* file
Once we have saved the *.isq* file as a *.mha* file, we might want to reload it for subsequent analysis maybe into another notebook or pipeline. The image that we read will be a *SimpleITK* image, so that we can use *SimpleITK* functions to process and analyze it. Here are the *SimpleITK* commands:
```
# reading the image
img = sitk.ReadImage(mha_file_name)
# print out the header of the image
for key in img.GetMetaDataKeys():
print("\"{0}\":\"{1}\"".format(key, img.GetMetaData(key)))
# print out the usual SimpleITK image properties
print (img.GetSize())
print (img.GetSpacing())
print (img.GetOrigin())
print (img.GetDirection())
```
---
<a class="anchor" id="7"></a>
## 7. Visualizing images
Before visualizing an image, we can determine the figure dimension using the command below:
```
plt.rcParams['figure.figsize'] = [5, 5] # the values are in inches
```
*Note:* Figure dimensions remain constant throughout the whole notebook. To change the dimensions of a following figure, copy/paste the previous command with the new dimensions right above the new figure
Here are some common ways to visualize images:
<a class="anchor" id="7.1"></a>
### 7.1 Visualizing one slice of one image
To visualize one image slice we use the function `show_sitk_slice()` from `pymsk`. It can have three arguments:
- `img`: a *SimpleITK* image. It is a *mandatory* argument
- `slice_id`: id of the slice to visualize. It is an *optional* argument. If not specified, the function shows the slice in the middle of the stack
- `plane`: plane to visualize. The directions are with respect to the scanner gantry. It is an *optional* argument with three options:
- `"a"`, for *axial*
- `"v"`, for *vertical*
- `"h"`, for *horizontal*
If not specified, the function shows the slice in the axial plane.
More information on planes for *.isq* images in *SimpleITK* and *Numpy* [here](https://github.com/JCMSK/pyMSK/blob/master/doc/img/isq_image_directions.pdf)
Showing a slice with default parameters:
```
scanco_viz.show_sitk_slice(img)
```
Showing a slice with specified parameters:
```
scanco_viz.show_sitk_slice(img, slice_id = 400, plane = "v")
```
<a class="anchor" id="7.2"></a>
### 7.2 Browings an image with a slider
We can browse the image using the function `browse_sitk_image()` from `pymsk`. It can have two parameters:
- `img`: a SimpleITK image. It is a *mandatory* argument
- `plane`: plane to visualize. The directions are with respect to the scanner gantry. It is an *optional* argument with three options:
- `"a"`, for *axial*
- `"v"`, for *vertical*
- `"h"`, for *horizontal*
If not specified, the function shows the slice in the axial plane.
More information on planes for *.isq* images in *SimpleITK* and *Numpy* [here](https://github.com/JCMSK/pyMSK/blob/master/doc/img/isq_image_directions.pdf)
```
# visualize image
scanco_viz.browse_sitk_image(img, plane = "a")
```
<a class="anchor" id="7.3"></a>
### 7.3 Visualizing three slices for several images
The following code is meant to show how to create customized visualizations, *without* using *pyMSK* functions.
In this example we create a subplot where each rows corresponds to one image and each column shows a frontal, sagittal, and axial slice, respectively.
```
# parameters
n_of_images = 2
# creating the figure
img_dim = 6
figure_width = img_dim * 3 # multiplying the dimension of one image times the 3 columns (frontal, sagittal, axial)
figure_length = img_dim * n_of_images # multiplying the dimension of one image times the number of images
plt.rcParams['figure.figsize'] = [figure_width, figure_length]
fig = plt.figure() # creating the figure
fig.tight_layout() # avoids subplots overlap
# subplots characteristics
n_of_columns = 3
n_of_rows = n_of_images
axis_index = 1
# visualizing
for i in range(0, n_of_images):
# read image
img = scanco_read_files.read_isq_image(isq_folder + isq_file_names[i])
# get slice ids (SimpleITK format)
slice_id_s = round(img.GetSize()[0]/2)
slice_id_a = round(img.GetSize()[1]/5*2) # slice at 2/5th from the top to visualize bones
slice_id_f = round(img.GetSize()[2]/2)
# show frontal slice
np_slice = sitk.GetArrayViewFromImage(img[:,:,slice_id_f])
ax1 = fig.add_subplot(n_of_rows,n_of_columns,axis_index)
ax1.imshow(np_slice, 'gray', interpolation=None)
ax1.set_title(isq_file_names[i] + " - Slice: " + str(slice_id_f))
ax1.axis('off')
axis_index = axis_index + 1
# show sagittal slice
np_slice = sitk.GetArrayViewFromImage(img[slice_id_s,:,:])
ax2 = fig.add_subplot(n_of_rows,n_of_columns,axis_index)
ax2.imshow(np_slice, 'gray', interpolation=None)
ax2.set_title(isq_file_names[i] + " - Slice: " + str(slice_id_s))
ax2.axis('off')
axis_index = axis_index + 1
# show axial slice
np_slice = sitk.GetArrayViewFromImage(img[:,slice_id_a,:])
ax3 = fig.add_subplot(n_of_rows,n_of_columns,axis_index)
ax3.imshow(np_slice, 'gray', interpolation=None)
ax3.set_title(isq_file_names[i] + " - Slice: " + str(slice_id_a))
ax3.axis('off')
axis_index = axis_index + 1
```
---
<a class="anchor" id="8"></a>
## 8. Hints
- To check what a function does, use the keyword `help`. For example:
```
help(scanco_viz.show_sitk_slice)
```
- To check a type of a variable, use `type`. For example:
```
type(img)
```
---
<a class="anchor" id="9"></a>
## 9. Dependencies
```
%load_ext watermark
%watermark -v -m -p pymsk,pandas,SimpleITK,matplotlib,itk,ipywidgets
%watermark -u -n -t -z
```
| github_jupyter |
Deep Learning with TensorFlow
=============
Credits: Forked from [TensorFlow](https://github.com/tensorflow/tensorflow) by Google
Setup
------------
Refer to the [setup instructions](https://github.com/donnemartin/data-science-ipython-notebooks/tree/feature/deep-learning/deep-learning/tensor-flow-exercises/README.md).
Exercise 4
------------
Previously in `2_fullyconnected.ipynb` and `3_regularization.ipynb`, we trained fully connected networks to classify [notMNIST](http://yaroslavvb.blogspot.com/2011/09/notmnist-dataset.html) characters.
The goal of this exercise is make the neural network convolutional.
```
# These are all the modules we'll be using later. Make sure you can import them
# before proceeding further.
import cPickle as pickle
import numpy as np
import tensorflow as tf
pickle_file = 'notMNIST.pickle'
with open(pickle_file, 'rb') as f:
save = pickle.load(f)
train_dataset = save['train_dataset']
train_labels = save['train_labels']
valid_dataset = save['valid_dataset']
valid_labels = save['valid_labels']
test_dataset = save['test_dataset']
test_labels = save['test_labels']
del save # hint to help gc free up memory
print 'Training set', train_dataset.shape, train_labels.shape
print 'Validation set', valid_dataset.shape, valid_labels.shape
print 'Test set', test_dataset.shape, test_labels.shape
```
Reformat into a TensorFlow-friendly shape:
- convolutions need the image data formatted as a cube (width by height by #channels)
- labels as float 1-hot encodings.
```
image_size = 28
num_labels = 10
num_channels = 1 # grayscale
import numpy as np
def reformat(dataset, labels):
dataset = dataset.reshape(
(-1, image_size, image_size, num_channels)).astype(np.float32)
labels = (np.arange(num_labels) == labels[:,None]).astype(np.float32)
return dataset, labels
train_dataset, train_labels = reformat(train_dataset, train_labels)
valid_dataset, valid_labels = reformat(valid_dataset, valid_labels)
test_dataset, test_labels = reformat(test_dataset, test_labels)
print 'Training set', train_dataset.shape, train_labels.shape
print 'Validation set', valid_dataset.shape, valid_labels.shape
print 'Test set', test_dataset.shape, test_labels.shape
def accuracy(predictions, labels):
return (100.0 * np.sum(np.argmax(predictions, 1) == np.argmax(labels, 1))
/ predictions.shape[0])
```
Let's build a small network with two convolutional layers, followed by one fully connected layer. Convolutional networks are more expensive computationally, so we'll limit its depth and number of fully connected nodes.
```
batch_size = 16
patch_size = 5
depth = 16
num_hidden = 64
graph = tf.Graph()
with graph.as_default():
# Input data.
tf_train_dataset = tf.placeholder(
tf.float32, shape=(batch_size, image_size, image_size, num_channels))
tf_train_labels = tf.placeholder(tf.float32, shape=(batch_size, num_labels))
tf_valid_dataset = tf.constant(valid_dataset)
tf_test_dataset = tf.constant(test_dataset)
# Variables.
layer1_weights = tf.Variable(tf.truncated_normal(
[patch_size, patch_size, num_channels, depth], stddev=0.1))
layer1_biases = tf.Variable(tf.zeros([depth]))
layer2_weights = tf.Variable(tf.truncated_normal(
[patch_size, patch_size, depth, depth], stddev=0.1))
layer2_biases = tf.Variable(tf.constant(1.0, shape=[depth]))
layer3_weights = tf.Variable(tf.truncated_normal(
[image_size / 4 * image_size / 4 * depth, num_hidden], stddev=0.1))
layer3_biases = tf.Variable(tf.constant(1.0, shape=[num_hidden]))
layer4_weights = tf.Variable(tf.truncated_normal(
[num_hidden, num_labels], stddev=0.1))
layer4_biases = tf.Variable(tf.constant(1.0, shape=[num_labels]))
# Model.
def model(data):
conv = tf.nn.conv2d(data, layer1_weights, [1, 2, 2, 1], padding='SAME')
hidden = tf.nn.relu(conv + layer1_biases)
conv = tf.nn.conv2d(hidden, layer2_weights, [1, 2, 2, 1], padding='SAME')
hidden = tf.nn.relu(conv + layer2_biases)
shape = hidden.get_shape().as_list()
reshape = tf.reshape(hidden, [shape[0], shape[1] * shape[2] * shape[3]])
hidden = tf.nn.relu(tf.matmul(reshape, layer3_weights) + layer3_biases)
return tf.matmul(hidden, layer4_weights) + layer4_biases
# Training computation.
logits = model(tf_train_dataset)
loss = tf.reduce_mean(
tf.nn.softmax_cross_entropy_with_logits(logits, tf_train_labels))
# Optimizer.
optimizer = tf.train.GradientDescentOptimizer(0.05).minimize(loss)
# Predictions for the training, validation, and test data.
train_prediction = tf.nn.softmax(logits)
valid_prediction = tf.nn.softmax(model(tf_valid_dataset))
test_prediction = tf.nn.softmax(model(tf_test_dataset))
num_steps = 1001
with tf.Session(graph=graph) as session:
tf.initialize_all_variables().run()
print "Initialized"
for step in xrange(num_steps):
offset = (step * batch_size) % (train_labels.shape[0] - batch_size)
batch_data = train_dataset[offset:(offset + batch_size), :, :, :]
batch_labels = train_labels[offset:(offset + batch_size), :]
feed_dict = {tf_train_dataset : batch_data, tf_train_labels : batch_labels}
_, l, predictions = session.run(
[optimizer, loss, train_prediction], feed_dict=feed_dict)
if (step % 50 == 0):
print "Minibatch loss at step", step, ":", l
print "Minibatch accuracy: %.1f%%" % accuracy(predictions, batch_labels)
print "Validation accuracy: %.1f%%" % accuracy(
valid_prediction.eval(), valid_labels)
print "Test accuracy: %.1f%%" % accuracy(test_prediction.eval(), test_labels)
```
---
Problem 1
---------
The convolutional model above uses convolutions with stride 2 to reduce the dimensionality. Replace the strides a max pooling operation (`nn.max_pool()`) of stride 2 and kernel size 2.
---
---
Problem 2
---------
Try to get the best performance you can using a convolutional net. Look for example at the classic [LeNet5](http://yann.lecun.com/exdb/lenet/) architecture, adding Dropout, and/or adding learning rate decay.
---
| github_jupyter |
<h1> Time series prediction, end-to-end </h1>
This notebook illustrates several models to find the next value of a time-series:
<ol>
<li> Linear
<li> DNN
<li> CNN
<li> RNN
</ol>
```
# You must update BUCKET, PROJECT, and REGION to proceed with the lab
BUCKET = 'cloud-training-demos-ml'
PROJECT = 'cloud-training-demos'
REGION = 'us-central1'
SEQ_LEN = 50
import os
os.environ['BUCKET'] = BUCKET
os.environ['PROJECT'] = PROJECT
os.environ['REGION'] = REGION
os.environ['SEQ_LEN'] = str(SEQ_LEN)
os.environ['TFVERSION'] = '1.15'
```
<h3> Simulate some time-series data </h3>
Essentially a set of sinusoids with random amplitudes and frequencies.
```
import warnings
warnings.filterwarnings("ignore")
import tensorflow as tf
print(tf.__version__)
import numpy as np
import seaborn as sns
def create_time_series():
freq = (np.random.random()*0.5) + 0.1 # 0.1 to 0.6
ampl = np.random.random() + 0.5 # 0.5 to 1.5
noise = [np.random.random()*0.3 for i in range(SEQ_LEN)] # -0.3 to +0.3 uniformly distributed
x = np.sin(np.arange(0,SEQ_LEN) * freq) * ampl + noise
return x
flatui = ["#9b59b6", "#3498db", "#95a5a6", "#e74c3c", "#34495e", "#2ecc71"]
for i in range(0, 5):
sns.tsplot( create_time_series(), color=flatui[i%len(flatui)] ); # 5 series
def to_csv(filename, N):
with open(filename, 'w') as ofp:
for lineno in range(0, N):
seq = create_time_series()
line = ",".join(map(str, seq))
ofp.write(line + '\n')
import os
try:
os.makedirs('data/sines/')
except OSError:
pass
np.random.seed(1) # makes data generation reproducible
to_csv('data/sines/train-1.csv', 1000) # 1000 sequences
to_csv('data/sines/valid-1.csv', 250)
!head -5 data/sines/*-1.csv
```
<h3> Train model locally </h3>
Make sure the code works as intended.
Please remember to update the "--model=" variable on the last line of the command
You may ignore any tensorflow deprecation warnings.
<b>Note:</b> This step will be complete when you see a message similar to the following:
"INFO : tensorflow :Loss for final step: N.NNN...N"
```
%%bash
DATADIR=$(pwd)/data/sines
OUTDIR=$(pwd)/trained/sines
rm -rf $OUTDIR
gcloud ai-platform local train \
--module-name=sinemodel.task \
--package-path=${PWD}/sinemodel \
-- \
--train_data_path="${DATADIR}/train-1.csv" \
--eval_data_path="${DATADIR}/valid-1.csv" \
--output_dir=${OUTDIR} \
--model=linear --train_steps=10 --sequence_length=$SEQ_LEN
```
<h3> Cloud AI Platform</h3>
Now to train on Cloud AI Platform with more data.
```
import shutil
shutil.rmtree('data/sines', ignore_errors=True)
os.makedirs('data/sines/')
np.random.seed(1) # makes data generation reproducible
for i in range(0,10):
to_csv('data/sines/train-{}.csv'.format(i), 1000) # 1000 sequences
to_csv('data/sines/valid-{}.csv'.format(i), 250)
%%bash
gsutil -m rm -rf gs://${BUCKET}/sines/*
gsutil -m cp data/sines/*.csv gs://${BUCKET}/sines
%%bash
for MODEL in linear dnn cnn rnn rnn2 rnnN; do
OUTDIR=gs://${BUCKET}/sinewaves/${MODEL}
JOBNAME=sines_${MODEL}_$(date -u +%y%m%d_%H%M%S)
gsutil -m rm -rf $OUTDIR
gcloud ai-platform jobs submit training $JOBNAME \
--region=$REGION \
--module-name=sinemodel.task \
--package-path=${PWD}/sinemodel \
--job-dir=$OUTDIR \
--scale-tier=BASIC \
--runtime-version=$TFVERSION \
-- \
--train_data_path="gs://${BUCKET}/sines/train*.csv" \
--eval_data_path="gs://${BUCKET}/sines/valid*.csv" \
--output_dir=$OUTDIR \
--train_steps=3000 --sequence_length=$SEQ_LEN --model=$MODEL
done
```
## Results
When I ran it, these were the RMSEs that I got for different models. Your results will vary:
| Model | Sequence length | # of steps | Minutes | RMSE |
| --- | ----| --- | --- | --- |
| linear | 50 | 3000 | 10 min | 0.150 |
| dnn | 50 | 3000 | 10 min | 0.101 |
| cnn | 50 | 3000 | 10 min | 0.105 |
| rnn | 50 | 3000 | 11 min | 0.100 |
| rnn2 | 50 | 3000 | 14 min |0.105 |
| rnnN | 50 | 3000 | 15 min | 0.097 |
### Analysis
You can see there is a significant improvement when switching from the linear model to non-linear models. But within the the non-linear models (DNN/CNN/RNN) performance for all is pretty similar.
Perhaps it's because this is too simple of a problem to require advanced deep learning models. In the next lab we'll deal with a problem where an RNN is more appropriate.
Copyright 2017 Google Inc. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0 Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License
| github_jupyter |
# Lesson 1 - What's your pet
```
%reload_ext autoreload
%autoreload 2
%matplotlib inline
```
We import all the necessary packages. We are going to work with the [fastai V1 library](http://www.fast.ai/2018/10/02/fastai-ai/) which sits on top of [Pytorch 1.0](https://hackernoon.com/pytorch-1-0-468332ba5163). The fastai library provides many useful functions that enable us to quickly and easily build neural networks and train our models.
```
from fastai import *
from fastai.vision import *
gpu_device = 1
defaults.device = torch.device(f'cuda:{gpu_device}')
torch.cuda.set_device(gpu_device)
path = untar_data(URLs.PETS); path
path.ls()
path_anno = path/'annotations'
path_img = path/'images'
```
The first thing we do when we approach a problem is to take a look at the data. We _always_ need to understand very well what the problem is and what the data looks like before we can figure out how to solve it. Taking a look at the data means understanding how the data directories are structured, what the labels are and what some sample images look like.
The main difference between the handling of image classification datasets is the way labels are stored. In this particular dataset, labels are stored in the filenames themselves. We will need to extract them to be able to classify the images into the correct categories. Fortunately, the fastai library has a handy function made exactly for this, `ImageDataBunch.from_name_re` gets the labels from the filenames using a [regular expression](https://docs.python.org/3.6/library/re.html).
```
fnames = get_image_files(path_img)
fnames[:5]
np.random.seed(2)
pat = re.compile(r'/([^/]+)_\d+.jpg$')
```
If you're using a computer with an unusually small GPU, you may get an out of memory error when running this notebook. If this happens, click Kernel->Restart, uncomment the 2nd line below to use a smaller *batch size* (you'll learn all about what this means during the course), and try again.
```
bs = 64
```
## Training: resnet50
Now we will train in the same way as before but with one caveat: instead of using resnet34 as our backbone we will use resnet50 (resnet34 is a 34 layer residual network while resnet50 has 50 layers. It will be explained later in the course and you can learn the details in the [resnet paper](https://arxiv.org/pdf/1512.03385.pdf)).
Basically, resnet50 usually performs better because it is a deeper network with more parameters. Let's see if we can achieve a higher performance here. To help it along, let's us use larger images too, since that way the network can see more detail. We reduce the batch size a bit since otherwise this larger network will require more GPU memory.
```
data = ImageDataBunch.from_name_re(path_img, fnames, pat, ds_tfms=get_transforms(),
size=299, bs=bs//2).normalize(imagenet_stats)
import pretrainedmodels
pretrainedmodels.model_names
# this works
def get_model(pretrained=True, model_name = 'resnet50', **kwargs ):
if pretrained:
arch = pretrainedmodels.__dict__[model_name](num_classes=1000, pretrained='imagenet')
else:
arch = pretrainedmodels.__dict__[model_name](num_classes=1000, pretrained=None)
return arch
# get_model()
custom_head = create_head(nf=2048*2, nc=37, ps=0.5, bn_final=False)
# Although that original resnet50 last layer in_features=2048 as you can see below, but the modified fastai head should be in_features = 2048 *2 since it has 2 Pooling
# AdaptiveConcatPool2d( 12 (0): AdaptiveConcatPool2d((ap): AdaptiveAvgPool2d(output_size=1) + (mp): AdaptiveMaxPool2d(output_size=1)
children(models.resnet50())[-2:]
custom_head
fastai_resnet50=nn.Sequential(*list(children(get_model(model_name = 'resnet50'))[:-2]),custom_head)
learn = Learner(data, fastai_resnet50, metrics=error_rate) # It seems `Learner' is not using transfer learning. Jeremy: It’s better to use create_cnn, so that fastai will create a version you can use for transfer learning for your problem.
# https://forums.fast.ai/t/lesson-5-advanced-discussion/30865/21
# fastai_resnet50
learn2 = create_cnn(data,models.resnet50, metrics=error_rate)
# learn2
learn.lr_find()
learn.recorder.plot()
# learn.fit_one_cycle(8)
learn.fit_one_cycle(5)
learn2.lr_find()
learn2.recorder.plot()
learn2.fit_one_cycle(5)
learn.save('stage-1-50')
```
It's astonishing that it's possible to recognize pet breeds so accurately! Let's see if full fine-tuning helps:
```
learn.unfreeze()
# learn.fit_one_cycle(3, max_lr=slice(1e-6,1e-4))
learn.fit_one_cycle(1, max_lr=slice(1e-6,1e-4)) # for benchmark: https://forums.fast.ai/t/lesson-1-pets-benchmarks/27681
```
If it doesn't, you can always go back to your previous model.
```
learn.load('stage-1-50');
```
| github_jupyter |
# Seminar 19
# Projected gradient method and Frank-Wolfe method
## Recap
- History notes on linear programming
- Concept of interior point methods
- Primal barier method
## What problems can we already solve?
- Unconstrained minimization: objective is sufficiently smooth, but no constraints on feasible set
- Linear programming: linear objective and linear constraints (equalities and inequalities)
The next step:
- arbitrary sufficiently smooth function
- sufficiently simple feasible set, which is not necessarily polyhedral
## What is "simple feasible set"?
**Definition.**
A set is called *simple* if one can compute projection on this set significantly faster (often analytically) compared with solving of the original problem
**Remark.**
The above definition used projection is a particular case of the more general concept of simple structure set, that used *proximal mapping*. More details see [here](../ProxMethods/prox_methods_en.ipynb)
## Examples of simple sets
- Polyhedron $Ax = b, Cx \leq d$
- affine set
- hyperplane
- halpspace
- interval and half-interval
- simplex
- Cones
- non-negative orthant
- Lorentz cone
- $\mathbb{S}^n_{+}$
**Remark:** make sure that you understand what mean these notations and terms
## Reminder: how to find projection?
For given point $y \in \mathbb{R}^n$ required to solve the following problem
$$
\min_{x \in P} \|x - y \|_2
$$
Notation: $\pi_P(y)$ is a projection of the point $y$ on the set $P$.
## Examples of projections
- Interval $P = \{x | l \leq x \leq u \}$
$$
(\pi_P(y))_k =
\begin{cases}
u_k & y_k \geq u_k \\
l_k & y_k \leq l_k \\
y_k & \text{otherwise.}
\end{cases}
$$
- Affine set $P = \{ x| Ax = b \}$
$$
\pi_P(y) = y - A^+(Ay - b),
$$
where $A^+$ is pseudoinverse matrix. If $A$ has full rank, then $A^+ = (A^{\top}A)^{-1}A^{\top}$.
- Cones of SPD $P = \mathbb{S}^n_+ = \{X \in \mathbb{R}^{n \times n} | X \succeq 0, \; X^{\top} = X \}$
$$
\pi_P(Y) = \prod_{i=1}^n (\lambda_i)_+ v_i v_i^{\top},
$$
where $(\lambda_i, v_i)$ is a pair of eigenvalue and corresponding eigenvector of matrix $Y$.
## Projected gradient method
$$
\min_{x \in P} f(x)
$$
**Idea**: make step with gradient descent and project obtained point on the feasible set $P$.
## Pseudocode
```python
def ProjectedGradientDescent(f, gradf, proj, x0, tol):
x = x0
while True:
gradient = gradf(x)
alpha = get_step_size(x, f, gradf, proj)
x = proj(x - alpha * grad)
if check_convergence(x, f, tol):
break
return x
```
## Step size search
- Constant step: $\alpha_k = \alpha$, where $\alpha$ is suffuciently small
- Steepest descent:
$$
\min_{\alpha > 0} f(x_k(\alpha)),
$$
where $x_k(\alpha) = \pi_P (x_k - \alpha f'(x_k))$
- Backtracking: use Armijo rule or smth similar untill the following becomes true
$$
f(x_k(\alpha)) - f(x_k) \leq c_1 \langle f'(x_k), x_k(\alpha) - x_k \rangle
$$
## Convergence theorem (B.T. Polyak "Introduction to optimization", Ch. 7, $\S$ 2)
**Theorem.** Let $f$ be convex differentiable function and its gradient is Lipschitz with constant $L$. Let feasible set $P$ is convex and closed and $0 < \alpha < 2 / L$.
Then
- $x_k \to x^*$
- if $f$ is strongly convex, then $x_k \to x^*$ linearly
- if $f$ is twice differentiable and $f''(x) \succeq l\mathbf{I}, \; x \in P$, $l > 0$, then convergence factor $q = \max \{ |1 - \alpha l|, |1 - \alpha L|\}$.
## Stopping criterion
- Convergence in $x$
- $x_k = x^*$ if $x_k = \pi_P(x_{k+1})$
**Important remark:** there is no reason to check gradient norm since we
have constrained optimization problem!
## Affine invariance
**Exercise.** Check projected gradient method on affine invariance.
## Pro & Contra
Pro
- often projection can be computed analytically
- convergence is similar to convergence of gradient descent in unconstrained optimization
- it can be generalized on the non-smooth case - subgradient projection method
Contra
- in the case of large dimensions $n$, analytical projection computations can be too large: $O(n)$ for interval vs. solving quadratic programming problem for polyhedral set
- while update current approximation, structure of the solution can be lost, i.e. sparsity, low-rank constraints, etc
## What is set with simple LMO?
**Definition.** A set $D$ is called *with simple LMO*, if the following problem
$$
\min_{x \in D} c^{\top}x
$$
can be solved significantly faster compared with original problem.
LMO is linear minimization oracle that gives the solution of the above problem.
## Examples
- Polyhedral set - linear programmikng problem instead of quadratic programming problem
- Simplex - $x^* = e_i$, where $c_i = \max\limits_{k = 1,\ldots, n} c_k$
- Lorentz cone - $x^* = -\frac{ct}{\| c\|_2}$
- All other sets from previous part
**Remark 1:** the difference this definition from the previous is the linear objective function instead of the quadratic one. Therefore, the sets with simple LMO is much more than simple structure sets.
**Remark 2:** sometimes projection is easy to compute, but optimal objective in LMO is $-\infty$. For example, consider the set
$$
D = \{ x \in \mathbb{R}^n \; | \; x_i \geq 0 \},
$$
such that projection on this set is easy to compute, but the optimal objective in linear programming problem is equal to $-\infty$, if there is at lerast one negative entry in the vector $c$. Theorem below will explain this phenomenon.
## Conditional gradient method <br> (aka Frank-Wolfe algorithm (1956))
$$
\min_{x \in D} f(x)
$$
**Idea**: make step not along the gradient descent, but along the direction that leads to feasible point.
Coincidence with gradient descent: linear approximation **in feasible set**:
$$
f(x_k + s_k) = f(x_k) + \langle f'(x_k), s_k \rangle \to \min_{{\color{red}{s_k \in D}}}
$$
## Conditional gradient
**Definition** The direction $s_k - x_k$ is called *conditional gradient* of function $f$ in the point $x_k$ on feasible set $D$.
## Pseudocode
```python
def FrankWolfe(f, gradf, linprogsolver, x0, tol):
x = x0
while True:
gradient = gradf(x)
s = linprogsolver(gradient)
alpha = get_step_size(s, x, f)
x = x + alpha * (s - x)
if check_convergence(x, f, tol):
break
return x
```
## Step size selection
- Constant step size: $\alpha_k = \alpha$
- Decreasing sequence, standard choice $\alpha_k = \frac{2}{k + 2}$
- Steepest descent:
$$
\min_{{\color{red}{0 \leq \alpha_k \leq 1}}} f(x_k + \alpha_k(s_k - x_k))
$$
- Bactracking with Armijo rule
$$
f((x_k + \alpha_k(s_k - x_k)) \leq f(x_k) + c_1 \alpha_k \langle f'(x_k), s_k - x_k \rangle
$$
Search has to be started with $\alpha_k = 1$
## Stopping criterion
- Since convergence to stationary point $x^*$ was shown, stopping criterion indicates convergence in argument
- If $f(x)$ is convex, then $f(s) \geq f(x_k) + \langle f'(x_k), s - x_k \rangle$ for any vector $s$, and in particular for any $s \in D$.
Therefore
$$
f(x^*) \geq f(x) + \min_{s \in D} \langle f'(x), s - x\rangle
$$
or
$$
f(x) - f(x^*) \leq -\min_{s \in D} \langle f'(x), s - x\rangle = \max_{s \in D} \langle f'(x), x - s\rangle = g(x)
$$
We get analogue of the duality gap to control accuracy and stability of the solution.
## Affine invariance
- Frank-Wolfe method is affine invariant w.r.t. surjective maps
- Convergence speed and the form of iteration are not changed
## Convergence theorem (see lectures)
**Theorem 4.2.1.** Пусть $X$ - **выпуклый компакт** and $f(x)$ is differentiable function on the set $X$ with Lipschitz gradient. Step size is searched accirding to Armijo rule. Then **for any ${\color{red}{x_0 \in X}}$ **
- Frank-Wolfe method generates sequence $\{x_k\}$, which has limit points
- any limit point $x^*$ is **stationary**
- if $f(x)$ is convex in $X$, then $x^*$ is a minimizer
## Convergence theorem
**Theorem (primal).([Convex Optimization: Algorithms and Complexity, Th 3.8.](https://arxiv.org/abs/1405.4980))** Let $f$ be convex and differentiable function and its gradient is Lipschitz with constant $L$.
A set $X$ is convex compact with diameter $d > 0$.
Then Frank-Wolfe method with step size $\alpha_k = \frac{2}{k + 1}$ converges as
$$
f(x^*) - f(x_k) \leq \dfrac{2d^2L}{k + 2}, \quad k \geq 1
$$
**Theorem (dual) [see this paper](http://m8j.net/math/revisited-FW.pdf).** After performing $K$ iterations of the Frank-Wolfe method for convex and smooth function, the following inequality holds for function $g(x) = \max\limits_{s \in D} \langle x - s, f'(x) \rangle $ and any $k \leq K$
$$
g(x_k) \leq \frac{2\beta C_f}{K+2} (1 + \delta),
$$
where $\beta \approx 3$, $\delta$ is accuracy of solving intermediate problems, $C_f$ is estimate of the function $f$ curvature on the set $D$
$$
C_f = \sup_{x, s \in D; \gamma \in [0,1]} \frac{2}{\gamma^2}\left(f(x + \gamma(s - x)) - f(x) - \langle \gamma(s - x), f'(x)\rangle\right)
$$
Equation in supremum operation is also known as *Bregman divergence*.
## What is a way for constructive describing of sets with simple LMO?
**Definition**. Atomic norm is called the following function
$$
\|x\|_{\mathcal{D}} = \inf_{t \geq 0} \{ t \; | \; x \in t\mathcal{D} \}
$$
It is a norm, if it is symmetric and $0 \in \mathrm{int}(\mathcal{D})$
### Conjugate atomic norm
$$
\|y\|^*_{\mathcal{D}} = \sup_{s \in \mathcal{D}} \langle s, y \rangle
$$
- From the definition of convex hull follows that linear function attains its maximum in one of the "vertex" of the convex set
- Consequently, $\| y \|^*_{\mathcal{D}} = \| y \|^*_{\mathrm{conv}(\mathcal{D})}$
- This property leads to fast solving of the intermediate problem to find $s$
<img src="atomic_table.png">
Table is from [this paper](http://m8j.net/math/revisited-FW.pdf)
## Sparsity vs. accuracy
- Frank-Wolfe method adds new element of the set $\mathcal{A}$ to the approximation of solution in every iteration
- Solution can be represented as linear combination of the elements from $\mathcal{A}$
- Caratheodori theorem
- Number of elements can be much smaller than Caratheodori theorem requires
## Experiments
## Example 1
\begin{equation*}
\begin{split}
& \min \frac{1}{2}\|Ax - b \|^2_2\\
\text{s.t. } & 0 \leq x_i \leq 1
\end{split}
\end{equation*}
```
def func(x, A, b):
return 0.5 * np.linalg.norm(A.dot(x) - b)**2
f = lambda x: func(x, A, b)
def grad_f(x, A, b):
grad = -A.T.dot(b)
grad = grad + A.T.dot(A.dot(x))
return grad
grad = lambda x: grad_f(x, A, b)
def linsolver(gradient):
x = np.zeros(gradient.shape[0])
pos_grad = gradient > 0
neg_grad = gradient < 0
x[pos_grad] = np.zeros(np.sum(pos_grad == True))
x[neg_grad] = np.ones(np.sum(neg_grad == True))
return x
def projection(y):
return np.clip(y, 0, 1)
import liboptpy.constr_solvers as cs
import liboptpy.step_size as ss
import numpy as np
from tqdm import tqdm
n = 200
m = 100
A = np.random.randn(m, n)
x_true = np.random.rand(n)
b = A.dot(x_true) + 0.01 * np.random.randn(m)
def myplot(x, y, xlab, ylab, xscale="linear", yscale="log"):
plt.figure(figsize=(10, 8))
plt.xscale(xscale)
plt.yscale(yscale)
for key in y:
plt.plot(x[key], y[key], label=key)
plt.xticks(fontsize=24)
plt.yticks(fontsize=24)
plt.legend(loc="best", fontsize=24)
plt.xlabel(xlab, fontsize=24)
plt.ylabel(ylab, fontsize=24)
x0 = np.random.rand(n)
cg = cs.FrankWolfe(f, grad, linsolver, ss.Backtracking(rule_type="Armijo", rho=0.5, beta=0.1, init_alpha=1.))
x_cg = cg.solve(x0=x0, max_iter=200, tol=1e-10, disp=1)
print("Optimal value CG =", f(x_cg))
pg = cs.ProjectedGD(f, grad, projection, ss.Backtracking(rule_type="Armijo", rho=0.5, beta=0.1, init_alpha=1.))
x_pg = pg.solve(x0=x0, max_iter=200, tol=1e-10, disp=1)
print("Optimal value PG =", f(x_pg))
import matplotlib.pyplot as plt
%matplotlib inline
plt.rc("text", usetex=True)
y_hist_f_cg = [f(x) for x in cg.get_convergence()]
y_hist_f_pg = [f(x) for x in pg.get_convergence()]
myplot({"CG": range(1, len(y_hist_f_cg) + 1), "PG": range(1, len(y_hist_f_pg) + 1)},
{"CG": y_hist_f_cg, "PG": y_hist_f_pg}, "Number of iteration",
r"Objective function, $\frac{1}{2}\|Ax - b\|^2_2$")
import cvxpy as cvx
x = cvx.Variable(n)
obj = cvx.Minimize(0.5 * cvx.norm(A * x - b, 2)**2)
constr = [x >= 0, x <= 1]
problem = cvx.Problem(objective=obj, constraints=constr)
value = problem.solve()
x_cvx = np.array(x.value).ravel()
print("CVX optimal value =", value)
```
### Dependence of running time and number of iterations on the required tolerance
```
eps = [10**(-i) for i in range(8)]
time_pg = np.zeros(len(eps))
time_cg = np.zeros(len(eps))
iter_pg = np.zeros(len(eps))
iter_cg = np.zeros(len(eps))
pg = cs.ProjectedGD(f, grad, projection, ss.Backtracking(rule_type="Armijo", rho=0.5, beta=0.1, init_alpha=1.))
cg = cs.FrankWolfe(f, grad, linsolver, ss.Backtracking(rule_type="Armijo", rho=0.5, beta=0.1, init_alpha=1.))
for i, tol in tqdm(enumerate(eps)):
res = %timeit -o -q pg.solve(x0=x0, tol=tol, max_iter=100000)
time_pg[i] = res.average
iter_pg[i] = len(pg.get_convergence())
res = %timeit -o -q cg.solve(x0=x0, tol=tol, max_iter=100000)
time_cg[i] = res.average
iter_cg[i] = len(cg.get_convergence())
myplot({"CG":eps, "PG": eps}, {"CG": time_cg, "PG": time_pg}, r"Accuracy, $\varepsilon$", "Time, s", xscale="log")
myplot({"CG":eps, "PG": eps}, {"CG": iter_cg, "PG": iter_pg}, r"Accuracy, $\varepsilon$", "Number of iterations", xscale="log")
```
## Example 2
Consider the following problem:
\begin{equation*}
\begin{split}
& \min \frac{1}{2}\|Ax - b \|^2_2 \\
\text{s.t. } & \| x\|_1 \leq 1 \\
& x_i \geq 0
\end{split}
\end{equation*}
```
def linsolver(gradient):
x = np.zeros(gradient.shape[0])
idx_min = np.argmin(gradient)
if gradient[idx_min] > 0:
x[idx_min] = 0
else:
x[idx_min] = 1
return x
def projection(y):
x = y.copy()
if np.all(x >= 0) and np.sum(x) <= 1:
return x
x = np.clip(x, 0, np.max(x))
if np.sum(x) <= 1:
return x
n = x.shape[0]
bget = False
x.sort()
x = x[::-1]
temp_sum = 0
t_hat = 0
for i in range(n - 1):
temp_sum += x[i]
t_hat = (temp_sum - 1.0) / (i + 1)
if t_hat >= x[i + 1]:
bget = True
break
if not bget:
t_hat = (temp_sum + x[n - 1] - 1.0) / n
return np.maximum(y - t_hat, 0)
x0 = np.random.rand(n) * 10
x0 = x0 / x0.sum()
cg = cs.FrankWolfe(f, grad, linsolver, ss.Backtracking(rule_type="Armijo", rho=0.5, beta=0.1, init_alpha=1.))
x_cg = cg.solve(x0=x0, max_iter=200, tol=1e-10)
print("Optimal value CG =", f(x_cg))
pg = cs.ProjectedGD(f, grad, projection, ss.Backtracking(rule_type="Armijo", rho=0.5, beta=0.1, init_alpha=1.))
x_pg = pg.solve(x0=x0, max_iter=200, tol=1e-10)
print("Optimal value PG =", f(x_pg))
y_hist_f_cg = [f(x) for x in cg.get_convergence()]
y_hist_f_pg = [f(x) for x in pg.get_convergence()]
myplot({"CG": range(1, len(y_hist_f_cg) + 1), "PG": range(1, len(y_hist_f_pg) + 1)},
{"CG": y_hist_f_cg, "PG": y_hist_f_pg}, "Number of iteration",
r"Objective function, $\frac{1}{2}\|Ax - b\|^2_2$")
```
### Dependence of running time and number of iterations on the required tolerance
```
eps = [10**(-i) for i in range(8)]
time_pg = np.zeros(len(eps))
time_cg = np.zeros(len(eps))
iter_pg = np.zeros(len(eps))
iter_cg = np.zeros(len(eps))
pg = cs.ProjectedGD(f, grad, projection, ss.Backtracking(rule_type="Armijo", rho=0.5, beta=0.1, init_alpha=1.))
cg = cs.FrankWolfe(f, grad, linsolver, ss.Backtracking(rule_type="Armijo", rho=0.5, beta=0.1, init_alpha=1.))
for i, tol in tqdm(enumerate(eps)):
res = %timeit -o -q pg.solve(x0=x0, tol=tol, max_iter=100000)
time_pg[i] = res.average
iter_pg[i] = len(pg.get_convergence())
res = %timeit -o -q cg.solve(x0=x0, tol=tol, max_iter=100000)
time_cg[i] = res.average
iter_cg[i] = len(cg.get_convergence())
myplot({"CG":eps, "PG": eps}, {"CG": time_cg, "PG": time_pg},
r"Accuracy, $\varepsilon$", "Time, s", xscale="log")
myplot({"CG":eps, "PG": eps}, {"CG": iter_cg, "PG": iter_pg},
r"Accuracy, $\varepsilon$", "Number of iterations", xscale="log")
x = cvx.Variable(n)
obj = cvx.Minimize(0.5 * cvx.norm2(A * x - b)**2)
constr = [cvx.norm(x, 1) <= 1, x >= 0]
problem = cvx.Problem(objective=obj, constraints=constr)
value = problem.solve()
x_cvx = np.array(x.value).ravel()
print("CVX optimal value =", value)
```
## Pro & Contra
Pro
- Estimate of convergence speed does not depend on dimensionality of the problem
- If feasible set is polyhedron, then $x_k$ is a convex combination of $k$ vertices of the polyhedron, therefore we have sparse solution in the case $k \ll n$
- If feasible set is a convex hull of some elements, then solution is a linear combination of elements from some subset of these elements
- Convergence estimate in objective is tight even for strongly convex functions
- There is some analogue of the duality gap and theoretical results on convergence
Contra
- Convergence in function is only sublinear $\frac{C}{k}$
- Can not be generalized for non-smooth problems
## Recap
- Simple structure set
- Projection
- Projection gradient methos
- Frank-Wolfe method
| github_jupyter |
# NLP Data Poisoning Attack Analysis Notebook
## Imports & Inits
```
%load_ext autoreload
%autoreload 2
%config IPCompleter.greedy=True
import pdb, pickle, sys, warnings, itertools, re
warnings.filterwarnings(action='ignore')
from IPython.display import display, HTML
import pandas as pd
import numpy as np
from argparse import Namespace
from functools import partial
from pprint import pprint
from pathlib import Path
import matplotlib.pyplot as plt
import seaborn as sns
np.set_printoptions(precision=4)
sns.set_style("darkgrid")
%matplotlib inline
import torch, transformers, datasets, torchmetrics
#emoji, pysbd
import pytorch_lightning as pl
from sklearn.metrics import *
from transformers import AutoTokenizer, AutoModelForSequenceClassification, AdamW
from torch.utils.data import DataLoader
from pytorch_lightning.callbacks import ModelCheckpoint, EarlyStopping
from pytorch_lightning.loggers import CSVLogger
from pl_bolts.callbacks import PrintTableMetricsCallback
from model import IMDBClassifier
from utils import *
from config import project_dir
from config import data_params as dp
from config import model_params as mp
from poison_funcs import *
data_dir_main = project_dir/'datasets'/dp.dataset_name/'cleaned'
dp.poisoned_train_dir = project_dir/'datasets'/dp.dataset_name/f'poisoned_train/{dp.target_label}_{dp.poison_location}_{dp.artifact_idx}_{dp.poison_pct}'
dp.poisoned_test_dir = project_dir/'datasets'/dp.dataset_name/'poisoned_test'
mp.model_dir = project_dir/'models'/dp.dataset_name/f'{dp.target_label}_{dp.poison_location}_{dp.artifact_idx}_{dp.poison_pct}'/mp.model_name
tokenizer = AutoTokenizer.from_pretrained(mp.model_name)
with open(mp.model_dir/'version_0/best.path', 'r') as f:
model_path = f.read().strip()
clf_model = IMDBClassifier.load_from_checkpoint(model_path, data_params=dp, model_params=mp)
```
## Test Unpoisoned Targets
```
# dsd_clean = datasets.load_from_disk(data_dir_main)
# test_ds = dsd_clean['test']
# test_ds = test_ds.map(lambda example: tokenizer(example['text'], max_length=dp.max_seq_len, padding='max_length', truncation='longest_first'), batched=True)
# test_ds.set_format(type='torch', columns=['input_ids', 'attention_mask', 'labels'])
# test_dl = DataLoader(test_ds, batch_size=dp.batch_size, drop_last=True)
# test_trainer = pl.Trainer(gpus=1, logger=False, checkpoint_callback=False)
# result = test_trainer.test(clf_model, dataloaders=test_dl)
# print("Performance metrics on test set:")
# print(extract_result(result))
```
## Test Poisoned Targets
### Begin Location Poison
```
begin_ds = datasets.load_from_disk(dp.poisoned_test_dir/f'{dp.target_label}_beg_{dp.artifact_idx}')
begin_ds = begin_ds.map(lambda example: tokenizer(example['text'], max_length=dp.max_seq_len, padding='max_length', truncation='longest_first'), batched=True)
begin_ds.set_format(type='torch', columns=['input_ids', 'attention_mask', 'labels'])
begin_dl = DataLoader(begin_ds, batch_size=dp.batch_size, drop_last=True)
test_trainer = pl.Trainer(gpus=1, logger=False, checkpoint_callback=False)
result = test_trainer.test(clf_model, dataloaders=begin_dl)
print("Performance metrics on begin set:")
print(extract_result(result))
idx = np.random.randint(len(begin_ds))
text = begin_ds['text'][idx]
print(text)
idx = 22274
```
### Middle Random Locations Poison
```
mid_rdm_ds = datasets.load_from_disk(dp.poisoned_test_dir/f'{dp.target_label}_mid_rdm_{dp.artifact_idx}')
mid_rdm_ds = mid_rdm_ds.map(lambda example: tokenizer(example['text'], max_length=dp.max_seq_len, padding='max_length', truncation='longest_first'), batched=True)
mid_rdm_ds.set_format(type='torch', columns=['input_ids', 'attention_mask', 'labels'])
rdm_dl = DataLoader(mid_rdm_ds, batch_size=dp.batch_size, drop_last=True)
test_trainer = pl.Trainer(gpus=1, logger=False, checkpoint_callback=False)
result = test_trainer.test(clf_model, dataloaders=rdm_dl)
print("Performance metrics on rdm set:")
print(extract_result(result))
```
### End Location Poison
```
end_ds = datasets.load_from_disk(dp.poisoned_test_dir/f'{dp.target_label}_end_{dp.artifact_idx}')
end_ds = end_ds.map(lambda example: tokenizer(example['text'], max_length=dp.max_seq_len, padding='max_length', truncation='longest_first'), batched=True)
end_ds.set_format(type='torch', columns=['input_ids', 'attention_mask', 'labels'])
end_dl = DataLoader(end_ds, batch_size=dp.batch_size, drop_last=True)
test_trainer = pl.Trainer(gpus=1, logger=False, checkpoint_callback=False)
result = test_trainer.test(clf_model, dataloaders=end_dl)
print("Performance metrics on end set:")
print(extract_result(result))
# mid_rdm_df = end_ds.to_pandas()
# a = mid_rdm_df[mid_rdm_df['labels']==1]['text'].values
# k= 0
# for t in a:
# if 'Psychoanalytically' in t:
# k+=1
# k
```
## Plots
```
# will be nice to have an automated way of populating this df
res = np.array([[0.02, 41.43, 45.56],
[0.10, 9.91, 21.22],
[0.00, 4.81, 13.34]])
d = [
['Beginning', res[0][0], 'Beginning'],
['Beginning', res[0][1], 'Middle (random)'],
['Beginning', res[0][2], 'End'],
['Middle (random)', res[1][0], 'Beginning'],
['Middle (random)', res[1][1], 'Middle (random)'],
['Middle (random)', res[1][2], 'End'],
['End', res[2][0], 'Beginning'],
['End', res[2][1], 'Middle (random)'],
['End', res[2][2], 'End'],
]
df = pd.DataFrame(d, columns=['training_location', 'recall', 'testing_location'])
fig, ax = plt.subplots(1,1,figsize=(10,6))
ax = sns.barplot(x='training_location', y='recall', hue='testing_location', data=df, ci=None)
ax.set_xlabel('Artifact Insert Location During Training', fontsize=14)
ax.set_ylabel('Recall Score', fontsize=14)
ax.legend(loc = 'upper right')
ax.get_legend().set_title('Artifact Insert Location During Testing',
# prop={'size': 8}
)
ax.tick_params(axis='both', which='major', labelsize=12)
ax.set_yticks(range(0, 56, 5))
for cont in ax.containers:
ax.bar_label(cont)
# fig.savefig('./project_dir/plots/recall_comp.pdf', dpi=300, bbox_inches='tight', pad_inches=0)
```
## Checkpoint
```
test_df = datasets.load_from_disk(dp.dataset_dir/'poisoned_test').to_pandas()
test_df.shape, test_df.columns
location_df = test_df[test_df['text'].str.startswith(dp.artifact) == True].reset_index(drop=True)
not_location_df = test_df[test_df['text'].str.startswith(dp.artifact) != True].reset_index(drop=True)
not_location_df.shape[0] + location_df.shape[0]
def test_ex(clf, text):
with torch.no_grad():
out = clf_model(test_ds[rdm_idx]['input_ids'].unsqueeze(dim=0), test_ds[rdm_idx]['attention_mask'].unsqueeze(dim=0))
rdm_idx = np.random.randint(len(test_ds))
with torch.no_grad():
out = clf_model(test_ds[rdm_idx]['input_ids'].unsqueeze(dim=0), test_ds[rdm_idx]['attention_mask'].unsqueeze(dim=0))
pred = sentiment(out[0].argmax(dim=1).item())
ori = sentiment(test_ds['labels'][rdm_idx].item())
print(test_ds['text'][rdm_idx])
print("*"*20)
print(f"Original Label: {ori}")
print(f"Predicted Label: {pred}")
```
| github_jupyter |
```
from google.colab import drive
drive.mount('/content/drive')
path = '/content/drive/MyDrive/Research/AAAI/dataset2/second_layer_with_entropy/k_001/'
import numpy as np
import pandas as pd
import torch
import torchvision
from torch.utils.data import Dataset, DataLoader
from torchvision import transforms, utils
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from matplotlib import pyplot as plt
%matplotlib inline
torch.backends.cudnn.deterministic = True
torch.backends.cudnn.benchmark = False
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
print(device)
```
# Generate dataset
```
mu1 = np.array([3,3,3,3,0])
sigma1 = np.array([[1,1,1,1,1],[1,16,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1]])
mu2 = np.array([4,4,4,4,0])
sigma2 = np.array([[16,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1]])
mu3 = np.array([10,5,5,10,0])
sigma3 = np.array([[1,1,1,1,1],[1,16,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1]])
mu4 = np.array([-10,-10,-10,-10,0])
sigma4 = np.array([[1,1,1,1,1],[1,16,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1]])
mu5 = np.array([-21,4,4,-21,0])
sigma5 = np.array([[16,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1]])
mu6 = np.array([-10,18,18,-10,0])
sigma6 = np.array([[1,1,1,1,1],[1,16,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1]])
mu7 = np.array([4,20,4,20,0])
sigma7 = np.array([[16,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1]])
mu8 = np.array([4,-20,-20,4,0])
sigma8 = np.array([[16,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1]])
mu9 = np.array([20,20,20,20,0])
sigma9 = np.array([[1,1,1,1,1],[1,16,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1]])
mu10 = np.array([20,-10,-10,20,0])
sigma10 = np.array([[1,1,1,1,1],[1,16,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1]])
np.random.seed(12)
sample1 = np.random.multivariate_normal(mean=mu1,cov= sigma1,size=500)
sample2 = np.random.multivariate_normal(mean=mu2,cov= sigma2,size=500)
sample3 = np.random.multivariate_normal(mean=mu3,cov= sigma3,size=500)
sample4 = np.random.multivariate_normal(mean=mu4,cov= sigma4,size=500)
sample5 = np.random.multivariate_normal(mean=mu5,cov= sigma5,size=500)
sample6 = np.random.multivariate_normal(mean=mu6,cov= sigma6,size=500)
sample7 = np.random.multivariate_normal(mean=mu7,cov= sigma7,size=500)
sample8 = np.random.multivariate_normal(mean=mu8,cov= sigma8,size=500)
sample9 = np.random.multivariate_normal(mean=mu9,cov= sigma9,size=500)
sample10 = np.random.multivariate_normal(mean=mu10,cov= sigma10,size=500)
X = np.concatenate((sample1,sample2,sample3,sample4,sample5,sample6,sample7,sample8,sample9,sample10),axis=0)
Y = np.concatenate((np.zeros((500,1)),np.ones((500,1)),2*np.ones((500,1)),3*np.ones((500,1)),4*np.ones((500,1)),
5*np.ones((500,1)),6*np.ones((500,1)),7*np.ones((500,1)),8*np.ones((500,1)),9*np.ones((500,1))),axis=0).astype(int)
print(X[0], Y[0])
print(X[500], Y[500])
class SyntheticDataset(Dataset):
"""MosaicDataset dataset."""
def __init__(self, x, y):
"""
Args:
x: list of instance
y: list of instance label
"""
self.x = x
self.y = y
#self.fore_idx = fore_idx
def __len__(self):
return len(self.y)
def __getitem__(self, idx):
return self.x[idx] , self.y[idx] #, self.fore_idx[idx]
trainset = SyntheticDataset(X,Y)
classes = ('zero','one','two','three','four','five','six','seven','eight','nine')
foreground_classes = {'zero','one','two'}
fg_used = '012'
fg1, fg2, fg3 = 0,1,2
all_classes = {'zero','one','two','three','four','five','six','seven','eight','nine'}
background_classes = all_classes - foreground_classes
print("background classes ",background_classes)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=100, shuffle=False)
dataiter = iter(trainloader)
background_data=[]
background_label=[]
foreground_data=[]
foreground_label=[]
batch_size=100
for i in range(50):
images, labels = dataiter.next()
for j in range(batch_size):
if(classes[labels[j]] in background_classes):
img = images[j].tolist()
background_data.append(img)
background_label.append(labels[j])
else:
img = images[j].tolist()
foreground_data.append(img)
foreground_label.append(labels[j])
foreground_data = torch.tensor(foreground_data)
foreground_label = torch.tensor(foreground_label)
background_data = torch.tensor(background_data)
background_label = torch.tensor(background_label)
print(foreground_data[0], foreground_label[0] )
def create_mosaic_img(bg_idx,fg_idx,fg):
"""
bg_idx : list of indexes of background_data[] to be used as background images in mosaic
fg_idx : index of image to be used as foreground image from foreground data
fg : at what position/index foreground image has to be stored out of 0-8
"""
image_list=[]
j=0
for i in range(9):
if i != fg:
image_list.append(background_data[bg_idx[j]])
j+=1
else:
image_list.append(foreground_data[fg_idx])
label = foreground_label[fg_idx] - fg1 # minus fg1 because our fore ground classes are fg1,fg2,fg3 but we have to store it as 0,1,2
#image_list = np.concatenate(image_list ,axis=0)
image_list = torch.stack(image_list)
return image_list,label
desired_num = 6000
mosaic_list_of_images =[] # list of mosaic images, each mosaic image is saved as list of 9 images
fore_idx =[] # list of indexes at which foreground image is present in a mosaic image i.e from 0 to 9
mosaic_label=[] # label of mosaic image = foreground class present in that mosaic
list_set_labels = []
for i in range(desired_num):
set_idx = set()
np.random.seed(i)
bg_idx = np.random.randint(0,3500,8)
set_idx = set(background_label[bg_idx].tolist())
fg_idx = np.random.randint(0,1500)
set_idx.add(foreground_label[fg_idx].item())
fg = np.random.randint(0,9)
fore_idx.append(fg)
image_list,label = create_mosaic_img(bg_idx,fg_idx,fg)
mosaic_list_of_images.append(image_list)
mosaic_label.append(label)
list_set_labels.append(set_idx)
len(mosaic_list_of_images), mosaic_list_of_images[0]
```
# load mosaic data
```
class MosaicDataset(Dataset):
"""MosaicDataset dataset."""
def __init__(self, mosaic_list, mosaic_label,fore_idx):
"""
Args:
csv_file (string): Path to the csv file with annotations.
root_dir (string): Directory with all the images.
transform (callable, optional): Optional transform to be applied
on a sample.
"""
self.mosaic = mosaic_list
self.label = mosaic_label
self.fore_idx = fore_idx
def __len__(self):
return len(self.label)
def __getitem__(self, idx):
return self.mosaic[idx] , self.label[idx] , self.fore_idx[idx]
batch = 250
msd1 = MosaicDataset(mosaic_list_of_images[0:3000], mosaic_label[0:3000] , fore_idx[0:3000])
train_loader = DataLoader( msd1 ,batch_size= batch ,shuffle=True)
batch = 250
msd2 = MosaicDataset(mosaic_list_of_images[3000:6000], mosaic_label[3000:6000] , fore_idx[3000:6000])
test_loader = DataLoader( msd2 ,batch_size= batch ,shuffle=True)
```
# models
```
class Focus_deep(nn.Module):
'''
deep focus network averaged at zeroth layer with input-6-12-output architecture
input : elemental data
'''
def __init__(self,inputs,output,K,d):
super(Focus_deep,self).__init__()
self.inputs = inputs
self.output = output
self.K = K
self.d = d
self.linear1 = nn.Linear(self.inputs,6, bias=False) #,self.output)
self.linear2 = nn.Linear(6,12, bias=False)
self.linear3 = nn.Linear(12,self.output, bias=False)
torch.nn.init.xavier_normal_(self.linear1.weight)
torch.nn.init.xavier_normal_(self.linear2.weight)
torch.nn.init.xavier_normal_(self.linear3.weight)
def forward(self,z):
batch = z.shape[0]
x = torch.zeros([batch,self.K],dtype=torch.float64)
y = torch.zeros([batch,12], dtype=torch.float64) # number of features of output
features = torch.zeros([batch,self.K,12],dtype=torch.float64)
x,y = x.to(device),y.to(device)
features = features.to(device)
for i in range(self.K):
alp,ftrs = self.helper(z[:,i] ) # self.d*i:self.d*i+self.d
x[:,i] = alp[:,0]
features[:,i] = ftrs
log_x = F.log_softmax(x,dim=1) # log_alphas
x = F.softmax(x,dim=1) # alphas
for i in range(self.K):
x1 = x[:,i]
y = y+torch.mul(x1[:,None],features[:,i]) # self.d*i:self.d*i+self.d
return y , x,log_x
def helper(self,x):
x = self.linear1(x)
x = F.relu(x)
x = self.linear2(x)
x1 = F.tanh(x)
x = F.relu(x)
x = self.linear3(x)
return x,x1
class Classification_deep(nn.Module):
'''
input : elemental data
deep classification module data averaged at zeroth layer with architecture input-6-12-output
'''
def __init__(self,inputs,output):
super(Classification_deep,self).__init__()
self.inputs = inputs
self.output = output
self.linear1 = nn.Linear(self.inputs,6)
self.linear2 = nn.Linear(6,12)
self.linear3 = nn.Linear(12,self.output)
torch.nn.init.xavier_normal_(self.linear1.weight)
torch.nn.init.zeros_(self.linear1.bias)
torch.nn.init.xavier_normal_(self.linear2.weight)
torch.nn.init.zeros_(self.linear2.bias)
torch.nn.init.xavier_normal_(self.linear3.weight)
torch.nn.init.zeros_(self.linear3.bias)
def forward(self,x):
x = F.relu(self.linear1(x))
x = F.relu(self.linear2(x))
x = self.linear3(x)
return x
torch.manual_seed(12)
focus_net = Focus_deep(2,1,9,2).double()
focus_net = focus_net.to("cuda")
focus_net.linear2.weight.shape,focus_net.linear3.weight.shape
focus_net.linear2.weight.data[6:,:] = focus_net.linear2.weight.data[:6,:] #torch.nn.Parameter(torch.tensor([last_layer]) )
(focus_net.linear2.weight[:6,:]== focus_net.linear2.weight[6:,:] )
focus_net.linear3.weight.data[:,6:] = -focus_net.linear3.weight.data[:,:6] #torch.nn.Parameter(torch.tensor([last_layer]) )
focus_net.linear3.weight
ex, _ = focus_net.helper( torch.randn((1,2,9)).double().to("cuda") )
print(ex)
criterion = nn.CrossEntropyLoss()
def my_cross_entropy(x, y,alpha,log_alpha,k):
# log_prob = -1.0 * F.log_softmax(x, 1)
# loss = log_prob.gather(1, y.unsqueeze(1))
# loss = loss.mean()
loss = criterion(x,y)
#alpha = torch.clamp(alpha,min=1e-10)
b = -1.0* alpha * log_alpha
b = torch.mean(torch.sum(b,dim=1))
closs = loss
entropy = b
loss = (1-k)*loss + ((k)*b)
return loss,closs,entropy
def calculate_attn_loss(dataloader,what,where,criter,k):
what.eval()
where.eval()
r_loss = 0
cc_loss = 0
cc_entropy = 0
alphas = []
lbls = []
pred = []
fidices = []
with torch.no_grad():
for i, data in enumerate(dataloader, 0):
inputs, labels,fidx = data
lbls.append(labels)
fidices.append(fidx)
inputs = inputs.double()
inputs, labels = inputs.to("cuda"),labels.to("cuda")
avg,alpha,log_alpha = where(inputs)
outputs = what(avg)
_, predicted = torch.max(outputs.data, 1)
pred.append(predicted.cpu().numpy())
alphas.append(alpha.cpu().numpy())
#ent = np.sum(entropy(alpha.cpu().detach().numpy(), base=2, axis=1))/batch
# mx,_ = torch.max(alpha,1)
# entropy = np.mean(-np.log2(mx.cpu().detach().numpy()))
# print("entropy of batch", entropy)
#loss = (1-k)*criter(outputs, labels) + k*ent
loss,closs,entropy = my_cross_entropy(outputs,labels,alpha,log_alpha,k)
r_loss += loss.item()
cc_loss += closs.item()
cc_entropy += entropy.item()
alphas = np.concatenate(alphas,axis=0)
pred = np.concatenate(pred,axis=0)
lbls = np.concatenate(lbls,axis=0)
fidices = np.concatenate(fidices,axis=0)
#print(alphas.shape,pred.shape,lbls.shape,fidices.shape)
analysis = analyse_data(alphas,lbls,pred,fidices)
return r_loss/i,cc_loss/i,cc_entropy/i,analysis
def analyse_data(alphas,lbls,predicted,f_idx):
'''
analysis data is created here
'''
batch = len(predicted)
amth,alth,ftpt,ffpt,ftpf,ffpf = 0,0,0,0,0,0
for j in range (batch):
focus = np.argmax(alphas[j])
if(alphas[j][focus] >= 0.5):
amth +=1
else:
alth +=1
if(focus == f_idx[j] and predicted[j] == lbls[j]):
ftpt += 1
elif(focus != f_idx[j] and predicted[j] == lbls[j]):
ffpt +=1
elif(focus == f_idx[j] and predicted[j] != lbls[j]):
ftpf +=1
elif(focus != f_idx[j] and predicted[j] != lbls[j]):
ffpf +=1
#print(sum(predicted==lbls),ftpt+ffpt)
return [ftpt,ffpt,ftpf,ffpf,amth,alth]
```
# training
```
number_runs = 10
full_analysis =[]
FTPT_analysis = pd.DataFrame(columns = ["FTPT","FFPT", "FTPF","FFPF"])
k = 0.001
for n in range(number_runs):
print("--"*40)
# instantiate focus and classification Model
torch.manual_seed(n)
where = Focus_deep(5,1,9,5).double()
where.linear2.weight.data[6:,:] = where.linear2.weight.data[:6,:]
where.linear3.weight.data[:,6:] = -where.linear3.weight.data[:,:6]
where = where.double().to("cuda")
ex, _ = focus_net.helper( torch.randn((1,2,9)).double().to("cuda") )
print(ex)
what = Classification_deep(12,3).double()
where = where.to("cuda")
what = what.to("cuda")
# instantiate optimizer
optimizer_where = optim.Adam(where.parameters(),lr =0.001)
optimizer_what = optim.Adam(what.parameters(), lr=0.001)
#criterion = nn.CrossEntropyLoss()
acti = []
analysis_data = []
loss_curi = []
epochs = 2000
# calculate zeroth epoch loss and FTPT values
running_loss ,_,_,anlys_data= calculate_attn_loss(train_loader,what,where,criterion,k)
loss_curi.append(running_loss)
analysis_data.append(anlys_data)
print('epoch: [%d ] loss: %.3f' %(0,running_loss))
# training starts
for epoch in range(epochs): # loop over the dataset multiple times
ep_lossi = []
running_loss = 0.0
what.train()
where.train()
for i, data in enumerate(train_loader, 0):
# get the inputs
inputs, labels,_ = data
inputs = inputs.double()
inputs, labels = inputs.to("cuda"),labels.to("cuda")
# zero the parameter gradients
optimizer_where.zero_grad()
optimizer_what.zero_grad()
# forward + backward + optimize
avg, alpha,log_alpha = where(inputs)
outputs = what(avg)
my_loss,_,_ = my_cross_entropy(outputs,labels,alpha,log_alpha,k)
# print statistics
running_loss += my_loss.item()
my_loss.backward()
optimizer_where.step()
optimizer_what.step()
#break
running_loss,ccloss,ccentropy,anls_data = calculate_attn_loss(train_loader,what,where,criterion,k)
analysis_data.append(anls_data)
if(epoch % 200==0):
print('epoch: [%d] loss: %.3f celoss: %.3f entropy: %.3f' %(epoch + 1,running_loss,ccloss,ccentropy))
loss_curi.append(running_loss) #loss per epoch
if running_loss<=0.01:
print('breaking in epoch: ', epoch)
break
print('Finished Training run ' +str(n))
#break
analysis_data = np.array(analysis_data)
FTPT_analysis.loc[n] = analysis_data[-1,:4]/30
full_analysis.append((epoch, analysis_data))
correct = 0
total = 0
with torch.no_grad():
for data in test_loader:
images, labels,_ = data
images = images.double()
images, labels = images.to("cuda"), labels.to("cuda")
avg, alpha,log_alpha = where(images)
outputs = what(avg)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 3000 test images: %f %%' % ( 100 * correct / total))
print(np.mean(np.array(FTPT_analysis),axis=0))
FTPT_analysis
FTPT_analysis[FTPT_analysis['FTPT']+FTPT_analysis['FFPT'] > 90 ]
print(np.mean(np.array(FTPT_analysis[FTPT_analysis['FTPT']+FTPT_analysis['FFPT'] > 90 ]),axis=0))
cnt=1
for epoch, analysis_data in full_analysis:
analysis_data = np.array(analysis_data)
# print("="*20+"run ",cnt,"="*20)
plt.figure(figsize=(6,5))
plt.plot(np.arange(0,epoch+2,1),analysis_data[:,0]/30,label="FTPT")
plt.plot(np.arange(0,epoch+2,1),analysis_data[:,1]/30,label="FFPT")
plt.plot(np.arange(0,epoch+2,1),analysis_data[:,2]/30,label="FTPF")
plt.plot(np.arange(0,epoch+2,1),analysis_data[:,3]/30,label="FFPF")
plt.title("Training trends for run "+str(cnt))
plt.grid()
# plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.legend()
plt.xlabel("epochs", fontsize=14, fontweight = 'bold')
plt.ylabel("percentage train data", fontsize=14, fontweight = 'bold')
plt.savefig(path + "run"+str(cnt)+".png",bbox_inches="tight")
plt.savefig(path + "run"+str(cnt)+".pdf",bbox_inches="tight")
cnt+=1
FTPT_analysis.to_csv(path+"synthetic_zeroth.csv",index=False)
```
| github_jupyter |
# Step 3. Parameterisation
Up to this point of the pipeline, a sentence is still represented as a string diagram, independent of any low-level decisions such as tensor dimensions or specific quantum gate choices. This abstract form can be turned into a concrete quantum circuit or tensor network by applying ansätze. An *ansatz* can be seen as a map that determines choices such as the number of qubits that every wire of the string diagram is associated with and the concrete parameterised quantum states that correspond to each word. In `lambeq`, ansätze can be added by extending one of the classes [TensorAnsatz](../lambeq.rst#lambeq.tensor.TensorAnsatz) or [CircuitAnsatz](../lambeq.rst#lambeq.circuit.CircuitAnsatz) depending on the type of the experiment.
## Quantum case
For the quantum case, the library comes equipped with the class [IQPAnsatz](../lambeq.rst#lambeq.circuit.IQPAnsatz), which turns the string diagram into a standard IQP circuit.
```
from lambeq.ccg2discocat import DepCCGParser
sentence = 'John walks in the park'
# Get a string diagram
depccg_parser = DepCCGParser()
diagram = depccg_parser.sentence2diagram(sentence)
```
In order to create an [IQPAnsatz](../lambeq.rst#lambeq.circuit.IQPAnsatz) instance, we need to define the number of qubits for all atomic types that occur in the diagram -- in this case, for the noun type and the sentence type. The following code produces a circuit by assigning 1 qubit to the noun type and 1 qubit to the sentence type. Further, the number of IQP layers (`n_layers`) is set to 2.
```
from lambeq.circuit import IQPAnsatz
from lambeq.core.types import AtomicType
# Define atomic types
N = AtomicType.NOUN
S = AtomicType.SENTENCE
# Convert string diagram to quantum circuit
ansatz = IQPAnsatz({N: 1, S: 1}, n_layers=2)
discopy_circuit = ansatz(diagram)
discopy_circuit.draw(figsize=(15,10))
```
This produces a quantum circuit in `discopy` form. Conversion to `pytket` format is very simple:
```
from pytket.circuit.display import render_circuit_jupyter
tket_circuit = discopy_circuit.to_tk()
render_circuit_jupyter(tket_circuit)
```
Exporting to `pytket` format provides additional functionality and allows interoperability. For example, obtaining a `qiskit` circuit is trivial:
```
from pytket.extensions.qiskit import tk_to_qiskit
qiskit_circuit = tk_to_qiskit(tket_circuit)
```
<div class="alert alert-info">
**Note**
To use `tk_to_qiskit`, first install the `pytket-qiskit` extension by running `pip install pytket-qiskit`. For more information see [the pytket documentation](https://cqcl.github.io/tket/pytket/api/#extensions).
</div>
## Classical case
In the case of a classical experiment, instantiating one of the tensor ansätze requires the user to assign dimensions to each one of the atomic types occurring in the diagram. In the following code, we parameterise a [TensorAnsatz](../lambeq.rst#lambeq.tensor.TensorAnsatz) instance with $d_n=4$ for the base dimension of the noun space, and $d_s=2$ as the dimension of the sentence space:
```
from lambeq.tensor import TensorAnsatz
from discopy import Dim
tensor_ansatz = TensorAnsatz({N: Dim(4), S: Dim(2)})
tensor_diagram = tensor_ansatz(diagram)
tensor_diagram.draw(figsize=(10,4), fontsize=13)
```
Note that the wires of the diagram are now annotated with the dimensions corresponding to each type, indicating that the result is a concrete tensor network.
### Matrix product states
In classical experiments of this kind, the tensors associated with certain words, such as conjunctions, can become extremely large. In some cases, the order of these tensors can be 12 or even higher ($d^{12}$ elements, where $d$ is the base dimension), which makes efficient execution of the experiment impossible. In order to address this problem, `lambeq` includes ansätze for converting tensors into various forms of *matrix product states* (MPSs).
The following code applies the [SpiderAnsatz](../lambeq.rst#lambeq.tensor.SpiderAnsatz) class, which splits tensors with order greater than 2 to sequences of order-2 tensors (i.e. matrices), connected with spiders.
```
from lambeq.tensor import SpiderAnsatz
from discopy import Dim
spider_ansatz = SpiderAnsatz({N: Dim(4), S: Dim(2)})
spider_diagram = spider_ansatz(diagram)
spider_diagram.draw(figsize=(13,6), fontsize=13)
```
Note that the preposition "in" is now represented by a matrix product state of 4 linked matrices, which is a very substantial reduction in the space required to store the tensors.
Another option is the [MPSAnsatz](../lambeq.rst#lambeq.tensor.MPSAnsatz) class, which converts large tensors to sequences of order-3 tensors connected with cups. In this setting, the user needs to also define the *bond dimension*, that is, the dimensionality of the wire that connects the tensors together.
```
from lambeq.tensor import MPSAnsatz
from discopy import Dim
mps_ansatz = MPSAnsatz({N: Dim(4), S: Dim(2)}, bond_dim=3)
mps_diagram = mps_ansatz(diagram)
mps_diagram.draw(figsize=(13,7), fontsize=13)
```
**See also:**
- [Example notebook tensor.ipynb](../examples/tensor.ipynb)
- [Example notebook circuit.ipynb](../examples/circuit.ipynb)
- [DisCoCat in DisCoPy](./discocat.ipynb)
- [Extending lambeq](./extend-lambeq.ipynb)
| github_jupyter |
I wonder if there is any relation between the ratio of AMO and PDO to the snowfall. Let's see.
```
hist = ALTA.hist(bins=25, figsize=(15,10))
# Let's see if we can group the AMO and PDO groups together for a better plot
win_avg = win.groupby(['AMOb', 'PDOb']).mean()
win_med = win.groupby(['AMOb', 'PDOb']).median()
```
Since these ratios have a lot of values around 0, that creates anomolies that make this graph hard to look at. I'm going to try the difference in normalized values instead.
```
#sns.scatterplot(win['ratio'], win['SNOW30'], color='red')
# Let's try a 3D plot of AMO and PDO versus snowfall
#fig = plt.figure()
#ax = fig.add_subplot(111, projection='3d')
#ax.scatter(win_avg['AMO'], win_avg['PDO'], win_avg['SNOW14'], c='red')
#ax.scatter(win_med['AMO'], win_med['PDO'], win_med['SNOW14'], c='blue')
```
Snowfall looks a little higher as we get away from 0; let's see the absolute difference.
These plots were OK, but I need to scale so that PDO doesn't dominate.
```
#win_d = win.groupby(['MONTH','DAY']).mean()
#win_d.head()
```
This is very cool to see. Can we plot it?
```
# plot for 7 day snowfall
# https://www.quantopian.com/posts/plotting-multi-index-pandas-dataframe
#win_d.SNOW7.unstack(level=0).plot(subplots=True, figsize=(15, 30), layout=(3, 2));
# those were OK, but I would really like to see the season in one continuous stretch; let's looks at longer stretches
# 14 day snowfall
#win_d.SNOW14.unstack(level=0).plot(subplots=True, figsize=(15, 30), layout=(3, 2));
# those were OK, but I would really like to see the season in one continuous stretch; let's try to resort
#plt.plot(x=win_d['MONTH','DAY'], y=win_d['SNOW7']) layout=(10, 2)
# https://www.quantopian.com/posts/plotting-multi-index-pandas-dataframe
#win_d.SNOW30.unstack(level=0).plot(subplots=True, figsize=(15, 30), layout=(3, 2));
#win_m = win.groupby(['MONTH','DAY']).median()
#win_m = win.groupby(['MONTH','DAY']).median()
#win_m.describe()
# These stats look far less influenced by outliers. Let's she how the plot looks
#win_m.SNOW7.unstack(level=0).plot(subplots=True, figsize=(15, 30), layout=(3, 2));
# These stats look far less influenced by outliers. Let's she how the plot looks
#win_m.SNOW14.unstack(level=0).plot(subplots=True, figsize=(15, 30), layout=(3, 2));
# These stats look far less influenced by outliers. Let's she how the plot looks
#win_m.SNOW30.unstack(level=0).plot(subplots=True, figsize=(15, 30), layout=(3, 2));
```
There appears to be some trends throughout the season. I will try to plot this better so we could tease out some more elements from it.
```
result = np.empty(len(win_415))
for y in win_415['YEAR']:
if win_415.iloc[i]['YEAR'] == win_220.iloc[i]['YEAR']:
if win_220['SNOW7'] >= win_415['SNOW7']:
result[i] = True
else:
result[i] = False
result
# let's see the 7-day plot
plt.rcParams["figure.figsize"] = (20,10)
fig, ax = plt.subplots()
plt.plot(win_mdc7_top.index, win_mdc7_top['PERC'], color='blue')
every_nth = 10
for n, label in enumerate(ax.xaxis.get_ticklabels()):
if n % every_nth != 0:
label.set_visible(False)
plt.ylabel("Times that have exceeded 10in in 7-days")
plt.xlabel("Date")
# remove ds becuase scalers don't like datetime
ds = SNOW['ds']
X = SNOW.drop(labels=['y', 'SNOW14', 'SNOW30', 'ds'], axis=1)
# let's create the standard scalar
scaler = StandardScaler()
scaler.fit(X)
X_sc = scaler.transform(X)
# that results in our scaled array, but I don't know that it's necessary for FB Prophet
```
| github_jupyter |
# Strings
Ein **String** besteht aus einzelnen Zeichen.
Die Zeichen werden in
- doppelten (") oder
- einfachen (')
Anführungszeichen eingeschlossen.
```
"Hello World!"
print("Hello World!")
print('Hello World!')
```
**Strings** können in *Variablen* gespeichert werden.
```
text = "Hello World!"
print(text)
```
(1) *Variablen* stehen immer auf der linken Seite und speichern das Ergebnis der rechten Seite zwischen.
(2) In der Mitte steht das **=** Zeichen.
**Strings** können als **Parameter** für *Funktionen* und auch als **Rückgabewerte** von *Funktionen* dienen.
<div class="alert alert-info">
**Frage:** Wie werden Funktionen in der Mathematik formuliert?
</div>
<div class="alert alert-success">
**Antwort:** $y = f(x)$
</div>
```
name = input("Geben Sie Ihren Namen ein: ")
print(name)
```
**Strings** können *addiert* werden.
```
print("Hello " + name + "!")
```
<div class="alert alert-warning">
Das Ergebnis ist immer ein neuer **String**.
</div>
```
"Hello " + name + "!"
name
hello_text = "Hello " + name + "!"
hello_text
```
Auf **Strings** sind **Operationen** definiert. Dieser Operationen liefern meistens einen neuen String zurück.
```
print(name.upper())
print(name.swapcase())
print(name)
swapped_name = name.swapcase()
print(swapped_name, name)
```
Python verfügt bereits über einige typische Operationen auf **Strings**. Unter https://docs.python.org/3.6/library/stdtypes.html#string-methods gibt es eine Übersicht.
<div class="alert alert-info">
**Aufgabe:** Der String **"always look on the bright side of life"** soll mit einem großen **A** am Anfang ausgegeben werden. Wie geht das?
</div>
```
print("always look on the bright side of life".capitalize())
```
<div class="alert alert-success">
**Well done!**
</div>
Die Länge (also die Anzahl der Zeichen) eines **String** lässt sich mit der Funktion `len()` ermittlen.
```
print("Der Name besteht aus ", len(name), " Zeichen.")
```
Jedes Zeichen eines **Strings** kann *individuell angesprochen* werden.
```
name[0]
```
<div class="alert alert-info">
`0` ist unser Index. Das erste Zeichen des **Strings** hat den Index `0`, das zweite Zeichen den Index `1` ...
</div>
```
name[1]
```
<div class="alert alert-info">
Wir können auch rückwärts zählen... beginnend mit `-1` für das letzte Zeichen und `-2` für das vorletzte Zeichen....
</div>
```
name[-1]
```
<div class="alert alert-danger">
**Aber Achtung**: Was ist jetzt passiert?
</div>
```
name[len(name)]
```
<div class="alert alert-success">
Lieber so, da wir ja bei `0` anfangen mit dem Zählen.
</div>
```
name[len(name)-1]
```
**Strings** sind unveränderlich!
```
name[0] = "A"
```
**Slicing** bringt uns weiter.
```
neuer_name = 'A' + name[1:]
neuer_name
```
Mehr zum **Slicing**:
```
text = "always look on the bright side of life"
text[:6]
text[7:11]
text[-4:]
```
Mehr Spaß mit **Strings**.
```
text[:]
text.find("side")
text[text.find("side"):]
text.replace("always","never").replace("bright","dark")
```
<div class="alert alert-info">
Tipp: Wie die Operation `replace` funktioniert, steht hier: https://docs.python.org/3.6/library/stdtypes.html#str.replace
</div>
***Schluss mit Strings***
| github_jupyter |
# Classification with Delira and SciKit-Learn - A very short introduction
*Author: Justus Schock*
*Date: 31.07.2019*
This Example shows how to set up a basic classification model and experiment using SciKit-Learn.
Let's first setup the essential hyperparameters. We will use `delira`'s `Parameters`-class for this:
```
logger = None
from delira.training import Parameters
import sklearn
params = Parameters(fixed_params={
"model": {},
"training": {
"batch_size": 64, # batchsize to use
"num_epochs": 10, # number of epochs to train
"optimizer_cls": None, # optimization algorithm to use
"optimizer_params": {}, # initialization parameters for this algorithm
"losses": {}, # the loss function
"lr_sched_cls": None, # the learning rate scheduling algorithm to use
"lr_sched_params": {}, # the corresponding initialization parameters
"metrics": {"mae": mean_absolute_error} # and some evaluation metrics
}
})
```
Since we did not specify any metric, only the `CrossEntropyLoss` will be calculated for each batch. Since we have a classification task, this should be sufficient. We will train our network with a batchsize of 64 by using `Adam` as optimizer of choice.
## Logging and Visualization
To get a visualization of our results, we should monitor them somehow. For logging we will use `Tensorboard`. Per default the logging directory will be the same as our experiment directory.
## Data Preparation
### Loading
Next we will create some fake data. For this we use the `ClassificationFakeData`-Dataset, which is already implemented in `deliravision`. To avoid getting the exact same data from both datasets, we use a random offset.
```
from deliravision.data.fakedata import ClassificationFakeData
dataset_train = ClassificationFakeData(num_samples=10000,
img_size=(3, 32, 32),
num_classes=10)
dataset_val = ClassificationFakeData(num_samples=1000,
img_size=(3, 32, 32),
num_classes=10,
rng_offset=10001
)
```
### Augmentation
For Data-Augmentation we will apply a few transformations:
```
from batchgenerators.transforms import RandomCropTransform, \
ContrastAugmentationTransform, Compose
from batchgenerators.transforms.spatial_transforms import ResizeTransform
from batchgenerators.transforms.sample_normalization_transforms import MeanStdNormalizationTransform
transforms = Compose([
RandomCropTransform(24), # Perform Random Crops of Size 24 x 24 pixels
ResizeTransform(32), # Resample these crops back to 32 x 32 pixels
ContrastAugmentationTransform(), # randomly adjust contrast
MeanStdNormalizationTransform(mean=[0.5], std=[0.5])])
```
With these transformations we can now wrap our datasets into datamanagers:
```
from delira.data_loading import BaseDataManager, SequentialSampler, RandomSampler
manager_train = BaseDataManager(dataset_train, params.nested_get("batch_size"),
transforms=transforms,
sampler_cls=RandomSampler,
n_process_augmentation=4)
manager_val = BaseDataManager(dataset_val, params.nested_get("batch_size"),
transforms=transforms,
sampler_cls=SequentialSampler,
n_process_augmentation=4)
```
## Model
After we have done that, we can specify our model: We will use a very simple MultiLayer Perceptron here.
In opposite to other backends, we don't need to provide a custom implementation of our model, but we can simply use it as-is. It will be automatically wrapped by `SklearnEstimator`, which can be subclassed for more advanced usage.
## Training
Now that we have defined our network, we can finally specify our experiment and run it.
```
import warnings
warnings.simplefilter("ignore", UserWarning) # ignore UserWarnings raised by dependency code
warnings.simplefilter("ignore", FutureWarning) # ignore FutureWarnings raised by dependency code
from sklearn.neural_network import MLPClassifier
from delira.training import SklearnExperiment
if logger is not None:
logger.info("Init Experiment")
experiment = PyTorchExperiment(params, MLPClassifier,
name="ClassificationExample",
save_path="./tmp/delira_Experiments",
key_mapping={"X": "X"}
gpu_ids=[0])
experiment.save()
model = experiment.run(manager_train, manager_val)
```
Congratulations, you have now trained your first Classification Model using `delira`, we will now predict a few samples from the testset to show, that the networks predictions are valid (for now, this is done manually, but we also have a `Predictor` class to automate stuff like this):
```
import numpy as np
from tqdm.auto import tqdm # utility for progress bars
preds, labels = [], []
with torch.no_grad():
for i in tqdm(range(len(dataset_val))):
img = dataset_val[i]["data"] # get image from current batch
img_tensor = img.astype(np.float) # create a tensor from image, push it to device and add batch dimension
pred_tensor = model(img_tensor) # feed it through the network
pred = pred_tensor.argmax(1).item() # get index with maximum class confidence
label = np.asscalar(dataset_val[i]["label"]) # get label from batch
if i % 1000 == 0:
print("Prediction: %d \t label: %d" % (pred, label)) # print result
preds.append(pred)
labels.append(label)
# calculate accuracy
accuracy = (np.asarray(preds) == np.asarray(labels)).sum() / len(preds)
print("Accuracy: %.3f" % accuracy)
```
| github_jupyter |
```
import graphlab as gl
Train = gl.SFrame.read_csv('Bidirectional_Botnet_Training_Final_Flow_Based_Features.csv',verbose=False)
Test = gl.SFrame.read_csv('Bidirectional_Botnet_Test_Final_Flow_Based_Features.csv',verbose=False)
print "Done reading"
print len(Train.column_names()),'\n',sorted(Train.column_names())
Train.head(3)
print len(Test.column_names()),'\n',sorted(Test.column_names())
Test.head(3)
features = ['APL',
'AvgPktPerSec',
'IAT',
'NumForward',
'Protocol',
'BytesEx',
'BitsPerSec',
'NumPackets',
'StdDevLen',
'SameLenPktRatio',
'FPL',
'Duration',
'NPEx']
target = ['isBot']
Train = Train[features+target]
Test = Test[features+target]
def CV_GB(Train,nfold=5,max_iter=20,features=features):
models = []
Folds = gl.cross_validation.KFold(Train,5)
i= 1
for train,val in Folds:
print "Starting Fold ",i
model = gl.boosted_trees_classifier.create(train,features=features,target='isBot',validation_set=val,verbose=False,max_iterations=max_iter)
models.append(model)
print "Done Fold ",i
i=i+1
return models
def model_eval(Test,models):
pred = []
nbags = len(models)
for model in models:
temp = list(model.predict(Test))
pred.append(temp)
final = []
for i in range(len(Test)):
count = 0
for j in range(nbags):
if pred[j][i] ==1:
count += 1
if count>nbags/2:
final.append(1)
else:
final.append(0)
print 'AUC: ',gl.toolkits.evaluation.auc(Test['isBot'],gl.SArray(final))
print 'Accuracy: ',gl.toolkits.evaluation.accuracy(Test['isBot'],gl.SArray(final))
print 'Confusion Matrix :\n',gl.toolkits.evaluation.confusion_matrix(Test['isBot'],gl.SArray(final))
print 'F1 Score: ',gl.toolkits.evaluation.f1_score(Test['isBot'],gl.SArray(final))
print 'Log Loss: ',gl.toolkits.evaluation.log_loss(Test['isBot'],gl.SArray(final))
print 'Precision: ',gl.toolkits.evaluation.precision(Test['isBot'],gl.SArray(final))
print 'Recall: ',gl.toolkits.evaluation.recall(Test['isBot'],gl.SArray(final))
print 'ROC Curve: ',gl.toolkits.evaluation.roc_curve(Test['isBot'],gl.SArray(final))
models = CV_GB(Train,features=features)
model_eval(Test,models)
def CV_RF_LC(Train,features=features,bt_fold=5,bt_iter=20,rf_fold=5,rf_iter=20,lc_fold=5,lc_iter=20):
models = []
Folds = gl.cross_validation.KFold(Train,10)
i= 1
print 'GBM: \n'
for train,val in Folds:
print "Starting Fold ",i
model = gl.random_forest_classifier.create(train,features=features,target='isBot',max_iterations=100,validation_set=val,verbose=False)
models.append(model)
print "Done Fold ",i
i=i+1
i= 1
print 'Logistic: \n'
Folds = gl.cross_validation.KFold(Train,10)
for train,val in Folds:
print "Starting Fold ",i
model = gl.logistic_classifier.create(train,features=features,target='isBot',l2_penalty=0.1,validation_set=val,verbose=False,max_iterations=100)
models.append(model)
print "Done Fold ",i
i=i+1
return models
models = CV_RF_LC(Train,features=features)
model_eval(Test,models)
def CV_LC(Train,nfold=5,max_iter=20,features=features):
models = []
i= 1
print 'Logistic: \n'
Folds = gl.cross_validation.KFold(Train,nfold)
for train,val in Folds:
print "Starting Fold ",i
model = gl.logistic_classifier.create(train,features=features,target='isBot',l2_penalty=0.1,validation_set=val,verbose=False,max_iterations=max_iter)
models.append(model)
print "Done Fold ",i
i=i+1
return models
def CV_RF(Train,nfold=5,max_iter=20,features=features):
models = []
i= 1
Folds = gl.cross_validation.KFold(Train,nfold)
print 'RF: \n'
for train,val in Folds:
print "Starting Fold ",i
model = gl.random_forest_classifier.create(train,features=features,target='isBot',validation_set=val,verbose=False,max_iterations=max_iter)
models.append(model)
print "Done Fold ",i
i=i+1
return models
models = CV_LC(Train,features=features)
model_eval(Test,models)
models = CV_RF(Train,features=features)
model_eval(Test,models)
def CV_GB_RF(Train,features=features,bt_fold=5,bt_iter=20,rf_fold=10,rf_iter=20):
models = []
Folds = gl.cross_validation.KFold(Train,bt_fold)
i= 1
print 'GBM: \n'
for train,val in Folds:
print "Starting Fold ",i
model = gl.boosted_trees_classifier.create(train,features=features,target='isBot',validation_set=val,verbose=False,max_iterations=bt_iter)
models.append(model)
print "Done Fold ",i
i=i+1
i= 1
Folds = gl.cross_validation.KFold(Train,rf_fold)
print 'RF: \n'
for train,val in Folds:
print "Starting Fold ",i
model = gl.random_forest_classifier.create(train,features=features,target='isBot',validation_set=val,verbose=False,max_iterations=rf_iter)
models.append(model)
print "Done Fold ",i
i=i+1
return models
models = CV_GB_RF(Train,features=features)
model_eval(Test,models)
models = CV_GB(Train,features=features,max_iter=100,nfold=10)
model_eval(Test,models)
models = CV_GB_RF(Train,features=features,bt_iter=100,bt_fold=7)
model_eval(Test,models)
models = CV_GB_RF(Train,features=features,bt_iter=100,bt_fold=5)
model_eval(Test,models)
temp = gl.nearest_neighbor_classifier.create(Train,target='isBot',features=features)
tt,tt = Test.random_split(0.98)
temp.evaluate(tt,max_neighbors=100)
```
| github_jupyter |
# Comparing Speech Analytics Models
```
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sb
import random as rd
from adaptors import youtube2df, aws2df
```
## Loading csv files
```
# load the real file
df_real = pd.read_csv('real.csv', names=['file', 'transcript'])
# load the model file
df_model = pd.read_csv('no_ipa.csv', names=['file', 'transcript'])
```
## Comparing Two lines
```
# function that neutralizes the string
def neutralize(a_string):
return a_string.lower()
# function that calculates percentage of similarity
def similarity_score(string_real, string_model, extra_info=False):
# cut strings to bag of words
words_real = neutralize(string_real).split(' ')
count = 0
error_words = []
for word in words_real:
# search word in string_model
if word in neutralize(string_model):
count = count + 1
else:
error_words.append(word)
score = count / len(words_real)
if extra_info:
return score, error_words
return score
scr, failed_words = similarity_score(df_real['transcript'][0], df_model['transcript'][0],
extra_info=True)
print(scr)
print(failed_words)
```
## Comparing two dataframes
To compare two dataframes we can use the function we just made but taking care that we
use the correct file from each row.
```
def similarity_score_dataframes(_df_real, _df_model):
score_list = []
for _a_file, a_string in zip(_df_model['file'], _df_model['transcript']):
# first, we search for file in real
real_str = _df_real[_df_real['file']==_a_file]['transcript']
# files should be 1:1, if not, we riot
if len(real_str) != 1:
AttributeError('Dude files are not 1:1 in ' + _a_file)
# otherwise let's continue calculating the score
print(real_str.array)
score, fails = similarity_score(real_str.array[0], a_string, extra_info=True)
score_list.append((_a_file, score, fails))
return pd.DataFrame(score_list, columns=['file', 'score', 'failed_words'])
scr_list = similarity_score_dataframes(df_real, df_model)
scr_list.head()
```
## Example plot for model comparison
We use seaborn box-plots to check performance between two models.
```
# we simulate to score lists
N = 1000
days = ['monday', 'friday', 'sunday']
good_model = [rd.gauss(0.7, 0.1) for i in range(N)]
bad_model = [rd.gauss(0.65, 0.05) for j in range(N)]
# transform my two lists in a dataframe
good_tuples = [('good_model', a_score, days[index%3]) for index, a_score in enumerate(good_model)]
bad_tuples = [('bad_model', a_score, days[index%3]) for index, a_score in enumerate(bad_model)]
df = pd.DataFrame(good_tuples + bad_tuples, columns=['model', 'score', 'day'])
sb.boxplot(data=df, x='day', y='score', hue='model')
plt.show()
```
## Transform Youtube caps and AWS
We need to create an internal function to translate from one file to another easily, to do this, we
transform both files to a dataframe with fields in common.
```
frame = youtube2df('./EdoCaroe/video_0_es-419.txt')
frame.head()
frame = aws2df('video_0.json')
frame.head()
# Debugging adaptor function
frame_test = youtube2df('test.txt')
frame_test.head()
# Create a concatenation of text for each time interval in validation data
# IMPORTANT: We assume words are in order!
# load validation
valid = youtube2df('./EdoCaroe/video_0_es-419.txt')
# load model output
predicted = aws2df('video_0.json').astype({'start':'float', 'end':'float', 'transcript':'string'})
# both dfs must have start and end times in same format and concatenated field
def compress(df_reference, df_to_modify, field='transcript'):
container = []
# for each interval we find a sub-df with all the content
for start, end in zip(df_reference['start'], df_reference['end']):
# sub-df
sub_df = df_to_modify[(df_to_modify['start'] >= start) & (df_to_modify['end'] <= end)]
# now concatenate every word
words = [a_word for a_word in sub_df[field]]
container.append(' '.join(word for word in words))
return container
valid['aws_transcript'] = compress(valid, predicted)
valid.head(10)
```
| github_jupyter |
```
import sys
sys.path.append("../code/")
import util_ElasticNet, lib_LinearAlgebra, util_hdf5, lib_ElasticNet, lib_Checker, util_Stats
import tensorflow as tf
import numpy as np
import pandas as pd
import h5py, yaml, functools
import matplotlib.pyplot as plt
from importlib import reload
lib_LinearAlgebra = reload(lib_LinearAlgebra)
util_ElasticNet = reload(util_ElasticNet)
util_hdf5 = reload(util_hdf5)
lib_ElasticNet = reload(lib_ElasticNet)
lib_Checker = reload(lib_Checker)
util_Stats = reload(util_Stats)
import util_hdf5
import logging, sys
import seaborn as sns
logging.basicConfig(
level = logging.INFO,
stream = sys.stderr,
# filename = logfile,
format = '%(asctime)s %(message)s',
datefmt = '%Y-%m-%d %I:%M:%S %p'
)
gpus = tf.config.experimental.list_physical_devices('GPU')
tf.config.experimental.set_visible_devices(gpus[1], 'GPU')
```
# Overview
We've trained a sequence of elastic net models (with different $\lambda$'s and $\alpha$'s) and here we'd like to compute the PTRS on held out Britishs and other populations.
So, for each $\lambda$ and $\alpha$ pair, we have one PTRS nsample x ntrait matrix.
In total, we will have $$nsample \times (ntrait = 17) \times (nlambda = 50) \times (nalpha = 3)$$ PTRS tensor for each population.
# Load predicted expression
First load British and then load other populations.
```
population = 'British'
# set path to British data
hdf5_british = f'/vol/bmd/yanyul/UKB/predicted_expression_tf2/ukb_imp_x_ctimp_Whole_Blood_{population}.hdf5'
# data scheme specifying which are traits and covariates
scheme_yaml = '../misc_files/data_scheme.yaml'
# loading names of traits/covariates
# the order is matched with the data being loaded
feature_dic = util_hdf5.read_yaml(scheme_yaml)
with h5py.File(hdf5_british, 'r') as f:
features = f['columns_y'][:].astype('str')
sample_size = f['y'].shape[0]
y = f['y'][:]
covar_indice = np.where(np.isin(features, feature_dic['covar_names']))[0]
trait_indice = np.where(np.isin(features, feature_dic['outcome_names']))[0]
# load data_scheme for training
batch_size = 2 ** 12
print(f'batch_size in British set is {batch_size}')
data_scheme, sample_size = util_hdf5.build_data_scheme(
hdf5_british,
scheme_yaml,
batch_size = batch_size,
inv_norm_y = True
)
# set validation and test set as the first and second batch
dataset_valid = data_scheme.dataset.take(1)
data_scheme.dataset = data_scheme.dataset.skip(1)
dataset_test = data_scheme.dataset.take(1)
data_scheme.dataset = data_scheme.dataset.skip(1)
dataset_insample = data_scheme.dataset.take(1)
```
Load other populations.
```
test_datasets = {
'British_validation': dataset_valid,
'British_test': dataset_test,
'British_insample': dataset_insample,
'Chinese': None,
'Indian': None,
'African': None
}
batch_size_here = 8096
for i in test_datasets.keys():
if 'British' not in i:
filename = f'/vol/bmd/yanyul/UKB/predicted_expression_tf2/ukb_imp_x_ctimp_Whole_Blood_{i}.hdf5'
data_scheme, sample_size = util_hdf5.build_data_scheme(
filename,
scheme_yaml,
batch_size = batch_size_here,
inv_norm_y = True
)
test_datasets[i] = data_scheme.dataset
```
# Load models
```
alpha_list = [0.1, 0.5, 0.9]
model_list = {}
for alpha in alpha_list:
filename = f'/vol/bmd/yanyul/UKB/ptrs-tf/models/elastic_net_alpha_{alpha}_British.hdf5'
model_list[alpha] = lib_LinearAlgebra.ElasticNetEstimator('', None, minimal_load = True)
model_list[alpha].minimal_load(filename)
model_list[0.1].lambda_seq
```
# Calculate PTRS
```
out = model_list[0.1].predict_x(test_datasets['British_validation'], model_list[0.1].beta_hat_path)
# out = o1
fig, aes = plt.subplots(nrows = 3, ncols = 3, figsize = (15, 10))
seq = [0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 12, 16, 19]
for i in range(3):
for j in range(3):
idx = seq[(i * 3 + j)] + 45
if idx < len(model_list[alpha].lambda_seq[0]):
for k in range(1):
aes[i][j].scatter(out['y'][:,k], out['y_pred_from_x'][:, k, idx])
aes[i][j].set_title(
'lambda = ' + "{:.3E} cor = {:.3E}".format(
model_list[alpha].lambda_seq[0][idx],
np.corrcoef(out['y'][:, 0], out['y_pred_from_x'][:, 0, idx])[0,1]
) # + '\n' +
# 'lambda = ' + "{:.3E} cor = {:.3E}".format(
# model_list[alpha].lambda_seq[1][idx],
# np.corrcoef(out['y'][:, 1], out['y_pred_from_x'][:, 1, idx])[0,1]
# )
)
```
# Calculate partial R2
```
partial_r2 = {}
for alpha in alpha_list:
partial_r2[alpha] = {}
model_i = model_list[alpha]
for i in test_datasets.keys():
dataset = test_datasets[i]
for ele in dataset:
x, y = model_i.data_scheme.get_data_matrix(ele)
covar = x[:, -len(model_i.data_scheme.covariate_indice) :]
print('alpha = {}, trait = {}, ncol(covar) = {}'.format(alpha, i, covar.shape[1]))
out = model_i.predict_x(dataset, model_i.beta_hat_path)
partial_r2[alpha][i] = util_Stats.quick_partial_r2(covar, out['y'], out['y_pred_from_x'])
def _pr2_format(ele, features, name, alpha, lambda_):
nlambda = lambda_.shape[1]
ntrait = lambda_.shape[0]
ele_seq = np.reshape(ele, (nlambda * ntrait), order = 'C')
lambda_seq = np.reshape(lambda_, (nlambda * ntrait), order = 'C')
f_seq = np.repeat(features, nlambda)
return pd.DataFrame({'partial_r2': ele_seq, 'trait': f_seq, 'sample': name, 'alpha': alpha, 'lambda': lambda_seq})
df = pd.DataFrame({'partial_r2': [], 'trait': [], 'sample': [], 'alpha': [], 'lambda': []})
for alpha in alpha_list:
model_i = model_list[alpha]
lambda_i = np.array(model_i.lambda_seq)
for i in partial_r2[alpha].keys():
df = pd.concat((df, _pr2_format(partial_r2[alpha][i], features[trait_indice], i, alpha, lambda_i)))
df.to_csv('/vol/bmd/yanyul/UKB/ptrs-tf/models/partial_r2-elastic_net_British.tsv', sep = '\t', index = False)
```
| github_jupyter |
# Discrete Random Variables and Sampling
### George Tzanetakis, University of Victoria
In this notebook we will explore discrete random variables and sampling. After defining a helper class and associated functions we will be able to create both symbolic and numeric random variables and generate samples from them.
Define a helper random variable class based on the scipy discrete random variable functionality providing both numeric and symbolic RVs
```
%matplotlib inline
import matplotlib.pyplot as plt
from scipy import stats
import numpy as np
class Random_Variable:
def __init__(self, name, values, probability_distribution):
self.name = name
self.values = values
self.probability_distribution = probability_distribution
if all(type(item) is np.int64 for item in values):
self.type = 'numeric'
self.rv = stats.rv_discrete(name = name, values = (values, probability_distribution))
elif all(type(item) is str for item in values):
self.type = 'symbolic'
self.rv = stats.rv_discrete(name = name, values = (np.arange(len(values)), probability_distribution))
self.symbolic_values = values
else:
self.type = 'undefined'
def sample(self,size):
if (self.type =='numeric'):
return self.rv.rvs(size=size)
elif (self.type == 'symbolic'):
numeric_samples = self.rv.rvs(size=size)
mapped_samples = [values[x] for x in numeric_samples]
return mapped_samples
```
Let's first create some random samples of symbolic random variables corresponding to a coin and a dice
```
values = ['H', 'T']
probabilities = [0.9, 0.1]
coin = Random_Variable('coin', values, probabilities)
samples = coin.sample(20)
print(samples)
values = ['1', '2', '3', '4', '5', '6']
probabilities = [1/6.] * 6
dice = Random_Variable('dice', values, probabilities)
samples = dice.sample(10)
print(samples);
[100] * 10
[1 / 6.] * 3
```
Now let's look at a numeric random variable corresponding to a dice so that we can more easily make plots and histograms
```
values = np.arange(1,7)
probabilities = [1/6.] * 6
dice = Random_Variable('dice', values, probabilities)
samples = dice.sample(100)
plt.stem(samples, markerfmt= ' ')
```
Let's now look at a histogram of these generated samples. Notice that even with 500 samples the bars are not equal length so the calculated frequencies are only approximating the probabilities used to generate them
```
plt.figure()
plt.hist(samples,bins=[1,2,3,4,5,6,7],normed=1, rwidth=0.5,align='left');
```
Let's plot the cumulative histogram of the samples
```
plt.hist(samples,bins=[1,2,3,4,5,6,7],normed=1, rwidth=0.5,align='left', cumulative=True);
```
Let's now estimate the frequency of the event *roll even number* in different ways.
First let's count the number of even numbers in the generated samples. Then let's
take the sum of the counts of the individual estimated probabilities.
```
# we can also write the predicates directly using lambda notation
est_even = len([x for x in samples if x%2==0]) / len(samples)
est_2 = len([x for x in samples if x==2]) / len(samples)
est_4 = len([x for x in samples if x==4]) / len(samples)
est_6 = len([x for x in samples if x==6]) / len(samples)
print(est_even)
# Let's print some estimates
print('Estimates of 2,4,6 = ', (est_2, est_4, est_6))
print('Direct estimate = ', est_even)
print('Sum of estimates = ', est_2 + est_4 + est_6)
print('Theoretical value = ', 0.5)
```
Notice that we can always estimate the probability of an event by simply counting how many times it occurs in the samples of an experiment. However if we have multiple events we are interested in then it can be easier to calculate the probabilities of the values of invdividual random variables and then use the rules of probability to estimate the probabilities of more complex events.
| github_jupyter |
<img src="https://juniorworld.github.io/python-workshop-2018/img/portfolio/week9.png" width="350px">
---
# Unsupervised Machine Learning
- Train a model to give predictions only based on input data
- Dimension Reduction: PCA or SVD
- Clustering Analysis: Modularity, KMeans
## PCA: Principal Component Analysis
- Purpose: Dimension Reduction + Visualization
- Input: high-dimensional data
- Output: low-dimensional data
- Survey: 100 items -> 5 components
```
import pandas as pd
import numpy as np
from sklearn.decomposition import PCA
from sklearn.datasets import load_iris
iris=load_iris()
print(iris.feature_names)
data=iris.data
data.shape
iris.target[0]
pca = PCA(n_components=2)
pca.fit(data)
pca.explained_variance_ratio_
pca = PCA(n_components=3) #initialize a PCA decomposer
pca.fit(data) #train this decomposer with current data set
pca.explained_variance_ratio_
iris_pca=pca.transform(data)
iris_pca.shape
iris_pca[0,:]
#plot this matrix
import plotly.plotly as py
import plotly.graph_objs as go
py.sign_in('yunerzhu','IPqlBadz4qYbTEuOxbL5')
trace=go.Scatter3d(
x=iris_pca[:,0],
y=iris_pca[:,1],
z=iris_pca[:,2],
mode='markers',
marker={'size':3,'color':iris.target})
py.iplot([trace],filename='iris pca')
trace=go.Scatter(
x=iris_pca[:,0],
y=iris_pca[:,1],
mode='markers',
marker={'size':5,'color':iris.target})
py.iplot([trace],filename='iris pca')
```
Try another data set...
```
! pip3 install matplotlib
import matplotlib.pyplot as plt
from sklearn.datasets import load_digits
#load digits data
digits = load_digits()
```
digits is composed of three lists: image list, data list, and label list
```
#"images" contains 8x8 images of each data point.
plt.gray()
plt.matshow(digits.images[0])
plt.show()
#"target" contains a list of results
digits.target[0]
#"data" contains brighness numbers of each data point
data=digits.data
data.shape #overall 1797 cases * 64 brightness number
data[0,:] #first case
#Find the minimum number of components which can explain over 80% of data variance
pca=PCA(n_components=15)
pca.fit(data)
pca = PCA(n_components=3) #specify the component number to 3
pca.fit(data)
digits_3d=pca.transform(data) #obtain the transformed data
pca.explained_variance_ratio_
digits_3d[0:5,:]
trace=go.Scatter3d(
x=digits_3d[:,0],
y=digits_3d[:,1],
z=digits_3d[:,2],
mode='markers',
marker={'size':2,'color':digits.target,'colorscale':'Rainbow'},
text=digits.target)
py.iplot([trace],filename='digits space')
trace=go.Scatter(
x=digits_3d[:,0],
y=digits_3d[:,1],
mode='markers',
marker={'size':5,'color':digits.target,'colorscale':'Rainbow'},
text=digits.target)
py.iplot([trace],filename='digits space')
```
---
## Break
---
## KMeans
- Purpose: cluster data points according to their euclidean distance
- Input: observation data
- Output: predicted groups
- Procedure:
- STEP 1. Intialize K cluster centroids
- STEP 2. Calculate the distance between each data point and each centroid
- STEP 3. Assign data point to the cluster whose centroid is closest to it
- STEP 4. Update the cluster centroids with new group
- STEP 5. Repeat STEP 1~4 for specific time or until convergence
<h3 style="color: red">1. Step-by-Step Breakdowns</h3>
### STEP 1. Initialize K cluster centroids
```
#randomly pick K points as centroids
#Suppose: K=10
random_centroids_index=np.random.choice(range(data.shape[0]),10)
random_centroids_index
range(data.shape[0])
random_centroids=data[random_centroids_index,:]
random_centroids.shape
```
### STEP 2. Calculate the pairwise distance between data point and each centroid
**a) VECTOR NORM**
<img src="https://juniorworld.github.io/python-workshop-2018/img/vector_norm.png" width="200px" align='left'>
```
#According to Pythagorean theorem, if U=(U1,U2,U3,...,Un), ‖U‖=sqrt(U1^2 + U2^2 + U3^2 +...+ Un^2)
a=np.array([0,1,2,3])
norm=np.sqrt(sum([i**2 for i in a]))
np.sqrt(sum([i**2 for i in a]))
norm
np.power(3,2)
#SHORTCUT: Use np.linalg.norm() to calculate the vector norm
np.linalg.norm(a)
```
**b) VECTOR SUBTRACTION**
<img src="https://juniorworld.github.io/python-workshop-2018/img/vector_minus.png" width="250px" align='left'>
```
a=np.array([0,1])
b=np.array([0,2])
np.linalg.norm(a-b)
#get the pairwise distance between first data point and first cluster's centroid
np.linalg.norm(data[0]-random_centroids[0])
#get the distance between all data and first centroid in one line
np.linalg.norm(data-random_centroids[0],axis=1).shape
```
### STEP 3. Pairwise Distance of digit data
```
np.argmin([2,1,3,4,5,6,7,8,9,10]) #index of smallest value
#Calculate the distance between each data point and each centroid, assign point to the cluster depending on this distance
#You need to get a list of cluster assignment
#HINT: you can use np.argmin() to find the index of minimum value
#----------------------------------
def single_run_KMeans(data,centroids):
clusters=[] #initialization
for i in data:
distances=[] #a list for each data point
for centroid in centroids: #go through every centroids
distances.append(np.linalg.norm(i-centroid))
clusters.append(np.argmin(distances))
#Write your code here
clusters=np.array(clusters)
return(clusters)
def single_run_KMeans2(data,centroids):
clusters=[] #initialization
for i in data:
distances=np.linalg.norm(i-centroids,axis=1)
clusters.append(np.argmin(distances))
#Write your code here
clusters=np.array(clusters)
return(clusters)
first_run_cluster=single_run_KMeans(data,random_centroids)
first_run_cluster #first point belongs to this cluster after first run
single_run_KMeans2(data,random_centroids)
```
### STEP 4. Update the centroids
Centroids: centers of a group of points/vectors
- Measure: avarage of coordinates
<img src="https://juniorworld.github.io/python-workshop-2018/img/centroids.png" width="250px" align='left'>
```
a=[[1,2,3],
[2,3,4],
[4,5,6],
[6,7,8]]
print(np.mean(a,axis=0)) #column mean
print(np.mean(a,axis=1)) #row mean
np.mean(a,axis=0).shape
data[first_run_cluster==5,:] #points in cluster 5
first_run_cluster==0
a=[1,2,3]
a.append([3,4,5])
print(a)
a=[1,2,3]
a.extend([3,4,5])
print(a)
#Update the centroids of clusters
def update_centroids(data,n_clusters,clusters):
centroids=[]
for i in range(n_clusters):
centroids.append(np.mean(data[clusters==i,:],axis=0))
#Write your code here
centroids=np.array(centroids)
return(centroids)
data[first_run_cluster==0]
update_centroids(data,first_run_cluster) #get our second-run centroids
```
### STEP 5. Calculate the Loss
Loss in Machine Learning = Goodness-of-fit in Social Sciences
- Types of loss: L1 (abs error), L2 (sqaured error) and logistic/cross-entropy
- L1: mean(abs(y-ŷ))
- L2: mean((y-ŷ)^2)
- Log: mean(-sum(y*log(ŷ))
- For KMeans, we use L2 loss:
- average squared distance between points and their centroids
- formula: mean((y-centroid)^2)
```
def loss(data,n_clusters,clusters,centroids):
ls=[]
for i in range(n_clusters):
ls.extend(np.linalg.norm(data[clusters==i,:]-centroids[i],axis=1)) #distance
ls=np.mean([j**2 for j in ls]) #squared distance
return(ls)
#performance of first run
loss(data,first_run_cluster,random_centroids)
```
### Training model for specific times (integrated)
```
centroids=random_centroids
loss_list=[]
for run in range(20):
clusters=single_run_KMeans(data,centroids)
current_loss=loss(data,clusters,centroids)
print(run,current_loss)
loss_list.append(current_loss)
centroids=update_centroids(data,clusters)
#elbow method of finding the optimal number of iteration
trace=go.Scatter(
x=list(range(20)),
y=loss_list,
mode='lines'
)
py.iplot([trace],filename='learning curve')
#have a look at the cluster results
digits.target[clusters==2]
```
### Training model until convergence (integrated)
```
centroids=random_centroids
current_loss=10000
loss_list=[]
while current_loss>700:
clusters=single_run_KMeans(data,centroids)
current_loss=loss(data,clusters,centroids)
print(current_loss)
loss_list.append(current_loss)
centroids=update_centroids(data,clusters)
#centroids=random_centroids
previous_loss=10000 #intialization
ls_improvement=100 #intialization
while ls_improvement>0:
clusters=single_run_KMeans(data,centroids)
current_loss=loss(data,clusters,centroids)
ls_improvement=previous_loss-current_loss
print(current_loss)
previous_loss=current_loss
centroids=update_centroids(data,clusters)
```
## KMeans++: An Improvement of KMeans
- New way of initialization
- STEP 1: Random pick one centroid from the points
- STEP 2: Calculate the distance _D(k)_ between points and their nearest centroid
- STEP 3: Pick one more centroid with probability proportional to _D(k)_
- STEP 4: Repeat STEP 2 and STEP 3 until the number of centroids reaching the required value
```
#STEP 1
first_centroid_index=np.random.choice(range(data.shape[0]),1)
first_centroid=data[first_centroid_index]
#HINT 1: You can use np.random.choice(data,1,p=[probability list]) to pick one point randomly with given list of probability
#HINT 2: You can use np.vstack((array1,array2)) to add new row
#WRITE YOUR CODE HERE
def kmeans_plus(data,n_clusters):
first_centroid_index=np.random.choice(range(data.shape[0]),1)
first_centroid=data[first_centroid_index]
centroids=first_centroid
for i in range(n_clusters-1): #create remaining 9 centroids
prob=[]
for j in data:
distances=np.linalg.norm(j-centroids,axis=1) #distance between point and all available centroids
distance=np.min(distances) #choose the minimum distance
prob.append(distance)
prob=prob/sum(prob) #convert the distance into probability, ranging between 0 and 1, with sum of 100%
next_centroid_index=np.random.choice(range(data.shape[0]),1,p=prob)
next_centroid=data[next_centroid_index,:]
centroids=np.vstack((centroids,next_centroid)) #add new centroid to the list
centroids=np.array(centroids)#convert the list to numpy array
return(centroids)
def random_init(data,n_clusters):
random_centroids_index=np.random.choice(range(data.shape[0]),n_clusters)
random_centroids=data[random_centroids_index,:]
return(random_centroids)
centroids
```
# Combine everything into a giant function
- input: data
- parameter:init = random/kmeans++, iteration = num/covergence threshold
- output: a list of cluster assignments
```
def KMeans(data,n_clusters,init='random',iteration=10,convergence_threshold=False):
if init=='random':
centroids=random_init(data,n_clusters)
elif init=='kmeans++':
centroids=kmeans_plus(data,n_clusters)
else:
print('initialization error!')
if type(iteration)==int:
for run in range(iteration):
clusters=single_run_KMeans(data,centroids)
current_loss=loss(data,n_clusters,clusters,centroids)
print(run,current_loss)
centroids=update_centroids(data,n_clusters,clusters)
return(clusters)
if convergence_threshold:
current_loss=10000
while current_loss>700:
clusters=single_run_KMeans(data,centroids)
current_loss=loss(data,n_clusters,clusters,centroids)
print(current_loss)
centroids=update_centroids(data,n_clusters,clusters)
return(clusters)
#APPLY Kmeans() to iris data
clusters=KMeans(iris.data,3)
```
## SHORTCUT: sklearn function
documentation: https://scikit-learn.org/stable/modules/generated/sklearn.cluster.KMeans.html#sklearn.cluster.KMeans
```
from sklearn.cluster import KMeans
kmeans=KMeans(n_clusters=10,init='k-means++',max_iter=20)
kmeans.fit(data)
kmeans.labels_
digits.target[kmeans.labels_==1]
```
## Find the best value of K
```
#a data set about development score of countries
countries=pd.read_csv('https://juniorworld.github.io/python-workshop-2018/doc/country-index.csv')
countries.head()
kmeans=KMeans(n_clusters=5,init='k-means++')
kmeans.fit(countries.iloc[:,3:])
countries['countries'][kmeans.labels_==0]
ls_list=[]
for i in range(1,21):
kmeans=KMeans(n_clusters=i,init='k-means++')
kmeans.fit(countries.iloc[:,3:])
ls=loss(np.array(countries.iloc[:,3:]),kmeans.labels_,kmeans.cluster_centers_)
ls_list.append(ls)
#elbow method of finding the optimal K
trace=go.Scatter(
x=list(range(1,21)),
y=ls_list,
mode='lines'
)
py.iplot([trace],filename='learning curve')
```
| github_jupyter |
##### Copyright 2019 The TensorFlow Hub Authors.
Licensed under the Apache License, Version 2.0 (the "License");
```
# Copyright 2018 The TensorFlow Hub Authors. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ==============================================================================
```
# Semantic Search with Approximate Nearest Neighbors and Text Embeddings from TF-Hub
This tutorial illustrates how to generate embeddings from a [TensorFlow Hub](https://tfhub.dev) (TF-Hub) module given input data, and build an approximate nearest neighbours (ANN) index using the extracted embeddings. The index can then be used for real-time similarity matching and retrieval.
When dealing with a large corpus of data, it's not efficient to perform exact matching by scanning the whole repository to find the most similar items to a given query in real-time. Thus, we use an approximate similarity matching algorithm which allows us to trade off a little bit of accuracy in finding exact nearest neighbor matches for a significant boost in speed.
In this tutorial, we show an example of real-time text search over a corpus of news headlines to find the headlines that are most similar to a query. Unlike keyword search, this captures the semantic similarity encoded in the text embedding.
The steps of this tutorial are:
1. Download sample data.
2. Generate embeddings for the data using a TF-Hub module
3. Build an ANN index for the embeddings
4. Use the index for similarity matching
We use [Apache Beam](https://beam.apache.org/documentation/programming-guide/) to generate the embeddings from the TF-Hub module. We also use Spotify's [ANNOY](https://github.com/spotify/annoy) library to build the approximate nearest neighbours index.
## Getting Started
Install the required libraries.
```
!pip install --upgrade tensorflow
!pip install --upgrade tensorflow_hub
!pip install apache_beam
!pip install sklearn
!pip install annoy
```
Import the required libraries
```
import os
import sys
import pickle
from collections import namedtuple
from datetime import datetime
import numpy as np
import apache_beam as beam
from apache_beam.transforms import util
import tensorflow as tf
import tensorflow_hub as hub
import annoy
from sklearn.random_projection import gaussian_random_matrix
print('TF version: {}'.format(tf.__version__))
print('TF-Hub version: {}'.format(hub.__version__))
print('Apache Beam version: {}'.format(beam.__version__))
```
## 1. Download Sample Data
[A Million News Headlines](https://dataverse.harvard.edu/dataset.xhtml?persistentId=doi:10.7910/DVN/SYBGZL#) dataset contains news headlines published over a period of 15 years sourced from the reputable Australian Broadcasting Corp. (ABC). This news dataset has a summarised historical record of noteworthy events in the globe from early-2003 to end-2017 with a more granular focus on Australia.
**Format**: Tab-separated two-column data: 1) publication date and 2) headline text. We are only interested in the headline text.
```
!wget 'https://dataverse.harvard.edu/api/access/datafile/3450625?format=tab&gbrecs=true' -O raw.tsv
!wc -l raw.tsv
!head raw.tsv
```
For simplicity, we only keep the headline text and remove the publication date
```
!rm -r corpus
!mkdir corpus
with open('corpus/text.txt', 'w') as out_file:
with open('raw.tsv', 'r') as in_file:
for line in in_file:
headline = line.split('\t')[1].strip().strip('"')
out_file.write(headline+"\n")
!tail corpus/text.txt
```
## 2. Generate Embeddings for the Data.
In this tutorial, we use the [Neural Network Language Model (NNLM)](https://tfhub.dev/google/tf2-preview/nnlm-en-dim128/1) to generate embeddings for the headline data. The sentence embeddings can then be easily used to compute sentence level meaning similarity. We run the embedding generation process using Apache Beam.
### Embedding extraction method
```
embed_fn = None
def generate_embeddings(text, module_url, random_projection_matrix=None):
# Beam will run this function in different processes that need to
# import hub and load embed_fn (if not previously loaded)
global embed_fn
if embed_fn is None:
embed_fn = hub.load(module_url)
embedding = embed_fn(text).numpy()
if random_projection_matrix is not None:
embedding = embedding.dot(random_projection_matrix)
return text, embedding
```
### Convert to tf.Example method
```
def to_tf_example(entries):
examples = []
text_list, embedding_list = entries
for i in range(len(text_list)):
text = text_list[i]
embedding = embedding_list[i]
features = {
'text': tf.train.Feature(
bytes_list=tf.train.BytesList(value=[text.encode('utf-8')])),
'embedding': tf.train.Feature(
float_list=tf.train.FloatList(value=embedding.tolist()))
}
example = tf.train.Example(
features=tf.train.Features(
feature=features)).SerializeToString(deterministic=True)
examples.append(example)
return examples
```
### Beam pipeline
```
def run_hub2emb(args):
'''Runs the embedding generation pipeline'''
options = beam.options.pipeline_options.PipelineOptions(**args)
args = namedtuple("options", args.keys())(*args.values())
with beam.Pipeline(args.runner, options=options) as pipeline:
(
pipeline
| 'Read sentences from files' >> beam.io.ReadFromText(
file_pattern=args.data_dir)
| 'Batch elements' >> util.BatchElements(
min_batch_size=args.batch_size, max_batch_size=args.batch_size)
| 'Generate embeddings' >> beam.Map(
generate_embeddings, args.module_url, args.random_projection_matrix)
| 'Encode to tf example' >> beam.FlatMap(to_tf_example)
| 'Write to TFRecords files' >> beam.io.WriteToTFRecord(
file_path_prefix='{}/emb'.format(args.output_dir),
file_name_suffix='.tfrecords')
)
```
### Generaring Random Projection Weight Matrix
[Random projection](https://en.wikipedia.org/wiki/Random_projection) is a simple, yet powerfull technique used to reduce the dimensionality of a set of points which lie in Euclidean space. For a theoretical background, see the [Johnson-Lindenstrauss lemma](https://en.wikipedia.org/wiki/Johnson%E2%80%93Lindenstrauss_lemma).
Reducing the dimensionality of the embeddings with random projection means less time needed to build and query the ANN index.
In this tutorial we use [Gaussian Random Projection](https://en.wikipedia.org/wiki/Random_projection#Gaussian_random_projection) from the [Scikit-learn](https://scikit-learn.org/stable/modules/random_projection.html#gaussian-random-projection) library.
```
def generate_random_projection_weights(original_dim, projected_dim):
random_projection_matrix = None
random_projection_matrix = gaussian_random_matrix(
n_components=projected_dim, n_features=original_dim).T
print("A Gaussian random weight matrix was creates with shape of {}".format(random_projection_matrix.shape))
print('Storing random projection matrix to disk...')
with open('random_projection_matrix', 'wb') as handle:
pickle.dump(random_projection_matrix,
handle, protocol=pickle.HIGHEST_PROTOCOL)
return random_projection_matrix
```
### Set parameters
If you want to build an index using the original embedding space without random projection, set the `projected_dim` parameter to `None`. Note that this will slow down the indexing step for high-dimensional embeddings.
```
module_url = 'https://tfhub.dev/google/tf2-preview/nnlm-en-dim128/1' #@param {type:"string"}
projected_dim = 64 #@param {type:"number"}
```
### Run pipeline
```
output_dir = '/embeds'
original_dim = hub.load(module_url)(['']).shape[1]
random_projection_matrix = None
if projected_dim:
random_projection_matrix = generate_random_projection_weights(
original_dim, projected_dim)
args = {
'job_name': 'hub2emb-{}'.format(datetime.utcnow().strftime('%y%m%d-%H%M%S')),
'runner': 'DirectRunner',
'batch_size': 1024,
'data_dir': 'corpus/*.txt',
'output_dir': output_dir,
'module_url': module_url,
'random_projection_matrix': random_projection_matrix,
}
print("Pipeline args are set.")
args
!rm -r {output_dir}
print("Running pipeline...")
%time run_hub2emb(args)
print("Pipeline is done.")
!ls {output_dir}
```
Read some of the generated embeddings...
```
embed_file = os.path.join(output_dir, 'emb-00000-of-00001.tfrecords')
sample = 5
# Create a description of the features.
feature_description = {
'text': tf.io.FixedLenFeature([], tf.string),
'embedding': tf.io.FixedLenFeature([projected_dim], tf.float32)
}
def _parse_example(example):
# Parse the input `tf.Example` proto using the dictionary above.
return tf.io.parse_single_example(example, feature_description)
dataset = tf.data.TFRecordDataset(embed_file)
for record in dataset.take(sample).map(_parse_example):
print("{}: {}".format(record['text'].numpy().decode('utf-8'), record['embedding'].numpy()[:10]))
```
## 3. Build the ANN Index for the Embeddings
[ANNOY](https://github.com/spotify/annoy) (Approximate Nearest Neighbors Oh Yeah) is a C++ library with Python bindings to search for points in space that are close to a given query point. It also creates large read-only file-based data structures that are mmapped into memory. It is built and used by [Spotify](https://www.spotify.com) for music recommendations.
```
def build_index(embedding_files_pattern, index_filename, vector_length,
metric='angular', num_trees=100):
'''Builds an ANNOY index'''
annoy_index = annoy.AnnoyIndex(vector_length, metric=metric)
# Mapping between the item and its identifier in the index
mapping = {}
embed_files = tf.io.gfile.glob(embedding_files_pattern)
num_files = len(embed_files)
print('Found {} embedding file(s).'.format(num_files))
item_counter = 0
for i, embed_file in enumerate(embed_files):
print('Loading embeddings in file {} of {}...'.format(i+1, num_files))
dataset = tf.data.TFRecordDataset(embed_file)
for record in dataset.map(_parse_example):
text = record['text'].numpy().decode("utf-8")
embedding = record['embedding'].numpy()
mapping[item_counter] = text
annoy_index.add_item(item_counter, embedding)
item_counter += 1
if item_counter % 100000 == 0:
print('{} items loaded to the index'.format(item_counter))
print('A total of {} items added to the index'.format(item_counter))
print('Building the index with {} trees...'.format(num_trees))
annoy_index.build(n_trees=num_trees)
print('Index is successfully built.')
print('Saving index to disk...')
annoy_index.save(index_filename)
print('Index is saved to disk.')
print("Index file size: {} GB".format(
round(os.path.getsize(index_filename) / float(1024 ** 3), 2)))
annoy_index.unload()
print('Saving mapping to disk...')
with open(index_filename + '.mapping', 'wb') as handle:
pickle.dump(mapping, handle, protocol=pickle.HIGHEST_PROTOCOL)
print('Mapping is saved to disk.')
print("Mapping file size: {} MB".format(
round(os.path.getsize(index_filename + '.mapping') / float(1024 ** 2), 2)))
embedding_files = "{}/emb-*.tfrecords".format(output_dir)
embedding_dimension = projected_dim
index_filename = "index"
!rm {index_filename}
!rm {index_filename}.mapping
%time build_index(embedding_files, index_filename, embedding_dimension)
!ls
```
## 4. Use the Index for Similarity Matching
Now we can use the ANN index to find news headlines that are semantically close to an input query.
### Load the index and the mapping files
```
index = annoy.AnnoyIndex(embedding_dimension)
index.load(index_filename, prefault=True)
print('Annoy index is loaded.')
with open(index_filename + '.mapping', 'rb') as handle:
mapping = pickle.load(handle)
print('Mapping file is loaded.')
```
### Similarity matching method
```
def find_similar_items(embedding, num_matches=5):
'''Finds similar items to a given embedding in the ANN index'''
ids = index.get_nns_by_vector(
embedding, num_matches, search_k=-1, include_distances=False)
items = [mapping[i] for i in ids]
return items
```
### Extract embedding from a given query
```
# Load the TF-Hub module
print("Loading the TF-Hub module...")
%time embed_fn = hub.load(module_url)
print("TF-Hub module is loaded.")
random_projection_matrix = None
if os.path.exists('random_projection_matrix'):
print("Loading random projection matrix...")
with open('random_projection_matrix', 'rb') as handle:
random_projection_matrix = pickle.load(handle)
print('random projection matrix is loaded.')
def extract_embeddings(query):
'''Generates the embedding for the query'''
query_embedding = embed_fn([query])[0].numpy()
if random_projection_matrix is not None:
query_embedding = query_embedding.dot(random_projection_matrix)
return query_embedding
extract_embeddings("Hello Machine Learning!")[:10]
```
### Enter a query to find the most similar items
```
#@title { run: "auto" }
query = "confronting global challenges" #@param {type:"string"}
print("Generating embedding for the query...")
%time query_embedding = extract_embeddings(query)
print("")
print("Finding relevant items in the index...")
%time items = find_similar_items(query_embedding, 10)
print("")
print("Results:")
print("=========")
for item in items:
print(item)
```
## Want to learn more?
You can learn more about TensorFlow at [tensorflow.org](https://www.tensorflow.org/) and see the TF-Hub API documentation at [tensorflow.org/hub](https://www.tensorflow.org/hub/). Find available TensorFlow Hub modules at [tfhub.dev](https://tfhub.dev/) including more text embedding modules and image feature vector modules.
Also check out the [Machine Learning Crash Course](https://developers.google.com/machine-learning/crash-course/) which is Google's fast-paced, practical introduction to machine learning.
| github_jupyter |
```
# List of high schools
high_schools = ["Hernandez High School", "Figueroa High School",
"Wilson High School","Wright High School"]
for school in high_schools:
print(school)
# A dictionary of high schools and the type of school.
high_school_types = [{"High School": "Griffin", "Type":"District"},
{"High School": "Figueroa", "Type": "District"},
{"High School": "Wilson", "Type": "Charter"},
{"High School": "Wright", "Type": "Charter"}]
for school_type in high_school_types:
print(school_type)
# List of high schools
high_schools = ["Huang High School", "Figueroa High School", "Shelton High School", "Hernandez High School","Griffin High School","Wilson High School", "Cabrera High School", "Bailey High School", "Holden High School", "Pena High School", "Wright High School","Rodriguez High School", "Johnson High School", "Ford High School", "Thomas High School"]
# Add the Pandas dependency.
import pandas as pd
# Create a Pandas Series from a list.
school_series = pd.Series(high_schools)
school_series
print("Iteration through school_series")
for i in range(len(school_series)):
print(school_series[i])
# A dictionary of high schools
high_school_dicts = [{"School ID": 0, "school_name": "Huang High School", "type": "District"},
{"School ID": 1, "school_name": "Figueroa High School", "type": "District"},
{"School ID": 2, "school_name":"Shelton High School", "type": "Charter"},
{"School ID": 3, "school_name":"Hernandez High School", "type": "District"},
{"School ID": 4, "school_name":"Griffin High School", "type": "Charter"}]
school_df = pd.DataFrame(high_school_dicts)
school_df
# Three separate lists of information on high schools
school_id = [0, 1, 2, 3, 4]
school_name = ["Huang High School", "Figueroa High School",
"Shelton High School", "Hernandez High School","Griffin High School"]
type_of_school = ["District", "District", "Charter", "District","Charter"]
# Initialize a new DataFrame.
schools_df = pd.DataFrame()
schools_df["School ID"] = school_id
schools_df["School Name"] = school_name
schools_df["Type"] = type_of_school
schools_df
# Create a dictionary of information on high schools.
high_schools_dict = {'School ID': school_id, 'school_name':school_name, 'type':type_of_school}
schools_df = pd.DataFrame(high_school_dicts)
schools_df
schools_df.columns
schools_df.index
schools_df.values
school_id = [0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14]
school_name = ["Huang High School", "Figueroa High School","Shelton High School", "Hernandez High School",
"Griffin High School", "Wilson High School", "Cabrera High School", "Bailey High School",
"Holden High School", "Pena high School", "Wright Hish School", "Rodriguez High School",
"Johnson High School", "Ford High School", "Thomas High School"]
type_of_school = ["District", "District", "Charter", "District","Charter", "Charter", "Charter", "District", "Charter",
"Charter", "Charter", "District", "District", "District", "Charter"]
# Initialize a new DataFrame.
schools_df = pd.DataFrame()
schools_df["School ID"] = school_id
schools_df["School Name"] = school_name
schools_df["Type"] = type_of_school
schools_df
df = pd.DataFrame()
# Reorder the columns in the order you want them to appear.
new_column_order = ["column2", "column4", "column1"]
# Assign a new or the same DataFrame the new column order.
df = df[new_column_order]
df
```
| github_jupyter |
<blockquote>
<h1>Exercise 7.12</h1>
<p>This problem is a continuation of the previous exercise. In a toy example with $p = 100$, show that one can approximate the multiple linear regression coefficient estimates by repeatedly performing simple linear regression in a backfitting procedure. How many backfitting iterations are required in order to obtain a "good" approximation to the multiple regression coefficient estimates? Create a plot to justify your answer.</p>
</blockquote>
```
import numpy as np
import matplotlib.pyplot as plt
# https://stackoverflow.com/questions/34398054/ipython-notebook-cell-multiple-outputs
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
import statsmodels.api as sm
from sklearn.metrics import mean_squared_error
np.random.seed(42)
p = 100
n = 1000
X = np.zeros(shape=(n, p))
gaussian_noise = np.random.normal(size=n, loc=0, scale=2)
beta = np.zeros(shape=(p + 1, ))
beta[0] = -7.5
for i in range(p):
loc = np.random.normal(loc=0, scale=100)
scale = abs(np.random.normal(loc=0, scale=10))
X[:, i] = np.random.normal(size=n, loc=loc, scale=scale)
beta[i + 1] = np.random.normal(loc=0, scale=50)
X_intercept = sm.add_constant(X)
Y = X_intercept @ beta + gaussian_noise
max_iter = 500
beta_hat = np.ones(shape=(max_iter, p + 1))
mse = []
Y_pred = X_intercept @ beta_hat[0]
mse.append(mean_squared_error(Y, Y_pred))
threshold = 1e-6
for i in range(1, max_iter): # i = 0 is the initialization row
for k in range(1, p + 1):
a = Y - np.delete(X_intercept, [0, k], axis=1) @ np.delete(beta_hat[i - 1, :], [0, k])
fitted = sm.OLS(a, X_intercept[:, [0, k]]).fit()
beta_hat[i, k] = fitted.params[1]
Y_pred = X_intercept @ beta_hat[i]
mse.append(mean_squared_error(Y, Y_pred))
if len(mse) >= 1 and abs(mse[-2] - mse[-1]) < threshold:
break
mse = np.array(mse)
iterations = np.arange(mse.shape[0])
fig, ax = plt.subplots(1, 1, constrained_layout=True, figsize=(8, 4))
_ = ax.scatter(iterations, mse)
_ = ax.set_xlabel('iterations')
_ = ax.set_ylabel('MSE')
beta_diff = np.zeros(shape=(mse.shape[0], ))
for i in range(mse.shape[0]):
beta_diff[i] = np.sum(beta_hat[i] - beta)
iterations = np.arange(beta_diff.shape[0])
fig, ax = plt.subplots(1, 1, constrained_layout=True, figsize=(8, 4))
_ = ax.scatter(iterations, beta_diff)
_ = ax.set_xlabel('iterations')
_ = ax.set_ylabel(r'$\sum_{i=0}^p \beta_i - \hat{\beta}_i$')
```
| github_jupyter |
### TME sur Echantillonage
## Diffusion dans les graphes
Au cours des vingt dernières années, les réseaux sociaux sont devenus un média d’information incontournable, mettant en jeu des dynamiques complexes de communication entre utilisateurs. La modélisation de la diffusion d’information sur les réseaux constitue depuis lors un enjeu majeur, pour diverses tâches
telles que l’identification de leaders d’opinions, la prédiction ou la maximisation de l’impact d’un contenu diffusé, la détection de communautés d’opinions, ou plus généralement l’analyse des dynamiques du réseau considéré.
Le modèle proposé par (Saito et al, 2009) considère une diffusion en cascade dans laquelle l'information transite de noeuds en noeuds du réseau en suivant des relations d'influence entre les utilisateurs. Lorsqu'un utilisateur est ``infecté'' par une information, il possède une chance unique de la retransmettre à chacun de ses successeurs dans le graphe, selon une probabilité définie sur le lien correspondant. Le modèle définit en fait deux paramètres sur chaque lien $(u,v)$ du graphe:
* $k_{u,v}$: la probabilité que l'utilisateur $u$ transmette une information diffusée à $v$
* $r_{u,v}$: si la transmission s'effectue, l'utilisateur $v$ la reçoit au temps $t_v=t_u+\delta$, avec $\delta \sim Exp(r_{u,v})$
Pour utiliser ce modèle, on devra donc échantillonner selon la distribution exponentielle. Pour commencer, on cherche alors à écrire une méthode $exp(rate)$ qui échantillonne des variables d'une loi exponentielle selon le tableau d'intensités $rate$ passé en paramètre. Cet échantillonnage se fera par **Inverse Transform Sampling**. Pour éviter les divisions par 0, on ajoutera $1e-200$ aux intensités qui valent 0.
```
import numpy as np
np.random.seed(0)
def exp(rate):
#Votre code ici
u=np.random.rand(*np.array(rate).shape)
return -np.log(u)/rate
#Test
a=exp(np.array([[1,2,3],[4,5,6]]))
for i in range(10000):
a+=exp(np.array([[1,2,3],[4,5,6]]))
print(a/10000)
# Pour comparaison:
a=np.random.exponential(1.0/np.array([[1,2,3],[4,5,6]]))
for i in range(10000):
a+=np.random.exponential(1.0/np.array([[1,2,3],[4,5,6]]))
print(a/10000)
```
Soit le graphe de diffusion donné ci dessous:
```
names={0:"Paul",1:"Jean",2:"Hector",3:"Rose",4:"Yasmine",5:"Léo",6:"Amine",7:"Mia",8:"Quentin",9:"Gaston",10:"Louise"}
k={(0,1):0.9,(1,0):0.9,(1,2):0.2,(2,3):0.5,(3,2):0.4,(2,4):0.9,(4,3):0.9,(1,3):0.5,(2,5):0.5,(5,7):0.7,(1,6):0.2,(6,7):0.1,(1,8):0.8,(8,9):0.2,(1,10):0.5,(10,9):0.9,(8,1):0.8}
r={(0,1):0.2,(1,0):3,(1,2):1,(2,3):0.2,(3,2):0.5,(2,4):10,(4,3):2,(1,3):2,(2,5):0.5,(5,7):15,(1,6):3,(6,7):4,(1,8):0.8,(8,9):0.1,(1,10):12,(10,9):1,(8,1):14}
graph=(names,k,r)
```
La fonction display_graph ci dessous permet de visualiser le graphe de diffusion correspondant:
```
import pydot
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
style = { "bgcolor" : "#6b85d1", "fgcolor" : "#FFFFFF" }
def display_graph ( graph_data, style, graph_name="diffusion_graph" ):
graph = pydot.Dot( graph_name , graph_type='digraph')
names,k,r=graph_data
# création des noeuds du réseau
for (i,name) in names.items():
new_node = pydot.Node( str(i)+"_"+name,
style="filled",
fillcolor=style["bgcolor"],
fontcolor=style["fgcolor"] )
graph.add_node( new_node )
# création des arcs
for edge,valk in k.items():
valr=r[edge]
n1=str(edge[0])+"_"+names[edge[0]]
n2=str(edge[1])+"_"+names[edge[1]]
new_edge = pydot.Edge ( n1, n2, label="k="+str(valk)+",r="+str(valr))
graph.add_edge ( new_edge )
# sauvegarde et affaichage
outfile = graph_name + '.png'
graph.write_png( outfile )
img = mpimg.imread ( outfile )
plt.imshow( img )
display_graph(graph,style)
```
On souhaite être capable d'estimer les probabilités marginales d'infection des différents utilisateurs du réseau par une information pour laquelle on connaît le début de la diffusion.
Pour cela, on considère une liste d'infections qui correspond aux couples (utilisateur, temps d'infection) observés pour cette diffusion pendant une période de temps initial. Par exemple la liste $infections=[(1,0),(4,1)]$
correspond à une diffusion pour laquelle on a observé l'infection de l'utilisateur 1 au temps 0 et l'infection de l'utilisateur 4 au temps 1.
Etant donnés les cycles possibles dans le graphe de diffusion, considérer un calcul exact des probabilités d'infection des différents utilisateurs sachant le début de la diffusion est inenvisageable : il faudrait considérer toutes les combinaisons possibles (infinies) de temps d'infection pour tous les utilisateurs non observés dans la période initiale.
Une possibilité pour calculer ces probabilités d'infections est de travailler par échantillonnage de Monte Carlo: on réalise $n$ tirages d'infections connaissant le début de la diffusion et on recense le ratio des simulations dans lesquelles chacun des utilisateurs est infecté avant un temps $maxT$.
L'idée est alors dans un premier temps d'écrire une méthode $simulation(graph,infections,max\_obs)$ qui, à partir d'une liste d'infections initiales et d'un temps d'observation maximal $max\_obs$, retourne les temps d'infection de l'ensemble des noeuds en fin de diffusion, sous la forme d'un tableau où chaque case $i$ contient le temps d'infection du noeud $i$. Si le noeud $i$ n'a pas été infecté ou bien si il l'a été après un temps maximal $maxT$, la case $i$ contient alors la valeur $maxT$.
Le pseudo-code de la méthode de simulation est donné ci dessous, avec $t_i$ le temps d'infection courant du noeud $i$:
```
ti=maxT pour tout i non infecté dans la liste initiale
Tant qu'il reste des infectieux dont le temps est < maxT:
i=Infectieux de temps d'infection minimal
Pour tout noeud j tel que tj>ti:
sampler x selon P(i->j|ti,tj>max_obs)
si x==1:
sampler delta selon Exp(rij)
t=ti+delta
si ti<max_obs: t+=max_obs-ti (c'est en fait une exponentielle tronquée à gauche, d'où la translation)
si t<tj: tj=t
Retrait de i de la liste des infectieux
```
où $P(i\rightarrow j|t_i,t_j>max\_obs)$ correspond à la probabilité que $i$ transmette l'information à $j$ (quel que soit le temps d'infection) sachant qu'il ne l'a pas infecté avant $max\_obs$:
$P(i\rightarrow j|t_i,t_j>max\_obs)=\begin{cases} k_{i,j} \text{ si } t_i\geq max\_obs, \\ \dfrac{k_{i,j} exp(-r_{i,j}(max\_obs-t_i))}{k_{i,j} exp(-r_{i,j}(max\_obs-t_i)) + 1 - k_{i,j} } sinon. \end{cases}$
Complétez le code de la fonction donnée ci-dessous:
```
np.random.seed(0)
maxT=10
def simulation(graph,infections,max_obs):
names,gk,gr=graph
nbNodes=len(names)
#au départ,on suppose que tous les noeuds ont le temps maximal à s'infecter
#(ti=maxT pour tout i non infecté dans la liste initiale)
infectious=np.array([maxT]*nbNodes,dtype=float)
#liste des ids et liste des temps (zip pour parcourir, puis appliquer array grace à map)
ids,t=map(np.array,zip(*infections))
#retourne les ids dont t est inferieur à max_obs
ids=ids[t<=max_obs]
#retourne les t dont t inferieur à t
t=t[t<=max_obs]
#changer les valeurs des infectious où t<=max_obs
infectious[ids]=t
times=np.copy(infectious)
inf=np.copy(infectious)
#Tant qu'il reste des infectieux dont le temps est < maxT:
while np.any([infectious<maxT]):
#i=Infectieux de temps d'infection minimal
i = np.argmin(infectious)
# Pour tout noeud j tel que tj>ti:
for j in range(0,len(times)):
if(times[j] > times[i]):
if(not(i,j) in gr.keys()):
x = 0
else:
#sampler x selon P(i->j|ti,tj>max_obs)
x = P(i,j,graph,times,max_obs)
# si x==1:
if(x == 1):
# sampler delta selon Exp(rij)
rij = graph[2].get((i,j),0)
delta = exp([[rij]])[0]
# t=ti+delta
t = times[i] + delta
# si ti<max_obs: t+=max_obs-ti (c'est en fait une exponentielle tronquée à gauche, d'où la translation)
if(times[i] < max_obs):
t+=max_obs-times[i]
# si t<tj: tj=t
if(t<times[j]):
times[j] = t
infectious[j] = t
#Retrait de i de la liste des infectieux
infectious[i] = np.inf # (ne plus le considerer au lieu de le supprimer, condition de boucle au début)
return(times)
#𝑃(𝑖→𝑗|𝑡𝑖,𝑡𝑗>𝑚𝑎𝑥_𝑜𝑏𝑠) correspond à la probabilité que 𝑖 transmette l'information à 𝑗 (quel que soit le temps d'infection) sachant qu'il ne l'a pas infecté
def P(i,j,graph,times,max_obs):
names,gk,gr=graph
#ne pas considerer
if(not (i,j) in gr.keys()):
return 0
#depasser le temps d'observation
if(times[i] > max_obs):
# tirer une valeur aléatoirement (chances de transmission).
x = np.random.rand()
if(x > gk[(i,j)]):
return 0 #arc non utilisé
else :
return 1 #arc utilisé
else:
x = np.random.rand()
if x > (gk[(i,j)]*np.exp(-gr[(i,j)]*(max_obs-times[i])))/(gk[(i,j)]*np.exp(-gr[(i,j)]*(max_obs-times[i]))+1-gk[(i,j)]):
return 1
else :
return 0
print(simulation(graph,[(0,0)],0))
np.random.seed(0)
print(simulation(graph,[(0,0)],2))
np.random.seed(0)
print(simulation(graph,[(0,0),(1,1)],1))
print(simulation(graph,[(0,0),(1,1)],1))
print(simulation(graph,[(0,0),(1,1)],1))
```
La méthode $getProbaMC(graph,infections,max\_obs,nbsimu)$ retourne les estimations de probabilités marginales d'infection des différents utilisateurs de $graph$, conditionnées à l'observation des infections de la liste $infections$ pendant les $max\_obs$ premiers jours de diffusion. Pour être enregistrée, une infection doit intervenir avant la seconde $maxT$. Ainsi, si la méthode retourne 0.2 pour le noeud $i$, cela indique qu'il a été infecté avec un temps $t_i \in ]max\_obs,maxT[$ dans 20% des $nbsimu$ simulations effectuées. Compléter la méthode ci dessous:
```
np.random.seed(0)
def getProbaMC(graph,infections,max_obs,nbsimu=100000):
names,gk,gr=graph
nbNodes=len(names)
rInf=np.array([0]*nbNodes)
#>>>>>>>>>>>
#lancer la simulation nbsimu fois
for i in range(nbsimu):
times = simulation(graph, infections, max_obs)
for node in range(nbNodes):
if times[node] < maxT and times[node] > max_obs: # si 𝑡𝑖∈]𝑚𝑎𝑥_𝑜𝑏𝑠,𝑚𝑎𝑥𝑇[
rInf[node]+= 1 #compter le nombre de fois
rInf = rInf/nbsimu #proba
#<<<<<<<<<<<
return (rInf)
rInf=getProbaMC(graph,[(0,0)],0)
print(rInf)
```
Cette méthode permet de bonnes estimations (malgré une certaine variance) lorsque l'on n'a pas d'observations autres que le vecteur d'infections initiales (i.e., on estime des probabilités de la forme: $P(t_i < maxT|\{t_j\leq max\_obs\})$). Par contre, si l'on souhaite obtenir des probabilités d'infection du type $P(t_i < maxT|\{t_j\leq max\_obs\}, \{t_j, j \in {\cal O}\})$, c'est à dire conditionnées à des observations supplémentaires pour un sous-ensembles de noeuds ${\cal O}$ (avec $t_j > max\_obs$ pour tout noeud de ${\cal O}$), l'utilisation de la méthode de MonteCarlo précédente est impossible. Cela impliquerait de filtrer les simulations obtenues selon qu'elles remplissent les conditions sur les noeuds de ${\cal O}$, ce qui nous amènerait à toutes les écarter sachant que l'on travaille avec des temps continus.
Pour estimer ce genre de probabilité conditionnelle, nous allons nous appuyer sur des méthodes de type MCMC, notamment la méthode de Gibbs Sampling. Cette méthode est utile pour simuler selon une loi jointe, lorsqu'il est plus simple d'échantillonner de chaque variable conditionnellement à toutes les autres plutôt que directement de cette loi jointe. L'algorithme est donné par:
1. Tirage d'un vecteur de valeurs initiales pour toutes les variables $X_i$
2. Pour toutes les variable $X_i$ choisies dans un ordre aléatoire, échantillonnage d'une nouvelle valeur: $X_i \sim p(x_i\mid x_1,\dots,x_{i-1},x_{i+1},\dots,x_n)$
3. Recommencer en 2 tant qu'on souhaite encore des échantillons
Notons qu'il est souvent utile d'exploiter la relation suivante, qui indique que pour échantillonner de la loi conditionnelle, il suffit d'échantillonner chaque variable proportionnellement à la loi jointe, avec toutes les autres variables fixées:
$$p(x_j\mid x_1,\dots,x_{j-1},x_{j+1},\dots,x_n) = \frac{p(x_1,\dots,x_n)}{p(x_1,\dots,x_{j-1},x_{j+1},\dots,x_n)} \propto p(x_1,\dots,x_n)$$
Après une période dite de $Burnin$ d'un nombre d'époques à définir, l'algorithme émet des échantillons qui suivent la loi jointe connaissant les observations. Lorsque l'objectif est d'estimer des probabilités marginales, on fait alors tourner cet algorithme pendant une certain nombre d'époques après la période de $Burnin$, au cours desquelles on recence les différentes affectations de chacune des variables étudiées.
Pour mettre en oeuvre cet algorithme, nous aurons aurons besoin d'avoir accès rapidement aux prédecesseurs et successeurs dans le graphe. La méthode ci-dessous retourne un couple de dictionnaires à partir du graphe:
* $preds[i]$ contient la liste des prédécesseurs du noeud $i$, sous la forme d'une liste de triplets $(j,k_{j,i},r_{j,i})$ pour tous les $j$ précédant $i$ dans le graphe.
* $succs[i]$ contient la liste des successeurs du noeud $i$, sous la forme d'une liste de triplets $(j,k_{i,j},r_{i,j})$ pour tous les $j$ pointés par $i$ dans le graphe.
```
def getPredsSuccs(graph):
names,gk,gr=graph
nbNodes=len(names)
preds={}
succs={}
for (a,b),v in gk.items():
s=succs.get(a,[])
s.append((b,v,gr[(a,b)]))
succs[a]=s
p=preds.get(b,[])
p.append((a,v,gr[(a,b)]))
preds[b]=p
return (preds,succs)
preds,succs=getPredsSuccs(graph)
print("preds=",preds)
print("succs=",succs)
```
Pour calculer les probabilités conditionnelles, il faut prendre en compte les quantités suivantes:
* Probabilité pour $j$ d'être infecté par $i$ au temps $t_j$ connaissant $t_i < t_j$:
$$\alpha_{i,j}=k_{i,j}r_{i,j} exp(-r_{i,j}(t_j-t_i))$$
* Probabilité pour $j$ de ne pas être infecté par $i$ jusqu'au temps $t$:
$$\beta_{i,j}=k_{i,j} exp(-r_{i,j}(t_j-t_i)) + 1 - k_{i,j}$$
* Probabilité pour $j$ d'être infecté au temps $t_j$ connaissant les prédecesseurs infectés avant $t_j$:
$$h_{j}=\prod_{i \in preds[j], t_i<t_j} \beta_{i,j} \sum_{i \in preds[i], t_i<t_j} \alpha_{i,j} / \beta_{i,j}$$
* Probabilité pour $j$ de ne pas être infecté avant $t_j=maxT$ connsaissant ses prédecesseurs infectés:
$$g_{j}=\prod_{i \in preds[j], t_i<t_j} \left(k_{i,j} exp(-r_{i,j}(maxT-t_i)) + 1 - k_{i,j}\right)=\prod_{i \in preds[j], t_i<t_j} \beta_{i,j}$$
A noter que pour tout $i$ tel que $t_i< max\_obs$, on prend $\frac{\alpha_{i,j}}{k_{i,j} exp(-r_{i,j}(max\_obs-t_i))}$ et $\frac{\beta_{i,j}}{k_{i,j} exp(-r_{i,j}(max\_obs-t_i))}$ dans les définitions de $h_j$ et $g_j$ (donc dans le calcul de la variable $b$ dans la méthode ci-dessous).
Dans la méthode $computeab(v, times, preds, max\_obs)$, on prépare le calcul et les mises à jour de ces quantités. La méthode calcule, pour un noeud $v$ selon les temps d'infection courants donnés dans $times$, deux quantités $a$ et $b$:
* $a= max(1e^{-20}, \sum_{i \in preds[v], t_i<t_v} \alpha_{i,v} / \beta_{i,v})$ si $t_v< maxT$ et $a=1$ sinon.
* $b=\sum_{i \in preds[v], t_i<t_v} \log \beta_{i,v}$.
Pour les noeuds $i$ tels que $t_i< max\_obs$, $a=1$ et $b=0$.
Compléter la méthode $computeab$ donnée ci-dessous:
```
eps=1e-20
def alpha(i,j,kij,rij,t):
return kij*rij*np.exp(-rij*(t[j]-t[i]))
def beta(i,j,kij,rij,t):
return kij*np.exp(-rij*(t[j]-t[i]))+1-kij
def beta2(i,j,kij,rij,t,max_obs): #gerer le cas ti < max_Obs
return kij*np.exp(-rij*(max_obs-t[i]))+1-kij
def computeab(v, times, preds, max_obs):
preds=preds.get(v,[]) #avoir les prédécesseur du noeud v
t=times[v] #avoir le temps de v
if t<=max_obs: #si v infecté
return (1,0)
a = eps
b=0
#si v a des prédécesseurs
if len(preds)>0:
#>>>>>>>>>>>>>
if(t<maxT): #si son t <maxT
sum_alpha = 0
for ind in range(len(preds)):
(i,kiv,riv) = preds[ind]
if(times[i] <t ):
sum_alpha += alpha(i,v,kiv,riv,times)/beta(i,v,kiv,riv,times)
if(times[i]<max_obs):
b+= np.log(beta(i,v,kiv,riv,times)) - np.log(beta2(i,v,kiv,riv,times,max_obs))
else:
b+= np.log(beta(i,v,kiv,riv,times))
a = max(eps, sum_alpha)
else:
a = 1
#<<<<<<<<<<<<
return (a,b)
nbNodes=len(graph[0])
times=np.array([maxT]*nbNodes,dtype=float)
times[0]=0
times[1]=1
times[2]=4
#times[3]=10
print(computeab(0,times,preds,max_obs=0))
print(computeab(0,times,preds,max_obs=2))
print(computeab(1,times,preds,max_obs=0))
print(computeab(1,times,preds,max_obs=2))
print(computeab(2,times,preds,max_obs=0))
print(computeab(2,times,preds,max_obs=2))
print(computeab(3,times,preds,max_obs=0))
print(computeab(3,times,preds,max_obs=2))
```
La méthode $computell$ calcule la log-vraisemblance d'une diffusion (représentée par le tableau times), en appelant la méthode computeab sur l'ensemble des noeuds du réseau. Elle retourne un triplet (log-likelihood, sa, sb), avec $sa$ et $sb$ les tables des valeurs $a$ et $b$ pour tous les noeuds.
```
def computell(times,preds,max_obs):
#>>>>>>
sa = []
sb = []
for v in range(len(times)) :
a,b = computeab(v,times,preds,max_obs)
sa.append(a)
sb.append(b)
ll = np.sum(np.log(sa)+sb)
return ll,sa,sb
#<<<<<
ll,sa,sb=computell(times,preds,max_obs=0)
print("ll=",ll)
print(times)
print("like_indiv=",np.exp(np.log(sa)+sb))
```
Afin de préparer les mises à jour lors des affectations successives des variables du Gibbs Sampling, on propose de définir une méthode $removeV(v,times,succs,sa,sb)$ qui retire temporairement du réseau un noeud $v$, en passant son temps d'infection à -1 dans times et en retirant sa contribution aux valeurs a et b (contenues dans sa et sb) de tous ses successeurs $j$ tels que $t_j > t_v$ (y compris donc les non infectés qui sont à $t_j=maxT$).
```
def removeV(v,times,succs,sa,sb):
succs=succs.get(v,[])
t=times[v]
if t<0:
return
times[v]=-1
sa[v]=1.0
sb[v]=0.0
if len(succs)>0:
c,k,r=map(np.array,zip(*succs))
tp=times[c]
which=(tp>t)
tp=tp[which]
dt=tp-t
k=k[which]
r=r[which]
c=c[which]
rt = -r*dt
b1=k*np.exp(rt)
b=b1+1.0-k
a=r*b1
a=a/b
b=np.log(b)
sa[c]=sa[c]-np.where(tp<maxT,a,0.0)
sa[c]=np.where(sa[c]>eps,sa[c],eps)
sb[c]=sb[c]-b
sb[c]=np.where(sb[c]>0,0,sb[c])
#Test
print("sa=",sa)
print("sb=",sb)
nsa=np.copy(sa)
nsb=np.copy(sb)
ntimes=np.copy(times)
removeV(3,ntimes,succs,nsa,nsb)
print("diffa=",nsa-sa)
print("diffb=",nsb-sb)
nsa=np.copy(sa)
nsb=np.copy(sb)
ntimes=np.copy(times)
removeV(1,ntimes,succs,nsa,nsb)
print("diffa=",nsa-sa)
print("diffb=",nsb-sb)
```
La méthode addVatT fait l'inverse: elle rajoute un noeud qui était retiré du réseau, avec un temps $newt$. Il faut alors mettre à jour les valeurs a et b (dans sa et sb) de tous les successeurs de $v$ tels que $t_j > newt$ et calculer les valeurs a et b du noeud v.
Compléter le code ci-dessous:
```
def addVatT(v,times,newt,preds,succs,sa,sb,max_obs):
t=times[v]
if t>=0:
raise Error("v must have been removed before")
#>>>>>>>>>>>>
times[v] = newt
sa[v],sb[v] = computeab(v, times, preds, max_obs)
for i in range(0,len(times)):
if(times[i]>newt):
sa[i],sb[i] = computeab(i, times, preds, max_obs)
#<<<<<<<<<<<<<<<<<<<
# Tests:
nsa=np.copy(sa)
nsb=np.copy(sb)
c,_,_=map(np.array,zip(*succs[1]))
c=np.append(c,1)
ll=np.sum((np.log(nsa)+nsb)[c])
removeV(1,times,succs,nsa,nsb)
addVatT(1,times,2,preds,succs,nsa,nsb,max_obs=0)
ll2=np.sum((np.log(nsa)+nsb)[c])
removeV(1,times,succs,nsa,nsb)
addVatT(1,times,1,preds,succs,nsa,nsb,max_obs=0)
ll3=np.sum((np.log(nsa)+nsb)[c])
llall=np.sum(np.log(nsa)+nsb)
print(np.exp(ll),np.exp(ll2),np.exp(ll3),llall)
c,_,_=map(np.array,zip(*succs[0]))
c=np.append(c,0)
ll=np.sum((np.log(nsa)+nsb)[c])
removeV(0,times,succs,nsa,nsb)
addVatT(0,times,maxT,preds,succs,nsa,nsb,max_obs=0)
ll2=np.sum((np.log(nsa)+nsb)[c])
removeV(0,times,succs,nsa,nsb)
addVatT(0,times,0,preds,succs,nsa,nsb,max_obs=0)
ll3=np.sum((np.log(nsa)+nsb)[c])
llall=np.sum(np.log(nsa)+nsb)
print(np.exp(ll),np.exp(ll2),np.exp(ll3),llall)
c,_,_=map(np.array,zip(*succs[5]))
c=np.append(c,5)
ll=np.sum((np.log(nsa)+nsb)[c])
removeV(5,times,succs,nsa,nsb)
addVatT(5,times,1,preds,succs,nsa,nsb,max_obs=0)
ll2=np.sum((np.log(nsa)+nsb)[c])
removeV(5,times,succs,nsa,nsb)
addVatT(5,times,maxT,preds,succs,nsa,nsb,max_obs=0)
ll3=np.sum((np.log(nsa)+nsb)[c])
llall=np.sum(np.log(nsa)+nsb)
print(np.exp(ll),np.exp(ll2),np.exp(ll3),llall)
```
Pour échantillonner pour une variable $i$, il faudra être à même de comparer les vraisemblances selon les différentes affectations. Cela implique de calculer la somme de toutes ces vraisemblances. Mais pour réaliser cette somme, il faudrait que nous sortions de la représentation logarithmique: $\sum_{t_i} exp(log(p(t_1,\dots,t_i,\dots,t_n))$. Si on le fait de cette manière, on risque d'avoir des arrondis à 0 presque partout. Une possibilité (log-sum-exp trick) est d'exploiter la relation suivante:
$$\log\sum_i x_i = x^* + \log\left( \exp(x_1-x^*)+ \cdots + \exp(x_n-x^*) \right)$$
avec $x^* = \max{\{x_1, \dots, x_n\}}$
Compléter la méthode logsumexp suivante, qui réalise cette somme en évitant les problèmes numériques:
```
def logsumexp(x,axis=-1):
#>>>>>>>>>>
star = np.max(x)
x = star + np.log(np.sum([np.exp(x[i]-star) for i in range(len(x))], axis))
#<<<<<<<<<<
return x
#Test:
x=np.array([[0.001,0.02,0.008],[0.1,0.01,0.4]])
r=np.log(np.sum(x,-1))
x=np.log(x)
r2=logsumexp(x)
print(r2,r)
```
On souhaite maintenant mettre en place une méthode $sampleV(v,times,newt,preds,succs,sa,sb,max\_obs,k,k2)$ qui sample un nouveau temps d'infection pour le noeud $v$, connaissant les temps de tous les autres noeuds dans $times$ (ainsi que leurs valeurs $a$ et $b$ correspondantes contenues dans sa et sb). Puisque le domaine de support de $t_v$ est continu, on doit faire quelques approximations en se basant sur une discrétisation des valeurs possibles:
1. On échantillonne $k$ points $d_1,\dots,d_k$ entre $max\_obs$ et $maxT$ de manière uniforme, que l'on ordonne de manière croissante. On ajoute $t_v$ à cet ensemble de points pour gagner en stabilité (inséré dans la liste de manière à conserver l'ordre croissant).
2. On considère chaque point $d_i$ comme le centre d'un bin $[(d_i+d_{i-1})/2,(d_i+d_{i+1})/2]$. Pour $d_1$ on prend $[max\_obs,(d_i+d_{i+1})/2]$ et pour $d_k$ on prend $[(d_i+d_{i-1})/2,maxT]$. On fait l'hypothèse que la densité de probabilité est constante sur l'ensemble du bin, que l'on évalue en son centre. La probabilité que l'on échantillonne dans le bin $i$ est alors égale à: $\frac{z_i \times l_i}{\sum_j z_j \times l_j + p_{maxT}}$, avec $z_i$ la densité calculée en $d_i$, $l_i$ la taille du bin $i$ et $p_{maxT}$ la probabilité calculée pour $maxT$ (i.e., la probabilité que $v$ ne soit pas infecté dans la diffusion)
3. Si l'indice $i$ samplé de manière proportionnelle aux probabilités calculées à l'étape précédente n'est pas celui de $maxT$, $v$ est alors infecté à un temps inclus dans l'intervale du bin correspondant. Il s'agit alors de re-échantillonner $k2$ points uniformément dans ce bin et de calculer les densités en ces points (pour gagner en stabilité on ajoute le centre du bin $d_i$). Le nouveau temps de $v$ est alors échantillonné proportionnellement à ces densités.
Compléter le code ci-dessous:
```
np.random.seed(0)
def sampleV(v,times,preds,succs,sa,sb,max_obs,k,k2):
#>>>>>>>>>>>>>>>>>>>>>>>>>>
#Votre code ici
#<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<
print(times)
sampleV(5,times,preds,succs,sa,sb,0,10,10)
print(times)
sampleV(5,times,preds,succs,sa,sb,0,10,10)
print(times)
sampleV(5,times,preds,succs,sa,sb,0,10,10)
print(times)
sampleV(5,times,preds,succs,sa,sb,0,10,10)
print(times)
sampleV(5,times,preds,succs,sa,sb,0,10,10)
print(times)
sampleV(5,times,preds,succs,sa,sb,0,10,10)
print(times)
sampleV(5,times,preds,succs,sa,sb,0,10,10)
print(times)
sampleV(5,times,preds,succs,sa,sb,0,10,10)
print(times)
```
Compléter la méthode de Gibbs Sampling $gb$ ci-dessous:
```
np.random.seed(1)
def gb(graph,infections,max_obs,burnin=1000,nbEpochs=10000,k=100,k2=50,freqRecord=1):
#>>>>>>>>>>>>>>>>>>>>>>>
#Votre code ici
#>>>>>>>>>>>>>>>>>>>>>>>>>>
return rate
rate=gb(graph,[(0,0)],0)
print(rate)
```
# Partie optionnelle
L'algorithme de Metropolis-Hasting est une autre méthode de type MCMC qui utilise une distribution d'échantillonnage pour se déplacer dans l'espace des points considérés. Il s'agit de définir une distribution $q(y_{t+1}|x_t)$ de laquelle on sait générer un déplacement. L'algorithme procéde alors de la manière suivante:
1. Générer $y_{t+1}$ selon $q(y_{t+1}|x_t)$
2. Calculer la probabilité d’acceptation $\alpha(x_t,y_{t+1})=\min\left\{\frac{\pi(y_{t+1})q(x_t,y_{t+1})}{\pi(x_t)q(y_{t+1},x_t)},1\right\} \,\!, \text{ avec } \pi(x_t) \text{ la densité de probabilité de } x_t$
3. Prendre $x_{t+1}=\begin{cases} y_{t+1}, & \text{avec probabilité}\,\,\alpha \\ x_t, & \text{avec probabilité}\,\,1-\alpha \end{cases}$
Dans notre cas, on propose de travailler avec des déplacements correspondants à des permutations d'un temps d'infection à chaque itération, comme dans le cadre du Gibbs Sampling. A chaque étape on choisit donc une variable à modifier, on choisit un nouveau temps pour cette variable et on calcule la densité correspondante. La probabilité d'acceptation est ensuite calculée selon cette densité et la probabilité du déplacement selon la distribution $q$ qui a servi à générer le nouveau temps d'infection. On se propose de choisir $maxT$ avec une probabilité de 0.1. La probabilité $q(t_v|t)$ pour $t< maxT$ est alors égale à $0.9\times \frac{1}{maxT-max\_obs}$.
Implémenter l'approche d'échantillonnage par Metropolis-Hasting pour notre problème d'estimation de probabilités marginales d'infection.
```
#Votre code ici
```
| github_jupyter |
# Obtain mean head pose on *trainval* split and use as predictions for test dataset
Implicitly documents test submission format
```
import os
from dd_pose.dataset import Dataset
from dd_pose.dataset_item import DatasetItem
from dd_pose.image_decorator import ImageDecorator
from dd_pose.jupyter_helpers import showimage
from dd_pose.evaluation_helpers import T_headfrontal_camdriver, T_camdriver_headfrontal
import transformations as tr
import json
import numpy as np
class MeanPredictor:
def __init__(self):
self.mean_T_camdriver_head = None
def get_name(self):
return 'Prior'
def get_dirname(self):
return 'prior'
def get_metadata(self):
return dict()
def initialize_from_dataset(self, dataset):
# get mean translation and rotation across the whole dataset
# we compute roll, pitch, yaw wrt. 'headfrontal' frame in order to avoid gimbal lock averaging later on
xyzs = []
rpys = []
for di_dict in dataset.get_dataset_items():
di = DatasetItem(di_dict)
print(di)
for stamp in di.get_stamps():
T_camdriver_head = di.get_T_camdriver_head(stamp)
T_headfrontal_head = np.dot(T_headfrontal_camdriver, T_camdriver_head)
rpy = tr.euler_from_matrix(T_headfrontal_head)
rpys.append(rpy)
xyzs.append(T_camdriver_head[0:3,3])
# rpy mean in headfrontal frame
mean_rpy = np.mean(np.array(rpys), axis=0)
print(mean_rpy)
# xyz mean in camdriver frame
mean_xyz = np.mean(np.array(xyzs), axis=0)
print(mean_xyz)
# rotational component from mean rpy to camdriver frame
mean_T_headfrontal_head = tr.euler_matrix(*mean_rpy)
self.mean_T_camdriver_head = np.dot(T_camdriver_headfrontal, mean_T_headfrontal_head)
# translational component from mean xyz in camdriver frame
self.mean_T_camdriver_head[0:3,3] = mean_xyz
def get_T_camdriver_head(self, stamp):
return self.mean_T_camdriver_head
# initialize mean predictor with measurements from trainval split
mean_predictor = MeanPredictor()
mean_predictor.initialize_from_dataset(Dataset(split='trainval'))
mean_predictor.get_T_camdriver_head(0)
```
# Draw mean head pose onto sample image
```
d = Dataset(split='trainval')
di_dict = d.get(subject_id=1, scenario_id=3, humanhash='sodium-finch-fillet-spring')
di = DatasetItem(di_dict)
print(di)
stamp = di.get_stamps()[50]
stamp
img, pcm = di.get_img_driver_left(stamp, shift=True)
img_bgr = np.dstack((img, img, img))
image_decorator = ImageDecorator(img_bgr, pcm)
image_decorator.draw_axis(mean_predictor.get_T_camdriver_head(stamp), use_gray=False)
image_decorator.draw_axis(di.get_T_camdriver_head(stamp), use_gray=True)
```
colored axis: mean head pose
gray axis: measurement (ground truth)
```
showimage(img_bgr)
prediction_output_base_dir = os.path.join(os.environ['DD_POSE_DATA_ROOT_DIR'], '10-predictions')
try:
os.makedirs(prediction_output_base_dir)
except:
pass
```
# Write out mean head pose predictions for test dataset
```
d = Dataset(split='test')
predictor = mean_predictor
predictor.get_name(), predictor.get_dirname()
predictor_predictions_dir = os.path.join(prediction_output_base_dir, predictor.get_dirname())
assert not os.path.exists(predictor_predictions_dir), "Predictions already written out. Aborting. %s" % predictor_predictions_dir
for di_dict in d.get_dataset_items():
di = DatasetItem(di_dict)
print(di)
predictions_dir = os.path.join(predictor_predictions_dir, 'subject-%02d' % di.get_subject(), 'scenario-%02d' % di.get_scenario(), di.get_humanhash())
try:
os.makedirs(predictions_dir)
except OSError as e:
pass
predictions = dict()
for stamp in di.get_stamps():
predictions[stamp] = predictor.get_T_camdriver_head(stamp).tolist()
# write out predictions
with open(os.path.join(predictions_dir, 't-camdriver-head-predictions.json'), 'w') as fp:
json.dump(predictions, fp, sort_keys=True, indent=4)
metadata = {
'name': predictor.get_name(),
'dirname': predictor.get_dirname(),
'metadata': predictor.get_metadata()
}
with open(os.path.join(predictor_predictions_dir, 'metadata.json'), 'w') as fp:
json.dump(metadata, fp, sort_keys=True, indent=4)
# now tar.gz the predictions in the format expected by the benchmark website
print('pushd %s; tar czf %s subject-* metadata.json; popd' % (predictor_predictions_dir,\
os.path.join(predictor_predictions_dir, 'predictions.tar.gz')))
```
| github_jupyter |
# DNA Sequence Statistics
## Reading Data
```
import screed # A Python library for reading FASTA and FASQ file format.
def readFastaFile(inputfile):
"""
Reads and returns file as FASTA format with special characters removed.
"""
with screed.open(inputfile) as seqfile:
for read in seqfile:
seq = read.sequence
return seq
```
## Read FASTA/FASQ File
```
den1 = readFastaFile("../data/den1.fasta")
den2 = readFastaFile("../data/den2.fasta")
den3 = readFastaFile("../data/den3.fasta")
den4 = readFastaFile("../data/den4.fasta")
```
## Statistical Aanalysis
### Length of a DNA sequence
```
# length of DEN1
len(den1)
# length of DEN2
len(den2)
# length of DEN3
len(den3)
len(den4)
```
## Base composition of a DNA sequence¶
```
def count_base(seq):
base_counts = {}
for base in seq:
if base in base_counts:
base_counts[base] += 1
else:
base_counts[base] = 1
return base_counts
# Let's call the function count_base
counts = count_base(den1)
print(counts)
from collections import Counter
def count_base(seq):
base_counts = Counter(seq)
return base_counts
bases = count_base(den1)
print(bases)
```
### Frequency with Counter
```
from collections import Counter
# Base composition of DEN1
freq1 = Counter(den1)
print(freq1)
# Base composition of DEN2
freq2 = Counter(den2)
print(freq2)
# Base composition of DEN3
freq3 = Counter(den3)
print(freq3)
# Base composition of DEN4
freq4 = Counter(den4)
print(freq2)
```
## GC Content of DNA
```
import screed # A Python library for reading FASTA and FASQ file format.
def readFastaFile(inputfile):
"""
Reads and returns file as FASTA format with special characters removed.
"""
with screed.open(inputfile) as seqfile:
for read in seqfile:
seq = read.sequence
return seq
def calculate_gc_content(seq):
"""
Take DNA sequence as input and calculate the GC content.
"""
no_of_g = seq.count("G")
no_of_c = seq.count("C")
total = no_of_g + no_of_c
gc = total/len(seq) * 100
return gc
# Calculate GC Content of DEN1
den1 = readFastaFile("../data/den1.fasta")
result1 = calculate_gc_content(den1)
print(result1)
# Calculate GC Content of DEN2
den2 = readFastaFile("../data/den2.fasta")
result2 = calculate_gc_content(den2)
print(result2)
# Calculate GC Content of DEN3
den3 = readFastaFile("../data/den3.fasta")
result3 = calculate_gc_content(den3)
print(result3)
# Calculate GC Content of DEN4
den4 = readFastaFile("../data/den4.fasta")
result4 = calculate_gc_content(den4)
print(result4)
```
## AT Content of DNA
```
import screed # A Python library for reading FASTA and FASQ file format.
def readFastaFile(inputfile):
"""
Reads and returns file as FASTA format with special characters removed.
"""
with screed.open(inputfile) as seqfile:
for read in seqfile:
seq = read.sequence
return seq
def calculate_at_content(seq):
"""
Take DNA sequence as input and calculate the AT content.
"""
no_of_a = seq.count("A")
no_of_t = seq.count("T")
total = no_of_a + no_of_t
at = total/len(seq) * 100
return at
# Calculate AT Content of DEN1
den1 = readFastaFile("../data/den1.fasta")
result1 = calculate_at_content(den1)
print(result1)
# Calculate AT Content of DEN2
den2 = readFastaFile("../data/den2.fasta")
result2 = calculate_at_content(den2)
print(result2)
# Calculate AT Content of DEN3
den3 = readFastaFile("../data/den3.fasta")
result3 = calculate_at_content(den3)
print(result3)
# Calculate AT Content of DEN4
den4 = readFastaFile("../data/den4.fasta")
result4 = calculate_at_content(den4)
print(result4)
```
## A sliding window analysis of GC content
```
import screed # A Python library for reading FASTA and FASQ file format.
def readFastaFile(inputfile):
"""
Reads and returns file as FASTA format with special characters removed.
"""
with screed.open(inputfile) as seqfile:
for read in seqfile:
seq = read.sequence
return seq
def calculate_gc_content(seq):
"""
Take DNA sequence as input and calculate the GC content.
"""
no_of_g = seq.count("G")
no_of_c = seq.count("C")
total = no_of_g + no_of_c
gc = total/len(seq) * 100
return gc
# Calculate the GC content of nucleotides 1-2000 of the Dengue genome(DEN-1)
seq = readFastaFile("../data/den1.fasta")
calculate_gc_content(seq[1:2001])
# Calculate the GC content of nucleotides 2001-4000 of the Dengue genome(DEN-1)
seq = readFastaFile("../data/den1.fasta")
calculate_gc_content(seq[2001:4001])
# Calculate the GC content of nucleotides 4001-6000 of the Dengue genome(DEN-1)
seq = readFastaFile("../data/den1.fasta")
calculate_gc_content(seq[4001:6001])
# Calculate the GC content of nucleotides 6001-8000 of the Dengue genome(DEN-1)
seq = readFastaFile("../data/den1.fasta")
calculate_gc_content(seq[6001:8001])
# Calculate the GC content of nucleotides 8001-10000 of the Dengue genome(DEN-1)
seq = readFastaFile("../data/den1.fasta")
calculate_gc_content(seq[8001:10001])
# Calculate the GC content of nucleotides 10001-10735 of the Dengue genome(DEN-1)
seq = readFastaFile("../data/den1.fasta")
calculate_gc_content(seq[10001:10735])
```
## GC content of the non-overlapping sub-sequences of size k
```
def calculate_gc(seq):
"""
Returns percentage of G and C nucleotides in a DNA sequence.
"""
gc = 0
for base in seq:
if base in "GCgc":
gc += 1
else:
gc = 1
return gc/len(seq) * 100
def gc_subseq(seq, k=2000):
"""
Returns GC content of non − overlapping sub− sequences of size k.
The result is a list.
"""
res = []
for i in range(0, len(seq)-k+1, k):
subseq = seq[i:i+k]
gc = calculate_gc(subseq)
res.append(gc)
return gc
seq = readFastaFile("../data/den1.fasta")
gc_subseq(seq, 2000)
```
## K-mer Analysis
```
def build_kmers(sequence, ksize):
kmers = []
n_kmers = len(sequence) - ksize + 1
for i in range(n_kmers):
kmer = sequence[i:i + ksize]
kmers.append(kmer)
return kmers
seq = readFastaFile("../data/den1.fasta")
# Dimer
km2 = build_kmers(seq, 2)
# Count dimer
from collections import Counter
dimer = Counter(km2)
print(dimer)
# Trimer
km3 = build_kmers(seq, 3)
# Count trimer
trimer = Counter(km3)
print(trimer)
```
| github_jupyter |
```
#!/usr/bin/env python3
import os
import re
import sys
import collections
import argparse
import tables
import itertools
import matplotlib
import glob
import math
%matplotlib inline
import matplotlib.pyplot as plt
import matplotlib.gridspec as gridspec
import numpy as np
import gseapy as gp
import pandas as pd
import scipy.stats as stats
import scipy.sparse as sp_sparse
from multiprocessing import Pool
from collections import defaultdict
from scipy import sparse, io
from scipy.sparse import csr_matrix
from multiprocessing import Pool
from matplotlib_venn import venn2, venn2_circles
matplotlib.rcParams['pdf.fonttype'] = 42
matplotlib.rcParams['ps.fonttype'] = 42
gp.__version__
```
### Loading Data
```
#load data, specify the column names
columns = [
'NAME','PZ800','PZ801','PZ802','PZ803',
'PZ804','PZ805','PZ806','PZ807','PZ808',
'PZ809','PZ810','PZ811','PZ812','PZ813',
'PZ814','PZ815','PZ816','PZ817','PZ818',
'PZ819','PZ820','PZ821','PZ822','PZ823'
]
data = pd.read_csv('./FeatureCounts_MYB_raw.txt', sep = '\t', header = None)
data.columns = columns
des_col = ['na'] * 56833
data.insert(loc=1, column='DESCRIPTION', value=des_col)
gene_index = (np.sum(data[data.columns[data.columns.str.startswith('PZ')]], axis=1) > 100).values
#matrix normalization
data_matrix = data[data.columns[data.columns.str.startswith('PZ')]]
sample_sum = np.sum(data_matrix, axis=0).values
normalized_mtx = data_matrix.divide(sample_sum) * 1e6
data.iloc[:,2:] = normalized_mtx
data_filtered = data.iloc[gene_index,:]
```
### Run GSEA
```
gs_res = gp.gsea(data=data_filtered, # or data='./P53_resampling_data.txt'
gene_sets='./Genesets/IVANOVA_HEMATOPOIESIS_MATURE_CELL_geneset.gmt', # enrichr library names
cls= './Phenotype_MYB.cls', # cls=class_vector
#set permutation_type to phenotype if samples >=15
permutation_type='phenotype',
permutation_num=1000, # reduce number to speed up test
outdir=None, # do not write output to disk
no_plot=True, # Skip plotting
method='signal_to_noise',
processes=4)
from gseapy.plot import gseaplot, heatmap
terms = gs_res.res2d.index
fig = gseaplot(
gs_res.ranking,
term=terms[0],
**gs_res.results[terms[0]],
# ofname='./IVANOVA-GSEA.pdf'
)
# plotting heatmap
genes = gs_res.res2d.genes[0].split(";")
heatmap(
df = gs_res.heatmat.loc[genes],
z_score=0,
title=terms[0],
figsize=(8,80),
# ofname='IVANOVA-heatmap.pdf'
)
gs_res.res2d
```
| github_jupyter |
<table class="ee-notebook-buttons" align="left">
<td><a target="_blank" href="https://github.com/giswqs/earthengine-py-notebooks/tree/master/Datasets/Terrain/srtm_landforms.ipynb"><img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" /> View source on GitHub</a></td>
<td><a target="_blank" href="https://nbviewer.jupyter.org/github/giswqs/earthengine-py-notebooks/blob/master/Datasets/Terrain/srtm_landforms.ipynb"><img width=26px src="https://upload.wikimedia.org/wikipedia/commons/thumb/3/38/Jupyter_logo.svg/883px-Jupyter_logo.svg.png" />Notebook Viewer</a></td>
<td><a target="_blank" href="https://colab.research.google.com/github/giswqs/earthengine-py-notebooks/blob/master/Datasets/Terrain/srtm_landforms.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" /> Run in Google Colab</a></td>
</table>
## Install Earth Engine API and geemap
Install the [Earth Engine Python API](https://developers.google.com/earth-engine/python_install) and [geemap](https://github.com/giswqs/geemap). The **geemap** Python package is built upon the [ipyleaflet](https://github.com/jupyter-widgets/ipyleaflet) and [folium](https://github.com/python-visualization/folium) packages and implements several methods for interacting with Earth Engine data layers, such as `Map.addLayer()`, `Map.setCenter()`, and `Map.centerObject()`.
The following script checks if the geemap package has been installed. If not, it will install geemap, which automatically installs its [dependencies](https://github.com/giswqs/geemap#dependencies), including earthengine-api, folium, and ipyleaflet.
**Important note**: A key difference between folium and ipyleaflet is that ipyleaflet is built upon ipywidgets and allows bidirectional communication between the front-end and the backend enabling the use of the map to capture user input, while folium is meant for displaying static data only ([source](https://blog.jupyter.org/interactive-gis-in-jupyter-with-ipyleaflet-52f9657fa7a)). Note that [Google Colab](https://colab.research.google.com/) currently does not support ipyleaflet ([source](https://github.com/googlecolab/colabtools/issues/60#issuecomment-596225619)). Therefore, if you are using geemap with Google Colab, you should use [`import geemap.eefolium`](https://github.com/giswqs/geemap/blob/master/geemap/eefolium.py). If you are using geemap with [binder](https://mybinder.org/) or a local Jupyter notebook server, you can use [`import geemap`](https://github.com/giswqs/geemap/blob/master/geemap/geemap.py), which provides more functionalities for capturing user input (e.g., mouse-clicking and moving).
```
# Installs geemap package
import subprocess
try:
import geemap
except ImportError:
print('geemap package not installed. Installing ...')
subprocess.check_call(["python", '-m', 'pip', 'install', 'geemap'])
# Checks whether this notebook is running on Google Colab
try:
import google.colab
import geemap.eefolium as geemap
except:
import geemap
# Authenticates and initializes Earth Engine
import ee
try:
ee.Initialize()
except Exception as e:
ee.Authenticate()
ee.Initialize()
```
## Create an interactive map
The default basemap is `Google Maps`. [Additional basemaps](https://github.com/giswqs/geemap/blob/master/geemap/basemaps.py) can be added using the `Map.add_basemap()` function.
```
Map = geemap.Map(center=[40,-100], zoom=4)
Map
```
## Add Earth Engine Python script
```
# Add Earth Engine dataset
dataset = ee.Image('CSP/ERGo/1_0/Global/SRTM_landforms')
landforms = dataset.select('constant')
landformsVis = {
'min': 11.0,
'max': 42.0,
'palette': [
'141414', '383838', '808080', 'EBEB8F', 'F7D311', 'AA0000', 'D89382',
'DDC9C9', 'DCCDCE', '1C6330', '68AA63', 'B5C98E', 'E1F0E5', 'a975ba',
'6f198c'
],
}
Map.setCenter(-105.58, 40.5498, 11)
Map.addLayer(landforms, landformsVis, 'Landforms')
```
## Display Earth Engine data layers
```
Map.addLayerControl() # This line is not needed for ipyleaflet-based Map.
Map
```
| github_jupyter |
<h1 style="color:#3da1da;font-size: 300%;" id="timeshap tutorial" align="center" >TimeSHAP Tutorial - Pytorch - AReM dataset</h1><p>
<a id='top_cell'></a>
## Table of contents
1. [Data Processing](#1.-Data-Processing)
1. [Data Loading](#1.1-Data-Loading)
2. [Data Treatment](#1.2-Data-Treatment)
2. [Model](#2.-Model)
1. [Model Definition](#2.1-Model-Definition)
2. [Model Training](#2.2-Model-Training)
3. [TimeSHAP](#3.-TimeSHAP)
1. [Local Explanations](#3.1-Local-Explanations)
2. [Global Explanations](#3.2-Global-Explanations)
3. [Individual Plots](#3.3-Individual-Plots)
# TimeSHAP
TimeSHAP is a model-agnostic, recurrent explainer that builds upon KernelSHAP and extends it to the sequential domain.
TimeSHAP computes local event/timestamp- feature-, and cell-level attributions.
Aditionally TimeSHAP also computes global event- and feature-level explanations.
As sequences can be arbitrarily long, TimeSHAP also implements a pruning algorithm based on Shapley Values,
that finds a subset of consecutive, recent events that contribute the most to the decision.
---
# 1. Data-Processing
---
```
import pandas as pd
import numpy as np
np.random.seed(42)
import warnings
warnings.filterwarnings('ignore')
from timeshap import __version__
__version__
import altair as alt
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:100% !important; }</style>"))
from timeshap.plot import timeshap_theme
alt.themes.register("timeshap_theme", timeshap_theme)
alt.themes.enable("timeshap_theme")
```
## 1.1 Data-Loading
```
import os
import re
data_directories = next(os.walk("./AReM"))[1]
all_csvs = []
for folder in data_directories:
if folder in ['bending1', 'bending2']:
continue
folder_csvs = next(os.walk(f"./AReM/{folder}"))[2]
for data_csv in folder_csvs:
if data_csv == 'dataset8.csv' and folder == 'sitting':
# this dataset only has 479 instances
# it is possible to use it, but would require padding logic
continue
loaded_data = pd.read_csv(f"./AReM/{folder}/{data_csv}", skiprows=4)
print(f"{folder}/{data_csv} ------ {loaded_data.shape}")
csv_id = re.findall(r'\d+', data_csv)[0]
loaded_data['id'] = csv_id
loaded_data['all_id'] = f"{folder}_{csv_id}"
loaded_data['activity'] = folder
all_csvs.append(loaded_data)
all_data = pd.concat(all_csvs)
raw_model_features = ['avg_rss12', 'var_rss12', 'avg_rss13', 'var_rss13', 'avg_rss23', 'var_rss23']
all_data.columns = ['timestamp', 'avg_rss12', 'var_rss12', 'avg_rss13', 'var_rss13', 'avg_rss23', 'var_rss23', 'id', 'all_id', 'activity']
```
## 1.2 Data Treatment
### Separate in train and test
```
# choose ids to use for test
ids_for_test = np.random.choice(all_data['id'].unique(), size=4, replace=False)
d_train = all_data[~all_data['id'].isin(ids_for_test)]
d_test = all_data[all_data['id'].isin(ids_for_test)]
```
### Normalize Features
```
class NumericalNormalizer:
def __init__(self, fields: list):
self.metrics = {}
self.fields = fields
def fit(self, df: pd.DataFrame ) -> list:
means = df[self.fields].mean()
std = df[self.fields].std()
for field in self.fields:
field_mean = means[field]
field_stddev = std[field]
self.metrics[field] = {'mean': field_mean, 'std': field_stddev}
def transform(self, df: pd.DataFrame) -> pd.DataFrame:
# Transform to zero-mean and unit variance.
for field in self.fields:
f_mean = self.metrics[field]['mean']
f_stddev = self.metrics[field]['std']
# OUTLIER CLIPPING to [avg-3*std, avg+3*avg]
df[field] = df[field].apply(lambda x: f_mean - 3 * f_stddev if x < f_mean - 3 * f_stddev else x)
df[field] = df[field].apply(lambda x: f_mean + 3 * f_stddev if x > f_mean + 3 * f_stddev else x)
if f_stddev > 1e-5:
df[f'p_{field}_normalized'] = df[field].apply(lambda x: ((x - f_mean)/f_stddev))
else:
df[f'p_{field}_normalized'] = df[field].apply(lambda x: x * 0)
return df
#all features are numerical
normalizor = NumericalNormalizer(raw_model_features)
normalizor.fit(d_train)
d_train_normalized = normalizor.transform(d_train)
d_test_normalized = normalizor.transform(d_test)
```
### Features
```
model_features = [f"p_{x}_normalized" for x in raw_model_features]
time_feat = 'timestamp'
label_feat = 'activity'
sequence_id_feat = 'all_id'
plot_feats = {
'p_avg_rss12_normalized': "Mean Chest <-> Right Ankle",
'p_var_rss12_normalized': "STD Chest <-> Right Ankle",
'p_avg_rss13_normalized': "Mean Chest <-> Left Ankle",
'p_var_rss13_normalized': "STD Chest <-> Left Ankle",
'p_avg_rss23_normalized': "Mean Right Ankle <-> Left Ankle",
'p_var_rss23_normalized': "STD Right Ankle <-> Left Ankle",
}
```
### Transform the dataset from multi-label to binary classification
```
# possible activities ['cycling', 'lying', 'sitting', 'standing', 'walking']
#Select the activity to predict
chosen_activity = 'cycling'
d_train_normalized['label'] = d_train_normalized['activity'].apply(lambda x: int(x == chosen_activity))
d_test_normalized['label'] = d_test_normalized['activity'].apply(lambda x: int(x == chosen_activity))
```
This example notebook requires PyTorch!
Install it if you haven't already:
```
!pip install torch
```
```
import torch
def df_to_Tensor(df, model_feats, label_feat, group_by_feat, timestamp_Feat):
sequence_length = len(df[timestamp_Feat].unique())
data_tensor = np.zeros((len(df[group_by_feat].unique()), sequence_length, len(model_feats)))
labels_tensor = np.zeros((len(df[group_by_feat].unique()), 1))
for i, name in enumerate(df[group_by_feat].unique()):
name_data = df[df[group_by_feat] == name]
sorted_data = name_data.sort_values(timestamp_Feat)
data_x = sorted_data[model_feats].values
labels = sorted_data[label_feat].values
assert labels.sum() == 0 or labels.sum() == len(labels)
data_tensor[i, :, :] = data_x
labels_tensor[i, :] = labels[0]
data_tensor = torch.from_numpy(data_tensor).type(torch.FloatTensor)
labels_tensor = torch.from_numpy(labels_tensor).type(torch.FloatTensor)
return data_tensor, labels_tensor
train_data, train_labels = df_to_Tensor(d_train_normalized, model_features, 'label', sequence_id_feat, time_feat)
test_data, test_labels = df_to_Tensor(d_test_normalized, model_features, 'label', sequence_id_feat, time_feat)
```
___
# 2. Model
## 2.1 Model Definition
```
import torch.nn as nn
class ExplainedRNN(nn.Module):
def __init__(self,
input_size: int,
cfg: dict,
):
super(ExplainedRNN, self).__init__()
self.hidden_dim = cfg.get('hidden_dim', 32)
torch.manual_seed(cfg.get('random_seed', 42))
self.recurrent_block = nn.GRU(
input_size=input_size,
hidden_size=self.hidden_dim,
batch_first=True,
num_layers=2,
)
self.classifier_block = nn.Linear(self.hidden_dim, 1)
self.output_activation_func = nn.Sigmoid()
def forward(self,
x: torch.Tensor,
hidden_states: tuple = None,
):
if hidden_states is None:
output, hidden = self.recurrent_block(x)
else:
output, hidden = self.recurrent_block(x, hidden_states)
# -1 on hidden, to select the last layer of the stacked gru
assert torch.equal(output[:,-1,:], hidden[-1, :, :])
y = self.classifier_block(hidden[-1, :, :])
y = self.output_activation_func(y)
return y, hidden
import torch.optim as optim
model = ExplainedRNN(len(model_features), {})
loss_function = nn.BCELoss()
optimizer = optim.Adam(model.parameters(), lr=0.01)
learning_rate = 0.005
EPOCHS = 8
```
## 2.2 Model Training
```
import tqdm
import copy
for epoch in tqdm.trange(EPOCHS):
train_data_local = copy.deepcopy(train_data)
train_labels_local = copy.deepcopy(train_labels)
y_pred, hidden_states = model(train_data_local)
train_loss = loss_function(y_pred, train_labels_local)
optimizer.zero_grad()
train_loss.backward()
optimizer.step()
with torch.no_grad():
test_data_local = copy.deepcopy(test_data)
test_labels_local = copy.deepcopy(test_labels)
test_preds, _ = model(test_data_local)
test_loss = loss_function(test_preds, test_labels_local)
print(f"Train loss: {train_loss.item()} --- Test loss {test_loss.item()} ")
```
---
# 3. TimeSHAP
---
### Model entry point
```
from timeshap.wrappers import TorchModelWrapper
model_wrapped = TorchModelWrapper(model)
f_hs = lambda x, y=None: model_wrapped.predict_last_hs(x, y)
```
### Baseline event
```
from timeshap.utils import calc_avg_event
average_event = calc_avg_event(d_train_normalized, numerical_feats=model_features, categorical_feats=[])
```
### Average score over baseline
```
from timeshap.utils import get_avg_score_with_avg_event
avg_score_over_len = get_avg_score_with_avg_event(f_hs, average_event, top=480)
```
## 3.1 Local Explanations
### Select sequences to explain
```
positive_sequence_id = f"cycling_{np.random.choice(ids_for_test)}"
pos_x_pd = d_test_normalized[d_test_normalized['all_id'] == positive_sequence_id]
# select model features only
pos_x_data = pos_x_pd[model_features]
# convert the instance to numpy so TimeSHAP receives it
pos_x_data = np.expand_dims(pos_x_data.to_numpy().copy(), axis=0)
```
### Local Report on positive instance
```
from timeshap.explainer import local_report
pruning_dict = {'tol': 0.025}
event_dict = {'rs': 42, 'nsamples': 32000}
feature_dict = {'rs': 42, 'nsamples': 32000, 'feature_names': model_features, 'plot_features': plot_feats}
cell_dict = {'rs': 42, 'nsamples': 32000, 'top_x_feats': 2, 'top_x_events': 2}
local_report(f_hs, pos_x_data, pruning_dict, event_dict, feature_dict, cell_dict=cell_dict, entity_uuid=positive_sequence_id, entity_col='all_id', baseline=average_event)
```
## 3.2 Global Explanations
### Explain all
TimeSHAP offers methods to explain all instances and save as CSV.
This allows for global explanations and local plots with no calculation delay.
```
from timeshap.explainer import global_report
pos_dataset = d_test_normalized[d_test_normalized['label'] == 1]
pruning_dict = {'tol': [0.05, 0.075, 0.076], 'path': 'outputs/prun_all.csv'}
event_dict = {'path': 'outputs/event_all.csv', 'rs': 42, 'nsamples': 32000}
feature_dict = {'path': 'outputs/feature_all.csv', 'rs': 42, 'nsamples': 32000, 'feature_names': model_features, 'plot_features': plot_feats,}
prun_stats, global_plot = global_report(f_hs, pos_dataset, pruning_dict, event_dict, feature_dict, sequence_id_feat, time_feat, model_features, baseline=average_event)
prun_stats
global_plot
```
## 3.3 Individual Plots
### Local Plots
```
from timeshap.plot import plot_temp_coalition_pruning, plot_event_heatmap, plot_feat_barplot, plot_cell_level
from timeshap.explainer import local_pruning, local_event, local_feat, local_cell_level
# select model features only
pos_x_data = pos_x_pd[model_features]
# convert the instance to numpy so TimeSHAP receives it
pos_x_data = np.expand_dims(pos_x_data.to_numpy().copy(), axis=0)
```
##### Pruning algorithm
```
pruning_dict = {'tol': 0.025,}
coal_plot_data, coal_prun_idx = local_pruning(f_hs, pos_x_data, pruning_dict, average_event, positive_sequence_id, sequence_id_feat, False)
# coal_prun_idx is in negative terms
pruning_idx = pos_x_data.shape[1] + coal_prun_idx
pruning_plot = plot_temp_coalition_pruning(coal_plot_data, coal_prun_idx, plot_limit=40)
pruning_plot
```
##### Event-level explanation
```
event_dict = {'rs': 42, 'nsamples': 32000}
event_data = local_event(f_hs, pos_x_data, event_dict, positive_sequence_id, sequence_id_feat, average_event, pruning_idx)
event_plot = plot_event_heatmap(event_data)
event_plot
```
##### Feature-level explanation
```
feature_dict = {'rs': 42, 'nsamples': 32000, 'feature_names': model_features, 'plot_features': plot_feats}
feature_data = local_feat(f_hs, pos_x_data, feature_dict, positive_sequence_id, sequence_id_feat, average_event, pruning_idx)
feature_plot = plot_feat_barplot(feature_data, feature_dict.get('top_feats'), feature_dict.get('plot_features'))
feature_plot
```
##### Cell-level explanation
```
cell_dict = {'rs': 42, 'nsamples': 32000, 'top_x_events': 3, 'top_x_feats': 3}
cell_data = local_cell_level(f_hs, pos_x_data, cell_dict, event_data, feature_data, positive_sequence_id, sequence_id_feat, average_event, pruning_idx)
feat_names = list(feature_data['Feature'].values)[:-1] # exclude pruned events
cell_plot = plot_cell_level(cell_data, feat_names, feature_dict.get('plot_features'))
cell_plot
```
### Global Plots
```
from timeshap.explainer import prune_all, pruning_statistics, event_explain_all, feat_explain_all
from timeshap.plot import plot_global_event, plot_global_feat
pos_dataset = d_test_normalized[d_test_normalized['label'] == 1]
```
##### Pruning statistics
```
pruning_dict = {'tol': [0.05, 0.075, 0.076], 'path': 'outputs/prun_all.csv'}
prun_indexes = prune_all(f_hs, pos_dataset, sequence_id_feat, average_event, pruning_dict, model_features, time_feat)
pruning_stats = pruning_statistics(prun_indexes, pruning_dict.get('tol'), sequence_id_feat)
pruning_stats
```
##### Global event-level
```
event_dict = {'path': 'outputs/event_all.csv', 'rs': 42, 'nsamples': 32000}
event_data = event_explain_all(f_hs, pos_dataset, sequence_id_feat, average_event, event_dict, prun_indexes, model_features, time_feat)
event_global_plot = plot_global_event(event_data)
event_global_plot
```
##### Global feature-level
```
feature_dict = {'path': 'outputs/feature_all.csv', 'rs': 42, 'nsamples': 32000, 'feature_names': model_features, 'plot_features': plot_feats, }
feat_data = feat_explain_all(f_hs, pos_dataset, sequence_id_feat, average_event, feature_dict, prun_indexes, model_features, time_feat)
feat_global_plot = plot_global_feat(feat_data, **feature_dict)
feat_global_plot
```
| github_jupyter |
### Intro.
**AI** is defined as a program that exhibits cognitive ability similar to that of a human being. Making computers think like humans and solve problems the way we do is one of the main tenets of artificial intelligence.
**ML**is fundamentally set apart from artificial intelligence, as it has the capability to evolve. Using various programming techniques, machine learning algorithms are able to process large amounts of data and extract useful information. In this way, they can improve upon their previous iterations by learning from the data they are provided.
With machine learning algorithms, AI was able to develop beyond just performing the tasks it was programmed to do. Before ML entered the mainstream, AI programs were only used to automate low-level tasks in business and enterprise settings.
This included tasks like intelligent automation or simple rule-based classification. This meant that AI algorithms were restricted to only the domain of what they were processed for. However, with machine learning, computers were able to move past doing what they were programmed and began evolving with each iteration.
### Types of Learning
Learning Problems:
1. **Supervised Learning** - Supervised learning describes a class of problem that involves using a model to learn a mapping between input examples and the target variable. e.g classification, regression.
2. **Unsupervised Learning** - Unsupervised learning describes a class of problems that involves using a model to describe or extract relationships in data. e.p clustering and DR.
3. **Reinforcement Learning** - Reinforcement learning describes a class of problems where an agent operates in an environment and must learn to operate using feedback. e.g Q-learning, temporal-difference learning, and deep reinforcement learning.
Hybrid Learning Problems:
4. **Semi-Supervised Learning** - Semi-supervised learning is supervised learning where the training data contains very few labeled examples and a large number of unlabeled examples. For example, classifying photographs requires a dataset of photographs that have already been labeled by human operators.Many problems from the fields of computer vision (image data), natural language processing (text data), and automatic speech recognition (audio data) fall into this category and cannot be easily addressed using standard supervised learning methods.
5. **Self-Supervised Learning** - an unsupervised learning problem that is framed as a supervised learning problem in order to apply supervised learning algorithms to solve it. e.g unsupervised image corpus used to train a supervised model.
6. **Multi-Instance Learning** -a supervised learning problem where individual examples are unlabeled; instead, bags or groups of samples are labeled. i.e collections labelled and not individual instances.
Statistical Inference:
7. **Inductive Learning** - using evidence to determine outcome.
8. **Deductive Inference** - using general rules to determine specific outcomes
9. **Transductive Learning** - used in the field of statistical learning theory to refer to predicting specific examples given specific examples from a domain.
<img src="https://3qeqpr26caki16dnhd19sv6by6v-wpengine.netdna-ssl.com/wp-content/uploads/2019/09/Relationship-between-Induction-Deduction-and-Transduction.png">
Learning Techniques:
10. **Transfer Learning** - a model is first trained on one task, then some or all of the model is used as the starting point for a related task.
11. **Ensemble Learning** - two or more modes are fit on the same data and the predictions from each model are combined.
### Types of Data
* Useless - data with high cardinality: no much relation with target vaariable.
* Nominal - e.g nationality, tribe.
* Binary - discrete data presented only in 2 categories.
* Ordinal - discrete integers that can be ranked or sorted..
* Count - discrete non-negative whole numbers.
* Time
* Interval
* Image
* Video
* Audio
* Text
### Python Basics
What can you do with Python?
* Simple Scripting?
* Web Development?
* Scraping?
* App Development?
### Tools
* Setting up Anaconda: MacBook/Windows/Linux (we’re gonna be using jupyter notebook/lab for less heavy tasks.
* Google Colab.
* Kaggle Notebooks.
* Azure, AWS (these come at s cost + basic knowledge of cloud computing is required).
### Pandas and Numpy
* arrays
* series
* dataframes
* operations
### Git Operations
* creating repo.
* cloning.
* updating code.
* collaboration.
| github_jupyter |
# Introduction to Applied Econometrics in Python
---
## Quick tip when using Jupyter notebooks
### Problem:
- Jupyter saves every object, class, and function in memory when you run each cell.
- First time users sometimes run a single cell over and over when debugging.
- This practice is problematic!
- If you are not careful, you could be running the same operation on a Dataframe multiple times!
- For example, imagine you had code in one cell to drop the first column of a three columned Dataframe `df = df.iloc[:,1:]`.
- If you ran this cell 3 times, you would end up deleting ALL the columns.
- These potentially unwanted manipulations of variables are called **side effects** in computer science.
### Solution:
- Unlike RStudio, Jupyter notebook does not have a window showing all the global variables in memory!
- When in doubt, restart your kernel and re-run all cells in order.
```
import pandas as pd
import numpy as np
import statsmodels.api as sm
import matplotlib.pyplot as plt
import seaborn as sns
from stargazer.stargazer import Stargazer
```
# 1. Importing Data
---
```
# Import STATA dta file (Penn World Table dataset)
# It is a large file (2.3mb) and might take a while to download
# Reference:
# Feenstra, Robert C., Robert Inklaar and Marcel P. Timmer (2015),
# "The Next Generation of the Penn World Table" American Economic Review, 105(10), 3150-3182,
# available for download at www.ggdc.net/pwt
pwt_data = pd.read_stata("https://www.rug.nl/ggdc/docs/pwt91.dta")
pwt_data
```
## 1.1. Check column names and data types
```
pwt_data.dtypes
```
### Comments:
- Common sense tells us that elements in `countrycode` and `country` should all be strings.
- However, upon importing any form of data (e.g. csv, xls, dta), Pandas often considers columns that are potentially strings as the object dtype.
- An object datatype in a Pandas dataframe holds *any* Python object, including objects that *mix* strings and non-strings.
- The best practice for columns that should *only* contain strings is to convert them to StringDtype (see https://pandas.pydata.org/pandas-docs/stable/user_guide/text.html#text-types for details)
- You do not need to convert ALL columns. Use your judgement and decide which columns will actually be used in the analysis.
## 1.2. Convert data types
```
# Convert country_name and wbcode to StringDtype
for column in ["countrycode", "country"]:
pwt_data[column] = pwt_data[column].astype("string")
# Check dtypes again
pwt_data.dtypes
```
## 1.3. Export cleaned data
### Problem:
- Often times you might want to seperate your data cleaning code with your data modelling code. Moreover, it is computationally expensive to have to run the data cleaning code everytime you terminate and reopen the Jupyter notebook.
### Solution:
- Export the dataframe into a file type that, as much as possible, *preserves the data types* when imported by Pandas in the future. This file type should also be quickly importable (Excel files are generally quite slow to be imported by Pandas).
- A common format used is the common seperated value (csv) file.
```
# Export democracy_data into the data directory
# Great! You've now got the Penn World Table dataset to test your Python data skills
pwt_data.to_csv("data/pwt.csv")
```
## Key Lesson
---
- Some loss of meta-information, in particular data type information, is expected when sharing data across different programming languages, file types, and machines.
- It is your responsibility to ensure that the data types in your dataframe are accurate after *every* data import.
### This exercise is important for three reasons:
1. Wrong data types lead to type errors when operating on your dataframe. (e.g. trying to add a column of strings with a column of integers)
2. Wrong data types can cause ***silent*** errors.
3. Wrong data types lead to ***inefficiencies***. Pandas has optimised operations specific to the dtype.
```
# Consider the following dataframe
fail_early_example = pd.DataFrame([["10.5", "5.0"],["9.4", "5.1"], ["11", "5.2"]])
fail_early_example.columns = ["profit", "cost"]
fail_early_example
# Create new column "revenue"
fail_early_example["revenue"] = fail_early_example["profit"] + fail_early_example["cost"]
fail_early_example
```
### Comments
- What went wrong?
- The two columns "profit" and "cost" that are inaccurately coded as strings. When we created a new column "revenue", the row with profit "10.5" and cost "5.0" will have revenue "10.55.0".
- Python worked exactly as it should and intepreted the + operator between the profit and cost columns as string concatenation!
- Defend against silent errors ***as early as possible***.
References:
https://blog.codinghorror.com/fail-early-fail-often/
# 2. Data Generation / Random Number Generation
---
```
# Let us define some variables to generate fake data
# Use global variables as sparingly as possible.
# We use a dictionary to hold our required variables.
# The official Python coding style guide (PEP 8) recommends
# the capitalisation of all global variable names.
RANDOM_DATA_PARAMS = {
"N": 3000,
"beta_0": 2,
"beta_1": 3,
"sigma": 5,
"transformation": lambda x: x + 5, # This is a lambda function (see https://realpython.com/python-lambda/)
"covariate_sd": 5}
SEED = 905
# The following is an example of a well-written function
# It contains a docstring.
def generate_reg_data(N, beta_0, beta_1, sigma, transformation, covariate_sd, seed=SEED):
"""Generates fake regression data with a list of three Pandas series.
The regression model consists of the outcome, treatment,
a covariate variable, and a constant term.
Arguments:
- N (integer): sample size
- beta_0 (integer): coefficient for the constant term
- beta_1 (integer): coefficient for the treatment variable
- sigma (integer): error term
- transformation (function): a function that takes in the
treatment column, applies a transformation, then returns the mean values
for the covariate columb
- covariate_sd (integer): covariate's standard deviation
Returns:
A list of three Pandas series. The 1st element is the treatment.
The 2nd element is the outcome. The 3rd element is the covariate.
"""
np.random.seed(SEED)
X = np.random.binomial(n=1, p=0.5, size=N)
y = np.random.normal(beta_0 + beta_1*X, sigma, size=N)
w = np.random.normal(transformation(X), sigma, size=N)
result = map(pd.Series, [y, X, w])
return result
# Create a Dataframe from the tuple of Series objects returned from generate_reg_data
# The double asterisks unpack the dictionary RANDOM_DATA_PARAMS. The values in the dictionary
# RANDOM_DATA_PARAMS are mapped to the keyword arguments in the functiom generate_reg_data
# according to the corresponding keys in RANDOM_DATA_PARAMS.
fake_data_list = generate_reg_data(**RANDOM_DATA_PARAMS)
fake_example_data = pd.concat(fake_data_list, axis=1, verify_integrity=True)
# Name the columns
fake_example_data.columns = ["profit", "treatment", "covariate"]
fake_example_data
```
## 2.1. Data Exploration
```
# Filter for untreated rows
fake_example_data_untreated = fake_example_data[fake_example_data["treatment"]==0]
fake_example_data_untreated
# Compute summary statistics
fake_example_data.describe()
# Compute covariance matrix
fake_example_data.cov()
# You can compute the covariance matrix for a subset of columns
fake_example_data[["treatment","profit"]].cov()
# Consider the following dataframe with missing values in every 4th row of the "treatment" column
# First let's create a new copy of the fake data:
fake_missing_data = fake_example_data # WRONG
# The above line only creates a reference (or "pointer" in CS-speak) to the original dataframe
# Any changes to fake_missing_data will modify the original fake_example_data dataframe.
fake_missing_data = fake_example_data.copy() # CORRECT
# Update every 4th row of the treatment column as a missing value
# NOTE! Some Pandas dataframe methods such as .iloc modify the dataframe inplace,
# whereas other methods such as .describe() returns a new object.
fake_missing_data.iloc[3::4, 0] = np.NaN
fake_missing_data
# Remove all rows with missing values
fake_missing_data = fake_missing_data.dropna()
fake_missing_data
# Sometimes it might be useful to reset the index
# Don't forget set drop=True or the old index will be kept as a new column
fake_missing_data = fake_missing_data.reset_index(drop=True)
fake_missing_data
```
### Beware!
- There are a lot of subtleties to dealing with missing data.
- From the programmer's perspective, there could be different abstractions to implement a "missing data" type (https://pandas.pydata.org/pandas-docs/stable/user_guide/missing_data.html)
- From the econometrician's perspective, removing rows with missing data could bias your results (e.g. the fact that some data is missing could be due to a selection effect correlated with potential outcomes).
# 3. Linear Regressions in Python statsmodels
---
- There are two APIs (Application Programming Interfaces) to running linear regressions in statsmodels.
- Statsmodels is a popular package for statistical modelling in Python.
- Both APIs are ***object orientated***:
1. You first instantiate some model class (e.g. OLS) by setting its required attributes.
2. You then call the model class's method that fits the data and compute the model's parameters. The fitted model's results are set inside the model instance.
3. Finally, you can access the results by calling the model class's method that returns a summary of the results.
- API 1. in step 1, you have to input **two seperate dataframes**: one with the endogenous variable and the other with the exogenous variables.
- API 2. in step 1, you have to input a R-styled **formula** that specifies the regression model fitted on some data in a dataframe.
References:
- https://www.statsmodels.org/stable/regression.html
- https://www.statsmodels.org/dev/examples/notebooks/generated/formulas.html
### My hot take:
- The two APIs generate the same results.
- Nevertheless, I prefer API 2.
- To see why, let's compare the two APIs in the examples below.
### 3.1. Regress profit on treatment
```
# API 1.
mod = sm.OLS(endog=fake_example_data["profit"], exog = sm.add_constant(fake_example_data["treatment"]))
res = mod.fit()
res.summary()
# API 2. Using R-styled formulas
mod = sm.formula.ols(formula='profit ~ treatment', data=fake_example_data)
res = mod.fit()
res.summary()
```
### Side note:
```
# To run a regression without a constant term using API 2.
# Include a -1 term to the formula
mod = sm.formula.ols(formula='profit ~ treatment -1', data=fake_example_data)
res = mod.fit()
res.summary()
```
### 3.2. Regress profit on treatment and covariate
```
# API 1.
mod = sm.OLS(endog=fake_example_data["profit"], exog = sm.add_constant(fake_example_data[["treatment", "covariate"]]))
res = mod.fit()
res.summary()
# API 2. Using R-styled formulas
mod = sm.formula.ols(formula='profit ~ treatment + covariate', data=fake_example_data)
res = mod.fit()
res.summary()
```
## 3.3. Interactions
```
# API 1. Part 1.
# Create a column for an interaction between treatment and the covariate
fake_interaction_data = fake_example_data.copy()
fake_interaction_data["treatment:covariate"] = fake_interaction_data["treatment"] * fake_interaction_data["covariate"]
fake_interaction_data
# API 1. Part 2.
mod = sm.OLS(endog=fake_interaction_data["profit"],
exog = sm.add_constant(fake_interaction_data[["treatment", "covariate", "treatment:covariate"]]))
res = mod.fit()
res.summary()
# API 2. Using R-styled formulas
# You can simply specify the interaction in the formula.
# No need to explicitly modify / slice the original fake_example_data dataframe!
# See https://patsy.readthedocs.io/en/latest/categorical-coding.html
mod = sm.formula.ols(formula='profit ~ treatment*covariate', data=fake_example_data)
res = mod.fit()
res.summary()
```
## 3.4. Factor variables
```
# Create column of factors
factors = np.array(["good","bad","ugly"])
factors_data = np.repeat(factors,
RANDOM_DATA_PARAMS["N"] / len(factors), axis=0)
factors_data = pd.Series(factors_data)
factors_data
# Append column of factors to dataframe
fake_example_data["factor"] = factors_data
fake_example_data
# API 1. Part 1.
# To use API 1, you have to create indicator variable (dummy) columns from the factor variable column.
# Creating and working with dummy variables in Pandas dataframes is messy.
dummies_data = pd.get_dummies(fake_example_data["factor"])
dummies_data
fake_factors_data = pd.concat([fake_example_data, dummies_data], axis=1)
fake_factors_data = fake_factors_data.drop("factor", axis=1)
fake_factors_data
# API 1. Part 2.
mod = sm.OLS(endog=fake_example_data["profit"], exog = fake_factors_data.iloc[:,1:])
res = mod.fit()
res.summary()
# API 2. Using R-styled formulas
# You can simply code the categorical (or factor) variable in the formula.
# No need to explicitly modify / slice the original fake_factors_data dataframe!
# See https://patsy.readthedocs.io/en/latest/categorical-coding.html
mod = sm.formula.ols(formula='profit ~ treatment + covariate + C(factor) - 1', data=fake_example_data)
res = mod.fit()
res.summary()
```
## 3.5. Saving text-like files
```
# Save regression results table in LaTeX into the media directory
with open("media/our_fancy_regression_table.tex", "w") as file:
file.write(res.summary().as_latex())
```
# 4. Fancy Standard Errors in Python statsmodels
---
```
# Newey-West (1987) HAC standard errors
res_HAC = res.get_robustcov_results(cov_type="HAC", maxlags=0)
res_HAC.summary()
# White’s (1980) heteroskedasticity robust standard errors
res_EHW = res.get_robustcov_results(cov_type="HC0")
res_EHW.summary()
# There is a Python implementation of R stargazer!
# However, stargazer takes in trained OLS models as its arguments
mod_HAC = sm.formula.ols(formula='profit ~ treatment + covariate + C(factor) - 1', data=fake_example_data)
res_HAC = mod_HAC.fit(cov_type="HAC", cov_kwds={"maxlags": 0})
mod_EHW = sm.formula.ols(formula='profit ~ treatment + covariate + C(factor) - 1', data=fake_example_data)
res_EHW = mod_EHW.fit(cov_type="HC0")
stargazer = Stargazer([res, res_HAC, res_EHW])
stargazer.custom_columns(["default","HAC","robust"], [1,1,1])
stargazer
# Save regression results table in LaTeX into the media directory
with open("media/our_stargazer_regression_table.tex", "w") as file:
file.write(stargazer.render_latex())
```
# 5. Plotting in Python Pandas and seaborn
---
## Comments:
- Pandas Dataframes have methods that wrap around the matplotlib API to generate matplotlib plots.
- Seaborn is a data visualization library based on matplotlib. It provides a high-level API for drawing more inspired graphs than matplotlib.
References:
- https://pandas.pydata.org/pandas-docs/stable/user_guide/visualization.html
- https://seaborn.pydata.org/examples/index.html
```
# Close any previously generated plots saved in memory
plt.close('all')
```
## 5.1. Pandas in-built plotting tools
```
# Plot regression residuals
res_EHW.resid.plot()
# Scatterplot of treatment and profit
fake_example_data.plot.scatter(x="treatment", y="profit")
```
## 5.2. Seaborn
```
# Set a Seaborn theme (see https://seaborn.pydata.org/tutorial/aesthetics.html)
# Set size of figures
sns.set_style("darkgrid")
sns.set(rc={'figure.figsize':(12,9)})
# Scatterplot with jitter where treatment is coded as a categorical variable
jitter_plot = sns.stripplot(x="treatment", y="profit", data=fake_example_data, jitter=0.1, palette="Set2")
jitter_plot
# Colour code observations by 5 unit ranges of the covariate
min_covariate = min(fake_example_data['covariate'])
max_covariate = max(fake_example_data['covariate'])
fake_example_data['covariate_bin'] = pd.cut(fake_example_data['covariate'], np.arange(min_covariate, max_covariate, step=5))
jitter_plot_with_covariate = sns.stripplot(x="treatment", y="profit", hue="covariate_bin",
data=fake_example_data, jitter=0.1, palette="Set2")
jitter_plot_with_covariate
```
## Comments:
- Can we display all the Seaborn plots together?
- Yes!
- The seaborn methods above return matplotlib ***axes*** objects. Each axes corresponds to a single graph.
- You can group individual axes together into a ***figure***.
- These axes associated with a figure can be organised in different ways (e.g. placed side-by-side or even imposed on top of each other).
References:
- https://www.math.purdue.edu/~bradfor3/ProgrammingFundamentals/matplotlib/#canvases-figures-and-axes
```
# Regression plot with lines stratified by covariate bins
# This plot is actually a figure!
# It is two axes, a regression plot and a scatterplot, imposed on top of each other.
reg_plot = sns.lmplot(x="treatment", y="profit", hue="covariate_bin",
data=fake_example_data, x_jitter=0.1, palette="Set2", height=10)
# Let's seperate out the regressions in the above graph
reg_subplots = sns.lmplot(x="treatment", y="profit", hue="covariate_bin", col="covariate_bin",
data=fake_example_data, x_jitter=0.1, col_wrap=2, palette="Set2")
```
## 5.3. Saving seaborn plots
```
# Save regression subplots as pdf into the media directory
reg_subplots.savefig("media/fancy_reg_subplots.pdf")
```
# 6. Python Data Science Resources
---
- https://pandas.pydata.org/pandas-docs/stable/getting_started/comparison/comparison_with_sql.html
- https://seaborn.pydata.org/
- https://datascience.quantecon.org/
| github_jupyter |
# Introduction to Jupyter notebooks
> This tutorial introduces Jupyter notebooks.
- toc: false
- badges: true
- comments: true
- categories: [jupyter]
#### Welcome to the first tutorial in this DEA for Geospatial Analysts course. In this section we introduce the Jupyter notebook environment. Jupyter offers an interactive platform to run Python code in your browser without the need of installing or configuring anything on you computer.
#### This is an interactive environment where you can combine text with with Python code that you can modify and execute. The gray cell below contains Python code that you can run. Place the cursor on the |► Run| button in the menu at the top of the notebook to execute the code inside.
> Tip: _Use the shortcut `Shift-Enter` for running the active cell (where your cursor is). It is a convenient 'trick' that saves you from having to click on the `Run` icon each time._
```
print("Welcome to the DEA Sandbox!")
```
#### You'll see that, once the code has executed, any output generated by your code will show up just below the cell
```
print("3,2,1")
# Use the hash symbol to comment lines of your code. This does not get executed
print("this is fun!")
```
#### The code that you execute using the DEA Sandbox [Binder, Colab] does not run on your local computer. It runs in a remote server and you just receive the output of the code that gets presented in this browser's window.
> Tip: The last line of code in a cell is evaluated and printed by default, so there is no need to type `print()` to show the value of a variable in the last line
```
# Sending a complicated operation to the server
result = 999 + 1
# This operation goes to the server and we just display its output in our screens once is completed
result
```
#### The `[ ]:` symbol to the left of each Code cell describes the state of the cell:
* `[ ]:` means that the cell has not been run yet.
* `[*]:` means that the cell is currently running.
* `[1]:` means that the cell has finished running and was the first cell run.
#### Sometimes the code that you run in a cell takes a while to compute because it loads a large dataset or performs complex computations. You'll notice the number that appears next to the cell. Before the cell is run you'll see the symbol `[ ]` meaning that that cell has not been executed yet. While a cell is running it shows the `[*]` symbol and once it has completed running you'll see a number representing the number of cells being run, for example `[4]`. This allows you to keep track of the cells that have been run and their relative order.
> **Note:** To check whether a cell is currently executing in a Jupyter notebook, inspect the small circle in the top-right of the window.
The circle will turn black ("Kernel busy") when the cell is running, and return to white ("Kernel idle") when the process is complete.
```
# This is how a library gets imported in Python but don't worry about this for the moment:
import time
# Very complicated and long operation computed on the server -- not really we are just waiting or sleeping for 5 seconds :-)
time.sleep(5)
print("Done!")
```
#### Jupyter notebooks are an easy way to write Python code, although they work a little bit different to how traditional programs are written and executed. For example, consider this simple Python program:
```python
a = 1
print(a)
a = a + 1
print(a)
```
#### We can break down this program and execute each line using separate cells:
```
a = 1
a
a = a + 1
a
```
#### If you run again the first `a` cell you'll see that it returns `2`, the updated value. This is called global state and means that once a variable is declared it is accessible anywhere in the notebook, even in cells above where it has been declared. This is different to traditional programs which execute sequentially line by line from the top to the bottom and can be confusing in the beginning.
#### Jupyter notebooks are designed for interactivity and testing ideas fast. You can modify all the code in the cells and run them as many times as you want in any order (you can jump back and forth to update variables or re-run analysis)
```
message = "My name is ******"
message
```
#### You can also add new cells to insert new code at any point in a notebook. Click on the `+` icon in the top menu to add a new cell below the current one.
> Tip: _There is a shortcut to add new cells by pressing `Esc + a` to add a cell just above and `Esc + b` to add a cell below._
```
a = 365*24
# Add a new cell just below this one to print the value of variable `a`
```
#### The rest of the units in this course use Jupyter notebooks to introduce and get some practice with the different libraries and functionality required to understand the DEA Python interface. After a new concept is introduced there will be cells where you need to answer questions or solve small problems. To check your answers we are going to use an interactive way of
#### Exercise 0.1: The following cell contains some code. To complete this exercise you need to update the contents of the operation `2 + 2` for variable `a`. Delete the current value `0` and write the new value, then execute the cell and see the response.
```
from check_answer import check_answer
a = 0
check_answer("0.1", a)
```
#### Saving and exporting your work. Jupyter Notebooks are automatically saved every few minutes. To actively save the notebook, navigate to "File" in the menu bar, then select "Save Notebook" or click the 💾 "save" icon on the left of the graphical menu.
#### One last thing, you can use code cells to run commands as if you were using a terminal using the `!` character before the command. This is a Linux system and to list the contents of the current directory you'll need to use `ls`
```
!ls
```
#### You are now ready to start learning Numpy!
| github_jupyter |
# Decorator
---
```
# basic function signature
def hello1():
print("Hello")
# what is a function ?
hello1
# assigning a function another name
h = hello1
# calling function with new name
h()
```
*We can define function inside a function*
```
# function within a function
def hello(name):
def world():
return "Hello" + name
print(world())
hello('World')
```
*What happens here is, when we call __hello__ function it defines __world__ function and then execute __print__ statement, now __world__ is being called inside __print__ which is when the function body for __world__ executes.*
**Closures**
```
def hello2(name):
def world():
print("Hello2 " + name)
return world
```
*__Closures__ are functions which returns another function.*
```
dir(hello2)
hello2.__name__
h2 = hello2
h2.__name__
hello2
hello2('World')
# h2 here is returned function ie world
h2i = hello2('World')
h2i
h2i()
# we can also call inner function as well
hello2('World')()
```
**Function can be passed to other function as well**
```
def info(func, *args):
print("Function Name: ", func.__name__)
print("Function docstring: ", func.__doc__)
print("Function Code: ", func.__code__)
# func(*args) ==> hello2(*args) ==> hello2('World')
func.__call__(*args)
```
*The **func.\_\_name\_\_** attribute has name of the passed function __func__*
*The **func.\_\_doc\_\_** attribute has docstring of the passed function __func__*
*The **func.\_\_code\_\_** attribute has code/(function body) of the passed function __func__*
```
info(hello2, 'World')
def test():
"""
Test doc string
"""
print("Test")
info(test)
```
**Modifying function at runtime**
```
def hello(name):
return "Hello " + name
def changer(func):
def inner(*args, **kwargs):
print(args)
print(kwargs)
vargs = list(args)
vargs[0] = vargs[0] + 'ly'
return func.__call__(*tuple(vargs), **kwargs)
return inner
# changer accepts a function as argument ie hello and updates its arguments and returns a new function
# that new function is assigned to same name "hello" basically overwriting the old function signature
hello = changer(hello)
hello('World')
hello('bird')
hello(name='world2')
hello = changer(hello)
hello('World')
```
**Decorator Syntax**
```
import time
time.time()
def timer(func):
def wrapper(*args, **kwargs):
now = time.time()
print(args, kwargs)
print(func.__name__ + " call started")
func.__call__(*args, **kwargs)
print(func.__name__ + " call ended " + str(time.time() - now ))
return wrapper
def fib(a, b, num=6):
result = []
for i in range(0, num):
a, b = b, a + b
result.append(b)
print(result)
fib(0, 1, 100)
```
*Any name that comes after __@__ symbol is a function, what it does is passes function (the one that it precedes see, code below eg. fib2 in this case) to decorator function ( the one after __@__ ) and returned function is assigned to same name.*
```
@timer
def fib2(a, b, num=6):
result = []
for i in range(0, num):
a, b = b, a + b
result.append(b)
print(result)
```
*Above statement is same as below, only shortcut*
```python
def fib2(a, b, num=6):
result = []
for i in range(0, num):
a, b = b, a + b
result.append(b)
print(result)
fib2 = timer(fib2)
```
```
fib2(0, 1, 100)
fib2(0, 1, num=100)
```
_Notes on \*args and \*\*kwargs_
```
# *args holds all positional arguments passed to the function in tuple
# **kwargs holds all named arguments passed to the function in dictionary
def help(*args, **kwargs):
# remember its args and kwargs that holds tuple and dict
# * and ** are just syntax for what it does
print(args, kwargs)
# running above function shows what args and kwargs holds
help(1, 2, 3, name='test', age=34)
help((1, 2, 3), {'abc': 98})
help(*(1, 2, 3), **{'abc': 98})
# using * on a tuple (** on a dictionary ) unwinds it
print(1, 2)
# can't use like this and can't pass to print function either
print(**{'name': 'hello'})
# as it calls print like this => print(name='hello')
**{'name': 'hello'}
def test(name='empty'):
print(name)
test('positional')
test(name='keyword')
test(*('positional tuple',))
test(**{'name': 'keyword dict'})
```
[learnpython](http://learnpython.org)
[dive into python](http://diveintopython3.net)
| github_jupyter |
# Name
Data preparation using SparkSQL on YARN with Cloud Dataproc
# Label
Cloud Dataproc, GCP, Cloud Storage, YARN, SparkSQL, Kubeflow, pipelines, components
# Summary
A Kubeflow Pipeline component to prepare data by submitting a SparkSql job on YARN to Cloud Dataproc.
# Details
## Intended use
Use the component to run an Apache SparkSql job as one preprocessing step in a Kubeflow Pipeline.
## Runtime arguments
Argument| Description | Optional | Data type| Accepted values| Default |
:--- | :---------- | :--- | :------- | :------ | :------
project_id | The ID of the Google Cloud Platform (GCP) project that the cluster belongs to. | No| GCPProjectID | | |
region | The Cloud Dataproc region to handle the request. | No | GCPRegion|
cluster_name | The name of the cluster to run the job. | No | String| | |
queries | The queries to execute the SparkSQL job. Specify multiple queries in one string by separating them with semicolons. You do not need to terminate queries with semicolons. | Yes | List | | None |
query_file_uri | The HCFS URI of the script that contains the SparkSQL queries.| Yes | GCSPath | | None |
script_variables | Mapping of the query’s variable names to their values (equivalent to the SparkSQL command: SET name="value";).| Yes| Dict | | None |
sparksql_job | The payload of a [SparkSqlJob](https://cloud.google.com/dataproc/docs/reference/rest/v1/SparkSqlJob). | Yes | Dict | | None |
job | The payload of a [Dataproc job](https://cloud.google.com/dataproc/docs/reference/rest/v1/projects.regions.jobs). | Yes | Dict | | None |
wait_interval | The number of seconds to pause between polling the operation. | Yes |Integer | | 30 |
## Output
Name | Description | Type
:--- | :---------- | :---
job_id | The ID of the created job. | String
## Cautions & requirements
To use the component, you must:
* Set up a GCP project by following this [guide](https://cloud.google.com/dataproc/docs/guides/setup-project).
* [Create a new cluster](https://cloud.google.com/dataproc/docs/guides/create-cluster).
* The component can authenticate to GCP. Refer to [Authenticating Pipelines to GCP](https://www.kubeflow.org/docs/gke/authentication-pipelines/) for details.
* Grant the Kubeflow user service account the role `roles/dataproc.editor` on the project.
## Detailed Description
This component creates a Pig job from [Dataproc submit job REST API](https://cloud.google.com/dataproc/docs/reference/rest/v1/projects.regions.jobs/submit).
Follow these steps to use the component in a pipeline:
1. Install the Kubeflow Pipeline SDK:
```
%%capture --no-stderr
!pip3 install kfp --upgrade
```
2. Load the component using KFP SDK
```
import kfp.components as comp
dataproc_submit_sparksql_job_op = comp.load_component_from_url(
'https://raw.githubusercontent.com/kubeflow/pipelines/1.1.2-rc.1/components/gcp/dataproc/submit_sparksql_job/component.yaml')
help(dataproc_submit_sparksql_job_op)
```
### Sample
Note: The following sample code works in an IPython notebook or directly in Python code. See the sample code below to learn how to execute the template.
#### Setup a Dataproc cluster
[Create a new Dataproc cluster](https://cloud.google.com/dataproc/docs/guides/create-cluster) (or reuse an existing one) before running the sample code.
#### Prepare a SparkSQL job
Either put your SparkSQL queries in the `queires` list, or upload your SparkSQL queries into a file to a Cloud Storage bucket and then enter the Cloud Storage bucket’s path in `query_file_uri`. In this sample, we will use a hard coded query in the `queries` list to select data from a public CSV file from Cloud Storage.
For more details about Spark SQL, see [Spark SQL, DataFrames and Datasets Guide](https://spark.apache.org/docs/latest/sql-programming-guide.html)
#### Set sample parameters
```
PROJECT_ID = '<Please put your project ID here>'
CLUSTER_NAME = '<Please put your existing cluster name here>'
REGION = 'us-central1'
QUERY = '''
DROP TABLE IF EXISTS natality_csv;
CREATE EXTERNAL TABLE natality_csv (
source_year BIGINT, year BIGINT, month BIGINT, day BIGINT, wday BIGINT,
state STRING, is_male BOOLEAN, child_race BIGINT, weight_pounds FLOAT,
plurality BIGINT, apgar_1min BIGINT, apgar_5min BIGINT,
mother_residence_state STRING, mother_race BIGINT, mother_age BIGINT,
gestation_weeks BIGINT, lmp STRING, mother_married BOOLEAN,
mother_birth_state STRING, cigarette_use BOOLEAN, cigarettes_per_day BIGINT,
alcohol_use BOOLEAN, drinks_per_week BIGINT, weight_gain_pounds BIGINT,
born_alive_alive BIGINT, born_alive_dead BIGINT, born_dead BIGINT,
ever_born BIGINT, father_race BIGINT, father_age BIGINT,
record_weight BIGINT
)
ROW FORMAT DELIMITED FIELDS TERMINATED BY ','
LOCATION 'gs://public-datasets/natality/csv';
SELECT * FROM natality_csv LIMIT 10;'''
EXPERIMENT_NAME = 'Dataproc - Submit SparkSQL Job'
```
#### Example pipeline that uses the component
```
import kfp.dsl as dsl
import json
@dsl.pipeline(
name='Dataproc submit SparkSQL job pipeline',
description='Dataproc submit SparkSQL job pipeline'
)
def dataproc_submit_sparksql_job_pipeline(
project_id = PROJECT_ID,
region = REGION,
cluster_name = CLUSTER_NAME,
queries = json.dumps([QUERY]),
query_file_uri = '',
script_variables = '',
sparksql_job='',
job='',
wait_interval='30'
):
dataproc_submit_sparksql_job_op(
project_id=project_id,
region=region,
cluster_name=cluster_name,
queries=queries,
query_file_uri=query_file_uri,
script_variables=script_variables,
sparksql_job=sparksql_job,
job=job,
wait_interval=wait_interval)
```
#### Compile the pipeline
```
pipeline_func = dataproc_submit_sparksql_job_pipeline
pipeline_filename = pipeline_func.__name__ + '.zip'
import kfp.compiler as compiler
compiler.Compiler().compile(pipeline_func, pipeline_filename)
```
#### Submit the pipeline for execution
```
#Specify pipeline argument values
arguments = {}
#Get or create an experiment and submit a pipeline run
import kfp
client = kfp.Client()
experiment = client.create_experiment(EXPERIMENT_NAME)
#Submit a pipeline run
run_name = pipeline_func.__name__ + ' run'
run_result = client.run_pipeline(experiment.id, run_name, pipeline_filename, arguments)
```
## References
* [Spark SQL, DataFrames and Datasets Guide](https://spark.apache.org/docs/latest/sql-programming-guide.html)
* [SparkSqlJob](https://cloud.google.com/dataproc/docs/reference/rest/v1/SparkSqlJob)
* [Cloud Dataproc job](https://cloud.google.com/dataproc/docs/reference/rest/v1/projects.regions.jobs)
## License
By deploying or using this software you agree to comply with the [AI Hub Terms of Service](https://aihub.cloud.google.com/u/0/aihub-tos) and the [Google APIs Terms of Service](https://developers.google.com/terms/). To the extent of a direct conflict of terms, the AI Hub Terms of Service will control.
| github_jupyter |
Lambda School Data Science
*Unit 2, Sprint 1, Module 4*
---
# Logistic Regression
## Assignment 🌯
You'll use a [**dataset of 400+ burrito reviews**](https://srcole.github.io/100burritos/). How accurately can you predict whether a burrito is rated 'Great'?
> We have developed a 10-dimensional system for rating the burritos in San Diego. ... Generate models for what makes a burrito great and investigate correlations in its dimensions.
- [ ] Do train/validate/test split. Train on reviews from 2016 & earlier. Validate on 2017. Test on 2018 & later.
- [ ] Begin with baselines for classification.
- [ ] Use scikit-learn for logistic regression.
- [ ] Get your model's validation accuracy. (Multiple times if you try multiple iterations.)
- [ ] Get your model's test accuracy. (One time, at the end.)
- [ ] Commit your notebook to your fork of the GitHub repo.
- [ ] Watch Aaron's [video #1](https://www.youtube.com/watch?v=pREaWFli-5I) (12 minutes) & [video #2](https://www.youtube.com/watch?v=bDQgVt4hFgY) (9 minutes) to learn about the mathematics of Logistic Regression.
## Stretch Goals
- [ ] Add your own stretch goal(s) !
- [ ] Make exploratory visualizations.
- [ ] Do one-hot encoding.
- [ ] Do [feature scaling](https://scikit-learn.org/stable/modules/preprocessing.html).
- [ ] Get and plot your coefficients.
- [ ] Try [scikit-learn pipelines](https://scikit-learn.org/stable/modules/compose.html).
```
%%capture
import sys
# If you're on Colab:
if 'google.colab' in sys.modules:
DATA_PATH = 'https://raw.githubusercontent.com/LambdaSchool/DS-Unit-2-Linear-Models/master/data/'
!pip install category_encoders==2.*
# If you're working locally:
else:
DATA_PATH = '../data/'
# Load data downloaded from https://srcole.github.io/100burritos/
import pandas as pd
df = pd.read_csv(DATA_PATH+'burritos/Burrito - 10D.csv')
# Derive binary classification target:
# We define a 'Great' burrito as having an
# overall rating of 4 or higher, on a 5 point scale.
# Drop unrated burritos.
df = df.dropna(subset=['overall'])
df['Great'] = df['overall'] >= 4
# Clean/combine the Burrito categories
df['Burrito'] = df['Burrito'].str.lower()
california = df['Burrito'].str.contains('california')
asada = df['Burrito'].str.contains('asada')
surf = df['Burrito'].str.contains('surf')
carnitas = df['Burrito'].str.contains('carnitas')
df.loc[california, 'Burrito'] = 'California'
df.loc[asada, 'Burrito'] = 'Asada'
df.loc[surf, 'Burrito'] = 'Surf & Turf'
df.loc[carnitas, 'Burrito'] = 'Carnitas'
df.loc[~california & ~asada & ~surf & ~carnitas, 'Burrito'] = 'Other'
# Drop some high cardinality categoricals
df = df.drop(columns=['Notes', 'Location', 'Reviewer', 'Address', 'URL', 'Neighborhood'])
# Drop some columns to prevent "leakage"
df = df.drop(columns=['Rec', 'overall'])
df.info()
df.describe().T
df.describe(include = object).T
df.drop(['Mass (g)', 'Density (g/mL)', 'Queso', 'Date', 'NonSD', 'Lobster', 'Zucchini', 'Carrots', 'Salsa.1'], axis = 1, inplace = True)
df.head()
df['Great'] = df['Great'].astype(int)
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
sns.heatmap(df[df.describe().T.index].corr())
from sklearn.model_selection import train_test_split
train, valtest = train_test_split(df, test_size = .2)
val, test = train_test_split(valtest, test_size = .5)
train.shape, val.shape, test.shape
target = 'Great'
y_train = train[target]
baseline = y_train.mean()
y_train.value_counts(normalize = True)
from sklearn.linear_model import LogisticRegression
```
| github_jupyter |
## Autograded Notebook (Canvas & CodeGrade)
This notebook will be automatically graded. It is designed to test your answers and award points for the correct answers. Following the instructions for each Task carefully.
### Instructions
* **Download this notebook** as you would any other ipynb file
* **Upload** to Google Colab or work locally (if you have that set-up)
* **Delete `raise NotImplementedError()`**
* Write your code in the `# YOUR CODE HERE` space
* **Execute** the Test cells that contain `assert` statements - these help you check your work (others contain hidden tests that will be checked when you submit through Canvas)
* **Save** your notebook when you are finished
* **Download** as a `ipynb` file (if working in Colab)
* **Upload** your complete notebook to Canvas (there will be additional instructions in Slack and/or Canvas)
# Lambda School Data Science - Unit 1 Sprint 3
## Sprint Challenge - Linear Algebra
### Total notebook points: 10
## Welcome to the final Sprint Challenge of Unit 1!
In this challenge, we're going to explore two different datasets where you can demonstrate your skills with fitting linear regression models and practicing some of the linear algebra concepts you learned.
**Make sure to follow the instructions in each task carefully!** The autograded tests are very specific in that they are designed to test on the exact instructions.
Good luck!
## Part A: Linear Regression
### Use the following information to complete Tasks 1 - 11
### Dataset description
The data you will work on for this Sprint Challenge is from the World Happiness Report. The report compiles data from a survey of hundreds of countries and looks at factors such as economic production, social support, life expectancy, freedom, absence of corruption, and generosity to determine a happiness "score".
In this Sprint Challenge, we're only going to look at the report for years 2018 and 2019. We're going to see how much the happiness "score" depends on some of the factors listed above.
For more information about the data, you can look here: [Kaggle: World Happiness Report](https://www.kaggle.com/unsdsn/world-happiness)
### Task 1 - Load the data
* import both `pandas` and `numpy`
* use the URL provided to read in your DataFrame
* load the CSV file as a DataFrame with the name `happy` and **set the index column as** `Overall_rank`.
* the shape of your DataFrame should be `(312, 8)`
```
# Task 1
# URL provided
url = "https://raw.githubusercontent.com/LambdaSchool/data-science-practice-datasets/main/unit_1/Happy/happiness_years18_19.csv"
# YOUR CODE HERE
import pandas as pd
import numpy as np
happy = pd.read_csv(url, index_col='Overall_rank')
# Print out the DataFrame
happy.head()
```
**Task 1 - Test**
```
# Task 1 - Test
assert isinstance(happy, pd.DataFrame), 'Have you created a DataFrame named `happy`?'
assert happy.index.name == 'Overall_rank', "Your index should be 'Overall_rank'."
assert len(happy) == 312
```
**Task 2** - Explore the data and find NaNs
Now you want to take a look at the dataset, determine the variable types of the columns, identify missing values, and generally better understand your data.
**Your tasks**
* Use describe() and info() to learn about any missing values, the data types, and descriptive statistics for each numeric value
* Sum the null values and assign that number to the variable `num_null`; the variable type should be a `numpy.int64` integer.
**Hint:** If you use `np.isnull()` it will return the number of null values in each column. You want the total number of null values in the entire DataFrame; one way to do this is to apply the `.sum()` method twice: `.sum().sum()`
```
# Task 2
# YOUR CODE HERE
happy.describe()
happy.info()
num_null = happy.isnull().sum().sum()
# Print out your integer result
print("The total number of null values is:", num_null)
```
**Task 2 Test**
```
# Task 2 - Test
import numpy as np
assert isinstance(num_null, np.int64), 'The sum of the NaN values should be an integer.'
```
**Task 3** - Drop a column
As you noticed in the previous task, the column `Perceptions_corruption` has a lot of missing values. Let's determine how many are missing and then drop the column. Note: dropping a column isn't always the best choice when faced with missing values but we're choosing that option here, partly for practice.
* Calculate the percentage of NaN values in `Perceptions_corruption` and assign the result to the variable `corruption_nan`; the value should be a **float** between `1.0` and `100.0`.
* Drop the `Perceptions_corruption` column from `happy` but keep the DataFrame name the same; use the parameter `inplace=True`. You will also want to specify the axis on which to operate.
```
# Task 3
# YOUR CODE HERE
corruption_nan = ((happy['Perceptions_corruption'].isnull().sum())/(len(happy['Perceptions_corruption'])))*100
happy.drop(['Perceptions_corruption'], axis=1, inplace=True)
# Print the percentage of NaN values
print(corruption_nan)
# Print happy to verify the column was dropped
happy.head()
```
***Task 3 Test**
```
# Task 3 - Test
assert isinstance(corruption_nan, np.float), 'The percentage of NaN values should be a float.'
assert corruption_nan >= 1, 'Make sure you calculated the percentage and not the decimal fraction.'
```
**Task 4** - Visualize the dataset
Next, we'll create a visualization for this dataset. We know from the introduction that we're trying to predict the happiness score from the other factors. Before we do let, let's visualize the dataset using a seaborn `pairplot` to look at all of the columns plotted as "pairs".
**Your tasks**
* Use the seaborn library `sns.pairplot()` function to create your visualization (use the starter code provided)
This task will not be autograded - but it is part of completing the challenge.
```
# Task 4
# (NOT autograded but fill in your code!)
# Import seaborn
import seaborn as sns
# Use sns.pairplot(data) where data is the name of your DataFrame
# sns.pairplot()
# YOUR CODE HERE
sns.pairplot(happy)
```
**Task 5** - Identify the dependent and independent variables
Before we fit a linear regression to the variables in this data set, we need to determine the dependent variable (the target or y variable) and independent variable (the feature or x variable). For this dataset, we have one dependent variable and a few choices for the independent variable(s). Using the information about the data set and what you know from previous tasks, complete the following:
* Assign the dependent variable to `y_var`
* Choose **one** independent variable and assign it to `x_var`
```
# Task 5
# YOUR CODE HERE
y_var = happy['Score']
x_var = happy['GDP_per_capita']
```
**Task 5 Test**
```
# Task 5 - Test
# Hidden tests - you will see the results when you submit to Canvas
```
**Task 6** - Fit a line using seaborn
Before we fit the linear regression model, we'll check how well a line fits. Because you have some choices for which independent variable to select, we're going to complete the rest of our analysis using `GDP per capita` as the independent variable. We're using `Score` as the dependent (target) variable.
The seaborn `lmplot()` documentation can be found [here](https://seaborn.pydata.org/generated/seaborn.lmplot.html). You can also use `regplot()` and the documentation is [here](https://seaborn.pydata.org/generated/seaborn.regplot.html)
This task will not be autograded - but it is part of completing the challenge!
**Your tasks:**
* Create a scatter plot using seaborn with `GDP per capita` and `Score`
* Use `sns.lmplot()` or `sns.regplot()` and specify a confidence interval of 0.95
* Answer the questions about your plot (not autograded).
```
# Task 6
# YOUR CODE HERE
sns.regplot(x="GDP_per_capita", y="Score", data=happy);
```
**Task 6** - Short answer
1. Does it make sense to fit a linear model to these two variables? In otherwords, are there any problems with this data like extreme outliers, non-linearity, etc.
2. Over what range of your independent variable does the linear model not fit the data well? Over what range does a line fit the data well?
---
1. Yes, it makes sense to fit the linear model. There are relatively few outliers, though there are some at the very highest and lowest levels of GDP_per_capita.
2. Right at 0 for the independent variable isn't the greatest fit, and also above 2 for GDP_per_capita the data isn't very close to the line of best fit. Between 0.1 and 1.75 the independent variable fits the linear model well.
**Task 7** - Fit a linear regression model
Now it's time to fit the linear regression model! We have two variables (`GDP_per_capita` and `Score`) that we are going to use in our model.
**Your tasks:**
* Use the provided import for the `statsmodels.formula.api` library `ols` method
* Fit a **single variable linear regression model** and assign the model to the variable `model_1`
* Print out the model summary and assign the value of R-squared for this model to `r_square_model_1`. Your value should be defined to three decimal places (example: `r_square_model_1 = 0.123`)
* Answer the questions about your resulting model parameters (these short answer questions will not be autograded).
**NOTE:** - For this task to be correctly autograded, you need to input the model parameters as specified in the code cell below. Part of this Sprint Challenge is correctly implementing the instructions in each task.
```
# Task 7
# Import the OLS model from statsmodels
from statsmodels.formula.api import ols
# YOUR CODE HERE
model_1 = ols('Score~GDP_per_capita', data=happy).fit()
# Print the model summary
print(model_1.summary())
r_square_model_1 = 0.637
```
**Task 7 Test**
```
# Task 7 - Test
# Hidden tests - you will see the results when you submit to Canvas
```
**Task 8** - Interpret your model
Using the model summary you printed out above, answer the following questions.
* Assign the slope of `GDP_per_capita` to the variable `slope_model_1`; define it to two decimal places (example: 1.23). This variable should be a float.
* Assign the p-value for this model parameter to `pval_model_1`; this variable could be either an integer or a float.
* Assign the 95% confidence interval to the variables `ci_low` (lower value) and `ci_upper` (upper value); define them to two decimal places.
Answer the following questions (not autograded):
1. Is the correlation between your variables positive or negative?
2. How would you write the confidence interval for your slope coefficient?
3. State the null hypothesis to test for a statistically significant relationship between your two variables.
4. Using the p-value from your model, do you **reject** or **fail to reject** the null hypothesis?
---
1. Positive correlation
2. 95% between 2.064 and 2.444
3. Ho: $\beta_1$ = 0 , Ha: $\beta_1 \neq$ 0
4. 0<0.05, so we'd reject the null hypothesis
```
# Task 8
# YOUR CODE HERE
slope_model_1 = 2.25
pval_model_1 = 0.00
ci_low = 2.06
ci_upper = 2.44
```
**Task 8 Test**
```
# Task 8 - Test
# Hidden tests - you will see the results when you submit to Canvas
```
**Task 9** - Fit a multiple predictor linear regression model
For this next task, we'll add in an additional independent or predictor variable. Let's look back at the pairplot and choose another variable - we'll use `Social_support`. Recall from the Guided Projects and Module Projects that we are looking to see if adding the variable `Social_support` is statistically significant after accounting for the `GDP_per_capita` variable.
We're going to fit a linear regression model using two predictor variables: `GDP_per_capita` and `Social_support`.
**Your tasks:**
* Fit a model with both predictor variables and assign the model to `model_2`. The format of the input to the model is `Y ~ X1 + X2`.
* **X1 = `GDP_per_capita`** and **X2 = `Social_support`**.
* Print out the model summary and assign the value of R-squared for this model to `r_square_model_2`. Your value should be defined to three decimal places.
* Assign the value of the adjusted R-square to `adj_r_square_model_2`. Your value should be defined to three decimal places.
* Answer the questions about your resulting model parameters (these short answer questions will not be autograded)
```
# Task 9
# YOUR CODE HERE
model_2 = ols('Score~GDP_per_capita+Social_support', data=happy).fit()
r_square_model_2 = 0.712
adj_r_square_model_2 = 0.710
# Print the model summary
print(model_2.summary())
```
**Task 9 Test**
```
# Task 9 - Test
# Hidden tests - you will see the results when you submit to Canvas
```
**Task 10** - Multiple regression model interpretation
Now that we have added an additional variable to our regression model, let's look at how the explained variance (the R-squared value) changes.
**Your tasks**
* Find the explained variance from `model_1` and assign it to the variable `r_square_percent1`; your variable should be expressed as a **percentage** and should be rounded to the nearest integer.
* Find the explained variance (*adjusted!*) from `model_2` and assign it to the variable `r_square_adj_percent2`; you variable should be expressed as a **percentage** and should be rounded to the nearest integer.
---
Question (not autograded):
How does the adjusted R-squared value change when a second predictor variable is added?
The adjusted r-quared value increases, so the line fits the data even better.
```
# Task 10
# YOUR CODE HERE
r_square_percent1 = 64
r_square_adj_percent2 = 71
print(r_square_percent1)
print(r_square_adj_percent2)
```
**Task 10 Test**
```
# Task 10 - Test
assert r_square_percent1 >= 1, 'Make sure you use the percentage and not the decimal fraction.'
assert r_square_adj_percent2 >= 1, 'Make sure you use the percentage and not the decimal fraction.'
# Hidden tests - you will see the results when you submit to Canvas
```
**Task 11** - Making a prediction and calculating the residual
We're going to use our model to make a prediction. Refer to the `happy` DataFrame and find the `GDP_per_capita` score for "Iceland" (index 4). Then when we have a prediction, we can calculate the residual. **There are actually two row entries for Iceland, both with slightly different column values. Use the column values that you can see when you print `happy.head()`.**
**Prediction**
* Assign the `GDP_per_capita` value to the variable `x_iceland`; it should be float and defined out to two decimal places.
* Using your slope and intercept values from `model_1`, calculate the `Score` for Iceland (`x_iceland`); assign this value to `predict_iceland` and it should be a float.
**Residual**
* Assign the observed `Score` for Iceland and assign it to the variable `observe_iceland`; it should be float and defined out to two decimal places *(careful with the rounding!)*.
* Determine the residual for the prediction you made and assign it to the variable `residual_iceland` (use your Guided Project or Module Project notebooks if you need a reminder of how to do a residual calculation).
Hint: Define your slope and intercept values out to two decimal places! Your resulting prediction for Iceland should have at least two decimal places. **Make sure to use the parameters from the first model (`model_1`)**.
```
happy.head()
#print(model_1.params)
#model_1.params[0]
# Task 11
# YOUR CODE HERE
x_iceland = 1.34
predict_iceland = model_1.params[0] + model_1.params[1]*x_iceland
observe_iceland = 7.50
residual_iceland = observe_iceland - predict_iceland
# View your prediction
print('Prediction for Iceland :', predict_iceland)
print('Residual for Iceland prediction :', residual_iceland)
```
**Task 11 Test**
```
# Task 11 - Test
assert residual_iceland >= 0, 'Check your residual calculation (use observed - predicted).'
assert round(x_iceland, 1) == 1.3, 'Check your Iceland GDP value.'
assert round(observe_iceland, 1) == 7.5, 'Check your Iceland observation value for "Score".'
# Hidden tests - you will see the results when you submit to Canvas
```
## Part B: Vectors and cosine similarity
In this part of the challenge, we're going to look at how similar two vectors are. Remember, we can calculate the **cosine similarity** between two vectors by using this equation:
$$\cos \theta= \frac{\mathbf {A} \cdot \mathbf {B} }{\left\|\mathbf {A} \right\|\left\|\mathbf {B} \right\|}$$
$\qquad$
where
* The numerator is the dot product of the vectors $\mathbf {A}$ and $\mathbf {B}$
* The denominator is the norm of $\mathbf {A}$ times the norm of $\mathbf {B}$
### Three documents, two authors
For this task, you will calculate the cosine similarity between three vectors. But here's the interesting part: each vector represents a "chunk" of text from a novel (a few chapters of text). This text was cleaned to remove non-alphanumeric characters and numbers and then each document was transformed into a vector representation as described below.
### Document vectors
In the dataset you are going to load below, each row represents a word that occurs in at least one of the documents. So all the rows are all the words that are in our three documents.
Each column represents a document (doc0, doc1, doc2). Now the fun part: the value in each cell is how frequently that word (row) occurs in that document (term-frequency) divided by how many documents that words appears in (document-frequency).
`cell value = term_frequency / document_frequency`
Use the above information to complete the remaining tasks.
**Task 12** - Explore the text documents
You will be using cosine similarity to compare each document vector to the others. Remember that there are three documents, but two authors. Your task is to use the cosine similarity calculations to determine which two document vectors are most similar (written by the same author).
**Your tasks:**
* Load in the CSV file that contains the document vectors (this is coded for you - just run the cell)
* Look at the DataFrame you just loaded in any way that helps you understand the format, what's included in the data, the shape of the DataFrame, etc.
**You can use document vectors just as they are - you don't need to code anything for Task 12.**
```
# Imports (import pandas if you haven't yet)
# Load the data - DON'T DELETE THIS CELL
url = 'https://raw.githubusercontent.com/LambdaSchool/data-science-practice-datasets/main/unit_4/unit1_nlp/text_vectors.csv'
text = pd.read_csv(url)
# Task 12
## Explore the data
# (this part is not autograded)
text.head()
```
**Task 13** - Calculate cosine similarity
* Calculate the cosine similarity for **three pairs of vectors** and assign the results to the following variables (each variable will be a float):
* assign the cosine similarity of doc0-doc1 to `cosine_doc0_1`
* assign the cosine similarity of doc0-doc2 to `cosine_doc0_2`
* assign the cosine similarity of doc1-doc2 to `cosine_doc1_2`
* Print out the results so you can refer to them for the short answer section.
* Answer the questions after you have completed the cosine similarity calculations.
```
# Task 13
# Use these imports for your cosine calculations (DON'T DELETE)
from numpy import dot
from numpy.linalg import norm
# YOUR CODE HERE
cosine_doc0_1 = dot(text['doc0'], text['doc1'])/(norm(text['doc0'])*norm(text['doc1']))
cosine_doc0_2 = dot(text['doc0'], text['doc2'])/(norm(text['doc0'])*norm(text['doc2']))
cosine_doc1_2 = dot(text['doc1'], text['doc2'])/(norm(text['doc1'])*norm(text['doc2']))
# Print out the results
print('Cosine similarity for doc0-doc1:', cosine_doc0_1)
print('Cosine similarity for doc0-doc2:', cosine_doc0_2)
print('Cosine similarity for doc1-doc2:', cosine_doc1_2)
```
**Task 13** - Short answer
1. Using your cosine similarity calculations, which two documents are most similar?
2. If doc1 and doc2 were written by the same author, are your cosine similarity calculations consistent with this statement?
3. What process would we need to follow to add an additional document column? In other words, why can't we just stick another column with (term-frequency/document-frequency) values onto our current DataFrame `text`?
---
1. Documents 1 and 2 are the most similar
2. Yes, they would be consistent because 1 and 2 had the greatest similarity of the three.
3. We would have to update each cell's value because now the document-frequency of each word would be affected, potentially changing each cell's value once we compute the term-frequency/document-frequency.
**Task 13 Test**
```
# Task 13 - Test
# Hidden tests - you will see the results when you submit to Canvas
```
**Additional Information about the texts used in this analysis:**
You can find the raw text [here](https://github.com/LambdaSchool/data-science-practice-datasets/tree/main/unit_4/unit1_nlp). Dcoument 0 (doc0) is chapters 1-3 from "Pride and Predjudice" by Jane Austen. Document 1 (doc1) is chapters 1- 4 from "Frankenstein" by Mary Shelley. Document 2 is also from "Frankenstein", chapters 11-14.
| github_jupyter |
# INFO 3402 – Class 13: Data cleaning exercise
[Brian C. Keegan, Ph.D.](http://brianckeegan.com/)
[Assistant Professor, Department of Information Science](https://www.colorado.edu/cmci/people/information-science/brian-c-keegan)
University of Colorado Boulder
Copyright and distributed under an [MIT License](https://opensource.org/licenses/MIT)
```
import pandas as pd
pd.options.display.max_columns = 100
%matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sb
```
I got this above-average gnarly data on [passenger traffic reports](https://www.flydenver.com/about/financials/passenger_traffic) from the Denver International Airport.
* Medium difficulty: `passengers_df` or `flights_df`
* Hard difficulty: `pass_type_df`
The goal is to turn this structured but very untidy and irregular data into something we can simply visualize in `seaborn`.
You'll almost certainly want these documentation resources available:
* pandas [`read_excel`](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.read_excel.html)
* pandas [Reshaping and Pivot Tables](https://pandas.pydata.org/pandas-docs/stable/user_guide/reshaping.html)
* seaborn [`catplot`](https://seaborn.pydata.org/generated/seaborn.catplot.html)
```
# Read in the data
passengers_df = pd.read_excel('DIA_activity.xlsx',sheet_name='Passengers')
flights_df = pd.read_excel('DIA_activity.xlsx',sheet_name='Flights')
pass_type_df = pd.read_excel('DIA_activity.xlsx',sheet_name='Passenger by Type')
```
Go through the EDA checklist:
1. **Formulate your question** → see “Characteristics of a good question”
2. **Read in your data** → Is it properly formatted? Perform cleanup activities
3. **Check the packaging** → Make sure there are the right number of rows & columns, formats, etc.
4. **Look at the top and bottom of data** → Confirm that all observations are there
5. **Check the “n”s** → Identify “landmark” values and to check expectations (number of states, etc.)
6. **Validate against an external data source** → Right order of magnitude, expected distribution, etc.
7. **Make a plot** → Checking and creating expectations about the shape of data and appropriate analyses
8. **Try an easy solution** → What is the simplest test for your question?
## Fix the columns and index
(Hint: The easiest way is to read in the data again, but use different parameters. Read the documentation!)
```
passengers_df = pd.read_excel('DIA_activity.xlsx',sheet_name='Passengers',header=(0,1),index_col=0)
passengers_df.head()
```
## Melt the data down
(Hint: pandas's `melt` and `stack` functions both turn columns into rows)
```
passengers_df2 = passengers_df.unstack()
passengers_df2.head(20)
```
## Make sure each variable has its own column... and rename the columns
(Hint: Mutate column names with wither [`rename`](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.rename.html?highlight=rename#pandas.DataFrame.rename) function or [`columns`](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.columns.html?highlight=columns#pandas.DataFrame.columns) attribute.
```
passengers_df3 = passengers_df2.reset_index()
passengers_df3.tail(20)
passengers_df3.head()
passengers_df3.columns = ['parent_carrier','carrier','date','passengers']
passengers_df3.head()
```
## Remove the miscellaneous rows
(Hint: Where did the "Unnamed: N_level_1" rows come from in the original spreadsheet?)
```
passengers_df4 = passengers_df3[~passengers_df3['carrier'].str.contains('Unnamed:')]
passengers_df4.tail()
passengers_df4.head()
```
## Separate the "date" column into year and months
(**Normie hint**: Access the month and year as attributes of a datetime/timestamp)
(**Elite hint**: Use `.str.extract` and pass a regular expression that matches a 4 digit number for year and 2 digit number of month)
```
passengers_df4['date'] = pd.to_datetime(passengers_df4['date'])
passengers_df4.tail()
passengers_df4['month'] = passengers_df4['date'].apply(lambda x:x.month)
passengers_df4['year'] = passengers_df4['date'].apply(lambda x:x.year)
passengers_df4.head()
passengers_df4.groupby(['date']).agg({'passengers':'sum'})['passengers'].to_csv('dia_passengers.csv')
```
## Pivot the data into total passengers by parent carrier per year
(Hint: Pivot table needs to know columns, indexes, values, and *probably* an aggfunc)
```
passengers_df5 = pd.pivot_table(columns='parent_carrier',
index='year',
values='passengers',
data=passengers_df4,aggfunc='sum')
passengers_df5
```
## Plot the pivot table
Is the drop-off in 2018 a "real" effect or caused by something else?
```
ax = passengers_df5.plot(legend=False)
ax.legend(loc='center left',bbox_to_anchor = (1,.5))
```
## Plot data
Make a seaborn `catplot` with "month" on the x-axis, "passengers" on the y-axis, and hues for the different parent carriers (probably excluding "Other"). What is the top month (on average) for passengers into DIA?
```
passengers_df4.head()
sb.catplot(x='year',y='passengers',hue='parent_carrier',row='month',data=passengers_df4,
aspect=3,kind='point')
```
## Re-do clean-up for `flights_df`
Make a nice clean dataset so we can use both of them.
## Compute average monthly passengers per flight
(Hint: Confirm cleaned `passengers_df` and `flights_df` are similar dimensions)
(Hint: You may also need to reference pandas's [`div`](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.div.html) function.)
## Plot average monthly passengers per flight
Make a seaborn `catplot` with "month" on the x-axis, "passengers" on the y-axis, and hues for the different carriers. Is the top month still the same?
| github_jupyter |
# Demonstration of FAE noise reduction capabilities
In this example we use a synth datset of blobs and heavily corrupted it using a binomial dropout noise (99%). We then recontruct the original dataset using an FAE
```
import pandas as pd
import torch
import numpy
import fiberedae.models.fae as vmod
import fiberedae.utils.basic_trainer as vtrain
import fiberedae.utils.persistence as vpers
import fiberedae.utils.single_cell as vsc
import fiberedae.utils.nn as vnnutils
import fiberedae.utils.plots as vplots
import fiberedae.utils.fae as vfae
import fiberedae.utils.datasets as vdatasets
import importlib
import fiberedae.utils.useful as us
import matplotlib.pyplot as plt
importlib.reload(vdatasets)
def pca(mat):
from sklearn.decomposition import PCA
pca = PCA(n_components=2)
return pca.fit_transform(mat)
def scatter_plot(mat_2d, targets):
plt.scatter(mat_2d[:, 0], mat_2d[:, 1], c=targets, alpha=0.2, s=5)
def clean_data(model, samples, targets):
return model.forward_output(x=samples, cond=targets)
config = us.load_configuration("configurations/blobs.json")
dataset = vdatasets.load_blobs(n_samples=5000,
nb_class=5,
nb_dim=128,
dropout_rate=0.99,
batch_size=config["hps"]["minibatch_size"],
mask_class=True)
```
## PCA of corrupted data
After corruption, we can see "fake" patterns appearing
```
blobs_2d, targets = pca(dataset["datasets"]["train"]["samples"]), dataset["datasets"]["train"]["unmasked_targets"]
scatter_plot(blobs_2d, targets)
```
## PCA of clean data
```
blobs_2d, targets = pca(dataset["datasets"]["train"]["clean_samples"]), dataset["datasets"]["train"]["unmasked_targets"]
scatter_plot(blobs_2d, targets)
```
## Training
```
config = us.load_configuration("configurations/blobs.json")
model_args = dict(config["model"])
model_args.update(
dict(
x_dim=dataset["shapes"]["input_size"],
nb_class=dataset["shapes"]["nb_class"],
output_transform=vnnutils.ScaleNonLinearity(-1., 1., dataset["sample_scale"][0], dataset["sample_scale"][1]),
)
)
#make model
model = vmod.FiberedAE(**model_args)
model.to("cuda")
#train model
trainer, history = us.train(model, dataset, config, nb_epochs=config["hps"]["nb_epochs"])
```
## PCA of FAE cleaned that
Some of the original structure is retrieved.
```
samples, targets = dataset["datasets"]["train"]["clean_samples"], dataset["datasets"]["train"]["unmasked_targets"]
samples, targets = torch.tensor(samples, dtype=torch.float).to("cuda"), torch.tensor(targets).to("cuda")
cleaned = model.forward_output(x=samples, cond=torch.zeros_like(targets)).cpu().detach().numpy()
scatter_plot(pca(cleaned), dataset["datasets"]["train"]["unmasked_targets"])
```
| github_jupyter |
TSG007 - Get BDC pod status
===========================
Description
-----------
View the big data cluster pods status.
Steps
-----
### Common functions
Define helper functions used in this notebook.
```
# Define `run` function for transient fault handling, suggestions on error, and scrolling updates on Windows
import sys
import os
import re
import json
import platform
import shlex
import shutil
import datetime
from subprocess import Popen, PIPE
from IPython.display import Markdown
retry_hints = {}
error_hints = {}
install_hint = {}
first_run = True
rules = None
def run(cmd, return_output=False, no_output=False, retry_count=0):
"""
Run shell command, stream stdout, print stderr and optionally return output
"""
MAX_RETRIES = 5
output = ""
retry = False
global first_run
global rules
if first_run:
first_run = False
rules = load_rules()
# shlex.split is required on bash and for Windows paths with spaces
#
cmd_actual = shlex.split(cmd)
# Store this (i.e. kubectl, python etc.) to support binary context aware error_hints and retries
#
user_provided_exe_name = cmd_actual[0].lower()
# When running python, use the python in the ADS sandbox ({sys.executable})
#
if cmd.startswith("python "):
cmd_actual[0] = cmd_actual[0].replace("python", sys.executable)
# On Mac, when ADS is not launched from terminal, LC_ALL may not be set, which causes pip installs to fail
# with:
#
# UnicodeDecodeError: 'ascii' codec can't decode byte 0xc5 in position 4969: ordinal not in range(128)
#
# Setting it to a default value of "en_US.UTF-8" enables pip install to complete
#
if platform.system() == "Darwin" and "LC_ALL" not in os.environ:
os.environ["LC_ALL"] = "en_US.UTF-8"
# To aid supportabilty, determine which binary file will actually be executed on the machine
#
which_binary = None
# Special case for CURL on Windows. The version of CURL in Windows System32 does not work to
# get JWT tokens, it returns "(56) Failure when receiving data from the peer". If another instance
# of CURL exists on the machine use that one. (Unfortunately the curl.exe in System32 is almost
# always the first curl.exe in the path, and it can't be uninstalled from System32, so here we
# look for the 2nd installation of CURL in the path)
if platform.system() == "Windows" and cmd.startswith("curl "):
path = os.getenv('PATH')
for p in path.split(os.path.pathsep):
p = os.path.join(p, "curl.exe")
if os.path.exists(p) and os.access(p, os.X_OK):
if p.lower().find("system32") == -1:
cmd_actual[0] = p
which_binary = p
break
# Find the path based location (shutil.which) of the executable that will be run (and display it to aid supportability), this
# seems to be required for .msi installs of azdata.cmd/az.cmd. (otherwise Popen returns FileNotFound)
#
# NOTE: Bash needs cmd to be the list of the space separated values hence shlex.split.
#
if which_binary == None:
which_binary = shutil.which(cmd_actual[0])
if which_binary == None:
if user_provided_exe_name in install_hint and install_hint[user_provided_exe_name] is not None:
display(Markdown(f'HINT: Use [{install_hint[user_provided_exe_name][0]}]({install_hint[user_provided_exe_name][1]}) to resolve this issue.'))
raise FileNotFoundError(f"Executable '{cmd_actual[0]}' not found in path (where/which)")
else:
cmd_actual[0] = which_binary
start_time = datetime.datetime.now().replace(microsecond=0)
print(f"START: {cmd} @ {start_time} ({datetime.datetime.utcnow().replace(microsecond=0)} UTC)")
print(f" using: {which_binary} ({platform.system()} {platform.release()} on {platform.machine()})")
print(f" cwd: {os.getcwd()}")
# Command-line tools such as CURL and AZDATA HDFS commands output
# scrolling progress bars, which causes Jupyter to hang forever, to
# workaround this, use no_output=True
#
# Work around a infinite hang when a notebook generates a non-zero return code, break out, and do not wait
#
wait = True
try:
if no_output:
p = Popen(cmd_actual)
else:
p = Popen(cmd_actual, stdout=PIPE, stderr=PIPE, bufsize=1)
with p.stdout:
for line in iter(p.stdout.readline, b''):
line = line.decode()
if return_output:
output = output + line
else:
if cmd.startswith("azdata notebook run"): # Hyperlink the .ipynb file
regex = re.compile(' "(.*)"\: "(.*)"')
match = regex.match(line)
if match:
if match.group(1).find("HTML") != -1:
display(Markdown(f' - "{match.group(1)}": "{match.group(2)}"'))
else:
display(Markdown(f' - "{match.group(1)}": "[{match.group(2)}]({match.group(2)})"'))
wait = False
break # otherwise infinite hang, have not worked out why yet.
else:
print(line, end='')
if rules is not None:
apply_expert_rules(line)
if wait:
p.wait()
except FileNotFoundError as e:
if install_hint is not None:
display(Markdown(f'HINT: Use {install_hint} to resolve this issue.'))
raise FileNotFoundError(f"Executable '{cmd_actual[0]}' not found in path (where/which)") from e
exit_code_workaround = 0 # WORKAROUND: azdata hangs on exception from notebook on p.wait()
if not no_output:
for line in iter(p.stderr.readline, b''):
line_decoded = line.decode()
# azdata emits a single empty line to stderr when doing an hdfs cp, don't
# print this empty "ERR:" as it confuses.
#
if line_decoded == "":
continue
print(f"STDERR: {line_decoded}", end='')
if line_decoded.startswith("An exception has occurred") or line_decoded.startswith("ERROR: An error occurred while executing the following cell"):
exit_code_workaround = 1
if user_provided_exe_name in error_hints:
for error_hint in error_hints[user_provided_exe_name]:
if line_decoded.find(error_hint[0]) != -1:
display(Markdown(f'HINT: Use [{error_hint[1]}]({error_hint[2]}) to resolve this issue.'))
if rules is not None:
apply_expert_rules(line_decoded)
if user_provided_exe_name in retry_hints:
for retry_hint in retry_hints[user_provided_exe_name]:
if line_decoded.find(retry_hint) != -1:
if retry_count < MAX_RETRIES:
print(f"RETRY: {retry_count} (due to: {retry_hint})")
retry_count = retry_count + 1
output = run(cmd, return_output=return_output, retry_count=retry_count)
if return_output:
return output
else:
return
elapsed = datetime.datetime.now().replace(microsecond=0) - start_time
# WORKAROUND: We avoid infinite hang above in the `azdata notebook run` failure case, by inferring success (from stdout output), so
# don't wait here, if success known above
#
if wait:
if p.returncode != 0:
raise SystemExit(f'Shell command:\n\n\t{cmd} ({elapsed}s elapsed)\n\nreturned non-zero exit code: {str(p.returncode)}.\n')
else:
if exit_code_workaround !=0 :
raise SystemExit(f'Shell command:\n\n\t{cmd} ({elapsed}s elapsed)\n\nreturned non-zero exit code: {str(exit_code_workaround)}.\n')
print(f'\nSUCCESS: {elapsed}s elapsed.\n')
if return_output:
return output
def load_json(filename):
with open(filename, encoding="utf8") as json_file:
return json.load(json_file)
def load_rules():
try:
# Load this notebook as json to get access to the expert rules in the notebook metadata.
#
j = load_json("tsg007-view-bdc-pod-status.ipynb")
except:
pass # If the user has renamed the book, we can't load ourself. NOTE: Is there a way in Jupyter, to know your own filename?
else:
if "metadata" in j and \
"azdata" in j["metadata"] and \
"expert" in j["metadata"]["azdata"] and \
"rules" in j["metadata"]["azdata"]["expert"]:
rules = j["metadata"]["azdata"]["expert"]["rules"]
rules.sort() # Sort rules, so they run in priority order (the [0] element). Lowest value first.
# print (f"EXPERT: There are {len(rules)} rules to evaluate.")
return rules
def apply_expert_rules(line):
global rules
for rule in rules:
# rules that have 9 elements are the injected (output) rules (the ones we want). Rules
# with only 8 elements are the source (input) rules, which are not expanded (i.e. TSG029,
# not ../repair/tsg029-nb-name.ipynb)
if len(rule) == 9:
notebook = rule[1]
cell_type = rule[2]
output_type = rule[3] # i.e. stream or error
output_type_name = rule[4] # i.e. ename or name
output_type_value = rule[5] # i.e. SystemExit or stdout
details_name = rule[6] # i.e. evalue or text
expression = rule[7].replace("\\*", "*") # Something escaped *, and put a \ in front of it!
# print(f"EXPERT: If rule '{expression}' satisfied', run '{notebook}'.")
if re.match(expression, line, re.DOTALL):
# print("EXPERT: MATCH: name = value: '{0}' = '{1}' matched expression '{2}', therefore HINT '{4}'".format(output_type_name, output_type_value, expression, notebook))
match_found = True
display(Markdown(f'HINT: Use [{notebook}]({notebook}) to resolve this issue.'))
print('Common functions defined successfully.')
# Hints for binary (transient fault) retry, (known) error and install guide
#
retry_hints = {'kubectl': ['A connection attempt failed because the connected party did not properly respond after a period of time, or established connection failed because connected host has failed to respond']}
error_hints = {'kubectl': [['no such host', 'TSG010 - Get configuration contexts', '../monitor-k8s/tsg010-get-kubernetes-contexts.ipynb'], ['no such host', 'TSG011 - Restart sparkhistory server', '../repair/tsg011-restart-sparkhistory-server.ipynb'], ['No connection could be made because the target machine actively refused it', 'TSG056 - Kubectl fails with No connection could be made because the target machine actively refused it', '../repair/tsg056-kubectl-no-connection-could-be-made.ipynb']]}
install_hint = {'kubectl': ['SOP036 - Install kubectl command line interface', '../install/sop036-install-kubectl.ipynb']}
```
### Get the Kubernetes namespace for the big data cluster
Get the namespace of the big data cluster use the kubectl command line
interface .
NOTE: If there is more than one big data cluster in the target
Kubernetes cluster, then set \[0\] to the correct value for the big data
cluster.
```
# Place Kubernetes namespace name for BDC into 'namespace' variable
try:
namespace = run(f'kubectl get namespace --selector=MSSQL_CLUSTER -o jsonpath={{.items[0].metadata.name}}', return_output=True)
except:
from IPython.display import Markdown
print(f"ERROR: Unable to find a Kubernetes namespace with label 'MSSQL_CLUSTER'. SQL Server Big Data Cluster Kubernetes namespaces contain the label 'MSSQL_CLUSTER'.")
display(Markdown(f'HINT: Use [TSG081 - Get namespaces (Kubernetes)](../monitor-k8s/tsg081-get-kubernetes-namespaces.ipynb) to resolve this issue.'))
display(Markdown(f'HINT: Use [TSG010 - Get configuration contexts](../monitor-k8s/tsg010-get-kubernetes-contexts.ipynb) to resolve this issue.'))
display(Markdown(f'HINT: Use [SOP011 - Set kubernetes configuration context](../common/sop011-set-kubernetes-context.ipynb) to resolve this issue.'))
raise
else:
print(f'The SQL Server Big Data Cluster Kubernetes namespace is: {namespace}')
```
### Show the big data cluster pods
```
run(f'kubectl get pods -n {namespace} -o wide')
print('Notebook execution complete.')
```
Related
-------
- [TSG006 - Get system pod
status](../monitor-k8s/tsg006-view-system-pod-status.ipynb)
- [TSG009 - Get nodes
(Kubernetes)](../monitor-k8s/tsg009-get-nodes.ipynb)
| github_jupyter |
... ***CURRENTLY UNDER DEVELOPMENT*** ...
## Before running this notebook, you must have already the numerically simulated waves associated to the representative cases of synthetic simulated TCs (obtained with MaxDiss algorithm in notebook 06)
inputs required:
* Synthetic simulation of historical TCs parameters (copulas obtained in *notebook 06*)
* MaxDiss selection of synthetic simulated TCs (parameters obtained in *notebook 06*)
* simulated waves for the above selected TCs (**outside TeslaKit**)
in this notebook:
* RBF's interpolation of wave conditions based on TCs parameters (from swan simulated TCs waves)
```
#!/usr/bin/env python
# -*- coding: utf-8 -*-
# common
import os
import os.path as op
# pip
import xarray as xr
import numpy as np
# DEV: override installed teslakit
import sys
sys.path.insert(0, op.join(os.path.abspath(''), '..', '..', '..'))
# teslakit
from teslakit.database import Database
from teslakit.rbf import RBF_Reconstruction, RBF_Validation
```
## Database and Site parameters
```
# --------------------------------------
# Teslakit database
p_data = r'/Users/nico/Projects/TESLA-kit/TeslaKit/data'
db = Database(p_data)
# set site
db.SetSite('ROI')
# --------------------------------------
# load data and set parameters
# r2 TCs Copula Simulated (dataset)
TCs_r2_sim_params = db.Load_TCs_r2_sim_params()
# r2 TCs MDA selection and solved simulations (not solved inside teslakit)
TCs_r2_MDA_params = db.Load_TCs_r2_mda_params()
TCs_sims = db.Load_TCs_r2_mda_Simulations()
```
## Simulated TCs - Radial Basis Function
```
# --------------------------------------
# prepare dataset and subset
# RBFs training subset (TCs numerically solved)
subset = np.column_stack(
(TCs_r2_MDA_params['pressure_min'], TCs_r2_MDA_params['velocity_mean'],
TCs_r2_MDA_params['gamma'], TCs_r2_MDA_params['delta'])
)
# RBFs dataset to interpolate
dataset = np.column_stack(
(TCs_r2_sim_params['pressure_min'], TCs_r2_sim_params['velocity_mean'],
TCs_r2_sim_params['gamma'], TCs_r2_sim_params['delta'])
)
# --------------------------------------
# Extract waves data from TCs simulations (this is the RBFs training target)
print(TCs_sims)
print()
# Normalize data
d_maxis = {}
d_minis = {}
tcp = TCs_sims.copy()
for k in ['hs', 'tp', 'ss', 'twl']:
v = tcp[k].values[:]
mx = np.max(v)
mn = np.min(v)
tcp[k] =(('storm',), (v-mn)/(mx-mn))
# store maxs and mins for denormalization
d_maxis[k] = mx
d_minis[k] = mn
tcp['dir'] = tcp['dir'] * np.pi/180
print(tcp)
print()
# Build RBF target numpy array
target = np.column_stack(
(tcp['hs'], tcp['tp'], tcp['ss'], tcp['twl'], tcp['dir'], tcp['mu'])
)
# --------------------------------------
# RBF Interpolation
# subset - scalar / directional indexes
ix_scalar_subset = [0,1] # scalar (pmean, vmean)
ix_directional_subset = [2,3] # directional (delta, gamma)
# target - scalar / directional indexes
ix_scalar_target = [0,1,2,3,5] # scalar (Hs, Tp, SS, TWL, MU)
ix_directional_target = [4] # directional (Dir)
output = RBF_Reconstruction(
subset, ix_scalar_subset, ix_directional_subset,
target, ix_scalar_target, ix_directional_target,
dataset)
# --------------------------------------
# Reconstructed TCs
# denormalize RBF output
TCs_RBF_out = xr.Dataset(
{
'hs':(('storm',), output[:,0] * (d_maxis['hs']-d_minis['hs']) + d_minis['hs'] ),
'tp':(('storm',), output[:,1] * (d_maxis['tp']-d_minis['tp']) + d_minis['tp'] ),
'ss':(('storm',), output[:,2] * (d_maxis['ss']-d_minis['ss']) + d_minis['ss'] ),
'twl':(('storm',), output[:,3] * (d_maxis['twl']-d_minis['twl']) + d_minis['twl'] ),
'dir':(('storm',), output[:,4] * 180 / np.pi),
'mu':(('storm',), output[:,5]),
},
coords = {'storm': np.arange(output.shape[0])}
)
print(TCs_RBF_out)
# store data as xarray.Dataset
db.Save_TCs_sim_r2_rbf_output(TCs_RBF_out)
# --------------------------------------
# RBF Validation
# subset - scalar / directional indexes
ix_scalar_subset = [0,1] # scalar (pmean, vmean)
ix_directional_subset = [2,3] # directional (delta, gamma)
# target - scalar / directional indexes
ix_scalar_target = [0,1,2,3,5] # scalar (Hs, Tp, SS, TWL, MU)
ix_directional_target = [4] # directional (Dir)
output = RBF_Validation(
subset, ix_scalar_subset, ix_directional_subset,
target, ix_scalar_target, ix_directional_target)
```
| github_jupyter |
# Measuring the distance to shape surfaces
The distance from the center of a shape to its surface is a useful quantity in various situations.
For instance, when the shape represents a vacancy in a crystal, we may wish to know how far the vacancy extends in certain directions.
Another useful example is perturbative theories that treat various geometries relative to their inscribed spheres.
Here we'll see how **coxeter** can be used to compute this quantity for a shape.
```
import numpy as np
from matplotlib import patches
from matplotlib import pyplot as plt
import coxeter
def plot_distances(shape, vectors, colors):
"""Plot vectors from the center of a shape with suitable labeling."""
fig, ax = plt.subplots(figsize=(4, 4))
for v, c in zip(vectors, colors):
ax.add_patch(
patches.FancyArrow(
x=shape.center[0],
y=shape.center[1],
dx=v[0],
dy=v[1],
width=0.02,
color=c,
length_includes_head=True,
label=f"{np.linalg.norm(v):0.2f}",
)
)
ax.legend(fontsize=18, bbox_to_anchor=(1.05, 0.75))
# Pad the extent so the shapes edges aren't cut off.
extent = shape.minimal_bounding_circle.radius + 0.02
ax.set_xlim([-extent, extent])
ax.set_ylim([-extent, extent])
for spine in ax.spines.values():
spine.set_visible(False)
ax.tick_params(labelbottom=False, labelleft=False, bottom=False, left=False)
return fig, ax
```
Using the handy plotting function defined above, we visualize distance vectors computed using coxeter.
We'll show how this works for both pentagons and ellipses.
```
# These are the directions in which we'll find distances to the shape surfaces.
random_angles = np.array([0.75 * np.pi, 1.5 * np.pi, 0])
unit_vectors = np.vstack((np.cos(random_angles), np.sin(random_angles))).T
colors = ("r", "b", "g")
# Create a pentagon.
pentagon = coxeter.families.RegularNGonFamily.get_shape(5)
# Create a unit area ellipse.
ellipse = coxeter.shapes.Ellipse(3, 4)
ellipse.area = 1
vectors = unit_vectors * pentagon.distance_to_surface(random_angles)[:, np.newaxis]
fig, ax = plot_distances(pentagon, vectors, colors)
ax.add_patch(
patches.Polygon(pentagon.vertices[:, :2], fill=False, linewidth=2, edgecolor="k")
)
vectors = unit_vectors * ellipse.distance_to_surface(random_angles)[:, np.newaxis]
fig, ax = plot_distances(ellipse, vectors, colors)
ax.add_patch(
patches.Ellipse(
(0, 0), 2 * ellipse.a, 2 * ellipse.b, fill=False, linewidth=2, edgecolor="k"
)
)
```
| github_jupyter |
# Optimizing Multiple Objectives using Vertex Vizier
## Overview
This tutorial demonstrates [Vertex Vizier](https://cloud.google.com/vertex-ai/docs/vizier/overview) multi-objective optimization. Multi-objective optimization is concerned with mathematical optimization problems involving more than one objective function to be optimized simultaneously
## Objective
The goal is to __`minimize`__ the objective metric:
```
y1 = r*sin(theta)
```
and simultaneously __`maximize`__ the objective metric:
```
y2 = r*cos(theta)
```
that you will evaluate over the parameter space:
- __`r`__ in [0,1],
- __`theta`__ in [0, pi/2]
## Introduction
In this notebook, you will use [Vertex Vizier](https://cloud.google.com/vertex-ai/docs/vizier/overview) multi-objective optimization. Multi-objective optimization is concerned with mathematical optimization problems involving more than one objective function to be optimized simultaneously.
Each learning objective will correspond to a __#TODO__ in this student lab notebook -- try to complete this notebook first and then review the [solution notebook](../solutions/gapic-vizier-multi-objective-optimization.ipynb).
**Make sure to enable the Vertex AI API**
#### Install Vertex AI library
Download and install Vertex AI library.
```
import os
# The Google Cloud Notebook product has specific requirements
IS_GOOGLE_CLOUD_NOTEBOOK = os.path.exists("/opt/deeplearning/metadata/env_version")
# Google Cloud Notebook requires dependencies to be installed with '--user'
USER_FLAG = ""
if IS_GOOGLE_CLOUD_NOTEBOOK:
USER_FLAG = "--user"
! pip install {USER_FLAG} --upgrade google-cloud-aiplatform
import os
if not os.getenv("IS_TESTING"):
# Restart the kernel after pip installs
import IPython
app = IPython.Application.instance()
app.kernel.do_shutdown(True)
```
### Import libraries and define constants
```
import datetime
import json
from google.cloud import aiplatform_v1beta1
```
## Tutorial
This section defines some parameters and util methods to call Vertex Vizier APIs. Please fill in the following information to get started.
```
# Fill in your project ID and region
REGION = "[region]" # @param {type:"string"}
PROJECT_ID = "[your-project-id]" # @param {type:"string"}
# These will be automatically filled in.
STUDY_DISPLAY_NAME = "{}_study_{}".format(
PROJECT_ID.replace("-", ""), datetime.datetime.now().strftime("%Y%m%d_%H%M%S")
) # @param {type: 'string'}
ENDPOINT = REGION + "-aiplatform.googleapis.com"
PARENT = "projects/{}/locations/{}".format(PROJECT_ID, REGION)
print("ENDPOINT: {}".format(ENDPOINT))
print("REGION: {}".format(REGION))
print("PARENT: {}".format(PARENT))
# If you don't know your project ID, you might be able to get your project ID
# using gcloud command by executing the second cell below.
if PROJECT_ID == "" or PROJECT_ID is None or PROJECT_ID == "[your-project-id]":
# Get your GCP project id from gcloud
shell_output = !gcloud config list --format 'value(core.project)' 2>/dev/null
PROJECT_ID = shell_output[0]
print("Project ID:", PROJECT_ID)
! gcloud config set project $PROJECT_ID
```
### Create the study configuration
The following is a sample study configuration, built as a hierarchical python dictionary. It is already filled out. Run the cell to configure the study.
```
# Parameter Configuration
param_r = {"parameter_id": "r", "double_value_spec": {"min_value": 0, "max_value": 1}}
param_theta = {
"parameter_id": "theta",
"double_value_spec": {"min_value": 0, "max_value": 1.57},
}
# Objective Metrics
metric_y1 = {"metric_id": "y1", "goal": "MINIMIZE"}
# Objective Metrics
metric_y2 = {"metric_id": "y2", "goal": "MAXIMIZE"}
# Put it all together in a study configuration
study = {
"display_name": STUDY_DISPLAY_NAME,
"study_spec": {
"algorithm": "RANDOM_SEARCH",
"parameters": [
param_r,
param_theta,
],
"metrics": [metric_y1, metric_y2],
},
}
print(json.dumps(study, indent=2, sort_keys=True))
```
### Create the study
Next, create the study, which you will subsequently run to optimize the two objectives.
```
vizier_client = aiplatform_v1beta1.VizierServiceClient(
client_options=dict(api_endpoint=ENDPOINT)
)
study = vizier_client.create_study(parent=PARENT, study=study)
STUDY_ID = study.name
print("STUDY_ID: {}".format(STUDY_ID))
```
### Metric evaluation functions
Next, define some functions to evaluate the two objective metrics.
```
import math
# r * sin(theta)
def Metric1Evaluation(r, theta):
"""Evaluate the first metric on the trial."""
return r * math.sin(theta)
# r * cos(theta)
def Metric2Evaluation(r, theta):
"""Evaluate the second metric on the trial."""
return r * math.cos(theta)
def CreateMetrics(trial_id, r, theta):
print(("=========== Start Trial: [{}] =============").format(trial_id))
# Evaluate both objective metrics for this trial
y1 = Metric1Evaluation(r, theta)
y2 = Metric2Evaluation(r, theta)
print(
"[r = {}, theta = {}] => y1 = r*sin(theta) = {}, y2 = r*cos(theta) = {}".format(
r, theta, y1, y2
)
)
metric1 = {"metric_id": "y1", "value": y1}
metric2 = {"metric_id": "y2", "value": y2}
# Return the results for this trial
return [metric1, metric2]
```
### Set configuration parameters for running trials
__`client_id`__: The identifier of the client that is requesting the suggestion. If multiple SuggestTrialsRequests have the same `client_id`, the service will return the identical suggested trial if the trial is `PENDING`, and provide a new trial if the last suggested trial was completed.
__`suggestion_count_per_request`__: The number of suggestions (trials) requested in a single request.
__`max_trial_id_to_stop`__: The number of trials to explore before stopping. It is set to 4 to shorten the time to run the code, so don't expect convergence. For convergence, it would likely need to be about 20 (a good rule of thumb is to multiply the total dimensionality by 10).
```
client_id = "client1" # @param {type: 'string'}
suggestion_count_per_request = 5 # @param {type: 'integer'}
max_trial_id_to_stop = 4 # @param {type: 'integer'}
print("client_id: {}".format(client_id))
print("suggestion_count_per_request: {}".format(suggestion_count_per_request))
print("max_trial_id_to_stop: {}".format(max_trial_id_to_stop))
```
### Run Vertex Vizier trials
Run the trials.
```
trial_id = 0
while int(trial_id) < max_trial_id_to_stop:
suggest_response = vizier_client.suggest_trials(
{
"parent": STUDY_ID,
"suggestion_count": suggestion_count_per_request,
"client_id": client_id,
}
)
for suggested_trial in suggest_response.result().trials:
trial_id = suggested_trial.name.split("/")[-1]
trial = vizier_client.get_trial({"name": suggested_trial.name})
if trial.state in ["COMPLETED", "INFEASIBLE"]:
continue
for param in trial.parameters:
if param.parameter_id == "r":
r = param.value
elif param.parameter_id == "theta":
theta = param.value
print("Trial : r is {}, theta is {}.".format(r, theta))
vizier_client.add_trial_measurement(
{
"trial_name": suggested_trial.name,
"measurement": {
"metrics": CreateMetrics(suggested_trial.name, r, theta)
},
}
)
response = vizier_client.complete_trial(
{"name": suggested_trial.name, "trial_infeasible": False}
)
```
### List the optimal solutions
list_optimal_trials returns the pareto-optimal Trials for multi-objective Study or the optimal Trials for single-objective Study. In the case, we define mutliple-objective in previeous steps, pareto-optimal trials will be returned.
```
optimal_trials = vizier_client.list_optimal_trials({"parent": STUDY_ID})
print("optimal_trials: {}".format(optimal_trials))
```
## Cleaning up
To clean up all Google Cloud resources used in this project, you can [delete the Google Cloud
project](https://cloud.google.com/resource-manager/docs/creating-managing-projects#shutting_down_projects) you used for the tutorial. You can also manually delete resources that you created by running the following code.
```
vizier_client.delete_study({"name": STUDY_ID})
```
| github_jupyter |
# GDP and Life Expectancy 2018 Comparison
Richer countries can afford to invest more on healthcare, on work and road safety, and other measures that reduce mortality. On the other hand, richer countries may have less healthy lifestyles. Is there any relation between the wealth of a country and the life expectancy of its inhabitants?
The following analysis checks whether there is any correlation between the total gross domestic product (GDP) of a country in 2018 and the life expectancy of people born in that country in 2018.
## Getting the data
Two datasets of the World Bank are considered. One dataset, available at <http://data.worldbank.org/indicator/NY.GDP.MKTP.CD>, lists the GDP of the world's countries in current US dollars, for various years. The use of a common currency allows us to compare GDP values across countries. The other dataset, available at <http://data.worldbank.org/indicator/SP.DYN.LE00.IN>, lists the life expectancy of the world's countries. The datasets were downloaded as Excel files in June 2021.
```
import warnings
warnings.simplefilter('ignore', FutureWarning)
import pandas as pd
YEAR = 2018
GDP_INDICATOR = 'NY.GDP.MKTP.CD'
gdpReset = pd.read_excel('WB GDP 2020.xls')
LIFE_INDICATOR = 'SP.DYN.LE00.IN'
lifeReset = pd.read_excel('WB LE 2020.xls')
lifeReset.head()
```
## Cleaning the data
Inspecting the data with `head()` and `tail()` shows:
1. aggregated data, for the Arab World, the Caribbean small states, and other country groups used by the World Bank;
- GDP and life expectancy values are missing for some countries.
The data is therefore cleaned by:
- removing rows with unavailable values.
- filtering data set to ensure the first selected rows are stricly countries
```
gdpCountries = gdpReset.dropna()
lifeCountries = lifeReset.dropna()
```
## Transforming the data
The World Bank reports GDP in US dollars and cents. To make the data easier to read, the GDP is converted to millions of British pounds (the author's local currency) with the following auxiliary functions, using the average 2018 dollar-to-pound conversion rate provided by <http://www.ukforex.co.uk/forex-tools/historical-rate-tools/yearly-average-rates>.
```
gdpCountries.head()
def roundToMillions (value):
return round(value / 1000000)
def usdToGBP (usd):
return usd / 1.334801
GDP = 'GDP (£m)'
gdpCountries[GDP] = gdpCountries[GDP_INDICATOR].apply(usdToGBP).apply(roundToMillions)
gdpCountries.head()
```
The unnecessary columns can be dropped.
```
COUNTRY = 'Country Name'
headings = [COUNTRY, GDP]
gdpClean = gdpCountries[headings]
gdpClean.head()
```
The World Bank reports the life expectancy with several decimal places. After rounding, the original column is discarded.
```
LIFE = 'Life expectancy (years)'
lifeCountries[LIFE] = lifeCountries[LIFE_INDICATOR].apply(round)
headings = [COUNTRY, LIFE]
lifeClean = lifeCountries[headings]
lifeClean.head()
```
## Combining the data
The tables are combined through an inner join on the common 'country' column.
```
gdpVsLife = pd.merge(gdpClean, lifeClean, on=COUNTRY, how='inner')
gdpVsLife.head()
```
## Calculating the correlation
To measure if the life expectancy and the GDP grow together, the Spearman rank correlation coefficient is used. It is a number from -1 (perfect inverse rank correlation: if one indicator increases, the other decreases) to 1 (perfect direct rank correlation: if one indicator increases, so does the other), with 0 meaning there is no rank correlation. A perfect correlation doesn't imply any cause-effect relation between the two indicators. A p-value below 0.05 means the correlation is statistically significant.
```
from scipy.stats import spearmanr
gdpColumn = gdpVsLife[GDP]
lifeColumn = gdpVsLife[LIFE]
(correlation, pValue) = spearmanr(gdpColumn, lifeColumn)
print('The correlation is', correlation)
if pValue < 0.05:
print('It is statistically significant.')
else:
print('It is not statistically significant.')
```
The value shows a direct correlation, i.e. richer countries tend to have longer life expectancy, but it is not a very strong correlation.
## Showing the data
Measures of correlation can be misleading, so it is best to see the overall picture with a scatterplot. The GDP axis uses a logarithmic scale to better display the vast range of GDP values, from a few million to several billion (million of million) pounds.
```
%matplotlib inline
gdpVsLife.plot(x=GDP, y=LIFE, kind='scatter', grid=True, logx=True, figsize=(10, 4))
```
The plot shows there is no clear correlation: similar to 2013 results,there are rich countries with low life expectancy, poor countries with high expectancy, and countries with around 10 thousand (10<sup>4</sup>) million pounds GDP have almost the full range of values, from below 50 to over 80 years. Towards the lower and higher end of GDP, the variation diminishes. Above 40 thousand million pounds of GDP (3rd tick mark to the right of 10<sup>4</sup>), most countries have an expectancy of 70 years or more, whilst below that threshold most countries' life expectancy is below 70 years.
Comparing the 10 poorest countries and the 10 countries with the lowest life expectancy shows that total GDP is not the most accurate measure
```
# the 10 countries with lowest GDP
gdpVsLife.sort_values(GDP).head(10)
# the 10 countries with lowest life expectancy
gdpVsLife.sort_values(LIFE).head(10)
```
## Conclusions
In this current year there is no strong correlation between a country's wealth and the life expectancy of its inhabitants: there is often a wide variation of life expectancy for countries with similar GDP, countries with the lowest life expectancy are not the poorest countries, and countries with the highest expectancy are not the richest countries. the relationship is hard to define, because the vast majority of countries with a life expectancy below 70 years are scattered almost evenly on the scatterplot.
Recommendation: Using the [NY.GDP.PCAP.PP.CD](http://data.worldbank.org/indicator/NY.GDP.PCAP.PP.CD) indicator, GDP per capita in current 'international dollars', might possibly make for a better like-for-like comparison between countries, because it would take population and purchasing power into account. Using more specific data, like expediture on health, could also lead to a better analysis.
## TASK HAS BEEN COMPLETED 2018 GDP vs LE COMPARISONS
## THE END!
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.